code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nothing But NumPy: A 2-layer neural network
# Part of the blog ["Nothing but NumPy: Understanding & Creating neural networks with computational graphs from scratch"](https://medium.com/@rafayak/nothing-but-numpy-understanding-creating-neural-networks-with-computational-graphs-from-scratch-6299901091b0)- by [<NAME>](https://twitter.com/RafayAK)
#
#
# In this notebook we'll create a 2-layer nueral network (i.e. a single hidden and output layer) and train it on the XOR data.
#
#
# First, let's import NumPy, our layers and helper functions.
#
# _Feel free to look into the helper functions in the `utils` directory_
import numpy as np
from util.utilities import *
from Layers.LinearLayer import LinearLayer
from Layers.ActivationLayer import SigmoidLayer
# to show all the generated plots inline in the notebook
# %matplotlib inline
# #### The XOR data:
#
# 
# +
# This is our XOR gate data
X = np.array([
[0, 0],
[0, 1],
[1, 0],
[1, 1]
])
Y = np.array([
[0],
[1],
[1],
[0]
])
# -
# Let's set up training data. Recall, data needs to be in $(features \times \text{number_of_examples})$ shape. So, we need to transpose `X` and `Y`.
X_train = X.T
Y_train = Y.T
# Recall, our neural net architecture with a single hidden layer with 3 neurons from the blog. We'll recreate it here.
#
#
# 
# +
# define training constants
learning_rate = 1
number_of_epochs = 5000
np.random.seed(48) # set seed value so that the results are reproduceable
# (weights will now be initailzaed to the same pseudo-random numbers, each time)
# Our network architecture has the shape:
# (input)--> [Linear->Sigmoid] -> [Linear->Sigmoid] -->(output)
#------ LAYER-1 ----- define hidden layer that takes in training data
Z1 = LinearLayer(input_shape=X_train.shape, n_out=3, ini_type='xavier')
A1 = SigmoidLayer(Z1.Z.shape)
#------ LAYER-2 ----- define output layer that take is values from hidden layer
Z2= LinearLayer(input_shape=A1.A.shape, n_out=1, ini_type='xavier')
A2= SigmoidLayer(Z2.Z.shape)
# see what random weights and bias were selected and their shape
# print(Z2.params)
# print(Z2.params)
# -
# Now we can start the training loop.
# +
costs = [] # initially empty list, this will store all the costs after a certian number of epochs
# Start training
for epoch in range(number_of_epochs):
# ------------------------- forward-prop -------------------------
Z1.forward(X_train)
A1.forward(Z1.Z)
Z2.forward(A1.A)
A2.forward(Z2.Z)
# ---------------------- Compute Cost ----------------------------
cost, dA2 = compute_cost(Y=Y_train, Y_hat=A2.A)
# print and store Costs every 100 iterations.
if (epoch % 100) == 0:
print("Cost at epoch#{}: {}".format(epoch, cost))
costs.append(cost)
# ------------------------- back-prop ----------------------------
A2.backward(dA2)
Z2.backward(A2.dZ)
A1.backward(Z2.dA_prev)
Z1.backward(A1.dZ)
# ----------------------- Update weights and bias ----------------
Z2.update_params(learning_rate=learning_rate)
Z1.update_params(learning_rate=learning_rate)
# See what the final weights and bias are training
# print(Z2.params)
# print(Z2.params)
# -
# `predict` helper functionin the cell below returns three things:
# - `p`: predicted labels (output 1 if predictded output is greater than 0.5)
# - `probas`: raw probabilities (how sure the neural net thinks the output is 1, this is just Y_hat)
# - `accuracy`: the number of correct predictions from total predictions
#
# In the cell below we are not storing `probas`( pythonic way is to use `_` it its place)
# +
# see the ouptput predictions
predicted_outputs, _, accuracy = predict(X=X_train, Y=Y_train, Zs=[Z1, Z2], As=[A1, A2])
print("The predicted outputs:\n {}".format(predicted_outputs))
print("The accuracy of the model is: {}%".format(accuracy))
# -
# #### The learning curve
plot_learning_curve(costs=costs, learning_rate=learning_rate, total_epochs=number_of_epochs)
# #### The decision boundary
plot_decision_boundary(lambda x:predict_dec(Zs=[Z1, Z2], As=[A1, A2], X=x.T), X_train.T, Y_train.T)
# #### The shaded decision boundary
plot_decision_boundary_shaded(lambda x:predict_dec(Zs=[Z1, Z2], As=[A1, A2], X=x.T), X_train.T, Y_train.T)
# ## Bonus
#
# Try to reinitialize the weights to `ini_type=plain` and see what happens.
| Understanding_and_Creating_NNs/2_layer_toy_network_XOR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [py27]
# language: python
# name: Python [py27]
# ---
# * Spearmint for analogy reasoning
# * Gaussian LDA
# * Evaluate word analogy reasoning
# * evalutate topic models
# * find background noise
# * find word pairs
# + [markdown] heading_collapsed=true
# # Setup
# + hidden=true
# %matplotlib notebook
import itertools
import logging
from functools import partial
import gensim
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pnd
from sklearn.cluster import *
from sklearn.decomposition import PCA, RandomizedPCA
from sklearn.manifold import TSNE
from codecs import open
import gc
from knub.thesis.util import *
matplotlib.style.use('ggplot')
# + hidden=true
from IPython.core.display import HTML
HTML("""
<style>
div.text_cell_render p, div.text_cell_render ul, table.dataframe {
font-size:1.3em;
line-height:1.1em;
}
</style>
""")
# + [markdown] heading_collapsed=true
# # Preprocessing
# + hidden=true
MODEL = "../models/topic-models/topic.full.alpha-1-100.256-400.model"
#MODEL = "../models/topic-models/topic.256-400.first-2000.alpha-001.beta-001.model"
# + hidden=true
print "Load vectors"
vectors = load_skip_gram()
model = TopicModelLoader(MODEL, vectors)
print "Load topic probs"
df_topic_probs_full = model.load_topic_probs()
print "Load topics"
df_topics = model.load_topics()
print "Load topic similars"
df_topic_similars = model.load_all_topic_similars()
# + hidden=true
word_prob_lower_threshold = df_topic_probs_full["word-prob"].quantile(0.4)
word_prob_upper_threshold = df_topic_probs_full["word-prob"].quantile(0.99)
# + [markdown] heading_collapsed=true
# # Topic Probs Analysis
# + hidden=true
df_topic_probs = df_topic_probs_full[df_topic_probs_full["word"].apply(lambda w: w in model.topic_words)].copy()
# + [markdown] hidden=true
# word-prob does not sum to one, because we only write out frequent words
# + hidden=true
df_topic_probs_full["word-prob"].sum()
# + hidden=true
def topic_prob_difference_from_first_to(row, n):
s = sorted(row, reverse=True)
return s[0] - s[n - 1]
for diff in [2, 5, 50]:
column_name = "diff-" + str(diff)
df_topic_probs_full[column_name] = df_topic_probs_full[model.prob_columns].apply(
partial(topic_prob_difference_from_first_to, n=diff), axis=1)
# + [markdown] hidden=true
# ## Strength of topic prevalence
# + [markdown] hidden=true
# ### Against second best topic
# + hidden=true
plt.figure()
df_topic_probs_full["diff-2"].hist(bins=20)
# + [markdown] hidden=true
# ### Against fifth best topic
# + hidden=true
plt.figure()
df_topic_probs_full["diff-5"].hist(bins=20)
# + [markdown] hidden=true
# ### Against fiftieth best topic
# + hidden=true
plt.figure()
df_topic_probs_full["diff-50"].hist(bins=20)
# + [markdown] hidden=true
# ## Most common words
# + hidden=true
df_topic_probs_full.sort_values(by="word-prob", ascending=False).head(10)[["word", "word-prob"]]
# + [markdown] hidden=true
# ## Highest std. dev.
# + hidden=true
df_topic_probs["stddev"] = df_topic_probs[model.prob_columns].std(axis=1)
df_topic_probs.sort_values(by="stddev", ascending=False).head(10)[["word", "stddev"]]
# + [markdown] hidden=true
# ## Lowest std. dev.
# + hidden=true
df_topic_probs["stddev"] = df_topic_probs[model.prob_columns].std(axis=1)
df_topic_probs.sort_values(by="stddev", ascending=True).head(10)[["word", "stddev"]]
# + [markdown] heading_collapsed=true
# # Correlation TM similarity and WE similarity
# + [markdown] hidden=true
# Topic model similarity evaluated using different probability distribution similarity measures (evaluated on the normalized word-topic distributions):
#
# * [Jensen-Shannon divergence](https://en.wikipedia.org/wiki/Jensen%E2%80%93Shannon_divergence)
# * [Hellinger distance](https://en.wikipedia.org/wiki/Hellinger_distance)
# * [Bhattacharyya coefficient](https://en.wikipedia.org/wiki/Bhattacharyya_distance#Bhattacharyya_coefficient)
# * Max difference
# * Sum difference ([total variation distance](https://en.wikipedia.org/wiki/Total_variation_distance_of_probability_measures))
# + [markdown] hidden=true
# ### Ten most similar words for each top-10-topic word
# + hidden=true
df_topic_similars["jensen-shannon"].head()
# + [markdown] hidden=true
# ### Correlation between TM and WE similarity
# + hidden=true
model.sim_functions = ["max", "sum", "bhattacharyya", "hellinger", "jensen-shannon"]
sim_corrs_spearman = []
sim_corrs_pearson = []
for sim_function in model.sim_functions:
corr_spearman = df_topic_similars[sim_function][["tm_sim", "we_sim"]].corr("spearman").ix[0,1]
corr_pearson = df_topic_similars[sim_function][["tm_sim", "we_sim"]].corr("pearson").ix[0,1]
sim_corrs_spearman.append(corr_spearman)
sim_corrs_pearson.append(corr_pearson)
df_tmp = pnd.DataFrame(model.sim_functions, columns=["sim_function"])
df_tmp["sim_corr_spearman"] = sim_corrs_spearman
df_tmp["sim_corr_pearson"] = sim_corrs_pearson
df_tmp
# + hidden=true
def correlation_in_group(corr_function):
def correlation(df_group):
return df_group.ix[:,-2:].corr(corr_function).ix[0,1]
return correlation
sim_corrs_spearman = []
sim_corrs_pearson = []
for sim_function in model.sim_functions:
df_tmp = df_topic_similars[sim_function]
df_group = df_tmp.groupby(np.arange(len(df_tmp)) // 10)
corr_spearman = df_group.apply(correlation_in_group("spearman")).mean()
corr_pearson = df_group.apply(correlation_in_group("pearson")).mean()
sim_corrs_spearman.append(corr_spearman)
sim_corrs_pearson.append(corr_pearson)
df_tmp = pnd.DataFrame(model.sim_functions, columns=["sim_function"])
df_tmp["sim_corr_spearman"] = sim_corrs_spearman
df_tmp["sim_corr_pearson"] = sim_corrs_pearson
df_tmp
# + [markdown] hidden=true
# **Note: Similar results Google vectors**
# + [markdown] hidden=true
# ### Distribution of TM similarity
# + hidden=true
plt.figure()
df_topic_similars["jensen-shannon"]["tm_sim"].hist(bins=100)
# + [markdown] hidden=true
# ### Distribution of WE similarity
# + hidden=true
plt.figure()
df_topic_similars["jensen-shannon"]["we_sim"].hist(bins=50)
# + hidden=true
plt.figure()
df_topic_similars["jensen-shannon"]["we_sim"].hist(bins=50, cumulative=True, normed=True)
# + hidden=true
def join_to_get_word_prob(df_param):
df_result = df_param.merge(df_topic_probs_full[["word", "word-prob"]],
left_on="similar_word", right_on="word",
suffixes=('', '_y'))
del df_result["word_y"]
return df_result
# + hidden=true
df_sim = join_to_get_word_prob(df_topic_similars["bhattacharyya"])
df_sim = df_sim[(df_sim["word-prob"] >= word_prob_lower_threshold) &
(df_sim["word-prob"] <= word_prob_upper_threshold)]
# + [markdown] heading_collapsed=true hidden=true
# ### High TM similarity, low WE similarity
# + hidden=true
df_high_tm_low_we = df_sim[(df_sim["we_sim"] < 0.4)]
df_high_tm_low_we.iloc[np.random.permutation(len(df_high_tm_low_we))]
# + [markdown] heading_collapsed=true hidden=true
# ### High TM similarity, high WE similarity
# + hidden=true
df_high_tm_low_we = df_sim[(df_sim["we_sim"] > 0.8)]
df_high_tm_low_we.iloc[np.random.permutation(len(df_high_tm_low_we))]
# + [markdown] hidden=true
# ### Low TM similarity, high WE similarity
# + hidden=true
df_embedding_similars = pnd.read_csv("../models/word-embeddings/embedding.model.skip-gram.similars.with-tm",
sep="\t", header=None)
df_embedding_similars.columns = ["word", "similar_word", "we_sim", "tm_sim"]
df_embedding_similars.head()
# + hidden=true
plt.figure()
df_embedding_similars["we_sim"].hist(bins=20)
# + hidden=true
plt.figure()
df_embedding_similars["tm_sim"].hist(bins=20)
# + hidden=true
plt.figure()
df_embedding_similars["tm_sim"].hist(bins=20, cumulative=True, normed=True)
# + hidden=true
df_sim2 = join_to_get_word_prob(df_embedding_similars)
df_sim2 = df_sim2[(df_sim2["word-prob"] >= word_prob_lower_threshold) &
(df_sim2["word-prob"] <= word_prob_upper_threshold)]
# + hidden=true
df_embedding_similars[df_embedding_similars["word"] == "france-based"]
# + hidden=true
df_low_tm_high_we = df_sim2[(df_sim2["tm_sim"] > 0.0) &
(df_sim2["tm_sim"] < 0.4)]
df_low_tm_high_we
# + [markdown] heading_collapsed=true
# # Findings
# + [markdown] hidden=true
# * syntatic variations play a bigger role in WE models, example:
#
# **(development, developed)**: TM-sim: 0.960519 WE-SIM: 0.360895
#
# **(composed, composers)** TM-SIM: 0.973376 WE-SIM: 0.329483
#
# **(works, working)** TM-SIM: 0.969470 WE-SIM: 0.274090
# * topic models are better at capturing loose relationships, such as:
#
# **(war, commander)** TM-SIM: 0.922352 WE-SIM: 0.187498
#
# **(living, households)** TM-SIM: 0.983162 WE-SIM: 0.207906
#
# **(county, rural)** TM-SIM: 0.882099 WE-SIM: 0.257984
#
# + [markdown] heading_collapsed=true
# # Concept categorization in TM and WE
# + [markdown] hidden=true
# Roughly the same results after using the same algorithm for both systems
# + hidden=true
def get_embedding_from_word_embedding(word):
try:
return vectors[word]
except:
return vectors["this"]
columns = [str(i) for i in range(256)]
def get_embedding_from_topics(word):
df_row = df_topic_probs_full[df_topic_probs_full["word"] == word]
assert len(df_row) == 1, "not exactly one row found: " + word + " " + len(df_row)
return df_row[columns].iloc[0,:].tolist()
def get_df_concept(embedding_function):
df_concept = pnd.read_csv(
"/home/knub/Repositories/master-thesis/data/concept-categorization/battig_concept-categorization.tsv",
sep="\t",
header=None)
df_concept.columns = ["word", "concept"]
df_concept["embeddings"] = df_concept["word"].apply(embedding_function)
return df_concept
df_we_concept = get_df_concept(get_embedding_from_word_embedding)
df_tm_concept = get_df_concept(get_embedding_from_topics)
df_tm_concept.head(2)
# + hidden=true
len(df_tm_concept.ix[0,"embeddings"])
# + hidden=true
from sklearn import metrics
# http://stats.stackexchange.com/questions/95731/how-to-calculate-purity
def single_cluster_purity(df_param):
return df_param["concept"].value_counts().max()
def calculate_purity(df_param):
purity = float(sum([single_cluster_purity(df_cluster_group)
for _, df_cluster_group
in df_param.groupby("cluster_id")])) / len(df_param)
return purity
def evaluate_clustering_algorithm(df_param, clustering):
X = np.array(df_param["embeddings"].tolist())
X_sim = metrics.pairwise.pairwise_distances(X, metric="cosine")
# sim or not sim? PCA or not PCA?
clusters = clustering.fit_predict(pca(X_sim, 20))
df_param["cluster_id"] = clusters
return calculate_purity(df_param)
# + hidden=true
for df_concept in [df_we_concept, df_tm_concept]:
print "-" * 100
for clustering in [KMeans(n_clusters=10, init="k-means++", n_jobs=1)]:
print clustering.__class__.__name__
print evaluate_clustering_algorithm(df_concept, clustering)
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
# + hidden=true
for df_concept in [df_we_concept, df_tm_concept]:
print "-" * 100
for clustering in [KMeans(n_clusters=10, init="k-means++", n_jobs=1),
AgglomerativeClustering(n_clusters=10, linkage="ward"),
AgglomerativeClustering(n_clusters=10, linkage="complete"),
AgglomerativeClustering(n_clusters=10, linkage="average"),
AffinityPropagation(damping=0.5),
AffinityPropagation(damping=0.6),
AffinityPropagation(damping=0.7),
AffinityPropagation(damping=0.8),
AffinityPropagation(damping=0.9),
SpectralClustering(n_clusters=3)]:
print clustering.__class__.__name__
print evaluate_clustering_algorithm(df_concept, clustering)
# -
# # Word Similarity
# + [markdown] heading_collapsed=true
# ## Similarity
# + hidden=true
def word_similarity(f):
try:
df_sim = pnd.read_csv(MODEL + f, sep="\t")
df_sim["embedding-sim"] = df_sim[["word1", "word2"]].apply(
lambda x: model.get_similarity(x["word1"], x["word2"], vectors), axis=1)
topic_sim_column = df_sim.columns[3]
topic_corr = df_sim[["human-sim", topic_sim_column]].corr("spearman").ix[0,1]
embedding_corr = df_sim[["human-sim", "embedding-sim"]].corr("spearman").ix[0, 1]
return pnd.DataFrame([[topic_corr, embedding_corr]],
columns=["topic_corr", "embedding_corr"],
index=[f])
except Exception as e:
return None
df_tmp = pnd.concat([word_similarity(".wordsim353-all-bhattacharyya"),
word_similarity(".wordsim353-all-hellinger"),
word_similarity(".wordsim353-all-jensen-shannon"),
word_similarity(".wordsim353-all-sum"),
word_similarity(".wordsim353-rel-bhattacharyya"),
word_similarity(".wordsim353-rel-hellinger"),
word_similarity(".wordsim353-rel-jensen-shannon"),
word_similarity(".wordsim353-rel-sum"),
word_similarity(".wordsim353-sim-bhattacharyya"),
word_similarity(".wordsim353-sim-hellinger"),
word_similarity(".wordsim353-sim-jensen-shannon"),
word_similarity(".wordsim353-sim-sum")])
df_tmp.sort_values(by="topic_corr", ascending=False)
# -
# ## Word Similarity performance with lower embedding dimensions
# ### Create word embeddings with different sizes
orig_vectors = load_skip_gram()
# +
#orig_vectors.save_word2vec_format("/home/knub/Repositories/master-thesis/data/word-similarity/wordsim353_sim_rel/dim-200.embedding", binary=False)
# -
with open("/home/knub/Repositories/master-thesis/data/word-similarity/wordsim353_sim_rel/dim-200.embedding", "r", encoding="utf-8") as f:
lines = [line.rstrip() for line in f]
count = int(lines[0].split(" ")[0])
lines = lines[1:]
words = []
vectors = []
for line in lines:
split = line.split(" ")
word = split[0]
words.append(word)
vector = [float(s) for s in split[1:]]
vectors.append(vector)
del lines
X = np.array(vectors)
print "Read embeddings"
print X.shape
print len(words)
# +
def project_down(n):
with open("/home/knub/Repositories/master-thesis/data/word-similarity/wordsim353_sim_rel/dim-%d.embedding" % n, "w", encoding="utf-8") as f:
f.write("%d %d\n" % (count, n))
pca_X = pca(X, n)
for i in range(count):
vector = pca_X[i,:]
output_vector = " ".join([str(v) for v in vector])
f.write("%s %s\n" % (words[i], output_vector))
DIMENSIONS = [110, 120, 130, 140]
for n in [d for d in DIMENSIONS if d != 200]:
print n
project_down(n)
gc.collect()
# -
# ### Evaluate performance
# +
df_wordsim353 = pnd.read_csv("/home/knub/Repositories/master-thesis/data/word-similarity/wordsim353_sim_rel/wordsim_all_goldstandard.txt",
sep="\t", header=None, names=["word1", "word2", "similarity"])
def get_similarity(word1, word2, v):
# ugly but works for now
if word1 not in v:
if word1.lower() in v:
word1 = word1.lower()
if word1.upper() in v:
word1 = word1.upper()
if word1.title() in v:
word1 = word1.title()
if word2 not in v:
if word2.lower() in v:
word2 = word2.lower()
if word2.upper() in v:
word2 = word2.upper()
if word2.title() in v:
word2 = word2.title()
try:
return v.similarity(word1, word2)
except KeyError:
print word1, word2
if word1 not in v:
print word1
if word2 not in v:
print word2
def evaluate():
for dim in DIMENSIONS:
gc.collect()
print dim
vectors = gensim.models.word2vec.Word2Vec.load_word2vec_format(
"/home/knub/Repositories/master-thesis/data/word-similarity/wordsim353_sim_rel/dim-%d.embedding" % dim,
binary=False)
df_wordsim353["dim-%d" % dim] = df_wordsim353[["word1", "word2"]].apply(
lambda x: get_similarity(x["word1"], x["word2"], vectors), axis=1)
evaluate()
gc.collect()
# -
for dim in DIMENSIONS:
print dim
print df_wordsim353["similarity"].corr(df_wordsim353["dim-%d" % dim])
| notebooks/Topics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.manifold import TSNE
import sklearn
import numpy as np
X = np.array([[1,2,3],[3,2,1],[4,5,6],[7,8,9]])
X.shape
tsne = TSNE(n_components=1)
tsne.fit_transform(X)
print(tsne.embedding_)
# +
from time import time
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.manifold import TSNE
def get_data():
digits = datasets.load_digits(n_class=6)
data = digits.data
label = digits.target
n_samples, n_features = data.shape
return data, label, n_samples, n_features
def plot_embedding(data, label, title):
x_min, x_max = np.min(data, 0), np.max(data, 0)
data = (data - x_min) / (x_max - x_min)
fig = plt.figure()
ax = plt.subplot(111)
for i in range(data.shape[0]):
plt.text(data[i, 0], data[i, 1], str(label[i]),
color=plt.cm.Set1(label[i] / 10.),
fontdict={'weight': 'bold', 'size': 9})
plt.xticks([])
plt.yticks([])
plt.title(title)
return fig
data, label, n_samples, n_features = get_data()
print('Computing t-SNE embedding')
tsne = TSNE(n_components=2, init='pca', random_state=0)
t0 = time()
result = tsne.fit_transform(data)
fig = plot_embedding(result, label,
't-SNE embedding of the digits (time %.2fs)'
% (time() - t0))
plt.show(fig)
# -
| Object Detection/TSNE-UMAP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def format_address(address_string):
# Declare variables
house_no = ''
address = ""
# Separate the address string into parts
input = address_string.split()
# Traverse through the address parts
for i in input:
if i.isnumeric():
house_no = i
else:
address += i + " "
# Determine if the address part is the
# house number or part of the street name
# Does anything else need to be done
# before returning the result?
# Return the formatted string
return "house number {} on street named {}".format(house_no, address)
print(format_address("123 Main Street"))
# Should print: "house number 123 on street named Main Street"
print(format_address("1001 1st Ave"))
# Should print: "house number 1001 on street named 1st Ave"
print(format_address("55 North Center Drive"))
# Should print "house number 55 on street named North Center Drive"
# +
def highlight_word(sentence, word):
return(sentence.replace(word, word.upper()))
print(highlight_word("Have a nice day", "nice"))
print(highlight_word("Shhh, don't be so loud!", "loud"))
print(highlight_word("Automating with Python is fun", "fun"))
# +
def combine_lists(list1, list2):
# Generate a new list containing the elements of list2
# Followed by the elements of list1 in reverse order
return list2 + list1[::-1]
Jamies_list = ["Alice", "Cindy", "Bobby", "Jan", "Peter"]
Drews_list = ["Mike", "Carol", "Greg", "Marcia"]
print(combine_lists(Jamies_list, Drews_list))
# +
def squares(start, end):
sq = []
while start<= end:
sq.append(start**2)
start += 1
return sq
print(squares(2, 3)) # Should be [4, 9]
print(squares(1, 5)) # Should be [1, 4, 9, 16, 25]
print(squares(0, 10)) # Should be [0, 1, 4, 9, 16, 25, 36, 49, 64, 81, 100]
# +
def car_listing(car_prices):
result = ""
for name, price in car_prices.items():
result += "{} costs {} dollars".format(name, price) + "\n"
return result
print(car_listing({"K<NAME>":19000, "<NAME>":55000, "Ford Fiesta":13000, "Toyota Prius":24000}))
# +
def combine_guests(guests1, guests2):
# Combine both dictionaries into one, with each key listed
# only once, and the value from guests1 taking precedence
guests = guests2.copy()
guests.update(guests1)
return guests
Rorys_guests = { "Adam": 2, "Brenda": 3, "David": 1, "Jose": 3, "Charlotte":2, "Terry":1, "Robert":4}
Taylors_guests = { "David":4, "Nancy":1, "Robert":2, "Adam":1, "Samantha":3, "Chris":5}
print(combine_guests(Rorys_guests, Taylors_guests))
# +
def count_letters(text):
result = {}
# Go through each letter in the text
for letter in text.lower():
# Check if the letter needs to be counted or not
if letter.isalpha() :
if letter not in result:
result[letter] = 0
# Add or increment the value in the dictionary
result[letter] += 1
return result
print(count_letters("AaBbCc"))
# Should be {'a': 2, 'b': 2, 'c': 2}
print(count_letters("Math is fun! 2+2=4"))
# Should be {'m': 1, 'a': 1, 't': 1, 'h': 1, 'i': 1, 's': 1, 'f': 1, 'u': 1, 'n': 1}
print(count_letters("This is a sentence."))
# Should be {'t': 2, 'h': 1, 'i': 2, 's': 3, 'a': 1, 'e': 3, 'n': 2, 'c': 1}
| Crash course on Pyhton/Week 4/Week 4 | Graded assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (datachallenges)
# language: python
# name: datachallenges
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
from insight_datachallenges.week2 import DATADIR
# -
# Overall approach
# ------
# I was provided with a chart showing user engagement over calendar time, as well as server log data on each user. I noticed right away that user engagement dropped at the end of July. Is this drop significant? And, what could be the reason for it?
#
# Conclusion
# ------------
# Vacation time in Europe is the likely cause of the decrease in user engagement.
events_csv_filename = os.path.join(DATADIR, 'yammer_events.csv')
events = pd.read_csv(events_csv_filename)
# I wasn't sure whether user engagement events were logged by user's local time or a time of day at the server's location. To infer this, I plotted the number of users signing up for Yammer as a function of time of day. If events were logged user-local time, I'd expect them to be distributed during typical waking hours. (This assumes a uniform distribution of users across the world).
events['occurred_at'] = pd.to_datetime(events['occurred_at'])
events['occurred_at_date'] = events['occurred_at'].apply(lambda x: x.date())
events['time_of_day'] = events['occurred_at'].apply(lambda x: x.hour)
engagement_mask = events['event_type'] == 'signup_flow'
event_time_groupby = events.loc[engagement_mask, :].groupby(['time_of_day'])
# a peek a the data (sanity check)
events.head()
# From the plot below, it's clear sign ups occur during normal waking hours, so the `occurred_at` column is indeed user local time.
counts_time_of_day = event_time_groupby.agg('count')
# df = counts_time_of_day.unstack(level=0)
fig, ax = plt.subplots(1, 1, figsize=(8, 8))
ax.set_ylabel('count', fontsize=18)
ax.set_xlabel('time of day (24 hour scale)', fontsize=18)
counts_time_of_day.loc[:, 'user_id'].plot(ax=ax)
# Let's just confirm the assumption that users are evenly distributed around the world. Below I've printed out the number of user events for each country.
location_groupby = events.groupby('location')
location_groupby.agg('count')['user_id']
# It looks like most users are from Europe, which suggests the end-of-July drop in engagement is due to Europeans going on holiday the month of August.
| others/Data_Challenge2/Vladimir_Shteyn_Yammer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from alpha_vantage.timeseries import TimeSeries
from datetime import datetime
import os
import sys
import pandas as pd
# -
# Define some symbol
symbol = "MSFT"
# Requires exporting the API Key
AV_API_KEY = os.environ.get("AV_API_KEY")
# Initialize time series object
timeSeries = TimeSeries(key=AV_API_KEY, output_format='pandas')
# Get data
data, meta = timeSeries.get_daily(symbol=symbol, outputsize='full')
# Name file
ofile = symbol + str(datetime.today().strftime('%Y-%m-%d'))
ofile
| lstm/Notebooks/.ipynb_checkpoints/prototyping-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Part of Speech Tagging
# Several corpora with manual part of speech (POS) tagging are included in NLTK. For this exercise, we'll use a sample of the Penn Treebank corpus, a collection of Wall Street Journal articles. We can access the part-of-speech information for either the Penn Treebank or the Brown as follows. We use sentences here because that is the preferred representation for doing POS tagging.
# +
from nltk.corpus import treebank, brown
print treebank.tagged_sents()[0]
print brown.tagged_sents()[0]
# -
# In NLTK, word/tag pairs are stored as tuples, the transformation from the plain text "word/tag" representation to the python data types is done by the corpus reader.
#
# The two corpus do not have the same tagset; the Brown was tagged with a more fine-grained tagset: for instance, instead of "DT" (determiner) as in the Penn Treebank, the word "the" is tagged as "AT" (article, which is a kind of determiner). We can actually convert them both to the Universal tagset.
print treebank.tagged_sents(tagset="universal")[0]
print brown.tagged_sents(tagset="universal")[0]
# Now, let's create a basic unigram POS tagger. First, we need to collect POS distributions for each word. We'll do this (somewhat inefficiently) using a dictionary of dictionaries.
# +
from collections import defaultdict
POS_dict = defaultdict(dict)
for word_pos_pair in treebank.tagged_words():
word = word_pos_pair[0].lower()
POS = word_pos_pair[1]
POS_dict[word][POS] = POS_dict[word].get(POS,0) + 1
# -
# Let's look at some words which appear with multiple POS, and their POS counts:
for word in POS_dict.keys()[:100]:
if len(POS_dict[word]) > 1:
print word
print POS_dict[word]
# Common ambiguities that we see here are between nouns and verbs (<i>increase</i>, <i>refunding</i>, <i>reports</i>), and, among verbs, between past tense and past participles (<i>contributed</i>, <i>reported</i>, <i>climbed</i>).
#
# To create an actual tagger, we just need to pick the most common tag for each
# +
tagger_dict = {}
for word in POS_dict:
tagger_dict[word] = max(POS_dict[word],key=lambda (x): POS_dict[word][x])
def tag(sentence):
return [(word,tagger_dict.get(word,"NN")) for word in sentence]
print tag(brown.sents()[0])
# -
# Though we'd probably want some better handling of capitalized phrases (backing off to NNP rather than NN when a word is capitalized), and there are a few other obvious errors, generally it's not too bad.
#
# NLTK has built-in support for n-gram taggers; Let's build unigram and bigram taggers, and test their performance. First we need to split our corpus into training and testing
size = int(len(treebank.tagged_sents()) * 0.9)
train_sents = treebank.tagged_sents()[:size]
test_sents = treebank.tagged_sents()[size:]
# Let's first compare a unigram and bigram tagger. All NLTK taggers have an evaluate method which prints out the accuracy on some test set.
# +
from nltk import UnigramTagger, BigramTagger
unigram_tagger = UnigramTagger(train_sents)
bigram_tagger = BigramTagger(train_sents)
print unigram_tagger.evaluate(test_sents)
print unigram_tagger.tag(brown.sents()[1])
print bigram_tagger.evaluate(test_sents)
print bigram_tagger.tag(brown.sents()[1])
# -
# The unigram tagger does way better. The reason is sparsity, the bigram tagger doesn't have counts for many of the word/tag context pairs; what's worse, once it can't tag something, it fails catastrophically for the rest of the sentence tag, because it has no counts at all for missing tag contexts. We can fix this by adding backoffs, including the default tagger with just tags everything as NN
# +
from nltk import DefaultTagger
default_tagger = DefaultTagger("NN")
unigram_tagger = UnigramTagger(train_sents,backoff=default_tagger)
bigram_tagger = BigramTagger(train_sents,backoff=unigram_tagger)
print bigram_tagger.evaluate(test_sents)
print bigram_tagger.tag(brown.sents()[1])
# -
# We see a 3% increase in performance from adding the bigram information on top of the unigram information.
#
# NLTK has interfaces to the Brill tagger (nltk.tag.brill) and also pre-build, state-of-the-art sequential POS tagging models, for instance the Stanford POS tagger (StanfordPOSTagger), which is what you should use if you actually need high-quality POS tagging for some application; if you are working on a computer with the Stanford CoreNLP tools installed and NLTK set up to use them (this is the case for the lab computers where workshops are held), the below code should work. If not, see the documentation <a href="https://github.com/nltk/nltk/wiki/Installing-Third-Party-Software"> here </a>
# +
from nltk import StanfordPOSTagger
stanford_tagger = StanfordPOSTagger('english-bidirectional-distsim.tagger')
print stanford_tagger.tag(brown.sents()[1])
| notebooks/.ipynb_checkpoints/part_of_speech_tagging-checkpoint (Jeremy Nicholson's conflicted copy 2017-03-06).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
class Network:
def __init__(self, xmin, xmax, ymin, ymax):
"""
xmin: 150,
xmax: 450,
ymin: 100,
ymax: 600
"""
self.StaticDiscipline = {
'xmin': xmin,
'xmax': xmax,
'ymin': ymin,
'ymax': ymax
}
def network(self, xsource, ysource = 100, Ynew = 600, divisor = 50): #ysource will always be 100
"""
For Network A
ysource: will always be 100
xsource: will always be between xmin and xmax (static discipline)
For Network B
ysource: will always be 600
xsource: will always be between xmin and xmax (static discipline)
"""
while True:
ListOfXsourceYSource = []
Xnew = np.random.choice([i for i in range(self.StaticDiscipline['xmin'], self.StaticDiscipline['xmax'])], 1)
#Ynew = np.random.choice([i for i in range(self.StaticDiscipline['ymin'], self.StaticDiscipline['ymax'])], 1)
source = (xsource, ysource)
target = (Xnew[0], Ynew)
#Slope and intercept
slope = (ysource - Ynew)/(xsource - Xnew[0])
intercept = ysource - (slope*xsource)
if (slope != np.inf) and (intercept != np.inf):
break
else:
continue
#print(source, target)
# randomly select 50 new values along the slope between xsource and xnew (monotonically decreasing/increasing)
XNewList = [xsource]
if xsource < Xnew:
differences = Xnew[0] - xsource
increment = differences /divisor
newXval = xsource
for i in range(divisor):
newXval += increment
XNewList.append(int(newXval))
else:
differences = xsource - Xnew[0]
decrement = differences /divisor
newXval = xsource
for i in range(divisor):
newXval -= decrement
XNewList.append(int(newXval))
#determine the values of y, from the new values of x, using y= mx + c
yNewList = []
for i in XNewList:
findy = (slope * i) + intercept#y = mx + c
yNewList.append(int(findy))
ListOfXsourceYSource = [(x, y) for x, y in zip(XNewList, yNewList)]
return XNewList, yNewList
# -
# Testing
net = Network(150,450,100,600)
NetworkA = net.network(300, ysource = 100, Ynew = 600) #Network A
NetworkB = net.network(200, ysource = 600, Ynew = 100) #Network B
#NetworkA
import seaborn as sns
import matplotlib.pyplot as plt
sns.jointplot(NetworkA[0], NetworkA[1])
plt.show()
sns.jointplot(NetworkB[0], NetworkB[1])
# +
#NetworkB
# +
DefaultPositionA = 300
DefaultPositionB = 300
def DefaultToPosition(x1, x2 = 300, divisor = 50):
XNewList = []
if x1 < x2:
differences = x2 - x1
increment = differences /divisor
newXval = x1
for i in range(divisor):
newXval += increment
XNewList.append(int(np.floor(newXval)))
else:
differences = x1 - x2
decrement = differences /divisor
newXval = x1
for i in range(divisor):
newXval -= decrement
XNewList.append(int(np.floor(newXval)))
return XNewList
out = DefaultToPosition(250)
print(out)
# -
j, x, y = next(zip(out, NetworkA[0], NetworkA[1]))
print(j, x, y)
# +
import pygame, sys
from pygame.locals import *
pygame.init()
import time
FPS = 50
fpsClock = pygame.time.Clock()
# set up the window
DISPLAYSURF = pygame.display.set_mode((600, 700), 0, 32)
pygame.display.set_caption('REINFORCEMENT LEARNING (Discrete Mathematics) - TABLE TENNIS')
# set up the colors
BLACK = ( 0,0,0)
WHITE = (255, 255, 255)
RED= (255,0,0)
GREEN = ( 0, 255,0)
BLUE = ( 0,0, 255)
# draw on the surface object
def display():
DISPLAYSURF.fill(WHITE)
pygame.draw.rect(DISPLAYSURF, GREEN, (150, 100, 300, 500))
pygame.draw.rect(DISPLAYSURF, RED, (150, 340, 300, 20))
pygame.draw.rect(DISPLAYSURF, BLACK, (0, 20, 600, 20))
pygame.draw.rect(DISPLAYSURF, BLACK, (0, 660, 600, 20))
return
# pixObj = pygame.PixelArray(DISPLAYSURF)
# del pixObj
PLAYERA = pygame.image.load('images/cap.jpg')
PLAYERA = pygame.transform.scale(PLAYERA, (50, 50))
PLAYERB = pygame.image.load('images/cap.jpg')
PLAYERB = pygame.transform.scale(PLAYERB, (50, 50))
ball = pygame.image.load('images/ball.png')
ball = pygame.transform.scale(ball, (15, 15))
playerax = 150
playerbx = 250
directionA = 'right'
directionB = 'right'
ballDirection = 'top'
ballx = 250
bally = 300
nextplayer = 'A'
lastxcoordinate = 350
count = 0
while True:
display()
if nextplayer == 'A':
#playerA should play
if count == 0:
#playerax = lastxcoordinate
NetworkA = net.network(lastxcoordinate, ysource = 100, Ynew = 600) #Network A
out = DefaultToPosition(lastxcoordinate)
#update lastxcoordinate
bally = NetworkA[1][count]
playerax = ballx
count += 1
# soundObj = pygame.mixer.Sound('sound/sound.wav')
# soundObj.play()
# time.sleep(0.4)
# soundObj.stop()
else:
ballx = NetworkA[0][count]
bally = NetworkA[1][count]
playerbx = ballx
playerax = out[count]
count += 1
#let playerB play after 50 new coordinate of ball movement
if count == 49:
count = 0
nextplayer = 'B'
else:
nextplayer = 'A'
else:
#playerB can play
if count == 0:
#playerbx = lastxcoordinate
NetworkB = net.network(lastxcoordinate, ysource = 600, Ynew = 100) #Network B
out = DefaultToPosition(lastxcoordinate)
#update lastxcoordinate
bally = NetworkB[1][count]
playerbx = ballx
count += 1
# soundObj = pygame.mixer.Sound('sound/sound.wav')
# soundObj.play()
# time.sleep(0.4)
# soundObj.stop()
else:
ballx = NetworkB[0][count]
bally = NetworkB[1][count]
playerbx = out[count]
playerax = ballx
count += 1
#update lastxcoordinate
#let playerA play after 50 new coordinate of ball movement
if count == 49:
count = 0
nextplayer = 'A'
else:
nextplayer = 'B'
#CHECK BALL MOVEMENT
DISPLAYSURF.blit(PLAYERA, (playerax, 50))
DISPLAYSURF.blit(PLAYERB, (playerbx, 600))
DISPLAYSURF.blit(ball, (ballx, bally))
#update last coordinate
lastxcoordinate = ballx
pygame.display.update()
fpsClock.tick(FPS)
for event in pygame.event.get():
if event.type == QUIT:
pygame.quit()
sys.exit()
# -
# # Define DQN Network
import keras
import tensorflow.compat.v1 as tf
from tensorflow.compat.v1.keras import models, Sequential, layers
def QNetwork(X_state, name):
with tf.variable_scope(name, reuse = tf.AUTO_REUSE) as scope:
hid1 = tf.layers.dense(X_state, 16, activation = 'relu')
hid2 = tf.layers.dense(hid1, 12 , activation= 'relu')
outputs = tf.layers.dense(hid2, 10, activation = 'softmax')
trainable_vars = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES,
scope = scope.name)
trainable_vars_by_name = {var.name[len(scope.name):]: var for var in trainable_vars}
return outputs, trainable_vars_by_name
# +
X_state = tf.placeholder(tf.float32, shape = (None, 1), name = 'X')
online_q_values, online_vars = QNetwork(X_state, name= 'q_networks/online')
target_q_values, target_vars = QNetwork(X_state, name= 'q_networks/target')
# -
copy_ops = [target_var.assign(online_vars[var_name]) for var_name, target_var in target_vars.items()]
copy_online_to_target = tf.group(*copy_ops)
# +
X_action = tf.placeholder(tf.int32, shape = [None])
q_value = tf.reduce_sum(target_q_values * tf.one_hot(X_action, 10),
axis = 1, keep_dims = True)
y = tf.placeholder(tf.float32, shape=[None, 1])
error = tf.abs(y - q_value)
clipped_error = tf.clip_by_value(error, 0.0, 1.0)
linear_error = 2 * (error - clipped_error)
loss = tf.reduce_mean(tf.square(clipped_error) + linear_error)
# -
learning_rate = 0.001
momentum = 0.95
global_step = tf.Variable(0, trainable=False, name='global_step')
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum, use_nesterov=True)
training_op = optimizer.minimize(loss, global_step=global_step)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
from collections import deque
replay_memory_size = 500000
replay_memory = deque([], maxlen=replay_memory_size)
def sample_memories(batch_size):
indices = np.random.permutation(len(replay_memory))[:batch_size]
cols = [[], [], [], [], []] # state, action, reward, next_state, continue
for idx in indices:
memory = replay_memory[idx]
for col, value in zip(cols, memory):
col.append(value)
cols = [np.array(col) for col in cols]
return (cols[0], cols[1], cols[2].reshape(-1, 1), cols[3],cols[4].reshape(-1, 1))
eps_min = 0.1
eps_max = 1.0
eps_decay_steps = 2000000
def epsilon_greedy(q_values, step):
epsilon = max(eps_min, eps_max - (eps_max-eps_min) * step/eps_decay_steps)
if np.random.rand() < epsilon:
return np.random.randint(n_outputs) # random action
else:
return np.argmax(q_values) # optimal action
import os, sys
os.getcwd()
n_steps = 4000000 # total number of training steps
training_start = 10000 # start training after 10,000 game iterations
training_interval = 4 # run a training step every 4 game iterations
save_steps = 1000 # save the model every 1,000 training steps
copy_steps = 10000 # copy online DQN to target DQN every 10,000 training steps
discount_rate = 0.99
skip_start = 90 # Skip the start of every game (it's just waiting time).
batch_size = 50
iteration = 0 # game iterations
checkpoint_path = "./my_dqn.ckpt"
done = True # env needs to be reset
with tf.Session() as sess:
if os.path.isfile(checkpoint_path + ".index"):
saver.restore(sess, checkpoint_path)
else:
init.run()
copy_online_to_target.run()
while True:
step = global_step.eval()
if step >= n_steps:
break
iteration += 1
if done: # game over, start again
obs = env.reset()
for skip in range(skip_start): # skip the start of each game
obs, reward, done, info = env.step(0)
state = preprocess_observation(obs)
# Online DQN evaluates what to do
q_values = online_q_values.eval(feed_dict={X_state: [state]})
action = epsilon_greedy(q_values, step)
# Online DQN plays
obs, reward, done, info = env.step(action)
next_state = preprocess_observation(obs)
# Let's memorize what just happened
replay_memory.append((state, action, reward, next_state, 1.0 - done))
state = next_state
if iteration < training_start or iteration % training_interval != 0:
continue # only train after warmup period and at regular intervals
# Sample memories and use the target DQN to produce the target Q-Value
X_state_val, X_action_val, rewards, X_next_state_val, continues = (sample_memories(batch_size))
next_q_values = target_q_values.eval(feed_dict={X_state: X_next_state_val})
max_next_q_values = np.max(next_q_values, axis=1, keepdims=True)
y_val = rewards + continues * discount_rate * max_next_q_values
# Train the online DQN
training_op.run(feed_dict={X_state: X_state_val,X_action: X_action_val, y: y_val})
# Regularly copy the online DQN to the target DQN
if step % copy_steps == 0:
copy_online_to_target.run()
# And save regularly
if step % save_steps == 0:
saver.save(sess, checkpoint_path)
| pytennis/game.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #!/usr/bin/python3
import os
import numpy as np
import string
import pandas as pd
from pandas import DataFrame
import random
import networkx as nx
import matplotlib
import matplotlib.pyplot as plt
import math
from collections import defaultdict
from sklearn import linear_model
from sklearn.metrics import mean_squared_error
from random import choice
# +
#This routine returns all the paths from the node v
def DFS(G,v,seen=None,path=None):
if seen is None: seen = []
if path is None: path = [v]
seen.append(v)
paths = []
for t in G[v]:
if t not in seen:
t_path = path + [t]
paths.append(tuple(t_path))
paths.extend(DFS(G, t, seen[:], t_path))
return paths
#Given a set of paths this routine returns the maximum length chain
def give_a_max_path(all_paths):
max_len = max(len(p) for p in all_paths)
max_paths = [p for p in all_paths if len(p) == max_len]
num_max_paths = len(max_paths)
chosen_path = max_paths[np.random.randint(num_max_paths)]
return chosen_path, max_len
def union(list1,list2):
return list(set(list1) | set(list2))
# +
# Defining the tree dictionary for each node
# Each block tree is a dictionary
# Each node is in nodes[] list
num_nodes = 10000
max_blocks = 100
#Chyeck list of lists
delay_vector= np.arange(0,5.25,0.25)
#delay_vector=[5]
num_delays=len(delay_vector)
num_trials = 4
MCL={}
#max_len_tracker = np.zeros((num_delays, max_blocks+1, num_trials))
D_tilde_factor = 0.1
threshold_coefficient=0.1
temporary_guess = x0_array
print(temporary_guess)
poll_L_array = [10]
#poll_L_array=[10]
delay_dict = {}
for i in poll_L_array:
delay_dict[i] = [0.5,1.5]
MCL[i] = {}
for j in delay_dict[i]:
MCL[i][j] = []
print(delay_dict)
print(MCL)
# +
for poll_L in poll_L_array:
print(poll_L)
delay_vector = delay_dict[poll_L]
num_delays=len(delay_vector)
for d in range(num_delays):
Delay = delay_vector[d]
data={}
D_tilde = D_tilde_factor*Delay
print("Starting with delay:", Delay)
for trial in range(num_trials):
nodes = num_nodes*[0]
#Tree variables
num_in_strings = [str(i) for i in range(max_blocks+1)]
for i in range(num_nodes):
nodes[i] = {i:[] for i in num_in_strings[:-1]}
# nodes[i] is a dictionary of block tree for node i which contains children of the blocks 0,1,....,T-1
god_node_children = {i:[] for i in num_in_strings}
god_node_parents = {i:"" for i in num_in_strings[:-1]}
rechability_matrix= np.zeros((num_nodes,max_blocks+1))
rechability_matrix[:,0]=1
delay_variables = np.random.exponential(Delay,(num_nodes,max_blocks+1))
poll_delay = np.random.exponential(D_tilde,(max_blocks+1,poll_L-1))
poll_delay = poll_delay + 0.1*D_tilde*(poll_L-1)
threshold_timer = threshold_coefficient*Delay
actual_poll_delay = np.full(max_blocks+1,threshold_timer)
maximum_poll_delay = np.amax(poll_delay,1)
actual_poll_delay = np.minimum(maximum_poll_delay,actual_poll_delay)
#print(D_tilde)
#print(np.mean(poll_delay))
#poll_delay = np.zeros((max_blocks+1))
#poll_delay = poll_delay+D_tilde
#Block arrival process
for t in range(1, max_blocks+1):
chosen_node = np.random.randint(num_nodes)
delay_variables[chosen_node,t] = 0 # Removing GOD model\n"
# Update block trees for all nodes "n"
if t==1:
god_node_children["0"].append(str(t))
god_node_parents[str(t)] = str(0)
else:
#Updating the block trees of all the nodes corresponding to the blocks 1,...t-1
for tau in range(1,t):
for k in range(num_nodes):
if delay_variables[k,tau] < t-tau-actual_poll_delay[tau]:
rechability_matrix[k,tau]=1
if str(tau) not in nodes[k][god_node_parents[str(tau)]]:
nodes[k][god_node_parents[str(tau)]].append(str(tau))
#Find the longest path for chosen node n starting from the genesis block 0,1,,
#If we poll we merge the dictionaries a.k.a local block trees and then chose the longest path
#graph = nodes[chosen_node]
#Updating the chosen node blocktree due to polling delay in proposal
for tau in range(1,t):
if delay_variables[chosen_node,tau] < t+actual_poll_delay[t]-tau-actual_poll_delay[tau]:
rechability_matrix[chosen_node,tau]=1
if str(tau) not in nodes[chosen_node][god_node_parents[str(tau)]]:
nodes[chosen_node][god_node_parents[str(tau)]].append(str(tau))
chosen_graph = nodes[chosen_node]
polled_node=chosen_node
polling_list=list(range(num_nodes))
polling_list.remove(polled_node)
for i in range(poll_L-1):
if poll_delay[t][i] <= actual_poll_delay[t]:
polled_node = choice(polling_list)
polled_graph = nodes[polled_node]
chosen_graph = {x:union(chosen_graph[x],polled_graph[x]) for x in chosen_graph}
polling_list.remove(polled_node)
all_paths = DFS(chosen_graph, "0")
if len(all_paths)==0:
chosen_leaf="0"
else:
chosen_path, max_len = give_a_max_path(all_paths)
chosen_leaf=chosen_path[-1]
#Attach this node in the Global tree
god_node_children[chosen_leaf].append(str(t))
god_node_parents[str(t)] = chosen_leaf
_, temp_length = give_a_max_path(DFS(god_node_children,"0"))
temp_length = temp_length-1
if t%25==0:
print(temp_length)
if t%100 == 0:
print("Trial {0}: Block {1} arrived : Growth rate of the main chain is {2}".format(trial, t, temp_length))
MCL[poll_L][Delay].append(temp_length)
#data["Trial%d"%(trial)]= max_len_tracker[d,:,trial]
#df = pd.DataFrame(data=data)
#df.to_csv('poll_delay_conservative/'+'updated_delay2_poll4_n%d_T%d_D%.2f.csv'%(num_nodes,max_blocks,Delay))
# -
print(MCL)
EMCL = {}
VMCL = {}
for i in range(10,11):
EMCL[i] = {}
VMCL[i] = {}
for j in [1-0.5,1+0.5]:
EMCL[i][j] = np.mean(np.array([MCL[i][j]]))
VMCL[i][j] = np.var(np.array([MCL[i][j]]))/4
print(VMCL[10])
# +
def calculate_x0(y1,y2,y0,x1,x2):
return(x1+ (y0-y1)*(x2-x1)/(y2-y1))
def calculate_var_x0(y1,y2,y0,x1,x2,vary1,vary2):
temp_term1 = ((x2-x1)**2)*((y0-y1)**2)/((y2-y1)**2)
temp_term2 = vary1/((y0-y1)**2) + (vary2+vary1)/((y2-y1)**2)
return temp_term1*temp_term2
# -
EMCL[1] = {}
VMCL[1] = {}
EMCL[1][0.5] = 88.0
EMCL[1][0.75] = 74.75
VMCL[1][0.5] = 1.5/4
VMCL[1][0.75] = 8.6875/4
# +
x0_array = []
std_x0_array = []
for i in range(10,11):
[x1,x2]=list(EMCL[i].keys())
[y1,y2]=list(EMCL[i].values())
[vary1,vary2] = list(VMCL[i].values())
y0 = 80
x0 = calculate_x0(y1,y2,y0,x1,x2)
var_x0=calculate_var_x0(y1,y2,y0,x1,x2,vary1,vary2)
x0_array.append(x0)
std_x0_array.append(np.sqrt(var_x0))
print(x0_array)
print(std_x0_array)
# -
x0_finalized = x0_array
std_x0_finalized = std_x0_array
#x0_finalized.pop()
#std_x0_finalized.pop()
x0_finalized.append(x0_array[0])
std_x0_finalized.append(std_x0_array[0])
print(x0_finalized)
print(std_x0_finalized)
plt.rcParams.update({'font.size': 14})
plt.figure(figsize=(10,7))
plt.errorbar(range(1,11),x0_finalized,std_x0_finalized,color=(0.5,0,0))
plt.xlabel('l')
plt.ylabel('D')
plt.legend(['0.01D(L-1)_connection_delay_0.1D_timer_delay'])
#plt.title('Lower_bound_tightness')
#plt.savefig('lower_bound1.fig')
plt.show()
np.save('DvsL_dict/0.01D_connect_0.1Dtimer_mean.npy',np.array(x0_finalized))
np.save('DvsL_dict/0.01D_connect_0.1Dtimer_std.npy',np.array(std_x0_finalized))
| poll_l_threshold_timer_conneciton_delay.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simultaneous phase and amplitude aberration sensing with a vector-Zernike wavefront sensor
#
# We will introduce the classical Zernike wavefront sensor (ZWFS) and a way to reconstruct phase aberrations. Then we will introduce the vector-Zernike WFS (vZWFS) and show how this version allows for simultaneous phase and amplitude aberration sensing.
# +
from hcipy import *
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
import os
# For notebook animations
from matplotlib import animation
from IPython.display import HTML
# +
pupil_grid = make_pupil_grid(256, 1.5)
aperture = make_magellan_aperture(True)
telescope_pupil = aperture(pupil_grid)
# -
# The classical Zernike wavefront sensor is implemented in hcipy. A ZWFS is a focal plane optic, but in HCIPy it is implemented as a pupil plane to pupil plane propagation, similar to the vortex coronagraph. This ensures optimal calculation speed using matrix Fourier transforms (MFT).
#
# First, we create the ZWFS optical element. In principle, the only parameter you have to give is the pupil grid. Other parameters that influence the performance are:
# 1. The phase step. For an optimal sensitivity, use $\pi/2$.
# 2. The phase dot diameter. For an optimal sensitivity, use 1.06 $\lambda/D$.
# 3. num_pix, sets the number of pixels the MFT uses.
# 4. The pupil diameter.
# 5. The reference wavelength.
#
ZWFS_ideal = ZernikeWavefrontSensorOptics(pupil_grid)
ZWFS_non_ideal = ZernikeWavefrontSensorOptics(pupil_grid, phase_step=0.45 * np.pi, phase_dot_diameter=1.2)
def plot_ZWFS(wavefront_in, wavefront_out):
'''Plot the input wavefront and ZWFS response.
Parameters
---------
wavefront_in : Wavefront
The aberrated wavefront coming in
wavefront_out : Wavefront
The wavefront_in propagated through the ZWFS
'''
# Plotting the phase pattern and the PSF
fig = plt.figure()
ax1 = fig.add_subplot(131)
im1 = imshow_field(wavefront_in.amplitude, cmap='gray')
ax1.set_title('Input amplitude')
divider = make_axes_locatable(ax1)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im1, cax=cax, orientation='vertical')
ax2 = fig.add_subplot(132)
im2 = imshow_field(wavefront_in.phase, cmap='RdBu')
ax2.set_title('Input phase')
divider = make_axes_locatable(ax2)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im2, cax=cax, orientation='vertical')
ax3 = fig.add_subplot(133)
im3 = imshow_field(wavefront_out.intensity, cmap='gray')
ax3.set_title('Output intensity')
divider = make_axes_locatable(ax3)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im3, cax=cax, orientation='vertical')
plt.show()
# Now it is time to measure the wavefront aberrations using the ZWFS. We create a random phase aberration with a power law distribution and propagate it through the ZWFS.
# +
# Creating the aberrated wavefront
phase_aberrated = make_power_law_error(pupil_grid, 0.2, 1)
phase_aberrated -= np.mean(phase_aberrated[telescope_pupil >= 0.5])
wf = Wavefront(telescope_pupil * np.exp(1j * phase_aberrated))
# Applying the ZWFS
wf_out = ZWFS_ideal.forward(wf)
plot_ZWFS(wf, wf_out)
# -
# From the intensity of the outcoming wavefront, it is clear that the intensity is dependent on the input phase. To show a simple reconstruction, we use the reconstruction algorithm of N'Diaye et al. 2013 [1]:
#
#
# $$\phi = −1 + \sqrt{2I_c}$$,
#
# where $\phi$ is the phase and $I_c$ the measured intensity.
#
# [1] N'Diaye et al. "Calibration of quasi-static aberrations in exoplanet direct-imaging instruments with a Zernike phase-mask sensor", Astronomy & Astrophysics 555 (2013)
def plot_reconstruction_phase(phase_in, phase_out, telescope_pupil):
'''Plot the incoming aberrated phase pattern and the reconstructed phase pattern
Parameters
---------
phase_in : Field
The phase of the aberrated wavefront coming in
phase_out : Field
The phase of the aberrated wavefront as reconstructed by the ZWFS
'''
# Calculating the difference of the reconstructed phase and input phase
diff = phase_out - phase_in
diff -= np.mean(diff[telescope_pupil >= 0.5])
# Plotting the phase pattern and the PSF
fig = plt.figure()
ax1 = fig.add_subplot(131)
im1 = imshow_field(phase_in, cmap='RdBu', vmin=-0.2, vmax=0.2, mask=telescope_pupil)
ax1.set_title('Input phase')
divider = make_axes_locatable(ax1)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im1, cax=cax, orientation='vertical')
ax2 = fig.add_subplot(132)
im2 = imshow_field(phase_out, cmap='RdBu', vmin=-0.2, vmax=0.2, mask=telescope_pupil)
ax2.set_title('Reconstructed phase')
divider = make_axes_locatable(ax2)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im2, cax=cax, orientation='vertical')
ax3 = fig.add_subplot(133)
im3 = imshow_field(diff, cmap='RdBu', vmin=-0.02, vmax=0.02, mask=telescope_pupil)
ax3.set_title('Difference')
divider = make_axes_locatable(ax3)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im3, cax=cax, orientation='vertical')
plt.show()
# +
phase_est = -1 + np.sqrt(2 * wf_out.intensity)
plot_reconstruction_phase(phase_aberrated, phase_est, telescope_pupil)
# -
# Using the simplified formula, we reconstruct the phase to a large extend and most of the structure in the phase pattern is reconstructed. However, the difference is larger than 10% of the original phase amplitude, and is structured in the shape similar to defocus.
#
# In N'Diaye et al. 2013, they propose a more accurate reconstruction:
#
# $$\phi = \sqrt{-1 + 3 − 2b −(1−I_c)/b},$$
#
# The $b$ is a term correction factor that takes into account the apodization from the Zernike mask. It can be approximated using a Strehl estimate, $S$, the Zernike mask geometry, $M$, and the telescope pupil geometry $P_0$:
#
#
# $$b \simeq S \widehat{M} \otimes P_0 = S b_0.$$
#
# Here $\widehat{M}$ is the Fourier transform of the Zernike mask geometry, $M$.
#
# In advance, the Strehl ratio is unknown. We set it to 1. Then we calculate the mask.
#
# +
S = 1
# Creating the mask M
focal_grid = make_focal_grid(20, 2)
M = Apodizer(circular_aperture(1.06)(focal_grid))
# Calculating b0
prop = FraunhoferPropagator(pupil_grid, focal_grid)
b0 = prop.backward(M(prop.forward(Wavefront(telescope_pupil)))).electric_field.real
# Calculating b
b = np.sqrt(S) * b0
# Estimating the phase using the equations from above
phase_est = -1 + np.sqrt(np.abs(3 - 2 * b - (1 - wf_out.intensity) / b))
plot_reconstruction_phase(phase_aberrated, phase_est, telescope_pupil)
# -
# It is clear that the ZWFS can accurately reconstruct the phase using the correction factor.
#
# However, one important limiting factor of the ZWFS reconstruction are amplitude errors, to which it is blind. This means that it reconstructs them as phase errors. To show this effect, we will propagate the phase aberration over a small distance using Fresnel propagation.
#
# In general, a ZWFS is least sensitive to tip-tilt modes and residual tip-tilt aberrations will dominate the reconstructed phase. In this notebook we will take the liberty to remove these tip-tilt modes. We also remove piston, as it is not important to reconstruct absolute phase.
# +
# Setting up the Fresnel propagator
prop_extra = FresnelPropagator(pupil_grid, distance=1e-4)
# Creating the power law error
phase_aberrated = make_power_law_error(pupil_grid, 0.3, 1)
phase_aberrated -= np.mean(phase_aberrated[telescope_pupil >= 0.5])
# Removing the piston and tip-tilt modes.
zbasis = make_zernike_basis(3, 1, pupil_grid)
for test in zbasis:
test*= telescope_pupil
phase_aberrated -= test * np.dot(phase_aberrated, test) / np.dot(test, test)
# Use super-Gaussian to avoid edge effects
p = telescope_pupil * np.exp(-(pupil_grid.as_('polar').r / 0.68)**20)
wf_new = prop_extra(Wavefront(p * np.exp(1j * phase_aberrated)))
# Updating reference phase and multiplying with the telescope pupil
phase_aberrated = (wf_new.phase - np.pi) % (2 * np.pi) - np.pi
wf_new.electric_field[telescope_pupil < 0.5] = 0
# Plotting the aberration
plt.figure()
plt.subplot(1,2,1)
imshow_field((wf_new.amplitude * telescope_pupil) - 1, vmin=-0.02, vmax=0.02, cmap='gray')
plt.title('Amplitude')
plt.subplot(1,2,2)
imshow_field(wf_new.phase*telescope_pupil, vmin=-0.2, vmax=0.2, cmap='RdBu', mask=telescope_pupil)
plt.title('Phase')
plt.show()
# -
# The Fresnel propagation added mostly high-frequency ampltitude variations and changed the phase a bit. To see the effect of these aberrations on the ZWFS, we redo the reconstruction.
# +
wf_out = ZWFS_ideal.forward(wf_new.copy())
S = 1
b = np.sqrt(S) * b0
phase_est = -1 + np.sqrt(np.abs(3 - 2 * b0 - (1 - wf_out.intensity) / b0))
plot_reconstruction_phase(phase_aberrated, phase_est, telescope_pupil)
# -
# As expected, the amplitude aberrations have been reconstructed as phase aberrations. The performance is worse and a lot of high-frequency noise has been added to the reconstructed phase.
#
# The rest of this notebook will introduce the vector-Zernike wavefront sensor, how the VZWFS output can be analysed and demonstrate the simultaneous phase and amplitude reconstruction.
#
# We start by creating the vZWFS optical element, similar to the ZWFS.
vZWFS_ideal = VectorZernikeWavefrontSensorOptics(pupil_grid, num_pix=128)
# To demonstrate the reconstructed amplitude, we generate the same plot function for amplitude as has been created for the phase.
def plot_reconstruction_amplitude(amplitude_in, amplitude_out, telescope_pupil):
'''Plot the incoming aberrated amplitude pattern and the reconstructed amplitude pattern
Parameters
---------
amplitude_in : Field
The phase of the aberrated wavefront coming in
amplitude_out : Field
The amplitude of the aberrated wavefront as reconstructed by the vZWFS
'''
amplitude_in = amplitude_in - 1
amplitude_out = amplitude_out - 1
# Plotting the phase pattern and the PSF
fig = plt.figure()
ax1 = fig.add_subplot(131)
im1 = imshow_field(amplitude_in, cmap='gray', vmin=-0.05, vmax=0.05, mask=telescope_pupil)
ax1.set_title('Input amplitude')
divider = make_axes_locatable(ax1)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im1, cax=cax, orientation='vertical')
ax2 = fig.add_subplot(132)
im2 = imshow_field(amplitude_out, cmap='gray', vmin=-0.05, vmax=0.05, mask=telescope_pupil)
ax2.set_title('Reconstructed amplitude')
divider = make_axes_locatable(ax2)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im2, cax=cax, orientation='vertical')
ax3 = fig.add_subplot(133)
im3 = imshow_field(amplitude_out - amplitude_in, cmap='gray', vmin=-0.01, vmax=0.01, mask=telescope_pupil)
ax3.set_title('Difference')
divider = make_axes_locatable(ax3)
cax = divider.append_axes('right', size='5%', pad=0.05)
fig.colorbar(im3, cax=cax, orientation='vertical')
plt.show()
# Now comes the most important part. The vZWFS applies $+\frac{\pi}{2}$ and $-\frac{\pi}{2}$ to the two orthogonal circular polarization states respectivily. Therefore, we have to add a circular polarizing beam splitter that separates these two polarization states.
#
# Then we follow the equations in Doelman et al. 2019 [1] to reconstruct the phase. We use both pupil intensities and combine these to calculate the phase and amplitude (or rather the square root of the intensity) from one snap shot.
#
# We start off by assuming a Strehl of 1. However, it is possible to update our estimate of b after calculating the phase and amplitude. We update the wavefront estimate that is used to calculate b and repeat the process. This allows for a slightly more accurate phase reconstruction.
#
# [1] Doelman, <NAME>., et al. "Simultaneous phase and amplitude aberration sensing with a liquid-crystal vector-Zernike phase mask." Optics letters 44.1 (2019)
def reconstructor(wavefront):
'''Plot the incoming aberrated phase pattern and the reconstructed phase pattern
Parameters
---------
phase_in : Field
The phase of the aberrated wavefront coming in
phase_out : Field
The phase of the aberrated wavefront as reconstructed by the ZWFS
'''
# Separate circular polarization states
CPBS = CircularPolarizingBeamSplitter()
wf_ch1, wf_ch2 = CPBS.forward(wavefront.copy())
I_L = wf_ch1.I
I_R = wf_ch2.I
# Creating masks for reconstruction
M = Apodizer(circular_aperture(1.06)(focal_grid))
b0 = np.abs(prop.backward(M(prop.forward(Wavefront(telescope_pupil)))).electric_field)
S = 1
b = np.sqrt(S) * b0
# Calculating the phase and amplitude aberrations.
for i in range(2):
amp_est = np.nan_to_num(np.sqrt(I_L + I_R + np.sqrt(np.abs(4 * b**2 * (I_R + I_L) - (I_R - I_L)**2 - 4 * b**4))))
phase_est = np.arcsin(I_L - I_R) / (2 * amp_est * b)
#Updating b for improved estimate
wf_est = Wavefront(amp_est * telescope_pupil * np.exp(1j * phase_est))
b = prop.backward(M(prop.forward(wf_est))).electric_field.real
return amp_est, phase_est
# +
amp_est, phase_est = reconstructor(vZWFS_ideal(wf_new.copy()))
I_est = amp_est**2
plot_reconstruction_phase(wf_new.phase, phase_est, telescope_pupil)
plot_reconstruction_amplitude(wf_new.I, I_est, telescope_pupil)
# -
# Both the phase and amplitude are reconstructed without cross-talk of phase and amplitude. This is the great advantage of a vector-Zernike wavefront sensor.
| doc/tutorial_notebooks/VectorZernikeWavefrontSensor/VectorZernikeWavefrontSensor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''passt_demo'': pyenv)'
# language: python
# name: python3
# ---
# +
from pathlib import Path
from typing import Dict, List
from hear21passt.base import load_model
import pandas as pd
from sklearn.metrics import f1_score
import torch
import torchaudio
from tqdm import tqdm
DEVICE = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
NUM_CLASSES = 527
TARGET_SR = 32_000
# -
class AudioSetDataset(torch.utils.data.Dataset):
def __init__(self, root: Path, labels: Dict[str, List[int]], audio_len: float = 10.0):
self._root = root
self._names = sorted(labels.keys())
self._labels = labels
self._num_samples = int(audio_len * TARGET_SR)
def __getitem__(self, index):
name = self._names[index]
path = self._root / (name + '.wav')
wav, sr = torchaudio.load(str(path))
assert wav.dim() == 2
wav = torch.mean(wav, dim=0)
if len(wav) > 0:
wav = torchaudio.functional.resample(wav, sr, TARGET_SR)
if len(wav) > self._num_samples:
wav = wav[:self._num_samples]
labels = self._labels[name]
target = torch.zeros(NUM_CLASSES)
for label in labels:
target[label] = 1.0
return wav, target
def __len__(self):
return len(self._names)
csv = pd.read_csv('data/audioset/class_labels_indices.csv')
label2ind = {}
for index, row in csv.iterrows():
label2ind[row.mid] = index
csv = pd.read_csv('data/audioset/val/valid.csv')
name2labels = {}
for _, row in csv.iterrows():
name2labels[row.YTID] = [label2ind[label] for label in row.positive_labels.split(',')]
ds = AudioSetDataset(Path('data/audioset/val/valid_wav'), name2labels)
loader = torch.utils.data.DataLoader(ds, batch_size=1, shuffle=False, num_workers=4, pin_memory=True)
model = load_model(mode='logits').eval().to(DEVICE)
def evaluate_thresholds(model, loader, thresholds):
scores = torch.zeros(len(thresholds))
model.eval()
for x, y in tqdm(loader):
if x.numel() == 0:
continue
x = x.to(DEVICE)
with torch.no_grad():
probs = torch.sigmoid(model(x)).cpu()
for i, threshold in enumerate(thresholds):
predictions = (probs > threshold).int()
scores[i] += f1_score(predictions.squeeze(), y.squeeze())
return scores / len(loader)
# +
import warnings
warnings.filterwarnings("ignore")
thresholds = torch.linspace(0.1, 0.9, 9)
scores = evaluate_thresholds(model, loader, thresholds)
print(scores)
# +
import matplotlib.pyplot as plt
_ = plt.title('Mean F1 score dependence on threshold')
_ = plt.xlabel('Threshold')
_ = plt.ylabel('Mean F1 score')
_ = plt.grid()
_ = plt.plot(thresholds, scores)
_ = plt.savefig('thresholds.jpg')
# +
import warnings
warnings.filterwarnings("ignore")
thresholds = torch.linspace(0.1, 0.3, 21)
scores = evaluate_thresholds(model, loader, thresholds)
print(scores)
# +
import matplotlib.pyplot as plt
_ = plt.title('Mean F1 score dependence on threshold')
_ = plt.xlabel('Threshold')
_ = plt.ylabel('Mean F1 score')
_ = plt.grid()
_ = plt.plot(thresholds, scores)
_ = plt.savefig('thresholds_small.jpg')
| threshold_search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <h1>Simulator</h1>
# <p>This notebook is meant to demonstrate the functionality of the simulator. I will provide a series of code examples along with visualizations to demonstrate what it enables</p>
# <h2>Import packages</h2>
# +
import os
from PIL import Image
import numpy as np
import trimesh
import warnings
# warnings.filterwarnings("ignore")
import meshplot as mp
from pterotactyl.simulator.scene import sampler
from pterotactyl.simulator.physics import grasping
from pterotactyl.utility import utils
import pterotactyl.objects as objects
# -
# <h2>Select object</h2>
OBJ_LOCATION = os.path.join(os.path.dirname(objects.__file__), "test_objects/0")
batch = [OBJ_LOCATION]
# <h2>Visualize object to be touched (OPTIONAL)</h2>
verts, faces = utils.load_mesh_touch(OBJ_LOCATION + '.obj')
plot = mp.plot(verts.data.cpu().numpy(), faces.data.cpu().numpy())
# <h2> Start the simulator and load the batch </h2>
# Here we setup the grasping environment, indicating that the batchsize is 1, that we want vision signals outputted, and that the resolution of the images should be 256 by 256. We then load the object into the simulator, and set the object scale to be 1/2.6 .
s = sampler.Sampler(grasping.Agnostic_Grasp, bs=1, vision=True, resolution = [256, 256])
s.load_objects(batch, from_dataset=False, scale = 2.6)
# <h2> Perform an action </h2>
action = [30]
parameters = [[[.3, .3, .3], [60, 0, 135]]]
signals = s.sample(action, touch=True, touch_point_cloud=True, vision=True, vision_occluded=True,parameters=parameters )
# <h2> Observe results </h2>
img_vision = Image.fromarray(signals["vision"][0])
display(img_vision)
img_vision_grasp = Image.fromarray(signals["vision_occluded"][0])
display(img_vision_grasp)
image = np.zeros((121*4, 121*2, 3)).astype(np.uint8)
for i in range(4):
print(f'Finger {i} has status {signals["touch_status"][0][i]}')
touch = signals["touch_signal"][0][i].data.numpy().astype(np.uint8)
image[i*121:i*121+121, :121] = touch
depth = utils.visualize_depth(signals["depths"][0][i].data.numpy()).reshape(121, 121, 1)
image[i*121:i*121+121, 121:] = depth
print(' ')
print(' TOUCH DEPTH')
display(Image.fromarray(image))
# <h2> Visualize the touches </h2>
# +
points = []
plot = mp.plot(verts.data.cpu().numpy(), faces.data.cpu().numpy())
for p in signals["touch_point_cloud"][0]:
if p.shape[0] >0:
points.append(p)
points = np.concatenate(points).reshape(-1,3)
plot.add_points(points, c=points.sum(axis=1), shading={ "point_size": 0.03})
# -
# <h2> Perfrom new actions </h2>
action = [40]
parameters = [[[0.35, -0.35, 0.3], [60, 0, 45]]]
signals = s.sample(action, touch=True, touch_point_cloud=False, vision=True, vision_occluded=True,parameters=parameters )
# <h2> Observe new results </h2>
img_vision = Image.fromarray(signals["vision"][0])
display(img_vision)
img_vision_grasp = Image.fromarray(signals["vision_occluded"][0])
display(img_vision_grasp)
# +
image = np.zeros((121*4, 121*2, 3)).astype(np.uint8)
for i in range(4):
print(f'Finger {i} has status {signals["touch_status"][0][i]}')
touch = signals["touch_signal"][0][i].data.numpy().astype(np.uint8)
image[i*121:i*121+121, :121] = touch
depth = utils.visualize_depth(signals["depths"][0][i].data.numpy()).reshape(121, 121, 1)
image[i*121:i*121+121, 121:] = depth
print(' ')
print(' TOUCH DEPTH')
display(Image.fromarray(image))
# -
| notebook/simulator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from datetime import date, datetime, timedelta
# import pandas as pd
import numpy as np
import glob
# %matplotlib notebook
import matplotlib.pyplot as plt
import sys
import datetime
# sys.path.insert(0, "../tools")
from covid19model.data.mobility import * # contains all necessary functions
from covid19model.visualization.output import *
from covid19model.data.sciensano import get_sciensano_COVID19_data_spatial
from covid19model.data.mobility import get_google_mobility_data
# OPTIONAL: Load the "autoreload" extension so that package code can change
# %load_ext autoreload
# OPTIONAL: always reload modules so that as you change code in src, it gets loaded
# %autoreload 2
# -
# Import Sciensano data
sciensano_df = get_sciensano_COVID19_data_spatial(agg='arr', values='hospitalised_IN', moving_avg=True).sum(axis=1)
sciensano_values= sciensano_df.values
sciensano_dates = sciensano_df.index
# # Google Mobility Data per activity
# +
start_date = datetime.datetime(2020, 1, 1, 0, 0)
end_date = datetime.datetime(2021, 9, 1, 0, 0)
start_calibration_wave1 = pd.Timestamp(2020, 3, 5, 0, 0)
end_calibration_wave1 = pd.Timestamp(2020, 8, 7, 0, 0)
start_calibration_wave2 = pd.Timestamp(2020, 9, 1, 0, 0)
end_calibration_wave2 = pd.Timestamp(2021, 2, 1, 0, 0)
fig, ax = plt.subplots(figsize=(9,3))
ax.grid(False)
ax.set_xlim(start_date, end_date)
ymin, ymax = (-100, 255)
ax.set_ylim(ymin, ymax)
color_timeframes(start_date, end_date, ax=ax)
# ax.set_title('test', size=14)
# Add shading denoting calibration periods
label_cal_size = 9
label_cal_move_up = 3
arrow_height = ymax
arrows = False
if arrows:
ax.annotate(s='', xy=(start_calibration_wave1,arrow_height), xytext=(end_calibration_wave1,arrow_height), \
arrowprops=dict(arrowstyle='<->'))
label_cal1_move_right = 44 # days
ax.text(start_calibration_wave1 + pd.Timedelta(days=label_cal1_move_right), \
ymax + label_cal_move_up, 'calibration wave 1', size=label_cal_size)
ax.annotate(s='', xy=(start_calibration_wave2,arrow_height), xytext=(end_calibration_wave2,arrow_height), \
arrowprops=dict(arrowstyle='<->'))
label_cal2_move_right = 41 # days
ax.text(start_calibration_wave2 + pd.Timedelta(days=label_cal2_move_right), \
ymax + label_cal_move_up, 'calibration wave 2', size=label_cal_size)
ticklabelsize = 12
plt.setp(ax.get_xticklabels(), Fontsize=ticklabelsize)
plt.setp(ax.get_yticklabels(), Fontsize=ticklabelsize)
plt.xticks(rotation=20)
# -
# **Insert Google Mobility parameters**
google_df = get_google_mobility_data(update=False)
google_df_dates = google_df.index.values
google_df.columns
# +
plt.style.use('seaborn-dark-palette')
linewidth=1
# colors = dict({'retail_recreation' : 'cornflowerblue',
# 'grocery' : 'firebrick',
# 'parks' : 'slategray',
# 'transport' : 'deeppink',
# 'work' : 'darkmagenta',
# 'residential' : 'darkturquoise'})
for loc in google_df.columns:
if loc=='retail_recreation':
ax.plot(google_df_dates, google_df[loc], linewidth=linewidth, label='retail & recreation') # color=colors[loc],
else:
ax.plot(google_df_dates, google_df[loc], linewidth=linewidth, label=loc) # color=colors[loc],
ax.legend(fontsize = ticklabelsize-2, loc='upper right', bbox_to_anchor=(0.27, 1))
ax.set_ylabel('mobility % to baseline', size=ticklabelsize)
plt.axhline(0, linestyle='--', linewidth=1, color='k', alpha=0.4)
plt.setp(ax.get_xticklabels(), Fontsize=ticklabelsize)
plt.setp(ax.get_yticklabels(), Fontsize=ticklabelsize)
plt.savefig('google-mobility-data.png', dpi=400, bbox_inches='tight')
# -
| notebooks/scratch/MR-visualise_timeline_google-mobility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Crash Course In Python And Manipulation of Data
# My objective for this lesson is to make you aware of python and its usefulness in wrangling with data.
#
# We are going to start slow with data types and expressions. Eventually, by the end of the lesson, we will programmatically genrerate data and then use a sophisticated library to help us manage and visualize it all.
#
# ### Data Types
# ##### Numbers
1
2 + 4
2 - 3
100 * 5
4 / 2
4 / 3
4 % 3
type(3)
type(3.5)
# ##### Strings
"Cats"
'Dogs'
'Hello World'
"I can't wait to program for the rest of my life"
"My name is " + "Andre"
type("Cats")
type("Doge")
# ##### Arrays
[1, 2, 3]
[100, 4, 12, 45]
["Cat", "Dog", "Zebra"]
# ##### Variables
# variable_name_here = expression_here
x = 4
x
x + 2
my_sweet_ride = "ferrari"
my_sweet_ride
my_name = "Andre"
"My name is " + my_name
type(x)
type(my_sweet_ride)
x = 4
y = 100
z = x * y
z
my_array = [100, 67, 4, 101]
my_array[0]
my_array[1]
my_array[2]
# This will error out because the index is asking for a location that is not in the
# array (out of range).
my_array[4]
# Access an index using a var!
pointless_var = 2
my_array[pointless_var]
my_array[0:3]
string_1 = "We are so lame."
string_1[0]
string_1[0:10]
new_string = string_1[0:10] + "cool."
new_string
# There are many ways to change the contents of lists besides assigning new values to individual elements:
#
odds = [1, 3, 5, 7]
print('odds are:', odds)
odds.append(11)
print('odds after adding a value:', odds)
del(odds[0])
print('odds after removing the first element:', odds)
odds.reverse()
print('odds after reversing:', odds)
odds = [1, 3, 5, 7]
print('odds are:', odds)
odds = odds + [11]
print('odds are:', odds)
# ##### Dictionaries
first_dict = {}
first_dict
first_dict["x"] = "some_var"
first_dict
first_dict[x] = 4
first_dict
second_dict = {"key_value": "value", "second_key_value": "second_value"}
second_dict
second_dict["key_value"]
second_dict["second_key_value"]
# ##### Exercise Block #1
# Create two python variables that will add up to 100.
# Add two python strings together to create a sentence.
# Create an array with number values only and assign it to a variable name. Then access the first element of that array.
# Create a dictionary and access a few values from it
# ##### Boolean Logic
True
False
1 == 1
2 == 1
1 < 10
1 <= 10
1 > 10
1 >= 10
"dogs are better than cats" == "dogs are better than cats"
"dogs" == "cats"
"abc" < "def"
"abc" > "def"
"a" < "b"
"a" > "b"
# Logic Flow
# +
x = 5
if x < 10:
print("This workshop is okay")
# +
x = 11
if x < 10:
print("This workshop is okay")
else:
print("This workshop is lame")
# +
str_1 = "Andre"
if str_1 == "Andre":
print("Welcome to the danger zone")
# +
my_var = 50
if my_var > 100:
print("Large value")
y = 10 # Example of doing different types of expressions and statements
elif my_var > 10:
print("Medium value")
y = 5
else:
print("Small value")
y = 0
# -
# ##### Looping
# +
list_of_elements = [1, 2, 3, 4]
# This could take a while to add everything by hand.
# This could take a while to add everything by hand.
for value in range(0, 10):
print(value)
# -
total_sum = 0
for value in range(0, 10):
total_sum = total_sum + value
total_sum
list_of_elements = []
for value in range(0, 10):
list_of_elements.append(value)
list_of_elements
# We can also start looping through all these values
for new_value in list_of_elements:
print(new_value * 2)
word = 'lead'
for char in word:
print(char)
# ###### For loops take the form of
# ```
# for variable in collection:
# do things with variable
# ```
# ##### Exercise Block #2
# Use a for-loop to convert the string “hello” into a list of letters:
# ```
# ["h", "e", "l", "l", "o"]
# ```
#
# Hint: You can create an empty list like this:
# ```
# my_list = []
# ```
| completed_lesson_part_1.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: Q 3.5
/ language: q
/ name: qpk
/ ---
/ # Machine Learning Demonstrations - 5
/
/ To run these notebooks, you need to install python dependencies:
/
/ With pip:
/
/ ```pip install numpy==1.14.0
/ pip install scipy==1.0.0
/ pip install graphviz==0.5.2
/ pip install matplotlib==2.0.2
/ pip install scikit_learn==0.19.1
/ pip install xgboost==0.7.post3```
/
/
/ Graphviz requires to be installed using *conda* also
/
/ With conda:
/
/ ```conda install -c anaconda graphviz```
/ ## Classification using Decision Trees
/
/ Decision Trees are a simple but effective algorithm, used for supervised classification and regression problems.
/
/ Decision Trees can
/ - Manage a mixture of discrete, continuous and categorical inputs
/ - Use data with no normalization/pre-processing (including missing data)
/ - Produce a highly-interpretable output, which can be easily explained and visualized
/
/ A decision tree is a collection of simple hierarchical decision rules, classifying datapoints into categories by *splitting* on feature values. The task of *fitting* a decision tree to data, is therefore the task of finding the sequence of feature splits and the optimal split values.
/
/ Further discussion of decision trees can be found on the [Wikipedia article](https://en.wikipedia.org/wiki/Decision_tree) or [Sci-Kit Learn documentation](http://scikit-learn.org/stable/modules/tree.html).
/ ## Breast Cancer Data
/ The Wisconsin Breast Cancer dataset is a set of 569 samples of fine needle aspirate (FNA) of breast mass. Each sample contains features describing characteristics of the cell nuclei , along with a classification of the sample as either benign or malignant.
/ ### Load data
/ Utility functions
\l ../utils/funcs.q
\l ../utils/graphics.q
data:.p.import[`sklearn.datasets;`:load_breast_cancer][]
X:data[`:data]`
y:data[`:target]`
featnames:getunicode data`:feature_names
/ Inspect data
-1"Shape of feature data is: ",sv[" x "]string shape X;
-1"";show 5#X;-1"";
-1"Distribution of target values is:\n";
show update pcnt:round[;.01]100*num%sum num from select num:count i by target from([]target:y);
/ Classes are quite unbalanced
/ - 37% are malignant (0)
/ - 63% are benign (1)
/ ### Split data
/ 50/50 training/test split
\S 123 / random seed
show count each datadict:traintestsplit[X;y;.5]
/ ### Fit model
/ We fit a DecisionTreeClassifier model, restricting the tree to a maximum depth of 3.
DecisionTreeClassifier:.p.import[`sklearn.tree]`:DecisionTreeClassifier
classifier:DecisionTreeClassifier[`max_depth pykw 3]
classifier[`:fit][datadict.xtrain;datadict.ytrain];
/ Using the graph visualization software [Graphviz](https://www.graphviz.org/), we can look at the structure of the resulting decision tree.
graphviz:.p.import`graphviz
exportgraphviz:.p.import[`sklearn.tree;`:export_graphviz]
dotdata:exportgraphviz[classifier;`out_file pykw (::);`feature_names pykw featnames]
graph:graphviz[`:Source]dotdata
display graph
/ The decision tree classifier produces a highly interpretable model which can be visualized and understood even by non-technical people.
/
/ The algorithm finds the best tree by following a *greedy* strategy.
/ It first finds the feature (mean concave points) and split value (0.052) that most effectively partitions the data.
/
/ This divides the dataset of 284 samples into two subsets of 176 samples and 108 samples.
/ - Of the 176 samples, 7 (4%) are malignant and 169 (96%) are benign.
/ - Of the 108 samples, 95 (88%) are malignant and 13 (12%) are benign.
/
/ So even one split value can provide a very powerful and predictive split between the classification labels.
/
/ The algorithm continues splitting the dataset at each node, by finding the feature and split value that most effectively partitions the benign from the malignant samples.
/ ### Evaluate model
/ The output of the decision tree is a class assignment.
/
/ We take a previously unseen sample and pass it through the decision tree. Following the appropriate branch at each split (based on the feature values of the test point), we eventually end up at a *leaf node*, at the bottom of the tree. At this point, we assign the test point the class value of the majority of the test examples included in that leaf.
/
/ We can therefore evaluate the performance of the decision tree, on the held-out test data.
/ +
yprob:classifier[`:predict_proba;<]datadict.xtest
ypred:raze {$[2=count x;1?;]x}each{x?max x}each yprob /#TODO ?
dtloss:logloss[datadict.ytest;yprob]
dtacc:accuracy[datadict.ytest;ypred]
-1"Performance of the classifier";
-1"log loss: ",(string dtloss),", accuracy: ",string dtacc;
/ -
/ The decision tree classifier achieves 95% accuracy on the test set, a strong performance from such a simple classifier.
/ ### Confusion matrix
/
/
/ With a Confusion Matrix, we can inspect the interaction between
/ - True positives
/ - True negatives
/ - False positives
/ - False negatives
show cnfM:cfm[datadict.ytest;ypred]
displayCM[value cnfM;`malignant`benign;"Test Set Confusion Matrix";()]
/ The classifier has
/
/ ```True Positive Rate TPR = TP/(TP+FN) = 99/(99+3) = 98%
/ False Positive Rate FPR = FP/(FP+TN) = 11/(11+172) = 6%```
/
/ n.b. We are using _positive_ here to denote the malignant case, which actually has the label 0, rather than 1, in the Wisconsin dataset.
/ ### ROC curve
/ Rather than using a _majority vote_ system, we could use a threshold other than 50% for assigning the points to classes at the leaf node.
/ - By modifying the threshold in favour of a _malignant_ diagnosis, we would increase the true positive rate, but would also increase the false positive rate.
/ - By modifying the threshold in favour of a _benign_ diagnosis, we would decrease the false positive rate, but would also decrease the true positive rate.
/
/ The realtionship between the true positive rate (sensitivity) and the false positive rate, is captured in the Receiver Operating Characteristic (ROC) curve.
/
/ n.b. 1 - FPR is called the _specificity_ of the model. We therefore have a trade-off between _sensitivity_ and _specificity_.
/
/ The area under the curve (AUC) is interpreted as the probability that a classifier will rank a randomly chosen _positive_ instance higher than a randomly chosen _negative_ one.
/
/ The optimal classifier would have a false positive rate of 0 and a true positive rate of 1, giving a an AUC curve of 1.
yprob:classifier[`:predict_proba;<;X]
displayROCcurve[y;yprob[;1]]
/ With an AUC of 0.96, our classifier is close to optimal.
| notebooks/ML05 Decision Trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lightning strikes
# First you have to import the meteomatics module and the datetime module
# +
import datetime as dt
import meteomatics.api as api
from __future__ import print_function
# -
# Input here your username and password from your meteomatics profile
###Credentials:
username = 'python-community'
password = '<PASSWORD>'
# Define a starting and an ending datetime
startdate_l = dt.datetime.utcnow() - dt.timedelta(days=1)
enddate_l = dt.datetime.utcnow() - dt.timedelta(minutes=5)
# Input here the limiting coordinates of the extract you want to look at.
lat_N_l = 90
lon_W_l = -180
lat_S_l = -90
lon_E_l = 180
# In the following, the request will start. If there is an error in the request as for example a wrong parameter or a date that doesn't exist, you get a message.
print("lightning strikes:")
try:
df_lightning = api.query_lightnings(startdate_l, enddate_l, lat_N_l, lon_W_l, lat_S_l, lon_E_l, username,
password)
print(df_lightning.head())
except Exception as e:
print("Failed, the exception is {}".format(e))
# You will get the data as a pandas dataframe. The index is made out of the validate, the latitude and the longitude. The column represents the stroke current. There are just events with lightning in the data frame. The time and points without lightning are not mentioned.
#
# 
# Now you can work on the data by using pandas commands. Here are some examples how you can access to the different datapoints.
maximum_stroke = df_lightning['stroke_current:kA'].max()
minimum_stroke = df_lightning['stroke_current:kA'].min()
mean_stroke = df_lightning['stroke_current:kA'].mean()
first_value = df_lightning.iloc[0]['stroke_current:kA']
just_after_18UTC = df_lightning[df_lightning.index.get_level_values('validdate').hour >= 18]
| examples/notebooks/07_Lightning_Strikes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## DeepExplain - Toy example of a model with multiple inputs
# ### Keras with Functional API
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import tempfile, sys, os
sys.path.insert(0, os.path.abspath('..'))
import numpy as np
import keras
from keras.datasets import mnist
from keras.models import Sequential, Model
from keras.layers import Dense, Dropout, Flatten, Activation, Input
from keras.layers import Conv2D, MaxPooling2D, Concatenate
from keras import backend as K
# Import DeepExplain
from deepexplain.tensorflow import DeepExplain
# +
# Define two simple numerical inputs
_x1 = np.array([[1]])
_x2 = np.array([[2]])
# Define model
# Here we implement y = sigmoid([w1*x1|w2*x2] * w3)
def init_model():
x1 = Input(shape=(1,))
x2 = Input(shape=(1,))
t1 = Dense(1)(x1)
t2 = Dense(1)(x2)
t3 = Concatenate()([t1, t2])
t4 = Dense(1)(t3)
y = Activation('sigmoid')(t4)
model = Model(inputs=[x1, x2], outputs=y)
model.compile(optimizer='rmsprop', loss='mse')
return model
model = init_model()
# This is a toy example. The random weight initialization will do just fine.
# model.fit(...)
# Make sure the model works
print ("Output: ", model.predict(x=[_x1, _x2]))
with DeepExplain(session=K.get_session()) as de: # <-- init DeepExplain context
# Need to reconstruct the graph in DeepExplain context, using the same weights.
input_tensors = model.inputs
fModel = Model(inputs = input_tensors, outputs = model.outputs)
target_tensor = fModel(input_tensors)
attributions = de.explain('grad*input', target_tensor, input_tensors, [_x1, _x2])
print ("Attributions:\n", attributions)
| examples/multiple_input_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Humpback Whale Identifaction -- public kernels summary
# ## 0. Some Exploration, including inspiration of data augmentation
# +
import math
from collections import Counter
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from PIL import Image
from tqdm import tqdm
# %matplotlib inline
INPUT_DIR = '../input'
def plot_images_for_filenames(filenames, labels, rows=4):
imgs = [plt.imread(f'{INPUT_DIR}/train/{filename}') for filename in filenames]
return plot_images(imgs, labels, rows)
def plot_images(imgs, labels, rows=4):
# Set figure to 13 inches x 8 inches
figure = plt.figure(figsize=(13, 8))
cols = len(imgs) // rows + 1
for i in range(len(imgs)):
subplot = figure.add_subplot(rows, cols, i + 1)
subplot.axis('Off')
if labels:
subplot.set_title(labels[i], fontsize=16)
plt.imshow(imgs[i], cmap='gray')
np.random.seed(42)
# -
train_df = pd.read_csv('../input/train.csv')
train_df.head()
# +
num_categories = len(train_df['Id'].unique())
print(f'Number of categories: {num_categories}')
# +
plt.figure(figsize=(10, 6))
plt.bar(range(len(size_buckets)), list(size_buckets.values())[::-1], align='center')
plt.xticks(range(len(size_buckets)), list(size_buckets.keys())[::-1])
plt.title("Num of categories by images in the training set")
plt.show()
# +
one_image_ids = train_df['Id'].value_counts().tail(8).keys()
one_image_filenames = []
labels = []
for i in one_image_ids:
one_image_filenames.extend(list(train_df[train_df['Id'] == i]['Image']))
labels.append(i)
plot_images_for_filenames(one_image_filenames, labels, rows=3)
# +
img_sizes = Counter([Image.open(f'{INPUT_DIR}/train/{i}').size for i in train_df['Image']])
size, freq = zip(*Counter({i: v for i, v in img_sizes.items() if v > 1}).most_common(20))
plt.figure(figsize=(10, 6))
plt.bar(range(len(freq)), list(freq), align='center')
plt.xticks(range(len(size)), list(size), rotation=70)
plt.title("Image size frequencies (where freq > 1)")
plt.show()
# -
from keras.preprocessing.image import (
random_rotation, random_shift, random_shear, random_zoom,
random_channel_shift, transform_matrix_offset_center, img_to_array)
# +
from keras import backend as K
intermediate_tensor_function = K.function([model.layers[0].input],[model.layers[layer_of_interest].output])
intermediate_tensor = intermediate_tensor_function([thisInput])[0]
# -
imgs = [
random_shear(img_arr, intensity=0.4, row_axis=0, col_axis=1, channel_axis=2, fill_mode='nearest') * 255
for _ in range(5)]
plot_images(imgs, None, rows=1)
imgs = [
random_zoom(img_arr, zoom_range=(1.5, 0.7), row_axis=0, col_axis=1, channel_axis=2, fill_mode='nearest') * 255
for _ in range(5)]
plot_images(imgs, None, rows=1)
# +
import random
def random_greyscale(img, p):
if random.random() < p:
return np.dot(img[...,:3], [0.299, 0.587, 0.114])
return img
imgs = [
random_greyscale(img_arr, 0.5) * 255
for _ in range(5)]
plot_images(imgs, None, rows=1)
# -
def augmentation_pipeline(img_arr):
img_arr = random_rotation(img_arr, 18, row_axis=0, col_axis=1, channel_axis=2, fill_mode='nearest')
img_arr = random_shear(img_arr, intensity=0.4, row_axis=0, col_axis=1, channel_axis=2, fill_mode='nearest')
img_arr = random_zoom(img_arr, zoom_range=(0.9, 2.0), row_axis=0, col_axis=1, channel_axis=2, fill_mode='nearest')
img_arr = random_greyscale(img_arr, 0.4)
return img_arr
imgs = [augmentation_pipeline(img_arr) * 255 for _ in range(5)]
plot_images(imgs, None, rows=1)
# ## 1. A model also inspiring
# +
batch_size = 128
num_classes = len(y_cat.toarray()[0])
epochs = 9
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
model = Sequential()
model.add(Conv2D(48, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(48, (3, 3), activation='sigmoid'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(48, (5, 5), activation='sigmoid'))
model.add(MaxPooling2D(pool_size=(3, 3)))
model.add(Dropout(0.33))
model.add(Flatten())
model.add(Dense(36, activation='sigmoid'))
model.add(Dropout(0.33))
model.add(Dense(36, activation='sigmoid'))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.summary()
model.fit_generator(image_gen.flow(x_train, y_train.toarray(), batch_size=batch_size),
steps_per_epoch= x_train.shape[0]//batch_size,
epochs=epochs,
verbose=1,
class_weight=class_weight_dic)
# -
# ## 2. Some methods for tackling data
# +
one_image_ids = train_df['Id'].value_counts().tail(8).keys()
one_image_filenames = []
labels = []
for i in one_image_ids:
one_image_filenames.extend(list(train_df[train_df['Id'] == i]['Image']))
labels.append(i)
plot_images_for_filenames(one_image_filenames, labels, rows=3)
# -
# ## 3. A way to improve model
# +
test_files = glob.glob("Whales/test/*.jpg")
l_image_name_test = [test_files[i].split('\\')[1] for i in range(len(test_files))]
l_class_data = [data['Id'][i] for i in range(len(data))] # data = file "train.csv"
# test_preds = predict of inference model for test images (data = for "train.csv" images)
test_image_dist_all = euclidean_distances(test_preds, data_preds)
preds_str = []
for ind in range(len(l_image_name_test)) :
test_image_dist = test_image_dist_all[ind] # distances between the test image and all the 'train.csv' images
vect_dist = [(l_class_data[i],test_image_dist[i]) for i in range(len(test_image_dist))] # create list of couples (class, distance)
vect_dist.append(("new_whale", 0.0)) # add "new_whale" ecach time
vect_dist.sort(key=lambda x: x[1]) # sort in order to have first the nearest
vect_dist = vect_dist[0:50] # best 50 nearest
vect_classes = [vect_dist[i][0] for i in range(len(vect_dist))]
# Maintain only one occurrence per class
vect_result = [vect_dist[0]] + [vect_dist[i] for i in range(1,len(vect_dist)) if vect_classes[i] not in vect_classes[0:i]]
vect_result = vect_result[:5] # take fist 5 nearest
preds_str.append(" ".join([x[0] for x in vect_result]))
# -
# ## 4. Another model but return no result
# +
from collections import defaultdict
import numpy as np
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
import os
import glob
from sklearn.neighbors import NearestNeighbors
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from keras import backend as K
from keras.models import Model
from keras.layers import Embedding, Flatten, Input, merge
from keras.optimizers import Adam
from keras.layers import Conv2D, MaxPooling2D, Input, Dense, Flatten, GlobalMaxPooling2D
from keras.models import Model
import glob
import os
from PIL import Image
from keras.callbacks import ModelCheckpoint, LearningRateScheduler, EarlyStopping, ReduceLROnPlateau, TensorBoard
from keras import optimizers, losses, activations, models
from keras.layers import Convolution2D, Dense, Input, Flatten, Dropout, MaxPooling2D, BatchNormalization, \
GlobalMaxPool2D, Concatenate, GlobalMaxPooling2D, GlobalAveragePooling2D, Lambda
from keras.applications.resnet50 import ResNet50
import pandas as pd
import numpy as np
import os
import glob
from sklearn.neighbors import NearestNeighbors
class sample_gen(object):
def __init__(self, file_class_mapping, other_class = "new_whale"):
self.file_class_mapping= file_class_mapping
self.class_to_list_files = defaultdict(list)
self.list_other_class = []
self.list_all_files = list(file_class_mapping.keys())
self.range_all_files = list(range(len(self.list_all_files)))
for file, class_ in file_class_mapping.items():
if class_ == other_class:
self.list_other_class.append(file)
else:
self.class_to_list_files[class_].append(file)
self.list_classes = list(set(self.file_class_mapping.values()))
self.range_list_classes= range(len(self.list_classes))
self.class_weight = np.array([len(self.class_to_list_files[class_]) for class_ in self.list_classes])
self.class_weight = self.class_weight/np.sum(self.class_weight)
def get_sample(self):
class_idx = np.random.choice(self.range_list_classes, 1, p=self.class_weight)[0]
examples_class_idx = np.random.choice(range(len(self.class_to_list_files[self.list_classes[class_idx]])), 2)
positive_example_1, positive_example_2 = \
self.class_to_list_files[self.list_classes[class_idx]][examples_class_idx[0]],\
self.class_to_list_files[self.list_classes[class_idx]][examples_class_idx[1]]
negative_example = None
while negative_example is None or self.file_class_mapping[negative_example] == \
self.file_class_mapping[positive_example_1]:
negative_example_idx = np.random.choice(self.range_all_files, 1)[0]
negative_example = self.list_all_files[negative_example_idx]
return positive_example_1, negative_example, positive_example_2
batch_size = 8
input_shape = (256, 256)
base_path = "../input/train/"
def identity_loss(y_true, y_pred):
return K.mean(y_pred - 0 * y_true)
def bpr_triplet_loss(X):
positive_item_latent, negative_item_latent, user_latent = X
# BPR loss
loss = 1.0 - K.sigmoid(
K.sum(user_latent * positive_item_latent, axis=-1, keepdims=True) -
K.sum(user_latent * negative_item_latent, axis=-1, keepdims=True))
return loss
def get_base_model():
latent_dim = 50
base_model = ResNet50(include_top=False) # use weights='imagenet' locally
# for layer in base_model.layers:
# layer.trainable = False
x = base_model.output
x = GlobalMaxPooling2D()(x)
x = Dropout(0.5)(x)
dense_1 = Dense(latent_dim)(x)
normalized = Lambda(lambda x: K.l2_normalize(x,axis=1))(dense_1)
base_model = Model(base_model.input, normalized, name="base_model")
return base_model
def build_model():
base_model = get_base_model()
positive_example_1 = Input(input_shape+(3,) , name='positive_example_1')
negative_example = Input(input_shape+(3,), name='negative_example')
positive_example_2 = Input(input_shape+(3,), name='positive_example_2')
positive_example_1_out = base_model(positive_example_1)
negative_example_out = base_model(negative_example)
positive_example_2_out = base_model(positive_example_2)
loss = merge(
[positive_example_1_out, negative_example_out, positive_example_2_out],
mode=bpr_triplet_loss,
name='loss',
output_shape=(1, ))
model = Model(
input=[positive_example_1, negative_example, positive_example_2],
output=loss)
model.compile(loss=identity_loss, optimizer=Adam(0.000001))
print(model.summary())
return model
model_name = "triplet_model"
file_path = model_name + "weights.best.hdf5"
def build_inference_model(weight_path=file_path):
base_model = get_base_model()
positive_example_1 = Input(input_shape+(3,) , name='positive_example_1')
negative_example = Input(input_shape+(3,), name='negative_example')
positive_example_2 = Input(input_shape+(3,), name='positive_example_2')
positive_example_1_out = base_model(positive_example_1)
negative_example_out = base_model(negative_example)
positive_example_2_out = base_model(positive_example_2)
loss = merge(
[positive_example_1_out, negative_example_out, positive_example_2_out],
mode=bpr_triplet_loss,
name='loss',
output_shape=(1, ))
model = Model(
input=[positive_example_1, negative_example, positive_example_2],
output=loss)
model.compile(loss=identity_loss, optimizer=Adam(0.000001))
model.load_weights(weight_path)
inference_model = Model(base_model.get_input_at(0), output=base_model.get_output_at(0))
inference_model.compile(loss="mse", optimizer=Adam(0.000001))
print(inference_model.summary())
return inference_model
def read_and_resize(filepath):
im = Image.open((filepath)).convert('RGB')
im = im.resize(input_shape)
im_array = np.array(im, dtype="uint8")[..., ::-1]
return np.array(im_array / (np.max(im_array)+ 0.001), dtype="float32")
def augment(im_array):
if np.random.uniform(0, 1) > 0.9:
im_array = np.fliplr(im_array)
return im_array
def gen(triplet_gen):
while True:
list_positive_examples_1 = []
list_negative_examples = []
list_positive_examples_2 = []
for i in range(batch_size):
positive_example_1, negative_example, positive_example_2 = triplet_gen.get_sample()
positive_example_1_img, negative_example_img, positive_example_2_img = read_and_resize(base_path+positive_example_1), \
read_and_resize(base_path+negative_example), \
read_and_resize(base_path+positive_example_2)
positive_example_1_img, negative_example_img, positive_example_2_img = augment(positive_example_1_img), \
augment(negative_example_img), \
augment(positive_example_2_img)
list_positive_examples_1.append(positive_example_1_img)
list_negative_examples.append(negative_example_img)
list_positive_examples_2.append(positive_example_2_img)
list_positive_examples_1 = np.array(list_positive_examples_1)
list_negative_examples = np.array(list_negative_examples)
list_positive_examples_2 = np.array(list_positive_examples_2)
yield [list_positive_examples_1, list_negative_examples, list_positive_examples_2], np.ones(batch_size)
num_epochs = 10
# Read data
data = pd.read_csv('../input/train.csv')
train, test = train_test_split(data, test_size=0.3, shuffle=True, random_state=1337)
file_id_mapping_train = {k: v for k, v in zip(train.Image.values, train.Id.values)}
file_id_mapping_test = {k: v for k, v in zip(test.Image.values, test.Id.values)}
train_gen = sample_gen(file_id_mapping_train)
test_gen = sample_gen(file_id_mapping_test)
# Prepare the test triplets
model = build_model()
#model.load_weights(file_path)
checkpoint = ModelCheckpoint(file_path, monitor='val_loss', verbose=1, save_best_only=True, mode='min')
early = EarlyStopping(monitor="val_loss", mode="min", patience=2)
callbacks_list = [checkpoint, early] # early
history = model.fit_generator(gen(train_gen), validation_data=gen(test_gen), epochs=3, verbose=2, workers=4, use_multiprocessing=True,
callbacks=callbacks_list, steps_per_epoch=300, validation_steps=30)
model_name = "triplet_loss"
def data_generator(fpaths, batch=16):
i = 0
for path in fpaths:
if i == 0:
imgs = []
fnames = []
i += 1
img = read_and_resize(path)
imgs.append(img)
fnames.append(os.path.basename(path))
if i == batch:
i = 0
imgs = np.array(imgs)
yield fnames, imgs
if i < batch:
imgs = np.array(imgs)
yield fnames, imgs
raise StopIteration()
data = pd.read_csv('../input/train.csv')
file_id_mapping = {k: v for k, v in zip(data.Image.values, data.Id.values)}
inference_model = build_inference_model()
train_files = glob.glob("../input/train/*.jpg")
test_files = glob.glob("../input/test/*.jpg")
train_preds = []
train_file_names = []
i = 1
for fnames, imgs in data_generator(train_files, batch=32):
print(i*32/len(train_files)*100)
i += 1
predicts = inference_model.predict(imgs)
predicts = predicts.tolist()
train_preds += predicts
train_file_names += fnames
train_preds = np.array(train_preds)
test_preds = []
test_file_names = []
i = 1
for fnames, imgs in data_generator(test_files, batch=32):
print(i * 32 / len(test_files) * 100)
i += 1
predicts = inference_model.predict(imgs)
predicts = predicts.tolist()
test_preds += predicts
test_file_names += fnames
test_preds = np.array(test_preds)
neigh = NearestNeighbors(n_neighbors=6)
neigh.fit(train_preds)
#distances, neighbors = neigh.kneighbors(train_preds)
#print(distances, neighbors)
distances_test, neighbors_test = neigh.kneighbors(test_preds)
distances_test, neighbors_test = distances_test.tolist(), neighbors_test.tolist()
preds_str = []
for filepath, distance, neighbour_ in zip(test_file_names, distances_test, neighbors_test):
sample_result = []
sample_classes = []
for d, n in zip(distance, neighbour_):
train_file = train_files[n].split(os.sep)[-1]
class_train = file_id_mapping[train_file]
sample_classes.append(class_train)
sample_result.append((class_train, d))
if "new_whale" not in sample_classes:
sample_result.append(("new_whale", 0.1))
sample_result.sort(key=lambda x: x[1])
sample_result = sample_result[:5]
preds_str.append(" ".join([x[0] for x in sample_result]))
df = pd.DataFrame(preds_str, columns=["Id"])
df['Image'] = [x.split(os.sep)[-1] for x in test_file_names]
df.to_csv("sub_%s.csv"%model_name, index=False)
| public kernels summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
# %matplotlib inline
import os,sys
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
from PIL import Image
import numpy as np
import torch
from torch.autograd import Variable
import timeit
# %load_ext autoreload
# %autoreload 2
# +
def apply_mirror_boundary_conditions(coord, dim):
"""
Return the correct coordinate according to mirror boundary conditions
coord: a coordinate (x or y) in the image
dim: the length of the axis of said coordinate
"""
# If the coordinate is outside of the bounds of the axis, take its reflection inside the image
if coord < 0:
coord = -coord
elif coord >= dim:
coord = 2*(dim-1) - coord % (2*(dim-1))
# Else, do nothing
return int(coord)
def get_window(image, window_size, centre_coordinates):
"""
Get a window in image taking into account boundary conditions
image: a numpy array representing our image
window_size: an odd number specifying the size of the window
centre_coordinates: a list containing the x-y coordinates of the window's central pixel
"""
# Get convenient variables
window_radius = (window_size - 1)/2
i_centre, j_centre = (centre_coordinates[0], centre_coordinates[1])
nrows, ncols, nchannels = image.shape
window = np.zeros((window_size, window_size, nchannels))
# Fill in the window array with pixels of the image
for i in range(window_size):
# Apply mirror boundary conditions on the x-coordinate
i_mirrored = apply_mirror_boundary_conditions(i_centre + i - window_radius, nrows)
for j in range(window_size):
# Same for the y-coordinate
j_mirrored = apply_mirror_boundary_conditions(j_centre + j - window_radius, ncols)
# Fill in the window with the corresponding pixel
window[i, j, :] = image[i_mirrored, j_mirrored, :]
return window
# -
for j in range(4, 3, -1):
print(j)
# +
def shift_to_the_right(image, window, centre_coordinates, step=1):
nrows, ncols, _ = image.shape
window_size = len(window)
window_radius = (window_size - 1)/2
#j_mirrored = apply_mirror_boundary_conditions(centre_coordinates[1] + step + window_radius, ncols)
shifted = np.roll(window, -step, axis=1)
for i in range(window_size):
i_mirrored = apply_mirror_boundary_conditions(centre_coordinates[0] + i - window_radius, nrows)
for j in range(window_size-step, window_size):
j_mirrored = apply_mirror_boundary_conditions(centre_coordinates[1] + j - window_radius + step, ncols)
shifted[i, j, :] = image[i_mirrored, j_mirrored, :]
return shifted
def shift_to_the_bottom(image, window, centre_coordinates, step=1):
nrows, ncols, _ = image.shape
window_size = len(window)
window_radius = (window_size - 1)/2
#i_mirrored = apply_mirror_boundary_conditions(centre_coordinates[0] + 1 + window_radius, nrows)
shifted = np.roll(window, -step, axis=0)
#print('shifted\n', shifted[:,:,0], '\n')
for j in range(window_size):
j_mirrored = apply_mirror_boundary_conditions(centre_coordinates[1] + j - window_radius, ncols)
for i in range(window_size-step, window_size):
i_mirrored = apply_mirror_boundary_conditions(centre_coordinates[0] + i - window_radius + step, nrows)
#print(i, i_mirrored)
shifted[i, j, :] = image[i_mirrored, j_mirrored, :]
return shifted
def sliding_window(image, window_size, step=1):
"""
Construct a list of sliding windows of given size on an image.
The windows will slide from left to right and from up to down.
image: a numpy array representing our image
window_size: an odd number specifying the size of the window
step: the value of the shift between windows
"""
nrows, ncols, _ = image.shape
windows = []
i = 0
row_windows = [get_window(image, window_size, [i, 0])]
for j in range(0, ncols-1, step):
row_windows += [shift_to_the_right(image, row_windows[-1], [i, j], step)]
windows += row_windows
for i in range(0, nrows-1, step):
row_windows = [shift_to_the_bottom(image, row_windows[int(j/step)], [i, j], step) for j in range(0, ncols, step)]
windows += row_windows
return windows
# +
# Directory and files name
train_dir = "training/"
tr_image_dir = train_dir + "images/"
tr_label_dir = train_dir + "groundtruth/"
tr_image_files = os.listdir(tr_image_dir)
tr_label_files = os.listdir(tr_label_dir)
# Number of training samples
N = len(tr_image_files)
# Load the images and ground truth
img_train = []
label_train = []
for i in range(N):
img = mpimg.imread(tr_image_dir + tr_image_files[i])
label = mpimg.imread(tr_label_dir + tr_label_files[i])
img_train.append(img)
label_train.append(label)
# Keep only sub-set of images
NUM_IMAGES = N
img_train = np.asarray(img_train[:NUM_IMAGES])
label_train = np.asarray(label_train[:NUM_IMAGES])
print(img_train.shape, label_train.shape)
# +
#im = img_train[0]
#for window_size in [3, 5, 7, 15, 35, 51]:
# %timeit sliding_window(im, window_size, step=int(window_size/2))
# -
def compute_all_windows(img_train, label_train, window_size, step):
train_data = []
train_labels = []
for im, labels, i in zip(img_train, label_train, range(len(img_train))):
w_im = sliding_window(im, window_size, step)
w_labels = sliding_window(labels[:, :, np.newaxis], window_size, step)
train_data += w_im
train_labels += w_labels
path = './windows_train/' + str(i)
os.makedirs(path, exist_ok=True)
for wi, wl, j in zip(w_im, w_labels, range(len(w_im))):
img_name = path + '/im_' + str(j) + '.png'
plt.imsave(img_name, wi)
label_name = path + '/label_' + str(j) + '.png'
plt.imsave(label_name, wl[:,:,0])
return train_data, train_labels
window_size = 51
step = int(window_size/2)
train_data, train_labels = compute_all_windows(img_train, label_train, window_size, step)
len(train_data) == len(train_labels)
train_d = np.array(train_data)
train_l = np.array(train_labels)[:, :, :, 0]
train_d = train_d.reshape((len(train_data)*51, 51, 3))
train_l = train_l.reshape((len(train_labels)*51, 51))
# +
#a = np.array(range(10*10))
#a = a.reshape((10, 10))
#print(a)
#aa = get_window(a[:,:,np.newaxis], 5, [5,5])[:,:,0]
#print('\n')
#print(aa)
#print('\n')
#print(shift_to_the_bottom(a[:,:,np.newaxis], aa[:,:,np.newaxis], [5,5], 2)[:,:,0])
# -
NUM_CHANNELS = 3 # RGB images
PIXEL_DEPTH = 255
NUM_LABELS = 2
IMG_PATCH_SIZE = 51
TRAINING_SIZE = 27300
BATCH_SIZE = 32
NUM_EPOCHS = 15
a = 0.00
from keras.models import Model, Sequential
from keras.layers import Dense, Dropout, Flatten, LeakyReLU, Input, Reshape, Permute, Average
from keras.layers import Conv2D, MaxPooling2D, Concatenate, Lambda, Activation, GlobalAveragePooling2D, Conv2DTranspose, GlobalAveragePooling1D
from keras.optimizers import SGD, Adam
# +
model = Sequential()
model.add(Conv2D(32, 2, input_shape=(IMG_PATCH_SIZE, IMG_PATCH_SIZE, NUM_CHANNELS)))
model.add(LeakyReLU(alpha=a))
model.add(Conv2D(32, 2))
model.add(LeakyReLU(alpha=a))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, 2))
model.add(LeakyReLU(alpha=a))
model.add(Conv2D(64, 2))
model.add(LeakyReLU(alpha=a))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128))
model.add(Dropout(0.5))
model.add(LeakyReLU(alpha=a))
model.add(Dense(1, activation='sigmoid'))
# -
from keras.engine.topology import Layer
def fire(x, squeeze, expand):
x = Conv2D(squeeze, 1)(x)
x = LeakyReLU(alpha=a)(x)
e11 = Conv2D(expand, 1)(x)
e11 = LeakyReLU(alpha=a)(e11)
e33 = Conv2D(expand, 1)(x)
e33 = LeakyReLU(alpha=a)(e33)
return Concatenate(axis=3)([e11, e33])
# +
inputs = Input(shape=(IMG_PATCH_SIZE, IMG_PATCH_SIZE, NUM_CHANNELS))
x = Conv2D(32, kernel_size=3, strides=2, input_shape=(IMG_PATCH_SIZE, IMG_PATCH_SIZE, NUM_CHANNELS))(inputs)
x = LeakyReLU(alpha=a)(x)
x = MaxPooling2D(pool_size=3, strides=2)(x)
x = fire(x, squeeze=8, expand=16)
x = fire(x, squeeze=8, expand=16)
x = MaxPooling2D(pool_size=3, strides=2)(x)
'''x = fire(x, squeeze=32, expand=128)
x = fire(x, squeeze=32, expand=128)
x = MaxPooling2D(pool_size=3, strides=2)(x)
x = fire(x, squeeze=48, expand=192)
x = fire(x, squeeze=48, expand=192)
x = fire(x, squeeze=64, expand=256)
x = fire(x, squeeze=64, expand=256)
x = MaxPooling2D(pool_size=3, strides=2)(x)'''
x = Flatten()(x)
x = Dense(1, activation='sigmoid')(x)
model = Model(inputs, x)
# +
model = Sequential()
model.add(Conv2D(32, 2, input_shape=(IMG_PATCH_SIZE, IMG_PATCH_SIZE, NUM_CHANNELS)))
model.add(LeakyReLU(alpha=a))
model.add(Conv2D(32, 2))
model.add(LeakyReLU(alpha=a))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, 2))
model.add(LeakyReLU(alpha=a))
model.add(Conv2D(64, 2))
model.add(LeakyReLU(alpha=a))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(2, 1))
model.add(Conv2DTranspose(1, kernel_size=IMG_PATCH_SIZE - 1, padding='valid'))
#model.add(Reshape((-1, IMG_PATCH_SIZE * IMG_PATCH_SIZE)))
#model.add(Permute((2,1)))
model.add(GlobalAveragePooling2D())
model.add(Activation('sigmoid'))
# -
for layer in model.layers:
print(layer.input_shape,layer.output_shape )
# +
LR = 0.001
DECAY = 0.00000
adam = Adam(lr=LR, decay=DECAY)
model.compile(loss='binary_crossentropy', optimizer=adam, metrics=['acc'])
model.fit(train_d, train_l, batch_size=BATCH_SIZE, epochs=NUM_EPOCHS)
# -
score = model.evaluate(train_data, train_labels)
score
| matt_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from uszipcode import Zipcode, SearchEngine
from geopy.distance import distance
covid = pd.read_csv('../data/california_covid.csv')
earthquake = pd.read_csv('../data/earthquakes_CA.csv')
fire = pd.read_csv('../data/fire_data.csv')
# -
covid.head(2)
# bringing in county population data as a column to the covid dataframe
population = pd.read_html('https://www.california-demographics.com/counties_by_population')[0]
population.head(2)
earthquake.head(2)
# Some fires need updated information for the acres burned. The inciweb has more updated information.
fyre = pd.read_html('https://inciweb.nwcg.gov/accessible-view/')[0]
fyre.head(3)
fyre = fyre[fyre['State'].str.contains('California')]
fyre = fyre[fyre['Type'] == 'Wildfire']
fyre.reset_index(drop=True, inplace=True)
fyre.head()
fyre.shape
fyre[fyre['Incident'] == 'Apple Fire']
fyre.iloc[16, 3]
fyre[fyre['Incident'].str.contains('Red')]
fire.head(3)
# subsetting rows that have 0 acres burned in the fire dataframe
fire_0 = fire[fire['AcresBurned'] == 0]
fire_0
len(fire_0)
# updating the `fire` dataframe with acres burned from inciweb (fyre dataframe)
for fire_name in fire_0['Name']:
# get the acres where the name matches
index1 = fyre[fyre['Incident'].str.contains(fire_name.split()[0])].index
# match only there is at least a match
if len(list(index1)) > 0:
acres = fyre.iloc[list(index1)[0], 3].split()[0]
index2 = fire[fire['Name'] == fire_name].index
fire.iloc[list(index2)[0], 4] = acres
# how many fires are active in Riverside
fire[fire['County'] == 'Riverside']
# check whether it's updated
fire_0 = fire[fire['AcresBurned'] == 0]
fire_0
fire[fire['PercentContained'] == 0]
# There are still fires that don't have the updated burned acres here. I can only find it through individual searches on inciweb. I will update the percent containment as well.
# The links are:
# [Slink Fire](https://inciweb.nwcg.gov/incident/article/7105/55963/)
# [Bobcat Fire](https://inciweb.nwcg.gov/incident/7152/)
# [Beach Fire](https://inciweb.nwcg.gov/incident/6987/)
# [W-5 Cold Springs](https://inciweb.nwcg.gov/incident/7010/)
#
# For percent containment:
# [Apple Fire, Red Salmon Complex, North Complex Fire, Dolan Fire, El Dorado Fire, Slater Fire (includes Devil Fire), and Fork Fire](https://www.sfchronicle.com/projects/california-fire-map/)
# [Blue Jay Fire](https://inciweb.nwcg.gov/incident/6888/)
# [Rattlesnake Fire](https://inciweb.nwcg.gov/incident/7131/)
# [Moraine Fire](https://inciweb.nwcg.gov/incident/7089/)
# [Bullfrog Fire](https://inciweb.nwcg.gov/incident/7191/)
# [Wolf Fire](https://inciweb.nwcg.gov/incident/7134/)
# [Hobo Fire](https://inciweb.nwcg.gov/incident/7126/)
# +
fire_update = {'Slink Fire': [26759, 90], 'Bobcat Fire ': [115796, 92],
'Beach Fire ': [3780, 100], 'W-5 Cold Springs': [84817, 100]}
for key, value in fire_update.items():
index = list(fire[fire['Name'] == key].index)
fire.iloc[index[0], 4] = value[0]
fire.iloc[index[0], 5] = value[1]
containment_update = {'Apple Fire': 95, 'Red Salmon Complex': 75, 'Blue Jay Fire': 50, 'North Complex Fire': 95,
'Dolan Fire': 98, 'Rattlesnake Fire': 40, 'Moraine Fire': 70, 'El Dorado Fire': 95,
'Slater Fire (includes Devil Fire)': 85, 'Fork Fire': 85, 'Bullfrog Fire': 60,
'Wolf Fire': 40, 'Hobo Fire ': 100}
for key, value in containment_update.items():
index = list(fire[fire['Name'] == key].index)
fire.iloc[index[0], 5] = value
# -
# verify
fire[fire['AcresBurned'] == 0]
# verify
fire[fire['PercentContained'] == 0]
fire.astype({'AcresBurned': 'int64'}).dtypes
fire.info()
# for some reason it's not letting the column to convert to int type, so I'm going to strip
# any possible white spaces and convert back to int
fire['AcresBurned'] = fire['AcresBurned'].apply(lambda x: int(str(x).split()[0]))
fire.info()
# This is fraction of california that has burned this year so far.
california_total_area = 99813760 # in acres; result from wikipedia (excludes water)
burned_area_2020 = sum(fire['AcresBurned'])
fraction_of_area_burned_2020 = burned_area_2020 / california_total_area
fraction_of_area_burned_2020
burned_area_2020 # total acres burned
# +
barh1 = california_total_area/california_total_area
barh2 = burned_area_2020/california_total_area
r = np.arange(barh1)
plt.figure(figsize=(1, 3))
plt.box(False)
plt.bar(r, california_total_area/california_total_area, color='lightgrey')
plt.bar(r, burned_area_2020/california_total_area, color='red')
plt.title('Burned Area in California in 2020')
# plt.text(x=r-0.6, y=barh1+0.01, s=str(california_total_area)+ ' acres')
plt.text(x=r+0.5, y=barh2, s='3,549,923 acres')
plt.text(x=r-0.1, y=barh2+0.02, s=str(int(fraction_of_area_burned_2020 *100))+'%')
plt.yticks([])
plt.xticks([])
plt.savefig('../figures/area_burned', bbox_inches='tight');
# -
# Number of fires this year
len(fire)
# Number of active fires as of October 17th
fire['IsActive'].sum()
covid.shape
population.shape
# the last row is not a county so drop it
population = population[:-1]
# the last row is not a county so drop it
population = population[population['County'].str.contains('County')]
# change `Population` data to int
population['Population'] = population['Population'].apply(lambda x: int(str(x).split()[0]))
population.dtypes
# fix county names so they don't contain 'County'
population['county'] = population['County'].apply(lambda x: ' '.join(x.split()[:-1]))
# drop 'Rank' and 'County' columns from population dataframe
population.drop(columns=['Rank', 'County'], inplace=True)
earthquake.shape
earthquake.info()
# number of earthquakes
len(earthquake)
sns.distplot(earthquake['Mag'])
plt.xlabel('Magnitude')
plt.ylabel('Frequency')
plt.title('Distribution of Earthquake Magnitudes')
plt.savefig('../figures/magnitude_distribution', bbox_inches='tight');
fire['Type'].value_counts()
# drop `Type` column because they are all wildfires
fire.drop(columns = ['Type'], inplace = True)
# The goal here is to merge dataframes based on county column as a common key. Covid data is clean and ready to go. Some fires extend to multiple counties. We are going to base our analysis on the county level, so we need to preserve the fire information for each county. Here, we are going to separate the county to individual counties while copying the fire data where the fire spans to multiple counties.
# first change the `County` column from str to a list
fire['County'] = fire['County'].apply(lambda x: x.split(', '))
# modified from https://stackoverflow.com/questions/24029659/python-pandas-replicate-rows-in-dataframe
# making copies of the rows with more than one county listed. The number of copies correspond to the number of counties
reps = [len(county) if len(county) > 1 else 1 for county in fire['County']]
fire = fire.loc[np.repeat(fire.index.values, reps)]
# reset index
fire.reset_index(inplace=True, drop=True)
# +
# reassign single counties to the `County` column
# index counter and an empty counties list
index = 0
counties = []
# loop to have an index track to not go out of index range
for indexer in range(len(fire)):
while index <= indexer:
# if there is only a single county name, append that county name to the list
if len(fire.iloc[index, 3]) == 1:
counties.append(fire.iloc[index, 3][0])
index += 1
# if there is more than one single county name, append each county name to the list
else:
for i in range(0, len(fire.iloc[index, 3])):
# append each indiviual county name of the multi-county list
counties.append(fire.iloc[index, 3][i])
# set the index, so that it goes to the next 'unique' item
index += len(fire.iloc[index, 3])
# reassign `County` column to this new list
fire['County'] = counties
# -
# The earthquake dataset does not include county names, so we are making a new column with the county names assigned from latitude and longitude using the python `uszipcode` library.
# make a function to retrive the county name from lat and long
def county_name(x):
'''Takes in latitude and longitude (as a literable) and returns the closest county name for the given coordinates'''
lat, long = x
search = SearchEngine()
result = search.by_coordinates(lat, long, radius=100)
# get the county name
try:
county = result[0].county
# this is in 'XXXX County', so fix the format so that it's only the county name without 'County' at the end
county = county.split()
# get everything but the last item (which is 'County')
county = county[:-1]
# return a string
return ' '.join(county)
except:
print(f'Something went wrong. Check your coordinates: {x}')
counties = earthquake[['Lat', 'Lon']].apply(county_name, axis=1)
# add a county column
earthquake['county'] = counties
# +
# making dictionaries to change column names before merging
covid_cols = {
'FIPS': 'fips',
'Admin2': 'county',
'Province_State': 'province_state',
'Country_Region': 'country',
'Last_Update': 'covid_last_update',
'Lat': 'county_latitude',
'Long_': 'county_longitudue',
'Confirmed': 'covid_confirmed',
'Deaths': 'covid_death',
'Recovered': 'covid_recovered',
'Active': 'covid_active',
'Combined_Key': 'combined_key',
'Incidence_Rate': 'covid_indidence_rate',
'Case-Fatality_Ratio': 'covid_case_fatality_ratio'
}
population_col = {'Population': 'county_population'}
earthquake_cols = {
'Event ID': 'earthquake_id',
'Name/Epicenter': 'eipicenter',
'Date': 'earthquake_date',
'Lat': 'earthquake_latidute',
'Lon': 'earthquake_longitude',
'Mag': 'magnitude'
}
fire_cols = {
'Name': 'fire_name',
'Final': 'fire_is_final',
'Started': 'fire_started',
'County': 'county',
'AcresBurned': 'acres_burned',
'PercentContained': 'fire_percent_contained',
'Longitude': 'fire_longitude',
'Latitude': 'fire_latitude',
'IsActive': 'fire_is_active',
'CalFireIncident': 'is_calfireincident',
'Location': 'fire_location'
}
# +
# rename all columns before merging
covid.rename(columns=covid_cols, inplace=True)
population.rename(columns=population_col, inplace=True)
earthquake.rename(columns=earthquake_cols, inplace=True)
fire.rename(columns=fire_cols, inplace=True)
# -
# Merge all three dataframes using `county` as the common key. In order to keep all information, we are using outer merge.
# +
# first merge the population data to covid data. this will be done as left join
covid = pd.merge(covid, population, on='county', how='left')
# merge covid and earthquake data
covid_earthquake = pd.merge(covid, earthquake, how='outer', on='county')
# merge fire data to the above
all_3 = pd.merge(covid_earthquake, fire, how='outer', on='county')
# write to file
all_3.to_csv('../data/covid_fire_earthquake.csv', index=False)
# -
# #### Summarize the all data on county-level
# add per capitat number for covid
covid['covid_death_per_capita'] = covid['covid_death'] / covid['county_population']
covid['covid_confirmed_per_capita'] = covid['covid_confirmed'] / covid['county_population']
covid['covid_active_cases_per_capita'] = covid['covid_active'] / covid['county_population']
# Here we want some county-level information on fires.
# +
# find how many fires per county in 2020
fires_per_county = fire.groupby('county')['fire_name'].size().to_frame()
# add fire per county data to covid
covid['fires_per_county_in_2020'] = covid.merge(fires_per_county, on='county', right_index=True)['fire_name']
# fill NaNs with 0 since those are the ones that had no fires this year
covid.fillna(0, inplace=True)
# +
# classify fires based on burned acres
def class_fire(acres_burned):
if (acres_burned > 0) and (acres_burned <= 200_000):
return 1
elif acres_burned <= 400_000:
return 2
elif acres_burned <= 600_000:
return 2
elif acres_burned <= 800_000:
return 2
elif acres_burned <= 1_000_000:
return 5
else:
return 6
# make a new column for fire classification
fire['fire_class'] = fire['acres_burned'].apply(class_fire)
# +
# add number of active fires per county to the covid data
# find how many fires per county in 2020
active_fires = fire[fire['fire_is_active']].groupby('county').size().to_frame()
# add fire per county data to covid
covid['active_fires_per_county'] = covid.merge(active_fires, on='county', right_index=True)[0]
# fill NaNs with 0 since those are the ones that had no fires this year
covid.fillna(0, inplace=True)
# -
# change`fire_is_active` column to True = 1 and False =0
fire['fire_is_active'] = np.where(fire['fire_is_active'] == True, 1, 0)
# We are going to give a score for fires. In this case, the score is based on the combination of number of fires, whether they are active or not, and the magnitude of the fire as classified above based on acres burned.
# $$ \text{fire score} = \text{number of fires} \times \text{active fire} + \text{number of fires} \times \text{fire class} $$
#
#
# +
# getting (number of fires) × (fire class)
fire_class_score = fire.groupby('county')['fire_class'].sum().to_frame()
# getting (number of fires) × (active fire)
fire_active_score = fire.groupby('county')['fire_is_active'].sum().to_frame()
# calculate the scores
fire_score = fire_class_score.merge(fire_active_score, on='county')
fire_score['score'] = fire_score['fire_class'] + fire_score['fire_is_active']
# get rid of unnecessary columns before merging
fire_score.drop(columns=['fire_class', 'fire_is_active'], inplace=True)
# add fire per county data to covid
covid['fire_score'] = covid.merge(fire_score, on='county', right_index=True)['score']
# fill NaNs with 0 since those are the ones that had no earthquakes this year
covid.fillna(0, inplace=True)
# -
# We also want county-level data on earthquakes
# +
# find how many earthquakes per county in 2020
earthquakes_per_county = earthquake.groupby('county')['earthquake_id'].size().to_frame()
# add fire per county data to covid
covid['earthquakes_per_county_in_2020'] = covid.merge(earthquakes_per_county, on='county', right_index=True)['earthquake_id']
# fill NaNs with 0 since those are the ones that had no earthquakes this year
covid.fillna(0, inplace=True)
# -
# We are going to classify earthquakes based on on their magnitude.
def class_earthquake(magnitude):
if magnitude >= 6:
return 3 # Strong category
elif magnitude >= 5:
return 2 # Moderate category
else:
return 1 # Light category
# make a column with earthquake classifications
earthquake['earthquake_class'] = earthquake['magnitude'].apply(class_earthquake)
# We want to give some metric of comparison for each county. For this, we are using the classification based on the magnitude and the number of earthquakes. For example, a county that has 2 earthquake in the category of 1 (lighht) would get a score of 2 (1 * 2) or a county with 3 earthquake in the category of 1 and 1 in the category of 2 (moderate) would get a score of 5 (3 * 1 + 1 * 2).
# +
earthquake_score = earthquake.groupby('county')['earthquake_class'].sum().to_frame()
# add fire per county data to covid
covid['earthquakes_score'] = covid.merge(earthquake_score, on='county', right_index=True)['earthquake_class']
# fill NaNs with 0 since those are the ones that had no earthquakes this year
covid.fillna(0, inplace=True)
# -
covid.drop(columns=['country', 'combined_key'], inplace = True)
pd.set_option('display.max_columns', 500)
covid.head(2)
# change dtypes to appropriate ones
covid = covid.astype({'fires_per_county_in_2020': 'int64',
'active_fires_per_county': 'int64',
'fire_score': 'int64',
'earthquakes_per_county_in_2020': 'int64',
'earthquakes_score': 'int64'})
covid.info()
# +
# write to file
#covid.to_csv('../data/covid_fire_earthquake_summary.csv', index=False)
# -
| code/merging_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <h1>CS4618: Artificial Intelligence I</h1>
# <h1>Regression Case Study</h1>
# <h2>
# <NAME><br>
# School of Computer Science and Information Technology<br>
# University College Cork
# </h2>
# + [markdown] slideshow={"slide_type": "skip"}
# <h1>Initialization</h1>
# $\newcommand{\Set}[1]{\{#1\}}$
# $\newcommand{\Tuple}[1]{\langle#1\rangle}$
# $\newcommand{\v}[1]{\pmb{#1}}$
# $\newcommand{\cv}[1]{\begin{bmatrix}#1\end{bmatrix}}$
# $\newcommand{\rv}[1]{[#1]}$
# $\DeclareMathOperator{\argmax}{arg\,max}$
# $\DeclareMathOperator{\argmin}{arg\,min}$
# $\DeclareMathOperator{\dist}{dist}$
# $\DeclareMathOperator{\abs}{abs}$
# + slideshow={"slide_type": "skip"}
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# + slideshow={"slide_type": "skip"}
import pandas as pd
import numpy as np
from pandas.plotting import scatter_matrix
from seaborn import scatterplot
from sklearn.model_selection import train_test_split
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import RobustScaler
from sklearn.preprocessing import OneHotEncoder
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.linear_model import Ridge
from sklearn.neighbors import KNeighborsRegressor
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import cross_validate
from sklearn.metrics import mean_absolute_error
from joblib import dump
# + [markdown] slideshow={"slide_type": "slide"}
# <h1>Introductory Remarks</h1>
# <ul>
# <li>In this lecture, we're going to rattle through the steps of a slightly larger case study.</li>
# <li>This will help bring out the importance of some steps we have not mentioned yet, including:
# <ul>
# <li>Dataset acquisition;</li>
# <li>Dataset exploration; and</li>
# <li>Dataset preprocessing.</li>
# </ul>
# </li>
# <li>If this were a module about data analytics, then each topic would fill several lectures. Since this is
# a module about AI, our treatment will be much more cursory.
# </li>
# </ul>
# -
# <h1>Identify the Problem</h1>
# <ul>
# <li>What are the business objectives?</li>
# <li>How do we expect to benefit by building and using an AI system for making predictions?
# How will it contribute to the business objectives?
# </li>
# </ul>
# <h1>Select Performance Measures</h1>
# <ul>
# <li>Select a performance measure:
# <ul>
# <li>E.g. for regression, we might select MAE.
# </ul>
# </li>
# <li>But we might also want a 'reference point' to know whether we have succeeded:
# <ul>
# <li>E.g. maybe we have succeeded if our MAE is 10% lower than the current system's MAE.</li>
# <li>E.g. maybe we must not only be better than the current system but statistically significantly
# better (according to some test of statistical significance).
# </li>
# <li>E.g. maybe we have succeeded if MAE is no greater than some threshold.</li>
# </ul>
# </li>
# <li>Sometimes there is more than one performance measure:
# <ul>
# <li>E.g. maybe in regression we want to separate the errors by measurng overestimates
# separately from underestimates.
# </li>
# <li>E.g. maybe we want to measure training time or memory used during training.</li>
# <li>E.g. maybe we want to measure prediction time or memory used when making a prediction.</li>
# <li>E.g. maybe we want an incremental learning algorithm that can easily accommodate new training
# examples when they arise.
# </li>
# <li>E.g. maybe we care about learning an interpretable model or being able to explain the system's
# predictions.
# </li>
# <li>And so on.</li>
# </ul>
# </li>
# </ul>
# + [markdown] slideshow={"slide_type": "slide"}
# <h1>Dataset Acquisition</h1>
# <ul>
# <li>Where will it come from? Existing databases and files? By scanning paper documents? By scraping web sites?
# …
# </li>
# <li>What quantities of data are available? How much do you need?</li>
# <li>What format is it in? What will you need to do to convert it?
# <ul>
# <li>E.g. you may need to de-normalize relational databases or flatten other data structures such as trees
# and graphs in order to produce the tabular (matrix) format that most learning algorithms expect.</li>
# <li>E.g. you may need to handle different character encodings.</li>
# </ul>
# </li>
# <li>Are there legal or ethical issues: copyright, authorization, privacy, bias? E.g. will you need to
# anonymize? Wil you need to de-bias?
# </li>
# <li>If you need a labeled dataset, are the labels available? Are they reliable? How will you
# obtain them if not? (Get an expert? Get several experts? Use crowdsourcing?)
# </li>
#
# </ul>
# -
# <h1>Data Cleaning</h1>
# <ul>
# <li>Take a quick look at the data: compute some summary statistics, look at a few examples.</li>
# <li>Identify any problems.</li>
# <li>Clean the data accordingly:
# <ul>
# <li>E.g. datasets may contain rows that violate your normal data validation criteria: perhaps
# a feature value is required but there are rows where it is missing; perhaps there is a
# maximum value for a feature and there are rows where this maximum is exceeded;
# perhaps a non-numeric
# feature has a finite set of allowed values but some rows contain illegal values for this
# feature. You must
# either fix these rows (if that is possible) or delete them.
# </li>
# <li>E.g. if we are doing supervised learning, we need target values: they cannot be missing and
# they need to be sensible.
# </li>
# <li>E.g. we may need to identify and handle duplicated examples.
# </ul>
# </li>
# <li>Note that this is one of several steps where it would be useful to be able to discuss with
# a domain expert.
# </li>
# </ul>
# + [markdown] slideshow={"slide_type": "slide"}
# <h2>Data cleaning for the Cork Property Prices Dataset</h2>
# <ul>
# <li>Let's illustrate data cleaning for a different (less clean) version of the Cork Property Prices Dataset.</li>
# </ul>
# +
# Use pandas to read the CSV file into a DataFrame
df = pd.read_csv("../datasets/dataset_corkB.csv")
# Shuffle the dataset
df = df.sample(frac=1, random_state=2)
df.reset_index(drop=True, inplace=True)
# -
# The dimensions
df.shape
# The columns
df.columns
# The datatypes
df.dtypes
# Maybe have a look at a few rows
df.head()
# Summary statistics
df.describe(include="all")
features = ["flarea", "bdrms", "bthrms", "floors", "type", "devment", "ber", "location"]
numeric_features = ["flarea", "bdrms", "bthrms", "floors"]
nominal_features = ["type", "devment", "ber", "location"]
# The values, in the case of nominal-valued features
for feature in nominal_features:
print(feature, df[feature].unique())
# Check for NaNs in the case of numeric features
for feature in numeric_features:
print(feature, df[feature].isnull().values.any())
# Check for NaN in the case of the target values
df["price"].isnull().values.any()
# <ul>
# <li>What problems do you see?</li>
# <li>Let's assume that, according to our normal data validation criteria, it is invalid to supply
# no value for $\mathit{devment}$ but it is OK to supply no value for $\mathit{ber}$. So we will delete
# rows that have NaN for their $\mathit{devment}$.
# </li>
# <li>Let's assume similarly that values for $\mathit{flarea}$ cannot be missing and must be greater
# than or equal to 40 and less than 750. So we will delete rows that violate these conditions too.
# </li>
# <li>Let's assume we also want to delete rows where $\mathit{bdrms}$ is NaN.
# </li>
# <li>We must delete the rows where $\mathit{price}$ is NaN because we are doing supervised learning!
# But I fear that some of these prices are wrong too. A scatter plot helps show why.
# </li>
# </ul>
plot = scatterplot(x="flarea", y="price", data=df)
# <ul>
# <li>So let's look at the row(s) that have the very high price tag:</li>
# </ul>
df[df["price"] >= 2000]
# <ul>
# <li>Undoubtedly a typo! So we will delete this row too.</li>
# </ul>
# +
# Delete examples where flarea, devment or price are NaN
df.dropna(subset=["flarea", "bdrms", "devment", "price"], inplace=True)
# Delete examples whose floor areas are too small or too big
df = (df[(df["flarea"] >= 40) & (df["flarea"] < 750)]).copy()
# Delete examples whose prices are too high
df = (df[df["price"] < 2000]).copy()
# Reset the index
df.reset_index(drop=True, inplace=True)
# Check the invalid data was filtered out
df.shape
# -
# <h1>Create a Test Set</h1>
# <ul>
# <li>Split off a test set.</li>
# <li>Don't look at it: everything we do until our final error estimation will be done on the rest of
# the dataset.
# </li>
# </ul>
# Split off the test set: 20% of the dataset.
dev_df, test_df = train_test_split(df, train_size=0.8, random_state=2)
# <h1>Dataset Exploration</h1>
# <ul>
# <li>Time spent just exploring the data is always worthwhile — especially if you have
# access to a domain expert while you are doing so.
# </li>
# <li><b>Visualization</b> is a great help at this stage.
# <ul>
# <li>Given a labeled dataset, we often want to see how target values are distributed
# with respect to values of one or more of the features.
# </li>
# <li>Given a labeled or unlabeled dataset, we might want to see how values of one feature are
# correlated with values of one or more other features.
# </li>
# </ul>
# A limitation is that visualizations are confined to two (or maybe three) columns of the dataset.
# </li>
# </ul>
# It can be good to do this on a copy of the dataset (excluding the test set, of course)
copy_df = dev_df.copy()
m = scatter_matrix(copy_df, figsize=(15, 15))
# <ul>
# <li>For any that look intesrting, we can draw a bigger scatter plot.
# </li>
# </ul>
plot = scatterplot(x="flarea", y="bdrms", data=copy_df)
# <ul>
# <li>We can compute correlations between numeric-valued columns.</li>
# <li>Most common is Pearson correlation, which measures linear correlation. It value lies between +1 and −1.
# A value of +1 is total positive linear correlation, 0 is no linear correlation, and −1 is total
# negative linear correlation.
# </li>
# </ul>
copy_df.corr()
# <ul>
# <li>We see that $\mathit{flarea}$ is strongly predictive of $\mathit{price}$.</li>
# <li>We see that $\mathit{flarea}$, $\mathit{bdrms}$ and $\mathit{bthrms}$ are quite strongly correlated
# with each other.
# </li>
# <li>We see that $\mathit{bdrms}$ and $\mathit{floors}$ are fairly strongly correlated.</li>
# <li>We can add other features, ones that are computed from the existing features — later, we call this
# <b>feature engineering</b>.
# <ul>
# <li>In general, these new features might be products or ratios of existing features.</li>
# <li>Or they might result from applying functions to existing features, e.g. squaring, square rooting,
# taking the log, …
# </li>
# <li>If you're going to be learning a linear model then it will not be useful to the linear model to add or subtract features or to multiply or divide by a scalar,
# since this just gives new features that are linearly correlated with existing features.
# </li>
# </ul>
# We can then produce visualizations and compute correlations to
# see whether these new features are predictive or not.
# </li>
# </ul>
copy_df["room_size"] = copy_df["flarea"] / (copy_df["bdrms"] + copy_df["bthrms"])
copy_df[["room_size", "flarea", "bdrms", "bthrms", "price"]].corr()
# <ul>
# <li>E.g. this new feature is quite predictive of price, but highly correlated with floor area.</li>
# </ul>
# <h1>Dataset Preparation</h1>
# <ul>
# <li>We need to prepare the dataset so that it is suitable for machine learning algorithms.</li>
# <li>An incomplete lists of activities follows.</li>
# <li>Missing values:
# <ul>
# <li>Most learning algorithms cannot handle missing values.</li>
# <li>We may have removed most of them during data cleaning.</li>
# <li>If any remain, we need a method to <b>impute</b> a value.
# <ul>
# <li>For numeric-valued features, we can replace missing values by the mean, for example.
# (In scikit-learn, use the <code>SimpleImputer</code> with <code>strategy="mean"</code>,
# which is the default.)
# </li>
# <li>For nominal-valued features, we can replace by the mode.
# (<code>strategy="most_frequent"</code>)
# </li>
# <li>There are other possibilities, e.g.: replace by some constant; binarize the feature
# (0 if the value was missing, 1 if not); predict the value from the other features.
# </li>
# </ul>
# </li>
# </ul>
# </li>
# <li>Scaling numeric-valued features:
# <ul>
# <li>Some algorithms (e.g. kNN) perform less well if features have different ranges; others
# (e.g. linear regression done using the normal equation) work equally well whether the
# data is scaled or not.
# <li>We've mentioned two ways to scale: min-max scaling and standardization. There are others.</li>
# </ul>
# </li>
# <li>Feature engineering:
# <ul>
# <li>In this module we use the phrase <b>feature engineering</b> for the following:
# augmenting the dataset with features that are computed from the other features.
# </li>
# <li>These may have been identified during dataset exploration above.</li>
# </ul>
# </li>
# <li>Feature selection:
# <ul>
# <li>There are methods for removing features that have low predictive power.
# </ul>
# </li>
# <li>Dimensionality reduction:
# <ul>
# <li>An example of this is Principal Component Analysis. Ignoring the details, it transforms
# the features into new features that are not linearly correlated with one another
# and enables us to discard the new features that contribute least.
# </li>
# </ul>
# </li>
# <li>Handling nominal-valued features:
# <ul>
# <li>Non-numeric features deserve separate explanation below.
# </ul>
# </li>
# </ul>
# <h2>Handling Nominal-Valued Features</h2>
# <ul>
# <li>Most AI algorithms work only with numeric-valued features.</li>
# <li>So, we will look at how to convert nominal-valued features to numeric-valued ones.</li>
# </ul>
# <h3>Binary-valued features</h3>
# <ul>
# <li>The simplest case, obviously, is a binary-valued feature, such as $\mathit{devment}$.</li>
# <li>We encode one value as 0 and the other as 1, e.g. "SecondHand" is 0 and "New" is 1.</li>
# </ul>
# <h3>Unordered nominal values</h3>
# <ul>
# <li>Suppose there are more than two values, e.g. Apartment, Detached, Semi-detached or Terraced.</li>
# <li>The obvious thing to do is to assign integers to each nominal value, e.g. 0 = Apartment, 1 = Detached,
# 2 = Semi-detached and 3 = Terraced.
# </li>
# <li>But often this is not the best encoding.
# <ul>
# <li>Algorithms may assume that the values themselves are meaningful, when they're actually arbitrary.
# <ul>
# <li>E.g. an algorithm might assume that Apartments (0) are more similar to Detached houses (1)
# than they are to Terraced houses (3).
# </li>
# </ul>
# </li>
# </ul>
# </li>
# <li>Instead, we use <b>one-hot encoding</b>.</li>
# </ul>
# <h3>One-Hot Encoding</h3>
# <ul>
# <li>If the original nominal-valued feature has $p$ values, then we use $p$ binary-valued features:
# <ul>
# <li>In each example, exactly one of them is set to 1 and the rest are zero.</li>
# </ul>
# </li>
# <li>For example, there are four types of dwelling, so we have four binary-valued features:
# <ul>
# <li>the first is set to 1 if and only if the type of dwelling is Apartment;</li>
# <li>the second is set to 1 if and only if the house is Detached;</li>
# <li>and so on.</li>
# </ul>
# So a detached house will be rpresented by $\rv{0, 1, 0, 0}$.
# </li>
# <li>
# In practice, it is not uncommon to be given a dataset where a nominal-valued feature has already been
# encoded numerically, one integer per value. You might be fooled into thinking that the feature is
# numeric-valued and overlook the need to use one-hot encoding on it. Watch out for this!
# </li>
# <li>One-hot encoding turns each $p$-valued non-numeric feature into $p$ binary-valued features. Thus, you
# can end up with a lot of features. Maybe you need to do some dimensionality reduction afterwards.
# </li>
# </ul>
# <h2>Data preprocessing in scikit-learn</h2>
# <ul>
# <li>Clearly, we want to automate all these data preparation activities.</li>
# <li>In scikit-learn, it is best to bring all of them together using a <code>ColumnTransformer</code>,
# <code>Pipelines</code> and, sometimes, <code>FeatureUnions</code>.
# </li>
# </ul>
# Create the preprocessor
preprocessor = ColumnTransformer([
("scaler", StandardScaler(),
numeric_features),
("nom", Pipeline([("imputer", SimpleImputer(missing_values=np.nan, strategy="most_frequent")),
("binarizer", OneHotEncoder(handle_unknown="ignore"))]),
nominal_features)],
remainder="passthrough")
# <ul>
# <li>Let's say we want to do some feature engineering. We could add extra features in the way we did
# when we were exploring the dataset, e.g.
# <pre>
# df["room_size"] = df["flarea"] / (df["bdrms"] + df["bthrms"])
# </pre>
# The problem is that this code is separate from our <code>ColumnTransformer</code>.
# And this leads to complications: for example, if we save our model, then this part of it won't be saved.
# If we can make it part of the <code>ColumnTransformer</code>, then it wll be saved along
# with everything else.
# </li>
# <li>The solution is to write our own transformer.</li>
# </ul>
class InsertRoomSize(BaseEstimator, TransformerMixin):
def __init__(self, insert=True):
self.insert = insert
def fit(self, X, y=None):
return self
def transform(self, X, y=None):
if self.insert:
X["room_size"] = X["flarea"] / (X["bdrms"] + X["bthrms"])
# If the new feature is intended to replace the existing ones,
# you could drop the existing ones here
# X.drop(["flarea", "bthrms", "bdrms"], axis=1)
X = X.replace( [ np.inf, -np.inf ], np.nan )
return X
# <ul>
# <li>Now our preprocessor might look like this:</li>
# </ul>
preprocessor = ColumnTransformer([
("num", Pipeline([("room_size", InsertRoomSize()),
("imputer", SimpleImputer(missing_values=np.nan, strategy="mean")),
("scaler", StandardScaler())]),
numeric_features),
("nom", Pipeline([("imputer", SimpleImputer(missing_values=np.nan, strategy="most_frequent")),
("binarizer", OneHotEncoder(handle_unknown="ignore"))]),
nominal_features)],
remainder="passthrough")
# <h1>Prepare for Model Selection</h1>
# <ul>
# <li>We have already split off our test set.</li>
# <li>But now we decide how to do model selection (see previous lecture):
# <ul>
# <li>Should we just split the remaining data into a training set and validation set?</li>
# <li>Or should we use $k$-fold cross-validation?</li>
# </ul>
# As discussed, it depends on how much data you have.
# <ul>
# <li>Since our dataset is quite small, we'll use $k$-fold cross-validation.</li>
# </ul>
# </li>
# <li>We must also decide which models we will experiment with.
# <ul>
# <li>We'll use linear regression and kNN.</li>
# </ul>
# </li>
# <li>And for each model, we must set up a grid of hyperparameter values.
# <ul>
# <li>For kNN, we need to try some different values for $k$.</li>
# <li>But we did something clever earlier when creating our own feature engineering transformer.
# We included an argument called <code>insert</code>. If this is True, then
# we add the extra feature; if it is False, we do not. Hence, this is a hyperparameter.
# We can try both values to see whether adding the feature helps or not.
# </li>
# </ul>
# </li>
# </ul>
# +
# Extract the features but leave as a DataFrame
dev_X = dev_df[features]
test_X = test_df[features]
# Target values, converted to a 1D numpy array
dev_y = dev_df["price"].values
test_y = test_df["price"].values
# +
# Create a pipeline that combines the preprocessor with kNN
knn = Pipeline([
("preprocessor", preprocessor),
("predictor", KNeighborsRegressor())])
# Create a dictionary of hyperparameters for kNN
knn_param_grid = {"predictor__n_neighbors": [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11],
"preprocessor__num__room_size__insert": [True, False]}
# Create the grid search object which will find the best hyperparameter values based on validation error
knn_gs = GridSearchCV(knn, knn_param_grid, scoring="neg_mean_absolute_error", cv=10)
# Run grid search by calling fit
knn_gs.fit(dev_X, dev_y)
# Let's see how well we did
knn_gs.best_params_, knn_gs.best_score_
# +
# Create a pipeline that combines the preprocessor with ridge regression
ridge = Pipeline([
("preprocessor", preprocessor),
("predictor", Ridge())])
# Create a dictionary of hyperparameters for ridge regression
ridge_param_grid = {"preprocessor__num__room_size__insert": [True, False],
"predictor__alpha": [0, 45.0, 50.0, 55.0]}
# Create the grid search object which will find the best hyperparameter values based on validation error
ridge_gs = GridSearchCV(ridge, ridge_param_grid, scoring="neg_mean_absolute_error", cv=10)
# Run grid search by calling fit
ridge_gs.fit(dev_X, dev_y)
# Let's see how well we did
ridge_gs.best_params_, ridge_gs.best_score_
# -
# <ul>
# <li>In fact, this gives me the idea of writing a transformer which takes in another transformer.
# Sounds uselesss. But, now that gives me a way of having transformers as hyperparameters.
# </li>
# <li>The example below uses this to select between a <code>StandardScaler</code>, a <code>MinMaxScaler</code>
# and a <code>RobustScaler</code>.
# </li>
# </ul>
class TransformerFromHyperP(BaseEstimator, TransformerMixin):
def __init__(self, transformer=None):
self.transformer = transformer
def fit(self, X, y=None):
if self.transformer:
self.transformer.fit(X, y)
return self
def transform(self, X, y=None):
if self.transformer:
return self.transformer.transform(X)
else:
return X
preprocessor = ColumnTransformer([
("num", Pipeline([("room_size", InsertRoomSize()),
("imputer", SimpleImputer(missing_values=np.nan, strategy="mean")),
("scaler", TransformerFromHyperP())]),
numeric_features),
("nom", Pipeline([("imputer", SimpleImputer(missing_values=np.nan, strategy="most_frequent")),
("binarizer", OneHotEncoder(handle_unknown="ignore"))]),
nominal_features)],
remainder="passthrough")
# +
# Create a pipeline that combines the preprocessor with kNN
knn = Pipeline([
("preprocessor", preprocessor),
("predictor", KNeighborsRegressor())])
# Create a dictionary of hyperparameters for kNN
knn_param_grid = {"predictor__n_neighbors": [8, 9, 10],
"preprocessor__num__room_size__insert": [True, False],
"preprocessor__num__scaler__transformer": [StandardScaler(), MinMaxScaler(), RobustScaler()]}
# Create the grid search object which will find the best hyperparameter values based on validation error
knn_gs = GridSearchCV(knn, knn_param_grid, scoring="neg_mean_absolute_error", cv=10)
# Run grid search by calling fit
knn_gs.fit(dev_X, dev_y)
# Let's see how well we did
knn_gs.best_params_, knn_gs.best_score_
# +
# Create a pipeline that combines the preprocessor with ridge regression
ridge = Pipeline([
("preprocessor", preprocessor),
("predictor", Ridge())])
# Create a dictionary of hyperparameters for rideg regression
ridge_param_grid = {"preprocessor__num__room_size__insert": [True, False],
"preprocessor__num__scaler__transformer": [StandardScaler(), MinMaxScaler(), RobustScaler()],
"predictor__alpha": [0, 45.0, 50.0, 55.0]}
# Create the grid search object which will find the best hyperparameter values based on validation error
ridge_gs = GridSearchCV(ridge, ridge_param_grid, scoring="neg_mean_absolute_error", cv=10)
# Run grid search by calling fit
ridge_gs.fit(dev_X, dev_y)
# Let's see how well we did
ridge_gs.best_params_, ridge_gs.best_score_
# -
# <h1>Tweak and Fine Tune</h1>
# <ul>
# <li>Now tweak and tune your model(s).
# <ul>
# <li>E.g. go back and change the grid of hyperparameter values.</li>
# <li>E.g. go back and add or remove transformers.</li>
# <li>E.g. go back and try regressors other than linear regression and kNN.</li>
# <li>E.g. you could even go back to the Dataset Acquisition step: maybe try to collect
# more features, for example.
# </li>
# </ul>
# </li>
# <li>You don't have to do this wholly at random. You can be guided in part by knowing whether you
# are underfitting or overfitting.
# <ul>
# <li>E.g. if you are underfitting, then collecting new features or doing feature engineering may help.
# </li>
# <li>But if you are overfitting then selecting among your features may be better.
# </li>
# </ul>
# See previous two lectures.
# </li>
# </ul>
knn.set_params(**knn_gs.best_params_)
scores = cross_validate(knn, dev_X, dev_y, cv=10,
scoring="neg_mean_absolute_error", return_train_score=True)
print("Training error: ", np.mean(np.abs(scores["train_score"])))
print("Validation error: ", np.mean(np.abs(scores["test_score"])))
ridge.set_params(**ridge_gs.best_params_)
scores = cross_validate(ridge, dev_X, dev_y, cv=10,
scoring="neg_mean_absolute_error", return_train_score=True)
print("Training error: ", np.mean(np.abs(scores["train_score"])))
print("Validation error: ", np.mean(np.abs(scores["test_score"])))
# <ul>
# <li>You can tweak and tune as much as you like!
# Keep doing this until you find a good model.
# </li>
# <li>Of course, if you're not careful, you'll overdo it and produce a model that overfits the training
# data.
# </li>
# <li>The main thing you shouldn't do, of course, is look at the test set until you have finished all
# this tweaking and tuning.
# </li>
# </ul>
# <h1>Evaluate on the Test Set</h1>
# <ul>
# <li>Once you've done, then, using the best model(s) we found, we can do error estimation on the test set.</li>
# </ul>
# +
# Now we re-train on train+validation and test on the test set
knn.set_params(**knn_gs.best_params_)
knn.fit(dev_X, dev_y)
mean_absolute_error(test_y, knn.predict(test_X))
# -
# <h1>Deploy</h1>
# <ul>
# <li>Make a decision: is this predictor good enough for real use?</li>
# <li>If it is, then re-train on the entire dataset, save the model and use it in your web app.</li>
# </ul>
knn.fit(df[features], df["price"].values)
dump(knn, 'models/my_model.pkl') # For this to work, create a folder called models!
# <ul>
# <li>Of course, even then your work is not finished:
# <ul>
# <li>You need to monitor peformance.</li>
# <li>You may need to re-train it when fresh data becomes available.</li>
# <li>And so on.</li>
# </ul>
# </li>
# </ul>
| ai1/lectures/AI1_12_regression_case_study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import glob
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.preprocessing.image import ImageDataGenerator, load_img, img_to_array, array_to_img
train_files = glob.glob('training_data/*')
train_files[:10]
# +
IMG_DIM = (150, 150)
train_files = glob.glob('training_data/*')
train_imgs = [img_to_array(load_img(img, target_size=IMG_DIM)) for img in train_files]
train_imgs = np.array(train_imgs)
train_labels = [fn.split('/')[1].split('.')[0].strip() for fn in train_files] # for linux
#train_labels = [fn.split('\\')[1].split('.')[0].strip() for fn in train_files] # for windows
validation_files = glob.glob('validation_data/*')
validation_imgs = [img_to_array(load_img(img, target_size=IMG_DIM)) for img in validation_files]
validation_imgs = np.array(validation_imgs)
validation_labels = [fn.split('/')[1].split('.')[0].strip() for fn in validation_files] # for linux
#validation_labels = [fn.split('\\')[1].split('.')[0].strip() for fn in validation_files] # for windows
print('Train dataset shape:', train_imgs.shape,
'\tValidation dataset shape:', validation_imgs.shape)
# +
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
le.fit(train_labels)
# encode wine type labels
train_labels_enc = le.transform(train_labels)
validation_labels_enc = le.transform(validation_labels)
print(train_labels[0:5], train_labels_enc[0:5])
# +
train_datagen = ImageDataGenerator(rescale=1./255, zoom_range=0.3, rotation_range=50,
width_shift_range=0.2, height_shift_range=0.2, shear_range=0.2,
horizontal_flip=True, fill_mode='nearest')
val_datagen = ImageDataGenerator(rescale=1./255)
# -
img_id = 2595
cat_generator = train_datagen.flow(train_imgs[img_id:img_id+1], train_labels[img_id:img_id+1],
batch_size=1)
cat = [next(cat_generator) for i in range(0,5)]
fig, ax = plt.subplots(1,5, figsize=(16, 6))
print('Labels:', [item[1][0] for item in cat])
l = [ax[i].imshow(cat[i][0][0]) for i in range(0,5)]
img_id = 1991
dog_generator = train_datagen.flow(train_imgs[img_id:img_id+1], train_labels[img_id:img_id+1],
batch_size=1)
dog = [next(dog_generator) for i in range(0,5)]
fig, ax = plt.subplots(1,5, figsize=(15, 6))
print('Labels:', [item[1][0] for item in dog])
l = [ax[i].imshow(dog[i][0][0]) for i in range(0,5)]
# +
train_generator = train_datagen.flow(train_imgs, train_labels_enc, batch_size=30)
val_generator = val_datagen.flow(validation_imgs, validation_labels_enc, batch_size=20)
input_shape = (150, 150, 3)
# +
from tensorflow.keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout
from tensorflow.keras.models import Sequential
from tensorflow.keras import optimizers
model = Sequential()
model.add(Conv2D(16, kernel_size=(3, 3), activation='relu',
input_shape=input_shape))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(128, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(128, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.3))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.3))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-4),
metrics=['accuracy'])
model.summary()
# -
history = model.fit_generator(train_generator, steps_per_epoch=100, epochs=100,
validation_data=val_generator, validation_steps=50, verbose=1)
# +
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
t = f.suptitle('CNN with Image Augmentation Performance', fontsize=12)
f.subplots_adjust(top=0.85, wspace=0.3)
epoch_list = list(range(1,101))
ax1.plot(epoch_list, history.history['acc'], label='Train Accuracy')
ax1.plot(epoch_list, history.history['val_acc'], label='Validation Accuracy')
ax1.set_xticks(np.arange(0, 101, 10))
ax1.set_ylabel('Accuracy Value')
ax1.set_xlabel('Epoch')
ax1.set_title('Accuracy')
l1 = ax1.legend(loc="best")
ax2.plot(epoch_list, history.history['loss'], label='Train Loss')
ax2.plot(epoch_list, history.history['val_loss'], label='Validation Loss')
ax2.set_xticks(np.arange(0, 101, 10))
ax2.set_ylabel('Loss Value')
ax2.set_xlabel('Epoch')
ax2.set_title('Loss')
l2 = ax2.legend(loc="best")
# -
model.save('cats_dogs_cnn_img_aug.h5')
| Chapter05/CNN with Image Augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Building a Chatbot with NLP and GRU model
# ### Importing the necessary libraries
import tensorflow as tf
import numpy as np
import json
import re
physical_devices = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], enable = True)
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input, Embedding, Dense, GRU, LSTM, Masking
from keras.preprocessing.text import tokenizer_from_json
# ### Importing preprocessed data
# +
with open('./preprocessed_data/questions.json', 'r') as f:
json_data = json.load(f)
question_corpus = tokenizer_from_json(json_data)
f.close()
with open('./preprocessed_data/answers.json', 'r') as f:
json_data = json.load(f)
answer_corpus = tokenizer_from_json(json_data)
f.close()
npzfile = np.load('./preprocessed_data/data.npz')
# -
# #### Some unknown reason make the corpus contain all words and labels in raw data rather than a vocabulary with limited size so we have to build the dict manually
q_word2ind={e:i for e, i in question_corpus.word_index.items() if i <= 8000}
q_ind2word={e:i for i, e in q_word2ind.items()}
a_word2ind={e:i for e, i in answer_corpus.word_index.items() if i <= 8000}
a_ind2word={e:i for i, e in a_word2ind.items()}
# ### Creating the encoder
def create_encoder(inputdim, embeddingsize, inputlen, n_units):
encoder_input = Input((inputlen,))
encoder_embed = Embedding(inputdim + 1, embeddingsize)(encoder_input)
encoder_mask = Masking()(encoder_embed)
encoder = GRU(n_units, return_state = True)
_, encoder_state = encoder(encoder_mask)
encoder = Model(encoder_input, encoder_state)
return encoder
# ### Creating the decoder
def create_decoder(inputdim, embeddingsize, n_units):
# The size of input at here is 1 because we want to predict the answer step by step, each time only input 1 word
decoder_input = Input((1,))
# Input of encoder state vectors
initial_state = Input((n_units,))
# Vectorizing input answers
decoder_embed = Embedding(inputdim + 1, embeddingsize)(decoder_input)
decoder_mask = Masking()(decoder_embed)
decoder = GRU(n_units, return_sequences = True, return_state = True)
# In inference model, we need decoder state
decoder_output, decoder_state = decoder(decoder_mask, initial_state = initial_state)
# Using activation function as softmax layer, predict the most potential sentence of reply
decoder_dense = Dense(inputdim, activation = 'softmax')
decoder_output_ = decoder_dense(decoder_output)
decoder = Model([decoder_input, initial_state], [decoder_output_, decoder_state])
return decoder
# ### Defining hyperparameters
n_unit = 256
embedding_size = 128
vocab_size = 8000
question_len = npzfile['arr_0'].shape[1]
answer_len = npzfile['arr_1'].shape[1]
encoder = create_encoder(vocab_size, embedding_size, question_len, n_unit)
encoder.summary()
encoder.load_weights('./trained_model/gru_encoder_test.h5')
decoder = create_decoder(vocab_size, embedding_size, n_unit)
decoder.summary()
decoder.load_weights('./trained_model/gru_decoder_test.h5')
def clean_text(text):
# remove unnecessary characters in sentences
text = text.lower().strip()
text = re.sub(r"i'm", "i am", text)
text = re.sub(r"he's", "he is", text)
text = re.sub(r"she's", "she is", text)
text = re.sub(r"it's", "it is", text)
text = re.sub(r"that's", "that is", text)
text = re.sub(r"what's", "what is", text)
text = re.sub(r"where's", "where is", text)
text = re.sub(r"there's", "there is", text)
text = re.sub(r"how's", "how is", text)
text = re.sub(r"\'ll", " will", text)
text = re.sub(r"\'ve", " have", text)
text = re.sub(r"\'re", " are", text)
text = re.sub(r"\'d", " would", text)
text = re.sub(r"\'re", " are", text)
text = re.sub(r"won't", "will not", text)
text = re.sub(r"can't", "cannot", text)
text = re.sub(r"n't", " not", text)
text = re.sub(r"n'", "ng", text)
text = re.sub(r"'bout", "about", text)
text = re.sub(r"'til", "until", text)
text = re.sub(r'[" "]+', " ", text)
text = re.sub(r"[-()\"#/@;:<>{}`+=~|.!?,]", "", text)
return text
# ### Evaluating the chat
def evaluate(sentence):
sentence = clean_text(sentence) # Clean the input text
encoder_inputs = []
# Converting the input text to index sequence and use unk replace the word not in vocabulary
for word in sentence.split():
if word in q_word2ind:
encoder_inputs.append(q_word2ind[word])
elif word not in q_word2ind:
encoder_inputs.append(q_word2ind['unk'])
# Initializing the encoder input
encoder_inputs = tf.keras.preprocessing.sequence.pad_sequences([encoder_inputs], maxlen = question_len, padding = 'post')
encoder_inputs = tf.convert_to_tensor(encoder_inputs)
encoder_state = encoder(encoder_inputs)
# Initializing the decoder input
decoder_inputs = tf.expand_dims([a_word2ind['bos']], 0)
hidden_state = encoder_state
result = ''
for t in range(answer_len):
pred,state = decoder([decoder_inputs, hidden_state])
pred = np.squeeze(pred)
pred_ind = tf.math.argmax(pred).numpy() + 1
# Once we get the 'eos' symbol, stop the loop
if a_ind2word[pred_ind] == 'eos':
return result
result += a_ind2word[pred_ind] + ' '
# Passing the predict index and state vectors to the next input
decoder_inputs = tf.expand_dims([pred_ind], 0)
hidden_state = state
return result
while True:
inputs = input('User :> ')
if inputs == 'quit':
break
result = evaluate(inputs)
print('Bot :> ' + result)
| gru_bot.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Lab Two
// ---
//
// Ok for this lab we're going to reiterate a lot of the things that we went over in class.
//
// Our Goals are:
// - Conditionals
// - If
// - Else
// - Else If
// +
// Make an if statement
if (true) {
System.out.println("Welcome to Software Dev I");
}
// +
// Make an if, else statement where the else statement triggers
if (false) {
System.out.println("Welcome to Software Dev I");
} else {
System.out.println("Coding is fun");
}
// +
// Make an if, else if, else statement where the else if statement triggers
if (!true) {
System.out.println("Welcome to Software Dev I");
} else if (true){
System.out.println("Marist 2024");
} else {
System.out.println("Coding is fun");
}
// -
// Make 2 variables and use them in an if else conditional
int x = 5;
int y = 10;
if (x+y > 13) {
System.out.println("It is big enough");
} else {
System.out.println("it is too small");
}
// +
// Make an if statement using 2 variables and an AND(&&) statement
boolean cheese = true;
boolean crackers = true;
if (cheese && crackers) {
System.out.println("I can make a cheese and crackers");
} else {
System.out.println("I cannot make cheese and crackers");
}
// +
// Make an if statement using 2 variables and an OR(||) statement
boolean blucheese = true;
boolean ranch = true;
if (!blucheese || ranch) {
System.out.println("My wings have sauce!");
} else {
System.out.println("My wings are dry");
}
| JupyterNotebooks/Labs/Lab 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hough Lines
# ### Import resources and display the image
# +
import numpy as np
import matplotlib.pyplot as plt
import cv2
# %matplotlib inline
# Read in the image
image = cv2.imread('images/phone.jpg')
# Change color to RGB (from BGR)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
plt.imshow(image)
# -
# ### Perform edge detection
# +
# Convert image to grayscale
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
# Define our parameters for Canny
low_threshold = 50
high_threshold = 100
edges = cv2.Canny(gray, low_threshold, high_threshold)
plt.imshow(edges, cmap='gray')
# -
# ### Find lines using a Hough transform
# +
# Define the Hough transform parameters
# Make a blank the same size as our image to draw on
rho = 1
theta = np.pi/180
threshold = 60
min_line_length = 100
max_line_gap = 5
line_image = np.copy(image) #creating an image copy to draw lines on
# Run Hough on the edge-detected image
lines = cv2.HoughLinesP(edges, rho, theta, threshold, np.array([]),
min_line_length, max_line_gap)
# Iterate over the output "lines" and draw lines on the image copy
for line in lines:
for x1,y1,x2,y2 in line:
cv2.line(line_image,(x1,y1),(x2,y2),(255,0,0),5)
plt.imshow(line_image)
# -
| Introduction to Computer Vision/Filters and Edge Detection/Hough lines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 파이썬 scipy.stats
#
# > "작성 완료"
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - author: 한재수
# - categories: [python, statistics]
# `-` 확률분포를 scipy.stats를 통해 그리는데 익숙하지 않아 기본 사용에 대해 알아볼 거임
#
# `-` 참고: [scipy.stats](https://docs.scipy.org/doc/scipy/reference/stats.html)
#
# `-` 참고: [scipy 확률분포](https://datascienceschool.net/02%20mathematics/08.01%20%EC%82%AC%EC%9D%B4%ED%8C%8C%EC%9D%B4%EB%A5%BC%20%EC%9D%B4%EC%9A%A9%ED%95%9C%20%ED%99%95%EB%A5%A0%EB%B6%84%ED%8F%AC%20%EB%B6%84%EC%84%9D.html)
# # 확률분포 클래스
# `-` scipy.stats를 통해 확률분포를 그려보자
#
# `-` 우선 확률분포에 대한 클래스 객체를 생성해야 함
#
# `-` 각 확률분포의 파라미터는 `scipy.stats.이름`을 통해 확인하자
#
# |종류|이름|확률분포|
# |:-:|:-:|:-:|
# |이산 |bernoulli |베르누이 분포|
# |이산|binom |이항 분포|
# |이산|poisson|포아송 분포|
# |이산|geom|기하 분포|
# |이산|nbinom|음이항 분포|
# |이산|hypergeom|초기하 분포|
# |이산 |multinomial |다항 분포|
# |연속 |norm |정규 분포|
# |연속 | uniform|균일 분포|
# |연속|expon|지수 분포|
# |연속 |gamma |감마 분포|
# |연속 |t |t 분포|
# | 연속|chi2 |카이제곱 분포|
# | 연속|f |f 분포|
# |연속 |beta |베타 분포|
# # 모수 지정
# `-` 확률분포의 모수는 종류별로 다르므로 문서를 참고하자
#
# `-` 하지만 대부분 확률분포가 공통적으로 가지는 모수가 있음
#
# |모수 이름|의미|
# |:-:|:-:|
# |loc|기댓값|
# |scale|표준편차|
# # 확률분포 methods
# `-` 확률분포 클래스 객체가 가지는 method가 있음
#
# `-` 정규분포를 예로 들어 ppf에 대한 설명을 해보면 norm.ppf(0.5)는 정규분포에서 $50$분위수에 해당하는 $x$값으로 $0$이다
#
# |메서드|기능|
# |:-:|:-:|
# |pmf|확률질량함수|
# |pdf|확률밀도함수|
# |cdf |누적분포함수 |
# |ppf |누적분포함수의 역함수(백분위 함수) |
# | sf|생존함수 = 1 $-$ 누적분포함수 |
# |isf |생존함수의 역함수 |
# |rvs |무작위 표본 생성 |
# # 확률분포 plot
# ## 정규 분포 pdf
# `-` 정규 분포 pdf를 그려보자
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
xx = np.linspace(-5, 5, 1000)
for scale in (0.5, 1.0, 2.0):
plt.plot(xx, norm(0, scale).pdf(xx), label = 'μ = 0, σ = ' + str(scale), lw = 2, alpha = 0.8)
plt.plot(xx, norm(-2, 0.5).pdf(xx), label = 'μ = -2, σ = 0.5', lw = 2, alpha = 0.8)
plt.xticks(np.arange(-5, 6))
plt.yticks(np.arange(0.0, 1.2, 0.2))
plt.title("normal distribution pdf")
plt.xlabel("$x$")
plt.ylabel("$f(x)$")
plt.grid()
plt.legend()
plt.show()
# -
# ## 정규 분포 cdf
# `-` 정규 분포 cdf를 그려보자
#
# `-` cdf에 대한 내용 정리 예정
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
xx = np.linspace(-5, 5, 1000)
for scale in (0.5, 1.0, 2.0):
plt.plot(xx, norm(0, scale).cdf(xx), label = 'μ = 0, σ = ' + str(scale), lw = 2, alpha = 0.8)
plt.plot(xx, norm(-2, 0.5).cdf(xx), label = 'μ = -2, σ = 0.5', lw = 2, alpha = 0.8)
plt.xticks(np.arange(-5, 6))
plt.yticks(np.arange(0.0, 1.2, 0.2))
plt.title("normal distribution cdf")
plt.xlabel("$x$")
plt.ylabel("$f(x)$")
plt.grid()
plt.legend()
plt.show()
# -
| _notebooks/2021-08-10-scipy-stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Test att hämta data [släktdata](https://www.slaktdata.org)
# * [Denna notebook](https://github.com/salgo60/Slaktdata/blob/master/Slaktdata.ipynb)
#
from datetime import datetime
start_time = datetime.now()
print("Last run: ", start_time)
# +
import urllib3, json
import pandas as pd
from pandas.io.json import json_normalize
http = urllib3.PoolManager()
urlbase= "https://www.slaktdata.org"
url = urlbase + "/?p=getregbyid&sldid=156206_F7_710"
print(url)
r = http.request('GET', urlbase + "?p=getregbyid&sldid=156206_F7_710",
headers={'Content-Type': 'application/json'})
df=pd.read_json(r.data)
df.columns
# -
#df["res"].to_frame().T
listscb = (113701, 138224, 140103, 141501, 141502,141503,141504,142703)
listbook = ("F","V","H","J","M")
# +
from tqdm.notebook import trange
dfList = []
for scb in listscb:
for book in listbook:
for booknr in range(1,13):
for slakdatanr in trange(0,1000,10): #test
url = urlbase + "/?p=getregbyid&sldid=" + str(scb) + "_" + str(book) + str(booknr) + "_" + str(slakdatanr)
#print (url,len(r.data))
r = http.request('GET', url,
headers={'Content-Type': 'application/json'})
if len(r.data) > 400:
#print (url,len(r.data))
df=pd.read_json(r.data)
dfList.append(df["res"].to_frame().T)
print(url)
dfTot = pd.concat(dfList, ignore_index=True)
dfTot.info()
# -
dfTot.shape
dfTot
dfTot["adexkluderat"].value_counts()
dfTot["adid"].value_counts()
dfTot["adress"].value_counts()
dfTot["enamn"].value_counts()
dfTot["fnamn"].value_counts()
dfTot["ovr1"].value_counts()
dfTot["scbkod"].value_counts()
dfTot["sdsuffix"].value_counts()
dfTot["sidnr"].value_counts()
dfTot["uppdaterat"].value_counts()
dfTot["web"].value_counts()
| Jupyter/Slaktdata.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Intelligence II - Team MensaNord
# ## Sheet 11
# - <NAME>
# - <NAME>
# - <NAME>
# - <NAME>
# - <NAME>
from __future__ import division, print_function
import matplotlib.pyplot as plt
# %matplotlib inline
import scipy.stats
import numpy as np
from scipy.ndimage import imread
import sys
# ### Exercise 1
# - Load the data into a vector and normalize it such that the values are between 0 and 1.
# - Create two new datasets by adding Gaussian noise with zero mean and standard deviation
# σ N ∈ {0.05, 0.1}.
# +
# import image
img_orig = imread('testimg.jpg').flatten()
print("$img_orig")
print("shape: \t\t", img_orig.shape) # = vector
print("values: \t from ", img_orig.min(), " to ", img_orig.max(), "\n")
# "img" holds 3 vectors
img = np.zeros((3,img_orig.shape[0]))
print("$img")
print("shape: \t\t",img.shape)
std = [0, 0.05, 0.1]
for i in range(img.shape[1]):
# normalize => img[0]
img[0][i] = img_orig[i] / 255
# gaussian noise => img[1] img[2]
img[1][i] = img[0][i] + np.random.normal(0, std[1])
img[2][i] = img[0][i] + np.random.normal(0, std[2])
print(img[:, 0:4])
# -
# - Create a figure showing the 3 histograms (original & 2 sets of noise corrupted data – use
# enough bins!). In an additional figure, show the three corresponding empirical distribution
# functions in one plot.
# histograms
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
for i, ax in enumerate(axes.flatten()):
plt.sca(ax)
plt.hist(img[i], 100, normed=1, alpha=0.75)
plt.xlim(-0.1, 1.1)
plt.ylim(0, 18)
plt.xlabel("value")
plt.ylabel("probability")
plt.title('img[{}]'.format(i))
# +
# divide probablity space in 100 bins
nbins = 100
bins = np.linspace(0, 1, nbins+1)
# holds data equivalent to shown histograms (but cutted from 0 to 1)
elementsPerBin = np.zeros((3,nbins))
for i in range(3):
ind = np.digitize(img[i], bins)
elementsPerBin[i] = [len(img[i][ind == j]) for j in range(nbins)]
# counts number of elements from bin '0' to bin 'j'
sumUptoBinJ = np.asarray([[0 for i in range(nbins)] for i in range(3)])
for i in range(3):
for j in range(nbins):
sumUptoBinJ[i][j] = np.sum(elementsPerBin[i][0:j+1])
# plot
plt.figure(figsize=(15, 5))
for i in range(3):
plt.plot(sumUptoBinJ[i], '.-')
plt.legend(['img[0]', 'img[1]', 'img[2]'])
plt.xlabel('bin')
plt.ylabel('empirical distribution functions');
# -
# - Take a subset of P = 100 observations and estimate the probability density p̂ of intensities
# with a rectangular kernel (“gliding window”) parametrized by window width h.
# - Plot the estimates p̂ resulting for (e.g. 10) different samples of size P
def H(vec, h):
"""
(rectangular) histogram kernel function
"""
vec = np.asarray(vec)
return np.asarray([1 if abs(x)<.5 else 0 for x in vec])
# ### $P(\underline{x}) = \frac{1}{h^n} \frac{1}{p} \Sigma_{\alpha=1}^{p} H(\frac{\underline{x} - \underline{x}^{(\alpha)}}{h})$
def P_est(x, h, data, kernel = H):
"""
returns the probability that data contains values @ (x +- h/2)
"""
n = 1 #= data.shape[1] #number of dimensions (for multidmensional data)
p = len(data)
return 1/(h**n)/p*np.sum(kernel((data - x)/h, h))
# take 10 data sets with 100 observations (indexes 100k to 101k)
# nomenclature: data_3(3, 10, 100) holds 3 times data(10, 100)
P = 100
offset = int(100000)
data_3 = np.zeros((3, 10,P))
for j in range(3):
for i in range(10):
data_3[j][i] = img[j][offset+i*P:offset+(i+1)*P]
print(data_3.shape)
# +
# calculate probability estimation for (center +- h/2) on the 10 data sets
h = .15
nCenters = 101
Centers = np.linspace(0,1,nCenters)
fig, ax = plt.subplots(2,5,figsize=(15,6))
ax = ax.ravel()
for i in range(10):
ax[i].plot([P_est(center,h,data_3[0][i]) for center in Centers])
# -
# - Calculate the negative log-likelihood per datapoint of your estimator using 5000
# samples from the data not used for the density estimation (i.e. the “test-set”). Get
# the average of the negative log-likelihood over the 10 samples.
# ### $P(\{\underline{x}^{(\alpha)}\};\underline{w}) = - \Sigma_{\alpha=1}^{p} ln P(\underline{x}^{(\alpha)};\underline{w})$
# +
testdata = img[0][50000:55000]
# calculate average negative log likelihood for
def avg_NegLL(data, h, kernel=H):
sys.stdout.write(".")
average = 0
for i in range(10):
L_prob = [np.log(P_est(x,h,data[i],kernel)) for x in testdata]
negLL = -1*np.sum(L_prob)
average += negLL
average /= 10
return average
# -
# 2) Repeat this procedure (without plotting) for a sequence of kernel widths h to get the mean
# log likelihood (averaged over the different samples) resulting for each value of h.
#
# (a) Apply this procedure to all 3 datasets (original and the two noise-corruped ones) to make
# a plot showing the obtained likelihoods (y-axis) vs. kernel width h (x-axis) as one line for
# each dataset.
# +
hs = np.linspace(0.001, 0.999, 20)
def plot_negLL(data_3=data_3, kernel=H):
fig = plt.figure(figsize=(12,8))
for j in range(3):
print("calc data[{}]".format(j))
LLs = [avg_NegLL(data_3[j],h,kernel=kernel) for h in hs]
plt.plot(hs,LLs)
print()
plt.legend(['img[0]', 'img[1]', 'img[2]'])
plt.show()
# -
plot_negLL()
# not plotted points have value = inf because:
#
# $negLL = - log( \Pi_\alpha P(x^\alpha,w) )$
#
# so if one single $P(x^\alpha,w) = 0$ occurs (x has 5000 elements)
#
# the result is -log(0)=inf (not defined)
#
# this only occurs with the histogram kernel.
# (b) Repeat the previous step (LL & plot) for samples of size P = 500.
# +
P = 500
data_3b = np.zeros((3, 10,P))
for j in range(3):
for i in range(10):
data_3b[j][i] = img[j][offset+i*P:offset+(i+1)*P]
plot_negLL(data_3=data_3b)
# -
# (c) Repeat the previous steps (a & b) for the Gaussian kernel with σ^2 = h.
def Gaussian(x,h):
"""
gaussian kernel function
"""
return np.exp(-x**2/h/2)/np.sqrt(2*np.pi*h)
# +
fig, ax = plt.subplots(2,5,figsize=(15,6))
h = .15
ax = ax.ravel()
for i in range(10):
ax[i].plot([P_est(center,h,data_3[0][i],kernel=Gaussian) for center in Centers])
# -
hs = np.linspace(0.001, 0.4, 20)
plot_negLL(kernel=Gaussian)
plot_negLL(data_3=data_3b, kernel=Gaussian)
# ## Exercise 2
# ### 1.1 Create dataset
# +
M = 2
w1, w2 = [2,2], [1,1] # means
sigma2 = 0.2 # standard deviations
N = 100
P1, P2 = 2/3, 1/3
def create_data(sigma1=0.7):
X = np.zeros((N, 2))
which_gaussian = np.zeros(N)
for n in range(N):
if np.random.rand() < P1: # sample from first Gaussian
X[n] = np.random.multivariate_normal(w1, np.eye(len(w1)) * sigma1**2)
which_gaussian[n] = 0
else: # sample from second Gaussian
X[n] = np.random.multivariate_normal(w2, np.eye(len(w2)) * sigma2**2)
which_gaussian[n] = 1
return X, which_gaussian
sigma1 = 0.7
X, which_gaussian = create_data(sigma1)
# +
def plot_data(X, which_gaussian, centers, stds):
plt.scatter(*X[which_gaussian == 0].T, c='r', label='Cluster 1')
plt.scatter(*X[which_gaussian == 1].T, c='b', label='Cluster 2')
plt.plot(centers[0][0], centers[0][1], 'k+', markersize=15, label='Centers')
plt.plot(centers[1][0], centers[1][1], 'k+', markersize=15)
plt.gca().add_artist(plt.Circle(centers[0], stds[0], ec='k', fc='none'))
plt.gca().add_artist(plt.Circle(centers[1], stds[1], ec='k', fc='none'))
plt.xlabel('x1')
plt.ylabel('x2')
plt.legend()
plot_data(X, which_gaussian, [w1, w2], [sigma1, sigma2])
plt.title('Ground truth')
# -
# ### 1.2 Run Expectation-Maximization algorithm
# See slide 18 of the lecture for an outline of the algorithm.
# +
from scipy.stats import multivariate_normal
def variance(X):
"""Calculate a single variance value for the vectors in X."""
mu = X.mean(axis=0)
return np.mean([np.linalg.norm(x - mu)**2 for x in X])
def run_expectation_maximization(X, w=None, sigma_squared=None, verbose=False):
# Initialization.
P_prior = np.ones(2) * 1 / M
P_likelihood = np.zeros((N, M))
P_posterior = np.zeros((M, N))
mu = X.mean(axis=0) # mean of the original data
var = variance(X) # variance of the original data
if w is None:
w = np.array([mu + np.random.rand(M) - 0.5, mu + np.random.rand(M) - 0.5])
if sigma_squared is None:
sigma_squared = np.array([var + np.random.rand() - 0.5,var + np.random.rand() - 0.5])
#sigma_squared = np.array([var, var])
if verbose:
print('Initial centers:', w)
print('Initial variances:', sigma_squared)
print()
print()
theta = 0.001
distance = np.inf
step = 0
# Optimization loop.
while distance > theta:
#for i in range(1):
step += 1
if verbose:
print('Step', step)
print('-'*50)
# Store old parameter values to calculate distance later on.
w_old = w.copy()
sigma_squared_old = sigma_squared.copy()
P_prior_old = P_prior.copy()
if verbose:
print('Distances of X[0] to proposed centers:', np.linalg.norm(X[0] - w[0]), np.linalg.norm(X[0] - w[1]))
# E-Step: Calculate likelihood for each data point.
for (alpha, q), _ in np.ndenumerate(P_likelihood):
P_likelihood[alpha, q] = multivariate_normal.pdf(X[alpha], w[q], sigma_squared[q])
if verbose:
print('Likelihoods of X[0]:', P_likelihood[0])
# E-Step: Calculate assignment probabilities (posterior) for each data point.
for (q, alpha), _ in np.ndenumerate(P_posterior):
P_posterior[q, alpha] = (P_likelihood[alpha, q] * P_prior[q]) / np.sum([P_likelihood[alpha, r] * P_prior[r] for r in range(M)])
if verbose:
print('Assignment probabilities of X[0]:', P_posterior[:, 0])
print()
distance = 0
# M-Step: Calculate new parameter values.
for q in range(M):
w[q] = np.sum([P_posterior[q, alpha] * X[alpha] for alpha in range(N)], axis=0) / np.sum(P_posterior[q])
#print(np.sum([P_posterior[q, alpha] * X[alpha] for alpha in range(N)], axis=0))
#print(np.sum(P_posterior[q]))
w_distance = np.linalg.norm(w[q] - w_old[q])
if verbose:
print('Distance of centers:', w_distance)
distance = max(distance, w_distance)
sigma_squared[q] = 1 / M * np.sum([np.linalg.norm(X[alpha] - w_old[q])**2 * P_posterior[q, alpha] for alpha in range(N)]) / np.sum(P_posterior[q])
sigma_squared_distance = np.abs(sigma_squared[q] - sigma_squared_old[q])
if verbose:
print('Distance of variances:', sigma_squared_distance)
distance = max(distance, sigma_squared_distance)
P_prior[q] = np.mean(P_posterior[q])
P_prior_distance = np.abs(P_prior[q] - P_prior_old[q])
if verbose:
print('Distance of priors:', P_prior_distance)
distance = max(distance, P_prior_distance)
if verbose:
print('Maximum distance:', distance)
print()
print('New centers:', w)
print('New variances:', sigma_squared)
print('New priors:', P_prior)
print('='*50)
print()
which_gaussian_EM = P_posterior.argmax(axis=0)
return which_gaussian_EM, w, np.sqrt(sigma_squared), step
which_gaussian_em, cluster_centers_em, cluster_stds_em, num_steps_em = run_expectation_maximization(X, verbose=True)
# -
plot_data(X, which_gaussian_em, cluster_centers_em, cluster_stds_em)
plt.title('Predicted by Expectation-Maximization')
# ### 1.3 Run K-means algorithm
# For simplicity, we use the sklearn version of K-means here. The detailed algorithm was already implemented in a previous exercise.
# +
from sklearn.cluster import KMeans
def run_k_means(X):
km = KMeans(2)
km.fit(X)
which_gaussian_km = km.predict(X)
cluster_stds = np.array([np.sqrt(variance(X[which_gaussian_km == 0])), np.sqrt(variance(X[which_gaussian_km == 1]))])
return which_gaussian_km, km.cluster_centers_, cluster_stds
which_gaussian_km, cluster_centers_km, cluster_stds_km = run_k_means(X)
plot_data(X, which_gaussian_km, cluster_centers_km, cluster_stds_km)
plt.title('Predicted by K-Means')
# -
# K-means clusters the data point by establishing a straight separation line. This cannot fully capture the nature of the data, e.g. the points around the lower left Gaussian, which actually belong to the upper right Gaussian.
# ### 1.4 Initialize EM algorithm with cluster parameters from K-Means
# +
_, _, _, num_steps_em_km = run_expectation_maximization(X, cluster_centers_km, cluster_stds_km**2)
print('Took', num_steps_em, 'steps with random initalization')
print('Took', num_steps_em_km, 'steps with initialization from K-means')
# -
# ### 1.5 Repeat analysis for different $\sigma_1$ values
# +
sigma1s = [0.1, 0.5, 1, 1.5]
fig, axes = plt.subplots(len(sigma1s), 3, figsize=(15, 15), sharex=True, sharey=True)
for i, (sigma1, horizontal_axes) in enumerate(zip(sigma1s, axes)):
X, which_gaussian = create_data(sigma1)
plt.sca(horizontal_axes[0])
plot_data(X, which_gaussian, [w1, w2], [sigma1, sigma2])
if i == 0:
plt.title('Ground truth')
which_gaussian_em, cluster_centers_em, cluster_stds_em, num_steps_em = run_expectation_maximization(X)
plt.sca(horizontal_axes[1])
plot_data(X, which_gaussian_em, cluster_centers_em, cluster_stds_em)
if i == 0:
plt.title('Predicted by Expectation-Maximization')
which_gaussian_km, cluster_centers_km, cluster_stds_km = run_k_means(X)
plt.sca(horizontal_axes[2])
plot_data(X, which_gaussian_km, cluster_centers_km, cluster_stds_km)
if i == 0:
plt.title('Predicted by K-Means')
# -
# Each row corresponds to increasing $\sigma_1$ (the values are 0.1, 0.5, 1, 1.5).
#
# K-means and Expectation-Maximization show similar results for small $\sigma_1$, i.e. if the clusters are clearly separated. With increasing $\sigma_1$, the Gaussians overlap more and more, and K-means fails to cluster them correctly.
| sheet11/sheet11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# [](https:pythonista.mx)
# ## Gestión de peticiones HTTP con *Requests*.
#
# El proyecto [*Requests*](http://docs.python-requests.org/en/master/) tiene como objetivo ofrecer una herramienta avanzada pero simple para realizar transacciones basadas en HTTP.
#
# Como su nombre lo indica, esta herramienta permite enviar peticiones (requests) a un servidor HTTP/1.1 y a su vez obtener las respuestas (responses) y mensajes enviados por dicho servidor.
# Entre otras cosas, *Requests* es capaz de:
#
# * Manejar cookies.
# * Levantar conexiones SSL.
# * Gestionar flujos (streams).
# * Enviar encabezados a la medida.
# * Autenticarse de forma avazanda.
# !pip install requests
import requests
help(requests)
# El paquete *requests* contiene funciones que emulan a los métodos propios de HTTP:
#
# * El método GET con *requests.get()*.
# * El método POST con *requests.post()*.
# * El método PUT con *requests.put()*.
# * El método PATCH con *requests.patch()*.
# * El método HEAD con *requests.head()*.
# * El método DELETE con *requests.delete()*.
# * El método CONNECT con *requests.connect()*.
# * El método OPTIONS con *requests.options()*.
#
# Cada una de estas funciones regresa un objeto con múltiples métodos y atributos con información sobre la comunicación entre el cliente y el servidor.
# ### El sitio https://httpbin.org.
#
# Este sitio ofrece un servidor de prueba que regresa determinados datos a partir de peticiones de diversas índoles.
#
# Se utilizará para ejemplificar algunos usos del módulo *requests*.
# ### La función *request.get()*.
# Esta función envía una petición utilizando el método GET con la siguiente sintaxis:
#
# ``` python
# requests.get('<URL>', params=<objeto tipo dict>)
# ```
# **Ejemplos:**
# Se utilizará *requests.get()* sin ningún parámetro adicional.
respuesta_get = requests.get('https://httpbin.org')
respuesta_get
dir(respuesta_get)
print(respuesta_get.headers)
print(respuesta_get.content)
print(respuesta_get.text)
print(respuesta_get.status_code)
respuesta_get.close()
# Se utilizarán algunos parámetros para *requests_get()*.
parametros = {'url': 'search-alias', 'field-keywords': 'python'}
respuesta_get = requests.get('https://amazon.com.mx', params=parametros)
print(respuesta_get.url)
print(respuesta_get.text)
# Se utilizará *request.get()* para obtener una imagen.
imagen = requests.get("https://httpbin.org/image/png")
print(imagen.content)
# Para desplegar una imagen se utilzará el módulo *Image* del paquete *iPython*.
from IPython.display import Image
Image(imagen.content)
imagen.close()
# ### La función *request.post()*.
# Esta función envía una petición utilizando el método PUT con la siguiente sintaxis:
#
# ``` python
# requests.post('<URL>', data=<objeto>, json=<objeto tipo dict>)
# ```
# **Ejemplo:**
import json
with requests.post('https://httpbin.org/post', data='Hola') as respuesta_post:
print(json.dumps(respuesta_post.json(), indent=2))
datos = {'nombre': 'Jose', 'apellido': 'Pérez'}
with requests.post('https://httpbin.org/post', json=datos) as respuesta_post:
print(respuesta_post.url)
print(json.dumps(respuesta_post.json(), indent=2))
# ## Acceso a API que requieren autenticación OAuth.
#
# ### Las bibliotecas *oauthlib* y *requests_oauthlib*.
# La biblioteca [*oauthlib*](https://oauthlib.readthedocs.io/en/latest/) es una biblioteca permite realizar la autenticación a un servidor hace uso del protocolo [OAuth](https://oauth.net/) para acceder a este. Sitios como Facebook, Twitter, Linkedin, GitHub y Google entre otros, utilizan este protocolo.
#
# Es posible acceder a sitios que utilizan dicho protocolo mediante el paquete *oauthlib* e incluso existe la biblioteca [*requests_oauthlib*](http://requests-oauthlib.readthedocs.io/en/latest/), la cual es una implementación de *requests* para acceder mediante este protocolo.
# En este ejemplo se hará una conexión a la API de Twitter, la cual utiliza la versión del protocolo OAuth 1.
# !pip install requests_oauthlib
import json
# * La función *credenciales_tw()* toma las credenciales de accceso a la API de twitter.
def credenciales_tw(fuente):
'''Regresa un objeto capaz de consumir la API de Twitter mediante
la lectura de un archivo que contiene las credenciales del desarrollador.'''
with open(fuente, 'r') as archivo:
(CONSUMER_KEY, CONSUMER_SECRET, OAUTH_TOKEN,
OAUTH_TOKEN_SECRET) = archivo.read().splitlines()
return CONSUMER_KEY, CONSUMER_SECRET, OAUTH_TOKEN, OAUTH_TOKEN_SECRET
# * Se ejecuta la función *credenciales_tw* y se le asigna el nombre correspondiente a cada credencial obtenida.
client_key, client_secret, resource_owner_key, resource_owner_secret = credenciales_tw('data/credenciales.txt')
# * Se importa la clase *OAuth1Session* de *requests_oauthlib* para iniciar la conexión con la API.
from requests_oauthlib import OAuth1Session
# * Se inicia la conexión y se le asigna el nombre *twitter* al objeto resultante.
twitter = OAuth1Session(client_key, client_secret, resource_owner_key, resource_owner_secret)
# * Se obtienen los datos de la cuenta del usuario al que pertencen las credenciales mediante el método GET, conforme a la especificación localizada en https://developer.twitter.com/en/docs/accounts-and-users/manage-account-settings/api-reference/get-account-settings
r = twitter.get('https://api.twitter.com/1.1/account/settings.json')
# * Se consultan algunos datos sobre la petición hecha a la API.
r.status_code
print(json.dumps(r.json(), indent=2))
print(r.url)
# * Se realiza una búsqueda de los tuits que coincidan con los parámetros especificados en el objeto tipo *dict* con nombre *elementos_busqueda* mediante el método GET conforme a la especificación loccalizada en https://developer.twitter.com/en/docs/tweets/search/api-reference/get-search-tweets
elementos_busqueda = {'q': "#python", 'count': 2, 'lang': 'es'}
busqueda = twitter.get('https://api.twitter.com/1.1/search/tweets.json', params=elementos_busqueda)
print(busqueda.url)
print(json.dumps(busqueda.json(), indent=2))
# * Se cierra la conexión.
twitter.close()
dir(r)
r.text
type(busqueda)
busqueda.content
busqueda.json()
busqueda.headers
busqueda.json()["statuses"]
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2019.</p>
| 14_introduccion_a_requests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The data block API
# + hide_input=true
from fastai.gen_doc.nbdoc import *
from fastai.vision import *
from fastai import *
# -
# The data block API lets you customize how to create a [`DataBunch`](/basic_data.html#DataBunch) by isolating the underlying parts of that process in separate blocks, mainly:
# - where are the inputs
# - how to label them
# - how to split the data into a training and validation set
# - what type of [`Dataset`](https://pytorch.org/docs/stable/data.html#torch.utils.data.Dataset) to create
# - possible transforms to apply
# - how to warp in dataloaders and create the [`DataBunch`](/basic_data.html#DataBunch)
#
# This is a bit longer than using the factory methods but is way more flexible. As usual, we'll begin with end-to-end examples, then switch to the details of each of those parts.
# ## Examples of use
# In [`vision.data`](/vision.data.html#vision.data), we create an easy [`DataBunch`](/basic_data.html#DataBunch) suitable for classification by simply typing:
path = untar_data(URLs.MNIST_SAMPLE)
tfms = get_transforms(do_flip=False)
data = ImageDataBunch.from_folder(path, ds_tfms=tfms, size=24)
# *ImageDataBunch* works with data in directories that follow the ImageNet style. In this style there is a *train* subdirectory and a *valid* subdirectory, each containing one subdirectory per class. The deepest subdirectories contain all the picture files.
#
# Here is the code for the code for the *data block* API to achieve the same result as the code in the cell above.
path = untar_data(URLs.MNIST_SAMPLE)
tfms = get_transforms(do_flip=False)
data = (ImageFileList.from_folder(path) #Where to find the data? -> in path and its subfolders
.label_from_folder() #How to label? -> depending on the folder of the filenames
.split_by_folder() #How to split in train/valid? -> use the folders
.datasets(ImageClassificationDataset) #How to convert to datasets? -> use ImageClassificationDataset
.transform(tfms, size=224) #Data augmetnation? -> use tfms with a size of 224
.databunch()) #Finally? -> use the defaults for conversion to ImageDataBunch
data.show_batch(rows=3, figsize=(5,5))
data.valid_ds.classes
# Let's look at another example from [`vision.data`](/vision.data.html#vision.data) with the planet dataset. This time, it's a multiclassification problem with the labels in a csv file and no given split between valid and train data, so we use a random split. The factory method is:
planet = untar_data(URLs.PLANET_TINY)
planet_tfms = get_transforms(flip_vert=True, max_lighting=0.1, max_zoom=1.05, max_warp=0.)
data = ImageDataBunch.from_csv(planet, folder='train', size=128, suffix='.jpg', sep = ' ', ds_tfms=planet_tfms)
# With the data block API we can rewrite this like that:
data = (ImageFileList.from_folder(planet)
#Where to find the data? -> in planet and its subfolders
.label_from_csv('labels.csv', sep=' ', folder='train', suffix='.jpg')
#How to label? -> use the csv file labels.csv in path,
#add .jpg to the names and take them in the folder train
.random_split_by_pct()
#How to split in train/valid? -> randomly with the defulat 20% in valid
.datasets(ImageMultiDataset)
#How to convert to datasets? -> use ImageMultiDataset
.transform(planet_tfms, size=128)
#Data augmetnation? -> use tfms with a size of 128
.databunch())
#Finally? -> use the defaults for conversion to databunch
data.show_batch(rows=3, figsize=(10,10), is_train=False)
# This new API also allows us to use dataset types for which there is no direct [`ImageDataBunch`](/vision.data.html#ImageDataBunch) factory method.
#
# For a segmentation task, for instance, we can use it to quickly get a [`DataBunch`](/basic_data.html#DataBunch). Let's take the example of the [camvid dataset](http://mi.eng.cam.ac.uk/research/projects/VideoRec/CamVid/). The images are in an 'images' folder and their corresponding masks are in a 'labels' folder.
camvid = untar_data(URLs.CAMVID_TINY)
path_lbl = camvid/'labels'
path_img = camvid/'images'
# We have a file that gives us the names of the classes (what each code inside the masks corresponds to: a pedestrian, a tree, a road, etc.).
codes = np.loadtxt(camvid/'codes.txt', dtype=str); codes
# And we define the following function that infers the mask filename from the image filename.
get_y_fn = lambda x: path_lbl/f'{x.stem}_P{x.suffix}'
# Then we can easily define a [`DataBunch`](/basic_data.html#DataBunch) using the data block API. Here we need to use `tfm_y=True` in the transform call because we need the same transforms to be applied to the target mask as were applied to the image.
data = (ImageFileList.from_folder(path_img) #Where are the input files? -> in path_img
.label_from_func(get_y_fn) #How to label? -> use get_y_fn
.random_split_by_pct() #How to split between train and valid? -> randomly
.datasets(SegmentationDataset, classes=codes) #How to create a dataset? -> use SegmentationDataset
.transform(get_transforms(), size=96, tfm_y=True) #Data aug -> Use standard tfms with tfm_y=True
.databunch(bs=64)) #Lastly convert in a databunch.
data.show_batch(rows=2, figsize=(5,5))
# One last example for object detection. We use our tiny sample of the [COCO dataset](http://cocodataset.org/#home) here. There is a helper function in the fastai library that reads the annotation file and returns the list of image names with the associated list of labelled bboxes.
#
# Next we convert the lists to a dictionary that maps image names with their bboxes and then write the function that will give us the target for each image filename.
coco = untar_data(URLs.COCO_TINY)
images, lbl_bbox = get_annotations(coco/'train.json')
img2bbox = {img:bb for img, bb in zip(images, lbl_bbox)}
get_y_func = lambda o:img2bbox[o.name]
# The following code is very similar to what we saw before. The only new addition is the use of a special function to collate the samples in batches. Our images may have multiple bounding boxes, so we need to pad them to the largest number of bounding boxes.
data = (ImageFileList.from_folder(coco)
#Where are the images? -> in coco
.label_from_func(get_y_func)
#How to find the labels? -> use get_y_func
.random_split_by_pct()
#How to split in train/valid? -> randomly with the default 20% in valid
.datasets(ObjectDetectDataset)
#How to create datasets? -> with ObjectDetectDataset
#Data augmentation? -> Standard transforms with tfm_y=True
.databunch(bs=16, collate_fn=bb_pad_collate))
#Finally we convert to a DataBunch and we use bb_pad_collate
data.show_batch(rows=3, is_train=False, figsize=(8,7))
# ## Provide inputs
# The inputs we want to feed our model are regrouped in the following class. The class contains methods to get the corresponding labels.
# + hide_input=true
show_doc(InputList, title_level=3, doc_string=False)
# -
# This class regroups the inputs for our model in `items` and saves a `path` attribute which is where it will look for any files (image files, csv file with labels...)
# + hide_input=true
show_doc(InputList.from_folder)
# -
# Note that [`InputList`](/data_block.html#InputList) is subclassed in vision by [`ImageFileList`](/vision.data.html#ImageFileList) that changes the default of `extensions` to image file extensions (which is why we used [`ImageFileList`](/vision.data.html#ImageFileList) in our previous examples).
# ## Labelling the inputs
# All the followings are methods of [`InputList`](/data_block.html#InputList). Note that some of them are primarly intended for inputs that are filenames and might not work in general situations.
# + hide_input=true
show_doc(InputList.label_from_csv)
# + hide_input=true
jekyll_note("This method will only keep the filenames that are both present in the csv file and in `self.items`.")
# + hide_input=true
show_doc(InputList.label_from_df)
# + hide_input=true
jekyll_note("This method will only keep the filenames that are both present in the dataframe and in `self.items`.")
# + hide_input=true
show_doc(InputList.label_from_folder)
# + hide_input=true
jekyll_note("This method looks at the last subfolder in the path to determine the classes.")
# + hide_input=true
show_doc(InputList.label_from_func)
# + hide_input=true
show_doc(InputList.label_from_re)
# + hide_input=true
show_doc(LabelList, title_level=3, doc_string=False)
# -
# A list of labelled inputs in `items` (expected to be tuples of input, label) with a `path` attribute. This class contains methods to create `SplitDataset`. In future development, it will contain factory methods to directly create a [`LabelList`](/data_block.html#LabelList) from a source of labelled data (a csv file or a dataframe with inputs and labels) for instance.
# ## Split the data between train and validation.
# The following functions are methods of [`LabelList`](/data_block.html#LabelList), to create a [`SplitData`](/data_block.html#SplitData) in different ways.
# + hide_input=true
show_doc(LabelList.random_split_by_pct)
# + hide_input=true
show_doc(LabelList.split_by_files)
# + hide_input=true
show_doc(LabelList.split_by_fname_file)
# + hide_input=true
show_doc(LabelList.split_by_folder)
# + hide_input=true
jekyll_note("This method looks at the folder immediately after `self.path` for `valid` and `train`.")
# + hide_input=true
show_doc(LabelList.split_by_idx)
# + hide_input=true
show_doc(SplitData, title_level=3)
# -
# ## Create datasets
# To create the datasets from [`SplitData`](/data_block.html#SplitData) we have the following class method.
# + hide_input=true
show_doc(SplitData.datasets)
# + hide_input=true
show_doc(SplitDatasets, title_level=3)
# -
# This class can be constructed directly from one of the following factory methods.
# + hide_input=true
show_doc(SplitDatasets.from_single)
# + hide_input=true
show_doc(SplitDatasets.single_from_c)
# + hide_input=true
show_doc(SplitDatasets.single_from_classes)
# -
# Then we can build the [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) around our [`Dataset`](https://pytorch.org/docs/stable/data.html#torch.utils.data.Dataset) like this.
# + hide_input=true
show_doc(SplitDatasets.dataloaders)
# -
# The methods `img_transform` and `img_databunch` used earlier are documented in [`vision.data`](/vision.data.html#vision.data).
# ## Utility classes
# + hide_input=true
show_doc(ItemList, title_level=3)
# + hide_input=true
show_doc(PathItemList, title_level=3)
| docs_src/data_block.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pkgutil as pu
import numpy as np
import matplotlib as mpl
import scipy as sp
import pydoc
print ("NumPy version", np.__version__)
print ("SciPy version", sp.__version__)
print ("Matplotlib version", mpl.__version__)
def clean(astr):
s = astr
# remove multiple spaces
s = ' '.join(s.split())
s = s.replace('=','')
return s
def print_desc(prefix, pkg_path):
for pkg in pu.iter_modules(path=pkg_path):
name = prefix + "." + pkg[1]
if pkg[2] == True:
try:
docstr = pydoc.plain(pydoc.render_doc(name))
docstr = clean(docstr)
start = docstr.find("DESCRIPTION")
docstr = docstr[start: start + 140]
print (name, docstr)
except:
continue
print_desc("numpy", np.__path__)
print (" ")
print (" ")
print (" ")
print_desc("scipy", sp.__path__)
# -
| PythonDAdata/3358OS_03_Code/3358OS_03_Code/pkg_check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # My first `scikit-learn` notebook
import pandas as pd
import numpy as np
from random import choices
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
# Load a dataset
forecast = pd.read_csv('data/Forecast.csv')
forecast.head()
# Setup the `numpy` arrays to use to train classifiers
y = forecast.pop('Go-Out').values # target feature
X = forecast.values # training data
type(X),type(y)
# Train a *k*-NN classifier
kNN = KNeighborsClassifier(n_neighbors=3)
kNN.fit(X,y)
X_test = np.array([[8,70,11],
[8,69,15]])
kNN.predict(X_test)
# All `sklearn` classifiers implement the `Estimator` API.
tree = DecisionTreeClassifier()
tree.fit(X,y)
tree.predict(X_test)
lr = LogisticRegression()
lr.fit(X,y)
lr.predict(X_test)
# Swapping between classifiers (Estimators) makes model selection easy.
cfrs = [kNN,tree,lr]
for cfr in cfrs:
cfr.fit(X,y)
print(cfr.predict(X_test))
# ## Preprocessing
# All preprocessing modules implement the `Transformer` API.
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(X) # standardise to zero mean and unit variance
X_scaled = scaler.transform(X)
X_test_scaled = scaler.transform(X_test)
X_test_scaled
mm_scaler = preprocessing.MinMaxScaler() # standardise to range [0,1]
mm_scaler.fit(X)
X_scaled = mm_scaler.transform(X)
X_test_scaled = mm_scaler.transform(X_test)
X_test_scaled
| Complete Modules/ML labs/.ipynb_checkpoints/01 sklearn Intro-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Simple Calculator
print('This is ESRG.')
import numpy as np
# +
# Program make a simple calculator
# This function adds two numbers
def add(x, y):
return x + y
# This function subtracts two numbers
def subtract(x, y):
return x - y
# This function multiplies two numbers
def multiply(x, y):
return x * y
# This function divides two numbers
def divide(x, y):
return x / y
print("Select operation.")
print("1.Add")
print("2.Subtract")
print("3.Multiply")
print("4.Divide")
while True:
# take input from the user
choice = input("Enter choice(1/2/3/4): ")
# check if choice is one of the four options
if choice in ('1', '2', '3', '4'):
num1 = float(input("Enter first number: "))
num2 = float(input("Enter second number: "))
if choice == '1':
print(num1, "+", num2, "=", add(num1, num2))
elif choice == '2':
print(num1, "-", num2, "=", subtract(num1, num2))
elif choice == '3':
print(num1, "*", num2, "=", multiply(num1, num2))
elif choice == '4':
print(num1, "/", num2, "=", divide(num1, num2))
# check if user wants another calculation
# break the while loop if answer is no
next_calculation = input("Let's do next calculation? (yes/no): ")
if next_calculation == "no":
break
else:
print("Invalid Input")
# -
| Simple Calculator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('.')
import model
# -
import matplotlib.pyplot as plt
import networkx as nx
import numpy as np
import pandas as pd
import random
import seaborn as sns
import statistics
# +
N = 100
K = 4
p_star = 0.256
g, _ = model.watts_strogatz_case_p_star(N, K, p_star)
green_cmap = plt.get_cmap('Greens')
orange_cmap = plt.get_cmap('Oranges')
plt.figure(3,figsize=(7,7))
nx.draw_circular(g,
node_size = 15,
node_color = ['c' if latitude(x) > 0 else 'b' for x in g.nodes()],
edge_color = ['c'
if latitude(e[0]) > 0 and latitude(e[1]) > 0
else 'k' for e in g.edges()]
)
# -
# We suspect that contact tracing is more effective when its adoption is _dense_.
#
# Let's consider a Watts-Strogatzz graph as before.
N = 2000
K = 4
p_star = 0.256
def ws_case_generator(N, K, p_star):
def wscg(**kwargs):
return model.watts_strogatz_case_p_star(N, K, p_star, **kwargs)
return wscg
g, _ = model.watts_strogatz_case_p_star(N, K, p_star)
# But this time we will segment the population into two regional halves, a 'north' and a 'south'.
def latitude(i):
dist_from_north_pole = min(
i,
abs(N / 4 - i) # 1/4 here is just rotating, for the drawing
)
return N / 4 - dist_from_north_pole
# +
green_cmap = plt.get_cmap('Greens')
orange_cmap = plt.get_cmap('Oranges')
plt.figure(3,figsize=(7,7))
nx.draw_circular(g,
node_size = 15,
node_color = ['c' if latitude(x) > 0 else 'b' for x in g.nodes()],
edge_color = ['c'
if latitude(e[0]) > 0 and latitude(e[1]) > 0
else 'k' for e in g.edges()]
)
# -
# We can compute the expected value of inter-group edges, and compare them with the empirical values.
#
# This is the computed number of north-north edges.
len(g.edges())
# +
both_north = [
e
for e in g.edges()
if latitude(e[0]) > 0 and latitude(e[1]) > 0
]
len(both_north)
# -
# We would expect that as function of $N$, $k$, and $p$, this value to be:
# - The chance of the first node being in the northern hemisphere, $1/2$, times
# - The chance that the linked node is in the northern hemisphere, which is, approximately:
# - $(1 - p)$, for the chance that the edge is "close" and therefore another "north" node
# - this will be offer by a small amount, proportional to $K / N$, due to "border nodes"
# - $p / 2$, for the chance that the edge is "distant" and the (randomly chosen) distant edge is also in the "north"
#
# Or, in total, $(NK/2) * 1/2 * ((1 - p) + p / 2)$ north-north edges.
(N * K / 2) * .5 * ((1 - p_star) + p_star / 2)
# Which we see is indeed very close to an "empirical" value based on randomized distant links.
# An elaboration of this would allow for regional partitions of the node space beyond half-half.
# -------------------------------
# Tying this back to contact tracing, we can consider what happens when the geographic regions (which might represent of forms of social categorization or partitioning) have different adoption rates.
#
# This difference in adoption could be due to group or activity based privacy concerns, such as those having to do with being an ethnic minority or attending political demonstrations.
# In the most extreme cases, the adoption rate of the first group is $A_0 = 1$ and the adoption rate of the second group is $A_1 = 0$.
#
# In this case, the number of "traced edges" is equal to the number of "north-north" edges.
# This is a function of $p$ but, notably, we are treating traced edges that are "close" and "distant" as the same.
#
# In general, if $b$ is the proportion of nodes in the first group, then the number of traced edges will be equal to:
#
# - The chance of being a north-north edge, $b * (1 - (1 - b)p)$ times the north-north tracing rate $A_0^2$
# - The chance of being a south-south edge, $(1 - b) * (1 - b p)$ times the south-south tracing rate $A_1^2$
# - The chance of being a north-south or south-north edge $b * p * (1 - b) + (1 - b) * p * b = 2pb(1-b)$ times the north-south tracing rate $A_0 A_1$
#
# If $b = 0.5$, then these come to
# $$(.5 - .25p) * A_0^2 + (.5 - .25p) * A_1^2 + 2pb(1-b) * A_0 A_1$$
# $$(.5 - .25p) * A_0^2 + (.5 - .25p) * A_1^2 + .5p * A_0 A_1$$
#
# Let $A_0 = \mu - \delta$ and $A_1 = \mu + \delta$
# Then total number of traced edges is:
#
# $$(.5 - .25p) (\mu - \delta)^2 + (.5 - .25p) (\mu + \delta)^2 + .5p (\mu - \delta) (\mu + \delta)$$
#
# $$(.5 - .25p) * (\mu^2 - 2 \mu \delta + \delta^2) + (.5 - .25p) * (\mu^2 + 2 \mu \delta + \delta^2 ) + .5p * (\mu^2 - \delta^2)$$
#
# $$(1 - .5p) * (\mu^2 + \delta^2 ) + .5p * (\mu^2 - \delta^2)$$
#
# $$\mu^2 + \delta^2 - .5p (\mu^2 + \delta^2) + .5p (\mu^2 - \delta^2)$$
#
# $$\mu^2 + \delta^2 - p \delta^2 = T$$
# ----------------------------------------------------------------
# One question we might ask is whether, for a given number of traced edges, whether it matters how much they are grouped together.
#
# We can operationalize this like so:
# - Given the _subgraph_ $G_T$ of $G$ which includes nodes with traced edges
# - and the _local clustering coefficient_ of each node in this subgraph (how close its neighbors are to being a clique, i.e. fully connected)
# - what is the _average local clustering coefficient_ of $G_T$. (This is the version of 'clustering coefficient' developed by Watts and Strogatz in the construction of the Watts-Strogatz model.)
# We can run the numbers with the simple case that $A_0 = 1$ and $A_1 = 1$ below.
# +
traced_nodes = set([x[0] for x in both_north]).union(set([x[1] for x in both_north]))
gt = g.subgraph(traced_nodes)
nx.draw_circular(gt,
node_size = 15,
node_color = ['c' if latitude(x) > 0 else 'b' for x in gt.nodes()],
edge_color = ['c'
if latitude(e[0]) > 0 and latitude(e[1]) > 0
else 'k' for e in gt.edges()]
)
# -
nx.average_clustering(gt)
# ---------------------------------------------------------------
#
N = 2000
K = 4
p_star = 0.256
## Population parameters:
base_params = {
# Node parameter
'A' : 1, # This will be modified in each case
# Edge parameter
'W' : .5, # probability of edge activation; 2/K
'C' : 1.0, ## all edges can be traced.
## Disease parameters
'beta_hat' : .4, # probability of transmission upon contact
'alpha' : .25, # probability of exposed becoming infectious
'gamma' : .1, # probability of infectious becoming recovered
'zeta' : .1, # probability of infectious becoming symptomatic
## Contact tracing parameters
'limit' : 10, # number of time steps the contact tracing system remembers
}
conditions = {
'mu-0.4_d-0.0' : {'A' : model.hemisphere_adoption(0.4, 0.0), 'mu' : 0.4, 'delta' : 0.0},
'mu-0.4_d-0.1' : {'A' : model.hemisphere_adoption(0.4, 0.1), 'mu' : 0.4, 'delta' : 0.1},
'mu-0.4_d-0.2' : {'A' : model.hemisphere_adoption(0.4, 0.0), 'mu' : 0.4, 'delta' : 0.2},
'mu-0.4_d-0.3' : {'A' : model.hemisphere_adoption(0.4, 0.0), 'mu' : 0.4, 'delta' : 0.3},
'mu-0.4_d-0.4' : {'A' : model.hemisphere_adoption(0.4, 0.0), 'mu' : 0.4, 'delta' : 0.4},
'mu-0.5_d-0.0' : {'A' : model.hemisphere_adoption(0.5, 0.0), 'mu' : 0.5, 'delta' : 0.0},
'mu-0.5_d-0.05' : {'A' : model.hemisphere_adoption(0.5, 0.1), 'mu' : 0.5, 'delta' : 0.05},
'mu-0.5_d-0.1' : {'A' : model.hemisphere_adoption(0.5, 0.1), 'mu' : 0.5, 'delta' : 0.1},
'mu-0.5_d-0.15' : {'A' : model.hemisphere_adoption(0.5, 0.1), 'mu' : 0.5, 'delta' : 0.15},
'mu-0.5_d-0.2' : {'A' : model.hemisphere_adoption(0.5, 0.2), 'mu' : 0.5, 'delta' : 0.2},
'mu-0.5_d-0.25' : {'A' : model.hemisphere_adoption(0.5, 0.1), 'mu' : 0.5, 'delta' : 0.25},
'mu-0.5_d-0.3' : {'A' : model.hemisphere_adoption(0.5, 0.3), 'mu' : 0.5, 'delta' : 0.3},
'mu-0.5_d-0.35' : {'A' : model.hemisphere_adoption(0.5, 0.1), 'mu' : 0.5, 'delta' : 0.35},
'mu-0.5_d-0.4' : {'A' : model.hemisphere_adoption(0.5, 0.4), 'mu' : 0.5, 'delta' : 0.4},
'mu-0.5_d-0.45' : {'A' : model.hemisphere_adoption(0.5, 0.1), 'mu' : 0.5, 'delta' : 0.45},
'mu-0.5_d-0.5' : {'A' : model.hemisphere_adoption(0.5, 0.5), 'mu' : 0.5, 'delta' : 0.5},
'mu-0.7_d-0.0' : {'A' : model.hemisphere_adoption(0.7, 0.0), 'mu' : 0.7, 'delta' : 0.0},
'mu-0.7_d-0.15' : {'A' : model.hemisphere_adoption(0.7, 0.15), 'mu' : 0.7, 'delta' : 0.15},
'mu-0.7_d-0.3' : {'A' : model.hemisphere_adoption(0.7, 0.3), 'mu' : 0.7, 'delta' : 0.3}
}
# +
def dfr(rs):
return pd.DataFrame(
[r for case in rs
for r in model.data_from_results(rs, case)])
runs = 200
base_params['A'] = 1
rs = model.experiment(
ws_case_generator(N, K, p_star),
base_params,
conditions,
runs)
temp = dfr(rs)
temp.to_csv('hemisphere_study.csv')
#del rs
# -
# -----------------------------------
#
# #### Analysis
#
# This is the analysis section
data = pd.read_csv('hemisphere_study.csv')
# +
data['traced_edges_close'] = data['traced_edges'] - data['traced_edges_distant']
data['traced_edges_ratio'] = data['traced_edges'] / (data['N'] * data['K'] / 2)
data['traced_edges_distant_ratio'] = data['traced_edges_distant'] / data['traced_edges']
data['T'] = data['mu'] ** 2 + data['delta'] ** 2 - data['p'] * data['delta'] ** 2
data['d/mu'] = data['delta'] / data['mu']
# +
data["delta-cat"] = data["delta"].apply(lambda x: f"delta = {x}")
splot = sns.lineplot(x='T', y='infected_ratio', hue="mu", data=data)
# +
data["delta-cat"] = data["delta"].apply(lambda x: f"delta = {x}")
splot = sns.lineplot(x='delta', y='infected_ratio', hue="mu", data=data)
# -
g = sns.scatterplot(
data = data,
y = 'infected_ratio',
x = 'traced_edges',
hue = "mu"
)
g, xyz, db = model.binned_heatmap(
data,
x = 'T',
x_base = 0.1,
y = 'd/mu',
y_base = .1,
z = 'infected_ratio'
)
# +
extreme = data[data["mu"] == 0.5]
extreme = data[data["delta"] == 0.5]
# -
plt.hist(extreme['infected_ratio'],bins=30)
g = sns.scatterplot(
data = extreme,
y = 'infected_ratio',
x = 'traced_edges_distant_ratio',
)
| contact-tracing/code/Python/Experiment--A12 on p* WS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="ku_logo_uk_v.png" alt="drawing" width="130" style="float:right"/>
#
# # <span style="color:#2c061f"> Exercise 4 </span>
#
# <br>
#
# ## <span style="color:#374045"> Introduction to Programming and Numerical Analysis </span>
#
#
# #### <span style="color:#d89216"> <br> <NAME> </span>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Plan for today
# <br>
#
# 1. Welcome
# 2. Plotting
# 3. Optimization
# 4. Problemset 1
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 2. Plotting
#
# There exist numerous different modules for plotting in Python. In this course we will work mainly with two of them. The first is the classic tool, `matplotlib`. The second tool is `seaborn` which is a newer module that allows to produce figures with less effort.
#
# Lets try it out:
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 2.1 import libraries
# + slideshow={"slide_type": "fragment"}
import matplotlib.pyplot as plt
from matplotlib import cm # for colormaps
import numpy as np
import seaborn as sns
#Center images in notebook (optional)
from IPython.core.display import HTML
HTML("""
<style>
.output_png {
display: table-cell;
text-align: center;
vertical-align: middle;
}
</style>
""")
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 2.1. Define function to plot
# + slideshow={"slide_type": "fragment"}
def f(x):
"""
Defines an equation.
Args:
x (list): list of variable arrays
Returns:
eq_1 (scalar): Function value
"""
if len(x)==0:
raise ValueError ("No variables defined")
elif len(x)==1: # to ensure input is valid - in this case only 1 allowed
eq_1 = np.sin(x[0])+0.05*x[0]**2
return eq_1
elif len(x)==2: # to ensure input is valid - in this case only 2 variables allowed
eq_1 = np.sin(x[0])+0.05*x[1]**2
return eq_1
elif len(x)>2:
raise ValueError ("Too many variables defined")
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 2.2 Generate data
# + slideshow={"slide_type": "fragment"}
N=100
x1_vec = np.linspace(-10,10,N) # draw N=100 evenly spaced numbers between -10 and 10
x2_vec = np.linspace(-10,10,N)
x1_grid,x2_grid = np.meshgrid(x1_vec,x2_vec,indexing='ij') # create coordinate matrix
x = ([x1_grid,x2_grid]) #because function takes a list of arrays
f_grid = f(x)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 2.3 Plot function values - 3-dimensional
# + slideshow={"slide_type": "fragment"}
fig = plt.figure(figsize=(9,9)) # define figure object and size in general 13/9
ax = fig.add_subplot(111, projection='3d') # define subplot and projection
ax.plot_surface(x1_grid,x2_grid,f_grid,cmap=cm.viridis) # plot 3d surface and colormap
ax.invert_xaxis()
ax.set_xlabel('$x_1$',fontsize=12) # set labels and fontsize
ax.set_ylabel('$x_2$',fontsize=12)
ax.set_zlabel('$f(x_1,x_2)$',fontsize=12)
#ax.set_title("Cool 3d-graph",fontsize=14)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 2.4 2-dimensional function
# + slideshow={"slide_type": "fragment"}
fig2 = plt.figure() # define new figure object
ax = fig2.add_subplot(111) # add subplot
ax.plot(x1_vec,f([x1_vec])) # plot function value for first instance in y_grid
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 2.4 2-dimensional function - grammar of graphics
# Now lets turn to how to construct a nice figure. I will follow the [_grammar of graphics_](http://vita.had.co.nz/papers/layered-grammar.pdf) framework. Things to keep in mind:
#
# 1. Keep it simple!
# - Anything unnecessary should be removed, see [this post](https://www.darkhorseanalytics.com/blog/data-looks-better-naked/).
#
# 2. Self explanatory
# - Contain axis label, title, footnotes in text containing relevant information.
# 3. Nice appereance
# - Choose the right plot type.
# - Make sure font type and size match. Beware of colors and line width.
# + slideshow={"slide_type": "subslide"}
fig3 = plt.figure(figsize=(10,5)) # define new figure object
ax = fig3.add_subplot(111) # add subplot
ax.plot(x1_vec,f([x1_vec])) # plot 2-dimensional function
#add lines
for y in range(-1, 6):
plt.plot(range(-10,11), [y] * len(range(-10, 11)), "--", lw=0.5, color="black", alpha=0.3)
ax.xaxis.label.set_fontsize(14) #set label fontsize to 14
ax.yaxis.label.set_fontsize(14)
ax.set(xlabel="$x_1$", ylabel = "$f(x_1)$",xlim = ([-10,10])) #set xlabel,ylabel and xlimit
for item in ax.get_yticklabels()+ax.get_xticklabels(): # set ticklabels to fontsize 14
item.set_fontsize(14)
#remove borders
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.spines['bottom'].set_visible(False)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 3.1 Optimization
#
# Let's try to find the global optimum of the 3d function. How can we do this?
# - We can loop through solutions and return optimum
# - Cumbersome and not guaranteed to yield optimum
# - Or we can use Scipy's optimization module
# - We use scipy.optimize.minimize_scalar when we only optimize one argument
# - For multidimensional functions we use scipy.optimize.minimize
#
# - **Note:** We use minimize even though we solve a maximization problem - just remember to subtract objective function!
# + slideshow={"slide_type": "subslide"}
# a. using scipy optmize
from scipy import optimize
x_guess = (0,0) # optimizer needs a starting point for the two values
obj = lambda x:f(x) #objective function to optimize - in this case minimize
#b.optimizing objective function
res = optimize.minimize(obj,x_guess,method="Nelder-Mead") #Nelder-mead is standard and simple method
print("-----------")
print(res.message)
print("-----------")
#c.unpacking results
x1_best_scipy = res.x[0]
x2_best_scipy = res.x[1]
f_best_scipy = res.fun
# d. print
print(f'Using numerical solver the optimal values are:')
print(f'Function = {f_best_scipy.item():.4f}; x1 = {x1_best_scipy:.4f}; x2 = {x2_best_scipy:.4f}')
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="nelder-mead.gif" style="float:right">
#
# # <span style="color:#2c061f"> 3.2 Whats going on under the hood? </span>
#
#
# <br>
# <span style="color:#374045"> - Largely depends on the method used! </span>
#
# <span style="color:#374045"> - Nelder-Mead is a downhill method, whereas other algorithm use a lagrangian method to incoorporate constraints - e.g. consumer budget constraint </span>
#
# <span style="color:#374045"> - Gif shows the Nelder-Mead method </span>
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="2d.gif" style="float:right">
#
# ## <span style="color:#2c061f"> 3.3 Optimizing 2-dimensional function </span>
# <br>
# <span style="color:#2c061f"> Nelder-Mead in action. Red dot starts at the initial guess $x_1=0$ and quickly converges to the global minimum. </span>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## <span style="color:#2c061f"> 3.3 bounded optimization </span>
# <br>
# <span style="color:#2c061f"> What if we know that the optimum lies within a given interval? For instance optimal consumption is bounded by individuals income. Lets turn to the 2d function for this. </span>
#
# <br>
# <br>
# <span style="color:#2c061f"> - Lets try to find the optimum in the interval [-10,-2.5] </span>
# + slideshow={"slide_type": "subslide"}
#a. bounded optimizer
x_guess = 0 # optimizer needs a starting point
bounds = (-10,-2.5) # interval optimum lies
obj = lambda x1_vec: f([x1_vec]) #objective function
#b.optimizing objective function
res = optimize.minimize_scalar(obj,x_guess,method="bounded",bounds=bounds) #we use minimize_scalar here!
print("-----------")
print(res.message)
print("-----------")
#c.unpacking results
x1_best_scipy = res.x
f_best_scipy = res.fun
# d. print
print(f'Using numerical solver the optimal values are:')
print(f'Function = {f_best_scipy.item():.4f}; x1 = {x1_best_scipy:.4f}')
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 3.4 Optimization tips and tricks
#
# - what if we have a function that takes more than one argument and we only want to optimize one argument?
# - Specify `args` in optimizer and type in those arguments that should be held constant e.g. $\alpha, \beta,$ income, prices etc.
#
# - Usually constraints can be expressed such that you can avoid using multi-dimensional constrainted solvers
# - Consumer budget constraint with two goods can be rewritten such that $x_2$ is an implicit function of $x_1$. See lecture notebook section 7.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## 4. Problemset 1
#
# Thats it for me today. Your turn to work with optimization and plotting!
| class 4/class 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir(os.path.join('..', 'notebook_format'))
from formats import load_style
load_style(plot_style = False)
# +
os.chdir(path)
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
# 4. a ipython magic to enable retina (high resolution) plots
# https://gist.github.com/minrk/3301035
# %matplotlib inline
# %load_ext watermark
# %load_ext autoreload
# %autoreload 2
# %config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import seaborn as sns
import scipy.stats as stats
import matplotlib.pyplot as plt
from statsmodels.stats.proportion import proportions_ztest
from statsmodels.stats.proportion import proportions_chisquare
# %watermark -a 'Ethen' -d -t -v -p numpy,scipy,pandas,matplotlib,statsmodels
# +
# setup the look and feel of the notebook
plt.rcParams['figure.figsize'] = 8, 6
sns.set_context('notebook', font_scale = 1.5, rc = {'lines.linewidth': 2.5})
sns.set_style('whitegrid')
sns.set_palette('deep')
# Create a couple of colors to use throughout the notebook
red = sns.xkcd_rgb['vermillion']
blue = sns.xkcd_rgb['dark sky blue']
# -
# Ideally, the reader should already understand or vaguely remember the statistic concepts such as z-score, p-value, hypothesis test, confidence interval. The warming-up section is a quick review of the concept, feel free to skip it if you're already acquainted with the concept.
#
# # Warming-up
#
# Statistical inference is the process of analyzing sample data to gain insight into the population from which the data was collected and to investigate differences between data samples. In data analysis, we are often interested in the characteristics of some large population, but collecting data on the entire population may be infeasible. For example, leading up to U.S. presidential elections it could be very useful to know the political leanings of every single eligible voter, but surveying every voter is not feasible. Instead, we could poll some subset of the population, such as a thousand registered voters, and use that data to make inferences about the population as a whole.
#
# ## Point Estimate
#
# Point estimates are estimates of population parameters based on sample data. For instance, if we wanted to know the average age of registered voters in the U.S., we could take a survey of registered voters and then use the average age of the respondents as a point estimate of the average age of the population as a whole. The average of a sample is known as the sample mean.
# The sample mean is usually not exactly the same as the population mean. This difference can be caused by many factors including poor survey design, biased sampling methods and the randomness inherent to drawing a sample from a population. Let's investigate point estimates by generating a population of random age data and then drawing a sample from it to estimate the mean:
# generate some random number to serve as our population
np.random.seed(10)
population_ages1 = stats.poisson.rvs(loc = 18, mu = 35, size = 150000)
population_ages2 = stats.poisson.rvs(loc = 18, mu = 10, size = 100000)
population_ages = np.concatenate((population_ages1, population_ages2))
print('population mean:', np.mean(population_ages))
np.random.seed(6)
sample_ages = np.random.choice(population_ages, size = 500)
print('sample mean:', np.mean(sample_ages))
# The experiment tells us that we'd expect the distribution of the population to be a similar shape to that of the sample, so we can assume that the mean of the sample and the population should have the same value. Note that we can't say that they exactly match, but it's the best estimate we can make.
#
# The population mean is often denoted as $\mu$, the estimated population mean as $\hat{\mu}$, mean of the sample $\bar{x}$. So here we're basically saying $\hat{\mu} = \bar{x}$, where we're using the sample mean to estimate the mean of the population and usually the larger the size of our sample, the more accurate our point estimator for the estimated population mean is going to be.
#
#
# ## Sampling Distributions and The Central Limit Theorem
#
# Many statistical procedures assume that data follows a normal distribution, because the normal distribution has nice properties like symmetricity and having the majority of the data clustered within a few standard deviations of the mean. Unfortunately, real world data is often not normally distributed and the distribution of a sample tends to mirror the distribution of the population. This means a sample taken from a population with a skewed distribution will also tend to be skewed.
fig = plt.figure(figsize = (12, 6))
plt.subplot(1, 2, 1)
plt.hist(population_ages)
plt.title('Population')
plt.subplot(1, 2, 2)
plt.hist(sample_ages)
plt.title('Sample')
plt.show()
# The plot reveals the data is clearly not normal: instead of one symmetric bell curve, it has as bimodal distribution with two high density peaks. Because of this, the sample we drew from this population should have roughly the same shape and skew.
#
# The sample has roughly the same shape as the underlying population. This suggests that we can't apply techniques that assume a normal distribution to this data set, since it is not normal. This leads to our next topic, the **central limit theorem**.
#
# The central limit theorem is one of the most important results of probability theory and serves as the foundation of many methods of statistical analysis. At a high level, the theorem states the distribution of many sample means, known as a sampling distribution, will be normally distributed. This rule holds even if the underlying distribution itself is not normally distributed. As a result we can treat the sample mean as if it were drawn normal distribution. To illustrate, let's create a sampling distribution by taking 200 samples from our population and then making 200 point estimates of the mean:
# +
np.random.seed(10)
samples = 200
point_estimates = [np.random.choice(population_ages, size = 500).mean()
for _ in range(samples)]
plt.hist(point_estimates)
plt.show()
# -
# The sampling distribution appears to be roughly normal, despite the bimodal population distribution that the samples were drawn from. In addition, the mean of the sampling distribution approaches the true population mean:
population_ages.mean() - np.mean(point_estimates)
# To hit the notion home, Central Limit Theorem states that that if we collect "a large number" of different samples mean from the population, the sampling distribution, the distribution of the samples mean you collected, will approximately take the shape of a normal distribution around the population mean no matter what the orginal population distribution is.
#
# Knowing that the sampling distribution will take the shape of a normal distribution is what makes the theorem so powerful, as it is the foundation of concepts such as confidence intervals and margins of error in frequentist statistics.
# ## Confidence Interval
#
# A point estimate can give us a rough idea of a population parameter like the mean, but estimates are prone to error. A confidence interval is a range of values above and below a point estimate that captures the true population parameter at some predetermined confidence level. For example, if you want to have a 95% chance of capturing the true population parameter with a point estimate and a corresponding confidence interval, we'd set our confidence level to 95%. Higher confidence levels result in a wider confidence intervals.
#
# The interval is computed using the formula:
#
# $$\text{point estimate} \pm z * SE$$
#
# Where
#
# - $z$ is called the **critical value** and it corresponds to the **confidence level** that we chose. Critical value is the number of standard deviations we'd have to go from the mean of the normal distribution to capture the proportion of the data associated with the desired confidence level. For instance, we know that roughly 95% of the data in a normal distribution lies within 2 standard deviations from the mean, so we could use 2 as the z-critical value for a 95% confidence interval (although it is more exact to get z-critical values with `stats.norm.ppf()`)
# - $SE$ represents the **standard error**. Generally the standard error for a point estimate is estimated from the data and computed using a formula. For example, the standard error for the sample mean is $\frac{s}{ \sqrt{n} }$, where $s$ is the standard deviation and $n$ is the number of samples.
# - The value $z * SE$ is called the **margin of error**.
# - Note that this constructing confidence intervals framework holds true for doing can be easily adapted for any estimator that has a nearly normal sampling distribution. e.g. sample mean, two sample mean, sample proportion and two sample proportion (we'll later see). All we have to do this is change the way that we're calculating the standard error.
# +
np.random.seed(10)
sample_size = 1000
sample = np.random.choice(population_ages, size = sample_size)
sample_mean = sample.mean()
confidence = 0.95
z_critical = stats.norm.ppf(q = confidence + (1 - confidence) / 2)
print('z-critical value:', z_critical)
pop_stdev = population_ages.std()
margin_of_error = z_critical * (pop_stdev / np.sqrt(sample_size))
confint = sample_mean - margin_of_error, sample_mean + margin_of_error
print('point esimate:', sample_mean)
print('Confidence interval:', confint)
# -
# Notice that the confidence interval we calculated captures the true population mean of 43.0023.
# Let's create several confidence intervals and plot them to get a better sense of what it means to "capture" the true mean:
# +
np.random.seed(12)
confidence = 0.95
sample_size = 1000
intervals = []
sample_means = []
for sample in range(25):
sample = np.random.choice(population_ages, size = sample_size)
sample_mean = sample.mean()
sample_means.append(sample_mean)
z_critical = stats.norm.ppf(q = confidence + (1 - confidence) / 2)
pop_stdev = population_ages.std()
margin_of_error = z_critical * (pop_stdev / np.sqrt(sample_size))
confint = sample_mean - margin_of_error, sample_mean + margin_of_error
intervals.append(confint)
plt.figure(figsize = (10, 8))
plt.errorbar(x = np.arange(0.1, 25, 1), y = sample_means,
yerr = [(top - bot) / 2 for top, bot in intervals], fmt = 'o')
plt.hlines(xmin = 0, xmax = 25,
y = population_ages.mean(),
linewidth = 2.0, color = red)
plt.show()
# -
# Notice that in the plot above, all but one of the 95% confidence intervals overlap the red line marking the true mean. This is to be expected: since a 95% confidence interval captures the true mean 95% of the time, we'd expect our interval to miss the true mean 5% of the time.
#
# More formally, the definition of a 95% confidence interval means that **95% of confidence intervals, created based on random samples of the same size from the same population will contain the true population parameter**.
# ## Hypothesis Testing
#
# Lets starts off with a motivating example that asks the question "If you toss a coin 30 times and see 22 heads, is it a fair coin?"
#
# We all know that a fair coin should come up heads roughly 15 out of 30 tosses, give or take, so it does seem unlikely to see so many heads. However, the skeptic might argue that even a fair coin could show 22 heads in 30 tosses from time-to-time. This could just be a chance event. So, the question would then be "how can you determine if we're tossing a fair coin?"
#
# Let's start by first considering the probability of a single coin flip coming up heads and work our way up to 22 out of 30.
#
# \begin{align}
# P(H) = \frac{1}{2}
# \end{align}
#
# As our equation shows, the probability of a single coin toss turning up heads is exactly 50% since there is an equal chance of either heads or tails turning up. Taking this one step further, to determine the probability of getting 2 heads in a row with 2 coin tosses, we would need to multiply the probability of getting heads by the probability of getting heads again since the two events are independent of one another.
#
# \begin{align}
# P(HH) = P(H) \cdot P(H) = P(H)^2 = \left(\frac{1}{2}\right)^2 = \frac{1}{4}
# \end{align}
#
# Let's now take a look at a slightly different scenario and calculate the probability of getting 2 heads and 1 tails with 3 coin tosses. To get the actual probability of tossing 2 heads and 1 tails we will have to add the probabilities for all of the possible permutations, of which there are exactly three: HHT, HTH, and THH.
#
# \begin{align}
# P(2H,1T) = P(HHT) + P(HTH) + P(THH) = \frac{1}{8} + \frac{1}{8} + \frac{1}{8} = \frac{3}{8}
# \end{align}
#
# Another way we could do this is to use the binomial distribution:
#
# \begin{align}
# P(N_H,N_T) = \binom{n}{k} p^{k} \left( 1 - p \right)^{n - k}
# \end{align}
#
# Where
#
# - $n$ is number of coin flips
# - $p$ is the probability of getting heads on each flip
#
# The $\binom{n}{k}$ tells us how many ways are there to get $k$ heads our of $n$ total number of coin flips?" and the $p^k(1-p)^{n-k}$ answers the question "how likely is any given $k$ heads and $n-k$ tails?", multiply them together and we get the probability of getting exactly $k$ heads.
#
# Now that we understand the classic method, let's use it to test whether we are actually tossing a fair coin.
# +
# Calculate the probability for every possible outcome
# of tossing a fair coin 30 k_range
k_range = range(1, 31) # number of heads appearing
n = 30 # number of k_range tossing the coin
p = 0.5 # probability of coin appearing up as head
prob = stats.binom(n = n, p = p).pmf(k = k_range)
# Plot the probability distribution using the probabilities list
# we created above.
plt.step(k_range, prob, where = 'mid', color = blue)
plt.xlabel('Number of heads')
plt.ylabel('Probability')
plt.plot((22, 22), (0, 0.1599), color = red)
plt.annotate('0.8%', xytext = (25, 0.08), xy = (22, 0.08),
va = 'center', color = red, size = 'large',
arrowprops = {'arrowstyle': '<|-', 'lw': 2,
'color': red, 'shrinkA': 10})
plt.show()
# -
# The visualization above shows the probability distribution for flipping a fair coin 30 times. Using this visualization we can now determine the probability of getting, say for example, 12 heads in 30 flips, which looks to be about 8%. Notice that we've labeled our example of 22 heads as 0.8%. If we look at the probability of flipping exactly 22 heads, it looks likes to be a little less than 0.8%, in fact if we calculate it using the function from above, we get 0.5%.
prob = stats.binom(n = n, p = p).pmf(k = 22)
print('Probability of flipping 22 heads: {:0.1f}%'.format(prob * 100))
# So, then why do we have 0.8% labeled in our probability distribution above? Well, that's because we are showing the probability of getting at least 22 heads, which is also known as the **p-value**.
#
# Let's pull back from our example and discuss formally about hypothesis testing. In standard frequentist statistic's hypothesis testing, we start with a null hypothesis that we usually call $H_0$ (pronouced as H naught), which represents our status quo. On the other hand, we also have an alternative hypothesis our $H_1$ that represents the question that we wish to answer, i.e. what we’re testing for.
#
# After setting up our null and alternative hypothesis, we conduct a hypothesis test under the assumption that the null hypothesis is true. If the test results suggest that the data do not provide convincing evidence for the alternative hypothesis, we stick with the null hypothesis. If they do, then we reject the null hypothesis in favor of the alternative.
#
# Frequentist statistic's hypothesis testing uses a p-value to weigh the strength of the evidence (what the data is telling you about the population). p-value is defined as **the probability of obtaining the observed or more extreme outcome, given that the null hypothesis is true (not the probability that the alternative hypthesis is true)**. It is a number between 0 and 1 and interpreted in the following way:
#
# - A small p-value (typically <= 0.05, 0.05 is a commonly used threshold, the threshold is often denoted as $\alpha$) indicates strong evidence against the null hypothesis, so we reject the null hypothesis. This means that something interesting is going on and it’s not just noise!
# - A large p-value (> 0.05) indicates weak evidence against the null hypothesis, so we fail to reject the null hypothesis. Although p-value is still in our favor, we cannot conclusively say that it was not due to random noise.
# - p-values very close to the cutoff (0.05) are considered to be marginal (could go either way). If you carefully read good papers on these kind of topics, you will always see the p-values being reported so that the readers can draw their own conclusions.
#
# **Example:**
#
# Let's say that a pizza place claims their delivery times are 30 minutes or less on average. Now we think it's actually takes more than 30 minutes. We conduct a hypothesis test because we believe the null hypothesis, that the mean delivery time is 30 minutes maximum, is incorrect. This means that our alternative hypothesis is the mean time is greater than 30 minutes. We randomly sample some delivery times and run the data through the hypothesis test, and our p-value turns out to be 0.01, which is much less than 0.05.
#
# In real terms, there is a probability of 0.001 that we will mistakenly reject the pizza place's claim that their delivery time is less than or equal to 30 minutes. Since typically we are willing to reject the null hypothesis when this probability is less than 0.05, we conclude that the pizza place is wrong; their delivery times are in fact more than 30 minutes on average.
# Back with our coin toss example, the null hypothesis assumes we have a fair coin, and the way we determine if this hypothesis is true or not is by calculating how often flipping this fair coin 30 times would result in 22 or more heads. If we then take the number of times that we got 22 or more heads and divide that number by the total of all possible permutations of 30 coin tosses, we get the probability of tossing 22 or more heads with a fair coin. This probability is essentially our p-value.
# +
def compute_pvalue(n, k, p):
"""Returns the p-value for binomial distribution"""
k_range = range(k, n + 1)
pvalue = stats.binom(n = n, p = p).pmf(k = k_range).sum()
return pvalue
pvalue = compute_pvalue(n = 30, k = 22, p = 0.5)
print('P-value: {:0.1f}%'.format(pvalue * 100))
# -
# The role of p-value is used to check the validity of the null hypothesis. The way this is done is by agreeing upon some predetermined upper limit for our p-value, below which we will assume that our null hypothesis is false.
#
# In other words, if our null hypothesis were true, and 22 heads in 30 flips could happen often enough by chance, we would expect to see it happen more often than the given threshold percentage of times. So, for example, if we chose 10% as our p-value threshold, then we would expect to see 22 or more heads show up at least 10% of the time to determine that this is a chance occurrence and not due to some bias in the coin. Historically, the generally accepted threshold has been 5%, and so if our p-value is less than 5%, we can then make the assumption that our coin may not be fair.
#
# Running the code above gives us a p-value of roughly 0.8%, which matches the value in our probability distribution above and is also less than the 5% threshold needed to reject our null hypothesis, so it does look like we may have a biased coin.
# we can also use the binom_test function from scipy to
# perform the hypothesis testing
pvalue = stats.binom_test(x = 22, n = 30, p = 0.5, alternative = 'greater')
print('P-value: {:0.1f}%'.format(pvalue * 100))
# ## Simulation
#
# Instead of using the stastistical approach, the code below seeks to answer the same question of whether or not our coin is fair by running a large number of simulated coin flips and calculating the proportion of these experiments that resulted in at least 22 heads or more.
# +
def coin_toss(n_simulation = 100000):
"""
computing a fair coin resulting in at
least 22 heads or more through simulation
"""
pvalue = 0
for i in range(n_simulation):
# trials: 1 denotes head, 0 denotes tail
trials = np.random.randint(2, size = 30)
if trials.sum() >= 22:
pvalue += 1
pvalue /= n_simulation
return pvalue
pvalue = coin_toss()
print('Simulated P-value: {:0.1f}%'.format(pvalue * 100))
# -
# The result of our simulations is 0.8%, the exact same result we got earlier when we calculated the p-value using the classical method above.
# # Frequentist A/B testing
#
# A/B testing is essentially a simple randomized trial. Randomized trials are (usually) considered the gold standard study design for evaluating the efficacy of new medical treatments, but they are also used much more widely in experimental research.
#
# For example, when someone visits a website, the site sends them to one of two (or possibly more) different landing or home pages, and which one they are sent to is chosen at random. The purpose is to determine which page version generates a superior outcome, e.g. which page generates more advertising revenue, or which which page leads a greater proportion of visitors to continue visiting the site.
#
# The key idea is that because we randomize which landing page (or treatment in the case of a randomized clinical trial) someone goes to, after a large number of visitors, the groups of people who visited the two pages are completely comparable in respect of all characteristics (e.g. age, gender, location, and anything else you can think of!). Because the two groups are comparable, we can compare the outcomes (e.g. amount of advertising revenue) between the two groups to obtain an unbiased, and fair, assessment of the relative effectiveness (in terms of our defined outcome) of the two designs.
#
# Suppose for the moment that we've had two visitors to our site, and one visitor has been randomized to page A, and the other visitor to page B (note that it is entirely possible, with simple randomization, that both visitors could have both been sent to page A). Suppose next that the visitor to page A generated revenue, but the visitor to page B generated no revenue. Should we conclude that page A is superior to page B, in terms of revenue generation? Of course not. Because we have only sampled two visitors, it is entirely possible that the visitor to page A would have generated revenue even if they had been sent to page B, perhaps because they are very interested in the site's content, whereas perhaps the visitor to page B was not particularly interested in the site content, and was never going to generate revenue. We can overcome this problem by running the A/B testing for a sufficiently large number of visitors, such that the probability that the scenario described above is sufficiently small.
#
# Scenario: We ran an A/B test with two different versions of a web page, a and b, for which we count the number of visitors and whether they convert or not. We can summarize this in a contingency table showing the frequency distribution of the events:
data = pd.DataFrame({
'version': ['A', 'B'],
'not_converted': [4514, 4473],
'converted': [486, 527]
})[['version', 'not_converted', 'converted']]
data
# It is trivial to compute the conversion rate of each version, 486/(486 + 4514) = 9.72% for a and 10.5% for b. With such a relatively small difference, however, can we convincingly say that the version b converts better? To test the statistical significance of a result like this, a hypothesis testing can be used.
# ## Comparing Two Proportions
#
# Let's formalize our thought process a little bit, suppose that we have obtained data from n visitors, $n_A$ of which have been (randomly) sent to page A, and $n_B$ of which have been sent to page B. Further, let $X_A$ and $X_B$ denote the number of visitors for whom we obtained a 'successful' outcome in the two groups. The proportion of successes in the two groups is then given by $\hat{p_A} = X_A/n_A$ and $\hat{p_B} = X_B/n_B$ respectively. The estimated difference in success rates is then give by the difference in proportions: $\hat{p_A} - \hat{p_B}$:
#
# To assess whether we have statistical evidence that the two pages' success rates truely differ, we can perform a hypothesis test. The null hypothesis that we want to test is that the two pages' true success rates are equal, whereas the alternative is that they differ (one is higher than the other). If $p_A$ = the proportion of the page A population whom we obtained a successful outcome and $p_B$ = the proportion of the page B population whom we obtained a successful outcome then we are interested in testing the following hypothesis:
#
# \begin{align}
# H_0:p_A = p_B \text{ versus } H_A: p_A \neq p_B
# \end{align}
#
# Or put it in another way, the null hypothesis says that the factors 'page type' and 'outcome' are statistically independent of each other. In words, this means knowing which page someone is sent to tells you nothing about the chance that they will have a successful outcome. Now that we know what hypothesis test we're interested in, we'll have to derive the appropriate test statistic.
#
# A test statistic is a single metric that can be used to evaluate the null hypothesis and the standard way to obtain this metric is to compute the z-score that measures how many standard deviations below or above the population mean a raw score is:
#
# \begin{align}
# z = \frac{x - \mu}{SE}
# \end{align}
#
# Where:
#
# - $\mu$ denotes the mean
# - $\sigma$ denotes the standard error, computed by $\frac{s}{\sqrt{n}}$, where $s$ denotes the standard error and $n$ denotes the number of samples
#
# The following link contains an example of where this is applied in proportion hypothesis testing for those who feels uncomfortable with this concept. [Notes: Eberly College of Science STAT 414/415: Test About Proportions](https://onlinecourses.science.psu.edu/stat414/node/265)
#
# For our test the underlying metric is a binary yes/no variable (event), which means the appropriate test statistic is a test for differences in proportions:
#
# \begin{align}
# Z = \frac{ (\hat{p_A} - \hat{p_B}) - (p_A - p_B) }{SE(p_A - p_B)}
# \end{align}
#
# The test statistic makes sense as it measuring the difference in the observed proportions and the estimated proportion, standardized by an estimate of the standard error of this quantity.
#
# To compute the test statistic, we first need to find the standard deviation/variance of $p_A - p_B$:
#
# \begin{align}
# Var(p_A - p_B)
# &= Var(p_A) + Var(p_B) \\
# &= \frac{p_A (1 - p_A)}{n_A} + \frac{p_B (1 - p_B)}{n_B} \\
# &= p (1 - p) \left( \frac{1}{n_A} + \frac{1}{n_B} \right)
# \end{align}
#
# - The first step stems from that fact that, given that we know:
# - The variance of a random variable X is defined as $Var(X) = E[X^2] - E[X]^2$
# - The covariance between two random variable X and Y is defined as $Cov(X, Y) = E[(X - u_x)(y - u_y)] = E[XY] - E[X]E[Y]$
# - When conducting hypothesis test, we know that the two groups should be independent of each other, i.e. the covariance between the two should be 0
#
# \begin{align}
# Var(X - Y)
# &= E[(X - Y)(X - Y)] - E[X - Y]^2 \\
# &= E[X^2 - 2XY + Y^2] - (u_x - u_y)^2 \\
# &= E[X^2 - 2XY + Y^2] - u_x^2 + 2u_xu_y - u_y^2 \\
# &= (E[X^2] - u_x^2) + (E[Y^2] - u_y^2) - 2(E[XY] - u_xu_y) \\
# &= Var(X) + Var(Y) - 2 Cov(X, Y)
# \end{align}
#
# - We're using the property that the variance of a binomial proportion is given by: $Var(p_A) = p_A (1 - p_A) / n_A$, the same can be applied for group B
# - The third step comes from the fact that if we assume that the null hypothesis, $p_A = p_B$ is true, then the population proportions equal some common value $p$, that is, $p_A = p_B = p$. Since we don't know the assumed common population proportion $p$ any more than we know the proportions $p_A$ and $p_B$ of each population, we can estimate $p$ using the proportion of "successes" in the two combined, $\hat{p} = (X_A + X_B)/(n_A + n_B)$, which is commonly referred to as the **pooled probability**
#
# During the third step, we utilized that fact that if we assume that the null hypothesis is true, then $p_A = p_B$, this also means $p_A - p_B = 0$. Given all of these information, the formula for our test statistic now becomes:
#
# \begin{align}
# Z
# &= \frac{ (\hat{p_A} - \hat{p_B}) - (p_A - p_B) }{SE(p_A - p_B)} \\
# &= \frac{ (\hat{p_A} - \hat{p_B}) - 0 }{\sqrt{\hat{p} (1 - \hat{p}) \left( \frac{1}{n_A} + \frac{1}{n_B} \right)}}
# \end{align}
#
# Where $\hat{p} = (X_A + X_B)/(n_A + n_B)$
def two_proprotions_test(success_a, size_a, success_b, size_b):
"""
A/B test for two proportions;
given a success a trial size of group A and B compute
its zscore and pvalue
Parameters
----------
success_a, success_b : int
Number of successes in each group
size_a, size_b : int
Size, or number of observations in each group
Returns
-------
zscore : float
test statistic for the two proportion z-test
pvalue : float
p-value for the two proportion z-test
"""
prop_a = success_a / size_a
prop_b = success_b / size_b
prop_pooled = (success_a + success_b) / (size_a + size_b)
var = prop_pooled * (1 - prop_pooled) * (1 / size_a + 1 / size_b)
zscore = np.abs(prop_b - prop_a) / np.sqrt(var)
one_side = 1 - stats.norm(loc = 0, scale = 1).cdf(zscore)
pvalue = one_side * 2
return zscore, pvalue
# +
success_a = 486
size_a = 5000
success_b = 527
size_b = 5000
zscore, pvalue = two_proprotions_test(success_a, size_a, success_b, size_b)
print('zscore = {:.3f}, pvalue = {:.3f}'.format(zscore, pvalue))
# +
# or we can use the implementation from statsmodels
# where we pass in the success (they call the argument counts)
# and the total number for each group (they call the argument nobs,
# number of observations)
counts = np.array([486, 527])
nobs = np.array([5000, 5000])
zscore, pvalue = proportions_ztest(counts, nobs, alternative = 'two-sided')
print('zscore = {:.3f}, pvalue = {:.3f}'.format(zscore, pvalue))
# -
# Based on the fact that our p-value is not smaller than the 0.05 commonly used threshold, the test statistic tells us we do not have strong evidence against our null hypothesis, i.e. we do not have strong evidence that the two pages are not equally effective.
#
# Apart from spitting out the p-value, we will also look at forming a confidence interval for $\hat{p_A} - \hat{p_B}$. If the number of trials in both groups is large, and the observed number of successes are not too small, we can calculate a 95% confidence interval using the formula:
#
# \begin{align}
# \text{point estimate} \pm z * SE
# &= (\hat{p_A} - \hat{p_B}) \pm z * \frac{p_A (1 - p_A)}{n_A} + \frac{p_B (1 - p_B)}{n_B}
# \end{align}
#
# Note that when calculating the confidence interval because we no longer have the assumption that $p_A = p_B$ from our null hypothesis, thus we can't leverage this property and use the pooled probability.
def two_proprotions_confint(success_a, size_a, success_b, size_b, significance = 0.05):
"""
A/B test for two proportions;
given a success a trial size of group A and B compute
its confidence interval;
resulting confidence interval matches R's prop.test function
Parameters
----------
success_a, success_b : int
Number of successes in each group
size_a, size_b : int
Size, or number of observations in each group
significance : float, default 0.05
Often denoted as alpha. Governs the chance of a false positive.
A significance level of 0.05 means that there is a 5% chance of
a false positive. In other words, our confidence level is
1 - 0.05 = 0.95
Returns
-------
prop_diff : float
Difference between the two proportion
confint : 1d ndarray
Confidence interval of the two proportion test
"""
prop_a = success_a / size_a
prop_b = success_b / size_b
var = prop_a * (1 - prop_a) / size_a + prop_b * (1 - prop_b) / size_b
se = np.sqrt(var)
# z critical value
confidence = 1 - significance
z = stats.norm(loc = 0, scale = 1).ppf(confidence + significance / 2)
# standard formula for the confidence interval
# point-estimtate +- z * standard-error
prop_diff = prop_b - prop_a
confint = prop_diff + np.array([-1, 1]) * z * se
return prop_diff, confint
prop_diff, confint = two_proprotions_confint(success_a, size_a, success_b, size_b)
print('estimate difference:', prop_diff)
print('confidence interval:', confint)
# Up till this point, we've been using the 5000 as the total number of observations/samples that are involved in the A/B testing process. The next question that we'll address is, in real world scenarios, how many obeservations do we need in order to draw a valid verdict on the test result. This leads us to our next topic **power**.
#
#
# ## Introducing Power
#
# In the world of hypothesis testing, rejecting the null hypothesis when it is actually true is called a type 1 error, often denoted as $\alpha$. Committing a type 1 error is a false positive because we end up recommending something that does not work. Conversely, a type 2 error, often denoted as $\beta$, occurs when you do not reject the null hypothesis when it is actually false. This is a false negative because we end up sitting on our hands when we should have taken action. We need to consider both of these types of errors when choosing the sample size.
#
# Two important probabilities related to type 1 and type 2 error are:
#
# - **Significance level:** Governs the chance of a false positive. A significance level of 0.05 means that there is a 5% chance of a false positive. Choosing level of significance is an arbitrary task, but for many applications, a level of 5% is chosen, for no better reason than that it is conventional
# - **Statistical power** Power of 0.80 means that there is an 80% chance that if there was an effect, we would detect it (or a 20% chance that we'd miss the effect). In other words, power is equivalent to $1 - \beta$. There are no formal standards for power, most researchers assess the power of their tests using 0.80 for adequacy
#
# | Scenario | $H_0$ is true | $H_0$ is false |
# |:--------------:|:----------------------------------:|:-------------------------:|
# | Accept $H_0$ | Correct Decision | Type 2 Error (1 - power) |
# | Reject $H_0$ | Type 1 Error (significance level) | Correct decision |
#
# The concepts of power and significance level can seem somewhat convoluted at first glance. A good way to get a feel for the underlying mechanics is to plot the probability distribution of $Z$ assuming that the null hypothesis is true. Then do the same assuming that the alternative hypothesis is true, and overlay the two plots.
#
# Consider the following example: $H_0: p_A = p_B, H_1: p_A > p_B$. A one-sided test was chosen here for charting-simplicity.
#
# - Total sample size, N=5,000 (assume equal sample sizes for the control and experiment groups, meaning exactly 2,500 in each group)
# - Say we decide that we need to observe a difference of 0.02 (detailed later) in order to be satisfied that the intervention worked (i.e., assuming that our original baseline, $p_B$ was 0.08, then we want $p_A = 0.10$). We will discuss how to make this decision later in the post
def plot_power(min_diff, prob_b, size_a, size_b, significance = 0.05):
"""illustrating power through a one-tailed hypothesis test"""
# obtain the z-score for the minimum detectable
# difference using proportion_ztest
prob_a = prob_b + min_diff
count_a = size_a * prob_a
count_b = size_b * prob_b
counts = np.array([count_a, count_b])
nobs = np.array([size_a, size_b])
zscore, _ = proportions_ztest(counts, nobs, alternative = 'larger')
# distribution for the null hypothesis, h0
# and alternative hypothesis, h1
h0 = stats.norm(loc = 0, scale = 1)
h1 = stats.norm(loc = zscore, scale = 1)
# points that are greater than the zscore for the
# specified significance level
x = np.linspace(-5, 6, num = 100)
threshold = h0.ppf(1 - significance)
mask = x > threshold
# power is the area after the thresold, i.e.
# 1 - the cumulative distribution function of that point
power = np.round(1 - h1.cdf(threshold), 2)
hypotheses = [h1, h0]
labels = ['$H_1$ is true', '$H_0$ is true']
for hypothesis, label in zip(hypotheses, labels):
y = hypothesis.pdf(x)
line = plt.plot(x, y, label = label)
plt.fill_between(x = x[mask], y1 = 0.0, y2 = y[mask],
alpha = 0.2, color = line[0].get_color())
title = 'p1: {}, p2: {}, size1: {}, size2: {}, power: {}'
plt.title(title.format(prob_a, prob_b, size_a, size_b, power))
plt.legend()
plt.tight_layout()
plt.show()
# +
prob_b = 0.08
min_diff = 0.02
size_a = 2500
size_b = 2500
plot_power(min_diff, prob_b, size_a, size_b)
# -
# The shaded green area denotes the significance region, while the the shaded blue area denotes the power (note that it includes the shaded green area). Note that if we pick a smaller N, or a smaller probability difference between the control and experiment group, the power drops (the shaded blue area decreases), meaning that if there’s is in fact a change, there’s lesser percent chance that we’ll detect it.
# +
# smaller N
prob_b = 0.08
min_diff = 0.02
size_a = 1250
size_b = 1250
plot_power(min_diff, prob_b, size_a, size_b)
# +
# smaller probability difference
prob_b = 0.08
min_diff = 0.001
size_a = 2500
size_b = 2500
plot_power(min_diff, prob_b, size_a, size_b)
# -
# The following link illustrates power for a two-sided hypothesis test for those interested. [Youtube: Calculating Power and the Probability of a Type II Error (A Two-Tailed Example)](https://www.youtube.com/watch?v=NbeHZp23ubs)
#
#
# ## Determining Sample Size
#
# Say we've followed the rule of thumb and require the significance level to be 5% and the power to be 80%. This means we have now specified two key components of a power analysis.
#
# - A decision rule of when to reject the null hypothesis. We reject the null when the p-value is less than 5%.
# - Our tolerance for committing type 2 error (1−80%=20%).
#
# To actually solve for the equation of finding the suitable sample size, we also need to specify the detectable difference, the level of impact we want to be able to detect with our test.
#
# In order to explain the dynamics behind this, we'll return to the definition of power: the power is the probability of rejecting the null hypothesis when it is false. Hence for us to calculate the power, we need to define what "false" means to us in the context of the study. In other words, how much impact, i.e., difference between test and control, do we need to observe in order to reject the null hypothesis and conclude that the action worked?
#
# Let's consider two illustrative examples: if we think that an event rate reduction of, say, $10^{-10}$ is enough to reject the null hypothesis, then we need a very large sample size to get a power of 80%. This is pretty easy to deduce from the charts above: if the difference in event rates between test and control is a small number like $10^{-10}$, the null and alternative probability distributions will be nearly indistinguishable. Hence we will need to increase the sample size in order to move the alternative distribution to the right and gain power. Conversely, if we only require a reduction of 0.02 in order to claim success, we can make do with a much smaller sample size.
#
# > The smaller the detectable difference, the larger the required sample size
#
# Here's how we could conduct a power test in python:
# +
import statsmodels.stats.api as sms
def compute_sample_size(prop1, min_diff, significance = 0.05, power = 0.8):
"""
Computes the sample sized required for a two-proportion A/B test;
result matches R's pwr.2p.test from the pwr package
Parameters
----------
prop1 : float
The baseline proportion, e.g. conversion rate
min_diff : float
Minimum detectable difference
significance : float, default 0.05
Often denoted as alpha. Governs the chance of a false positive.
A significance level of 0.05 means that there is a 5% chance of
a false positive. In other words, our confidence level is
1 - 0.05 = 0.95
power : float, default 0.8
Often denoted as beta. Power of 0.80 means that there is an 80%
chance that if there was an effect, we would detect it
(or a 20% chance that we'd miss the effect)
Returns
-------
sample_size : int
Required sample size for each group of the experiment
References
----------
R pwr package's vignette
- https://cran.r-project.org/web/packages/pwr/vignettes/pwr-vignette.html
Stackoverflow: Is there a python (scipy) function to determine parameters
needed to obtain a target power?
- https://stackoverflow.com/questions/15204070/is-there-a-python-scipy-function-to-determine-parameters-needed-to-obtain-a-ta
"""
prop2 = prop1 + min_diff
effect_size = sms.proportion_effectsize(prop1, prop2)
sample_size = sms.NormalIndPower().solve_power(
effect_size, power = power, alpha = significance, ratio = 1)
return sample_size
# -
sample_size = compute_sample_size(prop1 = 0.1, min_diff = 0.02)
print('sample size required per group:', sample_size)
# Note that the printed result is the sample size needed for each group!
#
# Unlike the significance level and the power, there are no plug-and-play values we can use for the detectable difference. The key is to define what "pay off" means for the study at hand, which depends on what the adverse event is a well as the cost of the action. Two guiding principles:
#
# - **Avoid wasteful sampling** Let’s say it takes an absolute difference of 0.02 between test and control in order for the treatment to pay off. In this case, aiming for a 0.01 detectable difference would just lead to more precision than we really need. Why have the ability to detect 0.01 if we don’t really care about a 0.01 difference? In many cases, sampling for unnecessary precision can be costly and a waste of time
# - **Avoid missed opportunities** Conversely, if we are analyzing a sensitive metric where small changes can have a large impact e.g. email campaigns, we have to aim for a small detectable difference. If we choose an insufficient sample size, we may end up sitting on our hands and missing an opportunity (type 2 error)
#
# Hence, choosing the minimum detectable difference should be a cross-functional analysis/discussion between the data scientist and the business stakeholder. Once there is a viable range for the detectable difference, we can evaluate the sample size required for each option. For example, let’s say that $p1=0.10$ and we want the detectable difference to be between 0.01 and 0.03. Clearly, we’d rather be able to detect a difference of 0.01, but it may be too costly and hence we want to evaluate more conservative options as well.
# +
# calculate the the required sample size
# for a range of minimum detectable difference
sample_sizes = []
min_diffs = np.arange(0.01, 0.03, 0.001)
for min_diff in min_diffs:
sample_size = compute_sample_size(prop1 = 0.1, min_diff = min_diff)
sample_sizes.append(sample_size)
plt.plot(min_diffs, sample_sizes)
plt.title('Sample Size Required for the Minimum Detectable Difference')
plt.ylabel('Sample Size')
plt.xlabel('Minimum Detectable Difference')
plt.tight_layout()
plt.show()
# -
# From the graph, we can see that we need roughly 10x more observations to get a detectable difference of 0.01 compared to 0.03.
#
# The following section is an alternative way of conducting a test statistic for proportional A/B test, feel free to skip it, it will not affect that understanding of later section.
# ## Alternative View of the Test Statistic
#
# There are two types of the chi-squared test, goodness of fit and test of independence, but it is the latter which is useful for the case in question. The reason as to why a test of “independence” is applicable becomes clear by converting the contingency table into a probability matrix by dividing each element by the grand total of frequencies:
cols = ['not_converted', 'converted']
data[cols] = data[cols] / data[cols].values.sum()
data
# We will denote $V$ as the version of the web page ($a$ or $b$) and $C$ as the conversion result, $f$ (false did not convert) or $t$ (true did in fact convert). The table that we computed above, which this the data that we observed can then be translated into this form:
#
#
# | Version (V) | $f$ (false did not convert) | $t$ (true did in fact convert) |
# |:-----------:|:----------------------------:|:------------------------------:|
# | A | $P(V = a, C = f)$ | $P(V = a, C = t)$ |
# | B | $P(V = b, C = f)$ | $P(V = b, C = t)$ |
#
#
# Now, our interest is whether the conversion $C$ depends on the page version $V$, and if it does, to learn which version converts better. In probability theory, the events $C$ and $V$ are said to be independent if the joint probability can be computed by $P(V, C) = P(V) \cdot P(C)$, where $P(V)$ and $P(C)$ are marginal probabilities of $V$ and $C$, respectively. It is straightforward to compute the marginal probabilities from row and column marginals:
#
# $$P(V = a) = \frac{4514 + 486}{10000} \hspace{1cm} P(V = b) = \frac{4473 + 527}{10000}$$
# $$P(C = f) = \frac{4514 + 4473}{10000} \hspace{1cm} P(V = b) = \frac{486 + 527}{10000}$$
#
# The null hypothesis is that $V$ and $C$ are independent, in which case the elements of the matrix, a.k.a the distribution that we're expecting is equivalent to:
#
# | Version (V) | $f$ (false did not convert) | $t$ (true did in fact convert) |
# |:-----------:|:----------------------------:|:------------------------------:|
# | A | $P(V = a)P(C = f)$ | $P(V = a)P(C = t)$ |
# | B | $P(V = b)P(C = f)$ | $P(V = b)P(C = t)$ |
#
#
# The conversion $C$ is said to be dependent on the version $V$ of the web site if this null hypothesis is rejected. Hence rejecting the null hypothesis means that one version is better at converting than the other. This is the reason why the test is on independence.
#
# When dealing with counts and investigating how far the observed counts are from the expected counts, we use a test statistic called the **chi-square test**. The chi-squared test compares an observed distribution $O_{ij}$ to an expected distribution $E_{ij}$:
#
# \begin{align}
# \chi^2 = \sum_{i,j} \frac{(O_{ij} - E_{ij})^2}{E_{ij}}
# \end{align}
#
# It's calculated as the observed minus the expected for each cell squared divided by the expected counts, the division with the expected counts makes final result proportional to our expected frequency. After performing the computation for each cell, we want to sum this over all of the cells (levels of the categorical variable).
#
# This $\chi^2$ probability distribution has only one parameter, the degrees of freedom. It influences the shape, the center and the spread of the chi-square distribution.
# +
# chi square distribution with varying degrees of freedom
fig = plt.figure(figsize = (8, 6))
x = np.linspace(0, 5, 1000)
deg_of_freedom = [1, 2, 3, 4]
for df in deg_of_freedom:
plt.plot(x, stats.chi2.pdf(x, df), label = '$df={}$'.format(df))
plt.xlim(0, 5)
plt.ylim(0, 0.5)
plt.xlabel('$\chi^2$')
plt.ylabel('$f(\chi^2)$')
plt.title('$\chi^2\ \mathrm{Distribution}$')
plt.legend()
plt.show()
# -
# chi-square distribution gives a way of measuring the difference between the frequencies we observe and the frequencies we expect. The smaller the value of $\chi^2$, the smaller the difference overall between the observed and expected frequencies. The way to compute the degree of freedom for the test of independence using a $r \times c$ contingency matrix is:
#
# \begin{align}
# df = (r - 1)(c - 1)
# \end{align}
#
# Where $r$ denotes the number of rows and $c$ denotes the number of columns. The rationale behind this calculation is because degrees of freedom is the number of expected frequencies we have to calculate independently after taking into account any restrictions. The restrictions come from the row and column sum constraints, but decreased by one because the last entry in the table/matrix is determined by either the row or column sum on that row/column.
#
# Fortunately it is very straightforward to carry out this hypothesis testing using packages. All we need is to supply the function with a contingency matrix and it will return the $\chi^2$ statistic and the corresponding p-value:
# +
# we can use the proportions_chisquare function,
# where we pass in the number of successes and
# the total number of trials/observation
count = np.array([486, 527])
nobs = np.array([5000, 5000])
# note that in this case (a two sample case with two sided
# alternative), the test produces the same value as porportions_ztest
# since the chi-square distribution is the square of a normal distribution
chisq, pvalue, table = proportions_chisquare(count, nobs)
print('chisq = {}, pvalue = {}'.format(chisq, pvalue))
# +
# or the chi2_contingency function where we pass
# in the observed contingency table
observed = np.array([[4514, 486], [4473, 527]])
# more about the correction = False parameter later
result = stats.chi2_contingency(observed, correction = False)
chisq, pvalue = result[:2]
print('chisq = {}, pvalue = {}'.format(chisq, pvalue))
# -
# The result for our experiment has a $\chi^2 = 1.74$ and $p = 0.185$. Since the p-value is greater than the standard threshold 0.05, we cannot reject the null hypothesis that the page version and the conversion is independent. Therefore the difference in the conversion rates is not statistically significant.
#
# For a 2 x 2 contingency table, Yate’s chi-squared test is commonly used. This applies a correction of the form:
#
# \begin{align}
# \chi^2_{Yate's} = \sum_{i,j} \frac{(\big|O_{ij} - E_{ij}\big| - 0.5)^2}{E_{ij}}
# \end{align}
#
# to account for an error between the observed discrete distribution and the continuous chi-squared distribution (the step of -0.5 is often referred to as continuity correction).
# we can use the correcction form, by specifying
# correction = True
result = stats.chi2_contingency(observed, correction = True)
chisq, pvalue = result[:2]
print('chisq = {}, pvalue = {}'.format(chisq, pvalue))
# Again, our pvalue is greater than the critical value, hence we simply would not reject the null hypothesis (that there is no relationship between the categorical variables).
#
# > Side note: in practice, we want to make sure that each particular scenario or cell has at least five expected counts before employing the chi-square test.
# # Frequentist A/B Testing Workflow
#
#
# After diving into the technical details of conducting a frequentist A/B testing, we will now introduce one possible template/workflow/thought-process for conducting A/B testing.
#
#
# ## Formulate Business Goals & Hypothesis Test
#
# **Define Business Goal**
#
# Every project or plan or test always starts with a goal e.g. A business objective for an online flower store is to "Increase our sales by receiving online orders for our bouquets"
#
# **Formulate A/B Test**
#
# The crux of A/B testing can be summarized into one sentence:
#
# > If **[Variable]**, then **[Result]**, because **[Rationale]**
#
# - **[Variable]** is the element such as call to action, media that we've modified
# - **[Result]** is basically what we expect to see, such as more clicks, more sign-ups. The effect size of [Result] will be determined by the data
# - **[Rationale]** what assumptions will be proven right/wrong after the experiment
#
#
# ### Result
#
# We start by asking ourselves, what result are we expecting out of this test? To do this, we need to:
#
# - **Define our Key Performance Indicators.** e.g. Our flower store’s business objective is to sell bouquets. Our KPI could be number of bouquets sold online.
# - **Define our target metrics.** e.g. For our imaginary flower store, we can define a monthly target of 175 bouquets sold.
# ### Rationale
#
# A lot of times, people have the idea that A/B testing is panacea, too many people think they'll just guess their way to great conversion and revenue, when trully successful tests are typically much more complicated than that.
#
# After defining the high level goal and knowing the result that we're aiming for, find out (not guess) which parts of our business are underperforming or trending and why. Ways to perform this step are:
#
# **Quantitative methods** We can start by looking at quantitative data if we have any. These methods do a much better job answering how many and how much types of questions.
#
# Say we're a website, we can take a look at our conversion funnel and examine the flow from the persuasive end (top of the funnel) and the transactional end (bottom of the funnel). e.g. We can identify problems by starting from the top 5 highest bounce rate pages. During the examination, segment to spot underlying underperformance or trends.
#
# - **Segment by source:** Separate people who arrive on your website from e-mail campaigns, google, twitter, youtube, etc. Find answers to questions like: Is there a difference between bounce rates for those segments? Is there a difference in Visitor Loyalty between those who came from Youtube versus those who came from Twitter? What products do people who come from Youtube care about more than people who come from Google?
# - **Segment by behavior:** Focus on groups of people who have similar behaviors For example, we can separate out people who visit more than ten times a month versus those that visit only twice. Do these people look for products in different price ranges? Are they from different regions? Or separate people out by the products they purchase, by order size, by people who have signed up.
#
# e.g. We're looking at our metric of total active users over time and we see a spike in one of the timelines. After confirming that this is not caused by seasonal variation, we can look at different segment of our visitors to see if one of the segment is causing the spike. Suppose we have chosen segment to be geographic, it might just happen that we’ve identify a large proportion of the traffic is generated by a specific region
#
# During the process we should ask ourselves: 1) Why is it happening? 2) How can we spread the success of other areas of the site. And it might be best for us to use qualitative methods to dig deeper and understand why, i.e. the rationale that behind the hypothesis test.
#
# **Qualitative methods:** Ideas for gathering qualitative data to understand the why a problem exists and how to potentially fix it:
#
# - Add an exit survey on our site, asking why our visitors did/didn't complete the goal
# - Track what customers are saying in social media and on review sites
# - User Experience Group (this is the preferred way as it is going really deep with a few users and ask qualitative questions such as what's holding them back from doing what we hope they'll do, e.g. converting)
# ### Variable
#
# Now that we've identify the overall business goal and the possible issue, it's time the determine the variable, which is the element that we'll be testing for. e.g. we've identified through quantitative method that less than one percent of visitors sign up for our newsletter and after conducting qualitative studies it's because the call to action wording does not resonate with the audience, then our variable will be changing the call to action's wording.
#
# Note that we may have multiple ideas for our variable, in that case we can collate all the ideas, prioritize them based on three simple metrics:
#
# - **Potential** How much potential for a conversion rate increase? We can check to see if this kind of idea worked before.
# - **Importance** How many visitors will be impacted from the test?
# - **Ease** How easy is it to implement the test? Go for the low-hanging fruit first.
#
# Every test that's developed should documented so that we can review and prioritize ideas that are inspired by winning tests. Some ideas worth experimenting are: Headlines, CTA (call to actions), check-out pages, forms and the elements include:
#
# - Wording. e.g. Call to action or value proposition.
# - Image. e.g. Replacing a general logistics image with the image of an actual employee.
# - Layout. e.g. Increased the size of the contact form or amount of content on the page.
#
# ---
#
# So given all of that a strong A/B test hypothesis may be:
#
# - If the call to action text is changed to "Complete My Order", the conversion rates in the checkout will icnrease, because the copy is more specific and personalized
# - If the navigation link is removed from checkout pages, the conversation rate will increase because our website analytics shows portions of our traffic drop out of the funnel by clicking on those links
# ## Quantitative A/B testing
#
# So now, suppose you're running an educational platform and your A/B testing hypothesis is : Will changing the "Start Now" button from orange to pink increase how many students explore the platform's courses. So in this case the metric that's use to evaluate the change's performance is the click through probability (unique visitors who click the button / unique visitors to page). Note that it is often times impractical to use metrices such as total number of students that completed the course as it often takes weeks or months before a student can do that.
#
# Next we will jot down the hypothesis that we wish to test out, in our case the our null and alternative hypothesis would be :
#
# - $H_0$: The experimental and control groups have the same probability of clicking the button. Or equivalent to saying that the differences of the two groups' probability is 0
# - $H_1$: The two groups have different probability of completing a clicking the button
#
# ### Define the Size and Duration
#
# Now that we've defined our hypothesis, the first question that comes into mind is how many tests do we need to run, or in a sense how long should the test last in order for us to make our decisions. To do that we can use a power analysis for two independent samples:
#
# Now suppose that our current baseline is 0.1, i.e. there's a 10 percent chance that people who saw the button will click it and we wish to detect a change of 2 percent in the click through rate (This change is quite high for online experiment).
sample_size = compute_sample_size(prop1 = 0.1, min_diff = 0.02)
print('sample size required per group:', sample_size)
# The result shows that we need at least 3841 sample size for each scenario to detect if there will actually be a 2 percent more than baseline click through probability. Note that this is only telling us the minimum sample size required per group, we still need to decide when do we want to run the experiment and for how long.
#
# e.g. Suppose we’ve chosen the goal to increase click-through rates, which is defined by the unique number of people who click the button versus the number of users who visited the page that the button was located. But to actually use the definition, we’ll also have to address some other questions. Such as, if the same user visits the page once and comes back a week or two later, do we still only want to count that once? Thus we’ll also need to specify a time period
#
# To account for this, if 99% of our visitors convert after 1 week, then we should do the following.
#
# - Run our test for two weeks
# - Include in the test only users who show up in the first week. If a user shows up on day 13, we have not given them enough time to convert (click-through)
# - At the end of the test, if a user who showed up on day 2 converts more than 7 days after he/she first arrived, he must be counted as a non-conversion
#
# There will be more discussion about this in the A/B Test Caveats & Advice section.
#
# For this step, there is also an online calculator that non-technical audience could use. [Online Calculator: Sample Size Calculator](http://www.evanmiller.org/ab-testing/sample-size.html)
# ## Define the Population
#
# Another consideration is what fraction of the traffic are we going to send through the experiment. The key is to identify which population of our users will be affected by our experiment, we might want to target our experiment to that traffic (e.g. changing features specific to one language’s users) so that the rest of the population won’t dilute the effect.
#
# Next, depending on the problem we're looking at, we might want to use a cohort instead of a population. A cohort makes much more sense than looking at the entire population when testing out learning effects, examining user retention or anything else that requires the users to be established for some reason.
#
# A quick note on cohort. The gist of cohort analysis is basically putting our customers into buckets so we can track their behaviours over a period of time. The term cohort stands for a group of customers grouped by the timeline (can be week, month) where they first made a purchase (can be a different action that’s valuable to the business). Having similar traits makes the two groups more comparable.
#
# e.g. You’re an educational platform has an existing course that’s already up and running. Some of the students have completed the course, some of them are midway through and there’re students who have not yet started. If you want to change the structure of of one of the lessons to see if it improves the completion rate of the entire course and they started the experiment at time X. For students who have started before the experiment initiated they may have already finished the lesson already leading to the fact that they may not even see the change. So taking the whole population of students and running the experiment on them isn’t what you want. Instead, you want to segment out the cohort, the group of customers, that started the lesson are the experiment was launched and split that into an experiment and control group.
# ## Evaluating Result
#
# Suppose we have ran the test and we've obtain the total number of sample sizes and the total number of successes for both groups. Given these variables we can use it to calculate whether the proportional change was due to variation or not.
# +
# made-up results
success_a = 386
size_a = 3834
success_b = 530
size_b = 3842
prob_diff, confint = two_proprotions_confint(success_a, size_a, success_b, size_b)
print('estimate difference:', prob_diff)
print('confidence interval:', confint)
# -
# In order to launch a change, the change should be larger than the minimum detectable change that we wished to detect. In our case, the value we’ve set was 0.02. Base on the result above, we can denote that since even the lower bound of the confidence interval is larger than the value, we’ll definitely launch the newer version of the click button.
#
# There is also an online calculator that we can use to perform the proportion test. [Online Calculator: AB Testguide](https://abtestguide.com/calc/)
#
#
# ## Sanity Check
#
# When running experiments, especially online experiments, it's a good idea to check whether the experiments were setup properly, i.e. are the users being split equally amongst the two groups. For instance, after running your experiment for a week, you've discovered that the total number of users assigned to the control group is 64454 and the total number of users assigned to the experiment group 61818. How would you figure out whether the difference is within expectation given that each user is randomly assigned to the control or experiment group with a probability of 0.5? It's usually good idea to check this.
#
# This is equivalent to saying out of a total 126272 (64454 + 61818) users, is it surprising to see if 64454 users are assigned to the control group? This is essentially a binomial distribution, thus, knowing this information, we can construct a confidence interval to test if the number lies within the confidence interval. The confidence interval can be calculated by the mean plus and minus the z-score times the standard error.
#
# \begin{align}
# mean \pm Z * \sqrt{np(1 - p)}
# \end{align}
#
# Where the mean is expected number of users in the control / experiment group, which is simply the total number of the two groups times 0.5, since the probability of a user falling into either group is 50%. And the standard error of a binomial distribution is $\sqrt{np(1-p)}$.
def sanity_check(size1, size2, significance = 0.05):
n = size1 + size2
confidence = 1 - significance
z = stats.norm.ppf(confidence + significance / 2)
confint = n * 0.5 + np.array([-1, 1]) * np.sqrt(n * 0.5 * 0.5)
return confint
size1 = 64454
size2 = 61818
sanity_check(size1, size2)
# The result shows that 64454 does not lie within the range of the computed 95 percent confidence interval and therefore it indicates the two groups may not be split equally.
#
# When this kind of situation happens it's usually best to go back to the day by day data to get a better idea of what could be going wrong. One good thing is to check whether any particular day stands out, or it is just an overall pattern. If it is an overall pattern, then it is suggested that we should check if something went wrong with the experiment setup before proceeding on to analyzing the result.
# # A/B Test Caveats & Advices
#
#
# ## Avoid Biased Stopping Times
#
# NO PEEKING. When running an A/B test, we should avoid stopping the experiment as soon as the results "look" significant. Using a stopping time that is dependent upon the results of the experiment can inflate our false-positive rate substantially.
#
# Recall that in many experiments, we set the significance threshold to be 5% (or a p-value threshold of 0.05). This means that we’ll accept that Variation A is better than Variation B if A beats B by a margin large enough that a false positive would only happen 5% of the time. If we, however, were to check the experiment with the intent of stopping it if it shows significance, then every time we perform the significance we're essentially inflating our false-positive rate. To be more explicit, every time we perform the test there's a 5% chance of false-positive, so in other words, 95% chance of drawing the right conclusion, if we perform it again then that means we need both test to be correct to draw the right conclusion, i.e. the probability of both test giving us the correct result now becomes (1 - 5%)(1 - 5%) and the probability of commiting a false positive error is now: 1 - (1 - 5%)(1 - 5%).
# +
# the false positive rate of conducting the test for n times
significance = 0.05
print('conducting the test 2 times', 1 - (1 - significance) ** 2)
print('conducting the test 10 times', 1 - (1 - significance) ** 10)
# -
# The easiest way to avoid this problem is to **choose a stopping time that's independent of the test results**. We could, for example, decide in advance to run the test for a fix amount of time, no matter the results we observed during the test's tenure. Thus just like in the template above, if 99% of your visitors convert after 1 week, then you should do the following.
#
# - Run your test for two weeks.
# - Include in the test only users who show up in the first week. If a user shows up on day 13, you have not given them enough time to convert.
# - At the end of the test, if a user who showed up on day 2 converts more than 7 days after he first arrived, he must be counted as a non-conversion.
#
# Or you could decide to run the test until each bucket has received more than 10,000 visitors, again ignoring the test results until that condition is met. There're tests like power tests that let's you determine how many tests you should run before you make a conclusion about the result. Although you should be very careful with this, because the truth is: It's not really the number of conversions that matters; it’s whether the time frame of the test is long enough to capture variations on your site.
#
# For instance, the website traffic may behave one way during the day and another way at night (the same holds on weekdays and weekends). Then it's worth noting that there are two effects that could occur when new features are introduced: **Primacy** and **Novelty** effect.
#
# - Primacy effect occurs when we change something and experienced users may be less efficient until they get used to the new feature, thus giving inherent advantage to the control (original version)
# - Novelty effect. Meaning when users are switched to a new experience, their initial reactions may not be their long-term reactions. In other words, if we are testing a new color for a button, the user may initially love the button and click it more often, just because it’s novel, but eventually he/she would get used to the new color and behave as he/she did before. It’s important to run the trial long enough to get past the period of the "shock of the new".
#
# In sum, you should setting a results-independent stopping time (a week) is the easiest and most reliable way to avoid biased stopping times. Note that running the test for a least a week is adviced since it'll make sure that the experiment captures the different user behaviour of weekdays, weekends and try to avoid holidays ....
# ## Do Follow Up Tests and Watch your Overall Success Rate
#
# If you're running a lot of A/B tests, you should run follow-up tests and pay attention to your base success rate.
#
# Let's talk about these in reverse order. Imagine that we've done everything right. We set our stopping time in advance, and keep it independent from the test results. We set a relatively high success criterion: A probability of at least 95% that the variant is better than the control (formally, $p \leq 0.05$). We do all of that.
#
# Then We run 100 tests, each with all the rigor just described. In the end, of those 100 tests, 5 of them claims that the variant will beat the control. How many of those variants do we think are really better than the control, though? If we run 20 tests in a row in which the "best" variant is worse or statistically indistinguishable from the control, then we should be suspicious when our 21st test comes out positive. If a button-color test failed to elicit a winner six months ago, but did produce one today, we should be skeptical. Why now but not then?
#
# Here's an intuitive way of thinking about this problem. Let’s say we have a class of students who
# each take a 100-item true/false test on a certain subject. Suppose each student chooses randomly on all
# questions. Each student would achieve a random score between 0 and 100, with an average of 50.
#
# Now take only the top scoring 10% of the class, and declaring them "winners", give them a second test, on
# which they again choose randomly. They will most likely score less on the second test than the first test. That's because, no matter what they scored on the first test they will still average 50 correct answers in the second test. This is what's called the **regression to the mean**. Meaning that tests which seem to be successful but then lose their uplift over time.
#
# It can be wise to run our A/B tests twice (a validation test). You'll find that doing so helps to eliminate illusory results. If the results of the first test aren’t robust, you’ll see noticeable decay with the second. But, if the uplift is real, you should still see uplift during the second test. This approach isn’t fail-safe but it will help check whether your results are robust. e.g. In a multiple testing, you tried out three variants, B, C, and D against the control A. Variant C won. Don't deploy it fully yet. Drive 50% of your traffic to Variant C and 50% to Variant A (or some modification on this; the percent split is not important as long as you will have reasonable statistical power within an acceptable time period). As this will give you more information about C's true performance relative to A.
#
# Given the situation above, it's better to keep a record of previous tests, when they were run, the variants that were tried, etc. Since these historical record gives you an idea of what's reasonable. Despite the fact that this information is not directly informative of the rates you should expect from future tests (The absolute numbers are extremely time dependent, so the raw numbers that you get today will be completely different than the ones you would have gotten six months later), it gives you an idea of what's plausible in terms of each test's relative performance.
#
# Also, by keeping a record of previous tests, we can avoid:
#
# - Falling into the trap of "We already tried that". A hypothesis can be implemented in so many different ways. If you just do one headline test and say "we tried that," you’re really selling yourself short.
# - Not testing continually or not retesting after months or years. Just because you tested a variation in the past doesn’t necessarily mean that those results are going to be valid a year or two from now (Because we have the record of what we did, we can easily reproduce the test).
# ## False Reporting
#
# Let's say you deploy a new feature to your product and wish to see if it increases the product's activation rate (or any other metric or KPI that's relevant to you). Currently the baseline of the product's activation rate is somewhere around 40%. After running the test, you realized that it WORKED, the activation went up to 50%. So you're like, YES! I just raised activation by 25%! and you sent this info to the head of product and ask for a raise.
#
# After two months, the head of product comes back to you and said "you told me you raised the activation rate by 25%, shouldn't this mean that I should see a big jump in the overall activation? What's going on?" Well, what's going on is, you did raised activation by 25%, but only for user who uses the product's feature. So if only 10 percent of your users use that product, then the overall increase in activation rate will probably only be around 2.5% (25% * 10%). Which is still probably very good, but the expectation that you've set by mis-reporting can get you into trouble.
#
#
# ## Seasonality / Not Running it Against the Correct Target
#
# Suppose you have different types of users (or users with different usage patterns) using your product. e.g. business user and students. Then what can happen is your A/B testing will have different result in July versus October. The reason may be in July all your student users are out on vacation (not using your product) and in October after school starts they start using it again. This is simply saying that the weighting of your user population may be different in different times of the year (seasonality). Thus, you should be clear with yourself about who you're targeting.
# ## Others
#
# Despite its useful functionality, there are still places where A/B testing isn't as useful. For example:
#
# - A/B testing can't tell us if we're missing something. Meaning it can tell you if A performs better B or vice versa, but it can't tell us that if we use C, then it will actually perform better than the former two.
# - Tesing out products that people rarely buy. e.g. cars, apartments. It might be too long before the user actually decides to take actions after seeing the information and we might be unaware of the actual motivation.
# - Optimizing for the funnel, rather than the product. Understanding what the customers want so that we can make the product better. Ultimately, we can’t simply test our headlines and get people to like our product more.
# - Non-Randomized Bucketing: Double check if you're actually randomly splitting you're users, this will most likely burn you if your system assigns user id to users in a systematical way. e.g. user id whose last two digits are 70 are all from a specific region.
# - Conflicting test: Two different product teams both deployed new feautures on your landing page and ran the A/B test at the same period of time. This is more of a organization problem. You should probably require the product teams to register for their test, and make sure that multiple tests on the same stuff are not running at the same time, or else you might be tracking the effect of the other test.
# - Optimizing the wrong metric. The best example is probably noting that higher click through rate doesn't necessary means higher relevance. To be explicit, poor search results means people perform more searches, and thereby click on more ads. While this seems good in the short term, it's terrible in the long term, as users get more and more frustrated with the search engine. A search engine's goal should be to help users find what they want as quickly as possible, and sessions per user (increasing sessions per user means users are satisfied and returning) should probably be the key metric to showcase instead.
# # Reference
#
# - [Youtube: Beautiful A/B Testing](https://www.youtube.com/watch?v=EvDg7ssY0M8)
# - [Notebook: Statistics for Hackers](http://nbviewer.jupyter.org/github/croach/statistics-for-hackers/blob/master/statistics-for-hackers.ipynb)
# - [Blog: What Are P-Values?](https://prateekvjoshi.com/2013/12/07/what-are-p-values/)
# - [Blog: Interpreting A/B Test using Python](http://okomestudio.net/biboroku/?p=2375)
# - [Blog: So, You Need a Statistically Significant Sample?](http://multithreaded.stitchfix.com/blog/2015/05/26/significant-sample/)
# - [Blog: How to Build a Strong A/B Testing Plan That Gets Results](https://conversionxl.com/how-to-build-a-strong-ab-testing-plan-that-gets-results/)
# - [Blog: A/B testing and Pearson's chi-squared test of independence](http://thestatsgeek.com/2013/07/22/ab-testing/)
# - [Blog: A/B testing - confidence interval for the difference in proportions using R](http://thestatsgeek.com/2014/02/15/ab-testing-confidence-interval-for-the-difference-in-proportions-using-r/)
# - [Blog: Python for Data Analysis Part 23: Point Estimates and Confidence Intervals](http://hamelg.blogspot.com/2015/11/python-for-data-analysis-part-23-point.html)
# - [Notes: MOST winning A/B test results are illusory](http://www.qubit.com/sites/default/files/pdf/mostwinningabtestresultsareillusory_0.pdf)
# - [Notes: Eberly College of Science STAT 414/415 Comparing Two Proportions](https://onlinecourses.science.psu.edu/stat414/node/268)
# - [Quora: When should A/B testing not be trusted to make decisions?](https://www.quora.com/When-should-A-B-testing-not-be-trusted-to-make-decisions)
# - [Forbes: How To Do A/B Testing Right And Avoid The Most Common Mistakes Marketers Make](https://www.forbes.com/sites/sujanpatel/2015/10/29/how-to-do-ab-testing-right-and-avoid-the-most-common-mistakes-marketers-make/)
# - [Paper: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME> (2012) Trustworthy Online Controlled Experiments: Five Puzzling Outcomes Explained](http://notes.stephenholiday.com/Five-Puzzling-Outcomes.pdf)
# - [Slideshare: 4 Steps Toward Scientific A/B Testing](https://www.slideshare.net/RJMetrics/4-steps-toward-scientific-ab-testing)
| ab_tests/frequentist_ab_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <font size=5 color="red" face="arial">
# <h1 align="center">Universidad de Medellín
# </font>
# <font size=4 color="RED" face="arial">
# <h1 align="center">PROGRAMACIÓN EN PYTHON - NIVEL BÁSICO</h1>
# </font>
#
# <font size=2 color="RED" face="arial">
# <h1 align="center">Sesión 06b - Módulos</h1>
# </font>
# ## Instructor:
# > <strong> *<NAME>, I.C. Ph.D.* </strong>
# ### Programación Modular y Módulos
# > <strong>Programación Modular:</strong> Técnica de diseño de software, que se basa en el principio general del *diseño modular*.
#
#
# > El *Diseño Modular* es un enfoque que se ha demostrado como indispensable en la ingeniería incluso mucho antes de las primeras computadoras.
#
#
# > El *Diseño Modular* significa que un sistema complejo se descompone en partes o componentes más pequeños, es decir módulos. Estos componentes pueden crearse y probarse independientemente. En muchos casos, pueden incluso utilizarse en otros sistemas.
#
#
# > Si desea desarrollar programas que sean legibles, fiables y mantenibles sin demasiado esfuerzo, debe utilizar algún tipo de diseño de software modular. Especialmente si su aplicación tiene un cierto tamaño.
#
#
# > Existe una variedad de conceptos para diseñar software en forma modular.
#
#
# > La programación modular es una técnica de diseño de software para dividir su código en partes separadas. Estas piezas se denominan módulos.
#
#
# > El enfoque para esta separación debe ser tener módulos con no o sólo algunas dependencias sobre otros módulos. En otras palabras: La minimización de las dependencias es la meta.
#
#
# > Al crear un sistema modular, varios módulos se construyen por separado y más o menos independientemente. La aplicación ejecutable se creará reuniéndolos.
#
# #### Importando Módulos
# > Cada archivo, que tiene la extensión de archivo `.py` y consta de código `Python` adecuado, se puede ver o es un módulo!
#
#
# > No hay ninguna sintaxis especial requerida para hacer que un archivo de este tipo sea un módulo.
#
#
# > Un módulo puede contener objetos arbitrarios, por ejemplo archivos, clases o atributos. Todos estos objetos se pueden acceder después de una importación.
#
#
# > Hay diferentes maneras de importar módulos. Demostramos esto con el módulo de matemáticas:
#
#
# Si quisiéramos obtener la raíz cudrada de un número, o el valor de pi, o alguna función trigonométrica simple...
#
x = sin(2*pi)
V = 4/3*pi*r**3
x = sqrt(4)
# > Para poder usar estas funciones es necesario importar el módulo correspondiente que las contiene. En este caso sería:
import math
# > El módulo matemático proporciona constantes y funciones matemáticas, `pi (math.pi)`, la función seno (`math.sin()`) y la función coseno (`math.cos()`). Cada atributo o función sólo se puede acceder poniendo "`math`" delante del nombre:
math.pi
math.sin(math.pi/2)
math.sqrt(4)
# > Se puede importar más de un módulo en una misma sentencia de importación. En este caso, los nombres de los módulos se deben separar por comas:
import math, random
random.
# > Las sentencias de importación pueden colocarse en cualquier parte del programa, pero es un buen estilo colocarlas directamente al principio de un programa.
#
#
# > Si sólo se necesitan ciertos objetos de un módulo, se pueden importar únicamente esos:
from math import sin, pi, sqrt
# > Los otros objetos, p.ej. `cos`, no estarán disponibles después de esta importación. Será posible acceder a las funciones `sin` y `pi` directamente, es decir, sin prefijarlos con `math`.
#
#
# > En lugar de importar explícitamente ciertos objetos de un módulo, también es posible importar todo en el espacio de nombres del módulo de importación. Esto se puede lograr usando un asterisco en la importación:
from math import *
e
pi
# > - No se recomienda utilizar la notación de asterisco en una instrucción de importación, excepto cuando se trabaja en el intérprete interactivo de `Python`.
#
#
# > - Una de las razones es que el origen de un nombre puede ser bastante oscuro, porque no se puede ver desde qué módulo podría haber sido importado. Demostramos otra complicación seria en el siguiente ejemplo:
from numpy import *
sin(3)
from math import *
sin(3)
sin(3)
# > Es usual la notación de asterisco, porque es muy conveniente. Significa evitar una gran cantidad de mecanografía tediosa.
#
#
# > Otra forma de reducir el esfuerzo de mecanografía consiste en usar alias.
import numpy as np
import matplotlib.pyplot as plt
np.
# > Ahora se pueden prefijar todos los objetos de `numpy` con `np`, en lugar de `numpy`
np.diag([3, 11, 7, 9])
# #### Diseñando y Escribiendo Módulos
# > Un módulo en `Python` es simplemente un archivo que contiene definiciones y declaraciones de `Python`.
#
#
# > El nombre del módulo se obtiene del nombre de archivo eliminando el sufijo `.py`.
#
#
# > - Por ejemplo, si el nombre del archivo es `fibonacci.py`, el nombre del módulo es `fibonacci`.
#
#
# > Para convertir las funciones `Fibonacci` en un módulo casi no hay nada que hacer, sólo guardar el siguiente código en un archivo `*.py`.
#
# + active=""
# def fib(n):
# if n == 0:
# return 0
# elif n == 1:
# return 1
# else:
# return fib(n-1) + fib(n-2)
# def ifib(n):
# a, b = 0, 1
# for i in range(n):
# a, b = b, a + b
# return a
# -
# > - El recién creado módulo `fibonacci` está listo para ser usado ahora.
#
#
# > - Podemos importar este módulo como cualquier otro módulo en un programa o script.
import fibonacci
fibonacci.fib(7)
fibonacci.ifib(7)
# > - Como podrá ver, es inconveniente si tiene que usar esas funciones a menudo en su programa y siempre hay que escribir el nombre completo, es decir, `fibonacci.fib(7)`.
#
#
# > - Una solución consiste en asignar un alias para obtener un nombre más corto:
fib = fibonacci.ifib
fib(10)
# #### Contenido de un Módulo
# > Con la función incorporada `dir()` y el nombre del módulo como argumento, puede listar todos los atributos y métodos válidos para ese módulo.
import math
dir(math)
# ### Paquetes en Python
# > Los módulos son archivos que contienen instrucciones y definiciones de `Python`, como definiciones de funciones y clases.
#
#
# > - En este capítulo aprenderemos a agrupar varios módulos para formar un paquete.
#
#
# > Un paquete es básicamente un directorio con archivos `Python` y un archivo con el nombre \__`init`__`.py`.
#
#
# > - Esto significa que cada directorio dentro de la ruta de acceso de Python, que contiene un archivo llamado \__init__.py, será tratado como un paquete por Python.
#
#
# > - Es posible poner varios módulos en un paquete.
#
#
# > Los paquetes son una forma de estructurar los nombres de módulos en Python usando "nombres de módulos punteados":
#
#
# > - A.B significa un submódulo B en un paquete A.
#
#
# > - Dos paquetes diferentes como P1 y P2 pueden tener módulos con el mismo nombre (A por ejemplo).
#
#
# > - El submódulo A del paquete P1 y el submódulo A del paquete P2 pueden ser totalmente diferentes.
#
#
# > Un paquete se importa como un módulo "normal".
# #### Ejemplo
# > Mostraremos cómo crear paquetes con un ejemplo muy simple:
#
#
# > - En primer lugar, necesitamos un directorio. El nombre de este directorio será el nombre del paquete, que queremos crear.
#
#
# > - Llamaremos nuestro paquete `simple_package`.
#
#
# >> - Este directorio debe contener un archivo con el nombre "\__init__.py".
#
#
# > - Este archivo puede estar vacío o puede contener código `Python` válido.
#
#
# > - Este código se ejecutará cuando se importe un paquete, por lo que se puede utilizar para inicializar un paquete, p.
#
#
# > - Ahora podemos poner en este directorio todos los archivos de Python que serán los submódulos de nuestro módulo.
#
#
# > - Creamos dos archivos sencillos a.py y b.py sólo para llenar el paquete con módulos.
#
#
# El contenido de a.py:
# + active=""
# def bar():
# print("Hello, function 'bar' from module 'a' calling")
# -
# El contenido de b.py:
# + active=""
# def foo():
# print("Hello, function 'foo' from module 'b' calling")
# -
# Veamos lo que pasa cuando importamos *simple_package* desde el shell interactivo de Python, suponiendo que el directorio *simple_package* está en el directorio desde el que llama al shell o que está contenido en la ruta de búsqueda o en la variable de entorno "PYTHONPATH":
import simple_package
simple_package
simple_package/a
simple_package/b
# > Podemos ver que el paquete *simple_package* ha sido cargado pero no el módulo "a" ni el módulo "b"!
#
#
# > Los módulos a y b se importan de la siguiente manera:
from simple_package import a, b
a.bar()
b.foo()
# > Sin embargo, hay una manera de cargar automáticamente estos módulos.
#
#
# > - Podemos usar el archivo __init__.py para este propósito.
#
#
# > - Todo lo que tenemos que hacer es añadir las siguientes líneas al archivo vacío \__init__.py:
# + active=""
# import simple_package.a
# import simple_package.b
# -
import simple_package
simple_package.a.bar()
simple_package.b.foo()
| images/Sesion06b_Modulos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from datetime import datetime
import os
PATH = os.getcwd()
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
sns.set()
import networkx
from networkx import *
import sys
from pathlib import Path
p = (Path('.') / 'seirsplus').resolve()
if str(p) not in sys.path:
sys.path.insert(0,str(p))
from seirsplus.models import *
from seirsplus.networks import *
from seirsplus.sim_loops import *
from seirsplus.utilities import *
from seirsplus.parallel_run import *
# ## Parameter setup
# ### Social network
def get_network(N, p = 0.2):
return {
"G": Defer(gaussian_random_partition_graph,N, 20, 5, 0.5, 5/(N-20), directed=False),
"N": N,
"isolation groups": None,
"p": p
}
def random_graph(N,d, p =0.2):
return {
"G": Defer(fast_gnp_random_graph,N,d/N),
"N": N,
"p": p,
"isolation groups": None
}
# ### Infection parameters
def infect_params(N,R0_mean, R0_coeffvar= None, average_introductions_per_day = 0, single_intro_by =None,):
latentPeriod_mean, latentPeriod_coeffvar = 3.0, 0.6
SIGMA = 1 / gamma_dist(latentPeriod_mean, latentPeriod_coeffvar, N)
presymptomaticPeriod_mean, presymptomaticPeriod_coeffvar = 2.2, 0.5
LAMDA = 1 / gamma_dist(presymptomaticPeriod_mean, presymptomaticPeriod_coeffvar, N)
symptomaticPeriod_mean, symptomaticPeriod_coeffvar = 4.0, 0.4
GAMMA = 1 / gamma_dist(symptomaticPeriod_mean, symptomaticPeriod_coeffvar, N)
infectiousPeriod = 1/LAMDA + 1/GAMMA
onsetToHospitalizationPeriod_mean, onsetToHospitalizationPeriod_coeffvar = 11.0, 0.45
ETA = 1 / gamma_dist(onsetToHospitalizationPeriod_mean, onsetToHospitalizationPeriod_coeffvar, N)
hospitalizationToDischargePeriod_mean, hospitalizationToDischargePeriod_coeffvar = 11.0, 0.45
GAMMA_H = 1 / gamma_dist(hospitalizationToDischargePeriod_mean, hospitalizationToDischargePeriod_coeffvar, N)
hospitalizationToDeathPeriod_mean, hospitalizationToDeathPeriod_coeffvar = 7.0, 0.45
MU_H = 1 / gamma_dist(hospitalizationToDeathPeriod_mean, hospitalizationToDeathPeriod_coeffvar, N)
PCT_FATALITY = 0.08
PCT_HOSPITALIZED = 0.035
P_GLOBALINTXN = 0.4
if R0_coeffvar is None:
R0_coeffvar = R0_mean/10
R0 = gamma_dist(R0_mean, R0_coeffvar, N)
BETA = 1/infectiousPeriod * R0
return dict(beta=BETA, sigma=SIGMA, lamda=LAMDA,
gamma=GAMMA,
gamma_asym=GAMMA, eta=ETA, gamma_H=GAMMA_H, mu_H=MU_H,
a=PCT_ASYMPTOMATIC, h=PCT_HOSPITALIZED, f=PCT_FATALITY,
isolation_time=14,
average_introductions_per_day = average_introductions_per_day,
introduction_days = [] if single_intro_by is None else Defer(single_introduction,single_intro_by))
def infect_params_orig(N,R0, average_introductions_per_day = 0, single_intro_by =None, skip_pre= False, pct_symptomatic= 0, pre_symptomatic_period=3, infectious_period = 6.5, incubation_period = 5.2):
ETA = 0
MU_H = 0
PCT_FATALITY = 0
PCT_HOSPITALIZED = 0
PCT_ASYMPTOMATIC = 1-pct_symptomatic
BETA = R0/infectious_period
if skip_pre:
lamda = 0
gamma = 1/infectious_period
else:
gamma = 1/pre_symptomatic_period
lamda = 1/(infectious_period-pre_symptomatic_period)
if not (single_intro_by is None):
print("***",single_intro_by)
introduction_days = Defer(single_introduction,single_intro_by)
else:
introduction_days = []
return dict(skip_pre=skip_pre,beta=BETA, sigma=1/incubation_period, lamda= lamda,
gamma=gamma,
gamma_asym=gamma, eta=ETA, gamma_H=gamma, mu_H=MU_H,
a=PCT_ASYMPTOMATIC, h=PCT_HOSPITALIZED, f=PCT_FATALITY,
isolation_time=14,
average_introductions_per_day = average_introductions_per_day,
introduction_days = introduction_days )
# ### Testing and intervention parameters
# +
def run_params(N,frac_tested,frequency, testing_compliance_symptomatic=0):
MAX_TIME = 105
D= dict(T = MAX_TIME,
runTillEnd = False,
intervention_start_pct_infected = 0/100,
initI = 0, # initial number of infected people
#testing_cadence = testing_cadence, # how often to do testing (other than self-reporting symptomatics who can get tested any day)
#cadence_testing_days = cadence_testing_days, # dictionary mapping testing_cadence to schedule (None = default weekly/workday/etc..)
pct_tested_per_day = frac_tested, # max daily test allotment defined as fraction of population size
test_falseneg_rate = 0.25, # test false negative rate, will use fn rate that varies with disease time
max_pct_tests_for_symptomatics = 1, # max percent of daily test allotment to use on self-reporting symptomatics
max_pct_tests_for_traces = 1, # max percent of daily test allotment to use on contact traces
random_testing_degree_bias = 0, # magnitude of degree bias in random selections for testing, none here
pct_contacts_to_trace = 0.0, # percentage of primary cases' contacts that are traced
tracing_lag = 2, # number of cadence testing days between primary tests and tracing tests
isolation_lag_symptomatic = 1, # number of days between onset of symptoms and self-isolation of symptomatics
isolation_lag_positive = 1, # test turn-around time (tat): number of days between administration of test and isolation of positive cases
isolation_lag_contact = 0, # number of days between a contact being traced and that contact self-isolating
testing_compliance_symptomatic = testing_compliance_symptomatic,
testing_compliance_traced = 0,
testing_compliance_random = 1, # assume employee testing is mandatory, so 100% compliance
tracing_compliance = 0.0,
isolation_compliance_symptomatic_individual = 0.0,
isolation_compliance_symptomatic_groupmate = 0.0,
isolation_compliance_positive_individual = 0.0,
isolation_compliance_positive_groupmate = 0.0, # isolate teams with a positive member, but suppose 20% of employees are essential workforce
isolation_compliance_positive_contact = 0.0,
isolation_compliance_positive_contactgroupmate = 0.0,
test_priority = 'last_tested',
stopping_policy = Defer(stop_at_detection,lag=1),
budget_policy = scale_to_pool # fraction of tests is determined as fraction of eligible pool
)
if isinstance(frequency,str):
D["testing_cadence"] = frequency
else:
D[("testing_cadence","cadence_testing_days","cadence_cycle_length")] = Defer(test_frequency,frequency)
return D
# -
# ## Set up experiment
# +
totals = [1,2,7,14,28,35,52]
Rs = [1.2, 1.6, 2, 2.4, 2.8,3.2,3.6,4.0,4.4,4.8]
torun = []
SKIP_PRE = True
INIT_INFECT = 1
INTRO_DAY = 0
# heat map budget
network = random_graph(500, 15)
for total in totals:
for f in [1,total]:
for R in Rs: #numpy.linspace(1.0,4.0,15):
N = network['N']
D = {"N":N,
"frequency":f,
"experiment" : "varying_total",
"type": f"freq={f}/tot={total}/R={R}",
"R":R,
"total": total,
**network,
**infect_params_orig(N,R,average_introductions_per_day = INTRO_DAY, single_intro_by=None, skip_pre=SKIP_PRE),
**run_params(N,frac_tested=f/total,frequency=f, testing_compliance_symptomatic=0),
"initE": INIT_INFECT,
"runTillEnd": INIT_INFECT>0,
}
torun.append(D)
# 28 days
for total in [28]:
for f in [1,7,14,total]:
for G in ["random","gaussian_random_partition"]:
for R in Rs: #numpy.linspace(1.0,4.0,15):
N = 500
if G=='random':
network = random_graph(500, 15)
else:
network = get_network(500,0.5)
D = {"N":N,
"frequency":f,
"experiment" : f"28_days_G_{G}",
"type": f"freq={f}/tot={total}/R={R}",
"R":R,
"total": total,
**network,
**infect_params_orig(N,R,average_introductions_per_day = INTRO_DAY, single_intro_by=None, skip_pre=SKIP_PRE),
**run_params(N,frac_tested=f/total,frequency=f, testing_compliance_symptomatic=0),
"initE": INIT_INFECT,
"runTillEnd": INIT_INFECT>0,
}
torun.append(D)
for total in [28]:
for f in [1,7,14,total]:
for R in Rs: #numpy.linspace(1.0,4.0,15):
N = 500
network= random_graph(500, 15)
D = {"N":N,
"frequency":f,
"experiment" : f"28_days_symptomatic",
"type": f"freq={f}/tot={total}/R={R}",
"R":R,
"total": total,
**network,
**infect_params_orig(N,R,average_introductions_per_day = INTRO_DAY, single_intro_by=None, skip_pre=False,pct_symptomatic=0.5),
**run_params(N,frac_tested=f/total,frequency=f, testing_compliance_symptomatic=0.25),
"initE": INIT_INFECT,
"runTillEnd": INIT_INFECT>0,
}
torun.append(D)
# no testing
for total in totals:
for N in [2000]:
for R in Rs: #numpy.linspace(1.0,4.0,15):
network = random_graph(2000, 15)
D = {"frequency":1,
"experiment" : f"effective_R_{N}_nodes",
"type": f"NoTesting/tot={total}/R={R}",
"R":R,
"total": total,
**network,
**infect_params_orig(N,R,single_intro_by=None,skip_pre=SKIP_PRE),
**run_params(N,frac_tested=0,frequency=1, testing_compliance_symptomatic=0.5),
"initE":INIT_INFECT,
"T": total,
"runnTillEnd": True,
#"intervention_start_pct_infected": 1.1 # don't do any interventions
}
torun.append(D)
# external infection
network = random_graph(500, 15)
for total in [28]:
for rate in [1/14,1/10,1/7,1/3.5,1/2]:
for f in [1,7,28]:# [1]+totals:
if total % f: continue
for R in Rs: #numpy.linspace(1.0,4.0,15):
N = network['N']
D = {"N":N,
"frequency":f,
"experiment" : "external_introductions",
"type": f"freq={f}/tot={total}/R={R}/ext={rate}",
"R":R,
"total": total,
**network,
**infect_params_orig(N,R,average_introductions_per_day = rate, single_intro_by=None, skip_pre=SKIP_PRE),
**run_params(N,frac_tested=f/total,frequency=f, testing_compliance_symptomatic=0),
"initI_asym": 0,
"runTillEnd": True
}
torun.append(D)
realizations = 400
file_prefix = f"all_versions"
file_prefix
# +
totals = [1,2,7,14,28,35,52]
Rs = [1.2, 1.6, 2, 2.4, 2.8,3.2,3.6,4.0,4.4,4.8]
torun = []
SKIP_PRE = True
INIT_INFECT = 1
INTRO_DAY = 0
for total in [28]:
for f in [1,7,14,total]:
for G in ["gaussian_random_partition"]:
for R in Rs: #numpy.linspace(1.0,4.0,15):
N = 500
if G=='random':
network = random_graph(500, 15)
else:
network = get_network(500)
D = {"N":N,
"frequency":f,
"experiment" : f"28_days_G_{G}",
"type": f"freq={f}/tot={total}/R={R}",
"R":R,
"total": total,
**network,
**infect_params_orig(N,R,average_introductions_per_day = INTRO_DAY, single_intro_by=None, skip_pre=SKIP_PRE),
**run_params(N,frac_tested=f/total,frequency=f, testing_compliance_symptomatic=0),
"initE": INIT_INFECT,
"runTillEnd": INIT_INFECT>0,
}
torun.append(D)
realizations = 400
file_prefix = f"all_versions"
file_prefix
# +
# Rs = [2,3.2,4.8]
# torun = []
# INIT_INFECT = 1
# INTRO_DAY = 0
# for total in [28]:
# for f in [1,14,total]:
# G = 'random'
# for pre_symptomatic_period in [3,6.4]:
# for R in Rs: #numpy.linspace(1.0,4.0,15):
# N = 500
# if G=='random':
# network = random_graph(500, 15)
# else:
# network = get_network(500,0.5)
# D = {"N":N,
# "frequency":f,
# "experiment" : f"28_days_G_{G}_pre_{pre_symptomatic_period}",
# "type": f"freq={f}/tot={total}/R={R}",
# "R":R,
# "total": total,
# **network,
# **infect_params_orig(N,R,average_introductions_per_day = INTRO_DAY, single_intro_by=None, skip_pre=False, pre_symptomatic_period=6.4),
# **run_params(N,frac_tested=f/total,frequency=f, testing_compliance_symptomatic=0),
# "initI_asym": INIT_INFECT,
# "runTillEnd": INIT_INFECT>0,
# }
# torun.append(D)
# for total in [28]:
# for f in [1,14,total]:
# G = 'random'
# for test_per in [0.1,0.5]:
# for pre_symptomatic_period in [3]:
# for R in Rs: #numpy.linspace(1.0,4.0,15):
# N = 500
# if G=='random':
# network = random_graph(500, 15)
# else:
# network = get_network(500,0.5)
# D = {"N":N,
# "frequency":f,
# "experiment" : f"28_days_G_{G}_symptomatic_{test_per}_pre_{pre_symptomatic_period}",
# "type": f"freq={f}/tot={total}/R={R}",
# "R":R,
# "total": total,
# **network,
# **infect_params_orig(N,R,average_introductions_per_day = INTRO_DAY, single_intro_by=None, skip_pre=False, pre_symptomatic_period=6.4,pct_symptomatic=0.5),
# **run_params(N,frac_tested=f/total,frequency=f, testing_compliance_symptomatic=test_per),
# "initI_asym": INIT_INFECT,
# "runTillEnd": INIT_INFECT>0,
# }
# torun.append(D)
# realizations = 400
# file_prefix = f"testing"
# file_prefix
# -
len(torun)*realizations
# ## Run several experiments in parallel
# +
os.chdir(PATH)
if not os.path.exists("data/"):
os.makedirs("data")
timestamp = datetime.now().strftime('%Y%m%d_%H_%M_%S')
datadir = f"data/{timestamp}_{realizations}"
os.makedirs(datadir)
torun_fname = datadir+"/torun.pickle"
save_to_file(torun,torun_fname)
print(datadir)
os.chdir("seirsplus")
# -
# %%time
# !python -m seirsplus.parallel_run --torun "../$torun_fname" --realizations $realizations --savename "../$datadir/data"
datadir ='data/20201020_14_12_29_400'
2
datadir
os.chdir(PATH)
data = read_from_file(datadir+"/data")
filename = file_prefix + "_" + datadir.split('/')[1]+".csv"
data.to_csv(filename)
print(filename)
len(data)
data.experiment.unique()
# +
def censor(df):
return df[df['numPositive/sum']>0]
def restrict(df,total,R):
return df[(df.total==total) & (df.R==R)]
def summarize(df, percentile=0.9):
print(f"Total: {df.total.unique()} R: {df.R.unique()} min(numPositive/sum): {df['numPositive/sum'].min()}")
print(f"Frequency: risk(mean),risk({percentile*100:.0f}%) | overall(mean),overall({percentile*100:.0f}%)")
for i in sorted(df.frequency.unique()):
s = df[df.frequency==i]
t=s['numInfectious/average']
u = s['overallInfected/last']
print(f"{i:02d}: {t.mean():.2f}, {t.quantile(percentile):.2f} "+
f"| {u.mean():.2f} , {u.quantile(percentile):.1f} "+
f"| count={t.count()} ")
summarize(censor(restrict(data,total=28,R=2.0)))
# -
def summary(df,fields = ['numInfectious/average','overallInfected/last','time/last'], p=0.9):
def top(x): return x.quantile(p)
print(f"Total: {df.total.unique()} R: {df.R.unique()} min(numPositive/sum): {df['numPositive/sum'].min()}")
D = {f: ['mean',top] for f in fields}
D[fields[0]].insert(0,'count')
return df.fillna(0).groupby('frequency')[fields].agg(D)
summary(censor(restrict(data,total=28,R=2.8)))
total = 28
show_violins(censor[(censor.total==total) & (censor.R==2.8)],
field = "numInfectious/average",
ylabel ="Social risk until 1st detection",
groupby="frequency",
title=f"Overall num of infected when testing 100% per {total} days")
# +
def panels(data, yfield):
daily = data[data.frequency==1]
batch = data[data.frequency==data.total]
panel1 = daily.groupby([yfield,"R"])["overallInfected/last"].mean().unstack()
panel2 = batch.groupby([yfield,"R"])["overallInfected/last"].mean().unstack()
panel3 = panel1/panel2*100
return panel1,panel2,panel3
# +
from matplotlib.colors import ListedColormap
def colormap(minval,midval,maxval, scale = "RdBu_r"):
minval,midval,maxval = int(minval), int(midval), int(maxval)
n = (maxval-minval)*10
L = sns.color_palette(scale, n).as_hex()
L1 = L[0:n//2:n//(2*(midval-minval))]
L2 = L[n//2:n:n//(2*(maxval-midval))]
return ListedColormap(L1+L2)
#greens = list(sns.color_palette("BuGn_d",(midval-minval)//2).as_hex())[::-1]
#L = greens + L1
#cmap_percentages = ListedColormap(L)
#L1 = sns.color_palette("RdBu_r", 2*MAX-10).as_hex()
#greens = list(reversed(sns.color_palette("BuGn_d",10).as_hex()))
#L = greens+L1[0:MAX:(MAX//10)] + L1[MAX:]
#from matplotlib.colors import ListedColormap
#cmap_numbers = ListedColormap(L)
# +
from matplotlib.ticker import StrMethodFormatter
def heatmap(data, yfield, ytitle, maxper = None, tight= False, maxval= 100):
panel1,panel2,panel3 = panels(data,yfield)
print(max(*panel1.values.flatten(),*panel2.values.flatten()))
if maxval is None:
maxval = max(*panel1.values.flatten(),*panel2.values.flatten())
if not maxper:
maxper = max(panel3.values.flatten())
cmap_numbers = colormap(0,10,max(11,maxval))
cmap_percentages = colormap(0,100,max(maxper,101), scale = 'RdYlGn_r')
print(data.total.unique())
size = 30
titlesize = 40
fig , axes = plt.subplots(1,3, sharex=True, sharey= True, figsize= (28,10))
cbar_ax = fig.add_axes([-0.05, 0.15, .03, .7])
#axes[0].title.set_text('28/4 vs baseline')
#cmap = sns.diverging_palette(150, 275, sep=1,s=80, l=55, n=25)
#fig.suptitle(f'Daily vs Batch testing - {statistic} overall infected individuals', fontsize=int(titlesize*1.1), color="blue")
#fig.subplots_adjust(top=1)
fmt = ".1f"
color = "white"
axes[0].set_title("a) Cost with daily testing",loc="left",fontsize=int(titlesize))
sns.heatmap(panel1,ax=axes[0],cbar=True, cbar_ax = cbar_ax , fmt=fmt, vmin=0,vmax= maxval, annot=False,cmap=cmap_numbers)# annot=True,fmt='.0f' ,cmap=cmap) # annot_kws={'color':color}
axes[1].set_title('b) Cost with batch testing', loc='left',fontsize=int(titlesize))
sns.heatmap(panel2,ax=axes[1],cbar=False, vmin=0,vmax= maxval, fmt=fmt, annot= False,cmap=cmap_numbers)# annot=True,fmt='.0f', cmap=cmap)
axes[2].set_title('c) Daily / Batch cost (%)', loc='left',fontsize=int(titlesize))
sns.heatmap(panel3,ax=axes[2],cbar=True, annot= False, fmt=fmt, vmin=0 ,vmax= max(100,maxper), cmap = cmap_percentages)# annot=True,fmt='.0f', cmap=cmap)
#sns.heatmap(percent,ax=axes[2],vmin=0,vmax=125,cbar=True, annot= True, fmt='.0f', cmap=cmap_percentages)# annot=True,fmt='.0f', cmap=cmap)
cbar = axes[0].collections[0].colorbar
cbar.ax.set_ylabel('# of individuals infected at detection',fontsize=size)
cbar.ax.yaxis.set_ticks_position('left')
cbar.ax.yaxis.set_label_position('left')
cbar_ax.tick_params(axis='y',labelsize=size)
cbar_ax = axes[2].collections[0].colorbar.ax
#cbar_ax.set_ylabel('daily cost as % of batch cost',fontsize=20, rotation = 270)
cbar_ax.text(4.5, 0.5, 'daily cost as % of batch cost', rotation=270, fontsize= size,
verticalalignment='center', horizontalalignment='right',
transform=cbar_ax.transAxes)
cbar_ax.tick_params(axis='y',labelsize=size)
from fractions import Fraction
def format_func(value, tick_number):
introductions = sorted(data.average_introductions_per_day.unique())
f = Fraction(introductions[tick_number]).limit_denominator(100)
return fr"$\frac{{{f.numerator}}}{{{f.denominator}}}$"
def format_func2(value,tick_number):
budgets = sorted(data[yfield].unique())
v = budgets[tick_number]
return fr"$\frac{{1}}{{{v}}}$"
for i in range(3):
axes[i].set_ylabel('' if i else ytitle, fontsize=size)
axes[i].set_xlabel('' if i!=1 else 'Reproductive number', fontsize=titlesize)
axes[i].tick_params(axis='x', rotation=45, labelsize= size)
axes[i].tick_params(axis='x', labelsize=size )
axes[i].tick_params(axis='y',labelsize=size )
if yfield == 'average_introductions_per_day':
axes[0].yaxis.set_major_formatter(plt.FuncFormatter(format_func))
else:
axes[0].yaxis.set_major_formatter(plt.FuncFormatter(format_func2))
axes[0].tick_params(axis='y',rotation=0 )
axes[0].tick_params(axis='y',labelsize= titlesize)
#fig.text(0.15,0,'* Dropping runs where all infected individuals recovered before detection', fontsize=12)
cbar = axes[2].collections[0].colorbar
#cbar.set_yticklabels([f"{int(i)}%" for i in cbar.get_ticks()]) # set ticks of your format
#plt.show()
if tight: fig.tight_layout() #rect=[0, 0, .9, 1])
# -
data = data[data.R < 4.1]
heatmap(data[data.experiment=='external_introductions'],'average_introductions_per_day','Mean daily external infections', maxper =200, tight = True)
heatmap(data[(data.experiment=='varying_total') & (data.total>1)],'total','Daily budget (fraction of population)', maxper = 200, tight=True)
data.experiment.unique()
# +
def cost_curves(df,BUDGET=28, FREQS = [1,14,28], title = None, ylim=None, percentile=None):
if title is None:
title = f"Overall infection per policy for budget of testing 100% every {BUDGET} days"
sns.set_style('ticks')
fontsmall = 14
fontlarge = 18
fig = plt.figure(figsize=(12, 6), dpi = 200)
ax = fig.add_subplot(111)
ax.spines["top"].set_visible(False)
ax.spines["right"].set_visible(False)
budget = df[df.total==BUDGET ]
Rs = sorted(budget.R.unique())
colors = sns.color_palette("tab10", len(FREQS))
i = 0
for f in FREQS[::-1]:
temp = budget[budget.frequency==f].groupby('R')['overallInfected/last']
if percentile is None:
cost = [temp.mean().loc[R] for R in Rs]
upper = [temp.quantile(0.8).loc[R] for R in Rs]
lower = [temp.quantile(0.3).loc[R] for R in Rs]
ax.plot(Rs,cost, color = colors[i], label = f"Test every {f} day{'s' if f>1 else ''} ({BUDGET//f} batch{'es' if f<BUDGET else ''})", zorder = 20-2*i)
ax.fill_between(Rs,lower,upper, color = colors[i], alpha = 0.2, zorder = 20-2*i-1)
else:
cost = [temp.quantile(percentile).loc[R] for R in Rs]
ax.plot(Rs,cost, linestyle='--',color = colors[i], label = f"Test every {f} day{'s' if f>1 else ''} ({BUDGET//f} batch{'es' if f<BUDGET else ''})", zorder = 20-2*i)
# {percentile*100:.0f}th percentile
i += 1
handles, labels = ax.get_legend_handles_labels()
ax.set_xticks(Rs)
l = ax.legend([handle for i,handle in enumerate(handles)], [label for i,label in enumerate(labels)], loc = 'upper left', frameon=False, fontsize= fontsmall)
l.set_zorder(50)
if percentile:
title += f' ({percentile*100:.0f}th percentile)'
#ax.set_title(title,fontsize=fontlarge)
ax.set_xlabel("Reproductive number", fontsize= fontsmall)
ax.set_ylabel("Overall number of infected individuals",fontsize= fontsmall)
ax.tick_params(axis='y', labelsize=fontsmall )
ax.tick_params(axis='x', labelsize=fontsmall )
if ylim:
ax.set_ylim(0,ylim)
fig.show()
#fig.text(0.15,0,'* Shaded area from 25th to 75th percentile. Dropping runs where all infected individuals recovered before detection', fontsize=8)
# -
data.experiment.unique()
# +
plots = [ ('28_days_G_random', ''), ('28_days_G_gaussian_random_partition','- random partition model')] # ,('28_days_symptomatic', '(symptomatic testing)')]
for t,text in plots:
df = data[data.experiment==t]
title = f"Outbreak size per policy " + text
cost_curves(df,title=title)
cost_curves(df,title=title, percentile = 0.95)
# -
data.experiment.unique()
df = data[data.experiment=='28_days_G_random']
df.groupby(['R','frequency'])['overallInfected/last'].mean().unstack()
temp = df[df.frequency==1]
df['numI_sym/sum'].sum()
network = random_graph(500, 15)
N = 500
R = 4.4
total = 28
f = 1
D = {"N":N,
"frequency":f,
"experiment" : f"28_days_G_{G}",
"type": f"freq={f}/tot={total}/R={R}",
"R":R,
"total": total,
**network,
**infect_params_orig(N,R,average_introductions_per_day = INTRO_DAY, single_intro_by=None, skip_pre=SKIP_PRE),
**run_params(N,frac_tested=f/total,frequency=f, testing_compliance_symptomatic=0),
"initE": INIT_INFECT,
"runTillEnd": INIT_INFECT>0,
}
hist, sum, m = run(D,True)
sum['numI_asym/sum']
hist['numI_asym']
data.set_index('time')['numTested'].fillna(0).plot()
data.set_index('time')['numPositive'].fillna(0).plot()
L = []
for n in range(m.numNodes):
istart = 0
for t in range(m.tidx+1):
if not istart and (m.Xseries[t,n]==m.I_sym):
istart = m.tseries[t]
if istart and (m.Xseries[t,n]==m.R):
L.append(m.tseries[t]-istart)
break
plt.hist(L,bins=range(13),density=True)
m.tidx
sum(L)/len(L)
len(L)
symptomaticPeriod_mean, symptomaticPeriod_coeffvar = 4.0, 0.4
GAMMA = gamma_dist(symptomaticPeriod_mean, symptomaticPeriod_coeffvar, 3400)
sum(GAMMA)/len(GAMMA)
plt.hist(list(GAMMA),bins=15,density=True)
| Varying_params.ipynb |
# # + 
# # **First Notebook: Virtual machine test and assignment submission**
# #### This notebook will test that the virtual machine (VM) is functioning properly and will show you how to submit an assignment to the autograder. To move through the notebook just run each of the cells. You will not need to solve any problems to complete this lab. You can run a cell by pressing "shift-enter", which will compute the current cell and advance to the next cell, or by clicking in a cell and pressing "control-enter", which will compute the current cell and remain in that cell. At the end of the notebook you will export / download the notebook and submit it to the autograder.
# #### ** This notebook covers: **
# #### *Part 1:* Test Spark functionality
# #### *Part 2:* Check class testing library
# #### *Part 3:* Check plotting
# #### *Part 4:* Check MathJax formulas
# #### *Part 5:* Export / download and submit
# ### ** Part 1: Test Spark functionality **
# #### ** (1a) Parallelize, filter, and reduce **
# +
# Check that Spark is working
largeRange = sc.parallelize(xrange(100000))
reduceTest = largeRange.reduce(lambda a, b: a + b)
filterReduceTest = largeRange.filter(lambda x: x % 7 == 0).sum()
print reduceTest
print filterReduceTest
# If the Spark jobs don't work properly these will raise an AssertionError
assert reduceTest == 4999950000
assert filterReduceTest == 714264285
# -
# #### ** (1b) Loading a text file **
# +
# Check loading data with sc.textFile
import os.path
baseDir = os.path.join('data')
inputPath = os.path.join('cs100', 'lab1', 'shakespeare.txt')
fileName = os.path.join(baseDir, inputPath)
rawData = sc.textFile(fileName)
shakespeareCount = rawData.count()
print shakespeareCount
# If the text file didn't load properly an AssertionError will be raised
assert shakespeareCount == 122395
# -
# ### ** Part 2: Check class testing library **
# #### ** (2a) Compare with hash **
# +
# TEST Compare with hash (2a)
# Check our testing library/package
# This should print '1 test passed.' on two lines
from test_helper import Test
twelve = 12
Test.assertEquals(twelve, 12, 'twelve should equal 12')
Test.assertEqualsHashed(twelve, '7b52009b64fd0a2a49e6d8a939753077792b0554',
'twelve, once hashed, should equal the hashed value of 12')
# -
# #### ** (2b) Compare lists **
# TEST Compare lists (2b)
# This should print '1 test passed.'
unsortedList = [(5, 'b'), (5, 'a'), (4, 'c'), (3, 'a')]
Test.assertEquals(sorted(unsortedList), [(3, 'a'), (4, 'c'), (5, 'a'), (5, 'b')],
'unsortedList does not sort properly')
# ### ** Part 3: Check plotting **
# #### ** (3a) Our first plot **
# #### After executing the code cell below, you should see a plot with 50 blue circles. The circles should start at the bottom left and end at the top right.
# +
# Check matplotlib plotting
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from math import log
# function for generating plot layout
def preparePlot(xticks, yticks, figsize=(10.5, 6), hideLabels=False, gridColor='#999999', gridWidth=1.0):
plt.close()
fig, ax = plt.subplots(figsize=figsize, facecolor='white', edgecolor='white')
ax.axes.tick_params(labelcolor='#999999', labelsize='10')
for axis, ticks in [(ax.get_xaxis(), xticks), (ax.get_yaxis(), yticks)]:
axis.set_ticks_position('none')
axis.set_ticks(ticks)
axis.label.set_color('#999999')
if hideLabels: axis.set_ticklabels([])
plt.grid(color=gridColor, linewidth=gridWidth, linestyle='-')
map(lambda position: ax.spines[position].set_visible(False), ['bottom', 'top', 'left', 'right'])
return fig, ax
# generate layout and plot data
x = range(1, 50)
y = [log(x1 ** 2) for x1 in x]
fig, ax = preparePlot(range(5, 60, 10), range(0, 12, 1))
plt.scatter(x, y, s=14**2, c='#d6ebf2', edgecolors='#8cbfd0', alpha=0.75)
ax.set_xlabel(r'$range(1, 50)$'), ax.set_ylabel(r'$\log_e(x^2)$')
pass
# -
# ### ** Part 4: Check MathJax Formulas **
# #### ** (4a) Gradient descent formula **
# #### You should see a formula on the line below this one: $$ \scriptsize \mathbf{w}_{i+1} = \mathbf{w}_i - \alpha_i \sum_j (\mathbf{w}_i^\top\mathbf{x}_j - y_j) \mathbf{x}_j \,.$$
#
# #### This formula is included inline with the text and is $ \scriptsize (\mathbf{w}^\top \mathbf{x} - y) \mathbf{x} $.
# #### ** (4b) Log loss formula **
# #### This formula shows log loss for single point. Log loss is defined as: $$ \begin{align} \scriptsize \ell_{log}(p, y) = \begin{cases} -\log (p) & \text{if } y = 1 \\\ -\log(1-p) & \text{if } y = 0 \end{cases} \end{align} $$
# ### ** Part 5: Export / download and submit **
# #### ** (5a) Time to submit **
# #### You have completed the lab. To submit the lab for grading you will need to download it from your IPython Notebook environment. You can do this by clicking on "File", then hovering your mouse over "Download as", and then clicking on "Python (.py)". This will export your IPython Notebook as a .py file to your computer.
# #### To upload this file to the course autograder, go to the edX website and find the page for submitting this assignment. Click "Choose file", then navigate to and click on the downloaded .py file. Now click the "Open" button and then the "Check" button. Your submission will be graded shortly and will be available on the page where you submitted. Note that when submission volumes are high, it may take as long as an hour to receive results.
| spark/lab0_student.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/emenriquez/python-challenge-exercises/blob/master/Persistent_Bugger_(6).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="cza5bAAoO4I9" colab_type="text"
# # Persistent Bugger (Rank 6)
#
# Write a function, `persistence`, that takes in a positive parameter `num` and returns its multiplicative persistence, which is the number of times you must multiply the digits in `num` until you reach a single digit.
#
# For example:
# ```
# persistence(39) => 3 # Because 3*9 = 27, 2*7 = 14, 1*4=4
# # and 4 has only one digit.
#
# persistence(999) => 4 # Because 9*9*9 = 729, 7*2*9 = 126,
# # 1*2*6 = 12, and finally 1*2 = 2.
#
# persistence(4) => 0 # Because 4 is already a one-digit number.
# ```
# + id="QpDrZYuPLYE2" colab_type="code" colab={}
def persistence(n):
# start with a counter value of zero, and count how many times we can iterate between results until only 1 digit is left
counter = 0
while len(str(n)) >1:
# separate digits into list items
separate_chars = [int(x) for x in str(n)]
result = 1
# iterate through list and multiply all list items together
for num in separate_chars:
result *= num
# update our number to our new product
n = result
# increase the counter to reflect 1 iteration through our loop
counter += 1
return counter
# + [markdown] id="IkKF1LRvOwnY" colab_type="text"
# ### Test Example
# + id="lNZFkxPtNK4Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="02d85236-3e18-4cbd-aad0-3b5409fee788"
persistence(39)
# + [markdown] id="-0VzLbWDPWeR" colab_type="text"
# Completed in 13 minutes
| Persistent_Bugger_(6).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Train multiple-steps model
#
# This notebook contains different examples of how to train different models based on a multiple-steps loss using the function ``` modules.full_pipeline_multiple_steps ```
# %load_ext autoreload
# %autoreload 2
# +
import sys
import time
sys.path.append('/'.join(sys.path[0].split('/')[:-1]))
from modules.full_pipeline_multiple_steps import main
# -
train_loss_ev, val_loss_ev, train_loss_steps_ev, test_loss_steps_ev, weight_variations_ev = \
main('config_residual_multiple_steps.json', load_model=False)
import matplotlib.pyplot as plt
from matplotlib import cm
import numpy as np
# +
len_epoch = len(train_loss_steps_ev[0]['t0'])
x_iter = np.arange(len_epoch)
plt.figure(figsize=(20,8))
colors = cm.Spectral(np.linspace(0,1,10))
for ep in range(1,len(train_loss_steps_ev)):
for i in range(8):
x_iter = np.arange(len(train_loss_steps_ev[ep]['t{}'.format(i)]))
x_iter_ep = x_iter + ep*len_epoch
plt.plot(x_iter_ep, train_loss_steps_ev[ep]['t{}'.format(i)], c=colors[i])
for (w, _, it) in weight_variations_ev[ep]:
plt.vlines(it + ep*len_epoch, 0.01, 0.09)
# -
for w in weight_variations_ev:
for new_w in w:
print('Epoch {} - Iteration {}'.format(new_w[1], new_w[2]))
print('WEIGHT VECTOR: ', new_w[0])
# ## Idem but with long connections architecture
# +
import sys
import time
sys.path.append('/'.join(sys.path[0].split('/')[:-1]))
from modules.full_pipeline_multiple_steps import main
# -
{
"directories":
{
"datadir": "../data/healpix/",
"input_dir": "5.625deg_nearest/",
"model_save_path": "models/",
"pred_save_path": "predictions/",
"obs_file_name": "observations_nearest.nc",
"rmse_weyn_name": "metrics/rmses_weyn.nc",
"constants": "constants/constants_5.625deg_standardized.nc",
"train_mean_file": "mean_train_features_dynamic.nc",
"train_std_file": "std_train_features_dynamic.nc",
"metrics_path": "metrics/",
"standardized_data": "None"
},
"training_constants":
{
"chunk_size": 521,
"train_years": ["1990","2012"],
"val_years": ["2013", "2016"],
"test_years": ["2017", "2018"],
"nodes":3072,
"max_lead_time": 120,
"nb_timesteps": 2,
"nb_epochs": 14,
"learning_rate": 0.008,
"batch_size": 10
},
"model_parameters":
{
"len_sqce": 2,
"delta_t": 6,
"in_features": 7,
"out_features":2,
"num_steps_ahead": 8,
"architecture_name": "loss_v0_8steps_increas_reinitialize_residual_l3_long_connections_per_epoch",
"resolution": 5.625,
"kernel_size_pooling": 4,
"initial_weights": [1, 0, 0, 0, 0, 0, 0, 0],
"model": "UNetSphericalHealpixResidualLongConnections"
}
}
train_loss_ev, val_loss_ev, train_loss_steps_ev, test_loss_steps_ev, weight_variations_ev = \
main('config_residual_multiple_steps.json', load_model=False)
for w in weight_variations_ev:
for new_w in w:
print('Epoch {} - Iteration {}'.format(new_w[1], new_w[2]))
print('WEIGHT VECTOR: ', new_w[0])
import json
import numpy as np
# +
best_epoch = np.argmin(val_loss_ev)
min_error = val_loss_ev[best_epoch]
from modules.mail import send_info_mail
# +
train_loss_json = '../data/healpix/models/train_loss_' + '8steps_increas_reinitialize_residual_l3_long_connections_per_epoch' + '.json'
with open(train_loss_json, "w") as outfile:
json.dump(train_loss_steps_ev, outfile)
test_loss_json = '../data/healpix/models/test_loss_' + '8steps_increas_reinitialize_residual_l3_long_connections_per_epoch' + '.json'
with open(test_loss_json, "w") as outfile:
json.dump(test_loss_steps_ev, outfile)
with open("../modules/confMail.json") as json_data_file:
mailConf = json.load(json_data_file)
mail = {
"sender": mailConf["sender"],
"receiver": mailConf["sender"],
"subject": "Finished training!",
"body": "Finished training model {}.\n Min loss {} at epoch {} " \
.format(description, min_error, best_epoch),
"fileAttaching": "Yes",
"file": ["../configs/" + 'config_residual_multiple_steps.json', train_loss_json, test_loss_json]
}
send_info_mail(mailInfo=mail, configFile=mailConf)
# +
train_loss_json = '../data/healpix/models/train_loss_' + '8steps_increas_reinitialize_residual_l3_long_connections_per_epoch' + '.json'
with open(train_loss_json, "w") as outfile:
json.dump(train_loss_steps_ev, outfile)
test_loss_json = '../data/healpix/models/test_loss_' + '8steps_increas_reinitialize_residual_l3_long_connections_per_epoch' + '.json'
with open(test_loss_json, "w") as outfile:
json.dump(test_loss_steps_ev, outfile)
# -
description = 'all_const_len2_delta_6_architecture_loss_v0_8steps_increas_reinitialize_residual_l3_long_connections_per_epoch'
import matplotlib.pyplot as plt
from matplotlib import cm
import numpy as np
# +
len_epoch = len(train_loss_steps_ev[0]['t0'])
x_iter = np.arange(len_epoch)
plt.figure(figsize=(20,8))
colors = cm.Spectral(np.linspace(0,1,10))
for ep in range(1,len(train_loss_steps_ev)):
for i in range(8):
x_iter = np.arange(len(train_loss_steps_ev[ep]['t{}'.format(i)]))
x_iter_ep = x_iter + ep*len_epoch
plt.plot(x_iter_ep, train_loss_steps_ev[ep]['t{}'.format(i)], c=colors[i])
for (w, _, it) in weight_variations_ev[ep]:
plt.vlines(it + ep*len_epoch, 0.01, 0.09)
# -
| notebooks/Restarting_weights_per_epoch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1. Introduction
#
# The sinking of the RMS Titanic is one of the most infamous shipwrecks in history. On April 15, 1912, during her maiden voyage, the Titanic sank after colliding with an iceberg, killing 1502 out of 2224 passengers and crew. This sensational tragedy shocked the international community and led to better safety regulations for ships.
#
# One of the reasons that the shipwreck led to such loss of life was that there were not enough lifeboats for the passengers and crew. Although there was some element of luck involved in surviving the sinking, some groups of people were more likely to survive than others, such as women, children, and the upper-class.
#
# The complete the analysis of what sorts of people were likely to survive and apply the tools of machine learning to predict which passengers survived the tragedy.
#
# - Defining the problem statement
# - Collecting the data
# - Data analysis
# - Data Visualization
# - Feature engineering
# - Modelling
# - Testing
#
#
# #### Goal
#
# The goal of this project to predict if a passenger survived the sinking of the Titanic or not.
# For each in the test set, you must predict a 0 or 1 value for the variable.
# +
# import required libraries
# pandas for data loading and analysis
#numpy for mathematical operations
#for plotting matplolib and seaborn
#for modeling, evalution and prediction scikit-learn
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
# -
# ## 2. Dataset
#
# The dataset can be acquired from this link https://www.kaggle.com/c/titanic, since i downloaded it already, i'm going to load it from local PC
# +
#load dataset
df_train = pd.read_csv('train.csv')
df_test = pd.read_csv('test.csv')
df = df_train.append(df_test, ignore_index = True)
PassengerIDs = df_test['PassengerId']
# -
# #### Data Dictionary
# - Survived: 0 = No, 1 = Yes
# - pclass: Ticket class 1 = 1st, 2 = 2nd, 3 = 3rd
# - sibsp: # of siblings / spouses aboard the Titanic
# - parch: # of parents / children aboard the Titanic
# - ticket: Ticket number
# - cabin: Cabin number
# - embarked: Port of Embarkation C = Cherbourg, Q = Queenstown, S = Southampton
#
#
#
# ## 3. Data Analysis
# head of data
df.head()
# description of data
df.describe()
# The statistics description of data above of numeric values of features telling the count, mean, standard deviation, minimum and maximum number of frequency of a feature. And it can be also seen that the Age and Survived count is less than as compared to other features.
df_train.Survived.value_counts()
# From counts of Survived people data, it seems like that in our training set the people died in disaster are more
# than the people survived
df.Sex.value_counts()
# it looks like that there are more male passengers as compared to female passengers
# shape of training set consist of 891 rows and 12 columns
df_train.shape
# shape of test set consist of 418 rows and 11 columns, it is goal to predict the survival of people so that's why
# the survival feature is not included in test set
df_test.shape
# the shape of merged data of both train and test sets contains 1309 rows and 12 columns
df.shape
df.info()
# As seen the information of the dataset we've total 12 features including target feature 'Survived', the feature have total entries of 1309 but there are some feature with null or NaN values, if not handled they can become problem, but before handling the data. Let's understand the data.
# ## 4. Data Visualization
# we have seen that more people died than survived so let's visualize
plt.figure(figsize = (10,8))
sns.kdeplot(df['Age'][df.Survived == 1], shade = True, color = 'r')
sns.kdeplot(df['Age'][df.Survived == 0], shade = True)
plt.legend(['Survived', 'Died'])
plt.title('Density Plot of Age for Surviving Population and Deceased Population')
plt.show()
def barplot(x, y, title):
sns.barplot(x = x, y = y)
plt.title(title)
barplot(df['Sex'], df['Survived'], 'Bar plot for Survival with respect to gender')
# The chart describes that women are more likely to survive that men
barplot(df['Embarked'], df['Survived'], 'Bar plot for Survival rate with respect to Port of Embarkation')
# - The Chart confirms a person aboarded from C slightly more likely survived
# - The Chart confirms a person aboarded from Q more likely dead
# - The Chart confirms a person aboarded from S more likely dead
barplot(df['Pclass'], df['Survived'], 'Bar plot for Survival rate with respect to passenger class')
# - The Chart confirms 1st class more likely survivied than other classes
# - The Chart confirms 3rd class more likely dead than other classes
# ## 5. Feature Engineering
#
# Feature engineering is the process of using domain knowledge of the data
# to create features (**feature vectors**) that make machine learning algorithms work.
#
# feature vector is an n-dimensional vector of numerical features that represent some object.
# Many algorithms in machine learning require a numerical representation of objects,
# since such representations facilitate processing and statistical analysis.
#
# We are not going to fill Survived feature values because that is our target to correctly predict missing values
df_train.info()
df_test.info()
# #### Feature Engineering Training Data
# +
# handling missing values in training set
df_train.Age = df_train.Age.fillna(df.Age.median())
df_train.Cabin = df_train.Cabin.fillna('U')
most_embarked = df.Embarked.value_counts().index[0]
df_train.Embarked = df_train.Embarked.fillna(most_embarked)
# -
# checking null values in training set
df_train.isnull().sum()
# +
#normalizing the name columns into a new title column
df_train['Title'] = df_train.Name.apply(lambda name: name.split(',')[1].split('.')[0].strip())
normalized_titles = {
"Capt": "Officer",
"Col": "Officer",
"Major": "Officer",
"Jonkheer": "Royalty",
"Don": "Royalty",
"Sir" : "Royalty",
"Dr": "Officer",
"Rev": "Officer",
"the Countess":"Royalty",
"Dona": "Royalty",
"Mme": "Mrs",
"Mlle": "Miss",
"Ms": "Mrs",
"Mr" : "Mr",
"Mrs" : "Mrs",
"Miss" : "Miss",
"Master" : "Master",
"Lady" : "Royalty"
}
df_train.Title = df_train.Title.map(normalized_titles)
df_train.Title.value_counts()
# -
df_train.Sex = df_train.Sex.map({'female': 0, 'male': 1})
df_train['FamilySize'] = df_train['SibSp'] + df_train['Parch'] + 1
df_train.Embarked = df_train.Embarked.map({'S': 0, 'C': 1, 'Q': 2})
df_train.Title = df_train.Title.map({'Mr': 0, 'Miss': 1, 'Mrs': 2, 'Master': 3, 'Officer': 4, 'Royalty': 5})
# #### Feature Engineering Test Data
# handling missing values in test set
df_test.Age = df_test.Age.fillna(df.Age.median())
df_test.Cabin = df_test.Cabin.fillna('U')
df_test.Fare = df_test.Fare.fillna(df.Fare.median())
df_test.isnull().sum()
# +
df_test['Title'] = df_test.Name.apply(lambda name: name.split(',')[1].split('.')[0].strip())
normalized_titles = {
"Capt": "Officer",
"Col": "Officer",
"Major": "Officer",
"Jonkheer": "Royalty",
"Don": "Royalty",
"Sir" : "Royalty",
"Dr": "Officer",
"Rev": "Officer",
"the Countess":"Royalty",
"Dona": "Royalty",
"Mme": "Mrs",
"Mlle": "Miss",
"Ms": "Mrs",
"Mr" : "Mr",
"Mrs" : "Mrs",
"Miss" : "Miss",
"Master" : "Master",
"Lady" : "Royalty"
}
df_test.Title = df_test.Title.map(normalized_titles)
df_test.Title.value_counts()
# -
df_test.Sex = df_test.Sex.map({'female': 0, 'male': 1})
df_test['FamilySize'] = df_test['SibSp'] + df_test['Parch'] + 1
df_test.Embarked = df_test.Embarked.map({'S': 0, 'C': 1, 'Q': 2})
df_test.Title = df_test.Title.map({'Mr': 0, 'Miss': 1, 'Mrs': 2, 'Master': 3, 'Officer': 4, 'Royalty': 5})
target = df_train['Survived']
train_data = df_train.drop(['PassengerId', 'Survived', 'Name', 'SibSp', 'Parch', 'Cabin', 'Ticket'], axis = 1)
train_data.head()
df_test = df_test.drop(['PassengerId', 'Name', 'SibSp', 'Ticket', 'Cabin', 'Parch'], axis = 1)
df_test.head()
# ## 6. Modeling
k_fold = KFold(n_splits=10, shuffle=True, random_state=0)
clf = KNeighborsClassifier(n_neighbors = 13)
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
round(np.mean(score)*100, 2)
clf = DecisionTreeClassifier(min_samples_leaf = 20, max_leaf_nodes = 7)
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
round(np.mean(score)*100, 2)
Rclf = RandomForestClassifier(n_estimators = 100, max_depth = 11)
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
round(np.mean(score)*100, 2)
clf = GradientBoostingClassifier(n_estimators = 60)
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
round(np.mean(score)*100, 2)
# ## 7. Testing
Rclf.fit(train_data, target)
prediction = Rclf.predict(df_test)
# For submitting the project result to www.kaggle.com
# +
submission = pd.DataFrame({
"PassengerId": PassengerIDs,
"Survived": prediction
})
submission.to_csv('submission.csv', index=False)
# -
submission = pd.read_csv('submission.csv')
submission.head()
| Machine Learning From Disaster (Classification)/Titanic Dataset Prediction (Classification).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Learning to rank
# We want to learn a function that can rank order a set of items.
# * Suppose we have a query $q$ and a set of documents $d^1,\ldots,d^m$ that might be relevant to $q$ (e.g. all documents that contain the string $q$).
# * We would like to sort these documents in decreasing order of relevance and show the top $k$ to the user.
#
# A standard way to measure the relevance of a document $d$ to a query $q$ is to use a probabilistic language model based on a bag of words model: <br>
# We define $\mathrm{sim}(q, d)\triangleq p(q|d) = \prod_{i=1}^n p(q_i|d)$, where $q_i$ is the $i$th word or term, and $p(q_i|d)$ is a multinoulli distribution estimated from document $d$.
#
# In practice, we need to smooth the estimated distribution, for example using a Dirichlet prior, representing the overall frequency of each word. This can be estimated from all documents in the system.
#
# $$
# p(t|d) = (1-\lambda)\frac{\text{TF}(t,d)}{\text{LEN}(d)} + \lambda p(t|\text{background})
# $$
#
# where $\text{TF}(t,d)$ is the frequency of term $t$ in document $d$, $\text{LEN}(d)$ is the number of words in $d$, and $0<\lambda<1$ is a smoothing parameter.
| murphy-book/chapter09/Learning-to-rank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Programming Exercise 6: Support Vector Machines
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import scipy.io #Used to load the OCTAVE *.mat files
from sklearn import svm #SVM software
import re #regular expression for e-mail processing
# This is one possible porter stemmer
# (note: I had to do a pip install stemming)
# https://pypi.python.org/pypi/stemming/1.0
from stemming.porter2 import stem
# This porter stemmer seems to more accurately duplicate the
# porter stemmer used in the OCTAVE assignment code
# (note: I had to do a pip install nltk)
# I'll note that both stemmers have very similar results
import nltk, nltk.stem.porter
# -
# ### 2 Spam Classification
# #### 2.1 Preprocessing Emails
print "emailSample1.txt:"
# !cat data/emailSample1.txt
def preProcess( email ):
"""
Function to do some pre processing (simplification of e-mails).
Comments throughout implementation describe what it does.
Input = raw e-mail
Output = processed (simplified) email
"""
# Make the entire e-mail lower case
email = email.lower()
# Strip html tags (strings that look like <blah> where 'blah' does not
# contain '<' or '>')... replace with a space
email = re.sub('<[^<>]+>', ' ', email);
#Any numbers get replaced with the string 'number'
email = re.sub('[0-9]+', 'number', email)
#Anything starting with http or https:// replaced with 'httpaddr'
email = re.sub('(http|https)://[^\s]*', 'httpaddr', email)
#Strings with "@" in the middle are considered emails --> 'emailaddr'
email = re.sub('[^\s]+@[^\s]+', 'emailaddr', email);
#The '$' sign gets replaced with 'dollar'
email = re.sub('[$]+', 'dollar', email);
return email
def email2TokenList( raw_email ):
"""
Function that takes in preprocessed (simplified) email, tokenizes it,
stems each word, and returns an (ordered) list of tokens in the e-mail
"""
# I'll use the NLTK stemmer because it more accurately duplicates the
# performance of the OCTAVE implementation in the assignment
stemmer = nltk.stem.porter.PorterStemmer()
email = preProcess( raw_email )
#Split the e-mail into individual words (tokens) (split by the delimiter ' ')
#but also split by delimiters '@', '$', '/', etc etc
#Splitting by many delimiters is easiest with re.split()
tokens = re.split('[ \@\$\/\#\.\-\:\&\*\+\=\[\]\?\!\(\)\{\}\,\'\"\>\_\<\;\%]', email)
#Loop over each word (token) and use a stemmer to shorten it,
#then check if the word is in the vocab_list... if it is,
#store what index in the vocab_list the word is
tokenlist = []
for token in tokens:
#Remove any non alphanumeric characters
token = re.sub('[^a-zA-Z0-9]', '', token);
#Use the Porter stemmer to stem the word
stemmed = stemmer.stem( token )
#Throw out empty tokens
if not len(token): continue
#Store a list of all unique stemmed words
tokenlist.append(stemmed)
return tokenlist
# ##### 2.1.1 Vocabulary List
def getVocabDict(reverse=False):
"""
Function to read in the supplied vocab list text file into a dictionary.
I'll use this for now, but since I'm using a slightly different stemmer,
I'd like to generate this list myself from some sort of data set...
Dictionary key is the stemmed word, value is the index in the text file
If "reverse", the keys and values are switched.
"""
vocab_dict = {}
with open("data/vocab.txt") as f:
for line in f:
(val, key) = line.split()
if not reverse:
vocab_dict[key] = int(val)
else:
vocab_dict[int(val)] = key
return vocab_dict
def email2VocabIndices( raw_email, vocab_dict ):
"""
Function that takes in a raw email and returns a list of indices corresponding
to the location in vocab_dict for each stemmed word in the email.
"""
tokenlist = email2TokenList( raw_email )
index_list = [ vocab_dict[token] for token in tokenlist if token in vocab_dict ]
return index_list
# #### 2.2 Extracting Features from Emails
def email2FeatureVector( raw_email, vocab_dict ):
"""
Function that takes as input a raw email, and returns a vector of shape
(n,1) where n is the size of the vocab_dict.
The first element in this vector is 1 if the vocab word with index == 1
is in the raw_email, 0 otherwise.
"""
n = len(vocab_dict)
result = np.zeros((n,1))
vocab_indices = email2VocabIndices( email_contents, vocab_dict )
for idx in vocab_indices:
result[idx] = 1
return result
# +
# " ... run your code on the email sample. You should see that the feature vector
# has length 1899 and 45 non-zero entries."
vocab_dict = getVocabDict()
email_contents = open( 'data/emailSample1.txt', 'r' ).read()
test_fv = email2FeatureVector( email_contents, vocab_dict )
print "Length of feature vector is %d" % len(test_fv)
print "Number of non-zero entries is: %d" % sum(test_fv==1)
# -
# #### 2.3 Training SVM for Spam Classification
# +
# Read in the training set and test set provided
# Note the feature vectors correspond to the stemming implementation
# done in the OCTAVE code... which may be different than mine.
# Training set
datafile = 'data/spamTrain.mat'
mat = scipy.io.loadmat( datafile )
X, y = mat['X'], mat['y']
#NOT inserting a column of 1's in case SVM software does it for me automatically...
#X = np.insert(X ,0,1,axis=1)
# Test set
datafile = 'data/spamTest.mat'
mat = scipy.io.loadmat( datafile )
Xtest, ytest = mat['Xtest'], mat['ytest']
# -
pos = np.array([X[i] for i in xrange(X.shape[0]) if y[i] == 1])
neg = np.array([X[i] for i in xrange(X.shape[0]) if y[i] == 0])
print 'Total number of training emails = ',X.shape[0]
print 'Number of training spam emails = ',pos.shape[0]
print 'Number of training nonspam emails = ',neg.shape[0]
# +
# Run the SVM training (with C = 0.1) using SVM software.
# First we make an instance of an SVM with C=0.1 and 'linear' kernel
linear_svm = svm.SVC(C=0.1, kernel='linear')
# Now we fit the SVM to our X matrix, given the labels y
linear_svm.fit( X, y.flatten() )
# +
# "Once the training completes, you should see that the classifier gets a
# training accuracy of about 99.8% and a test accuracy of about 98.5%"
train_predictions = linear_svm.predict(X).reshape((y.shape[0],1))
train_acc = 100. * float(sum(train_predictions == y))/y.shape[0]
print 'Training accuracy = %0.2f%%' % train_acc
test_predictions = linear_svm.predict(Xtest).reshape((ytest.shape[0],1))
test_acc = 100. * float(sum(test_predictions == ytest))/ytest.shape[0]
print 'Test set accuracy = %0.2f%%' % test_acc
# -
# #### 2.4 Top Predictors for Spam
# +
# Determine the words most likely to indicate an e-mail is a spam
# From the trained SVM we can get a list of the weight coefficients for each
# word (technically, each word index)
vocab_dict_flipped = getVocabDict(reverse=True)
#Sort indicies from most important to least-important (high to low weight)
sorted_indices = np.argsort( linear_svm.coef_, axis=None )[::-1]
print "The 15 most important words to classify a spam e-mail are:"
print [ vocab_dict_flipped[x] for x in sorted_indices[:15] ]
print
print "The 15 least important words to classify a spam e-mail are:"
print [ vocab_dict_flipped[x] for x in sorted_indices[-15:] ]
print
# Most common word (mostly to debug):
most_common_word = vocab_dict_flipped[sorted_indices[0]]
print '# of spam containing \"%s\" = %d/%d = %0.2f%%'% \
(most_common_word, sum(pos[:,1190]),pos.shape[0], \
100.*float(sum(pos[:,1190]))/pos.shape[0])
print '# of NON spam containing \"%s\" = %d/%d = %0.2f%%'% \
(most_common_word, sum(neg[:,1190]),neg.shape[0], \
100.*float(sum(neg[:,1190]))/neg.shape[0])
# +
# Note my SVM gets some different predictor words for spam than shown in the
# assignment PDF... I've done debugging and I'm confident it's due to a different
# SVM software package, not because of a bug or something in my code.
# Also note the optional exercises "Try your own emails" and "Build your own
# dataset" I will be doing seperately in a blog post... Check out
# blog.davidkaleko.com/svm-email-filter-implementation.html to have a look!
| machine-learning/Exercise 6 - Support Vector Machines/ex6_spam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .ps1
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .NET (PowerShell)
# language: PowerShell
# name: .net-powershell
# ---
Install-Module pskit -force
# +
function ConvertTo-MarkdownTable {
param($targetData)
$names = $targetData[0].psobject.Properties.name
$all = @()
1..$names.count | foreach {
if($_ -eq $names.count) {
$all += '|'
} else {
$all += '|---'
}
}
$result = foreach($record in $targetData) {
$inner=@()
foreach($name in $names) {
$inner+=$record.$name
}
'|' + ($inner -join '|') + '|' + "`n"
}
@"
$('|' + ($names -join '|') + '|')
$($all)
$($result)
"@ | ConvertFrom-Markdown | % html | Get-HtmlContent | Out-Display
}
# +
$date=(Get-Date).AddDays(-1).ToString('MM-dd-yyyy')
$url = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/{0}.csv' -f $date
$data = ConvertFrom-Csv (irm $url)
$table = New-LookupTable $data Country/Region
$result = foreach ($key in $table.keys) {
[pscustomobject][ordered]@{
Key = $key
TotalConfirmed=($table.$key| measure confirmed -Sum).Sum
}
}
# +
$data = ($result | sort-object TotalConfirmed -Descending | select -First 10 -skip 3)
ConvertTo-MarkdownTable $data
$ConfirmedSeries = [Graph.Bar]@{
name = "TotalConfirmed"
x = $data.Key
y = $data.TotalConfirmed
}
$chart = $ConfirmedSeries | New-PlotlyChart -Title "Total Confirmed - COVID-19"
Out-Display $chart
| COVID-19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="yl5YDWvEeu4a" colab_type="text"
# The purpose is to use Dropout in VAE in order to generate data.
#
# Origin VAE uses latent values to generate the data.
# However, with Dropout VAE, we use the mean of the latents to generate the data.
#
#
# The dataset used in notebook: [Customer Support](https://www.ibm.com/communities/analytics/watson-analytics-blog/guide-to-sample-datasets/)
#
# More example about VAE: [Modeling Telecom customer churn](https://towardsdatascience.com/modeling-telecom-customer-churn-with-variational-autoencoder-4e5cf6194871)
#
# Origin code for VAE in keras: [Building Autoencoders in keras](https://blog.keras.io/building-autoencoders-in-keras.html)
# + [markdown] id="fdT9BTrmT3ro" colab_type="text"
# # Import Libraries
# + id="CQNUpL6YT32X" colab_type="code" outputId="eaa36e85-ecc3-4860-8638-45e7b9457d0d" colab={"base_uri": "https://localhost:8080/", "height": 34}
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
import random
import argparse
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder
from tensorflow import set_random_seed
from keras import regularizers
from keras import backend as K
from keras.models import Model
from keras.utils import plot_model
from keras.losses import mse, binary_crossentropy
from keras.layers import Lambda, Input, Dense, Dropout
set_random_seed(1)
np.random.seed(1)
random.seed(1)
# + [markdown] id="sNKVLBOBgR3W" colab_type="text"
# # Path to dataset
# + id="u4QUbkrWUl2L" colab_type="code" colab={}
# Access to resources
path = '/Dataset/WA_Fn-UseC_-Telco-Customer-Churn.csv'
# + id="C8zWTy_ePGg7" colab_type="code" colab={}
intermediate_dim = 256
# + [markdown] id="8VTGWwfObeoT" colab_type="text"
# ## Read Data
# + id="DfcE15dwhyaL" colab_type="code" outputId="1cd188b0-a4be-4a28-dbec-1e0f1d83bd5e" colab={"base_uri": "https://localhost:8080/", "height": 595}
import pandas as pd
na_values = {'?', ' '}
df = pd.read_csv(path,
sep=',',
na_filter=True,
verbose=False,
skip_blank_lines=True,
na_values=na_values,
keep_default_na=False)
df.fillna(method='ffill', inplace=True)
df.dropna(axis=1, how='any', inplace=True)
df.drop(['customerID'], axis=1, inplace=True)
df = df.reset_index(drop=True)
print(df.info())
print(df.head())
# + [markdown] id="ptmiXDEjbhmp" colab_type="text"
# ## Recognizing Categorical data
# + id="pfL6yO7HZR7E" colab_type="code" outputId="cbe8ac86-4153-430e-c0b2-4a77b311cf37" colab={"base_uri": "https://localhost:8080/", "height": 153}
colnums = len(df.columns)
for i in df.columns:
try:
if df[i].dtype.name == 'object':
df[i] = df[i].astype('category')
else:
df[i].astype('float32')
except:
continue
print(df.head())
# + id="BpGxC2v2CDi1" colab_type="code" outputId="c3cedf9c-b6cc-49ac-c514-bf05ea17e239" colab={"base_uri": "https://localhost:8080/", "height": 153}
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler, StandardScaler
np.random.seed(1)
vals = df.values.copy()
total_nums = len(vals)
train, validation = train_test_split(df, test_size=0.5,
random_state=42,
shuffle=True)
validation = validation.reindex(sorted(validation.columns), axis=1)
validation.to_csv(path + '_For_Test.csv')
print(validation.head())
# + id="G-52cnSrZHGm" colab_type="code" outputId="04d75d85-891b-40ae-876a-3b2bd90bac3a" colab={"base_uri": "https://localhost:8080/", "height": 102}
df = train
categorical = df.select_dtypes(['category']).columns
print(categorical)
for f in categorical:
dummies = pd.get_dummies(df[f], prefix = f, prefix_sep = '_')
df = pd.concat([df, dummies], axis = 1)
# drop original categorical features
df.drop(categorical, axis = 1, inplace = True)
df.to_csv(path + '_For_Training.csv', index=False)
# + [markdown] id="OnJXo9LfUGaJ" colab_type="text"
# # Define VAE
# + id="LmIPmd5wjAM7" colab_type="code" colab={}
np.random.seed(1)
df = pd.read_csv(path + '_For_Training.csv')
train = df.values.copy()
train.astype('float32')
scaler = MinMaxScaler()
train = scaler.fit_transform(train)
x_train, x_test = train_test_split(train, test_size=0.5,
random_state=42,
shuffle=True)
# + id="uNf-3EMl2usm" colab_type="code" outputId="c0cf40ca-564b-4ae4-c952-6495d77c066d" colab={"base_uri": "https://localhost:8080/", "height": 51}
original_dim = x_train.shape[1]
x_train = np.reshape(x_train, [-1, original_dim])
x_test = np.reshape(x_test, [-1, original_dim])
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
print(x_train.shape)
print(x_test.shape)
# + id="uy-JqW7Jpuyz" colab_type="code" colab={}
np.random.seed(1)
set_random_seed(1)
class VAE:
def __init__(self, input_shape=(original_dim,),
intermediate_dim=128, latent_dim=2, summary=False):
self._build_model(input_shape,
intermediate_dim,
latent_dim, summary)
def _build_model(self, input_shape, intermediate_dim, latent_dim,
summary=False):
inputs = Input(shape=input_shape, name='encoder_input')
x = inputs
x = Dense(intermediate_dim, activation='relu')(x)
x = Dense(intermediate_dim//2, activation='relu')(x)
z_mean = Dense(latent_dim, name='z_mean')(x)
z_log_var = Dense(latent_dim, name='z_log_var')(x)
z = Lambda(self.sampling, output_shape=(latent_dim,),
name='z')([z_mean, z_log_var])
self.encoder = Model(inputs, [z_mean, z_log_var, z],
name='encoder')
latent_inputs = Input(shape=(latent_dim,),
name='z_sampling')
x = latent_inputs
x = Dense(intermediate_dim//2, activation='relu')(x)
x = Dense(intermediate_dim, activation='relu')(x)
outputs = Dense(original_dim, activation='sigmoid')(x)
self.decoder = Model(latent_inputs, outputs, name='decoder')
outputs = self.decoder(self.encoder(inputs)[2])
self.vae = Model(inputs, outputs, name='vae_mlp')
reconstruction_loss = binary_crossentropy(inputs, outputs)
reconstruction_loss *= original_dim
kl_loss = 1 + z_log_var - K.square(z_mean) - K.exp(z_log_var)
kl_loss = K.sum(kl_loss, axis=-1)
kl_loss *= -0.5
vae_loss = K.mean(reconstruction_loss + kl_loss)
self.vae.add_loss(vae_loss)
self.vae.compile(optimizer='adam')
if summary:
print(self.vae.summary())
def sampling(self, args):
z_mean, z_log_var = args
batch = K.shape(z_mean)[0]
dim = K.int_shape(z_mean)[1]
epsilon = K.random_normal(shape=(batch, dim))
return z_mean + K.exp(0.5 * z_log_var) * epsilon
def fit(self, x_train, x_test, epochs=100, batch_size=100,
verbose=1):
self.vae.fit(x_train,
shuffle=True,
epochs=epochs,
batch_size=batch_size,
verbose=verbose,
validation_data=(x_test, None))
def encoder_predict(self, x_test, batch_size=100):
return self.encoder.predict(x_test,
batch_size=batch_size)
def generate(self, latent_val, batch_size=100):
return self.decoder.predict(latent_val)
def predict(self, x_test, batch_size=100):
prediction = self.vae.predict(x_test,
batch_size=batch_size)
return prediction
# + [markdown] id="KXhRjx02UYZN" colab_type="text"
# ## Training VAE
# + [markdown] id="Wd02caRJWGG8" colab_type="text"
# Just let the last value to test
# + id="XBbIF17p8iY2" colab_type="code" outputId="8a6e63b1-56df-4705-aa61-5117b94e4516" colab={"base_uri": "https://localhost:8080/", "height": 4406}
np.random.seed(1)
set_random_seed(1)
latent_dim = original_dim//2
if latent_dim < 2:
latent_dim = 2
vae = VAE(intermediate_dim=intermediate_dim, latent_dim=latent_dim)
vae.fit(x_train, x_test, epochs=120)
# + id="qTfvSN_R2c76" colab_type="code" outputId="9efc5c86-43a0-4827-aa96-668950769406" colab={"base_uri": "https://localhost:8080/", "height": 51}
import time
np.random.seed(1)
start = time.time()
x_test = np.reshape(x_test, (-1, original_dim))
x_test_encoded = vae.encoder.predict(x_test)
x_test_encoded = np.asarray(x_test_encoded)
total_nums = 2
results = []
for i in range(x_test_encoded.shape[1]):
latent_gen = []
for _ in range(total_nums):
epsilon = np.random.normal(0., 1., x_test_encoded.shape[2])
latent_gen.extend([x_test_encoded[0, i, :] + np.exp(x_test_encoded[1, i, :]*0.5)*epsilon])
latent_gen = np.asarray(latent_gen)
results.append(vae.generate(latent_gen))
results = np.asarray(results)
results = np.reshape(results, (-1, original_dim))
print(results.shape)
results = scaler.inverse_transform(results)
end = time.time()
print('Total time=', end-start)
# + id="fC6DiGV8MV9W" colab_type="code" colab={}
d = {}
names = list(df)
for i, name in enumerate(names):
d[name] = results[:, i]
df = pd.DataFrame(data=d)
# + id="49_bbWWCsrHG" colab_type="code" outputId="1590e1aa-7d2c-4871-a4ab-f7a6f7fbd51a" colab={"base_uri": "https://localhost:8080/", "height": 238}
names = list(df)
c_dict = {}
for n in names:
if '_' in n:
index = n.index('_')
c_dict[n[:index]] = [c for c in names if n[:index+1] in c]
values = []
for key, items in c_dict.items():
dummies = df[items]
d_names = list(dummies)
c_dict = {}
for n in d_names:
c_dict[n] = n[n.index('_')+1:]
dummies.rename(columns=c_dict,
inplace=True)
df[key] = dummies.idxmax(axis=1)
df.drop(items, axis=1, inplace=True)
print(df.head())
# + id="1ljK-ceANBih" colab_type="code" colab={}
df = df.reindex(sorted(df.columns), axis=1)
df.to_csv(path + '_vae.csv')
# + [markdown] colab_type="text" id="SZr05ADu6LQP"
# # Dropout VAE
# + colab_type="code" id="9KXhkZujT29r" colab={}
np.random.seed(1)
df = pd.read_csv(path + '_For_Training.csv')
train = df.values.copy()
train.astype('float32')
scaler = MinMaxScaler()
train = scaler.fit_transform(train)
x_train, x_test = train_test_split(train, test_size=0.5,
random_state=42,
shuffle=True)
# + colab_type="code" outputId="802d6f66-3565-421f-c044-65e29193a5cc" id="fa5UJs9cT29v" colab={"base_uri": "https://localhost:8080/", "height": 51}
original_dim = x_train.shape[1]
x_train = np.reshape(x_train, [-1, original_dim])
x_test = np.reshape(x_test, [-1, original_dim])
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
print(x_train.shape)
print(x_test.shape)
# + colab_type="code" id="MnKFnBX-6LQR" colab={}
from keras.regularizers import l2
from keras.losses import categorical_crossentropy
np.random.seed(1)
class DropoutVAE:
def __init__(self, input_shape=(original_dim,),
intermediate_dim=32, latent_dim=3, dropout=0.05,
summary=False):
self._build_model(input_shape,
intermediate_dim,
latent_dim, summary,
dropout)
def _build_model(self, input_shape, intermediate_dim, latent_dim,
summary=False, dropout=0.05):
inputs = Input(shape=input_shape, name='encoder_input')
x = inputs
x = Dense(intermediate_dim, activation='relu')(x)
x = Dense(intermediate_dim//2, activation='relu')(x)
z_mean = Dense(latent_dim, name='z_mean')(x)
z_log_var = Dense(latent_dim, name='z_log_var')(x)
self.encoder = Model(inputs, [z_mean, z_log_var], name='encoder')
latent_inputs = Input(shape=(latent_dim,), name='z_sampling')
x = latent_inputs
x = Dense(intermediate_dim//2, activation='relu',
kernel_regularizer=l2(1e-4),
bias_regularizer=l2(1e-4))(x)
x = Dropout(dropout)(x)
x = Dense(intermediate_dim, activation='relu',
kernel_regularizer=l2(1e-4),
bias_regularizer=l2(1e-4))(x)
x = Dropout(dropout)(x)
outputs = Dense(original_dim, activation='sigmoid',
kernel_regularizer=l2(1e-4),
bias_regularizer=l2(1e-4))(x)
self.decoder = Model(latent_inputs,
outputs,
name='decoder')
outputs = self.decoder(self.encoder(inputs)[0])
self.vae = Model(inputs, outputs,
name='vae_mlp')
reconstruction_loss = binary_crossentropy(inputs, outputs)
reconstruction_loss *= original_dim
kl_loss = 1 + z_log_var - K.square(z_mean) - K.exp(z_log_var)
kl_loss = K.sum(kl_loss, axis=-1)
kl_loss *= -0.5
vae_loss = K.mean(reconstruction_loss + kl_loss)
self.vae.add_loss(vae_loss)
self.vae.compile(optimizer='adam')
if summary:
print(self.vae.summary())
def fit(self, x_train, x_test, epochs=100, batch_size=100,
verbose=1):
self.vae.fit(x_train,
shuffle=True,
epochs=epochs,
batch_size=batch_size,
verbose=verbose,
validation_data=(x_test, None))
def encoder_predict(self, x_test, batch_size=100):
return self.encoder.predict(x_test,
batch_size=batch_size)
def generate(self, latent_val, batch_size=100):
return self.decoder.predict(latent_val)
def predict(self, x_test, batch_size=1, nums=1000):
predict_stochastic = K.function([self.vae.layers[0].input,
K.learning_phase()],
[self.vae.get_output_at(0)])
Yt_hat = []
for _ in range(nums):
Yt_hat.append(predict_stochastic([x_test, 1]))
return np.asarray(Yt_hat)
def mean_predict(self, x_test, batch_size=1, nums=1000):
predict_stochastic = K.function([self.decoder.layers[0].input,
K.learning_phase()],
[self.decoder.get_output_at(0)])
latents = self.encoder.predict(x_test)[0]
Yt_hat = []
for _ in range(nums):
Yt_hat.append(predict_stochastic([latents, 1]))
return np.asarray(Yt_hat)
# + [markdown] colab_type="text" id="TsARTKz36LQZ"
# ## Train and evaluate Dropout VAE
# + colab_type="code" id="pkdmVggK6LQp" outputId="4402b56f-b0f9-40aa-faea-7a4a48fa7fd5" colab={"base_uri": "https://localhost:8080/", "height": 3743}
np.random.seed(1)
set_random_seed(1)
latent_dim = original_dim//2
if latent_dim < 2:
latent_dim = 2
vae = DropoutVAE(intermediate_dim=intermediate_dim,
dropout=0.1, latent_dim=latent_dim,
summary=True)
vae.fit(x_train, x_test, epochs=100)
# + colab_type="code" id="2HndA3MG6LQs" outputId="96e937b3-e76e-4712-c649-e53ef0d97439" colab={"base_uri": "https://localhost:8080/", "height": 51}
np.random.seed(1)
x_test = np.reshape(x_test, (-1, original_dim))
print(x_test.shape)
print(x_test[0].reshape(-1, original_dim).shape)
# + id="_sgO9JKSqWsW" colab_type="code" outputId="94ebe509-31e1-4806-de87-0098810afb5d" colab={"base_uri": "https://localhost:8080/", "height": 68}
import time
np.random.seed(1)
set_random_seed(1)
start = time.time()
total_nums = 2
results = []
x_test_encoded = vae.mean_predict(x_test, nums=total_nums, batch_size=100)
results = x_test_encoded
results = np.asarray(results)
print(results.shape)
results = results.reshape(total_nums*results.shape[2], original_dim)
results = scaler.inverse_transform(results)
print(results.shape)
end = time.time()
print('Total time=', end-start)
# + [markdown] id="POfJxgHamlmx" colab_type="text"
# ## Results
# + colab_type="code" id="YIlGSzekAYlO" colab={}
d = {}
names = list(df)
for i, name in enumerate(names):
d[name] = results[:, i]
df = pd.DataFrame(data=d)
# + colab_type="code" outputId="4f17fd64-e9d7-46b3-fc6b-ee8a777dd89f" id="shurAQCSAYlV" colab={"base_uri": "https://localhost:8080/", "height": 238}
names = list(df)
c_dict = {}
for n in names:
if '_' in n:
index = n.index('_')
c_dict[n[:index]] = [c for c in names if n[:index+1] in c]
values = []
for key, items in c_dict.items():
dummies = df[items]
d_names = list(dummies)
c_dict = {}
for n in d_names:
c_dict[n] = n[n.index('_')+1:]
dummies.rename(columns=c_dict,
inplace=True)
df[key] = dummies.idxmax(axis=1)
df.drop(items, axis=1, inplace=True)
print(df.head())
# + colab_type="code" id="n8dLIoyVW_Kn" colab={}
df = df.reindex(sorted(df.columns), axis=1)
df.to_csv(path + '_dropout.csv')
# + [markdown] id="xL6N8v2sYTNm" colab_type="text"
# # Transform categorical -> number
# + id="FgVhmliYYzwS" colab_type="code" colab={}
np.random.seed(1)
validation.to_csv(path + '_For_Test.csv')
df = pd.read_csv(path + '_For_Test.csv',
na_filter=True,
verbose=False,
skip_blank_lines=True,
na_values=na_values,
keep_default_na=False)
df_mc = pd.read_csv(path + '_dropout.csv',
na_filter=True,
verbose=False,
skip_blank_lines=True,
na_values=na_values,
keep_default_na=False)
df_vae = pd.read_csv(path + '_vae.csv',
na_filter=True,
verbose=False,
skip_blank_lines=True,
na_values=na_values,
keep_default_na=False)
df.drop('Unnamed: 0', axis=1, inplace=True)
df_mc.drop('Unnamed: 0', axis=1, inplace=True)
df_vae.drop('Unnamed: 0', axis=1, inplace=True)
names = list(df)
# + id="HSbyWbb0ZjAJ" colab_type="code" outputId="c120b248-d395-406b-fb27-9f05ac3ed6ed" colab={"base_uri": "https://localhost:8080/", "height": 102}
from sklearn.preprocessing import LabelEncoder
colnums = len(df.columns)
for i in df.columns:
try:
if df[i].dtype.name == 'object':
df[i] = df[i].astype('category')
except:
continue
cat_columns = df.select_dtypes(['category']).columns
print(cat_columns)
for col in cat_columns:
le = LabelEncoder()
df[col] = le.fit_transform(df[col].values)
df_mc[col] = le.transform(df_mc[col].values)
df_vae[col] = le.transform(df_vae[col].values)
# + id="J-scTKTHZ9Wk" colab_type="code" colab={}
df = df.reindex(sorted(df.columns), axis=1)
df_mc = df_mc.reindex(sorted(df_mc.columns), axis=1)
df_vae = df_vae.reindex(sorted(df_vae.columns), axis=1)
df.to_csv(path + '_For_Test_encoded.csv')
df_mc.to_csv(path + '_dropout_encoded.csv')
df_vae.to_csv(path + '_vae_encoded.csv')
# + [markdown] id="gLy-z65OUnRj" colab_type="text"
# # Predicting with generated data
# + id="-W-tX8ysUuOC" colab_type="code" colab={}
df = pd.read_csv(path + '_For_Test_encoded.csv',
na_filter=True,
verbose=False,
skip_blank_lines=True,
na_values=na_values,
keep_default_na=False)
df_mc = pd.read_csv(path + '_dropout_encoded.csv',
na_filter=True,
verbose=False,
skip_blank_lines=True,
na_values=na_values,
keep_default_na=False)
df_vae = pd.read_csv(path + '_vae_encoded.csv',
na_filter=True,
verbose=False,
skip_blank_lines=True,
na_values=na_values,
keep_default_na=False)
df.drop('Unnamed: 0', axis=1, inplace=True)
df_mc.drop('Unnamed: 0', axis=1, inplace=True)
df_vae.drop('Unnamed: 0', axis=1, inplace=True)
# + id="8F40hUKFVC3E" colab_type="code" outputId="868e331b-d5f4-456c-fa1e-c1100b18363a" colab={"base_uri": "https://localhost:8080/", "height": 425}
print(df.head())
print(df_mc.head())
print(df_vae.head())
# + id="0tyhyfWHXbn4" colab_type="code" outputId="3d7fd433-154f-4ec9-c735-9dd979ac84c0" colab={"base_uri": "https://localhost:8080/", "height": 51}
y = df['Churn'].values
df.drop(['Churn'], axis=1, inplace=True)
X = df.values
print(y.shape)
print(X.shape)
# + id="rwzr1m5TX5QC" colab_type="code" outputId="39d6ca89-2020-4aed-e167-edf6e3b32302" colab={"base_uri": "https://localhost:8080/", "height": 51}
y_mc = df_mc['Churn'].values
df_mc.drop(['Churn'], axis=1, inplace=True)
X_mc = df_mc.values
print(y_mc.shape)
print(X_mc.shape)
# + id="xbq6H2ZnYXqz" colab_type="code" outputId="74692b43-eaef-4c76-ed92-8f85ea7788d0" colab={"base_uri": "https://localhost:8080/", "height": 51}
y_vae = df_vae['Churn'].values
df_vae.drop(['Churn'], axis=1, inplace=True)
X_vae = df_vae.values
print(y_vae.shape)
print(X_vae.shape)
# + [markdown] id="iWMcVL29ggWg" colab_type="text"
# ### Original data
# + id="ob3Et0uia9YZ" colab_type="code" outputId="37a8a4ef-07d8-45a5-a26e-62fbd971b139" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
scores = cross_val_score(clf, X, y, cv=10)
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# + colab_type="code" id="ySOOpbfD5RP1" outputId="0e0625ea-e349-4182-8105-9061b2db55e9" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
scores = cross_val_score(clf, X_mc, y_mc, cv=10)
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
# + colab_type="code" id="lycl8s_A5Vxf" outputId="c8a38e16-2442-4873-969f-d24286470ba6" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
scores = cross_val_score(clf, X_vae, y_vae, cv=10)
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(),
scores.std() * 2))
# + [markdown] colab_type="text" id="hSUz0c0ngkj5"
# ### DropoutVAE generated data
# + colab_type="code" id="y4-GzQ_bbPCo" outputId="4bd608e5-9042-48bc-fba3-adff537b221c" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
clf.fit(X_mc, y_mc)
print(clf.score(X, y))
# + [markdown] colab_type="text" id="NL3OtgaGgxQB"
# ### VAE generated data
# + id="Crz0AfZ-bSg-" colab_type="code" outputId="1ba2015d-d893-4b8d-9e6e-97b570b14300" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
clf.fit(X_vae, y_vae)
print(clf.score(X, y))
# + [markdown] colab_type="text" id="mrCqDtssg4nB"
# ### Original data with Dropout VAE outcome
# + colab_type="code" id="mH3gS3nacvV7" outputId="1af32f59-80e2-4397-c3b0-2257d0145b51" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
clf.fit(X, y)
print(clf.score(X_mc, y_mc))
# + [markdown] colab_type="text" id="YlER2gGnhBGJ"
# ### Original data with VAE outcome
# + colab_type="code" id="MezW_-9ZcvWA" outputId="2bd60e48-8253-409c-ca06-51f8a2c904c1" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
clf.fit(X, y)
print(clf.score(X_vae, y_vae))
# + id="JCm6BBN1zB-0" colab_type="code" outputId="1be2c881-9bd9-4ce5-843a-c388e91f4a98" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
clf.fit(X_mc, y_mc)
print(clf.score(X_vae, y_vae))
# + colab_type="code" id="XcarR61z4UqD" outputId="3d661cf2-eb9a-4d8f-be9d-ff7f9bdb4fe4" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(1)
from sklearn.model_selection import cross_val_score
clf = RandomForestClassifier(n_estimators=100,
max_depth=2, random_state=42)
clf.fit(X_vae, y_vae)
print(clf.score(X_mc, y_mc))
| DropoutVAE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''deformable_detr'': conda)'
# name: python3
# ---
import argparse
import datetime
import json
import random
import time
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import torch
from torch.utils.data import DataLoader
import datasets
import util.misc as utils
import datasets.samplers as samplers
from datasets import build_dataset, get_coco_api_from_dataset
from engine import evaluate, train_one_epoch
from models import build_model
import torchvision.transforms.functional as functional
import torch.nn.functional as F
from util.misc import nested_tensor_from_tensor_list
# + tags=[]
class Compose(object):
def __init__(self, transforms):
self.transforms = transforms
def __call__(self, image):
for t in self.transforms:
image = t(image)
return image
def __repr__(self):
format_string = self.__class__.__name__ + "("
for t in self.transforms:
format_string += "\n"
format_string += " {0}".format(t)
format_string += "\n)"
return format_string
class Normalize(object):
def __init__(self, mean, std):
self.mean = mean
self.std = std
def __call__(self, image):
image = functional.normalize(image, mean=self.mean, std=self.std)
return image
class ToTensor(object):
def __call__(self, img):
return functional.to_tensor(img)
def resize(image, size, max_size=None):
# size can be min_size (scalar) or (w, h) tuple
def get_size_with_aspect_ratio(image_size, size, max_size=None):
w, h = image_size
if max_size is not None:
min_original_size = float(min((w, h)))
max_original_size = float(max((w, h)))
if max_original_size / min_original_size * size > max_size:
size = int(round(max_size * min_original_size / max_original_size))
if (w <= h and w == size) or (h <= w and h == size):
return (h, w)
if w < h:
ow = size
oh = int(size * h / w)
else:
oh = size
ow = int(size * w / h)
return (oh, ow)
def get_size(image_size, size, max_size=None):
if isinstance(size, (list, tuple)):
return size[::-1]
else:
return get_size_with_aspect_ratio(image_size, size, max_size)
size = get_size(image.size, size, max_size)
rescaled_image = functional.resize(image, size)
return rescaled_image
class Resize(object):
def __init__(self, sizes, max_size=None):
assert isinstance(sizes, (list, tuple))
self.sizes = sizes
self.max_size = max_size
def __call__(self, img):
size = self.sizes
return resize(img, size, self.max_size)
# -
checkpoint = torch.load("exps/r50_deformable_detr/checkpoint0019.pth", map_location='cuda:0')
device = torch.device("cuda")
args = checkpoint["args"]
model, _, _ = build_model(args)
model.load_state_dict(checkpoint['model'])
model.to(device)
# +
# load image
raw_img = plt.imread('data/tinydata/train2017/0000000.jpg')
h, w = raw_img.shape[0], raw_img.shape[1]
orig_size = torch.as_tensor([int(h), int(w)])
# normalize image
normalize = Compose([
ToTensor(),
Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225]),
Resize([800],max_size=1333),
])
img = normalize(raw_img)
inputs = nested_tensor_from_tensor_list([img]).to(device)
plt.axis('off')
plt.imshow(raw_img)
# -
outputs = model(inputs)
outputs["pred_logits"].shape
# +
out_prob = outputs['pred_logits'].softmax(-1)
out_prob
# -
keep = out_prob[0,:,0] > 0.6
keep
# +
out_bezierboxes = outputs['pred_bezierboxes'][0]
out_bezierboxes[keep,:]
# +
keep = labels = out_prob.max(-1)
img_h, img_w = orig_size.unbind(0)
# -
out_bezierboxes
scale_fct = torch.unsqueeze(torch.stack([img_w, img_h, img_w, img_h,img_w, img_h, img_w, img_h,img_w, img_h, img_w, img_h,img_w, img_h, img_w, img_h], dim=0), dim=0)
bezierboxes = out_bezierboxes * scale_fct[:, None, :]
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="lP6JLo1tGNBg"
# # Artificial Neural Network
# + [markdown] colab_type="text" id="gWZyYmS_UE_L"
# ### Importing the libraries
# -
#for debug purpose
# %qtconsole --style solarized-dark
# + colab={} colab_type="code" id="MxkJoQBkUIHC"
import numpy as np
import pandas as pd
import tensorflow as tf
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 2400, "status": "ok", "timestamp": 1590257449959, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="ZaTwK7ojXr2F" outputId="0b27a96d-d11a-43e8-ab4b-87c1f01896fe"
tf.__version__
# + [markdown] colab_type="text" id="1E0Q3aoKUCRX"
# ## Part 1 - Data Preprocessing
# + [markdown] colab_type="text" id="cKWAkFVGUU0Z"
# ### Importing the dataset
# + colab={} colab_type="code" id="MXUkhkMfU4wq"
dataset = pd.read_csv('Churn_Modelling.csv')
X = dataset.iloc[:, 3:-1].values
y = dataset.iloc[:, -1].values
# + colab={"base_uri": "https://localhost:8080/", "height": 137} colab_type="code" executionInfo={"elapsed": 2396, "status": "ok", "timestamp": 1590257449961, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="VYP9cQTWbzuI" outputId="797e7a64-9bac-436a-8c9c-94437e5e7587"
print(X)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 2391, "status": "ok", "timestamp": 1590257449961, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="38vKGE6Nb2RR" outputId="a815e42a-e0dd-4cb5-ab97-b17ead98fbc3"
print(y)
# + [markdown] colab_type="text" id="N6bQ0UgSU-NJ"
# ### Encoding categorical data
# + [markdown] colab_type="text" id="le5MJreAbW52"
# Label Encoding the "Gender" column
# + colab={} colab_type="code" id="PxVKWXxLbczC"
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
X[:, 2] = le.fit_transform(X[:, 2])
# + colab={"base_uri": "https://localhost:8080/", "height": 137} colab_type="code" executionInfo={"elapsed": 2719, "status": "ok", "timestamp": 1590257450295, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="-M1KboxFb6OO" outputId="e2b8c7e8-0cbc-4cdf-f4eb-7f0853a00b88"
print(X)
# + [markdown] colab_type="text" id="CUxGZezpbMcb"
# One Hot Encoding the "Geography" column
# + colab={} colab_type="code" id="AMXC8-KMVirw"
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
ct = ColumnTransformer(transformers=[('encoder', OneHotEncoder(), [1])], remainder='passthrough')
X = np.array(ct.fit_transform(X))
# + colab={"base_uri": "https://localhost:8080/", "height": 137} colab_type="code" executionInfo={"elapsed": 2713, "status": "ok", "timestamp": 1590257450296, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="ZcxwEon-b8nV" outputId="23a98af4-5e33-4b26-c27b-f06e3c5d2baf"
print(X)
# + [markdown] colab_type="text" id="vHol938cW8zd"
# ### Splitting the dataset into the Training set and Test set
# + colab={} colab_type="code" id="Z-TDt0Y_XEfc"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# + [markdown] colab_type="text" id="RE_FcHyfV3TQ"
# ### Feature Scaling
# + colab={} colab_type="code" id="ViCrE00rV8Sk"
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + [markdown] colab_type="text" id="-zfEzkRVXIwF"
# ## Part 2 - Building the ANN
# + [markdown] colab_type="text" id="KvdeScabXtlB"
# ### Initializing the ANN
# + colab={} colab_type="code" id="3dtrScHxXQox"
ann = tf.keras.models.Sequential()
# + [markdown] colab_type="text" id="rP6urV6SX7kS"
# ### Adding the input layer and the first hidden layer
# + colab={} colab_type="code" id="bppGycBXYCQr"
ann.add(tf.keras.layers.Dense(units=6, activation='relu'))
# + [markdown] colab_type="text" id="BELWAc_8YJze"
# ### Adding the second hidden layer
# + colab={} colab_type="code" id="JneR0u0sYRTd"
ann.add(tf.keras.layers.Dense(units=6, activation='relu'))
# + [markdown] colab_type="text" id="OyNEe6RXYcU4"
# ### Adding the output layer
# + colab={} colab_type="code" id="Cn3x41RBYfvY"
ann.add(tf.keras.layers.Dense(units=1, activation='sigmoid'))
# + [markdown] colab_type="text" id="JT4u2S1_Y4WG"
# ## Part 3 - Training the ANN
# + [markdown] colab_type="text" id="8GWlJChhY_ZI"
# ### Compiling the ANN
# + colab={} colab_type="code" id="fG3RrwDXZEaS"
ann.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# + [markdown] colab_type="text" id="0QR_G5u7ZLSM"
# ### Training the ANN on the Training set
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 33685, "status": "ok", "timestamp": 1590257481284, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="nHZ-LKv_ZRb3" outputId="718cc4b0-b5aa-40f0-9b20-d3d31730a531"
ann.fit(X_train, y_train, batch_size = 32, epochs = 100)
# + [markdown] colab_type="text" id="tJj5k2MxZga3"
# ## Part 4 - Making the predictions and evaluating the model
# + [markdown] colab_type="text" id="84QFoqGYeXHL"
# ### Predicting the result of a single observation
# + [markdown] colab_type="text" id="CGRo3eacgDdC"
# **Homework**
#
# Use our ANN model to predict if the customer with the following informations will leave the bank:
#
# Geography: France
#
# Credit Score: 600
#
# Gender: Male
#
# Age: 40 years old
#
# Tenure: 3 years
#
# Balance: \$ 60000
#
# Number of Products: 2
#
# Does this customer have a credit card? Yes
#
# Is this customer an Active Member: Yes
#
# Estimated Salary: \$ 50000
#
# So, should we say goodbye to that customer?
# + [markdown] colab_type="text" id="ZhU1LTgPg-kH"
# **Solution**
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 33990, "status": "ok", "timestamp": 1590257481594, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="2d8IoCCkeWGL" outputId="957f3970-e197-4c3b-a150-7f69dc567f5d"
print(ann.predict(sc.transform([[1, 0, 0, 600, 1, 40, 3, 60000, 2, 1, 1, 50000]])) > 0.5)
# + [markdown] colab_type="text" id="wGjx94g2n7OV"
# Therefore, our ANN model predicts that this customer stays in the bank!
#
# **Important note 1:** Notice that the values of the features were all input in a double pair of square brackets. That's because the "predict" method always expects a 2D array as the format of its inputs. And putting our values into a double pair of square brackets makes the input exactly a 2D array.
#
# **Important note 2:** Notice also that the "France" country was not input as a string in the last column but as "1, 0, 0" in the first three columns. That's because of course the predict method expects the one-hot-encoded values of the state, and as we see in the first row of the matrix of features X, "France" was encoded as "1, 0, 0". And be careful to include these values in the first three columns, because the dummy variables are always created in the first columns.
# + [markdown] colab_type="text" id="u7yx47jPZt11"
# ### Predicting the Test set results
# + colab={"base_uri": "https://localhost:8080/", "height": 137} colab_type="code" executionInfo={"elapsed": 33987, "status": "ok", "timestamp": 1590257481595, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="nIyEeQdRZwgs" outputId="82330ba8-9bdc-4fd1-d3cf-b6d78ee7c2a3"
y_pred = ann.predict(X_test)
y_pred = (y_pred > 0.5)
print(np.concatenate((y_pred.reshape(len(y_pred),1), y_test.reshape(len(y_test),1)),1))
# + [markdown] colab_type="text" id="o0oyfLWoaEGw"
# ### Making the Confusion Matrix
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 33981, "status": "ok", "timestamp": 1590257481595, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhEuXdT7eQweUmRPW8_laJuPggSK6hfvpl5a6WBaA=s64", "userId": "15047218817161520419"}, "user_tz": -240} id="ci6K_r6LaF6P" outputId="4d854e9e-22d5-432f-f6e5-a102fe3ae0bd"
from sklearn.metrics import confusion_matrix, accuracy_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
accuracy_score(y_test, y_pred)
| MachineLearningAZ/Part8DeepLearning/ANN/Python/artificial_neural_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chinese Poetry Generator
#
# Generates Tang poetry using the CharRNN model
# Data: from https://github.com/chinese-poetry/chinese-poetry
# +
import random
from collections import Counter
import torch
from torch.utils.data import Dataset, DataLoader
from torch import nn
import torch.optim as optim
from zhconv import convert
from tqdm.auto import tqdm
import numpy as np
# -
from data_loader import ParseRawData
# This loads the json data and only takes the main body of each poem
data_all = ParseRawData()
len(data_all)
# We have 57598 Tang Poems in total.
data_all[0]
# # preprocess data
# Filter and prepare data:
# - convert traditional Chinese to simplified Chinese for better readability
# - only take poems with 5 character lines - easier for model to learn the poem structure
comma = data_all[0][5]
data = [convert(x, "zh-hans") for x in data_all if len(x.split(comma)[0])==5]
len(data)
data[0]
char_counter = Counter([x for poem in data for x in poem])
len([x for x in char_counter if char_counter[x] < 5])
char_to_ix = {}
min_freq = 5 # a relatively large number is chosen to make the dataset smaller to fit my laptop
# EOP_TOKEN = '$'
for poem in data:
for char in poem:
# only take chars that appeared at least min_freq times
if (char not in char_to_ix) and (char_counter[char] >= min_freq):
char_to_ix[char] = len(char_to_ix)
# char_to_ix['$'] = len(char_to_ix)
# char_to_ix['<START>'] = len(char_to_ix)
ix_to_char = dict((i, w) for (w, i) in char_to_ix.items())
# +
# get input and target for training.
# use first 12 chars as input to predict the next char.
seq_len = 12
input_data = []
target = []
# EOP_TOKEN = '$'
PADDING = 'O'
char_to_ix[PADDING] = len(char_to_ix)
for poem in data:
# add EOP token
# poem += EOP_TOKEN
# Add 6 chars padding before the poem.
poem = PADDING * 6 + poem
for i in range(len(poem) - seq_len):
target.append(poem[i + seq_len])
input_data.append(poem[i:i+seq_len])
print("Number of training samples: {}".format(len(input_data)))
# -
# plus one to count for the unknown chars
vocab_size = len(char_to_ix) + 1
print("vocab size: {}".format(vocab_size))
input_data[0]
# +
def transform_X(text, char_to_ix, onehot=False):
"""Transforms one row of input text to index or onehot encoded arrays
input format: '秦川雄帝宅,'
output format: if onehot == False, output an index array same length as text.
otherwise, output onehot encoded array of shape (len(text), vocab_size).
"""
if onehot:
X = np.zeros((len(text), len(char_to_ix)+1))
for i, char in enumerate(text):
X[i, char_to_ix.get(char, len(char_to_ix))] = 1
else:
X = np.array([char_to_ix.get(char, len(char_to_ix)) for char in text])
return X
# def transform_y(text, char_to_ix):
# """Transform the target text into onehot encoded tensor"""
# y = np.zeros((len(char_to_ix)+1))
# y[char_to_ix.get(text, len(char_to_ix))] = 1
# return y
# For Pytorch crossentropyloss, the target does not need to be onehot encoded.
def transform_y(text, char_to_ix):
return char_to_ix.get(text, len(char_to_ix))
# dataset is too big for my laptop. Build a custom dataset
class PoemDataset(Dataset):
def __init__(self, data, target, transform_X, transform_y, char_to_ix):
self.data = data
self.target = target
self.transform_X = transform_X
self.transform_y = transform_y
self.char_to_ix = char_to_ix
def __len__(self):
"""Total number of samples"""
return len(self.data)
def __getitem__(self, index):
"""Generate one sample of data"""
X = self.transform_X(self.data[index], self.char_to_ix)
y = self.transform_y(self.target[index], self.char_to_ix)
sample = {"input": X, "target": y}
return sample
# -
poem_dataset = PoemDataset(input_data, target, transform_X, transform_y, char_to_ix)
batch_size = 128
dataloader = DataLoader(poem_dataset, batch_size=batch_size, shuffle=True)
# +
# # check we have the right shape
# for i, sample_batched in enumerate(dataloader):
# print(i, sample_batched['input'].size(),
# sample_batched['target'].size(),
# sample_batched['target'].dtype)
# if i == 3:
# break
# -
class PoemGenerationModel(nn.Module):
def __init__(self, vocab_size, embed_dim, hidden_dim):
super(PoemGenerationModel, self).__init__()
self.embedding = nn.Embedding(vocab_size, embed_dim)
self.lstm = nn.LSTM(embed_dim, hidden_dim, num_layers=2, batch_first=True)
self.dropout = nn.Dropout(0.2)
self.linear = nn.Linear(hidden_dim, vocab_size)
def forward(self, input_data):
embeds = self.embedding(input_data)
lstm_out, _ = self.lstm(embeds)
lstm_drop = self.dropout(lstm_out)
logits = self.linear(lstm_drop[:, -1, :].squeeze())
return logits
embed_dim = 256
hidden_dim = 256
lr = 0.001
device = 'cuda' if torch.cuda.is_available() else 'cpu'
device
model = PoemGenerationModel(vocab_size, embed_dim, hidden_dim).float().to(device)
loss_function = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=lr)
print(model)
def sample(preds, temperature=0.8):
"""Sample the output based predicted probabilities.
preds: 1D tensor. Logits from the model
temperature: When temperature is low, tend to choose the most likely words.
When temperature is high, model will be more adventurous.
"""
# helper function to sample an index from a probability array
preds = torch.nn.functional.softmax(preds, dim=0).detach().cpu().numpy()
exp_preds = np.power(preds, 1./temperature)
preds = exp_preds / np.sum(exp_preds)
pro = np.random.choice(range(len(preds)), 1, p=preds)
return int(pro.squeeze())
# +
def generate_poem(model, input_text, output_length=18, temperature=1):
"""Given input_text, generate a poem.
input_text need to be 6 chars, where last one is a comma.
Example input: "我有紫霞想,"
"""
if len(input_text) < seq_len:
input_text = PADDING * (seq_len - len(input_text)) + input_text
generated = ""
for i in range(output_length):
pred = generate_one_char(model, input_text, temperature=temperature)
generated += pred
input_text = input_text[1:] + pred
return generated
def generate_one_char(model, input_text, temperature=1):
# X_test = np.zeros((1, seq_len, vocab_size))
# for t, char in enumerate(input_text):
# X_test[0, t, char_to_ix.get(char, len(char_to_ix))] = 1
X_test = transform_X(input_text, char_to_ix)
pred = model(torch.from_numpy(X_test).unsqueeze(0).to(device)) #use less precision for laptop
next_index = sample(pred, temperature)
next_char = ix_to_char.get(next_index, "?")
return next_char
# -
def generate_sample():
# Prints generated sample text. Used during training to check the model performance
print('\n----- Generating text:')
# randomly pick the starting line of a poem as the seed
poem_index = random.randint(0, len(data))
print("Generating with seed: {}".format(data[poem_index][:seq_len]))
seed_text = data[poem_index][:seq_len]
for temperature in [0.2, 0.5, 0.7, 1.0]:
print('----- temperature:', temperature)
generated = generate_poem(model, seed_text, output_length=12, temperature=temperature)
print(generated)
seed_text = PADDING*6 + data[poem_index][:seq_len-6]
print("Generating with seed: {}".format(seed_text))
for temperature in [0.2, 0.5, 0.7, 1.0]:
print('----- temperature:', temperature)
generated = generate_poem(model, seed_text, temperature=temperature)
print(generated)
# +
n_epochs = 5
for epoch in range(n_epochs):
# model.train()
step = 0
# Dataloader returns the batches
for samples in tqdm(dataloader):
cur_batch_size = len(samples)
batch_X = samples['input'].to(device)
batch_y = samples['target'].to(device)
# Zero out the gradients before backpropagation
model.zero_grad()
y_pred = model(batch_X.long())
# Compute loss and update gradients
loss = loss_function(y_pred, batch_y.long())
loss.backward()
# `clip_grad_norm` helps prevent the exploding gradient problem in RNNs / LSTMs.
# nn.utils.clip_grad_norm_(model.parameters(), 1)
optimizer.step()
if step % 1000 == 0:
print("\n\nLoss at epoch {} step {}: {}".format(epoch, step, loss))
generate_sample()
step += 1
# -
save_path = "saved_model/charrnn_pytorch_embedding_model_v2"
# +
# torch.save(model.state_dict(), save_path)
# -
torch.save(model, save_path)
# 1st try of model with embedding:
# `PoemGenerationModel(
# (embedding): Embedding(5195, 128)
# (lstm): LSTM(128, 128, batch_first=True)
# (dropout): Dropout(p=0.2, inplace=False)
# (linear): Linear(in_features=128, out_features=5195, bias=True)
# )
# `
# seq=6
#
# 2nd try of model with embedding:
# `PoemGenerationModel(
# (embedding): Embedding(5196, 256)
# (lstm): LSTM(256, 256, num_layers=2, batch_first=True)
# (dropout): Dropout(p=0.2, inplace=False)
# (linear): Linear(in_features=256, out_features=5196, bias=True)
# )
# `
# seq=12
#
# # Load the saved model and test
# trained_model = PoemGenerationModel(vocab_size, embed_dim, hidden_dim)
# trained_model.load_state_dict(torch.load(save_path))
trained_model = torch.load(save_path)
trained_model.eval()
seed_text = "我有紫霞想,"
generated = generate_poem(model, seed_text, temperature=0.5)
print(generated)
generated = generate_poem(trained_model, seed_text, temperature=0.3)
print(generated)
generated = generate_poem(trained_model, seed_text, output_length=42, temperature=0.5)
print(generated)
generated = generate_poem(trained_model, "明月几时有,", output_length=18, temperature=0.3)
print(generated)
generated = generate_poem(trained_model, "八月湖水平,", output_length=18, temperature=0.2)
print(generated)
| generator_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
data = pd.read_csv('https://github.com/albahnsen/PracticalMachineLearningClass/raw/master/datasets/dataTrain_carListings.zip')
data_test = pd.read_csv('https://github.com/albahnsen/PracticalMachineLearningClass/raw/master/datasets/dataTest_carListings.zip', index_col=0)
Big_data=data.append(pd.DataFrame(data=data_test), ignore_index=True,sort=False)
Big_data['ind']=Big_data.index
Big_data.head(5)
Big_data.tail(5)
Big_data.shape
import category_encoders as ce
Big_data['Year_s']=Big_data[str('Year')]
X_1=pd.get_dummies(Big_data['Year_s'], prefix='Y')
data2=pd.DataFrame(Big_data['State'])
X_2 = ce.BinaryEncoder().fit_transform(data2)
data3=pd.DataFrame(Big_data['Make'])
X_3 = ce.BinaryEncoder().fit_transform(data3)
data4=pd.DataFrame(Big_data['Model'])
X_4 = ce.BinaryEncoder().fit_transform(data4)
New_data =pd.concat([Big_data[['ind','Price','Mileage']],X_1,X_2,X_3,X_4],axis=1)
New_data.shape
New_data.head(5)
train = New_data[New_data['ind']<500000]
test = New_data[New_data['ind']>=500000]
X_train=train.drop(['Price','ind'],axis=1)
X_train.head(5)
y_train = train['Price']
y_train.head(5)
X_train.shape
y_train.shape
X_test=test.drop(['Price','ind'],axis=1)
X_test.head(5)
X_test.shape
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import accuracy_score
from sklearn.ensemble import RandomForestRegressor
import numpy as np
from sklearn.metrics import mean_squared_error
RFreg = RandomForestRegressor()
RFreg
# max_features=6 is best and n_estimators=200 is sufficiently large
RFreg = RandomForestRegressor(bootstrap=True,criterion='mse',max_features=24, max_depth=20,n_estimators=200,
random_state=1, n_jobs=-1)
# fit and predict
RFreg.fit(X_train, y_train)
y_pred = RFreg.predict(X_test)
y_pred
y_pred=pd.DataFrame(y_pred)
y_pred['Price']=y_pred.iloc[:,0]
y_pred['ind']=y_pred.index
y_pred.head(5)
y_pred.shape
X_test.head(5)
X_test = X_test.reset_index(drop=True)
X_test['ind']=X_test.index
X_test.head(5)
data_test['ind']=data_test.index
y_est=pd.merge(data_test,y_pred,on='ind',how='left')
y_est.head(5)
y_est.shape
Price_model=data.groupby('Model')['Price'].mean()
Price_model.head(10)
Price_pred=y_est.groupby('Model')['Price'].mean()
Price_pred.head(10)
Price_model=pd.DataFrame(Price_model)
Price_model.head(5)
Price_pred=pd.DataFrame(Price_pred)
Price_pred.head(5)
Price_eval=pd.merge(Price_model,Price_pred,on="Model")
Price_eval
y_pred_f=y_pred["Price"]
y_pred_f.head(5)
y_pred_f.shape
y_pred_f.to_csv('test_submission.csv', index_label='ID')
Price_eval.to_csv('Price_eval.csv', index_label='ID')
| P1/New P1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# There are two basic types of graphs/visualizations: ones that summarize all the information for the basin and those that summarize data at a particular flow monitor.
#
# Summary graphics include:
# 1. dot matrix which compares number of problematic monitors to the number in other basins
# * illustrates how this basin compares to the other basins, which helps when comparing priority across basins
# 1. dot matrix which compares performance metrics and the number of badly performing flow monitor locations for each metric
# * helps illustrate what the most prevalent issues are in this basin
# * Are we reaching capacity? Is it infiltration? Is in inflow? RDII?
# 1. heatmap of performance metrics and flow monitors
# * each flow monitor gets a ranking for each performance metric that is either good, bad, or really bad
# * purpose: highlight most problematic flow monitor/basin locations
# * (probably create more official sounding rankings)
# * this allows for priority sorting
# 1. box plot of Q/Qc for each flow monitor
# * purpose: highlight potential capacity issues
# * contains information about Qmax, Qavg, and Qmin
# * Could force Qavg to be dry weather avg - need to ask
#
# Wet weather graphics include:
# 1. horizontal barchart with all storm dates, sorted by average event rainfall across the flow monitors in decreasing order
# 1. vertical barchart with Q vs I projects for worst performing flow monitors
# 1. heatmap of each flow monitor's net I&I compared across the 5 biggest storm events
#
# Flow monitor graphics include:
# 1. bullet graph of performance metrics
# * d/D dry, d/D wet, base infiltration as percent of average flow, normalized RDII, and capture coefficient
# 1. map of monitor location
# 1. box plot of dry weather data comparing weekday and weekend
# 1. gross Q (same y axis limits as net)
# 1. net Q (same y axis limits as gross)
# 1. d/D
# 1. vertical bar chart for net I&I at the flow monitor for each storm event
# 1. scatterplot/line chart of rainfall inches vs. net I&I
#
# NOTE: key missing item in the reports is how previous work has impacted the system. Are we making a difference?
# How can we include this information?
# +
# IMPORTS, DATA READING
import pickle
import math
import numpy as np
from os import mkdir
import pandas as pd
import datetime as dt
import scipy.stats as stats
import matplotlib.pyplot as plt
import matplotlib
homeDir = 'P:\\PW-WATER SERVICES\\TECHNICAL SERVICES\\Anna'
pickleLocation = homeDir + '\\2018\\Python Objects\\'
fmdataFile = homeDir + '\\FMdata.csv'
saveDir = homeDir + '\\2018\\Yearly Report'
# +
'''
basinDryWeather is a dictionary with flow monitors names as keys that
contain the following dictionaries:
* 'Weekday' : {
'd/D' : a numpy array
'Gross Q' : a pandas series (index: datetime, values: Q)
'DataFrame' : pandas data frame (index:time, columns: date,
values: q)
'Gross Diurnal' : pandas series (index: time, values: Q)
'Net Diurnal' : pandas series (index: time, values: Q)
}
* 'Weekend' : {
'd/D' : a numpy array
'Gross Q' :
'DataFrame' : pandas data frame (index:time, columns: date,
values: q)
'Gross Diurnal' : pandas series (index: time, values: Q)
'Net Diurnal' : pandas series (index: time, values: Q)
}
* 'Overall' : {
'd/D' : a numpy array
'Base Infiltration' : numpy float
}'''
with open(pickleLocation + 'basinDryWeather.pickle', 'rb') as handle:
basinDryWeather = pickle.load(handle)
'''
stormsDict is a dictionary with flow monitors as keys that contain a pandas
dataframe, dfStorms, with storm start times as indices and the following
columns:
* Storm RT : storm rain total in inches
* Storm Dur : storm duration in hours
* Event RT : event rain total in inches
* Event Dur : event duration in hours
* Gross Vol: gross I&I volume in MG
* Net Vol : net I&I volume in MG
* Capt Coeff : capture coefficient (vol I&I/vol rainfall)
* RDII : rain dependent I&I ranking (gal/in(rain)/in-mi(pipe))'''
with open(pickleLocation + 'stormsDict.pickle', 'rb') as handle:
stormDict = pickle.load(handle)
'''gageStorms is a dictionary with rain gages as keys that contain a pandas
dataframe, dfStorms, with storm start tiems as indices and the following
columns:
* Storm RT : storm rain total in inches
* Storm Dur : storm duration in hours
* Event RT : event rain total in inches
* Event Dur : event duration in hours'''
with open(pickleLocation + 'gageStorms.pickle', 'rb') as handle:
gageStorms = pickle.load(handle)
# -
basinDryWeather['BC32']['Weekday']['Net Q']
# +
df = (basinDryWeather['BC12']['Weekday']['DataFrame']
- basinDryWeather['BC18']['Weekday']['DataFrame']
- basinDryWeather['BC13']['Weekday']['DataFrame'])
fig, ax = plt.subplots()
df.plot(ax = ax,
color = 'xkcd:light grey',
legend = False)
df = (basinDryWeather['BC12']['Weekday']['DataFrame']
- basinDryWeather['BC13']['Weekday']['DataFrame'])
df.plot(color = 'xkcd:light grey',
legend = False)
df = (basinDryWeather['BC12']['Weekday']['DataFrame']
- basinDryWeather['BC18']['Weekday']['DataFrame'])
df.plot(color = 'xkcd:light grey',
legend = False)
basinDryWeather['BC13']['Weekday']['DataFrame'].plot(
color = 'xkcd:light grey',
legend = False)
basinDryWeather['BC12']['Weekday']['DataFrame'].plot(
color = 'xkcd:light grey',
legend = False)
# +
def prettyxTime(ax):
ticks = ax.get_xticks()
ax.set_xticks(np.linspace(ticks[0],24*3600,5))
ax.set_xticks(np.linspace(ticks[0],24*3600,25),minor=True)
fmname = 'BC32'
df = basinDryWeather[fmname]['Weekday']['DataFrame']
fig, ax = plt.subplots(figsize = (7.5,2))
df.plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:light grey')
df.mean(axis=1).plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:leaf green')
ax.set_title(fmname)
prettyxTime(ax)
fmname = 'BC37'
df = basinDryWeather[fmname]['Weekday']['DataFrame']
fig, ax = plt.subplots(figsize = (7.5,2))
df.plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:light grey')
df.mean(axis=1).plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:leaf green')
ax.set_title(fmname)
prettyxTime(ax)
fmname = 'BC44'
df = basinDryWeather[fmname]['Weekday']['DataFrame']
fig, ax = plt.subplots(figsize = (7.5,2))
df.plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:light grey')
df.mean(axis=1).plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:leaf green')
ax.set_title(fmname)
prettyxTime(ax)
fmname = 'BC38'
df = basinDryWeather[fmname]['Weekday']['DataFrame']
fig, ax = plt.subplots(figsize = (7.5,2))
df.plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:light grey')
df.mean(axis=1).plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:leaf green')
ax.set_title(fmname)
prettyxTime(ax)
# +
fmname = 'BC32'
df = basinDryWeather[fmname]['Weekday']['DataFrame']
ens_mean = df.mean(axis = 1)
ens_std = df.std(axis = 1)
df2 = df.copy()
df2[df < (ens_mean - 2 * ens_std)] = float('NaN')
df2[df > (ens_mean + 2 * ens_std)] = float('NaN')
new_mean = df2.mean(axis = 1)
fmname = 'BC32'
df = basinDryWeather[fmname]['Weekday']['DataFrame']
fig, ax = plt.subplots(figsize = (7.5,2))
df.plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:light grey')
df.mean(axis=1).plot(ax = ax,
kind = 'line',
legend = False,
color = 'xkcd:leaf green')
ax.set_title(fmname)
prettyxTime(ax)
(ens_mean - 2 * ens_std).plot(ax = ax,
kind = 'line',
color = 'xkcd:scarlet')
(ens_mean + 2 * ens_std).plot(ax = ax,
kind = 'line',
color = 'xkcd:scarlet')
df2.mean(axis=1).plot(ax = ax,
kind = 'line',
linestyle = ':',
legend = False,
color = 'xkcd:charcoal')
fig, ax = plt.subplots(figsize = (7.5,2))
df.plot(ax = ax,
kind = 'line',
marker = '.',
legend = False,
linewidth = 0,
color = 'xkcd:light grey')
df2[df < (ens_mean - 2 * ens_std)].plot(
ax = ax,
kind = 'line',
marker = '.',
legend = False,
linewidth = 0,
color = 'xkcd:scarlet')
prettyxTime(ax)
# -
def readFMdetails(filename):
#column names: 'Rain Gage', 'Diameter', 'Linear Feet', 'Basin Area (Ac)', 'Basin Footprint (in-mi)', 'Total Footage (LF)'
df = pd.read_csv(filename,
index_col=0,
dtype = {
'Flow Monitor' : 'object',
'Rain Gage' : 'object',
'Diameter' : np.float64,
'Linear Feet' : np.float64,
'Basin Area (Ac)' : np.float64,
'Bassin Footprint (in-mi)' : np.float64,
'Total Footage (LF)' : np.float64
})
df[df==9999.0] = float('NaN')
return(df)
# ADDITIONAL CALCULATIONS
# find Qc for each flow monitor pipe
dfmDetails = readFMdetails(
filename=fmdataFile)
diameters = dfmDetails.loc[:, 'Diameter']
dfmDetails.head()
def circular_manningsEQ(k, n, D, D_units, S):
convToFeet = {
'in' : 1/12,
'ft' : 1,
'mm' : 1/304.8,
'cm' : 1/30.48
}
D = D * convToFeet[D_units]
A = math.pi() * D**2/4
P = math.pi() * D
Rh = A/P
Q = (k/n) * A * Rh**(2/3) * S**0.5
return(Q)
# find 5 biggest storms
# +
# KPI HEATMAP
# +
# BOXPLOT Q/Qc
# +
# STORM DATES HORZ BAR CHART
# +
# Q vs i VERT BAR CHART
# +
# NET I&I HEATMAP (5 biggest storms)
# +
# BULLET GRAPHS - performance metrics
# d/D dry (%) - use median
# d/D wet (%) - use median
# Base infiltration (%)
# RDII (gal/in/mile)? - use mean
# Capture Coefficient (%) - use mean
fmname = 'BC38'
def constructDict(keyList, keyVals, dictionary = {}):
for idx, key in enumerate(keyList):
dictionary[key] = keyVals[idx]
return(dictionary)
def bulletGraph_fms(fmname, basinDryWeather, stormDict, saveDir = []):
# light yellow, light orange, red-orange
palette = ['#fed98e', '#fe9929', '#cc4c02']
metrics = ['d/D Dry', 'd/D Wet', 'Base Infil.','C Coeff']
limits = [[0.3,0.45,1], [0.5,0.65,1], [0.25,0.5,1], [0.05,0.1,1]]
labels = ['Good', 'OK', 'Poor']
# construct limit dictionary
limitDict = constructDict(
keyList = metrics,
keyVals = limits)
#find that data
dD_dry = min(np.quantile(
basinDryWeather[fmname]['Overall']['d/D Dry'],
0.95),
1)
dD_wet = min(np.quantile(
basinDryWeather[fmname]['Overall']['d/D Wet'],
0.95),
1)
baseInfil = basinDryWeather[fmname]['Overall']['Base Infiltration']
#RDII = stormDict[fmname]['RDII'].mean()
cc = stormDict[fmname]['Capt Coeff'].mean()
vals = [dD_dry, dD_wet, baseInfil, cc]
fig, axarr = plt.subplots(
nrows = len(metrics),
ncols = 1,
sharex = True,
figsize = (5,2))
for metricIdx, metric in enumerate(metrics):
h = limitDict[metric][-1] / 10
ax = axarr[metricIdx]
# format
ax.set_aspect('equal')
ax.set_yticks([1])
ax.set_yticklabels([metric])
ax.spines['bottom'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
prev_limit = 0
for limIdx, lim in enumerate(limitDict[metric]):
ax.barh([1],
lim - prev_limit,
left = prev_limit,
height = h,
color = palette[limIdx])
prev_limit = lim
ax.barh([1], vals[metricIdx], color = 'xkcd:chocolate', height = h / 3)
if not saveDir:
plt.show()
else:
plt.tight_layout()
saveName = saveDir + '\\' + fmname + '\\' + fmname + '_bullet.png'
plt.savefig(saveName)
plt.close(fig)
fmname = 'BC70A'
bulletGraph_fms(
fmname = fmname,
basinDryWeather = basinDryWeather,
stormDict = stormDict)
# +
# DRY WEATHER BOXPLOTS (weekday vs weekend)
fmname = 'BC01'
def removeNans(ser):
return(ser.values[~np.isnan(ser.values)])
# SET FONT
font = {'family' : 'DejaVu Sans',
'weight' : 'normal',
'size' : 12}
matplotlib.rc('font', **font)
def dryBoxplots(data, ylabel, title, showyticks,
topLim, bottomLim, annotate,
saveDir = [], saveName = []):
fig,ax = plt.subplots(figsize=(2.2,1.75))
boxwidth = 0.3
bp = ax.boxplot(data,
labels = ['WKD','WKE'],
patch_artist = True,
showfliers = False,
widths = boxwidth,
whis = [5,95],
showcaps = False)
plt.setp(bp['boxes'],color = 'xkcd:clay', linewidth = 2.0)
plt.setp(bp['whiskers'],color = 'xkcd:clay', linewidth = 2.0)
plt.setp(bp['medians'],color = 'xkcd:charcoal', linewidth = 2.5)
plt.setp(bp['caps'],color = 'xkcd:clay', linewidth = 3)
# plot weekend and weekday differently
colors = ['xkcd:clay', 'xkcd:white']
for box, color in zip(bp['boxes'],colors):
box.set(facecolor = color)
ax.yaxis.grid(True,
linestyle = '-',
which = 'major',
color = 'xkcd:warm grey',
alpha = 0.5)
ax.set_ylim(top = topLim,
bottom = min(bottomLim,0))
ax.set_title(title)
if showyticks:
ax.set_ylabel(ylabel)
else:
plt.tick_params(
axis = 'y', # changes apply to the x-axis
which = 'both', # both major and minor ticks are affected
left = True, # ticks along the bottom edge are off
right = False, # ticks along the top edge are off
labelleft = False) # labels along the bottom edge are off
if annotate:
for values in data:
labelNums - np.quantile(values,[0.05,0.5,0.95])
if not saveDir:
plt.show()
else:
saveName = saveDir + '\\' + fmname + '\\' + fmname + '_' + saveName + '.png'
plt.savefig(saveName)
plt.close(fig)
plt.show()
# gross Q
grossQ_wkd = removeNans(basinDryWeather[fmname]['Weekday']['Gross Q'])
grossQ_wke = removeNans(basinDryWeather[fmname]['Weekend']['Gross Q'])
dryBoxplots(
data = [grossQ_wkd, grossQ_wke],
ylabel = 'Q (MGD)',
title = 'Gross Q',
showyticks = True,
topLim = round(1.5 * max(
np.quantile(grossQ_wkd,0.95),
np.quantile(grossQ_wke,0.95)),1),
bottomLim = round(1.2 * min(
np.quantile(grossQ_wkd,0.05),
np.quantile(grossQ_wke,0.05))),
annotate = False,
savedir = [],
savename = [])
# net Q
netQ_wkd = removeNans(basinDryWeather[fmname]['Weekday']['Net Q'])
netQ_wke = removeNans(basinDryWeather[fmname]['Weekend']['Net Q'])
dryBoxplots(
data = [netQ_wkd, netQ_wke],
ylabel = 'Q (MGD)',
title = 'Net Q',
showyticks = True,
topLim = round(1.5 * max(
np.quantile(netQ_wkd,0.95),
np.quantile(netQ_wke,0.95)),1),
bottomLim = round(1.2 * min(
np.quantile(netQ_wkd,0.05),
np.quantile(netQ_wke,0.05))),
annotate = False)
# d/D
dD_wkd = basinDryWeather[fmname]['Weekday']['d/D']
dD_wke = basinDryWeather[fmname]['Weekend']['d/D']
dryBoxplots(
data = [dD_wkd, dD_wke],
ylabel = 'd/D',
title = 'Dry Capacity',
showyticks = True,
topLim = 1,
bottomLim = 0,
annotate = False)
# +
def pltDryGrossQ(basinDryWeather,fmname,saveDir = []):
# gross Q
grossQ_wkd = removeNans(basinDryWeather[fmname]['Weekday']['Gross Q'])
grossQ_wke = removeNans(basinDryWeather[fmname]['Weekend']['Gross Q'])
dryBoxplots(
data = [grossQ_wkd, grossQ_wke],
ylabel = 'Q (MGD)',
title = 'Net Q',
showyticks = True,
topLim = round(1.5 * max(
np.quantile(netQ_wkd,0.95),
np.quantile(netQ_wke,0.95)),1),
bottomLim = round(1.2 * min(
np.quantile(netQ_wkd,0.05),
np.quantile(netQ_wke,0.05))),
annotate = False,
savedir = saveDir,
savename = 'grossQ')
def pltDryNetQ(basinDryWeather,fmname,saveDir = []):
# net Q
netQ_wkd = removeNans(basinDryWeather[fmname]['Weekday']['Net Q'])
netQ_wke = removeNans(basinDryWeather[fmname]['Weekend']['Net Q'])
dryBoxplots(
data = [netQ_wkd, netQ_wke],
ylabel = 'Q (MGD)',
title = 'Net Q',
showyticks = True,
topLim = round(1.5 * max(
np.quantile(netQ_wkd,0.95),
np.quantile(netQ_wke,0.95)),1),
bottomLim = round(1.2 * min(
np.quantile(netQ_wkd,0.05),
np.quantile(netQ_wke,0.05))),
annotate = False,
savedir = saveDir,
savename = 'netQ')
def pltDrydD(basinDryWeather,fmname,saveDir = []):
# d/D
dD_wkd = basinDryWeather[fmname]['Weekday']['d/D']
dD_wke = basinDryWeather[fmname]['Weekend']['d/D']
dryBoxplots(
data = [dD_wkd, dD_wke],
ylabel = 'd/D',
title = 'Dry Capacity',
showyticks = True,
topLim = 1,
bottomLim = 0,
annotate = False,
savedir = saveDir,
savename = 'dD')
# +
# NET I&I VERT BAR CHART
def netii_bar(data, topNum, yLims, saveDir = []):
df = data.copy()
# sort in descending order of net I&I
df.sort_values(
by='Net Vol',
ascending=False,
inplace=True)
netii = df['Net Vol']
# take the top number of storms, e.g., 20
ii = netii[:topNum]
ii = ii.sort_index(ascending = True)
# assign color
colors = []
for date in ii.index:
if ((date >= dt.datetime(date.year,5,1))
& (date < dt.datetime(date.year,10,15))):
color = 'xkcd:seafoam blue'
else:
color = 'xkcd:stormy blue'
colors.append(color)
# format the index
ii.index = ii.index.strftime('%b %d')
# plot
fig,ax = plt.subplots(figsize = (7.25,1.5))
barPlot = ii.plot.bar(
ax = ax,
color = colors)
#ax.xaxis.set_major_formatter(dates.DateFormatter('%b %d'))
ax.set_ylabel('Net I&I (MG)')
ax.set_yscale('log')
ax.set_ylim(top = yLims[1],
bottom = yLims[0])
ax.yaxis.grid(True,
linestyle = '-',
which = 'major',
color = 'xkcd:charcoal',
alpha = 0.4)
if not saveDir:
plt.show()
else:
saveName = saveDir + '\\' + fmname + '\\' + fmname + '_netIIVol_bar.png'
plt.savefig(saveName)
plt.close(fig)
fmname = 'BC01'
netii_bar(
data = stormDict[fmname],
topNum = 12,
yLims = (.01,50))
# -
# NET I&I vs i FITS (summer vs. winter)
def df_rainComp(stormDict, fmname, col, ylabel,
saveDir = [], fitData = {}, fit = True):
df = stormDict[fmname].copy()
df = df.loc[df.loc[:,col] > 0]
colors = []
summer_ii = []
summer_rain = []
winter_ii = []
winter_rain = []
for date, value, rain in zip(
df.index,
df[col].values,
df['Storm Rain'].values):
if ((date >= dt.datetime(date.year,5,1))
& (date < dt.datetime(date.year,10,15))):
color = 'xkcd:seafoam blue'
summer_ii.append(value)
summer_rain.append(rain)
else:
color = 'xkcd:stormy blue'
winter_ii.append(value)
winter_rain.append(rain)
colors.append(color)
fig,ax = plt.subplots(figsize = (7.25,1.5))
# plot data points
ax.scatter(
x = df['Storm Rain'].values,
y = df[col].values,
c = colors,
alpha = 0.8)
ax.set_yscale('linear')
ax.set_ylabel(ylabel)
topLim = round(1.2 * max(df[col].values),1)
ax.set_ylim(top = topLim,
bottom = 0)
ax.set_xscale('linear')
ax.set_xlabel('Rain (in)')
rightLim = round(1.2 * max(df['Storm Rain'].values))
ax.set_xlim(right = rightLim,
left = 0)
if fit:
# summer fit
m_summer, b_summer, r_summer, p, err = stats.linregress(
x = summer_rain,
y = summer_ii)
# winter fit
m_winter, b_winter, r_winter, p, err = stats.linregress(
x = winter_rain,
y = winter_ii)
# update dictionary
fitData[fmname] = {
'Winter' : {
'slope' : m_winter,
'intercept' : b_winter,
'r-squared' : r_winter},
'Summer' : {
'slope' : m_summer,
'intercept' : b_summer,
'r-squared' : r_summer}
}
# plot fits
x = np.array([0,rightLim])
y_summer = m_summer * x + b_summer
y_winter = m_winter * x + b_winter
ax.plot(x_vals,
y_summer,
linewidth = 2.0,
linestyle = '-',
color = 'xkcd:seafoam blue',
label = 'summer')
ax.plot(x_vals,
y_winter,
linewidth = 2.0,
linestyle = '-',
color = 'xkcd:stormy blue',
label = 'winter')
ax.legend(loc = 'upper left')
if not saveDir:
plt.show()
else:
saveName = saveDir + '\\' + fmname + '\\' + fmname + '_netIIvsi.png'
plt.savefig(saveName)
plt.close(fig)
return(fitData)
fitDict = df_rainComp(
stormDict = stormDict,
fmname = 'BC01',
col = 'Net Vol',
ylabel = 'Net I&I (MG)',
fit = True)
# +
fmname = 'RSPSM'
netQ_wkd = removeNans(basinDryWeather[fmname]['Weekday']['Net Q'])
data = netQ_wkd
maxBinEdge = 1.1 * round(data.max(),2)
minBinEdge = np.min(1.1 * round(data.min(),2), 0)
nbins = 20
binEdges = np.linspace(minBinEdge, maxBinEdge, nbins)
fig,ax = plt.subplots()
cc_hist = ax.hist(x = netQ_wkd,
bins = binEdges,
facecolor = 'xkcd:light grey',
edgecolor = 'xkcd:charcoal',
density = True,
cumulative = True,
align = 'right')
ax.yaxis.grid(True,
linestyle = '-',
which = 'major',
color = 'xkcd:charcoal',
alpha = 0.4)
ax.set_xlabel('Net Q (MGD)')
ax.set_title(fmname)
ax.set_ylabel('% Less Than')
for patch, binEdge in zip(cc_hist[2],cc_hist[1][1:]):
if binEdge < 0:
patch.set_fc(color = 'xkcd:cornflower')
print(netQ_wkd.mean())
# -
colors =
# +
# UPSTREAM DRY WEATHER MEANS, turn into a function
def prettyxTime(ax):
ticks = ax.get_xticks()
ax.set_xticks(np.linspace(ticks[0],24*3600,5))
ax.set_xticks(np.linspace(ticks[0],24*3600,25),minor=True)
bc19a = basinDryWeather['BC19A']['Weekday']['DataFrame'].mean(axis = 1)
bc70a = basinDryWeather['BC70A']['Weekday']['DataFrame'].mean(axis = 1)
bc20m = basinDryWeather['BC20M']['Weekday']['DataFrame'].mean(axis = 1)
bc32 = basinDryWeather['BC32']['Weekday']['DataFrame'].mean(axis = 1)
bc65 = basinDryWeather['BC65']['Weekday']['DataFrame'].mean(axis = 1)
fig, ax = plt.subplots(figsize = (7.5,3))
df_up = pd.DataFrame({
'BC20M' : bc20m,
'BC32' : bc32,
'BC65' : bc65,
}, index = bc19a.index)
df_up.plot.area(ax = ax,
stacked = True,
color = ['#deebf7','#9ecae1','#3182bd'])
ax.set_ylabel('Q (MGD)')
ax.set_xlabel('Time of Day')
prettyxTime(ax)
df_down = pd.DataFrame(data = {'BC19A' : bc19a})
df_down.plot(kind = 'line',
color = 'xkcd:charcoal',
linestyle = ':',
ax = ax)
ax.set_ylim(top = 1.1 * bc19a.max())
ax.legend(loc = 'lower right')
# +
def readUpstreamFile(filename):
df = pd.read_csv(filename,
index_col=0)
return(df)
def findUpstreamFMs(df, fmname):
usfms = df.loc[fmname, 'USFM']
if usfms=='None':
usfms = [] #return an empty list
else:
usfms = usfms.split(',') # return the list of upstream flow monitors
return(usfms)
def prettyxTime(ax):
ticks = ax.get_xticks()
ax.set_xticks(np.linspace(ticks[0],24*3600,5))
ax.set_xticks(np.linspace(ticks[0],24*3600,25),minor=True)
upstreamFile = homeDir + '\\FMtoUpstream.csv'
dfUpstream = readUpstreamFile(
filename = upstreamFile)
def plotUpstreamFlows(fmname, basinDryWeather, usfmList, saveDir = []):
fmMean = basinDryWeather[fmname]['Weekday']['DataFrame'].mean(axis = 1)
df_down = pd.DataFrame(
data = {fmname: fmMean},
index = fmMean.index)
data = {}
colors = seaborn.color_palette('Set2',
len(usfmList))[::-1]
for usfm in usfmList:
data[usfm] = basinDryWeather[usfm]['Weekday']['DataFrame'].mean(axis = 1)
df_up = pd.DataFrame(data = data, index = fmMean.index)
# plot
fig, ax = plt.subplots(figsize = (8.5,4))
df_up.plot.area(ax = ax,
stacked = True,
color = colors)
ax.set_ylabel('Q (MGD)')
prettyxTime(ax)
df_down.plot(kind = 'line',
color = 'xkcd:charcoal',
linestyle = ':',
linewidth = 2.0,
ax = ax)
ax.set_ylim(top = 1.2 * fmMean.max())
ax.set_xlabel('Time of Day')
ax.legend(loc = 'lower right')
if not saveDir:
plt.show()
else:
saveName = saveDir + '\\' + fmname + '\\' + fmname + '_wUpstream.png'
plt.savefig(saveName)
plt.close(fig)
plotUpstreamFlows(fmname = '20M',
basinDryWeather = basinDryWeather,
dfUpstream = dfUpstream)
# -
def findTextFiles(readDir):
d = []
f = []
t = []
c = []
for (root, dirs, files) in walk(readDir, topdown=True):
d.extend(dirs)
f.extend(files)
for x in f:
if x.endswith('.txt'):
t.extend([x])
elif x.endswith('csv'):
c.extend([x])
d = sorted(d)
t = sorted(t)
c = sorted(c)
return(d, t, c)
# +
import pickle
import numpy as np
from os import makedirs
from os import walk
import pandas as pd
import datetime as dt
import scipy.stats as stats
import matplotlib.pyplot as plt
import matplotlib
homeDir = 'P:\\PW-WATER SERVICES\\TECHNICAL SERVICES\\Anna'
pickleLocation = homeDir + '\\2018\\Python Objects\\'
fmdataFile = homeDir + '\\FMdata.csv'
saveDir = homeDir + '\\2018\\Yearly Report'
upstreamFile = homeDir + '\\FMtoUpstream.csv'
# SET FONT
font = {'family' : 'DejaVu Sans',
'weight' : 'normal',
'size' : 12}
matplotlib.rc('font', **font)
dfUpstream = readUpstreamFile(
filename = upstreamFile)
folders, txt, csv = findTextFiles(saveDir)
# +
'''
basinDryWeather is a dictionary with flow monitors names as keys that
contain the following dictionaries:
* 'Weekday' : {
'd/D' : a numpy array
'Gross Q' : a pandas series (index: datetime, values: Q)
'DataFrame' : pandas data frame (index:time, columns: date,
values: q)
'Gross Diurnal' : pandas series (index: time, values: Q)
'Net Diurnal' : pandas series (index: time, values: Q)
}
* 'Weekend' : {
'd/D' : a numpy array
'Gross Q' :
'DataFrame' : pandas data frame (index:time, columns: date,
values: q)
'Gross Diurnal' : pandas series (index: time, values: Q)
'Net Diurnal' : pandas series (index: time, values: Q)
}
* 'Overall' : {
'd/D' : a numpy array
'Base Infiltration' : numpy float
}'''
with open(pickleLocation + 'basinDryWeather.pickle', 'rb') as handle:
basinDryWeather = pickle.load(handle)
'''
stormsDict is a dictionary with flow monitors as keys that contain a pandas
dataframe, dfStorms, with storm start times as indices and the following
columns:
* Storm RT : storm rain total in inches
* Storm Dur : storm duration in hours
* Event RT : event rain total in inches
* Event Dur : event duration in hours
* Gross Vol: gross I&I volume in MG
* Net Vol : net I&I volume in MG
* Capt Coeff : capture coefficient (vol I&I/vol rainfall)
* RDII : rain dependent I&I ranking (gal/in(rain)/in-mi(pipe))'''
with open(pickleLocation + 'stormsDict.pickle', 'rb') as handle:
stormDict = pickle.load(handle)
'''gageStorms is a dictionary with rain gages as keys that contain a pandas
dataframe, dfStorms, with storm start tiems as indices and the following
columns:
* Storm RT : storm rain total in inches
* Storm Dur : storm duration in hours
* Event RT : event rain total in inches
* Event Dur : event duration in hours'''
with open(pickleLocation + 'gageStorms.pickle', 'rb') as handle:
gageStorms = pickle.load(handle)
# -
fitDict = {}
for fmname in stormDict:
if fmname not in folders:
#make the directory
makedirs(saveDir + "\\" + fmname)
bulletGraph_fms(
fmname = fmname,
basinDryWeather = basinDryWeather,
stormDict = stormDict,
saveDir = saveDir)
pltDryGrossQ(
basinDryWeather = basinDryWeather,
fmname = fmname,
saveDir = saveDir)
pltDryNetQ(
basinDryWeather = basinDryWeather,
fmname = fmname,
saveDir = saveDir)
pltDrydD(
basinDryWeather = basinDryWeather,
fmname = fmname,
saveDir = saveDir)
netii_bar(
data = stormDict[fmname],
topNum = 12,
yLims = (.01,50),
saveDir = saveDir)
fitDict[fmname] = df_rainComp(
stormDict = stormDict,
fmname = fmname,
col = 'Net Vol',
ylabel = 'Net I&I (MG)',
fit = True,
saveDir = saveDir,
fitDict = fitDict)
usfmList = findUpstreamFMs(
df = dfUpstream,
fmname = fmname)
if not usfmList:
pass
else:
plotUpstreamFlows(
fmname = fmname,
basinDryWeather = basinDryWeather,
usfmList,
saveDir = saveDir)
| .ipynb_checkpoints/yearly reports-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Searching the UniProt database and saving fastas:
#
# This notebook is really just to demonstrate how Andrew finds the sequences for the datasets. <br>
#
# If you do call it from within our github repository, you'll probably want to add the fastas to the `.gitignore` file.
# Import bioservices module, to run remote UniProt queries
# (will probably need to pip install this to use)
from bioservices import UniProt
# ## Connecting to UniProt using bioservices:
#
service = UniProt()
fasta_path = 'refined_query_fastas/' #optional file organization param
# ## Query with signal_peptide
# +
def data_saving_function_with_SP(organism,save_path=''):
secreted_query = f'(((organism:{organism} OR host:{organism}) annotation:("signal peptide") keyword:secreted) NOT annotation:(type:transmem)) AND reviewed:yes'
secreted_result = service.search(secreted_query, frmt="fasta")
secreted_outfile = f'{save_path}{organism}_secreted_SP_new.fasta'
with open(secreted_outfile, 'a') as ofh:
ofh.write(secreted_result)
cytoplasm_query = f'(((organism:{organism} OR host:{organism}) locations:(location:cytoplasm)) NOT (annotation:(type:transmem) OR annotation:("signal peptide"))) AND reviewed:yes'
cytoplasm_result = service.search(cytoplasm_query, frmt="fasta")
cytoplasm_outfile = f'{save_path}{organism}_cytoplasm_SP_new.fasta'
with open(cytoplasm_outfile, 'a') as ofh:
ofh.write(cytoplasm_result)
membrane_query = f'(((organism:{organism} OR host:{organism}) annotation:(type:transmem)) annotation:("signal peptide")) AND reviewed:yes'
membrane_result = service.search(membrane_query, frmt="fasta")
membrane_outfile = f'{save_path}{organism}_membrane_SP_new.fasta'
with open(membrane_outfile, 'a') as ofh:
ofh.write(membrane_result)
# -
data_saving_function_with_SP('human',fasta_path)
data_saving_function_with_SP('escherichia',fasta_path)
# ## Query without signal_peptide
# +
def data_saving_function_without_SP(organism,save_path=''):
# maybe new:
secreted_query = f'(((organism:{organism} OR host:{organism}) AND (keyword:secreted OR goa:("extracellular region [5576]"))) NOT (annotation:(type:transmem) OR goa:("membrane [16020]") OR locations:(location:cytoplasm) OR goa:("cytoplasm [5737]") )) AND reviewed:yes'
secreted_result = service.search(secreted_query, frmt="fasta")
secreted_outfile = f'{save_path}{organism}_secreted_noSP_new_new.fasta'
with open(secreted_outfile, 'a') as ofh:
ofh.write(secreted_result)
cytoplasm_query = f'(((organism:{organism} OR host:{organism}) AND (locations:(location:cytoplasm) OR goa:("cytoplasm [5737]")) ) NOT (annotation:(type:transmem) OR goa:("membrane [16020]") OR keyword:secreted OR goa:("extracellular region [5576]") )) AND reviewed:yes'
cytoplasm_result = service.search(cytoplasm_query, frmt="fasta")
cytoplasm_outfile = f'{save_path}{organism}_cytoplasm_noSP_new_new.fasta'
with open(cytoplasm_outfile, 'a') as ofh:
ofh.write(cytoplasm_result)
membrane_query= f'(((organism:{organism} OR host:{organism}) AND ( annotation:(type:transmem) OR goa:("membrane [16020]") )) NOT ( keyword:secreted OR goa:("extracellular region [5576]") OR locations:(location:cytoplasm) OR goa:("cytoplasm [5737]") )) AND reviewed:yes'
membrane_result = service.search(membrane_query, frmt="fasta")
membrane_outfile = f'{save_path}{organism}_membrane_noSP_new_new.fasta'
with open(membrane_outfile, 'a') as ofh:
ofh.write(membrane_result)
# -
data_saving_function_without_SP('human',fasta_path)
data_saving_function_without_SP('yeast',fasta_path)
data_saving_function_without_SP('escherichia',fasta_path)
# ## Query ALL SHIT (warning: do not do unless you have lots of free time and computer memory)
# +
def data_saving_function_without_SP_full_uniprot(save_path=''):
# maybe new:
secreted_query = f'((keyword:secreted OR goa:("extracellular region [5576]")) NOT (annotation:(type:transmem) OR goa:("membrane [16020]") OR locations:(location:cytoplasm) OR goa:("cytoplasm [5737]") )) AND reviewed:yes'
secreted_result = service.search(secreted_query, frmt="fasta")
secreted_outfile = f'{save_path}all_secreted_noSP_new_new.fasta'
with open(secreted_outfile, 'a') as ofh:
ofh.write(secreted_result)
cytoplasm_query = f'(( locations:(location:cytoplasm) OR goa:("cytoplasm [5737]") ) NOT (annotation:(type:transmem) OR goa:("membrane [16020]") OR keyword:secreted OR goa:("extracellular region [5576]") )) AND reviewed:yes'
cytoplasm_result = service.search(cytoplasm_query, frmt="fasta")
cytoplasm_outfile = f'{save_path}all_cytoplasm_noSP_new_new.fasta'
with open(cytoplasm_outfile, 'a') as ofh:
ofh.write(cytoplasm_result)
membrane_query= f'(( annotation:(type:transmem) OR goa:("membrane [16020]") ) NOT ( keyword:secreted OR goa:("extracellular region [5576]") OR locations:(location:cytoplasm) OR goa:("cytoplasm [5737]") )) AND reviewed:yes'
membrane_result = service.search(membrane_query, frmt="fasta")
membrane_outfile = f'{save_path}all_membrane_noSP_new_new.fasta'
with open(membrane_outfile, 'a') as ofh:
ofh.write(membrane_result)
# -
data_saving_function_without_SP_full_uniprot(fasta_path)
| notebooks/UniProt_refined_query.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import sys
from pathlib import Path
sys.path.append(str(Path.cwd().parent))
# -
from load_dataset import Dataset
# +
# Dataset класс, принимающий на вход директорию, содержащую csv файлы формата index, value, позволяющий удобно
# оперировать с большим количеством временных рядов.
# -
dataset = Dataset('../data/dataset/')
# метод __repr__ (именно он вызывается, когда мы запускаем ячейку) показывает все ряды из датасета
dataset
# чтобы достать временной ряд, можно просто использовать subscriptable нотацию
dataset["day_2690.csv"]
# +
# dataset является iterable обьектом, на каждом шаге возвращается tuple из имени ряда (имя файла в папке)
# и самого временного ряда, обьекта pd.Series()
# -
for key, ts in dataset:
print(key)
# +
# чтобы выгрузить весь в оперативную память, можно использовать метод load
# -
dataset.load()
| week_1/dataset_usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="sU6UjIJbl9Qe"
# #Introduction
# + [markdown] id="_YeS8fbTmCHz"
# Linear algebra is a field of mathematics that is widely used in various disciplines. Linear algebra plays an important role in data science and machine
# learning. A solid understanding of linear algebra concepts can enhance the
# understanding of many data science and machine learning algorithms. This
# chapter introduces basic concepts for data science and includes vector spaces,
# orthogonality, eigenvalues, matrix decomposition and further expanded to include linear regression and principal component analysis where linear algebra
# plays a central role for solving data science problems. More advanced concepts
# and applications of linear algebra can be found in many references $[1, 2, 3, 4]$.
# + [markdown] id="BOtMuUHAl9Xb"
# #Elements of Linear Algebra
#
# + [markdown] id="K9xLEBQRmqlK"
# ##Linear Spaces
# + [markdown] id="XJ58jcOamqe7"
# ###Linear Combinations
# + colab={"base_uri": "https://localhost:8080/"} id="9u4AmhBgoojI" outputId="1475e474-dfa4-4689-a05d-390b8f1cb645"
import numpy as np
v = np.array([[3.3],[11]])
w = np.array([[-2],[-40]])
a = 1.5
b = 4.9
u = a*v + b*w
u
# + colab={"base_uri": "https://localhost:8080/"} id="SsZ-jHdGnLml" outputId="6bfc2db6-7dde-421e-af49-62f1f75209ae"
import numpy as np
x = np.array([[4, 8, 1],
[2, 11, 0],
[1, 7.4, 0.2]])
y = ([3.65, 1.55, 3.42])
result = np.linalg.solve(x, y)
result
# + [markdown] id="nV43pejOmqOA"
# ###Linear Independence and Dimension
#
# + colab={"base_uri": "https://localhost:8080/"} id="H_i1l1ExnMZn" outputId="48a0d109-8b12-4e80-ab15-62da91500118"
import sympy
import numpy as np
matrix = np.array([[2,1,7],[2,9,11]])
_, inds = sympy.Matrix(matrix).T.rref()
print(inds)
# + [markdown] id="-lZ2rcwts2fC"
# This means vetor 0 and vector 1 are linearly independent.
# + colab={"base_uri": "https://localhost:8080/"} id="E8INoUvuss3L" outputId="77fcc50c-db35-45dd-89ab-aa82c951d428"
matrix = np.array([[0,1,0,0],[0,0,1,0],[0,1,1,0],[1,0,0,1]])
_, inds = sympy.Matrix(matrix).T.rref()
print(inds)
# + [markdown] id="b5dIh30Hsblj"
# This means vetor 0, vector 1, and vector 3 are lineatly independent while vector 2 is linearly dependent.
# + [markdown] id="B7D5pfd4tHhJ"
# ##Orthogonality
# + [markdown] id="c5lGX8m1vFcq"
# Inner product
# + colab={"base_uri": "https://localhost:8080/"} id="5PL7oE51ue6c" outputId="bee29261-8bcc-4b47-e5a6-3bdee3454879"
a = np.array([1.6,2.5,3.9])
b = np.array([4,1,11])
np.inner(a, b)
# + [markdown] id="g2zseWBcvRk2"
# Norm
# + colab={"base_uri": "https://localhost:8080/"} id="sbrhNUS4vOmr" outputId="99c253d3-120f-40c8-8682-204d765673db"
from numpy import linalg as LA
c = np.array([[ 1.3, -7.2, 12.1],
[-1, 0, 4]])
LA.norm(c)
# + [markdown] id="vYkf2gcnyS1e"
# Orthogonality
# + colab={"base_uri": "https://localhost:8080/"} id="uuizTL3xwrla" outputId="8cf2766c-b383-4d1f-d3b0-97e134c0606e"
v1 = np.array([1,-2, 4])
v2 = np.array([2, 5, 2])
dot_product = np.dot(v1,v2)
if dot_product == 0:
print('v1 and v2 are orthorgonal')
else: print('v1 and v2 are not orthorgonal')
# + colab={"base_uri": "https://localhost:8080/"} id="q-_6IqAKxJIH" outputId="7b4ed4e7-e22c-4703-eccf-3cefcbb1e019"
n1 = LA.norm(v1)
n2 = LA.norm(v2)
if n1 == 1 and n2 == 1:
print('v1 and v2 are orthornormal')
else: print('v1 and v2 are not orthornormal')
# + [markdown] id="K4c_1y4stL5r"
# ##Gram-Schmidt Process
# + id="1qZRJcYNyUrh"
import numpy as np
def gs(X):
Q, R = np.linalg.qr(X)
return Q
# + colab={"base_uri": "https://localhost:8080/"} id="tZqg1DRQ_2Mu" outputId="862c299d-36f0-421e-8338-3b2568aab10e"
X = np.array([[3,-1,0],[1.8,11.3,-7.5], [4,13/4,-7/3]])
gs(X)
# + [markdown] id="bsADBIPktMQr"
# ##Eigenvalues and Eigenvectors
# + colab={"base_uri": "https://localhost:8080/"} id="p4PGgISABCpe" outputId="91d0aa1d-fea4-4947-e506-6cd99405c403"
import numpy as np
from numpy.linalg import eig
a = np.array([[2.1, -5/2, 11.4],
[1, 3, 5],
[2.4, 3.5, 7.4]])
u,v=eig(a)
print('E-value:', u)
print('E-vector', v)
# + [markdown] id="n_3ZcgCMmRBQ"
# #Linear Regression
# + [markdown] id="vzIOY0AetuC3"
# ##QR Decomposition
# + colab={"base_uri": "https://localhost:8080/"} id="tggLlHF4CGJi" outputId="73bf7922-2f96-4b66-8859-fb2d1edf620d"
import numpy as np
from numpy.linalg import qr
m = np.array([[1/2, -2.8, 5/3],
[2.5, 3, 9],
[8.3, 4, -5.2]])
q, r = qr(m)
print('Q:', q)
print('R:', r)
n = np.dot(q, r)
print('QR:', n)
# + [markdown] id="OoGOn6AGtyPA"
# ##Least-squares Problems
#
# + [markdown] id="zKieKaePDA5V"
# Use direct inverse method
# + colab={"base_uri": "https://localhost:8080/"} id="VRYB6hQ1DHym" outputId="5d0a38c4-6ac3-4eca-c4c9-775fc6474597"
import numpy as np
from scipy import optimize
import matplotlib.pyplot as plt
plt.style.use('seaborn-poster')
x = np.linspace(0, 10, 500)
y = 1/2 + x * np.random.random(len(x))
A = np.vstack([x, np.ones(len(x))]).T
y = y[:, np.newaxis]
lst_sqr = np.dot((np.dot(np.linalg.inv(np.dot(A.T,A)),A.T)),y)
print(lst_sqr)
# + [markdown] id="V5zeLAuZE2GO"
# Use the pseudoinverse
# + colab={"base_uri": "https://localhost:8080/"} id="PIUOaBbiD8fC" outputId="fe7e519d-b99a-4360-8b8f-f5770caa2f69"
pinv = np.linalg.pinv(A)
lst_sqr = pinv.dot(y)
print(lst_sqr)
# + [markdown] id="gSRDrRF4GB8O"
# Use numpy.linalg.lstsq
# + colab={"base_uri": "https://localhost:8080/"} id="PYQCcJD8FL76" outputId="75715e92-d719-4f82-fad3-0fabdd363d0a"
lst_sqr = np.linalg.lstsq(A, y, rcond=None)[0]
print(lst_sqr)
# + [markdown] id="AWTlyMUCF9bF"
# Use optimize.curve_fit from scipy
# + colab={"base_uri": "https://localhost:8080/"} id="F88SGNvxFiT5" outputId="e123a98b-7e42-4878-9f13-64678a74ad85"
x = np.linspace(0, 10, 500)
y = 1/2 + x * np.random.random(len(x))
def func(x, a, b):
y = a*x + b
return y
lst_sqr = optimize.curve_fit(func, xdata = x, ydata = y)[0]
print(lst_sqr)
# + [markdown] id="Sr_eosHEt1J_"
# ## Linear Regression
#
# + colab={"base_uri": "https://localhost:8080/"} id="YR_cZkifJE0M" outputId="1dfe38c9-0ca3-467e-ff83-66cb1fdb25e0"
import numpy as np
from sklearn.linear_model import LinearRegression
x = np.array([5.3, 15.2, 25.8, 35.4, 45.5, 54.9]).reshape((-1, 1))
y = np.array([4.7, 20.4, 31/2, 33.2, 22, 38.6])
model = LinearRegression().fit(x, y)
r_sq = model.score(x, y)
print('coefficient of determination:', r_sq)
# + colab={"base_uri": "https://localhost:8080/"} id="nDlD5lNkJGEx" outputId="268885d1-5a88-441e-dd93-f283c7182d1c"
print('intercept:', model.intercept_)
# + colab={"base_uri": "https://localhost:8080/"} id="CrgIWCAaJO6K" outputId="ddcddd62-7e5e-4e7e-ded8-1c8a7b2d2123"
print('slope:', model.coef_)
# + colab={"base_uri": "https://localhost:8080/"} id="XxwcQ-93JlE4" outputId="1c821918-c343-4927-dd63-1d8652469fc7"
y_pred = model.predict(x)
print('predicted response:', y_pred, sep='\n')
# + [markdown] id="-wX2BTummTGA"
# #Principal Component Analysis
# + [markdown] id="vac8GQN4uBRP"
# ##Singular Value Decomposition
# + colab={"base_uri": "https://localhost:8080/"} id="rb0tj0_uKKId" outputId="891c842f-039a-49a6-bfa2-9df42333d23b"
from numpy import array
from scipy.linalg import svd
A = array([[3, -2, 5],
[1, 0, -3],
[4, 6, -1]])
print('Matrix A:')
print(A)
U, sigma, VT = svd(A)
print('The m × m orthogonal matrix:')
print(U)
print('The m × n diagonal matrix:')
print(sigma)
print('The n × n orthogonal matrix:')
print(VT)
# + [markdown] id="q7pUxKAmuHum"
# ##Principal Component Analysis
# + [markdown] id="OPMtOY8UPwHj"
# Covariance Matrix
# + colab={"base_uri": "https://localhost:8080/"} id="TJtVKau7Vzn2" outputId="da716ad0-aceb-4853-a34a-0d878aca1940"
A = array([[3, -2, 5],
[1, 0, -3],
[4, 6, -1]])
covMatrix = np.cov(A,bias=True)
print('Covariance matrix of A:')
print(covMatrix)
# + [markdown] id="j2yOTfQyXLBr"
# Principal Component Analysis
# + id="97IBOZEPXOOT" colab={"base_uri": "https://localhost:8080/"} outputId="2b0e5cf4-f8b4-4423-ec01-957e19802ca7"
import numpy as np
from numpy.linalg import eig
X = np.random.randint(10,50,100).reshape(20,5)
X_meaned = X - np.mean(X , axis = 0)
covMatrix = np.cov(X_meaned, rowvar = False)
val,vec = eig(covMatrix)
s_index = np.argsort(val)[::-1]
s_val = val[s_index]
s_vec = vec[:,s_index]
n_components = 8
vec_sub = s_vec[:,0:n_components]
#Transform the data
X_reduced = np.dot(vec_sub.transpose(), X_meaned.transpose()).transpose()
X_reduced
# + [markdown] id="ygFegkIUWEhr"
# Total Variance
# + id="-0WYtXbEWFzK" colab={"base_uri": "https://localhost:8080/"} outputId="32398bf3-3e62-4898-c030-d9022ed36bf3"
B = covMatrix
total_var_matr = B.trace()
print('Total variance of A:')
print(total_var_matr)
| Module_D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use('ggplot')
df = pd.read_csv("newly_truncated_value.csv")
df['7'] = df['7'].replace(0.99998999999999993,0)
df.head()
df.shape
X = np.array(df.drop(['5','6','7'],1))
y = np.array(df['7'])
y = y.astype("int")
from sklearn.model_selection import train_test_split
# +
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.4,random_state=42, stratify=y)
# +
from sklearn.neighbors import KNeighborsClassifier
neighbors = np.arange(1,7)
train_accuracy =np.empty(len(neighbors))
test_accuracy = np.empty(len(neighbors))
for i,k in enumerate(neighbors):
#Setup a knn classifier with k neighbors
knn = KNeighborsClassifier(n_neighbors=k)
#Fit the model
knn.fit(X_train, y_train)
#Compute accuracy on the training set
train_accuracy[i] = knn.score(X_train, y_train)
#Compute accuracy on the test set
test_accuracy[i] = knn.score(X_test, y_test)
# +
#Generate plot
plt.title('k-NN Varying number of neighbors')
plt.plot(neighbors, test_accuracy, label='Testing Accuracy')
plt.plot(neighbors, train_accuracy, label='Training accuracy')
plt.legend()
plt.xlabel('Number of neighbors')
plt.ylabel('Accuracy')
plt.show()
# -
#Setup a knn classifier with k neighbors
knn = KNeighborsClassifier(n_neighbors=2)
# +
#Fit the model
knn.fit(X_train,y_train)
# +
#Get accuracy. Note: In case of classification algorithms score method represents accuracy.
knn.score(X_test,y_test)
# +
#import confusion_matrix
from sklearn.metrics import confusion_matrix
# +
#let us get the predictions using the classifier we had fit above
y_pred = knn.predict(X_test)
# -
confusion_matrix(y_test,y_pred)
# +
pd.crosstab(y_test, y_pred, rownames=['True'], colnames=['Predicted'], margins=True)
# +
#import classification_report
from sklearn.metrics import classification_report
# -
print(classification_report(y_test,y_pred))
# +
y_pred_proba = knn.predict_proba(X_test)[:,1]
# -
from sklearn.metrics import roc_curve
# +
fpr, tpr, thresholds = roc_curve(y_test, y_pred_proba)
# -
plt.plot([0,1],[0,1],'k--')
plt.plot(fpr,tpr, label='Knn')
plt.xlabel('fpr')
plt.ylabel('tpr')
plt.title('Knn(n_neighbors=7) ROC curve')
plt.show()
# +
#Area under ROC curve
from sklearn.metrics import roc_auc_score
roc_auc_score(y_test,y_pred_proba)
# +
#import GridSearchCV
from sklearn.model_selection import GridSearchCV
# +
#In case of classifier like knn the parameter to be tuned is n_neighbors
param_grid = {'n_neighbors':np.arange(1,50)}
# +
knn = KNeighborsClassifier()
knn_cv= GridSearchCV(knn,param_grid,cv=5)
knn_cv.fit(X,y)
# +
knn_cv.best_score_
# +
knn_cv.best_params_
# -
from pandas.plotting import parallel_coordinates
plt.figure(figsize=(15,10))
parallel_coordinates(df,"7")
plt.title('Parallel Coordinates Plot', fontsize=20, fontweight='bold')
plt.xlabel('Features', fontsize=15)
plt.ylabel('Features values', fontsize=15)
plt.legend(loc=1, prop={'size': 15}, frameon=True,shadow=True, facecolor="white", edgecolor="black")
plt.show()
# +
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
plt.figure()
sns.pairplot(df, hue = "7", height=3, markers=["o", "s"])
plt.show()
# -
| ExampleSets/actual_knn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dependencies
# + _kg_hide-input=true
import os, warnings, shutil
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from transformers import AutoTokenizer
from sklearn.model_selection import StratifiedKFold
SEED = 0
warnings.filterwarnings("ignore")
# -
# # Parameters
MAX_LEN = 192
tokenizer_path = 'jplu/tf-xlm-roberta-base'
# # Load data
# + _kg_hide-input=true
train_df = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/jigsaw-toxic-comment-train.csv",
usecols=['comment_text', 'toxic'])
train_df = pd.concat([train_df[['comment_text', 'toxic']].query('toxic==1'),
train_df[['comment_text', 'toxic']].query('toxic==0').sample(n=100000, random_state=SEED)
]).reset_index()
print('Train samples %d' % len(train_df))
display(train_df.head())
# -
# # Tokenizer
# + _kg_hide-output=true
tokenizer = AutoTokenizer.from_pretrained('jplu/tf-xlm-roberta-base', lowercase=True)
# -
# # Data generation sanity check
# + _kg_hide-input=true
for idx in range(5):
print('\nRow %d' % idx)
max_seq_len = 22
comment_text = train_df['comment_text'].loc[idx]
enc = tokenizer.encode_plus(comment_text, return_token_type_ids=False, pad_to_max_length=True, max_length=max_seq_len)
print('comment_text : "%s"' % comment_text)
print('input_ids : "%s"' % enc['input_ids'])
print('attention_mask: "%s"' % enc['attention_mask'])
assert len(enc['input_ids']) == len(enc['attention_mask']) == max_seq_len
# -
# # 5-Fold split
# + _kg_hide-input=true
folds = StratifiedKFold(n_splits=5, shuffle=True, random_state=SEED)
for fold_n, (train_idx, val_idx) in enumerate(folds.split(train_df, train_df['toxic'])):
print('Fold: %s, Train size: %s, Validation size %s' % (fold_n+1, len(train_idx), len(val_idx)))
train_df[('fold_%s' % str(fold_n+1))] = 0
train_df[('fold_%s' % str(fold_n+1))].loc[train_idx] = 'train'
train_df[('fold_%s' % str(fold_n+1))].loc[val_idx] = 'validation'
# -
# # Label distribution
# + _kg_hide-input=true
for fold_n in range(folds.n_splits):
fold_n += 1
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(18, 6))
fig.suptitle('Fold %s' % fold_n, fontsize=22)
sns.countplot(x="toxic", data=train_df[train_df[('fold_%s' % fold_n)] == 'train'], palette="GnBu_d", ax=ax1).set_title('Train')
sns.countplot(x="toxic", data=train_df[train_df[('fold_%s' % fold_n)] == 'validation'], palette="GnBu_d", ax=ax2).set_title('Validation')
sns.despine()
plt.show()
# -
# # Output 5-fold set
# + _kg_hide-input=true
train_df.to_csv('5-fold.csv', index=False)
display(train_df.head())
for fold_n in range(folds.n_splits):
fold_n += 1
base_path = 'fold_%d/' % fold_n
# Create dir
os.makedirs(base_path)
x_train = tokenizer.batch_encode_plus(train_df[train_df[('fold_%s' % fold_n)] == 'train']['comment_text'].values,
return_token_type_ids=False,
pad_to_max_length=True,
max_length=MAX_LEN)
x_train = np.array([np.array(x_train['input_ids']),
np.array(x_train['attention_mask'])])
x_valid = tokenizer.batch_encode_plus(train_df[train_df[('fold_%s' % fold_n)] == 'validation']['comment_text'].values,
return_token_type_ids=False,
pad_to_max_length=True,
max_length=MAX_LEN)
x_valid = np.array([np.array(x_valid['input_ids']),
np.array(x_valid['attention_mask'])])
y_train = train_df[train_df[('fold_%s' % fold_n)] == 'train']['toxic'].values
y_valid = train_df[train_df[('fold_%s' % fold_n)] == 'validation']['toxic'].values
np.save(base_path + 'x_train', np.asarray(x_train))
np.save(base_path + 'y_train', y_train)
np.save(base_path + 'x_valid', np.asarray(x_valid))
np.save(base_path + 'y_valid', y_valid)
print('\nFOLD: %d' % (fold_n))
print('x_train shape:', x_train.shape)
print('y_train shape:', y_train.shape)
print('x_valid shape:', x_valid.shape)
print('y_valid shape:', y_valid.shape)
# Compress logs dir
# !tar -cvzf fold_1.tar.gz fold_1
# !tar -cvzf fold_2.tar.gz fold_2
# !tar -cvzf fold_3.tar.gz fold_3
# !tar -cvzf fold_4.tar.gz fold_4
# !tar -cvzf fold_5.tar.gz fold_5
# Delete logs dir
shutil.rmtree('fold_1')
shutil.rmtree('fold_2')
shutil.rmtree('fold_3')
shutil.rmtree('fold_4')
shutil.rmtree('fold_5')
# -
# # Validation set
# + _kg_hide-input=true
valid_df = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/validation.csv", usecols=['comment_text', 'toxic', 'lang'])
display(valid_df.head())
x_valid = tokenizer.batch_encode_plus(valid_df['comment_text'].values,
return_token_type_ids=False,
pad_to_max_length=True,
max_length=MAX_LEN)
x_valid = np.array([np.array(x_valid['input_ids']),
np.array(x_valid['attention_mask'])])
y_valid = valid_df['toxic'].values
np.save('x_valid', np.asarray(x_valid))
np.save('y_valid', y_valid)
print('x_valid shape:', x_valid.shape)
print('y_valid shape:', y_valid.shape)
# -
# # Test set
# + _kg_hide-input=true
test_df = pd.read_csv("/kaggle/input/jigsaw-multilingual-toxic-comment-classification/test.csv", usecols=['content'])
display(test_df.head())
x_test = tokenizer.batch_encode_plus(test_df['content'].values,
return_token_type_ids=False,
pad_to_max_length=True,
max_length=MAX_LEN)
x_test = np.array([np.array(x_test['input_ids']),
np.array(x_test['attention_mask'])])
np.save('x_test', np.asarray(x_test))
print('x_test shape:', x_test.shape)
| Datasets/jigsaw-dataset-split-toxic-roberta-base-192.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Advanced Regular Expressions Lab
#
# Complete the following set of exercises to solidify your knowledge of regular expressions.
import re
# ### 1. Use a regular expression to find and extract all vowels in the following text.
text = "This is going to be a sentence with a good number of vowels in it."
pattern = '[aeiou]'
print(re.findall(pattern, text))
# ### 2. Use a regular expression to find and extract all occurrences and tenses (singular and plural) of the word "puppy" in the text below.
text = "The puppy saw all the rest of the puppies playing and wanted to join them. I saw this and wanted a puppy of my own!"
pattern = 'puppy|puppies'
print(re.findall(pattern, text))
# ### 3. Use a regular expression to find and extract all tenses (present and past) of the word "run" in the text below.
text = "I ran the relay race the only way I knew how to run it."
pattern = 'run|ran'
print(re.findall(pattern, text))
# ### 4. Use a regular expression to find and extract all words that begin with the letter "r" from the previous text.
pattern = 'r\w*'
print(re.findall(pattern, text))
# ### 5. Use a regular expression to find and substitute the letter "i" for the exclamation marks in the text below.
text = "Th!s !s a sentence w!th spec!al characters !n !t."
re.sub('!', 'i', text)
# ### 6. Use a regular expression to find and extract words longer than 4 characters in the text below.
text = "This sentence has words of varying lengths."
pattern = '[A-z]{4,}'
print(re.findall(pattern, text))
# ### 7. Use a regular expression to find and extract all occurrences of the letter "b", some letter(s), and then the letter "t" in the sentence below.
text = "I bet the robot couldn't beat the other bot with a bat, but instead it bit me."
pattern = 'b[A-z]+t'
print(re.findall(pattern, text))
# ### 8. Use a regular expression to find and extract all words that contain either "ea" or "eo" in them.
text = "During many of the peaks and troughs of history, the people living it didn't fully realize what was unfolding. But we all know we're navigating breathtaking history: Nearly every day could be — maybe will be — a book."
pattern = '\w*e[ao]\w*'
print(re.findall(pattern, text))
# ### 9. Use a regular expression to find and extract all the capitalized words in the text below individually.
text = "<NAME> and <NAME> walk into a bar."
pattern = '[A-Z]\w*'
print(re.findall(pattern, text))
# ### 10. Use a regular expression to find and extract all the sets of consecutive capitalized words in the text above.
pattern = '[A-Z]\w* [A-Z]\w*'
print(re.findall(pattern, text))
# ### 11. Use a regular expression to find and extract all the quotes from the text below.
#
# *Hint: This one is a little more complex than the single quote example in the lesson because there are multiple quotes in the text.*
text = 'Roosevelt says to Lincoln, "I will bet you $50 I can get the bartender to give me a free drink." Lincoln says, "I am in!"'
pattern = '\"(.*?)\"'
print(re.findall(pattern, text))
pattern = '\"' #'["]'
print(re.findall(pattern, text))
# ### 12. Use a regular expression to find and extract all the numbers from the text below.
text = "There were 30 students in the class. Of the 30 students, 14 were male and 16 were female. Only 10 students got A's on the exam."
pattern = '\d+'
print(re.findall(pattern, text))
# ### 13. Use a regular expression to find and extract all the social security numbers from the text below.
text = """
Henry's social security number is 876-93-2289 and his phone number is (847)789-0984.
Darlene's social security number is 098-32-5295 and her phone number is (987)222-0901.
"""
pattern = '\d\d\d-\d\d-\d\d\d\d'
print(re.findall(pattern, text))
pattern = '\d{3}-\d{2}-\d{4}'
print(re.findall(pattern, text))
# ### 14. Use a regular expression to find and extract all the phone numbers from the text below.
pattern = '\W\d\d\d\W\d\d\d-\d\d\d\d'
print(re.findall(pattern, text))
pattern = '\W\d{3}\W\d{3}-\d{4}'
print(re.findall(pattern, text))
# ### 15. Use a regular expression to find and extract all the formatted numbers (both social security and phone) from the text below.
pattern = '\d\d\d-\d\d-\d\d\d\d|\W\d\d\d\W\d\d\d-\d\d\d\d'
print(re.findall(pattern, text))
pattern = '\d{3}-\d{2}-\d{4}|\W\d{3}\W\d{3}-\d{4}'
print(re.findall(pattern, text))
| module-1/Advanced-Regex/your-code/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="1v2PmJNYxxTA" colab_type="code" colab={}
#Instalamos la libreria tflearn
# !pip install tflearn
# + id="1JbvrCSFyyIo" colab_type="code" colab={}
#Importamos todo lo que necesitamos de tflearn
import tflearn
from tflearn.layers.core import input_data, fully_connected
from tflearn.layers.estimator import regression
from tflearn.data_utils import to_categorical
#Importamos el dataset y la funcion para separar el dataset en entrenamiento y prueba
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
# + id="p07Lyvt7zRch" colab_type="code" colab={}
#Cargamos el dataset
breast_cancer = load_breast_cancer()
#Lo separamos en entrenamiento y prueba
X_train, X_test, Y_train, Y_test = train_test_split(breast_cancer.data, breast_cancer.target)
# + id="7WdqiJz26B0n" colab_type="code" colab={}
#Convertimos las variables Y_train y Y_test en cateforicas de dos clases
Y_train = to_categorical(Y_train, nb_classes=2)
Y_test = to_categorical(Y_test, nb_classes=2)
# + id="rLmIuPR9_I8f" colab_type="code" colab={}
#Definimos la estructura de la red neuronal
net = input_data(shape=[None, 30])
net = fully_connected(net, 32, activation='ReLU', regularizer='L2')
net = fully_connected(net, 32, activation='ReLU', regularizer='L2')
net = fully_connected(net, 2, activation='softmax')
net = regression(net, optimizer='sgd', learning_rate=0.001, loss='categorical_crossentropy')
model = tflearn.DNN(net)
# + id="o0TMjdf9An0v" colab_type="code" colab={}
#Entrenamos el modelo y lo validamos
model.fit(X_train,Y_train, validation_set=(X_test, Y_test), show_metric=True, n_epoch=100)
| DNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
#SVD
'''
Singular Value Decomposition
uses:
-get rid of redudant data (dimensionality reduction)
-speed up training on redundant data by removing redundancy
details:
-eigenvalues tell you which data is important and which is redundant
-going from n features to k features, where each k is a linear combinarion of some of the n features
-similar in use to PCA
-"things should be made as simple as possible, but not simpler"
-leverages eigendecomposition to find 'principal component vectors'
-eigenvalues are the 'singular values' of the matrix
-think of it as breakdown into scaling and rotation
math:
> start with matrix A (data)
> find V,S, U such that A=U*sum(*V.T) (U are eigenvectors, V are eigenvalues)
> mask off eigenvalues below some threshold, and then
use the remaining eigenvalues to recompose the image,
now in lower dimension! (lower matrix rank)
LIN ALG: -recall eigenvalue is roughly the "magnitude" of an eigenvector,
which tells us on which axis the transormation
described by the matrix hinges. So eigenvectors tell us how strongly
the transorfmation hinges in the direction given by the eigenvector.
'''
# +
#SVD example
import numpy as np
import scipy.linalg as SL
import matplotlib.pyplot as plt
Pxx = np.genfromtxt('mBtasJLD.txt')
U, s, Vh = SL.svd(Pxx, full_matrices=False)
assert np.allclose(Pxx, np.dot(U, np.dot(np.diag(s), Vh)))
s[2:] = 0
new_a = np.dot(U, np.dot(np.diag(s), Vh))
#sample of data
print(new_a)
#should be less data in the reduced matrix
print(np.mean(Pxx))
print(np.mean(new_a))
#full data
plt.plot(Pxx)
plt.title("Full")
plt.show()
#reduced
plt.plot(new_a)
plt.title("After SVD")
plt.show()
| svd/SVD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notes from Linked In Learning
# ## Become a Business Intelligence Specialist
# ### Learning Power BI Desktop
#
# - Display recommendations: 1440x900 or 1600x900
# - Need 8 GB of RAM or better for the best Power BI experience
# - Initially built as a series of add-ins for Excel
# - Power BI Desktop can acquire, model, visualize and share data
# - Models built in Excel can be imported and used in Power BI
# - To share reports with colleagues, upload from Power BI Desktop to Power BI Service
# - can explore data using natural language Q&A
# - create and view dashboards
# - works with Microsoft Dynamics 365 or Salesforce
#
# Free Power BI:
# - 1 GB data limit (10 GB for Pro)
# - Daily data refresh (hourly for Pro)
# - 10k rows/hr streaming data (1 million rows/hr for Pro)
# - Data via Power BI Service (Pro: direct interactive connection)
#
#
#
#
#
#
# ### Financial Forecasting with Big Data
#
# - project cash flows, product pricing, equipment failures, defaults at a bank, customer couponing, facility locations
# - using regression to make forecasts
# - Use Excel Analysis Toolpak
#
# ### AVOID THE HiPPO (highest paid person's opinion)
# - business decision making is a balance between data and our best professional judgment
# - put those things together to make an informed and reasonable decision for any big data question
#
# ### Using BIG DATA in industries:
# - insurance (for pricing risk)
# - manufacturing firms have traditionally been reluctant to use big data
# - banks -- data on people borrowing money
#
# # STEPS in ALL Big Data Projects:
# - Gather/Clean Data
# - Analyze Data
# - Test Choices with Data
# - Make a Decision
#
# ## What is business intelligence?
# - term coined in 1993
# - a group of statistical tools and processes used to help make decisions for a company
# - takes you from raw data to business insight
# - business intelligence combines data and models to help make choices
#
# 
#
#
#
#
# ## The twin paradigms that make things more difficult are:
# - uncertainty: all things that could happen that we hadn't thought about
# - complexity: having too much data at once
#
# ## Advantages of Business Intelligence
# - you avoid guessing - you are making an objective choice based on the best available information that's out there.
# - improves business forecasting
# - helps ensure business continuity (what is the highest pays person or decision maker wanders off to work for another company)
# - reduces subjectivity (gives confidence to our investors and lenders -- if we are able to explain the process)
#
# ## Disadvantages of BI:
# - complexity (don't lose forest for the trees)
# - requires an upfront investment - BI is a process, not a purchase
# - analysis paralysis - paralyzed with fear because we have so many different and competing strings of information coming at us.
# - risk of black boxes (relying too much on software to make choices -- not really understanding what's going on and how the decisions are being made)
#
# # REGRESSION ANALYSIS
#
# - model/tool for predictions and forecasts
# - y=ax+b
# - **simple regression** fits a straight line to the data.
# - in the real world, there are many factors that could impact our decision
# - you can incorporate all of the many factors into a single model called a **multiple regression**
#
#
# # STEPS TO PREDICTION:
# - run regression
# - save coefficients from output (for each of the different factors) for example, this will help us to measure the impact of each additional dollar spent on training.
# - use the coefficients and expected values for the future to get a prediction
#
# # Regressions help us to minimize prediction errors
#
#
#
| BI-Specialist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9 (tensorflow)
# language: python
# name: tensorflow
# ---
# + [markdown] id="LSIM-PITWYFa"
# <a href="https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_05_tpu.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YDTXd8-Lmp8Q"
# # T81-558: Applications of Deep Neural Networks
# **Module 6: Convolutional Neural Networks (CNN) for Computer Vision**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# + [markdown] id="ncNrAEpzmp8S"
# # Module 13 Video Material
#
# * Part 13.1: Flask and Deep Learning Web Services [[Video]](https://www.youtube.com/watch?v=H73m9XvKHug&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_01_flask.ipynb)
# * Part 13.2: Interrupting and Continuing Training [[Video]](https://www.youtube.com/watch?v=kaQCdv46OBA&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_02_checkpoint.ipynb)
# * Part 13.3: Using a Keras Deep Neural Network with a Web Application [[Video]](https://www.youtube.com/watch?v=OBbw0e-UroI&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_03_web.ipynb)
# * Part 13.4: When to Retrain Your Neural Network [[Video]](https://www.youtube.com/watch?v=K2Tjdx_1v9g&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_04_retrain.ipynb)
# * **Part 13.5: Tensor Processing Units (TPUs)** [[Video]](https://www.youtube.com/watch?v=Ygyf3NUqvSc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_13_05_tpu.ipynb)
#
#
# + [markdown] id="lux_6KOXMU94"
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow.
# + colab={"base_uri": "https://localhost:8080/"} id="fU9UhAxTmp8S" outputId="8a0287ac-a84c-41c0-c9c1-a8a51600370d"
# Detect Colab if present
try:
from google.colab import drive
COLAB = True
print("Note: using Google CoLab")
# %tensorflow_version 2.x
except:
print("Note: not using Google CoLab")
COLAB = False
# + [markdown] id="TX_m2PyCWn3e"
# To use Tensor Processing Units (TPUs), you must grant access to Google Cloud Platform (GCP) drives. If this access is not successfully, you will likely see this error:
#
# ```
# InvalidArgumentError: Unable to parse tensor proto
# ```
#
# From Google CoLab, issue the following command:
# + id="Xomyq-3zQAaz"
from google.colab import auth
auth.authenticate_user()
# + [markdown] id="Q09yMGGcmp9N"
# # Part 13.5: Tensor Processing Units (TPUs)
#
# This book focuses primarily on NVIDIA Graphics Processing Units (GPUs) for deep learning acceleration. NVIDIA GPUs are not the only option for deep learning acceleration. TensorFlow continues to gain additional support for AMD and Intel GPUs. TPUs are also available from Google cloud platforms to accelerate deep learning. The focus of this book and course is upon NVIDIA GPUs because of their wide availability on both local and cloud systems.
#
# Though this book focuses on NVIDIA GPUs, we will briefly examine Google Tensor Processing Units (TPUs). These devices are an AI accelerator Application-Specific Integrated Circuit (ASIC) developed by Google. They were designed specifically for neural network machine learning, mainly using Google's TensorFlow software. Google began using TPUs internally in 2015, and in 2018 made them available for third-party use, both as part of its cloud infrastructure and by offering a smaller version of the chip for sale.
#
# The full use of a TPU is a complex topic that I only introduced in this part. Supporting TPUs are slightly more complicated than GPUs because specialized coding is needed. Changes are rarely required to adapt CPU code to GPU for most relatively simple mainstream GPU tasks in TensorFlow. I will cover the mild code changes needed to utilize in this part.
#
# We will create a regression neural network to count paper clips in this part. I demonstrated this dataset and task several times previously in this book. The focus of this part is upon the utilization of TPUs and not the creation of neural networks. I covered the design of computer vision previously in this book.
#
# + colab={"base_uri": "https://localhost:8080/"} id="c8ixjIi5p8Uy" outputId="f82eed1a-b2d1-4d84-edf0-13c7b3650e2e"
# HIDE OUTPUT
import os
import pandas as pd
URL = "https://github.com/jeffheaton/data-mirror/"
DOWNLOAD_SOURCE = URL+"releases/download/v1/paperclips.zip"
DOWNLOAD_NAME = DOWNLOAD_SOURCE[DOWNLOAD_SOURCE.rfind('/')+1:]
if COLAB:
PATH = "/content"
else:
# I used this locally on my machine, you may need different
PATH = "/Users/jeff/temp"
EXTRACT_TARGET = os.path.join(PATH,"clips")
SOURCE = os.path.join(EXTRACT_TARGET, "paperclips")
# Download paperclip data
# !wget -O {os.path.join(PATH,DOWNLOAD_NAME)} {DOWNLOAD_SOURCE}
# !mkdir -p {SOURCE}
# !mkdir -p {TARGET}
# !mkdir -p {EXTRACT_TARGET}
# !unzip -o -j -d {SOURCE} {os.path.join(PATH, DOWNLOAD_NAME)} >/dev/null
# Add filenames
df_train = pd.read_csv(os.path.join(SOURCE, "train.csv"))
df_train['filename'] = "clips-" + df_train.id.astype(str) + ".jpg"
# + [markdown] id="pw_ysSVRiZcP"
# ## Preparing Data for TPUs
#
# To present the paperclips dataset to the TPU we will convert the images to a Keras Dataset. Because we will load the entire dataset to RAM we will only utilize the first 1,000 images. We previously loaded the labels from the **train.csv** file. The following code loads these images and converts them to a Keras dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="RY4NU-vz_Eis" outputId="586c3d29-080d-4d38-cbe7-3b25bac78963"
import tensorflow as tf
import keras_preprocessing
import glob, os
import tqdm
import numpy as np
from PIL import Image
IMG_SHAPE = (128,128)
BATCH_SIZE = 32
# Resize each image and convert the 0-255 ranged RGB values to 0-1 range.
def load_images(files, img_shape):
cnt = len(files)
x = np.zeros((cnt,)+img_shape+(3,),dtype=np.float32)
i = 0
for file in tqdm.tqdm(files):
img = Image.open(file)
img = img.resize(img_shape)
img = np.array(img)
img = img/255
x[i,:,:,:] = img
i+=1
return x
# Process training data
df_train = pd.read_csv(os.path.join(SOURCE, "train.csv"))
df_train['filename'] = "clips-" + df_train.id.astype(str) + ".jpg"
# Use only the first 1000 images
df_train = df_train[0:1000]
# Load images
images = [os.path.join(SOURCE,x) for x in df_train.filename]
x = load_images(images, IMG_SHAPE)
y = df_train.clip_count.values
# Convert to dataset
dataset = tf.data.Dataset.from_tensor_slices((x, y))
dataset = dataset.batch(BATCH_SIZE)
# + [markdown] id="ig6mQ4tBlDsp"
# TPUs are typically Cloud TPU workers, different from the local process running the user's Python program. Thus, it would be best to do some initialization work to connect to the remote cluster and initialize the TPUs. The TPU argument to tf.distribute.cluster_resolver.TPUClusterResolver is a unique address just for Colab. If you are running your code on Google Compute Engine (GCE), you should instead pass in the name of your Cloud TPU. The following code performs this initialization.
# + colab={"base_uri": "https://localhost:8080/"} id="7O04z13LV0Dp" outputId="f405a27a-562a-4441-ce9c-6cf752686025"
# HIDE OUTPUT
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver.connect()
print("Device:", tpu.master())
strategy = tf.distribute.TPUStrategy(tpu)
except:
strategy = tf.distribute.get_strategy()
print("Number of replicas:", strategy.num_replicas_in_sync)
# + [markdown] id="8rVZJBMIgqJq"
# We will now use a ResNet neural network as a basis for our neural network. We will redefine both the input shape and output of the ResNet model, so we will not transfer the weights. Since we redefine the input, the weights are of minimal value. We begin by loading, from Keras, the ResNet50 network. We specify **include_top** as False because we will change the input resolution. We also specify **weights** as false because we must retrain the network after changing the top input layers.
# + colab={"base_uri": "https://localhost:8080/"} id="6MJpmLyhtahJ" outputId="f92c5133-c710-498f-df5a-1614e7be7cbe"
from tensorflow.keras.applications.resnet50 import ResNet50
from tensorflow.keras.layers import Input
from tensorflow.keras.layers import Dense, GlobalAveragePooling2D
from tensorflow.keras.models import Model
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.metrics import RootMeanSquaredError
def create_model():
input_tensor = Input(shape=IMG_SHAPE+(3,))
base_model = ResNet50(
include_top=False, weights=None, input_tensor=input_tensor,
input_shape=None)
x=base_model.output
x=GlobalAveragePooling2D()(x)
x=Dense(1024,activation='relu')(x)
x=Dense(1024,activation='relu')(x)
model=Model(inputs=base_model.input,outputs=Dense(1)(x))
return model
with strategy.scope():
model = create_model()
model.compile(loss = 'mean_squared_error', optimizer='adam',
metrics=[RootMeanSquaredError(name="rmse")])
history = model.fit(dataset, epochs=100, steps_per_epoch=32,
verbose = 1)
# + [markdown] id="9v9hQ59MUwI2"
# You might receive the following error while fitting the neural network.
#
# ```
# InvalidArgumentError: Unable to parse tensor proto
# ```
#
# If you do receive this error, it is likely because you are missing proper authentication to access Google Drive to store your datasets.
| t81_558_class_13_05_tpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Neural ODE のデモンストレーション (レーザービーム)
# +
import sys
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import time
from scipy.integrate import solve_ivp
def f(t,z):
"""
definition of rhs of ode using tensorflow
args z: 4 dim tensor
returns: 4dim tensor
"""
g = 9.8
return (tf.concat( [z[2:4],[0,-g]],axis=0))
def jacobian(t, f, x):
""" return jacobian matrix of f at x"""
n = x.shape[-1].value
fx = f(t, x)
if x.shape[-1].value != fx.shape[-1].value:
print('For calculating Jacobian matrix',
'dimensions of f(x) and x must be the same')
return
return tf.concat([tf.gradients(fx[i], x) for i in range(0, n)], 0)
def Df(t,f,z):
return jacobian(t,f,z)
class F_np:
''' calc rhs of ode (numpy)'''
def __init__(self,sess,f,z_ph):
self.f_tf=f(0,z_ph)
self.sess=sess
self.z_ph=z_ph
def __call__(self,t,z):
return( self.sess.run(self.f_tf,
feed_dict={self.z_ph:z}) )
class F_with_adjoint_np:
'''calc ode and adjoint ode (numpy function)'''
def __init__(self,sess,f,z_ph, a_ph):
"""
args:
sess : session
f: main function of ode
z_ph: placeholder of main variable z
a_ph placeholdero f ajoint variable a
"""
self.dim = 4
self.fz=f(0,z_ph)
self.Df=Df(0,f,z_ph)
self.sess=sess
self.z_ph=z_ph
self.a_ph=a_ph
self.Df_a=-tf.linalg.matvec(self.Df, self.a_ph, transpose_a=True)
def __call__(self,t,za):
fzv,dav = self.sess.run([self.fz, self.Df_a],
feed_dict={self.z_ph:za[0:4], self.a_ph:za[4:8]})
return np.concatenate([fzv,dav])
# -
xy_target = (100,0) # target
t_end = 3 # duration of integration
ts=np.arange(0,t_end+0.1,0.1)
dim=4 # dimension of ode
# +
tf.reset_default_graph()
# forward calculation
c = tf.Variable([35,np.pi/4], dtype=tf.float64,
name='c') # set initial speed and angle here (unit km/h to m/s)
z_0 = tf.concat([tf.Variable([0,0,],dtype=tf.float64), [c[0]*tf.cos(c[1])], [c[0]*tf.sin(c[1])]], axis=0)
z_T= tf.placeholder(shape=[dim],dtype=tf.float64)
L = tf.reduce_sum(tf.square(z_T[0:2]-xy_target) )
# backward calculation
dLdz_T = tf.gradients(L, z_T)[0]
dLdz_0 = tf.placeholder(shape=[dim],dtype=tf.float64)
#Ldummy
Ldummy0 = tf.reduce_sum(dLdz_0* z_0)
# optimizer and training operator
opt = tf.train.GradientDescentOptimizer(1e-5)
train_op = opt.minimize(Ldummy0, var_list=[c])
# +
## tensors etc used in ODE solvers
z_ph = tf.placeholder(shape=[dim],dtype=tf.float64)
a_ph = tf.placeholder(shape=[dim],dtype=tf.float64)
sess = tf.Session()
f_np = F_np(sess,f,z_ph)
f_with_adj = F_with_adjoint_np(sess,f,z_ph,a_ph)
# variables of numpy arrays end with _val
za_val =np.zeros(shape=[dim*2],dtype=np.float64)
sess.run(tf.initializers.global_variables())
# df= Df(0,f,z_ph)
# a_ph=a_ph
# Df_a=-tf.linalg.matvec(df,a_ph, transpose_a=True)
# check
# print(sess.run(df, feed_dict={z_ph:[0,0,0,0]}))
# print(sess.run(Df_a, feed_dict={z_ph:[0,0,0,0],a_ph:[1,1,1,1]}))
# -
# # 前向きと後ろ向き計算の定義
# +
def forward(ts=None):
z0_val, c_val = sess.run([z_0,c])
sol = solve_ivp(fun=f_np,
t_span=[0, t_end], y0=z0_val, t_eval=ts) #,
zt_val = sol['y']
z_T_val = sol['y'][:,-1]
L_val = sess.run(L,feed_dict = {z_T:z_T_val})
return L_val, z_T_val, zt_val,
def backward(z_T_val):
[dLdz_T_val] = sess.run([dLdz_T],feed_dict={z_T:z_T_val})
za_val[0:dim] = z_T_val[0:dim]
za_val[dim:2*dim] = dLdz_T_val[:]
# backward integration
sol_back = solve_ivp(fun=f_with_adj,
t_span=[t_end, 0], y0=za_val, t_eval=ts[::-1]) #,
za_0_val = sol_back['y'][:,-1]
dLdz_0_val = za_0_val[dim:2*dim]
#update c
_,c_val = sess.run([train_op,c], feed_dict={dLdz_0:dLdz_0_val})
return c_val, dLdz_T_val, dLdz_0_val, za_0_val
# -
# # 前向き計算の確認
#
#
# +
#forward calculation
L_val, z_T_val, zt_val = forward(ts=ts)
# z_0 and c value
z0_val, c_val = sess.run([z_0,c])
# change to degree
print('(v_0, phi(deg)) = ({:.4}, {:.4}'.format(c_val[0], 360*c_val[1]/(2*np.pi)))
np.set_printoptions(3)
print('z(0)={}'.format(z0_val))
# z(T)
print('z(T)={}'.format(z_T_val))
print('L= {}'.format(L_val))
#sess.close()
# -
# ## 軌道の図示
plt.plot(zt_val[0,:],zt_val[1,:])
plt.xlabel('x')
plt.ylabel('y')
# # 後ろ向き計算の確認
# +
#backward calculation
c_val, dLdz_T_val, dLdz_0_val, za_0_val = backward(z_T_val)
# [dLdz_T_val] = sess.run([dLdz_T],feed_dict={z_T:z_T_val})
print('dL/dz(T) = {}'.format(dLdz_T_val))
print('dL/dz(0) = {}'.format(dLdz_0_val))
print('(v_0, phi(deg)) = ({:.3}, {:.3})'.format(c_val[0], 360*c_val[1]/(2*np.pi)))
# -
# # メインの学習ループ
# 最初の10回は1回ごとに軌道の保存(図を書くため)
# +
orbits = []
n_itr = []
for i in range(11):
L_val,z_T_val, zt_val = forward()
c_val, *other = backward(z_T_val)
L_val,z_T_val, zt_val = forward(ts=ts)
print('iteration:{}'.format(i))
print('(v_0[m/s], phi[deg]) = ({:.10}, {:.10})'.format(c_val[0], 360*c_val[1]/(2*np.pi)))
print('L= {:.4}'.format(L_val))
print('z(T)={}'.format(z_T_val))
orbits.append(zt_val)
n_itr.append(i)
# -
# 軌道が改良されていくことの確認
orbits[0].shape
fig=plt.figure()
ax=fig.add_subplot(1,1,1)
for ( i,orb) in zip(n_itr,orbits):
ax.plot(orb[0],orb[1], label=i)
ax.legend(loc = 'upper left')
ax.set_xlabel('x')
ax.set_ylabel('y')
# ループの続き
# +
for i in range(11, 10001):
L_val,z_T_val, zt_val = forward()
c_val, *other = backward(z_T_val)
if i % 1000 ==0:
L_val,z_T_val, zt_val = forward(ts=ts)
print('iteration:{}'.format(i))
print('(v_0[m/s], phi[deg]) = ({:.10}, {:.10})'.format(c_val[0], 360*c_val[1]/(2*np.pi)))
print('L= {:.4}'.format(L_val))
print('z(T)={}'.format(z_T_val))
# -
# # 結果確認
#
# 最終的に得られた速度,角度など
print('(v_0[m/s], phi[deg]) = ({:.10}, {:.10})'.format(c_val[0], 360*c_val[1]/(2*np.pi)))
print('L= {:.4}'.format(L_val))
print('z(T)={}'.format(z_T_val))
# 解析解との比較
# +
v0_ana = np.sqrt( (100.0/3.0)**2 + (9.8*t_end/2.0)**2 ) #analytical solution of v_0
print('v_0_ana = {}'.format(v0_ana))
phi_ana = np.arctan2(9.8*3/2, 100.0/3)
print('phi_0_ana = {:.10}[rad]={:.10}[deg]'.format(phi_ana, 360*phi_ana/(2*np.pi)))
#error
er_v_0 = c_val[0]-v0_ana
er_phi = c_val[1]-phi_ana
print('error = {:.8}, {:.8}'.format(er_v_0,er_phi))
# +
L_val,z_T_val, zt_val = forward(ts=ts)
plt.plot(zt_val[0,:],zt_val[1,:])
plt.xlabel('x')
plt.ylabel('y')
# -
| laser-beam-neural-ode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introducing tf.estimator.train_and_evaluate()
#
# **Learning Objectives**
# - Introduce new type of input function (`serving_input_reciever_fn()`) which supports remote access to our model via REST API
# - Use the `tf.estimator.train_and_evaluate()` method to periodically evaluate *during* training
# - Practice using TensorBoard to visualize training and evaluation loss curves
# ## Introduction
#
# In this notebook, we'll see how to use the `train_and_evaluate` method within `tf.estimator` to train and evaluate our machin learning model.
#
# Run the following cell and reset the session if needed:
import tensorflow as tf
import shutil
print(tf.__version__)
# ## Train and Evaluate Input Functions
#
# We'll use the same train and evaluation input functions that we created before.
CSV_COLUMN_NAMES = ["fare_amount","dayofweek","hourofday","pickuplon","pickuplat","dropofflon","dropofflat"]
CSV_DEFAULTS = [[0.0],[1],[0],[-74.0], [40.0], [-74.0], [40.7]]
# +
def parse_row(row):
fields = tf.decode_csv(records = row, record_defaults = CSV_DEFAULTS)
features = dict(zip(CSV_COLUMN_NAMES, fields))
label = features.pop("fare_amount")
return features, label
def read_dataset(csv_path):
dataset = tf.data.TextLineDataset(filenames = csv_path).skip(count = 1) # skip header
dataset = dataset.map(map_func = parse_row)
return dataset
def train_input_fn(csv_path, batch_size = 128):
dataset = read_dataset(csv_path)
dataset = dataset.shuffle(buffer_size = 1000).repeat(count = None).batch(batch_size = batch_size)
return dataset
def eval_input_fn(csv_path, batch_size = 128):
dataset = read_dataset(csv_path)
dataset = dataset.batch(batch_size = batch_size)
return dataset
# -
# ## Feature Columns
#
# We also create the feature columns for the model the same as before.
# +
FEATURE_NAMES = CSV_COLUMN_NAMES[1:] # all but first column
feature_cols = [tf.feature_column.numeric_column(key = k) for k in FEATURE_NAMES]
feature_cols
# -
# ## Serving Input Receiver Function
#
# In a prior notebook we used the `estimator.predict()` function to get taxifare predictions. This worked fine because we had done our model training on the same machine.
#
# However in a production setting this won't usually be the case. Our clients may be remote web servers, mobile apps and more. Instead of having to ship our model files to every client, it would be better to host our model in one place, and make it remotely accesible for prediction requests using a REST API.
#
# The TensorFlow solution for this is a project called [TF Serving](https://www.tensorflow.org/serving/), which is part of the larger [Tensorflow Extended (TFX)](https://www.tensorflow.org/tfx/) platform that extends TensorFlow for production environments.
#
# The interface between TensorFlow and TF Serving is a `serving_input_receiver_fn()`. It has two jobs:
# - To add `tf.placeholder`s to the graph to specify what type of tensors TF Serving should recieve during inference requests. The placeholders are specified as a dictionary object
# - To add any additional ops needed to convert data from the client into the tensors expected by the model.
#
# The function must return a `tf.estimator.export.ServingInputReceiver` object, which packages the placeholders and the neccesary transformations together.
# #### **Exercise 1**
#
# In the cell below, implement a `serving_input_receiver_fn` function that returns an instance of
# `tf.estimator.export.ServingInputReceiver(features, receiver_tensors)`. Have a look at [the documentation for Tensorflow's ServingInputReceiver](https://www.tensorflow.org/api_docs/python/tf/estimator/export/ServingInputReceiver). Here `receiver_tensors` is a dictionary describing the JSON object received by the Cloud ML Engine API, and is a dictionary `features` that has the structure as the feature dictionary accepted by our estimator.
#
# Here we keep things simple by assuming that the API receives a JSON object that has already the correct structure
# (i.e. `features = receiver_tensors`):
def serving_input_receiver_fn():
receiver_tensors = # TODO: Your code goes here
features = receiver_tensors
return tf.estimator.export.ServingInputReceiver(features = features, receiver_tensors = receiver_tensors)
# ## Train and Evaluate
#
# One issue with the previous notebooks is we only evaluate on our validation data once training is complete. This means we can't tell at what point overfitting began. What we really want is to evaluate at specified intervals *during* the training phase.
#
# The Estimator API way of doing this is to replace `estimator.train()` and `estimator.evaluate()` with `estimator.train_and_evaluate()`. This causes an evaluation to be done after every training checkpoint. However by default Tensorflow only checkpoints once every 10 minutes. Since this is less than the length of our total training we'd end up with the same behavior as before which is just one evaluation at the end of training.
#
# To remedy this we speciy in the `tf.estimator.RunConfig()` that TensorFlow should checkpoint every 100 steps.
#
# The default evaluation metric `average_loss` is MSE, but we want RMSE. Previously we just took the square root of the final `average_loss`. However it would be better if we could calculate RMSE not just at the end, but for every intermediate checkpoint and plot the change over time in TensorBoard. [`tf.contrib.estimator.add_metrics()`](https://www.tensorflow.org/api_docs/python/tf/contrib/estimator/add_metrics) allows us to do this. We wrap our estimator with it, and provide a custom evaluation function.
#
# `train_and_evaluate()` also allows us to use our `serving_input_receiver_fn()` to export our models in the SavedModel format required by TF Serving.
#
# *Note: Training will be slower than the last notebook because we are now evaluating after every 100 train steps. Previously we didn't evaluate until training finished.*
# #### **Exercise 2**
#
# In the cell below, create a instance of `tf.estimator.RunConfig` named `config` and pass to its
# constructor information concerning:
# - the directory where we want the trained model and its checkpoints to be saved
# - the random seed which we want to be set to 1
# - the cadence at which we want the model to create checkpoints (every 100 steps)
#
# To remind yourself what arguments `tf.estimator.RunConfig` takes have a look at [the documentation](https://www.tensorflow.org/api_docs/python/tf/estimator/RunConfig).
# +
OUTDIR = "taxi_trained"
config = tf.estimator.RunConfig(
# TODO: Your code goes here
)
# -
# #### **Exercise 3**
#
# In the cell below, create a `DNNRegressor` model with two layers of 10 neurons each using
# the `RunConfig` instance and the `feature_cols` list you just created.
#
# Note that we do not need to pass the model directory directly to the estimator constructor, since that info should
# already be wrapped into the `RunConfig` instance.
model = tf.estimator.DNNRegressor(
# TODO: Your code goes here
)
# ### Adding custom evaluation metrics
#
# If we want to add a custom evaluation metric (one not included automatically with the canned `DNNRegressor` estimator) we will can do that by wrapping our model with our custom metric function using the `contrib` function `.add_metrics`. We will implement a `my_rmse` function that
# - takes as input a tensor of `labels` and a tensor of `predictions`
# - returns a dictionary with the single key `rmse` and with value the root mean square error between the labels and the predictions
#
# You can have a look at this blog post by <NAME> on ["How to extend a canned TensorFlow Estimator"](https://towardsdatascience.com/how-to-extend-a-canned-tensorflow-estimator-to-add-more-evaluation-metrics-and-to-pass-through-ddf66cd3047d) for more information.
# #### **Exercise 4**
#
# Implement a `my_rmse` function that
# - takes as input a tensor of `labels` and a tensor of `predictions`
# - returns a dictionary with the single key `rmse` and with value the root mean square error between the labels and the predictions
#
# **Hint:** Have a look at [the Tensorflow documentation](https://www.tensorflow.org/api_docs/python/tf/metrics/root_mean_squared_error) for `tf.metrics.root_mean_squared_error`. You will have to do some preliminary step to `predictions` before you can compute the RMSE. In fact, you may notice that you get a shape error if you try to use the prediction values as is. It may help to use `tf.squeeze`. Have a closer look at what `tf.sqeeze` does in [the docs here](https://www.tensorflow.org/api_docs/python/tf/squeeze).
def my_rmse(labels, predictions):
pred_values = # TODO: Your code goes here
return {
"rmse": # TODO: Your code goes here
}
# Run the following cell to add the custom metric you defined above to the `model`:
model = tf.contrib.estimator.add_metrics(estimator = model, metric_fn = my_rmse)
# #### **Exercise 5**
#
# In the cell block below, create an instance of a `tf.estimator.TrainSpec` using the `train_input_fn` defined at the top of this file and
# with a `max_steps` of 500. Note, the training data should be loaded from `./taxi-train.csv`. See the details of how to implement a Tensorflow `TrainSpec` in [the documentation](https://www.tensorflow.org/api_docs/python/tf/estimator/TrainSpec).
#
# **Hint:** You may need to use a `lambda` function to pass the
# training input function correctly.
train_spec = tf.estimator.TrainSpec(
input_fn = # TODO: Your code goes here
max_steps = # TODO: Your code goes here
)
# #### **Exercise 6**
#
# Next, create an exporter using the `serving_input_receiver_fn` defined at the beginning of this notebook. You want to export the trained model and its checkpoints in the './exporter' subdirectory. Use `tf.estimator.FinalExporter` to create the exporter intance. Have a look at [the documentation for FinalExporter](https://www.tensorflow.org/api_docs/python/tf/estimator/FinalExporter) to ensure proper usage.
#
# **Note:** You may alternatively use `tf.estimator.BestExporter`
# to export at every checkpoint that has lower loss than the previous checkpoint, instead
# of exporting only the last checkpoint.
exporter = # TODO: Your code goes here
# #### **Exercise 7**
#
# In the cell below, create an instance of an `EvalSpec` to which you specify that
# - the data should be loaded from `/.taxi-valid.csv` during evaluation (use the correct input function!)
# - the exporter you defined above should be used
# - the first evaluation should start after 1 second of training
# - and then be repeated every 1 second
#
# Look at [the documentaiton for tf.estimator.EvalSpec](https://www.tensorflow.org/api_docs/python/tf/estimator/EvalSpec) to help.
#
# **Note:** We use the checkpoint setting above because we want to evaluate after every checkpoint.
# As long as checkpoints are > 1 sec apart this ensures the throttling never kicks in.
eval_spec = # TODO: Your code goes here
# #### **Exercise 8**
#
# Finally we use `tf.estimator.train_and_evaluate` to start the training and evaluation as you specified them above. Complete the code in the cell below, providing the necessary arguments. Have a look at [the documentation for the train_and_evaluate method](https://www.tensorflow.org/api_docs/python/tf/estimator/train_and_evaluate) to make sure you pass everything it needs.
# +
tf.logging.set_verbosity(tf.logging.INFO)
shutil.rmtree(path = OUTDIR, ignore_errors = True)
tf.summary.FileWriterCache.clear() # ensure filewriter cache is clear for TensorBoard events file
tf.estimator.train_and_evaluate(# TODO: Your code goes here
)
# -
# ## Inspect Export Folder
#
# Now in the output directory, in addition to the checkpoint files, you'll see a subfolder called 'export'. This contains one or models in the SavedModel format which is compatible with TF Serving. In the next notebook we will deploy the SavedModel behind a production grade REST API.
# !ls -R taxi_trained/export
# ## Monitoring with TensorBoard
#
# [TensorBoard](https://www.tensorflow.org/guide/summaries_and_tensorboard) is a web UI that allows us to visualize various aspects of our model, including the training and evaluation loss curves. Although you won't see the loss curves yet, it is best to launch TensorBoard *before* you start training so that you may see them update during a long running training process.
#
# To get Tensorboard to work within a Deep Learning VM or Colab, we need to create a tunnel connection to your local machine. To do this we'll set up a tunnel connection with `ngrok`. Using ngrok we'll then create a tunnel connection to our virtual machine's port 6006. We can view the Tensorboard results by following the link provided by ngrok after executing the following cell.
# +
get_ipython().system_raw(
"tensorboard --logdir {} --host 0.0.0.0 --port 6006 &"
.format(OUTDIR)
)
get_ipython().system_raw("../assets/ngrok http 6006 &")
# -
# !curl -s http://localhost:4040/api/tunnels | python3 -c \
# "import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
# ### Tensorboard cleanup
#
# To close the tunnel connection to Tensorboard, we can find the PIDs for ngrok and Tensorboard and stop them.
# this will kill the processes for Tensorboard
# !ps aux | grep tensorboard | awk '{print $2}' | xargs kill
# this will kill the processes for ngrok
# !ps aux | grep ngrok | awk '{print $2}' | xargs kill
# ## Challenge exercise
#
# Modify your solution to the challenge exercise in d_csv_input.ipynb appropriately.
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/02_tensorflow/labs/e_traineval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # マップ圧縮コード
# ## マップを配列に格納
#
# https://labo-code.com/python/image-analysis/image-to-list/
# +
from PIL import Image
import numpy as np
import cv2
img = Image.open('tsudanuma.pgm')
width, height = img.size
image_array = np.empty((height, width), dtype = int)
for y in range(height):
for x in range(width):
image_array[y][x] = img.getpixel((x, y))
# -
# ## マップのサイズ入力
m = 6240
n = 4000
# ## マップのサイズの変更する倍率を設定
z = 10
# ## マップの行列が格納できているのかを確認
#
# https://www.sejuku.net/blog/75248
a = image_array
print(a)
# ## z列の行列に変換
#
# https://note.nkmk.me/python-list-ndarray-1d-to-2d/
b = a.reshape(-1, z)
print(b)
# ## 行ごとの最小値を求める
#
# https://deepage.net/features/numpy-min.html
c = np.amin(b, axis=1)
print(c)
# ## 列が1/zの形にする
x1 = int(m/z)
d = c.reshape(-1, x1)
print(d)
# ## 転置する
#
# https://note.nkmk.me/python-numpy-transpose/
e = d.T
print(e)
# ## 繰り返し
f = e.reshape(-1, z)
g = np.amin(f, axis=1)
y1 = int(n/z)
h = g.reshape(-1, y1)
i = h.T
print(i)
# ## マップを保存する
#
# https://tomokichi.blog/%E3%80%90python%E3%80%91%E5%8A%A0%E5%B7%A5%E3%81%97%E3%81%9F%E7%94%BB%E5%83%8F%E3%82%92%E4%BF%9D%E5%AD%98%E3%81%97%E3%81%A6%E3%81%BF%E3%82%88%E3%81%86/
cv2.imwrite('new.pgm',i)
| map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This notebook is an exercise in the [Geospatial Analysis](https://www.kaggle.com/learn/geospatial-analysis) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/coordinate-reference-systems).**
#
# ---
#
# # Introduction
#
# You are a bird conservation expert and want to understand migration patterns of purple martins. In your research, you discover that these birds typically spend the summer breeding season in the eastern United States, and then migrate to South America for the winter. But since this bird is under threat of endangerment, you'd like to take a closer look at the locations that these birds are more likely to visit.
#
# <center>
# <img src="https://i.imgur.com/qQcS0KM.png" width="1000"><br/>
# </center>
#
# There are several [protected areas](https://www.iucn.org/theme/protected-areas/about) in South America, which operate under special regulations to ensure that species that migrate (or live) there have the best opportunity to thrive. You'd like to know if purple martins tend to visit these areas. To answer this question, you'll use some recently collected data that tracks the year-round location of eleven different birds.
#
# Before you get started, run the code cell below to set everything up.
# +
import pandas as pd
import geopandas as gpd
from shapely.geometry import LineString
from learntools.core import binder
binder.bind(globals())
from learntools.geospatial.ex2 import *
# -
# # Exercises
#
# ### 1) Load the data.
#
# Run the next code cell (without changes) to load the GPS data into a pandas DataFrame `birds_df`.
# Load the data and print the first 5 rows
birds_df = pd.read_csv("../input/geospatial-learn-course-data/purple_martin.csv", parse_dates=['timestamp'])
print("There are {} different birds in the dataset.".format(birds_df["tag-local-identifier"].nunique()))
birds_df.head()
# There are 11 birds in the dataset, where each bird is identified by a unique value in the "tag-local-identifier" column. Each bird has several measurements, collected at different times of the year.
#
# Use the next code cell to create a GeoDataFrame `birds`.
# - `birds` should have all of the columns from `birds_df`, along with a "geometry" column that contains Point objects with (longitude, latitude) locations.
# - Set the CRS of `birds` to `{'init': 'epsg:4326'}`.
# +
# Create the GeoDataFrame
birds = gpd.GeoDataFrame(birds_df, geometry=gpd.points_from_xy(birds_df["location-long"], birds_df["location-lat"]))
# Set the CRS to {'init': 'epsg:4326'}
birds.crs = {'init' :'epsg:4326'}
# Check your answer
q_1.check()
# +
# Lines below will give you a hint or solution code
#q_1.hint()
#q_1.solution()
# -
# ### 2) Plot the data.
#
# Next, we load in the `'naturalearth_lowres'` dataset from GeoPandas, and set `americas` to a GeoDataFrame containing the boundaries of all countries in the Americas (both North and South America). Run the next code cell without changes.
# Load a GeoDataFrame with country boundaries in North/South America, print the first 5 rows
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
americas = world.loc[world['continent'].isin(['North America', 'South America'])]
americas.head()
# Use the next code cell to create a single plot that shows both: (1) the country boundaries in the `americas` GeoDataFrame, and (2) all of the points in the `birds_gdf` GeoDataFrame.
#
# Don't worry about any special styling here; just create a preliminary plot, as a quick sanity check that all of the data was loaded properly. In particular, you don't have to worry about color-coding the points to differentiate between birds, and you don't have to differentiate starting points from ending points. We'll do that in the next part of the exercise.
# Your code here
ax = americas.plot(figsize=(10,10), color="w", linestyle=':', edgecolor = 'gray')
birds.plot(ax=ax, markersize = 15, color="black", marker = 'o')
# Uncomment to see a hint
#q_2.hint()
# +
# Get credit for your work after you have created a map
q_2.check()
# Uncomment to see our solution (your code may look different!)
##q_2.solution()
# -
# ### 3) Where does each bird start and end its journey? (Part 1)
#
# Now, we're ready to look more closely at each bird's path. Run the next code cell to create two GeoDataFrames:
# - `path_gdf` contains LineString objects that show the path of each bird. It uses the `LineString()` method to create a LineString object from a list of Point objects.
# - `start_gdf` contains the starting points for each bird.
# +
# GeoDataFrame showing path for each bird
path_df = birds.groupby("tag-local-identifier")['geometry'].apply(list).apply(lambda x: LineString(x)).reset_index()
path_gdf = gpd.GeoDataFrame(path_df, geometry=path_df.geometry)
path_gdf.crs = {'init' :'epsg:4326'}
# GeoDataFrame showing starting point for each bird
start_df = birds.groupby("tag-local-identifier")['geometry'].apply(list).apply(lambda x: x[0]).reset_index()
start_gdf = gpd.GeoDataFrame(start_df, geometry=start_df.geometry)
start_gdf.crs = {'init' :'epsg:4326'}
# Show first five rows of GeoDataFrame
start_gdf.head()
# -
# Use the next code cell to create a GeoDataFrame `end_gdf` containing the final location of each bird.
# - The format should be identical to that of `start_gdf`, with two columns ("tag-local-identifier" and "geometry"), where the "geometry" column contains Point objects.
# - Set the CRS of `end_gdf` to `{'init': 'epsg:4326'}`.
# +
# Your code here
end_df = birds.groupby("tag-local-identifier")['geometry'].apply(list).apply(lambda x: x[-1]).reset_index()
end_gdf = gpd.GeoDataFrame(end_df, geometry=end_df.geometry)
end_gdf.crs = {'init': 'epsg:4326'}
# Check your answer
q_3.check()
# +
# Lines below will give you a hint or solution code
#q_3.hint()
#q_3.solution()
# -
# ### 4) Where does each bird start and end its journey? (Part 2)
#
# Use the GeoDataFrames from the question above (`path_gdf`, `start_gdf`, and `end_gdf`) to visualize the paths of all birds on a single map. You may also want to use the `americas` GeoDataFrame.
# +
# Your code here
ax = americas.plot(figsize=(10, 10), color='white', linestyle=':', edgecolor='gray')
start_gdf.plot(ax=ax, color='red', markersize=30)
path_gdf.plot(ax=ax, cmap='tab20b', linestyle='-', linewidth=1, zorder=1)
end_gdf.plot(ax=ax, color='black', markersize=30)
# Uncomment to see a hint
#q_4.hint()
# +
# Get credit for your work after you have created a map
q_4.check()
# Uncomment to see our solution (your code may look different!)
#q_4.solution()
# -
# ### 5) Where are the protected areas in South America? (Part 1)
#
# It looks like all of the birds end up somewhere in South America. But are they going to protected areas?
#
# In the next code cell, you'll create a GeoDataFrame `protected_areas` containing the locations of all of the protected areas in South America. The corresponding shapefile is located at filepath `protected_filepath`.
# +
# Path of the shapefile to load
protected_filepath = "../input/geospatial-learn-course-data/SAPA_Aug2019-shapefile/SAPA_Aug2019-shapefile/SAPA_Aug2019-shapefile-polygons.shp"
# Your code here
protected_areas = gpd.read_file(protected_filepath)
# Check your answer
q_5.check()
# +
# Lines below will give you a hint or solution code
#q_5.hint()
#q_5.solution()
# -
# ### 6) Where are the protected areas in South America? (Part 2)
#
# Create a plot that uses the `protected_areas` GeoDataFrame to show the locations of the protected areas in South America. (_You'll notice that some protected areas are on land, while others are in marine waters._)
# +
# Country boundaries in South America
south_america = americas.loc[americas['continent']=='South America']
# Your code here: plot protected areas in South America
ax = south_america.plot(figsize=(10,10), color = "white", edgecolor = 'grey')
protected_areas.plot(ax=ax, alpha=0.4)
# Uncomment to see a hint
#q_6.hint()
# +
# Get credit for your work after you have created a map
q_6.check()
# Uncomment to see our solution (your code may look different!)
#q_6.solution()
# -
# ### 7) What percentage of South America is protected?
#
# You're interested in determining what percentage of South America is protected, so that you know how much of South America is suitable for the birds.
#
# As a first step, you calculate the total area of all protected lands in South America (not including marine area). To do this, you use the "REP_AREA" and "REP_M_AREA" columns, which contain the total area and total marine area, respectively, in square kilometers.
#
# Run the code cell below without changes.
P_Area = sum(protected_areas['REP_AREA']-protected_areas['REP_M_AREA'])
print("South America has {} square kilometers of protected areas.".format(P_Area))
# Then, to finish the calculation, you'll use the `south_america` GeoDataFrame.
south_america.head()
# Calculate the total area of South America by following these steps:
# - Calculate the area of each country using the `area` attribute of each polygon (with EPSG 3035 as the CRS), and add up the results. The calculated area will be in units of square meters.
# - Convert your answer to have units of square kilometeters.
# Your code here: Calculate the total area of South America (in square kilometers)
totalArea = sum(south_america.geometry.to_crs(epsg=3035).area) / 10**6
# Check your answer
display(totalArea)
# Check your answer
q_7.check()
# +
# Lines below will give you a hint or solution code
#q_7.hint()
#q_7.solution()
# -
# Run the code cell below to calculate the percentage of South America that is protected.
# What percentage of South America is protected?
percentage_protected = P_Area/totalArea
print('Approximately {}% of South America is protected.'.format(round(percentage_protected*100, 2)))
# ### 8) Where are the birds in South America?
#
# So, are the birds in protected areas?
#
# Create a plot that shows for all birds, all of the locations where they were discovered in South America. Also plot the locations of all protected areas in South America.
#
# To exclude protected areas that are purely marine areas (with no land component), you can use the "MARINE" column (and plot only the rows in `protected_areas[protected_areas['MARINE']!='2']`, instead of every row in the `protected_areas` GeoDataFrame).
# Your code here
ax = south_america.plot(figsize=(10,10), color='white', edgecolor='gray')
protected_areas[protected_areas['MARINE']!='2'].plot(ax=ax, alpha=0.4, zorder=1)
birds[birds.geometry.y < 0].plot(ax=ax, color='red', alpha=0.6, markersize=10, zorder=2)
# Uncomment to see a hint
#q_8.hint()
# +
# Get credit for your work after you have created a map
q_8.check()
# Uncomment to see our solution (your code may look different!)
#q_8.solution()
# -
# # Keep going
#
# Create stunning **[interactive maps](https://www.kaggle.com/alexisbcook/interactive-maps)** with your geospatial data.
# ---
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/161464) to chat with other Learners.*
| exercise2-coordinate-reference-systems.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sb
import matplotlib.pyplot as plt
# +
split = 'chemical_may_cv3'
train_assay_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/assay_matrix_discrete_train_scaff.csv'
test_assay_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/assay_matrix_discrete_test_scaff.csv'
aucs_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/2021_03_evaluation_all_data_python.csv'
train_assay_df = pd.read_csv(train_assay_file)
test_assay_df = pd.read_csv(test_assay_file)
aucs_df = pd.read_csv(aucs_file).set_index('assay_id')
modalities = ['mowh_es_op', 'cp_es_op', 'ge_es_op', 'late_fusion_cs_ge', 'late_fusion_cs_ge_mowh', 'late_fusion_cs_mowh', 'late_fusion_ge_mowh']
#modalities = ['late_fusion_cs_ge', 'late_fusion_cs_ge_mowh', 'late_fusion_cs_mowh', 'late_fusion_ge_mowh']
aucs_df = aucs_df[aucs_df['descriptor'].isin(modalities)]
modalities_dict = {'mowh_es_op':'MO', 'cp_es_op':'CS', 'ge_es_op':'GE', 'late_fusion_cs_ge':'CS+GE', 'late_fusion_cs_ge_mowh':'CS+GE+MO', 'late_fusion_cs_mowh':'CS+MO', 'late_fusion_ge_mowh':'GE+MO'}
#modalities_dict = {'late_fusion_cs_ge':'CS+GE', 'late_fusion_cs_ge_mowh':'CS+GE+MO', 'late_fusion_cs_mowh':'CS+MO', 'late_fusion_ge_mowh':'GE+MO'}
aucs_df['descriptor'].replace(modalities_dict, inplace=True)
assays = train_assay_df.columns[1:]
# +
readouts = {}
readouts_compounds = {}
hits = {}
hit_rate = {}
hits_compounds = {}
for a in assays:
readouts[a] = np.count_nonzero(~np.isnan(test_assay_df[a].to_numpy()))
readouts_compounds[a] = set(test_assay_df[test_assay_df[a].notna()]['smiles'])
hits[a] = int(np.nansum(test_assay_df[a].to_numpy()))
hits_compounds[a] = set(test_assay_df[test_assay_df[a] > 0]['smiles'])
if readouts[a] != 0:
hit_rate[a] = hits[a] / readouts[a]
else:
hit_rate[a] = 'NA'
readouts_df = pd.Series(readouts, name="readouts").to_frame()
hits_df = pd.Series(hits, name="hits").to_frame()
hit_rate_df = pd.Series(hit_rate, name="hit_rate").to_frame()
# -
aucs_df_readouts_hits_df = pd.merge(aucs_df, readouts_df, left_index=True, right_index=True)
aucs_df_readouts_hits_df = pd.merge(aucs_df_readouts_hits_df, hits_df, left_index=True, right_index=True)
aucs_df_readouts_hits_df = pd.merge(aucs_df_readouts_hits_df, hit_rate_df, left_index=True, right_index=True)
aucs_df_readouts_hits_df.index.name = 'assay_id'
aucs_df_readouts_hits_df
hits_compounds['100_277']
# +
top_hit_rate_dict = {'CS':{}, 'GE': {}, 'MO':{}, 'CS+GE':{}, 'GE+MO':{}, 'CS+MO':{}, 'CS+GE+MO': {} }
list_dataframe = []
# 3200 compounds / rank / rank only readouts / remove others
# hit rate normalized / total number of hits (top number of hits), basically it will top (number of hits) instead of top 1%
predictions_cs_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/predictions/predictions_cp_es_op.csv'
predictions_cs_df = pd.read_csv(predictions_cs_file)
for a in assays:
if hits[a] != 0:
cut = predictions_cs_df[predictions_cs_df['smiles'].isin(readouts_compounds[a])]
cut = cut.sort_values(by=a, ascending=False, na_position='last').head(hits[a])[['smiles',a]]
hits_top1 = set(cut['smiles'])
#top_hit_rate_dict['CS'][a] = len(hits_compounds[a].intersection(hits_top1)) / int(len(predictions_cs_df) / 100)
top_hit_rate_dict['CS'][a] = len(hits_compounds[a].intersection(hits_top1)) / hits[a]
list_dataframe.append([a,'CS', top_hit_rate_dict['CS'][a]])
else:
top_hit_rate_dict['CS'][a] = 'NA'
list_dataframe.append([a,'CS','NA'])
predictions_mo_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/predictions/predictions_mowh_es_op.csv'
predictions_mo_df = pd.read_csv(predictions_cs_file)
for a in assays:
if hits[a] != 0:
cut = predictions_mo_df[predictions_mo_df['smiles'].isin(readouts_compounds[a])]
cut = cut.sort_values(by=a, ascending=False, na_position='last').head(hits[a])[['smiles',a]]
# cut = predictions_mo_df.sort_values(by=a, ascending=False, na_position='last').head(int(len(predictions_mo_df) / 100))[['smiles',a]]
hits_top1 = set(cut['smiles'])
#top_hit_rate_dict['MO'][a] = len(hits_compounds[a].intersection(hits_top1)) / int(len(predictions_mo_df) / 100)
top_hit_rate_dict['MO'][a] = len(hits_compounds[a].intersection(hits_top1)) / hits[a]
list_dataframe.append([a,'MO', top_hit_rate_dict['MO'][a]])
else:
top_hit_rate_dict['MO'][a] = 'NA'
list_dataframe.append([a,'MO','NA'])
predictions_ge_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/predictions/predictions_ge_es_op.csv'
predictions_ge_df = pd.read_csv(predictions_cs_file)
for a in assays:
if hits[a] != 0:
#cut = predictions_ge_df.sort_values(by=a, ascending=False, na_position='last').head(int(len(predictions_ge_df) / 100))[['smiles',a]]
cut = predictions_ge_df[predictions_ge_df['smiles'].isin(readouts_compounds[a])]
cut = cut.sort_values(by=a, ascending=False, na_position='last').head(hits[a])[['smiles',a]]
hits_top1 = set(cut['smiles'])
#top_hit_rate_dict['GE'][a] = len(hits_compounds[a].intersection(hits_top1)) / int(len(predictions_ge_df) / 100)
top_hit_rate_dict['GE'][a] = len(hits_compounds[a].intersection(hits_top1)) / hits[a]
list_dataframe.append([a,'GE', top_hit_rate_dict['GE'][a]])
else:
top_hit_rate_dict['GE'][a] = 'NA'
list_dataframe.append([a,'GE','NA'])
predictions_gemo_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/predictions/late_fusion_ge_mowh.csv'
predictions_gemo_df = pd.read_csv(predictions_cs_file)
for a in assays:
if hits[a] != 0:
#cut = predictions_gemo_df.sort_values(by=a, ascending=False, na_position='last').head(int(len(predictions_gemo_df) / 100))[['smiles',a]]
cut = predictions_gemo_df[predictions_gemo_df['smiles'].isin(readouts_compounds[a])]
cut = cut.sort_values(by=a, ascending=False, na_position='last').head(hits[a])[['smiles',a]]
hits_top1 = set(cut['smiles'])
#top_hit_rate_dict['GE+MO'][a] = len(hits_compounds[a].intersection(hits_top1)) / int(len(predictions_gemo_df) / 100)
top_hit_rate_dict['GE+MO'][a] = len(hits_compounds[a].intersection(hits_top1)) / hits[a]
list_dataframe.append([a,'GE+MO', top_hit_rate_dict['GE+MO'][a]])
else:
top_hit_rate_dict['GE+MO'][a] = 'NA'
list_dataframe.append([a,'GE+MO','NA'])
predictions_csmo_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/predictions/late_fusion_cs_mowh.csv'
predictions_csmo_df = pd.read_csv(predictions_cs_file)
for a in assays:
if hits[a] != 0:
#cut = predictions_csmo_df.sort_values(by=a, ascending=False, na_position='last').head(int(len(predictions_csmo_df) / 100))[['smiles',a]]
cut = predictions_csmo_df[predictions_csmo_df['smiles'].isin(readouts_compounds[a])]
cut = cut.sort_values(by=a, ascending=False, na_position='last').head(hits[a])[['smiles',a]]
hits_top1 = set(cut['smiles'])
#top_hit_rate_dict['CS+MO'][a] = len(hits_compounds[a].intersection(hits_top1)) / int(len(predictions_csmo_df) / 100)
top_hit_rate_dict['CS+MO'][a] = len(hits_compounds[a].intersection(hits_top1)) / hits[a]
list_dataframe.append([a,'CS+MO', top_hit_rate_dict['CS+MO'][a]])
else:
top_hit_rate_dict['CS+MO'][a] = 'NA'
list_dataframe.append([a,'CS+MO','NA'])
predictions_csge_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/predictions/late_fusion_cs_ge.csv'
predictions_csge_df = pd.read_csv(predictions_cs_file)
for a in assays:
if hits[a] != 0:
#cut = predictions_csge_df.sort_values(by=a, ascending=False, na_position='last').head(int(len(predictions_csge_df) / 100))[['smiles',a]]
cut = predictions_csge_df[predictions_csge_df['smiles'].isin(readouts_compounds[a])]
cut = cut.sort_values(by=a, ascending=False, na_position='last').head(hits[a])[['smiles',a]]
hits_top1 = set(cut['smiles'])
#top_hit_rate_dict['CS+GE'][a] = len(hits_compounds[a].intersection(hits_top1)) / int(len(predictions_csge_df) / 100)
top_hit_rate_dict['CS+GE'][a] = len(hits_compounds[a].intersection(hits_top1)) / hits[a]
list_dataframe.append([a,'CS+GE', top_hit_rate_dict['CS+GE'][a]])
else:
top_hit_rate_dict['CS+GE'][a] = 'NA'
list_dataframe.append([a,'CS+GE','NA'])
predictions_csgemo_file = '/home/jupyter-nmoshkov@broadinst-04e9f/PUMA/' + split + '/predictions/late_fusion_cs_ge_mowh.csv'
predictions_csgemo_df = pd.read_csv(predictions_cs_file)
for a in assays:
if hits[a] != 0:
#cut = predictions_csgemo_df.sort_values(by=a, ascending=False, na_position='last').head(int(len(predictions_csgemo_df) / 100))[['smiles',a]]
cut = predictions_csgemo_df[predictions_csgemo_df['smiles'].isin(readouts_compounds[a])]
cut = cut.sort_values(by=a, ascending=False, na_position='last').head(hits[a])[['smiles',a]]
hits_top1 = set(cut['smiles'])
#top_hit_rate_dict['CS+GE+MO'][a] = len(hits_compounds[a].intersection(hits_top1)) / int(len(predictions_csgemo_df) / 100)
top_hit_rate_dict['CS+GE+MO'][a] = len(hits_compounds[a].intersection(hits_top1)) / hits[a]
list_dataframe.append([a,'CS+GE+MO', top_hit_rate_dict['CS+GE+MO'][a]])
else:
top_hit_rate_dict['CS+GE+MO'][a] = 'NA'
list_dataframe.append([a,'CS+GE+MO','NA'])
# +
#aucs_df_readouts_hits_df.to_csv('consolidated_data_latefusion_' + split + '.csv')
# -
top1_hit_rate_df = pd.DataFrame(list_dataframe, columns=['assay_id', 'descriptor','top_rank_hit_rate'])
top1_hit_rate_df
final_df = pd.merge(aucs_df_readouts_hits_df.reset_index(level=0), top1_hit_rate_df)
final_df
final_df.to_csv('toprank_' + split + '_hitsnorm.csv')
| analysis/tables_for_HitRates_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 09: Deep Learning Part I: Fully Connected Neural Networks
#
# In class, we have developed the mathematics and programming techniques for binary classification using fully connected neural networks
# having one or more hidden layers.
#
# Today, we'll expand on that to consider (small) image classification using again fully connected neural networks with a multinomial
# (softmax) output layer.
#
# ## MNIST Data
#
# An image is a 2D array of pixels. Pixels can be scalar intensities (for a grayscale / black and white image) or a vector indicating a
# point in a color space such as RGB or HSV.
#
# Today we'll consider 8x8 grayscale images of digits from the famous "MNIST" dataset, which was considered a benchmark for machine learning algorithms
# up to the early 2000s, before the advent of large-scale image classification datasets.
#
# This dataset in SciKit-Learn has 10 classes, with 180 samples per class in most cases, for a total of 1797 samples.
#
# Let's load the dataset and plot an example.
# +
import numpy as np
from sklearn.datasets import load_digits
import matplotlib.pyplot as plt
# Load data
data = load_digits()
def convert_to_one_hot(y):
y_vect = np.zeros((len(y), 10))
for i in range(len(y)):
y_vect[i, int(y[i])] = 1
return y_vect
# Convert target indices to one-hot representation
y_indices = data.target
y = convert_to_one_hot(y_indices)
X = np.matrix(data.data)
M = X.shape[0]
N = X.shape[1]
# Plot an example
plt.imshow(np.reshape(X[0,:],(8,8)), 'gray')
plt.title('Example MNIST sample (category %d)' % y_indices[0])
# -
# ## Hand-Coded Fully Connected Neural Network
#
# OK, now let's modify our code from class to work with this dataset and run 100 epochs of training.
# The main change is to use a one-hot encoding of the 10 classes at the output layer and to use
# the softmax activation function at the output. Some minor changes are required to calculate multinomial
# cross entropy loss rather than binary cross entropy loss.
# +
import random
import warnings
warnings.filterwarnings("ignore")
# Normalize each input feature
def normalize(X):
M = X.shape[0]
XX = X - np.tile(np.mean(X,0),[M,1])
XX = np.divide(XX, np.tile(np.std(XX,0),[M,1]))
return np.nan_to_num(XX, copy=True,nan=0.0)
XX = normalize(X)
idx = np.arange(0,M)
# Partion data into training and testing dataset
random.shuffle(idx)
percent_train = .6
m_train = int(M * percent_train)
train_idx = idx[0:m_train]
test_idx = idx[m_train:M+1]
X_train = XX[train_idx,:];
X_test = XX[test_idx,:];
y_train = y[train_idx];
y_test = y[test_idx];
y_test_indices = y_indices[test_idx]
# Let's start with a 3-layer network with sigmoid activation functions,
# 6 units in layer 1, and 5 units in layer 2.
h2 = 5
h1 = 6
W = [[], np.random.normal(0,0.1,[N,h1]),
np.random.normal(0,0.1,[h1,h2]),
np.random.normal(0,0.1,[h2,10])]
b = [[], np.random.normal(0,0.1,[h1,1]),
np.random.normal(0,0.1,[h2,1]),
np.random.normal(0,0.1,[10,1])]
L = len(W)-1
def sigmoid_act(z):
return 1/(1+np.exp(-z))
def softmax_act(z):
exps = np.exp(z)
return exps / np.sum(exps)
def sigmoid_actder(z):
az = sigmoid_act(z)
prod = np.multiply(az,1-az)
return prod
def ff(x,W,b):
L = len(W)-1
a = x
for l in range(1,L+1):
z = W[l].T*a+b[l]
if (l == L):
a = softmax_act(z)
else:
a = sigmoid_act(z)
return a
def loss(y, yhat):
return - np.dot(y, np.log(yhat))
# Train for 100 epochs with mini-batch size 1
cost_arr = []
alpha = 0.01
max_iter = 100
for iter in range(0, max_iter):
loss_this_iter = 0
order = np.random.permutation(m_train)
for i in range(0, m_train):
# Grab the pattern order[i]
x_this = X_train[order[i],:].T
y_this = y_train[order[i],:]
# Feed forward step
a = [x_this]
z = [[]]
delta = [[]]
dW = [[]]
db = [[]]
for l in range(1,L+1):
z.append(W[l].T*a[l-1]+b[l])
if (l == L):
a.append(softmax_act(z[l]))
else:
a.append(sigmoid_act(z[l]))
# Just to give arrays the right shape for the backprop step
delta.append([]); dW.append([]); db.append([])
loss_this_pattern = loss(y_this, a[L])
loss_this_iter = loss_this_iter + loss_this_pattern
# Backprop step. Note that derivative of multinomial cross entropy
# loss is the same as that of binary cross entropy loss. See
# https://levelup.gitconnected.com/killer-combo-softmax-and-cross-entropy-5907442f60ba
# for a nice derivation.
delta[L] = a[L] - np.matrix(y_this).T
for l in range(L,0,-1):
db[l] = delta[l].copy()
dW[l] = a[l-1] * delta[l].T
if l > 1:
delta[l-1] = np.multiply(sigmoid_actder(z[l-1]), W[l] *
delta[l])
# Check delta calculation
if False:
print('Target: %f' % y_this)
print('y_hat: %f' % a[L][0,0])
print(db)
y_pred = ff(x_this,W,b)
diff = 1e-3
W[1][10,0] = W[1][10,0] + diff
y_pred_db = ff(x_this,W,b)
L1 = loss(y_this,y_pred)
L2 = loss(y_this,y_pred_db)
db_finite_difference = (L2-L1)/diff
print('Original out %f, perturbed out %f' %
(y_pred[0,0], y_pred_db[0,0]))
print('Theoretical dW %f, calculated db %f' %
(dW[1][10,0], db_finite_difference[0,0]))
for l in range(1,L+1):
W[l] = W[l] - alpha * dW[l]
b[l] = b[l] - alpha * db[l]
cost_arr.append(loss_this_iter[0,0])
print('Epoch %d train loss %f' % (iter, loss_this_iter))
plt.plot(np.arange(1,max_iter+1,1), cost_arr)
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.show()
# Get test set accuracy
def predict_y(W, b, X):
M = X.shape[0]
y_pred = np.zeros(M)
for i in range(X.shape[0]):
y_pred[i] = np.argmax(ff(X[i,:].T, W, b))
return y_pred
y_test_predicted = predict_y(W, b, X_test)
y_correct = y_test_predicted == y_test_indices
test_accuracy = np.sum(y_correct) / len(y_correct)
print('Test accuracy: %.4f' % (test_accuracy))
# -
# ## In-class exercise
#
# Modify the code above to plot both training loss and test loss as a function of epoch number.
# Use early stopping to obtain the best model according to the validation set.
# Experiment with the hyperparameters (learning rate, number of layers, number of units per layer) to get
# the best result you can.
#
# Describe your experiments and results in your lab report.
#
# ## PyTorch tutorial
#
# Is there an easier way to build this type of model? One way is to learn a framework such as TensorFlow or PyTorch. Both of these frameworks have their
# pros and cons, but PyTorch is probably the most productive neural network framework for research purposes. We'll use it here.
#
# The material for this tutorial is from
# [<NAME>aha's PyTorch tutorial](https://github.com/anandsaha/deep.learning.with.pytorch).
# ### Tensors and Tensor operations
#
# Let's get some hands on experience with tensor creation and operations.
# The torch package contains the necessary data structures to create multidimensional tensors.
# It also defines the mathematical operations that can be performed on these.
import torch
print(torch.__version__)
# ### Tensor creation
# Create a (2x3) dimentional Tensor.
#
# Note that a) You get back a FloatTensor b) The values are uninitialized
t = torch.Tensor(2, 3)
print(t)
# The above call was equivalent to
t = torch.FloatTensor(2, 3)
print(t)
# Inspect type of an element
t[0][0]
type(t[0][0])
# Inspect `t`'s dimensions
print(t.size())
print(t.dim())
print(len(t.size()) == t.dim())
# Set values
t[0][0] = 1
t[0][1] = 2
t[0][2] = 3
t[1][0] = 4
t[1][1] = 5
t[1][2] = 6
print(t)
# Let's cast a FloatTensor to IntTensor
t = torch.FloatTensor([1.1, 2.2])
print(t)
t.type(torch.IntTensor)
# Let's explore some other ways of creating a tensor
# +
# From another Tensor
t2 = torch.Tensor(t)
print(t2)
# +
# From a Python list
t3 = torch.IntTensor([[1, 2],[3, 4]])
print(t3)
# +
# From a NumPy array
import numpy as np
a = np.array([55, 66])
t4 = torch.Tensor(a)
print(t4)
# +
# Create a Tensor with all zeros
t5 = torch.zeros(2, 3)
print(t5)
# +
# Create a Tensor with all ones
t6 = torch.ones(2, 3)
print(t6)
# +
# Create a Tensor with all ones with dimensions
# of another Tensor
t7 = torch.ones_like(t4)
print(t7)
# -
# ### Tensor operations
# Add two Tensors
t1 = torch.ones(2, 2)
t2 = torch.ones(2, 2)
t = t1 + t2
print(t)
# Inplace/out-of-place operations
t1.add(t2)
print(t1)
t1.add_(t2)
print(t1)
# Class methods and package functions
t1.cos()
torch.cos(t1)
# A few more operations
# +
# Create a one-dimensional tensor of steps equally
# spaced points between start and end
torch.linspace(3, 10, steps=5)
# +
# Create a 1-D Tensor with values from [start, end)
torch.arange(0, 5)
# +
# Create a (2x3) Tensor with random values sampled
# from uniform distrubution on the interval [0,1)
torch.rand((2,3))
# +
# Create a (2x3) Tensor with random values sampled
# from normal distrubution with 0 mean and variance 1
torch.randn((2,3))
# +
# Do a matrix multiply
a = torch.rand((2, 3))
b = torch.rand((3, 2))
torch.mm(a, b)
# -
# ### Variables
#
# Next, let's understand variables in PyTorch and the operations we can perform on them.
import torch
from torch.autograd import Variable
# Let's create a small computation graph
x = Variable(torch.FloatTensor([11.2]), requires_grad=True)
y = 2 * x
print(x)
print(y)
print(x.data)
print(y.data)
print(x.grad_fn)
print(y.grad_fn)
y.backward() # Calculates the gradients
print(x.grad)
print(y.grad)
# ### Working with PyTorch and NumPy
import torch
import numpy as np
# Convert a NumPy array to Tensor
n = np.array([2, 3])
t = torch.from_numpy(n)
print(n)
print(t)
# Change a Tensor value, and see the change in
# corresponding NumPy array
n[0] = 100
print(t)
# Convert a Tensor to NumPy array
t = torch.FloatTensor([5, 6])
n = t.numpy()
print(t)
print(n)
# Change a Tensor value, and see the change in corresponding NumPy array
t[0] = 100
print(n)
# ### Tensors on GPU
# Check if your machine has GPU support
if torch.cuda.is_available():
print("GPU Supported")
else:
print("GPU Not Supported")
# Check the number of GPUs attached to this machine
torch.cuda.device_count()
# Get device name
torch.cuda.get_device_name(0)
# Moving a Tensor to GPU
t = torch.FloatTensor([2, 3])
print(t)
t = t.cuda(0)
# Creating a Tensor on GPU, directly
t = torch.cuda.FloatTensor([2, 3])
print(t)
# Bring it back to CPU
t = t.cpu()
print(t)
# Use device context
with torch.cuda.device(0):
t = torch.cuda.FloatTensor([2, 3])
print(t)
# + [markdown] _cell_guid="ff5c0b1b-7df0-49e9-ac50-ffcb7c7ce68b" _uuid="d32496f0fda13237f626529660ef77ca6812abfe"
#
# ## MNIST digit recognition using PyTorch
#
# This part of the lab was
# taken from the [Kaggle tutorial on MNIST with PyTorch]('https://www.kaggle.com/justuser/mnist-with-pytorch-fully-connected-network).
#
# We will use a fully connected neural network and a batch learning algorithm and explain each
# step along the way.
#
# So, with that being said, let's start with imports that we will need.
# First of all, we need to import PyTorch. There are some common names for torch modules (like numpy is always named np): torch.nn.functional is imported as F, torch.nn is the core module, and is simply imported as nn.
# Also, we need numpy. We also use pyplot and seaborn for visualization, but they are not required for the network itself.
# And finally, we use pandas for importing and transforming data.
#
# + _cell_guid="9c846800-05b5-40e4-9899-e3c5e6a149ba" _uuid="2945bec027bb7b48141bc406ccb6ecb19d240aaf"
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Variable
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
# + [markdown] _cell_guid="7ef86f98-c394-4595-8675-e2539ea2332d" _uuid="dc797b7df7d886a974e0fd36a7e30da01159875b"
# Now we can import and transform the data. I decided to split it into input and labels right away at this step:
# + _cell_guid="6d14bbd6-47d0-4b67-b3d4-d2cf21d46aab" _uuid="1bf6c2fde0bb1799d2c551cdeda74bb2a5e02bca"
print("Reading the data...")
data = pd.read_csv('train_mnist.csv', sep=",")
test_data = pd.read_csv('test_mnist.csv', sep=",")
print("Reshaping the data...")
dataFinal = data.drop('label', axis=1)
labels = data['label']
dataNp = dataFinal.to_numpy()
labelsNp = labels.to_numpy()
test_dataNp = test_data.to_numpy()
print("Data is ready")
# + [markdown] _cell_guid="728ef5da-7b0f-4a32-9a0e-10d75e10173e" _uuid="d8441dfce526382bc0c9a0658add9df793f06ed1"
# Now that data is ready, we can take a look at what we're dealing with. I will be using heatmaps from seaborn, which is an excellent tool for matrix visualization. But first, since the images in the MNIST dataset are represented as a long 1d arrays of pixels, we will need to reshape it into 2d array. That's where .reshape() from numpy comes in handy. The pictures are 28 x 28 pixels, so these will be the parameters.
#
# Let's select a couple random samples and visualize them. I will also print their labels, so we can compare images with their actual value:
# + _cell_guid="54405d05-313e-48b9-8984-14337df18c7d" _uuid="7c796bf8cece8a8fef7ab459c0e7624159f2f1e2"
plt.figure(figsize=(14, 12))
pixels = dataNp[10].reshape(28, 28)
plt.subplot(321)
sns.heatmap(data=pixels)
pixels = dataNp[11].reshape(28, 28)
plt.subplot(322)
sns.heatmap(data=pixels)
pixels = dataNp[20].reshape(28, 28)
plt.subplot(323)
sns.heatmap(data=pixels)
pixels = dataNp[32].reshape(28, 28)
plt.subplot(324)
sns.heatmap(data=pixels)
pixels = dataNp[40].reshape(28, 28)
plt.subplot(325)
sns.heatmap(data=pixels)
pixels = dataNp[52].reshape(28, 28)
plt.subplot(326)
sns.heatmap(data=pixels)
print(labels[10], " / ", labels[11])
print(labels[20], " / ", labels[32])
print(labels[40], " / ", labels[52])
# + [markdown] _cell_guid="b7a5f805-f8dd-4288-b44d-8984f479ae7f" _uuid="147ebd083edc2cf96c8e0aefb3893dc971fc5f0b"
# PyTorch has it's own way to store data - those are called tensors, and they are just like numpy arrays, but are suited for PyTorch needs. If we want to feed the data to the network, we need to transform the dataset into those tensors. The good news is that PyTorch can easily do that by transforming numpy arrays or regular lists into tensors.
# + _cell_guid="410167b2-846d-4363-831b-84349053ecfd" _uuid="af8849f388dd69082bdcf52ea48547eb0ea1a483"
x = torch.FloatTensor(dataNp.tolist())
y = torch.LongTensor(labelsNp.tolist())
# + [markdown] _cell_guid="4131196e-07e4-49a5-8380-f0fcfccba081" _uuid="fcd60bddacdb2146d7d3fa272eb3dc766d8314fe"
# Before we start writing the actual network, we need to determine what will be the hyperparameters. Those will not be adjusted during training, so we need to be careful how we set them up.
#
# Here's what we will specify:
# * **input_size** - size of the input layer, it is always fixed (784 pixels)
# * **output_size** - size of the output layer, also fixed size (10 for every possible digit)
# * **hidden_size** - size of the hidden layer, this parameter determines structure of the network. 200 worked for me, but it is worth to play with this parameter to see what works for you
# * **epochs** - how many times will the network go through the entire dataset during training.
# * **learning_rate** - determines how fast will the network learn. You should be very careful about this parameter, because if it is too high, the network won't learn at all, if it is too low, the net will learn too long. I's always about balance. Usualy 10^-3 - 10^-5 works just fine.
# * **batch_size** - size of mini batches during training
# + _cell_guid="80dd6a9a-c28f-4a16-b8fe-6c2c4986422c" _uuid="fdc55b6ed97d0d641046f977cb7a578ad80fbc57"
# hyperparameters
input_size = 784
output_size = 10
hidden_size = 200
epochs = 20
batch_size = 50
learning_rate = 0.00005
# + [markdown] _cell_guid="4d2b9f6c-8fb0-4468-b7ce-88cb3d28fcef" _uuid="507d4fb6939a3a9093ff3eabf1dd841ba5880b97"
# Now we can finally write the actual network. To make it all work, the Network class needs to inherit the *nn.Module*, which gives it the basic functionality required, and allows PyTorch to work with it as expected.
#
# When writing a PyTorch neural network, some things must always be there:
# * \__init\__(self) - initializes the net and creates an instance of that *nn.Module*. Here we define the structure of the network.
# * forward(self, x) - defines forward propagation and how the data flow through the network. Of course, it is based on the structure that is defined in the previous function.
#
# In the initialization, first of all, we need to initialize super (or base) module that the net inherits. After that first line, is the definition of structure. You can experiment with (put more layers or change hidden layer size, etc.), but this structure worked for me just fine.
#
# In forward propagation we simply reassign the value of x as it flows through the layers and return the [softmax](https://en.wikipedia.org/wiki/Softmax_function) at the end.
# + _cell_guid="f238d14f-05bb-4a49-b031-5eaa47bddc64" _uuid="dc77322a324a27ef7a7f30b0432bfd0d3da3e567"
class Network(nn.Module):
def __init__(self):
super(Network, self).__init__()
self.l1 = nn.Linear(input_size, hidden_size)
self.relu = nn.ReLU()
self.l3 = nn.Linear(hidden_size, output_size)
def forward(self, x):
x = self.l1(x)
x = self.relu(x)
x = self.l3(x)
return F.log_softmax(x)
# + [markdown] _cell_guid="02a3fe74-b0d4-4d78-bcee-4fffc6acca89" _uuid="e4dbac71f5f491cf8e79b127cbe18a4e90c65d71"
# After we've defined the network, we can initialize it.
# Also, if we "print" the instance of the net, we can see the structure of it in a neat format:
# + _cell_guid="2376a5b3-3f93-46ac-8e19-e22cf3fb3f01" _uuid="b70b7208f45abe5b9324735a515c6259cc4fb3e3"
net = Network()
print(net)
# + [markdown] _cell_guid="75eaeb97-f840-4862-8819-109b023660a7" _uuid="4cb26c4f84a53abdd5f4c3f7fc0a625ef7d805c4"
# Now it's time to set up the [optimizer](http://pytorch.org/docs/master/optim.html) and a loss function.
#
# *There are quite a lot of things happening behind these two lines of code, so if you don't know what is going on here, don't worry too much for now, it will get clearer eventualy.*
#
# Optimizer is what updates the parameters of the network. I will be using Stochastic Gradient Descent with momentum. Also, the optimizer takes the network parameters as an argument, but it's not a big deal since we can get those with .parameters() function.
#
# I decided to use [Cross Entropy Loss](https://en.wikipedia.org/wiki/Cross_entropy) for this problem, but again, there are many options and you are free to choose whatever suits you best.
# + _cell_guid="f9fe7b0f-7fec-49f3-8cd2-a6fedd4c8048" _uuid="d4fd65ae06c5003428ab90b1557142f015152b7c"
optimizer = optim.SGD(net.parameters(), lr=learning_rate, momentum=0.9)
loss_func = nn.CrossEntropyLoss()
# + [markdown] _cell_guid="465d2e54-0a22-446d-9f87-d687f48140f3" _uuid="9684f943703fb995be47a7d92d1d281acc3dae32"
# Now that everything is ready, our network can start learning. I will separate data into minibatches and feed it to the network. It has many advantages over single batch learning, but that is a different story.
#
# Also, I will use loss_log list to keep track of the loss function during the training process.
# + _cell_guid="3b646efc-4cae-4f94-9350-e059e4c89446" _uuid="c1247ff34caf53e9894a1c2ea5c435c0e446d048"
loss_log = []
for e in range(epochs):
for i in range(0, x.shape[0], batch_size):
x_mini = x[i:i + batch_size]
y_mini = y[i:i + batch_size]
x_var = Variable(x_mini)
y_var = Variable(y_mini)
optimizer.zero_grad()
net_out = net(x_var)
loss = loss_func(net_out, y_var)
loss.backward()
optimizer.step()
if i % 100 == 0:
loss_log.append(loss.item())
print('Epoch: {} - Loss: {:.6f}'.format(e, loss.item()))
# + [markdown] _cell_guid="548ac762-97c0-48b9-98ff-93559b293c9a" _uuid="e3e6d56f52974ccb3120340695aed4b9bea0907b"
# So, let's go line by line and see what is happening here:
#
# This is the main loop that goes through all the epochs of training. An epoch is one full training on the full dataset.
#
# for e in range(epochs):
# This is the inner loop that simply goes through the dataset batch by batch:
#
# for i in range(0, x.shape[0], batch_size):
# Here is where we get the batches out of our data and simply assign them to variables for further work:
#
# x_mini = x[i:i + batch_size]
# y_mini = y[i:i + batch_size]
# These two lines are quite *important*. Remember I told you about tensors and how PyTorch stores data in them? That's not the end of story. Actually, to allow the network to work with data, we need a wrapper for those tensors called Variable. It has some additional properties, like allowing automatic gradient computation when backpropagating. It is required for the proper work of PyTorch, so we will add them here and supply tensors as parameters:
#
# x_var = Variable(x_mini)
# y_var = Variable(y_mini)
# This line just resets the gradient of the optimizer:
#
# optimizer.zero_grad()
# Remember the *forward(self, x)* function that we previously defined? The next line is basically calling this function and does the forward propagation:
#
# net_out = net(x_var)
# This line computes the loss function based on predictions of the net and the correct answers:
#
# loss = loss_func(net_out, y_var)
# Here we compute the gradient based on the loss that we've got. It will be used to adjust parameters of the network.
#
# loss.backward()
# And here is where we finally update our network with new adjusted parameters:
#
# optimizer.step()
# The rest is just logging, which might be helpful to observe how well the network is performing.
#
# After the network is done with training, we can take a look at the loss function, and how it behaved during training:
# + _cell_guid="5073f433-dbae-452b-8b7f-647e20c2d0dc" _uuid="4b635e76f4bb761ceef632c381546a2286442818"
plt.figure(figsize=(10,8))
plt.plot(loss_log)
# + [markdown] _cell_guid="91f7e895-9ff7-4363-82d4-c2dc847aa9c3" _uuid="c5e3981a9d1e16ad5b8dcf5916031a22ac3336db"
# At this point, the network should be trained, and we can make a prediction using the test dataset. All we need to do is wrap the data into the Variable and feed it to the trained net, so nothing new here.
# + _cell_guid="9792a7ca-d5a8-499b-87d0-077ecf77685d" _uuid="72b3a1b85275cc4c1f5bc7f442bf81004014b118"
test = torch.FloatTensor(test_dataNp.tolist())
test_var = Variable(test)
net_out = net(test_var)
print(torch.max(net_out.data, 1)[1].numpy())
# + [markdown] _cell_guid="f34cc0fb-c950-4346-b742-259b86d9374c" _uuid="fed2362db49fc9991a1fd0be98d29c55b03b6b82"
# Now we have out predictions that are ready to be submitted. Before that, we can take a look at predictions and compare them to the actual pictures of digits, just like at the start with training data:
# + _cell_guid="bf0126d4-3eab-45ce-b84e-2f07c8fe9663" _uuid="41378dcf1951b3cbb7f929bec0a288bad007b72d"
plt.figure(figsize=(14, 12))
pixels = test_dataNp[1].reshape(28, 28)
plt.subplot(321)
sns.heatmap(data=pixels)
test_sample = torch.FloatTensor(test_dataNp[1].tolist())
test_var_sample = Variable(test_sample)
net_out_sample = net(test_var_sample)
pixels = test_dataNp[10].reshape(28, 28)
plt.subplot(322)
sns.heatmap(data=pixels)
test_sample = torch.FloatTensor(test_dataNp[10].tolist())
test_var_sample = Variable(test_sample)
net_out_sample = net(test_var_sample)
pixels = test_dataNp[20].reshape(28, 28)
plt.subplot(323)
sns.heatmap(data=pixels)
test_sample = torch.FloatTensor(test_dataNp[20].tolist())
test_var_sample = Variable(test_sample)
net_out_sample = net(test_var_sample)
pixels = test_dataNp[30].reshape(28, 28)
plt.subplot(324)
sns.heatmap(data=pixels)
test_sample = torch.FloatTensor(test_dataNp[30].tolist())
test_var_sample = Variable(test_sample)
net_out_sample = net(test_var_sample)
pixels = test_dataNp[100].reshape(28, 28)
plt.subplot(325)
sns.heatmap(data=pixels)
test_sample = torch.FloatTensor(test_dataNp[100].tolist())
test_var_sample = Variable(test_sample)
net_out_sample = net(test_var_sample)
pixels = test_dataNp[2000].reshape(28, 28)
plt.subplot(326)
sns.heatmap(data=pixels)
test_sample = torch.FloatTensor(test_dataNp[1].tolist())
test_var_sample = Variable(test_sample)
net_out_sample = net(test_var_sample)
print("Prediction: {} / {}".format(torch.max(net_out.data, 1)[1].numpy()[1], torch.max(net_out.data, 1)[1].numpy()[10]))
print("Prediction: {} / {}".format(torch.max(net_out.data, 1)[1].numpy()[20], torch.max(net_out.data, 1)[1].numpy()[30]))
print("Prediction: {} / {}".format(torch.max(net_out.data, 1)[1].numpy()[100], torch.max(net_out.data, 1)[1].numpy()[2000]))
# + _cell_guid="22c40d7b-a3a4-4118-9aa5-d5191876ecd1" _uuid="c60e5abb1be0166b2c22aaad560885a74db21da9"
output = (torch.max(net_out.data, 1)[1]).numpy()
#np.savetxt("out.csv", np.dstack((np.arange(1, output.size+1),output))[0],"%d,%d",header="ImageId,Label")
# + [markdown] _cell_guid="13d2eeae-86ea-499e-923a-642527f23ce0" _uuid="97523e84c3d8ee0be9aea776f3f7e6c109f05cdd"
# And that is about it, we've made a simple neural network using PyTorch that can recognize handwritten digits. Not so bad!
#
# When I was writing this notebook, this model scorred 96.6%, which is not perfect by any means, but it's not that bad either.
#
# I hope this was useful for some of you. If you are totally new to deep learning, I suggest you learn how the neural networks actually work from the inside, especially the backpropagation algorithm.
#
# These videos explain [neural nets](https://www.youtube.com/watch?v=aircAruvnKk&t=708s) and [backpropagation](https://www.youtube.com/watch?v=Ilg3gGewQ5U) quite well.
#
# Also I suggest you to take a look at this [online book](http://neuralnetworksanddeeplearning.com/chap1.html) (it's absolutely free, btw), where neural networks are explained in great detail, and it even has an implementation of the MNIST problem from scratch, using only numpy.
#
# If you have any feedback, feel free to leave comments down below, and good luck with your deep learning adventures :)
# -
# ## Take-home exercise
#
# Make sure you can run the PyTorch examples of MNIST classification, then apply the PyTorch example to another
# classification problem you've worked with this semester, for example the breast cancer dataset. Get familiar with
# working with models in PyTorch.
#
# Report your experiments and results in your brief lab report.
| Labs/09-DeepLearning-I/09-DeepLearning-I.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# ### NetworKIN Result Files
# - See Code/PreprocessingPredictionData/Running_NetworKIN.md for instructions on installing and running NetworKIN for preparing the initial Raw NetworKIN results file
#
# ### About the NetworKIN Raw Results File
# - Version: Last Run in KinPred Nov 2020
# - NetworKIN result files contain the following columns:
# - #Name
# - Position
# - Tree
# - NetPhorest Group
# - Kinase/Phosphatase/Phospho-binding domain
# - NetworKIN score
# - NetPhorest probability
# - STRING score
# - Target STRING ID
# - Kinase/Phosphatase/Phospho-binding domain
# - STRING ID
# - Target description
# - Kinase/Phosphatase/Phospho-binding domain description
# - Target Name
# - Kinase/Phosphatase/Phospho-binding domain Name
# - Peptide sequence window
# - Intermediate nodes
#
# ### Preprocessing NetworKIN Raw
# 1. Filtering Date: only keep the results of kinase prediction (TREE == 'KIN')
# 2. Mapping accessions:
# - **Mapping substrate accessions:** Get the UniprotID from the sequence identifiers of the original fasta files that submitted in NetworKIN for prediction.
# - **Mapping kinase accessions:** get uniprotID for the kinases using 'Kinase/Phosphatase/Phospho-binding domain description' column
# - correcting duplicated names found above
# 3. **Mapping sites:**
# - formatting pep seq (e.g. ----MsGSKSV --> ____MSGSKSV)
# - update 'site' if the orignal sequence submitted for perdiction is different from the reference sequence due to changes causing shift in position
#
# **Output Files Dataframe:**
# - **substrate_id:** unique IDs for the substrate phosphorylation site (substrate_acc + position)
# - **substrate_name:** substrate gene name used in NetworKIN
# - **substrate_acc:** substrate uniprotID
# - **site:** aa + position in protein sequence
# - **Position:** position in protein sequence
# - **pep:** +/- 5 AA
# - **kinase_name:** kinase name used in NetworKIN
# - **kinase_acc:** kinase uniprotID
# - **score:** NetworKIN score
#
#
#
# ### Creating Resource Files
# 1. **'globalKinaseMap.csv':**
# - creat a new or add unique kinases from NetworKIN to the globel substrate resource file.
# - get and add the Kinase Name that would use across all perdictors for the kinases to the result files
#
# ### Standard Formatted NetworKIN
# **'NetworKIN_formatted.csv':** Standardize the preprocessed file with following columns:
# - **substrate_id** - unique IDs for the substrate phosphorylation site (substrate_acc + position)
# - **substrate_name** - gene name for the substrates
# - **substrate_acc** - mapped UniprotIDs for the substrates
# - **site** - phosphorylation site
# - **pep** - +/- 5 AA peptide sequence around the site
# - **score**
# - **Kinase Name**
#
# +
# IMPORTS
import pandas as pd
import os
import re
import glob
import humanProteomesReference, networKin_convert, getUniprotID, checkSite
#only need when testing the code
import time
# +
# DEFINE FILE NAMES/DIRs
##################
# Version (Date) #
##################
version = '2019-12-11'
##################
# File Location #
##################
# local (../../)
base = '../../'
##################################################
# For Prepare Fasta Files to Submit in NetworKIN #
##################################################
# Human Proteome fasta file
HP_fasta = base + 'Data/Raw/HumanProteome/humanProteome_' + version + '.fasta'
# Dir for splited Human Proteome fasta files
HP_dir = base + 'Data/Raw/HumanProteome/'
# human proteome referece file
HP_csv = base + 'Data/Map/humanProteome_' + version + '.csv'
####################################################
# For Preprocessing NetworKIN Prediction Results #
#--------------------------------------------------#
# . The files submitted for NetworKIN predictor is #
# NOT the up-to-date human proteom sequences #
# . There has been an update in human proteomes #
# from the time the perdiction results were got #
# to running the preporcessing steps. #
####################################################
# NetworKIN results dir
NW_dir = base + 'Data/Raw/NetworKIN/'
NW_update_dir = base + 'Data/Raw/NetworKIN/updated/'
# NetworKIN temp dir
NW_temp_dir_acc = base + 'Data/Temp/NetworKIN/mappedAcc/'
NW_temp_dir_site = base + 'Data/Temp/NetworKIN/mappedSite/'
NW_temp_dir_acc_update = base + 'Data/Temp/NetworKIN/mappedAcc/updated/'
NW_temp_dir_site_update = base + 'Data/Temp/NetworKIN/mappedSite/updated/'
# Resource Files
HK_org = base + 'Data/Raw/HumanKinase/globalKinaseMap.txt' # orginal manually created kinase file
KinaseMap = base + 'Data/Map/globalKinaseMap.csv' # add all unique kinase in HPRD to the global file
# Standard formatted output file
NW_formatted = base + 'Data/Formatted/NetworKIN/NetworKIN_formatted_' + version + '.csv' # preprocessed file with cloumns: substrate_id/substrate/substrate_acc/kinase/site/pep/score
# -
# # Preprocessing NetworKIN Raw
# In the NetworKIN raw file, most of the names in the 'Kinase/Phosphatase/Phospho-binding domain' column can not use to retrieve the uniprotID for the kinase, but all the names in the 'Kinase/Phosphatase/Phospho-binding domain description' column can. However, there are different 'Kinase/Phosphatase/Phospho-binding domain' with the same 'Kinase/Phosphatase/Phospho-binding domain description'. We need to identify those and correct them if need.
# ### Get unique kinases in NetworKIN
# +
all_results = glob.glob(NW_dir + '*.tsv')
#create empty df to store unique kinases
df_unique_kin = pd.DataFrame()
for filename in all_results:
df = pd.read_csv(filename, usecols = ['Tree','Kinase/Phosphatase/Phospho-binding domain', 'Kinase/Phosphatase/Phospho-binding domain description'], sep = '\t')
# the only type of perdiction we are intreseted in is 'KIN'
df = df[df.Tree == 'KIN']
df = df[['Kinase/Phosphatase/Phospho-binding domain', 'Kinase/Phosphatase/Phospho-binding domain description']].drop_duplicates()
# append unique kinases found in each result files
df_unique_kin = df_unique_kin.append(df)
# drop any duplicated kinases
df_unique_kin = df_unique_kin.drop_duplicates()
# get the kinase(s) with the same domain description
duplicateRowsDF = df_unique_kin[df_unique_kin.duplicated(['Kinase/Phosphatase/Phospho-binding domain description'],keep=False)]
duplicateRowsDF.sort_values(['Kinase/Phosphatase/Phospho-binding domain description','Kinase/Phosphatase/Phospho-binding domain'])
# -
# After manually check for the above kinases, MST4 should associate with STK26 not STK3. Create a dictionary to correct that in the result files.
correct_kinase = {'MST4' : 'STK26'}
# ### Mapping Accessions (UniprotID) and Site
# 1. Filtering Date: only keep the results of kinase prediction (TREE == 'KIN')
# 2. Mapping accessions:
# - **Mapping substrate accessions:** Get the UniprotID from the sequence identifiers of the original fasta files that submitted in NetworKIN for prediction.
# - **Mapping kinase accessions:** get uniprotID for the kinases using 'Kinase/Phosphatase/Phospho-binding domain description' column
# - correcting duplicated names found above
# 3. **Mapping sites:**
# - formatting pep seq (e.g. ----MsGSKSV --> ____MSGSKSV)
# - update 'site' if the orignal sequence submitted for perdiction is different from the reference sequence due to changes causing shift in position
#
# **Output Files Dataframe:**
# - **substrate_id:** unique IDs for the substrate phosphorylation site (substrate_acc + position)
# - **substrate_name:** substrate gene name used in NetworKIN
# - **substrate_acc:** substrate uniprotID
# - **site:** aa + position in protein sequence
# - **Position:** position in protein sequence
# - **pep:** +/- 5 AA
# - **kinase_name:** kinase name used in NetworKIN
# - **kinase_acc:** kinase uniprotID
# - **score:** NetworKIN score
# **Mapping Accessions**
# convert substrate_acc and kinase_acc
convert_type = 'acc'
networKin_convert.kin_convert_directory(NW_dir, 'na', NW_temp_dir_acc, convert_type)
# **Mapping Site**
# map the site to the updated (new) human proteome reference
convert_type = 'site'
networKin_convert.kin_convert_directory(NW_temp_dir_acc, HP_csv, NW_temp_dir_site, convert_type)
# **Remove unmapped/outdated results**
# - Check if there is any unmapped substrate/site due to outdated uniprot sequence records
# - get a list of outdated uniprotID
# +
all_results = glob.glob(NW_temp_dir_site + '*.csv')
updatedSub_li = []
for filename in all_results:
start = time.time()
df_unmapped = pd.read_csv(filename, usecols = ['substrate_id','substrate_acc', 'site'])
df_unmapped = df_unmapped[~(df_unmapped['site'].str.contains('S|T|Y', na=False)) | (df_unmapped['substrate_id'] == 'outdated')]
df_unmapped = df_unmapped.substrate_acc.drop_duplicates()
unmapped_li = df_unmapped.values.tolist()
updatedSub_li.extend(unmapped_li)
end = time.time()
print (f"Time\t{(end-start):.3f}")
updatedSub_li
# -
# - Remove any record with unmapped substrate_acc/site in '*_mappedSite.csv'
# +
all_results = glob.glob(NW_temp_dir_site + '*.csv')
for filename in all_results:
start = time.time()
df_mapSite = pd.read_csv(filename)
# remove the outdated records from df_subMap
df_update = df_mapSite[~df_mapSite['substrate_acc'].isin(updatedSub_li)]
df_update.to_csv(filename, chunksize=100000, index=False)
end = time.time()
print (f"chunk time\t{(end-start):.3f}")
# -
# ### Update NetworKIN results
# - The next 6 cells is only for NetworKIN results from outdated Human Proteomes sequences (input sequences for NetworKIN prediction an earlier version than the referece human proteome sequence)
# 1. download the sequence fasta file of the above UniprotID (updatedSub_li) from Uniprot.org, save it as '../Data/Raw/HumanProteome/NetworKIN_updateSub.fasta'. Submit the 'NetworKIN_updateSub.fasta' in NetworKIN again
# 2. run mapAcc and mapSite function for the result file from 'NetworKIN_updateSub.fasta'
# convert substrate_acc and kinase_acc
convert_type = 'acc'
networKin_convert.kin_convert_directory(NW_update_dir, 'na', NW_temp_dir_acc_update, convert_type)
# map the site to the updated (new) human proteome reference
convert_type = 'site'
networKin_convert.kin_convert_directory(NW_temp_dir_acc_update, HP_csv, NW_temp_dir_site_update, convert_type)
# 3. save '2020_mappedSite.csv' under the same dir as other *_mappedSite.csv files
# - NetworKIN ignorgs the 'U'(Selenocysteine) in some substrate sequences, this causes frame shift of the downstream sequences
# - those unmapped sites will not be included
start = time.time()
df_mapUpdateSite = pd.read_csv(NW_temp_dir_site_update + '2020_mappedSite.csv')
df_mapUpdateSite = df_mapUpdateSite[(df_mapUpdateSite['site'].str.contains('S|T|Y', na=False)) & (df_mapUpdateSite['substrate_id'] != 'outdated')]
df_mapUpdateSite.to_csv(NW_temp_dir_site + '2020_mappedSite.csv')
end = time.time()
print (f"Time\t{(end-start):.3f}")
df_mapUpdateSite
# ### Get the Gene Name of the Substrates from the Reference Human Proteome
# - get and add the Gene Name that would use across all perdictors for the substrates to the result files
# get the protein gene names and accessions from the Reference Human Proteome
df_unique_sub = pd.read_csv(HP_csv, usecols = ['UniprotID','Gene Name'], sep = '\t')
df_unique_sub
# +
# add the Gene Name that would use across all perdictors for the substrates to the result files
start = time.time()
all_results = glob.glob(NW_temp_dir_site + '*.csv')
for filename in all_results:
df = pd.read_csv(filename, usecols = ['substrate_id', 'substrate_acc', 'substrate_name', 'site', 'Position', 'pep', 'score', 'kinase_name', 'kinase_acc'])
# merge df_subsMap with df_unique_sub to add the common substrate gene name to the df
df = df.merge(df_unique_sub, left_on=['substrate_acc'], right_on=['UniprotID'], how = 'left')
df = df.drop(columns = ['UniprotID'])
df.to_csv(filename, index=False)
df
# -
# ### Creating Resource Files
# **globalKinaseMap**
# - creat a new or add unique kinases from NetworKIN to the globel kinase resource file.
# - get and add the Kinase Name that would use across all perdictors for the kinases to the result files
# +
# get unique kinases in the NetworKIN result files
all_results = glob.glob(NW_temp_dir_site + '*.csv')
df_unique_kin = pd.DataFrame()
for filename in all_results:
df = pd.read_csv(filename, usecols = ['kinase_acc', 'kinase_name'])
df = df.drop_duplicates()
df_unique_kin = df_unique_kin.append(df, ignore_index=True)
df_unique_kin = df_unique_kin.drop_duplicates()
df_unique_kin
# +
# start = time.time()
unmapped_list = pd.DataFrame()
if os.path.isfile(KinaseMap):
df_humanKinase = pd.read_csv(KinaseMap)
# if globalkinaseMap.csv file does not exist, create an new df using orginal human kinase map
else:
df_humanKinase = pd.read_csv(HK_org, usecols = ['Kinase Name', 'Preferred Name', 'UniprotID', 'Type', 'description'], sep = '\t')
df_humanKinase['description'].replace(regex=True,inplace=True,to_replace=r'\[Source.+\]',value=r'')
# add unique kinases from NetworKIN to the globel kinases resource file
for index, row in df_unique_kin.iterrows():
kinase = df_unique_kin.at[index, 'kinase_acc']
# if the kinase/other enzyme already in the globalKinaseMap.csv file
if any(df_humanKinase.UniprotID == kinase):
# get the index of the substrate in the globalKinaseMap.csv file
idx = df_humanKinase.index[df_humanKinase.UniprotID == kinase].values[0]
df_humanKinase.at[idx, 'NetworKIN_kinase_name'] = df_unique_kin.at[index, 'kinase_name']
# if the kinase is not in the globalKinaseMap.csv file, we need a list to check annotations manullay
else:
unmapped_list = unmapped_list.append(row,sort=False).reset_index(drop=True)
print (unmapped_list)
# -
# - Manually check the above unmapped kinase. Create a dictionay for the one(s) that are kinases to add in the globalKinaseMap.csv. Create a list of the one(s) that are not kinases to drop records from the NetworKIN result files.
# +
new_kinase = {'EPHB6':'Ephrin type-B receptor 6'}
not_kinase = ['PDK2','PDK3','PDK4','LCA5']
# -
# - add the 1 protein kinases to the globalKinaseMap.csv using above dictionary
# +
# get the length (last index) of the current df_humanKinase
len = df_humanKinase.UniprotID.count()
# add the 1 new kinase in the globalKinaseMap.csv
for key in new_kinase:
# add the kinase name and description for the new kinase
df_humanKinase = df_humanKinase.append({'Kinase Name': key}, ignore_index=True)
df_humanKinase.at[len,'Preferred Name'] = key
df_humanKinase.at[len,'description'] = new_kinase[key]
# get the index of where the new kinase is in the df_unique_kinase
i = df_unique_kin.index[df_unique_kin['kinase_name'] == key].values[0]
# add the uniprotID for the new kinase
df_humanKinase.at[len,'UniprotID'] = df_unique_kin.at[i, 'kinase_acc']
# add the accs used in NetworKIN for the new kinase
df_humanKinase.at[len,'NetworKIN_kinase_name'] = df_unique_kin.at[i, 'kinase_name']
len += 1
df_humanKinase.to_csv(KinaseMap,index = False)
df_humanKinase
# -
# get the new list kinase with common kinase name that would use across all referece and the uniprotID for these kinase
df_unique_kin = df_humanKinase[['Kinase Name','UniprotID']]
df_unique_kin
# - remove the 4 enzyme records that are not protein kinase from the NetworKIN result files
# - add the Kinase Name that would use across all perdictors for the kinase to the result files
# +
all_results = glob.glob(NW_temp_dir_site + '*.csv')
for filename in all_results:
df = pd.read_csv(filename)
# remove the 4 enzyme records that are not protein kinase from the df_kinaseMap
df = df[~df['kinase_name'].isin(not_kinase)]
# merge with df_unique_kinase to add the common kinase name to the df
df = df.merge(df_unique_kin, left_on='kinase_acc', right_on='UniprotID', how = 'left')
# drop the duplicated uniprot id for kinases
df = df.drop(columns = 'UniprotID')
df.to_csv(filename,index=False)
df
# -
# # Standard Formatted NetWorKIN
# ### 'NetworKIN_formatted.csv'
# Standardize the preprocessed file with following columns:
#
# - substrate_id - unique IDs for the substrate phosphorylation site (substrate_acc + position)
# - substrate_name - gene name for the substrates
# - substrate_acc - mapped UniprotIDs for the substrates
# - site - phosphorylation site
# - pep - +/- 5 AA peptide sequence around the site
# - score - perdiction score
# - Kinase Name - Kinase name
# +
all_results = glob.glob(NW_temp_dir_site + '*.csv')
NW = []
for filename in all_results:
df = pd.read_csv(filename, usecols = ['substrate_id','substrate_acc','Gene Name', 'site','pep', 'score', 'Kinase Name'])
df = df.rename(columns={'Gene Name': 'substrate_name'})
NW.append(df)
df_final = pd.concat(NW)
df_final = df_final.drop_duplicates()
df_final.to_csv(NW_formatted, chunksize = 1000000, index = False)
df_final
# -
| Code/PreprocessingPredictionData/FormattingNetworKIN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from scipy.io import loadmat
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
rois = ['V1', 'V2', 'V3', 'hV4', 'IOG', 'pFus', 'mFus']
def filter_voxels(res, cutoff=50):
# as in Kay et al., select non-noisy voxels with at least 50% variance explained
idx = res['aggregatedtestperformance'][0] >= cutoff
return np.median(res['params'][..., idx], axis=0)
params = dict()
for hemi in ['L', 'R']:
for roi in rois:
ok_voxs = []
for s in range(1, 4):
res = loadmat(f'../output/sub-{s:02d}_{hemi}{roi}.mat')
ok_voxs.append(filter_voxels(res))
ok_voxs = np.hstack(ok_voxs)
params[f'{hemi}{roi}'] = ok_voxs
for roi, val in params.items():
print(f'{roi}: {val.shape[1]}')
params.keys()
# +
# the parameters of the CSS model are [R C S G N] where
# R is the row index of the center of the 2D Gaussian
# C is the column index of the center of the 2D Gaussian
# S is the standard deviation of the 2D Gaussian
# G is a gain parameter
# N is the exponent of the power-law nonlinearity
# +
def eccentricity(x, y, res=100, width_degree=12.5):
m = res/2
scaling = width_degree/res
x = (x-m) * scaling
y = (y-m) * scaling
return np.sqrt(x**2 + y**2)
def prf_size(sd, n, res=100, width_degree=12.5):
scaling = width_degree/res
return scaling * sd/np.sqrt(n)
# -
ecc_size = dict()
for roi, pp in params.items():
ecc_size[roi] = np.vstack((eccentricity(*pp[:2]), prf_size(pp[2], pp[-1])))
fig, axs = plt.subplots(2, 7, figsize=(12, 8))
for i, hemi in enumerate(['L', 'R']):
axs_ = axs[i]
for ax, roi in zip(axs_, rois):
roi_name = f'{hemi}{roi}'
pp = ecc_size[roi_name]
sns.regplot(x=pp[0], y=pp[1], scatter_kws={'s': 2}, ax=ax, )
ax.set_xlim([0, 12.5]);
ax.set_ylim([0, 12.5]);
ax.set_aspect('equal');
ax.set_title(roi_name)
ax.set_xlabel('Eccentricity')
ax.set_ylabel('Size')
plt.tight_layout()
# combine hemispheres
ecc_size_combined = dict()
for roi in rois:
ecc_size_combined[roi] = np.hstack((ecc_size[f'L{roi}'], ecc_size[f'R{roi}']))
ecc_size_combined['V1'].shape
fig, axs = plt.subplots(1, 7, figsize=(12, 8))
for ax, roi in zip(axs, rois):
roi_name = roi
pp = ecc_size_combined[roi_name]
sns.regplot(x=pp[0], y=pp[1], scatter_kws={'s': 2}, ax=ax, )
ax.set_xlim([0, 12.5]);
ax.set_ylim([0, 12.5]);
ax.set_aspect('equal');
ax.set_title(roi_name)
ax.set_xlabel('Eccentricity')
ax.set_ylabel('Size')
plt.tight_layout()
from scipy.stats import linregress
slope_int = dict()
for roi, (ecc, size) in ecc_size_combined.items():
reg = linregress(ecc, size)
slope_int[roi] = reg[:2]
slope_int
fig, ax = plt.subplots(1, 1, figsize=(4, 4))
for roi in rois:
slope, intercept = slope_int[roi]
ax.plot([0, 12.5], [intercept, 12.5*slope + intercept], label=roi)
ax.set_title(hemi)
ax.set_xlim([0, 12.5]);
ax.set_ylim([0, 12.5]);
ax.set_aspect('equal');
ax.set_xlabel('Eccentricity')
ax.set_ylabel('Size')
ax.legend()
fig, axs = plt.subplots(1, 2, figsize=(12, 12))
for ax, hemi in zip(axs, ['L', 'R']):
for roi in rois:
slope, intercept = slope_int[f'{hemi}{roi}']
ax.plot([0, 12.5], [intercept, 12.5*slope + intercept], label=roi)
ax.set_title(hemi)
ax.set_xlim([0, 12.5]);
ax.set_ylim([0, 12.5]);
ax.set_aspect('equal');
ax.set_xlabel('Eccentricity')
ax.set_ylabel('Size')
ax.legend()
# +
median_ecc = dict()
for roi, (ecc, size) in ecc_size.items():
median_ecc[roi] = np.median(ecc)
# -
import pandas as pd
df_ecc = pd.DataFrame(list(median_ecc.items()), columns=['roi', 'eccentricity'])
df_ecc['hemi'] = [r[0] for r in df_ecc.roi]
df_ecc['roi'] = [r[1:] for r in df_ecc.roi]
sns.pointplot(x='roi', y='eccentricity', hue='hemi', data=df_ecc, join=False)
# +
# save parameters for later use
header = ['row', 'col', 'std', 'gain', 'n']
for roi, param in params.items():
np.savetxt(f'../output/{roi}_median_param.txt', param, header=' '.join(header))
| vtcdata/code/check_params.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tirgul 5
# The topics are:
# - Plots
# - Histograms
# - BoxPlots
# - Lots of Fun!
# ## matplotlib - library used for graphics
# 1. For matplotlib video tutorials: https://www.youtube.com/watch?v=UO98lJQ3QGI
# 1. Matplotlib tutorial: https://www.datacamp.com/community/tutorials/matplotlib-tutorial-python
# 1. Matplitlib Docs: https://matplotlib.org/stable/contents.html
#
# + id="npYZUBPHByeI"
# Import matplotlib, we usually work with pyplot
import matplotlib.pyplot as plt
# Plot size in jupyter
plt.rcParams['figure.figsize'] = [10, 5]
# + colab={"base_uri": "https://localhost:8080/", "height": 440} id="UiaPWw0g_LAu" outputId="be1b69ff-8ddc-47b1-b71a-76f0f9896ee7"
xValues = [10,20,30]
yValues = [100,120,110]
fig = plt.figure()
plt.plot(xValues,yValues)
plt.plot([3,4])
#plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 470} id="Yy9tJCjj_LBF" outputId="fea52a57-b96f-4cf8-e4f2-716fa5f286b4"
# We can set titles for x\y labels
plt.title('this is a nice title')
plt.xlabel('x values')
plt.ylabel('y values')
plt.plot(xValues,yValues)
plt.show()
# + id="MSdBhg08DcX6"
# We can save the figure as an image
fig.savefig('my_figure.png') # remember to refresh files display
# + [markdown] id="zE8o_kql_LBG"
# ## Let's plot some practicle plots
# Read the *trees.csv* file and display height vs volume graph
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="diC1v6hy_LBH" outputId="0d7a0553-0671-42f8-f785-3f0aab46d990"
import pandas as pd
url = 'trees.csv'
treesData = pd.read_csv(url)
treesData.head()
# + id="2ttxAubp_LBI"
treesHeight = treesData['Height']
treesVolume = treesData['Volume']
# + colab={"base_uri": "https://localhost:8080/", "height": 470} id="R010zqxo_LBJ" outputId="46d48e75-cbb1-4ba6-8dc1-da45f27b9019"
plt.plot(treesHeight,treesVolume,'bo') # b- is for 'bule', 'o' sets the marker shape to be a circle. For more colors/shapes look a the docs
plt.title('Trees Volume vs Height')
plt.ylabel('Volume')
plt.xlabel('Height')
plt.show()
# -
# ### Display the *boxplot*
# + O is an outlier
# + \- represents the max/min values that are not outliers
# + red line is the medial
# + The box top/bottom represent the 2nd and 3rd quarters of the data
# + colab={"base_uri": "https://localhost:8080/", "height": 440} id="ULEwNBAERGSM" outputId="aa38ba8e-f46f-46fa-96a2-88b3a95cd550"
myData = [treesData['Height'] , treesData['Volume']]
fig=plt.boxplot(myData)
# -
# ### Histogram:
# + colab={"base_uri": "https://localhost:8080/", "height": 487} id="74pyCNAPVT6e" outputId="24400a35-1d4a-4fcd-b8b2-6bc612ddbbf7"
plt.hist([treesData['Height'],treesData['Volume']])
plt.ylabel('# num of appearances')
plt.xlabel('height')
plt.title('Histogram of trees height')
# + [markdown] id="XCl4Tq79_LBK"
# ### Matplotlib excercises:
# https://pynative.com/python-matplotlib-exercise/
#
# + colab={"base_uri": "https://localhost:8080/", "height": 457} id="lrGDAosf_Pw4" outputId="38b685aa-d5aa-44f3-f7b8-27fbff3a93a1"
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0,10*np.pi , 1000)
x
#print("X: {}...{}".format(x[:5],x[-5:]))
y = np.sin(x)
#print("Y: {}...{}".format(y[:5],y[-5:]))
plt.plot(x,y)
# -
# ### Subplots
# + colab={"base_uri": "https://localhost:8080/", "height": 469} id="gmiGW8V8_Sv3" outputId="8ba9acf0-eff8-43e7-986d-7d574231694e"
# We can plot several subplots in one figure
fig = plt.figure() # create a plot figure
# create the first of two panels and set current axis
import numpy as np
x = np.linspace(-2*np.pi, 2*np.pi, 100)
plt.subplot(2, 2, 1) # (rows, columns, panel number)
plt.plot(x, np.sin(x))
plt.title('sin x')
# create the second panel and set current axis
plt.subplot(2, 2, 2)
plt.plot(x, np.cos(x), color = 'red' , linestyle = '--'); # --r will also work
plt.title('cos x',color = 'green', fontsize = 20)
# -
# ### self excercise:
# + id="3FSpvVmtO-O9"
# plot y=x^2 from -10 to 10
# with x and y labels and title and legend
# with dashed yellow line
# + colab={"base_uri": "https://localhost:8080/", "height": 504} id="WT8R3rEzPSDG" outputId="7efadef7-f1ed-4992-efc8-954d4f1d33c0"
x = np.linspace(-10,10,1000)
plt.plot(x,x**2,'--y')
plt.xlabel('x')
plt.ylabel('y=x^2')
plt.title('y=x^2')
# -
# ### Plot X/Y axis limits
# + colab={"base_uri": "https://localhost:8080/", "height": 440} id="1VBZAqqwG32C" outputId="442f4668-7c68-4004-c045-146ce98482b7"
# we can control the axis limits:
plt.plot(x, np.sin(x))
plt.xlim(-4,4)
plt.ylim(-2,2)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 434} id="UG5VUXY-ISdW" outputId="9dcefc8d-7b70-4389-c2a8-3046cb9558c2"
plt.plot(x, np.sin(x))
plt.axis('tight');
# + colab={"base_uri": "https://localhost:8080/", "height": 434} id="oJawspv4IxI2" outputId="e534ed64-030e-44a1-db7b-16264c45787d"
# Set equal ratio
plt.plot(x, np.sin(x))
plt.axis('equal');
# -
# ### Plot Legend
# + colab={"base_uri": "https://localhost:8080/", "height": 440} id="BG5EjZNvI9Pz" outputId="db4427a9-eef2-4cac-c839-13edc895d70c"
plt.plot(x, np.sin(x), 'g', label='sin(x)')
plt.plot(x, np.cos(x), ':r', label='cos(x)')
plt.legend();
# -
| tirgul/tirgul5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda]
# language: python
# name: conda-env-anaconda-py
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ***
# ***
# # 基于机器学习的情感分析
# ***
# ***
#
# 王成军
#
# <EMAIL>
#
# 计算传播网 http://computational-communication.com
# + [markdown] slideshow={"slide_type": "slide"}
# 情感分析(sentiment analysis)和意见挖掘(opinion mining)虽然相关,但是从社会科学的角度而言,二者截然不同。这里主要是讲情感分析(sentiment or emotion),而非意见挖掘(opinion, 后者通过机器学习效果更可信)。
#
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# # classify emotion
# Different types of emotion: anger, disgust, fear, joy, sadness, and surprise. The classification can be performed using different algorithms: e.g., naive Bayes classifier trained on Carlo Strapparava and <NAME>’s emotions lexicon.
#
# # classify polarity
#
# To classify some text as positive or negative. In this case, the classification can be done by using a naive Bayes algorithm trained on Janyce Wiebe’s subjectivity lexicon.
# + [markdown] slideshow={"slide_type": "slide"}
# 
#
# + [markdown] slideshow={"slide_type": "subslide"}
#
# # LIWC & TextMind
#
# http://ccpl.psych.ac.cn/textmind/
#
# “文心(TextMind)”中文心理分析系统是由中科院心理所计算网络心理实验室研发的,针对中文文本进行语言分析的软件系统,通过“文心”,您可以便捷地分析文本中使用的不同类别语言的程度、偏好等特点。针对中国大陆地区简体环境下的语言特点,参照LIWC2007和正體中文C-LIWC词库,我们开发了“文心(TextMind)”中文心理分析系统。“文心”为用户提供从简体中文自动分词,到语言心理分析的一揽子分析解决方案,其词库、文字和符号等处理方法专门针对简体中文语境,词库分类体系也与LIWC兼容一致。
# + [markdown] slideshow={"slide_type": "slide"}
# # Preparing the data
# + [markdown] slideshow={"slide_type": "subslide"}
# # NLTK
# Anaconda自带的(默认安装的)第三方包。http://www.nltk.org/
#
# > NLTK is a leading platform for building Python programs to work with human language data. It provides easy-to-use interfaces to over 50 corpora and lexical resources such as WordNet, along with a suite of text processing libraries for classification, tokenization, stemming, tagging, parsing, and semantic reasoning, wrappers for industrial-strength NLP libraries, and an active discussion forum.
# + slideshow={"slide_type": "subslide"}
import nltk
pos_tweets = [('I love this car', 'positive'),
('This view is amazing', 'positive'),
('I feel great this morning', 'positive'),
('I am so excited about the concert', 'positive'),
('He is my best friend', 'positive')]
neg_tweets = [('I do not like this car', 'negative'),
('This view is horrible', 'negative'),
('I feel tired this morning', 'negative'),
('I am not looking forward to the concert', 'negative'),
('He is my enemy', 'negative')]
# + slideshow={"slide_type": "subslide"}
tweets = []
for (words, sentiment) in pos_tweets + neg_tweets:
words_filtered = [e.lower() for e in words.split() if len(e) >= 3]
tweets.append((words_filtered, sentiment))
tweets[:2]
# + slideshow={"slide_type": "subslide"}
test_tweets = [
(['feel', 'happy', 'this', 'morning'], 'positive'),
(['larry', 'friend'], 'positive'),
(['not', 'like', 'that', 'man'], 'negative'),
(['house', 'not', 'great'], 'negative'),
(['your', 'song', 'annoying'], 'negative')]
# + [markdown] slideshow={"slide_type": "slide"}
# # Extracting Features
# Then we need to get the unique word list as the features for classification.
#
# + code_folding=[] slideshow={"slide_type": "subslide"}
# get the word lists of tweets
def get_words_in_tweets(tweets):
all_words = []
for (words, sentiment) in tweets:
all_words.extend(words)
return all_words
# get the unique word from the word list
def get_word_features(wordlist):
wordlist = nltk.FreqDist(wordlist)
word_features = wordlist.keys()
return word_features
word_features = get_word_features(get_words_in_tweets(tweets))
' '.join(word_features)
# + [markdown] slideshow={"slide_type": "subslide"}
# To create a classifier, we need to decide what features are relevant. To do that, we first need a feature extractor.
# + slideshow={"slide_type": "subslide"}
def extract_features(document):
document_words = set(document)
features = {}
for word in word_features:
features['contains(%s)' % word] = (word in document_words)
return features
# + slideshow={"slide_type": "skip"}
help(nltk.classify.util.apply_features)
# + slideshow={"slide_type": "skip"}
training_set[0]
# + slideshow={"slide_type": "slide"}
training_set = nltk.classify.util.apply_features(extract_features,\
tweets)
classifier = nltk.NaiveBayesClassifier.train(training_set)
# + slideshow={"slide_type": "subslide"}
# You may want to know how to define the ‘train’ method in NLTK here:
def train(labeled_featuresets, estimator=nltk.probability.ELEProbDist):
# Create the P(label) distribution
label_probdist = estimator(label_freqdist)
# Create the P(fval|label, fname) distribution
feature_probdist = {}
model = NaiveBayesClassifier(label_probdist, feature_probdist)
return model
# + slideshow={"slide_type": "slide"}
tweet_positive = '<NAME>'
classifier.classify(extract_features(tweet_positive.split()))
# + slideshow={"slide_type": "fragment"}
tweet_negative = '<NAME> not my friend'
classifier.classify(extract_features(tweet_negative.split()))
# + slideshow={"slide_type": "subslide"}
# Don’t be too positive, let’s try another example:
tweet_negative2 = 'Your song is annoying'
classifier.classify(extract_features(tweet_negative2.split()))
# + slideshow={"slide_type": "slide"}
def classify_tweet(tweet):
return classifier.classify(extract_features(tweet))
# nltk.word_tokenize(tweet)
total = accuracy = float(len(test_tweets))
for tweet in test_tweets:
if classify_tweet(tweet[0]) != tweet[1]:
accuracy -= 1
print('Total accuracy: %f%% (%d/20).' % (accuracy / total * 100, accuracy))
# + [markdown] slideshow={"slide_type": "slide"}
# # 使用sklearn的分类器
# + slideshow={"slide_type": "subslide"}
# nltk有哪些分类器呢?
nltk_classifiers = dir(nltk)
for i in nltk_classifiers:
if 'Classifier' in i:
print(i)
# + slideshow={"slide_type": "subslide"}
from sklearn.svm import LinearSVC
from nltk.classify.scikitlearn import SklearnClassifier
classif = SklearnClassifier(LinearSVC())
svm_classifier = classif.train(training_set)
# + slideshow={"slide_type": "subslide"}
# Don’t be too positive, let’s try another example:
tweet_negative2 = 'Your song is annoying'
svm_classifier.classify(extract_features(tweet_negative2.split()))
# + [markdown] slideshow={"slide_type": "slide"}
# # 作业1:
#
# 使用另外一种sklearn的分类器来对tweet_negative2进行情感分析
#
# # 作业2:
#
# 使用https://github.com/victorneo/Twitter-Sentimental-Analysis 所提供的推特数据进行情感分析,可以使用其代码 https://github.com/victorneo/Twitter-Sentimental-Analysis/blob/master/classification.py
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# # 推荐阅读:
#
# movies reviews情感分析 http://nbviewer.jupyter.org/github/rasbt/python-machine-learning-book/blob/master/code/ch08/ch08.ipynb
#
# Sentiment analysis with machine learning in R http://chengjun.github.io/en/2014/04/sentiment-analysis-with-machine-learning-in-R/
#
# 使用R包sentiment进行情感分析 https://site.douban.com/146782/widget/notes/15462869/note/344846192/
#
# 中文的手机评论的情感分析 https://github.com/computational-class/Review-Helpfulness-Prediction
#
# 基于词典的中文情感倾向分析 https://site.douban.com/146782/widget/notes/15462869/note/355625387/
# + [markdown] slideshow={"slide_type": "slide"}
# # Sentiment Analysis using TextBlob
# -
# # 安装textblob
# https://github.com/sloria/TextBlob
#
# > pip install -U textblob
#
# > python -m textblob.download_corpora
# + slideshow={"slide_type": "subslide"}
from textblob import TextBlob
text = '''
The titular threat of The Blob has always struck me as the ultimate movie
monster: an insatiably hungry, amoeba-like mass able to penetrate
virtually any safeguard, capable of--as a doomed doctor chillingly
describes it--"assimilating flesh on contact.
Snide comparisons to gelatin be damned, it's a concept with the most
devastating of potential consequences, not unlike the grey goo scenario
proposed by technological theorists fearful of
artificial intelligence run rampant.
'''
blob = TextBlob(text)
blob.tags # [('The', 'DT'), ('titular', 'JJ'),
# ('threat', 'NN'), ('of', 'IN'), ...]
blob.noun_phrases # WordList(['titular threat', 'blob',
# 'ultimate movie monster',
# 'amoeba-like mass', ...])
for sentence in blob.sentences:
print(sentence.sentiment.polarity)
# 0.060
# -0.341
blob.translate(to="es") # 'La amenaza titular de The Blob...'
# + [markdown] slideshow={"slide_type": "slide"}
# # Sentiment Analysis Using Turicreate
# + [markdown] slideshow={"slide_type": "fragment"}
# In this notebook, I will explain how to develop sentiment analysis classifiers that are based on a bag-of-words model.
# Then, I will demonstrate how these classifiers can be utilized to solve Kaggle's "When Bag of Words Meets Bags of Popcorn" challenge.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Code Recipe: Creating Sentiment Classifier
# + [markdown] slideshow={"slide_type": "fragment"}
# Using <del>GraphLab</del> Turicreate it is very easy and straight foward to create a sentiment classifier based on bag-of-words model. Given a dataset stored as a CSV file, you can construct your sentiment classifier using the following code:
# + slideshow={"slide_type": "fragment"}
import turicreate as tc
train_data = tc.SFrame.read_csv(traindata_path,header=True,
delimiter='\t',quote_char='"',
column_type_hints = {'id':str,
'sentiment' : int,
'review':str } )
train_data['1grams features'] = tc.text_analytics.count_ngrams(
train_data['review'],1)
train_data['2grams features'] = tc.text_analytics.count_ngrams(
train_data['review'],2)
cls = tc.classifier.create(train_data, target='sentiment',
features=['1grams features',
'2grams features'])
# + [markdown] slideshow={"slide_type": "fragment"}
# In the rest of this notebook, we will explain this code recipe in details, by demonstrating how this recipe can used to create IMDB movie reviews sentiment classifier.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Set up
# + [markdown] slideshow={"slide_type": "fragment"}
# Before we begin constructing the classifiers, we need to import some Python libraries: turicreate (tc), and IPython display utilities.
# + slideshow={"slide_type": "fragment"}
import turicreate as tc
from IPython.display import display
from IPython.display import Image
# + [markdown] slideshow={"slide_type": "slide"}
# ## IMDB movies reviews Dataset
#
# > # Bag of Words Meets Bags of Popcorn
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Throughout this notebook, I will use Kaggle's IMDB movies reviews datasets that is available to download from the following link: https://www.kaggle.com/c/word2vec-nlp-tutorial/data. I downloaded labeledTrainData.tsv and testData.tsv files, and unzipped them to the following local files.
#
# ### DeepLearningMovies
#
# Kaggle's competition for using Google's word2vec package for sentiment analysis
#
# https://github.com/wendykan/DeepLearningMovies
# + slideshow={"slide_type": "slide"}
traindata_path = "/Users/datalab/bigdata/cjc/kaggle_popcorn_data/labeledTrainData.tsv"
testdata_path = "/Users/datalab/bigdata/cjc/kaggle_popcorn_data/testData.tsv"
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Loading Data
# + [markdown] slideshow={"slide_type": "fragment"}
# We will load the data with IMDB movie reviews to an SFrame using SFrame.read_csv function.
# + slideshow={"slide_type": "slide"}
movies_reviews_data = tc.SFrame.read_csv(traindata_path,header=True,
delimiter='\t',quote_char='"',
column_type_hints = {'id':str,
'sentiment' : str,
'review':str } )
# + [markdown] slideshow={"slide_type": "fragment"}
# By using the SFrame show function, we can visualize the data and notice that the train dataset consists of 12,500 positive and 12,500 negative, and overall 24,932 unique reviews.
# + slideshow={"slide_type": "slide"}
movies_reviews_data
# + [markdown] slideshow={"slide_type": "slide"}
# ## Constructing Bag-of-Words Classifier
# + [markdown] slideshow={"slide_type": "fragment"}
# One of the common techniques to perform document classification (and reviews classification) is using Bag-of-Words model, in which the frequency of each word in the document is used as a feature for training a classifier. GraphLab's text analytics toolkit makes it easy to calculate the frequency of each word in each review. Namely, by using the count_ngrams function with n=1, we can calculate the frequency of each word in each review. By running the following command:
# + slideshow={"slide_type": "fragment"}
movies_reviews_data['1grams features'] = tc.text_analytics.count_ngrams(movies_reviews_data ['review'],1)
# + [markdown] slideshow={"slide_type": "fragment"}
# By running the last command, we created a new column in movies_reviews_data SFrame object. In this column each value is a dictionary object, where each dictionary's keys are the different words which appear in the corresponding review, and the dictionary's values are the frequency of each word.
# We can view the values of this new column using the following command.
# + slideshow={"slide_type": "slide"}
movies_reviews_data#[['review','1grams features']]
# + [markdown] slideshow={"slide_type": "slide"}
# We are now ready to construct and evaluate the movie reviews sentiment classifier using the calculated above features. But first, to be able to perform a quick evaluation of the constructed classifier, we need to create labeled train and test datasets. We will create train and test datasets by randomly splitting the train dataset into two parts. The first part will contain 80% of the labeled train dataset and will be used as the training dataset, while the second part will contain 20% of the labeled train dataset and will be used as the testing dataset. We will create these two dataset by using the following command:
# + slideshow={"slide_type": "fragment"}
train_set, test_set = movies_reviews_data.random_split(0.8, seed=5)
# + [markdown] slideshow={"slide_type": "slide"}
# We are now ready to create a classifier using the following command:
# + slideshow={"slide_type": "fragment"}
model_1 = tc.classifier.create(train_set, target='sentiment', \
features=['1grams features'])
# + [markdown] slideshow={"slide_type": "slide"}
# We can evaluate the performence of the classifier by evaluating it on the test dataset
# + slideshow={"slide_type": "fragment"}
result1 = model_1.evaluate(test_set)
# + [markdown] slideshow={"slide_type": "fragment"}
# In order to get an easy view of the classifier's prediction result, we define and use the following function
# + slideshow={"slide_type": "slide"}
def print_statistics(result):
print( "*" * 30)
print( "Accuracy : ", result["accuracy"])
print( "Confusion Matrix: \n", result["confusion_matrix"])
print_statistics(result1)
# + [markdown] slideshow={"slide_type": "fragment"}
# As can be seen in the results above, in just a few relatively straight foward lines of code, we have developed a sentiment classifier that has accuracy of about ~0.88. Next, we demonstrate how we can improve the classifier accuracy even more.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Improving The Classifier
# + [markdown] slideshow={"slide_type": "fragment"}
# One way to improve the movie reviews sentiment classifier is to extract more meaningful features from the reviews. One method to add additional features, which might be meaningful, is to calculate the frequency of every two consecutive words in each review. To calculate the frequency of each two consecutive words in each review, as before, we will use turicreate's count_ngrams function only this time we will set n to be equal 2 (n=2) to create new column named '2grams features'.
# + slideshow={"slide_type": "fragment"}
movies_reviews_data['2grams features'] = tc.text_analytics.count_ngrams(movies_reviews_data['review'],2)
# -
movies_reviews_data
# + [markdown] slideshow={"slide_type": "fragment"}
# As before, we will construct and evaluate a movie reviews sentiment classifier. However, this time we will use both the '1grams features' and the '2grams features' features
# + slideshow={"slide_type": "slide"}
train_set, test_set = movies_reviews_data.random_split(0.8, seed=5)
model_2 = tc.classifier.create(train_set, target='sentiment', features=['1grams features','2grams features'])
result2 = model_2.evaluate(test_set)
# + slideshow={"slide_type": "subslide"}
print_statistics(result2)
# + [markdown] slideshow={"slide_type": "slide"}
# Indeed, the new constructed classifier seems to be more accurate with an accuracy of about ~0.9.
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Unlabeled Test File
# + [markdown] slideshow={"slide_type": "fragment"}
# To test how well the presented method works, we will use all the 25,000 labeled IMDB movie reviews in the train dataset to construct a classifier. Afterwards, we will utilize the constructed classifier to predict sentiment for each review in the unlabeled dataset. Lastly, we will create a submission file according to Kaggle's guidelines and submit it.
# + slideshow={"slide_type": "slide"}
traindata_path = "/Users/datalab/bigdata/cjc/kaggle_popcorn_data/labeledTrainData.tsv"
testdata_path = "/Users/datalab/bigdata/cjc/kaggle_popcorn_data/testData.tsv"
#creating classifier using all 25,000 reviews
train_data = tc.SFrame.read_csv(traindata_path,header=True, delimiter='\t',quote_char='"',
column_type_hints = {'id':str, 'sentiment' : int, 'review':str } )
train_data['1grams features'] = tc.text_analytics.count_ngrams(train_data['review'],1)
train_data['2grams features'] = tc.text_analytics.count_ngrams(train_data['review'],2)
cls = tc.classifier.create(train_data, target='sentiment', features=['1grams features','2grams features'])
#creating the test dataset
test_data = tc.SFrame.read_csv(testdata_path,header=True, delimiter='\t',quote_char='"',
column_type_hints = {'id':str, 'review':str } )
test_data['1grams features'] = tc.text_analytics.count_ngrams(test_data['review'],1)
test_data['2grams features'] = tc.text_analytics.count_ngrams(test_data['review'],2)
#predicting the sentiment of each review in the test dataset
test_data['sentiment'] = cls.classify(test_data)['class'].astype(int)
#saving the prediction to a CSV for submission
test_data[['id','sentiment']].save("/Users/datalab/bigdata/cjc/kaggle_popcorn_data/predictions.csv", format="csv")
# + [markdown] slideshow={"slide_type": "slide"}
# We then submitted the predictions.csv file to the Kaggle challange website and scored AUC of about 0.88.
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Further Readings
# + [markdown] slideshow={"slide_type": "fragment"}
# Further reading materials can be found in the following links:
#
# http://en.wikipedia.org/wiki/Bag-of-words_model
#
# https://dato.com/products/create/docs/generated/graphlab.SFrame.html
#
# https://dato.com/products/create/docs/graphlab.toolkits.classifier.html
#
# https://www.kaggle.com/c/word2vec-nlp-tutorial/details/part-1-for-beginners-bag-of-words
#
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. (2011). "Learning Word Vectors for Sentiment Analysis." The 49th Annual Meeting of the Association for Computational Linguistics (ACL 2011).
#
| code/11.sentiment_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 对象关系映射
# 数据库中的记录可以与一个 `Python` 对象对应。
#
# 例如对于上一节中的数据库:
#
# Order|Date|Stock|Quantity|Price
# --|--|--|--|--
# A0001|2013-12-01|AAPL|1000|203.4
# A0002|2013-12-01|MSFT|1500|167.5
# A0003|2013-12-02|GOOG|1500|167.5
#
# 可以用一个类来描述:
#
# Attr.|Method
# --|--
# Order id| Cost
# Date|
# Stock|
# Quant.|
# Price|
#
# 可以使用 `sqlalchemy` 来实现这种对应:
# +
from sqlalchemy.ext.declarative import declarative_base
from sqlalchemy import Column, Date, Float, Integer, String
Base = declarative_base()
class Order(Base):
__tablename__ = 'orders'
order_id = Column(String, primary_key=True)
date = Column(Date)
symbol = Column(String)
quantity = Column(Integer)
price = Column(Float)
def get_cost(self):
return self.quantity*self.price
# -
# 生成一个 `Order` 对象:
import datetime
order = Order(order_id='A0004', date=datetime.date.today(), symbol='MSFT', quantity=-1000, price=187.54)
# 调用方法:
order.get_cost()
# 使用上一节生成的数据库产生一个 `session`:
# +
from sqlalchemy import create_engine
from sqlalchemy.orm import sessionmaker
engine = create_engine("sqlite:///my_database.sqlite") # 相当于 connection
Session = sessionmaker(bind=engine) # 相当于 cursor
session = Session()
# -
# 使用这个 `session` 向数据库中添加刚才生成的对象:
session.add(order)
session.commit()
# 显示是否添加成功:
for row in engine.execute("SELECT * FROM orders"):
print row
# 使用 `filter` 进行查询,返回的是 `Order` 对象的列表:
for order in session.query(Order).filter(Order.symbol=="AAPL"):
print order.order_id, order.date, order.get_cost()
# 返回列表的第一个:
order_2 = session.query(Order).filter(Order.order_id=='A0002').first()
order_2.symbol
| lijin-THU:notes-python/05-advanced-python/05.07-object-relational-mappers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 13.3. Simulating a Brownian motion
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
n = 5000
k = 10
x = np.cumsum(np.random.randn(n))
y = np.cumsum(np.random.randn(n))
# +
# We add 10 intermediary points between two
# successive points. We interpolate x and y.
x2 = np.interp(np.arange(n * k), np.arange(n) * k, x)
y2 = np.interp(np.arange(n * k), np.arange(n) * k, y)
# + podoc={"output_text": "<matplotlib.figure.Figure at 0x7734b38>"}
fig, ax = plt.subplots(1, 1, figsize=(8, 8))
# Now, we draw our points with a gradient of colors.
ax.scatter(x2, y2, c=range(n * k), linewidths=0,
marker='o', s=3, cmap=plt.cm.jet,)
ax.axis('equal')
ax.set_axis_off()
# -
| chapter13_stochastic/03_brownian.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
interim_path = "../data/interim/"
# -
df = pd.read_csv(interim_path+"sp500sub.csv")
df.head()
df.info()
print(f"""Qty of stocks: {df["Name"].nunique()}""")
# +
date_min = df["Date"].min()
date_max = df["Date"].max()
print(f"""Range of the data: from {date_min} to {date_max} with {df["Date"].nunique()} unique dates.""")
# +
df_close_prices = pd.pivot_table(df, values='Close', columns=['Name'], index=['Date'])
print(f"""After getting only close prices and pivoting table, we get {df_close_prices.shape[0]} observations (or dates), and {df_close_prices.shape[1]} columns (or stocks).
""")
df_close_prices.head()
# +
print("CAUTION! The following stocks have NaN values.")
lst_null_stocks = (df_close_prices.isnull().sum()[df_close_prices.isnull().sum()>0].sort_values(ascending=False)).index
lst_null_stocks = lst_null_stocks.tolist()
df_close_prices.isnull().sum()[df_close_prices.isnull().sum()>0].sort_values(ascending=False)
# -
df_close_prices.to_parquet(interim_path+"sp500sub_close.pqt")
# +
df_close_prices_2 = pd.read_parquet(interim_path+"sp500sub_close.pqt")
print(f"""Loading data and parsing by date, we get {df_close_prices_2.shape[0]} observations (or dates), and {df_close_prices_2.shape[1]} columns (or stocks).""")
df_close_prices_2.head()
# -
# # Dealing with missing data
# +
fig = plt.figure(figsize=(20, 5))
ax = fig.add_subplot(111)
for stock in lst_null_stocks:
ax.plot(df_close_prices_2[stock], label=stock)
ax.xaxis.set_major_locator(plt.MaxNLocator(30))
ax.legend(loc='lower left', frameon=False)
plt.xticks(rotation=90)
plt.show();
# -
# ## Removing holidays and weekends
df_close_prices_cleaning = df_close_prices_2.dropna(axis=0, how='all').copy()
# +
fig = plt.figure(figsize=(20, 5))
ax = fig.add_subplot(111)
for stock in lst_null_stocks:
ax.plot(df_close_prices_cleaning[stock], label=stock)
ax.xaxis.set_major_locator(plt.MaxNLocator(30))
ax.legend(loc='lower left', frameon=False)
plt.xticks(rotation=90)
plt.show();
# +
print("CAUTION! The following stocks have NaN values.")
df_close_prices_cleaning.isnull().sum()[df_close_prices_cleaning.isnull().sum()>0].sort_values(ascending=False)
# -
# ## Foward fill and backward fill
print(f"""Total NaNs before foward fill: {df_close_prices_cleaning.isna().sum().sum()}""")
# +
df_close_prices_cleaning.fillna(method='ffill', inplace=True)
print(f"""Total NaNs AFTER foward fill: {df_close_prices_cleaning.isna().sum().sum()}""")
# +
df_close_prices_cleaning.fillna(method='bfill', inplace=True)
print(f"""Total NaNs AFTER backward fill: {df_close_prices_cleaning.isna().sum().sum()}""")
# +
fig = plt.figure(figsize=(20, 5))
ax = fig.add_subplot(111)
for stock in lst_null_stocks:
ax.plot(df_close_prices_cleaning[stock], label=stock)
ax.xaxis.set_major_locator(plt.MaxNLocator(30))
ax.legend(loc='lower left', frameon=False)
plt.xticks(rotation=90)
plt.show();
# -
# # Normalised stock values
df_close_prices_normalised = df_close_prices_cleaning/df_close_prices_cleaning.iloc[0]
# +
fig = plt.figure(figsize=(20, 5))
ax = fig.add_subplot(111)
for stock in lst_null_stocks:
ax.plot(df_close_prices_normalised[stock], label=stock)
ax.xaxis.set_major_locator(plt.MaxNLocator(30))
ax.legend(loc='upper left', frameon=False)
plt.xticks(rotation=90)
plt.show();
# -
| section_02_financial_basics/s02_02_understanding_financial_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="80H-_h1_Yzy-" colab_type="code" colab={}
#importing necessary libraries
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import tensorflow_hub as hub
import tensorflow_datasets as tfds
from tensorflow.keras import layers
import logging
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
# + id="uPhVg8obZNF0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 593, "referenced_widgets": ["3c0750d5021d4a31b21706e5f53815dd", "ef6ea4c8c5104ba9a05f899291dc8b7a", "f4e265f910ad45e1a600ec1cc202fdd5", "f08a82607b0e4151b5d6ab585fc4d6be", "0af3ffd79eca46248fd6da037ef29657", "b8f7e3caac82435f8f440628836eb999", "7b42bda76c244dc2b8f9ee21e3e462ba", "c61b95441af9412d9724ad63cf879bee"]} outputId="3618d3cc-5f38-470d-fc17-db5e3f3c15db"
#Downloading and splitting the dataset
(training_set, validation_set), dataset_info = tfds.load(
'tf_flowers',
split=['train[:70%]','train[70%:]'],
with_info=True,
as_supervised=True)
print(dataset_info)
# + id="npN9sIIoZXMB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="687cb8cf-bac4-4655-af74-bfe9396725ff"
num_classes = dataset_info.features['label'].num_classes
im_shape = dataset_info.features['image'].shape
print(im_shape)
num_training_examples = 0
for example in training_set:
num_training_examples += 1
num_validation_examples = 0
for example in validation_set:
num_validation_examples += 1
print('Total Number of Classes: {}'.format(num_classes))
print('Total Number of Training Images: {}'.format(num_training_examples))
print('Total Number of Validation Images: {} \n'.format(num_validation_examples))
# + id="LwrwFIJFZc11" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="a3da0556-9f3a-4a8a-aff2-ae774953accd"
for i, example in enumerate(training_set.take(5)):
print('Image {} shape: {} label: {}'.format(i+1, example[0].shape, example[1]))
# + id="gOH2KZ-wZgEj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="40a883c0-e49c-4fbb-f51c-63d05d24e8cf"
#Reformatting images and creating batches
IMAGE_RES = 224
def format_image(image, label):
#image = tf.image.resize(image, (IMAGE_RES, IMAGE_RES))/.255 this gives lower accuracy in the model I tried beleive me!!~~
image = tf.image.resize(image, (IMAGE_RES, IMAGE_RES))/255.0
return image, label
BATCH_SIZE = 32
train_batches = training_set.shuffle(num_training_examples//4).map(format_image).batch(BATCH_SIZE).prefetch(1)
validation_batches = validation_set.map(format_image).batch(BATCH_SIZE).prefetch(1)
print(train_batches)
print(validation_batches)
# + id="ksW5osKIZpFL" colab_type="code" colab={}
#Simple transfer learning from hub
URL = 'https://tfhub.dev/google/tf2-preview/mobilenet_v2/feature_vector/4'
feature_extractor = hub.KerasLayer(URL, input_shape = (IMAGE_RES, IMAGE_RES,3),trainable=False)# Also Freezed the pre-trained model
# + id="LjcgxdkcaA56" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="a0053640-d998-4642-8bb6-bd1dc3d10dc5"
#Attaching a classification head
model = tf.keras.Sequential([
feature_extractor,
layers.Dense(num_classes)
#can also be done this way
#tf.keras.layers.Dense(num_classes)
])
model.summary()
# + id="fwc2H1muaODq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="599473ab-01e5-409e-abcc-7adc38dd1f82"
#Training the model
model.compile(
optimizer = 'adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits = True),
metrics=['accuracy'])
EPOCHS = 6
history = model.fit(
train_batches,
epochs=EPOCHS,
validation_data = validation_batches)
# + id="0cybZAEoabmQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 499} outputId="4bcf1d80-2204-4fb2-94fe-c06bc401610d"
#Plotting validation and training graph
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(EPOCHS)
plt.figure(figsize = (8,8))
plt.subplot(1,2,1)
plt.plot(epochs_range, acc, label = 'Training accuracy')
plt.plot(epochs_range, val_acc, label = 'validation accuracy')
plt.legend(loc = 'lower right')
plt.title('Training and Validation accuracy')
plt.subplot(1,2,2)
plt.plot(epochs_range, loss, label = 'Training loss')
plt.plot(epochs_range, val_loss, label = 'validation loss')
plt.legend(loc = 'upper right')
plt.title('Training and Validation loss')
plt.show()
# + id="SI-2Xumza8l9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="48f2abbe-55f3-4580-c6ba-1fb985d1c5fb"
#checking prediction
class_names = np.array(dataset_info.features['label'].names)
print(class_names)
# + id="QOWjixnCbDXp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="1d5d2a81-8df9-4a60-b26b-f96dd882b537"
#creating an image batch amd checking prediction
image_batch, label_batch = next(iter(train_batches))
image_batch = image_batch.numpy()
label_batch = label_batch.numpy()
predicted_batch = model.predict(image_batch)
predicted_batch = tf.squeeze(predicted_batch).numpy()
predicted_ids = np.argmax(predicted_batch, axis=-1)
predicted_class_names = class_names[predicted_ids]
print(predicted_class_names)
# + id="5wkOZT-1bNmv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="8be68412-3eff-4ca7-84b1-7e0a085c32b1"
#printing True labels and prediction indices
print("Labels: ", label_batch)
print("Predicted labels: ", predicted_ids)
# + id="2OLvF30kbz90" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 585} outputId="7830af20-9732-4009-dccd-52bce841ddfc"
#Plotting model predictions
plt.figure(figsize=(10,9))
for n in range(30):
plt.subplot(6,5,n+1)
plt.subplots_adjust(hspace = 0.3)
plt.imshow(image_batch[n])
color = "blue" if predicted_ids[n] == label_batch[n] else "red"
plt.title(predicted_class_names[n].title(), color=color)
plt.axis('off')
_ = plt.suptitle("Model predictions (blue: correct, red: incorrect)")
# + id="_v8G7xIGb73e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="5834b645-9267-4b57-cf08-fafb9a09a278"
#performing Transfer learning with inception model
IMAGE_RESOLUTION = 299
(training_set,validation_set), dataset_in = tfds.load(
'tf_flowers',
split = ['train[:70%]','train[70%:]'],
with_info=True,
as_supervised=True,
)
def for_i(image,label):
image = tf.image.resize(image, (IMAGE_RESOLUTION, IMAGE_RESOLUTION))/255.0
return image,label
train_batches=training_set.shuffle(num_training_examples//4).map(for_i).batch(BATCH_SIZE).prefetch(1)
validation_batches=validation_set.map(for_i).batch(BATCH_SIZE).prefetch(1)
URL = 'https://tfhub.dev/google/tf2-preview/inception_v3/feature_vector/4'
feature_ex = hub.KerasLayer(URL, input_shape=(IMAGE_RESOLUTION,IMAGE_RESOLUTION,3),trainable=False)
#feature_ex.trainable = False
model_ec = tf.keras.Sequential(
[feature_ex,
#layers.Dense(num_classes)
tf.keras.layers.Dense(num_classes)]
)
model_ec.summary()
# + id="iI3bevpScVDn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="d08ad25c-c607-4bbf-8d1a-68c3b228e43f"
#training the inception model
model_ec.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics = ['accuracy']
)
EP = 6
history_inception = model_ec.fit(
train_batches,
epochs=EP,
validation_data = validation_batches)
# + id="Hg0y5m10cZ2P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 499} outputId="07372ff3-3594-4001-e253-fae7072eed7e"
#plotting the inception model sets of accuracy and loss
acc = history_inception.history['accuracy']
val_acc = history_inception.history['val_accuracy']
loss = history_inception.history['loss']
val_loss = history_inception.history['val_loss']
epochs_range = range(EP)
plt.figure(figsize = (8,8))
plt.subplot(1,2,1)
plt.plot(epochs_range, acc, label = 'Training accuracy')
plt.plot(epochs_range, val_acc, label = 'validation accuracy')
plt.legend(loc = 'lower right')
plt.title('Training and Validation accuracy')
plt.subplot(1,2,2)
plt.plot(epochs_range, loss, label = 'Training loss')
plt.plot(epochs_range, val_loss, label = 'validation loss')
plt.legend(loc = 'upper right')
plt.title('Training and Validation loss')
plt.show()
# + id="Xk690QL-dxAn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1f441499-851a-4f03-944f-a93ef177d558"
#checking prediction with inception model
class_name=np.array(dataset_in.features['label'].names)
print(class_name)
# + id="tRBwdQjiefns" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="4ec139e2-5f29-4149-db0a-a521ce6dcd68"
#creating image batches and checking them with inception model
image_batch, label_batch = next(iter(train_batches))
image_batch = image_batch.numpy()
label_batch = label_batch.numpy()
predicted_batch = model_ec.predict(image_batch)
predicted_batch = tf.squeeze(predicted_batch).numpy()
predicted_ids = np.argmax(predicted_batch, axis=-1)
predicted_class_name = class_name[predicted_ids]
print(predicted_class_name)
# + id="NDC1x2L0e0bY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="9e245f85-17cd-47d9-c4d0-f7f97c5fbce8"
#printing true labels and predicted indices
print("Labels: ", label_batch)
print("Predicted labels: ", predicted_ids)
# + id="sU-N29Vre9-v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 585} outputId="76c21e66-a320-427d-c9e8-f81efe79951e"
#plotting Inception model prediction for better comparison between two models
plt.figure(figsize=(10,9))
for n in range(30):
plt.subplot(6,5,n+1)
plt.subplots_adjust(hspace = 0.3)
plt.imshow(image_batch[n])
color = "green" if predicted_ids[n] == label_batch[n] else "red"
plt.title(predicted_class_name[n].title(), color=color)
plt.axis('off')
_ = plt.suptitle("Model predictions (green: correct, red: incorrect)")
| ML_Training_udacity/5. TRANSFER LEARNING/Transfer_learning_for_flowers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="iz2Tuf8C2hoc"
# # 05 - Python Finance
#
# **Capitulo 05**: Calcular o desempenho da carteira de ações. Índice Sharpe, Beta, CAGR, Volatilidade Anual e Drawdown são alguns dos indicadores de desempenho que serão calculados.
# + [markdown] id="GoQ7GXYMt1kL"
# ## 1. Importando bibliotecas
#
# Instalando o YFinance
#
# + id="betAK92nshET"
# !pip install yfinance --upgrade --no-cache-dir
# + [markdown] id="7foPovCAuVBA"
# Importando o YFinance e sobrescrevendo os métodos do pandas_datareader
# + id="-mxNCu2vuXCi"
import yfinance as yf
import pandas_datareader.data as web
yf.pdr_override()
# + [markdown] id="Tg17VivDuQ3q"
# Importando as Bibliotecas
# + id="JHEYijH-uRiR"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="zI4n2rsC0W1o"
# Instalando o PyFolio
# + id="DePP77QF0MVV"
# !pip install pyfolio
# + id="-_6nVffU6hVm"
# Para corrgir o bug: AttributeError: 'numpy.int64' object has no attribute 'to_pydatetime'
# !pip install git+https://github.com/quantopian/pyfolio
# + [markdown] id="7jcWdhPA0c3M"
# Importando as Bibliotecas
# + id="oxECJvGm6ozz"
import pyfolio as pf
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="ABiLJwk9uZYM"
# ## 2. Obtendo e tratando os dados
# + colab={"base_uri": "https://localhost:8080/"} id="WsyD8CAc1Fi8" outputId="08fff807-3209-4abe-b984-3cd4617b604d"
tickers = "PETR3.SA PRIO3.SA BBDC4.SA ITUB4.SA BBAS3.SA BIDI4.SA ITSA4.SA VALE3.SA MGLU3.SA MDIA3.SA ALPA4.SA WEGE3.SA TAEE11.SA TRPL4.SA ALUP11.SA EGIE3.SA EQTL3.SA ENGI11.SA SAPR11.SA MRVE3.SA EZTC3.SA TRIS3.SA ABEV3.SA QUAL3.SA FLRY3.SA RADL3.SA HYPE3.SA GNDI3.SA VIVT3.SA RENT3.SA RAIL3.SA JBSS3.SA LWSA3.SA TOTS3.SA SUZB3.SA CSAN3.SA UNIP6.SA NTCO3.SA ^BVSP"
dados_yahoo = yf.download(tickers=tickers, period="5y")['Adj Close']
# + id="KXLutTei6zsj"
#tickers = ["tickers = "PETR3.SA", "PRIO3.SA", "BBDC4.SA", "ITUB4.SA", "BBAS3.SA", "BIDI4.SA", "ITSA4.SA", "VALE3.SA", "MGLU3.SA", "MDIA3.SA", "ALPA4.SA", "WEGE3.SA", "TAEE11.SA", "TRPL4.SA", "ALUP11.SA", "EGIE3.SA", "EQTL3.SA", "ENGI11.SA", "SAPR11.SA", "MRVE3.SA", "EZTC3.SA", "TRIS3.SA", "ABEV3.SA", "QUAL3.SA", "FLRY3.SA", "RADL3.SA", "HYPE3.SA", "GNDI3.SA", "VIVT3.SA", "RENT3.SA", "RAIL3.SA", "JBSS3.SA", "LWSA3.SA", "TOTS3.SA", "SUZB3.SA", "CSAN3.SA", "UNIP6.SA", "NTCO3.SA", '^BVSP']
#tickers1 = "PETR3.SA PRIO3.SA"
#tickers2 = "BBDC4.SA ITUB4.SA BBAS3.SA BIDI4.SA ITSA4.SA"
#tickers3 = "VALE3.SA"
#tickers4 = "MGLU3.SA MDIA3.SA ALPA4.SA"
#tickers5 = "WEGE3.SA"
#tickers6 = "TAEE11.SA TRPL4.SA ALUP11.SA EGIE3.SA EQTL3.SA ENGI11.SA"
#tickers7 = "SAPR11.SA"
#tickers8 = "MRVE3.SA EZTC3.SA TRIS3.SA"
#tickers9 = "ABEV3.SA"
#tickers10 = "QUAL3.SA FLRY3.SA RADL3.SA HYPE3.SA GNDI3.SA"
#tickers11 = "VIVT3.SA"
#tickers12 = "RENT3.SA RAIL3.SA"
#tickers13 = "JBSS3.SA"
#tickers14 = "LWSA3.SA TOTS3.SA"
#tickers15 = "SUZB3.SA CSAN3.SA"
#tickers16 = "UNIP6.SA"
#tickers17 = "NTCO3.SA"
#dados_yahoo = web.get_data_yahoo(tickers, period="5y")["Adj Close"]
# + [markdown] id="Cq_bdZwSugoC"
# Exibindo dados
# + id="rLE88uaS3tUw" colab={"base_uri": "https://localhost:8080/", "height": 640} outputId="9637763d-decc-46cb-8630-1609c7c021f8"
#dados do fechamento ajustados dos ultimos cincos anos
dados_yahoo
# + id="Q2T_Ucmc5C6s" colab={"base_uri": "https://localhost:8080/", "height": 640} outputId="bb679037-6bdc-4cc3-f59a-1904e64eb059"
#tabela retorno, calcular o retorno diario de todos os ativos
retorno = dados_yahoo.pct_change()
retorno
# + id="pqjdmfrb5WjK" colab={"base_uri": "https://localhost:8080/", "height": 640} outputId="cc3ec529-1c92-4971-e475-0991019d1767"
#tabela retorno acumulado produtorio da tabela retorno
retorno_acumulado = (1 + retorno).cumprod()
retorno_acumulado.iloc[0] = 1
retorno_acumulado
# + id="E7n4HlxZ4Cd4" colab={"base_uri": "https://localhost:8080/", "height": 640} outputId="cb9b2a09-c11f-4c8b-9935-973a80bba05b"
#carteira R$ 1000, excluir coluna ibovespa, inserir coluna saldo e coluna retorno
carteira = 1000 * retorno_acumulado.iloc[:, :38]
carteira["saldo"] = carteira.sum(axis=1)
carteira["retorno"] = carteira["saldo"].pct_change()
carteira
# + [markdown] id="hRORuCvU38C8"
# ## 3. Resultados
# + id="Arb0t61r5ryS" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="1e403dd0-2c9e-4a40-871b-5910a625f384"
#retornos da carteira comparados ibovespa
pf.create_full_tear_sheet(carteira["retorno"], benchmark_rets=retorno["^BVSP"])
#59 meses, retorno anual composto 57%, retorno total 848%, volatividade anual 32%, sharpe 1.57, carteira abaixo saldo no máximo 49%
# + id="pBvtxDY651vb" colab={"base_uri": "https://localhost:8080/", "height": 420} outputId="5c06c247-a0b9-449e-ddeb-a516c5b1dbda"
fig, ax1 = plt.subplots(figsize=(16,8))
pf.plot_rolling_beta(carteira["retorno"], factor_returns=retorno["^BVSP"], ax=ax1)
plt.ylim((0.5, 1.4));
# + [markdown] id="0jcgZ38c8cRz"
# ## 4. Análise
| 05_Python_Finance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:datascience]
# language: python
# name: conda-env-datascience-py
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import seaborn as sns
import umap
# %matplotlib inline
#sns.set(style='white', rc={'figure.figsize':(12,8)})
import requests
import zipfile
import imageio
import os
import umap
import MulticoreTSNE
import time
import sklearn.manifold
sns.set(context='poster', rc={'figure.figsize':(12, 10)})
# -
# ### Pull the data from the internet and write it to a file
# %%time
if not os.path.isfile('coil20.zip'):
results = requests.get('http://www.cs.columbia.edu/CAVE/databases/SLAM_coil-20_coil-100/coil-20/coil-20-proc.zip')
with open("coil20.zip", "wb") as code:
code.write(results.content)
# ### Read in from file and transform into a vector space
# We add a filter to ensure that we only parse png files from our download zip.
import re
images_zip = zipfile.ZipFile('coil20.zip')
mylist = images_zip.namelist()
r = re.compile(".*\.png$")
filelist = list(filter(r.match, mylist))
# Next we pull the object id's from the file names using pandas
if not os.path.isfile('coil-20-proc/obj10__1.png'):
# !unzip coil20.zip
# %%time
feature_vectors = []
for filename in filelist:
im = imageio.imread(filename)
feature_vectors.append(im.flatten())
# ### Now we have our data in a list of vectors. Let's extract the object id's from the files and cast to data frame (in case we want to explore things further)
#
# We could leave this data in numpy arrays for improved efficiency but at data sizes this small the added functionality of pandas for potential exploration is appealing.
labels = pd.Series(filelist).str.extract("obj([0-9]+)", expand=False)
data = np.array(pd.DataFrame(feature_vectors, index=labels))
data.shape
# ### Now let's use UMAP to embed these points into a two dimensional space.
# +
timings = []
dims = []
dimensions = [2,4,6,8]
for dim in dimensions:
for rep in range(5):
start = time.time()
embedding = umap.UMAP(n_components=dim, n_neighbors=5, metric='euclidean').fit_transform(data)
total_time = time.time()- start
dims.append(dim)
timings.append(total_time)
print("dim: {} timing:{}".format(dim,total_time))
# -
d_umap = pd.DataFrame({'dimension':dims,'timing':timings})
# ### T-SNE
# +
tsne_timings = []
tsne_dims = []
dimensions = [2,4,6,8]
for dim in dimensions:
for rep in range(5):
start = time.time()
fit_tsne = MulticoreTSNE.MulticoreTSNE(n_jobs=1, n_components=dim).fit_transform(data)
#embedding = umap.UMAP(n_components=dim, n_neighbors=5, metric='euclidean').fit_transform(data)
total_time = time.time()- start
tsne_dims.append(dim)
tsne_timings.append(total_time)
print("dim: {} timing:{}".format(dim,total_time))
# -
#d.to_csv('timingScalingWithEmbeddingDimension_coil.csv')
d_tsne = pd.DataFrame({'dimension':tsne_dims, 'timing':tsne_timings})
# +
fig = plt.figure()
ax = fig.add_subplot(111)
sns.regplot(x='dimension',y='timing',data=np.log2(d_tsne),ax=ax, label='t-SNE', order=2)
sns.regplot(x='dimension',y='timing',data=np.log2(d_umap),ax=ax, label='UMAP')
plt.ylabel('timing log2(seconds)')
plt.xlabel('log2(dimension)')
plt.legend(loc='best', frameon=True, fancybox=True, fontsize=20)
plt.title('Scaling with dimensionality on Coil20', fontsize=20)
plt.savefig('scalingWithDimensionalityComparisonCoil20_log.png')
# +
fig = plt.figure()
ax = fig.add_subplot(111)
sns.regplot(x='dimension',y='timing',data=d_tsne,ax=ax, label='t-SNE', order=2)
sns.regplot(x='dimension',y='timing',data=d_umap,ax=ax, label='UMAP')
plt.ylabel('timing (seconds)')
plt.legend(loc='best', frameon=True, fancybox=True, fontsize=20)
plt.title('Scaling with dimensionality on Coil20', fontsize=20)
plt.savefig('scalingWithDimensionalityComparisonCoil20.png')
# -
# ## Test UMAP scaling to much higher dimensions.
# +
large_timings = []
large_dims = []
dimensions = [2,4,8, 16, 32,64,128,256,512,1028]
for dim in dimensions:
for rep in range(5):
start = time.time()
embedding = umap.UMAP(n_components=dim, n_neighbors=5, metric='euclidean').fit_transform(data)
total_time = time.time()- start
large_dims.append(dim)
large_timings.append(total_time)
print("dim: {} timing:{}".format(dim,total_time))
# -
d_large = pd.DataFrame({'dimension':large_dims, 'timing':large_timings})
# +
fig = plt.figure()
ax = fig.add_subplot(111)
sns.regplot(x='dimension',y='timing',data=d_large,ax=ax, label='UMAP')
plt.ylabel('timing (seconds)')
#plt.legend(loc='best', frameon=True, fancybox=True, fontsize=20)
plt.title('UMAP scaling with dimensionality on Coil20', fontsize=20)
plt.savefig('scalingWithDimensionalityUMAPCoil20.png')
# -
| UMAP Coil20-EmbeddingDimensionScaling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import math
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn import svm
from sklearn.ensemble import BaggingClassifier
basePre = pd.read_csv('./bases/base_pre.csv')
baseScaled = pd.read_csv('./bases/base_scaled.csv')
basePCACompleta = pd.read_csv('./bases/base_train_completa.csv')
basePCAInversa = pd.read_csv('./bases/base_train_correlacao_inversa.csv')
basePCAProporcional = pd.read_csv('./bases/base_train_correlacao_proporcional.csv')
basePca70 = pd.read_csv('./bases/base_train_70.csv')
basePca50 = pd.read_csv('./bases/base_train_50.csv')
cv = 5
Y = basePre['target']
clf = svm.SVC(gamma='scale', kernel='linear', C=1)
clf
# -
# SINGLE EXECUTION
# Applying in baseScaled
sc = cross_val_score(clf, baseScaled, Y, cv=cv)
accArray = np.array([[sc.mean(), sc.std()*2]])
# Applying in basePCAInversa
sc = cross_val_score(clf, basePCAInversa, Y, cv=cv)
accArray = np.append(accArray, [[sc.mean(), sc.std()*2]], axis=0)
# Applying in basePCAProporcional
sc = cross_val_score(clf, basePCAProporcional, Y, cv=cv)
accArray = np.append(accArray, [[sc.mean(), sc.std()*2]], axis=0)
# PCA com 70%
sc = cross_val_score(clf, basePca70, Y, cv=cv)
accArray = np.append(accArray, [[sc.mean(), sc.std()*2]], axis=0)
# PCA com 50%
sc = cross_val_score(clf, basePca50, Y, cv=cv)
accArray = np.append(accArray, [[sc.mean(), sc.std()*2]], axis=0)
dfAcc = pd.DataFrame(accArray, columns=['mean', 'std'], index=None)
dfAcc = (dfAcc*100).apply(np.floor)
dfAcc
# +
from plt import *
single(dfAcc, 'svmSingle.png', '#0000FF', '#FFA500')
# -
# BAGGING com a melhor single
# +
model = BaggingClassifier(clf, n_estimators=5, random_state=0)
sc = cross_val_score(model, basePca70, Y, cv=cv)
model
# -
accArray = np.array([[sc.mean(), sc.std()*2]])
# +
model = BaggingClassifier(clf, n_estimators=10, random_state=0)
sc = cross_val_score(model, basePca70, Y, cv=cv)
model
# -
accArray = np.append(accArray, [[sc.mean(), sc.std()*2]], axis=0)
# +
model = BaggingClassifier(clf, n_estimators=20, random_state=0)
sc = cross_val_score(model, basePca70, Y, cv=cv)
model
# -
accArray = np.append(accArray, [[sc.mean(), sc.std()*2]], axis=0)
# +
model = BaggingClassifier(clf, n_estimators=30, random_state=0)
sc = cross_val_score(model, basePca70, Y, cv=cv)
model
# -
accArray = np.append(accArray, [[sc.mean(), sc.std()*2]], axis=0)
dfAcc = pd.DataFrame(accArray, columns=['mean', 'std'], index=None)
dfAcc = (dfAcc*100).apply(np.floor)
dfAcc
bagging(dfAcc, 'svmBagging.png', '#0000FF', '#FFA500')
| alg-SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import copy
import math
import os
import torch
from torch.utils.data import DataLoader
import numpy as np
import matplotlib.pyplot as plt
import librosa
import librosa.display
from SAMAF import SAMAF
from SinhalaSongsDataset import SinhalaSongsDataset
# -
# ### Utility Methods
def draw_mfccs(*mfccs):
plots = len(mfccs)
plt.figure()
for i, mfcc in enumerate(mfccs):
plt.subplot(1, plots, i+1)
librosa.display.specshow(mfcc.transpose(0,1).numpy(), x_axis="time")
plt.title("MFCC")
plt.tight_layout()
plt.colorbar()
# ## Trainer
def train_model(train_dataset, validation_dataset, epochs, device, embedding_dimension=128, save_path="", start_state=None):
def mseLoss(pred, true):
return torch.nn.functional.mse_loss(pred, true)
def hashLoss(embeddings):
embeddings_repeated_1 = embeddings.repeat(1, embeddings.shape[1], 1) # [1,2,3] => [1,1,2,2,3,3]
embeddings_repeated_2 = embeddings.repeat_interleave(embeddings.shape[1], dim=1) # [1,2,3] => [1,2,3,1,2,3]
cosine_similarity = torch.nn.functional.cosine_similarity(embeddings_repeated_1, embeddings_repeated_2, dim=2)
# print("Cosine similarity values", cosine_similarity.shape)
cosine_similarity = cosine_similarity.view(-1, embeddings.shape[1], embeddings.shape[1])
multiplier = (torch.ones(embeddings.shape[1]) - torch.eye(embeddings.shape[1])).unsqueeze(0)
cosine_similarity = cosine_similarity * multiplier * (1/0.55)
cosine_similarity[cosine_similarity < 0] = 0.0
cosine_similarity[cosine_similarity > 1] = 1.0
# print("Cosine similarity values", cosine_similarity.shape)
# print(cosine_similarity[0])
l2_norm = torch.linalg.norm(embeddings.unsqueeze(1)-embeddings.unsqueeze(2), ord=2, dim=3)
l2_norm_squared = torch.square(l2_norm)
# print("Squared L2 Norm ", l2_norm_squared.shape)
neumerator = torch.sum(cosine_similarity * l2_norm_squared, dim=(1,2))
# print("Neumerator ", neumerator.shape)
denominator = torch.count_nonzero(cosine_similarity.detach(), dim=(1,2))
# print("Denominator ", denominator.shape)
return torch.mean(neumerator / denominator)
def bitwiseEntropyLoss(embeddings):
# TODO: Implement
return 0
model = SAMAF(embedding_dim=embedding_dimension).to(device)
optimizer = torch.optim.Adagrad(model.parameters(), lr=1e-2)
history = dict(train=[], validation=[])
best_model_weights = copy.deepcopy(model.state_dict())
best_loss = 100000000.0
start_epoch = 1
if start_state:
model.load_state_dict(start_state["model_state_dict"])
optimizer.load_state_dict(start_state["optimizer_state_dict"])
start_epoch = start_state["epoch"]
history = start_state["history"]
best_model_weights = start_state["best_model_weights"]
best_loss = start_state["best_loss"]
for epoch in range(start_epoch, epochs+1):
train_losses = []
model = model.train()
for i, (_, seq_true) in enumerate(train_dataset):
seq_true = seq_true.to(device)
optimizer.zero_grad()
embeddings, seq_pred = model(seq_true)
loss = 1.0 * mseLoss(seq_pred, seq_true) + 1.0 * hashLoss(embeddings) + 1.0 * bitwiseEntropyLoss(embeddings) # criterion(seq_pred, seq_true).to(device)
loss.backward()
optimizer.step()
train_losses.append(loss.item())
if i % 100 == 99:
print("Epoch {} batch {}: train loss {}".format(epoch, i+1, loss.item()))
validation_losses=[]
model = model.eval()
with torch.no_grad():
for i, (_, seq_true) in enumerate(validation_dataset):
seq_true = seq_true.to(device)
_, seq_pred = model(seq_true)
loss = 1.0 * mseLoss(seq_pred, seq_true) + 1.0 * hashLoss(embeddings) + 1.0 * bitwiseEntropyLoss(embeddings)
validation_losses.append(loss.item())
if i % 100 == 99:
print("Epoch {} batch {}: validation loss {}".format(epoch, i+1, loss.item()))
train_loss = np.mean(train_losses)
validation_loss = np.mean(validation_losses)
history['train'].append(train_loss)
history['validation'].append(validation_loss)
print("Epoch {}: train loss {}, validation loss {}".format(epoch, train_loss, validation_loss))
torch.save({
"epoch": epoch,
"model_state_dict": model.state_dict(),
"optimizer_state_dict": optimizer.state_dict(),
"train_loss": train_loss,
"validation_loss": validation_loss,
"history": history,
"best_model_weights": best_model_weights,
"best_loss": best_loss
}, os.path.join(save_path, "snapshot-{}.pytorch".format(epoch)))
if validation_loss < best_loss:
best_loss = validation_loss
best_model_weights = copy.deepcopy(model.state_dict())
x = [*range(1, len(history['train'])+1)]
plt.clf()
plt.plot(x, history['train'], label="Train Loss")
plt.plot(x, history['validation'], label="Validation Loss")
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.title("Model Performance upto epoch {}".format(epoch))
plt.legend()
plt.savefig(os.path.join(save_path, "model-performance-{}.png".format(epoch)))
return best_model_weights, history
# + tags=[]
# test_dataset = SinhalaSongsDataset(root_dir="/home/pasinducw/Downloads/Research-Datasets/Sinhala-Songs/features", trim_seconds=40, test=True)
# dataloader = DataLoader(test_dataset, batch_size=128, shuffle=False)
# (song_id, mfccs) = iter(dataloader).next()
# print(song_id.shape)
# print(mfccs.shape)
# transformed = mfccs.view(-1, 1, 100, 13).squeeze(1)
# print(transformed.shape)
# draw_mfccs(transformed[50], transformed[51])
# draw_mfccs(mfccs[1][10], mfccs[1][11])
# model = SAMAF()
# embeddings, decoder_outputs = model(mfccs)
# print("Input shape ", mfccs.shape)
# print("Embeddings ", embeddings.shape)
# print("Decoder outputs ", decoder_outputs.shape)
# torch.nn.functional.mse_loss(decoder_outputs, mfccs)
# +
# Dataset tests
train_dataset = SinhalaSongsDataset(root_dir="/home/pasinducw/Downloads/Research-Datasets/Sinhala-Songs/features", trim_seconds=40)
train_dataloader = torch.utils.data.DataLoader(train_dataset, batch_size=256, shuffle=False)
validation_dataset = SinhalaSongsDataset(root_dir="/home/pasinducw/Downloads/Research-Datasets/Sinhala-Songs/features", trim_seconds=40, validation=True)
validation_dataloader = torch.utils.data.DataLoader(validation_dataset, batch_size=256, shuffle=False)
device = torch.device("cpu")
best_model, history = train_model(train_dataloader, validation_dataloader, 50, device, 256, "/home/pasinducw/Documents/research/src/samaf/data/L1-D256-B256-E50-EXP1")
# -
device = torch.device("cpu")
checkpoint = torch.load("../data/L1-D196-B20-E100-EXP2/snapshot-3.pytorch", map_location=device)
test_model = SAMAF(embedding_dim=196)
test_model.load_state_dict(checkpoint['best_model_weights'])
test_dataloader = torch.utils.data.DataLoader(SinhalaSongsDataset(root_dir="/home/pasinducw/Downloads/Research-Datasets/Sinhala-Songs/features", trim_seconds=10, test=True), shuffle=True)
with torch.no_grad():
_, mfccs = iter(test_dataloader).next()
embeddings, seq_pred = test_model(mfccs)
print(mfccs.shape, seq_pred.shape)
draw_mfccs(mfccs[0, 0], seq_pred[0,0])
# draw_mfccs(mfccs[0,1], seq_pred[0,1])
a = torch.randn(3,3)
b = lambda v: v if 0 <= v <= 0.55 else 0
c = torch.vmap(b)
| samaf/model/trainer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. What is the most common Unicode encoding when moving data between systems?
# ##### Ans: UTF-8
# #### 2. What is the decimal (Base-10) numeric value for the upper case letter "G" in the ASCII character set?
print(ord('G'))
# ##### Ans: 71
# #### 3. What word does the following sequence of numbers represent in ASCII:
#
# >108, 105, 115, 116
# ##### Ans: list
# #### 4. How are strings stored internally in Python 3?
# ##### Ans: Unicode
# #### 5. When reading data across the network (i.e. from a URL) in Python 3, what method must be used to convert it to the internal format used by strings?
# ##### Ans: decode()
| Coursera/Using Databases with Python/Week-1/Quiz/Using-Encoded-Data-in-Python-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import tensorflow as tf
from matplotlib import pyplot as plt
from tensorflow.contrib import learn
from sklearn.metrics import mean_squared_error
from lstm import lstm_model
from data_processing import x_sin, generate_data,
# -
LOG_DIR = './ops_logs/x_sin'
TIMESTEPS = 10
RNN_LAYERS = [{'num_units': 10}, {'num_units': 5}]
DENSE_LAYERS = None
TRAINING_STEPS = 100000
PRINT_STEPS = TRAINING_STEPS / 10
BATCH_SIZE = 100
regressor = learn.Estimator(model_fn=lstm_model(TIMESTEPS, RNN_LAYERS, DENSE_LAYERS),
model_dir=LOG_DIR)
X, y = generate_data(x_sin, np.linspace(0, 100, 10000, dtype=np.float32), TIMESTEPS, seperate=False)
# create a lstm instance and validation monitor
validation_monitor = learn.monitors.ValidationMonitor(X['val'], y['val'],
every_n_steps=PRINT_STEPS,
early_stopping_rounds=10000)
regressor.fit(X['train'], y['train'],
monitors=[validation_monitor],
batch_size=BATCH_SIZE,
steps=TRAINING_STEPS)
predicted = regressor.predict(X['test'])
rmse = np.sqrt(((predicted - y['test']) ** 2).mean(axis=0))
score = mean_squared_error(predicted, y['test'])
print ("MSE: %f" % score)
plot_predicted, = plt.plot(predicted, label='predicted')
plot_test, = plt.plot(y['test'], label='test')
plt.legend(handles=[plot_predicted, plot_test])
plt.plot(regressor.predict(X['train']))
plt.plot(y['train'])
| lstm_xsin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import mysql.connector
from IPython.display import Markdown, display
db = mysql.connector.connect(
host="localhost",
user="root",
password="<PASSWORD>",
port="3306",
database="serlo"
)
import pandas as pd
def anzahl_show(title, ids):
display(Markdown(f"# {title}"))
ids_sql = ",".join(map(str, ids))
df = pd.read_sql(f"""
select actor_id, id from event_log
where event_id in ({ids_sql})
and date > now()-interval 3 month
""", db)
df = df.groupby("actor_id").count()
return len(df[df["id"] > 10])
anzahl_show("Aktuelle Anzahl aktiver Autor:innen", [5, 4])
# -
anzahl_show("Aktuelle Anzahl aktiver Reviewer:innen", [6, 11])
anzahl_show("Aktuelle Anzahl aktiver Taxonomybauer:innen", [1,2,12,15,17])
anzahl_show("Aktuelle Anzahl aktiver Moderator:innen", [9,14,16])
anzahl_show("Aktuelle Anzahl aktiver Admins (mit vorsicht zu genießen)", [10,13])
| 2021/2021-06-30 Anzahl aktuell aktive Community.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # _*Qiskit Aqua: qGANs for Loading Random Distributions*_
#
# The latest version of this notebook is available on https://github.com/Qiskit/qiskit-iqx-tutorials.
#
# ***
# ### Contributors
# <NAME><sup>[1,2]</sup>, <NAME><sup>[1]</sup>
# ### Affiliation
# - <sup>[1]</sup>IBMQ
# - <sup>[2]</sup>ETH Zurich
#
# ### Introduction
# Given $k$-dimensional data samples, we employ a quantum Generative Adversarial Network (qGAN) to learn the data's underlying random distribution and to load it directly into a quantum state:
#
# $$ \big| g_{\theta}\rangle = \sum_{j=0}^{2^n-1} \sqrt{p_{\theta}^{j}}\big| j \rangle $$
#
# where $p_{\theta}^{j}$ describe the occurrence probabilities of the basis states $\big| j\rangle$.
#
# The aim of the qGAN training is to generate a state $\big| g_{\theta}\rangle$ where $p_{\theta}^{j}$, for $j\in \left\{0, \ldots, {2^n-1} \right\}$, describe a probability distribution that is close to the distribution underlying the training data $X=\left\{x^0, \ldots, x^{k-1} \right\}$.
#
# For further details please refer to <a href="https://arxiv.org/abs/1904.00043">Quantum Generative Adversarial Networks for Learning and Loading Random Distributions. Zoufal, <NAME>. 2019.</a>
#
# How to use a trained qGAN in an application, i.e., pricing of financial derivatives, is illustrated here:
# <a href="../finance/machine_learning/qgan_option_pricing.ipynb">qGAN Option Pricing</a>.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import time
start = time.time()
from torch import optim
from qiskit import QuantumRegister, QuantumCircuit
from qiskit.aqua.components.optimizers import ADAM
from qiskit.aqua.components.uncertainty_models import UniformDistribution, UnivariateVariationalDistribution
from qiskit.aqua.components.variational_forms import RY
from qiskit.aqua.algorithms.adaptive import QGAN
from qiskit.aqua.components.neural_networks.quantum_generator import QuantumGenerator
from qiskit.aqua.components.neural_networks.numpy_discriminator import NumpyDiscriminator
from qiskit.aqua import aqua_globals, QuantumInstance
from qiskit.aqua.components.initial_states import Custom
from qiskit import BasicAer
# -
# ### Load the Training Data
# First, we need to load the $k$-dimensional training data samples (here k=1). <br/>
# Next, the data resolution is set, i.e. the min/max data values and the number of qubits used to represent each data dimension.
# +
# Number training data samples
N = 1000
# Load data samples from log-normal distribution with mean=1 and standard deviation=1
mu = 1
sigma = 1
real_data = np.random.lognormal(mean = mu, sigma=sigma, size=N)
# Set the data resolution
# Set upper and lower data values as list of k min/max data values [[min_0,max_0],...,[min_k-1,max_k-1]]
bounds = np.array([0.,3.])
# Set number of qubits per data dimension as list of k qubit values[#q_0,...,#q_k-1]
num_qubits = [2]
k = len(num_qubits)
# -
# ### Initialize the qGAN
# The qGAN consists of a quantum generator $G_{\theta}$, a variational quantum circuit, and a classical discriminator $D_{\phi}$, a neural network. <br/>
# To implement the quantum generator, we choose a depth-$1$ variational form that implements $R_Y$ rotations and $CZ$ gates which takes a uniform distribution as an input state. Notably, for $k>1$ the generator's parameters must be chosen carefully. For example, the circuit depth should be $>1$ because higher circuit depths enable the representation of more complex structures.<br/>
# The classical discriminator is given by a $3$-layer neural network that applies linear transformations, leaky ReLU functions in the hidden layers and a sigmoid function in the output layer. Notably, the neural network is implemented with PyTorch. Please refer to https://pytorch.org/get-started/locally/ for PyTorch installation instructions.<br/>
# Here, both networks are updated with the ADAM optimization algorithm.
# +
# Set number of training epochs
# Note: The algorithm's runtime can be shortened by reducing the number of training epochs.
num_epochs = 3000
# Batch size
batch_size = 100
# Initialize qGAN
qgan = QGAN(real_data, bounds, num_qubits, batch_size, num_epochs, snapshot_dir=None)
qgan.seed = 1
# Set quantum instance to run the quantum generator
quantum_instance = QuantumInstance(backend=BasicAer.get_backend('statevector_simulator'))
# Set entangler map
entangler_map = [[0, 1]]
# Set an initial state for the generator circuit
init_dist = UniformDistribution(sum(num_qubits), low=bounds[0], high=bounds[1])
q = QuantumRegister(sum(num_qubits), name='q')
qc = QuantumCircuit(q)
init_dist.build(qc, q)
init_distribution = Custom(num_qubits=sum(num_qubits), circuit=qc)
var_form = RY(int(np.sum(num_qubits)), depth=1, initial_state = init_distribution,
entangler_map=entangler_map, entanglement_gate='cz')
# Set generator's initial parameters
init_params = aqua_globals.random.rand(var_form._num_parameters) * 2 * np.pi
# Set generator circuit
g_circuit = UnivariateVariationalDistribution(int(sum(num_qubits)), var_form, init_params,
low=bounds[0], high=bounds[1])
# Set quantum generator
qgan.set_generator(generator_circuit=g_circuit)
# Set classical discriminator neural network
discriminator = NumpyDiscriminator(len(num_qubits))
qgan.set_discriminator(discriminator)
# -
# ### Run the qGAN Training
# During the training the discriminator's and the generator's parameters are updated alternately w.r.t the following loss functions:
# $$ L_G\left(\phi, \theta\right) = -\frac{1}{m}\sum\limits_{l=1}^{m}\left[\log\left(D_{\phi}\left(g^{l}\right)\right)\right] $$
# and
# $$ L_D\left(\phi, \theta\right) =
# \frac{1}{m}\sum\limits_{l=1}^{m}\left[\log D_{\phi}\left(x^{l}\right) + \log\left(1-D_{\phi}\left(g^{l}\right)\right)\right], $$
# with $m$ denoting the batch size and $g^l$ describing the data samples generated by the quantum generator.
#
# Please note that the training will take a while ($\sim 20$ min).
# +
# Run qGAN
qgan.run(quantum_instance)
# Runtime
end = time.time()
print('qGAN training runtime: ', (end - start)/60., ' min')
# -
# ### Training Progress & Outcome
# Now, we plot the evolution of the generator's and the discriminator's loss functions during the training as well as the progress in the relative entropy between the trained and the target distribution.
# <br/> Finally, we also compare the cumulative distribution function (CDF) of the trained distribution to the CDF of the target distribution.
# +
# Plot progress w.r.t the generator's and the discriminator's loss function
t_steps = np.arange(num_epochs)
plt.figure(figsize=(6,5))
plt.title("Progress in the loss function")
plt.plot(t_steps, qgan.g_loss, label = "Generator loss function", color = 'mediumvioletred', linewidth = 2)
plt.plot(t_steps, qgan.d_loss, label = "Discriminator loss function", color = 'rebeccapurple', linewidth = 2)
plt.grid()
plt.legend(loc = 'best')
plt.xlabel('time steps')
plt.ylabel('loss')
plt.show()
# Plot progress w.r.t relative entropy
plt.figure(figsize=(6,5))
plt.title("Relative Entropy ")
plt.plot(np.linspace(0, num_epochs, len(qgan.rel_entr)), qgan.rel_entr, color ='mediumblue', lw=4, ls=':')
plt.grid()
plt.xlabel('time steps')
plt.ylabel('relative entropy')
plt.show()
#Plot the PDF of the resulting distribution against the target distribution, i.e. log-normal
log_normal = np.random.lognormal(mean=1, sigma=1, size=100000)
log_normal = np.round(log_normal)
log_normal = log_normal[log_normal <= bounds[1]]
temp = []
for i in range(int(bounds[1]+1)):
temp += [np.sum(log_normal==i)]
log_normal = np.array(temp / sum(temp))
plt.figure(figsize=(6,5))
plt.title("CDF")
samples_g, prob_g = qgan.generator.get_output(qgan.quantum_instance, shots=10000)
samples_g = np.array(samples_g)
samples_g = samples_g.flatten()
num_bins = len(prob_g)
plt.bar(samples_g, np.cumsum(prob_g), color='royalblue', width= 0.8, label='simulation')
plt.plot( np.cumsum(log_normal),'-o', label='log-normal', color='deepskyblue', linewidth=4, markersize=12)
plt.xticks(np.arange(min(samples_g), max(samples_g)+1, 1.0))
plt.grid()
plt.xlabel('x')
plt.ylabel('p(x)')
plt.legend(loc='best')
plt.show()
# -
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
| qiskit/advanced/aqua/machine_learning/qgans_for_loading_random_distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.Objectives
# <NAME>, Helsinki Metropolia University of Applied Sciences
# Cognitive Systems for Health Technology Applications
# The object is to create and train a dense neural network to predict the presence of heart disease
# Edited 7.2.2018
# # 2. Required libraries
# Importing the required libraries
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas as pd
import sklearn as sk
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
import keras
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout
from keras import models
from keras import layers
# -
# # 3. Data description and preprocessing
# +
filename = 'https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.cleveland.data'
filename2 = 'https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.hungarian.data'
filename3 = 'https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.va.data'
filename4 = 'https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.switzerland.data'
colnames = ['age','sex','cp','tresbps','chol',
'fbs','restecg','thalach','exang',
'oldpeak','slope','ca','thal','num']
# read data to data frame
df1 = pd.read_csv(filename,
names = colnames,
na_values = '?')
df2 = pd.read_csv(filename2,
names = colnames,
na_values = '?')
df3 = pd.read_csv(filename3,
names = colnames,
na_values = '?')
df4 = pd.read_csv(filename4,
names = colnames,
na_values = '?')
df1.head()
# +
frames = [df1,df2,df3,df4]
df = pd.concat(frames)
df.index = range(920)
# +
#Ikäjakauma
df['age'].hist(bins = np.arange(28, 90, 5))
plt.xlabel('Age (Years)')
plt.ylabel('Count')
plt.show()
# +
#Missing values
df = df.where(~np.isnan(df), df.mode(), axis = 'columns')
label = (df['num'] > 0).values
# +
#Data normalization
df = (df - df.min())/(df.max() - df.min())
df.head()
# -
columns = ['age', 'sex', 'cp', 'tresbps',
'chol', 'fbs', 'restecg', 'thalach',
'exang', 'oldpeak', 'slope', 'ca', 'thal']
data = df[columns].values
# +
ndata = 920
ntrain= int(0.8*ndata) #80 testing, 10 training
train_data = data[:ntrain]
train_labels = label[:ntrain]
test_data = data[ntrain:]
test_labels = label[ntrain:]
# -
# # 4. Modeling and compilation
# Modeling and compilating the network
# +
from keras import models, layers
network = models.Sequential()
network.add(layers.Dense(6, activation = 'relu' , input_shape=(13, )))
network.add(layers.Dense(6, activation = 'relu'))
network.add(layers.Dense(1, activation = 'relu')) # output 0 ja 1
# +
from keras.utils.np_utils import to_categorical
categorical_labels = to_categorical(label, num_classes=None)
# -
network.compile(optimizer ='rmsprop',
loss = 'binary_crossentropy',
metrics = ['accuracy'])
# # 5. Training and Validation
# Training the network with fit method. 120 epochs and batch size 128
# +
N = 120
h = network.fit(train_data, train_labels, verbose = 0, epochs = 120, batch_size = 128, validation_data=(test_data, test_labels))
# -
# # 6. Evaluation
# Evaluate the network with network.evaluate. Testing the losses and the accuracy
# +
test_loss, test_acc = network.evaluate(test_data, test_labels, batch_size = 128)
print('test_accuracy', test_acc)
# -
# # 7. Results
# Movin the results to accuracy plot
# +
# Plot the results
epochs = range(1, N + 1)
acc = h.history['acc']
val_acc = h.history['val_acc']
loss = h.history['loss']
val_loss = h.history['val_loss']
# Accuracy plot
plt.figure(figsize = (20, 5))
plt.plot(epochs, acc, 'bo', label='Training')
plt.plot(epochs, val_acc, 'b', label = 'Validation')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.ylim([0, 1])
plt.grid()
plt.legend()
plt.show()
# -
# # 8. Conclusions
# The accuracy was pretty bad. I tested with many different combinations and these were one of the best results. The accuracy gets stable after 80 epochs and after that it looks pretty stable. I wouldnt trust this results because the accuracy doesnt even go to 80%
| Case1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from pandas import Series, DataFrame
# # Series
sample_pandas_data = pd.Series([0,10,20,30,40,50,60,70,80,90])
print(sample_pandas_data)
#印出資料值
print('資料值:',sample_pandas_data.values)
#印出索引值
print('資料值:',sample_pandas_data.index)
# # Specify index in Series
# +
sample_pandas_index_data = pd.Series(
[0, 10,20,30,40,50,60,70,80,90],
#輸入自訂的文字index
['a','b','c','d','e','f','g','h','i','j']
)
print(sample_pandas_index_data)
# -
print('資料值:', sample_pandas_index_data.values)
print('索引值:', sample_pandas_index_data.index)
print(sample_pandas_index_data.index[3])
# # DataFrame
# +
attri_data1 = {'ID':['100','101','102','103','104'],
'City':['Taipei','Taipei','Hsinchu','Tainan','Kaohsiung'],
'Birth_year':[1990,1989,1992,1997,1982],
'Name':['Sally','Hanks','Hannah','Kitty','Steve']}
attri_data_frame1 = DataFrame(attri_data1)
print(attri_data_frame1)
# -
# # Specify Index in DataFrame
attri_data_frame_index1 = DataFrame(attri_data1,
#輸入自訂的index
index=['a','b','c','d','e']
)
print(attri_data_frame_index1)
attri_data_frame_index1
# # Transpose
#attri_data_frame1的行列交換
attri_data_frame1.T
# # Get One Column
#取得Birth_year行
#attri_data_frame1.Birth_year
# # Get Many Column
#取得ID與Birth_year行
attri_data_frame1[['ID','Birth_year']]
# # Filter
# attri_data_frame1中找出City為Taipei的資料
attri_data_frame1[attri_data_frame1['City'] == 'Taipei']
attri_data_frame1['City'] == 'Taipei'
# # Filter for Many Requiements
#attri_data_frame1中找出City為台北和台南的資料
attri_data_frame1[attri_data_frame1['City'].isin(['Taipei','Tainan'])]
# # Pratice
#找出attri_data_frame1的Birth_year小於1990
attri_data_frame1[attri_data_frame1['Birth_year'] <= 1990]
# # Drop Column
#去掉Birth_year這一行
print(attri_data_frame1.drop([0],axis = 0))
print(attri_data_frame1.drop(['Birth_year'],axis = 1))
# +
attri_data1 = {'ID':['100','101','102','103','104'],
'City':['Taipei','Taipei','Hsinchu','Tainan','Kaohsiung'],
'Birth_year':[1990,1989,1992,1997,1982],
'Name':['Sally','Hanks','Hannah','Kitty','Steve']}
attri_data_frame1 = DataFrame(attri_data1)
print(attri_data_frame1)
attri_data_frame1.drop(['Birth_year'], axis = 1)
print(attri_data_frame1)
#使用assign,使得attri_data_frame1的資料中真正拿掉Birth_year行
attri_data_frame1 = attri_data_frame1.drop(['Birth_year'], axis = 1)
print(attri_data_frame1)
# -
# # Combine DataFrame
# +
#準備資料
attri_data1 = {'ID':['100','101','102','103','104'],
'City':['Taipei','Taipei','Hsinchu','Tainan','Kaohsiung'],
'Birth_year':[1990,1989,1992,1997,1982],
'Name':['Sally','Hanks','Hannah','Kitty','Steve']}
attri_data_frame1 = DataFrame(attri_data1)
print(attri_data_frame1)
attri_data2 = {'ID':['100','101','102','105','107'],
'Math':[50,43,33,76,98],
'English':[90,30,20,50,30],
'Sex':['M','F','F','M','M']}
attri_data_frame2 = DataFrame(attri_data2)
print(attri_data_frame2)
# -
#合併資料
pd.merge(attri_data_frame1,attri_data_frame2)
# # Statistics
#將「Sex」的行設為基準來計算出數學成績的平均分數
attri_data_frame2.groupby('Sex')['Math'].mean()
# # Practice
#將「Sex」的行設為基準來計算出英文成績的最大值與最小值。
print(attri_data_frame2.groupby('Sex')['English'].max())
print(attri_data_frame2.groupby('Sex')['English'].min())
# # Sort
#準備資料
attri_data2 = {'ID':['100','101','102','103','104'],
'City':['Taipei','Taipei','Hsinchu','Tainan','Kaohsiung'],
'Birth_year':[1990,1989,1992,1997,1982],
'Name':['Sally','Hanks','Hannah','Kitty','Steve']}
attri_data_frame_index2 = DataFrame(attri_data2,index=['e','b','a','d','c'])
attri_data_frame_index2
#sort by index
attri_data_frame_index2.sort_index()
#sort by values
attri_data_frame_index2.Birth_year.sort_values()
# # Check nan
#確認是否存在該值
attri_data_frame_index2.isin(['Taipei'])
# +
# 處理遺漏值
# 將Name設成nan
import numpy as np
attri_data_frame_index2['Name'] = np.nan
#判斷是否為nan
attri_data_frame_index2.isnull()
# -
#計算nan的總數
attri_data_frame_index2.isnull().sum()
# # 練習
# +
np.random.seed(2)
array2 = np.random.randint(2,size = 100)
array = []
for i in range(0,100):
if array2[i] == 0:
array.append('F');
else:
array.append('M');
np.random.seed(3)
array3 = np.random.normal(1000,10,size = 100)
attri_data = { 'ID':range(1,101),
'Sex':array,
'Money':array3
}
attri_data_frame_index2 = DataFrame(attri_data)
attri_data_frame_index2
print(attri_data_frame_index2[attri_data_frame_index2['Money'] == attri_data_frame_index2['Money'].min()])
print(attri_data_frame_index2[attri_data_frame_index2['Money'] > 1010])
print(attri_data_frame_index2[attri_data_frame_index2['Money'] > 1010].sort_values('Money',ascending = False))
# -
# # 資料補值:遺漏資料的處理
#資料的準備
import numpy as np
from numpy import nan as NA
import pandas as pd
np.random.seed(0)
df = pd.DataFrame(np.random.rand(10, 4))
# 設定為NA
df.iloc[1,0] = NA
df.iloc[2:3,2] = NA
df.iloc[5:,3] = NA
print(df)
#成批刪除(list-wise deletion)
df.dropna()
#逐對刪除 拿想要的 再刪除
df[[0,1]].dropna()
#填補
print(df.fillna(0))
#拿前一個值
print(df.fillna(method = 'ffill'))
#平均值來填補
print(df.fillna(df.mean()))
# 練習
import numpy as np
from numpy import nan as NA
import pandas as pd
import numpy.random as random
random.seed(0)
df2 = pd.DataFrame(np.random.rand(15, 6))
df2.iloc[2,0] = NA
df2.iloc[5:8,2] = NA
df2.iloc[7:9,3] = NA
df2.iloc[10,5] = NA
df2
df2.dropna()
print(df2.fillna(0))
print(df2.fillna(df2.mean()))
# # 資料標註
# +
#用labal box 標註
| climb/2Pandas_class_forstudents.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
#
# This notebook generates fake data for a few of the questions! If you can reverse engineer it within 35 minutes, props to you. Otherwise, still props to you. It means you can read code and know enough math to understand whats going on! I'd say thats pretty great.
# +
import random
import numpy as np
import pandas as pd
from faker import Faker
fake = Faker()
# -
# ### Q3 Data Generation
Q3_FILE = './data/Q3-Sentence-Classifier-Results.pickle'
fake_cateogories = ["class_" + str(i) for i in range(1,10)]
# +
def generate_probability(val):
return np.random.uniform(0, val)
def generate_fake_category_data(fake_cats):
"""
Sorry, but limited comments because I want you to struggle a little.
"""
# some variables we use to help you figure out whats going on
MEAN_OF_PRIMARY_CLASS_DIST = .73
STD_OF_DIST = .25
# what will be returned or updated
probabilities = {}
classes = fake_cats.copy()
total_probs = 0
# select primary class
primary_class = random.choice(classes)
# generate a probability for this class
prob = np.random.normal(loc = MEAN_OF_PRIMARY_CLASS_DIST,
scale = STD_OF_DIST,
size = 1)
# regenerate until its less than 1
while prob > 1:
prob = np.random.normal(loc = MEAN_OF_PRIMARY_CLASS_DIST,
scale = STD_OF_DIST,
size = 1)
classes.remove(primary_class)
total_probs += prob
probabilities[primary_class] = prob
for remaining in classes:
prob_left = 1 - total_probs
prob = generate_probability(prob_left)
probabilities[remaining] = prob
total_probs += prob
return probabilities, primary_class
# +
q3_df = pd.DataFrame()
for i in range(1,101):
id = i
proba_data, primary_class = generate_fake_category_data(fake_cateogories)
q3_df = q3_df.append(
pd.DataFrame({
'id' : [id],
'predicted_class' : [primary_class],
'probabiltiies' : [proba_data],
})
)
# -
q3_df.head()
q3_df.to_pickle(Q3_FILE)
| Data Generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Monte Carlo Methods
# +
import datetime as dt
import matplotlib.pylab as plt
import numpy as np
import pandas_datareader.data as web
import seaborn as sns
import timeit
sns.set(context='notebook')
# -
# ## Doel
#
# In de tweede helft van de workshop ga je door middel van Monte Carlo onderzoeken hoe de spreiding van een aandeel zich zal verlopen. Aandelen kunnen zich op nagenoeg oneindig veel manieren ontwikkelen. Hoe de prijs van morgen, de state, ten opzichte van vandaag eruit ziet is niet te voorspellen. Door middel van Monte Carlo gaan we het verloop honderden tot duizenden malen simuleren en een kansverdeling opstellen. Op het einde van de workshop gaan we een groot nadeel van de Monte Carlo Method ondervinden, hoeveel herhalingen van de simulatie kan jouw computer aan?
#
# Aan het einde van deze workshop is het de bedoeling dat je plots er als volgt uitzien. Deze foto zal je waarschijnlijk herkennen van de presentatie:
# <img src="attachments/photo1.png" align="center">
#
# ***
#
# ## Deel 1: Functies schrijven
# Voor dit eerste deel schrijven we een functie die het verloop van een willekeurig aandeel berekent. De return-waarde dient een lijst te zijn van lengte $n$ te zijn. Hierbij staat $n$ voor het aantal dagen dat de Monte Carlo-simulatie vooruit gaat voorspellen. In de code heet deze variabele `days`. Verder dient er een startprijs, `start_price`, voor de voorspelling te worden ingegeven. Vanaf deze prijs zal voor $n$ dagen het verloop worden voorspeld. Het laatste argument is `STD`. Dit is de standaard deviatie met welke voorspellende waardes gegeven een normaalverdeling worden gegenereerd.
#
# De formule om de prijs voor morgen, gegeven de prijs van vandaag, te berekenen is als volgt:
#
# $$ Price_{n+1} = Price_{n} \times (1 + {\mathcal{N}(0,\,\sigma)})$$
#
# Oftewel, de prijs van vandaag vermenigvuldigd met een waarde. Deze waarde zijnde $1$ plus een willekeurige getal uit een normaalverdeling met $\mu=0$ en een door de gebruiker gegeven $\sigma$. Hoe de waarde voor $\sigma$ wordt bepaald, zal verderop in het document worden toegelicht. Voorlopig is het doel om de functie werkende te krijgen, verderop in de notebook gaan we realistische waarden genereren.
def calc_price(start_price=25, days=100, STD=0.05):
# breid hier de code verder uit
return prices
# Met bovenstaande functie `calc_price` kunnen we één herhaling simuleren. Uiteraard hebben we hier niet genoeg aan.
#
# Schrijf een functie die bovenstaande functie $n$ maal herhaalt. De return waarde hiervan moet een lijst zijn. Deze lijst dient $n$ lijsten te bevatten welke ieder `days` aan elementen bevat. Oftewel, de return bestaat uit een lijst van simulaties.
def calc_all_prices(n=100, start_price=25, days=100, STD=0.05):
# breid hier de code verder uit
return all_prices
# Als het goed is kan je nu met `calc_all_prices` data genereren. Met de argumenten `n` en `days` mag je zelf experimenteren. Wij raden voorlopig 100 resp. 100 voor deze twee variabelen aan.
#
# ***
#
# ## Deel 2: Realistische waarden
#
# Voor de variabelen `start_price` en `STD` zijn echter nog geen realistische waarden gekozen. Pas onderstaande functie zo aan dat deze de laatst bekende sluitingsprijs en de standaard deviatie van de procentuele verandering van het afgelopen jaar returnen. Deze twee waarden kunnen wij vervolgens gebruiken om de functie `calc_all_prices` te verbeteren met realistischere waarden.
#
# Enkele leuke stocks om mee te experimenteren zijn:
# - NKE
# - DIS
# - CSCO
# - MCD
# - AAPL
# - IBM
#
# > Let op! Voor de standaard deviatie van de procentuele verandering (Percentage Change) van de aandelen dien je twee achtereenvolgende functies uit te voeren op de lijst `stocks`.
def get_stocks_2019(stock):
start = dt.datetime(2019, 1, 1) # Startdatum van data
end = dt.datetime(2019, 12, 31) # Einddatum van data
stocks = web.DataReader(stock, 'yahoo', start, end)['Close'] # Haalt de kolom 'Close' van stock 'stock' op
# Pas onderstaande variabelen aan
last_price = stocks[-1] # Laatst bekende prijs van het aandeel
std_of_pct_change = stocks.pct_change().std() # Standaard deviatie van de procentuele verandering van het aandeel
return last_price, std_of_pct_change
# Probeer in onderstaande cell eens om voor 100 herhalingen/simulaties, 100 dagen aan **AAPL** (Apple) stocks te voorspellen:
#
# > Let op! Het ophalen van stockdata kan enkele seconden duren.
# +
last_price, std = get_stocks_2019("AAPL")
all_prices = calc_all_prices(start_price=last_price, STD=std)
# -
# Als het goed is heb je nu een lijst met 100 simulaties van het vermoedelijke beursverloop van **AAPL**.
#
# ***
#
#
# ## Deel 3: Gegevens visualiseren
# Nu je data kunt genereren, dient dit nog te worden gevisualiseerd. Maak een Seaborn-lineplot welke lijkt op de foto bovenaan deze notebook. Op de x-as dient het tijdsverloop en op de y-as de prijs te komen. Als input dien je het resultaat van `calc_all_prices` te geven.
#
# Is er al een trend te ontdekken in de plot?
#
# > Bonuspunten: Voor degene met een mooie titel en as-labels
# +
def plot_verloop(data):
# breid hier de code verder uit
# gebruik sns.lineplot voor het plotten van iedere individuele prijs lijn
plt.show()
plot_verloop(all_prices)
# -
# Mooi! Nu kan -als het goed is- het mogelijke verloop van de stockprijs laten zien. Echter kunnen we hier nog geen conclusies uit trekken. Door 100 simulaties uit te voeren hebben we hopelijk genoeg data kunnen genereren om een globale trend te kunnen herkennen. Dit is één van de doelen van Monte Carlo Methods: uit veel repetities van een simulatie een trend kunnen halen.
#
# In onderstaande cell dien je een normaalverdeling te plotten van de verzamelde data. Ook hier geldt als input het resultaat van `calc_all_prices`. Zorg dat in de titel het gemiddelde ($\mu$) en de standaard deviatie ($\sigma$) staan.
def plot_norm_dist(data):
# breid hier de code verder uit
plt.show()
plot_norm_dist(all_prices)
# ***
#
# ## Deel 4: tijdscomplexiteit
#
# Gefeliciteerd! In de vorige delen heb je het volledige proces van Monte Carlo doorlopen. In het vierde en laatste deel van deze helft van de workshop ga je een van de nadelen van Monte Carlo ondervinden.
#
# Ga met behulp van de `timeit`-module meten hoelang het genereren van data voor verschillende samplesizes duurt. Gebruik weer de functie `calc_all_prices` Kies enkele verschillende waarden voor het aantal simulaties, $n$. Bijvoorbeeld 100, 1.000 en 10.000.
#
# En? heeft je computer al moeite met 10.000? Dit is het grote nadeel van Monte Carlo waar we het eerder in de presentatie over hadden.
last_price, std
# Time hier hoelang het duur om ```calc_all_prices()``` te runnen met n = 100. Gebruik hiervoor ```%timeit```
# Time hier hoelang het duur om ```calc_all_prices()``` te runnen met n = 1_000. Gebruik hiervoor ```%timeit```
# Time hier hoelang het duur om ```calc_all_prices()``` te runnen met n = 10_000. Gebruik hiervoor ```%timeit```
# Te zien is dat de tijd met $n$ toeneemt.
#
# ***
#
# ## Bronnen
#
# - https://www.youtube.com/watch?v=_T0l015ecK4
# - https://gym.openai.com/docs/
# - https://www.youtube.com/watch?v=TvO0Sa-6UVc
# - https://www.youtube.com/watch?v=e8ofon3sg8E
# - https://towardsdatascience.com/playing-blackjack-using-model-free-reinforcement-learning-in-google-colab-aa2041a2c13d
| MonteCarlo/Monte-Carlo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IMDB movie review sentiment example
#
# Build a quick LSTM model to learn whether a movie review is positive or negative using nutshell library
#
# Validation accuracy: 87.2%
# +
import pandas as pd
import numpy as np
from nutshell import ModelData, Learner, TextReader
# -
# ## Parse movie review txt files into lists of words
# +
# read imdb movie review files into a list
# download data from github - https://github.com/jalbertbowden/large-movie-reviews-dataset/tree/master/acl-imdb-v1
# # copy train & test subdirectories to this directory
reader = TextReader()
pos_texts = reader.read_text_files('./train/pos/*.txt')
neg_texts = reader.read_text_files('./train/neg/*.txt')
texts = pos_texts + neg_texts
labels = ([1] * len(pos_texts)) + ([0] * len(neg_texts))
# search and replace these values in each review
# treat periods and commas like words and strip off some characters
replacements = {'<br />': '', '"': '', '(': '( ',')': ' )', "'s ": " 's ",
'?': ' ? ', '-': ' ', ', ': ' , ', '. ': ' . ', '*': ''}
for i in range(0,len(texts)):
texts[i] = texts[i].lower()
texts[i] = reader.multi_replace(texts[i], replacements)
# parse review text into lists of words (delimited by " ")
word_lists = []
for text in texts:
word_list = text.split(' ')
if len(word_list) > 1:
word_lists.append(word_list)
print('Parsed', len(word_lists), 'reviews')
# -
# ## Format data for building a simple LSTM for classification
# ### - one that is able to predict whether the review sentiment is positive or negative
#
# - The single input is a list of word token ids
# - The words in the review were tokenized in the prepare_data
# - The label is a 1 for positive and 0 for negative
# - The model will output a floating point number between 0 and 1
# - Values >= .5 can be considered positive reviews
#
dfInput = pd.DataFrame()
dfInput['words'] = word_lists
dfInput['label'] = labels
data = ModelData(dfInput)
data.category_columns = ['words'] # indicates the contents are categories, not numeric values
data.sequence_columns = ['words'] # indicates the column contains a list of category values
data.label_column = 'label'
data.sequence_length = 1000 # almost all reviews are < 1000 words
data.validation_split = .10
data.prepare_data()
data.split_data(shuffle=True)
# ## Define Keras Model
#
# Learner object will choose LSTM/Dropout layer sets for the sequential inputs
# build model
learner = Learner(data)
learner.hidden_layers = 2 # number of lstm/dropout layer pairs
learner.dropout_rate = .30
learner.batch_size = 256
learner.lstm_units = 256
learner.gpu = True
learner.build_model()
learner.train_model(filename='imdb_simple', epochs=3)
learner.train_model(filename='imdb_simple', learning_rate=.0001, epochs=1)
| Imdb - Simple LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# # Exploratory Data Analysis and Visualization
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Exploratory Data Analysis
#
# - <NAME> (1915-2000) was a mathematician
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Fast Fourier Transform (Cooley-Tukey algorithm)
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Coined the phrase Exploratory Data Analysis (EDA)
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Exploratory Data Analysis
#
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Exploratory Data Analysis
#
# > 'Exploratory data analysis' is an attitude, a state of flexibility, a willingness to look for those things that we believe are not there, as well as those we believe to be there. - Exploratory Data Analysis
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# > If we need a short suggestion of what exploratory data analysis is, I would suggest that: 1. it is an attitude, AND 2. a flexibility, AND 3. some graph paper (or transparencies, or both). - The collected works of <NAME>: Philosophy and principles of data analysis
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# ### EDA should involve lots of visualization
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Bad Visualization
#
# 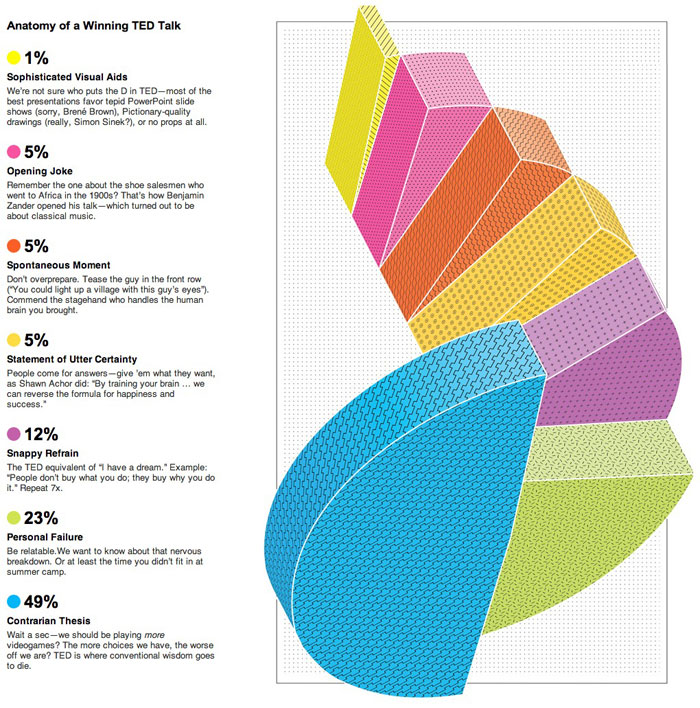
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# 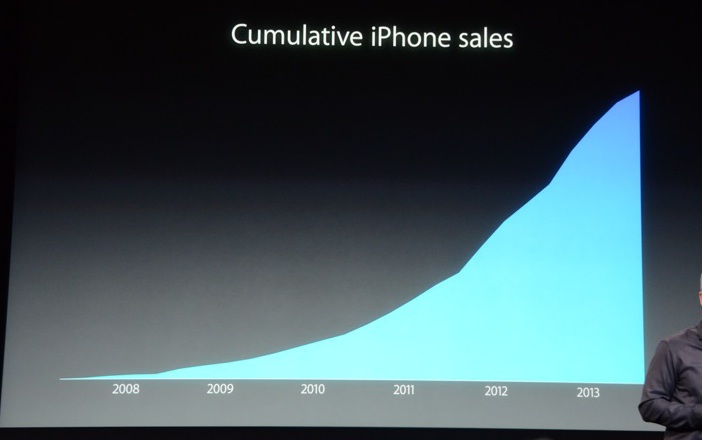
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# 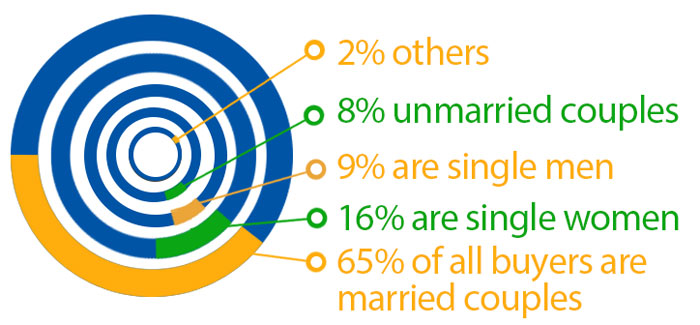
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# [Bad visualizations](https://www.google.com/search?q=bad+visualizations&source=lnms&tbm=isch&sa=X&ved=0ahUKEwiU2fG8qtnhAhXIna0KHXfsALkQ_AUIDigB&biw=1744&bih=1863)
#
# [WTF Visualizations](http://viz.wtf/)
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Good Visualization
#
# - [Gapminder Example](https://www.gapminder.org/tools/#$state$time$value=2018;&marker$axis_x$scaleType=linear;;;&chart-type=bubbles)
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Interactivity
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Good use of color, shapes, and sizes
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Clear labeling of components
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Axis (log vs. linear)
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Six categories of principles
#
# - scale
# - conditioning
# - perception
# - transformation
# - context
# - smoothing
#
# [Berkeley's DS-100 textbook](https://github.com/DS-100/textbook)
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Principles of Scale
#
# - Representative Chaffetz discussed in a congressional hearing [[full report](https://oversight.house.gov/interactivepage/plannedparenthood.)]
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Two numbers from Planned Parenthood programs:
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Number of abortions and cancer screenings
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - In reality the plot looked like this:
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
# HIDDEN
import warnings
# Ignore numpy dtype warnings. These warnings are caused by an interaction
# between numpy and Cython and can be safely ignored.
# Reference: https://stackoverflow.com/a/40846742
warnings.filterwarnings("ignore", message="numpy.dtype size changed")
warnings.filterwarnings("ignore", message="numpy.ufunc size changed")
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# %matplotlib inline
import ipywidgets as widgets
from ipywidgets import interact, interactive, fixed, interact_manual
import nbinteract as nbi
sns.set()
sns.set_context('talk')
np.set_printoptions(threshold=20, precision=2, suppress=True)
pd.options.display.max_rows = 7
pd.options.display.max_columns = 8
pd.set_option('precision', 2)
# This option stops scientific notation for pandas
# pd.set_option('display.float_format', '{:.2f}'.format)
pp = pd.read_csv("https://raw.githubusercontent.com/DS-100/textbook/master/notebooks/06/data/plannedparenthood.csv")
plt.plot(pp['year'], pp['screening'], linestyle="solid", marker="o", label='Cancer')
plt.plot(pp['year'], pp['abortion'], linestyle="solid", marker="o", label='Abortion')
plt.title('Planned Parenthood Procedures')
plt.xlabel("Year")
plt.ylabel("Service")
plt.xticks([2006, 2013])
plt.legend();
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Zoom to appropriate scale
#
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Principles of Conditioning
#
# - The US Bureau of Labor Statistics oversees scientific surveys related to the economic health of the US
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Following compares median weekly earnings split by sex: [US Bureau of Labor Statistics](https://www.bls.gov/data/)
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Comparison between Men and Women are easier (wage gap)
#
# - Lines are usually appropriate for ordinal and numerical data
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
cps = pd.read_csv("data/edInc2.csv")
ax = sns.pointplot(x="educ", y="income", hue="gender", data=cps)
ticks = ["<HS", "HS", "<BA", "BA", ">BA"]
ax.set_xticklabels(ticks)
ax.set_xlabel("Education")
ax.set_ylabel("Income")
ax.set_title("2014 Median Weekly Earnings\nFull-Time Workers over 25 years old");
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Principles of Perception
#
# ### Color perception
#
# - Humans perceive colors differently
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# 
#
# [Web color 787877](https://www.color-hex.com/color/787877)
# [Checker Shadow illusion](https://en.wikipedia.org/wiki/Checker_shadow_illusion)
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Not good!
# 
#
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Qualitative colors:
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
sns.palplot(sns.color_palette())
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Circular colors:
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
sns.palplot(sns.color_palette("husl", 8))
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Color Brewer interactive color chooser:
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
temp = sns.choose_colorbrewer_palette("sequential")
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
sns.palplot(sns.color_palette(temp))
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Color pallette from color names: e.g., [xkcd colors](https://xkcd.com/color/rgb/)
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
colors = ["puke green", "pistachio", "bubblegum pink", "mud brown", "ugly yellow", "dusty purple"]
sns.palplot(sns.xkcd_palette(colors))
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Some sequential colors can be used as `colormaps`: e.g., `cubehelix_palette()`
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
x, y = np.random.multivariate_normal([0, 0], [[1, -.5], [-.5, 1]], size=300).T
cmap = sns.cubehelix_palette(light=1, as_cmap=True)
sns.kdeplot(x, y, cmap=cmap, shade=True);
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Diverging colors:
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
sns.palplot(sns.color_palette("coolwarm", 7))
# -
# - [NYTimes Middle-class Jobs visualization](https://www.nytimes.com/interactive/2015/02/23/business/economy/the-changing-nature-of-middle-class-jobs.html)
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
def sinplot(flip=1):
x = np.linspace(0, 14, 100)
for i in range(1, 7):
plt.plot(x, np.sin(x + i * .5) * (7 - i) * flip)
sns.set_palette("husl")
sinplot()
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
with sns.color_palette("PuBuGn_d"):
sinplot()
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ### Area vs Length perception
#
# - Human's perception of lengths is better than areas
#
# - South Africa is about twice that of Algeria
#
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - South Africa is about twice that of Algeria
#
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ### Moving target
#
# - Human eyes do not detect changing baseline well
#
# - Carbon dioxide emissions over time split by country.
#
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Does UK's emissions have increased or decreased over time?
#
# - Changing baseline makes it difficult to see
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# 
# + hideCode=false hidePrompt=true
co2 = pd.read_csv("data/CAITcountryCO2.csv", skiprows = 2,
names = ["Country", "Year", "CO2"])
last_year = co2.Year.iloc[-1]
q = f"Country != 'World' and Country != 'European Union (15)' and Year == {last_year}"
top14_lasty = co2.query(q).sort_values('CO2', ascending=False).iloc[:14]
top14 = co2[co2.Country.isin(top14_lasty.Country) & (co2.Year >= 1950)]
from cycler import cycler
linestyles = (['-', '--', ':', '-.']*3)[:7]
colors = sns.color_palette('colorblind')[:4]
lines_c = cycler('linestyle', linestyles)
color_c = cycler('color', colors)
fig, ax = plt.subplots(figsize=(9, 9))
ax.set_prop_cycle(lines_c * color_c)
x, y ='Year', 'CO2'
for name, df in top14.groupby('Country'):
ax.semilogy(df[x], df[y], label=name)
ax.set_xlabel(x)
ax.set_ylabel(y + "Emissions [Million Tons]")
ax.legend(ncol=2, frameon=True);
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ## Principles of Transformation
#
# - Data transformation reveal non-linear relationships between variables
#
# - Distribution of ticket fares on Titanic is right-skewed (right-tailed)
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
sns.set_palette(sns.color_palette("muted"))
ti = sns.load_dataset('titanic')
sns.distplot(ti['fare'])
plt.title('Fares for Titanic Passengers')
plt.xlabel('Fare in USD')
plt.ylabel('Density');
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
sns.distplot(np.log(ti.loc[ti['fare'] > 0, 'fare']), bins=25)
plt.title('log(Fares) for Titanic Passengers')
plt.xlabel('log(Fare) in USD')
plt.ylabel('Density');
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Kepler recorded planets distances to the sun and their periods of orbit
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Basis for Third Law of Planetary Motion
# + hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
planets = pd.read_csv("data/planets.data", delim_whitespace=True, comment="#", usecols=[0, 1, 2])
planets
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
sns.lmplot(x='mean_dist', y='period', data=planets, ci=False);
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
sns.lmplot(x='mean_dist', y='period',
data=np.log(planets.iloc[:, [1, 2]]),
ci=False);
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# $$
# \begin{aligned}
# \log(period) &= m \log(dist) + b \\
# period &= e^{m \log(dist) + b} & \text{Taking the exponent of both sides} \\
# period &= e^b dist^m \\
# period &= C \cdot dist^m
# \end{aligned}
# $$
#
# - Constant $ C = e^b $
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Two variables have a polynomial relationship, so log of the variables are linearly related
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Degree of the polynomial is slope of the line
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Slope is 1.5 which gives us Kepler's third law: $ period \propto dist^{1.5} $
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Similarly, if $ \log(y) $ and $ x $ have linear dependence, $x$ and $y$ have exponential relationship: $ y = a^x $
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ### Principles of Context
#
# - Add as much relevant context as possible
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Following shows provides little context of what is being plotted
#
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Plots can be self-explanatory
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Add title, caption, axes labels, units for the axes, and labels for the plotted lines
#
# 
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ([This blog post](https://www.dataquest.io/blog/making-538-plots/) explains how to make these modifications using `matplotlib`.)
#
# In general, we provide context for a plot through:
#
# - Plot title
# - Axes labels
# - Reference lines and markers for important values
# - Labels for interesting points and unusual observations
# - Captions that describe the data and its important features
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ### Principles of Smoothing
#
# - Smoothing can help when there are many data points
#
# - Histograms are a type of smoothing for rugplots: e.g. age of the passengers in the Titanic.
# + hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
ages = ti['age'].dropna()
sns.rugplot(ages, height=0.2);
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# - Histogram is helpful and _kernel density estimation_ (KDE) can be useful
# + hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
sns.distplot(ages, kde=True); # defaults to adding smoothing
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ### Gaussian Kernel Smoothing
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
points = np.array([2, 3, 5])
sns.rugplot(points, height=0.2)
plt.xlim(0, 7);
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
from scipy.stats import norm
def gaussians(points, scale=True, sd=0.5):
x_vals = [np.linspace(point - 2, point + 2, 100) for point in points]
y_vals = [norm.pdf(xs, loc=point, scale=sd) for xs, point in zip(x_vals, points)]
if scale:
y_vals = [ys / len(points) for ys in y_vals]
return zip(x_vals, y_vals)
for xs, ys in gaussians(points, scale=True):
plt.plot(xs, ys, c=sns.color_palette()[0])
sns.rugplot(points, height=0.2)
plt.xlim(0, 7)
plt.ylim(0, 1);
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# HIDDEN
sns.rugplot(points, height=0.2)
sns.kdeplot(points, bw=0.5)
plt.xlim(0, 7)
plt.ylim(0, 1);
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
sns.kdeplot(ages)
sns.rugplot(ages);
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
sns.kdeplot(ages, bw=0.5) # narrower Gaussian yields less smoothing
sns.rugplot(ages);
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ### Cherry Blossom Run data
#
# - Annual 10-mile run in Washington D.C.
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Each runner can report their age and their race time
# + hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
runners = pd.read_csv('data/cherryBlossomMen.csv').dropna()
runners
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
sns.lmplot(x='age', y='time', data=runners, fit_reg=False);
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Many points make it difficult to see any trend
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Two dimensional kernel smoothing!
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# ### Two dimensional kernel density estimation
# + hideCode=true hidePrompt=true slideshow={"slide_type": "-"}
# Plot three points
two_d_points = pd.DataFrame({'x': [1, 3, 4], 'y': [4, 3, 1]})
sns.lmplot(x='x', y='y', data=two_d_points, fit_reg=False)
plt.xlim(-2, 7)
plt.ylim(-2, 7);
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# Place a Gaussian at each point and use a contour plot to show each one
sns.kdeplot(two_d_points['x'], two_d_points['y'], bw=0.4)
plt.xlim(-2, 7)
plt.ylim(-2, 7);
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
# Place a Gaussian at each point and use a contour plot to show each one
sns.kdeplot(two_d_points['x'], two_d_points['y'])
plt.xlim(-2, 7)
plt.ylim(-2, 7);
# + hideCode=true hidePrompt=true slideshow={"slide_type": "slide"}
sns.kdeplot(runners['age'], runners['time'])
plt.xlim(-10, 70)
plt.ylim(3000, 8000);
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Most runners were between 25 and 50
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Most ran between 1 to 2 hours (4000-7000 seconds)
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Slight up-trend as age increases?
# + [markdown] hideCode=true hidePrompt=true slideshow={"slide_type": "fragment"}
# - Strange group of age between 0 and 10 years old
# + [markdown] slideshow={"slide_type": "slide"}
# ### Seaborn visualization documentaion
#
# - Relational: [API](https://seaborn.pydata.org/api.html#relational-api) / [Tutorial](https://seaborn.pydata.org/tutorial/relational.html#relational-tutorial)
# - Categorical: [API](https://seaborn.pydata.org/api.html#categorical-api) / [Tutorial](https://seaborn.pydata.org/tutorial/categorical.html#categorical-tutorial)
# - Distributions: [API](https://seaborn.pydata.org/api.html#distribution-api) / [Tutorial](https://seaborn.pydata.org/tutorial/distributions.html#distribution-tutorial)
# - Regressions: [API](https://seaborn.pydata.org/api.html#regression-api) / [Tutorial](https://seaborn.pydata.org/tutorial/regression.html#regression-tutorial)
# - Multiples: [API](https://seaborn.pydata.org/api.html#grid-api) / [Tutorial](https://seaborn.pydata.org/tutorial/axis_grids.html#grid-tutorial)
# - Style: [API](https://seaborn.pydata.org/api.html#style-api) / [Tutorial](https://seaborn.pydata.org/tutorial/aesthetics.html#aesthetics-tutorial)
# - Color: [API](https://seaborn.pydata.org/api.html#palette-api) / [Tutorial](https://seaborn.pydata.org/tutorial/color_palettes.html#palette-tutorial)
| lecture-notes/06-Exploratory-Data-Analysis-and-Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Anomaly detection with an Autoencoder
# +
# %matplotlib inline
import tensorflow as tf
from tensorflow.keras.layers.experimental import preprocessing
import seaborn as sns
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import datasets, metrics, model_selection
# +
digits = datasets.load_digits()
fig, axes = plt.subplots(nrows=1, ncols=10, figsize=(10, 3))
for ax, image, label in zip(axes, digits.images, digits.target):
ax.set_axis_off()
ax.imshow(image, cmap=plt.cm.gray_r)
ax.set_title('%i' % label)
# +
target = digits.target
data = digits.images
print("min value: {}".format(np.amin(data)))
print("max value: {}".format(np.amax(data)))
print("shape: {}".format(np.shape(data)))
# +
X_train, X_test, y_train, y_test = model_selection.train_test_split(
data, target, test_size=0.5)
X_train = X_train.astype('float32') / 16.
X_test = X_test.astype('float32') / 16.
df_train = pd.DataFrame(y_train, columns=['target'])
df_train['type'] = 'train'
df_test = pd.DataFrame(y_test, columns=['target'])
df_test['type'] = 'test'
df_set = df_train.append(df_test)
_ = sns.countplot(x='target', hue='type', data=df_set)
print('train samples:', len(X_train))
print('test samples', len(X_test))
# +
class Autoencoder(tf.keras.models.Model):
def __init__(self):
super(Autoencoder, self).__init__()
self.encoder = tf.keras.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(16, activation='relu'),
tf.keras.layers.Dense(8, activation='relu'),
])
self.decoder = tf.keras.Sequential([
tf.keras.layers.Dense(16, activation='relu'),
tf.keras.layers.Dense(64, activation='sigmoid'),
tf.keras.layers.Reshape((8, 8))
])
def call(self, x):
encoded = self.encoder(x)
decoded = self.decoder(encoded)
return decoded
autoencoder = Autoencoder()
autoencoder.compile(optimizer='adam', loss='mse')
# -
# %%time
history = autoencoder.fit(X_train, X_train,
epochs=100,
validation_split = 0.2,
validation_data=(X_test, X_test),
verbose=0)
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
# +
def plot_loss(history):
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label='val_loss')
plt.xlabel('Epoch')
plt.ylabel('Error')
plt.legend()
plt.grid(True)
plot_loss(history)
# +
reconstructions = autoencoder.predict(digits.images)
fig, axes = plt.subplots(nrows=1, ncols=10, figsize=(10, 3))
for ax, image, label in zip(axes, reconstructions, digits.target):
ax.set_axis_off()
ax.imshow(image, cmap=plt.cm.gray_r)
ax.set_title('%i' % label)
# +
reconstruction_error_train = np.mean(tf.keras.losses.mae(autoencoder.predict(X_train), X_train), axis=-1)
reconstruction_error_test = np.mean(tf.keras.losses.mae(autoencoder.predict(X_test), X_test), axis=-1)
df_train = pd.DataFrame(reconstruction_error_train, columns=['reconstruction_error'])
df_train['type'] = 'train'
df_test = pd.DataFrame(reconstruction_error_test, columns=['reconstruction_error'])
df_test['type'] = 'test'
df_set = df_train.append(df_test)
fig, axs = plt.subplots(nrows=2, figsize=(10, 5))
fig.suptitle('Reconstruction error', fontsize=16)
p_threshold = 99
threshold = np.percentile(reconstruction_error_test, p_threshold)
x_max = np.max(reconstruction_error_test) + np.std(reconstruction_error_test)
axs[0].axvline(threshold, color='r', ls='--')
axs[0].set(xlim=(0, x_max))
axs[0].text(0.85, 0.2, 'threshold {:.3f}\n(percentile: {})'.format(threshold, p_threshold),
horizontalalignment='left', verticalalignment='center', transform=axs[0].transAxes)
axs[1].axvline(threshold, color='r', ls='--')
axs[1].set(xlim=(0, x_max))
_ = sns.kdeplot(data=df_set, x='reconstruction_error' ,hue='type', ax=axs[0])
_ = sns.boxplot(data=df_set, x='reconstruction_error', y='type', orient='h', ax=axs[1])
# +
anomalies_index = np.argwhere(reconstruction_error_test > threshold).flatten()
anomalies_x = np.array(X_test)[anomalies_index]
anomalies_y = np.array(y_test)[anomalies_index]
fig, axes = plt.subplots(nrows=1, ncols=len(anomalies_x), figsize=(10, 3))
fig.suptitle('Samples with reconstruction error > {:.3f} (percentile: {})'.format(threshold, p_threshold), fontsize=16)
for ax, image, label, in zip(axes, anomalies_x, anomalies_y):
ax.set_axis_off()
ax.imshow(image, cmap=plt.cm.gray_r)
ax.set_title('%i' % label)
# -
_ = sns.countplot(x=anomalies_y).set_title('Reconstruction error by target')
# +
flipped_images = np.array([np.transpose(x) for x in digits.images[0:10]])
flipped_images = flipped_images / 16.
flipped_images
reconstruction_error_flipped_images = np.mean(tf.keras.losses.mae(autoencoder.predict(flipped_images), flipped_images), axis=-1)
is_anomaly = reconstruction_error_flipped_images > threshold
# -
fig, axes = plt.subplots(nrows=1, ncols=len(flipped_images), figsize=(10, 2))
fig.suptitle('Flipped images'.format(threshold, p_threshold), fontsize=16)
for ax, image, anomaly in zip(axes, flipped_images, is_anomaly):
ax.set_axis_off()
ax.imshow(image, cmap=plt.cm.gray_r)
if anomaly:
ax.set_title('anomaly')
pd.DataFrame(reconstruction_error_flipped_images, columns=['reconstruction_error'])
| notebooks/unsupervised/neural_net/anomaly_detection_with_autoencoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
"""
Necessary Libraries
"""
import cv2
import numpy as np
# +
"""
Camera Selection
"""
cam_01=False; cam_02=True; cam_03=True; cam_04=True; cam_05=True
FPS = 10;
TestMode=False;
# -
"""
Video Parameters for Each Camera
"""
if cam_01:
if TestMode:
cap_01= cv2.VideoCapture(0)
else:
cap_01= cv2.VideoCapture('rtsp://service:core_PTZcam-123!@192.168.3.101/1')
width_01= int(cap_01.get(cv2.CAP_PROP_FRAME_WIDTH))
height_01= int(cap_01.get(cv2.CAP_PROP_FRAME_HEIGHT))
if cam_02:
cap_02= cv2.VideoCapture('rtsp://service:core_PTZcam-123!@192.168.3.102/1')
width_02= int(cap_02.get(cv2.CAP_PROP_FRAME_WIDTH))
height_02= int(cap_02.get(cv2.CAP_PROP_FRAME_HEIGHT))
if cam_03:
cap_03= cv2.VideoCapture('rtsp://service:core_PTZcam-123!@192.168.3.103/1')
width_03= int(cap_03.get(cv2.CAP_PROP_FRAME_WIDTH))
height_03= int(cap_03.get(cv2.CAP_PROP_FRAME_HEIGHT))
if cam_04:
cap_04= cv2.VideoCapture('rtsp://service:core_PTZcam-123!@192.168.3.104/1')
width_04= int(cap_04.get(cv2.CAP_PROP_FRAME_WIDTH))
height_04= int(cap_04.get(cv2.CAP_PROP_FRAME_HEIGHT))
if cam_05:
cap_05= cv2.VideoCapture('rtsp://service:core_PTZcam-123!@192.168.3.105/1')
width_05= int(cap_05.get(cv2.CAP_PROP_FRAME_WIDTH))
height_05= int(cap_05.get(cv2.CAP_PROP_FRAME_HEIGHT))
# +
if cam_01:
writer_01= cv2.VideoWriter('camera_01_1.mkv', cv2.VideoWriter_fourcc(*'DIVX'), FPS, (width_01,height_01))
if cam_02:
writer_02= cv2.VideoWriter('camera_02_2.mkv', cv2.VideoWriter_fourcc(*'DIVX'), FPS, (width_02,height_02))
if cam_03:
writer_03= cv2.VideoWriter('camera_03_3.mkv', cv2.VideoWriter_fourcc(*'DIVX'), FPS, (width_03,height_03))
if cam_04:
writer_04= cv2.VideoWriter('camera_04_4.mkv', cv2.VideoWriter_fourcc(*'DIVX'), FPS, (width_04,height_04))
if cam_05:
writer_05= cv2.VideoWriter('camera_05_5.mkv', cv2.VideoWriter_fourcc(*'DIVX'), FPS, (width_05,height_05))
while True:
if cam_01:
ret_01,frame_01= cap_01.read()
writer_01.write(frame_01)
cv2.imshow('Camera-01',frame_01)
if cam_02:
ret_02,frame_02= cap_02.read()
writer_02.write(frame_02)
cv2.imshow('Camera-02',frame_02)
if cam_03:
ret_03,frame_03= cap_03.read()
writer_03.write(frame_03)
cv2.imshow('Camera-03',frame_03)
if cam_04:
ret_04,frame_04= cap_04.read()
writer_04.write(frame_04)
cv2.imshow('Camera-04',frame_04)
if cam_05:
ret_05,frame_05= cap_05.read()
writer_05.write(frame_05)
cv2.imshow('Camera-05',frame_05)
if cv2.waitKey(1) & 0xFF == ord('q'):
break
if cam_01:
cap_01.release()
writer_01.release()
if cam_02:
cap_02.release()
writer_02.release()
if cam_03:
cap_03.release()
writer_03.release()
if cam_04:
cap_04.release()
writer_04.release()
if cam_05:
cap_05.release()
writer_05.release()
cv2.destroyAllWindows()
# -
| src/data_collection/Robofootball_Match_Record.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# ## Building an end-to-end ML Pipeline with AWS Sagemaker & API
#
# This Notebook shows a basic example how to build an end-to-end machine learing pipeline on AWS by using the [AWS Sagemaker Python SDK](https://sagemaker.readthedocs.io/en/stable/).
#
# The AWS Sagemaker Components provide a service for running your python scripts in docker containers, using either AWS maintained images or custom images. Additionally the Sagemaker Python SDK implements many convenience functions for handling parameters such as instance size, IO-Handling, and deployment. They are great building blocks for scalable, consistent, reproducable ml pipelines. They can easily be orchestarated by either using an open source Workflow Tool (Airlflow, Prefect) or AWS Step Functions. They provide a perfect fit for ordinary ml workflows with medium sized datasets, tabular data. They support images for the most widely used ml frameworks, eg. scikit-learn, tensorflow and pytorch.
#
# The Notebook contains both the source code for preprocessing, training and deployment, as well as the calls to the sagemaker API that are executing the jobs.
#
# ### The Pipeline
#
# ##### Data Processing:
# - The flow begins with a preprocessing script that uses `pandas` and `scikit-learn` to read a csv, apply transformations to the data, splits the data into train and test set, and saves the data to S3.
# - The preprocessing file will be executed with the `SKLearnProcessor`, where instance size and IO paths will be configured.
#
# ##### Model Training & Deployment:
# - Next, another script for model training and deployment will be created. This script includes the algorithm, the training rutine, the serialization of the model, and the serving functions that will be used for model deployment.
# - This script will be executed with the `SKLearn` estimator class. When calling `fit()` on it, model training will be executed. When calling `deploy()`, the model will be deployed.
#
# ##### Model Serving:
# - A lmabda function is created as an intermediate layer between your sagemaker model endpoint and your REST API.
# - A REST API is configured with API Gateway. It consists of a simple `POST` method that calls the lambda function with live data as payload. The live data is passed to the enpoint and predictions are receid and returned to the caller.
#
# <img src="img/flowchart_ml_pipeline.png" alt="Flowchart" width="1200" height="675" style="horizontal-align:middle">
# -
# ### Prerequisites
# To run this demo, you will need access to an AWS account, a user that let's you access the ressource needed, and roles to grand permessions for the services. This demo will not cover how to set up IAM roles and permissions.
#
# To run this demo, you will need access to an AWS account, create a user with a policy that grants permissions to all services that will be used in this example. The notbook can be run on any environment, given that authentication is provided, however the recommended, and easiest way is to run this tutorial on an AWS sagemakeer notebook instance. You can find more information about setting that up [here](https://docs.aws.amazon.com/sagemaker/latest/dg/howitworks-create-ws.html).
# #### Load Environment Variables
#
# I am using [dotenv](https://github.com/theskumar/python-dotenv) to handle my environment variables. You could either directly define them in the notebook below, replacing the `os.getenv()` call (e.g. `script_path = "path/to/your/script"`), or you define them in an `.env` file in your root directory.
import os
# +
# %load_ext dotenv
# %dotenv
sagemaker_role= os.getenv("SAGEMAKER_ROLE") # Sagemaker Role TODO: Define Separate Roles for SageMaker, Lambda and Gateway
raw_data_bucket=os.getenv("RAW_DATA_PATH")
processed_data_bucket=os.getenv("PREPROCESSING_OUTPUT_PATH")
# +
# Environment variables
#sagemaker_role = "XXXXXXXXXXXXXXXX"
#lambda_role = "XXXXXXXXXXXXXXXX"
#raw_data_path="XXXXXXXXXXXXXXXX"
#preprocessing_output_path="XXXXXXXXXXXXXXXX"
# -
# ## Get Data
#
# In this example we will be using the Plamer Penguin Dataset, which provides a good alternative to the frequently used Iris dataset. It contains information about various penguins. You can read more about it [here](https://allisonhorst.github.io/palmerpenguins/articles/intro.html).
#
# The objective we will be solving with our machine learning algorithm is to predict the gender of a penguin by using all other columns as features.
from palmerpenguins import load_penguins
import pandas as pd
# !pip install palmerpenguins
penguins = load_penguins()
penguins.head(3)
# Write csv to raw data s3 bucket
file_path = os.path.join(raw_data_bucket, "penguins.csv")
penguins.to_csv(file_path)
print(f"Stored raw data in '{file_path}'.")
# ## Preprocessing
# Load Sagemaker classes
from sagemaker.sklearn.processing import SKLearnProcessor
from sagemaker.sklearn.estimator import SKLearn
from sagemaker.sklearn import SKLearnModel
from sagemaker.processing import ProcessingInput, ProcessingOutput
# #### Develop preprocessing script
#
# This is an example preprocessing script. It will read the data into a Pandas DataFrame, and apply a scikit-learn column transformer pipeline, one-hot-encoding categorical variables and scaling interval-scaled variables. The it will split the data into training and test set and write the data to a flat file.
#
# When executing the cell, the magic command `%%writefile filename.py` will save the file the code as a python file in your current working directory. This will allow the SageMaker preprocessing job to use the script in a seperate docker container, where the preprocessing will be executed.
# +
# %%writefile preprocessing.py
import argparse
import os
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder, StandardScaler
from sklearn.compose import make_column_transformer
features = [
"bill_length_mm",
"bill_depth_mm",
"flipper_length_mm",
"species",
"island",
]
target = "sex"
columns = features + [target]
if __name__ == "__main__":
# Parse Arguments
parser = argparse.ArgumentParser()
parser.add_argument("--train-test-split", type=float, default=0.3)
args, _ = parser.parse_known_args()
split = args.train_test_split
print("Arguments {}".format(args))
# Process input data
input_data_path = os.path.join("/opt/ml/processing/input", "penguins.csv")
df = pd.read_csv(input_data_path)
df = pd.DataFrame(data=df, columns=columns)
df.dropna(inplace=True)
df.drop_duplicates(inplace=True)
# Create sklearn preprocessing pipeline
preprocess_pipeline = make_column_transformer(
(["bill_length_mm", "bill_depth_mm", "flipper_length_mm"], StandardScaler()),
(["species", "island"], OneHotEncoder(sparse=False)),
)
# Apply Pipeline
X = preprocess_pipeline.fit_transform(df.drop(columns=target))
# Split data into training and test set
X_train, X_test, y_train, y_test = train_test_split(
pd.DataFrame(X),
df[target],
test_size=split,
random_state=42,
)
train_features_output_path: str = os.path.join(
"/opt/ml/processing/train", "train_features.csv"
)
train_labels_output_path: str = os.path.join(
"/opt/ml/processing/train", "train_labels.csv"
)
test_features_output_path: str = os.path.join(
"/opt/ml/processing/test", "test_features.csv"
)
test_labels_output_path: str = os.path.join(
"/opt/ml/processing/test", "test_labels.csv"
)
# Save processed data as csv
print("Training features path {}".format(train_features_output_path))
X_train.to_csv(train_features_output_path, header=False, index=False)
print("Test features path {}".format(test_features_output_path))
X_test.to_csv(test_features_output_path, header=False, index=False)
print("Training labels path {}".format(train_labels_output_path))
y_train.to_csv(train_labels_output_path, header=False, index=False)
print("Test labels path {}".format(test_labels_output_path))
y_test.to_csv(test_labels_output_path, header=False, index=False)
# -
# #### Define & Run SKLearn Preprocessor
#
# The `SKLearnProcessor` lets you configure the preprocessing job, including the `framework_version`, the `instance_type` and the number of instances. You could also pass a custom docker image to the object that would be used instead of the scikit-learn image maintained by AWS.
#
# When calling `run()` the preprocessing job will be executed. The function accepts the path to the preprocessing script that was defined in the cell above as input. Additionally, data input and output paths will be defined. S3 buckets can be used for retrieval of raw data and storing of proceeesed data. With the objects `ProccesingInput` and `ProcessingOutput` we make sure that the paths in S3 and in our docker container are mapped accordingly.
# +
sklearn_processor = SKLearnProcessor(
framework_version="0.20.0",
base_job_name="preprocessing",
role=sagemaker_role,
instance_type="ml.m5.xlarge",
instance_count=1,
)
docker_base_path: str = "/opt/ml/processing/"
sklearn_processor.run(
code="preprocessing.py",
inputs=[
ProcessingInput(
source=raw_data_bucket,
destination=os.path.join(docker_base_path, "input")
),
],
outputs=[
ProcessingOutput(
destination=processed_data_bucket,
output_name="train_data",
source=os.path.join(docker_base_path, "train")
),
ProcessingOutput(
destination=processed_data_bucket,
output_name="test_data",
source=os.path.join(docker_base_path, "test")
),
],
)
preprocessing_job_description = sklearn_processor.jobs[-1].describe()
# -
# #### Inspect generated training data
#
# Let's have a look at our processed data.
training_features = pd.read_csv(processed_data_bucket + "train_features.csv", nrows=10, header=None)
print("Training features shape: {}".format(training_features.shape))
training_features.head(n=3)
# ## Model Training
# + [markdown] tags=[]
# #### Create SKLearn training and deploy script
#
# In order to execetue model training and deployment of the trained model, we need to write another script.
#
# The script will comprise of the training routine, which will ingest the processed training data that was generated in the Sagemaker Processing step above. It reads the data, instanciates the model - here a simple `LogisticRegression` and calls `fit` on the model. The model is then serialized and saved in our working directory. The `SKLearn` object will then move the artifacts to the desired output path in S3. If no output path is specified, Sagemaker will create a new bucket to store the artifacts of the training job.
#
# The script also contasins several serving functions that Sagemaker requires for model serving via the sagemaker model endpoint service. These functions comprise of `model_fn()` ensuring that the model gets loaded from file, `input_fn()` handling the input in a way that it can be used for calling the `predict()` function on the model, the `predict_fn()` which actually calls `predict` on the model and the `output_fn()`, which will convert the model output to a format that can be send back to the caller.
#
# The script will also be saved to disc with the `%%writefile` magic command.
# +
# %%writefile train_and_deploy.py
import os
import pandas as pd
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.externals import joblib
"""
Define model serving functions. More aboutthese functions at:
https://sagemaker.readthedocs.io/en/stable/frameworks/sklearn/using_sklearn.html#load-a-model
"""
def model_fn(model_dir):
model = joblib.load(os.path.join(model_dir, "model.joblib"))
return model
def input_fn(request_body, content_type):
if content_type == 'text/csv':
samples = []
for r in request_body.split('|'):
samples.append(list(map(float,r.split(','))))
return np.array(samples)
else:
raise ValueError("Thie model only supports text/csv input")
def predict_fn(input_data, model):
return model.predict(input_data)
def output_fn(prediction, content_type):
return str(prediction)
if __name__ == "__main__":
training_data_directory = "/opt/ml/input/data/train"
train_features_data = os.path.join(training_data_directory, "train_features.csv")
train_labels_data = os.path.join(training_data_directory, "train_labels.csv")
X_train = pd.read_csv(train_features_data, header=None)
y_train = pd.read_csv(train_labels_data, header=None)
model = LogisticRegression(class_weight="balanced", solver="lbfgs")
model.fit(X_train, y_train)
model_output_directory = os.path.join("/opt/ml/model", "model.joblib")
print("Model saing path {}".format(model_output_directory))
joblib.dump(model, model_output_directory)
# -
# The `SKLearn` object is the standard interface for scheduling and defining model training and deployment of scikit-learn models. After specifying the ressources needed, the framework version and the entry_point, we can call `fit()` in order to execute the training job. We pass a dictionary with a single keyword `"train"` that specifies the path to the processed data in S3.
sklearn = SKLearn(
entry_point="train_and_deploy.py",
framework_version="0.20.0",
instance_type="ml.m5.xlarge",
role=sagemaker_role
)
sklearn.fit({"train": processed_data_bucket})
# #### Evaluate Model Performance
# Another script is created in order to evaluate the perfomance of the model created above. The evaluation step will again be executed as an individual step in our ml pipeline. It loads both the model and the processed test data, collects several metrics (classification report, roc_auc score, accuracy) and stores them in a JSON file.
# Because we did not specify a bucket, where our model artifact should be stored, the training job created a new one. The uri can be retrieved from the metadata contained in the `sklearn` object:
# Get model data in order to load model
model_data_s3_uri = sklearn.output_path + sklearn.latest_training_job.name + "/output/model.tar.gz"
model_data_s3_uri
#
# Execute model evaluation using the same processing configurations as for the preprocessing job and the same object instantiated above. Two inputs are specified, one for the model and another one for the test data.
# + [markdown] tags=[]
# #### Evaluate Model Performance
#
# Another script is created in order to evaluate the perfomance of the model created above. The evaluation step will again be executed as an individual step in our ml pipeline. It loads both the model and the processed test data, collects several metrics (classification report, roc_auc score, accuracy) and stores them in a JSON file.
# +
# %%writefile evaluate.py
import json
import os
import tarfile
import pandas as pd
from sklearn.externals import joblib
from sklearn.metrics import classification_report, accuracy_score
if __name__ == "__main__":
model_path = os.path.join("/opt/ml/processing/model", "model.tar.gz")
print("Extracting model from path: {}".format(model_path))
with tarfile.open(model_path) as tar:
tar.extractall(path=".")
print("Loading model")
model = joblib.load("model.joblib")
print("Loading test input data")
test_features_data = os.path.join("/opt/ml/processing/test", "test_features.csv")
test_labels_data = os.path.join("/opt/ml/processing/test", "test_labels.csv")
X_test = pd.read_csv(test_features_data, header=None)
y_test = pd.read_csv(test_labels_data, header=None)
predictions = model.predict(X_test)
report = classification_report(y_test, predictions, output_dict=True)
report["accuracy"] = accuracy_score(y_test, predictions)
print("Classification report:\n{}".format(report))
eval_output_path = os.path.join(
"/opt/ml/processing/evaluation", "evaluation.json"
)
print("Evaluation output path: {}".format(eval_output_path))
with open(eval_output_path, "w") as f:
f.write(json.dumps(report))
# -
# Execute model evaluation using the same processing configurations as for the preprocessing job and the same object instantiated above. Two inputs are specified, one for the model and another one for the test data.
# +
import json
from sagemaker.s3 import S3Downloader
sklearn_processor.run(
code="evaluate.py",
inputs=[
ProcessingInput(source=model_data_s3_uri, destination="/opt/ml/processing/model"),
ProcessingInput(source=processed_data_bucket, destination="/opt/ml/processing/test"),
],
outputs=[ProcessingOutput(output_name="evaluation", source="/opt/ml/processing/evaluation")],
)
evaluation_job_description = sklearn_processor.jobs[-1].describe()
# + [markdown] tags=[]
# #### Inspect Evaluation result
#
# The JSON file that was created in the evaluation job can now be read and inspected.
# +
import boto3
client = boto3.client('s3')
s3_path=evaluation_job_description["ProcessingOutputConfig"]["Outputs"][0]["S3Output"]["S3Uri"]
bucket, key = s3_path.split("//")[1].split("/",1)
result = client.get_object(Bucket=bucket, Key= key + '/evaluation.json')
json.loads(result['Body'].read().decode('utf-8'))
# + [markdown] tags=[]
# ## Model Deployment
# -
# #### Deploy Estimator to Sagemaker Endpoint
#
# After evaluating our model, we can now go on and deploy it. In order to do so, we only have to call `deploy()` on the `sklearn` object that we used for model training.
# #### Test Sagemaker Endpoint
# We can now run our first test against our model endpoint directly from our jupyter notebook. To do so, we can simply take some of the training features, add them to a request and then call our model by using the Sagemaker client with the `invoce_endpoint` method.
# +
# Load in two rows from the training data
training_data = training_features.head(2).values.tolist()
# Format the deploy_test data features
request_body = ""
for sample in training_data:
request_body += ",".join([str(n) for n in sample]) + "|"
request_body = request_body[:-1]
print("*"*20)
print(f"Calling Sagemaker Endopint with the following request_body: {request_body}")
# create sagemaker client using boto3
client = boto3.client('sagemaker-runtime')
# Specify endpoint and content_type
endpoint_name = predictor.endpoint
content_type = 'text/csv'
# Make call to endpoint
response = client.invoke_endpoint(
EndpointName=endpoint_name,
Body=request_body,
ContentType=content_type
)
response_from_endpoint = response['Body'].read().decode("utf-8")
print("*"*20)
print(f"Response from Endpoint: {response_from_endpoint}")
# -
# #### Delete Endpoint, if no longer in use
#
# Because your endpoint has incurring costs while in use, it is advisable to delete it as soon as it is no longer needed. If you follow this tutorial for testing purposes, make sure that your endpoint is deleted as soon as you stop working on it.
# +
# This call will delete the endpoint
# predictor.delete_endpoint()
# -
# Beware that directly calling the model endpoint should only be done for testing purposes. If you want to make your model available for live predictions, it is advisable to add a proper REST API that handles incoming requests. How this can be done, will be described in the next step.
# ## Build REST API
# #### Create Lambda Function for handling API <-> Sagemaker Endpoint traffic
#
# First we will write a lambda function for handling the traffic between our REST API and our model enpoint. It will be receiving requests from the API as input, invoke the model endpoint and return the results.
# +
# %%writefile serving_lambda.py
import os
import boto3
import json
endpoint_name = os.environ['ENDPOINT_NAME']
runtime= boto3.client('runtime.sagemaker')
def lambda_handler(event, context):
print("Received event: " + json.dumps(event, indent=2))
data = json.loads(json.dumps(event))
payload = json.loads(data['data'])
print(payload)
# Format the data so that it can be processed by the model endpoint
body = ""
for sample in payload:
body += ",".join([str(n) for n in sample]) + "|"
body = body[:-1]
print("request_body: ", body)
response = runtime.invoke_endpoint(EndpointName=endpoint_name,
ContentType='text/csv',
Body=body)
label = response['Body'].read().decode('utf-8').strip("[]").strip("'")
return label
# -
# Print out predictor endpoint and add string to lambda function as environment variable
predictor.endpoint
# #### Configure API Gateway
#
# This tutrial will walk you through setting up API Gateway via the management console. If you whish to run this in production, it is advisable to provision and configure this ressource with an infrastructure management tool, such as AWS Cloud Formation or Terraform.
#
# ##### Step I: Go to API Gateway & Select Create new REST Endpoint
#
# 
#
# ##### Step II: Choose a name and create a new API
#
# 
#
# ##### Step III: Create a new method of type POST and choose your lambda as target
#
# 
#
# ##### Step IV: Deploy API
#
# 
#
# ##### Step V: Go to APIs --> Stages --> Inspect your newly created stage and collect Invocation Endpoint
# Set invocation endpoint URL as environemnt variable `"API_URL"`
# #### Invoke Request against REST API
#
# After you have successfully created your REST API with API Gateway, you can now test it.
# +
import requests
import json
url = os.getenv("API_URL")
payload = json.dumps({"data":"[[-0.6396528091784842, 0.3738717119645826, -0.9980179785096928, 1.0, 0.0, 0.0, 1.0, 0.0, 0.0]]"})
print(f"Calling model REST API with the following payload {payload}")
response = requests.post(url, data=payload)
print("*"*20)
print(f"Return Message. Status code: {response.status_code}, Message: {response.text}")
# -
# ### Outro
#
# That's it. After following all steps, you should now have successfully created an end-to-end ml pipeline with AWS Sagemaker and configured a REST API that serves your predictions online. WOW!!!
| notebooks/ml_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Variables and Data Structures
# ### BIOINF 575 - Fall 2020
# ### Interactive python apps commands
#
#   python – open a plain python console in current terminal <br>
#   http://jupyter.org/ - The evolved form of iPython <br>
#   ipython or jupyter-console – open an iPython console in the current terminal <br>
#   jupyter-qtconsole – open an iPython session in a qt console (console uses qt GUI library, more functional window)
# <br>
#   jupyter notebook – start a notebook session and open in in a web browser <br>
#   Jupyter-lab – start a jupyter lab session, the future of the notebook <br>
#   spyder – launch the Spyder IDE <br>
#    https://www.spyder-ide.org/ <br>
#   anaconda-navigator – launch the Anaconda Navigator <br>
# https://www.dataquest.io/blog/jupyter-notebook-tutorial/
# A <b>markup language</b> is a system for annotating a document where the test does not read very well but the result makes the content easier to understand<br>
# Example: using color, different font styles, different font size, indentation, itemizing, numbering
# The idea started from the "marking up" of paper manuscripts.
#
# <b>Markdown</b> is a tool to convert text to HTML.<br>
# Markdown is lightweight markup language with plain text formatting syntax.<br>
# Its design allows it to be converted to many output formats, but the original tool by the same name only supports HTML.<br>
# Markdown is often used to format readme files, for writing messages in online discussion forums, and to create rich text using a plain text editor.<br><br>
# https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet<br>
# #### The Zen of Python, by <NAME>
import this
# ### Variables
# Naming conventions
# * underscore separated - variable_name - preference
# * Camel case - VariableName
#
# There are many others:
# * https://en.wikipedia.org/wiki/Naming_convention_(programming)
#
# Use meaningful names, your future self will thank you!
# <b>Assign</b> a value to a variable. <br>
# The assignment operator is: "=" <br><br>
message = "Let's get started!"
# Check the value of a variable
message
print(message)
# <font color = "red">EXERCISE</font> <br>
# <br>
# Assign the value 10 to a variable.<br>
# Assign the value True to a variable.<br><br>
# Assign the value 100 to the variable attendance_percentage.<br>
# Assign the value of the variable attendance_percentage to the variable quiz_percentage.<br><br>
# <b>Multiple assignment:</b> assign the value 20 to two variables at the same time.<br>
# Then check the value of the two variables.<br><br>
cat_weight = 10
print(cat_weight)
is_Wednesday = True
print(is_Wednesday)
attendance_percentage = 100
print(attendance_percentage)
quiz_percentage = attendance_percentage
print(quiz_percentage)
attendance_percentage = 90
print(attendance_percentage)
# Remove the value from a variable using <b>None</b>.
message
message = None
message
# Remove a variable from the environment using the function <b>del()</b>.
del(message)
# Now let's try to see what value variable <i>message</i> has<br>
# What do we expect?
message
# python complained!
# <img src='http://greenlining.org/wp-content/uploads/2013/10/COMPLAIN.jpg' width="200"/>
# http://greenlining.org/wp-content/uploads/2013/10/COMPLAIN.jpg
# List the type of a variable with the function <b>type()</b>.
message = "Let's test the type."
message
type(message)
message = 10
message
type(message)
# Typing is dynamic but strict in python (objects have type).<br><br>
# The principal built-in types are numerics, sequences, mappings, classes, instances and exceptions.<br>
# Numeric Types — int (subtype: bool), float, complex.<br>
# Sequence Types — string, list, tuple, range.<br><br>
# There are two major categories: mutable and immutable types.<br>
# <b>Immutable</b> variables <b>cannot be changed</b>, while <b>Mutable</b> variables <b>can be changed</b>.<br>
# Integer, Floats, Strings, Tuples, Boolean are immutable.<br>
# List are mutable types.<br>
#
# https://docs.python.org/3/library/stdtypes.html
#
# List functions and attributes in strings using the <b>dir(str)</b> function.
dir(str)
"test".capitalize()
# Funtions preceeded by \_\_ (e.g. \_\_lt\_\_ - less than) are special internal funtions used by python funtions and operators
# <font color = "red">EXERCISE</font> <br><br>
# Test more of the string functions: upper(), strip(), ...<br><br>
"test".upper()
" test text ".strip()
"test".index("es")
# Get more information regarding strings using the <b>help(str)</b> function.
help(str)
# ### Numeric
# Basic math operations: <b>int</b> (natural numbers), <b>float</b> (decimal numbers), <b>complex</b><br>
# Integer, arbitrarily long, convert from floats or strings with int()<br>
# Floats, convert from str or int types with float()<br>
# Convert to string with str()<br>
# <b>No automatic coercion of types, explicit conversion only</b><br>
# <b>Exception: print() - converts to strings where possible</b>
dir(int)
# Computing the area of a rectangle.
length = 2
width = 4
area = length * width
area
type(4)
# Float division
type(4/2)
type(4//2)
4//2
4/2
# Explicit conversion
float(4)
float("4.5")
float("DNA sequence")
int("4")
repr(4)
str(4)
# remainder/modulo
5%2
# exponent
5**2
message = "test"
number = 10
message * number
# ### Boolean
# Boolean is a subtype of integer<br>
# <b>True</b>, <b>False</b> values<br>
# 1 == True, 0 == False<br>
# empty structure == False<br>
type(True)
dir(bool)
cond_result = True
cond_result
cond_result1 = False
cond_result1
# Logical operators <br><br>
# <b>not</b> <br>
# <b>and</b> <br>
# <b>or</b> <br>
not cond_result
cond_result and cond_result1
cond_result or cond_result1
# Comparison operators <br><br>
# <b><</b> strictly less than<br>
# <b><=</b> less than or equal<br>
# <b>\></b> strictly greater than<br>
# <b>>=</b> greater than or equal<br>
# <b>==</b> equal<br>
# <b>!=</b> not equal<br>
# <b>is</b> object identity<br>
# <b>is not</b> negated object identity<br>
#
# not cond_result
cond_result = 2>=3
cond_result
int(True)
# <font color = "red">EXERCISE</font> <br><br>
# Assign a numeric value to a variable.<br>
# Write a composite condition to check if the variable is in the range 10 to 100.<br>
# Write a composite condition to check if the variable is lower than -10 or higher than 10.<br>
# Write a composite condition to check if the variable is lower than -10 or higher than 10 and if it can be divided by 5 or 7. <br>
class_size = 15
print(class_size > 10 and class_size < 100)
print(class_size < -10 or class_size > 10)
print(abs(class_size) > 10)
print((class_size < -10 or class_size > 10) and (class_size%5 == 0 or class_size%7 == 0))
# ### Sequence types: <b>String</b>, <b>List</b>, and <b>Tuple</b> - are iterable
# ### String
# Sequence of characters - immutable
# Concatenation
"Complex " + 'text'
# Simple, double or triple quotes:
"Don't worry about apostrophes"
'She said: "Good day!"'
test_message = """Trying the triple quotes"""
test_message
'''
I am testing the
triple quotes - a comment
'''
# this is a COMMENT
test_message = '''Trying the simple triple quotes'''
test_message
# <b>Common Sequence operations</b>
# <br>
#
# Operation Description:<br><br>
# <b>x in s</b>
# True if an item of s is equal to x, else False
#
# <b>x not in s</b>
# False if an item of s is equal to x, else True
#
# <b>s + t</b>
# the concatenation of s and t
#
# <b>s * n or n * s</b>
# equivalent to adding s to itself n times
#
# <b>s[i]</b>
# ith item of s, origin 0
#
# <b>s[i:j]</b>
# slice of s from i to j
#
# <b>s[i:j:k]</b>
# slice of s from i to j with step k
#
# <b>len(s)</b>
# length of s
#
# <b>min(s)</b>
# smallest item of s
#
# <b>max(s)</b>
# largest item of s
#
# <b>s.index(x[, i[, j]])</b>
# index of the first occurrence of x in s (at or after index i and before index j)
#
# <b>s.count(x)</b>
# total number of occurrences of x in s
"ll" in "collection"
"test this, string." in "Collection"
"rep"*4
"collection"[0]
"collection"[1:3]
"collection"[1:3]
"testing the subsetting"[3:16:3]
"testing the subsetting"[3:1000]
test_string = "testing the subsetting"
test_string[:len(test_string)]
len(test_string)
# #### Subsetting s[i:j:k]
#
# If i or j is negative, the index is relative to the end of sequence s: len(s) + i or len(s) + j is substituted. But note that -0 is still 0.
#
# The slice of s from i to j is defined as the sequence of items with index k such that i <= k < j. <br>
# If i or j is greater than len(s), use len(s). <br>
# If i is omitted or None, use 0. <br>
# If j is omitted or None, use len(s). <br>
# If i is greater than or equal to j, the slice is empty.<br>
#
# The slice of s from i to j with step k is defined as the sequence of items with index x = i + n\*k such that 0 <= n < (j-i)/k. <br>
# In other words, the indices are i, i+k, i+2\*k, i+3\*k and so on, stopping when j is reached (but never including j). <br>
# When k is positive, i and j are reduced to len(s) if they are greater. <br>
# When k is negative, i and j are reduced to len(s) - 1 if they are greater. <br>
# If i or j are omitted or None, they become “end” values (which end depends on the sign of k). <br>
# Note, k cannot be zero. <br>
# If k is None, it is treated like 1.<br>
#
# https://docs.python.org/3/library/stdtypes.html
#
len("collection")
"collection".index("ll")
min("collection")
"A" < "a"
ord("a")
ord("A")
max("collecTion")
"collection".count("l")
cooper = 20
Cooper
s = "collection"
s
s.index("e")
# <font color = "red">Exercise</font> <br><br>
# Create the sequence "blablabla" by using the operator *. <br>
# Find the position of the second o in "collection".<br>
# Retrieve the word "other" from "immunotherapy" using subsetting.<br><br>
"bla"*3
s = "another test"
subs = "t"
s.index(subs, s.index(subs)+1)
# <b>String formatting</b> - much more in a future session.
#
#
# Python uses C-style string formatting to create new, formatted strings. The "%" operator is used to format a set of variables enclosed in a "tuple" (a fixed size list), together with a format string, which contains normal text together with "argument specifiers", special symbols like "%s" and "%d".
#
#
# # %s - String (or any object with a string representation, like numbers) <br>
# # %d - Integers<br>
# # %f - Floating point numbers<br>
# %.<n>f - Floating point numbers with a fixed (n) amount of digits to the right of the dot.
#
#
name = "John"
print("Hello, %s!" % name)
# ### Range - an immutable sequence of numbers
# It is commonly used for looping a specific number of times.
# range(stop)<br>
# range(start, stop[, step])
dir(range)
# +
list(range(10))
# -
range(10)
list(range(5, 31, 5))
# ### List - a collection of elements, allows duplicates, is unorderred, and is mutable (changeable)
# A list may be constructed in several ways:<br>
#
# Using a pair of square brackets to denote the empty list: []<br>
# Using square brackets, separating items with commas: [a], [a, b, c]<br>
# Using a list comprehension: [x for x in iterable]<br>
# Using the type constructor: list() or list(iterable)<br>
dir(list)
patient_BMIs = [25, 18, 30, 16, 22, 32, 28]
patient_BMIs
# Size/length of a list
len(patient_BMIs)
# Save the list in a different variable
patient_BMIs_followup = patient_BMIs
patient_BMIs_followup
# Retrieve elements from a list subsetting/slicing
patient_BMIs_followup[2:4]
# Negative indexing - retrieving elements from the end of the list
patient_BMIs[-4:-1]
# Change a value in a list
patient_BMIs_followup[1] = 20
patient_BMIs_followup
patient_BMIs
# patient_BMIs_followup and patient_BMIs are references to the same object<br>
# <b>When the referred object changes the values display for both referring variables change.</b>
# To make a copy of a list:<br>
# copy_list = initial_list[:] <br><br>
# Similarly we can do <br>
# copy_list = list(initial_list)
patient_BMIs_followup = patient_BMIs[:]
# patient_BMIs_followup = list(patient_BMIs)
patient_BMIs_followup[2] = 27
patient_BMIs_followup
patient_BMIs
# <img src="http://henry.precheur.org/python/list1.png" width=200 />
# <img src="http://henry.precheur.org/python/list2.png" width=200 />
# To copy a list through subsetting works ... until it doesn't.<br>
# When the list contains sublists.
# The copy module has a deepcopy funtion.
# Add elements to a list
patient_BMIs.append(26)
patient_BMIs
patient_BMIs.sort()
patient_BMIs
patient_BMIs.append("test")
patient_BMIs
patient_BMIs.append([35,"text"])
patient_BMIs
patient_BMIs.extend([22,26,33])
patient_BMIs
"""insert an element at a certain position
insert(position, element)"""
patient_BMIs.insert(3,29)
patient_BMIs
# Change list elements
patient_BMIs[3:5] = [25,25]
patient_BMIs
patient_BMIs[3:6] = [20,20,30,31]
patient_BMIs
patient_BMIs.sort()
patient_BMIs
patient_BMIs.index(20)
patient_BMIs.count(20)
20 in patient_BMIs
# Remove list elements
# <b>remove()</b> removes the first matching value, not a specific index.
patient_BMIs.remove(22)
patient_BMIs
# <b>del</b> removes the item at a specific index.
del patient_BMIs[4]
patient_BMIs
# <b>pop()</b> removes the item at a specific index and returns it.
patient_BMIs.pop(4)
patient_BMIs
# <b>clear()</b> removes all the elements of a list
patient_BMIs.clear()
patient_BMIs
# <b>String join()</b>
",".join(["Make","a","sentence."])
# <b>String split()</b>
"Get the, words".split(",")
| class_notebooks/variables_datatypes_clean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import csv
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
from collections import OrderedDict
# %matplotlib inline
fname='elc_output.csv'
colfilter = ['Process','Period','PV']
xlocs = [2013]
xlocs=xlocs+list(np.arange(2020,2101,10))
a = pd.read_csv(fname, delimiter=';', usecols = colfilter)
a = a[colfilter]
a=a.groupby(['Process', 'Period']).sum() #sum values for same period, process
a=a.reset_index()
a.set_index('Period', inplace=True)
procs = a.Process.unique()
newyrs=pd.Series(range(2013,2101))
pltdict=OrderedDict()
netgen=0
for prc in procs:
b=a.loc[a['Process'] == prc].reindex(newyrs,fill_value=0)
pltdict[prc]=b['PV']
netgen+=(pltdict[prc].sum())
#a.to_csv('df_test.csv', sep='\t')
#print(procs)
ordlegend=['Hydro','Nuclear',
'Coal','Oil', 'LNG',
'SOFC','CCS-Gas',
'Geothermal','Solar','Emerging Solar',
'Onshore Wind','Offshore Fixed',#,'Offshore Floating',
'Li-ion'
]
colours = ['aqua','orangered',
'black', 'saddlebrown', 'slategray',
'salmon','tan',
'green','yellow','maroon',
'blue','dodgerblue',#,'teal',
'violet'
]
gridRow=2
gridColumn=6
legendCols=5
fig = plt.figure(figsize=(20,10))
gs = fig.add_gridspec(gridRow,gridColumn)
ax1 = fig.add_subplot(gs[0,2:4])
ax1.set_title('Electricity supplied for demand',fontsize='18')
ax2 = fig.add_subplot(gs[1,1:3])
ax2.set_title('Capacity of supply & storage',fontsize='18')
ax3 = fig.add_subplot(gs[1,3:5])
ax3.set_title('Emission sources',fontsize='18')
fig.tight_layout()
fig.subplots_adjust(hspace=0.4, wspace=0.35)
pltyrs=range(2013,2101)
ax1.stackplot(pltyrs, pltdict['FTE-DIMEHYD13'],pltdict['FTE-DIMEONUC13'],
pltdict['ELCTECOA'],pltdict['ELCTEOIL'], pltdict['ELCTELNG']+pltdict['FTE-DIMECC17'],
pltdict['SOFC30'], pltdict['FTE-DIMECCSG22'],
pltdict['ELCTEGEO'],
pltdict['ELCTEPVS']+pltdict['FTE-DIMEPVN17'],pltdict['FTE-DIMESOL17'],
pltdict['ELCTEWND']+pltdict['FTE-DIMEWON17'],pltdict['FTE-DIMEWOFX17'],#pltdict['FTE-DIMEWOFL19'],
pltdict['FTE-ELCSTG17'],
labels=ordlegend, colors=colours)
ax1.tick_params(axis="x", labelsize=10)
ax1.tick_params(axis="y", labelsize=12)
ax1.get_yaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(lambda x, p: format(int(x), ',')))
ax1.set_xlim((2013, 2100))
ax1.legend(loc='upper center', bbox_to_anchor=(0.5, -0.05), shadow=True, ncol=legendCols)
#ax1.set_xlabel('Years',fontsize='18')
ax1.set_ylabel('Electricity supplied (GWh)',fontsize='16')
ax1.set_xticks(ticks=xlocs)
fname='co2_emi_sources.csv'
colfilter = ['Process','Period','PV']
cdf = pd.read_csv(fname, delimiter=';', usecols = colfilter)
cdf = cdf[colfilter]
cdf=cdf.groupby(['Process', 'Period']).sum() #sum values for same period, process
cdf=cdf.reset_index()
cdf.set_index('Period', inplace=True)
procs2 = cdf.Process.unique()
pltdict2={}
for prc in procs2:
c=cdf.loc[cdf['Process'] == prc].reindex(newyrs,fill_value=0)
pltdict2[prc]=c['PV']
#print(procs2)
fname='net_cap.csv'
colfilter = ['Process','Period','PV']
capdf = pd.read_csv(fname, delimiter=';', usecols = colfilter)
capdf = capdf[colfilter]
capdf=capdf.groupby(['Process', 'Period']).sum() #sum values for same period, process
capdf=capdf.reset_index()
capdf.set_index('Period', inplace=True)
procs3 = capdf.Process.unique()
pltdict4={}
for prc in procs3:
c=capdf.loc[capdf['Process'] == prc].reindex(newyrs,fill_value=0)
pltdict4[prc]=c['PV']
#print(procs3)
ordlegend2=['Hydro','Nuclear',
'Coal','Oil', 'LNG',
'AEC','PEMEC','SOEC','PWS',
'SOFC',
'CCS-Gas',
'Geothermal','Solar','Emerging Solar',
'Onshore Wind','Offshore Fixed',
'Li-ion'
]
colours2 = ['aqua','orangered',
'black', 'saddlebrown', 'slategray',
'olive','indigo','lightpink','lawngreen','salmon',
'tan',
'green','yellow','maroon',
'blue','dodgerblue',
'violet'
]
pltyrs2=range(2013,2101)
l2 = ax2.stackplot(pltyrs2, pltdict4['ELCTEHYD'],pltdict4['ELCTENUC'],
pltdict4['ELCTECOA'],pltdict4['ELCTEOIL'], pltdict4['ELCTELNG']+pltdict4['ELCCC17'],
pltdict4['ECALK22'],pltdict4['ECPEM22'],pltdict4['ECSOEC50'],pltdict4['PWS50'],pltdict4['SOFC30'],
pltdict4['LNGCCS22'],
pltdict4['ELCTEGEO'],
pltdict4['ELCTEPVS']+pltdict4['ELCPVN17'],pltdict4['EMSOL17'],
pltdict4['ELCTEWND']+pltdict4['ELCWON17'],pltdict4['ELCWOFX17'],
pltdict4['STGLION17'],
labels=ordlegend2, colors=colours2)
ax2.set_xlim((2013, 2100))
ax2.legend(loc='upper center', bbox_to_anchor=(0.5, -0.05), shadow=True, ncol=legendCols-1)
ax2.set_ylabel('Nameplate Capacity (GW)',fontsize='16')
ax2.set_xticks(ticks=xlocs)
ordlegend3=['Hydro','Nuclear',
'Coal','Oil', 'LNG',
'Alkaline Electrolyzer','PEM Electrolyzer','SOEC','PWS','SOFC',
'CCS-Gas',
'Geothermal','Solar','Emerging Solar',
'Onshore Wind','Offshore Fixed',
'Li-ion'
]
colours3 = ['aqua','orangered',
'black', 'saddlebrown', 'slategray',
'olive','indigo','lightpink','lawngreen','salmon',
'tan',
'green','yellow','maroon',
'blue','dodgerblue',
'violet'
]
pltyrs3=range(2013,2101)
ax3.stackplot(pltyrs3, pltdict2['ELCTEHYD'],pltdict2['ELCTENUC'],
pltdict2['ELCTECOA'],pltdict2['ELCTEOIL'], pltdict2['ELCTELNG']+pltdict2['ELCCC17'],
pltdict2['FTE-EALKH22'],pltdict2['FTE-EPEMH22'],pltdict2['FTE-ESOECH50'],pltdict2['PWS50'],
pltdict2['SOFC30'],pltdict2['LNGCCS22'],
pltdict2['ELCTEGEO'],
pltdict2['ELCTEPVS']+pltdict2['ELCPVN17'],pltdict2['EMSOL17'],
pltdict2['ELCTEWND']+pltdict2['ELCWON17'],pltdict2['ELCWOFX17'],
pltdict2['FTE-ELCSTG17'],
labels=ordlegend3, colors=colours3)
ax3.set_xlim((2013, 2100))
ax3.set_ylabel('Carbon Emissions (million t)',fontsize='16')
ax3.legend(loc='upper center', bbox_to_anchor=(0.5, -0.05), shadow=True, ncol=legendCols-1)
ax3.set_xticks(ticks=xlocs)
fig.savefig('lowtrltech_nonuc.png',bbox_inches='tight', dpi=300)
# +
fname='selc_sources.csv'
colfilter = ['Process','Period','PV']
sdf = pd.read_csv(fname, delimiter=';', usecols = colfilter)
sdf = sdf[colfilter]
sdf=sdf.groupby(['Process', 'Period']).sum() #sum values for same period, process
sdf=sdf.reset_index()
sdf.set_index('Period', inplace=True)
procs4 = sdf.Process.unique()
pltdict4={}
stgperc={}
stgsum=0
for prc in procs4:
s=sdf.loc[sdf['Process'] == prc].reindex(newyrs,fill_value=0)
pltdict4[prc]=s['PV']
stgsum+=(pltdict4[prc].sum())
print('Total storage electricity:',stgsum)
for prc in procs4:
stgperc[prc]=(pltdict4[prc].sum())/stgsum
print('Process:',prc)
print('Percentage:',stgperc[prc]*100)
# +
fname='h2_sources.csv'
colfilter = ['Process','Period','PV']
hdf = pd.read_csv(fname, delimiter=';', usecols = colfilter)
hdf = hdf[colfilter]
hdf=hdf.groupby(['Process', 'Period']).sum() #sum values for same period, process
hdf=hdf.reset_index()
hdf.set_index('Period', inplace=True)
procs5 = hdf.Process.unique()
pltdict5={}
h2perc={}
h2sum=0
for prc in procs5:
h=hdf.loc[hdf['Process'] == prc].reindex(newyrs,fill_value=0)
pltdict5[prc]=h['PV']
h2sum+=(pltdict5[prc].sum())
print('Total H2:',h2sum)
for prc in procs5:
h2perc[prc]=(pltdict5[prc].sum())/h2sum
print('Process:',prc)
print('Percentage:',h2perc[prc]*100)
# -
ccs=pltdict['FTE-DIMECCSG22'].sum()
print(ccs)
print(ccs/netgen)
| simulated_scenarios/low_trl_tech-no_new_nuclear/output2/plotter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Linear Algebra Examples
# ##### Note
# For this notebook an elmentwise matrix multiply is indicated with a $\cdot$ and a matrix product is indicated with an $*$. So
#
# $C= A \cdot B \rightarrow \left[ c_{i,j} \right] \leftarrow \left[a_{i,j} \right] \cdot \left[b_{i,j} \right]$ .
#
# and
#
# $C \leftarrow A * B \rightarrow \left[ c_{i,k} \right] = \sum_{i=0}^{M-1} \sum_{j=0}^{N-1} \left[a_{i,j} \right] \left[ b_{j,k} \right]$.
#
import pyJvsip as pjv
a=pjv.create('vview_d',10).fill(1.)
b=pjv.create('vview_d',10).ramp(0.,1.)
print('dot(a,b) = %.1f'%a.dot(b))
print('sumval(b) = %.1f'%b.sumval)
a=pjv.create('cvview_d',10).fill(1.)
b=pjv.create('cvview_d',10).randn(3)
a.mprint('%.2f');b.mprint('%.2f')
print('jdot(a,b) = (%.1f,%.1fi)'%(a.jdot(b).real,a.jdot(b).imag))
help(pjv.gemp)
# #### General Matrix Product
# gemp
#
# $C = \alpha \cdot op(A) * op(B) + \beta \cdot C$
#
# Where $\alpha$ and $\beta$ are scalar values.
# Make up some data
A=pjv.create('mview_d',4,3);
B=pjv.create('mview_d',4,4);
C=pjv.create('mview_d',3,4);
A.block.vector.ramp(1,1);
B.randn(4);
C.fill(1.0)
alpha=2;
beta=3;
# Do general matrix product
#
# alpha * transpose(A).prod(B) + beta * C
#
# longhand
opA=A.trans
opB=B
out=C.empty.fill(1.0)
out=alpha*opA.prod(opB)+beta*out
out.mprint('%.1f')
# Do general matrix product using gemp.
pjv.gemp(alpha,A,'TRANS',B,'NTRANS',beta,C)
C.mprint('%.1f')
# #### General Matrix Sum
# gems
#
# $C \leftarrow \alpha \cdot op(A) + \beta \cdot C$
A=pjv.create('mview_d',4,4);
C=A.empty;
A.block.vector.ramp(1,1);
C.fill(1.0)
alpha=2;
beta=3;
# Do it long hand
opA= A.trans
out=C.copy
pjv.mul(alpha,opA,opA)
pjv.mul(beta,out,out);
pjv.add(opA,out,out);
out.mprint('%.1f')
# Do it with gems
pjv.gems(alpha,A,'TRANS',beta,C).mprint('%.1f')
# ### Conjugate Dot Product
# #### **jdot**
# For vectors **a** and **b**
#
# $ \alpha = \sum \limits_{i=0}^{N-1} a_i \cdot$ opConj$(b_i) $
a=pjv.create('cvview_d',10).randn(4)
b=pjv.create('cvview_d',10).fill(complex(1,1))
a.jdot(b)
a.realview.jdot(b)
a.jdot(b.imagview)
a.realview.sumval
| doc/notebooks/LinearAlgebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="P9kMxNEtXPxK"
# # ***Support Vector Machine (SVM)***
# -
# Support vector machines (SVMs) are a set of supervised learning methods used for classification, regression and outliers detection.<br>
# The support vector machines in scikit-learn support both dense (numpy.ndarray and convertible to that by numpy.asarray) and sparse (any scipy.sparse) sample vectors as input. However, to use an SVM to make predictions for sparse data, it must have been fit on such data. For optimal performance, use C-ordered numpy.ndarray (dense) or scipy.sparse.csr_matrix (sparse) with dtype=float64.<br>
# They analyze the large amount of data to identify patterns from them.<br>
# SVMs are based on the idea of finding a hyperplane that best divides a dataset into two classes.
# <br><img src="https://miro.medium.com/max/600/0*0o8xIA4k3gXUDCFU.png" style="width:25%">
# ## **<ins>Problem</ins>**
# Breast cancer is the most common malignancy among women, accounting for nearly 1 in 3 cancers diagnosed among women in the United States, and it is the second leading cause of cancer death among women. Breast Cancer occurs as a results of abnormal growth of cells in the breast tissue, commonly referred to as a Tumor. A tumor does not mean cancer - tumors can be benign (not cancerous), pre-malignant (pre-cancerous), or malignant (cancerous). Tests such as MRI, mammogram, ultrasound and biopsy are commonly used to diagnose breast cancer performed.
#
# ## **<ins>Expected outcome</ins>**
# Given breast cancer results from breast fine needle aspiration (FNA) test (is a quick and simple procedure to perform, which removes some fluid or cells from a breast lesion or cyst (a lump, sore or swelling) with a fine needle similar to a blood sample needle). Since this build a model that can classify a breast cancer tumor using two training classification:
#
# 1= Malignant (Cancerous) - Present
# 0= Benign (Not Cancerous) -Absent
#
# ## **<ins>Objective</ins>**
# Since the labels in the data are discrete, the predication falls into two categories, (i.e. Malignant or benign). In machine learning this is a classification problem.
#
# Thus, the goal is to classify whether the breast cancer is benign or malignant and predict the recurrence and non-recurrence of malignant cases after a certain period. To achieve this we have used machine learning classification methods to fit a function that can predict the discrete class of new input.
# + [markdown] id="REpRoJiZX8Sb"
# ## Importing libraries
# + id="IICDFBCCEWrZ"
#Load libraries for data processing
import pandas as pd
import numpy as np
## Supervised learning.
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn import metrics
from sklearn.metrics import classification_report
# visualization
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="O2zZHCkaX_nL"
# ## Importing dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 240} id="lKXVOdeHGdUz" outputId="3388aa28-026c-4e6d-b6db-67cf0cbe316c"
# loading brest cancer risk prediction dataset
data = pd.read_csv('/content/breast-cancer-risk-prediction-dataset.csv', index_col=False)
data.drop('Unnamed: 0',axis=1, inplace=True)
# viewing the dataset
data.head()
# + [markdown] id="kQ-R3tKpYXpI"
# ## Spliting data into attributes and labels
# + id="PL0RdYFldpJI"
# Assign predictors to a variable of ndarray (matrix) type
array = data.values
X = array[:,1:31] # features
y = array[:,0] # labels
# + [markdown] id="x9ew8Oa_Yni_"
# ## Encoding the labels
# + id="pXWQZm7edrTL"
# transform the class labels from their original string representation (M and B) into integers using encoder
le = LabelEncoder()
y = le.fit_transform(y)
# + [markdown] id="qtuzfcwaYqzw"
# ## Normalizing the data
# + id="8ksIecmYGd5Z"
# Normalize the data (center around 0 and scale to remove the variance).
scaler =StandardScaler()
Xs = scaler.fit_transform(X)
# + [markdown] id="2GC9k6MYYgNe"
# ## Divide the data into training and testing sets.
# + id="s3a15AKVdmkL"
# Divide records in training and testing sets.
X_train, X_test, y_train, y_test = train_test_split(Xs, y, test_size=0.3, random_state=2, stratify=y)
# + [markdown] id="HN4r6WGdYiLe"
# ## Training the SVM algorithm
# + colab={"base_uri": "https://localhost:8080/"} id="tvzBXIICGg6i" outputId="5d5b1a1d-fdf7-4750-9c8a-f55a408c1497"
# Create an SVM classifier and train it on 70% of the data set.
clf = SVC(probability=True)
clf.fit(X_train, y_train)
# Analyze accuracy of predictions on 30% of the holdout test sample.
classifier_score = clf.score(X_test, y_test)
print('\nThe classifier accuracy score is {:03.2f}\n'.format(classifier_score))
# + [markdown] id="8aPKFCWWY39F"
# ## Plotting confusion matrix
# + colab={"base_uri": "https://localhost:8080/", "height": 486} id="TchwxPkgGrtu" outputId="577ab326-0c80-4165-9e23-a6e629dd50f0"
# displaying confustion matrix with the classification report
y_pred = clf.fit(X_train, y_train).predict(X_test)
cm = metrics.confusion_matrix(y_test, y_pred)
fig, ax = plt.subplots(figsize=(5, 5))
ax.matshow(cm, cmap=plt.cm.Reds, alpha=0.3)
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(x=j, y=i,
s=cm[i, j],
va='center', ha='center')
plt.xlabel('Predicted Values', )
plt.ylabel('Actual Values')
plt.show()
print(classification_report(y_test, y_pred ))
# + [markdown] id="eh7mTaXpXy-d"
# ## Optimizing the SVM classifier
# + colab={"base_uri": "https://localhost:8080/"} id="f5DEtXdkWhd7" outputId="c0da42ca-71a6-4823-dd2d-8b0cc0174c3b"
from sklearn.model_selection import GridSearchCV
# Train classifiers.
kernel_values = [ 'linear' , 'poly' , 'rbf' , 'sigmoid' ]
param_grid = {'C': np.logspace(-3, 2, 6), 'gamma': np.logspace(-3, 2, 6),'kernel': kernel_values}
grid = GridSearchCV(SVC(), param_grid=param_grid, cv=5)
grid.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="anu3xQEwWjTG" outputId="6db58969-99f3-4f36-ab7b-de17ed4197ab"
print("The best parameters are %s with a score of %0.2f"% (grid.best_params_, grid.best_score_))
# + colab={"base_uri": "https://localhost:8080/", "height": 486} id="ny4Up2uaWmZF" outputId="e3f539bf-b7fc-4894-ddd9-3e21d1f552bb"
grid.best_estimator_.probability = True
clf = grid.best_estimator_
# plotting confustion matrix for optimized classifier
y_pred = clf.fit(X_train, y_train).predict(X_test)
cm = metrics.confusion_matrix(y_test, y_pred)
#print(cm)
print(classification_report(y_test, y_pred ))
fig, ax = plt.subplots(figsize=(5, 5))
ax.matshow(cm, cmap=plt.cm.Reds, alpha=0.3)
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(x=j, y=i,
s=cm[i, j],
va='center', ha='center')
plt.xlabel('Predicted Values', )
plt.ylabel('Actual Values')
plt.show()
# + [markdown] id="WO28TXSCZ7r-"
# ## Decision boundaries of different classifiers
# + colab={"base_uri": "https://localhost:8080/", "height": 551} id="8zVJ3ZSzKYUl" outputId="cc354b69-ee36-455d-90d9-7a8eae0d3b55"
Xtrain = X_train[:, :2] # we only take the first two features.
C = 1.0 # SVM regularization parameter
svm = SVC(kernel='linear', random_state=0, gamma=0.1, C=C).fit(Xtrain, y_train)
rbf_svc = SVC(kernel='rbf', gamma=0.7, C=C).fit(Xtrain, y_train)
poly_svc = SVC(kernel='poly', degree=3, C=C).fit(Xtrain, y_train)
plt.rcParams['figure.figsize'] = (15, 9)
plt.rcParams['axes.titlesize'] = 'large'
# create a mesh to plot in
x_min, x_max = Xtrain[:, 0].min() - 1, Xtrain[:, 0].max() + 1
y_min, y_max = Xtrain[:, 1].min() - 1, Xtrain[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.1), np.arange(y_min, y_max, 0.1))
# title for the plots
titles = ['SVC with linear kernel',
'SVC with RBF kernel',
'SVC with polynomial (degree 3) kernel']
for i, clf in enumerate((svm, rbf_svc, poly_svc)):
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
plt.subplot(2, 2, i + 1)
plt.subplots_adjust(wspace=0.4, hspace=0.4)
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.coolwarm, alpha=0.8)
# Plot also the training points
plt.scatter(Xtrain[:, 0], Xtrain[:, 1], c=y_train, cmap=plt.cm.coolwarm)
plt.xlabel('radius_mean')
plt.ylabel('texture_mean')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.xticks(())
plt.yticks(())
plt.title(titles[i])
plt.show()
# -
#
# ## **<ins>Conclusion</ins>**
# This work demonstrates the modelling of breast cancer as classification task using Support Vector Machine
#
# The SVM performs better when the dataset is standardized so that all attributes have a mean value of zero and a standard deviation of one. We can calculate this from the entire training dataset and apply the same transform to the input attributes from the validation dataset.
# + id="nqd62-VKKuTK"
| Datascience_With_Python/Machine Learning/Libraries/Support Vector Machines/support_vector_machines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import pandas as pd
import numpy as np
from scipy import stats
from xgboost import XGBRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import RandomizedSearchCV
from sklearn.decomposition import PCA
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
from sklearn.base import TransformerMixin, BaseEstimator, clone
PROJECT_PATH = os.path.join(os.getcwd(), '../')
if PROJECT_PATH not in sys.path:
sys.path.append(PROJECT_PATH)
from server.ml_models.all_model import AllModelData
from server.ml_models.match_model import CATEGORY_COLS
from server.ml_models import EnsembleModel
from server.ml_models.data_config import TEAM_NAMES, ROUND_TYPES, VENUES
from src.model.metrics import yearly_performance_scores
from src.model.charts import graph_yearly_model_performance
SEED = 42
N_ITER = 5
CV = 5
np.random.seed(SEED)
# -
data = AllModelData(train_years=(None, 2015), test_years=(2016, 2016))
X_train, y_train = data.train_data()
X_train
# ## Test performance with new features
# +
model = EnsembleModel()
params = {
'pipeline__baggingregressor__n_estimators': stats.randint(5, 15),
'pipeline__baggingregressor__base_estimator__booster': ['gbtree', 'gblinear', 'dart'],
'pipeline__baggingregressor__base_estimator__colsample_bylevel': stats.uniform(0.8, 0.2),
'pipeline__baggingregressor__base_estimator__colsample_bytree': stats.uniform(0.8, 0.2),
'pipeline__baggingregressor__base_estimator__learning_rate': stats.uniform(0.075, 0.05),
'pipeline__baggingregressor__base_estimator__max_depth': stats.randint(2, 10),
'pipeline__baggingregressor__base_estimator__n_estimators': stats.randint(75, 150),
'pipeline__baggingregressor__base_estimator__reg_alpha': stats.uniform(0.0, 0.2),
'pipeline__baggingregressor__base_estimator__reg_lambda': stats.uniform(0.75, 0.5),
'pipeline__baggingregressor__base_estimator__subsample': stats.uniform(0.8, 0.2),
}
bag_cv = RandomizedSearchCV(model, params, scoring='neg_mean_absolute_error', n_jobs=1, cv=CV,
n_iter=N_ITER, random_state=SEED, verbose=1)
# -
bag_cv.fit(*data.train_data())
bag_cv.best_score_, bag_cv.refit_time_, bag_cv.best_params_
# +
data.train_years = (None, 2016)
X_train, y_train = data.train_data()
scores = yearly_performance_scores([('ensemble', model, {})], X_train, y_train, data_frame=True, parallel=False)
scores
# -
scores.groupby('model').mean()[['error', 'accuracy']]
# ### Performance is in the middle of prior models
# It's better than the first version of the ensemble model, but a mixed compared to the model with just the `elo_rating` feature added (higher accuracy & error).
# ## Reduce features via correlation with labels
# I had pretty good results using this when I had far fewer features, reducing from 76 to 18 with an increase in error of 0.16. Also, it's faster than PCA or `feature_importance` methods, because it doesn't require an extra model-training step in the pipeline.
# +
data.train_years = (None, 2015)
X_train, y_train = data.train_data()
X_train['margin'] = y_train
correlations = X_train.corr().fillna(0)['margin'].abs().sort_values(ascending=False)
correlations
# +
threshold_min = 0.001
threshold_max = 0.05
threshold_best = 0.038121046704238715
print(f'threshold={threshold_min}')
print(f'Feature count:', sum(correlations > threshold_min))
print(f'% feature sample:', sum(correlations > threshold_min) / len(correlations))
print()
print(f'threshold={threshold_max}')
print(f'Feature count:', sum(correlations > threshold_max))
print(f'% feature sample:', sum(correlations > threshold_max) / len(correlations))
print()
print(f'threshold={threshold_best}')
print(f'Feature count:', sum(correlations > threshold_best))
print(f'% feature sample:', sum(correlations > threshold_best) / len(correlations))
# +
class CorrelationChooser(TransformerMixin, BaseEstimator):
def __init__(self, labels, cols_to_keep=[], threshold=None):
self.labels = labels
self.threshold = threshold
self.cols_to_keep = cols_to_keep
self._cols_to_keep = self.cols_to_keep
def transform(self, X):
return X[self._cols_to_keep]
def fit(self, X, *_):
df = pd.concat([X, self.labels], axis=1).drop(self.cols_to_keep, axis=1)
self._cols_to_keep = df.columns[df.corr().fillna(0)[df.columns[-1]].abs() > self.threshold]
self._cols_to_keep = self.cols_to_keep + [col for col in self._cols_to_keep if col in X.columns]
return self
reduced_model = clone(model)
reduced_model.pipeline.steps = [
(
"correlationchooser",
CorrelationChooser(
y_train,
threshold=0.1,
cols_to_keep=["team", "oppo_team", "round_type", "venue"],
),
)
] + reduced_model.pipeline.steps
reduced_params = {
'pipeline__correlationchooser__threshold': stats.uniform(0.025, 0.024),
'pipeline__baggingregressor__n_estimators': stats.randint(5, 15),
'pipeline__baggingregressor__base_estimator__booster': ['gbtree', 'gblinear', 'dart'],
'pipeline__baggingregressor__base_estimator__colsample_bylevel': stats.uniform(0.8, 0.2),
'pipeline__baggingregressor__base_estimator__colsample_bytree': stats.uniform(0.8, 0.2),
'pipeline__baggingregressor__base_estimator__learning_rate': stats.uniform(0.075, 0.05),
'pipeline__baggingregressor__base_estimator__max_depth': stats.randint(2, 10),
'pipeline__baggingregressor__base_estimator__n_estimators': stats.randint(75, 150),
'pipeline__baggingregressor__base_estimator__reg_alpha': stats.uniform(0.0, 0.2),
'pipeline__baggingregressor__base_estimator__reg_lambda': stats.uniform(0.75, 0.5),
'pipeline__baggingregressor__base_estimator__subsample': stats.uniform(0.8, 0.2),
}
reduced_cv = RandomizedSearchCV(reduced_model, reduced_params, scoring='neg_mean_absolute_error', n_jobs=1, cv=CV,
n_iter=N_ITER, random_state=SEED, verbose=1)
# -
reduced_cv.fit(X_train.drop('margin', axis=1), y_train)
reduced_cv.best_score_, reduced_cv.refit_time_, reduced_cv.best_params_
# ## Reduce features with feature_importances_
# +
X_train, y_train = data.train_data()
X_train_numeric = X_train.drop(['team', 'oppo_team', 'round_type', 'venue'], axis=1)
dt = DecisionTreeRegressor()
dt.fit(X_train_numeric, y_train)
feature_importances = (
pd.Series(dt.feature_importances_, index=X_train_numeric.columns.values).sort_values(ascending=False)
)
feature_importances
# +
threshold_min = 0.0001
threshold_max = 0.0035
threshold_best = 0.0027294074748835753
print(f'threshold={threshold_min}')
print(f'Feature count:', sum(feature_importances > threshold_min))
print(f'% feature sample:', sum(feature_importances > threshold_min) / len(feature_importances))
print()
print(f'threshold={threshold_max}')
print(f'Feature count:', sum(feature_importances > threshold_max))
print(f'% feature sample:', sum(feature_importances > threshold_max) / len(feature_importances))
print()
print(f'threshold={threshold_best}')
print(f'Feature count:', sum(feature_importances > threshold_best))
print(f'% feature sample:', sum(feature_importances > threshold_best) / len(feature_importances))
# +
X_train, y_train = data.train_data()
class ImportanceChooser(TransformerMixin, BaseEstimator):
def __init__(self, labels, cols_to_keep=[], threshold=None, model=DecisionTreeRegressor()):
self.labels = labels
self.threshold = threshold
self.cols_to_keep = cols_to_keep
self._cols_to_keep = self.cols_to_keep
self.model = model
def transform(self, X):
return X[self._cols_to_keep]
def fit(self, X, *_):
_X = X.drop(self.cols_to_keep, axis=1)
_y = self.labels.loc[_X.index]
self.model.fit(_X, _y)
feature_importances = pd.Series(dt.feature_importances_, index=_X.columns.values)
self._cols_to_keep = feature_importances[feature_importances > self.threshold].index.values
self._cols_to_keep = self.cols_to_keep + [col for col in self._cols_to_keep if col in X.columns]
return self
import_model = clone(model)
import_model.pipeline.steps = [
(
"importancechooser",
ImportanceChooser(
y_train,
threshold=0.0001,
cols_to_keep=["team", "oppo_team", "round_type", "venue"],
),
)
] + import_model.pipeline.steps
import_params = {
'pipeline__importancechooser__threshold': stats.uniform(0.0018, 0.0017),
'pipeline__baggingregressor__n_estimators': stats.randint(5, 15),
'pipeline__baggingregressor__base_estimator__booster': ['gbtree', 'gblinear', 'dart'],
'pipeline__baggingregressor__base_estimator__colsample_bylevel': stats.uniform(0.8, 0.2),
'pipeline__baggingregressor__base_estimator__colsample_bytree': stats.uniform(0.8, 0.2),
'pipeline__baggingregressor__base_estimator__learning_rate': stats.uniform(0.075, 0.05),
'pipeline__baggingregressor__base_estimator__max_depth': stats.randint(2, 10),
'pipeline__baggingregressor__base_estimator__n_estimators': stats.randint(75, 150),
'pipeline__baggingregressor__base_estimator__reg_alpha': stats.uniform(0.0, 0.2),
'pipeline__baggingregressor__base_estimator__reg_lambda': stats.uniform(0.75, 0.5),
'pipeline__baggingregressor__base_estimator__subsample': stats.uniform(0.8, 0.2),
}
import_cv = RandomizedSearchCV(import_model, import_params, scoring='neg_mean_absolute_error', n_jobs=1, cv=CV,
n_iter=N_ITER, random_state=SEED, verbose=1)
# -
import_cv.fit(X_train, y_train)
import_cv.best_score_, import_cv.refit_time_, import_cv.best_params_
# ### Dimension reduction with PCA
# +
X_train, y_train = data.train_data()
pca = PCA()
pca.fit_transform(pd.get_dummies(X_train))
np.cumsum(pca.explained_variance_ratio_)
# +
import matplotlib.pyplot as plt
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.title('Scree Plot')
plt.xlabel('Principal Component (k)')
plt.ylabel('% of Variance Explained < = k')
None
# +
n_features = len(X_train.columns)
pca_model = clone(model)
pca_model.pipeline.steps = [
(
'columntransformer',
ColumnTransformer(
[
(
"onehotencoder",
OneHotEncoder(
categories=[TEAM_NAMES, TEAM_NAMES, ROUND_TYPES, VENUES], sparse=False
),
list(range(4)),
),
],
remainder=PCA(),
),
),
] + pca_model.pipeline.steps[1:]
pca_params = {
'pipeline__columntransformer__remainder__n_components': stats.randint(7, n_features - 1),
'pipeline__baggingregressor__n_estimators': stats.randint(5, 15),
'pipeline__baggingregressor__base_estimator__booster': ['gbtree', 'gblinear', 'dart'],
'pipeline__baggingregressor__base_estimator__colsample_bylevel': stats.uniform(0.8, 0.2),
'pipeline__baggingregressor__base_estimator__colsample_bytree': stats.uniform(0.8, 0.2),
'pipeline__baggingregressor__base_estimator__learning_rate': stats.uniform(0.075, 0.05),
'pipeline__baggingregressor__base_estimator__max_depth': stats.randint(2, 10),
'pipeline__baggingregressor__base_estimator__n_estimators': stats.randint(75, 150),
'pipeline__baggingregressor__base_estimator__reg_alpha': stats.uniform(0.0, 0.2),
'pipeline__baggingregressor__base_estimator__reg_lambda': stats.uniform(0.75, 0.5),
'pipeline__baggingregressor__base_estimator__subsample': stats.uniform(0.8, 0.2),
}
pca_cv = RandomizedSearchCV(pca_model, pca_params, scoring='neg_mean_absolute_error', n_jobs=1, cv=CV,
n_iter=N_ITER, random_state=SEED, verbose=1)
# -
pca_cv.fit(X_train, y_train)
pca_cv.best_score_, pca_cv.refit_time_, pca_cv.best_params_
| notebooks/2019_season/10.3-engineered-features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Anaconda 3.7
# language: python
# name: python3
# ---
from uproot_methods import TVector2, TVector3, TLorentzVector
# Getting ROOT, Python3, and Jupyter happy together on macOS was hard enough. Just going to grab HEPvector from disk. Don't do this in your code, just use pip to install.
import sys
sys.path.insert(0, '..')
from hepvector import Vector, Vector2D, Vector3D, LorentzVector
import numpy as np
# We need a way to compare a hepvector to an uproot vector:
def compare(npvec, rvec):
rveclist = [rvec.x, rvec.y]
if hasattr(rvec, 'z'):
rveclist.append(rvec.z)
if hasattr(rvec, 't'):
rveclist.append(rvec.t)
rvecarr = np.array(rveclist, np.double)
return np.allclose(npvec, rvecarr)
# # 2D vector comparison
v = Vector2D(1,2)
v2 = Vector2D(.3, .1)
r = TVector2(1,2)
r2 = TVector2(.3, .1)
# +
assert v.x == r.x
assert v.y == r.y
assert v[0] == r.x
assert v[1] == r.y
# -
assert v.phi == r.phi
assert v.rho == r.rho
assert v.mag == r.mag
assert v.mag2 == r.mag2
assert compare(v.unit, r.unit)
assert np.isclose(v.angle(v2), r.angle(r2))
# r.Norm(r2)
# r.Ort()
# r.Proj(r2)
# r.Rotate(phi)
# v.pt()
# v.pt2()
# # 3D vector comparison
v = Vector3D(1,2,3)
v1 = Vector3D(.4,-.1,.9)
r = TVector3(1,2,3)
r1 = TVector3(.4,-.1,.9)
assert v.x == r.x
assert v.y == r.y
assert v.z == r.z
r.cottheta
np.tan(v.theta)
assert v.mag == r.mag # get magnitude (=rho=Sqrt(x*x+y*y+z*z)))
assert v.mag2 == r.mag2 # get magnitude squared
assert v.theta == r.theta # get polar angle
#assert np.cos(v.theta) == r.CosTheta() # get cos of theta
assert np.isclose(np.tan(v.theta), r.cottheta)
assert v.phi == r.phi # get azimuth angle
#assert v.pt == r.Perp() # get transverse component
#assert v.pt2 == r.Perp2() # get transverse component**2
# +
# v.Perp(v1);
# v.Perp2(v1);
# +
# v.PseudoRapidity();
# Patch for bug in uproot_methods:
TVector3.TVector3 = TVector3
assert compare(v.cross(v1), r.cross(r1))
assert v.dot(v1) == r.dot(r1)
assert compare(v + v1, r + r1)
assert compare(v - v1, r - r1)
# r.Rotate* are oddly only inplace
# v.rotate_euler(phi=30)
# -
# # Lorentz vector comparison
v = LorentzVector(1,2,3,.2)
v1 = LorentzVector(.4,.1,-.7,2)
r = TLorentzVector(1,2,3,.2)
r1 = TLorentzVector(.4,.1,-.7,2)
assert v.x == r.x
assert v.y == r.y
assert v.z == r.z
assert v.t == r.t
assert v.e == r.t
assert v.beta == r.beta
assert np.isclose(v.dot(v1), r.dot(r1))
assert compare(v + v1, r + r1)
assert compare(v - v1, r - r1)
#assert v.pt == r.Perp()
#assert v.pt2 == r.Perp2()
assert compare(v.boostp3, r.boostp3)
assert v.delta_r(v1) == r.delta_r(r1)
assert v.eta == r.eta
assert v1.rapidity == r1.rapidity
# # Defined methods
#
# Let's look at all methods available (classmethods too):
# +
import inspect
from itertools import zip_longest
ignore = {x for x,y in inspect.getmembers(np.ndarray)}
ignore |= {'__module__', '_repr_html_', '__slots__'}
insp = lambda cls: {x for x,y in inspect.getmembers(cls)} - ignore
v0 = insp(Vector)
v2 = insp(Vector2D) - v0
v3 = insp(Vector3D) - v2
l4 = insp(LorentzVector) - v3
mc = lambda x: max(max(map(len, x)), 8)
a,b,c,d = 'Vector Vector2D Vector3D LorentzVector'.split()
print(f'{a:{mc(v0)}} | {b:{mc(v2)}} | {c:{mc(v3)}} | {d:{mc(l4)}}')
print(f'{"":-^{mc(v0)}}-|-{"":-^{mc(v2)}}-|-{"":-^{mc(v3)}}-|-{"":-^{mc(l4)}}')
for a,b,c,d in zip_longest(v0, v2, v3, l4, fillvalue=''):
print(f'{a:{mc(v0)}} | {b:{mc(v2)}} | {c:{mc(v3)}} | {d:{mc(l4)}}')
# -
| notebooks/CompareWithUprootMethods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
# %matplotlib inline
plt.rcParams['figure.figsize'] = [10, 10]
import os
import sys
import inspect
module_path = os.path.abspath(os.path.join('../../..'))
if module_path not in sys.path:
sys.path.append(module_path)
from dataset.dataset import get_test_datasets, get_validation_datasets, get_dataset_shapes
from experiment.model import get_scalar_model, get_vector_model
from experiment.learning import validate_model, validate_model_multiple, test_model, test_model_multiple
# +
train_dataset, test_dataset = get_test_datasets('cardiotocography2', random_seed=200, feature_range=(0,1))
datasets_generator_fun = lambda: get_validation_datasets('cardiotocography2', random_seed=200, feature_range=(0,1))
dataset_shapes = get_dataset_shapes('cardiotocography2')
num_tries = 15
dataset_shapes
# -
# # Scalar network without hidden layers
# +
model_fun = lambda: get_scalar_model(dataset_shapes, hidden_layer_units=[], activation='relu', output_activation=None, \
kernel_initializer='random_normal', bias_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# # Scalar network with one hidden layer
# ### S(2)
# +
model_fun = lambda: get_scalar_model(dataset_shapes, hidden_layer_units=[2], activation='relu', output_activation=None, \
kernel_initializer='random_normal', bias_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# ### S(5)
# +
model_fun = lambda: get_scalar_model(dataset_shapes, hidden_layer_units=[5], activation='relu', output_activation=None, \
kernel_initializer='random_normal', bias_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# ### S(15)
# +
model_fun = lambda: get_scalar_model(dataset_shapes, hidden_layer_units=[15], activation='relu', output_activation=None, \
kernel_initializer='random_normal', bias_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# ### S(30)
# +
model_fun = lambda: get_scalar_model(dataset_shapes, hidden_layer_units=[30], activation='relu', output_activation=None, \
kernel_initializer='random_normal', bias_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# # Scalar network with three hidden layers
# ### S(2,2,2)
# +
model_fun = lambda: get_scalar_model(dataset_shapes, hidden_layer_units=[2,2,2], activation='relu', output_activation=None, \
kernel_initializer='random_normal', bias_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# ### S(10,7,5)
# +
model_fun = lambda: get_scalar_model(dataset_shapes, hidden_layer_units=[10,7,5], activation='relu', output_activation=None, \
kernel_initializer='random_normal', bias_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# # Vector network with one hidden layer (c=1), unique weights
# ### V1(2):U(2)
# +
model_fun = lambda: get_vector_model(dataset_shapes, fractal_depth=1, hidden_layer_units=(2,), inner_hidden_layer_units=(2,), \
activation='relu', output_activation=None, \
weight_type="unique", weight_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# ### V1(5):U(2)
# +
model_fun = lambda: get_vector_model(dataset_shapes, fractal_depth=1, hidden_layer_units=(5,), inner_hidden_layer_units=(2,), \
activation='relu', output_activation=None, \
weight_type="unique", weight_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# # Vector network with one hidden layer (c=1), shared weights
# ### V1(2):S(2)
# +
model_fun = lambda: get_vector_model(dataset_shapes, fractal_depth=1, hidden_layer_units=(2,), inner_hidden_layer_units=(2,), \
activation='relu', output_activation=None, \
weight_type="shared", weight_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
# -
# ### V1(5):S(2)
# +
model_fun = lambda: get_vector_model(dataset_shapes, fractal_depth=1, hidden_layer_units=(5,), inner_hidden_layer_units=(2,), \
activation='relu', output_activation=None, \
weight_type="shared", weight_initializer='random_normal', \
optimizer=keras.optimizers.RMSprop(), loss=keras.losses.MeanSquaredError(), metrics=[keras.metrics.MeanSquaredError()])
validate_model_multiple(model_fun, datasets_generator_fun, epochs=175, num_tries=5, \
loss_name="mean_squared_error", measure_name="val_mean_squared_error")
| VNN/notebooks/network_experiments/cardiotocography2/validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
from __future__ import division
# +
import numpy as np
from phasor.utilities.ipynb.displays import *
from phasor.utilities.ipynb.sympy import *
import declarative
from declarative.bunch import (
DeepBunch
)
import phasor.math.dispatched as dmath
#import phasor.math.dispatch_sympy
# +
import phasor.utilities.version as version
print(version.foundations_version())
from phasor.utilities.np import logspaced
from phasor import optics
from phasor import base
from phasor import signals
from phasor import system
from phasor import readouts
import scipy.signal
# +
F_nyquist = 16384 / 2
F_AC = F_nyquist * 2 * np.arange(0, 1002) / 1002
ZPKz = (
[
.9,
-.3,
],
[
#.9,
.99,
#-.2,
],
10
)
b,a = scipy.signal.zpk2tf(*ZPKz)
Fb = mplfigB(Nrows=2)
w, h = scipy.signal.freqz_zpk(*ZPKz , worN = F_AC / F_nyquist * np.pi)
F_exact = h
Fb.ax0.loglog(F_AC, abs(h), label = 'discrete')
Fb.ax1.semilogx(F_AC, np.angle(h), label = 'discrete')
w, h = scipy.signal.freqz(b, a , worN = F_AC / F_nyquist * np.pi)
F_exact = h
Fb.ax0.loglog(F_AC, abs(h), label = 'discrete')
Fb.ax1.semilogx(F_AC, np.angle(h), label = 'discrete')
Fb.ax0.legend()
# -
F_AC[1] + F_AC[-1]
np.fft.fft(b, len(F_exact)) / F_exact
np.fft.ifft(F_exact)
fj = F_exact
n = 2
m = 300
N = len(fj)
col = np.fft.ifft(fj)/(N+1);
col = col
row = np.fft.ifft(fj.conjugate()).conjugate()/(N+1);
row = row
Z = scipy.linalg.toeplitz(col,row[0:n]);
[U,S,V] = np.linalg.svd(Z[m:,:]);
arat = V[:,-1];
qj = np.fft.fft(arat, N);
bh = np.fft.ifft(qj * fj);
brat = bh[0:m+0];
pj = np.fft.fft(brat,N+0);
print(S)
qj
brat / b
arat / a
# +
Fb = mplfigB(Nrows=2)
w, h = scipy.signal.freqz(b, a, worN = F_AC / F_nyquist * np.pi)
Fb.ax0.loglog(F_AC, abs(h), label = 'discrete')
Fb.ax1.semilogx(F_AC, np.angle(h), label = 'discrete')
h = F_exact
Fb.ax0.loglog(F_AC, abs(h), label = 'Fex')
Fb.ax1.semilogx(F_AC, np.angle(h), label = 'Fex')
w, h = scipy.signal.freqz(brat, arat, worN = F_AC / F_nyquist * np.pi)
Fb.ax0.loglog(F_AC, abs(h), label = 'fit', color = 'green')
Fb.ax1.semilogx(F_AC, np.angle(h), label = 'fit', color = 'green')
Fb.ax0.loglog(F_AC, abs(pj/qj), label = 'fit', color = 'cyan')
#Fb.ax0.set_xscale('linear')
#Fb.ax1.semilogx(F_AC, np.angle(h), label = 'fit', color = 'green')
Fb.ax0.legend()
# -
R = np.random.randn(len(F_AC)//2-1) + 1j*np.random.randn(len(F_AC)//2-1)
R2 = np.concatenate([[0], R, [0], R[::-1].conjugate()])
len(R2)
len(F_AC)
fj = F_exact * (1 + R2 / 100)
n = 2
m = 3
N = len(fj)
col = np.fft.ifft(fj)/(N+1);
col = col
row = np.fft.ifft(fj.conjugate()).conjugate()/(N+1);
row = row
Z = scipy.linalg.toeplitz(col,row[0:n]);
[U,S,V] = np.linalg.svd(Z[m:,:]);
arat = V[:,-1];
qj = np.fft.fft(arat, N);
bh = np.fft.ifft(qj * fj);
brat = bh[0:m+0];
pj = np.fft.fft(brat,N+0);
print(S)
qj
fj = F_exact * (1 + R2 / 100)
n = 2
m = 3
N = len(fj)
col = np.fft.ifft(fj)/(N+1);
col = col
row = np.fft.ifft(fj.conjugate()).conjugate()/(N+1);
row = row
Z = scipy.linalg.toeplitz(col,row[0:n]);
[U,S,V] = np.linalg.svd(Z[m:,:]);
arat = V[:,-1];
qj = np.fft.fft(arat, N);
bh = np.fft.ifft(qj * fj);
brat = bh[0:m+0];
pj = np.fft.fft(brat,N+0);
print(S)
qj
# +
Fb = mplfigB(Nrows=2)
w, h = scipy.signal.freqz(b, a, worN = F_AC / F_nyquist * np.pi)
Fb.ax0.loglog(F_AC, abs(h), label = 'discrete')
Fb.ax1.semilogx(F_AC, np.angle(h), label = 'discrete')
h = F_exact
Fb.ax0.loglog(F_AC, abs(h), label = 'Fex')
Fb.ax1.semilogx(F_AC, np.angle(h), label = 'Fex')
w, h = scipy.signal.freqz(brat, arat, worN = F_AC / F_nyquist * np.pi)
Fb.ax0.loglog(F_AC, abs(h), label = 'fit', color = 'green')
Fb.ax1.semilogx(F_AC, np.angle(h), label = 'fit', color = 'green')
Fb.ax0.loglog(F_AC, abs(pj/qj), label = 'fit', color = 'cyan')
#Fb.ax0.set_xscale('linear')
#Fb.ax1.semilogx(F_AC, np.angle(h), label = 'fit', color = 'green')
Fb.ax0.legend()
# -
| phasor/signals/test_direct_methods_true_ratdisk.ipynb |