code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise Notebook (DS)
# this code conceals irrelevant warning messages
import warnings
warnings.simplefilter('ignore', FutureWarning)
import numpy as np
elements = [3,5,6,8,2,9]
print(type(elements))
print(elements)
elements = np.array(elements)
print(type(elements))
print(elements)
# #### NumPy - Array Attributes
#
# `ndarray.shape`
# This array attribute returns a tuple consisting of array dimensions. It can also be used to resize the array.
#
# **Examples**
a = np.array([[1,2,3],[4,5,6]])
print (a.shape)
a
# +
# this resizes the ndarray
a = np.array([[1,2,3],[4,5,6]])
a.shape = (3,2)
print (a)
# -
# NumPy also provides a reshape function to resize an array.
a = np.array([[1,2,5],[4,5,6]])
b = a.reshape(3,2)
print (b)
# #### ndarray.ndim
# This array attribute returns the number of array dimensions.
#
# `Example`
elements.ndim
# an array of evenly spaced numbers
a = np.arange(24)
print (a)
# +
# this is one dimensional array
a = np.arange(24)
print(a.ndim)
# now reshape it
b = a.reshape(2,4,3)
print (b)
# b is having three dimensions
print(b.ndim)
# -
b[1][1][2]
# array of five zeros. Default dtype is float
x = np.zeros(5)
print (x)
# array of five ones. Default dtype is float
x = np.ones(5)
print (x)
# #### NumPy - Array From Existing Data
#
# **numpy.asarray**
# This function is similar to numpy.array except for the fact that it has fewer parameters. This routine is useful for converting Python sequence into ndarray.
# +
# convert list to ndarray
x = [1,2,3]
a = np.asarray(x)
print (a)
# +
# ndarray from list of tuples
x = [(1,2,3),(4,5)]
a = np.asarray(x)
print (a)
# -
# #### numpy.frombuffer
# This function interprets a buffer as one-dimensional array. Any object that exposes the buffer interface is used as parameter to return an ndarray.
#
# `numpy.frombuffer(buffer, dtype = float, count = -1, offset = 0)`
# ##### The constructor takes the following parameters.
#
# Sr.No. Parameter & Description
# 1. buffer
#
# Any object that exposes buffer interface
#
# 2. dtype
#
# Data type of returned ndarray. Defaults to float
#
# 3. count
#
# The number of items to read, default -1 means all data
#
# 4. offset
#
# The starting position to read from. Default is 0
# #### Operations on Numpy Array
# 2D Numpy Array
a = np.array([[1,2,3], [4,1,5]])
print (a)
# Addition
a+3
# Multiplication
a*2
# Subtraction
a-2
# Division
a/3
# **NumPy - Indexing & Slicing**
#
# Contents of ndarray object can be accessed and modified by indexing or slicing, just like Python's in-built container objects.
#
# As mentioned earlier, items in ndarray object follows zero-based index. Three types of indexing methods are available − field access, basic slicing and advanced indexing.
#
# Basic slicing is an extension of Python's basic concept of slicing to n dimensions. A Python slice object is constructed by giving **start, stop, and step** parameters to the built-in slice function. This slice object is passed to the array to extract a part of array.
a = np.arange(10)
print(a)
s = slice(2,7,2)
print(a[s])
# + active=""
# In the above example, an ndarray object is prepared by arange() function. Then a slice object is defined with start, stop, and step values 2, 7, and 2 respectively. When this slice object is passed to the ndarray, a part of it starting with index 2 up to 7 with a step of 2 is sliced.
#
# The same result can also be obtained by giving the slicing parameters separated by a colon : (start:stop:step) directly to the ndarray object.
# -
a = np.arange(10)
print(a)
b = a[2:7:2]
print (b)
# +
a = np.array([[1,2,3],[3,4,5],[4,5,6]])
print (a )
# slice items starting from index
print('Now we will slice the array from the index a[1:]')
print(a[1:])
# -
# #### Numpy 2D Array
a = np.array([[1,2,3],[3,4,5]])
print(a)
a = np.array([[1,2,3], [4,1,5]])
print (a)
# Addition
a+3
# Multiplication
a*2
# Subtraction
a-2
# Division
a/3
# ##### Task
# 1. Write a NumPy program to test element-wise for NaN of a given array.
import numpy as np
a = np.array([1, 0, 4, 8, np.nan, np.inf])
print("Original array")
print(a)
print("Test element-wise for NaN:")
print(np.isnan(a))
# 2. Write a NumPy program to test element-wise for complex number, real number of a given array. Also test whether a given number is a scalar type or not.
import numpy as np
a = np.array([7+1j, 1+0j, 4.5, 3, 3+2j, 7j])
print("Original array")
print(a)
print("Checking for complex number:")
print(np.iscomplex(a))
print("Checking for real number:")
print(np.isreal(a))
print("Checking for scalar type:")
print(np.isscalar(3.1))
print(np.isscalar([3.1]))
| Omobolanle Adeyemi WT 192/10.ipynb |
# # Create General KLDCriterion
# ## Verify forward pass
require 'BKLDCriterion'
require 'KLDCriterion'
bkld = nn.BKLDCriterion()
kld = nn.KLDCriterion()
mu = torch.randn(3,4)
lv = torch.randn(3,4):pow(2):log()
pmu = torch.zeros(3,4)
plv = torch.zeros(3,4)
print(bkld:forward(mu, lv))
print(kld:forward({pmu, plv}, {mu, lv})) -- make sure you use the right one as "target"!
# ## Verify backward pass
mu = torch.randn(3,4)
lv = torch.randn(3,4):pow(2):log()
pmu = torch.randn(3,4)
plv = torch.randn(3,4):pow(2):log()
dpmu, dplv, dmu, dlv = unpack(kld:backward({pmu, plv}, {mu, lv}))
h = 1e-5
for i = 1,mu:size(1) do
for j = 1,mu:size(2) do
mu[{i, j}] = mu[{i, j}] + h
fph = kld:forward({pmu, plv}, {mu, lv})
mu[{i, j}] = mu[{i, j}] - h - h
fmp = kld:forward({pmu, plv}, {mu, lv})
mu[{i, j}] = mu[{i, j}] + h
print((fph - fmp)/2/h - dmu[{i, j}])
end
end
for i = 1,lv:size(1) do
for j = 1,lv:size(2) do
lv[{i, j}] = lv[{i, j}] + h
fph = kld:forward({pmu, plv}, {mu, lv})
lv[{i, j}] = lv[{i, j}] - h - h
fmp = kld:forward({pmu, plv}, {mu, lv})
lv[{i, j}] = lv[{i, j}] + h
print((fph - fmp)/2/h - dlv[{i, j}])
end
end
for i = 1,pmu:size(1) do
for j = 1,pmu:size(2) do
pmu[{i, j}] = pmu[{i, j}] + h
fph = kld:forward({pmu, plv}, {mu, lv})
pmu[{i, j}] = pmu[{i, j}] - h - h
fmp = kld:forward({pmu, plv}, {mu, lv})
pmu[{i, j}] = pmu[{i, j}] + h
print((fph - fmp)/2/h - dpmu[{i, j}])
end
end
for i = 1,plv:size(1) do
for j = 1,plv:size(2) do
plv[{i, j}] = plv[{i, j}] + h
fph = kld:forward({pmu, plv}, {mu, lv})
plv[{i, j}] = plv[{i, j}] - h - h
fmp = kld:forward({pmu, plv}, {mu, lv})
plv[{i, j}] = plv[{i, j}] + h
print((fph - fmp)/2/h - dplv[{i, j}])
end
end
# # Basic matrix stuff
-- check my implementation is correct
-- create mu's
mu1 = torch.randn(3,1)
mu2 = torch.randn(3,1)
-- create sigma
sig = torch.diag(torch.randn(3):pow(2))
sig_lin = torch.diag(sig):reshape(3,1)
-- first approach
(mu1 - mu2):t() * (torch.inverse(sig)) * (mu1 - mu2)
-- second approach
torch.Tensor(1):fill((mu1 - mu2):pow(2):cdiv(sig_lin):sum())
# # Test inplace
x = torch.Tensor(1):fill(3)
x:add(3):add(x)
| notebooks/gradient_checker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/maxaubel/Scientific-Computing/blob/master/Tarea_1_CC.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="aBPGOpUM11V1"
#
# <center>
# <h1> INF285 - Computación Científica </h1>
# <h1> Tarea N°1, V0.3 </h1>
#
# </center>
#
# <p>
#
# </p>
# + [markdown] id="B7MCAVSDRk3l"
# ## Instrucciones
#
# * La tarea es individual, sin embargo se invita a todos l@s estudiantes a que discutan entre ustedes las preguntas pero luego implementen de forma individual su tarea.
# * Las consultas sobre las tareas se deben realizar por medio de la plataforma Aula.
# * La tarea debe ser realizada en `Jupyter Notebook` (`Python3`).
# * Se evaluará la correcta utilización de librerias `NumPy`, `SciPy`, entre otras, así como la correcta implementación de algoritmos de forma vectorizada.
# * **No modifique la firma de las funciones** (a menos que se le diga lo contrario) y respete el output que se le exije. **En caso de no respetar esta regla la función se considerará errónea.**
# * **El archivo de entrega debe denominarse ROL-tarea-numero.ipynb**. _De no respetarse este formato existirá un descuento de **50 puntos**_
# * La fecha de entrega es el jueves 6 de mayo a las **18:00 hrs**.
# * Debe citar cualquier código ajeno utilizado (incluso si proviene de los Jupyter Notebooks del curso).
# * Puede agregar funciones extras siempre y cuando **no interfieran en las firmas** de las funciones principales establecidas.
# + [markdown] id="PNhvmtRq2Orx"
# ## Introducción
#
# En esta primera tarea de INF-285, versión 2021-1, estudiaremos la importancia de los primeros temas estudiados en el curso, los cuales son: Representación de Punto Flotante, Pérdida de Importancia, Errores de Cancelación y Búsqueda de Ceros.
# El desarrollo de cada uno de esos temas se presenta en una serie de preguntas donde deberá ir decidiendo, pregunta a pregunta, cómo cada uno de los temas se aplica.
# En general, los temas no se analizan por separado, sino de manera acoplada.
# Es muy importante que cada uno de los problemas sea analizado teóricamente primero, para luego poner su conocimiento en acción.
# Cada problema puede ser desarrollado de diversas formas, sin embargo, es muy importante determinar al final si el camino elegido resuelve la pregunta presentada.
# Es decir, debe aplicar la metodología IDEA!
# + [markdown] id="lAtn1U0qRg2S"
# ## Problemas
# + [markdown] id="3znHUOxOUB0-"
# ### 1. Simulador (60 ptos)
#
# Dada la familia de polinomios de grado 3 con la forma:
#
#
# \begin{equation}
# f(x)=-1 + x + 2 A^2 x - 3 A^2 x^2 + A^2 x^3
# \end{equation}
#
# Se pide implementar un algoritmo que reciba como parámetros el valor de $A$ (con $|A|>>1$), 'bits_mant' que corresponde a la cantidad de bits que tiene la mantisa y 'bits_exp' que corresponde a la cantidad de bits que tiene el exponente.
# El algoritmo se ejecutará en un computador ficticio que representa los números con esa cantidad de bits para el estándar de punto flotante IEEE.
# Debe controlar los casos especiales correctamente.
# Esta implementación debe calcular las raíces de $f$ sin pérdida de **importancia** con la cantidad de bits disponibles para la mantisa y la cantidad de bits para el exponente.
# Para obtener las raíces de $f$ usted debe encontrar de forma algebraica sus raíces y luego proponer un algoritmo basado en las _fórmulas_ obtenidas.
#
# El computador ficticio cuenta con las operaciones matemáticas necesarias para obtener las raíces.
# Considere como límites de bits para la mantisa en el intervalo $[2,52]$ y en el intervalo $[2,11]$ para los bits del exponente.
#
# Ejemplo:
#
# ```python
# # Algoritmo de ejemplo
# a = 9819824.624837
# b = 148736.523476
# c = a+b
#
# # Implementación de ejemplo con Representación de Punto Flotante de 'bits_mant' bits en la mantisa y 'bits_exp' bits en el exponente.
# am = f_new_rep(9819824.624837,bits_mant,bits_exp) # Aproximar el input en la nueva representación.
# bm = f_new_rep(148736.523476,bits_mant,bits_exp) # Aproximar el input en la nueva representación.
# cm = f_suma(am,bm,bits_mant,bits_exp) # Aproximar el output de la suma y cada operación en la nueva representación.
# ```
#
# + id="l1Yapf7cOIPf" colab={"base_uri": "https://localhost:8080/"} outputId="d6a42ed7-527a-4c7d-e1dd-838359d463d5"
# # !pip install bitstring # to run on google colab
import bitstring as bs
import numpy as np
# + id="2efm5YD2UBGD"
"""
input
x : (double) valor a evaluar
bits_mant : (int) cantidad de bits de la mantisa
bits_exp : (int) cantidad de bits del exponente
output
result : (double) resultado obtenido
"""
def f_new_rep(x, bits_mant, bits_exp):
# Algoritmo de representación de punto flotante modificada.
bin_num = bs.pack('>d', x).bin
sign_og, exp_og, mant_og = bin_num[0], bin_num[1:12], bin_num[12:12+bits_mant]
sign_dec = 1
if (sign_og == "1"): sign_dec = -1
# Check first for special cases
if exp_og == "1"*len(exp_og):
if mant_og == "0"*len(mant_og):
return np.inf*sign_dec # Infinity and beyond
else: return np.nan # Not a Number
if bin_num == "0"*len(bin_num): return 0 # Zero
# Mantissa
mant_dec = 0
for i, n in enumerate(mant_og):
if(n == "1"): mant_dec += 2**-(i+1)
# Exponent
offset_og = int("1"*10, 2)
dec_og = int(exp_og, 2) - offset_og
if dec_og > 2**(bits_exp-1):
dec_og = 2**(bits_exp-1)
elif dec_og < -2**(bits_exp-1):
return 0
# Finally, "carefully" calculate the decimal
num = sign_dec * (1+mant_dec) * (2**dec_og)
return num
"""
input
n1 : (double) valor a sumar
n2 : (double) valor a sumar
bits_mant : (int) cantidad de bits de la mantisa
bits_exp : (int) cantidad de bits del exponente
output
result : (double) resultado obtenido
"""
def f_suma(n1,n2,bits_mant,bits_exp):
#Algoritmo que calcula la suma de dos numeros manteniendo el estandar asignado
n1, n2 = f_new_rep(n1, bits_mant, bits_exp), f_new_rep(n2, bits_mant, bits_exp)
result = f_new_rep(n1 + n2, bits_mant, bits_exp)
return result
"""
input
n1 : (double) valor a multiplicar
n2 : (double) valor a multiplicar
bits_mant : (int) cantidad de bits de la mantisa
bits_exp : (int) cantidad de bits del exponente
output
result : (double) resultado obtenido
"""
def f_multiplicacion(n1,n2,bits_mant,bits_exp):
#Algoritmo que calcula la multiplicación de dos numeros manteniendo el estandar asignado
n1, n2 = f_new_rep(n1, bits_mant, bits_exp), f_new_rep(n2, bits_mant, bits_exp)
result = f_new_rep(n1 * n2, bits_mant, bits_exp)
return result
"""
input
n1 : (double) valor a multiplicar
n2 : (double) valor a multiplicar
bits_mant : (int) cantidad de bits de la mantisa
bits_exp : (int) cantidad de bits del exponente
output
result : (double) resultado obtenido
"""
def f_division(n1,n2,bits_mant,bits_exp):
#Algoritmo que calcula la división de dos numeros manteniendo el estandar asignado
n1, n2 = f_new_rep(n1, bits_mant, bits_exp), f_new_rep(n2, bits_mant, bits_exp)
result = f_new_rep(n1 / n2, bits_mant, bits_exp)
return result
"""
input
A : (double) valor a evaluar
bits_mant : (int) cantidad de bits de la mantisa
bits_exp : (int) cantidad de bits del exponente
output
x_roots : (array) raices del polinomio ordenadas de menor a mayor
"""
def f_find_roots(A,bits_mant,bits_exp, with_sqrt=True):
# Algoritmo para encontrar raíces de f(x).
r_1 = 1 # Trivial root
square = lambda x: f_multiplicacion(x, x, bits_mant, bits_exp)
n = f_new_rep(square(A)-1, bits_mant, bits_exp)
if not with_sqrt:
non_trivial_root = f_new_rep(f_bisection(n, 1, n, square, bits_mant, bits_exp, depth=500), bits_mant, bits_exp)
else:
non_trivial_root = f_new_rep(np.sqrt(n), bits_mant, bits_exp)
fraction = f_division(non_trivial_root, abs(A), bits_mant, bits_exp)
r_0 = 1 - fraction
r_2 = 1 + fraction
return (r_0, r_1, r_2)
"""
input
n : (double) numero al que se le buscará la raíz
a : (double) cota inferior de la biseccion
b : (double) cota superior de la biseccion
f : (function) funcion para la que se buscará la raiz
bits_mant : (int) cantidad de bits de la mantisa
bits_exp : (int) cantidad de bits del exponente
output
x_roots : (double) raiz
"""
def f_bisection(n, a, b, f, bits_mant, bits_exp, depth):
# Se implementó este método para calcular la raiz antes de que se dijera que se podía usar sqrt
c = f_division( f_suma(a, b, bits_mant, bits_exp), 2, bits_mant, bits_exp);
f_c = f(c); # c^2
depth -= 1
if ((f_c == n) or (depth <= 0)):
return c
elif (f_c < n):
return f_bisection(n, c, b, f, bits_mant, bits_exp, depth)
else:
return f_bisection(n, a, c, f, bits_mant, bits_exp, depth)
# + colab={"base_uri": "https://localhost:8080/"} id="EV6RWjvbuqR5" outputId="5c7b6d4d-d35d-4877-df95-c7ba9e0f9930"
["%.55f" % root for root in f_find_roots(A=3,bits_mant=5,bits_exp=5)]
# + [markdown] id="Nw-jJfD-VPMa"
# ### Sinusoidal de Oreman (40 puntos)
# Considere la siguiente función en 3 variables:
# \begin{equation}
# T(t,\alpha,\beta) = \alpha\cdot \cos(t)+\beta\cdot \sin(\log(t)).
# \end{equation}
#
# La cual permite calcular la temperatura de una cámara de ambiente dado los parámetros $\alpha$ y $\beta$ los cuales corresponden a las condiciones iniciales del experimento.
# Por otro lado $t$ representa el tiempo en días transcurridos desde que se inició.
#
# A usted le permiten utilizar la cámara por un tiempo limitado, para esto planea terminar a los 25 días pero debe asegurarse que las sustancias sean extraídas a temperatura 0.
#
# Cree un programa que reciba los parametros $\alpha$ y $\beta$ y que retorne el día en el cual deba ser extraído el material cumpliendo la condición de ser el día más cercano a 25 (ya sea antes o después).
#
# + id="x0rfc_-6JTrG"
"""
input
alpha: (float) parametro alpha
beta: (float) parametro beta
tol: (float) tolerancia
output
root: (double) raiz obtenida
"""
# Code based on Newton's method approach in https://github.com/tclaudioe/Scientific-Computing/blob/master/SC1v2/03_roots_of_1D_equations.ipynb
# Modified to work on both sides of the requested x value
def raiz_de_oreman(alpha, beta, tol=1e-10):
fp = lambda x: beta*np.cos(np.log(x))/x - alpha*np.sin(x)
f = lambda x: alpha*np.cos(x) + beta*np.sin(np.log(x))
x0_low, x0_up = 23, 25.25
hybrid_error_low = 100
hybrid_error_up = 100
error_i_low = np.inf
error_i_up = np.inf
max_iters = 100
i = 1
while (((hybrid_error_low > tol and hybrid_error_low < 1e12) or
(hybrid_error_up > tol and hybrid_error_up < 1e12)) and i<=max_iters):
x1_low = x0_low - f(x0_low) / fp(x0_low)
x1_up = x0_up - f(x0_up ) / fp(x0_up )
if f(x1_low) == 0.0 or f(x1_up) == 0.0:
hybrid_error = 0.0
break
hybrid_error_low = abs(x1_low-x0_low) / np.max([abs(x1_low), 1e-12])
hybrid_error_up = abs(x1_up-x0_up) / np.max([abs(x1_up), 1e-12])
error_iminus1_low = error_i_low
error_iminus1_up = error_i_up
error_i_low = abs(x1_low-x0_low)
error_i_up = abs(x1_up-x0_up)
x0_low = x1_low
x0_up = x1_up
i+=1
if (hybrid_error_low < tol) and (abs(x1_low-25) <= abs(x1_up-25)):
return x1_low
if (hybrid_error_up < tol) and (abs(x1_up-25) <= abs(x1_low-25)):
return x1_up
if i>=max_iters:
print('Newtons Method did not converge. Too many iterations!!')
return None
else:
print('Newtons Method did not converge!!')
print(alpha, beta)
return None
# + colab={"base_uri": "https://localhost:8080/"} id="q05zWGYizKdb" outputId="3e655a95-57e6-43ed-c623-5f3e5c9cf611"
raiz_de_oreman(1.6, 20)
# + [markdown] id="gVOkWN_zrvR9"
# # Referencias
#
# https://github.com/tclaudioe/Scientific-Computing/blob/master/SC1v2/03_roots_of_1D_equations.ipynb
| Tarea_1_CC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="exterior-meaning"
# # Deviations from Normality
# + id="sized-joseph" outputId="e87bce30-c739-4c43-e334-64e40e44cb7d"
# %load_ext autoreload
# %autoreload 2
import pandas as pd
import edhec_risk_kit as erk
# + id="valid-sequence" outputId="29855759-e76a-4647-fc32-27053518e02d"
hfi = erk.get_hfi_returns()
hfi.head(4)
# + id="radio-actress" outputId="4cd0d3a2-fc84-4e2b-c560-2d8990cc9e53"
pd.concat([hfi.mean(), hfi.median(), hfi.mean() > hfi.median()], axis='columns')
# + [markdown] id="fourth-departure"
# ## measuring skewness
# + id="indian-regulation" outputId="28d2d157-5f78-4655-d265-d4ff56e411e8"
erk.skewness(hfi).sort_values()
# + id="effective-shield"
import scipy.stats
# + id="armed-distributor" outputId="b4b6f7d1-c23a-4820-bede-884dd7a9753c"
scipy.stats.skew(hfi)
# + id="female-coordinator"
import numpy as np
# + id="reduced-resort" outputId="085cb34e-78dc-40f1-8a23-daf32530b6d9"
hfi.shape
# + id="competent-identity"
normal_rets = np.random.normal(0, .15, (263, 1))
# + id="covered-guide" outputId="59600474-040f-457f-b49f-b6881371f145"
erk.skewness(normal_rets)
# + [markdown] id="thorough-examination"
# ## Kurtosis
# + id="alone-france" outputId="91a83eeb-72bc-469b-be44-391df1f22397"
erk.kurtosis(normal_rets)
# + id="institutional-doctrine" outputId="41009bea-1e1b-49a0-ebcf-be000a3e7a70"
erk.kurtosis(hfi)
# + id="modern-terrace" outputId="1601a0b1-be4c-4e5f-a72a-d4de95603273"
scipy.stats.kurtosis(normal_rets)
# + id="greater-variance" outputId="e35567ea-a3d7-479c-d594-110ec7648869"
scipy.stats.kurtosis(hfi)
# + id="generic-addition" outputId="d3a00e4c-ed2b-45c1-e169-6969a3843566"
scipy.stats.jarque_bera(normal_rets)
# + id="weekly-hungarian" outputId="33a5832d-3e78-47c9-8d62-3ff79f4a4626"
scipy.stats.jarque_bera()
# + id="embedded-signal" outputId="4544d077-34e9-4e06-b612-d4322c303138"
erk.is_normal(normal_rets)
# + id="lonely-belarus" outputId="a5862440-ba04-4462-9fe3-ec5af030177e"
hfi.apply(erk.is_normal, axis=0)
# + id="polish-invalid" outputId="e6fe7fa2-59aa-4ea7-c12c-cf3218a333f9"
hfi.aggregate(erk.is_normal)
# + id="religious-vatican" outputId="c8383e42-aa06-4143-8aa8-5700b8750aea"
ffme = erk.get_ffme_returns()
erk.skewness(ffme)
# + id="moving-ethics" outputId="e49fb9b3-b07d-4c65-c90d-9de03032da4f"
erk.kurtosis(ffme)
# + id="cathedral-charity" outputId="5f5702cd-094a-4e21-9d99-1fee6271911d"
ffme.aggregate(erk.is_normal)
# + id="internal-atlanta"
| course1portfolio_construction/Copy_of_deviationsfromNormality.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from PyQt5.QtCore import QDate, QTime, QDateTime, Qt
# +
# The QDate is a class for working with calendar date in the Gregorian calendar.
# The QTime class works with a clock time.
# The QDateTime is a class that combines both QDate and QTime objects into one object.
# -
# ## Current date and time
# +
now = QDate.currentDate()
print(now.toString(Qt.ISODate))
print(now.toString(Qt.DefaultLocaleLongDate))
datetime = QDateTime.currentDateTime()
print(datetime.toString())
time = QTime.currentTime()
print(time.toString(Qt.DefaultLocaleLongDate))
# This example prints the current date, date and time, and time in various formats.
# -
now = QDate.currentDate()
# The currentDate method returns the current date.
print(now.toString(Qt.ISODate))
print(now.toString(Qt.DefaultLocaleLongDate))
datetime = QDateTime.currentDateTime()
# The currentDateTime returns current date and time.
print(datetime.toString(Qt.ISODate))
print(datetime.toString(Qt.DefaultLocaleLongDate))
time = QTime.currentTime()
# The currentTime method returns the current time
print(time.toString(Qt.ISODate))
print(time.toString(Qt.DefaultLocaleLongDate))
# ## UTC Time
# +
now = QDateTime.currentDateTime()
print('Local datetime: ', now.toString(Qt.ISODate))
print('Universal datetime: ', now.toUTC().toString(Qt.ISODate))
print(f'The offset from UTC is: {now.offsetFromUtc()} seconds')
# This example determines the current universal and local date and time.
# +
# The currentDateTime returns the current date and time expressed as local time.
# The toUTC method returns the universal time.
# The offsetFromUtc gives the difference between universal time and local time in seconds.
# -
# ## Number of days
# +
now = QDate.currentDate()
d = QDate(1945, 5, 7)
print(f'Days in month: {d.daysInMonth()}')
print(f'Days in year: {d.daysInYear()}')
# This example prints the number of days in a month and year for the chosen date.
# -
# ## Difference in days
# +
xmas1 = QDate(2020, 12, 24)
xmas2 = QDate(2021, 12, 24)
now = QDate.currentDate()
daypassed = xmas1.daysTo(now)
print(f'{daypassed} days have passed since last XMas.')
nofdays = now.daysTo(xmas2)
print(f'There are {nofdays} days until next XMas.')
# This example calculate the number of days passed from the last XMas and the number of days until the next XMas.
# -
# ## Datetime arithmetic
# +
now = QDateTime.currentDateTime()
print(f'Today: {now.toString(Qt.ISODate)}')
print(f'Adding 12 days: {now.addDays(12).toString(Qt.ISODate)}')
print(f'Subtracting 22 days: {now.addDays(-22).toString(Qt.ISODate)}')
print(f'Adding 50 seconds: {now.addSecs(50).toString(Qt.ISODate)}')
print(f'Adding 3 months: {now.addMonths(3).toString(Qt.ISODate)}')
print(f'Adding 12 years: {now.addYears(12).toString(Qt.ISODate)}')
# The example determines the current datetime and add ot subtract days, seconds, months and years.
# -
# ## Daylight saving time
# +
now = QDateTime.currentDateTime()
print(f'Time zone: {now.timeZoneAbbreviation()}')
if now.isDaylightTime():
print(f'The current date falls into DST time.')
else:
print(f'The current date does not fall into DST time.')
# This example checks if the datetime is in the daylight savng time.
# +
# The timeZoneAbbreviation() method returns the time zone abbreviation for the datetime.
# the isDayLightTime() method returns if the datetime falls in daylight saving time.
# -
# ## Unix epoch
# +
# The Unix epoch is the time 00:00:00 UTC on 1 January 1970.
# The date and time in a computer is determined according to the number of seconds or clock
# ticks that have elapsed since the defined epoch for that computer platform.
# Unix time is the number of seconds elapsed since Unix epoch.
# +
now = QDateTime.currentDateTime()
unix_time = now.toSecsSinceEpoch()
print(unix_time)
d = QDateTime.fromSecsSinceEpoch(unix_time)
print(d.toString(Qt.ISODate))
# This example prints the Unix time and converts it back to the QDateTime.
# The toSecsSinceEpoch() returns the Unix time.
# the fromSecsSinceEpoch() converts the Unix time to QDateTime.
# -
# ## Julian Day
now = QDate.currentDate()
print(f'Gregorian date for today: {now.toString(Qt.ISODate)}')
print(f'Julian day for today: {now.toJulianDay()}')
# In this example we compute the Gregorian date and the Julian day for today.
# The Julian day is returned with the toJulianDay() method.
# ## Historical battles
# +
borodino_battle = QDate(1812, 9, 7)
slavkov_battle = QDate(1805, 12, 2)
now = QDate.currentDate()
j_today = now.toJulianDay()
j_bordino = borodino_battle.toJulianDay()
j_slavkov = slavkov_battle.toJulianDay()
d1 = j_today - j_slavkov
d2 = j_today - j_bordino
print(f'Days since Slavkov battle: {d1}')
print(f'Days since Borodino Battle: {d2}')
# This example counts the number of days passed since two historical events.
| 01 PyQt5 date and time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import re
import matplotlib.pyplot as plt
attacks = pd.read_csv("input/attacks.csv",encoding='utf-8')
attacks.Activity.value_counts().head(10)
import src.helpers as helpers
# -
# ## Get month, and season for Dataframe
# +
attacks['Month']= attacks['Date'].map(helpers.get_month)
attacks['Season']= attacks['Month'].map(helpers.get_season)
# -
# ## Parse shark type and add total area attacks
# +
attacks['Shark type']= attacks.apply(helpers.fix_species_name, axis='columns')
attacks['Area Total']=attacks['Area'].groupby(attacks['Area']).transform('count')
# -
# ## Attacks by activity
attacks.Activity.value_counts()
# ## Parse activities and divide them by above surface (Kayak, surf ...) and below surface (swimming, diving ..)
attacks['activity_clean']= attacks['Activity'].map(helpers.clean_activity)
attacks.activity_clean.value_counts()
# ## Which activities have higher death rate ?
# +
by_activity = attacks.groupby(['Activity','Fatal (Y/N)']).size().reset_index(name='counts').sort_values(by='counts',ascending=False)
by_activity[by_activity['Fatal (Y/N)']=='Y'][:9]
# +
by_activity = attacks.groupby(['activity_clean','Fatal (Y/N)']).size().reset_index(name='counts').sort_values(by='counts',ascending=False)
by_activity[:5]
# +
labels = 'Survived','Dead',
below_surface =by_activity[ by_activity['activity_clean']=='below_surface'][:2]['counts']
fig1, ax1 = plt.subplots()
ax1.pie(below_surface, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
ax1.axis('equal') # Equal aspect ratio ensures that pie is drawn as a circle.
plt.title("Below Surface Fatal (Y/N)")
plt.show()
# +
labels = 'Survived','Dead',
below_surface =by_activity[ by_activity['activity_clean']=='above_surface'][:2]['counts']
fig1, ax1 = plt.subplots()
ax1.pie(below_surface, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
ax1.axis('equal') # Equal aspect ratio ensures that pie is drawn as a circle.
plt.title("Above Surface Fatal (Y/N)")
plt.show()
# -
attacks[attacks['Fatal (Y/N)']=='Y'][['Area']].value_counts()
attacks.groupby(['Area','activity_clean']).size().reset_index(name='counts').sort_values(by='counts',ascending=False)[:20]
# +
usa_attacks = helpers.get_data_by_country(attacks,'USA')
by_activity = usa_attacks.groupby(['activity_clean','Fatal (Y/N)','Season']).size().reset_index(name='counts').sort_values(by='counts',ascending=False)
by_activity
# -
# ## Most common Shark Species By Area
# +
by_shark_type= attacks.groupby(['Area','Area Total']).apply(helpers.count_species).reset_index().sort_values(by=["Area Total",'Shark type'],ascending=False)
by_shark_type.groupby(['Area']).apply(lambda df: df.loc[df['Shark type'].idxmax()] ).sort_values(by=["Area Total",'Shark type'],ascending=False)[:20]
# -
by_shark = attacks.groupby(['Shark type','Fatal (Y/N)']).size().reset_index(name='counts').sort_values(by='counts',ascending=False)
by_shark
by_shark[by_shark['Fatal (Y/N)']=='Y'][:20]
| data-wrangling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # SLU19 - k-Nearest Neighbours (kNN)
#
# In this notebook we will be covering the following:
#
# - k-Nearest Neighbours Algorithm
# - A Primer on Distance
# - Some considerations about kNN
# - Using kNN
#
# ## 1. k-Nearest Neighbours Algorithm
#
# k-Nearest Neighbours (or kNN) is a supervised learning algorithm, that is mainly used for classification, but can also be used for regression tasks.
#
# Its main advantages are that it is very simple to understand and implement (as you'll see here!) and that it can be used in a wide range of problems.
#
# Its main disadvantages are that it doesn't scale very well to large datasets and to high dimensional spaces (without some optimisations).
# ### 1.1 How does it work
#
# The main intuition behind the algorithm is that neighbors are similar to each other.
#
# For example, a cat is likely to be more similar to other cats than to a dog.
#
# And if we want to classify whether it's a cat or a dog based on some parameters (e.g., sharpness of claws and length of ears), we can take a look at a few most similar neighbors and see whether they are dogs or cats.
#
# If 4 out of 5 most similar animals are cats, it's very likely that this one is a cat too, right?
#
#
# 
# More formally, the principle behind k-nearest neighbour methods is to find a predefined number of training samples closest to the point we want to find a prediction for, and predict the label from these. The predefined number of samples is a user-defined constant `k`.
#
# In the example above, we decided to take a look on the 5 most similar animals, so `k` was 5.
#
# The assumption here is that if two points are similar, i.e, close together in the features space, then their labels should also be similar.
#
# We'll be more careful in the definition of similarity, but let's first begin with one more example on how can we use kNN for classification and another for regression.
# 
# In the **Classification with kNN** figure, we can see how kNN can be used for classification.
#
# We have a point for which we want to predict a class, in this case it's the yellow star. We start by finding the _k_ points in the training data that are the closest to the star: these are the k-nearest neighbours. Then, we select as the predicted class for the star, the most common class among the k-nearest neighbours.
#
# In this example above, if we use _k_ = _3_, the star's nearest neighbours are the two red circles and the blue square that are inside the smallest dashed circle. The most common class among the nearest neighbours is class B (red circles), so that is the class that we'll predict for the star.
#
# Can you figure out what would be the predicted class if we used _k_ = _7_?
# You might ask a reasonable question: what if k=4 and we have 2 red circles and 2 blue squares out of 4 nearest neighbors?
#
# Well, for binary classification we're usually trying to select odd values for k. It solves the problem completely: one of the classes will always appear at least 1 more time than the other one.
#
# But it's not only a case when k is even. For 3-classes classification (e.g. cats, dogs and parrots) we might have k=5, and out of 5 nearest neighbors there might be 2 cats, 2 dogs and 1 parrot.
#
# In this case the algorithms usually select one of 2 options:
#
# 1. Choose a random class (cat or dog)
# 2. Choose the one that has the lowest average distance: if 2 cats are nearer than 2 dogs, the label is cat.
#
# We're not going to implement this corner case, but keep it in mind.
# In the **Regression with kNN** figure, we can see how kNN can be used for regression.
#
# We have a point for which we know the x-value and want to predict the y-value (which is the star!). Again we need to find the k-nearest neighbours, and then, select as the predicted value, **the average y-value of the k-nearest neighbours.**
#
# In this example above, we know the star's x-value, which is 8, and we want to predict its y-value.
# If we use _k_ = _3_, the star's nearest neighbours (measured in the x-axis) are the three points within the shaded area.
# To get the predicted y-value for the star, we average the y-values of the nearest neighbours:
#
# $$\frac{2+4+5}{3} = 3.67$$
#
# Can you figure out what would be the predicted y-value for the star if we used _k_ = _5_?
# ## 2. A Primer on Distance
#
# As we mentioned before, in order to find the k-nearest neighbours of a point, we need to have a measure of similarity, in order to actually understand how "near" two points are.
#
# The most common way to handle this is to use a **distance function**, that gives us a numerical measurement of how far apart two points are. Once we have such a function, we can select the nearest neighbours of a certain point by finding the neighbours for which the distance is the smallest.
#
# In most of the cases, using a distance function to measure distances between data points requires all the features in a dataset to be numerical. So, in case you have any categorical variables, you will need to represent them as numbers (or drop them!), before measuring distances in your dataset.
#
# Let's see some examples.
# ### 2.1 Euclidean distance
#
# Remember when you were in high school and had to compute vector norms and distances between vectors? Cool, because you were using the Euclidean distance back then!
#
# Let's define it.
# **The one dimensional case**
#
# In the one dimensional case, we have two one-dimensional points **p** and **q**.
#
# $$d(\mathbf{p}, \mathbf{q}) = \sqrt{(q - p)^2} = |q - p|$$
# **The two dimensional case**
#
# In the two dimensional case, we have two two-dimensional points or vectors $\mathbf{p}$ and $\mathbf{q}$.
#
# $$d(\mathbf{p}, \mathbf{q}) = \sqrt{(q_1 - p_1)^2 + (q_2 - p_2)^2}$$
# **The n dimensional case**
#
# In the n dimensional case, we have two n-dimensional points or vectors $\mathbf{p}$ and $\mathbf{q}$.
#
# $$d(\mathbf{p}, \mathbf{q}) = \sqrt{(q_1 - p_1)^2 + (q_2 - p_2)^2 + ... + (q_n - p_n)^2} = \sqrt{ \sum_{i=1}^n (q_i - p_i)^2}$$
# The euclidean distance is a good choice when the features are more or less in the same range of values. Can you figure out why this is the case?
# ### 2.2 Dot product (*)
#
# The dot product between two n-dimensional vectors $\mathbf{u}$ and $\mathbf{v}$ is defined as
#
# $$\mathbf{u} \; . \mathbf{v} = \sum_{i=1}^n u_i v_i = u_1v_1 + u_2v_2 + ... + u_nv_n$$
#
# Given the angle formed by the two vectors, $\theta$, and the norms of the vectors $|.|$, we can also define the dot product between $\mathbf{u}$ and $\mathbf{v}$ as
#
# $$\mathbf{u} \; . \mathbf{v} = |\mathbf{u}| \; |\mathbf{v}| \; cos(\theta)$$
#
#
# In fact, this second definition makes it easier to understand how the dot product can be used as a distance.
# The dot product measures the projection of one vector into the other, which basically means that we're measuring the vectors' norms and how much the two vectors are pointing in the same direction.
#
# **Note:** $\theta$ is the angle between vectors $\mathbf{u}$ and $\mathbf{v}$, so $cos(\theta) = cos(\mathbf{u}, \mathbf{v})$
#
# Let's use the following image and consider some particular cases to get a better intuition on this.
#
# 
#
# This image shows a representation of the dot product between vectors $\mathbf{u}$ and $\mathbf{v}$.
#
# Consider the following cases:
#
# * $\theta = 0$:
# In this case, $cos(\theta) = 1 $, which means the two vectors are collinear. This is when $\mathbf{u} \; . \mathbf{v} = |\mathbf{u}| \; |\mathbf{v}|$ has the maximum value.
#
#
# * $0 < \theta < \frac{\pi}{2}$:
# In this case, $0 < cos(\theta) < 1$, meaning that $|\mathbf{u}| \; |\mathbf{v}|$ is multiplied by a number between 0 and 1, and it gets smaller. The wider the angle (or difference in direction) between the two vectors, the smaller the dot product gets.
#
#
# * $\theta = \frac{\pi}{2}$:
# In this case, $cos(\theta) = 0$, which means the two vectors are orthogonal. This is when $\mathbf{u} \; . \mathbf{v} = 0$.
#
#
# * $\frac{\pi}{2} < \theta < \pi $:
# In this case, $-1 < cos(\theta) < 0$, meaning that $|\mathbf{u}| \; |\mathbf{v}|$ is multiplied by a number between -1 and 0, and it gets smaller in absolute value, and negative. This means that the two vectors have started to point in very different directions. Again, the wider the angle between the two vectors, the smaller the dot product gets.
#
#
# * $\theta = \pi$:
# In this case, $cos(\theta) = -1$, which means the two vectors are parallel, but pointing in opposite directions. This is when $\mathbf{u} \; . \mathbf{v} = -|\mathbf{u}| \; |\mathbf{v}|$ has the minimum value.
# ### 2.3 Cosine distance (*)
#
# As we saw above, there is a relationship between the dot product and the cosine of two vectors:
#
# $$cos(\theta) = \frac{\mathbf{u} \; . \mathbf{v}}{|\mathbf{u}| \; |\mathbf{v}|}$$
#
# With the cosine, we are measuring how similar is the direction of the two vectors, and disregarding the vectors' norms.
#
# Now we just need to convert this similarity into a distance. Since the domain of the cosine function is $[-1, 1]$, we can do this in the following way:
#
# $$cos\_dist(\mathbf{u}, \mathbf{v}) = 1 - cos(\mathbf{u}, \mathbf{v})$$
#
# The cosine distance works very well in cases where the features have values in different ranges. This is because dividing the dot product by the norms of the vectors works as a kind of normalization!
# **(*) Note for the math nerds**
#
# Neither the dot product nor the cosine are distances, as per the mathematical definition of a [distance function](https://en.wikipedia.org/wiki/Distance#General_metric). Because of that, we cannot use some of scikit's optimisations that make kNN run faster. But that is a bit out of scope here!
# ## 3. Some considerations about kNN
#
# Now that we have some intuition on how kNN works, and we've seen some functions that can be used as a distance (i.e, a measure of similarity), let's go through some considerations about this algorithm.
#
#
# ### Non-parametric
#
# kNN is a non-parametric model because its structure is not specified a priori but is instead determined from the data.
#
# To better understand what this means, we can think of a counter example: Linear Regression, which is a parametric model, assumes that the data follows a linear distribution.
#
#
# ### No learning
#
# When we described how does kNN work, you may have noticed a key difference between this algorithm and other algorithms that you've seen before, like Linear Regression or Logistic Regression: in kNN we don't actually learn anything!
#
# Taking Linear Regression as an example, in the training phase, we used training data to learn some parameters ($\beta$) that were later used in the prediction phase to make predictions on unseen data.
#
# In kNN we don't learn any parameters, and in the training phase we don't do more than just loading the training dataset into memory. Instead, most of the action takes place at prediction time, when we determine the nearest neighbours (using training data), and make predictions based on them. This is why we say that kNN is a **lazy** method.
#
#
# ### How to chose the value of _k_
#
# The optimal choice of the value *k* is highly data-dependent: in general a larger _k_ suppresses the effects of noise, but making it too large results in more prediction errors.
#
# In `SLU15 - Hyperparameter Tuning`, we'll learn how to systematically find the best value for _k_.
#
#
# ### kNN and high dimensional spaces
#
# When we increase the number of features in our model, we need more dimensions to represent the data points.
#
# The problem with high dimensional spaces is that the data gets very sparse, and consequently, points tend not to be close to each other. In particular, the k-nearest neighbours of a point won't be much closer to it than any other random points, which breaks the algorithm's assumption that points that are close are also similar.
#
# This phenomenon, called **curse of dimensionality**, is very well explained with an example [here](http://www.cs.cornell.edu/courses/cs4780/2018fa/lectures/lecturenote02_kNN.html).
#
# This problem exists when we use the euclidean distance and the dot product, as these two distances measure amplitudes.
# The same doesn't happen with the cosine distance, that just considers directions.
#
# Another way to avoid this is to use dimensionality reduction techniques, in order to simplify our data. This won't be covered in this course.
#
#
# ### kNN and large datasets
#
# Given a dataset with _N_ training points, when we try to get a prediction for a certain point, we need to load the entire dataset into memory and compute the distance between this point and all the other points.
#
# This means that the time that it takes to yield a prediction, depends on the dataset size. In fact, it grows linearly with it!
#
# Given these considerations, it's easy to understand that kNN is not the best choice when the dataset is very large.
#
# There are some ways to make kNN run faster, but these are out of the scope of this SLU!
#
# ### Theoretically perfect
#
# K-nearest neighbors is a well-studied approach. There are many important theorems claiming that, on "endless" datasets, it is the optimal method of classification.
#
# The authors of the classic book "The Elements of Statistical Learning" consider kNN to be a theoretically ideal algorithm which usage is only limited by computation power and the curse of dimensionality.
#
# ### kNN in the real world
#
# kNN can serve as a good starting approach (baseline) in some cases.
#
# kNN might also be used in recommendation systems. The initial approach might be to recommend a product that is popular among K most similar people to a client.
#
# On Kaggle competitions, kNN is often used as a part of more complicated models that combine different approaches (those methods are called stacking and blending, but they are out of this course scope)
# ## 4. Using kNN
#
# Let's now see how can we use kNN in classification and regression problems.
#
# Let's start with the usual imports.
# +
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# ### 4.1 Classification on the iris dataset
#
# We'll use kNN to solve the iris classification problem.
#
# The [iris flower dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set) is a well known dataset for classification.
#
# In the dataset, each row (or observation) represents an iris flower. The features that describe the flower are _sepal length,_ _sepal width,_ _petal length_ and _petal width._
#
# The goal is to predict the iris' type, that can be one of _setosa,_ _versicolor_ and _virginica._
# 
# Here we're doing a bunch of imports:
#
# * scikit's datasets: this is a package that allows up to load the iris dataset
# * accuracy score: this is to evaluate our classification model
# * train_test_split: this is to split out dataset into training and testing sets
# * **KNeighborsClassifier**: this is our kNN classifier
from sklearn import datasets
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
# We'll start by loading the iris dataset and then split the dataset into training and testing.
# +
# Loading the iris dataset
iris = datasets.load_iris()
# Splitting the data into train and test
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size=0.33, random_state=42)
print(f"Train data size: {len(X_train)}\nTest data size: {len(X_test)}")
# -
# We'll create a DataFrame with the features and target in the training set, just to quickly check their values and check their ranges with a boxplot.
df_train = pd.DataFrame(X_train, columns=iris['feature_names'])
df_train['target'] = y_train
df_train.head()
df_train.boxplot(vert=False);
plt.xlabel("Values (cm)");
plt.ylabel("Features");
plt.title("Iris feature value's analysis");
# Now, our goal is to predict labels for the data points in the testing set.
#
# First, let's get a baseline, which is the accuracy of the simplest model we can think of. Our model must be better than that!
#
# The simplest model is to always predict the most frequent class. So let's see how to do that.
# +
test_size = len(y_test)
most_common_target = df_train.target.value_counts(sort=True).index[0]
y_pred = np.ones(test_size) * most_common_target
y_pred
# -
accuracy_score(y_test, y_pred)
# So, we need to improve upon a 0.3 accuracy score. Let's see if we can do it with our kNN classifier.
#
# For each point in the testing set, kNN will search for the k nearest neighbours in the training set, and predict the most frequent label among the the neighbours.
#
# For now let's use the default value for k, which is 5.
clf = KNeighborsClassifier(n_neighbors=5)
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
accuracy_score(y_test, y_pred)
# 0.98 accuracy! That's way better than our baseline. kNN did a good job :)
# ### 4.2 Regression on the diabetes dataset
# For the example with regression, we'll use the [diabetes dataset](https://scikit-learn.org/stable/datasets/toy_dataset.html#diabetes-dataset).
#
# Each data point represents one person who has diabetes. The features have information like the person's age, sex, body mass index, and other health metrics. The target is a quantitative measure of disease progression one year after a certain baseline.
#
# Our goal is to predict this quantitative measure for unseen data.
# We'll start with some additional imports:
#
# * **KNeighborsRegressor**: this is our kNN regression model
# * mean_squared_error: this is to evaluate our model
# * scipy's cosine: this is for us to try the cosine distance in the kNN algorithm
from sklearn.neighbors import KNeighborsRegressor
from sklearn.metrics import mean_squared_error
from scipy.spatial.distance import cosine as cos_dist
# As we did before, we'll load the dataset from scikit's datasets and then, do a train and test split.
diabetes = datasets.load_diabetes()
X_train, X_test, y_train, y_test = train_test_split(diabetes.data, diabetes.target, test_size=0.33, random_state=42)
print(f"Train data size: {len(X_train)}\nTest data size: {len(X_test)}")
# Here we're loading the features into a DataFrame in order to quickly visualise them with the help of a boxplot.
df_X_train = pd.DataFrame(X_train, columns=diabetes['feature_names'])
df_X_train.head()
df_X_train.boxplot(vert=False);
plt.xlabel("Values");
plt.ylabel("Features");
plt.title("Diabetes feature value's analysis");
# Again, let's get a baseline. In this case, we'll always predict the mean value of the targets in the training set.
#
# We'll be using the mean_squared_error to evaluate our model.
# baseline with predicting the average
y_pred = np.ones(len(y_test)) * y_train.mean()
mean_squared_error(y_test, y_pred)
# Now let's get the predictions using the kNN regressor. We'll use k=5, which is the default value, as a starting point.
reg = KNeighborsRegressor()
reg.fit(X_train, y_train)
y_pred = reg.predict(X_test)
mean_squared_error(y_test, y_pred)
# This was a big improvement upon our baseline! Let's see if we can still do better, by choosing a different value for k.
#
# We'll learn how to do hyperparameter tuning properly in a later SLU, but for now let's go with this method.
#
# Plotting values of k vs the mean squared error will show us how does the error change with k. Then, we can select the value of k that minimises the error.
# +
# this list will save the different values for k and mean squared error
k_rmse = []
# for each value of k between 1 and 100, we'll compute the mean squared error
# and save it in the k_rmse list like [(k1, error1), (k2, error2), ..., (k100, error100)]
for k in range(1, 100):
reg = KNeighborsRegressor(k)
reg.fit(X_train, y_train)
y_pred = reg.predict(X_test)
k_rmse.append((k, mean_squared_error(y_test, y_pred)))
# here we're separating the k values from the error values into two lists
k_values, rmse_values = zip(*k_rmse)
# and here we're plotting k vs the error
plt.plot(k_values, rmse_values);
plt.xlabel('k');
plt.ylabel('mean squared error');
plt.title('Mean squared error vs number of nearest neighbours (k)');
# -
# We can see that at first, the value of the error decreases a lot by adding more neighbours, and then we reach a point, around k=20, where by adding more neighbours we actually make the error worse!
#
# So let's stick to `k=20` and see what's our mean squared error.
reg = KNeighborsRegressor(n_neighbors=20)
reg.fit(X_train, y_train)
y_pred = reg.predict(X_test)
mean_squared_error(y_test, y_pred)
# We still managed to improve a bit from our first attempt with k=5.
#
# Let's finish by checking how can we use the cosine distance instead of the euclidean distance (which is the default in scikit).
reg = KNeighborsRegressor(metric=cos_dist)
reg.fit(X_train, y_train)
y_pred = reg.predict(X_test)
mean_squared_error(y_test, y_pred)
# The mean squared error decreases slightly, but not significantly.
# As we saw before, the cosine distance tends to work better than the euclidean distance when we have a lot of features (because of the curse of dimensionality) or when the features are within very different ranges.
#
# Neither of these two situations verifies here, so there is no great benefit in using the cosine distance.
#
# In fact, using the cosine distance is probably worse, as we'll not benefit from scikit's built in optimisations for kNN. If you try to find the best k in this scenario, using the method that we used above, you'll see that it will run much slower than with the euclidean distance.
| S01 - Bootcamp and Binary Classification/SLU19 - k-Nearest Neighbors (kNN)/Learning notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from spellchecker import SpellChecker
import re
spell = SpellChecker()
text_file = open("Downloads/stopwords.txt", "r")
stopwords = text_file.read().split('\n')
df = pd.read_csv('Downloads/spell_corrected_industry_counts.csv')
df.columns = ['Industry','count']
custom_stop = ["agency","animal","autobody","automobile","beverage","bpo","business","call","camp","canadian","car","center","centre","commercial","community","control","course","culture","customer","development","employed","employment","equipment","fire","firm","foods","glass","group","home","house","housing","human","immigration","industry","information","interior","keeping","living","local","long","maintenance","natural","non","none","office","organization","pet","private","processing","production","protection","real","safety","sands","save","sector","self","senior","service","services","social","support","temp","term","tire","trade","trades","trading","various","video","windows","work","worker","works"]
stopwords.extend(custom_stop)
df_new = df[df['Industry'].notnull()]
df_new['Processed'] = 'hello'
df_new['combined'] = False
print('a')
for index, row in df_new.iterrows():
x = re.findall(r"[\w']+", row['Industry'])
x = [spell.correction(w) for w in x]
df_new.at[index, 'Processed'] = ' '.join(x)
top_entries = list(df_new.iloc[0:500,0])
print('finihsed')
# +
df_new['new_count']=0
df_new['match'] = "NA"
df_new['association'] = "NA"
n=0
import warnings
warnings.filterwarnings('ignore')
for index, row in df_new.iloc[501:,].iterrows():
n=n+1
x = row['Processed'].split()
x = [w for w in x if not w in stopwords]
for xx in x:
if row['combined'] == False:
for tt in top_entries:
t = tt.split()
t = [w for w in t if not w in stopwords]
for z in t:
if xx == z:
if row['combined'] == False:
row['combined'] = True
df_new['new_count'][df_new.Industry == tt] = df_new['new_count'][df_new.Industry == tt] + row['count']
df_new['combined'][df_new.Industry == row['Industry']] = True
df_new['count'][df_new.Industry == row['Industry']] = 0
df_new['match'][df_new.Industry == row['Industry']] = xx
df_new['association'][df_new.Industry == row['Industry']] = tt
if n < 40:
print('we got a match')
print(n)
print(xx)
print(row)
print(df_new[df_new.Industry == z])
print(row['count'])
print('finished')
# -
df_new.head()
df_new['total'] = df_new['count'] + df_new['new_count']
df_new.to_csv('Downloads/second_pass_at_industry_groupings_w_match_and_association.csv')
df_new['match'].value_counts().to_csv('Downloads/second_industry_matches_count.csv')
| notebooks/industry grouping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import seaborn as sns
iris = sns.load_dataset('iris')
iris.shape
iris = pd.DataFrame(iris, columns=iris.columns)
iris.head()
iris['sepal_length'].value_counts().hist(bins=10)
species = {'setosa': 0, 'versicolor': 1}
iris['species_number'] = iris['species'].apply(lambda x: species.get(x, 2))
iris['species_number'].value_counts().plot(kind='bar')
len(iris)
from sklearn.neighbors import KNeighborsClassifier
help(KNeighborsClassifier)
model = KNeighborsClassifier(n_neighbors=3)
X = iris[['sepal_length', 'sepal_width', 'petal_length', 'petal_width']]
Y = iris['species']
model.fit(X, Y)
desconhecido = pd.DataFrame([[5.1, 3.5, 1.4, 0.2]])
model.predict(desconhecido)
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split?
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size=0.3)
model = KNeighborsClassifier(n_neighbors=3)
model.fit(x_train, y_train)
model.score(x_test, y_test)
| notebooks/iris.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import LinearSVC
from sklearn.pipeline import make_pipeline
from sklearn.metrics import confusion_matrix
# -
from spectra_analysis.preprocessing import read_spectra
from spectra_analysis.plotting import plot_spectra, plot_spectra_by_type, plot_cm, plot_regression
from spectra_analysis.regression import regression_experiment, fit_params, transform
# ### IO: Reading and preprocess the data
# We can define a function which will read the data and process them.
# +
# read the frequency and get a pandas serie
frequency = pd.read_csv('data/freq.csv')['freqs']
# read all data for training
filenames = ['data/spectra_{}.csv'.format(i)
for i in range(4)]
spectra, concentration, molecule = [], [], []
for filename in filenames:
spectra_file, concentration_file, molecule_file = read_spectra(filename)
spectra.append(spectra_file)
concentration.append(concentration_file)
molecule.append(molecule_file)
# Concatenate in single DataFrame and Serie
spectra = pd.concat(spectra)
concentration = pd.concat(concentration)
molecule = pd.concat(molecule)
# -
# ### Plot helper functions
# We can create two functions: (i) to plot all spectra and (ii) plot the mean spectra with the std intervals.
# We will make a "private" function which will be used by both plot types.
fig, ax = plot_spectra(frequency, spectra, 'All training spectra')
fig, ax = plot_spectra_by_type(frequency, spectra, molecule)
ax.set_title('Mean spectra in function of the molecules')
fig, ax = plot_spectra_by_type(frequency, spectra, concentration,
'Mean spectra in function of the concentrations')
# ### Reusability for new data:
# +
spectra_test, concentration_test, molecule_test = read_spectra('data/spectra_4.csv')
plot_spectra(frequency, spectra_test,
'All training spectra')
plot_spectra_by_type(frequency, spectra_test, molecule_test,
'Mean spectra in function of the molecules')
plot_spectra_by_type(frequency, spectra_test, concentration_test,
'Mean spectra in function of the concentrations');
# -
# ## Training and testing a machine learning model for classification
for clf in [RandomForestClassifier(random_state=0),
LinearSVC(random_state=0)]:
pipeline = make_pipeline(StandardScaler(),
PCA(n_components=100, random_state=0),
clf)
y_pred = pipeline.fit(spectra, molecule).predict(spectra_test)
fig, ax = plot_cm(
confusion_matrix(molecule_test, y_pred),
pipeline.classes_,
'Confusion matrix using {}'.format(clf.__class__.__name__))
print('Accuracy score: {0:.2f}'.format(pipeline.score(spectra_test,
molecule_test)))
# ## Training and testing a machine learning model for regression
regression_experiment(spectra, spectra_test,
concentration, concentration_test)
# compute the statistics on the training data
med, var = fit_params(spectra)
# transform the training and testing data
spectra_scaled = transform(spectra, med, var)
spectra_test_scaled = transform(spectra_test, med, var)
regression_experiment(spectra_scaled, spectra_test_scaled,
concentration, concentration_test)
| Day_2_Software_engineering_best_practices/solutions/04_modules/04_modules.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Data
import numpy as np
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
import datetime as dt
import matplotlib.pyplot as plt
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
df_1 = pd.read_csv('banknifty.csv')
# ## Preprocessing Data
def Date_Time(dataFrame):
dateTime = dataFrame['date'].map(str)+dataFrame['time']
k = pd.to_datetime(dateTime, format='%Y%m%d%H:%M')
dataFrame['DateTime'] = k
dataFrame['Day'] = dataFrame['DateTime'].dt.day
dataFrame['Month'] = dataFrame['DateTime'].dt.month
dataFrame['Year'] = dataFrame['DateTime'].dt.year
#dataFrame['Hour'] = dataFrame['DateTime'].dt.hour
#dataFrame['Minute'] = dataFrame['DateTime'].dt.minute
dataFrame = dataFrame.drop(labels=['DateTime'], axis=1)
dataFrame['group']= dataFrame['Year'].map(str) + dataFrame['Month'].map(str)+ dataFrame['Day'].map(str)
dataFrame = dataFrame[['open', 'high', 'low', 'Day', 'Month', 'Year','group','close']]
dataFrame= dataFrame.sort_values(by=['Year','Month','Day'])
dataFrame= dataFrame.reset_index(drop=True)
return(dataFrame)
def processing(dataframe):
df = dataframe
day_group = df['group'].unique() # extract unique hour values to form group based on days, month and year
d_group_index = np.arange(1,len(day_group)+1)# for reindexing hour group values from 1 to number of groups.
#As indexing starts from 0 so 1 is added
# replacing hour group values with new indexing for extracting hour groups
#(This step will take 20 minutes due to 3 hundred thousand samples)
# it is already done once and results are saved in file hour.npy
# so instead of running again, load this file
for i in range(len(day_group)):
df['group'] = df['group'].replace([day_group[i]],d_group_index[i])
df1 = pd.DataFrame(df, index= day_group) # this data frame has day group as index values for extracting its index
count_index = df['close'].groupby(df['group']).count() # counting each day group values
day_index = [] # extracting months index
w=0
for i in count_index:
w = i+w
day_index.append(w)
day_index = np.array(day_index) -1
# above steps are adding count values(in other words "commulative count_index")
# we need commulative count_index as count_index are absolute values from which required values cant be extracted
# extracting close values which is last value of each month group
close = []
for i in day_index:
t = df.loc[i,'close']
close.append(t)
close = np.array(close)
#extracting low, high, month, year values of each month group
low = pd.DataFrame(df['low'].groupby(df['group']).min()).reset_index(drop=True)
high = pd.DataFrame(df['high'].groupby(df['group']).max()).reset_index(drop=True)
Day = pd.DataFrame(df['Day'].groupby(df['group']).max()).reset_index(drop=True)
Month = pd.DataFrame(df['Month'].groupby(df['group']).max()).reset_index(drop=True)
Year = pd.DataFrame(df['Year'].groupby(df['group']).max()).reset_index(drop=True)
#extracting first value of open from each month group
openn = []
for i in (day_index-count_index+1):
r = df.loc[i,'open']
openn.append(r)
openn = np.array(openn)
#creating new data frame with extracted values
df2 = pd.DataFrame()
df2['open'] = openn
df2['high'] = high
df2['low'] = low
df2['Day'] = Day
df2['Month'] = Month
df2['Year'] = Year
df2['close'] = close
# rearranging data into ascending form
df2 = df2.sort_values(by=['Year','Month','Day'])
df2 = df2.reset_index(drop=True) # reset index
return(df2)
def scaling(dataFrame):
close = np.array(dataFrame['close']).reshape(-1,1)
stock_df = dataFrame.drop(labels=['close'], axis = 1)
scaler = MinMaxScaler(feature_range=(0,1))
scaler.fit(stock_df)
scaled_df = scaler.transform(stock_df)
scaler2 = MinMaxScaler(feature_range=(0,1))
scaler2.fit(close)
scaled_close = scaler2.transform(close)
scaled_df = pd.DataFrame(scaled_df, columns=stock_df.columns)
scaled_df['close'] = scaled_close
return(scaled_df, scaler, scaler2)
stock_df1 = Date_Time(df_1)
stock_df1.head()
stock_df1_1 = processing(stock_df1)
stock_df1_1
stock_df2, in_scaler, out_scaler = scaling(stock_df1_1)
stock_df2.head()
# ## Data Plots
import seaborn as sns
# #### Time Series Distribution For Month
sns.set(rc={'figure.figsize':(11,4)})
stock_df1_1[['open','high','low']].plot(linewidth=0.8, title='Days Series')
plt.xlabel('Days (2012-2016)')
plt.ylabel('Stock Rate')
cols_plot = ['open', 'high','low']
axes = stock_df1_1[cols_plot].plot(marker='o', alpha=0.8, linestyle='-', figsize=(11, 9), subplots=True)
for ax in axes:
ax.set_ylabel('Stock Rate')
ax.set_xlabel('Days (2012-2016)')
# #### Box Pots
fig, axes = plt.subplots(3, 1, figsize=(12, 10), sharex=True)
for name, ax in zip(['open', 'high', 'low'], axes):
sns.boxplot(data=stock_df1_1, x='Day', y=name, ax=ax)
ax.set_ylabel('Stock Rate')
ax.set_title(name)
# #### Heat Map
sns.heatmap(stock_df1_1[['open','high','low']])
# #### Histograms and Curve Distribution
fig, axes = plt.subplots(1,3, figsize=(15,5))
for name, ax in zip(['open', 'high', 'low'], axes):
sns.distplot(stock_df1_1[name], ax=ax)
# #### Correlation
plt.matshow(stock_df1_1.corr())
plt.show()
# #### Scatter Plot
plt.scatter(stock_df1_1['Day'],stock_df1_1['open'])
plt.scatter(stock_df1_1['Day'],stock_df1_1['high'])
plt.scatter(stock_df1_1['Day'],stock_df1_1['low'])
plt.legend(['Open','High','Low'])
plt.xlabel('Dayss')
plt.ylabel('Stock Rate')
plt.show()
plt.plot(stock_df1_1['open'].rolling(window=150, center=True, min_periods=30).mean())
plt.plot(stock_df1_1['high'].rolling(window=150, center=True, min_periods=30).mean())
plt.plot(stock_df1_1['low'].rolling(window=150, center=True, min_periods=30).mean())
plt.legend(['Open','High','Low'])
plt.title('Trend Line')
plt.xlabel('Days')
plt.boxplot(stock_df2['close'], showmeans=True)
plt.show()
# ## Splitting Data into Train/Test
def train_test_data(data):
x = np.array(data.iloc[:,:-1])
y = np.array(data.iloc[:,-1])
x_train, x_test, y_train, y_test = train_test_split(x,y, test_size = 0.2, shuffle= True)
return (x_train, x_test, y_train, y_test)
x_train_df1, x_test_df1, y_train_df1, y_test_df1 = train_test_data(stock_df2)
x_train_df1.shape
y_train_df1.shape
# ## Random Forest
from sklearn.ensemble import RandomForestRegressor
rf = RandomForestRegressor(verbose=1)
rf.fit(x_train_df1, y_train_df1)
pred = rf.predict(x_test_df1)
# Output value is scaled. To get actual value undo scaled value of output
print('Scaled Value Predicted: %.2f' %pred[2])
print('Actual Predicted Value: %.2f'%out_scaler.inverse_transform([[pred[2]]]))
print('True Value: %.2f' %out_scaler.inverse_transform([[y_test_df1[2]]]))
# ### Random Forest Evaluation
print('R_2 Score: %.7f' %r2_score(y_test_df1, pred))
print('Mean Absolute Error: %.7f' %mean_absolute_error(y_test_df1, pred))
print('Mean Square Error: %.7f' %mean_squared_error(y_test_df1, pred))
print('Root Mean Square Error: %.7f' %np.sqrt(mean_squared_error(y_test_df1, pred)))
# ### Random Forest Plots
plt.plot(y_test_df1, 'b')
plt.plot( pred, 'y')
plt.legend(['True', 'Pred'])
plt.title('Predicted vs True')
plt.show()
# +
plt.figure(figsize=(12,4))
plt.subplot(1,2,1)
plt.hist(y_test_df1)
plt.title('True')
plt.subplot(1,2,2)
plt.hist(pred, color='grey')
plt.title('Predicted')
plt.show()
# -
plt.boxplot(pred, showmeans=True)
plt.show()
# ## Time Based Prediction Plot
true = out_scaler.inverse_transform([y_test_df1]) # true values without scaling
result = out_scaler.inverse_transform([pred]) # predicted values without scaling
tm = in_scaler.inverse_transform(x_test_df1) # descaling test input values
tm1 = tm[:,3] # descaling Test set Day values
tm2 = tm[:,4] # descaling Test set Month values
tm3 = tm[:,5] # descaling Test set Year values
# Result DataFrame without scaled values for true and predicted
res = pd.DataFrame()
res['Day'] = tm1.astype(int)
res['Month'] = tm2.astype(int)
res['Year'] = tm3.astype(int)
res['True'] = np.squeeze(true)
res['Predicted'] = np.squeeze(result)
res['Time'] = res['Day'].map(str) +'-' +res['Month'].map(str) +'-' + res['Year'].map(str)
# Sorting result values according to time in ascending order and resetting index
res1 = res.sort_values(by=['Year','Month','Day'])
res1 = res1.reset_index(drop=True)
res1.head()
# Complete Test Set Plot
plt.plot(res1['True'], 'b')
plt.plot( res1['Predicted'], 'y')
plt.legend(['True', 'Pred'])
plt.title('Predicted vs True')
plt.ylabel('Stock Value')
plt.xlabel('Time (Day-Month-Year)')
plt.show()
# First 40 Test Set Points Plot
plt.plot(res1.loc[:40,'True'], 'b')
plt.plot( res1.loc[:40,'Predicted'], 'y')
plt.legend(['True', 'Pred'])
plt.title('Predicted vs True')
# for displaying day time x-axis labels uncomment the next line
plt.xticks(ticks=np.arange(40), labels=res1.loc[:40,'Time'], rotation = 45)
plt.ylabel('Stock Value')
plt.xlabel('Time (Day-Month-Year)')
plt.show()
# ## User Input Data
def user():
u = input('Do you want to enter custom data sample? Enter "y" or "n" :')
if u == 'y':
print('\nStock value range is 1400-21000')
openn = float(input('Enter Opening Stock Value of the day: '))
high = float(input('Enter Highest Stock Value of the day: '))
low = float(input('Enter Lowest Stock Value of the day: '))
date = str(input('Enter Date and time in format (21/03/201914:02) : '))
new_data = np.array([[openn, high, low, date]])
else:
new_data = None
return new_data
def u_output(u_dt):
user_df = pd.DataFrame(u_dt, columns=['Open','High', 'Low', 'Date'])
user_dt = pd.to_datetime(user_df['Date'], format='%d/%m/%Y%H:%M')
user_df2 = user_df.drop(labels=['Date'], axis=1)
user_df2['Day'] = user_dt.dt.day
user_df2['Month'] = user_dt.dt.month
user_df2['Year'] = user_dt.dt.year
sca_user_df1 = in_scaler.transform(user_df2)
user_df1 = pd.DataFrame(sca_user_df1, columns=user_df2.columns)
u_pred = rf.predict(np.array(user_df1))
u_real1 = out_scaler.inverse_transform([u_pred])
return(u_real1)
user_data = user()
if user_data is None:
print('Process Completed')
else:
o1 = u_output(user_data)
print('Stock Closing is at: %.2f'%o1)
| Stock_RF_day_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#from yahoo_finance import Share
# changed to https://github.com/cgoldberg/ystockquote
# changed to yfinance
import yfinance as yf
import ystockquote
from matplotlib import pyplot as plt
import numpy as np
import tensorflow as tf
import random
import pandas_datareader.data as web
import pandas as pd
def get_prices(share_symbol, start_date, end_date,
cache_filename='../data/cache/stock_prices.npy', force=False):
try:
if force:
raise IOError
else:
stock_prices = np.load(cache_filename)
except IOError:
#share = Share(share_symbol)
#stock_hist = ystockquote.get_historical_prices(share_symbol, start_date, end_date)
stock_hist = yf.download(share_symbol, start=start_date, end=end_date)
#stock_hist = share.get_historical(start_date, end_date)
stock_prices = []
stock_hist = stock_hist.sort_values(by=['Date'])
for index, day in stock_hist.iterrows():
stock_val = day['Open'].astype(float)
stock_prices.append(stock_val)
stock_prices = np.asarray(stock_prices)
#stock_prices = [stock_price['Open'] for stock_price in stock_hist]
np.save(cache_filename, stock_prices)
return stock_prices.astype(float)
def plot_prices(prices):
plt.title('Opening stock prices')
plt.xlabel('day')
plt.ylabel('price ($)')
plt.plot(prices)
plt.savefig('../data/cache/prices.png')
plt.show()
prices = get_prices('MSFT', '1992-07-22', '2016-07-22', force=True)
plot_prices(prices)
df = web.get_data_yahoo("MSFT", '1992-07-22', '2016-07-22')
new_df = df.drop(['High', 'Low', 'Close', 'Volume', 'Adj Close'], axis=1)
plot_prices(new_df.values)
# does not look like the one in the book
| TFv2/ch13/Listing 13.01 - 13.04.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# +
#install.packages("geojsonio", lib="/usr/local/lib/R/site-library/")
# + [markdown] tags=[]
# # Data needs to be under `/shared_volume/`. Is located in `hsi-kale` bucket
# +
#loads
library(hsi)
library(magrittr)
library(raster)
library(usethis)
library(devtools)
library(rgdal)
# -
## Localidades en shapefile de la especies con los anios
poncaloc<-rgdal::readOGR("/shared_volume/Ponca_DV_loc/","poncadav2")
#Se reproyectan las localidades
poncaloc<-sp::spTransform(poncaloc,
CRSobj = "+proj=lcc +lat_1=17.5 +lat_2=29.5 +lat_0=12 +lon_0=-102 +x_0=2500000 +y_0=0 +datum=WGS84 +units=m +no_defs +ellps=WGS84 +towgs84=0,0,0")
colnames(poncaloc@coords)
#Información general de las localidades
print(poncaloc)
plot(poncaloc)
ponca_mask <- raster::raster("/shared_volume/Ponca_DV/poncamask.tif")
print(ponca_mask)
plot(ponca_mask)
plot(poncaloc,add=T)
#histograma de frecuencia, reclasificacion de anios al periodo de reproyeccion
test_sp<-sp_temporal_data(occs=poncaloc,longitude = "coords.x1",
latitude = "coords.x2",sp_year_var="Year",
layers_by_year_dir ="/shared_volume/forest_jEquihua_mar/",
layers_ext = "*.tif$",reclass_year_data = T)
#Filtrar las localidades que se usaran mediante la mascara
test_sp_mask <- occs_filter_by_mask(test_sp,ponca_mask)
#Limpia localidades duplicadas por anio
test_sp_clean <- clean_dup_by_year(this_species = test_sp,
threshold = res(ponca_mask)[1])
# +
#save(test_sp_clean,file = "/Users/Mariana/Google Drive/andonue/Ponca_DV/new_model/test_ponca_clean.RData")
#Obtiene los valores ambientales mediante un extract de los puntos ocn las variables ambientales por año
# -
e_test<-extract_by_year(this_species=test_sp_clean,layers_pattern="_mar")
e_test
# +
#save(e_test,file = "/shared_volume/e_puma_test.RData")
# +
#next is not working:
#best_model_2004<-find_best_model(this_species =e_test,
# cor_threshold = 0.8,
# ellipsoid_level = 0.975,
# nvars_to_fit = 3,E = 0.05,
# RandomPercent = 70,
# NoOfIteration = 1000,
# parallel = TRUE, #Run process in parallel
# n_cores = 2,
# plot3d = FALSE)
# + [markdown] tags=[]
# ## Debugging
# -
library(hsi)
library(magrittr)
library(raster)
library(usethis)
library(devtools)
library(rgdal)
load(file ="/shared_volume/e_puma_test.RData" )
stopifnot(inherits(e_test, "sp.temporal.env"))
n_nas <- floor(dim(e_test$env_data_train)[1]*0.1)
n_nas
env_train <- e_test$env_data_train
rm_layers <- unlist(sapply( 1:dim(env_train)[2], function(x){
if(length(which(is.na(env_train[,x]))) > n_nas) return(x)
} ))
if(!is.null(rm_layers)){
env_train <- stats::na.omit(env_train[,-rm_layers])
}
numericIDs <- which(sapply(env_train, is.numeric))
cor_matrix <- stats::cor(env_train[,numericIDs])
cor_threshold=0.9
ellipsoid_level=0.975
nvars_to_fit=3
plot3d=FALSE
E = 0.05
RandomPercent = 50
NoOfIteration=1000
parallel=FALSE
n_cores=4
find_cor <- correlation_finder(cor_mat = cor_matrix,
threshold = cor_threshold,
verbose = F)
cor_filter <- find_cor$descriptors
cor_filter
combinatoria_vars <- combn(length(cor_filter),nvars_to_fit)
combinatoria_vars
# +
year_to_search <- min(as.numeric(names(e_test$layers_path_by_year)))
# -
year_to_search
# +
cat("The total number of models to be tested are: ", dim(combinatoria_vars)[2],"...\n\n")
env_layers <- raster::stack(e_test$layers_path_by_year[[paste0(year_to_search)]])
# -
env_layers
combinatoria_vars
this_species <- e_test
#cell to test if sprintf is working per iteration
mifun <- function(x){
cat(sprintf("Doing model: %d of %d \n", x , dim(combinatoria_vars)[2]))
# Varaibles filtadas por combinatiria de las mas representativas
vars_model <- cor_filter[combinatoria_vars[,x]]
ellip <- try(cov_center(env_train[,vars_model],
level = ellipsoid_level ,vars = vars_model),silent = T)
if(class(ellip)=="try-error") return()
# Datos de presencia de la sp en el ambiente
occs_env <- this_species$env_data_train[,vars_model]
# Ajuste del modelo de elipsoide
sp_model <- ellipsoidfit(data = env_layers[[vars_model]],
centroid =ellip$centroid,
covar = ellip$covariance,
level = ellipsoid_level,
size = 3,
plot = plot3d)
valData <- this_species$test_data[,c(1,2)]
valData$sp_name <- "sp"
valData <- valData[,c(3,1,2)]
p_roc<- PartialROC(valData = valData,
PredictionFile = sp_model$suitRaster,
E = E,
RandomPercent = RandomPercent,
NoOfIteration = NoOfIteration)
p_roc$auc_pmodel <- paste0(x)
return(list(model = sp_model$suitRaster,
pRoc=p_roc[,c("auc_ratio","auc_pmodel")],
metadata=ellip))
}
l <- list()
for (i in 1:2){
print("hola")
print(i)
l[[i]] <- mifun(i)
}
mifun2 <- function(x){
cat(sprintf("Doing model: %d of %d \n", x , dim(combinatoria_vars)[2]))
}
for (i in 1:10){
mifun2(i)
x <- 1
}
modelos <- lapply(1:50,function(x){
cat(sprintf("Doing model: %d of %d \n", x , dim(combinatoria_vars)[2]))
print("Varaibles filtadas por combinatiria de las mas representativas")
vars_model <- cor_filter[combinatoria_vars[,x]]
ellip <- try(cov_center(env_train[,vars_model],
level = ellipsoid_level ,vars = vars_model),silent = T)
if(class(ellip)=="try-error") return("what to return?")
print("Datos de presencia de la sp en el ambiente")
occs_env <- this_species$env_data_train[,vars_model]
print("Ajuste del modelo de elipsoide")
sp_model <- ellipsoidfit(data = env_layers[[vars_model]],
centroid =ellip$centroid,
covar = ellip$covariance,
level = ellipsoid_level,
size = 3,
plot = plot3d)
valData <- this_species$test_data[,c(1,2)]
valData$sp_name <- "sp"
valData <- valData[,c(3,1,2)]
print("PartialROC")
p_roc<- PartialROC(valData = valData,
PredictionFile = sp_model$suitRaster,
E = E,
RandomPercent = RandomPercent,
NoOfIteration = NoOfIteration)
p_roc$auc_pmodel <- paste0(x)
return(list(model = sp_model$suitRaster,
pRoc=p_roc[,c("auc_ratio","auc_pmodel")],
metadata=ellip))
})
# + [markdown] tags=[]
# # Next some executions of lines in `find_best_model` function
# -
procs <- lapply(1:length(modelos),function(x) {
proc <- modelos[[x]][[2]]
})
procs <- do.call("rbind.data.frame",procs)
procs$auc_pmodel <- as.factor(procs$auc_pmodel)
m1 <- lm(auc_ratio ~ auc_pmodel, data = procs)
model_means <- sapply(levels(procs$auc_pmodel), function(y){
model_index <- which(procs$auc_pmodel == y)
media_model <- mean(procs[model_index,1],na.rm=T)
return(media_model)
})
best_model <-names(model_means)[which(model_means==max(model_means,na.rm = TRUE))]
best_model
models_meta_data <- lapply(1:length(modelos), function(x){
matadata <- modelos[[x]][[3]]
})
best_model_metadata <- modelos[[as.numeric(best_model)]][[3]]
sp.temp.best.model <- list(sp_coords = this_species$sp_coords,
coords_env_data_all = this_species$coords_env_data_all,
env_data_train = this_species$env_data_train,
env_data_test = this_species$env_data_test,
test_data = this_species$test_data,
sp_occs_year = this_species$sp_occs_year,
oocs_data = this_species$oocs_data,
lon_lat_vars = this_species$lon_lat_vars,
layers_path_by_year = this_species$layers_path_by_year,
best_model_metadata= best_model_metadata,
ellipsoid_level =ellipsoid_level,
pROC_table = procs,
models_meta_data=models_meta_data)
class(sp.temp.best.model) <- c("list", "sp.temporal.modeling","sp.temporal.env","sp.temp.best.model")
best_model_2004 <- sp.temp.best.model
temporal_projection(this_species = best_model_2004,
save_dir = "/shared_volume/Ponca_DV/new_model",
sp_mask = ponca_mask,
crs_model = NULL,
sp_name ="pan_onca",
plot3d = FALSE)
# + [markdown] tags=[]
#
# -
vars_model <- cor_filter[combinatoria_vars[,1]]
vars_model
ellip <- try(cov_center(env_train[,vars_model],
level = ellipsoid_level ,vars = vars_model),silent = T)
occs_env <- e_test$env_data_train[,vars_model]
sp_model <- ellipsoidfit(data = env_layers[[vars_model]],
centroid =ellip$centroid,
covar = ellip$covariance,
level = ellipsoid_level,
size = 3,
plot = plot3d)
valData <- e_test$test_data[,c(1,2)]
valData$sp_name <- "sp"
valData <- valData[,c(3,1,2)]
p_roc<- PartialROC(valData = valData,
PredictionFile = sp_model$suitRaster,
E = E,
RandomPercent = RandomPercent,
NoOfIteration = NoOfIteration)
modelos <- list(list(model = sp_model$suitRaster,
pRoc=p_roc[,c("auc_ratio","auc_pmodel")],
metadata=ellip))
1:length(modelos)
procs <- lapply(1:length(modelos),function(x) {
proc <- modelos[[x]][[2]]
})
procs <- do.call("rbind.data.frame",procs)
procs$model <- as.factor(procs$auc_pmodel)
m1 <- lm(auc_ratio ~ model, data = procs)
model_means <- sapply(levels(procs$model), function(y){
model_index <- which(procs$model == y)
media_model <- mean(procs[model_index,1],na.rm=T)
return(media_model)
})
best_model <-names(model_means)[which(model_means==max(model_means,na.rm = TRUE))]
models_meta_data <- lapply(1:length(modelos), function(x){
matadata <- modelos[[x]][[3]]
})
best_model_metadata <- modelos[[as.numeric(best_model)]][[3]]
sp.temp.best.model <- list(sp_coords = e_test$sp_coords,
coords_env_data_all = e_test$coords_env_data_all,
env_data_train = e_test$env_data_train,
env_data_test = e_test$env_data_test,
test_data = e_test$test_data,
sp_occs_year = e_test$sp_occs_year,
oocs_data = e_test$oocs_data,
lon_lat_vars = e_test$lon_lat_vars,
layers_path_by_year = e_test$layers_path_by_year,
best_model_metadata= best_model_metadata,
ellipsoid_level =ellipsoid_level,
pROC_table = procs,
models_meta_data=models_meta_data)
class(sp.temp.best.model) <- c("list", "sp.temporal.modeling","sp.temporal.env","sp.temp.best.model")
class(sp.temp.best.model)
modelos <- lapply(1:dim(combinatoria_vars)[2],function(x){
cat("Doing model: ", x," of ", dim(combinatoria_vars)[2],"\n")
#MODELO DE CALIDAD DE HABITAT
system.time({best_model_2004<-find_best_model(this_species =e_test,
cor_threshold = 0.8,
ellipsoid_level = 0.975,
nvars_to_fit = 3,E = 0.05,
RandomPercent = 70,
NoOfIteration = 1000,
parallel = TRUE, #Run process in parallel
n_cores = 2,
plot3d = FALSE)})
# +
#el numero de cores depende de la maquina donde se corra
#y sirve para la paralizacion de procesos
#save(best_model_2004,file = "~/Dropbox/CONABIO_PROYECTO/PresentacionFinal/best_modelPuma.RData")
#REPROYECCION TEMPORAL
system.time({
temporal_projection(this_species = best_model_2004,
save_dir = "/Users/Mariana/Google Drive/andonue/Ponca_DV/new_model",
sp_mask = ponca_mask,
crs_model = NULL,
sp_name ="pan_onca",
plot3d = FALSE)
})
| hsi/notebooks/hsi_pipeline_debugging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import pandas_profiling
titanicRawTrain = pd.read_csv('../data/raw/train.csv')
# ## Load Rob's Titanic cleaning and feature engineering
import sys
sys.path.append('/home/rob/Dropbox/DataProject/titanicKaggle/src/features')
import build_features
titanicCleanTrain = build_features.execute_cleaning(titanicRawTrain)
titanicColumns = build_features.produce_columns(titanicCleanTrain)
titanicCleanTrain.head()
titanicCleanTrain.columns
# ## Investigate variability of factors
# ## Load new model
from sklearn.model_selection import train_test_split
from sklearn import linear_model
from sklearn.model_selection import cross_val_score
X_train, X_test, y_train, y_test = train_test_split(titanicCleanTrain[titanicColumns],
titanicCleanTrain.Survived, test_size = 0.4, random_state =0)
clf = linear_model.LogisticRegression().fit(X_train, y_train)
clf.score(X_test, y_test)
from sklearn.cross_validation import KFold
from sklearn.metrics import make_scorer, accuracy_score
def run_kfold(clf, X_all, y_all):
kf = KFold(891, n_folds = 10)
outcomes = []
fold = 0
for train_i, test_i in kf:
fold += 1
X_train, X_test = X_all.values[train_i], X_all.values[test_i]
y_train, y_test = y_all.values[train_i], y_all.values[test_i]
clf.fit(X_train, y_train)
predictions = clf.predict(X_test)
accuracy = accuracy_score(y_test, predictions)
outcomes.append(accuracy)
print("Fold {0} accuracy: {1}".format(fold, accuracy))
mean_outcome = np.mean(outcomes)
print("Mean Accuracy: {0}". format(mean_outcome))
run_kfold(clf, titanicCleanTrain[titanicColumns], titanicCleanTrain['Survived'])
# ## Test it out
titanicRawTest = pd.read_csv('../data/raw/test.csv')
data_test = build_features.execute_cleaning(titanicRawTest)
# +
ids = data_test['PassengerId']
predictions = clf.predict(data_test.drop('PassengerId', axis=1))
output = pd.DataFrame({ 'PassengerId' : ids, 'Survived': predictions })
output.to_csv('titanic-predictions.csv', index = False)
# -
output.head()
# ## ToDo:
# New features (ANOVA investigation)
#
# Normalization
#
#
| notebooks/3.0-rah-updatedLogisticRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import pickle
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
from sklearn.metrics import f1_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import recall_score
from sklearn.metrics import precision_score
from sklearn.metrics import accuracy_score
experiments = [('bgl', True, True, 0.001, False),
('bgl', True, False, 0.001, False),
('bgl', False, True, 0.001, False),
('bgl', False, False, 0.001, False),
('bgl', True, True, 0.05, False),
('bgl', True, False, 0.05, False),
('bgl', False, True, 0.05, False),
('bgl', False, False, 0.05, False),
('bgl', True, True, 0.1, False),
('bgl', True, False, 0.1, False),
('bgl', False, True, 0.1, False),
('bgl', False, False, 0.1, False),
('bgl', True, True, 0.2, False),
('bgl', True, False, 0.2, False),
('bgl', False, True, 0.2, False),
('bgl', False, False, 0.2, False),
('bgl', True, True, 0.4, False),
('bgl', True, False, 0.4, False),
('bgl', False, True, 0.4, False),
('bgl', False, False, 0.4, False),
('bgl', True, True, 0.6, False),
('bgl', True, False, 0.6, False),
('bgl', False, True, 0.6, False),
('bgl', False, False, 0.6, False),
('bgl', True, True, 0.8, False),
('bgl', True, False, 0.8, False),
('bgl', False, True, 0.8, False),
('bgl', False, False, 0.8, False)]
# +
from collections import defaultdict
def evaluate_greater(labels_test, max_distances, start_epoch, end_epoch):
res_ = defaultdict(dict)
for epoch in range(start_epoch, end_epoch):
print("Best results per epoch {}".format(epoch+1))
print("-------"*10)
f1_scores = []
precision_scores = []
recall_scores = []
auc_score = []
d = {}
fpr, tpr, thresholds = roc_curve(labels_test, max_distances[epoch], pos_label=1)
# if len(thresholds)>300:
# top_th = 100
# else:
# top_th = 50
top_th = 50
# top_th = 30
print("Number of th to evaluate {}".format(top_th))
# candidate_th = np.sort(np.abs(np.gradient(thresholds)))[(-1)*top_th:]
# candidate_th_indx = np.where((tpr - (1-fpr))==0, True, False)
# print(candidate_th_indx)
print("MEANS ", np.argmax(np.sqrt(tpr * (1-fpr))))
candidate_th = np.array(thresholds[np.argmax(np.sqrt(tpr * (1-fpr)))]).reshape(-1, 1)
candidate_th = np.array(thresholds[np.argsort((tpr-fpr))][(-1)*top_th:]).reshape(-1, 1)
# print(type(candidate_th))
# # plt.figure()
# # plt.plot(labels_test, c="b")
# # print(np.unique(labels_test, return_counts=True))
# # plt.plot(max_distances[epoch],c="r")
# # plt.figure()
# # plt.scatter(fpr, tpr)
# # plt.show()
# # plt.figure(1)
# plt.plot(tpr, c="b")
# plt.plot(1-fpr, c="r")
# plt.plot((tpr - (1-fpr)))
# plt.scatter(np.arange(tpr.shape[0]), tpr-fpr, c="g")
# plt.scatter(np.argsort((tpr-fpr))[(-1)*top_th:], (tpr-fpr)[np.argsort((tpr-fpr))[(-1)*top_th:]], c="purple")
# plt.show()
# plt.plot(thresholds)
# plt.plot(np.abs(np.gradient(thresholds)))
# plt.show()
# plt.figure(2)
# plt.scatter(fpr, tpr)
# # plt.show()
# plt.scatter(np.arange(0,1, 0.05), np.arange(0,1, 0.05))
# plt.show()
# plt.figure(1)
# plt.scatter(np.arange(len(max_distances[epoch])), max_distances[epoch])
# plt.show()
# plt.figure(2)
# plt.scatter(np.arange(len(max_distances[epoch])), labels_test)
# plt.show()
# print(np.unique(candidate_th))
# print(thresholds)
best_f1 = 0
preds = []
for th in candidate_th:
preds = np.where((max_distances[epoch]>th)==True, 1, 0)
print("The F1 score is {} the threshold is {}".format(f1_score(labels_test, preds), th))
# auc_score.append(roc_auc_score(labels_test, max_distances[epoch]))
f1_scores.append(f1_score(labels_test, preds))
if f1_score(labels_test, preds) > best_f1:
best_preds = preds
best_f1 = f1_score(labels_test, preds)
# precision_scores.append(precision_score(labels_test, preds))
# recall_scores.append(recall_score(labels_test, preds))
d["f1"] = np.max(f1_scores)
d["precision"] = precision_score(labels_test, best_preds)
d["recall"] = recall_score(labels_test, best_preds)
d["thresholds"] = candidate_th[np.argmax(f1_scores)]
d["AUC"] = roc_auc_score(labels_test, max_distances[epoch])
ind = np.argmax(f1_scores)
d["best_index_per_epoch"] = ind
print("THE AUC score is {}".format(roc_auc_score(labels_test, max_distances[epoch])))
# print("THE MAXIMAL VALUE FOR the th is {}".format(candidate_th[ind]))
# d["precision"] = precision_scores[ind]
# d["recall"] = recall_scores[ind]
# d["auc"] = auc_score[ind]
res_[epoch] = d
return res_
# -
experiment = ('spirit', True, True, 0.2, False)
# +
dataset_name = experiment[0]
path = "./log_anomaly_detection_evaluation/" + dataset_name + "/"
model_name = "transfer_"
phase = "distances_"
with open (path + model_name + phase + "_".join([str(x) for x in experiment]) + ".pickle", "rb") as file:
max_distances = pickle.load(file)
path = "./log_anomaly_detection_evaluation/" + dataset_name + "/"
model_name = "transfer_"
phase = "labels_test_"
with open (path + model_name + phase + "_".join([str(x) for x in experiment]) + ".pickle", "rb") as file:
labels_test = pickle.load(file)
# -
# +
def read_experiment(experiment):
dataset_name = experiment[0]
path = "./log_anomaly_detection_evaluation/" + dataset_name + "/run1/"
model_name = "transfer_"
phase = "distances_"
print("reading distances")
with open (path + model_name + phase + "_".join([str(x) for x in experiment]) + ".pickle", "rb") as file:
max_distances = pickle.load(file)
print("reading labels")
path = "./log_anomaly_detection_evaluation/" + dataset_name + "/run1/"
model_name = "transfer_"
phase = "labels_test_"
with open (path + model_name + phase + "_".join([str(x) for x in experiment]) + ".pickle", "rb") as file:
labels_test = pickle.load(file)
print("calculating_results")
res_ = evaluate_greater(labels_test, max_distances, start_epoch=0, end_epoch=len(max_distances))
print("------"*10)
return res_
# +
set_exp = defaultdict(dict)
for idx, experiment in enumerate(experiments):
print("_".join([str(x) for x in experiment]))
set_exp["_".join([str(x) for x in experiment])] = read_experiment(experiment)
# -
| evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
# # Conversion Rate
df = pd.read_csv('../data/bank-additional-full.csv', sep=';')
df.shape
df.tail()
df['conversion'] = df['y'].apply(lambda x: 1 if x == 'yes' else 0)
df['y']
df.head()
# ### 1. Aggregate Conversion Rate
print('total conversions: %i out of %i' % (df.conversion.sum(), df.shape[0]))
print('conversion rate: %0.2f%%' % (df.conversion.sum() / df.shape[0] * 100.0))
# ### 2. Conversion Rates by Number of Contacts
pd.DataFrame(
df.groupby(
by='campaign'
)['conversion'].sum()
)
pd.DataFrame(
df.groupby(
by='campaign'
)['conversion'].count()
)
conversions_by_contacts = df.groupby(
by='campaign'
)['conversion'].sum() / df.groupby(
by='campaign'
)['conversion'].count() * 100.0
pd.DataFrame(conversions_by_contacts)
# +
ax = conversions_by_contacts[:10].plot(
grid=True,
figsize=(10, 7),
xticks=conversions_by_contacts.index[:10],
title='Conversion Rates by Number of Contacts'
)
ax.set_ylim([0, 15])
ax.set_xlabel('number of contacts')
ax.set_ylabel('conversion rate (%)')
plt.show()
# -
# ### 3. Conversion Rates by Age
# #### - Line Chart
pd.DataFrame(
df.groupby(
by='age'
)['conversion'].sum()
)
pd.DataFrame(
df.groupby(
by='age'
)['conversion'].count()
)
conversions_by_age = df.groupby(
by='age'
)['conversion'].sum() / df.groupby(
by='age'
)['conversion'].count() * 100.0
pd.DataFrame(conversions_by_age)
# +
ax = conversions_by_age.plot(
grid=True,
figsize=(10, 7),
title='Conversion Rates by Age'
)
ax.set_xlabel('age')
ax.set_ylabel('conversion rate (%)')
plt.show()
# -
# #### - Age Groups
df['age_group'] = df['age'].apply(
lambda x: '[18, 30)' if x < 30 else '[30, 40)' if x < 40 \
else '[40, 50)' if x < 50 else '[50, 60)' if x < 60 \
else '[60, 70)' if x < 70 else '70+'
)
df.head()
pd.DataFrame(
df.groupby(
by='age_group'
)['conversion'].sum()
)
pd.DataFrame(
df.groupby(
by='age_group'
)['conversion'].count()
)
conversions_by_age_group = df.groupby(
by='age_group'
)['conversion'].sum() / df.groupby(
by='age_group'
)['conversion'].count() * 100.0
pd.DataFrame(conversions_by_age_group)
# +
ax = conversions_by_age_group.loc[
['[18, 30)', '[30, 40)', '[40, 50)', '[50, 60)', '[60, 70)', '70+']
].plot(
kind='bar',
color='skyblue',
grid=True,
figsize=(10, 7),
title='Conversion Rates by Age Groups'
)
ax.set_xlabel('age')
ax.set_ylabel('conversion rate (%)')
plt.show()
# -
# ### 4. Conversions vs. Non-Conversions
# #### 4.1. Marital Status
conversions_by_marital_status_df = pd.pivot_table(df, values='y', index='marital', columns='conversion', aggfunc=len)
conversions_by_marital_status_df
conversions_by_marital_status_df.columns = ['non_conversions', 'conversions']
conversions_by_marital_status_df
df.groupby('conversion')['duration'].describe()
df.loc[df['conversion'] == 0, 'duration'].reset_index(drop=True)
df.columns
pd.concat([
df.loc[df['conversion'] == 0, 'duration'].reset_index(drop=True),
df.loc[df['conversion'] == 1, 'duration'].reset_index(drop=True)
],axis=1)
# +
duration_df=pd.concat([
df.loc[df['conversion'] == 0, 'duration'].reset_index(drop=True),
df.loc[df['conversion'] == 1, 'duration'].reset_index(drop=True)
],axis=1)
duration_df.columns = ['conversions', 'non_conversions']
# -
ax=duration_df.plot(
kind='box',
grid=True,
figsize=(10, 10),
)
ax.set_ylabel('last contact duration (hours)')
ax.set_xlabel('last contact duration')
# +
conversions_by_marital_status_df.plot(
kind='pie',
figsize=(15, 7),
startangle=90,
subplots=True,
autopct=lambda x: '%0.1f%%' % x
)
plt.show()
# -
# #### 4.2. Education
conversions_by_education_df = pd.pivot_table(df, values='y', index='education', columns='conversion', aggfunc=len)
conversions_by_education_df
conversions_by_education_df.columns = ['non_conversions', 'conversions']
conversions_by_education_df
# +
conversions_by_education_df.plot(
kind='pie',
figsize=(15, 7),
startangle=90,
subplots=True,
autopct=lambda x: '%0.1f%%' % x,
legend=False
)
plt.show()
# -
# #### 4.3. Last Contact Duration
df.groupby('conversion')['duration'].describe()
# +
duration_df = pd.concat([
df.loc[df['conversion'] == 1, 'duration'].reset_index(drop=True),
df.loc[df['conversion'] == 0, 'duration'].reset_index(drop=True)
], axis=1)
duration_df.columns = ['conversions', 'non_conversions']
duration_df = duration_df / (60*60)
# -
duration_df
# +
ax = duration_df.plot(
kind='box',
grid=True,
figsize=(10, 10),
)
ax.set_ylabel('last contact duration (hours)')
ax.set_title('Last Contact Duration')
plt.show()
# -
# ### 5. Conversions by Age Groups & Marital Status
age_marital_df = df.groupby(['age_group', 'marital'])['conversion'].sum().unstack('marital').fillna(0)
age_marital_df = age_marital_df.divide(
df.groupby(
by='age_group'
)['conversion'].count(),
axis=0
)
age_marital_df
# +
ax = age_marital_df.loc[
['[18, 30)', '[30, 40)', '[40, 50)', '[50, 60)', '[60, 70)', '70+']
].plot(
kind='bar',
grid=True,
figsize=(10,7)
)
ax.set_title('Conversion rates by Age & Marital Status')
ax.set_xlabel('age group')
ax.set_ylabel('conversion rate (%)')
plt.show()
# +
ax = age_marital_df.loc[
['[18, 30)', '[30, 40)', '[40, 50)', '[50, 60)', '[60, 70)', '70+']
].plot(
kind='bar',
stacked=True,
grid=True,
figsize=(10,7)
)
ax.set_title('Conversion rates by Age & Marital Status')
ax.set_xlabel('age group')
ax.set_ylabel('conversion rate (%)')
plt.show()
| Chapter02/ConversionRate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training from scratch on Imagewoof with the efficientnet_pytorch repo
# https://github.com/lukemelas/EfficientNet-PyTorch
# !pip install efficientnet_pytorch
# +
from fastai.script import *
from fastai.vision import *
from fastai.callbacks import *
from fastai.distributed import *
from fastprogress import fastprogress
from torchvision.models import *
from efficientnet_pytorch import EfficientNet
import sys
torch.backends.cudnn.benchmark = True
fastprogress.MAX_COLS = 80
def get_data(size, woof, bs, workers=None):
if size<=128: path = URLs.IMAGEWOOF_160 if woof else URLs.IMAGENETTE_160
elif size<=224: path = URLs.IMAGEWOOF_320 if woof else URLs.IMAGENETTE_320
else : path = URLs.IMAGEWOOF if woof else URLs.IMAGENETTE
path = untar_data(path)
n_gpus = num_distrib() or 1
if workers is None: workers = min(8, num_cpus()//n_gpus)
return (ImageList.from_folder(path).split_by_folder(valid='val')
.label_from_folder().transform(([flip_lr(p=0.5)], []), size=size)
.databunch(bs=bs, num_workers=workers)
.presize(size, scale=(0.35,1))
.normalize(imagenet_stats))
# -
# Change image size and batch size below depending on model:
data = get_data(300,1,16) #240, bs=32 for B1, 300, bs=16 for B3
opt_func = partial(optim.Adam, betas=(0.9,0.99), eps=1e-6)
# Pick model below:
m = EfficientNet.from_name('efficientnet-b3')
m._fc = nn.Linear(m._fc.in_features, out_features=10, bias=True)
nn.init.kaiming_normal_(m._fc.weight);
# +
learn = (Learner(data, m, wd=1e-5, opt_func=opt_func,
metrics=[accuracy,top_k_accuracy],
bn_wd=False, true_wd=True,
loss_func = LabelSmoothingCrossEntropy())
)
# +
mixup = 0
if mixup: learn = learn.mixup(alpha=mixup)
learn = learn.to_fp16(dynamic=True)
# -
#B1
learn.lr_find()
learn.recorder.plot()
# ## 5 epochs B1 (xresnet50 gets ~ 62%; xresnet50 + self attention gets 67%)
learn.fit_one_cycle(5, 1e-4, div_factor=10, pct_start=0.3)
# restarted from scratch
learn.fit_one_cycle(5, 1e-3, div_factor=10, pct_start=0.3)
# restarted from scratch
learn.fit_one_cycle(5, 5e-3, div_factor=10, pct_start=0.3)
# ## 80 epochs B1 (xresnet50 gets 89.9% on 256px)
# restarted from scratch, mixup =0.2
learn.fit_one_cycle(80, 1e-3, div_factor=10, pct_start=0.3)
# # 5 epochs B3
learn.lr_find()
learn.recorder.plot()
learn.fit_one_cycle(5, 1e-4, div_factor=10, pct_start=0.3)
#restart
learn.fit_one_cycle(5, 1e-3, div_factor=10, pct_start=0.3)
#restart
learn.fit_one_cycle(5, 5e-5, div_factor=10, pct_start=0.3)
| Imagewoof from scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center>Построение простейших моделей прогнозирования</center>
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Загрузим и отобразим данные об объемах пассажирских авиаперевозок.
passengers = pd.read_csv('data/passengers.csv', index_col='date', parse_dates=True)
passengers['num_passengers'].plot(figsize=(12,5))
plt.title('International airline passengers: monthly totals in thousands. Jan 49 – Dec 60')
plt.show()
# Временной ряд обладает трендом и годовой сезонностью, которые должны быть учтены при построении моделей прогнозирования. Наличие тренда и сезонности проявляется на коррелограмме. Для построения коррелограммы воспользуемся функцией `plot_acf` из библиотеки `statsmodels`.
from statsmodels.graphics.tsaplots import plot_acf
# Опция `lags` позволяет задавать количество лагов для вычисления значений автокорреляции.
fig, ax = plt.subplots(figsize=(12,5))
plot_acf(passengers['num_passengers'], lags=40, ax=ax)
plt.show()
# <div class="alert alert-info">
#
# <h3> Задание 1</h3>
# <p></p>
# Постройте наивную сезонную модель прогнозирования. Данная модель предполагает, что количество пассажиров в январе 1961 г. будет таким же, как и в январе 1960 г. С помощью полученной модели получите прогноз на 3 года вперед и отобразите результат на графике.
# <p></p>
#
# </div>
# <div class="alert alert-info">
#
# <h3> Задание 2</h3>
# <p></p>
# Наивная сезонная модель прогнозирования не учитывает тренд временного ряда. Воспользуйтесь методом экстраполяции тренда, постройте прогноз также на 3 года и отобразите результат на графике.
#
# <em>Комментарий: пока реализуйте только метод экстраполяции тренда! Не спешите объединять модели.</em>
# <p></p>
#
# </div>
# <div class="alert alert-info">
#
# <h3> Задание 3</h3>
# <p></p>
# В результате выполнения предыдущих заданий вы получили две модели, каждая из которых описывает только одну из компонент ряда: либо тренд, либо сезонность. Попробуем их объединить. Для этого:
# <p></p>
# <li>вычтите из исходного ряда трендовую составляющую, постройте коррелограмму полученного ряда остатков;</li>
# <li>для ряда остатков постройте наивную сезонную модель.</li>
#
# Получите прогноз на 3 года вперед с помощью комбинации моделей. Результат отобразите на графике.
# <p></p>
#
# </div>
# ### Мультипликативная сезонность
# Во временном ряду пассажирских авиаперевозок наблюдается **мультипликативная сезонность**, которая характеризуется увеличением амплитуды сезонных колебаний. В случае когда амплитуда сезонных колебаний не меняется, говорят об **аддитивной сезонности**.
# Ряд с трендом и аддитивной сезонностью может быть представлен в виде суммы его компонент:
#
# $$y(t)=b(t)+s(t)+\epsilon(t),$$
#
# где $b(t)$ – трендовая, $s(t)$ – сезонная, а $\epsilon(t)$ – случайная компоненты ряда.
# Ряд с мультипликативной сезонностью представляет собой произведение данных компонент:
#
# $$y(t)=b(t)\times s(t)\times \epsilon(t).$$
# <div class="alert alert-info">
#
# <h3> Задание 4</h3>
# <p></p>
# Постройте мультипликативную модель прогнозирования, получите прогноз на 3 года вперед и отобразите результат на графике.
# <p></p>
#
# </div>
| Forecasting-Methods/1. Naive forecasting models 2/1. Naive forecasting models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# This notebook was used to perform training and evaluation of a simple two-hidden-layer and three-hidden layers on
# MNIST and Fashion MNIST datasets.
#
# No external dependencies are needed, simply open the notebook in Jupyter server and run all cells. Note that the simulation
# is relatively long (each model took approx. 30 - 60 minutes on one dataset) and thus strong GPU or cloud computation
# is recommended.
#
# Part of the code is adapted from the SpyTorch GitHub repository: https://github.com/fzenke/spytorch
# -
from typing import List, Tuple
import torch.nn as nn
import torch
import os
import torchvision
import numpy as np
import pandas as pd
from sklearn import metrics
# + pycharm={"name": "#%%\n"}
# Set seed for consistent behavior (though different results will still occur due to computation on GPU - if present)
seed = 0
torch.manual_seed(seed)
np.random.seed(seed)
# Set the device for computation
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
device
# + pycharm={"name": "#%%\n"}
# Set data type for computations in the neural network
dtype = torch.float
# + pycharm={"name": "#%%\n"}
# Function to convert the inputs into spikes
def convert_to_firing_time(x, tau=20, thr=0.2, tmax=1.0, epsilon=1e-7):
idx = x < thr
x = np.clip(x, thr + epsilon, 1e9)
T = tau * np.log(x / (x - thr))
T[idx] = tmax
return T
# This function is used to generate features and labels for the simulator in batches
def sparse_data_generator(x, y, batch_size, num_steps, num_units, time_step=1e-3, shuffle=True):
labels_ = np.array(y, dtype=np.int)
number_of_batches = len(x) // batch_size
sample_index = np.arange(len(x))
# compute discrete firing times
tau_eff = 20e-3 / time_step
firing_times = np.array(convert_to_firing_time(x, tau=tau_eff, tmax=num_steps), dtype=np.int)
unit_numbers = np.arange(num_units)
# shuffle if set to True
if shuffle:
np.random.shuffle(sample_index)
counter = 0
while counter < number_of_batches:
batch_index = sample_index[batch_size * counter:batch_size * (counter + 1)]
coo = [[] for _ in range(3)]
for bc, idx in enumerate(batch_index):
c = firing_times[idx] < num_steps
times, units = firing_times[idx][c], unit_numbers[c]
batch = [bc for _ in range(len(times))]
coo[0].extend(batch)
coo[1].extend(times)
coo[2].extend(units)
i = torch.LongTensor(coo).to(device)
v = torch.FloatTensor(np.ones(len(coo[0]))).to(device)
X_batch = torch.sparse.FloatTensor(i, v, torch.Size([batch_size, num_steps, num_units])).to(device)
y_batch = torch.tensor(labels_[batch_index], device=device, dtype=torch.long)
yield X_batch.to(device=device), y_batch.to(device=device)
counter += 1
class SurrGradSpike(torch.autograd.Function):
scale = 100.0 # controls steepness of surrogate gradient
@staticmethod
def forward(ctx, input):
ctx.save_for_backward(input)
out = torch.zeros_like(input)
out[input > 0] = 1.0
return out
@staticmethod
def backward(ctx, grad_output):
input, = ctx.saved_tensors
grad_input = grad_output.clone()
grad = grad_input / (SurrGradSpike.scale * torch.abs(input) + 1.0) ** 2
return grad
# + pycharm={"name": "#%%\n"}
class DeepSNNModel:
"""
This class implements a simple deep snn model that comprises a set of fully connected layers
"""
def __init__(self, units: List[int], weight_scale: Tuple = (7.0, 1.0), recurrent=False, time_step=1e-3,
tau_mem=10e-3,
tau_syn=5e-3):
"""
:param units: list of units in each layer
:param weight_scale: tuple of one or two values, controls the scaling of the weights
:param recurrent: whether the network is recurrent one - additional recurrent weights are used
:param time_step: how long does one time step take (seconds)
:param tau_mem: membrane tau parameter
:param tau_syn: synapse tau parameter
"""
self.units = units
self.tau_mem = tau_mem
self.tau_syn = tau_syn
self.alpha = float(np.exp(-time_step / tau_syn))
self.beta = float(np.exp(-time_step / tau_mem))
self.is_recurrent = recurrent
if len(weight_scale) == 2:
self.weight_scale = weight_scale[0] * (weight_scale[1] - self.beta)
else:
self.weight_scale = weight_scale[0]
self.weights = self._init_weights()
self.recurrent_weights = None if not recurrent else self._init_recurrent_weights()
def _init_weights(self):
"""
Initializes weights between layers
:return: list of torch tensors representing weights
"""
weights = []
for i in range(len(self.units) - 1):
wi = torch.empty((self.units[i], self.units[i + 1]), device=device, dtype=dtype, requires_grad=True)
torch.nn.init.normal_(wi, mean=0.0, std=self.weight_scale / np.sqrt(self.units[i]))
weights.append(wi)
return weights
def _init_recurrent_weights(self):
"""
Initializes recurrent weights for recurrent network for every hidden layer
:return: list of torch tensors representing recurrent weights
"""
recurrent_weights = []
for i in range(1, len(self.units) - 1):
vi = torch.empty((self.units[i], self.units[i]), device=device, dtype=dtype, requires_grad=True)
torch.nn.init.normal_(vi, mean=0.0, std=self.weight_scale / np.sqrt(self.units[i]))
recurrent_weights.append(vi)
return recurrent_weights
def run_recurrent(self, inputs, batch_size, steps, spike_fn=SurrGradSpike.apply):
"""
Runs recurrent network
:param inputs: input data
:param batch_size: batch size
:param steps: number of timesteps
:param spike_fn: surrogate gradient function
:return: output layer results
"""
layer_outputs = [inputs]
mem_recs = []
spike_recs = []
# compute activity in each hidden layer
for i in range(1, len(self.units) - 1): # the current hidden layer index
hidden_units = self.units[i] # neurons in the current hidden layer
syn_hidden_i = torch.zeros((batch_size, hidden_units), device=device, dtype=dtype)
mem_hidden_i = torch.zeros((batch_size, hidden_units), device=device, dtype=dtype)
mem_rec_hidden_i = [mem_hidden_i]
spike_rec_hidden_i = [mem_hidden_i]
hi = torch.zeros((batch_size, hidden_units), device=device, dtype=dtype)
hi_from_prev_layer = torch.einsum('abc, cd -> abd', (layer_outputs[i - 1], self.weights[i - 1]))
for dt in range(steps):
mem_threshold = mem_hidden_i - 1.0
spike_out = spike_fn(mem_threshold)
rst = torch.zeros_like(mem_hidden_i)
c = (mem_threshold > 0)
rst[c] = torch.ones_like(mem_hidden_i)[c]
hi = hi_from_prev_layer[:, dt] + torch.einsum('ab, bc -> ac', (hi, self.recurrent_weights[i - 1]))
new_syn = self.alpha * syn_hidden_i + hi
new_mem = self.beta * mem_hidden_i + syn_hidden_i - rst
mem_hidden_i = new_mem
syn_hidden_i = new_syn
mem_rec_hidden_i.append(mem_hidden_i)
spike_rec_hidden_i.append(spike_out)
spike_rec_hidden_i = torch.stack(spike_rec_hidden_i, dim=1)
mem_rec_hidden_i = torch.stack(mem_rec_hidden_i, dim=1)
layer_outputs.append(spike_rec_hidden_i) # append output so it can be fed to the next hidden layer
mem_recs.append(mem_rec_hidden_i)
spike_recs.append(spike_rec_hidden_i)
# output layer
hn = torch.einsum('abc, cd -> abd', (layer_outputs[-1], self.weights[-1]))
flt = torch.zeros((batch_size, self.units[-1]), device=device, dtype=dtype)
spike_out = torch.zeros((batch_size, self.units[-1]), device=device, dtype=dtype)
out_rec = [spike_out]
for dt in range(steps):
new_flt = self.alpha * flt + hn[:, dt]
new_out = self.beta * spike_out + flt
flt = new_flt
spike_out = new_out
out_rec.append(spike_out)
out_rec = torch.stack(out_rec, dim=1)
return out_rec, spike_recs, layer_outputs, mem_recs
def run_feed_forward(self, inputs, batch_size, steps, spike_fn=SurrGradSpike.apply):
"""
Run feed forward network in the same manner as a recurrent one except that no recurrent weights are calculated
:param inputs: input data
:param batch_size: batch size
:param steps: number of time steps
:param spike_fn: surrogate gradient function
:return: output layer results
"""
layer_outputs = [inputs]
mem_recs = []
spike_recs = []
# compute activity of every hidden layer
# hidden layers are stored from index 1 to index len(self.units) - 2
for i in range(1, len(self.units) - 1):
hidden_units = self.units[i] # units in next layer
# sum of output from previous layer and weights of current hidden layer
hi = torch.einsum('abc,cd->abd', (layer_outputs[i - 1], self.weights[i - 1]))
syn_hidden_i = torch.zeros((batch_size, hidden_units), device=device, dtype=dtype) # synapses
mem_hidden_i = torch.zeros((batch_size, hidden_units), device=device, dtype=dtype) # membranes
mem_rec_hidden_i = [mem_hidden_i]
spike_rec_hidden_i = [mem_hidden_i]
for dt in range(steps):
mem_threshold = mem_hidden_i - 1.0
spike_out = spike_fn(mem_threshold)
rst = torch.zeros_like(mem_hidden_i)
c = (mem_threshold > 0)
rst[c] = torch.ones_like(mem_hidden_i)[c]
new_syn = self.alpha * syn_hidden_i + hi[:, dt]
new_mem = self.beta * mem_hidden_i + syn_hidden_i - rst
mem_hidden_i = new_mem
syn_hidden_i = new_syn
mem_rec_hidden_i.append(mem_hidden_i)
spike_rec_hidden_i.append(spike_out)
spike_rec_hidden_i = torch.stack(spike_rec_hidden_i, dim=1)
mem_rec_hidden_i = torch.stack(mem_rec_hidden_i, dim=1)
layer_outputs.append(spike_rec_hidden_i) # append output so it can be fed to the next hidden layer
mem_recs.append(mem_rec_hidden_i)
spike_recs.append(spike_rec_hidden_i)
# readout layer
hn = torch.einsum('abc,cd->abd', (layer_outputs[-1], self.weights[-1]))
flt = torch.zeros((batch_size, self.units[-1]), device=device, dtype=dtype)
spike_out = torch.zeros((batch_size, self.units[-1]), device=device, dtype=dtype)
out_rec = [spike_out]
for dt in range(steps):
new_flt = self.alpha * flt + hn[:, dt]
new_out = self.beta * spike_out + flt
flt = new_flt
spike_out = new_out
out_rec.append(spike_out)
out_rec = torch.stack(out_rec, dim=1)
return out_rec, spike_recs, layer_outputs, mem_recs
def train(self, x_data, y_data, batch_size, num_steps=100, time_step=1e-3, lr=1e-3, num_epochs=10):
"""
Train the network
:param x_data: features (training data)
:param y_data: labels (training)
:param batch_size: number of elements per batch
:param num_steps: number of time steps
:param time_step: how long does a time step take (s)
:param lr: learning rate
:param num_epochs: number of epochs for training
:return:
"""
# Create optimizer, and loss function
optimizer = torch.optim.Adam(self.weights, lr=lr, betas=(0.9, 0.999))
log_softmax_fn = nn.LogSoftmax(dim=1)
loss_fn = nn.NLLLoss()
# Begin training
loss_hist = []
for epoch in range(num_epochs):
local_loss = []
for x_local, y_local in sparse_data_generator(x_data, y_data, batch_size, num_steps, self.units[0],
time_step):
if not self.is_recurrent: # if the network does not contain recurrent weights run it as feedforward
output, spike_recs, layer_outputs, mem_recs = self.run_feed_forward(x_local.to_dense(), batch_size,
num_steps)
else:
output, spike_recs, layer_outputs, mem_recs = self.run_recurrent(x_local.to_dense(), batch_size,
num_steps)
output_max, _ = torch.max(output, 1)
log_p_y = log_softmax_fn(output_max) # apply log softmax function
loss_val = loss_fn(log_p_y, y_local) # calculate the loss
# Perform the optimization
optimizer.zero_grad()
loss_val.backward()
optimizer.step()
local_loss.append(loss_val.item())
mean_loss = np.mean(local_loss)
print('Epoch {}: loss={:.5f}'.format(epoch + 1, mean_loss)) # log the loss to console
loss_hist.append(mean_loss)
return loss_hist
# + pycharm={"name": "#%%\n"}
def get_metrics(x_test, y_test, batch_size, snn_model, time_step=1e-3, num_steps=100):
"""
Calculate metrics for the network
:param x_test: testing features
:param y_test: testing labels
:param batch_size: number of elements per batch
:param snn_model: reference to the trained snn model
:param time_step: how long does a time step take (s)
:param num_steps: number of time steps
:return: accuracy, precision, recall, f1 metric and confusion matrix
"""
predictions = []
# Truncate the remaining number of samples according to batch size
samples = (x_test.shape[0] // batch_size ) * batch_size
x_test, y_test = x_test[:samples], y_test[:samples]
for x_local, y_local in sparse_data_generator(x_test, y_test, batch_size, num_steps, snn_model.units[0], time_step,
False):
# Run feedforward/recurrent network and get the output
output, spike_recs, layer_outputs, mem_recs = snn_model.run_recurrent(x_local.to_dense(), batch_size, num_steps) \
if snn_model.recurrent_weights else snn_model.run_feed_forward(x_local.to_dense(), batch_size, num_steps)
output_max, _ = torch.max(output, 1)
_, output_argmax = torch.max(output_max, 1)
# Append the current batch
predictions.append(output_argmax)
# Concatenate to a single tensor
predictions = torch.cat(predictions)
# Map to numpy
predictions = predictions.cpu().detach().numpy()
# Now calculate the metrics
# Use the "micro" average - i.e. the calculation is done globally, counting total number of tp, fp and fn for
# each class
precision = metrics.precision_score(y_true=y_test, y_pred=predictions, average='macro')
recall = metrics.recall_score(y_true=y_test, y_pred=predictions, average='macro')
f1 = metrics.f1_score(y_true=y_test, y_pred=predictions, average='macro')
accuracy = metrics.accuracy_score(y_true=y_test, y_pred=predictions)
confusion_matrix = metrics.confusion_matrix(y_true=y_test, y_pred=predictions)
# Return the metrics
return accuracy, precision, recall, f1, confusion_matrix
# + pycharm={"name": "#%%\n"}
# Download datasets through PyTorch (if necessary)
dataset_folder = os.path.join('cached_datasets')
train_dataset = torchvision.datasets.MNIST(dataset_folder, train=True,
transform=None, target_transform=None, download=True)
test_dataset = torchvision.datasets.MNIST(dataset_folder, train=False,
transform=None, target_transform=None, download=True)
# + pycharm={"name": "#%%\n"}
# Declaration of models that will be tested on the MNIST and the Fashion MNIST datasets
models_mnist = [
DeepSNNModel([28*28, 256, 128, 10]),
DeepSNNModel([28*28, 256, 128, 64, 10]),
DeepSNNModel([28*28, 256, 128, 10], recurrent=True),
DeepSNNModel([28*28, 256, 128, 64, 10], recurrent=True)
]
# Standardize the MNIST dataset
mnist_x_train = np.array(train_dataset.data, dtype=np.float)
mnist_x_train = mnist_x_train.reshape(mnist_x_train.shape[0], -1) / 255
mnist_x_test = np.array(test_dataset.data, dtype=np.float)
mnist_x_test = mnist_x_test.reshape(mnist_x_test.shape[0], -1) / 255
mnist_y_train = np.array(train_dataset.targets, dtype=np.int)
mnist_y_test = np.array(test_dataset.targets, dtype=np.int)
'The MNIST dataset is ready.'
# + pycharm={"name": "#%%\n"}
print('Running benchmark on MNIST dataset')
# Array for statistics of each model
mnist_stats = []
# Run each model
for i, model in enumerate(models_mnist):
model.train(mnist_x_train, mnist_y_train, 256, num_epochs=30)
accuracy, precision, recall, f1, confusion_matrix = get_metrics(mnist_x_test, mnist_y_test, 256, model)
stats = {
'accuracy': accuracy,
'precision': precision,
'recall': recall,
'f1': f1,
'confusion_matrix': confusion_matrix
}
print(f'Model {i+1}: {stats}')
mnist_stats.append(stats)
# + pycharm={"name": "#%%\n"}
# Fashion MNIST
train_dataset = torchvision.datasets.FashionMNIST(dataset_folder, train=True,
transform=None, target_transform=None, download=True)
test_dataset = torchvision.datasets.FashionMNIST(dataset_folder, train=False,
transform=None, target_transform=None, download=True)
# Create the same models for the Fashion MNIST dataset
models_fashion_mnist = [
DeepSNNModel([28*28, 256, 128, 10]),
DeepSNNModel([28*28, 256, 128, 64, 10]),
DeepSNNModel([28*28, 256, 128, 10], recurrent=True),
DeepSNNModel([28*28, 256, 128, 64, 10], recurrent=True)
]
# Standardize the data
fmnist_x_train = np.array(train_dataset.data, dtype=np.float)
fmnist_x_train = fmnist_x_train.reshape(fmnist_x_train.shape[0], -1) / 255
fmnist_x_test = np.array(test_dataset.data, dtype=np.float)
fmnist_x_test = fmnist_x_test.reshape(fmnist_x_test.shape[0], -1) / 255
fmnist_y_train = np.array(train_dataset.targets, dtype=np.int)
fmnist_y_test = np.array(test_dataset.targets, dtype=np.int)
fashion_mnist_stats = []
i = 1
for i, model in enumerate(models_fashion_mnist):
model.train(mnist_x_train, mnist_y_train, 256, num_epochs=30) # train the model
# Get metrics
accuracy, precision, recall, f1, confusion_matrix = get_metrics(mnist_x_test, mnist_y_test, 256, model)
# Create dictionary for later use
stats = {
'accuracy': accuracy,
'precision': precision,
'recall': recall,
'f1': f1,
'confusion_matrix': confusion_matrix
}
# Print and save the stats
print(f'Model {i+1}: {stats}')
fashion_mnist_stats.append(stats)
# + pycharm={"name": "#%%\n"}
mnist_df = pd.DataFrame({
'model': [f'{i}' for i in range(1, len(models_mnist) + 1)],
'accuracy': [x['accuracy'] for x in mnist_stats],
'precision': [x['precision'] for x in mnist_stats],
'recall': [x['recall'] for x in mnist_stats],
'f1': [x['f1'] for x in mnist_stats],
})
mnist_df
# + pycharm={"name": "#%%\n"}
# Print confusion matrix for each model
print('Confusion Matrices:')
for confusion_matrix in map(lambda x: x['confusion_matrix'], mnist_stats):
print(confusion_matrix)
# + pycharm={"name": "#%%\n"}
# Create output directory to save the results
output_path = 'surrogate_gradient_results'
os.makedirs(output_path, exist_ok=True)
# + pycharm={"name": "#%%\n"}
# Save as XLSX (excel) file
mnist_df.to_excel(os.path.join(output_path, 'mnist_results.xlsx'))
# + pycharm={"name": "#%%\n"}
fmnist_df = pd.DataFrame({
'model': [f'{i}' for i in range(1, len(models_fashion_mnist))],
'accuracy': [x['accuracy'] for x in fashion_mnist_stats],
'precision': [x['precision'] for x in fashion_mnist_stats],
'recall': [x['recall'] for x in fashion_mnist_stats],
'f1': [x['f1'] for x in fashion_mnist_stats],
})
fmnist_df
# + pycharm={"name": "#%%\n"}
# Print confusion matrix for each model
print('Confusion Matrices:')
for confusion_matrix in map(lambda x: x['confusion_matrix'], fashion_mnist_stats):
print(confusion_matrix)
# + pycharm={"name": "#%%\n"}
# Save as XLSX
fmnist_df.to_excel(os.path.join(output_path, 'fashion_mnist_results.xlsx'))
# -
| experiments/surrogate_gradient_experiment/surrogate_gradient_deep_snns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %env CUDA_DEVICE_ORDER=PCI_BUS_ID
# %env CUDA_VISIBLE_DEVICES=1
from cuml.manifold.umap import UMAP as cumlUMAP
import numpy as np
import numpy as np
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
import pandas as pd
from cuml.manifold.umap import UMAP as cumlUMAP
from avgn.utils.paths import DATA_DIR, most_recent_subdirectory, ensure_dir
from avgn.signalprocessing.create_spectrogram_dataset import flatten_spectrograms
# ### load data
DATASET_ID = 'batsong_segmented'
df_loc = DATA_DIR / 'syllable_dfs' / DATASET_ID / 'fruitbat.pickle'
syllable_df = pd.read_pickle(df_loc)
syllable_df[:3]
len(syllable_df)
np.shape(syllable_df.spectrogram.values[0])
len(syllable_df.indv.unique())
syllable_df['indv_nonneg'] = [i if '-' not in i else i[1:] for i in syllable_df.indv.values]
syllable_df.indv_nonneg.unique()
# ### project
for indv in tqdm(syllable_df.indv_nonneg.unique()):
subset_df = syllable_df[syllable_df.indv_nonneg == indv]
specs = list(subset_df.spectrogram.values)
specs = [i/np.max(i) for i in tqdm(specs)]
specs_flattened = flatten_spectrograms(specs)
print(np.shape(specs_flattened))
embedding = cumlUMAP(0.01).fit_transform(specs_flattened)
subset_df['umap-0.01'] = list(embedding)
embedding = cumlUMAP(min_dist=0.1).fit_transform(specs_flattened)
subset_df['umap-0.1'] = list(embedding)
embedding = cumlUMAP(min_dist=0.5).fit_transform(specs_flattened)
subset_df['umap-0.5'] = list(embedding)
unique_labs = np.unique(subset_df.context.values)
unique_labs_dict = {lab:i for i, lab in enumerate(unique_labs)}
lab_list = [unique_labs_dict[i] for i in subset_df.context.values]
fig, ax = plt.subplots()
ax.scatter(embedding[:,0], embedding[:,1], s=1, c=lab_list, cmap=plt.cm.tab20, alpha = 0.25)
ax.set_xlim([-8,8])
ax.set_ylim([-8,8])
plt.show()
ensure_dir(DATA_DIR / 'embeddings' / 'multiple_spread' / DATASET_ID / 'indvs')
subset_df.to_pickle(DATA_DIR / 'embeddings' / 'multiple_spread' / DATASET_ID / 'indvs' / (indv + '.pickle'))
embedding
| notebooks/02.5-make-projection-dfs/multiple_spread/batsong-umap-indv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''.venv'': venv)'
# language: python
# name: python3
# ---
# +
import pandas as pd
import json
import os
from decouple import config
import pyodbc
import numpy as np
# -
data = pd.read_csv('water_data.csv')
data_unpivot = data.melt(id_vars=['ID', 'INTERFACE_ID', 'DATETIME'], var_name='aid', value_name='value')
data_unpivot['aid'] = data_unpivot['aid'].map(lambda x: x.lstrip('AI'))
data_unpivot = data_unpivot.rename(columns={"INTERFACE_ID" : "unitID", "aid" : "analogID", "value" : "readingValue"})
data_unpivot = data_unpivot.astype({'analogID': 'int64'})
# +
#data_unpivot.to_csv('water_data_unpivot.csv', index=False)
# +
BASE_URL = config('SEL_API_URL')
USER_KEY = config('SEL_USER_KEY')
API_KEY = config('SEL_API_KEY')
DB_DRIVER = config('DB_DRIVER')
DB_URL = config('AZURE_DB_SERVER')
DB_BATABASE = config('AZURE_DB_DATABASE')
DB_USR = config('AZURE_DB_USR')
DB_PWD = config('AZURE_DB_PWD')
# Formatted connection string for the SQL DB.
SQL_CONN_STR = "DRIVER={0};SERVER={1};Database={2};UID={3};PWD={4};".format(DB_DRIVER, DB_URL, DB_BATABASE, DB_USR, DB_PWD)
# -
conn = pyodbc.connect(SQL_CONN_STR)
sql = """
SELECT DISTINCT [unitGUID], [unitID], [unitName]
FROM [SEL_UNITS]
"""
units = pd.read_sql(sql, conn)
sql = """
SELECT DISTINCT r.[sensorGUID], r.[unitGUID], r.[analogID], r.[mUnitGUID], s.sensorName
FROM [SEL_READINGS] as r
JOIN [SEL_SENSORS] as s
ON (r.[sensorGUID] = s.[sensorGUID])
"""
sensors = pd.read_sql(sql, conn)
sql = """
SELECT DISTINCT [mUnitGUID], [mUnitName]
FROM [SEL_MEASURE_UNITS]
"""
m_units = pd.read_sql(sql, conn)
# Find and process IDs here
join_df = data_unpivot.merge(units, on="unitID", how="inner")
join_df = join_df.merge(sensors, on=["unitGUID", "analogID"])
join_df = join_df.merge(m_units, on=["mUnitGUID"])
join_df
join_df = join_df[['DATETIME', 'analogID', 'readingValue', 'unitName', 'sensorName', 'mUnitName']].copy()
join_df.to_csv('water_data_join.csv', index=False)
new_df = join_df[["unitGUID","mUnitGUID", "sensorGUID", "analogID", "readingValue"]].copy()
cursor = conn.cursor()
cursor.fast_executemany = True
sql = """
SET NOCOUNT ON;
DECLARE @readingGUID UNIQUEIDENTIFIER
SET @readingGUID = NULL
SET @readingGUID = NEWID()
INSERT INTO [SEL_READINGS] (readingGUID, unitGUID, mUnitGUID, sensorGUID, analogID, readingValue)
OUTPUT Inserted.readingGUID
VALUES (@readingGUID, ?, ?, ?, ?, ?);
"""
params = new_df.values.tolist()
unitGUID = cursor.executemany(sql, params)
# +
try:
first_result = cursor.fetchall()
except pyodbc.ProgrammingError:
first_result = None
result_sets = []
while cursor.nextset():
result_sets.append(cursor.fetchall())
all_inserted_ids = np.array(result_sets).flatten()
# -
cursor.commit()
sql = """
INSERT INTO [SEL_UPDATES] (readingGUID, lastUpdate) VALUES (?, ?)
"""
params = (unitGUID, data_unpivot[['DATETIME']])
cursor.executemany(sql, params)
# +
try:
first_result = cursor.fetchall()
except pyodbc.ProgrammingError:
first_result = None
result_sets = []
while cursor.nextset():
result_sets.append(cursor.fetchall())
all_inserted_ids = np.array(result_sets).flatten()
# -
cursor.commit()
cursor.close()
conn.close()
| SEL/misc/water_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2-3 Intro Python Practice
# ## power iteration of sequences
#
# <font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - Iterate through Lists using **`for`** and **`in`**
# - Use **`for` *`count`* `in range()`** in looping operations
# - Use list methods **`.extend()`, `+, .reverse(), .sort()`**
# - convert between lists and strings using **`.split()`** and **`.join()`**
# - cast strings to lists **/** direct multiple print outputs to a single line. ** `print("hi", end='')`**
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
# ## list iteration: `for in`
# ### `for item in list:`
# +
# [ ] print out the "physical states of matter" (matter_states) in 4 sentences using list iteration
# each sentence should be of the format: "Solid - is state of matter #1"
matter_states = ['solid', 'liquid', 'gas', 'plasma']
# +
# [ ] iterate the list (birds) to see any bird names start with "c" and remove that item from the list
# print the birds list before and after removals
birds = ["turkey", "hawk", "chicken", "dove", "crow"]
# +
# the team makes 1pt, 2pt or 3pt baskets
# [ ] print the occurace of each type of basket(1pt, 2pt, 3pt) & total points using the list baskets
baskets = [2,2,2,1,2,1,3,3,1,2,2,2,2,1,3]
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
# ## iteration with `range(start)` & `range(start,stop)`
#
# +
# [ ] using range() print "hello" 4 times
# +
# [ ] find spell_list length
# [ ] use range() to iterate each half of spell_list
# [ ] label & print the first and second halves
spell_list = ["Tuesday", "Wednesday", "February", "November", "Annual", "Calendar", "Solstice"]
# +
# [ ] build a list of numbers from 20 to 29: twenties
# append each number to twenties list using range(start,stop) iteration
# [ ] print twenties
twenties = []
# +
# [ ] iterate through the numbers populated in the list twenties and add each number to a variable: total
# [ ] print total
total = 0
# +
# check your answer above using range(start,stop)
# [ ] iterate each number from 20 to 29 using range()
# [ ] add each number to a variable (total) to calculate the sum
# should match earlier task
total = 0
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 3</B></font>
# ## iteration with `range(start:stop:skip)`
#
# +
# [ ] create a list of odd numbers (odd_nums) from 1 to 25 using range(start,stop,skip)
# [ ] print odd_nums
# hint: odd numbers are 2 digits apart
# +
# [ ] create a Decending list of odd numbers (odd_nums) from 25 to 1 using range(start,stop,skip)
# [ ] print odd_nums, output should resemble [25, 23, ...]
# +
# the list, elements, contains the names of the first 20 elements in atomic number order
# [ ] print the even number elements "2 - Helium, 4 - Beryllium,.." in the list with the atomic number
elements = ['Hydrogen', 'Helium', 'Lithium', 'Beryllium', 'Boron', 'Carbon', 'Nitrogen', 'Oxygen', 'Fluorine', \
'Neon', 'Sodium', 'Magnesium', 'Aluminum', 'Silicon', 'Phosphorus', 'Sulfur', 'Chlorine', 'Argon', \
'Potassium', 'Calcium']
# +
# [ ] # the list, elements_60, contains the names of the first 60 elements in atomic number order
# [ ] print the odd number elements "1 - Hydrogen, 3 - Lithium,.." in the list with the atomic number elements_60
elements_60 = ['Hydrogen', 'Helium', 'Lithium', 'Beryllium', 'Boron', 'Carbon', 'Nitrogen', \
'Oxygen', 'Fluorine', 'Neon', 'Sodium', 'Magnesium', 'Aluminum', 'Silicon', \
'Phosphorus', 'Sulfur', 'Chlorine', 'Argon', 'Potassium', 'Calcium', 'Hydrogen', \
'Helium', 'Lithium', 'Beryllium', 'Boron', 'Carbon', 'Nitrogen', 'Oxygen', 'Fluorine', \
'Neon', 'Sodium', 'Magnesium', 'Aluminum', 'Silicon', 'Phosphorus', 'Sulfur', 'Chlorine', \
'Argon', 'Potassium', 'Calcium', 'Scandium', 'Titanium', 'Vanadium', 'Chromium', 'Manganese', \
'Iron', 'Cobalt', 'Nickel', 'Copper', 'Zinc', 'Gallium', 'Germanium', 'Arsenic', 'Selenium', \
'Bromine', 'Krypton', 'Rubidium', 'Strontium', 'Yttrium', 'Zirconium']
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 4</B></font>
# ## combine lists with `+` and `.extend()`
# +
# [ ] print the combined lists (numbers_1 & numbers_2) using "+" operator
numbers_1 = [20, 21, 22, 23, 24, 25, 26, 27, 28, 29]
# pythonic casting of a range into a list
numbers_2 = list(range(30,50,2))
print("numbers_1:",numbers_1)
print("numbers_2",numbers_2)
# +
# [ ] print the combined element lists (first_row & second_row) using ".extend()" method
first_row = ['Hydrogen', 'Helium']
second_row = ['Lithium', 'Beryllium', 'Boron', 'Carbon', 'Nitrogen', 'Oxygen', 'Fluorine', 'Neon']
print("1st Row:", first_row)
print("2nd Row:", second_row)
# -
# ## Project: Combine 3 element rows
# Choose to use **"+" or ".extend()" **to build output similar to
#
# ```
# The 1st three rows of the Period Table of Elements contain:
# ['Hydrogen', 'Helium', 'Lithium', 'Beryllium', 'Boron', 'Carbon', 'Nitrogen', 'Oxygen', 'Fluorine', 'Neon', 'Sodium', 'Magnesium', 'Aluminum', 'Silicon', 'Phosphorus', 'Sulfur', 'Chlorine', 'Argon']
#
# The row breakdown is
# Row 1: Hydrogen, Helium
# Row 2: Lithium, Beryllium, Boron, Carbon, Nitrogen, Oxygen, Fluorine, Neon
# Row 3: Sodium, Magnesium, Aluminum, Silicon, Phosphorus, Sulfur, Chlorine, Argon
# ```
# +
# [ ] create the program: combined 3 element rows
elem_1 = ['Hydrogen', 'Helium']
elem_2 = ['Lithium', 'Beryllium', 'Boron', 'Carbon', 'Nitrogen', 'Oxygen', 'Fluorine', 'Neon']
elem_3 = ['Sodium', 'Magnesium', 'Aluminum', 'Silicon', 'Phosphorus', 'Sulfur', 'Chlorine', 'Argon']
# +
# [ ] .extend() jack_jill with "next_line" string - print the result
jack_jill = ['Jack', 'and', 'Jill', 'went', 'up', 'the', 'hill']
next_line = ['To', 'fetch', 'a', 'pail', 'of', 'water']
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 5</B></font>
# ## .reverse() : reverse a list in place
# +
# [ ] use .reverse() to print elements starting with "Calcium", "Chlorine",... in reverse order
elements = ['Hydrogen', 'Helium', 'Lithium', 'Beryllium', 'Boron', 'Carbon', 'Nitrogen', 'Oxygen', 'Fluorine', \
'Neon', 'Sodium', 'Magnesium', 'Aluminum', 'Silicon', 'Phosphorus', 'Sulfur', 'Chlorine', 'Argon', \
'Potassium', 'Calcium']
# +
# [ ] reverse order of the list... Then print only words that are 8 characters or longer from the now reversed order
spell_list = ["Tuesday", "Wednesday", "February", "November", "Annual", "Calendar", "Solstice"]
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 6</B></font>
# ## .sort() and sorted()
# +
# [ ] sort the list element, so names are in alphabetical order and print elements
elements = ['Hydrogen', 'Helium', 'Lithium', 'Beryllium', 'Boron', 'Carbon', 'Nitrogen', 'Oxygen', 'Fluorine', \
'Neon', 'Sodium', 'Magnesium', 'Aluminum', 'Silicon', 'Phosphorus', 'Sulfur', 'Chlorine', 'Argon', \
'Potassium', 'Calcium']
# +
# [ ] print the list, numbers, sorted and then below print the original numbers list
numbers = [2,2,2,1,2,1,3,3,1,2,2,2,2,1,3]
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 7</B></font>
# ## Converting a string to a list with `.split()`
# +
# [ ] split the string, daily_fact, into a list of word strings: fact_words
# [ ] print each string in fact_words in upper case on it's own line
daily_fact = "Did you know that there are 1.4 billion students in the world?"
# +
# [ ] convert the string, code_tip, into a list made from splitting on the letter "o"
# +
# [ ] split poem on "b" to create a list: poem_words
# [ ] print poem_words by iterating the list
poem = "The bright brain, has bran!"
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 8</B></font>
# ## `.join()`
# ### build a string from a list
# +
# [ ] print a comma separated string output from the list of Halogen elements using ".join()"
halogens = ['Chlorine', 'Florine', 'Bromine', 'Iodine']
# +
# [ ] split the sentence, code_tip, into a words list
# [ ] print the joined words in the list with no spaces in-between
# [ ] Bonus: capitalize each word in the list before .join()
code_tip ="Read code aloud or explain the code step by step to a peer"
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 8</B></font>
# ## `list(string)` & `print("hello",end=' ')`
#
# - **Cast a string to a list**
# - **print to the same line**
long_word = 'neumonoultramicroscopicsilicovolcanoconiosis'
long_word="neumonoultramicroscopicsilicovolcanoconiosis"
letters=list(long_word)
for i in letters:
print(i)
# +
# [ ] use use end= in print to output each string in questions with a "?" and on new lines
string_questions = ["What's the closest planet to the Sun", "How deep do Dolphins swim", "What time is it"]
string_questions =["Whats the closest planet to sun","How deep do Dolphins swim","What time is it"]
for i in string_questions:
print(i,end="?\n")
# +
# [ ] print each item in foot bones
# - capitalized, both words if two word name
# - separated by a comma and space
# - and keeping on a single print line
foot_bones = ["calcaneus", "talus", "cuboid", "navicular", "lateral cuneiform",
"intermediate cuneiform", "medial cuneiform"]
for i in foot_bones:
if " " in i:
print(i.title(),end=", ")
else:
print(i,end=", ")
# -
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Python Fundamentals/Module_3_Practice_Python_Fundamentals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Imports
import numpy as np
import pandas as pd
import torch
import transformers as ppb # pytorch transformers
from sklearn.linear_model import LogisticRegression
# ## Data Extraction
train_data = pd.read_csv('./archive/Train.csv')
test_data = pd.read_csv('./archive/Test.csv')
validation_data = pd.read_csv('./archive/Valid.csv')
train_data.head()
x_tr, y_tr = train_data['text'].values, train_data['label'].values
x_val, y_val = validation_data['text'].values, validation_data['label'].values
# +
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer(max_features=10000)
word_to_ix = vectorizer.fit(train_data.text)
# +
VOCAB_SIZE = len(word_to_ix.vocabulary_)
NUM_LABELS = 2
tr_data_vecs = torch.FloatTensor(word_to_ix.transform(x_tr).toarray())
tr_labels = y_tr.tolist()
val_data_vecs = torch.FloatTensor(word_to_ix.transform(x_val).toarray())
val_labels = y_val.tolist()
tr_data_loader = [(sample, label) for sample, label in zip(tr_data_vecs, tr_labels)]
val_data_loader = [(sample, label) for sample, label in zip(val_data_vecs, val_labels)]
# +
BATCH_SIZE = 20
from torch.utils.data import DataLoader
train_iterator = DataLoader(tr_data_loader,
batch_size=BATCH_SIZE,
shuffle=True,
)
valid_iterator = DataLoader(val_data_loader,
batch_size=BATCH_SIZE,
shuffle=False,
)
# -
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# +
from torch import nn
class BoWClassifier(nn.Module): # inheriting from nn.Module!
def __init__(self, num_labels, vocab_size):
super(BoWClassifier, self).__init__()
self.linear = nn.Linear(vocab_size, num_labels)
def forward(self, bow_vec):
return F.log_softmax(self.linear(bow_vec), dim=1)
# +
INPUT_DIM = VOCAB_SIZE
OUTPUT_DIM = 2
model = BoWClassifier(OUTPUT_DIM, INPUT_DIM)
# +
import torch.optim as optim
optimizer = optim.Adam(model.parameters(), lr=1e-3)
# +
from collections import defaultdict
criterion = nn.NLLLoss()
model = model.to(device)
criterion = criterion.to(device)
metrics = defaultdict(list)
# -
from sklearn.metrics import classification_report
def class_accuracy(preds, y):
rounded_preds = preds.argmax(1)
correct = (rounded_preds == y).float()
acc = correct.sum() / len(correct)
return acc
# +
import torch.nn.functional as F
def train(model, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
model.train()
for texts, labels in iterator:
texts = texts.to(device)
labels = labels.to(device)
optimizer.zero_grad()
predictions = model(texts)
loss = criterion(predictions, labels)
acc = class_accuracy(predictions, labels)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# +
# The evaluation is done on the validation dataset
def evaluate(model, iterator, criterion):
epoch_loss = 0
epoch_acc = 0
# On the validation dataset we don't want training so we need to set the model on evaluation mode
model.eval()
# Also tell Pytorch to not propagate any error backwards in the model
# This is needed when you only want to make predictions and use your model in inference mode!
with torch.no_grad():
# The remaining part is the same with the difference of not using the optimizer to backpropagation
for texts, labels in iterator:
# We copy the text and label to the correct device
texts = texts.to(device)
labels = labels.to(device)
predictions = model(texts)
loss = criterion(predictions, labels)
acc = class_accuracy(predictions, labels)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# +
import time
# This is just for measuring training time!
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# +
# Define the epoch number parameter
N_EPOCHS = 50
best_valid_loss = float('inf')
patience = 2
patience_counter = 0
previous_loss = 100
previous_accuracy = 0
# We loop forward on the epoch number
for epoch in range(N_EPOCHS):
start_time = time.time()
# Train the model on the training set using the dataloader
train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
metrics['train_loss'].append(train_loss)
metrics['train_accuracy'].append(train_acc)
# And validate your model on the validation set
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
metrics['dev_loss'].append(valid_loss)
metrics['dev_accuracy'].append(valid_acc)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
# If we find a better model, we save the weights so later we may want to reload it
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut1-model.pt')
if train_acc <= previous_accuracy and train_loss <= previous_loss:
patience_counter += 1
previous_accuracy = train_acc
previous_loss = train_loss
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
if patience_counter == patience:
break
# -
import matplotlib.pyplot as plt
import seaborn as sns
# +
fig, ax = plt.subplots(1, 2, figsize=(16, 4))
sns.lineplot(data=metrics['train_loss'], ax=ax[0], label='train loss')
sns.lineplot(data=metrics['dev_loss'], ax=ax[0], label='dev loss')
sns.lineplot(data=metrics['train_accuracy'], ax=ax[1], label='train acc')
sns.lineplot(data=metrics['dev_accuracy'], ax=ax[1], label='dev acc')
| NLP-Project(Bag of Words model).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from astropy.io import ascii
import numpy as np
import scipy.stats
import matplotlib
from matplotlib.pyplot import *
# %matplotlib inline
from ecdf import ecdf
cat = ascii.read("matched_cat_0.5arcsec_2arcsec_uniq.cat")
cat = cat[cat['SFR_tot']>0]
cat = cat[cat['gg']!='--']
cat = cat[cat['m20']!='--']
cat = cat[cat['cc']!='--']
cat = cat[cat['aa']!='--']
cat.colnames
zx,zy,zvar = ecdf(cat["z_peak"])
plot(zx,zy)
#semilogy(cat["LMASS"][::20],cat["SFR_tot"][::20],'.')
#ylim(.1,1000)
#xlim(8,11.5)
#MAKE Contour plots
slope, intercept, loslope, highslope = scipy.stats.theilslopes(x=cat['LMASS'][::20],y=np.log10(cat["SFR_tot"][::20]))
plot(np.linspace(8,12,100),slope*np.linspace(8,12,100) + intercept)
hexbin(cat["LMASS"],np.log10(cat["SFR_tot"]))
xlim(8,11.5)
ylim(-1,3)
#now try clustering to get rid of red cloud clump
semilogx(cat["SFR_tot"],cat["gg"],'.')
semilogx(cat["SFR_tot"],cat["m20"],'.')
semilogx(cat["SFR_tot"],cat["cc"],'.')
semilogx(cat["SFR_tot"],cat["aa"],'.')
| cosmos_ultravista_data/mainseq_morphology.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# + [markdown] deletable=true editable=true
# # tmp-API-check
# The `clustergrammer_gl` class is now being loaded into the `Network` class. The class and widget instance are saved in th `Network` instance, `net`. This allows us to load data, cluster, and finally produce a new widget instance using the `widget` method. The instance of the widget is saved in `net` and can be used to grab the data from the clustergram as a Pandas DataFrame using the `widget_df` method. The exported DataFrame will reflect any filtering or imported categories that were added on the front end.
#
# In these examples, we will filter the matrix using the brush crop tool, export the filtered matrix as a DataFrame, and finally visualize this as a new clustergram widget.
# + deletable=true editable=true
import numpy as np
import pandas as pd
from clustergrammer_gl import *
net = Network(clustergrammer_gl)
# + [markdown] deletable=true editable=true
# # Make widget using new API
# + deletable=true editable=true
net.load_file('rc_two_cats.txt')
net.cluster()
net.widget()
# + [markdown] deletable=true editable=true
# Above, we have filtered the matrix to a region of interest using the brush cropping tool. Below we will get export this region of interest, defined on the front end, to a DataFrame, `df_genes`. This demonstrates the two-way communication capabilities of widgets.
# + deletable=true editable=true
df_genes = net.widget_df()
df_genes.shape
# + deletable=true editable=true
net.load_df(df_genes)
net.cluster()
net.widget()
# + [markdown] deletable=true editable=true
# Above, we made a new widget visualizing this region of interest.
# + [markdown] deletable=true editable=true
# # Generate random DataFrame
# Here we will genrate a DataFrame with random data and visualize it using the widget.
# + deletable=true editable=true
# generate random matrix
num_rows = 500
num_cols = 10
np.random.seed(seed=100)
mat = np.random.rand(num_rows, num_cols)
# make row and col labels
rows = range(num_rows)
cols = range(num_cols)
rows = [str(i) for i in rows]
cols = [str(i) for i in cols]
# make dataframe
df = pd.DataFrame(data=mat, columns=cols, index=rows)
# + deletable=true editable=true
net.load_df(df)
net.cluster()
net.widget()
# + [markdown] deletable=true editable=true
# Above, we selected a region of interest using the front-end brush crop tool and export to DataFrame, df_random. Below we will visualize it using a new widget.
# + deletable=true editable=true
df_random = net.widget_df()
df_random.shape
# + deletable=true editable=true
net.load_df(df_random)
net.cluster()
net.widget()
# + deletable=true editable=true
| tmp-API-check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pragmatic color describers
__author__ = "<NAME>"
__version__ = "CS224u, Stanford, Spring 2021"
# ## Contents
#
# 1. [Overview](#Overview)
# 1. [Set-up](#Set-up)
# 1. [The corpus](#The-corpus)
# 1. [Corpus reader](#Corpus-reader)
# 1. [ColorsCorpusExample instances](#ColorsCorpusExample-instances)
# 1. [Displaying examples](#Displaying-examples)
# 1. [Color representations](#Color-representations)
# 1. [Utterance texts](#Utterance-texts)
# 1. [Far, Split, and Close conditions](#Far,-Split,-and-Close-conditions)
# 1. [Toy problems for development work](#Toy-problems-for-development-work)
# 1. [Core model](#Core-model)
# 1. [Toy dataset illustration](#Toy-dataset-illustration)
# 1. [Predicting sequences](#Predicting-sequences)
# 1. [Listener-based evaluation](#Listener-based-evaluation)
# 1. [Other prediction and evaluation methods](#Other-prediction-and-evaluation-methods)
# 1. [Cross-validation](#Cross-validation)
# 1. [Baseline SCC model](#Baseline-SCC-model)
# 1. [Modifying the core model](#Modifying-the-core-model)
# 1. [Illustration: LSTM Cells](#Illustration:-LSTM-Cells)
# 1. [Illustration: Deeper models](#Illustration:-Deeper-models)
# ## Overview
#
# This notebook is part of our unit on grounding. It illustrates core concepts from the unit, and it provides useful background material for the associated homework and bake-off.
# ## Set-up
from colors import ColorsCorpusReader
import os
import pandas as pd
from sklearn.model_selection import train_test_split
import torch
from torch_color_describer import (
ContextualColorDescriber, create_example_dataset)
import utils
from utils import START_SYMBOL, END_SYMBOL, UNK_SYMBOL
utils.fix_random_seeds()
# The [Stanford English Colors in Context corpus](https://cocolab.stanford.edu/datasets/colors.html) (SCC) is included in the data distribution for this course. If you store the data in a non-standard place, you'll need to update the following:
COLORS_SRC_FILENAME = os.path.join(
"data", "colors", "filteredCorpus.csv")
# ## The corpus
# The SCC corpus is based in a two-player interactive game. The two players share a context consisting of three color patches, with the display order randomized between them so that they can't use positional information when communicating.
#
# The __speaker__ is privately assigned a target color and asked to produce a description of it that will enable the __listener__ to identify the speaker's target. The listener makes a choice based on the speaker's message, and the two succeed if and only if the listener identifies the target correctly.
#
# In the game, the two players played repeated reference games and could communicate with each other in a free-form way. This opens up the possibility of modeling these repeated interactions as task-oriented dialogues. However, for this unit, we'll ignore most of this structure. We'll treat the corpus as a bunch of independent reference games played by anonymous players, and we will ignore the listener and their choices entirely.
#
# For the bake-off, we will be distributing a separate test set. Thus, all of the data in the SCC can be used for exploration and development.
# ### Corpus reader
# The corpus reader class is `ColorsCorpusReader` in `colors.py`. The reader's primary function is to let you iterate over corpus examples:
corpus = ColorsCorpusReader(
COLORS_SRC_FILENAME,
word_count=None,
normalize_colors=True)
# The two keyword arguments have their default values here.
#
# * If you supply `word_count` with an interger value, it will restrict to just examples where the utterance has that number of words (using a whitespace heuristic). This creates smaller corpora that are useful for development.
#
# * The colors in the corpus are in [HLS format](https://en.wikipedia.org/wiki/HSL_and_HSV). With `normalize_colors=False`, the first (hue) value is an integer between 1 and 360 inclusive, and the L (lightness) and S (saturation) values are between 1 and 100 inclusive. With `normalize_colors=True`, these values are all scaled to between 0 and 1 inclusive. The default is `normalize_colors=True` because this is a better choice for all the machine learning models we'll consider.
examples = list(corpus.read())
# We can verify that we read in the same number of examples as reported in [Monroe et al. 2017](https://transacl.org/ojs/index.php/tacl/article/view/1142):
# +
# Should be 46994:
len(examples)
# +
# Showing off an example
examples[5].display()
# -
# ### ColorsCorpusExample instances
# The examples are `ColorsCorpusExample` instances:
ex1 = next(corpus.read())
# These objects have a lot of attributes and methods designed to help you study the corpus and use it for our machine learning tasks. Let's review some highlights.
# #### Displaying examples
# You can see what the speaker saw, with the utterance they chose printed above the patches:
ex1.display(typ='speaker')
# This is the original order of patches for the speaker. The target happens to be the leftmost patch, as indicated by the black box around it.
#
# Here's what the listener saw, with the speaker's message printed above the patches:
ex1.display(typ='listener')
# The listener isn't shown the target, of course, so no patches are highlighted.
# If `display` is called with no arguments, then the target is placed in the final position and the other two are given in an order determined by the corpus metadata:
ex1.display()
# This is the representation order we use for our machine learning models.
# #### Color representations
# For machine learning, we'll often need to access the color representations directly. The primary attribute for this is `colors`:
ex1.colors
# In this display order, the third element is the target color and the first two are the distractors. The attributes `speaker_context` and `listener_context` return the same colors but in the order that those players saw them. For example:
ex1.speaker_context
# #### Utterance texts
# Utterances are just strings:
ex1.contents
# There are cases where the speaker made a sequences of utterances for the same trial. We follow [Monroe et al. 2017](https://transacl.org/ojs/index.php/tacl/article/view/1142) in concatenating these into a single utterance. To preserve the original information, the individual turns are separated by `" ### "`. Example 3 is the first with this property – let's check it out:
ex3 = examples[2]
ex3.contents
# The method `parse_turns` will parse this into individual turns:
ss
# For examples consisting of a single turn, `parse_turns` returns a list of length 1:
ex1.parse_turns()
# ### Far, Split, and Close conditions
# The SCC contains three conditions:
#
# __Far condition__: All three colors are far apart in color space. Example:
# +
print("Condition type:", examples[1].condition)
examples[1].display()
# -
# __Split condition__: The target is close to one of the distractors, and the other is far away from both of them. Example:
# +
print("Condition type:", examples[3].condition)
examples[3].display()
# -
# __Close condition__: The target is similar to both distractors. Example:
# +
print("Condition type:", examples[2].condition)
examples[2].display()
# -
# These conditions go from easiest to hardest when it comes to reliable communication. In the __Far__ condition, the context is hardly relevant, whereas the nature of the distractors reliably shapes the speaker's choices in the other two conditions.
#
# You can begin to see how this affects speaker choices in the above examples: "purple" suffices for the __Far__ condition, a more marked single word ("lime") is used in the __Split__ condition, and the __Close__ condition triggers a pretty long, complex description.
# The `condition` attribute provides access to this value:
ex1.condition
# The following verifies that we have the same number of examples per condition as reported in [Monroe et al. 2017](https://transacl.org/ojs/index.php/tacl/article/view/1142):
pd.Series([ex.condition for ex in examples]).value_counts()
# ## Toy problems for development work
# The SCC corpus is fairly large and quite challenging as an NLU task. This means it isn't ideal when it comes to testing hypotheses and debugging code. Poor performance could trace to a mistake, but it could just as easily trace to the fact that the problem is very challenging from the point of view of optimization.
#
# To address this, the module `torch_color_describer.py` includes a function `create_example_dataset` for creating small, easy datasets with the same basic properties as the SCC corpus.
#
# Here's a toy problem containing just six examples:
tiny_contexts, tiny_words, tiny_vocab = create_example_dataset(
group_size=2, vec_dim=2)
tiny_vocab
tiny_words
tiny_contexts
# Each member of `tiny_contexts` contains three vectors. This is meant to be an easy problem, so the final (target) vector always has values that unambiguously determine which utterance is produced. Thus, the model basically just needs to learn to ignore the distractors and find the association between the target vector and the corresponding sequence.
#
# All the models we study have a capacity to solve this task with very little data, so you should see perfect or near perfect performance on reasonably-sized versions of this task.
# ## Core model
# Our core model for this problem is implemented in `torch_color_describer.py` as `ContextualColorDescriber`. At its heart, this is a pretty standard encoder–decoder model:
#
# * `Encoder`: Processes the color contexts as a sequence. We always place the target in final position so that it is closest to the supervision signals that we get when decoding.
#
# * `Decoder`: A neural language model whose initial hidden representation is the final hidden representation of the `Encoder`.
#
# * `EncoderDecoder`: Coordinates the operations of the `Encoder` and `Decoder`.
#
# Finally, `ContextualColorDescriber` is a wrapper around these model components. It handles the details of training and implements the prediction and evaluation functions that we will use.
#
# Many additional details about this model are included in the slides for this unit.
# ### Toy dataset illustration
# To highlight the core functionality of `ContextualColorDescriber`, let's create a small toy dataset and use it to train and evaluate a model:
toy_color_seqs, toy_word_seqs, toy_vocab = create_example_dataset(
group_size=50, vec_dim=2)
toy_color_seqs_train, toy_color_seqs_test, toy_word_seqs_train, toy_word_seqs_test = \
train_test_split(toy_color_seqs, toy_word_seqs)
# `ContextualColorDescriber` is a subclass of `TorchModelBase`, so all of the optimization parameters from that model are available here; see [torch_model_base.py](torch_model_base.py) for full details.
#
# Here is a simple use of `ContextualColorDescriber`:
toy_mod = ContextualColorDescriber(toy_vocab, max_iter=200)
_ = toy_mod.fit(toy_color_seqs_train, toy_word_seqs_train)
# ### Predicting sequences
# The `predict` method takes a list of color contexts as input and returns model descriptions:
toy_preds = toy_mod.predict(toy_color_seqs_test)
toy_preds[0]
# We can then check that we predicted all correct sequences:
# +
toy_correct = sum(1 for x, p in zip(toy_word_seqs_test, toy_preds) if x == p)
toy_correct / len(toy_word_seqs_test)
# -
# For real problems, this is too stringent a requirement, since there are generally many equally good descriptions. This insight gives rise to metrics like [BLEU](https://en.wikipedia.org/wiki/BLEU), [METEOR](https://en.wikipedia.org/wiki/METEOR), [ROUGE](https://en.wikipedia.org/wiki/ROUGE_(metric)), [CIDEr](https://arxiv.org/pdf/1411.5726.pdf), and others, which seek to relax the requirement of an exact match with the test sequence. These are reasonable options to explore, but we will instead adopt a communcation-based evaluation, as discussed in the next section.
# ### Listener-based evaluation
# `ContextualColorDescriber` implements a method `listener_accuracy` that we will use for our primary evaluations in the assignment and bake-off. The essence of the method is that we can calculate
#
# $$
# c^{*} = \text{argmax}_{c \in C} P_S(\text{utterance} \mid c)
# $$
#
#
# where $P_S$ is our describer model and $C$ is the set of all permutations of all three colors in the color context. We take $c^{*}$ to be a correct prediction if it is one where the target is in the privileged final position. (There are two such contexts; we try both in case the order of the distractors influences the predictions, and the model is correct if one of them has the highest probability.)
#
# Here's the listener accuracy of our toy model:
toy_mod.listener_accuracy(toy_color_seqs_test, toy_word_seqs_test)
# ### BLEU scores
# The listener-based evaluation scheme has the unusual property that, in some sense, it assesses the model's ability to communicate with itself. This creates a danger that it will drift far from English as we know it but still succeed in signaling the target color. Ideally, we would train a separate listener model to help prevent this, but doing so would be cumbersome and could limit creative system development. Thus, as a quick check that our systems are still going to be able to communicate with us, we can calculate a BLEU score:
# +
bleu_score, predicted_texts = toy_mod.corpus_bleu(toy_color_seqs_test, toy_word_seqs_test)
bleu_score
# -
# For discussion of BLEU scores, see the [evaluation metrics notebook](evaluation_metrics.ipynb).
# ### Other prediction and evaluation methods
# You can get the perplexities for test examples with `perpelexities`:
toy_perp = toy_mod.perplexities(toy_color_seqs_test, toy_word_seqs_test)
toy_perp[0]
# You can use `predict_proba` to see the full probability distributions assigned to test examples:
toy_proba = toy_mod.predict_proba(toy_color_seqs_test, toy_word_seqs_test)
# +
# 4 tokens, each assigned a distribution over 5 vocab items:
print(toy_word_seqs_test[0])
toy_proba[0].shape
# -
for timestep in toy_proba[0]:
print(dict(zip(toy_vocab, timestep)))
# ### Cross-validation
# You can use `utils.fit_classifier_with_hyperparameter_search` to cross-validate these models. Just be sure to set `scoring=None` so that the sklearn model selection methods use the `score` method of `ContextualColorDescriber`, which is an alias for `listener_accuracy`:
best_mod = utils.fit_classifier_with_hyperparameter_search(
toy_color_seqs_train,
toy_word_seqs_train,
toy_mod,
cv=2,
scoring=None,
param_grid={'hidden_dim': [10, 20]})
# ## Baseline SCC model
# Just to show how all the pieces come together, here's a very basic SCC experiment using the core code and very simplistic assumptions (which you will revisit in the assignment) about how to represent the examples:
# To facilitate quick development, we'll restrict attention to the two-word examples:
dev_corpus = ColorsCorpusReader(COLORS_SRC_FILENAME, word_count=2)
dev_examples = list(dev_corpus.read())
len(dev_examples)
# Here we extract the raw colors and texts (as strings):
dev_cols, dev_texts = zip(*[[ex.colors, ex.contents] for ex in dev_examples])
# To tokenize the examples, we'll just split on whitespace, taking care to add the required boundary symbols:
dev_word_seqs = [[START_SYMBOL] + text.split() + [END_SYMBOL] for text in dev_texts]
dev_word_seqs
# We'll use a random train–test split:
dev_cols_train, dev_cols_test, dev_word_seqs_train, dev_word_seqs_test = \
train_test_split(dev_cols, dev_word_seqs)
# Our vocab is determined by the train set, and we take care to include the `$UNK` token:
# +
dev_vocab = sorted({w for toks in dev_word_seqs_train for w in toks})
dev_vocab += [UNK_SYMBOL]
# -
# And now we're ready to train a model:
dev_mod = ContextualColorDescriber(
dev_vocab,
embed_dim=10,
hidden_dim=10,
early_stopping=True)
# %time _ = dev_mod.fit(dev_cols_train, dev_word_seqs_train)
# And finally an evaluation in terms of listener accuracy and BLEU scores. The `evaluate` method combines these:
dev_mod_eval = dev_mod.evaluate(dev_cols_test, dev_word_seqs_test)
dev_mod_eval['listener_accuracy']
dev_mod_eval['corpus_bleu']
# ## Modifying the core model
# The first few assignment problems concern how you preprocess the data for your model. After that, the goal is to subclass model components in `torch_color_describer.py`. For the bake-off submission, you can do whatever you like in terms of modeling, but my hope is that you'll be able to continue subclassing based on `torch_color_describer.py`.
#
# This section provides some illustrative examples designed to give you a feel for how the code is structured and what your options are in terms of creating subclasses. The principles are the same as those reviewed for a wider range of models in [tutorial_pytorch_models.ipynb](tutorial_pytorch_models.ipynb).
# ### Illustration: LSTM Cells
# Both the `Encoder` and the `Decoder` of `torch_color_describer` are currently GRU cells. Switching to another cell type is easy:
# __Step 1__: Subclass the `Encoder`; all we have to do here is change `GRU` from the original to `LSTM`:
# +
import torch.nn as nn
from torch_color_describer import Encoder
class LSTMEncoder(Encoder):
def __init__(self, color_dim, hidden_dim):
super().__init__(color_dim, hidden_dim)
self.rnn = nn.LSTM(
input_size=self.color_dim,
hidden_size=self.hidden_dim,
batch_first=True)
# -
# __Step 2__: Subclass the `Decoder`, making the same simple change as above:
# +
import torch.nn as nn
from torch_color_describer import Encoder, Decoder
class LSTMDecoder(Decoder):
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self.rnn = nn.LSTM(
input_size=self.embed_dim,
hidden_size=self.hidden_dim,
batch_first=True)
# -
# __Step 3__:`ContextualColorDescriber` has a method called `build_graph` that sets up the `Encoder` and `Decoder`. The needed revision just uses `LSTMEncoder`:
# +
from torch_color_describer import EncoderDecoder
class LSTMContextualColorDescriber(ContextualColorDescriber):
def build_graph(self):
# Use the new Encoder:
encoder = LSTMEncoder(
color_dim=self.color_dim,
hidden_dim=self.hidden_dim)
# Use the new Decoder:
decoder = LSTMDecoder(
vocab_size=self.vocab_size,
embed_dim=self.embed_dim,
embedding=self.embedding,
hidden_dim=self.hidden_dim)
return EncoderDecoder(encoder, decoder)
# -
# Here's an example run:
lstm_mod = LSTMContextualColorDescriber(
toy_vocab,
embed_dim=10,
hidden_dim=10)
_ = lstm_mod.fit(toy_color_seqs_train, toy_word_seqs_train)
lstm_mod.listener_accuracy(toy_color_seqs_test, toy_word_seqs_test)
# ### Illustration: Deeper models
# The `Encoder` and `Decoder` are both currently hard-coded to have just one hidden layer. It is straightforward to make them deeper as long as we ensure that both the `Encoder` and `Decoder` have the same depth; since the `Encoder` final states are the initial hidden states for the `Decoder`, we need this alignment.
#
# (Strictly speaking, we could have different numbers of `Encoder` and `Decoder` layers, as long as we did some kind of averaging or copying to achieve the hand-off from `Encoder` to `Decocer`. I'll set this possibility aside.)
# __Step 1__: We need to subclass the `Encoder` and `Decoder` so that they have `num_layers` argument that is fed into the RNN cell:
# +
import torch.nn as nn
from torch_color_describer import Encoder, Decoder
class DeepEncoder(Encoder):
def __init__(self, *args, num_layers=2, **kwargs):
super().__init__(*args, **kwargs)
self.num_layers = num_layers
self.rnn = nn.GRU(
input_size=self.color_dim,
hidden_size=self.hidden_dim,
num_layers=self.num_layers,
batch_first=True)
class DeepDecoder(Decoder):
def __init__(self, *args, num_layers=2, **kwargs):
super().__init__(*args, **kwargs)
self.num_layers = num_layers
self.rnn = nn.GRU(
input_size=self.embed_dim,
hidden_size=self.hidden_dim,
num_layers=self.num_layers,
batch_first=True)
# -
# __Step 2__: As before, we need to update the `build_graph` method of `ContextualColorDescriber`. The needed revision just uses `DeepEncoder` and `DeepDecoder`. To expose this new argument to the user, we also add a new keyword argument to `ContextualColorDescriber`:
# +
from torch_color_describer import EncoderDecoder
class DeepContextualColorDescriber(ContextualColorDescriber):
def __init__(self, *args, num_layers=2, **kwargs):
self.num_layers = num_layers
super().__init__(*args, **kwargs)
def build_graph(self):
encoder = DeepEncoder(
color_dim=self.color_dim,
hidden_dim=self.hidden_dim,
num_layers=self.num_layers) # The new piece is this argument.
decoder = DeepDecoder(
vocab_size=self.vocab_size,
embed_dim=self.embed_dim,
embedding=self.embedding,
hidden_dim=self.hidden_dim,
num_layers=self.num_layers) # The new piece is this argument.
return EncoderDecoder(encoder, decoder)
# -
# An example/test run:
mod_deep = DeepContextualColorDescriber(
toy_vocab,
embed_dim=10,
hidden_dim=10)
_ = mod_deep.fit(toy_color_seqs_train, toy_word_seqs_train)
mod_deep.listener_accuracy(toy_color_seqs_test, toy_word_seqs_test)
| colors_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="p4yIVZCyJyL0" executionInfo={"status": "ok", "timestamp": 1604559119722, "user_tz": -540, "elapsed": 757, "user": {"displayName": "\uc720\ub2e4\ube48", "photoUrl": "", "userId": "17470031872640109060"}}
import pandas as pd
import requests
from bs4 import BeautifulSoup
# + id="qjh1_PrBJ0tW" executionInfo={"status": "ok", "timestamp": 1604564388393, "user_tz": -540, "elapsed": 732, "user": {"displayName": "\uc720\ub2e4\ube48", "photoUrl": "", "userId": "17470031872640109060"}}
def total_pages(url):
# page 찾기
naver_movie_url = url
# request 보냄.
response = requests.get(naver_movie_url)
# HTML 텍스트 추출
html = response.text.strip()
# BeautifulSoup 객체 생성
soup = BeautifulSoup(markup=html, features='html5lib')
# 찾고자 하는 element의 selector
page_selector = 'div.score_total em'
# element 찾기
search_pages = soup.select(page_selector)
for link in search_pages:
if ',' in link.text:
a = link.text
a1 = a.split(',')[0]
a2 = a.split(',')[1]
total_pages = int(a1+a2)
else:
total_pages = int(link.text)
pages = int(total_pages/10)
# print(pages)
return pages
# + id="BmCBn0NyJ2CU" executionInfo={"status": "ok", "timestamp": 1604565120056, "user_tz": -540, "elapsed": 751, "user": {"displayName": "\uc720\ub2e4\ube48", "photoUrl": "", "userId": "17470031872640109060"}}
def crawling_movie(page, crawling_url, title):
score_list = []
review_list = []
for page in range(1, pages + 1): # pages + 1
# URL
naver_movie_url = crawling_url + f'{page}'
# print(naver_movie_url) # 확인용
# request 보냄.
response = requests.get(naver_movie_url)
# print(response)
# HTML 텍스트 추출
html = response.text.strip()
# BeautifulSoup 객체 생성
soup = BeautifulSoup(markup=html, features='html5lib')
# 찾고자 하는 element의 selector
## 평점 찾기
score_selector = 'div.star_score em'
# body > div > div > div.score_result > ul > li:nth-child(1) > div.star_score > em
# element 찾기
search_scores = soup.select(score_selector)
# 찾고자 하는 element의 selector
## 리뷰 찾기
for num in range(0, 11):
review_selector = f'div.score_reple p span#_filtered_ment_{num}'
#_filtered_ment_0
# element 찾기
search_reviews = soup.select(review_selector)
for review in search_reviews:
review_list.append(review.text.strip())
review = pd.Series(review_list, name='review')
for score in search_scores:
score_list.append(score.text)
score = pd.Series(score_list, name='score')
# 출력
# for score, review in zip(score_list, review_list):
# print(score, review)
movie_review = pd.merge(score, review, left_index=True, right_index=True)
movie_review.to_csv(f'movie_review_{title}.csv', index=False)
# + [markdown] id="PqJUfhFkNArt"
# 1. https://movie.naver.com/
# 2. 아무 영화 클릭
# 3. 아래 리뷰 탭 클릭
# 4. url을 아래와 같이 review부분을 pointWriteFormList로 바꿔 주세요!
# * https://movie.naver.com/movie/bi/mi/review.nhn?code=193194
# * https://movie.naver.com/movie/bi/mi/pointWriteFormList.nhn?code=136990&type=after
# 5. 첫번째 함수 url : 위에서 바꾼 url
# 6. 두번째 함수 crawling_url :
# * f'https://movie.naver.com/movie/bi/mi/pointWriteFormList.nhn?code=193194&page=' 이 형태로 저장해서 함수 실행시켜주세요!
# 7. 두번쨰 함수에서 title은 파일 제목이에요!
# + [markdown] id="4acmB0PijfIZ"
# ### 도굴
# + id="d3StekeJNDzD" executionInfo={"status": "ok", "timestamp": 1604559590388, "user_tz": -540, "elapsed": 1853, "user": {"displayName": "\uc720\ub2e4\ube48", "photoUrl": "", "userId": "17470031872640109060"}} outputId="f36c3f17-49ee-403a-f54a-1d2cffd859c8" colab={"base_uri": "https://localhost:8080/"}
naver_movie_page_url = 'https://movie.naver.com/movie/bi/mi/pointWriteFormList.nhn?code=193194&page=1'
pages = int(total_pages(naver_movie_page_url))
pages
# + id="11FpnBAgLcWu"
naver_movie_url = 'https://movie.naver.com/movie/bi/mi/pointWriteFormList.nhn?code=193194&page='
crawling_movie(pages, naver_movie_url, a)
# + [markdown] id="Gzl7Qu6Ch6Ya"
# ### 곰돌이 푸 다시만나 행복해
# + id="kjefu3gQh6Cz" executionInfo={"status": "ok", "timestamp": 1604565083089, "user_tz": -540, "elapsed": 1839, "user": {"displayName": "\uc720\ub2e4\ube48", "photoUrl": "", "userId": "17470031872640109060"}} outputId="f397f042-503c-4fb4-a1c6-93fa18e1ae5d" colab={"base_uri": "https://localhost:8080/"}
naver_movie_page_url = 'https://movie.naver.com/movie/bi/mi/pointWriteFormList.nhn?code=168023&page=1'
pages = int(total_pages(naver_movie_page_url))
pages
# + id="HYTEBtv_iM5R"
naver_movie_url = 'https://movie.naver.com/movie/bi/mi/pointWriteFormList.nhn?code=168023&page='
crawling_movie(pages, naver_movie_url, 'pooh')
# + id="sG6BjkuTidZV"
| google_colab/movie_crawling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center> Interpolación de Lagrange <center>
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import Image
# ## 1. Ejemplo 3 puntos
#Hacemos la interpolación con tres puntos de la tabla
#(205,1724.3);(210,1907.7);(215,2105.9)
X=np.array([205,210,215])
Y=np.array([1724.3,1907.7,2105.9])
datos=pd.read_csv("vapor_Saturado.csv",delimiter=",")
print(datos)
X=datos['T(°C)']
# ## Formula para los coeficientes de interpolación:
# x: Punto a interpolar ; n: número de puntos-1
# $$ L_{i}(x)=\Pi_{j=0,j\neq i}^{n}\frac{(x-x_{j})}{(x_{i}-x_{j})} $$ $ i=0,...,n$
# $$ p(x)=\sum_{i=0}^{n}y_{i}*L_{i}(x)$$
#Coeficientes de p(x
#x: punto a interpolar
#X: datos en x, Y datos en y
def L0(x,X):#i=0, j=1,2
return (x-X[1])*(x-X[2])/((X[0]-X[1])*(X[0]-X[2]))
def L1(x,X):#i=1,j=0,2
return (x-X[0])*(x-X[2])/((X[1]-X[0])*(X[1]-X[2]))
def L2(x,X):#i=1,j=0,1
return (x-X[0])*(x-X[1])/((X[2]-X[0])*(X[2]-X[1]))
#sumatoria para el polinomio
def p_lagrange(x,X,Y):
return L0(x,X)*Y[0]+L1(x,X)*Y[1]+L2(x,X)*Y[2]
x=207.5#K; valor a interpolar
p_lagrange(x,X,Y)
import sympy as sy
x=sy.symbols('x')
L0(x,X)
print(L1(x,X))
print(L2(x,X))
print(p_lagrange(x,X,Y))
sy.expand(p_lagrange(x,X,Y))
# ## 2. Interpolación de P vs T y generalización programa Lagrange
#base del polinomio L_i
def L(i,x,X,n):#i=0, j=1,2
prod=1
for j in range(n):
if(i!=j):
prod*=(x-X[j])/(X[i]-X[j])
return prod
#Calculo de los polinomios haciendo una sumatoria de los productos L_iy_i
def p_lagrange(x,X,Y):
n=len(X)
suma=0
for i in range(n):
# print(type(L(i,x,X,n)))
# print(type(Y[i]))
suma+=L(i,x,X,n)*Y[i]
return suma
# ## Datos
# Tomados de la tabla A-5 del Cengel para las propiedades del agua saturada, obtenidos mediante cálculoscon la formulación de la IAPWS para las propiedades del agua.
# https://doi.org/10.1063/1.1461829 \
# http://www.iapws.org/newform.html
datos=pd.read_csv("thermodynamical_tables_water.csv",delimiter="\t",header=[0,1,2])
datos
# Opcional: Modificamos los indices para que queden los multiíndices de forma adecuada
a=[x[j] for x in datos.columns for j in range(3)]
a=np.reshape(a,[int(len(a)/3),3])
a[3][0]=a[2][0]
a[5][0]=a[4][0]
a[6][0]=a[4][0]
a[8][0]=a[7][0]
a[9][0]=a[7][0]
a[11][0]=a[10][0]
a[12][0]=a[10][0]
a=[tuple(a[i]) for i in range(len(a))]
col_names =pd.MultiIndex.from_tuples(a)
datos.columns=col_names
datos
T=datos["Temp. (°C)"]["Temp."]["T"]
P=datos["Sat. press. (kPa)"]["Sat. press."]["Psat"]
v_f=datos['Specific volume (m3/kg)']['Sat. liquid.']['v_f']
v_g=datos['Specific volume (m3/kg)']['Sat.vapor.']['v_g']
s_f=datos['Entropy (kJ/kg·K)']['Sat. liquid.']['s_f']
s_fg=datos['Entropy (kJ/kg·K)']['Evap.']['s_fg']
s_g=datos['Entropy (kJ/kg·K)']['Sat.vapor.']['s_g']
T_inter=np.linspace(7.5,372.5,74)
# ### Tabla interpolación P vs T
P_pred_complete=p_lagrange(T_inter,T,P)#[p_lagrange(t,T,P) for t in T_inter]
results_complete=pd.DataFrame({"T(°C)":T_inter,"P predicted (kPa)":P_pred_complete})
P_pred_pag914=p_lagrange(T_inter[40:],np.array(T[40:]),np.array(P[40:]))#[p_lagrange(t,T,P) for t in T_inter]
# P_pred_pag914=p_lagrange(T_inter[50:-20],np.array(T[50:-20]),np.array(P[50:-20]))
# results_pag914=pd.DataFrame({"T(°C)":T_inter[40:],"P predicted (kPa)":P_pred_pag914})
print(results_complete)
# ### 5. a) Gráfica
plt.figure()
plt.plot(T_inter[12:-13],P_pred_complete[12:-13])
plt.scatter(T,P)
plt.xlabel('T(°C)')
plt.ylabel('P(kPa)')
plt.savefig("P_vs_T_completo.png")
plt.figure()
plt.plot(T_inter[41:-2],P_pred_pag914[1:-2])
plt.xlabel('T(°C)')
plt.ylabel('P(kPa)')
# plt.plot(T_inter[50:-20],P_pred_pag914[:])
plt.scatter(T[40:],P[40:])
plt.savefig("P_vs_T_pag_914.png")
# ### 3. Tabla interpolación $v_f$ vs T
v_f_pred_complete=p_lagrange(T_inter,T,v_f)#[p_lagrange(t,T,P) for t in T_inter]
results_complete=pd.DataFrame({"T(°C)":T_inter,"v_f predicted (m^3/kg)":v_f_pred_complete})
v_f_pred_pag914=p_lagrange(T_inter[40:],np.array(T[40:]),np.array(v_f[40:]))#[p_lagrange(t,T,P) for t in T_inter]
# results_pag914=pd.DataFrame({"T(°C)":T_inter[40:],"P predicted (kPa)":P_pred_pag914})
print(results_complete)
# ### 5. b) Gráfica
plt.figure()
plt.plot(T_inter[13:-15],v_f_pred_complete[13:-15])
plt.scatter(T,v_f)
plt.savefig("v_f_vs_T_completo.png")
plt.xlabel('T(°C)')
plt.ylabel('v_f(m^3/kg)')
plt.figure()
plt.plot(T_inter[42:-3],v_f_pred_pag914[2:-3])
plt.scatter(T[40:],v_f[40:])
plt.xlabel('T(°C)')
plt.ylabel('v_f(m^3/kg)')
plt.savefig("v_f_vs_T_pag_914.png")
# ### 4. Tabla interpolación $v_g$ vs T
v_g_pred_complete=p_lagrange(T_inter,T,v_g)#[p_lagrange(t,T,P) for t in T_inter]
results_complete=pd.DataFrame({"T(°C)":T_inter,"v_g predicted (m^3/kg)":v_g_pred_complete})
v_g_pred_pag914=p_lagrange(T_inter[40:],np.array(T[40:]),np.array(v_g[40:]))#[p_lagrange(t,T,P) for t in T_inter]
# results_pag914=pd.DataFrame({"T(°C)":T_inter[40:],"P predicted (kPa)":P_pred_pag914})
print(results_complete)
# ### 5. c) Gráfica
plt.figure()
plt.plot(T_inter[13:-13],v_g_pred_complete[13:-13])
plt.scatter(T,v_g)
plt.xlabel('T(°C)')
plt.ylabel('v_g(m^3/kg)')
plt.savefig("v_g_vs_T_completo.png")
plt.figure()
plt.plot(T_inter[41:-2],v_g_pred_pag914[1:-2])
plt.scatter(T[40:],v_g[40:])
plt.xlabel('T(°C)')
plt.ylabel('v_g(m^3/kg)')
plt.savefig("v_g_vs_T_pag_914.png")
# ## Laboratorio métodos
# ### 3. Tabla interpolación $s_f$ vs T
s_f_pred_complete=p_lagrange(T_inter,T,s_f)#[p_lagrange(t,T,P) for t in T_inter
results_complete=pd.DataFrame({"T(°C)":T_inter,"s_f predicted (m^3/kg)":s_f_pred_complete})
print(results_complete)
# ### 5. b) Gráfica
plt.figure()
plt.plot(T_inter[13:-14],s_f_pred_complete[13:-14])
plt.scatter(T,s_f)
plt.xlabel('T(°C)')
plt.ylabel('s_f(kJ/(kg.K))')
plt.savefig("s_f_vs_T_completo.png")
# +
s_fg_pred_complete=p_lagrange(T_inter,T,s_fg)#[p_lagrange(t,T,P) for t in T_inter
results_complete=pd.DataFrame({"T(°C)":T_inter,"s_fg predicted (m^3/kg)":s_fg_pred_complete})
print(results_complete)
plt.figure()
plt.plot(T_inter[13:-14],s_fg_pred_complete[13:-14])
plt.scatter(T,s_fg)
plt.xlabel('T(°C)')
plt.ylabel('s_fg(kJ/(kg.K))')
plt.savefig("s_fg_vs_T_completo.png")
# -
# ### 4. Tabla interpolación $s_g$ vs T
# +
s_g_pred_complete=p_lagrange(T_inter,T,s_g)#[p_lagrange(t,T,P) for t in T_inter]
results_complete=pd.DataFrame({"T(°C)":T_inter,"s_g predicted (m^3/kg)":s_g_pred_complete})
print(results_complete)
# -
# ### 5. c) Gráfica
plt.figure()
plt.plot(T_inter[12:-13],s_g_pred_complete[12:-13])
plt.scatter(T,s_g)
plt.xlabel('T(°C)')
plt.ylabel('s_f(kJ/(kg.K))')
plt.savefig("s_g_vs_T_completo.png")
# ## 6 a) Resultados y análisis
# Observamos en prácticamente todas las gráficas que los polinomios de interpolación de Lagrange muestran resultados similares a los valores proporcionados en el apéndice, sin embargo en los puntos extremos el polinomio de Lagrange tiene a diverger oscilatoriamente. Este fenómeno se conoce como el fenómeno de Runge, el cual ocurre porque aunque exista un polinomio de interpolación a la función verdadera. no se conoce si el polinomio de Lagrange se acercará a dicho polinomio. Además, al tener un número considerable de datos n=76, el polinomio interpolado va a tener una derivada de alto orden que va a ir aumentando si aumentamos el número de puntos y nos alejamos del origen, con lo cual las oscilaciones van a aumentar a medida que nos alejamos de los datos.
plt.figure()
plt.plot(s_f_pred_complete[15:-16],T_inter[15:-16],c='r',label="Interpolación s_f")
plt.scatter(s_f,T,c='tab:blue',label='Datos')
plt.plot(s_g_pred_complete[15:-16],T_inter[15:-16],c='g',label="Interpolación s_g")
plt.scatter(s_g,T,c='tab:blue')
plt.ylabel('T (°C)')
plt.xlabel('s (kJ/(kg.K))')
plt.legend()
plt.savefig("diagrama_TS_completo.png")
Image(url= "TS_diagram_cengel.png", width=600, height=600)
# Por otro lado, al considerar el diagrama TS (T vs s_f y T vs s_g) tanto para los datos como para el polinomido de interpolación, vemos que se puede reproducir el diagrama TS para la región del vapor saturado y que corresponde a la curva de color negro en el Cengel p 924. Para las regiones interiores a la curva se debe conocer los porcentajes que hay en la mezcla de f (fluido) con g (gas) de agua. Si miramos la interpolación, vemos que no el polinomio no puede predecir valores de temperatura bajos ni valores cercanos al punto crítico (donde coexisten las fases)
# ## 6 b) Conclusiones
# * Exceptuando los primeros 16 y los últimos 16 datos, el polinomios de interpolación de Lagrange muestran resultados similares a los valores proporcionados en el apéndice.
# * En los puntos extremos el polinomio de Lagrange presenta el fenómeno de Runge, observándose oscilaciones.
# * El polinomio de Lagrange es útil para una cantidad baja de puntos, como 3 puntos.
# * Se puede revisar el error porcentual para conocer el margen de error con respecto a los teóricos y no solo estimar la diferencia en orden de magnitud o gráficamente.
# * Se pueden realizar pruebas con diferente cantidad de puntos para conocer cuando es suficientemente buena la interpolación de Lagrange.
# * Los puntos de la entropía proporcionan una mejor interpretación de los datos para el análisis del estudiante puesto que existen diagramas TS en la literatura, así como diagramas HS (diagrama de Mollier)
# * Para una primera instancia se recomienda realizar análisis con las tablas de gas ideal para estudiantes novicios, para los otros se pueden dejar estos datos y/o analizar las tablas de vapor sobrecalentada (curvas de nivel)
| Act_4_lagrange/.ipynb_checkpoints/interpolacion-checkpoint.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .rs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Rust
// language: rust
// name: rust
// ---
// :dep showata = "0.1.0"
// :dep mime = "*"
extern crate showata;
extern crate mime;
use showata::*;
std::fs::read_dir("res").unwrap().for_each(|x| println!("{:?}", x));
show_file_in_jupyter("res/rustacean-flat-happy.png", mime::IMAGE_PNG, false)
show_file_in_jupyter("res/rustacean-flat-happy.svg", mime::IMAGE_SVG, true)
| examples/sample_file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp cifar_loader
# -
# # CIFAR Loader
#
# STATUS: SUPER EARLY ALPHA
#
# This package is NOT yet ready for PUBLIC CONSUMPTION. Use at your own RISK!!!!
#
# Everything, including the API (and even
# the existence of this module) are subject
# to breaking change...
#
# These are utilities
#colab
# !pip install -Uqq git+https://github.com/fastai/fastai.git
#colab
# !curl -s https://course19.fast.ai/setup/colab | bash
# !pip freeze | grep torch
# !pip freeze | grep fast
#
# Start of kernel
#exporti
import numpy as np
import torchvision.datasets.utils as tv_utils
import torchvision.datasets.cifar as cifar_dsets
import pickle
#export
def download_cifar10_dsets(path):
"""Download cifar10 datasets using torchvision utils
Arguments:
path (pathlib.Path): path to download the dataset (aka root)
"""
train_dataset = cifar_dsets.CIFAR10(
root=path,
train=True,
download=True
)
test_dataset = cifar_dsets.CIFAR10(
root=path,
train=False,
download=True,
)
return train_dataset,test_dataset
#hide
from fastai.data.external import Config
from pathlib import Path
# patch pathlib with fastai goodies
import fastcore.xtras
cfg = Config()
cifar_root = cfg.data_path/'cifar'
# cifar_root = Path('/tmp/cifar')
cifar_root.mkdir(parents=True,exist_ok=True)
cifar_root.ls()
_ = download_cifar10_dsets(cifar_root)
cifar_root.ls()
#export
def load_cifar_items(downloaded_list, path, check=False):
"""loads cifar test/train items into tuple(data, target)
scrobbled together from torch.data.utils.datasets.CIFAR10 code
https://pytorch.org/docs/stable/_modules/torchvision/datasets/cifar.html#CIFAR10
Arguments:
downloaded_list : a list of file names with their checksum, see CIFAR10.train_list or CIFAR10.test_list.
path (pathlib.Path): the root path where the dataset was downloaded
check(bool, optional): whether to perform an integrity check on the downloaded files (default: False)
"""
data = []
targets = []
# now load the picked numpy arrays
for file_name, checksum in downloaded_list:
file_path = path/cifar_dsets.CIFAR10.base_folder/file_name
if check and not tv_utils.check_integrity(file_path, checksum):
raise RuntimeError(
f'Data checksum failed for file:{file_path} checksum:{checksum}')
with open(file_path, 'rb') as f:
entry = pickle.load(f, encoding='latin1')
data.append(entry['data'])
if 'labels' in entry:
targets.extend(entry['labels'])
else:
targets.extend(entry['fine_labels'])
data = np.vstack(data).reshape(-1, 3, 32, 32)
data = data.transpose((0, 2, 3, 1)) # convert to HWC
return data, targets
#hide
tst_data, tst_targets = load_cifar_items(cifar_dsets.CIFAR10.test_list,cifar_root, check=True)
#hide
from fastcore.test import *
#hide
test_eq(tst_data.shape,(10000,32,32,3))
test_eq(len(tst_targets),10000)
assert isinstance(tst_data,np.ndarray)
assert isinstance(tst_targets[0], int)
# +
#export
# TODO: incorporate list of classes into dataloaders vocab and decodes
from fastcore.foundation import L
def load_classes(path):
"""Load classes to used to map categories to target labels"""
base_folder = cifar_dsets.CIFAR10.base_folder
meta = cifar_dsets.CIFAR10.meta
file_path = path/base_folder/meta['filename']
if not tv_utils.check_integrity(file_path, meta['md5']):
raise RuntimeError('Dataset metadata file not found or corrupted.' +
' You can use download=True to download it')
data = {}
with open(file_path, 'rb') as infile:
data = pickle.load(infile, encoding='latin1')
classes = data[meta['key']]
# class_for i, _class in enumerato_idx = {_class: i te(classes)}
return L(classes)
# -
#hide
classes = load_classes(cifar_root)
#hide
test_eq(len(classes),10); classes
#hide
train_items = load_cifar_items(cifar_dsets.CIFAR10.train_list, cifar_root, check=True)
#exporti
from fastcore.transform import Transform
import torchvision.transforms.functional as TVF
import torch
import PIL
#export
class CifarNP2ImageTransform(Transform):
def encodes(self, o:np.ndarray) -> None:
return PIL.Image.fromarray(o)
#export
class Int2TensorTransform(Transform):
def encodes(self, o: int) -> None:
return torch.tensor(o)
#export
class CifarImageTransform(Transform):
def encodes(self, o: PIL.Image) -> None:
return TVF.to_tensor(o)
#export
class CifarImage2FloatTransform(Transform):
def encodes(self, o: torch.Tensor) -> None:
return o.float().div_(255.)
#exporti
import torchvision as thv
#export
def make_torch_tfms():
norm = thv.transforms.Normalize(
mean=(0.4914, 0.4822, 0.4465), std=(0.2023, 0.1994, 0.2010))
transform_train = thv.transforms.Compose([
thv.transforms.RandomCrop(32, padding=4),
thv.transforms.RandomHorizontalFlip(),
thv.transforms.ToTensor(),
norm,
])
transform_test = thv.transforms.Compose([
thv.transforms.ToTensor(),
norm,
])
return transform_train, transform_test
th_train_tfms, th_test_tfms = make_torch_tfms()
#exporti
from fastcore.transform import ItemTransform
from fastcore.basics import store_attr
#export
class CifarTupleTransform(ItemTransform):
def __init__(self, x_tfm, y_tfm):
store_attr()
def encodes(self,xy):
return [self.x_tfm(xy[0]), self.y_tfm(xy[1])]
#exporti
import torch.utils.data as th_data
from torch.utils.data import Dataset
# +
#export
# TODO: Use TupleTorchDS to create torch dataloaders
class TupleTorchDS(th_data.Dataset):
def __init__(self, items, x_tfm=None, y_tfm=None):
store_attr()
def __len__(self):
return len(self.items)
def __getitem__(self, index):
x,y = self.items[index]
x = self.x_tfm(x) if self.x_tfm is not None else x
y = self.y_tfm(y) if self.y_tfm is not None else y
return (x,y)
# -
#exporti
from fastcore.transform import Pipeline
# +
#export
i2t_tfm = Int2TensorTransform() # cnvt int -> torch.tensor
cfnp2img_tfm = CifarNP2ImageTransform() # cnvt ndarray -> PIL.Image
cfimg_tfm = CifarImageTransform() # cnvt PIL.Image -> torch.tensor
cfimg2float_tfm = CifarImage2FloatTransform() # cnvt tensor int -> float + div 255
def make_cifar_item_tfm(th_img_tfms=None):
img_tfms = [cfnp2img_tfm]
if th_img_tfms is not None:
# assumes th_img_tfms incl ToTensor (cnvt2 PIL.Image -> tensor + div by 255)
img_tfms += [th_img_tfms]
else:
img_tfms += [cfimg_tfm, cfimg2float_tfm]
return CifarTupleTransform(x_tfm=Pipeline(img_tfms), y_tfm=i2t_tfm)
# -
all_fastai_tfms = make_cifar_item_tfm()
mixed_fastai_train_tfms = make_cifar_item_tfm(th_train_tfms)
#exporti
from fastai.data.core import TfmdDL
from fastai.data.core import TfmdLists
from fastcore.foundation import L
#export
def make_cifar_tls(file_list, path, item_tfm, check=True):
data, targets = load_cifar_items(file_list, path, check=check)
item_tuples = L(data,targets).zip()
tls = TfmdLists(item_tuples,[item_tfm])
return tls
#export
def make_cifar_dl(file_list, path, th_img_tfms=None, check=True, bs=64, **kwargs):
item_tfm = make_cifar_item_tfm(th_img_tfms)
tls = make_cifar_tls(file_list, path, item_tfm, check=check)
dl = TfmdDL(tls,bs=bs, **kwargs)
return dl
train_tls = make_cifar_tls(cifar_dsets.CIFAR10.train_list,
cifar_root,mixed_fastai_train_tfms)
train_tls.tfms[0].x_tfm[1]
train_dl1 = make_cifar_dl(cifar_dsets.CIFAR10.train_list, cifar_root)
#exporti
from fastai.data.core import DataLoaders
#export
def make_fastai_cifar_dls(path, bs=64, check=True, device=None, **kwargs):
train_tfm, test_tfm = make_torch_tfms()
train_dl = make_cifar_dl(
cifar_dsets.CIFAR10.train_list,
path,
train_tfm,
check=check, bs=bs,
shuffle=True)
test_dl = make_cifar_dl(
cifar_dsets.CIFAR10.test_list,
path,
test_tfm,
check=check, bs=bs,
shuffle=False)
dls = DataLoaders(train_dl, test_dl, device=device)
return dls
import torch.cuda
import torch
device = torch.device(torch.cuda.current_device()) if torch.cuda.is_available() else torch.device('cpu')
device
cifar_dls = make_fastai_cifar_dls(cifar_root, device=device)
(cifar_root/'cifar-10-batches-py').ls()
cifar_dsets.CIFAR10.train_list
xb, yb = cifar_dls.one_batch()
xb.dtype
xb.shape
yb.dtype
yb.shape
xb.device
yb.device
cifar_dls.train.after_batch
type(cifar_dls.train.dataset)
hasattr(cifar_dls, 'device')
cifar_dls.device is None
cifar_dls.path
hasattr(cifar_dls.loaders[0],'to')
cifar_dls.loaders[0].device is None
from fastai.vision.learner import cnn_learner
import fastai.callback.progress
from fastai.vision.models import resnet18
from fastai.metrics import accuracy
import torch.nn as nn
learner = cnn_learner(cifar_dls, resnet18, n_out=10, pretrained=False,
normalize=False,
loss_func=nn.CrossEntropyLoss(),metrics=accuracy)
learner.show_training_loop()
learner.summary()
#colab
# %%time
learner.fit(20)
| nbs/02_cifar_loader.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise List 6
#
# Utilizando o Apache Spark e demais ferramentas correlatas, além do grafo construído na lista 5, implemente as seguintes operações:
#
# 1. Determine o nó central através do grau.
# 2. Determine o nó central por centralidade utilizando a distância de Wasserman e a distância harmônica (consultar material).
# 3. Determine o nó central por intermediação.
# **Setup**
# +
from pyspark import SparkConf
from pyspark.context import SparkContext
from pyspark.sql.session import SparkSession
conf = SparkConf().setAppName('appName').setMaster('local')
sc = SparkContext.getOrCreate(conf)
spark = SparkSession(sc)
# -
# **Crie um CSV para armazenar as cidades, com: `id` (nome da cidade), `latitude`, `longitude` e `população`**
cidades = spark.read.format("csv").option("header", "true").load("data/transport/transport-nodes.csv")
cidades.show()
# **Crie outro CSV para armazenar a distância entre essas cidades, com: `src`, `dst` e `relationship` como campos**
distancias = spark.read.format("csv").option("header", "true").load("data/transport/transport-relationships.csv")
distancias.show(30)
# **Utilizando as bibliotecas do Spark, crie um objeto GraphFrame a partir desses dois CSVs.**
# +
from graphframes import *
g = GraphFrame(cidades, distancias)
# -
g.vertices.show()
g.edges.show()
# ### 1. Determine o nó central através do grau.
total_degree = g.degrees
in_degree = g.inDegrees
out_degree = g.outDegrees
(total_degree
.join(in_degree, "id", how="left")
.join(out_degree, "id", how="left")
.fillna(0)
.sort("inDegree", ascending=False)
.show())
# Por grau, os nós centrais são: Ouricuri e Salgueiro.
# ### 2. Determine o nó central por centralidade utilizando a distância de Wasserman e a distância harmônica (consultar material).
# +
import sys
# install a pip package in the current Jupyter kernel
# !{sys.executable} -m pip install networkx
# -
import networkx as nx
import pandas as pd
import json
df = pd.read_csv('data/transport/transport-relationships-networkx.csv')
Graphtype = nx.DiGraph()
gnx = nx.from_pandas_edgelist(df, edge_attr='cost', create_using=Graphtype)
def run_algorithm(runnable, parameter, return_max_only=True):
result = runnable(parameter)
print(json.dumps(result, indent=2))
return result if (return_max_only == False) else max(result, key=result.get)
# **Distância de [Wasserman](https://networkx.github.io/documentation/stable/reference/algorithms/generated/networkx.algorithms.centrality.closeness_centrality.html?highlight=wasserman)**
run_algorithm(nx.closeness_centrality, gnx)
# **Distancia [Harmônica](https://networkx.github.io/documentation/stable/reference/algorithms/generated/networkx.algorithms.centrality.harmonic_centrality.html?highlight=harmonic#networkx.algorithms.centrality.harmonic_centrality)**
run_algorithm(nx.harmonic_centrality, gnx)
# ### 3. Determine o nó central por intermediação ([betweenness](https://networkx.github.io/documentation/stable/reference/algorithms/generated/networkx.algorithms.bipartite.centrality.betweenness_centrality.html?highlight=betweenness#networkx.algorithms.bipartite.centrality.betweenness_centrality))
run_algorithm(nx.betweenness_centrality, gnx)
# **Extra: Determine o nó central por [PageRank](https://graphframes.github.io/graphframes/docs/_site/user-guide.html#pagerank)**
results = g.pageRank(resetProbability=0.15,
maxIter=20)
results.vertices.sort("pagerank", ascending=False).show()
# Nó central utilizando PageRank: Salgueiro.
| pyspark/exercise_list_06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Exploring Ensemble Methods
# # Question 1
# <img src="images/lec5_quiz01_pic01.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 2
# <img src="images/lec5_quiz01_pic02.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 3
# <img src="images/lec5_quiz01_pic03.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 4
# <img src="images/lec5_quiz01_pic04.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 5
# <img src="images/lec5_quiz01_pic05.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 6
# <img src="images/lec5_quiz01_pic06.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 7
# <img src="images/lec5_quiz01_pic07.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 8
# <img src="images/lec5_quiz01_pic08.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 9
# <img src="images/lec5_quiz01_pic09.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
# # Question 10
# <img src="images/lec5_quiz01_pic10.png">
#
# *Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/FgzAt/exploring-ensemble-methods)*
#
# <!--TEASER_END-->
| machine_learning/3_classification/assigment/week5/quiz-week5-assignment1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Understanding Factors in Animal Shelter Pet Adoption - Data Wrangling
#
# In efforts to understand trends in pet adoption outcomes, the Austin Animal Center has provided data relating to the pets in their adoption center. Understanding this data and using it to model the factors that influence pet adoption could lead to recommendations that improve the performance of the center and help more pets find homes.
#
# ### Objective
#
# In this project I will be exploring the dataset and using various data wrangling techniques to prepare the data via basic data wrangling techniques in order to prepare the data for analysis. This will include the following steps:
#
# 1. Loading the data and extracting general info and structure
# 2. Verifying that data is tidy
# 3. Identifying & dealing with missing values/outliers
# ### 1. Data Info and Structure
#
# First I will start by loading the required packages, as well as the dataset which can be found **[here](https://data.austintexas.gov/Health-and-Community-Services/Austin-Animal-Center-Outcomes/9t4d-g238)**
#
# **Note:** This dataset is updated hourly, and was accessed on Sunday, December 12th 2017 at 19:00 UTC for this project.
# +
# For working with dataframes and manipulation
import numpy as np
import pandas as pd
# Used to create graphics and plots
import matplotlib.pyplot as plt
# -
# Instead of the raw data, I will be working with the data that was cleaned in "Data Wrangling - Pet Adoption"
# Load the dataset locally
data = pd.read_csv('data/Austin_Animal_Center_Outcomes.csv')
# Display number of entries and features in the data
print('# of entries in this dataset: {0}'.format(data.shape[0]))
print('# of features per entry: {0}'.format(data.shape[1]))
# Display general information on dataset
data.info()
# There are a few details to mention here. Firstly we can see that within our 12 columns, there seem to be some missing entries in several of the columns, which may need to be addressed later on. In addition, all of the row types are classified as 'object', which can most likely be handled more efficiently if we are able to parse out specific types such as the 'DateTime' column. This column can be handled with much more functionality if we are able to convert it to a Datetime object in our dataframe.
#
# In order to get more information, we will preview the first few rows of the data.
# Display first 10 entries
data.head(10)
# This table gives a much better look at what is going on in the data. Starting from the leftmost column are the following observations:
#
# 1. **Animal ID** - This is a unique identifier for each entry that is a letter combined with a number. This seems well-formatted.
#
# 2. **Name** - Some entries are missing here, and there are also some entries with asterisks before the names (e.g. \*Pebbles, \*Johnny). It will be useful if we can find out the meaning of the asterisk in this field.
#
# 3. **DateTime** and **MonthYear** - These columns look like datetime objects, but they look identical for the entries we see. If we verify that the columns are identical, we may be better served removing one.
#
# 4. **Date of Birth** - This year may also be converted into a datetime object, so that we can perform time-series analysis with this information.
#
# 5. **Outcome Type** - There are several categories in this column, and we may be able to convert the entries into categories for easier handling.
#
# 6. **Outcome Subtype** - This has many missing entries, and we only see categories for Euthanasia and Transfer corresponding outcome types. Depending on the number of subtypes for these types, it may be more efficient to integrate them into the outcome type category.
#
# 7. **Animal Type** - In addition to cats and dogs, there is an 'Other' category here in the third entry, corresponding to a 'Raccoon Mix' breed of animal. With the small amount of types of animals, this column is likely to perform better as categorical values.
#
# 8. **Age upon Outcome** - If we are able to convert this into a uniform value (e.g. age in months) we can work with these values as numbers which will make analyzing this data easier. It should also be noted that if we work with the Outcome DateTime and Date of Birth columns as datetime objects, columns like this can be generated by arithmetic operations.
#
# 9. **Breed** and **Color** - These columns look well-formatted, but more investigation is needed in order to determine whether they will perform better as categorical values, or what additional ways they can be transformed in order to yield more information.
#
# ### 2. Data Cleaning
#
# Before we start cleaning the data, we will look at the distribution of Animal Types that the Austin Animal Center works with.
# Display number of occurences of animal types
data['Animal Type'].value_counts().plot(kind='pie', legend='Best', labels=None)
plt.axis('equal')
plt.show()
# For this project we will only be considering cats and dogs, since they make up the majority of the activity at the animal center and thus require the most resources to shelter and find homes.
cat_data = data['Animal Type'] == 'Cat'
dog_data = data['Animal Type'] == 'Dog'
data = data[cat_data | dog_data]
# Next let's look at the 'Name' column of the data. There are entries with asterisks and entries without. I have contacted the custodians of this dataset, and recieved the following reply:
#
# "...ASO Staff are supposed to be using the asterisk when they name a pet at their review – after they come in.
# If they came in with a name, then no asterisk."
#
# Since the asterisks only denotes whether or not the name of the animal was given before they arrived at the shelter it is not a factor of interest for this analysis so they will simply be removed
# +
# Remove asterisks from names
no_asterisks = []
for name in data['Name']:
if type(name) == str:
no_asterisks.append(name.lstrip('*'))
else:
no_asterisks.append(None)
# Replace list of names without asterisks
data['Name'] = no_asterisks
# -
# Now the names will appear without asterisks.
#
# Next up is the DateTime and MonthYear columns:
# Check if all values in the DateTime and MonthYear columns are identical
(data.DateTime == data.MonthYear).value_counts()
# Since these columns are identical, we can remove the MonthYear column for now.
# Remove MonthYear columnn from the dataset
data = data.drop('MonthYear', axis=1);
# Now we can convert the 'DateTime' column into a datetime format. Since 'Date of Birth' needs this as well, we will perform this action on both columns.
# Convert DateTime and Date of Birth into datetime format
data['DateTime'] = pd.to_datetime(data['DateTime'], format='%m/%d/%Y %I:%M:%S %p')
data['Date of Birth'] = pd.to_datetime(data['Date of Birth'], format='%m/%d/%Y')
# Next let's look at the outcome types and subtypes.
# +
# Display number of occurences of outcome types
print('Outcome Types\n', data['Outcome Type'].value_counts())
# Display number of occurences of outcome subtypes
print('\nOutcome Subtypes\n', data['Outcome Subtype'].value_counts())
# -
# For the outcome types we will mainly be investigating Adoption, Transfers and we will consider all other outcomes as undesirable. Euthanasia and Death must be accounted for so they will remain, but we will drop the small number of entries for animals that either went missing or were brought to the center for disposal.
# Remove 'Missing' and 'Disposal Entries
data = data[data['Outcome Type'] != 'Missing']
data = data[data['Outcome Type'] != 'Disposal']
# Additionally, we will combine the rto-adopt entries into the class of adoption:
# Replace 'Rto-Adopt' entries with 'Adoption' class
data['Outcome Type'].replace('Rto-Adopt', 'Adoption');
# This will help to significantly reduce the number of classes we are working with and thus improve the simplicity of the model.
#
# Before moving on, we should convert our 'Outcome Type' column into categorical values, since we have a managable number of classes and it is our main feature of interest.
# Convert column to categorical entries
data['Outcome Type'] = pd.Categorical(data['Outcome Type'], ordered=False)
# Similarly, the 'Animal Type' and 'Sex upon Outcome' columns should be considered as categorical features
# +
# Display number of occurences of animal types
print('Animal Types\n', data['Animal Type'].value_counts())
# Display number of occurences of sex types
print('\nSex upon Outcome\n', data['Sex upon Outcome'].value_counts())
# -
# Convert columns to categorical entries
data['Animal Type'] = pd.Categorical(data['Animal Type'], ordered=False)
data['Sex upon Outcome'] = pd.Categorical(data['Sex upon Outcome'], ordered=False)
# As previously discussed, we can calculate the outcome age of any animal in the table (with a bit more granularity than we were provided) by converting this column into a timedelta format.
# Replace age column with calculation from birthdate and outcome date
data['Age upon Outcome'] = pd.to_timedelta(data['DateTime'] - data['Date of Birth'])
# Next we consider the breed and color columns:
# +
# Display number of occurences of breeds
print('Breeds\n', data['Breed'].value_counts())
# Display number of occurences of animal colors
print('\nColors\n', data['Color'].value_counts())
# -
# We can see that the list of breeds and colors are very long and many categories only have one entry. There are a few options to take here. First, for breeds, we can cut down the number of entries by reducing the number of mixed breed entries of the format 'breed/breed'.
# +
# Reset the index of the dataframe
data.reset_index(drop=True, inplace=True);
# Print the number of distinct breeds found in the data
print('# of unique breeds in the original dataset: {0}'.format(len(data['Breed'].unique())))
# Print the number of mixed breeds with the format 'breed/breed'
print('# of mixed breeds classified by "breed/breed": {0}'.format(len(data[data['Breed'].str.contains('/').fillna(False)]['Breed'].unique())))
# Iterate over the list to find and replace all mixed breeds with 'first_breed/second_breed' to format 'first_breed Mix'
mixed_breeds = []
for breed in data['Breed']:
if '/' in str(breed):
mixed_breeds.append(breed.split('/')[0] + ' Mix')
else:
mixed_breeds.append(breed)
# Check to see whether the result is of appropriate length
assert len(mixed_breeds) == len(data['Breed'])
# Replace 'Breed' data with reduced category set
data['Breed'] = mixed_breeds
# Display the number of distinct breeds after replacement
print('# of unique breeds after replacement: {0}'.format(len(data['Breed'].unique())))
# -
# Now we have significantly reduced the number of breed categories in the dataset. For the 'Color' categories, we may lose some important information that can be useful. For example, do white and orange cats get adopted more often than white and black cats? In order to keep as much information as possible, but provide some simpler avenues for analysis I will split the 'Color' column into 'Primary Color' and 'Secondary Color' values for the animals.
# +
# Print the number of distinct colors found in the dataset
print('# of unique colors in the original dataset: {0}'.format(len(data['Color'].unique())))
# Print the number of mixed breeds with the format 'color/color'
print('# of mixed colors classified by "color/color": {0}'.format(len(data[data['Color'].str.contains('/')]['Color'].unique())))
# Iterate over the list to find and replace all mixed breeds with 'first_breed/second_breed' to format 'first_breed Mix'
primary_colors = []
secondary_colors = []
for color in data['Color']:
if '/' in color:
primary_colors.append(color.split('/')[0])
secondary_colors.append(color.split('/')[1])
else:
primary_colors.append(color)
secondary_colors.append(None)
# Check to see whether the result is of appropriate length
assert len(primary_colors) == len(data['Color'])
assert len(secondary_colors) == len(data['Color'])
# Replace 'Color' with 'Primary Color' and 'Secondary Color' data
data = data.drop('Color', axis=1)
data['Primary Color'] = pd.Series(primary_colors)
data['Secondary Color'] = pd.Series(secondary_colors)
# Display the number of distinct colors after replacement
print('# of unique colors after replacement: {0}'.format(len(pd.Series(primary_colors).unique())))
print('# of unique colors after replacement: {0}'.format(len(pd.Series(secondary_colors).unique())))
# -
# Now the number of color categories has been reduced to about 10% of its original number. Now we can re-check the values for these categories.
# +
# Display number of occurences of breeds
print('Breeds\n', data['Breed'].value_counts())
# Display number of occurences of animal colors
print('\nPrimary Colors\n', data['Primary Color'].value_counts())
# -
# And finally we set these columns as categorical variables.
# Convert columns to categorical entries
data['Breed'] = pd.Categorical(data['Breed'], ordered=False)
data['Primary Color'] = pd.Categorical(data['Primary Color'], ordered=False)
data['Secondary Color'] = pd.Categorical(data['Secondary Color'], ordered=False)
# Now that we have applied some formatting to each column, we can revisit the dataset information to get a summary of our results.
# +
# Display number of entries and features in the formatted dataset
print('# of entries in this dataset: {0}'.format(data.shape[0]))
print('# of features per entry: {0}\n'.format(data.shape[1]))
# Display formatted dataset information
data.info()
# -
# Not only is the data better formatted for analysis. After replacements, we ended up with the same number of features as before, though these features have a richer set of data for analysis.
# ### 3. Missing Values
#
# Below are the columns with missing data:
# +
# Identify columns with any missing values
NA_columns = data.columns[data.isnull().any()]
# Count missing entries in each column
NA_data_counts = data.isnull().sum()[NA_columns]
# Display columns and counts of missing entries
pd.DataFrame(data = NA_data_counts, columns = ['# of missing entries'])
# -
# We can see that out of the 5 columns that contain missing entries, 'Outcome Subtype' and 'Secondary Color' have empty entries by necessity, since some outcome types do not have subclasses, and some animals do not have secondary colors. In addition, names may not be crucial to the other animal attributes, but it is crucial to keep records of animals with and without given names, since this fact may also influence the outcome type for that animal.
#
# What is left are a handful of entries without outcome type, which is critical information that we are interested in, and without sex information. Since we are currently working with a dataset of more than 70,000 entries, we can remove these entries without any significant impact on our data.
# +
# Remove entries from columns with missing data for either Outcome Type or Sex
data = data.dropna(subset=['Outcome Type', 'Sex upon Outcome'])
# Display columns and counts of missing entries
NA_columns = data.columns[data.isnull().any()]
NA_data_counts = data.isnull().sum()[NA_columns]
pd.DataFrame(data = NA_data_counts, columns = ['# of missing entries'])
# -
# ### 4. Outliers
# Although many outliers were corrected with the previous formatting, there is one main column of interest here that I will explore: 'Outcome Subtype'.
# Finally, we can also classify the marginal subtypes as 'Other' in order to reduce the number of subtypes we are dealing with:
# Display number of occurences of outcome subtypes
print('Outcome Subtypes\n', data['Outcome Subtype'].value_counts())
# +
# Reset the index of the dataframe
data.reset_index(drop=True, inplace=True);
# Create list with subtypes that have less than 100 occurrences each
others = data['Outcome Subtype'].value_counts() < 100
others = others[others == True].index.astype(str)
# Convert Series back into string type temporarily for replacement
data['Outcome Subtype'] = data['Outcome Subtype'].astype(str)
# Initialize empty list and i for loop
i = 0
others_list = []
# replace categories with less than 100 occurences each with 'Other' subtype
for entry in data['Outcome Subtype'].isin(others):
if entry == True:
others_list.append('Other')
elif data.iloc[i, 5] == 'nan':
others_list.append(None)
else:
others_list.append(data.iloc[i, 5])
i += 1
others_list = pd.Categorical(others_list, ordered = False)
# Merge replacements
data['Outcome Subtype'] = others_list
# Display occurrences of outcome subtypes after replacement
print(data['Outcome Subtype'].value_counts())
# -
# Convert columns to categorical entries
data['Outcome Subtype'] = pd.Categorical(data['Outcome Subtype'], ordered=False)
data.info()
# Before wrapping up, the only step remaining is to store the newly formatted dataframe information for uses in analysis.
# Store formatted data as pickle file
data.to_pickle('data/data_clean.pkl')
# ## Closing Remarks
#
# In this project, the Austin Animal Center dataset was explored and prepared for analysis. By converting columns into their appropriate formatting, removing duplicate information, and correcting missing values/outliers where possible, the data can yield more information in analysis. This includes the dataset taking less space in memory to work more quickly, as well as having a richer set of data which we can probe.
#
# ### Thanks for Reading!
| Capstone Project 1/Data Wrangling - Pet Adoption V2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DC Criminalistics: Transit Recommender
# ## Imports
import joblib
import pickle
import pandas as pd
import numpy as np
import datetime
import requests
import censusgeocode as cg
import ipywidgets as widgets
from ipywidgets import HBox, VBox
pd.options.mode.chained_assignment = None
# ## Import Transit data
df_bus = pd.read_csv('../data/wmata-data/bus_station_data_full.csv')
df_rail = pd.read_csv('../data/wmata-data/rail_station_data_full.csv')
df_cabi = pd.read_csv('../data/cabi-station-data/cabi_station_data_full.csv')
# +
df_bus['census_index'] = df_bus['census_index'].apply(str)
df_bus['census_index'] = df_bus['census_index'].apply(lambda x: x.zfill(7))
df_rail['census_index'] = df_rail['census_index'].apply(str)
df_rail['census_index'] = df_rail['census_index'].apply(lambda x: x.zfill(7))
df_cabi['census_index'] = df_cabi['census_index'].apply(str)
df_cabi['census_index'] = df_cabi['census_index'].apply(lambda x: x.zfill(7))
# -
# ## Load model, encoder, and scaler
# +
filename = '../model/BaggingClassifier.sav'
loaded_model = joblib.load(filename)
bg_encoder_file = open('../model/bg_cat_encoder.sav','rb')
bg_encoder = pickle.load(bg_encoder_file)
bg_encoder_file.close()
scaler_file = open('../model/scaler_final.sav','rb')
scaler = pickle.load(scaler_file)
scaler_file.close()
# -
# ## Defining Functions
# #### Geocode address and retrieve census tract and block
def addressLookup(address_input):
address = cg.onelineaddress(address_input + ", Washington, DC")
addressLookup.block_group = address[0]['geographies']['2010 Census Blocks'][0]['BLKGRP']
addressLookup.tract = address[0]['geographies']['2010 Census Blocks'][0]['TRACT']
addressLookup.lat = address[0]['coordinates']['y']
addressLookup.lon = address[0]['coordinates']['x']
# #### Retrieve weather data
def darkSkyAPICall(lat,lon,date_time):
base_url = 'https://api.darksky.net/forecast/'
api_key = 'c9274e7c52c1a5b7e99be6f22db98855'
exclude = 'minutely, hourly, daily, flags'
params = {'exclude': exclude}
lat_address = lat
lon_address = lon
query = "/{},{},{}".format(lat_address,lon_address,date_time)
url = base_url + api_key + query
try:
response = requests.get(url, params=params)
except ConnectionError:
pass
try:
response_json = response.json()
except:
response_json = {}
darkSkyAPICall.summary = response_json['currently']['summary']
#darkSkyAPICall.precip_intensity = response_json['currently']['precipIntensity']
#darkSkyAPICall.precip_probability = response_json['currently']['precipProbability']
darkSkyAPICall.temp = response_json['currently']['temperature']
darkSkyAPICall.dewpoint = response_json['currently']['dewPoint']
darkSkyAPICall.humidity = response_json['currently']['humidity']
#darkSkyAPICall.pressure = response_json['currently']['pressure']
#darkSkyAPICall.wind_speed = response_json['currently']['windSpeed']
#darkSkyAPICall.wind_gust = response_json['currently']['windGust']
#darkSkyAPICall.wind_bearing = response_json['currently']['windBearing']
#darkSkyAPICall.cloud_cover = response_json['currently']['cloudCover']
darkSkyAPICall.uv_index = response_json['currently']['uvIndex']
#darkSkyAPICall.visibility = response_json['currently']['visibility']
weather = "Summary: {} / Temp: {} ".format(darkSkyAPICall.summary, darkSkyAPICall.temp)
#include print of weather to make sure all tests are working
print(weather)
# #### Transforming user input date and time data
def cleanDateTimeInput(date,time,am_pm):
time_transformed = time + ':00' + am_pm
if time_transformed[-2:] == "AM" and time_transformed[:2] == "12":
cleanDateTimeInput.time24 = "00" + time_transformed[2:-2]
elif time_transformed[-2:] == "AM":
cleanDateTimeInput.time24 = time_transformed[:-2]
elif time_transformed[-2:] == "PM" and time_transformed[:2] == "12":
cleanDateTimeInput.time24 = time_transformed[:-2]
else:
cleanDateTimeInput.time24 = str(int(time_transformed[:2]) + 12) + time_transformed[2:8]
cleanDateTimeInput.time24_clean = datetime.datetime.strptime(cleanDateTimeInput.time24, '%H:%M:%S').time()
cleanDateTimeInput.datetime_combined = datetime.datetime.combine(date, cleanDateTimeInput.time24_clean)
cleanDateTimeInput.date_time_clean = cleanDateTimeInput.datetime_combined.isoformat()
return cleanDateTimeInput.date_time_clean
return cleanDateTimeInput.time24
# #### Categorize time of day into appropriate bucket
def timeOfDayBucket(mydatetime):
if 23 <= mydatetime.hour:
timeOfDayBucket.tod_num = 8
if 0 <= mydatetime.hour < 2:
timeOfDayBucket.tod_num = 8
elif 2 <= mydatetime.hour < 5:
timeOfDayBucket.tod_num = 1
elif 5 <= mydatetime.hour < 8:
timeOfDayBucket.tod_num = 2
elif 8 <= mydatetime.hour < 11:
timeOfDayBucket.tod_num = 3
elif 11 <= mydatetime.hour < 14:
timeOfDayBucket.tod_num = 4
elif 14 <= mydatetime.hour < 17:
timeOfDayBucket.tod_num = 5
elif 17 <= mydatetime.hour < 20:
timeOfDayBucket.tod_num = 6
elif 20 <= mydatetime.hour < 23:
timeOfDayBucket.tod_num = 7
# #### Function to return transit recommendation
def transitOrLyft(predicted):
if predicted == 'High' or predicted == 'Med-High':
print('Take a cab!')
else:
bus_options = pd.merge(df['BlockGroup'], df_bus, how='left',
left_on='BlockGroup', right_on='census_index')
rail_options = pd.merge(df['BlockGroup'], df_rail, how='left',
left_on='BlockGroup', right_on='census_index')
capitol_bike_share_options = pd.merge(df['BlockGroup'], df_cabi, how='left',
left_on='BlockGroup', right_on='census_index')
print('Bus Options:')
for index, row in bus_options.iterrows():
if(pd.isnull(row['Stop_Name'])):
print('No nearby bus options')
else:
print(row['Stop_Name'], row['Routes_Available'])
print(' ')
print('Metro Rail Options:')
for index, row in rail_options.iterrows():
if(pd.isnull(row['Description'])):
print('No nearby Metro Rail options')
else:
print(row['Description'], row['Station_Entrance'])
print(' ')
print('Capitol Bike Share Options:')
for index, row in capitol_bike_share_options.iterrows():
if(pd.isnull(row['name'])):
print('No nearby Capitol Bike Share Options')
else:
print(row['name'])
# ## Setting up widgets
# #### Address input widget
# +
address = widgets.Text(
value='Ex: 640 Massachusetts Ave NW',
placeholder='',
description='Address: ',
disabled=False
)
date = widgets.DatePicker(
description='Pick a Date',
disabled=False
)
time = widgets.Dropdown(
options=['12:00', '12:30', '01:00', '01:30', '02:00', '02:30', '03:00',
'03:30', '04:00', '04:30','05:00', '05:30', '06:00', '06:30', '07:00', '07:30',
'08:00', '08:30', '09:00', '09:30', '10:00', '10:30', '11:00', '11:30'],
value='09:00',
description='Depart Time:',
disabled=False,
)
am_pm = widgets.Dropdown(
options=['AM','PM'],
value='AM',
description='AM or PM? ',
disabled=False,
)
# -
# # User Inputs
address
date
HBox([time, am_pm])
# ## Clean
# ## Return travel recommendation
# +
cleanDateTimeInput(date.value,time.value,am_pm.value)
addressLookup(address.value)
darkSkyAPICall(addressLookup.lat,addressLookup.lon,cleanDateTimeInput.date_time_clean)
timeOfDayBucket(cleanDateTimeInput.time24_clean)
weekday = cleanDateTimeInput.datetime_combined.weekday()
day = cleanDateTimeInput.datetime_combined.day
month = cleanDateTimeInput.datetime_combined.month
# +
tract_input = addressLookup.tract.rjust(6, '0')
block_group_input = addressLookup.block_group
block_group = addressLookup.tract + addressLookup.block_group
bg_cat = tract_input + ' ' + block_group_input
list_values = [bg_cat, weekday, timeOfDayBucket.tod_num, darkSkyAPICall.uv_index,
darkSkyAPICall.temp, day, block_group]
list_columns = ['bg_cat', 'weekday', 'tod_num', 'uv_index',
'temperature', 'day', 'BlockGroup']
df = pd.DataFrame([list_values],columns=list_columns)
df_modeling = df[['bg_cat', 'weekday', 'tod_num', 'uv_index',
'temperature', 'day']]
df_modeling['bg_cat'] = bg_encoder.transform(df_modeling['bg_cat'])
scaled_df = scaler.transform(df_modeling)
predicted = loaded_model.predict(scaled_df)
transitOrLyft(predicted)
| demo/Capstone Demo Notebook_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gizemzeynep/BBY162-2020/blob/master/calisma_defteri.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="6IOyCRzqRgBu" colab_type="text"
# Bu çalışma defteri Stepnik eğitim platformunda yer alan Jetbrains'in "Introduction to Python" dersi temel alınarak hazırlanmıştır. (https://stepik.org/course/238)
#
# + [markdown] id="AE4EYdj6VPvq" colab_type="text"
# # Bölüm 01: Giriş
# + [markdown] id="4nLn-agoUdfh" colab_type="text"
# ## İlk Bilgisayar Programımız
# Geleneksel olarak herhangi bir programlama dilinde yazdığınız ilk program "Merhaba Dünya!"'dır.
# + [markdown] id="xhY6ovVFSBp1" colab_type="text"
# **Örnek Uygulama:**
# ```
# print("Merhaba Dünya!")
# ```
# + [markdown] id="MuYQpCfoh7In" colab_type="text"
# **Görev:** Kendinizi dünyaya tanıtacak ilk bilgisayar programınızı yazın!
# + id="CBHkVbIlUXgW" colab_type="code" cellView="both" outputId="58a91757-4a09-410f-d43f-d972c97201d1" colab={"base_uri": "https://localhost:8080/", "height": 34}
print("Merhaba dünyacığım")
# + [markdown] id="0S8fTkAqVb4v" colab_type="text"
# ## Yorumlar
# Python'daki yorumlar # "hash" karakteriyle başlar ve fiziksel çizginin sonuna kadar uzanır. Yorum yapmak için kullandığımız # "hash" karakteri kod satırlarını geçici olarak devre dışı bırakmak amacıyla da kullanılabilir.
# + [markdown] colab_type="text" id="ql-GYfPYWpUW"
# **Örnek Uygulama:**
# ```
# # Bu ilk bilgisayar programım için ilk yorumum
# print("# bu bir yorum değildir")
# print("Merhaba!") # yorumlar kod satırının devamında da yapılabilir.
# #print("Bu kod geçici olarak devre dışı bırakılmıştır.")
# ```
#
# + [markdown] id="4ufK03hEiP76" colab_type="text"
# **Görev:** Bilgisayar kodunuza yeni yorum ekleyin, kodları geçici olarak devre dışı bırakın!
# + id="QzUppEjoXLPM" colab_type="code" outputId="ad4cddb7-bcff-407a-d27d-1f2069a86601" colab={"base_uri": "https://localhost:8080/", "height": 34}
print("#Merhaba! artık devre dışıyım")
# + [markdown] id="9JeaGlvQYylS" colab_type="text"
# # Bölüm 02: Değişkenler
# + [markdown] id="VzeGixS6Y919" colab_type="text"
# ## Değişken Nedir?
# Değişkenler değerleri depolamak için kullanılır, böylece daha sonra bu değişkenleri program içinde çağırarak kullanabilir. Değişkenler etiketlere benzer ve atama operatörü olarak adlandırılan eşittir ( = ) operatörü ile bir değişkene bir değer atanabilir. Bir değer ataması zincirleme şeklinde gerçekleştirilebilir. Örneğin: a = b = 2
# + [markdown] id="7tWAdnNlZtQr" colab_type="text"
# **Örnek Uygulama:**
# ```
# a = b = 2 # Bu bir "zincirleme atamadır". Değer olarak atanan 2 hem "a" değişkenine, hem de "b" değişkenine atanmaktadır.
# print("a = " + str(a))
# print("b = " + str(b))
# # str(a) ve str(b) ifadesi eğitimin ilerleyen bölümlerinde anlatılacaktır.
# # Şu an için "a" ve "b" değişkenini metin formatına çevirmek için kullanılmakta olduğunu bilmeniz yeterlidir.
#
# adSoyad = "<NAME>"
# print("Adı Soyadı: " + adSoyad)
# ```
# + [markdown] id="szRkWMg9idf9" colab_type="text"
# **Görev:** "eposta" adlı bir değişken oluşturun. Oluşturduğunuz bu değişkene bir e-posta adresi atayın. Daha sonra atadığınız bu değeri, yazdırın. Örneğin: "E-posta: orcun[at]madran.net"
# + id="RmIa5BJsZ2EI" colab_type="code" outputId="ca89f409-c2ba-4f64-e583-a8242aaf263a" colab={"base_uri": "https://localhost:8080/", "height": 34}
eposta = "zynptzl0[at]gmail.com"
print("E-posta:" + " " + eposta)
# + [markdown] colab_type="text" id="2irKEi_raxaB"
# ## Değişken Tanımlama
# Değişken adları yalnızca harf, rakam ve / veya alt çizgi karakteri içerebilir ve bir rakamla başlayamaz. Python'ın 3. sürümünden itibaren değişken adlarında latin dışı karakterler (Ör: İ, ı, ç, ö, ş, ğ vb.) kullanılabilir. Ancak bu programlama dilleri genelinde tavsiye edilmez.
# + [markdown] id="pOxCruNHdp9M" colab_type="text"
# **Örnek Uygulama:**
# ```
# degisken = 1
# ```
# + [markdown] id="SVwQ0Tf-isfz" colab_type="text"
# **Görev:** Henüz tanımlanmamış bir değişken kullanırsanız ne olacağını kontrol edin!
# + id="1W14OfkxeGuN" colab_type="code" outputId="0ae52acc-ed42-4408-ff80-a1670b7676fb" colab={"base_uri": "https://localhost:8080/", "height": 180}
degisken1 = "Veri"
print(degisken2)
# + [markdown] id="GwLgmn1fi04Q" colab_type="text"
# **Görev:** Tanımladığınız değişkeni ekrana yazdırın!
# + id="JrNUlQAyeqHQ" colab_type="code" outputId="38de25df-8ab2-44e8-d3b1-3b1594042efd" colab={"base_uri": "https://localhost:8080/", "height": 34}
degisken1 = "Veri"
print(degisken1)
# + [markdown] id="_6eRj3kmeyGw" colab_type="text"
# ## Değişken Türleri
# Python'da iki ana sayı türü vardır; tam sayılar ve ondalık sayılar.
#
# ---
#
#
# **Dikkat:** Ondalık sayıların yazımında Türkçe'de *virgül* (,) kullanılmasına rağmen, programlama dillerinin evrensel yazım kuralları içerisinde ondalık sayılar *nokta* (.) ile ifade edilir.
# + [markdown] id="37MxozrGf1-F" colab_type="text"
# **Örnek Uygulama:**
# ```
# tamSayi = 5
# print(type(tamSayi)) # tamSayi değişkeninin türünü yazdırır
#
# ondalikSayi = 7.4
# print(type(ondalikSayi) # ondalikSayi değişkeninin türünü yazdırır
# ```
# + [markdown] id="asiOZVmbn_TO" colab_type="text"
# **Görev:** "sayi" değişkeninin türünü belirleyerek ekrana yazdırın!
# + id="UrGfF38CgjdQ" colab_type="code" cellView="both" outputId="96e00d15-644f-405b-d500-f8cb27180d2d" colab={"base_uri": "https://localhost:8080/", "height": 34}
sayi = 9.0
print(type(sayi))
# + [markdown] id="i3Qht_zmjLf2" colab_type="text"
# ## Değişken Türü Dönüştürme
# Bir veri türünü diğerine dönüştürmenize izin veren birkaç yerleşik fonksiyon (built-in function) vardır. Bu fonksiyonlar ("int()", "str()", "float()") uygulandıkları değişkeni dönüştürerek yeni bir nesne döndürürler.
# + [markdown] id="F-Zp8zfmkzJX" colab_type="text"
# **Örnek Uygulama**
# ```
# sayi = 6.5
# print(type(sayi)) # "sayi" değişkeninin türünü ondalık olarak yazdırır
# print(sayi)
#
# sayi = int(sayi) # Ondalık sayı olan "sayi" değişkenini tam sayıya dönüştürür
# print(type(sayi))
# print(sayi)
#
# sayi = float(sayi) # Tam sayı olan "sayi" değişkenini ondalık sayıya dönüştürür
# print(type(sayi))
# print(sayi)
#
# sayi = str(sayi) # "sayi" değişkeni artık düz metin halini almıştır
# print(type(sayi))
# print(sayi)
# ```
# + [markdown] id="7j51gKaIlM-u" colab_type="text"
# **Görev:** Ondalık sayıyı tam sayıya dönüştürün ve ekrana değişken türünü ve değeri yazdırın!
# + id="o9ZQ6Jr3lL6A" colab_type="code" outputId="b0b35b8c-5032-45bd-cf69-14d7edb1420e" colab={"base_uri": "https://localhost:8080/", "height": 51}
sayi = int(3.14)
print(type(sayi))
print(sayi)
# + [markdown] id="f5MK6vZOl6dG" colab_type="text"
# ## Aritmetik Operatörler
# Diğer tüm programlama dillerinde olduğu gibi, toplama (+), çıkarma (-), çarpma (yıldız) ve bölme (/) operatörleri sayılarla kullanılabilir. Ek olarak Python'un üs (çift yıldız) ve mod (%) operatörleri vardır.
#
# ---
# **Not:** Matematik işlemlerinde geçerli olan aritmetik operatörlerin öncelik sıralamaları (çarpma, bölme, toplama, çıkarma) ve parantezlerin önceliği kuralları Python içindeki matematiksel işlemler için de geçerlidir.
#
# + [markdown] id="1THb0XpGmwSD" colab_type="text"
# **Örnek Uygulama:**
# ```
# # Toplama işlemi
# sayi = 7.0
# sonuc = sayi + 3.5
# print(sonuc)
#
# # Çıkarma işlemi
# sayi = 200
# sonuc = sayi - 35
# print(sonuc)
#
# # Çarpma işlemi
# sayi = 44
# sonuc = sayi * 10
# print(sonuc)
#
# # Bölme işlemi
# sayi = 30
# sonuc = sayi / 3
# print(sonuc)
#
# # Üs alma işlemi
# sayi = 30
# sonuc = sayi ** 3
# print(sonuc)
#
# # Mod alma işlemi
# sayi = 35
# sonuc = sayi % 4
# print(sonuc)
# ```
# + [markdown] id="jp3PvDghnJkY" colab_type="text"
# **Görev:** Aşağıda değer atamaları tamamlanmış olan değişkenleri kullanarak ürünlerin peşin satın alınma bedelini TL olarak hesaplayınız ve ürün adı ile birlikte ekrana yazdırınız! İpucu: Ürün adını ve ürün bedelini tek bir satırda yazdırmak isterseniz ürün bedelini str() fonksiyonu ile düz metin değişken türüne çevirmeniz gerekir.
# + id="oMY_MZjmndQj" colab_type="code" outputId="4e8a7888-abd7-4b96-a24d-20fd90659c8f" colab={"base_uri": "https://localhost:8080/", "height": 34}
urunAdi = "Bisiklet"
urunBedeliAvro = 850
pariteAvroTL = 7
urunAdet = 3
pesinAdetIndirimTL = 500
aAvro=850*3
aTL=aAvro*7
print("Almak istediğiniz " +str(urunAdi)+ " " +str(aTL)+ " "+ "TL'dir")
# + [markdown] id="OUWeht_8nyCx" colab_type="text"
# ## Artırılmış Atama Operatörleri
# Artırılmış atama, bir değişkenin mevcut değerine belirlenen değerin eklenerek (+=) ya da çıkartılarak (-=) atanması işlemidir.
#
# + [markdown] id="r-6ImrKCobv_" colab_type="text"
# **Örnek Uygulama**
#
# ```
# sayi = 8
# sayi += 4 # Mevcut değer olan 8'e 4 daha ekler.
# print(sayi)
#
# sayi -= 6 # Mevcut değer olan 12'den 6 eksiltir.
# print("number = " + str(number))
# ```
#
#
# + [markdown] id="Du5w88EaovzQ" colab_type="text"
# **Görev:** "sayi" değişkenine 20 ekleyip, 10 çıkartarak değişkenin güncel değerini ekrana yazdırın!
# + id="ro0-FwgDpCkx" colab_type="code" outputId="aa78df80-02c3-4476-b100-557d38f26030" colab={"base_uri": "https://localhost:8080/", "height": 34}
sayi = 55
sayi +=20
sayi -=10
print(sayi)
# + [markdown] id="C0G673j0pQD4" colab_type="text"
# ## Boolean Operatörleri
# Boolean, yalnızca **Doğru (True)** veya **Yanlış (False)** olabilen bir değer türüdür. Eşitlik (==) operatörleri karşılaştırılan iki değişkenin eşit olup olmadığını kontrol eder ve *True* ya da *False* değeri döndürür.
# + [markdown] id="Trd3P6X4pwbu" colab_type="text"
# **Örnek Uygulama:**
# ```
# deger1 = 10
# deger2 = 10
# esitMi = (deger1 == deger2) # Eşit olup olmadıkları kontrol ediliyor
# print(esitMi) # Değişken "True" olarak dönüyor
#
# deger1 = "Python"
# deger2 = "Piton"
# esitMi = (deger1 == deger2) # Eşit olup olmadıkları kontrol ediliyor
# print(esitMi) # Değişken "False" olarak dönüyor
# ```
# + [markdown] id="QSvUSLN4p7Ra" colab_type="text"
# **Görev:** Atamaları yapılmış olan değişkenler arasındaki eşitliği kontrol edin ve sonucu ekrana yazıdırın!
# + id="ihVP2eQpttpv" colab_type="code" outputId="eb697f97-bd0a-4be0-c21e-c107979c0e91" colab={"base_uri": "https://localhost:8080/", "height": 34}
sifre = "Python2020"
sifreTekrar = "Piton2020"
esitMi = (sifre == sifreTekrar)
print(esitMi)
# + [markdown] id="NU2bhXlwuUtj" colab_type="text"
# ## Karşılaştırma Operatörleri
# Python'da, >=, <= , >, < vb. dahil olmak üzere birçok operatör bulunmaktadır. Python'daki tüm karşılaştırma operatörleri aynı önceliğe sahiptir. Karşılaştırma sonucunda boole değerleri (*True* ya da *False*) döner. Karşılaştırma operatörleri isteğe bağlı olarak arka arkaya da (zincirlenerek) kullanılabilir.
# + [markdown] id="kUkH4Qe6vBIm" colab_type="text"
# **Örnek Uygulama:**
# ```
# deger1 = 5
# deger2 = 7
# deger3 = 9
#
# print(deger1 < deger2 < deger3) # Sonuç "True" olarak dönecektir
# ```
#
#
# + [markdown] id="JrbnTUNavEsJ" colab_type="text"
# **Görev:** Aşağıda değer atamaları tamamlanmış olan değişkenleri kullanarak ürünlerin peşin satın alınma bedelini TL olarak hesaplayın. Toplam satın alma bedeli ile bütçenizi karşılaştırın. Satın alma bedelini ve bütçenizi ekrana yazdırın. Ödeme bütçenizi aşıyorsa ekrana "False", aşmıyorsa "True" yazdırın.
# + id="chsHnUn9vWDA" colab_type="code" colab={}
urunAdi = "Bisiklet"
urunBedeliAvro = 850
kurAvro = 7
urunAdet = 3
pesinAdetIndirimTL = 500
butce = 15000
# + [markdown] id="qhLUJ8wJM79g" colab_type="text"
# # Part 03: Metin Katarları
# + [markdown] id="9UNmx1IUNSrB" colab_type="text"
# ## Birbirine Bağlama
# Birbirine bağlama artı (+) işlemini kullanarak iki metin katarının birleştirilmesi işlemine denir.
# + [markdown] id="6usSXTKYze8y" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="1CkYIUTqNhSR" colab_type="text"
#
#
# ```
# deger1 = "Merhaba"
# deger2 = "Dünya"
#
# selamlama = deger1 + " " + deger2
# print(selamlama) # Çıktı: Merhaba Dünya
# ```
#
#
# + [markdown] id="n9rA4HAHNqI7" colab_type="text"
# **Görev:** *ad*, *soyad* ve *hitap* değişkenlerini tek bir çıktıda birleştirecek kodu yazın!
# + id="xyNs9VEFOHJc" colab_type="code" outputId="d0a6987a-1401-4a6b-da14-3bf5e751f6a8" colab={"base_uri": "https://localhost:8080/", "height": 34}
hitap = "<NAME>."
ad = "Orçun"
soyad = "Madran"
# Çıktı: Öğr. Gör. Or<NAME>
print(hitap + " " + ad+ " " + soyad)
# + [markdown] id="ipcnECBCOKWX" colab_type="text"
# ## Metin Katarı Çarpımı
# Python, metin katarlarının çarpım sayısı kadar tekrar ettirilmesini desteklemektedir.
# + [markdown] id="hKQAzEYOzyIk" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="HXxuq9kMO2xm" colab_type="text"
#
#
# ```
# metin = "Hadi! "
# metniCarp = metin * 4
# print(metniCarp) # Çıktı: Hadi! Hadi! Hadi! Hadi!
# ```
#
#
# + [markdown] id="56NercDUOh8x" colab_type="text"
# **Görev:** Sizi sürekli bekleten arkadaşınızı uyarabilmek için istediğiniz sayıda "Hadi!" kelimesini ekrana yazdırın!
# + id="j_CdqS1bPHAb" colab_type="code" outputId="c3fdfaa3-80f2-4b45-a945-4d4ee113f986" colab={"base_uri": "https://localhost:8080/", "height": 34}
metin = "Hadi! "
# Çıktı: Hadi! Hadi! Hadi! Hadi! ... Hadi!
metnicarp = metin * 9
print(metnicarp)
# + [markdown] id="Xo6sOrv-PpgY" colab_type="text"
# ##Metin Katarı Dizinleme
# Konumunu biliniyorsa, bir metin katarındaki ilgili karaktere erişebilebilir. Örneğin; str[index] metin katarındaki indeks numarasının karşılık geldiği karakteri geri döndürecektir. İndekslerin her zaman 0'dan başladığı unutulmamalıdır. İndeksler, sağdan saymaya başlamak için negatif sayılar da olabilir. -0, 0 ile aynı olduğundan, negatif indeksler -1 ile başlar.
# + [markdown] id="93mxLsQ0z9bB" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="VV71MAQFQaPs" colab_type="text"
#
#
# ```
# metin = "Python Programlama Dili"
# print("h " + metin[3]) # Not: indeksler 0'dan başlar
# ```
#
# + [markdown] id="55Lviae-QTax" colab_type="text"
# **Görev:** İndeks numarasını kullanarak metin katarındaki ikinci "P" harfini ekrana yazdırın!
#
# + id="dHJ48v5WQe4c" colab_type="code" outputId="7da0a634-c567-40f0-9470-e513f3253853" colab={"base_uri": "https://localhost:8080/", "height": 34}
metin = "Python Programlama Dili"
# Çıktı: P
print(metin[7])
# + [markdown] id="op2D6v9xQsRe" colab_type="text"
# ## Metin Katarı Negatif Dizinleme
# Metin katarının sonlarında yer alan bir karaktere daha rahat erişebilmek için indeks numarası negatif bir değer olarak belirlenebilir.
# + [markdown] id="FR64ir-m0Hw2" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="kZTrtqD6RM7j" colab_type="text"
#
# ```
# metin = "Python Programlama Dili"
# dHarfi = metin[-4]
# print(dHarfi)
# ```
#
#
# + [markdown] id="ZM6vbbChQ_4R" colab_type="text"
# **Task:** Metin katarının sonunda yer alan "i" harfini ekrana yazdırın!
# + id="d49Cfw-FRJMT" colab_type="code" outputId="ba777715-8ae5-486d-9120-954514e68fee" colab={"base_uri": "https://localhost:8080/", "height": 34}
metin = "Python Programlama Dili"
#Çıktı: i
iHarfi = metin[-1]
print(iHarfi)
# + [markdown] id="xPgrx41FRbrw" colab_type="text"
# ##Metin Katarı Dilimleme
# Dilimleme, bir metin katarından birden çok karakter (bir alt katar) almak için kullanılır. Söz dizimi indeks numarası ile bir karaktere erişmeye benzer, ancak iki nokta üst üste işaretiyle ayrılmış iki indeks numarası kullanılır. Ör: str[ind1:ind2].
#
# Noktalı virgülün solundaki indeks numarası belirlenmezse ilk karakterden itibaren (ilk karakter dahil) seçimin yapılacağı anlamına gelir. Ör: str[:ind2]
#
# Noktalı virgülün sağındaki indeks numarası belirlenmezse son karaktere kadar (son karakter dahil) seçimin yapılacağı anlamına gelir. Ör: str[ind1:]
# + [markdown] id="cDh07_U90Kan" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="X_MHtUgUSFpY" colab_type="text"
#
#
# ```
# metin = "Python Programlama Dili"
# dilimle = metin[:6]
# print(dilimle) # Çıktı: Python
#
# metin = "Python Programlama Dili"
# print(metin[7:]) # Çıktı: Programlama Dili
#
# ```
#
#
# + [markdown] id="SjPJOtqUR7rT" colab_type="text"
# **Görev:** Metin katarını dilemleyerek katarda yer alan üç kelimeyi de ayrı ayrı (alt alta) ekrana yazdırın!.
# + id="bHxs-WnaSKie" colab_type="code" outputId="d25cc490-7823-46f6-eeb0-443a6af47b95" colab={"base_uri": "https://localhost:8080/", "height": 68}
metin = "Python Programlama Dili"
# Çıktı:
# Python
# Programlama
# Dili
dilimleme = metin[7:18]
print(metin[:6])
print(metin[7:18])
print(metin[19:])
dilimleme2 = metin[:6]
dilimleme3 = metin[19:]
# + [markdown] id="aIfLylzcShEG" colab_type="text"
# ##In Operatörü
# Bir metin katarının belirli bir harf ya da bir alt katar içerip içermediğini kontrol etmek için, in anahtar sözcüğü kullanılır.
# + [markdown] id="oToLWF8F0Mx8" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="pZZxStolS-Rg" colab_type="text"
#
#
# ```
# metin = "Python Programlama Dili"
# print("Programlama" in metin) # Çıktı: True
# ```
#
#
# + [markdown] id="551pDT0YS14M" colab_type="text"
# **Görev:** Metin katarında "Python" kelimesinin geçip geçmediğini kontrol ederek ekrana yazdırın!
# + id="2CpxVwHMTENL" colab_type="code" outputId="f275ba75-f8b6-427d-874d-ff85c4b21152" colab={"base_uri": "https://localhost:8080/", "height": 34}
metin = "Python Programlama Dili"
# Çıktı: True
print("Python"in metin)
# + [markdown] id="mKLQLw8_TJyz" colab_type="text"
# ##Metin Katarının Uzunluğu
# Bir metin katarının kaç karakter içerdiğini saymak için len() yerleşik fonksiyonu kullanılır.
# + [markdown] id="TO1s9wYS0Qhx" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="aa4IkB7CTuyr" colab_type="text"
#
#
# ```
# metin = "Python programlama dili"
# print(len(metin)) # Çıktı: 23
# ```
#
#
# + [markdown] id="BF5DPvyuTa4R" colab_type="text"
# **Görev:** Metin katarındaki cümlenin ilk yarısını ekrana yazdırın!
# + id="Uvnr2f1XT05N" colab_type="code" outputId="dd04e78f-6685-4735-a3af-19055cf0b953" colab={"base_uri": "https://localhost:8080/", "height": 34}
metin = "Python programlama dili, dünyada eğitim amacıyla en çok kullanılan programlama dillerinin başında gelir."
# Çıktı: Python programlama dili, dünyada eğitim amacıyla en
print(metin[:int(len(metin)/2)])
# + [markdown] id="T_u1X0aKVAO2" colab_type="text"
# ## Özel Karakterlerden Kaçma
# Metin katarları içerisinde tek ve çift tırnak kullanımı kimi zaman sorunlara yol açmaktadır. Bu karakterin metin katarları içerisinde kullanılabilmesi için "Ters Eğik Çizgi" ile birlikte kullanılırlar.
#
# Örneğin: 'Önümüzdeki ay "Ankara'da Python Eğitimi" gerçekleştirilecek' cümlesindeki tek tırnak kullanımı soruna yol açacağından 'Önümüzdeki ay "Ankara\'da Python Eğitimi" gerçekleştirilecek' şeklinde kullanılmalıdır.
#
# **Not:** Tek tırnaklı metin katarlarından kaçmak için çift tırnak ya da tam tersi kullanılabilir.
# + [markdown] id="WF9cEZEE0T4q" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="bJX6J5bvVbZa" colab_type="text"
#
#
# ```
# metin = 'Önümüzdeki ay "Ankara'da Python Eğitimi" gerçekleştirilecektir.'
# print(metin) # Çıktı: Geçersiz söz dizimi hatası dönecektir.
#
# metin = 'Önümüzdeki ay "Ankara\'da Python Eğitimi" gerçekleştirilecektir.'
# print(metin) #Çıktı: Önümüzdeki ay "Ankara\'da Python Eğitimi" gerçekleştirilecektir.
# ```
#
#
# + [markdown] id="5520MewzVPmx" colab_type="text"
# **Task:** Metin katarındaki cümlede yer alan noktalama işaretlerinden uygun şekilde kaçarak cümleyi ekrana yazdırın!
# + id="A8Gpec2uVhxU" colab_type="code" outputId="9776a9e3-66b5-4a42-ca2e-3b36e236c7df" colab={"base_uri": "https://localhost:8080/", "height": 34}
metin = 'Bilimsel çalışmalarda "Python" kullanımı Türkiye\'de çok yaygınlaştı!'
print(metin)
# + [markdown] id="cft6m1VaVqVH" colab_type="text"
# ##Basit Metin Katarı Metodları
# Python içinde birçok yerleşik metin katarı fonksiyonu vardır. En çok kullanılan fonksiyonlardan bazıları olarak;
#
# * tüm harfleri büyük harfe dönüştüren upper(),
# * tüm harfleri küçük harfe dönüştüren lower(),
# * sadece cümlenin ilk harfini büyük hale getiren capitalize() sayılabilir.
#
# **Not:** Python'daki yerleşik fonksiyonların bir listesini görüntüleyebilmek için metin katarından sonra bir nokta (.) koyulur ve uygun olan fonksiyonlar arayüz tarafından otomatik olarak listelenir. Bu yardımcı işlevi tetiklemek için CTRL + Bolşuk tuş kombinasyonu da kullanılabilir.
# + [markdown] id="5_V3UGn20YGV" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="S3-V5gloV3MM" colab_type="text"
#
#
# ```
# metin = "Python Programlama Dili"
# print(metin.lower()) # Çıktı: python programlama dili
# print(metin.upper()) # Çıktı: PYTHON PROGRAMLAMA DILI
# print(metin.capitalize()) # Çıktı: Python programlama dili
# ```
#
#
# + [markdown] id="-ko2Gpq_Vz23" colab_type="text"
# **Task:** *anahtarKelime* ve *arananKelime* değişkenlerinde yer alan metinler karşılaştırıldığında birbirlerine eşit (==) olmalarını sağlayın ve dönen değerin "True" olmasını sağlayın!
# + id="XfaN4psBV8wr" colab_type="code" outputId="7f323f62-4dde-4d06-8dd8-33db530e071a" colab={"base_uri": "https://localhost:8080/", "height": 51}
anahtarKelime = "Makine Öğrenmesi"
arananKelime = "makine öğrenmesi"
print(anahtarKelime == arananKelime) # Çıktı: True
print(anahtarKelime.lower() == arananKelime)
# + [markdown] id="16m1oC6LWEdJ" colab_type="text"
# ##Metin Katarı Biçimlendirme
# Bir metin katarından sonraki % operatörü, bir metin katarını değişkenlerle birleştirmek için kullanılır. % operatörü, bir metin katarıdanki % s öğesini, arkasından gelen değişkenle değiştirir. % d özel sembolü ise, sayısal veya ondalık değerler için yer tutucu olarak kullanılır.
# + [markdown] id="Qr673Q4m0czk" colab_type="text"
# **Örnek Uygulama:**
# + [markdown] id="1l70j4-3WP4b" colab_type="text"
#
#
# ```
# adsoyad = "<NAME>"
# dogumTarihi = 1976
#
# print("Merhaba, ben %s!" % adsoyad) # Çıktı: Merhaba, ben <NAME>!
# print("Ben %d doğumluyum" % dogumTarihi) # Ben 1976 doğumluyum.
# ```
#
#
# + [markdown] id="b7ny_Q7uWJJX" colab_type="text"
# **Görev:** <NAME>, bu dönemki dersiniz "Programlama Dilleri". Başarılar! cümlesini ekrana biçimlendirmeyi kullanarak (artı işaretini kullanmadan) yazdırın!
# + id="HxYY3XqGWYsi" colab_type="code" outputId="95a8d865-b909-4e9d-88ea-b84136f28f89" colab={"base_uri": "https://localhost:8080/", "height": 34}
ad = "Orçun"
soyad = "Madran"
ders = "Programlama Dilleri"
# Çıktı: <NAME> Madran, bu dönemki dersiniz "Programlama Dilleri". Başarılar!
print("Merhaba, ben %s " % ad)
# + [markdown] id="nTiryVt-EuJP" colab_type="text"
# # Bölüm 04: Veri Yapıları
# + [markdown] id="H78J_wNDE1Id" colab_type="text"
# ## Listeler
# Liste, farklı bilgi parçalarının bir koleksiyonunu tek bir değişken adı altında saklamak için kullanabileceğiniz bir veri yapısıdır. Bir liste köşeli parantez arasında virgülle ayrılmış değerler dizisi olarak yazılabilir. Ör: liste = [deger1, deger2]. Listeler farklı türden öğeler içerebilir, ancak genellikle listedeki tüm öğeler aynı türdedir. Metin katarları gibi listeler de dizine eklenebilir ve dilimlenebilir (bkz. Bölüm 3).
# + [markdown] id="T_nubI5kGCs3" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="O4TC5Qs2Ge8H" colab_type="text"
#
#
# ```
# acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"] # acikListe adında yeni bir liste oluşturur
#
# print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak']
# ```
#
#
# + [markdown] colab_type="text" id="ZqYRBLZSIJvi"
# **Görev 1:** acikListe içinde yer alan 3. liste öğesini ekrana yazıdırın!
# + id="bVSMLdnLG7Kx" colab_type="code" outputId="2c4c0685-f4b1-4ed8-bcac-bb7e308ef667" colab={"base_uri": "https://localhost:8080/", "height": 34}
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"]
print(acikListe[2])
# + [markdown] colab_type="text" id="pZ4hTBC0JCln"
# **Görev 2:** acikListe içinde yer alan 4. ve 5. liste öğesini ekrana yazıdırın!
# + colab_type="code" id="_DgLPxODI3aP" outputId="6b10d78a-6ac9-4158-befa-46936778c63c" colab={"base_uri": "https://localhost:8080/", "height": 34}
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"]
print(acikListe[3:])
# + [markdown] id="D_Vt3_mQJq9Q" colab_type="text"
# ## Liste İşlemleri
# Append () yöntemini ya da birleştirmeyi += kullanarak listenin sonuna yeni öğeler (değerler) eklenebilir. Metin katarlarından farklı olarak, listeler değiştirilebilir bir türdür, yani liste[indeksNo] = yeni_deger kullanarak içeriklerini değiştirmek mümkündür.
# + [markdown] id="m7Y-NBAALfyk" colab_type="text"
# **Örnek Uygulama**
# + [markdown] colab_type="text" id="3xGEUhipL11G"
#
#
# ```
# acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"] # acikListe adında yeni bir liste oluşturur
# print(acikliste) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak']
#
# acikListe += ["Açık Donanım", "Açık İnovasyon"] # listeye iki yeni öğe ekler
# print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak', 'Açık Donanım', 'Açık İnovasyon']
#
# acikListe.append("Açık Veri Gazeteciliği") # listeye yeni bir öğe ekler
# print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak', 'Açık Donanım', 'Açık İnovasyon', 'Açık Veri Gazeteciliği']
#
# acikListe[4] = "Açık Kaynak Kod" # listenin 5. öğesini değiştirir
# print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', '<NAME>', '<NAME>', 'Açık İnovasyon', 'Açık Veri Gazeteciliği']
# ```
#
#
# + [markdown] id="5hgUnMj7ODY1" colab_type="text"
# **Görev:** bilgiBilim adlı bir liste oluşturun. Bu listeye bilgi bilim disiplini ile ilgili 3 adet anahtar kelime ya da kavram ekleyin. Bu listeyi ekrana yazdırın. Listeye istediğiniz bir yöntem ile (append(), +=) 2 yeni öğe ekleyin. Ekrana listenin son durumunu yazdırın. Listenizdeki son öğeyi değiştirin. Listenin son halini ekrana yazıdırn.
# + id="_gzwR_UGLkL4" colab_type="code" outputId="3800a768-1b3e-41c3-b7aa-0a0a8f249f13" colab={"base_uri": "https://localhost:8080/", "height": 85}
#bilgiBilim
acikListe1 = ["açık bilim", "açık veri", "açık erişim"]
print(acikListe1)
acikListe1 += [" Donanım", "inovasyon"]
print(acikListe1)
acikListe1.append("gazetecilik")
print(acikListe1)
acikListe1[4] = "kod"
print(acikListe1)
# + [markdown] id="qiE0PiXLPQmM" colab_type="text"
# ## Liste Öğeleri
# + [markdown] id="S7yYWbwTPXQ8" colab_type="text"
# Liste öğelerini dilimleme (slice) yaparak da atamak mümkündür. Bu bir listenin boyutunu değiştirebilir veya listeyi tamamen temizleyebilir.
# + [markdown] id="rujdtp_XPp9N" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="_ooqgLJbPsXC" colab_type="text"
#
#
# ```
# acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"] # acikListe adında yeni bir liste oluşturur
# print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak']
#
# acikListe[2:4] = ["Açık İnovasyon"] # "Açık Veri" ve "Açık Eğitim" öğelerinin yerine tek bir öğe ekler
# print(acikListe) #Çıktı: ["Açık Bilim", "Açık Erişim", "Açık İnovasyon", "Açık Kaynak"]
#
# acikListe[:2] = [] # listenin ilk iki öğesini siler
# print(acikListe) #Çıktı: ["Açık İnovasyon", "Açık Kaynak"]
#
# acikListe[:] = [] # listeyi temizler
# print(acikListe) #Çıktı: []
# ```
#
#
# + [markdown] id="4C3xLijJSKdE" colab_type="text"
# **Görev:** Önceki görevde oluşturulan "bilgiBilim" adlı listenin istediğiniz öğesini silerek listenin güncel halini ekrana yazdırın. Listeyi tamamen temizleyerek listenin güncel halini ekrana yazdırın.
# + id="uQPbQgJAQ8nq" colab_type="code" outputId="56801723-3bb0-42f5-89cf-23c764e98f8d" colab={"base_uri": "https://localhost:8080/", "height": 68}
#bilgiBilim
acikListe = ["Bilim", "Erişim", "Veri", "Eğitim"]
print(acikListe)
acikListe[:2] = []
print(acikListe)
acikListe[:] = []
print(acikListe)
# + [markdown] id="J_f83oCDS-Hz" colab_type="text"
# ## Demetler (Tuples)
# + [markdown] id="9er0WsfhTBpI" colab_type="text"
# Demetler neredeyse listelerle aynı. Demetler ve listeler arasındaki tek önemli fark, demetlerin değiştirilememesidir. Demetlere öğe eklenmez, öğe değiştirilmez veya demetlerden öğe silinemez. Demetler, parantez içine alınmış bir virgül operatörü tarafından oluşturulur. Ör: demet = ("deger1", "deger2", "deger3"). Tek bir öğe demetinde ("d",) gibi bir virgül olmalıdır.
# + [markdown] id="yvlCXOLOTq0o" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="h95qzA42Tti-" colab_type="text"
#
#
# ```
# ulkeKodlari = ("TR", "US", "EN", "JP")
# print(ulkeKodlari) # Çıktı: ('TR', 'US', 'EN', 'JP')
# ```
#
#
# + [markdown] id="ON1kJ2-nU2ZO" colab_type="text"
# **Görev:** <NAME>phanesi konu başlıkları listesinin kodlarından oluşan bir demet oluşturun ve ekrana yazdırın! Oluşturulan demet içindeki tek bir öğeyi ekrana yazdırın!
# + id="p4i7BTX1Uv9S" colab_type="code" outputId="5a0e55dc-83f7-44a9-fe67-6aba1feb1bb7" colab={"base_uri": "https://localhost:8080/", "height": 34}
#konuBasliklari
kkod = ("CB", "CT", "DS")
print(kkod)
# + [markdown] id="HkDm5do0Vduc" colab_type="text"
# ## Sözlükler
# Sözlük, listeye benzer, ancak sözlük içindeki değerlere indeks numarası yerine bir anahtara bakarak erişilebilir. Bir anahtar herhangi bir metin katarı veya rakam olabilir. Sözlükler ayraç içine alınır. Ör: sozluk = {'anahtar1': "değer1", 'anahtar2': "değer2"}.
# + [markdown] id="KAS6zUyhV6BQ" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="e1BbS2f3V9le" colab_type="text"
#
#
# ```
# adresDefteri = {"Hacettepe Üniversitesi": "hacettepe.edu.tr", "ODTÜ": "odtu.edu.tr", "Bilkent Üniversitesi": "bilkent.edu.tr"} # yeni bir sözlük oluşturur
# print(adresDefteri) # Çıktı: {'Hacettepe Üniversitesi': 'hacettepe.edu.tr', 'ODTÜ': 'odtu.edu.tr', 'Bilkent Üniversitesi': 'bilkent.edu.tr'}
#
# adresDefteri["Ankara Üniversitesi"] = "ankara.edu.tr" #sözlüğe yeni bir öğe ekler
# print(adresDefteri) # Çıktı: {'Hacettepe Üniversitesi': 'hacettepe.edu.tr', 'ODTÜ': 'odtu.edu.tr', 'Bilkent Üniversitesi': 'bilkent.edu.tr', 'Ankara Üniversitesi': 'ankara.edu.tr'}
#
# del adresDefteri ["Ankara Üniversitesi"] #sözlükten belirtilen öğeyi siler
# print(adresDefteri) # Çıktı: {'Hacettepe Üniversitesi': 'hacettepe.edu.tr', 'ODTÜ': 'odtu.edu.tr', 'Bilkent Üniversitesi': 'bilkent.edu.tr'}
# ```
#
#
# + [markdown] id="fDWQzlKTXYbo" colab_type="text"
# **Görev:** İstediğin herhangi bir konuda 5 öğeye sahip bir sözlük oluştur. Sözlüğü ekrana yazdır. Sözlükteki belirli bir öğeyi ekrana yazdır. Sözlükteki belirli bir öğeyi silerek sözlüğün güncel halini ekrana yazdır!
# + id="t4-oPZTRXEuQ" colab_type="code" outputId="76cff985-fc1d-4e14-cca6-d9ba53a9423c" colab={"base_uri": "https://localhost:8080/", "height": 68}
#sozluk
makyaj = {"Rimel": "kirpik", "far": "göz", "ruj": "dudak"}
print(makyaj)
makyaj["allık"] = "yüz"
print(makyaj)
del makyaj ["Rimel"]
print(makyaj)
# + [markdown] id="9qqctZNZX6q4" colab_type="text"
# ## Sözlük Değerleri ve Anahtarları
# Sözlüklerde values() ve keys() gibi birçok yararlı fonksiyon vardır. Bir sozlük adı ve ardından noktadan sonra çıkan listeyi kullanarak geri kalan fonksiyolar incelenebilir.
# + [markdown] id="SSfVp3ITYa42" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="EaYu0M-1Ydia" colab_type="text"
#
#
# ```
# adresDefteri = {"Hacettepe Üniversitesi": "hacettepe.edu.tr", "ODTÜ": "odtu.edu.tr", "Bilkent Üniversitesi": "bilkent.edu.tr"} # yeni bir sözlük oluşturur
# print(adresDefteri) # Çıktı: {'Hacettepe Üniversitesi': 'hacettepe.edu.tr', 'ODTÜ': 'odtu.edu.tr', 'Bilkent Üniversitesi': 'bilkent.edu.tr'}
#
# print(adresDefteri.values()) # Çıktı: dict_values(['hacettepe.edu.tr', 'odtu.edu.tr', 'bilkent.edu.tr'])
#
# print(adresDefteri.keys()) # Çıktı: dict_keys(['Hacettepe Üniversitesi', 'ODTÜ', 'Bilkent Üniversitesi'])
# ```
#
#
# + [markdown] id="twxTKZ-XY_5l" colab_type="text"
# **Görev:** İstediğin bir konuda istediğin öğe saysına sahip bir sözlük oluştur. Sözlükler ile ilgili farklı fonksiyoları dene. Sonuçları ekrana yazdır!
# + id="CcNcGoOaZfja" colab_type="code" outputId="77ab4f0c-0d8a-4f0d-d2f2-fca15c5a043a" colab={"base_uri": "https://localhost:8080/", "height": 68}
#yeniSozluk
meyve = {"Elma": "elma.com", "erik": "erik.com", "çilek": "çilek.com"}
print(meyve)
print(meyve.values())
print(meyve.keys())
# + [markdown] id="iFlaNCzFpdIj" colab_type="text"
# ##In Anahtar Kelimesi
# "In" anahtar sözcüğü, bir listenin veya sözlüğün belirli bir öğe içerip içermediğini kontrol etmek için kullanılır. Daha önce metin katarlarındaki kullanıma benzer bir kullanımı vardır. "In" anahtar sözcüğü ile öğe kontrolü kontrolü yapıldıktan sonra sonuç öğe listede ya da sözlükte yer alıyorsa *True* yer almıyorsa *False* olarak geri döner.
#
# **Önemli**: Aranan öğe ile liste ya da sözlük içinde yer alan öğelerin karşılaştırılması sırasında büyük-küçük harf duyarlılığı bulunmaktadır. Ör: "Bilgi" ve "bilgi" iki farklı öğe olarak değerlendirilir.
# + [markdown] id="0oPHzw0wq5rw" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="rZ-pjki1q6bs" colab_type="text"
#
#
# ```
# bilgiKavramları = ["indeks", "erişim", "koleksiyon"] # yeni bir liste oluşturur
#
# print("Erişim" in bilgiKavramları) # Çıktı: False
#
# bilgiSozlugu = {"indeks": "index", "erişim": "access", "koleksiyon": "collection"} # yeni bir sozluk oluşturur
#
# print("koleksiyon" in bilgiSozlugu.keys()) # çıktı: True
# ```
#
#
# + [markdown] id="Pdn_zG-mq9Sx" colab_type="text"
# **Görev:** Bir liste ve bir sözlük oluşturun. Liste içinde istediğiniz kelimeyi aratın ve sonucunu ekrana yazdırın! Oluşturduğunuz sözlüğün içinde hem anahtar kelime (keys()) hem de değer (values()) kontrolü yaptırın ve sonucunu ekrana yazdırın!
# + id="3Sl_XODcq1mL" colab_type="code" outputId="ac9430f2-2332-43b9-d598-6f4f5db21843" colab={"base_uri": "https://localhost:8080/", "height": 68}
#yeniListe
#yeniSozluk
kavramlar = ["indeks", "erişim", "koleksiyon"]
print(kavramlar)
print("erişim" in kavramlar)
sozluk = ["indeks", "erişim", "koleksiyon"]
print("Koleksiyon" in sozluk)
# + [markdown] id="o68yxOoTx1vA" colab_type="text"
# #Bölüm 05: Koşullu İfadeler
# + [markdown] id="0R6l7TiLyI1H" colab_type="text"
# ##Mantıksal Operatörler
# Mantıksal operatörler ifadeleri karşılaştırır ve sonuçları *True* ya da *False* değerleriyle döndürür. Python'da üç tane mantıksal operatör bulunur:
#
# 1. "and" operatörü: Her iki yanındaki ifadeler doğru olduğunda *True* değerini döndürür.
# 2. "or" operatörü: Her iki tarafındaki ifadelerden en az bir ifade doğru olduğunda "True" değerini döndürür.
# 3. "not" operatörü: İfadenin tam tersi olarak değerlendirilmesini sağlar.
#
# + [markdown] id="Rp86gPKQ1oau" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="6HZvGmjG1rxk" colab_type="text"
#
#
# ```
# kullaniciAdi = "orcunmadran"
# sifre = 123456
# print(kullaniciAdi == "orcunmadran" and sifre == 123456) # Çıktı: True
#
# kullaniciAdi = "orcunmadran"
# sifre = 123456
# print(kullaniciAdi == "orcunmadran" and not sifre == 123456) # Çıktı: False
#
# cepTel = "05321234567"
# ePosta = "<EMAIL>"
# print(cepTel == "" or ePosta == "<EMAIL>" ) # Çıktı: True
# ```
#
#
# + [markdown] id="nwkgY8TZ5NcI" colab_type="text"
# **Görev:** Klavyeden girilen kullanıcı adı ve şifrenin kayıtlı bulunan kullanıcı adı ve şifre ile uyuşup uyuşmadığını kontrol edin ve sonucu ekrana yazdırın!
# + id="XMHB4_dY3B1v" colab_type="code" outputId="63370aa2-0279-4038-bf41-dd1a019b88a1" colab={"base_uri": "https://localhost:8080/", "height": 68}
#Sistemde yer alan bilgiler:
sisKulAdi = "yonetici"
sisKulSifre = "bby162"
#Klavyeden girilen bilgiler:
girKulAdi = input("Kullanıcı Adı: ")
girKulSifre = input("Şifre: ")
#Kontrol
#Sonuç
print(sisKulAdi == "yonetici" and not sisKulSifre =="bby162")
# + [markdown] id="rd-bsVch9Dxb" colab_type="text"
# ## If Cümleciği
# "If" anahtar sözcüğü, verilen ifadenin doğru olup olmadığını kontrol ettikten sonra belirtilen kodu çalıştıran bir koşullu ifade oluşturmak için kullanılır. Python'da kod bloklarının tanımlanması için girinti kullanır.
# + [markdown] id="3Cba5E7H976e" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="VujnXgWV-B_u" colab_type="text"
#
#
# ```
# acikKavramlar = ["bilim", "erişim", "veri", "eğitim"]
# kavram = input("Bir açık kavramı yazın: ")
# if kavram in acikKavramlar:
# print(kavram + " açık kavramlar listesinde yer alıyor!")
# ```
#
#
# + [markdown] id="mwAnF_eiHj-g" colab_type="text"
# **Görev:** "acikSozluk" içinde yer alan anahtarları (keys) kullanarak eğer klavyeden girilen anahtar kelime sözlükte varsa açıklamasını ekrana yazdırın!
# + id="-Knt3IBI_BRm" colab_type="code" outputId="f11ae7b7-f8dd-4a60-b3e5-83fc2a83817f" colab={"base_uri": "https://localhost:8080/", "height": 51}
acikSozluk = {
"Açık Bilim" : "Bilimsel bilgi kamu malıdır. Bilimsel yayınlara ve verilere açık erişim bir haktır." ,
"Açık Erişim" : "Kamu kaynakları ile yapılan araştırmalar sonucunda üretilen yayınlara ücretsiz erişim" ,
"Açık Veri" : "Kamu kaynakları ile yapılan araştırma sonucunda üretilen verilere ücretsiz ve yeniden kullanılabilir biçimde erişim"
}
anahtar = input("Anahtar Kelime: ")
if anahtar in acikSozluk:
print(anahtar + " kelimesi yer alıyor, tebrikler!")
#If
# + [markdown] id="gBj0pSOoLvWu" colab_type="text"
# ## Else ve Elif Kullanımı
# "If" cümleciği içinde ikinci bir ifadenin doğruluğunun kontrolü için "Elif" ifadesi kullanılır. Doğruluğu sorgulanan ifadelerden hiçbiri *True* döndürmediği zaman çalışacak olan kod bloğu "Else" altında yer alan kod bloğudur.
# + [markdown] id="B6Dc3QHlMcRF" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="k-jKSvQcMeu6" colab_type="text"
#
#
# ```
# gunler = ["Pazartesi", "Çarşamba", "Cuma"]
# girilen = input("Gün giriniz: ")
# if girilen == gunler[0]:
# print("Programlama Dilleri")
# elif girilen == gunler[1]:
# print("Kataloglama")
# elif girilen == gunler[2]:
# print("Bilimsel İletişim")
# else :
# print("Kayıtlı bir gün bilgisi girmediniz!")
# ```
#
#
# + [markdown] id="5JzZRGxWr3Sm" colab_type="text"
# **Görev:** Klavyeden girilen yaş bilgisini kullanarak ekrana aşağıdaki mesajları yazdır:
#
# * 21 yaş altı ve 64 yaş üstü kişilere: "Sokağa çıkma yasağı bulunmaktadır!"
# * Diğer tüm kişilere: "Sokağa çıkma yasağı yoktur!"
# * Klavyeden yaş harici bir bilgi girişi yapıldığında: "Yaşınızı rakam olarak giriniz!"
# + id="vEFr1ub_thtK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="f17a5c53-ff88-448d-d068-2851d6c9c126"
yas = int(input("yaşınızı giriniz: "))
if yas == yas < 20:
print("Sokağa çıkamazsın!")
elif yas == yas > 64:
print("Sokağa çıkamazsın!")
else:
print("yasak yoktur!")
# + [markdown] id="mLq4XzRPHdHl" colab_type="text"
# # Bölüm 06: Döngüler
#
# + [markdown] id="yuku4yibHvy5" colab_type="text"
# ## for Döngüsü
# for döngüleri belirli komut satırını ya da satırlarını yinelemek (tekrar etmek) için kullanılır. Her yinelemede, for döngüsünde tanımlanan değişken listedeki bir sonraki değere otomatik olarak atanacaktır.
# + [markdown] id="oPU0aQyFIcmf" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="JjwFU3rcIfzO" colab_type="text"
#
#
# ```
# for i in range(5): # i değerine 0-4 arası indeks değerleri otomatik olarak atanır
# print(i) # Çıktı: Bu komut satırı toplam 5 kere tekrarlanır ve her satırda yeni i değeri yazdırılır
#
#
# konular = ["<NAME>", "<NAME>", "<NAME>"] # yeni bir liste oluşturur
#
# for konu in konular:
# print(konu) #Çıktı: Her bir liste öğesi alt alta satırlara yazdırılır
#
# ```
#
#
# + [markdown] id="cbpRcW1aKFYu" colab_type="text"
# **Görev:** Bir liste oluşturun. Liste öğelerini "for" döngüsü kullanarak ekrana yazdırın!
# + id="ry1L7GJ_J_wZ" colab_type="code" outputId="36f1a013-33bd-4186-c203-d2c5340b34c2" colab={"base_uri": "https://localhost:8080/", "height": 85}
#liste
konular = ["renk", "biçim", "doku", "vücut"]
for konu in konular:
print(konu)
# + [markdown] id="rHicjY1WKk5o" colab_type="text"
# ## Metin Katarların for Döngüsü Kullanımı
# Metin Katarları üzerinde gerçekleştirilebilecek işlemler Python'daki listelerle büyük benzerlik taşırlar. Metin Katarını oluşturan öğeler (harfler) liste elemanları gibi "for" döngüsü yardımıyla ekrana yazdırılabilir.
# + [markdown] id="CC6FYNwGLa61" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="UlkXx7vOLdhn" colab_type="text"
#
#
# ```
# cumle = "Bisiklet hem zihni hem bedeni dinç tutar!"
#
# for harf in cumle: # Cümledeki her bir harfi ekrana satır satır yazdırır
# print(harf)
# ```
#
#
# + [markdown] id="y_7DFSavMJO4" colab_type="text"
# **Görev:** İçinde metin katarı bulunan bir değişken oluşturun. Bu değişkende yer alan her bir harfi bir satıra gelecek şekilde "for" döngüsü ile ekrana yazdırın!
# + id="83a-E21mMEUr" colab_type="code" outputId="4003131d-8eb3-40ac-e22b-aaa44fbbb48a" colab={"base_uri": "https://localhost:8080/", "height": 544}
#degisken
cumle = "Nemlendirici krem almam gerek.."
for harf in cumle:
print(harf)
# + [markdown] id="mSp_IsCyM7Ed" colab_type="text"
# ## while Döngüsü
# "While" döngüsü "if" cümleciğinin ifade şekline benzer. Koşul doğruysa döngüye bağlı kod satırı ya da satırları yürütülür (çalıştırılır). Temel fark, koşul doğru (True) olduğu olduğu sürece bağlı kod satırı ya da satırları çalışmaya devam eder.
# + [markdown] id="9IL4WNXrOEJ2" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="O0pVXw8jOGjc" colab_type="text"
#
#
# ```
# deger = 1
#
# while deger <= 10:
# print(deger) # Bu satır 10 kez tekrarlanacak
# deger += 1 # Bu satır da 10 kez tekrarlanacak
#
# print("Program bitti") # Bu satır sadece bir kez çalıştırılacak
# ```
#
#
# + [markdown] id="CBGdLeJbSEev" colab_type="text"
# ## break Anahtar Kelimesi
# Asla bitmeyen döngüye sonsuz döngü adı verilir. Döngü koşulu daima doğru (True) olursa, böyle bir döngü sonsuz olur. "Break" anahtar kelimesi geçerli döngüden çıkmak için kullanılır.
# + [markdown] id="99WbOWjYSk7U" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="ihLhjXWcSnv8" colab_type="text"
#
#
# ```
# sayi = 0
#
# while True: # bu döngü sonsuz bir döngüdür
# print(sayi)
# sayi += 1
# if sayi >= 5:
# break # sayı değeri 5 olduğunda döngü otomatik olarak sonlanır
# ```
#
#
# + colab_type="code" id="xXDmHHe2TVLJ" outputId="a3f082b8-131d-4bf6-84bc-a24163c48660" colab={"base_uri": "https://localhost:8080/", "height": 102}
#Örnek Uygulamayı çalıştırarak gözlemleyelim.
sayi = 0
while True: # bu döngü sonsuz bir döngüdür
print(sayi)
sayi += 1
if sayi >= 5:
break # sayı değeri 5 olduğunda döngü otomatik olarak sonlanır
# + [markdown] id="ySFedvaOTqjR" colab_type="text"
# ## continue Anahtar Kelimesi
# "continue" anahtar kelimesi, o anda yürütülen döngü için döngü içindeki kodun geri kalanını atlamak ve "for" veya "while" deyimine geri dönmek için kullanılır.
# + [markdown] id="Ge8XX1mbT0Me" colab_type="text"
#
#
# ```
# for i in range(5):
# if i == 3:
# continue # i değeri 3 olduğu anda altta yer alan "print" komutu atlanıyor.
# print(i)
# ```
#
#
# + id="8kRu3DpFT4pe" colab_type="code" outputId="039c5e37-82c5-4ae6-c39e-52ab9adaa521" colab={"base_uri": "https://localhost:8080/", "height": 170}
#Örnek Uygulamayı çalıştırarak gözlemleyelim.
for i in range(10):
if i == 3:
continue
print(i)
# + [markdown] id="bvvGF4PTO3MX" colab_type="text"
# **Görev: Tahmin Oyunu**
#
# "while" döngüsü kullanarak bir tahmin oyunu tasarlayın. Bu tahmin oyununda, önceden belirlenmiş olan kelime ile klavyeden girilen kelime karşılaştırılmalı, tahmin doğru ise oyun "Bildiniz..!" mesajı ile sonlanmalı, yanlış ise tahmin hakkı bir daha verilmeli.
# + colab_type="code" id="aPxS3byjrjEN" colab={"base_uri": "https://localhost:8080/", "height": 129} outputId="a3f2a233-6382-47b2-a540-250998723bac"
#Tahmin Oyunu
kelime = "ruj"
tahmin = ""
while kelime != tahmin
tahmin = input("Tahmin ediniz: ")
if kelime == tahmin :
print("Tebrikler")
# + [markdown] id="yWfAoCUc-D9d" colab_type="text"
# # Bölüm 07: Fonksiyonlar
# + [markdown] id="YJrL9elb-LvL" colab_type="text"
# ## Tanımlama (Definition)
# Fonksiyonlar, yazılan kodu faydalı bloklara bölmenin, daha okunabilir hale getirmenin ve tekrar kullanmaya yardımcı olmanın kullanışlı bir yoludur. Fonksiyonlar "def" anahtar sözcüğü ve ardından fonksiyonun adı kullanılarak tanımlanır.
# + [markdown] id="pfj4kKSW-4DJ" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="AY162tKL-6Nd" colab_type="text"
#
#
# ```
# def merhaba_dunya(): # fonksiyon tanımlama, isimlendirme
# print("<NAME>!") #fonksiyona dahil kod satırları
#
# for i in range(5):
# merhaba_dunya() # fonksiyon 5 kere çağırılacak
# ```
#
#
# + [markdown] id="UnHQO9MR_pII" colab_type="text"
# ##Fonksiyolarda Parametre Kullanımı
# Fonksiyon parametreleri, fonksiyon adından sonra parantez () içinde tanımlanır. Parametre, iletilen bağımsız değişken için değişken adı görevi görür.
# + [markdown] id="pov3U3OQ_1_l" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="IDgYf17j_4fu" colab_type="text"
#
#
# ```
# def foo(x): # x bir fonksiyon parametresidir
# print("x = " + str(x))
#
# foo(5) # 5 değeri fonksiyona iletilir ve değer olarak kullanılır.
# ```
#
#
# + id="BKerIDjcpfyE" colab_type="code" outputId="64e1e98b-cebd-49cd-8061-ebfae661e9be" colab={"base_uri": "https://localhost:8080/", "height": 51}
def karsila(kAd, kSoyad):
print("Hoşgeldin, "+kAd+" "+kSoyad)
karsila("Orçun","Madran")
karsila("Ayşe","Gel")
# + [markdown] id="LKBMne6BAhid" colab_type="text"
# ##Return Değeri
# Fonksiyonlar, "return" anahtar sözcüğünü kullanarak fonksiyon sonucunda bir değer döndürebilir. Döndürülen değer bir değişkene atanabilir veya sadece yazdırmak için kullanılabilir.
# + [markdown] id="h5kjonbFA7ey" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="fJuWpMooBV0x" colab_type="text"
#
#
# ```
# def iki_sayi_topla(a, b):
# return a + b # hesaplama işleminin sonucu değer olarak döndürülüyor
#
# print(iki_sayi_topla(3, 12)) # ekrana işlem sonucu yazdırılacak
# ```
#
#
# + colab_type="code" outputId="da2c011d-23be-40dd-cf39-7af1db3c0036" id="xRukn_KfrsA3" colab={"base_uri": "https://localhost:8080/", "height": 34}
def karsilama(kAd, kSoyad):
return "Hoşgeldin, "+kAd+" "+kSoyad
print(karsilama("Orçun","Madran"))
# + [markdown] id="FXXW68lICB7S" colab_type="text"
# ##Varsayılan Parametreler
# Bazen bir veya daha fazla fonksiyon parametresi için varsayılan bir değer belirtmek yararlı olabilir. Bu, ihtiyaç duyulan parametrelerden daha az argümanla çağrılabilen bir fonksiyon oluşturur.
# + [markdown] id="FgcKjR-wCeAX" colab_type="text"
# **Örnek Uygulama**
# + [markdown] id="fCHKSZpcCglT" colab_type="text"
#
#
# ```
# def iki_sayi_carp(a, b=2):
# return a * b
#
# print(iki_sayi_carp(3, 47)) # verilen iki degeri de kullanır
# print(iki_sayi_carp(3)) # verilmeyen 2. değer yerine varsayılanı kullanır
# ```
#
#
# + [markdown] colab_type="text" id="nbKueZdw2uZp"
# **Örnek Uygulama 2**
# + id="PRtF6Bd_b2Xg" colab_type="code" outputId="73fa9f36-717c-4d86-f452-a4dff038b2e4" colab={"base_uri": "https://localhost:8080/", "height": 119}
#Sayısal Loto Örnek Uygulama
from random import randint
def tahminEt(rakam=6, satir=6, baslangic=1, bitis=49):
i = 0
secilenler = []
for liste in range(rakam):
secilenler.append(0)
for olustur in range(satir):
while i < len(secilenler):
secilen = randint(baslangic, bitis)
if secilen not in secilenler:
secilenler[i] = secilen
i+=1
print(sorted(secilenler))
i=0
tahminEt(10,6,1,60)
# + [markdown] id="lpT_y9p9DB_U" colab_type="text"
# **Görev:** Bu görev genel olarak fonksiyon bölümünü kapsamaktadır. Daha önce yapmış olduğunuz "<NAME>" projesini fonksiyonlar kullanarak oyun bittiğinde tekrar başlatmaya gerek duyulmadan yeniden oynanabilmesine imkan sağlayacak şekilde yeniden kurgulayın. Oyunun farklı sekansları için farklı fonksiyonlar tanımlayarak oyunu daha optimize hale getirmeye çalışın.
# + id="JfxS1H8rEKuR" colab_type="code" colab={}
#Fonksiyonlu <NAME>
# + [markdown] id="Uocer9uZoa_6" colab_type="text"
# # Bölüm 08: Sınıflar ve Nesneler
#
# Bu bölümde:
#
# * Sınıf ve nesne tanımlama,
# * Değişkenlere erişim,
# * self parametresi,
# * init metodu yer almaktadır.
# + [markdown] id="nSklAub4oh9p" colab_type="text"
# ## Sınıf ve Nesne Tanımlama
# Bir nesne değişkenleri ve fonksiyonları tek bir varlıkta birleştirir. Nesneler değişkenlerini ve fonksiyonlarını sınıflardan alır. Sınıflar bir anlamda nesnelerinizi oluşturmak için kullanılan şablonlardır. Bir nesneyi, fonksiyonların yanı sıra veri içeren tek bir veri yapısı olarak düşünebilirsiniz. Nesnelerin fonksiyonlarına yöntem (metod) denir.
#
# **İpucu:** Sınıf isimlerinin baş harfi büyük yazılarak Python içindeki diğer öğelerden (değişken, fonksiyon vb.) daha rahat ayırt edilmeleri sağlanır.
# + [markdown] id="ShzCIh8VpgM0" colab_type="text"
# **Örnek Uygulama**
#
#
# ```
# class BenimSinifim: # yeni bir sınıfın tanımlanması
#
# bsDegisken = 4 # sınıf içinde yer alan bir değişken
#
# def bsFonksiyon(self): #sınıf içinde yer alan bir fonksiyon
# print("Benim sınıfımın fonksiyonundan Merhaba!")
#
# benimNesnem = BenimSinifim()
# ```
#
#
# + [markdown] id="5NwBvq6S0r4v" colab_type="text"
# ##Değişkenlere ve Fonksiyonlara Erişim
# Sınıftan örneklenen bir nesnenin içindeki bir değişkene ya da fonksiyona erişmek için öncelikle nesnenin adı daha sonra ise değişkenin ya da fonkiyonun adı çağırılmalıdır (Ör: nesneAdi.degiskenAdi). Bir sınıfın farklı örnekleri (nesneleri) içinde tanımlanan değişkenlerin değerleri değiştirebilir.
# + [markdown] colab_type="text" id="NAcj1XjQn89D"
# **Örnek Uygulama 1**
#
#
# ```
# class BenimSinifim: # yeni bir sınıf oluşturur
# bsDegisken = 3 # sınıfın içinde bir değişken tanımlar
# def bsFonksiyon(self): #sınıfın içinde bir fonksiyon tanımlar
# print("Benim sınıfımın fonksiyonundan Merhaba!")
#
# benimNesnem = BenimSinifim() #sınıftan yeni bir nesne oluşturur
#
# for i in range(benimNesnem.bsDegisken): # oluşturulan nesne üzerinden değişkene ve fonksiyona ulaşılır
# benimNesnem.bsFonksiyon()
#
# benimNesnem.bsDegisken = 5 # sınıfın içinde tanımlanan değişkene yeni değer atanması
#
# for i in range(benimNesnem.bsDegisken):
# benimNesnem.bsFonksiyon()
# ```
#
#
# + id="LpNM7SptoLK-" colab_type="code" colab={}
# Örnek uygulama 1'i gözlemleyelim
class BenimSinifim:
bsDegisken = 3
def bsFonksiyon(self):
print("Benim sınıfımın fonksiyonundan Merhaba!")
benimNesnem = BenimSinifim()
for i in range(benimNesnem.bsDegisken):
benimNesnem.bsFonksiyon()
benimNesnem.bsDegisken = 5
for i in range(benimNesnem.bsDegisken):
benimNesnem.bsFonksiyon()
# + [markdown] id="E0v9sloQ22oP" colab_type="text"
# **Örnek Uygulama 2**
#
#
# ```
# class Bisiklet:
# renk = "Kırmızı"
# vites = 1
# def ozellikler(self):
# ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
# return ozellikDetay
#
# bisiklet1 = Bisiklet()
# bisiklet2 = Bisiklet()
#
# print("Bisiklet 1: " + bisiklet1.ozellikler())
#
# bisiklet2.renk = "Sarı"
# bisiklet2.vites = 22
#
# print("Bisiklet 2: " + bisiklet2.ozellikler())
# ```
#
#
# + id="RPIbXyYvrTlk" colab_type="code" colab={}
# Örnek uygulama 2'i gözlemleyelim
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
return ozellikDetay
bisiklet1 = Bisiklet()
bisiklet2 = Bisiklet()
print("Bisiklet 1: " + bisiklet1.ozellikler())
bisiklet2.renk = "Sarı"
bisiklet2.vites = 22
print("Bisiklet 2: " + bisiklet2.ozellikler())
# + [markdown] id="o8f-zk-y3KvD" colab_type="text"
# ##self Parametresi
# "self" parametresi bir Python kuralıdır. "self", herhangi bir sınıf yöntemine iletilen ilk parametredir. Python, oluşturulan nesneyi belirtmek için self parametresini kullanır.
# + [markdown] id="ZoxlmJep4C6j" colab_type="text"
# **Örnek Uygulama**
#
# Aşağıdaki örnek uygulamada **Bisiklet** sınıfının değişkenleri olan *renk* ve *bisiklet*, sınıf içindeki fonksiyonda **self** parametresi ile birlikte kullanılmaktadır. Bu kullanım şekli sınıftan oluşturulan nesnelerin tanımlanmış değişkenlere ulaşabilmeleri için gereklidir.
# ```
# class Bisiklet:
# renk = "Kırmızı"
# vites = 1
# def ozellikler(self):
# ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
# return ozellikDetay
# ```
#
#
# + id="2gqeiPB9ys1l" colab_type="code" colab={}
# Örnek uygulamada "self" tanımlaması yapılmadığı zaman döndürülen hata kodunu inceleyin
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (renk, vites) #tanımlama eksik
return ozellikDetay
bisiklet1 = Bisiklet()
bisiklet2 = Bisiklet()
print("Bisiklet 1: " + bisiklet1.ozellikler())
bisiklet2.renk = "Sarı"
bisiklet2.vites = 22
print("Bisiklet 2: " + bisiklet2.ozellikler())
# + [markdown] id="1B4iU-FI4wRy" colab_type="text"
# ##__init__ Metodu
# __init__ fonksiyonu, oluşturduğu nesneleri başlatmak için kullanılır. init "başlat" ın kısaltmasıdır. __init__() her zaman yaratılan nesneye atıfta bulunan en az bir argüman alır: "self".
#
# + [markdown] id="FMtmqmiD44ZH" colab_type="text"
# **Örnek Uygulama**
#
# Aşağıdaki örnek uygulamada *sporDali* sınıfının içinde tanımlanan **init** fonksiyonu, sınıf oluşturulduğu anda çalışmaya başlamaktadır. Fonksiyonun ayrıca çağırılmasına gerek kalmamıştır.
# ```
# class sporDali:
# sporlar = ["Yüzme", "Bisiklet", "Koşu"]
# def __init__(self):
# for spor in self.sporlar:
# print(spor + " bir triatlon branşıdır.")
#
# triatlon = sporDali()
# ```
#
#
# + id="5zwuicQu0wm7" colab_type="code" colab={}
# Örnek uygulamayı çalıştır
class sporDali:
sporlar = ["Yüzme", "Bisiklet", "Koşu"]
def __init__(self):
for spor in self.sporlar:
print(spor + " bir triatlon branşıdır.")
triatlon = sporDali()
# + [markdown] id="UbKP1Z_c0d9k" colab_type="text"
# #Bölüm 09: Modüller ve Paketler
# + [markdown] id="KyJG2zkhzLD7" colab_type="text"
# ##Modülün İçe Aktarılması
# Python'daki modüller, Python tanımlarını (sınıflar, fonksiyonlar vb.) ve ifadelerini (değişkenler, listeler, sözlükler vb.) içeren .py uzantısına sahip Python dosyalarıdır.
#
# Modüller, *import* anahtar sözcüğü ve uzantı olmadan dosya adı kullanılarak içe aktarılır. Bir modül, çalışan bir Python betiğine ilk kez yüklendiğinde, modüldeki kodun bir kez çalıştırılmasıyla başlatılır.
# + [markdown] id="15prWEGSz1SN" colab_type="text"
# **Örnek Uygulama**
#
#
#
# ```
# #bisiklet.py adlı modülün içeriği
# """
# Bu modül içinde Bisiklet sınıfı yer almaktadır.
# """
# class Bisiklet:
# renk = "Kırmızı"
# vites = 1
# def ozellikler(self):
# ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
# return ozellikDetay
# ```
#
#
#
# ```
# #bisikletler.py adlı Python dosyasının içeriği
#
# import bisiklet
#
# bisiklet1 = bisiklet.Bisiklet()
#
# print("Bisiklet 1: " + bisiklet1.ozellikler())
#
# ```
#
#
# + [markdown] id="lmb0EwAx-xYG" colab_type="text"
# **PyCharm Örneği**
#
#
# 
#
# bisiklet.py
#
#
#
# ---
#
#
#
# 
#
# bisikletler.py
# + [markdown] id="btdYo-btCCu5" colab_type="text"
# ##Colab'de Modülün İçe Aktarılması
#
# Bir önceki bölümde (Modülün İçe Aktarılması) herhangi bir kişisel bilgisayarın sabit diski üzerinde çalışırken yerleşik olmayan (kendi yazdığımız) modülün içe aktarılması yer aldı.
#
# Bu bölümde ise Colab üzerinde çalışırken yerleşik olmayan bir modülü nasıl içe aktarılacağı yer almakta.
# + [markdown] colab_type="text" id="4KYGmiV4EQwz"
# **Örnek Uygulama**
#
# Aşağıda içeriği görüntülenen *bisiklet.py* adlı Python dosyası Google Drive içerisinde "BBY162_Python_a_Giris.ipynb" dosyasının ile aynı klasör içinde bulunmaktadır.
#
# ```
# #bisiklet.py adlı modülün içeriği
# """
# Bu modül içinde Bisiklet sınıfı yer almaktadır.
# """
# class Bisiklet:
# renk = "Kırmızı"
# vites = 1
# def ozellikler(self):
# ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
# return ozellikDetay
# ```
#
# + id="slM_q141DkRK" colab_type="code" colab={}
# Google Drive'ın bir disk olarak görülmesi
from google.colab import drive
drive.mount('gdrive') # bağlanan diskin 'gdrive' adı ile tanımlanması.
import sys # bağlanan diskin fiziksel yolunun tespit edilmesi ve bağlantı yoluna eklenmesi
sys.path.append('/content/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/')
import bisiklet # bisiklet.py içerisindeki 'bisiklet' sınıfının içe aktarılması
bisiklet1 = bisiklet.Bisiklet()
print("Bisiklet 1: " + bisiklet1.ozellikler())
# + [markdown] id="tVPUaDwZBfrc" colab_type="text"
# ##Yerleşik Modüller (built-in)
# Python aşağıdaki bağlantıda yer alan standart modüllerle birlikte gelir. Bu modüllerin *import* anahtar kelimesi ile çağrılması yeterlidir. Ayrıca bu modüllerin yüklenmesine gerek yoktur.
#
# [Python Standart Modülleri](https://docs.python.org/3/library/)
# + [markdown] id="1UvfkosdCOQW" colab_type="text"
# **Örnek Uygulama**
#
#
# ```
# import datetime
# print(datetime.datetime.today())
# ```
#
#
# + id="CKTfLO0PH3mF" colab_type="code" colab={}
# Örnek uygulamayı çalıştır
import datetime
print(datetime.datetime.today())
# + [markdown] id="nqSNESNxDoVL" colab_type="text"
# ##from import Kullanımı
# İçe aktarma ifadesinin bir başka kullanım şekli *from* anahtar kelimesinin kullanılmasıdır. *from* ifadesi ile modül adları paketin içinde alınarak direkt kullanıma hazır hale getirilir. Bu şekilde, içe aktarılan modül, modül_adı öneki olmadan doğrudan kullanılır.
# + [markdown] colab_type="text" id="ccZnbUVLE9hg"
# **Örnek Uygulama**
#
#
#
# ```
# #bisiklet.py adlı modülün içeriği
# """
# Bu modül içinde Bisiklet sınıfı yer almaktadır.
# """
# class Bisiklet:
# renk = "Kırmızı"
# vites = 1
# def ozellikler(self):
# ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
# return ozellikDetay
# ```
#
# + colab_type="code" id="HxSSqtqMIYpS" colab={}
# Google Drive'ın bir disk olarak görülmesi
from google.colab import drive
drive.mount('gdrive') # bağlanan diskin 'gdrive' adı ile tanımlanması.
import sys # bağlanan diskin fiziksel yolunun tespit edilmesi ve bağlantı yoluna eklenmesi
sys.path.append('/content/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/')
from bisiklet import Bisiklet # bisiklet.py içerisindeki 'bisiklet' sınıfının içe aktarılması
bisiklet1 = Bisiklet() # bisiklet ön tanımlamasına gerek kalmadı
print("Bisiklet 1: " + bisiklet1.ozellikler())
# + [markdown] id="8s0bn98LFdh4" colab_type="text"
# #Bölüm 10: Dosya İşlemleri
# + [markdown] id="cTf7unkXFtRi" colab_type="text"
# ##Dosya Okuma
# Python, bilgisayarınızdaki bir dosyadan bilgi okumak ve yazmak için bir dizi yerleşik fonksiyona sahiptir. **open** fonksiyonu bir dosyayı açmak için kullanılır. Dosya, okuma modunda (ikinci argüman olarak "r" kullanılarak) veya yazma modunda (ikinci argüman olarak "w" kullanılarak) açılabilir. **open** fonksiyonu dosya nesnesini döndürür. Dosyanın saklanması için kapatılması gerekir.
# + [markdown] id="SV9N24JsRqbk" colab_type="text"
# **Örnek Uygulama**
#
#
# ```
# #Google Drive Bağlantısı
# from google.colab import drive
# drive.mount('/gdrive')
#
# dosya = "/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/metin.txt"
#
# f = open(dosya, "r")
#
# for line in f.readlines():
# print(line)
#
# f.close()
# ```
#
# Dosyanın sağlıklı şekilde okunabilmesi için Google Drive ile bağlantının kurulmuş olması ve okunacak dosyanın yolunun tam olarak belirtilmesi gerekmektedir.
#
# 
# + id="3KXhzd_FKMWt" colab_type="code" colab={}
#Google Drive Bağlantısı
from google.colab import drive
drive.mount('/gdrive')
dosya = "/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/metin.txt"
f = open(dosya, "r")
for line in f.readlines():
print(line)
f.close()
# + [markdown] id="pToEIEVUR16I" colab_type="text"
# ##Dosya Yazma
# Bir dosyayı ikinci argüman olarak "w" (yazma) kullanarak açarsanız, yeni bir boş dosya oluşturulur. Aynı ada sahip başka bir dosya varsa silineceğini unutmayın. Mevcut bir dosyaya içerik eklemek istiyorsanız "a" (ekleme) değiştiricisini kullanmalısınız.
# + [markdown] id="aserUhlGcBVX" colab_type="text"
# **Örnek Uygulama**
#
# Aşağıdaki örnekte dosya 'w' parametresi ile açıldığı için var olan dosyanın içindekiler silinir ve yeni veriler dosyaya yazılır. Dosyanın içindeki verilerin kalması ve yeni verilerin eklenmesi isteniyorsa dosya 'a' parametresi ile açılmalıdır.
#
# ```
# #Google Drive Bağlantısı
# from google.colab import drive
# drive.mount('/gdrive')
#
# dosya = "/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/cikti.txt"
#
# f = open(dosya, 'w') # Mevcut veriye ek veri yazılması için parametre: 'a'
# f.write("test") # Her yeni verinin bir alt satıra yazdırılması "test\n"
# f.close()
# ```
#
# Kod çalıştırıldıktan sonra eğer *cikti.txt* adında bir dosya yoksa otomatik olarak oluşturulur ve istenilen içerik yazılır.
#
# 
# + id="JGu95coVKdTn" colab_type="code" colab={}
#Google Drive Bağlantısı
from google.colab import drive
drive.mount('/gdrive')
dosya = "/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/cikti.txt"
f = open(dosya, 'w') # Mevcut veriye ek veri yazılması için parametre: 'a'
f.write("test") # Her yeni verinin bir alt satıra yazdırılması "test\n"
f.close()
| calisma_defteri.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gevent
# +
def foo():
print('Running in foo')
gevent.sleep(0)
print('Explicit context switch to foo again')
def bar():
print('Explicit context to bar')
gevent.sleep(0)
print('Implicit context switch back to bar')
gevent.joinall([
gevent.spawn(foo),
gevent.spawn(bar),
])
# -
import time
import gevent
from gevent import select
# +
start = time.time()
tic = lambda: 'at %1.1f seconds' % (time.time() - start)
def gr1():
# Busy waits for a second, but we don't want to stick around...
print('Started Polling: %s' % tic())
select.select([], [], [], 2)
print('Ended Polling: %s' % tic())
def gr2():
# Busy waits for a second, but we don't want to stick around...
print('Started Polling: %s' % tic())
select.select([], [], [], 2)
print('Ended Polling: %s' % tic())
def gr3():
print("Hey lets do some stuff while the greenlets poll, %s" % tic())
gevent.sleep(1)
gevent.joinall([
gevent.spawn(gr1),
gevent.spawn(gr2),
gevent.spawn(gr3),
])
# -
import threading
x = 0
# +
def foo():
global x
for i in range(10000000):
x += 1
def bar():
global x
for i in range(10000000):
x -= 1
t1 = threading.Thread(target=foo)
t2 = threading.Thread(target=bar)
t1.start()
t2.start()
t1.join()
t2.join()
print(x)
# -
# `print(x)` 의 결과가 0으로 나오는 게 정상적으로 작동할 것이라 생각이 들지만, 실제 계산을 통해 나온 값은 전혀 이상한 숫자가 된다.
# 전역 변수 x 에 두 개의 thread가 동시에 접근해서 각자의 작업을 하면서 어느 한 쪽의 작업 결과가 반영이 되지 않기 때문이다.
# 이렇게 여러 thread가 공유된 데이터를 변경함으로써 발생하는 문제를 `race condition`이라고도 부른다.
# ### mutex
# Thread-safe한 코드를 만들기 위해서 사용하는 것 중 하나가 mutex (mutual exclusion) 이다. 위에서 본 참사를 막기 위해서, 공유되는 메모리의 데이터를 여러 thread가 동시에 사용할 수 없도록 잠그는 일을 mutex가 맡는다.
#
# **mutex에 대한 좋은 비유**
# > 휴대폰이 없던 시절에는 공중 전화를 주로 이용했었다. 거리의 모든 남자들은 각자의 아내에게 전화를 너무나 걸고 싶어한다.
# 어떤 한 남자가 처음으로 공중 전화 부스에 들어가서 그의 사랑하는 아내에게 전화를 걸었다면, 그는 꼭 전화 부스의 문을 꼭 잡고 있어야 한다. 왜냐하면 사랑에 눈이 먼 다른 남자들이 전화를 걸기 위해 시도때도 없이 달려들고 있기 때문이다. 줄 서는 질서 문화 따위는 없다. 심지어 그 문을 놓친다면, 전화 부스에 들이닥친 남자들이 수화기를 뺏어 당신의 아내에게 애정 표현을 할 지도 모른다.
# 아내와의 즐거운 통화를 무사히 마쳤다면, 이제 문을 잡고 있던 손을 놓고 부스 밖으로 나가면 된다. 그러면 공중 전화를 쓰기 위해 달려드는 다른 남자들 중 제일 빠른 한 명이 부스에 들어가서 똑같이 문을 꼭 잡고 그의 아내와 통화할 수 있다.
#
# - thread: 각 남자들
# - mutex: 공중 전화 부스의 문
# - lock: 그 문을 잡고 있는 남자의 손
# - resource: 공중 전화
import sys
a = []
b = a
sys.getrefcount(a)
# - `a`가 처음 만들어 졌을 때의 reference 개수가 하나,
# - `b`에 `a`의 reference를 할당했으므로, 그 개수가 하나 늘어나서 두 개,
# - `sys.getrefcount`함수에 argument로 `a`가 들어가서, 이 함수 내부에서 `a`의 reference 개수를 하나 늘리므로 세 개 (그리고 이 함수가 끝날 때 다시 reference 개수를 하나 줄일 것이다)
#
# 그리고 이 개수가 0이 되면 CPython이 알아서 메모리를 회수한다고 생각할 수 있다.
import gevent
import random
# +
def task(pid):
gevent.sleep(random.randint(0, 2) * 0.001)
print('Task %s done' % pid)
def synchronous():
for i in range(1, 10):
task(i)
def asynchronous():
threads = [gevent.spawn(task, i) for i in range(10)]
gevent.joinall(threads)
print('Synchronous: ')
synchronous()
print('Asynchronous: ')
asynchronous()
# -
import gevent.monkey
gevent.monkey.patch_socket()
import gevent
import requests
# import simplejson as json
import json
# +
def fetch(pid):
response = requests.get('https://api.github.com/events')
json_result = response.json()
# json_result = json.loads(result)
result_id = json_result[0]['id']
print('Process %s: %s' % (pid, result_id))
return result_id
def synchronous():
for i in range(1, 10):
fetch(i)
def asynchronous():
threads = []
for i in range(1, 10):
threads.append(gevent.spawn(fetch, i))
gevent.joinall(threads)
# -
print('Synchronous:')
synchronous()
print('Asynchronous:')
asynchronous()
import time
def echo(i):
time.sleep(0.001)
return i
from multiprocessing import Pool
p = Pool(10)
run1 = [a for a in p.imap_unordered(echo, range(10))]
run2 = [a for a in p.imap_unordered(echo, range(10))]
run3 = [a for a in p.imap_unordered(echo, range(10))]
run4 = [a for a in p.imap_unordered(echo, range(10))]
print(run1 == run2 == run3 == run4)
from gevent.pool import Pool
p = Pool(10)
run1 = [a for a in p.imap_unordered(echo, range(10))]
run2 = [a for a in p.imap_unordered(echo, range(10))]
run3 = [a for a in p.imap_unordered(echo, range(10))]
run4 = [a for a in p.imap_unordered(echo, range(10))]
print(run1 == run2 == run3 == run4)
| Gevent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
# ___
# # NLP (Natural Language Processing) with Python
#
# This is the notebook that goes along with the NLP video lecture!
#
# In this lecture we will discuss a higher level overview of the basics of Natural Language Processing, which basically consists of combining machine learning techniques with text, and using math and statistics to get that text in a format that the machine learning algorithms can understand!
#
# Once you've completed this lecture you'll have a project using some Yelp Text Data!
#
# **Requirements: You will need to have NLTK installed, along with downloading the corpus for stopwords. To download everything with a conda installation, run the cell below. Or reference the full video lecture**
# +
# ONLY RUN THIS CELL IF YOU NEED
# TO DOWNLOAD NLTK AND HAVE CONDA
# WATCH THE VIDEO FOR FULL INSTRUCTIONS ON THIS STEP
# Uncomment the code below and run:
# # !conda install nltk #This installs nltk
# import nltk # Imports the library
# nltk.download() #Download the necessary datasets
# -
# ## Get the Data
# We'll be using a dataset from the [UCI datasets](https://archive.ics.uci.edu/ml/datasets/SMS+Spam+Collection)! This dataset is already located in the folder for this section.
# The file we are using contains a collection of more than 5 thousand SMS phone messages. You can check out the **readme** file for more info.
#
# Let's go ahead and use rstrip() plus a list comprehension to get a list of all the lines of text messages:
messages = [line.rstrip() for line in open('smsspamcollection/SMSSpamCollection')]
print(len(messages))
# A collection of texts is also sometimes called "corpus". Let's print the first ten messages and number them using **enumerate**:
for message_no, message in enumerate(messages[:10]):
print(message_no, message)
print('\n')
# Due to the spacing we can tell that this is a [TSV](http://en.wikipedia.org/wiki/Tab-separated_values) ("tab separated values") file, where the first column is a label saying whether the given message is a normal message (commonly known as "ham") or "spam". The second column is the message itself. (Note our numbers aren't part of the file, they are just from the **enumerate** call).
#
# Using these labeled ham and spam examples, we'll **train a machine learning model to learn to discriminate between ham/spam automatically**. Then, with a trained model, we'll be able to **classify arbitrary unlabeled messages** as ham or spam.
#
# From the official SciKit Learn documentation, we can visualize our process:
# <img src='http://www.astroml.org/sklearn_tutorial/_images/plot_ML_flow_chart_3.png' width=600/>
# Instead of parsing TSV manually using Python, we can just take advantage of pandas! Let's go ahead and import it!
import pandas as pd
# We'll use **read_csv** and make note of the **sep** argument, we can also specify the desired column names by passing in a list of *names*.
messages = pd.read_csv('smsspamcollection/SMSSpamCollection', sep='\t',
names=["label", "message"])
messages.head()
# ## Exploratory Data Analysis
#
# Let's check out some of the stats with some plots and the built-in methods in pandas!
messages.describe()
# Let's use **groupby** to use describe by label, this way we can begin to think about the features that separate ham and spam!
messages.groupby('label').describe()
# As we continue our analysis we want to start thinking about the features we are going to be using. This goes along with the general idea of [feature engineering](https://en.wikipedia.org/wiki/Feature_engineering). The better your domain knowledge on the data, the better your ability to engineer more features from it. Feature engineering is a very large part of spam detection in general. I encourage you to read up on the topic!
#
# Let's make a new column to detect how long the text messages are:
messages['length'] = messages['message'].apply(len)
messages.head()
# ### Data Visualization
# Let's visualize this! Let's do the imports:
# +
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
messages['length'].plot(bins=50, kind='hist')
# Play around with the bin size! Looks like text length may be a good feature to think about! Let's try to explain why the x-axis goes all the way to 1000ish, this must mean that there is some really long message!
messages.length.describe()
# Woah! 910 characters, let's use masking to find this message:
messages[messages['length'] == 910]['message'].iloc[0]
# Looks like we have some sort of Romeo sending texts! But let's focus back on the idea of trying to see if message length is a distinguishing feature between ham and spam:
messages.hist(column='length', by='label', bins=50,figsize=(12,4))
# Very interesting! Through just basic EDA we've been able to discover a trend that spam messages tend to have more characters. (Sorry Romeo!)
#
# Now let's begin to process the data so we can eventually use it with SciKit Learn!
# ## Text Pre-processing
# Our main issue with our data is that it is all in text format (strings). The classification algorithms that we've learned about so far will need some sort of numerical feature vector in order to perform the classification task. There are actually many methods to convert a corpus to a vector format. The simplest is the the [bag-of-words](http://en.wikipedia.org/wiki/Bag-of-words_model) approach, where each unique word in a text will be represented by one number.
#
#
# In this section we'll convert the raw messages (sequence of characters) into vectors (sequences of numbers).
#
# As a first step, let's write a function that will split a message into its individual words and return a list. We'll also remove very common words, ('the', 'a', etc..). To do this we will take advantage of the NLTK library. It's pretty much the standard library in Python for processing text and has a lot of useful features. We'll only use some of the basic ones here.
#
# Let's create a function that will process the string in the message column, then we can just use **apply()** in pandas do process all the text in the DataFrame.
#
# First removing punctuation. We can just take advantage of Python's built-in **string** library to get a quick list of all the possible punctuation:
# +
import string
mess = 'Sample message! Notice: it has punctuation.'
# Check characters to see if they are in punctuation
nopunc = [char for char in mess if char not in string.punctuation]
# Join the characters again to form the string.
nopunc = ''.join(nopunc)
# -
# Now let's see how to remove stopwords. We can impot a list of english stopwords from NLTK (check the documentation for more languages and info).
from nltk.corpus import stopwords
stopwords.words('english')[0:10] # Show some stop words
nopunc.split()
# Now just remove any stopwords
clean_mess = [word for word in nopunc.split() if word.lower() not in stopwords.words('english')]
clean_mess
# Now let's put both of these together in a function to apply it to our DataFrame later on:
def text_process(mess):
"""
Takes in a string of text, then performs the following:
1. Remove all punctuation
2. Remove all stopwords
3. Returns a list of the cleaned text
"""
# Check characters to see if they are in punctuation
nopunc = [char for char in mess if char not in string.punctuation]
# Join the characters again to form the string.
nopunc = ''.join(nopunc)
# Now just remove any stopwords
return [word for word in nopunc.split() if word.lower() not in stopwords.words('english')]
# Here is the original DataFrame again:
messages.head()
# Now let's "tokenize" these messages. Tokenization is just the term used to describe the process of converting the normal text strings in to a list of tokens (words that we actually want).
#
# Let's see an example output on on column:
#
# **Note:**
# We may get some warnings or errors for symbols we didn't account for or that weren't in Unicode (like a British pound symbol)
# Check to make sure its working
messages['message'].head(5).apply(text_process)
# Show original dataframe
messages.head()
# ### Continuing Normalization
#
# There are a lot of ways to continue normalizing this text. Such as [Stemming](https://en.wikipedia.org/wiki/Stemming) or distinguishing by [part of speech](http://www.nltk.org/book/ch05.html).
#
# NLTK has lots of built-in tools and great documentation on a lot of these methods. Sometimes they don't work well for text-messages due to the way a lot of people tend to use abbreviations or shorthand, For example:
#
# 'Nah dawg, IDK! Wut time u headin to da club?'
#
# versus
#
# 'No dog, I don't know! What time are you heading to the club?'
#
# Some text normalization methods will have trouble with this type of shorthand and so I'll leave you to explore those more advanced methods through the [NLTK book online](http://www.nltk.org/book/).
#
# For now we will just focus on using what we have to convert our list of words to an actual vector that SciKit-Learn can use.
# ## Vectorization
# Currently, we have the messages as lists of tokens (also known as [lemmas](http://nlp.stanford.edu/IR-book/html/htmledition/stemming-and-lemmatization-1.html)) and now we need to convert each of those messages into a vector the SciKit Learn's algorithm models can work with.
#
# Now we'll convert each message, represented as a list of tokens (lemmas) above, into a vector that machine learning models can understand.
#
# We'll do that in three steps using the bag-of-words model:
#
# 1. Count how many times does a word occur in each message (Known as term frequency)
#
# 2. Weigh the counts, so that frequent tokens get lower weight (inverse document frequency)
#
# 3. Normalize the vectors to unit length, to abstract from the original text length (L2 norm)
#
# Let's begin the first step:
# Each vector will have as many dimensions as there are unique words in the SMS corpus. We will first use SciKit Learn's **CountVectorizer**. This model will convert a collection of text documents to a matrix of token counts.
#
# We can imagine this as a 2-Dimensional matrix. Where the 1-dimension is the entire vocabulary (1 row per word) and the other dimension are the actual documents, in this case a column per text message.
#
# For example:
#
# <table border = “1“>
# <tr>
# <th></th> <th>Message 1</th> <th>Message 2</th> <th>...</th> <th>Message N</th>
# </tr>
# <tr>
# <td><b>Word 1 Count</b></td><td>0</td><td>1</td><td>...</td><td>0</td>
# </tr>
# <tr>
# <td><b>Word 2 Count</b></td><td>0</td><td>0</td><td>...</td><td>0</td>
# </tr>
# <tr>
# <td><b>...</b></td> <td>1</td><td>2</td><td>...</td><td>0</td>
# </tr>
# <tr>
# <td><b>Word N Count</b></td> <td>0</td><td>1</td><td>...</td><td>1</td>
# </tr>
# </table>
#
#
# Since there are so many messages, we can expect a lot of zero counts for the presence of that word in that document. Because of this, SciKit Learn will output a [Sparse Matrix](https://en.wikipedia.org/wiki/Sparse_matrix).
from sklearn.feature_extraction.text import CountVectorizer
# There are a lot of arguments and parameters that can be passed to the CountVectorizer. In this case we will just specify the **analyzer** to be our own previously defined function:
# +
# Might take awhile...
bow_transformer = CountVectorizer(analyzer=text_process).fit(messages['message'])
# Print total number of vocab words
print(len(bow_transformer.vocabulary_))
# -
# Let's take one text message and get its bag-of-words counts as a vector, putting to use our new `bow_transformer`:
message4 = messages['message'][3]
print(message4)
# Now let's see its vector representation:
bow4 = bow_transformer.transform([message4])
print(bow4)
print(bow4.shape)
# This means that there are seven unique words in message number 4 (after removing common stop words). Two of them appear twice, the rest only once. Let's go ahead and check and confirm which ones appear twice:
print(bow_transformer.get_feature_names()[4073])
print(bow_transformer.get_feature_names()[9570])
# Now we can use **.transform** on our Bag-of-Words (bow) transformed object and transform the entire DataFrame of messages. Let's go ahead and check out how the bag-of-words counts for the entire SMS corpus is a large, sparse matrix:
messages_bow = bow_transformer.transform(messages['message'])
print('Shape of Sparse Matrix: ', messages_bow.shape)
print('Amount of Non-Zero occurences: ', messages_bow.nnz)
sparsity = (100.0 * messages_bow.nnz / (messages_bow.shape[0] * messages_bow.shape[1]))
print('sparsity: {}'.format(round(sparsity)))
# After the counting, the term weighting and normalization can be done with [TF-IDF](http://en.wikipedia.org/wiki/Tf%E2%80%93idf), using scikit-learn's `TfidfTransformer`.
#
# ____
# ### So what is TF-IDF?
# TF-IDF stands for *term frequency-inverse document frequency*, and the tf-idf weight is a weight often used in information retrieval and text mining. This weight is a statistical measure used to evaluate how important a word is to a document in a collection or corpus. The importance increases proportionally to the number of times a word appears in the document but is offset by the frequency of the word in the corpus. Variations of the tf-idf weighting scheme are often used by search engines as a central tool in scoring and ranking a document's relevance given a user query.
#
# One of the simplest ranking functions is computed by summing the tf-idf for each query term; many more sophisticated ranking functions are variants of this simple model.
#
# Typically, the tf-idf weight is composed by two terms: the first computes the normalized Term Frequency (TF), aka. the number of times a word appears in a document, divided by the total number of words in that document; the second term is the Inverse Document Frequency (IDF), computed as the logarithm of the number of the documents in the corpus divided by the number of documents where the specific term appears.
#
# **TF: Term Frequency**, which measures how frequently a term occurs in a document. Since every document is different in length, it is possible that a term would appear much more times in long documents than shorter ones. Thus, the term frequency is often divided by the document length (aka. the total number of terms in the document) as a way of normalization:
#
# *TF(t) = (Number of times term t appears in a document) / (Total number of terms in the document).*
#
# **IDF: Inverse Document Frequency**, which measures how important a term is. While computing TF, all terms are considered equally important. However it is known that certain terms, such as "is", "of", and "that", may appear a lot of times but have little importance. Thus we need to weigh down the frequent terms while scale up the rare ones, by computing the following:
#
# *IDF(t) = log_e(Total number of documents / Number of documents with term t in it).*
#
# See below for a simple example.
#
# **Example:**
#
# Consider a document containing 100 words wherein the word cat appears 3 times.
#
# The term frequency (i.e., tf) for cat is then (3 / 100) = 0.03. Now, assume we have 10 million documents and the word cat appears in one thousand of these. Then, the inverse document frequency (i.e., idf) is calculated as log(10,000,000 / 1,000) = 4. Thus, the Tf-idf weight is the product of these quantities: 0.03 * 4 = 0.12.
# ____
#
# Let's go ahead and see how we can do this in SciKit Learn:
# +
from sklearn.feature_extraction.text import TfidfTransformer
tfidf_transformer = TfidfTransformer().fit(messages_bow)
tfidf4 = tfidf_transformer.transform(bow4)
print(tfidf4)
# -
# We'll go ahead and check what is the IDF (inverse document frequency) of the word `"u"` and of word `"university"`?
print(tfidf_transformer.idf_[bow_transformer.vocabulary_['u']])
print(tfidf_transformer.idf_[bow_transformer.vocabulary_['university']])
# To transform the entire bag-of-words corpus into TF-IDF corpus at once:
messages_tfidf = tfidf_transformer.transform(messages_bow)
print(messages_tfidf.shape)
# There are many ways the data can be preprocessed and vectorized. These steps involve feature engineering and building a "pipeline". I encourage you to check out SciKit Learn's documentation on dealing with text data as well as the expansive collection of available papers and books on the general topic of NLP.
# ## Training a model
# With messages represented as vectors, we can finally train our spam/ham classifier. Now we can actually use almost any sort of classification algorithms. For a [variety of reasons](http://www.inf.ed.ac.uk/teaching/courses/inf2b/learnnotes/inf2b-learn-note07-2up.pdf), the Naive Bayes classifier algorithm is a good choice.
# We'll be using scikit-learn here, choosing the [Naive Bayes](http://en.wikipedia.org/wiki/Naive_Bayes_classifier) classifier to start with:
from sklearn.naive_bayes import MultinomialNB
spam_detect_model = MultinomialNB().fit(messages_tfidf, messages['label'])
# Let's try classifying our single random message and checking how we do:
print('predicted:', spam_detect_model.predict(tfidf4)[0])
print('expected:', messages.label[3])
# Fantastic! We've developed a model that can attempt to predict spam vs ham classification!
#
# ## Part 6: Model Evaluation
# Now we want to determine how well our model will do overall on the entire dataset. Let's begin by getting all the predictions:
all_predictions = spam_detect_model.predict(messages_tfidf)
print(all_predictions)
# We can use SciKit Learn's built-in classification report, which returns [precision, recall,](https://en.wikipedia.org/wiki/Precision_and_recall) [f1-score](https://en.wikipedia.org/wiki/F1_score), and a column for support (meaning how many cases supported that classification). Check out the links for more detailed info on each of these metrics and the figure below:
# <img src='https://upload.wikimedia.org/wikipedia/commons/thumb/2/26/Precisionrecall.svg/700px-Precisionrecall.svg.png' width=400 />
from sklearn.metrics import classification_report
print (classification_report(messages['label'], all_predictions))
# There are quite a few possible metrics for evaluating model performance. Which one is the most important depends on the task and the business effects of decisions based off of the model. For example, the cost of mis-predicting "spam" as "ham" is probably much lower than mis-predicting "ham" as "spam".
# In the above "evaluation",we evaluated accuracy on the same data we used for training. **You should never actually evaluate on the same dataset you train on!**
#
# Such evaluation tells us nothing about the true predictive power of our model. If we simply remembered each example during training, the accuracy on training data would trivially be 100%, even though we wouldn't be able to classify any new messages.
#
# A proper way is to split the data into a training/test set, where the model only ever sees the **training data** during its model fitting and parameter tuning. The **test data** is never used in any way. This is then our final evaluation on test data is representative of true predictive performance.
#
# ## Train Test Split
# +
from sklearn.model_selection import train_test_split
msg_train, msg_test, label_train, label_test = \
train_test_split(messages['message'], messages['label'], test_size=0.2)
print(len(msg_train), len(msg_test), len(msg_train) + len(msg_test))
# -
# The test size is 20% of the entire dataset (1115 messages out of total 5572), and the training is the rest (4457 out of 5572). Note the default split would have been 30/70.
#
# ## Creating a Data Pipeline
#
# Let's run our model again and then predict off the test set. We will use SciKit Learn's [pipeline](http://scikit-learn.org/stable/modules/pipeline.html) capabilities to store a pipeline of workflow. This will allow us to set up all the transformations that we will do to the data for future use. Let's see an example of how it works:
# +
from sklearn.pipeline import Pipeline
pipeline = Pipeline([
('bow', CountVectorizer(analyzer=text_process)), # strings to token integer counts
('tfidf', TfidfTransformer()), # integer counts to weighted TF-IDF scores
('classifier', MultinomialNB()), # train on TF-IDF vectors w/ Naive Bayes classifier
])
# -
# Now we can directly pass message text data and the pipeline will do our pre-processing for us! We can treat it as a model/estimator API:
pipeline.fit(msg_train,label_train)
predictions = pipeline.predict(msg_test)
print(classification_report(predictions,label_test))
# Now we have a classification report for our model on a true testing set! There is a lot more to Natural Language Processing than what we've covered here, and its vast expanse of topic could fill up several college courses! I encourage you to check out the resources below for more information on NLP!
# ## More Resources
#
# Check out the links below for more info on Natural Language Processing:
#
# [NLTK Book Online](http://www.nltk.org/book/)
#
# [Kaggle Walkthrough](https://www.kaggle.com/c/word2vec-nlp-tutorial/details/part-1-for-beginners-bag-of-words)
#
# [SciKit Learn's Tutorial](http://scikit-learn.org/stable/tutorial/text_analytics/working_with_text_data.html)
# # Good Job!
| Udemy/Refactored_Py_DS_ML_Bootcamp-master/20-Natural-Language-Processing/01-NLP (Natural Language Processing) with Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kalz2q/mycolabnotebooks/blob/master/chartmathb03sequence.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="d98TC-brMOSs"
# メモ
#
# チャート式数学Bの数列のところを読む
# + [markdown] id="toF9pCLrPRoh"
# # 数列
# sequence of numbers か? array, list, sequence, series, 関連して set というのもあるか。
#
# $1,3,5,7,9, \cdots \cdots $
#
# ($\cdots \cdots$ は無限に続くことを表す)
#
# のように、数を 1列に並べたものを、数列といい、その各数を数列の項という。
#
# ということは、数以外のリストは数列とはいわない、ということか。 だから array や list ではなくて、 sequence of numbers なのかも。
#
# 最初の項を初項、第2項、$\cdots\ $ $n$ 番目の項を第 $n$ 項という。
#
# 項といえば、多項式 polynomial という言葉があったが、ここでは関係ないか。
#
# 数列の第 $n$ 項が $n$ の式で表されるとき、**一般項**という。
#
# 有限数列、無限数列は言葉の通り。
#
# $\{ a_n \}$ という略記もある。
#
# + [markdown] id="qhdB-a2gSBGI"
# **等差数列**
#
# 定義 $\quad a_{n+1} = a_n + d \quad $ すなわち $\quad a_{n+1} - a_n = d $ (一定) $\quad d$ は一定
#
# 一般項 $\quad a_n= a+ (n-1)d \quad \longleftarrow$ (第$n$項)=(初項)+($n-1)\times$ (交差)
#
#
# + [markdown] id="FHtdttn4V0ag"
# **等差中項**
# + id="MJe0fYX3MNv4"
| chartmathb03sequence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in this lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/02_Python_Datatypes/tree/main/006_Python_Sets_Methods)**
# </i></small></small>
# # Python Set `symmetric_difference_update()`
#
# The **`symmetric_difference_update()`** method finds the symmetric difference of two sets and updates the set calling it.
#
# The symmetric difference of two sets **`A`** and **`B`** is the set of elements that are in either **`A`** or **`B`**, but not in their intersection.
#
# <div>
# <img src="img/symmetric_difference_update.png" width="250"/>
# </div>
# **Syntax**:
#
# ```python
# A.symmetric_difference_update(B)
# ```
# ## Return Value from `symmetric_difference_update()`
#
# The **`symmetric_difference_update()`** returns None (returns nothing). Rather, it updates the set calling it.
# +
# Example: Working of symmetric_difference_update()
A = {'a', 'c', 'd'}
B = {'c', 'd', 'e' }
result = A.symmetric_difference_update(B)
print('A =', A)
print('B =', B)
print('result =', result)
# -
# Here, the set **`A`** is updated with the symmetric difference of set **`A`** and **`B`**. However, the set **`B`** is unchanged.
# **Recommended Reading:** **[Python Set symmetric_difference()](https://github.com/milaan9/02_Python_Datatypes/blob/main/006_Python_Sets_Methods/014_Python_Set_symmetric_difference%28%29.ipynb)**
| 006_Python_Sets_Methods/015_Python_Set_symmetric_difference_update().ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="SZS0wkHPlyFO" colab_type="text"
# # A: Studying Higgs Boson Analysis. Signal and Background
#
# Sloping background as in real experiment
# + [markdown] id="ZvjolHp0lvEG" colab_type="text"
# ## Part 1 The Background
#
# This file contains the code for the unit "The Elusive Mr. Higgs". It explains the experiment with the Higgs signal and background signal under different settings
#
# In this part we look at the Background
# + id="GBEQu5Gpl-eq" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pylab
# + id="ek9DNgCsla97" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="adbf41b6-2d20-48a8-9a83-ebfa56fe6c7f"
''' Create the background for the setup and plot the same'''
testrand = np.random.rand(42000) #A random array have 42000 elements according to the uniform distribution. The numbers generated will between 0 and 1
Base = 110 + 30* np.random.rand(42000) #Generating a uniform background(base) between a 110 GeV and 140 GeV
index = (1.0 - 0.5* (Base-110)/30) > testrand #To create a boolean index which has True for 100% samples at 110GeV; This percentage reduces linearly as the value for background increases and has ture for 50% samples as 140 GeV
sloping = Base[index] #This generates a sloping background. Here the values in base corresponding the True values in "index" are retained rest are discarded. So this has a distribution as desribed above
####Plotting - Sloping Background.
####See The Plot named "Sloping"
plt.figure("Sloping")
plt.hist(sloping, bins=15, range =(110,140), color = "green")
plt.title("Sloping Background from 42000 events", backgroundcolor = "white")
plt.show()
# + [markdown] id="lL-W-AQSmAwS" colab_type="text"
# ## Part 2 The Higgs Signal
#
# Create a Gaussian Higgs signal(Gaussian signal is what we will get due to error in measurement). The width of the Gaussian indicated the extent of measurement error. This signal has 300 samples.
#
# We compare two histogram choices -- 2 GeV or 0.5 GeV -- each for twop measurement errors 0.5 and 2 GeV
#
# In the next part Part 3, we combine signal and background
# + id="xPG03DMtlnRb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="6c8f2ddb-2c9c-45bd-abc8-f795cc9b0635"
gauss = 2 * np.random.randn(300) +126 #The signal is centered at 126 GeV and has a width of 2.
narrowGauss = 0.5 * np.random.randn(300) +126 #A signal with centered at 126 GeV and width of 0.5(Error in measurement is less than previous signal"
####Plotting - The Higgs Signal with width 2.0.
####See The Plot named "HiggsAlone"
plt.figure("HiggsAlone-1")
plt.hist(gauss, bins=15, range =(110,140), color = "red")
plt.hist(narrowGauss, bins=15, range =(110,140), color = "black")
plt.title("2 Gev and 0.5 GeV Higgs in 2 GeV bins on their own", backgroundcolor = "white")
plt.show()
plt.figure("HiggsAlone-2")
plt.hist(gauss, bins=60, range =(110,140), color = "red")
plt.hist(narrowGauss, bins=60, range =(110,140), color = "black")
plt.title("2 Gev and 0.5 GeV Higgs in 0.5 GeV bins on their own", backgroundcolor = "white")
plt.show()
# + [markdown] id="_8_oudPvmeB-" colab_type="text"
# ## Part 3 Higgs plus Background
#
# Create the actual Signal by combining the Higgs signals(Gaussian shape) and the sloping background signal.
#
# We look at 2 different widths for the Higgs
#
# Note Higgs width is entirely due to measurement error. The Higgs has a natural width as it is NOT stable but rather decays. This natural width can only be estimated at present from observed decay rate and it is < 10 MeV - a factor of more than 100 lower than observed width.
#
# As width coming from measurement errors, it is expected to be Gaussian. It would be a different shape (Breit Wigner) if coming from its Quantum properties
#
# The three plots fix the measurement with at 0.5 GeV and vary your choice as a data analyst; namely the bin size of histogram. The 0.5 GeV bin size seems best
# + id="dH9MuRnrmZkN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 809} outputId="5cbddfee-b407-4e4d-a33a-3a3f81ef7e77"
total = np.concatenate((sloping, gauss)) #Accumalate the sloping Background and gauss(width = 2.0)
narrowTotal = np.concatenate((sloping, narrowGauss)) #Accumalate the sloping background and narrowGauss(width = 0.5)
####Plotting - The 3 kinds of signals(Sloping Background, Higgs Signal, Combined Signal) where Higgs Signal has width 0.5. The plot has 15 bins(each bin represents 2 GeV).
####See The Plot named "Total Narrow Higgs"
plt.figure("Total Narrow Higgs")
plt.hist(narrowTotal, bins=15, range =(110,140), alpha = 0.5, color = "blue") #Total Signal
plt.hist(sloping, bins=15, range =(110,140), alpha = 0.5, color = "green") #Sloping Signal.
plt.hist(narrowGauss, bins=15, range =(110,140), alpha = 0.5, color = "red") #Only the Higgs Signal.
plt.title("0.5 Gev Higgs in 2 GeV bins with Sloping Background", backgroundcolor = "white")
plt.show()
####Plotting - The 3 kinds of signals(Sloping Background, Higgs Signal, Combined Signal) where Higgs Signal has width 0.5. The plot has 30 bins(each bin represents 1GeV).
####See The Plot named "Total Narrow Higgs Bin 1 GeV"
plt.figure("Total Narrow Higgs Bin 1 GeV")
plt.hist(narrowTotal, bins=30, range =(110,140), alpha = 0.5, color = "blue") #Total Signal
plt.hist(sloping, bins=30, range =(110,140), alpha = 0.5, color = "green") #Sloping Signal.
plt.hist(narrowGauss, bins=30, range =(110,140), alpha = 0.5, color = "red") #Only the Higgs Signal.
plt.title("0.5 Gev Higgs in 1 GeV bins with Sloping Background", backgroundcolor = "white")
plt.show()
####Plotting - The 3 kinds of signals(Sloping Background, Higgs Signal, Combined Signal) where Higgs Signal has width 0.5. The plot has 60 bins(each bin represents 0.5GeV).
####See The Plot named "Total Narrow Higgs Bin 0.5 GeV"
plt.figure("Total Narrow Higgs Bin 0.5 GeV")
plt.hist(narrowTotal, bins=60, range =(110,140), alpha = 0.5, color = "blue") #Total Signal
plt.hist(sloping, bins=60, range =(110,140), alpha = 0.5, color = "green") #Sloping Signal.
plt.hist(narrowGauss, bins=60, range =(110,140), alpha = 0.5, color = "red") #Only the Higgs Signal.
plt.title("0.5 Gev Higgs in 0.5 GeV bins with Sloping Background", backgroundcolor = "white")
plt.show()
# + [markdown] id="46PhUyqSHXhP" colab_type="text"
# Here we look at a 2 GeV wide Higgs Boson. It does not stand out!
# + id="3dKVyEtsHhvp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="e68a918d-a467-47f1-8dce-b21d286ce75e"
####Plotting - The 3 kinds of signals(Sloping Background, Higgs Signal, Combined Signal) signal where Higgs Signal has width 2.0. The plot has 15 bins(each bin represents 2GeV).
####See The Plot named "Total Wide Higgs Bin 2 GeV"
plt.figure("Total Wide Higgs Bin 2 GeV")
values, binedges, junk = plt.hist(total, bins=15, range =(110,140), alpha = 0.5, color = "blue") #Total Signal. We also store the histogram values(number of members per bin) and binedges.
plt.hist(sloping, bins=15, range =(110,140), alpha = 0.5, color = "green") #Sloping Signal.
plt.hist(gauss, bins=15, range =(110,140), alpha = 0.5, color = "red") #Only the Higgs Signal.
plt.title("2 Gev Higgs in 2 GeV bins with Sloping Background", backgroundcolor = "white")
plt.show()
# + [markdown] id="PI2-nEdzmrbx" colab_type="text"
# ## Part 4 Error Estimates
#
# Computing the bin Centers and Errors where latter come from rule for counting experiments that an observation of N events has an error sqrt(N)
#
# The error bars are quite small as 2500 has error 50 which is 2%). They are bigger at the end when we reduce total number of events.
# + id="0yFjx39fmpmJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="1e40f63c-7910-4239-d63b-522154d0ae0e"
centers = 0.5*(binedges[1:] + binedges[:-1]) #Computing bin center as the average of its 2 bin-edges.
errors = np.sqrt(values) #Computing expected error as the square root of values.
####Plotting - The 3 kinds of signals(Sloping Background, Higgs Signal, Combined Signal) and expected errors where Higgs Signal has width 2.0. The plot has 60 bins(each bin represents 0.5GeV).
####See The Plot named "Total Wide Higgs Bin 2 GeV with errors"
plt.figure("Total Wide Higgs Bin 2 GeV with errors")
plt.hist(total, bins=15, range =(110,140), alpha = 0.5, color = "blue") #The total signal
plt.hist(sloping, bins=15, range =(110,140), alpha = 0.5, color = "green") #Sloping Signal.
plt.hist(gauss, bins=15, range =(110,140), alpha = 0.5, color = "red") #Only the Higgs Signal.
plt.errorbar(centers, values, yerr = errors, ls='None', marker ='x', color = 'black', markersize= 6.0 ) #The error bar
plt.title("2 Gev Higgs in 2 GeV bins with Sloping Background + Errors", backgroundcolor = "white")
plt.show()
# + [markdown] id="7_8_PR7Em3iC" colab_type="text"
# ## Part 5 A Larger Signal
#
# Creating a Higgs Signal with 30000 elements. If we use this against the original backgrounds, we study the effect of making the Higgs 100 times more likely(The original number of Higgs samples was 300).
# + id="LKRu6vtlm0xA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 597} outputId="35524415-24a5-4d78-fd91-d697ae7bf137"
print('Higgs on its own with no background')
gaussbig = 2 * np.random.randn(30000) +126 #Creating a Higgs Signal with 30000 elements and width of 2.0
gaussnarrowbig = 0.5 * np.random.randn(30000) +126 #Creating a Higgs Signal with 30000 elements and width of 0.5(less error in measurement than previous case)
totalbig = np.concatenate((sloping, gaussbig))
####Plotting - The Pure Higgs Signals with weight 0.5 and 2. The signal with 30,000 elements is used. Plot has 60 bins(each of size .5GeV).
####See The Plot named "30000 Higgs in 0.5 GeV bins"
plt.figure("30000 Higgs in 0.5 GeV bins")
plt.hist(gaussnarrowbig, bins=60, range =(110,140), alpha = 0.5) #Higgs signal
plt.hist(gaussbig, bins=60, range =(110,140), alpha = 0.5)
plt.title("30000 Narrow and Wide Higgs in 0.5 GeV bins", backgroundcolor = "white")
plt.show()
print('\nLarge Higgs signal on original size background')
####Plotting - The 3 kinds of signals(Sloping Background, Higgs Signal, Combined Signal) where Higgs Signal has width 2.0 and is 100 times more frequent. The plot has 15 bins(each bin represents 2GeV).
####See The Plot named "Total Wide Higgs Bin 2 GeV 100 times Higgs"
plt.figure("Total Wide Higgs Bin 2 GeV 100 times Higgs")
plt.hist(totalbig, bins=15, range =(110,140), alpha = 0.5) #Total signal with Higgs being 100 times more frequent
plt.hist(sloping, bins=15, range =(110,140), alpha = 0.5) #sloping background
plt.hist(gaussbig, bins=15, range =(110,140), alpha = 0.5) #Higgs signal with 30,000 elements
plt.title("Total Wide Higgs Bin 2 GeV 100 times Higgs", backgroundcolor = "white")
plt.show()
# + [markdown] id="5LFswql-m_N8" colab_type="text"
# ## Part 6 A Small Experiment (1% original)
#
# Creating a setup with 1% data. Now the background has initially 420 elements
#
# This data fluctuates too much and you wont see any signal from Higgs (only 3 particles expected)
# + id="b9iXy_4am-V7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="c337c742-cad5-4753-ac79-99b98ad870b8"
testrand420 = np.random.rand(420) #Creating random array with 420 elements.
Base420 = 110 + 30* np.random.rand(420) #Creating Background with 420 events
index420 = (1.0 - 0.5* (Base420-110)/30) > testrand420 #Creating index to get the sloping background.
Sloping420 = Base420[index420] #Creating Sloping Background
#### Plot - Plotting the 1% background data. Plot has 15 bins(each 2GeV)
####See The Plot named "Sloping 420 Events"
plt.figure("Sloping 420 Events")
plt.hist(Sloping420, bins=15, range =(110,140))
plt.title("Sloping Background from 420 events (1%)", backgroundcolor = "white")
plt.show()
# + [markdown] id="Fgkwr3u9nOP1" colab_type="text"
# ## Part 7 Medium Size Experiment
#
# Creating setup with 10% data(4200 elements for background, 30 elements for Higgs signal) and computing the errors.
# + id="T0o9s-mRnKtm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="d66dcaf7-ac21-4f01-b82a-6c785e83ae56"
testrand4200 = np.random.rand(4200) #Creating random array with 4200 elements.
Base4200 = 110 + 30* np.random.rand(4200) #Creating Background with 4200 events
index4200 = (1.0 - 0.5* (Base4200-110)/30) > testrand4200 #Creating index to get the sloping background.
Sloping4200 = Base4200[index4200] #Creating Sloping Background
gauss30 = 2 * np.random.randn(30) +126 #Creating Higgs signal with width 2.0 and center at 126GeV
total10percent = np.concatenate((Sloping4200, gauss30)) #Combining the Higgs signal and sloping background
####Plotting - The 3 kinds of signals(Sloping Background, Higgs Signal, Combined Signal) where Higgs Signal has width 2.0 for 10% of data. The plot has 15 bins(each bin represents 2GeV).
####See The Plot named "Total Sloping Background 10% Data"
plt.figure("Total Sloping Background 10% Data")
values10percent, binedges10percent, junk = plt.hist(total10percent, bins=15, range =(110,140), alpha = 0.5, color = "blue") #Total signal and storing the number values/bin and the bin edges.
plt.hist(Sloping4200, bins=15, range =(110,140), alpha = 0.5, color = "green") #Sloping background
plt.hist(gauss30, bins=15, range =(110,140), alpha = 0.5, color = "red") #Higgs Signal(width 2.0)
plt.title("Total with Sloping Background from 4200 events (10%)", backgroundcolor = "white")
plt.show()
####Plotting - The 3 kinds of signals(Sloping Background, Higgs Signal, Combined Signal) and the error where Higgs Signal has width 2.0 for 10% of data. The plot has 15 bins(each bin represents 2GeV).
####See The Plot named "Total Sloping Background 10% Data with errors"
plt.figure("Total Sloping Background 10% Data with errors")
centers10percent = 0.5*(binedges10percent[1:] + binedges10percent[:-1]) #Computing the bin centers
errors10percent = np.sqrt(values10percent) #Computing expected errors
plt.hist(total10percent, bins=15, range =(110,140), alpha = 0.5, color = "blue") #Plotting the total signal
plt.hist(Sloping4200, bins=15, range =(110,140), alpha = 0.5, color = "green") #Plotting the sloping background
plt.hist(gauss30, bins=15, range =(110,140), alpha = 0.5, color = "red") #Plotting the Higgs Signal
plt.errorbar(centers10percent, values10percent, yerr = errors10percent, ls='None', marker ='x', color = 'black', markersize= 6.0 ) #Plotting the error
plt.title("Total with Sloping Background from 4200 events (10%) + Errors", backgroundcolor = "white")
plt.show()
| content/en/modules/notebooks/E534_Higgs_Discovery_A.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 訓練情感模型
# 分析這篇文章中的李+賢是否有AML風險_將2-2處理好的300個字資料集進行訓練
import transformers
from transformers import BertModel, BertTokenizer, AdamW, get_linear_schedule_with_warmup
import torch
import numpy as np
import pandas as pd
import seaborn as sns
from pylab import rcParams
import matplotlib.pyplot as plt
from matplotlib import rc
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, classification_report
from collections import defaultdict
from textwrap import wrap
from torch import nn, optim
from torch.utils.data import Dataset, DataLoader
# ## 情感分析模型
df = pd.read_csv('step2_2_output_train_data_300.csv')
# 正負資料比較及
class_names = ['negative', 'positive']
ax = sns.countplot(df.article_tags)
plt.xlabel('review sentiment')
ax.set_xticklabels(class_names);
tokenizer = BertTokenizer.from_pretrained("bert-base-chinese")#, do_lower_case=False)
sample_txt = df.iloc[0].article_truncate
tokens = tokenizer.tokenize(sample_txt)
token_ids = tokenizer.convert_tokens_to_ids(tokens)
print(f' Sentence: {sample_txt}')
print(f' Tokens: {tokens}')
print(f'Token IDs: {token_ids}')
MAX_LEN = 300
class GPReviewDataset(Dataset):
def __init__(self, reviews, targets, tokenizer, max_len):
self.reviews = reviews
self.targets = targets
self.tokenizer = tokenizer
self.max_len = max_len
def __len__(self):
return len(self.reviews)
def __getitem__(self, item):
review = str(self.reviews[item])
target = self.targets[item]
encoding = self.tokenizer.encode_plus(
review,
add_special_tokens=True,
max_length=self.max_len,
return_token_type_ids=False,
pad_to_max_length=True,
return_attention_mask=True,
return_tensors='pt',
)
return {
'review_text': review,
'input_ids': encoding['input_ids'].flatten(),
'attention_mask': encoding['attention_mask'].flatten(),
'targets': torch.tensor(target, dtype=torch.long)
}
df_train, df_test = train_test_split(
df,
test_size=0.1,
random_state=38
)
df_val = df_test.copy()
df_train.article_tags.value_counts()#0.08869431270014999
df_test.article_tags.value_counts() #0.09140767824497258
df_test.head()
def create_data_loader(df, tokenizer, max_len, batch_size):
ds = GPReviewDataset(
reviews=df.article_truncate.to_numpy(),
targets=df.article_tags.to_numpy(),
tokenizer=tokenizer,
max_len=max_len
)
return DataLoader(
ds,
batch_size=batch_size,
num_workers=0
)
BATCH_SIZE = 16
train_data_loader = create_data_loader(df_train, tokenizer, MAX_LEN, BATCH_SIZE)
val_data_loader = create_data_loader(df_test, tokenizer, MAX_LEN, BATCH_SIZE)
test_data_loader = create_data_loader(df_test, tokenizer, MAX_LEN, BATCH_SIZE)
# +
PRE_TRAINED_MODEL_NAME = "bert-base-chinese"
bert_model = BertModel.from_pretrained(PRE_TRAINED_MODEL_NAME)
# -
class SentimentClassifier(nn.Module):
def __init__(self, n_classes):
super(SentimentClassifier, self).__init__()
self.bert = BertModel.from_pretrained(PRE_TRAINED_MODEL_NAME)
self.drop = nn.Dropout(p=0.3)
self.out = nn.Linear(self.bert.config.hidden_size, n_classes)
def forward(self, input_ids, attention_mask):
_, pooled_output = self.bert(
input_ids=input_ids,
attention_mask=attention_mask
)
output = self.drop(pooled_output)
return self.out(output)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model = SentimentClassifier(len(class_names))
model = model.to(device)
EPOCHS = 10
optimizer = AdamW(model.parameters(), lr=2e-5, correct_bias=False)
total_steps = len(train_data_loader) * EPOCHS
scheduler = get_linear_schedule_with_warmup(
optimizer,
num_warmup_steps=0,
num_training_steps=total_steps
)
loss_fn = nn.CrossEntropyLoss().to(device)
def train_epoch(
model,
data_loader,
loss_fn,
optimizer,
device,
scheduler,
n_examples
):
model = model.train()
losses = []
correct_predictions = 0
for d in data_loader:
input_ids = d["input_ids"].to(device)
attention_mask = d["attention_mask"].to(device)
targets = d["targets"].to(device)
outputs = model(
input_ids=input_ids,
attention_mask=attention_mask
)
_, preds = torch.max(outputs, dim=1)
loss = loss_fn(outputs, targets)
correct_predictions += torch.sum(preds == targets)
losses.append(loss.item())
loss.backward()
nn.utils.clip_grad_norm_(model.parameters(), max_norm=1.0)
optimizer.step()
scheduler.step()
optimizer.zero_grad()
return correct_predictions.double() / n_examples, np.mean(losses)
def eval_model(model, data_loader, loss_fn, device, n_examples):
model = model.eval()
losses = []
correct_predictions = 0
with torch.no_grad():
for d in data_loader:
input_ids = d["input_ids"].to(device)
attention_mask = d["attention_mask"].to(device)
targets = d["targets"].to(device)
outputs = model(
input_ids=input_ids,
attention_mask=attention_mask
)
_, preds = torch.max(outputs, dim=1)
loss = loss_fn(outputs, targets)
correct_predictions += torch.sum(preds == targets)
losses.append(loss.item())
return correct_predictions.double() / n_examples, np.mean(losses)
# %%time
history = defaultdict(list)
best_accuracy = 0
for epoch in range(EPOCHS):
print(f'Epoch {epoch + 1}/{EPOCHS}')
print('-' * 10)
train_acc, train_loss = train_epoch(
model,
train_data_loader,
loss_fn,
optimizer,
device,
scheduler,
len(df_train)
)
print(f'Train loss {train_loss} accuracy {train_acc}')
val_acc, val_loss = eval_model(
model,
val_data_loader,
loss_fn,
device,
len(df_val)
)
print(f'Val loss {val_loss} accuracy {val_acc}')
print()
history['train_acc'].append(train_acc)
history['train_loss'].append(train_loss)
history['val_acc'].append(val_acc)
history['val_loss'].append(val_loss)
if val_acc > best_accuracy:
torch.save(model.state_dict(), 'best_model_state.bin')
best_accuracy = val_acc
# save model
import os
PATH = os.getcwd()+str('/step3_output_bert_senti.pth')
torch.save(model, PATH)
# +
PATH = os.getcwd()+str('/step3_output_bert_senti.pth')
model2 = torch.load(PATH)
model2.eval()
# -
plt.plot(history['train_acc'], label='train accuracy')
plt.plot(history['val_acc'], label='validation accuracy')
plt.title('Training history')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend()
plt.ylim([0, 1]);
test_acc, _ = eval_model(
model,
test_data_loader,
loss_fn,
device,
len(df_test)
)
#test_acc.item()
test_acc.item() #0.9728643216080403
def get_predictions(model, data_loader):
model = model.eval()
review_texts = []
predictions = []
prediction_probs = []
real_values = []
with torch.no_grad():
for d in data_loader:
texts = d["review_text"]
input_ids = d["input_ids"].to(device)
attention_mask = d["attention_mask"].to(device)
targets = d["targets"].to(device)
outputs = model(
input_ids=input_ids,
attention_mask=attention_mask
)
_, preds = torch.max(outputs, dim=1)
review_texts.extend(texts)
predictions.extend(preds)
prediction_probs.extend(outputs)
real_values.extend(targets)
predictions = torch.stack(predictions).cpu()
prediction_probs = torch.stack(prediction_probs).cpu()
real_values = torch.stack(real_values).cpu()
return review_texts, predictions, prediction_probs, real_values
y_review_texts, y_pred, y_pred_probs, y_test = get_predictions(
model,
test_data_loader
)
print(classification_report(y_test, y_pred, target_names=class_names))
def show_confusion_matrix(confusion_matrix):
hmap = sns.heatmap(confusion_matrix, annot=True, fmt="d", cmap="Blues")
hmap.yaxis.set_ticklabels(hmap.yaxis.get_ticklabels(), rotation=0, ha='right')
hmap.xaxis.set_ticklabels(hmap.xaxis.get_ticklabels(), rotation=30, ha='right')
plt.ylabel('True sentiment')
plt.xlabel('Predicted sentiment');
cm = confusion_matrix(y_test, y_pred)
df_cm = pd.DataFrame(cm, index=class_names, columns=class_names)
show_confusion_matrix(df_cm)
idx = 7
review_text = y_review_texts[idx]
true_sentiment = y_test[idx]
pred_df = pd.DataFrame({
'class_names': class_names,
'values': y_pred_probs[idx]
})
print("\n".join(wrap(review_text)))
print()
print(f'True sentiment: {class_names[true_sentiment]}')
df_test2 = df_test.reset_index()#[df_test.article_tags==1]
df_test2[df_test2.article_tags==1]
pd.Series(np.array(y_test)).value_counts()
y_pred_probs
pd.DataFrame(np.array(y_pred_probs.tolist())).min()
pd.DataFrame(np.array(y_pred_probs.tolist())).max()
# # 預測
# + jupyter={"outputs_hidden": true}
# Model class must be defined somewhere
PATH = 'step3_output_bert_senti.pth'
model2 = torch.load(PATH)
model2.eval()
# -
review_text = "富二代網路警察李家賢查獲一間詐騙集團,協助玉山銀行進行洗錢防制"
encoded_review = tokenizer.encode_plus(
review_text,
max_length=100,#MAX_LEN,
add_special_tokens=True,
return_token_type_ids=False,
pad_to_max_length=True,
return_attention_mask=True,
return_tensors='pt',
)
input_ids = encoded_review['input_ids'].to(device)
attention_mask = encoded_review['attention_mask'].to(device)
output = model2(input_ids, attention_mask)
_, prediction = torch.max(output, dim=1)
print(f'Review text: {review_text}')
print(f'Sentiment : {class_names[prediction]}')
prediction.tolist()[0]
np.sum(output)
np.sum(np.array(output.tolist()).squeeze())
a, prediction = torch.max(output, dim=1)
df_test2.head()
# +
predict=[]
for i in range(len(df_test2)):
review_text = df_test2.iloc[i].article_truncate
encoded_review = tokenizer.encode_plus(
review_text,
max_length=MAX_LEN,
add_special_tokens=True,
return_token_type_ids=False,
pad_to_max_length=True,
return_attention_mask=True,
return_tensors='pt',
)
input_ids = encoded_review['input_ids'].to(device)
attention_mask = encoded_review['attention_mask'].to(device)
output = model(input_ids, attention_mask)
_, prediction = torch.max(output, dim=1)
predict.append( prediction.tolist()[0] )
#predict.append( np.sum(np.array(output.tolist()).squeeze()) )
# -
len(predict)
len(df_test2)
df_test2['predict'] = predict
group_test =df_test2[['article_index','article_tags','article_tags_name','predict']].groupby(['article_index','article_tags_name']).mean()
group_test.predict.value_counts()
group_test.reset_index(inplace=True)
uniq_article = np.unique(group_test.article_index.values) # 文章獨一無二的量
group_test[(group_test.article_index==2)&(group_test.article_tags==1)].article_tags_name.tolist()
# + jupyter={"outputs_hidden": true}
group_test
# + jupyter={"outputs_hidden": true}
predd_list = []
answer_list = []
for i in uniq_article:
print('article:{f}'.format(f = i))
predd = group_test[(group_test.article_index==i)&(group_test.predict>=0.5)].article_tags_name.tolist()
predd_list.append(predd)
print('預測 = '+str(predd))
answer = group_test[(group_test.article_index==i)&(group_test.article_tags==1)].article_tags_name.tolist()
print('答案 = '+str(answer))
answer_list.append(answer)
# + jupyter={"outputs_hidden": true}
predd_list = []
answer_list = []
for i in uniq_article:
print('article:{f}'.format(f = i))
predd = group_test[(group_test.article_index==i)&(group_test.predict>=0.2)].article_tags_name.tolist()
predd_list.append(predd)
print('預測 = '+str(predd))
answer = group_test[(group_test.article_index==i)&(group_test.article_tags==1)].article_tags_name.tolist()
print('答案 = '+str(answer))
answer_list.append(answer)
# + jupyter={"outputs_hidden": true}
predd_list = []
answer_list = []
for i in uniq_article:
print('article:{f}'.format(f = i))
predd = group_test[(group_test.article_index==i)&(group_test.predict>=1.0)].article_tags_name.tolist()
predd_list.append(predd)
print('預測 = '+str(predd))
answer = group_test[(group_test.article_index==i)&(group_test.article_tags==1)].article_tags_name.tolist()
print('答案 = '+str(answer))
answer_list.append(answer)
# -
def score(truth, predict):
if truth == [] and predict != []:
return 0
if truth != [] and predict == []:
return 0
if truth == [] and predict == []:
return 1
recall = len([i for i in truth if i in predict])/len(truth)
precision = len([i for i in predict if i in truth])/len(predict)
try:
return 2/((1/recall)+(1/precision))
except:
return 0
score(answer_list, predd_list)0.6
score(answer_list, predd_list)0.2
score(answer_list, predd_list)#0.8
# +
predict=[]
for i in range(len(df_test2)):
review_text = df_test2.iloc[i].article_truncate
encoded_review = tokenizer.encode_plus(
review_text,
max_length=MAX_LEN,
add_special_tokens=True,
return_token_type_ids=False,
pad_to_max_length=True,
return_attention_mask=True,
return_tensors='pt',
)
input_ids = encoded_review['input_ids'].to(device)
attention_mask = encoded_review['attention_mask'].to(device)
output = model2(input_ids, attention_mask)
_, prediction = torch.max(output, dim=1)
predict.append( prediction.tolist()[0] )
#predict.append( np.sum(np.array(output.tolist()).squeeze()) )
# -
df_test2['predict'] = predict
group_test =df_test2[['article_index','article_tags','article_tags_name','predict']].groupby(['article_index','article_tags_name']).mean()
group_test.reset_index(inplace=True)
uniq_article = np.unique(group_test.article_index.values) # 文章獨一無二的量
# + jupyter={"outputs_hidden": true}
predd_list = []
answer_list = []
for i in uniq_article:
print('article:{f}'.format(f = i))
predd = group_test[(group_test.article_index==i)&(group_test.predict>=0.5)].article_tags_name.tolist()
predd_list.append(predd)
print('預測 = '+str(predd))
answer = group_test[(group_test.article_index==i)&(group_test.article_tags==1)].article_tags_name.tolist()
print('答案 = '+str(answer))
answer_list.append(answer)
# -
score(answer_list, predd_list)#0.2
| step3_bertmodel_300.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !dir
# +
from sklearn import linear_model
import numpy as np
import matplotlib.pyplot as plt
# linear regression
data = np.array([
[0.05,0.12],
[0.18,0.22],
[0.31,0.35],
[0.42,0.38],
[0.5,0.49]])
#split into inputs and putputs
X, y = data[:,0], data[:,1]
X = X.reshape((len(X),1))
lr = linear_model.LinearRegression()
lr.fit(X,y)
#predict
yhat = lr.predict(X)
#plot data and predictions
plt.scatter(X,y)
plt.plot(X,yhat,color='red')
plt.show()
# -
#generar aproximación
lr.predict([[0.42]])
# !start .
import pandas as pd
df = pd.read_excel("Formulario sin título (respuestas).xlsx")
df.head()
# +
X, y = df["Peso"].values, df["Altura"].values
X = X.reshape((len(X),1))
lr = linear_model.LinearRegression()
lr.fit(X,y)
#predict
yhat = lr.predict(X)
#plot data and predictions
plt.scatter(X,y)
plt.plot(X,yhat,color='red')
plt.show()
# +
#nomina de 0-3000
names = ["Fecha","Nombre", "Dependencia", "Puesto", "Estatus", "Tipo", "Percepciones","Deducciones", "Neto"]
df = pd.read_csv("https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_0_3000.csv", names=names, header = None)
df.head()
# +
#nomina de 10001
names = ["Fecha","Nombre", "Dependencia", "Puesto", "Estatus", "Tipo", "Percepciones","Deducciones", "Neto"]
df = pd.read_csv("https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_10001_.csv", names=names, header = None)
df.head()
# -
#nomina de 30001 a 4000
names = ["Fecha","Nombre", "Dependencia", "Puesto", "Estatus", "Tipo", "Percepciones","Deducciones", "Neto"]
df=pd.read_csv("https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_3001_4000.csv", names=names, header = None)
df.head()
#nomina de 4001 a 4500
names = ["Fecha","Nombre", "Dependencia", "Puesto", "Estatus", "Tipo", "Percepciones","Deducciones", "Neto"]
df=pd.read_csv("https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_4001_4500.csv", names=names, header = None)
df.head()
#nomina de 4901 a 4999
names = ["Fecha","Nombre", "Dependencia", "Puesto", "Estatus", "Tipo", "Percepciones","Deducciones", "Neto"]
df=pd.read_csv("https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_4901_4999.csv", names=names, header = None)
df.head()
#nomina de 5001 a 7000
names = ["Fecha","Nombre", "Dependencia", "Puesto", "Estatus", "Tipo", "Percepciones","Deducciones", "Neto"]
df =pd.read_csv("https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_5001_7000.csv", names=names, header = None)
df.head()
#nomina de 7000 a 10000
names = ["Fecha","Nombre", "Dependencia", "Puesto", "Estatus", "Tipo", "Percepciones","Deducciones", "Neto"]
df =pd.read_csv("https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_7000_10000.csv", names=names, header = None)
df.head()
df.describe()
# !dir /b
# +
NominaList = ["https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_0_3000.csv",
"https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_10001_.csv",
"https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_3001_4000.csv",
"https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_4001_4500.csv",
"https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_4901_4999.csv",
"https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_5001_7000.csv",
"https://raw.githubusercontent.com/RoboticaIndustrial/Robotica_2020/master/NominaJalisco/nomina_7000_10000.csv"]
mi_df = lambda x: pd.read_csv(x,usecols = [1,2,3,4,5,6,7,8],names=names,header=None,skiprows=2)
df_from_each_file = (mi_df(f) for f in NominaList)
nomina = pd.concat(df_from_each_file,ignore_index=True)
nomina.head()
# -
nomina.describe()
nomina.plot.scatter("Percepciones","Deducciones",figsize=(12,9))
# +
X, y = nomina["Percepciones"].values, nomina["Deducciones"].values
X = X.reshape((len(X),1))
lr = linear_model.LinearRegression()
lr.fit(X,y)
#predict
yhat = lr.predict(X)
#plot data and predictions
plt.scatter(X,y)
plt.plot(X,yhat,color='red')
plt.show()
| Linear Regressions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Symbolic Partial Derivative Routine
#
# ## Authors: <NAME> & <NAME>
#
# ## This module contains a routine for computing partial derivatives of a mathematical expression that is written as several subexpressions.
#
# **Notebook Status:** <font color='green'><b> Validated </b></font>
#
# **Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented [below](#code_validation). Additionally, this notebook has been validated by checking that results are consistent with exact derivative expressions used in the SEOBNRv3_opt approixment of [LALSuite](https://git.ligo.org/lscsoft/lalsuite).
#
# ### NRPy+ Source Code for this module: [SEOBNR_Derivative_Routine.py](../edit/SEOBNR/SEOBNR_Derivative_Routine.py)
#
# ## Introduction
# $$\label{intro}$$
#
# This notebook documents the symbolic partial derivative routine used to generate analytic derivatives of the [SEOBNRv3](https://git.ligo.org/lscsoft/lalsuite) Hamiltonian (documented [here](../Tutorial-SEOBNR_v3_Hamiltonian.ipynb)) and described in [this article](https://arxiv.org/abs/1803.06346). In general, this notebook takes as input a file of inter-dependent mathematical expressions (in SymPy syntax), a file listing the names of values within those expressions, and a file listing all variables with which to take partial derivatives of each expression. The output is a text file containing the original expression and those for each partial derivative computation. The intention is to perform CSE on these expressions to create efficient partial derivative code!
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Initialize core Python/NRPy+ modules
# 1. [Step 2:](#read_expressions) Read in Hamiltonian expressions from `Hamstring.txt`
# 1. [Step 3:](#list_constants) Specify constants and variables in Hamiltonian expression
# 1. [Step 4:](#list_free_symbols) Extract free symbols
# 1. [Step 5:](#convert_to_func) Convert variables to function notation; e.g., `var` goes to `var(xx)`
# 1. [Step 6:](#differentiate) Differentiate with respect to `xx`
# 1. [Step 7:](#remove_zeros) Remove derivatives (of constants) that evaluate to zero, simplifying derivative expressions
# 1. [Step 8:](#partial_derivative) Simplify derivatives with respect to a specific variable
# 1. [Step 9:](#store_results) Store partial derivatives to SymPy notebook `partial_derivatives.txt-VALIDATION.txt`
# 1. [Step 10:](#code_validation) Validate against LALSuite and trusted `SEOBNR_Derivative_Routine` NRPy+ module
# 1. [Step 11:](#latex_pdf_output) Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Initialize core Python/NRPy+ modules \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
# Let's start by importing all the needed modules from Python/NRPy+ and creating the output directory (if it does not already exist):
# +
# Step 1.a: import all needed modules from Python/NRPy+:
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import sys, os # Standard Python modules for multiplatform OS-level functions
from outputC import superfast_uniq, lhrh # Remove duplicate entries from a Python array; store left- and right-
# hand sides of mathematical expressions
# Step 1.b: Check for a sufficiently new version of SymPy (for validation)
# Ignore the rc's and b's for release candidates & betas.
sympy_version = sp.__version__.replace('rc', '...').replace('b', '...')
sympy_version_decimal = float(sympy_version.split(".")[0]) + float(sympy_version.split(".")[1])/10.0
if sympy_version_decimal < 1.2:
print('Error: NRPy+ does not support SymPy < 1.2')
sys.exit(1)
# Step 1.c: Name of the directory containing the input file
inputdir = "SEOBNR"
# -
# <a id='read_expressions'></a>
#
# # Step 2: Read in Hamiltonian expressions from `Hamstring.txt` \[Back to [top](#toc)\]
# $$\label{read_expressions}$$
#
# We read in the expressions of which we will compute partial derivatives in a single large string before splitting the string by line (carriage return) and by "=". Doing so allows us to manipulate the right- and left-hand sides of the expressions appropriately. We store the left- and right-hand sides in the array `lr`, which consists of `lhrh` arrays with left-hand sides `lhs` and right-hand sides `rhs`. Note that `Lambda` is a protected keyword in Python, so the variable $\Lambda$ in the Hamiltonian is renamed `Lamb`.
# +
# Step 2.a: Read in expressions as a (single) string
with open(os.path.join(inputdir,'Hamstring.txt'), 'r') as file:
expressions_as_lines = file.readlines()
# Step 2.b: Create and populate the "lr" array, which separates each line into left- and right-hand sides
# Each entry is a string of the form lhrh(lhs='',rhs='')
lr = []
for i in range(len(expressions_as_lines)):
# Ignore lines with 2 or fewer characters and those starting with #
if len(expressions_as_lines[i]) > 2 and expressions_as_lines[i][0] != "#":
# Split each line by its equals sign
split_line = expressions_as_lines[i].split("=")
# Append the line to "lr", removing spaces, "sp." prefixes, and replacing Lambda->Lamb
# (Lambda is a protected keyword):
lr.append(lhrh(lhs=split_line[0].replace(" ","").replace("Lambda","Lamb"),
rhs=split_line[1].replace(" ","").replace("sp.","").replace("Lambda","Lamb")))
# Step 2.c: Separate and sympify right- and left-hand sides into separate arrays
lhss = []
rhss = []
# As of April 2021, "sp.sympify("Q+1")" fails because Q is a reserved keyword.
# This is the workaround, courtesy <NAME>.
custom_global_dict = {}
exec('from sympy import *', custom_global_dict)
del custom_global_dict['Q']
if sympy_version_decimal >= 1.6:
custom_parse_expr = lambda expr: sp.parse_expr(expr, global_dict=custom_global_dict)
else:
custom_parse_expr = lambda expr: sp.sympify(expr)
for i in range(len(lr)):
lhss.append(custom_parse_expr(lr[i].lhs))
rhss.append(custom_parse_expr(lr[i].rhs))
# -
# <a id='list_constants'></a>
#
# # Step 3: Specify constants and variables in Hamiltonian expression \[Back to [top](#toc)\]
# $$\label{list_constants}$$
#
# We read in and declare as SymPy symbols the constant values; derivatives with respect to these variables will be set to zero. We then read in the variables with respect to which we want to take derivatives and declare those as SymPy variables as well.
# +
# Step 3.a: Create `input_constants` array and populate with SymPy symbols
m1,m2,tortoise,eta,KK,k0,k1,EMgamma,d1v2,dheffSSv2 = sp.symbols('m1 m2 tortoise eta KK k0 k1 EMgamma d1v2 dheffSSv2',
real=True)
input_constants = [m1,m2,tortoise,eta,KK,k0,k1,EMgamma,d1v2,dheffSSv2]
# Step 3.b: Create `dynamic_variables` array and populate with SymPy symbols
x,y,z,px,py,pz,s1x,s1y,s1z,s2x,s2y,s2z = sp.symbols('x y z px py pz s1x s1y s1z s2x s2y s2z', real=True)
dynamic_variables = [x,y,z,px,py,pz,s1x,s1y,s1z,s2x,s2y,s2z]
# -
# <a id='list_free_symbols'></a>
#
# # Step 4: Extract free symbols \[Back to [top](#toc)\]
# $$\label{list_free_symbols}$$
#
# By ''free symbols'' we mean the variables in the right-hand sides. We first create a list of all such terms (using SymPy's built-in free_symbol attribute), including duplicates, and then strip the duplicates. We then remove input constants from the symbol list.
# +
# Step 4.a: Prepare array of "free symbols" in the right-hand side expressions
full_symbol_list_with_dups = []
for i in range(len(lr)):
for variable in rhss[i].free_symbols:
full_symbol_list_with_dups.append(variable)
# Step 4.b: Remove duplicate free symbols
full_symbol_list = superfast_uniq(full_symbol_list_with_dups)
# Step 4.c: Remove input constants from symbol list
for inputconst in input_constants:
for symbol in full_symbol_list:
if str(symbol) == str(inputconst):
full_symbol_list.remove(symbol)
# -
# <a id='convert_to_func'></a>
#
# # Step 5: Convert variables to function notation; e.g., `var` goes to `var(xx)` \[Back to [top](#toc)\]
# $$\label{convert_to_func}$$
#
# In order to compute the partial derivative of each right-hand side, we mark each variable (left-hand side) and each free symbol (in right-hand sides) as a function with argument $\texttt{xx}$.
# +
# Step 5.a: Convert each left-hand side to function notation
# while separating and simplifying left- and right-hand sides
xx = sp.Symbol('xx',real=True)
func = []
for i in range(len(lr)):
func.append(sp.sympify(sp.Function(lr[i].lhs,real=True)(xx)))
# Step 5.b: Mark each free variable as a function with argument xx
full_function_list = []
for symb in full_symbol_list:
func = sp.sympify(sp.Function(str(symb),real=True)(xx))
full_function_list.append(func)
for i in range(len(rhss)):
for var in rhss[i].free_symbols:
if str(var) == str(symb):
rhss[i] = rhss[i].subs(var,func)
# -
# <a id='differentiate'></a>
#
# # Step 6: Differentiate with respect to `xx` \[Back to [top](#toc)\]
# $$\label{differentiate}$$
#
# Now we differentiate the right-hand expressions with respect to `xx`. We use the SymPy $\texttt{diff}$ command, differentiating with respect to $\texttt{xx}$. After so doing, we remove $\texttt{(xx)}$ and "Derivative" (which is output by $\texttt{diff}$), and use "prm" suffix to denote the derivative with respect to $\texttt{xx}$.
# Step 6: Use SymPy's diff function to differentiate right-hand sides with respect to xx
# and append "prm" notation to left-hand sides
lhss_deriv = []
rhss_deriv = []
for i in range(len(rhss)):
lhss_deriv.append(custom_parse_expr(str(lhss[i])+"prm"))
newrhs = custom_parse_expr(str(sp.diff(rhss[i],xx)).replace("(xx)","").replace(", xx","prm").replace("Derivative",""))
rhss_deriv.append(newrhs)
# <a id='remove_zeros'></a>
#
# # Step 7: Remove derivatives (of constants) that evaluate to zero, simplifying derivative expressions \[Back to [top](#toc)\]
# $$\label{remove_zeros}$$
#
# We declare a function to simply the derivative expressions. In particular, we want to remove terms equal to zero.
# +
# Step 7.a: Define derivative simplification function
def simplify_deriv(lhss_deriv,rhss_deriv):
# Copy expressions into another array
lhss_deriv_simp = []
rhss_deriv_simp = []
for i in range(len(rhss_deriv)):
lhss_deriv_simp.append(lhss_deriv[i])
rhss_deriv_simp.append(rhss_deriv[i])
# If a right-hand side is 0, substitute value 0 for the corresponding left-hand side in later terms
for i in range(len(rhss_deriv_simp)):
if rhss_deriv_simp[i] == 0:
for j in range(i+1,len(rhss_deriv_simp)):
for var in rhss_deriv_simp[j].free_symbols:
if str(var) == str(lhss_deriv_simp[i]):
rhss_deriv_simp[j] = rhss_deriv_simp[j].subs(var,0)
zero_elements_to_remove = []
# Create array of indices for expressions that are zero
for i in range(len(rhss_deriv_simp)):
if rhss_deriv_simp[i] == sp.sympify(0):
zero_elements_to_remove.append(i)
# When removing terms that are zero, we need to take into account their new index (after each removal)
count = 0
for i in range(len(zero_elements_to_remove)):
del lhss_deriv_simp[zero_elements_to_remove[i]+count]
del rhss_deriv_simp[zero_elements_to_remove[i]+count]
count -= 1
return lhss_deriv_simp,rhss_deriv_simp
# Step 7.b: Call the simplication function and then copy results
lhss_deriv_simp,rhss_deriv_simp = simplify_deriv(lhss_deriv,rhss_deriv)
lhss_deriv = lhss_deriv_simp
rhss_deriv = rhss_deriv_simp
# -
# <a id='partial_derivative'></a>
#
# # Step 8: Simplify derivatives with respect to a specific variable \[Back to [top](#toc)\]
# $$\label{partial_derivative}$$
#
# In [Step 6](#differentiate) we took a generic derivative of each expression, assuming all variables were functions of `xx`. We now define a function that will select a specific dynamic variable (element of `dynamic_variables`) and set the derivative of the variable to 1 and all others to 0.
# +
# Step 8.a: Define onevar derivative function
def deriv_onevar(lhss_deriv,rhss_deriv,variable_list,index):
# Denote each variable with prm
variableprm_list = []
for variable in variable_list:
variableprm_list.append(str(variable)+"prm")
# Copy expressions into another array
lhss_deriv_new = []
rhss_deriv_new = []
for i in range(len(rhss_deriv)):
lhss_deriv_new.append(lhss_deriv[i])
rhss_deriv_new.append(rhss_deriv[i])
# For each free symbol's derivative, replace it with:
# 1, if we are differentiating with respect to the variable, or
# 0, if we are note differentiating with respect to that variable
for i in range(len(rhss_deriv_new)):
for var in variableprm_list:
if variableprm_list.index(str(var))==index:
rhss_deriv_new[i] = rhss_deriv_new[i].subs(var,1)
else:
rhss_deriv_new[i] = rhss_deriv_new[i].subs(var,0)
# Simplify derivative expressions again
lhss_deriv_simp,rhss_deriv_simp = simplify_deriv(lhss_deriv_new,rhss_deriv_new)
return lhss_deriv_simp,rhss_deriv_simp
# Step 8.b: Call the derivative function and populate dictionaries with the result
lhss_derivative = {}
rhss_derivative = {}
for index in range(len(dynamic_variables)):
lhss_temp,rhss_temp = deriv_onevar(lhss_deriv,rhss_deriv,dynamic_variables,index)
lhss_derivative[dynamic_variables[index]] = lhss_temp
rhss_derivative[dynamic_variables[index]] = rhss_temp
# -
# <a id='store_results'></a>
#
# # Step 9: Store partial derivatives to SymPy notebook `partial_derivatives.txt-VALIDATION.txt` \[Back to [top](#toc)\]
# $$\label{store_results}$$
#
# We write the resulting derivatives in SymPy syntax. Each partial derivative is output in its own file, in a similar format to the input expressions.
# Step 9: Output original expression and each partial derivative expression in SymPy snytax
with open(os.path.join(inputdir,'partial_derivatives.txt-VALIDATION'), 'w') as output:
for i in range(len(lr)):
right_side = lr[i].rhs
right_side_in_sp = right_side.replace("sqrt(","sp.sqrt(").replace("log(","sp.log(").replace("pi",
"sp.pi").replace("sign(","sp.sign(").replace("Abs(",
"sp.Abs(").replace("Rational(","sp.Rational(")
output.write(str(lr[i].lhs)+" = "+right_side_in_sp)
for var in dynamic_variables:
for i in range(len(lhss_derivative[var])):
right_side = str(rhss_derivative[var][i])
right_side_in_sp = right_side.replace("sqrt(","sp.sqrt(").replace("log(","sp.log(").replace("pi",
"sp.pi").replace("sign(","sp.sign(").replace("Abs(",
"sp.Abs(").replace("Rational(","sp.Rational(").replace("prm",
"prm_"+str(var))
output.write(str(lhss_derivative[var][i]).replace("prm","prm_"+str(var))+" = "+right_side_in_sp+"\n")
# <a id='code_validation'></a>
#
# # Step 10: Validate against LALSuite and trusted `SEOBNR_Derivative_Routine` NRPy+ module \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# We validate the output of this notebook against known LALSuite values of the Hamiltonian partial derivatives and the output of the `SEOBNR_Derivative_Routine` NRPy+ module. We note that due to cancellations in the deriavtive terms, various versions of SymPy may result in relative errors that differ as much as an order of magnitude. Furthermore, even changing the set of input pararameters can affect the relative error by as many as two orders of magnitude. Therefore we look for agreement with LALSuite to at least 10 significant digits.
#
# When comparing the notebook output to that of the NRPy+ module, we compare term-by-term using SymPy to check that each right-hand side side is equivalent.
# +
# Define a function to return a set of reasonable input parameters
# This function contains three distinct sets of input parameters, and index differentiates between them
def reset_values(tort_value, index):
# Check that a reasonable tortoise value has been passed
if tort_value!=1 and tort_value!=2:
print("Error: tortoise must be 1 or 2.")
sys.exit(1)
# Each index corresponds to a specific set of input parameters
if index==0:#-M 13 -m 11 -X 0.1 -Y -0.2 -Z 0.3 -x -0.3 -y 0.2 -z -0.1
values = {'m1': 1.300000000000000e+01, 'm2': 1.100000000000000e+01, 'eta': 2.482638888888889e-01,
'x': 1.658426645098320e+01, 'y': 3.975021008701605e-02, 'z': -1.820682538442627e-07,
's1x': 2.934751675254397e-02, 's1y': -5.867672205485316e-02, 's1z': 8.802097562761332e-02,
's2x': -6.302678133897792e-02, 's2y': 4.200490780215727e-02, 's2z': -2.100705983874398e-02,
'KK': 3.913980338468737e-01, 'k0': -7.447639215330089e-01, 'k1': -6.380586501824999e-01,
'd1v2': -7.476323019145448e+01,'dheffSSv2':2.105103187692902e+01,
'EMgamma': 0.577215664901532860606512090082402431}
# Note that we transform the momentum based on the tortoise values
if tort_value==1:
values.update({'px': -1.517631642228534e-03, 'py': 2.693180445886167e-01, 'pz': -1.320499830947482e-04,
'tortoise': 1})
else:
values.update({'px': -1.633028076483384e-03, 'py': 2.693177679992048e-01, 'pz': -1.320499918278832e-04,
'tortoise': 2})
elif index==1:#-M 25 -m 10 -X 0.1 -Y -0.0 -Z 0.1 -x -0.2 -y 0.0 -z -0.2
values = {'m1': 2.500000000000000e+01, 'm2': 1.000000000000000e+01, 'eta': 2.040816326530612e-01,
'x': 1.289689003662444e+01, 'y': 5.495441315063273e-03, 'z': -1.717482806041791e-11,
's1x': 5.102040816179230e-02, 's1y': 9.846215537206260e-07, 's1z': 5.102040816473832e-02,
's2x': -1.632653061189792e-02, 's2y': -6.762952223804450e-07, 's2z': -1.632653061259188e-02,
'KK': 5.642540639599580e-01, 'k0': -1.063532077165767e+00, 'k1': -8.835684149841774e-01,
'd1v2': -8.041179092044979e+01,'dheffSSv2':1.125986130778842e+01,
'EMgamma': 0.577215664901532860606512090082402431}
if tort_value==1:
values.update({'px': -1.898773926867491e-03, 'py': 3.160984442121970e-01, 'pz': 1.171602901570564e-07,
'tortoise': 1})
else:
values.update({'px': -2.209215477700561e-03, 'py': 3.160983119312114e-01, 'pz': 1.171602905704723e-07,
'tortoise': 2})
elif index==2:#-M 7 -m 5 -X 0.01 -Y -0.5 -Z 0.03 -x -0.04 -y 0.05 -z -0.06
values = {'m1': 7.000000000000000e+00, 'm2': 5.000000000000000e+00, 'eta': 2.430555555555556e-01,
'x': 2.633506161699224e+01, 'y': 7.574563213724998e-02, 'z': -2.789625823248071e-08,
's1x': 3.417297286269225e-03, 's1y': -1.701385963191495e-01, 's1z': 1.020835932957879e-02,
's2x': -6.945454346305877e-03, 's2y': 8.679766617922793e-03, 's2z': -1.041665076794264e-02,
'KK': 4.052853693162246e-01, 'k0': -7.706473492549312e-01, 'k1': -6.587426366263742e-01,
'd1v2': -7.555647472993827e+01,'dheffSSv2':1.972817669753086e+01,
'EMgamma': 0.577215664901532860606512090082402431}
if tort_value==1:
values.update({'px': -7.883793607066706e-04, 'py': 2.068742709904638e-01, 'pz': -7.338789145500886e-04,
'tortoise': 1})
else:
values.update({'px': -8.039726989861640e-04, 'py': 2.068742261404732e-01, 'pz': -7.338789145335709e-04,
'tortoise': 2})
else:
# If an improper index is passed, exit
print("Error: invalid index (only three sets of input parameters available).")
sys.exit(1)
# Return the input values
return values
# Numerically evaluate right-hand sides using input values
def evaluate_expression(left_sides,right_sides,input_values):
new_right_sides = []
for i in range(len(right_sides)):
term = custom_parse_expr(str(right_sides[i]).replace("(xx)",""))
# Only look for the free variables in each expression to reduce computation time
free_vars = term.free_symbols
for variable in free_vars:
term = term.subs(variable, input_values[str(variable)])
# Evaluate each term to reduce computation time
new_right_sides.append(sp.sympify(term.evalf()))
# Store each subexpression in values numerically
input_values[str(left_sides[i])] = new_right_sides[i]
# Return the input values dictionary with all numerical right-hand added
return input_values
# Create array of trusted LALSuite derivative values
# Note that position in the array corresponds to the index of the corresponding input values
LALSuite_validated_values = []
#-M 13 -m 11 -X 0.1 -Y -0.2 -Z 0.3 -x -0.3 -y 0.2 -z -0.1
LALSuite_validated_values.append({'Hreal': 9.928923110195770e-01,'dHreal_dx': 9.932484846748471e-04,
'dHreal_dy': 2.813294366789505e-06, 'dHreal_dz': 1.926378549762488e-06,
'dHreal_dpx': -3.710666135737856e-04, 'dHreal_dpy': 6.116199124763537e-02,
'dHreal_dpz': -5.600910364542288e-07, 'dHreal_ds1x': -1.438467658934620e-05,
'dHreal_ds1y': -1.319462868057848e-06, 'dHreal_ds1z': 7.665413183773232e-04,
'dHreal_ds2x': -2.075691477548065e-05,'dHreal_ds2y': 2.456427688083135e-06,
'dHreal_ds2z': 8.762835349889455e-04})
#-M 25 -m 10 -X 0.1 -Y -0.0 -Z 0.1 -x -0.2 -y 0.0 -z -0.2
LALSuite_validated_values.append({'Hreal': 9.926852598351464e-01, 'dHreal_dx': 1.397519118422771e-03,
'dHreal_dy': 1.928133240540033e-06, 'dHreal_dz': -1.215449398950413e-06,
'dHreal_dpx': -4.004159849919695e-04, 'dHreal_dpy': 5.701850933742150e-02,
'dHreal_dpz': 4.329487960716782e-08, 'dHreal_ds1x': 2.259457049322466e-06,
'dHreal_ds1y': -2.544122765762015e-09, 'dHreal_ds1z': 9.834156257814124e-04,
'dHreal_ds2x': 5.185557993931246e-06,'dHreal_ds2y': 2.437768415468806e-10,
'dHreal_ds2z': 2.111169766641698e-03})
#-M 7 -m 5 -X 0.01 -Y -0.5 -Z 0.03 -x -0.04 -y 0.05 -z -0.06
LALSuite_validated_values.append({'Hreal': 9.955293642650920e-01, 'dHreal_dx': 3.734697245297603e-04,
'dHreal_dy': 1.105998063449349e-06, 'dHreal_dz': 5.367207414282669e-08,
'dHreal_dpx': -1.848412708548443e-04, 'dHreal_dpy': 4.754239153769983e-02,
'dHreal_dpz': -3.549083643069269e-08, 'dHreal_ds1x': -4.819261725948465e-07,
'dHreal_ds1y': 3.333280059627902e-06, 'dHreal_ds1z': 2.201786563823208e-04,
'dHreal_ds2x': -7.576810957551029e-07,'dHreal_ds2y': 6.818093508597533e-06,
'dHreal_ds2z': 2.922663340179887e-04})
# Sort variables by which tortoise value we use to compute the derivatives
variables_tort2 = [x,y,z]
variables_tort1 = [px,py,pz,s1x,s1y,s1z,s2x,s2y,s2z]
# Call evaluation function
print("Computing the difference between notebook output and trusted LALSuite derivative values...")
for index in range(3):
values = reset_values(2,index)
values = evaluate_expression(lhss,rhss,values)
Hreal = values['Hreal']
trusted_Hreal = LALSuite_validated_values[index]['Hreal']
relative_difference = (trusted_Hreal - Hreal)/trusted_Hreal
if abs(relative_difference) > 1e-9:
print("The relative difference in Hreal is too large: %.15e" % relative_difference)
sys.exit(1)
for var in variables_tort2:
Hrealprm = evaluate_expression(lhss_derivative[var],rhss_derivative[var],values)['Hrealprm']
trusted_Hrealprm = LALSuite_validated_values[index]['dHreal_d'+str(var)]
relative_difference = (trusted_Hrealprm - Hrealprm)/trusted_Hrealprm
if abs(relative_difference) > 1e-9:
print("The relative difference in dHreal_d%s is too large: %.15e" % (var,relative_difference))
sys.exit(1)
values = reset_values(1,index)
values = evaluate_expression(lhss,rhss,values)
for var in variables_tort1:
Hrealprm = evaluate_expression(lhss_derivative[var],rhss_derivative[var],values)['Hrealprm']
trusted_Hrealprm = LALSuite_validated_values[index]['dHreal_d'+str(var)]
relative_difference = (trusted_Hrealprm - Hrealprm)/trusted_Hrealprm
if abs(relative_difference) > 1e-9:
print("The relative difference in dHreal_d%s is too large: %.15e" % (var,relative_difference))
sys.exit(1)
print("Test passed: the notebook agrees with LALSuite to at least 10 significant digits!")
print("Printing difference between notebook output and trusted NRPy+ module output...")
# Open the files to compare
file = 'partial_derivatives.txt'
outfile = 'partial_derivatives.txt-VALIDATION'
print("Checking file " + outfile)
with open(os.path.join(inputdir,file), "r") as file1, open(os.path.join(inputdir,outfile), "r") as file2:
# Read the lines of each file
file1_lines = file1.readlines()
file2_lines = file2.readlines()
# Compare right-hand sides of the expressions by computing the difference between them
num_diffs = 0
for i in range(len(file1_lines)):
expr_new = custom_parse_expr(file1_lines[i].split("=")[1].replace("sp.",""))
expr_validated = custom_parse_expr(file2_lines[i].split("=")[1].replace("sp.",""))
difference = sp.simplify(expr_new - expr_validated)
if difference != 0:
num_diffs += 1
print(difference)
if num_diffs == 0:
print("No difference. TEST PASSED!")
else:
print("ERROR: Disagreement found with the trusted file. See differences above.")
sys.exit(1)
# -
# <a id='latex_pdf_output'></a>
#
# # Step 11: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-SEOBNR_Derivative_Routine.pdf](Tutorial-SEOBNR_Derivative_Routine.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-SEOBNR_Derivative_Routine")
| Tutorial-SEOBNR_Derivative_Routine.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: F#
// language: fsharp
// name: ifsharp
// ---
// (*** hide ***)
// +
#load "../../bin/net45/Deedle.fsx"
open System
// -
// (**
// Creating lazily loaded series
// =============================
//
// When loading data from an external data source (such as a database), you might
// want to create a _virtual_ time series that represents the data source, but
// does not actually load the data until needed. If you apply some range restriction
// (like slicing) to the data series before using the values, then it is not
// necessary to load the entire data set into memory.
//
// Deedle supports lazy loading through the `DelayedSeries.FromValueLoader`
// method. It returns an ordinary data series of type `Series<K, V>` which has a
// delayed internal representation.
//
// ## Creating lazy series
//
// We will not use a real database in this tutorial, but let's say that you have the
// following function which loads data for a given day range:
// *)
// +
open Deedle
/// Given a time range, generates random values for dates (at 12:00 AM)
/// starting with the day of the first date time and ending with the
/// day after the second date time (to make sure they are in range)
let generate (low:DateTime) (high:DateTime) =
let rnd = Random()
let days = int (high.Date - low.Date).TotalDays + 1
seq { for d in 0 .. days ->
KeyValue.Create(low.Date.AddDays(float d), rnd.Next()) }
// -
// (**
// Using random numbers as the source in this example is not entirely correct, because
// it means that we will get different values each time a new sub-range of the series
// is required - but it will suffice for the demonstration.
//
// Now, to create a lazily loaded series, we need to open the `Indices` namespace,
// specify the minimal and maximal value of the series and use `DelayedSeries.FromValueLoader`:
// *)
// +
open Deedle.Indices
// Minimal and maximal values that can be loaded from the series
let min, max = DateTime(2010, 1, 1), DateTime(2013, 1, 1)
// Create a lazy series for the given range
let ls = DelayedSeries.FromValueLoader(min, max, fun (lo, lob) (hi, hib) -> async {
printfn "Query: %A - %A" (lo, lob) (hi, hib)
return generate lo hi })
// -
// (**
// To make the diagnostics easier, we print the required range whenever a request
// is made. After running this code, you should not see any output yet.
// The parameter to `DelayedSeries.FromValueLoader` is a function that takes 4 arguments:
//
// - `lo` and `hi` specify the low and high boundaries of the range. Their
// type is the type of the key (e.g. `DateTime` in our example)
// - `lob` and `hib` are values of type `BoundaryBehavior` and can be either
// `Inclusive` or `Exclusive`. They specify whether the boundary value should
// be included or not.
//
// Our sample function does not handle boundaries correctly - it always includes the
// boundary (and possibly more values). This is not a problem, because the lazy loader
// automatically skips over such values. But if you want, you can use `lob` and `hib`
// parameters to build a more optimal SQL query.
//
// ## Using un-evaluated series
//
// Let's now have a look at the operations that we can perform on un-evaluated series.
// Any operation that actually accesses values or keys of the series (such as `Series.observations`
// or lookup for a specific key) will force the evaluation of the series.
//
// However, we can use range restrictions before accessing the data:
// *)
// +
// Get series representing January 2012
let jan12 = ls.[DateTime(2012, 1, 1) .. DateTime(2012, 2, 1)]
// Further restriction - only first half of the month
let janHalf = jan12.[.. DateTime(2012, 1, 15)]
// Get value for a specific date
janHalf.[DateTime(2012, 1, 1)]
// [fsi: Query: (1/1/2012, Inclusive) - (1/15/2012, Inclusive)]
// [fsi: val it : int = 1127670994]
janHalf.[DateTime(2012, 1, 2)]
// [fsi: val it : int = 560920727]
// -
// (**
// As you can see from the output on line 9, the series obtained data for the
// 15 day range that we created by restricting the original series. When we requested
// another value within the specified range, it was already available and it was
// returned immediately. Note that `janHalf` is restricted to the specified 15 day
// range, so we cannot access values outside of the range. Also, when you access a single
// value, entire series is loaded. The motivation is that you probably need to access
// multiple values, so it is likely cheaper to load the whole series.
//
// Another operation that can be performed on an unevaluated series is to add it
// to a data frame with some existing key range:
// *)
// +
// Create empty data frame for days of December 2011
let dec11 = Frame.ofRowKeys [ for d in 1 .. 31 -> DateTime(2011, 12, d) ]
// Add series as the 'Values' column to the data frame
dec11?Values <- ls
// [fsi: Query: (12/1/2011, Inclusive) - (12/31/2011, Inclusive)]
// -
// (**
// When adding lazy series to a data frame, the series has to be evaluated (so that
// the values can be properly aligned) but it is first restricted to the range of the
// data frame. In the above example, only one month of data is loaded.
//
// *)
| notebooks/other/Deedle/lazysource.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS" tags=[]
# # Script format of function calls
# + [markdown] kernel="SoS" tags=[]
# * **Difficulty level**: easy
# * **Time need to lean**: 10 minutes or less
# * **Key points**:
# * A Python function with a string as the first parameter can be written in a script style
# * Scripts are dedented, and included in verbatim by default
# * `expand=True` turns the script into a Python f-string
# * `expand="l r"` can be used to specify an alternative delimiter for string interpolation
# + [markdown] kernel="SoS" tags=[]
# ## SoS "actions"
# + [markdown] kernel="SoS" tags=[]
# SoS defines a number of **actions**, which are simply Python functions that follow a specific set of conventions. For example, an action `sh` is a python function that executes its first parameter as a shell script:
# + kernel="SoS" tags=[]
sh('echo Hello world')
# + [markdown] kernel="SoS" tags=[]
# When you execute a SoS script from command line or SoS Notebook, these functions are automatically imported and can be used directly, and you can use them just like any other Python functions. For example, you can compose a script using Python string formatting:
# + kernel="SoS" tags=[]
greeting = "Hello world"
sh(f'echo {greeting}')
# + [markdown] kernel="SoS" tags=[]
# Here we define a Python string `greeting` and use the `sh` action to execute a shell script. A [Python f-string](https://www.python.org/dev/peps/pep-0498/) is used to compose the script with the defined variable.
# + [markdown] kernel="SoS" tags=[]
# <div class="bs-callout bs-callout-primary" role="alert">
# <h4>Python f-string</h4>
# <p>SoS uses Python f-string extensively. Please read <a href="https://www.python.org/dev/peps/pep-0498/">PEP498</a> or one of the online tutorials on how to use Python f-strings if you are not familiar with it.</p>
# </div>
# + [markdown] kernel="SoS" tags=[] workflow_cell=true
# When the scripts get longer, you can use Python multi-line strings to pass them to the action. Here is an example for the use of a `R` action to execute a longer `R` script:
# + kernel="SoS" tags=[]
R("""\
x <- 5
if(x >= 0) {
print("Non-negative number")
} else {
print("Negative number")
}
""")
# + [markdown] kernel="SoS" tags=[]
# The use of format string in these cases become more complicated. First, you will need to use multi-line f-string (`f'''x'''` and `f"""x"""`). Second, when the script itself contains braces, they will need to be doubled to avoid being interpreted as Python expressions.
#
# Consequently, a `R` script that uses a Python variable `my_num` needs to be written as follows:
# + kernel="SoS" tags=[]
my_num = -1
R(f"""\
x <- {my_num}
if(x >= 0) {{
print("Non-negative number")
}} else {{
print("Negative number")
}}
""")
# + [markdown] kernel="SoS" tags=[]
# ## Script style function calls <a id="Script_style_function_call"></a>
# + [markdown] kernel="SoS" tags=[]
# The f-string in the last example is not quite readable, error-prone, and difficult to maintain, especially when the script contains multiple braces and variables. For this reason, SoS introduces a special syntax that allows you to write Python functions that accept a script (string) as the first parameter in a special script format.
#
# For example,
# + kernel="SoS" tags=[]
R("""\
x <- 5
if(x >= 0) {
print("Non-negative number")
} else {
print("Negative number")
}
""")
# + [markdown] kernel="SoS" tags=[]
# can be written as
# + kernel="SoS" tags=[]
R:
x <- 5
if(x >= 0) {
print("Non-negative number")
} else {
print("Negative number")
}
# + [markdown] kernel="SoS" tags=[]
# but as the warning message shows, better included with indentation:
# + kernel="SoS" tags=[]
R:
x <- 5
if(x >= 0) {
print("Non-negative number")
} else {
print("Negative number")
}
# + [markdown] kernel="SoS" tags=[]
# <div class="bs-callout bs-callout-info" role="alert">
# <h4>Indentation of scripts</h4>
# <p>The indentation of scripts in the script style is optional but highly recommended because it makes the scripts much easier to identify and read </p>
# <pre>
# R:
# cat('this is R')
# </pre>
# <pre>
# python:
# print('this is python')
# </pre>
# </div>
# + [markdown] kernel="SoS" tags=[]
# ## <a id="option-expand"></a> The `expand` option
# + [markdown] kernel="SoS" tags=[]
# When option `expand=True` is specified, the included script will be treated as a Python f-string. For example,
# + kernel="SoS" tags=[]
my_num = -1
R(f"""\
x <- {my_num}
if(x >= 0) {{
print("Non-negative number")
}} else {{
print("Negative number")
}}
""")
# + [markdown] kernel="SoS" tags=[]
# can be written as
# + kernel="SoS" tags=[]
my_num = -1
R: expand=True
x <- {my_num}
if(x >= 0) {{
print("Non-negative number")
}} else {{
print("Negative number")
}}
# + [markdown] kernel="SoS" tags=[]
# Note that SoS Notebook automatically highlights the interpolated parts of the included script, which makes it much easier to differentiate Python expressions from the original R script.
# + [markdown] kernel="SoS" tags=[]
# <div class="bs-callout bs-callout-info" role="alert">
# <h4>Use of alternative sigils</h4>
# <p>When the included scripts have braces, it is easier to use an alternative sigil for string interpolation.</p>
# </div>
# + [markdown] kernel="SoS" tags=[]
# Because the included script has two pairs of braces, it is necessary to double them so that they are not treated as Python expressions. In these cases, it is actually easier to use a different set of sigil (delimiters) for string interpolation. This can be done using the `expand` option as follows:
# + kernel="SoS" tags=[]
my_num = -1
R: expand='${ }'
x <- ${my_num}
if(x >= 0) {
print("Non-negative number")
} else {
print("Negative number")
}
# + [markdown] kernel="SoS" tags=[]
# The sigil should be specified as a string with left and right sigil separated by a space. You can use any pair of sigils as long as they do not cause confusion.
# + [markdown] kernel="SoS" tags=[]
# ## Indented script format
# + [markdown] kernel="SoS" tags=[]
# What you have seen so far are function calls without Python control structure. Because SoS allows you to execute the same statements repeatedly for different inputs ([substeps](input_substeps.html) and stop the execution of substeps through actions such as [`skip_if` and `done_if`](control_actions.html), there is rarely a need to call SoS actions within the `if...else...` block or `for` or `while` loops.
#
# However, if you do need to call SoS actions inside a Python control structure, you can still use the script format for it. For example, instead of writing
# + kernel="SoS" tags=[]
for i in range(3):
sh(f'echo {i}')
# + [markdown] kernel="SoS" tags=[]
# You can write the `sh` action in the format of
# + kernel="SoS" tags=[]
for i in range(3):
sh: expand=True
echo {i}
# + [markdown] kernel="SoS" tags=[]
# or in more complex case
# + kernel="SoS" tags=[]
# %run --var 2 -v1
parameter: var=int
if var > 1:
python:
print('Python script for var > 1')
else:
python:
print('Python script for var <= 1')
| src/user_guide/script_format.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: initPython383
# language: python
# name: initpython383
# ---
# <img align="left" style="padding-right:10px;" width="150" src="https://upload.wikimedia.org/wikipedia/commons/thumb/6/6c/Star_Wars_Logo.svg/320px-Star_Wars_Logo.svg.png" />
#
# *elaborado por <NAME>.*
# < [Colecciones de objetos: listas, tuplas y diccionarios](modulo1_tema4_Py_30_colec_obj.ipynb) | [Índice](modulo1_tema4_Py_00_indice.ipynb) | [Colecciones de objetos: pandas](modulo1_tema4_Py_32_pandas.ipynb) >
#
# __[Abre en Colab](https://colab.research.google.com/github/griu/init_python_b1/blob/master/modulo1_tema4_Py_31_numpy.ipynb)__ *: <span style="color:rgba(255, 99, 71, 0.8)">Padawan! Cuando inicies sesión en Colab, prepara el entorno ejecutando el siguiente código.</span>*
# + [markdown] id="Dy11Y3WrHRHS"
# # Preparación del entorno
#
# ¡Padawan! Cuando inicies sesión en Colab, prepara el entorno ejecutando el siguiente código:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 25678, "status": "ok", "timestamp": 1604317087349, "user": {"displayName": "alumnos bigdata", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="2SBF5dlSTi-h" outputId="b85ff6ea-725c-464c-abf4-fc481a6882de"
if 'google.colab' in str(get_ipython()):
# !git clone https://github.com/griu/init_python_b1.git /content/init_python_b1
# !git -C /content/init_python_b1 pull
# %cd /content/init_python_b1
# + [markdown] id="F9n3xfvpF5df"
# # 5 - Numpy
#
# El package [numpy](https://numpy.org/) es la solución más popular dentro de Python para realizar computación científica.
#
# Recoge las mejores prácticas introducidas en las Listas y organizadas para realizar cálculos de forma eficiente. Se estructuran como vectores o arrays de N, dimensiones de un mismo tipo de dato.
#
# El primer paso que realizaremos será cargar la librería, habitualmente, con *namespace* `np`:
# + id="UAORIi9sF5do"
import numpy as np
# + [markdown] id="5KSnwU5BF5eG"
# ##### 5.1. Actividad Guiada 2.3
#
# Antes de entrar en materia, vamos a presentar la actividad guiada de *numpy*.
#
# Se trata de seleccionar la nave interestelar más rápida en MGLT (Megaluz/Hora), que pueda llevar a 4 tripulantes (como los protagonistas: Han Solo, Leia, Luke y Chewbacca). Además, el coste de la nave debe ser com máximo 1M de créditos interestelares.
#
# > «Es la chatarra más veloz de la galaxia.»
# ―<NAME>
#
# Lando se refería al <NAME>. ¿Será verdad?
#
# Para esta actividad contamos con los siguiente datos em formato pandas:
# + colab={"base_uri": "https://localhost:8080/", "height": 520} executionInfo={"elapsed": 1040, "status": "ok", "timestamp": 1604326572174, "user": {"displayName": "alumnos bigdata", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="BaU5fHBcF5eM" outputId="442bb833-a3a7-475d-c506-9767cad6b5db"
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn; seaborn.set() # para el estilo de graficos
entidades = ['planets','starships','vehicles','people','species']
entidades_df = {x: pd.read_pickle('www/' + x + '_df.pkl') for x in entidades}
starships_df = entidades_df['starships'][["cost_in_credits","MGLT","crew"]].dropna()
starships_df
# + [markdown] id="13fi6uN0F5es"
# Transformamos la matriz numpy con las variables numéricas:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 713, "status": "ok", "timestamp": 1604326575819, "user": {"displayName": "alumnos bigdata", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="fQiWePljF5ex" outputId="de4d0c6e-766e-4d98-f125-cf8da423123b"
starship_np = starships_df.values
starship_np # las filas son las naves y las columnas: cost_in_credits, MGLT y crew
# + [markdown] id="Q4fFdcarF5fM"
# Transformamos los nombres de las naves a array numpy:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 745, "status": "ok", "timestamp": 1604326578827, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="Q6z9DYktF5fU" outputId="f66aebc2-96a1-4dec-c89e-9a035e26ea31"
starship_names = starships_df.index.values
starship_names
# + [markdown] id="B15GpaTMF5gN"
# #### 5.2. Tipo de datos en Numpy
# Habitualmente, se crean a partir de listas o tuplas con elementos homogéneos:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 734, "status": "ok", "timestamp": 1604326581553, "user": {"displayName": "al<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="NsocN1GTF5gQ" outputId="4c132292-ffb9-4fab-b984-52700e69a9e2"
b_np = np.array([True, False, False]) # booleano
b_np
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 892, "status": "ok", "timestamp": 1604326583621, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="pOQLJO9OF5gl" outputId="cd73a83a-7683-4723-afed-ff44f66e2ad5"
s_np = np.array(['f', 'h', 'j']) # string
s_np
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 731, "status": "ok", "timestamp": 1604326585848, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="4VIR41zaF5g4" outputId="8eb45873-260f-4fb8-80af-17281c62bf5b"
i_np = np.array(range(3)) # entero
i_np
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 751, "status": "ok", "timestamp": 1604326587902, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="rjqaLwdVF5hQ" outputId="0959152b-204a-43da-b2f2-30cca64652d1"
f_np = np.array([2.4, 1.5, 3.0]) # coma flotante
f_np
# + [markdown] id="CWiR0JHnF5hm"
# Los 4 objetos comparten ser de tipo `numpy.ndarray`:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 796, "status": "ok", "timestamp": 1604326590333, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="AHDkYUyVF5hp" outputId="de59b063-95b5-417f-d4e8-7f234ed04b07"
ej_np = (b_np, s_np, i_np, f_np)
[type(x) for x in ej_np]
# + [markdown] id="YmD1yb3QF5h_"
# Sin embargo, cada uno tiene un tipo interno accesible con el método `.dtype`:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 803, "status": "ok", "timestamp": 1604326592802, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="nWGnAhHnF5iD" outputId="769a9027-81db-4f8c-cbc2-ccf94443cf27"
[x.dtype for x in ej_np]
# + [markdown] id="u8pW_uuBF5ig"
# Observa que numpy define sus propios nombres para los tipos básicos donde añade el tamaño en bits:
#
# - 'U1' es texto Unicode de 1 bit.
# - 'int64', 'float64' numéricos de 64 bits.
#
# Se puede definir este tipo en su construcción:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 847, "status": "ok", "timestamp": 1604326595682, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="oYg-U6sZF5ij" outputId="abba9382-2da6-479f-ccbf-0baf912b774e"
i_np = np.array(range(3), dtype='int16')
i_np
# + [markdown] id="gfrUpGgsF5i0"
# O convertir el numpy de un tipo interno a otro en un nuevo objeto con `.astype()`:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 798, "status": "ok", "timestamp": 1604326598341, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="pXsMA8j_F5i4" outputId="d6513e22-f6a0-4297-bec9-513f9fefab05"
s_i_np = i_np.astype(str)
s_i_np
# + [markdown] id="02bF7ixjF5jO"
# El tipo de la matriz numérica de las naves es:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1064, "status": "ok", "timestamp": 1604326600859, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="28pFlxnCF5jT" outputId="f42511f5-8487-4693-bdac-1266ed015d3e"
starship_np.dtype
# + [markdown] id="_6DtdXyCF5jj"
# El vector de nombres tiene dtype object. Este tipo se utiliza para numpy de texto:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1096, "status": "ok", "timestamp": 1604326603660, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="yK_B_kEdF5jm" outputId="4b978f7e-6792-4b41-afd5-67a2dadfa2a8"
starship_names.dtype
# + [markdown] id="I6NbQeQcF5j5"
# Así, se ve mejor:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 737, "status": "ok", "timestamp": 1604326606038, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="MgGEDHKJF5j7" outputId="fd754ac3-f120-4ebf-8d67-188cc39ece79"
print(starship_names.dtype)
# + [markdown] id="TuyYCGHgF5kP"
# #### 5.3. Funciones Universales
#
# En numpy, las funciones universales o *ufunc*, son un conjunto de funciones que vectorizan, es decir, que al aplicarlas sobre un objeto numpy, se aplica sobre cada uno de los elementos que lo componen.
#
# > **Recuerda**: Ya comentamos las funciones que vectorizan, en el capitulo de funciones de R.
#
# Típicamente, las funciones de este tipo son las funciones aritméticas:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 566, "status": "ok", "timestamp": 1604326609791, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="e9GJVCZTF5kU" outputId="4ef41cd6-1f2e-45a0-9ce0-a3a5e5e66af4"
np.array([1,2,3]) * 2 # multiplicación por escalar
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1130, "status": "ok", "timestamp": 1604326612662, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="sFYu2QbuF5kj" outputId="618af358-ccf5-4a8c-d854-903892879595"
np.array([1,2,3]) + np.array([4,5,6]) # suma vectores
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1009, "status": "ok", "timestamp": 1604326615330, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="9cxNT5sxF5k5" outputId="e19e418d-4a33-4c79-d04f-12b1be2e075f"
np.array([1,2,3]) * np.array([4,5,6]) # producto elemento a elemento
# + [markdown] id="XwEDsOQEF5lL"
# NumPy ofrece una extensa colección de funciones matemáticas que se aplican de forma vectorial. Algunas de ellas son: `abs`,`sign`, `sqrt`, `log`, `log10`, `exp`, `sin`, `cos`, `tan`, `arcsin`, `arccos`, `arctan`, `sinh`, `cosh`, `tanh`, `arcsinh`, `arccosh` y `arctanh`.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 790, "status": "ok", "timestamp": 1604326618268, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="9a0g9Tr-F5lN" outputId="11513eb5-a2c2-47b6-e0fc-05dab944e749"
np.sqrt(np.array([1,2,3,4]))
# + [markdown] id="of0yoElrF5lf"
# #### 5.4. Secuencias en Numpy
#
# La función propia de numpy es `np.arange(ini,fin,paso)`:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 753, "status": "ok", "timestamp": 1604326623937, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="o2S-lYZKF5lg" outputId="5e65f234-950d-492e-cc5a-a2f40a3401ce"
np.arange(0,6,2)
# + [markdown] id="JbsfFFn0F5l6"
# Secuencias de ceros:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 824, "status": "ok", "timestamp": 1604326627070, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="P5mx3T7vF5mI" outputId="0be34b84-8f3c-440c-9e29-009013147536"
np.zeros(3)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 755, "status": "ok", "timestamp": 1604326629981, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="Y5RQA4axF5mZ" outputId="cb2346cd-fec8-4469-9233-85e05d17cc0a"
np.ones(3)
# + [markdown] id="dQ--OIeTF5mu"
# Repetir todo el vector:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 723, "status": "ok", "timestamp": 1604326633361, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="KUOAd3QyF5mw" outputId="cc462648-593f-4040-8fa8-cd530785c0e1"
a = [1,2,3]
b = np.tile(a, 4)
b
# + [markdown] id="srRSu5prF5m-"
# Repetir cada elemento de un vector:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 712, "status": "ok", "timestamp": 1604326635651, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="8QBZbs1iF5nD" outputId="b02b9afe-b6b5-4078-de73-74cf1bc2d1ff"
a = [1,2,3]
b = np.repeat(a, 4)
b
# + [markdown] id="FW0ufl7VF5nR"
# #### 5.5. Dimensiones en Numpy
# + id="mYSsHlVDF5nT"
np.random.seed(123) # Semilla para poder reproducir
x1 = np.random.randint(10, size=6) # array 1-dimensional
x2 = np.random.randint(10, size=(3, 4)) # array 2-dimensional
x3 = np.random.randint(10, size=(3, 4, 5)) # array 3-dimensional
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 928, "status": "ok", "timestamp": 1604326640842, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="PS_a8nRpF5nj" outputId="8ecd8efe-9467-4929-922c-7bc47ebe7915"
print("x3 número dimensiones: ", x3.ndim)
print("x3 shape:", x3.shape)
print("x3 número de elmentos: ", x3.size)
print("X3 cantidad de bytes:", x3.nbytes, "bytes")
# + [markdown] id="R2qk6tm6F5nz"
# Nuestra matriz de naves interestelares:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 698, "status": "ok", "timestamp": 1604326643735, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="aLwc3zQgF5n2" outputId="5d76cdbb-06b9-4778-a57e-0ed21767021e"
print("Naves número dimensiones: ", starship_np.ndim)
print("Nombres número dimensiones: ", starship_names.ndim)
print("Naves shape:", starship_np.shape)
print("Nombres shape:", starship_names.shape)
print("Naves número de elmentos: ", starship_np.size)
print("Nombres número de elmentos: ", starship_names.size)
print("Naves cantidad de bytes:", starship_np.nbytes, "bytes")
print("Nombres cantidad de bytes:", starship_names.nbytes, "bytes")
# + [markdown] id="7a2sTa97F5oC"
# ### 5.6. Filtros en Numpy
#
# Siguen las mismas reglas que en las listas pero, ahora, con más dimensiones:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 724, "status": "ok", "timestamp": 1604326647250, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="bvqonHw9F5oF" outputId="6a7fe86c-de44-428b-af55-525e2869a3b8"
x2
# + [markdown] id="I_yUJTgzF5oX"
# Elemento de la matriz:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 971, "status": "ok", "timestamp": 1604326650163, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="HKZCQhkxF5oa" outputId="733ad73e-d5b4-42b9-e11e-f5c4b0c01360"
x2[0, 0]
# + [markdown] id="17gJYTNsF5os"
# #### 5.6.1. Filtros Slicing
# + [markdown] id="Qdp_uJw5F5ou"
# Vector segunda columna:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 706, "status": "ok", "timestamp": 1604326653067, "user": {"displayName": "al<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="SCxDxiIbF5ox" outputId="b523737e-6ff3-40ab-9025-93f463857b7e"
x2[:,1]
# + [markdown] id="t76C2F36F5o-"
# Vector tercera fila:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 833, "status": "ok", "timestamp": 1604326655646, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="yrwqGVAZF5pA" outputId="6eeed2e1-e627-48bb-f0f6-e750b0ab26da"
x2[2,:]
# + [markdown] id="YsQ7OHqdF5pN"
# O, la misma fila, por omisión del segundo elemento:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 695, "status": "ok", "timestamp": 1604326657671, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="VjYzpEaLF5pO" outputId="224ccde6-2d68-4b13-ad43-f8acecf54ed3"
x2[2]
# + [markdown] id="-lLsvMBcF5pa"
# En nuestras naves, las columnas eran: cost_in_credits, MGLT, crew.
#
# Para acceder a la velocidad (MGLT):
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 716, "status": "ok", "timestamp": 1604326661283, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="gbTrT3laF5pc" outputId="429e7c0b-e74d-4aaf-8f55-cacccd4ecacd"
starship_np[:,1]
# + [markdown] id="IqlTodMXF5po"
# Para seleccionar la 5ª nave, se selecciona una fila 4:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 923, "status": "ok", "timestamp": 1604326663930, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="q3rFk6VLF5pq" outputId="f02db35c-9e5d-4bc5-c9f8-7fe1d8aab16d"
starship_np[4,:]
# + [markdown] id="tALLtgjWF5p1"
# Para ver su nombre:
# + colab={"base_uri": "https://localhost:8080/", "height": 36} executionInfo={"elapsed": 699, "status": "ok", "timestamp": 1604326666983, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="I4w7HcbNF5p3" outputId="143b261e-a7df-4cff-9bd1-cf2d543f8893"
starship_names[4]
# + [markdown] id="LKCQie2nF5qF"
# #### 5.6.2. Indexación Elegante
#
# + [markdown] id="I3NKSRueNa7w"
#
# ##### 5.6.2.1. Filtros Posicionales
#
# Otra mejora respecto a las listas es la capacidad de indexar por una lista o array.
#
# Para filtrar por un array unidimensional de posiciones:
# + colab={"base_uri": "https://localhost:8080/"} eval=false executionInfo={"elapsed": 847, "status": "ok", "timestamp": 1604326671822, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="Bzxtn7D4F5qH" outputId="ab463c47-4550-490a-9b81-5fbc3fc7b06b"
x1
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 724, "status": "ok", "timestamp": 1604326674354, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="IF_5ggmxF5qU" outputId="dc19836c-8f7c-4ca0-f9a0-31b061f5f391"
ind = [0,2,5]
x1[ind]
# + [markdown] id="hzVN9mz0F5qg"
# > **Recuerda**: En el objeto list, esta operación no es posible. En su lugar, se utilizan las list comprehension.
# + [markdown] id="6fTjZ7ilF5qj"
# En dos dimensiones, a diferencia de lo visto en R, se aparean por coordenadas filas y columnas, es lo que se llama Broadcasting. Lo veremos más adelante:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1339, "status": "ok", "timestamp": 1604326677782, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="ZkIXVobTF5ql" outputId="29ebccac-bac5-4d5f-ee20-2db1107bb1fa"
x2
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 759, "status": "ok", "timestamp": 1604326681014, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="p6KYPXh8F5qz" outputId="ab6407c3-99ed-4356-ee68-e108e5449143"
fils = np.array([0,1])
cols = np.array([0,3])
x2[fils,cols]
# + [markdown] id="qyWa2jaNF5q-"
# > **Recuerda**: en arrays numpy los filtros por posición, se seleccionan apareando coordenadas fila y columna. Ve a la sección de Broadcasting, para saber más.
#
# En R se obtendría el siguiente resultado:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 745, "status": "ok", "timestamp": 1604326683774, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="gfNySjsKF5rB" outputId="a845e65c-575d-43b7-a9c8-0f7856a126bc"
x2[fils,:][:,cols]
# + [markdown] id="hgR3nmqKF5rR"
# ##### 5.6.2.2. Filtros Lógicos
#
# Otro mecanismo *fancy indexing* son los filtros lógicos.
#
# Antes, recordemos que para obtener el número de tripulantes (crew):
# + colab={"base_uri": "https://localhost:8080/"} eval=false executionInfo={"elapsed": 740, "status": "ok", "timestamp": 1604326687668, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="aeogC5IpF5rU" outputId="63c6d1d1-8372-49c1-b1c2-2c37af25c15f"
starship_np[:,2]
# + [markdown] id="9k7-i9XhF5rz"
# Para saber cuáles son las naves de 4 o más tripulantes:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 732, "status": "ok", "timestamp": 1604326690810, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="CmjmNg6vF5r6" outputId="8606e42c-31a2-47fd-be75-e8b93b897729"
starship_np[:,2] >= 4
# + [markdown] id="9Ii6jneXF5sF"
# Si queremos rescatar sus nombres:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1050, "status": "ok", "timestamp": 1604326694346, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="4nBBBfLWF5sH" outputId="5a142e92-b073-4cd3-a367-f0a15c9fa161"
starship_names[starship_np[:,2] >= 4]
# + [markdown] id="GHAPA4YbF5sY"
# Continúan funcionando los operadores distinto `!=`, en `in`, no en `not in`.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 745, "status": "ok", "timestamp": 1604326697476, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="wEnCpgtmF5sc" outputId="a91f9e11-61dd-42e2-8186-e14cf9181ccb"
'<NAME>' in starship_names[starship_np[:,2] >= 4]
# + [markdown] id="JSW1NIoEF5sm"
# > Para reproducir el mismo comportamiento que el operador `a %in% b` de R se utiliza `np.isin(a, b)`:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 777, "status": "ok", "timestamp": 1604326701288, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="ka4sThFQF5so" outputId="4a2f7ca5-578f-4d95-a572-dd08dda8552c"
a = np.array([1, 2, 3, 4])
b = np.array([3, 4, 5])
np.isin(a,b)
# + [markdown] id="pCLknuxWF5s2"
# ##### 5.6.2.3. Operaciones Booleanas
#
# Utilizamos los `&`, `|` y `~` en vez de los `and`, `or`, `not` (respec.) utilizados en los tipos lógicos. Será necesario cerrar las expresiones entre paréntesis.
#
# Por ejemplo:
#
# ```
# ( A > 1 ) | ( B < 5)
# ```
#
# Respecto a nuestras naves, las que tienen 4 o más tripulantes y que cuestan menos de 1M de créditos:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 821, "status": "ok", "timestamp": 1604326707326, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="rtyiW1uSF5s5" outputId="c7f95a1d-3329-4f13-9268-3e07f04c9b28"
starship_names_sel = starship_names[(starship_np[:,2] >= 4) & (starship_np[:,0] <10**6)]
starship_names_sel
# + [markdown] eval=false id="kA2SqD2gF5tH"
# > **Recuerda**: las dos expresiones van entre paréntesis.
#
# Ahora, filtramos la matriz de datos de estas naves:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 708, "status": "ok", "timestamp": 1604326710800, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="lqt9A89qF5tJ" outputId="091867dc-3cf8-4580-affb-2d0b1d1aea3e"
starship_np_sel = starship_np[(starship_np[:,2] >= 4) & (starship_np[:,0] <10**6), :]
starship_np_sel
# + [markdown] id="qUH_kQRhF5tZ"
# Gráficamente:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 724, "status": "ok", "timestamp": 1604326718507, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="Xsp2l1BlF5ta" outputId="42a81388-8aee-4a78-a6b0-fcbb4f6e7291"
starship_names_sel
# + colab={"base_uri": "https://localhost:8080/", "height": 276} executionInfo={"elapsed": 877, "status": "ok", "timestamp": 1604326722483, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="wjqHoIwWF5uF" outputId="1a05acf4-cc5f-4fc8-f917-1bbcf3120fa1"
ind = np.arange(len(starship_names_sel)) # posiciones
plt.bar(ind, starship_np_sel[:,1] );
for i in ind:
plt.text(ind[i],starship_np_sel[i,1], int(starship_np_sel[i,1]),size=9)
plt.gcf().subplots_adjust(left=0.15,bottom=0.15)
plt.ylabel('Velocidad (MGLT)')
plt.title('Naves Interestelares seleccionadas')
p = plt.xticks(ind, starship_names_sel,fontsize=9);
# + [markdown] id="gWMz4gPhF5uQ"
# ¡Parece que el Halcón Milenario con 75 MGLT, es la nave que andábamos buscando!
# + [markdown] id="eAet3mf2F5uR"
# ### 5.7. Modificar Arrays
# + [markdown] id="7x4YB74LF5uT"
# Para modificar un valor de un array:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 748, "status": "ok", "timestamp": 1604326730789, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="RY6yeyrwF5uW" outputId="b827bab1-71a4-41c4-ef29-524d8cf32062"
x2
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 906, "status": "ok", "timestamp": 1604326733980, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="qUqrIZilF5ug" outputId="75aaaf1e-652a-4c9f-91d9-0e6aeacbd6f6"
x2[1,1] = 20
x2
# + [markdown] id="5A9YGcX_F5uq"
# Mucha precaución con el tipo interno del numpy ya que es fijo. No se va a modificar:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 591, "status": "ok", "timestamp": 1604326736979, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="JWr3f2NaF5us" outputId="da061563-f518-4b9b-8712-b2fdda2648b4"
x2[1,1] = 20.765
x2
# + [markdown] id="Xmbpao2cF5u3"
# > **Recuerda**: Si intentas insertar un valor en coma flotante sobre un numpy con tipo entero, éste se va a cortar.
# + [markdown] id="w47oaaNuF5u5"
# Se pueden modificar N valores con un vector de N posiciones.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 988, "status": "ok", "timestamp": 1604326743157, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="rAC7e-K-F5u7" outputId="37d11c0f-2505-4797-b78b-ac09fa9298da"
x2[1,:] = x2[0,:] * 10
x2
# + [markdown] id="uG-EMgQhF5vE"
# También, se pueden modificar varios a la vez, con un único valor.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 704, "status": "ok", "timestamp": 1604326746856, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="7rpUJ1htF5vG" outputId="f90b6f7b-1ad4-4143-d336-aa68734d3d78"
x2[[1,2],[2,2]] = 55
x2
# + [markdown] id="wSDlx02cF5vO"
# #### 5.7.1. Vistas no copiadas
#
# Cuando se asigna un array numpy a otro objeto, ya sea filtrado con `[,]` o no, éste retorna una vista, no una copia. Es muy importante tenerlo en cuenta, cuando se modifica el objeto:
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 713, "status": "ok", "timestamp": 1604326751570, "user": {"displayName": "al<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="dHJBZufYF5vP" outputId="ff3ea001-1ebe-4ee6-f045-b391f32e0ae3"
a = np.array([1, 'm', [2, 3, 4]], dtype=object)
b = a
b[0] = 10
a
# + [markdown] id="DPDNr5lkF5v4"
# > **Importante**: Los cambios en una vista no copiada, afectan a todas las copias.
# + [markdown] id="spX1SkU1F5v6"
# #### 5.7.2. Copias de Arrays
#
# Para copiar un array a otro objeto utiliza `copy()`:
#
# + id="-3M275Q2F5v8"
c = a.copy() # COPIAS DE ARRAYS
c[0] = 20
print(a)
# + [markdown] id="vcKacYLNF5wk"
# Ahora, el objeto original, no cambia. Pero no es perfecto!
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 824, "status": "ok", "timestamp": 1604326766577, "user": {"displayName": "alumnos bigdata", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="Qh8haXqlF5wc" outputId="1cd9da91-89b1-47c6-af0e-349edb4424f8"
c[2][0] = 30
print(a)
# + [markdown] id="vcKacYLNF5wk"
# Internamente continua utilizando referencias a los mismos objetos.
# + [markdown] id="spX1SkU1F5v6"
# #### 5.7.2.1 Copias profundas de Arrays
#
# Veamos en que consiste.
# + id="-3M275Q2F5v8"
import copy
d = copy.deepcopy(a)
d[2][0] = 40
print(a)
# + [markdown] id="vcKacYLNF5wk"
# Ahora copia la estructura interna.
# + [markdown] id="avjeAduPF5wn"
# #### 5.7.3. Cambios de *Shape*
#
# Con `.reshape()` se puede modificar las dimensiones del array:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 747, "status": "ok", "timestamp": 1604326769644, "user": {"displayName": "alumnos bigdata", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="mHQoGxAxF5wo" outputId="a32eabc5-dbc3-4d6d-e56b-5f7040086e31"
x1
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 767, "status": "ok", "timestamp": 1604326771591, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="AjoOiMuoF5wy" outputId="cbee51f9-b27b-4397-9e7b-9cc0084bfdc9"
x1.reshape(2,3)
# + [markdown] id="UVgmVZHeF5w-"
# #### 5.7.4. Concatenar Arrays
#
# Para concatenar 2 arrays unidimensionales:
#
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 762, "status": "ok", "timestamp": 1604326774356, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="xySi2-EQF5xA" outputId="6c3bec0f-3923-4286-f165-0290ec3c1c14"
x = np.array([1, 2, 3])
y = np.array([3, 2, 1])
np.concatenate([x, y])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 969, "status": "ok", "timestamp": 1604326777319, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="204vVEu7F5xH" outputId="17d27b42-4333-4b08-c926-316ad48ef647"
grid = np.arange(6).reshape(2,3)
# concatenar filas, (axis = 0)
np.concatenate([grid, grid])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1026, "status": "ok", "timestamp": 1604326780277, "user": {"displayName": "alumnos bigdata", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="OBHUUFrXF5xQ" outputId="750f6d7e-48d1-4ab2-ea30-1b02b0821ec0"
# concatenar columnas, (axis = 1)
np.concatenate([grid, grid], axis=1)
# + [markdown] id="igos5GLqF5xY"
# Lo mismo con vstack y hstack, pero con arrays de distinta dimensión:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 886, "status": "ok", "timestamp": 1604326782473, "user": {"displayName": "al<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="IL2fzMqPF5xZ" outputId="d6a755d4-3527-47d9-ce0d-d208afe197f9"
x = np.array([1, 2, 3])
grid = np.array([[9, 8, 7],
[6, 5, 4]])
# apilado vertical
np.vstack([x, grid])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 770, "status": "ok", "timestamp": 1604326784825, "user": {"displayName": "alumnos bigdata", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="itnAKYXSF5xi" outputId="c684bdf1-c97f-4c04-df2d-0afc32bb376a"
# apilado horizontal
y = np.array([[99],
[99]])
np.hstack([grid, y])
# + [markdown] id="w1XZYlwiF5xp"
# #### 5.7.5. Broadcasting de Arrays
#
# El Broadcasting es una operación de alineamiento que sigue, de forma estricta, las siguientes reglas de interacción entre 2 arrays:
#
# - **Regla 1**: si dos arrays difieren en el número de dimensiones, el shape (dimensiones) del que tenga menor dimensión se aumenta con unos por la izquierda:
# ```
# [3,1,6] x [2,6] ----> [3,1,6] x [1,2,6]
# [3,5,4] x [3,5] ----> [3,5,4] x [1,3,5]
# ```
# - **Regla 2**: si dos arrays tienen shape distinto en alguna dimensión, el array con shape igual a 1 se estira (copia) para que concuerde con el de la misma dimensión del otro array:
#
# ```
# [3,1,6] x [1,2,6] ----> [3,2,6] x [3,2,6]
# [3,5,4] x [1,3,5] ----> [3,5,4] x [3,3,5]
# ```
# - **Regla 3**: si alguna dimensión tiene tamaño distinto y no hay ninguna dimensión 1, se lanza un error:
# ```
# [3,2,6] x [3,2,6] ----> Concuerdan
# [3,5,4] x [3,3,5] ----> Error !!!
# ```
#
# En la práctica, esto significa facilitar algunas operaciones:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 758, "status": "ok", "timestamp": 1604326791997, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="_kr8Q8n-F5xr" outputId="ffed0d85-c117-419d-88f9-f74b8b42bcf7"
a = np.ones((2,2))
a
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 733, "status": "ok", "timestamp": 1604326794222, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="4AQXc-Z7F5xz" outputId="5f3362bc-1daa-46b8-c94b-93a9565f19e3"
b = np.tile(0.5, 2)
b
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 762, "status": "ok", "timestamp": 1604326796582, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="jjD0XQ-7F5x9" outputId="f94c9bba-860f-4756-9203-4cc895f72643"
a - b
# + [markdown] id="JBfNveiBF5yI"
# #### 5.7.6. Conversión a Listas
#
# Los numpy se convierten a lista con `.tolist()`:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 705, "status": "ok", "timestamp": 1604326798410, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="fRK9StENF5yK" outputId="d9251008-808d-4d3e-fc06-b0d520b47878"
a = np.array([2,3,7,3])
b = a.tolist()
b
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 936, "status": "ok", "timestamp": 1604326800935, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="Eil45oqQF5yY" outputId="42bbab1b-9a96-4a93-f3d7-e7fba6e68cd2"
type(b)
# + [markdown] id="Bp8rxnCNF5yg"
# #### 5.7.7. Valores Faltantes
#
# Por definición, un NaN es un número en coma flotante, distinto a cualquier otro número:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 797, "status": "ok", "timestamp": 1604326805833, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="JN99POGzF5yh" outputId="54816b7b-6d56-4879-9b6c-bef32f868cb7"
np.nan != np.nan
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 724, "status": "ok", "timestamp": 1604326808237, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="jSkP_yiNF5yq" outputId="85223352-3d0b-4ec0-c5c4-47b77681a418"
a = np.arange(6).reshape(3,2) * 1.0
a[[1,2],[1] ] = np.nan
a
# + [markdown] id="HvgRXXB7F5yy"
# > **Observa**: se ha aplicado broadcast al asignar `np.nan`.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1055, "status": "ok", "timestamp": 1604326811467, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10710302360204190833"}, "user_tz": -60} id="kSLDP5V8F5y0" outputId="1469adde-c76e-46e9-d82d-f7d10380ffa0"
np.isnan(a)
# + [markdown] id="l5ykUksQF5y7"
# Numpy es la base de la computación con arrays. Veamos ahora qué aportan los objetos **pandas**.
# -
# < [Colecciones de objetos: listas, tuplas y diccionarios](modulo1_tema4_Py_30_colec_obj.ipynb) | [Índice](modulo1_tema4_Py_00_indice.ipynb) | [Colecciones de objetos: pandas](modulo1_tema4_Py_32_pandas.ipynb) >
#
# __[Abre en Colab](https://colab.research.google.com/github/griu/init_python_b1/blob/master/modulo1_tema4_Py_31_numpy.ipynb)__
| modulo1_tema4_Py_31_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: geocomp
# language: python
# name: geocomp
# ---
# imports
import numpy as np
import timeit
# read files
f1 = np.load('entropy_20180610.npy')
f2 = np.load('mid_unit.npy')
f3 = np.load('realisation_1_10.npy')
# file shape
f1.shape, f2.shape, f3.shape
# file size
f1.size, f2.size, f3.size
# save as array
f1_array = np.array(f1)
f2_array = np.array(f2)
f3_array = np.array(f3)
# flatten to 2D array
f1_array_flat = f1_array.reshape(250,16200)
f2_array_flat = f2_array.reshape(250,16200)
f3_array_flat = f3_array.reshape(250, 3240000)
# reshape to original dimensions
f1_array_reshaped = f1_array_flat.reshape(250, 162, 100)
f2_array_reshaped = f2_array_flat.reshape(250, 162, 100)
f3_array_reshaped = f3_array_flat.reshape(250, 162, 100, 200)
# check if arrays equal after reshape
np.array_equal(f1_array, f1_array_reshaped),
np.array_equal(f2_array, f2_array_reshaped),
np.array_equal(f3_array, f3_array_reshaped)
# +
# save to csv
#np.savetxt("f1_array_flat.csv", f1_array_flat, delimiter=',')
#np.savetxt("f2_array_flat.csv", f2_array_flat, delimiter=',')
#np.savetxt("f3_array_flat.csv", f3_array_flat, delimiter=',')
# -
# save to npy
np.save("f1_array_flat", f1_array_flat, allow_pickle=False)
np.save("f2_array_flat", f2_array_flat, allow_pickle=False)
np.save("f3_array_flat", f3_array_flat, allow_pickle=False)
# +
# #%timeit np.save("testnpy", f1_array_flat, allow_pickle=False)
# +
# #%timeit np.savetxt("testcsv.csv", f1_array_flat, delimiter=',')
# -
| static/Pickles_to_csv_flatten_arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: phm08ds
# language: python
# name: phm08ds
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import preprocessing
from sklearn import metrics
import os, sys
from time import time
from phm08ds.models import experiment
# -
# ## Load Dataset
folderpath = '../../../data/interim/'
data_op_04 = pd.read_csv(folderpath + 'data_op_04.csv')
data_op_04.head()
# ## Data preprocessing
# ### Get rid of informations there are not sensor readings
# Wang (2008) reports Sensor 15 has importat information. However, there are no relevant informations of this sensor. The data seems to be corrupted like this:
# Let's remove it from our database creating an object transformer.
# +
from phm08ds.features.feature_selection import RemoveSensor
tf_remove_sensor_15 = RemoveSensor(sensors=[15])
data_op_04 = tf_remove_sensor_15.fit_transform(data_op_04)
data_op_04.head()
# -
# Before feeding to the classifier, let's remove unwanted information, such as unit, time_step and operational settings.
# +
from phm08ds.features.feature_selection import RemoveInfo
tf_remove_info = RemoveInfo()
data_with_features = tf_remove_info.fit_transform(data_op_04)
data_with_features.head()
# -
# We need to normalize our data. Let's use Z-score standardization.
# +
from sklearn.preprocessing import StandardScaler
tf_std_scaller = preprocessing.StandardScaler()
data_with_features_std = tf_std_scaller.fit_transform(data_with_features.drop(labels='Health_state', axis=1))
data_with_features_std
# -
labels = np.array(data_with_features['Health_state'])
labels
# # Classification steps
from phm08ds.models import experiment
# ## Load best mlp from random search
# +
from sklearn.externals import joblib
random_search_mlp = joblib.load('Results/old_models/clf_mlp.pkl')
# random_search_svm = joblib.load('Results/old_models/clf_svm.pkl')
# -
clf_mlp = random_search_mlp['MLP'].best_estimator_
# ## Put all clf in a dictionary:
classifiers = {'MLP': clf_mlp}
# Since we are using SVM and MLP we need to extract all power from those methods. Let's perform a Random Search to parameters optimizations.
kfolds = 10
clf_outputs = experiment.run_classifiers(data_with_features_std, labels, classifiers, kfolds)
# ## Performance assessment
# Saving variables in a dictionary:
results = {}
results['train'] = experiment.results_clf(4, clf_outputs['train']['true'], clf_outputs['train']['pred'])
results['test'] = experiment.results_clf(4, clf_outputs['test']['true'], clf_outputs['test']['pred'])
# ## Savel results, models and pipeline to a .pkl file
# +
from sklearn.pipeline import Pipeline
data_preprocessing = Pipeline([('remove_sensor_15', tf_remove_sensor_15),
('remove_info', tf_remove_info),
('std_scaler', tf_std_scaller)
])
# -
experiment.save_models(classifiers)
experiment.save_pipeline(data_preprocessing)
# ## Save results to CSVs and figures
experiment.export_results(results['test'], 'test')
experiment.export_results(results['train'], 'train')
# ## Savel results, models and pipeline to a .pkl file
# +
from sklearn.pipeline import Pipeline
data_preprocessing = Pipeline([('remove_sensor_15', tf_remove_sensor_15),
('remove_info', tf_remove_info),
('std_scaler', tf_std_scaller)
])
# -
experiment.save_models(clf_mlp, name='clf_mlp')
experiment.save_pipeline(data_preprocessing)
| notebooks/E03_PHM08-train-best_MLP_and_SVM/model_selection-OP_04-navarmn/0.1-model_selection_MLP--OP_04-navarmn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import display, HTML
import pygments
display(HTML("<style>.container { width:290mm !important; }</style>")) # to set cell widths
#to get this into the dissertation,
#1. widht (above) was changed from 290 to 266 mm
#2. exported with jupyter notebook as html (space gets wider),
#3. converted to a pdf with https://www.sejda.com/de/html-to-pdf (space gets wider again),
#4. placed as a Verknüpfung into Adobe illustrator, where for each page separate Zeichenflächen
# have been chosen. For their placement the coordinate origin should be changed from "centered"
# to "left".
#5. Each Zeichenfläche was saved separately and loaded in LaTeX
formatter = pygments.formatters.get_formatter_by_name('html', linenos='inline')
import numpy as np
import pandas as pd
# #### Calculation: Fiber volume fraction & Relative Density
# +
### Parameters ##########################
# warp fiber radius:
r_wa = 75 #[µm],
# horizontal distance between surfaces of fiber 1 (left) and fiber 2 (right):
dh_f1s_f2s = np.linspace(0,240,500) #[µm],
# vertical distance between the ground surface and the surfaces of the lower fiber (here fiber 1):
d_gs_f1s = np.linspace(0,140,int(500*140/200)) #[µm],
# vertical distance between the lower and higher fiber:
vertical_displ = np.linspace(0,140,3) #[µm]
def calc_fiber_vol_frac(r_wa, dh_f1s_f2s, d_gs_f1s, vertical_displ):
# fiber
fiber_area = 0.5*np.pi * r_wa**2 #[µm²]
# horizontal distance between the fiber centers:
b = dh_f1s_f2s + 2 * r_wa #[µm]
# area, in which the fiber volume fraction is calculated
controll_area = b * (r_wa + d_gs_f1s + vertical_displ/2) #[µm²]
# fiber volume fraction:
fiber_vol_frac = fiber_area / controll_area #[-]
return fiber_vol_frac
def calc_rel_density(r_wa, dh_f1s_f2s, d_gs_f1s, vertical_displ):
# distance between surfaces of ground and higher fiber
d_gs_f2s = d_gs_f1s + vertical_displ #[µm],
# horizontal distance between the the fiber centers
b = dh_f1s_f2s + 2 * r_wa #[µm],
# distance between surfaces of fiber 1 (left) and fiber 2 (right):
d_f1s_f2s = (b**2 + vertical_displ**2)**0.5 - 2 * r_wa
# radius until "CVD surface-to-surface contact" for uniform growth
r_grown = r_wa + d_f1s_f2s/2
# area, in which the relative density is calculated
controll_area = b * (r_wa + d_gs_f1s + vertical_displ/2) #[µm²]
# part of the radius "h", for which the circle (fiber) overlaps with the ground,
h1 = d_f1s_f2s - d_gs_f1s
h2 = d_f1s_f2s - d_gs_f2s
def calc_A(h):
'''
to calculate the part of the left or right fiber area (A/2) that needs to be removed
see https://de.wikipedia.org/wiki/Kreissegment
'''
if isinstance(h, int) or isinstance(h, float):
if h < 0: h = 0
else: #numpy array:
h[h < 0] = 0
A = r_grown**2 * np.arccos(1 - h/r_grown) - (r_grown - h) * (2*r_grown*h - h**2)**0.5
return A
# solid area = grown fiber area - overlap + grown ground area
solid_area = np.pi/2 * r_grown**2 - (calc_A(h1) + calc_A(h2))/2 + b*d_f1s_f2s/2
rel_density = solid_area / controll_area
return rel_density
# -
# #### Contour plots
import matplotlib.pyplot as plt
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter, AutoMinorLocator)
from matplotlib.colors import ListedColormap
def contourPlot(vertical_displ = 0, markers_x = [], markers_y = []):
fig, ax = plt.subplots(1,1)
### Plot labeling and settings #########################
ax.set_title('$Y^{´}_{\!\!ff}$ = ' + str(vertical_displ) + ' µm', fontsize = 12)
ax.set_xlabel('$X_{ff}$ [µm]')
ax.set_ylabel('$Y_{fg}$ [µm]')
# grid and ticks
ax.grid(alpha = 0.4)
ax.set_xlim(0, 245)
ax.set_xticks(np.arange(0, 260, 20).tolist())
ax.xaxis.set_minor_locator(MultipleLocator(4))
ax.yaxis.set_minor_locator(MultipleLocator(4))
# rel. density and fiber vol. fraction labels
ax.text(160, 130 - vertical_displ/2, r'$\rho_{rel}$', color = 'g',
backgroundcolor = 'w', fontsize = 14)
ax.text(90, 130 - vertical_displ/2, r'$\Phi_{V_f}$', color = 'b',
backgroundcolor = 'w', fontsize = 14)
########################################################
# for manual contour label positions:
def getLabelPosis(x1, x2, y1, y2, n = 6):
y1b = y1 + 8 - 0.42*vertical_displ
y2b = y2 + 8 - 0.42*vertical_displ
m = (y2b - y1b) / (x2 - x1)
b = y2b - x2*(y2b - y1b) / (x2 - x1)
f = lambda x: m*x + b
x = np.linspace(x1, x2, n)
result = []
for i in range(n):
result += [(x[i], f(x[i]))]
return result
# slice contour plot color map
def sliceCmap(cmap_name, lo = 0.5, hi = 0.9):
cmap = plt.cm.get_cmap(cmap_name, 512)
return ListedColormap(cmap(np.linspace(lo, hi, 256)))
x = dh_f1s_f2s
y = d_gs_f1s
X, Y = np.meshgrid(x, y)
# cotour plot for rel. density
Z = calc_rel_density(r_wa, X, Y, vertical_displ)
rel_dens_levels=[0.97,0.98,0.99,0.996,0.999,1]
CS = plt.contour(X, Y, Z, levels=rel_dens_levels, linewidths = 2, cmap = sliceCmap('Greens'))
label_x_pos = 200
ax.clabel(CS, inline=1, fontsize=11,
manual = getLabelPosis(x1 = 205, x2 = 215, y1 = 120, y2 = 25, n = len(rel_dens_levels)))
# cotour plot for fiber vol. fraction
Z = calc_fiber_vol_frac(r_wa, X, Y, vertical_displ)
f_frac_levels = [0.2,0.25,0.30,0.35]
CS = plt.contour(X, Y, Z, levels= f_frac_levels, linewidths = 2, cmap = sliceCmap('Blues'))
label_x_pos = 30
ax.clabel(CS, inline=1, fontsize=11,
manual = getLabelPosis(x1 = 20, x2 = 60, y1 = 60, y2 = 125, n = len(f_frac_levels)))
# xy marker lines
marker_colors = ['r', 'orange']
for i in range(len(markers_x)):
ax.plot((0, markers_x[i]),(markers_y[i], markers_y[i]), color = marker_colors[i], zorder = -1)
ax.plot((markers_x[i], markers_x[i]),(0, markers_y[i]), color = marker_colors[i], zorder = -1)
contourPlot(vertical_displ = 27, markers_x = [], markers_y = [])
contourPlot(vertical_displ = 25, markers_x = [125,210], markers_y = [40,64])
| Python scripts/theoretical rel density for uniform growth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import shap
import sklearn
from sklearn.cluster import KMeans
from sklearn.manifold import TSNE
from sklearn.decomposition import PCA
import bhtsne
import matplotlib
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
import seaborn as sns
shap.initjs()
# -
# <h2> Data Loading and Preprocessing </h2>
counts = pd.read_csv('/Users/vincentliu/Desktop/PeerLab/SCRAS/Data/pbmc_4k_dense.csv', index_col=0)
counts = counts.astype(np.int32)
# <h3> Cell and Gene Filtering </h3>
# +
ms = np.log10( counts.sum(axis=1) )
# Figure
plt.hist(ms, 100)
print(counts.shape)
# +
# Cells passing the threshold
ms = counts.sum(axis = 1)
use_cells = ms.index[np.log10(ms) > 3]
# Remove zero sum genes
use_genes = counts.columns[counts.loc[use_cells,:].sum() > 0]
# Update counts
counts = counts.loc[use_cells, use_genes]
# -
# Figure
ms = np.log10(counts.sum(axis=1))
plt.hist(ms, 100)
print(counts.shape)
ms = counts.sum(axis=1)
norm_df = counts.div(ms, axis=0).mul(np.median(ms), axis=0) # normalization based on median
# Visualize gene distribution
gene_counts = (norm_df > 0).sum()
plt.figure()
sns.kdeplot(np.log10(gene_counts))
# +
use_genes = gene_counts.index[gene_counts > 10]
percent = len(use_genes) / len(gene_counts)
print("percent genes used: {}".format(str(percent)))
counts = counts.loc[:, use_genes]
norm_df = norm_df.loc[:, use_genes]
log_df = np.log2(norm_df+0.1)
# -
# <h2> Dimensionality Reduction </h2>
# <h3> PCA </h3>
pca = PCA(n_components=10, svd_solver='randomized')
pcaproj = pd.DataFrame(pca.fit_transform(norm_df), index=norm_df.index)
print(np.sum(pca.explained_variance_ratio_))
# <h3> tSNE </h3>
# +
from copy import deepcopy
data = deepcopy(norm_df)
tsne = pd.DataFrame(bhtsne.tsne(data, perplexity=150), index=data.index, columns=['x', 'y'])
# -
# Library size plot
plt.figure(figsize=[7, 6])
plt.scatter(tsne['x'], tsne['y'], s=5, edgecolors='none',
cmap=matplotlib.cm.Spectral_r, c=np.log10(ms[tsne.index]))
plt.colorbar()
plt.title("Molecule count per cell")
# <h2> Clustering </h2>
# +
import phenograph
# Cluster and cluster centrolds
communities, graph, Q = phenograph.cluster(pcaproj, k=200)
communities = pd.Series(communities, index=pcaproj.index)
# +
# Best prediction
fig = plt.figure(figsize=[4*4, 2*4])
for i, cluster in enumerate(set(communities)):
cells = communities.index[communities == cluster]
ax = fig.add_subplot(3, 4, i+1)
ax.scatter(tsne['x'], tsne['y'],
s=3, edgecolors='none', color='lightgrey')
ax.scatter(tsne.loc[cells, 'x'], tsne.loc[cells,'y'],
s=5, edgecolors='none')
# Clean up
ax.set_axis_off()
ax.set_title(cluster)
sns.despine()
# -
def plot_genes(exp_mat, tsne_data, genes, title):
# Set up figure grid
n = len(genes); max_cols = 5
nrows = int(np.ceil(n / max_cols)); ncols = int(min((max_cols, n)))
fig = plt.figure(figsize=[6 * ncols, 4*nrows])
for i, gene in enumerate(genes):
ax = fig.add_subplot(nrows, ncols, i+1)
ax.scatter(tsne_data.loc[:, 'x'], tsne_data.loc[:, 'y'], s=3,
cmap=matplotlib.cm.Spectral_r, c=exp_mat.loc[tsne_data.index, gene])
ax.set_title(gene)
ax.set_axis_off()
plt.suptitle(title)
plot_genes(norm_df, tsne, ['MS4A1'], 'B Cell')
# <h2> Explain using Cluster Labels </h2>
# 0 if not in cluster 2, 1 otherwise
clus_binary = [0 if x != 2 else 1 for x in communities]
knn = sklearn.neighbors.KNeighborsClassifier()
knn.fit(pcaproj, clus_binary)
f = lambda x: knn.predict_proba(pca.transform(x))[:,1]
X_rep = shap.kmeans(pcaproj, 10)
explainer = shap.KernelExplainer(f, X_rep)
shap_values_single = explainer.shap_values(pcaproj.iloc[0,:], nsamples=1000)
shap.force_plot(shap_values_single, pcaproj)
| pbmc_clustering_anaysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.datasets import imdb
from keras import preprocessing
max_features = 10000
maxlen = 20
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
x_train.shape, y_train.shape, x_test.shape, y_test.shape
# turns lists of integers into a 2D integer tensor of shape
x_train = preprocessing.sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = preprocessing.sequence.pad_sequences(x_test, maxlen=maxlen)
x_train.shape, x_test.shape
# #### building model
from keras.models import Sequential
from keras.layers import Flatten, Dense, Embedding
# +
model = Sequential()
# why 8 dimensional
model.add(Embedding(10000,8, input_length=maxlen))
# has to be flatten before connecting to dense layer
model.add(Flatten())
# We add the classifier on top
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
model.summary()
history = model.fit(x_train, y_train,
epochs=10,
batch_size=32,
validation_split=0.2)
# -
# ## Using pre-trained word embeddings
pwd
data_path='/Users/i846240/projects/_deep_learning/fastai/courses/dl1/data/aclimdb/'
# ls {data_path}
import os
train_dir = os.path.join(data_path,'train')
# ls {train_dir}
def load_data(dir_name):
"""
load data from directory to list
expects pos and neg subdirectories in dir_name
returns texts, labels as lists
"""
labels = []
texts = []
for label_type in ['neg', 'pos']:
dir_name = os.path.join(train_dir,label_type)
for fname in os.listdir(dir_name):
if fname[-4:]=='.txt':
f = open(os.path.join(dir_name,fname))
texts.append(f.read())
f.close()
if label_type == 'neg':
labels.append(0)
else:
labels.append(1)
return texts, labels
texts, labels = load_data(train_dir)
len(texts)
texts[0]
# ### Tokenize the data
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
import numpy as np
maxlen = 100 # We will cut reviews after 100 words
training_samples = 200
validation_samples = 10000
max_words = 10000 # we will only consider the top 10,000 words in the dataset
# #### https://faroit.github.io/keras-docs/1.2.2/preprocessing/text/
tokenizer = Tokenizer(num_words=max_words)
tokenizer.fit_on_texts(texts)
sequences = tokenizer.texts_to_sequences(texts)
len(sequences)
# #### get word index from tokenizer
word_index = tokenizer.word_index
print('found %s unique tokens.' % len(word_index))
word_index['man']
data = pad_sequences(sequences, maxlen=maxlen)
labels = np.asarray(labels)
data.shape, labels.shape
x_train = data[:training_samples]
y_train = labels[:training_samples]
x_val = data[training_samples: training_samples + validation_samples]
y_val = labels[training_samples: training_samples + validation_samples]
# #### Download the GloVe word embeddings
# https://nlp.stanford.edu/projects/glove/
glove_dir='/Users/i846240/projects/_deep_learning/data/glove.6B/'
# ls {glove_dir}
# !tail {os.path.join(glove_dir,'glove.6B.100d.txt')}
# #### build embeddings index with word as key and embedding index as value
# +
embeddings_index = {}
f = open(os.path.join(glove_dir,'glove.6B.100d.txt'))
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:],dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Found %s word vectors.' % len(embeddings_index))
# -
# #### build an embedding matrix that will be load into an Embedding layer
embeddings_index['man'].size
# +
embedding_dim = 100
# initialize (10000, 100) matrix with zeros
embedding_matrix = np.zeros((max_words, embedding_dim))
# loop through word in index
# (all unique words from document in numeric value)
for word, i in word_index.items():
# find word in index and index as value
embedding_vector = embeddings_index.get(word)
if i < max_words:
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
# -
max(word_index.values())
| 2019-01-23-word-embedding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Chiebukar/Deep-Learning/blob/main/dog_cat_classification_CNN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="knIlYiCo7kK_"
from google.colab import files
files.upload()
# + id="CocIgyFo8CdE"
# !mkdir -p ~/.kaggle
# !cp kaggle.json ~/.kaggle/
# !chmod 600 ~/.kaggle/kaggle.json
# + colab={"base_uri": "https://localhost:8080/"} id="gBSBeJDF8u6u" outputId="d0ca4c64-dc5f-440b-d369-1363206e0649"
# !kaggle datasets download -d arnaudeq/cats-vs-dogs-5000
# + colab={"base_uri": "https://localhost:8080/"} id="ZqvpNuzX-lAw" outputId="048c6b09-b745-4b8c-b58d-5739d7c50afc"
# !ls -d $PWD/*
# + id="sYTvg0wP_db2"
# !unzip \*.zip && rm *.zip
# + colab={"base_uri": "https://localhost:8080/"} id="5-8uCCie_f0_" outputId="122d9fa4-0041-43e3-c3e6-<PASSWORD>"
# !ls -d $PWD/*
# + id="i3XHtMAzOncd"
train_dir = 'dogs_cats_sample_5000/train'
validation_dir = 'dogs_cats_sample_5000/valid'
# + id="sH_lMNZD_sNp"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dense, Flatten, Conv2D, MaxPooling2D, Dropout
from keras.preprocessing.image import ImageDataGenerator
from keras import optimizers
# + colab={"base_uri": "https://localhost:8080/"} id="WLEQvmfjBpPu" outputId="cb7a0b42-bcfe-4699-a197-7456ea76148f"
model = Sequential()
model.add(Conv2D(32, kernel_size = (3,3), activation='relu', input_shape = (150, 150, 3)))
model.add(MaxPooling2D(2,2))
model.add(Conv2D(64, kernel_size=(3,3), activation='relu'))
model.add(MaxPooling2D(2,2))
model.add(Conv2D(128, kernel_size=(3,3), activation='relu'))
model.add(MaxPooling2D(2,2))
model.add(Conv2D(512, kernel_size=(3,3), activation='relu'))
model.add(MaxPooling2D(2,2))
model.add(Flatten())
model.add(Dense(512, activation ='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# + id="jtt37YrQE7k2"
model.compile(loss= 'binary_crossentropy', optimizer= 'rmsprop', metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="8leShvX8Ig0D" outputId="a9b273ef-3edd-4b20-bae2-b6409956b952"
# create data generator object and set scaling
train_datagen = ImageDataGenerator(rescale = 1./255)
val_datagen = ImageDataGenerator(rescale = 1./255)
train_generator = train_datagen.flow_from_directory(
train_dir,
target_size = (150,150),
batch_size = 20,
class_mode = 'binary'
)
validation_generator = val_datagen.flow_from_directory(
validation_dir,
target_size = (150,150),
batch_size = 20,
class_mode = 'binary'
)
# + colab={"base_uri": "https://localhost:8080/"} id="BrF3tQ-GPnsc" outputId="99d04ebc-8a77-477e-dae7-5ce739158707"
for data_batch, labels_batch in train_generator:
print('data batch shape', data_batch.shape)
print('data label shape', labels_batch.shape)
break
# + id="3b-dUtuyTMbH"
history = model.fit_generator(train_generator, steps_per_epoch=100,
epochs=30, validation_data=validation_generator,
validation_steps=50)
# + id="Rb-JpJ-3sFiS"
model.save('cats_and_dogs_small_1.h5')
# + id="USpFzLH_sWc_"
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="7M6dRYzCtD7d" outputId="3a04451e-c52e-4cd4-8814-03f98f90a329"
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="p4pfvMz8tUzz" outputId="0b29195f-3528-4a22-bf30-2742d3af96c3"
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="43AM478cx2yC"
# Data Augmentation
# + id="IaJY5I9Ly75q"
train_datagen = ImageDataGenerator(rescale=1./255, rotation_range=40, width_shift_range=0.2,
height_shift_range=0.2, shear_range=0.2, zoom_range=0.2,
horizontal_flip=True)
val_datagen = ImageDataGenerator(rescale=1./255)
# + colab={"base_uri": "https://localhost:8080/"} id="v5NJ1OUw1jnq" outputId="4008a680-6a7d-42db-8637-4806c607e003"
train_generator= train_datagen.flow_from_directory(train_dir,
target_size = (150,150),
batch_size = 32,
class_mode = 'binary'
)
validation_generator= val_datagen.flow_from_directory(validation_dir,
target_size = (150,150),
batch_size = 32,
class_mode = 'binary'
)
# + colab={"base_uri": "https://localhost:8080/"} id="3oBOV5Ih0Fr3" outputId="a5378804-e74e-4af3-aad5-805649fc4397"
for data_batch, labels_batch in train_generator:
print('data batch shape', data_batch.shape)
print('data label shape', labels_batch.shape)
break
# + id="edSeqkg7BstQ"
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(150, 150, 3)))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(128, (3, 3), activation='relu'))
model.add(MaxPooling2D((2, 2)))
model.add(Conv2D(128, (3, 3), activation='relu'))
model.add(MaxPooling2D((2, 2)))
model.add(Flatten())
model.add(Dropout(0.5))
model.add(Dense(512, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer=optimizers.RMSprop(lr=1e-4), metrics=['acc'])
# + id="TGZ6Rqnh7Ma2"
history = model.fit_generator(train_generator, steps_per_epoch=100,
epochs=50, validation_data=validation_generator,
validation_steps=50)
# + id="uh2fKxsY7fBA"
model.save('cats_and_dogs_small_2.h5')
# + id="-ZJrZj3G705M"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="VM1CoyKB78j2" outputId="6b2789b0-2fdc-4e2d-970b-a0599636c457"
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="w08pY-pd8GRW" outputId="3ca910b6-3fff-4bc5-8a69-72c28744202e"
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
| classification/dog_cat_classification_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook walks you through how to use accelerators for Kubeflow Pipelines steps.
#
# # Preparation
#
# If you installed Kubeflow via [kfctl](https://www.kubeflow.org/docs/gke/customizing-gke/#common-customizations), these steps will have already been done, and you can skip this section.
#
# If you installed Kubeflow Pipelines via [Google Cloud AI Platform Pipelines UI](https://console.cloud.google.com/ai-platform/pipelines/) or [Standalone manifest](https://github.com/kubeflow/pipelines/tree/master/manifests/kustomize), you willl need to follow these steps to set up your GPU enviroment.
#
# ## Add GPU nodes to your cluster
#
# To see which accelerators are available in each zone, run the following command or check the [document](https://cloud.google.com/compute/docs/gpus#gpus-list)
#
# ```
# gcloud compute accelerator-types list
# ```
#
# You may also check or edit the GCP's **GPU Quota** to make sure you still have GPU quota in the region.
#
# To reduce costs, you may want to create a zero-sized node pool for GPU and enable autoscaling.
#
# Here is an example to create a P100 GPU node pool for a cluster.
#
# ```shell
# # You may customize these parameters.
# export GPU_POOL_NAME=p100pool
# export CLUSTER_NAME=existingClusterName
# export CLUSTER_ZONE=us-west1-a
# export GPU_TYPE=nvidia-tesla-p100
# export GPU_COUNT=1
# export MACHINE_TYPE=n1-highmem-16
#
#
# # Node pool creation may take several minutes.
# gcloud container node-pools create ${GPU_POOL_NAME} \
# --accelerator type=${GPU_TYPE},count=${GPU_COUNT} \
# --zone ${CLUSTER_ZONE} --cluster ${CLUSTER_NAME} \
# --num-nodes=0 --machine-type=${MACHINE_TYPE} --min-nodes=0 --max-nodes=5 --enable-autoscaling \
# --scopes=cloud-platform
# ```
#
# Here in this sample, we specified **--scopes=cloud-platform**. More info is [here](https://cloud.google.com/sdk/gcloud/reference/container/node-pools/create#--scopes). This scope will allow node pool jobs to use the GCE Default Service Account to access GCP APIs (like GCS, etc.). You can also use [Workload Identity](https://cloud.google.com/kubernetes-engine/docs/how-to/workload-identity) or [Application Default Credentials](https://cloud.google.com/docs/authentication/production) to replace **--scopes=cloud-platform**.
#
# ## Install NVIDIA device driver to the cluster
#
# After adding GPU nodes to your cluster, you need to install NVIDIA’s device drivers to the nodes. Google provides a GKE `DaemonSet` that automatically installs the drivers for you.
#
# To deploy the installation DaemonSet, run the following command. You can run this command any time (even before you create your node pool), and you only need to do this once per cluster.
#
# ```shell
# kubectl apply -f https://raw.githubusercontent.com/GoogleCloudPlatform/container-engine-accelerators/master/nvidia-driver-installer/cos/daemonset-preloaded.yaml
# ```
#
# # Consume GPU via Kubeflow Pipelines SDK
#
# Once your cluster is set up to support GPUs, the next step is to indicate which steps in your pipelines should use accelerators, and what type they should use.
# Here is a [document](https://www.kubeflow.org/docs/gke/pipelines/enable-gpu-and-tpu/) that describes the options.
#
# The following is an example 'smoke test' pipeline, to see if your cluster setup is working properly.
#
# +
import kfp
from kfp import dsl
def gpu_smoking_check_op():
return dsl.ContainerOp(
name='check',
image='tensorflow/tensorflow:latest-gpu',
command=['sh', '-c'],
arguments=['nvidia-smi']
).set_gpu_limit(1)
@dsl.pipeline(
name='GPU smoke check',
description='smoke check as to whether GPU env is ready.'
)
def gpu_pipeline():
gpu_smoking_check = gpu_smoking_check_op()
if __name__ == '__main__':
kfp.compiler.Compiler().compile(gpu_pipeline, 'gpu_smoking_check.yaml')
# -
# You may see a warning message from Kubeflow Pipeline logs saying "Insufficient nvidia.com/gpu". If so, this probably means that your GPU-enabled node is still spinning up; please wait for few minutes. You can check the current nodes in your cluster like this:
#
# ```
# kubectl get nodes -o wide
# ```
#
# If everything runs as expected, the `nvidia-smi` command should list the CUDA version, GPU type, usage, etc. (See the logs panel in the pipeline UI to view output).
#
# > You may also notice that after the pipeline step's GKE pod has finished, the new GPU cluster node is still there. GKE autoscale algorithm will free that node if no usage for certain time. More info is [here](https://cloud.google.com/kubernetes-engine/docs/concepts/cluster-autoscaler).
# # Multiple GPUs pool in one cluster
#
# It's possible you want more than one type of GPU to be supported in one cluster.
#
# - There are several types of GPUs.
# - Certain regions often support a only subset of the GPUs ([document](https://cloud.google.com/compute/docs/gpus#gpus-list)).
#
# Since we can set `--num-nodes=0` for certain GPU node pool to save costs if no workload, we can create multiple node pools for different types of GPUs.
#
# ## Add additional GPU nodes to your cluster
#
#
# In a previous section, we added a node pool for P100s. Here we add another pool for V100s.
#
# ```shell
# # You may customize these parameters.
# export GPU_POOL_NAME=v100pool
# export CLUSTER_NAME=existingClusterName
# export CLUSTER_ZONE=us-west1-a
# export GPU_TYPE=nvidia-tesla-v100
# export GPU_COUNT=1
# export MACHINE_TYPE=n1-highmem-8
#
#
# # Node pool creation may take several minutes.
# gcloud container node-pools create ${GPU_POOL_NAME} \
# --accelerator type=${GPU_TYPE},count=${GPU_COUNT} \
# --zone ${CLUSTER_ZONE} --cluster ${CLUSTER_NAME} \
# --num-nodes=0 --machine-type=${MACHINE_TYPE} --min-nodes=0 --max-nodes=5 --enable-autoscaling
# ```
#
# ## Consume certain GPU via Kubeflow Pipelines SDK
#
# If your cluster has multiple GPU node pools, you can explicitly specify that a given pipeline step should use a particular type of accelerator.
# This example shows how to use P100s for one pipeline step, and V100s for another.
# +
import kfp
from kfp import dsl
def gpu_p100_op():
return dsl.ContainerOp(
name='check_p100',
image='tensorflow/tensorflow:latest-gpu',
command=['sh', '-c'],
arguments=['nvidia-smi']
).add_node_selector_constraint('cloud.google.com/gke-accelerator', 'nvidia-tesla-p100').container.set_gpu_limit(1)
def gpu_v100_op():
return dsl.ContainerOp(
name='check_v100',
image='tensorflow/tensorflow:latest-gpu',
command=['sh', '-c'],
arguments=['nvidia-smi']
).add_node_selector_constraint('cloud.google.com/gke-accelerator', 'nvidia-tesla-v100').container.set_gpu_limit(1)
@dsl.pipeline(
name='GPU smoke check',
description='Smoke check as to whether GPU env is ready.'
)
def gpu_pipeline():
gpu_p100 = gpu_p100_op()
gpu_v100 = gpu_v100_op()
if __name__ == '__main__':
kfp.compiler.Compiler().compile(gpu_pipeline, 'gpu_smoking_check.yaml')
# -
# You should see different "nvidia-smi" logs from the two pipeline steps.
# ## Using Preemptible GPUs
#
# A [Preemptible GPU resource](https://cloud.google.com/compute/docs/instances/preemptible#preemptible_with_gpu) is cheaper, but use of these instances means that a pipeline step has the potential to be aborted and then retried. This means that pipeline steps used with preemptible instances must be idempotent (the step gives the same results if run again), or creates some kind of checkpoint so that it can pick up where it left off. To use preemptible GPUs, create a node pool as follows. Then when specifying a pipeline, you can indicate use of a preemptible node pool for a step.
#
# The only difference in the following node-pool creation example is that the **--preemptible** and **--node-taints=preemptible=true:NoSchedule** parameters have been added.
#
# ```
# export GPU_POOL_NAME=v100pool-preemptible
# export CLUSTER_NAME=existingClusterName
# export CLUSTER_ZONE=us-west1-a
# export GPU_TYPE=nvidia-tesla-v100
# export GPU_COUNT=1
# export MACHINE_TYPE=n1-highmem-8
#
# gcloud container node-pools create ${GPU_POOL_NAME} \
# --accelerator type=${GPU_TYPE},count=${GPU_COUNT} \
# --zone ${CLUSTER_ZONE} --cluster ${CLUSTER_NAME} \
# --preemptible \
# --node-taints=preemptible=true:NoSchedule \
# --num-nodes=0 --machine-type=${MACHINE_TYPE} --min-nodes=0 --max-nodes=5 --enable-autoscaling
# ```
#
# Then, you can define a pipeline as follows (note the use of `use_preemptible_nodepool()`).
# +
import kfp
import kfp.gcp as gcp
from kfp import dsl
def gpu_p100_op():
return dsl.ContainerOp(
name='check_p100',
image='tensorflow/tensorflow:latest-gpu',
command=['sh', '-c'],
arguments=['nvidia-smi']
).add_node_selector_constraint('cloud.google.com/gke-accelerator', 'nvidia-tesla-p100').container.set_gpu_limit(1)
def gpu_v100_op():
return dsl.ContainerOp(
name='check_v100',
image='tensorflow/tensorflow:latest-gpu',
command=['sh', '-c'],
arguments=['nvidia-smi']
).add_node_selector_constraint('cloud.google.com/gke-accelerator', 'nvidia-tesla-v100').container.set_gpu_limit(1)
def gpu_v100_preemptible_op():
v100_op = dsl.ContainerOp(
name='check_v100_preemptible',
image='tensorflow/tensorflow:latest-gpu',
command=['sh', '-c'],
arguments=['nvidia-smi'])
v100_op.container.set_gpu_limit(1)
v100_op.add_node_selector_constraint('cloud.google.com/gke-accelerator', 'nvidia-tesla-v100')
v100_op.apply(gcp.use_preemptible_nodepool(hard_constraint=True))
return v100_op
@dsl.pipeline(
name='GPU smoking check',
description='Smoking check whether GPU env is ready.'
)
def gpu_pipeline():
gpu_p100 = gpu_p100_op()
gpu_v100 = gpu_v100_op()
gpu_v100_preemptible = gpu_v100_preemptible_op()
if __name__ == '__main__':
kfp.compiler.Compiler().compile(gpu_pipeline, 'gpu_smoking_check.yaml')
# -
# # TPU
# Google's TPU is awesome. It's faster and lower TOC. To consume TPUs, there is no need to create a node-pool; just call KFP SDK to use it. Here is a [doc](https://www.kubeflow.org/docs/gke/pipelines/enable-gpu-and-tpu/#configure-containerop-to-consume-tpus). Note that not all regions have TPU yet.
#
#
| samples/tutorials/gpu/gpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn import metrics
# read in the iris data
iris = load_iris()
# create X (features) and y (response)
X = iris.data
Y = iris.target
print(type(X))
print(f"X = {X}")
# -
print(f'{len(X[0])}')
# +
len_x = len(X)
len_y = len(Y)
print(f'len_x = {len_x}')
print(f'len_y = {len_y}')
# +
# use train/test split with different random_state values# use t
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, random_state=4)
len_x_train = len(X_train)
len_x_test = len(X_test)
len_y_train = len(Y_train)
len_y_test = len(Y_test)
print(f'len_x_train = {len_x_train}')
print(f'len_x_test = {len_x_test}')
print(f'len_y_train = {len_y_train}')
print(f'len_y_trst = {len_y_test}')
print(X_train)
# +
print(Y_train)
one_count = list(Y_train).count(1)
two_count = list(Y_train).count(2)
zero_count = list(Y_train).count(0)
print(f'1 num = {one_count}')
print(f'2 num = {two_count}')
print(f'0 num = {zero_count}')
'''
Y_train[Y_train==2] = 0
print(Y_train)
one_count = list(Y_train).count(1)
two_count = list(Y_train).count(2)
zero_count = list(Y_train).count(0)
print(f'1 num = {one_count}')
print(f'2 num = {two_count}')
print(f'0 num = {zero_count}')
'''
# -
# check classification accuracy of KNN with K=5
knn = KNeighborsClassifier(n_neighbors=5)
knn.fit(X_train, Y_train)
Y_pred = knn.predict(X_test)
print(metrics.accuracy_score(Y_test, Y_pred))
# +
# simulate splitting a dataset of 25 observations into 5 folds
#from sklearn.model_selection import KFold
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import KFold
skf = StratifiedKFold(n_splits=5, random_state=25, shuffle=False)
#skf = KFold(n_splits=5, random_state=25, shuffle=False)
# print the contents of each training and testing set
print('{} {:^61} {}'.format('Iteration', 'Training set observations', 'Testing set observations'))
iteration = 0
for train_index, test_index in skf.split(X_train, Y_train):
print('{} {} {}'.format(iteration, train_index, test_index))
list_test = [Y_train[x] for x in test_index]
one_count = list_test.count(1)
two_count = list_test.count(2)
zero_count = list_test.count(0)
print(f'list_test = {list_test}')
print(f'1 num = {one_count}')
print(f'2 num = {two_count}')
print(f'0 num = {zero_count}')
iteration += 1
#for iteration, data in enumerate(kf, start=1):
# print('{:^9} {} {:^25}'.format(iteration, data[0], data[1]))
# -
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
from sklearn.metrics import classification_report
# Set the parameters by cross-validation
tuned_parameters = [{'kernel': ['rbf'], 'gamma': [1e-3, 1e-4],
'C': [1, 10, 100, 1000]},
{'kernel': ['linear'], 'C': [1, 10, 100, 1000]}]
# +
#scores = ['accuracy', 'precision_macro', 'recall_macro', 'f1_macro']
scores = ['precision_macro', 'recall_macro', 'accuracy', 'f1_macro']
for score in scores:
print("# Tuning hyper-parameters for %s" % score)
print()
skf = StratifiedKFold(n_splits=10, random_state=25, shuffle=True)
clf = GridSearchCV(SVC(), tuned_parameters, cv=skf, scoring = score)
clf.fit(X_train, Y_train)
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print()
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
Y_true, Y_pred = Y_test, clf.predict(X_test)
print(classification_report(Y_true, Y_pred))
print()
# +
import numpy as np
from sklearn.model_selection import StratifiedKFold
X = np.array([[1, 2], [3, 4], [1, 2], [3, 4]])
y = np.array([0, 0, 1, 1])
skf = StratifiedKFold(n_splits=2)
skf.get_n_splits(X, y)
print(skf)
StratifiedKFold(n_splits=2, random_state=None, shuffle=False)
for train_index, test_index in skf.split(X, y):
print("train_index = ", train_index, "\ntest_index = ", test_index)
print(f'type(train_index) = {type(train_index)}')
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
print(f"TRAIN_X_VAL = {X_train}")
print(f"TRAIN_Y_VAL = {y_train}")
print(f"TEST_X_VAL = {X_test}")
print(f"TEST_Y_VAL = {y_test}")
print('------')
# -
index = np.array([1, 3])
print(type(index))
#X[index]
| Cross_Validation_Grid_Search_StratifiedFold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Packages
# +
import numpy as np
import sys
import os
import pandas as pd
import boto3
# Data storage
from sqlalchemy import create_engine # SQL helper
import psycopg2 as psql # PostgreSQL DBs
sys.path.append("..")
# -
import keys
import data_fcns as dfc
# ## Options
# So that items in pandas columns don't show truncated values
pd.set_option('display.max_colwidth', -1)
# ## Create Official item factors matrix or dataframe
item_factors_df = pd.read_pickle('support_data/item_factors.pkl')
item_factors_df.head()
# ## Get comics data from DB
# +
# Define path to secret
secret_path_aws = os.path.join(os.environ['HOME'], '.secret',
'aws_ps_flatiron.json')
secret_path_aws
aws_keys = keys.get_keys(secret_path_aws)
user = aws_keys['user']
ps = aws_keys['password']
host = aws_keys['host']
db = aws_keys['db_name']
aws_ps_engine = ('postgresql://' + user + ':' + ps + '@' + host + '/' + db)
# Setup PSQL connection
conn = psql.connect(
database=db,
user=user,
password=ps,
host=host,
port='5432'
)
# -
# Instantiate cursor
cur = conn.cursor()
# +
# Count records.
query = """
SELECT * FROM comics
"""
# Execute the query
cur.execute(query)
conn.commit()
# -
# Check results
comics_pdf = pd.DataFrame(cur.fetchall())
comics_pdf.columns = [col.name for col in cur.description]
comics_pdf.head()
# ## Merge item factors and comics
# Change column names for item_factors_df
item_factors_df.columns = ['comic_id', 'features']
combo = item_factors_df.merge(comics_pdf, left_on='comic_id', right_on='comic_id', how='inner', )
combo.set_index(['comic_id'], inplace=True)
combo.head()
comics_pdf.loc[comics_pdf['comic_id']==20]
combo.shape
combo.drop(['img_url'], axis=1, inplace=True)
combo.shape
combo.head()
combo.columns = ['features', 'comic_title', 'img_url']
combo.head()
# ## Create pickle file
combo.to_pickle('support_data/comics_factors_201908.pkl')
# # Test 2019.08.12 fix
# +
# Read new pickle
# -
cf_new = pd.read_pickle('support_data/comics_factors_201908.pkl')
# +
# Read old pickle
# -
cf_old = pd.read_pickle('support_data/comics_factors.pkl')
cf_new.shape
cf_old.shape
cf_new.head()
cf_old.head()
| comrx/dev/deprecated/comics_rx-03c_build_comics_factors_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Short Bursts
#
# We look at the behavior of using short bursts to increase number of opportunity districts for bvap and it projection into the line model
import matplotlib.pyplot as plt
from gerrychain import (GeographicPartition, Partition, Graph, MarkovChain,
proposals, updaters, constraints, accept, Election)
from gerrychain.proposals import recom, propose_random_flip
from gerrychain.tree import recursive_tree_part
from gerrychain.metrics import mean_median, efficiency_gap, polsby_popper, partisan_gini
from functools import (partial, reduce)
import pandas
import geopandas as gp
import numpy as np
import networkx as nx
import pickle
import seaborn as sns
import pprint
import operator
import scipy
from mpl_toolkits.mplot3d import Axes3D
from sklearn.decomposition import PCA
from sklearn.preprocessing import scale, normalize
import random
from nltk.util import bigrams
from nltk.probability import FreqDist
import pylab
## This function takes a name of a shapefile and returns a tuple of the graph
## and its associated dataframe
def build_graph(filename):
print("Pulling in Graph from Shapefile: " + filename)
graph = Graph.from_file(filename)
df = gp.read_file(filename)
return(graph, df)
def config_markov_chain(initial_part, iters=1000, epsilon=0.05, compactness=True, pop="TOT_POP"):
# The recom proposal needs to know the ideal population for the districts so
# that we can improve speed by bailing early on unbalanced partitions.
ideal_population = sum(initial_part["population"].values()) / len(initial_part)
# We use functools.partial to bind the extra parameters (pop_col, pop_target,
# epsilon, node_repeats) of the recom proposal.
proposal = partial(recom,
pop_col=pop,
pop_target=ideal_population,
epsilon=epsilon,
node_repeats=1)
# To keep districts about as compact as the original plan, we bound the number
# of cut edges at 2 times the number of cut edges in the initial plan.
if compactness:
compactness_bound = constraints.UpperBound(lambda p: len(p["cut_edges"]),
2*len(initial_part["cut_edges"]))
cs = [constraints.within_percent_of_ideal_population(initial_part, epsilon), compactness_bound]
else:
cs = [constraints.within_percent_of_ideal_population(initial_part, epsilon)]
# Configure the MarkovChain.
return MarkovChain(proposal=proposal, constraints=cs, accept=accept.always_accept,
initial_state=initial_part, total_steps=iters)
# ## Functions to Simulate Short Bursts
## This function takes a partition and returns the number o
def num_opportunity_dists(part, minority_prec="bvap_prec"):
dist_precs = part[minority_prec].values()
return sum(list(map(lambda v: v >= 0.35, dist_precs)))
def short_burst_run(starting_part, score_func, num_bursts=10, num_steps=100, verbose=False,
pop="TOT_POP"):
max_part = (starting_part, num_opportunity_dists(starting_part))
observed_num_ops = np.zeros((num_bursts, num_steps))
for i in range(num_bursts):
if verbose: print("Burst:", i)
chain = config_markov_chain(max_part[0], iters=num_steps, epsilon=0.1, pop=pop)
for j, part in enumerate(chain):
part_score = num_opportunity_dists(part)
observed_num_ops[i][j] = part_score
max_part = (part, part_score) if part_score >= max_part[1] else max_part
return (max_part, observed_num_ops)
# ## Pull in PA data and set up partitions
# graph_PA, df_PA = build_graph("PA_shape/PA_VTD.shp")
graph_PA = pickle.load(open("PA_graph.p", "rb"))
df_PA = pickle.load(open("PA_df.p", "rb"))
PA_updaters = {"population": updaters.Tally("TOT_POP", alias="population"),
"bvap": updaters.Tally("BLACK_POP", alias="bvap"),
"vap": updaters.Tally("VAP", alias="vap"),
"bvap_prec": lambda part: {k: part["bvap"][k] / part["population"][k] for k in part["bvap"]}}
total_population_PA = sum(df_PA.TOT_POP.values)
ideal_population_PA = total_population_PA / 50
seed_part_senate = recursive_tree_part(graph_PA, range(50), pop_col="TOT_POP",
pop_target=ideal_population_PA,
epsilon=0.01, node_repeats=1)
PA_seed_seante = GeographicPartition(graph_PA, assignment=seed_part_senate,updaters=PA_updaters)
seed_part_house = recursive_tree_part(graph_PA, range(203), pop_col="TOT_POP",
pop_target=ideal_population_PA,
epsilon=0.01, node_repeats=1)
PA_seed_house = GeographicPartition(graph_PA, assignment=seed_part_house,updaters=PA_updaters)
enacted_senate = GeographicPartition(graph_PA, assignment="SSD", updaters=PA_updaters)
num_opportunity_dists(enacted_senate)
# ## Pull in AR data and set up partitions
graph_AR, df_AR = build_graph("AR_shape/AR.shp")
AR_updaters = {"population": updaters.Tally("TOTPOP", alias="population"),
"bvap": updaters.Tally("BVAP", alias="bvap"),
"vap": updaters.Tally("VAP", alias="vap"),
"bvap_prec": lambda part: {k: part["bvap"][k] / part["vap"][k]
for k in part["bvap"]}}
AR_enacted_senate = GeographicPartition(graph_AR, assignment="SSD", updaters=AR_updaters)
num_opportunity_dists(AR_enacted_senate)
total_population_AR = sum(df_AR.TOTPOP.values)
ideal_population_AR = total_population_AR / 35
senate_seed = recursive_tree_part(graph_AR, range(35), pop_col="TOTPOP",
pop_target=ideal_population_AR,
epsilon=0.01, node_repeats=1)
AR_seed_senate = GeographicPartition(graph_AR, assignment=senate_seed,updaters=AR_updaters)
num_opportunity_dists(AR_seed_senate)
house_seed = recursive_tree_part(graph_AR, range(100), pop_col="TOTPOP",
pop_target=ideal_population_AR,
epsilon=0.01, node_repeats=1)
AR_seed_house = GeographicPartition(graph_AR, assignment=house_seed,updaters=AR_updaters)
# ### How does burst length affect findings
# +
plt.figure(figsize=(10,8))
plt.xlim(-.5, 8)
plt.xlabel("Number of opportunity districts")
plt.ylabel("Steps")
plt.title("PA short bursts of different lengths")
total_steps = 1000
for color, len_burst in [("k", 1), ("b", 5), ("r", 25), ("g", 50), ("y", 100),
("cyan", 1000)]:
print(len_burst)
num_bursts = int(total_steps / len_burst)
_, observations = short_burst_run(enacted_senate, num_opportunity_dists,
num_bursts=num_bursts, num_steps=len_burst)
for i in range(num_bursts):
plt.plot(observations[i], range(len_burst*i, len_burst*(i+1)),
color=color, alpha=0.5, marker=".", markevery=[0,-1])
plt.plot([], color=color, label=("Burst_len " + str(len_burst)))
plt.legend()
plt.show()
# +
plt.figure(figsize=(10,8))
plt.xlim(-.5, 8)
plt.xlabel("Number of opportunity districts")
plt.ylabel("Steps")
plt.title("AR short bursts of different lengths")
total_steps = 1000
for color, len_burst in [("k", 1), ("b", 5), ("r", 25), ("g", 50), ("y", 100),
("cyan", 1000)]:
print(len_burst)
num_bursts = int(total_steps / len_burst)
_, observations = short_burst_run(AR_seed_senate, num_opportunity_dists,
num_bursts=num_bursts, num_steps=len_burst, pop="TOTPOP")
for i in range(num_bursts):
plt.plot(observations[i], range(len_burst*i, len_burst*(i+1)),
color=color, alpha=0.5, marker=".", markevery=[0,-1])
plt.plot([], color=color, label=("Burst_len " + str(len_burst)))
plt.legend()
plt.show()
# -
# ## Run on PA state senate and visualizing results
num_bursts = 50
num_steps = 10
max_part, observed_num_ops = short_burst_run(enacted_senate, num_opportunity_dists,
num_bursts=num_bursts, num_steps=num_steps)
_, observed_num_ops_flat = short_burst_run(enacted_senate, num_opportunity_dists,
num_bursts=1, num_steps=500)
plt.hist(observed_num_ops.flatten())
plt.hist(observed_num_ops_flat.flatten())
for i, row in enumerate(observed_num_ops):
plt.figure()
plt.title("Observations at burst " + str(i + 1))
plt.xlabel("Number of opportunity districts")
plt.ylabel("Frequency")
plt.xlim(0,10)
bins = np.linspace(0, 10, 11)
sns.distplot(row, kde=False, bins=bins)
#plt.savefig("plots/PA_senate_10_100_seed_3/short_burst_" + str(i+1) + "_hist.png")
names = list(map(lambda i: "Burst " + str(i), range(1,num_bursts + 1)))
plt.figure(figsize=(10,8))
plt.xlim(-.5, 8)
plt.xlabel("Number of opportunity districts")
plt.ylabel("Steps")
for i in range(num_bursts):
plt.plot(observed_num_ops[i], range(num_steps*i, num_steps*(i+1)))#, label=names[i])
# plt.legend()
# plt.savefig("plots/burst_walk_100_100.png")
# +
plt.figure(figsize=(10,8))
colors = ["red", "orange", "yellow", "lime", "green",
"cyan", "blue", "indigo", "purple", "fuchsia"]
names = list(map(lambda i: "Burst " + str(i), range(1,num_bursts + 1)))
plt.hist(list(observed_num_ops), color=colors, label=names, align='mid', rwidth=10)
plt.xlim(1,7)
plt.legend()
plt.show()
# -
# ## Reprojecting bursts onto the line
# ## AR
_, observed_num_ops_AR = short_burst_run(AR_seed_senate, num_opportunity_dists,
num_bursts=1000, num_steps=10, pop="TOTPOP")
transitions_AR = []
for row in observed_num_ops_AR:
transitions_AR.extend(bigrams(row))
fdist_AR = FreqDist(transitions_AR)
probs = {}
N = np.matrix(observed_num_ops_AR).astype(int)
dim = N.max()
A = np.zeros((dim, dim))
for k, v in fdist_AR.items():
p = v / fdist_AR.N()
probs[k] = p
A[int(k[0]) - 1][int(k[1]) - 1] = p
A_normed = normalize(A, norm="l1")
G = nx.from_numpy_array(A_normed, create_using=nx.DiGraph)
mapping = {n: n+1 for n in G.nodes}
G1 = nx.relabel_nodes(G, mapping)
AR_line_proj=dict([((u,v,), round(d['weight'],4))
for u,v,d in G1.edges(data=True)])
AR_line_proj
# ## PA
transitions = []
for row in observed_num_ops:
transitions.extend(bigrams(row))
fdist = FreqDist(transitions)
fdist
probs = {}
N = np.matrix(observed_num_ops).astype(int)
dim = N.max()
A = np.zeros((dim, dim))
for k, v in fdist.items():
p = v / fdist.N()
probs[k] = p
A[int(k[0]) - 1][int(k[1]) - 1] = p
A_normed = normalize(A, norm="l1")
A_normed[2]
G = nx.from_numpy_array(A_normed, create_using=nx.DiGraph)
G.edges(data=True)
mapping = {n: n+1 for n in G.nodes}
G1 = nx.relabel_nodes(G, mapping)
PA_line_proj=dict([((u,v,), round(d['weight'],4))
for u,v,d in G1.edges(data=True)])
PA_line_proj
plt.figure()
pos=nx.shell_layout(G1)
nx.draw(G1, pos=pos, edge_cmap=plt.cm.Reds)
nx.draw_networkx_edge_labels(G1,pos=pos,edge_labels=edge_labels)
plt.show()
H = nx.DiGraph()
for (n1,n2) in probs:
H.add_edges_from([(n1, n2)], weight=probs[n1,n2])
edge_labels=dict([((u,v,), round(d['weight'],4))
for u,v,d in H.edges(data=True)])
edge_labels
plt.figure()
pos=nx.shell_layout(H)
nx.draw(H, pos=pos, edge_cmap=plt.cm.Reds)
nx.draw_networkx_edge_labels(H,pos=pos,edge_labels=edge_labels)
plt.show()
from networkx.drawing.nx_agraph import write_dot
plt.figure(figsize=(10,10))
nx.draw_networkx(
G1, pos=nx.circular_layout(G1),
node_shape='o', node_size=1000, font_size=10,
edge_color='#555555', width=3.0
)
nx.draw_networkx_edge_labels(
G1, pos=nx.circular_layout(G1),
edge_labels=edge_labels,
font_size=10, label_pos=0.25, rotate=False
)
# plt.axis('off')
write_dot(G,'graph.dot')
# +
from __future__ import division
import matplotlib as mpl
G = H
pos = nx.layout.spring_layout(G)
# node_sizes = [3 + 10 * i for i in range(len(G))]
M = G.number_of_edges()
edge_colors = range(2, M + 2)
edge_alphas = [round(d['weight'],4)*2 for u,v,d in H.edges(data=True)]
nodes = nx.draw_networkx_nodes(G, pos, node_color='blue')
edges = nx.draw_networkx_edges(G, pos, arrowstyle='->',
arrowsize=10, edge_color=edge_colors,
edge_cmap=plt.cm.Blues, width=2)
# set alpha value for each edge
for i in range(M):
edges[i].set_alpha(edge_alphas[i])
pc = mpl.collections.PatchCollection(edges, cmap=plt.cm.Blues)
pc.set_array(edge_colors)
plt.colorbar(pc)
ax = plt.gca()
ax.set_axis_off()
plt.show()
# -
| misc/simple_random_walks_and_short_bursts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# <font size="+5">Flexible Thinking</font>
#
# # The challenge
# -
# <div class="alert alert-success">
# 3 different lines of code to get the same result
# </div>
# # The covered solution
import matplotlib.pyplot as plt
import seaborn as sns
df = sns.load_dataset('car_crashes')
df.head()
# +
# code 1 ??
# -
plt.scatter(x=df.alcohol, y=df.total);
# +
# code 2 ??
# -
plt.scatter(x=df['alcohol'], y=df['total']);
# +
# code 3 ??
# -
plt.scatter(x='alcohol', y='total', data=df);
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# # What will we learn?
#
# - Fundamental Teaching 1
# - Fundamental Teaching 2
# -
# # Which concepts will we use?
#
# - Fundamental Concept 1
# - Fundamental Concept 2
# # Requirements?
#
# - Mandatory Requirement 1
# - Mandatory Requirement 2
# + [markdown] tags=[]
# # The starting *thing*
# -
# # Syllabus for the Material
# # The uncovered solution
# # References
#
# > Links gathered while developing the solution
#
# - Link Reference 1
# - Link Reference 2
| 00_Introduction Motivation & Philosophy/01 Flexible Thinking/00 syllabus template recursive learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import sqlite3
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# -
conn = sqlite3.connect("/data/movie-lens.db")
movies = pd.read_sql("select * from movies", conn)
movies.head()
ratings = pd.read_sql("select * from ratings", conn)
ratings.head()
ratings.rating.value_counts().sort_index().plot.bar()
plt.title("Frequency plot for rating")
plt.xlabel("Rating")
plt.ylabel("Frequency")
plt.savefig("/tmp/rating.jpeg")
agg = ratings.groupby("movieId").rating.agg([np.mean, len])
agg.head()
agg.to_sql("rating_avg", conn, if_exists="append") # Save the dataframe to database
| Appendix B - Working with database.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Activity 1.02: Forest Fire Size and Temperature Analysis
# In this activity, we will use pandas features to derive some insights from a forest fire dataset. We will get the mean size of forest fires, what the largest recorded fire in our dataset is, and whether the amount of forest fires grows proportionally to the temperature in each month.
# #### Loading the dataset
# importing the necessary dependencies
import pandas as pd
# loading the Dataset
dataset = pd.read_csv('../../Datasets/forestfires.csv')
# looking at the first two rows of the dataset
dataset[0:2]
# The dataset contains:
# - X - x-axis spatial coordinate within the Montesinho park map: 1 to 9
# - Y - y-axis spatial coordinate within the Montesinho park map: 2 to 9
# - **month - month of the year: 'jan' to 'dec'**
# - day - day of the week: 'mon' to 'sun'
# - FFMC - FFMC index from the FWI system: 18.7 to 96.20
# - DMC - DMC index from the FWI system: 1.1 to 291.3
# - DC - DC index from the FWI system: 7.9 to 860.6
# - ISI - ISI index from the FWI system: 0.0 to 56.10
# - **temp - temperature in Celsius degrees: 2.2 to 33.30**
# - RH - relative humidity in %: 15.0 to 100
# - wind - wind speed in km/h: 0.40 to 9.40
# - rain - outside rain in mm/m2 : 0.0 to 6.4
# - **area - the burned area of the forest (in ha): 0.00 to 1090.84**
#
#
# **Note:**
# The fields that we'll be working with are highlighted in the listing.
# ---
# #### Getting insights into the sizes of forest fires.
#
# When looking at the first two rows of our dataset we can already see that it contains entries in which the area is 0.
# For this first task we only care about fires that have an area of more than 0.
#
# Create a new dataset that only contains the entries with an area value of > 0.
# +
# filter the dataset for rows that have an area > 0
area_dataset= dataset[dataset["area"] > 0]
area_dataset
# -
# After filtering out the zero area entries, we can simply use the `mean` method of pandas to get the mean area size of the forest fires for the filtered down dataset not containing zero area sizes.
#
# Get the mean value for the `area` column of out filtered dataset.
# get the mean value for the area column
area_dataset["area"].mean()
# In addition to that, looking at the largest and smallest non-zero area can help us understand the range of possible area sizes.
# Let's get more insights into that.
#
# - Use the `min` and `max` methods to see the smallest and largest area that has been affected by a forest fire.
# - Use the `std` method to get insights into how much variation there is in our dataset.
# get the smallest area value from our dataset
area_dataset["area"].min()
# get the largest area value from our dataset
area_dataset["area"].max()
# get the standard deviation of values in our dataset
area_dataset["area"].std()
# The largest value is much larger than our mean.
# The standard deviation also is quite large which indicates that the difference between our mean and the "middle value" will be quite high.
#
# Let's look at the last 20 values of our sorted dataset to see if we have more than one very large value.
# Sort the filtered dataset by the `area` column and output the last 20 entries from it.
# sorting the filtered dataset and printing the last 20 elements
area_dataset.sort_values(by=["area"]).tail(20)
# As we can see here, only 11 out of the 270 rows contain values that are larger than 100.
# After 20 values we are close to the area value of 60.
#
# Let's imagine our dataset contained only 1 or 2 values that were much higher than the other ones, e.g. an area size value of 10254.91. Simply by observing the dataset, this feels like there might have been an error on adding this to the dataset.
# In a smaller dataset, the mean value would get heavily distored by this one entry. A more stable value to use in such a case is the median value of the dataset.
#
# Get the median value for the ´area´ column.
# calculate the median value for the area column
area_dataset["area"].median()
# **Note:**
# Remember that the median is not the same as then mean of your dataset. While the median is simple the "value in the middle", the mean is much more prone to distortion by outliers.
# ---
# #### Finding the month with the most forest fires
# In this second task we want to quickly see which months have the most forest fires and whether or not the temperature has a direct connection to it.
#
# Get a list of month values that are present in our dataset.
# +
# get a list of month values from the dataset
months = dataset["month"].unique()
months
# -
# In addition to the unique values we also want use the shape element of our dataset to determine how many rows it has.
#
# Filter the dataset for only rows that contain the month `mar` and print the number of rows using `shape`.
# get the number of forest fires for the month of march
dataset[dataset["month"] == "mar"].shape[0]
# The last step to fulfil the task is to iterate over all months, filtering our dataset for the rows containing the given month and calculating the mean temperature.
#
# - Iterate over the months from the unique list we created
# - Filter our dataset for the rows containing the given month
# - Get the number of rows from `shape`
# - Get the mean temperature for the given month
# - Print a statement with the number of fires, mean temperature and the month
# iterate over the months list
# get number of forest fires for each month
# get mean temperature for each month
# print out number of fires and mean temperature
for month in months:
month_dataset = dataset[dataset["month"] == month]
fires_in_month = month_dataset.shape[0]
avg_tmp_in_month = int(month_dataset["temp"].mean())
print(str(fires_in_month) + " fires in " + month + " with a mean temperature of ~" + str(avg_tmp_in_month) + "°C")
| Chapter01/Activity1.02/Activity1.02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from pandas import Series,DataFrame
from numpy.random import randn
# stats
from scipy import stats
#plot
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
data1 = randn(100)
plt.hist(data1)
data2 = randn(100)
plt.hist(data2, color = 'indianred')
plt.hist(data1,normed=True, color = 'indianred', alpha = 0.5, bins=10)
plt.hist(data2,normed=True, alpha = 0.5, bins=10)
sns.jointplot(data1,data2)
sns.jointplot(data1,data2,kind= 'hex')
sns.regplot(data1,data2)
sns.rugplot(data1)
sns.distplot(data1,kde_kws={'color':'red','label':'HIST'})
ser1 = Series(data1,name= 'myname')
sns.distplot(ser1,bins = 25)
sns.boxplot(ser1)
sns.violinplot(ser1)
df = pd.DataFrame(np.random.randint(0,1000,size=(100, 2)), columns=list('AB'))
df
sns.lmplot('A','B',df,order=4,
scatter_kws = {'marker':'o','color': 'indianred'},
line_kws = {'linewidth':1,'color':'green' })
sns.heatmap(df)
sns.clustermap(df)
df.head()
| plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Get csv ready
# +
import pandas as pd
import numpy as np
import os
house_data_path = os.path.join(os.path.pardir,'data','raw','MelbourneHousingSnapshot')
house_csv_path = os.path.join(house_data_path,'melb_data.csv')
house_csv_df = pd.read_csv(house_csv_path)
# -
# 1. Get Info
house_csv_df.info()
# 2. get 2 Rows
house_csv_df.head(2)
house_csv_df.Suburb
house_csv_df.Suburb.unique()
house_csv_df[['Suburb','Type']]
house_csv_df.loc[5:15,]
house_csv_df.loc[5:15,'Suburb':'SellerG']
preston_properties = house_csv_df.loc[house_csv_df.Suburb == 'Preston',]
preston_properties
print('Number of sold houses in Preston ', len(preston_properties))
preston_townhouses = house_csv_df.loc[(house_csv_df.Suburb == 'Preston') & (house_csv_df.Type =='t'),]
preston_townhouses
house_1m_df = house_csv_df.loc[(house_csv_df.Price > 1000000),['Address','Rooms']]
house_1m_df
house_csv_df.Price.describe()
| notebooks/data_visualizations_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import tensorflow as tf
sess = tf.InteractiveSession()
x = tf.placeholder(tf.float32,shape=[None,784])
y_ = tf.placeholder(tf.float32,shape=[None,10])
W = tf.Variable(tf.zeros([784,10]))
b = tf.Variable(tf.zeros([10]))
sess.run(tf.initialize_all_variables())
y = tf.nn.softmax(tf.matmul(x,W)+b)
cross_entropy = -tf.reduce_sum(y_*tf.log(y))
Opt = tf.train.GradientDescentOptimizer(0.01)
train_step = Opt.minimize(cross_entropy)
def weight_variable(shape):
initial = tf.truncated_normal(shape,stddev=0.1)
return tf.Variable(initial)
def bias_variable(shape):
initial = tf.truncated_normal(shape,stddev=0.1)
return tf.Variable(initial)
def conv2d(x,W):
return tf.nn.conv2d(x,W,strides=[1,1,1,1],padding="SAME")
def max_pool_2x2(x):
return tf.nn.max_pool(x,ksize=[1,2,2,1],strides=[1,2,2,1],padding="SAME")
W_conv1 = weight_variable([5,5,1,32])
b_conv1 = bias_variable([32])
x_image = tf.reshape(x,[-1,28,28,1])
h_conv1 = tf.nn.relu(conv2d(x_image,W_conv1)+b_conv1)
h_pool1 = max_pool_2x2(h_conv1)
W_conv2 = weight_variable([5,5,32,64])
b_conv2 = bias_variable([64])
h_conv2 = tf.nn.relu(conv2d(h_pool1,W_conv2)+b_conv2)
h_pool2 = max_pool_2x2(h_conv2)
W_fc1 = weight_variable([7*7*64,1024])
b_fc1 = bias_variable([1024])
h_pool2_flat = tf.reshape(h_pool2,[-1,7*7*64])
h_fc1 = tf.nn.relu(tf.matmul(h_pool2_flat,W_fc1)+b_fc1)
keep_prob = tf.placeholder(tf.float32)
h_fc1_drop = tf.nn.dropout(h_fc1,keep_prob)
W_fc2 = weight_variable([1024,10])
b_fc2 = bias_variable([10])
y_conv = tf.nn.softmax(tf.matmul(h_fc1_drop,W_fc2)+b_fc2)
cross_entropy = -tf.reduce_sum(y_*tf.log(y_conv))
train_step = tf.train.GradientDescentOptimizer(1e-4).minimize(cross_entropy)
correct_prediction = tf.equal(tf.argmax(y_conv,1),tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction,tf.float32))
sess.run(tf.initialize_all_variables())
for i in xrange(20000):
batch = minist.train.next_batch(50)
if i%100 == 0:
train_accuracy = accuracy.eval(feed_dict={x:batch[0],y_:batch[1],keep_prob:1.0})
print("step %d, trainning accuracy %g" %(i,train_accuracy))
train_step.run(feed_dict={x:batch[0],y:batch[1],keep_prob:0.5})
print("test accuracy %g" % accuracy.eval(feed_dict={x:mnist.test.images,y_:mnist.test.labels,keep_prob:1.0}))
| learn/tensorflow/public/MNIST_for_experts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import sys
import random
import matplotlib.pyplot as plt
import os
import networkx as nx
import NetworkAlgorithm as na
from networkx.algorithms import approximation as approx
karatepath = "./Data/karate/karate.gml"
powerpath = "./Data/power/power.gml"
dolphinspath = "./Data/dolphins/dolphins.gml"
adjnounpath = "./Data/adjnoun/adjnoun.gml"
# -
def statistic(G):
nodenum = len(G.nodes)
edgenum = len(G.edges)
degree = G.degree()
degree = [i[1] for i in degree]
degreeaver = sum(degree)/len(degree)
averClustering = approx.clustering_coefficient.average_clustering(G)
shortest_aver = nx.algorithms.shortest_paths.generic.average_shortest_path_length(G)
print("node number:", nodenum)
print("edge number", edgenum)
print("average degree:", degreeaver)
print("average clustering coefficient:", averClustering)
print("average shortest path", shortest_aver)
karateG = nx.read_gml(karatepath, label=None)
power = nx.read_gml(powerpath, label=None)
dolphins = nx.read_gml(dolphinspath, label=None)
adjnoun = nx.read_gml(adjnounpath, label=None)
print("karate")
statistic(karateG)
print("dolphins")
statistic(dolphins)
print("adjnoun")
statistic(adjnoun)
print("power")
statistic(power)
# +
# karateG = nx.read_gml(karatepath, label=None)
# print(len(karateG.nodes))
# print(len(karateG.edges))
# degree = karateG.degree()
# degree = [i[1] for i in degree]
# degreeaver = sum(degree)/len(degree)
# print(karateG.degree())
# print(degreeaver)
# averClustering = approx.clustering_coefficient.average_clustering(karateG)
# print(averClustering)
# betweenness = nx.algorithms.centrality.betweenness_centrality(karateG, normalized=True)
# # print(betweenness)
# betweenness = [betweenness[i] for i in betweenness]
# averbetween = sum(betweenness)/len(betweenness)
# print(averbetween)
# shortest = nx.algorithms.shortest_paths.generic.average_shortest_path_length(karateG)
# print("shortest",shortest)
# path = nx.algorithms.shortest_paths.generic.shortest_path_length(karateG)
# pathsum = []
# for i in path:
# path = [i[1][j] for j in i[1]]
# # print(sum(path)/(len(path) -1))
# # print(sum(path))
# pathsum.append(sum(path))
# aversum = sum(pathsum)/len(pathsum)
# print(aversum)
# betweenness = nx.algorithms.centrality.betweenness_centrality(karateG, normalized=False)
# betweenness = [betweenness[i] for i in betweenness]
# averbetween = sum(betweenness)/len(betweenness)
# print(betweenness)
# betentro = na.entropy(betweenness, isNormalization=False)
# print(betentro)
# print(pathsum)
# pathentro = na.entropy(pathsum, isNormalization=False)
# print(pathentro)
# -
| Code/Statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mohan-mj/Random-Forest-Classification/blob/master/Random%20Forest%20Classification-Transfusion.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="LfJaYztaHN-I"
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="w-RDgbLJHN-N" outputId="7111af85-83d6-4499-b019-0cdf78bba358"
# Importing the dataset
dataset = pd.read_csv('https://raw.githubusercontent.com/mohan-mj/Random-Forest-Classification/master/transfusion.csv')
dataset.head()
# + colab={"base_uri": "https://localhost:8080/"} id="IdX8kdciHj8Z" outputId="a700c555-6a2d-4aa8-8d8f-acfbd951a93e"
# dataset info
dataset.info()
# + id="_eRP4eMLHN-P"
# split the data into X and y
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
# + colab={"base_uri": "https://localhost:8080/"} id="cnBscnvqHN-Q" outputId="775eaae4-94f0-4560-d6f7-2aeed260bdc5"
# shape of X
X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="_ecc2KtRHN-R" outputId="4489c99a-222e-405e-b817-3063c49af77c"
# shape of y
y.shape
# + id="cox8-TWOHN-R"
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state = 40)
# + id="5vi31Ro7HN-S"
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="TxsZjAF2HN-T" outputId="455bb8af-516f-48de-9674-109f3a6ba92a"
# Fitting Random Forest Classification to the Training set
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(n_estimators = 30, criterion = 'entropy', random_state = 40)
classifier.fit(X_train, y_train)
# + id="4chPK9xOHN-V"
# Predict
y_pred = classifier.predict(X_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="4j1Wt3o6HN-X" outputId="8268d7ff-36f6-4335-c01d-63856263b6ed"
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_train, y_pred)
sns.heatmap(cm, annot=True, fmt="d")
# + colab={"base_uri": "https://localhost:8080/"} id="hFm8hFm3HN-X" outputId="7be3b4e9-44f8-4f3d-9364-43b1f5c08c18"
# (cm.diagonal().sum()/cm.sum())*100
classifier.score(X_train,y_train)*100
# + colab={"base_uri": "https://localhost:8080/"} id="T68lbFMLHN-Y" outputId="614ef561-2782-44e1-bcee-718875b0244a"
classifier.score(X_test,y_test)*100
# + colab={"base_uri": "https://localhost:8080/"} id="FunWKO7JHN-Y" outputId="2ca051ec-bcd1-4a2c-f1b2-173344dceeb7"
cm1 = confusion_matrix(y_test, classifier.predict(X_test))
cm1
# + colab={"base_uri": "https://localhost:8080/"} id="gQ0gfNb4HN-Z" outputId="65d26c0e-d50e-458c-f5dd-8d726b28b01c"
(cm1.diagonal().sum()/cm1.sum())*100
# + id="1cuQB9d8HN-a"
| Random Forest Classification-Transfusion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# # Project "Training a cat to catch a Mouse using Q-learning and Qiskit"
#
# Participants:
#
# ## 1. Intoduction
# ### 1.1 Idea
# The idea of this project is to train a novice(single agent) to catch a mouse in a grid environment 3*3.
#
# We placed the cat in the lower right corner and the mouse in the upper left corner. Our task is to find the best way for the cat to catch a motionless (and unsuspecting danger) mouse. We want to test whether it is possible to solve such a problem using quantum computing.
#
# <table>
# <tr>
# <th>MOUSE</th>
# <td>EMPTY</td>
# <td>EMPTY</td>
# </tr>
#
# <tr>
# <td>EMPTY</td>
# <td>EMPTY</td>
# <td>EMPTY</td>
# </tr>
#
# <tr>
# <td>EMPTY</td>
# <td>EMPTY</td>
# <th>CAT</th>
# </tr>
#
# </table>
#
#
# ### 1.2 Q-Learning
# Q-Learning is a reinforcement learning algorithm to learn the value of an action in a particular state.
# When Q-learning is performed we create what’s called a **q-table** or matrix that follows the shape of *[state, action]* and we initialize our values to zero. We then update and store our q-values after an episode. This q-table becomes a reference table for our agent to select the best action based on the q-value.
#
#
# An agent interacts with the environment in 1 of 2 ways. The first is to use the q-table as a reference and view all possible actions for a given state. The agent then selects the action based on the max value of those actions. This is known as **exploiting** since we use the information we have available to us to make a decision.
#
#
# The second way to take action is to act randomly. This is called **exploring**. Instead of selecting actions based on the max future reward we select an action at random. Acting randomly is important because it allows the agent to explore and discover new states that otherwise may not be selected during the exploitation process. You can balance exploration/exploitation using epsilon (ε) and setting the value of how often you want to explore vs exploit. Here’s some rough code that will depend on how the state and action space are setup.
#
# Q-table is updated in every episode of training via **Bellman Equation**:
# $$
# \begin{align}
# \underbrace{\text{New}Q(s,a)}_{\scriptstyle\text{New Q-Value}}=Q(s,a)+\mkern-34mu\underset{\text{New Q-Value}}{\underset{\Bigl|}{\alpha}}\mkern-30mu \underbrace{R(s,a)}_{\scriptstyle\text{Reward}}+ \mkern-30mu\underset{\text{Discount rate}}{\underset{\Biggl |}{\gamma}}\mkern-75mu * \overbrace{\max Q'(s',a')}^{\scriptstyle\substack{\text{Maximum predicted reward, given} \\ \text{new state and all possible actions}}} * \mkern-45mu-Q(s,a)
# \end{align}
# $$
#
# ### 1.3 Variational Quantum Eigensolver (VQE)
#
# VQE is an application of the variational method of quantum mechanics.
# $$
# \begin{align}
# \lambda_{min} \le \langle H \rangle_{\psi} = \langle \psi | H | \psi \rangle = \sum_{i = 1}^{N} \lambda_i | \langle \psi_i | \psi\rangle |^2
# \end{align}
# $$
# The above equation is known as the **variational method**.
#
# When the Hamiltonian of a system is described by the Hermitian matrix $H$ the ground state energy of that system, $E_{gs}$, is the smallest eigenvalue associated with $H$. By arbitrarily selecting a wave function $|\psi \rangle$ (called an *ansatz*) as an initial guess approximating $|\psi_{min}\rangle$, calculating its expectation value, $\langle H \rangle_{\psi}$, and iteratively updating the wave function, arbitrarily tight bounds on the ground state energy of a Hamiltonian may be obtained.
#
# A systematic approach to varying the ansatz is required to implement the variational method on a quantum computer. VQE does so through the use of a parameterized circuit with a fixed form. Such a circuit is often called a *variational form*, and its action may be represented by the linear transformation $U(\theta)$. A variational form is applied to a starting state $|\psi\rangle$ (such as the vacuum state $|0\rangle$, or the Hartree Fock state) and generates an output state $U(\theta)|\psi\rangle\equiv |\psi(\theta)\rangle$. Iterative optimization over $|\psi(\theta)\rangle$ aims to yield an expectation value $\langle \psi(\theta)|H|\psi(\theta)\rangle \approx E_{gs} \equiv \lambda_{min}$. Ideally, $|\psi(\theta)\rangle$ will be close to $|\psi_{min}\rangle$ (where 'closeness' is characterized by either state fidelity, or Manhattan distance) although in practice, useful bounds on $E_{gs}$ can be obtained even if this is not the case.
#
# ## 2. Solution scheme via Q-learning & Quantum Calculations
#
# The scheme of our solution a as follows: Each point of our medium of size 3*3 corresponds to a quantum circuit with configurable parameters. Each such scheme consists of several U3-gates, the type of which is shown below.
#
# $$
# \begin{align}
# U3(\theta, \phi, \lambda) = \begin{pmatrix}\cos(\frac{\theta}{2}) & -e^{i\lambda}\sin(\frac{\theta}{2}) \\ e^{i\phi}\sin(\frac{\theta}{2}) & e^{i\lambda + i\phi}\cos(\frac{\theta}{2}) \end{pmatrix}
# \end{align}
# $$
#
# One of the options for implementing such a quantum circuit looks like this:
#
# 
#
from copy import deepcopy
import numpy as np
import random
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister
from qiskit import Aer, transpile, assemble
from qiskit.providers import backend
from qiskit.aqua.components.optimizers import COBYLA
import matplotlib.pyplot as plt
import itertools
# ### 2.3 Basic Classes
#
# **State**
#
# This class is needed for describing the position of the cat and operating with it.
#
# **GridWorld**
#
# We use this for operating with out grid where the cat and the mouse are located.
#
# **QNet**
#
# Here we can perfrom all the quantum computation needed for updating the q-tables in each point.
#
# **Cat**
#
# This class describes the cat's actions and update q-circuits accordingly.
#
# **PetSchool**
#
# In this class we can train our model and show the results.
# +
# TYPES:
CAT = "c"
MOUSE = "m"
EMPTY = "emp"
# ACTIONS:
UP = "00"
DOWN = "01"
LEFT = "10"
RIGHT = "11"
ACTIONS = [UP, DOWN, LEFT, RIGHT]
# random seed
random.seed(10)
np.random.seed(10)
# -
# state of cat
class State:
def __init__(self, catP):
#self.row = catP[0]
#self.column = catP[1]
self.catP = catP
def __eq__(self, other):
return isinstance(other, State) and self.catP == other.catP
def __hash__(self):
return hash(str(self.catP))
def __str__(self):
return f"State(cat_pos={self.catP})"
# GridWorld
# e.g.
# MOUSE | EMPTY | EMPTY
# EMPTY | EMPTY | EMPTY
# EMPTY | EMPTY | CAT
class GridWorld:
def __init__(self, s, catP, mouseP):
self.numRows = s[0]
self.numColumns = s[1]
self.catP = catP
self.mouseP = mouseP
# self.dogP = dogP
assert(not self.compaireList(self.catP, self.mouseP))
def getItem(self, p):
if p[0]>=self.numRows or p[0]<0:
return None
if p[1]>=self.numColumns or p[1]<0:
return None
if self.compaireList(p, catP):
return CAT
elif self.compaireList(p, mouseP):
return MOUSE
# elif self.compaireList(p, DOG):
# return DOG
else:
return EMPTY
def compaireList(self, l1,l2):
for i, j in zip(l1, l2):
if i!=j:
return False
return True
def getNumRows(self):
return self.numRows
def getNumColumns(self):
return self.numColumns
def getMouse(self):
return self.mouse
def getCatP(self):
return self.catP
def setCatP(self, p):
self.catP = p
def setMouseP(self, p):
self.mouseP = p
def initCatState(self, rd = False):
# init cat position
if not rd:
catP = [self.getNumRows() - 1, self.getNumColumns() - 1]
else:
catP = [random.randint(0, self.getNumRows()), random.randint(0, self.getNumColumns())]
while self.getItem(catP) != EMPTY and self.getItem(catP) != CAT:
catP = [random.randint(0, self.getNumRows()), random.randint(0, self.getNumColumns())]
self.setCatP(catP)
return State(catP)
def show(self):
output = ""
for i in range(self.numRows):
for j in range(self.numColumns):
if self.compaireList([i,j], self.catP):
output += CAT + " "
if self.compaireList([i,j], self.mouseP):
output += MOUSE + " "
if not self.compaireList([i,j], self.catP) and not self.compaireList([i,j], self.mouseP):
output += EMPTY + " "
output += "\n"
print(output)
# QNet
class QNet:
def __init__(self, qTable, gridWorld:GridWorld, alpha=0.1, gamma=1.0, eps=0.2, actions=[UP, DOWN, LEFT, RIGHT], numParams=6):
self.gw = gridWorld
self.qt = qTable
self.eps = eps
self.backend = Aer.get_backend("qasm_simulator")
self.NUM_SHOTS = 1000 # number of measurements
self.optimizer = COBYLA(maxiter=500, tol=0.0001) # off the shelf
self.gamma = gamma
self.alpha = alpha
self.ACTIONS = actions
# self.rets = {(0,0):([0,..,0],0.0,0), ...}
self.rets = dict() # resulting parameters after optimization for all points in the grid
self.state = None
for i in range(self.gw.getNumRows()):
for j in range(self.gw.getNumColumns()):
self.rets[i, j] = (np.random.rand(numParams), 0.0, 0)
def qcMaker(self, params):
qr = QuantumRegister(2, name="q")
cr = ClassicalRegister(2, name="c")
qc = QuantumCircuit(qr, cr)
qc.u3(params[0], params[1], params[2], qr[0])
qc.u3(params[3], params[4], params[5], qr[1])
# qc.cx(qr[0], qr[1]) # create entangelment
qc.measure(qr, cr)
return qc
def newPosition(self, state, action):
p = deepcopy(state.catP)
if action == UP:
p[0] = max(0, p[0] - 1)
elif action == DOWN:
p[0] = min(self.gw.getNumRows() - 1, p[0]+1)
elif action == LEFT:
p[1] = max(0, p[1] - 1)
elif action == RIGHT:
p[1] = min(self.gw.getNumColumns() - 1, p[1] + 1)
else:
raise ValueError(f"Unkown action {action}")
return p
def getReward(self, p):
grid = self.gw.getItem(p)
if grid == EMPTY:
reward = -1
elif grid == MOUSE:
reward = 1000
elif grid == CAT:
reward = -1
else:
raise ValueError(f"Unknown grid item {grid}")
return reward
def selectAction(self, state, training):
if random.uniform(0, 1) < self.eps:
return random.choice(self.ACTIONS)
else:
if training:
self.state = deepcopy(state)
self.updateCircuit(state)
return self.ACTIONS[np.argmax(self.qt[self.state.catP[0], self.state.catP[1]])]
def lossFunction(self, params):
action = ""
qc = self.qcMaker(params=params)
t_qc = transpile(qc, self.backend)
job = assemble(t_qc, shots=self.NUM_SHOTS)
rlt = self.backend.run(job).result()
counts = rlt.get_counts(qc)
# speedup training, cross the ravine
if random.uniform(0, 1) < self.eps:
action = random.choice(self.ACTIONS)
else:
action = max(counts, key = counts.get)
nextPosition = self.newPosition(self.state, action) # handle the
reward = self.getReward(nextPosition)
targetQvalue = reward + self.gamma * np.max(self.qt[nextPosition[0],nextPosition[1]])
predictedQvalue = self.calculateQvalue(action, nextPosition, reward, targetQvalue, self.state)
# update q-table
self.updateQtable(predictedQvalue, action)
return targetQvalue - self.qt[self.state.catP[0],self.state.catP[1]][int(action,2)]
def updateQtable(self, predictedQvalue, action):
if self.qt[(self.state.catP[0],self.state.catP[1])][int(action,2)] < predictedQvalue:
self.qt[self.state.catP[0],self.state.catP[1]][int(action,2)] = predictedQvalue
def calculateQvalue(self, action, nextPosition, reward, targetQvalue, state:State):
targetQvalue = reward + self.gamma * np.max(self.qt[nextPosition[0],nextPosition[1]])
return self.qt[state.catP[0], state.catP[1]][int(action,2)] + self.alpha * (targetQvalue - self.qt[state.catP[0],state.catP[1]][int(action,2)]) # update q-table
def updateCircuit(self, state:State):
self.rets[state.catP[0], state.catP[1]] = self.optimizer.optimize(num_vars=6, objective_function=self.lossFunction, initial_point=self.rets[state.catP[0], state.catP[1]][0])
def setAlpha(self, alpha):
self.alpha = alpha
# @Daniel-Molpe
def drawVectors(self, hasdiagonals):
# Draw vectors representing the cat's desired direction for each place in the grid based on the Qtable
x = np.linspace(0, self.gw.getNumColumns()-1, self.gw.getNumColumns())
y = np.linspace(0, self.gw.getNumRows()-1, self.gw.getNumRows())
vecx=np.zeros([len(x),len(y)])
vecy=np.zeros([len(x),len(y)])
for i in range(len(x)):
for j in range(len(y)):
vecx[i,j] = self.qt[(x[i], y[j])][3]-self.qt[(x[i], y[j])][2]
vecy[i,j] = self.qt[(x[i], y[j])][0]-self.qt[(x[i], y[j])][1]
norm = np.sqrt(vecx[i,j]**2 + vecy[i,j]**2)
vecx[i,j]=-vecx[i,j]/norm
vecy[i,j]=-vecy[i,j]/norm
pts = itertools.product(x, y)
plt.scatter(*zip(*pts), marker='o', s=30, color='red')
X, Y = np.meshgrid(x, y)
QP = plt.quiver(X, Y, vecx, vecy)
plt.grid()
plt.show()
# Agent: cat
class Cat:
def __init__(self, qNet: QNet, training=True, eps = 0.2, actions = [UP, DOWN, LEFT, RIGHT]):
self.eps = eps
self.training = training
self.qNet = qNet
self.ACTIONS = actions
self.state = None
def newPosition(self, state, action):
p = deepcopy(state.catP)
if action == UP:
p[0] = max(0, p[0] - 1)
elif action == DOWN:
p[0] = min(self.qNet.gw.getNumRows() - 1, p[0] + 1)
elif action == LEFT:
p[1] = max(0, p[1] - 1)
elif action == RIGHT:
p[1] = min(self.qNet.gw.getNumColumns() - 1, p[1] + 1)
else:
raise ValueError(f"Unkown action {self.ACTIONS[action]}")
return p
def getReward(self, p):
grid = self.qNet.gw.getItem(p)
if grid == MOUSE:
reward = 1000
end = True
elif grid == EMPTY:
reward = -1
end = False
elif grid == CAT:
reward = -1
end = False
else:
raise ValueError(f"Unknown grid item {grid}")
return reward, end
def act(self, state, action):
p = self.newPosition(state, action)
reward, end = self.getReward(p)
return p, reward, end
def updateQtable(self, action, p, reward, state):
pqv = self.qNet.calculateQvalue(action, p, reward, state)
self.qNet.updateQtable(pqv, action)
def setTraining(self, training):
self.Training = training
# The pet school
class PetSchool:
def __init__(self, cat:Cat, numEpisodes, maxEpisodeSteps, training=True, minAlpha = 0.02, eps = 0.2):
self.cat = cat
self.training = training
self.NUM_EPISODES = numEpisodes
self.MAX_EPISODE_STEPS = maxEpisodeSteps
self.alphas = np.linspace(1.0, minAlpha, self.NUM_EPISODES)
self.eps = eps
def train(self):
a=[]
b=[]
counter = 0
rd = True
for e in range(self.NUM_EPISODES): # episode: a rund for agent
print("episode: ", e)
if e > int(self.NUM_EPISODES/2):
rd = False
state = self.cat.qNet.gw.initCatState(rd=rd) # default is rd = False
self.cat.qNet.setAlpha(self.alphas[e])
total_reward = 0
step = 0
end = False
for _ in range(self.MAX_EPISODE_STEPS): # step: a time step for agent
action = self.cat.qNet.selectAction(deepcopy(state), self.training)
p, reward, end = self.cat.act(state, action)
self.catMoveTo(p)
# self.cat.updateQtable(action, p, reward, state) # speedup learning
total_reward += reward
step += 1
counter += 1
if end:
print("catch the mouse!!!")
print("total reward: ", total_reward, "steps: ", step)
break
a.append(step)
b.append(e)
print("counter: ", counter)
plt.plot(b,a)
def catMoveTo(self, p):
self.cat.qNet.gw.setCatP(p)
def show(self):
self.cat.qNet.gw.show()
print("qTable: ", self.cat.qNet.qt)
print("\nparams: ", self.cat.qNet.rets)
self.cat.qNet.drawVectors(False)
def result(self):
return self.cat.qNet.qt, self.cat.qNet.rets
def initqTable(self, actions, size):
d = {}
for i in range(size[0]):
for j in range(size[1]):
d[i,j] = np.zeros(len(actions))
return d
# @Daniel-Molpe
def mouseMove(p,oldPos): # goal (mouse) moves randomly with prob p every time the cat moves
side = min(self.cat.qNet.gw.getNumColumns(), self.cat.qNet.gw.getNumRows()) # Number of cells per side of the grid
if np.random.random() < p:
n = np.random.random()
if n < 0.25:
newPos = (max(0, oldPos[0]-1),oldPos[1])
elif n < 0.5:
newPos = (min(side - 1, oldPos[0]+1),oldPos[1])
elif n < 0.75:
newPos = (oldPos[0],max(0, oldPos[1]-1))
else:
newPos = (oldPos[0],min(side - 1, oldPos[1]+1))
else:
newPos = oldPos
return newPos
# Super Pparameter
gridSize = [3, 3]
catP = [gridSize[0]-1, gridSize[0]-1]
mouseP = [0, 0]
EPS = 50 # 500 for more training time
MAX_EPS_STEP = 50 # 1000 for more training time
sizeOfParams = 6
gamma = 0.98
# +
def initqTable(size, actions=[UP, DOWN, LEFT, RIGHT]):
d = {}
for i in range(size[0]):
for j in range(size[1]):
d[i,j] = np.zeros(len(actions))
return d
# initGridWorld
gridWorld = GridWorld(gridSize, catP=catP, mouseP=mouseP)
# init q Table
qt = initqTable(gridSize)
# init q Circuit
qNet = QNet(qt, gridWorld, gamma=gamma)
# init cat
cat = Cat(qNet=qNet)
# init pet school
petSchool = PetSchool(cat, EPS, MAX_EPS_STEP)
# start training
petSchool.train()
# -
# show what have been learned
petSchool.show()
# +
# @<NAME>
backend = Aer.get_backend("qasm_simulator")
NUM_SHOTS = 1000 # number of measurements
optimizer = COBYLA(maxiter=500, tol=0.0001) # off the shelf
def qcMaker(params):
qr = QuantumRegister(2, name="q")
cr = ClassicalRegister(2, name="c")
qc = QuantumCircuit(qr, cr)
qc.u3(params[0], params[1], params[2], qr[0])
qc.u3(params[3], params[4], params[5], qr[1])
qc.measure(qr, cr)
return qc
def calcActions(d):
actions = dict()
for key in d.keys():
qc = qcMaker(d[key][0])
t_qc = transpile(qc, backend)
job = assemble(t_qc, shots=NUM_SHOTS)
rlt = backend.run(job).result()
counts = rlt.get_counts(qc)
action = max(counts, key = counts.get)
actions[key] = action
return actions
# qt is a dict with all q-values
def calcPerformance(d, actions: dict, qt: dict):
diff = 0
# the difference between two action is defined as the steps difference between two action
# e.g. left needs two steps turn to right.
def difference(a, b):
if a == UP:
if b == DOWN:
return (2*2)/5
else:
return 1/5
elif a == DOWN:
if b == UP:
return (2*2)/5
else:
return 1/5
elif a == LEFT:
if b == RIGHT:
return (2*2)/5
else:
return 1/5
elif a == RIGHT:
if b == LEFT:
return (2*2)/5
else:
return 1/5
for key in d.keys():
action = ACTIONS[int(actions[key],2)] # "00","01" ..
targetAction = ACTIONS[np.argmax(qt[key])] # "00","01" ..
if action != targetAction:
diff += difference(action, targetAction)
return diff/len(d.keys())
qt, rets = petSchool.result()
print("MSE of trained circuit: ", calcPerformance(rets, calcActions(rets), qt))
| catHiddenMouse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercise 01.1
#
# Degrees Fahrenheit ($T_F$) are converted to degrees Celsius ($T_c$) using the formula:
#
# $$
# T_c = 5(T_f - 32)/9
# $$
#
# Write a program to convert 78 degrees Fahrenheit to degrees Celsius and print the result.
# Write your program such that you can easily change the temperature in Fahrenheit that you are converting from.
#
# Use the variable name `T_c` for the temperature in degrees Celsius.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "9aad06999d15d78fbcee70dcbca838c7", "grade": false, "grade_id": "cell-ca2d6af7fabf2918", "locked": false, "schema_version": 3, "solution": true}
# YOUR CODE HERE
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "298ddee2ca0d1dd09bc995505dac5c63", "grade": true, "grade_id": "cell-a7542313c8cdf4ed", "locked": true, "points": 0, "schema_version": 3, "solution": false}
assert round(T_c - 25.55555555555555, 10) == 0.0
# -
# ## Exercise 01.2
#
# You have been tasked with developing a mortgage affordability test tool. A component is to compute the monthly
# interest payments for a range of scenarios.
#
# Interest on a particular mortgage is charged at fixed rate above the Bank of England (BoE) 'official Bank Rate'.
# Interest is computed per annum, and interest payments spread equally over each month of the year.
# Write a program that computes the interest payable each month, with variables for:
#
# 1. Loan principal (amount borrowed)
# 1. Official Bank Rate (percentage, expressed per annum)
# 1. Rate over the official Bank Rate (percentage, expressed per annum)
#
# Test your program with a loan principal of £150,000, BoE rate of 0.5%, and rate over the
# BoE rate of 1.49%. Use the variable name `interest` for the monthly interest amount.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "c26b7adb39cb9665b26873573b1f0a6c", "grade": false, "grade_id": "cell-aaab53918bb9627d", "locked": false, "schema_version": 3, "solution": true}
# Assign data to variables
# YOUR CODE HERE
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "d01c1d6775e975163ba8b456945ee4bc", "grade": true, "grade_id": "cell-258e3a2c80f6a765", "locked": true, "points": 0, "schema_version": 3, "solution": false}
assert round(interest - 248.75, 10) == 0.0
| Assignment/01 Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from keras.applications import InceptionV3, ResNet50
from keras.layers import Dense, Flatten, GlobalAveragePooling2D
from keras import Model
# # Original InceptionV3 Model with pretrained weights loaded from imagenet
orig = InceptionV3(weights='imagenet', )
# +
# orig.summary()
# -
# # InceptionV3 model with a custom input_shape, here `480, 480, 3` and with no output layer
model = InceptionV3(input_shape=(480, 480, 3), weights='imagenet', include_top=False)
# +
# model.summary()
# -
# ### Create `128` category output
last_layer_output = model.layers[-1].output
x = GlobalAveragePooling2D()(last_layer_output)
x = Dense(128, activation='softmax', name='predictions')(x)
new_model = Model(inputs=model.input, outputs=x)
# +
# new_model.summary()
# -
# ### Freeze layers whose `name` that start with `conv` in the last `20` layers of the network
for l in new_model.layers[-20:]:
if l.name.startswith('conv'):
print l.name
l.trainable = False
# +
# new_model.summary()
# -
| notes/Import_Pretrained_Models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import cv2
import glob
import matplotlib.pyplot as plt
# %matplotlib qt
# %matplotlib inline
from mpl_toolkits.axes_grid1 import ImageGrid
# prepare object points, like (0,0,0), (1,0,0), (2,0,0) ....,(8,5,0)
objp = np.zeros((6*9,3), np.float32)
objp[:,:2] = np.mgrid[0:9, 0:6].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
# Make a list of calibration images
images = glob.glob('camera_cal/calibration*.jpg')
i = 0
fig = plt.figure(1, (30, 30))
grid = ImageGrid(fig, 111, # similar to subplot(111)
nrows_ncols=(7, 3), # creates 2x2 grid of axes
axes_pad=0.2, # pad between axes in inch.
)
print("Total images :" , len(images))
# Step through the list and search for chessboard corners
for idx, fname in enumerate(images):
img = cv2.imread(fname)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray, (9,6), None)
# If found, add object points, image points
if ret == True:
objpoints.append(objp)
imgpoints.append(corners)
# Draw and display the corners
cv2.drawChessboardCorners(img, (9,6), corners, ret)
ax = grid[i]
ax.imshow(img)
i = i +1
plt.show()
#cv2.destroyAllWindows()
#print(ret)
#print(corners)
print("Done")
# -
# If the above cell ran sucessfully, you should now have objpoints and imgpoints needed for camera calibration. Run the cell below to calibrate, calculate distortion coefficients, and test undistortion on an image!
# +
import pickle
# %matplotlib inline
# Test undistortion on an image
img = cv2.imread('camera_cal/test_image.jpg')
img_size = (img.shape[1], img.shape[0])
# Do camera calibration given object points and image points
ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints, img_size,None,None)
dst = cv2.undistort(img, mtx, dist, None, mtx)
cv2.imwrite('camera_cal/test_undist.jpg',dst)
# Save the camera calibration result for later use (we won't worry about rvecs / tvecs)
dist_pickle = {}
dist_pickle["mtx"] = mtx
dist_pickle["dist"] = dist
pickle.dump( dist_pickle, open( "camera_cal/wide_dist_pickle.p", "wb" ) )
#dst = cv2.cvtColor(dst, cv2.COLOR_BGR2RGB)
# Visualize undistortion
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,10))
ax1.imshow(img)
ax1.set_title('Original Image', fontsize=30)
ax2.imshow(dst)
ax2.set_title('Undistorted Image', fontsize=30)
# -
# ## Perspective transform
# +
#import pickle
#import cv2
#import numpy as np
#import matplotlib.pyplot as plt
#import matplotlib.image as mpimg
# Read in the saved camera matrix and distortion coefficients
# These are the arrays you calculated using cv2.calibrateCamera()
dist_pickle = pickle.load( open( "camera_cal/wide_dist_pickle.p", "rb" ) )
mtx = dist_pickle["mtx"]
dist = dist_pickle["dist"]
# Read in an image
img = cv2.imread('test_images/straight_lines1.jpg')
nx = 9 # the number of inside corners in x
ny = 6 # the number of inside corners in y
# MODIFY THIS FUNCTION TO GENERATE OUTPUT
# THAT LOOKS LIKE THE IMAGE ABOVE
print(dist)
plt.imshow(img)
img = cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
plt.figure()
plt.imshow(img)
def corners_unwarp(img, nx, ny, mtx, dist):
# 1) Undistort using mtx and dist.
undis = cv2.undistort(img, mtx, dist, None, mtx)
# 2) Convert to grayscale
plt.figure()
plt.imshow(undis)
gray = cv2.cvtColor(undis, cv2.COLOR_BGR2GRAY)
# 3) Find the chessboard corners
offset=10
src_bottom_left = [260,680]
src_bottom_right = [1040,680]
src_top_left = [581,460]
src_top_right = [700,460]
destination_bottom_left = [100,700]
destination_bottom_right = [1000,700]
destination_top_left = [100,50]
destination_top_right = [1000,50]
#src = np.float32([corners[0],corners[nx-1],corners[-1],corners[-nx]])
src = np.float32([[src_top_left,src_top_right,src_bottom_right,src_bottom_left]])
print(src)
print(img.shape)
dst_points = np.float32([[destination_top_left,destination_top_right,destination_bottom_right,destination_bottom_left]])
#dst_points = np.float32([[offset,offset],
# [img.shape[1]-offset,offset],
# [img.shape[1]-offset,img.shape[0]-offset],
# [offset,img.shape[0]-offset]])
#dst_points = np.float32([(offset,offset),(1200-offset,offset),()])
M = cv2.getPerspectiveTransform(src, dst_points)
warped = cv2.warpPerspective(undis, M, (img.shape[1],img.shape[0]), flags=cv2.INTER_LINEAR)
#delete the next two lines
#M = None
#warped = np.copy(img)
return warped, M
top_down, perspective_M = corners_unwarp(img, nx, ny, mtx, dist)
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
f.tight_layout()
ax1.imshow(img)
ax1.set_title('Original Image', fontsize=50)
ax2.imshow(top_down)
ax2.set_title('Undistorted and Warped Image', fontsize=50)
plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
# -
| NoteBook1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import json
import math
import datetime
from time import sleep, strptime
from collections import OrderedDict
import pandas as pd
import numpy as np
import requests
#import plotly.offline as pyo
#pyo.init_notebook_mode()
import plotly.express as px
import plotly.graph_objects as go
# -
# ### Load csv files provided by Wienerlinien
# +
# Info about line ids and stop ids. It also contains order of the stops
fahrwegverlaeufe = pd.read_csv('../../resources/wienerlinien_csv/1_wienerlinien-ogd-fahrwegverlaeufe.csv', sep=";")
# Polygon coordinates between stops. Also contains distances in meters.
gps_punkte = pd.read_csv('../../resources/wienerlinien_csv/2_wienerlinien-ogd-gps-punkte.csv', sep=";")
# Stops. Contains StopID which is used for realtime data requests.
haltepunkte = pd.read_csv('../../resources/wienerlinien_csv/3_wienerlinien-ogd-haltepunkte.csv', sep=";")
# Similar to stops but on DIVA level. DIVA - Summary of multiple Stops of a Station area
haltestellen = pd.read_csv('../../resources/wienerlinien_csv/4_wienerlinien-ogd-haltestellen.csv', sep=";")
# Lines info
linien = pd.read_csv('../../resources/wienerlinien_csv/5_wienerlinien-ogd-linien.csv', sep=";")
# Staircase info
steige = pd.read_csv('../../resources/wienerlinien_csv/6_wienerlinien-ogd-steige.csv', sep=";")
# Distances between stops. Similar to gps_punkte but without inbetween points. Also probably more accurate since not polygon.
verbindungen = pd.read_csv('../../resources/wienerlinien_csv/7_wienerlinien-ogd-verbindungen.csv', sep=";")
# Data validity. Not needed.
version = pd.read_csv('../../resources/wienerlinien_csv/8_wienerlinien-ogd-version.csv', sep=";")
# No description. Not needed.
teilstrecken_lonlat = pd.read_csv('../../resources/wienerlinien_csv/wienerlinien-ogd-teilstrecken-lonlat.csv', sep=";")
teilstrecken_wkt = pd.read_csv('../../resources/wienerlinien_csv/wienerlinien-ogd-teilstrecken-wkt.csv', sep=";")
# +
# Prepare the ubahn_df (same as in data retrieval notebook)
ubahn_linien = linien[linien.MeansOfTransport == "ptMetro"]
ubahn_df = fahrwegverlaeufe.merge(ubahn_linien, how="inner", on="LineID")
ubahn_df = ubahn_df.merge(haltepunkte, how="inner", on="StopID")
# clean up
# Meidling has 2 different StopText names. Lets unify them
ubahn_df.loc[ubahn_df.StopText.str.contains("Meidling"), "StopText"] = "Meidling Hauptstraße"
# turns out some stations are in the wrong order
ubahn_df.loc[ubahn_df[(ubahn_df.LineText=="U2") & (ubahn_df.PatternID==1) & (ubahn_df.StopText=="Donaustadtbrücke")].index,"StopSeqCount"] = 8
ubahn_df.loc[ubahn_df[(ubahn_df.LineText=="U2") & (ubahn_df.PatternID==1) & (ubahn_df.StopText=="Donaumarina")].index,"StopSeqCount"] = 9
#ubahn_df.head()
# -
# ### Functions
# +
def get_diva_from_name(diva_df, stop_text):
"""
Accepts name of the station.
Returns DIVA id of the station.
"""
ubahn_df_reduced = diva_df[["DIVA", "StopText"]].drop_duplicates()
diva = ubahn_df_reduced[ubahn_df_reduced.StopText == stop_text].DIVA.astype(int)
if len(diva) > 1:
raise ValueError("stop_text is not unique")
if len(diva) == 0:
raise ValueError("stop_text not found")
return str(diva.values[0])
def get_name_from_diva(diva_df, diva):
"""
Accepts DIVA id of the station.
Returns name of the station.
"""
ubahn_df_reduced = diva_df[["DIVA", "StopText"]].drop_duplicates()
name = ubahn_df_reduced[ubahn_df_reduced.DIVA == diva].StopText
return str(name.values[0])
# -
# ### Data preparation
# +
# load data from file (data was retireved for 1st of December 2021 at 12h)
travel_df = pd.read_csv("../../resources/data/retrieved_data/ubahn_travel_times_2021_12_01__12_00.csv")
travel_df.set_index("idx", inplace=True)
travel_df.index.name = None
# lets remove stations that were skipped
diva_ids_to_remove = []
# both Neue Donau DIVA ids
diva_ids_to_remove.append(60200455)
diva_ids_to_remove.append(60201668)
diva_ids_to_remove.append(int(get_diva_from_name(ubahn_df, "Museumsquartier")))
diva_ids_to_remove.append(int(get_diva_from_name(ubahn_df, "Rathaus")))
travel_df.drop(labels=diva_ids_to_remove, axis=0, inplace=True)
travel_df.drop(labels=[str(x) for x in diva_ids_to_remove], axis=1, inplace=True)
#travel_df.head()
# +
# transform matrix into a long format dataframe
travel_df_long = travel_df.unstack()
travel_df_long = pd.DataFrame(travel_df_long)
travel_df_long.reset_index(inplace=True)
travel_df_long.columns = ["source", "destination", "time"]
travel_df_long.source = travel_df_long.source.astype(int)
travel_df_long.head()
# +
# Let's add station names to the dataframe
# create a "DIVA <-> StopName" mapping dataframe
diva_stop_mapping = ubahn_df[["DIVA", "StopText"]].drop_duplicates()
diva_stop_mapping = diva_stop_mapping[diva_stop_mapping.DIVA.isin(list(travel_df_long.source))]
travel_df_long = travel_df_long.merge(diva_stop_mapping, left_on="source", right_on="DIVA")
travel_df_long.drop("DIVA", axis=1, inplace=True)
travel_df_long.columns = ["source", "destination", "time", "source_name"]
travel_df_long = travel_df_long.merge(diva_stop_mapping, left_on="destination", right_on="DIVA")
travel_df_long.drop("DIVA", axis=1, inplace=True)
travel_df_long.columns = ["source", "destination", "time", "source_name", "destination_name"]
travel_df_long.head()
# +
# Lets prepare dataset with coordinates
travel_df_long = travel_df_long[["source","destination","time","source_name","destination_name"]]
# coordinates dataframe
coord_df = travel_df_long.merge(ubahn_df[["DIVA", "Longitude", "Latitude"]].groupby("DIVA").mean().reset_index(), how="inner", left_on="source", right_on="DIVA")
coord_df.drop("DIVA", axis=1, inplace=True)
coord_df.rename(mapper={"Longitude":"source_longitude", "Latitude":"source_latitude"}, axis=1, inplace=True)
coord_df = coord_df.merge(ubahn_df[["DIVA", "Longitude", "Latitude"]].groupby("DIVA").mean().reset_index(), how="inner", left_on="destination", right_on="DIVA")
coord_df.drop("DIVA", axis=1, inplace=True)
coord_df.rename(mapper={"Longitude":"destination_longitude", "Latitude":"destination_latitude"}, axis=1, inplace=True)
coord_df.head()
# -
# The dataset with stations, travel times and coordinates is now ready.
# Next we need to perform a transformation from space coordinates to time coordinates
# ### Time-space tranformation
# +
# Let's make a copy of the coord_df dataframe
transformed_df = coord_df.copy()
# Translate destination coordinates to the center (0,0)
# (Subtract source coordinates from destination coordinates)
transformed_df['destination_latitude_center'] = transformed_df['destination_latitude'] - transformed_df['source_latitude']
transformed_df["destination_longitude_center"] = transformed_df['destination_longitude'] - transformed_df['source_longitude']
# Let's scale
# calculate max coordinate in each row and scale both longitude and latitude to that value
max_coord = abs(transformed_df[["destination_longitude_center","destination_latitude_center"]]).max(axis=1)
transformed_df.destination_latitude_center /= max_coord
transformed_df.destination_longitude_center /= max_coord
# lastly lets scale each coordinate with travel time to get time dependent coordinates
#transformed_df.destination_longitude_center *= transformed_df.time_scaled
#transformed_df.destination_latitude_center *= transformed_df.time_scaled
# time into minutes
transformed_df.time = transformed_df.time/60
# we want the distance from the origin (0,0) to be same as travel time
# first we need to calculate the angle and then recalculate the lat (x) and long (y)
transformed_df["angle"] = np.arctan(abs(transformed_df.destination_longitude_center)/abs(transformed_df.destination_latitude_center))
#transformed_df.destination_latitude_center *= transformed_df.time
#transformed_df.destination_longitude_center *= transformed_df.time
transformed_df.destination_latitude_center = (np.cos(transformed_df.angle) * transformed_df.time) * np.copysign(1,transformed_df.destination_latitude_center)
transformed_df.destination_longitude_center = (np.sin(transformed_df.angle) * transformed_df.time) * np.copysign(1,transformed_df.destination_longitude_center)
transformed_df.destination_latitude_center.fillna(0, inplace=True)
transformed_df.destination_longitude_center.fillna(0, inplace=True)
# add station info
transformed_df = transformed_df.merge(ubahn_df[["DIVA","LineText"]].drop_duplicates().groupby(['DIVA']).agg(lambda col: ','.join(col)), how="inner", left_on="destination", right_on="DIVA").drop_duplicates()
transformed_df
# +
# Let's make a copy of the coord_df dataframe
transformed_df = coord_df.copy()
# Translate destination coordinates to the center (0,0)
# (Subtract source coordinates from destination coordinates)
transformed_df['destination_latitude_center'] = transformed_df['destination_latitude'] - transformed_df['source_latitude']
transformed_df["destination_longitude_center"] = transformed_df['destination_longitude'] - transformed_df['source_longitude']
# Transform time from seconds into minutes
transformed_df.time = transformed_df.time/60
# We want the distance from the origin (0,0) to be the same as the travel time
# First we need to calculate the angle and then recalculate the latitude (x) and longitude (y)
transformed_df["angle"] = np.arctan(abs(transformed_df.destination_longitude_center)/abs(transformed_df.destination_latitude_center))
# Let's calculate the final latitude and longitude
# by calculating sine and cosine of the angle and multiplying them by time
# Use np.copysign to determine the sign of the coordinate
transformed_df.destination_latitude_center = (np.cos(transformed_df.angle) * transformed_df.time) * np.copysign(1,transformed_df.destination_latitude_center)
transformed_df.destination_longitude_center = (np.sin(transformed_df.angle) * transformed_df.time) * np.copysign(1,transformed_df.destination_longitude_center)
transformed_df.destination_latitude_center.fillna(0, inplace=True)
transformed_df.destination_longitude_center.fillna(0, inplace=True)
# Add line info
transformed_df = transformed_df.merge(ubahn_df[["DIVA","LineText"]].drop_duplicates().groupby(['DIVA']).agg(lambda col: ','.join(col)), how="inner", left_on="destination", right_on="DIVA").drop_duplicates()
transformed_df
# -
# Store data as csv
transformed_df.to_csv("../../resources/data/prepared_data/time_space_map_ubahn_2021_12_01__12_00.csv", index=False)
# ### Interactive chart
# Unfortunatelly it is not possible to run interactive charts in jupyter notebooks.
#
# For this purpose the demo was created: https://martinvolk91.github.io/time-space-maps/
| src/data_preparation/.ipynb_checkpoints/data_preparation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nanoparticle‐microglial interaction in the ischemic brain is modulated by injury duration and treatment Workflow
# ##### Citation
# <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., 2020. Nanoparticle‐microglial interaction in the ischemic brain is modulated by injury duration and treatment. Bioengineering & Translational Medicine.. doi:10.1002/btm2.10175
# ## Purpose: To split tile scans, pick training and testing image sets, and prepping for running the full VAMPIRE Workflow
# The notebook as is was the skeleton used to produce the precise notebook for the publication cited above. This state with the data uploaded on our lab drive under this publication title will reproduce the results from our paper when run following protocol on the shared desktop folder location specific in user inputs.
# # Experimental Steps Necessary Before Notebook
# Experimental Steps:
# 1. Move/Download the images for testing and training into a new folder
# 2. Rename images to insure they include the condition somewhere in them
# 3. Add a folder for each of your stains into the folder created in step 1
# 4. Input the name of that folder into 'folder_location' below
# 5. Input the names of the nuclear stain into 'stain1' and the cell stain into 'stain2' below
# 6. Insert the number of slices into the np.arange command below
# 7. Insert the number of slices that you want to split each image into in 'slice number'
# 8. Increase the random_state_num by one
# 8. Add a folder labeled 'train' to your desktop
# 9. Add a folder labeled 'test' to your desktop
# 10. Within folder 'test' create a folder for each of your conditions
# # Steps the Notebook will need to follow
# *Step 1: Import necessary packages*
# *Step 2: User Inputs*
# *Step 3: Split the Image(s) into Color Channels*
# *Step 4: Split the Images*
# *Step 5: Moving the DAPI and Iba images into their own folders*
# *Step 6: Choose training and testing data sets*
# *Step 7: Moving the testing and training DAPI data sets into test and train folders*
# *Step 8: Renaming the DAPI and Iba datasets according to proper VAMPIRE naming modality*
# *Step 9: Splitting the test group into the appropriate conditions*
# *Step 10: Renaming the test images and getting their appropriate iba stain*
#
#
# *Final Experimentalist Steps:*
# 1. Follow Python 2.7 Protocol on shared Computer for VAMPIRE Analysis
# 2. Upload all files to the appropriate paper folder in the Shared Google Drive
# # Initial Experimental Design Plans (Used to Help Build Notebook)
# ### Personnel Workflow
# (1) Slicing - <NAME>
#
# (2) Staining/Imaging - <NAME>/<NAME>
#
# (3) Image Processing - <NAME> via Python & Jupyter Notebook
#
# (4) Data Analysis - <NAME>
#
# (5) Data Analysis Results To be sent to <NAME> for final Visualization
# ### Data Locations and File Types
#
# Original Images: Computer Attached to Confocal Microscope (as .nd2)
# --> Later move to the Shared Nance Lab Google Drive (as .tiff)
#
# Images for Processing: Shared Nance Lab Google Drie (as .tiff)
#
# Images to be converted for VAMPIRE workflow: Temporary Storage on communal or personal desktop (as .png)
# --> Long term storage on Shared Nance Lab Google Drive (as .png before and after segmentation)
#
# Cell Morphology Quantification Data: Temporary Storage on communal or personal desktop (as .csv)
# --> Long term storage on Shared Nance Lab Google Dirve (as. csv)
#
# Data Visualization: Completed for this paper using GraphPad Prism
# ### Treatment Groups
# (1) Non-treated Control
#
# (2) Oxygen-Glucose Deprivation 0.5h
#
# (3) Oxygen-Glucose Deprivation 1.5h
#
# (4) Treatment - Azithromycin
#
# (5) Treatment - SOD
# ### Brain Regions of Interest
# Details: Sprague-Dawley Post Natal Day 7 Rats (sex controlled)
# 1. Hippocampus
# 2. Cortex
# 3. Thalamus
# ### Stains Used in Step 3
# 1. iba1 - green channel
# 2. dapi - blue channel
# 3. PI - red channel
#
# ### Pertinent Confocal Settings
# Zoom: 40x for all images
# ### Number of Expected Images
# 1. 5 Treatment Groups
# 2. 3 Regions Per Treatment
# 3. 3 Images per Region
# 4. Quadrant the Images for Analysis
# 5. 2 Stains
#
# Approximatey 360 final images
| RAW_Joseph_Liao_OGD_Nanoparticle_interaction-Raw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
dataset = pd.read_csv('Position_Salaries.csv')
X = dataset.iloc[:, 1:2]
y = dataset.iloc[:, 2]
from sklearn.preprocessing import StandardScaler
scalar_x = StandardScaler()
scalar_y = StandardScaler()
X = scalar_x.fit_transform(X)
y = scalar_y.fit_transform(np.array(y).reshape(-1,1))
from sklearn.svm import SVR
regressor = SVR()
regressor.fit(X,y)
scalar_y.inverse_transform(regressor.predict(scalar_x.transform([[6.5]])))
plt.scatter(X, y, color='red')
plt.plot(X, regressor.predict(X), color='blue')
plt.title('SV Regression')
plt.xlabel('Level')
plt.ylabel('Salary')
plt.show()
X_grid = np.arange(min(X), max(X), 0.01) # choice of 0.01 instead of 0.1 step because the data is feature scaled
X_grid = X_grid.reshape((len(X_grid), 1))
plt.scatter(X, y, color='red')
plt.plot(X_grid, regressor.predict(X_grid).reshape(-1,1), color='blue')
plt.show()
| Machine Learning A-Z Template Folder/Part 2 - Regression/Section 7 - Support Vector Regression (SVR)/practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="H7LoMj4GA4n_"
# # Train a GPT-2 Text-Generating Model w/ GPU For Free
#
# inspired from (https://github.com/minimaxir/gpt-2-simple)
#
# + id="KBkpRgBCBS2_" colab={"base_uri": "https://localhost:8080/"} outputId="ec1e5863-0bf6-4c59-e97e-4bc46fa897fe"
# %tensorflow_version 1.x
# !pip install -q gpt-2-simple
import gpt_2_simple as gpt2
from datetime import datetime
from google.colab import files
# + [markdown] id="Bj2IJLHP3KwE"
# ## GPU
#
# Colaboratory uses either a Nvidia T4 GPU or an Nvidia K80 GPU. The T4 is slightly faster than the old K80 for training GPT-2, and has more memory allowing you to train the larger GPT-2 models and generate more text.
#
# You can verify which GPU is active by running the cell below.
# + id="sUmTooTW3osf" colab={"base_uri": "https://localhost:8080/"} outputId="6a137620-71a9-47d2-9c75-e2baec090e58"
# !nvidia-smi
# + [markdown] id="0wXB05bPDYxS"
# ## Downloading GPT-2
#
# If you're retraining a model on new text, you need to download the GPT-2 model first.
#
# There are three released sizes of GPT-2:
#
# * `124M` (default): the "small" model, 500MB on disk.
# * `355M`: the "medium" model, 1.5GB on disk.
# * `774M`: the "large" model, cannot currently be finetuned with Colaboratory but can be used to generate text from the pretrained model (see later in Notebook)
# * `1558M`: the "extra large", true model. Will not work if a K80 GPU is attached to the notebook. (like `774M`, it cannot be finetuned).
#
# Larger models have more knowledge, but take longer to finetune and longer to generate text. You can specify which base model to use by changing `model_name` in the cells below.
#
# The next cell downloads it from Google Cloud Storage and saves it in the Colaboratory VM at `/models/<model_name>`.
#
# This model isn't permanently saved in the Colaboratory VM; you'll have to redownload it if you want to retrain it at a later time.
# + id="P8wSlgXoDPCR" colab={"base_uri": "https://localhost:8080/"} outputId="7bad95fe-5670-413b-cb4c-61688229cc4f"
gpt2.download_gpt2(model_name="124M")
# + [markdown] id="N8KXuKWzQSsN"
# ## Mounting Google Drive
#
# The best way to get input text to-be-trained into the Colaboratory VM, and to get the trained model *out* of Colaboratory, is to route it through Google Drive *first*.
#
# Running this cell (which will only work in Colaboratory) will mount your personal Google Drive in the VM, which later cells can use to get data in/out. (it will ask for an auth code; that auth is not saved anywhere)
# + id="puq4iC6vUAHc" colab={"base_uri": "https://localhost:8080/"} outputId="38bef57e-e1ea-4f1e-85be-c553faf83754"
gpt2.mount_gdrive()
# + [markdown] id="BT__brhBCvJu"
# ## Uploading a Text File to be Trained to Colaboratory
#
# In the Colaboratory Notebook sidebar on the left of the screen, select *Files*. From there you can upload files:
#
# 
#
# Upload **any smaller text file** (<10 MB) and update the file name in the cell below, then run the cell.
# + id="6OFnPCLADfll"
file_name = "jokes.txt"
# + [markdown] id="HeeSKtNWUedE"
# If your text file is larger than 10MB, it is recommended to upload that file to Google Drive first, then copy that file from Google Drive to the Colaboratory VM.
# + id="-Z6okFD8VKtS"
gpt2.copy_file_from_gdrive(file_name)
# + [markdown] id="LdpZQXknFNY3"
# ## Finetune GPT-2
#
# The next cell will start the actual finetuning of GPT-2. It creates a persistent TensorFlow session which stores the training config, then runs the training for the specified number of `steps`. (to have the finetuning run indefinitely, set `steps = -1`)
#
# The model checkpoints will be saved in `/checkpoint/run1` by default. The checkpoints are saved every 500 steps (can be changed) and when the cell is stopped.
#
# The training might time out after 4ish hours; make sure you end training and save the results so you don't lose them!
#
# **IMPORTANT NOTE:** If you want to rerun this cell, **restart the VM first** (Runtime -> Restart Runtime). You will need to rerun imports but not recopy files.
#
# Other optional-but-helpful parameters for `gpt2.finetune`:
#
#
# * **`restore_from`**: Set to `fresh` to start training from the base GPT-2, or set to `latest` to restart training from an existing checkpoint.
# * **`sample_every`**: Number of steps to print example output
# * **`print_every`**: Number of steps to print training progress.
# * **`learning_rate`**: Learning rate for the training. (default `1e-4`, can lower to `1e-5` if you have <1MB input data)
# * **`run_name`**: subfolder within `checkpoint` to save the model. This is useful if you want to work with multiple models (will also need to specify `run_name` when loading the model)
# * **`overwrite`**: Set to `True` if you want to continue finetuning an existing model (w/ `restore_from='latest'`) without creating duplicate copies.
# + id="aeXshJM-Cuaf" colab={"base_uri": "https://localhost:8080/"} outputId="58277dab-77c3-4f2f-d910-c6a45b86e1b0"
sess = gpt2.start_tf_sess()
gpt2.finetune(sess,
dataset=file_name,
model_name='124M',
steps=1000,
restore_from='fresh',
run_name='run1',
print_every=10,
sample_every=200,
save_every=500
)
# + [markdown] id="IXSuTNERaw6K"
# After the model is trained, you can copy the checkpoint folder to your own Google Drive.
#
# If you want to download it to your personal computer, it's strongly recommended you copy it there first, then download from Google Drive. The checkpoint folder is copied as a `.rar` compressed file; you can download it and uncompress it locally.
# + id="VHdTL8NDbAh3"
gpt2.copy_checkpoint_to_gdrive(run_name='run1')
# + [markdown] id="qQJgV_b4bmzd"
# You're done! Feel free to go to the **Generate Text From The Trained Model** section to generate text based on your retrained model.
# + [markdown] id="pel-uBULXO2L"
# ## Load a Trained Model Checkpoint
#
# Running the next cell will copy the `.rar` checkpoint file from your Google Drive into the Colaboratory VM.
# + id="DCcx5u7sbPTD"
gpt2.copy_checkpoint_from_gdrive(run_name='run1')
# + [markdown] id="RTa6zf3e_9gV"
# The next cell will allow you to load the retrained model checkpoint + metadata necessary to generate text.
#
# **IMPORTANT NOTE:** If you want to rerun this cell, **restart the VM first** (Runtime -> Restart Runtime). You will need to rerun imports but not recopy files.
# + id="-fxL77nvAMAX"
sess = gpt2.start_tf_sess()
gpt2.load_gpt2(sess, run_name='run1')
# + [markdown] id="ClJwpF_ACONp"
# ## Generate Text From The Trained Model
#
# After you've trained the model or loaded a retrained model from checkpoint, you can now generate text. `generate` generates a single text from the loaded model.
# + id="4RNY6RBI9LmL"
gpt2.generate(sess, run_name='run1')
# + [markdown] id="oF4-PqF0Fl7R"
# If you're creating an API based on your model and need to pass the generated text elsewhere, you can do `text = gpt2.generate(sess, return_as_list=True)[0]`
#
# You can also pass in a `prefix` to the generate function to force the text to start with a given character sequence and generate text from there (good if you add an indicator when the text starts).
#
# You can also generate multiple texts at a time by specifing `nsamples`. Unique to GPT-2, you can pass a `batch_size` to generate multiple samples in parallel, giving a massive speedup (in Colaboratory, set a maximum of 20 for `batch_size`).
#
# Other optional-but-helpful parameters for `gpt2.generate` and friends:
#
# * **`length`**: Number of tokens to generate (default 1023, the maximum)
# * **`temperature`**: The higher the temperature, the crazier the text (default 0.7, recommended to keep between 0.7 and 1.0)
# * **`top_k`**: Limits the generated guesses to the top *k* guesses (default 0 which disables the behavior; if the generated output is super crazy, you may want to set `top_k=40`)
# * **`top_p`**: Nucleus sampling: limits the generated guesses to a cumulative probability. (gets good results on a dataset with `top_p=0.9`)
# * **`truncate`**: Truncates the input text until a given sequence, excluding that sequence (e.g. if `truncate='<|endoftext|>'`, the returned text will include everything before the first `<|endoftext|>`). It may be useful to combine this with a smaller `length` if the input texts are short.
# * **`include_prefix`**: If using `truncate` and `include_prefix=False`, the specified `prefix` will not be included in the returned text.
# + id="8DKMc0fiej4N"
gpt2.generate(sess,
length=250,
temperature=0.7,
prefix="LORD",
nsamples=5,
batch_size=5
)
# + [markdown] id="zjjEN2Tafhl2"
# For bulk generation, you can generate a large amount of text to a file and sort out the samples locally on your computer. The next cell will generate a generated text file with a unique timestamp.
#
# You can rerun the cells as many times as you want for even more generated texts!
| gpt2_finetune.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2020년 7월 1일 수요일
# ### BaekJoon - 8진수 2진수 (Python)
# ### 문제 : https://www.acmicpc.net/problem/1212
# ### 블로그 : https://somjang.tistory.com/entry/BaekJoon-1212%EB%B2%88-8%EC%A7%84%EC%88%98-2%EC%A7%84%EC%88%98-Python
# ### 첫번째 시도
print(bin(int(input(), 8))[2:])
| DAY 101 ~ 200/DAY146_[BaekJoon] 8진수 2진수 (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to use `dialogExtend` with jp_proxy_widgets
#
# The `dialogExtend` `jQueryUI` plugin extends the dialog component with various features
# such as maximization, minimization, and so forth. Below is a screen shot of an extended dialog
#
# <img src="dialogextend.png" width="600"/>
#
# <a href="https://www.jqueryscript.net/other/jQuery-jQuery-UI-Dialog-Enhancement-Plugin-dialogextend.html">Following examples from https://www.jqueryscript.net/other/jQuery-jQuery-UI-Dialog-Enhancement-Plugin-dialogextend.html</a>.
#
# The code for the plugin is designed to be loaded using the `require` module loader.
# The following demo shows how to load and use the module using jp_proxy_widgets.
#
# A quick demo:
# +
import jp_proxy_widget
cdn_url = "https://cdn.jsdelivr.net/npm/binary-com-jquery-dialogextended@1.0.0/jquery.dialogextend.js"
module_id = "dialogExtend"
# Load the module using a widget (any widget -- the module loads to the global jQuery object).
loader = jp_proxy_widget.JSProxyWidget()
# Configure the module to be loaded.
loader.require_js(module_id, cdn_url)
# Load the module
loader.js_init("""
element.requirejs([module_identifier], function(module_value) {
element.html("loaded " + module_identifier + " : " + module_value);
});
""", module_identifier=module_id)
loader
# +
widget = jp_proxy_widget.JSProxyWidget()
#widget.element.html("Hello world")
# xxxx I'm not sure why this didn't work...
#widget.element.dialog({ "title" : "Your Title" }).dialogExtend({
# "maximizable" : True,
# "dblclick" : "maximize",
# "icons" : { "maximize" : "ui-icon-arrow-4-diag" },
# })
widget.js_init("""
// but the Javascript equivalent works.
element.html("Hello world")
element.dialog({ "title" : "Your Title" }).dialogExtend({
"maximizable" : true,
"dblclick" : "maximize",
"icons" : { "maximize" : "ui-icon-arrow-4-diag" },
});
""")
widget
# -
| notebooks/misc/JQueryUI dialogextend plugin demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="wYtuKeK0dImp"
# ATTENTION: Please do not alter any of the provided code in the exercise. Only add your own code where indicated
# ATTENTION: Please do not add or remove any cells in the exercise. The grader will check specific cells based on the cell position.
# ATTENTION: Please use the provided epoch values when training.
import csv
import numpy as np
import tensorflow as tf
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from os import getcwd
# + colab={} colab_type="code" id="4kxw-_rmcnVu"
def get_data(filename):
# You will need to write code that will read the file passed
# into this function. The first line contains the column headers
# so you should ignore it
# Each successive line contians 785 comma separated values between 0 and 255
# The first value is the label
# The rest are the pixel values for that picture
# The function will return 2 np.array types. One with all the labels
# One with all the images
#
# Tips:
# If you read a full line (as 'row') then row[0] has the label
# and row[1:785] has the 784 pixel values
# Take a look at np.array_split to turn the 784 pixels into 28x28
# You are reading in strings, but need the values to be floats
# Check out np.array().astype for a conversion
with open(filename) as training_file:
# Your code starts here
images = []
labels = []
contents = training_file.readlines()
contents.remove(contents[0])
for item in contents:
item = item.split(',')
labels.append(item[0])
images.append(np.array_split(item[1:785],28))
labels = np.array(labels, dtype = float)
images = np.array(images, dtype = float)
# Your code ends here
return images, labels
path_sign_mnist_train = f"{getcwd()}/../tmp2/sign_mnist_train.csv"
path_sign_mnist_test = f"{getcwd()}/../tmp2/sign_mnist_test.csv"
training_images, training_labels = get_data(path_sign_mnist_train)
testing_images, testing_labels = get_data(path_sign_mnist_test)
# Keep these
print(training_images.shape)
print(training_labels.shape)
print(testing_images.shape)
print(testing_labels.shape)
# Their output should be:
# (27455, 28, 28)
# (27455,)
# (7172, 28, 28)
# (7172,)
# + colab={} colab_type="code" id="awoqRpyZdQkD"
# In this section you will have to add another dimension to the data
# So, for example, if your array is (10000, 28, 28)
# You will need to make it (10000, 28, 28, 1)
# Hint: np.expand_dims
training_images = np.expand_dims(training_images,axis = training_images.ndim)# Your Code Here
testing_images = np.expand_dims(testing_images, axis = testing_images.ndim) # Your Code Here
# Create an ImageDataGenerator and do Image Augmentation
train_datagen = ImageDataGenerator(
rescale = 1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest'
# Your Code Here
)
validation_datagen = ImageDataGenerator(rescale = 1./255.
# Your Code Here
)
# Keep These
print(training_images.shape)
print(testing_images.shape)
# Their output should be:
# (27455, 28, 28, 1)
# (7172, 28, 28, 1)
# + colab={} colab_type="code" id="Rmb7S32cgRqS"
# Define the model
# Use no more than 2 Conv2D and 2 MaxPooling2D
model = tf.keras.models.Sequential([
# Your Code Here
tf.keras.layers.Conv2D(64,(3,3),activation = 'relu',input_shape = (28,28,1)),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(64,(3,3),activation = 'relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(256,activation = 'relu'),
tf.keras.layers.Dense(26,activation = 'softmax')
])
# Compile Model.
model.compile(loss = 'sparse_categorical_crossentropy',
optimizer = 'Adam',
metrics = ['accuracy'])
# Train the Model
history = model.fit_generator(train_datagen.flow(training_images, training_labels, batch_size=32),
steps_per_epoch=len(training_images) / 32,
epochs=2,
validation_data=validation_datagen.flow(testing_images, testing_labels, batch_size=32),
validation_steps=len(testing_images) / 32)
model.evaluate(testing_images, testing_labels, verbose=0)
# + colab={} colab_type="code" id="_Q3Zpr46dsij"
# Plot the chart for accuracy and loss on both training and validation
# %matplotlib inline
import matplotlib.pyplot as plt
acc = history.history['accuracy']# Your Code Here
val_acc =history.history['val_accuracy'] # Your Code Here
loss = history.history['loss']# Your Code Here
val_loss = history.history['val_loss']# Your Code Here
epochs = range(len(acc))
plt.plot(epochs, acc, 'r', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'r', label='Training Loss')
plt.plot(epochs, val_loss, 'b', label='Validation Loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# -
# # Submission Instructions
# +
# Now click the 'Submit Assignment' button above.
# -
# # When you're done or would like to take a break, please run the two cells below to save your work and close the Notebook. This will free up resources for your fellow learners.
# + language="javascript"
# <!-- Save the notebook -->
# IPython.notebook.save_checkpoint();
# + language="javascript"
# IPython.notebook.session.delete();
# window.onbeforeunload = null
# setTimeout(function() { window.close(); }, 1000);
| Coursera/Exercise_4_Multi_class_classifier_Question-My_Resolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Smoketest for exercises in Informationvisualisation IVIS
#
# This test installs an verifies the proper functioning of the exercices.
# In case of any issue send me a mail (<NAME> at FHNW) or post a message in Teams,
#
# The test can be run
# * on mybinder.org or
# * on an Anaconda JupiterLab installation or
# * in a Python Notebook
#
# ## Setup for mybinder.org
#
# Nothing to do. Skip to "[Test Altair](#Test-Altair)".
#
# ## Setup for DeepNote.com
#
# Nothing to do. Skip to "[Test Altair](#Test-Altair)".
#
#
# ## Setup for Anaconda
#
# ### Prerequisites
#
# 1. Install [Anaconda](https://www.anaconda.com/products/individual).
# 2. Start Anaconda Navigator
#
#
# ### JupyterLab
#
# 3. Start JupyterLab
# 4. Load this smoketest.ipynb
# 5. Uncomment the statements in the following cell
# 6. Execute smoketest.ipynb
#
# +
#################################################################################
## NOTE: This code has been tested with JupyterLab installed through Anaconda. ##
## It may fail in other environments. ##
## Uncomment the statements (remove #)
#################################################################################
# import sys
# # !conda install --yes --prefix {sys.prefix} -c conda-forge altair vega-datasets vega
# now wait until cell finished loading
# -
# ### JupyterNotebook
#
# 3. Install `altair and vega-datasets` (You can uses either 'conda' or `pip` for this).
#
# # Test Altair
#
# 1. Execute the following cells
# 2. The response should look as follows <br/>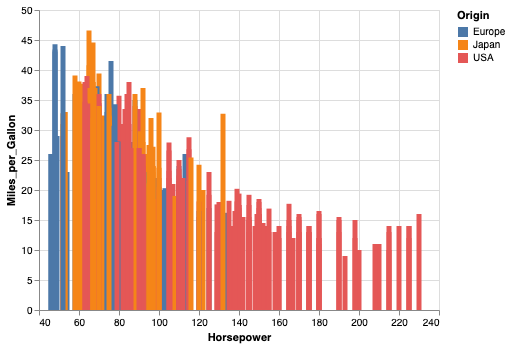
# 3. Get back to me if this does not work.
import altair as alt
# load a simple dataset as a pandas DataFrame
from vega_datasets import data
cars = data.cars()
alt.Chart(cars).mark_bar().encode(
x='Horsepower',
y='Miles_per_Gallon',
color='Origin',
).interactive()
# # Test Vega-Lite
#
# 1. Execute the following 2 cells.
# 2. The result should look like this:<br/>
# 
#
import altair as alt
alt.Chart.from_dict({
"$schema": "https://vega.github.io/schema/vega-lite/v4.json",
"data": {
"url": "https://gist.githubusercontent.com/marcosoldati/37b652aff7476c6ab77f16f4f808b243/raw/6c2125013811db238170f5dc0d728e858e40989d/olympics.csv"
},
"mark": "tick",
"encoding": {
"x": {
"field": "Year",
"type": "temporal",
}
}
})
# # PyPlot testen
#
# Matplotlib `Pyplot` should be already installed an can be use right away.
import matplotlib.pyplot as plt
plt.plot([1, 2, 3, 4], [1, 4, 9, 16])
plt.ylabel('some numbers')
plt.axis([1, 4, 0, 20])
plt.show()
# # Seaborn testen
#
# `Seaborn` bases on `matlibplot` and is already installed in most setups. Seaborn is not required for these exercises.
import seaborn as sns
# +
# Load an example dataset
tips = sns.load_dataset("tips")
# Create a visualization
sns.relplot(
data=tips,
x="total_bill", y="tip", col="time",
hue="smoker", style="smoker", size="size",
)
# -
| 0_smoketest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Normalized Sense Function
#
# In this notebook, let's go over the steps a robot takes to help localize itself from an initial, uniform distribution to sensing and updating that distribution and finally normalizing that distribution.
#
# 1. The robot starts off knowing nothing; the robot is equally likely to be anywhere and so `p` is a uniform distribution.
# 2. Then the robot senses a grid color: red or green, and updates this distribution `p` according to the values of pHit and pMiss.
# 3. **We normalize `p` such that its components sum to 1.**
#
# <img src='images/robot_sensing.png' width=50% height=50% />
#
# importing resources
import matplotlib.pyplot as plt
import numpy as np
# A helper function for visualizing a distribution.
def display_map(grid, bar_width=1):
if(len(grid) > 0):
x_labels = range(len(grid))
plt.bar(x_labels, height=grid, width=bar_width, color='b')
plt.xlabel('Grid Cell')
plt.ylabel('Probability')
plt.ylim(0, 1) # range of 0-1 for probability values
plt.title('Probability of the robot being at each cell in the grid')
plt.xticks(np.arange(min(x_labels), max(x_labels)+1, 1))
plt.show()
else:
print('Grid is empty')
# ### QUIZ: Modify your code so that it normalizes the output for the sense function.
#
# This means that the entries in `q` should sum to one.
#
# Note that `pHit` refers to the probability that the robot correctly senses the color of the square it is on, so if a robot senses red *and* is on a red square, we'll multiply the current location probability (0.2) with pHit. Same goes for if a robot senses green *and* is on a green square.
# +
# given initial variables
p=[0.2, 0.2, 0.2, 0.2, 0.2]
# the color of each grid cell in the 1D world
world=['green', 'red', 'red', 'green', 'green']
# Z, the sensor reading ('red' or 'green')
Z = 'red'
pHit = 0.6
pMiss = 0.2
## Complete this function
def sense(p, Z):
''' Takes in a current probability distribution, p, and a sensor reading, Z.
Returns a *normalized* distribution after the sensor measurement has been made, q.
This should be accurate whether Z is 'red' or 'green'. '''
q=[]
##TODO: normalize q
# loop through all grid cells
for i in range(len(p)):
# check if the sensor reading is equal to the color of the grid cell
# if so, hit = 1
# if not, hit = 0
hit = (Z == world[i])
q.append(p[i] * (hit * pHit + (1-hit) * pMiss))
q = [i/sum(q) for i in q]
return q
q = sense(p,Z)
print(q)
display_map(q)
# -
| Object Tracking and Localization/Robot Localization/Normalized Sense Function/Normalized Sense Function, exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
#open file
file = open('../AData/energy.txt','r')
#read from the file
text = file.readlines()
energy = []
for line in text:
energy.append(float(line[:-1]))
file.close()
#
#open file
file = open('../AData/temperatur.txt','r')
#read from the file
text = file.readlines()
temperatur = []
for line in text:
temperatur.append(float(line[:-1]))
#visulize
plt.rcParams['figure.figsize'] = [16,9]
f = plt.figure()
ax = f.add_subplot(111)
plt.text(0.2,0.9,'First try. probably not relaxed enough, the p structur needs a rework',horizontalalignment='center',verticalalignment='center', transform = ax.transAxes)
plt.plot(temperatur,energy)
plt.xlabel("Temperatur [K]")
plt.ylabel("Energy [eV]")
plt.savefig('plot_m7firstTry.png')
plt.show()
| AJupyter/energyVisulizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mariellemiron/Linear-Algebra_ChE_2nd-Sem-2021-2022/blob/main/Assignment_6.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="1eV26_aYTrTA"
# # Linear Algebra for ChE
# ## Laboratory 6 : Matrix Operations
# + [markdown] id="zlGP-Q4BTvcu"
# ## Objectives
# In this activity, the students will be able to acquire the follow:
# 1. Familiarize fundamental matrix operations.
# 2. Apply the operations to solve intermediate equations.
# 3. Apply matrix algebra in engineering solutions.
# + [markdown] id="3_IhtAZ1Tfi0"
# ## Discussion
# + id="CCQkS1eLN6xI"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] id="DAPjO3M1T6cM"
# ## Transposition
# + [markdown] id="jtvvEY-w5Ru5"
# Transposition is a fundamental operation in matrix algebra. The values of a matrix's elements are flipped over its diagonals to transpose it. The rows and columns of the original matrix will be switched as a result of this. As a result, the transpose of a matrix $C$ is denoted by $C^T$. This may now be done programmatically with `np.transpose()` or the `T` function. For instance:
# + [markdown] id="IGa-28MkT-vf"
# $$A = \begin{bmatrix} 5 & -1 & 2\\0 & -3 &2 \\ 1 & 3 & -9\end{bmatrix} $$
# + [markdown] id="aPnbLGE9UMqh"
# $$ A^T = \begin{bmatrix} 5 & 0 & 1\\-1 & -3 &3 \\ 2& 2 & -9\end{bmatrix}$$
# + colab={"base_uri": "https://localhost:8080/"} id="FUAFw0NlVQZk" outputId="e593b61e-4862-49a9-ca4c-8f9e8176f4a7"
A = np.array([
[2 ,5, 0],
[4, -9, 3],
[1, 1, 7]
])
A
# + colab={"base_uri": "https://localhost:8080/"} id="5diWWrhkVWeP" outputId="8a586961-535d-46df-8778-2c36ea8e5555"
AT1 = np.transpose(A)
AT1
# + colab={"base_uri": "https://localhost:8080/"} id="kAcy10CZVg7r" outputId="e917e456-5733-4665-86a1-774390c02106"
AT2 = A.T
AT2
# + colab={"base_uri": "https://localhost:8080/"} id="jGbPUgKzWEj1" outputId="f53ec4b6-8973-4a05-9d98-5e9475724d6c"
np.array_equiv(AT1, AT2)
# + colab={"base_uri": "https://localhost:8080/"} id="N-1_oLM9WHGd" outputId="91208071-0140-4109-ed64-53ecaac9d14e"
B = np.array([
[9,1,-9],
[5,-8,5],
])
B.shape
# + colab={"base_uri": "https://localhost:8080/"} id="fx7M78ZFWR74" outputId="b7990e31-a712-4700-d88d-38fb901b42bb"
np.transpose(B).shape
# + colab={"base_uri": "https://localhost:8080/"} id="3tG5mcGbWUE0" outputId="2950b050-e032-46f6-e033-9f57f926ee2e"
B.T.shape
# + [markdown] id="JK9LteIDWgLW"
# ## Test / Trial
# + colab={"base_uri": "https://localhost:8080/"} id="KFss-T2OWhVS" outputId="a8ac1a34-50ab-4d73-9d66-068b1e21a971"
M=np.array([
[4,6,4,5],
[8,9,7,-9]
])
M.shape
# + colab={"base_uri": "https://localhost:8080/"} id="6BoN7ysKWwwT" outputId="84d3da79-7cde-4faf-9b78-f7dc92dc67fb"
np.transpose(M).shape
# + colab={"base_uri": "https://localhost:8080/"} id="3RhpBPebW0FZ" outputId="4582e5d6-4cb4-4808-814c-20f8999d7974"
M.T.shape
# + colab={"base_uri": "https://localhost:8080/"} id="iQYqGjx_W2sD" outputId="5d453a5d-20f6-4e83-c4ea-dc6b3935301e"
MT = M.T
MT
# + [markdown] id="zUI2M_OAXEDr"
# ## Dot Product / Inner Product
# + [markdown] id="sVI2NfzpXNq_"
# If you remember the dot product from the laboratory activity, we'll try to do the same thing with matrices. We'll retrieve the sum of products of the vectors by row-column pairings in matrix dot product. So, if we have two matrices, $X$ and $Y$, we can deduce:
#
# $$X = \begin{bmatrix}x_{(0,0)}&x_{(0,1)}\\ x_{(1,0)}&x_{(1,1)}\end{bmatrix}, Y = \begin{bmatrix}y_{(0,0)}&y_{(0,1)}\\ y_{(1,0)}&y_{(1,1)}\end{bmatrix}$$
#
# The dot product will then be computed as:
# $$X \cdot Y= \begin{bmatrix} x_{(0,0)}*y_{(0,0)} + x_{(0,1)}*y_{(1,0)} & x_{(0,0)}*y_{(0,1)} + x_{(0,1)}*y_{(1,1)} \\ x_{(1,0)}*y_{(0,0)} + x_{(1,1)}*y_{(1,0)} & x_{(1,0)}*y_{(0,1)} + x_{(1,1)}*y_{(1,1)}
# \end{bmatrix}$$
#
# So if we assign values to $X$ and $Y$:
# $$X = \begin{bmatrix}1&2\\ 0&1\end{bmatrix}, Y = \begin{bmatrix}-1&0\\ 2&2\end{bmatrix}$$
#
# $$X \cdot Y= \begin{bmatrix} 1*-1 + 2*2 & 1*0 + 2*2 \\ 0*-1 + 1*2 & 0*0 + 1*2 \end{bmatrix} = \begin{bmatrix} 3 & 4 \\2 & 2 \end{bmatrix}$$
#
# Using np.dot(), np.matmul(), or the @ operator.
# + id="EujOgrumXSx5"
Q = np.array([
[5,4],
[2,0]
])
W = np.array([
[-5,-4],
[-2,2]
])
# + colab={"base_uri": "https://localhost:8080/"} id="Sh1lQJi1XV7L" outputId="837c3155-de39-4667-9cb6-bf37fa05121c"
np.array_equiv(Q, W)
# + colab={"base_uri": "https://localhost:8080/"} id="ki309bQlXhNb" outputId="1bdd5be9-7829-45f7-f377-6b0a218b843c"
np.dot(Q,W)
# + colab={"base_uri": "https://localhost:8080/"} id="84KaZ1adXjV3" outputId="357d2ab1-f9a2-427d-dca1-425ee98d060f"
Q.dot(W)
# + colab={"base_uri": "https://localhost:8080/"} id="VCk3HBuJXk-h" outputId="5bf327e1-28b7-4a13-f793-06701782f047"
Q @ W
# + colab={"base_uri": "https://localhost:8080/"} id="0D68lnSRXm6F" outputId="1d3263d1-7b34-46a6-cfd3-8e500786843f"
np.matmul(Q,W)
# + id="5WX745psXo5y"
D = np.array([
[0,1,2],
[-3,-4,-5],
[6,7,8]
])
K = np.array([
[-9,-10,-11],
[0,1,2],
[-3,4,-5]
])
# + colab={"base_uri": "https://localhost:8080/"} id="YlP2GwRJX255" outputId="f6b473f2-ccbf-4416-e5e0-d451ee762579"
D @ K
# + colab={"base_uri": "https://localhost:8080/"} id="QGTfSN-pX4ky" outputId="2a77ca2b-5fbb-410b-af0c-179abdb7f11c"
D.dot(K)
# + colab={"base_uri": "https://localhost:8080/"} id="9upWySebYDmE" outputId="f8d46ec8-0fcb-42bb-f563-e03d23725792"
np.matmul(D, K)
# + colab={"base_uri": "https://localhost:8080/"} id="95_N8Ov0YGo2" outputId="60616806-77ec-4c20-c0df-45ace5b07606"
np.dot(D, K)
# + [markdown] id="YXZ3DCgM4pMq"
# In comparison to vector dot products, matrix dot products have additional rules. There are fewer limits because vector dot products are only one dimensional. Since we're dealing with Rank 2 vectors, there are a few rules to keep in mind:
# + [markdown] id="tQKMcXP2ZZcU"
# ### Rule 1: The inner dimensions of the two matrices in question must be the same.
# + [markdown] id="4Wf3XD724tOI"
# Assume you have a matrix A with the shape $(a,b)$, where $a$ and $b$ are any integers. Matrix $B$ should have the shape $(b,c)$, where $b$ and $c$ are any integers, if we want to do a dot product between A and another matrix $B$. As a result, for the following matrices:
#
# $$A = \begin{bmatrix}2&4\\5&-2\\0&1\end{bmatrix}, B = \begin{bmatrix}1&1\\3&3\\-1&-2\end{bmatrix}, C = \begin{bmatrix}0&1&1\\1&1&2\end{bmatrix}$$
#
# So in this case $A$ has a shape of $(3,2)$, $B$ has a shape of $(3,2)$ and $C$ has a shape of $(2,3)$. So the only matrix pairs that is eligible to perform dot product is matrices $A \cdot C$, or $B \cdot C$.
# + colab={"base_uri": "https://localhost:8080/"} id="BMg10X5gZc6n" outputId="f8d5ecf9-0426-4109-c045-d79e2f7c6655"
X = np.array([
[9, 9,9,4],
[8, -7,4,7],
[-5, 6,2,1]
])
Y = np.array([
[5,-5,7,4],
[4,-3,1,5],
[2,0,-1,9]
])
Z = np.array([
[0,1,1],
[1,1,2],
[0,5,9],
[7,1,2]
])
print(X.shape)
print(Y.shape)
print(Z.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="DSJ3OF_ZZyDK" outputId="fcfd6b3b-b728-4d06-8561-20d3e9e22e85"
X @ Z
# + colab={"base_uri": "https://localhost:8080/"} id="RQRziWdAZ7LF" outputId="4321de28-9000-4018-ce62-95b30829317f"
Y @ Z
# + colab={"base_uri": "https://localhost:8080/", "height": 183} id="5-u6vlwvaFHp" outputId="5efb5c1c-60fb-4947-e0d2-e61f4fe6dff0"
X @ Y
# + [markdown] id="T__mvkEgaR4c"
# If you would notice the shape of the dot product changed and its shape is not the same as any of the matrices we used. The shape of a dot product is actually derived from the shapes of the matrices used. So recall matrix $A$ with a shape of $(a,b)$ and matrix $B$ with a shape of $(b,c)$, $A \cdot B$ should have a shape $(a,c)$.
# + colab={"base_uri": "https://localhost:8080/"} id="NTSDwVcDaSgn" outputId="e00d0206-67ad-4af5-a7a2-bf0951dc16c9"
X @ Y.T
# + colab={"base_uri": "https://localhost:8080/"} id="RGmTHgueaYPd" outputId="e04c5568-2a7d-49ff-a8f2-d734dee924ae"
M = np.array([
[9,-8,7,-6]
])
G = np.array([
[5,-4,3,-2]
])
print(M.shape)
print(G.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="jGG9HpIIakmA" outputId="4039e86b-8cde-420a-8b08-51a4bd57b0c5"
G.T @ M
# + colab={"base_uri": "https://localhost:8080/"} id="2Nrl1vU9an8j" outputId="1d30a760-bb8e-4a3f-e072-f9f2ff31b9cc"
M @ G.T
# + [markdown] id="dIHpZyJ6axeY"
# ### Rule 2: Dot Product has special properties
# + [markdown] id="fePYlNmV5aia"
# Dot products are common in matrix algebra, which means they have various distinct qualities that should be taken into account while formulating solutions:
# 1. $A \cdot B \neq B \cdot A$
# 2. $A \cdot (B \cdot C) = (A \cdot B) \cdot C$
# 3. $A\cdot(B+C) = A\cdot B + A\cdot C$
# 4. $(B+C)\cdot A = B\cdot A + C\cdot A$
# 5. $A\cdot I = A$
# 6. $A\cdot \emptyset = \emptyset$
# + id="i-ZAKu3oa0pA"
A = np.array([
[1,2,3],
[4,5,6],
[9,8,7]
])
B = np.array([
[9,7,8],
[8,7,9],
[2,3,5]
])
C = np.array([
[8,7,8],
[6,5,4],
[3,8,7]
])
# + colab={"base_uri": "https://localhost:8080/"} id="fAR12wUAbMcY" outputId="5be759d4-c241-4fe2-e8da-529ddcd09067"
np.eye(3)
# + colab={"base_uri": "https://localhost:8080/"} id="msNDw69tbPLk" outputId="31dd63a3-58a0-4757-ec46-eef47d225cb0"
A.dot(np.eye(3))
# + colab={"base_uri": "https://localhost:8080/"} id="MjI4qgEfbRAa" outputId="0f253f87-cb93-45a3-ef0a-59e46319f848"
np.array_equal(A@B, B@A)
# + colab={"base_uri": "https://localhost:8080/"} id="SpUPRuhBbUDx" outputId="50759849-3ab6-46d7-f4a9-572ad271ca01"
D = A @ (B @ C)
D
# + colab={"base_uri": "https://localhost:8080/"} id="JaIroI54bZNV" outputId="c717aa96-05f4-4dd2-9491-74b9dca7ec12"
E = (A @ B) @ C
E
# + colab={"base_uri": "https://localhost:8080/"} id="mxtgE3IUbh8s" outputId="633e8232-f8dd-42c3-a7e8-664a0d5a0992"
np.array_equal(E, M)
# + colab={"base_uri": "https://localhost:8080/"} id="smCwXludbrWU" outputId="84a114fe-83a6-4271-fb87-edc0e13cddd8"
np.array_equiv(D, E)
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="UxjDsKxvb0ot" outputId="ef43e69c-e343-4df5-ab1e-6b82fd36052c"
np.eye(A)
# + colab={"base_uri": "https://localhost:8080/"} id="7WEnvKOmb5Yh" outputId="04b31170-0a7a-491c-f5b9-a6bd61880a70"
A @ D
# + colab={"base_uri": "https://localhost:8080/"} id="RwZ3Ei0tb8LJ" outputId="b4d97b44-acaf-4069-c90c-2ed979337b7e"
z_mat = np.zeros(A.shape)
z_mat
# + colab={"base_uri": "https://localhost:8080/"} id="13Lwd9DSb-28" outputId="aee1b31d-f5e9-4fdd-8704-5d1d45bc0b1b"
a_dot_z = A.dot(np.zeros(A.shape))
a_dot_z
# + colab={"base_uri": "https://localhost:8080/"} id="BACWoiUfcBhN" outputId="11792d27-ffd6-4c38-b8dc-cec058ad344d"
np.array_equal(a_dot_z,z_mat)
# + colab={"base_uri": "https://localhost:8080/"} id="ntpINHu7cDcN" outputId="27483e0f-ecb7-429d-a5e1-c9886045a0e2"
null_mat = np.empty(A.shape, dtype=float)
null = np.array(null_mat,dtype=float)
print(null)
np.allclose(a_dot_z,null)
# + [markdown] id="x3CzoJnmcGwm"
# ## Determinant
# + [markdown] id="3G7GfxvtcJBg"
# A determinant is a scalar value that can be calculated using a square matrix. In matrix algebra, the determinant is a fundamental and crucial value. Although it will not be clear how it may be utilized realistically in this laboratory, it will be extensively employed in future lessons.
#
# The determinant of some matrix $A$ is denoted as $det(A)$ or $|A|$. So let's say $A$ is represented as:
# $$A = \begin{bmatrix}a_{(0,0)}&a_{(0,1)}\\a_{(1,0)}&a_{(1,1)}\end{bmatrix}$$
# We can compute for the determinant as:
# $$|A| = a_{(0,0)}*a_{(1,1)} - a_{(1,0)}*a_{(0,1)}$$
# So if we have $A$ as:
# $$A = \begin{bmatrix}1&4\\0&3\end{bmatrix}, |A| = 3$$
#
# But what about square matrices that aren't in the shape (2,2)? Several strategies, such as co-factor expansion and the minors method, can be used to solve this problem. This can be taught in a laboratory lecture, but we can use Python to perform the difficult computation of high-dimensional matrices programmatically. Using `np.linalg.det()`, we can accomplish this.
# + colab={"base_uri": "https://localhost:8080/"} id="OUgDmBc7cIcR" outputId="aecc8179-c5bf-4c17-e33c-61a6c113dabf"
S = np.array([
[3,2],
[3,4]
])
np.linalg.det(S)
# + colab={"base_uri": "https://localhost:8080/"} id="mFtmtGl-cVVQ" outputId="2ad71090-801f-44ff-f493-f7d4a4aee126"
V = np.array([
[4, 5, 6],
[9, 3 ,2],
[1, -2, -1]
])
np.linalg.det(V)
# + colab={"base_uri": "https://localhost:8080/"} id="_2d1OhKddc0U" outputId="7d5164f4-3d8f-4a4b-f9df-fc7bd1bc4e53"
T = np.array([
[1,2,3,8],
[3,4,6,4],
[1,6,9,2],
[0,8,3,0]
])
np.linalg.det(T)
# + [markdown] id="MoWnj2Xvd6Fv"
# ## Inverse
# + [markdown] id="muAsmUWqd9Z1"
# Another essential operation in matrix algebra is the inverse of a matrix. We can identify a matrix's solvability and characteristic as a system of linear equations by determining its inverse — we'll go over this more in the nect module. The inverse matrix can also be used to solve the problem of divisibility amongst matrices. Although element-by-element division is possible, dividing matrices as a whole is not. Inverse matrices allow a similar process that might be thought of as "splitting" matrices.
#
# Now to determine the inverse of a matrix we need to perform several steps. So let's say we have a matrix $M$:
# $$M = \begin{bmatrix}1&7\\-3&5\end{bmatrix}$$
# First, we need to get the determinant of $M$.
# $$|M| = (1)(5)-(-3)(7) = 26$$
# Next, we need to reform the matrix into the inverse form:
# $$M^{-1} = \frac{1}{|M|} \begin{bmatrix} m_{(1,1)} & -m_{(0,1)} \\ -m_{(1,0)} & m_{(0,0)}\end{bmatrix}$$
# So that will be:
# $$M^{-1} = \frac{1}{26} \begin{bmatrix} 5 & -7 \\ 3 & 1\end{bmatrix} = \begin{bmatrix} \frac{5}{26} & \frac{-7}{26} \\ \frac{3}{26} & \frac{1}{26}\end{bmatrix}$$
# For higher-dimension matrices you might need to use co-factors, minors, adjugates, and other reduction techinques. To solve this programmatially we can use `np.linalg.inv()`.
# + colab={"base_uri": "https://localhost:8080/"} id="Rx0oLalAeA32" outputId="a627d71a-76c9-419a-c5fe-ddca6e3f73f0"
C = np.array([
[8,6],
[6,-8]
])
np.array(C @ np.linalg.inv(C), dtype=int)
# + colab={"base_uri": "https://localhost:8080/"} id="TSMGXU47eDce" outputId="1cdfcf30-fc72-4b11-8491-31ed56a4e29c"
R = np.array([
[1, 3, 5],
[7, 9, 2],
[-4, 6, 8]
])
T = np.linalg.inv(R)
T
# + colab={"base_uri": "https://localhost:8080/"} id="DAlgu5pteFYp" outputId="e34f1bfa-415c-44f6-8f53-a1345012055f"
R @ T
# + colab={"base_uri": "https://localhost:8080/"} id="tTGGeG1aeG1n" outputId="2b30437c-01de-44fb-f3a7-21364d82c84f"
M = np.array([
[1,2,2,3,4,4,5],
[20,6,6,7,8,8,9],
[0,1,1,2,3,3,4],
[10,11,11,10,9,9,11],
[9,16,8,9,7,7,8],
[-1,13,13,0,0,1,-1],
[20,3,0,4,5,15,12],
])
M_inv = np.linalg.inv(M)
np.array(M @ M_inv,dtype=int)
# + [markdown] id="Zcza-ZPveLJZ"
# To validate the wether if the matrix that you have solved is really the inverse, we follow this dot product property for a matrix $M$:
#
# $$M\cdot M^{-1} = I$$
# + colab={"base_uri": "https://localhost:8080/"} id="Q_Q__wjCeL6M" outputId="41024cd6-3b44-4dd2-e3c7-c3f08ce04786"
squad = np.array([
[1.2, 1.0, 0.8],
[0.5, 0.3, 1.0],
[0.9, 0.5, 1.9]
])
weights = np.array([
[0.5, 0.6, 0.7]
])
p_grade = squad @ weights.T
p_grade
# + [markdown] id="YWAKrwXJhnIG"
# ## Activity
# + [markdown] id="4CEFy5y7ho4A"
# ### Task 1
# + [markdown] id="-mbKmYAshrX8"
# Prove and implement the remaining 6 matrix multiplication properties. You may create your own matrices in which their shapes should not be lower than $(3,3)$.
# In your methodology, create individual flowcharts for each property and discuss the property you would then present your proofs or validity of your implementation in the results section by comparing your result to present functions from NumPy.
#
# 1. $A \cdot B \neq B \cdot A$
# 2. $A \cdot (B \cdot C) = (A \cdot B) \cdot C$
# 3. $A\cdot(B+C) = A\cdot B + A\cdot C$
# 4. $(B+C)\cdot A = B\cdot A + C\cdot A$
# 5. $A\cdot I = A$
# 6. $A\cdot \emptyset = \emptyset$
# + id="eh0glBqpmIHg"
D = np.array([
[8, 4, 24],
[8, 6, -5],
[9, 1, 15]
])
O = np.array([
[17, 35, 79],
[6, 51, 5],
[38, -1, -6]
])
G = np.array([
[23, 70, -34],
[24, -9, 8],
[34, 2, 46]
])
# + [markdown] id="UbF4g5XqpB5P"
# Property no.1: $A \cdot B \neq B \cdot A$
# + colab={"base_uri": "https://localhost:8080/"} id="xf0r3ys6mO-1" outputId="7f88c606-1244-454f-f25a-9693abaab8bc"
W = D @ O
W
# + colab={"base_uri": "https://localhost:8080/"} id="S4xlVcYWmtSY" outputId="bf7e508a-e6fe-41b5-d7a4-af3d888d4bdf"
E = O @ D
E
# + colab={"base_uri": "https://localhost:8080/"} id="RiSBsVI7mvc_" outputId="3ec9e0a9-dba2-44ad-9bf7-346d76ce87b5"
np.array_equiv(W, E)
# + [markdown] id="qNsYJ6DKpL9O"
# Property no.2: $A \cdot (B \cdot C) = (A \cdot B) \cdot C$
# + colab={"base_uri": "https://localhost:8080/"} id="nLQ5ObZPmw3B" outputId="e8b593d3-1e27-44fc-d989-cbfbf954adf2"
M = D @ (O @ G)
M
# + colab={"base_uri": "https://localhost:8080/"} id="o8Rc20hlmz4C" outputId="f813336c-4182-41af-b7b8-e10e37b976ef"
Y = (D @ O) @ G
Y
# + colab={"base_uri": "https://localhost:8080/"} id="wooC77rzm9YI" outputId="07780625-8dcc-4505-b092-e6005578ad38"
np.array_equiv(M,Y)
# + [markdown] id="JzGFeqyIpU67"
# Property no.3: $A\cdot(B+C) = A\cdot B + A\cdot C$
# + colab={"base_uri": "https://localhost:8080/"} id="QGKOiOUtnAo6" outputId="f160eaaa-f11e-49a2-9d2c-483ac6283276"
I = D @ (O + G)
I
# + colab={"base_uri": "https://localhost:8080/"} id="_0sZ993OnQA0" outputId="0305b6fd-f5be-4c61-a4a5-fc88db8744bc"
S = D @ O + D @ G
S
# + colab={"base_uri": "https://localhost:8080/"} id="P3NDB6oLnSYt" outputId="98059fc3-dea1-42d2-9687-df8e81ebf60d"
np.array_equiv(I, S)
# + [markdown] id="holcdFD4prrq"
# Propery no.4: $(B+C)\cdot A = B\cdot A + C\cdot A$
# + colab={"base_uri": "https://localhost:8080/"} id="9QmbyW5anUeh" outputId="11387ebd-a164-4028-a706-b85251afd702"
L = (O + G) @ D
L
# + colab={"base_uri": "https://localhost:8080/"} id="Dki-GGvonZlG" outputId="58c29fb5-ff30-40a2-f3af-f2552e4dd510"
U = O @ D + G @ D
U
# + colab={"base_uri": "https://localhost:8080/"} id="JcAX0s66ndgm" outputId="af5329e4-cf8e-4871-f520-4dd68df0d908"
np.array_equiv(L, U)
# + [markdown] id="S9u1rwgupzf_"
# Property no.5: $A\cdot I = A$
# + colab={"base_uri": "https://localhost:8080/"} id="kuJHwKWSnhGC" outputId="a7315819-bb91-44c8-8a5b-9b91414b0a78"
D @ np.eye(3)
# + colab={"base_uri": "https://localhost:8080/"} id="5e15tFrjnkCQ" outputId="f303ec55-81b7-4035-c13e-1f0679b9f84a"
np.array_equiv(D, D @ np.eye(3))
# + [markdown] id="CxDPe2c3r36h"
# Property no.6: $A\cdot \emptyset = \emptyset$
# + colab={"base_uri": "https://localhost:8080/"} id="b0vqZpzfnoAQ" outputId="59e10462-ecec-4a8b-ce1b-6d77fb307a0d"
D @ np.zeros ((3,3))
# + colab={"base_uri": "https://localhost:8080/"} id="2pHFMNwGnqVd" outputId="82a693d8-3c86-4186-cbef-4b078dd5476e"
np.array_equiv(D@np.zeros((3,3)), np.zeros((3,3)))
# + colab={"base_uri": "https://localhost:8080/"} id="IeM7wxFHnseR" outputId="1bc6b0c5-89c3-499b-a562-88cb619c84bf"
D.dot(np.zeros(D.shape))
# + colab={"base_uri": "https://localhost:8080/"} id="UVQ2WnhBnuWh" outputId="ded47880-21b7-42c5-f4a3-1ceb4ced4799"
z_mat = np.zeros (D.shape)
z_mat
# + colab={"base_uri": "https://localhost:8080/"} id="HV3vLzdPnwHY" outputId="10551e7f-abf6-489a-938d-716cef40bbc9"
i_dot_z = D.dot(np.zeros(D.shape))
i_dot_z
# + colab={"base_uri": "https://localhost:8080/"} id="00U76jzdnyPM" outputId="9435d981-09bc-48e8-9dac-b92f7f67bf19"
np.array_equal(i_dot_z,z_mat)
# + colab={"base_uri": "https://localhost:8080/"} id="nAON8fP2nzWX" outputId="030a43ab-efcf-4149-8517-68fce55c11e3"
null_mat = np.empty(D.shape, dtype = float)
null = np.array(null_mat,dtype = float)
print(null)
np.allclose(i_dot_z , null)
| Assignment_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import boto3
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from s3ssp import get_file
s3 = boto3.resource('s3')
get_file(s3, "s3ssp", download_file="Analysis_Data/master_audiofeatures_track_uri.csv", rename_file="audiofeatures.csv")
df = pd.read_csv("audiofeatures.csv", sep = '|')
df.head()
df.hist(['danceability'])
df.hist(['valence'])
df.hist(['energy'])
df.hist(['tempo'])
fig, axes = plt.subplots(1, 2)
df.hist(['valence'], bins=50, ax=axes[0])
df.hist(['energy'], bins=50, ax=axes[1])
df.hist(['valence', 'danceability', 'energy'], bins=50)
df['valence'].corr(df['energy'])
df['valence'].corr(df['danceability'])
df['energy'].corr(df['loudness'])
df['danceability'].corr(df['tempo'])
df['danceability'].corr(df['tempo'])
df['tempo'].corr(df['energy'])
df['instrumentalness'].corr(df['speechiness'])
df.hist(['tempo'])
| 4_Data_Analysis/audiofeatures_analysis_valence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
import re
import seaborn as sns
from wordcloud import WordCloud
os.getcwd()
os.chdir(r"C:\Users\amit.srivastava\Desktop\mine\Python\EXAMS")
reviews = pd.read_csv("Consumer_Reviews_of_Amazon_Products_May19.csv")
reviews.head(5)
#Answer no 1
reviews.shape
#Answer no 2
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
plt.figure(figsize=(6, 6))
ax=sns.countplot(x="primaryCategories",data=reviews)
# Answer No 3
import numpy as np
import matplotlib.pyplot as plt
sns.countplot(y="primaryCategories",data=reviews, color='Green')
# +
# #!pip install pip -U --user
# #!pip install setuptools -U --user
# #!pip install -U spacy --user
# #!python -m spacy download en_core_web_sm
# +
#Answer No 4
import spacy
from spacy import displacy
import en_core_web_sm
nlp = en_core_web_sm.load()
res = []
reviews_combined = " ".join(reviews[reviews['reviews_rating']==1].reviews_text)
doc = nlp(reviews_combined[:51])
for token in doc:
res.append([token.text, token.lemma_, token.pos_, token.tag_, token.is_alpha, token.is_stop])
pd.DataFrame(res,columns=["text","lemma_","pos","tag","is_alpha","is_stop"])
# -
reviews.reviews_rating.value_counts()
#Answer No 5
text = ' '.join(reviews[reviews['reviews_rating']==1].reviews_text)
wordcloud = WordCloud().generate(text)
# Generate plot
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
#Answer No 6
text = ' '.join(reviews[reviews['reviews_rating']==5].reviews_text)
wordcloud = WordCloud().generate(text)
# Generate plot
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
#Answer No 7
text = ' '.join(reviews[reviews['reviews_rating']==3].reviews_text)
wordcloud = WordCloud().generate(text)
# Generate plot
plt.figure(figsize=[8,8])
plt.imshow(wordcloud)
plt.show()
#Answer No 8
r5=reviews[reviews['reviews_rating']==5]
r5e=r5[r5["primaryCategories"]=="Electronics"]
r5e.head()
from wordcloud import WordCloud
r5e_combined = " ".join(r5e.reviews_text.values)
len(r5e_combined)
word_cloudr5e = WordCloud().generate(r5e_combined)
word_cloudr5e
word_cloudr5e = WordCloud(width=800,height=800,
background_color='white',
max_words=150).\
generate(r5e_combined)
plt.figure(figsize=[6,6])
plt.imshow(word_cloudr5e)
plt.show()
#Answer No 9
import nltk
from nltk import word_tokenize
positive_rating=reviews[(reviews['reviews_rating']==5)]
positive_rating_Electrnics=positive_rating[positive_rating['primaryCategories']=='Electronics']
text_reviw_title=" ".join(positive_rating_Electrnics.reviews_title.values)
tokens = word_tokenize(text_reviw_title)
nltk.pos_tag(tokens)[:50]
| AMIT_SRIVASTAVA_PYTHON_CODE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 3 Fast Learning SVM
# ## Preparing datasets
import os
print "Current directory is: \"%s\"" % (os.getcwd())
# +
import urllib2 # import urllib.request as urllib2 in Python3
import requests, io, os, StringIO
import numpy as np
import tarfile, zipfile, gzip
def unzip_from_UCI(UCI_url, dest=''):
"""
Downloads and unpacks datasets from UCI in zip format
"""
response = requests.get(UCI_url)
compressed_file = io.BytesIO(response.content)
z = zipfile.ZipFile(compressed_file)
print ('Extracting in %s' % os.getcwd()+'\\'+dest)
for name in z.namelist():
if '.csv' in name:
print ('\tunzipping %s' %name)
z.extract(name, path=os.getcwd()+'\\'+dest)
def gzip_from_UCI(UCI_url, dest=''):
"""
Downloads and unpacks datasets from UCI in gzip format
"""
response = urllib2.urlopen(UCI_url)
compressed_file = io.BytesIO(response.read())
decompressed_file = gzip.GzipFile(fileobj=compressed_file)
filename = UCI_url.split('/')[-1][:-3]
with open(os.getcwd()+'\\'+filename, 'wb') as outfile:
outfile.write(decompressed_file.read())
print ('File %s decompressed' % filename)
def targzip_from_UCI(UCI_url, dest='.'):
"""
Downloads and unpacks datasets from UCI in tar.gz format
"""
response = urllib2.urlopen(UCI_url)
compressed_file = StringIO.StringIO(response.read())
tar = tarfile.open(mode="r:gz", fileobj = compressed_file)
tar.extractall(path=dest)
datasets = tar.getnames()
for dataset in datasets:
size = os.path.getsize(dest+'\\'+dataset)
print ('File %s is %i bytes' % (dataset,size))
tar.close()
def load_matrix(UCI_url):
"""
Downloads datasets from UCI in matrix form
"""
return np.loadtxt(urllib2.urlopen(UCI_url))
# -
# ###Bike Sharing Dataset Data Set
UCI_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/00275/Bike-Sharing-Dataset.zip'
unzip_from_UCI(UCI_url, dest='bikesharing')
# ###Covertype Data Set
UCI_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/covtype/covtype.data.gz'
gzip_from_UCI(UCI_url)
# ## Understanding Scikit-learn SVM implementation
from sklearn import datasets
iris = datasets.load_iris()
X_i, y_i = iris.data, iris.target
from sklearn.svm import SVC
from sklearn.cross_validation import cross_val_score
import numpy as np
h_class = SVC(kernel='rbf', C=1.0, gamma=0.7, random_state=101)
scores = cross_val_score(h_class, X_i, y_i, cv=20, scoring='accuracy')
print 'Accuracy: %0.3f' % np.mean(scores)
h_class.fit(X_i,y_i)
print h_class.support_
import numpy as np
from sklearn.datasets import load_boston
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
boston = load_boston()
shuffled = np.random.permutation(boston.target.size)
X_b = scaler.fit_transform(boston.data[shuffled,:])
y_b = boston.target[shuffled]
from sklearn.svm import SVR
from sklearn.cross_validation import cross_val_score
h_regr = SVR(kernel='rbf', C=20.0, gamma=0.001, epsilon=1.0)
scores = cross_val_score(h_regr, X_b, y_b, cv=20, scoring='mean_squared_error')
print 'Mean Squared Error: %0.3f' % abs(np.mean(scores))
# ## Pursuing non linear SVM by sub-sampling
from random import seed, randint
SAMPLE_COUNT = 5000
TEST_COUNT = 20000
seed(0) # allows repeatable results
sample = list()
test_sample = list()
for index, line in enumerate(open('covtype.data','rb')):
if index < SAMPLE_COUNT:
sample.append(line)
else:
r = randint(0, index)
if r < SAMPLE_COUNT:
sample[r] = line
else:
k = randint(0, index)
if k < TEST_COUNT:
if len(test_sample) < TEST_COUNT:
test_sample.append(line)
else:
test_sample[k] = line
import numpy as np
from sklearn.preprocessing import StandardScaler
for n,line in enumerate(sample):
sample[n] = map(float,line.strip().split(','))
y = np.array(sample)[:,-1]
scaling = StandardScaler()
X = scaling.fit_transform(np.array(sample)[:,:-1])
for n,line in enumerate(test_sample):
test_sample[n] = map(float,line.strip().split(','))
yt = np.array(test_sample)[:,-1]
Xt = scaling.transform(np.array(test_sample)[:,:-1])
from sklearn.svm import SVC
h = SVC(kernel='rbf', C=250.0, gamma=0.0025, random_state=101)
h.fit(X,y)
prediction = h.predict(Xt)
from sklearn.metrics import accuracy_score
print accuracy_score(yt, prediction)
# ## Achieving SVM at scale with SGD
# +
import csv, time, os
import numpy as np
from sklearn.linear_model import SGDRegressor
from sklearn.feature_extraction import DictVectorizer
from sklearn.preprocessing import MinMaxScaler
from scipy.sparse import csr_matrix
def explore(target_file, separator=',', fieldnames= None, binary_features=list(), numeric_features=list(), max_rows=20000):
"""
Generate from an online style stream a DictVectorizer and a MinMaxScaler.
Parameters
‐‐‐‐‐‐‐‐‐‐
target file = the file to stream from
separator = the field separator character
fieldnames = the fields' labels (can be ommitted and read from file)
binary_features = the list of qualitative features to consider
numeric_features = the list of numeric futures to consider
max_rows = the number of rows to be read from the stream (can be None)
"""
features = dict()
min_max = dict()
vectorizer = DictVectorizer(sparse=False)
scaler = MinMaxScaler()
with open(target_file, 'rb') as R:
iterator = csv.DictReader(R, fieldnames, delimiter=separator)
for n, row in enumerate(iterator):
# DATA EXPLORATION
for k,v in row.iteritems():
if k in binary_features:
if k+'_'+v not in features:
features[k+'_'+v]=0
elif k in numeric_features:
v = float(v)
if k not in features:
features[k]=0
min_max[k] = [v,v]
else:
if v < min_max[k][0]:
min_max[k][0]= v
elif v > min_max[k][1]:
min_max[k][1]= v
else:
pass # ignore the feature
if max_rows and n > max_rows:
break
vectorizer.fit([features])
A = vectorizer.transform([{f:0 if f not in min_max else min_max[f][0] for f in vectorizer.feature_names_},
{f:1 if f not in min_max else min_max[f][1] for f in vectorizer.feature_names_}])
scaler.fit(A)
return vectorizer, scaler
# -
def pull_examples(target_file, vectorizer, binary_features, numeric_features, target, min_max=None, separator=',',
fieldnames=None, sparse=True):
"""
Reads a online style stream and returns a generator of normalized feature vectors
Parameters
‐‐‐‐‐‐‐‐‐‐
target file = the file to stream from
vectorizer = a DictVectorizer object
binary_features = the list of qualitative features to consider
numeric_features = the list of numeric features to consider
target = the label of the response variable
min_max = a MinMaxScaler object, can be omitted leaving None
separator = the field separator character
fieldnames = the fields' labels (can be ommitted and read from file)
sparse = if a sparse vector is to be returned from the generator
"""
with open(target_file, 'rb') as R:
iterator = csv.DictReader(R, fieldnames, delimiter=separator)
for n, row in enumerate(iterator):
# DATA PROCESSING
stream_row = {}
response = np.array([float(row[target])])
for k,v in row.iteritems():
if k in binary_features:
stream_row[k+'_'+v]=1.0
else:
if k in numeric_features:
stream_row[k]=float(v)
if min_max:
features = min_max.transform(vectorizer.transform([stream_row]))
else:
features = vectorizer.transform([stream_row])
if sparse:
yield(csr_matrix(features), response, n)
else:
yield(features, response, n)
source = '\\bikesharing\\hour.csv'
local_path = os.getcwd()
b_vars = ['holiday','hr','mnth', 'season','weathersit','weekday','workingday','yr']
n_vars = ['hum', 'temp', 'atemp', 'windspeed']
std_row, min_max = explore(target_file=local_path+'\\'+source, binary_features=b_vars, numeric_features=n_vars)
print 'Features: '
for f,mv,mx in zip(std_row.feature_names_, min_max.data_min_, min_max.data_max_):
print '%s:[%0.2f,%0.2f] ' % (f,mv,mx)
# +
from sklearn.linear_model import SGDRegressor
SGD = SGDRegressor(loss='epsilon_insensitive', epsilon=0.001, penalty=None, random_state=1, average=True)
val_rmse = 0
val_rmsle = 0
predictions_start = 16000
def apply_log(x): return np.log(x + 1.0)
def apply_exp(x): return np.exp(x) - 1.0
for x,y,n in pull_examples(target_file=local_path+'\\'+source,
vectorizer=std_row, min_max=min_max,
binary_features=b_vars, numeric_features=n_vars, target='cnt'):
y_log = apply_log(y)
# MACHINE LEARNING
if (n+1) >= predictions_start:
# HOLDOUT AFTER N PHASE
predicted = SGD.predict(x)
val_rmse += (apply_exp(predicted) - y)**2
val_rmsle += (predicted - y_log)**2
if (n-predictions_start+1) % 250 == 0 and (n+1) > predictions_start:
print n,
print '%s holdout RMSE: %0.3f' % (time.strftime('%X'), (val_rmse / float(n-predictions_start+1))**0.5),
print 'holdout RMSLE: %0.3f' % ((val_rmsle / float(n-predictions_start+1))**0.5)
else:
# LEARNING PHASE
SGD.partial_fit(x, y_log)
print '%s FINAL holdout RMSE: %0.3f' % (time.strftime('%X'), (val_rmse / float(n-predictions_start+1))**0.5)
print '%s FINAL holdout RMSLE: %0.3f' % (time.strftime('%X'), (val_rmsle / float(n-predictions_start+1))**0.5)
# -
source = 'shuffled_covtype.data'
local_path = os.getcwd()
n_vars = ['var_'+'0'*int(j<10)+str(j) for j in range(54)]
std_row, min_max = explore(target_file=local_path+'\\'+source, binary_features=list(),
fieldnames= n_vars+['covertype'], numeric_features=n_vars, max_rows=50000)
print 'Features: '
for f,mv,mx in zip(std_row.feature_names_, min_max.data_min_, min_max.data_max_):
print '%s:[%0.2f,%0.2f] ' % (f,mv,mx)
from sklearn.linear_model import SGDClassifier
SGD = SGDClassifier(loss='hinge', penalty=None, random_state=1, average=True)
accuracy = 0
accuracy_record = list()
predictions_start = 50
sample = 5000
early_stop = 50000
for x,y,n in pull_examples(target_file=local_path+'\\'+source,
vectorizer=std_row,
min_max=min_max,
binary_features=list(), numeric_features=n_vars,
fieldnames= n_vars+['covertype'], target='covertype'):
# LEARNING PHASE
if n > predictions_start:
accuracy += int(int(SGD.predict(x))==y[0])
if n % sample == 0:
accuracy_record.append(accuracy / float(sample))
print '%s Progressive accuracy at example %i: %0.3f' % (time.strftime('%X'), n, np.mean(accuracy_record[-sample:]))
accuracy = 0
if early_stop and n >= early_stop:
break
SGD.partial_fit(x, y, classes=range(1,8))
# ## Including non-linearities in SGD
# +
from sklearn.linear_model import SGDRegressor
from sklearn.preprocessing import PolynomialFeatures
source = '\\bikesharing\\hour.csv'
local_path = os.getcwd()
b_vars = ['holiday','hr','mnth', 'season','weathersit','weekday','workingday','yr']
n_vars = ['hum', 'temp', 'atemp', 'windspeed']
std_row, min_max = explore(target_file=local_path+'\\'+source, binary_features=b_vars, numeric_features=n_vars)
poly = PolynomialFeatures(degree=2, interaction_only=False, include_bias=False)
SGD = SGDRegressor(loss='epsilon_insensitive', epsilon=0.001, penalty=None, random_state=1, average=True)
val_rmse = 0
val_rmsle = 0
predictions_start = 16000
def apply_log(x): return np.log(x + 1.0)
def apply_exp(x): return np.exp(x) - 1.0
for x,y,n in pull_examples(target_file=local_path+'\\'+source,
vectorizer=std_row, min_max=min_max, sparse = False,
binary_features=b_vars, numeric_features=n_vars, target='cnt'):
y_log = apply_log(y)
# Extract only quantitative features and expand them
num_index = [j for j, i in enumerate(std_row.feature_names_) if i in n_vars]
x_poly = poly.fit_transform(x[:,num_index])[:,len(num_index):]
new_x = np.concatenate((x, x_poly), axis=1)
# MACHINE LEARNING
if (n+1) >= predictions_start:
# HOLDOUT AFTER N PHASE
predicted = SGD.predict(new_x)
val_rmse += (apply_exp(predicted) - y)**2
val_rmsle += (predicted - y_log)**2
if (n-predictions_start+1) % 250 == 0 and (n+1) > predictions_start:
print n,
print '%s holdout RMSE: %0.3f' % (time.strftime('%X'), (val_rmse / float(n-predictions_start+1))**0.5),
print 'holdout RMSLE: %0.3f' % ((val_rmsle / float(n-predictions_start+1))**0.5)
else:
# LEARNING PHASE
SGD.partial_fit(new_x, y_log)
print '%s FINAL holdout RMSE: %0.3f' % (time.strftime('%X'), (val_rmse / float(n-predictions_start+1))**0.5)
print '%s FINAL holdout RMSLE: %0.3f' % (time.strftime('%X'), (val_rmsle / float(n-predictions_start+1))**0.5)
# -
# ## Trying explicit high dimensional mappings
# +
source = 'shuffled_covtype.data'
local_path = os.getcwd()
n_vars = ['var_'+str(j) for j in range(54)]
std_row, min_max = explore(target_file=local_path+'\\'+source, binary_features=list(),
fieldnames= n_vars+['covertype'], numeric_features=n_vars, max_rows=50000)
from sklearn.linear_model import SGDClassifier
from sklearn.kernel_approximation import RBFSampler
SGD = SGDClassifier(loss='hinge', penalty=None, random_state=1, average=True)
rbf_feature = RBFSampler(gamma=0.5, n_components=300, random_state=0)
accuracy = 0
accuracy_record = list()
predictions_start = 50
sample = 5000
early_stop = 50000
for x,y,n in pull_examples(target_file=local_path+'\\'+source,
vectorizer=std_row,
min_max=min_max,
binary_features=list(),
numeric_features=n_vars,
fieldnames= n_vars+['covertype'], target='covertype', sparse=False):
rbf_x = rbf_feature.fit_transform(x)
# LEARNING PHASE
if n > predictions_start:
accuracy += int(int(SGD.predict(rbf_x))==y[0])
if n % sample == 0:
accuracy_record.append(accuracy / float(sample))
print '%s Progressive accuracy at example %i: %0.3f' % (time.strftime('%X'), \
n, np.mean(accuracy_record[-sample:]))
accuracy = 0
if early_stop and n >= early_stop:
break
SGD.partial_fit(rbf_x, y, classes=range(1,8))
# -
# ## Hyperparameters tuning
# +
from sklearn.linear_model import SGDRegressor
from sklearn.grid_search import ParameterSampler
source = '\\bikesharing\\hour.csv'
local_path = os.getcwd()
b_vars = ['holiday','hr','mnth', 'season','weathersit','weekday','workingday','yr']
n_vars = ['hum', 'temp', 'atemp', 'windspeed']
std_row, min_max = explore(target_file=local_path+'\\'+source, binary_features=b_vars, numeric_features=n_vars)
val_rmse = 0
val_rmsle = 0
predictions_start = 16000
tmp_rsmle = 10**6
def apply_log(x): return np.log(x + 1.0)
def apply_exp(x): return np.exp(x) - 1.0
param_grid = {'penalty':['l1', 'l2'], 'alpha': 10.0**-np.arange(2,5)}
random_tests = 3
search_schedule = list(ParameterSampler(param_grid, n_iter=random_tests, random_state=5))
results = dict()
for search in search_schedule:
SGD = SGDRegressor(loss='epsilon_insensitive', epsilon=0.001, penalty=None, random_state=1, average=True)
params =SGD.get_params()
new_params = {p:params[p] if p not in search else search[p] for p in params}
SGD.set_params(**new_params)
print str(search)[1:-1]
for iterations in range(200):
for x,y,n in pull_examples(target_file=local_path+'\\'+source,
vectorizer=std_row, min_max=min_max, sparse = False,
binary_features=b_vars, numeric_features=n_vars, target='cnt'):
y_log = apply_log(y)
# MACHINE LEARNING
if (n+1) >= predictions_start:
# HOLDOUT AFTER N PHASE
predicted = SGD.predict(x)
val_rmse += (apply_exp(predicted) - y)**2
val_rmsle += (predicted - y_log)**2
else:
# LEARNING PHASE
SGD.partial_fit(x, y_log)
examples = float(n-predictions_start+1) * (iterations+1)
print_rmse = (val_rmse / examples)**0.5
print_rmsle = (val_rmsle / examples)**0.5
if iterations == 0:
print 'Iteration %i - RMSE: %0.3f - RMSE: %0.3f' % (iterations+1, print_rmse, print_rmsle)
if iterations > 0:
if tmp_rmsle / print_rmsle <= 1.01:
print 'Iteration %i - RMSE: %0.3f - RMSE: %0.3f\n' % (iterations+1, print_rmse, print_rmsle)
results[str(search)]= {'rmse':float(print_rmse), 'rmsle':float(print_rmsle)}
break
tmp_rmsle = print_rmsle
# -
# ## Other alternatives for SVM fast learning
# ###Useful dataset examples
# +
with open('house_dataset','wb') as W:
W.write("0 | price:.23 sqft:.25 age:.05 2006\n")
W.write("1 2 'second_house | price:.18 sqft:.15 age:.35 1976\n")
W.write("0 1 0.5 'third_house | price:.53 sqft:.32 age:.87 1924\n")
with open('house_dataset','rb') as R:
for line in R:
print line.strip()
# -
# ###A way to call VW from Python
# +
import subprocess
def execute_vw(parameters):
execution = subprocess.Popen('vw '+parameters, \
shell=True, stderr=subprocess.PIPE)
line = ""
history = ""
while True:
out = execution.stderr.read(1)
history += out
if out == '' and execution.poll() != None:
print '------------ COMPLETED ------------\n'
break
if out != '':
line += out
if '\n' in line[-2:]:
print line[:-2]
line = ''
return history.split('\r\n')
# -
params = "house_dataset"
results = execute_vw(params)
# ###Processing examples
# +
import csv
def vw_convert(origin_file, target_file, binary_features, numeric_features, target, transform_target=lambda(x):x,
separator=',', classification=True, multiclass=False, fieldnames= None, header=True, sparse=True):
"""
Reads a online style stream and returns a generator of normalized feature vectors
Parameters
‐‐‐‐‐‐‐‐‐‐
original_file = the csv file you are taken the data from
target file = the file to stream from
binary_features = the list of qualitative features to consider
numeric_features = the list of numeric features to consider
target = the label of the response variable
transform_target = a function transforming the response
separator = the field separator character
classification = a Boolean indicating if it is classification
multiclass = a Boolean for multiclass classification
fieldnames = the fields' labels (can be ommitted and read from file)
header = a boolean indicating if the original file has an header
sparse = if a sparse vector is to be returned from the generator
"""
with open(target_file, 'wb') as W:
with open(origin_file, 'rb') as R:
iterator = csv.DictReader(R, fieldnames, delimiter=separator)
for n, row in enumerate(iterator):
if not header or n>0:
# DATA PROCESSING
response = transform_target(float(row[target]))
if classification and not multiclass:
if response == 0:
stream_row = '-1 '
else:
stream_row = '1 '
else:
stream_row = str(response)+' '
quantitative = list()
qualitative = list()
for k,v in row.iteritems():
if k in binary_features:
qualitative.append(str(k)+'_'+str(v)+':1')
else:
if k in numeric_features and (float(v)!=0 or not sparse):
quantitative.append(str(k)+':'+str(v))
if quantitative:
stream_row += '|n '+' '.join(quantitative)
if qualitative:
stream_row += '|q ' + ' '.join(qualitative)
W.write(stream_row+'\n')
# -
# ###Examples with toys datasets
import numpy as np
from sklearn.datasets import load_iris, load_boston
from random import seed
iris = load_iris()
seed(2)
re_order = np.random.permutation(len(iris.target))
with open('iris_versicolor.vw','wb') as W1:
for k in re_order:
y = iris.target[k]
X = iris.values()[1][k,:]
features = ' |f '+' '.join([a+':'+str(b) for a,b in zip(map(lambda(a): a[:-5].replace(' ','_'), iris.feature_names),X)])
target = '1' if y==1 else '-1'
W1.write(target+features+'\n')
boston = load_boston()
seed(2)
re_order = np.random.permutation(len(boston.target))
with open('boston.vw','wb') as W1:
for k in re_order:
y = boston.target[k]
X = boston.data[k,:]
features = ' |f '+' '.join([a+':'+str(b) for a,b in zip(map(lambda(a): a[:-5].replace(' ','_'), iris.feature_names),X)])
W1.write(str(y)+features+'\n')
# ###Binary Iris
# +
params = '--ksvm --l2 0.000001 --reprocess 2 -b 18 --kernel rbf --bandwidth=0.1 -p iris_bin.test -d iris_versicolor.vw'
results = execute_vw(params)
accuracy = 0
with open('iris_bin.test', 'rb') as R:
with open('iris_versicolor.vw', 'rb') as TRAIN:
holdouts = 0.0
for n,(line, example) in enumerate(zip(R,TRAIN)):
if (n+1) % 10==0:
predicted = float(line.strip())
y = float(example.split('|')[0])
accuracy += np.sign(predicted)==np.sign(y)
holdouts += 1
print 'holdout accuracy: %0.3f' % ((accuracy / holdouts)**0.5)
# -
# ###Boston dataset
params = 'boston.vw -f boston.model --loss_function squared -k --cache_file cache_train.vw --passes=20 --nn 5 --dropout'
results = execute_vw(params)
params = '-t boston.vw -i boston.model -k --cache_file cache_test.vw -p boston.test'
results = execute_vw(params)
val_rmse = 0
with open('boston.test', 'rb') as R:
with open('boston.vw', 'rb') as TRAIN:
holdouts = 0.0
for n,(line, example) in enumerate(zip(R,TRAIN)):
if (n+1) % 10==0:
predicted = float(line.strip())
y = float(example.split('|')[0])
val_rmse += (predicted - y)**2
holdouts += 1
print 'holdout RMSE: %0.3f' % ((val_rmse / holdouts)**0.5)
# ## Faster bikesharing
# +
import os
import numpy as np
def apply_log(x):
return np.log(x + 1.0)
def apply_exp(x):
return np.exp(x) - 1.0
local_path = os.getcwd()
b_vars = ['holiday','hr','mnth', 'season','weathersit','weekday','workingday','yr']
n_vars = ['hum', 'temp', 'atemp', 'windspeed']
source = '\\bikesharing\\hour.csv'
origin = target_file=local_path+'\\'+source
target = target_file=local_path+'\\'+'bike.vw'
vw_convert(origin, target, binary_features=b_vars, numeric_features=n_vars, target = 'cnt', transform_target=apply_log,
separator=',', classification=False, multiclass=False, fieldnames= None, header=True)
# -
params = 'bike.vw -f regression.model -k --cache_file cache_train.vw --passes=100 --hash strings --holdout_after 16000'
results = execute_vw(params)
# +
params = '-t bike.vw -i regression.model -k --cache_file cache_test.vw -p pred.test'
results = execute_vw(params)
val_rmse = 0
val_rmsle = 0
with open('pred.test', 'rb') as R:
with open('bike.vw', 'rb') as TRAIN:
holdouts = 0.0
for n,(line, example) in enumerate(zip(R,TRAIN)):
if n > 16000:
predicted = float(line.strip())
y_log = float(example.split('|')[0])
y = apply_exp(y_log)
val_rmse += (apply_exp(predicted) - y)**2
val_rmsle += (predicted - y_log)**2
holdouts += 1
print 'holdout RMSE: %0.3f' % ((val_rmse / holdouts)**0.5)
print 'holdout RMSLE: %0.3f' % ((val_rmsle / holdouts)**0.5)
# -
# ## Covertype dataset crunched by VW
import os
local_path = os.getcwd()
n_vars = ['var_'+'0'*int(j<10)+str(j) for j in range(54)]
source = 'shuffled_covtype.data'
origin = target_file=local_path+'\\'+source
target = target_file=local_path+'\\'+'covtype.vw'
vw_convert(origin, target, binary_features=list(), fieldnames= n_vars+['covertype'], numeric_features=n_vars,
target = 'covertype', separator=',', classification=True, multiclass=True, header=False, sparse=False)
params = 'covtype.vw --ect 7 -f multiclass.model -k --cache_file cache_train.vw --passes=2 -l 1.0 --cubic nnn'
results = execute_vw(params)
params = '-t covtype.vw -i multiclass.model -k --cache_file cache_test.vw -p covertype.test'
results = execute_vw(params)
accuracy = 0
with open('covertype.test', 'rb') as R:
with open('covtype.vw', 'rb') as TRAIN:
holdouts = 0.0
for n,(line, example) in enumerate(zip(R,TRAIN)):
if (n+1) % 10==0:
predicted = float(line.strip())
y = float(example.split('|')[0])
accuracy += predicted ==y
holdouts += 1
print 'holdout accuracy: %0.3f' % (accuracy / holdouts)
| Module 3/Chapter 3/Chapter_3_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# ## Gated PixelCNN
# PixelCNN is an autoregressive likelihood model for the task of image modeling, and Gated PixelCNN is an improved version, which uses GatedMaskedConv.
#
# ### Reference
# - https://keras.io/examples/generative/pixelcnn/
# - Pixel Recurrent Neural Networks: https://arxiv.org/abs/1601.06759
# - Conditional Image Generation with PixelCNN Decoders: https://arxiv.org/abs/1606.05328
# - https://uvadlc-notebooks.readthedocs.io/en/latest/tutorial_notebooks/tutorial12/Autoregressive_Image_Modeling.html
# +
import os
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
from torchvision import datasets, transforms
from sklearn.manifold import TSNE
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
class MaskedConv2d(nn.Conv2d):
"""
Implements a conv2d with mask applied on its weights.
Args:
mask (torch.Tensor): the mask tensor.
in_channels (int) – Number of channels in the input image.
out_channels (int) – Number of channels produced by the convolution.
kernel_size (int or tuple) – Size of the convolving kernel
"""
def __init__(self, mask, in_channels, out_channels, kernel_size, **kwargs):
super().__init__(in_channels, out_channels, kernel_size, **kwargs)
self.register_buffer('mask', mask[None, None])
def forward(self, x):
self.weight.data *= self.mask # mask weights
return super().forward(x)
class VerticalStackConv(MaskedConv2d):
def __init__(self, mask_type, in_channels, out_channels, kernel_size, **kwargs):
# Mask out all pixels below. For efficiency, we could also reduce the kernel
# size in height (k//2, k), but for simplicity, we stick with masking here.
self.mask_type = mask_type
if isinstance(kernel_size, int):
kernel_size = (kernel_size, kernel_size)
mask = torch.zeros(kernel_size)
mask[:kernel_size[0]//2, :] = 1.0
if self.mask_type == "B":
mask[kernel_size[0]//2, :] = 1.0
super().__init__(mask, in_channels, out_channels, kernel_size, **kwargs)
class HorizontalStackConv(MaskedConv2d):
def __init__(self, mask_type, in_channels, out_channels, kernel_size, **kwargs):
# Mask out all pixels on the left. Note that our kernel has a size of 1
# in height because we only look at the pixel in the same row.
self.mask_type = mask_type
if isinstance(kernel_size, int):
kernel_size = (1, kernel_size)
assert kernel_size[0] == 1
if "padding" in kwargs:
if isinstance(kwargs["padding"], int):
kwargs["padding"] = (0, kwargs["padding"])
mask = torch.zeros(kernel_size)
mask[:, :kernel_size[1]//2] = 1.0
if self.mask_type == "B":
mask[:, kernel_size[1]//2] = 1.0
super().__init__(mask, in_channels, out_channels, kernel_size, **kwargs)
# +
# visualize the receptive field for VerticalStackConv and HorizontalStackConv
# we can compute the gradients of the input to imshow the receptive field
inp_img = torch.zeros(1, 1, 11, 11)
inp_img.requires_grad_()
def show_center_recep_field(img, out):
"""
Calculates the gradients of the input with respect to the output center pixel,
and visualizes the overall receptive field.
Inputs:
img - Input image for which we want to calculate the receptive field on.
out - Output features/loss which is used for backpropagation, and should be
the output of the network/computation graph.
"""
# Determine gradients, the center pixel
loss = out[0, :, img.shape[2]//2, img.shape[3]//2].sum() # L1 loss for simplicity
# Retain graph as we want to stack multiple layers and show the receptive field of all of them
loss.backward(retain_graph=True)
img_grads = img.grad.abs()
img.grad.fill_(0) # Reset grads
# Plot receptive field
img = img_grads.squeeze().cpu().numpy()
fig, ax = plt.subplots(1, 2)
pos = ax[0].imshow(img)
ax[1].imshow(img > 0)
# Mark the center pixel in red if it doesn't have any gradients,
# which is the case for standard autoregressive models)
show_center = (img[img.shape[0]//2, img.shape[1]//2] == 0)
if show_center:
center_pixel = np.zeros(img.shape + (4,))
center_pixel[center_pixel.shape[0]//2, center_pixel.shape[1]//2, :] = np.array([1.0, 0.0, 0.0, 1.0])
for i in range(2):
ax[i].axis('off')
if show_center:
ax[i].imshow(center_pixel)
ax[0].set_title("Weighted receptive field")
ax[1].set_title("Binary receptive field")
plt.show()
plt.close()
# we don't use conv, so the receptive field is only the center pixel
show_center_recep_field(inp_img, inp_img)
# +
# we first visualize the original masked_conv
kernel_size = 3
mask_A = torch.zeros((3, 3))
mask_A[:kernel_size//2, :] = 1.0
mask_A[kernel_size//2, :kernel_size//2] = 1.0
masked_conv = MaskedConv2d(mask_A, 1, 1, 3, padding=1)
masked_conv.weight.data.fill_(1)
masked_conv.bias.data.fill_(0)
masked_conv_img = masked_conv(inp_img)
show_center_recep_field(inp_img, masked_conv_img)
# +
# use mask_type B
mask_B = mask_A.clone()
mask_B[kernel_size//2, kernel_size//2] = 1.
masked_conv = MaskedConv2d(mask_B, 1, 1, 3, padding=1)
masked_conv.weight.data.fill_(1)
masked_conv.bias.data.fill_(0)
for l_idx in range(4):
masked_conv_img = masked_conv(masked_conv_img)
print(f"Layer {l_idx+2}")
show_center_recep_field(inp_img, masked_conv_img)
# there is a “blind spot” on the right upper side
# -
# visualize HorizontalStackConv
horiz_conv = HorizontalStackConv("A", 1, 1, 3, padding=1)
horiz_conv.weight.data.fill_(1)
horiz_conv.bias.data.fill_(0)
horiz_img = horiz_conv(inp_img)
show_center_recep_field(inp_img, horiz_img)
# visualize VerticalStackConv
vert_conv = VerticalStackConv("A", 1, 1, 3, padding=1)
vert_conv.weight.data.fill_(1)
vert_conv.bias.data.fill_(0)
vert_img = vert_conv(inp_img)
show_center_recep_field(inp_img, vert_img)
# combine the two by adding, which is what we expect
horiz_img = vert_img + horiz_img
show_center_recep_field(inp_img, horiz_img)
# +
# Initialize convolutions with equal weight to all input pixels
horiz_conv = HorizontalStackConv("B", 1, 1, 3, padding=1)
horiz_conv.weight.data.fill_(1)
horiz_conv.bias.data.fill_(0)
vert_conv = VerticalStackConv("B", 1, 1, 3, padding=1)
vert_conv.weight.data.fill_(1)
vert_conv.bias.data.fill_(0)
# note we use mask_type A for the first layer, but after first layer we should use mask_type B
# We reuse our convolutions for the 4 layers here. Note that in a standard network,
# we don't do that, and instead learn 4 separate convolution. As this cell is only for
# visualization purposes, we reuse the convolutions for all layers.
for l_idx in range(4):
vert_img = vert_conv(vert_img)
horiz_img = horiz_conv(horiz_img) + vert_img
print(f"Layer {l_idx+2}")
show_center_recep_field(inp_img, horiz_img)
# -
# check the vert_conv
show_center_recep_field(inp_img, vert_img)
class GatedMaskedConv(nn.Module):
def __init__(self, in_channels, kernel_size=3, dilation=1):
"""
Gated Convolution block implemented the computation graph shown above.
"""
super().__init__()
padding = dilation * (kernel_size - 1) // 2
self.conv_vert = VerticalStackConv("B", in_channels, 2*in_channels, kernel_size, padding=padding,
dilation=dilation)
self.conv_horiz = HorizontalStackConv("B", in_channels, 2*in_channels, kernel_size, padding=padding,
dilation=dilation)
self.conv_vert_to_horiz = nn.Conv2d(2*in_channels, 2*in_channels, kernel_size=1)
self.conv_horiz_1x1 = nn.Conv2d(in_channels, in_channels, kernel_size=1)
def forward(self, v_stack, h_stack):
# Vertical stack (left)
v_stack_feat = self.conv_vert(v_stack)
v_val, v_gate = v_stack_feat.chunk(2, dim=1)
v_stack_out = torch.tanh(v_val) * torch.sigmoid(v_gate)
# Horizontal stack (right)
h_stack_feat = self.conv_horiz(h_stack)
h_stack_feat = h_stack_feat + self.conv_vert_to_horiz(v_stack_feat)
h_val, h_gate = h_stack_feat.chunk(2, dim=1)
h_stack_feat = torch.tanh(h_val) * torch.sigmoid(h_gate)
h_stack_out = self.conv_horiz_1x1(h_stack_feat)
h_stack_out = h_stack_out + h_stack
return v_stack_out, h_stack_out
# +
# GatedPixelCNN
class GatedPixelCNN(nn.Module):
def __init__(self, in_channels, channels, out_channels):
super().__init__()
# Initial first conv with mask_type A
self.conv_vstack = VerticalStackConv("A", in_channels, channels, 3, padding=1)
self.conv_hstack = HorizontalStackConv("A", in_channels, channels, 3, padding=1)
# Convolution block of PixelCNN. use dilation instead of
# downscaling used in the encoder-decoder architecture in PixelCNN++
self.conv_layers = nn.ModuleList([
GatedMaskedConv(channels),
GatedMaskedConv(channels, dilation=2),
GatedMaskedConv(channels),
GatedMaskedConv(channels, dilation=4),
GatedMaskedConv(channels),
GatedMaskedConv(channels, dilation=2),
GatedMaskedConv(channels)
])
# Output classification convolution (1x1)
self.conv_out = nn.Conv2d(channels, out_channels, kernel_size=1)
def forward(self, x):
# first convolutions
v_stack = self.conv_vstack(x)
h_stack = self.conv_hstack(x)
# Gated Convolutions
for layer in self.conv_layers:
v_stack, h_stack = layer(v_stack, h_stack)
# 1x1 classification convolution
# Apply ELU before 1x1 convolution for non-linearity on residual connection
out = self.conv_out(F.elu(h_stack))
return out
# -
# visualize GaGatedPixelCNN
test_model = GatedPixelCNN(1, 64, 1)
inp = torch.zeros(1, 1, 28, 28)
inp.requires_grad_()
out = test_model(inp)
show_center_recep_field(inp, out.squeeze(dim=2))
del inp, out, test_model
# +
image_size = 28
in_channels = 1
out_channels = 1
channels = 128 # hidden channels
batch_size = 64
epochs = 10
transform=transforms.Compose([
transforms.ToTensor()
])
dataset1 = datasets.MNIST('/data', train=True, download=True,
transform=transform)
dataset2 = datasets.MNIST('/data', train=False,
transform=transform)
train_loader = torch.utils.data.DataLoader(dataset1, batch_size=batch_size, shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset2, batch_size=batch_size)
model = GatedPixelCNN(in_channels, channels, out_channels).cuda()
optimizer = torch.optim.Adam(model.parameters(), lr=5e-4)
# -
print_freq = 1000
for epoch in range(epochs):
print("Start training epoch {}".format(epoch,))
for i, (images, labels) in enumerate(train_loader):
images = (images > 0.33).float() # convert to 0, 1
images = images.cuda()
logits = model(images)
loss = F.binary_cross_entropy_with_logits(logits, images)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if (i + 1) % print_freq == 0 or (i + 1) == len(train_loader):
print("\t [{}/{}]: loss {}".format(i, len(train_loader), loss.item()))
# +
## generate new images by PixelCNN
n_cols, n_rows = 8, 8
C = 1
H = 28
W = 28
# Create an empty array of pixels.
pixels = torch.zeros(n_cols * n_rows, C, H, W).cuda()
model.eval()
with torch.no_grad():
# Iterate over the pixels because generation has to be done sequentially pixel by pixel.
for h in range(H):
for w in range(W):
for c in range(C):
# Feed the whole array and retrieving the pixel value probabilities for the next pixel.
logits = model(pixels)[:, c, h, w]
probs = logits.sigmoid()
# Use the probabilities to pick pixel values and append the values to the image frame.
pixels[:, c, h, w] = torch.bernoulli(probs)
generated_imgs = pixels.cpu().numpy()
generated_imgs = np.array(generated_imgs * 255, dtype=np.uint8).reshape(n_rows, n_cols, H, W)
fig = plt.figure(figsize=(8, 8), constrained_layout=True)
gs = fig.add_gridspec(n_rows, n_cols)
for n_col in range(n_cols):
for n_row in range(n_rows):
f_ax = fig.add_subplot(gs[n_row, n_col])
f_ax.imshow(generated_imgs[n_row, n_col], cmap="gray")
f_ax.axis("off")
# +
# we can use pixelcnn to autocomplete images
test_loader_iter = iter(test_loader)
images, labels = next(test_loader_iter)
n_samples = 8
n_rows = 12 # the number of rows to keep
images = (images[:n_samples] > 0.33).float() # convert to 0, 1
sub_images = images.clone()
sub_images[:, :, n_rows:, :] = 0 # mask some pixels
pixels = sub_images.cuda()
with torch.no_grad():
# Iterate over the pixels because generation has to be done sequentially pixel by pixel.
for h in range(n_rows, H):
for w in range(W):
for c in range(C):
# Feed the whole array and retrieving the pixel value probabilities for the next pixel.
logits = model(pixels)[:, c, h, w]
probs = logits.sigmoid()
# Use the probabilities to pick pixel values and append the values to the image frame.
pixels[:, c, h, w] = torch.bernoulli(probs)
original_imgs = images.cpu().numpy()
original_imgs = np.array(original_imgs * 255, dtype=np.uint8).reshape(n_samples, H, W)
sub_imgs = sub_images.cpu().numpy()
sub_imgs = np.array(sub_imgs * 255, dtype=np.uint8).reshape(n_samples, H, W)
generated_imgs = pixels.cpu().numpy()
generated_imgs = np.array(generated_imgs * 255, dtype=np.uint8).reshape(n_samples, H, W)
fig = plt.figure(figsize=(3, 8), constrained_layout=True)
gs = fig.add_gridspec(n_samples, 3)
for n_row in range(n_samples):
f_ax = fig.add_subplot(gs[n_row, 0])
f_ax.imshow(original_imgs[n_row], cmap="gray")
f_ax.axis("off")
f_ax = fig.add_subplot(gs[n_row, 1])
f_ax.imshow(sub_imgs[n_row], cmap="gray")
f_ax.axis("off")
f_ax = fig.add_subplot(gs[n_row, 2])
f_ax.imshow(generated_imgs[n_row], cmap="gray")
f_ax.axis("off")
| models/gated_pixelcnn.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.0
# language: julia
# name: julia-1.6
# ---
# +
using Pkg
if isfile("../Project.toml") && isfile("../Manifest.toml")
Pkg.activate("..");
end
using Random
using Distributions
using Plots
using prml
Random.seed!(1234);
gr();
# +
function create_toy_data(func, sample_size, std)
x = collect(range(0, stop=1.0, length=sample_size));
noise = rand(Normal(0.0, std), sample_size);
return x, func(x) + noise
end
function sinusoidal(x)
return sin.(2 * pi * x)
end
# -
# add some Gaussian noise to the data points generated from sinusoidal function
#
# $$
# y \sim \sin(2 \pi x) + \mathcal{N}(0, \sigma^2)
# $$
# +
x_train, y_train = create_toy_data(sinusoidal, 10, 0.25);
x_test = collect(range(0, stop=1.0, length=100));
y_test = sinusoidal(x_test);
plot(x_train, y_train, seriestype=:scatter, label="training data", legend=:topright)
plot!(x_test, y_test, label="sin(2π x)")
# -
# <img src="images/ch1/image1.png">
# We fit the data points with the linear cohesion of polynomial functions
#
# $$
# t_n \sim \boldsymbol{\phi}(x_n) \boldsymbol{w}.
# $$
#
# The answer is given by
#
# $$
# \boldsymbol{w} = (\boldsymbol{\Phi}^{T} \boldsymbol{\Phi})^{-1} \boldsymbol{\Phi}^{T} \boldsymbol{t}
# $$
#
# where $\boldsymbol{\Phi}$ is
#
# $$
# \boldsymbol{\Phi} = \begin{bmatrix} \boldsymbol{\phi}^{T} (\boldsymbol{x_1}) \\ \boldsymbol{\phi}^{T} (\boldsymbol{x_2}) \\ \vdots \\ \boldsymbol{\phi}^{T}(\boldsymbol{x_N}) \end{bmatrix}
# $$
feature = PolynomialFeature(9);
X_train = transform(feature, x_train);
X_test = transform(feature, x_test);
#x = collect(reshape(range(0, stop=1.0, length=10), 10));
model = LinearRegressor([0], 0);
#Phi = collect(reshape(transpose(X_train), size(X_train)[2], size(X_train)[1]));
fitting(model, X_train, y_train);
#tmp = collect(reshape(transpose(X_test), size(X_test)[2], size(X_test)[1]));
y, y_std = predict(model, X_test, true);
# +
l = @layout [a b; c d]
plots = []
for i in [0, 1, 3, 9]
feature = PolynomialFeature(i);
X_train = transform(feature, x_train);
X_test = transform(feature, x_test);
#x = range(0, stop=1.0, length=10);
model = LinearRegressor([0], 0);
fitting(model, X_train, y_train);
y = predict(model, X_test, false);
p = plot(x_test, y, label="predicted", legend=:topright);
p = plot!(x_train, y_train, label="train", seriestype=:scatter);
p = plot!(x_test, y_test, label="sin");
push!(plots, p);
end
plot(plots[1], plots[2], plots[3], plots[4], layout=l)
# -
# <img src="images/ch1/image2.png">
# +
using LinearAlgebra
function rmse(a::Array{Float64, 1}, b::Array{Float64, 1})
return sum((b - a).^2) / size(a)[1]
end
training_errors = [];
test_errors = [];
for i in 0:10
feature = PolynomialFeature(i);
X_train = transform(feature, x_train);
X_test = transform(feature, x_test);
model = LinearRegressor([0], 0);
fitting(model, X_train, y_train);
y_trained = predict(model, X_train, false);
push!(training_errors, rmse(predict(model, X_train, false), y_train));
push!(test_errors, rmse(predict(model, X_test, false), y_test + rand(Normal(0.0, 0.25), size(y_test)[1])));
end
plot(training_errors, xlabel="degree", ylabel="RMSE", lw=2, linecolor=:red, label="Training")
plot!(test_errors, lw=2, linecolor=:blue, label="Test")
plot!(training_errors, seriestype=:scatter, markercolor=:red, label=nothing)
plot!(test_errors, seriestype=:scatter, markercolor=:blue, label=nothing)
# -
# <img src="images/ch1/image3.png">
| notebook/ch01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (myenv)
# language: python
# name: myenv
# ---
# Import SQLAlchemy and other dependencies here
import sqlalchemy
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, inspect, func
from sqlalchemy import Column, Float, Integer, String, Date
from sqlalchemy.ext.declarative import declarative_base
Base2 = declarative_base()
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sqlalchemy import create_engine
db_user = "Progress"
db_password = "<PASSWORD>"
connection_string = (f'postgresql://{db_user}:{db_password}@localhost:5432/Query_SQL')
engine = create_engine(connection_string)
connection = engine.connect()
connection
# +
# Create DataFrames from each table in Database
df_titles = pd.read_sql_table("titles",connection)
df_departments = pd.read_sql_table("departments",connection)
df_employees = pd.read_sql_table("employees",connection)
df_salaries = pd.read_sql_table("salaries",connection)
df_dept_emp = pd.read_sql_table("dept_emp",connection)
df_dept_manager = pd.read_sql_table("dept_manager",connection)
# -
# Create the inspector and connect it to the engine
inspector = inspect(engine)
# Collect the names of tables within the database
inspector.get_table_names()
# Using the inspector to print the column names within the 'employees' table and its types
columns = inspector.get_columns('employees')
for column in columns:
print(column["name"], column["type"])
#create employees class
class employees(Base2):
__tablename__ = 'employees'
emp_no = Column(Integer, primary_key=True)
birth_date = Column(Date)
first_name = Column(String)
last_name = Column(String)
sex = Column(String)
hire_date = Column(Date)
session = Session(engine)
# +
# print sums by gender
male = session.query(employees).filter_by(sex = 'M').count()
female = session.query(employees).filter_by(sex = 'F').count()
print(male)
print(female)
# -
# Using the inspector to print the column names within the 'Salaries' table and its types
columns = inspector.get_columns('salaries')
for column in columns:
print(column["name"], column["type"])
#create salaries class
class salaries(Base2):
__tablename__ = 'salaries'
emp_no = Column(Integer, primary_key=True)
salary = Column(Integer)
#query the salaries table
x = session.query(salaries.salary)
# Plot the Results in a Matplotlib bar chart
df = pd.DataFrame(x, columns=['salary'])
a = np.array(df)
x = a[0:,0]
# +
fig, ax = plt.subplots(figsize=(12, 8))
# the histogram of the data
n, bins, patches = plt.hist(x, 12, facecolor='purple', alpha=0.75)
plt.xlabel('Salary')
plt.ylabel('Frequency')
plt.title('Histogram of Employee Salaries')
plt.grid(True)
plt.show()
fig.savefig('Employee_Salary_Histogram.png')
# -
# Using the inspector to print the column names within the 'titles' table and its types
columns = inspector.get_columns('titles')
for column in columns:
print(column["name"], column["type"])
#create titles class
class titles(Base2):
__tablename__ = 'titles'
emp_no = Column(Integer, primary_key=True)
title = Column(String)
from_date = Column(Date)
to_date = Column(Date)
# +
# Create a DataFrame to store salary by title data from a query
query = "select title,avg(salary) as salary from salaries join employees on salaries.emp_no=employees.emp_no join titles on titles.title_id=employees.emp_title_id group by title order by salary desc;"
df_salary_title = pd.read_sql_query(query,connection)
# Create Bar Chart of Average Salary by Title
fig, ax = plt.subplots(figsize=(12, 8))
plt.xticks(rotation=90)
plt.ylabel('Avg. Salary ($)')
plt.title('Avg. Salary by Title')
plt.xticks(rotation=45)
plt.show()
fig.savefig('Avg_Salary_Title.png')
# -
| EmployeeSQL/.ipynb_checkpoints/Analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import time
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib
from random import randint
# -
device = "cpu" # please change it to "gpu" if the model needs to be run on cuda.
# +
protoFile = "pose/coco/pose_deploy_linevec.prototxt"
weightsFile = "pose/coco/pose_iter_440000.caffemodel"
nPoints = 18
# COCO Output Format
keypointsMapping = ['Nose', 'Neck', 'R-Sho', 'R-Elb', 'R-Wr', 'L-Sho',
'L-Elb', 'L-Wr', 'R-Hip', 'R-Knee', 'R-Ank', 'L-Hip',
'L-Knee', 'L-Ank', 'R-Eye', 'L-Eye', 'R-Ear', 'L-Ear']
POSE_PAIRS = [[1,2], [1,5], [2,3], [3,4], [5,6], [6,7],
[1,8], [8,9], [9,10], [1,11], [11,12], [12,13],
[1,0], [0,14], [14,16], [0,15], [15,17],
[2,17], [5,16] ]
# index of pafs correspoding to the POSE_PAIRS
# e.g for POSE_PAIR(1,2), the PAFs are located at indices (31,32) of output, Similarly, (1,5) -> (39,40) and so on.
mapIdx = [[31,32], [39,40], [33,34], [35,36], [41,42], [43,44],
[19,20], [21,22], [23,24], [25,26], [27,28], [29,30],
[47,48], [49,50], [53,54], [51,52], [55,56],
[37,38], [45,46]]
colors = [ [0,100,255], [0,100,255], [0,255,255], [0,100,255], [0,255,255], [0,100,255],
[0,255,0], [255,200,100], [255,0,255], [0,255,0], [255,200,100], [255,0,255],
[0,0,255], [255,0,0], [200,200,0], [255,0,0], [200,200,0], [0,0,0]]
# -
# Find the Keypoints using Non Maximum Suppression on the Confidence Map
def getKeypoints(probMap, threshold=0.1):
mapSmooth = cv2.GaussianBlur(probMap,(3,3),0,0)
mapMask = np.uint8(mapSmooth>threshold)
keypoints = []
#find the blobs
contours, _ = cv2.findContours(mapMask, cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
#for each blob find the maxima
for cnt in contours:
blobMask = np.zeros(mapMask.shape)
blobMask = cv2.fillConvexPoly(blobMask, cnt, 1)
maskedProbMap = mapSmooth * blobMask
_, maxVal, _, maxLoc = cv2.minMaxLoc(maskedProbMap)
keypoints.append(maxLoc + (probMap[maxLoc[1], maxLoc[0]],))
return keypoints
# ## Use the following equation for finding valid joint-pairs
#
# 
#
# In the above equation:
#
# L is the PAF;
#
# d is the vector joining two joints;
#
# p is the interpolated point between two joints;
#
# It is implemented using the dot product between the PAF and the vector $d_{ij}$
# Find valid connections between the different joints of a all persons present
def getValidPairs(output):
valid_pairs = []
invalid_pairs = []
n_interp_samples = 10
paf_score_th = 0.1
conf_th = 0.7
# loop for every POSE_PAIR
for k in range(len(mapIdx)):
# A->B constitute a limb
pafA = output[0, mapIdx[k][0], :, :]
pafB = output[0, mapIdx[k][1], :, :]
pafA = cv2.resize(pafA, (frameWidth, frameHeight))
pafB = cv2.resize(pafB, (frameWidth, frameHeight))
# Find the keypoints for the first and second limb
candA = detected_keypoints[POSE_PAIRS[k][0]]
candB = detected_keypoints[POSE_PAIRS[k][1]]
nA = len(candA)
nB = len(candB)
# If keypoints for the joint-pair is detected
# check every joint in candA with every joint in candB
# Calculate the distance vector between the two joints
# Find the PAF values at a set of interpolated points between the joints
# Use the above formula to compute a score to mark the connection valid
if( nA != 0 and nB != 0):
valid_pair = np.zeros((0,3))
for i in range(nA):
max_j=-1
maxScore = -1
found = 0
for j in range(nB):
# Find d_ij
d_ij = np.subtract(candB[j][:2], candA[i][:2])
norm = np.linalg.norm(d_ij)
if norm:
d_ij = d_ij / norm
else:
continue
# Find p(u)
interp_coord = list(zip(np.linspace(candA[i][0], candB[j][0], num=n_interp_samples),
np.linspace(candA[i][1], candB[j][1], num=n_interp_samples)))
# Find L(p(u))
paf_interp = []
for k in range(len(interp_coord)):
paf_interp.append([pafA[int(round(interp_coord[k][1])), int(round(interp_coord[k][0]))],
pafB[int(round(interp_coord[k][1])), int(round(interp_coord[k][0]))] ])
# Find E
paf_scores = np.dot(paf_interp, d_ij)
avg_paf_score = sum(paf_scores)/len(paf_scores)
# Check if the connection is valid
# If the fraction of interpolated vectors aligned with PAF is higher then threshold -> Valid Pair
if ( len(np.where(paf_scores > paf_score_th)[0]) / n_interp_samples ) > conf_th :
if avg_paf_score > maxScore:
max_j = j
maxScore = avg_paf_score
found = 1
# Append the connection to the list
if found:
valid_pair = np.append(valid_pair, [[candA[i][3], candB[max_j][3], maxScore]], axis=0)
# Append the detected connections to the global list
valid_pairs.append(valid_pair)
else: # If no keypoints are detected
print("No Connection : k = {}".format(k))
invalid_pairs.append(k)
valid_pairs.append([])
print(valid_pairs)
return valid_pairs, invalid_pairs
# This function creates a list of keypoints belonging to each person
# For each detected valid pair, it assigns the joint(s) to a person
# It finds the person and index at which the joint should be added. This can be done since we have an id for each joint
def getPersonwiseKeypoints(valid_pairs, invalid_pairs):
# the last number in each row is the overall score
personwiseKeypoints = -1 * np.ones((0, 19))
for k in range(len(mapIdx)):
if k not in invalid_pairs:
partAs = valid_pairs[k][:,0]
partBs = valid_pairs[k][:,1]
indexA, indexB = np.array(POSE_PAIRS[k])
for i in range(len(valid_pairs[k])):
found = 0
person_idx = -1
for j in range(len(personwiseKeypoints)):
if personwiseKeypoints[j][indexA] == partAs[i]:
person_idx = j
found = 1
break
if found:
personwiseKeypoints[person_idx][indexB] = partBs[i]
personwiseKeypoints[person_idx][-1] += keypoints_list[partBs[i].astype(int), 2] + valid_pairs[k][i][2]
# if find no partA in the subset, create a new subset
elif not found and k < 17:
row = -1 * np.ones(19)
row[indexA] = partAs[i]
row[indexB] = partBs[i]
# add the keypoint_scores for the two keypoints and the paf_score
row[-1] = sum(keypoints_list[valid_pairs[k][i,:2].astype(int), 2]) + valid_pairs[k][i][2]
personwiseKeypoints = np.vstack([personwiseKeypoints, row])
return personwiseKeypoints
image1 = cv2.imread("images/image1.jpg")
frameWidth = image1.shape[1]
frameHeight = image1.shape[0]
# #### Load the network and pass the image through the network
# +
t = time.time()
net = cv2.dnn.readNetFromCaffe(protoFile, weightsFile)
if device == "cpu":
net.setPreferableBackend(cv2.dnn.DNN_TARGET_CPU)
print("Using CPU device")
elif device == "gpu":
net.setPreferableBackend(cv2.dnn.DNN_BACKEND_CUDA)
net.setPreferableTarget(cv2.dnn.DNN_TARGET_CUDA)
print("Using GPU device")
# Fix the input Height and get the width according to the Aspect Ratio
inHeight = 368
inWidth = int((inHeight/frameHeight)*frameWidth)
inpBlob = cv2.dnn.blobFromImage(image1, 1.0 / 255, (inWidth, inHeight),
(0, 0, 0), swapRB=False, crop=False)
net.setInput(inpBlob)
output = net.forward()
print("Time Taken = {}".format(time.time() - t))
# -
# #### Slice a probability map ( for e.g Nose ) from the output for a specific keypoint and plot the heatmap ( after resizing ) on the image itself
i = 0
probMap = output[0, i, :, :]
probMap = cv2.resize(probMap, (frameWidth, frameHeight))
plt.figure(figsize=[14,10])
plt.imshow(cv2.cvtColor(image1, cv2.COLOR_BGR2RGB))
plt.imshow(probMap, alpha=0.6)
plt.colorbar()
plt.axis("off")
# +
detected_keypoints = []
keypoints_list = np.zeros((0,3))
keypoint_id = 0
threshold = 0.1
for part in range(nPoints):
probMap = output[0,part,:,:]
probMap = cv2.resize(probMap, (image1.shape[1], image1.shape[0]))
# plt.figure()
# plt.imshow(255*np.uint8(probMap>threshold))
keypoints = getKeypoints(probMap, threshold)
print("Keypoints - {} : {}".format(keypointsMapping[part], keypoints))
keypoints_with_id = []
for i in range(len(keypoints)):
keypoints_with_id.append(keypoints[i] + (keypoint_id,))
keypoints_list = np.vstack([keypoints_list, keypoints[i]])
keypoint_id += 1
detected_keypoints.append(keypoints_with_id)
# -
frameClone = image1.copy()
for i in range(nPoints):
for j in range(len(detected_keypoints[i])):
cv2.circle(frameClone, detected_keypoints[i][j][0:2], 3, [0,0,255], -1, cv2.LINE_AA)
plt.figure(figsize=[15,15])
plt.imshow(frameClone[:,:,[2,1,0]])
valid_pairs, invalid_pairs = getValidPairs(output)
personwiseKeypoints = getPersonwiseKeypoints(valid_pairs, invalid_pairs)
# +
for i in range(17):
for n in range(len(personwiseKeypoints)):
index = personwiseKeypoints[n][np.array(POSE_PAIRS[i])]
if -1 in index:
continue
B = np.int32(keypoints_list[index.astype(int), 0])
A = np.int32(keypoints_list[index.astype(int), 1])
cv2.line(frameClone, (B[0], A[0]), (B[1], A[1]), colors[i], 3, cv2.LINE_AA)
plt.figure(figsize=[15,15])
plt.imshow(frameClone[:,:,[2,1,0]])
# -
"""
Question - Run the openpose on the images provided in pat-1 folder and try to reason about
why it’s not working for some of the images and what’s the way to fix it?
"""
| Vision Meets ML/HW4/Part_1_origin/multi-person-openpose_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import confusion_matrix
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
dataset = pd.read_csv("/home/praveen/Desktop/Python/Machine learning/K-Nearest Neighbour/dataset/diabetes.csv")
dataset.head()
#replace zeros
zero_not_accepted = ['Glucose','BloodPressure','SkinThickness','BMI','Insulin']
for column in zero_not_accepted:
dataset[column] = dataset[column].replace(0,np.NaN)
mean = int(dataset[column].mean(skipna=True))
dataset[column] = dataset[column].replace(np.NaN,mean)
x = dataset.iloc[:,0:8]
y = dataset.iloc[:,8]
x_train,x_test,y_train,y_test = train_test_split(x,y,random_state=0,test_size=0.2)
sc_X = StandardScaler()
x_train = sc_X.fit_transform(x_train)
x_test = sc_X.fit_transform(x_test)
x_train
import math
math.sqrt(len(y_test))
classifier = KNeighborsClassifier(n_neighbors=11,p=2,metric='euclidean')
classifier.fit(x_train,y_train)
y_pred=classifier.predict(x_test)
y_pred
cm = confusion_matrix(y_test,y_pred)
print(cm)
print(f1_score(y_test,y_pred))
print(accuracy_score(y_test,y_pred))
| K-Nearest Neighbour/KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from dl import authClient as ac, queryClient as qc, storeClient as sc
from dl.helpers.utils import convert
from dl.helpers import crossmatch
from getpass import getpass
import pandas as pd
token = ac.login(input("User name: "),getpass("Password: "))
query = "select ra,dec,dered_mag_g,dered_mag_r from ls_dr5.tractor limit 1000"
res = qc.query(token,query)
print(res)
df = convert(res,'pandas')
print(df.head())
| project/.ipynb_checkpoints/RNtutorial-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import os
import numpy as np
import pandas as pd
#feature extraction in pipeline--------->
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.naive_bayes import MultinomialNB
# + deletable=true editable=true
ppt=[]
os.chdir("S:\\op_spam_train\\positive_polarity\\true")
for filename in os.listdir(os.getcwd()):
with open(filename,'r') as file:
ppt.append(file.read())
ppf=[]
os.chdir("S:\\op_spam_train\\positive_polarity\\false")
for filename in os.listdir(os.getcwd()):
with open(filename,'r') as file:
ppf.append(file.read())
npt=[]
os.chdir("S:\\op_spam_train\\negative_polarity\\true")
for filename in os.listdir(os.getcwd()):
with open(filename,'r') as file:
npt.append(file.read())
npf=[]
os.chdir("S:\\op_spam_train\\negative_polarity\\false")
for filename in os.listdir(os.getcwd()):
with open(filename,'r') as file:
npf.append(file.read())
deceptive=[]
deceptive=ppf+npf
truthful=[]
truthful=ppt+npt
data=[]
data=deceptive+truthful
data=np.array(data)
#train_data contains all the deceptive reviews from the positive part as a list
#train_data[0]='first review' and so on
# + deletable=true editable=true
features=['review']
data_df = pd.DataFrame(data, columns=features)
for label, row in data_df.iterrows():
data_df.loc[label, 'review_len']=len(row['review'])
index=np.arange(1600)
data_df['index'] = index
for label, row in data_df.iterrows():
if (int(row['index'])<800):
data_df.loc[label, 'label']=0
else:
data_df.loc[label, 'label']=1
data_df
# -
X = data_df.review
y = data_df.label
print(X.shape)
#right now X is a 1-d we will pass it to the vectorizer to convert it into 2-d
print(y.shape)
# +
# Now we split X and y into training and testing sets
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# + deletable=true editable=true
#Vectorizing the dataset
from sklearn.feature_extraction.text import CountVectorizer
vect = CountVectorizer(ngram_range=(1,2))
X_train_dtm = vect.fit_transform(X_train)
X_test_dtm = vect.transform(X_test)
mnb = MultinomialNB()
mnb.fit(X_train_dtm, y_train)
#making pedictions
y_predict = mnb.predict(X_test_dtm)
from sklearn import metrics
print(metrics.accuracy_score(y_test, y_predict))
print(metrics.confusion_matrix(y_test, y_predict))
'''Confusion matrix
[TN FP
FN TP]'''
# + deletable=true editable=true
'''Pipepline implementation'''
pipeline = Pipeline([
('features', FeatureUnion([
('ngram_tf_idf', Pipeline([
('counts_ngram', CountVectorizer(ngram_range=(1,5), analyzer='char')),
('tf_idf_ngram', TfidfTransformer())
])),
])),
('classifier', MultinomialNB())
])
pipeline.fit(X_train, y_train)
pipeline.predict(X_test)
print(metrics)
print(metrics.confusion_matrix(y_test, y_predict))
| model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Note: You need to reset the kernel for the keras installation to take place
#Todo: Remove this line once it is installed, reset the kernel: Menu > Kernel > Reset & Clear Output
# !git clone https://github.com/fchollet/keras.git && cd keras && python setup.py install --user
import keras
from keras.applications.inception_resnet_v2 import InceptionResNetV2
from keras.preprocessing import image
from keras.engine import Layer
from keras.applications.inception_resnet_v2 import preprocess_input
from keras.layers import Conv2D, UpSampling2D, InputLayer, Conv2DTranspose, Input, Reshape, merge, concatenate, Activation, Dense, Dropout, Flatten
from keras.layers.normalization import BatchNormalization
from keras.callbacks import TensorBoard
from keras.models import Sequential, Model
from keras.layers.core import RepeatVector, Permute
from keras.preprocessing.image import ImageDataGenerator, array_to_img, img_to_array, load_img
from skimage.color import rgb2lab, lab2rgb, rgb2gray, gray2rgb
from skimage.transform import resize
from skimage.io import imsave
import numpy as np
import os
import random
import tensorflow as tf
# +
# Get images
# Change to '/data/images/Train/' to use all the 10k images
X = []
for filename in os.listdir('colornet/'):
X.append(img_to_array(load_img('colornet/'+filename)))
X = np.array(X, dtype=float)
Xtrain = 1.0/255*X
#Load weights
inception = InceptionResNetV2(weights=None, include_top=True)
inception.load_weights('/data/inception_resnet_v2_weights_tf_dim_ordering_tf_kernels.h5')
inception.graph = tf.get_default_graph()
# +
embed_input = Input(shape=(1000,))
#Encoder
encoder_input = Input(shape=(256, 256, 1,))
encoder_output = Conv2D(64, (3,3), activation='relu', padding='same', strides=2)(encoder_input)
encoder_output = Conv2D(128, (3,3), activation='relu', padding='same')(encoder_output)
encoder_output = Conv2D(128, (3,3), activation='relu', padding='same', strides=2)(encoder_output)
encoder_output = Conv2D(256, (3,3), activation='relu', padding='same')(encoder_output)
encoder_output = Conv2D(256, (3,3), activation='relu', padding='same', strides=2)(encoder_output)
encoder_output = Conv2D(512, (3,3), activation='relu', padding='same')(encoder_output)
encoder_output = Conv2D(512, (3,3), activation='relu', padding='same')(encoder_output)
encoder_output = Conv2D(256, (3,3), activation='relu', padding='same')(encoder_output)
#Fusion
fusion_output = RepeatVector(32 * 32)(embed_input)
fusion_output = Reshape(([32, 32, 1000]))(fusion_output)
fusion_output = concatenate([encoder_output, fusion_output], axis=3)
fusion_output = Conv2D(256, (1, 1), activation='relu', padding='same')(fusion_output)
#Decoder
decoder_output = Conv2D(128, (3,3), activation='relu', padding='same')(fusion_output)
decoder_output = UpSampling2D((2, 2))(decoder_output)
decoder_output = Conv2D(64, (3,3), activation='relu', padding='same')(decoder_output)
decoder_output = UpSampling2D((2, 2))(decoder_output)
decoder_output = Conv2D(32, (3,3), activation='relu', padding='same')(decoder_output)
decoder_output = Conv2D(16, (3,3), activation='relu', padding='same')(decoder_output)
decoder_output = Conv2D(2, (3, 3), activation='tanh', padding='same')(decoder_output)
decoder_output = UpSampling2D((2, 2))(decoder_output)
model = Model(inputs=[encoder_input, embed_input], outputs=decoder_output)
# +
#Create embedding
def create_inception_embedding(grayscaled_rgb):
grayscaled_rgb_resized = []
for i in grayscaled_rgb:
i = resize(i, (299, 299, 3), mode='constant')
grayscaled_rgb_resized.append(i)
grayscaled_rgb_resized = np.array(grayscaled_rgb_resized)
grayscaled_rgb_resized = preprocess_input(grayscaled_rgb_resized)
with inception.graph.as_default():
embed = inception.predict(grayscaled_rgb_resized)
return embed
# Image transformer
datagen = ImageDataGenerator(
shear_range=0.2,
zoom_range=0.2,
rotation_range=20,
horizontal_flip=True)
#Generate training data
batch_size = 20
def image_a_b_gen(batch_size):
for batch in datagen.flow(Xtrain, batch_size=batch_size):
grayscaled_rgb = gray2rgb(rgb2gray(batch))
embed = create_inception_embedding(grayscaled_rgb)
lab_batch = rgb2lab(batch)
X_batch = lab_batch[:,:,:,0]
X_batch = X_batch.reshape(X_batch.shape+(1,))
Y_batch = lab_batch[:,:,:,1:] / 128
yield ([X_batch, create_inception_embedding(grayscaled_rgb)], Y_batch)
#Train model
tensorboard = TensorBoard(log_dir="/output")
model.compile(optimizer='adam', loss='mse')
model.fit_generator(image_a_b_gen(batch_size), callbacks=[tensorboard], epochs=1000, steps_per_epoch=20)
# -
# Save model
model_json = model.to_json()
with open("model.json", "w") as json_file:
json_file.write(model_json)
model.save_weights("color_tensorflow_real_mode.h5")
# +
#Make predictions on validation images
# Change to '/data/images/Test/' to use all the 500 test images
color_me = []
for filename in os.listdir('../Test/'):
color_me.append(img_to_array(load_img('../Test/'+filename)))
color_me = np.array(color_me, dtype=float)
color_me = 1.0/255*color_me
color_me = gray2rgb(rgb2gray(color_me))
color_me_embed = create_inception_embedding(color_me)
color_me = rgb2lab(color_me)[:,:,:,0]
color_me = color_me.reshape(color_me.shape+(1,))
# Test model
output = model.predict([color_me, color_me_embed])
output = output * 128
# Output colorizations
for i in range(len(output)):
cur = np.zeros((256, 256, 3))
cur[:,:,0] = color_me[i][:,:,0]
cur[:,:,1:] = output[i]
imsave("result/img_"+str(i)+".png", lab2rgb(cur))
| floydhub/Full-version/full_version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/evaneschneider/parallel-programming/blob/master/cuda_julia_set.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 114} colab_type="code" id="TNAaaptVFD6o" outputId="f8ac8f2a-9ae3-4efd-91e7-684a65ba555e"
# !/usr/local/cuda/bin/nvcc --version
# + colab={} colab_type="code" id="F_zdIqOjdXep"
# !pip install git+git://github.com/andreinechaev/nvcc4jupyter.git
# + colab={} colab_type="code" id="LqYHrt-DFP87"
# %load_ext nvcc_plugin
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="LIBJsZDzFWE8" outputId="87fbba25-39d3-4e93-b67e-f849080c87f4"
# %%cu
#include <iostream>
int main() {
std::cout << "Hello world!";
return 0;
}
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="RGextq50nrbZ" outputId="01b725f0-de82-4353-c1d9-db413a349d3b"
# %%cu
#include<stdio.h>
#include<stdlib.h>
#include<iostream>
#define DIM 1000
struct cuComplex {
float r;
float i;
__device__ cuComplex( float a, float b ) : r(a), i(b) {}
__device__ float magnitude2( void ) {
return r * r + i * i;
}
__device__ cuComplex operator*(const cuComplex& a) {
return cuComplex(r*a.r - i*a.i, i*a.r + r*a.i);
}
__device__ cuComplex operator+(const cuComplex& a) {
return cuComplex(r+a.r, i+a.i);
}
};
__device__ int julia( int x, int y ) {
const float scale = 1.5;
float jx = scale * (float)(DIM/2 - x)/(DIM/2);
float jy = scale * (float)(DIM/2 - y)/(DIM/2);
cuComplex c(-0.8, 0.156);
cuComplex a(jx, jy);
int i = 0;
for (i=0; i<200; i++) {
a = a * a + c;
if (a.magnitude2() > 1000)
return 0;
}
return 1;
}
__global__ void kernel( int *ptr ) {
// map from threadIdx/BlockIdx to pixel position
int x = blockIdx.x;
int y = blockIdx.y;
int offset = x + y * gridDim.x;
// now calculate the value at that position
int juliaValue = julia( x, y );
ptr[offset] = juliaValue;
}
int main( void ) {
int bitmap[DIM*DIM];
int *dev_bitmap;
cudaMalloc( (void**)&dev_bitmap, DIM*DIM*sizeof(int) );
dim3 grid(DIM,DIM);
kernel<<<grid,1>>>( dev_bitmap );
cudaMemcpy( bitmap, dev_bitmap, DIM*DIM*sizeof(int), cudaMemcpyDeviceToHost );
cudaFree( dev_bitmap );
//for (int i=0; i<DIM*DIM; i++) printf("%d ", bitmap[i]);
FILE *myfile;
myfile=fopen("/tmp/test.bin","wb");
if (!myfile) {
printf("Unable to open file!");
return 1;
}
fwrite(bitmap, sizeof(int), DIM*DIM, myfile);
fclose(myfile);
std::cout << "Success!";
}
# + colab={"base_uri": "https://localhost:8080/", "height": 504} colab_type="code" id="98rMyqVOvtnW" outputId="dbe580d4-0e32-444f-a605-57396ac55eeb"
import matplotlib.pyplot as plt
import numpy as np
bitmap = np.fromfile('/tmp/test.bin', dtype=np.int32)
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111)
ax.imshow(a.T, origin="lower")
# + colab={"base_uri": "https://localhost:8080/", "height": 65} colab_type="code" id="m6dvvGPtdsko" outputId="3cfd1f56-30c5-402b-ca68-9b37e1d022c0"
# !rm /tmp/test.bin
| Session8/Day5/cuda_julia_set.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Ruby 2.5.3
# language: ruby
# name: ruby
# ---
require 'daru/view'
Daru::View.plotting_library = :googlecharts
# +
data = [
['Director (Year)', '<NAME>', 'IMDB'],
['<NAME> (1935)', 8.4, 7.9],
['<NAME> (1959)', 6.9, 6.5],
['<NAME> (1978)', 6.5, 6.4],
['<NAME> (2008)', 4.4, 6.2]
]
stepped_area_chart_table = Daru::View::Table.new(data)
stepped_area_chart_table.show_in_iruby
# +
stepped_area_chart_options = {
type: :steppedArea, height: 400
}
stepped_area_chart_chart = Daru::View::Plot.new(stepped_area_chart_table.table, stepped_area_chart_options)
stepped_area_chart_chart.show_in_iruby
# +
stepped_area_chart_options = {
title: 'The decline of \'The 39 Steps\'',
vAxis: {title: 'Accumulated Rating'},
isStacked: true,
type: :steppedArea, height: 400
}
stepped_area_chart_chart = Daru::View::Plot.new(stepped_area_chart_table.table, stepped_area_chart_options)
stepped_area_chart_chart.show_in_iruby
# +
stepped_area_chart_options = {
backgroundColor: '#ddd',
legend: { position: 'bottom' },
connectSteps: false,
colors: ['#4374E0', '#53A8FB'],
isStacked: true,
type: :steppedArea, height: 400
}
stepped_area_chart_chart = Daru::View::Plot.new(stepped_area_chart_table.table, stepped_area_chart_options)
stepped_area_chart_chart.show_in_iruby
# -
data = [
['Month', 'Bolivia', 'Ecuador', 'Madagascar', 'Papua New Guinea', 'Rwanda', 'Average'],
['2004/05', 165, 938, 522, 998, 450, 614.6],
['2005/06', 135, 1120, 599, 1268, 288, 682],
['2006/07', 157, 1167, 587, 807, 397, 623],
['2007/08', 139, 1110, 615, 968, 215, 609.4],
['2008/09', 136, 691, 629, 1026, 366, 569.6]
]
stepped_area_table = Daru::View::Table.new(data)
stepped_area_table.show_in_iruby
# +
stepped_area_chart_options = {
isStacked: true,
legend: {position: 'top', maxLines: 3},
vAxis: {minValue: 0},
type: :steppedArea, height: 400
}
stepped_area_chart = Daru::View::Plot.new(stepped_area_table.table, stepped_area_chart_options)
stepped_area_chart.show_in_iruby
# +
stepped_area_chart_options = {
isStacked: 'relative',
height: 300,
legend: {position: 'top', maxLines: 3},
vAxis: {
minValue: 0,
ticks: [0, 0.3, 0.6, 0.9, 1]
},
type: :steppedArea, height: 400
}
stepped_area_chart = Daru::View::Plot.new(stepped_area_table.table, stepped_area_chart_options)
stepped_area_chart.show_in_iruby
# -
| spec/dummy_iruby/Google Charts - Stepped Area Chart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 1: Introduction to Exploratory Data Analysis
# In this notebook we dive into some plotting methods commonly used for Exploratory Data Analysis (EDA).
#
# Our goals for EDA are to open-mindedly explore the data, and see what insights we may find.
#
# The purpose of the EDA approach is to:
# - maximize insight into a data set
# - uncover underlying structure
# - extract important variables
# - detect outliers and anomalies
# - test underlying assumptions
# - develop parsimonious models
# - determine optimal factor settings
# In this notebook we'll investigate these plotting techniques:
#
# 1. Scatter Plot
# 1. Scatter Matrix
# 1. Histogram
# 1. Bar Plot
# 1. Box Plot
# 1. Time Series
# ### Setup
# <a id='setup'></a>
# +
from datetime import datetime
import dateutil.parser
import re
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# The command below means that the output of multiple commands in a cell will be output at once
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# The command below tells jupyter to display up to 80 columns, this keeps everything visible
pd.set_option('display.max_columns', 80)
pd.set_option('expand_frame_repr', True)
sns.set_palette("hls")
# %matplotlib inline
# -
# With each notebook, we need to read in our dataset
path = '../data/'
filename = 'loans.csv'
df = pd.read_csv(path+filename)
# Before diving into our exploratory data analysis, it is worth reiterating that this whole process is about understanding the distribution of data and relationships between different features.
#
# When we move on to use machine learning algorithms, we will be asking a question and trying to answer it using the statistical relationships between different features in the data. The EDA analysis will help us shape this question and have a clear idea about how to approach building the algorithm!
#
# With that in mind, let's look at several visualization methods to examine the data and any relationships between features…
# ### 1. Scatter plot
# To start, the scatter plot! This is a very popular and powerful way to visualize the relationship between two continuous features. Essentially this plot shows us how feature Y changes when feature X is changed. If there is a clear pattern formed in the scatter plot, we say that x and y are **correlated**.
#
# There are several outcomes we see on a scatter plot:
# - Positive Linear = When X increases, y increases and the data points follow an approximate straight line
# - Negative Linear = When X increase, y decreases and the data points follow an approximate straight line
# - Non-Linear = When X increases, there is a consistent change in Y but this is not linear. It could be quadratic or exponential for example.
# - No correlation = When X increases, there is no clear pattern to how y changes, The data points form a random distribution.
#
# Let's try this out on our data and choose two continuous variables to plot. First lets extract all the continuous variables from our dataset.
numeric_vars = df.select_dtypes(include=[np.number]).columns.tolist()
for variable in numeric_vars:
print(variable)
# To start, let's look if there is a relationship between lender_count and loan_amount... intuition suggests that bigger loans much have more lenders. If this is true, we'll see this in the scatter plot!
ax = sns.regplot(x='lender_count', y='loan_amount', data=df)
# Where does the data follow the line?
#
# Where does the data __not__ follow the line?
#
# What are possible reasons that data does __not__ follow the line?
# -----
# Let's explore another relationship.
# ------
#
# How about the repayment term and the loan amount?
#
# What kind of relationship would you expect between the repayment term and the loan amount?
ax = sns.regplot(x='repayment_term',
y='loan_amount',
data=df)
# Where does the data follow the line?
#
# Where does the data __not__ follow the line?
#
# What are possible reasons that data does __not__ follow the line?
# ### 2. Scatter Matrix
#
# When we have lots of continuous variables, we could go through them one by one to see the relationship or we could use a scatterplot matrix! This creates a scatter plot between every combination of variables in a list.
#
# Another interesting quality of the scatter matrix is that the diagonals give a histogram of the variable in question.
# +
# Let's choose only a couple of columns to examine:
columns = ['loan_amount', 'funded_amount', 'status']
num_df = df[columns]
# Remove the NaN rows so Seaborn can plot
num_df = num_df.dropna(axis=0, how='any')
# Create the scatter plot and let's color the data point by their status.
sns.pairplot(num_df, hue='status');
# -
# What can say about the data?
#
# <br>
# <br>
# <br>
# ### 4. Histogram
#
# A histogram is useful for looking at the distribution of values for a single variable and also identifying outliers. It shows us the count of data.
#
# The plot below shows the data distribution of loan_amount using both bars and a continuous line. Without going into too much detail about the value on the y-axis, what we can take away from this is there is a much higher occurrence of small loans (high bar/peak in the line) and that large loans are much rarer (low bars/drop in the line).
sns.distplot(df['loan_amount'].dropna(axis = 0));
# Let's just look at those under 5K
small_loans_df = df[(df['loan_amount'] < 5000)]
sns.distplot(small_loans_df['loan_amount']);
# Looking at the loans less than 5000 we see a much clearer distribution, although it is still left-hand skewed.
# ### 5. Bar Plot
#
# Bar plots are useful for understanding how categorical groups are different with respect to a continuous variable.
#
p = sns.barplot(x='sector', y = 'loan_amount', data=df, estimator=np.mean);
p.set(title='Average loan amount by sector')
p.set_xticklabels(p.get_xticklabels(), rotation=-45);
# Which sector is the largest? Why?
p = sns.barplot(x='sector', y = 'loan_amount', data=df, estimator=np.sum);
p.set(title='Total loan amount by sector')
p.set_xticklabels(p.get_xticklabels(), rotation=-45);
# Which sector is the largest? Why?
#
# <br>
# <br>
# ### 6. Box Plots
#
# A box plot describes the distribution of data based on five important summary numbers: the minimum, first quartile, median, third quartile, and maximum. In the simplest box plot the central rectangle spans the first quartile to the third quartile (the interquartile range or IQR). A segment inside the rectangle shows the median and "whiskers" above and below the box show the locations of the minimum and maximum.
#
# Lets use this to look at the distribution of borrowers counts by each sector for different loan status for different partners. First lets look at how many loans come from different partners.
df_retail = df[df.sector=='Retail']
df_retail.head()
sector = 'Retail'
df_retail = df[df.sector==sector]
p = sns.boxplot(x='sector',
y='loan_amount',
data=df_retail);
p.set(title = f'Loan amounts for {sector}');
p.set_xticklabels(p.get_xticklabels(), rotation=-45);
# Try this - Select other sectors and see how they look
# Aha! It looks like we are onto something here... we can see different trends for different partners! We'll look into this further in feature_engineering to see how we can use to create powerful features.
# ### 7. Time dependancy
# Quite often it's useful to see how a variable changes over time. This means creating a plot with time on the x-axis and the variable on the y-axis.
#
# Lets have a look at how the average loan amount changes over time on a monthly basis.
# Convert posted date to a datetime object
time_column = 'funded_date'
df[time_column] = pd.to_datetime(df[time_column])
# Resample the date to monthly intervals , taking the mean of loan_amount
# This creates an array where the index is the timestamp and the value is the mean of loan amount
time_data = df.resample('M', on=time_column)['loan_amount'].mean().fillna(0)
fig, ax = plt.subplots(figsize=(15,8))
ax.plot(time_data)
plt.title('Mean loan_amount over time');
# We can look at different timefrance by changing the parameter in resample. Lets look on a weekly basis!
# Resample the date to monthly intervals , taking the mean of loan_amount
# This creates an array where the index is the timestamp and the value is the mean of loan amount
time_data = df.resample('7D', on=time_column)['loan_amount'].mean().fillna(0)
fig, ax = plt.subplots(figsize=(15,8))
ax.plot(time_data)
plt.title('Mean loan_amount over time');
# What is next
# ------
#
# Next we move on to feature engineering, where we create variables from what we've found!
# <br>
# <br>
# <br>
#
# ----
| module_1_introduction/1_4_exploratory_data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="mvePf4fVelQ0" colab_type="code" colab={}
def getKey(bs):
# print(bs)
pp = bs.replace('/home/ishadij/TexyGan/gen_data/', '').replace('coco/','').replace('\n','').replace('.txt','')
# pp = bs.replace('/home/ishadij/TexyGan/gen_data/', '').replace('emnlp/','').replace('\n','').replace('.txt','')
gan_pp, key_pp = pp.split('/')
gan_pp, key_pp
key_pp, val = key_pp.split(':')
val = float(val)
return gan_pp,key_pp, val
# getKey('/home/ishadij/TexyGan/gen_data/mle/coco/1_200_0.001.txt: 0.27467031433851313\n')
# + id="v1ivilFvlA_d" colab_type="code" colab={}
temps = [0.001,0.5,1.0,1.5,2.0]
sizes = [200, 400, 600, 800, 1000]
gans = ['seqgan', 'gsgan','mle', 'leakgan', 'textgan']
sbs = list()
with open('selfBLEU') as sfbl:
for hypothesis in sfbl:
sbs.append(hypothesis)
tbs = list()
with open('testBLEU') as sfbl:
for hypothesis in sfbl:
tbs.append(hypothesis)
# + id="wbg9iFbiHBN2" colab_type="code" colab={}
gan_tb_dict = dict()
gan_sb_dict = dict()
tb_dict = dict()
sb_dict = dict()
for gan in gans:
tb_dict[gan] = dict()
sb_dict[gan] = dict()
gan_tb_dict[gan] = dict()
gan_sb_dict[gan] = dict()
for tb in tbs:
gan_name, key, val = getKey(tb)
gan_tb_dict[gan_name][key] = val
for sb in sbs:
gan_name, key, val = getKey(sb)
gan_sb_dict[gan_name][key] = val
# + id="MlGy_8IOed-G" colab_type="code" outputId="2ddadc3a-257f-4470-9262-7e494bd412c1" colab={"base_uri": "https://localhost:8080/", "height": 2142}
for gan in gans:
gan_tb_val = gan_tb_dict[gan]
gan_sb_val = gan_sb_dict[gan]
for size in sizes:
for temp in temps:
half_key = str(size)+'_'+str(temp)
val_1 = gan_tb_val[str(1)+'_'+half_key]
val_2 = gan_tb_val[str(2)+'_'+half_key]
val_3 = gan_tb_val[str(3)+'_'+half_key]
meanVal = (val_1+val_2+val_3)/3
tb_dict[gan][half_key] = 1-meanVal
val_1 = gan_sb_val[str(1)+'_'+half_key]
val_2 = gan_sb_val[str(2)+'_'+half_key]
val_3 = gan_sb_val[str(3)+'_'+half_key]
meanVal2 = (val_1+val_2+val_3)/3
sb_dict[gan][half_key] = meanVal2
print(gan,half_key, 1- meanVal,meanVal2 )
# + id="dpqHTWbxmdMG" colab_type="code" outputId="a9fa77a2-af2c-45c5-e6cf-d1bb73c53158" colab={"base_uri": "https://localhost:8080/", "height": 442}
import numpy as np
from numpy import trapz
import matplotlib.pyplot as plt
from scipy.integrate import simps
for gan in gans:
gan_tb_val = tb_dict[gan]
gan_sb_val = sb_dict[gan]
for size in sizes:
x = list()
y = list()
x_ord = list()
y_ord = list()
x_ord1 = list()
y_ord1 = list()
for temp in temps:
half_key = str(size)+'_'+str(temp)
x.append(tb_dict[gan][half_key])
y.append(sb_dict[gan][half_key])
x_args = np.argsort(x)
for id in x_args:
x_ord.append(x[id])
y_ord.append(y[id])
y_args = np.argsort(y)
for id1 in y_args:
x_ord1.append(y[id1])
y_ord1.append(x[id1])
area1 = trapz(y_ord, dx = 1)
area2 = trapz(y_ord1, dx = 1)
area = area1+area2
# run once without the division, then run again dividing by the max float seen in the output. This is for scaling
# print(gan,',', size,',',area2/3.999969293219355)
# print(gan,',', size,',',area1/3.9892631102131144)
print(gan,',', size,',',area/7.958946710431023)
| Texygen-master/AreaUtil.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook illustrates how to perform Linear Regression using scikit-learn. In simple linear regression we examine the relationship between an independent numerical variable and a dependent numerical variable.
| MACHINE_LEARNING/.ipynb_checkpoints/Linear Regression in Python-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AC Notes
# ## Files
# ### 20200303_ForestPotential_Samples.csv
# - system:index
# - CHELSA_Annual_Mean_Temperature
# - CHELSA_Annual_Precipitation
# - CHELSA_Mean_Temperature_of_Warmest_Quarter
# - CHELSA_Precipitation_Seasonality
# - CHELSA_Precipitation_of_Driest_Quarter
# - EarthEnvTopoMed_Eastness
# - EarthEnvTopoMed_Elevation
# - EarthEnvTopoMed_Northness
# - Lat
# - Long
# - Resolve_Biome
# - SG_Depth_to_bedrock
# - SG_Sand_Content_000cm
# - SG_Sand_Content_005cm
# - shrubcover
# - treecover
# - .geo
#
# ### 20200303_ForestPotential_PointSampling
# No idea - Javascript file
#
# Import the necessary modules for both Earth Engine work as well as map visualization within a Jupyter notebook
import ee as ee
ee.Initialize()
import folium
import geehydro #AC This isnt used anywhere
import time
import datetime
import numpy as np
import subprocess
# ### users/devinrouth/Resolve_Biomes_30ArcSec dataset
# Has 14 biomes:
# https://developers.google.com/earth-engine/datasets/catalog/RESOLVE_ECOREGIONS_2017
# ### users/devinrouth/Future_BioClim_Ensembles/rcp* datasets
# These are WorldClim data with added variables for ecology.
# BIOCLIM variables https://www.edenextdata.com/?q=content/bioclim-worldclim-bioclimatic-variables-2000-20-50-80
# The years are 2030,2050,2070,2080
# The RCPS are 4.5 and 8.5
# Have 19 variables:
# Annual_Mean_Temperature: 29.585416666666667
# Mean_Diurnal_Range: 9.506770833333333
# Isothermality: 68.97395833333333
# Temperature_Seasonality: 119.45416666666667
# Max_Temperature_of_Warmest_Month: 37.36770833333333
# Min_Temperature_of_Coldest_Month: 23.684375
# Temperature_Annual_Range: 13.683333333333334
# Mean_Temperature_of_Wettest_Quarter: 28.03333333333333
# Mean_Temperature_of_Driest_Quarter: 30.833854166666665
# Mean_Temperature_of_Warmest_Quarter: 31.144791666666666
# Mean_Temperature_of_Coldest_Quarter: 27.9859375
# Annual_Precipitation: 1932.15625
# Precipitation_of_Wettest_Month: 277.9375
# Precipitation_of_Driest_Month: 15.052083333333334
# Precipitation_Seasonality: 58.651041666666664
# Precipitation_of_Wettest_Quarter: 782.515625
# Precipitation_of_Driest_Quarter: 77.85416666666667
# Precipitation_of_Warmest_Quarter: 140.3125
# Precipitation_of_Coldest_Quarter: 780.9010416666666
# ### "users/devinrouth/ETH_Composites/CrowtherLab_Composite_30ArcSec" dataset
# This has 302 bands!! Including:
# Aridity index
# depth to water table
# EVI
# Cloud cover statistics
# Texture measurements including Shannon and Simpson
# Aspect, elevation, roughness, slope
# FPAR
# Above ground biomass
# Hansen Forest cover
# Human footprint
# Landcover class
# Mycorrhizae
# NDVI ?
# Nematodes
# Lots of soil properties
# Tree density
# Chelsa Climate data http://chelsa-climate.org/
# +
# Load the datasets of interest
resolveBiomes = ee.Image("users/devinrouth/Resolve_Biomes_30ArcSec")
rcp45_2080s_Mean = ee.Image("users/devinrouth/Future_BioClim_Ensembles/rcp45_2080s_Mean")
rcp45_2030s_Mean = ee.Image("users/devinrouth/Future_BioClim_Ensembles/rcp45_2030s_Mean")
rcp45_2050s_Mean = ee.Image("users/devinrouth/Future_BioClim_Ensembles/rcp45_2050s_Mean")
rcp45_2070s_Mean = ee.Image("users/devinrouth/Future_BioClim_Ensembles/rcp45_2070s_Mean")
rcp85_2030s_Mean = ee.Image("users/devinrouth/Future_BioClim_Ensembles/rcp85_2030s_Mean")
rcp85_2050s_Mean = ee.Image("users/devinrouth/Future_BioClim_Ensembles/rcp85_2050s_Mean")
rcp85_2070s_Mean = ee.Image("users/devinrouth/Future_BioClim_Ensembles/rcp85_2070s_Mean")
rcp85_2080s_Mean = ee.Image("users/devinrouth/Future_BioClim_Ensembles/rcp85_2080s_Mean")
composite = ee.Image("users/devinrouth/ETH_Composites/CrowtherLab_Composite_30ArcSec")
# Create an unbounded geometry for later use
# unboundedGeo = ee.Geometry.Rectangle([-180, -90, 180, 90], "EPSG:4326", False)
unboundedGeo = ee.Geometry.Polygon([-180, 88, 0, 88, 180, 88, 180, -88, 0, -88, -180, -88], None, False);
# !! Change testGeo when running full pipeline
testGeo = ee.Geometry.Polygon(
[[[8.453552577781124, 47.44586460369053],
[8.453552577781124, 47.2896051444241],
[8.728210780906124, 47.2896051444241],
[8.728210780906124, 47.44586460369053]]], None, False);
# Decide what geometry to use for exports (i.e., determining the extent of the maps)
exportingGeometry = unboundedGeo
# Create a list of biome number designations for later use
biomeNumberList = [1,2,3,4,5,6,7,8,9,10,11,12,13,14]
# Input the username folder where all of the images and collections will be saved
# usernameFolderString = 'devinrouth_backup'
usernameFolderString = 'acottam'
# Input the name of the project folder wherein all (top-level) items will be saved
# !! This folder must be created in order for the script to run
projectFolder = 'ETH_Biome_Future_Predictions'
# Input the name of the folder that will hold the bootstrapped samples
bootstrapCollFolder = 'Bootstrap_Samples'
# Select how many points per biome you'd like to sample
pointsPerBiome = 2500
# Select a tileScale to use across sampling calls
# AC The default is 1 and this may be why it takes so long?
tileScaleToUse = 16
# Input the normal wait time (in seconds) for "wait and break" cells
normalWaitTime = 60
# Input the long wait time (in seconds) for "wait and break" cells
longWaitTime = 600
# +
# Write a list of random numbers to serve as the seed for the bootstrap collections; the number of seeds will determine the number of bootstraps
# !! This list determines how many bootstrap samples will be created / modeled
#seedList = list(range(1,1001))
seedList = list(range(1,11))
print(seedList)
print('\n\n\n')
# Choose how many collections/images to use/make at a time when bootstrapping
n = 10
seedChunkListToMap = [seedList[i:i + n] for i in range(0, len(seedList), n)]
print(seedChunkListToMap)
print('\n\n\n')
# -
# Create bash variables in order to create/check/delete Earth Engine Assets
bashFunction = 'earthengine'
arglist_CreateCollection = ['--no-use_cloud_api','create','collection']
arglist_CreateFolder = ['--no-use_cloud_api','create','folder']
arglist_Detect = ['--no-use_cloud_api','asset','info']
arglist_Delete = ['--no-use_cloud_api','rm','-r']
stringsOfInterest = ['Asset does not exist or is not accessible']
bashCommandList_Detect = [bashFunction]+arglist_Detect
bashCommandList_Delete = [bashFunction]+arglist_Delete
bashCommandList_CreateCollection = [bashFunction]+arglist_CreateCollection
bashCommandList_CreateFolder = [bashFunction]+arglist_CreateFolder
# Make a scaling image to ensure the scale of the CHELSA images matches the other images
scalingDictionary = ee.Dictionary(
{
'CHELSA_Annual_Mean_Temperature':0.1,
'CHELSA_Annual_Precipitation':1,
'CHELSA_Isothermality':0.1,
'CHELSA_Max_Temperature_of_Warmest_Month':0.1,
'CHELSA_Mean_Diurnal_Range':0.1,
'CHELSA_Mean_Temperature_of_Coldest_Quarter':0.1,
'CHELSA_Mean_Temperature_of_Driest_Quarter':0.1,
'CHELSA_Mean_Temperature_of_Warmest_Quarter':0.1,
'CHELSA_Mean_Temperature_of_Wettest_Quarter':0.1,
'CHELSA_Min_Temperature_of_Coldest_Month':0.1,
'CHELSA_Precipitation_Seasonality':1,
'CHELSA_Precipitation_of_Coldest_Quarter':1,
'CHELSA_Precipitation_of_Driest_Month':1,
'CHELSA_Precipitation_of_Driest_Quarter':1,
'CHELSA_Precipitation_of_Warmest_Quarter':1,
'CHELSA_Precipitation_of_Wettest_Month':1,
'CHELSA_Precipitation_of_Wettest_Quarter':1,
'CHELSA_Temperature_Annual_Range':0.1,
'CHELSA_Temperature_Seasonality':0.1
});
# Scale the CHELSA bands from the composite to match the future predictions on these bands
# AC This is taking the above bands and multiplying their values by a constant image with the scale factors! Nice.
chelsaMultibandImage = composite.select(scalingDictionary.keys()).multiply(scalingDictionary.toImage());
# Compute lists of other variables to include in the models (i.e., soil and topographic variables)
# AC topComp has the topographic variables: slope, elevation and aspect
topComp = composite.select(['EarthEnvTopoMed_Slope','EarthEnvTopoMed_Elevation','EarthEnvTopoMed_Eastness','EarthEnvTopoMed_Northness']);
# AC soilComp has the soil variables: organic carbon, pH, density, sand, water capacity, cation exchange capacity (CEC)
soilComp = composite.select(['SG_CEC_000cm','SG_SOC_Density_000cm','SG_Soil_pH_H2O_000cm','SG_Bulk_density_000cm','SG_Sand_Content_000cm','SG_H2O_Capacity_000cm']);
# Concatenate the two images into a single variable
totComp = chelsaMultibandImage.addBands(soilComp).addBands(topComp);
# AC so the independent variables in the model are the following:
# CHELSA climate, topographic and soil variables
# These are used to train the Random Forest classifier
covariateNames = totComp.bandNames()
# First, make a dictionary to translate the band names
# AC Between the CHELSA climate data dataset and the RCP scenario data (e.g. users/devinrouth/Future_BioClim_Ensembles/rcp45_2080s_Mean)
chelsaDict =ee.Dictionary({
'Annual_Mean_Temperature':'CHELSA_Annual_Mean_Temperature',
'Mean_Diurnal_Range':'CHELSA_Mean_Diurnal_Range',
'Isothermality':'CHELSA_Isothermality',
'Temperature_Seasonality':'CHELSA_Temperature_Seasonality',
'Max_Temperature_of_Warmest_Month':'CHELSA_Max_Temperature_of_Warmest_Month',
'Min_Temperature_of_Coldest_Month':'CHELSA_Min_Temperature_of_Coldest_Month',
'Temperature_Annual_Range':'CHELSA_Temperature_Annual_Range',
'Mean_Temperature_of_Wettest_Quarter':'CHELSA_Mean_Temperature_of_Wettest_Quarter',
'Mean_Temperature_of_Driest_Quarter':'CHELSA_Mean_Temperature_of_Driest_Quarter',
'Mean_Temperature_of_Warmest_Quarter':'CHELSA_Mean_Temperature_of_Warmest_Quarter',
'Mean_Temperature_of_Coldest_Quarter':'CHELSA_Mean_Temperature_of_Coldest_Quarter',
'Annual_Precipitation':'CHELSA_Annual_Precipitation',
'Precipitation_of_Wettest_Month':'CHELSA_Precipitation_of_Wettest_Month',
'Precipitation_of_Driest_Month':'CHELSA_Precipitation_of_Driest_Month',
'Precipitation_Seasonality':'CHELSA_Precipitation_Seasonality',
'Precipitation_of_Wettest_Quarter':'CHELSA_Precipitation_of_Wettest_Quarter',
'Precipitation_of_Driest_Quarter':'CHELSA_Precipitation_of_Driest_Quarter',
'Precipitation_of_Warmest_Quarter':'CHELSA_Precipitation_of_Warmest_Quarter',
'Precipitation_of_Coldest_Quarter':'CHELSA_Precipitation_of_Coldest_Quarter'
});
# +
# Prepare all of the future imagery
# AC This selects the climate fields in the Bioclim data and renames them
rcp45_2030s_Mean = rcp45_2030s_Mean.select(chelsaDict.keys(),chelsaDict.values());
rcp45_2050s_Mean = rcp45_2050s_Mean.select(chelsaDict.keys(),chelsaDict.values());
rcp45_2070s_Mean = rcp45_2070s_Mean.select(chelsaDict.keys(),chelsaDict.values());
rcp45_2080s_Mean = rcp45_2080s_Mean.select(chelsaDict.keys(),chelsaDict.values());
rcp85_2030s_Mean = rcp85_2030s_Mean.select(chelsaDict.keys(),chelsaDict.values());
rcp85_2050s_Mean = rcp85_2050s_Mean.select(chelsaDict.keys(),chelsaDict.values());
rcp85_2070s_Mean = rcp85_2070s_Mean.select(chelsaDict.keys(),chelsaDict.values());
rcp85_2080s_Mean = rcp85_2080s_Mean.select(chelsaDict.keys(),chelsaDict.values());
# AC This adds the soil and topographic variables to the Bioclim data
rcp45_2030s_Mean_Soil = rcp45_2030s_Mean.addBands(soilComp).addBands(topComp);
rcp45_2050s_Mean_Soil = rcp45_2050s_Mean.addBands(soilComp).addBands(topComp);
rcp45_2070s_Mean_Soil = rcp45_2070s_Mean.addBands(soilComp).addBands(topComp);
rcp45_2080s_Mean_Soil = rcp45_2080s_Mean.addBands(soilComp).addBands(topComp);
rcp85_2030s_Mean_Soil = rcp85_2030s_Mean.addBands(soilComp).addBands(topComp);
rcp85_2050s_Mean_Soil = rcp85_2050s_Mean.addBands(soilComp).addBands(topComp);
rcp85_2070s_Mean_Soil = rcp85_2070s_Mean.addBands(soilComp).addBands(topComp);
rcp85_2080s_Mean_Soil = rcp85_2080s_Mean.addBands(soilComp).addBands(topComp);
# +
# Compute the training and test samples
trainSamplesToExport = totComp.addBands(resolveBiomes.rename('ResolveBiome').int()).stratifiedSample(
numPoints=pointsPerBiome,
classBand='ResolveBiome',
region=unboundedGeo,
seed=11111,
tileScale=tileScaleToUse,
geometries=True
)
validateSamplesToExport = totComp.addBands(resolveBiomes.rename('ResolveBiome').int()).stratifiedSample(
numPoints=pointsPerBiome,
classBand='ResolveBiome',
region=unboundedGeo,
seed=22222,
tileScale=tileScaleToUse,
geometries=True
)
testSamplesToExport = totComp.addBands(resolveBiomes.rename('ResolveBiome').int()).stratifiedSample(
numPoints=pointsPerBiome,
classBand='ResolveBiome',
region=unboundedGeo,
seed=33333,
tileScale=tileScaleToUse,
geometries=True
)
# +
# Perform the export then wait for it to finish before moving on
trainingSampleExport = ee.batch.Export.table.toAsset(
collection=trainSamplesToExport,
description='trainingSamples',
assetId='users/'+usernameFolderString+'/'+projectFolder+'/trainingSamples'
);
trainingSampleExport.start()
validateSampleExport = ee.batch.Export.table.toAsset(
collection=validateSamplesToExport,
description='validateSamples',
assetId='users/'+usernameFolderString+'/'+projectFolder+'/validateSamples'
);
validateSampleExport.start()
testSampleExport = ee.batch.Export.table.toAsset(
collection=testSamplesToExport,
description='testSamples',
assetId='users/'+usernameFolderString+'/'+projectFolder+'/testSamples'
);
testSampleExport.start()
# Sleep to ensure the jobs have been queued
time.sleep(normalWaitTime/2)
# + jupyter={"outputs_hidden": true}
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# -
# Load the collections
trainSamples = ee.FeatureCollection('users/'+usernameFolderString+'/'+projectFolder+'/trainingSamples')
validateSamples = ee.FeatureCollection('users/'+usernameFolderString+'/'+projectFolder+'/validateSamples')
testSamples = ee.FeatureCollection('users/'+usernameFolderString+'/'+projectFolder+'/testSamples')
# +
# Instantiate a selection of random forest classifiers to determine the best model (using features for wrapping, so the entire process can be processed via an export task)
rf_VP2 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP2','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=2,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP3 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP3','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=3,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP4 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP4','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=4,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP5 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP5','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=5,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP6 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP6','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=6,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP7 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP7','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=7,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP8 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP8','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=8,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP9 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP9','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=9,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP10 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP10','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=10,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP12 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP12','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=12,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
rf_VP14 = ee.Feature(ee.Geometry.Point([0,0])).set('cName','rf_VP14','c',ee.Classifier.smileRandomForest(
numberOfTrees=100,
variablesPerSplit=14,
bagFraction=0.632
).setOutputMode('CLASSIFICATION').train(trainSamples,'ResolveBiome',covariateNames))
# +
# Map through the trained classifiers, classify the validation samples, then compute their accuracy to compare them
classifierFC = ee.FeatureCollection([rf_VP2,
rf_VP3,
rf_VP4,
rf_VP5,
rf_VP6,
rf_VP7,
rf_VP8,
rf_VP9,
rf_VP10,
rf_VP12,
rf_VP14])
def outputValidationAccuracy(classiferFeature):
return ee.Feature(classiferFeature).set('OverallAccuracy',validateSamples.classify(ee.Feature(classiferFeature).get('c'),'PredictedBiome').errorMatrix('ResolveBiome','PredictedBiome').accuracy()).select(['OverallAccuracy','cName'])
# !! Export the accuracy FC
accuracyFC = classifierFC.map(outputValidationAccuracy).sort('OverallAccuracy',False)
finalClassifierFCExport = ee.batch.Export.table.toAsset(
collection=ee.FeatureCollection(accuracyFC),
description='finalClassifierFC',
assetId='users/'+usernameFolderString+'/'+projectFolder+'/finalClassifierFC'
);
finalClassifierFCExport.start()
# Sleep to ensure the job has been queued
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# !! Export the best model information separately
bestModelDictionary = ee.Feature(accuracyFC.first()).toDictionary()
finalValidationFCExport = ee.batch.Export.table.toAsset(
collection=ee.FeatureCollection(ee.Feature(ee.Geometry.Point([0,0])).set(bestModelDictionary)),
description='finalValidationFC',
assetId='users/'+usernameFolderString+'/'+projectFolder+'/finalValidationFC'
);
finalValidationFCExport.start()
# Sleep to ensure the job has been queued
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# Compute the final accuracy on the test collection
bestModelName = ee.Feature(ee.FeatureCollection('users/'+usernameFolderString+'/'+projectFolder+'/finalValidationFC').first()).get('cName')
bestModel = ee.Feature(classifierFC.filterMetadata('cName','equals',bestModelName).first()).get('c')
finalAccuracy = testSamples.classify(ee.Classifier(bestModel),'PredictedBiome').errorMatrix('ResolveBiome','PredictedBiome',biomeNumberList).accuracy()
kappaAccuracy = testSamples.classify(ee.Classifier(bestModel),'PredictedBiome').errorMatrix('ResolveBiome','PredictedBiome',biomeNumberList).kappa()
# !! Export the final accuracy
finalAccuracyFCExport = ee.batch.Export.table.toAsset(
collection=ee.FeatureCollection(ee.Feature(ee.Geometry.Point([0,0])).set('FinalAccuracy',finalAccuracy).set('KappaAccuracy',kappaAccuracy)),
description='finalAccuracyFC',
assetId='users/'+usernameFolderString+'/'+projectFolder+'/finalAccuracyFC'
);
finalAccuracyFCExport.start()
# Sleep to ensure the job has been queued
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# Return accuracy values here
finalAccuracyToPrint = ee.Feature(ee.FeatureCollection('users/'+usernameFolderString+'/'+projectFolder+'/finalAccuracyFC').first()).get('FinalAccuracy')
print('FinalAccuracy');
print(finalAccuracyToPrint.getInfo(),'\n')
kappaAccuracyToPrint = ee.Feature(ee.FeatureCollection('users/'+usernameFolderString+'/'+projectFolder+'/finalAccuracyFC').first()).get('KappaAccuracy')
print('KappaAccuracy');
print(kappaAccuracyToPrint.getInfo(),'\n')
# +
# Show the raw confusion matrix for the test data
# !! Use a while loop here to repeat the code in case there are Computation time out errors
while True:
try:
confusionMatrix = testSamples.classify(ee.Classifier(bestModel),'PredictedBiome').errorMatrix('ResolveBiome','PredictedBiome',biomeNumberList).array().getInfo()
print(confusionMatrix)
print('\n')
except Exception as e:
print(e,' : ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(30)
continue
else:
break
for r in confusionMatrix:
print(*r)
# +
# Show the consumers' accuracy for the test data
# !! Use a while loop here to repeat the code in case there are Computation time out errors
while True:
try:
consumersMatrix = testSamples.classify(ee.Classifier(bestModel),'PredictedBiome').errorMatrix('ResolveBiome','PredictedBiome',biomeNumberList).consumersAccuracy().getInfo()[0]
print(consumersMatrix)
print('\n')
except Exception as e:
print(e,' : ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(30)
continue
else:
break
for b in biomeNumberList:
print('Biome ',b,': ',consumersMatrix[b-1])
# +
# Print the producers' accuracy values
# !! Use a while loop here to repeat the code in case there are Computation time out errors
while True:
try:
producersMatrix = testSamples.classify(ee.Classifier(bestModel),'PredictedBiome').errorMatrix('ResolveBiome','PredictedBiome',biomeNumberList).producersAccuracy().getInfo()
print(producersMatrix)
print('\n')
except Exception as e:
print(e,' : ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(30)
continue
else:
break
for b in biomeNumberList:
print('Biome ',b,': ',producersMatrix[b-1][0])
# -
# # Bootstrapping
# +
# Create a folder to house the bootstrapped feature collection
# Turn the folder string into an assetID and perform the deletion
assetIDToCreate_Folder = 'users/'+usernameFolderString+'/'+projectFolder+'/'+bootstrapCollFolder
print(assetIDToCreate_Folder,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateFolder+[assetIDToCreate_Folder])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate_Folder], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# Write a function load each of the collections in, chunk by chunk, and use them to predict a composite
# Create the bootstrap collections for training
for seed in seedList:
assetIDString = str('BootstrapSamples_')+str(seed).zfill(4)
bootstrapSamples = totComp.addBands(resolveBiomes.rename('ResolveBiome').int()).stratifiedSample(
numPoints=pointsPerBiome,
classBand='ResolveBiome',
region=unboundedGeo,
seed=seed,
tileScale=tileScaleToUse,
geometries=True
)
bootstrapSampleExport = ee.batch.Export.table.toAsset(
collection=bootstrapSamples,
description=assetIDString,
assetId='users/'+usernameFolderString+'/'+projectFolder+'/'+bootstrapCollFolder+'/'+assetIDString
);
bootstrapSampleExport.start()
# + jupyter={"source_hidden": true}
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# +
# Convert the categorical image to a multi-band indicator variable style image
# AC This is the WWF Global 200 - https://en.wikipedia.org/wiki/Global_200
resolveBiomesDict = {
1: 'Tropical and subtropical moist broadleaf forests',
2: 'Tropical and subtropical dry broadleaf forests',
3: 'Tropical and subtropical coniferous forests',
4: 'Temperate broadleaf and mixed forests',
5: 'Temperate conifer forests',
6: 'Boreal forests or taiga',
7: 'Tropical and subtropical grasslands, savannas, and shrublands',
8: 'Temperate grasslands, savannas, and shrublands',
9: 'Flooded Grasslands and Savannas',
10: 'Montane grasslands and shrublands',
11: 'Tundra',
12: 'Mediterranean forests, woodlands, and scrub',
13: 'Deserts and xeric shrublands',
14: 'Mangroves'
};
# Instantiate each of the desired function arguments before the scope of the function so as to have only a single argument to the function
classBandToMap = 'Resolve_Biome';
classDictToMap = resolveBiomesDict;
imageNameForBands = 'ResolveBiome';
def categoricalToIndicatorWithoutRefToApply(inputCatImage):
def makeBooleanRasterFromValue(i):
return ee.Image(inputCatImage).select(classBandToMap).eq(ee.Image(ee.Number.parse(i))).rename(ee.String(imageNameForBands).cat(ee.String('_')).cat(ee.Number.parse(i).format('%02d')));
# AC Make a boolean image for each biome and add it to an image list
catClassImageList = ee.Dictionary(classDictToMap).keys().map(makeBooleanRasterFromValue);
def makeRasterFromImageList(i,b):
return ee.Image(b).addBands(i)
#AC Converts the image list to a single image with bands ResolveBiome_01, ResolveBiome_02 etc
unsortedBandImage = ee.Image(catClassImageList.iterate(makeRasterFromImageList,ee.Image()))
return unsortedBandImage.select(unsortedBandImage.bandNames().remove('constant').sort());
# +
# Define a function to make the bootstrap indicator images and save them to a recipient image collection
def makeBootstrapIC(imageToClassify,parentColl,listOfLists):
for seeds in listOfLists:
seedChunkList = seeds
indexString = str(listOfLists.index(seeds)).zfill(2)
fcAssetIDList = []
for seed in seedChunkList:
assetIDString = str('BootstrapSamples_')+str(seed).zfill(4)
# AC load the bootstrap training samples
assetId='users/'+usernameFolderString+'/'+projectFolder+'/'+bootstrapCollFolder+'/'+assetIDString
fcAssetIDList.append(assetId)
def returnFCFromString(string):
return ee.FeatureCollection(string)
# AC creates an image collection with the image to classify with a property for the bootstrap samples
imageCollWithTrainingColls = ee.ImageCollection(ee.List(list(map(returnFCFromString,fcAssetIDList)))
.map(lambda s: imageToClassify.set('TrainingCollection',ee.FeatureCollection(s),
'TrainingString',s)))
def classifyImage(imageWithColl):
trainingCollToUse = ee.FeatureCollection(imageWithColl.get('TrainingCollection'))
trainedModel = ee.Classifier(bestModel).setOutputMode('CLASSIFICATION').train(trainingCollToUse,'ResolveBiome',covariateNames)
classifiedImage = imageWithColl.classify(trainedModel,'PredictedBiome').rename('Resolve_Biome')
return categoricalToIndicatorWithoutRefToApply(classifiedImage.copyProperties(imageWithColl,['TrainingString']))
classifiedImageCollSum = ee.ImageCollection(imageCollWithTrainingColls.map(classifyImage)).sum()
ImageExport = ee.batch.Export.image.toAsset(
image=classifiedImageCollSum,
description=parentColl+'_'+'BootstrapImage_'+indexString,
assetId='users/'+usernameFolderString+'/'+projectFolder+'/'+parentColl+'/BootstrapImage_'+indexString,
crs='EPSG:4326',
crsTransform='[0.008333333333333333,0,-180,0,-0.008333333333333333,90]',
region=exportingGeometry.getInfo()['coordinates'],
maxPixels=int(1e13)
);
ImageExport.start()
# -
# # Bootstrap each of the time periods, sum the collection, then delete the image collection (to make room in the account asset storage)
# usernameFolderString = 'devinrouth_backup'
usernameFolderString = 'acottam'
projectFolder = 'ETH_Biome_Future_Predictions'
# Create bash variables in order to create/check/delete Earth Engine Assets
bashFunction = 'earthengine'
arglist_CreateCollection = ['--no-use_cloud_api','create','collection']
arglist_CreateFolder = ['--no-use_cloud_api','create','folder']
arglist_Detect = ['--no-use_cloud_api','asset','info']
arglist_Delete = ['--no-use_cloud_api','rm','-r']
stringsOfInterest = ['Asset does not exist or is not accessible']
bashCommandList_Detect = [bashFunction]+arglist_Detect
bashCommandList_Delete = [bashFunction]+arglist_Delete
bashCommandList_CreateCollection = [bashFunction]+arglist_CreateCollection
bashCommandList_CreateFolder = [bashFunction]+arglist_CreateFolder
# ## rcp45_2030s
# +
# rcp45_2030s Image Collection creation
rcp45_2030s_Coll = 'rcp45_2030s_Coll';
# Turn the coll string into an assetID and perform the deletion
currentCollToCreate = rcp45_2030s_Coll
assetIDToCreate = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToCreate
print(assetIDToCreate,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateCollection+[assetIDToCreate])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# +
# Make the bootstrap images inside of the collection
# AC params to makeBootstrapIC are: image to classify, parent collection, list of lists
rcp45_2030s_Classified = makeBootstrapIC(rcp45_2030s_Mean_Soil,rcp45_2030s_Coll,seedChunkListToMap);
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# Make a function to load in an image collection (by asset string name) then sum it an export the resultant image
def sumImageCollection(collStringName):
pathOfImageCollToSum = 'users/'+usernameFolderString+'/'+projectFolder+'/'+collStringName;
outputPathOfSummedImage = 'users/'+usernameFolderString+'/'+projectFolder+'/'+collStringName+'_SummedImage';
summedImageToExport = ee.ImageCollection(pathOfImageCollToSum).sum()
summedImageExportTask = ee.batch.Export.image.toAsset(
image=summedImageToExport,
description=collStringName+'_SummedImage',
assetId=outputPathOfSummedImage,
crs='EPSG:4326',
crsTransform='[0.008333333333333333,0,-180,0,-0.008333333333333333,90]',
region=exportingGeometry.getInfo()['coordinates'],
maxPixels=int(1e13)
);
summedImageExportTask.start()
# Sum each of the image collections to get a finalized map
sumImageCollection(rcp45_2030s_Coll)
# !! Break and wait
# Wait while all queued tasks finish
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# rcp45_2030s_Coll
currentCollToDelete = rcp45_2030s_Coll
# Turn the coll string into an assetID and perform the deletion
assetIDToDelete = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToDelete
print(assetIDToDelete,'being deleted')
# Delete the image collection after summing the
subprocess.run(bashCommandList_Delete+[assetIDToDelete])
while not all(x in subprocess.run(bashCommandList_Detect+[assetIDToDelete], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for the asset to delete...')
time.sleep(5)
print('Collection is deleted!')
# -
# ## rcp45_2050s
# +
# rcp45_2050s Image Collection creation
rcp45_2050s_Coll = 'rcp45_2050s_Coll';
# Turn the coll string into an assetID and perform the deletion
currentCollToCreate = rcp45_2050s_Coll
assetIDToCreate = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToCreate
print(assetIDToCreate,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateCollection+[assetIDToCreate])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# +
# Make the bootstrap images inside of the collection
rcp45_2050s_Classified = makeBootstrapIC(rcp45_2050s_Mean_Soil,rcp45_2050s_Coll,seedChunkListToMap);
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# Sum each of the image collections to get a finalized map
sumImageCollection(rcp45_2050s_Coll)
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# rcp45_2050s_Coll
currentCollToDelete = rcp45_2050s_Coll
# Turn the coll string into an assetID and perform the deletion
assetIDToDelete = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToDelete
print(assetIDToDelete,'being deleted')
# Delete the image collection after summing the
subprocess.run(bashCommandList_Delete+[assetIDToDelete])
while not all(x in subprocess.run(bashCommandList_Detect+[assetIDToDelete], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for the asset to delete...')
time.sleep(5)
print('Collection is deleted!')
# -
# ## rcp45_2070s
# +
# rcp45_2070s_Coll Image Collection creation
rcp45_2070s_Coll = 'rcp45_2070s_Coll';
# Turn the coll string into an assetID and perform the deletion
currentCollToCreate = rcp45_2070s_Coll
assetIDToCreate = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToCreate
print(assetIDToCreate,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateCollection+[assetIDToCreate])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# +
# Make the bootstrap images inside of the collection
rcp45_2070s_Classified = makeBootstrapIC(rcp45_2070s_Mean_Soil,rcp45_2070s_Coll,seedChunkListToMap);
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# Sum each of the image collections to get a finalized map
sumImageCollection(rcp45_2070s_Coll)
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# rcp45_2070s_Coll
currentCollToDelete = rcp45_2070s_Coll
# Turn the coll string into an assetID and perform the deletion
assetIDToDelete = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToDelete
print(assetIDToDelete,'being deleted')
# Delete the image collection after summing the
subprocess.run(bashCommandList_Delete+[assetIDToDelete])
while not all(x in subprocess.run(bashCommandList_Detect+[assetIDToDelete], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for the asset to delete...')
time.sleep(5)
print('Collection is deleted!')
# -
# ## rcp45_2080s
# +
# rcp45_2080s_Coll Image Collection creation
rcp45_2080s_Coll = 'rcp45_2080s_Coll';
# Turn the coll string into an assetID and perform the deletion
currentCollToCreate = rcp45_2080s_Coll
assetIDToCreate = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToCreate
print(assetIDToCreate,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateCollection+[assetIDToCreate])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# +
# Make the bootstrap images inside of the collection
rcp45_2080s_Classified = makeBootstrapIC(rcp45_2080s_Mean_Soil,rcp45_2080s_Coll,seedChunkListToMap);
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# Sum each of the image collections to get a finalized map
sumImageCollection(rcp45_2080s_Coll)
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# rcp45_2080s_Coll
currentCollToDelete = rcp45_2080s_Coll
# Turn the coll string into an assetID and perform the deletion
assetIDToDelete = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToDelete
print(assetIDToDelete,'being deleted')
# Delete the image collection after summing the
subprocess.run(bashCommandList_Delete+[assetIDToDelete])
while not all(x in subprocess.run(bashCommandList_Detect+[assetIDToDelete], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for the asset to delete...')
time.sleep(5)
print('Collection is deleted!')
# -
# ## rcp85_2030s
# +
# rcp85_2030s_Coll Image Collection creation
rcp85_2030s_Coll = 'rcp85_2030s_Coll';
# Turn the coll string into an assetID and perform the deletion
currentCollToCreate = rcp85_2030s_Coll
assetIDToCreate = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToCreate
print(assetIDToCreate,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateCollection+[assetIDToCreate])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# +
# Make the bootstrap images inside of the collection
rcp85_2030s_Classified = makeBootstrapIC(rcp85_2030s_Mean_Soil,rcp85_2030s_Coll,seedChunkListToMap);
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# Sum each of the image collections to get a finalized map
sumImageCollection(rcp85_2030s_Coll)
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# rcp85_2030s_Coll
currentCollToDelete = rcp85_2030s_Coll
# Turn the coll string into an assetID and perform the deletion
assetIDToDelete = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToDelete
print(assetIDToDelete,'being deleted')
# Delete the image collection after summing the
subprocess.run(bashCommandList_Delete+[assetIDToDelete])
while not all(x in subprocess.run(bashCommandList_Detect+[assetIDToDelete], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for the asset to delete...')
time.sleep(5)
print('Collection is deleted!')
# -
# ## rcp85_2050s
# +
# rcp85_2050s_Coll Image Collection creation
rcp85_2050s_Coll = 'rcp85_2050s_Coll';
# Turn the coll string into an assetID and perform the deletion
currentCollToCreate = rcp85_2050s_Coll
assetIDToCreate = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToCreate
print(assetIDToCreate,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateCollection+[assetIDToCreate])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# +
# Make the bootstrap images inside of the collection
rcp85_2050s_Classified = makeBootstrapIC(rcp85_2050s_Mean_Soil,rcp85_2050s_Coll,seedChunkListToMap);
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
# Wait while all queued tasks finish
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# Sum each of the image collections to get a finalized map
sumImageCollection(rcp85_2050s_Coll)
# !! Break and wait
# Wait while all queued tasks finish
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# rcp85_2050s_Coll
currentCollToDelete = rcp85_2050s_Coll
# Turn the coll string into an assetID and perform the deletion
assetIDToDelete = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToDelete
print(assetIDToDelete,'being deleted')
# Delete the image collection after summing the
subprocess.run(bashCommandList_Delete+[assetIDToDelete])
while not all(x in subprocess.run(bashCommandList_Detect+[assetIDToDelete], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for the asset to delete...')
time.sleep(5)
print('Collection is deleted!')
# -
# ## rcp85_2070s
# +
# rcp85_2070s_Coll Image Collection creation
rcp85_2070s_Coll = 'rcp85_2070s_Coll';
# Turn the coll string into an assetID and perform the deletion
currentCollToCreate = rcp85_2070s_Coll
assetIDToCreate = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToCreate
print(assetIDToCreate,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateCollection+[assetIDToCreate])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# +
# Turn the coll string into an assetID and perform the deletion
rcp85_2070s_Classified = makeBootstrapIC(rcp85_2070s_Mean_Soil,rcp85_2070s_Coll,seedChunkListToMap);
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# Sum each of the image collections to get a finalized map
sumImageCollection(rcp85_2070s_Coll)
# !! Break and wait
# Wait while all queued tasks finish
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# rcp85_2070s_Coll
currentCollToDelete = rcp85_2070s_Coll
# Turn the coll string into an assetID and perform the deletion
assetIDToDelete = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToDelete
print(assetIDToDelete,'being deleted')
# Delete the image collection after summing the
subprocess.run(bashCommandList_Delete+[assetIDToDelete])
while not all(x in subprocess.run(bashCommandList_Detect+[assetIDToDelete], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for the asset to delete...')
time.sleep(5)
print('Collection is deleted!')
# -
# ## rcp85_2080s
# +
# rcp85_2080s_Coll Image Collection creation
rcp85_2080s_Coll = 'rcp85_2080s_Coll';
# Turn the coll string into an assetID and perform the deletion
currentCollToCreate = rcp85_2080s_Coll
assetIDToCreate = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToCreate
print(assetIDToCreate,'being created...')
# Create the image collection before classifying each of the bootstrap images
subprocess.run(bashCommandList_CreateCollection+[assetIDToCreate])
while any(x in subprocess.run(bashCommandList_Detect+[assetIDToCreate], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for asset to be created...')
time.sleep(normalWaitTime)
print('Asset created!')
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# +
# Create the image collection before classifying each of the bootstrap images
rcp85_2080s_Classified = makeBootstrapIC(rcp85_2080s_Mean_Soil,rcp85_2080s_Coll,seedChunkListToMap);
# Sleep to allow the server time to receive incoming requests
time.sleep(normalWaitTime/2)
# -
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(longWaitTime)
print('Moving on...')
# Sum each of the image collections to get a finalized map
sumImageCollection(rcp85_2080s_Coll)
# !! Break and wait
while any(x in str(ee.batch.Task.list()) for x in ['RUNNING','READY']):
print('You have jobs running! ',datetime.datetime.fromtimestamp(time.time()).strftime('%Y-%m-%d %H:%M:%S'))
time.sleep(normalWaitTime)
print('Moving on...')
# +
# rcp85_2080s_Coll
currentCollToDelete = rcp85_2080s_Coll
# Turn the coll string into an assetID and perform the deletion
assetIDToDelete = 'users/'+usernameFolderString+'/'+projectFolder+'/'+currentCollToDelete
print(assetIDToDelete,'being deleted')
# Delete the image collection after summing the
subprocess.run(bashCommandList_Delete+[assetIDToDelete])
while not all(x in subprocess.run(bashCommandList_Detect+[assetIDToDelete], stdout=subprocess.PIPE).stdout.decode('utf-8') for x in stringsOfInterest):
print('Waiting for the asset to delete...')
time.sleep(5)
print('Collection is deleted!')
| notebooks/crowther/Scripts/Automated_Pipeline_Examples/Biome_Future_Predictions_iMacProVersion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introdução ao Python - <NAME><img src="https://octocat-generator-assets.githubusercontent.com/my-octocat-1626096942740.png" width="324" height="324" align="right">
# ## Link para download: https://github.com/AnabeatrizMacedo241/Python-101
# ## Github: https://github.com/AnabeatrizMacedo241
# ## Linkedin: https://www.linkedin.com/in/ana-beatriz-oliveira-de-macedo-85b05b215/
#
# <img src="https://cdn.jsdelivr.net/gh/devicons/devicon/icons/python/python-original.svg" alt="rails" width='150' height='150' style='max-width: 100%;'></img>
# ## Nessa terceira parte veremos:
# 1. Listas -> Símbolo: []
# 2. Tuplas -> Símbolo: ()
# 3. Dicionários -> Símbolo: {}
# ### Listas
lista_mercado = ['leite', 'manteiga', 'ovo', 'yogurt']
lista_mercado
# Agora vamos falar dos índices dentro de uma Lista.
lista_mercado[0] #Com o index [0], será retornado o primeiro índice da lista.
lista_mercado[-1] #[-1] será retornado o último item da lista.
# Podemos alterar uma lista, porque elas são mutáveis!
lista_mercado[2] = 'suco' #Com o index [2] mudamos o ovo por um suco
lista_mercado
# Também podemos deletar os itens de uma lista pelo seu index usando a função `del`
del lista_mercado[2] #Aqui deletamos o index do suco
lista_mercado
# Listas aninhadas(Listas dentro de listas)
listas = [['leite','manteiga','suco'], [1,2,3]]
listas
listas[0]
listas[1]
#Podemos acessar um índice dentro de um segundo índice usando um segundo index
listas[0][2]
#Assim como strings, podemos concatenar listas
lista_concatenada = listas + lista_mercado
lista_concatenada
# Como checar se há um elemento específico na lista? Usando o operador `in`
'leite'in lista_mercado
'massa'in lista_mercado
# Agora irei mostar alguma funções que podem facilitar ao criar um código...
#A função len mostra a 'largura'e quantidade de elementos em um código.
len(lista_concatenada)
numeros = [1, 32, 23, 7, 24] #A função max retorna o maior elemento da lista
max(numeros)
min(numeros) #min retorna o menor elemento
#A função append adiciona um novo elemento a uma determinada lista
lista_mercado.append('frutas')
lista_mercado
#E drop() retira um elemento
letras = ['a', 'b', 'd', 'b']
letras.pop(0) #Retira o primeiro elemento
letras
#count() quantas vezes certo elemento está presente
letras.count('b')
#reverse() inverte a ordem dos elementos
cores = ['azul', 'rosa', 'vermelho']
cores.reverse()
cores
#sort deixa em ordem
num = [2, 1, 78, 65]
num.sort()
num
#Colocando itens de uma lista anterior em uma nova
for item in num:
numeros.append(item)
numeros
# ### Tuplas
# Diferente de listas, tuplas não podem ser alteradas. Elas são IMUTÁVEIS!
tupla = ('carro', 'moto', 'aviao')
tupla
len(tupla)
tupla.index('carro')
# Slicing[:] pega apenas os elementos que você definie para uma lista, tipla, dicionário, etc...
tupla[1:] #Nesse caso pega tudo do segud=ndo elemento para frente
# Usando a função list() para converter uma tupla para lista
nova_lista = list(tupla)
nova_lista
# ### Dicionários
# +
# Atenção, pois chave e valor podem ser iguais, mas representam coisas diferentes.
# -
dicionário = {"Sapato":36, "Blusa":'P', "Anel":14, "Calça":40}
dicionário
dicionário["Blusa"]
#Adicionando novo elemento
dicionário["Óculos"] = 2.5
dicionário
dicionário.keys()
dicionário.values()
# Dicionário de listas
dict = {'key1':20,'key2':[1,2,3],'key3':['leite','manteiga','suco']}
# Acessando um item da lista, dentro do dicionário
dict['key3'][0].upper()
# Operações com itens da lista, dentro do dicionário
operaçao = dict['key2'][0] - 2
operaçao
#Dicionários aninhados
dict_aninhado = {'key1':{'key2_aninhada':{'key3_aninhada':'Dicionário aninhado'}}}
dict_aninhado['key1']['key2_aninhada']['key3_aninhada']
# ### Slicing
string = 'Listas, Tuplas e Dicionários''
# ### Exercícios
# 1. Crie uma lista com 5 elementos e depois imprima.
# 2. Converta a lista para uma tupla
# 3. Crie um dicionário com 2 keys e 2 values. Depois tente adcionar mais duas keys e dois values.
# 4. Imprimas, apenas, 'Python' da string: 'Estou aprendendo Python' usando o método de slicing.
# 5. Crie uma lista que dentro dela haja: um dicionário, uma tupla e uma segunda lista.
# ### Faça seus próprios exemplos para praticar e bons estudos!
# ## <NAME>
| Aulas/.ipynb_checkpoints/Caderno-03-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/PacktPublishing/Modern-Computer-Vision-with-PyTorch/blob/master/Chapter02/Fetching_values_of_intermediate_layers.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="e1gtGb85daHv"
import torch
x = [[1,2],[3,4],[5,6],[7,8]]
y = [[3],[7],[11],[15]]
# + id="ZaYGlxtQdbd1"
X = torch.tensor(x).float()
Y = torch.tensor(y).float()
# + id="q5DT95H_dcwi"
device = 'cuda' if torch.cuda.is_available() else 'cpu'
X = X.to(device)
Y = Y.to(device)
# + id="rHq7VwgDdeJ-"
import torch.nn as nn
# + id="clpig_05dfYK"
class MyNeuralNet(nn.Module):
def __init__(self):
super().__init__()
self.input_to_hidden_layer = nn.Linear(2,8)
self.hidden_layer_activation = nn.ReLU()
self.hidden_to_output_layer = nn.Linear(8,1)
def forward(self, x):
x = self.input_to_hidden_layer(x)
x = self.hidden_layer_activation(x)
x = self.hidden_to_output_layer(x)
return x
# + id="McmLsQstdnxr"
torch.random.manual_seed(10)
mynet = MyNeuralNet().to(device)
# + id="8cyG-B-AdoBB"
loss_func = nn.MSELoss()
# + colab={"base_uri": "https://localhost:8080/"} id="QRTf8vdKdqmP" outputId="55370c9b-a407-423b-8baa-aaeeae811b22"
_Y = mynet(X)
loss_value = loss_func(_Y,Y)
print(loss_value)
# + id="dTOdsvFydsQK"
from torch.optim import SGD
opt = SGD(mynet.parameters(), lr = 0.001)
# + id="xyHCZwfYduGO"
loss_history = []
for _ in range(50):
opt.zero_grad()
loss_value = loss_func(mynet(X),Y)
loss_value.backward()
opt.step()
loss_history.append(loss_value)
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="DiO6I53udwvY" outputId="bac08c80-f41e-4d8f-93d1-294e47f3bcb3"
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(loss_history)
plt.title('Loss variation over increasing epochs')
plt.xlabel('epochs')
plt.ylabel('loss value')
# + [markdown] id="0-Sn8tA11MKR"
# ### 1. Fetching intermediate values by directly calling the intermediate layer
# + colab={"base_uri": "https://localhost:8080/"} id="jmfHZmKXdyND" outputId="7c3a7be9-46c7-461e-8e37-19e28ac07ff1"
mynet.input_to_hidden_layer(X)
# + [markdown] id="bPkK_o_r2J0y"
# ### 2. Fetching intermediate values by returning them in `nn.Module` definition
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="yarZlkt71Qrw" outputId="e4129b1f-2291-417a-b025-d47d73e23beb"
torch.random.manual_seed(10)
class MyNeuralNet(nn.Module):
def __init__(self):
super().__init__()
self.input_to_hidden_layer = nn.Linear(2,8)
self.hidden_layer_activation = nn.ReLU()
self.hidden_to_output_layer = nn.Linear(8,1)
def forward(self, x):
hidden1 = self.input_to_hidden_layer(x)
hidden2 = self.hidden_layer_activation(hidden1)
x = self.hidden_to_output_layer(hidden2)
return x, hidden1
mynet = MyNeuralNet().to(device)
loss_func = nn.MSELoss()
_Y, _Y_hidden = mynet(X)
loss_value = loss_func(_Y,Y)
opt = SGD(mynet.parameters(), lr = 0.001)
loss_history = []
for _ in range(50):
opt.zero_grad()
loss_value = loss_func(mynet(X)[0],Y)
loss_value.backward()
opt.step()
loss_history.append(loss_value)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(loss_history)
plt.title('Loss variation over increasing epochs')
plt.xlabel('epochs')
plt.ylabel('loss value')
# + colab={"base_uri": "https://localhost:8080/"} id="sNQxAxOM1oaX" outputId="f2219f64-dd84-4ba7-f2fd-f36a96e348c1"
mynet(X)[1]
# + id="vXZvZI-U1pcK"
| Chapter02/Fetching_values_of_intermediate_layers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import os
import us
import numpy as np
import pandas as pd
# pd.set_option('display.max_rows', None)
import matplotlib.pyplot as plt
weights = pd.read_csv('../data/ref/parsed_usa_intnt_users.csv')
# +
class Transform:
# Requirements:
# import os
# import pickle
# import numpy as np
# import pandas as pd
# import matplotlib.pyplot as plt
def __init__(self, keyword):
self.keyword = keyword
def agg(self, df, kind, **kwargs):
if (kind=='arithmetic mean'):
return df.mean(axis=0)
if kind in ['weighted mean', 'weighted']:
temp = df.merge(kwargs['weights'], how='left', on=kwargs['key'])
temp['intnt_pop_norm'] = temp['intnt_pop'].apply(lambda x: x / temp['intnt_pop'].sum())
for col in df.convert_dtypes().select_dtypes(include=['int', 'float']).columns:
temp[col] = temp[col] * temp['intnt_pop_norm']
temp = temp.drop(['intnt_pop_norm', 'intnt_pop'], axis=1)
if kind == 'weighted':
weighted = temp.melt(id_vars=[self.geoName])
weighted['combine'] = weighted['variable'].str.replace(' ', '_') \
.str.lower() + '_' + weighted[self.geoName]
return weighted[['combine', 'value']].round(3).set_index('combine')
if kind == 'weighted mean':
if self.grouping == 'none':
return temp.mean(numeric_only=True)
return temp.groupby('region').mean()
def cleanColumns(self, df, grouping):
df.columns = df.columns.str.replace('\"', '').str.replace(' ', '_')
df['date'] = pd.to_datetime(df['date'])
df = df.sort_values(by='date').reset_index(drop=True)
# Reordering columns to have 'date' as the first column
col_order = ['date'] + (df.columns.tolist()[:-1])
df = df[col_order]
if self.geoName == 'state':
return df
if self.geoName == 'country_code':
new_cols = ['date']
for col in df.columns[1:]:
if grouping == 'none':
brand, denom = col.lower().split('_')
new_col = denom + '_' + brand
else:
region, brand, denom = col.lower().split('_')
new_col = denom + '_' + brand + '_' + region
new_cols.append(new_col)
new_df = df.rename(columns=dict(zip(df.columns,new_cols)))
return new_df
def aggFromPickles(self, inputPath, geoName, aggFunc, grouping=pd.DataFrame({'None': [0,0]}), **kwargs):
# grouping(pd.DataFrame()): dataframe denoting regions to be grouped by, defaults to 'None'
# aggFunc(str): aggregation method of choice (e.g. 'arithmetic mean')
df = pd.DataFrame()
for root, _, files in os.walk(inputPath):
for file in sorted(files):
if file.endswith(".pkl"):
year = file.split('-')[0]
month = file.split('-')[1]
with open(os.path.join(root, file), 'rb') as f:
df_temp = pickle.load(f)
if geoName == 'state':
self.geoName = 'state'
df_temp = df_temp.rename(columns={'geoName': geoName})
conv = us.states.mapping('name', 'abbr')
df_temp[geoName] = df_temp[geoName].map(conv)
else:
self.geoName = 'country_code'
df_temp = df_temp.rename(columns={'geoName': geoName})
df_temp = df_temp.loc[df_temp[geoName]!='U.S. Outlying Islands', :]
df_temp[geoName] = cc.convert(names=df_temp[geoName].tolist(),
to='ISO3',
not_found='not there')
df_temp = df_temp.loc[df_temp[geoName]!='not there', :]
if grouping.equals(pd.DataFrame({'None': [0,0]})):
self.grouping = 'none'
if aggFunc in ['weighted', 'weighted mean']:
df_temp = pd.DataFrame(self.agg(df_temp,
aggFunc,
weights=kwargs['weights'],
key=kwargs['key'])).T
else:
df_temp = pd.DataFrame(self.agg(df_temp.iloc[:, 1:], aggFunc)).T
else:
df_temp_merged = df_temp.merge(grouping, on=geoName)
if aggFunc == 'weighted mean':
df_temp = self.agg(df_temp_merged,
aggFunc,
weights=kwargs['weights'],
key=kwargs['key'])
df_temp = df_temp.reset_index() \
.melt(id_vars='region')
else:
df_temp = df_temp_merged.groupby(['region']) \
.apply(self.agg, aggFunc) \
.reset_index() \
.melt(id_vars='region')
df_temp['Group'] = df_temp['region'] + '_' + df_temp['variable']
df_temp = df_temp.rename(columns={'value': aggFunc})
df_temp = df_temp[['Group', aggFunc]].set_index('Group').T
df_temp['date'] = year + '-' + month
df = df.append(df_temp)
df = self.cleanColumns(df, self.grouping)
self.df = df
def lineplot(self):
fig, ax = plt.subplots(figsize=(10,7))
for i in range(1, len(self.df.columns)):
ax.plot(self.df.iloc[:,0], self.df.iloc[:, i], label=self.df.columns[i])
# plt.xticks(np.arange(0, len(self.df), 12),
# list(map(str, self.df['date'].dt.year.unique())),
# rotation=60)
ax.set_ylim(bottom=0)
ax.set_xlabel('Year')
ax.set_ylabel('Mean')
ax.set_title('Keyword: \'{}\''.format(self.keyword.capitalize()))
ax.legend()
def toPickle(self, outputPath):
fileName = 'global-arimean-{}'.format(self.keyword)
pathName = '{}/{}.pkl'.format(outputPath, fileName)
with open(pathName, 'wb') as f:
pickle.dump(self.df, f)
def toCSV(self, outputPath, aggFunc, grouping):
fileName = '{}-{}-{}'.format(self.keyword, aggFunc, grouping)
pathName = '{}/{}.csv'.format(outputPath, fileName)
self.df.to_csv(pathName, index=False)
class Transform_keyword(Transform):
# Subclass of class Transform
def __init__(self, keyword):
super().__init__(keyword)
def path(self):
return '../data/raw/{}'.format(self.keyword)
def aggFromPickles(self):
super().aggFromPickles(self.path())
# -
t = Transform('cuisine')
t.aggFromPickles(inputPath='../data/raw',
geoName='state',
aggFunc='weighted',
weights= weights,
key='state')
t.df
t.toCSV('../data/agg', 'wtd', 'usa')
| src/processing/transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import pickle
from tqdm.notebook import tqdm as log_progress
from ipywidgets import widgets # for buttons and dropdown menus
from IPython.display import display, clear_output # for buttons and dropdown menus
import seaborn as sns
import matplotlib.pyplot as plt
from scalecast.Forecaster import Forecaster
data = pd.read_csv('avocado.csv',parse_dates=['Date'])
data = data.sort_values(['region','type','Date'])
sns.set(rc={'figure.figsize':(12,8)})
def results_vis(f_dict,plot_type='forecast',order_by='LevelTestSetMAPE',level=True):
""" visualize the forecast results
leverages Jupyter widgets
level arg only works for plot_type = 'forecast' since plot_set_set only plots at the level of the forecast
"""
def display_user_selections(ts_selection,mo_selection):
""" displays graphs with seaborn based on what user selects from dropdown menus
"""
sns.set(rc={'figure.figsize':(18,10)})
selected_data = f_dict[ts_selection]
if plot_type == 'forecast':
print(ts_selection)
selected_data.plot(models=f'top_{mo_selection}',order_by=order_by,level=level,
print_attr=['TestSetRMSE','TestSetR2','LevelTestSetRMSE','TestSetMAPE','LevelTestSetR2','LevelTestSetMAPE','Scaler','HyperParams','Xvars','models','Integration'])
elif plot_type == 'test':
print(ts_selection)
selected_data.plot_test_set(models=f'top_{mo_selection}',order_by=order_by,include_train=52)
def on_button_clicked(b):
""" passes the user options to the display_user_selections function after the button is pressed
"""
mo_selection = mo_dd.value
ts_selection = ts_dropdown.value
with output:
clear_output()
display_user_selections(ts_selection,mo_selection)
all_models = models + ('weighted1','weighted2','avg')
ts_dropdown = widgets.Dropdown(options=f_dict.keys(), description = 'Time Series:')
mo_dd = widgets.Dropdown(options=range(1,len(all_models)+1), description = 'No. Models')
# never changes
button = widgets.Button(description="Select Time Series")
output = widgets.Output()
display(ts_dropdown,mo_dd)
display(button, output)
button.on_click(on_button_clicked)
avc = {}
for reg in data.region.unique():
for typ in data.type.unique():
data_slice = data.loc[(data['region'] == reg) & (data['type'] == typ)]
load_dates = pd.date_range(start=data_slice['Date'].min(),end=data_slice['Date'].max(),freq='W')
data_load = pd.DataFrame({'Date':load_dates})
data_load['Vol'] = data_load.merge(data_slice,how='left',on='Date')['Total Volume'].values
data_load.fillna(0,inplace=True)
f = Forecaster(y=data_load['Vol'],current_dates=data_load['Date'],type=typ,region=reg)
avc[f"{reg}-{typ}"] = f
# summaries
print(f'number of series to forecast: {len(avc)}')
print(f'number of actual weeks to forecast with: {len(f.y)}')
f.plot_acf(diffy=True,lags=26)
plt.title(f'{f.type} {f.region} ACF Plot')
plt.show()
f.plot_pacf(diffy=True,lags=26)
plt.title(f'{f.type} {f.region} PACF Plot')
plt.show()
print(f'{f.type} {f.region} Seasonal Decomposition')
f.seasonal_decompose(diffy=True).plot()
plt.show()
# + tags=[]
models = ('mlr','knn','xgboost','gbt','elasticnet','mlp','prophet')
#avc = pickle.load(open('avocado_results.pckl','rb'))
for k, f in log_progress(avc.items()):
f.generate_future_dates(52)
f.set_test_length(26)
f.set_validation_length(13)
f.add_ar_terms(3)
f.add_AR_terms((1,26))
if not f.adf_test():
f.diff()
f.add_seasonal_regressors('week','month','quarter',sincos=True)
f.add_seasonal_regressors('year')
f.add_time_trend()
for m in models:
f.set_estimator(m)
f.tune() # by default, will pull the grid with the same name as the estimator (mlr will pull the mlr grid, etc.)
f.auto_forecast()
# combine models and run manually specified models of other varieties
f.set_estimator('combo')
f.manual_forecast(how='weighted',models=models,determine_best_by='ValidationMetricValue',call_me='weighted') # weighted average of all models based on determine_best_by
f.manual_forecast(how='simple',models='top_5',determine_best_by='ValidationMetricValue',call_me='avg') # a simple average of the top 5 best performing models based on determine_best_by
# +
forecast_info = pd.DataFrame()
for k, f in avc.items():
df = f.export(dfs='model_summaries',determine_best_by='LevelTestSetMAPE')
df['Name'] = k
df['Region'] = f.region
df['Type'] = f.type
forecast_info = pd.concat([forecast_info,df],ignore_index=True)
forecast_info.to_csv('avocado_model_summaries.csv',index=False)
# -
forecast_info['LevelTestSetMAPE'].mean()
forecast_info.loc[forecast_info['best_model'] == True, 'LevelTestSetMAPE'].mean()
forecast_info.groupby('ModelNickname')['best_model'].sum().plot.bar()
plt.show()
results_vis(avc,'test','TestSetRMSE')
results_vis(avc,'forecast','TestSetRMSE')
with open('avocado_results.pckl','wb') as f:
pickle.dump(avc,f)
| .ipynb_checkpoints/avocados-checkpoint.ipynb |