code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://blog.csdn.net/weixin_41712499/article/details/82875574
# # Normalization(正则化、规范化) and Scaler(归一化)
# normalizer主要用于样本;
#
# scaler 主要用于特征
# 正则化的过程是将每个样本缩放到单位范数(每个样本的范数为1),如果后面要使用如二次型(点积)或者其它核方法计算两个样本之间的相似性这个方法会很有用。
#
# 对于两个TF-IDF向量的l2-norm进行点积,就可以得到这两个向量的余弦相似性。
#
# Normalization主要思想是对每个样本计算其p-范数,然后对该样本中每个元素除以该范数,这样处理的结果是使得每个处理后样本的p-范数(l1-norm,l2-norm)等于1。
# ## 补充
# 
import numpy as np
x = np.arange(-4, 5).reshape((3, 3))
x
# ## Normalization
from sklearn.preprocessing import Normalizer
normalizer = Normalizer(norm='l2')
normalizer.fit(x)
normalizer.transform(x)
from sklearn.preprocessing import normalize
# ### 最大值置为1(除以最大值)
normalize(x, 'max')
normalize(x, 'max', axis=0)
# ### 向量的1范数:元素绝对值和为1
normalize(x, 'l1', axis=0)
# ### 向量的2范数:元素平方和为1
normalize(x, 'l2', axis=0)
# ## Scaler(归一化)
# ### StandardScaler:Z-Score标准化: 均值为0,方差为1
# Z-Score通过(x-μ)/σ将两组或多组数据转化为无单位的Z-Score分值,使得数据标准统一化,提高了数据可比性,削弱了数据解释性。
# 公式为:(X-mean)/std 计算时对每个属性/每列分别进行。
#
# 将数据按期属性(按列进行)减去其均值,并处以其方差。得到的结果是,对于每个属性/每列来说所有数据都聚集在0附近,方差为1。
#
# 实现时,有两种不同的方式:
# 使用sklearn.preprocessing.StandardScaler类,使用该类的好处在于可以保存训练集中的参数(均值、方差)直接使用其对象转换测试集数据。
x
# +
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(x)
# -
scaler.mean_
scaler.scale_
# 方差
scaler.var_
scaler.transform(x)
# 使用sklearn.preprocessing.scale()函数,可以直接将给定数据进行标准化。
from sklearn.preprocessing import scale
scale(x)
# ### MinMaxScaler
# 除了上述介绍的方法之外,另一种常用的方法是将属性缩放到一个指定的最大和最小值(通常是1-0)之间,这可以通过preprocessing.MinMaxScaler类实现。
#
# 使用这种方法的目的包括:
#
# 1、对于方差非常小的属性可以增强其稳定性。
#
# 2、维持稀疏矩阵中为0的条目。
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
scaler.fit(x)
# 原始的每个特征值范围
scaler.data_range_
scaler.transform(x)
# # 处理时间数据
# ## 提取每个时间字段的月,日,weekday
for i in date_cols:
df2[f'{i}_day'] = df2[i].dt.day
df2[f'{i}_month'] = df2[i].dt.month
df2[f'{i}_weekday'] = df2[i].dt.weekday
# ## 日期间隔
import itertools
for s, e in itertools.combinations(date_cols, 2):
df2[f'{e}-{s}'] = (df2[e] - df2[s]).dt.days
num_cols.append(f'{e}-{s}')
# ## 删除时间字段
df2.drop(columns=date_cols, inplace=True)
| feature_engineering/data_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import sys
sys.path.append(".")
import ngm
# ## Read in metabric data from deeptype paper
# +
import scipy.io
import numpy as np
import pandas as pd
mat = scipy.io.loadmat('BRCA1View20000.mat')
data = mat['data']
mat = {k:v for k, v in mat.items() if k[0] != '_'}
print(len(mat['targets']))
#mat = {k: pd.Series(v[0]) for k, v in mat.items() if len(v) == 0}
mat = {k: pd.Series(v.flatten()) for k, v in mat.items()}
targets = pd.DataFrame.from_dict(mat).dropna()[['id', 'targets']].astype(int)
# -
data.shape
X = data.T
X.shape
# ## Read in our adjacency matrix
import pandas as pd
import io
import requests
url="https://metabric.s3-us-west-1.amazonaws.com/cat2vec/cat2vec_adj.csv"
s=requests.get(url).content
c=pd.read_csv(io.StringIO(s.decode('utf-8')))
c['id'] = c['Patient ID'].apply(lambda x: int(x.split("-")[1]))
c_merged = c.merge(targets, on='id')
c_merged = c_merged.set_index('Patient ID')
c_merged = c_merged[c_merged.index]
c_merged.shape
c_merged
c_merged.index = list([int(c.split("-")[1]) for c in c_merged.index])
c_merged.columns = c_merged.index
c_merged
# ## Fix things so they aren't weird
mat.keys()
genes = [g[0] for g in mat['gene']]
pd.Series(genes)
X_df = pd.DataFrame(X)
X_df.columns = genes
X_df = X_df.set_index(mat['id'])
X_df
y = targets.set_index('id')['targets'].astype(int) # Richa: I wanted to change this to string but it broke pytroch
y = y - y.min() # make sure 0 is the first class. Richa: let's fix this as well
y.value_counts()
A=c_merged
# ## Put data into the pytorch format
import torch
from sklearn.model_selection import train_test_split
from torch.utils.data import DataLoader
c_merged
X_df.loc[A.index]
y.loc[A.index]
batch_size = 5 # The size of input data took for one iteration
X_train, X_test, y_train, y_test = train_test_split(X_df.loc[A.index], y.loc[A.index], test_size=0.1, random_state=42)
X_train = torch.tensor(X_train.values).float()
y_train = torch.tensor(y_train.values).float()
X_test = torch.tensor(X_test.values).float()
y_test = torch.tensor(y_test.values).float()
dataset = ngm.base.MyDataset(X_train, y_train)
dataloader = DataLoader(dataset, batch_size=batch_size)
testset = ngm.base.MyDataset(X_test, y_test)
testloader = DataLoader(testset, batch_size=batch_size, shuffle=True)
# ## Now construct the model
input_size = X_df.shape[1]
hidden_size = 50 # The number of nodes at the hidden layer
num_classes = len(y.unique()) # The number of output classes. In this case, from 1 to 39
model = ngm.base.Net(input_size, hidden_size, num_classes)
# ## Train the model
learning_rate = 0.001 # The speed of convergence
num_epochs=30
ngm.base.train(model,A,dataloader,dataset,learning_rate,num_epochs)
# ## Evaluation
from sklearn.metrics import classification_report
ypred,ylabels = ngm.base.prepare_for_evaluation(model,testloader)
print(classification_report(ylabels, ypred))
pd.Series(ypred).value_counts()
| metabric.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Anaconda3]
# language: python
# name: conda-env-Anaconda3-py
# ---
import numpy as np
a=np.array([[1,0,0,1],[0,0,0,0]])
print(a)
b=np.array([1,1,0,1])
np.matmul(a,b)
H=np.array([[0,0,0,1,1,1,1,0,1,0,1,1,0,0,1],[0,0,1,0,0,0,1,1,1,1,0,1,0,1,1],[0,1,0,0,0,1,1,1,1,0,1,0,1,1,0],[1,0,0,0,1,1,1,1,0,1,0,1,1,0,0],[0,1,1,0,0,0,1,1,0,0,0,1,1,0,0],[0,0,1,1,0,0,0,1,1,0,0,0,1,1,0],[0,0,1,0,1,0,0,1,0,1,0,0,1,0,1],[1,0,1,1,1,1,0,1,1,1,1,0,1,1,1]])
x=np.array([0,0,1,1,0,0,1,0,1,0,1,1,0,0,1])
# +
y=list(np.matmul(H,x))
[x%2 for x in y]
# -
| double-error correcting code on GF(16).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/danio2010/ON2022/blob/main/Praca_z_plikami.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="wT3FV-zthalh"
# + [markdown] id="dependent-norman"
# ## Praca z plikami na lokalnej maszynie
#
# [Własne pliki w Colab](https://neptune.ai/blog/google-colab-dealing-with-files)
#
# + colab={"base_uri": "https://localhost:8080/"} id="zWyBD_GQwSvh" outputId="74966497-a5e5-4eab-f62f-e2ec38b09128"
#czytanie komendami bash
# !head -n 5 sample_data/README.md
# + colab={"base_uri": "https://localhost:8080/"} id="velvet-champion" outputId="02a71851-3342-4c30-b188-34dae56832b9"
with open('sample_data/README.md', 'r') as moj_plik:
# tu można robić cokolwiek z plikiem
# 'r' to tryb odczytu - domyślny
# 'w' to tryb do zapisu
# 'a' - tryb dopisywania
print(moj_plik.readline())
print(moj_plik.readline())
print(moj_plik.readline())
print(moj_plik.readline())
# + id="f6g1dNYLpGbo"
# + colab={"base_uri": "https://localhost:8080/"} id="proud-compression" outputId="8ff94bcd-b8f5-43d4-968b-3c4e120cd23a"
moj_plik2=open('sample_data/README.md', 'r')
wiersze=moj_plik2.readlines()
for line in wiersze:
print(line)
moj_plik2.close()
# + [markdown] id="1o5deLu2t3Ug"
# #### Zapisywanie do plików
# + id="jSvmpsewt2HJ"
with open('sample_data/nowy_plik.txt', 'w') as plik:
plik.write('Hej!')
plik.write('Jak się masz?')
# + colab={"base_uri": "https://localhost:8080/"} id="pOtNC9PoofS5" outputId="728205c4-de0a-41f8-8d0f-fe2ab68c270a"
# !head -n 5 sample_date/mnist_train_small.csv
# + [markdown] id="h_C9PYuYsDXG"
# ### Zadanie
# Wczytaj plik ```mnist_train_small.csv```. Wypisz jego pięć pierwszych wierszy stawiając na początku numer wiersza, a na końcu wykrzyknik (po każdym wierszu).
#
# Zapisz też to wszystko w nowym pliku txt.
# + colab={"base_uri": "https://localhost:8080/"} id="UwbQ_dmmsfqB" outputId="535bcadb-e1d6-4c7f-cc65-242c663382c9"
wiersze=[]
with open('sample_data/mnist_train_small.csv') as plik:
for i in range(5):
nowy_wiersz=f'{i}: {plik.readline()[:-2]}!'
wiersze.append(nowy_wiersz)
print(nowy_wiersz)
with open('sample_data/nowe_wiersze.txt', 'w') as plik:
for el in wiersze:
plik.write(el+'\n')
# + colab={"base_uri": "https://localhost:8080/"} id="y9lv9AwZvIml" outputId="3b804ceb-bbd4-4f41-f6e6-4061a2efa6b2"
# + [markdown] id="duplicate-divide"
# ## Praca z plikami z sieci
# [Zewnętrzne dane w Colab](https://towardsdatascience.com/7-ways-to-load-external-data-into-google-colab-7ba73e7d5fc7)
# + [markdown] id="baking-dryer"
# ### Wczytywanie plików do pd.DataFrame (pandas)
# + id="cross-tuesday"
import pandas as pd
url ='https://raw.githubusercontent.com/danio2010/ON2022/main/statystyki.csv'
plik=pd.read_csv(url)
# + id="unavailable-chair" outputId="91897e37-f480-4b75-fce1-e5439f22a74d"
type(plik)
# + [markdown] id="computational-westminster"
# ### Czytanie surowych stron internetowych/plików pod url
# + id="received-difference"
import requests
czytaj=requests.get('https://www-users.mat.umk.pl/~danio/')
# + id="organic-terrace" outputId="6937026d-04d0-49e4-e0fd-c04725eafb01" colab={"base_uri": "https://localhost:8080/"}
type(czytaj)
# + id="tested-grade" outputId="17d50c1b-5517-4f82-8f3b-9b3c7b59e2bd"
czytaj.text
# + id="interstate-pencil"
url='https://raw.githubusercontent.com/danio2010/ON2022/main/zad1.6.txt'
czytaj_tekst=requests.get(url)
# + id="hundred-traveler" outputId="7d3d6cb2-5694-4fda-b1fa-775fe097615c" colab={"base_uri": "https://localhost:8080/", "height": 35}
czytaj_tekst.text
# + id="accessible-today" outputId="8c12aa46-ff58-4a09-d8be-01793078b04b"
for line in czytaj_tekst:
decoded_line = line.decode("utf-8")
print(decoded_line)
# + id="identical-learning"
import urllib.request
file=urllib.request.urlopen(url)
# + id="parliamentary-complaint" outputId="7f184f21-1c3c-4ea1-fd41-80c0e580d767"
file.readline()
# + id="0zqOR-75rwQD"
# + [markdown] id="PXSbVZjMrwl2"
# ## Dekodery
# CSV
# + id="FnL54oNOiVwS"
import numpy as np
import pandas as pd
import json
# + id="H2yWC5OqrxmN"
import csv
with open("sample_data/california_housing_test.csv") as plik:
czytaj_csv = csv.reader(plik)
rows2=list(czytaj_csv)
# + colab={"base_uri": "https://localhost:8080/"} id="f2fCn2BLflN8" outputId="2af89395-0e4a-4e41-d075-8619b7ce2e5e"
rows2[0]
# + id="9ww00wUufmKC"
# zapisywanie
data = np.random.randn(100, 3)
np.savetxt('losowe_dane.csv',data, delimiter=",", header="x, y, z", comments="# Losowe współrzędne x, y, z\n")
# + id="szOO-qJ_f12T"
data_load = np.loadtxt("losowe_dane.csv", skiprows=2, delimiter=",")
# + colab={"base_uri": "https://localhost:8080/"} id="bhFe1CFmh9OG" outputId="eb2b04a3-a07b-4a24-e119-baf282373c72"
(data == data_load).all()
# + id="ZccoIOnwh5QH"
tabelka=pd.read_csv('losowe_dane.csv',skiprows=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="GfpG4iqJi2B5" outputId="f819c0dd-5226-4bde-91f1-aaa30ec39bb2"
tabelka
# + colab={"base_uri": "https://localhost:8080/"} id="ZSd2q3Pbi3F7" outputId="19f3ca75-0032-48fa-cb68-4e6908aa04af"
tabelka.info()
# + id="gaMG-ZM-jCYy"
#zapisywanie
tabelka.to_csv('losowe_dane2.csv')
# + [markdown] id="KeKGkIxZj9eD"
# #### JSON
# + id="S6Lwurevj_4V"
data = {"one": [1],
"two": [1, 2],
"three": [1, 2, 3]}
data_json = json.dumps(data, indent=True) #dumps tworzy string z danych słownikowych, a to już możemy normalnie zapisywać do pliku
# + colab={"base_uri": "https://localhost:8080/"} id="QMwqyacjstHw" outputId="a228879e-8423-4958-8834-2da0b87d600a"
type(data_json)
# + id="kDJBJ4A-kTX6"
with open('sample_data/anscombe.json') as plik:
dane_z_pliku=json.load(plik)
# + id="6V9JailEkfZb"
with open("dane_json2.json", "w") as f:
json.dump(dane_z_pliku[:5], f) #UWAGA: dump, a nie dumps!
# + [markdown] id="pNaonIGPoBXG"
# #### XLSX i JSON w Pandas
# ```pd.read_excel```
#
# ```pd.read_json```
#
# ```df.to_json```
#
# ```df.to_excel```
# + id="vjnfUiT3lA4F"
jsonDF=pd.read_json('sample_data/anscombe.json')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="-XlF9TyDo5eT" outputId="fe0824a9-aacf-4007-d381-9bc94277328b"
jsonDF
# + id="-G_tQKhKo6LF"
jsonDF.to_excel('dane_z_json.xlsx',index=False)
# + id="-9PVb9fgpsUK"
#przypomnienie - ładowanie pliku bezpośrednio z url
plik=pd.read_json('https://github.com/danio2010/ON2022/raw/main/tokyo-metro.json')
# + id="d64CQ6qApwOY" colab={"base_uri": "https://localhost:8080/", "height": 291} outputId="47762dba-9ba9-4c4d-9547-9e6e7ca41918"
plik
# + id="m1C-ilEEtVsC"
| Praca_z_plikami.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clusterization methods
# <h3> Plan </h3>
# +
import os
import time
import warnings
import datetime
import numpy as np
import pandas as pd
import random
from sklearn.metrics import accuracy_score as acc
from sklearn.model_selection import cross_val_score
import pylab
from matplotlib import pyplot as plt
from matplotlib.colors import ListedColormap
from IPython.display import Image, SVG
# %matplotlib inline
# -
#
# ** Questions **
# * What is clusterization?
# * What main steps are in K-Means?
# ## k-means demo
# <a href='https://www.naftaliharris.com/blog/visualizing-k-means-clustering/'> k-means </a>
#
# ## DBSCAN demo
# <a href='https://www.naftaliharris.com/blog/visualizing-dbscan-clustering/'> DBSCAN </a>
#
# ---------
# <h1 align="center"> K-Means </h1>
# ** k-means steps: **
# - 1. update clusters (reassign objects to clusters):
# ## $$ y_i := \arg\min\limits_{y\in Y} \rho (x_i ; \mu_y),~~i = 1,\dots,\ell;$$
# - 2: update clusters weights
# ## $$ \mu_{yj} := \frac{\sum_{i=1}^\ell [y_i = y]\cdot f_j(x_i)} {\sum_{i=1}^\ell[y_i = y]}$$
#
#
class KMeans():
def __init__(self, K, X=None, N=0):
'''
K - number of clusters
X - dataset (if X is None then X is generated from gauss distribution)
N - a number of samples to generate if X is None
'''
self.K = K
if X is None:
if N == 0:
raise Exception("If no data is provided, \
a parameter N (number of points) is needed")
else:
self.N = N
self.X = self._init_board_gauss(N, K)
else:
self.X = X
self.N = len(X)
# initialization
self.mu = None # a list of centers of clusters
self.clusters = None # labels of samples
self.method = None # method for sampling initial centers of clusters
def _init_board_gauss(self, N, k):
'''
N - a number of samples to generate
k - a number of clusters
'''
n = float(N)/k
X = []
for i in range(k):
c = (random.uniform(-1,1), random.uniform(-1,1))
s = random.uniform(0.05,0.15)
x = []
while len(x) < n:
a,b = np.array([np.random.normal(c[0],s),np.random.normal(c[1],s)])
# Continue drawing points from the distribution in the range [-1,1]
if abs(a) and abs(b)<1:
x.append([a,b])
X.extend(x)
X = np.array(X)[:N]
return X
def plot_board(self, fig_size=(10,7) ):
'''
'''
X = self.X
fig = plt.figure(figsize = fig_size)
plt.xlim(-1,1)
plt.ylim(-1,1)
if self.mu and self.clusters:
mu = self.mu
clus = self.clusters
# print(clus)
K = self.K
for m, clu in clus.items():
cmap = plt.cm.get_cmap("Spectral")
cs = cmap(1.*m/self.K)
plt.plot(mu[m][0], mu[m][1], 'o', marker='*', \
markersize=20, color=cs)
# print(zip(clus[m]))
plt.plot([x[0] for x in clus[m]], [x[1] for x in clus[m]], '.', \
markersize=8, color=cs, alpha=0.5)
else:
plt.plot(X[:,0], X[:,1], '.', alpha=0.5)
if self.method == '++':
tit = 'K-means++'
else:
tit = 'K-means with random initialization'
pars = 'N=%s, K=%s' % (str(self.N), str(self.K))
plt.title('\n'.join([pars, tit]), fontsize=16)
plt.savefig('kpp_N%s_K%s.png' % (str(self.N), str(self.K)), \
bbox_inches='tight', dpi=200)
def _reevaluate_centers(self):
'''
Maximization step in Kmeans
'''
clusters = self.clusters
newmu = []
keys = sorted(self.clusters.keys())
for k in keys:
newmu.append(np.mean(clusters[k], axis = 0))
self.mu = newmu
def _cluster_points(self):
'''
expectation step in Kmeans
'''
mu = self.mu
clusters = {}
for x in self.X:
bestmukey = # YOUR CODE!!!For x find index of the nearest cluster for it (use np.linalg.norm)!!!
try:
clusters[bestmukey].append(# YOUR CODE!!!Add x to the find cluster's list of elements!!!)
except KeyError:
clusters[bestmukey] = [x]
if len(clusters) < self.K:
for k in range(len(self.K)):
if k not in clusters.keys():
clusters[k] = mu[k] # Not update cluster
self.clusters = clusters
def find_centers(self, method='random'):
self.method = method
X = self.X
K = self.K
#print(X)
self.oldmu = random.sample(list(X), K)
if method != '++':
# Initialize to K random centers
self.mu = random.sample(list(X), K)
while not self._has_converged():
self.oldmu = self.mu # remember previous cluster centers
# Assign all points in X to clusters
self._cluster_points()
# Reevaluate centers
self._reevaluate_centers()
def _has_converged(self):
'''
condition of convergence of cluster points
'''
K = len(self.oldmu)
return(# YOUR CODE!!!Check that our optimization has converged (use self.oldmu)!!!
and len(set([tuple(a) for a in self.mu])) == K)
kmeans = KMeans(K=3, N=200)
kmeans.find_centers()
kmeans.plot_board(fig_size=(10,7))
kmeans.find_centers()
kmeans.plot_board(fig_size=(15,4))
# ### Improve to Kmeans++
class KPlusPlus(KMeans):
def _dist_from_centers(self):
cent = self.mu
X = self.X
D2 = np.array([min([np.linalg.norm(x-c)**2 for c in cent]) for x in X])
self.D2 = D2
def _choose_next_center(self):
self.probs = self.D2/self.D2.sum()
self.cumprobs = self.probs.cumsum()
r = (random.random()+1.0)/2
ind = np.where(self.cumprobs >= r)[0][0]
return(self.X[ind])
def init_centers(self):
self.mu = random.sample(list(self.X), 1)
while len(self.mu) < self.K:
self._dist_from_centers()
self.mu.append(self._choose_next_center())
def plot_init_centers(self, fig_size = (10,7)):
X = self.X
fig = plt.figure(figsize=(10,5))
plt.xlim(-1,1)
plt.ylim(-1,1)
plt.plot(X[:,0], X[:,1], '.', alpha=0.5)
plt.plot([x[0] for x in self.mu], [x[1] for x in self.mu], 'ro')
plt.savefig('kpp_init_N%s_K%s.png' % (str(self.N),str(self.K)), \
bbox_inches='tight', dpi=200)
kplusplus = KPlusPlus(K=5, N=400)
kplusplus.plot_board(fig_size = (15,5))
# Random initialization
kplusplus.find_centers(method='random')
kplusplus.plot_board(fig_size = (15,3))
# k-means++ initialization
kplusplus.init_centers()
kplusplus.plot_init_centers(fig_size=(10,3))
kplusplus.find_centers(method='++')
kplusplus.plot_board(fig_size = (15,3))
# -------
# <h1 align="center">Text clusterization</h1>
# ## Sample
from sklearn.datasets import fetch_20newsgroups
train_all = fetch_20newsgroups(subset='train')
print (train_all.target_names)
simple_dataset = fetch_20newsgroups(
subset='train',
categories=['comp.sys.mac.hardware', 'soc.religion.christian', 'rec.sport.hockey'])
print (simple_dataset.data[0])
print (simple_dataset.data[-1])
print (simple_dataset.data[-2])
print (len(simple_dataset.data))
# ### Extract features from text
# +
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
vectorizer = TfidfVectorizer(max_df=500, min_df=10)
matrix = vectorizer.fit_transform(simple_dataset.data)
matrix.shape
# -
# ## AgglomerativeClustering, Neighbour joining
# +
from sklearn.cluster.hierarchical import AgglomerativeClustering
model = AgglomerativeClustering(n_clusters=3, affinity='cosine', linkage='complete')
preds = model.fit_predict(matrix.toarray())
# -
print(list(preds)[:5])
print(matrix[0])
vectorizer.get_feature_names()
vectorizer.get_feature_names()[877]
simple_dataset.data[0]
simple_dataset.target
preds
# Assessement
mapping = {2 : 1, 1: 2, 0: 0}
mapped_preds = [mapping[pred] for pred in preds]
# print (float(sum(mapped_preds != simple_dataset.target)) / len(simple_dataset.target))
print(acc(mapped_preds, simple_dataset.target))
import itertools
def validate_with_mappings(preds, target):
permutations = itertools.permutations([0, 1, 2])
for a, b, c in permutations:
mapping = {2 : a, 1: b, 0: c}
mapped_preds = [mapping[pred] for pred in preds]
# print (float(sum(mapped_preds != target)) / len(target))
print(acc(mapped_preds, target))
validate_with_mappings(preds, simple_dataset.target)
# ## KMeans
# +
from sklearn.cluster import KMeans
model = KMeans(n_clusters=3, random_state=1)
preds = model.fit_predict(matrix.toarray())
print (preds)
print (simple_dataset.target)
validate_with_mappings(preds, simple_dataset.target)
# -
# Compare with Linear Regression
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression()
print (cross_val_score(clf, matrix, simple_dataset.target).mean())
# ** Вопрос: ** Very big quality of K-Means, nearly as supervised algorithm, why?
# ## More complex dataset
noteasy_dataset = fetch_20newsgroups(
subset='train',
categories=['comp.sys.mac.hardware', 'comp.os.ms-windows.misc', 'comp.graphics'])
matrix = vectorizer.fit_transform(noteasy_dataset.data)
model = KMeans(n_clusters=3, random_state=1)
preds = model.fit_predict(matrix.toarray())
print (preds)
print (noteasy_dataset.target)
validate_with_mappings(preds, noteasy_dataset.target)
clf = LogisticRegression()
print (cross_val_score(clf, matrix, noteasy_dataset.target).mean())
# ## SVD + KMeans
# +
from sklearn.decomposition import TruncatedSVD
model = KMeans(n_clusters=3, random_state=42)
svd = TruncatedSVD(n_components=1000, random_state=123)
features = svd.fit_transform(matrix)
preds = model.fit_predict(features)
validate_with_mappings(preds, noteasy_dataset.target)
# -
model = KMeans(n_clusters=3, random_state=42)
svd = TruncatedSVD(n_components=200, random_state=321)
features = svd.fit_transform(matrix)
preds = model.fit_predict(features)
validate_with_mappings(preds, noteasy_dataset.target)
# # Quality of clusterization
# --------
#
# ### Homogeneity: each cluster contains only members of a single class
#
# ### Completeness: all members of a given class are assigned to the same cluster
#
# ### V-measure:
# ### $$v = 2 \cdot \frac{(homogeneity \cdot completeness)}{ (homogeneity + completeness)}$$
# +
from sklearn.metrics.cluster import homogeneity_score, completeness_score,v_measure_score
print(completeness_score(noteasy_dataset.target, preds))
print(homogeneity_score(noteasy_dataset.target, preds))
print(v_measure_score(noteasy_dataset.target, preds))
# -
# ### Results
# 1. Good results for both text datasets
# 2. On easy data clusterization methods work well
| day_14_Unsupervised/14_Unsupervised.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# cd '/Users/zoltan/Dropbox/Chronostratigraphy/stratigraph'
# +
import stratigraph as sg
from mayavi import mlab
import matplotlib.pyplot as plt
import numpy as np
import h5py
# set up graphics:
# %matplotlib qt
plt.rcParams['svg.fonttype'] = 'none'
# -
# load data from hdf5 file (this is a 'meanderpy' model)
fname = '/Users/zoltan/Dropbox/Chronostratigraphy/meanderpy_strat_model_example_1.hdf5'
f = h5py.File(fname, 'r')
model = f['model']
topo = np.array(model['topo'])
strat = np.array(model['strat'])
facies = np.array(model['facies'])
porosity = np.array(model['porosity'])
facies_code = {}
facies_code[int(np.array(model['point bar']))] = 'point bar'
facies_code[int(np.array(model['levee']))] = 'levee'
dx = float(np.array(model['dx']))
f.close()
# create facies volume (the facies data that comes from meanderpy is a 1D array and we need to expand it to 3D)
facies3d = np.zeros((strat.shape[0], strat.shape[1], strat.shape[2]-1))
for i in range(len(facies)):
facies3d[:,:,i] = facies[i] - 1 # facies codes should start at 0
# ## Simple block diagram, facies mode
# +
mlab.figure(bgcolor = (1,1,1))
ve = 5.0
scale = 1
strat_switch = 1
layers_switch = 1
contour_switch = 0
dx = 10.0
bottom = np.min(strat) - 2
colors = [[0.9,0.9,0],[0.5,0.25,0]]
line_thickness = 0.5
gap = 50
color_mode = 'facies'
h = 6.0
export = 0
topo_min = np.min(topo[:,:,-1])
topo_max = np.max(topo[:,:,-1])
ci = 1.0 # contour interval
sg.create_exploded_view(strat, None, facies3d, x0=0, y0=0, nx=1, ny=1,
gap=gap, dx=dx, ve=ve, scale=scale, plot_strat=True, plot_surfs=True, plot_contours=False,
plot_sides=True, color_mode = color_mode, colors=colors, colormap='viridis', line_thickness=line_thickness,
bottom=bottom, export=0, topo_min=topo_min, topo_max=topo_max, ci=ci, opacity=1.0)
# -
# ## Exploded view with 2 x 2 blocks, facies mode
# display 4 blocks, 2 in each direction (this takes a while)
mlab.figure(bgcolor = (1,1,1))
sg.create_exploded_view(strat, None, facies3d, x0=0, y0=0, nx=2, ny=2,
gap=gap, dx=dx, ve=ve, scale=scale, plot_strat=True, plot_surfs=True, plot_contours=False,
plot_sides=True, color_mode = color_mode, colors=colors, colormap='viridis', line_thickness=line_thickness,
bottom=bottom, export=0, topo_min=topo_min, topo_max=topo_max, ci=ci, opacity=1.0)
# ## Fence diagram, facies mode
mlab.figure(bgcolor = (1,1,1))
sg.create_fence_diagram(strat, None, facies3d, x0=0, y0=0, nx=2, ny=2, dx=dx, ve=ve, scale=scale,
plot_surfs=True, plot_sides=True, color_mode=color_mode,
colors=colors, colormap = 'viridis', line_thickness=line_thickness, bottom=bottom,
export=0, opacity=1.0)
# ## Exploded view with 2 x 2 blocks, property mode
mlab.figure(bgcolor = (1,1,1))
sg.create_exploded_view(strat, porosity, facies3d, x0=0, y0=0, nx=2, ny=2,
gap=gap, dx=dx, ve=ve, scale=scale, plot_strat=True, plot_surfs=True, plot_contours=False,
plot_sides=True, color_mode = 'property', colors=colors, colormap='viridis',
line_thickness=line_thickness,
bottom=bottom, export=0, topo_min=topo_min,topo_max=topo_max, ci=ci, opacity=1.0)
# ## Fence diagram, facies mode
mlab.figure(bgcolor = (1,1,1))
sg.create_fence_diagram(strat, porosity, facies3d, x0=0, y0=0, nx=2, ny=2, dx=dx, ve=ve, scale=scale,
plot_surfs=True, plot_sides=True, color_mode='property',
colors=colors, colormap = 'viridis', line_thickness=line_thickness, bottom=bottom,
export=0, opacity=1.0)
# ## Time-elevation curve ('Barrell plot')
# create time array (assumes that every point bar - overbank couplet was deposited in 5 years)
dt = 5.0
time = np.linspace(0, np.round(dt*(topo.shape[2]-1)/3), int((topo.shape[2]-1)/3)+1)
# only consider point bar - overbank couplets so that surfaces represent constant time increments:
elevation = topo[200, 200, ::3]
fig = sg.plot_strat_diagram(time, elevation, 'years', 'm', max(time), 2.0)
# ## Chronostratigraphic (Wheeler) diagram
np.shape(strat)
np.shape(strat)
# use every third surface in the topography array and every second surface in the stratigraphy array:
strat, wheeler, wheeler_strat, vacuity = sg.create_wheeler_diagram(topo[:,:,::3])
# strike section
plt.figure(figsize=(15,10))
plt.imshow(wheeler[:, 100, :].T, cmap='RdBu', vmin = -8, vmax = 8, extent = [0, dx*strat.shape[0], time[-1], 0],
interpolation='none', aspect='auto')
plt.gca().invert_yaxis()
plt.colorbar()
plt.xlabel('distance (m)', fontsize = 14)
plt.ylabel('time (years)', fontsize = 14)
plt.title('chronostratigraphic diagram', fontsize = 14)
plt.tight_layout()
# strike section, showing only what is preserved
plt.figure(figsize=(15,10))
plt.imshow(wheeler_strat[:, 100, :].T, cmap='RdBu', vmin = -8, vmax = 8, extent = [0, dx*strat.shape[0], time[-1], 0],
interpolation='none', aspect='auto')
plt.gca().invert_yaxis()
plt.colorbar()
plt.xlabel('distance (m)', fontsize = 14)
plt.ylabel('time (years)', fontsize = 14)
plt.title('chronostratigraphic diagram', fontsize = 14)
plt.tight_layout()
# dip section
plt.figure(figsize=(15,10))
plt.imshow(wheeler[200, :, :].T, cmap='RdBu', vmin = -8, vmax = 8, extent = [0, dx*strat.shape[0], time[-1], 0],
interpolation='none', aspect='auto')
plt.gca().invert_yaxis()
plt.colorbar()
plt.xlabel('distance (m)', fontsize = 14)
plt.ylabel('time (years)', fontsize = 14)
plt.title('chronostratigraphic diagram', fontsize = 14)
plt.tight_layout()
# time section
plt.figure(figsize=(15,10))
plt.imshow(wheeler[:, :, 60], cmap='RdBu', vmin = -8, vmax = 8, extent = [0, dx*strat.shape[1], 0, dx*strat.shape[0]],
interpolation='none')
plt.colorbar()
plt.xlabel('distance (m)', fontsize = 14)
plt.ylabel('distance (m)', fontsize = 14)
plt.title('chronostratigraphic diagram', fontsize = 14)
plt.axis('equal')
plt.tight_layout()
# 3D visualization with isosurface
mlab.figure(bgcolor = (1,1,1))
source = mlab.pipeline.scalar_field(np.swapaxes(wheeler, 0, 1))
source.spacing = [1,1,2]
mlab.pipeline.iso_surface(source, contours=[-1, 1], opacity=1, colormap='RdBu', vmin = -1.5, vmax = 1.5);
# 3D visualization with plane widgets
mlab.figure()
source = mlab.pipeline.scalar_field(np.swapaxes(wheeler, 0, 1))
source.spacing = [1,1,4]
for axis in ['x', 'y', 'z']:
plane = mlab.pipeline.image_plane_widget(source, plane_orientation = '{}_axes'.format(axis),
slice_index=i, colormap='RdBu', vmin = -8, vmax = 8);
# ## Cross sections (no need for Mayavi)
# load data from hdf5 file (this is a 'meanderpy' model); it needs to be reloaded as
# running 'sg.create_wheeler_diagram' has changed the dimensions of the 'strat' array but it did not change
# the 'facies3d' array
fname = '/Users/zoltan/Dropbox/Chronostratigraphy/meanderpy_strat_model_example_1.hdf5'
f = h5py.File(fname, 'r')
model = f['model']
topo = np.array(model['topo'])
strat = np.array(model['strat'])
facies = np.array(model['facies'])
porosity = np.array(model['porosity'])
facies_code = {}
facies_code[int(np.array(model['point bar']))] = 'point bar'
facies_code[int(np.array(model['levee']))] = 'levee'
dx = float(np.array(model['dx']))
f.close()
fig = sg.plot_model_cross_section_EW(strat, porosity, facies3d, dx, 260, color_mode='facies',
flattening_ind = False, ve = 5, list_of_colors = ['yellow', 'brown'])
fig = sg.plot_model_cross_section_NS(strat, porosity, facies3d, dx, 160, color_mode='facies',
flattening_ind = False, ve = 10, list_of_colors = ['yellow', 'brown'])
# ## XES 02 Experiment
# +
import os
dirname = '/Users/zoltan/Dropbox/Chronostratigraphy/XES_02/final_topography_data/topography/'
T = np.zeros((261,111,101))
filenames = os.listdir(dirname)
for filename in filenames:
surf_no = int(filename[8:-4])
xyz = np.loadtxt(dirname+filename)
z = xyz[:,2]
T[:,:,surf_no-1] = np.reshape(z,(261,111))
dirname = '/Users/zoltan/Dropbox/Chronostratigraphy/XES_02/final_topography_data/basement_topography/'
B = np.zeros((261,111,101))
filenames = os.listdir(dirname)
for filename in filenames:
surf_no = int(filename[:-4])
xyz = np.loadtxt(dirname+filename)
z = xyz[:,2]
B[:,:,surf_no-1] = np.reshape(z,(261,111))
# np.shape(T)
# -
import pandas as pd
df = pd.read_csv('/Users/zoltan/Dropbox/Chronostratigraphy/XES_02/final_topography_data/sealevel_and_scantimes.csv')
df[:10]
# +
def convert_to_seconds(string):
h = int(string.split(':')[0])
m = int(string.split(':')[1])
s = int(string.split(':')[2])
return h*60*60 + m*60 + s
exp_time = np.nan * np.ones((T.shape[2],))
sea_level = np.nan * np.ones((T.shape[2],))
for i in range(len(exp_time)):
if len(df.loc[df['Scan number'] == i+1]) > 0:
exp_time[i] = convert_to_seconds(df.loc[df['Scan number'] == i+1]['run time (hhh:mm:ss)'].values[0])
sea_level[i] = df.loc[df['Scan number'] == i+1]['sl(mm)']
missing_times = np.where(np.isnan(exp_time)==1)[0]
for i in missing_times:
exp_time[i] = (exp_time[i-1] + exp_time[i+1]) * 0.5
sea_level[i] = (sea_level[i-1] + sea_level[i+1]) * 0.5
exp_time = np.delete(exp_time, np.array([2, 5, 12, 25, 30, 31, 48, 60])) # get rid of locations with missing data
sea_level = np.delete(sea_level, np.array([2, 5, 12, 25, 30, 31, 48, 60])) # get rid of locations with missing data
plt.figure(figsize=(10, 6))
plt.plot(exp_time, sea_level, 'o-')
plt.xlabel('time (seconds)', fontsize = 16)
plt.ylabel('sea level (mm)', fontsize = 16);
T = np.delete(T, np.array([2, 5, 12, 25, 30, 31, 48, 60]), axis=2) # get rid of locations with no data
B = np.delete(B, np.array([2, 5, 12, 25, 30, 31, 48, 60]), axis=2) # get rid of locations with no data
# +
# create resampled and smooth sea level curve
sampling_rate = 900 # resample at every 900 seconds
time1, sea_level_rs1 = sg.resample_elevation_spl(exp_time, sea_level, sampling_rate)
plt.figure()
plt.plot(exp_time, sea_level, '.-')
plt.plot(time1, sea_level_rs1, '.-')
time2, sea_level_rs2 = sg.resample_elevation_int1d(exp_time, sea_level, sampling_rate)
plt.plot(time2, sea_level_rs2, '.-')
# sea_level_rs1[804:868] = sea_level_rs2[804:868] # replace sea level 1 w/ sea level 2
# sea_level_rs1[972:1215] = sea_level_rs2[972:1215] # replace sea level 1 w/ sea level 2
# sea_level_rs1[267:289] = sea_level_rs2[267:289] # replace sea level 1 w/ sea level 2
# sea_level_rs1[324:405] = sea_level_rs2[324:405] # replace sea level 1 w/ sea level 2
sea_level_rs1[535:578] = sea_level_rs2[535:578] # replace sea level 1 w/ sea level 2
sea_level_rs1[648:810] = sea_level_rs2[648:810] # replace sea level 1 w/ sea level 2
plt.plot(time1, sea_level_rs1, 'b.-')
time = time1.copy()
sea_level_rs = sea_level_rs1.copy()
# +
# resample topography and subsidence arrays:
topo = np.zeros((T.shape[0], T.shape[1], len(time)))
for j in range(T.shape[1]):
for i in range(T.shape[0]):
elevation = T[i, j, :].copy()
time, elevation = sg.resample_elevation_int1d(exp_time, elevation, sampling_rate)
topo[i,j,:] = elevation
subsid = np.zeros((B.shape[0], B.shape[1], len(time)))
for j in range(B.shape[1]):
for i in range(B.shape[0]):
elevation = B[i, j, :].copy()
time, elevation = sg.resample_elevation_int1d(exp_time, elevation, sampling_rate)
subsid[i,j,:] = elevation
# adjust topographic and subsidence surfaces in the proximal corners (for aestethic reasons only)
inds = np.indices(np.shape(subsid[:,:,0]))
inds2 = np.argwhere(inds[0] < -4.8*inds[1] + 120)
inds3 = np.argwhere(inds[0] > 5.13*inds[1] + 142)
for i in range(np.shape(subsid)[2]):
subsid[:, :, i][inds2[:,0], inds2[:,1]] = -229.6
subsid[:, :, i][inds3[:,0], inds3[:,1]] = -229.6
for i in range(np.shape(topo)[2]):
topo[:, :, i][inds2[:,0], inds2[:,1]] = 0
topo[:, :, i][inds3[:,0], inds3[:,1]] = 0
# account for subsidence:
topo_s = topo.copy()
for i in range(0, topo.shape[2]):
topo_s[:,:,i] = topo_s[:,:,i]+(subsid[:,:,-1]-subsid[:,:,i])
# -
# QC topographic surfaces (corrected for subsidence)
plt.figure(figsize=(18, 10))
for i in range(0, topo_s.shape[2], 20):
plt.plot(topo_s[130, :, i], 'k', linewidth=0.5)
# create a number of time-elevation plots, from proximal to distal locations
fig = sg.plot_strat_diagram(time, topo_s[125, 20, :], 'seconds', 'mm', time[-1], np.max(topo_s[125, 20, :]))
fig = sg.plot_strat_diagram(time, topo_s[125, 60, :], 'seconds', 'mm', time[-1], np.max(topo_s[125, 60, :]))
fig = sg.plot_strat_diagram(time, topo_s[125, 100, :], 'seconds', 'mm', time[-1], np.max(topo_s[125, 100, :]))
# convert topographic surfaces to stratigraphy
strat = sg.topostrat(topo_s)
# create wheeler diagram(s)
strat, wheeler, wheeler_strat, vacuity = sg.create_wheeler_diagram(topo_s)
plt.figure(figsize=(20,15))
plt.imshow(wheeler[125,:,:].T, extent = [0, 111, time[-1], 0], cmap='RdBu', vmin = -4, vmax = 4,
interpolation='none', aspect='auto')
plt.gca().invert_yaxis()
plt.colorbar();
sl = 20 * (sea_level_rs - np.min(sea_level_rs))/213.5 # sea level curve
plt.plot(2 +np.max(sl)-sl, time, 'k', linewidth=3)
plt.ylim(0, time[-1])
plt.xlim(2, 111)
plt.title('chronostratigraphic diagram', fontsize=20);
plt.figure(figsize=(20,15))
plt.imshow(wheeler_strat[125,:,:].T, extent = [0, 111, time[-1], 0], cmap='RdBu', vmin = -4, vmax = 4,
interpolation='none', aspect='auto')
plt.gca().invert_yaxis()
plt.colorbar();
sl = 20 * (sea_level_rs - np.min(sea_level_rs))/213.5
plt.plot(2 +np.max(sl)-sl, time, 'k', linewidth=3)
plt.ylim(0, time[-1])
plt.xlim(2, 111)
plt.title('chronostratigraphic diagram w/ vacuity', fontsize=20);
# 3D visualization with isosurface
mlab.figure()
source = mlab.pipeline.scalar_field(np.swapaxes(wheeler, 0, 1))
source.spacing = [5, 1, 0.1]
mlab.pipeline.iso_surface(source, contours=[-2, 2], opacity=0.5, colormap='RdBu', vmin = -4, vmax = 4);
# 3D visualization with plane widgets
mlab.figure()
source = mlab.pipeline.scalar_field(np.swapaxes(wheeler, 0, 1))
source.spacing = [5, 1, 0.2]
for axis in ['x', 'y', 'z']:
plane = mlab.pipeline.image_plane_widget(source, plane_orientation = '{}_axes'.format(axis),
slice_index=i, colormap='RdBu', vmin = -4, vmax = 4);
# create 3D facies array (as a function of water depth in this case)
ny, nx, nz = np.shape(strat)
facies = np.zeros((ny, nx, nz))
for i in range(facies.shape[2]):
topo_sl = topo[:, :, i] - sea_level_rs[i]
facies_sl = np.zeros(np.shape(topo_sl))
facies_sl[topo_sl >= 0] = 0
facies_sl[(topo_sl < 0) & (topo_sl >= -100)] = 1
facies_sl[topo_sl < -100] = 2
facies[:, :, i] = facies_sl
# eliminating nans from 'strat' array
for i in range(np.shape(strat)[2]):
t = strat[:, :, i]
t[np.isnan(t) == 1] = t[136, 1]
strat[:, :, i] = t
# plot cross section colored by water depth (= facies in this case)
fig = sg.plot_model_cross_section_EW(strat, facies, facies, dx = 50, xsec = 135, color_mode = 'facies',
map_aspect = 0.2,
line_freq = 5,
flattening_ind = False, units = 'mm')
fig = sg.plot_model_cross_section_NS(strat, facies, facies, dx = 10, xsec = 50, color_mode = 'facies',
map_aspect = 0.2,
line_freq = 5,
flattening_ind = False, units = 'mm')
| stratigraph/Stratigraph_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Creating-XML-for-NETCONF-via-LXML" data-toc-modified-id="Creating-XML-for-NETCONF-via-LXML-1"><span class="toc-item-num">1 </span>Creating XML for NETCONF via LXML</a></div><div class="lev1 toc-item"><a href="#Alternative-One:-Namespaces" data-toc-modified-id="Alternative-One:-Namespaces-2"><span class="toc-item-num">2 </span>Alternative One: Namespaces</a></div><div class="lev2 toc-item"><a href="#Namespaces-we-care-about" data-toc-modified-id="Namespaces-we-care-about-21"><span class="toc-item-num">2.1 </span>Namespaces we care about</a></div><div class="lev2 toc-item"><a href="#Register-the-namespaces-with-ElementTree" data-toc-modified-id="Register-the-namespaces-with-ElementTree-22"><span class="toc-item-num">2.2 </span>Register the namespaces with ElementTree</a></div><div class="lev3 toc-item"><a href="#Using-xpath-to-retrieve-Elements" data-toc-modified-id="Using-xpath-to-retrieve-Elements-221"><span class="toc-item-num">2.2.1 </span>Using xpath to retrieve Elements</a></div><div class="lev1 toc-item"><a href="#Alternative-Two:-No-(Less)-Namespaces" data-toc-modified-id="Alternative-Two:-No-(Less)-Namespaces-3"><span class="toc-item-num">3 </span>Alternative Two: No (Less) Namespaces</a></div><div class="lev1 toc-item"><a href="#Writing-XML-to-a-File" data-toc-modified-id="Writing-XML-to-a-File-4"><span class="toc-item-num">4 </span>Writing XML to a File</a></div>
# -
# # Creating XML for NETCONF via LXML
# LXML is a Python library to deal with XML element trees (among other things). It provides quite a lot of capabilities and in the context of NETCONF here's a little sample code that shows some ways how to create XML data for NETCONF via a script (vs. manually copyingand pasting it).
#
# But first of all let's lay some ground work:
from lxml import etree as ET
def pp(element):
print(ET.tostring(element, pretty_print=True).decode('utf-8'))
# # Alternative One: Namespaces
# This approach actually uses namespaces throughout the creation of the objects. This might look a bit over the top but it actually helps a lot to understand where those namespaces come into play. When looking at e.g. the `ietf-interfaces.yang` model, we can see pretty much at the top:
#
# module ietf-interfaces {
#
# namespace "urn:ietf:params:xml:ns:yang:ietf-interfaces";
# prefix if;
#
# These statements define the namespace and the prefix for the given namespace. We will see these two elements later in our XML creation for an interface element in the `ietf-interfaces` YANG model.
#
# ## Namespaces we care about
# The following three namespaces are relevant int the context of `ietf-interfaces` YANG model or more specifically to describe an interface within that model.
if_ns = ('if', 'urn:ietf:params:xml:ns:yang:ietf-interfaces')
it_ns = ('ianaift', 'urn:ietf:params:xml:ns:yang:iana-if-type')
ip_ns = ('ip', 'urn:ietf:params:xml:ns:yang:ietf-ip')
# ## Register the namespaces with ElementTree
# If we register the namespace with ET automatically creates the proper prefix ('if' for ietf-interfaces) when creating an element without explicitely specifying a namespace map IF the element tag is properly specified **with** a namespace designator. This looks like
#
# {urn:ietf:params:xml:ns:yang:ietf-interfaces}interfaces
#
# The below is a bit of a hack to coerce the namespace tuples we defined above into function paramenters. Actually, `register_namespace()` takes two:
#
# 1. the prefix -and-
# 2. the actual URN for the namespace.
ET.register_namespace(*if_ns)
ET.register_namespace(*it_ns)
ET.register_namespace(*ip_ns)
# Now, let's create a 'config' container with several sub-containers which eventually make up an ietf-interfaces interface tree describing a loopback interface with a name, a state and some IP addresses.
# +
root = ET.Element('config')
# Interface container
interfaces = ET.SubElement(root, '{%s}interfaces' % if_ns[1])
interface = ET.SubElement(interfaces, '{%s}interface' % if_ns[1])
# Interface Name
e = ET.SubElement(interface, '{%s}name' % if_ns[1])
e.text = 'Loopback99'
# # enabled?
e = ET.SubElement(interface, '{%s}enabled' % if_ns[1])
e.text = 'true'
# Interface Type
e = ET.SubElement(interface, '{%s}type' % if_ns[1], nsmap=dict((if_ns, it_ns)))
e.text = '%s:softwareLoopback' % it_ns[0]
# IPv4 Element
ip = ET.SubElement(interface, '{%s}ipv4' % ip_ns[1])
address = ET.SubElement(ip, '{%s}address' % ip_ns[1])
e = ET.SubElement(address, '{%s}ip' % ip_ns[1])
e.text = '1.2.3.4'
e = ET.SubElement(address, '{%s}netmask' % ip_ns[1])
e.text = '255.0.0.0'
# IPv6 Element
ip = ET.SubElement(interface, '{%s}ipv6' % ip_ns[1])
address = ET.SubElement(ip, '{%s}address' % ip_ns[1])
e = ET.SubElement(address, '{%s}ip' % ip_ns[1])
e.text = '2001:db8:99::1'
e = ET.SubElement(address, '{%s}prefix-length' % ip_ns[1])
e.text = '64'
# -
# Finally, let's print the resulting ElementTree:
pp(root)
# As you can see, all of the elements within the `<config>` container are properly prefixed with the namespace and the proper `xmlns` attribute.
#
# ### Using xpath to retrieve Elements
# We can use the `xpath` method to extract information from the ElementTree. Note that providing the proper namespace maps **and** prefixing the elements with their associated namespaces is required. Results are returned as a list.
root.xpath('/config/if:interfaces/if:interface/ip:ipv6/ip:address/ip:ip/text()', namespaces=dict((if_ns, ip_ns)))
pp(root.xpath('//*[local-name() = "ipv4"]')[0])
# If we want to specify a particular element in the tree without specifying namespaces then we have to use the `local-name()` function:
root.xpath('//*[local-name() = "ipv6"]/*[local-name()="address"]/*[local-name()="ip"]/text()')
root.xpath('//*[local-name() = "ipv6/address/ip"]/text()')
root.xpath('//*[local-name() = "ip"]/text()')
# `xpath` is a powerful way to retrieve information from an XML tree. It can use expression functions, variable replacesment, regular expression and more. See [the documentation page](http://lxml.de/xpathxslt.html#xpath) for more examples and information. And some additional [examples on xpath functions](https://msdn.microsoft.com/en-us/library/ms256086(v=vs.110).aspx).
#
# # Alternative Two: No (Less) Namespaces
# Or at least minimize the use of namespaces and operate primarily with the default namesspace. The default namespace has no prefix but elements can still carry the xmlns attributes to distinguish between different namespaces, where appropriate.
if_ns = {None: 'urn:ietf:params:xml:ns:yang:ietf-interfaces'}
it_ns = {'ianaift': 'urn:ietf:params:xml:ns:yang:iana-if-type'}
ip_ns = {None: 'urn:ietf:params:xml:ns:yang:ietf-ip'}
# +
# root element
root = ET.Element('config')
# Interface container
interfaces = ET.SubElement(root, 'interfaces', nsmap=if_ns)
interface = ET.SubElement(interfaces, 'interface')
# Interface Name
e = ET.SubElement(interface, 'name')
e.text = 'Loopback99'
# # enabled?
e = ET.SubElement(interface, 'enabled')
e.text = 'true'
# Interface Type
e = ET.SubElement(interface, 'type', nsmap={**if_ns, **it_ns})
e.text = 'ianaift:softwareLoopback'
# IPv4 Element
ip = ET.SubElement(interface, 'ipv4', nsmap=ip_ns)
address = ET.SubElement(ip, 'address')
e = ET.SubElement(address, 'ip')
e.text = '1.2.3.4'
e = ET.SubElement(address, 'netmask')
e.text = '255.0.0.0'
# IPv6 Element
ip = ET.SubElement(interface, 'ipv6', nsmap=ip_ns)
address = ET.SubElement(ip, 'address')
e = ET.SubElement(address, 'ip')
e.text = '2001:db8:99::1'
e = ET.SubElement(address, 'prefix-length')
e.text = '64'
# -
pp(root)
# The resulting XML above looks a bit more concise then the previous result. Syntactically and also from a functionality point of view (in the ncclient / NETCONF context) they are exactly the same. It's up to the reader which method they prefer.
#
# The idea here is to only pass a namespace map (`nsmap`) when required, e.g. when defining the first element within the tree with a different namespace than the previous element used.
#
# There is one oddity in the above code worth mentioning: The construct `nsmap={**if_ns, **it_ns}`. What does this do? This is one way of 'merging' two dictionaries into one. Essentially `nsmap = if_ns + if_ns`. But this isn't defined in Python.
#
# The reason why we need two namespaces for the `<type>` element is that the element itself is within the previous namespace ('if'). The value of the element, however, is from a different namespace ('ianaift'). This needs to be designated and hence the dual-namespace map.
# # Writing XML to a File
# Finally, if we want to write the entire Element Tree into a file, we can do something like this:
filename = '../tmp/output.xml'
with open(filename,'w') as f:
f.write(ET.tostring(root).decode('utf-8'))
| DOCKER/sections/LXML-NETCONF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Chapter 3 – Classification**
#
# _This notebook contains all the sample code and solutions to the exercises in chapter 3._
# # Setup
# First, let's make sure this notebook works well in both python 2 and 3, import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures:
# + pycharm={"name": "#%%\n", "is_executing": false}
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "classification"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
# -
# # MNIST
# **Warning**: `fetch_mldata()` is deprecated since Scikit-Learn 0.20. You should use `fetch_openml()` instead. However, it returns the unsorted MNIST dataset, whereas `fetch_mldata()` returned the dataset sorted by target (the training set and the test test were sorted separately). In general, this is fine, but if you want to get the exact same results as before, you need to sort the dataset using the following function:
# + pycharm={"name": "#%%\n", "is_executing": false}
def sort_by_target(mnist):
reorder_train = np.array(sorted([(target, i) for i, target in enumerate(mnist.target[:60000])]))[:, 1]
reorder_test = np.array(sorted([(target, i) for i, target in enumerate(mnist.target[60000:])]))[:, 1]
mnist.data[:60000] = mnist.data[reorder_train]
mnist.target[:60000] = mnist.target[reorder_train]
mnist.data[60000:] = mnist.data[reorder_test + 60000]
mnist.target[60000:] = mnist.target[reorder_test + 60000]
# + pycharm={"name": "#%%\n", "is_executing": false}
try:
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1, cache=True)
mnist.target = mnist.target.astype(np.int8) # fetch_openml() returns targets as strings
sort_by_target(mnist) # fetch_openml() returns an unsorted dataset
except ImportError:
from sklearn.datasets import fetch_mldata
mnist = fetch_mldata('MNIST original')
mnist["data"], mnist["target"]
# + pycharm={"name": "#%%\n", "is_executing": false}
mnist.data.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
X, y = mnist["data"], mnist["target"]
X.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
y.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
def plot_digit(data):
image = data.reshape(28, 28)
plt.imshow(image, cmap = mpl.cm.binary,
interpolation="nearest")
plt.axis("off")
# + pycharm={"name": "#%%\n", "is_executing": false}
# EXTRA
def plot_digits(instances, images_per_row=10, **options):
size = 28
images_per_row = min(len(instances), images_per_row)
images = [instance.reshape(size,size) for instance in instances]
n_rows = (len(instances) - 1) // images_per_row + 1
row_images = []
n_empty = n_rows * images_per_row - len(instances)
images.append(np.zeros((size, size * n_empty)))
for row in range(n_rows):
rimages = images[row * images_per_row : (row + 1) * images_per_row]
row_images.append(np.concatenate(rimages, axis=1))
image = np.concatenate(row_images, axis=0)
plt.imshow(image, cmap = mpl.cm.binary, **options)
plt.axis("off")
# + pycharm={"name": "#%%\n", "is_executing": false}
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
# + pycharm={"name": "#%%\n", "is_executing": false}
import numpy as np
shuffle_index = np.random.permutation(60000)
X_train, y_train = X_train[shuffle_index], y_train[shuffle_index]
# + pycharm={"name": "#%%\n", "is_executing": false}
from sklearn.model_selection import StratifiedKFold
from sklearn.base import clone
from sklearn.model_selection import cross_val_score
from sklearn.base import BaseEstimator
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_score, recall_score
from sklearn.metrics import f1_score
from sklearn.metrics import precision_recall_curve
# + pycharm={"name": "#%%\n", "is_executing": false}
def plot_precision_recall_vs_threshold(precisions, recalls, thresholds):
plt.plot(thresholds, precisions[:-1], "b--", label="Precision", linewidth=2)
plt.plot(thresholds, recalls[:-1], "g-", label="Recall", linewidth=2)
plt.xlabel("Threshold", fontsize=16)
plt.legend(loc="upper left", fontsize=16)
plt.ylim([0, 1])
# + pycharm={"name": "#%%\n", "is_executing": false}
def plot_precision_vs_recall(precisions, recalls):
plt.plot(recalls, precisions, "b-", linewidth=2)
plt.xlabel("Recall", fontsize=16)
plt.ylabel("Precision", fontsize=16)
plt.axis([0, 1, 0, 1])
# + pycharm={"name": "#%%\n", "is_executing": false}
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
def plot_roc_curve(fpr, tpr, label=None):
plt.plot(fpr, tpr, linewidth=2, label=label)
plt.plot([0, 1], [0, 1], 'k--')
plt.axis([0, 1, 0, 1])
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
# -
# **Note**: we set `n_estimators=10` to avoid a warning about the fact that its default value will be set to 100 in Scikit-Learn 0.22.
# + pycharm={"name": "#%%\n", "is_executing": false}
from sklearn.multiclass import OneVsOneClassifier
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train.astype(np.float64))
# + pycharm={"name": "#%%\n", "is_executing": false}
def plot_confusion_matrix(matrix):
"""If you prefer color and a colorbar"""
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111)
cax = ax.matshow(matrix)
fig.colorbar(cax)
# + pycharm={"name": "#%%\n", "is_executing": false}
# from sklearn.model_selection import GridSearchCV
# param_grid = [
# {'n_neighbors': [3, 4, 5], 'weights': ['distance', 'uniform']}
# ]
# #
from sklearn.neighbors import KNeighborsClassifier
# neigh = KNeighborsClassifier()
# #
# grid_search = GridSearchCV(neigh, param_grid, cv=2, n_jobs=-1, verbose=100)
# grid_search.fit(X_train, y_train)
# + pycharm={"name": "#%%\n", "is_executing": false}
# grid_search.best_params_
#
# + pycharm={"name": "#%%\n", "is_executing": false}
# grid_search.best_score_
# + pycharm={"name": "#%%\n", "is_executing": false}
from sklearn.metrics import accuracy_score
# y_pred = grid_search.predict(X_test)
neigh = KNeighborsClassifier(n_neighbors=4, weights='distance', n_jobs=-1)
# neigh.fit(X_train, y_train)
# y_pred = neigh.predict(X_test)
# accuracy_score(y_test, y_pred)
# + pycharm={"name": "#%%\n", "is_executing": false}
# Write a function that can shift an MNIST image in any direction (left, right, up, or
# down) by one pixel. Then, for each image in the training set, create four shifted
# copies (one per direction) and add them to the training set. Finally, train your best
# model on this expanded training set and measure its accuracy on the test set.
#You can use the shift() function from the scipy.ndimage.interpolation module. For example,
# shift(image, [2, 1], cval=0) shifts the image 2 pixels down and 1 pixel to the right.
from scipy.ndimage.interpolation import shift
def shift_arr(arr, reshape_value = 28):
arr2 = arr.reshape(reshape_value, reshape_value)
return [
shift(arr2, [1, 0]), #down
shift(arr2, [0, 1]), #right
shift(arr2, [-1, 0]),#up
shift(arr2, [0, -1]) #left
]
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr = [1, 2, 3, 4]
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr0 = np.array(my_arr).reshape(2, 2).tolist()
my_arr0
# + pycharm={"name": "#%%\n", "is_executing": false}
# for img in X_train:
# shifted = shift_arr(img)
my_arr1 = np.array(my_arr0)
my_arr1
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr2 = shift_arr(np.array([1, 2, 3, 4]), 2)
my_arr2
# + pycharm={"name": "#%%\n", "is_executing": false}
#my_arr2.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr3 = np.array([my_arr0])
my_arr3
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr3.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr4 = [my_arr1] + my_arr2
my_arr4
#my_arr2.shape()
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr1.flatten()
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr5 = np.array(my_arr4)
my_arr5
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr6 = map(lambda e: e.flatten(), my_arr4)
my_arr7 = list(my_arr6)
my_arr7
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr8 = np.array(my_arr7)
my_arr8
# + pycharm={"name": "#%%\n", "is_executing": false}
np.array([[[1,2], [3,4]], [[5,6], [7,8]]]).flatten()
#-> need [[1,2], [3,4], [5,6], [7,8]]
# + pycharm={"name": "#%%\n", "is_executing": false}
[[1,2], [3,4]] + [[5,6], [7,8]]
# + pycharm={"name": "#%%\n", "is_executing": false}
from functools import reduce
list(reduce(lambda e1, e2: e1 + e2, [[[1,2], [3,4]], [[5,6], [7,8]]]))
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr8.tolist()
# + pycharm={"name": "#%%\n", "is_executing": false}
def shift_arr2(arr, reshape_value = 28):
arr0 = np.array(arr)
arr1 = arr0.reshape(reshape_value, reshape_value)
arr2 = [arr1] + shift_arr(arr0, reshape_value)
arr3 = map(lambda e: e.flatten(), arr2)
arr4 = list(arr3)
arr5 = np.array(arr4).tolist()
return arr5
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr9 = shift_arr2([1, 2, 3, 4], 2)
my_arr9
# + pycharm={"name": "#%%\n", "is_executing": false}
X_train2 = X_train[:3]
X_train2
# + pycharm={"name": "#%%\n", "is_executing": false}
X_train2.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
X_train3 = np.array(list(map(shift_arr2, X_train2.tolist())))
Xt3size = X_train3.shape
Xt3size
# list(map(shift_arr2, X_train2.tolist()))
# + pycharm={"name": "#%%\n", "is_executing": false}
X_train4 = reduce(lambda e1, e2: e1 + e2, X_train3.tolist())
X_t4_size = np.array(X_train4).shape
X_t4_size
# + pycharm={"name": "#%%\n", "is_executing": false}
X_t5 = np.array(X_train4).reshape(Xt3size[0] * Xt3size[1], Xt3size[2])
X_t5
# + pycharm={"name": "#%%\n", "is_executing": false}
X_t5.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
X_train_aug0 = list(map(shift_arr2, X_train.tolist()))
X_train_aug1 = np.array(X_train_aug0)
X1_size = X_train_aug1.shape
X_train_aug = X_train_aug1.reshape(X1_size[0] * X1_size[1], X1_size[2])
X_train_aug.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr10 = [1,2,3]
aug_size = 5
my_arr11 = np.array(list(map(lambda e: [e] * aug_size, my_arr10)))
my_arr11
# + pycharm={"name": "#%%\n", "is_executing": false}
my_arr11.reshape(aug_size * my_arr10.__len__())
# + pycharm={"name": "#%%\n", "is_executing": false}
Y_train_aug = np.array(list(map(lambda e: [e] * aug_size, y_train))).reshape(aug_size * y_train.shape[0])
Y_train_aug.shape
# + pycharm={"name": "#%%\n", "is_executing": false}
neigh.fit(X_train_aug, Y_train_aug)
# + pycharm={"name": "#%%\n", "is_executing": false}
y_pred = neigh.predict(X_test)
# + pycharm={"name": "#%%\n", "is_executing": false}
accuracy_score(y_test, y_pred)
# + pycharm={"name": "#%%\n"}
| 03_exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Economic Data Processing (MADDISON)
#
# >Maddison Project Database, version 2018. Bolt, Jutta, <NAME>, <NAME> and <NAME> (2018), “Rebasing ‘Maddison’: new income comparisons and the shape of long-run economic development”, Maddison Project Working paper 10
#
# ## Data Dictionary
#
# | Full data | Data in single table |
# |-------------------|---------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
# | countrycode | 3-letter ISO country code |
# | country | Country name |
# | year | Year |
# | cgdppc | Real GDP per capita in 2011US\$, multiple benchmarks (suitable for cross-country income comparisons) |
# | rgdpnapc | Real GDP per capita in 2011US\$, 2011 benchmark (suitable for cross-country growth comparisons) |
# | pop | Population, mid-year (thousands) |
# | i_cig | 0/1/2: observation is extrapolated (0), benchmark (1), or interpolated (2) |
# | i_bm | For benchmark observations: 1: ICP PPP estimates, 2: Historical income benchmarks, 3: Real wages and urbanization, 4: Multiple of subsistence, 5: Braithwaite (1968) PPPs |
# | Partial countries | Data for selected sub-national units with long time series |
# +
import pandas as pd
import pycountry
# %matplotlib inline
pd.set_option('display.float_format', lambda x: '%.3f' % x)
# -
# ## Load The File
df = pd.read_excel("../data/external/Economy/MADDISON/mpd2018.xlsx",
sheet_name='Full data')
df.sample(5)
# ## Standardize Country Codes
""" Only Select rows with valid country codes
"""
country_locations = []
for country in df['countrycode']:
try:
pycountry.countries.lookup(country)
country_locations.append(True)
except LookupError:
country_locations.append(False)
df = df[country_locations]
# ## Standardize Indexes
# ### Years (1995≤ x ≥2017)
df = df[df['year'] >= 1995]
df = df[df['year'] <= 2017]
# ### Reindex & Rename
df.rename(
{
"year": "Year",
"countrycode": "Country Code",
"cgdppc": "Maddison GDPPC"
},
axis='columns',
inplace=True)
df.set_index(["Country Code", "Year"], inplace=True)
# ## Clean Data
# ### Remove unneeded variables
df.drop(["country", "i_cig", "i_bm", "rgdpnapc", "pop"],
axis='columns',
inplace=True)
# ### Data Types
df.dtypes
# ## Save Data
df.to_pickle("../data/processed/Economic_MADDISON.pickle")
| notebooks/4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
DATE = "2021-08-22"
TASK = "iterative-correction-via-resampling"
DIR = mkpath("$(homedir())/$(DATE)")
# +
import Pkg
pkgs = [
# "BioAlignments",
"BioSequences",
# "Clustering",
# "CodecZlib",
# "Colors",
# "Combinatorics",
# "DataFrames",
# "DataStructures",
"Dates",
# "DelimitedFiles",
# "Distances",
# "EzXML",
"FASTX",
# "GFF3",
# "HTTP",
# "Impute",
# "JSON",
"Graphs",
"LSHFunctions",
# "Measures",
"MetaGraphs",
"https://github.com/cjprybol/Mycelia.git",
# "NumericIO",
# "PlotlyJS",
# "Plots",
"Primes",
# "Printf",
# "ProgressMeter",
"Random",
"Revise",
"SparseArrays",
# "Statistics",
"StatsBase",
# "StatsPlots",
# "StringDistances",
# "uCSV",
# "XLSX",
]
unregistered_packages = filter(pkg -> occursin(r"(^https|git$)", pkg), pkgs)
registered_packages = setdiff(pkgs, unregistered_packages)
for pkg in registered_packages
try
eval(Meta.parse("import $(pkg)"))
catch
Pkg.add(pkg)
Pkg.build(pkg)
eval(Meta.parse("import $(pkg)"))
end
end
for pkg_url in unregistered_packages
pkg_name = replace(basename(pkg_url), ".git" => "")
try
eval(Meta.parse("import $(pkg_name)"))
catch
Pkg.develop(url=pkg_url)
Pkg.build(pkg_name)
eval(Meta.parse("import $(pkg_name)"))
end
end
# -
if Sys.isapple()
BANDAGE = "/Applications/Bandage.app/Contents/MacOS/Bandage"
else
BANDAGE = "Bandage"
end
function assess_kmer_sparsity_in_reads(k, observations)
# @show k, observations
canonical_kmer_counts = Mycelia.count_canonical_kmers(BioSequences.DNAMer{k}, observations)
# canonical_kmer_counts = Mycelia.count_canonical_kmers(BioSequences.DNAMer{k}, first(observations))
# # @show canonical_kmer_counts
# for i in 2:length(observations)
# canonical_kmer_counts_ = Mycelia.count_canonical_kmers(BioSequences.DNAMer{k}, observations[i])
# canonical_kmer_counts = merge!(+, canonical_kmer_counts, canonical_kmer_counts_)
# end
# canonical_kmer_counts = map(o -> Mycelia.count_canonical_kmers(BioSequences.DNAMer{k}, o), observations)...)
total_observed_canonical_kmers = length(keys(canonical_kmer_counts))
total_possible_canonical_kmers = (4^k)/2
sparsity = round(total_observed_canonical_kmers/total_possible_canonical_kmers*100, digits=2)
@show sparsity
return sparsity
end
function sequence_to_canonical_kmers(kmer_type, sequence)
return (BioSequences.canonical(kmer.fw) for kmer in BioSequences.each(kmer_type, sequence))
end
function determine_edge_weights(graph)
outgoing_edge_probabilities = determine_edge_weights(graph, true)
incoming_edge_probabilities = determine_edge_weights(graph, false)
return Dict(true => outgoing_edge_probabilities, false => incoming_edge_probabilities)
end
function determine_edge_weights(graph, strand)
kmers = [graph.vprops[v][:kmer] for v in Graphs.vertices(graph)]
outgoing_edge_weights = SparseArrays.spzeros(length(kmers), length(kmers))
for (kmer_index, kmer) in enumerate(kmers)
if !strand
kmer = BioSequences.reverse_complement(kmer)
end
downstream_neighbors = []
downstream_neighbor_weights = []
for neighbor in BioSequences.neighbors(kmer)
canonical_neighbor = BioSequences.canonical(neighbor)
neighbor_orientation = neighbor == canonical_neighbor
neighbor_index_range = searchsorted(kmers, canonical_neighbor)
if !isempty(neighbor_index_range)
@assert length(neighbor_index_range) == 1
neighbor_index = first(neighbor_index_range)
if Graphs.has_edge(graph, Graphs.Edge(kmer_index, neighbor_index))
edge_orientations = graph.eprops[Graphs.Edge(kmer_index, neighbor_index)][:orientations]
this_edge_orientation = (source_orientation = strand, destination_orientation = neighbor_orientation)
if this_edge_orientation in edge_orientations
edge_weight = graph.eprops[Graphs.Edge(kmer_index, neighbor_index)][:weight]
outgoing_edge_weights[kmer_index, neighbor_index] = edge_weight
end
end
end
end
end
return outgoing_edge_weights
end
function determine_edge_probabilities(edge_weights)
edge_probabilities = deepcopy(edge_weights)
for orientation in [true, false]
for row in 1:size(edge_probabilities[orientation], 1)
edge_weights = edge_probabilities[orientation][row, :]
edge_total = sum(edge_weights)
for (col, edge_weight) in enumerate(edge_weights)
if edge_total > 0
edge_probabilities[orientation][row, col] = edge_weight/edge_total
else
edge_probabilities[orientation][row, col] = 0.0
end
end
end
end
return edge_probabilities
end
function random_step(current_vertex, current_orientation, step_probabilities)
outgoing_edge_likelihoods = step_probabilities[initial_orientation][current_vertex, :]
chosen_step = StatsBase.sample(outgoing_edge_likelihoods.nzind, StatsBase.weights(outgoing_edge_likelihoods.nzval))
possible_orientations = observation_graph.eprops[Graphs.Edge(current_vertex, chosen_step)][:orientations]
possible_orientations = filter(o -> o.source_orientation == current_orientation, possible_orientations)
chosen_orientation = rand(possible_orientations).destination_orientation
chosen_oriented_step = (vertex = chosen_step, orientation = chosen_orientation)
return chosen_oriented_step
end
function random_walk(observation_graph, initial_vertex, initial_orientation, step_probabilities; n_steps=size(step_probabilities[true], 1))
step_count = 0
outgoing_edge_likelihoods = step_probabilities[initial_orientation][initial_vertex, :]
walk = Vector{NamedTuple{(:vertex, :orientation), Tuple{Int64, Bool}}}()
done = (step_count > n_steps) || (sum(outgoing_edge_likelihoods) == 0)
current_vertex = initial_vertex
current_orientation = initial_orientation
while !done
# @show "choosing"
chosen_step = StatsBase.sample(outgoing_edge_likelihoods.nzind, StatsBase.weights(outgoing_edge_likelihoods.nzval))
chosen_step
possible_orientations = observation_graph.eprops[Graphs.Edge(current_vertex, chosen_step)][:orientations]
possible_orientations
current_orientation
possible_orientations = filter(o -> o.source_orientation == current_orientation, possible_orientations)
possible_orientations
chosen_orientation = rand(possible_orientations).destination_orientation
chosen_oriented_step = (vertex = chosen_step, orientation = chosen_orientation)
push!(walk, chosen_oriented_step)
current_vertex = chosen_step
current_orientation = chosen_orientation
outgoing_edge_likelihoods = step_probabilities[last(walk).orientation][last(walk).vertex, :]
step_count += 1
# @show outgoing_edge_likelihoods
done = (step_count > n_steps) || (sum(outgoing_edge_likelihoods) == 0)
end
return walk
end
# generate a genome
sequence_length = 10
sequence = BioSequences.randdnaseq(Random.seed!(sequence_length), sequence_length)
sequence_id = string(hash(sequence))
fasta_record = FASTX.FASTA.Record(sequence_id, sequence)
reference_fasta_file = "$(DIR)/LENGTH-$(sequence_length).fasta"
open(reference_fasta_file, "w") do io
fastx_io = FASTX.FASTA.Writer(io)
write(fastx_io, fasta_record)
close(fastx_io)
end
# +
# randomly sample reads with errors from the genome
# 1%
error_rate = 0.01
n_reads = 100
# observations = [Mycelia.observe(FASTX.sequence(record), error_rate = error_rate) for i in 1:n_reads]
observations = [Mycelia.observe(fasta_record, error_rate = error_rate) for i in 1:n_reads];
# +
# build assembly graph
min_k = 7
max_k = min(minimum(length(observations)), 61)
k_options = Primes.primes(min_k, max_k)
k_index = findfirst(k -> assess_kmer_sparsity_in_reads(k, observations) < 100, k_options)
k = k_options[k_index]
observations_file = "$(DIR)/LENGTH-$(sequence_length)-DEPTH-$(n_reads).fastq"
open(observations_file, "w") do io
fastq_io = FASTX.FASTQ.Writer(io)
for record in observations
write(fastq_io, record)
end
close(fastq_io)
end
# -
reference_graph = Mycelia.fastx_to_kmer_graph(BioSequences.DNAMer{k}, reference_fasta_file)
reference_kmers = [reference_graph.vprops[v][:kmer] for v in Graphs.vertices(reference_graph)]
observation_graph = Mycelia.fastx_to_kmer_graph(BioSequences.DNAMer{k}, observations_file)
# +
# visualize
gfa_file = observations_file * ".k-$k.gfa"
Mycelia.graph_to_gfa(observation_graph, gfa_file)
run(`$(BANDAGE) image $(gfa_file) $(gfa_file).svg --depwidth .9 --deppower .9`)
# --nodewidth <float> Average node width (0.5 to 1000, default: 5)
# --depwidth <float> Depth effect on width (0 to 1, default: 0.5)
# --deppower <float> Power of depth effect on width (0 to 1, default: 0.5)
html_path_to_svg = "./" * repeat("../", length(split(pwd(), '/')) - 3)
html_path_to_svg *= replace("$(gfa_file).svg", "$(homedir())/" => "")
x = "<img src=$(html_path_to_svg)>"
display("text/html", x)
# display("image/svg+xml", read("$(gfa_file).svg", String))
# -
kmer_counts = Dict(observation_graph.vprops[v][:kmer] => observation_graph.vprops[v][:weight] for v in Graphs.vertices(observation_graph))
total_observed_kmers = sum(values(kmer_counts))
kmer_probabilities = Dict(k => v/total_observed_kmers for (k,v) in kmer_counts)
scale = 250
Mycelia.plot_kmer_frequency_spectra(values(kmer_counts), size=(2scale,scale), title="kmer frequencies")
distance_to_reference_graph = 1 - LSHFunctions.jaccard(Set(reference_kmers), Set(keys(kmer_counts)))
# +
# step 1, hard-filter singletons
# -
filtered_vertices = findall(v -> observation_graph.vprops[v][:weight] >= 5, Graphs.vertices(observation_graph))
filtered_observation_graph, vertex_map = Graphs.induced_subgraph(observation_graph, filtered_vertices)
# +
# visualize
gfa_file = observations_file * ".k-$k.filtered.gfa"
Mycelia.graph_to_gfa(filtered_observation_graph, gfa_file)
run(`$(BANDAGE) image $(gfa_file) $(gfa_file).svg --depwidth .9 --deppower .9`)
# --nodewidth <float> Average node width (0.5 to 1000, default: 5)
# --depwidth <float> Depth effect on width (0 to 1, default: 0.5)
# --deppower <float> Power of depth effect on width (0 to 1, default: 0.5)
html_path_to_svg = "./" * repeat("../", length(split(pwd(), '/')) - 3)
html_path_to_svg *= replace("$(gfa_file).svg", "$(homedir())/" => "")
x = "<img src=$(html_path_to_svg)>"
display("text/html", x)
# display("image/svg+xml", read("$(gfa_file).svg", String))
# -
kmer_counts = Dict(filtered_observation_graph.vprops[v][:kmer] => filtered_observation_graph.vprops[v][:weight] for v in Graphs.vertices(filtered_observation_graph))
total_observed_kmers = sum(values(kmer_counts))
kmer_probabilities = Dict(k => v/total_observed_kmers for (k,v) in kmer_counts)
scale = 250
Mycelia.plot_kmer_frequency_spectra(values(kmer_counts), size=(2scale,scale), title="kmer frequencies")
distance_to_reference_graph = 1 - LSHFunctions.jaccard(Set(reference_kmers), Set(keys(kmer_counts)))
# +
# step 2, re-simulate reads as a means of error correction
# -
kmers = [filtered_observation_graph.vprops[v][:kmer] for v in Graphs.vertices(filtered_observation_graph)]
kmer_weights = [filtered_observation_graph.vprops[v][:weight] for v in Graphs.vertices(filtered_observation_graph)]
kmer_probabilities = kmer_weights ./ sum(kmer_weights)
edge_weights = determine_edge_weights(filtered_observation_graph)
edge_weights[true]
edge_weights = determine_edge_weights(filtered_observation_graph)
edge_probabilities = determine_edge_probabilities(edge_weights)
# +
kmers = [filtered_observation_graph.vprops[v][:kmer] for v in Graphs.vertices(filtered_observation_graph)]
kmer_weights = [filtered_observation_graph.vprops[v][:weight] for v in Graphs.vertices(filtered_observation_graph)]
kmer_probabilities = kmer_weights ./ sum(kmer_weights)
edge_weights = determine_edge_weights(filtered_observation_graph)
edge_probabilities = determine_edge_probabilities(edge_weights)
step_probabilities = deepcopy(edge_probabilities)
for orientation in [true, false]
# @show orientation
oriented_step_probabilities = step_probabilities[orientation]
for row in 1:size(step_probabilities[orientation], 1)
step_probabilities_ = step_probabilities[orientation][row, :] .* kmer_probabilities
if sum(step_probabilities_) > 0
step_probabilities[orientation][row, :] .= step_probabilities_ ./ sum(step_probabilities_)
end
end
end
# -
new_records = FASTX.FASTQ.Record[]
for observation in observations
read_length = length(FASTX.sequence(observation))
max_steps = read_length - filtered_observation_graph.gprops[:k]
# apply a squaring penalty to kmer_weights?
initial_vertex = StatsBase.sample(1:length(kmers), StatsBase.weights(kmer_weights.^2))
# @show "choosing initial vertex $initial_vertex with probability $(kmer_weights[initial_vertex])"
initial_orientation = rand(Bool)
forward_walk = random_walk(filtered_observation_graph, initial_vertex, initial_orientation, step_probabilities, n_steps=max_steps)
remaining_steps = max_steps - length(forward_walk)
reverse_walk = random_walk(filtered_observation_graph, initial_vertex, !initial_orientation, step_probabilities, n_steps = remaining_steps)
full_walk = [[(vertex=x.vertex, orientation=!x.orientation) for x in reverse(reverse_walk)]..., [(vertex = initial_vertex, orientation = initial_orientation)]..., forward_walk...]
oriented_path = [(x.vertex, x.orientation) for x in full_walk]
new_seq = Mycelia.oriented_path_to_sequence(filtered_observation_graph, oriented_path)
new_record = FASTX.FASTQ.Record(
FASTX.identifier(observation)*"-k$k",
FASTX.identifier(observation),
new_seq,
StatsBase.sample(FASTX.quality(observation), length(new_seq))
)
# @show "here"
push!(new_records, new_record)
end
new_records;
corrected_fastq_file = replace(observations_file, r"\.fastq" => ".k$k.fastq")
open(corrected_fastq_file, "w") do io
fastq_writer = FASTX.FASTQ.Writer(io)
for record in new_records
write(fastq_writer, record)
end
close(fastq_writer)
end
corrected_observation_graph = Mycelia.fastx_to_kmer_graph(BioSequences.DNAMer{k}, corrected_fastq_file)
# +
# visualize
gfa_file = corrected_fastq_file * ".k-$k.gfa"
Mycelia.graph_to_gfa(corrected_observation_graph, gfa_file)
run(`$(BANDAGE) image $(gfa_file) $(gfa_file).svg --depwidth .9 --deppower .9`)
# --nodewidth <float> Average node width (0.5 to 1000, default: 5)
# --depwidth <float> Depth effect on width (0 to 1, default: 0.5)
# --deppower <float> Power of depth effect on width (0 to 1, default: 0.5)
html_path_to_svg = "./" * repeat("../", length(split(pwd(), '/')) - 3)
html_path_to_svg *= replace("$(gfa_file).svg", "$(homedir())/" => "")
x = "<img src=$(html_path_to_svg)>"
display("text/html", x)
# display("image/svg+xml", read("$(gfa_file).svg", String))
# -
kmer_counts = Dict(corrected_observation_graph.vprops[v][:kmer] => corrected_observation_graph.vprops[v][:weight] for v in Graphs.vertices(corrected_observation_graph))
total_observed_kmers = sum(values(kmer_counts))
kmer_probabilities = Dict(k => v/total_observed_kmers for (k,v) in kmer_counts)
scale = 250
Mycelia.plot_kmer_frequency_spectra(values(kmer_counts), size=(2scale,scale), title="kmer frequencies")
# +
# compare the new graph to the error-free graph to the error-corrected graph
# -
distance_to_reference_graph = 1 - LSHFunctions.jaccard(Set(reference_kmers), Set(keys(kmer_counts)))
| docs/_src/5.Development/2021-08-22-iterative-correction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.9 64-bit
# language: python
# name: python3
# ---
import tensorflow as tf
# +
# Session 写法一
tens1 = tf.constant([1, 2, 3])
# 创建一个会话
session1 = tf.Session()
# 使用该创建会话执行想要得到的结果语句
# 如:session.run(result)
# 此处为得到上述张量的结果
try:
print(session1.run(tens1))
except:
print("Exception !")
# 关闭会话,释放资源
finally:
session1.close()
# +
# Session 写法二
node1 = tf.constant(3.0, tf.float32, name="node1")
node2 = tf.constant(4.0, tf.float32, name="node2")
result = tf.add(node1, node2)
# 通过 Python 上下文管理器管理会话
# 此方法不需要调用 session.close()方法 关闭会话,系统将自动释放资源
with tf.Session() as sessions:
# 使用该会话计算结果
print(sessions.run(result))
# +
# Session 默认会话
# 当指定默认会话时,可通过 tf.Tensor.eval() 计算所需张量值
node1 = tf.constant(3.0, tf.float32, name="node1")
node2 = tf.constant(4.0, tf.float32, name="node2")
result = tf.add(node1, node2)
session2 = tf.compat.v1.Session()
with session2.as_default():
print(result.eval())
session3 = tf.compat.v1.Session()
# 下面写法均可得到相同的输出结果
print(session3.run(result))
# 无缺省会话,需要手动指定
print(result.eval(session=session3))
# +
# Session 交互环境下设置默认会话
node1 = tf.constant(3.0, tf.float32, name="node1")
node2 = tf.constant(4.0, tf.float32, name="node2")
result = tf.add(node1, node2)
# tf.compat.v1.InteractiveSession() 会将生成的会话自动注册为一个默认会话
session4 = tf.compat.v1.InteractiveSession()
print(result.eval())
session4.close()
| Course/1.0/Basic/5-Session.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '1'
# +
import sys
SOURCE_DIR = os.path.dirname(os.path.dirname(os.path.abspath(__name__)))
sys.path.insert(0, SOURCE_DIR)
# -
import malaya_speech
from pysptk import sptk
import numpy as np
# +
import tensorflow as tf
# tf.compat.v1.enable_eager_execution()
# -
vggvox_v2 = malaya_speech.gender.deep_model(model = 'vggvox-v2')
speaker_model = malaya_speech.speaker_vector.deep_model('vggvox-v2')
freqs = {'female': [100, 600], 'male': [50, 250]}
# +
from scipy.signal import get_window
from scipy import signal
import soundfile as sf
import random
sr = 22050
def butter_highpass(cutoff, fs, order=5):
nyq = 0.5 * fs
normal_cutoff = cutoff / nyq
b, a = signal.butter(order, normal_cutoff, btype='high', analog=False)
return b, a
b, a = butter_highpass(30, sr, order=5)
# +
from math import ceil
def speaker_normalization(f0, index_nonzero, mean_f0, std_f0):
f0 = f0.astype(float).copy()
f0[index_nonzero] = (f0[index_nonzero] - mean_f0) / std_f0
f0[index_nonzero] = np.clip(f0[index_nonzero], -3, 4)
return f0
def preprocess_wav(x):
if x.shape[0] % 256 == 0:
x = np.concatenate((x, np.array([1e-06])), axis=0)
y = signal.filtfilt(b, a, x)
wav = y * 0.96 + (np.random.uniform(size = y.shape[0]) - 0.5)*1e-06
return wav
def get_f0(wav, lo, hi):
f0_rapt = sptk.rapt(wav.astype(np.float32)*32768, sr, 256, min=lo, max=hi, otype=2)
index_nonzero = (f0_rapt != -1e10)
mean_f0, std_f0 = np.mean(f0_rapt[index_nonzero]), np.std(f0_rapt[index_nonzero])
return speaker_normalization(f0_rapt, index_nonzero, mean_f0, std_f0)
def pad_seq(x, base = 8):
len_out = int(base * ceil(float(x.shape[0]) / base))
len_pad = len_out - x.shape[0]
assert len_pad >= 0
return np.pad(x, ((0, len_pad), (0, 0)), 'constant'), x.shape[0]
def get_speech(f, hop_size = 256):
x, fs = malaya_speech.load(f, sr = sr)
wav = preprocess_wav(x)
lo, hi = freqs.get(vggvox_v2(x), [50, 250])
f0 = np.expand_dims(get_f0(wav, lo, hi), -1)
mel = malaya_speech.featurization.universal_mel(wav)
batch_max_steps = random.randint(16384, 110250)
batch_max_frames = batch_max_steps // hop_size
if len(mel) > batch_max_frames:
interval_start = 0
interval_end = len(mel) - batch_max_frames
start_frame = random.randint(interval_start, interval_end)
start_step = start_frame * hop_size
wav = wav[start_step : start_step + batch_max_steps]
mel = mel[start_frame : start_frame + batch_max_frames, :]
f0 = f0[start_frame : start_frame + batch_max_frames, :]
v = speaker_model([wav])[0]
v = v / v.max()
return wav, mel, f0, v
# -
wav, mel, f0, v = get_speech('../speech/example-speaker/female.wav')
wav_1, mel_1, f0_1, v_1 = get_speech('../speech/example-speaker/khalil-nooh.wav')
mels, mel_lens = malaya_speech.padding.sequence_nd([mel, mel_1], dim = 0, return_len = True)
mels.shape, mel_lens
f0s, f0_lens = malaya_speech.padding.sequence_nd([f0, f0_1], dim = 0, return_len = True)
f0s.shape, f0_lens
vs = malaya_speech.padding.sequence_nd([v, v_1], dim = 0)
vs.shape
X = tf.placeholder(tf.float32, [None, None, 80])
X_f0 = tf.placeholder(tf.float32, [None, None, 1])
len_X = tf.placeholder(tf.int32, [None])
V = tf.placeholder(tf.float32, [None, 512])
from malaya_speech.train.model.fastspeechsplit import inference as fastspeechsplit
# from malaya_speech.train.model.fastspeechsplit import inference as fastspeechsplit
from malaya_speech.train.model import speechsplit, fastspeechsplit, fastspeech
hparams = speechsplit.hparams
config = malaya_speech.config.fastspeech_config
config = fastspeech.Config(vocab_size = 1, **config)
interplnr = speechsplit.InterpLnr(hparams)
model = fastspeechsplit.Model(config, hparams)
model_F0 = fastspeechsplit.Model_F0(config, hparams)
bottleneck_speaker = tf.keras.layers.Dense(hparams.dim_spk_emb)
speaker_dim = bottleneck_speaker(V)
x_f0_intrp = interplnr(tf.concat([X, X_f0], axis = -1), len_X)
x_f0_intrp.shape
f0_org_intrp = speechsplit.quantize_f0_tf(x_f0_intrp[:,:,-1])
x_f0_intrp_org = tf.concat((x_f0_intrp[:,:,:-1], f0_org_intrp), axis=-1)
f0_org = speechsplit.quantize_f0_tf(X_f0[:,:,0])
f0_org_intrp, x_f0_intrp_org, X, speaker_dim, f0_org
o = model(x_f0_intrp_org, X, speaker_dim, len_X)
o
_, _, _, f0_target = model_F0(X, f0_org, len_X)
f0_target
sess = tf.Session()
sess.run(tf.global_variables_initializer())
o_ = sess.run(o, feed_dict = {
X: mels, X_f0: f0s, len_X: mel_lens, V: vs
})
o_[0].shape, o_[1].shape, o_[2].shape, o_[3].shape, o_[4].shape
o = sess.run([f0_target], feed_dict = {
X: mels, X_f0: f0s, len_X: mel_lens, V: vs
})
o[0].shape
saver = tf.train.Saver()
saver.save(sess, 'test/model.ckpt')
# !ls -lh test
# !rm -rf test
| test/test-fastspeechsplit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Fundamentals of Computer Science for Neuroengineering
#
# ## Lecture 2: Conditional Branching and Iteration
#
# This notebook is adapted from <NAME>'s Scipy course:
# https://github.com/nickdelgrosso/SciPyCourse2016
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## This Lecture's Learning Goals:
# - How do **if-else statements** work in Python?
# - How do you perform **conditional variable assignment** in Python?
# - How do you **Index** and **Slice** collections in Python?
# - How can you **copy** lists and dicts?
# - How do you perform **C-Style For Loops** in Python?
# - How do you use **List Comprehensions** to **filter** lists?
# + [markdown] slideshow={"slide_type": "slide"}
# # Programmatic Control Flow: the If-elif-else statements
# When you want your program to do one thing **if** something is **True**, and something **else** **if** some other condition is **True**, or **else** do some final thing if all those conditions before were **False**...
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Comparison Always Comes down to True vs False
#
# ### Comparison Operators: <, >, ==, !=, and, or
# + slideshow={"slide_type": "fragment"}
a, b, c = -2, 0, 2
a < b
a <= b
a != b
a < b and b < c
b < a or b < c
a < b < c
# b == b < c
# b == (b < c)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## To Be Safe, You Can Use Parentheses ( ) when performing multiple comparisons
# + slideshow={"slide_type": "fragment"}
(a < b) or (b < c)
# + [markdown] slideshow={"slide_type": "fragment"}
# But make sure both sides are comparisons!
# + slideshow={"slide_type": "fragment"}
a or b < c
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Conditional Assignment: Single-line "if" statements
# You can choose to assign one value to a variable if a condition is True, and another value if it is False!
# + slideshow={"slide_type": "fragment"}
subject_code = 20
cond = "Experimental" if subject_code < 30 else "Control"
cond
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Multi-line If-else Statements
# + slideshow={"slide_type": "fragment"}
a, b, c, a_string, a_bool = -2, 0, 2, 'hello', True
if a > b:
print('The first statement was True!')
elif c > a:
print('The second statement was True')
else:
print('Nothing was True')
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Special Operator for collections: "in"
# + slideshow={"slide_type": "fragment"}
animals = ['dog', 'cat', 'mouse', 'rat', 'bat']
'bat' in animals
['bat'] in animals
'o' in 'mouse'
situation = 'Science' if 'rat' in animals else 'Pet'
# situation
# + [markdown] slideshow={"slide_type": "slide"}
# # Indexing and Slicing Collections
#
# When you have a collection of something, you often will want to select a specific object from the collection. If you know where the object is located in the collection (it's **index**, or **key**), you only need to give that index and you'll get your object! In Python, you index a collection using the **square brackets: [ ]**
#
# **Note:** Python is a **Zero-Indexed** language, so the first **Element** of a collection has an index value of 0.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Indexing Ordered Collections: List and Tuple
# + slideshow={"slide_type": "fragment"}
subjects = ['A1', 'A2', 'B2', 'B3']
subjects[0]
# + slideshow={"slide_type": "fragment"}
subjects[-1]
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Indexing Unordered Collections: Dict
# Unordered collections are indexed using the same **syntax**, but by inserting the **key** instead.
# + slideshow={"slide_type": "fragment"}
conditions = {'knockout': [1, 2, 1, 2],
'control': [5, 6, 5],}
conditions.keys()
# +
# conditions[]
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Slicing: Indexing multiple values in an Ordered Collection
#
# You can get a continuous subset of a collection by doing **Slicing**. This is done using the **Colon :**
#
# **Note:** Python uses a **Half-Open** Slicing Convention, which means that the final index given is ignored, but the first value is used.
# -
subjects = ('A1', 'B2', 'C3', 'D4', 'E5', 'F6', 'G7')
subjects[1:-2]
subjects[::2]
# + slideshow={"slide_type": "skip"}
subjects[:4]
# + slideshow={"slide_type": "skip"}
subjects[1:-1]
# + [markdown] slideshow={"slide_type": "skip"}
# You can also skip values using a second colon:
# + slideshow={"slide_type": "skip"}
subjects[1:-1:2]
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Indexed Assignment
#
# With Mutable objects, you can modify an element of a collection in-place via indexing and slicing!
# + slideshow={"slide_type": "fragment"}
subjects = ['A1', 'B2', 'C3', 'D4', 'E5', 'F6', 'G7']
print(subjects)
subjects[3] = 'New Subject'
print(subjects)
# -
subjects[0:2] = ['a', 'b', 'c']
subjects
a = [1, 2, 3]
b = (1, 2, 3)
id(a)
id(b)
a.append(5)
id(a)
# + [markdown] slideshow={"slide_type": "slide"}
# # Mutability and Identity of Variables
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Object Memory Identity: "id()" and the "is" operator
#
# While the comparison operators above examine the **value** of a variable, one can also check to see if two variables are the same piece of data in memory.
#
# We can get the numeric memory address of the variable using the built-in **id()** function
# + slideshow={"slide_type": "fragment"}
a = [50.23, 10]
b = a
b[1] = 1000
# + slideshow={"slide_type": "fragment"}
id(a)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Mutable vs Immutable
# - **Immutable** types cannot be modified. Even if just their value would be changing, the entire variable is created completely new, with a new memory address.
# - Important Immutable classes: str, int, float, tuple
# - **Mutable** types can be modified. When their value is changed, they retain their identity.
# - Important mutable classes: list, dict
# + [markdown] slideshow={"slide_type": "subslide"}
# # Aside: Things to Consider When Choosing a Proper Data Collection Type
# - Do these pieces of data belong together?
# - Is the size of my collection conceivably variable?
# - How independent are these values, in terms of what they represent?
# - Essentially: Is it important for my collection to be mutable?
# - What technical requirements do I have for working with this data?
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Constant IDs are a very useful feature of Mutable objects
# - Helps you avoid copying data
# - Keeps code more readable
# + slideshow={"slide_type": "fragment"}
a = [1, 2, 3]
b = a
b = [1,2, 3]
c = a.append(4)
type(c)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Gotcha: Ints and Floats Have Unusual id() assignment behaviors!
# Only rely on the id() of an object if it is mutable. Basically dicts and lists. ints and floats, on the other hand, are treated as mostly immutable objects, except in certain cases. See the examples below; do they behave as you'd expect?
# + slideshow={"slide_type": "fragment"}
a = 3
b = a
a is b
# + slideshow={"slide_type": "fragment"}
a = 3
b = a
a = 4
a is b
# + slideshow={"slide_type": "fragment"}
3 is 3
# + slideshow={"slide_type": "fragment"}
a = 3
b = 3
a is b
# + slideshow={"slide_type": "fragment"}
a, b = 3, 3
a is b
# + slideshow={"slide_type": "fragment"}
a = 500 # ints above 255 are treated different here.
b = 500
a is b
# + slideshow={"slide_type": "fragment"}
a, b = 500, 500
a is b
# + [markdown] slideshow={"slide_type": "slide"}
# # Loops
#
# Loops are used to repeat lines of code.
# - To repeat the code until a certain condition is fulfilled, use a **while** loop
# - To repeat the code a predetermined number of times, use a **for** loop.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## While Loops
#
# While loops work basically the same as the if statement.
#
# **Caution**: It is possible to get infinite loops if you aren't careful! Use the Ctrl-C command to stop the program, if you find yourself in that situation!
# + slideshow={"slide_type": "fragment"}
var = 0
while var < 10:
print('var is currently {}!'.format(var))
var += 1
# + [markdown] slideshow={"slide_type": "subslide"}
# # Iteration and the "For" Loop
#
# ### Aside: Terminology
# - **Iteration** is the process of going through each element of a group, one at a time. If you can iterate through the object, it is said to be **Iterable**.
# - For example, a list is an iterable, since it is a collection of multiple items. Strings are also iterable.
# - An **Iterator** is an iterable object.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### The "For loop" requests each element from an iterator, one at a time, until all elements have been requested.
# + slideshow={"slide_type": "fragment"}
subjects = ['A1', 4, [''], 'C3', '94', 'E5', 'F6', 'G7']
print(subjects)
for subject in subjects:
print(type(subject))
print(subject)
print('Finished!')
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Generators
#
# A **Generator** is a function that can create iterators that are not collections. This means that each value is calculated and given only at the time that is requested. This has huge computational benefits!
# - In Python 3, the **range()**, **dict.keys()**, and **dict.values()** functions are generators. To make it a list, use the list constructor: list(dict.keys())
# + slideshow={"slide_type": "fragment"}
print(subjects)
for i in range(len(subjects)):
print(subjects[i])
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Counting your Elements: the enumerate() function
#
# **enumerate()** takes an iterator as input, and outputs an iterator of tuples: (index, element)
# + slideshow={"slide_type": "fragment"}
for idx, subject in enumerate(subjects):
print( 'Subject {}: {}'.format(idx, subject))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Iterating through Multiple Iterators: the zip() function
#
# The **zip()** function takes multiple iterators and returns the elements of each of them at a time!
# + slideshow={"slide_type": "fragment"}
subjects = ['A1', 'B2', 'C3', 'D4', 's1']
conditions = ['Exp', 'Ctl', 'Exp', 'Ctl']
for subject, condition in zip(subjects, conditions):
print( 'Subject {} was in Condition {}'.format(subject, condition))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Advanced Example: Combining zip() and enumerate()
# + slideshow={"slide_type": "fragment"}
subjects = ['A1', 'B2', 'C3', 'D4']
conditions = ['Exp', 'Ctl', 'Exp', 'Ctl']
for index, (subject, condition) in enumerate(zip(subjects, conditions)):
print( '{}: Subject {} was in Condition {}'.format(index, subject, condition))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Iterating through Dicts
# By default, each element of a dict will simply give its key. For example:
# + slideshow={"slide_type": "fragment"}
subj_count = {'A': 6, 'B': 8, 'C': 2}
for condition in subj_count:
print(subj_count[condition])
# + [markdown] slideshow={"slide_type": "subslide"}
# To get values alone, use **dict.values()**:
# + slideshow={"slide_type": "fragment"}
subj_count = {'A': 6, 'B': 8, 'C': 2}
for total in subj_count.values():
print(total)
# + [markdown] slideshow={"slide_type": "fragment"}
# To get the key, value pairs, use **dict.items()**:
# + slideshow={"slide_type": "fragment"}
subj_count = {'A': 6, 'B': 8, 'C': 2}
for condition, total in subj_count.items():
print('{} Subjects were in Condition {}'.format(total, condition))
# + [markdown] slideshow={"slide_type": "slide"}
# # List Comprehensions: Single-Line For Loops
#
# Sometimes, you just want to do a simple calculation and build a new list out of an old list! Python's **List Comprehensions** feature is really great for this!
# + slideshow={"slide_type": "subslide"}
subject_letter = []
for subject in subjects:
subject_letter.append(subjects[1])
subject_letter
# + slideshow={"slide_type": "subslide"}
subjects = ['A1', 'B2', 'C3', 'D4']
subject_letter = [subject[0] for subject in subjects]
subject_letter
# + slideshow={"slide_type": "fragment"}
full_subject_names = ['Subj' + subject[0] for subject in subjects]
full_subject_names
# + [markdown] slideshow={"slide_type": "subslide"}
# # Conditional List Comprehensions
# You can also only include values for going in the new list if a given statement is True!
# + slideshow={"slide_type": "fragment"}
subjects = ['A1', 'B2', 'C3', 'D4']
high_subjects = {subject[0]: int(subject[1]) for subject in subjects if int(subject[1]) - 1}
high_subjects
# + [markdown] slideshow={"slide_type": "slide"}
# ## This Lecture's Learning Goals:
#
# - How do **if-else statements** work in Python?
# - How do you perform **conditional variable assignment** in Python?
# - How do you **Index** and **Slice** collections in Python?
# - What are **Iterators**? How do you iterate in Python?
# - How do you perform **C-Style For Loops** in Python?
# - How do you use **List Comprehensions** to **filter** lists?
# + [markdown] slideshow={"slide_type": "slide"}
# # Questions / Discussion
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercises
#
# - https://www.practicepython.org/exercise/2014/03/19/07-list-comprehensions.html
# - https://www.practicepython.org/solution/2014/07/25/13-fibonacci-solutions.html
# - https://www.practicepython.org/exercise/2017/01/24/33-birthday-dictionaries.html
# -
| notebooks/block1/block1_session3_programming_concepts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
using Random
using Base: @kwdef
using Parameters: @unpack
using Plots
# +
@kwdef struct SOMParameter{}
N::UInt32 = 20 # Linear size of 2D map
n_teacher::UInt32 = 10000 # of teacher signal
Random.seed!(100)
#rand(1,1)
end
function update!(param::SOMParameter, I::Vector, dt)
@unpack N, n_teacher, = param
nodes = rand(N,N,3)
p3=wireframe(x,y,z,title="wireframe")
plot(p3,layout=(1,1),size=(250,250),fmt=:png)
"""
Learning
"""
teacher = rand(n_teacher, 3)
for n = 1:n_teacher
end
| SNN/SOM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Cleaning
#
# The WESAD Dataset is huge (12GB). But all of it is not useful. As mentioned in the original paper some part of the signal, that have labels > 3 are not important for stress data or are due to device malfuction.
#
# In this notebook I have cleaned the dataset to only the relevant that is needed for this project. This reduces the dimensionality of the dataset and also ensures better results.
#
# I have further done certain analysis on the data, which are also shown here
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.123993, "end_time": "2021-08-27T16:23:32.354902", "exception": false, "start_time": "2021-08-27T16:23:32.230909", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
import pickle
import gc
import os
import ctypes
# + [markdown] papermill={"duration": 0.105177, "end_time": "2021-08-27T16:23:32.567318", "exception": false, "start_time": "2021-08-27T16:23:32.462141", "status": "completed"} tags=[]
# # User Input
# ### Change the below cell and input the number of the Subject you want
#
# There were 17 participants in the test, but unfortunately device malfunctioned for two of them. So to get the neat and useful data of a paticular subject, set the file number accordingly.
# From
# [2,3,4,5,6,7,8,9,10,11,13,14,15,16,17]
# + papermill={"duration": 0.113013, "end_time": "2021-08-27T16:23:32.788082", "exception": false, "start_time": "2021-08-27T16:23:32.675069", "status": "completed"} tags=[]
file_num:int = 8
# -
# Change the path below as well
# + papermill={"duration": 0.113901, "end_time": "2021-08-27T16:23:33.008124", "exception": false, "start_time": "2021-08-27T16:23:32.894223", "status": "completed"} tags=[]
filename_ = "../input/wesad-stress-detection/PKL Datas/S{}.pkl".format(file_num)
# + papermill={"duration": 0.116768, "end_time": "2021-08-27T16:23:33.229686", "exception": false, "start_time": "2021-08-27T16:23:33.112918", "status": "completed"} tags=[]
print(file_num)
# + papermill={"duration": 0.117672, "end_time": "2021-08-27T16:23:33.453594", "exception": false, "start_time": "2021-08-27T16:23:33.335922", "status": "completed"} tags=[]
def create_dataframe(filename):
"It returns a pandas Dataframe consisting of chest data"
# reading the data
infile = open(filename,'rb')
new_dict = pickle.load(infile, encoding= 'latin1')
infile.close()
#
data = pd.DataFrame()
# Acceleration data
data['ACCX'] = new_dict['signal']['chest']['ACC'][:,0]
data['ACCY'] = new_dict['signal']['chest']['ACC'][:,1]
data['ACCZ'] = new_dict['signal']['chest']['ACC'][:,2]
# Reading input data
for i in new_dict['signal']['chest'].keys() :
if i != 'ACC' :
data[i] = new_dict['signal']['chest'][i]
# labels data
data['label'] = new_dict['label']
return data
# + papermill={"duration": 13.680124, "end_time": "2021-08-27T16:23:47.238130", "exception": false, "start_time": "2021-08-27T16:23:33.558006", "status": "completed"} tags=[]
data = create_dataframe(filename_)
# + papermill={"duration": 4.404172, "end_time": "2021-08-27T16:23:52.708602", "exception": false, "start_time": "2021-08-27T16:23:48.304430", "status": "completed"} tags=[]
import pandas as pd
import numpy as np
import seaborn as sns
from pylab import rcParams
import matplotlib.pyplot as plt
from matplotlib import rc
from matplotlib.ticker import MaxNLocator
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
# + papermill={"duration": 0.140641, "end_time": "2021-08-27T16:23:53.190304", "exception": false, "start_time": "2021-08-27T16:23:53.049663", "status": "completed"} tags=[]
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
sns.set(style='whitegrid', palette='muted', font_scale=1.2)
HAPPY_COLORS_PALETTE = ['#01BEFE', '#FFDD00', '#FF7D00', '#FF006D', '#ADFF02', '#8F00FF']
sns.set_palette(sns.color_palette(HAPPY_COLORS_PALETTE))
rcParams['figure.figsize'] = 16,10
# + papermill={"duration": 0.137491, "end_time": "2021-08-27T16:23:53.433229", "exception": false, "start_time": "2021-08-27T16:23:53.295738", "status": "completed"} tags=[]
data.head()
# + papermill={"duration": 0.124801, "end_time": "2021-08-27T16:23:53.663933", "exception": false, "start_time": "2021-08-27T16:23:53.539132", "status": "completed"} tags=[]
data.columns
# + papermill={"duration": 1.58083, "end_time": "2021-08-27T16:23:55.387587", "exception": false, "start_time": "2021-08-27T16:23:53.806757", "status": "completed"} tags=[]
data.describe()
# + papermill={"duration": 1.019817, "end_time": "2021-08-27T16:23:56.518949", "exception": false, "start_time": "2021-08-27T16:23:55.499132", "status": "completed"} tags=[]
plt.plot(data['label'])
plt.show()
# + papermill={"duration": 0.486874, "end_time": "2021-08-27T16:23:57.117545", "exception": false, "start_time": "2021-08-27T16:23:56.630671", "status": "completed"} tags=[]
label_1 = data[data['label'] == 1 ]
label_1.describe()
# + papermill={"duration": 0.315013, "end_time": "2021-08-27T16:23:57.545165", "exception": false, "start_time": "2021-08-27T16:23:57.230152", "status": "completed"} tags=[]
label_2 = data[data['label'] == 2 ]
label_2.describe()
# + papermill={"duration": 0.25127, "end_time": "2021-08-27T16:23:57.905389", "exception": false, "start_time": "2021-08-27T16:23:57.654119", "status": "completed"} tags=[]
label_3 = data[data['label'] == 3 ]
label_3.describe()
# + papermill={"duration": 0.146899, "end_time": "2021-08-27T16:23:58.168171", "exception": false, "start_time": "2021-08-27T16:23:58.021272", "status": "completed"} tags=[]
label_3
# + papermill={"duration": 0.19425, "end_time": "2021-08-27T16:23:58.474443", "exception": false, "start_time": "2021-08-27T16:23:58.280193", "status": "completed"} tags=[]
label_3.columns
# + papermill={"duration": 0.122351, "end_time": "2021-08-27T16:23:58.708736", "exception": false, "start_time": "2021-08-27T16:23:58.586385", "status": "completed"} tags=[]
sns.set()
# + papermill={"duration": 0.173855, "end_time": "2021-08-27T16:23:58.998450", "exception": false, "start_time": "2021-08-27T16:23:58.824595", "status": "completed"} tags=[]
label_1.count()[0], label_2.count()[0], label_3.count()[0]
# + papermill={"duration": 0.149229, "end_time": "2021-08-27T16:23:59.263088", "exception": false, "start_time": "2021-08-27T16:23:59.113859", "status": "completed"} tags=[]
label_1
# + [markdown] papermill={"duration": 0.114838, "end_time": "2021-08-27T16:23:59.494543", "exception": false, "start_time": "2021-08-27T16:23:59.379705", "status": "completed"} tags=[]
# Amout of data available
# + papermill={"duration": 0.180658, "end_time": "2021-08-27T16:23:59.790318", "exception": false, "start_time": "2021-08-27T16:23:59.609660", "status": "completed"} tags=[]
label_1.count()[0]/700, label_2.count()[0]/700, label_3.count()[0]/700
# + [markdown] papermill={"duration": 0.116161, "end_time": "2021-08-27T16:24:00.019678", "exception": false, "start_time": "2021-08-27T16:23:59.903517", "status": "completed"} tags=[]
# ### Run only for S4,S7 and ignore the warning
# As it has ```700x+1``` data in label 1, 3 respectively
# + papermill={"duration": 0.122981, "end_time": "2021-08-27T16:24:00.255758", "exception": false, "start_time": "2021-08-27T16:24:00.132777", "status": "completed"} tags=[]
# only for S4 ( it has one extra data)
if file_num == 4 :
n = 1
label_1.drop(label_1.tail(n).index,inplace=True)
if file_num == 7 :
n = 1
label_3.drop(label_3.tail(n).index,inplace=True)
# + papermill={"duration": 0.180022, "end_time": "2021-08-27T16:24:00.562321", "exception": false, "start_time": "2021-08-27T16:24:00.382299", "status": "completed"} tags=[]
label_1.count()[0]/700, label_2.count()[0]/700, label_3.count()[0]/700
# + [markdown] papermill={"duration": 0.114646, "end_time": "2021-08-27T16:24:01.258472", "exception": false, "start_time": "2021-08-27T16:24:01.143826", "status": "completed"} tags=[]
# ----------------------------------------------------------------------------------------------------
# + [markdown] papermill={"duration": 0.113122, "end_time": "2021-08-27T16:24:01.484753", "exception": false, "start_time": "2021-08-27T16:24:01.371631", "status": "completed"} tags=[]
# ### Code To Generate Sequence Data
# + [markdown] papermill={"duration": 0.113741, "end_time": "2021-08-27T16:24:01.712691", "exception": false, "start_time": "2021-08-27T16:24:01.598950", "status": "completed"} tags=[]
# - Given **Sampling Rate = 700 Hz**. Therefore we have 700 samples per second
# - Now, for 1 sample represent 1/700s <br/>
# - Thus *x* samples represent *x/700s*
#
# -----------------------------
#
# - We have total data of 2121 seconds ie approximately 35 minutes for this participant (for S2)
# - To know the code for particular subject see the cell below
# + papermill={"duration": 0.173592, "end_time": "2021-08-27T16:24:02.000068", "exception": false, "start_time": "2021-08-27T16:24:01.826476", "status": "completed"} tags=[]
# We have total data of
cnt = (label_1.count() + label_2.count() + label_3.count() )[0]/700
cnt
# + papermill={"duration": 0.126549, "end_time": "2021-08-27T16:24:02.242040", "exception": false, "start_time": "2021-08-27T16:24:02.115491", "status": "completed"} tags=[]
cnt/60
# + [markdown] papermill={"duration": 0.113784, "end_time": "2021-08-27T16:24:02.468703", "exception": false, "start_time": "2021-08-27T16:24:02.354919", "status": "completed"} tags=[]
# ## Appending the data frames
# + papermill={"duration": 0.122216, "end_time": "2021-08-27T16:24:02.704370", "exception": false, "start_time": "2021-08-27T16:24:02.582154", "status": "completed"} tags=[]
seq_data = label_1
# + papermill={"duration": 0.414099, "end_time": "2021-08-27T16:24:03.233432", "exception": false, "start_time": "2021-08-27T16:24:02.819333", "status": "completed"} tags=[]
seq_data = seq_data.append(label_2).append(label_3)
# + papermill={"duration": 0.165082, "end_time": "2021-08-27T16:24:03.513545", "exception": false, "start_time": "2021-08-27T16:24:03.348463", "status": "completed"} tags=[]
seq_data.count()[0]/700
# + papermill={"duration": 0.140121, "end_time": "2021-08-27T16:24:03.770296", "exception": false, "start_time": "2021-08-27T16:24:03.630175", "status": "completed"} tags=[]
seq_data.head()
# + papermill={"duration": 0.737266, "end_time": "2021-08-27T16:24:04.623729", "exception": false, "start_time": "2021-08-27T16:24:03.886463", "status": "completed"} tags=[]
seq_data.describe()
# -
# # High Class Imbalance
# + papermill={"duration": 0.654451, "end_time": "2021-08-27T16:24:05.436444", "exception": false, "start_time": "2021-08-27T16:24:04.781993", "status": "completed"} tags=[]
seq_data.label.value_counts().plot(kind='bar')
plt.xticks(rotation=45)
plt.show()
# + papermill={"duration": 0.128521, "end_time": "2021-08-27T16:24:05.681932", "exception": false, "start_time": "2021-08-27T16:24:05.553411", "status": "completed"} tags=[]
len(seq_data)/700
# + papermill={"duration": 3.764804, "end_time": "2021-08-27T16:24:09.565014", "exception": false, "start_time": "2021-08-27T16:24:05.800210", "status": "completed"} tags=[]
seq = np.array([], dtype='int')
for i in range(int(len(seq_data)/700)) :
temp = np.ones(shape = 700,dtype='int')*i
seq = np.append(seq,temp)
# + papermill={"duration": 0.126728, "end_time": "2021-08-27T16:24:09.814008", "exception": false, "start_time": "2021-08-27T16:24:09.687280", "status": "completed"} tags=[]
seq.shape
# + papermill={"duration": 0.129576, "end_time": "2021-08-27T16:24:10.061496", "exception": false, "start_time": "2021-08-27T16:24:09.931920", "status": "completed"} tags=[]
seq
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
# ## Preprocessing
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
# converting labels to 0,1,2
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
seq_data.head()
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
seq_data['label'] -= 1
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
# Creating X and Y
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
y_train = pd.DataFrame()
y_train['series_id'] = seq_data['series_id']
y_train['label'] = seq_data['label']
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
# The Lables are already integers so we dont need to encode them
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
FEATURE_COLUMNS = seq_data.columns.tolist()[1:]
FEATURE_COLUMNS
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
(seq_data.series_id.value_counts() == 700 ).sum() == len(y_train)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
y_train = y_train.groupby(['series_id']).mean()
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
temp = range(int(len(seq_data)/700))
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
y_train['series_id'] = temp
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
# ## ASSERT STATEMENT
# Don't proceed if the following cell outputs False
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
(seq_data.series_id.value_counts() == 700 ).sum() == len(y_train)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
seq_data
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
name = 'S{}_train.csv'.format(file_num)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
seq_data.to_csv(name)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
y_train
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
tname = 'S{}_label.csv'.format(file_num)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "pending"} tags=[]
y_train.to_csv(tname)
# -
# From the two cells above I have used the cleaned output data
# Authored By - <NAME> ( <EMAIL> )
| train_dir/data_cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Select patients for CFR model: Split patients in train, val and test sets ###
# +
import os
import glob
import numpy as np
import pandas as pd
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
# -
cfr_data_root = os.path.normpath('/mnt/obi0/andreas/data/cfr')
pet_data_dir = os.path.normpath('/mnt/obi0/phi/pet/pet_cfr')
meta_date = '200617'
meta_dir = os.path.join(cfr_data_root, 'metadata_'+meta_date)
print(meta_dir)
# +
match_view_filename = 'pet_match365_diff_files_'+meta_date+'.parquet'
files_cfr = pd.read_parquet(os.path.join(meta_dir, match_view_filename))
print('Total number of patients {}'.format(len(files_cfr.mrn.unique())))
print('Total number of echo studies {}'.format(len(files_cfr.study.unique())))
print('Total number of PET studies {}'.format(len(files_cfr.petmrn_identifier.unique())))
print('Total number of echos {}'.format(len(files_cfr.filename.unique())))
files_cfr.head(2)
# -
# ### Filter data sets: GLOBAL and NON-DEFECT variables ###
# ### Global variables ###
#
# Notebook 3/17/2020: global_pet_cfr
# File used: pet_cfr_petdata_02_26_2020_withperfandseg7.xlsx
#
# As described above, 2871 after excluding CABG, transplant, and those with missing perfusion data
# File used: post_2018_pets_with_clinical_cfr_all.csv
#
# After excluding CABG, transplant, and missing CFR values, 167 PETs with any perfusion
# Merge:
#
# After combining, 3038 PETs
# File saved as pets_with_echos_global_all.parquet
# Use notes:
#
# NOTE- there are petmrn_identifiers that have two rows- the one with post_2018==0 is the one to use, and the one with post_2018==1 should be excluded
# Variables to use: rest_global_mbf, stress_global_mbf, global_cfr_calc
# Other variables- myocardial_perfusion, segmental data, perfusion data, TID, gated SPECT results, calcium score, height, weight
# For "cleaner" data, would exclude those with post-2018==1
#
# #### Update 6/14/2020 ####
#
# Created a revised version of the above that includes CABG cases (from pre-2018), and excludes post-2018 cases that are already in the pre-2018 dataset (n=7). Total 3718 rows. File at /mnt/obi0/phi/pet/pet_cfr/pets_with_echos_global_all_withcabg.parquet
# Update 6/17/2020:
#
# Created a revised version that has a column tracer_obi for the tracer used. Also, the 7 post-2018 duplicate cases are excluded so there are are 3031 studies.
# File used- aiCohort_withPerfusion_addRadiopharm.xlsx
# The ammonia cases before 7/25/2011 have the value 'listed as ammonia', and the rubidium cases after 7/25/2011 have the value 'listed as rubidium. For the remaining discrepant values (i.e. missing values, FDG, sestamibi), the tracer was assumed to be the tracer in use at the time.
# Rubidium 1,740, ammonia 1,276, listed as ammonia 4, listed as rubidium 2
# File saved as /mnt/obi0/phi/pet/pet_cfr/pets_with_echos_global_all_withtracer.parquet
global_pet_file = 'pets_with_echos_global_all_withtracer.parquet'
global_pet = pd.read_parquet(os.path.join(pet_data_dir, global_pet_file))
global_pet = global_pet.astype({'pet_date': 'datetime64[ns]'})
print(f'PET studies: {len(global_pet.petmrn_identifier.unique())}')
print(os.path.join(pet_data_dir, global_pet_file))
global_pet.head()
global_pet[global_pet.petmrn_identifier=='1414556_2018-10-30']
# +
# Exclude files without frame_time
file_cfr_meta = files_cfr.loc[~files_cfr.frame_time.isnull()]
# Add echo data to the pet studies (inner join, to keep only keys in both dataframes)
global_pet_echo = global_pet.merge(file_cfr_meta, on = ['mrn', 'pet_date', 'petmrn_identifier'], how='inner')
print(f'PET studies: {len(global_pet_echo.petmrn_identifier.unique())}')
print(f'ECHO studies: {len(global_pet_echo.study.unique())}')
# +
print(f'All data: patients: {len(file_cfr_meta.mrn.unique())}')
print(f'All data: PET studies: {len(file_cfr_meta.petmrn_identifier.unique())}')
print(f'All data: ECHO studies: {len(file_cfr_meta.study.unique())}')
print(f'All data: videos: {len(file_cfr_meta.filename.unique())}')
print()
print(f'Global data: patients: {len(global_pet_echo.mrn.unique())}')
print(f'Global data: PET studies: {len(global_pet_echo.petmrn_identifier.unique())}')
print(f'Global data: ECHO studies: {len(global_pet_echo.study.unique())}')
print(f'Global data: videos: {len(global_pet_echo.filename.unique())}')
excluded_pet_studies = len(file_cfr_meta.petmrn_identifier.unique()) - len(global_pet_echo.petmrn_identifier.unique())
print(f'Excluded PET studies: {excluded_pet_studies}')
# +
# Find out which echo studies do not have a4c views
# Get all studies WITH a4c views
a4c_study_list = list(global_pet_echo[global_pet_echo.max_view=='a4c'].study.unique())
print(len(a4c_study_list))
# Remove all studies from global list that have a4c views
global_pet_echo_no_a4c = global_pet_echo[~global_pet_echo.study.isin(a4c_study_list)]
print(f'Number of ECHO studies without a4c view: {len(global_pet_echo_no_a4c.study.unique())}')
print(f'For this number of patients: {len(global_pet_echo_no_a4c.mrn.unique())}')
print(f'With this number of videos: {len(global_pet_echo_no_a4c.filename.unique())}')
# +
# Exclusions and filters
# A4C VIEW
global_pet_echo_a4c = global_pet_echo.loc[global_pet_echo.max_view=='a4c']
global_pet_echo_a4c = global_pet_echo_a4c.loc[~global_pet_echo_a4c.frame_time.isnull()]
global_a4c_pre18 = global_pet_echo_a4c.loc[global_pet_echo_a4c['post-2018']==0]
print(f'Global data: patients: {len(global_pet_echo.mrn.unique())}')
print(f'Global data: PET studies: {len(global_pet_echo.petmrn_identifier.unique())}')
print(f'Global data: ECHO studies: {len(global_pet_echo.study.unique())}')
print(f'Global data: videos: {len(global_pet_echo.filename.unique())}')
print()
print('After a4c and post-2018 filters:')
print(f'Global data: patients: {len(global_a4c_pre18.mrn.unique())}')
print(f'Global data: PET studies: {len(global_a4c_pre18.petmrn_identifier.unique())}')
print(f'Global data: ECHO studies: {len(global_a4c_pre18.study.unique())}')
print(f'Global data: videos: {len(global_a4c_pre18.filename.unique())}')
print()
print(f'Lost {len(global_pet_echo_a4c.filename.unique())-len(global_a4c_pre18.filename.unique())} due to post-2018 filtering.')
# +
# Global_pet_echo table with the variables to use (drop rows with na in any of those variables)
global_pet_echo.head(2)
global_pet_variables_target = ['rest_global_mbf',
'stress_global_mbf',
'global_cfr_calc',
'post-2018',
'tracer_obi']
global_pet_variables = global_pet_variables_target.copy()
global_pet_variables.extend(list(files_cfr.columns))
global_pet_echo_variables = global_a4c_pre18[global_pet_variables].dropna(subset=global_pet_variables_target,
axis=0)
print(f'Global data: patients: {len(global_pet_echo_variables.mrn.unique())}')
print(f'Global data: PET studies: {len(global_pet_echo_variables.petmrn_identifier.unique())}')
print(f'Global data: ECHO studies: {len(global_pet_echo_variables.study.unique())}')
print(f'Global data: videos: {len(global_pet_echo_variables.filename.unique())}')
print(f'Tracer values: {len(global_pet_echo_variables.tracer_obi.unique())}')
# -
# Complete list of unique petmrn_identifier
petmrn_identifier_list = list(global_pet.petmrn_identifier.unique())
petmrn_identifier_set = list(set(petmrn_identifier_list))
# +
# Let's filter Rahuls list of missing echos
missing_echo = pd.read_parquet(os.path.join(meta_dir, 'mrn_pet_missing_echo_file.parquet'))
print(f'petmrn_identifier in original list: {len(missing_echo.petmrn_identifier.unique())}')
missing_echo.tail()
missing_echo_filtered = missing_echo[missing_echo.petmrn_identifier.isin(petmrn_identifier_set)].\
drop(columns=['pet_measurement'], axis=1).reset_index(drop=True)
print(f'petmrn_identifier in filtered list: {len(missing_echo_filtered.petmrn_identifier.unique())}')
missing_no_echo_date = missing_echo_filtered.loc[missing_echo_filtered.echo_date.isnull()]
print(f'petmrn_identifier without echo dates: {len(missing_no_echo_date.petmrn_identifier.unique())}')
missing_echo_filtered_file = 'mrn_pet_missing_echo_file_filtered.parquet'
missing_echo_filtered.to_parquet(os.path.join(meta_dir, missing_echo_filtered_file))
# -
# ### Split the patients in train, validate and test sets ###
# Although each view might have a little different patient pupulation distribution, because not all views are in each study. However, we want the same MRNS in each data set and for all views so that we can directly compare the performance of the algorithm for the same patients. We can expand the data frame above to add the splits.
def patientsplit(patient_list):
train_test_split = 0.85
train_eval_split = 0.90
# Take a test set from all patients
patient_list_train = np.random.choice(patient_list,
size = int(np.floor(train_test_split*len(patient_list))),
replace = False)
patient_list_test = list(set(patient_list).difference(patient_list_train))
train_test_intersection = set(patient_list_train).intersection(set(patient_list_test)) # This should be empty
print('Intersection of patient_list_train and patient_list_test:', train_test_intersection)
# Further separate some patients for evaluation from the training list
patient_list_eval = np.random.choice(patient_list_train,
size = int(np.ceil((1-train_eval_split)*len(patient_list_train))),
replace = False)
patient_list_train = set(patient_list_train).difference(patient_list_eval)
train_eval_intersection = set(patient_list_train).intersection(set(patient_list_eval))
print('Intersection of patient_list_train and patient_list_eval:', train_eval_intersection)
# Show the numbers
print('total patients:', len(patient_list))
print()
print('patients in set:', np.sum([len(patient_list_train),
len(patient_list_eval),
len(patient_list_test)]))
print()
print('patients in train:', len(patient_list_train))
print('patients in eval:', len(patient_list_eval))
print('patients in test:', len(patient_list_test))
return patient_list_train, patient_list_eval, patient_list_test
# +
dataset = global_pet_echo_variables
dataset_filename = 'global_pet_echo_dataset_'+meta_date+'.parquet'
global_pet_echo_variables.head()
# Get a patient list
patient_list = list(dataset.sample(frac=1).mrn.unique())
patient_list_train, patient_list_eval, patient_list_test = patientsplit(patient_list)
patient_split = {'train': patient_list_train,
'eval': patient_list_eval,
'test': patient_list_test}
print('Patient IDs in train:', len(patient_split['train']))
print('Patient IDs in eval:', len(patient_split['eval']))
print('Patient IDs in test:', len(patient_split['test']))
print()
print('Intersection train-test:', set(patient_split['train']).intersection(set(patient_split['test'])))
print('Intersection train-eval:', set(patient_split['train']).intersection(set(patient_split['eval'])))
print('Intersection eval-test:', set(patient_split['eval']).intersection(set(patient_split['test'])))
# +
# Add mode column to files_cfr
split_list = []
for dset in patient_split.keys():
dset_mrn_list = list(patient_split[dset])
split_list.append(pd.DataFrame({'mrn': dset_mrn_list,
'dset_mode': [dset]*len(dset_mrn_list)}))
split_df = pd.concat(split_list, ignore_index = True)
dataset_split = dataset.merge(right = split_df, on = 'mrn', how = 'left').\
sample(frac = 1).\
reset_index(drop = True)
# +
print(f'TOTAL patients: {len(dataset_split.mrn.unique())}')
print(f'TOTAL PET studies: {len(dataset_split.petmrn_identifier.unique())}')
print(f'TOTAL ECHO studies: {len(dataset_split.study.unique())}')
print(f'TOTAL videos: {len(dataset_split.filename.unique())}')
dset_list = ['train', 'eval', 'test']
for dset in dset_list:
df_dset = dataset_split[dataset_split.dset_mode==dset]
print()
print(f'patients in {dset}: {len(df_dset.mrn.unique())}')
print(f'PET studies in {dset}: {len(df_dset.petmrn_identifier.unique())}')
print(f'ECHO studies in {dset}: {len(df_dset.study.unique())}')
print(f'videos in {dset}: {len(df_dset.filename.unique())}')
# +
# Load actual video files after TFR conversion
tfr_dir = os.path.join(cfr_data_root, 'tfr_200617', 'cfr')
train_files = sorted(glob.glob(os.path.join(tfr_dir, 'cfr_a4c_train_200617_*.parquet')))
train_files_failed = sorted(glob.glob(os.path.join(tfr_dir, 'cfr_a4c_train_200617_*.failed')))
eval_files = sorted(glob.glob(os.path.join(tfr_dir, 'cfr_a4c_eval_200617_*.parquet')))
test_files = sorted(glob.glob(os.path.join(tfr_dir, 'cfr_a4c_test_200617_*.parquet')))
train_df = pd.concat([pd.read_parquet(file) for file in train_files])
eval_df = pd.concat([pd.read_parquet(file) for file in eval_files])
test_df = pd.concat([pd.read_parquet(file) for file in test_files])
df_tf = pd.concat([train_df, eval_df, test_df]).reset_index(drop=True)
# Lets look at the failed training files
train_failed_df = pd.concat([pd.read_parquet(file) for file in train_files_failed])
print((train_failed_df.err.unique()))
dset_list = ['train', 'eval', 'test']
for dset in dset_list:
df_dset = df_tf[df_tf.dset_mode==dset]
print()
print(f'patients in {dset}: {len(df_dset.mrn.unique())}')
print(f'PET studies in {dset}: {len(df_dset.petmrn_identifier.unique())}')
print(f'ECHO studies in {dset}: {len(df_dset.study.unique())}')
print(f'videos in {dset}: {len(df_dset.filename.unique())}')
# -
train_failed_df[(train_failed_df.err=='deltaXY') & (train_failed_df.deltaX<0)].shape
# Let's look at the video tracer numbers
tracer_list = dataset_split.tracer_obi.unique()
print(tracer_list)
for tracer in tracer_list:
df_tracer = dataset_split[dataset_split.tracer_obi==tracer]
for dset in dset_list:
df_dset = df_tracer[df_tracer.dset_mode==dset]
print()
print(f'{dset}: patients for {tracer}: {len(df_dset.mrn.unique())}')
print(f'{dset}: PET studies for {tracer}: {len(df_dset.petmrn_identifier.unique())}')
print(f'{dset}: ECHO studies for {tracer}: {len(df_dset.study.unique())}')
print(f'{dset}: videos for {tracer}: {len(df_dset.filename.unique())}')
dataset_split.columns
# +
# Prepare the final set that we will use for writing TFR files. We don't want any rows with NAs in some columns.
col_set = ['rest_global_mbf', 'rest_global_mbf', 'global_cfr_calc', 'tracer_obi',
'pet_measurement', 'sum_views', 'dset_mode']
dataset_split_tfr = dataset_split.dropna(subset = col_set, axis=0)
print('Dropped {} rows.'.format(dataset_split.shape[0]-dataset_split_tfr.shape[0]))
# Add some other information that we need and shuffle the whole thing
dataset_split_tfr = dataset_split_tfr.assign(rate = np.round(1/dataset_split_tfr.frame_time*1e3, decimals = 1))
dataset_split_tfr = dataset_split_tfr.assign(dur = dataset_split_tfr.frame_time*1e-3*dataset_split_tfr.number_of_frames)
dataset_split_tfr = dataset_split_tfr.sample(frac = 1)
dataset_split.loc[~dataset_split.index.isin(dataset_split_tfr.index)]
# -
print(list(dataset_split_tfr.tracer_obi.unique()))
dataset_split_tfr.to_parquet(os.path.join(meta_dir, dataset_filename))
print('Saved to file:', dataset_filename)
print(dataset_split_tfr.shape)
dataset_split_tfr.head()
# +
minframes = 40
max_frame_time = 33.34
print('Minimum frames: {}'.format(minframes))
print('Maximum frame_time: {}'.format(max_frame_time))
minduration = max_frame_time*minframes*1e-3
print('Minimum duration: {}'.format(minduration))
#minframes = int(np.ceil(minrate*minduration))
maxrows = dataset_split_tfr.shape[0]
dataset_disqualified = dataset_split_tfr[(dataset_split_tfr.frame_time > max_frame_time) | (dataset_split_tfr.dur < minduration)]
dataset_qualified = dataset_split_tfr[(dataset_split_tfr.frame_time <= max_frame_time) & (dataset_split_tfr.dur >= minduration)]
n_videos = len(dataset_split_tfr.filename.unique())
n_disqualified = len(dataset_disqualified.filename.unique())
n_qualified = len(dataset_qualified.filename.unique())
print()
print('Total videos: {}'.format(n_videos))
print('Disqualified videos {}, fraction:{:.1f}'.format(n_disqualified,
np.round(n_disqualified/n_videos*100), decimals=1))
print('Qualified videos {}, fraction:{:.1f}'.format(n_qualified,
np.round(n_qualified/n_videos*100), decimals=1))
# -
# Report train val test numbers that qualify....
tracer_list = dataset_split.tracer_obi.unique()
print(tracer_list)
for dset in dset_list:
df_dset = dataset_qualified[dataset_qualified.dset_mode==dset]
print()
print(f'{dset}: patients for TOTAL: {len(df_dset.mrn.unique())}')
print(f'{dset}: PET studies for TOTAL: {len(df_dset.petmrn_identifier.unique())}')
print(f'{dset}: ECHO studies for TOTAL: {len(df_dset.study.unique())}')
print(f'{dset}: videos for TOTAL: {len(df_dset.filename.unique())}')
for tracer in tracer_list:
df_tracer = df_dset[df_dset.tracer_obi==tracer]
print()
print(f'{dset}: patients for {tracer}: {len(df_dset.mrn.unique())}')
print(f'{dset}: PET studies for {tracer}: {len(df_dset.petmrn_identifier.unique())}')
print(f'{dset}: ECHO studies for {tracer}: {len(df_dset.study.unique())}')
print(f'{dset}: videos for {tracer}: {len(df_dset.filename.unique())}')
| notebooks/cfr_datasets_patientsplit_global.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <ul>
# <div style="text-align: LEFT"> <div style="color:BLUE"> <font size="13"> <p><b>FUNCTIONS PROGRAMS</b></p></div> </div></font>
# <div style="color:white"> <font size="12">
# </div></font>
# </ul>
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>1.program to <strong>find the sum of two numbers</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to find the sum of two numbers
a=int(input("ENTER THE FIRST NUMBER"))#prompting the user for input and performing type-casting to convert the input from
b=int(input("ENTER THE SECOND NUMBER")) # string to int
def add():#function definition of add function
sum=a+b#adding two numbers and storing the result in the variable sum
return sum #returning the sum when the function is called
def main():#function definition of main function
result=add()#calling the function add and storing the returning value in the variable result
print(f"THE SUM OF TWO NUMBERS IS {result}")#printing the sum of two numbers
if(__name__=="__main__"):
main()#calling the main function
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>2.program to <strong>check whether the given number is even or odd</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to check whether the given number is even or odd
n=int(input("ENTER THE NUMBER"))#prompting the user for input and performing type-casting to convert the input from string to int
def even_or_odd():#function definition of the function even_or_odd
if(n%2==0):#conditional statement to check whether the number is divisible by 2
print(f"{n} is even")#printing the number is even if the condition is true
else:
print(f"{n} is odd")#printing the number is odd if the condition is false
def main():#function definition of the main function
even_or_odd()#calling the even_or_odd function
if(__name__=="__main__"):
main()#calling the main function
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>3.program to <strong>find the square of a number</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to find the square of a number
n=int(input("ENTER THE NUMBER"))#prompting the user for input and performing type casting to convert the input from string to int
def square():#function definition of the square function
result=n**2#performing the square of a number and storing it in the variable result
return result#returning the result
def main():#function definition of the main function
sq=square()#calling the square function and storing the returning value in variable square
print(f"THE SQUARE OF {n} IS {sq}")#printing the number and square of the number
if(__name__=="__main__"):
main()#calling the main function
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>4.program to <strong>swap two numbers</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to swap two numbers
x=int(input("ENTER THE 1st NUMBER"))#prompting the user for input and performing type-casting to convert the input from string to int
y=int(input("ENTER THE 2nd NUMBER"))
def swap(x,y):#function definition of the function swap with two arguments x and y
print(f"before swaping value of x is{x} and y is {y}")#printing the values of x and y before swapping
temp=x#swaping the values of x and y by taking one temp variable
x=y
y=temp
print(f"AFTER SWAPING VALUE OF x IS {x} AND y IS {y}")#printing the values of x and y after swapping
def main():#function definition of main function
swap(x,y)#calling the swap function by passing two arguments x and y
if(__name__=="__main__"):
main()#calling the main function
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>5.program to <strong>find the power of m rise to n</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to find the power of m rise to n
m=int(input("ENTER THE BASE NUMBER"))#prompting the user for input and performing type-casting to convert the user input from string to int
n=int(input("ENTER THE POWER TO WHICH THE NUMBER HAS TO BE RAISED"))
def power(m,n):#function definition of the power function with two arguments m and n
result=m**n#performing m power n and storing the result in variable result
return result#returning the result when the function is called
def main():#function definition of the main function
res=power(m,n)#calling the function power by passing 2 arguments m and n and storing the returning value in the variable res
print(f"THE RESULT IS {res}")#printing the result
if(__name__=="__main__"):
main()#calling the main function
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>6.program to <strong>find the number is niven</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to find the number is niven
#the number is niven if it is divisible by the sum of its digits
n=int(input("ENTER THE NUMBER"))#prompting the user for input
def check_niven(n):#function definition of the function check_niven with one argument n
temp=n#assigning the value of n to temp
sum=0#initialising the value of sum to 0
while(temp>0):#conditional statement to check the number is greater than 0
last_digit=temp%10#accessing the last digit of the number
sum=sum+last_digit#calculating the sum of digits
temp=temp//10#accessing the remaining numbers leavimg the last digit
if(n%sum==0):#checking whether the number is completely divisible by the sum of digits
print(f"{n} IS NIVEN NUMBER")#printing the number is niven if the condition is true
else:
print(f"{n} IS NOT NIVEN NUMBER")#printing the number is not niven if the condition is false
def main():#function definition of the main function
check_niven(n)#calling the function check_niven by passing one parameter n
if(__name__=="__main__"):
main()#calling main function
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>7.program to <strong>find the number is palindrome</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to check the number is palindrome
n=int(input("ENTER THE NUMBER"))#prompting the user for input and performing type-casting to convert the user input from string to int
def palindrome(n):#function definition of the function palindrome with one argument n
temp=n#assigning the value of n to variable temp
rn=0#initialising the value of rn to o
while(temp>0):#checking whether the number is greater than 0
last_digit=temp%10#accessing the last digit of the number
rn=rn*10+last_digit
temp=temp//10#accessing the number leaving the last digit
if(n==rn):#checking whether n is equal to rn
print(f"{n} IS PALINDROME")#printing the number is palindrome if the condition is true
else:
print(f"{n} IS NOT PALINDROME")#printing the number is not palindrome if the condition is not true
def main():#function definition of the main function
palindrome(n)#calling the function palindrome by passing one argument n
if(__name__=="__main__"):
main()#calling the main function
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>8.program to <strong>find the largest of 3 numbers</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to find the largest of 3 numbers
a=int(input("ENTER 1st NUMBER "))#prompting the user for input and performing type-casting to convert the user input from string to int
b=int(input("ENTER 2nd NUMBER"))
c=int(input("ENTER 3rd NUMBER"))
def largest():#function definition of the function largest
if(a>b and a>c):#checking whether a is greater than b and c
print(f"{a} is the largest")#printing a is the largest if the condition is true
elif(b>a and b>c):#checking whether b is greater than both a and c
print(f"{b} is the largest")#printing b is the largest if the condition is true
else:
print(f"{c} is the largest")#printing c is the largest if both the condition is not true
largest()#calling the function largest
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>9.program to <strong>Find the factors of the number</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 program to find the factors of the number
n=int(input("ENTER THE NUMBER"))#prompting the user for input and performing type-casting to convert the user input from string to int
def factors():#function definition of the function factors
print(f"THE FACTORS OF {n} ARE")#printing the factors of number are
for i in range (1,n):#looping statement to access all the values between 1 and the number
if(n%i==0):#checking whether n is divisible by i
print(i)#printing i if the condition is true
factors()#calling the function factors
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="text-align: LEFT"> <div style="color:BLUE"> <font size="13"> <p><b>STRINGS PROGRAMS</b></p></div> </div></font>
# <div style="color:white"> <font size="12">
# </div></font>
# </ul>
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>10.program to <strong>find the length of the string</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#program to find the length of the string
str=input("ENTER THE STRING")#prompting the user for input a string
count=0#initialising count variable to 0
for each_char in str:#loop construct for accessing each character in the string
count+=1#incrementing the value of count
print(f"THE NUMBER OF CHARACTERS IN {str} IS {count}")#printing the string and number of characters in it
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>11.program to <strong>reverse the string</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#program to reverse the string
str=input("ENTER THE STRING")#prompting the user to input a string
rev=str[::-1]#performing slicing operation to reverse the string and storing it in variable rev
print(f"THE REVERSED STRING IS {rev}")#printing the reversed string
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>12.program to <strong>print the even length words in the string</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python3 code to print the even length words in the string
s=input("ENTER THE STRING")#prompting the user to input a string
def printWords(s):#function definition of the function printWords
s=s.split()#spliting the string
for word in s:#iterating in words of string
if len(word)%2==0:#checking if the length is even
print(word)#printing the word if the condition is true
printWords(s)#calling the function printWords
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="text-align: LEFT"> <div style="color:BLUE"> <font size="13"> <p><b>LISTS PROGRAMS</b></p></div> </div></font>
# <div style="color:white"> <font size="12">
# </div></font>
# </ul>
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>13.program to <strong>find the sum of elements in the list</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python program to find the sum of elements in the list
def sum_list(items):#function definition of the function sum_list with one argument items(here items is the list)
sum=0#initialising sum to 0
for i in items:#loop construct to iterate through each element in the list
sum+=i#computing sum is equal to sum + each element i
print(f"the sum of elements is {sum}")#printing the sum
sum_list([1,3,6])#calling the function sum_list by passing list as the argument
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>14.program to <strong>find the product of the elements in the list</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python program to find the product of the elements in the list
def product(items):#function definition of the function product with one list argument
pro=1#initialising the value of pro to 1
for i in items:#iterating through each element in the list
pro=pro*i#computing product
print(f"the product is {pro}")#printing the product
product([1,2,3])#calling the function product by passing list as the argument
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>15.program to <strong>find the largest element in the list</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python program to find the largest element in the list
def largest(a):#function definition of the function largest with one list argument
large=a[0]#initialising large value with the first element of the list
for i in a:#iterating through eacg element in the list
if(i>large):#checking whether i is greater than the large element
large=i#updating the value of large to i if the condition is true
print(f"the largest element is {large}")#printing the largest element
largest([1,4,8])#calling the function largest by passing list as the argument
# + [markdown] pycharm={"name": "#%% md\n"}
# <ul>
# <div style="color:BLACK"> <font size="12">
# <li>16.program to <strong>find the smallest element in the list</strong></li>
# </div></font>
# </ul>
# + pycharm={"name": "#%%\n"}
#python program to find the smallest element in the list
def smallest(a):#function definition of the function smallest with one list argument
small=a[0]#initialising small value with the first element of the list
for i in a:#iterating through each element in the list
if(i<small):#checking whether i is lesser than the small element
small=i#updating the value of small to i if the condition is true
print(f"the smallest element is {small}")#printing the largest element
smallest([1,4,8])#calling the function smallest by passing list as the argument
| DAY2-AR-FSLA-PROGRAMS WITH SOLUTION.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 循环
# - 循环是一种控制语句块重复执行的结构
# - while 适用于广度遍历
# - for 开发中经常使用
# ## while 循环
# - 当一个条件保持真的时候while循环重复执行语句
# - while 循环一定要有结束条件,否则很容易进入死循环
# - while 循环的语法是:
#
# while loop-contunuation-conndition:
#
# Statement
i = 0
while i<10:
print('hahaha')
i += 1
# ## 示例:
# sum = 0
#
# i = 1
#
# while i <10:
#
# sum = sum + i
# i = i + 1
# ## 错误示例:
# sum = 0
#
# i = 1
#
# while i <10:
#
# sum = sum + i
#
# i = i + 1
# - 一旦进入死循环可按 Ctrl + c 停止
# ## EP:
# 
# 
# # 验证码
# - 随机产生四个字母的验证码,如果正确,输出验证码正确。如果错误,产生新的验证码,用户重新输入。
# - 验证码只能输入三次,如果三次都错,返回“别爬了,我们小网站没什么好爬的”
# - 密码登录,如果三次错误,账号被锁定
#
import random
n = random.randint(65,122)
N = ""
i = 0
while 1:
if 91<=n<=96:
n = random.randint(65,122)
else:
N += chr(n)
n = random.randint(65,122)
i += 1
if i == 4:
break
print(N)
count = 0
for i in range(1000):
a = random.randint(0,1000) / 1000
if 0<a<0.001
# ## 尝试死循环
# ## 实例研究:猜数字
# - 你将要编写一个能够随机生成一个0到10之间的且包括两者的数字程序,这个程序
# - 提示用户连续地输入数字直到正确,且提示用户输入的数字是过高还是过低
# ## 使用哨兵值来控制循环
# - 哨兵值来表明输入的结束
# - 
# ## 警告
# 
# ## for 循环
# - Python的for 循环通过一个序列中的每个值来进行迭代
# - range(a,b,k), a,b,k 必须为整数
# - a: start
# - b: end
# - k: step
# - 注意for 是循环一切可迭代对象,而不是只能使用range
for i in range(100):
print('Joker is a better man!')
a = 100
bb = 'JOker'
bb.__iter__()
c = [1,2,3]
c.__iter__
{'key':'value'}.__iter__
(1,3,43).__iter__
{1,2,43}.__iter__
for i in range(5):
print(i)
# # 在Python里面一切皆对象
# ## EP:
# - 
i = 1
sum_ = 0
while sum_ < 10000:
sum_ += i
i += 1
print(sum_)
sum_ = 0
for i in range(1,10001):
sum_ += i
if sum_ > 10000:
break
print(sum_)
sum = 0
i = 0
while i < 1001:
sum = sum + i
i += 1
print(sum)
# ## 嵌套循环
# - 一个循环可以嵌套另一个循环
# - 每次循环外层时,内层循环都会被刷新重新完成循环
# - 也就是说,大循环执行一次,小循环会全部执行一次
# - 注意:
# > - 多层循环非常耗时
# - 最多使用3层循环
# ## EP:
# - 使用多层循环完成9X9乘法表
# - 显示50以内所有的素数
# ## 关键字 break 和 continue
# - break 跳出循环,终止循环
# - continue 跳出此次循环,继续执行
for i in range(1,10):
for j in range(1,i+1):
print(j,'X',i,'=',i*j,end=' ')
print()
# ## 注意
# 
# 
# # Homework
# - 1
# 
# +
zhengshu = 0
fushu = 0
sum_ = 0
cishu = 0
data = 1
while data !=0 :
data = eval(input(">>"))
if data > 0:
zhengshu += 1
if data < 0:
fushu += 1
sum_ += data
if data != 0:
cishu += 1
print(zhengshu)
print(fushu)
print(sum_ / cishu)
# -
# - 2
# 
# dorlla = 10000
# for i in range(14):
# dorlla = dorlla * 0.05 + dorlla
# if i == 9:
# print(dorlla)
# print(dorlla)
def dorlla(dorlla):
sum = 0
for i in range(14):
dorlla = dorlla * 0.05 + dorlla
if i > 9:
sum = sum + dorlla
if i == 9:
print('第十年学费为:%f'%dorlla)
print('十年后大学四年的总学费为:%f'%sum)
dorlla(10000)
# - 3
# 
# +
sum1 = 0
sum2 = 0
sum3 = 0
count = 0
def average(number):
global count
global sum1
global sum2
global sum3
sum3 = sum3 + number
if number != 0:
count +=1
if number > 0:
sum1 +=1
if number < 0:
sum2 +=1
else:
print('正数有%d个,负数有%d个,平均值为:%f'%(sum1,sum2,sum3 / count))
while True:
number = int(input('Enter an integer,the input ends if it is 0:'))
average(number)
if number == 0:
break
# -
# - 4
# 
def zhengchu():
count = 0
for i in range(100,1001):
if i % 5==0 and i % 6 == 0:
print(i,end=" ")
count += 1
if count % 10 == 0:
print()
zhengchu()
# - 5
# 
def dayu():
n = 0
while n **2 < 12000:
n +=1
print(n)
def xiaoyu():
n = 0
while n **3 < 12000:
n +=1
print(n-1)
dayu()
xiaoyu()
# - 6
# 
amount=eval(input('Loan Amount:'))
year=eval(input('Number of Years:'))
print('Interest Rate Monthly Payment Total Payment')
rate=5
while 1:
tpay=amount
tm=year*12
y=rate/100/12
mpay=(amount*y*(1+y)**tm)/((1+y)**tm-1)
tpay=mpay*tm
print('%.3f%% %.2f %.2f '%(rate,mpay,tpay))
rate=rate+1/8
if rate>8:
break
# - 7
# 
def he():
res = 0
for i in range(50000,0,-1):
res += 1/i
print(res)
he()
# - 8
# 
res = 0
for i in range(1,98,2):
res += i/ (i+2)
print(res)
# - 9
# 
# +
def pi():
for i in range(10000,100001,10000):
pi =0
for j in range(1,i+1):
pi = pi + ((-1) ** (j + 1)) / (2 * j - 1)
print( 4 * pi)
pi()
# -
# - 10
# 
def shu(a,b):
for i in range(a,b):
res = 0
for j in range(1,i):
if i % j == 0:
res += j
if i == res:
print(i)
shu(1,10000)
# - 11
# 
def zh():
count = 0
for i in range(1,8):
for j in range(i + 1,8):
print(i,j)
count +=1
print('The total number of all combinations is %d'%count)
zh()
# - 12
# 
import math
print('enter ten numbers:')
ls=[]
sum1=0.0
devi=0.0
for i in range(10):
num=eval(input(''))
ls.append(num)
part1=0.0
part2=0.0
for i in range(10):
sum1=sum1+ls[i]
mean=sum1/10
for i in range(10):
part1=part1+(ls[i]-mean)**2
#part2=sum1**2
#print(part1,part2)
devi=math.sqrt(part1/9)
print('The mean is %.2f'%(mean))
print('The standerd deviation %f'%(devi))
| 7.19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
'''
Let's make plots for 5 stem categories and describe percent of degrees granted to each gender during some period(in years)
We want to plot each degree-gender category percentage next to each other, all in one row
We want to make end gender gaps sorted in decreasing manner
'''
women_degrees = pd.read_csv('percent-bachelors-degrees-women-usa.csv')
#tuples that define color for each line (men and women) in RGB values
cb_dark_blue = (0/255,107/255,164/255)
cb_orange = (255/255, 128/255, 14/255)
stem_cats = ['Engineering', 'Computer Science', 'Psychology', 'Biology', 'Physical Sciences', 'Math and Statistics']
fig = plt.figure(figsize=(18, 3))
for sp in range(0,6):
ax = fig.add_subplot(1,6,sp+1)
ax.plot(women_degrees['Year'], women_degrees[stem_cats[sp]], c=cb_dark_blue, label='Women', linewidth=3)
ax.plot(women_degrees['Year'], 100-women_degrees[stem_cats[sp]], c=cb_orange, label='Men', linewidth=3)
#set all four spines(left,right,top,bottom) to unvisible to improve data-ink ratio
for key,value in ax.spines.items():
value.set_visible(False)
ax.set_xlim(1968, 2011)
ax.set_ylim(0,100)
ax.set_title(stem_cats[sp])
ax.tick_params(bottom="off", top="off", left="off", right="off")
if sp == 0:
ax.text(2005, 87, 'Men')
ax.text(2002, 8, 'Women')
elif sp == 5:
ax.text(2005, 62, 'Men')
ax.text(2001, 35, 'Women')
# +
stem_cats = ['Psychology', 'Biology', 'Math and Statistics', 'Physical Sciences'
, 'Computer Science', 'Engineering']
lib_arts_cats = ['Foreign Languages', 'English', 'Communications and Journalism', 'Art and Performance'
, 'Social Sciences and History']
other_cats = ['Health Professions', 'Public Administration', 'Education',
'Agriculture','Business', 'Architecture']
cb_dark_blue = (0/255,107/255,164/255)
cb_orange = (255/255, 128/255, 14/255)
cb_y_midline = (177/255,177/255,177/255)
figure = plt.figure(figsize=(9,20))
for i in range(0,len(stem_cats)):
ax = figure.add_subplot(6,3,i*3 + 1)
#plot line for women degree percentage
ax.plot(women_degrees['Year'], women_degrees[stem_cats[i]], c=cb_dark_blue, linewidth=3)
#plot line for men degree percentage ( 100 - women)
ax.plot(women_degrees['Year'], 100-women_degrees[stem_cats[i]], c=cb_orange, linewidth=3)
#we wont turn off labels on left side of plot for this column cuz its elements are first displayed in each row
#other columns elements have labels on left side off
ax.tick_params(bottom='off', top='off', left='off', right='off', labelbottom='off')
ax.set_title(stem_cats[i])
ax.set_xlim(1968,2011)
ax.set_ylim(0,100)
#draw mid line, where gender percentages are equal
ax.axhline(y=50, c=cb_y_midline, alpha=0.3)
# reduce y ticks(percentage) to just 2
ax.set_yticks([0,100])
# set spines to unvisible
for key,val in ax.spines.items():
val.set_visible(False)
if i == 0:
ax.text(2005, 80, 'Women')
ax.text(1998, 15, 'Man')
elif i == len(stem_cats)-1:
ax.text(2005, 72, 'Men')
ax.text(2004, 25, 'Women')
#we should turn x labels for the last row
ax.tick_params(labelbottom='on')
for i in range(0,len(lib_arts_cats)):
ax = figure.add_subplot(6,3,i*3 + 2)
#plot line for women degree percentage
ax.plot(women_degrees['Year'], women_degrees[lib_arts_cats[i]], c=cb_dark_blue, linewidth=3)
#plot line for men degree percentage ( 100 - women)
ax.plot(women_degrees['Year'], 100-women_degrees[lib_arts_cats[i]], c=cb_orange, linewidth=3)
ax.tick_params(bottom='off', top='off', left='off', right='off', labelbottom='off', labelleft='off')
ax.set_title(lib_arts_cats[i])
ax.set_xlim(1968,2011)
ax.set_ylim(0,100)
#draw mid line, where gender percentages are equal
ax.axhline(y=50, c=cb_y_midline, alpha=0.3)
# reduce y ticks(percentage) to just 2
ax.set_yticks([0,100])
for key,val in ax.spines.items():
val.set_visible(False)
if i == 0:
ax.text(2005, 80, 'Women')
ax.text(1998, 15, 'Men')
elif i==len(lib_arts_cats)-1:
#we should turn x labels for the last row
ax.tick_params(labelbottom='on')
for i in range(0,len(other_cats)):
ax = figure.add_subplot(6,3,i*3 + 3)
#plot line for women degree percentage
ax.plot(women_degrees['Year'], women_degrees[other_cats[i]], c=cb_dark_blue, linewidth=3)
#plot line for men degree percentage ( 100 - women)
ax.plot(women_degrees['Year'], 100-women_degrees[other_cats[i]], c=cb_orange, linewidth=3)
ax.tick_params(bottom='off', top='off', left='off', right='off', labelbottom='off',labelleft='off')
ax.set_title(other_cats[i])
ax.set_xlim(1968,2011)
ax.set_ylim(0,100)
#draw mid line, where gender percentages are equal
ax.axhline(y=50, c=cb_y_midline, alpha=0.3)
# reduce y ticks(percentage) to just 2
ax.set_yticks([0,100])
for key,val in ax.spines.items():
val.set_visible(False)
if i == 0:
ax.text(2005, 75, 'Women')
ax.text(2001, 20, 'Men')
elif i == len(other_cats)-1:
ax.text(2005, 72, 'Men')
ax.text(2004, 25, 'Women')
#we should turn x labels for the last row
ax.tick_params(labelbottom='on')
# export plots show on the figure to png file
figure.savefig('gender_degrees.png')
# -
| data-science/data_visualization/Visualizing The Gender Gap In College Degrees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import matplotlib.pyplot as plt
import joblib
import numpy as np
import pandas as pd
plt.style.use("fivethirtyeight")
# -
# !pip install joblib
class Perceptron:
def __init__(self,eta: float=None,epochs: int=None):
self.weights = np.random.randn(3) * 1e-4
self.eta=eta #Learning rate
self.epochs=epochs #number of interations
def _z_outcome(self,inputs,weights):
return np.dot(inputs,weights) #matrix multiplication
def activation_function(self,z):
return np.where(z>0,1,0)
def fit(self, X, y):
self.X=X
self.y=y
X_with_bias = np.c_[self.X, -np.ones((len(self.X),1))]
print(f"X with bias: \n{X_with_bias}")
for epochs in range(self.epochs):
print("--"*10)
print(f"for epoch >> {epochs +1}")
print("--"*10)
z= self._z_outcome(X_with_bias,self.weights)
y_hat = self.activation_function(z)
print(f"predicted value after forward pass: \n{y_hat}")
self.error =self.y - y_hat
print(f"error: \n{self.error}")
self.weights = self.weights + self.eta * np.dot(X_with_bias.T,self.error)
print(f"updated weights after epoch: {epochs +1}/{self.epochs} : \n{self.weights}")
print("##"*10)
def predict(self, X):
X_with_bias = np.c_[X, -np.ones((len(X),1))]
z=self._z_outcome(X_with_bias,self.weights)
return self.activation_function(z)
# +
OR = {
"x1":[0,0,1,1],
"x2":[0,1,0,1],
"y":[0,1,1,1]
}
df_OR=pd.DataFrame(OR)
df_OR
# -
def prepare_data(df, target_col="y"):
X= df.drop(target_col, axis=1)
y=df[target_col]
return X,y
X,y =prepare_data(df_OR)
X
y
# +
X,y =prepare_data(df_OR)
ETA=0.1
EPOCHS=10
model_or = Perceptron(eta=ETA,epochs=EPOCHS)
model_or.fit(X,y)
# -
model_or.predict(X)
model_or.predict(X=[[1,0]])
# +
AND = {
"x1":[0,0,1,1],
"x2":[0,1,0,1],
"y":[0,0,0,1]
}
df_AND=pd.DataFrame(AND)
df_AND
# +
X,y =prepare_data(df_AND)
ETA=0.1
EPOCHS=10
model_and = Perceptron(eta=ETA,epochs=EPOCHS)
model_and.fit(X,y)
# -
model_and.predict(X)
model_or.predict(X=[[1,0]])
model_and.predict(X=[[1,1]])
# +
XOR = {
"x1": [0,0,1,1],
"x2": [0,1,0,1],
"y" : [0,1,1,0]
}
df_XOR = pd.DataFrame(XOR)
df_XOR
X, y = prepare_data(df_XOR)
ETA = 0.1
EPOCHS = 10
model_xor = Perceptron(eta=ETA, epochs=EPOCHS)
model_xor.fit(X, y)
| Research_env/Perceptron_implimentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decorators
#
#
# A decorator involves concepts of functional programming. Although it is the same underlying concept, **we have two different types of decorators** in Python:
#
# * **functional** decorators.
#
# * **class** decorators
#
# A **decorator** in Python is **any** callable Python object that is used **to modify a function or a class**. A reference to a `func` function or a `C` class is passed to a decorator and the decorator returns a modified function or class. Modified functions or classes generally contain calls to the original `func` function or `C` class.
#
# First, **function names are references** to functions and **we can assign multiple names to the same function**. It is possible to **nest functions** within functions. It is also **possible** to make one **function** the **argument of another**.
#
# The **decoration occurs on the line before the function header**. The `@` is **followed** by the **decorator function name**.
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# show all ouputs of the same cell in the notebook, not only the last
# +
def our_decorator(func):
def function_wrapper(x):
print("Before calling {}".format(func.__name__))
func(x)
print("After calling {}".format(func.__name__))
return function_wrapper
@our_decorator
def foo(x):
print("foo has been called with: {}".format(x))
foo('he hey')
# -
# It can also be used with functions already created in any package.
# +
from math import sin, cos
def our_decorator(func):
def function_wrapper(x):
print("Before calling {}".format(func.__name__))
res = func(x)
print(res)
print("After calling {}".format(func.__name__))
return function_wrapper
# Alternative way to invoke the decorator
sin = our_decorator(sin)
cos = our_decorator(cos)
for f in [sin, cos]:
f(3.1415)
# -
# The `function_wrapper` above only works for **functions with exactly one parameter**. **We provide a generalized version** of `function_wrapper`, **which accepts functions with arbitrary parameters** in the following example.
# +
from random import randint, choice
def our_decorator(func):
def function_wrapper(*args, **kwargs):
print("Before calling {}".format(func.__name__))
res = func(*args, **kwargs)
print(res)
print("After calling {}".format(func.__name__))
return function_wrapper
# Decorated functions
randint = our_decorator(randint)
choice = our_decorator(choice)
randint(3, 8)
choice([4, 5, 6])
# -
# To summarize, we can say that a decorator in Python is a callable Python object that is used to modify a function, method or class definition. The original object, the one to be modified, is passed to a decorator as an argument. The decorator returns a modified object, e.g. a modified function, which is bound to the name used in the definition.
#
# Let's see some interesting applications. For argument parsing, check that the input number for calculating the factorial is natural.
# +
def argument_test_natural_number(f):
def helper(x):
if type(x) == int and x > 0:
return f(x)
else:
raise Exception("Argument is not an integer")
return helper
@argument_test_natural_number
def factorial(n):
if n == 1:
return 1
else:
return n * factorial(n-1)
# recurrence function
factorial(4)
factorial(-1)
# -
# To count the number of times a function has been invoked.
# +
def call_counter(func):
def helper(*args, **kwargs):
helper.calls += 1
# method to count the calls
return func(*args, **kwargs)
helper.calls = 0
return helper
@call_counter
def succ(x):
return x + 1
@call_counter
def mul1(x, y=1):
return x*y + 1
for i in range(3):
mul1(i,i+1)
print(f'Number of calls {mul1.calls}')
# -
# It is possible to make the **decorator depend on a parameter** that **particularizes the decoration according to the need**. We will achieve this by **wrapping the decorator in a function**, which could already depend on several parameters although in the example we make it depend on only one.
# +
def greeting(expr):
# function that wraps the decorator
def greeting_decorator(func):
# decorator function, note that it has func argument
def function_wrapper(x):
# real action performed
print(f'I say {expr}, a my function {func.__name__} returns: ')
func(x)
return function_wrapper
return greeting_decorator
@greeting("καλημερα")
def foo(x):
print('Another thing')
foo('Needs an useless argument')
# +
# Alternative way
def foo(x):
print('Another thing')
greeting2 = greeting("καλημερα")
foo = greeting2(foo)
foo('Needs an useless argument')
# -
# By using this previous dependency on a function, a decorator can show the results of a generator.
# +
def go_throughInf(n):
def go_through(gen):
# note that gen is the object to wrap
def gtAux(*args, **kwargs):
g = gen(*args, **kwargs)
# expect random number of inputs
sentinel = True
for _ in range(n):
try:
print(next(g))
except StopIteration:
print('empty generator')
return gtAux
return go_through
@go_throughInf(10)
# display first 10 number of fibonacci
def fib(x=0,y=1):
a, b = x, y
while True:
yield a
a, b = b, a + b
fib()
# -
1e6
# To measure the performance of a function in terms of time and resources.
# +
import psutil
import time
import os
def manytimeit(num):
def timeit(func):
def timed(*args, **kw):
accumulatortime = []
accumulatorcpu = []
for _ in range(num):
ts = time.time()
pid = os.getppid()
result = func(*args, **kw)
te = time.time()
p = psutil.Process(pid)
nowcpu = p.cpu_percent(interval=1) / psutil.cpu_count()
accumlatortime = accumulatortime.append(te-ts)
accumlatorcpu = accumulatorcpu.append(nowcpu)
meantime = sum(accumulatortime)/len(accumulatortime)
meancpu = sum(accumulatorcpu)/len(accumulatorcpu)
# maxmem = psutil.virtual_memory()[3]/(1e6)
# total memory in use for the system
# info = (pid, func.__name__, meantime, maxcpu)
print(f'PID:{pid}\nName:{func.__name__}\nTime:{meantime:.10f} sec over {num} times\nCPU:{meancpu:.3f} %')
return result
return timed
return timeit
@manytimeit(3)
# perform that number of times the timing
def fibIte(n):
def fibonacciIte(n):
a, b = 0, 1
for i in range(n):
a, b = b, a + b
return a
return fibonacciIte(n)
fibIte(10)
# -
# To study in detail the memory consumed by each line of your code you must run it as a script and use the `memory_profiler` package. Run `python example.py` when by using the below decorator, and when not explicitly adding the package just use `python -m memory_profiler example.py`.
# +
from memory_profiler import profile
@profile
def fibIte(n):
def fibonacciIte(n):
a, b = 0, 1
for i in range(n):
a, b = b, a + b
return a
return fibonacciIte(n)
# -
# When you want to access certain attributes of the function being wrapped the best solution is the following.
# +
from functools import wraps
# this function wraps the attributes of the main function to avoid presenting the attributes of the decorator
def greeting(func):
@wraps(func)
def function_wrapper(x):
""" function_wrapper of greeting """
print("Hi, " + func.__name__ + " returns:")
return func(x)
return function_wrapper
@greeting
def f(x):
""" just some silly function """
return x + 4
print(f(10))
print("function name: " + f.__name__)
print("docstring: " + f.__doc__)
print("module name: " + f.__module__)
# -
# So far we have used functions as decorators. Before we can define a **decorator as a class**, **we have to introduce the class method** `__call__`. We already mentioned that a decorator is simply an invocable object that takes a function as an input parameter. **A function is an invocable object, but many Python programmers do not know that there are other invocable objects**. An invocable object is an object that can be used and behaves like a function but might not be a function. It is possible to **define classes** so that **instances** are **invocable objects**. The `__call__` method is called, if the instance is called "like a function", i.e., using parentheses.
# +
class decorator2:
def __init__(self, f):
self.f = f
print('Initializing decorator class')
def __call__(self):
print("Decorating", self.f.__name__)
self.f()
@decorator2
def foo():
print("inside foo()")
foo()
# -
| A_decorators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: RPyc-Python
# language: Python
# name: rpyc
# ---
import os
print(os.uname())
# +
from maix import display, camera
while True:
tmp = camera.capture()
if tmp:
display.show(tmp)
# -
| tests/general/usage_display_hook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Data Science with Spark - Analyzing Stock Prices and Twitter Data
#
# * We need to import the relevant Quandl and Twitter Data
# * Then we choose a data storage method to persist the data
# * We use PySpark to generate descriptive stats
# * After which we use some modelling techniques
# * Optional (ML in R or Spark)
# * Port to Shiny app
#
# ## Ingest the Data Using API's
import quandl
import pandas as pd
quandl.ApiConfig.api_key = 'cxtmPDusyCfZtdF29pyA'
# +
STOCK_SYMBOLS = ["AAPL", "FB", "TSLA", "ANF", "UAL","RIGL", "AMZN", "MSFT", "GOOG", "XOM", "JNJ", "GE", "WFC", "WMT", "PFE", "KO", "ORCL", "BAC", "INTC", "V", "DIS", 'MRK']
for stock in STOCK_SYMBOLS:
str_code = 'WIKI/' + stock
print str_code
data = quandl.get(str_code.strip(), start_date = '2014-01-01')
pandas_df = pd.DataFrame(data)
pandas_df['TICKER'] = stock
all_df.append(pandas_df)
# -
augmented_df = pd.concat(all_df)
augmented_df.count()
| data/notebooks/nb_724669.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:RoboND]
# language: python
# name: conda-env-RoboND-py
# ---
# +
# Import some packages from matplotlib
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
# Uncomment the next line for use in a Jupyter notebook
# %matplotlib inline
# Define the filename, read and plot the image
filename = 'sample.jpg'
image = mpimg.imread(filename)
plt.imshow(image)
plt.show()
| code/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import geopandas
import numpy as np
municipalities = geopandas.read_file("/Users/jaydenfont/Desktop/Code/Spark/ACLU/CENSUS2010TOWNS_SHP/CENSUS2010TOWNS_POLY.shp", encoding="utf-8")
municipalities
municipalities[["TOWN2", "TYPE"]].head(20)
# +
def check_match(x, towns):
if x in list(towns.departments):
return float(towns[towns.departments == x].crime_discrepancy)
else:
return np.nan
def plot_map(municipalities, year):
midsize = pd.read_excel("/Users/jaydenfont/Desktop/Code/Spark/ACLU/midsize_areas_ranking.xlsx", f"midsize_areas_ranking_{year}")
urban = pd.read_excel("/Users/jaydenfont/Desktop/Code/Spark/ACLU/urban_areas_breakdown.xlsx", f"urban_areas_breakdown_{year}")
merged = pd.concat([midsize, urban]).reset_index(drop=True)
municipalities['discrepancy'] = municipalities.TOWN2.apply(check_match, args=(merged,))
municipalities[['TOWN2', "discrepancy"]]
map = municipalities.plot(column='discrepancy', figsize=(12, 9),
legend=True, legend_kwds={
"label": "Arrest Discrepancy",
"orientation": "horizontal"
}, edgecolor="black",
cmap="winter")
map.set_title(f"Map of Arrest Discrepancies in MA by Municipality {year}")
map.figure.savefig(f"Map_{year}")
# -
for i in range(2016, 2020):
plot_map(municipalities, i)
all_years = []
for year in range(2016, 2020):
midsize = pd.read_excel("/Users/jaydenfont/Desktop/Code/Spark/ACLU/midsize_areas_ranking.xlsx", f"midsize_areas_ranking_{year}")
urban = pd.read_excel("/Users/jaydenfont/Desktop/Code/Spark/ACLU/urban_areas_breakdown.xlsx", f"urban_areas_breakdown_{year}")
merged = pd.concat([midsize, urban]).reset_index(drop=True)
all_years.append(merged)
all_years_merged = pd.concat(all_years)
year_groups = all_years_merged.groupby(by="year")
# +
total_change = {"2016-2017": [], "2017-2018": [], "2018-2019": []}
percent_change = {"2016-2017": [], "2017-2018": [], "2018-2019": []}
for i in range(0, 3):
if i == 0:
y1 = year_groups.get_group(2016)
y2 = year_groups.get_group(2017)
for town in y2.departments:
try:
v1 = float(list(y1[y1.departments == town].crime_discrepancy)[0])
v2 = float(list(y2[y2.departments == town].crime_discrepancy)[0])
change = v2-v1
total_change["2016-2017"].append(change)
try:
percent_change["2016-2017"].append(100 * (change/v1))
except ZeroDivisionError:
percent_change["2016-2017"].append(np.nan)
except IndexError:
total_change["2016-2017"].append(np.nan)
percent_change["2016-2017"].append(np.nan)
elif i == 1:
y1 = year_groups.get_group(2017)
y2 = year_groups.get_group(2018)
for town in y2.departments:
try:
v1 = float(list(y1[y1.departments == town].crime_discrepancy)[0])
v2 = float(list(y2[y2.departments == town].crime_discrepancy)[0])
change = v2-v1
total_change["2017-2018"].append(change)
try:
percent_change["2017-2018"].append(100 * (change/v1))
except ZeroDivisionError:
percent_change["2017-2018"].append(np.nan)
except IndexError:
total_change["2017-2018"].append(np.nan)
percent_change["2017-2018"].append(np.nan)
else:
y1 = year_groups.get_group(2018)
y2 = year_groups.get_group(2019)
for town in y2.departments:
try:
v1 = float(list(y1[y1.departments == town].crime_discrepancy)[0])
v2 = float(list(y2[y2.departments == town].crime_discrepancy)[0])
change = v2-v1
total_change["2018-2019"].append(change)
try:
percent_change["2018-2019"].append(100 * (change/v1))
except ZeroDivisionError:
percent_change["2018-2019"].append(np.nan)
except IndexError:
total_change["2018-2019"].append(np.nan)
percent_change["2018-2019"].append(np.nan)
# -
for year in range(2017, 2020):
midsize = pd.read_excel("/Users/jaydenfont/Desktop/Code/Spark/ACLU/midsize_areas_ranking.xlsx", f"midsize_areas_ranking_{year}")
urban = pd.read_excel("/Users/jaydenfont/Desktop/Code/Spark/ACLU/urban_areas_breakdown.xlsx", f"urban_areas_breakdown_{year}")
merged = pd.concat([midsize, urban]).reset_index(drop=True)
if year == 2017:
merged["total_change_2016-2017"] = total_change["2016-2017"]
merged["pct_change_2016-2017"] = percent_change["2016-2017"]
elif year == 2018:
merged["total_change_2017-2018"] = total_change["2017-2018"]
merged["pct_change_2017-2018"] = percent_change["2017-2018"]
else:
merged["total_change_2018-2019"] = total_change["2018-2019"]
merged["pct_change_2018-2019"] = percent_change["2018-2019"]
merged.to_excel(f"data_with_pct_change_{year}.xlsx", index=False)
| Police Arrest Analysis/maps_and_pct_change.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 4344, "status": "ok", "timestamp": 1624910861530, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="pFhy95XbZqOS" outputId="accf5617-2ac2-4be7-d3b1-7d0ee4950615"
import torch
from torch.autograd import grad
import torch.nn as nn
from numpy import genfromtxt
import torch.optim as optim
import matplotlib.pyplot as plt
import torch.nn.functional as F
sidr_data = genfromtxt('sidr_50.csv', delimiter=',') #in the form of [t,S,I,D,R]
torch.manual_seed(1234)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 713, "status": "ok", "timestamp": 1624910862231, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="AD6iFgYfZqOa" outputId="1ef08aca-e086-4e9e-fd6c-b87c82c74649"
# %%time
PATH = 'covid_50_pts'
class DINN(nn.Module):
def __init__(self, t, S_data, I_data, D_data, R_data): #[t,S,I,D,R]
super(DINN, self).__init__()
self.N = 59e6 #population size
self.t = torch.tensor(t, requires_grad=True)
self.t_float = self.t.float()
self.t_batch = torch.reshape(self.t_float, (len(self.t),1)) #reshape for batch
self.S = torch.tensor(S_data)
self.I = torch.tensor(I_data)
self.D = torch.tensor(D_data)
self.R = torch.tensor(R_data)
self.losses = []
self.save = 3 #which file to save to
self.alpha_tilda = torch.nn.Parameter(torch.rand(1, requires_grad=True)) #0.191
self.beta_tilda = torch.nn.Parameter(torch.rand(1, requires_grad=True)) #0.05
self.gamma_tilda = torch.nn.Parameter(torch.rand(1, requires_grad=True)) #0.0294
#find values for normalization
self.S_max = max(self.S)
self.I_max = max(self.I)
self.D_max = max(self.D)
self.R_max = max(self.R)
self.S_min = min(self.S)
self.I_min = min(self.I)
self.D_min = min(self.D)
self.R_min = min(self.R)
#unnormalize
self.S_hat = (self.S - self.S_min) / (self.S_max - self.S_min)
self.I_hat = (self.I - self.I_min) / (self.I_max - self.I_min)
self.D_hat = (self.D - self.D_min) / (self.D_max - self.D_min)
self.R_hat = (self.R - self.R_min) / (self.R_max - self.R_min)
#matrices (x4 for S,I,D,R) for the gradients
self.m1 = torch.zeros((len(self.t), 4)); self.m1[:, 0] = 1
self.m2 = torch.zeros((len(self.t), 4)); self.m2[:, 1] = 1
self.m3 = torch.zeros((len(self.t), 4)); self.m3[:, 2] = 1
self.m4 = torch.zeros((len(self.t), 4)); self.m4[:, 3] = 1
#NN
self.net_sidr = self.Net_sidr()
self.params = list(self.net_sidr.parameters())
self.params.extend(list([self.alpha_tilda, self.beta_tilda, self.gamma_tilda]))
#force parameters to be in a range
@property
def alpha(self):
return torch.tanh(self.alpha_tilda)*0.191*20
@property
def beta(self):
return torch.tanh(self.beta_tilda)*0.05*20
@property
def gamma(self):
return torch.tanh(self.gamma_tilda)*0.0294*20
#nets
class Net_sidr(nn.Module): # input = [t]
def __init__(self):
super(DINN.Net_sidr, self).__init__()
self.fc1=nn.Linear(1, 20) #takes 100 t's
self.fc2=nn.Linear(20, 20)
self.fc3=nn.Linear(20, 20)
self.fc4=nn.Linear(20, 20)
self.out=nn.Linear(20, 4) #outputs S, I, D, R
def forward(self, t_batch):
sidr=F.relu(self.fc1(t_batch))
sidr=F.relu(self.fc2(sidr))
sidr=F.relu(self.fc3(sidr))
sidr=F.relu(self.fc4(sidr))
sidr=self.out(sidr)
return sidr
def net_f(self, t_batch):
sidr_hat = self.net_sidr(t_batch)
S_hat, I_hat, D_hat, R_hat = sidr_hat[:,0], sidr_hat[:,1], sidr_hat[:,2], sidr_hat[:,3]
#S_t
sidr_hat.backward(self.m1, retain_graph=True)
S_hat_t = self.t.grad.clone()
self.t.grad.zero_()
#I_t
sidr_hat.backward(self.m2, retain_graph=True)
I_hat_t = self.t.grad.clone()
self.t.grad.zero_()
#D_t
sidr_hat.backward(self.m3, retain_graph=True)
D_hat_t = self.t.grad.clone()
self.t.grad.zero_()
#R_t
sidr_hat.backward(self.m4, retain_graph=True)
R_hat_t = self.t.grad.clone()
self.t.grad.zero_()
#unnormalize
S = self.S_min + (self.S_max - self.S_min) * S_hat
I = self.I_min + (self.I_max - self.I_min) * I_hat
D = self.D_min + (self.D_max - self.D_min) * D_hat
R = self.R_min + (self.R_max - self.R_min) * R_hat
f1_hat = S_hat_t - (-(self.alpha / self.N) * S * I) / (self.S_max - self.S_min)
f2_hat = I_hat_t - ((self.alpha / self.N) * S * I - self.beta * I - self.gamma * I ) / (self.I_max - self.I_min)
f3_hat = D_hat_t - (self.gamma * I) / (self.D_max - self.D_min)
f4_hat = R_hat_t - (self.beta * I ) / (self.R_max - self.R_min)
return f1_hat, f2_hat, f3_hat, f4_hat, S_hat, I_hat, D_hat, R_hat
def load(self):
# Load checkpoint
try:
checkpoint = torch.load(PATH + str(self.save)+'.pt')
print('\nloading pre-trained model...')
self.load_state_dict(checkpoint['model'])
self.optimizer.load_state_dict(checkpoint['optimizer_state_dict'])
self.scheduler.load_state_dict(checkpoint['scheduler'])
epoch = checkpoint['epoch']
self.losses = checkpoint['losses']
except RuntimeError :
print('changed the architecture, ignore')
pass
except FileNotFoundError:
pass
def train(self, n_epochs):
#try loading
self.load()
#train
print('\nstarting training...\n')
for epoch in range(n_epochs):
#lists to hold the output (maintain only the final epoch)
S_pred_list = []
I_pred_list = []
D_pred_list = []
R_pred_list = []
f1, f2, f3, f4, S_pred, I_pred, D_pred, R_pred = self.net_f(self.t_batch)
self.optimizer.zero_grad()
S_pred_list.append(self.S_min + (self.S_max - self.S_min) * S_pred)
I_pred_list.append(self.I_min + (self.I_max - self.I_min) * I_pred)
D_pred_list.append(self.D_min + (self.D_max - self.D_min) * D_pred)
R_pred_list.append(self.R_min + (self.R_max - self.R_min) * R_pred)
loss = (torch.mean(torch.square(self.S_hat - S_pred))+
torch.mean(torch.square(self.I_hat - I_pred))+
torch.mean(torch.square(self.D_hat - D_pred))+
torch.mean(torch.square(self.R_hat - R_pred))+
torch.mean(torch.square(f1))+
torch.mean(torch.square(f2))+
torch.mean(torch.square(f3))+
torch.mean(torch.square(f4))
)
loss.backward()
self.optimizer.step()
self.scheduler.step()
self.losses.append(loss.item())
if epoch % 1000 == 0:
print('\nEpoch ', epoch)
#loss + model parameters update
if epoch % 4000 == 9999:
#checkpoint save
print('\nSaving model... Loss is: ', loss)
torch.save({
'epoch': epoch,
'model': self.state_dict(),
'optimizer_state_dict': self.optimizer.state_dict(),
'scheduler': self.scheduler.state_dict(),
#'loss': loss,
'losses': self.losses,
}, PATH + str(self.save)+'.pt')
if self.save % 2 > 0: #its on 3
self.save = 2 #change to 2
else: #its on 2
self.save = 3 #change to 3
print('epoch: ', epoch)
print('alpha: (goal 0.191 ', self.alpha)
print('beta: (goal 0.05 ', self.beta)
print('gamma: (goal 0.0294 ', self.gamma)
return S_pred_list, I_pred_list, D_pred_list, R_pred_list
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1595, "status": "ok", "timestamp": 1624910863824, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="_P1obOwWZqOc" outputId="2d2bbe1b-bcb6-45f2-f659-96d18ba62341"
# %%time
dinn = DINN(sidr_data[0], sidr_data[1], sidr_data[2], sidr_data[3],
sidr_data[4]) #in the form of [t,S,I,D,R]
learning_rate = 1e-5
optimizer = optim.Adam(dinn.params, lr = learning_rate)
dinn.optimizer = optimizer
scheduler = torch.optim.lr_scheduler.CyclicLR(dinn.optimizer, base_lr=1e-6, max_lr=1e-3, step_size_up=1000, mode="exp_range", gamma=0.85, cycle_momentum=False)
dinn.scheduler = scheduler
try:
S_pred_list, I_pred_list, D_pred_list, R_pred_list = dinn.train(1) #train
except EOFError:
if dinn.save == 2:
dinn.save = 3
S_pred_list, I_pred_list, D_pred_list, R_pred_list = dinn.train(1) #train
elif dinn.save == 3:
dinn.save = 2
S_pred_list, I_pred_list, D_pred_list, R_pred_list = dinn.train(1) #train
# + colab={"base_uri": "https://localhost:8080/", "height": 296} executionInfo={"elapsed": 608, "status": "ok", "timestamp": 1624910864429, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="WwqBVtEM9FYG" outputId="fd9639ad-3782-4e46-e1a2-f5d39e7dac58"
plt.plot(dinn.losses[300000:], color = 'teal')
plt.xlabel('Epochs')
plt.ylabel('Loss'),
# + colab={"base_uri": "https://localhost:8080/", "height": 722} executionInfo={"elapsed": 522, "status": "ok", "timestamp": 1624910864948, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="pJrvoRWQZqOd" outputId="7aa1d9b2-d3c9-4c97-9583-bec2b3504db2"
fig = plt.figure(figsize=(12,12))
ax = fig.add_subplot(111, facecolor='#dddddd', axisbelow=True)
ax.set_facecolor('xkcd:white')
ax.scatter(sidr_data[0], sidr_data[1], c = 'black', alpha=0.5, lw=2, label='Susceptible Data')
ax.plot(sidr_data[0], S_pred_list[0].detach().numpy(), 'red', alpha=0.9, lw=2, label='Susceptible Prediction', linestyle='dashed')
ax.scatter(sidr_data[0], sidr_data[2], c = 'violet', alpha=0.5, lw=2, label='Infected Data')
ax.plot(sidr_data[0], I_pred_list[0].detach().numpy(), 'dodgerblue', alpha=0.9, lw=2, label='Infected Prediction', linestyle='dashed')
ax.scatter(sidr_data[0], sidr_data[3], c = 'darkgreen', alpha=0.5, lw=2, label='Dead Data')
ax.plot(sidr_data[0], D_pred_list[0].detach().numpy(), 'green', alpha=0.9, lw=2, label='Dead Prediction', linestyle='dashed')
ax.scatter(sidr_data[0], sidr_data[4], c = 'blue', alpha=0.5, lw=2, label='Recovered Data')
ax.plot(sidr_data[0], R_pred_list[0].detach().numpy(), 'teal', alpha=0.9, lw=2, label='Recovered Prediction', linestyle='dashed')
ax.set_xlabel('Time /days',size = 20)
ax.set_ylabel('Number',size = 20)
#ax.set_ylim([-1,50])
ax.yaxis.set_tick_params(length=0)
ax.xaxis.set_tick_params(length=0)
plt.xticks(size = 20)
plt.yticks(size = 20)
# ax.grid(b=True, which='major', c='black', lw=0.2, ls='-')
legend = ax.legend(prop={'size':20})
legend.get_frame().set_alpha(0.5)
for spine in ('top', 'right', 'bottom', 'left'):
ax.spines[spine].set_visible(False)
plt.savefig('covid_50_pts.pdf')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 8, "status": "ok", "timestamp": 1624910864948, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="MBbyd2AgPrwe" outputId="0dad25e2-90ca-485d-b9e9-77ce1f9fb86e"
#calculate relative MSE loss
import math
import numpy as np
S_total_loss = 0
S_den = 0
I_total_loss = 0
I_den = 0
D_total_loss = 0
D_den = 0
R_total_loss = 0
R_den = 0
t = np.linspace(0, 500, 50)
for timestep in range(len(t)):
S_value = sidr_data[1][timestep] - S_pred_list[0].detach().numpy()[timestep]
S_total_loss += S_value**2
S_den += (sidr_data[1][timestep])**2
I_value = sidr_data[2][timestep] - I_pred_list[0].detach().numpy()[timestep]
I_total_loss += I_value**2
I_den += (sidr_data[2][timestep])**2
D_value = sidr_data[3][timestep] - D_pred_list[0].detach().numpy()[timestep]
D_total_loss += D_value**2
D_den += (sidr_data[3][timestep])**2
R_value = sidr_data[4][timestep] - R_pred_list[0].detach().numpy()[timestep]
R_total_loss += R_value**2
R_den += (sidr_data[4][timestep])**2
S_total_loss = math.sqrt(S_total_loss/S_den)
I_total_loss = math.sqrt(I_total_loss/I_den)
D_total_loss = math.sqrt(D_total_loss/D_den)
R_total_loss = math.sqrt(R_total_loss/R_den)
print('S_total_loss: ', S_total_loss)
print('I_total_loss: ', I_total_loss)
print('D_total_loss: ', D_total_loss)
print('R_total_loss: ', R_total_loss)
# + colab={"base_uri": "https://localhost:8080/", "height": 774} executionInfo={"elapsed": 1022, "status": "ok", "timestamp": 1624910866084, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="iUzZI6VMZqOe" outputId="309d00a2-f6e8-41bb-d36f-01a408a8774d"
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
# Initial conditions
N = 59e6
S0 = N - 1
I0 = 1
D0 = 0
R0 = 0
# A grid of time points (in days)
t = np.linspace(0, 500, 50)
#parameters
alpha = dinn.alpha
beta = dinn.beta
gamma = dinn.gamma
print(alpha)
print(beta)
print(gamma)
# The SIR model differential equations.
def deriv(y, t, alpha, betta, gamma):
S, I, D, R = y
dSdt = - (alpha / N) * S * I
dIdt = (alpha / N) * S * I - beta * I - gamma * I
dDdt = gamma * I
dRdt = beta * I
return dSdt, dIdt, dDdt, dRdt
# Initial conditions vector
y0 = S0, I0, D0, R0
# Integrate the SIR equations over the time grid, t.
ret = odeint(deriv, y0, t, args=(alpha, beta, gamma))
S, I, D, R = ret.T
# Plot the data on two separate curves for S(t), I(t)
fig = plt.figure(facecolor='w', figsize=(12,12))
ax = fig.add_subplot(111, facecolor='#dddddd', axisbelow=True)
ax.plot(t, S, 'violet', alpha=0.5, lw=2, label='Learnable Param Susceptible', linestyle='dashed')
ax.plot(t, sidr_data[1], 'dodgerblue', alpha=0.5, lw=2, label='Susceptible')
ax.plot(t, I, 'darkgreen', alpha=0.5, lw=2, label='Learnable Param Infected', linestyle='dashed')
ax.plot(t, sidr_data[2], 'gold', alpha=0.5, lw=2, label='Susceptible')
ax.plot(t, D, 'red', alpha=0.5, lw=2, label='Learnable Param Dead', linestyle='dashed')
ax.plot(t, sidr_data[3], 'salmon', alpha=0.5, lw=2, label='Dead')
ax.plot(t, R, 'blue', alpha=0.5, lw=2, label='Learnable Param Recovered', linestyle='dashed')
ax.plot(t, sidr_data[4], 'wheat', alpha=0.5, lw=2, label='Recovered')
ax.set_xlabel('Time /days')
ax.yaxis.set_tick_params(length=0)
ax.xaxis.set_tick_params(length=0)
ax.grid(b=True, which='major', c='w', lw=2, ls='-')
legend = ax.legend()
legend.get_frame().set_alpha(0.5)
for spine in ('top', 'right', 'bottom', 'left'):
ax.spines[spine].set_visible(False)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 8, "status": "ok", "timestamp": 1624910866084, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="R-zofRIm2RNz" outputId="0dbda0e4-6064-4e6f-928d-ec60233b13bc"
#calculate relative MSE loss
import math
S_total_loss = 0
S_den = 0
I_total_loss = 0
I_den = 0
D_total_loss = 0
D_den = 0
R_total_loss = 0
R_den = 0
for timestep in range(len(t)):
S_value = sidr_data[1][timestep] - S[timestep]
S_total_loss += S_value**2
S_den += (sidr_data[1][timestep])**2
I_value = sidr_data[2][timestep] - I[timestep]
I_total_loss += I_value**2
I_den += (sidr_data[2][timestep])**2
D_value = sidr_data[3][timestep] - D[timestep]
D_total_loss += D_value**2
D_den += (sidr_data[3][timestep])**2
R_value = sidr_data[4][timestep] - R[timestep]
R_total_loss += R_value**2
R_den += (sidr_data[4][timestep])**2
S_total_loss = math.sqrt(S_total_loss/S_den)
I_total_loss = math.sqrt(I_total_loss/I_den)
D_total_loss = math.sqrt(D_total_loss/D_den)
R_total_loss = math.sqrt(R_total_loss/R_den)
print('S_total_loss: ', S_total_loss)
print('I_total_loss: ', I_total_loss)
print('D_total_loss: ', D_total_loss)
print('R_total_loss: ', R_total_loss)
# + executionInfo={"elapsed": 7, "status": "ok", "timestamp": 1624910866085, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiXQKL7UiRoL28-GMShElFe0PuFh4NWnMP9hbDD=s64", "userId": "12455150063240177220"}, "user_tz": 360} id="6lFJLEj4LFVw"
| Experiments/more_or_less_data/covid_50_pts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import IPython.display as ipd
print("original(оригинал)")
ipd.display(ipd.Audio('/content/Multi-Tacotron-Voice-Cloning/book/Alexandr_Slobodskoy/Alexandr_Slobodskoy_1.wav')) #filepath to original voices (Путь до оригинального файла)
print("cloned(клонированный)")
ipd.display(ipd.Audio('/home/vogorjachko/logs-my_run/wavs'))#filepath to synthesized voices (Путь до синтезированного файла)
# +
import IPython.display as ipd
ipd.display(ipd.Audio('/home/vogorjachko/logs-my_run/wavs/step-300-wave-from-mel.wav'))#filepath to synthesized voices (Путь до синтезированного файла)
# +
ipd.display(ipd.Audio('/home/vogorjachko/logs-gst_run/wavs/step-300-wave-from-mel.wav'))#filepath to synthesized voices (Путь до синтезированного файла)
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Graph
# ### <img src="https://raw.githubusercontent.com/aadishgoel2013/Must-Do-Coding-Questions/master/graph_diagram.png">
# +
n = 6
graph = {
1:[(7,2), (9,3), (14,6)],
2:[(7,1), (10,3), (15,4)],
3:[(9,1), (10,2), (11,4), (2,6)],
4:[(15,2), (11,3), (6,5)],
5:[(6,4),(9,6)],
6:[(14,1),(2,3), (9,5)],
}
graph2 = [
(1,2,7),
(1,3,9),
(1,6,14),
(2,3,10),
(2,4,15),
(3,4,11),
(3,6,2),
(4,5,6),
(5,6,9),
]
graph3 = [
(1,2),
(1,3),
(1,6),
(2,3),
(3,6),
(4,5),
]
# -
# ## Dijkstra Algo
# +
from heapq import heappush, heappop
def dijkstra(graph, start):
weights = [None]* (len(graph)+1)
heap = [(0, start)]
while heap:
path_len, min_node = heappop(heap)
if weights[min_node] is None:
weights[min_node] = path_len
for edge_len, node in graph[min_node]:
if weights[node] is None:
heappush(heap, (edge_len+path_len, node))
return weights
dijkstra(graph1, 1)
# -
# ## Kruskal
# +
def kruskal(graph, n):
gp = {i:i for i in range(1,n+1)}
sets = { i:[i] for i in range(1,n+1)}
graph.sort(key=lambda x:x[2])
edges_used = []
for u,v,w in graph:
if gp[u]!=gp[v]:
temp = gp[v]
for node in sets[gp[v]]:
gp[node] = gp[u]
sets[gp[u]].append(node)
del sets[temp]
edges_used.append((u,v,w))
return edges_used
kruskal(graph2, n)
# -
# ### Connected Components (using kruskal)
# +
def connected_components(graph, n):
gp = {i:i for i in range(1,n+1)}
sets = { i:[i] for i in range(1,n+1)}
for u,v in graph:
if gp[u]!=gp[v]:
temp = gp[v]
for node in sets[gp[v]]:
gp[node] = gp[u]
sets[gp[u]].append(node)
del sets[temp]
return len(sets)
connected_components(graph3, n)
# -
| Graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# 
import numpy as np
import pandas as pd
from IPython.display import Image
# ## Predictive Modeling
# What we saw above is a common setup. We have $\mathbf{X}$ and $\mathbf{y}$ data from the past and $\mathbf{X}$ data for the present for which we want to **predict** the future $\mathbf{y}$ values.
#
# We can generalize this notion of past / present data into what's generally called **train** and **test** data.
#
# * **Training Data** -- A dataset that we use to train our model. We have both $\mathbf{X}$ and $\mathbf{y}$
# * **Testing Data** -- A dataset which only has $\mathbf{X}$ values and for which we need to predict $\mathbf{y}$ values. We might also have access to the real $\mathbf{y}$ values so that we can test how well our model will perform on data it hasn't seen before.
#
# ### <span style="color:red">Model Fitting Exercise</span>
# 1. Partner up. On one computer:
# 1. Write a function with the call signature `predict_test_values(model, x_train, y_train, x_test)` where `model` is a scikit learn model
# 1. Fit the model on `x_train` and `y_train`
# 1. Predict the y values for `X_test`
# 1. Return a vector of predicted y values
# 1. Write a second function with the call signature `calc_train_and_test_error(model, x_train, y_train, x_test, y_test)`
# 1. Fit the model on `x_train` and `y_train`
# 1. Predict the y values for `x_test`
# 1. Predict the y values for `x_train`
# 1. Calculate the `mean_squared_error` on both the train and test data.
# 1. Return the train error and test error
# 1. Describe to your partner the situations in which you might use each function
# +
def mean_squared_error(y_true, y_pred):
"""
calculate the mean_squared_error given a vector of true ys and a vector of predicted ys
"""
diff = y_true - y_pred
return np.dot(diff, diff) / len(diff)
def predict_test_values(model, X_train, y_train, X_test):
model.fit(X_train, y_train)
return model.predict(X_test)
def calc_train_and_test_error(model, X_train, y_train, X_test, y_test):
model.fit(X_train, y_train)
y_pred_train = model.predict(X_train)
y_pred_test = model.predict(X_test)
return mean_squared_error(y_train, y_pred_train), mean_squared_error(y_test, y_pred_test)
# -
# ## The Central Theses of Machine Learning
# ### <span style="color:green">**1) A predictive model is only as good as its predictions on unseen data **</span>
#
# ### <span style="color:green">**2) Error on the dataset we trained on is not a good predictor of error on future data**</span>
#
# Why isn't error on the training data a good indicator of future performance? Overfitting.
# ### Overfitting in One Picture
Image(url='http://radimrehurek.com/data_science_python/plot_bias_variance_examples_2.png')
# ## How to Fight Overfitting?
# Ultimately we don't want to build a model which performs well on data we've already seen, we want to build a model which will perform well on data we haven't seen.
#
# There are two linked strategies for to accomplish this: **regularization** and **model selection**.
# ## Regularization
# The idea in regularization is that we're going to modify our loss function to penalize it for being too complex. Simple models are better.
#
# One way to do this is to try to keep our regression coefficients small. Why would we want to do this? One intuitive explanation is that if we have big regression coefficients we'll get large changes in the predicted values from small changes in input value. That's bad. Intuitively, our predictions should vary smoothly with the data.
#
# So a model with smaller coefficients makes smoother predictions. It is simpler, which means it will have a harder time overfitting.
#
# We can change our linear regression loss function to help us reduce overfitting:
#
# ### Linear Regression Loss Function
# \begin{eqnarray*}
# Loss(\beta) = MSE &=& \frac{1}{N} \sum_{i=1}^{N} (y_i - \hat y_i)^2 \\
# &=& \frac{1}{N} \sum_{i=1}^{N} (y_i - x_i^T\beta)^2 \\
# \end{eqnarray*}
#
# ### L2 Regularized Linear Regression Loss Function -- "Ridge"
# \begin{eqnarray*}
# Loss(\beta) &=& \frac{1}{N} \sum_{i=1}^{N} (y_i - x_i^T\beta)^2 + \alpha ||\beta||_2^2\\
# &=& \frac{1}{N} \sum_{i=1}^{N} (y_i - x_i^T\beta)^2 + \alpha \beta^T \beta\\
# &=& \frac{1}{N} \sum_{i=1}^{N} (y_i - x_i^T\beta)^2 + \alpha \sum_{d=1}^D \beta_d^2\\
# \end{eqnarray*}
# We won't get into details, but a ridge regression model can be optimized in much the same way as an unregularized linear regression: either with using some form of gradient descent or matrix-based solutions.
# +
# Ridge Regression in scikit-learn
from sklearn import linear_model
model_ridge = linear_model.Ridge(alpha = .5)
# once it's been fit, you can look at the learned beta values of the model with: model_ridge.coef_
# -
# ### <span style="color:red">Ridge Regression Errors</span>
# 1. Partner up. On one computer:
# 1. Using your `calc_train_and_test_error` function from the previous exercise:
# 1. Calculate the training and testing error for a LinearRegression model on the dataset below
# 1. Calculate the training and testing error for a Ridge regression model with `alpha=1` on the dataset below
# 1. Add up the absolute values of the coefficients of each model. Which is bigger?
#
# **Note:** If you have a fit model called `m`, then you can access a vector holding its learned coefficients with `m.coef_`.
#
# **Note:** Check out the functions `np.sum()` and `np.abs()`
#
# 1. Discuss with your partner what's happening here
# +
# load overfitting data
with np.load('data/overfitting_data.npz') as data:
x_train = data['x_train']
y_train = data['y_train']
x_test = data['x_test']
y_test = data['y_test']
model_lr = linear_model.LinearRegression()
model_ridge = linear_model.Ridge(alpha=1)
print "Linear Regression Training and Test Errors:"
print calc_train_and_test_error(model_lr, x_train, y_train, x_test, y_test)
print
print "Ridge Regression Training and Test Errors:"
print calc_train_and_test_error(model_ridge, x_train, y_train, x_test, y_test)
print
print "Sum of Linear Regression Coefficients:"
print np.sum(np.abs(model_lr.coef_))
print
print "Sum of Ridge Regression Coefficients:"
print np.sum(np.abs(model_ridge.coef_))
print
# -
# ?linear_model.Ridge
# ### L1 Regularized Linear Regression Loss Function -- "LASSO"
# LASSO is another regularization method. It penalizes not with the square of the regression coefficients (the $\beta$s) but with their absolute values.
#
# LASSO has the additional property that it tends to push beta values of unimportant dimensions all the way to exactly 0. This has the beneficial property of enforcing sparsity in our model. If having lots of small coefficients leads to a simpler model, having lots of 0-valued coefficients lead to even simpler models.
#
# \begin{eqnarray*}
# Loss(\beta) &=& \frac{1}{N} \sum_{i=1}^{N} (y_i - x_i^T\beta)^2 + \alpha ||\beta||_1\\
# &=& \frac{1}{N} \sum_{i=1}^{N} (y_i - x_i^T\beta)^2 + \alpha \sum_{d=1}^D |\beta_d|\\
# \end{eqnarray*}
# LASSO in scikit-learn
from sklearn import linear_model
model_lasso = linear_model.Lasso(alpha = 0.5)
# ### <span style="color:red">LASSO Coefficients and Errors</span>
# 1. Partner up. On one computer:
# 1. Using your `calc_train_and_test_error` again, calculate the training and testing error for a LASSO model with `alpha=1` on the dataset from the previous exercise
# 1. Add up the absolute values of the coefficients of the LASSO model. Compare it to the coefficient sums from the `LinearRegression` and `Ridge` models.
# 1. Look at the first 10 coefficients of the `LinearRegression`, `Ridge`, and `LASSO` models.
# 1. Discuss with your partner what's happening here
# +
# Write your code here
model_lasso = linear_model.Lasso(alpha=1)
print "Ridge Regression Training and Test Errors:"
print calc_train_and_test_error(model_lasso, x_train, y_train, x_test, y_test)
print
print "Sum of Ridge Regression Coefficients:"
print np.sum(np.abs(model_lasso.coef_))
print
# +
n_disp_coefs = 10
print 'Linear Regression Coefficients:'
print model_lr.coef_[:n_disp_coefs]
print
print 'Ridge Regression Coefficients:'
print model_ridge.coef_[:n_disp_coefs]
print
print 'LASSO Coefficients:'
print model_lasso.coef_[:n_disp_coefs]
print
# -
# ### L1 + L2 Regularized Linear Regression Loss Function -- "ElasticNet"
# \begin{eqnarray*}
# Loss(\beta) &=& \frac{1}{2N} \sum_{i=1}^{N} (y_i - x_i^T\beta)^2 + \alpha \rho ||\beta||_1 + \frac{\alpha (1 - \rho)}{2} ||\beta||_2^2\\\\
# &=& \frac{1}{2N} \sum_{i=1}^{N} (y_i - x_i^T\beta)^2 + \alpha \rho \sum_{d=1}^D |\beta_d| + \frac{\alpha (1 - \rho)}{2} \sum_{d=1}^D \beta_d^2\\
# \end{eqnarray*}
# +
from sklearn import linear_model
model_en = linear_model.ElasticNet(alpha=0.5, l1_ratio=0.1)
# note: scikit learn's current implementation of ElasticNet isn't stable with l1_ratio <= 0.01
# -
# ### <span style="color:red">ElasticNet Coefficients and Errors</span>
# 1. Partner up. On one computer:
# 1. Using your `calc_train_and_test_error` again, calculate the training and testing error for an `ElasticNet` model with `alpha=1` and `l1_ratio=0.5` on the dataset from the previous exercises
# 1. Add up the absolute values of the first 10 coefficients of the `ElasticNet` model. Compare it to the sums from the `LinearRegression`, `Ridge`, and `LASSO` models.
# 1. Look at the first 10 coefficients of the `ElasticNet` model.
# 1. Discuss with your partner what's happening here
# +
# Write your code here
model_en = linear_model.ElasticNet(alpha=1, l1_ratio=0.5)
print 'ElasticNet Errors:'
print calc_train_and_test_error(model_en, x_train, y_train, x_test, y_test)
print
print 'Sum of ElasticNet Coefficients'
print np.sum(np.abs(model_en.coef_))
print
n_disp_coefs = 10
print 'ElasticNet Coefficients:'
print model_en.coef_[:n_disp_coefs]
print
# -
# ## Cross Validation
# Now we know three types of regularization for linear regression: ridge regression, LASSO, and elastic net.
#
# All of our regularized models had better test error that simple linear regression. But how should we choose which model to ultimatley use or which parameters to use? The answer is through careful use of **cross validation**.
#
# There are many forms of cross validation, but the basic idea of each is to _train_ your model on some data and _estimate it's future performance_ on other data.
# ## Types of Cross Validation
# ### Validation Set Cross Validation
# 1. Pick an amount of training data to be in your validation data set (e.g. 10%)
# 2. Randomly split datapoints into training points (90%) and validation points (10%)
# 3. Train your model on the training data
# 4. Test your model on the validation data, record the validation error
# 5. Estimated future errors is the validation error
#
#
# * **Good:** Easy and computationally cheap
# * **Bad:** Statistically noisy and wastes data
#
#
# **Aside:** So far we've been calculating error on out test dataset. This is conceptually almost identical to using a validation set, but with two significant differences:
# 1. We don't have to peek at our test data set. This is good because if we do that too much, we can actually still overfit to our test data and still perform poorly on future unseen data.
# 1. We don't have to be given the $\mathbf{y}$ vector for our test dataset. Validation set cross validation only requires a training dataset.
# +
# a helper function for performing validation set cross validation
from sklearn.cross_validation import train_test_split
validation_portion = 0.1
seed = 1234
x_train_small, x_valid, y_train_small, y_valid = \
train_test_split(x_train, y_train, test_size=validation_portion, random_state=seed)
print 'Original Training Set Size:'
print x_train.shape, y_train.shape
print
print 'Reducted Training Set Size:'
print x_train_small.shape, y_train_small.shape
print
print 'Validation Set Size:'
print x_valid.shape, y_valid.shape
print
# -
# #### <span style="color:red">Validation Set Cross Validation Exercise</span>
# 1. Partner up. On one computer:
# 1. Write a function with the call signature `validation_set_error(model, x_train, y_train, validation_portion=0.1, seed=1234)` which returns the validation set estimate of the future error for the given `model`. This function should:
# 1. Split the data into a reduced training set and a validation set
# 1. Train on the reduced training set
# 1. Estimate the mean squared error on the validation set
# 1. Return that estimate
# 1. Use your `calc_train_and_test_error(model, x_train, y_train, x_test, y_test)` function to calculate training and test set errors for these
# 1. Use this your `validation_set_error` function to estimate the future error on the overfitting data for:
# 1. A linear regression model
# 1. A ridge regression models with `alpha` = 10
# 1. Do this for multiple random seeds
# 1. Does validation error do a good job of predicting test error?
# 1. If you have time: How does changing the validation_portion affect the similarity between the validation and test error?
#
# +
def validation_set_error(model, x_train, y_train, validation_portion=0.1, seed=1234):
# FILL IN YOUR CODE HERE
x_train_small, x_valid, y_train_small, y_valid = \
train_test_split(x_train, y_train, test_size=validation_portion, random_state=seed)
model.fit(x_train_small, y_train_small)
y_pred_valid = model.predict(x_valid)
return mean_squared_error(y_valid, y_pred_valid)
# set up models
model_lr_valid = linear_model.LinearRegression()
model_ridge_valid = linear_model.Ridge(alpha=10)
# calculate errors
valid_portion = .1
n_seeds = 5
print "Linear Regression Training and Test Errors:"
# FILL IN YOUR CODE HERE
print calc_train_and_test_error(model_lr_valid, x_train_small, y_train_small, x_test, y_test)
print
print "Linear Regression Validation Errors:"
# FILL IN YOUR CODE HERE
print validation_set_error(model_lr_valid, x_train, y_train, validation_portion=0.1, seed=1234)
print
for seed in range(n_seeds):
print validation_set_error(model_lr_valid, x_train, y_train, validation_portion=valid_portion, seed=seed)
print
print "Ridge Regression Training and Test Errors:"
# FILL IN YOUR CODE HERE
print calc_train_and_test_error(model_ridge_valid, x_train_small, y_train_small, x_test, y_test)
print
print "Ridge Regression Validation Errors:"
# FILL IN YOUR CODE HERE
print validation_set_error(model_ridge_valid, x_train, y_train, validation_portion=0.1, seed=1234)
print
for seed in range(n_seeds):
print validation_set_error(model_ridge_valid, x_train, y_train, validation_portion=valid_portion, seed=seed)
print
# -
# ### K-Fold Cross Validation
# K-Fold cross validation is another cross validation method for estimating the out-of-sample error of a model. It works like this:
#
# 1. Partition the training data into K folds
# 2. For each fold k in 1 to K:
# 1. Train the model on all your data except the data in fold k
# 2. Record the error on the the data in fold k
# 3. Estimate future error as average error across all folds
Image(url='https://chrisjmccormick.files.wordpress.com/2013/07/10_fold_cv.png')
# * **Good:** Only wastes 100/k% of the data at a time
# * **Bad:** Takes k times long as just training one model, still wastes 100/k% of the data
# +
# scikit learn provides a useful object to help you perform kfold cross validation
from sklearn.cross_validation import KFold
n_data = len(y_train)
fold_count = 0
for train_reduced_row_ids, valid_row_ids in KFold(n_data, n_folds=4):
print
print
print "FOLD %d:" % fold_count
print "-------"
print("train_ids:\n%s\n\nvalid_ids\n%s" % (train_reduced_row_ids, valid_row_ids))
x_train_reduced = x_train[train_reduced_row_ids]
y_train_reduced = y_train[train_reduced_row_ids]
x_valid = x_train[valid_row_ids]
y_valid = y_train[valid_row_ids]
fold_count += 1
# -
# NOTE: KFolds isn't random at all. It's important to shuffle your data first before using it.
from sklearn.utils import shuffle
x_train_shuffled, y_train_shuffled = shuffle(x_train, y_train)
# #### <span style="color:red">K-Fold Cross Validation Exercise</span>
# 1. Partner up. On one computer:
# 1. Write a function with the call signature `kfold_error(model, x_train, y_train, k=4, seed=1234)` which returns the k-fold cross validation estimate of the future error for the given `model`. This function should:
# 1. Shuffle the training data set (both $\mathbf{x}$ and $\mathbf{y}$ in unison)
# 1. For each fold:
# 1. Split the data into a reduced training set and a validation set
# 1. Train on the reduced training set
# 1. Estimate the mean squared error on the validation set
# 1. Add the estimated error to a running sum of the estimated total error
# 1. Return the average error across folds: i.e.: the estimated total error divided by the number of folds
# 1. Use your `calc_train_and_test_error(model, x_train, y_train, x_test, y_test)` function to calculate training and test set errors for these
# 1. Use your `kfold_error` function with k=5 to estimate the future error on the overfitting data for:
# 1. A linear regression model
# 1. A ridge regression models with `alpha` = 10
# 1. Do this for multiple random seeds
# 1. Does k-fold error do a good job of predicting test error?
# +
def kfold_error(model, x_train, y_train, k=4, seed=1234):
# FILL IN YOUR CODE HERE
# shuffle training data
x_train_shuffled, y_train_shuffled = shuffle(x_train, y_train, random_state=seed)
n_data = len(y_train)
error_sum = 0
for train_reduced_row_ids, valid_row_ids in KFold(n_data, n_folds=k):
x_train_reduced = x_train_shuffled[train_reduced_row_ids]
y_train_reduced = y_train_shuffled[train_reduced_row_ids]
x_valid = x_train_shuffled[valid_row_ids]
y_valid = y_train_shuffled[valid_row_ids]
model.fit(x_train_reduced, y_train_reduced)
y_valid_pred = model.predict(x_valid)
error_sum += mean_squared_error(y_valid, y_valid_pred)
return error_sum*1.0 / k
# set up models
model_lr_valid = linear_model.LinearRegression()
model_ridge_valid = linear_model.Ridge(alpha=10)
# calculate errors
n_seeds = 3
k = 5
print "Linear Regression Training and Test Errors:"
# FILL IN YOUR CODE HERE
print calc_train_and_test_error(model_lr_valid, x_train, y_train, x_test, y_test)
print
print "Linear Regression K-Fold Errors:"
# FILL IN YOUR CODE HERE
print
for seed in range(n_seeds):
print kfold_error(model_lr_valid, x_train, y_train, k=k, seed=seed)
print
print
print "Ridge Regression Training and Test Errors:"
# FILL IN YOUR CODE HERE
print calc_train_and_test_error(model_ridge_valid, x_train, y_train, x_test, y_test)
print
print "Ridge Regression K-Fold Errors:"
# FILL IN YOUR CODE HERE
print
for seed in range(n_seeds):
print kfold_error(model_ridge_valid, x_train, y_train, k=k, seed=seed)
print
# -
# ## Putting It All Together: Model and Hyperparameter Selection with Cross Validation
# 1. For each model and hyperparameter combo you're willing to consider:
# 1. Estimate the model's performance on future data using cross validation
# 2. Pick the model with the best estimated future performance
# 3. Train the best model from scratch on the full dataset. This is your final model
[np.nan] + [1,2]
# +
def model_name(model):
s = model.__str__().lower()
if "linearregression" in s:
return 'LinearRegression'
elif "lasso" in s:
return 'Lasso(a=%g)' % model.alpha
elif "ridge" in s:
return 'Ridge(a=%g)' % model.alpha
elif "elastic" in s:
return 'ElasticNet(a=%g, r=%g)' % (model.alpha, model.l1_ratio)
else:
raise ValueError("Unknown Model Type")
def create_models(alphas=(.01, .03, .1, .3, 1, 3), l1_ratios=(.7, .5, .3)):
models = [linear_model.LinearRegression()]
models.extend([linear_model.Ridge(a) for a in alphas])
models.extend([linear_model.Lasso(a) for a in alphas])
models.extend([linear_model.ElasticNet(a, l1_ratio=l) for a in alphas for l in l1_ratios])
return models
def results_df(models, betas_true, x_train, y_train, x_test, y_test, k=4):
n_data, n_dim = x_train.shape
n_zeros = n_dim - len(betas_true)
betas_true = np.concatenate([betas_true, np.zeros(n_zeros)])
# fit models to training data
[m.fit(x_train, y_train) for m in models]
betas = np.vstack([betas_true] + [m.coef_ for m in models])
beta_names = ['Beta ' + str(i) for i in range(n_dim)]
# set up model names
model_names = ["True Coefs"] + [model_name(m) for m in models]
df = pd.DataFrame(data=betas, columns=beta_names, index=model_names)
# calculate training errors
y_preds = [m.predict(x_train) for m in models]
errors = [np.nan] + [mean_squared_error(y_train, y_pred) for y_pred in y_preds]
df['Train Error'] = errors
# calculate validation errors
errors = [np.nan] + [kfold_error(m, x_train, y_train, k=k) for m in models]
df['Cross Validation Error'] = errors
# calculate test errors
y_preds = [m.predict(x_test) for m in models]
errors = [np.nan] + [mean_squared_error(y_test, y_pred) for y_pred in y_preds]
df['Test Error'] = errors
return df
# these are some of the magic parameters that I used to actually
# generate the overfitting dataset
n_dim = 598
n_dim_meaningful = 3
n_dim_disp_extra = 2
# the actual betas used to generate the y values. the rest were 0.
betas_true = np.arange(n_dim_meaningful) + 1
# create a whole bunch of untrained models
models = create_models(alphas=(.01, .03, .1, .3, 1), l1_ratios=(.9, .7, .5))
#
all_results = results_df(models, betas_true, x_train, y_train, x_test, y_test, k=4)
# decide which columns we want to display
disp_cols = ["Beta " + str(i) for i in range(n_dim_meaningful + n_dim_disp_extra)]
disp_cols += ['Train Error', 'Cross Validation Error', 'Test Error']
# display the results
all_results[disp_cols]
# -
# %matplotlib inline
import matplotlib.pyplot as plt
f = plt.figure()
plt.scatter(all_results['Cross Validation Error'], all_results['Test Error'])
plt.xlabel('Cross Validation Error')
plt.ylabel('Test Error')
f.set_size_inches(8, 8)
plt.show()
# scikit learn includes some functions for making cross validation easier
# and computationally faster for a some models
from sklearn import linear_model
model_ridge_cv = linear_model.RidgeCV(alphas=[0.1, 1.0, 10.0])
model_lasso_cv = linear_model.LassoCV(alphas=[0.1, 1.0, 10.0])
model_en_cv = linear_model.ElasticNetCV(l1_ratio=[.9], n_alphas=100)
# ## Caveats:
# * You can still overfit with intensive cross validation based model selection!
# * But it's much better than without
# ## Summary:
# * **The Central Theses of Machine Learning:**
# * We're only interested in predictive performance on unseen data, not on seen data.
# * **Training error** estimates error on **seen** data
# * **Cross validation error** estimates error on **unseen** data
# * **Regularization** strategies change how to train a model so that it will perform better on unseen data
# * We talked about three forms of regularization for linear regression:
# * **Ridge Regression** (L2 Penalty)
# * **LASSO** (L1 Penalty)
# * **ElasticNet** (L1 + L2 Penalties)
# * We talked about two kinds of cross validation error:
# * **Validation Error** -- split your training set into a reduced training set and a validation set
# * **K-Fold Error** -- Split your training data into k reduced training sets and a validation sets
# * Regularization introduces new hyperparameters
# * Use a cross validated estimate of future performance to choose your model and hyperparameter settings
| Session 2 - Overfitting_Regularization_ModelSelection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
from IPython.display import display, HTML
from IPython.core.interactiveshell import InteractiveShell
#InteractiveShell.ast_node_interactivity = "all"
InteractiveShell.ast_node_interactivity = "last_expr"
# %autoreload 2
import pandas as pd
pd.set_option("display.precision",2)
# # Heating Model
# This notebook uses the weather and consumption data we have from 2015-2018 to build a simple linear heating model for the house.
#
# The model is, simply put, too simple. Here are the major failings:
# - DHW load is probably a huge fraction of the base load. But many of these measure could reduce it.
# - Passive solar gain is neglected.
# - Simply subtracting average base load is a gross oversimplification.
#
# Both of these are amenable to refinement, given sufficient time.
# Load the data we have
import pandas as pd
data = pd.read_pickle('wxhydro2018.pickle')
# ## Determine baseline electrical consumption
# The house's electrical consumption is shown in the center histogram below. It's a very obvious bimodal distribution, that looks a lot like a mixture of gaussians, and I intuitively know what it means: there is a significant base load (centered on the left peak), and a very separate heating load (centered on the right peak).
#
# #### Known base load
# Fortunately, we have an easy way to learn a lot about the base-load -- I happen to know that in this house, essentially no mechanical heating or cooling is ever required when the OAT is between 16C and 20C (and actually a fair bit outside of that range), so we will use that to discover our baseline. Most of it is domestic hot water, which is quite random. But there are also some lights, computers, and other appliances. To make things easier to handle, we'll plot the non-heating consumption on the basis of a 24-hr sliding-window average. The left plot shows our averaged no-heat power data. Sure enough, it looks like a slightly skewed gaussian, with a nice little low-load peak on the left that probably indicates the load during unoccupied periods longer than a day.
#
# #### First guess at heating load
# These two distributions are so distinct that it's pretty easy to imagine that they're gaussians and visually disentangle them. By inspection, we have a pretty good idea that the mean base load is 1.1 kW, and the mean total load *when there is heating active* is 2.4 kW. By looking at the relative areas of the two peaks, it's pretty clear that there are a lot more data points at base load than with heating active.
#
# A good first guess would be to subtract the mean of our *known* base load. This is quite obviously not a great fit -- it commingles a lot of base load points with the heating load, and our resulting heating model will be quite a bit on the low side.
#
# A good second estimate is to *filter* that plot to remove points that we suspect are probably base load. This has the opposite problem -- it will remove a lot of low-power heating points, biasing our model on the high side.
#
# #### Heating load
#
# The plot on the right shows the distribution of (power minus baseload), or as we're now calling it, heating load.
# + hideCode=true
# Compute load series
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# Set up the data we'll be using
# Rolling averages
data['OATAvg'] = data['OAT'].rolling(24).mean()
data['consAvg'] = data['cons'].rolling(24).mean()
#baseline=data[(data.OATAvg>=16)&(data.OATAvg<=22)] # Using calendar instead
baseline = data[(data.index.dayofyear>=167)&(data.index.dayofyear<259)] # Summer: June 15 - Sept 14
winter = data[(data.index.dayofyear<167)|(data.index.dayofyear>=259)] # Non-summer
#Compute
meanbaseload = np.mean(baseline.consAvg)
std = np.sqrt(np.var(baseline.consAvg))
baseload = (meanbaseload + 2.5 * std)
#baseload = meanbaseload
# Print mean, stddev, and baseload
print("Mean: %.2f kW StdDev: %.2f"%(meanbaseload,std) )
print("Offset: %.2f kW Baseload: %.2f kW" % (2.7*std,baseload))
# Calculated series
data['hmloadAvg']=data.eval('(consAvg - @meanbaseload)*(consAvg>=@meanbaseload)')
data['hloadAvg']=data.eval('(consAvg - @baseload)*(consAvg>=@baseload)')
data['hfloadAvg']=data.eval('(consAvg - @meanbaseload)*(consAvg>=@baseload)')
data['hload']=data.eval('(cons - @baseload)*(cons>@baseload)')
# -
# +
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(12,4),sharey=True)
ha = data.cons['2017-1-01'].plot(ax=axes[0])
ha.set_xlabel("Time")
ha.title.set_text('January 1 - Hourly Power')
ha = data.cons['2016-06-01'].plot(ax=axes[1])
ha.set_xlabel("Time")
ha.set_ylabel("Average power(kW)")
ha.title.set_text('June 1 - Hourly Power')
# +
# Plot consumption and load histograms
def allpower(a):
ha = data.consAvg.hist(bins=60,ax=a)
ha.set_xlabel("Averaged power(kW)")
ha.set_ylabel("hours")
ha.title.set_text('All electrical consumption')
for devs in range(-2,5):
if devs == 0: col = 'g'
else: col='r'
vl=ha.axvline(x=baseline.consAvg.mean()+devs*std, color=col, alpha=0.5)
def sumpow(a):
baseline=data[(data.index.month==7)|(data.index.month==8)]
ha = baseline.consAvg.hist(bins=60,ax=a)
ha.set_xlabel("Averaged power(kW)")
ha.set_ylabel("hours")
ha.title.set_text('Non-Heating (Where 16-20C)')
for devs in range(-2,3):
if devs == 0: col = 'g'
else: col='r'
vl=ha.axvline(x=baseline.consAvg.mean()+devs*std, color=col, alpha=0.5)
def nonheat(a):
ha = baseline.consAvg.hist(bins=60,ax=a)
ha.set_xlabel("Averaged power(kW)")
ha.set_ylabel("hours")
ha.title.set_text('Non-Heating (Jun 15-Sep 14)')
for devs in range(-2,3):
if devs == 0: col = 'g'
else: col='r'
vl=ha.axvline(x=baseline.consAvg.mean()+devs*std, color=col, alpha=0.5)
def meanheat(a):
mdshl= data['hmloadAvg'][data['hmloadAvg'].gt(0)]
hlp = mdshl.hist(bins=60,ax=a)
hlp.title.set_text('Heating (mean baseload of %.2f kW)'%meanbaseload)
hlp.set_xlabel("Nonzero heating power")
hlp.set_ylabel("hours at power")
def baseheat(a):
# Plot agg cons
dshl= data['hfloadAvg'][data['hfloadAvg'].gt(0)]
binrange=np.arange(0.0,4.0,5/60)
hlp = dshl.hist(bins=binrange,ax=a)
hlp.title.set_text('Heating ( baseload of %.2f kW)'%baseload)
hlp.set_xlabel("Nonzero heating power (filtered)")
hlp.set_ylabel("hours at power")
def plot4():
# Set up the 4-plot figure
fig, axes = plt.subplots(nrows=2, ncols=2,figsize=(15,10))
#fig.suptitle("Heating vs Non-Heating Hourly consumption",size=14)
allpower(axes[0,0])
nonheat(axes[0,1])
#sumpow(axes[0,1])
meanheat(axes[1,0])
baseheat(axes[1,1])
def plot3():
# Set up the 4-plot figure
fig, axes = plt.subplots(nrows=1, ncols=3,figsize=(15,5))
fig.suptitle("Heating vs Non-Heating Hourly consumption",size=14)
allpower(axes[0])
nonheat(axes[1])
meanheat(axes[2])
def plot2():
# Set up the figure
fig, axes = plt.subplots(nrows=2, ncols=1,figsize=(7,10))
#fig.suptitle("Heating vs Non-Heating Hourly consumption",size=14)
allpower(axes[0])
nonheat(axes[1])
#meanheat(axes[2])
def plotRes2():
# Set up the figure
fig, axes = plt.subplots(nrows=2, ncols=1,figsize=(7,10))
#fig.suptitle("Heating vs Non-Heating Hourly consumption",size=14)
meanheat(axes[0])
#baseheat(axes[1])
#meanheat(axes[2])
def plotRes():
# Set up the figure
#fig, axes = plt.subplots(nrows=2, ncols=1,figsize=(7,10))
#fig.suptitle("Heating vs Non-Heating Hourly consumption",size=14)
meanheat(axes[0])
#baseheat(axes[1])
#meanheat(axes[2])
plot2()
plt.show()
# +
def plotRes2():
# Set up the figure
fig, axes = plt.subplots(nrows=2, ncols=1,figsize=(7,10))
#fig.suptitle("Heating vs Non-Heating Hourly consumption",size=14)
meanheat(axes[0])
#baseheat(axes[1])
#meanheat(axes[2])
def plotRes():
# Set up the figure
fig, axes = plt.subplots(nrows=1, ncols=1,figsize=(7,7))
#fig.suptitle("Heating vs Non-Heating Hourly consumption",size=14)
meanheat(axes)
#baseheat(axes[1])
#meanheat(axes[2])
plotRes()
plt.show()
# -
# ## GMM visualization
#
# Sometimes it's useful to see if there's obvious clustering. Applying the sklearn Gaussian Mixture model to this data does indeed separate it roughly where we're suggesting -- somewhere along the 2 kW line.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
import numpy as np
xt = data[['cons','OAT']]['2015'].fillna(0)
from sklearn.mixture import GaussianMixture
gmm = GaussianMixture(n_components=2,covariance_type="full").fit(xt)
labels = gmm.predict(xt)
plt.scatter(xt.OAT,xt.cons, c=labels, s=4,cmap='RdBu'); # , cmap='viridis'
# -
# ## Scatter Plots
# Now that we have a reasonable value for *heating load* for every hour over our study period, our next job is to treat that as a dependent variable, and figure out which independent variables influence it.
#
# Typical environmental factors influencing heating load are things like OAT, wind speed, and solar irradiance (which we unfortunately don't have for this study period). There are human factors too, such as temperature setpoint and occupancy, but my house has a constant setpoint of 22C, and I have no occupancy data, so we'll have to neglect that.
#
# A good way to discover pairwise relationships is to look at scatter plots. So here are scatter plots of heating load (x-axis) against various weather data, for all points where heating load is non-zero.
# + hideCode=true
fig, axes = plt.subplots(nrows=3, ncols=3,figsize=(14,14))
COLS = ['OAT', 'OATAvg', 'Dewpoint','Relhum', 'Windir', 'Winspd', 'Press', 'Hmdx', 'Winchill']
r=0
s=1
for i in range (0,9):
c=i%3
colname=COLS[i]
if r>0:
s=4
data.plot.scatter(x='hloadAvg',y=colname,ax=axes[r,c],s=s)
if(c==2):
r=r+1
# -
# ## Visualize heating power versus Heating-degree-hours
# It's clear in the scatter matrix that an inverse correlation with OAT is really the only significant weather relationship we have -- dewpoint is really too strongly linked to temperature to be useful, and we can see that there isn't much of a correlation with relative humidity.
#
# It's common in the industry to straighten out this inverse correlation by using heating degree-days or degree-hours instead of temperature. That's just the sum-product of hours below 18C.
#
# The plot below shows a sliding window average of heating load versus HDHs. It looks like a pretty good fit.
# + hideCode=true
# Some weather stuff that we don't need
data['hdh']=data.eval('(18-OAT)*(18>OAT)')
data['hdhAvg']= data['hdh'].rolling(24).mean()
rs=[['20151001','20160401']
,['20161001','20170401']
,['20171001','20180301']]
pdata=data[data.hloadAvg>0][['hdhAvg','hloadAvg']]
fig,axes=plt.subplots(3,1,figsize=(16,12),sharex=False)
for i in range (0,3):
yr=2015+i
yd = pdata[rs[i][0]:rs[i][1]].sort_index()
yd.plot(yd.index,secondary_y='hloadAvg',ax=axes[i])
label = str(yr)+"-"+str(yr+1)
axes[i].set_ylabel(label)
#axes[i].plot(yd.hdh)
#ax2=axes[i].twinx()
#ax2.plot(yd.hloadAvg,'g-')
# -
# ## Linear Regression
# We are now reasonably confident that we now have a pretty good relationship between a trailing-window HDH and heating load. The data is very noisy, so it's not practical to try to model the behaviour precisely. A simple linear regression will probably be as good as anything else.
#
# Here, we've run linear regressions against both the *filtered* and *unfiltered* estimates of heating load. We know that one is biased high, and the other low, so for lack of enough time to do better, we'll just average them.
# +
import scipy.stats as stats
def plot1(a):
# Plot 1
h2data=data[(data.hdh>0)][['OATAvg','hmloadAvg']].replace([np.inf,-np.inf],np.nan).dropna()
lrr = stats.linregress(h2data.OATAvg,h2data.hmloadAvg)
slope, intercept, r_value, p_value, std_err = lrr
h2p = h2data.plot.scatter('OATAvg','hmloadAvg',s=3,ax=a)
h2p.title.set_text('Heating vs OAT')
f=lambda x: intercept + slope*x
h2p.plot(h2data.OATAvg,f(h2data.OATAvg),c='r')
x0 = -intercept/slope
display(lrr)
#print("X-intercept 1 is at y=%f kW"%f(0))
#print("Y-intercept 1 is at x=%f °C OAT"%x0)
return (intercept, slope)
def plot2(a):
# Plot 2
h3data=data[(data.hdh>0)&(data.hfloadAvg>0)][['OATAvg','hfloadAvg']].replace([np.inf,-np.inf],np.nan).dropna()
lrr = stats.linregress(h3data.OATAvg,h3data.hfloadAvg)
slope, intercept, r_value, p_value, std_err = lrr
h3p = h3data.plot.scatter('OATAvg','hfloadAvg',s=3,ax=a)
h3p.title.set_text('Filtered heating: Linear regression against OAT')
h3p.get_figure().suptitle("Linear regression of non-zero heating load against OAT")
f=lambda x: intercept + slope*x
h3p.plot(h3data.OATAvg,f(h3data.OATAvg),c='r')
x0 = -intercept/slope
display(lrr)
#print("X-intercept 2 is at y=%f kW"%f(0))
#print("Y-intercept 2 is at x=%f °C OAT"%x0)
return (intercept, slope)
def plotw(a):
# Plot 3: Winter hloadAvg
h3data=winter[['OATAvg','hloadAvg']].replace([np.inf,-np.inf],np.nan).dropna()
lrr = stats.linregress(h3data.OATAvg,h3data.hloadAvg)
slope, intercept, r_value, p_value, std_err = lrr
h3p = h3data.plot.scatter('OATAvg','hloadAvg',s=3,ax=a)
h3p.title.set_text('Filtered heating: Linear regression against OAT')
h3p.get_figure().suptitle("Linear regression of non-zero heating load against OAT")
f=lambda x: intercept + slope*x
h3p.plot(h3data.OATAvg,f(h3data.OATAvg),c='r')
x0 = -intercept/slope
display(lrr)
#print("X-intercept 2 is at y=%f kW"%f(0))
#print("Y-intercept 2 is at x=%f °C OAT"%x0)
return (intercept, slope)
def plotCompare():
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(15,7), sharey=True)
fig.suptitle("Linear Regression",size=14)
Heat01, HeatSlope1 = plot1(axes[0])
Heat02, HeatSlope2 = plotw(axes[1])
fig, axes = plt.subplots(nrows=1, ncols=1,figsize=(7,7))
fig.suptitle("Linear Regression",size=14)
Heat01, HeatSlope1 = plot1(axes)
#Heat02, HeatSlope2 = plotw(axes[1])
BaseThreshold=baseload
MeanBaseload=meanbaseload
#Heat0=(Heat01+Heat02)/2
#HeatSlope=(HeatSlope1+HeatSlope2)/2
Heat0=Heat01
HeatSlope=HeatSlope1
#Heat0=Heat01
#HeatSlope=HeatSlope1
# -
# ### Heating model
# So, the linear heating model we're going to use as a function of OAT in degC:
#
# $HeatLoad = max(0, Heat0 + HeatSlope \times OAT)\:kW \\
# Cons_{BAU} = (HeatLoad+BaseThreshold)\:kW$
# +
#heat=lambda temp: Heat0 + HeatSlope*temp
print("Heat0=%.2f"% Heat0)
print("HeatSlope=%.3f"% HeatSlope)
print("BaseThreshold=%.2f"%BaseThreshold)
print("MeanBaseload=%.2f"%MeanBaseload)
tempAtHeat=lambda h: (h - Heat0)/HeatSlope
print("HeatCutoff=%.2f"%tempAtHeat(0.0))
# -
# ## Validation
#
# +
# Validate
def doBAU(dfIn):
#df = pd.DataFrame( dfIn.loc[dfIn.index.year == year] )
df = dfIn
df['hload'] = np.maximum(0, Heat0 + HeatSlope * dfIn.OAT )
#df['bcons'] = df.hload + BaseThreshold # MeanBaseload
df['bcons'] = dfIn.hload + MeanBaseload
df['err'] = (df.cons - df.bcons) / df.cons *100
return df
def valYears(df):
for i in range(2015,2019):
#df = doBAU(data,year=i)
real = df.loc[df.index.year==i].cons.sum()
model = df.loc[df.index.year==i].bcons.sum()
print("Year %d total consumption (kWh): Real %d Model %d Error %.1f%%"%(i,real,model,abs(100*(model-real)/real)))
df = doBAU(data)
valYears(df)
# Total error:
real = df.cons.sum()
model = df.bcons.sum()
print("All years total consumption (kWh): Real %d Model %d Error %.1f%%"%(real,model,abs(100*(model-real)/real)))
# -
#hy.cons.groupby((hy.index.year,hy.index.month)).sum().plot(x=hy.index,y=hy.cons,figsize=(18,6))
df.err.groupby((df.index.year,df.index.month)).mean().plot(x=df.index,y=df.err)
#df.groupby((df.index.year,df.index.month)).mean().plot(x=df.index,y=['bcons'])
# ## Conclusions
#
# We could clearly have a better heating model -- our error in 2017 is quite high. I see two primary counfounding factors:
#
# 1. I've used a naiive approach to defining "heating" data points. As the baseload has a wide probability distribution, simply filtering out a "typical" baseload still leaves all of the baseload variability in the so-called "heating data."
#
# 2. The domestic hot water load is entangled with the heating load, and I haven't done a good enough job of separating them.
#
# Having a trustworthy heating model is key to making our following decisions, but perfecting it isn't really the purpose of this project. For now, we'll proceed with what we've got.
| .ipynb_checkpoints/AHeatingModel-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image classifier using least squares method
import numpy as np
# Let's build a simple classifier based on least square method.
#
# The data which we are using is written into two files - mnist_train.csv and mnist_test.csv.
# You can download it here: https://pjreddie.com/projects/mnist-in-csv/
#
# Attention! Reading data from csv files requires a lot of memory (up to couple of gigabytes) and computation power, run the following block of code only once.
# +
def load_train_data():
"""
reads csv file with train dataset
csv file should be located in the same directory with notebook
"""
return np.genfromtxt("mnist_train.csv", delimiter=',')
def load_test_data():
"""
reads csv file with test dataset
csv file should be located in the same directory with notebook
"""
return np.genfromtxt("mnist_test.csv", delimiter=',')
# -
train_data = load_train_data()
test_data = load_test_data()
# +
def select_images_and_labels_from_dataset(data, shape):
"""selects rundom images from dataset"""
old_shape = data.shape[0]
indexces = np.random.choice(old_shape, shape, replace=False)
labels = data[indexces, 0].astype(int)
images = data[indexces, 1:]
return labels, images
def get_train_labels_and_images():
"""selects random images from train data"""
return select_images_and_labels_from_dataset(train_data, N_train)
def get_test_labels_and_images():
"""selects random images from test data"""
return select_images_and_labels_from_dataset(test_data, N_test)
# -
# Now, let's define the amount of images we will use to train the classifier and test it.
# After that we will load train and test data and select rundom images from those datasets.
#
# As you will see, there are 60k and 10k vectors in train and test data respectively. Each vector contains 785 values and represents an image 28X28 with a lable. First value in a vector is lables of an image - digit from 0 to 9, next 784 values are image data.
N_train = 10000
N_test = 1000
train_labels, train_images = get_train_labels_and_images()
test_labels, test_images = get_test_labels_and_images()
# The algorythm of classfifier is the following:
# 1. Select test image from test images
# 2. Find the distances from selected image to train images
# 3. Find minimum distance and return it's label from train labels
def classify_image(test_image, train_images, train_labels):
DM = np.square(test_image - train_images).sum(axis=1)
index = DM.argmin(axis=0)
return train_labels[index]
predicted_results = [classify_image(test_image, train_images, train_labels) for test_image in test_images]
# Let's calculate the accuracy of our algorythm:
accuracy = (predicted_results == test_labels).sum() / N_test
print("SAMPLES COUNT:", N_train)
print("TEST COUNT:", N_test)
print("ACCURACY:", np.round(accuracy * 100, 2), '%')
# However, how we can maximise the accuracy? We will see how the size of train data influences the accuracy of image prediction.
# +
def run_train_itteration(N_train, test_images, test_labels):
train_labels, train_images = select_images_and_labels_from_dataset(train_data, N_train)
predicted_results = [classify_image(test_image, train_images, train_labels) for test_image in test_images]
accuracy = (predicted_results == test_labels).sum() / test_images.shape[0]
return accuracy
def get_average_accuracy(N_train, amount_of_iterations):
accuracies = [run_train_itteration(N_train, test_images, test_labels) for _ in range(amount_of_iterations)]
return sum(accuracies) / amount_of_iterations
# +
size_of_subset_v = [50, 200, 500, 1000, 2000, 5000, 10000]
count_of_iters_v = [100, 40, 20, 10, 5, 2, 1]
test_labels, test_images = get_test_labels_and_images()
accuracies = [get_average_accuracy(size, iters)
for size, iters in zip(size_of_subset_v, count_of_iters_v)]
# -
# Now let's visualize results with mathplotlib. Note that x-axis is logarithmically scaled.
# +
import matplotlib.pyplot as plt
plt.semilogx(size_of_subset_v, accuracies, color = "g")
plt.scatter(size_of_subset_v, accuracies, color = "b")
plt.xlabel("Size of train subset")
plt.ylabel("Accuracy")
plt.grid(axis="y", which="both", linestyle='--')
plt.show()
# -
| practise1/Practice1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 1. Importing Libraries
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sn
# ### 2. Read in data
dataset = pd.read_csv('P39-Financial-Data.csv')
dataset.head()
dataset.describe()
# ### data quality check
dataset.isna().any()
# ### 3. Count distribution analysis
# +
dataset2 = dataset.drop(columns = ['entry_id', 'pay_schedule', 'e_signed'])
fig = plt.figure(figsize=(15, 12))
plt.suptitle('Histograms of Numerical Columns', fontsize=20)
for i in range(dataset2.shape[1]):
plt.subplot(6, 3, i + 1)
f = plt.gca()
f.set_title(dataset2.columns.values[i])
vals = np.size(dataset2.iloc[:, i].unique())
if vals >= 100:
vals = 100
plt.hist(dataset2.iloc[:, i], bins=vals, color='#3F5D7D')
plt.tight_layout(rect=[0, 0.03, 1, 0.95])
# -
# ### 4. Correlation with Response Variable
dataset2.corrwith(dataset.e_signed).plot.bar(
figsize = (20, 10), title = "Correlation with E Signed", fontsize = 15,
rot = 35, grid = True)
# ### 5. Correlation bwt predictors
# +
## Correlation Matrix
sn.set(style="white")
# Compute the correlation matrix
corr = dataset2.corr()
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(18, 15))
# Generate a custom diverging colormap
cmap = sn.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sn.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# -
| Loan-prediction/.ipynb_checkpoints/eloan-prediction_EDA-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (WalkForwardBacktrader)
# language: python
# name: pycharm-a45eb1e2
# ---
# + pycharm={"is_executing": true}
from __future__ import (absolute_import, division, print_function,
unicode_literals)
import pandas as pd
import numpy as np
import statsmodels.formula.api as sm
import backtrader as bt
import backtrader.talib as talib
import backtrader.feeds as btfeeds
import backtrader.feed
import backtrader.indicators as btind
import backtrader.analyzers as btanalyzers
import backtrader.feeds as btfeeds
import backtrader.strategies as btstrats
import itertools
from IPython.display import display, HTML
import matplotlib.pyplot as plt
import matplotlib.dates as mpdates
import time;
from datetime import timedelta
import datetime
from sklearn import linear_model
import blackbox as bb
import math
from pprint import pprint
from backtrader import Indicator
import os
candle_freq = '4Hour'
coin_name = 'ETH'
datapath = os.path.abspath('..') + '/Data/GDAX/'
# %tb
# + pycharm={"is_executing": true}
candle_freq_to_seconds_map = {}
candle_freq_to_seconds_map['1Min'] = 60
candle_freq_to_seconds_map['3Min'] = 180
candle_freq_to_seconds_map['5Min'] = 300
candle_freq_to_seconds_map['15Min'] = 900
candle_freq_to_seconds_map['1Hour'] = 3600
candle_freq_to_seconds_map['4Hour'] = 14400
candle_freq_to_seconds_map['6Hour'] = 21600
# -
#
#
# ### 1) First we do the preleminary in sample testing and make sure the results are same with the trading view results (without slippage and commissions)
# + pycharm={"is_executing": true}
class AcctStats(bt.Analyzer):
"""A simple analyzer that gets the gain in the value of the account; should be self-explanatory"""
def __init__(self):
self.start_val = self.strategy.broker.get_value()
self.end_val = None
def stop(self):
self.end_val = self.strategy.broker.get_value()
def get_analysis(self):
return {"start": self.start_val, "end": self.end_val}
# + pycharm={"is_executing": true}
class ValueStats(bt.Analyzer):
"""A simple analyzer that gets the gain in the value of the account; should be self-explanatory"""
val = []
def __init__(self):
self.val = []
def next(self):
self.val.append(self.strategy.broker.get_value())
def get_analysis(self):
return self.val
# + pycharm={"is_executing": true}
class LinearRegression(Indicator):
alias = ('LR',)
lines = ('linear_regression','ma')
params = (
('len', 300),
)
iter = 0
def changeLen(self,length):
self.params.len = length
def next(self):
if (self.iter > self.params.len):
raw_prices = self.data.get(size=self.params.len)
prices = np.array(raw_prices).reshape(-1, 1)
x_line = np.array([i for i in range(0, self.params.len)]).reshape(-1, 1)
# Create linear regression object
regr = linear_model.LinearRegression()
# Train the model using the training sets
regr.fit(x_line, prices)
prediction = regr.predict(np.array([self.params.len]).reshape(-1, 1))
self.lines.linear_regression[0] = prediction
self.iter += 1
# + pycharm={"is_executing": true}
class MyStrategy(bt.Strategy):
def_params = (
('linear_reg_length', 20),
)
params = (
('interval_params', [(999999999999999.0, def_params)]),
('printlog', False),
('usable_cash_ratio' , 0.5),
)
def log(self, txt, dt=None, tm=None, doprint=False):
''' Logging function fot this strategy'''
if self.params.printlog or doprint:
dt = dt or self.datas[0].datetime.date(0)
tm = tm or self.datas[0].datetime.time(0)
print('%s, %s, %s' % (dt, tm, txt))
def get_params_for_time(self):
time_now_string = str(self.datas[0].datetime.date(0)).strip()
time_now = time.mktime(time.strptime(time_now_string, "%Y-%m-%d"));
if(self.sorted_params[self.interval_index][0] < time_now):
self.interval_index += 1
self.log('Params changed to : ' + str(self.sorted_params[self.interval_index][1]))
return self.sorted_params[self.interval_index][1]
def __init__(self):
self.long = False
self.dataclose = self.datas[0].close
self.datalow = self.datas[0].low
self.sorted_params = sorted(self.params.interval_params)
self.cash_buffer = (self.broker.getvalue()*self.params.usable_cash_ratio)
self.interval_index = 0
self.params_to_use = self.sorted_params[self.interval_index][1]
self.LR_low_trend = LinearRegression(self.datalow, len=self.params_to_use['linear_reg_length'])
self.LR_low_trend_to_use = self.LR_low_trend
def next(self):
self.params_to_use = self.get_params_for_time()
self.LR_low_trend.changeLen(self.params_to_use['linear_reg_length'])
if self.LR_low_trend_to_use < self.data.close and (not self.long):
self.size_to_buy = int((self.broker.getvalue()-self.cash_buffer) / self.dataclose[0])
self.order = self.buy(exectype=bt.Order.Market, size=self.size_to_buy)
self.long = True
elif self.LR_low_trend_to_use > self.data.close and self.long:
self.order = self.sell(exectype=bt.Order.Market, size=self.size_to_buy)
self.long = False
# + pycharm={"is_executing": true}
def getData(start_timestamp, end_timestamp):
path = datapath + str(coin_name) + '/' + str(candle_freq) + '.csv'
frame = pd.read_csv(path)
frame['time'] = pd.to_datetime(frame['time'], unit='s')
frame = frame[(frame['time'] >= pd.to_datetime(start_timestamp, unit='s')) \
& (frame['time'] <= pd.to_datetime(end_timestamp, unit='s'))]
frame = frame.sort_values(by=['time'])
frame = frame.rename(columns={'time': 'datetime'})
frame.set_index('datetime', inplace=True)
return frame
# + pycharm={"is_executing": true}
def RunBackTest(coin_name,candle_freq,capital,start_timestamp,end_timestamp,params\
,shouldPlot=False, shouldPrint=False):
frame_to_add = getData(start_timestamp, end_timestamp)
pd_frame = bt.feeds.PandasData(dataname=frame_to_add)
# Create a cerebro entity
cerebro = bt.Cerebro()
# Add a strategy
cerebro.addstrategy(MyStrategy ,interval_params = params['interval_params']\
,printlog = shouldPrint)
# Set commision and slippage
if('slippage' in params):
cerebro.broker.set_slippage_perc(perc=params['slippage'])
if('commision' in params):
cerebro.broker.setcommission(commission=params['commision'])
# Analyzer
cerebro.addanalyzer(btanalyzers.TradeAnalyzer, _name='TradeAnalysis')
cerebro.addanalyzer(btanalyzers.SharpeRatio,timeframe=bt.TimeFrame.Days,riskfreerate=0.0,\
_name='SharpeAnalysis')
cerebro.addanalyzer(btanalyzers.DrawDown , _name='DrawDownAnalysis')
cerebro.addanalyzer(AcctStats, _name='ActualAnalysis')
cerebro.addanalyzer(ValueStats, _name='ValueAnalysis')
cerebro.adddata(pd_frame)
cerebro.broker.setcash(capital)
starting_portfolio_value = cerebro.broker.getvalue()
# Print out the starting conditions
if(shouldPrint): print('Starting Portfolio Value: %.2f' % starting_portfolio_value)
# Run over everything
results = cerebro.run(runonce=False)
ending_portfolio_value = cerebro.broker.getvalue()
# Print out the final result
if(shouldPrint): print('Final Portfolio Value: %.2f' % ending_portfolio_value)
result = results[0]
trade_anlaysis = result.analyzers.TradeAnalysis.get_analysis()
sharpe_anlaysis = result.analyzers.SharpeAnalysis.get_analysis()
#calmar_anlaysis = result.analyzers.CalmarAnalysis.get_analysis()
drawdown_analysis = result.analyzers.DrawDownAnalysis.get_analysis()
returns_anlaysis = result.analyzers.ActualAnalysis.get_analysis()
value_anlaysis = result.analyzers.ValueAnalysis.get_analysis()
try :
total_pnl = returns_anlaysis['end'] - returns_anlaysis['start'] # ending_portfolio_value - starting_portfolio_value # trade_anlaysis['pnl']['gross']['total']
num_won_trades = trade_anlaysis['won']['total']
num_lost_trades = trade_anlaysis['lost']['total']
sharperatio = sharpe_anlaysis['sharperatio']
win_ratio = num_won_trades/(num_won_trades+num_lost_trades)
used_capital = (capital*0.5)
percentage_pnl = (total_pnl/ used_capital)*100
max_drawdown = drawdown_analysis['max']['drawdown']
calmarratio = percentage_pnl/math.sqrt((1+max_drawdown))
if(shouldPlot): cerebro.plot(iplot=False,style='candle')#,style='line')
return {'win_ratio':win_ratio, 'percentage_pnl':percentage_pnl,\
'sharperatio': sharperatio, 'calmarratio' :calmarratio, 'trade_analysis':trade_anlaysis,\
'value_anlaysis' : value_anlaysis}
except:
if(shouldPrint): print("Probably no trades were made")
return {'win_ratio':1.0, 'percentage_pnl':0.0,'sharperatio' : 0.0, 'calmarratio' :-1.0\
, 'trade_analysis':{}, 'value_anlaysis' : []}
# + pycharm={"is_executing": true}
# %matplotlib notebook
tstart = time.mktime(time.strptime("20.01.2018 00:00:00", "%d.%m.%Y %H:%M:%S"));
tend = time.mktime(time.strptime("10.03.2019 11:05:02", "%d.%m.%Y %H:%M:%S"));
interval_params_one_time = time.mktime(time.strptime("01.02.2019 21:05:02", "%d.%m.%Y %H:%M:%S"));
interval_params_one = {}
interval_params_one['linear_reg_length'] = 300
interval_params_two_time = tend
interval_params_two = {}
interval_params_two['linear_reg_length'] = 300
params={}
params['interval_params'] = [(interval_params_one_time , interval_params_one),\
(interval_params_two_time , interval_params_two)]
params['commision'] = 0
params['slippage'] = 0
max_lookback_buffer_param = max(interval_params_one.values())
needed_lookback_buffer_in_seconds = max_lookback_buffer_param*candle_freq_to_seconds_map[candle_freq]
start_date_with_buffer = tstart - needed_lookback_buffer_in_seconds
limitedEntryTestResult = RunBackTest(coin_name,candle_freq,200000.0,start_date_with_buffer,tend,params,True,True)
print(limitedEntryTestResult['win_ratio'])
print('PnL: ' + str(limitedEntryTestResult['percentage_pnl']) + '%')
print(limitedEntryTestResult['sharperatio'])
# -
# # Part 3 - Walk Forward Analysis
# + pycharm={"is_executing": true}
def get_params_to_optimize_ranges(optimization_params, tstart, tend):
a = optimization_params['linear_reg_length']
b = [tstart,tstart]
c = [tend,tend]
list_values = [ a,b,c]
return list_values
# + pycharm={"is_executing": true}
def getOptimizedParamsBlackBox(optimized_params):
params={}
params['linear_reg_length'] = int(optimized_params[0])
return params
# + pycharm={"is_executing": true}
def BlackBoxParallelWalkForwardAnalysis(start_date, end_date, optimization_period, out_of_sample_period\
,optimization_params, param_n, param_m, param_batch, OptFun\
,unanchored = True ):
optimization_start_date = start_date - optimization_period
optimization_end_date = start_date
testing_start_date = start_date
testing_end_date = start_date + out_of_sample_period
optimized_parameters = {}
out_of_sample_result = {}
while optimization_end_date < end_date:
params_to_optimize_ranges = get_params_to_optimize_ranges(optimization_params\
,optimization_start_date\
,optimization_end_date)
# Get the optimized params
optimization_start_date_key = str(time.strftime("%Y-%m-%d %H:%M:%S",time.localtime(optimization_start_date)))
optimization_end_date_key = str(time.strftime("%Y-%m-%d %H:%M:%S",time.localtime(optimization_end_date)))
optimization_key = optimization_start_date_key + "_" + optimization_end_date_key
print()
print("*****************************************")
print()
print("Optimizing for : ", optimization_start_date_key.split(" ")[0], " - ", \
optimization_end_date_key.split(" ")[0], " ...." )
# Run optimization for in Sample Period.
optimized_params = bb.search(f=OptFun, # given function
box=params_to_optimize_ranges, # range of values for each parameter (2D case)
n=param_n, # number of function calls on initial stage (global search)
m=param_m, # number of function calls on subsequent stage (local search)
batch=param_batch, # number of calls that will be evaluated in parallel
resfile= os.getcwd() + '/output.csv')
# text file where results will be saved
# Get the top 10 performing params for later analysis.
optimized_parameters[optimization_key] = optimized_params[0:20]
print("Optimization for start date : "\
+ str(optimization_start_date_key)\
+ " to end date : " +\
str(optimization_end_date_key)\
+ " is completed.")
print("testing dates for these optimization params are : ")
print("testing start date: " + str(time.strftime("%Y-%m-%d %H:%M:%S",time.localtime(testing_start_date))))
print("testing end date: " + str(time.strftime("%Y-%m-%d %H:%M:%S",time.localtime(testing_end_date))))
if(unanchored):
optimization_start_date += out_of_sample_period
optimization_end_date += out_of_sample_period
testing_start_date += out_of_sample_period
testing_end_date += out_of_sample_period
return {'optimized_parameters' : optimized_parameters\
,'out_of_sample_result' : out_of_sample_result}
# -
# ### BlackBox WalkForward
# #### optimization_period = timedelta(days=30) out_of_sample_period = timedelta(days=6)
# + pycharm={"is_executing": true}
def convertOptParamsToInputParams(paramsToOptimize):
params={}
params['linear_reg_length'] = int(paramsToOptimize[0])
return params
# + pycharm={"is_executing": true}
def opt_fun(paramsToOptimize):
tstart = paramsToOptimize[1]
tend = paramsToOptimize[2]
params=convertOptParamsToInputParams(paramsToOptimize)
interval_params=convertOptParamsToInputParams(paramsToOptimize)
params['interval_params'] = [(tend, interval_params)]
params['commision'] = 0.0005
params['slippage'] = 0.001
testResult = RunBackTest(coin_name,candle_freq,200000.0,tstart,tend\
,params,False,False)
return -testResult['percentage_pnl']
# + pycharm={"is_executing": true}
def getEquityCurve(WalkForwardAnalysisResults, start_ind):
equity_curve = []
total_pnl = 100
trade_date_keys = sorted(WalkForwardAnalysisResults.iterkeys())
for ind , key in enumerate(trade_date_keys):
if(ind < start_ind) : continue
this_result = WalkForwardAnalysisResults[key]
this_pnl = (this_result['percentage_pnl'] +100)/100
total_pnl = total_pnl*this_pnl
equity_curve.append(total_pnl-100)
return equity_curve
# + pycharm={"is_executing": true}
def getIntervalParamsFromOptimizedParams(optimized_params_list, out_of_sample_period, which_best):
sorted_opt_params = sorted(optimized_params_list)
interval_params = []
out_of_sample_start_date_with_buffer = 0
out_of_sample_start_date_to_log = 0
end_date = 0
for optimization_date_key in sorted_opt_params:
optimized_params = optimized_params_list[optimization_date_key][which_best]
optimization_end_date_key = optimization_date_key.split('_')[1]
optimization_end_date = time.mktime(time.strptime(optimization_end_date_key, "%Y-%m-%d %H:%M:%S"));
out_of_sample_start_date = optimization_end_date
out_of_sample_end_date = out_of_sample_start_date + out_of_sample_period
optimized_params_for_this_run = getOptimizedParamsBlackBox(optimized_params)
# This takes the first one and then never updates.
if(out_of_sample_start_date_with_buffer == 0):
#We need a warm start for the indicators to get ready.
max_lookback_buffer_param = max(optimized_params_for_this_run.values())
needed_lookback_buffer_in_seconds = max_lookback_buffer_param*candle_freq_to_seconds_map[candle_freq]
out_of_sample_start_date_with_buffer = out_of_sample_start_date - needed_lookback_buffer_in_seconds
out_of_sample_start_date_to_log = out_of_sample_start_date
# This updates everytime and ends up with the last iteration.
end_date = out_of_sample_end_date
interval_params.append((out_of_sample_end_date, optimized_params_for_this_run ))
return [interval_params, out_of_sample_start_date_with_buffer, out_of_sample_start_date_to_log, end_date]
# + pycharm={"is_executing": true}
def generateAlternateOutOfSampleResultsWithInterval(optimized_params_list, out_of_sample_period, top_best, shouldPlot):
out_of_sample_results = []
for i in range(top_best):
out_of_sample_result = {}
print("Doing " + str(i) + " th best params")
[out_of_sample_interval_params_for_this_run,\
out_of_sample_start_date_with_buffer,\
out_of_sample_start_date,\
out_of_sample_end_date] = getIntervalParamsFromOptimizedParams(optimized_params_list, out_of_sample_period,i)
optimized_params_for_this_run = {}
optimized_params_for_this_run['interval_params'] = out_of_sample_interval_params_for_this_run
optimized_params_for_this_run['commision'] = 0.0005
optimized_params_for_this_run['slippage'] = 0.001
out_of_sample_end_date_key = str(time.strftime("%<KEY>",time.localtime(out_of_sample_end_date)))
out_of_sample_start_date_with_buffer_key =\
str(time.strftime("%Y-%m-%d %H:%M:%S",time.localtime(out_of_sample_start_date_with_buffer)))
out_of_sample_start_date_key =\
str(time.strftime("%Y-%m-%d %H:%M:%S",time.localtime(out_of_sample_start_date)))
out_of_sample_key = out_of_sample_start_date_with_buffer_key +\
"_" + out_of_sample_end_date_key
# Run the out of sample Period with the optimized params
OutOfSampleResults = RunBackTest(coin_name\
,candle_freq\
,200000.0\
,out_of_sample_start_date_with_buffer\
,out_of_sample_end_date\
,optimized_params_for_this_run\
,shouldPlot\
,False)
print("The interval end dates and params are: ")
for param in out_of_sample_interval_params_for_this_run:
print(str(time.strftime("%Y-%m-%d %H:%M:%S", time.localtime(param[0]))) , param[1])
print("Out of sample testing for start date " + out_of_sample_start_date_key + " and" +\
" for the best : " + str(i) + "th param has finished!")
print("Buffered Start Date was : " + out_of_sample_start_date_with_buffer_key)
print()
print('***********************************')
print()
out_of_sample_result[out_of_sample_key] = OutOfSampleResults
out_of_sample_results.append(out_of_sample_result)
return out_of_sample_results
# -
# ## Walking Forward
# + pycharm={"is_executing": true}
tstart = time.mktime(time.strptime("20.01.2018 00:00:00", "%d.%m.%Y %H:%M:%S"));
tend = time.mktime(time.strptime("10.03.2019 11:05:02", "%d.%m.%Y %H:%M:%S"));
optimization_period = timedelta(days=200).total_seconds()
out_of_sample_period = timedelta(days=90).total_seconds()
start_time = time.time()
optimization_params={}
optimization_params['linear_reg_length'] = [10,1000]
optimization_params['commision'] = 0.0005
optimization_params['slippage'] = 0.001
WalkForwardAnalysisResultsPNLAll_LastYear = BlackBoxParallelWalkForwardAnalysis(tstart\
,tend\
,optimization_period\
,out_of_sample_period\
,optimization_params\
,100\
,100\
,4\
,opt_fun)
end_time = time.time()
print(str(time.strftime("%Y-%m-%d %H:%M:%S", time.localtime(end_time))))
print("Walk Forward Analysis Completed it took ")
diff = end_time - start_time
print(diff)
print("Seconds")
# + pycharm={"is_executing": true}
out_of_sample_period = timedelta(days=90).total_seconds()
interval_results_lastyear = generateAlternateOutOfSampleResultsWithInterval(WalkForwardAnalysisResultsPNLAll_LastYear['optimized_parameters']\
,out_of_sample_period,10, False)
# + pycharm={"is_executing": true}
# %matplotlib notebook
capital = 100000
value_to_plot = []
end_values = []
first_n = 10
for ind, result in enumerate(interval_results_lastyear):
if(ind>first_n): break
for key in result: # there is only 1 value
value_to_plot = ((np.array(result[key]['value_anlaysis']) - capital) /capital)*100
end_values.append(value_to_plot[-1])
plt.plot(value_to_plot, label='equity_curve' + str(ind))
plt.legend()
plt.show()
plt.figure()
plt.plot(end_values,'*')
plt.show()
print(end_values)
print('average: ' + str(sum(end_values)/len(end_values)))
print('min: ' + str(min(end_values)))
print('max: ' + str(max(end_values)))
| WalkForward/WalkForwardWorksheet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
from tqdm import tqdm
import os
import seaborn as sns
import sys
sys.path.insert(0,'/home/gsoc0/Adversarial_CapsNet_Pytorch/')
from model.net import *
from model.cnn_net import *
from utils.training import *
from data.data import *
# -
# ## Experiment Config
# +
model_path = os.path.join(os.getcwd(), "weights")
args = {
'DATASET_NAME':'mnist',
'num_classes':10,
'USE_CUDA': True if torch.cuda.is_available() else False,
'BATCH_SIZE': 512,
# 'N_EPOCHS': 100,
# 'LEARNING_RATE_1': 1e-2,
# 'LEARNING_RATE_2': 1e-2,
# 'WEIGHTDECAY':5e-6,
# #Scheduler
# 'USE_SCHEDULER':True,
# 'sched_milestones':[14,25,30,40,50],
# 'sched_gamma':0.1,
# #Loss Function
# 'LAMBDA_recon': 0.5,
# 'LAMBDA_class': 1,
##For Decoder
'num_features':160,
'LReLU_negative_slope':0.1,
'input_height':28,
'input_width':28,
'input_channel':1,
}
#Setting Default Cuda Device
torch.cuda.set_device(0)
# -
# ## Model Loading
# +
#Modified Reconstructor
class Reconstructor(nn.Module):
def __init__(self, net, args):
super(Reconstructor, self).__init__()
self.args = args
self.reconstraction_layers = net.decoder.reconstraction_layers
self.mean = torch.tensor(0.1307)
self.std = torch.tensor(0.3081)
if(args['USE_CUDA']):
self.mean = self.mean.cuda()
self.std = self.std.cuda()
self.unnormalize = UnNormalize(self.mean, self.std)
def forward(self, x, data):
with torch.no_grad():
classes = torch.sqrt((x ** 2).sum(2))
classes = F.softmax(classes.squeeze(), dim=1)
_, max_length_indices = classes.max(dim=1)
eye = torch.sparse.torch.eye(10)
if USE_CUDA:
eye = eye.cuda()
reconstructions_of_digits = []
list_of_indices = [max_length_indices]
for i, index_list in enumerate(list_of_indices):
masked = eye.index_select(dim=0, index=index_list.squeeze().data)
reconstructions_of_digits.append(self.reconstraction_layers((x * masked[:, :, None, None]).view(x.size(0), 1, -1)))
reconstructions_of_digits = torch.cat(reconstructions_of_digits, dim=1)
expanded_data = data.view(x.size(0), 1, -1).expand(-1, len(list_of_indices), -1)
unnormalized_data = self.unnormalize(expanded_data)
l2_distances = torch.square((unnormalized_data - reconstructions_of_digits)).sum(dim=(list(range(0, len(expanded_data.shape)))[2:]))
return l2_distances
def class_conditional_recon(self, x, data):
with torch.no_grad():
classes = torch.sqrt((x ** 2).sum(2))
classes = F.softmax(classes.squeeze(), dim=1)
_, max_length_indices = classes.max(dim=1)
eye = torch.sparse.torch.eye(10)
if USE_CUDA:
eye = eye.cuda()
reconstructions_of_digits = []
list_of_indices = [max_length_indices]
for i, index_list in enumerate(list_of_indices):
masked = eye.index_select(dim=0, index=index_list.squeeze().data)
reconstructions_of_digits.append(self.reconstraction_layers((x * masked[:, :, None, None]).view(x.size(0), 1, -1)))
reconstructions_of_digits = torch.cat(reconstructions_of_digits, dim=1)
expanded_data = data.view(x.size(0), 1, -1).expand(-1, len(list_of_indices), -1)
unnormalized_data = self.unnormalize(expanded_data)
l2_distances = torch.square((unnormalized_data - reconstructions_of_digits)).sum(dim=(list(range(0, len(expanded_data.shape)))[2:]))
return l2_distances
def imshow(img):
# img = img * 0.3081 + 0.1307 # unnormalize
npimg = img.cpu().numpy()
# plt.imshow(np.transpose(npimg)#, (1, 2, 0)))
plt.imshow(npimg)
plt.show()
# +
class Config:
def __init__(self, dataset='mnist'):
# CNN (cnn)
self.cnn_in_channels = 1
self.cnn_out_channels = 12
self.cnn_kernel_size = 15
# Primary Capsule (pc)
self.pc_num_capsules = 1
self.pc_in_channels = 12
self.pc_out_channels = 16
self.pc_kernel_size = 8
self.pc_num_routes = 7 * 7
# Digit Capsule 1 (dc)
self.dc_num_capsules = 49
self.dc_num_routes = 7 * 7
self.dc_in_channels = 16
self.dc_out_channels = 16 #1
# Digit Capsule 2 (dc)
self.dc_2_num_capsules = 10
self.dc_2_num_routes = 7 * 7
self.dc_2_in_channels = 16 #1
self.dc_2_out_channels = 16
# Decoder
self.input_width = 28
self.input_height = 28
torch.manual_seed(1)
config = Config()
net = CapsNet(args, config)
# capsule_net = torch.nn.DataParallel(capsule_net)
if args['USE_CUDA']:
net = net.cuda()
net.load_state_dict(torch.load(os.path.join(model_path, 'CapsNet_mnist.pth'), map_location='cpu'))
# -
# ## Loading Dataset
trainloader, testloader = dataset(args)
for data in trainloader:
print(data[0][0].min())
print(data[0][0].max())
break
# ## Untargeted WhiteBox Attack for CapsuleNet
a = torch.tensor([], dtype=torch.int16)
b = torch.tensor([1,2,3], dtype=torch.int16)
torch.cat((a,b))
# +
import matplotlib.pyplot as plt
from advertorch.attacks import LinfPGDAttack, GradientSignAttack, CarliniWagnerL2Attack, LinfBasicIterativeAttack
class Model_for_Adversary(nn.Module):
def __init__(self, net):
super(Model_for_Adversary, self).__init__()
self.net = net
def forward(self, x):
output, recons, masked = self.net(x)
classes = torch.sqrt((output ** 2).sum(2)).squeeze()
return classes
def WhiteBox_Attacks_Targeted(net, dataloader, adversary_dict, args):
net.eval()
n_batch = len(dataloader)
Success_Rate = {key:0.0 for key in adversary_dict.keys()}
Undetected_Rate = {key:0.0 for key in adversary_dict.keys()}
Und_l2 = {key:torch.tensor([],dtype=torch.int16).cuda() for key in adversary_dict.keys()}
for adversary in adversary_dict.keys():
for batch_id, (data, labels) in enumerate(tqdm(dataloader)):
target = torch.randint(0,10,size=(labels.size(0),), dtype=labels.dtype).cuda()
if(args['USE_CUDA']):
data, labels = data.cuda(), labels.cuda()
adv_data = adversary_dict[adversary].perturb(data, target)
with torch.no_grad():
output, reconstructions, max_length_indices = net(adv_data)
unnormalized_data = net.decoder.unnormalize(adv_data)
l2_distances = ((reconstructions.view(adv_data.size(0),-1)-unnormalized_data.view(adv_data.size(0), -1))**2).sum(1).squeeze().detach()
theta = np.percentile(l2_distances.cpu().numpy(), 95)
if(adversary=='Clean'):
Und_l2[adversary] = torch.cat((Und_l2[adversary],l2_distances))
else:
Und_l2[adversary] = torch.cat((Und_l2[adversary],l2_distances[max_length_indices == target]))
Success_Rate[adversary]+=torch.sum(max_length_indices == target).item()
Undetected_Rate[adversary]+=torch.sum(l2_distances[max_length_indices == target]<=theta).item()
Und_l2[adversary] = Und_l2[adversary].cpu().numpy()
Success_Rate[adversary]/=100
Undetected_Rate[adversary]/=100
return Success_Rate, Undetected_Rate, Und_l2
def WhiteBox_Attacks_Untargeted(net, dataloader, adversary_dict, args):
net.eval()
n_batch = len(dataloader)
Success_Rate = {key:0.0 for key in adversary_dict.keys()}
Undetected_Rate = {key:0.0 for key in adversary_dict.keys()}
Und_l2 = {key:torch.tensor([],dtype=torch.int16).cuda() for key in adversary_dict.keys()}
for adversary in adversary_dict.keys():
for batch_id, (data, labels) in enumerate(tqdm(dataloader)):
if(args['USE_CUDA']):
data, labels = data.cuda(), labels.cuda()
adv_data = adversary_dict[adversary].perturb(data)
with torch.no_grad():
output, reconstructions, max_length_indices = net(adv_data)
unnormalized_data = net.decoder.unnormalize(adv_data)
l2_distances = ((reconstructions.view(adv_data.size(0),-1)-unnormalized_data.view(adv_data.size(0), -1))**2).sum(1).squeeze().detach()
theta = np.percentile(l2_distances.cpu().numpy(), 95)
if(adversary=='Clean'):
Und_l2[adversary] = torch.cat((Und_l2[adversary],l2_distances))
else:
Und_l2[adversary] = torch.cat((Und_l2[adversary],l2_distances[max_length_indices != labels]))
Success_Rate[adversary]+=torch.sum(max_length_indices != labels).item()
Undetected_Rate[adversary]+=torch.sum(l2_distances[max_length_indices!=labels]<=theta).item()
# print(Success_Rate[adversary])
# print(Undetected_Rate[adversary])
# print(theta)
Und_l2[adversary] = Und_l2[adversary].cpu().numpy()
Success_Rate[adversary]/=100
Undetected_Rate[adversary]/=100
return Success_Rate, Undetected_Rate, Und_l2
# +
from advertorch.attacks.base import Attack, LabelMixin
from advertorch.utils import clamp
class CleanAttack(Attack, LabelMixin):
def __init__(self, clip_min=0., clip_max=1.):
super(CleanAttack, self).__init__(None,None,clip_min, clip_max)
def perturb(self, x, y=None):
return x
def Attack_Table(L2_Distances, theta_percentile):
theta = np.percentile(L2_Distances['Clean'], theta_percentile)
Success_Rate = {key:0.0 for key in L2_Distances.keys()}
Undetected_Rate = {key:0.0 for key in L2_Distances.keys()}
for adversary in L2_Distances.keys():
Success_Rate[adversary] = L2_Distances[adversary].shape[0]/ 100
Undetected_Rate[adversary] = np.sum(L2_Distances[adversary]<=theta).item()/100
# print("---------------------------------------------------------")
print(theta, ":", theta_percentile ," ", Success_Rate['Clean'],"/",Undetected_Rate['Clean'],"----",Success_Rate['FGSM'],"/",Undetected_Rate['FGSM'],"----",Success_Rate['BIM'],"/",Undetected_Rate['BIM'],"----",Success_Rate['PGD'],"/",Undetected_Rate['PGD'])
# +
model_for_adversary = Model_for_Adversary(net)
linf_eps = 0.3
fgsm_step = 0.05
bim_pgd_step = 0.01
adversary_dict = {}
adversary_dict['Clean'] = CleanAttack(clip_min=-0.4242, clip_max=2.8215)
adversary_dict['PGD'] = LinfPGDAttack(
model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081),
nb_iter=100, eps_iter=(bim_pgd_step/0.3081), rand_init=True, clip_min=-0.4242, clip_max=2.8215,
targeted=False)
adversary_dict['FGSM'] = GradientSignAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(fgsm_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=False)
adversary_dict['BIM'] = LinfBasicIterativeAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081), nb_iter=100, eps_iter=(bim_pgd_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=False)
# adversary_dict['CW'] = CarliniWagnerL2Attack(model_for_adversary, num_classes=args['num_classes'], confidence=0, targeted=False, learning_rate=0.01, binary_search_steps=9, max_iterations=1000, abort_early=True, initial_const=0.001, clip_min=-0.4242, clip_max=2.8215, loss_fn=None)
Success_Rate, Undetected_Rate, Und_L2 = WhiteBox_Attacks_Untargeted(net, testloader, adversary_dict, args)
# +
print(np.percentile(Und_L2['Clean'],95))
for adversary in adversary_dict.keys():
if(adversary!='Clean'):
x = np.linspace(0, 100, 20)
# z = np.array([np.sum(Und_L2['Clean']>((i_x/100))*(np.percentile(Und_L2['Clean'],95)*2)).item()/100 for i_x in x])
y = np.array([np.sum(Und_L2[adversary]<=((i_x/100))*(np.percentile(Und_L2['Clean'],95)*2)).item()/100 for i_x in x])
plt.plot(x, y, 'go-', label='line1', linewidth=2)
# -
Und_L2_Untargeted_Capsnet = Und_L2
print('theta-----------percentile----Clean--------------FGSM------------BIM------------PGD')
for i in range(0,101):
Attack_Table(Und_L2, i)
# +
model_for_adversary = Model_for_Adversary(net)
linf_eps = 0.3
fgsm_step = 0.05
bim_pgd_step = 0.01
adversary_dict = {}
adversary_dict['Clean'] = CleanAttack(clip_min=-0.4242, clip_max=2.8215)
adversary_dict['PGD'] = LinfPGDAttack(
model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081),
nb_iter=100, eps_iter=(bim_pgd_step/0.3081), rand_init=True, clip_min=-0.4242, clip_max=2.8215,
targeted=True)
adversary_dict['FGSM'] = GradientSignAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(fgsm_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=True)
adversary_dict['BIM'] = LinfBasicIterativeAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081), nb_iter=100, eps_iter=(bim_pgd_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=True)
# adversary_dict['CW'] = CarliniWagnerL2Attack(model_for_adversary, num_classes=args['num_classes'], confidence=0, targeted=False, learning_rate=0.01, binary_search_steps=9, max_iterations=1000, abort_early=True, initial_const=0.001, clip_min=-0.4242, clip_max=2.8215, loss_fn=None)
Success_Rate, Undetected_Rate, Und_L2 = WhiteBox_Attacks_Targeted(net, testloader, adversary_dict, args)
# -
for adversary in adversary_dict.keys():
if(adversary!='Clean'):
x = np.linspace(0, 100, 20)
z = np.array([np.sum(Und_L2['Clean']>((i_x/100))*((np.percentile(Und_L2['Clean'],95)))).item()/100 for i_x in x])
y = np.array([np.sum(Und_L2[adversary]<=((i_x/100))*((np.percentile(Und_L2['Clean'],95)))).item()/100 for i_x in x])
plt.plot(z, y, 'go-', label='line1', linewidth=2)
Und_L2_Targeted_Capsnet = Und_L2
for i in range(0,100):
Attack_Table(Und_L2, i)
np.save('Und_L2_Targeted_Capsnet.npy', Und_L2_Targeted_Capsnet)
# np.save('Und_L2_Untargeted_Capsnet.npy', Und_L2_Untargeted_Capsnet)
# ## Histograms for CNN + CR
# +
args = {
'DATASET_NAME':'mnist',
'num_classes':10,
'USE_CUDA': True if torch.cuda.is_available() else False,
'BATCH_SIZE': 256,
'N_EPOCHS': 100,
'LEARNING_RATE_1': 1e-2,
'LEARNING_RATE_2': 1e-2,
'WEIGHTDECAY':5e-6,
#Scheduler
'USE_SCHEDULER':True,
'sched_milestones':[8,20,30,40,50],
'sched_gamma':0.1,
#Loss Function
'LAMBDA_recon': 0.5,
'LAMBDA_class': 1,
#For Decoder
'num_features':160,
'LReLU_negative_slope':0.1,
'input_height':28,
'input_width':28,
'input_channel':1,
'type':'plusCR',
}
# +
class Config:
def __init__(self, dataset='mnist'):
# CONV1
self.conv1_in = 1
self.conv1_out = 12
self.conv1_kernel_size = 15
# CONV2
self.conv2_in = 12
self.conv2_out = 16
self.conv2_kernel_size = 8
# FC1
self.fc1_in = 7 * 7 * 16
self.fc1_out = 784
# FC1
self.fc2_in = 784
self.fc2_out = 160
torch.manual_seed(2)
config = Config()
net = CNNnet(args, config)
# net = torch.nn.DataParallel(net)
net.load_state_dict(torch.load(os.path.join(model_path, 'CNNplusCR_mnist.pth'), map_location='cpu'))
if args['USE_CUDA']:
net = net.cuda()
# +
import matplotlib.pyplot as plt
from advertorch.attacks import LinfPGDAttack, GradientSignAttack, CarliniWagnerL2Attack, LinfBasicIterativeAttack
class Model_for_Adversary(nn.Module):
def __init__(self, net):
super(Model_for_Adversary, self).__init__()
self.net = net
def forward(self, x):
output, recons, masked = self.net(x)
classes = output.sum(2)
return classes
# +
from advertorch.attacks.base import Attack, LabelMixin
from advertorch.utils import clamp
class CleanAttack(Attack, LabelMixin):
def __init__(self, clip_min=0., clip_max=1.):
super(CleanAttack, self).__init__(None, None, clip_min, clip_max)
self.clip_min = clip_min
self.clip_max = clip_max
self.targeted = False
def perturb(self, x, y=None):
return x
model_for_adversary = Model_for_Adversary(net)
linf_eps = 0.3
fgsm_step = 0.05
bim_pgd_step = 0.01
adversary_dict = {}
adversary_dict['Clean'] = CleanAttack(clip_min=-0.4242, clip_max=2.8215)
adversary_dict['PGD'] = LinfPGDAttack(
model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081),
nb_iter=1000, eps_iter=(bim_pgd_step/0.3081), rand_init=True, clip_min=-0.4242, clip_max=2.8215,
targeted=False)
adversary_dict['FGSM'] = GradientSignAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(fgsm_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=False)
adversary_dict['BIM'] = LinfBasicIterativeAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081), nb_iter=1000, eps_iter=(bim_pgd_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=False)
# adversary_dict['CW'] = CarliniWagnerL2Attack(model_for_adversary, num_classes=args['num_classes'], confidence=0, targeted=False, learning_rate=0.01, binary_search_steps=9, max_iterations=1000, abort_early=True, initial_const=0.001, clip_min=-0.4242, clip_max=2.8215, loss_fn=None)
Success_Rate, Undetected_Rate, Und_L2 = WhiteBox_Attacks_Untargeted(net, testloader, adversary_dict, args)
# -
Und_L2_Untargeted_plusCR = Und_L2
for adversary in adversary_dict.keys():
if(adversary!='Clean'):
x = np.linspace(0, 100, 20)
z = np.array([np.sum(Und_L2['Clean']>((i_x/100))*(np.percentile(Und_L2['Clean'],95))).item()/100 for i_x in x])
y = np.array([np.sum(Und_L2[adversary]<=((i_x/100))*(np.percentile(Und_L2['Clean'],95))).item()/100 for i_x in x])
plt.plot(z, y, 'go-', label='line1', linewidth=2)
for i in range(0,100):
Attack_Table(Und_L2, i)
# +
model_for_adversary = Model_for_Adversary(net)
linf_eps = 0.3
fgsm_step = 0.05
bim_pgd_step = 0.01
adversary_dict = {}
adversary_dict['Clean'] = CleanAttack(clip_min=-0.4242, clip_max=2.8215)
adversary_dict['PGD'] = LinfPGDAttack(
model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081),
nb_iter=1000, eps_iter=(bim_pgd_step/0.3081), rand_init=True, clip_min=-0.4242, clip_max=2.8215,
targeted=True)
adversary_dict['FGSM'] = GradientSignAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(fgsm_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=True)
adversary_dict['BIM'] = LinfBasicIterativeAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081), nb_iter=1000, eps_iter=(bim_pgd_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=True)
# adversary_dict['CW'] = CarliniWagnerL2Attack(model_for_adversary, num_classes=args['num_classes'], confidence=0, targeted=False, learning_rate=0.01, binary_search_steps=9, max_iterations=1000, abort_early=True, initial_const=0.001, clip_min=-0.4242, clip_max=2.8215, loss_fn=None)
Success_Rate, Undetected_Rate, Und_L2 = WhiteBox_Attacks_Targeted(net, testloader, adversary_dict, args)
# -
Und_L2_Targeted_plusCR = Und_L2
for adversary in adversary_dict.keys():
if(adversary!='Clean'):
x = np.linspace(0, 100, 20)
z = np.array([np.sum(Und_L2['Clean']>((i_x/100))*(np.percentile(Und_L2['Clean'],95))).item()/100 for i_x in x])
y = np.array([np.sum(Und_L2[adversary]<=((i_x/100))*(np.percentile(Und_L2['Clean'],95))).item()/100 for i_x in x])
plt.plot(z, y, 'go-', label='line1', linewidth=2)
for i in range(0,100):
Attack_Table(Und_L2, i)
np.save('Und_L2_Targeted_plusCR.npy', Und_L2_Targeted_plusCR)
np.save('Und_L2_Untargeted_plusCR.npy', Und_L2_Untargeted_plusCR)
# ## Histograms for CNN plus R
# +
args = {
'DATASET_NAME':'mnist',
'num_classes':10,
'USE_CUDA': True if torch.cuda.is_available() else False,
'BATCH_SIZE': 256,
'N_EPOCHS': 100,
'LEARNING_RATE_1': 1e-2,
'LEARNING_RATE_2': 1e-2,
'WEIGHTDECAY':5e-6,
#Scheduler
'USE_SCHEDULER':True,
'sched_milestones':[8,20,30,40,50],
'sched_gamma':0.1,
#Loss Function
'LAMBDA_recon': 0.5,
'LAMBDA_class': 1,
#For Decoder
'num_features':160,
'LReLU_negative_slope':0.1,
'input_height':28,
'input_width':28,
'input_channel':1,
'type':'plusR',
}
# +
class Config:
def __init__(self, dataset='mnist'):
# CONV1
self.conv1_in = 1
self.conv1_out = 12
self.conv1_kernel_size = 15
# CONV2
self.conv2_in = 12
self.conv2_out = 16
self.conv2_kernel_size = 8
# FC1
self.fc1_in = 7 * 7 * 16
self.fc1_out = 784
# FC1
self.fc2_in = 784
self.fc2_out = 160
torch.manual_seed(2)
config = Config()
net = CNNnet(args, config)
# net = torch.nn.DataParallel(net)
net.load_state_dict(torch.load(os.path.join(model_path, 'CNNplusR_mnist.pth'), map_location='cpu'))
if args['USE_CUDA']:
net = net.cuda()
# +
model_for_adversary = Model_for_Adversary(net)
linf_eps = 0.3
fgsm_step = 0.05
bim_pgd_step = 0.01
adversary_dict = {}
adversary_dict['Clean'] = CleanAttack(clip_min=-0.4242, clip_max=2.8215)
adversary_dict['PGD'] = LinfPGDAttack(
model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081),
nb_iter=1000, eps_iter=(bim_pgd_step/0.3081), rand_init=True, clip_min=-0.4242, clip_max=2.8215,
targeted=False)
adversary_dict['FGSM'] = GradientSignAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(fgsm_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=False)
adversary_dict['BIM'] = LinfBasicIterativeAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081), nb_iter=1000, eps_iter=(bim_pgd_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=False)
# adversary_dict['CW'] = CarliniWagnerL2Attack(model_for_adversary, num_classes=args['num_classes'], confidence=0, targeted=False, learning_rate=0.01, binary_search_steps=9, max_iterations=1000, abort_early=True, initial_const=0.001, clip_min=-0.4242, clip_max=2.8215, loss_fn=None)
Success_Rate, Undetected_Rate, Und_L2 = WhiteBox_Attacks_Untargeted(net, testloader, adversary_dict, args)
# -
Und_L2_Untargeted_plusR = Und_L2
for adversary in adversary_dict.keys():
if(adversary!='Clean'):
x = np.linspace(0, 100, 20)
z = np.array([np.sum(Und_L2['Clean']>((i_x/100))*(np.percentile(Und_L2['Clean'],95))).item()/100 for i_x in x])
y = np.array([np.sum(Und_L2[adversary]<=((i_x/100))*(np.percentile(Und_L2['Clean'],95))).item()/100 for i_x in x])
plt.plot(z, y, 'go-', label='line1', linewidth=2)
for i in range(0,100):
Attack_Table(Und_L2, i)
# +
model_for_adversary = Model_for_Adversary(net)
linf_eps = 0.3
fgsm_step = 0.05
bim_pgd_step = 0.01
adversary_dict = {}
adversary_dict['Clean'] = CleanAttack(clip_min=-0.4242, clip_max=2.8215)
adversary_dict['PGD'] = LinfPGDAttack(
model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081),
nb_iter=1000, eps_iter=(bim_pgd_step/0.3081), rand_init=True, clip_min=-0.4242, clip_max=2.8215,
targeted=True)
adversary_dict['FGSM'] = GradientSignAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(fgsm_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=True)
adversary_dict['BIM'] = LinfBasicIterativeAttack(model_for_adversary, loss_fn=nn.CrossEntropyLoss(reduction="sum"), eps=(linf_eps/0.3081), nb_iter=1000, eps_iter=(bim_pgd_step/0.3081), clip_min=-0.4242, clip_max=2.8215, targeted=True)
# adversary_dict['CW'] = CarliniWagnerL2Attack(model_for_adversary, num_classes=args['num_classes'], confidence=0, targeted=False, learning_rate=0.01, binary_search_steps=9, max_iterations=1000, abort_early=True, initial_const=0.001, clip_min=-0.4242, clip_max=2.8215, loss_fn=None)
Success_Rate, Undetected_Rate, Und_L2 = WhiteBox_Attacks_Targeted(net, testloader, adversary_dict, args)
# -
Und_L2_Targeted_plusR = Und_L2
for adversary in adversary_dict.keys():
if(adversary!='Clean'):
x = np.linspace(0, 100, 20)
z = np.array([np.sum(Und_L2['Clean']>((i_x/100))*(np.percentile(Und_L2['Clean'],95))).item()/100 for i_x in x])
y = np.array([np.sum(Und_L2[adversary]<=((i_x/100))*(np.percentile(Und_L2['Clean'],95))).item()/100 for i_x in x])
plt.plot(z, y, 'go-', label='line1', linewidth=2)
for i in range(0,100):
Attack_Table(Und_L2, i)
# +
# np.save('Und_L2_Targeted_plusR.npy', Und_L2_Targeted_plusR)
# np.save('Und_L2_Untargeted_plusR.npy', Und_L2_Untargeted_plusR)
# +
def plot_Ud_FPR(adversary, Und_L2_list):
color_list = ['green', 'orange','blue']
model_list = ['CapsNet', 'plusCR', 'plusR']
for i,Und_L2 in enumerate(Und_L2_list):
if(adversary!='Clean'):
x = np.linspace(0, 100, 50)
# z = np.array([((1 - i_x/100))*(np.percentile(Und_L2['Clean'],95)*2).item() for i_x in x])
# z = np.array([np.sum(Und_L2['Clean']>((i_x/100))*(np.percentile(Und_L2['Clean'],100))).item()/100 for i_x in x])
# y = np.array([np.sum(Und_L2[adversary]<=((i_x/100))*(np.percentile(Und_L2['Clean'],100))).item()/100 for i_x in x])
z = np.array([np.sum(Und_L2['Clean']>((i_x/100))*(80)).item()/100 for i_x in x])
y = np.array([np.sum(Und_L2[adversary]<=((i_x/100))*(80)).item()/100 for i_x in x])
plt.plot(z, y,color=color_list[i], linestyle='-', marker='o', label=model_list[i], linewidth=1)
plot_Ud_FPR('PGD',[Und_L2_Targeted_Capsnet,Und_L2_Targeted_plusCR,Und_L2_Targeted_plusR])
# -
plot_Ud_FPR('BIM',[Und_L2_Targeted_Capsnet,Und_L2_Targeted_plusCR,Und_L2_Targeted_plusR])
plot_Ud_FPR('FGSM',[Und_L2_Targeted_Capsnet,Und_L2_Targeted_plusCR,Und_L2_Targeted_plusR])
# +
Und_L2_Targeted_Capsnet = np.load('Und_L2_Targeted_Capsnet.npy', allow_pickle=True).item()
Und_L2_Untargeted_Capsnet = np.load('Und_L2_Untargeted_Capsnet.npy', allow_pickle=True).item()
# Und_L2_Targeted_plusCR = np.load('Und_L2_Targeted_plusCR.npy', allow_pickle=True).item()
# Und_L2_Untargeted_plusCR = np.load('Und_L2_Untargeted_plusCR.npy', allow_pickle=True).item()
# Und_L2_Targeted_plusR = np.load('Und_L2_Targeted_plusR.npy', allow_pickle=True).item()
# Und_L2_Untargeted_plusR = np.load('Und_L2_Untargeted_plusR.npy', allow_pickle=True).item()
# +
from tabulate import tabulate
def Print_Table(Und_L2_dict, percentile=95, theta=None):
print('Percentile :', percentile)
table = []
for network in Und_L2_dict.keys():
if theta is None:
theta = np.percentile(Und_L2_dict[network]['Clean'], percentile)
else:
theta = theta
Success_Rate = {key:0.0 for key in Und_L2_dict[network].keys()}
Undetected_Rate = {key:0.0 for key in Und_L2_dict[network].keys()}
for adversary in Und_L2_dict[network].keys():
Success_Rate[adversary] = Und_L2_dict[network][adversary].shape[0]/ 100
Undetected_Rate[adversary] = np.sum(Und_L2_dict[network][adversary]<=theta).item()/100
table.append([network, str(Success_Rate['FGSM'])+"/"+str(Undetected_Rate['FGSM']),str(Success_Rate['BIM'])+"/"+str(Undetected_Rate['BIM']),str(Success_Rate['PGD'])+"/"+str(Undetected_Rate['PGD'])])
print(tabulate(table, headers=['Networks', 'FGSM', 'BIM', 'PGD']))
Und_L2_dict = {'CapsNet':Und_L2_Untargeted_Capsnet, 'CNN+CR':Und_L2_Untargeted_plusCR, 'CNN+R':Und_L2_Untargeted_plusR}
Print_Table(Und_L2_dict)
# -
Und_L2_dict = {'CapsNet':Und_L2_Targeted_Capsnet, 'CNN+CR':Und_L2_Targeted_plusCR, 'CNN+R':Und_L2_Targeted_plusR}
Print_Table(Und_L2_dict)
Und_L2_dict = {'CapsNet':Und_L2_Untargeted_Capsnet, 'CNN+CR':Und_L2_Untargeted_plusCR, 'CNN+R':Und_L2_Untargeted_plusR}
Print_Table(Und_L2_dict, theta=45)
Und_L2_dict = {'CapsNet':Und_L2_Targeted_Capsnet, 'CNN+CR':Und_L2_Targeted_plusCR, 'CNN+R':Und_L2_Targeted_plusR}
Print_Table(Und_L2_dict, theta=45)
| Standard-Attacks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 9. Thresholding exercises
# 1. Import the relevant packages: numpy, matplotlib and skimage
# 2. Import the same image as in the main notebook (myoblast.tif)
# 3. Create a new variable called ```image_green``` that contains only the green channel by slicing the array
# 4. Plot this image
# 5. Plot a histogram of the image
# 6. Try to determine an appropriate threshold to recover the large structures in the image, create the corresponding mask and plot it
# 7. There are other algorithms for thresholding than the one of Otsu. Try to find one in the [Scikit-image documentation](http://scikit-image.org/docs/dev/api/skimage.filters.html) and to use it
| e09-Thresholding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import scipy.optimize as opt
from scipy import interpolate
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from PIL import Image
# %matplotlib notebook
def get_image(image_path, scale):
"""Get a numpy array of an image so that one can access values[x][y]."""
image = Image.open(image_path, "r")
width, height = image.size
l = max(width, height)
c = l/scale
x = np.linspace(0, c, l)
x, y = np.meshgrid(x, x)
pixel_values = list(image.getdata())
if image.mode == "RGB":
channels = 3
elif image.mode == "L":
channels = 1
else:
print("Unknown mode: %s" % image.mode)
return None
pixel_values = np.array(pixel_values).reshape((width, height, channels))
return pixel_values, x, y, width, height, l, c
im, x, y, width, height, l, c = get_image("images/image3.jpg", 1)
R = im[:,:,0]
R = R.reshape(width, height)
buffer = np.zeros((l-width,l))
print(R.shape, buffer.shape)
R = np.concatenate((R,buffer), axis=0)
#R = R.reshape(R.shape[0], R.shape[1],1)
print(R.shape)
plt.imshow(R,aspect='auto')
plt.show()
p = R[:,:]
print(p.shape)
#im = p[:,:,0]#+p[:,:,1]+p[:,:,2]
plt.imshow(p)
plt.show()
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(x, y, p, cmap="magma")
#plt.savefig("test2.jpg")
plt.show()
def twoD_Gaussian(mesh, amplitude, x_0, y_0, sigma, offset):
(x, y) = mesh
x_0 = float(x_0)
y_0 = float(y_0)
g = offset + amplitude*np.exp(-0.5*(((x-x_0)**2) + ((y-y_0)**2))/(sigma**2))
return g.ravel()
data = R.ravel()
plt.figure()
plt.imshow(data.reshape(l,l), extent=[0,c,0,c], cmap=plt.cm.jet)
plt.colorbar()
initial_guess = (150,c/2,c/2,1,100)
popt, pcov = opt.curve_fit(twoD_Gaussian, (x, y), data, p0=initial_guess)
data_fitted = twoD_Gaussian((x, y), *popt)
print(popt)
fig, ax = plt.subplots(1, 1)
ax.imshow(data.reshape(l, l), cmap=plt.cm.jet, extent=[0,c,0,c])
ax.contour(x, y, data_fitted.reshape(l, l), 8, colors='black')
plt.show()
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(x, y, data_fitted.reshape(l, l), cmap="magma")
plt.savefig("test1.jpg")
#w(z)
print(np.sqrt(4*(popt[3])**2))
| Lab_1_-_Data_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [binaryen](https://github.com/WebAssembly/binaryen) is a Compiler infrastructure and toolchain for WASM
# # Installation
#
# ```shell
# # cd ~/local
# git clone https://github.com/WebAssembly/binaryen.git binaryen-git
# # cd binaryen-git
# # mkdir _build
# # cd _build
# cmake .. -DCMAKE_BUILD_TYPE=Release -DCMAKE_INSTALL_PREFIX=/home/obs/local/binaryen/
# make
# make install
# ```
# # wasm-dis
#
# Dissasemble and print to text for wasm file:
# ```shell
# wasm-dis fact.wasm
# ```
#
# Or save it to a file
# ```shell
# wasm-dis fact.wasm -o fact.wat
# ```
# # wasm-opt
#
# Optimize wasm binary code:
#
# ```shell
# wasm-opt -O3 input.wasm -o output.wasm
# ```
#
# Optimisations passes: ```-O0``` to ```-O4```, ```-Os``` and ```-Oz``` to optimize size
| refs/wasm/binaryen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Practical 3: Modules and Functions - Introducing Numpy!
#
# <div class="alert alert-block alert-success">
# <b>Objectives:</b> In this practical the overarching objective is for you to practice using modules and functions. Following on from a discussion in class, this will be done through 4 sections in this notebook, each with its own exercise for you to practice:
#
# - 1) [Part 1: Learning how to use the Numpy module](#Part1)
# * [Exercise 1: Create a 2D array and populate with specific values](#Exercise1)
# - 2) [Part 2: Using conditional statements to modify values in an array using Numpy](#Part2)
# * [Exercise 2: Create a 2D array and populate with values according to an equation and condition](#Exercise2)
# - 3) [Part 3: Creating your own function](#Part3)
# * [Exercise 3: Create a spooky type-writer](#Exercise3)
# * [Exercise 4: Build a function that creates a 2D surface plot of a scalar function](#Exercise4)
# - 4) [Part 4: Numpy operations on entire arrays](#Part4)
# * [Exercise 5: Re-create a function that creates a 2D surface plot of a scalar function](#Exercise5)
#
# As with our other notebooks, we will provide you with a template for plotting the results. Also please note that you should not feel pressured to complete every exercise in class. These practicals are designed for you to take outside of class and continue working on them. Do try to finish as much as you can whilst we are here to help. Proposed solutions to all exercises can be found in the 'solutions' folder.
# </div>
# ## First, why modules and functions?
#
# As we saw in the lecture for today, a typical computer program is contructed in a modular way. This is beneficial as your program gets bigger. It also allows you to develop and test routines seperately, making improvements or upgrades in a clear and transparent way.
#
# A module is a file containing Python definitions and statements. The file name is the module name with the suffix <code> .py </code> appended. It will also likely contain functions we wish to use. For example, if we want to use a module we would use the following command:
#
# ```python
# import *module name*
# ```
# Where the module name is the same as the file
#
# > *module name.py*
#
# and, if not in the same directory, can be found by the Python interpreter. We will not cover this here, but you can also specify the location of modules you write yourself by changing what is called the Python path or by importing another module that adds the location in the code directly.
#
# From this point on in our course, will be using the [Numpy](https://numpy.org) module. We can import it by including the following statement in our program or by writing this into the command line Python shell:
import numpy
# If you position your mouse in the above box and then click the <code> Run </code> button from the notebook menu, nothing appears to have happened. But it has! We now have access to all of Numpy's functionality. We can also import any module and change its name. This is useful if we constantly want to write and refer to a module. If you search for Python examples online, you will invariably see the following:
import numpy as np
# This is just common practice and proves to be much easier when using lots of functions held within Numpy, which we will get to shortly. We will also be using a number of other widely used modules in our course, including:
#
# - [Pandas](https://pandas.pydata.org): A module dedicated to data structures and data analysis tools.
# - [Scikit-learn](https://scikit-learn.org/stable/): A module dedicate to developing and deploying machine learning functions
# - [Scipy](https://www.scipy.org): (pronounced “<NAME>ie”) is a Python-based ecosystem of open-source software for mathematics, science, and engineering [including Numpy]
# - [Matplotlib](https://matplotlib.org): A module used for plotting, which we will use throughout the course.
#
# On the plotting module, we will not provide specific training on this but provide you with templates that cover most basic examples from line, scatter to 3D plots. You can take these templates and modify them for your own benefit. You can also search for help online or via the official Matplotlib webpage: https://matplotlib.org
#
# We normally import all of the modules we need at the start of any program. For example, if we want to import all of the above I would write:
import numpy as np
import pandas as pd
import sklearn #the name of the module is different from the official title, but the .py will have this name
import matplotlib
# Sometimes we do not want to import an entire module if we are only using one function out of many. As an example, in the Scikit-learn package I might only want to import a function that allows me to perform a technique known as K-means clustering. By checking the online documentation, I find that I should import this as:
from sklearn.cluster import KMeans
# Does the syntax make sense? The use of '.' is important. This is allowing me to access functions within the entire package. We will use this today as we explore the functionality of the Numpy package.
# # Part 1: Learning how to use the Numpy module <a name="Part1"></a>
#
# Numpy is a package used for numerical computations. It is used inplace of the vanilla Python approaches, as we used in the first lecture, on account of its speed and flexible functionality. For the remaining part of the course we will use Numpy to create and manipulate arrays. Let us go through some example and practice assigments on creating and manipulating 1D and 2D arrays
#
# ### 1.1) Creating 1D arrays
#
# In the following code we import Numpy and use it to intialise two 1D arrays. One will contain integers, the other floating points. Please note that, unlike lists and tuples in Python, we **cannot** blend data types in one array via Numpy. This is ok for our course, however, and related to the efficiency of Numpy arrays. Please see the code snippet below and notice where we use the **'.'** extension to the imported packaged to access a function. We will write our own functions later on in this practical:
# +
# Lets import the numpy package and call it np
import numpy as np
# We can use an internal Numpy function to create a 1D array of fixed length. Lets say we want to create arrays
# with 15 elements. For this Im going to use the function numpy.zeros() to create an array filled with 0's.
# The official documentation for this function can be found here: https://docs.scipy.org/doc/numpy/reference/generated/numpy.zeros.html
# Lets look at our example for now
# Im going to use my own variable names for each array. This dosnt matter:
Integer_array = np.zeros((15,), dtype=int)
Float_array = np.zeros((15,), dtype=float)
# -
# Before we print these arrays to the screen, let us see what has happened here. The function **numpy.zeros()** is expecting at least two bits of information
#
# > numpy.zeros((rows,columns), type of data)
#
# Where, in our case, we have specified we want 15 entries but only in one dimension as we have ignored the 'columns' variable. If we wanted to specify a 2D array, or matrix, as we will do later with 15 rows and 4 columns, we would write:
#
# > Integer_array = np.zeros((15,4), dtype=int)
#
# Notice we also specified our data type in each case. The first uses **'dtype=int'** and the second **'dtype=float'**. The function **expects** this syntax, as you might see in the official documentation. Let us now print our arrays to the screen to see what we have created. The notebook will automatically run code in each block when we first load it. However if you change anything, click inside the box and then click the 'Run' button above:
# +
# Simply print the arrays to screen
print("The integer array looks like > ",Integer_array)
print("The floating point array looks like > ",Float_array)
# You should be able to see a difference in format, thus precision.
# -
# ### 1.2) 2D Numpy arrays and accessing/changing values
#
# Now we can practice creating 2D arrays but also accessing and modifying/creating values. Let us create a 2D matrix that has 3 rows and 3 columns and then put a floating point number in every cell. Note the use of the **':'** operator that allows us to straddle the start and end indices of either a set of rows of columns.
#
#
#
#
# Now let us read through the code in the box below.
# +
# Create an empty matrix, of 3 rows and 3 columns, using the numpy.zeros() function
matrix = np.zeros((3,3), dtype=float)
# Now lets put the value 298.15 in each cell. There are multiple ways we could do this. I demonstrate 4
# below, where one method is significantly quicker than the other from a data entry perspective
# 1) Enter all values manually. 1st row and 1st column, 1st row and 2nd column etc etc
matrix[0,0] = 298.15; matrix[0,1] = 298.15; matrix[0,2] = 298.15
matrix[1,0] = 298.15; matrix[1,1] = 298.15; matrix[1,2] = 298.15
matrix[2,0] = 298.15; matrix[2,1] = 298.15; matrix[2,2] = 298.15
# Lets print the matrix to the screen
print(" first entry method result = ", matrix)
# 2) Assign the value to every column as a function of each row using the ':' operator
matrix = np.zeros((3,3), dtype=float)
matrix[0,0:3] = 298.15
matrix[1,0:3] = 298.15
matrix[2,0:3] = 298.15
# Lets print the matrix to the screen
print(" second entry method result = ", matrix)
# 3) Assign the value to every row as a function of each column using the ':' operator
matrix = np.zeros((3,3), dtype=float)
matrix[0:3,0] = 298.15
matrix[0:3,1] = 298.15
matrix[0:3,2] = 298.15
# Lets print the matrix to the screen
print(" third entry method result = ", matrix)
# 4) Assign every cell using the ':' operator in 2 dimensions
matrix = np.zeros((3,3), dtype=float)
matrix[0:3,0:3] = 298.15
# Lets print the matrix to the screen
print(" fourth entry method result = ", matrix)
# -
# Remember that, in Python, indexing starts at 0. For a 2D Numpy array, a nice schematic is provided by [O'Reilly](https://www.oreilly.com/library/view/python-for-data/9781449323592/ch04.html) below.
#
# <img src="images/Nump_array_indexing.png" alt="Numpy array indexing" style="width: 400px;"/>
#
#
#
# <div class="alert alert-block alert-success">
# <b> Exercise 1: Create a 2D array and add specific values. <a name="Exercise1"></a> </b> Now it is your turn to practice these operations. In the box below, you are tasked with the following:
#
# - Create an empty 2D array with 5 rows and 6 columns, containing floating point values.
# - For the first two rows the array will contain a value of 2.5
# - For the remaining rows to contain a value of 5.5.
# - Print the array to the screen so you can check your answer.
#
# Your answer should look like this:
#
# ```python
# matrix result = [[2.5 2.5 2.5 2.5 2.5 2.5]
# [2.5 2.5 2.5 2.5 2.5 2.5]
# [5.5 5.5 5.5 5.5 5.5 5.5]
# [5.5 5.5 5.5 5.5 5.5 5.5]
# [5.5 5.5 5.5 5.5 5.5 5.5]]
# ```
# </div>
#
# <div class="alert alert-block alert-warning">
# <b>HINT</b>: Rather than writing an entry for each cell, why not try using the technique demonstrated in example 2 and 3 above. </div>
# +
#------'INSERT CODE HERE'------
# Initialise an empty 2D array using the np.zeros function
practice_matrix = np.zeros((5,6), dtype=float)
# Populate the array with appropriate values
#------------------------------
# Lets print the matrix to the screen
print(" matrix result = ", practice_matrix)
# -
# # Part 2: Using conditional statements to modify values in an array using Numpy <a name="Part2"></a>
#
# In the first two practicals we came across **for loops** and conditional statements. We can combine these with Numpy arrays to start building a basis for numerical models. We can then turn a given combination of the above into a function that we can call over and over again. Let us start with an example. In the code snippet below, you can see a loop that populates a 1D Numpy array <code> Y </code> with 100 cells, according to the function:
#
# \begin{eqnarray}
# Y = X^{2.0}
# \end{eqnarray}
#
# Where <code> X </code> is an array that contains values from 1 to 100. This example combines a number of operations we have covered so far, including:
#
# - Initialising a 1D Numpy array of specific size, designed for floating point values
# - Creating a 'for' loop that cycles through integer values
# - Accessing a specific entry in a 1D array through an index value, remembering that Python indexing starts at 0.
# - Raising a number to a power
# - Print values to the screen
#
# When you run the code below you will see the output:
#
# ```python
# the value of the 98th component of x is 98.0
# the value of the 98th component of y is 9604.0
# the value of the 99th component of x is 99.0
# the value of the 99th component of y is 9801.0
# the value of the 100th component of x is 100.0
# the value of the 100th component of y is 10000.0
# ```
# +
# First create an empty array for both x and y
x = np.zeros((100,), dtype=float)
y = np.zeros((100,), dtype=float)
# Now implement a *for* loop that cycles through 100 values and creates the X and thus Y values
for step in range(100):
x[step] = step+1 # We need to create the x array starting from 1
y[step] = x[step]**2.0
# print the final 3 values for both x and y
print("the value of the 98th component of x is ", x[97])
print("the value of the 98th component of y is ", y[97])
print("the value of the 99th component of x is ", x[98])
print("the value of the 99th component of y is ", y[98])
print("the value of the 100th component of x is ", x[99])
print("the value of the 100th component of y is ", y[99])
# -
# What about accessing and modifying values in a 2D array? We can 'nest' multiple *for* loops to deal with each dimension. Below is an example where we populate values in a 2D array, of 100 rows and 100 columns, according to the following function:
#
# \begin{align*}
# z = x+y
# \end{align*}
#
# Where variable <code> X </code> represents a row index and <code> Y </code> represents the column index. <code> Z </code> is a 2D array which is then plotted using the matplotlib package, assuming you use a 2D Numpy array for <code> Z </code>. Read through the comments associated with each step, can you recognise the stages of constructing this set of operations?
# +
# Import the modules we will be using
import numpy as np
import matplotlib.pyplot as plt
# Initialise the Numpy arrays
x = np.zeros((100,), dtype=float)
y = np.zeros((100,), dtype=float)
z = np.zeros((100,100), dtype=float)
# Lets populate our x and y arrays with values from 1 to 100
for step in range(100):
x[step]=step+1 # why do we add 1?
y[step]=step+1 # why do we add 1?
# Now create a nested loop that cycles through each dimension to produce our z array
# Thus the first loop cycles through each row
for row in range(100):
# The second loop cycles through each column.
for column in range(100):
z[row,column]=x[row]+y[column]
# Plot the results. Dont worry about the Matplotlib template for now.
plt.contourf(x, y, z, 20, cmap='RdGy')
plt.colorbar();
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
# Before we move on to the second exercise, lets look at what is happening in our 'nested' loop. In English, the operation is designed to populate values in our 2D array, or matrix, as follows:
#
# 1) Specify a for loop for the row index. Lets call this variable 'row'. This cycles through values from 1 to 100. The index is our row number.
#
# 2) For each row number, lets cycle through each column index. Lets call this variable 'col'. This is equivalent to scanning across a matrix left-right.
#
# 3) Now we have a value for a row and column index. Use these indices to extract values from our X and Y arrays, as X[row] and Y[col].
#
# 4) Use these values to create an entry into our 2D array 'Z' as Z[row,col] = X[row]+Y[col].
#
# <div class="alert alert-block alert-success">
# <b> Exercise 2: Create a 2D array and populate with values according to an equation <a name="Exercise2"></a> </b>
#
# In this practice exercise, your goal is to use a *for* loop to populate a 2D Numpy array with 200 rows and 300 columns based on the following equation:
#
# \begin{align*}
# z = x+y*x
# \end{align*}
#
# Where variable <code> X </code> is an array that holds the row index and <code> Y </code> is an array that holds the column index. <code> Z </code> is a 2D array which is then plotted using the matplotlib package. If successful, your simulation should generate a plot given by the figure below.
#
# 
#
#
# </div>
#
# <div class="alert alert-block alert-warning">
# <b>HINT</b> You will need to use a nested 'for' loop to cycle through the rows and columns. Check out our previous example above. Remember that Python expects indentation after the ':' operator.</div>
# +
# Import the modules we will be using
import numpy as np
import matplotlib.pyplot as plt
# Initialise the arrays
x = np.zeros((100,), dtype=float)
y = np.zeros((100,), dtype=float)
z = np.zeros((100,100), dtype=float)
# ------- 'INSERT CODE HERE' ---------
# Lets populate our x and y arrays with values from 1 to 100
# ------- 'INSERT CODE HERE' ---------
# Now create a nested loop that cycles through each dimension to produce our z array
# Thus the first loop cycles through each row
for row in range(100):
# The second loop cycles through each column.
for column in range(100):
# Plot the results. Dont worry about the Matplotlib template for now.
plt.contourf(x, y, z, 20, cmap='RdGy')
plt.colorbar();
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
# ## Part 3: Creating your own function <a name="Part3"></a>
#
# In the same way we have started to use functions included within the Numpy package, we can write our own functions to perform specific tasks. This might be a numerical operation, or string manipulation task. This brings us closer to modular software development. In the example below we define a function with the keyword **'def'** and then call the function to receive its output. For example:
# +
# Lets define a function called 'test'. This function will simply write a message to the screen.
def test():
print("All work and no play makes Jack a dull boy")
return
# Call the function
test()
# -
# Let us understand a few important features used in defining the function:
#
# - The 'def' keyword defined a function by its name.
# - The parenthesis () can take a list of inputs that the function needs.
# - The function name and input list is terminated with a colon ':'.
# - The next line, thus internal workings of the function, is *idented* by 4 spaces.
# - A 'return' statement closes the function definition.
#
# <div class="alert alert-block alert-success">
# <b> Exercise 3: Re-create the repeated output from the typerwriter in the Shinning... <a name="Exercise3"></a> </b>
#
# Using the example function above, combine this function call with a <code> FOR </code> loop to print 30 instances of the sentence to the screen.
#
# </div>
#
# <div class="alert alert-block alert-warning">
# <b>HINT</b>: Remember that a <code> FOR </code> loop takes the format: 'for "variable" in range(number):' </div>
# +
# Lets define a function called 'test'. This function will simply write a message to the screen.
def test():
print("All work and no play makes Jack a dull boy")
return
# Call the function in a 'for' loop, 30 times
# ------- 'INSERT CODE HERE' ---------
# ------------------------------------
# -
# Before we move on to the final exercise, let's look at another example on populating a 2D array with the function:
#
# \begin{align*}
# z = x+y
# \end{align*}
#
# Where we are going to implement this equation as a function, which is passed arrays <code> X </code> and <code> Y </code> and returns <code> Z </code>. Read through the code below and see if this makes sense. Can you notice where we have now put the nested loops inside a new function?
# +
# Import the modules we will be using
import numpy as np
import matplotlib.pyplot as plt
# Initialise the arrays
x = np.zeros((100,), dtype=float)
y = np.zeros((100,), dtype=float)
# Lets populate our x and y arrays with values from 1 to 100
for step in range(100):
x[step]=step+1 # why do we add 1?
y[step]=step+1 # why do we add 1?
# Implement the equation z=x+y as a function
def create_z_array(x,y):
# Initialise the z array
z = np.zeros((100,100), dtype=float)
# Now create a nested loop that cycles through each dimension to produce our z array
# Thus the first loop cycles through each row
for row in range(100):
# The second loop cycles through each column.
for column in range(100):
z[row,column]=x[row]+y[column]
# return the z array
return z
# Call the function that generates our z array
z=create_z_array(x,y)
# Plot the results. Dont worry about the Matplotlib template for now.
plt.contourf(x, y, z, 20, cmap='RdGy')
plt.colorbar();
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
# <div class="alert alert-block alert-success">
# <b> Exercise 4: Build a function that creates a 2D surface plot of a scalar function <a name="Exercise4"></a> </b>
#
# In this practice assignment, you are tasked with inserting all of the code [except the plotting function] to replicate the function used in Practice '2':
#
# \begin{align*}
# z = x+y*x
# \end{align*}
#
# Where variable <code> X </code> represents the row number and <code> Y </code> represented the column number. In this instance we have 200 rows and 200 columns and the procedure to populate our <code> Z </code> array should be included as a function.
#
# </div>
#
# <div class="alert alert-block alert-warning">
# <b>HINT</b>: The majority of this code has already been written earlier in this practical, but you need to reformat this to be called as a function. Also note the change in number of rows and columns. </div>
# +
# Import the modules we will be using
import numpy as np
import matplotlib.pyplot as plt
# Initialise the arrays
x = np.zeros((300,), dtype=float)
y = np.zeros((300,), dtype=float)
z = np.zeros((300,300), dtype=float)
# ------- 'INSERT CODE HERE' ---------
# Lets populate our x and y arrays with values from 1 to 100
# ------------------------------------
# Now create a nested loop that cycles through each dimension to produce our z array
# Thus the first loop cycles through each row
def create_z(x,y,z):
# ------- 'INSERT CODE HERE' ---------
# ------------------------------------
return z
z=create_z(x,y,z)
# Plot the results. Dont worry about the Matplotlib template for now.
plt.contourf(x, y, z, 20, cmap='RdGy')
plt.colorbar();
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
# # Part 4: Numpy operations on entire arrays <a name="Part4"></a>
#
# Once we start writing *large* modules we need to start thinking about how efficient our procedures are. In many areas of research, time of computation is a fundamental hurdle to increasing complexity. We will not cover optimisation in this course, but it is useful to start considering where we might use more efficient options in numerical computation. The standard Numpy library offers us multiple options which also helps to reduce the size of our code and, potentially, make it more readable!
#
# ### 4.1) Operations on a 1D Array
#
# A [nice example](https://www.pluralsight.com/guides/overview-basic-numpy-operations) is provided by using arithmetic operators with Numpy. If we have 2 arrays, array1 and array2, created through:
#
# ```python
# array1 = np.array([10,20,30,40,50], dtype=int)
# array2 = np.array([0,1,2,3,4], dtype=int)
# array3 = array1+array2
# ```
# what might our answer look like? Lets run this code below and print to the screen
# +
import numpy as np
array1 = np.array([10,20,30,40,50], dtype=int)
array2 = np.array([0,1,2,3,4], dtype=int)
array3 = array1+array2
print("array3 = ", array3)
# -
# Rather than cycling through each element using a loop, this was much easier to write. What exactly is going on? The [visualisation](https://www.pluralsight.com/guides/overview-basic-numpy-operations) below should help:
#
# <img src="images/adding_arrays.png" alt="Numpy array indexing" style="width: 500px;"/>
#
# ### 4.2) Operations on a 2D Array
#
# Unsurprisingly, we can also perform common arithmetic operations on 2D arrays. For example, let us add two 2D arrays [matrices] tother as per the following example:
#
# +
# Initiliase two empty matrices, A and B
A = np.zeros((3,3), dtype=float)
B = np.zeros((3,3), dtype=float)
# Set all cells in A to 2.4 and all cells in B to 1.6.
A[:,:]=2.4
B[:,:]=1.6
# Lets print the result of adding both together
print(A+B)
# -
# All the arithmetic operations work in a similar way. We can also multiply or divide the arrays. In the example below, see how we multiply matrix <code> A </code> by matrix <code> B </code>.
print(A*B)
# We can demonstrate how each cell is multiplied together with another example:
# Create two new matrices by manually writing out a Numpy array
C = np.array([[3,2],[0,1]])
D = np.array([[3,1],[2,1]])
# Multiply them both together. Can you confirm each cell has been acted on independently?
print(C*D)
# <div class="alert alert-block alert-success">
# <b> Exercise 5: Re-create a function that creates a 2D surface plot of a scalar function <a name="Exercise5"></a> </b>
#
# In this practice assignment, you are tasked with inserting all of the code [except the plotting function] to replicate the function used in Exercise '4':
#
# \begin{align*}
# z = x+y*x
# \end{align*}
#
# Where variable <code> X </code> represents the row number and <code> Y </code> represented the column number. In this instance we have 200 rows and 200 columns and the procedure to populate our <code> Z </code> array should be included as a function. To start you off, I have initialised both <code> X </code> and <code> Y </code> as 2D arrays for you to use
#
# </div>
#
# <div class="alert alert-block alert-warning">
# <b>HINT</b>: The majority of this code has already been written earlier in this practical, but you need to reformat this to be called as a function. Also note the change in number of rows and columns. </div>
# +
# Import the modules we will be using
import numpy as np
import matplotlib.pyplot as plt
# Initialise the arrays
x = np.zeros((300,300), dtype=float)
y = np.zeros((300,300), dtype=float)
z = np.zeros((300,300), dtype=float)
# Lets populate our x and y arrays with values from 1 to 100
for step in range(300):
x[step,:]=step+1 # why do we add 1?
y[:,step]=step+1 # why do we add 1?
# ------- 'INSERT CODE HERE' ---------
# Now create a nested loop that cycles through each dimension to produce our z array
# Thus the first loop cycles through each row
def create_z(x,y,z):
return z
# ------------------------------------
z=create_z(x,y,z)
# Plot the results. Dont worry about the Matplotlib template for now.
plt.contourf(x, y, z, 20, cmap='RdGy')
plt.colorbar();
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
| Practical_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FMU Data Acquisition Code
from serial import *
import time
import sys
import numpy as np
import matplotlib.pyplot as plt
from threading import Thread
# ## Connect
# +
COM = 'COM4'# /dev/ttyACM0 (Linux)
BAUD = 9600
last_received = ''
stop_threading = False
def receiving(ser):
global last_received, stop_threading
buffer_string = ''
while True:
buffer_string = buffer_string + str(ser.read(ser.inWaiting()),'utf-8')
if '\n' in buffer_string:
lines = buffer_string.split('\n') # Guaranteed to have at least 2 entries
last_received = lines[-2]
#If the Arduino sends lots of empty lines, you'll lose the
#last filled line, so you could make the above statement conditional
#like so: if lines[-2]: last_received = lines[-2]
buffer_string = lines[-1]
if stop_threading==True:
break
COM = 'COM4'
ser = Serial(
port=COM,
baudrate=9600,
bytesize=EIGHTBITS,
parity=PARITY_NONE,
stopbits=STOPBITS_ONE,
timeout=0.1,
xonxoff=0,
rtscts=0,
interCharTimeout=None
)
thread=Thread(target=receiving, args=(ser,))
thread.start()
print('Waiting for device')
print(ser.name)
# time.sleep(7)
for i in range(7):
sys.stdout.write("\r"+str(i+1)+" sec")
time.sleep(1)
print("\nConnection Established!")
def is_number(s):
try:
float(s)
return True
except ValueError:
return False
# -
# ## Acquisition
# +
# timeFinal = float(input("Duration of acquisition:")) # seconds
timeFinal = 10 # seconds
sensorValue=[]
sensorValue2=[]
timeValue=[]
sensorValue.clear()
sensorValue2.clear()
timeValue.clear()
timeInitial=time.time()
timePassed=0
i=0
timeValue_last=0
print("Starting measurement...")
while (timePassed<timeFinal):
timeCurrent = time.time()
timePassed=timeCurrent-timeInitial
serialValue = last_received.split() #Capture serial output as a decoded string
if float(serialValue[0])!=timeValue_last:
if (len(serialValue) == 3):
if is_number(serialValue[0] and serialValue[1]):
timeValue.append(float(serialValue[0]))
sensorValue.append(float(serialValue[1]))
sensorValue2.append(float(serialValue[2]))
timeValue_last=float(serialValue[0])
sys.stdout.write("\r"+"Measuring: " + str("{0:.0f}".format(np.floor(100*timePassed/timeFinal)))+"% "+str("{0:.1f}".format(timePassed))+" seconds passed")
print("\nMeasurement finished.")
sensorValue = np.array(sensorValue)
sensorValue2 = np.array(sensorValue2)
# sensorValue3 = np.array(sensorValue3)
timeValue = np.array(timeValue)
timeValue = timeValue-np.ones((len(timeValue),))*timeValue[0]
timeValue=timeValue*1E-3 #put in seconds (original in mili)
fig=plt.gcf()
plt.clf()
fig.tight_layout()
plt.ylabel("Force (g)")
plt.xlabel("Time (s)")
plt.plot(timeValue, sensorValue, label="Bottom Sensor")
plt.plot(timeValue, sensorValue2, label="Top Sensors")
plt.legend()
# -
# # Save
#
# !!! **IMPORTANT**: Change *numberOfSample* otherwise files get overwritten
# +
wafer = "test-sultan"
numberOfSample = ""
fname = wafer+"-"+str(numberOfSample)
# fname="test"
try:
np.savetxt("sample-"+fname+".csv",np.c_[timeValue,sensorValue,sensorValue2],delimiter=",") #np.c_[] to put lines in columns in csv
fig.savefig("sample-"+fname+".png",dpi=120)
print("Files saved!")
except ValueError:
print(ValueError)
# -
# # Test Variables
# +
# print(timeValue[1000])
print(last_received)
plt.clf()
fig=plt.gcf()
fig.tight_layout()
plt.ylabel("Force (g)")
plt.xlabel("Time (s)")
# plt.plot(timeValue, sensorValue, label="Bottom Sensor")
plt.plot(timeValue, sensorValue2,color='orange', label="Top Sensors")
plt.legend()
# -
# # Close Connection COM4
stop_threading=True
thread.join()
sys.stdout.write("\rThread killed!")
ser.close()
| read_serial_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: feat_select
# language: python
# name: feat_select
# ---
# # Feature Selection
# ---
# Feature selection is the process of selecting the features that hold the most predictive power to the target. By removing unnecessary features, we reduce model complexity and minimize the computational resources required for training and inference. Feature selection is a crucial step that can have a great impact on model efficiency in production settings.
#
# The methods of feature selection we will perform are:
# * Filter Methods
# * Correlation
# * Univariate Feature Selection
# * Wrapper Methods
# * Forward Selection
# * Backward Selection
# * Recursive Feature Elimination
# * Embedded Methods
# * Feature Importance (Tree-based)
# * L1 Regularization
#
# In this notebook, we will demonstrate the feature selection methods above on the [Census Income](https://archive.ics.uci.edu/ml/datasets/Census-Income+%28KDD%29) dataset from the UCI repository. The dataset contains both numerical and categorical features with the goal to predict whether a person's salary is greater than or equal to $50k.
# # Imports
# +
# for data processing and manipulation
import pandas as pd
import numpy as np
# scikit-learn modules for feature selection and model evaluation
from sklearn.ensemble import RandomForestClassifier
from sklearn.feature_selection import RFE, SelectKBest, SelectFromModel, SequentialFeatureSelector, chi2, f_classif
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, roc_auc_score, precision_score, recall_score, f1_score
from sklearn.feature_selection import SelectFromModel
from sklearn.preprocessing import StandardScaler, MinMaxScaler
# libraries for visualization
import seaborn as sns
import matplotlib
import matplotlib.pyplot as plt
| feature_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import os
import matplotlib.pyplot as plt
proj_path = os.path.join((os.getcwd()),os.path.pardir)
raw_path = os.path.join(proj_path, "data","raw")
train_file_path=os.path.join(raw_path, "train.csv")
test_file_path=os.path.join(raw_path, "test.csv")
train_df = pd.read_csv(train_file_path, index_col="PassengerId")
test_df = pd.read_csv(test_file_path, index_col="PassengerId")
titanic_df = pd.concat((train_df,test_df))
# %matplotlib inline
fig, axs = plt.subplots(1,2)
titanic_df.Age.plot(figsize=(15,5),kind="hist",title="Age of Passengers",bins=20,
color="lightgreen", edgecolor='white', linewidth=2, ax = axs[0]);
titanic_df.Age.plot(figsize=(15,5),kind="kde",title="Age of Passengers",ax=axs[1]);
fig, axs = plt.subplots(1,2)
titanic_df.Fare.plot(figsize=(15,5),kind="hist",title="Ticket Fare Distribution",bins=20,
color="lightgreen", edgecolor='white', linewidth=2, ax = axs[0]);
titanic_df.Fare.plot(figsize=(15,5),kind="kde",title="Ticket Fare Distribution",ax=axs[1]);
print('skewness for age: {0:0.2f}'.format(titanic_df.Age.skew()))
print('skewness for age: {0:0.2f}'.format(titanic_df.Fare.skew()))
fig, axs = plt.subplots(1,2)
titanic_df.plot.scatter(figsize=(15,4),x="Age", y="Fare", title="Age vs Fare Distribution",
color="darkgreen", alpha=0.2, ax=axs[0]);
titanic_df.plot.scatter(figsize=(15,4),x="Pclass", y="Fare", title="Age vs Fare Distribution",
color="darkgreen", alpha=0.1, ax=axs[1]);
# ### GROUPING AND AGGREGATION
from pprint import pprint as pp
titanic_df.groupby("Age").mean()
titanic_df.groupby("Sex").Age.median()
titanic_df.groupby(["Pclass"]).Age.median()
titanic_df.groupby(["Pclass"])["Fare","Age"].median()
titanic_df.groupby(["Pclass"]).agg({"Fare":"mean","Age":"median"})
aggregations = {
"Fare":{
"mean_fare":"mean",
"median_fare":"median",
"max_fare":"max",
"min_fare":np.min
},
"Age":{
"median_age":"median",
"max_age":"max",
"min_age":"min",
"range_age": lambda x: max(x)-min(x)
}
}
titanic_df.groupby(["Pclass"]).agg(aggregations)
titanic_df.groupby(["Pclass","Embarked"]).Fare.median()
titanic_df.groupby(["Pclass"]).agg(aggregations)
pd.crosstab(titanic_df.Sex,titanic_df.Pclass)
titanic_df.groupby(["Sex","Pclass"])["Sex"].agg("count").to_frame().iloc[0:6
,0:3].unstack()
fig, axs = plt.subplots(1,2)
pd.crosstab(titanic_df.Sex,titanic_df.Pclass).plot(kind="hist", ax=axs[0], figsize=(15,4));
pd.crosstab(titanic_df.Sex,titanic_df.Pclass).plot(kind="bar", ax=axs[1], figsize=(15,4))
titanic_df.pivot_table(index="Sex", columns = "Pclass", values="Age", aggfunc="mean")
titanic_df.groupby(["Pclass","Sex"])["Age"].mean()
titanic_df.groupby(["Pclass","Sex"])["Age"].mean().unstack()
| notebooks/distribution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env
# language: python
# name: env
# ---
# +
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
#from keras import backend as K
import matplotlib.pyplot as plt
import pandas as pd
import datetime
from matplotlib.ticker import MaxNLocator
from matplotlib import cm
from mpl_toolkits.mplot3d import Axes3D
import os
import random
# -
path = '/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/TwoClassModelsV2'
Main_path = '/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow'
# +
train_X = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainX13.csv', header=None)
train_Y = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainY13.csv', header=None)
trainY = train_Y.to_numpy()
trainX = train_X.to_numpy()
trainX = np.expand_dims(trainX,axis=2)
minimum = np.amin(trainX)
maximum = np.amax(trainX)
# -
accuracy_df = pd.read_csv(path+"/Accuracy.csv")
accuracy_df
classification_model = tf.keras.models.load_model('TrainedModel/trainedModel.h5')
# ## Define bivariate normal distribution pdf
def biVariate_Prob(coVariance,mean,x):
LHS = np.power(np.linalg.det(2*np.pi*coVariance),-0.5)
RHS = np.exp(-0.5*np.dot(((x-mean).T),(np.dot(np.linalg.inv(coVariance),(x-mean)))))
return LHS*RHS
# + tags=[]
# #%matplotlib widget
# #%matplotlib inline
for index, row in accuracy_df.iterrows():
#Checking if the specific class Tuple has enough points for both A and B
if((row['Correct?'] > 120) and (row['A_Count'] > 50) and (row['B_Count'] > 50)):
#Extracting classA classB tuple from Accuracy.csv
my_list = row['Class Tuples'].split(",")
classA = int(my_list[0])
classB = int(my_list[1])
#Uncomment below if you need to force class A and class B to be certain values. Dont forget to add break at the end of for loop
classA = 3
classB = 11
#Loading the corresponding decoder of class A, classB combination
decoder = tf.keras.models.load_model(path+"/"+str(classA)+"/"+str(classB)+"/modeldecoder.h5")
points = pd.read_csv(path+"/"+str(classA)+"/"+str(classB)+"/points.csv")
points = points.drop(['Unnamed: 0'], axis=1)
X_A = points["X_A"].to_numpy()
Y_A = points["Y_A"].to_numpy()
X_B = points["X_B"].to_numpy()
Y_B = points["Y_B"].to_numpy()
X_A = X_A[~np.isnan(X_A)]
Y_A = Y_A[~np.isnan(Y_A)]
X_B = X_B[~np.isnan(X_B)]
Y_B = Y_B[~np.isnan(Y_B)]
#Might be implemented incorrectly, check while debugging! Update: Honestly, this is a disgusting way to do it and I'm ashamed but it works so I'm not touching it!
APoints = np.vstack((X_A, Y_A))
covA = np.cov(APoints)
BPoints = np.vstack((X_B, Y_B))
covB = np.cov(BPoints)
#Parameters to set ---- A
mu_x_A = np.mean(X_A)
variance_x_A = covA[0][0]
mu_y_A = np.mean(Y_A)
variance_y_A = covA[1][1]
mu_A = np.array([mu_x_A,mu_y_A])
#Parameters to set ---- B
mu_x_B = np.mean(X_B)
variance_x_B = covB[0][0]
mu_y_B = np.mean(Y_B)
variance_y_B = covB[1][1]
mu_B = np.array([mu_x_B,mu_y_B])
#Printing covariances and means for debugging purposes
#print("Class A")
#print("Covariance: "+ str(covA))
#print("Mean: "+ str(mu_A))
#print("-----------")
#print("Class B")
#print("Covariance: "+ str(covB))
#print("Mean: "+ str(mu_B))
maxY = np.amax([np.amax(Y_A),np.amax(Y_B)])
maxX = np.amax([np.amax(X_A),np.amax(X_B)])
minY = np.amin([np.amin(Y_A),np.amin(Y_B)])
minX = np.amin([np.amin(X_A),np.amin(X_B)])
#decreaseRate= how much should the box be smaller than the normal enclosing box that normally would be. This is done because the box is, well... literally a box. The edges are not even between two clusters-
#and so the edges of the box correspond to points outside of the two clusters, and therefore E[clusterA]-E[clusterB] still gives super small numbers but end up being wrong.
#POSSIBLE OPTIMIZATION: make the linspace a circle instead of a box, should technically help a lot!
decreaseRate = 0.9
x = np.linspace(maxX-decreaseRate, minX+decreaseRate, num=50)
y = np.linspace(maxY-decreaseRate, minY+decreaseRate, num=50)
x_array = []
y_array = []
z_array = []
bestPoints_for_normal = []
#print(bestPoints_for_normal.shape)
#print(np.expand_dims(bestPoints_for_normal,axis=1).shape)
summa = 0
for x_i in x:
best_Point_x = []
best_Point_y = []
best_Point_difference = []
for y_i in y:
#calculating differences with respect to biVariate expectation, and then appending it to z_array
difference = abs(biVariate_Prob(covA,mu_A,[x_i,y_i])-biVariate_Prob(covB,mu_B,[x_i,y_i]))
#Append to global array
x_array.append(x_i)
y_array.append(y_i)
z_array.append(difference)
#Append to array that exists only for specific y
best_Point_x.append(x_i)
best_Point_y.append(y_i)
best_Point_difference.append(difference)
YPoints = np.append(np.array([best_Point_x]).T,np.array([best_Point_y]).T,axis=1)
smallestPoint = YPoints[np.argmin(best_Point_difference)]
#print(smallestPoint)
#break
#print(YPoints[np.argmin(best_Point_difference)].shape)
#break
#best_PointFor_Y = np.append(best_PointFor_Y,np.array([best_Point_difference]).T,axis=1)
bestPoints_for_normal.append(smallestPoint)
#bestPoints_for_normal = np.append(bestPoints_for_normal,smallestPoint,axis=1)
bestPoints_for_normal = np.array(bestPoints_for_normal)
#Appending x_array, y_array, and z_array together to create a matrix
Points = np.append(np.array([x_array]).T,np.array([y_array]).T,axis=1)
Points = np.append(Points,np.array([z_array]).T,axis=1)
#Taking first two columns of Points (x,y), and reconstructing and classifying and taking the confidences of classA and class B and putting it back to x,y array as the 3rd column. For some reason all this can be done in 4 lines
#Magic of Python and Numpy
pointsForClassification = Points[:,0:2]
reconstructions = decoder.predict(Points[:,0:2])
confidences = classification_model.predict(reconstructions)
differencesWRTClassification = abs(confidences[:, [classA]]-confidences[:, [classB]])
#THESE ARE THE TWO POINT ARRAYS WHERE THE 3RD COLUMN IS EITHER DIFFERENCE WRT NORMAL OR DIFFERENCE WRT CLASSIFICATION MODEL
PointsNormal = Points
PointsClassification = np.append(Points[:,0:2],differencesWRTClassification,axis=1)
#SORTING THE ARRAYS WRT 3RD COLUMN(DIFFERENCES)
sortedPointsClassification = PointsClassification[PointsClassification[:,2].argsort()]
sortedPointsNormal = PointsNormal[PointsNormal[:,2].argsort()]
amountToCut = 50
#Sampling multivariate random from given covariance and mean for class A
randomSamples = []
for i in range(amountToCut):
randomSample = np.random.multivariate_normal(mu_A,covA)
randomSamples.append(randomSample)
randomSamples = np.array(randomSamples).T
reconstructions_From_Middle = decoder.predict(bestPoints_for_normal)
reconstructions_From_Random = decoder.predict(randomSamples.T)
NoiseA = reconstructions_From_Middle-reconstructions_From_Random
#Sampling multivariate random from given covariance and mean for class B
randomSamples = []
for i in range(amountToCut):
randomSample = np.random.multivariate_normal(mu_B,covB)
randomSamples.append(randomSample)
randomSamples = np.array(randomSamples).T
reconstructions_From_Middle = decoder.predict(bestPoints_for_normal)
reconstructions_From_Random = decoder.predict(randomSamples.T)
NoiseB = reconstructions_From_Middle-reconstructions_From_Random
if(True):
isExist = os.path.exists(Main_path+"/Noise/"+str(classA))
if not isExist:
# Create a new directory because it does not exist
os.makedirs(Main_path+"/Noise/"+str(classA))
isExist = os.path.exists(Main_path+"/Noise/"+str(classB))
if not isExist:
# Create a new directory because it does not exist
os.makedirs(Main_path+"/Noise/"+str(classB))
#print(np.squeeze(NoiseA,axis=2).shape)
np.savetxt(Main_path+"/Noise/"+str(classA)+"/"+str(classB)+".csv", np.squeeze(NoiseA * (maximum-minimum)+minimum,axis=2), delimiter=",")
np.savetxt(Main_path+"/Noise/"+str(classB)+"/"+str(classA)+".csv", np.squeeze(NoiseB * (maximum-minimum)+minimum,axis=2), delimiter=",")
#MOST BOTTOM IF STATEMENTS ARE TESTS
#------------------------------------------------------
if(False):
train_X = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainX13.csv', header=None)
train_Y = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainY13.csv', header=None)
trainY = train_Y.to_numpy()
trainX = train_X.to_numpy()
trainX = np.expand_dims(trainX,axis=2)
minimum = np.amin(trainX)
maximum = np.amax(trainX)
amt = 20
x = np.linspace(mu_x_A, mu_x_B, num=amt)
y = np.linspace(mu_y_A, mu_y_B, num=amt)
together = np.append([x],[y],axis=0).T
reconstructed = decoder.predict(together)
reconstructed = reconstructed * (maximum-minimum)+minimum
for i in range(amt):
fig = plt.figure(figsize=(35,5))
plt.plot(reconstructed[i].T[0])
plt.legend()
plt.yticks(np.arange(0, 9, 1))
plt.xticks(np.arange(0, 3000, 500))
plt.grid()
plt.xlabel("5 ms")
plt.ylabel("PnP timing")
fig.savefig("Graphs/"+str(i)+".png",dpi=200)
#print(reconstructed[0].T[0])
#-------------------------------------------------------
if(True):
train_X = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainX13.csv', header=None)
train_Y = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainY13.csv', header=None)
amt = 20
x = np.linspace(mu_x_A, mu_x_B, num=amt)
y = np.linspace(mu_y_A, mu_y_B, num=amt)
together = np.append([x],[y],axis=0).T
#print(together[7].tolist())
#print([[mu_x_A,mu_y_A]])
reconstruction_test = decoder.predict([together[11].tolist()])
specificReconstruction = reconstructions_From_Middle[25]
trainY = train_Y.to_numpy()
trainX = train_X.to_numpy()
trainX = np.expand_dims(trainX,axis=2)
minimum = np.amin(trainX)
maximum = np.amax(trainX)
print(minimum)
print(maximum)
#print(randomSamples.T[25])
fig = plt.figure(figsize=(35,5))
#print(reconstruction_test[0].T[0])
#print(specificReconstruction.T)
#plt.plot(NoiseA[25].T[0]*(maximum-minimum)+minimum)
#plt.plot(specificReconstruction.T[0]*(maximum-minimum)+minimum)
plt.plot(reconstruction_test[0].T[0]*(maximum-minimum)+minimum)
#reconst = np.expand_dims(reconstruction_test[0].T[0],axis=1)
print(classification_model.predict(reconstruction_test))
plt.legend()
plt.yticks(np.arange(0, 9, 1))
plt.xticks(np.arange(0, 3000, 500))
plt.grid()
#plt.axhline(linewidth=1, color='r')
plt.xlabel("5 ms")
plt.ylabel("PnP timing")
fig.savefig("Graphs/"+str(i)+".png",dpi=200)
#-------------------------------------------------------
break
#print(NoiseA)
#print("---------")
#print(NoiseB)
#THIS IS ALL AFTER NOISE IS CALCULATED FOR CLASS A AND B
if(False):
train_X = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainX13.csv', header=None)
train_Y = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainY13.csv', header=None)
trainY = train_Y.to_numpy()
trainX = train_X.to_numpy()
trainX = np.expand_dims(trainX,axis=2)
minimum = np.amin(trainX)
maximum = np.amax(trainX)
trainX_normalized = (trainX-minimum)/(maximum-minimum)
trainX_tensor = tf.convert_to_tensor(trainX_normalized)
output = tf.nn.max_pool1d(trainX_tensor, 2, 2, padding='VALID')
trainA = pooled_dataset[classA::14]
trainB = pooled_dataset[classB::14]
trainA_Out = trainY[classA::14]
trainB_Out = trainY[classB::14]
finalX = np.append(trainA,trainB,axis=0)
finalY = np.append(trainA_Out,trainB_Out)
if(False):
i = 0
for rec in reconstructions_From_Middle:
fig = plt.figure(figsize=(35,5))
#plt.plot(results)
#sampleToPredict = 15
#plt.plot(reconstructed_x[sampleToPredict],label='Reconstruction')
#plt.plot(trainXCUT[sampleToPredict],label='Sample')
#plt.plot(data3[0],label=3)
#plt.plot(data4[0],label=4)
plt.plot(rec.T[0])
plt.legend()
plt.yticks(np.arange(0, 1, 0.1))
plt.xticks(np.arange(0, 3000, 500))
plt.grid()
#plt.axhline(linewidth=1, color='r')
plt.xlabel("5 ms")
plt.ylabel("PnP timing")
#figure(figsize=(8, 6), dpi=80)
fig.savefig("Graphs/"+str(i)+".png",dpi=200)
i = i +1
#PLOTTING
if(False):
fig = plt.figure(figsize=(13,10))
plt.scatter(X_A,Y_A,color='r',label="Class "+str(classA))
plt.scatter(X_B,Y_B,color='b',label="Class "+str(classB))
#plt.scatter(sortedPointsNormal[0:amountToCut,0:1].T[0],sortedPointsNormal[0:amountToCut,1:2].T[0],color='y',label="Middle According to Normal")
plt.scatter(sortedPointsClassification[0:amountToCut,0:1].T[0],sortedPointsClassification[0:amountToCut,1:2].T[0],color='g',label="Middle According to Classification")
plt.scatter(bestPoints_for_normal[:,0],bestPoints_for_normal[:,1],color='y',label="Middle According to Berk")
plt.scatter(randomSamples[0],randomSamples[1],color='k')
plt.axhline(y=0, color='k', linestyle='-')
plt.axvline(x=0, color='k', linestyle='-')
plt.legend()
print(row)
# -
# ## Test Noise Array
train_X = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainX13.csv', header=None)
train_Y = pd.read_csv('/home/ege/Repo/SideChannel-AdversarialAI/Tensorflow/DataSet/trainY13.csv', header=None)
trainY = train_Y.to_numpy()
trainX = train_X.to_numpy()
trainX = np.expand_dims(trainX,axis=2)
trainX.shape
trainX_tensor = tf.convert_to_tensor(trainX)
finalX = tf.nn.max_pool1d(trainX, 2, 2, padding='VALID')
finalX = finalX.numpy()
finalX.shape
# +
numOfClasses = 14
NoiseToAdd = np.zeros(finalX.shape)
NoiseToAdd = np.squeeze(NoiseToAdd)
eachClassSize = int(len(trainY)/numOfClasses)
for i in range(numOfClasses):
print("Class: "+str(i))
isExist = os.path.exists(Main_path+"/Noise/"+str(i))
if isExist:
target_files = os.listdir(Main_path+"/Noise/"+str(i))
noiseArrayForSingleClass = []
for file in target_files:
single_target = pd.read_csv(Main_path+"/Noise/"+str(i)+"/"+file, header=None).to_numpy()
noiseArrayForSingleClass.append(single_target)
noiseForSingleClass = np.reshape(np.array(noiseArrayForSingleClass), (len(target_files)*50,3000))
j = i
while(j<len(trainY)):
#print(random.randint(0,len(noiseForSingleClass)-1))
NoiseToAdd[j] = noiseForSingleClass[random.randint(0,len(noiseForSingleClass)-1)]
j = j +numOfClasses
NoiseToAdd = np.expand_dims(NoiseToAdd,axis=2)
print(NoiseToAdd.shape)
# -
NoiseToAdd[NoiseToAdd < 0] = 0
finalXwNoise = finalX + 10*NoiseToAdd
# +
minimum = np.amin(trainX)
maximum = np.amax(trainX)
trainX_normalized = (finalXwNoise-minimum)/(maximum-minimum)
# -
trainX_normalized.shape
classification_model.evaluate(trainX_normalized,trainY)
# ### Train New Model w Noise
numberOfWebsites = 14
def my_model_sddec():
input_1 = keras.Input(shape = (3000,1))
conv1d_1 = layers.Conv1D(256,16,strides=3,padding='valid',activation='relu',use_bias=True,kernel_initializer='VarianceScaling',bias_initializer = 'Zeros')(input_1)#possibly update kernel_initializer
max_pooling1d_1 = layers.MaxPooling1D(pool_size = 4,strides = 4, padding = 'same')(conv1d_1)
conv1d_2 = layers.Conv1D(128,8,strides=3,padding='valid',activation='relu',use_bias=True,kernel_initializer='VarianceScaling',bias_initializer = 'Zeros')(max_pooling1d_1)#possibly update kernel_initializer
max_pooling1d_2 = layers.MaxPooling1D(pool_size = 4,strides = 4, padding = 'same')(conv1d_2)
conv1d_3 = layers.Conv1D(32,8,strides=3,padding='same',activation='relu',use_bias=True,kernel_initializer='VarianceScaling',bias_initializer = 'Zeros')(max_pooling1d_2)#possibly update kernel_initializer
max_pooling1d_2 = layers.MaxPooling1D(pool_size = 4,strides = 4, padding = 'same')(conv1d_3)
# conv1d_1 = layers.Conv1D(256,16,strides=3,padding='valid',activation='relu',use_bias=True,kernel_initializer='VarianceScaling',bias_initializer = 'Zeros')(input_1)#possibly update kernel_initializer
# max_pooling1d_1 = layers.MaxPooling1D(pool_size = 4,strides = 4, padding = 'same')(conv1d_1)
# conv1d_2 = layers.Conv1D(32,8,strides=3,padding='same',activation='relu',use_bias=True,kernel_initializer='VarianceScaling',bias_initializer = 'Zeros')(max_pooling1d_1)#possibly update kernel_initializer
# max_pooling1d_2 = layers.MaxPooling1D(pool_size = 4,strides = 4, padding = 'same')(conv1d_2)
#lstm_1 = layers.LSTM(32,activation='tanh',recurrent_activation='hard_sigmoid',use_bias=True,kernel_initializer='VarianceScaling',recurrent_initializer = 'orthogonal',bias_initializer='Zeros', return_sequences = True)(max_pooling1d_2) #Variance Scaling
flatten_1 = layers.Flatten()(max_pooling1d_2)
#x = layers.Dense(200,activation = 'softmax')(flatten_1)
#dropout_1 = layers.Dropout(0.3)(flatten_1)
#dense_1 = layers.Dense(300,activation = 'relu')(dropout_1)
#dropout_2 = layers.Dropout(0.5)(x)
dense_2= layers.Dense(numberOfWebsites, kernel_regularizer = 'l2',activation = 'softmax', kernel_initializer = 'VarianceScaling', bias_initializer = 'zeros')(flatten_1)
model = keras.Model(inputs = input_1, outputs = dense_2)
return model
# + tags=[]
model = my_model_sddec()
model.summary()
# + tags=[]
model.compile(
loss=keras.losses.SparseCategoricalCrossentropy(),
optimizer=keras.optimizers.Adam(),
metrics=["accuracy"]
)
history = model.fit(trainX_normalized, trainY,validation_split = 0.3, batch_size=32, epochs=25, verbose=1)
# +
fig = plt.figure(figsize=(10,5))
#plotting
plt.locator_params(axis="x", nbins=20)
plt.locator_params(axis="y", nbins=10)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['training','validation'], loc='upper left')
fig.savefig('plot.png',dpi=200)
# -
np.var(trainX_normalized,axis=0)
| Tensorflow/Fit Normal to Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <p style="text-align: center;"> Part Six: Fuzzy Matching </p>
#
# 
from IPython.core.display import HTML
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
The raw code for this IPython notebook is by default hidden for easier reading.
To toggle on/off the raw code, click <a href="javascript:code_toggle()">here</a>.''')
# # <p style="text-align: center;"> Table of Contents </p>
# - ## 1. [Introduction](#Introduction)
# - ### 1.1 [Abstract](#abstract)
# - ## 2 [Fuzzy Matching](#fz)
# - ### 2.1 [Ratio](#r)
# - ### 2.2 [Partial Ratio](#pr)
# - ### 2.3 [Token Sort Ration](#tr)
# - ### 2.4 [Token Set Ratio](#ts)
# - ### 2.5 [Fuzzy Process Extract](#fz1)
# - ## 3. [Conclusion](#Conclusion)
# - ## 4. [Contribution](#Contribution)
# - ## 5. [Citation](#Citation)
# - ## 6. [License](#License)
# # <p style="text-align: center;"> 1.0 Introduction </p> <a id='Introduction'> </a>
#
# ## 1.1 Abstract <a id="abstract"> </a>
#
# As a data scientist, you are forced to retrieve information from various sources by either leveraging publicly available API’s, asking for data, or by simply scraping your own data from a web page. All this information is useful if we are able to combine it and not have any duplicates in the data. But how do we make sure that there are no duplicates?
#
# I know … “duh! you can just use a function that retrieves all the unique information thus removing duplicates”. Well, that’s one way, but our function probably can’t tell that a name like “<NAME>” is the same as “<NAME>” right? (Assuming we were retrieving names of the most famous people in the world). We can clearly tell that these names are different but they are probably referring to the same person. So, how do we match these names?
#
# This is where Fuzzy String Matching comes in. This post will explain what Fuzzy String Matching is together with its use cases and give examples using Python’s Library Fuzzywuzzy.
#
# 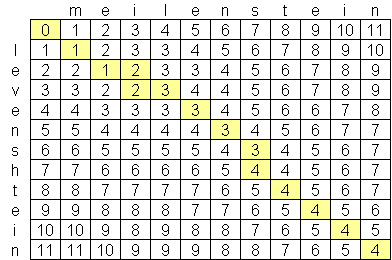
# ## 1.2 Importing Libaries <a id="importing_libraries"> </a>
import numpy as np
import pandas as pd
# ## <p style="text-align: center;"> 2.0 Fuzzy Matching </p> <a id='fz'> </a>
#
# We know we need to match records to identify duplicates and link records for entity resolution. But how exactly do we go about identifying matching records? What properties should we focus on?
#
# #### Deterministic Data Matching
# Let’s start with ‘unique identifiers’. These are properties in the records you want to match that are unlikely to change over time, Customer Name for instance. You can assign weights to each property to improve your matching process. Think about it; if you are migrating customer data from one system to another and need to check for duplicates pre- and post-migration, you could, for instance, choose Name as the one unique identifier and phone number as the second. Now it’s just a matter of running a search for matching Customer IDs and phone numbers and you have all potential matches identified. That method is known as ‘deterministic data matching’.
#
# #### Problems with Deterministic Data Matching?
#
# Although effective in theory, the method is rarely used because of its inflexibility: The approach assumes that all entries are free of mistakes and standardized across systems – which is almost never the case in real-world linkage scenarios.
#
# #### How do you go about determining a match when so many variations exist?
#
# By performing probabilistic data matching, that’s how. More commonly known as fuzzy matching’, this approach permits the user to account for variations like spelling errors, nicknames, punctuation differences, and many more by combining a variety of algorithms.
#
# ### Fuzzy matching is a computer-assisted technique to score the similarity of data.
#
# 
#
#
# ### Fuzzy matching is a technique used in computer-assisted translation as a special case of record linkage. It works with matches that may be less than 100% perfect when finding correspondences between segments of a text and entries in a database of previous translations. It usually operates at sentence-level segments, but some translation technology allows matching at a phrasal level. It is used when the translator is working with translation memory.
#
# Here’s a list of the various fuzzy matching techniques that are in use today:
#
# - Levenshtein Distance (or Edit Distance)
# - Damerau-Levenshtein Distance
# - Jaro-Winkler Distance
# - Keyboard Distance
# - Kullback-Leibler Distance
# - Jaccard Index
# - Metaphone 3
# - Name Variant
# - Syllable Alignment
# - Acronym
#
# #### We won't be going much in detail of each technique
df_fuzzy_match=pd.read_csv("Datasets/room_type.csv")
df_fuzzy_match.head(3)
# #### Importing Library for doing fuzzy match
from fuzzywuzzy import fuzz
# #### Looks like we do not have that installed . So, Module needs installation and here is a command for that!
# !pip3 install fuzzywuzzy
# !python -m pip install --trusted-host pypi.org --trusted-host files.pythonhosted.org fuzzywuzzy
# ## 2.1 RATIO - Compares the entire string similarity <a id="r"> </a>
#
# Ratio function computes the standard Levenshtein distance similarity ratio between two sequences
from fuzzywuzzy import fuzz
Str1 = "Apple Inc."
Str2 = "apple Inc"
Ratio = fuzz.ratio(Str1.lower(),Str2.lower())
print(Ratio)
fuzz.ratio('Deluxe Room, 1 King Bed', 'Deluxe King Room')
fuzz.ratio('Traditional Double Room, 2 Double Beds', 'Double Room with Two Double Beds')
fuzz.ratio('Room, 2 Double Beds (19th to 25th Floors)', 'Two Double Beds - Location Room (19th to 25th Floors)')
# ## 2.2 PARTIAL RATIO - Compares partial string similarity <a id="pr"> </a>
#
# It is a powerful function that allows us to deal with more complex situations such as substring matching
# If the short string has length k and the longer string has the length m, then the algorithm seeks the score of the best matching length-k substring.
Str1 = "Los Angeles Lakers"
Str2 = "Lakers"
Ratio = fuzz.ratio(Str1.lower(),Str2.lower())
Partial_Ratio = fuzz.partial_ratio(Str1.lower(),Str2.lower())
print(Ratio)
print(Partial_Ratio)
fuzz.partial_ratio('Deluxe Room, 1 King Bed', 'Deluxe King Room')
fuzz.partial_ratio('Traditional Double Room, 2 Double Beds', 'Double Room with Two Double Beds')
fuzz.partial_ratio('Room, 2 Double Beds (19th to 25th Floors)', 'Two Double Beds - Location Room (19th to 25th Floors)')
# ## 2.3 TOKEN SORT RATIO - Ignores word order <a id="tr"> </a>
#
# #### What happens when the strings comparison the same, but they are in a different order?
#
# The fuzz.token functions have an important advantage over ratio and partial_ratio. They tokenize the strings and preprocess them by turning them to lower case and getting rid of punctuation. In the case of fuzz.token_sort_ratio(), the string tokens get sorted alphabetically and then joined together. After that, a simple fuzz.ratio() is applied to obtain the similarity percentage.
Str1 = "united states v. nixon"
Str2 = "Nixon v. United States"
Ratio = fuzz.ratio(Str1.lower(),Str2.lower())
Partial_Ratio = fuzz.partial_ratio(Str1.lower(),Str2.lower())
Token_Sort_Ratio = fuzz.token_sort_ratio(Str1,Str2)
print(Ratio)
print(Partial_Ratio)
print(Token_Sort_Ratio)
fuzz.token_sort_ratio('Deluxe Room, 1 King Bed', 'Deluxe King Room')
fuzz.token_sort_ratio('Traditional Double Room, 2 Double Beds', 'Double Room with Two Double Beds')
fuzz.token_sort_ratio('Room, 2 Double Beds (19th to 25th Floors)', 'Two Double Beds - Location Room (19th to 25th Floors)')
# ## 2.4 TOKEN SET RATIO - Ignore duplicate words similarly to token sort ratio <a id="ts"> </a>
#
# #### What happens if these two strings are of widely differing lengths?
#
# That's where fuzz.token_set_ratio() comes in.
# Instead of just tokenizing the strings, sorting and then pasting the tokens back together, token_set_ratio performs a set operation that takes out the common tokens (the intersection) and then makes fuzz.ratio() pairwise comparisons between the following new strings:
#
# - s1 = Sorted_tokens_in_intersection
# - s2 = Sorted_tokens_in_intersection + sorted_rest_of_str1_tokens
# - s3 = Sorted_tokens_in_intersection + sorted_rest_of_str2_tokens
#
# The logic behind these comparisons is that since Sorted_tokens_in_intersection is always the same, the score will tend to go up as these words make up a larger chunk of the original strings or the remaining tokens are closer to each other.
#
Str1 = "The supreme court case of Nixon vs The United States"
Str2 = "Nixon v. United States"
Ratio = fuzz.ratio(Str1.lower(),Str2.lower())
Partial_Ratio = fuzz.partial_ratio(Str1.lower(),Str2.lower())
Token_Sort_Ratio = fuzz.token_sort_ratio(Str1,Str2)
Token_Set_Ratio = fuzz.token_set_ratio(Str1,Str2)
print(Ratio)
print(Partial_Ratio)
print(Token_Sort_Ratio)
print(Token_Set_Ratio)
fuzz.token_set_ratio('Deluxe Room, 1 King Bed', 'Deluxe King Room')
fuzz.token_set_ratio('Traditional Double Room, 2 Double Beds', 'Double Room with Two Double Beds')
fuzz.token_set_ratio('Room, 2 Double Beds (19th to 25th Floors)', 'Two Double Beds - Location Room (19th to 25th Floors)')
# #### As TOKEN SET RATIO is the best for this dataset, let's explore it a bit more.
# +
def get_ratio(row):
name1 = row['Expedia']
name2 = row['Booking.com']
return fuzz.token_set_ratio(name1, name2)
rated = df_fuzzy_match.apply(get_ratio, axis=1)
rated.head(10)
# -
# #### Which ones got a set ratio greater than 70%?
greater_than_70_percent =df_fuzzy_match[rated > 70]
greater_than_70_percent.count()
greater_than_70_percent.head(10)
len(greater_than_70_percent) / len(df_fuzzy_match)
# ##### More than 90% of the records have a score greater than 70%.
greater_than_70_percent = df_fuzzy_match[rated > 60]
greater_than_70_percent.count()
len(greater_than_70_percent) / len(df_fuzzy_match)
# ##### And more than 98% of the records have a score greater than 60%.
# ## 2.5 Fuzzy Process Extract <a id="fz1" > </a>
#
# A module called process that allows you to calculate the string with the highest similarity out of a vector of strings
#
from fuzzywuzzy import process
str2Match = "apple inc"
strOptions = ["Apple Inc.","apple park","apple incorporated","iphone"]
Ratios = process.extract(str2Match,strOptions)
print(Ratios)
# You can also select the string with the highest matching percentage
highest = process.extractOne(str2Match,strOptions)
print(highest)
# # <p style="text-align: center;"> 3.0 Conclusion </p> <a id="Conclusion"> </a>
#
# The world of fuzzy string matching has come a long way. There are a lot more advanced ways that incorporate these concepts into their fuzzy string searches, and there is more room for efficiency
# # <p style="text-align: center;"> 4.0 Contribution</p> <a id='Contribution'> </a>
#
#
#
# This was a fun project in which we explore the idea of Data cleaning and Data Preprocessing. We take inspiration from kaggle learning course and create our own notebook enhancing the same idea and supplementing it with our own contributions from our experiences and past projects.
#
# - Code by self : 65%
# - Code from external Sources : 35%
# # <p style="text-align: center;"> 5.0 Citations <a id="Citation"> </a>
#
# - https://dataladder.com/fuzzy-matching-101/
# - https://medium.com/tim-black/fuzzy-string-matching-at-scale-41ae6ac452c2
# - https://www.kaggle.com/leandrodoze/fuzzy-string-matching-with-hotel-rooms
# - https://medium.com/@julientregoat/an-introduction-to-fuzzy-string-matching-178805cca2ab
# # <p style="text-align: center;"> 6.0 License <a id="License"> </a>
#
# Copyright (c) 2020 <NAME>, <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
| Data Science/Data_Preprocessing/Notebooks/.ipynb_checkpoints/INFO7390_Assignment_3_Mini_Project_One_Part_Six-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Validation report for dmu26_XID+PACS_COSMOS_20170303
# The data product dmu26_XID+PACS_COSMOS_20170303, contains three files:
#
# 1. dmu26_XID+PACS_COSMOS_20170303.fits: The catalogue file
# 2. dmu26_XID+PACS_COSMOS_20170303_Bayes_pval_PACS100.fits: The Bayesian pvalue map
# 3. dmu26_XID+PACS_COSMOS_20170303_Bayes_pval_PACS160.fits: The Bayesian pvalue map
# ## Catalogue Validation
# Validation of the catalogue should cover the following as a minimum:
#
# * Compare XID+ Fluxes with previous catalogues
# * Check for sources with poor convergence (i.e. $\hat{R}$ >1.2 and $n_{eff}$ <40)
# * Check for sources with strange error (i.e. small upper limit and large lower limit, which would be indicating prior is limiting flux)
# * Check for sources that return prior (i.e. probably very large flux and large error)
# * Check background estimate is similar across neighbouring tiles (will vary depending on depth of prior list)
#
#
from astropy.table import Table
import numpy as np
import pylab as plt
# %matplotlib inline
table=Table.read('/Users/williamp/validation/cosmos/PACS/dmu26_XID+PACS_COSMOS_20170303.fits', format='fits')
table[:10].show_in_notebook()
import seaborn as sns
# ### Comparison to previous catalogues
# Using COSMOS2015 catalogue and matching to closest PACS objects within 1''
# +
#table.sort('help_id')
COSMOS2015 = Table.read('/Users/williamp/validation/cosmos/PACS/COSMOS2015_Laigle+v1.1_wHELPids_PACS_red.fits')
COSMOS2015.sort('HELP_ID')
# +
plt.scatter(np.log10(COSMOS2015['F_PACS_160']), np.log10(COSMOS2015['FLUX_160']))
plt.xlabel('$\log_{10}S_{160 \mathrm{\mu m}, XID+}$')
plt.ylabel('$\log_{10}S_{160 \mathrm{\mu m}, COSMOS2015}$')
plt.show()
plot=sns.jointplot(x=np.log10(COSMOS2015['F_PACS_160']), y=np.log10(COSMOS2015['FLUX_160']), xlim=(-1,3), kind='hex')
plot.set_axis_labels('$\log_{10}S_{160 \mathrm{\mu m}, XID+}$', '$\log_{10}S_{160 \mathrm{\mu m}, COSMOS2015}$')
plt.show()
# -
# Agreement is reasonable good. Lower flux objects can be given lower flux density by XID+ as would be expecetd. High flux density objects have a convergance in XID+ and COSMOS2015 as would be expected.
# ### Convergence Statistics
# e.g. How many of the objects satisfy critera?
# (note Some of the $\hat{R}$ values are NaN. This is a PyStan bug. They are most likely 1.0
plt.figure(figsize=(20,10))
plt.subplot(1,2,1)
Rhat=plt.hist(np.isfinite(table['Rhat_PACS_160']), bins=np.arange(0.9,1.2,0.01))
plt.xlabel(r'$\hat{R}$')
plt.subplot(1,2,2)
neff=plt.hist(table['n_eff_PACS_160'])
plt.yscale('log')
plt.xlabel(r'$n_{eff.}$')
# +
numRhat = 0
numNeff = 0
for i in range(0, len(table)):
if table['Rhat_PACS_160'][i] > 1.2 and np.isfinite(table['Rhat_PACS_160']):
numRhat += 1
if table['n_eff_PACS_160'][i] < 40:
numNeff += 1
print(str(numRhat)+' objects have $\hat{R}$ > 1.2')
print(str(numNeff)+' objects have n$_{eff}$ < 40')
# -
# All objects have good $\hat{R}$ and n$_{eff}$ values
# ### Skewness
plot=sns.jointplot(x=np.log10(table['F_PACS_160']), y=(table['FErr_PACS_160_u']-table['F_PACS_160'])/(table['F_PACS_160']-table['FErr_PACS_160_l*1000.0']), xlim=(-1,2), kind='hex')
plot.set_axis_labels(r'$\log_{10}S_{160 \mathrm{\mu m}}$ ', r'$(84^{th}-50^{th})/(50^{th}-16^{th})$ percentiles')
# ### Sources where posterior=prior
# Suggest looking at size of errors to diagnose. How many appear to be returning prior? Where are they on Bayesian P value map? Does it make sense why?
# The lower errors appear to be very, very small. The catalogue appears to have them multiplied by 1000 and they are still 10\,000 times smaller than the upper errors.
# ### Background value
# Are all the background values similar? For those that aren't is it obvious why? (e.g. edge of map, extended source not fitted well etc)
plt.hist(table['Bkg_PACS_160'])
plt.xlabel(r'Background (MJy/sr)')
plt.show()
# The background seems to have quite a large scatter but roughly consistent around ~-1.0
# -------------
# ## Bayesian P value map
# The Bayesian P value map can be thought of as a more robust residual map. They provide the probability our model would obtain the pixel data value, having inferred our model on the data. The probabilitiy value is expressed in terms of a sigma value.
#
# * a value of < -2 indicates that our model cannot explain why there is so little flux in the map
# * a value of > -2 indicates our model cannot explain why there is so much flux in the map
# * a value of ~ 0 indicates a good fit
#
# Plotting the distribution of the Bayesian P value map can indicate whether the map has been been fit well in general. If the distribution is centered on 0 and roughly symmetric then it is a good fit.
#
# Suggested validation checks:
#
# * Check distribution is reasonable
# * Check for strange areas in map
import aplpy
from astropy.io import fits
hdulist=fits.open('/Users/williamp/validation/cosmos/PACS/dmu26_XID+PACS_COSMOS_20170303_Bayes_pval_PACS160.fits')
plt.figure()
Bayes_hist=plt.hist(np.isfinite(hdulist[1].data), bins=np.arange(-6,6.1,0.05))
plt.xlabel(r'$\sigma$ value')
plt.show()
# #Checking the Positions
# Using the PACS map to check where the high flux objects and objects with highish bakground are.
# +
hdulist = fits.open('/Users/williamp/dev/XID_plus/input/cosmosKs/pep_COSMOS_red_Map.DR1.fits')
hdulist[1].header['CTYPE1'] = 'RA'
hdulist[1].header['CTYPE2'] = 'DEC'
# -
# PACS_red map is in greyscale, object positions are in blue, objects with fluxes > 1e3 are in red, objects with background values > 1.25 MJy/sr are in yellow, objects with P-values > 0.5 are in green.
# +
vmin=0.0001
vmax=0.02
fig = plt.figure(figsize=(30,10))
pltut = aplpy.FITSFigure(hdulist[1], figure=fig)
pltut.show_colorscale(vmin=vmin,vmax=vmax,cmap='Greys',stretch='log')
pltut.show_circles(table['RA'],table['Dec'], radius=0.0025, color='b')
bRA = []
bDec = []
for i in range(0, len(table)):
if table['F_PACS_160'][i] > 1e3:
bRA.append(table['RA'][i])
bDec.append(table['Dec'][i])
if len(bRA) > 0:
pltut.show_circles(bRA, bDec, radius=0.005, color='r')
bgRA = []
bgDec = []
for i in range(0, len(table)):
if table['Bkg_PACS_160'][i] > 1.25:
bgRA.append(table['RA'][i])
bgDec.append(table['Dec'][i])
if len(bgRA) > 0:
pltut.show_circles(bgRA, bgDec, radius=0.005, color='y')
pRA = []
pDec = []
for i in range(0, len(table)):
if table['Pval_res_160'][i] > 0.5:
pRA.append(table['RA'][i])
pDec.append(table['Dec'][i])
if len(pRA) > 0:
pltut.show_circles(pRA, pDec, radius=0.005, color='g')
plt.show()
# -
# The high flux value objects (red) appear to mainly cluster arround the edges of the masked regions, so may just be artefacts of bad masking.
#
# The high background objects (yellow) appear to be a property of individual tiles as they form squares on the image above. These tiles are on the edge of the map and the high background is probably a result of this.
#
# The high P-value objects (green) appear arround brighter objects (but not all bright objects) which makes sense; bright blended objects are harder to deblend than non-blended objects. They also often coincide with the high background tiles.
| dmu26/dmu26_XID+PACS_COSMOS/validation_report-PACS_red.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.0
# language: julia
# name: julia-0.4
# ---
# # Lex-Minimizer #
# In this notebook, we will demonstrate the use of functions that computes the inf-minimizer (`CompInfMin`) and lex-minimizer (`CompLexMin`), as well as other experiments. You can find all source code in `lex.jl`.
pwd()
push!(LOAD_PATH,"../src")
using Laplacians
using PyPlot
include("../src/lex.jl") # for development
# ### IterLex Functions ###
# * `simIterLexUnwtd(...)` runs IterLex on a **uniformly weighted** graph for a specific number of iterations;
#
# * `checkLexUnwtd(...)` checks the correctness of the assignment up to `LEX_EPS`;
#
# * `simIterLex(...)` runs IterLex on any weighted graph;
#
# * `checkLex(...)` checks the correctness of the assignment up to `LEX_EPS`;
# ## Simple examples ##
# ### Path Graphs ###
# A path graph with `n` vertices. Let vertex `1` and `n` be terminals, with voltages `0` and `1` respectively.
# +
# Set up
n = 10
Pn = pathGraph(n)
isTerm = zeros(Bool, n)
isTerm[1] = true
isTerm[n] = true
initVal = zeros(n)
initVal[n] = 1.0
# +
# inf-minimizer
infMinVolt = CompInfMin(Pn, isTerm, initVal)
println(infMinVolt)
println(MaxEdgeGrad(Pn, infMinVolt))
# lex-minimizer
lexMinVolt = CompLexMin(Pn, isTerm, initVal)
println(lexMinVolt)
# -
# default error tolerance
LEX_EPS
# +
t1 = 5
asgnmt = simIterLexUnwtd(t1, Pn, isTerm, initVal)
@printf("After %d iterations: %s\n", t1, checkLexUnwtd(Pn, isTerm, initVal, asgnmt, fatal = false))
t2 = 500
asgnmt = simIterLexUnwtd(t2, Pn, isTerm, initVal)
@printf("After %d iterations: %s\n", t2, checkLexUnwtd(Pn, isTerm, initVal, asgnmt, fatal = false))
# -
iterVolt = copy(initVal)
i = 0
# the last parameter is to tell CheckLex not to complain when it sees a value that's worse than epsilon approx
while (!CheckLex(Pn, isTerm, initVal, iterVolt, LEX_EPS, false))
i += 1
iterVolt = simIterLexUnwtd(1, Pn, isTerm, iterVolt)
end
@printf("After %d iterations, IterLex on path graph with %d vertices has an error <= %.2e", i, n, LEX_EPS)
# Plotting the number of iteration needed vs. number of nodes in the path graph, we can see that it takes about O(n^2) iterations.
# +
MAXN = 20
iterArr = zeros(Int64, MAXN)
for n in 3:MAXN
Pn = pathGraph(n)
isTerm = zeros(Bool, n)
isTerm[1] = true
isTerm[n] = true
initVal = zeros(n)
initVal[n] = 1.0
iterVolt = copy(initVal)
i = 0
while (!CheckLex(Pn, isTerm, initVal, iterVolt, 1e-14, false))
i += 1
iterVolt = simIterLexUnwtd(1, Pn, isTerm, iterVolt)
end
iterArr[n] = i
end
x = collect(1:20)
x2 = x .* x * 5 # estimate
y = copy(iterArr)
plot(x, y, linewidth=1.0, "o-", x, x2, "g^--")
# -
# ### Star Graph ###
# +
# Star Graph: simplest example:
# picking the right pair of neighbors to average
n = 5
Sn = zeros(n, n)
Sn[1,:] = a = [0, 1/20, 1/20, 1/10, 1/18]
Sn[:,1] = a'
Sn = sparse(Sn)
isTerm = ones(Bool, n)
isTerm[1] = false
initVal = [0.0, 20, -5, -5, 17]
asgnmt = simIterLex(1, Sn, isTerm, initVal)
# -
checkLex(Sn, isTerm, initVal, asgnmt, fatal = false)
# ### Random Graph ###
n = 20
G = chimera(n)
# +
isTerm = zeros(Bool, n)
# arbitrary terminal values
isTerm[1] = true
isTerm[5] = true
isTerm[11] = true
isTerm[18] = true
initVal = zeros(Float64, n)
initVal[1] = 0.0
initVal[5] = 13
initVal[11] = 7
initVal[18] = 11
# -
infMinVolt = CompInfMin(G, isTerm, initVal)
println(infMinVolt)
println(MaxEdgeGrad(G, infMinVolt))
lexMinVolt = simIterLex(500, G, isTerm, initVal)
println(lexMinVolt)
println(MaxEdgeGrad(G, lexMinVolt))
println(checkLex(G, isTerm, initVal, lexMinVolt))
# ## Lex Algorithm Related ##
# * `termFreeShortestPaths` gives the shortest paths from the vertex `start` to every other vertex without going through a terminal;
# *
G = readIJV("testLexGraph.txt")
n = G.n
# +
isTerm = zeros(Bool, n)
# arbitrary terminal values
isTerm[1] = true
isTerm[5] = true
isTerm[19] = true
initVal = zeros(Float64, n)
initVal[1] = 0.0
initVal[5] = 13
initVal[19] = 11
# -
include("../src/lex.jl")
lexMinVolt = CompLexMin(G, isTerm, initVal)
checkLex(G, isTerm, initVal, lexMinVolt)
setLexDebugFlag(true)
LEX_DEBUG
lexMinVolt
MaxEdgeGrad(G, lexMinVolt)
| notebooks/Lex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Session 2 Homework
# =================
#
# ## Exercise 1
#
# In the first class, we examined a dataset containing molecular descriptors and the results from the ESOL solubility model. In this homework, we are going to use the `tabula` library to extract a table from the associated paper. In your `pdf` folder, you have a file called `Delaney_paper.pdf`. Your task for this homework is to pull information from table 4 and create plots using seaborn showing the correlation between the various solubility models and the experimental values.
#
# Using the file `pdfs/Delaney_paper.pdf` and `tabula-py`, read in the data from Table 4. Save this file in your data folder as `delaney_table4.csv`.
#
# ## Exercise 2
#
# Read in your saved data and clean the table using pandas.
#
# - Rename the columns to have easy and descriptive names.
# - Use `pd.to_numeric` with the appropriate options to cast columns to floats.
# - Drop rows or columns if necessary.
#
# Your dataframe should have the following column names before you save it:
#
# ```
# ['common name', 'CAS no.', 'experimental values', 'ESOL', 'Liu',
# 'Huuskonen', 'Kuhne', 'Wegner', 'Gasteiger', 'Tetko', 'GSE']
# ```
#
# Save your cleaned dataframe as `delaney_table4_clean.csv`
#
# ## Exercise 3
#
# Use seaborn `lmplot` to create a plot showing `experimental values` vs each of the models. You should have one plot per model. Create a plot which has two columns per row.
#
# **Bonus** - Use seaborn to visualize the correlation between experimental values and the models. Which one has the highest correlation? The method `df.corr` gives you Pearson's correlation coefficient (R). Table 4 reports $R^2$ - can you get the values to match? (how?)
| book/2-homework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Darkdev88/Corso_fuzzy_2021-1/blob/main/Tesina_Logica_Fuzzy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qK1Mg4J2YM3I"
# **Introduzione**
#
# La logica è nata dalla necessità di formalizzare il linguaggio parlato.
# In passato, quando si andava a porre un quesito o ad esporre una situazione, si incappava sempre nel solito problema: la difficoltà nell'esprimere un'idea nel modo più oggettivo possibile.
# Per esprimere i formalismi più basilari furono introdotti i sillogismi.
# Nel momento in cui entrò in gioco la matematica però, si riscontrarono importanti innovazioni anche dal punto di vista linguistico.
# Un linguaggio unico però, che mettesse tutti d'accordo, ancora non esisteva.
# Fu Boole, il primo a formalizzare e a studiare la logica da un punto di vista algebrico tra il 1847 e il 1850.
# Suoi gli enunciati che spiegano la prassi per la trascrizione delle formule e la riproduzione dei connettivi.
# La logica matematica quindi, inizialmente si focalizzò sull'aspetto sintattico della lingua, ossia sul modo in cui si descrivono le cose.
# Boole in particolare introdusse algebre costituite da due estremi: 0 e 1. Lo "0" va interpretato come falso e l' "1" come vero.
# In questo modo abbiamo un formalismo e possiamo parlare di più oggetti e la logica inizia ad avere un suo ruolo attivo nella matematica.
# La Logica proposizionale si occupa invece di studiare la verità e la falsità delle affermazioni, che possono essere formulate con l'ausilio della matematica.
# In ambito logico, si definisce "proposizione" qualsiasi affermazione di cui si possa determinare un valore di verità (vero o falso), senza che ciò comporti ambiguità.
# In seguito Lukasiewicz, a partire dalla Logica a due valori introdusse un nuovo concetto, in virtù del quale un costrutto deve fornirmi un valore di verità diverso da 0 e 1.
# Introdusse inoltre il "non lo so"; vale a dire il terzo valore compreso tra 0 e 1; quello tra bianco assoluto e nero assoluto a cui fanno riferimento tutte le gradazioni di grigio, ossia infiniti valori.
# La logica fuzzy è invece una branca della matematica in cui si può attribuire a ciascuna proposizione un grado di verità diverso da 0 e 1 e compreso tra di loro. È una logica polivalente, ossia un'estensione della logica booleana.
# È legata alla "Teoria degli insiemi sfocati con grado di verità o valore di appartenenza" con cui si analizza quanto è vera una proprietà che può essere, oltre che vera (= a valore 1) o falsa (= a valore 0) come nella logica classica, anche parzialmente vera e parzialmente falsa.
#
#
# + [markdown] id="bk5pepvdlH3s"
# **Connettivi Logici in Python**
# + id="yVTY1Qr9kPQS"
def Congiunzione(var1,var2):
return var1 and var2
def Disgiunzione(var1,var2):
return var1 or var2
def ImplicazioneSemplice(ipotesi,tesi):
return (not ipotesi or tesi)
def oppure(var1,var2):
return (not var1 and var2) or (var1 and not var2)
# + id="nhZBNspPS-6C"
def stampa_tabella(formula):
valori_possibili = (False, True)
for A in valori_possibili:
for B in valori_possibili:
print(f'{formula(A,B)}')
# + colab={"base_uri": "https://localhost:8080/"} id="yE5c1sQNVAND" outputId="8dcd377c-5e0a-4705-a789-1e6a8ea66bd7"
stampa_tabella(oppure)
# + [markdown] id="Yjs0Rt8_se8X"
# **Tabella di verità**
#
# Si è preferito utilizzare questo codice dal momento che è quello risulta essere più chiaro ad un programmatore evitando interpretazioni errate.
# Più in dettaglio è stata utilizzata la funzione AND.
# Va previsato inoltre che l' utilizzo di due cicli for risulti particolarmente dispendioso per python dove non vengono dichiarate le variabili a differenza degli altri linguaggi, ad esempio C.
# + colab={"base_uri": "https://localhost:8080/"} id="xfx9aZp7sxaA" outputId="fa9397d1-f626-4eab-dcf4-38dfc5aa3207"
valori_possibili = (False, True)
for A in valori_possibili:
for B in valori_possibili:
print(f'{A} AND {B} = {A and B}')
# + [markdown] id="bgs_EOAPDVQT"
# **Funzioni di verità delle seguenti formule :**
#
# **A or (not A)**
#
# **not A -> B**
# + id="Cgs1SAh3IVgP"
#Funzione 1
def uno(var1):
return var1 or (not var1)
#Funzione 2
def due(var1, var2):
return (not (not var1) or var2)
# + id="yL1I23oNH8d5"
def stampa_tabella(formula):
valori = (0,1)
print('\nTabella di verita\':')
if formula == 'uno':
for A in valori:
print(f'{A} | {eval(formula+"(A)")}')
else:
for A in valori:
for B in valori:
print(f' {A} | {B} | {eval(formula+"(A,B)")}')
# + colab={"base_uri": "https://localhost:8080/"} id="otv73auyIbvk" outputId="0222522f-47cf-4a45-d3e7-d3e213621ce3"
print('Stampa Funzione')
risposta = 's'#per terminare il while
while risposta == 's':
#input utente
operazione = input('Quale delle due operazioni vuoi effettuare ? "uno" "due"')
#concateno le stringhe per richiamare delle funzioni
stampa_tabella(operazione)
risposta = input('\nUn\'altra Operazione? Si(s), no(n)')
print('\n\nGrazie per avermi utilizzato!')
| Tesina_Logica_Fuzzy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook uses [the world population data from World Bank](https://data.worldbank.org/indicator/SP.POP.TOTL)
#
# The code below loads the data from the CSV file
import pandas as pd
data = pd.read_csv('data/WorldBank_world_population.csv', skiprows=3).set_index('Country Name')
data.head()
data.index
data.columns
data.dtypes
pop2018 = data['2018']
pop2018.head()
type(pop2018)
# # Indexing and selection with Series
#
# `pop2018` is a series that contains population data for year 2018
#
# Answer the following questions using indexing and selection operators
pop2018['United States']
pop2018.loc['United States']
# - What is the population of the `United States`?
pop2018['United States'] = pop2018['United States'] + 1
# - Increase that number by 1
pop2018[ ['United States', 'Canada', 'Mexico'] ]
pop2018.loc[ ['United States', 'Canada', 'Mexico'] ]
# - Select the population values of the `United States`, `Canada`, and `Mexico`?
pop2018.head(5)
pop2018.iloc[:5]
pop2018[:5]
# - Select the population values of the first five countries in the series
pop2018.tail()
pop2018[-5:]
pop2018.iloc[-5:]
# - Select the population values of the last five countries in the series
pop2018[pop2018 > 1e9]
# - Select the population values that are larger than one billion
# # Indexing and selection with DataFrame
# - Select data of years 2017 and 2018
# +
#data['2017':'2018']
# -
data[['2017', '2018']]
data.loc[:, ['2017', '2018']]
data.loc[:, '2017':'2018']
# - Select entries of countries `United States`, `Canada`, and `Mexico`
# data.loc[['United States', 'Canada', 'Mexico'] # or
data.loc[['United States', 'Canada', 'Mexico'], :]
# - Select data of countries `United States`, `Canada`, and `Mexico` in years 2017 and 2018
data.loc[['United States', 'Canada', 'Mexico'], '2017':'2018']
data.loc[['United States', 'Canada', 'Mexico'], ['2017', '2018']]
# - Select data of years 2000 to 2018
data.loc[:, '2000':'2018']
# - Select entries of countries from `Aruba` to `Azerbaijan`?
data.loc['Aruba':'Azerbaijan', :]
# - Select entries of countries that had more that a billion people in year 2000?
data[data['2000'] > 1e9]
data.loc[data['2000'] > 1e9]
# - Select the first five rows
data.head()
data[:5]
data.iloc[:5]
# - Select the first five rows and the first five columns
data.iloc[:5, :5]
| 15-warmup-solution_dataframe_and_series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: aiffel
# language: python
# name: aiffel
# ---
# # Google Review Keyword Frequency
#
# ## 수행 요소
# 1. keyword별 빈도수 리스트 파일로 저장
# ## Data path 확인
# !pwd # 현재 경로 확인
# 아래 tree 명령어로 상위 폴더(jungcheck)내 파일 구성을 볼 수 있습니다.
# !ls . # 현재 위치의 file list
# !tree -L 3 -N ./data/google_review # jungcheck 폴더내 경로 확인
# ## Library Import
# +
import os
import glob
from tqdm import tqdm
import re
import pandas as pd
import copy # deep copy
# -
# ## Data Path & Load
root = os.getenv('HOME') + '/jungcheck'
root
data_path = os.path.join(root, 'data/google_review/')
data_path
folders = os.listdir(data_path)
print(folders)
print(len(folders)) # 17개 대상 관광지 중 15개
# +
keywords_ko = [file_name[2:] for file_name in folders]
keywords_en = ['jangtae_mountain', 'gyejok_mountain', 'dongchundang', 'uineungjeongi_street', 'ppuri_park',
'expo_science_park', 'sungsimdang_bakery', 'water_barrel', 'yuseong_hotspring', 'hanbat_arboretum',
'science_museum','daecheong_lake', 'art_culture_complex', 'observatory', 'oworld_zoo']
keywords = {k:v for k, v in zip(keywords_ko, keywords_en)}
keywords
# -
filtered_path = os.path.join(root, 'filtered/google_review/')
filtered_path
folders = os.listdir(filtered_path)
print(folders)
print(len(folders)) # 17개 대상 관광지 중 15개
# key <=> value
keywords = {v:k for k, v in keywords.items()}
keywords
# 👉 영어 keyword로 한글로 된 keyword를 살펴볼거기 때문에 key와 value를 바꿔줍니다.
# root_path내의 dataframe을 합해주는 함수
def concatCsv(root_path:str, files_list:list):
data_list = []
for file in files_list:
file_path = os.path.join(root_path, file)
print(file_path)
df = pd.read_csv(file_path, encoding='utf-8') # csv 파일 읽기
data_list.append(df)
df = pd.concat(data_list, axis=0)
return df
filtered_df = concatCsv(filtered_path, folders)
len(filtered_df)
for key_en, key_ko in keywords.items():
cnt = filtered_df[filtered_df['keyword']==key_en]['search'].count()
print(f'{key_en}과 연관된 검색어는 {cnt}개')
print(list(set(filtered_df[filtered_df['keyword']==key_en]['search'])))
print('-'*40)
# ## Frequency
# +
from soynlp.noun import LRNounExtractor
from soynlp.noun import LRNounExtractor_v2
from soynlp.noun import NewsNounExtractor
def get_noun_sorted_by_frequency(noun_extractor):
"""
Description
이미 학습되어 있는 noun_extractor의 noun을
frequency 내림차순으로 정렬하여 리턴함.
"""
return_lst = list()
nouns = noun_extractor.extract()
for (each_noun, each_noun_info) in nouns.items():
each_noun_dict = {
"noun": each_noun,
"frequency": each_noun_info.frequency,
"score": each_noun_info.score
}
return_lst.append(each_noun_dict)
return_lst = sorted(return_lst, key=lambda x: x['frequency'], reverse=True)
return return_lst
# -
noun_extractor_dict = {
"LRNounExtractor": LRNounExtractor(verbose=False),
"LRNounExtractor_v2": LRNounExtractor_v2(verbose=False),
"NewsNounExtractor": NewsNounExtractor(verbose=False)
}
# ### Sample
target = 'oworld_zoo'
test_list = filtered_df[filtered_df['keyword']==target]['comment']
# filtered_list = ' '.join(test_list)
for noun_ext_name, noun_ext in noun_extractor_dict.items():
# train
noun_ext.train(test_list)
print(f"== extractor: {noun_ext_name}")
# noun을 빈도 내림차순으로 정렬하여 리턴
noun_lst = get_noun_sorted_by_frequency(noun_ext)
for noun in noun_lst[:20]:
print(noun)
print("--" * 50)
# 👉 NewsNounExtractor가 명사 중심으로 추출을 잘해내는 것을 확인했으므로 해당 명사추출기를 사용합니다.
for noun_dict in noun_lst:
target_freq = 0
if target in noun_dict['noun']:
target_freq = noun_dict.get('frequency')
continue
print(noun_dict.get('noun'))
print('-'*20)
print(f'검색 관광지명인 {target}가 {target_freq}번 언급됐습니다.')
# +
noun_ext = noun_extractor_dict["NewsNounExtractor"]
noun_lst = get_noun_sorted_by_frequency(noun_ext)
data_list = []
for noun_dict in noun_lst:
noun = noun_dict.get('noun')
freq = noun_dict.get('frequency')
score = noun_dict.get('score')
data_list.append([target, noun, freq, score])
# -
noun_dict.keys()
pd.DataFrame(data=data_list, columns=['target']+list(noun_dict.keys()))
# ### 전체 Set에 적용 후 저장
keywords
save_path = os.path.join(os.getcwd(), 'frequency/')
save_path
# +
noun_ext = noun_extractor_dict["NewsNounExtractor"]
for key_en, _ in keywords.items():
noun_ext.train(filtered_df[filtered_df['keyword']==key_en]['comment'])
noun_lst = get_noun_sorted_by_frequency(noun_ext)
data_list = []
for noun_dict in noun_lst:
noun = noun_dict.get('noun')
freq = noun_dict.get('frequency')
score = noun_dict.get('score')
data_list.append([key_en, noun, freq, score])
df = pd.DataFrame(data=data_list, columns=['keyword']+list(noun_dict.keys()))
df.to_csv(save_path + key_en + '.csv', header=True, index=False)
# -
# 👉 명사 추출기를 썼음에도 좋아요와 같은 동사가 남는 현상이 있었습니다.
# 먼저, Mecab의 품사태깅을 활용해 한번더 명사를 정제해줬습니다.
# ### 데이터 다시 정제 후 품사태깅
test = filtered_df['comment'].iloc[0]
test
# +
from konlpy.tag import Kkma, Hannanum, Komoran, Mecab, Twitter
mecab = Mecab()
## 전체 품사 태깅
speech_pos = mecab.pos(filtered_df['comment'].iloc[0])
print(speech_pos)
# -
# 👉 Mecab으로 하더라도 문장의 띄어쓰기 문제가 해결되지 않은 경우에는 품사태깅이 잘 이뤄지지 않음을 알 수 있습니다. 데이터 정제 방법을 고민했을 때 구글 검색에서 검색어를 정제해주는 기능을 활용하기 위해 셀레니움을 통해 읽어오는 방향을 시도해볼까 고민중..
# 
| googleKeywordFrequency.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.5.0
# language: julia
# name: julia-0.5
# ---
using FileIO
using MeshIO
using GeometryTypes
using StaticArrays
using Plots
gr()
using DrakeVisualizer
using Interact
DrakeVisualizer.new_window()
mesh = load("/Users/rdeits/locomotion/drake-distro/drake/examples/Atlas/urdf/meshes/pelvis_chull.obj")
Visualizer(mesh)
import AdaptiveDistanceFields
reload("AdaptiveDistanceFields")
s = AdaptiveDistanceFields.ConvexMesh.signed_distance(mesh)
origin = SVector(-2., -2, -2)
widths = SVector(4., 4, 4)
adaptive = AdaptiveDistanceFields.AdaptiveDistanceField(s, origin, widths, 1e-2, 1e-2)
# +
verts = Point{3, Float64}[]
faces = Face{3, Int, 0}[]
for cell in RegionTrees.allcells(adaptive.root)
for face in RegionTrees.faces(cell.boundary)
i = length(verts)
for vert in face
push!(verts, Point(vert[1], vert[2], vert[3]))
end
# push!(faces, Face(i+1, i+2, i+1))
# push!(faces, Face(i+1, i+3, i+1))
# push!(faces, Face(i+2, i+4, i+2))
# push!(faces, Face(i+3, i+4, i+3))
push!(faces, Face(i+1, i+2, i+4))
push!(faces, Face(i+4, i+3, i+1))
end
end
boxmesh = HomogenousMesh(verts, faces)
Visualizer(boxmesh);
# -
xs = linspace(-2, 2)
plot(xs, [s(SVector(x, 0.0, 0.0)) for x in xs], ylim=(-1, 2))
plot!(xs, [adaptive(SVector(x, 0.0, 0.0)) for x in xs], ylim=(-1, 2))
@manipulate for isolevel in linspace(-1.0, 1.0, 101)
Visualizer(contour_mesh(s, SVector(-1., -1, -1), SVector(1., 1, 1), isolevel))
end
@manipulate for isolevel in linspace(-1.0, 1.0, 101)
Visualizer(contour_mesh(adaptive, SVector(-1., -1, -1), SVector(1., 1, 1), isolevel, 0.02))
end
using BenchmarkTools
p = SVector(0.25, 0, 0)
@benchmark $adaptive($p)
| mesh_distance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/120Davies/DS-Unit-2-Kaggle-Challenge/blob/master/Ro_Davies_assignment_kaggle_challenge_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="7IXUfiQ2UKj6" colab_type="text"
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Kaggle Challenge, Module 3
#
# ## Assignment
# - [ ] [Review requirements for your portfolio project](https://lambdaschool.github.io/ds/unit2/portfolio-project/ds6), then choose your dataset, and [submit this form](https://forms.gle/nyWURUg65x1UTRNV9), due today at 4pm Pacific.
# - [ ] Continue to participate in our Kaggle challenge.
# - [ ] Try xgboost.
# - [ ] Get your model's permutation importances.
# - [ ] Try feature selection with permutation importances.
# - [ ] Submit your predictions to our Kaggle competition. (Go to our Kaggle InClass competition webpage. Use the blue **Submit Predictions** button to upload your CSV file. Or you can use the Kaggle API to submit your predictions.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
# ## Stretch Goals
#
# ### Doing
# - [ ] Add your own stretch goal(s) !
# - [ ] Do more exploratory data analysis, data cleaning, feature engineering, and feature selection.
# - [ ] Try other categorical encodings.
# - [ ] Try other Python libraries for gradient boosting.
# - [ ] Look at the bonus notebook in the repo, about monotonic constraints with gradient boosting.
# - [ ] Make visualizations and share on Slack.
#
# ### Reading
#
# Top recommendations in _**bold italic:**_
#
# #### Permutation Importances
# - _**[Kaggle / <NAME>: Machine Learning Explainability](https://www.kaggle.com/dansbecker/permutation-importance)**_
# - [<NAME>: Interpretable Machine Learning](https://christophm.github.io/interpretable-ml-book/feature-importance.html)
#
# #### (Default) Feature Importances
# - [<NAME>: Selecting good features, Part 3, Random Forests](https://blog.datadive.net/selecting-good-features-part-iii-random-forests/)
# - [<NAME>, et al: Beware Default Random Forest Importances](https://explained.ai/rf-importance/index.html)
#
# #### Gradient Boosting
# - [A Gentle Introduction to the Gradient Boosting Algorithm for Machine Learning](https://machinelearningmastery.com/gentle-introduction-gradient-boosting-algorithm-machine-learning/)
# - _**[A Kaggle Master Explains Gradient Boosting](http://blog.kaggle.com/2017/01/23/a-kaggle-master-explains-gradient-boosting/)**_
# - [_An Introduction to Statistical Learning_](http://www-bcf.usc.edu/~gareth/ISL/ISLR%20Seventh%20Printing.pdf) Chapter 8
# - [Gradient Boosting Explained](http://arogozhnikov.github.io/2016/06/24/gradient_boosting_explained.html)
# - _**[Boosting](https://www.youtube.com/watch?v=GM3CDQfQ4sw) (2.5 minute video)**_
#
# #### Categorical encoding for trees
# - [Are categorical variables getting lost in your random forests?](https://roamanalytics.com/2016/10/28/are-categorical-variables-getting-lost-in-your-random-forests/)
# - [Beyond One-Hot: An Exploration of Categorical Variables](http://www.willmcginnis.com/2015/11/29/beyond-one-hot-an-exploration-of-categorical-variables/)
# - _**[Categorical Features and Encoding in Decision Trees](https://medium.com/data-design/visiting-categorical-features-and-encoding-in-decision-trees-53400fa65931)**_
# - _**[Coursera — How to Win a Data Science Competition: Learn from Top Kagglers — Concept of mean encoding](https://www.coursera.org/lecture/competitive-data-science/concept-of-mean-encoding-b5Gxv)**_
# - [Mean (likelihood) encodings: a comprehensive study](https://www.kaggle.com/vprokopev/mean-likelihood-encodings-a-comprehensive-study)
# - [The Mechanics of Machine Learning, Chapter 6: Categorically Speaking](https://mlbook.explained.ai/catvars.html)
#
# #### Imposter Syndrome
# - [Effort Shock and Reward Shock (How The Karate Kid Ruined The Modern World)](http://www.tempobook.com/2014/07/09/effort-shock-and-reward-shock/)
# - [How to manage impostor syndrome in data science](https://towardsdatascience.com/how-to-manage-impostor-syndrome-in-data-science-ad814809f068)
# - ["I am not a real data scientist"](https://brohrer.github.io/imposter_syndrome.html)
# - _**[Imposter Syndrome in Data Science](https://caitlinhudon.com/2018/01/19/imposter-syndrome-in-data-science/)**_
#
#
#
#
#
# + [markdown] id="Wd6T-m6bneLS" colab_type="text"
# ### Python libraries for Gradient Boosting
# - [scikit-learn Gradient Tree Boosting](https://scikit-learn.org/stable/modules/ensemble.html#gradient-boosting) — slower than other libraries, but [the new version may be better](https://twitter.com/amuellerml/status/1129443826945396737)
# - Anaconda: already installed
# - Google Colab: already installed
# - [xgboost](https://xgboost.readthedocs.io/en/latest/) — can accept missing values and enforce [monotonic constraints](https://xiaoxiaowang87.github.io/monotonicity_constraint/)
# - Anaconda, Mac/Linux: `conda install -c conda-forge xgboost`
# - Windows: `conda install -c anaconda py-xgboost`
# - Google Colab: already installed
# - [LightGBM](https://lightgbm.readthedocs.io/en/latest/) — can accept missing values and enforce [monotonic constraints](https://blog.datadive.net/monotonicity-constraints-in-machine-learning/)
# - Anaconda: `conda install -c conda-forge lightgbm`
# - Google Colab: already installed
# - [CatBoost](https://catboost.ai/) — can accept missing values and use [categorical features](https://catboost.ai/docs/concepts/algorithm-main-stages_cat-to-numberic.html) without preprocessing
# - Anaconda: `conda install -c conda-forge catboost`
# - Google Colab: `pip install catboost`
# + [markdown] id="o1nF5eU9nJwL" colab_type="text"
# ### Categorical Encodings
#
# **1.** The article **[Categorical Features and Encoding in Decision Trees](https://medium.com/data-design/visiting-categorical-features-and-encoding-in-decision-trees-53400fa65931)** mentions 4 encodings:
#
# - **"Categorical Encoding":** This means using the raw categorical values as-is, not encoded. Scikit-learn doesn't support this, but some tree algorithm implementations do. For example, [Catboost](https://catboost.ai/), or R's [rpart](https://cran.r-project.org/web/packages/rpart/index.html) package.
# - **Numeric Encoding:** Synonymous with Label Encoding, or "Ordinal" Encoding with random order. We can use [category_encoders.OrdinalEncoder](https://contrib.scikit-learn.org/categorical-encoding/ordinal.html).
# - **One-Hot Encoding:** We can use [category_encoders.OneHotEncoder](http://contrib.scikit-learn.org/categorical-encoding/onehot.html).
# - **Binary Encoding:** We can use [category_encoders.BinaryEncoder](http://contrib.scikit-learn.org/categorical-encoding/binary.html).
#
#
# **2.** The short video
# **[Coursera — How to Win a Data Science Competition: Learn from Top Kagglers — Concept of mean encoding](https://www.coursera.org/lecture/competitive-data-science/concept-of-mean-encoding-b5Gxv)** introduces an interesting idea: use both X _and_ y to encode categoricals.
#
# Category Encoders has multiple implementations of this general concept:
#
# - [CatBoost Encoder](http://contrib.scikit-learn.org/categorical-encoding/catboost.html)
# - [James-Stein Encoder](http://contrib.scikit-learn.org/categorical-encoding/jamesstein.html)
# - [Leave One Out](http://contrib.scikit-learn.org/categorical-encoding/leaveoneout.html)
# - [M-estimate](http://contrib.scikit-learn.org/categorical-encoding/mestimate.html)
# - [Target Encoder](http://contrib.scikit-learn.org/categorical-encoding/targetencoder.html)
# - [Weight of Evidence](http://contrib.scikit-learn.org/categorical-encoding/woe.html)
#
# Category Encoder's mean encoding implementations work for regression problems or binary classification problems.
#
# For multi-class classification problems, you will need to temporarily reformulate it as binary classification. For example:
#
# ```python
# encoder = ce.TargetEncoder(min_samples_leaf=..., smoothing=...) # Both parameters > 1 to avoid overfitting
# X_train_encoded = encoder.fit_transform(X_train, y_train=='functional')
# X_val_encoded = encoder.transform(X_train, y_val=='functional')
# ```
#
# **3.** The **[dirty_cat](https://dirty-cat.github.io/stable/)** library has a Target Encoder implementation that works with multi-class classification.
#
# ```python
# dirty_cat.TargetEncoder(clf_type='multiclass-clf')
# ```
# It also implements an interesting idea called ["Similarity Encoder" for dirty categories](https://www.slideshare.net/GaelVaroquaux/machine-learning-on-non-curated-data-154905090).
#
# However, it seems like dirty_cat doesn't handle missing values or unknown categories as well as category_encoders does. And you may need to use it with one column at a time, instead of with your whole dataframe.
#
# **4. [Embeddings](https://www.kaggle.com/learn/embeddings)** can work well with sparse / high cardinality categoricals.
#
# _**I hope it’s not too frustrating or confusing that there’s not one “canonical” way to encode categorcals. It’s an active area of research and experimentation! Maybe you can make your own contributions!**_
# + id="o9eSnDYhUGD7" colab_type="code" colab={}
# If you're in Colab...
import os, sys
in_colab = 'google.colab' in sys.modules
if in_colab:
# Install required python packages:
# category_encoders, version >= 2.0
# eli5, version >= 0.9
# pandas-profiling, version >= 2.0
# plotly, version >= 4.0
# !pip install --upgrade category_encoders eli5 pandas-profiling plotly
# Pull files from Github repo
os.chdir('/content')
# !git init .
# !git remote add origin https://github.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge.git
# !git pull origin master
# Change into directory for module
os.chdir('module3')
# + id="QJBD4ruICm1m" colab_type="code" colab={}
import pandas as pd
from sklearn.model_selection import train_test_split
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv('../data/tanzania/train_features.csv'),
pd.read_csv('../data/tanzania/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv('../data/tanzania/test_features.csv')
sample_submission = pd.read_csv('../data/tanzania/sample_submission.csv')
train, val = train_test_split(train, train_size = 0.8, test_size = 0.2, stratify = train['status_group'], random_state = 1)
import numpy as np
def wrangle(X):
X = X.copy()
X['latitude']=X['latitude'].replace(-2e-08, 0)
cols_with_zeroes = ['longitude', 'latitude', 'construction_year', 'gps_height', 'population']
for col in cols_with_zeroes:
X[col] = X[col].replace(0, np.nan)
X[col+'_MISSING'] = X[col].isnull()
duplicates = ['quantity_group', 'payment_type']
X = X.drop(columns = duplicates)
unusable_variance = ['recorded_by', 'id']
X = X.drop(columns = unusable_variance)
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.month
X = X.drop(columns = 'date_recorded')
X['years'] = X['year_recorded'] - X['construction_year']
X['years_MISSING'] = X['years'].isnull()
return X
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
# + id="W88LDOv1ZnMc" colab_type="code" colab={}
target = 'status_group'
X_train = train.drop(columns=target)
y_train = train[target]
X_val = val.drop(columns = target)
y_val = val[target]
X_test = test
# + id="QmMQvKRvZ5Pf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="92900828-c7e1-4aea-8be8-76a7cbb6d8cb"
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy = 'median'),
RandomForestClassifier(n_estimators = 100, random_state = 1, n_jobs =-1)
)
pipeline.fit(X_train, y_train)
print('accuracy:', pipeline.score(X_val, y_val))
# + id="IwBM2kC8bAfv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 607} outputId="99e9166e-5711-4f06-aeea-021453946c72"
rf = pipeline.named_steps['randomforestclassifier']
importances = pd.Series(rf.feature_importances_, X_train.columns)
# %matplotlib inline
import matplotlib.pyplot as plt
n = 20
plt.figure(figsize = (10, 10))
plt.title(f'Top {n} features')
importances.sort_values()[-n:].plot.barh(color='grey');
# + id="ReKVc4hSb3kD" colab_type="code" colab={}
| Ro_Davies_assignment_kaggle_challenge_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.6 64-bit
# name: python3
# ---
from data import current_path_file
import os
import pandas as pd
df = pd.read_csv(os.path.join(current_path_file,'my_data.csv'))
# +
import json
with open(os.path.join(current_path_file,'my_data.json'),'r') as json_file:
json_data = json.load(json_file)
df_json = pd.DataFrame(json_data)
# -
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyomo.environ as pe
import numpy as np
import pandas as pd
m = pe.ConcreteModel()
m.a = pe.Set(initialize=[1, 2, 3])
m.x = pe.Var(m.a, initialize=0, bounds=(-10,10))
m.y = pe.Var(m.a)
def c_rule(m, i):
return m.x[i] >= m.y[i]
m.c = pe.Constraint(m.a, rule=c_rule)
m.c.pprint()
m.z = pe.Var()
m.c2 = pe.Constraint(pe.Any)
m.c2[1] = m.x[1] == 5 * m.z
m.c2[8] = m.x[2] == m.z * m.y[2]
m.c2.pprint()
# Initialize sets and components
m.A = pe.Set(initialize=[2,3,5])
m.B = pe.Set(initialize=set([2,3,5]))
m.C = pe.Set(initialize=(2,3,5))
m.A.pprint()
# using generators for inititalization
m.D = pe.Set(initialize=range(9))
m.E = pe.Set(initialize=(i for i in m.B if i%2 == 0))
m.E.pprint()
# using numpy
f = np.array([2, 3, 5])
m.F = pe.Set(initialize=f)
# using functions for initialization
def g(model):
return [2,3,5]
m.G = pe.Set(initialize=g)
m.G.pprint()
# +
# using indexed sets
H_init = {}
H_init[2] = [1,3,5]
H_init[3] = [2,4,6]
H_init[4] = [3,5,7]
m.H = pe.Set([2,3,4],initialize=H_init)
# -
# parameters
m.a = pe.Param(initialize=1.1)
m.b = pe.Param([1,2,3], initialize={1:1, 2:2, 3:3})
def c(model):
return {1:1, 2:2, 3:3}
m.c = pe.Param([1,2,3], initialize=c)
# The following commands are used to declare data:
# - The set command declares set data.
# - The param command declares a table of parameter data, which can also include the declaration of the set data used to index the parameter data.
# - The table command declares a two-dimensional table of parameter data.
# - The load command defines how set and parameter data is loaded from external data sources, including ASCII table files, CSV files, XML files, YAML files, JSON files, ranges in spreadsheets, and database tables.
#
# The following commands are also used in data command files:
# - The include command specifies a data command file that is processed immediately.
# - The data and end commands do not perform any actions, but they provide compatibility with AMPL scripts that define data commands.
# - The namespace keyword allows data commands to be organized into named groups that can be enabled or disabled during model construction.
#
# The following data types can be represented in a data command file:
# - Numeric value: Any Python numeric value (e.g. integer, float, scientific notation, or boolean).
# - Simple string: A sequence of alpha-numeric characters.
# - Quoted string: A simple string that is included in a pair of single or double quotes. A quoted string can include quotes within the quoted string
# +
# sodacan-pyomo.py
from pyomo.environ import *
from math import pi
m = ConcreteModel()
m.r = Var(bounds=(0,None))
m.h = Var(bounds=(0,None))
m.o = Objective(expr=2*pi*m.r*(m.r + m.h))
m.c = Constraint(expr=pi*m.h*m.r**2 == 355)
solver = SolverFactory('ipopt')
status = solver.solve(m)
print("Status = %s" % status.solver.termination_condition)
print("%s = %f" % (m.r, value(m.r)))
print("%s = %f" % (m.h, value(m.h)))
print("Objective = %f" % value(m.o))
# -
# ## Telecom Tower Problem
# +
from pyomo.environ import *
from math import pi
m = ConcreteModel()
m.x = Var(initialize=1e-10)#(bounds=(1e-10, None))
m.y = Var(initialize=1e-10)#(bounds=(1e-10, None))
m.a = Param([1,2,3,4], initialize={1:0, 2:-110, 3:-130, 4:-250})
m.b = Param([1,2,3,4], initialize={1:0, 2:60, 3:20, 4:100})
m.c1 = Constraint(expr= ((m.x - m.a[1])**2 + (m.y - m.b[1])**2)**0.5 <= 160)
m.c2 = Constraint(expr= ((m.x - m.a[2])**2 + (m.y - m.b[2])**2)**0.5 <= 160)
m.c3 = Constraint(expr= ((m.x - m.a[3])**2 + (m.y - m.b[3])**2)**0.5 <= 160)
m.c4 = Constraint(expr= ((m.x - m.a[4])**2 + (m.y - m.b[4])**2)**0.5 <= 160)
m.o = Objective(expr=
((m.x - m.a[1])**2 + (m.y - m.b[1])**2)**0.5 + \
((m.x - m.a[2])**2 + (m.y - m.b[2])**2)**0.5 + \
((m.x - m.a[3])**2 + (m.y - m.b[3])**2)**0.5 + \
((m.x - m.a[4])**2 + (m.y - m.b[4])**2)**0.5 , sense=minimize)
solver = SolverFactory('ipopt')
status = solver.solve(m)
print("Status = %s" % status.solver.termination_condition)
print("%s = %f" % (m.x, value(m.x)))
print("%s = %f" % (m.y, value(m.y)))
print("Objective = %f" % value(m.o))
# -
m.a[2]
(250**2 + 0**2)**0.5
| optimization/NLP-pyomo-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Refresher
#
# From http://scipy-lectures.org:
#
# Python is:
#
# - an **interpreted** (as opposed to compiled) language. Contrary to e.g. C or Fortran, one does not compile Python code before executing it. In addition, Python can be used interactively: many Python interpreters are available, from which commands and scripts can be executed.
# - **free software** released under an open-source license: Python can be used and distributed free of charge, even for building commercial software.
# - **multi-platform**: Python is available for all major operating systems, Windows, Linux/Unix, MacOS X, most likely your mobile phone OS, etc.
# - a very **readable language** with clear, non-verbose syntax (that maps closely to English!)
# - a language for which a **large variety of high-quality packages** are available for various applications, from web frameworks to scientific computing.
# - a language very **easy to interface** with other languages, in particular C and C++.
#
# In this lesson we will refresh your memory, the code cells have instructions. We will work through this together.
#
# **Ask questions freely; stop me at any time for explanations!**
# +
# Assign a to the string 'hello world' and print it.
#Now print only the second word 'world'
# -
greeting = 'hello world'
print(greeting)
greeting
greeting[6:]
greeting.split()[-1]
# +
# Assign 7 to an integer n, and 3.1415 to a float x.
# Print both of them. Print their sum. What type is the result?
# -
n = 7
x = 3.1415
import math
x = math.pi
x
# ### Containers
#
# There are three basic containers in python: tuples created with `()`, lists created with `[]`, and dictionaries (key-value stores) created with `{}`. There are more: `sets` and variations on these basic ones.
#
# Note that the notebook is linear, values created in a previous cell can be accessed in subsequent ones.
# +
import math
a = 'hello world'
n = 7
x = math.pi
# +
# create a tuple `f` that holds a, n, x. Print the first item from the tuple.
# create a list `e' that holds a, n, x. Print the first item from the list.
# create a dictionary `d` that has keys `a`, `n`, `x` and values `a`, `n`, `x` respectively.
# Print the value for the key `a`
# create two sets, g and h, that contains some numbers; find their intersection
# -
f = (a, n, x)
f
e = [a, n, x]
e
d = {'hello': 'a greeting in the English language', 1: 'one'}
d
d = {'a': a, 'n': n, 'x': x}
d
d['a']
set([1, 2, 3])
set0 = {1, 2, 3}
set1 = {1, 5, 7}
print(set0, set1)
set0.intersection(set1)
[1, 2, 3] + [4, 5, 6]
# ### Mutability
#
# Strings, ints, floats in Python are immutable, they cannot be changed in-place.
# What about lists, dictionaries, tuples?
# +
# Apend `f` to `e`, and add both `e` and `f` to `d`
# Can you append `e` to `f`?
# -
f
e
e + e
x = [e, f]
x
e[0] = 'goodbye world'
x
x = [d, e]
x
d['a'] = 'another value of a'
x
d
d1 = {'hello': 'world'}
d2 = {'hi': 'there', 4: 5}
print(d1, d2)
d1.update(d2)
print(d1)
{ name: value for (name, value) in ... }
# +
d0 = {1: 3}
d1 = {1: 4}
d0.update(d1)
print(d0)
# +
# Add a to a, print it. Do the same for f and e. Can you add `d` to `d`? Why or why not?
# -
# Assign 'e' to a new variable `j`. Now change `e`, set its second value to `f`.
# What happens to `g`?
# +
# Go back to the 4th cell and re-evaluate it.
# Print `e` and `j`. What happened?
# -
# ### Slices and Operators
#
# Indexing in python starts at ... ? Slices take the form `[start:stop:step]` where `stop` and `step` can be implied. `start`, `stop` can be negative, which then starts from the end.
# +
# create r as the concatenation of `e`, `e`, `e`, `e`. Can you do it without `+`?
# print `r` reversed using indexing
# print the last 5 values of `r`
# +
# create r as the integers from 0 to 9. Print the sum.
# -
lst = [1, 2, 3, 'a', 'b', 'c']
lst
lst[::-1]
for i in range(6):
print(i)
list(range(6))
lst
lst[2:4]
# ### Functions
# +
import math
def hypotenuse(a, b=1):
"""Calculate the hypotenuse of a right triangle.
Parameters
----------
a : float
Length of first edge.
b : float, optional
Length of second edge.
Returns
-------
h : float
Hypotenuse of the triangle.
"""
return math.sqrt(a**2 + b**2)
hypotenuse(1)
# -
# #### Generators
def square_range(n):
for i in range(n):
yield i**2
# ### Getting Help
#
# google, [the docs](https://www.python.org/doc/), [language reference](https://docs.python.org/3/), [stackoverflow](https://stackoverflow.com/)
range()
# +
# type `rang` and then hit tab
# type SHIFT-tab to get a docstring
# type range? and then execute the cell (SHIFT or CTRL-enter)
# -
# ---
#
# ### Exercise: Programming syntax
#
# Blocks in Python are denoted by indentation. In order to start a block, the previous line ends with `:`. Used for function definitions, if/else, for/while, exception handling.
#
# Remember the function syntax:
#
# +
# write a loop that sums the even numbers of `r`. Use `for` and `if`
# +
# write a function `sum_even` that accepts a list and returns a list of the sum of the even numbers
# +
# write a function `cumsum` that changes a list in-place to the sum of the numbers before it and returns it
# +
# create a new variable p, and assign to it a list of the numbers from 1 to 10
# call `cumsum` on `p`
# now look at the value of `p`'s value
# what does this tell us about the way that values get passed to Python functions?
# -
# Write a function `divmod(a, b)` that returns both the `div` and the `mod` of `a` by `b`. Now call the function on `a=5` and `b=2` and check that the results are what you'd expect.
# ---
# ### Exceptions
#
# Exceptions take the form of
# ```
# try:
# # here is what you want to happen
# except Exception as e:
# # here is where you end up if something goes wrong
# ```
# +
# rewrite cumsum to raise an exception (`ValueError`) if there is a non-number in the input
# -
# ### List comprension
#
# Python is an interpreted language, which is slow, but it has programming constructs to speed things up and make them look nicer. One very commonly used idiom is `[x for x in lst if has_property(x)]
#
# 1. Square the values in `range(15)` using list comprehension.
# 2. Use list comprehension to find all the prime numbers under 500, given the following function to identify primes.
def is_prime(n):
""""pre-condition: n is a nonnegative integer
post-condition: return True if n is prime and False otherwise.
See https://stackoverflow.com/a/15285590/214686
"""
if n < 2:
return False;
if n % 2 == 0:
return n == 2 # return False
k = 3
while k*k <= n:
if n % k == 0:
return False
k += 2
return True
# ### modules and libraries
#
# Useful functions can be put into a text file with the suffix `.py` and loaded into python via the keyword `import`.
# +
# import the math module. What does it contain?
# +
# now type math. and hit tab
# -
# The `.` keeps the `math` functions inside the `math` *namespace*. Namespaces are great, embrace them. Don't ignore or try to work around them (*never* use `from math import *`).
# use list comprehension to calculate a list of natural logs of the numbers in r
# ### Enumerate
animals = ['cat', 'dog', 'elephant']
for idx, item in enumerate(animals):
print(idx, item)
# ### zip
z = zip([1, 2, 3], ['one', 'two', 'three'])
list(z)
# ### Write your own Python module
#
# Write a Python module, `calc.py`, that defines `add`, `sub`, `mul`, `div` for `+`, `-`, `*`, `/`. Import the module here, and use its functions to calculate (5 + 10) / (3 * 7).
#
# [Note: if you modify your `calc.py` file, you may need to do `%reload calc` or restart the kernel (press 00) to reload the new file. Note that the Jupyter notebook has a "Run all above" option in the Cell menu.]
# ### Tests
#
# Because python is so dynamic, tests are critical to keep your software healthy. Unfortunately, the Notebook framework does not really encourage tests, but we can write some tests.
#
# A test is typically of the form:
#
# ```python
# def test_my_func():
# x = ...
# y = ...
#
# assert my_func(x, y) == 3 # some assertion
# ```
#
# - Write a function, `test_cumsum` that checks `cumsum`. Start by verifying that it turns `[2, 5, 10]` into `[2, 7, 17]`.
#
# - Now, try and think of some corner cases, and test for those as well.
#
# - Test to make sure errors get raised under the right conditions.
# ### For more info about tests
#
# - Builtin framework for tests - [unittest](https://docs.python.org/3/library/unittest.html)
# - [Pytest](https://docs.pytest.org/en/latest/assert.html) - **use this instead**
# Let's run the NumPy test suite. Notice how many tests there are? You may stop the tests with "Kernel -> Interrupt".
np.__file__
# !pytest /home/stefan/envs/py36/lib/python3.6/site-packages/numpy
| notebooks/python_refresher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Imports for the excercise
import pandas as pd
import logging
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn import linear_model
from arcus.azureml.environment.aml_environment import AzureMLEnvironment
# -
# ## Connecting to a remote AzureML environment
azure_config_file = '../.azureml/config.json'
work_env = AzureMLEnvironment.Create(datastore_path='arcus_partition_test', config_file=azure_config_file)
# ## Loading a partitioned dataframe
partition_df = work_env.load_tabular_partition('test-partitioning/stock_AT*', columns=['Close', 'High', 'Isin', 'ItemDate', 'Low', 'Market', 'Open', 'Ticker', 'Volume'])
partition_df.tail(5)
df = work_env.load_tabular_dataset('smart-devops-changesets')
df.head(5)
# ## Starting an experiment with runs and logging to it
trainer = work_env.start_experiment('arcus-demo')
idx = 0
# +
for c_value in [1, 10, 100, 1000]:
print('Classifier with C=', c_value)
idx += 1
_run = trainer.new_run('C=' + str(c_value))
# trainen van de logistic regression classifier
logreg = linear_model.LogisticRegression(C=c_value,solver='liblinear')
logreg.fit(X_train, y_train)
trainer.evaluate_classifier(logreg, X_test, y_test, show_roc = True, upload_model = True)
# -
# ## Build simple classification model
df = pd.read_csv('../tests/resources/datasets/student-admission.csv')
y = df.Admission.values
X = np.asarray(df.drop(['Admission'],axis=1))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
df.tail(5)
| samples/interactive_experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Tensors
#
# * constant tensor `tf.constant()`
# * Value of tensor never changes, hence *constant*.
# * `tf.constant(1234)` is a 0-dimensional int32 tensor
# * `tf.constant([1,2,3,4])` is a 4-dimensional int32 tensor
#
# Sample code:
# +
import tensorflow as tf
# Create TensorFlow object called tensor
hello_constant = tf.constant('Hello World!')
# -
# ## Session
# * An environment for running a graph. In charge of allocating the operations to GPU(s) and/or CPU(s).
#
# Continuing our example:
with tf.Session() as sess:
output = sess.run(hello_constant)
print(output)
# ## Input
#
# * `tf.placeholder()`: returns a tensor that gets it’s value from data passed to the `tf.session.run()` function, allowing you to set the input right before the session runs.
# * `feed_dict`: Use the feed_dict parameter in tf.session.run() to set the placeholder tensor.
#
# Example:
#
a = tf.placeholder(tf.string)
b = tf.placeholder(tf.int32)
c = tf.placeholder(tf.float32)
with tf.Session() as sess:
output = sess.run(a, feed_dict={a: 'hi', b: 23, c: 32.0})
print(output)
# It also works if you feed it only `{a: 'hi'}`, i.e. the relevant placeholder value(s).
#
#
# ## Maths
# +
# Add, subtract, multiply and divide operations
add = tf.add(5, 2) # 7
sub = tf.sub(10, 4) # 6
mul = tf.mul(2, 5) # 10
div = tf.div(10, 5) # 2
with tf.Session() as sess:
output = [sess.run(add), sess.run(sub), sess.run(mul),
sess.run(div)]
print(output)
# -
# [TF Math documentation](https://www.tensorflow.org/versions/r0.11/api_docs/python/math_ops.html)
#
# ## Variables
#
# * `tf.Variable()` function creates a tensor with an initial value that can be modified later, much like a normal Python variable. This tensor stores it’s state in the session, so you must use the `tf.initialize_all_variables()` function to initialize the state of the tensor.
#
# +
# Initialisation
def variables():
output = None
x = tf.Variable([1, 2, 3, 4])
# Initialise all variables
init = tf.initialize_all_variables()
with tf.Session() as sess:
sess.run(init)
output = sess.run(x)
return output
variables()
# +
# Logistic Regression
def logits():
output = None
x_data = [[1.0, 2.0], [2.5, 6.3]]
test_weights = [[-0.3545495, -0.17928936], [-0.63093454, 0.74906588]]
class_size = 2
x = tf.placeholder(tf.float32)
weights = tf.Variable(test_weights)
biases = tf.Variable(tf.zeros([class_size]))
# ToDo: Implement wx + b in TensorFlow
logits = tf.matmul(weights, x)
init = tf.initialize_all_variables()
with tf.Session() as sess:
sess.run(init)
output = sess.run(logits, feed_dict={x: x_data})
return output
logits()
# -
# ## Softmax
#
# Turns logits into probabilities that sum to 1.
# * `tf.nn.softmax()`.
#
# Example of how it works:
#
# ```
# # logits is a one-dimensional array with 3 elements
# logits = [1.0, 2.0, 3.0]
# # softmax will return a one-dimensional array with 3 elements
# print softmax(logits)
#
# [ 0.09003057 0.24472847 0.66524096]
#
# # logits is a two-dimensional array
# logits = np.array([
# [1, 2, 3, 6],
# [2, 4, 5, 6],
# [3, 8, 7, 6]])
# # softmax will return a two-dimensional array with the same shape
# print softmax(logits)
#
#
# [
# [ 0.09003057 0.00242826 0.01587624 0.33333333]
# [ 0.24472847 0.01794253 0.11731043 0.33333333]
# [ 0.66524096 0.97962921 0.86681333 0.33333333]
# ]
# ```
# +
# Softmax function in ram Python
import numpy as np
def softmax(x):
"""Compute softmax values for each sets of scores in x."""
# TODO: Compute and return softmax(x)
# S(y_i) = (e**(y_i) / sum_over_j(e**y_j))
return np.exp(x) / np.sum(np.exp(x), axis=0)
# -
# That's some elegant Numpy code.
# +
# Softmax with TF
import tensorflow as tf
def run():
output = None
logit_data = [2.0, 1.0, 0.1]
logits = tf.placeholder(tf.float32)
# ToDo: Calculate the softmax of the logits
softmax = tf.nn.softmax(logits)
with tf.Session() as sess:
# ToDo: Feed in the logits data
output = sess.run(softmax, feed_dict={logits: logit_data})
return output
# -
# Scaling and Softmax
# * When you divide all the logits by e.g. 10, the probabilities get closer to the uniform distribution.
# * When you multiply all the logits by e.g. 10, the probabilities get closer to 0.0 or 1.0.
#
# ## One-Hot Encodings
# * Vectors with one 1.0 and 0.0 everywhere else.
#
# ## ReLUs: f(x) = max(0,x)
# *Adding nonlinearities*
#
# A Rectified linear unit (ReLU) is type of **activation function** that is defined as `f(x) = max(0, x)`. The function returns 0 if `x` is negative, otherwise it returns `x`. TensorFlow provides the ReLU function as `tf.nn.relu()`, as shown below.
#
# 
# +
# Hidden Layer with ReLU activation function
hidden_layer = tf.add(tf.matmul(features, weights), biases)
hidden_layer = tf.nn.relu(hidden_layer)
output = tf.add(tf.matmul(hidden_layer, weights), biases)
# -
# The above code applies the `tf.nn.relu()` function to the `hidden_layer`, effectively turning off any negative weights and acting like an on/off switch. Adding additional layers, like the output layer, after an activation function turns the model into a nonlinear function. This nonlinearity allows the network to solve more complex problems.
#
#
# It's interesting how you just add `hidden_layer=tf.nn.relu(hidden_layer)`.
# Solution is available in the other "solution.py" tab
def run():
output = None
hidden_layer_weighats = [
[0.1, 0.2, 0.4],
[0.4, 0.6, 0.6],
[0.5, 0.9, 0.1],
[0.8, 0.2, 0.8]]
out_weights = [
[0.1, 0.6],
[0.2, 0.1],
[0.7, 0.9]]
# Weights and biases
weights = [
tf.Variable(hidden_layer_weights),
tf.Variable(out_weights)]
biases = [
tf.Variable(tf.zeros(3)),
tf.Variable(tf.zeros(2))]
# Input
features = tf.Variable([[1.0, 2.0, 3.0, 4.0], [-1.0, -2.0, -3.0, -4.0], [11.0, 12.0, 13.0, 14.0]])
# Model
hidden_layer = tf.matmul(features, weights[0]) + biases[0]
# ToDo: Apply activation using a single Relu
hidden_layer = tf.nn.relu(hidden_layer)
logits = tf.matmul(hidden_layer, weights[1]) + biases[1]
# Calculate logits
with tf.Session() as sess:
sess.run(tf.initialize_all_variables())
output = sess.run(logits)
return output
# ## DNN in Tensorflow
import tensorflow as tf
# ### Import data
help(input_data.read_data_sets)
mnist
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets(".", one_hot=True)
# Udacity version included reshape=False but this got the
# 'unexpected keyword' error
# ### Learning parameters
# +
# Learning Parameters
## Usually we have to find these.
learning_rate = 0.001
training_epochs = 15
batch_size = 100
display_step = 1
n_input = 784 # MNIST data input (img shape: 28*28)
n_classes = 10 # MNIST total classes (0-9 digits)
# -
# ### Hidden layer width
n_hidden_layer = 256 # layer number of features (width of a layer)
# ### Weights and biases
# Store layers weight & bias
weights = {
'hidden_layer': tf.Variable(tf.random_normal([n_input, n_hidden_layer])),
'out': tf.Variable(tf.random_normal([n_hidden_layer, n_classes]))
}
biases = {
'hidden_layer': tf.Variable(tf.random_normal([n_hidden_layer])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
# ### Input
# tf Graph input
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
x_flat = x
tf.shape(x)
x_flat2 = tf.reshape(x, [-1, n_input])
tf.shape(x_flat2)
# + active=""
# # tf Graph input
# x = tf.placeholder("float", [None, 784, 1])
# y = tf.placeholder("float", [None, n_classes])
# x_flat = x
# + active=""
# # tf Graph input
# x = tf.placeholder("float", [None, 28, 28, 1])
# y = tf.placeholder("float", [None, n_classes])
#
# # The MNIST data is made up of 28px by 28px images with a single channel.
# # `tf.reshape` reshapes a batch of 28*28 pixels, x, to a batch of 784 pixels.
# x_flat = tf.reshape(x, [-1, n_input])
# -
# ### Multilayer Perceptron
# Hidden layer with RELU activation
layer_1 = tf.add(tf.matmul(x_flat, weights['hidden_layer']), biases['hidden_layer'])
layer_1 = tf.nn.relu(layer_1)
# Output layer with linear activation
logits = tf.matmul(layer_1, weights['out']) + biases['out']
# *So we're putting RELUs in between layers with weights and biases in them to allow for more complexity. Here we have two layers sandwiching a ReLU.*
# Define loss and optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits, y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate).minimize(cost)
# +
# Initializing the variables
init = tf.initialize_all_variables()
# Launch the graph
with tf.Session() as sess:
sess.run(init)
# Training cycle
for epoch in range(training_epochs):
total_batch = int(mnist.train.num_examples/batch_size)
# Loop over all batches
for i in range(total_batch):
batch_x, batch_y = mnist.train.next_batch(batch_size)
# Run optimization op (backprop) and cost op (to get loss value)
sess.run(optimizer, feed_dict={x: batch_x, y: batch_y})
# +
'''
A Multilayer Perceptron implementation example using TensorFlow library.
This example is using the MNIST database of handwritten digits
(http://yann.lecun.com/exdb/mnist/)
Author: <NAME>
Project: https://github.com/aymericdamien/TensorFlow-Examples/
'''
from __future__ import print_function
# Import MNIST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
import tensorflow as tf
# Parameters
learning_rate = 0.001
training_epochs = 15
batch_size = 100
display_step = 1
# Network Parameters
n_hidden_1 = 256 # 1st layer number of features
n_hidden_2 = 256 # 2nd layer number of features
n_input = 784 # MNIST data input (img shape: 28*28)
n_classes = 10 # MNIST total classes (0-9 digits)
# tf Graph input
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
# Create model
def multilayer_perceptron(x, weights, biases):
# Hidden layer with RELU activation
layer_1 = tf.add(tf.matmul(x, weights['h1']), biases['b1'])
layer_1 = tf.nn.relu(layer_1)
# Hidden layer with RELU activation
layer_2 = tf.add(tf.matmul(layer_1, weights['h2']), biases['b2'])
layer_2 = tf.nn.relu(layer_2)
# Output layer with linear activation
out_layer = tf.matmul(layer_2, weights['out']) + biases['out']
return out_layer
# Store layers weight & bias
weights = {
'h1': tf.Variable(tf.random_normal([n_input, n_hidden_1])),
'h2': tf.Variable(tf.random_normal([n_hidden_1, n_hidden_2])),
'out': tf.Variable(tf.random_normal([n_hidden_2, n_classes]))
}
biases = {
'b1': tf.Variable(tf.random_normal([n_hidden_1])),
'b2': tf.Variable(tf.random_normal([n_hidden_2])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
# Construct model
pred = multilayer_perceptron(x, weights, biases)
# Define loss and optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(pred, y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
# Initializing the variables
init = tf.initialize_all_variables()
# Launch the graph
with tf.Session() as sess:
sess.run(init)
# Training cycle
for epoch in range(training_epochs):
avg_cost = 0.
total_batch = int(mnist.train.num_examples/batch_size)
# Loop over all batches
for i in range(total_batch):
batch_x, batch_y = mnist.train.next_batch(batch_size)
# Run optimization op (backprop) and cost op (to get loss value)
_, c = sess.run([optimizer, cost], feed_dict={x: batch_x,
y: batch_y})
# Compute average loss
avg_cost += c / total_batch
# Display logs per epoch step
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1), "cost=", \
"{:.9f}".format(avg_cost))
print("Optimization Finished!")
# Test model
correct_prediction = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
# Calculate accuracy
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print("Accuracy:", accuracy.eval({x: mnist.test.images, y: mnist.test.labels}))
# -
# ## Dropout
#
# Dropout is a regularization technique for reducing overfitting. The technique temporarily drops units (Artificial Neurons) from the network, along with all its incoming and outgoing connections as shown in Figure 1.
# *Presumably it then drops those units if it obtains better results when dropping those units.*
#
# `tf.nn.dropout()`
#
# +
keep_prob = tf.placeholder(tf.float32) # probability to keep units
hidden_layer = tf.add(tf.matmul(features, weights[0]), biases[0])
hidden_layer = tf.nn.relu(hidden_layer)
hidden_layer = tf.nn.dropout(hidden_layer, keep_prob)
logits = tf.add(tf.matmul(hidden_layer, weights[1]), biases[1])
# -
# The first parameter, hidden_layer, is the tensor that is regularized using dropout.
#
# The second parameter, keep_prob, is the probability of keeping (i.e. not dropping) any given unit.
#
# keep_prob allows you to adjust the number of units to drop. In order to compensate for dropped units, tf.nn.dropout() multiplies all units that are kept (i.e. not dropped) by 1/keep_prob.
#
# During training, a good starting value for keep_prob is 0.5.
#
# During testing, use a keep_prob value of 1.0 to keep all units and maximize the power of the model.
# Added:
# * `hidden_layer = tf.nn.dropout(hidden_layer, keep_prob)`
# and
# * `feed_dict` portion of
# `output = sess.run(logits, feed_dict={keep_prob: 0.5})`
# +
# Solution is available in the other "solution.py" tab
import tensorflow as tf
def run():
output = None
hidden_layer_weights = [
[0.1, 0.2, 0.4],
[0.4, 0.6, 0.6],
[0.5, 0.9, 0.1],
[0.8, 0.2, 0.8]]
out_weights = [
[0.1, 0.6],
[0.2, 0.1],
[0.7, 0.9]]
# Weights and biases
weights = [
tf.Variable(hidden_layer_weights),
tf.Variable(out_weights)]
biases = [
tf.Variable(tf.zeros(3)),
tf.Variable(tf.zeros(2))]
keep_prob = tf.placeholder(tf.float32)
# Input
features = tf.Variable([[0.0, 2.0, 3.0, 4.0], [0.1, 0.2, 0.3, 0.4], [11.0, 12.0, 13.0, 14.0]])
# Model
hidden_layer = tf.matmul(features, weights[0]) + biases[0]
hidden_layer = tf.nn.relu(hidden_layer)
# TODO: Add dropout
hidden_layer = tf.nn.dropout(hidden_layer, keep_prob)
logits = tf.matmul(hidden_layer, weights[1]) + biases[1]
# Calculate logits
with tf.Session() as sess:
sess.run(tf.initialize_all_variables())
output = sess.run(logits, feed_dict={keep_prob: 0.5})
return output
# -
# ## Convolution layer
# * `tf.nn.conv2d()`
# * Computes convolution. TensorFlow uses a stride for each input dimension, [batch, input_height, input_width, input_channels].
# * `tf.nn.bias_add()`
# * adds a 1-d bias to the last dimension in a matrix.
#
#
#
# You'll focus on changing input_height and input_width while setting batch and input_channels to 1. The input_height and input_width strides are for striding the filter over input. In the example code, I'm using a stride of 2 with 5x5 filter over input.
#
#
# +
# Output depth
k_output = 64
# Image Properties
image_width = 10
image_height = 10
color_channels = 3
# Convolution filter
filter_size_width = 5
filter_size_height = 5
# Input/Image
input = tf.placeholder(
tf.float32,
shape=[None, image_width, image_height, color_channels])
# Weight and bias
weight = tf.Variable(tf.truncated_normal(
[filter_size_width, filter_size_height, color_channels, k_output]))
bias = tf.Variable(tf.zeros(k_output))
# Apply Convolution
conv_layer = tf.nn.conv2d(input, weight, strides=[1, 2, 2, 1], padding='SAME')
# Add bias
conv_layer = tf.nn.bias_add(conv_layer, bias)
# Apply activation function
conv_layer = tf.nn.relu(conv_layer)
# -
# ## Max Pooling
#
# `tf.nn.max_pool()`
#
# The image above is an example of max pooling with a 2x2 filter and stride of 2. The four 2x2 colors represent each time the filter was applied to find the maximum value.
#
# * **Benefits of max pooling**: reduces the size of the input, and allow the neural network to focus on only the most important elements.
# * **Method**: Max pooling does this by only retaining the maximum value for each filtered area, and removing the remaining values.
#
#
conv_layer = tf.nn.conv2d(input, weight, strides=[1, 2, 2, 1], padding='SAME')
conv_layer = tf.nn.bias_add(conv_layer, bias)
conv_layer = tf.nn.relu(conv_layer)
# Apply Max Pooling
conv_layer = tf.nn.max_pool(
conv_layer,
ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1],
padding='SAME')
# The tf.nn.max_pool() function performs max pooling with the ksize parameter as the size of the filter and the strides parameter as the length of the stride. 2x2 filters with a stride of 2x2 are common in practice.
#
# The ksize and strides parameters are structured as 4-element lists, with each element corresponding to a dimension of the input tensor ([batch, height, width, channels]). For both ksize and strides, the batch and channel dimensions are typically set to 1.
| Tensorflow/TensorFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Battle Of Neighborhoods
# ### Introduction:
# New York City's demographics show that it is a large and ethnically diverse metropolis. It is the largest city in the United States with a long history of international immigration. New York City was home to nearly 8.5 million people in 2014, accounting for over 40% of the population of New York State and a slightly lower percentage of the New York metropolitan area, home to approximately 23.6 million. Over the last decade the city has been growing faster than the region. The New York region continues to be by far the leading metropolitan gateway for legal immigrants admitted into the United States.
#
# New York City has also been a major point of entry for immigrants; the term "melting pot" was coined to describe densely populated immigrant neighbourhoods on the Lower East Side. As many as 800 languages are spoken in New York, making it the most linguistically diverse city in the world. English remains the most widely spoken language, although there are areas in the outer boroughs in which up to 25% of people speak English as an alternate language, and/or have limited or no English language fluency. English is least spoken in neighbourhoods such as Flushing, Sunset Park, and Corona.
#
# With its diverse culture, comes diverse food items. There are many restaurants in New York City, each belonging to different categories like Chinese, Indian, and French etc.
# ### Problem:
# To find the answers to the following questions:
# - List and visualize all major parts of New York City that has great Indian restaurants.
# - What is best location in New York City for Indian Cuisine?
# - Which areas have potential Indian Restaurant Market?
# - Which all areas lack Indian Restaurants?
# - Which is the best place to stay if you prefer Indian Cuisine?
# ### Data Section:
# New York City's demographics show that it is a large and ethnically diverse metropolis. With its diverse culture, comes diverse food items. There are many restaurants in New York City, each belonging to different categories like Chinese, Indian, and French etc.
#
# For this project we need the following data:
# New York City data that contains list Boroughs, Neighbourhoods along with their latitude and longitude.
# - Data source : https://cocl.us/new_york_dataset
# - Description: This data set contains the required information in JSON format. And we will use this data set to explore various neighbourhoods of New York City.
# - Fields - Feature type, geometry of the feature , coordinates of the feature, name, borough, neighborhood and bounding box coordinates.
# Indian restaurants in each neighbourhood of New York City.
# - Data source : Foursquare API
# - Description : By using this API we will get all the venues in each neighbourhood. We can filter these venues to get only Indian restaurants.
# We have already come across the FourSquare API, so nothing new to explain as we need to link the coordinates data from New York City dataset with the API and get the neighborhood details to explore the influence of Indian restaurants in the vicinity.
# # The Approach
#
# - We begin by collecting the New York city data from "https://cocl.us/new_york_dataset".
# - We will find all venues for each neighborhood using FourSquare API.
# - We will then filter out all Indian Restuarant venues.
# - Next using FourSquare API, we will find the Ratings, Tips, and Like count for all the Indian Resturants.
# - Next we will sort the data keeping Ratings as the constraint.
# - Finally, we will visualize the Ranking of neighborhoods using python's Folium library.
# ### Import Libraries
# +
import pandas as pd
import numpy as np
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
import requests
from bs4 import BeautifulSoup
import os
import geocoder
import folium
from geopy.geocoders import Nominatim
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib.colors as colors
# %matplotlib inline
print('Libraries imported!')
# -
# ### FourSquare API details
radius=1000
LIMIT=100
CLIENT_ID = ''
CLIENT_SECRET = ''
VERSION = '20200401'
# ### Define a function to get the coordinates of a given location using geopy.
def geo_location(address):
'''
To get the coordinates of a given location using geopy
'''
geolocator = Nominatim(user_agent="ny_explorer")
location = geolocator.geocode(address)
latitude = location.latitude
longitude = location.longitude
return latitude,longitude
# ### Define a function to interact with FourSquare API and get top 100 venues within a radius of 1000 metres for a given coordinate. Below function will return us the venue id , venue name and category.
def get_venues(lat,lng):
'''
To interact with FourSquare API and get top 100 venues id, name and category
within a radius of 1000 metres for a given coordinate
'''
#url to fetch data from foursquare api
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}'.format(
CLIENT_ID,
CLIENT_SECRET,
VERSION,
lat,
lng,
radius,
LIMIT)
# get all the data
results = requests.get(url).json()
venue_data=results["response"]['groups'][0]['items']
venue_details=[]
for row in venue_data:
try:
venue_id=row['venue']['id']
venue_name=row['venue']['name']
venue_category=row['venue']['categories'][0]['name']
venue_details.append([venue_id,venue_name,venue_category])
except KeyError:
pass
column_names=['ID','Name','Category']
df = pd.DataFrame(venue_details,columns=column_names)
return df
# ### Define a function to get venue details like like count , rating , tip counts for a given venue id to be used for ranking
def get_venue_details(venue_id):
'''
To get venue details like like count , rating , tip counts for a given venue id
'''
#url to fetch data from foursquare api
url = 'https://api.foursquare.com/v2/venues/{}?&client_id={}&client_secret={}&v={}'.format(
venue_id,
CLIENT_ID,
CLIENT_SECRET,
VERSION)
# get all the data
results = requests.get(url).json()
venue_data=results['response']['venue']
venue_details=[]
try:
venue_id=venue_data['id']
venue_name=venue_data['name']
venue_likes=venue_data['likes']['count']
venue_rating=venue_data['rating']
venue_tips=venue_data['tips']['count']
venue_details.append([venue_id,venue_name,venue_likes,venue_rating,venue_tips])
except KeyError:
pass
column_names=['ID','Name','Likes','Rating','Tips']
df = pd.DataFrame(venue_details,columns=column_names)
return df
# ### Define a funtion to get the New York City data such as boroughs, neighborhoods along with their coordinates
def get_new_york_data():
'''
To get the New York City data such as boroughs, neighborhoods along with their coordinates
'''
url='https://cocl.us/new_york_dataset'
resp=requests.get(url).json()
# all data is present in features label
features=resp['features']
# define the dataframe columns
column_names = ['Borough', 'Neighborhood', 'Latitude', 'Longitude']
# instantiate the dataframe
new_york_data = pd.DataFrame(columns=column_names)
for data in features:
borough = data['properties']['borough']
neighborhood_name = data['properties']['name']
neighborhood_latlon = data['geometry']['coordinates']
neighborhood_lat = neighborhood_latlon[1]
neighborhood_lon = neighborhood_latlon[0]
new_york_data = new_york_data.append({'Borough': borough,
'Neighborhood': neighborhood_name,
'Latitude': neighborhood_lat,
'Longitude': neighborhood_lon}, ignore_index=True)
return new_york_data
# #### Call the above funtion to get the new york city data.
new_york_data=get_new_york_data()
new_york_data.head()
new_york_data.shape
# ## There are 306 different Neighborhoods in New York City
# ### Plot to show different neighborhoods in New York City
plt.figure(figsize=(9,5), dpi = 100)
# title
plt.title('Number of Neighborhood for each Borough in New York City')
#On x-axis
plt.xlabel('Borough', fontsize = 15)
#On y-axis
plt.ylabel('No.of Neighborhood', fontsize=15)
#giving a bar plot
new_york_data.groupby('Borough')['Neighborhood'].count().plot(kind='bar')
#legend
plt.legend()
#displays the plot
plt.show()
# ### From the above Bar Plot, we can see that Queens has highest number of neighborhoods.
# ### Lets collect Indian resturants for each neighborhood.
# prepare neighborhood list that contains indian resturants
column_names=['Borough', 'Neighborhood', 'ID','Name']
indian_rest_ny=pd.DataFrame(columns=column_names)
count=1
for row in new_york_data.values.tolist():
Borough, Neighborhood, Latitude, Longitude=row
venues = get_venues(Latitude,Longitude)
indian_resturants=venues[venues['Category']=='Indian Restaurant']
print('(',count,'/',len(new_york_data),')','Indian Resturants in '+Neighborhood+', '+Borough+':'+str(len(indian_resturants)))
for resturant_detail in indian_resturants.values.tolist():
id, name , category=resturant_detail
indian_rest_ny = indian_rest_ny.append({'Borough': Borough,
'Neighborhood': Neighborhood,
'ID': id,
'Name' : name
}, ignore_index=True)
count+=1
indian_rest_ny.head()
indian_rest_ny.shape
# ### From the above result, we see that there are 144 Indian Resturants across New York City.
# ### Create a bar plot to show Number of Indian Resturants for each Borough in New York City.
plt.figure(figsize=(9,5), dpi = 100)
# title
plt.title('Number of Indian Resturants for each Borough in New York City')
#On x-axis
plt.xlabel('Borough', fontsize = 15)
#On y-axis
plt.ylabel('No.of Indian Resturants', fontsize=15)
#giving a bar plot
indian_rest_ny.groupby('Borough')['ID'].count().plot(kind='bar')
#legend
plt.legend()
#displays the plot
plt.show()
# ### From the above Bar Plot, we can see that Queens has highest number of Indian resturants.
plt.figure(figsize=(9,5), dpi = 100)
# title
plt.title('Number of Indian Resturants for each Neighborhood in New York City')
#On x-axis
plt.xlabel('Neighborhood', fontsize = 15)
#On y-axis
plt.ylabel('No.of Indian Resturants', fontsize=15)
#giving a bar plot
indian_rest_ny.groupby('Neighborhood')['ID'].count().nlargest(5).plot(kind='bar')
#legend
plt.legend()
#displays the plot
plt.show()
indian_rest_ny[indian_rest_ny['Neighborhood']=='Floral Park']
indian_rest_ny[indian_rest_ny['Neighborhood']=='Richmond Hill']
# ### We can see that, Floral Park and Richmond Hill in Queens have the highest number of Indian Resturants with a total count of 8 each.
# ### Get the ranking of each resturant for further analysis
# +
# prepare neighborhood list that contains indian resturants
column_names=['Borough', 'Neighborhood', 'ID','Name','Likes','Rating','Tips']
indian_rest_stats_ny=pd.DataFrame(columns=column_names)
count=1
for row in indian_rest_ny.values.tolist():
Borough,Neighborhood,ID,Name=row
try:
venue_details=get_venue_details(ID)
print(venue_details)
id,name,likes,rating,tips=venue_details.values.tolist()[0]
except:
print('No data available for id=',ID)
# we will assign 0 value for these resturants as they may have been
#recently opened or details does not exist in FourSquare Database
id,name,likes,rating,tips=[0]*5
print('(',count,'/',len(indian_rest_ny),')','processed')
indian_rest_stats_ny = indian_rest_stats_ny.append({'Borough': Borough,
'Neighborhood': Neighborhood,
'ID': id,
'Name' : name,
'Likes' : likes,
'Rating' : rating,
'Tips' : tips
}, ignore_index=True)
count+=1
# -
indian_rest_stats_ny.head()
indian_rest_stats_ny.shape
indian_rest_ny.shape
# Now that we got data for all resturants Now lets save this data to a csv sheet. In case we by mistake modify it. As the number of calls to get details for venue are premium call and have limit of 500 per day, we will refer to saved data sheet csv if required.
| Week5/Capstone - The Battle of Neighborhoods - Part 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification
#
# *지도학습* 기법에는 일련의 변수와 알고 있는 라벨 값이 포함된 데이터 세트를 학습하여 *라벨*을 예측합니다.***y***는 예측하려는 레이블을 나타내고 ***X*** 는 모델을 예측하기 위해 사용하는 변수의 벡터를 나타냅니다.
#
# $$ y = f([x_1, x_2, x_3, ...]) $$
#
#
# *분류*는 예측 변수(***X***)를 학습하여 레이블(***y***)을 예측하는 지도 머신 러닝의 한 형태입니다. 가능한 각 클래스에 속하는 관찰 된 케이스의 확률과 적절한 레이블을 예측합니다. 가장 간단한 분류 형식은 이진분류이며, 여기서 레이블은 두 클래스 중 하나를 나타내는 0 또는 1입니다.(예 : "True" 또는 "False"; "내부" 또는 "외부"; "수익성" 또는 "수익성 없음" 등).
# ## 이진 분류
#
# 모델이 두 클래스 중 하나에 속하는 레이블을 예측해야 하는 이진 분류의 예를 살펴 보겠습니다. 이 연습에서는 일부 의료 데이터를 기반으로 환자가 당뇨병 검사를 받아야하는지 여부를 예측하기 위해 이진 분류기를 훈련합니다.
#
# ### 데이터 탐색
#
# 다음 셀을 실행하여 특허 데이터의 CSV 파일을*Pandas*데이터 프레임에 로드 합니다.
#
# >*인용*: 이 연습에 사용 된 당뇨병 데이터 세트는 원래 the National Institute of Diabetes and Digestive and Kidney Diseases에서 수집한 데이터를 기반으로 합니다.
# +
import pandas as pd
# load the training dataset
diabetes = pd.read_csv('data/diabetes.csv')
diabetes.head()
# -
# 이 데이터는 당뇨병 검사를 받은 일부 환자에 대한 진단 정보로 구성된다. 필요한 경우 오른쪽으로 스크롤하고 데이터 세트의 마지막 열(**Diabetic**)에 당뇨병 음성 테스트 환자의 경우 값 *0*, 양성 테스트 환자의 경우 값이 *1*이 포함되어 있습니다. 이것은 예측하기 위해 모델을 훈련시킬 레이블입니다. 대부분의 다른 열 (**Pregnancies**,**PlasmaGlucose**,**DiastolicBloodPressure** 등)은 *당뇨병* 라벨을 예측하는데 사용할 특징입니다.
#
# 전체 변수에서 레이블에서 분리 해보겠습니다. 예측 변수를 **X**, 레이블 **y**라고 합니다.
# + tags=[]
# Separate features and labels
features = ['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']
label = 'Diabetic'
X, y = diabetes[features].values, diabetes[label].values
for n in range(0,4):
print("Patient", str(n+1), "\n Features:",list(X[n]), "\n Label:", y[n])
# -
# 이제 각 레이블 값에 대한 변수들의 분포를 비교해 보겠습니다.
# +
from matplotlib import pyplot as plt
# %matplotlib inline
features = ['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']
for col in features:
diabetes.boxplot(column=col, by='Diabetic', figsize=(6,6))
plt.title(col)
plt.show()
# -
# 일부 특징의 경우 각 라벨 값의 분포에 눈에 띄는 차이가 있습니다. 특히 **Pregnancies**과 **Age**은 당뇨병 환자의 분포가 비당뇨병 환자와 현저하게 다릅니다. 이러한 변수는 환자가 당뇨병인지 여부를 예측하는데 도움이 될 수 있습니다.
#
# ### 데이터 분할
#
# 우리의 데이터 세트에는 레이블에 대해 알려진 값이 포함되어 있으므로 이를 사용하여 분류기를 훈련시켜 기능과 레이블 값 간의 통계적 관계를 찾을 수 있습니다. 하지만 우리 모델이 좋은지 어떻게 알 수 있을까요? 또한 훈련되지 않은 새로운 데이터와 함께 사용할 때 올바르게 예측할 수 있는지 어떻게 알 수 있을까요? 우리는 알려진 레이블 값이 있는 대규모 데이터 세트가 있다는 사실을 활용하고, 일부만 사용하여 모델을 훈련시키고, 훈련된 모델을 테스트하기 위해 일부 데이터를 사용하지 않고 예측된 레이블을 실제 레이블과 비교할 수 있습니다.
#
# Python에서 **scikit-learn** 패키지에는 학습 및 테스트 데이터를 통계적으로 무작위로 분할 할 수 있도록 보장하는 **train_test_split** 함수를 포함하여 머신 러닝 모델을 구축하는데 사용할 수 있는 많은 함수가 포함되어 있습니다. 이를 사용하여 학습을 위해 데이터를 70%로 분할하고 테스트를 위해 30%를 남깁니다.
# + tags=[]
from sklearn.model_selection import train_test_split
# Split data 70%-30% into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
print ('Training cases: %d\nTest cases: %d' % (X_train.shape[0], X_test.shape[0]))
# -
# ### 이진 분류 모델 학습 및 평가
# 이제 학습할 변수(**X_train**)를 학습 레이블(**y_train**)에 맞춰 모델을 학습시킬 준비가 되었습니다. 모델 훈련에 사용할 수있는 다양한 알고리즘이 있습니다. 이 예에서는 분류를 위해 잘 알려진 알고리즘인 *로지스틱 회귀*를 사용합니다. 훈련 변수 및 레이블 외에도 aregularization 매개 변수를 설정 해야합니다. 이는 샘플의 모든 편향에 대응하고 모델을 학습 데이터에 과적합하지 않도록 하여 모델이 잘 일반화되도록하는데 사용됩니다.
#
# > **참고** : 기계 학습 알고리즘의 파라미터는 일반적으로 ashyperparameters라고합니다 (데이터사이언티스트에게 파라미터는 데이터 자체의 값입니다. 하이퍼 파라미터는 데이터에서 외부적으로 정의됩니다!)
# + tags=[]
# Train the model
from sklearn.linear_model import LogisticRegression
# Set regularization rate
reg = 0.01
# train a logistic regression model on the training set
model = LogisticRegression(C=1/reg, solver="liblinear").fit(X_train, y_train)
print (model)
# -
# 이제 훈련 데이터를 사용하여 모델을 훈련 시켰으므로 남겨 놓은 테스트 데이터를 사용하여 얼마나 잘 예측하는지 평가할 수 있습니다. **scikit-learn**은 이 작업을 수행하는 데 도움이 될 수 있습니다. 먼저 모델을 사용하여 테스트 세트의 레이블을 예측하고 예측된 레이블을 실제 레이블과 비교해 보겠습니다.
# + tags=[]
predictions = model.predict(X_test)
print('Predicted labels: ', predictions)
print('Actual labels: ' ,y_test)
# -
# 레이블 배열이 너무 길어서 노트북 출력에 표시 할 수 없으므로 몇 가지 값만 비교할 수 있습니다. 예측 및 실제 라벨을 모두 출력하더라도 모델을 평가하는 합리적인 방법으로 만들기에는 라벨이 너무 많습니다. 다행히 **scikit-learn**에는 몇 가지 트릭이 더 있으며 모델을 평가하는 데 사용할 수있는 몇 가지 메트릭을 제공합니다.
#
# 가장 분명한 것은 예측의 *정확도*를 확인하는 것입니다. 간단히 말해서 모델이 올바르게 예측 한 레이블의 비율은 얼마일까요?
# + tags=[]
from sklearn.metrics import accuracy_score
print('Accuracy: ', accuracy_score(y_test, predictions))
# -
# 정확도는 10 진수 값으로 반환됩니다. 값이 1.0이면 모델이 예측의 100 %를 올바르게 얻었음을 의미합니다. 0.0의 정확도는 쓸모 없습니다!
#
# 정확성은 평가할 수있는 합리적인 지표처럼 보이지만 분류 정확성에서 명확한 결론이라고 도출하는데에는 다른 지표도 필요 합니다. 얼마나 많은 케이스가 올바르게 예측되었는지 측정한 것임을 기억하십시오. 인구의 3%만이 당뇨병이라고 가정합니다. 항상 0을 예측하는 분류기를 만들 수 있으며 97 % 정확하지만 당뇨병 환자를 식별하는데 크게 도움이되지는 않습니다!
#
# 다행히도 모델의 성능에 대해 좀 더 자세히 보여주는 다른 측정 항목이 있습니다. Scikit-Learn에는 원래 정확도 보다 더 많은 통찰력을 제공하는 분류 보고서(classification report)를 만드는 기능이 포함되어 있습니다.
# + tags=[]
from sklearn. metrics import classification_report
print(classification_report(y_test, predictions))
# -
# 분류 보고서에는 각 클래스 (0과 1)에 대한 다음 메트릭이 포함됩니다.
#
# > 헤더 행이 값과 일치하지 않을 수 있습니다.
#
# * *정밀도(Precision)* :이 클래스에 대한 모델의 예측 중 올바른 비율
# * *재현율(Recall)* : 테스트 데이터 세트에 있는 이 클래스의 모든 인스턴스 중에서 모델이 식별한 비율
# * *F1-Score* : 정밀도와 재현율을 모두 고려하는 평균 측정 항목
# * *Support* : 테스트 데이터 세트에 각 클래스의 데이터 개수
#
# 분류 보고서에는 각 클래스의 케이스 수에서 불균형을 허용하는 가중 평균을 포함하여 이러한 메트릭에 대한 평균도 포함됩니다.
#
# 이것은 *이진* 분류 문제이기 때문에 ***1*** 클래스는 *양성*으로 간주되며 정밀도와 재현율이 특히 흥미롭습니다. 실제로 다음 질문에 답합니다.
#
# - 모델이 예측 한 모든 환자 중 실제로 당뇨병이있는 환자는 몇 명입니까?
# - 실제로 당뇨병 환자 중 몇 명을 모델이 식별 했습니까?
#
# scikit-learn에서 **precision_score** 및 **recall_score** 측정 항목을 사용하여 자체적으로 이러한 값을 검색 할 수 있습니다.(기본적으로 이진 분류 모델을 가정)
# + tags=[]
from sklearn.metrics import precision_score, recall_score
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
# -
# 정밀도 및 재현율 메트릭은 다음과 같은 네 가지 가능한 예측 결과에서 파생됩니다.
# * *True Positives* : 예측 라벨과 실제 라벨이 모두 1입니다.
# * *False Positives* : 예측 라벨은 1이지만 실제 라벨은 0입니다.
# * *False Negatives* : 예측 라벨은 0이지만 실제 라벨은 1입니다.
# * *True Negatives* : 예측 라벨과 실제 라벨이 모두 0입니다.
#
# 이러한 측정 항목은 일반적으로 테스트 세트에 대해 표로 작성되며 다음과 같은 형식을 취하는 혼동 행렬로 함께 표시됩니다.
#
# <table style="border: 1px solid black;">
# <tr style="border: 1px solid black;">
# <td style="border: 1px solid black;color: black;" bgcolor="lightgray">TN</td><td style="border: 1px solid black;color: black;" bgcolor="white">FP</td>
# </tr>
# <tr style="border: 1px solid black;">
# <td style="border: 1px solid black;color: black;" bgcolor="white">FN</td><td style="border: 1px solid black;color: black;" bgcolor="lightgray">TP</td>
# </tr>
# </table>
#
# 올바른 (*true*) 예측은 왼쪽 상단에서 오른쪽 하단으로 대각선을 형성합니다. 모델이 좋은 경우이 수치는 *false* 예측보다 훨씬 높아야합니다.
#
# Python에서는 **sklearn.metrics.confusion_matrix** 함수를 사용하여 학습된 분류(classifier)에 대해 다음 값을 찾을 수 있습니다.
# + tags=[]
from sklearn.metrics import confusion_matrix
# Print the confusion matrix
cm = confusion_matrix(y_test, predictions)
print (cm)
# -
# 지금까지 모델의 예측을 1 또는 0 클래스 레이블로 간주했습니다. 사실, 상황은 그것보다 조금 더 복잡합니다. 로지스틱 회귀와 같은 통계적 기계 학습 알고리즘은 *확률* 을 기반으로합니다. 따라서 이진 분류기에 의해 실제로 예측되는 것은 레이블이 참일 확률 (**P(y)**)과
# 레이블이 거짓 일 확률 (1-**P(y)**)입니다. 임계 값 0.5는 예측 된 레이블이 1 (*P(y) > 0.5*)인지 0 (*P(y) <= 0.5*)인지 결정하는 데 사용됩니다. **predict_proba** 방법을 사용하여 각 케이스의 확률 쌍을 볼 수 있습니다.
# + tags=[]
y_scores = model.predict_proba(X_test)
print(y_scores)
# -
# 예측 점수를 1 또는 0으로 결정하는 것은 예측 확률이 비교되는 임계 값에 따라 다릅니다. 임계 값을 변경하면 예측에 영향을 미칩니다. 따라서 혼동 행렬의 메트릭을 변경합니다. 분류자를 평가하는 일반적인 방법은 가능한 임계 값 범위에 대해 *true positive 비율* (재현율)과 *false positive 비율* 을 조사하는 것입니다. 이러한 비율은 가능한 모든 임계 값에 대해 플롯되어 다음과 같이 *ROC (received operator character)차트* 로 알려진 차트를 형성합니다.
# +
from sklearn.metrics import roc_curve
from sklearn.metrics import confusion_matrix
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
# ROC 차트는 0과 1 사이의 다른 임계 값에 대한 참 및 거짓 긍정 비율의 곡선을 보여줍니다. 완벽한 분류자는 왼쪽을 곧장 올라가서 위쪽을 가로지르는 곡선을 가지고 있을 것이다. 차트의 대각선은 50/50 무작위 예측으로 올바르게 예측할 확률을 나타냅니다. 따라서 곡선이 그보다 더 높기를 원합니다(또는 모델이 단순히 추측하는 것보다 낫지 않습니다!).
#
# 곡선 아래 영역 (AUC)은 모델의 전체 성능을 정량화하는 0에서 1 사이의 값입니다. 이 값이 1에 가까울수록 모델이 더 좋습니다. 다시 한 번 scikit-Learn에는이 메트릭을 계산하는 함수가 포함되어 있습니다.
# + tags=[]
from sklearn.metrics import roc_auc_score
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
# -
# ### 파이프라인에서 전처리 수행
#
# 이 경우 ROC 곡선과 AUC는 랜덤 추측보다 모형이 더 잘 수행된다는 것을 나타내며, 이는 데이터에 대한 전처리를 거의 수행하지 않았다는 점을 고려할 때 나쁘지 않습니다.
#
# 실제로는 알고리즘이 모델에 더 쉽게 맞출 수 있도록 데이터에 대한 몇 가지 전처리를 수행하는 것이 일반적입니다. 데이터를 모델링할 수 있도록 하기 위해 수행할 수 있는 광범위한 전처리 방법들은 다음과 같은 몇 가지 일반적인 기법으로 제한됩니다.
#
# - 동일한 스케일에 있도록 수치형 변수를 스케일링합니다. 따라서 값이 큰 형상이 예측에 불균형적으로 영향을 미치는 계수를 생성하지 않습니다.
# - 범주형 변수를 인코딩합니다. 예를 들어 *one-hot encoding* 을 사용하여 가능한 각 범주 값에 대해 개별 이진(참/거짓) 특징을 생성할 수 있습니다.
#
# 이러한 전처리 전환을 적용하기 위해 *파이프라인*이라는 Scikit-Learn 기능을 사용할 것입니다. 이를 통해 알고리즘으로 끝나는 일련의 전처리 단계를 정의할 수 있습니다. 그런 다음 전체 파이프라인을 데이터에 적합시켜 모델이 모든 전처리 단계와 회귀 알고리즘을 캡슐화할 수 있습니다. 모델을 사용하여 새 데이터의 값을 예측하려면 동일한 변환을 적용해야 하기 때문에 이 방법이 유용합니다(교육 데이터에 사용되는 동일한 통계 분포 및 범주 인코딩을 기반으로 함).
#
# >**참고**: *파이프라인*이라는 용어는 기계 학습에서 광범위하게 사용되며, 매우 다른 의미를 갖는 경우가 많습니다! 이러한 맥락에서 Seagate는 Scikit-Learn에서 파이프라인 개체를 참조하는 데 사용되지만 다른 곳에서 다른 의미를 갖는 경우도 있습니다.
# + tags=[]
# Train the model
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.linear_model import LogisticRegression
import numpy as np
# Define preprocessing for numeric columns (normalize them so they're on the same scale)
numeric_features = [0,1,2,3,4,5,6]
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
# Define preprocessing for categorical features (encode the Age column)
categorical_features = [7]
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
# Combine preprocessing steps
preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)])
# Create preprocessing and training pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('logregressor', LogisticRegression(C=1/reg, solver="liblinear"))])
# fit the pipeline to train a logistic regression model on the training set
model = pipeline.fit(X_train, (y_train))
print (model)
# -
# 파이프라인은 전처리 단계와 모델 교육을 캡슐화합니다.
#
# 이 파이프라인에서 학습한 모델을 사용하여 테스트 세트의 레이블을 예측하고 성능 메트릭을 이전에 생성한 기본 모델과 비교해 보겠습니다.
# + tags=[]
# Get predictions from test data
predictions = model.predict(X_test)
y_scores = model.predict_proba(X_test)
# Get evaluation metrics
cm = confusion_matrix(y_test, predictions)
print ('Confusion Matrix:\n',cm, '\n')
print('Accuracy:', accuracy_score(y_test, predictions))
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
# 결과가 조금 더 좋아 보이므로 데이터를 전처리하는 것이 분명히 차이를 만들어 냈습니다.
#
# ### 다른 알고리즘을 사용해 보십시오.
#
# 이제 다른 알고리즘을 시도해 보겠습니다. 이전에는 *선형* 알고리즘인 로지스틱 회귀 알고리즘을 사용했습니다. 시도할 수 있는 분류 알고리즘은 다음과 같습니다.
#
# - **Support Vector Machine algorithms**: 클래스를 구분하는 *hyperplane*을 정의하는 알고리즘
# - **Tree-based algorithms**: 예측에 도달하기 위한 의사결정 트리를 구축하는 알고리즘
# - **Ensemble algorithms** : 복수의 기본 알고리즘의 산출물을 조합하여 일반화 해가며 개선하는 알고리즘
#
# 이번에는 이전과 동일한 전처리 단계를 사용하겠지만, 복수의 랜덤 의사결정 트리의 결과들을 결합하는 *Random Forest* 알고리즘을 사용하여 모델을 학습하겠습니다(자세한 내용은 [Sikit-Learn 설명서](https://scikit-learn.org/stable/modules/ensemble.html#forests-of-randomized-trees) 참조).
# + tags=[]
from sklearn.ensemble import RandomForestClassifier
# Create preprocessing and training pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('logregressor', RandomForestClassifier(n_estimators=100))])
# fit the pipeline to train a random forest model on the training set
model = pipeline.fit(X_train, (y_train))
print (model)
# -
# 새로운 모델의 성능 지표를 살펴보겠습니다.
# + tags=[]
predictions = model.predict(X_test)
y_scores = model.predict_proba(X_test)
cm = confusion_matrix(y_test, predictions)
print ('Confusion Matrix:\n',cm, '\n')
print('Accuracy:', accuracy_score(y_test, predictions))
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
auc = roc_auc_score(y_test,y_scores[:,1])
print('\nAUC: ' + str(auc))
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# -
# 더 좋아 보이네요!
#
# ## 추론을 위한 모델 사용
# 이제 상당히 유용한 학습 모델을 확보했으므로 나중에 새 데이터에 대한 라벨을 예측하는데 사용할 수 있습니다.
# +
import joblib
# Save the model as a pickle file
filename = './models/diabetes_model.pkl'
joblib.dump(model, filename)
# -
# 라벨을 모르는 새로운 데이터가 있는 경우 모델을 로드하여 라벨에 대한 값을 예측할 수 있습니다.
# + tags=[]
# Load the model from the file
model = joblib.load(filename)
# predict on a new sample
# The model accepts an array of feature arrays (so you can predict the classes of multiple patients in a single call)
# We'll create an array with a single array of features, representing one patient
X_new = np.array([[2,180,74,24,21,23.9091702,1.488172308,22]])
print ('New sample: {}'.format(list(X_new[0])))
# Get a prediction
pred = model.predict(X_new)
# The model returns an array of predictions - one for each set of features submitted
# In our case, we only submitted one patient, so our prediction is the first one in the resulting array.
print('Predicted class is {}'.format(pred[0]))
# -
# ## 다중 클래스 분류
#
# 이진 분류 기법은 데이터 관찰이 "참" 또는 "거짓"과 같은 두 가지 클래스 또는 범주 중 하나에 속할 때 잘 작동합니다. 데이터를 세 개 이상의 클래스로 분류할 수 있는 경우 다중 클래스 분류 알고리즘을 사용해야 합니다.
#
# 다중 클래스 분류는 여러 개의 이진 분류자의 조합으로 생각할 수 있습니다. 문제에 접근하는 방법에는 두 가지가 있습니다.
#
# - **One vs Rest (OVR)*** 각 클래스 값에 대해 분류기가 생성되며, 예측이 특정 클래스인 경우 긍정적인 결과를, 예측이 다른 클래스인 경우 부정적인 예측이 생성됩니다. 네 가지 형상 클래스(*제곱*, *원*, *삼각형*, *육각형*)를 사용할 수 있는 분류 문제를 해결하려면 다음을 예측하는 네 개의 classifier(분류자)가 필요합니다.
# - *square* or not
# - *circle* or not
# - *triangle* or not
# - *hexagon* or not
#
# - **One vs One(OVO)*** 각 클래스 쌍에 대한 분류기가 생성됩니다. 네 가지 모양 클래스에서 발생하는 분류 문제에는 다음과 같은 이진 분류자가 필요합니다.
# - *square* or *circle*
# - *square* or *triangle*
# - *square* or *hexagon*
# - *circle* or *triangle*
# - *circle* or *hexagon*
# - *triangle* or *hexagon*
#
# 두 접근 방식 모두에서 분류자를 결합하는 전체 모델은 개별 이진 분류기에서 생성된 확률을 사용하여 예측할 클래스를 결정하는 예측 벡터를 생성합니다.
#
# 다행히 scikit-learn을 포함한 대부분의 기계 학습 프레임 워크에서 다중 클래스 분류 모델을 구현하는 것은 이진 분류보다 훨씬 복잡하지 않습니다. 대부분의 경우 이진 분류에 사용되는 추정기는 OVR 알고리즘 인 OVO를 추상화하여 다중 클래스 분류를 암시적으로 지원합니다. 알고리즘을 사용하거나 둘 중 하나를 선택할 수 있습니다.
#
# > **추가 정보**: Scikit-Learn에서 다중 클래스 분류에 대한 estimator 지원에 대한 자세한 내용은 [Scikit-Learn 설명서](https://scikit-learn.org/stable/modules/multiclass.html)를 참조하십시오.
#
# 여러 클래스의 관찰을 포함하는 데이터 세트를 조사하여 시작하겠습니다. 우리는 서로 다른 세 종의 펭귄에 대한 관찰 데이터 세트를 사용할 것입니다.
#
# > **인용**: 이 연습에 사용된 펭귄 데이터 세트
# [Dr. <NAME>](https://www.uaf.edu/cfos/people/faculty/detail/kristen-gorman.php)
# and the [Palmer Station, Antarctica LTER](https://pal.lternet.edu/), a
# member of the [Long Term Ecological Research
# Network](https://lternet.edu/).
# +
import pandas as pd
# load the training dataset
penguins = pd.read_csv('data/penguins.csv')
# Display a random sample of 10 observations
sample = penguins.sample(10)
sample
# -
# 데이터 세트에는 다음 열로 구성 되어있습니다/
# * **CulmenLength**: 펭귄의 부리 길이.
# * **Culmen Depth**: 펭귄의 부리 깊이(mm).
# * **FlipperLength**: 펭귄 지느러미의 길이(mm).
# * **BodyMass**: 펭귄의 체질량(g 단위).
# * **Species**: 펭귄의 종을 나타내는 정수 값.
#
# * *Species* 열은 예측하기 위해 모델을 교육하려는 레이블입니다. 데이터 세트에는 가능한 세 가지 종(0, 1, 2)이 포함됩니다. 실제 종 이름은 아래 코드로 확인할 수 있습니다.
# + tags=[]
penguin_classes = ['Adelie', 'Gentoo', 'Chinstrap']
print(sample.columns[0:5].values, 'SpeciesName')
for index, row in penguins.sample(10).iterrows():
print('[',row[0], row[1], row[2], row[3], int(row[4]),']',penguin_classes[int(row[4])])
# -
# 이제 데이터의 특징과 레이블이 무엇을 나타내는 지 알았으니 데이터 세트를 살펴 보겠습니다. 먼저 누락된 (*null*) 값이 있는지 살펴 보겠습니다.
# + tags=[]
# Count the number of null values for each column
penguins.isnull().sum()
# -
# 누락된 변수 값은 일부 있지만 레이블은 누락 되지 않았습니다. Null이 포함된 행을 살펴보겠습니다.
# + tags=[]
# Show rows containing nulls
penguins[penguins.isnull().any(axis=1)]
# -
# 변수 값을 전혀 포함하지 않는 행이 두 개 있으므로 (*NaN*은 "숫자가 아님"을 나타냄) 모델 학습에 유용하지 않습니다. 데이터 세트에서 삭제하겠습니다.
# Drop rows containing NaN values
penguins=penguins.dropna()
#Confirm there are now no nulls
penguins.isnull().sum()
# 이제 누락 된 값을 처리 했으므로 상자 그림을 만들어 변수들이 레이블과 어떻게 관련되는지 살펴 보겠습니다.
# +
from matplotlib import pyplot as plt
# %matplotlib inline
penguin_features = ['CulmenLength','CulmenDepth','FlipperLength','BodyMass']
penguin_label = 'Species'
for col in penguin_features:
penguins.boxplot(column=col, by=penguin_label, figsize=(6,6))
plt.title(col)
plt.show()
# -
# 상자 그림에서 종 0과 2 (Amelie 및 Chinstrap)는 부리 깊이, 플리퍼 길이 및 체질량에 대해 유사한 데이터 프로필을 가지고있는 것처럼 보이지만 Chinstrap은 더 긴 부리를 갖는 경향이 있습니다. 1 종 (Gentoo)은 다른 것들과 상당히 분명하게 구별되는 특징을 가지고 있습니다. 좋은 분류 모델을 훈련하는 데 도움이 될 것입니다.
#
# ### 데이터 준비
#
# 이진 분류와 마찬가지로 모델을 훈련하기 전에 기능과 레이블을 분리 한 다음 훈련 및 검증을 위해 데이터를 하위 집합으로 분할해야 합니다. 또한 훈련 및 검증 데이터 세트에서 각 레이블 값의 비율을 유지하기 위해 데이터를 분할 할 때 *stratification(층화)* 기술을 적용합니다.
# + tags=[]
from sklearn.model_selection import train_test_split
# Separate features and labels
penguins_X, penguins_y = penguins[penguin_features].values, penguins[penguin_label].values
# Split data 70%-30% into training set and test set
x_penguin_train, x_penguin_test, y_penguin_train, y_penguin_test = train_test_split(penguins_X, penguins_y,
test_size=0.30,
random_state=0,
stratify=penguins_y)
print ('Training Set: %d, Test Set: %d \n' % (x_penguin_train.shape[0], x_penguin_test.shape[0]))
# -
# ### 다중 클래스 분류 훈련 및 평가
#
# 이제 학습 기능 세트와 해당 학습 레이블이 있으므로 데이터에 다중 클래스 분류 알고리즘을 적용하여 모델을 만들 수 있습니다. 대부분의 scikit-learn 분류 알고리즘은 본질적으로 다중 클래스 분류를 지원합니다. 로지스틱 회귀 알고리즘을 시도해 보겠습니다.
# + tags=[]
from sklearn.linear_model import LogisticRegression
# Set regularization rate
reg = 0.1
# train a logistic regression model on the training set
multi_model = LogisticRegression(C=1/reg, solver='lbfgs', multi_class='auto', max_iter=10000).fit(x_penguin_train, y_penguin_train)
print (multi_model)
# -
# 이제 훈련 된 모델을 사용하여 테스트 기능의 레이블을 예측하고 예측 된 레이블을 실제 레이블과 비교할 수 있습니다.
# + tags=[]
penguin_predictions = multi_model.predict(x_penguin_test)
print('Predicted labels: ', penguin_predictions[:15])
print('Actual labels : ' ,y_penguin_test[:15])
# -
# 분류 보고서(classification report)를 살펴 보겠습니다.
# + tags=[]
from sklearn. metrics import classification_report
print(classification_report(y_penguin_test, penguin_predictions))
# -
# 이진 분류와 마찬가지로 보고서에는 각 클래스에 대한 *정밀도* 및 *재현율* 메트릭이 포함됩니다. 그러나 이진 분류를 사용하면 *positive* 클래스의 점수에 집중할 수 있습니다. 이 경우에는 여러 클래스가 있으므로 전체 메트릭(매크로 또는 가중 평균)을 검토하여 세 클래스 모두에서 모델이 얼마나 잘 수행되는지 파악해야합니다.
#
# scikit-learn 메트릭 점수 클래스를 사용하여 보고서와 별도로 전체 메트릭을 가져올 수 있지만 다중 클래스 결과를 사용하려면 정밀도 및 재현율에 사용할 평균 메트릭을 지정해야합니다.
# + tags=[]
from sklearn.metrics import accuracy_score, precision_score, recall_score
print("Overall Accuracy:",accuracy_score(y_penguin_test, penguin_predictions))
print("Overall Precision:",precision_score(y_penguin_test, penguin_predictions, average='macro'))
print("Overall Recall:",recall_score(y_penguin_test, penguin_predictions, average='macro'))
# -
# 이제 모델에 대한 혼동 행렬(confusion_matrix)을 살펴 보겠습니다.
# + tags=[]
from sklearn.metrics import confusion_matrix
# Print the confusion matrix
mcm = confusion_matrix(y_penguin_test, penguin_predictions)
print(mcm)
# -
# 혼동 행렬은 각 클래스에 대한 예측 및 실제 레이블 값의 교차점을 보여줍니다. 간단히 말해서 왼쪽 상단에서 오른쪽 하단까지의 대각선 교차는 올바른 예측 수를 나타냅니다.
#
# 여러 클래스를 처리 할 때 일반적으로 다음과 같이 이것을 히트맵으로 시각화하는 것이 더 직관적입니다.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.imshow(mcm, interpolation="nearest", cmap=plt.cm.Blues)
plt.colorbar()
tick_marks = np.arange(len(penguin_classes))
plt.xticks(tick_marks, penguin_classes, rotation=45)
plt.yticks(tick_marks, penguin_classes)
plt.xlabel("Predicted Species")
plt.ylabel("Actual Species")
plt.show()
# -
# 혼동 행렬의 어두운 정사각형은 사례 수가 많다는 것을 나타내며, 예측 레이블과 실제 레이블이 동일한 경우를 보고자 하면 정사각형의 대각선 어두움 정도를 보면 됩니다.
#
# 다중 클래스 분류 모델의 경우 참 양성 비율과 위양성 비율을 보여주는 단일 ROC 곡선은 불가능합니다. 그러나 OVR (One vs Rest) 비교에서 각 클래스의 One vs Rest을 사용하여 각 클래스에 대한 ROC 차트를 만들 수 있습니다.
# +
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
# Get class probability scores
penguin_prob = multi_model.predict_proba(x_penguin_test)
# Get ROC metrics for each class
fpr = {}
tpr = {}
thresh ={}
for i in range(len(penguin_classes)):
fpr[i], tpr[i], thresh[i] = roc_curve(y_penguin_test, penguin_prob[:,i], pos_label=i)
# Plot the ROC chart
plt.plot(fpr[0], tpr[0], linestyle='--',color='orange', label=penguin_classes[0] + ' vs Rest')
plt.plot(fpr[1], tpr[1], linestyle='--',color='green', label=penguin_classes[1] + ' vs Rest')
plt.plot(fpr[2], tpr[2], linestyle='--',color='blue', label=penguin_classes[2] + ' vs Rest')
plt.title('Multiclass ROC curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive rate')
plt.legend(loc='best')
plt.show()
# -
# ROC 성능을 정량화하기 위해 모든 OVR 곡선에서 평균화 된 곡선 점수 아래의 집계 영역을 계산할 수 있습니다.
auc = roc_auc_score(y_penguin_test,penguin_prob, multi_class='ovr')
print('Average AUC:', auc)
# ### 파이프라인의 데이터 전처리
#
# 이진 분류와 마찬가지로 파이프라인을 사용하여 데이터를 모델 학습 전에 데이터에 전처리 단계를 적용할 수 있습니다. 훈련 전에 변환 단계에서 수치형 특성을 스케일링하여 펭귄 예측를 개선 할 수 있는지 살펴 보겠습니다. 이번에는 다른 알고리즘 (서포트 벡터 머신)을 시도하겠습니다.
# + tags=[]
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC
# Define preprocessing for numeric columns (scale them)
feature_columns = [0,1,2,3]
feature_transformer = Pipeline(steps=[
('scaler', StandardScaler())
])
# Create preprocessing steps
preprocessor = ColumnTransformer(
transformers=[
('preprocess', feature_transformer, feature_columns)])
# Create training pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('regressor', SVC(probability=True))])
# fit the pipeline to train a linear regression model on the training set
multi_model = pipeline.fit(x_penguin_train, y_penguin_train)
print (multi_model)
# -
# 이제 새로운 모델을 평가할 수 있습니다.
# + tags=[]
# Get predictions from test data
penguin_predictions = multi_model.predict(x_penguin_test)
penguin_prob = multi_model.predict_proba(x_penguin_test)
# Overall metrics
print("Overall Accuracy:",accuracy_score(y_penguin_test, penguin_predictions))
print("Overall Precision:",precision_score(y_penguin_test, penguin_predictions, average='macro'))
print("Overall Recall:",recall_score(y_penguin_test, penguin_predictions, average='macro'))
print('Average AUC:', roc_auc_score(y_penguin_test,penguin_prob, multi_class='ovr'))
# Confusion matrix
plt.imshow(mcm, interpolation="nearest", cmap=plt.cm.Blues)
plt.colorbar()
tick_marks = np.arange(len(penguin_classes))
plt.xticks(tick_marks, penguin_classes, rotation=45)
plt.yticks(tick_marks, penguin_classes)
plt.xlabel("Predicted Species")
plt.ylabel("Actual Species")
plt.show()
# -
# ### 학습한 모델에 새 데이터 관측치를 이용
#
# 이제 훈련 된 모델을 저장하여 나중에 다시 사용할 수 있도록 하겠습니다.
# +
import joblib
# Save the model as a pickle file
filename = './models/penguin_model.pkl'
joblib.dump(multi_model, filename)
# -
# 이제 훈련 된 모델이 생겼습니다. 새로운 펭귄 관측 데이터 클래스를 예측하는데 사용하겠습니다.
# + tags=[]
# Load the model from the file
multi_model = joblib.load(filename)
# The model accepts an array of feature arrays (so you can predict the classes of multiple penguin observations in a single call)
# We'll create an array with a single array of features, representing one penguin
x_new = np.array([[50.4,15.3,224,5550]])
print ('New sample: {}'.format(x_new[0]))
# The model returns an array of predictions - one for each set of features submitted
# In our case, we only submitted one penguin, so our prediction is the first one in the resulting array.
penguin_pred = multi_model.predict(x_new)[0]
print('Predicted class is', penguin_classes[penguin_pred])
# -
# 또한 펭귄 관측치 배치 만큼을 모델에 제출하고 예측을 받아올 수 있습니다.
# + tags=[]
# This time our input is an array of two feature arrays
x_new = np.array([[49.5,18.4,195, 3600],
[38.2,20.1,190,3900]])
print ('New samples:\n{}'.format(x_new))
# Call the web service, passing the input data
predictions = multi_model.predict(x_new)
# Get the predicted classes.
for prediction in predictions:
print(prediction, '(' + penguin_classes[prediction] +')')
# -
# ## 더 읽을 거리
#
# 분류는 기계 학습의 가장 일반적인 형태 중 하나이며,이 노트북에서 논의한 기본 원칙을 따르면 scikit-learn으로 분류 모델을 학습하고 평가할 수 있습니다. 분류 알고리즘을 좀 더 깊이 조사하는데 시간을 할애 할 가치가 있으며, 좋은 출발점은 [Scikit-Learn 설명서](https://scikit-learn.org/stable/user_guide.html) 입니다.
#
# ## 과제 : 와인 분류
#
# 분류 모델을 교육하고 싶으십니까? [/challenge/03 - Wine Classification Challenge.ipynb](/Challenge/03%20-%20Wine%20Classification%20Challenge.ipynb) 노트북에서 도전을 시도하여 와인을 포도 품종으로 분류할 수 있는지 확인해 보십시오!
#
# > **참고**: 이 선택적 과제를 완료하는 시간은 이 연습의 예상 시간에 포함되지 않습니다. 원하는 만큼 시간을 적게 또는 많이 할애할 수 있습니다!
| 03 - Classification(KR).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext nb_black
# +
import numpy as np
import matplotlib.pyplot as plt
from quantum_systems import ODQD, GeneralOrbitalSystem
# +
l = 10
grid_length = 10
num_grid_points = 2001
omega = 0.25
odho = ODQD(
l, grid_length, num_grid_points, a=0.25, alpha=1, potential=ODQD.HOPotential(omega)
)
# -
# \begin{align}
# \epsilon_i = \hbar \omega \left(i + \frac{1}{2}\right)
# \end{align}
np.diag(odho.h)
# +
fig = plt.figure(figsize=(16, 10))
plt.plot(odho.grid, ODQD.HOPotential(omega)(odho.grid))
for i in range(l):
plt.plot(
odho.grid,
np.abs(odho.spf[i]) ** 2 + odho.h[i, i].real,
label=r"$\psi_{" + f"{i}" + r"}$",
)
plt.grid()
plt.legend()
plt.show()
# -
print(f"l = {odho.l}")
print(f"grid shape = {odho.grid.shape}")
print(f"h shape = {odho.h.shape}")
print(f"u shape = {odho.u.shape}")
print(f"x shape = {odho.position.shape}")
print(f"spf shape = {odho.spf.shape}")
system = GeneralOrbitalSystem(n=2, basis_set=odho, anti_symmetrize_u=True)
print(f"l = {system.l}")
print(f"grid shape = {system._basis_set.grid.shape}")
print(f"h shape = {system.h.shape}")
print(f"u shape = {system.u.shape}")
print(f"x shape = {system.position.shape}")
print(f"spf shape = {system.spf.shape}")
# __Note:__ `system.u` in the `GeneralOrbitalSystem` are the anti-symmetric Coulomb matrix elements.
# \begin{align}
# \hat{H}
# &= \hat{t} + \hat{v} + \hat{u}
# = \hat{h} + \hat{u}
# = \sum_{i}\left(
# -\frac{1}{2}\frac{d^2}{dx^2}
# + \frac{1}{2}\omega^2 x^2
# \right)
# + \sum_{i < j} \frac{\alpha}{\sqrt{(x_i - x_j)^2 + a^2}}.
# \end{align}
#
# \begin{align}
# \hat{H} \Psi(x_1, \dots, x_N) = E \Psi(x_1, \dots, x_N)
# \end{align}
# In Hartree-Fock:
# \begin{align}
# \Psi(x_1, \dots, x_N) \approx \Phi(x_1, \dots, x_N)
# = \begin{vmatrix}
# \phi_1(x_1) & \dots & \phi_1(x_N) \\
# \vdots & \ddots & \vdots \\
# \phi_N(x_1) & \dots & \phi_N(x_N)
# \end{vmatrix}
# \end{align}
#
# Variational principle:
# \begin{align}
# E_{gs} \leq E[\Psi] = \langle \Psi | \hat{H} | \Psi \rangle
# \end{align}
#
# Minimization:
# \begin{align}
# L = \langle \Phi | \hat{H} | \Phi \rangle - \lambda_{ji}\left( \langle \phi_i | \phi_j \rangle - \delta_{ij} \right)
# \end{align}
#
# $\implies$ Modern Quantum Chemistry - Szabo & Östlund $\implies$
#
# \begin{align}
# \hat{f}|\phi_i\rangle = \varepsilon_i |\phi_i\rangle.
# \end{align}
# Atomic orbitals (harmonic oscillator orbitals) $\{\chi_{\alpha}\}_{\alpha = 1}^{l}$ to the molecular orbitals (Hartree-Fock orbitals):
#
# \begin{align}
# |\phi_i \rangle = C_{\alpha i} | \chi_{\alpha} \rangle,
# \end{align}
#
# where $\{\phi_i\}_{i = 1}^{N}$ and $N$ is not necessarily equal to $l$.
# Inserting the basis transformation into the Hartree-Fock equations:
# \begin{gather}
# \hat{f} C_{\alpha i} | \chi_{\alpha} \rangle = \varepsilon_i C_{\alpha i} | \chi_{\alpha} \rangle
# \end{gather}
#
# Left-projecting with $\chi_{\beta}$:
# \begin{gather}
# C_{\alpha i} \langle \chi_{\beta} | \hat{f} | \chi_{\alpha} \rangle
# = \varepsilon_i C_{\alpha i} \langle \chi_{\beta} | \chi_\alpha \rangle
# = \varepsilon_i C_{\beta i},
# \end{gather}
# as $\langle \chi_{\beta} | \chi_\alpha \rangle = \delta_{\beta \alpha}$ in our case.
# Define the matrices $\mathbf{F}$ and $\mathbf{C}$, and the vector $\boldsymbol{\varepsilon}$. The elements of $\mathbf{F}$ are
# \begin{gather}
# [\mathbf{F}]_{\beta \alpha} \equiv \langle \chi_{\beta} | \hat{f} | \chi_{\alpha} \rangle.
# \end{gather}
#
# This lets us write the Roothan-Hall equations:
# \begin{align}
# \mathbf{F} \mathbf{C} = \mathbf{C} \boldsymbol{\varepsilon},
# \end{align}
# which is a generalized eigenvalue equation.
# The Fock operator:
# \begin{align}
# \hat{f} = \hat{h} + \hat{u}^{direct} + \hat{u}^{exchange}
# \end{align}
#
# The matrix elements are:
# \begin{align}
# \langle \chi_{\beta} | \hat{f} | \chi_{\alpha} \rangle
# = h_{\beta \alpha} + u^{direct}_{\beta \alpha} + u^{exchange}_{\beta \alpha},
# \end{align}
# where the one-body Hamiltonian is:
# \begin{align}
# h_{\beta \alpha}
# = \int dx \chi_{\beta}(x) h \chi_{\alpha}(x),
# \end{align}
# (this is found in `odho.h` in the code.)
# the direct interaction is:
# \begin{align}
# u^{direct}_{\beta \alpha}
# = \sum_{i}^{N} \langle \chi_{\beta} \phi_i | \hat{u} | \chi_{\alpha} \phi_i \rangle,
# \end{align}
# the exchange interaction is:
# \begin{align}
# u^{exchange}_{\beta \alpha}
# = \sum_{i}^{N} \langle \chi_{\beta} \phi_i | \hat{u} | \phi_i \chi_{\alpha} \rangle,
# \end{align}
# \begin{gather}
# \langle \chi_{\beta} | \hat{f} | \chi_{\alpha} \rangle
# = h_{\beta \alpha} + C^{*}_{??} C_{??} u_{\beta ? ? \alpha}
# \end{gather}
#
# The matrix elements $u^{\alpha \beta}_{\gamma \delta}$ can be found in `odho.u`. They are labelled:
# \begin{align}
# u^{\alpha \beta}_{\gamma \delta}
# = \int d x_1 d x_2 \chi^{*}_{\alpha}(x_1) \chi^{*}_{\beta}(x_2) u(x_1, x_2) \chi_{\gamma}(x_1) \chi_{\delta}(x_2)
# \end{align}
| odho-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 수학 기초
# ## Day 1 함수
import numpy as np
a = np.arange(1,6)
print(a)
b = np.arange(1, 10, 2)
print(b)
a.sum()
a.prod()
b.sum()
b.prod()
def identity(x):
y = x
return y
identity(2)
[identity(x) for x in a]
def constant(x):
y = 1
return y
[constant(x) for x in a]
def square(x):
y = x ** 2
return y
[square(x) for x in a]
def double(x):
y = x * 2
return y
[double(x) for x in a]
def composition(f, g):
return lambda x: f(g(x))
comp1 = composition(double, square)
comp2 = composition(square, double)
comp1(2)
comp2(2)
def summation(x, y):
z = x + y
return z
summation(10, 20)
def production(x, y):
z = x * y
return z
production(10,20)
np.exp(0)
np.exp(1)
np.exp(2+3)
np.exp(7-2)
np.exp(7) / np.exp(2)
def exponential(a, b):
return a ** b
exponential(2,3)
exponential(2, 1+2)
exponential(2, 1) * exponential(2, 2)
np.log(10)
np.log2(10)
np.log10(10)
np.log10(2 * 5)
np.log10(2) + np.log10(5)
np.log10(20/2)
np.log10(20) - np.log10(2)
import math
theta = math.radians(90)
math.sin(theta)
math.cos(theta)
math.tan(theta)
| python/sem0/Math Python Day1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.1
# language: julia
# name: julia-1.3
# ---
# # Test if MendelImpute can impute untyped SNPs
using Revise
using VCFTools
using MendelImpute
using GeneticVariation
using Random
using StatsBase
# ## Generate subset of markers for prephasing
cd("/Users/biona001/.julia/dev/MendelImpute/data/1000_genome_phase3_v5/filtered")
function filter_and_mask()
for chr in [22]
# filter chromosome data for unique snps
# data = "../raw/ALL.chr$chr.phase3_v5.shapeit2_mvncall_integrated.noSingleton.genotypes.vcf.gz"
# full_record_index = .!find_duplicate_marker(data)
# @time VCFTools.filter(data, full_record_index, 1:nsamples(data),
# des = "chr$chr.uniqueSNPs.vcf.gz")
# summarize data
total_snps, samples, _, _, _, maf_by_record, _ = gtstats("chr$chr.uniqueSNPs.vcf.gz")
# generate target file with 250 samples and 100k snps with maf>0.05
n = 250
p = 100000
record_idx = falses(total_snps)
large_maf = findall(x -> x > 0.05, maf_by_record)
shuffle!(large_maf)
record_idx[large_maf[1:p]] .= true
sample_idx = falses(samples)
sample_idx[1:n] .= true
shuffle!(sample_idx)
@time VCFTools.filter("chr$chr.uniqueSNPs.vcf.gz", record_idx, sample_idx,
des = "target.chr$chr.typedOnly.vcf.gz")
# generate target panel with all snps
@time VCFTools.filter("chr$chr.uniqueSNPs.vcf.gz",
1:total_snps, sample_idx, des = "target.chr$chr.full.vcf.gz")
# also generate reference panel without target samples
@time VCFTools.filter("chr$chr.uniqueSNPs.vcf.gz",
1:total_snps, .!sample_idx, des = "ref.chr$chr.excludeTarget.vcf.gz")
# unphase and mask 1% entries in target file
masks = falses(p, n)
missingprop = 0.01
for j in 1:n, i in 1:p
rand() < missingprop && (masks[i, j] = true)
end
@time mask_gt("target.chr$chr.typedOnly.vcf.gz", masks,
des="target.chr$chr.typedOnly.masked.vcf.gz", unphase=true)
# generate subset of reference file that matches target file
@time conformgt_by_pos("ref.chr$chr.excludeTarget.vcf.gz",
"target.chr$(chr).typedOnly.masked.vcf.gz",
"chr$chr.aligned", "$chr", 1:typemax(Int))
if nrecords("chr$chr.aligned.tgt.vcf.gz") == p
rm("chr$chr.aligned.tgt.vcf.gz", force=true) # perfect match
else
error("target file has SNPs not matching in reference file! Shouldn't happen!")
end
mv("chr22.aligned.ref.vcf.gz", "ref.chr22.aligned.vcf.gz", force=true)
end
end
Random.seed!(2020)
@time filter_and_mask()
# ### Missing rate
#
# In typed markers, 1% of data is missing at random. In addition, only 84% of all markers are not typed (i.e. systematically missing).
tgtfile = "target.chr22.typedOnly.masked.vcf.gz"
reffile = "ref.chr22.excludeTarget.vcf.gz"
missing_rate = 1 - nrecords(tgtfile) / nrecords(reffile)
# # MendelImpute on untyped markers with dp
Threads.nthreads()
tgtfile = "target.chr22.typedOnly.masked.vcf.gz"
reffile = "ref.chr22.excludeTarget.vcf.gz"
reffile_aligned = "ref.chr22.aligned.vcf.gz"
X_typedOnly_complete = "target.chr22.typedOnly.vcf.gz"
X_full_complete = "target.chr22.full.vcf.gz"
@show nrecords(tgtfile), nsamples(tgtfile)
@show nrecords(reffile), nsamples(reffile)
@show nrecords(reffile_aligned), nsamples(reffile_aligned)
@show nrecords(X_typedOnly_complete), nsamples(X_typedOnly_complete)
@show nrecords(X_full_complete), nsamples(X_full_complete)
# dp method using incomplete ref panel
cd("/Users/biona001/.julia/dev/MendelImpute/data/1000_genome_phase3_v5/filtered")
Random.seed!(2020)
function run()
X_complete = convert_gt(Float32, "target.chr22.full.vcf.gz")
n, p = size(X_complete)
chr = 22
for width in [500, 1000, 2000]
println("Imputing typed + untyped SNPs with dynamic programming, width = $width")
tgtfile = "target.chr$chr.typedOnly.masked.vcf.gz"
reffile = "ref.chr$chr.excludeTarget.vcf.gz"
outfile = "mendel.imputed.dp$width.vcf.gz"
@time phase(tgtfile, reffile, impute=true,
outfile = outfile, width = width, fast_method=false)
X_mendel = convert_gt(Float32, outfile)
println("error overall = $(sum(X_mendel .!= X_complete) / n / p) \n")
end
end
run()
# # Beagle 5.0 on incomplete ref panel
# beagle 5
cd("/Users/biona001/.julia/dev/MendelImpute/data/1000_genome_phase3_v5/filtered")
function beagle()
chr = 22
tgtfile = "target.chr$chr.typedOnly.masked.vcf.gz"
reffile = "ref.chr$chr.excludeTarget.vcf.gz"
outfile = "beagle.imputed"
Base.run(`java -Xmx15g -jar beagle.28Sep18.793.jar gt=$tgtfile ref=$reffile out=$outfile nthreads=4`)
# beagle error rate
X_complete = convert_gt(Float32, "target.chr22.full.vcf.gz")
X_beagle = convert_gt(Float32, "beagle.imputed.vcf.gz")
n, p = size(X_complete)
println("error overall = $(sum(X_beagle .!= X_complete) / n / p) \n")
end
beagle()
# # Eagle 2 + Minimac4
#
# In order to use the reference panel in Eagle 2's prephase option, one must first convert it to `.bcf` format via e.g. `htslib` which is *extremely* difficult to install. Even after we went through all the hard work to obtain the final `.bcf` reference file (see commands below), eagle 2.4 STILL SAYS the file is not acceptable (not bgzipped or some processing error). Therefore, I have no choice but to prephase without the reference panel.
# THIS CHUNK OF CODE DID NOT WORK
module load htslib/1.9 # load htslib on hoffman
gunzip ref.chr22.excludeTarget.vcf.gz # unzip .vcf.gz
bgzip ref.chr22.excludeTarget.vcf # bgzip the vcf file
bcftools index -f ref.chr22.excludeTarget.vcf.gz # load some required index, not sure if needed
bcftools view ref.chr22.excludeTarget.vcf > ref.chr22.excludeTarget.bcf # create .bcf -> CREATED FILE BUT says not bgzipped
bgzip ref.chr22.excludeTarget.bcf # try to bgzip .bcf file -> cannot do so
# run eagle 2.4: 3367.79 sec on amd-2382 machine (can only run on linux systems)
eagle --vcf=target.chr22.typedOnly.masked.vcf.gz --outPrefix=eagle.phased --numThreads=4 --geneticMapFile=../Eagle_v2.4.1/tables/genetic_map_hg19_withX.txt.gz
# convert ref file to m3vcf format (Total Run completed in 1 hours, 46 mins, 24 seconds)
/u/home/b/biona001/haplotype_comparisons/Minimac3/bin/Minimac3 --refHaps ref.chr22.excludeTarget.vcf.gz --processReference --prefix ref.chr22.excludeTarget
# run minimac4 (2619 seconds)
minimac4 --refHaps ref.chr22.excludeTarget.m3vcf.gz --haps eagle.phased.vcf.gz --prefix minimac.imputed.chr22 --format GT --cpus 4
# minimac4 error rate
X_complete = convert_gt(Float32, "target.chr22.full.vcf.gz")
X_minimac = convert_gt(Float32, "minimac.imputed.chr22.dose.vcf.gz")
n, p = size(X_complete)
println("error overall = $(sum(X_minimac .!= X_complete) / n / p) \n")
| data/1000_genome_phase3_v5/filtered/Impute_untyped_chrom22.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["remove-cell"] active=""
# ---
# title: "Basic queries"
# teaching: 3000
# exercises: 0
# questions:
# - "How can we select and download the data we want from the Gaia server?"
#
# objectives:
# - "Compose a basic query in ADQL/SQL."
# - "Use queries to explore a database and its tables."
# - "Use queries to download data."
# - "Develop, test, and debug a query incrementally."
# keypoints:
# - "If you can't download an entire dataset (or it's not practical) use queries to select the data you need."
#
# - "Read the metadata and the documentation to make sure you understand the tables, their columns, and what they mean."
#
# - "Develop queries incrementally: start with something simple, test it, and add a little bit at a time."
#
# - "Use ADQL features like `TOP` and `COUNT` to test before you run a query that might return a lot of data."
#
# - "If you know your query will return fewer than 3000 rows, you can
# run it synchronously, which might complete faster (but it doesn't seem to make much difference). If it might return more than 3000 rows, you should run it asynchronously."
#
# - "ADQL and SQL are not case-sensitive, so you don't have to
# capitalize the keywords, but you should."
#
# - "ADQL and SQL don't require you to break a query into multiple
# lines, but you should."
#
# ---
#
# {% include links.md %}
#
# -
# # 1. Queries
#
# This is the first in a series of lessons about working with astronomical data.
#
# As a running example, we will replicate parts of the analysis in a recent paper, "[Off the beaten path: Gaia reveals GD-1 stars outside of the main stream](https://arxiv.org/abs/1805.00425)" by <NAME> and <NAME>.
# ## Outline
#
# This lesson demonstrates the steps for selecting and downloading data from the Gaia Database:
#
# 1. First we'll make a connection to the Gaia server,
#
# 2. We will explore information about the database and the tables it contains,
#
# 3. We will write a query and send it to the server, and finally
#
# 4. We will download the response from the server.
#
# ## Query Language
#
# In order to select data from a database, you have to compose a query, which is a program written in a "query language".
# The query language we'll use is ADQL, which stands for "Astronomical Data Query Language".
#
# ADQL is a dialect of [SQL](https://en.wikipedia.org/wiki/SQL) (Structured Query Language), which is by far the most commonly used query language. Almost everything you will learn about ADQL also works in SQL.
#
# [The reference manual for ADQL is here](http://www.ivoa.net/documents/ADQL/20180112/PR-ADQL-2.1-20180112.html).
# But you might find it easier to learn from [this ADQL Cookbook](https://www.gaia.ac.uk/data/gaia-data-release-1/adql-cookbook).
# ## Using Jupyter
#
# If you have not worked with Jupyter notebooks before, you might start with [the tutorial on from Jupyter.org called "Try Classic Notebook"](https://jupyter.org/try), or [this tutorial from DataQuest](https://www.dataquest.io/blog/jupyter-notebook-tutorial/).
#
# There are two environments you can use to write and run notebooks:
#
# * "Jupyter Notebook" is the original, and
#
# * "Jupyter Lab" is a newer environment with more features.
#
# For these lessons, you can use either one.
# If you are too impatient for the tutorials, here are the most important things to know:
#
# 1. Notebooks are made up of code cells and text cells (and a few other less common kinds). Code cells contain code; text cells, like this one, contain explanatory text written in [Markdown](https://www.markdownguide.org/).
#
# 2. To run a code cell, click the cell to select it and press Shift-Enter. The output of the code should appear below the cell.
#
# 3. In general, notebooks only run correctly if you run every code cell in order from top to bottom. If you run cells out of order, you are likely to get errors.
#
# 4. You can modify existing cells, but then you have to run them again to see the effect.
#
# 5. You can add new cells, but again, you have to be careful about the order you run them in.
#
# 6. If you have added or modified cells and the behavior of the notebook seems strange, you can restart the "kernel", which clears all of the variables and functions you have defined, and run the cells again from the beginning.
#
# * If you are using Jupyter notebook, open the `Kernel` menu and select "Restart and Run All".
#
# * In Jupyter Lab, open the `Kernel` menu and select "Restart Kernel and Run All Cells"
#
# * In Colab, open the `Runtime` menu and select "Restart and run all"
#
# Before you go on, you might want to explore the other menus and the toolbar to see what else you can do.
# + [markdown] tags=["remove-cell"]
# ## Installing libraries
#
# If you are running this notebook on Colab, you should run the following cell to install the libraries we'll need.
#
# If you are running this notebook on your own computer, you might have to install these libraries yourself.
# + tags=["remove-cell"]
# If we're running on Colab, install libraries
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
# !pip install astroquery
# -
# ## Connecting to Gaia
#
# The library we'll use to get Gaia data is [Astroquery](https://astroquery.readthedocs.io/en/latest/).
# Astroquery provides `Gaia`, which is an [object that represents a connection to the Gaia database](https://astroquery.readthedocs.io/en/latest/gaia/gaia.html).
#
# We can connect to the Gaia database like this:
from astroquery.gaia import Gaia
# This import statement creates a [TAP+](http://www.ivoa.net/documents/TAP/) connection; TAP stands for "Table Access Protocol", which is a network protocol for sending queries to the database and getting back the results.
# ## Databases and Tables
#
# What is a database, anyway? Most generally, it can be any collection of data, but when we are talking about ADQL or SQL:
#
# * A database is a collection of one or more named tables.
#
# * Each table is a 2-D array with one or more named columns of data.
#
# We can use `Gaia.load_tables` to get the names of the tables in the Gaia database. With the option `only_names=True`, it loads information about the tables, called "metadata", not the data itself.
tables = Gaia.load_tables(only_names=True)
# The following `for` loop prints the names of the tables.
# + tags=["hide-output", "truncate-output"]
for table in tables:
print(table.name)
# -
# So that's a lot of tables. The ones we'll use are:
#
# * `gaiadr2.gaia_source`, which contains Gaia data from [data release 2](https://www.cosmos.esa.int/web/gaia/data-release-2),
#
# * `gaiadr2.panstarrs1_original_valid`, which contains the photometry data we'll use from PanSTARRS, and
#
# * `gaiadr2.panstarrs1_best_neighbour`, which we'll use to cross-match each star observed by Gaia with the same star observed by PanSTARRS.
#
# We can use `load_table` (not `load_tables`) to get the metadata for a single table. The name of this function is misleading, because it only downloads metadata, not the contents of the table.
meta = Gaia.load_table('gaiadr2.gaia_source')
meta
# Jupyter shows that the result is an object of type `TapTableMeta`, but it does not display the contents.
#
# To see the metadata, we have to print the object.
print(meta)
# ## Columns
#
# The following loop prints the names of the columns in the table.
# + tags=["hide-output", "truncate-output"]
for column in meta.columns:
print(column.name)
# -
# You can probably infer what many of these columns are by looking at the names, but you should resist the temptation to guess.
# To find out what the columns mean, [read the documentation](https://gea.esac.esa.int/archive/documentation/GDR2/Gaia_archive/chap_datamodel/sec_dm_main_tables/ssec_dm_gaia_source.html).
#
# If you want to know what can go wrong when you don't read the documentation, [you might like this article](https://www.vox.com/future-perfect/2019/6/4/18650969/married-women-miserable-fake-paul-dolan-happiness).
# ### Exercise
#
# One of the other tables we'll use is `gaiadr2.panstarrs1_original_valid`. Use `load_table` to get the metadata for this table. How many columns are there and what are their names?
# + tags=["hide-cell"]
# Solution
meta2 = Gaia.load_table('gaiadr2.panstarrs1_original_valid')
print(meta2)
for column in meta2.columns:
print(column.name)
# -
# ## Writing queries
#
# By now you might be wondering how we download these tables. With tables this big, you generally don't. Instead, you use queries to select only the data you want.
#
# A query is a string written in a query language like SQL; for the Gaia database, the query language is a dialect of SQL called ADQL.
#
# Here's an example of an ADQL query.
query1 = """SELECT
TOP 10
source_id, ra, dec, parallax
FROM gaiadr2.gaia_source
"""
# **Python note:** We use a [triple-quoted string](https://docs.python.org/3/tutorial/introduction.html#strings) here so we can include line breaks in the query, which makes it easier to read.
#
# The words in uppercase are ADQL keywords:
#
# * `SELECT` indicates that we are selecting data (as opposed to adding or modifying data).
#
# * `TOP` indicates that we only want the first 10 rows of the table, which is useful for testing a query before asking for all of the data.
#
# * `FROM` specifies which table we want data from.
#
# The third line is a list of column names, indicating which columns we want.
#
# In this example, the keywords are capitalized and the column names are lowercase. This is a common style, but it is not required. ADQL and SQL are not case-sensitive.
#
# Also, the query is broken into multiple lines to make it more readable. This is a common style, but not required. Line breaks don't affect the behavior of the query.
# To run this query, we use the `Gaia` object, which represents our connection to the Gaia database, and invoke `launch_job`:
job = Gaia.launch_job(query1)
job
# The result is an object that represents the job running on a Gaia server.
#
# If you print it, it displays metadata for the forthcoming results.
print(job)
# Don't worry about `Results: None`. That does not actually mean there are no results.
#
# However, `Phase: COMPLETED` indicates that the job is complete, so we can get the results like this:
results = job.get_results()
type(results)
# The `type` function indicates that the result is an [Astropy Table](https://docs.astropy.org/en/stable/table/).
#
# **Optional detail:** Why is `table` repeated three times? The first is the name of the module, the second is the name of the submodule, and the third is the name of the class. Most of the time we only care about the last one. It's like the Linnean name for gorilla, which is *Gorilla gorilla gorilla*.
# An Astropy `Table` is similar to a table in an SQL database except:
#
# * SQL databases are stored on disk drives, so they are persistent; that is, they "survive" even if you turn off the computer. An Astropy `Table` is stored in memory; it disappears when you turn off the computer (or shut down this Jupyter notebook).
#
# * SQL databases are designed to process queries. An Astropy `Table` can perform some query-like operations, like selecting columns and rows. But these operations use Python syntax, not SQL.
#
# Jupyter knows how to display the contents of a `Table`.
results
# Each column has a name, units, and a data type.
#
# For example, the units of `ra` and `dec` are degrees, and their data type is `float64`, which is a 64-bit [floating-point number](https://en.wikipedia.org/wiki/Floating-point_arithmetic), used to store measurements with a fraction part.
#
# This information comes from the Gaia database, and has been stored in the Astropy `Table` by Astroquery.
# ### Exercise
#
# Read [the documentation](https://gea.esac.esa.int/archive/documentation/GDR2/Gaia_archive/chap_datamodel/sec_dm_main_tables/ssec_dm_gaia_source.html) of this table and choose a column that looks interesting to you. Add the column name to the query and run it again. What are the units of the column you selected? What is its data type?
# +
# Solution
# Let's add
#
# radial_velocity : Radial velocity (double, Velocity[km/s] )
#
# Spectroscopic radial velocity in the solar barycentric
# reference frame.
#
# The radial velocity provided is the median value of the
# radial velocity measurements at all epochs.
query = """SELECT
TOP 10
source_id, ra, dec, parallax, radial_velocity
FROM gaiadr2.gaia_source
"""
# -
# ## Asynchronous queries
#
# `launch_job` asks the server to run the job "synchronously", which normally means it runs immediately. But synchronous jobs are limited to 2000 rows. For queries that return more rows, you should run "asynchronously", which mean they might take longer to get started.
#
# If you are not sure how many rows a query will return, you can use the SQL command `COUNT` to find out how many rows are in the result without actually returning them. We'll see an example in the next lesson.
#
# The results of an asynchronous query are stored in a file on the server, so you can start a query and come back later to get the results.
# For anonymous users, files are kept for three days.
#
# As an example, let's try a query that's similar to `query1`, with these changes:
#
# * It selects the first 3000 rows, so it is bigger than we should run synchronously.
#
# * It selects two additional columns, `pmra` and `pmdec`, which are proper motions along the axes of `ra` and `dec`.
#
# * It uses a new keyword, `WHERE`.
query2 = """SELECT
TOP 3000
source_id, ra, dec, pmra, pmdec, parallax
FROM gaiadr2.gaia_source
WHERE parallax < 1
"""
# A `WHERE` clause indicates which rows we want; in this case, the query selects only rows "where" `parallax` is less than 1. This has the effect of selecting stars with relatively low parallax, which are farther away.
# We'll use this clause to exclude nearby stars that are unlikely to be part of GD-1.
#
# `WHERE` is one of the most common clauses in ADQL/SQL, and one of the most useful, because it allows us to download only the rows we need from the database.
#
# We use `launch_job_async` to submit an asynchronous query.
job = Gaia.launch_job_async(query)
job
# And here are the results.
results = job.get_results()
results
# You might notice that some values of `parallax` are negative. As [this FAQ explains](https://www.cosmos.esa.int/web/gaia/archive-tips#negative%20parallax), "Negative parallaxes are caused by errors in the observations." They have "no physical meaning," but they can be a "useful diagnostic on the quality of the astrometric solution."
# ### Exercise
#
# The clauses in a query have to be in the right order. Go back and change the order of the clauses in `query2` and run it again.
# The modified query should fail, but notice that you don't get much useful debugging information.
#
# For this reason, developing and debugging ADQL queries can be really hard. A few suggestions that might help:
#
# * Whenever possible, start with a working query, either an example you find online or a query you have used in the past.
#
# * Make small changes and test each change before you continue.
#
# * While you are debugging, use `TOP` to limit the number of rows in the result. That will make each test run faster, which reduces your development time.
#
# * Launching test queries synchronously might make them start faster, too.
# +
# Solution
# In this example, the WHERE clause is in the wrong place
query = """SELECT
TOP 3000
WHERE parallax < 1
source_id, ref_epoch, ra, dec, parallax
FROM gaiadr2.gaia_source
"""
# -
# ## Operators
#
# In a `WHERE` clause, you can use any of the [SQL comparison operators](https://www.w3schools.com/sql/sql_operators.asp); here are the most common ones:
#
# | Symbol | Operation
# |--------| :---
# | `>` | greater than
# | `<` | less than
# | `>=` | greater than or equal
# | `<=` | less than or equal
# | `=` | equal
# | `!=` or `<>` | not equal
#
# Most of these are the same as Python, but some are not. In particular, notice that the equality operator is `=`, not `==`.
# Be careful to keep your Python out of your ADQL!
# You can combine comparisons using the logical operators:
#
# * AND: true if both comparisons are true
# * OR: true if either or both comparisons are true
#
# Finally, you can use `NOT` to invert the result of a comparison.
# ### Exercise
#
# [Read about SQL operators here](https://www.w3schools.com/sql/sql_operators.asp) and then modify the previous query to select rows where `bp_rp` is between `-0.75` and `2`.
# + tags=["hide-cell"]
# Solution
# Here's a solution using > and < operators
query = """SELECT
TOP 10
source_id, ref_epoch, ra, dec, parallax
FROM gaiadr2.gaia_source
WHERE parallax < 1
AND bp_rp > -0.75 AND bp_rp < 2
"""
# And here's a solution using the BETWEEN operator
query = """SELECT
TOP 10
source_id, ref_epoch, ra, dec, parallax
FROM gaiadr2.gaia_source
WHERE parallax < 1
AND bp_rp BETWEEN -0.75 AND 2
"""
# -
# `bp_rp` contains BP-RP color, which is the difference between two other columns, `phot_bp_mean_mag` and `phot_rp_mean_mag`.
# You can [read about this variable here](https://gea.esac.esa.int/archive/documentation/GDR2/Gaia_archive/chap_datamodel/sec_dm_main_tables/ssec_dm_gaia_source.html).
#
# This [Hertzsprung-Russell diagram](https://sci.esa.int/web/gaia/-/60198-gaia-hertzsprung-russell-diagram) shows the BP-RP color and luminosity of stars in the Gaia catalog (Copyright: ESA/Gaia/DPAC, CC BY-SA 3.0 IGO).
#
# <img width="300" src="https://github.com/AllenDowney/AstronomicalData/raw/main/images/1567214809100-ESA_Gaia_DR2_HRD_Gaia_625.jpg">
#
# Selecting stars with `bp-rp` less than 2 excludes many [class M dwarf stars](https://xkcd.com/2360/), which are low temperature, low luminosity. A star like that at GD-1's distance would be hard to detect, so if it is detected, it it more likely to be in the foreground.
# ## Formatting queries
#
# The queries we have written so far are string "literals", meaning that the entire string is part of the program.
# But writing queries yourself can be slow, repetitive, and error-prone.
#
# It is often better to write Python code that assembles a query for you. One useful tool for that is the [string `format` method](https://www.w3schools.com/python/ref_string_format.asp).
#
# As an example, we'll divide the previous query into two parts; a list of column names and a "base" for the query that contains everything except the column names.
#
# Here's the list of columns we'll select.
columns = 'source_id, ra, dec, pmra, pmdec, parallax'
# And here's the base; it's a string that contains at least one format specifier in curly brackets (braces).
query3_base = """SELECT
TOP 10
{columns}
FROM gaiadr2.gaia_source
WHERE parallax < 1
AND bp_rp BETWEEN -0.75 AND 2
"""
# This base query contains one format specifier, `{columns}`, which is a placeholder for the list of column names we will provide.
#
# To assemble the query, we invoke `format` on the base string and provide a keyword argument that assigns a value to `columns`.
query3 = query3_base.format(columns=columns)
# In this example, the variable that contains the column names and the variable in the format specifier have the same name.
# That's not required, but it is a common style.
#
# The result is a string with line breaks. If you display it, the line breaks appear as `\n`.
query3
# But if you print it, the line breaks appear as... line breaks.
print(query3)
# Notice that the format specifier has been replaced with the value of `columns`.
#
# Let's run it and see if it works:
job = Gaia.launch_job(query3)
print(job)
results = job.get_results()
results
# Good so far.
# ### Exercise
#
# This query always selects sources with `parallax` less than 1. But suppose you want to take that upper bound as an input.
#
# Modify `query3_base` to replace `1` with a format specifier like `{max_parallax}`. Now, when you call `format`, add a keyword argument that assigns a value to `max_parallax`, and confirm that the format specifier gets replaced with the value you provide.
# + tags=["hide-cell"]
# Solution
query_base = """SELECT
TOP 10
{columns}
FROM gaiadr2.gaia_source
WHERE parallax < {max_parallax} AND
bp_rp BETWEEN -0.75 AND 2
"""
query = query_base.format(columns=columns,
max_parallax=0.5)
print(query)
# -
# ## Summary
#
# This notebook demonstrates the following steps:
#
# 1. Making a connection to the Gaia server,
#
# 2. Exploring information about the database and the tables it contains,
#
# 3. Writing a query and sending it to the server, and finally
#
# 4. Downloading the response from the server as an Astropy `Table`.
#
# In the next lesson we will extend these queries to select a particular region of the sky.
# ## Best practices
#
# * If you can't download an entire dataset (or it's not practical) use queries to select the data you need.
#
# * Read the metadata and the documentation to make sure you understand the tables, their columns, and what they mean.
#
# * Develop queries incrementally: start with something simple, test it, and add a little bit at a time.
#
# * Use ADQL features like `TOP` and `COUNT` to test before you run a query that might return a lot of data.
#
# * If you know your query will return fewer than 2000 rows, you can run it synchronously, which might complete faster. If it might return more than 2000 rows, you should run it asynchronously.
#
# * ADQL and SQL are not case-sensitive, so you don't have to capitalize the keywords, but you should.
#
# * ADQL and SQL don't require you to break a query into multiple lines, but you should.
# Jupyter notebooks can be good for developing and testing code, but they have some drawbacks. In particular, if you run the cells out of order, you might find that variables don't have the values you expect.
#
# To mitigate these problems:
#
# * Make each section of the notebook self-contained. Try not to use the same variable name in more than one section.
#
# * Keep notebooks short. Look for places where you can break your analysis into phases with one notebook per phase.
| soln/01_query.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false
# 
#
# Exercise material of the MSc-level course **Numerical Methods in Geotechnical Engineering**.
# Held at Technische Universität Bergakademie Freiberg.
#
# Comments to:
#
# *Prof. Dr. <NAME>
# Chair of Soil Mechanics and Foundation Engineering
# Geotechnical Institute
# Technische Universität Bergakademie Freiberg.*
#
# https://tu-freiberg.de/en/soilmechanics
#
# -
# # Iteration schemes and their convergence: Newton-Raphson and Picard
# ## Example problem
#
# Given a function $f(x)$ we seek the solution $x^*$ such that $f(x^*) = 0$.
#
# Here, for illustrative purposes, we seek the solution to $f(x) = \sin x$. Analytically, we find for this periodic function $x^*_n = n \pi$ for $n \in Z$.
# +
import numpy as np #numerical methods
import matplotlib.pyplot as plt #plotting
from ipywidgets import widgets
from ipywidgets import interact
#Some plot settings
import plot_functions.plot_settings
# %run plot_functions/iteration_schemes_plots.ipynb
# -
# Let's define the function:
# ## Newton-Raphson scheme
#
# Anticipating an iterative scheme, we depart from the Taylor series expansion around the given solution $x_i$ in order to find the next solution $x_{i+1}$, which we truncate at first order and for which we demand equality to zero, as we're seeking $f(x^*) = 0$:
#
# $$
# f(x_{i+1}) \approx f(x_i) + \left. \frac{\partial f(x)}{\partial x} \right|_{x_i} (x_{i+1} - x_{i}) \overset{!}{=} 0
# $$
#
# By rearrangement, we find the iterative update procecture
#
# $$
# x_{i+i} = x_i - \left( \left. \frac{\partial f(x)}{\partial x} \right|_{x_i} \right)^{-1} f(x_i)
# $$
#
# The iteration is started at a starting guess $x_0$ which, ideally, is close to the sought solution. We will investigate it's role in the sequel.
#
# We stop the iteration once we're sufficiently close to zero:
#
# $$
# |f(x_i)| < \varepsilon_\text{tol} \quad \text{with } \varepsilon_\text{tol} \ll 1 \quad \rightarrow \quad x^* \approx x_i
# $$
# For the chosen problem we have:
#
# $$
# f(x) = \sin x \quad \rightarrow \quad f'(x) = \cos x
# $$
plot_interactive_newton()
# ### Tasks
#
# Increase the maximum iteration number one by one to see the iteration unfold. You can also play with the tolerance value.
#
# Now, keeping the tolerance fixed at $10^{-1}$ and the maximum iteration number at 10, answer the following questions.
#
# What happens -- and why -- if you
#
# * set the starting value to $x_0 = 2$?
# * set the starting value to $x_0 = 1.6$?
# * set the starting value to $x_0 = \pi/2$ or multiples thereof?
# * increase the starting value from 1.15 to 1.2 in steps of 0.01?
#
# Note your observations in terms of the solution found, the rate of convergence, etc.
# ## Picard iterations
#
# You're already familiar with this type of iterations. Remember, for example, the iteration rule for the method of slices for the simplified Bishop's method:
# Dies führt auf die Formel nach DIN4084:
#
# \begin{align*}
# \mu = \frac{E_\text{M,d}}{R_\text{M,d}} = \frac{\sum \limits_i \left[ P_\text{vi,d} + G_\text{i,d} \right] \sin \vartheta_i}{\displaystyle \sum \limits_i \frac{(G_{i,\text{d}} + P_{\text{v},i,\text{d}} - u_{i,\text{d}}b_i) \tan \varphi_{i,\text{d}} + c_{i,\text{d}}b_i}{\cos\vartheta_i + \mu \tan \varphi_{i,\text{d}} \sin \vartheta_i}}
# \end{align*}
#
# Vorgehen: Einsetzen von $\mu_0 = 1$ oder einem anderen geschätzten Startwert. Dann wiederholtes Einsetzen und Ausrechnen von $\mu$ bis
# \begin{align*}
# \left| \frac{\mu_{i+1}-\mu_i}{\mu_i} \right| < 0.03
# \end{align*}
#
# Konvergenz wird i.d.R. innerhalb weniger Iterationen erzielt.
# We observe, that we have an iteration update of the form
#
# $$
# x_{i+1} = g(x_i)
# $$
#
# Therefore, we re-cast our example function $f(x) = \sin x$ into the following form by using $f(x) = \sin x + x - x$:
#
# $$
# x = f(x) + x = g(x) = \sin x + x
# $$
#
# In other words, finding $x^*$ for $f(x^*) = 0$ is now the fixed point problem of identifying $x^* = g(x^*)$.
#
# That's it -- we can start looking at how it works:
plot_interactive_picard()
# ## Tasks
#
# * What do you observe for the identified solution as you vary the starting guess? Why is this?
# * What can you say about the convergence rate?
#
# Now let's repeat this by rearranging $f(x)$ slightly differently:
#
# $$
# g(x) = x - \sin(x)
# $$
#
# This definition is equivalent in terms of the roots ($\sin x$ and $-\sin x$ share the same roots).
plot_interactive_picard_inv()
# The attractors change, however the principal problem remains the same as before ....
#
#
# ## Additional material
#
# ### Modified Newton scheme
#
# In the lecture we saw the modified Newton schemes and the initial stiffness schemes. Here we look at such a modification by keeping keeping the slope constant:
#
# $$
# x_{i+i} = x_i - \left( \left. \frac{\partial f(x)}{\partial x} \right|_{x_0} \right)^{-1} f(x_i)
# $$
plot_interactive_newton_modified()
# ### Tasks
#
# Play with the starting values and observe the convergence rate. Especially, try $x_0 = [1.3, 1.1, 1.0, 0.8, 0.5, 0.2]$. What do you observe?
# ## Another function ..
#
# We now look at
#
# $$
# f(x) = e^{-x} - x
# $$
#
# ### Newton:
#
# The 'Jacobian' is given as $f'(x) = -e^{-x} - 1$
#
# ### Picard:
#
# The fixed point equation is given ax $x = g(x) = e^{-x}$.
plot_interactive_picard_2()
plot_interactive_newton_2()
# ### Tasks:
#
# * Do the methods converge to the same solution?
# * How quickly do they converge? Check how quickly the residuals decrease.
# * How would the modified Newton scheme perform in this simpler case?
| 04a_Newton_Picard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Ingest data with Athena
# This notebook demonstrates how to set up a database with Athena and query data with it.
#
# Amazon Athena is a serverless interactive query service that makes it easy to analyze your S3 data with standard SQL. It uses S3 as its underlying data store, and uses Presto with ANSI SQL support, and works with a variety of standard data formats, including CSV, JSON, ORC, Avro, and Parquet. Athena is ideal for quick, ad-hoc querying but it can also handle complex analysis, including large joins, window functions, and arrays.
#
# To get started, you can point to your data in Amazon S3, define the schema, and start querying using the built-in query editor. Amazon Athena allows you to tap into all your data in S3 without the need to set up complex processes to extract, transform, and load the data (ETL).
#
# ## Set up Athena
# First, we are going to make sure we have the necessary policies attached to the role that we used to create this notebook to access Athena. You can do this through an IAM client as shown below, or through the AWS console.
#
# **Note: You would need IAMFullAccess to attach policies to the role.**
# #### Attach IAMFullAccess Policy from Console
#
# **1.** Go to **SageMaker Console**, choose **Notebook instances** in the navigation panel, then select your notebook instance to view the details. Then under **Permissions and Encryption**, click on the **IAM role ARN** link and it will take you to your role summary in the **IAM Console**.
# <div>
# <img src="image/athena-iam-1.png" width="300"/>
# </div>
# **2.** Click on **Create Policy** under **Permissions**.
# <div>
# <img src="image/athena-iam-2.PNG" width="300"/>
# </div>
# **3.** In the **Attach Permissions** page, search for **IAMFullAccess**. It will show up in the policy search results if it has not been attached to your role yet. Select the checkbox for the **IAMFullAccess** Policy, then click **Attach Policy**. You now have the policy successfully attached to your role.
#
# <div>
# <img src="image/athena-iam-3.PNG" width="500"/>
# </div>
# %pip install -qU 'sagemaker>=2.15.0' 'PyAthena==1.10.7' 'awswrangler==1.2.0'
# +
import io
import boto3
import sagemaker
import json
from sagemaker import get_execution_role
import os
import sys
from sklearn.datasets import fetch_california_housing
import pandas as pd
from botocore.exceptions import ClientError
# Get region
session = boto3.session.Session()
region_name = session.region_name
# Get SageMaker session & default S3 bucket
sagemaker_session = sagemaker.Session()
bucket = sagemaker_session.default_bucket() # replace with your own bucket name if you have one
iam = boto3.client("iam")
s3 = sagemaker_session.boto_session.resource("s3")
role = sagemaker.get_execution_role()
role_name = role.split("/")[-1]
prefix = "data/tabular/california_housing"
filename = "california_housing.csv"
# -
# ### Download data from online resources and write data to S3
# This example uses the California Housing dataset, which was originally published in:
#
# > Pace, <NAME>, and <NAME>. "Sparse spatial autoregressions." Statistics & Probability Letters 33.3 (1997): 291-297.
# +
# helper functions to upload data to s3
def write_to_s3(filename, bucket, prefix):
# put one file in a separate folder. This is helpful if you read and prepare data with Athena
filename_key = filename.split(".")[0]
key = "{}/{}/{}".format(prefix, filename_key, filename)
return s3.Bucket(bucket).upload_file(filename, key)
def upload_to_s3(bucket, prefix, filename):
url = "s3://{}/{}/{}".format(bucket, prefix, filename)
print("Writing to {}".format(url))
write_to_s3(filename, bucket, prefix)
# +
tabular_data = fetch_california_housing()
tabular_data_full = pd.DataFrame(tabular_data.data, columns=tabular_data.feature_names)
tabular_data_full["target"] = pd.DataFrame(tabular_data.target)
tabular_data_full.to_csv("california_housing.csv", index=False)
upload_to_s3(bucket, "data/tabular", filename)
# -
# ### Set up IAM roles and policies
# When you run the following command, you will see an error that you cannot list policies if `IAMFullAccess` policy is not attached to your role. Please follow the steps above to attach the IAMFullAccess policy to your role if you see an error.
# check if IAM policy is attached
try:
existing_policies = iam.list_attached_role_policies(RoleName=role_name)["AttachedPolicies"]
if "IAMFullAccess" not in [po["PolicyName"] for po in existing_policies]:
print(
"ERROR: You need to attach the IAMFullAccess policy in order to attach policy to the role"
)
else:
print("IAMFullAccessPolicy Already Attached")
except ClientError as e:
if e.response["Error"]["Code"] == "AccessDenied":
print(
"ERROR: You need to attach the IAMFullAccess policy in order to attach policy to the role."
)
else:
print("Unexpected error: %s" % e)
# ### Create Policy Document
# We will create policies we used to access S3 and Athena. The two policies we will create here are:
# * S3FullAccess: `arn:aws:iam::aws:policy/AmazonS3FullAccess`
# * AthenaFullAccess: `arn:aws:iam::aws:policy/AmazonAthenaFullAccess`
#
# You can check the policy document in the IAM console and copy the policy file here.
athena_access_role_policy_doc = {
"Version": "2012-10-17",
"Statement": [
{"Effect": "Allow", "Action": ["athena:*"], "Resource": ["*"]},
{
"Effect": "Allow",
"Action": [
"glue:CreateDatabase",
"glue:DeleteDatabase",
"glue:GetDatabase",
"glue:GetDatabases",
"glue:UpdateDatabase",
"glue:CreateTable",
"glue:DeleteTable",
"glue:BatchDeleteTable",
"glue:UpdateTable",
"glue:GetTable",
"glue:GetTables",
"glue:BatchCreatePartition",
"glue:CreatePartition",
"glue:DeletePartition",
"glue:BatchDeletePartition",
"glue:UpdatePartition",
"glue:GetPartition",
"glue:GetPartitions",
"glue:BatchGetPartition",
],
"Resource": ["*"],
},
{"Effect": "Allow", "Action": ["lakeformation:GetDataAccess"], "Resource": ["*"]},
],
}
# create IAM client
iam = boto3.client("iam")
# create a policy
try:
response = iam.create_policy(
PolicyName="myAthenaPolicy", PolicyDocument=json.dumps(athena_access_role_policy_doc)
)
except ClientError as e:
if e.response["Error"]["Code"] == "EntityAlreadyExists":
print("Policy already created.")
else:
print("Unexpected error: %s" % e)
# get policy ARN
sts = boto3.client("sts")
account_id = sts.get_caller_identity()["Account"]
policy_athena_arn = f"arn:aws:iam::{account_id}:policy/myAthenaPolicy"
# ### Attach Policy to Role
# Attach a role policy
try:
response = iam.attach_role_policy(PolicyArn=policy_athena_arn, RoleName=role_name)
except ClientError as e:
if e.response["Error"]["Code"] == "EntityAlreadyExists":
print("Policy is already attached to your role.")
else:
print("Unexpected error: %s" % e)
# ## Intro to PyAthena
#
# We are going to leverage [PyAthena](https://pypi.org/project/PyAthena/) to connect and run Athena queries. PyAthena is a Python DB API 2.0 (PEP 249) compliant client for Amazon Athena. **Note that you will need to specify the region in which you created the database/table in Athena, making sure your catalog in the specified region that contains the database.**
from pyathena import connect
from pyathena.pandas_cursor import PandasCursor
from pyathena.util import as_pandas
# Set Athena database name
database_name = "tabular_california_housing"
# Set S3 staging directory -- this is a temporary directory used for Athena queries
s3_staging_dir = "s3://{0}/athena/staging".format(bucket)
# write the SQL statement to execute
statement = "CREATE DATABASE IF NOT EXISTS {}".format(database_name)
print(statement)
# connect to s3 using PyAthena
cursor = connect(region_name=region_name, s3_staging_dir=s3_staging_dir).cursor()
cursor.execute(statement)
# ### Register Table with Athena
# When you run a CREATE TABLE query in Athena, you register your table with the AWS Glue Data Catalog.
#
# To specify the path to your data in Amazon S3, use the LOCATION property, as shown in the following example: `LOCATION s3://bucketname/folder/`
#
# The LOCATION in Amazon S3 specifies all of the files representing your table. Athena reads all data stored in `s3://bucketname/folder/`. If you have data that you do not want Athena to read, do not store that data in the same Amazon S3 folder as the data you want Athena to read. If you are leveraging partitioning, to ensure Athena scans data within a partition, your WHERE filter must include the partition. For more information, see [Table Location and Partitions](https://docs.aws.amazon.com/athena/latest/ug/tables-location-format.html).
prefix = "data/tabular"
filename_key = "california_housing"
data_s3_location = "s3://{}/{}/{}/".format(bucket, prefix, filename_key)
table_name_csv = "california_housing_athena"
# +
# SQL statement to execute
statement = """CREATE EXTERNAL TABLE IF NOT EXISTS {}.{}(
MedInc double,
HouseAge double,
AveRooms double,
AveBedrms double,
Population double,
AveOccup double,
Latitude double,
Longitude double,
MedValue double
) ROW FORMAT DELIMITED FIELDS TERMINATED BY ',' LINES TERMINATED BY '\\n' LOCATION '{}'
TBLPROPERTIES ('skip.header.line.count'='1')""".format(
database_name, table_name_csv, data_s3_location
)
# -
# Execute statement using connection cursor
cursor = connect(region_name=region_name, s3_staging_dir=s3_staging_dir).cursor()
cursor.execute(statement)
# +
# verify the table has been created
statement = "SHOW TABLES in {}".format(database_name)
cursor.execute(statement)
df_show = as_pandas(cursor)
df_show.head(5)
# -
# run a sample query
statement = """SELECT * FROM {}.{}
LIMIT 100""".format(
database_name, table_name_csv
)
# Execute statement using connection cursor
cursor = connect(region_name=region_name, s3_staging_dir=s3_staging_dir).cursor()
cursor.execute(statement)
df = as_pandas(cursor)
df.head(5)
# ## Alternatives: Use AWS Data Wrangler to query data
import awswrangler as wr
# #### Glue Catalog
for table in wr.catalog.get_tables(database=database_name):
print(table["Name"])
# #### Athena
# %%time
df = wr.athena.read_sql_query(
sql="SELECT * FROM {} LIMIT 100".format(table_name_csv), database=database_name
)
df.head(5)
# ### Citation
#
# Data Science On AWS workshops, <NAME>, <NAME>, https://www.datascienceonaws.com/
| ingest_data/ingest-with-aws-services/ingest_data_with_Athena.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nlp-dl-env]
# language: python
# name: conda-env-nlp-dl-env-py
# ---
from hyperopt import hp
from hyperopt import fmin, tpe, space_eval
# ## Non-hierarchical params
# Define an objective function
def objective(params):
# Here you train your model and return some metric, e.g. accuracy
x = params['x']
y = params['y']
return (x - y)**2
# +
# Define a search space
space = {
'x': hp.uniform('x', -10, 1),
'y': hp.uniform('y', -10, 1)
}
# Minimize the objective over the space
best = fmin(objective, space, algo=tpe.suggest, max_evals=1000)
# -
best
# In the example below we define two continuous uniform (`hp.uniform`) variables `x` and `y`.
#
# To understand how to define other types of variables (`hp.loguniform`, `hp.quniform`, `hp.choice`, ...) please refer to:
#
# http://hyperopt.github.io/hyperopt/getting-started/search_spaces/
# ## Hierarchical params
# Define an objective for hierarchical params
def obj_hier(params):
print(params) # Check print output to understand the `params` staructure
return 0
# +
# Define a hierarchical search space
space_hier = hp.choice('case',
[
('case_1', {
'loss_func': hp.choice('loss_func_1', ['hinge']),
'penalty': hp.choice('penalty_1', ['l1', 'l2'])
}),
('case_2', {
'loss_func': hp.choice('loss_func_2', ['hinge', 'hinge2']),
'penalty': hp.choice('penalty_2', ['l1', 'l2', 'l1000', 'l13'])
})
])
# Minimize the objective over the space
best = fmin(obj_hier, space_hier, algo=tpe.suggest, max_evals=10)
# -
best
| [IMPL]_Hyperopt_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center>Multiscale Geographically Weighted Regression - Poisson dependent variable</center>
#
#
# The model has been explored and tested for multiple parameters on real and simulated datasets. The research includes the following outline with separate notebooks for each part.
#
#
# **Notebook Outline:**
#
# **Introduction (current)**
# - [Introduction](#Introduction)
# - [Introduction to the problem](#Introduction-to-the-project)
# - [Statistical Equations](#Statistical-Equations)
# - [Approaches Explored](#Approaches-Explored)
# - [Other notebooks](#Other-notebooks)
# - [References](#References)
# [Back to the main page](https://mehak-sachdeva.github.io/MGWR_book/)
# ---
#
# # Introduction
#
# ***
#
# ## Introduction to the problem
#
# A recent addition to the local statistical models in PySAL is the implementation of Multiscale Geographically Weighted Regression (MGWR) model, a multiscale extension to the widely used approach for modeling process spatial heterogeneity - Geographically Weighted Regression (GWR). GWR is a local spatial multivariate statistical modeling technique embedded within the regression framework that is calibrated and estimates covariate parameters at each location using borrowed data from neighboring observations. The extent of neighboring observations used for calibration is interpreted as the indicator of scale for the spatial processes and is assumed to be constant across covariates in GWR. MGWR, using a back-fitting algorithm relaxes the assumption that all processes being modeled operate at the same spatial scale and estimates a unique indicator of scale for each process.
# The GWR model in PySAL can currently estimate Gaussian, Poisson and Logistic models though the MGWR model is currently limited to only Gaussian models. This project aims to expand the MGWR model to nonlinear local spatial regression modeling techniques where the response outcomes may be discrete (following a Poisson distribution). This will enable a richer and holistic local statistical modeling framework to model multi-scale process heterogeneity for the open source community.
#
# ## Statistical Equations
#
# A conventional Poisson regression model is written as:
#
# \begin{align}
# O_i {\sim} Poisson[E_i exp ({\sum} {\beta} & _k x _{k,i})] \\
# \end{align}
#
# where $x_{k,1}$ is the kth explanatory variable in place i and the ${\beta}_ks$ are the parameters and Poisson indicates a Poisson distribution with mean $\lambda$.
#
# Nakaya et.al. (2005) introduced the concept of allowing parameter values to vary with geographical location ($u_i$), which is a vector of two dimensional co-ordinates describing the location *i*. The Poisson model for geographically varying parameters can be written as:
#
# \begin{align}
# O_i {\sim} Poisson[E_i exp ({\sum} {\beta} & _k (u_i) x _{k,i})] \\
# \end{align}
#
# The Geographically Weighted Poisson Regression model (GWPR) is estimated using a modified local Fisher scoring procedure, a form of iteratively reweighted least squares (IRLS). In this procedure, the following matrix computation of weighted least squares should be repeated to update parameter estimates until they converge (Nakaya et.al., 2005):
#
# \begin{align}
# \beta^{(l+1)} (u_i) = (X^{t} W (u_i) A(u_i)^{(l)} X)^{-1} X^{t} W (u_i) A (u_i) ^{(l)} z (u_i){(l)} \\
# \end{align}
# # Approaches Explored
# **Expected theoretical model calibration:**<br><br>
# The calibration methodology for modeling response variables with a Poisson distribution in MGWR, through references from Geographically Weighted Poisson Regression (GWPR) (Nakaya, et al., 2005) and literature on Generalized Additive Models (Hastie & Tibshirani, 1986), is expected to be as follows:<br>
#
# 1. Initialize using GWPR estimates $𝛽_𝑘(𝑢_𝑖)^0: f^1_0,f^2_0, … , f^𝐾_0, $
# where $f_𝑘^0 = (𝑥_{1 𝑘}𝛽_𝑘(𝑢_1)^0 … 𝑥_{𝑁𝑘}𝛽_𝑘(𝑢_𝑁)^0)$<br><br>
# 2. Update for each location ($𝑢_𝑖$) an adjusted dependent variable: <br>
# $z(𝑢_𝑖)^{(𝑙)} = (z_1(𝑢_𝑖)^{(𝑙)}, z_2(𝑢_𝑖)^{(𝑙)}, … ,z_𝑁(𝑢_𝑖)^{(𝑙)})^𝑡$<br>
# $z_𝑗(𝑢_𝑖)^{(𝑙)} = Σ𝛽_𝑘(𝑢_𝑖)^{(𝑙)} 𝑥_{𝑗,𝑘} + O_𝑗 − Ô_𝑗(𝛽(𝑢_𝑖)^{(𝑙)}) / Ô_𝑗(𝛽(𝑢_𝑖)^{(𝑙)})$<br><br>
# 3. Construct weights as follows:
# $A(𝑢_𝑖)^{(𝑙)}$ which is a diagonal matrix with values ($Ô_1(𝛽(
# 𝑢_𝑖)^{(𝑙)}), Ô_2(𝛽(
# 𝑢_𝑖)^{(𝑙)}), ... ,Ô_𝑁(𝛽(𝑢_𝑖)^{(𝑙)}))$<br><br>
# 4. Fit an MGWR model to $z(𝑢_𝑖)^{(𝑙)}$ to update $𝛽_𝑘(𝑢_𝑖)$ and $z(𝑢_𝑖)^{(𝑙)}$, using the new weight matrix:<br>
# $𝑊_𝑘∗(𝑢_𝑖)^{(𝑙)}=𝑊_𝑘(𝑢_𝑖)^{(𝑙)} A(𝑢_𝑖)^{(𝑙)}$<br><br>
# 5. Repeat steps (2) through (4) until convergence.
# # Other notebooks
# Below are links to the tests and exploration for the finalized MGWR model for Poisson dependent variables:
# ## Tests with real and simulated data
# ***
# **[Initial module changes and univariate model check ](http://mehak-sachdeva.github.io/MGWR_book/Html/Poisson_MGWR_univariate_check)**
# - Setup with libraries
# - Fundamental equations for Poisson MGWR
# - Example Dataset
# - Helper functions
# - Univariate example
# - Parameter check
# - Bandwidths check
#
# **[Simulated Data example](http://mehak-sachdeva.github.io/MGWR_book/Html/Simulated_data_example_Poisson-MGWR)**
# - Setup with libraries
# - Create Simulated Dataset
# - Forming independent variables
# - Creating y variable with Poisson distribution
# - Univariate example
# - Bandwidth: Random initialization check
# - Parameters check
# - Multivariate example
# - Bandwidths: Random initialization check
# - Parameters check
# - Global model parameter check
#
# **[Real Data example](http://mehak-sachdeva.github.io/MGWR_book/Html/Real_data_example_Poisson-MGWR)**
#
# - Setup with libraries
# - Tokyo Mortality Dataset
# - Univariate example
# - Bandwidth: Random initialization check
# - Parameter check
# - Multivariate example
# Bandwidths: Random initialization check
# - MGWR bandwidths
# - AIC, AICc, BIC check
#
# ## Monte Carlo Test
# ***
#
# **[Monte Carlo Simulation Visualization](http://mehak-sachdeva.github.io/MGWR_book/Html/Poisson_MGWR_MonteCarlo_Results)**
#
# - Setup with libraries
# - List bandwidths from pickles
# - Parameter functions
# - GWR bandwidth
# - MGWR bandwidths
# - AIC, AICc, BIC check
# - AIC, AICc, BIC Boxplots for comparison
# - Parameter comparison from MGWR and GWR
# # References
# 1. <NAME>., <NAME>., & <NAME>. (2017). Multiscale Geographically Weighted Regression (MGWR). Annals of the American Association of Geographers, 107(6), 1247–1265. https://doi.org/10.1080/24694452.2017.1352480
#
#
# 2. <NAME>., <NAME>., <NAME>., & <NAME>. (2005). Geographically weighted Poisson regression for disease association mapping. Statistics in Medicine, 24(17), 2695–2717. https://doi.org/10.1002/sim.2129
#
#
# 3. <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2019). Inference in Multiscale Geographically Weighted Regression. Geographical Analysis, gean.12189. https://doi.org/10.1111/gean.12189
#
#
# 4. <NAME>., & <NAME>. (1986). Generalized Additive Models. Statistical Science, 1(3), 297–310. https://doi.org/10.1214/ss/1177013604
#
#
# <NAME>. (2006). Generalized additive models : an introduction with <NAME> & Hall/CRC.
# [Back to the main page](https://mehak-sachdeva.github.io/MGWR_book/)
| Notebooks/.ipynb_checkpoints/Poisson_main-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Keras tutorial
# Tutorial from [here](https://keras.io/examples/vision/mnist_convnet/).
import numpy as np
from tensorflow import keras
from tensorflow.keras import layers
# ## Load data
# +
num_classes = 10
input_shape = (28,28,1)
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
# Scale images to [0,1] range
x_train = x_train.astype("float32") / 255
x_test = x_test.astype("float32") / 255
# -
x_train.shape, y_train.shape
# +
# Change input shape
x_train = np.expand_dims(x_train, -1)
x_test = np.expand_dims(x_test, -1)
x_train.shape, x_test.shape
# -
y_test
# Encode classes as vectors
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
y_train
# +
## Model
model = keras.Sequential(
[
keras.Input(shape=input_shape),
layers.Conv2D(32, kernel_size=(3,3), activation="relu"),
layers.MaxPooling2D(pool_size=(2,2)),
layers.Conv2D(64, kernel_size=(3,3), activation="relu"),
layers.MaxPooling2D(pool_size=(2,2)),
layers.Flatten(),
layers.Dropout(0.5),
layers.Dense(num_classes,activation="softmax")
]
)
# -
model.summary()
# +
batch_size = 128
epochs = 15
model.compile(loss="categorical_crossentropy", optimizer="adam", metrics=["accuracy"])
model.fit(x_train, y_train, batch_size=batch_size, epochs=epochs, validation_split=0.1)
# +
score = model.evaluate(x_test, y_test, verbose=0)
print("Test loss:", score[0])
print("Test accuracy:", score[1])
# -
| mnist/Keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ravi-prakash1907/Machine-Learning-for-Cyber-Security/blob/main/Spam%20Filtering/Emails/main.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="6oISEkl1eBit"
# ## Spam EMail Classification using SVM
# + id="GN3hgJc4epGJ"
#Import Library
from sklearn import svm
import pandas as pd
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import confusion_matrix
import seaborn as sns
import matplotlib.pyplot as plt
import requests
import warnings
warnings.filterwarnings('ignore')
# + id="uGvjRR2oN8Dm"
## fun. to download the data from any url
def downloadData(fileURL, saveAs='downloaded'):
req = requests.get(fileURL)
fileURLContent = req.content
data = open(saveAs, 'wb')
data.write(fileURLContent)
data.close()
# + id="JETIcQYKN-wt"
## location of the file
testURL = 'https://raw.githubusercontent.com/ravi-prakash1907/Machine-Learning-for-Cyber-Security/main/Spam%20Filtering/Emails/dataset/test.csv?token=<KEY> <PASSWORD>'
trainURL = 'https://raw.githubusercontent.com/ravi-prakash1907/Machine-Learning-for-Cyber-Security/main/Spam%20Filtering/Emails/dataset/train.csv?token=<KEY>'
## downloading the from url
downloadData(testURL,'test.csv')
downloadData(trainURL,'train.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 246} id="Evxw8cfqeAmo" outputId="c1ef9694-4f63-49a3-c1d9-6be030a5479a"
#Load Train and Test datasets
#Identify feature and response variable(s) and values must be numeric and numpy arrays
train=pd.read_csv('train.csv')
train.head()
# + colab={"base_uri": "https://localhost:8080/"} id="Rr0b3Rbcg6WQ" outputId="b11f40cf-9600-4ca5-d896-ad929052e1f5"
train.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 336} id="_XNKpMp5g8AW" outputId="5d5f1876-8486-4f68-b78c-20ff26044935"
train.describe()
# + id="ZY5cd2_rXg5j" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="91571121-8ac2-4ce4-8a7d-517c6bf5ee74"
zeros = train[train['Prediction'] == 0]
ones = train[train['Prediction'] == 1]
x0 = zeros.index
y0 = zeros['Prediction']
x1 = ones.index
y1 = ones['Prediction']
plt.scatter(x0, y0, s=10, c='b', marker="s", label='first')
plt.scatter(x1,y1, s=10, c='r', marker="o", label='second')
plt.legend(loc='upper left');
plt.show()
# + id="5XzOfdKbe2dt"
train_y=train['Prediction'] ## predictions
train_x=train.drop(['Prediction'],axis=1) ## dependent var
# + colab={"base_uri": "https://localhost:8080/", "height": 246} id="cGdXmopee6NC" outputId="ab1a2d04-a9b8-48f5-dba6-c6454353629f"
test=pd.read_csv('test.csv')
test.head()
# + id="ETiPe-kafQ6T"
test_y=test['Prediction']
test_x=test.drop(['Prediction'],axis=1)
# + id="hgSqJPS6fzVb"
# Create Linear SVM object
support = svm.LinearSVC(random_state=20)
# + colab={"base_uri": "https://localhost:8080/"} id="5cbXUFquf6En" outputId="6f5c44b1-9b2b-4114-e938-5b587c084f84"
# Train the model using the training sets and check score on test dataset
support.fit(train_x, train_y)
# + colab={"base_uri": "https://localhost:8080/"} id="OYUnClQegRB_" outputId="b1f127e1-4111-429b-cfbf-0d87c0c4faec"
predicted= support.predict(test_x)
predicted[:10]
# + colab={"base_uri": "https://localhost:8080/"} id="kcL4VSLtghDM" outputId="f9ad311e-c9dc-43ea-d6aa-0953a6ce1457"
given = test_y[:10]
list(given)
# + id="wm6VTwhegZda"
f1Score=f1_score(test_y,predicted)
accuracy=accuracy_score(test_y,predicted)
precision=precision_score(test_y,predicted)
recall=recall_score(test_y,predicted)
# + colab={"base_uri": "https://localhost:8080/"} id="oXtsD8OYgnNw" outputId="6a963a3c-0862-42b9-ca17-fe426d116938"
print("Model's Performance Measures:\n")
print("F1-Score: {} \nAccuracy: {} \nPrecision: {} \nRecall: {}".format(f1Score,accuracy,precision,recall))
# + id="A149Ruh_yIa6" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="0fce22ec-f518-4325-ea51-34d6a093be43"
cf_matrix = confusion_matrix(test_y,predicted)
print("Confusion matrix: \n", cf_matrix, "\n")
sns.heatmap(cf_matrix, annot=True)
# + id="1AJjMVg_grM_"
train.to_csv( "pred.csv")
| Spam Filtering/Emails/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
plt.style.use('ggplot')
import scipy.stats as st
# +
## download data
#import requests
#payload='target=GHO/MH_12&profile=crosstable&filter=COUNTRY:*;REGION:*&x-sideaxis=COUNTRY&x-topaxis=GHO;YEAR;SEX'
#suicide_rate_url='http://apps.who.int/gho/athena/data/xmart.csv?'
#with open('who_suicide_rates.csv', 'wb') as fout:
# fout.write(requests.get(suicide_rate_url+payload).content)
# -
rates = pd.read_csv('who_suicide_rates.csv', names=['country', 'both', 'male', 'female'], skiprows=3)
rates.head(10)
rates.plot.hist(stacked=True, y=['male', 'female'], bins=30)
print rates['male'].mean(), rates['female'].mean()
print rates[rates['both'] > 40]
_ = rates.boxplot()
def plot_cdf(data, plot_range=None, scale_to=None, **kwargs):
num_bins = len(data)
sorted_data = np.array(sorted(data), dtype=np.float64)
data_range = sorted_data[-1] - sorted_data[0]
counts, bin_edges = np.histogram(sorted_data, bins=num_bins)
xvalues = bin_edges[1:]
yvalues = np.cumsum(counts)
if plot_range is None:
xmin = sorted_data[0]
xmax = sorted_data[-1]
else:
xmin, xmax = plot_range
#pad the arrays
xvalues = np.concatenate([[xmin, xvalues[0]], xvalues, [xmax]])
yvalues = np.concatenate([[0.0, 0.0], yvalues, [yvalues.max()]])
if scale_to is not None:
yvalues = yvalues / len(data) * scale_to
#print xvalues.shape, yvalues.shape
return plt.plot(xvalues, yvalues, **kwargs)
plt.figure(figsize=(9, 3))
plt.subplot(1, 2, 1)
plot_cdf(rates['both'])
plt.title('cdf')
plt.subplot(1, 2, 2)
_ = st.probplot(rates['both'], dist='norm', plot=plt)
beta, loc, eta = st.weibull_min.fit(rates['both'], floc=0)
print beta, loc, eta
rvWb = st.weibull_min(beta, scale=eta)
plt.figure(figsize=(9, 3))
plt.subplot(1, 3, 1)
st.probplot(rates['both'], dist=rvWb, plot=plt)
plt.subplot(1, 3, 2)
rates['both'].hist(bins=30)
plt.hist(rvWb.rvs(len(rates['both'])), bins=30, alpha=0.5)
plt.title('hist')
plt.subplot(1, 3, 3)
plot_cdf(rates['both'])
plot_cdf(rvWb.rvs(len(rates['both'])))
plt.title('cdf')
# test the assumption that the amount of sunlight in each country is directly proportional to the latitude
# “A simple countries centroid can be found on the Gothos web page: http://gothos.info/2009/02/centroids-for-countries/”
coords = pd.read_csv('country_centroids/country_centroids_primary.csv', sep='\t')
print coords.keys()
print coords.head()
rates['lat'] = ''
rates['lng'] = ''
for i in coords.index:
ind = rates['country'].isin([coords['SHORT_NAME'][i]])
val = coords.loc[i, ['LAT', 'LONG']].values.astype('float')
rates.loc[ind, ['lat', 'lng']] = list(val)
rates.loc[rates['lat'].isin(['']), ['lat']] = np.nan
rates.loc[rates['lng'].isin(['']), ['lng']] = np.nan
print rates[['lat', 'lng']].describe()
rates[['lat', 'lng']] = rates[['lat', 'lng']].astype('float')
#rates.head()
rates['dfe'] = np.abs(rates['lat'])
rates.head()
# countries within +/-23.5 degrees away from the equator get an equal amount of
# sunlight throughout the year, so they should be considered to have the same
# suicide rate according to our hypothesis
plt.plot(rates['dfe'], rates['both'], 'b.')
plt.fill_between([0, 23.5], [100, 100], alpha=0.5, color='yellow')
plt.ylim(0, rates['both'].max()*1.1)
plt.xlabel('dfe')
plt.ylabel('both')
plt.plot(rates['dfe'], rates['both'], 'b.')
plt.fill_between([0, 23.5], [100, 100], alpha=0.5, color='yellow')
plt.ylim(0, rates['both'].max()*1.1)
plt.xlabel('dfe')
plt.ylabel('both')
bins = np.arange(23.5, rates['dfe'].max()+1, 10, dtype='float')
grpRates = rates.groupby(np.digitize(rates['dfe'], bins))
plt.errorbar(grpRates.mean()['dfe'], grpRates.mean()['both'], yerr=grpRates.std()['both'])
# +
from sklearn.linear_model import LinearRegression
sel = ~rates['dfe'].isnull() * rates['dfe'] > 23.5
mat = rates[sel].as_matrix(columns=['dfe', 'both'])
x = mat[:, 0].reshape(-1, 1)
y = mat[:, 1].reshape(-1, 1)
model = LinearRegression().fit(x, y)
xx = np.linspace(23.5, rates['dfe'].max()*1.1, 200).reshape(-1, 1)
plt.plot(xx, model.predict(xx), '--', lw=3)
plt.plot(rates['dfe'], rates['both'], 'b.')
plt.fill_between([0, 23.5], [100, 100], alpha=0.5, color='yellow')
plt.ylim(0, rates['both'].max()*1.1)
plt.xlabel('dfe')
plt.ylabel('both')
bins = np.arange(23.5, rates['dfe'].max()+1, 10, dtype='float')
grpRates = rates.groupby(np.digitize(rates['dfe'], bins))
plt.errorbar(grpRates.mean()['dfe'], grpRates.mean()['both'], yerr=grpRates.std()['both'])
# -
from pandas.io import wb
# wb.search('gdp.*capita.*').iloc[:, :2]
dat = wb.download(indicator='NY.GDP.PCAP.PP.CD', country='all', start=2014, end=2014)
dat.head()
countries = np.array(dat.index.tolist())[:, 0]
gdp = np.array(dat['NY.GDP.PCAP.PP.CD'])
data = pd.DataFrame(data=np.hstack((countries.reshape(-1, 1), gdp.reshape(-1, 1))), columns=['countries', 'gdp'])
data.head()
rates['gdp'] = ''
for i in np.arange(len(data)):
ind = rates['country'].isin([data['countries'][i]])
val = data.loc[i, ['gdp']].values.astype('float')
rates.loc[ind, ['gdp']] = val
rates.loc[rates['gdp'].isin(['']), ['gdp']] = np.nan
print rates[rates['country'] == 'Sweden']
print data[data['countries'] == 'Sweden']
sel = (~rates['dfe'].isnull()) * (rates['dfe'] > 23.5)
sel *= (~rates['gdp'].isnull())
plt.plot(rates[sel]['gdp'], rates[sel]['both'], '.', ms=10)
plt.xlabel('gdp')
plt.ylabel('suicide rate')
plt.scatter(rates[sel]['gdp']/1000, rates[sel]['dfe'], s=rates[sel]['both']**1.5)
plt.xlabel('gdp/1000')
plt.ylabel('dfe')
from mpl_toolkits.mplot3d import Axes3D
mat = rates[sel].as_matrix(columns=['dfe', 'gdp'])
mat[:, 1] /= 1000
z = rates[sel].as_matrix(columns=['both'])
model = LinearRegression().fit(mat, z)
xx, yy = np.meshgrid(np.linspace(mat[:, 0].min(), mat[:, 0].max(), 100),
np.linspace(mat[:, 1].min(), mat[:, 1].max(), 100))
z2 = model.predict(np.hstack((xx.reshape(-1, 1), yy.reshape(-1, 1)))).reshape(100, 100)
# z2 = xx*model.coef_[0][0] + yy*model.coef_[0][1] + model.intercept_[0]
# print z2.shape, model.coef_, model.intercept_
fig = plt.figure(figsize=(12, 8))
ax = Axes3D(fig, azim=-135, elev=15)
surf = ax.plot_surface(xx, yy, z2, cmap=plt.cm.RdBu, alpha=0.6, linewidth=0)
ax.scatter(mat[:, 0], mat[:, 1], z)
ax.set_xlabel('dfe')
ax.set_ylabel('gdp')
ax.set_zlabel('both')
# +
# data = pd.DataFrame(rates[['country', 'both', 'male', 'female', 'dfe', 'gdp']])[~rates['dfe'].isnull()]
# data.to_hdf('ch4data.h5', 'ch4data', mode='w', table=True, encoding='utf8')
# data.head()
# +
# d2 = pd.read_hdf('ch4data.h5')
# d2.head()
# -
k, m = 1.0, -5.0
y = lambda x: k*x + m
p = lambda x: 1.0 / (1 + np.exp(-(k*x + m)))
xx = np.linspace(0, 10, 100)
plt.plot(xx, y(xx), label='linear')
plt.plot(xx, p(xx), label='logistic')
plt.plot([0, abs(m)], [0.5, 0.5], '--', lw=3, color='.7')
plt.plot([abs(m), abs(m)], [-0.5, 0.5], '--', lw=3, color='.7')
plt.ylim(-0.5, 1.5)
plt.legend()
studytime=[0,0,1.5,2,2.5,3,3.5,4,4,4,5.5,6,6.5,7,7,8.5,9,9,9,10.5,10.5,12,12,12,12.5,13,14,15,16,18]
passed=[0,0,0,0,0,0,0,0,0,0,0,1,0,1,1,0,1,1,0,1,1,1,1,1,1,1,1,1,1,1]
data = pd.DataFrame(data=np.array([studytime, passed]).T, columns=['time', 'passed'])
plt.figure(figsize=(9, 4))
plt.subplot(1, 2, 1)
data['time'].hist(bins=6)
plt.xlabel('time')
plt.ylabel('# students')
plt.subplot(1, 2, 2)
plt.plot(data['time'], data['passed'], 'o', ms=7)
plt.xlabel('time')
plt.ylabel('passed or not')
plt.ylim(-0.1, 1.1)
plt.xlim(-2, 20)
# +
from sklearn.linear_model import LogisticRegression
model = LogisticRegression().fit(data['time'].values.reshape(-1, 1), data['passed'].values.reshape(-1, 1))
xx = np.linspace(0, data.time.max()).reshape(-1, 1)
k, m = model.coef_[0][0], model.intercept_[0]
print k, m
y = lambda x: 1. / (1 + np.exp(-1*(k*x+m)))
plt.plot(xx, y(xx), lw=2, label='fit')
plt.plot(xx, model.decision_function(xx))
plt.plot(data['time'], data['passed'], 'o', ms=7)
plt.xlabel('time')
plt.ylabel('passed or not')
plt.ylim(-0.1, 1.1)
plt.xlim(-2, 20)
# +
import statsmodels.api as sm
model = sm.Logit(data.passed, sm.add_constant(data.time, prepend=True))
fit_result = model.fit()
print fit_result.summary()
print fit_result.params
intercept, slope = fit_result.params.const, fit_result.params.time
intercept_err, slope_err = np.diag(fit_result.cov_params()) ** 0.5
y = lambda x, k, m: 1. / (1 + np.exp(-(k*x+m)))
xx = np.linspace(0, data.time.max())
l1 = plt.plot(xx, y(xx, slope, intercept), lw=2, label='fit')
plt.fill_between(xx, y(xx, slope+slope_err**2, intercept+intercept_err),
y(xx, slope-slope_err**2, intercept-intercept_err), alpha=0.15, color=l1[0].get_color())
plt.plot(data['time'], data['passed'], 'o', ms=7, label='data')
plt.xlabel('time')
plt.ylabel('passed or not')
plt.ylim(-0.1, 1.1)
plt.xlim(-2, 20)
plt.legend(loc=2)
# -
| jupyterNotebooks/LinearRegression/LinearRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FloPy
#
# ## Basic plotting of MODFLOW 6 Results for a single GWF model
#
# This notebook demonstrates use of a MODFLOW 6 binary grid (grd) file to plot structured and unstructured models and use of the `plot_cvfd()` and `contour_array_cvfd()` mapping methods in flopy.
# +
import os
import sys
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
# -
workspace = os.path.join('..', 'data', 'mfgrd_test')
# ### Single layer structured model
fn = os.path.join(workspace, 'nwtp3.dis.grb')
grd = flopy.utils.MfGrdFile(fn, verbose=True)
iverts, verts = grd.get_verts()
mg = grd.get_modelgrid()
extents = mg.extent
vertc = grd.get_centroids()
print(vertc[0, :])
print(len(iverts))
print(iverts[0])
print(verts.shape)
print(verts[0:5,:])
print(mg.extent)
fn = os.path.join(workspace, 'nwtp3.hds.cmp')
ho = flopy.utils.HeadFile(fn)
times = ho.get_times()
print(times)
h = ho.get_data(totim=times[-1]).reshape(80*80)
print(h.shape)
mm = flopy.plot.PlotMapView(modelgrid=mg, layer=0)
ax = plt.gca()
ax.set_xlim(extents[:2])
ax.set_ylim(extents[2:])
mm.plot_cvfd(verts, iverts, a=h)
cs = mm.contour_array_cvfd(vertc, h, colors='white')
plt.clabel(cs, fmt='%.1f', colors='white', fontsize=11);
# ### Multi-layer structured model
fn = os.path.join(workspace, 'uzfp3_lakmvr_v2.dis.grb')
grd = flopy.utils.MfGrdFile(fn, verbose=True)
iverts, verts = grd.get_verts()
mg = grd.get_modelgrid()
extents = mg.extent
vertc = grd.get_centroids()
fn = os.path.join(workspace, 'uzfp3_lakmvr.hds.cmp')
ho = flopy.utils.HeadFile(fn)
times = ho.get_times()
print(times)
h = ho.get_data(totim=times[-1]).reshape(2*15*10)
print(h.shape)
print(vertc.shape)
mm = flopy.plot.PlotMapView(modelgrid=mg, layer=0)
ax = plt.gca()
ax.set_xlim(extents[:2])
ax.set_ylim(extents[2:])
v = mm.plot_cvfd(verts, iverts, a=h, ncpl=150, masked_values=[6999.], cmap='viridis')
cs = mm.contour_array_cvfd(vertc, h, ncpl=150, masked_values=[6999.],
levels=[1024], colors='white')
plt.clabel(cs, fmt='%.0f', colors='white', fontsize=11)
plt.colorbar(v, shrink=0.5);
vertc = grd.get_centroids()
mm = flopy.plot.PlotMapView(modelgrid=mg, layer=1)
ax = plt.gca()
ax.set_xlim(extents[:2])
ax.set_ylim(extents[2:])
v = mm.plot_cvfd(verts, iverts, a=h, ncpl=150, masked_values=[6999.], cmap='viridis')
cs = mm.contour_array_cvfd(vertc, h, ncpl=150, masked_values=[6999.],
levels=[1020, 1021, 1022, 1023, 1024, 1025, 1026], colors='white')
plt.clabel(cs, fmt='%.0f', colors='white', fontsize=11)
plt.colorbar(v, shrink=0.5);
# ### Single-layer unstructured model
fn = os.path.join(workspace, 'flow.disv.grb')
grd = flopy.utils.MfGrdFile(fn, verbose=True)
iverts, verts = grd.get_verts()
vertc = grd.get_centroids()
mg = grd.get_modelgrid()
fn = os.path.join(workspace, 'flow.hds.cmp')
ho = flopy.utils.HeadFile(fn)
times = ho.get_times()
print(times)
h = ho.get_data(totim=times[-1]).reshape(218)
print(h.shape)
mm = flopy.plot.PlotMapView(modelgrid=mg)
ax = plt.gca()
ax.set_xlim(0,700)
ax.set_ylim(0,700)
v = mm.plot_cvfd(verts, iverts, edgecolor='black', a=h)
cs = mm.contour_array_cvfd(vertc, h, colors='white')
plt.clabel(cs, fmt='%.1f', colors='white', fontsize=11)
plt.colorbar(v, shrink=0.5);
# ### Single-layer unstructured model with xt3d
fn = os.path.join(workspace, 'flowxt3d.disv.grb')
grd = flopy.utils.MfGrdFile(fn, verbose=True)
iverts, verts = grd.get_verts()
vertc = grd.get_centroids()
mg = grd.get_modelgrid()
fn = os.path.join(workspace, 'flowxt3d.hds.cmp')
ho = flopy.utils.HeadFile(fn, precision='double')
times = ho.get_times()
print(times)
h = ho.get_data(totim=times[-1]).reshape(218)
print(h.shape)
mm = flopy.plot.PlotMapView(modelgrid=mg)
ax = plt.gca()
ax.set_xlim(0,700)
ax.set_ylim(0,700)
v = mm.plot_cvfd(verts, iverts, edgecolor='black', a=h)
cs = mm.contour_array_cvfd(vertc, h, colors='white')
plt.clabel(cs, fmt='%.1f', colors='white', fontsize=11)
plt.colorbar(v, shrink=0.5);
# ### Single-layer unstructured model with a well
fn = os.path.join(workspace, 'flowwel.disv.grb')
grd = flopy.utils.MfGrdFile(fn, verbose=True)
iverts, verts = grd.get_verts()
vertc = grd.get_centroids()
mg = grd.get_modelgrid()
fn = os.path.join(workspace, 'flowwel.hds.cmp')
ho = flopy.utils.HeadFile(fn, precision='double')
times = ho.get_times()
print(times)
h = ho.get_data(totim=times[-1]).reshape(218)
print(h.shape)
mm = flopy.plot.PlotMapView(modelgrid=mg)
ax = plt.gca()
ax.set_xlim(0,700)
ax.set_ylim(0,700)
v = mm.plot_cvfd(verts, iverts, edgecolor='black', a=h)
cs = mm.contour_array_cvfd(vertc, h, colors='white')
plt.clabel(cs, fmt='%.1f', colors='white', fontsize=11)
plt.colorbar(v, shrink=0.5);
# ### Single-layer unstructured model with a well and xt3d
fn = os.path.join(workspace, 'flowwelxt3d.disv.grb')
grd = flopy.utils.MfGrdFile(fn, verbose=True)
iverts, verts = grd.get_verts()
vertc = grd.get_centroids()
mg = grd.get_modelgrid()
fn = os.path.join(workspace, 'flowwelxt3d.hds.cmp')
ho = flopy.utils.HeadFile(fn, precision='double')
times = ho.get_times()
print(times)
h2 = ho.get_data(totim=times[-1]).reshape(218)
print(h2.shape)
mm = flopy.plot.PlotMapView(modelgrid=mg)
ax = plt.gca()
ax.set_xlim(0,700)
ax.set_ylim(0,700)
v = mm.plot_cvfd(verts, iverts, edgecolor='black', a=h2, cmap='jet_r')
cs = mm.contour_array_cvfd(vertc, h2, colors='white', linewidths=2)
cs = mm.contour_array_cvfd(vertc, h, linestyles='dashed', colors='cyan', linewidths=2)
plt.clabel(cs, fmt='%.1f', colors='white', fontsize=11)
plt.colorbar(v, shrink=0.5);
d = h - h2
mm = flopy.plot.PlotMapView(modelgrid=mg)
ax = plt.gca()
ax.set_xlim(0,700)
ax.set_ylim(0,700)
v = mm.plot_cvfd(verts, iverts, edgecolor='black', a=d, cmap='jet_r')
plt.colorbar(v, shrink=0.5);
# ### Single-layer nested model with a well
fn = os.path.join(workspace, 'flowquadwel.disv.grb')
grd = flopy.utils.MfGrdFile(fn, verbose=True)
iverts, verts = grd.get_verts()
vertc = grd.get_centroids()
mg = grd.get_modelgrid()
fn = os.path.join(workspace, 'flowquadwel.hds.cmp')
ho = flopy.utils.HeadFile(fn, precision='double')
times = ho.get_times()
print(times)
h = ho.get_data(totim=times[-1]).flatten()
print(h.shape)
mm = flopy.plot.PlotMapView(modelgrid=mg)
ax = plt.gca()
ax.set_xlim(0,700)
ax.set_ylim(0,700)
v = mm.plot_cvfd(verts, iverts, edgecolor='black', a=h)
cs = mm.contour_array_cvfd(vertc, h, colors='white')
plt.clabel(cs, fmt='%.1f', colors='white', fontsize=11)
plt.colorbar(v, shrink=0.5);
# ### Single-layer nested model with a well and xt3d
fn = os.path.join(workspace, 'flowquadwelxt3d.disv.grb')
grd = flopy.utils.MfGrdFile(fn, verbose=True)
iverts, verts = grd.get_verts()
vertc = grd.get_centroids()
mg = grd.get_modelgrid()
fn = os.path.join(workspace, 'flowquadwelxt3d.hds.cmp')
ho = flopy.utils.HeadFile(fn, precision='double')
times = ho.get_times()
print(times)
h2 = ho.get_data(totim=times[-1]).flatten()
print(h2.shape)
mm = flopy.plot.PlotMapView(modelgrid=mg)
ax = plt.gca()
ax.set_xlim(0,700)
ax.set_ylim(0,700)
v = mm.plot_cvfd(verts, iverts, edgecolor='black', a=h2)
cs = mm.contour_array_cvfd(vertc, h2, colors='white')
plt.clabel(cs, fmt='%.1f', colors='white', fontsize=11)
cs = mm.contour_array_cvfd(vertc, h, linestyles='dashed', colors='cyan', linewidths=2)
plt.colorbar(v, shrink=0.5);
d = h - h2
mm = flopy.plot.PlotMapView(modelgrid=mg)
ax = plt.gca()
ax.set_xlim(0,700)
ax.set_ylim(0,700)
v = mm.plot_cvfd(verts, iverts, edgecolor='black', a=d, cmap='jet_r')
plt.colorbar(v, shrink=0.5);
| examples/Notebooks/flopy3_mf6_BasicPlotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from nltk.tokenize import TreebankWordTokenizer
sentence = "The faster Harry got to the store, the faster Harry, the faster, would get home."
tokenizer = TreebankWordTokenizer()
token_sequence = tokenizer.tokenize(sentence.lower())
print(token_sequence)
# -
from collections import Counter
bag_of_words = Counter(token_sequence)
print(bag_of_words)
word_list = bag_of_words.most_common() # Passing an integer as an argument will give you that many from the top of the list
print(word_list)
# +
times_harry_appears = bag_of_words['harry']
total_words = len(word_list) # The number of tokens from our original source.
tf = times_harry_appears/total_words
print(tf)
# -
kite_text = "A kite is traditionally a tethered heavier-than-air craft with wing surfaces that react against the air to create lift and drag. A kite consists of wings, tethers, and anchors. Kites often have a bridle to guide the face of the kite at the correct angle so the wind can lift it. A kite's wing also may be so designed so a bridle is not needed; when kiting a sailplane for launch, the tether meets the wing at a single point. A kite may have fixed or moving anchors. Untraditionally in technical kiting, a kite consists of tether-set-coupled wing sets; even in technical kiting, though, a wing in the system is still often called the kite. The lift that sustains the kite in flight is generated when air flows around the kite's surface, producing low pressure above and high pressure below the wings. The interaction with the wind also generates horizontal drag along the direction of the wind. The resultant force vector from the lift and drag force components is opposed by the tension of one or more of the lines or tethers to which the kite is attached. The anchor point of the kite line may be static or moving (e.g., the towing of a kite by a running person, boat, free-falling anchors as in paragliders and fugitive parakites or vehicle). The same principles of fluid flow apply in liquids and kites are also used under water. A hybrid tethered craft comprising both a lighter-than-air balloon as well as a kite lifting surface is called a kytoon. Kites have a long and varied history and many different types are flown individually and at festivals worldwide. Kites may be flown for recreation, art or other practical uses. Sport kites can be flown in aerial ballet, sometimes as part of a competition. Power kites are multi-line steerable kites designed to generate large forces which can be used to power activities such as kite surfing, kite landboarding, kite fishing, kite buggying and a new trend snow kiting. Even Man-lifting kites have been made."
# +
from collections import Counter
from nltk.tokenize import TreebankWordTokenizer
tokenizer = TreebankWordTokenizer()
# kite_text = "A kite is traditionally ..." # Step left to user, so we aren't repeating ourselves
tokens = tokenizer.tokenize(kite_text.lower())
token_sequence = Counter(tokens)
print(token_sequence)
# +
import nltk
nltk.download('stopwords')
stopwords = nltk.corpus.stopwords.words('english')
tokens = [x for x in tokens if x not in stopwords]
kite_count = Counter(tokens)
print(kite_count)
# +
document_vector = []
doc_length = len(tokens)
for key, value in kite_count.most_common():
document_vector.append(value / doc_length)
print(document_vector)
# -
doc_0 = "The faster Harry got to the store, the faster Harry, the faster, would get home."
doc_1 = "Harry is hairy and faster than Jill."
doc_2 = "Jill is not as hairy as Harry."
# +
tokens_0 = tokenizer.tokenize(doc_0.lower())
tokens_1 = tokenizer.tokenize(doc_1.lower())
tokens_2 = tokenizer.tokenize(doc_2.lower())
lexicon = set(tokens_0 + tokens_1 + tokens_2)
print(lexicon)
# -
print(len(lexicon))
# +
from collections import OrderedDict
vector_template = OrderedDict((token, 0) for token in lexicon)
print(vector_template)
# +
import copy
document_vectors = []
for doc in [doc_0, doc_1, doc_2]:
vec = copy.copy(vector_template) # So we are dealing with new objects, not multiple references to the same object
tokens = tokenizer.tokenize(doc.lower())
token_counts = Counter(tokens)
for key, value in token_counts.items():
vec[key] = value / len(lexicon)
document_vectors.append(vec)
# +
import math
def cosine_sim(vec1, vec2):
"""
Since our vectors are dictionaries, lets convert them to lists for easier mathing.
"""
vec1 = [val for val in vec1.values()]
vec2 = [val for val in vec2.values()]
dot_prod = 0
for i, v in enumerate(vec1):
dot_prod += v * vec2[i]
mag_1 = math.sqrt(sum([x**2 for x in vec1]))
mag_2 = math.sqrt(sum([x**2 for x in vec2]))
return dot_prod / (mag_1 * mag_2)
# -
from nltk.corpus import brown
print(len(brown.words())) # words is a builtin method of the nltk corpus object that gives a list of tokens
# +
from collections import Counter
puncs = [',', '.', '--', '-', '!', '?', ':', ';', '``', "''", '(', ')', '[', ']']
word_list = [x.lower() for x in brown.words() if x not in puncs]
token_counts = Counter(word_list)
print(token_counts.most_common(20))
# -
history_text = 'Kites were invented in China, where materials ideal for kite building were readily available: silk fabric for sail material; fine, high-tensile-strength silk for flying line; and resilient bamboo for a strong, lightweight framework. The kite has been claimed as the invention of the 5th-century BC Chinese philosophers Mozi (also Mo Di) and Lu Ban (also Gongshu Ban). By 549 AD paper kites were certainly being flown, as it was recorded that in that year a paper kite was used as a message for a rescue mission. Ancient and medieval Chinese sources describe kites being used for measuring distances, testing the wind, lifting men, signaling, and communication for military operations. The earliest known Chinese kites were flat (not bowed) and often rectangular. Later, tailless kites incorporated a stabilizing bowline. Kites were decorated with mythological motifs and legendary figures; some were fitted with strings and whistles to make musical sounds while flying. From China, kites were introduced to Cambodia, Thailand, India, Japan, Korea and the western world. After its introduction into India, the kite further evolved into the fighter kite, known as the patang in India, where thousands are flown every year on festivals such as Makar Sankranti. Kites were known throughout Polynesia, as far as New Zealand, with the assumption being that the knowledge diffused from China along with the people. Anthropomorphic kites made from cloth and wood were used in religious ceremonies to send prayers to the gods. Polynesian kite traditions are used by anthropologists get an idea of early "primitive" Asian traditions that are believed to have at one time existed in Asia.'
# intro_text = "A kite is traditionally ..." # Step left to user, as above
intro_text = kite_text.lower()
intro_tokens = tokenizer.tokenize(intro_text)
# history_text = "Kites were invented in China, ..." # Also as above
history_text = history_text.lower()
history_tokens = tokenizer.tokenize(history_text)
intro_total = len(intro_tokens)
history_total = len(history_tokens)
intro_tf = {}
history_tf = {}
intro_counts = Counter(intro_tokens)
intro_tf['kite'] = intro_counts['kite'] / intro_total
history_counts = Counter(history_tokens)
history_tf['kite'] = history_counts['kite'] / history_total
print('Term Frequency of "kite" in intro is: {}'.format(intro_tf['kite']))
print('Term Frequency of "kite" in history is: {}'.format(history_tf['kite']))
intro_tf['and'] = intro_counts['and'] / intro_total
history_tf['and'] = history_counts['and'] / history_total
print('Term Frequency of "and" in intro is: {}'.format(intro_tf['and']))
print('Term Frequency of "and" in history is: {}'.format(history_tf['and']))
num_docs_containing_and = 0
for doc in [intro_tokens, history_tokens]:
if 'and' in doc:
num_docs_containing_and += 1
num_docs_containing_kite = 0
for doc in [intro_tokens, history_tokens]:
if 'kite' in doc:
num_docs_containing_kite += 1
num_docs_containing_china = 0
for doc in [intro_tokens, history_tokens]:
if 'china' in doc:
num_docs_containing_china += 1
intro_tf['china'] = intro_counts['china'] / intro_total
history_tf['china'] = history_counts['china'] / history_total
num_docs = 2
intro_idf = {}
history_idf = {}
intro_idf['and'] = num_docs / num_docs_containing_and
history_idf['and'] = num_docs / num_docs_containing_and
intro_idf['kite'] = num_docs / num_docs_containing_kite
history_idf['kite'] = num_docs / num_docs_containing_kite
intro_idf['china'] = num_docs / num_docs_containing_china
history_idf['china'] = num_docs / num_docs_containing_china
# +
intro_tfidf = {}
intro_tfidf['and'] = intro_tf['and'] * intro_idf['and']
intro_tfidf['kite'] = intro_tf['kite'] * intro_idf['kite']
intro_tfidf['china'] = intro_tf['china'] * intro_idf['china']
# +
history_tfidf = {}
history_tfidf['and'] = history_tf['and'] * history_idf['and']
history_tfidf['kite'] = history_tf['kite'] * history_idf['kite']
history_tfidf['china'] = history_tf['china'] * history_idf['china']
# -
document_tfidf_vectors = []
documents = [doc_0, doc_1, doc_2]
for doc in documents:
vec = copy.copy(vector_template) # So we are dealing with new objects, not multiple references to the same object
tokens = tokenizer.tokenize(doc.lower())
token_counts = Counter(tokens)
for key, value in token_counts.items():
docs_containing_key = 0
for _doc in documents:
if key in _doc:
docs_containing_key += 1
tf = value / len(lexicon)
if docs_containing_key:
idf = len(documents) / docs_containing_key
else:
idf = 0
vec[key] = tf * idf
document_tfidf_vectors.append(vec)
# +
query = "How long does it take to get to the store?"
query_vec = copy.copy(vector_template)
query_vec = copy.copy(vector_template) # So we are dealing with new objects, not multiple references to the same object
tokens = tokenizer.tokenize(query.lower())
token_counts = Counter(tokens)
for key, value in token_counts.items():
docs_containing_key = 0
for _doc in documents:
if key in _doc.lower():
docs_containing_key += 1
if docs_containing_key == 0: # We didn't find that token in the lexicon go to next key
continue
tf = value / len(tokens)
idf = len(documents) / docs_containing_key
query_vec[key] = tf * idf
print(cosine_sim(query_vec, document_tfidf_vectors[0]))
print(cosine_sim(query_vec, document_tfidf_vectors[1]))
print(cosine_sim(query_vec, document_tfidf_vectors[2]))
# +
from sklearn.feature_extraction.text import TfidfVectorizer
corpus = [doc_0, doc_1, doc_2]
vectorizer = TfidfVectorizer(min_df=1)
model = vectorizer.fit_transform(corpus)
print(model.todense()) # The model becomes a sparse numpy matrix, as in a large corpus there would be mostly zeros to deal with. todense() brings it back to a regular numpy matrix for our viewing pleasure.
| src/nlpia/book/examples/ch03_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Single Index Quantile Regression
#
# Author: <NAME> (Supervised by Prof. <NAME>)
#
# August 18th, 2019
#
# This is a tutorial on how to use the Single Index Quantile Regression model package. The packages almost identically replicates the profile optimization discussed in Ma and He (2016). See the paper as
# > https://pdfs.semanticscholar.org/9324/e31866435d446f147320f80acde682e8e614.pdf
#
# Environment: Python 3
#
# Package requirements: Numpy 1.16.3, Pandas 0.22.0, Scipy 1.3.1, Matplotlib 3.1.1.
# ### 1. Generate B-spline
# According to Ma and He (2016, p4) and de Boor (2001), the nonparametric single index function $G_{\tau}(\cdot)$ can be approximated well
# by a spline function such that $G_{\tau}(\cdot) \approx B(\cdot)^T\theta_{\tau} $, w.h. $B(\cdot)$ is the basis splines with respect to a given degree, smoothness, and domain partition.
#
# **Part I** provides a python code to generate the B-splines for any given interval and knots.The construction of B-Splines follows the Cox-de Boor recursion formula. See the description as
# > https://en.wikipedia.org/wiki/B-spline
# \begin{align}
# B_{i,1}(x) &= \left\{
# \begin{array}{rl}
# 1 & \text{if } t_i \le x < t_{i+1} \\
# 0 & \text{if otherwise}.
# \end{array} \right. \\
# B_{i,k+1}(x) &= \dfrac{x-t_i}{t_{i+k}-t_i}B_{i,k}(x)+\dfrac{t_{i+k+1}-x}{t_{i+k+1}-t_{i+1}}B_{i+1,k}(x)
# \end{align}
# The derivatives which might be used latter in the sensitivity analysis can be easily put in
# \begin{align}
# B'_{i,k+1}(x) = \dfrac{1}{t_{i+k}-t_i}B_{i,k}(x)+\dfrac{x-t_i}{t_{i+k}-t_i}B'_{i,k}(x)+\dfrac{-1}{t_{i+k+1}-t_{i+1}}B_{i+1,k}(x)+\dfrac{t_{i+k+1}-x}{t_{i+k+1}-t_{i+1}}B'_{i+1,k}(x)
# \end{align}
import numpy as np; import pandas as pd
from pandas import DataFrame
import matplotlib.pyplot as plt
from scipy.optimize import minimize, Bounds, LinearConstraint, NonlinearConstraint, BFGS
# +
def indicator(x,a,b,right_close=False):
if not right_close:
if (x >= a) & (x < b):
return 1
else:
return 0
else:
if (x >= a) & (x <= b):
return 1
else:
return 0
def I(a,b,right_close=False):
'''
return indicator function for a <= x < (or <=) b
'''
def inter(x):
return indicator(x,a,b,right_close)
return inter
def add(I1,I2):
'''
define the addition
'''
def inter(x):
return I1(x) + I2(x)
return inter
def mult(I1,I2):
'''
define the multiplication
'''
def inter(x):
return I1(x)*I2(x)
return inter
def scalar(I,alpha):
'''
define the scalar multiplication
'''
def inter(x):
return alpha*I(x)
return inter
def f(x, t_1, t_2, x_large = True):
if t_1 != t_2:
if x_large:
return (x - t_1)/(t_2 - t_1)
else:
return (t_2 - x)/(t_2 - t_1)
else:
return 0
def recur(t_1, t_2, x_large = True):
'''
return the recursion polynomial in the Cox-de Boor's algorithm
'''
def inter(x):
return f(x, t_1, t_2, x_large)
return inter
# +
def partition(a,b,N):
'''
interval [a,b] is evenly partitioned into a = t_0 < t_1 < ... < t_N < b = t_N+1
return the knots [t_0, t_1, ..., t_N+1]
'''
h = (b - a)/(N + 1)
return [a + i*h for i in range(0,N+2)]
def extend(t, p):
'''
extend the original t of length N+1 into a dictionary of length N+1+2p, convinient for de Boor algorithm
p is the final degree of the polynomials, i.e., p = m - 1 where m is the order of B-splines
'''
dic = {}
N = len(t) - 1
for i in range(p):
dic[i-p] = t[0]
for i in range(N+1):
dic[i] = t[i]
for i in range(p):
dic[N+i+1] = t[N]
return dic
# -
def deBoor(a, b, m, N, deri = False):
'''
a, b : the infimum and supremum , or minimum and maximum, of the scalar product <X, \beta>
m : the order of B-spline (>= 2)
N : the number of partition, i.e., [t0(=a), t1], [t1, t2], ... , [tN, tN+1(=b)]
deri : when True, return the derivatives. Default is False
'''
# the choice of N follow the implementation in Ma and He (2016, p9)
p = m - 1
t = partition(a,b,N)
t = extend(t,p)
if not deri:
B_k_1 = {}
for i in range(-p, N + p + 1) :
B_k_1[i] = I(t[i],t[i+1])
for k in range(1, p + 1):
B_k_0 = B_k_1
B_k_1 = {}
for i in range(-p, N + p + 1 - k):
recursion0 = mult( B_k_0[i] , recur(t[i], t[i+k], True) )
recursion1 = mult( B_k_0[i+1] , recur(t[i+1], t[i+k+1], False) )
B_k_1[i] = add(recursion0, recursion1)
return B_k_1
else:
B_k_1 = {}
b_k_1 = {}
for i in range(-p, N + p + 1) :
B_k_1[i] = I(t[i],t[i+1])
b_k_1[i] = I(0.,0.)
for k in range(1, p + 1):
B_k_0 = B_k_1
b_k_0 = b_k_1
B_k_1 = {}
b_k_1 = {}
for i in range(-p, N + p + 1 - k):
recursion0 = mult( B_k_0[i] , recur(t[i], t[i+k], True) )
recursion1 = mult( B_k_0[i+1] , recur(t[i+1], t[i+k+1], False) )
B_k_1[i] = add(recursion0, recursion1)
deri1 = mult( b_k_0[i] , recur(t[i], t[i+k], True) )
deri2 = mult( b_k_0[i+1] , recur(t[i+1], t[i+k+1], False) )
deri3 = scalar( B_k_0[i] , recur(t[i], t[i+k], True)(t[i]+1) )
deri4 = scalar( B_k_0[i+1] , recur(t[i+1], t[i+k+1], False)(t[i+k+1]+1) )
b_k_1[i] = add( add(deri1,deri2) , add(deri3,deri4) )
return B_k_1, b_k_1
# an example is provided
a, b, m, N = 0, 12, 4, 3
B_spline, b_deri = deBoor(a, b, m, N, True)
B_spline
plt.figure(figsize=(20,16))
for i in list(B_spline.keys()):
plt.subplot(3,3,i - list(B_spline.keys())[0] + 1)
X = np.arange(0,12,0.05)
Y = [B_spline[i](j) for j in X]
l = 'B(' + str(i) + ',' + str(m) + ')'
plt.plot(X,Y,label=l)
plt.legend()
plt.show()
plt.figure(figsize=(20,16))
for i in list(b_deri.keys()):
plt.subplot(3,3,i - list(b_deri.keys())[0] + 1)
X = np.arange(0,12,0.05)
Y = [b_deri[i](j) for j in X]
l = 'b(' + str(i) + ',' + str(m) + ')'
plt.plot(X,Y,label=l)
plt.legend()
plt.show()
# +
# sanity check: the sum of B-splines should be 1 over the domain
ss = lambda x : 0
for i in list(B_spline.keys()):
ss = add(ss,B_spline[i])
x = np.arange(0,12,0.05)
y = [ss(j) for j in x]
plt.figure(figsize=(4,4))
plt.plot(x,y)
plt.show()
# -
# ### 2. Determine the infimum and supremum of $x^T\beta$
# Y:
# + the log return of 000001.SZ
#
# X:
# + the log return of other main commercial banks (listed before 2011)
# + 000001.SZ specific characteristics
# + macro state variables
data1 = pd.read_csv('results/log_return.csv').dropna()
data2 = pd.read_csv('results/000001_specific.csv').dropna()
data3 = pd.read_csv('results/macro_state.csv').dropna()
X = pd.concat([data1[data1.columns[2:]], data2[data2.columns[1:]], data3[data3.columns[1:]]], axis = 1)
Y = data1[data1.columns[1]]
X = np.array(X)
Y = np.array(Y)
# sanity check
print(X.shape)
print(Y.shape)
# +
def u(x):
def inter(beta):
return np.dot(x,beta)/np.sqrt((beta**2).sum())
return inter
def v(x):
def inter(beta):
return -1. * np.dot(x,beta)/np.sqrt((beta**2).sum())
return inter
def min_max(x, min_ = True):
d = len(x)
beta0 = np.ones(d)
# define the linear constraint beta_0 > 0
ub = np.ones(d)*np.inf
lb = - np.ones(d)*np.inf
lb[0] = 0.
bou = Bounds(lb, ub)
if min_:
res = minimize(u(x), beta0, method='L-BFGS-B',bounds = bou)
else:
res = minimize(v(x), beta0, method='L-BFGS-B',bounds = bou)
return u(x)(res.x)
def inf_sup(X):
n = X.shape[0]
d = X.shape[1]
inf, sup = [], []
for i in range(n):
inf = inf + [min_max(X[i], min_ = True)]
sup = sup + [min_max(X[i], min_ = False)]
return np.array(inf).min(),np.array(sup).max()
# -
a, b = inf_sup(X)
print(a,b)
# ### 3. Define the loss function
n = X.shape[0]
m = 4
N = round(n**(1/(2*m+1))) + 1
dB = deBoor(a, b, m, N)
B = [i for i in dB.values()]
tau = 0.95
# +
def linear(B,theta):
'''
B : list of basis splines, dimension J = N + m
theta : control points of basis splines, (J,) array
'''
J = len(theta)
lin = scalar(B[0],theta[0])
for i in range(1,J):
lin = add(lin, scalar(B[i],theta[i]))
return lin
def rho(s,tau):
'''
define the pinball loss
'''
if s >= 0:
return tau*s
if s < 0:
return (tau - 1)*s
def SIQ_loss(X,Y,beta,B,theta,tau):
'''
X : sample input, (n, d) array
Y : sample output, (n,) array
beta : index, (d,) array
B : list of basis splines, dimension J = N + m
theta : control points of basis splines, (J,) array
tau : quantile to be estimated
'''
n = X.shape[0]
L = 0.
for i in range(n):
lin = linear(B, theta)
s = Y[i] - lin( u(X[i])(beta) )
L += rho(s,tau)
return L/n
# -
# ### 4. Optimization for nonparametric function $G(\cdot)$ given index $\beta$
# +
def loss_on_theta(X,Y,beta,B,tau):
def inter(theta):
return SIQ_loss(X,Y,beta,B,theta,tau)
return inter
def theta_on_others(X,Y,beta,B,tau,theta0):
J = len(B)
res = minimize(loss_on_theta(X,Y,beta,B,tau), theta0, method='BFGS')
return res.x
# -
# ### 5. Optimization for Index $\beta$
# +
def loss_on_beta_(X,Y,B,theta,tau):
def inter(beta):
return SIQ_loss(X,Y,beta,B,theta,tau)
return inter
def beta_on_others_(X,Y,B,theta,tau,beta0):
d = X.shape[1]
# define the linear constraint beta_0 > 0
ub = np.ones(d)*np.inf
lb = - np.ones(d)*np.inf
lb[0] = 0.
bou = Bounds(lb, ub)
res = minimize(loss_on_beta_(X,Y,B,theta,tau), beta0, method='L-BFGS-B',bounds = bou)
return res.x
# -
# ### 4*. Optimization for both $\beta$ and $G(\cdot)$
# ### 5. Optimization for nonparametric function $G(\cdot)$
# \begin{align}
# BIC(N_n) = \log\{\text{Loss}\} + \dfrac{\log n}{2n}(N_n+m)
# \end{align}
record = {}
for N_n in range(2,2*N):
dB_n = deBoor(a, b, m, N_n)
B_n = [i for i in dB_n.values()]
theta_n = theta_on_others(X,Y,beta,B_n,tau)
BIC_n = np.log(SIQ_loss(X,Y,beta,B_n,theta_n,tau)) + np.log(n)/(2*n)*(N_n+m)
record[N_n] = [theta, BIC_n]
record_df = DataFrame(record, index=['theta','BIC']).T
record_df
| Single Index Quantile Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="n-bz83O4K9Xp"
import numpy as np
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="d6D-kiST7uWE" outputId="0133437a-f0da-4d7d-ec45-bb9546800169"
url = 'https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/09_Time_Series/Apple_Stock/appl_1980_2014.csv'
apple = pd.read_csv(url)
apple.head()
# + id="CZzsNy-r7uRs" colab={"base_uri": "https://localhost:8080/"} outputId="115b7cd0-b4b6-4c1e-baf2-90294223cdd0"
apple.dtypes
# + id="YJLqPmhq7uMv"
apple['Date'] = pd.to_datetime(apple.Date)
# + colab={"base_uri": "https://localhost:8080/"} id="r8qlD404inHG" outputId="ce47973d-7ad6-4c87-8411-30bfcdd8aec2"
apple.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="1PB4JR5uinEn" outputId="54fd56c0-e29f-4e0e-c0b8-d997eb85b733"
apple.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 236} id="B03fyZjtinCG" outputId="46439bb7-9fc6-4c11-c6a1-bd29db1006ec"
apple = apple.set_index('Date')
apple.head()
# + colab={"base_uri": "https://localhost:8080/"} id="giqcxchnim_e" outputId="e7520e16-705a-4377-9832-b245631af1f5"
apple.index.is_unique
# + colab={"base_uri": "https://localhost:8080/", "height": 236} id="cZr5Emnxi6Vf" outputId="221ae826-bbd3-4fa1-e8ad-d558742694a1"
apple.sort_index(ascending = True).head()
# + colab={"base_uri": "https://localhost:8080/", "height": 454} id="4JB3JnfujOKv" outputId="5cb76a7a-78e6-4fd1-890c-8883f023f640"
apple_month = apple.resample("BM").mean()
apple_month
# + colab={"base_uri": "https://localhost:8080/"} id="akCgSfYmjnmP" outputId="4bb8e31d-218d-48b6-d2e9-ee9ad479ef5b"
(apple.index.max() - apple.index.min()).days
# + colab={"base_uri": "https://localhost:8080/"} id="6sKYXh5Vjrph" outputId="03090607-4fe2-491e-8c58-9653220a3e54"
apple_months = apple.resample('BM').mean()
len(apple_months.index)
| Exercise18.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
This part of code is the Q learning brain, which is a brain of the agent.
All decisions are made in here.
View more on my tutorial page: https://morvanzhou.github.io/tutorials/
"""
import numpy as np
import pandas as pd
class QLearningTable:
def __init__(self, actions, learning_rate=0.01, reward_decay=0.9, e_greedy=0.9):
self.actions = actions # a list
self.lr = learning_rate
self.gamma = reward_decay
self.epsilon = e_greedy
self.q_table = pd.DataFrame(columns=self.actions, dtype=np.float64)
def choose_action(self, observation):
self.check_state_exist(observation)
# action selection
if np.random.uniform() < self.epsilon:
# choose best action
state_action = self.q_table.loc[observation, :]
# some actions may have the same value, randomly choose on in these actions
action = np.random.choice(state_action[state_action == np.max(state_action)].index)
else:
# choose random action
action = np.random.choice(self.actions)
return action
def learn(self, s, a, r, s_):
self.check_state_exist(s_)
q_predict = self.q_table.loc[s, a]
if s_ != 'terminal':
q_target = r + self.gamma * self.q_table.loc[s_, :].max() # next state is not terminal
else:
q_target = r # next state is terminal
self.q_table.loc[s, a] += self.lr * (q_target - q_predict) # update
def check_state_exist(self, state):
if state not in self.q_table.index:
# append new state to q table
self.q_table = self.q_table.append(
pd.Series(
[0]*len(self.actions),
index=self.q_table.columns,
name=state,
)
)
# -
| Q Learn算法更新/RL_brain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="40733c39647bac5056ba0dada8409d6184fefb68"
# ## Tutorial Keras: Transfer Learning with ResNet50 for image classification on Cats & Dogs dataset
#
# ### <NAME>
# + [markdown] _uuid="0182259264d23ccfb3c27d530ed10fb4ba7a35da"
# This kernel is intended to be a tutorial on Keras around image files handling for Transfer Learning using pre-trained weights from ResNet50 convnet.
#
# Though loading all train & test images resized (224 x 224 x 3) in memory would have incurred ~4.9GB of memory, the plan was to batch source image data during the training, validation & testing pipeline. Keras ImageDataGenerator supports batch sourcing image data for all training, validation and testing. Actually, it is quite clean and easy to use Keras ImageDataGenerator except few limitations (listed at the end).
#
# Keras ImageDataGenerator expects labeled training images to be available in certain folder heirarchy, 'train' data was manually split into 10k for training & 2.5k for validation and re-arranged into the desired folder hierarchy. Even 'test' images had to rearranged due to a known issue in flow_from_directory.
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import cv2
import os
# + [markdown] _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# ### Global Constants
# + _uuid="d157dc1311928d1ada13ed2ef4a34c4ea5c0b538"
# Fixed for our Cats & Dogs classes
NUM_CLASSES = 2
# Fixed for Cats & Dogs color images
CHANNELS = 3
IMAGE_RESIZE = 224
RESNET50_POOLING_AVERAGE = 'avg'
DENSE_LAYER_ACTIVATION = 'softmax'
OBJECTIVE_FUNCTION = 'categorical_crossentropy'
# Common accuracy metric for all outputs, but can use different metrics for different output
LOSS_METRICS = ['accuracy']
# EARLY_STOP_PATIENCE must be < NUM_EPOCHS
NUM_EPOCHS = 10
EARLY_STOP_PATIENCE = 3
# These steps value should be proper FACTOR of no.-of-images in train & valid folders respectively
# Training images processed in each step would be no.-of-train-images / STEPS_PER_EPOCH_TRAINING
STEPS_PER_EPOCH_TRAINING = 10
STEPS_PER_EPOCH_VALIDATION = 10
# These steps value should be proper FACTOR of no.-of-images in train & valid folders respectively
# NOTE that these BATCH* are for Keras ImageDataGenerator batching to fill epoch step input
BATCH_SIZE_TRAINING = 100
BATCH_SIZE_VALIDATION = 100
# Using 1 to easily manage mapping between test_generator & prediction for submission preparation
BATCH_SIZE_TESTING = 1
# + _uuid="ee0da10690e2848537102cb0c74f9fe19a50431b"
from tensorflow.python.keras.applications import ResNet50
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.layers import Dense
###
### Below systax is available with TensorFlow 1.11 onwards but this upgrade is not available for Kaggle kernel yet
###
#import tensorflow as tf
#print(tf.__version__)
#import tensorflow as tf
#from tf.keras.applications import ResNet50
#from tf.keras.models import Sequential
# + [markdown] _uuid="b651539464d36c7443a601e914479f0d2153f435"
# ### ResNet50
# * Notice that resnet50 folder has 2 pre-trained weights files... xyz_tf_kernels.h5 & xyz_tf_kernels_NOTOP.h5
# * The xyz_tf_kernels.h5 weights is useful for pure prediction of test image and this prediction will rely completely on ResNet50 pre-trained weights, i.e., it does not expected any training from our side
# * Out intention in this kernel is Transfer Learning by using ResNet50 pre-trained weights except its TOP layer, i.e., the xyz_tf_kernels_NOTOP.h5 weights... Use this weights as initial weight for training new layer using train images
# + _uuid="2fc47c3f887e6713b5a2be38c02b38a8ec80bf97"
resnet_weights_path = '../input/resnet50/resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5'
# + [markdown] _uuid="4ebad39fff164624f5616a83cd8831308431da7d"
# ### Define Our Transfer Learning Network Model Consisting of 2 Layers
#
# Here, we are preparing specification or blueprint of the TensorFlow DAG (directed acyclcic graph) for just the MODEL part.
# + _uuid="0bb4a8eccd70874ef21f0809f478f993fa127fb2"
#Still not talking about our train/test data or any pre-processing.
model = Sequential()
# 1st layer as the lumpsum weights from resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5
# NOTE that this layer will be set below as NOT TRAINABLE, i.e., use it as is
model.add(ResNet50(include_top = False, pooling = RESNET50_POOLING_AVERAGE, weights = resnet_weights_path))
# 2nd layer as Dense for 2-class classification, i.e., dog or cat using SoftMax activation
model.add(Dense(NUM_CLASSES, activation = DENSE_LAYER_ACTIVATION))
# Say not to train first layer (ResNet) model as it is already trained
model.layers[0].trainable = False
# + _uuid="b617a28f0f89b272a0aa2af6cf72f2dd642ee052"
model.summary()
# + [markdown] _uuid="3587981a5b6b1d6ff8221738d51d78cafb2dd02c"
# ### Compile Our Transfer Learning Model
# + _uuid="670238611770f43a056332cc06efff44e20ad124"
from tensorflow.python.keras import optimizers
sgd = optimizers.SGD(lr = 0.01, decay = 1e-6, momentum = 0.9, nesterov = True)
model.compile(optimizer = sgd, loss = OBJECTIVE_FUNCTION, metrics = LOSS_METRICS)
# + [markdown] _uuid="8d003a99fe4ff58b8905c7f6b5286d97a852cb7a"
# ### Prepare Keras Data Generators
#
# Keras *ImageDataGenerator(...)* generates batches of tensor image data with real-time data augmentation. The data will be looped over (in batches). It is useful with large dataset to source, pre-process (resize, color conversion, image augmentation, batch normalize) & supply resulting images in batches to downstream Keras modeling components, namely *fit_generator(...)* & *predict_generator(...)* -vs- *fit(...)* & *predict(...)* for small dataset.
#
# Kaggle competition rule expects Dog & Cat to be labeled as 1 & 0. Keras >> ImageDataGenerator >> flow_from_directory takes in 'classes' list for mapping it to LABEL indices otherwise treats sub-folders enumerated classes in alphabetical order, i.e., Cat is 0 & Dog is 1.
# + _uuid="ac9ebd909fee81c8c8d9b9fccb6590944d8106eb"
from keras.applications.resnet50 import preprocess_input
from keras.preprocessing.image import ImageDataGenerator
image_size = IMAGE_RESIZE
# preprocessing_function is applied on each image but only after re-sizing & augmentation (resize => augment => pre-process)
# Each of the keras.application.resnet* preprocess_input MOSTLY mean BATCH NORMALIZATION (applied on each batch) stabilize the inputs to nonlinear activation functions
# Batch Normalization helps in faster convergence
data_generator = ImageDataGenerator(preprocessing_function=preprocess_input)
# flow_From_directory generates batches of augmented data (where augmentation can be color conversion, etc)
# Both train & valid folders must have NUM_CLASSES sub-folders
train_generator = data_generator.flow_from_directory(
'../input/catsdogs-trainvalid-80pc-prepd/trainvalidfull4keras/trainvalidfull4keras/train',
target_size=(image_size, image_size),
batch_size=BATCH_SIZE_TRAINING,
class_mode='categorical')
validation_generator = data_generator.flow_from_directory(
'../input/catsdogs-trainvalid-80pc-prepd/trainvalidfull4keras/trainvalidfull4keras/valid',
target_size=(image_size, image_size),
batch_size=BATCH_SIZE_VALIDATION,
class_mode='categorical')
# + _uuid="5014f91b6aeae661f0bc5db5d546089a07ca5b9d"
# Max number of steps that these generator will have opportunity to process their source content
# len(train_generator) should be 'no. of available train images / BATCH_SIZE_TRAINING'
# len(valid_generator) should be 'no. of available train images / BATCH_SIZE_VALIDATION'
(BATCH_SIZE_TRAINING, len(train_generator), BATCH_SIZE_VALIDATION, len(validation_generator))
# + [markdown] _uuid="f56e7cf9eae4c86c72468e322e7f00859a2ce9cd"
# ### Train Our Model With Cats & Dogs Train (splitted) Data Set
# + _uuid="2dfad78129d725f42110cde0270c32d7373d6d1d"
# Early stopping & checkpointing the best model in ../working dir & restoring that as our model for prediction
from tensorflow.python.keras.callbacks import EarlyStopping, ModelCheckpoint
cb_early_stopper = EarlyStopping(monitor = 'val_loss', patience = EARLY_STOP_PATIENCE)
cb_checkpointer = ModelCheckpoint(filepath = '../working/best.hdf5', monitor = 'val_loss', save_best_only = True, mode = 'auto')
# + _uuid="d108d3598210930bab431f43c6865488b01d467b"
# Grid Search is an ideal candidate for distributed machine learning
# Pseudo code for hyperparameters Grid Search
from sklearn.grid_search import ParameterGrid
param_grid = {'epochs': [5, 10, 15], 'steps_per_epoch' : [10, 20, 50]}
grid = ParameterGrid(param_grid)
# Accumulate history of all permutations (may be for viewing trend) and keep watching for lowest val_loss as final model
for params in grid:
print(params)
# + _uuid="cf85fe3c0653aa56503b7da058ea8acf445eec6e"
fit_history = model.fit_generator(
train_generator,
steps_per_epoch=STEPS_PER_EPOCH_TRAINING,
epochs = NUM_EPOCHS,
validation_data=validation_generator,
validation_steps=STEPS_PER_EPOCH_VALIDATION,
callbacks=[cb_checkpointer, cb_early_stopper]
)
model.load_weights("../working/best.hdf5")
# + [markdown] _uuid="1ef4dd95f6d12f9277255576645e8bfce5270b81"
# ### Training Metrics
#
# One of the default callbacks that is registered when training all deep learning models is the History callback. It records training metrics (training accuracy, training loss, validation loss & validation accuracy) for each epoch. Note that training accuracy & loss during epoch steps are somewhat incomplete information and they are not recorded in history.
#
# Observe that training uses early stopping, hence metrics is available for epochs run, not for NUM_EPOCHS.
# + _uuid="383d7a0aa87e1508a46fcebfcd5a80d7aafb840f"
print(fit_history.history.keys())
# + _uuid="00e51568f7e6022c6738c0e1a41080bc7152c257"
plt.figure(1, figsize = (15,8))
plt.subplot(221)
plt.plot(fit_history.history['acc'])
plt.plot(fit_history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'valid'])
plt.subplot(222)
plt.plot(fit_history.history['loss'])
plt.plot(fit_history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'valid'])
plt.show()
# + _uuid="3941e7c5d983e2ad6ae6d6da0a1415ea27e1db7e"
# NOTE that flow_from_directory treats each sub-folder as a class which works fine for training data
# Actually class_mode=None is a kind of workaround for test data which too must be kept in a subfolder
# batch_size can be 1 or any factor of test dataset size to ensure that test dataset is samples just once, i.e., no data is left out
test_generator = data_generator.flow_from_directory(
directory = '../input/test-files-prepd/test4keras/test4keras',
target_size = (image_size, image_size),
batch_size = BATCH_SIZE_TESTING,
class_mode = None,
shuffle = False,
seed = 123
)
# Try batch size of 1+ in test_generator & check batch_index & filenames in resulting batches
'''
for i in test_generator:
#print(test_generator.batch_index, test_generator.batch_size)
idx = (test_generator.batch_index - 1) * test_generator.batch_size
print(test_generator.filenames[idx : idx + test_generator.batch_size])
'''
# + [markdown] _uuid="8c6e7f35552f87b522aa32397ffb71ea732ce80b"
# ### Observe Prediction Time With Different Batch Size
#
# With GPU, 97s for full prediction with batch_size=100 -vs- 264s with 1. But note that to avoid ImageDataGenerator iterator repeatability, we need to use 1 as batch_size.
# + _uuid="9911ff38ec4b82f5282f078aae63b2a510a9a6cc"
# Reset before each call to predict
test_generator.reset()
pred = model.predict_generator(test_generator, steps = len(test_generator), verbose = 1)
predicted_class_indices = np.argmax(pred, axis = 1)
# + _uuid="e54bc22bd1a2af7d704f9164b624b79b8cbd4f56"
TEST_DIR = '../input/test-files-prepd/test4keras/test4keras/'
f, ax = plt.subplots(5, 5, figsize = (15, 15))
for i in range(0,25):
imgBGR = cv2.imread(TEST_DIR + test_generator.filenames[i])
imgRGB = cv2.cvtColor(imgBGR, cv2.COLOR_BGR2RGB)
# a if condition else b
predicted_class = "Dog" if predicted_class_indices[i] else "Cat"
ax[i//5, i%5].imshow(imgRGB)
ax[i//5, i%5].axis('off')
ax[i//5, i%5].set_title("Predicted:{}".format(predicted_class))
plt.show()
# + _uuid="39bac850385ba7129c97c1c0d389b36e7f2f0dfe"
results_df = pd.DataFrame(
{
'id': pd.Series(test_generator.filenames),
'label': pd.Series(predicted_class_indices)
})
results_df['id'] = results_df.id.str.extract('(\d+)')
results_df['id'] = pd.to_numeric(results_df['id'], errors = 'coerce')
results_df.sort_values(by='id', inplace = True)
results_df.to_csv('submission.csv', index=False)
results_df.head()
# + [markdown] _uuid="2e5600300a330725b29c74c813135144a22cee56"
# ### Keras Limitations
#
# * [10/02/2018] The *validation_split* is not supported in *fit_generator*, hence its expects ImageDataGenerator for pre-splitted train & valid.
# * [10/02/2018] Model learning through *fit_generator* is not compatible for Sklearn *GridSearchCV* again *mostly* due to no support for *validation_split*.
# + [markdown] _uuid="60a34703bd652a314983948d93ee2dfe6d760ec3"
# ### Followup Plan
#
# 1. Scale and pad and avoid aspect ratio change of original image through Keras ImageDataGenerator pre-processing insfrastructure
# 2. Image augmentation
# 3. Pipeline
# 4. Distributed ML for Grid Search on Spark Cluster
# + [markdown] _uuid="422beefc8f408582e85292cec3fe8225228f2ab5"
# ### References
#
# 1. [Transfer Learning by <NAME>](https://www.kaggle.com/dansbecker/transfer-learning)
| Tutorial Keras_ Transfer Learning with ResNet50.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// # Клас контейнер на *C#*
// Створимо базовий клас `String`, який буде зберігатися у класі `Text`
class String
{
protected string value;
public String(string value)
{
this.value = value;
}
public int len()
{
return this.value.Length;
}
public void all_upper()
{
this.value = this.value.ToUpper();
}
public int compare(string str)
{
if (this.value == str) return 1;
else return 0;
}
public string get()
{
return this.value;
}
}
// Цей клас містить такі методи:
// * `String` *конструктор*: передає у клас рядок;
// * `len`: повертає довжину рядка;
// * `all_upper`: переводить рядок у uppercase;
// * `compare`: приймає 1 рядок та порівнює рядок класу з ним. Повертає 1 або 0 якщо вони однакові чи ні відповідно;
// * `get`: повертає значення рядка, що зберігається у класі;
// Створимо клас-контейнер `Text`
// +
using System.Collections.Generic;
class Text
{
private List<String> Strings;
public Text() {this.Strings = new List<String>();}
private int length()
{
return this.Strings.Count;
}
public void show()
{
if (length() == 0)
{
Console.WriteLine("EMPTY!");
}
else
{
foreach (String str in Strings)
{
Console.WriteLine(str.get());
}
}
}
public void add(String str)
{
this.Strings.Add(str);
}
public void delete_one(int index = -1)
{
if (length() > 0)
{
if (index < 0)
{
this.Strings.RemoveAt(this.Strings.Count + index);
}
else
{
this.Strings.RemoveAt(index);
}
}
}
public void delete_all()
{
this.Strings.Clear();
}
public void delete_len(int del_len)
{
List<int> result;
result = new List<int>();
for (int i = 0; i < length(); i++)
{
if (this.Strings[i].len() == del_len) result.Add(i);
}
for (int i = 0; i < result.Count; i++) this.Strings.RemoveAt(result[i] - i);
}
public void upper()
{
foreach (String str in Strings)
{
str.all_upper();
}
}
public void search(string search_str)
{
int result = 0;
foreach (String str in Strings)
{
result += str.compare(search_str);
}
Console.WriteLine($"{result} '{search_str}' в тексті");
}
}
// -
// Цей клас містить наступні методи:
// * `Text` *конструктор*;
// * `length`: повертає кількість елементів тексту;
// * `delete_all`: очищує весь клас;
// * `delete_one`: видаляє елемент за індексом чи останній елемент;
// * `delete_len`: видаляє усі елементи заданої довжини;
// * `add`: додає елемент в кінець тексту;
// * `show`: виводить текст у консоль;
// * `upper`: переводить усі елементи у верхній регістр;
// * `search`: приймає рядок, виводить кількість співпадань у тексті;
// Перевіримо роботу классів. Для цього створимо 4 рядка та 4 тексти з них.
// +
var s1 = new String("Foo Bar");
var s2 = new String("Spam Eggs");
var s3 = new String("Pupa Lupa");
var s4 = new String("C++ is harder");
var t1 = new Text();
var t2 = new Text();
var t3 = new Text();
var t4 = new Text();
// -
// Циклом додамо до кожного тексту усі рядки
// +
List<Text> T;
List<String> S;
S = new List<String>();
T = new List<Text>();
T.Add(t1);
T.Add(t2);
T.Add(t3);
T.Add(t4);
S.Add(s1);
S.Add(s2);
S.Add(s3);
S.Add(s4);
foreach (Text t in T)
{
foreach (String s in S)
{
t.add(s);
}
}
// -
// Повне очищення тексту
t1.delete_all();
t1.show();
// Видалення 1 елементу за індексом та без
t2.delete_one();
t2.delete_one(1);
t2.show();
// Видалення за довжиною
t3.delete_len(9);
t3.show();
// Пошук у тексті, перевод тексту у верхній регістр
t4.add(s3);
t4.search("Spam Eggs");
t4.search("Pupa Lupa");
t4.upper();
t4.show();
| Notebooks/3hird/C_Sharp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.0
# language: julia
# name: julia-1.5
# ---
# Example 5.4 of [AJPR14]. The JSR is 3.917384715148.
#
# [AJPR14] <NAME>, <NAME>, <NAME> and <NAME>,
# *Joint spectral radius and path-complete graph Lyapunov functions.*
# SIAM J. CONTROL OPTIM 52(1), 687-717, **2014**.
using HybridSystems
A1 = [-1 -1
-4 0]
A2 = [ 3 3
-2 1]
s = discreteswitchedsystem([A1, A2])
# Pick an SDP solver.
using CSDP
using JuMP
optimizer_constructor = optimizer_with_attributes(CSDP.Optimizer, MOI.Silent() => true);
# We first try with CQLF and find the smp.
using SwitchOnSafety
soslyapb(s, 1, optimizer_constructor=optimizer_constructor, tol=1e-4)
seq = sosbuildsequence(s, 1, p_0=:Primal)
psw = findsmp(seq)
# Now with quartic forms the upper bound get closer.
soslyapb(s, 2, optimizer_constructor=optimizer_constructor, tol=1e-4)
seq = sosbuildsequence(s, 2, p_0=:Primal)
psw = findsmp(seq)
| examples/AJPR14e54.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Issue 10 -- Waveform Time Shifting
#
# Double check / Fix waveform time-shifting
# Setup ipython environment
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# The Important Things
from numpy import *
from positive import *
from positive.physics import pn
from positive.maths import lim
from matplotlib.pyplot import *
from nrutils import scsearch,gwylm,pad_wfarr,tshift_wfarr,plot_wfarr
from positive.physics import mishra
# Find a Simulation
A = scsearch(nonspinning=True,q=1,keyword='hrq',verbose=True)
# Load Simulation Data
y = gwylm(A[0],lm=[(2,2),(3,2)],verbose=True)
# Extract a waveform array for testing
wfarr = y.lm[2,2]['psi4'].wfarr.copy()
# Test padding behavior
wfarr2 = pad_wfarr( wfarr, 1, verbose=True )
t,p,c = wfarr2.T
plot( t,p )
plot( t,c )
# +
# Test time shifting on arrays --- Index based
wfarr3 = tshift_wfarr( wfarr, -diff(lim(t))/8, verbose=True )
t = wfarr[:,0]
# plot
plot( t, wfarr[:,1], label='original' )
plot( t, wfarr3[:,1], label='shifted' )
xlim( lim(t) )
legend()
# +
# Test time shifting on arrays --- Index based
wfarr4 = tshift_wfarr( wfarr3, diff(lim(t))/8, verbose=True )
t = wfarr[:,0]
# plot
plot( t, wfarr3[:,1], label='shifted' )
plot( t, wfarr4[:,1], label='shifted back!' )
plot( t, wfarr[:,1], label='original', color='k', ls='--', alpha=0.5 )
xlim( lim(t) )
legend()
# +
# Time shift gwf
yy = y.lm[2,2]['psi4']
yy1 = yy.tshift(diff(lim(t))/8,verbose=True,method='index')
# plot
yy.plot( ref_gwf=yy1, labels=('original','shifted') )
# -
| issues/open/issue_10_waveform_time_shifting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="1OLxnpePKtcI"
# # Building an Estimator
#
# Once we have our data in the "right" format (i.e. files containing
# `tf.train.Example` protocol buffers), Tensorflow makes the actual
# ML part straightforward.
#
# In this notebook we will first set up the computational graph that
# reads the input data and transforms it into tensors, and then use
# Tensorflow's "canned" estimator to learn from there.
#
# This notebook contains a whole series of exciting bonus sections
# that you certainly won't have time to solve during the workshop,
# but you might want to come back to these after the workshop to create
# more sophisticated models that solve the task better.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 2620, "status": "ok", "timestamp": 1526712849326, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="ul42dk3WKtcJ" outputId="33c8a779-4f19-4a26-c017-d0b735f5f2d1"
from __future__ import division, print_function
import itertools
import numpy as np
import tensorflow as tf
from matplotlib import pyplot
# !mkdir -p _derived # (Needed for running in Colab environment.)
# %matplotlib inline
# Always make sure you are using running the expected version.
# There are considerable differences between versions...
# Tested with 1.4.1 and 1.7.0
tf.__version__
# + [markdown] colab_type="text" id="2HJ-v7BHKtcO"
# # 1 Reading the data
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 561} colab_type="code" executionInfo={"elapsed": 1915, "status": "ok", "timestamp": 1526712855645, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="-wOh5H8jKtcQ" outputId="8d151d58-ecea-474a-8b68-814014b3d7a9"
# Load data generated in 1_qd_data:
data_path = '../data/dataset_img'
# Show files in dataset.
# !ls -lh $data_path
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 221} colab_type="code" executionInfo={"elapsed": 983, "status": "ok", "timestamp": 1526712875627, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="iVl9e7Y7KtcU" outputId="336371e8-3463-4529-86f3-6aa5f54d266c"
# It's customary to identify the label by an integer number. These
# numbers map to the different classes as follows:
classes = open('%s/labels.txt' % data_path).read().splitlines()
print('%d label classes:\n' % len(classes))
for i, label in enumerate(classes):
print('%d -> %s' % (i, label))
# + [markdown] colab_type="text" id="YJ3wTbh5KtcX"
# ## 1.1 Reading the data using pure Python
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 136} colab_type="code" executionInfo={"elapsed": 982, "status": "ok", "timestamp": 1526712904783, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="KTxdM9p0KtcX" outputId="88b2982a-737e-4e49-a376-f2284d3e77ca"
# Step 1 : Read a single record from the first file.
train_files = tf.gfile.Glob('%s/train-*' % data_path)
record = next(tf.python_io.tf_record_iterator(train_files[0]))
# Step 2 : Parse record into tf.train.Example proto.
example = tf.train.Example.FromString(record)
# We need the image "img_64" and the "label". Both are of type
# int64_list: the label is stored by its index and the image is
# a 64x64 list of integers that are the pixel values.
for name, feature in example.features.feature.items():
print('%20s (%s)' % (name, feature.WhichOneof('kind')))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="UJ9laaC2Ktcb"
# Step 3 : Read features.
img_64 = example.features.feature['img_64'].int64_list.value
img_64 = np.array(img_64).reshape((64, 64)) / 255.
label = example.features.feature['label'].int64_list.value[0]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 280} colab_type="code" executionInfo={"elapsed": 1283, "status": "ok", "timestamp": 1526712912903, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="GaAAj3XQKtcd" outputId="46098ce1-ec95-417c-ecd0-ae1f5fc9f2ac"
# Visualize drawing.
def show_img(img_64, title, ax=None):
(ax if ax else pyplot).matshow(img_64, cmap='gray')
ax = ax if ax else pyplot.gca()
ax.set_xticks([])
ax.set_yticks([])
ax.set_title(title)
show_img(img_64, classes[label])
# + [markdown] colab_type="text" id="I5kGHQupKtch"
# ## 1.2 Reading the data using Tensorflow
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1128, "status": "ok", "timestamp": 1526712921788, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="Jip76pl4Ktci" outputId="2cddae33-defe-412b-e2bf-78068f548e59"
# %%writefile _derived/4_input_fn_img.py
# (Written into separate file for sharing with cloud code.)
# Read input data from sharded files using 1.4 API.
# This is copied and slightly simplified from tf_snippets.ipynb
# Note that this cell simply defines the functions but does not yet call
# them. The functions are executed in the next cell within a tf.Graph()
# context.
# This dictionary specifies what "features" we want to extract from the
# tf.train.Example protos (i.e. what they look like on disk). We only
# need the image data "img_64" and the "label". Both features are tensors
# with a fixed length.
# You need to specify the correct "shape" and "dtype" parameters for
# these features...
feature_spec = {
# Single label per example => shape=[1] (we could also use shape=() and
# then do a transformation in the input_fn).
'label': tf.FixedLenFeature(shape=[1], dtype=tf.int64),
# The bytes_list data is parsed into tf.string.
'img_64': tf.FixedLenFeature(shape=[64, 64], dtype=tf.int64),
}
def parse_example(serialized_example):
# Convert string to tf.train.Example and then extract features/label.
features = tf.parse_single_example(serialized_example, feature_spec)
# Important step: remove "label" from features!
# Otherwise our classifier would simply learn to predict
# label=features['label']...
label = features.pop('label')
# Convert int64 [0..255] to float [0..1]
features['img_64'] = tf.cast(features['img_64'], tf.float32) / 255.
return features, label
# Common Tensorflow pattern : wrap function to specify parameters.
# The estimator interface expects a "input_fn" as a parameter, and not the
# tensors (features, labels) so it can re-create the tensors in different
# graphs later on.
def make_input_fn(files_pattern, batch_size=100):
def input_fn():
# Signature input_fn: () -> features=Dict[str, Tensor], labels=Tensor
ds = tf.data.TFRecordDataset(tf.gfile.Glob(files_pattern))
ds = ds.map(parse_example).batch(batch_size)
ds = ds.shuffle(buffer_size=5*batch_size).repeat()
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
return input_fn
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 158} colab_type="code" executionInfo={"elapsed": 1204, "status": "ok", "timestamp": 1526712931620, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="LUiFMFAlKtcl" outputId="c0249db8-e69c-4170-adc8-dc2275e287bf"
# Define a graph, load a batch of examples and display one of them.
# %run -i _derived/4_input_fn_img.py
# First, we create the input_fn -- this defines which files to read from
# and the batch size, and can later be passed around.
batch_size = 5
input_fn = make_input_fn(files_pattern='%s/train-*' % data_path,
batch_size=batch_size)
with tf.Graph().as_default():
# After registering a default graph, we call the previously defined
# input_fn.
# This yields the dictionary of feature tensors and the label tensor.
features, labels = input_fn()
with tf.train.MonitoredSession() as sess:
# Note: We create a tf.train.MonitoredSession instead of a normal
# tf.Session because Tensorflow is using pipelines under the hood
# to read data in a streaming fashion from disk into our computational
# Graph. If we used a plain old tf.Session(), then we would have to
# take care of initializing the threads for the pipelines ourselves...
img_64_, labels_ = sess.run([features['img_64'], labels])
# Note that first dimension is the batch dimension.
for i in range(batch_size):
ax = pyplot.subplot(1, batch_size, i+1)
show_img(img_64_[i], classes[labels_[i][0]], ax)
# Also note how content changes upon every re-execution of this cell.
# That is because of the randomization in the input_fn.
# + [markdown] colab_type="text" id="ry51bTZ-Ktco"
# # 2 Canned estimators
#
# Once we have our `input_fn`, it's a piece of cake to do some
# machine learning on top of it using Tensorflow's "canned
# estimators". Let's start with a simple
# [LinearClassifier](https://www.tensorflow.org/versions/master/api_docs/python/tf/estimator/LinearClassifier).
#
# A LinearClassifier implements simple **logistic regression** that is basically
# a neural network where every input neuron (pixel image) is directly connected
# to all output neurons (probabilities for different classes). Every output neuron
# effectively remembers a "mask" in the input pixel space, an approach that
# quickly learns to identify drawings that have similar pixel activations, but
# fails to learn anything more complicated (e.g. an elephant's trunk could
# activate the giraffe's "neck mask" if it happened to be in the same pixel
# region).
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 88} colab_type="code" executionInfo={"elapsed": 819, "status": "ok", "timestamp": 1526712949974, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="UFfD2AROKtcp" outputId="1ac771c9-4030-43ed-d798-fbb53f1360a2"
# "Feature columns" tell our canned estimator how to make use of the
# data returned by input_fn(). In our case we have a single feature that
# is of numerical (defaults to tf.float32) type with shape=(64, 64).
feature_columns = [
# Add a feature column of numeric type and correct shape for the "img_64" feature.
tf.feature_column.numeric_column('img_64', shape=(64, 64))
]
# Note that feature columns can also be used to generate the "feature_spec"
# used to parsing the tf.train.Example protocol buffers, and that there are
# other feature column types that can do complicated data transformation
# (such as looking up strings and creating embeddings) -- but that's a story
# for another workshop ;-)
# (we also need to specify number of output classes)
linear_estimator = tf.estimator.LinearClassifier(feature_columns=feature_columns,
n_classes=len(classes))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 272} colab_type="code" executionInfo={"elapsed": 7959, "status": "ok", "timestamp": 1526712987872, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="4IBQ1eWTKtct" outputId="530ef497-2d36-4e9d-f3fc-c40726c93a1a"
# Connect it to the input_fn with the training data and train the classifier.
### YOUR ACTION REQUIRED:
# You have to specify what data to use for training (which input_fn), and how
# many steps you want to train for. The number of examples used for training is
# = steps * batch_size.
input_fn = make_input_fn('%s/train-*' % data_path) #input_fn = make_input_fn(...)
### YOUR ACTION REQUIRED:
# Try different number of training steps and see how this influences training time
# (and accuracy in cell below). Note that you have to re-run the previous cell
# to "reset" the classifier (otherwise the estimator will simply *continue* the
# training with an additional number of steps).
steps = 300 #steps =
linear_estimator.train(input_fn=input_fn, steps=steps)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 428} colab_type="code" executionInfo={"elapsed": 3364, "status": "ok", "timestamp": 1526712994230, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="P726iOBsKtcy" outputId="c33992e2-3d33-42a2-f955-ee4d261d0279"
# Evaluate the fitted classifier.
### YOUR ACTION REQUIRED:
# Find the correct values for below variables. Note that you should *not*
# evaluate with the same data used for training.
input_fn = make_input_fn('%s/eval-*' % data_path) #input_fn =
steps = 100 #steps =
linear_estimator.evaluate(input_fn=input_fn, steps=steps)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 800} colab_type="code" executionInfo={"elapsed": 1842, "status": "ok", "timestamp": 1526713007059, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="nYmfTgvrKtc2" outputId="fed911ce-a294-4ea4-cbc6-5e96fa753aca"
# The simple LinearClassifier does not perform too well... Let's examine
# some classifications so we can see which cases were failing.
# Because estimator.predict() only returns class probabilities and not the
# input data, we parse the features in Python (which gives us a handle on
# the input data) and then create an input_fn using numpy_input_fn().
def get_label_img(files_pattern, n):
"""Reads batch of "img_64" and "label" features.
Args:
files_pattern: Pattern matching files containing tf.train.Example.
n: Number of examples to parse.
Returns:
(imgs, labels) with shapes (n, 64, 64) and (n, 1). The image pixel values
fall in the range 0..1.
"""
imgs = np.zeros((n, 64, 64), dtype=np.float32)
labels = np.zeros((n, 1), dtype=np.int64)
i = 0
for filename in tf.gfile.Glob(files_pattern):
for record in tf.python_io.tf_record_iterator(filename):
example = tf.train.Example.FromString(record)
labels[i, 0] = example.features.feature['label'].int64_list.value[0]
img_64 = example.features.feature['img_64'].int64_list.value
imgs[i, :, :] = np.array(img_64).reshape((64, 64)) / 255.
i += 1
if i >= n:
return imgs, labels
def show_predictions(estimator, rows=4, cols=4, predict_keys=None):
"""Shows predictions + labels for a couple of examples."""
# Read data in Python.
imgs, labels = get_label_img('%s/test-*' % data_path, n=rows*cols)
# Create input_fn from Python data.
input_fn = tf.estimator.inputs.numpy_input_fn({'img_64': imgs}, shuffle=False)
# Get predictions.
predict_iter = estimator.predict(input_fn=input_fn, predict_keys=predict_keys)
_, axs = pyplot.subplots(rows, cols, figsize=(cols*3, rows*3))
for (i, prediction) in enumerate(predict_iter):
# Plot image, predicted label + correct label.
predicted = np.argmax(prediction['probabilities'])
label = labels[i, 0]
title = '%s? %s!' % (classes[predicted], classes[label])
ax = axs[i//cols][i%cols]
show_img(imgs[i], title, ax)
ax.title.set_position([0.5, 1.0])
show_predictions(linear_estimator)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1635} colab_type="code" executionInfo={"elapsed": 167314, "status": "ok", "timestamp": 1526713557417, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="vhTUnm3GKtc7" outputId="3b2c2eca-34fa-4b99-fa9c-10d6245d8da4"
# Let's use another canned estimator : The "DNNClassifier" (where DNN stands
# for "deep neural network"). Since we're using the same estimator interface,
# we can use the same input_fn() as before.
### YOUR ACTION REQUIRED:
# Try to find a configuration that outperforms our linear classifier.
steps = 3000 #steps =
hidden_units = [1000] #hidden_units =
dnn_estimator = tf.estimator.DNNClassifier(
hidden_units=hidden_units,
feature_columns=feature_columns,
n_classes=len(classes))
dnn_estimator.train(input_fn=make_input_fn('%s/train-*' % data_path),
steps=steps)
dnn_estimator.evaluate(input_fn=make_input_fn('%s/eval-*' % data_path),
steps=100)
# + [markdown] colab_type="text" id="-XsvGEQzKtc_"
# # 3 Custom convolutional classifier – bonus!<a name="_3 custom convolutional classifier – bonus!"></a><a name="_3 custom convolutional classifier – bonus!"></a>
# + [markdown] colab_type="text" id="y4fJuVO8Ktc_"
# In the previous section we used a "canned" linear estimator that
# computed the predictions from the pixel values directly by logistic
# regression.
#
# Of course we know that the pixel values are not randomly distributed,
# but actually form a two-dimensional image and we can leverage our
# understanding of the data by using
# [2D convolutions](https://en.wikipedia.org/wiki/Multidimensional_discrete_convolution).
#
# We will still be using the `tf.estimator` interface, but this time we
# specify the computational graph that computes the predictions from the
# raw pixel values by hand, using 2D convolutions and max pooling (max pooling
# is used to reduce the convoluted image's spatial dimensions).
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 634, "status": "ok", "timestamp": 1526713963159, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="6zTvKT8lKtdB" outputId="2681517a-4550-4cb8-df9a-32c23684153f"
# %%writefile _derived/4_get_logits_img.py
# (Written into separate file for sharing with cloud code.)
# Define a function that computes "logits" from features.
# The "logits" are unbound numbers that will be used as the
# input to the softmax function (which is basically a sigmoid
# function extended to more than two classes). The more positive
# a logit, the closer to one the corresponding probability that
# is the output of the softmax.
# https://en.wikipedia.org/wiki/Softmax_function
def get_logits_img(features, n_classes, mode, params):
"""Computes logits for provided features.
Args:
features: A dictionary of tensors that are the features
and whose first dimension is batch (as returned by input_fn).
n_classes: Number of classes from which to predict (i.e. the number
of different values in the "labels" tensor returned by the
input_fn).
mode: A tf.estimator.ModeKeys.
params: Hyper parameters: "convs" specifying the configuration of the
convolutions, and "hidden" specifying the configuration of the
dense layers after the convolutions.
Returns:
The logits tensor with shape=[batch, n_classes].
"""
# The parameter "convs" specifies (kernel, stride, filters)
# of successive convolution layers.
convs = params.get('convs', ((10, 4, 32), (5, 4, 64)))
# The parameter "hidden" specifies the number of neurons of
# successive fully connected layers (after convolution).
hidden = params.get('hidden', (256,))
# The function tf.layers.conv2d expects the tensor to have format
# [batch, height, width, channels] -- since our "img_64" tensor
# has format [batch, height, width], we need to expand the tensor
# to get [batch, height, width, channels=1].
last_layer = tf.cast(tf.expand_dims(features['img_64'], axis=3), tf.float32)
# We start with dims=width=height=64 and filters=channels=1 and then
# successively reduce the number of dimensions while increasing the
# number of filters in every convolutional/maxpooling layer.
dim = 64
filters = 1
for kernel, stride, filters in convs:
conv = tf.layers.conv2d(
inputs=last_layer, filters=filters, kernel_size=[kernel, kernel],
padding='same', activation=tf.nn.relu)
last_layer = tf.layers.max_pooling2d(
inputs=conv, pool_size=[stride, stride], strides=stride)
dim //= stride
# "Flatten" the last layer to get shape [batch, *]
last_layer = tf.reshape(last_layer, [-1, filters * dim * dim])
# Add some fully connected layers.
for units in hidden:
dense = tf.layers.dense(inputs=last_layer, units=units,
activation=tf.nn.relu)
# Regularize using dropout.
training = mode == tf.estimator.ModeKeys.TRAIN
last_layer = tf.layers.dropout(inputs=dense, rate=0.4,
training=training)
# Finally return logits that is activation of neurons in last layer.
return tf.layers.dense(inputs=last_layer, units=n_classes)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 776, "status": "ok", "timestamp": 1526713966357, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="9TTuubJUKtdI" outputId="3b6484be-999b-479e-f17a-d7837dbb0b2d"
# %%writefile _derived/4_make_model_fn.py
# (Written into separate file for sharing with cloud code.)
# Warning : Boilerplate code cell...
# This cell defines a function that connects the "get_logits_fn"
# to the estimator interface and adds some useful output for our problem.
# By specifying different parameters, the same function can be reused
# in section 5 where we use a recurrent neural network to compute the
# logits.
def make_model_fn(get_logits_fn, n_classes):
"""Creates a model_fn.
Args:
get_logits_fn: Function that computes logits from features.
n_classes: Number of classes.
Returns:
A model_fn to be used with an estimator.
"""
def model_fn(features, labels, mode, params):
"""The model_fn is passed as an argument to the estimator.
Args:
features: Dictionary mapping feature names to feature tensors.
labels: Optional labels (`None` during inference).
mode: A `tf.estimator.ModeKeys`.
params: Optional dictionary of hyper parameters.
Returns:
A `tf.estimator.EstimatorSpec`.
"""
# Create logits from features using RNN.
logits = get_logits_fn(features, n_classes=n_classes, mode=mode,
params=params)
# Convert logits to probabilities.
probabilities = tf.nn.softmax(logits)
# Extract class with highest probability.
predictions = tf.argmax(probabilities, axis=1)
onehot_labels = loss = train_op = eval_metric_ops = None
if labels is not None:
onehot_labels = tf.one_hot(tf.squeeze(labels), n_classes)
loss = tf.losses.softmax_cross_entropy(onehot_labels, logits)
if mode == tf.estimator.ModeKeys.TRAIN:
# Compute loss.
global_step = tf.train.get_global_step()
# Minimize.
train_op = tf.train.AdamOptimizer().minimize(loss, global_step=global_step)
tf.summary.scalar('loss', loss)
# Output number of parameters for educational purposes.
trainable_params = 0
for var in tf.trainable_variables():
tf.logging.info('Variable "%s" : %s.', var.name,
var.get_shape().as_list())
trainable_params += np.prod(var.get_shape().as_list())
tf.logging.info('Total params : %d.', trainable_params)
if mode == tf.estimator.ModeKeys.EVAL:
# Report accuracy when evaluating.
eval_metric_ops = {
'accuracy': tf.metrics.accuracy(labels, predictions),
}
return tf.estimator.EstimatorSpec(
loss=loss,
mode=mode,
predictions={
'probabilities': probabilities,
'predictions': predictions,
},
export_outputs={
'prediction': tf.estimator.export.PredictOutput(outputs={
'probabilities': probabilities,
'predictions': predictions,
}),
},
train_op=train_op,
eval_metric_ops=eval_metric_ops,
)
return model_fn
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 105} colab_type="code" executionInfo={"elapsed": 1097, "status": "ok", "timestamp": 1526713969192, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="Tbi_n0ZiKtdL" outputId="0da313b2-461f-41a1-ed31-549798622449"
# %run -i _derived/4_get_logits_img.py
# %run -i _derived/4_make_model_fn.py
# Instead of creating a tf.estimator.LinearClassifier, we now create
# a tf.estimator.Estimator and specify our custom model_fn that will
# be used by the estimator to compute the predictions from the features.
run_config = tf.estimator.RunConfig(save_summary_steps=10)
model_fn = make_model_fn(get_logits_fn=get_logits_img, n_classes=len(classes))
cnn_estimator = tf.estimator.Estimator(model_fn=model_fn, config=run_config)
# -- from here on, the code is exactly the same as in the previous section ...
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 357} colab_type="code" executionInfo={"elapsed": 115350, "status": "ok", "timestamp": 1526714085496, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="VId2TE6gKtdN" outputId="6d066cc6-7fdd-498f-d0e6-76e8cce4efae"
# Note: You can get detailed information about the training by pointing a
# Tensorboard instance to the training directory.
#
# You start Tensorboard via a terminal, for example by clicking on "New"
# -> "Terminal" in the Jupyter directory view (this also works when
# Jupyter server is running inside a Docker image).
#
# The following command will start Tensorboard and show all experimental
# data below the "/tmp" directory (which is where new model directories
# are created if we do not specify a model path):
#
# $ tensorboard --logdir /tmp/
#
# And then view the Tensorboard in your browser:
# http://localhost:6006
# Bonus challenge (don't do this during the workshop) : Try to increase the
# number of training steps, batch_size, and/or parameters (hint: check out
# the "params" argument when creating the estimator in the previous cell)
# -- what accuracy can achieve with this convolutional network?
input_fn = make_input_fn('%s/train-*' % data_path, batch_size=100)
cnn_estimator.train(input_fn=input_fn, steps=100)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 4372, "status": "ok", "timestamp": 1526714134054, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="1Ua0CEAOKtdQ" outputId="73d20d21-1013-4391-e3eb-c5e963219aa9"
cnn_estimator.evaluate(input_fn=make_input_fn('%s/eval-*' % data_path),
steps=1)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 800} colab_type="code" executionInfo={"elapsed": 2563, "status": "ok", "timestamp": 1526714154116, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="vhzbioUkKtdV" outputId="f5490ce7-c53d-4d41-fb86-5b37a8870bbd"
input_fn = make_input_fn('%s/test-*' % data_path, batch_size=1)
show_predictions(cnn_estimator, predict_keys='probabilities')
# + [markdown] colab_type="text" id="KGNhs-ACKtdY"
# # 4 Keras – bonus!<a name="_4 keras – bonus!"></a><a name="_4 keras – bonus!"></a>
# + [markdown] colab_type="text" id="Qd2EksWCKtdZ"
# "Keras is a high-level neural networks API, written in Python and capable of
# running on top of TensorFlow, CNTK, or Theano. It was developed with a focus
# on enabling fast experimentation. Being able to go from idea to result with
# the least possible delay is key to doing good research."
# (from https://keras.io/)
#
# In this section we reconstruct the same convolutional model as in the last
# section using Keras and then convert the Keras model to an estimator so we
# can use the same input functions reading data in a streaming fashion from our
# data on disk.
#
# More useful links about Keras (not needed for this section):
#
# - https://keras.io/models/about-keras-models/
# - https://keras.io/models/sequential/
# - https://keras.io/models/model/
# - [Converting Keras model to Tensorflow estimator](https://cloud.google.com/blog/big-data/2017/12/new-in-tensorflow-14-converting-a-keras-model-to-a-tensorflow-estimator)
# - [Example using functional API with input_fn](https://github.com/keras-team/keras/blob/master/examples/mnist_tfrecord.py)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 374} colab_type="code" executionInfo={"elapsed": 922, "status": "ok", "timestamp": 1526714174789, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="_cWVQm43KtdZ" outputId="8357c73d-1aee-4aa9-cf80-37ab9ba4a218"
# Keras is part of Tensorflow
from tensorflow import keras
# The model is specified as a succession of layers.
# The layers here specify exactly the same model as in the previous section.
model = keras.models.Sequential([
# Note that we need to name the first layer (to match features, see next
# cell) and specify the input shape.
keras.layers.Conv2D(filters=32, kernel_size=10, activation='relu',
padding='same', name='firstlayer',
input_shape=(64, 64, 1)),
keras.layers.MaxPooling2D(pool_size=4),
keras.layers.Conv2D(filters=64, kernel_size=5, activation='relu',
padding='same'),
keras.layers.MaxPooling2D(pool_size=4),
keras.layers.Flatten(),
keras.layers.Dense(units=256, activation='relu'),
# We also need to name the output layer.
tf.keras.layers.Dense(units=len(classes), activation='softmax',
name='labels'),
])
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# Neat :-)
model.summary()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 309} colab_type="code" executionInfo={"elapsed": 115057, "status": "ok", "timestamp": 1526714293302, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="bDIiMXoGKtdd" outputId="907e2919-b5c9-466f-ed01-d85b313bde4b"
# Helper function to wrap input_fn().
def keras_make_input_fn(*args, **kwargs):
def wrapper():
input_fn = make_input_fn(*args, **kwargs)
features, labels = input_fn()
# We need to specify which feature is used as the input to
# which layer. In our case we have a single input feature
# that is read by layer "firstlayer". Note that we also need
# to add an additional dimension to get shape=(64, 64, 1).
features = {
'firstlayer_input': tf.expand_dims(features['img_64'], axis=3),
}
# Labels are expected in one_hot format.
labels = tf.one_hot(tf.squeeze(labels), 10)
return features, labels
return wrapper
# From now on we proceed exactly as before...
keras_estimator = keras.estimator.model_to_estimator(model)
input_fn = keras_make_input_fn('%s/train-*' % data_path, batch_size=100)
keras_estimator.train(input_fn=input_fn, steps=100)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 2740, "status": "ok", "timestamp": 1526714346211, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="x3-dlv-1Ktdg" outputId="10e5e77e-9221-4ef8-de07-e1a4b1775660"
keras_estimator.evaluate(input_fn=keras_make_input_fn('%s/eval-*' % data_path),
steps=1)
# + [markdown] colab_type="text" id="cGjez0EpKtdi"
# # 5 Recurrent neural network – bonus!
#
# So far we have been processing two dimensional image data. That
# data can be converted nicely into a dense tensor that is then the
# input layer of a linear classifier or convolutional network.
#
# If we want to process the raw stroke data, we need a different
# network architecture that reads in coordinate by coordinate, while
# updating its internal state and finally outputs a prediction --
# a **recurrent network**.
#
# Explaining the architecture for recurrent networks (specifically,
# we use a LSTM in this section) is out of the scope of this workshop,
# please read the following articles if you want to know more about the
# architecture:
#
# - http://colah.github.io/posts/2015-08-Understanding-LSTMs/
# - http://karpathy.github.io/2015/05/21/rnn-effectiveness/
#
# Using recurrent networks requires some tensor shape black magic and
# this is the main focus of this section.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 561} colab_type="code" executionInfo={"elapsed": 2036, "status": "ok", "timestamp": 1526714360421, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="MqoLvLIOKtdi" outputId="4cf370dc-10ad-43dc-bbc2-1e4afa4781fc"
# Load stroke data generated in 1_qd_data (bonus section):
stroke_data_path = '../data/dataset_stroke'
# !ls -lh $stroke_data_path
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 221} colab_type="code" executionInfo={"elapsed": 671, "status": "ok", "timestamp": 1526714405858, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="E8G33l_PKtdm" outputId="9a6a778f-ae79-4529-d9a2-c98aeb9f4cc4"
# Label classes...
classes = open('%s/labels.txt' % stroke_data_path).read().splitlines()
print('%d label classes:\n' % len(classes))
for i, label in enumerate(classes):
print('%d -> %s' % (i, label))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 119} colab_type="code" executionInfo={"elapsed": 844, "status": "ok", "timestamp": 1526714421000, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="-5d9psaAKtdr" outputId="81c29bcc-2976-4803-8aac-0cc54e3a5cdb"
# Some remarks on "sparse tensors": Conceptually, these
# tensors are an efficient representation of a tensor that
# has mostly "0" values. In the example below we initialize
# a sparse tensor by specifying only non-"0" values and then
# print the tensor as well as its dense representation.
# If we read "variable length" tensors from disk then these
# tensors will also be represented as sparse tensors (although
# the tensors might be "dense" in a sense that they have very
# few "0" values).
with tf.Graph().as_default():
# Only specify non-"0" values of 3x3 diagnal matrix.
sparse = tf.SparseTensor(indices=[[0, 0], [1, 1], [2, 2]], values=[1, 2, 3],
dense_shape=[3, 3])
# Convert to dense representation. Note that we can specify any
# "dense_shape" (resulting dense tensor is zero padded).
dense = tf.sparse_to_dense(sparse.indices, sparse.dense_shape,
sparse.values)
with tf.Session() as sess:
print(sparse.eval())
print(dense.eval())
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1006, "status": "ok", "timestamp": 1526714424544, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="ecQWAyARKtdw" outputId="24f2046b-9414-4d27-b33b-ef901983867d"
# %%writefile _derived/4_convert_sparse.py
# (Written into separate file for sharing with cloud code.)
# Now let's define a helper function that limits variable length
# sparse tensors to a maximum length and converts them to dense
# tensors. We need to convert sparse tensors to dense tensors before
# we can use them as input in the recurrent neural network.
def convert_sparse(sparse, max_len):
"""Converts batched sparse tensor to dense tensor with specified size.
Args:
sparse: tf.SparseTensor instance of shape=[n].
max_len: Truncates / zero-pads the dense tensor the specified max_len.
"""
# Convert to dense tensor.
dense = tf.sparse_to_dense(sparse.indices, sparse.dense_shape,
sparse.values)
# Discard values above max_len.
dense = dense[:max_len]
# Zero-pad if length < max_len.
dense = tf.pad(dense, [[0, max_len - tf.shape(dense)[0]]])
return dense
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 1069, "status": "ok", "timestamp": 1526714437371, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="SZ-8dOEVKtdz" outputId="efedacdf-e0d6-4164-9e70-f622fdf1a569"
# %run -i _derived/4_convert_sparse.py
with tf.Graph().as_default():
# Manually define sparse X-coordinates [1,2,3,4,5].
# Note that our stroke coordinate "sparse tensors" have a single dimension
# and do not contain any zeros at all...
stroke_x = tf.SparseTensor(
indices=[[0], [1], [2], [3], [4]],
values=[1, 2, 3, 4, 5],
dense_shape=[5])
# Extract both shorter and longer dense tensor.
dense_short = convert_sparse(stroke_x, max_len=3)
dense_long = convert_sparse(stroke_x, max_len=10)
with tf.Session() as sess:
print(dense_short.eval())
print(dense_long.eval())
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 865, "status": "ok", "timestamp": 1526714447805, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="553nkfckKtd2" outputId="78fd2ba5-65ef-4b50-9211-641d1405212c"
# %%writefile _derived/4_input_fn_stroke.py
# (Written into separate file for sharing with cloud code.)
# Because the data is stored in a different format (strokes instead of pixels)
# we need a new input_fn.
# Maximum number of points in concatenated strokes.
MAX_LEN = 256
# Because every drawing has a different number of points, we use "VarLenFeature"
# and not "FixedLenFeature" for the stroke data. This will create
# "SparseTensor".
feature_spec = {
'stroke_x': tf.VarLenFeature(dtype=tf.float32),
'stroke_y': tf.VarLenFeature(dtype=tf.float32),
'stroke_z': tf.VarLenFeature(dtype=tf.float32),
'stroke_len': tf.FixedLenFeature([], tf.int64),
'label': tf.FixedLenFeature([], tf.int64),
}
def parse_example_stroke(serialized_example):
features = tf.parse_single_example(serialized_example, feature_spec)
label = features.pop('label')
# The we create a 'stroke' tensor with shape [3, MAX_LEN] where the first
# dimension indicates whether the values are X, Y, or Z coordinates.
stroke = tf.stack([
convert_sparse(features['stroke_x'], max_len=MAX_LEN),
convert_sparse(features['stroke_y'], max_len=MAX_LEN),
convert_sparse(features['stroke_z'], max_len=MAX_LEN),
])
# Also truncate the "stroke_len" to MAX_LEN if needed.
stroke_len = tf.minimum(tf.cast(MAX_LEN, tf.int64), features['stroke_len'])
return dict(stroke=stroke, stroke_len=stroke_len), label
# Copied from above Section "1.2 Reading the data using Tensorflow"
def make_input_fn_stroke(files_pattern, batch_size=100):
def input_fn():
dataset = tf.data.TFRecordDataset(tf.gfile.Glob(files_pattern))
dataset = dataset.map(parse_example_stroke).batch(batch_size)
dataset = dataset.shuffle(buffer_size=5*batch_size).repeat()
features, labels = dataset.make_one_shot_iterator().get_next()
return features, labels
return input_fn
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 206} colab_type="code" executionInfo={"elapsed": 1407, "status": "ok", "timestamp": 1526714454895, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="CGGUHXF9Ktd4" outputId="ad316dd0-df55-4735-aeea-74c3c3877c81"
# Read some examples with above input_fn and plot data.
# %run -i _derived/4_input_fn_stroke.py
# (Modified code from section 1.1 and 1.2)
def show_stroke_img(stroke, label, ax=None):
"""Plots stroke data.
Args:
stroke: Array of shape=[3, n] where the second dimension
is time and the first dimension indicates X/Y coordinates
and Z-dimension that is set to 1 when a stroke ends and
0 otherwise (the array actually represents an array of
concatenated strokes and the Z-dimension is needed to tell
the individual strokes apart).
"""
ax = ax if ax else pyplot.gca()
xy = stroke[:2, :].cumsum(axis=1)
ax.plot(*xy)
# Plot all the strokes, including connecting line between strokes.
pxy = xy[:, stroke[2] != 0]
# Red dots mark end of individual strokes.
ax.plot(pxy[0], pxy[1], 'ro')
ax.set_xticks([])
ax.set_yticks([])
ax.set_title(label)
batch_size = 5
input_fn = make_input_fn_stroke(files_pattern='%s/train-*' % stroke_data_path,
batch_size=batch_size)
with tf.Graph().as_default():
features, labels = input_fn()
with tf.train.MonitoredSession() as sess:
strokes_, labels_ = sess.run([features['stroke'], labels])
pyplot.figure(figsize=(10, 2))
for i in range(batch_size):
ax = pyplot.subplot(1, batch_size, i+1)
show_stroke_img(strokes_[i], classes[labels_[i]], ax)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 897, "status": "ok", "timestamp": 1526714457639, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="RCeOIUs_Ktd6" outputId="c1ec261b-db16-444d-c28b-02b4c2a95df7"
# %%writefile _derived/4_get_nth.py
# (Written into separate file for sharing with cloud code.)
# Another helper function: This function will return the "nth element"
# with different "n" for every element in the batch.
# We will later need this function to get the prediction in the output
# of the dynamic_rnn.
def get_nth(tensor, ns, last_dim):
"""Tensor has shape [batch_size, max_len, last_dim]."""
shape = tf.shape(tensor)
batch_size, max_len = shape[0], shape[1]
# Flatten first two dimensions.
tensor = tf.reshape(tensor, [-1, last_dim])
# Calculate indices within flattened tensor.
idxs = tf.range(0, batch_size) * max_len + (tf.cast(ns, tf.int32) - 1)
# Return nth elements.
# TODO get rid of error UserWarning : https://stackoverflow.com/questions/35892412
return tf.gather(tensor, idxs)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 968, "status": "ok", "timestamp": 1526714464266, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="VUgsf7mUKtd9" outputId="0159569a-c363-4551-fb1d-ff6ba708a8f1"
# %run -i _derived/4_get_nth.py
with tf.Graph().as_default(), tf.Session(graph=tf.Graph()) as sess:
# Define a batch with shape=[batch_size=2, max_len=5, last_dim=2]
batch = tf.constant([
# First tensor in batch (shape=[5, 2])
[[1, 1], [2, 2], [3, 3], [4, 4], [5, 5]],
# Second tensor in batch (shape=[5, 2])
[[-1, -1], [-2, -2], [-3, -3], [-4, -4], [-5, -5]],
])
# Return the second and third element.
lens = tf.constant([2, 3], dtype=tf.int64)
print(get_nth(batch, lens, last_dim=2).eval())
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 841, "status": "ok", "timestamp": 1526714466948, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="Mhz9hKA1Ktd_" outputId="7c18203f-fea7-4c45-fb29-fcd0d400e670"
# %%writefile _derived/4_get_logits_stroke.py
# (Written into separate file for sharing with cloud code.)
# This function creates creates the logits fro the stroke features.
def get_logits_stroke(features, n_classes, mode, params):
"""Computes logits for provided features.
Args:
features: A dictionary of tensors that are the features
and whose first dimension is batch (as returned by input_fn).
n_classes: Number of classes from which to predict (i.e. the number
of different values in the "labels" tensor returned by the
input_fn).
mode: A tf.estimator.ModeKeys.
params: Hyper parameters: "cell_size" specifying the state size of
the LSTM cells, and "hidden" specifying the configuration of the
dense layers after recurrent network.
Returns:
The logits tensor with shape=[batch, n_classes].
"""
cell_size = params.get('cell_size', 256)
hidden = params.get('hidden', ())
# First we convert our data from "coords major" to "time major",
# as required by the dynamic_rnn API.
# [batch, coords, time] -> [batch, time, coords]
stroke = tf.transpose(features['stroke'], perm=[0, 2, 1])
stroke_len = features['stroke_len']
# Construct a bi-directional dynamic recurrent NN with LSTM
# cells.
outputs, states = tf.nn.bidirectional_dynamic_rnn(
cell_fw=tf.nn.rnn_cell.LSTMCell(cell_size),
cell_bw=tf.nn.rnn_cell.LSTMCell(cell_size),
inputs=stroke,
sequence_length=stroke_len,
dtype=tf.float32,
)
# Use helper function from last cell to extract RNN output values.
outputs = tf.concat((get_nth(outputs[0], stroke_len, last_dim=cell_size),
get_nth(outputs[1], stroke_len, last_dim=cell_size)),
axis=1)
# Add fully connected layers on top.
for units in hidden:
outputs = tf.layers.dense(inputs=outputs, units=units,
activation=tf.nn.relu)
# Logits are activations of last fully connected layer.
return tf.layers.dense(inputs=outputs, units=n_classes)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 751} colab_type="code" executionInfo={"elapsed": 54520, "status": "ok", "timestamp": 1526714528479, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "115804658599019978174"}, "user_tz": -120} id="OHZEIewvKteB" outputId="3a96161a-349a-40bc-fa39-4c41db18c464"
# %run -i _derived/4_get_logits_stroke.py
# Same as before : Create estimator, train, evaluate.
# Note that we need *much* more time and/or CPU power to get
# decent results with the RNN, but eventually it will outperform
# the convolutional classifier...
# Use make_model_fn from section "3 Custom convolutional classifier"
model_fn = make_model_fn(get_logits_fn=get_logits_stroke, n_classes=len(classes))
config = tf.estimator.RunConfig(save_summary_steps=1)
rnn_estimator = tf.estimator.Estimator(model_fn=model_fn, config=config)
# We use smaller batch size to keep memory usage below ~1G (the
# RNN has a large memory footprint because of the temporal unrolling).
# To get good performance with this type of network we need much longer
# training, which is impractical in a notebook. See next notebook
# "5_qd_cloud" that describes how to train the network on Google Cloud
# Environment using Cloud ML...
input_fn = make_input_fn_stroke('%s/train-*' % stroke_data_path, batch_size=10)
rnn_estimator.train(input_fn=input_fn, steps=100)
input_fn = make_input_fn_stroke('%s/eval-*' % stroke_data_path, batch_size=10)
rnn_estimator.evaluate(input_fn=input_fn, steps=10)
# + [markdown] colab_type="text" id="OkpS7QOqKteH"
# # A References
#
# - https://www.tensorflow.org/versions/master/get_started/feature_columns
# - https://arxiv.org/abs/1704.03477 : A Neural Representation of Sketch Drawings
# - https://cloud.google.com/blog/big-data/2017/01/learn-tensorflow-and-deep-learning-without-a-phd : Nice tutorial explaining convolutions, recurrent networks, and some deep learning tricks – Challenge: try to improve the models in this notebook with the techniques described in this presentation!
| extras/amld/notebooks/solutions/4_qd_estimator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ecdf-guide
# language: python
# name: ecdf-guide
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# -
# # Introduction
#
# In this notebook, I am going to compare the use of histograms and ECDFs in interpreting data.
#
# Histograms are probably the most popular distribution visualization tool. But it comes fraught with one major hidden danger that can be decieving: binning bias.
#
# Using the default binning in matplotlib, we get the following plot for draws from a standard normal distribution:
# +
# Set random seed for deterministic reproducibility.
np.random.seed(42)
# Generate a single Gaussian distribution, but with
data = np.random.normal(loc=0, scale=1, size=(20))
plt.hist(data)
plt.show()
# -
# If we increased the number of bins, each bin gets smaller:
plt.hist(data, bins=100)
plt.show()
# Suddenly, this standard normal distribution looks a bit more like a Laplace distribution centered on -0.5 - which we know is incorrect!
#
# Moreover, it's difficult to intuit important statistical properties of the data: e.g. the range (min-max), inter-quartile range (25th-75th percentile), the median. They are simply not immediately visible from the histogram.
#
# What if, instead, we looked at the ECDF of the data?
def ecdf(data):
x, y = np.sort(data), np.arange(1, len(data)+1) / len(data)
return x, y
x, y = ecdf(data)
p = [25, 50, 75]
fractions = np.array(p)/100
percs = np.percentile(data, p, interpolation='lower')
plt.scatter(x, y, label='data')
plt.vlines(percs, min(y), fractions, linestyles='--')
plt.hlines(fractions, min(x), percs, linestyles='--')
# A few things are immediately obvious! We can see read off the range approx. $(-1.0, 0.5)$ as containing approximately 50% of the probability density associated with the data.
x, y = ecdf(data)
plt.scatter(x, y, label='data')
x_n, y_n = ecdf(np.random.normal(0, 1, size=20000))
plt.plot(x_n, y_n, color='orange', label='normal', ls='--')
x_l, y_l = ecdf(np.random.laplace(-0.5, 1, size=20000))
plt.plot(x_l, y_l, color='red', label='laplace', ls='--')
plt.legend()
plt.show()
# By comparing the ECDF of the data against a standard normal CDF and a shifted laplace distribution, we can immediately see that the shifted laplace might have a poorer distributional fit compared to the standard normal distribution.
# ## Bimodal Distributions
#
# Bimodal distributions can be easily discovered in an ECDF. It manifests as a plateau in the data.
#
# Let's create a bimodal gaussian mixture, which are well-separated.
# +
mode1 = np.random.normal(loc=0, scale=1, size=1000)
mode2 = np.random.normal(loc=4, scale=1, size=100)
data_bimodal = np.concatenate([mode1, mode2])
fig = plt.figure(figsize=(6, 4))
ax1 = fig.add_subplot(211)
ax2 = fig.add_subplot(212)
ax1.hist(data_bimodal)
ax1.set_title('histogram')
x, y = ecdf(data_bimodal)
ax2.set_title('ecdf')
ax2.plot(x, y)
dy = y[1:] - y[:-1]
idxs = np.argsort(dy)
idxs = idxs[idxs > 820]
ax2.vlines(x=x[idxs[0]], ymin=min(y), ymax=y[idxs[0]])
ax2.hlines(y=y[idxs[0]], xmin=min(x), xmax=x[idxs[0]])
plt.tight_layout()
plt.show()
# -
# By going to the first plateau observed, we can roughly guess a 9 or 10-to-one ratio of mixture components.
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Assignment_3_Task_3
########
#Assignment is done by - Md <NAME> (220100676), <EMAIL>
######## <NAME> (220203317), <EMAIL>
# +
import matplotlib.pyplot as plt
import pandas as pd
import csv
plt.style.use('bmh')
# +
#Importing dataset in dataframe
df_height = pd.read_csv('data3.csv')
df_height = pd.DataFrame(df_height)
df_height
# +
#Plotting the dataset with a histogram
x = df_height['height']
plt.xlabel('height', fontsize=25)
plt.title('Height of People in Kung San', fontsize=20)
plt.xlabel('Height', fontsize=15)
plt.ylabel('Number of People', fontsize=15)
plt.hist(x)
plt.show()
| Assignment 3/Assignment_ 3_Task_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [Boosters] Raiffeisen Data Cup. Baseline
# Общий подход:
# - Добавляем к каждой транзакции столбец: is_work (если транзакция находится в пределах 0.02 от дома клиента)
# - Добавляем к каждой транзакции столбец: is_home (если транзакция находится в пределах 0.02 от работы клиента)
# - Обучаем классификатор предсказывающий вероятность (is_home == 1) для транзакции
# - Обучаем классификатор предсказывающий вероятность (is_work == 1) для транзакции
#
# Точность определения местоположения:
# - для классификатора is_home: ~3x%
# - для классификатора is_work: ~2x%
# - общая оценка на Public Leaderboard: ???
#
# Примечание
# * Требуется Python версии 3.5
# * Требуется библиотека xgboost (для обучения использовалась xgboost версии 0.7.post3)
# * Требуются файлы: test_set.csv, train_set.csv в одном каталоге с данным скриптом
# * Требования к памяти: должно работать с 2Гб свободного RAM
# * Время работы: ~3 минуты (тестировалось на процессоре Intel Core i7-4770)
# +
# %load_ext autoreload
# %autoreload 2
import sys
MODULES_PATH = '../code/'
if MODULES_PATH not in sys.path:
sys.path.append(MODULES_PATH)
import mfuncs
import pandas as pd
import numpy as np
from tqdm import tqdm
tqdm.pandas()
pd.options.display.max_columns = 1000
import lightgbm as lgb
from sklearn.neighbors import NearestNeighbors
# %pylab inline
# +
# Определим типы колонок для экономии памяти
dtypes = {
'transaction_date': str,
'atm_address': str,
'country': str,
'city': str,
'amount': np.float32,
'currency': np.float32,
'mcc': str,
'customer_id': str,
'pos_address': str,
'atm_address': str,
'pos_adress_lat': np.float32,
'pos_adress_lon': np.float32,
'pos_address_lat': np.float32,
'pos_address_lon': np.float32,
'atm_address_lat': np.float32,
'atm_address_lon': np.float32,
'home_add_lat': np.float32,
'home_add_lon': np.float32,
'work_add_lat': np.float32,
'work_add_lon': np.float32,
}
# для экономии памяти будем загружать только часть атрибутов транзакций
usecols_train = ['customer_id','transaction_date','amount','country', 'city', 'currency', 'mcc', 'pos_adress_lat', 'pos_adress_lon', 'atm_address_lat', 'atm_address_lon','home_add_lat','home_add_lon','work_add_lat','work_add_lon']
usecols_test = ['customer_id','transaction_date','amount','country', 'city', 'currency', 'mcc', 'pos_address_lat', 'pos_address_lon', 'atm_address_lat', 'atm_address_lon']
# -
# ## Читаем train_set, test_set, соединяем в один датасет
# +
dtypes = {
'transaction_date': str,
'atm_address': str,
'country': str,
'city': str,
'amount': np.float32,
'currency': np.float32,
'mcc': str,
'customer_id': str,
'pos_address': str,
'atm_address': str,
'pos_adress_lat': np.float32,
'pos_adress_lon': np.float32,
'pos_address_lat': np.float32,
'pos_address_lon': np.float32,
'atm_address_lat': np.float32,
'atm_address_lon': np.float32,
'home_add_lat': np.float32,
'home_add_lon': np.float32,
'work_add_lat': np.float32,
'work_add_lon': np.float32,
}
rnm = {
'atm_address_lat': 'atm_lat',
'atm_address_lon': 'atm_lon',
'pos_adress_lat': 'pos_lat',
'pos_adress_lon': 'pos_lon',
'pos_address_lat': 'pos_lat',
'pos_address_lon': 'pos_lon',
'home_add_lat': 'home_lat',
'home_add_lon': 'home_lon',
'work_add_lat': 'work_lat',
'work_add_lon': 'work_lon',
}
# +
df_train = pd.read_csv('../data/train_set.csv', dtype=dtypes)
df_test = pd.read_csv('../data/test_set.csv', dtype=dtypes)
df_train.rename(columns=rnm, inplace=True)
df_test.rename(columns=rnm, inplace=True)
# -
# удалим чувак с множественными адресами
print(df_train.shape)
gb = df_train.groupby('customer_id')['work_lat'].agg('nunique')
cid_incorrect = gb[gb == 2].index
df_train = df_train[~df_train.customer_id.isin(cid_incorrect.values)]
print(df_train.shape)
gb = df_train.groupby('customer_id')['home_lat'].agg('nunique')
cid_incorrect = gb[gb == 2].index
df_train = df_train[~df_train.customer_id.isin(cid_incorrect.values)]
print(df_train.shape)
# +
# соединяем test/train в одном DataFrame
df_train['is_train'] = np.int32(1)
df_test['is_train'] = np.int32(0)
df_all = pd.concat([df_train, df_test])
del df_train, df_test
# -
# ### Обрабатываем дату транзакции и категориальные признаки
# +
df_all['currency'] = df_all['currency'].fillna(-1).astype(np.int32)
df_all['mcc'] = df_all['mcc'].apply(lambda x: int(x.replace(',', ''))).astype(np.int32)
df_all['city'] = df_all['city'].factorize()[0].astype(np.int32)
df_all['country'] = df_all['country'].factorize()[0].astype(np.int32)
# удаляем транзакции без даты
df_all = df_all[~df_all['transaction_date'].isnull()]
df_all['transaction_date'] = pd.to_datetime(df_all['transaction_date'], format='%Y-%m-%d')
# -
# ### Фичи для даты
df_all['month'] = df_all.transaction_date.dt.month
df_all['day'] = df_all.transaction_date.dt.day
df_all['dayofyear'] = df_all.transaction_date.dt.dayofyear
df_all['dayofweek'] = df_all.transaction_date.dt.dayofweek
# ### Приводим адрес транзакции для pos и atm-транзакций к единообразному виду
# Просто объединяем в одну колонку и добавляем фичу - это атм или пос
dfs = []
for cid in tqdm(df_all.customer_id.unique()):
df_an = df_all[df_all.customer_id == cid]
df_an = mfuncs.add_dist_to_neighbours(df_an)
dfs.append(df_an)
df_knn = pd.concat(dfs)
df_knn.head()
df_knn['pos2pos_1', 'pos2pos_2', 'atm2pos_1', 'atm2pos_2', 'pos2atm_1',
'pos2atm_2', 'pos2atm_1', 'pos2atm_2']
df_knn.to_csv('../data/df_knn.csv', index=None)
df_all = df_knn.copy()
# +
df_all['is_atm'] = (~df_all['atm_lat'].isnull()).astype(np.int8)
df_all['is_pos'] = (~df_all['pos_lat'].isnull()).astype(np.int8)
df_all['add_lat'] = df_all['atm_lat'].fillna(0) + df_all['pos_lat'].fillna(0)
df_all['add_lon'] = df_all['atm_lon'].fillna(0) + df_all['pos_lon'].fillna(0)
df_all.drop(['atm_lat','atm_lon','pos_lat','pos_lon'], axis=1, inplace=True)
df_all = df_all[~((df_all['add_lon'] == 0) & (df_all['add_lon'] == 0))]
# -
# ### Генерируем признаки is_home, is_work
# TODO: удалить чуваков у которых несколько домов
# +
lat = df_all['home_lat'] - df_all['add_lat']
lon = df_all['home_lon'] - df_all['add_lon']
df_all['is_home'] = (np.sqrt((lat ** 2) + (lon ** 2)) <= 0.02).astype(np.int8)
df_all['has_home'] = (~df_all['home_lon'].isnull()).astype(np.int8)
lat = df_all['work_lat'] - df_all['add_lat']
lon = df_all['work_lon'] - df_all['add_lon']
df_all['is_work'] = (np.sqrt((lat ** 2) + (lon ** 2)) <= 0.02).astype(np.int8)
df_all['has_work'] = (~df_all['work_lon'].isnull()).astype(np.int8)
df_all.drop(['work_lat','work_lon','home_lat','home_lon'], axis=1, inplace=True)
# -
# ### Генерируем категориальный признак для адреса
df_all['address'] = df_all['add_lat'].apply(lambda x: "%.02f" % x) + ';' + df_all['add_lon'].apply(lambda x: "%.02f" % x)
df_all['address'] = df_all['address'].factorize()[0].astype(np.int32)
# ### Генерируем несколько абонентских фич
# +
# количество транзакций каждого клиента
df_all = df_all.merge(df_all.groupby('customer_id')['amount'].count().reset_index(name='cid_trans_count'), how='left')
df_all['cid_trans_count'] = df_all['cid_trans_count'].astype(np.int32)
df_all = df_all.merge(df_all.groupby(['customer_id','address'])['amount'].count().reset_index(name='cid_add_trans_count'),
how='left')
df_all['cid_add_trans_count'] = df_all['cid_add_trans_count'].astype(np.int32)
# какая часть транзакций клиента приходится на данный адрес
# TODO: БОЛЬШЕ ТАКИХ ФИЧ
df_all['ratio1'] = df_all['cid_add_trans_count'] / df_all['cid_trans_count']
# -
# ## Мои фичи
df_gb[['amount', 'add_lat', 'add_lon']].agg(['mean', 'max', 'min'])
df_all[['customer_id','amount', 'add_lat', 'add_lon']]
df_gb['amount', 'add_lat', 'add_lon'].agg(['mean', 'max', 'min'])
df_all.reset_index(inplace=True, drop=True)
df_all[['customer_id','amount', 'add_lat', 'add_lon']].groupby('customer_id').agg('max')
# добавим признаки после групбая
df_gb = df_all[['customer_id','amount', 'add_lat', 'add_lon']].groupby('customer_id')
coord_stat_df = df_gb.agg(['mean', 'max', 'min'])
coord_stat_df['transactions_per_user'] = df_gb.agg('size')
coord_stat_df.columns = ['_'.join(col).strip() for col in coord_stat_df.columns.values]
coord_stat_df.reset_index(inplace=True)
df_all = pd.merge(df_all, coord_stat_df, on='customer_id', how='left')
cols = ['add_lat', 'add_lon']
types = ['min', 'max', 'mean']
for c in cols:
for t in types:
df_all['{}_ratio_{}'.format(c, t)] = np.abs(df_all[c] / df_all['{}_{}'.format(c, t)])
df_all = pd.concat([df_all, pd.get_dummies(df_all['mcc'], prefix='mcc')], axis=1)
del df_all['mcc']
# # LightGBM
df_all = df_all.loc[:,~df_all.columns.duplicated()]
# +
from sklearn.model_selection import train_test_split
ys = ['is_home', 'is_work']
drop_cols = ['atm_address', 'customer_id', 'pos_address', 'terminal_id', 'transaction_date',
'is_home' ,'has_home', 'is_work', 'has_work', 'is_train']
drop_cols += ['pred:is_home', 'pred:is_work']
y_cols = ['is_home', 'is_work']
usecols = df_all.drop(drop_cols, 1, errors='ignore').columns
# +
params = {
'objective': 'binary',
'num_leaves': 63,
'learning_rate': 0.01,
'metric' : 'binary_logloss',
'feature_fraction': 0.8,
'bagging_fraction': 0.8,
'bagging_freq': 1,
'num_threads': 12,
'verbose': 0,
}
model = {}
# +
y_col = 'is_home'
cust_train = df_all[df_all['is_train']==1].groupby('customer_id')[y_col.replace('is_','has_')].max()
cust_train = cust_train[cust_train > 0].index
cust_train, cust_valid = train_test_split(cust_train, test_size=0.2, shuffle=True, random_state=111)
df_train = pd.DataFrame(cust_train, columns=['customer_id']).merge(df_all, how='left')
df_valid = pd.DataFrame(cust_valid, columns=['customer_id']).merge(df_all, how='left')
lgb_train = lgb.Dataset(df_train[usecols], df_train[y_col])
lgb_valid = lgb.Dataset(df_valid[usecols], df_valid[y_col])
gbm_h = lgb.train(params,
lgb_train,
valid_sets=[lgb_valid],
num_boost_round=2000,
verbose_eval=30,
early_stopping_rounds=300)
model[y_col] = gbm_h
# +
y_col = 'is_work'
cust_train = df_all[df_all['is_train']==1].groupby('customer_id')[y_col.replace('is_','has_')].max()
cust_train = cust_train[cust_train > 0].index
cust_train, cust_valid = train_test_split(cust_train, test_size=0.2, shuffle=True, random_state=111)
df_train = pd.DataFrame(cust_train, columns=['customer_id']).merge(df_all, how='left')
df_valid = pd.DataFrame(cust_valid, columns=['customer_id']).merge(df_all, how='left')
lgb_train = lgb.Dataset(df_train[usecols], df_train[y_col])
lgb_valid = lgb.Dataset(df_valid[usecols], df_valid[y_col])
gbm_w = lgb.train(params,
lgb_train,
valid_sets=[lgb_valid],
num_boost_round=2000,
verbose_eval=30,
early_stopping_rounds=300)
model[y_col] = gbm_w
# -
lgb.plot_importance(gbm_w, max_num_features=15)
# +
def _best(x):
ret = None
for col in ys:
pred = ('pred:%s' % col)
if pred in x:
i = (x[pred].idxmax())
cols = [pred, 'add_lat', 'add_lon']
if col in x:
cols.append(col)
tmp = x.loc[i,cols]
tmp.rename({
'add_lat':'%s:add_lat' % col,
'add_lon':'%s:add_lon' % col,
}, inplace = True)
if ret is None:
ret = tmp
else:
ret = pd.concat([ret, tmp])
return ret
def predict_proba(dt, ys=['is_home', 'is_work']):
for col in ys:
pred = ('pred:%s' % col)
dt[pred] = model[col].predict(dt[usecols])
return dt.groupby('customer_id').apply(_best).reset_index()
def score(dt, ys=['is_home', 'is_work']):
dt_ret = predict_proba(dt, ys)
mean = 0.0
for col in ys:
col_mean = dt_ret[col].mean()
mean += col_mean
if len(ys) == 2:
mean = mean / len(ys)
return mean
# +
print ("Train accuracy:", score(df_train, ys=['is_home']))
print ("Test accuracy:", score(df_valid, ys=['is_home']))
print ("Train accuracy:", score(df_train, ys=['is_work']))
print ("Test accuracy:", score(df_valid, ys=['is_work']))
# -
# # Predict
# +
cust_test = df_all[df_all['is_train'] == 0]['customer_id'].unique()
df_test = pd.DataFrame(cust_test, columns = ['customer_id']).merge(df_all, how = 'left')
df_test = predict_proba(df_test)
df_test.rename(columns = {
'customer_id':'_ID_',
'is_home:add_lat': '_HOME_LAT_',
'is_home:add_lon': '_HOME_LON_',
'is_work:add_lat': '_WORK_LAT_',
'is_work:add_lon': '_WORK_LON_'}, inplace = True)
df_test = df_test[['_ID_', '_WORK_LAT_', '_WORK_LON_', '_HOME_LAT_', '_HOME_LON_']]
df_test.head()
# -
# # Формируем submission-файл
# +
# Заполняем пропуски
df_ = pd.read_csv('../data/test_set.csv', dtype=dtypes, usecols=['customer_id'])
submission = pd.DataFrame(df_['customer_id'].unique(), columns=['_ID_'])
submission = submission.merge(df_test, how='left').fillna(0)
# Пишем файл submission
submission.to_csv('../submissions/base_2_47_32.csv', index=None)
| Raif/notebooks/Baseline_my.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reproduce Figure 1
# If you haven't seen it already, take a look at other tutorials to setup and install the progressive learning package `Installation-and-Package-Setup-Tutorial.ipynb`
#
# # Analyzing the UncertaintyForest Class by Reproducing Figure 1
# ## *Goal: Run the UncertaintyForest class to produce the results from Figure 1*
# *Note: Figure 1 refers to Figure 1 from [this paper](https://arxiv.org/pdf/1907.00325.pdf)*
# ### First, we'll import the necessary packages that will be required
# +
import numpy as np
from sklearn.ensemble import RandomForestClassifier
from sklearn.calibration import CalibratedClassifierCV
from proglearn.forest import UncertaintyForest
import sys
sys.path.append('../../docs/tutorials/functions')
from unc_forest_tutorials_functions import generate_data, estimate_posterior, plot_posterior, plot_variance, plot_fig1
# -
# ### Now, we'll specify some parameters
# Here are the "Real Parameters"
n = 6000 # number of data points
mean = 1 # mean of the data
var = 1 # variance of the data
num_trials = 100 # number of trials to run
X_eval = np.linspace(-2, 2, num = 30).reshape(-1, 1) # the evaluation span (over X) for the plot
n_estimators = 300 # the number of estimators
num_plotted_trials = 10 # the number of "fainter" lines to be displayed on the figure
# ### Now, we'll specify which learners we'll compare. Figure 1 uses three different learners specified below.
# +
# Algorithms used to produce figure 1
algos = [
{
'instance': RandomForestClassifier(n_estimators = n_estimators),
'label': 'CART',
'title': 'CART Forest',
'color': "#1b9e77",
},
{
'instance': CalibratedClassifierCV(base_estimator=RandomForestClassifier(n_estimators = n_estimators // 5),
method='isotonic',
cv = 5),
'label': 'IRF',
'title': 'Isotonic Reg. Forest',
'color': "#fdae61",
},
{
'instance': UncertaintyForest(n_estimators = n_estimators, tree_construction_proportion = 0.4, kappa = 3.0),
'label': 'UF',
'title': 'Uncertainty Forest',
'color': "#F41711",
},
]
# Plotting parameters
parallel = True
# -
# ### Now, we'll run the code to obtain the results that will be displayed in Figure1
# This is the code that actually generates data and predictions.
for algo in algos:
algo['predicted_posterior'] = estimate_posterior(algo, n, mean, var, num_trials, X_eval, parallel = parallel)
# ### Finally, create figure 1.
plot_fig1(algos, num_plotted_trials, X_eval)
| benchmarks/uf_posterior_visualization/uncertaintyforest_fig1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Replaying the Interaction Dataset
#
# This notebook shows how to use the interaction dataset in bark. Details on the dataset: https://arxiv.org/abs/1910.03088
from bark.runtime.scenario.scenario_generation.interaction_dataset_scenario_generation import \
InteractionDatasetScenarioGeneration
from bark.runtime.commons.parameters import ParameterServer
from bark.runtime.viewer.matplotlib_viewer import MPViewer
from bark.runtime.viewer.video_renderer import VideoRenderer
import os
import os.path
import argparse
import matplotlib.pyplot as plt
# Make sure you have access to the repository `interaction_dataset_fortiss_internal` and make it available by commenting out the command
# ```
# git_repository(
# name = "interaction_dataset_fortiss_internal",
# commit = "<PASSWORD>",
# remote = "https://git.fortiss.org/autosim/interaction_dataset"
# )
# ```
#
# in the top level Workspace file.
#
# Then, start the notebook server with `bazel run docs/tutorials:run --define interaction_dataset_present=true`
# Here, we specify which slize of the dataset shall be loaded. **Note that the files have to exist!** The dataset itself is not included in bark! We here specify the parameters:
# - The map: "MapFilename"
# - The vehicle trajectories: "TrackFilename"
# - Which vehicles: "TrackIds"
# - Start time: "StartTs"
# - End Time: "EndTs"
# - Who's treated as ego agent (aka. where are metrics evaluated): "EgoTrackId"
#
# Lets define the secnario:
param_server = ParameterServer()
param_server["Scenario"]["Generation"]["InteractionDatasetScenarioGeneration"]["MapFilename"] = os.path.expanduser('~') +"/bark-simulator/interaction_dataset_fortiss_internal/DR_DEU_Merging_MT/map/DR_DEU_Merging_MT_v01_shifted.xodr"
param_server["Scenario"]["Generation"]["InteractionDatasetScenarioGeneration"]["TrackFilename"] = os.path.expanduser('~') +"/bark-simulator/interaction_dataset_fortiss_internal/DR_DEU_Merging_MT/tracks/vehicle_tracks_013.csv"
param_server["Scenario"]["Generation"]["InteractionDatasetScenarioGeneration"]["TrackIds"] = [63,64,65,66,67,68]
param_server["Scenario"]["Generation"]["InteractionDatasetScenarioGeneration"]["StartTs"] = 232000
param_server["Scenario"]["Generation"]["InteractionDatasetScenarioGeneration"]["EndTs"] = 259000
param_server["Scenario"]["Generation"]["InteractionDatasetScenarioGeneration"]["EgoTrackId"] = 65
# Create the scenario
scenario_generation = InteractionDatasetScenarioGeneration(num_scenarios=1, random_seed=0, params=param_server)
scenario = scenario_generation.create_single_scenario()
# Initialize
sim_step_time = 0.2
sim_time_steps = 5
# +
# TODO maybe someone finds a way to reuse one plot
# Run the scenario in a loop
world_state = scenario.GetWorldState()
for _ in range(0, sim_time_steps):
world_state.PlanAgents(sim_step_time)
fig = plt.figure(figsize=[10, 10])
viewer = MPViewer(params=param_server, use_world_bounds=True, axis=fig.gca())
viewer.drawWorld(world_state, scenario._eval_agent_ids)
world_state.DoExecution(sim_step_time)
# -
# # Replacing an Agent
#
# Based on the above, we replace one agent with another agent model. For this, we just add a parameter to the server. Here we chose the replate the agent 67 with an IDM with standard parameters.
# Insert IDM Agent
param_server_idm = param_server
param_server_idm["Scenario"]["Generation"]["InteractionDataset"]["BehaviorModel"] = {"67":"BehaviorIDMClassic"}
# Override the scenario
scenario_generation_idm = InteractionDatasetScenarioGeneration(num_scenarios=1, random_seed=0, params=param_server_idm)
scenario = scenario_generation_idm.create_single_scenario()
# +
# TODO maybe someone finds a way to reuse one plot
# Run the scenario
world_state = scenario.GetWorldState()
for _ in range(0, sim_time_steps):
world_state.PlanAgents(sim_step_time)
fig = plt.figure(figsize=[10, 10])
viewer = MPViewer(params=param_server, use_world_bounds=True, axis=fig.gca())
viewer.drawWorld(world_state, scenario._eval_agent_ids)
world_state.DoExecution(sim_step_time)
# -
# That's it. We now have replayed one slice of the interaction dataset in bark and inserted an IDM agent, that is abel to follow the given traffic.
| docs/tutorials/04_interaction_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="VofbcErR9HwM"
# #Recurrent Neural Networks
# + colab={} colab_type="code" id="0R_nrGHiub5K"
tensorflow_version 2.x
# + [markdown] colab_type="text" id="0tNFJJspfUXS"
# ##Introduction to Recurrent Neural Networks
# ###Simple RNNs
#
# + colab={"base_uri": "https://localhost:8080/", "height": 185} colab_type="code" id="l71m9jmKM4BQ" outputId="499d2b75-301c-4f75-e543-81d2d8fbe1f7"
import numpy as np
import tensorflow as tf
from tensorflow.keras import Sequential
from tensorflow.keras.layers import SimpleRNN, Dense, Embedding
model = Sequential()
model.add(SimpleRNN(10, input_shape=(5, 2)))
model.summary()
# + [markdown] colab_type="text" id="r7yNZL-Ufrag"
# ###Embedding Layers
# + colab={"base_uri": "https://localhost:8080/", "height": 185} colab_type="code" id="XpVLJY99xxgG" outputId="dcedeea3-2422-4189-cef0-e2df44006ae7"
import numpy as np
import tensorflow as tf
from tensorflow.keras import Sequential
from tensorflow.keras.layers import SimpleRNN, Dense, Embedding
vocab_size = 30
embddng_dim = 10
seqnc_lngth = 5
model = Sequential()
model.add(Embedding(vocab_size, embddng_dim, input_length=seqnc_lngth))
model.summary()
# + [markdown] colab_type="text" id="FPgWVZybfxn5"
# ###Word Embedding + RNN on IMDB
# + colab={"base_uri": "https://localhost:8080/", "height": 171} colab_type="code" id="qfGD9MFTEFNy" outputId="3893b5d5-246a-4f7c-9f75-010abac6c4e3"
from keras.datasets import imdb
from keras.preprocessing import sequence
inpt_dim = 128
index_from = 3
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=10000,
start_char=1,
oov_char=2,
index_from=index_from,
skip_top=20)
x_train = sequence.pad_sequences(x_train, maxlen=inpt_dim)
x_test = sequence.pad_sequences(x_test, maxlen=inpt_dim)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
print(' '.join(str(int(id)) for id in x_train[7]))
word_to_id = imdb.get_word_index()
word_to_id = {k:(v+index_from) for k,v in word_to_id.items()}
word_to_id["<PAD>"] = 0
word_to_id["<START>"] = 1
word_to_id["<UNK>"] = 2
word_to_id["<UNUSED>"] = 3
id_to_word = {value:key for key,value in word_to_id.items()}
print(' '.join(id_to_word[id] for id in x_train[7]))
# + colab={"base_uri": "https://localhost:8080/", "height": 454} colab_type="code" id="QtacPS1t8f5W" outputId="88a880e9-bed6-41c1-8ac8-bc34e18d3be7"
from tensorflow.keras.models import Model
from tensorflow.keras.layers import SimpleRNN, Embedding, BatchNormalization
from tensorflow.keras.layers import Dense, Activation, Input, Dropout
from keras.datasets import imdb
from keras.preprocessing import sequence
import numpy as np
seqnc_lngth = 128
embddng_dim = 64
vocab_size = 10000
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=vocab_size, skip_top=20)
x_train = sequence.pad_sequences(x_train, maxlen=seqnc_lngth)
x_test = sequence.pad_sequences(x_test, maxlen=seqnc_lngth)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# now with batch norm
inpt_vec = Input(shape=(seqnc_lngth,))
l1 = Embedding(vocab_size, embddng_dim, input_length=seqnc_lngth)(inpt_vec)
l2 = Dropout(0.3)(l1)
l3 = SimpleRNN(32)(l2)
l4 = BatchNormalization()(l3)
l5 = Dropout(0.2)(l4)
output = Dense(1, activation='sigmoid')(l5)
# model that takes input and encodes it into the latent space
rnn = Model(inpt_vec, output)
rnn.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
rnn.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 827} colab_type="code" id="J4XhUgZvGyFL" outputId="cda2f135-d6bb-4553-92c5-5b60a60964f0"
# Fitting the RNN to the data
from tensorflow.keras.callbacks import ReduceLROnPlateau, EarlyStopping
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=3,
min_delta=1e-4, mode='min', verbose=1)
stop_alg = EarlyStopping(monitor='val_loss', patience=7,
restore_best_weights=True, verbose=1)
hist = rnn.fit(x_train, y_train, batch_size=100, epochs=1000,
callbacks=[stop_alg, reduce_lr], shuffle=True,
validation_data=(x_test, y_test))
rnn.save_weights("rnn.hdf5")
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10,6))
plt.plot(hist.history['loss'], color='#785ef0')
plt.plot(hist.history['val_loss'], color='#dc267f')
plt.title('Model Loss Progress')
plt.ylabel('Brinary Cross-Entropy Loss')
plt.xlabel('Epoch')
plt.legend(['Training Set', 'Test Set'], loc='upper right')
plt.savefig('ch.13.rnn.imdb.loss.png', dpi=350, bbox_inches='tight')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 306} colab_type="code" id="8-hnblyJCPcR" outputId="e2ec99a4-52c4-4971-aece-8793fc4864c4"
# Generate predictions
predictions = rnn.predict(x_train[0:8])
print(predictions)
for i in range(2):
INDEX_FROM=3 # word index offset
word_to_id = imdb.get_word_index()
word_to_id = {k:(v+INDEX_FROM) for k,v in word_to_id.items()}
word_to_id["<PAD>"] = 0
word_to_id["<START>"] = 1
word_to_id["<UNK>"] = 2
word_to_id["<UNUSED>"] = 3
id_to_word = {value:key for key,value in word_to_id.items()}
print('=================================================')
print(f'Sample = {i} | Length = {len(x_test[i])}')
print('=================================================')
print(' '.join(id_to_word[id] for id in x_test[i] ))
# + colab={"base_uri": "https://localhost:8080/", "height": 471} colab_type="code" id="ddjdsnhCHNOz" outputId="041ce961-5db1-4c57-98a9-03fc87904769"
from sklearn.metrics import confusion_matrix
from sklearn.metrics import balanced_accuracy_score
from sklearn.metrics import roc_curve, auc
import matplotlib.pyplot as plt
import numpy as np
y_hat = rnn.predict(x_test)
# Compute ROC curve and ROC area for each class
fpr, tpr, thresholds = roc_curve(y_test, y_hat)
roc_auc = auc(fpr, tpr)
fig = plt.figure(figsize=(10,6))
plt.plot(fpr, tpr, color='#785ef0', label='ROC curve (AUC = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='#dc267f', linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver Operating Characteristic Curve')
plt.legend(loc="lower right")
plt.savefig('ch.13.rnn.imdb.roc.png', dpi=350, bbox_inches='tight')
plt.show()
optimal_idx = np.argmax(tpr - fpr)
optimal_threshold = thresholds[optimal_idx]
print("Threshold value is:", optimal_threshold)
y_pred = np.where(y_hat>=optimal_threshold, 1, 0)
print(balanced_accuracy_score(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# + [markdown] colab_type="text" id="AwE4UOJqlR-3"
# ##Long Short-Term Memory Models
# + colab={"base_uri": "https://localhost:8080/", "height": 454} colab_type="code" id="4Hc3tLUsCjgR" outputId="e7cf3f1f-6fba-44bb-ab45-3c7d7c8e71a2"
from tensorflow.keras.models import Model
from tensorflow.keras.layers import LSTM, Embedding, BatchNormalization
from tensorflow.keras.layers import Dense, Activation, Input, Dropout
from keras.datasets import imdb
from keras.preprocessing import sequence
import numpy as np
seqnc_lngth = 128
embddng_dim = 64
vocab_size = 10000
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=vocab_size, skip_top=20)
x_train = sequence.pad_sequences(x_train, maxlen=seqnc_lngth)
x_test = sequence.pad_sequences(x_test, maxlen=seqnc_lngth)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# now with batch norm
inpt_vec = Input(shape=(seqnc_lngth,))
l1 = Embedding(vocab_size, embddng_dim, input_length=seqnc_lngth)(inpt_vec)
l2 = Dropout(0.3)(l1)
l3 = LSTM(32)(l2)
l4 = BatchNormalization()(l3)
l5 = Dropout(0.2)(l4)
output = Dense(1, activation='sigmoid')(l5)
# model that takes input and encodes it into the latent space
lstm = Model(inpt_vec, output)
lstm.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
lstm.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 827} colab_type="code" id="iY4_JvYVHPaK" outputId="df78449d-355c-48bc-86e1-804a165b5091"
# Fitting the LSTM to the data
from tensorflow.keras.callbacks import ReduceLROnPlateau, EarlyStopping
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=3,
min_delta=1e-4, mode='min', verbose=1)
stop_alg = EarlyStopping(monitor='val_loss', patience=7,
restore_best_weights=True, verbose=1)
hist = lstm.fit(x_train, y_train, batch_size=100, epochs=1000,
callbacks=[stop_alg, reduce_lr], shuffle=True,
validation_data=(x_test, y_test))
lstm.save_weights("lstm.hdf5")
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10,6))
plt.plot(hist.history['loss'], color='#785ef0')
plt.plot(hist.history['val_loss'], color='#dc267f')
plt.title('Model Loss Progress')
plt.ylabel('Brinary Cross-Entropy Loss')
plt.xlabel('Epoch')
plt.legend(['Training Set', 'Test Set'], loc='upper right')
plt.savefig('ch.13.lstm.imdb.loss.png', dpi=350, bbox_inches='tight')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 306} colab_type="code" id="GA9e4k_iGTQj" outputId="ccff0781-9599-462f-e7cd-67f29b5d3dfb"
# Generate predictions
predictions = lstm.predict(x_train[0:8])
print(predictions)
for i in range(2):
INDEX_FROM=3 # word index offset
word_to_id = imdb.get_word_index()
word_to_id = {k:(v+INDEX_FROM) for k,v in word_to_id.items()}
word_to_id["<PAD>"] = 0
word_to_id["<START>"] = 1
word_to_id["<UNK>"] = 2
word_to_id["<UNUSED>"] = 3
id_to_word = {value:key for key,value in word_to_id.items()}
print('=================================================')
print(f'Sample = {i} | Length = {len(x_test[i])}')
print('=================================================')
print(' '.join(id_to_word[id] for id in x_test[i] ))
# + colab={"base_uri": "https://localhost:8080/", "height": 471} colab_type="code" id="UPNQCByhN4qK" outputId="a9437cb4-dcac-4698-bb10-67ff95d8a512"
from sklearn.metrics import confusion_matrix
from sklearn.metrics import balanced_accuracy_score
from sklearn.metrics import roc_curve, auc
from sklearn.metrics import roc_auc_score
import matplotlib.pyplot as plt
import numpy as np
y_hat = lstm.predict(x_test)
# Compute ROC curve and ROC area for each class
fpr, tpr, thresholds = roc_curve(y_test, y_hat)
roc_auc = auc(fpr, tpr)
fig = plt.figure(figsize=(10,6))
plt.plot(fpr, tpr, color='#785ef0', label='ROC curve (AUC = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='#dc267f', linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver Operating Characteristic Curve')
plt.legend(loc="lower right")
plt.savefig('ch.13.lstm.imdb.roc.png', dpi=350, bbox_inches='tight')
plt.show()
optimal_idx = np.argmax(tpr - fpr)
optimal_threshold = thresholds[optimal_idx]
print("Threshold value is:", optimal_threshold)
y_pred = np.where(y_hat>=optimal_threshold, 1, 0)
print(balanced_accuracy_score(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# + [markdown] colab_type="text" id="3ArC1yvzgB4c"
# ##Sequence to Vector Models
# ###Unsupervised Model
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="71n1rZprrLaJ" outputId="bcd45281-d9b5-4c01-f293-c0aea7956bf3"
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Dense, Activation, Input
from tensorflow.keras.layers import BatchNormalization, Dropout
from tensorflow.keras.layers import Embedding, LSTM
from tensorflow.keras.layers import RepeatVector, TimeDistributed
from tensorflow.keras.datasets import mnist
import numpy as np
from tensorflow.keras.callbacks import ReduceLROnPlateau, EarlyStopping
seqnc_lngth = 28
ltnt_dim = 2
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
inpt_vec = Input(shape=(seqnc_lngth, seqnc_lngth,))
l1 = Dropout(0.1)(inpt_vec)
l2 = LSTM(seqnc_lngth, activation='tanh',
recurrent_activation='sigmoid')(l1)
l3 = BatchNormalization()(l2)
l4 = Dropout(0.1)(l3)
l5 = Dense(ltnt_dim, activation='sigmoid')(l4)
# model that takes input and encodes it into the latent space
encoder = Model(inpt_vec, l5)
l6 = RepeatVector(seqnc_lngth)(l5)
l7 = LSTM(seqnc_lngth, activation='tanh', recurrent_activation='sigmoid',
return_sequences=True)(l6)
l8 = BatchNormalization()(l7)
l9 = TimeDistributed(Dense(seqnc_lngth, activation='sigmoid'))(l8)
autoencoder = Model(inpt_vec, l9)
autoencoder.compile(loss='binary_crossentropy', optimizer='adam')
autoencoder.summary()
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=5,
min_delta=1e-4, mode='min', verbose=1)
stop_alg = EarlyStopping(monitor='val_loss', patience=15,
restore_best_weights=True, verbose=1)
hist = autoencoder.fit(x_train, x_train, batch_size=100, epochs=1000,
callbacks=[stop_alg, reduce_lr], shuffle=True,
validation_data=(x_test, x_test))
# + [markdown] colab_type="text" id="kXGM075ZmTYM"
# ###Results
# + colab={"base_uri": "https://localhost:8080/", "height": 404} colab_type="code" id="18GxIyuPrMzZ" outputId="0406e339-2cc5-40bc-ee07-e1bad5e41464"
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10,6))
plt.plot(hist.history['loss'], color='#785ef0')
plt.plot(hist.history['val_loss'], color='#dc267f')
plt.title('Model Loss Progress')
plt.ylabel('Brinary Cross-Entropy Loss')
plt.xlabel('Epoch')
plt.legend(['Training Set', 'Test Set'], loc='upper right')
plt.savefig('ch.13.ae.lstm.mnist.loss.png', dpi=350, bbox_inches='tight')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 671} colab_type="code" id="UCRQbImWWq-u" outputId="f7784495-eb94-42c3-8946-00025b28133c"
import matplotlib.pyplot as plt
import numpy as np
encdd = encoder.predict(x_test)
x_hat = autoencoder.predict(x_test)
smp_idx = [3,2,1,18,4,8,11,0,61,9]
plt.figure(figsize=(12,6))
for i, (img, y) in enumerate(zip(x_hat[smp_idx].reshape(10, 28, 28), y_test[smp_idx])):
plt.subplot(2,5,i+1)
plt.imshow(img, cmap='gray')
plt.xticks([])
plt.yticks([])
plt.title(y)
plt.savefig('ch.13.ae.lstm.mnist.reconstructed.png', bbox_inches='tight', dpi=350)
plt.show()
plt.figure(figsize=(12,6))
for i, (img, y) in enumerate(zip(x_test[smp_idx].reshape(10, 28, 28), y_test[smp_idx])):
plt.subplot(2,5,i+1)
plt.imshow(img, cmap='gray')
plt.xticks([])
plt.yticks([])
plt.title(y)
plt.savefig('ch.13.ae.lstm.mnist.original.png', bbox_inches='tight', dpi=350)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 530} colab_type="code" id="d2V_aR3L6KYg" outputId="6736e846-d015-446f-805a-72e580655f88"
import matplotlib.pyplot as plt
y_ = list(map(int, y_test))
X_ = encdd
print(X_.shape)
plt.figure(figsize=(10,8))
plt.title('LSTM-based Encoder')
plt.scatter(X_[:,0], X_[:,1], s=5.0, c=y_, alpha=0.75, cmap='tab10')
plt.xlabel('First encoder dimension')
plt.ylabel('Second encoder dimension')
plt.colorbar()
plt.savefig('ch.13.ae.lstm.png', bbox_inches='tight', dpi=350)
# + [markdown] colab_type="text" id="Xa1ZWW4Xmd7l"
# ##Vector to Sequence Models
# ###Bidirectional LSTM
# ###Implementation and Results
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="hu37iJMmSc6d" outputId="bf5bec96-c202-4623-f459-5f0d00ba9915"
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Dense, Activation, Input
from tensorflow.keras.layers import BatchNormalization, Dropout
from tensorflow.keras.layers import Bidirectional, LSTM
from tensorflow.keras.layers import RepeatVector, TimeDistributed
from tensorflow.keras.datasets import mnist
import numpy as np
from tensorflow.keras.callbacks import ReduceLROnPlateau, EarlyStopping
seqnc_lngth = 28
ltnt_dim = 100
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
inpt_vec = Input(shape=(seqnc_lngth, seqnc_lngth,))
l1 = Dropout(0.5)(inpt_vec)
l2 = Bidirectional(LSTM(seqnc_lngth, activation='tanh',
recurrent_activation='sigmoid'))(l1)
l3 = BatchNormalization()(l2)
l4 = Dropout(0.5)(l3)
l5 = Dense(ltnt_dim, activation='sigmoid')(l4)
# model that takes input and encodes it into the latent space
encoder = Model(inpt_vec, l5, name='encoder')
encoder.summary()
ltnt_vec = Input(shape=(ltnt_dim,))
l6 = Dropout(0.1)(ltnt_vec)
l7 = RepeatVector(seqnc_lngth)(l6)
l8 = Bidirectional(LSTM(seqnc_lngth, activation='tanh', recurrent_activation='sigmoid',
return_sequences=True))(l7)
l9 = BatchNormalization()(l8)
l10 = TimeDistributed(Dense(seqnc_lngth, activation='sigmoid'))(l9)
decoder = Model(ltnt_vec, l10, name='decoder')
decoder.summary()
recon = decoder(encoder(inpt_vec))
autoencoder = Model(inpt_vec, recon, name='ae')
autoencoder.compile(loss='binary_crossentropy', optimizer='adam')
autoencoder.summary()
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=5,
min_delta=1e-4, mode='min', verbose=1)
stop_alg = EarlyStopping(monitor='val_loss', patience=15,
restore_best_weights=True, verbose=1)
hist = autoencoder.fit(x_train, x_train, batch_size=100, epochs=1000,
callbacks=[stop_alg, reduce_lr], shuffle=True,
validation_data=(x_test, x_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 404} colab_type="code" id="3WZzPr6iW5fW" outputId="5aabc48e-d86d-4c61-fcd3-cf643e45f5a9"
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10,6))
plt.plot(hist.history['loss'], color='#785ef0')
plt.plot(hist.history['val_loss'], color='#dc267f')
plt.title('Model Loss Progress')
plt.ylabel('Brinary Cross-Entropy Loss')
plt.xlabel('Epoch')
plt.legend(['Training Set', 'Test Set'], loc='upper right')
plt.savefig('ch.13.ae.bilstm.mnist.loss.png', dpi=350, bbox_inches='tight')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 671} colab_type="code" id="2mTJaRG_XATq" outputId="87f6d3ad-9edd-49ed-d437-42cae64973a1"
import matplotlib.pyplot as plt
import numpy as np
encdd = encoder.predict(x_test)
x_hat = autoencoder.predict(x_test)
smp_idx = [3,2,1,18,4,8,11,0,61,9]
plt.figure(figsize=(12,6))
for i, (img, y) in enumerate(zip(x_hat[smp_idx].reshape(10, 28, 28), y_test[smp_idx])):
plt.subplot(2,5,i+1)
plt.imshow(img, cmap='gray')
plt.xticks([])
plt.yticks([])
plt.title(y)
plt.savefig('ch.13.ae.bilstm.mnist.reconstructed.png', bbox_inches='tight', dpi=350)
plt.show()
plt.figure(figsize=(12,6))
for i, (img, y) in enumerate(zip(x_test[smp_idx].reshape(10, 28, 28), y_test[smp_idx])):
plt.subplot(2,5,i+1)
plt.imshow(img, cmap='gray')
plt.xticks([])
plt.yticks([])
plt.title(y)
plt.savefig('ch.13.ae.bilstm.mnist.original.png', bbox_inches='tight', dpi=350)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 546} colab_type="code" id="LVyhptpAXL3N" outputId="7eb853de-19ec-4d28-c812-a479a350ad27"
import matplotlib.pyplot as plt
import umap
y_ = list(map(int, y_test))
X_ = encdd
print(X_.shape)
X_ = umap.UMAP().fit_transform(encdd)
print(X_.shape)
plt.figure(figsize=(10,8))
plt.title('UMAP of a BiLSTM Encoder')
plt.scatter(X_[:,0], X_[:,1], s=5.0, c=y_, alpha=0.75, cmap='tab10')
plt.xlabel('First UMAP dimension')
plt.ylabel('Second UMAP dimension')
plt.colorbar()
plt.savefig('ch.13.ae.bilstm.umap.png', bbox_inches='tight', dpi=350)
# + colab={"base_uri": "https://localhost:8080/", "height": 298} colab_type="code" id="5bYo86L8YWtH" outputId="9810b579-f846-4cda-eb6e-e48ef2379583"
z = np.random.rand(1,100)
x_ = decoder.predict(z)
print(x_.shape)
plt.imshow(x_[0], cmap='gray')
# + [markdown] colab_type="text" id="6NroIdYDnIXr"
# ##Sequence to Sequence Models
# + colab={"base_uri": "https://localhost:8080/", "height": 344} colab_type="code" id="Ku8rlvWg8pXy" outputId="7e46b192-1090-4737-b89b-9f78d059acf8"
plt.figure(figsize=(12,6))
for i in range(10):
plt.subplot(2,5,i+1)
rnd_vec = np.round(np.mean(x_test[y_test==i],axis=0))
rnd_vec = np.reshape(rnd_vec, (1,28,28))
z = encoder.predict(rnd_vec)
decdd = decoder.predict(z)
plt.imshow(decdd[0], cmap='gray')
plt.xticks([])
plt.yticks([])
plt.title(i)
plt.savefig('ch.13.ae.bilstm.mnist.v2s.png', bbox_inches='tight', dpi=350)
plt.show()
| Chapter13/Chapter_13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 라이브러리 불러오기
import sklearn
print(sklearn.__version__)
# !pip install sklearn
import sklearn
print(sklearn.__version__)
import numpy as np
import pandas as pd
# +
# -----------------------------------
# 학습 데이터, 테스트 데이터 읽기
# -----------------------------------
# 학습 데이터, 테스트 데이터 읽기
train = pd.read_csv('../input/ch01-titanic/train.csv')
test = pd.read_csv('../input/ch01-titanic/test.csv')
# 학습 데이터을 종속 변수와 목적 변수로 나누기
train_x = train.drop(['Survived'], axis=1)
train_y = train['Survived']
# 테스트 데이터와 종속변수만 있기 때문에, 그대로 괜찮음.
test_x = test.copy()
# +
# -----------------------------------
# 피처 추출(피처 엔지니어링)
# -----------------------------------
from sklearn.preprocessing import LabelEncoder
# 변수 PassengerId을 제거
train_x = train_x.drop(['PassengerId'], axis=1)
test_x = test_x.drop(['PassengerId'], axis=1)
# 변수Name, Ticket, Cabin을 제거
train_x = train_x.drop(['Name', 'Ticket', 'Cabin'], axis=1)
test_x = test_x.drop(['Name', 'Ticket', 'Cabin'], axis=1)
# 범주형 변수를 label encoding을 적용.
for c in ['Sex', 'Embarked']:
# 학습 데이터를 기반으로 어떻게 변환할지를 정한다.
le = LabelEncoder()
le.fit(train_x[c].fillna('NA'))
# 학습 데이터, 테스트 데이터를 변환
train_x[c] = le.transform(train_x[c].fillna('NA'))
test_x[c] = le.transform(test_x[c].fillna('NA'))
# +
# -----------------------------------
# 모델 만들기
# -----------------------------------
from xgboost import XGBClassifier
# 모델 만들기 및 학습 데이터를 가지고 학습
model = XGBClassifier(n_estimators=20, random_state=71)
model.fit(train_x, train_y)
# 테스트 데이터의 예측치를 확률로 출력한다.
pred = model.predict_proba(test_x)[:, 1]
# 테스트 데이터의 예측치를 두개의 값(1,0)으로 변환
pred_label = np.where(pred > 0.5, 1, 0)
# 제출용 파일의 작성
submission = pd.DataFrame({'PassengerId': test['PassengerId'], 'Survived': pred_label})
submission.to_csv('submission_first.csv', index=False)
# +
# score:0.7799(이 수치는 사용자에 따라다를 수 있음.)
# +
# -----------------------------------
# 모델 평가
# -----------------------------------
from sklearn.metrics import log_loss, accuracy_score
from sklearn.model_selection import KFold
# 각 fold의 스코어를 저장을 위한 빈 리스트 작성
scores_accuracy = []
scores_logloss = []
# 교차 검증(Cross-validation)을 수행
# 01 학습 데이터를 4개로 분할
# 02 그중의 하나를 평가용 데이터 셋으로 한다.
# 03 이후 평가용 데이터 셋을 하나씩 옆으로 옮겨가며 검증을 수행
kf = KFold(n_splits=4, shuffle=True, random_state=71)
for tr_idx, va_idx in kf.split(train_x):
# 학습 데이터를 학습 데이터와 평가용 데이터 셋으로 나눈다.
tr_x, va_x = train_x.iloc[tr_idx], train_x.iloc[va_idx]
tr_y, va_y = train_y.iloc[tr_idx], train_y.iloc[va_idx]
# 모델 학습을 수행
model = XGBClassifier(n_estimators=20, random_state=71)
model.fit(tr_x, tr_y)
# 평가용 데이터의 예측 결과를 확률로 출력
va_pred = model.predict_proba(va_x)[:, 1]
# 평가용 데이터의 스코어를 계산
logloss = log_loss(va_y, va_pred)
accuracy = accuracy_score(va_y, va_pred > 0.5)
# 각 fold의 스코어를 저장
scores_logloss.append(logloss)
scores_accuracy.append(accuracy)
# 각 fold의 스코어의 평균을 출력
logloss = np.mean(scores_logloss)
accuracy = np.mean(scores_accuracy)
print(f'logloss: {logloss:.4f}, accuracy: {accuracy:.4f}')
# +
# logloss: 0.4270, accuracy: 0.8148(결과값은 이 책과 다를 가능성이 있습니다.)
# +
# -----------------------------------
# 모델 튜닝
# -----------------------------------
import itertools
# 튜닝 후보로 삼는 파라미터 값을 준비
param_space = {
'max_depth': [3, 5, 7],
'min_child_weight': [1.0, 2.0, 4.0]
}
# 탐색할 하이퍼 파라미터 값의 조합
param_combinations = itertools.product(param_space['max_depth'], param_space['min_child_weight'])
# 각 파라미터의 조합, 그에 대한 스코어를 보존하는 빈 리스트
params = []
scores = []
# 각 파라미터 조합별로 교차 검증(Cross-validation)으로 평가를 수행
for max_depth, min_child_weight in param_combinations:
score_folds = []
# 교차 검증(Cross-validation)을 수행
# 학습 데이터를 4개로 분할한 후,
# 그중 하나를 평가용 데이터로 하는 것을 데이터를 바꾸어 가면서 반복한다.
kf = KFold(n_splits=4, shuffle=True, random_state=123456)
for tr_idx, va_idx in kf.split(train_x):
# 학습 데이터를 학습 데이터와 평가용 데이터로 분할
tr_x, va_x = train_x.iloc[tr_idx], train_x.iloc[va_idx]
tr_y, va_y = train_y.iloc[tr_idx], train_y.iloc[va_idx]
# 모델의 학습을 수행
model = XGBClassifier(n_estimators=20, random_state=71,
max_depth=max_depth, min_child_weight=min_child_weight)
model.fit(tr_x, tr_y)
# 평가용 데이터의 스코어를 계산한 후, 저장
va_pred = model.predict_proba(va_x)[:, 1]
logloss = log_loss(va_y, va_pred)
score_folds.append(logloss)
# 각 fold의 스코어의 평균을 구한다.
score_mean = np.mean(score_folds)
# 파라미터를 조합, 그에 대한 스코어를 저장.
params.append((max_depth, min_child_weight))
scores.append(score_mean)
# 가장 스코어가 좋은 것을 베스트 파라미터로 한다.
# 最もスコアが良いものをベストなパラメータとする
best_idx = np.argsort(scores)[0]
best_param = params[best_idx]
print(f'max_depth: {best_param[0]}, min_child_weight: {best_param[1]}')
# max_depth=7, min_child_weight=2.0의 스코어가 가장 좋았음.
# -
# ### 1.5.6 책에는 빠져 있음
# +
# -----------------------------------
# 로지스틱 회귀용 feature 만들기
# -----------------------------------
from sklearn.preprocessing import OneHotEncoder
# 원 데이터를 복사한다.
train_x2 = train.drop(['Survived'], axis=1)
test_x2 = test.copy()
# 변수 PassengerId를 제거한다.
train_x2 = train_x2.drop(['PassengerId'], axis=1)
test_x2 = test_x2.drop(['PassengerId'], axis=1)
# 변수Name, Ticket, Cabin을 제거
train_x2 = train_x2.drop(['Name', 'Ticket', 'Cabin'], axis=1)
test_x2 = test_x2.drop(['Name', 'Ticket', 'Cabin'], axis=1)
# -
train_x2.info()
# 범주형 변수를 label encoding을 적용.
for c in ['Sex', 'Embarked']:
# 학습 데이터를 기반으로 어떻게 변환할지를 정한다.
le = LabelEncoder()
le.fit(train_x2[c].fillna('NA'))
# 학습 데이터, 테스트 데이터를 변환
train_x2[c] = le.transform(train_x2[c].fillna('NA'))
test_x2[c] = le.transform(test_x2[c].fillna('NA'))
train_x2.head()
# one-hot encoding을 수행
cat_cols = ['Embarked', 'Pclass', 'Sex']
# ohe = OneHotEncoder(categories='auto', sparse=False)
ohe = OneHotEncoder(sparse=False)
ohe.fit(train_x2[cat_cols].fillna('NA'))
train_x2.Sex.unique()
# one-hot encoding의 더미 변수 컬럼명을 작성한다.
ohe_columns = []
for i, c in enumerate(cat_cols):
print(i,c)
# ohe_columns += [f'{c}_{v}' for v in ohe.categories_[i]]
ohe_columns += [f'{c}_{v}' for v in train_x2[c].unique()]
ohe_columns.sort()
ohe_columns
train_x2.head()
# one-hot encodingによる変換を行う
ohe_train_x2 = pd.DataFrame(ohe.transform(train_x2[cat_cols].fillna('NA')), columns=ohe_columns)
ohe_test_x2 = pd.DataFrame(ohe.transform(test_x2[cat_cols].fillna('NA')), columns=ohe_columns)
ohe_train_x2
# +
# one-hot encoding 끝난 변수를 제외하가
train_x2 = train_x2.drop(cat_cols, axis=1)
test_x2 = test_x2.drop(cat_cols, axis=1)
# one-hot encoding로 변환된 변수를 추가
train_x2 = pd.concat([train_x2, ohe_train_x2], axis=1)
test_x2 = pd.concat([test_x2, ohe_test_x2], axis=1)
# 수치 변수의 결측치를 학습 데이터 평균으로 메우기
num_cols = ['Age', 'SibSp', 'Parch', 'Fare']
for col in num_cols:
train_x2[col].fillna(train_x2[col].mean(), inplace=True)
test_x2[col].fillna(train_x2[col].mean(), inplace=True)
# 변수 Fare을 로그 변환하기
train_x2['Fare'] = np.log1p(train_x2['Fare'])
test_x2['Fare'] = np.log1p(test_x2['Fare'])
# -
# ### 1.5.6 앙상블
# +
# -----------------------------------
# 앙상블(ensemble)
# -----------------------------------
from sklearn.linear_model import LogisticRegression
# xgboost 모델
model_xgb = XGBClassifier(n_estimators=20, random_state=71)
model_xgb.fit(train_x, train_y)
pred_xgb = model_xgb.predict_proba(test_x)[:, 1]
# 로지스틱 회귀 모델
# xgboost 모델과는 다른 종속변수를 넣어야 하는 필요가 있으므로 별도로 train_x2, test_x2를 작성.
model_lr = LogisticRegression(solver='lbfgs', max_iter=300)
model_lr.fit(train_x2, train_y)
pred_lr = model_lr.predict_proba(test_x2)[:, 1]
# 예측 결과의 가중 평균을 취하다.
pred = pred_xgb * 0.8 + pred_lr * 0.2
pred_label = np.where(pred > 0.5, 1, 0)
# -
pred_label
| ch01/.ipynb_checkpoints/ch01_01_titanic_sk_ver0_22-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from tensorflow.keras.datasets import cifar10
import numpy
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D, Input, BatchNormalization
import os
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Conv2D, MaxPool2D , Flatten
from tensorflow.keras import optimizers
import numpy as np
(trainX, trainy), (testX, testy) = cifar10.load_data()
# print to make sure we have the correct shapes + number of images for training
print("number of train pictures:", trainX.shape)
print("number of trained picture values:", trainy.shape)
# divide by 255 to make [0,255] into [0,1] + print to make sure!
trainy = tf.keras.utils.to_categorical(trainy,10)
testy = tf.keras.utils.to_categorical(testy,10)
trainX = trainX/255.0
testX = testX/255.0
# +
import tensorflow_model_optimization as tfmot
LastValueQuantizer = tfmot.quantization.keras.quantizers.LastValueQuantizer
MovingAverageQuantizer = tfmot.quantization.keras.quantizers.MovingAverageQuantizer
class ModifiedDenseQuantizeConfig(tfmot.quantization.keras.QuantizeConfig):
def get_weights_and_quantizers(self, layer):
return [(layer.kernel, LastValueQuantizer(num_bits=4, symmetric=True, narrow_range=False, per_axis=False))]
def get_activations_and_quantizers(self, layer):
return [(layer.activation, MovingAverageQuantizer(num_bits=4, symmetric=False, narrow_range=False, per_axis=False))]
def set_quantize_weights(self, layer, quantize_weights):
# Add this line for each item returned in `get_weights_and_quantizers`
# , in the same order
layer.kernel = quantize_weights[0]
def set_quantize_activations(self, layer, quantize_activations):
# Add this line for each item returned in `get_activations_and_quantizers`
# , in the same order.
layer.activation = quantize_activations[0]
# Configure how to quantize outputs (may be equivalent to activations).
def get_output_quantizers(self, layer):
return []
def get_config(self):
return {}
# -
# ### Quantizing vgg-16
# +
annotate = tfmot.quantization.keras.quantize_annotate_layer
quant_vgg16 = tf.keras.Sequential()
# Only annotated layers will be quantized
#block-1
quant_vgg16.add(annotate(Conv2D(input_shape=(32,32,3),
filters=64,kernel_size=(3,3),
padding="same",
activation="relu",
name='block1_conv1'), quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.3))
quant_vgg16.add(annotate(Conv2D(filters=64,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block1_conv2'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.4))
quant_vgg16.add(MaxPool2D(pool_size=(2,2), strides=(2,2), name='block1_pool'))
#block-2
quant_vgg16.add(annotate(Conv2D(filters=128,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block2_conv1'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.4))
quant_vgg16.add(annotate(Conv2D(filters=128,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block2_conv2'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(MaxPool2D(pool_size=(2,2),strides=(2,2), name='block2_pool'))
#block-3
quant_vgg16.add(annotate(Conv2D(filters=256,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block3_conv1'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.4))
quant_vgg16.add(annotate(Conv2D(filters=256,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block3_conv2'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.4))
quant_vgg16.add(annotate(Conv2D(filters=256,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block3_conv3'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(MaxPool2D(pool_size=(2,2),strides=(2,2), name='block3_pool'))
#block-4
quant_vgg16.add(annotate(Conv2D(filters=512,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block4_conv1'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.4))
quant_vgg16.add(annotate(Conv2D(filters=512,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block4_conv2'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.4))
quant_vgg16.add(annotate(Conv2D(filters=512,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block4_conv3'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(MaxPool2D(pool_size=(2,2),strides=(2,2), name='block4_pool'))
#block-5
quant_vgg16.add(annotate(Conv2D(filters=512,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block5_conv1'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(Dropout(0.4))
quant_vgg16.add(annotate(Conv2D(filters=512,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block5_conv2'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.4))
quant_vgg16.add(annotate(Conv2D(filters=512,
kernel_size=(3,3),
padding="same",
activation="relu",
name='block5_conv3'),
quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(MaxPool2D(pool_size=(2,2),strides=(2,2), name='block5_pool'))
#fc1, fc2 and predictions
quant_vgg16.add(Dropout(0.5))
quant_vgg16.add(annotate(Flatten(name='flatten')))
quant_vgg16.add(annotate(Dense(units=512,activation="relu",name='fc1'), quantize_config=ModifiedDenseQuantizeConfig()))
quant_vgg16.add(BatchNormalization())
quant_vgg16.add(Dropout(0.5))
quant_vgg16.add(Dense(units=10, activation="softmax",name='predictions'))
quantize_scope = tfmot.quantization.keras.quantize_scope
# `quantize_apply` requires mentioning `DefaultDenseQuantizeConfig` with `quantize_scope`
# as well as the custom Keras layer.
with quantize_scope(
{'ModifiedDenseQuantizeConfig':ModifiedDenseQuantizeConfig}):
# Use `quantize_apply` to actually make the model quantization aware.
vgg_quant_model = tfmot.quantization.keras.quantize_apply(quant_vgg16)
vgg_quant_model.summary()
# +
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.optimizers import SGD
opt = SGD(learning_rate=0.1, decay=1e-6, momentum=0.9, nesterov=True)
# Compile the model
vgg_quant_model.compile(optimizer=opt, loss=tf.keras.losses.categorical_crossentropy,metrics=['accuracy'])
# Fit data to model
vgg_quant_model.fit(trainX, trainy,
batch_size=50,
epochs=100,
verbose=1,
validation_split=0.2)
# -
vgg_quant_model.save_weights('cifar10vgg_quant4.h5')
score = vgg_quant_model.evaluate(testX, testy, verbose=1)
print("Test loss {:.4f}, accuracy {:.2f}%".format(score[0], score[1] * 100))
52
| partitioned_vgg16/vgg_INT4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''project-tinder-env'': conda)'
# name: python3
# ---
# # Agent-Based Simulation: Balanced Flow Convergence
# +
import matplotlib.pyplot as plt
from pathlib import Path
import random
import pandas as pd
from matplotlib.collections import LineCollection
import matplotlib as mpl
import numpy as np
from scipy import optimize as opt
from scipy import integrate as intg
from scipy.optimize import least_squares
from scipy.stats import beta
from scipy.stats import bernoulli
from scipy.stats import cumfreq, beta
from utils import simulate_game
# %matplotlib inline
plt.style.use('../notebook.mplstyle')
SEED = 1
# -
# ## Exogenous Parameters
#
# +
# Setting exogenous parameters
def reset_exog_params():
global Bm, Bw, bm_vals, bw_vals, δ, Um, Uw, Fm, Fw, λm, λw
Bm = 10
Bw = 10
bm_vals = range(1,Bm+1)
bw_vals = range(1,Bw+1)
δ = 0.97
Um = lambda θ : θ
Uw = lambda θ : θ
Fm = beta(3,3)
Fw = beta(3,3)
λm = 50
λw = 50
def exog_params():
return (Bm, Bw, bm_vals, bw_vals, δ, Um, Uw, Fm, Fw, λm, λw)
# -
# ## Steady State with Cutoff Strategies
def steady_state_men(μ):
# Computing z's
zm = []
for b in bm_vals:
if b==Bm:
zm.append(1-δ*Fw.cdf(μ[Bm-1]))
else:
z=1
for i in range(1, Bm-b+1):
z *= ((δ*(1-Fw.cdf(μ[Bm-i])))/(1-δ*Fw.cdf(μ[Bm-i-1])))
zm.append(z)
# Computing steady-state mass
Nm = (λm) * ((zm[Bm-1] - δ * zm[0] * (1 - Fw.cdf(μ[0]))) / ((1-δ) * zm[Bm-1]))
# Computing steady state distribution over budgets
Pbm = [((λm) / (Nm * zm[Bm-1])) * zm[b] for b in range(Bm-1)]
Pbm.append(((λm) / (Nm * zm[Bm-1])))
return Nm, Pbm
def steady_state_women(ω):
# Computing z's
zw = []
for b in bw_vals:
if b==Bw:
zw.append(1-δ*Fm.cdf(ω[Bw-1]))
else:
z=1
for i in range(1, Bw-b+1):
z *= ((δ * (1-Fm.cdf(ω[Bw-i])))/(1-δ*Fm.cdf(ω[Bw-i-1])))
zw.append(z)
# Computing steady-state mass
Nw = (λw) * ((zw[Bw-1] - δ * zw[0] * (1 - Fm.cdf(ω[0]))) / ((1-δ) * zw[Bw-1]))
# Computing steady state distribution over budgets
Pbw = [((λw) / (Nw * zw[Bw-1])) * zw[b] for b in range(Bw-1)]
Pbw.append(((λw) / (Nw * zw[Bw-1])))
return Nw, Pbw
def steady_state(μ, ω, verbose=False):
# Computing masses and distributions
Nm, Pbm = steady_state_men(μ)
Nw, Pbw = steady_state_women(ω)
# Computing tightness and alpha
if Nw>Nm:
τm = 1
else:
τm = Nw/Nm
τw = τm *(Nm/Nw)
αm = (τm*δ)/(1-δ*(1-τm))
αw = (τw*δ)/(1-δ*(1-τw))
return Nm, Nw, Pbm, Pbw, τm, τw, αm, αw
# ## Two-Sided Search Equilibrium Conditions
#
# Optimality conditions
def SSE(x):
# Compute steady state
Nm, Nw, Pbm, Pbw, τm, τw, αm, αw = steady_state(x[:Bm], x[Bm:])
# Initialysing system of equilibrium equations
E = np.empty(2*Bm + 2*Bw + 2)
# Initial conditions
E[0] = (Um(x[0])
- αm * Um(x[0]) * Fw.cdf(x[0])
- αm * intg.quad(lambda t: Um(t) * Fw.pdf(t), x[0], 1)[0])
E[Bm] = (Uw(x[Bm])
- αw * Uw(x[Bm]) * Fm.cdf(x[Bm])
- αw * intg.quad(lambda t: Uw(t) * Fm.pdf(t), x[Bm], 1)[0])
# Intertemporal optimality conditions for men
for b in range(1,Bm):
E[b] = (Um(x[b])
- αm * Um(x[b]) * Fw.cdf(x[b])
- αm * Um(x[b-1])*(1-Fw.cdf(x[b-1]))
- αm * intg.quad(lambda t : Um(t) * Fw.pdf(t), x[b], x[b-1])[0])
# Intertemporal optimality conditions for women
for b in range(1,Bw):
E[Bm+b] = (Uw(x[Bm+b])
- αw * Uw(x[Bm+b]) * Fm.cdf(x[Bm+b])
- αw * Uw(x[Bm+b-1])*(1-Fm.cdf(x[Bm+b-1]))
- αw * intg.quad(lambda t : Uw(t) * Fm.pdf(t), x[Bm+b], x[Bm+b-1])[0])
# PMF unity sum conditions
E[Bm+Bw] = sum(Pbm)-1
E[Bm+Bw+1] = sum(Pbw)-1
# PMF non-negativity conditions
for b in range(Bm):
E[Bm+Bw+2+b] = Pbm[b]-abs(Pbm[b])
for b in range(Bw):
E[Bm+Bw+2+Bm+b] = Pbw[b]-abs(Pbw[b])
return E
# ## Solving For Steady State Equilibria
# +
reset_exog_params()
m_test = np.random.rand(Bm)#*0.5
w_test = np.random.rand(Bw)#*0.5
print('μ0: ', m_test)
print('ω0: ', w_test)
print('')
x_start = np.concatenate((m_test, w_test), axis=None)
solution = least_squares(SSE, x_start, bounds = (0,1), verbose=1)
μ_star = solution.x[:Bm]
ω_star = solution.x[Bm:]
print('')
print('μ*', μ_star)
print('ω*', ω_star)
print('Loss:', round(solution.cost,6))
# +
# Computing steady state
Nm, Nw, Pbm, Pbw, τm, τw, αm, αw = steady_state(μ_star, ω_star, True)
print('Stock: ', round(Nm), round(Nw))
print('PMF check:', round(sum(Pbm),3), round(sum(Pbw),3))
print('Tightness: ', round(τm,2), round(τw,2))
print('Alphas: ', round(αm,2), round(αw,2))
ρm = sum([(1 - Fw.cdf(μ_star[b]))*Pbm[b] for b in range(Bm)])
ρw = sum([(1 - Fm.cdf(ω_star[b]))*Pbw[b] for b in range(Bw)])
print('Average Swipe Rate: ', round(ρm, 3), round(ρw, 3))
# -
# ## Single Batch Agent-Based Simulation
reset_exog_params()
T = 100
simulation = simulate_game(T, μ_star, ω_star, exog_params())
batch = pd.DataFrame(simulation)
batch.head(5)
# +
mass_by_sex = batch.groupby(['time', 'sex'], as_index=False).size()
Nm_series = mass_by_sex[mass_by_sex.sex=='Male']['size']
Nw_series = mass_by_sex[mass_by_sex.sex=='Female']['size']
fig, ax = plt.subplots()
ax.set_xlim(0,T)
ax.plot(range(0,T), Nm_series, color='tab:blue', label='Male')
ax.plot(range(0,T), Nw_series, color='tab:pink', label='Female')
ax.axhline(y=Nm, color='tab:blue', linestyle='--', label='Male Steady State')
ax.axhline(y=Nw, color='tab:pink', linestyle='--', label='Female Steady State')
ax.set(xlabel=r'Time ($t$)', ylabel=r'Agent Stock ($N_t$)')
ax.legend()
plt.show()
# -
# ## Multiple Batch Agent-Based Simulation
# +
simulations = []
batches = 10
for i in range(batches):
game = simulate_game(T, μ_star, ω_star, exog_params(), i)
simulations += game
data = pd.DataFrame(simulations)
# +
mass = data.groupby(['batch','time','sex'], as_index=False).size()
fig, axs = plt.subplots(1,2, figsize=(6.267704441677044, 2.6368271881975483), constrained_layout=True, sharey=True)
axs[0].set_xlim(0,T)
axs[1].set_xlim(0,T)
for i in range(batches):
if i==0:
axs[0].plot(range(0,T), mass[(mass.batch==i)&(mass.sex=='Male')]['size'], color='tab:blue', label='Male')
axs[1].plot(range(0,T), mass[(mass.batch==i)&(mass.sex=='Female')]['size'], color='tab:pink', label='Female')
else:
axs[0].plot(range(0,T), mass[(mass.batch==i)&(mass.sex=='Male')]['size'], color='tab:blue')
axs[1].plot(range(0,T), mass[(mass.batch==i)&(mass.sex=='Female')]['size'], color='tab:pink')
axs[0].set(ylabel=r'Agent Stock ($N_t$)')
axs[0].axhline(y=Nm, color='r', linestyle='--', label='Steady State')
axs[1].axhline(y=Nw, color='r', linestyle='--', label='Steady State')
axs[0].legend()
axs[1].legend()
fig.supxlabel('Time ($t$)', size=10)
#plt.savefig('../../figures/abm-conv.png', bbox_inches='tight')
plt.show()
# -
# ## Varying Sample Sizes
# +
simulations = []
sstates = []
arrivals = [10, 100, 1000]
for i, l in enumerate(arrivals):
# Setting arrivals rate
print(f'Arrivals rate: {l}')
λm = l
λw = l
# Solve for steady state
m_test = np.random.rand(Bm)
w_test = np.random.rand(Bw)
x_start = np.concatenate((m_test, w_test), axis=None)
solution = least_squares(SSE, x_start, bounds = (0,1), verbose=1)
μ_star = solution.x[:Bm]
ω_star = solution.x[Bm:]
Nm, Nw, Pbm, Pbw, τm, τw, αm, αw = steady_state(μ_star, ω_star, True)
print(f'Steady State Mass: {Nm + Nw}')
sstates.append(Nm+Nw)
# Simulating game
game = simulate_game(T, μ_star, ω_star, exog_params(), i)
simulations += game
data = pd.DataFrame(simulations)
# +
mass = data.groupby(['batch','time'], as_index=False).size()
fig, axs = plt.subplots(1,3, figsize=(6.267704441677044, 1.8912181254650321), constrained_layout=True)
for i, l in enumerate(arrivals):
axs[i].set_xlim(0,T)
axs[i].plot(range(0,T), mass[(mass.batch==i)]['size'], color='k', label='Total Stock')
axs[i].axhline(y=sstates[i], color='r', linestyle='--', label='Steady State')
#axs[i].set(title=r'$\lambda=$' + str(l))
axs[i].legend()
axs[0].set(ylabel=r'Number of Agents ($N_t$)')
fig.supxlabel('Time ($t$)', size=10)
#plt.savefig('../../figures/abm-conv-ssize.png', bbox_inches='tight')
plt.show()
| code/abm/b-conv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="s_qNSzzyaCbD" pycharm={"name": "#%% md\n"}
# ##### Copyright 2019 The TensorFlow Authors and <NAME> 2020
#
# + pycharm={"name": "#%%\n"}
# ! which python
# ! python --version
# + cellView="form" id="jmjh290raIky" pycharm={"name": "#%%\n"}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + colab={"base_uri": "https://localhost:8080/"} id="EV1qWhrmI1nF" outputId="684fb7e9-4cea-4595-8da1-15cfa0ebaae6" pycharm={"name": "#%%\n"}
from cloudmesh.common.util import banner
import os
import sys
import socket
import pathlib
import humanize
from cloudmesh.common.console import Console
from cloudmesh.common.Shell import Shell
from cloudmesh.common.dotdict import dotdict
from cloudmesh.common.Printer import Printer
from cloudmesh.common.StopWatch import StopWatch
from cloudmesh.common.util import readfile
from cloudmesh.gpu.gpu import Gpu
from pprint import pprint
import sys
from IPython.display import display
import tensorflow_datasets as tfds
import tensorflow as tf
from tqdm.keras import TqdmCallback
from tqdm import tnrange
from tqdm import notebook
from tqdm import tqdm
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import LSTM
from tensorflow.keras.layers import GRU
from tensorflow.keras.layers import Dense
import os
import subprocess
import gc
from csv import reader
from csv import writer
import sys
import random
import math
import numpy as np
import matplotlib.pyplot as plt
from textwrap import wrap
import pandas as pd
import io as io
import string
import time
import datetime
from datetime import timedelta
from datetime import date
# TODO: better would be to distinguish them and not overwritethe one datetime with the other.
from datetime import datetime
import yaml
from typing import Dict
from typing import Tuple
from typing import Optional
from typing import List
from typing import Union
from typing import Callable
import matplotlib
import matplotlib.patches as patches
# import matplotlib.pyplot as plt
from matplotlib.figure import Figure
from matplotlib.path import Path
import matplotlib.dates as mdates
import scipy as sc
import scipy.linalg as solver
from matplotlib import colors
import enum
import pandas as pd
import abc
import json
import psutil
gregor = True
# %load_ext autotime
StopWatch.start("total")
in_colab = 'google.colab' in sys.modules
in_rivanna = "hpc.virginia.edu" in socket.getfqdn()
in_nonstandard = not in_colab and not in_rivanna
config = dotdict()
content = readfile("config.yaml")
program_config = dotdict(yaml.safe_load(content))
config.update(program_config)
print(Printer.attribute(config))
if in_colab:
# Avoids scroll-in-the-scroll in the entire Notebook
#test
from IPython.display import Javascript
def resize_colab_cell():
display(Javascript('google.colab.output.setIframeHeight(0, true, {maxHeight: 20000})'))
get_ipython().events.register('pre_run_cell', resize_colab_cell)
from google.colab import drive
drive.mount('/content/gdrive')
if in_rivanna or gregor:
tf.config.set_soft_device_placement(config.set_soft_device_placement)
tf.debugging.set_log_device_placement(config.debugging_set_log_device_placement)
# tf.test.is_gpu_available
import re
if in_rivanna:
test_localscratch = re.search('localscratch', str(sys.executable))
if test_localscratch != None:
in_localscratch = test_localscratch.group(0) == 'localscratch'
else:
in_localscratch = False
else:
in_localscratch = False
# + pycharm={"name": "#%%\n"}
def TIME_start(name):
banner(f"Start timer {name}")
StopWatch.start(name)
def TIME_stop(name):
StopWatch.stop(name)
t = StopWatch.get(name)
h = humanize.naturaldelta(timedelta(seconds=t))
banner(f"Stop timer {name}: {t}s or {h}")
# + pycharm={"name": "#%%\n"}
# who am i
config.user = Shell.run('whoami').strip()
try:
config.user_id = Shell.run('id -u').strip()
config.group_id = Shell.run('id -g').strip()
except subprocess.CalledProcessError:
print("The command <id> is not on your path.")
print(Printer.attribute(config))
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
StopWatch.benchmark()
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
r = Gpu().system()
try:
## Once Cloudmesh GPU PR2 is merged, the above block can be removed and the below be used.
config.gpuname = [x['product_name'] for x in r]
config.gpuvendor = [x.get('vendor', "Unknown Vendor") for x in r]
except:
pass
print (Printer.attribute(config))
# + pycharm={"name": "#%%\n"}
def SAVEFIG(plt, filename):
if ".png" in filename:
_filename = filename.replace(".png", "")
else:
_filename = filename
plt.savefig(f'{filename}.png', format='png')
plt.savefig(f'{filename}.pdf', format='pdf')
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
# Set Runname
RunName = 'EARTHQ-newTFTv29'
RunComment = ' TFT Dev on EarthQuakes -- 2 weeks 3 months 6 months 1 year d_model 160 dropout 0.1 Location Based Validation BS 64 Simple 2 layers CUDA LSTM MSE corrected MHA Plot 28'
#
# StopWatch.benchmark(sysinfo=True)
# make sure we have free memory in it
# replace the following and if needed read from StopWatch
memory = StopWatch.get_sysinfo()["mem.available"]
print(f'Your runtime has {memory} of available RAM\n')
config.runname = RunName
# + [markdown] id="LWNb05uZ7V9I" pycharm={"name": "#%% md\n"}
# # Initial System Code
# + colab={"base_uri": "https://localhost:8080/", "height": 852} id="w4cXSlPV7hNG" outputId="7fae2abb-b562-4da8-d7c6-5b581b32421f" pycharm={"name": "#%%\n"}
startbold = "\033[1m"
resetfonts = "\033[0m"
startred = '\033[31m'
startpurple = '\033[35m'
startyellowbkg = '\033[43m'
banner("System information")
r = StopWatch.systeminfo()
print (r)
# + pycharm={"name": "#%%\n"}
banner("nvidia-smi")
gpu_info = Shell.run("nvidia-smi")
if gpu_info.find('failed') >= 0:
print('Select the Runtime > "Change runtime type" menu to enable a GPU accelerator, ')
else:
print(gpu_info)
# + [markdown] id="J0Qjg6vuaHNt" pycharm={"name": "#%% md\n"}
# # Transformer model for science data based on original for language understanding
# + [markdown] id="AOpGoE2T-YXS" pycharm={"name": "#%% md\n"}
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/text/transformer">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/text/transformer.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/transformer.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/text/transformer.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="HA3Lx2aY1xeg" pycharm={"name": "#%% md\n"}
# ## Science Data Parameters and Sizes
# + [markdown] id="EMY9LokXwa9K" pycharm={"name": "#%% md\n"}
# -------
# Here is structure of science time series module. We will need several arrays that will need to be flattened at times. Note Python defaults to row major i.e. final index describes contiguous positions in memory
#
#
# At highest level data is labeled by Time and Location
#
# * Ttot is total number of time steps
# * Tseq is length of each sequence in time steps
# * Num_Seq is number of sequences in time: Num_Seq = Ttot-Tseq + 1
# * Nloc is Number of locations. The locations could be a 1D list or have array structure such as an image.
# * Nsample is number of data samples Nloc * Num_Seq
#
#
#
#
# Input data is at each location
# * Nprop time independent properties describing the location
# * Nforcing is number of time dependent forcing features INPUT at each time value
#
#
# Output (predicted) data at each location and for each time sequence is
# * Npred predicted time dependent values defined at every time step
# * Recorded at Nforecast time values measured wrt final time value of sequence
# * ForecastDay is an array of length Nforecast defining how many days into future prediction is. Typically ForecastDay[0] = 1 and Nforecast is often 1
# * There is also a class of science problems that are more similar to classic Seq2Seq. Here Nforecast = Tseq and ForecastDay = [-Tseq+1 ... 0]
# * We also support Nwishful predictions of events in future such probability of an earthquake of magnitude 6 in next 3 years. These are defined by araays EventType and Timestart, TimeInterval of length Nwishful. EventType is user defined and Timestart, TimeInterval is measured in time steps
# * Any missing output values should be set to NaN and Loss function must ensure that these points are ignored in derivative calculation and value calculation
#
# We have an input module that supports either LSTM or Transformer (multi-head attention) models
#
# Example Problem AICov
#
# * Ttot = 114
# * Tseq = 9
# * Num_Seq = 106
# * Nloc = 110
#
#
# * Nprop = 35
# * Nforcing = 5 including infections, fatalities, plus 3 temporal position variables (last 3 not in current version)
#
#
# * Npred = 2 (predicted infections and fatalities). Could be 5 if predicted temporal position of output)
# * Nforecast= 15
# * ForecastDay = [1, 2, .......14, 15]
# * Nwishful = 0
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
# + [markdown] id="_UlOJMJ31SoG" pycharm={"name": "#%% md\n"}
# ## Science Data Arrays
# + [markdown] id="BdszPs9on5gk" pycharm={"name": "#%% md\n"}
# Typical Arrays
#
#
# [ time, Location ] as Pandas array with label [name of time-dependent variable] as an array or just name of Pandas array
#
# time labels rows indexed by datetime or the difference datetime - start
#
# Non windowed data is stored with propert name as row index and location as column index
# [ static property, Location]
#
# Covid Input is
# [Sequence number 0..Num_Seq-1 ] [ Location 0..Nloc-1 ] [position in time sequence Tseq] [ Input Features]
#
# Covid Output is
# [Sequence number Num_Seq ] [ Location Nloc ] [ Output Features]
#
# Output Features are [ ipred = 0 ..Npred-1 ] [ iforecast = 0 ..Nforecast-1 ]
#
# Input Features are static fields followed by if present by dynamic system fields (cos-theta sin-theta linear) chosen followed by cases, deaths. In fact this is user chosen as they set static and dynamic system properties to use
# + [markdown] id="8-iizX9OKmI3" pycharm={"name": "#%% md\n"}
# We will have various numpy and pandas arrays where we designate label
#
# [Ttot] is all time values
#
# [Num_Seq] is all sequences of window size ***Tseq***
#
# We can select time values or sequences [Ttot-reason] [Num_Seq-reason] for a given "reason"
#
# [Num_Seq][Tseq] is all time values in all sequences
#
# [Nloc] is all locations while [Nloc-reason] is subset of locations for given "reason"
#
# [Model1] is initial embedding of each data point
#
# [Model1+TrPosEnc] is initial embedding of each data point with Transformer style positional encoding
#
# [Nforcing] is time dependent input parameters and [Nprop] static properties while [ExPosEnc] are explicit positional (temporal) encoding.
#
# [Nforcing+ExPosEnc+Nprop] are all possible inputs
#
# [Npred] is predicted values with [Npred+ExPosEnc] as predictions plus encodings with actually used [Predvals] = [Npred+ExPosEnc-Selout]
#
# [Predtimes] = [Forecast time range] are times forecasted with "time range" separately defined
#
#
# + [markdown] id="NdH4W3OJTLyj" pycharm={"name": "#%% md\n"}
# ### Define Basic Control parameters
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="mZAL5yNsC_UK" outputId="b45b3c2d-676a-4c27-dde9-4e28a18bbb06" pycharm={"name": "#%%\n"}
def wraptotext(textinput,size=None):
if size is None:
size = 120
textlist = wrap(textinput,size)
textresult = textlist[0]
for itext in range(1,len(textlist)):
textresult += '\n'+textlist[itext]
return textresult
def timenow():
now = datetime.now()
return now.strftime("%m/%d/%Y, %H:%M:%S") + " UTC"
def float32fromstrwithNaN(instr):
if instr == 'NaN':
return NaN
return np.float32(instr)
def printexit(exitmessage):
print(exitmessage)
sys.exit()
def strrnd(value):
return str(round(value,4))
NaN = np.float32("NaN")
ScaleProperties = False
ConvertDynamicPredictedQuantity = False
ConvertDynamicProperties = True
GenerateFutures = False
GenerateSequences = False
PredictionsfromInputs = False
RereadMay2020 = False
UseOLDCovariates = False
Dropearlydata = 0
NIHCovariates = False
UseFutures = True
Usedaystart = False
PopulationNorm = False
SymbolicWindows = False
Hydrology = False
Earthquake = False
EarthquakeImagePlots = False
AddSpecialstoSummedplots = False
UseRealDatesonplots = False
Dumpoutkeyplotsaspics = False
OutputNetworkPictures = False
NumpredbasicperTime = 2
NumpredFuturedperTime = 2
NumTimeSeriesCalculated = 0
Dailyunit = 1
TimeIntervalUnitName = 'Day'
InitialDate = datetime(2000,1,1)
NumberofTimeunits = 0
Num_Time =0
FinalDate = datetime(2000,1,1)
GlobalTrainingLoss = 0.0
GlobalValidationLoss = 0.0
# Type of Testing
LocationBasedValidation = False
LocationValidationFraction = 0.0
LocationTrainingfraction = 1.0
RestartLocationBasedValidation = False
global SeparateValandTrainingPlots
SeparateValandTrainingPlots = True
Plotsplitsize = -1 # if > 1 split time in plots
GarbageCollect = True
GarbageCollectionLimit = 0
current_time = timenow()
print(startbold + startred + current_time + ' ' + f'{RunName} {RunComment}' + resetfonts)
Earthquake = True
# + [markdown] id="DefDaYecDhIM" pycharm={"name": "#%% md\n"}
# ## Define input structure
#
# Read in data and set it up for Tensorflow with training and validation
#
# + [markdown] id="9kj1DvDTneDZ" pycharm={"name": "#%% md\n"}
# Set train_examples, val_examples as science training and validatioon set.
#
# The shuffling of Science Data needs some care. We have ***Tseq*** * size of {[Num_Seq][Nloc]} locations in each sample. In simplease case the last is just a decomposition over location; not over time. Let's Nloc-sel be number of locations per sample. It will be helpful if Nloc-sel is divisable by 2.
#
# Perhaps Nloc-sel = 2 6 or 10 is reasonable.
#
# Then you shuffle locations every epoch and divide them into groups of size Nloc-sel with 50% overlap so you get locations
#
# 0 1 2 3 4 5;
#
# 3 4 5 6 7 8;
#
# 6 7 8 9 10 11 etc.
#
# Every locations appears twice in an epoch (for each time value). You need to randomly add locations at end of sequence so it is divisiuble by Nloc-sel e.g add 4 random positions to the end if Nloc=110 and Nloc-sel = 6. Note last group of 6 has members 112 113 114 0 1 2
#
# After spatial structure set up, randomly shuffle in Num_Seq where there is an argument to do all locations for a partcular time value together.
#
# For validation, it is probably best to select validation location before chopping them into groups of size Nloc-sel
#
# How one groups locations for inference is not clear. One idea is to take trained network and use it to find for each location which other locations have the most attention with it. Use those locations in prediction
# + [markdown] id="YKm_MgRMdcTT" pycharm={"name": "#%% md\n"}
# More general input.
# NaN allowed value
#
# * Number time values
# * Number locations
# * Number driving values
# * Number predicted values
#
# For COVID driving same as predicted
#
# * a) Clean up >=0 daily
# * b) Normalize
# * c) Add Futures
# * d) Add time/location encoding
#
# + [markdown] id="9KJIxYoMDZOu" pycharm={"name": "#%% md\n"}
# ### Setup File Systems
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="ShbYhXJbKCDT" outputId="f9f98b16-db80-441c-8809-ba54de7cf782" pycharm={"name": "#%%\n"}
if in_colab:
# find the data
COLABROOTDIR="/content/gdrive/My Drive/Colab Datasets"
elif in_rivanna:
if in_localscratch:
#Shell.mkdir(f'{config["run"]["datadir"]}/EarthquakeDec2020/Outputs')
Shell.mkdir(f'{config["run.datadir"]}/EarthquakeDec2020/Outputs')
COLABROOTDIR=str(pathlib.Path(config["run.datadir"]).resolve())
else:
Shell.mkdir('../data/EarthquakeDec2020/Outputs')
COLABROOTDIR=str(pathlib.Path("../data").resolve())
else:
#Shell.mkdir(f'{config["run"]["datadir"]}/EarthquakeDec2020/Outputs')
Shell.mkdir(f'{config["run.datadir"]}/EarthquakeDec2020/Outputs')
COLABROOTDIR=str(pathlib.Path(config["run.datadir"]).resolve())
print (COLABROOTDIR)
if not os.path.exists(COLABROOTDIR):
Console.error(f"Missing data directory: {COLABROOTDIR}")
sys.exit(1)
os.environ["COLABROOTDIR"] = COLABROOTDIR
APPLDIR=f"{COLABROOTDIR}/EarthquakeDec2020"
CHECKPOINTDIR = f"{APPLDIR}/checkpoints/{RunName}dir/"
Shell.mkdir(CHECKPOINTDIR)
print(f'Checkpoint set up in directory {CHECKPOINTDIR}')
config.checkpointdir = CHECKPOINTDIR
config.appldir = APPLDIR
config.colabrootdir = COLABROOTDIR
print(Printer.attribute(config))
# + [markdown] id="vX4_pGSonAyz" pycharm={"name": "#%% md\n"}
# ### Space Filling Curves
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="z-FrNiAY0ALF" outputId="7954dcdf-a19c-43d5-dfbc-670f0e10414e" pycharm={"name": "#%%\n"}
def cal_gilbert2d(width: int, height: int) -> List[Tuple[int, int]]:
coordinates: List[Tuple[int, int]] = []
def sgn(x: int) -> int:
return (x > 0) - (x < 0)
def gilbert2d(x: int, y: int, ax: int, ay: int, bx: int, by: int):
"""
Generalized Hilbert ('gilbert') space-filling curve for arbitrary-sized
2D rectangular grids.
"""
w = abs(ax + ay)
h = abs(bx + by)
(dax, day) = (sgn(ax), sgn(ay)) # unit major direction
(dbx, dby) = (sgn(bx), sgn(by)) # unit orthogonal direction
if h == 1:
# trivial row fill
for i in range(0, w):
coordinates.append((x, y))
(x, y) = (x + dax, y + day)
return
if w == 1:
# trivial column fill
for i in range(0, h):
coordinates.append((x, y))
(x, y) = (x + dbx, y + dby)
return
(ax2, ay2) = (ax // 2, ay // 2)
(bx2, by2) = (bx // 2, by // 2)
w2 = abs(ax2 + ay2)
h2 = abs(bx2 + by2)
if 2 * w > 3 * h:
if (w2 % 2) and (w > 2):
# prefer even steps
(ax2, ay2) = (ax2 + dax, ay2 + day)
# long case: split in two parts only
gilbert2d(x, y, ax2, ay2, bx, by)
gilbert2d(x + ax2, y + ay2, ax - ax2, ay - ay2, bx, by)
else:
if (h2 % 2) and (h > 2):
# prefer even steps
(bx2, by2) = (bx2 + dbx, by2 + dby)
# standard case: one step up, one long horizontal, one step down
gilbert2d(x, y, bx2, by2, ax2, ay2)
gilbert2d(x + bx2, y + by2, ax, ay, bx - bx2, by - by2)
gilbert2d(x + (ax - dax) + (bx2 - dbx), y + (ay - day) + (by2 - dby), -bx2, -by2, -(ax - ax2), -(ay - ay2))
if width >= height:
gilbert2d(0, 0, width, 0, 0, height)
else:
gilbert2d(0, 0, 0, height, width, 0)
return coordinates
def lookup_color(unique_colors, color_value: float) -> int:
ids = np.where(unique_colors == color_value)
color_id = ids[0][0]
return color_id
def plot_gilbert2d_space_filling(
vertices: List[Tuple[int, int]],
width: int,
height: int,
filling_color: Optional[np.ndarray] = None,
color_map: str = "rainbow",
figsize: Tuple[int, int] = (12, 8),
linewidth: int = 1,
) -> None:
fig, ax = plt.subplots(figsize=figsize)
patch_list: List = []
if filling_color is None:
cmap = matplotlib.cm.get_cmap(color_map, len(vertices))
for i in range(len(vertices) - 1):
path = Path([vertices[i], vertices[i + 1]], [Path.MOVETO, Path.LINETO])
patch = patches.PathPatch(path, fill=False, edgecolor=cmap(i), lw=linewidth)
patch_list.append(patch)
ax.set_xlim(-1, width)
ax.set_ylim(-1, height)
else:
unique_colors = np.unique(filling_color)
# np.random.shuffle(unique_colors)
cmap = matplotlib.cm.get_cmap(color_map, len(unique_colors))
for i in range(len(vertices) - 1):
x, y = vertices[i]
fi, fj = x, height - 1 - y
color_value = filling_color[fj, fi]
color_id = lookup_color(unique_colors, color_value)
path = Path(
[rescale_xy(x, y), rescale_xy(vertices[i + 1][0], vertices[i + 1][1])], [Path.MOVETO, Path.LINETO]
)
# path = Path([vertices[i], vertices[i + 1]], [Path.MOVETO, Path.LINETO])
patch = patches.PathPatch(path, fill=False, edgecolor=cmap(color_id), lw=linewidth)
patch_list.append(patch)
ax.set_xlim(-120 - 0.1, width / 10 - 120)
ax.set_ylim(32 - 0.1, height / 10 + 32)
collection = matplotlib.collections.PatchCollection(patch_list, match_original=True)
# collection.set_array()
# plt.colorbar(collection)
ax.add_collection(collection)
ax.set_aspect("equal")
plt.show()
return
def rescale_xy(x: int, y: int) -> Tuple[float, float]:
return x / 10 - 120, y / 10 + 32
def remapfaults(InputFaultNumbers, Numxlocations, Numylocations, SpaceFillingCurve):
TotalLocations = Numxlocations*Numylocations
OutputFaultNumbers = np.full_like(InputFaultNumbers, -1, dtype=int)
MaxOldNumber = np.amax(InputFaultNumbers)
mapping = np.full(MaxOldNumber+1, -1,dtype=int)
newlabel=-1
for sfloc in range(0, TotalLocations):
[x,y] = SpaceFillingCurve[sfloc]
pixellocation = y*Numxlocations + x
pixellocation1 = y*Numxlocations + x
oldfaultnumber = InputFaultNumbers[pixellocation1]
if mapping[oldfaultnumber] < 0:
newlabel += 1
mapping[oldfaultnumber] = newlabel
OutputFaultNumbers[pixellocation] = mapping[oldfaultnumber]
MinNewNumber = np.amin(OutputFaultNumbers)
if MinNewNumber < 0:
printexit('Incorrect Fault Mapping')
print('new Fault Labels generated 0 through ' + str(newlabel))
plot_gilbert2d_space_filling(SpaceFillingCurve,Numxlocations, Numylocations, filling_color = np.reshape(OutputFaultNumbers,(40,60)), color_map="gist_ncar")
return OutputFaultNumbers
def annotate_faults_ndarray(pix_faults: np.ndarray, figsize=(10, 8), color_map="rainbow"):
matplotlib.rcParams.update(matplotlib.rcParamsDefault)
plt.rcParams.update({"font.size": 12})
unique_colors = np.unique(pix_faults)
np.random.shuffle(unique_colors)
cmap = matplotlib.cm.get_cmap(color_map, len(unique_colors))
fig, ax = plt.subplots(figsize=figsize)
height, width = pix_faults.shape
for j in range(height):
for i in range(width):
x, y = i / 10 - 120, (height - j - 1) / 10 + 32
ax.annotate(str(pix_faults[j, i]), (x + 0.05, y + 0.05), ha="center", va="center")
color_id = lookup_color(unique_colors, pix_faults[j, i])
ax.add_patch(patches.Rectangle((x, y), 0.1, 0.1, color=cmap(color_id), alpha=0.5))
ax.set_xlim(-120, width / 10 - 120)
ax.set_ylim(32, height / 10 + 32)
plt.show()
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 6319} id="3WchUjkEnMxD" outputId="6648f4af-5d60-4713-a0dd-7f11bd9c2c9f" pycharm={"name": "#%% md\n"}
# # CELL READ DATA
# + pycharm={"name": "#%%\n"}
def makeadateplot(plotfigure,plotpointer, Dateaxis=None, datemin=None, datemax=None, Yearly=True, majoraxis = 5):
if not Yearly:
sys.exit('Only yearly supported')
plt.rcParams.update({'font.size': 9})
years5 = mdates.YearLocator(majoraxis) # every 5 years
years_fmt = mdates.DateFormatter('%Y')
plotpointer.xaxis.set_major_locator(years5)
plotpointer.xaxis.set_major_formatter(years_fmt)
if datemin is None:
datemin = np.datetime64(Dateaxis[0], 'Y')
if datemax is None:
datemax = np.datetime64(Dateaxis[-1], 'Y') + np.timedelta64(1, 'Y')
plotpointer.set_xlim(datemin, datemax)
plotfigure.autofmt_xdate()
return datemin, datemax
def makeasmalldateplot(figure,ax, Dateaxis):
plt.rcParams.update({'font.size': 9})
months = mdates.MonthLocator(interval=2) # every month
datemin = np.datetime64(Dateaxis[0], 'M')
datemax = np.datetime64(Dateaxis[-1], 'M') + np.timedelta64(1, 'M')
ax.set_xlim(datemin, datemax)
months_fmt = mdates.DateFormatter('%y-%b')
locator = mdates.AutoDateLocator()
locator.intervald['MONTHLY'] = [2]
formatter = mdates.ConciseDateFormatter(locator)
# ax.xaxis.set_major_locator(locator)
# ax.xaxis.set_major_formatter(formatter)
ax.xaxis.set_major_locator(months)
ax.xaxis.set_major_formatter(months_fmt)
figure.autofmt_xdate()
return datemin, datemax
def Addfixedearthquakes(plotpointer,graphmin, graphmax, ylogscale = False, quakecolor = None, Dateplot = True, vetoquake = None):
if vetoquake is None: # Vetoquake = True means do not plot this quake
vetoquake = np.full(numberspecialeqs, False, dtype = bool)
if quakecolor is None: # Color of plot
quakecolor = 'black'
Place =np.arange(numberspecialeqs, dtype =int)
Place[8] = 11
Place[10] = 3
Place[12] = 16
Place[7] = 4
Place[2] = 5
Place[4] = 14
Place[11] = 18
ymin, ymax = plotpointer.get_ylim() # Or work with transform=ax.transAxes
for iquake in range(0,numberspecialeqs):
if vetoquake[iquake]:
continue
# This is the x position for the vertical line
if Dateplot:
x_line_annotation = Specialdate[iquake] # numpy date format
else:
x_line_annotation = Numericaldate[iquake] # Float where each interval 1 and start is 0
if (x_line_annotation < graphmin) or (x_line_annotation > graphmax):
continue
# This is the x position for the label
if Dateplot:
x_text_annotation = x_line_annotation + np.timedelta64(5*Dailyunit,'D')
else:
x_text_annotation = x_line_annotation + 5.0
# Draw a line at the position
plotpointer.axvline(x=x_line_annotation, linestyle='dashed', alpha=1.0, linewidth = 0.5, color=quakecolor)
# Draw a text
if Specialuse[iquake]:
ascii = str(round(Specialmags[iquake],1)) + '\n' + Specialeqname[iquake]
if ylogscale:
yminl = max(0.01*ymax,ymin)
yminl = math.log(yminl,10)
ymaxl = math.log(ymax,10)
logyplot = yminl + (0.1 + 0.8*(float(Place[iquake])/float(numberspecialeqs-1)))*(ymaxl-yminl)
yplot = pow(10, logyplot)
else:
yplot = ymax - (0.1 + 0.8*(float(Place[iquake])/float(numberspecialeqs-1)))*(ymax-ymin)
if Dateplot:
if x_text_annotation > graphmax - np.timedelta64(2000, 'D'):
x_text_annotation = graphmax - np.timedelta64(2000, 'D')
else:
if x_text_annotation > graphmax - 100:
x_text_annotation = graphmax - 100
# print(str(yplot) + " " + str(ymin) + " " + str(ymax) + " " + str(x_text_annotation) + " " + str(x_line_annotation)) + " " + ascii
plotpointer.text(x=x_text_annotation, y=yplot, s=wraptotext(ascii,size=10), alpha=1.0, color='black', fontsize = 6)
def quakesearch(iquake, iloc):
# see if top earthquake iquake llies near location iloc
# result = 0 NO; =1 YES Primary: locations match exactly; = -1 Secondary: locations near
# iloc is location before mapping
xloc = iloc%60
yloc = (iloc - xloc)/60
if (xloc == Specialxpos[iquake]) and (yloc == Specialypos[iquake]):
return 1
if (abs(xloc - Specialxpos[iquake]) <= 1) and (abs(yloc - Specialypos[iquake]) <= 1):
return -1
return 0
# Read Earthquake Data
def log_sum_exp10(ns, sumaxis =0):
max_v = np.max(ns, axis=None)
ds = ns - max_v
sum_of_exp = np.power(10, ds).sum(axis=sumaxis)
return max_v + np.log10(sum_of_exp)
def log_energyweightedsum(nvalue, ns, sumaxis = 0):
max_v = np.max(ns, axis=None)
ds = ns - max_v
ds = np.power(10, 1.5*ds)
dvalue = (np.multiply(nvalue,ds)).sum(axis=sumaxis)
ds = ds.sum(axis=0)
return np.divide(dvalue,ds)
# Set summed magnitude as log summed energy = 10^(1.5 magnitude)
def log_energy(mag, sumaxis =0):
return log_sum_exp10(1.5 * mag, sumaxis = sumaxis) / 1.5
def AggregateEarthquakes(itime, DaysDelay, DaysinInterval, Nloc, Eqdata, Approach, weighting = None):
if (itime + DaysinInterval + DaysDelay) > NumberofTimeunits:
return np.full([Nloc],NaN,dtype = np.float32)
if Approach == 0: # Magnitudes
if MagnitudeMethod == 0:
TotalMagnitude = log_energy(Eqdata[itime +DaysDelay:itime+DaysinInterval+DaysDelay])
else:
TotalMagnitude = Eqdata[itime +DaysDelay:itime+DaysinInterval+DaysDelay,:].sum(axis=0)
return TotalMagnitude
if Approach == 1: # Depth -- energy weighted
WeightedResult = log_energyweightedsum(Eqdata[itime +DaysDelay:itime+DaysinInterval+DaysDelay],
weighting[itime +DaysDelay:itime+DaysinInterval+DaysDelay])
return WeightedResult
if Approach == 2: # Multiplicity -- summed
SimpleSum = Eqdata[itime +DaysDelay:itime+DaysinInterval+DaysDelay,:].sum(axis=0)
return SimpleSum
def TransformMagnitude(mag):
if MagnitudeMethod == 0:
return mag
if MagnitudeMethod == 1:
return np.power(10, 0.375*(mag-3.29))
return np.power(10, 0.75*(mag-3.29))
# Change Daily Unit
# Accumulate data in Dailyunit chunks.
# This changes data so it looks like daily data bu really collections of chunked data.
# For earthquakes, the aggregations uses energy averaging for depth and magnitude. It just adds for multiplicity
def GatherUpData(OldInputTimeSeries):
Skipped = NumberofTimeunits%Dailyunit
NewInitialDate = InitialDate + timedelta(days=Skipped)
NewNum_Time = int(Num_Time/Dailyunit)
NewFinalDate = NewInitialDate + Dailyunit * timedelta(days=NewNum_Time-1)
print(' Daily Unit ' +str(Dailyunit) + ' number of ' + TimeIntervalUnitName + ' Units ' + str(NewNum_Time)+ ' ' +
NewInitialDate.strftime("%d/%m/%Y") + ' To ' + NewFinalDate.strftime("%d/%m/%Y"))
NewInputTimeSeries = np.empty([NewNum_Time,Nloc,NpropperTimeDynamicInput],dtype = np.float32)
for itime in range(0,NewNum_Time):
NewInputTimeSeries[itime,:,0] = AggregateEarthquakes(Skipped + itime*Dailyunit,0,Dailyunit, Nloc,
BasicInputTimeSeries[:,:,0], 0)
NewInputTimeSeries[itime,:,1] = AggregateEarthquakes(Skipped + itime*Dailyunit,0,Dailyunit, Nloc,
BasicInputTimeSeries[:,:,1], 1,
weighting = BasicInputTimeSeries[:,:,0])
NewInputTimeSeries[itime,:,2] = AggregateEarthquakes(Skipped + itime*Dailyunit,0,Dailyunit, Nloc,
BasicInputTimeSeries[:,:,2], 2)
NewInputTimeSeries[itime,:,3] = AggregateEarthquakes(Skipped + itime*Dailyunit,0,Dailyunit, Nloc,
BasicInputTimeSeries[:,:,3], 2)
return NewInputTimeSeries, NewNum_Time, NewNum_Time, NewInitialDate, NewFinalDate
# Daily Read in Version
if Earthquake:
read1950 = True
Eigenvectors = 2
UseEarthquakeEigenSystems = False
Dailyunit = 14
addwobblingposition = False
r = Shell.ls(config.appldir)
print (r)
if read1950:
MagnitudeDataFile = APPLDIR + '/1950start/SC_1950-2019.freq-D-25567x2400-log_eng.multi.csv'
DepthDataFile = APPLDIR + '/1950start/SC_1950-2019.freq-D-25567x2400-w_depth.multi.csv'
MultiplicityDataFile = APPLDIR + '/1950start/SC_1950-2019.freq-D-25567x2400-n_shock.multi.csv'
RundleMultiplicityDataFile = APPLDIR + '/1950start/SC_1950-2019.freq-D-25567x2400-n_shock-mag-3.29.multi.csv'
NumberofTimeunits = 25567
InitialDate = datetime(1950,1,1)
else:
MagnitudeDataFile = APPLDIR + '/SC_1990-2019.freq-D-10759x2400.csv'
DepthDataFile = APPLDIR + '/SC_1990-2019.freq-D-w_depth-10759x2400.multi.csv'
MultiplicityDataFile = APPLDIR + '/SC_1990-2019.freq-D-num_evts-10759x2400.csv'
RundleMultiplicityDataFile = APPLDIR + '/SC_1990-2019.freq-D-10755x2400-n_shock-mag-3.29.multi.csv'
NumberofTimeunits = 10759
InitialDate = datetime(1990,1,1)
Topearthquakesfile = APPLDIR + '/topearthquakes_20.csv'
FaultLabelDataFile = APPLDIR + '/pix_faults_SmallJan21.csv'
MagnitudeMethod = 0
ReadFaultMethod = 2 # one set of x values for each input row
Numberxpixels = 60
Numberypixels = 40
Numberpixels = Numberxpixels*Numberypixels
Nloc = Numberpixels
Nlocdimension = 2
Nlocaxislengths = np.array((Numberxpixels,Numberypixels), ndmin = 1, dtype=int) # First row is top (north)
vertices = cal_gilbert2d(Numberxpixels,Numberypixels)
# print(vertices[0], vertices[1],vertices[2399], vertices[1198], vertices[1199],vertices[1200], vertices[1201])
sfcurvelist = vertices
plot_gilbert2d_space_filling(sfcurvelist, Numberxpixels, Numberypixels)
Dropearlydata = 0
FinalDate = InitialDate + timedelta(days=NumberofTimeunits-1)
print(startbold + startred + InitialDate.strftime("%d/%m/%Y") + ' To ' + FinalDate.strftime("%d/%m/%Y")
+ ' days ' + str(NumberofTimeunits) + resetfonts)
print( ' Pixels ' + str(Nloc) + ' x dimension ' + str(Nlocaxislengths[0]) + ' y dimension ' + str(Nlocaxislengths[1]) )
# Set up location information
Num_Time = NumberofTimeunits
NFIPS = Numberpixels
Locationname = [''] * NFIPS
Locationstate = [' '] * NFIPS
Locationpopulation = np.ones(NFIPS, dtype=int)
Locationfips = np.empty(NFIPS, dtype=int) # integer version of FIPs
Locationcolumns = [] # String version of FIPS
FIPSintegerlookup = {}
FIPSstringlookup = {}
for iloc in range (0, Numberpixels):
localfips = iloc
xvalue = localfips%Nlocaxislengths[0]
yvalue = np.floor(localfips/Nlocaxislengths[0])
Stringfips = str(xvalue) + ',' + str(yvalue)
Locationcolumns.append(Stringfips)
Locationname[iloc] = Stringfips
Locationfips[iloc] = localfips
FIPSintegerlookup[localfips] = localfips
FIPSstringlookup[Stringfips] = localfips
# TimeSeries 0 magnitude 1 depth 2 Multiplicity 3 Rundle Multiplicity
NpropperTimeDynamicInput = 4
BasicInputTimeSeries = np.empty([Num_Time,Nloc,NpropperTimeDynamicInput],dtype = np.float32)
# StaticProps 0...NumFaultLabels-1 Fault Labels
NumFaultLabels = 4
BasicInputStaticProps = np.empty([Nloc,NumFaultLabels],dtype = np.float32)
RawFaultData = np.empty(Nloc,dtype = int)
# Read in Magnitude Data into BasicInputTimeSeries
with open(MagnitudeDataFile, 'r') as read_obj:
csv_reader = reader(read_obj)
header = next(csv_reader)
Ftype = header[0]
if Ftype != '':
printexit('EXIT: Wrong header on line 1 ' + Ftype + ' of ' + MagnitudeDataFile)
itime = 0
for nextrow in csv_reader:
if len(nextrow)!=Numberpixels + 1:
printexit('EXIT: Incorrect row length Magnitude ' + str(itime) + ' ' +str(len(nextrow)))
localtime = nextrow[0]
if itime != int(localtime):
printexit('EXIT: Unexpected Time in Magnitude ' + localtime + ' ' +str(itime))
for iloc in range(0, Numberpixels):
BasicInputTimeSeries[itime,iloc,0] = TransformMagnitude(float(nextrow[iloc + 1]))
itime += 1
if itime != Num_Time:
printexit('EXIT Inconsistent time lengths in Magnitude Data ' +str(itime) + ' ' + str(Num_Time))
print('Read Magnitude data locations ' + str(Nloc) + ' Time Steps ' + str(Num_Time))
# End Reading in Magnitude data
# Read in Depth Data into BasicInputTimeSeries
with open(DepthDataFile, 'r') as read_obj:
csv_reader = reader(read_obj)
header = next(csv_reader)
Ftype = header[0]
if Ftype != '':
printexit('EXIT: Wrong header on line 1 ' + Ftype + ' of ' + DepthDataFile)
itime = 0
for nextrow in csv_reader:
if len(nextrow)!=Numberpixels + 1:
printexit('EXIT: Incorrect row length Depth ' + str(itime) + ' ' +str(len(nextrow)))
localtime = nextrow[0]
if itime != int(localtime):
printexit('EXIT: Unexpected Time in Depth ' + localtime + ' ' +str(itime))
for iloc in range(0, Numberpixels):
BasicInputTimeSeries[itime,iloc,1] = nextrow[iloc + 1]
itime += 1
if itime != Num_Time:
printexit('EXIT Inconsistent time lengths in Depth Data ' +str(itime) + ' ' + str(Num_Time))
print('Read Depth data locations ' + str(Nloc) + ' Time Steps ' + str(Num_Time))
# End Reading in Depth data
# Read in Multiplicity Data into BasicInputTimeSeries
with open(MultiplicityDataFile, 'r') as read_obj:
csv_reader = reader(read_obj)
header = next(csv_reader)
Ftype = header[0]
if Ftype != '':
printexit('EXIT: Wrong header on line 1 ' + Ftype + ' of ' + MultiplicityDataFile)
itime = 0
for nextrow in csv_reader:
if len(nextrow)!=Numberpixels + 1:
printexit('EXIT: Incorrect row length Multiplicity ' + str(itime) + ' ' +str(len(nextrow)))
localtime = nextrow[0]
if itime != int(localtime):
printexit('EXIT: Unexpected Time in Multiplicity ' + localtime + ' ' +str(itime))
for iloc in range(0, Numberpixels):
BasicInputTimeSeries[itime,iloc,2] = nextrow[iloc + 1]
itime += 1
if itime != Num_Time:
printexit('EXIT Inconsistent time lengths in Multiplicity Data ' +str(itime) + ' ' + str(Num_Time))
print('Read Multiplicity data locations ' + str(Nloc) + ' Time Steps ' + str(Num_Time))
# End Reading in Multiplicity data
# Read in Rundle Multiplicity Data into BasicInputTimeSeries
with open(RundleMultiplicityDataFile, 'r') as read_obj:
csv_reader = reader(read_obj)
header = next(csv_reader)
Ftype = header[0]
if Ftype != '':
printexit('EXIT: Wrong header on line 1 ' + Ftype + ' of ' + RundleMultiplicityDataFile)
itime = 0
for nextrow in csv_reader:
if len(nextrow)!=Numberpixels + 1:
printexit('EXIT: Incorrect row length Rundle Multiplicity ' + str(itime) + ' ' +str(len(nextrow)))
localtime = nextrow[0]
if itime != int(localtime):
printexit('EXIT: Unexpected Time in Rundle Multiplicity ' + localtime + ' ' +str(itime))
for iloc in range(0, Numberpixels):
BasicInputTimeSeries[itime,iloc,3] = nextrow[iloc + 1]
itime += 1
if itime != Num_Time:
printexit('EXIT Inconsistent time lengths in Rundle Multiplicity Data ' +str(itime) + ' ' + str(Num_Time))
print('Read Rundle Multiplicity data locations ' + str(Nloc) + ' Time Steps ' + str(Num_Time))
# End Reading in Rundle Multiplicity data
# Read in Top Earthquake Data
numberspecialeqs = 20
Specialuse = np.full(numberspecialeqs, True, dtype=bool)
Specialuse[14] = False
Specialuse[15] = False
Specialuse[18] = False
Specialuse[19] = False
Specialmags = np.empty(numberspecialeqs, dtype=np.float32)
Specialdepth = np.empty(numberspecialeqs, dtype=np.float32)
Speciallong = np.empty(numberspecialeqs, dtype=np.float32)
Speciallat = np.empty(numberspecialeqs, dtype=np.float32)
Specialdate = np.empty(numberspecialeqs, dtype = 'datetime64[D]')
Specialxpos = np.empty(numberspecialeqs, dtype=np.int32)
Specialypos = np.empty(numberspecialeqs, dtype=np.int32)
Specialeqname = []
with open(Topearthquakesfile, 'r') as read_obj:
csv_reader = reader(read_obj)
header = next(csv_reader)
Ftype = header[0]
if Ftype != 'date':
printexit('EXIT: Wrong header on line 1 ' + Ftype + ' of ' + Topearthquakesfile)
iquake = 0
for nextrow in csv_reader:
if len(nextrow)!=6:
printexit('EXIT: Incorrect row length Special Earthquakes ' + str(iquake) + ' ' +str(len(nextrow)))
Specialdate[iquake] = nextrow[0]
Speciallong[iquake] = nextrow[1]
Speciallat[iquake] = nextrow[2]
Specialmags[iquake] = nextrow[3]
Specialdepth[iquake] = nextrow[4]
Specialeqname.append(nextrow[5])
ixpos = math.floor((Speciallong[iquake]+120.0)*10.0)
ixpos = max(0,ixpos)
ixpos = min(59,ixpos)
iypos = math.floor((36.0-Speciallat[iquake])*10.0)
iypos = max(0,iypos)
iypos = min(39,iypos)
Specialxpos[iquake] = ixpos
Specialypos[iquake] = iypos
iquake += 1
for iquake in range(0,numberspecialeqs):
line = str(iquake) + ' mag ' + str(round(Specialmags[iquake],1)) + ' Lat/Long '
line += str(round(Speciallong[iquake],2)) + ' ' + str(round(Speciallong[iquake],2)) + ' ' + np.datetime_as_string(Specialdate[iquake])
line += Specialeqname[iquake]
print(line)
# Possibly change Unit
current_time = timenow()
print(startbold + startred + current_time + ' Data read in ' + RunName + ' ' + RunComment + resetfonts)
if Dailyunit != 1:
if Dailyunit == 14:
TimeIntervalUnitName = 'Fortnight'
if Dailyunit == 28:
TimeIntervalUnitName = 'LunarMonth'
BasicInputTimeSeries, NumberofTimeunits, Num_Time, InitialDate, FinalDate = GatherUpData(BasicInputTimeSeries)
current_time = timenow()
print(startbold + startred + current_time + ' Data unit changed ' +RunName + ' ' + RunComment + resetfonts)
Dateaxis = np.empty(Num_Time, dtype = 'datetime64[D]')
Dateaxis[0] = np.datetime64(InitialDate).astype('datetime64[D]')
for idate in range(1,Num_Time):
Dateaxis[idate] = Dateaxis[idate-1] + np.timedelta64(Dailyunit,'D')
for idate in range(0,Num_Time):
Dateaxis[idate] = Dateaxis[idate] + np.timedelta64(int(Dailyunit/2),'D')
print('Mid unit start time ' + np.datetime_as_string(Dateaxis[0]))
Totalmag = np.zeros(Num_Time,dtype = np.float32)
Totalefourthroot = np.zeros(Num_Time,dtype = np.float32)
Totalesquareroot = np.zeros(Num_Time,dtype = np.float32)
Totaleavgedmag = np.zeros(Num_Time,dtype = np.float32)
Totalmult = np.zeros(Num_Time,dtype = np.float32)
Totalmag[:] = BasicInputTimeSeries[:,:,0].sum(axis=1)
Totaleavgedmag = log_energy(BasicInputTimeSeries[:,:,0], sumaxis=1)
Totalmult[:] = BasicInputTimeSeries[:,:,3].sum(axis=1)
MagnitudeMethod = 1
Tempseries = TransformMagnitude(BasicInputTimeSeries[:,:,0])
Totalefourthroot = Tempseries.sum(axis=1)
MagnitudeMethod = 2
Tempseries = TransformMagnitude(BasicInputTimeSeries[:,:,0])
Totalesquareroot = Tempseries.sum(axis=1)
MagnitudeMethod = 0
basenorm = Totalmult.max(axis=0)
magnorm = Totalmag.max(axis=0)
eavgedmagnorm = Totaleavgedmag.max(axis=0)
efourthrootnorm = Totalefourthroot.max(axis=0)
esquarerootnorm = Totalesquareroot.max(axis=0)
print('Maximum Mult ' + str(round(basenorm,2)) + ' Mag 0.15 ' + str(round(magnorm,2))
+ ' E-avg 0.5 ' + str(round(eavgedmagnorm,2)) + ' E^0.25 1.0 ' + str(round(efourthrootnorm,2))
+ ' E^0.5 1.0 ' + str(round(esquarerootnorm,2)) )
Totalmag = np.multiply(Totalmag, 0.15*basenorm/magnorm)
Totaleavgedmag = np.multiply(Totaleavgedmag, 0.5*basenorm/eavgedmagnorm)
Totalefourthroot= np.multiply(Totalefourthroot, basenorm/efourthrootnorm)
Totalesquareroot= np.multiply(Totalesquareroot, basenorm/esquarerootnorm)
plt.rcParams["figure.figsize"] = [16,8]
figure, ax = plt.subplots()
datemin, datemax = makeadateplot(figure, ax, Dateaxis)
ax.plot(Dateaxis, Totalmult, label='Multiplicity')
ax.plot(Dateaxis, Totalmag, label='Summed Magnitude')
ax.plot(Dateaxis, Totaleavgedmag, label='E-averaged Magnitude')
ax.plot(Dateaxis, Totalefourthroot, label='Summed E^0.25')
ax.plot(Dateaxis, Totalesquareroot, label='Summed E^0.5')
ax.set_title('Observables summed over space')
ax.set_xlabel("Years")
ax.set_ylabel("Mult/Mag/Energy")
ax.grid(True)
ax.legend(loc='upper right')
Addfixedearthquakes(ax, datemin, datemax)
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
figure.tight_layout()
plt.show()
else:
print(' Data unit is the day and input this way')
Dateaxis = np.empty(Num_Time, dtype = 'datetime64[D]')
Dateaxis[0] = np.datetime64(InitialDate).astype('datetime64[D]')
for idate in range(1,Num_Time):
Dateaxis[idate] = Dateaxis[idate-1] + np.timedelta64(Dailyunit,'D')
for idate in range(0,Num_Time):
Dateaxis[idate] = Dateaxis[idate] + np.timedelta64(int(Dailyunit/2),'D')
print('Mid unit start time ' + np.datetime_as_string(Dateaxis[0]))
# Read in Fault Label Data into BasicInputStaticProps
# No header for data
with open(FaultLabelDataFile, 'r') as read_obj:
csv_reader = reader(read_obj)
iloc = 0
if ReadFaultMethod ==1:
for nextrow in csv_reader:
if len(nextrow)!=1:
printexit('EXIT: Incorrect row length Fault Label Data ' + str(iloc) + ' ' + str(len(nextrow)))
RawFaultData[iloc] = nextrow[0]
iloc += 1
else:
for nextrow in csv_reader:
if len(nextrow)!=Numberxpixels:
printexit('EXIT: Incorrect row length Fault Label Data ' + str(iloc) + ' ' + str(len(nextrow)) + ' ' + str(Numberxpixels))
for jloc in range(0, len(nextrow)):
RawFaultData[iloc] = nextrow[jloc]
iloc += 1
if iloc != Nloc:
printexit('EXIT Inconsistent location lengths in Fault Label Data ' +str(iloc) + ' ' + str(Nloc))
print('Read Fault Label data locations ' + str(Nloc))
# End Reading in Fault Label data
if NumFaultLabels == 1:
BasicInputStaticProps[:,0] = RawFaultData.astype(np.float32)
else: # remap fault label more reasonably
unique, counts = np.unique(RawFaultData, return_counts=True)
num = len(unique)
print('Number Fault Collections ' + str(num))
# for i in range(0,num):
# print(str(unique[i]) + ' ' + str(counts[i]))
BasicInputStaticProps[:,0] = remapfaults(RawFaultData, Numberxpixels,Numberypixels, sfcurvelist).astype(np.float32)
pix_faults = np.reshape(BasicInputStaticProps[:,0],(40,60)).astype(int)
annotate_faults_ndarray(pix_faults,figsize=(24, 16))
sfcurvelist2 = []
for yloc in range(0, Numberypixels):
for xloc in range(0, Numberxpixels):
pixellocation = yloc*Numberxpixels + xloc
[x,y] = sfcurvelist[pixellocation]
sfcurvelist2.append([x,39-y])
BasicInputStaticProps[:,1] = remapfaults(RawFaultData, Numberxpixels,Numberypixels, sfcurvelist2).astype(np.float32)
sfcurvelist3 = []
for yloc in range(0, Numberypixels):
for xloc in range(0, Numberxpixels):
pixellocation = yloc*Numberxpixels + xloc
[x,y] = sfcurvelist[pixellocation]
sfcurvelist3.append([59-x,y])
BasicInputStaticProps[:,2] = remapfaults(RawFaultData, Numberxpixels,Numberypixels, sfcurvelist3).astype(np.float32)
sfcurvelist4 = []
for yloc in range(0, Numberypixels):
for xloc in range(0, Numberxpixels):
pixellocation = yloc*Numberxpixels + xloc
[x,y] = sfcurvelist[pixellocation]
sfcurvelist4.append([59-x,39-y])
BasicInputStaticProps[:,3] = remapfaults(RawFaultData, Numberxpixels,Numberypixels, sfcurvelist4).astype(np.float32)
NpropperTimeDynamicCalculated = 11
NpropperTimeDynamic = NpropperTimeDynamicInput + NpropperTimeDynamicCalculated
NpropperTimeStatic = NumFaultLabels
# NumpredbasicperTime = NpropperTimeDynamic
NumpredbasicperTime = 1 # Can be 1 upto NpropperTimeDynamic
NumpredFuturedperTime = NumpredbasicperTime
# Setup Transformed Data
MagnitudeMethodTransform = 1
TransformName = 'E^0.25'
NpropperTime = NpropperTimeStatic + NpropperTimeDynamic
InputPropertyNames = [' '] * NpropperTime
DynamicNames = ['Magnitude Now',
'Depth Now',
'Multiplicity Now',
'Mult >3.29 Now',
'Mag 2/3 Month Back',
'Mag 1.5 Month Back',
'Mag 3 Months Back',
'Mag 6 Months Back',
'Mag Year Back',
TransformName + ' Now',
TransformName+' 2/3 Month Back',
TransformName+' 1.5 Month Back',
TransformName+' 3 Months Back',
TransformName+' 6 Months Back',
TransformName+' Year Back']
if Dailyunit == 14:
DynamicNames = ['Magnitude 2 weeks Now',
'Depth 2 weeks Now',
'Multiplicity 2 weeks Now',
'Mult >3.29 2 weeks Now',
'Mag 4 Weeks Back',
'Mag 2 Months Back',
'Mag 3 Months Back',
'Mag 6 Months Back',
'Mag Year Back',
TransformName+ ' 2 weeks Back',
TransformName+' 4 weeks Back',
TransformName+' 2 Months Back',
TransformName+' 3 Months Back',
TransformName+' 6 Months Back',
TransformName+' Year Back']
Property_is_Intensive = np.full(NpropperTime, True, dtype = bool)
for iprop in range(0, NpropperTimeStatic):
InputPropertyNames[iprop] = 'Fault ' +str(iprop)
for iprop in range(0, NpropperTimeDynamic):
InputPropertyNames[iprop+NpropperTimeStatic] = DynamicNames[iprop]
Num_Extensive = 0
ScaleProperties = True
GenerateFutures = False
GenerateSequences = True
PredictionsfromInputs = True
ConvertDynamicPredictedQuantity = False
AddSpecialstoSummedplots = True
UseRealDatesonplots = True
EarthquakeImagePlots = False
UseFutures = False
PopulationNorm = False
OriginalNloc = Nloc
MapLocation = False
# Add summed magnitudes as properties to use in prediction and Calculated Properties for some
# Calculated Properties are sums starting at given time and are set to NaN if necessary
NumTimeSeriesCalculatedBasic = 9
NumTimeSeriesCalculated = 2*NumTimeSeriesCalculatedBasic + 1
NamespredCalculated = ['Mag 2/3 Month Ahead',
'Mag 1.5 Month Ahead',
'Mag 3 Months Ahead',
'Mag 6 Months Ahead',
'Mag Year Ahead Ahead',
'Mag 2 Years Ahead',
'Mag 4 years Ahead',
'Mag Skip 1, Year ahead',
'Mag 2 years 2 ahead',
TransformName+' Daily Now',
TransformName+' 2/3 Month Ahead',
TransformName+' 1.5 Month Ahead',
TransformName+' 3 Months Ahead',
TransformName+' 6 Months Ahead',
TransformName+' Year Ahead',
TransformName+' 2 Years Ahead',
TransformName+' 4 years Ahead',
TransformName+' Skip 1, Year ahead',
TransformName+' 2 years 2 ahead']
Unitjumps = [ 23, 46, 92, 183, 365, 730, 1460, 365, 730]
Unitdelays = [ 0, 0, 0, 0, 0, 0, 0, 365, 730]
Plottingdelay = 1460
if Dailyunit == 14:
NumTimeSeriesCalculatedBasic = 9
NumTimeSeriesCalculated = 2*NumTimeSeriesCalculatedBasic + 1
NamespredCalculated = ['Mag 4 Weeks Ahead',
'Mag 2 Month Ahead',
'Mag 3 Months Ahead',
'Mag 6 Months Ahead',
'Mag Year Ahead',
'Mag 2 Years Ahead',
'Mag 4 years Ahead',
'Mag Skip 1, Year ahead',
'Mag 2 years 2 ahead',
TransformName+' 2 Weeks Now',
TransformName+' 4 Weeks Ahead',
TransformName+' 2 Months Ahead',
TransformName+' 3 Months Ahead',
TransformName+' 6 Months Ahead',
TransformName+' Year Ahead',
TransformName+' 2 Years Ahead',
TransformName+' 4 years Ahead',
TransformName+' Skip 1, Year ahead',
TransformName+' 2 years 2 ahead']
Unitjumps = [ 2, 4, 7, 13, 26, 52, 104, 26, 52]
Unitdelays = [ 0, 0, 0, 0, 0, 0, 0, 26, 52]
Plottingdelay = 104
NumpredbasicperTime += NumTimeSeriesCalculated
CalculatedTimeSeries = np.empty([Num_Time,Nloc,NumTimeSeriesCalculated],dtype = np.float32)
for icalc in range (0, NumTimeSeriesCalculatedBasic):
newicalc = icalc+1+NumTimeSeriesCalculatedBasic
for itime in range(0,Num_Time):
MagnitudeMethod = 0
CalculatedTimeSeries[itime,:,icalc] = AggregateEarthquakes(itime,Unitdelays[icalc],Unitjumps[icalc], Nloc,
BasicInputTimeSeries[:,:,0], 0)
MagnitudeMethod = MagnitudeMethodTransform
CalculatedTimeSeries[itime,:,newicalc] = TransformMagnitude(CalculatedTimeSeries[itime,:,icalc])
MagnitudeMethod = 0
current_time = timenow()
print(startbold + startred + 'Earthquake ' + str(icalc) + ' ' + NamespredCalculated[icalc] + ' ' + current_time + ' ' +RunName + resetfonts)
print(startbold + startred + 'Earthquake ' + str(newicalc) + ' ' + NamespredCalculated[newicalc] + ' ' + current_time + ' ' +RunName + resetfonts)
MagnitudeMethod = MagnitudeMethodTransform
CalculatedTimeSeries[:,:,NumTimeSeriesCalculatedBasic] = TransformMagnitude(BasicInputTimeSeries[:,:,0])
MagnitudeMethod = 0
print(startbold + startred + 'Earthquake ' + str(NumTimeSeriesCalculatedBasic) + ' ' + NamespredCalculated[NumTimeSeriesCalculatedBasic] + ' ' + current_time + ' ' +RunName + resetfonts)
for iprop in range(0,NumTimeSeriesCalculated):
InputPropertyNames.append(NamespredCalculated[iprop])
# + [markdown] id="4w6y73vmEleC" pycharm={"name": "#%% md\n"}
# ### Earthquake Eigensystems
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="ogCpNPUMEtiK" outputId="27c30056-ccb2-4233-bcc1-d24016b7b302" pycharm={"name": "#%%\n"}
if UseEarthquakeEigenSystems:
version = sc.version.version
print(f'SciPy version {version}')
#x = np.array([[1,2.0],[2.0,0]])
#w, v = solver.eigh(x, driver='evx')
#print(w)
#print(v)
# + [markdown] id="4KwljLkzTikB" pycharm={"name": "#%% md\n"}
# ### Multiplicity Data
# + colab={"base_uri": "https://localhost:8080/", "height": 1826} id="z86OVQYxTqwp" outputId="ccfee076-00ed-40c0-d56c-f55d44bf9199" pycharm={"name": "#%%\n"}
def histogrammultiplicity(Type, numbins, Data):
hitcounts = np.zeros(Nloc, dtype=int)
rawcounts = np.zeros(Nloc, dtype=int)
for iloc in range(0,Nloc):
rawcounts[iloc] = int(0.1+Data[:,iloc].sum(0))
hitcounts[iloc] = int(min(numbins, rawcounts[iloc]))
matplotlib.rcParams.update(matplotlib.rcParamsDefault)
plt.rcParams.update({'font.size': 9})
plt.rcParams["figure.figsize"] = [8,6]
plt.hist(hitcounts, numbins, facecolor='b', alpha=0.75, log=True)
plt.title('\n'.join(wrap(RunComment + ' ' + RunName + ' ' + Type + ' Earthquake Count per location ',70)))
plt.xlabel('Hit Counts')
plt.ylabel('Occurrences')
plt.grid(True)
plt.show()
return rawcounts
def threebythree(pixellocation,numxlocations,numylocations):
indices = np.empty([3,3], dtype=int)
y = int(0.1 + pixellocation/numxlocations)
x = pixellocation - y*numxlocations
bottomx = max(0,x-1)
bottomx = min(bottomx,numxlocations-3)
bottomy = max(0,y-1)
bottomy = min(bottomy,numylocations-3)
for ix in range(0,3):
for iy in range(0,3):
x= bottomx+ix
y= bottomy+iy
pixellocation = y*numxlocations + x
indices[ix,iy] = pixellocation
return indices
if Earthquake:
MappedLocations = np.arange(0,Nloc, dtype=int)
LookupLocations = np.arange(0,Nloc, dtype=int)
MappedNloc = Nloc
histogrammultiplicity('Basic', 100, BasicInputTimeSeries[:,:,2])
nbins = 10
if read1950:
nbins= 50
rawcounts1 = histogrammultiplicity('Rundle > 3.29', nbins, BasicInputTimeSeries[:,:,3])
TempTimeSeries = np.zeros([Num_Time,Nloc],dtype = np.float32)
for iloc in range (0,Nloc):
indices = threebythree(iloc,60,40)
for itime in range(0,Num_Time):
sum3by3 = 0.0
for ix in range(0,3):
for iy in range(0,3):
pixellocation = indices[ix,iy]
sum3by3 += BasicInputTimeSeries[itime,pixellocation,3]
TempTimeSeries[itime,iloc] = sum3by3
nbins =40
if read1950:
nbins= 150
rawcounts2 = histogrammultiplicity('3x3 Rundle > 3.29', nbins, TempTimeSeries)
#
# Define "Interesting Locations"
if read1950:
singleloccut = 25
groupedloccut = 110
singleloccut = 7.1
groupedloccut = 34.1
# groupedloccut = 1000000000
else:
singleloccut = 5.1
groupedloccut = 24.9
MappedLocations.fill(-1)
MappedNloc = 0
ct1 = 0
ct2 = 0
for iloc in range (0,Nloc):
if rawcounts1[iloc] >= singleloccut:
ct1 += 1
if rawcounts2[iloc] >= groupedloccut:
ct2 += 1
if rawcounts1[iloc] < singleloccut and rawcounts2[iloc] < groupedloccut:
continue
MappedLocations[iloc] = MappedNloc
MappedNloc += 1
LookupLocations = None
LookupLocations = np.empty(MappedNloc, dtype=int)
for iloc in range (0,Nloc):
jloc = MappedLocations[iloc]
if jloc >= 0:
LookupLocations[jloc] = iloc
TempTimeSeries = None
print('Total ' + str(MappedNloc) +
' Single location multiplicity cut ' + str(singleloccut) +
' ' + str(ct1) + ' 3x3 ' + str(groupedloccut) + ' ' + str(ct2))
if UseEarthquakeEigenSystems:
if Eigenvectors > 0:
UseTopEigenTotal = 16
UseTopEigenLocal = 0
if Eigenvectors > 1:
UseTopEigenLocal = 4
Num_EigenProperties = UseTopEigenTotal + UseTopEigenLocal
EigenTimeSeries = np.empty([Num_Time,MappedNloc],dtype = np.float32)
PsiTimeSeries = np.empty([Num_Time,MappedNloc],dtype = np.float32)
FiTimeSeries = np.empty([Num_Time,MappedNloc],dtype = np.float32)
EigenTimeSeries[:,:] = BasicInputTimeSeries[:,LookupLocations,3]
StoreEigenvectors = np.zeros([Num_Time,MappedNloc,MappedNloc],dtype = np.float32)
StoreEigencorrels = np.zeros([Num_Time,MappedNloc,MappedNloc],dtype = np.float32)
StoreNormingfactor = np.zeros([Num_Time],dtype = np.float32)
StoreNormingfactor1 = np.zeros([Num_Time],dtype = np.float32)
StoreNormingfactor2 = np.zeros([Num_Time],dtype = np.float32)
current_time = timenow()
print(startbold + startred + 'Start Eigen Earthquake '
+ current_time + ' ' +RunName + resetfonts)
for itime in range (0,Num_Time):
imax = itime
imin = max(0, imax-25)
Result = np.zeros(MappedNloc, dtype = np.float64)
Result = AggregateEarthquakes(imin,0,imax-imin+1, MappedNloc, EigenTimeSeries[:,:], 2)
PsiTimeSeries[itime,:] = Result
FiTimeSeries[itime,:] = EigenTimeSeries[itime,:]
current_time = timenow()
print(startbold + startred + 'End Eigen Earthquake 1 '
+ current_time + ' ' +RunName + resetfonts)
Eigenvals = np.zeros([Num_Time,MappedNloc], dtype = np.float32)
Chi1 = np.zeros(Num_Time, dtype = np.float32)
Chi2 = np.zeros(Num_Time, dtype = np.float32)
Sumai = np.zeros(Num_Time, dtype = np.float32)
Bestindex = np.zeros(Num_Time, dtype = int)
Numbereigs = np.zeros(Num_Time, dtype = int)
Besttrailingindex = np.zeros(Num_Time, dtype = int)
Eig0coeff = np.zeros(Num_Time, dtype = np.float32)
meanmethod = 0
if meanmethod == 1:
Meanovertime = np.empty(MappedNloc, dtype = np.float32)
sigmaovertime = np.empty(MappedNloc, dtype = np.float32)
Meanovertime = FiTimeSeries.mean(axis=0)
Meanovertime = Meanovertime.reshape(1,MappedNloc)
sigmaovertime = FiTimeSeries.std(axis=0)
sigmaovertime = sigmaovertime.reshape(1,MappedNloc)
countbad = 0
OldActualNumberofLocationsUsed = -1
for itime in range (25,Num_Time):
LocationCounts = FiTimeSeries[0:itime,:].sum(axis=0)
NumLocsToday = np.count_nonzero(LocationCounts)
Nonzeromapping = np.zeros(NumLocsToday, dtype = int)
#gregor
# Nonzeromapping = np.zeros(NumLocsToday, dtype = int)
ActualNumberofLocationsUsed = 0
for ipos in range (0,MappedNloc):
if LocationCounts[ipos] == 0:
continue
Nonzeromapping[ActualNumberofLocationsUsed] = ipos
ActualNumberofLocationsUsed +=1
if ActualNumberofLocationsUsed <= 1:
print(str(itime) + ' Abandoned ' + str(ActualNumberofLocationsUsed))
continue
FiHatTimeSeries = np.empty([itime+1,ActualNumberofLocationsUsed], dtype = np.float32)
if meanmethod == 1:
FiHatTimeSeries[:,:] = np.divide(np.subtract(FiTimeSeries[0:(itime+1),Nonzeromapping],Meanovertime[0,Nonzeromapping]),
sigmaovertime[0,Nonzeromapping])
else:
FiHatTimeSeries[:,:] = FiTimeSeries[0:(itime+1),Nonzeromapping]
# FiHatTimeSeries[:,:] = PsiTimeSeries[0:(itime+1),Nonzeromapping]
CorrelationMatrix = np.corrcoef(FiHatTimeSeries, rowvar =False)
bad = np.count_nonzero(np.isnan(CorrelationMatrix))
if bad > 0:
countbad += 1
continue
evalues, evectors = solver.eigh(CorrelationMatrix)
Newevector = evectors[:,ActualNumberofLocationsUsed-1]
Newevalue = evalues[ActualNumberofLocationsUsed-1]
debug = False
if debug:
if OldActualNumberofLocationsUsed == ActualNumberofLocationsUsed:
Mapdiff = np.where(np.not_equal(OldNonzeromapping,Nonzeromapping),1,0.).sum()
if Mapdiff > 0:
print(str(itime) + ' Change in mapping ' + str(ActualNumberofLocationsUsed) + ' Change ' + str(Mapdiff))
else:
Corrdiff = np.absolute(np.subtract(OldCorrelationMatrix,CorrelationMatrix)).sum()
Corrorg = np.absolute(CorrelationMatrix).sum()
yummy = CorrelationMatrix.dot(Oldevector)
vTMv = yummy.dot(Oldevector)
Doubleyummy = CorrelationMatrix.dot(Newevector)
newvTMv = Doubleyummy.dot(Newevector)
print(str(itime) + ' Change in correlation ' + str(ActualNumberofLocationsUsed) + ' Change '
+ str(Corrdiff) + ' original ' + str(Corrorg) + ' eval ' + str(Oldevalue) + ' new '
+ str(Newevalue) + ' vTMv ' + str(vTMv) + ' New ' + str(newvTMv))
else:
print(str(itime) + ' Change in size ' + str(OldActualNumberofLocationsUsed) + ' ' +
str(ActualNumberofLocationsUsed))
OldActualNumberofLocationsUsed = ActualNumberofLocationsUsed
OldNonzeromapping = Nonzeromapping
OldCorrelationMatrix = CorrelationMatrix
Oldevector = Newevector
Oldevalue = Newevalue
normcoeff = 100.0/evalues.sum()
evalues = np.multiply(evalues, normcoeff)
Numbereigs[itime] = ActualNumberofLocationsUsed
for ieig in range(0,ActualNumberofLocationsUsed):
Eigenvals[itime, ieig] = evalues[ActualNumberofLocationsUsed-ieig-1]
chival = 0.0
sumaieig = 0.0
Checkvector = np.zeros(ActualNumberofLocationsUsed,dtype = np.float32)
largesteigcoeff = -1.0
largestindex = -1
Keepaisquared = np.zeros(ActualNumberofLocationsUsed, dtype=np.float32)
for ieig in range(0,ActualNumberofLocationsUsed):
aieig = 0.0
backwards = ActualNumberofLocationsUsed-ieig-1
for vectorindex in range(0,ActualNumberofLocationsUsed):
StoreEigenvectors[itime,backwards,Nonzeromapping[vectorindex]] = evectors[vectorindex,ieig]
aieig += evectors[vectorindex,ieig]*PsiTimeSeries[itime,Nonzeromapping[vectorindex]]
for vectorindex in range(0,ActualNumberofLocationsUsed):
Checkvector[vectorindex] += aieig*evectors[vectorindex, ieig]
aieig *= aieig
chival += aieig*evalues[ieig]
sumaieig += aieig
Keepaisquared[backwards] = aieig
for ieig in range(0,ActualNumberofLocationsUsed):
backwards = ActualNumberofLocationsUsed-ieig-1
aieig = Keepaisquared[backwards]
aieig = aieig/sumaieig
if backwards == 0:
Eig0coeff[itime] = aieig
test = evalues[ieig]*aieig
if test > largesteigcoeff:
largesteigcoeff = test
largestindex = backwards
Bestindex[itime] = largestindex
discrep = 0.0
for vectorindex in range(0,ActualNumberofLocationsUsed):
discrep += pow(Checkvector[vectorindex] - PsiTimeSeries[itime,Nonzeromapping[vectorindex]], 2)
if discrep > 0.01:
print('Eigendecomposition Failure ' + str(itime) + ' ' + str(discrep))
Chi1[itime] = chival
Chi2[itime] = chival/sumaieig
Sumai[itime] = sumaieig
largesteigcoeff = -1.0
largestindex = -1
sumaieig = 0.0
Trailingtimeindex = itime-3
if itime > 40:
Trailinglimit = Numbereigs[Trailingtimeindex]
KeepTrailingaisquared = np.zeros(Trailinglimit, dtype=np.float32)
for ieig in range(0,Trailinglimit):
aieig = 0.0
for vectorindex in range(0,MappedNloc):
# aieig += StoreEigenvectors[Trailingtimeindex,ieig,vectorindex]*PsiTimeSeries[itime,vectorindex]
aieig += StoreEigenvectors[Trailingtimeindex,ieig,vectorindex]*StoreEigenvectors[itime,
Bestindex[itime],vectorindex]
aieig *= aieig
sumaieig += aieig
KeepTrailingaisquared[ieig] = aieig
for ieig in range(0,Trailinglimit):
aieig = KeepTrailingaisquared[ieig]
aieig = aieig/sumaieig
test = Eigenvals[Trailingtimeindex, ieig]*aieig
if test > largesteigcoeff:
largesteigcoeff = test
largestindex = ieig
Besttrailingindex[itime] = largestindex
if itime >40: # Calculate eigenvector tracking
Leader = StoreEigenvectors[itime,:,:]
Trailer = StoreEigenvectors[itime-3,:,:]
StoreEigencorrels[itime,:,:] = np.tensordot(Leader, Trailer, (1, (1)))
StrippedDown = StoreEigencorrels[itime,Bestindex[itime],:]
Normingfactor = np.multiply(StrippedDown,StrippedDown).sum()
Normingfactor1 = np.multiply(StrippedDown[0:8],StrippedDown[0:8]).sum()
Normingfactor2 = np.multiply(StrippedDown[0:30],StrippedDown[0:30]).sum()
StoreNormingfactor[itime] = Normingfactor
StoreNormingfactor1[itime] = Normingfactor1
StoreNormingfactor2[itime] = Normingfactor2
averagesumai = Sumai.mean()
Chi1 = np.divide(Chi1,averagesumai)
print('Bad Correlation Matrices ' + str(countbad))
print(startbold + startred + 'End Eigen Earthquake 2 '
+ current_time + ' ' +RunName + resetfonts)
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="IG7_hFuDb0oM" outputId="95ea5c3e-d8eb-4fad-93cb-a105bb44150f" pycharm={"name": "#%%\n"}
def makeasmalldateplot(figure,ax, Dateaxis):
plt.rcParams.update({'font.size': 9})
months = mdates.MonthLocator(interval=2) # every month
datemin = np.datetime64(Dateaxis[0], 'M')
datemax = np.datetime64(Dateaxis[-1], 'M') + np.timedelta64(1, 'M')
ax.set_xlim(datemin, datemax)
months_fmt = mdates.DateFormatter('%y-%b')
locator = mdates.AutoDateLocator()
locator.intervald['MONTHLY'] = [2]
formatter = mdates.ConciseDateFormatter(locator)
# ax.xaxis.set_major_locator(locator)
# ax.xaxis.set_major_formatter(formatter)
ax.xaxis.set_major_locator(months)
ax.xaxis.set_major_formatter(months_fmt)
figure.autofmt_xdate()
return datemin, datemax
def plotquakeregions(HalfSize,xaxisdates, SetofPlots, Commontitle, ylabel, SetofColors, Startx, ncols):
numplotted = SetofPlots.shape[1]
totusedquakes = 0
for iquake in range(0,numberspecialeqs):
x_line_index = Specialindex[iquake]
if (x_line_index <= Startx) or (x_line_index >= Num_Time-1):
continue
if Specialuse[iquake]:
totusedquakes +=1
nrows = math.ceil(totusedquakes/ncols)
sortedquakes = np.argsort(Specialindex)
jplot = 0
kplot = -1
for jquake in range(0,numberspecialeqs):
iquake = sortedquakes[jquake]
if not Specialuse[iquake]:
continue
x_line_annotation = Specialdate[iquake]
x_line_index = Specialindex[iquake]
if (x_line_index <= Startx) or (x_line_index >= Num_Time-1):
continue
kplot +=1
if kplot == ncols:
SAVEFIG(plt, f'{APPLDIR}/Outputs/QRegions' + str(jplot) + f'{RunName}.png')
plt.show()
kplot = 0
jplot +=1
if kplot == 0:
plt.rcParams["figure.figsize"] = [16,6]
figure, axs = plt.subplots(nrows=1, ncols=ncols, squeeze=False)
beginplotindex = x_line_index - HalfSize
beginplotindex = max(beginplotindex, Startx)
endplotindex = x_line_index + HalfSize
endplotindex = min(endplotindex, Num_Time-1)
eachplt = axs[0,kplot]
ascii = ''
if Specialuse[iquake]:
ascii = np.datetime_as_string(Specialdate[iquake]) + ' ' + str(round(Specialmags[iquake],1)) + ' ' + Specialeqname[iquake]
eachplt.set_title(str(iquake) + ' ' + RunName + ' Best Eigenvalue (Black) Trailing (Red) \n' + ascii)
datemin, datemax = makeasmalldateplot(figure, eachplt, xaxisdates[beginplotindex:endplotindex+1])
for curves in range(0,numplotted):
eachplt.plot(xaxisdates[beginplotindex:endplotindex+1], SetofPlots[beginplotindex:endplotindex+1,curves],
'o', color=SetofColors[curves], markersize =1)
ymin, ymax = eachplt.get_ylim()
if ymax >= 79.9:
ymax = 82
eachplt.set_ylim(bottom=-1.0, top=max(ymax,20))
eachplt.set_ylabel(ylabel)
eachplt.set_xlabel('Time')
eachplt.grid(True)
eachplt.set_yscale("linear")
eachplt.axvline(x=x_line_annotation, linestyle='dashed', alpha=1.0, linewidth = 2.0, color='red')
for kquake in range(0,numberspecialeqs):
if not Specialuse[kquake]:
continue
if kquake == iquake:
continue
anotherx_line_index = Specialindex[kquake]
if (anotherx_line_index < beginplotindex) or (anotherx_line_index >= endplotindex):
continue
eachplt.axvline(x=Specialdate[kquake], linestyle='dashed', alpha=1.0, linewidth = 1.0, color='purple')
eachplt.tick_params('x', direction = 'in', length=15, width=2, which='major')
SAVEFIG(plt, f'{APPLDIR}/Outputs/QRegions' + str(jplot) + f'{RunName}.png')
plt.show()
EigenAnalysis = False
if Earthquake and EigenAnalysis:
UseTopEigenTotal = 40
FirstTopEigenTotal = 10
PLTlabels = []
for ieig in range(0,UseTopEigenTotal):
PLTlabels.append('Eig-' + str(ieig))
plt.rcParams["figure.figsize"] = [12,10]
figure, ax = plt.subplots()
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
plt.rcParams["figure.figsize"] = [12,10]
for ieig in range(0,FirstTopEigenTotal):
ax.plot(Dateaxis[26:],np.maximum(Eigenvals[26:, ieig],0.1))
def gregor_plot(RunName, scale="log"): # linear
ax.set_title(RunName + ' Multiplicity Eigenvalues')
ax.set_ylabel('Eigenvalue')
ax.set_xlabel('Time')
ax.set_yscale(scale)
ax.grid(True)
ax.legend(PLTlabels[0:FirstTopEigenTotal], loc='upper right')
Addfixedearthquakes(ax, datemin, datemax,ylogscale=True )
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
plt.show()
# gregor_plot(RunName,scale="log")
ax.set_title(RunName + ' Multiplicity Eigenvalues')
ax.set_ylabel('Eigenvalue')
ax.set_xlabel('Time')
ax.set_yscale("log")
ax.grid(True)
ax.legend(PLTlabels[0:FirstTopEigenTotal], loc='upper right')
Addfixedearthquakes(ax, datemin, datemax,ylogscale=True )
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
plt.show()
# end gregor plot
plt.rcParams["figure.figsize"] = [12,10]
figure, ax = plt.subplots()
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
plt.rcParams["figure.figsize"] = [12,10]
for ieig in range(FirstTopEigenTotal,UseTopEigenTotal):
ax.plot(Dateaxis[26:],np.maximum(Eigenvals[26:, ieig],0.1))
# gregor_plot(RunName,scale="linear")
ax.set_title(RunName + ' Multiplicity Eigenvalues')
ax.set_ylabel('Eigenvalue')
ax.set_xlabel('Time')
ax.set_yscale("linear")
ax.grid(True)
ax.legend(PLTlabels[FirstTopEigenTotal:], loc='upper right')
Addfixedearthquakes(ax, datemin, datemax,ylogscale=False )
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
plt.show()
# end gregor plot
ShowEigencorrels = False
if ShowEigencorrels:
for mastereig in range(0, UseTopEigenTotal):
figure, ax = plt.subplots()
plt.rcParams["figure.figsize"] = [12,8]
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
for ieig in range(0,UseTopEigenTotal):
alpha = 1.0
width = 3
if ieig == mastereig:
alpha=0.5
width = 1
ax.plot(Dateaxis[26:],np.power(StoreEigencorrels[26:,mastereig,ieig],2), alpha=alpha, linewidth = width)
ax.set_title(RunName + ' Eigenvalue ' + str(mastereig) + ' Current versus Past Total Correlation')
ax.set_ylabel('Norm')
ax.set_xlabel('Time')
ax.grid(True)
ax.legend(PLTlabels, loc='upper right')
Addfixedearthquakes(ax, datemin, datemax,ylogscale=False )
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
plt.show()
def gregor_plot_normfacor(title,normfactor=StoreNormingfactor):
figure, ax = plt.subplots()
plt.rcParams["figure.figsize"] = [12,8]
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
alpha = 1.0
width = 0.5
ax.plot(Dateaxis[26:],StoreNormingfactor[26:], alpha=alpha, linewidth = width)
ax.set_title(f'{RunName} Eigenvalue Full Norming Factor with Past')
ax.set_ylabel('Norming Factor')
ax.set_xlabel('Time')
ax.grid(True)
Addfixedearthquakes(ax, datemin, datemax,ylogscale=False )
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
plt.show()
# Gregor: creat functions for plots
# gregor_plot_normfactor(f"{RunName} Eigenvalue Full Norming Factor with Past", StoreNormingfactor)
figure, ax = plt.subplots()
plt.rcParams["figure.figsize"] = [12,8]
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
alpha = 1.0
width = 0.5
ax.plot(Dateaxis[26:],StoreNormingfactor[26:], alpha=alpha, linewidth = width)
ax.set_title(f'{RunName} Eigenvalue Full Norming Factor with Past')
ax.set_ylabel('Norming Factor')
ax.set_xlabel('Time')
ax.grid(True)
Addfixedearthquakes(ax, datemin, datemax,ylogscale=False )
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
plt.show()
# Gregor: creat functions for plots
# gregor_plot_normfactor(f"{RunName} Eigenvalue First 8 Norming Factor with Past", StoreNormingfactor1)
figure, ax = plt.subplots()
plt.rcParams["figure.figsize"] = [12,8]
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
alpha = 1.0
width = 0.5
ax.plot(Dateaxis[26:],StoreNormingfactor1[26:], alpha=alpha, linewidth = width)
ax.set_title(f"{RunName} Eigenvalue First 8 Norming Factor with Past")
ax.set_ylabel('Norming Factor')
ax.set_xlabel('Time')
ax.grid(True)
Addfixedearthquakes(ax, datemin, datemax,ylogscale=False )
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
plt.show()
# gregor_plot_normfactor(f"{RunName} Eigenvalue First 38 Norming Factor with Past", StoreNormingfactor2)
# Gregor: creat functions for plots
figure, ax = plt.subplots()
plt.rcParams["figure.figsize"] = [12,8]
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
alpha = 1.0
width = 0.5
ax.plot(Dateaxis[26:],StoreNormingfactor2[26:], alpha=alpha, linewidth = width)
ax.set_title(RunName + ' Eigenvalue First 30 Norming Factor with Past')
ax.set_ylabel('Norming Factor')
ax.set_xlabel('Time')
ax.grid(True)
Addfixedearthquakes(ax, datemin, datemax,ylogscale=False )
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
plt.show()
# Gregor: creat functions for plots
figure, ax = plt.subplots()
plt.rcParams["figure.figsize"] = [12,8]
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
plt.rcParams["figure.figsize"] = [12,8]
ax.plot(Dateaxis[26:],Chi1[26:])
ax.set_title(RunName + ' Correlations Normalized on average over time')
ax.set_ylabel('Chi1')
ax.set_xlabel('Time')
ax.grid(True)
Addfixedearthquakes(ax, datemin, datemax)
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
ax.set_yscale("linear")
plt.show()
figure, ax = plt.subplots()
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
plt.rcParams["figure.figsize"] = [12,8]
ax.plot(Dateaxis[26:],Chi2[26:])
ax.set_title(RunName + ' Correlations Normalized at each time')
ax.set_ylabel('Chi2')
ax.set_xlabel('Time')
ax.grid(True)
Addfixedearthquakes(ax, datemin, datemax)
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
ax.set_yscale("linear")
plt.show()
figure, ax = plt.subplots()
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
plt.rcParams["figure.figsize"] = [12,8]
norm = np.amax(Chi1[26:])
Maxeig = 80
# ax.plot(Dateaxis[26:],Chi1[26:]*Maxeig/norm)
ax.plot(Dateaxis[26:], 0.5 + np.minimum(Maxeig, Bestindex[26:]), 'o', color='black', markersize =1)
ax.plot(Dateaxis[26:], np.minimum(Maxeig, Besttrailingindex[26:]), 'o', color='red', markersize =1)
ax.set_title(RunName + ' Best Eigenvalue (Black) Trailing (Red)')
ax.set_ylabel('Eig#')
ax.set_xlabel('Time')
ax.grid(True)
Addfixedearthquakes(ax, datemin, datemax)
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
ax.set_yscale("linear")
plt.show()
SetofPlots = np.empty([len(Bestindex),2], dtype=np.float32)
SetofPlots[:,0] = 0.5 + np.minimum(Maxeig, Bestindex[:])
SetofPlots[:,1] = np.minimum(Maxeig, Besttrailingindex[:])
SetofColors = ['black', 'red']
plotquakeregions(25, Dateaxis, SetofPlots,
RunName + ' Best Eigenvalue (Black) Trailing (Red)', 'Eig#', SetofColors, 26,2)
plt.rcParams["figure.figsize"] = [12,8]
figure, ax = plt.subplots()
datemin, datemax = makeadateplot(figure, ax, Dateaxis[26:])
ax.plot(Dateaxis[26:], Eig0coeff[26:], 'o', color='black', markersize =2)
ymin, ymax = ax.get_ylim()
ax.plot(Dateaxis[26:], Chi1[26:]*ymax/norm)
ax.set_title(RunName + ' Fraction Largest Eigenvalue')
ax.set_ylabel('Eig 0')
ax.set_xlabel('Time')
ax.grid(True)
Addfixedearthquakes(ax, datemin, datemax)
ax.tick_params('x', direction = 'in', length=15, width=2, which='major')
ax.xaxis.set_minor_locator(mdates.YearLocator(1))
ax.tick_params('x', direction = 'in', length=10, width=1, which='minor')
ax.set_yscale("linear")
plt.show()
# + [markdown] id="rk4-HhAzdezY" pycharm={"name": "#%% md\n"}
# ### End of Earthquake. Reset Timing
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="5g9IO3Y3ddQR" outputId="1c114488-56af-41a5-f826-f6251e53cf40" pycharm={"name": "#%%\n"}
# Reset Start Date by a year so first entry has a 365 day sample ending at that day and so can be made an input as can all
# lower time intervals
# Do NOT include 2 year or 4 year in input stream
# So we reset start date by one year skipping first 364 daya except to calculate the first one year (and lower limit) observables
# Time indices go from 0 to NumberofTimeunits-1
# Sequence Indices go from Begin to Begin+Tseq-1 where Begin goes from 0 to NumberofTimeunits-1-Tseq
# So Num_Seq = Numberodays-Tseq and Begin has Num_Seq values
if Earthquake:
SkipTimeUnits = 364
if Dailyunit == 14:
SkipTimeUnits = 25
Num_Time_old = NumberofTimeunits
NumberofTimeunits = NumberofTimeunits - SkipTimeUnits
Num_Time = NumberofTimeunits
InitialDate = InitialDate + timedelta(days=SkipTimeUnits*Dailyunit)
FinalDate = InitialDate + timedelta(days=(NumberofTimeunits-1)*Dailyunit)
print('Skip ' +str(SkipTimeUnits) + ' New dates: ' + InitialDate.strftime("%d/%m/%Y") + ' To '
+ FinalDate.strftime("%d/%m/%Y")+ ' days ' + str(NumberofTimeunits*Dailyunit))
DynamicPropertyTimeSeries = np.empty([Num_Time,Nloc,NpropperTimeDynamic],dtype = np.float32)
CountNaN = np.zeros(NpropperTimeDynamic, dtype=int)
# Skewtime makes certain propert ENDS at given cell and is the cell itself if size = DailyUnit
SkewTime = [0] * NpropperTimeDynamicInput
if Dailyunit == 1:
SkewTime = SkewTime + [22,45,91,182,364,0,22,45,91,182,364]
if Dailyunit == 14:
SkewTime = SkewTime + [1, 3, 6, 12, 25,0,1, 3, 6, 12, 25]
i = 0
total = NumberofTimeunits * Nloc * NpropperTimeDynamic
for itime in range(0,NumberofTimeunits):
for iloc in range(0,Nloc):
for iprop in range(0,NpropperTimeDynamic):
i = i + 1
addtime = SkipTimeUnits - SkewTime[iprop]
if iprop < NpropperTimeDynamicInput:
# BUG HERE
if i % 1000 == 0:
print(itime+addtime,f"{i}/{total}", iloc,iprop)
localval = BasicInputTimeSeries[itime+addtime,iloc,iprop]
elif iprop < (NpropperTimeDynamic-5):
localval = CalculatedTimeSeries[itime+addtime,iloc,iprop-NpropperTimeDynamicInput]
else:
localval = CalculatedTimeSeries[itime+addtime,iloc,iprop-NpropperTimeDynamicInput+4]
if np.math.isnan(localval):
localval = NaN
CountNaN[iprop] +=1
DynamicPropertyTimeSeries[itime,iloc,iprop] = localval
print(startbold+startred+'Input NaN values ' + resetfonts)
# Add E^0.25 Input Quantities
MagnitudeMethod = MagnitudeMethodTransform
jprop = 9
for iprop in range(0,9):
line = ''
if iprop == 0 or iprop > 3:
DynamicPropertyTimeSeries[:,:,jprop] = TransformMagnitude(DynamicPropertyTimeSeries[:,:,iprop])
jprop += 1
line = ' New ' + str(jprop) + ' ' + InputPropertyNames[jprop+NpropperTimeStatic] + ' NaN ' + str(CountNaN[iprop])
print(str(iprop) + ' ' + InputPropertyNames[iprop+NpropperTimeStatic] + ' NaN ' + str(CountNaN[iprop]) + line)
NpropperTimeDynamic = jprop
MagnitudeMethod = 0
NewCalculatedTimeSeries = np.empty([Num_Time,Nloc,NumTimeSeriesCalculated],dtype = np.float32)
# NewCalculatedTimeSeries = CalculatedTimeSeries[SkipTimeUnits:Num_Time+SkipTimeUnits]
NewCalculatedTimeSeries = TransformMagnitude(CalculatedTimeSeries[SkipTimeUnits:Num_Time+SkipTimeUnits])
CalculatedTimeSeries = None
CalculatedTimeSeries = NewCalculatedTimeSeries
BasicInputTimeSeries = None
if GarbageCollect:
gc.collect()
MagnitudeMethod = 0
current_time = timenow()
print(startbold + startred + 'Earthquake Setup ' + current_time + ' ' +RunName + ' ' + RunComment + resetfonts)
# + [markdown] id="My9HJsTkbCRo" pycharm={"name": "#%% md\n"}
# ### Set Earthquake Execution Mode
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="kWmTr4vUbRcA" outputId="54c61c7f-13de-49a9-9c13-5a5354f46370" pycharm={"name": "#%%\n"}
if Earthquake:
SymbolicWindows = True
Tseq = 26
Tseq = config.Tseq #num_encoder_steps
if Dailyunit == 14:
GenerateFutures = True
UseFutures = True
# + [markdown] id="qLL834JvEjLd" pycharm={"name": "#%% md\n"}
# ### Plot Earthquake Images
# + colab={"base_uri": "https://localhost:8080/", "height": 9283} id="LfCR1vKwEpVw" outputId="a4ed54b7-88ac-4c9a-e787-a7d977b89b55" pycharm={"name": "#%%\n"}
# Tom: added local min and max to graphs not based on absolute values.
# added localmin and localmax values to the plotimages function and modified last line of code to add those values in.
def plotimages(Array,Titles,nrows,ncols,localmin,localmax):
usedcolormap = "YlGnBu"
plt.rcParams["figure.figsize"] = [16,6*nrows]
figure, axs = plt.subplots(nrows=nrows, ncols=ncols, squeeze=False)
iplot=0
images = []
norm = colors.Normalize(vmin=localmin, vmax=localmax)
for jplot in range(0,nrows):
for kplot in range (0,ncols):
eachplt = axs[jplot,kplot]
if MapLocation:
Plotit = np.zeros(OriginalNloc, dtype = np.float32)
for jloc in range (0,Nloc):
Plotit[LookupLocations[jloc]] = Array[iplot][jloc]
TwoDArray = np.reshape(Plotit,(40,60))
else:
TwoDArray = np.reshape(Array[iplot],(40,60))
extent = (-120,-114, 36,32)
images.append(eachplt.imshow(TwoDArray, cmap=usedcolormap, norm=norm,extent=extent))
eachplt.label_outer()
eachplt.set_title(Titles[iplot])
iplot +=1
figure.colorbar(images[0], ax=axs, orientation='vertical', fraction=.05)
plt.show()
if Earthquake:
# DynamicPropertyTimeSeries and CalculatedTimeSeries are dimensione by time 0 ...Num_Time-1
# DynamicPropertyTimeSeries holds values upto and including that time
# CalculatedTimeSeries holds values STARTING at that time
fullmin = np.nanmin(CalculatedTimeSeries)
fullmax = np.nanmax(CalculatedTimeSeries)
fullmin = min(fullmin,np.nanmin(DynamicPropertyTimeSeries[:,:,0]))
fullmax = max(fullmax,np.nanmax(DynamicPropertyTimeSeries[:,:,0]))
print('Full Magnitude Ranges ' + str(fullmin) + ' ' + str(fullmax))
Num_Seq = NumberofTimeunits-Tseq
dayindexmax = Num_Seq-Plottingdelay
Numdates = 4
denom = 1.0/np.float64(Numdates-1)
for plotdays in range(0,Numdates):
dayindexvalue = math.floor(0.1 + (plotdays*dayindexmax)*denom)
if dayindexvalue < 0:
dayindexvalue = 0
if dayindexvalue > dayindexmax:
dayindexvalue = dayindexmax
dayindexvalue += Tseq
InputImages =[]
InputTitles =[]
InputImages.append(DynamicPropertyTimeSeries[dayindexvalue,:,0])
ActualDate = InitialDate + timedelta(days=dayindexvalue)
localmax1 = DynamicPropertyTimeSeries[dayindexvalue,:,0].max()
localmin1 = DynamicPropertyTimeSeries[dayindexvalue,:,0].min()
InputTitles.append('Day ' +str(dayindexvalue) + ' ' + ActualDate.strftime("%d/%m/%Y") + ' One day max/min '
+ str(round(localmax1,3)) + ' ' + str(round(localmin1,3)))
for localplot in range(0,NumTimeSeriesCalculated):
localmax1 = CalculatedTimeSeries[dayindexvalue,:,0].max()
localmin1 = CalculatedTimeSeries[dayindexvalue,:,0].min()
InputImages.append(CalculatedTimeSeries[dayindexvalue,:,localplot])
InputTitles.append('Day ' +str(dayindexvalue) + ' ' + ActualDate.strftime("%d/%m/%Y") + NamespredCalculated[localplot] + ' max/min '
+ str(round(localmax1,3)) + ' ' + str(round(localmin1,3)))
print(f'Local Magnitude Ranges {round(localmin1,3)} - {round(localmax1,3)}')
plotimages(InputImages,InputTitles,5,2, round(localmin1,3), round(localmax1,3))
# + [markdown] id="7EdsHkx7jLJX" pycharm={"name": "#%% md\n"}
# ### Read and setup NIH Covariates August 2020 and January, April 2021 Data
#
# new collection of time dependent covariates (even if constant).
#
# cases and deaths and location property from previous data
#
# + [markdown] id="awLjz1adEXr3" pycharm={"name": "#%% md\n"}
# ## Process Input Data in various ways
#
#
# + [markdown] id="H1fLYj-KBAjF" pycharm={"name": "#%% md\n"}
# ### Set TFT Mode
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="hH6D2TmcBE4u" outputId="721ae2e2-1b7b-4613-a59f-e18c414445e0" pycharm={"name": "#%%\n"}
UseTFTModel = True
# + [markdown] id="wrugyhFU66md" pycharm={"name": "#%% md\n"}
# ### Convert Cumulative to Daily
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="ipjkf86A6imL" outputId="edaa6cd3-4460-4654-9744-166c8078df4a" pycharm={"name": "#%%\n"}
# Gregor: DELETE
# Convert cumulative to Daily.
# Replace negative daily values by zero
# remove daily to sqrt(daily) and Then normalize maximum to 1
if ConvertDynamicPredictedQuantity:
NewBasicInputTimeSeries = np.empty_like(BasicInputTimeSeries, dtype=np.float32)
Zeroversion = np.zeros_like(BasicInputTimeSeries, dtype=np.float32)
Rolleddata = np.roll(BasicInputTimeSeries, 1, axis=0)
Rolleddata[0,:,:] = Zeroversion[0,:,:]
NewBasicInputTimeSeries = np.maximum(np.subtract(BasicInputTimeSeries,Rolleddata),Zeroversion)
originalnumber = np.sum(BasicInputTimeSeries[NumberofTimeunits-1,:,:],axis=0)
newnumber = np.sum(NewBasicInputTimeSeries,axis=(0,1))
print('Original summed counts ' + str(originalnumber) + ' become ' + str(newnumber)+ ' Cases, Deaths')
BasicInputTimeSeries = NewBasicInputTimeSeries
# + [markdown] id="aSDyT65ly4Q-" pycharm={"name": "#%% md\n"}
# ### Normalize All Static and Dynamic Properties
#
# for Static Properties BasicInputStaticProps[Nloc,NpropperTimeStatic] converts to NormedInputStaticProps[Nloc,NpropperTimeStatic]
# + colab={"base_uri": "https://localhost:8080/", "height": 832} id="NGbBzf47zv1m" outputId="d9860f1a-063a-4356-b22b-6aafb6ec4807" pycharm={"name": "#%%\n"}
# Gregor: DELETE some portions of this to be reviewed
def SetTakeroot(x,n):
if np.isnan(x):
return NaN
if n == 3:
return np.cbrt(x)
elif n == 2:
if x <= 0.0:
return 0.0
return np.sqrt(x)
return x
def DynamicPropertyScaling(InputTimeSeries):
Results = np.full(7, 0.0,dtype=np.float32)
Results[1] = np.nanmax(InputTimeSeries, axis = (0,1))
Results[0] = np.nanmin(InputTimeSeries, axis = (0,1))
Results[3] = np.nanmean(InputTimeSeries, axis = (0,1))
Results[4] = np.nanstd(InputTimeSeries, axis = (0,1))
Results[2] = np.reciprocal(np.subtract(Results[1],Results[0]))
Results[5] = np.multiply(Results[2],np.subtract(Results[3],Results[0]))
Results[6] = np.multiply(Results[2],Results[4])
return Results
NpropperTimeMAX = NpropperTime + NumTimeSeriesCalculated
print(NpropperTimeStatic,NpropperTime,NumTimeSeriesCalculated, NpropperTimeMAX)
if ScaleProperties:
QuantityTakeroot = np.full(NpropperTimeMAX,1,dtype=int)
# Scale data by roots if requested
for iprop in range(0, NpropperTimeMAX):
if QuantityTakeroot[iprop] >= 2:
if iprop < NpropperTimeStatic:
for iloc in range(0,Nloc):
BasicInputStaticProps[iloc,iprop] = SetTakeroot(BasicInputStaticProps[iloc,iprop],QuantityTakeroot[iprop])
elif iprop < NpropperTime:
for itime in range(0,NumberofTimeunits):
for iloc in range(0,Nloc):
DynamicPropertyTimeSeries[itime,iloc,iprop-NpropperTimeStatic] = SetTakeroot(
DynamicPropertyTimeSeries[itime,iloc,iprop-NpropperTimeStatic],QuantityTakeroot[iprop])
else:
for itime in range(0,NumberofTimeunits):
for iloc in range(0,Nloc):
CalculatedTimeSeries[itime,iloc,iprop-NpropperTime] =SetTakeroot(
CalculatedTimeSeries[itime,iloc,iprop-NpropperTime],QuantityTakeroot[iprop])
QuantityStatisticsNames = ['Min','Max','Norm','Mean','Std','Normed Mean','Normed Std']
QuantityStatistics = np.zeros([NpropperTimeMAX,7], dtype=np.float32)
if NpropperTimeStatic > 0:
print(BasicInputStaticProps.shape)
max_value = np.amax(BasicInputStaticProps, axis = 0)
min_value = np.amin(BasicInputStaticProps, axis = 0)
mean_value = np.mean(BasicInputStaticProps, axis = 0)
std_value = np.std(BasicInputStaticProps, axis = 0)
normval = np.reciprocal(np.subtract(max_value,min_value))
normed_mean = np.multiply(normval,np.subtract(mean_value,min_value))
normed_std = np.multiply(normval,std_value)
QuantityStatistics[0:NpropperTimeStatic,0] = min_value
QuantityStatistics[0:NpropperTimeStatic,1] = max_value
QuantityStatistics[0:NpropperTimeStatic,2] = normval
QuantityStatistics[0:NpropperTimeStatic,3] = mean_value
QuantityStatistics[0:NpropperTimeStatic,4] = std_value
QuantityStatistics[0:NpropperTimeStatic,5] = normed_mean
QuantityStatistics[0:NpropperTimeStatic,6] = normed_std
NormedInputStaticProps =np.empty_like(BasicInputStaticProps)
for iloc in range(0,Nloc):
NormedInputStaticProps[iloc,:] = np.multiply((BasicInputStaticProps[iloc,:] - min_value[:]),normval[:])
if (NpropperTimeDynamic > 0) or (NumTimeSeriesCalculated>0):
for iprop in range(NpropperTimeStatic,NpropperTimeStatic+NpropperTimeDynamic):
QuantityStatistics[iprop,:] = DynamicPropertyScaling(DynamicPropertyTimeSeries[:,:,iprop-NpropperTimeStatic])
for iprop in range(0,NumTimeSeriesCalculated):
QuantityStatistics[iprop+NpropperTime,:] = DynamicPropertyScaling(CalculatedTimeSeries[:,:,iprop])
NormedDynamicPropertyTimeSeries = np.empty_like(DynamicPropertyTimeSeries)
for iprop in range(NpropperTimeStatic,NpropperTimeStatic+NpropperTimeDynamic):
NormedDynamicPropertyTimeSeries[:,:,iprop - NpropperTimeStatic] = np.multiply((DynamicPropertyTimeSeries[:,:,iprop - NpropperTimeStatic]
- QuantityStatistics[iprop,0]),QuantityStatistics[iprop,2])
if NumTimeSeriesCalculated > 0:
NormedCalculatedTimeSeries = np.empty_like(CalculatedTimeSeries)
for iprop in range(NpropperTime,NpropperTimeMAX):
NormedCalculatedTimeSeries[:,:,iprop - NpropperTime] = np.multiply((CalculatedTimeSeries[:,:,iprop - NpropperTime]
- QuantityStatistics[iprop,0]),QuantityStatistics[iprop,2])
CalculatedTimeSeries = None
BasicInputStaticProps = None
DynamicPropertyTimeSeries = None
print(startbold + "Properties scaled" +resetfonts)
line = 'Name '
for propval in range (0,7):
line += QuantityStatisticsNames[propval] + ' '
print('\n' + startbold +startpurple + line + resetfonts)
for iprop in range(0,NpropperTimeMAX):
if iprop == NpropperTimeStatic:
print('\n')
line = startbold + startpurple + str(iprop) + ' ' + InputPropertyNames[iprop] + resetfonts + ' Root ' + str(QuantityTakeroot[iprop])
for propval in range (0,7):
line += ' ' + str(round(QuantityStatistics[iprop,propval],3))
print(line)
# + [markdown] id="yW9bPWExf4YK" pycharm={"name": "#%% md\n"}
# ### Set up Futures
#
# -- currently at unit time level
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="1uwExtALgrsW" outputId="17ff430e-1c78-47b6-afd8-332ab1d32d54" pycharm={"name": "#%%\n"}
class Future:
def __init__(self, name, daystart = 0, days =[], wgt=1.0, classweight = 1.0):
self.name = name
self.days = np.array(days)
self.daystart = daystart
self.wgts = np.full_like(self.days,wgt,dtype=float)
self.size = len(self.days)
self.classweight = classweight
LengthFutures = 0
Unit = "Day"
if Earthquake:
Unit = "2wk"
if GenerateFutures:
Futures =[]
daylimit = 14
if Earthquake:
daylimit = 25
for ifuture in range(0,daylimit):
xx = Future(Unit + '+' + str(ifuture+2), days=[ifuture+2])
Futures.append(xx)
LengthFutures = len(Futures)
Futuresmaxday = 0
Futuresmaxweek = 0
for i in range(0,LengthFutures):
j = len(Futures[i].days)
if j == 1:
Futuresmaxday = max(Futuresmaxday, Futures[i].days[0])
else:
Futuresmaxweek = max(Futuresmaxweek, Futures[i].days[j-1])
Futures[i].daystart -= Dropearlydata
if Futures[i].daystart < 0: Futures[i].daystart = 0
if Earthquake:
Futures[i].daystart = 0
# + [markdown] id="Kdm4DDFL92NJ" pycharm={"name": "#%% md\n"}
# ### Set up mappings of locations
#
# In next cell, we map locations for BEFORE location etc added
#
# In cell after that we do same for sequences
# + colab={"base_uri": "https://localhost:8080/", "height": 687} id="JRZm-x13980a" outputId="ebb1d654-9690-44eb-c102-c3cdd26ef57f" pycharm={"name": "#%%\n"}
OriginalNloc = Nloc
if Earthquake:
MapLocation = True
MappedDynamicPropertyTimeSeries = np.empty([Num_Time,MappedNloc,NpropperTimeDynamic],dtype = np.float32)
MappedNormedInputStaticProps = np.empty([MappedNloc,NpropperTimeStatic],dtype = np.float32)
MappedCalculatedTimeSeries = np.empty([Num_Time,MappedNloc,NumTimeSeriesCalculated],dtype = np.float32)
print(LookupLocations)
MappedDynamicPropertyTimeSeries[:,:,:] = NormedDynamicPropertyTimeSeries[:,LookupLocations,:]
NormedDynamicPropertyTimeSeries = None
NormedDynamicPropertyTimeSeries = MappedDynamicPropertyTimeSeries
MappedCalculatedTimeSeries[:,:,:] = NormedCalculatedTimeSeries[:,LookupLocations,:]
NormedCalculatedTimeSeries = None
NormedCalculatedTimeSeries = MappedCalculatedTimeSeries
MappedNormedInputStaticProps[:,:] = NormedInputStaticProps[LookupLocations,:]
NormedInputStaticProps = None
NormedInputStaticProps = MappedNormedInputStaticProps
Nloc = MappedNloc
if GarbageCollect:
gc.collect()
print('Number of locations reduced to ' + str(Nloc))
else:
MappedLocations = np.arange(0,Nloc, dtype=int)
LookupLocations = np.arange(0,Nloc, dtype=int)
MappedNloc = Nloc
# + [markdown] id="MTUIpVT3vris" pycharm={"name": "#%% md\n"}
# ###Property and Prediction Data Structures
#
# Two important Lists Properties and Predictions that are related
#
# * Data stored in series is for properties, the calculated value occuring at or ending that day
# * For predictions, the data is the calculated value from that date or later.
#
# * We store data labelled by time so that
# * for inputs we use time 0 upto last value - 1 i.e. position [length of array - 1]
# * for outputs (predictions) with sequence Tseq, we use array locations [Tseq] to [length of array -1]
# * This implies Num_Seq = Num_Time - Tseq
#
#
# **Properties**
#
# Everything appears in Property list -- both input and output (predicted)
# DynamicPropertyTimeSeries holds input property time series where value is value at that time using data before this time for aggregations
# * NpropperTimeStatic is the number of static properties -- typically read in or calculated from input information
# * NpropperTimeDynamicInput is total number of input time series
# * NpropperTimeDynamicCalculated is total number of calculated dynamic quantities used in Time series analysis as input properties and/or output predictions
# * NpropperTimeDynamic = NpropperTimeDynamicInput + NpropperTimeDynamicCalculated ONLY includes input properties
# * NpropperTime = NpropperTimeStatic + NpropperTimeDynamic will not include futures and NOT include calculated predictions
# * InputPropertyNames is a list of size NpropperTime holding names
# * NpropperTimeMAX = NpropperTime + NumTimeSeriesCalculated has calculated predictions following input properties ignoring futures
# * QuantityStatistics has 7 statistics used in normalizing for NpropperTimeMAX properties
# * Normalization takes NpropperTimeStatic static features in BasicInputStaticProps and stores in NormedInputStaticProps
# * Normalization takes NpropperTimeDynamicInput dynamic features in BasicInputTimeSeries and stores in NormedInputTimeSeries
# * Normalization takes NpropperTimeDynamicCalculated dynamic features in DynamicPropertyTimeSeries and stores in NormedDynamicPropertyTimeSeries
#
# **Predictions**
#
# * NumpredbasicperTime can be 1 upto NpropperTimeDynamic and are part of dynamic input series. It includes input values that are to be predicted (these MUST be at start) plus NumTimeSeriesCalculated calculated series
# * NumpredFuturedperTime is <= NumpredbasicperTime and is the number of input dynamic series that are futured
# * NumTimeSeriesCalculated is number of calculated (not as futures) time series stored in CalculatedTimeSeries and names in NamespredCalculated
# * Typically NumpredbasicperTime = NumTimeSeriesCalculated + NumpredFuturedperTime (**Currently this is assumed**)
# * Normalization takes NumTimeSeriesCalculated calculated series in CalculatedTimeSeries and stores in NormedCalculatedTimeSeries
# * Predictions per Time are NpredperTime = NumpredbasicperTime + NumpredFuturedperTime*LengthFutures
# * Predictions per sequence Npredperseq = NpredperTime
#
#
# + [markdown] id="dGvEtAj5xHhR" pycharm={"name": "#%% md\n"}
# ### Set Requested Properties Predictions Encodings
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="lycrtgBHxQCq" outputId="f9028b06-230d-45ba-9da2-14dc4ed084db" pycharm={"name": "#%%\n"}
# FuturePred = -1 Means NO FUTURE >= 0 FUTURED
# BASIC EARTHQUAKE SET JUST LOG ENERGY AND MULTIPLICITY
# PARAMETER IMPORTANT
if Earthquake:
InputSource = ['Static','Static','Static','Static','Dynamic','Dynamic','Dynamic','Dynamic'
,'Dynamic','Dynamic','Dynamic','Dynamic','Dynamic']
InputSourceNumber = [0,1,2,3,0,1,2,3,4,5,6,7,8]
PredSource = ['Dynamic','Calc','Calc','Calc','Calc','Calc','Calc','Calc','Calc','Calc']
PredSourceNumber = [0,0,1,2,3,4,5,6,7,8]
FuturedPred = [-1]*len(PredSource)
# Earthquake Space-Time
PropTypes = ['Spatial', 'TopDown', 'TopDown','TopDown','TopDown','TopDown','BottomUp','BottomUp','BottomUp','BottomUp']
PropValues = [0, 0, 1, 2, 3,4, 8,16,32,64]
PredTypes = ['Spatial', 'TopDown', 'TopDown','TopDown','TopDown','TopDown','BottomUp','BottomUp','BottomUp','BottomUp']
PredValues = [0, 0, 1, 2, 3,4, 8,16,32,64]
if UseTFTModel:
InputSource = ['Static','Static','Static','Static','Dynamic','Dynamic','Dynamic','Dynamic'
,'Dynamic','Dynamic','Dynamic','Dynamic','Dynamic']
InputSourceNumber = [0,1,2,3,0,1,2,3,4,5,6,7,8]
PredSource = ['Dynamic','Dynamic']
PredSourceNumber = [0,7]
PredTypes =[]
PredValues = []
FuturedPred = [1,1]
#TFT2 1 year
PredSource = ['Dynamic','Dynamic','Dynamic','Dynamic']
PredSourceNumber = [0,6,7,8]
FuturedPred = [1,1,1,1]
# + [markdown] id="DZbYR4a2lGCe" pycharm={"name": "#%% md\n"}
# ### Choose Input and Predicted Quantities
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="tXz5CLaOlOnn" outputId="de0a8387-ef3a-44ce-d5e9-7710410318f0" pycharm={"name": "#%%\n"}
# Gregor: DELETE some portions of this, review and identify
# PARAMETER. SUPER IMPORTANT. NEEDS TO BE STUDIED
if len(InputSource) != len(InputSourceNumber):
printexit(' Inconsistent Source Lengths ' + str(len(InputSource)) + ' ' +str(len(InputSourceNumber)) )
if len(PredSource) != len(PredSourceNumber):
printexit(' Inconsistent Prediction Lengths ' + str(len(PredSource)) + ' ' + str(len(PredSourceNumber)) )
# Executed by all even if GenerateFutures false except for direct Romeo data
if not UseFutures:
LengthFutures = 0
print(startbold + "Number of Futures -- separate for each regular prediction " +str(LengthFutures) + resetfonts)
Usedaystart = False
if len(PredSource) > 0: # set up Predictions
NumpredbasicperTime = len(PredSource)
FuturedPointer = np.full(NumpredbasicperTime,-1,dtype=int)
NumpredFuturedperTime = 0
NumpredfromInputsperTime = 0
for ipred in range(0,len(PredSource)):
if PredSource[ipred] == 'Dynamic':
NumpredfromInputsperTime += 1
countinputs = 0
countcalcs = 0
for ipred in range(0,len(PredSource)):
if not(PredSource[ipred] == 'Dynamic' or PredSource[ipred] == 'Calc'):
printexit('Illegal Prediction ' + str(ipred) + ' ' + PredSource[ipred])
if PredSource[ipred] == 'Dynamic':
countinputs += 1
else:
countcalcs += 1
if FuturedPred[ipred] >= 0:
if LengthFutures > 0:
FuturedPred[ipred] = NumpredFuturedperTime
FuturedPointer[ipred] = NumpredFuturedperTime
NumpredFuturedperTime += 1
else:
FuturedPred[ipred] = -1
else: # Set defaults
NumpredfromInputsperTime = NumpredFuturedperTime
FuturedPointer = np.full(NumpredbasicperTime,-1,dtype=int)
PredSource =[]
PredSourceNumber = []
FuturedPred =[]
futurepos = 0
for ipred in range(0,NumpredFuturedperTime):
PredSource.append('Dynamic')
PredSourceNumber.append(ipred)
futured = -1
if LengthFutures > 0:
futured = futurepos
FuturedPointer[ipred] = futurepos
futurepos += 1
FuturedPred.append(futured)
for ipred in range(0,NumTimeSeriesCalculated):
PredSource.append('Calc')
PredSourceNumber.append(ipred)
FuturedPred.append(-1)
print('Number of Predictions ' + str(len(PredSource)))
PropertyNameIndex = np.empty(NpropperTime, dtype = np.int32)
PropertyAverageValuesPointer = np.empty(NpropperTime, dtype = np.int32)
for iprop in range(0,NpropperTime):
PropertyNameIndex[iprop] = iprop # names
PropertyAverageValuesPointer[iprop] = iprop # normalizations
# Reset Source -- if OK as read don't set InputSource InputSourceNumber
# Reset NormedDynamicPropertyTimeSeries and NormedInputStaticProps
# Reset NpropperTime = NpropperTimeStatic + NpropperTimeDynamic
if len(InputSource) > 0: # Reset Input Source
NewNpropperTimeStatic = 0
NewNpropperTimeDynamic = 0
for isource in range(0,len(InputSource)):
if InputSource[isource] == 'Static':
NewNpropperTimeStatic += 1
if InputSource[isource] == 'Dynamic':
NewNpropperTimeDynamic += 1
NewNormedDynamicPropertyTimeSeries = np.empty([Num_Time,Nloc,NewNpropperTimeDynamic],dtype = np.float32)
NewNormedInputStaticProps = np.empty([Nloc,NewNpropperTimeStatic],dtype = np.float32)
NewNpropperTime = NewNpropperTimeStatic + NewNpropperTimeDynamic
NewPropertyNameIndex = np.empty(NewNpropperTime, dtype = np.int32)
NewPropertyAverageValuesPointer = np.empty(NewNpropperTime, dtype = np.int32)
countstatic = 0
countdynamic = 0
for isource in range(0,len(InputSource)):
if InputSource[isource] == 'Static':
OldstaticNumber = InputSourceNumber[isource]
NewNormedInputStaticProps[:,countstatic] = NormedInputStaticProps[:,OldstaticNumber]
NewPropertyNameIndex[countstatic] = PropertyNameIndex[OldstaticNumber]
NewPropertyAverageValuesPointer[countstatic] = PropertyAverageValuesPointer[OldstaticNumber]
countstatic += 1
elif InputSource[isource] == 'Dynamic':
OlddynamicNumber =InputSourceNumber[isource]
NewNormedDynamicPropertyTimeSeries[:,:,countdynamic] = NormedDynamicPropertyTimeSeries[:,:,OlddynamicNumber]
NewPropertyNameIndex[countdynamic+NewNpropperTimeStatic] = PropertyNameIndex[OlddynamicNumber+NpropperTimeStatic]
NewPropertyAverageValuesPointer[countdynamic+NewNpropperTimeStatic] = PropertyAverageValuesPointer[OlddynamicNumber+NpropperTimeStatic]
countdynamic += 1
else:
printexit('Illegal Property ' + str(isource) + ' ' + InputSource[isource])
else: # pretend data altered
NewPropertyNameIndex = PropertyNameIndex
NewPropertyAverageValuesPointer = PropertyAverageValuesPointer
NewNpropperTime = NpropperTime
NewNpropperTimeStatic = NpropperTimeStatic
NewNpropperTimeDynamic = NpropperTimeDynamic
NewNormedInputStaticProps = NormedInputStaticProps
NewNormedDynamicPropertyTimeSeries = NormedDynamicPropertyTimeSeries
# + [markdown] id="Yb8-aCUg3Ry5" pycharm={"name": "#%% md\n"}
# ### Calculate Futures
#
# Start Predictions
# + colab={"base_uri": "https://localhost:8080/", "height": 7601} id="Mx4PkF7nkLu_" outputId="404dfa50-efa7-4e63-a60d-bbd3346a1718" pycharm={"name": "#%%\n"}
# Order of Predictions *****************************
# Basic "futured" Predictions from property dynamic arrays
# Additional predictions without futures and NOT in property arrays including Calculated time series
# LengthFutures predictions for first NumpredFuturedperTime predictions
# Special predictions (temporal, positional) added later
NpredperTime = NumpredbasicperTime + NumpredFuturedperTime*LengthFutures
Npredperseq = NpredperTime
Predictionbasicname = [' '] * NumpredbasicperTime
for ipred in range(0,NumpredbasicperTime):
if PredSource[ipred] == 'Dynamic':
Predictionbasicname[ipred] = InputPropertyNames[PredSourceNumber[ipred]+NpropperTimeStatic]
else:
Predictionbasicname[ipred]= NamespredCalculated[PredSourceNumber[ipred]]
TotalFutures = 0
if NumpredFuturedperTime <= 0:
GenerateFutures = False
if GenerateFutures:
TotalFutures = NumpredFuturedperTime * LengthFutures
print(startbold + 'Predictions Total ' + str(Npredperseq) + ' Basic ' + str(NumpredbasicperTime) + ' Of which futured are '
+ str(NumpredFuturedperTime) + ' Giving number explicit futures ' + str(TotalFutures) + resetfonts )
Predictionname = [' '] * Npredperseq
Predictionnametype = [' '] * Npredperseq
Predictionoldvalue = np.empty(Npredperseq, dtype=int)
Predictionnewvalue = np.empty(Npredperseq, dtype=int)
Predictionday = np.empty(Npredperseq, dtype=int)
PredictionAverageValuesPointer = np.empty(Npredperseq, dtype=int)
Predictionwgt = [1.0] * Npredperseq
for ipred in range(0,NumpredbasicperTime):
Predictionnametype[ipred] = PredSource[ipred]
Predictionoldvalue[ipred] = PredSourceNumber[ipred]
Predictionnewvalue[ipred] = ipred
if PredSource[ipred] == 'Dynamic':
PredictionAverageValuesPointer[ipred] = NpropperTimeStatic + Predictionoldvalue[ipred]
else:
PredictionAverageValuesPointer[ipred] = NpropperTime + PredSourceNumber[ipred]
Predictionwgt[ipred] = 1.0
Predictionday[ipred] = 1
extrastring =''
Predictionname[ipred] = 'Next ' + Predictionbasicname[ipred]
if FuturedPred[ipred] >= 0:
extrastring = ' Explicit Futures Added '
print(str(ipred)+ ' Internal Property # ' + str(PredictionAverageValuesPointer[ipred]) + ' ' + Predictionname[ipred]
+ ' Weight ' + str(round(Predictionwgt[ipred],3)) + ' Day ' + str(Predictionday[ipred]) + extrastring )
for ifuture in range(0,LengthFutures):
for ipred in range(0,NumpredbasicperTime):
if FuturedPred[ipred] >= 0:
FuturedPosition = NumpredbasicperTime + NumpredFuturedperTime*ifuture + FuturedPred[ipred]
Predictionname[FuturedPosition] = Predictionbasicname[ipred] + ' ' + Futures[ifuture].name
Predictionday[FuturedPosition] = Futures[ifuture].days[0]
Predictionwgt[FuturedPosition] = Futures[ifuture].classweight
Predictionnametype[FuturedPosition] = Predictionnametype[ipred]
Predictionoldvalue[FuturedPosition] = Predictionoldvalue[ipred]
Predictionnewvalue[FuturedPosition] = Predictionnewvalue[ipred]
PredictionAverageValuesPointer[FuturedPosition] = PredictionAverageValuesPointer[ipred]
print(str(iprop)+ ' Internal Property # ' + str(PredictionAverageValuesPointer[FuturedPosition]) + ' ' +
Predictionname[FuturedPosition] + ' Weight ' + str(round(Predictionwgt[FuturedPosition],3))
+ ' Day ' + str(Predictionday[FuturedPosition]) + ' This is Explicit Future ')
Predictionnamelookup = {}
print(startbold + '\nBasic Predicted Quantities' + resetfonts)
for ipred in range(0,Npredperseq):
Predictionnamelookup[Predictionname[ipred]] = ipred
iprop = Predictionnewvalue[ipred]
line = startbold + startred + Predictionbasicname[iprop]
line += ' Weight ' + str(round(Predictionwgt[ipred],4))
if (iprop < NumpredFuturedperTime) or (iprop >= NumpredbasicperTime):
line += ' Day= ' + str(Predictionday[ipred])
line += ' Name ' + Predictionname[ipred]
line += resetfonts
jpred = PredictionAverageValuesPointer[ipred]
line += ' Processing Root ' + str(QuantityTakeroot[jpred])
for proppredval in range (0,7):
line += ' ' + QuantityStatisticsNames[proppredval] + ' ' + str(round(QuantityStatistics[jpred,proppredval],3))
print(wraptotext(line,size=150))
print(line)
# Note that only Predictionwgt and Predictionname defined for later addons
# + [markdown] id="4V0SXGd-nVfX" pycharm={"name": "#%% md\n"}
# ### Set up Predictions
#
# first for time arrays; we will extend to sequences next. Sequences include the predictions for final time in sequence.
#
# This is prediction for sequence ending one day before the labelling time index. So sequence must end one unit before last time value
#
# Note this is "pure forecast" which are of quantities used in driving data allowing us to iitialize prediction to input
#
# NaN represents non existent data
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="kXMefJVkkFL7" outputId="95982d7c-f933-4696-b55b-6acd54fd834d" pycharm={"name": "#%%\n"}
if PredictionsfromInputs:
InputPredictionsbyTime = np.zeros([Num_Time, Nloc, Npredperseq], dtype = np.float32)
for ipred in range (0,NumpredbasicperTime):
if Predictionnametype[ipred] == 'Dynamic':
InputPredictionsbyTime[:,:,ipred] = NormedDynamicPropertyTimeSeries[:,:,Predictionoldvalue[ipred]]
else:
InputPredictionsbyTime[:,:,ipred] = NormedCalculatedTimeSeries[:,:,Predictionoldvalue[ipred]]
# Add Futures based on Futured properties
if LengthFutures > 0:
NaNall = np.full([Nloc],NaN,dtype = np.float32)
daystartveto = 0
atendveto = 0
allok = NumpredbasicperTime
for ifuture in range(0,LengthFutures):
for itime in range(0,Num_Time):
ActualTime = itime+Futures[ifuture].days[0]-1
if ActualTime >= Num_Time:
for ipred in range (0,NumpredbasicperTime):
Putithere = FuturedPred[ipred]
if Putithere >=0:
InputPredictionsbyTime[itime,:,NumpredbasicperTime + NumpredFuturedperTime*ifuture + Putithere] = NaNall
atendveto +=1
elif Usedaystart and (itime < Futures[ifuture].daystart):
for ipred in range (0,NumpredbasicperTime):
Putithere = FuturedPred[ipred]
if Putithere >=0:
InputPredictionsbyTime[itime,:,NumpredbasicperTime + NumpredFuturedperTime*ifuture + Putithere] = NaNall
daystartveto +=1
else:
for ipred in range (0,NumpredbasicperTime):
Putithere = FuturedPred[ipred]
if Putithere >=0:
if Predictionnametype[ipred] == 'Dynamic':
InputPredictionsbyTime[itime,:,NumpredbasicperTime + NumpredFuturedperTime*ifuture + Putithere] \
= NormedDynamicPropertyTimeSeries[ActualTime,:,Predictionoldvalue[ipred]]
else:
InputPredictionsbyTime[itime,:,NumpredbasicperTime + NumpredFuturedperTime*ifuture + Putithere] \
= NormedCalculatedTimeSeries[ActualTime,:,Predictionoldvalue[ipred]]
allok += NumpredFuturedperTime
print(startbold + 'Futures Added: Predictions set from inputs OK ' +str(allok) +
' Veto at end ' + str(atendveto) + ' Veto at start ' + str(daystartveto) + ' Times number of locations' + resetfonts)
# + [markdown] id="VlGIiaIWIrYm" pycharm={"name": "#%% md\n"}
# ### Clean-up Input quantities#################
# + colab={"base_uri": "https://localhost:8080/", "height": 90} id="0Gq6G5JjIw_g" outputId="188e12b1-2e42-4314-f8b9-ae4ba3b3b3d1" pycharm={"name": "#%%\n"}
def checkNaN(y):
countNaN = 0
countnotNaN = 0
ctprt = 0
if y is None:
return
if len(y.shape) == 2:
for i in range(0,y.shape[0]):
for j in range(0,y.shape[1]):
if np.math.isnan(y[i, j]):
countNaN += 1
else:
countnotNaN += 1
else:
for i in range(0,y.shape[0]):
for j in range(0,y.shape[1]):
for k in range(0,y.shape[2]):
if np.math.isnan(y[i, j, k]):
countNaN += 1
ctprt += 1
print(str(i) + ' ' + str(j) + ' ' + str(k))
if ctprt > 10:
sys.exit(0)
else:
countnotNaN += 1
percent = (100.0*countNaN)/(countNaN + countnotNaN)
print(' is NaN ',str(countNaN),' percent ',str(round(percent,2)),' not NaN ', str(countnotNaN))
# Clean-up Input Source
if len(InputSource) > 0:
PropertyNameIndex = NewPropertyNameIndex
NewPropertyNameIndex = None
PropertyAverageValuesPointer = NewPropertyAverageValuesPointer
NewPropertyAverageValuesPointer = None
NormedInputStaticProps = NewNormedInputStaticProps
NewNormedInputStaticProps = None
NormedDynamicPropertyTimeSeries = NewNormedDynamicPropertyTimeSeries
NewNormedDynamicPropertyTimeSeries = None
NpropperTime = NewNpropperTime
NpropperTimeStatic = NewNpropperTimeStatic
NpropperTimeDynamic = NewNpropperTimeDynamic
print('Static Properties')
if NpropperTimeStatic > 0 :
checkNaN(NormedInputStaticProps)
else:
print(' None Defined')
print('Dynamic Properties')
checkNaN(NormedDynamicPropertyTimeSeries)
# + [markdown] id="VZyZD9mEio0z" pycharm={"name": "#%% md\n"}
# ###Setup Sequences and UseTFTModel
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="kTQVBsqmix8O" outputId="7f2a12ec-c2a5-427e-a542-8928580c3f24" pycharm={"name": "#%%\n"}
Num_SeqExtraUsed = Tseq-1
Num_Seq = Num_Time - Tseq
Num_SeqPred = Num_Seq
TSeqPred = Tseq
TFTExtraTimes = 0
Num_TimeTFT = Num_Time
if UseTFTModel:
TFTExtraTimes = 1 + LengthFutures
SymbolicWindows = True
Num_SeqExtraUsed = Tseq # as last position needed in input
Num_TimeTFT = Num_Time +TFTExtraTimes
Num_SeqPred = Num_Seq
TseqPred = Tseq
# If SymbolicWindows, sequences are not made but we use same array with that dimension (RawInputSeqDimension) set to 1
# reshape can get rid of this irrelevant dimension
# Predictions and Input Properties are associated with sequence number which is first time value used in sequence
# if SymbolicWindows false then sequences are labelled by sequence # and contain time values from sequence # to sequence# + Tseq-1
# if SymbolicWindows True then sequences are labelled by time # and contain one value. They are displaced by Tseq
# If TFT Inputs and Predictions do NOT differ by Tseq
# Num_SeqExtra extra positions in RawInputSequencesTOT for Symbolic windows True as need to store full window
# TFTExtraTimes are extra times
RawInputSeqDimension = Tseq
Num_SeqExtra = 0
if SymbolicWindows:
RawInputSeqDimension = 1
Num_SeqExtra = Num_SeqExtraUsed
# + [markdown] id="XYgeVR4S11pc" pycharm={"name": "#%% md\n"}
# ###Generate Sequences from Time labelled data
# given Tseq set above
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="KUnmDWwS3Iai" outputId="e4555c6d-5220-487d-fab2-fa8325da18d4" pycharm={"name": "#%%\n"}
if GenerateSequences:
UseProperties = np.full(NpropperTime, True, dtype=bool)
Npropperseq = 0
IndexintoPropertyArrays = np.empty(NpropperTime, dtype = int)
for iprop in range(0,NpropperTime):
if UseProperties[iprop]:
IndexintoPropertyArrays[Npropperseq] = iprop
Npropperseq +=1
RawInputSequences = np.zeros([Num_Seq + Num_SeqExtra, Nloc, RawInputSeqDimension, Npropperseq], dtype =np.float32)
RawInputPredictions = np.zeros([Num_SeqPred, Nloc, Npredperseq], dtype =np.float32)
locationarray = np.empty(Nloc, dtype=np.float32)
for iseq in range(0,Num_Seq + Num_SeqExtra):
for windowposition in range(0,RawInputSeqDimension):
itime = iseq + windowposition
for usedproperty in range (0,Npropperseq):
iprop = IndexintoPropertyArrays[usedproperty]
if iprop>=NpropperTimeStatic:
jprop =iprop-NpropperTimeStatic
locationarray = NormedDynamicPropertyTimeSeries[itime,:,jprop]
else:
locationarray = NormedInputStaticProps[:,iprop]
RawInputSequences[iseq,:,windowposition,usedproperty] = locationarray
if iseq < Num_SeqPred:
RawInputPredictions[iseq,:,:] = InputPredictionsbyTime[iseq+TseqPred,:,:]
print(startbold + 'Sequences set from Time values Num Seq ' + str(Num_SeqPred) + ' Time ' +str(Num_Time) + resetfonts)
NormedInputTimeSeries = None
NormedDynamicPropertyTimeSeries = None
if GarbageCollect:
gc.collect()
GlobalTimeMask = np.empty([1,1,1,Tseq,Tseq],dtype =np.float32)
# + [markdown] id="lprQwdZFby5Y" pycharm={"name": "#%% md\n"}
# ### Define Possible Temporal and Spatial Positional Encodings
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="Tu9Oy46Nb4LO" outputId="49a56c08-5d4b-4952-bcfe-f7e1f3f5ae8e" pycharm={"name": "#%%\n"}
# PARAMETER. Possible functions as input MLCOMMONS RELEVANT
def LinearLocationEncoding(TotalLoc):
linear = np.empty(TotalLoc, dtype=float)
for i in range(0,TotalLoc):
linear[i] = float(i)/float(TotalLoc)
return linear
def LinearTimeEncoding(Dateslisted):
Firstdate = Dateslisted[0]
numtofind = len(Dateslisted)
dayrange = (Dateslisted[numtofind-1]-Firstdate).days + 1
linear = np.empty(numtofind, dtype=float)
for i in range(0,numtofind):
linear[i] = float((Dateslisted[i]-Firstdate).days)/float(dayrange)
return linear
def P2TimeEncoding(numtofind):
P2 = np.empty(numtofind, dtype=float)
for i in range(0,numtofind):
x = -1 + 2.0*i/(numtofind-1)
P2[i] = 0.5*(3*x*x-1)
return P2
def P3TimeEncoding(numtofind):
P3 = np.empty(numtofind, dtype=float)
for i in range(0,numtofind):
x = -1 + 2.0*i/(numtofind-1)
P3[i] = 0.5*(5*x*x-3)*x
return P3
def P4TimeEncoding(numtofind):
P4 = np.empty(numtofind, dtype=float)
for i in range(0,numtofind):
x = -1 + 2.0*i/(numtofind-1)
P4[i] = 0.125*(35*x*x*x*x - 30*x*x + 3)
return P4
def WeeklyTimeEncoding(Dateslisted):
numtofind = len(Dateslisted)
costheta = np.empty(numtofind, dtype=float)
sintheta = np.empty(numtofind, dtype=float)
for i in range(0,numtofind):
j = Dateslisted[i].date().weekday()
theta = float(j)*2.0*math.pi/7.0
costheta[i] = math.cos(theta)
sintheta[i] = math.sin(theta)
return costheta, sintheta
def AnnualTimeEncoding(Dateslisted):
numtofind = len(Dateslisted)
costheta = np.empty(numtofind, dtype=float)
sintheta = np.empty(numtofind, dtype=float)
for i in range(0,numtofind):
runningdate = Dateslisted[i]
year = runningdate.year
datebeginyear = datetime(year, 1, 1)
displacement = (runningdate-datebeginyear).days
daysinyear = (datetime(year,12,31)-datebeginyear).days+1
if displacement >= daysinyear:
printexit("EXIT Bad Date ", runningdate)
theta = float(displacement)*2.0*math.pi/float(daysinyear)
costheta[i] = math.cos(theta)
sintheta[i] = math.sin(theta)
return costheta, sintheta
def ReturnEncoding(numtofind,Typeindex, Typevalue):
Dummy = costheta = np.empty(0, dtype=float)
if Typeindex == 1:
return LinearoverLocationEncoding, Dummy, ('LinearSpace',0.,1.0,0.5,0.2887), ('Dummy',0.,0.,0.,0.)
if Typeindex == 2:
if Dailyunit == 1:
return CosWeeklytimeEncoding, SinWeeklytimeEncoding, ('CosWeekly',-1.0, 1.0, 0.,0.7071), ('SinWeekly',-1.0, 1.0, 0.,0.7071)
else:
return Dummy, Dummy, ('Dummy',0.,0.,0.,0.), ('Dummy',0.,0.,0.,0.)
if Typeindex == 3:
return CosAnnualtimeEncoding, SinAnnualtimeEncoding, ('CosAnnual',-1.0, 1.0, 0.,0.7071), ('SinAnnual',-1.0, 1.0, 0.,0.7071)
if Typeindex == 4:
if Typevalue == 0:
ConstArray = np.full(numtofind,0.5, dtype = float)
return ConstArray, Dummy, ('Constant',0.5,0.5,0.5,0.0), ('Dummy',0.,0.,0.,0.)
if Typevalue == 1:
return LinearovertimeEncoding, Dummy, ('LinearTime',0., 1.0, 0.5,0.2887), ('Dummy',0.,0.,0.,0.)
if Typevalue == 2:
return P2TimeEncoding(numtofind), Dummy, ('P2-Time',-1.0, 1.0, 0.,0.4472), ('Dummy',0.,0.,0.,0.)
if Typevalue == 3:
return P3TimeEncoding(numtofind), Dummy, ('P3-Time',-1.0, 1.0, 0.,0.3780), ('Dummy',0.,0.,0.,0.)
if Typevalue == 4:
return P4TimeEncoding(numtofind), Dummy, ('P4-Time',-1.0, 1.0, 0.,0.3333), ('Dummy',0.,0.,0.,0.)
if Typeindex == 5:
costheta = np.empty(numtofind, dtype=float)
sintheta = np.empty(numtofind, dtype=float)
j = 0
for i in range(0,numtofind):
theta = float(j)*2.0*math.pi/Typevalue
costheta[i] = math.cos(theta)
sintheta[i] = math.sin(theta)
j += 1
if j >= Typevalue:
j = 0
return costheta, sintheta,('Cos '+str(Typevalue)+ ' Len',-1.0, 1.0,0.,0.7071), ('Sin '+str(Typevalue)+ ' Len',-1.0, 1.0,0.,0.7071)
# Dates set up in Python datetime format as Python LISTS
# All encodings are Numpy arrays
print("Total number of Time Units " + str(NumberofTimeunits) + ' ' + TimeIntervalUnitName)
if NumberofTimeunits != (Num_Seq + Tseq):
printexit("EXIT Wrong Number of Time Units " + str(Num_Seq + Tseq))
Dateslist = []
for i in range(0,NumberofTimeunits + TFTExtraTimes):
Dateslist.append(InitialDate+timedelta(days=i*Dailyunit))
LinearoverLocationEncoding = LinearLocationEncoding(Nloc)
LinearovertimeEncoding = LinearTimeEncoding(Dateslist)
if Dailyunit == 1:
CosWeeklytimeEncoding, SinWeeklytimeEncoding = WeeklyTimeEncoding(Dateslist)
CosAnnualtimeEncoding, SinAnnualtimeEncoding = AnnualTimeEncoding(Dateslist)
# Encodings
# linearlocationposition
# Supported Time Dependent Probes that can be in properties and/or predictions
# Special
# Annual
# Weekly
#
# Top Down
# TD0 Constant at 0.5
# TD1 Linear from 0 to 1
# TD2 P2(x) where x goes from -1 to 1 as time goes from start to end
#
# Bottom Up
# n-way Cos and sin theta where n = 4 7 8 16 24 32
EncodingTypes = {'Spatial':1, 'Weekly':2,'Annual':3,'TopDown':4,'BottomUp':5}
PropIndex =[]
PropNameMeanStd = []
PropMeanStd = []
PropArray = []
PropPosition = []
PredIndex =[]
PredNameMeanStd = []
PredArray = []
PredPosition = []
Numberpropaddons = 0
propposition = Npropperseq
Numberpredaddons = 0
predposition = Npredperseq
numprop = len(PropTypes)
if numprop != len(PropValues):
printexit('Error in property addons ' + str(numprop) + ' ' + str(len(PropValues)))
for newpropinlist in range(0,numprop):
Typeindex = EncodingTypes[PropTypes[newpropinlist]]
a,b,c,d = ReturnEncoding(Num_Time + TFTExtraTimes,Typeindex, PropValues[newpropinlist])
if c[0] != 'Dummy':
PropIndex.append(Typeindex)
PropNameMeanStd.append(c)
InputPropertyNames.append(c[0])
PropArray.append(a)
PropPosition.append(propposition)
propposition += 1
Numberpropaddons += 1
line = ' '
for ipr in range(0,20):
line += str(round(a[ipr],4)) + ' '
# print('c'+line)
if d[0] != 'Dummy':
PropIndex.append(Typeindex)
PropNameMeanStd.append(d)
InputPropertyNames.append(d[0])
PropArray.append(b)
PropPosition.append(propposition)
propposition += 1
Numberpropaddons += 1
line = ' '
for ipr in range(0,20):
line += str(round(b[ipr],4)) + ' '
# print('d'+line)
numpred = len(PredTypes)
if numpred != len(PredValues):
printexit('Error in prediction addons ' + str(numpred) + ' ' + str(len(PredValues)))
for newpredinlist in range(0,numpred):
Typeindex = EncodingTypes[PredTypes[newpredinlist]]
a,b,c,d = ReturnEncoding(Num_Time + TFTExtraTimes,Typeindex, PredValues[newpredinlist])
if c[0] != 'Dummy':
PredIndex.append(Typeindex)
PredNameMeanStd.append(c)
PredArray.append(a)
Predictionname.append(c[0])
Predictionnamelookup[c] = predposition
PredPosition.append(predposition)
predposition += 1
Numberpredaddons += 1
Predictionwgt.append(0.25)
if d[0] != 'Dummy':
PredIndex.append(Typeindex)
PredNameMeanStd.append(d)
PredArray.append(b)
Predictionname.append(d[0])
Predictionnamelookup[d[0]] = predposition
PredPosition.append(predposition)
predposition += 1
Numberpredaddons += 1
Predictionwgt.append(0.25)
# + [markdown] id="ANMrg0vjoPxS" pycharm={"name": "#%% md\n"}
# ### Add in Temporal and Spatial Encoding
# + colab={"base_uri": "https://localhost:8080/", "height": 2807} id="I977Ffv_obEC" outputId="d90c52b6-4a19-4ac4-90b2-5ea4cbeb2d39" pycharm={"name": "#%%\n"}
def SetNewAverages(InputList): # name min max mean std
results = np.empty(7, dtype = np.float32)
results[0] = InputList[1]
results[1] = InputList[2]
results[2] = 1.0
results[3] = InputList[3]
results[4] = InputList[4]
results[5] = InputList[3]
results[6] = InputList[4]
return results
NpropperseqTOT = Npropperseq + Numberpropaddons
# These include both Property and Prediction Variables
NpropperTimeMAX =len(QuantityTakeroot)
NewNpropperTimeMAX = NpropperTimeMAX + Numberpropaddons + Numberpredaddons
NewQuantityStatistics = np.zeros([NewNpropperTimeMAX,7], dtype=np.float32)
NewQuantityTakeroot = np.full(NewNpropperTimeMAX,1,dtype=int) # All new ones aare 1 and are set here
NewQuantityStatistics[0:NpropperTimeMAX,:] = QuantityStatistics[0:NpropperTimeMAX,:]
NewQuantityTakeroot[0:NpropperTimeMAX] = QuantityTakeroot[0:NpropperTimeMAX]
# Lookup for property names
NewPropertyNameIndex = np.empty(NpropperseqTOT, dtype = np.int32)
NumberofNames = len(InputPropertyNames)-Numberpropaddons
NewPropertyNameIndex[0:Npropperseq] = PropertyNameIndex[0:Npropperseq]
NewPropertyAverageValuesPointer = np.empty(NpropperseqTOT, dtype = np.int32)
NewPropertyAverageValuesPointer[0:Npropperseq] = PropertyAverageValuesPointer[0:Npropperseq]
for propaddons in range(0,Numberpropaddons):
NewPropertyNameIndex[Npropperseq+propaddons] = NumberofNames + propaddons
NewPropertyAverageValuesPointer[Npropperseq+propaddons] = NpropperTimeMAX + propaddons
NewQuantityStatistics[NpropperTimeMAX + propaddons,:] = SetNewAverages(PropNameMeanStd[propaddons])
# Set extra Predictions metadata for Sequences
NpredperseqTOT = Npredperseq + Numberpredaddons
NewPredictionAverageValuesPointer = np.empty(NpredperseqTOT, dtype = np.int32)
NewPredictionAverageValuesPointer[0:Npredperseq] = PredictionAverageValuesPointer[0:Npredperseq]
for predaddons in range(0,Numberpredaddons):
NewPredictionAverageValuesPointer[Npredperseq +predaddons] = NpropperTimeMAX + +Numberpropaddons + predaddons
NewQuantityStatistics[NpropperTimeMAX + Numberpropaddons + predaddons,:] = SetNewAverages(PredNameMeanStd[predaddons])
RawInputSequencesTOT = np.empty([Num_Seq + Num_SeqExtra + TFTExtraTimes, Nloc, RawInputSeqDimension, NpropperseqTOT], dtype =np.float32)
flsize = np.float(Num_Seq + Num_SeqExtra)*np.float(Nloc)*np.float(RawInputSeqDimension)* np.float(NpropperseqTOT)* 4.0
print('Total storage ' +str(round(flsize,0)) + ' Bytes')
for i in range(0,Num_Seq + Num_SeqExtra):
for iprop in range(0,Npropperseq):
RawInputSequencesTOT[i,:,:,iprop] = RawInputSequences[i,:,:,iprop]
for i in range(Num_Seq + Num_SeqExtra,Num_Seq + Num_SeqExtra + TFTExtraTimes):
for iprop in range(0,Npropperseq):
RawInputSequencesTOT[i,:,:,iprop] = NaN
for i in range(0,Num_Seq + Num_SeqExtra + TFTExtraTimes):
for k in range(0,RawInputSeqDimension):
for iprop in range(0, Numberpropaddons):
if PropIndex[iprop] == 1:
continue
RawInputSequencesTOT[i,:,k,PropPosition[iprop]] = PropArray[iprop][i+k]
for iprop in range(0, Numberpropaddons):
if PropIndex[iprop] == 1:
for j in range(0,Nloc):
RawInputSequencesTOT[:,j,:,PropPosition[iprop]] = PropArray[iprop][j]
# Set extra Predictions for Sequences
RawInputPredictionsTOT = np.empty([Num_SeqPred + TFTExtraTimes, Nloc, NpredperseqTOT], dtype =np.float32)
for i in range(0,Num_SeqPred):
for ipred in range(0,Npredperseq):
RawInputPredictionsTOT[i,:,ipred] = RawInputPredictions[i,:,ipred]
for i in range(Num_SeqPred, Num_SeqPred + TFTExtraTimes):
for ipred in range(0,Npredperseq):
RawInputPredictionsTOT[i,:,ipred] = NaN
for i in range(0,Num_SeqPred + TFTExtraTimes):
for ipred in range(0, Numberpredaddons):
if PredIndex[ipred] == 1:
continue
actualarray = PredArray[ipred]
RawInputPredictionsTOT[i,:,PredPosition[ipred]] = actualarray[i+TseqPred]
for ipred in range(0, Numberpredaddons):
if PredIndex[ipred] == 1:
for j in range(0,Nloc):
RawInputPredictionsTOT[:,j,PredPosition[ipred]] = PredArray[ipred][j]
PropertyNameIndex = None
PropertyNameIndex = NewPropertyNameIndex
QuantityStatistics = None
QuantityStatistics = NewQuantityStatistics
QuantityTakeroot = None
QuantityTakeroot = NewQuantityTakeroot
PropertyAverageValuesPointer = None
PropertyAverageValuesPointer = NewPropertyAverageValuesPointer
PredictionAverageValuesPointer = None
PredictionAverageValuesPointer = NewPredictionAverageValuesPointer
print('Time and Space encoding added to input and predictions')
if SymbolicWindows:
SymbolicInputSequencesTOT = np.empty([Num_Seq, Nloc], dtype =np.int32) # This is sequences
for iseq in range(0,Num_Seq):
for iloc in range(0,Nloc):
SymbolicInputSequencesTOT[iseq,iloc] = np.left_shift(iseq,16) + iloc
ReshapedSequencesTOT = np.transpose(RawInputSequencesTOT,(1,0,3,2))
ReshapedSequencesTOT = np.reshape(ReshapedSequencesTOT,(Nloc,Num_Seq + Num_SeqExtra + TFTExtraTimes,NpropperseqTOT))
# To calculate masks (identical to Symbolic windows)
SpacetimeforMask = np.empty([Num_Seq, Nloc], dtype =np.int32)
for iseq in range(0,Num_Seq):
for iloc in range(0,Nloc):
SpacetimeforMask[iseq,iloc] = np.left_shift(iseq,16) + iloc
print(PropertyNameIndex)
print(InputPropertyNames)
for iprop in range(0,NpropperseqTOT):
line = 'Property ' + str(iprop) + ' ' + InputPropertyNames[PropertyNameIndex[iprop]]
jprop = PropertyAverageValuesPointer[iprop]
line += ' Processing Root ' + str(QuantityTakeroot[jprop])
for proppredval in range (0,7):
line += ' ' + QuantityStatisticsNames[proppredval] + ' ' + str(round(QuantityStatistics[jprop,proppredval],3))
print(wraptotext(line,size=150))
for ipred in range(0,NpredperseqTOT):
line = 'Prediction ' + str(ipred) + ' ' + Predictionname[ipred] + ' ' + str(round(Predictionwgt[ipred],3))
jpred = PredictionAverageValuesPointer[ipred]
line += ' Processing Root ' + str(QuantityTakeroot[jpred])
for proppredval in range (0,7):
line += ' ' + QuantityStatisticsNames[proppredval] + ' ' + str(round(QuantityStatistics[jpred,proppredval],3))
print(wraptotext(line,size=150))
RawInputPredictions = None
RawInputSequences = None
if SymbolicWindows:
RawInputSequencesTOT = None
if GarbageCollect:
gc.collect()
# + [markdown] id="B0FxRdZa81_Z" pycharm={"name": "#%% md\n"}
# ###Set up NNSE and Plots including Futures
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="tdFW7f6l3Uo-" outputId="ccaf5744-03b5-4632-dd54-ad1a6004fd18" pycharm={"name": "#%%\n"}
#Set up NNSE Normalized Nash Sutcliffe Efficiency
CalculateNNSE = np.full(NpredperseqTOT, False, dtype = bool)
PlotPredictions = np.full(NpredperseqTOT, False, dtype = bool)
for ipred in range(0,NpredperseqTOT):
CalculateNNSE[ipred] = True
PlotPredictions[ipred] = True
# + [markdown] id="hytLQj7QW3gx" pycharm={"name": "#%% md\n"}
# ## Location Based Validation
# + colab={"base_uri": "https://localhost:8080/", "height": 777} id="s2g_-MHEhyGr" outputId="b6accc74-5e61-4cea-8c2a-8c095bc5a392" pycharm={"name": "#%%\n"}
LocationBasedValidation = False
LocationValidationFraction = 0.0
RestartLocationBasedValidation = False
RestartRunName = RunName
if Earthquake:
LocationBasedValidation = True
LocationValidationFraction = 0.2
RestartLocationBasedValidation = True
RestartRunName = 'EARTHQN-Transformer3'
FullSetValidation = False
global SeparateValandTrainingPlots
SeparateValandTrainingPlots = True
if not LocationBasedValidation:
SeparateValandTrainingPlots = False
LocationValidationFraction = 0.0
NlocValplusTraining = Nloc
ListofTrainingLocs = np.arange(Nloc, dtype = np.int32)
ListofValidationLocs = np.full(Nloc, -1, dtype = np.int32)
MappingtoTraining = np.arange(Nloc, dtype = np.int32)
MappingtoValidation = np.full(Nloc, -1, dtype = np.int32)
TrainingNloc = Nloc
ValidationNloc = 0
if LocationBasedValidation:
if RestartLocationBasedValidation:
InputFileName = APPLDIR + '/Validation' + RestartRunName
with open(InputFileName, 'r', newline='') as inputfile:
Myreader = reader(inputfile, delimiter=',')
header = next(Myreader)
LocationValidationFraction = np.float32(header[0])
TrainingNloc = np.int32(header[1])
ValidationNloc = np.int32(header[2])
ListofTrainingLocs = np.empty(TrainingNloc, dtype = np.int32)
ListofValidationLocs = np.empty(ValidationNloc, dtype = np.int32)
nextrow = next(Myreader)
for iloc in range(0, TrainingNloc):
ListofTrainingLocs[iloc] = np.int32(nextrow[iloc])
nextrow = next(Myreader)
for iloc in range(0, ValidationNloc):
ListofValidationLocs[iloc] = np.int32(nextrow[iloc])
LocationTrainingfraction = 1.0 - LocationValidationFraction
if TrainingNloc + ValidationNloc != Nloc:
printexit('EXIT: Inconsistent location counts for Location Validation ' +str(Nloc)
+ ' ' + str(TrainingNloc) + ' ' + str(ValidationNloc))
print(' Validation restarted Fraction ' +str(round(LocationValidationFraction,4)) + ' ' + RestartRunName)
else:
LocationTrainingfraction = 1.0 - LocationValidationFraction
TrainingNloc = math.ceil(LocationTrainingfraction*Nloc)
ValidationNloc = Nloc - TrainingNloc
np.random.shuffle(ListofTrainingLocs)
ListofValidationLocs = ListofTrainingLocs[TrainingNloc:Nloc]
ListofTrainingLocs = ListofTrainingLocs[0:TrainingNloc]
for iloc in range(0,TrainingNloc):
jloc = ListofTrainingLocs[iloc]
MappingtoTraining[jloc] = iloc
MappingtoValidation[jloc] = -1
for iloc in range(0,ValidationNloc):
jloc = ListofValidationLocs[iloc]
MappingtoValidation[jloc] = iloc
MappingtoTraining[jloc] = -1
if ValidationNloc <= 0:
SeparateValandTrainingPlots = False
if not RestartLocationBasedValidation:
OutputFileName = APPLDIR + '/Validation' + RunName
with open(OutputFileName, 'w', newline='') as outputfile:
Mywriter = writer(outputfile, delimiter=',')
Mywriter.writerow([LocationValidationFraction, TrainingNloc, ValidationNloc] )
Mywriter.writerow(ListofTrainingLocs)
Mywriter.writerow(ListofValidationLocs)
print('Training Locations ' + str(TrainingNloc) + ' Validation Locations ' + str(ValidationNloc))
if ValidationNloc <=0:
LocationBasedValidation = False
if Earthquake:
StartDate = np.datetime64(InitialDate).astype('datetime64[D]') + np.timedelta64(Tseq*Dailyunit + int(Dailyunit/2),'D')
dayrange = np.timedelta64(Dailyunit,'D')
Numericaldate = np.empty(numberspecialeqs, dtype=np.float32)
PrimaryTrainingList = []
SecondaryTrainingList = []
PrimaryValidationList = []
SecondaryValidationList = []
for iquake in range(0,numberspecialeqs):
Numericaldate[iquake] = max(0,math.floor((Specialdate[iquake] - StartDate)/dayrange))
Trainingsecondary = False
Validationsecondary = False
for jloc in range(0,Nloc):
iloc = LookupLocations[jloc] # original location
result = quakesearch(iquake, iloc)
if result == 0:
continue
kloc = MappingtoTraining[jloc]
if result == 1: # Primary
if kloc >= 0:
PrimaryTrainingList.append(iquake)
Trainingsecondary = True
else:
PrimaryValidationList.append(iquake)
Validationsecondary = True
else: # Secondary
if kloc >= 0:
if Trainingsecondary:
continue
Trainingsecondary = True
SecondaryTrainingList.append(iquake)
else:
if Validationsecondary:
continue
Validationsecondary = True
SecondaryValidationList.append(iquake)
iloc = Specialxpos[iquake] + 60*Specialypos[iquake]
jloc = MappedLocations[iloc]
kloc = -2
if jloc >= 0:
kloc = LookupLocations[jloc]
line = str(iquake) + " " + str(Trainingsecondary) + " " + str(Validationsecondary) + " "
line += str(iloc) + " " + str(jloc) + " " + str(kloc) + " " + str(round(Specialmags[iquake],1)) + ' ' + Specialeqname[iquake]
print(line)
PrimaryTrainingvetoquake = np.full(numberspecialeqs,True, dtype = bool)
SecondaryTrainingvetoquake = np.full(numberspecialeqs,True, dtype = bool)
PrimaryValidationvetoquake = np.full(numberspecialeqs,True, dtype = bool)
SecondaryValidationvetoquake = np.full(numberspecialeqs,True, dtype = bool)
for jquake in PrimaryTrainingList:
PrimaryTrainingvetoquake[jquake] = False
for jquake in PrimaryValidationList:
PrimaryValidationvetoquake[jquake] = False
for jquake in SecondaryTrainingList:
if not PrimaryTrainingvetoquake[jquake]:
continue
SecondaryTrainingvetoquake[jquake] = False
for jquake in SecondaryValidationList:
if not PrimaryValidationvetoquake[jquake]:
continue
SecondaryValidationvetoquake[jquake] = False
for iquake in range(0,numberspecialeqs):
iloc = Specialxpos[iquake] + 60*Specialypos[iquake]
line = str(iquake) + " Loc " + str(iloc) + " " + str(MappedLocations[iloc]) + " Date " + str(Specialdate[iquake]) + " " + str(Numericaldate[iquake])
line += " " + str(PrimaryTrainingvetoquake[iquake]) + " " + str(SecondaryTrainingvetoquake[iquake])
line += " Val " + str(PrimaryValidationvetoquake[iquake]) + " " + str(SecondaryValidationvetoquake[iquake])
print(line)
# + [markdown] id="33FLmGmcilz5" pycharm={"name": "#%% md\n"}
# ## LSTM Control Parameters EDIT
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="Ds28euHRi5vt" outputId="7477183b-feef-4eee-ccf0-01cebc4aed45" pycharm={"name": "#%%\n"}
CustomLoss = 1
UseClassweights = True
PredictionTraining = False
# Gregor: MODIFY
if (not Hydrology) and (not Earthquake) and (NpredperseqTOT <=2):
useFutures = False
CustomLoss = 0
UseClassweights = False
number_of_LSTMworkers = 1
TFTTransformerepochs = 10
LSTMbatch_size = TrainingNloc
LSTMbatch_size = min(LSTMbatch_size, TrainingNloc)
LSTMactivationvalue = "selu"
LSTMrecurrent_activation = "sigmoid"
LSTMoptimizer = 'adam'
LSTMdropout1=0.2
LSTMrecurrent_dropout1 = 0.2
LSTMdropout2=0.2
LSTMrecurrent_dropout2 = 0.2
number_LSTMnodes= 16
LSTMFinalMLP = 64
LSTMInitialMLP = 32
LSTMThirdLayer = False
LSTMSkipInitial = False
LSTMverbose = 0
AnyOldValidation = 0.0
if LocationBasedValidation:
AnyOldValidation = LocationBasedValidation
LSTMvalidationfrac = AnyOldValidation
# + [markdown] id="dJFhD-nq0fO0" pycharm={"name": "#%% md\n"}
# ## Important Parameters defining Transformer project EDIT
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="oOxm7gWkyjIj" outputId="dc91896f-a382-4216-ed4c-3992b3e9ddb9" pycharm={"name": "#%%\n"}
ActivateAttention = False
DoubleQKV = False
TimeShufflingOnly = False
Transformerbatch_size = 1
Transformervalidationfrac = 0.0
UsedTransformervalidationfrac = 0.0
Transformerepochs = 200
Transformeroptimizer ='adam'
Transformerverbose = 0
TransformerOnlyFullAttention = True
d_model =64
d_Attention = 2 * d_model
if TransformerOnlyFullAttention:
d_Attention = d_model
d_qk = d_model
d_intermediateqk = 2 * d_model
num_heads = 2
num_Encoderlayers = 2
EncoderDropout= 0.1
EncoderActivation = 'selu'
d_EncoderLayer = d_Attention
d_merge = d_model
d_ffn = 4*d_model
MaskingOption = 0
PeriodicInputTemporalEncoding = 7 # natural for COVID
LinearInputTemporalEncoding = -1 # natural for COVID
TransformerInputTemporalEncoding = 10000
UseTransformerInputTemporalEncoding = False
# + [markdown] id="6CdCNdQ_yGWV" pycharm={"name": "#%% md\n"}
# ## General Control Parameters
# + colab={"base_uri": "https://localhost:8080/"} id="fwkXnZZGgJ_1" outputId="2195cab1-2e52-44c5-dc1a-0bb136c6f77b" pycharm={"name": "#%%\n"}
OuterBatchDimension = Num_Seq * TrainingNloc
IndividualPlots = False
Plotrealnumbers = False
PlotsOnlyinTestFIPS = True
ListofTestFIPS = ['36061','53033','17031','6037']
if Earthquake:
ListofTestFIPS = ['','']
Plotrealnumbers = True
StartDate = np.datetime64(InitialDate).astype('datetime64[D]') + np.timedelta64(Tseq*Dailyunit + int(Dailyunit/2),'D')
dayrange = np.timedelta64(Dailyunit,'D')
CutoffDate = np.datetime64('1989-01-01')
NumericalCutoff = math.floor((CutoffDate - StartDate)/dayrange)
print('Start ' + str(StartDate) + ' Cutoff ' + str(CutoffDate) + " sequence index " + str(NumericalCutoff))
TimeCutLabel = [' All Time ',' Start ',' End ']
# + colab={"base_uri": "https://localhost:8080/"} id="4V88mmqms1pq" outputId="9d9543d2-346b-491a-c1d9-cbbde26944a6" pycharm={"name": "#%%\n"}
print("Size of sequence window Tseq ", str(Tseq))
print("Number of Sequences in time Num_Seq ", str(Num_Seq))
print("Number of locations Nloc ", str(Nloc))
print("Number of Training Sequences in Location and Time ", str(OuterBatchDimension))
print("Number of internal properties per sequence including static or dynamic Npropperseq ", str(Npropperseq))
print("Number of internal properties per sequence adding in explicit space-time encoding ", str(NpropperseqTOT))
print("Total number of predictions per sequence NpredperseqTOT ", str(NpredperseqTOT))
# + [markdown] id="ikdmffIpA6AC" pycharm={"name": "#%% md\n"}
# ## Useful Time series utilities
# + [markdown] id="g2QTzC0vnSGP" pycharm={"name": "#%% md\n"}
# ### DLprediction
#
# Prediction and Visualization LSTM+Transformer
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="aarkiMHirB1S" outputId="24df045e-f1f3-4a9e-ae2a-1e72ea531cbd" pycharm={"name": "#%%\n"}
def DLprediction(Xin, yin, DLmodel, modelflag, LabelFit =''):
# modelflag = 0 LSTM = 1 Transformer
# Input is the windows [Num_Seq] [Nloc] [Tseq] [NpropperseqTOT] (SymbolicWindows False)
# Input is the sequences [Nloc] [Num_Time-1] [NpropperseqTOT] (SymbolicWindows True)
# Input Predictions are always [Num_Seq] [NLoc] [NpredperseqTOT]
current_time = timenow()
print(startbold + startred + current_time + ' ' + RunName + " DLPrediction " +RunComment + resetfonts)
FitPredictions = np.zeros([Num_Seq, Nloc, NpredperseqTOT], dtype =np.float32)
# Compare to RawInputPredictionsTOT
RMSEbyclass = np.zeros([NpredperseqTOT,3], dtype=np.float64)
RMSETRAINbyclass = np.zeros([NpredperseqTOT,3], dtype=np.float64)
RMSEVALbyclass = np.zeros([NpredperseqTOT,3], dtype=np.float64)
RMSVbyclass = np.zeros([NpredperseqTOT], dtype=np.float64)
AbsEbyclass = np.zeros([NpredperseqTOT], dtype=np.float64)
AbsVbyclass = np.zeros([NpredperseqTOT], dtype=np.float64)
ObsVbytimeandclass = np.zeros([Num_Seq, NpredperseqTOT,3], dtype=np.float64)
Predbytimeandclass = np.zeros([Num_Seq, NpredperseqTOT,3], dtype=np.float64)
countbyclass = np.zeros([NpredperseqTOT,3], dtype=np.float64)
countVALbyclass = np.zeros([NpredperseqTOT,3], dtype=np.float64)
countTRAINbyclass = np.zeros([NpredperseqTOT,3], dtype=np.float64)
totalcount = 0
overcount = 0
weightedcount = 0.0
weightedovercount = 0.0
weightedrmse1 = 0.0
weightedrmse1TRAIN = 0.0
weightedrmse1VAL = 0.0
closs = 0.0
dloss = 0.0
eloss = 0.0
floss = 0.0
sw = np.empty([Nloc,NpredperseqTOT],dtype = np.float32)
for iloc in range(0,Nloc):
for k in range(0,NpredperseqTOT):
sw[iloc,k] = Predictionwgt[k]
global tensorsw
tensorsw = tf.convert_to_tensor(sw, np.float32)
Ctime1 = 0.0
Ctime2 = 0.0
Ctime3 = 0.0
samplebar = notebook.trange(Num_Seq, desc='Predict loop', unit = 'sequences')
countingcalls = 0
for iseq in range(0, Num_Seq):
StopWatch.start('label1')
if SymbolicWindows:
if modelflag == 2:
InputVector = np.empty((Nloc,2), dtype = int)
for iloc in range (0,Nloc):
InputVector[iloc,0] = iloc
InputVector[iloc,1] = iseq
else:
InputVector = Xin[:,iseq:iseq+Tseq,:]
else:
InputVector = Xin[iseq]
Time = None
if modelflag == 0:
InputVector = np.reshape(InputVector,(-1,Tseq,NpropperseqTOT))
elif modelflag == 1:
InputVector = np.reshape(InputVector,(1,Tseq*Nloc,NpropperseqTOT))
BasicTimes = np.full(Nloc,iseq, dtype=np.int32)
Time = SetSpacetime(np.reshape(BasicTimes,(1,-1)))
StopWatch.stop('label1')
Ctime1 += StopWatch.get('label1', digits=4)
StopWatch.start('label2')
PredictedVector = DLmodel(InputVector, training = PredictionTraining, Time=Time)
StopWatch.stop('label2')
Ctime2 += StopWatch.get('label2', digits=4)
StopWatch.start('label3')
PredictedVector = np.reshape(PredictedVector,(Nloc,NpredperseqTOT))
TrueVector = yin[iseq]
functionval = numpycustom_lossGCF1(TrueVector,PredictedVector,sw)
closs += functionval
PredictedVector_t = tf.convert_to_tensor(PredictedVector)
yin_t = tf.convert_to_tensor(TrueVector)
dloss += weightedcustom_lossGCF1(yin_t,PredictedVector_t,tensorsw)
eloss += custom_lossGCF1spec(yin_t,PredictedVector_t)
OutputLoss = 0.0
FitPredictions[iseq] = PredictedVector
for iloc in range(0,Nloc):
yy = yin[iseq,iloc]
yyhat = PredictedVector[iloc]
sum1 = 0.0
for i in range(0,NpredperseqTOT):
overcount += 1
weightedovercount += Predictionwgt[i]
if math.isnan(yy[i]):
continue
weightedcount += Predictionwgt[i]
totalcount += 1
mse1 = ((yy[i]-yyhat[i])**2)
mse = mse1*sw[iloc,i]
if i < Npredperseq:
floss += mse
sum1 += mse
AbsEbyclass[i] += abs(yy[i] - yyhat[i])
RMSVbyclass[i] += yy[i]**2
AbsVbyclass[i] += abs(yy[i])
RMSEbyclass[i,0] += mse
countbyclass[i,0] += 1.0
if iseq < NumericalCutoff:
countbyclass[i,1] += 1.0
RMSEbyclass[i,1] += mse
else:
countbyclass[i,2] += 1.0
RMSEbyclass[i,2] += mse
if LocationBasedValidation:
if MappingtoTraining[iloc] >= 0:
ObsVbytimeandclass [iseq,i,1] += abs(yy[i])
Predbytimeandclass [iseq,i,1] += abs(yyhat[i])
RMSETRAINbyclass[i,0] += mse
countTRAINbyclass[i,0] += 1.0
if iseq < NumericalCutoff:
RMSETRAINbyclass[i,1] += mse
countTRAINbyclass[i,1] += 1.0
else:
RMSETRAINbyclass[i,2] += mse
countTRAINbyclass[i,2] += 1.0
if MappingtoValidation[iloc] >= 0:
ObsVbytimeandclass [iseq,i,2] += abs(yy[i])
Predbytimeandclass [iseq,i,2] += abs(yyhat[i])
RMSEVALbyclass[i,0] += mse
countVALbyclass[i,0] += 1.0
if iseq < NumericalCutoff:
RMSEVALbyclass[i,1] += mse
countVALbyclass[i,1] += 1.0
else:
RMSEVALbyclass[i,2] += mse
countVALbyclass[i,2] += 1.0
ObsVbytimeandclass [iseq,i,0] += abs(yy[i])
Predbytimeandclass [iseq,i,0] += abs(yyhat[i])
weightedrmse1 += sum1
if LocationBasedValidation:
if MappingtoTraining[iloc] >= 0:
weightedrmse1TRAIN += sum1
if MappingtoValidation[iloc] >= 0:
weightedrmse1VAL += sum1
OutputLoss += sum1
StopWatch.stop('label3')
Ctime3 += StopWatch.get('label3', digits=4)
OutputLoss /= Nloc
countingcalls += 1
samplebar.update(1)
samplebar.set_postfix( Call = countingcalls, TotalLoss = OutputLoss)
print('Times ' + str(round(Ctime1,5)) + ' ' + str(round(Ctime3,5)) + ' TF ' + str(round(Ctime2,5)))
weightedrmse1 /= (Num_Seq * Nloc)
floss /= (Num_Seq * Nloc)
if LocationBasedValidation:
weightedrmse1TRAIN /= (Num_Seq * TrainingNloc)
if ValidationNloc>0:
weightedrmse1VAL /= (Num_Seq * ValidationNloc)
dloss = dloss.numpy()
eloss = eloss.numpy()
closs /= Num_Seq
dloss /= Num_Seq
eloss /= Num_Seq
current_time = timenow()
line1 = ''
global GlobalTrainingLoss, GlobalValidationLoss, GlobalLoss
GlobalLoss = weightedrmse1
if LocationBasedValidation:
line1 = ' Training ' + str(round(weightedrmse1TRAIN,6)) + ' Validation ' + str(round(weightedrmse1VAL,6))
GlobalTrainingLoss = weightedrmse1TRAIN
GlobalValidationLoss = weightedrmse1VAL
print( startbold + startred + current_time + ' DLPrediction Averages' + ' ' + RunName + ' ' + RunComment + resetfonts)
line = LabelFit + ' ' + RunName + ' Weighted sum over predicted values ' + str(round(weightedrmse1,6))
line += ' No Encoding Preds ' + str(round(floss,6)) + line1
line += ' from loss function ' + str(round(closs,6)) + ' TF version ' + str(round(dloss,6)) + ' TFspec version ' + str(round(eloss,6))
print(wraptotext(line))
print('Count ignoring NaN ' +str(round(weightedcount,4))+ ' Counting NaN ' + str(round(weightedovercount,4)), 70 )
print(' Unwgt Count no NaN ',totalcount, ' Unwgt Count with NaN ',overcount, ' Number Sequences ', Nloc*Num_Seq)
ObsvPred = np.sum( np.abs(ObsVbytimeandclass-Predbytimeandclass) , axis=0)
TotalObs = np.sum( ObsVbytimeandclass , axis=0)
SummedEbyclass = np.divide(ObsvPred,TotalObs)
RMSEbyclass1 = np.divide(RMSEbyclass,countbyclass) # NO SQRT
RMSEbyclass2 = np.sqrt(np.divide(RMSEbyclass[:,0],RMSVbyclass))
RelEbyclass = np.divide(AbsEbyclass, AbsVbyclass)
extracomments = []
line1 = '\nErrors by Prediction Components -- class weights not included except in final Loss components\n Name Count without NaN, '
line2 = 'sqrt(sum errors**2/sum target**2), sum(abs(error)/sum(abs(value), abs(sum(abs(value)-abs(pred)))/sum(abs(pred)'
print(wraptotext(startbold + startred + line1 + line2 + resetfonts))
countbasic = 0
for i in range(0,NpredperseqTOT):
line = startbold + startred + ' AVG MSE '
for timecut in range(0,3):
line += TimeCutLabel[timecut] + 'Full ' + str(round(RMSEbyclass1[i,timecut],6)) + resetfonts
if LocationBasedValidation:
RTRAIN = np.divide(RMSETRAINbyclass[i],countTRAINbyclass[i])
RVAL = np.full(3,0.0, dtype =np.float32)
if countVALbyclass[i,0] > 0:
RVAL = np.divide(RMSEVALbyclass[i],countVALbyclass[i])
for timecut in range(0,3):
line += startbold + startpurple + TimeCutLabel[timecut] + 'TRAIN ' + resetfonts + str(round(RTRAIN[timecut],6))
line += startbold + ' VAL ' + resetfonts + str(round(RVAL[timecut],6))
else:
RTRAIN = RMSEbyclass1[i]
RVAL = np.full(3,0.0, dtype =np.float32)
print(wraptotext(str(i) + ' ' + startbold + Predictionname[i] + resetfonts + ' All Counts ' + str(round(countbyclass[i,0],0)) + ' IndE^2/IndObs^2 '
+ str(round(100.0*RMSEbyclass2[i],2)) + '% IndE/IndObs ' + str(round(100.0*RelEbyclass[i],2)) + '% summedErr/SummedObs ' + str(round(100.0*SummedEbyclass[i,0],2)) + '%' +line ) )
Trainline = 'AVG MSE F=' + str(round(RTRAIN[0],6)) + ' S=' + str(round(RTRAIN[1],6)) + ' E=' + str(round(RTRAIN[2],6)) + ' TOTAL summedErr/SummedObs ' + str(round(100.0*SummedEbyclass[i,1],2)) + '%'
Valline = 'AVG MSE F=' + str(round(RVAL[0],6)) + ' S=' + str(round(RVAL[1],6)) + ' E=' + str(round(RVAL[2],6)) + ' TOTAL summedErr/SummedObs ' + str(round(100.0*SummedEbyclass[i,2],2)) + '%'
extracomments.append([Trainline, Valline] )
countbasic += 1
if countbasic == NumpredbasicperTime:
countbasic = 0
print(' ')
# Don't use DLPrediction for Transformer Plots. Wait for DL2B,D,E
if modelflag == 1:
return FitPredictions
FindNNSE(yin, FitPredictions)
print('\n Next plots come from DLPrediction')
PredictedQuantity = -NumpredbasicperTime
for ifuture in range (0,1+LengthFutures):
increment = NumpredbasicperTime
if ifuture > 1:
increment = NumpredFuturedperTime
PredictedQuantity += increment
if not PlotPredictions[PredictedQuantity]:
continue
Dumpplot = False
if PredictedQuantity ==0:
Dumpplot = True
Location_summed_plot(ifuture, yin, FitPredictions, extracomments = extracomments, Dumpplot = Dumpplot)
if IndividualPlots:
ProduceIndividualPlots(yin, FitPredictions)
if Earthquake and EarthquakeImagePlots:
ProduceSpatialQuakePlot(yin, FitPredictions)
# Call DLprediction2F here if modelflag=0
DLprediction2F(Xin, yin, DLmodel, modelflag)
return FitPredictions
# + [markdown] id="ZW3dd6kVriWQ" pycharm={"name": "#%% md\n"}
# ### Spatial Earthquake Plots
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="QgmRznp-ryGp" outputId="44c87f89-3775-4ccd-99dc-273206ca228a" pycharm={"name": "#%%\n"}
def ProduceSpatialQuakePlot(Observations, FitPredictions):
current_time = timenow()
print(startbold + startred + current_time + ' Produce Spatial Earthquake Plots ' + RunName + ' ' + RunComment + resetfonts)
dayindexmax = Num_Seq-Plottingdelay
Numdates = 4
denom = 1.0/np.float64(Numdates-1)
for plotdays in range(0,Numdates):
dayindexvalue = math.floor(0.1 + (plotdays*dayindexmax)*denom)
if dayindexvalue < 0:
dayindexvalue = 0
if dayindexvalue > dayindexmax:
dayindexvalue = dayindexmax
FixedTimeSpatialQuakePlot(dayindexvalue,Observations, FitPredictions)
def EQrenorm(casesdeath,value):
if Plotrealnumbers:
predaveragevaluespointer = PredictionAverageValuesPointer[casesdeath]
newvalue = value/QuantityStatistics[predaveragevaluespointer,2] + QuantityStatistics[predaveragevaluespointer,0]
rootflag = QuantityTakeroot[predaveragevaluespointer]
if rootflag == 2:
newvalue = newvalue**2
if rootflag == 3:
newvalue = newvalue**3
else:
newvalue=value
return newvalue
def FixedTimeSpatialQuakePlot(PlotTime,Observations, FitPredictions):
Actualday = InitialDate + timedelta(days=(PlotTime+Tseq))
print(startbold + startred + ' Spatial Earthquake Plots ' + Actualday.strftime("%d/%m/%Y") + ' ' + RunName + ' ' + RunComment + resetfonts)
NlocationsPlotted = Nloc
real = np.zeros([NumpredbasicperTime,NlocationsPlotted])
predict = np.zeros([NumpredbasicperTime,NlocationsPlotted])
print('Ranges for Prediction numbers/names/property pointer')
for PredictedQuantity in range(0,NumpredbasicperTime):
for iloc in range(0,NlocationsPlotted):
real[PredictedQuantity,iloc] = EQrenorm(PredictedQuantity,Observations[PlotTime, iloc, PredictedQuantity])
predict[PredictedQuantity,iloc] = EQrenorm(PredictedQuantity,FitPredictions[PlotTime, iloc, PredictedQuantity])
localmax1 = real[PredictedQuantity].max()
localmin1 = real[PredictedQuantity].min()
localmax2 = predict[PredictedQuantity].max()
localmin2 = predict[PredictedQuantity].min()
predaveragevaluespointer = PredictionAverageValuesPointer[PredictedQuantity]
expectedmax = QuantityStatistics[predaveragevaluespointer,1]
expectedmin = QuantityStatistics[predaveragevaluespointer,0]
print(' Real max/min ' + str(round(localmax1,3)) + ' ' + str(round(localmin1,3))
+ ' Predicted max/min ' + str(round(localmax2,3)) + ' ' + str(round(localmin2,3))
+ ' Overall max/min ' + str(round(expectedmax,3)) + ' ' + str(round(expectedmin,3))
+ str(PredictedQuantity) + ' ' + Predictionbasicname[PredictedQuantity] + str(predaveragevaluespointer))
InputImages =[]
InputTitles =[]
for PredictedQuantity in range(0,NumpredbasicperTime):
InputImages.append(real[PredictedQuantity])
InputTitles.append(Actualday.strftime("%d/%m/%Y") + ' Observed ' + Predictionbasicname[PredictedQuantity])
InputImages.append(predict[PredictedQuantity])
InputTitles.append(Actualday.strftime("%d/%m/%Y") + ' Predicted ' + Predictionbasicname[PredictedQuantity])
plotimages(InputImages,InputTitles,NumpredbasicperTime,2)
# + [markdown] id="ZIHPso_LrPJy" pycharm={"name": "#%% md\n"}
# ###Organize Location v Time Plots
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="1WsspqAef_yR" outputId="fc27d527-155d-46df-c1f2-7d4ea6a684cd" pycharm={"name": "#%%\n"}
def ProduceIndividualPlots(Observations, FitPredictions):
current_time = timenow()
print(startbold + startred + current_time + ' Produce Individual Plots ' + RunName + ' ' + RunComment + resetfonts)
# Find Best and Worst Locations
fips_b, fips_w = bestandworst(Observations, FitPredictions)
if Hydrology or Earthquake:
plot_by_fips(fips_b, Observations, FitPredictions)
plot_by_fips(fips_w, Observations, FitPredictions)
else:
plot_by_fips(6037, Observations, FitPredictions)
plot_by_fips(36061, Observations, FitPredictions)
plot_by_fips(17031, Observations, FitPredictions)
plot_by_fips(53033, Observations, FitPredictions)
if (fips_b!=6037) and (fips_b!=36061) and (fips_b!=17031) and (fips_b!=53033):
plot_by_fips(fips_b, Observations, FitPredictions)
if (fips_w!=6037) and (fips_w!=36061) and (fips_w!=17031) and (fips_w!=53033):
plot_by_fips(fips_w, Observations, FitPredictions)
# Plot top 10 largest cities
sortedcities = np.flip(np.argsort(Locationpopulation))
for pickout in range (0,10):
Locationindex = sortedcities[pickout]
fips = Locationfips[Locationindex]
if not(Hydrology or Earthquake):
if fips == 6037 or fips == 36061 or fips == 17031 or fips == 53033:
continue
if fips == fips_b or fips == fips_w:
continue
plot_by_fips(fips, Observations, FitPredictions)
if LengthFutures > 1:
plot_by_futureindex(2, Observations, FitPredictions)
if LengthFutures > 6:
plot_by_futureindex(7, Observations, FitPredictions)
if LengthFutures > 11:
plot_by_futureindex(12, Observations, FitPredictions)
return
def bestandworst(Observations, FitPredictions):
current_time = timenow()
print(startbold + startred + current_time + ' ' + RunName + " Best and Worst " +RunComment + resetfonts)
keepabserrorvalues = np.zeros([Nloc,NumpredbasicperTime], dtype=np.float64)
keepRMSEvalues = np.zeros([Nloc,NumpredbasicperTime], dtype=np.float64)
testabserrorvalues = np.zeros(Nloc, dtype=np.float64)
testRMSEvalues = np.zeros(Nloc, dtype=np.float64)
real = np.zeros([NumpredbasicperTime,Num_Seq], dtype=np.float64)
predictsmall = np.zeros([NumpredbasicperTime,Num_Seq], dtype=np.float64)
c_error_props = np.zeros([NumpredbasicperTime], dtype=np.float64)
c_error_props = np.zeros([NumpredbasicperTime], dtype=np.float64)
for icity in range(0,Nloc):
validcounts = np.zeros([NumpredbasicperTime], dtype=np.float64)
RMSE = np.zeros([NumpredbasicperTime], dtype=np.float64)
for PredictedQuantity in range(0,NumpredbasicperTime):
for itime in range (0,Num_Seq):
if not math.isnan(Observations[itime, icity, PredictedQuantity]):
real[PredictedQuantity,itime] = Observations[itime, icity, PredictedQuantity]
predictsmall[PredictedQuantity,itime] = FitPredictions[itime, icity, PredictedQuantity]
validcounts[PredictedQuantity] += 1.0
RMSE[PredictedQuantity] += (Observations[itime, icity, PredictedQuantity]-FitPredictions[itime, icity, PredictedQuantity])**2
c_error_props[PredictedQuantity] = cumulative_error(predictsmall[PredictedQuantity], real[PredictedQuantity]) # abs(error) as percentage
keepabserrorvalues[icity,PredictedQuantity] = c_error_props[PredictedQuantity]
keepRMSEvalues[icity,PredictedQuantity] = RMSE[PredictedQuantity] *100. / validcounts[PredictedQuantity]
testabserror = 0.0
testRMSE = 0.0
for PredictedQuantity in range(0,NumpredbasicperTime):
testabserror += c_error_props[PredictedQuantity]
testRMSE += keepRMSEvalues[icity,PredictedQuantity]
testabserrorvalues[icity] = testabserror
testRMSEvalues[icity] = testRMSE
sortingindex = np.argsort(testabserrorvalues)
bestindex = sortingindex[0]
worstindex = sortingindex[Nloc-1]
fips_b = Locationfips[bestindex]
fips_w = Locationfips[worstindex]
current_time = timenow()
print( startbold + "\n" + current_time + " Best " + str(fips_b) + " " + Locationname[bestindex] + " " + Locationstate[bestindex] + ' ABS(error) ' +
str(round(testabserrorvalues[bestindex],2)) + ' RMSE ' + str(round(testRMSEvalues[bestindex],2)) + resetfonts)
for topcities in range(0,10):
localindex = sortingindex[topcities]
printstring = str(topcities) + ") " + str(Locationfips[localindex]) + " " + Locationname[localindex] + " ABS(error) Total " + str(round(testabserrorvalues[localindex],4)) + " Components "
for PredictedQuantity in range(0,NumpredbasicperTime):
printstring += ' ' + str(round(keepabserrorvalues[localindex,PredictedQuantity],2))
print(printstring)
print("\nlist RMSE")
for topcities in range(0,9):
localindex = sortingindex[topcities]
printstring = str(topcities) + ") " + str(Locationfips[localindex]) + " " + Locationname[localindex] + " RMSE Total " + str(round(testRMSEvalues[localindex],4)) + " Components "
for PredictedQuantity in range(0,NumpredbasicperTime):
printstring += ' ' + str(round(keepRMSEvalues[localindex,PredictedQuantity],2))
print(printstring)
print( startbold + "\n" + current_time + " Worst " + str(fips_w) + " " + Locationname[worstindex] + " " + Locationstate[worstindex] + ' ABS(error) ' +
str(round(testabserrorvalues[worstindex],2)) + ' RMSE ' + str(round(testRMSEvalues[worstindex],2)) + resetfonts)
for badcities in range(Nloc-1,Nloc-11,-1):
localindex = sortingindex[badcities]
printstring = str(badcities) + ") " + str(Locationfips[localindex]) + " " + Locationname[localindex] + " ABS(error) Total " + str(round(testabserrorvalues[localindex],4)) + " Components "
for PredictedQuantity in range(0,NumpredbasicperTime):
printstring += ' ' + str(round(keepabserrorvalues[localindex,PredictedQuantity],2))
print(printstring)
print("\nlist RMSE")
for badcities in range(0,9):
localindex = sortingindex[badcities]
printstring = str(badcities) + ") " + str(Locationfips[localindex]) + " " + Locationname[localindex] + " RMSE Total " + str(round(testRMSEvalues[localindex],4)) + " Components "
for PredictedQuantity in range(0,NumpredbasicperTime):
printstring += ' ' + str(round(keepRMSEvalues[localindex,PredictedQuantity],2))
print(printstring)
return fips_b,fips_w
# + [markdown] pycharm={"name": "#%% md\n"}
#
# + [markdown] id="0S2QaUybnLTb" pycharm={"name": "#%% md\n"}
# ### Summed & By Location Plots
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="GrWzXpoTa18c" outputId="236d9feb-7ebd-4e73-9a07-14356818a1ee" pycharm={"name": "#%%\n"}
def setValTrainlabel(iValTrain):
if SeparateValandTrainingPlots:
if iValTrain == 0:
Overalllabel = 'Training '
if GlobalTrainingLoss > 0.0001:
Overalllabel += str(round(GlobalTrainingLoss,5)) + ' '
if iValTrain == 1:
Overalllabel = 'Validation '
if GlobalValidationLoss > 0.0001:
Overalllabel += str(round(GlobalValidationLoss,5)) + ' '
else:
Overalllabel = 'Full ' + str(round(GlobalLoss,5)) + ' '
Overalllabel += RunName + ' '
return Overalllabel
def Location_summed_plot(selectedfuture, Observations, FitPredictions, fill=True, otherlabs= [], otherfits=[], extracomments = None, Dumpplot = False):
# plot sum over locations
current_time = timenow()
print(wraptotext(startbold + startred + current_time + ' Location_summed_plot ' + RunName + ' ' + RunComment + resetfonts))
otherlen = len(otherlabs)
basiclength = Num_Seq
predictlength = LengthFutures
if (not UseFutures) or (selectedfuture > 0):
predictlength = 0
totallength = basiclength + predictlength
if extracomments is None:
extracomments = []
for PredictedQuantity in range(0,NpredperseqTOT):
extracomments.append([' ',''])
NumberValTrainLoops = 1
if SeparateValandTrainingPlots:
NumberValTrainLoops = 2
selectedfield = NumpredbasicperTime + NumpredFuturedperTime*(selectedfuture-1)
selectednumplots = NumpredFuturedperTime
if selectedfuture == 0:
selectedfield = 0
selectednumplots = NumpredbasicperTime
ActualQuantity = np.arange(selectednumplots,dtype=np.int32)
if selectedfuture > 0:
for ipred in range(0,NumpredbasicperTime):
ifuture = FuturedPointer[ipred]
if ifuture >= 0:
ActualQuantity[ifuture] = ipred
real = np.zeros([selectednumplots,NumberValTrainLoops,basiclength])
predictsmall = np.zeros([selectednumplots,NumberValTrainLoops,basiclength])
predict = np.zeros([selectednumplots,NumberValTrainLoops,totallength])
if otherlen!=0:
otherpredict = np.zeros([otherlen,selectednumplots,NumberValTrainLoops, totallength])
for PlottedIndex in range(0,selectednumplots):
PredictedPos = PlottedIndex+selectedfield
ActualObservable = ActualQuantity[PlottedIndex]
for iValTrain in range(0,NumberValTrainLoops):
for iloc in range(0,Nloc):
if SeparateValandTrainingPlots:
if iValTrain == 0:
if MappingtoTraining[iloc] < 0:
continue
if iValTrain == 1:
if MappingtoTraining[iloc] >= 0:
continue
for itime in range (0,Num_Seq):
if np.math.isnan(Observations[itime, iloc, PredictedPos]):
real[PlottedIndex,iValTrain,itime] += FitPredictions[itime, iloc, PredictedPos]
else:
real[PlottedIndex,iValTrain,itime] += Observations[itime, iloc, PredictedPos]
predict[PlottedIndex,iValTrain,itime] += FitPredictions[itime, iloc, PredictedPos]
for others in range (0,otherlen):
otherpredict[others,PlottedIndex,iValTrain,itime] += FitPredictions[itime, iloc, PredictedPos] + otherfits[others,itime, iloc, PredictedPos]
if selectedfuture == 0:
if FuturedPointer[PlottedIndex] >= 0:
for ifuture in range(selectedfuture,LengthFutures):
jfuture = NumpredbasicperTime + NumpredFuturedperTime*ifuture
predict[PlottedIndex,iValTrain,Num_Seq+ifuture] += FitPredictions[itime, iloc,
FuturedPointer[PlottedIndex] + jfuture]
for others in range (0,otherlen):
otherpredict[others,PlottedIndex,iValTrain,Num_Seq+ifuture] += FitPredictions[itime, iloc, PlottedIndex + jfuture] + otherfits[others, itime, iloc, PlottedIndex + jfuture]
for itime in range(0,basiclength):
predictsmall[PlottedIndex,iValTrain,itime] = predict[PlottedIndex,iValTrain,itime]
error = np.absolute(real - predictsmall)
xsmall = np.arange(0,Num_Seq)
neededrows = math.floor((selectednumplots*NumberValTrainLoops +1.1)/2)
iValTrain = -1
PlottedIndex = -1
for rowloop in range(0,neededrows):
plt.rcParams["figure.figsize"] = [16,6]
figure, (ax1,ax2) = plt.subplots(nrows=1, ncols=2)
for kplot in range (0,2):
if NumberValTrainLoops == 2:
iValTrain = kplot
else:
iValTrain = 0
if iValTrain == 0:
PlottedIndex +=1
if PlottedIndex > (selectednumplots-1):
PlottedIndex = selectednumplots-1
Overalllabel = setValTrainlabel(iValTrain)
PredictedPos = PlottedIndex+selectedfield
ActualObservable = ActualQuantity[PlottedIndex]
eachplt = ax1
if kplot == 1:
eachplt = ax2
Overalllabel = 'Full '
if SeparateValandTrainingPlots:
if iValTrain == 0:
Overalllabel = 'Training '
if GlobalTrainingLoss > 0.0001:
Overalllabel += str(round(GlobalTrainingLoss,5)) + ' '
if iValTrain == 1:
Overalllabel = 'Validation '
if GlobalValidationLoss > 0.0001:
Overalllabel += str(round(GlobalValidationLoss,5)) + ' '
else:
Overalllabel += RunName + ' ' + str(round(GlobalLoss,5)) + ' '
maxplot = np.float32(totallength)
if UseRealDatesonplots:
StartDate = np.datetime64(InitialDate).astype('datetime64[D]') + np.timedelta64(Tseq*Dailyunit + math.floor(Dailyunit/2),'D')
EndDate = StartDate + np.timedelta64(totallength*Dailyunit)
datemin, datemax = makeadateplot(figure,eachplt, datemin=StartDate, datemax=EndDate)
Dateplot = True
Dateaxis = np.empty(totallength, dtype = 'datetime64[D]')
Dateaxis[0] = StartDate
for idate in range(1,totallength):
Dateaxis[idate] = Dateaxis[idate-1] + np.timedelta64(Dailyunit,'D')
else:
Dateplot = False
datemin = 0.0
datemax = maxplot
sumreal = 0.0
sumerror = 0.0
for itime in range(0,Num_Seq):
sumreal += abs(real[PlottedIndex,iValTrain,itime])
sumerror += error[PlottedIndex,iValTrain,itime]
c_error = round(100.0*sumerror/sumreal,2)
if UseRealDatesonplots:
eachplt.plot(Dateaxis[0:real.shape[-1]],real[PlottedIndex,iValTrain,:], label=f'real')
eachplt.plot(Dateaxis,predict[PlottedIndex,iValTrain,:], label='prediction')
eachplt.plot(Dateaxis[0:error.shape[-1]],error[PlottedIndex,iValTrain,:], label=f'error', color="red")
for others in range (0,otherlen):
eachplt.plot(Dateaxis[0:otherpredict.shape[-1]],otherpredict[others,PlottedIndex,iValTrain,:], label=otherlabs[others])
if fill:
eachplt.fill_between(Dateaxis[0:predictsmall.shape[-1]], predictsmall[PlottedIndex,iValTrain,:],
real[PlottedIndex,iValTrain,:], alpha=0.1, color="grey")
eachplt.fill_between(Dateaxis[0:error.shape[-1]], error[PlottedIndex,iValTrain,:], alpha=0.05, color="red")
else:
eachplt.plot(real[PlottedIndex,iValTrain,:], label=f'real')
eachplt.plot(predict[PlottedIndex,iValTrain,:], label='prediction')
eachplt.plot(error[PlottedIndex,iValTrain,:], label=f'error', color="red")
for others in range (0,otherlen):
eachplt.plot(otherpredict[others,PlottedIndex,iValTrain,:], label=otherlabs[others])
if fill:
eachplt.fill_between(xsmall, predictsmall[PlottedIndex,iValTrain,:], real[PlottedIndex,iValTrain,:],
alpha=0.1, color="grey")
eachplt.fill_between(xsmall, error[PlottedIndex,iValTrain,:], alpha=0.05, color="red")
if Earthquake and AddSpecialstoSummedplots:
if NumberValTrainLoops == 2:
if iValTrain == 0:
Addfixedearthquakes(eachplt, datemin, datemax, quakecolor = 'black', Dateplot = Dateplot,
vetoquake = PrimaryTrainingvetoquake)
Addfixedearthquakes(eachplt, datemin, datemax, quakecolor = 'purple', Dateplot = Dateplot,
vetoquake = SecondaryTrainingvetoquake)
else:
Addfixedearthquakes(eachplt, datemin, datemax, quakecolor = 'black', Dateplot = Dateplot,
vetoquake = PrimaryValidationvetoquake)
Addfixedearthquakes(eachplt, datemin, datemax, quakecolor = 'purple', Dateplot = Dateplot,
vetoquake = SecondaryValidationvetoquake)
else:
vetoquake = np.full(numberspecialeqs,False, dtype = bool)
Addfixedearthquakes(eachplt, datemin, datemax, quakecolor = 'black', Dateplot = Dateplot,
vetoquake = vetoquake)
extrastring = Overalllabel + current_time + ' ' + RunName + " "
extrastring += f"Length={Num_Seq}, Location Summed Results {Predictionbasicname[ActualObservable]}, "
yaxislabel = Predictionbasicname[ActualObservable]
if selectedfuture > 0:
yaxislabel = Predictionname[PredictedPos]
extrastring += " FUTURE " + yaxislabel
newyaxislabel = yaxislabel.replace("Months","Months\n")
newyaxislabel = newyaxislabel.replace("weeks","weeks\n")
newyaxislabel = newyaxislabel.replace("year","year\n")
eachplt.text(0.05,0.75,"FUTURE \n" + newyaxislabel,transform=eachplt.transAxes, color="red",fontsize=14, fontweight='bold')
extrastring += extracomments[PredictedPos][iValTrain]
eachplt.set_title('\n'.join(wrap(extrastring,70)))
if Dateplot:
eachplt.set_xlabel('Years')
else:
eachplt.set_xlabel(TimeIntervalUnitName+'s')
eachplt.set_ylabel(yaxislabel, color="red",fontweight='bold')
eachplt.grid(False)
eachplt.legend()
figure.tight_layout()
if Dumpplot and Dumpoutkeyplotsaspics:
VT = 'Both'
if NumberValTrainLoops == 1:
VT='Full'
SAVEFIG(plt, APPLDIR +'/Outputs/DLResults' + VT + str(PredictedPos) +RunName + '.png')
plt.show()
# Produce more detailed plots in time
# ONLY done for first quantity
splitsize = Plotsplitsize
if splitsize <= 1:
return
Numpoints = math.floor((Num_Seq+0.001)/splitsize)
extraone = Num_Seq%Numpoints
neededrows = math.floor((splitsize*NumberValTrainLoops +1.1)/2)
iValTrain = -1
PlottedIndex = 0
iseqnew = 0
counttimes = 0
for rowloop in range(0,neededrows):
plt.rcParams["figure.figsize"] = [16,6]
figure, (ax1,ax2) = plt.subplots(nrows=1, ncols=2)
for kplot in range (0,2):
if NumberValTrainLoops == 2:
iValTrain = kplot
else:
iValTrain = 0
Overalllabel = setValTrainlabel(iValTrain)
eachplt = ax1
if kplot == 1:
eachplt = ax2
sumreal = 0.0
sumerror = 0.0
if iValTrain == 0:
iseqold = iseqnew
iseqnew = iseqold + Numpoints
if counttimes < extraone:
iseqnew +=1
counttimes += 1
for itime in range(iseqold,iseqnew):
sumreal += abs(real[PlottedIndex,iValTrain,itime])
sumerror += error[PlottedIndex,iValTrain,itime]
c_error = round(100.0*sumerror/sumreal,2)
eachplt.plot(xsmall[iseqold:iseqnew],predict[PlottedIndex,iValTrain,iseqold:iseqnew], label='prediction')
eachplt.plot(xsmall[iseqold:iseqnew],real[PlottedIndex,iValTrain,iseqold:iseqnew], label=f'real')
eachplt.plot(xsmall[iseqold:iseqnew],error[PlottedIndex,iValTrain,iseqold:iseqnew], label=f'error', color="red")
if fill:
eachplt.fill_between(xsmall[iseqold:iseqnew], predictsmall[PlottedIndex,iValTrain,iseqold:iseqnew], real[PlottedIndex,iseqold:iseqnew], alpha=0.1, color="grey")
eachplt.fill_between(xsmall[iseqold:iseqnew], error[PlottedIndex,iValTrain,iseqold:iseqnew], alpha=0.05, color="red")
extrastring = Overalllabel + current_time + ' ' + RunName + " " + f"Range={iseqold}, {iseqnew} Rel Error {c_error} Location Summed Results {Predictionbasicname[PredictedPos]}, "
eachplt.set_title('\n'.join(wrap(extrastring,70)))
eachplt.set_xlabel(TimeIntervalUnitName+'s')
eachplt.set_ylabel(Predictionbasicname[PredictedPos])
eachplt.grid(True)
eachplt.legend()
figure.tight_layout()
plt.show()
def normalizeforplot(casesdeath,Locationindex,value):
if np.math.isnan(value):
return value
if Plotrealnumbers:
predaveragevaluespointer = PredictionAverageValuesPointer[casesdeath]
newvalue = value/QuantityStatistics[predaveragevaluespointer,2] + QuantityStatistics[predaveragevaluespointer,0]
rootflag = QuantityTakeroot[predaveragevaluespointer]
if rootflag == 2:
newvalue = newvalue**2
if rootflag == 3:
newvalue = newvalue**3
else:
newvalue = value
if PopulationNorm:
newvalue *= Locationpopulation[Locationindex]
return newvalue
# PLOT individual city data
def plot_by_fips(fips, Observations, FitPredictions, dots=True, fill=True):
Locationindex = FIPSintegerlookup[fips]
current_time = timenow()
print(startbold + startred + current_time + ' plot by location ' + str(Locationindex) + ' ' + str(fips) + ' ' + Locationname[Locationindex] + ' ' +RunName + ' ' + RunComment + resetfonts)
basiclength = Num_Seq
predictlength = LengthFutures
if not UseFutures:
predictlength = 0
totallength = basiclength + predictlength
real = np.zeros([NumpredbasicperTime,basiclength])
predictsmall = np.zeros([NumpredbasicperTime,basiclength])
predict = np.zeros([NumpredbasicperTime,totallength])
for PredictedQuantity in range(0,NumpredbasicperTime):
for itime in range (0,Num_Seq):
if np.math.isnan(Observations[itime, Locationindex, PredictedQuantity]):
Observations[itime, Locationindex, PredictedQuantity] = FitPredictions[itime, Locationindex, PredictedQuantity]
else:
real[PredictedQuantity,itime] = normalizeforplot(PredictedQuantity, Locationindex, Observations[itime, Locationindex, PredictedQuantity])
predict[PredictedQuantity,itime] = normalizeforplot(PredictedQuantity, Locationindex, FitPredictions[itime, Locationindex, PredictedQuantity])
if FuturedPointer[PredictedQuantity] >= 0:
for ifuture in range(0,LengthFutures):
jfuture = NumpredbasicperTime + NumpredFuturedperTime*ifuture
predict[PredictedQuantity,Num_Seq+ifuture] += normalizeforplot(PredictedQuantity,Locationindex,
FitPredictions[itime, Locationindex, FuturedPointer[PredictedQuantity] + jfuture])
for itime in range(0,basiclength):
predictsmall[PredictedQuantity,itime] = predict[PredictedQuantity,itime]
error = np.absolute(real - predictsmall)
xsmall = np.arange(0,Num_Seq)
neededrows = math.floor((NumpredbasicperTime +1.1)/2)
iplot = -1
for rowloop in range(0,neededrows):
plt.rcParams["figure.figsize"] = [16,6]
figure, (ax1,ax2) = plt.subplots(nrows=1, ncols=2)
for kplot in range (0,2):
iplot +=1
if iplot > (NumpredbasicperTime-1):
iplot = NumpredbasicperTime-1
eachplt = ax1
if kplot == 1:
eachplt = ax2
sumreal = 0.0
sumerror = 0.0
for itime in range(0,Num_Seq):
sumreal += abs(real[iplot,itime])
sumerror += error[iplot,itime]
c_error = round(100.0*sumerror/sumreal,2)
RMSEstring = ''
if not Plotrealnumbers:
sumRMSE = 0.0
count = 0.0
for itime in range(0,Num_Seq):
sumRMSE += (real[iplot,itime] - predict[iplot,itime])**2
count += 1.0
RMSE_error = round(100.0*sumRMSE/count,4)
RMSEstring = ' RMSE ' + str(RMSE_error)
x = list(range(0, totallength))
if dots:
eachplt.scatter(x, predict[iplot])
eachplt.scatter(xsmall, real[iplot])
eachplt.plot(predict[iplot], label=f'{fips} prediction')
eachplt.plot(real[iplot], label=f'{fips} real')
eachplt.plot(error[iplot], label=f'{fips} error', color="red")
if fill:
eachplt.fill_between(xsmall, predictsmall[iplot], real[iplot], alpha=0.1, color="grey")
eachplt.fill_between(xsmall, error[iplot], alpha=0.05, color="red")
name = Locationname[Locationindex]
if Plotrealnumbers:
name = "Actual Numbers " + name
stringpopulation = " "
if not Hydrology:
stringpopulation = " Population " +str(Locationpopulation[Locationindex])
titlestring = current_time + ' ' + RunName + f" {name}, Label={fips}" + stringpopulation + f" Length={Num_Seq}, Abs Rel Error={c_error}%" + RMSEstring + ' ' + RunName
eachplt.set_title('\n'.join(wrap(titlestring,70)))
eachplt.set_xlabel(TimeIntervalUnitName+'s')
eachplt.set_ylabel(Predictionbasicname[iplot])
eachplt.grid(True)
eachplt.legend()
figure.tight_layout()
plt.show();
def cumulative_error(real,predicted):
error = np.absolute(real-predicted).sum()
basevalue = np.absolute(real).sum()
return 100.0*error/basevalue
# Plot summed results by Prediction Type
# selectedfuture one more than usual future index
def plot_by_futureindex(selectedfuture, Observations, FitPredictions, fill=True, extrastring=''):
current_time = timenow()
print(startbold + startred + current_time + ' plot by Future Index ' + str(selectedfuture) + ' ' + RunName + ' ' + RunComment + resetfonts)
selectedfield = NumpredbasicperTime + NumpredFuturedperTime*(selectedfuture-1)
if selectedfuture == 0:
selectedfield = 0
real = np.zeros([NumpredFuturedperTime,Num_Seq])
predictsmall = np.zeros([NumpredFuturedperTime,Num_Seq])
validdata = 0
for PredictedQuantity in range(0,NumpredFuturedperTime):
for iloc in range(0,Nloc):
for itime in range (0,Num_Seq):
real[PredictedQuantity,itime] += Observations[itime, iloc, selectedfield+PredictedQuantity]
predictsmall[PredictedQuantity,itime] += FitPredictions[itime, iloc, selectedfield+PredictedQuantity]
for itime in range (0,Num_Seq):
if np.math.isnan(real[PredictedQuantity,itime]):
real[PredictedQuantity,itime] = predictsmall[PredictedQuantity,itime]
else:
if PredictedQuantity == 0:
validdata += 1
error = np.absolute(real - predictsmall)
xsmall = np.arange(0,Num_Seq)
neededrows = math.floor((NumpredFuturedperTime +1.1)/2)
iplot = -1
for rowloop in range(0,neededrows):
plt.rcParams["figure.figsize"] = [16,6]
figure, (ax1,ax2) = plt.subplots(nrows=1, ncols=2)
for kplot in range (0,2):
iplot +=1
if iplot > (NumpredbasicperTime-1):
iplot = NumpredbasicperTime-1
eachplt = ax1
if kplot == 1:
eachplt = ax2
sumreal = 0.0
sumerror = 0.0
for itime in range(0,Num_Seq):
sumreal += abs(real[iplot,itime])
sumerror += error[iplot,itime]
c_error = round(100.0*sumerror/sumreal,2)
eachplt.plot(predictsmall[iplot,:], label='prediction')
eachplt.plot(real[iplot,:], label=f'real')
eachplt.plot(error[iplot,:], label=f'error', color="red")
if fill:
eachplt.fill_between(xsmall, predictsmall[iplot,:], real[iplot,:], alpha=0.1, color="grey")
eachplt.fill_between(xsmall, error[iplot,:], alpha=0.05, color="red")
errorstring= " Error % " + str(c_error)
printstring = current_time + " Future Index " + str(selectedfuture) + " " + RunName
printstring += " " + f"Length={Num_Seq}, Location Summed Results {Predictionbasicname[iplot]}, " + errorstring + " " + extrastring
eachplt.set_title('\n'.join(wrap(printstring,70)))
eachplt.set_xlabel(TimeIntervalUnitName+'s')
eachplt.set_ylabel(Predictionbasicname[iplot])
eachplt.grid(True)
eachplt.legend()
figure.tight_layout()
plt.show()
# + [markdown] id="MVxWS_-p5T_N" pycharm={"name": "#%% md\n"}
# ### Calculate NNSE
#
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="dHr9p5LC5Z-k" outputId="9895ae52-c438-48b9-b5e9-f495cf1fb5c3" pycharm={"name": "#%%\n"}
# Calculate NNSE
# Sum (Obsevations - Mean)^2 / [Sum (Obsevations - Mean)^2 + Sum(Observations-Predictions)^2]
def FindNNSE(Observations, FitPredictions, Label=''):
NNSEList = np.empty(NpredperseqTOT, dtype = int)
NumberNNSEcalc = 0
for ipred in range(0,NpredperseqTOT):
if CalculateNNSE[ipred]:
NNSEList[NumberNNSEcalc] = ipred
NumberNNSEcalc +=1
if NumberNNSEcalc == 0:
return
StoreNNSE = np.zeros([Nloc,NumberNNSEcalc], dtype = np.float64)
basiclength = Num_Seq
current_time = timenow()
print(wraptotext(startbold + startred + current_time + ' Calculate NNSE ' + Label + ' ' +RunName + ' ' + RunComment + resetfonts))
for NNSEpredindex in range(0,NumberNNSEcalc):
PredictedQuantity = NNSEList[NNSEpredindex]
averageNNSE = 0.0
averageNNSETraining = 0.0
averageNNSEValidation = 0.0
line = ''
for Locationindex in range(0, Nloc):
QTObssq = 0.0
QTDiffsq = 0.0
QTObssum = 0.0
for itime in range (0,Num_Seq):
Observed = Observations[itime, Locationindex, PredictedQuantity]
if np.math.isnan(Observed):
Observed = FitPredictions[itime, Locationindex, PredictedQuantity]
real = normalizeforplot(PredictedQuantity, Locationindex, Observed)
predict = normalizeforplot(PredictedQuantity, Locationindex, FitPredictions[itime,
Locationindex, PredictedQuantity])
QTObssq += real**2
QTDiffsq += (real-predict)**2
QTObssum += real
Obsmeasure = QTObssq - (QTObssum**2 / Num_Seq )
StoreNNSE[Locationindex,NNSEpredindex] = Obsmeasure / (Obsmeasure +QTDiffsq )
if MappingtoTraining[Locationindex] >= 0:
averageNNSETraining += StoreNNSE[Locationindex,NNSEpredindex]
if MappingtoValidation[Locationindex] >= 0:
averageNNSEValidation += StoreNNSE[Locationindex,NNSEpredindex]
averageNNSE += StoreNNSE[Locationindex,NNSEpredindex]
line += str(round(StoreNNSE[Locationindex,NNSEpredindex],3)) + ' '
if ValidationNloc > 0:
averageNNSEValidation = averageNNSEValidation / ValidationNloc
averageNNSETraining = averageNNSETraining / TrainingNloc
averageNNSE = averageNNSE / Nloc
# Location Summed
QTObssq = 0.0
QTDiffsq = 0.0
QTObssum = 0.0
QTObssqT = 0.0
QTDiffsqT = 0.0
QTObssumT = 0.0
QTObssqV = 0.0
QTDiffsqV = 0.0
QTObssumV = 0.0
for itime in range (0,Num_Seq):
real = 0.0
predict = 0.0
realT = 0.0
predictT = 0.0
realV = 0.0
predictV = 0.0
for Locationindex in range(0, Nloc):
Observed = Observations[itime, Locationindex, PredictedQuantity]
if np.math.isnan(Observed):
Observed = FitPredictions[itime, Locationindex, PredictedQuantity]
localreal = normalizeforplot(PredictedQuantity, Locationindex, Observed)
localpredict = normalizeforplot(PredictedQuantity, Locationindex, FitPredictions[itime,
Locationindex, PredictedQuantity])
real += localreal
predict += localpredict
if MappingtoTraining[Locationindex] >= 0:
realT += localreal
predictT += localpredict
if MappingtoValidation[Locationindex] >= 0:
realV += localreal
predictV += localpredict
QTObssq += real**2
QTDiffsq += (real-predict)**2
QTObssum += real
QTObssqT += realT**2
QTDiffsqT += (realT-predictT)**2
QTObssumT += realT
QTObssqV += realV**2
QTDiffsqV += (realV-predictV)**2
QTObssumV += realV
Obsmeasure = QTObssq - (QTObssum**2 / Num_Seq )
SummedNNSE = Obsmeasure / (Obsmeasure +QTDiffsq )
ObsmeasureT = QTObssqT - (QTObssumT**2 / Num_Seq )
SummedNNSET = ObsmeasureT / (ObsmeasureT +QTDiffsqT )
ObsmeasureV = QTObssqV - (QTObssumV**2 / Num_Seq )
if ValidationNloc > 0:
SummedNNSEV = ObsmeasureV / (ObsmeasureV +QTDiffsqV )
else:
SummedNNSEV = 0.0
line = ''
if PredictedQuantity >= NumpredbasicperTime:
line = startred + 'Future ' + resetfonts
print(wraptotext(line + 'NNSE ' + startbold + Label + ' ' + str(PredictedQuantity) + ' ' + Predictionname[PredictedQuantity] + startred + ' Averaged ' +
str(round(averageNNSE,3)) + resetfonts + ' Training ' + str(round(averageNNSETraining,3)) +
' Validation ' + str(round(averageNNSEValidation,3)) + startred + startbold + ' Summed ' +
str(round(SummedNNSE,3)) + resetfonts + ' Training ' + str(round(SummedNNSET,3)) +
' Validation ' + str(round(SummedNNSEV,3)), size=200))
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="WOO6uzE1FUa1" outputId="71bffc4f-b39b-4dc5-b958-e3efa0358c9e" pycharm={"name": "#%%\n"}
def weightedcustom_lossGCF1(y_actual, y_pred, sample_weight):
tupl = np.shape(y_actual)
flagGCF = tf.math.is_nan(y_actual)
y_actual = y_actual[tf.math.logical_not(flagGCF)]
y_pred = y_pred[tf.math.logical_not(flagGCF)]
sw = sample_weight[tf.math.logical_not(flagGCF)]
tensordiff = tf.math.reduce_sum(tf.multiply(tf.math.square(y_actual-y_pred),sw))
if len(tupl) >= 2:
tensordiff /= tupl[0]
if len(tupl) >= 3:
tensordiff /= tupl[1]
if len(tupl) >= 4:
tensordiff /= tupl[2]
return tensordiff
def numpycustom_lossGCF1(y_actual, y_pred, sample_weight):
tupl = np.shape(y_actual)
flagGCF = np.isnan(y_actual)
y_actual = y_actual[np.logical_not(flagGCF)]
y_pred = y_pred[np.logical_not(flagGCF)]
sw = sample_weight[np.logical_not(flagGCF)]
tensordiff = np.sum(np.multiply(np.square(y_actual-y_pred),sw))
if len(tupl) >= 2:
tensordiff /= tupl[0]
if len(tupl) >= 3:
tensordiff /= tupl[1]
if len(tupl) >= 4:
tensordiff /= tupl[2]
return tensordiff
def weightedcustom_lossGCF1(y_actual, y_pred, sample_weight):
tupl = np.shape(y_actual)
flagGCF = tf.math.is_nan(y_actual)
y_actual = y_actual[tf.math.logical_not(flagGCF)]
y_pred = y_pred[tf.math.logical_not(flagGCF)]
sw = sample_weight[tf.math.logical_not(flagGCF)]
tensordiff = tf.math.reduce_sum(tf.multiply(tf.math.square(y_actual-y_pred),sw))
if len(tupl) >= 2:
tensordiff /= tupl[0]
if len(tupl) >= 3:
tensordiff /= tupl[1]
if len(tupl) >= 4:
tensordiff /= tupl[2]
return tensordiff
# + [markdown] id="zeDyzoVynCHL" pycharm={"name": "#%% md\n"}
# ### Custom Loss Functions
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="lJylkkL9AvsV" outputId="69dc04fe-2e2d-4664-d13c-6537b1498b2a" pycharm={"name": "#%%\n"}
def custom_lossGCF1(y_actual,y_pred):
tupl = np.shape(y_actual)
flagGCF = tf.math.is_nan(y_actual)
y_actual = y_actual[tf.math.logical_not(flagGCF)]
y_pred = y_pred[tf.math.logical_not(flagGCF)]
tensordiff = tf.math.reduce_sum(tf.math.square(y_actual-y_pred))
if len(tupl) >= 2:
tensordiff /= tupl[0]
if len(tupl) >= 3:
tensordiff /= tupl[1]
if len(tupl) >= 4:
tensordiff /= tupl[2]
return tensordiff
@tf.autograph.experimental.do_not_convert
def custom_lossGCF1spec(y_actual,y_pred):
global tensorsw
tupl = np.shape(y_actual)
flagGCF = tf.math.is_nan(y_actual)
y_actual = y_actual[tf.math.logical_not(flagGCF)]
y_pred = y_pred[tf.math.logical_not(flagGCF)]
sw = tensorsw[tf.math.logical_not(flagGCF)]
tensordiff = tf.math.reduce_sum(tf.multiply(tf.math.square(y_actual-y_pred),sw))
if len(tupl) >= 2:
tensordiff /= tupl[0]
if len(tupl) >= 3:
tensordiff /= tupl[1]
if len(tupl) >= 4:
tensordiff /= tupl[2]
return tensordiff
def custom_lossGCF1A(y_actual,y_pred):
print(np.shape(y_actual), np.shape(y_pred))
flagGCF = tf.math.is_nan(y_actual)
y_actual = y_actual[tf.math.logical_not(flagGCF)]
y_pred = y_pred[tf.math.logical_not(flagGCF)]
tensordiff = tf.math.square(y_actual-y_pred)
return tf.math.reduce_mean(tensordiff)
# Basic TF does NOT supply sample_weight
def custom_lossGCF1B(y_actual,y_pred,sample_weight=None):
tupl = np.shape(y_actual)
flagGCF = tf.math.is_nan(y_actual)
y_actual = y_actual[tf.math.logical_not(flagGCF)]
y_pred = y_pred[tf.math.logical_not(flagGCF)]
sw = sample_weight[tf.math.logical_not(flagGCF)]
tensordiff = tf.math.reduce_sum(tf.multiply(tf.math.square(y_actual-y_pred),sw))
if len(tupl) >= 2:
tensordiff /= tupl[0]
if len(tupl) >= 3:
tensordiff /= tupl[1]
if len(tupl) >= 4:
tensordiff /= tupl[2]
return tensordiff
def custom_lossGCF4(y_actual,y_pred):
tensordiff = y_actual-y_pred
newtensordiff = tf.where(tf.math.is_nan(tensordiff), tf.zeros_like(tensordiff), tensordiff)
return tf.math.reduce_mean(tf.math.square(newtensordiff))
# + [markdown] id="tIWDP9I8myNQ" pycharm={"name": "#%% md\n"}
# ### Utility: Shuffle, Finalize
#
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="OIB3yMlo7kFI" outputId="fd1c9512-c0d4-4bb8-e2c7-bc246d07be34" pycharm={"name": "#%%\n"}
def SetSpacetime(BasicTimes):
global GlobalTimeMask
Time = None
if (MaskingOption == 0) or (not GlobalSpacetime):
return Time
NumTOTAL = BasicTimes.shape[1]
BasicTimes = BasicTimes.astype(np.int16)
BasicTimes = np.reshape(BasicTimes,(BasicTimes.shape[0],NumTOTAL,1))
addons = np.arange(0,Tseq,dtype =np.int16)
addons = np.reshape(addons,(1,1,Tseq))
Time = BasicTimes+addons
Time = np.reshape(Time,(BasicTimes.shape[0], NumTOTAL*Tseq))
BasicPureTime = np.arange(0,Tseq,dtype =np.int16)
BasicPureTime = np.reshape(BasicPureTime,(Tseq,1))
GlobalTimeMask = tf.where( (BasicPureTime-np.transpose(BasicPureTime))>0, 0.0,1.0)
GlobalTimeMask = np.reshape(GlobalTimeMask,(1,1,1,Tseq,Tseq))
return Time
def shuffleDLinput(Xin,yin,AuxiliaryArray=None, Spacetime=None):
# Auxiliary array could be weight or location/time tracker
# These are per batch so sorted axis is first
np.random.seed(int.from_bytes(os.urandom(4), byteorder='little'))
trainingorder = list(range(0, len(Xin)))
random.shuffle(trainingorder)
Xinternal = list()
yinternal = list()
if AuxiliaryArray is not None:
AuxiliaryArrayinternal = list()
if Spacetime is not None:
Spacetimeinternal = list()
for i in trainingorder:
Xinternal.append(Xin[i])
yinternal.append(yin[i])
if AuxiliaryArray is not None:
AuxiliaryArrayinternal.append(AuxiliaryArray[i])
if Spacetime is not None:
Spacetimeinternal.append(Spacetime[i])
X = np.array(Xinternal)
y = np.array(yinternal)
if (AuxiliaryArray is None) and (Spacetime is None):
return X, y
if (AuxiliaryArray is not None) and (Spacetime is None):
AA = np.array(AuxiliaryArrayinternal)
return X,y,AA
if (AuxiliaryArray is None) and (Spacetime is not None):
St = np.array(Spacetimeinternal)
return X,y,St
AA = np.array(AuxiliaryArrayinternal)
St = np.array(Spacetimeinternal)
return X,y,AA,St
# Simple Plot of Loss from history
def finalizeDL(ActualModel, recordtrainloss, recordvalloss, validationfrac, X_in, y_in, modelflag, LabelFit =''):
# Ouput Loss v Epoch
histlen = len(recordtrainloss)
trainloss = recordtrainloss[histlen-1]
plt.rcParams["figure.figsize"] = [8,6]
plt.plot(recordtrainloss)
if (validationfrac > 0.001) and len(recordvalloss) > 0:
valloss = recordvalloss[histlen-1]
plt.plot(recordvalloss)
else:
valloss = 0.0
current_time = timenow()
print(startbold + startred + current_time + ' ' + RunName + ' finalizeDL ' + RunComment +resetfonts)
plt.title(LabelFit + ' ' + RunName+' model loss ' + str(round(trainloss,7)) + ' Val ' + str(round(valloss,7)))
plt.ylabel('loss')
plt.xlabel('epoch')
plt.yscale("log")
plt.grid(True)
plt.legend(['train', 'val'], loc='upper left')
plt.show()
# Setup TFT
if modelflag == 2:
global SkipDL2F, IncreaseNloc_sample, DecreaseNloc_sample
SkipDL2F = True
IncreaseNloc_sample = 1
DecreaseNloc_sample = 1
TFToutput_map = TFTpredict(TFTmodel,TFTtest_datacollection)
VisualizeTFT(TFTmodel, TFToutput_map)
else:
FitPredictions = DLprediction(X_in, y_in,ActualModel,modelflag, LabelFit = LabelFit)
for debugfips in ListofTestFIPS:
if debugfips != '':
debugfipsoutput(debugfips, FitPredictions, X_in, y_in)
return
def debugfipsoutput(debugfips, FitPredictions, Xin, Observations):
print(startbold + startred + 'debugfipsoutput for ' + str(debugfips) + RunName + ' ' + RunComment +resetfonts)
# Set Location Number in Arrays
LocationNumber = FIPSstringlookup[debugfips]
# Sequences to look at
Seqcount = 5
Seqnumber = np.empty(Seqcount, dtype = int)
Seqnumber[0] = 0
Seqnumber[1] = int(Num_Seq/4)-1
Seqnumber[2] = int(Num_Seq/2)-1
Seqnumber[3] = int((3*Num_Seq)/4) -1
Seqnumber[4] = Num_Seq-1
# Window Positions to look at
Wincount = 5
Winnumber = np.empty(Wincount, dtype = int)
Winnumber[0] = 0
Winnumber[1] = int(Tseq/4)-1
Winnumber[2] = int(Tseq/2)-1
Winnumber[3] = int((3*Tseq)/4) -1
Winnumber[4] = Tseq-1
if SymbolicWindows:
InputSequences = np.empty([Seqcount,Wincount, NpropperseqTOT], dtype=np.float32)
for jseq in range(0,Seqcount):
iseq = Seqnumber[jseq]
for jwindow in range(0,Wincount):
window = Winnumber[jwindow]
InputSequences[jseq,jwindow] = Xin[LocationNumber,iseq+jseq]
else:
InputSequences = Xin
# Location Info
print('\n' + startbold + startred + debugfips + ' # ' + str(LocationNumber) + ' ' +
Locationname[LocationNumber] + ' ' + Locationstate[LocationNumber] + ' Pop '
+ str(Locationpopulation[LocationNumber]) + resetfonts)
plot_by_fips(int(debugfips), Observations, FitPredictions)
if PlotsOnlyinTestFIPS:
return
# Print Input Data to Test
# Static Properties
print(startbold + startred + 'Static Properties ' + debugfips + ' ' +
Locationname[LocationNumber] + resetfonts)
line = ''
for iprop in range(0,NpropperTimeStatic):
if SymbolicWindows:
val = InputSequences[0,0,iprop]
else:
val = InputSequences[0,LocationNumber,0,iprop]
line += startbold + InputPropertyNames[PropertyNameIndex[iprop]] + resetfonts + ' ' + str(round(val,3)) + ' '
print('\n'.join(wrap(line,200)))
# Dynamic Properties
for iprop in range(NpropperTimeStatic, NpropperTime):
print('\n')
for jwindow in range(0,Wincount):
window = Winnumber[jwindow]
line = startbold + InputPropertyNames[PropertyNameIndex[iprop]] + ' W= '+str(window) +resetfonts
for jseq in range(0,Seqcount):
iseq = Seqnumber[jseq]
line += startbold + startred + ' ' + str(iseq) + ')' +resetfonts
if SymbolicWindows:
val = InputSequences[jseq,jwindow,iprop]
else:
val = InputSequences[iseq,LocationNumber,window,iprop]
line += ' ' + str(round(val,3))
print('\n'.join(wrap(line,200)))
# Total Input
print('\n')
line = startbold + 'Props: ' + resetfonts
for iprop in range(0,NpropperseqTOT):
if iprop%5 == 0:
line += startbold + startred + ' ' + str(iprop) + ')' + resetfonts
line += ' ' + InputPropertyNames[PropertyNameIndex[iprop]]
print('\n'.join(wrap(line,200)))
for jseq in range(0,Seqcount):
iseq = Seqnumber[jseq]
for jwindow in range(0,Wincount):
window = Winnumber[jwindow]
line = startbold + 'Input: All in Seq ' + str(iseq) + ' W= ' + str(window) + resetfonts
for iprop in range(0,NpropperseqTOT):
if iprop%5 == 0:
line += startbold + startred + ' ' + str(iprop) + ')' +resetfonts
if SymbolicWindows:
val = InputSequences[jseq,jwindow,iprop]
else:
val = InputSequences[iseq,LocationNumber,window,iprop]
result = str(round(val,3))
line += ' ' + result
print('\n'.join(wrap(line,200)))
# Total Prediction
print('\n')
line = startbold + 'Preds: ' + resetfonts
for ipred in range(0,NpredperseqTOT):
if ipred%5 == 0:
line += startbold + startred + ' ' + str(ipred) + ')' + resetfonts
line += ' ' + Predictionname[ipred]
for jseq in range(0,Seqcount):
iseq = Seqnumber[jseq]
line = startbold + 'Preds: All in Seq ' + str(iseq) + resetfonts
for ipred in range(0,NpredperseqTOT):
fred = Observations[iseq,LocationNumber,ipred]
if np.math.isnan(fred):
result = 'NaN'
else:
result = str(round(fred,3))
if ipred%5 == 0:
line += startbold + startred + ' ' + str(ipred) + ')' + resetfonts
line += ' ' + result
print('\n'.join(wrap(line,200)))
# + [markdown] id="HaflIIrbKjRJ" pycharm={"name": "#%% md\n"}
# ### DLPrediction2E printloss ?DEL
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="1vuo893BKprz" outputId="86581fb0-6d7f-43a1-b565-9f18d2609242" pycharm={"name": "#%%\n"}
def printloss(name,mean,var,SampleSize, lineend =''):
mean /= SampleSize
var /= SampleSize
std = math.sqrt(var - mean**2)
print(name + ' Mean ' + str(round(mean,5)) + ' Std Deviation ' + str(round(std,7)) + ' ' + lineend)
def DLprediction2E(Xin, yin, DLmodel, modelflag):
# Form restricted Attention separately over Training and Validation
if not LocationBasedValidation:
return
if UsedTransformervalidationfrac < 0.001 or ValidationNloc <= 0:
return
if SkipDL2E:
return
if GarbageCollect:
gc.collect()
SampleSize = 1
FitRanges_PartialAtt = np.zeros([Num_Seq, Nloc, NpredperseqTOT,5], dtype =np.float32)
FRanges = np.full(NpredperseqTOT, 1.0, dtype = np.float32)
# 0 count 1 mean 2 Standard Deviation 3 Min 4 Max
print(wraptotext(startbold+startred+ 'DLPrediction2E Partial Attention ' +current_time + ' ' + RunName + RunComment + resetfonts))
global OuterBatchDimension, Nloc_sample, d_sample, max_d_sample
global FullSetValidation
saveFullSetValidation = FullSetValidation
FullSetValidation = False
X_predict, y_predict, Spacetime_predict, X_val, y_val, Spacetime_val = setSeparateDLinput(1, Spacetime = True)
FullSetValidation = saveFullSetValidation
Nloc_sample = TrainingNloc
OuterBatchDimension = Num_Seq
d_sample = Tseq * TrainingNloc
max_d_sample = d_sample
UsedValidationNloc = ValidationNloc
if SymbolicWindows:
X_Transformertraining = np.reshape(X_predict, (OuterBatchDimension, Nloc_sample))
else:
X_Transformertraining = np.reshape(X_predict, (OuterBatchDimension, d_sample, NpropperseqTOT))
y_Transformertraining = np.reshape(y_predict, (OuterBatchDimension, Nloc_sample, NpredperseqTOT))
Spacetime_Transformertraining = np.reshape(Spacetime_predict, (OuterBatchDimension, Nloc_sample))
if SymbolicWindows:
X_Transformerval = np.reshape(X_val, (OuterBatchDimension, UsedValidationNloc))
else:
X_Transformerval = np.reshape(X_val, (OuterBatchDimension, UsedValidationNloc*Tseq, NpropperseqTOT))
y_Transformerval = np.reshape(y_val, (OuterBatchDimension, UsedValidationNloc, NpredperseqTOT))
Spacetime_Transformerval = np.reshape(Spacetime_val, (OuterBatchDimension, UsedValidationNloc))
if UseClassweights:
sw_Transformertraining = np.empty_like(y_predict, dtype=np.float32)
for i in range(0,sw_Transformertraining.shape[0]):
for j in range(0,sw_Transformertraining.shape[1]):
for k in range(0,NpredperseqTOT):
sw_Transformertraining[i,j,k] = Predictionwgt[k]
sw_Transformerval = np.empty_like(y_val, dtype=np.float32)
for i in range(0,sw_Transformerval.shape[0]):
for jloc in range(0,sw_Transformerval.shape[1]):
for k in range(0,NpredperseqTOT):
sw_Transformerval[i,jloc,k] = Predictionwgt[k]
else:
sw_Transformertraining = []
sw_Transformerval = []
if SymbolicWindows:
X_Transformertrainingflat2 = np.reshape(X_Transformertraining, (-1, TrainingNloc))
X_Transformertrainingflat1 = np.reshape(X_Transformertrainingflat2, (-1))
else:
X_Transformertrainingflat2 = np.reshape(X_Transformertraining, (-1, TrainingNloc,Tseq, NpropperseqTOT))
X_Transformertrainingflat1 = np.reshape(X_Transformertrainingflat2, (-1, Tseq, NpropperseqTOT))
y_Transformertrainingflat1 = np.reshape(y_Transformertraining, (-1,NpredperseqTOT) )
Spacetime_Transformertrainingflat1 = np.reshape(Spacetime_Transformertraining,(-1))
if UseClassweights:
sw_Transformertrainingflat1 = np.reshape(sw_Transformertraining, (-1,NpredperseqTOT) )
if SymbolicWindows:
X_Transformervalflat2 = np.reshape(X_Transformerval, (-1, UsedValidationNloc))
X_Transformervalflat1 = np.reshape(X_Transformervalflat2, (-1))
else:
X_Transformervalflat2 = np.reshape(X_Transformerval, (-1, UsedValidationNloc,Tseq, NpropperseqTOT))
X_Transformervalflat1 = np.reshape(X_Transformervalflat2, (-1, Tseq, NpropperseqTOT))
y_Transformervalflat1 = np.reshape(y_Transformerval, (-1,NpredperseqTOT) )
Spacetime_Transformervalflat1 = np.reshape(Spacetime_Transformerval,(-1))
if UseClassweights:
sw_Transformervalflat1 = np.reshape(sw_Transformerval, (-1,NpredperseqTOT) )
meanvalue2 = 0.0
meanvalue3 = 0.0
meanvalue4 = 0.0
variance2= 0.0
variance3= 0.0
variance4= 0.0
# START LOOP OVER SAMPLES
samplebar = notebook.trange(SampleSize, desc='Full Samples', unit = 'sample')
epochsize = 2*OuterBatchDimension
if IncreaseNloc_sample > 1:
epochsize = int(epochsize/IncreaseNloc_sample)
elif DecreaseNloc_sample > 1:
epochsize = int(epochsize*DecreaseNloc_sample)
bbar = notebook.trange(epochsize, desc='Batch loop', unit = 'sample')
for shuffling in range (0,SampleSize):
if GarbageCollect:
gc.collect()
# TRAINING SET
if TimeShufflingOnly:
X_train, y_train, sw_train, Spacetime_train = shuffleDLinput(X_Transformertraining,
y_Transformertraining, sw_Transformertraining, Spacetime = Spacetime_Transformertraining)
else:
X_train, y_train, sw_train, Spacetime_train = shuffleDLinput(X_Transformertrainingflat1,
y_Transformertrainingflat1, sw_Transformertrainingflat1, Spacetime = Spacetime_Transformertrainingflat1)
Nloc_sample = TrainingNloc
OuterBatchDimension = Num_Seq
Totaltodo = Nloc_sample*OuterBatchDimension
if IncreaseNloc_sample > 1:
Nloc_sample = int(Nloc_sample*IncreaseNloc_sample)
elif DecreaseNloc_sample > 1:
Nloc_sample = int(Nloc_sample/DecreaseNloc_sample)
OuterBatchDimension = int(Totaltodo/Nloc_sample)
if OuterBatchDimension * Nloc_sample != Totaltodo:
printexit('Inconsistent Nloc_sample ' + str(Nloc_sample))
d_sample = Tseq * Nloc_sample
max_d_sample = d_sample
if SymbolicWindows:
X_train = np.reshape(X_train, (OuterBatchDimension, Nloc_sample))
else:
X_train = np.reshape(X_train, (OuterBatchDimension, d_sample, NpropperseqTOT))
y_train = np.reshape(y_train, (OuterBatchDimension, Nloc_sample, NpredperseqTOT))
sw_train = np.reshape(sw_train, (OuterBatchDimension, Nloc_sample, NpredperseqTOT))
Spacetime_train = np.reshape(Spacetime_train, (OuterBatchDimension, Nloc_sample))
quan3 = 0.0
quan4 = 0.0
losspercallVl = 0.0
losspercallTr = 0.0
TotalTr = 0.0
TotalVl = 0.0
for Trainingindex in range(0, OuterBatchDimension):
if GarbageCollect:
gc.collect()
X_trainlocal = X_train[Trainingindex]
if SymbolicWindows:
X_trainlocal = np.reshape(X_trainlocal,[1,X_trainlocal.shape[0]])
else:
X_trainlocal = np.reshape(X_trainlocal,[1,X_trainlocal.shape[0],X_trainlocal.shape[1]])
Numinbatch = X_trainlocal.shape[0]
NuminAttention = X_trainlocal.shape[1]
NumTOTAL = Numinbatch*NuminAttention
# SymbolicWindows X_train is indexed by Batch index, Location List for Attention. Missing 1(replace by Window), 1 (replace by properties)
if SymbolicWindows:
X_trainlocal = np.reshape(X_trainlocal,NumTOTAL)
iseqarray = np.right_shift(X_trainlocal,16)
ilocarray = np.bitwise_and(X_trainlocal, 0b1111111111111111)
X_train_withSeq = list()
for iloc in range(0,NumTOTAL):
X_train_withSeq.append(ReshapedSequencesTOT[ilocarray[iloc],iseqarray[iloc]:iseqarray[iloc]+Tseq])
X_train_withSeq = np.array(X_train_withSeq)
X_train_withSeq = np.reshape(X_train_withSeq,(Numinbatch, d_sample, NpropperseqTOT))
Time = None
if modelflag==1:
Time = SetSpacetime(np.reshape(iseqarray,[Numinbatch,-1]))
PredictedVector = DLmodel(X_train_withSeq, training = PredictionTraining, Time=Time )
else:
Spacetime_trainlocal = Spacetime_train[Trainingindex]
iseqarray = np.right_shift(Spacetime_trainlocal,16)
ilocarray = np.bitwise_and(Spacetime_trainlocal, 0b1111111111111111)
Time = SetSpacetime(np.reshape(iseqarray,[Numinbatch,-1]))
PredictedVector = DLmodel(X_trainlocal, training = PredictionTraining, Time=Time )
PredictedVector = np.reshape(PredictedVector,(1,Nloc_sample,NpredperseqTOT))
TrueVector = y_train[Trainingindex]
TrueVector = np.reshape(TrueVector,(1,Nloc_sample,NpredperseqTOT))
sw_trainlocal = sw_train[Trainingindex]
sw_trainlocal = np.reshape(sw_trainlocal,[1,sw_trainlocal.shape[0],sw_trainlocal.shape[1]])
losspercallTr = numpycustom_lossGCF1(TrueVector,PredictedVector,sw_trainlocal)
quan3 += losspercallTr
for iloc_sample in range(0,Nloc_sample):
LocLocal = ilocarray[iloc_sample]
SeqLocal = iseqarray[iloc_sample]
yyhat = PredictedVector[0,iloc_sample]
if FitRanges_PartialAtt [SeqLocal, LocLocal, 0, 0] < 0.1:
FitRanges_PartialAtt [SeqLocal,LocLocal,:,3] = yyhat
FitRanges_PartialAtt [SeqLocal,LocLocal,:,4] = yyhat
else:
FitRanges_PartialAtt [SeqLocal,LocLocal,:,3] = np.maximum(FitRanges_PartialAtt[SeqLocal,LocLocal,:,3],yyhat)
FitRanges_PartialAtt [SeqLocal,LocLocal,:,4] = np.minimum(FitRanges_PartialAtt[SeqLocal,LocLocal,:,4],yyhat)
FitRanges_PartialAtt [SeqLocal,LocLocal,:,0] += FRanges
FitRanges_PartialAtt[SeqLocal,LocLocal,:,1] += yyhat
FitRanges_PartialAtt[SeqLocal,LocLocal,:,2] += np.square(yyhat)
fudge = 1.0/(1+Trainingindex)
TotalTr = quan3 *fudge
bbar.set_postfix(TotalTr = TotalTr, Tr = losspercallTr)
bbar.update(Transformerbatch_size)
# END Training Batch Loop
TotalTr= quan3/OuterBatchDimension
# VALIDATION SET
Nloc_sample = UsedValidationNloc
OuterBatchDimension = Num_Seq
Totaltodo = Nloc_sample*OuterBatchDimension
if IncreaseNloc_sample > 1:
Nloc_sample = int(Nloc_sample*IncreaseNloc_sample)
elif DecreaseNloc_sample > 1:
Nloc_sample = int(Nloc_sample/DecreaseNloc_sample)
OuterBatchDimension = int(Totaltodo/Nloc_sample)
if OuterBatchDimension * Nloc_sample != Totaltodo:
printexit('Inconsistent Nloc_sample ' + str(Nloc_sample))
d_sample = Tseq * Nloc_sample
max_d_sample = d_sample
if TimeShufflingOnly:
X_val, y_val, sw_val, Spacetime_val = shuffleDLinput(
X_Transformerval, y_Transformerval, sw_Transformerval, Spacetime_Transformerval)
else:
X_val, y_val, sw_val, Spacetime_val = shuffleDLinput(
X_Transformervalflat1, y_Transformervalflat1, sw_Transformervalflat1, Spacetime_Transformervalflat1)
if SymbolicWindows:
X_val = np.reshape(X_val, (OuterBatchDimension, Nloc_sample))
else:
X_val = np.reshape(X_val, (OuterBatchDimension, d_sample, NpropperseqTOT))
y_val = np.reshape(y_val, (OuterBatchDimension, Nloc_sample, NpredperseqTOT))
sw_val = np.reshape(sw_val, (OuterBatchDimension, Nloc_sample, NpredperseqTOT))
Spacetime_val = np.reshape(Spacetime_val, (OuterBatchDimension, Nloc_sample))
# START VALIDATION Batch Loop
for Validationindex in range(0,OuterBatchDimension):
X_valbatch = X_val[Validationindex]
y_valbatch = y_val[Validationindex]
sw_valbatch = sw_val[Validationindex]
Spacetime_valbatch = Spacetime_val[Validationindex]
if SymbolicWindows:
X_valbatch = np.reshape(X_valbatch,[1,X_valbatch.shape[0]])
else:
X_valbatch = np.reshape(X_valbatch,[1,X_valbatch.shape[0],X_valbatch.shape[1]])
y_valbatch = np.reshape(y_valbatch,[1,y_valbatch.shape[0],y_valbatch.shape[1]])
sw_valbatch = np.reshape(sw_valbatch,[1,sw_valbatch.shape[0],sw_valbatch.shape[1]])
Numinbatch = X_valbatch.shape[0]
NuminAttention = X_valbatch.shape[1]
NumTOTAL = Numinbatch*NuminAttention
if SymbolicWindows:
X_valbatch = np.reshape(X_valbatch,NumTOTAL)
iseqarray = np.right_shift(X_valbatch,16)
ilocarray = np.bitwise_and(X_valbatch, 0b1111111111111111)
X_valbatch_withSeq = list()
for iloc in range(0,NumTOTAL):
X_valbatch_withSeq.append(ReshapedSequencesTOT[ilocarray[iloc],iseqarray[iloc]:iseqarray[iloc]+Tseq])
X_valbatch_withSeq = np.array(X_valbatch_withSeq)
X_valbatch_withSeq = np.reshape(X_valbatch_withSeq,(Numinbatch, d_sample, NpropperseqTOT))
Time = SetSpacetime(np.reshape(iseqarray,[Numinbatch,-1]))
PredictedVector = DLmodel(X_valbatch_withSeq, training = PredictionTraining, Time=Time )
else:
Spacetime_valbatch = np.reshape(Spacetime_valbatch,-1)
iseqarray = np.right_shift(Spacetime_valbatch,16)
ilocarray = np.bitwise_and(Spacetime_valbatch, 0b1111111111111111)
Time = SetSpacetime(np.reshape(iseqarray,[Numinbatch,-1]))
PredictedVector = DLmodel(X_valbatch, training = PredictionTraining, Time=Time )
PredictedVector = np.reshape(PredictedVector,(1,Nloc_sample,NpredperseqTOT))
TrueVector = np.reshape(y_valbatch,(1,Nloc_sample,NpredperseqTOT))
sw_valbatch = np.reshape(sw_valbatch,(1,Nloc_sample,NpredperseqTOT))
losspercallVl = numpycustom_lossGCF1(TrueVector,PredictedVector,sw_valbatch)
quan4 += losspercallVl
for iloc_sample in range(0,Nloc_sample):
LocLocal = ilocarray[iloc_sample]
SeqLocal = iseqarray[iloc_sample]
yyhat = PredictedVector[0,iloc_sample]
if FitRanges_PartialAtt [SeqLocal, LocLocal, 0, 0] < 0.1:
FitRanges_PartialAtt [SeqLocal,LocLocal,:,3] = yyhat
FitRanges_PartialAtt [SeqLocal,LocLocal,:,4] = yyhat
else:
FitRanges_PartialAtt [SeqLocal,LocLocal,:,3] = np.maximum(FitRanges_PartialAtt[SeqLocal,LocLocal,:,3],yyhat)
FitRanges_PartialAtt [SeqLocal,LocLocal,:,4] = np.minimum(FitRanges_PartialAtt[SeqLocal,LocLocal,:,4],yyhat)
FitRanges_PartialAtt [SeqLocal,LocLocal,:,0] += FRanges
FitRanges_PartialAtt[SeqLocal,LocLocal,:,1] += yyhat
FitRanges_PartialAtt[SeqLocal,LocLocal,:,2] += np.square(yyhat)
TotalVl = quan4/(1+Validationindex)
losspercall = (TotalTr*TrainingNloc+TotalVl*ValidationNloc)/Nloc
bbar.update(Transformerbatch_size)
bbar.set_postfix(Loss = losspercall, TotalTr = TotalTr, TotalVl= TotalVl, Vl = losspercallVl)
# END VALIDATION BATCH LOOP
# Processing at the end of Sampling Loop
fudge = 1.0/OuterBatchDimension
quan2 = (quan3*TrainingNloc + quan4*ValidationNloc)/Nloc
quan2 *= fudge
meanvalue2 += quan2
variance2 += quan2**2
if LocationBasedValidation:
quan3 *= fudge
quan4 *= fudge
meanvalue3 += quan3
meanvalue4 += quan4
variance3 += quan3**2
variance4 += quan4**2
samplebar.update(1)
if LocationBasedValidation:
samplebar.set_postfix(Shuffle=shuffling, Loss = quan2, Tr = quan3, Val = quan4)
else:
samplebar.set_postfix(Shuffle=shuffling, Loss = quan2)
bbar.reset()
# End Shuffling loop
printloss(' Full Loss ',meanvalue2,variance2,SampleSize)
printloss(' Training Loss ',meanvalue3,variance3,SampleSize)
printloss(' Validation Loss ',meanvalue4,variance4,SampleSize)
global GlobalTrainingLoss, GlobalValidationLoss, GlobalLoss
GlobalLoss = meanvalue2
GlobalTrainingLoss = meanvalue3
GlobalValidationLoss = meanvalue4
FitRanges_PartialAtt[:,:,:,1] = np.divide(FitRanges_PartialAtt[:,:,:,1],FitRanges_PartialAtt[:,:,:,0])
FitRanges_PartialAtt[:,:,:,2] = np.sqrt(np.maximum(np.divide(FitRanges_PartialAtt[:,:,:,2],FitRanges_PartialAtt[:,:,:,0]) -
np.square(FitRanges_PartialAtt[:,:,:,1]), 0.0))
FitPredictions = np.zeros([Num_Seq, Nloc, NpredperseqTOT], dtype =np.float32)
for iseq in range(0,Num_Seq):
for iloc in range(0,Nloc):
FitPredictions[iseq,iloc,:] = FitRanges_PartialAtt[iseq,iloc,:,1]
DLprediction3(yin, FitPredictions, ' Separate Attention mean values')
FindNNSE(yin, FitPredictions, Label='Separate Attention' )
print(startbold+startred+ 'END DLPrediction2E ' +current_time + ' ' + RunName + RunComment +resetfonts)
return
# + [markdown] id="pPM9420zCDSO" pycharm={"name": "#%% md\n"}
# ### DLPrediction2F Sensitivity
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="_bv4pFAeCXZu" outputId="6b4b51e7-94cc-4d35-ebc9-75dc4c6ff360" pycharm={"name": "#%%\n"}
def DLprediction2F(Xin, yin, DLmodel, modelflag):
# Input is the windows [Num_Seq] [Nloc] [Tseq] [NpropperseqTOT] (SymbolicWindows False)
# Input is the sequences [Nloc] [Num_Time-1] [NpropperseqTOT] (SymbolicWindows True)
# Input Predictions are always [Num_Seq] [NLoc] [NpredperseqTOT]
# Label Array is always [Num_Seq][Nloc] [0=Window(first sequence)#, 1=Location]
if SkipDL2F:
return
if GarbageCollect:
gc.collect()
global OuterBatchDimension, Nloc_sample, d_sample, max_d_sample
SensitivityAnalyze = np.full((NpropperseqTOT), False, dtype = bool)
SensitivityChange = np.zeros ((NpropperseqTOT), dtype = np.float32)
SensitvitybyPrediction = False
something = 0
SensitivityList = []
for iprop in range(0,NpropperseqTOT):
if SensitivityAnalyze[iprop]:
something +=1
SensitivityList.append(iprop)
if something == 0:
return
ScaleProperty = 0.99
SampleSize = 1
SensitivityFitPredictions = np.zeros([Num_Seq, Nloc, NpredperseqTOT, 1 + something], dtype =np.float32)
FRanges = np.full((NpredperseqTOT), 1.0, dtype = np.float32)
current_time = timenow()
print(wraptotext(startbold+startred+ 'DLPrediction2F ' +current_time + ' ' + RunName + RunComment + resetfonts))
sw = np.empty_like(yin, dtype=np.float32)
for i in range(0,sw.shape[0]):
for j in range(0,sw.shape[1]):
for k in range(0,NpredperseqTOT):
sw[i,j,k] = Predictionwgt[k]
labelarray =np.empty([Num_Seq, Nloc, 2], dtype = np.int32)
for iseq in range(0, Num_Seq):
for iloc in range(0,Nloc):
labelarray[iseq,iloc,0] = iseq
labelarray[iseq,iloc,1] = iloc
Totaltodo = Num_Seq*Nloc
Nloc_sample = Nloc # default
if IncreaseNloc_sample > 1:
Nloc_sample = int(Nloc_sample*IncreaseNloc_sample)
elif DecreaseNloc_sample > 1:
Nloc_sample = int(Nloc_sample/DecreaseNloc_sample)
if Totaltodo%Nloc_sample != 0:
printexit('Invalid Nloc_sample ' + str(Nloc_sample) + " " + str(Totaltodo))
d_sample = Tseq * Nloc_sample
max_d_sample = d_sample
OuterBatchDimension = int(Totaltodo/Nloc_sample)
print(' Predict with ' +str(Nloc_sample) + ' sequences per sample and batch size ' + str(OuterBatchDimension))
print(startbold+startred+ 'Sensitivity using Property ScaleFactor ' + str(round(ScaleProperty,3)) + resetfonts)
for Sensitivities in range(0,1+something):
if Sensitivities == 0: # BASIC unmodified run
iprop = -1
print(startbold+startred+ 'Basic Predictions' + resetfonts)
if SymbolicWindows:
ReshapedSequencesTOTmodified = ReshapedSequencesTOT # NOT used if modelflag == 2
if modelflag == 2:
DLmodel.MakeMapping()
else:
Xinmodified = Xin
else:
iprop = SensitivityList[Sensitivities-1]
maxminplace = PropertyNameIndex[iprop]
lastline = ''
if iprop < Npropperseq:
lastline = ' Normed Mean ' +str(round(QuantityStatistics[maxminplace,5],4))
print(startbold+startred+ 'Property ' + str(iprop) + ' ' + InputPropertyNames[maxminplace] + resetfonts + lastline)
if SymbolicWindows:
if modelflag == 2:
DLmodel.SetupProperty(iprop)
DLmodel.ScaleProperty(ScaleProperty)
DLmodel.MakeMapping()
else:
ReshapedSequencesTOTmodified = np.copy(ReshapedSequencesTOT)
ReshapedSequencesTOTmodified[:,:,iprop] = ScaleProperty * ReshapedSequencesTOTmodified[:,:,iprop]
else:
Xinmodified = np.copy(Xin)
Xinmodified[:,:,:,iprop] = ScaleProperty*Xinmodified[:,:,:,iprop]
CountFitPredictions = np.zeros([Num_Seq, Nloc, NpredperseqTOT], dtype =np.float32)
meanvalue2 = 0.0
meanvalue3 = 0.0
meanvalue4 = 0.0
variance2= 0.0
variance3= 0.0
variance4= 0.0
samplebar = notebook.trange(SampleSize, desc='Full Samples', unit = 'sample')
bbar = notebook.trange(OuterBatchDimension, desc='Batch loop', unit = 'sample')
for shuffling in range (0,SampleSize):
if GarbageCollect:
gc.collect()
yuse = yin
labeluse = labelarray
y2= np.reshape(yuse, (-1, NpredperseqTOT)).copy()
labelarray2 = np.reshape(labeluse, (-1,2))
if SymbolicWindows:
# Xin X2 X3 not used rather ReshapedSequencesTOT
labelarray2, y2 = shuffleDLinput(labelarray2, y2)
ReshapedSequencesTOTuse = ReshapedSequencesTOTmodified
else:
Xuse = Xinmodified
X2 = np.reshape(Xuse, (-1, Tseq, NpropperseqTOT)).copy()
X2, y2, labelarray2 = shuffleDLinput(X2, y2,labelarray2)
X3 = np.reshape(X2, (-1, d_sample, NpropperseqTOT))
y3 = np.reshape(y2, (-1, Nloc_sample, NpredperseqTOT))
sw = np.reshape(sw, (-1, Nloc_sample, NpredperseqTOT))
labelarray3 = np.reshape(labelarray2, (-1, Nloc_sample, 2))
quan2 = 0.0
quan3 = 0.0
quan4 = 0.0
for Batchindex in range(0, OuterBatchDimension):
if GarbageCollect:
gc.collect()
if SymbolicWindows:
if modelflag == 2: # Note first index of InputVector Location, Second is sequence number; labelarray3 is opposite
InputVector = np.empty((Nloc_sample,2), dtype = np.int32)
for iloc_sample in range(0,Nloc_sample):
InputVector[iloc_sample,0] = labelarray3[Batchindex, iloc_sample,1]
InputVector[iloc_sample,1] = labelarray3[Batchindex, iloc_sample,0]
else:
X3local = list()
for iloc_sample in range(0,Nloc_sample):
LocLocal = labelarray3[Batchindex, iloc_sample,1]
SeqLocal = labelarray3[Batchindex, iloc_sample,0]
X3local.append(ReshapedSequencesTOTuse[LocLocal,SeqLocal:SeqLocal+Tseq])
InputVector = np.array(X3local)
else:
InputVector = X3[Batchindex]
Labelsused = labelarray3[Batchindex]
Time = None
if modelflag == 0:
InputVector = np.reshape(InputVector,(-1,Tseq,NpropperseqTOT))
elif modelflag == 1:
Time = SetSpacetime(np.reshape(Labelsused[:,0],(1,-1)))
InputVector = np.reshape(InputVector,(1,Tseq*Nloc_sample,NpropperseqTOT))
PredictedVector = DLmodel(InputVector, training = PredictionTraining, Time=Time )
PredictedVector = np.reshape(PredictedVector,(1,Nloc_sample,NpredperseqTOT))
swbatched = sw[Batchindex,:,:]
if LocationBasedValidation:
swT = np.zeros([1,Nloc_sample,NpredperseqTOT],dtype = np.float32)
swV = np.zeros([1,Nloc_sample,NpredperseqTOT],dtype = np.float32)
for iloc_sample in range(0,Nloc_sample):
fudgeT = Nloc/TrainingNloc
fudgeV = Nloc/ValidationNloc
iloc = Labelsused[iloc_sample,1]
if MappingtoTraining[iloc] >= 0:
swT[0,iloc_sample,:] = swbatched[iloc_sample,:]*fudgeT
else:
swV[0,iloc_sample,:] = swbatched[iloc_sample,:]*fudgeV
TrueVector = y3[Batchindex]
TrueVector = np.reshape(TrueVector,(1,Nloc_sample,NpredperseqTOT))
swbatched = np.reshape(swbatched,(1,Nloc_sample,NpredperseqTOT))
losspercall = numpycustom_lossGCF1(TrueVector,PredictedVector,swbatched)
quan2 += losspercall
bbar.update(1)
if LocationBasedValidation:
losspercallTr = numpycustom_lossGCF1(TrueVector,PredictedVector,swT)
quan3 += losspercallTr
losspercallVl = numpycustom_lossGCF1(TrueVector,PredictedVector,swV)
quan4 += losspercallVl
for iloc_sample in range(0,Nloc_sample):
LocLocal = Labelsused[iloc_sample,1]
SeqLocal = Labelsused[iloc_sample,0]
yyhat = PredictedVector[0,iloc_sample]
CountFitPredictions [SeqLocal,LocLocal,:] += FRanges
SensitivityFitPredictions [SeqLocal,LocLocal,:,Sensitivities] += yyhat
fudge = 1.0/(1.0 + Batchindex)
mean2 = quan2 * fudge
if LocationBasedValidation:
mean3 = quan3 * fudge
mean4 = quan4 * fudge
bbar.set_postfix(AvLoss = mean2, AvTr = mean3, AvVl = mean4, Loss = losspercall, Tr = losspercallTr, Vl = losspercallVl)
else:
bbar.set_postfix(Loss = losspercall, AvLoss = mean2 )
# Processing at the end of Sampling Loop
fudge = 1.0/OuterBatchDimension
quan2 *= fudge
quan3 *= fudge
quan4 *= fudge
meanvalue2 += quan2
variance2 += quan2**2
variance3 += quan3**2
variance4 += quan4**2
if LocationBasedValidation:
meanvalue3 += quan3
meanvalue4 += quan4
samplebar.update(1)
if LocationBasedValidation:
samplebar.set_postfix(Shuffle=shuffling, Loss = quan2, Tr = quan3, Val = quan4)
else:
samplebar.set_postfix(Shuffle=shuffling, Loss = quan2)
bbar.reset()
# End Shuffling loop
if Sensitivities == 0:
iprop = -1
lineend = startbold+startred+ 'Basic Predictions' + resetfonts
else:
iprop = SensitivityList[Sensitivities-1]
nameplace = PropertyNameIndex[iprop]
maxminplace = PropertyAverageValuesPointer[iprop]
lastline = ' Normed Mean ' +str(round(QuantityStatistics[maxminplace,5],4))
lineend= startbold+startred + 'Property ' + str(iprop) + ' ' + InputPropertyNames[nameplace] + resetfonts + lastline
if modelflag == 2:
DLmodel.ResetProperty()
meanvalue2 /= SampleSize
global GlobalTrainingLoss, GlobalValidationLoss, GlobalLoss
printloss(' Full Loss ',meanvalue2,variance2,SampleSize, lineend = lineend)
meanvalue2 /= SampleSize
GlobalLoss = meanvalue2
GlobalTrainingLoss = 0.0
GlobalValidationLoss = 0.0
if LocationBasedValidation:
printloss(' Training Loss ',meanvalue3,variance3,SampleSize, lineend = lineend)
printloss(' Validation Loss ',meanvalue4,variance4,SampleSize, lineend = lineend)
meanvalue3 /= SampleSize
meanvalue4 /= SampleSize
GlobalTrainingLoss = meanvalue3
GlobalValidationLoss = meanvalue4
label = 'Sensitivity ' +str(Sensitivities)
Location_summed_plot(0, yin, SensitivityFitPredictions[:,:,:,Sensitivities] , extracomments = [label,label], Dumpplot = False)
# Sequence Location Predictions
SensitivityFitPredictions[:,:,:,Sensitivities] = np.divide(SensitivityFitPredictions[:,:,:,Sensitivities],CountFitPredictions[:,:,:])
if Sensitivities == 0:
Goldstandard = np.sum(np.abs(SensitivityFitPredictions[:,:,:,Sensitivities]), axis =(0,1))
TotalGS = np.sum(Goldstandard)
continue
Change = np.sum(np.abs(np.subtract(SensitivityFitPredictions[:,:,:,Sensitivities],SensitivityFitPredictions[:,:,:,0])), axis =(0,1))
TotalChange = np.sum(Change)
SensitivityChange[iprop] = TotalChange
print(str(round(TotalChange,5)) + ' GS ' + str(round(TotalGS,5)) + ' ' +lineend)
if SensitvitybyPrediction:
for ipred in range(0,NpredperseqTOT):
print(str(round(Change[ipred],5)) + ' GS ' + str(round(Goldstandard[ipred],5))
+ ' ' + str(ipred) + ' ' + Predictionname[ipred] + ' wgt ' + str(round(Predictionwgt[ipred],3)))
print(startbold+startred+ '\nSummarize Changes Total ' + str(round(TotalGS,5))+ ' Property ScaleFactor ' + str(round(ScaleProperty,3)) + resetfonts )
for Sensitivities in range(1,1+something):
iprop = SensitivityList[Sensitivities-1]
nameplace = PropertyNameIndex[iprop]
maxminplace = PropertyAverageValuesPointer[iprop]
lastline = ' Normed Mean ' +str(round(QuantityStatistics[maxminplace,5],4))
lastline += ' Normed Std ' +str(round(QuantityStatistics[maxminplace,6],4))
TotalChange = SensitivityChange[iprop]
NormedChange = TotalChange/((1-ScaleProperty)*TotalGS)
stdmeanratio = 0.0
stdchangeratio = 0.0
if np.abs(QuantityStatistics[maxminplace,5]) > 0.0001:
stdmeanratio = QuantityStatistics[maxminplace,6]/QuantityStatistics[maxminplace,5]
if np.abs(QuantityStatistics[maxminplace,6]) > 0.0001:
stdchangeratio = NormedChange/QuantityStatistics[maxminplace,6]
lratios = ' Normed Change '+ str(round(NormedChange,5)) + ' /std ' + str(round(stdchangeratio,5))
lratios += ' Std/Mean ' + str(round(stdmeanratio,5))
print(str(iprop) + ' Change '+ str(round(TotalChange,2)) + startbold + lratios
+ ' ' + InputPropertyNames[nameplace] + resetfonts + lastline)
current_time = timenow()
print(startbold+startred+ '\nEND DLPrediction2F ' + current_time + ' ' + RunName + RunComment +resetfonts)
return
# + [markdown] id="rMIeYApPybFA" pycharm={"name": "#%% md\n"}
# # TFT Model
# + [markdown] id="GkiL51xZ3XUr" pycharm={"name": "#%% md\n"}
# ##Set up TFT
# + [markdown] id="GqBfMQNwQj1z" pycharm={"name": "#%% md\n"}
# ###Data and Input Types
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="-tUyF2ZhQ3-C" outputId="001794d6-c375-4294-c87f-3b04b0d100b3" pycharm={"name": "#%%\n"}
# Type defintions
class DataTypes(enum.IntEnum):
"""Defines numerical types of each column."""
REAL_VALUED = 0
CATEGORICAL = 1
DATE = 2
NULL = -1
STRING = 3
BOOL = 4
class InputTypes(enum.IntEnum):
"""Defines input types of each column."""
TARGET = 0 # Known before and after t for training
OBSERVED_INPUT = 1 # Known upto time t
KNOWN_INPUT = 2 # Known at all times
STATIC_INPUT = 3 # By definition known at all times
ID = 4 # Single column used as an entity identifier
TIME = 5 # Single column exclusively used as a time index
NULL = -1
def checkdfNaN(label, AttributeSpec, y):
countNaN = 0
countnotNaN = 0
if y is None:
return
names = y.columns.tolist()
count = np.zeros(y.shape[1])
for j in range(0,y.shape[1]):
colname = names[j]
if AttributeSpec.loc[colname,'DataTypes'] != DataTypes.REAL_VALUED:
continue
for i in range(0,y.shape[0]):
if np.math.isnan(y.iloc[i, j]):
countNaN += 1
count[j] += 1
else:
countnotNaN += 1
percent = (100.0*countNaN)/(countNaN + countnotNaN)
print(label + ' is NaN ',str(countNaN),' percent ',str(round(percent,2)),' not NaN ', str(countnotNaN))
for j in range(0,y.shape[1]):
if count[j] == 0:
continue
print(names[j] + ' has NaN ' + str(count[j]))
# + [markdown] id="FOXCqWirQxZb" pycharm={"name": "#%% md\n"}
# ###Convert FFFFWNPF to TFT
# + colab={"base_uri": "https://localhost:8080/", "height": 5828} id="98Wz3J3y3b2z" outputId="6537b4ae-a931-4ce5-d0f2-6cef6316b232" pycharm={"name": "#%%\n"}
if UseTFTModel:
# Pick Values setting InputType
# Currently ONLY pick from properties BUT
# If PropPick = 0 (target) then these should be selected as predictions in FFFFWNPF and futured of length LengthFutures
# Set Prediction Property mappings and calculations
# PredictionTFTAction -2 a Future -1 Ignore 0 Futured Basic Prediction, 1 Nonfutured Simple Sum, 2 Nonfutured Energy Averaged Earthquake
# CalculatedPredmaptoRaw is Raw Prediction on which Calculated Prediction based
# PredictionCalcLength is >1 if Action=1,2 and says action based on this number of consequitive predictions
# PredictionTFTnamemapping if a non trivial string it is that returned by TFT in output map; if ' ' it isd a special extra prediction
PredictionTFTnamemapping =np.full(NpredperseqTOT,' ',dtype=object)
PredictionTFTAction = np.full(NpredperseqTOT, -1, dtype = np.int32)
for ipred in range(0,NpredperseqTOT):
if ipred >= NumpredbasicperTime:
PredictionTFTAction[ipred] = -2
elif FuturedPointer[ipred] >= 0:
PredictionTFTAction[ipred] = 0
# Default is -1
CalculatedPredmaptoRaw = np.full(NpredperseqTOT, -1, dtype = np.int32)
PredictionCalcLength = np.full(NpredperseqTOT, 1, dtype = np.int32)
# TFT Pick flags
# 0 Target and observed input
# 1 Observed Input NOT predicted
# 2 Known Input
# 3 Static Observed Input
#
# Data Types 0 Float or Integer converted to Float
# Assuming Special non futured 6 months forward prediction defined but NOT directly predicted by TFT
PropPick = [3,3,3,3,0,1,1,1,1,1,0,0,0,2,2,2,2,2,2,2,2,2,2,2,2,2,2]
PropDataType = [0] * NpropperseqTOT
# Dataframe is overall label (real starting at 0), Location Name, Time Input Properties, Predicted Properties Nloc times Num_Time values
# Row major order in Location-Time Space
Totalsize = (Num_Time + TFTExtraTimes) * Nloc
RawLabel = np.arange(0, Totalsize, dtype =np.float32)
LocationLabel = []
FFFFWNPFUniqueLabel = []
RawTime = np.empty([Nloc,Num_Time + TFTExtraTimes], dtype = np.float32)
RawTrain = np.full([Nloc,Num_Time + TFTExtraTimes], True, dtype = bool)
RawVal = np.full([Nloc,Num_Time + TFTExtraTimes], True, dtype = bool)
# print('Times ' + str(Num_Time) + ' ' + str(TFTExtraTimes))
ierror = 0
for ilocation in range(0,Nloc):
# locname = Locationstate[LookupLocations[ilocation]] + ' ' + Locationname[LookupLocations[ilocation]]
locname = Locationname[LookupLocations[ilocation]] + ' ' + Locationstate[LookupLocations[ilocation]]
if locname == "":
printexit('Illegal null location name ' + str(ilocation))
for idupe in range(0,len(FFFFWNPFUniqueLabel)):
if locname == FFFFWNPFUniqueLabel[idupe]:
print(' Duplicate location name ' + str(ilocation) + ' ' + str(idupe) + ' ' + locname)
ierror += 1
FFFFWNPFUniqueLabel.append(locname)
# print(str(ilocation) + ' ' +locname)
for jtime in range(0,Num_Time + TFTExtraTimes):
RawTime[ilocation,jtime] = np.float32(jtime)
LocationLabel.append(locname)
if LocationBasedValidation:
if MappingtoTraining[ilocation] >= 0:
RawTrain[ilocation,jtime] = True
else:
RawTrain[ilocation,jtime] = False
if MappingtoValidation[ilocation] >= 0:
RawVal[ilocation,jtime] = True
else:
RawVal[ilocation,jtime] = False
if ierror > 0:
printexit(" Duplicate Names " + str(ierror))
RawTime = np.reshape(RawTime,-1)
RawTrain = np.reshape(RawTrain,-1)
RawVal = np.reshape(RawVal,-1)
TFTdf1 = pd.DataFrame(RawLabel, columns=['RawLabel'])
if LocationBasedValidation:
TFTdf2 = pd.DataFrame(RawTrain, columns=['TrainingSet'])
TFTdf3 = pd.DataFrame(RawVal, columns=['ValidationSet'])
TFTdf4 = pd.DataFrame(LocationLabel, columns=['Location'])
TFTdf5 = pd.DataFrame(RawTime, columns=['Time from Start'])
TFTdfTotal = pd.concat([TFTdf1,TFTdf2,TFTdf3,TFTdf4,TFTdf5], axis=1)
else:
TFTdf2 = pd.DataFrame(LocationLabel, columns=['Location'])
TFTdf3 = pd.DataFrame(RawTime, columns=['Time from Start'])
TFTdfTotal = pd.concat([TFTdf1,TFTdf2,TFTdf3], axis=1)
TFTdfTotalSpec = pd.DataFrame([['RawLabel', DataTypes.REAL_VALUED, InputTypes.NULL]], columns=['AttributeName', 'DataTypes', 'InputTypes'])
if LocationBasedValidation:
TFTdfTotalSpec.loc[len(TFTdfTotalSpec.index)] = ['TrainingSet', DataTypes.BOOL, InputTypes.NULL]
TFTdfTotalSpec.loc[len(TFTdfTotalSpec.index)] = ['ValidationSet', DataTypes.BOOL, InputTypes.NULL]
TFTdfTotalSpec.loc[len(TFTdfTotalSpec.index)] = ['Location', DataTypes.STRING, InputTypes.ID]
TFTdfTotalSpec.loc[len(TFTdfTotalSpec.index)] = ['Time from Start', DataTypes.REAL_VALUED, InputTypes.TIME]
ColumnsProp=[]
for iprop in range(0,NpropperseqTOT):
line = str(iprop) + ' ' + InputPropertyNames[PropertyNameIndex[iprop]]
jprop = PropertyAverageValuesPointer[iprop]
if QuantityTakeroot[jprop] > 1:
line += ' Root ' + str(QuantityTakeroot[jprop])
ColumnsProp.append(line)
QuantityStatisticsNames = ['Min','Max','Norm','Mean','Std','Normed Mean','Normed Std']
TFTInputSequences = np.reshape(ReshapedSequencesTOT,(-1,NpropperseqTOT))
TFTPropertyChoice = np.full(NpropperseqTOT, -1, dtype = np.int32)
TFTNumberTargets = 0
for iprop in range(0,NpropperseqTOT):
if PropPick[iprop] >= 0:
if PropPick[iprop] == 0:
TFTNumberTargets += 1
nextcol = TFTInputSequences[:,iprop]
dftemp = pd.DataFrame(nextcol, columns=[ColumnsProp[iprop]])
TFTdfTotal = pd.concat([TFTdfTotal,dftemp], axis=1)
jprop = TFTdfTotal.columns.get_loc(ColumnsProp[iprop])
print('Property column ' + str(jprop) + ' ' + ColumnsProp[iprop])
TFTPropertyChoice[iprop] = jprop
TFTdfTotalSpec.loc[len(TFTdfTotalSpec.index)] = [ColumnsProp[iprop], PropDataType[iprop], PropPick[iprop]]
FFFFWNPFNumberTargets = TFTNumberTargets
ReshapedPredictionsTOT = np.transpose(RawInputPredictionsTOT,(1,0,2))
TFTdfTotalSpec = TFTdfTotalSpec.set_index('AttributeName', drop= False)
TFTdfTotalshape = TFTdfTotal.shape
TFTdfTotalcols = TFTdfTotal.columns
print(TFTdfTotalshape)
print(TFTdfTotalcols)
pd.set_option('display.max_rows', 100)
display(TFTdfTotalSpec)
print('Prediction mapping')
ifuture = 0
itarget = 0
for ipred in range(0,NpredperseqTOT):
predstatus = PredictionTFTAction[ipred]
if (predstatus == -1) or (predstatus > 0):
PredictionTFTnamemapping[ipred] = ' '
text = 'NOT PREDICTED DIRECTLY'
elif (predstatus == -2) or (predstatus == 0):
text = f't+{ifuture}-Obs{itarget}'
PredictionTFTnamemapping[ipred] = text
itarget += 1
if itarget >= TFTNumberTargets:
itarget = 0
ifuture += 1
fp = -2
if ipred < NumpredbasicperTime:
fp = FuturedPointer[ipred]
line = startbold + startpurple + str(ipred) + ' ' + Predictionname[ipred] + ' ' + text + resetfonts + ' Futured ' +str(fp) + ' '
line += 'Action ' + str(predstatus) + ' Property ' + str(CalculatedPredmaptoRaw[ipred]) + ' Length ' + str(PredictionCalcLength[ipred])
jpred = PredictionAverageValuesPointer[ipred]
line += ' Processing Root ' + str(QuantityTakeroot[jpred])
for proppredval in range (0,7):
line += ' ' + QuantityStatisticsNames[proppredval] + ' ' + str(round(QuantityStatistics[jpred,proppredval],3))
print(wraptotext(line,size=150))
# Rescaling done by that appropriate for properties and predictions
TFTdfTotalSpecshape = TFTdfTotalSpec.shape
TFTcolumn_definition = []
for i in range(0,TFTdfTotalSpecshape[0]):
TFTcolumn_definition.append((TFTdfTotalSpec.iloc[i,0],TFTdfTotalSpec.iloc[i,1],TFTdfTotalSpec.iloc[i,2]))
print(TFTcolumn_definition)
print(TFTdfTotalSpec.columns)
print(TFTdfTotalSpec.index)
# Set Futures to be calculated
PlotFutures = np.full(1+LengthFutures,False, dtype=bool)
PlotFutures[0] = True
PlotFutures[6] = True
PlotFutures[12] = True
PlotFutures[25] = True
PredictedQuantity = -NumpredbasicperTime
for ifuture in range (0,1+LengthFutures):
increment = NumpredbasicperTime
if ifuture > 1:
increment = NumpredFuturedperTime
PredictedQuantity += increment
for j in range(0,increment):
PlotPredictions[PredictedQuantity+j] = PlotFutures[ifuture]
CalculateNNSE[PredictedQuantity+j] = PlotFutures[ifuture]
# + [markdown] id="fbNT-soy5zkY" pycharm={"name": "#%% md\n"}
# ###TFT Setup
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="qVkN0Mn9526c" outputId="042c723d-5b1d-4b2d-ff7c-9a774771efe7" pycharm={"name": "#%%\n"}
if gregor:
content = readfile("config.yaml")
program_config = dotdict(yaml.safe_load(content))
config.update(program_config)
print(config)
DLAnalysisOnly = config.DLAnalysisOnly
DLRestorefromcheckpoint = config.DLRestorefromcheckpoint
DLinputRunName = RunName
DLinputCheckpointpostfix = config.DLinputCheckpointpostfix
TFTTransformerepochs = config.TFTTransformerepochs #num_epochs
#TFTTransformerepochs = 10 # just temporarily
#TFTTransformerepochs = 2 # just temporarily
# set transformer epochs to lower values for faster runs
# set them to 60 for now I set them to 20 so we get faster run
if False:
DLAnalysisOnly = True
DLRestorefromcheckpoint = True
DLinputRunName = RunName
DLinputRunName = 'EARTHQ-newTFTv28'
DLinputCheckpointpostfix = '-67'
TFTTransformerepochs = 40
TFTdropout_rate = 0.1
TFTdropout_rate = config.TFTdropout_rate #dropout_rate
TFTTransformerbatch_size = 64
TFTTransformerbatch_size = config.TFTTransformerbatch_size #minibatch_size
TFTd_model = 160
TFTd_model = config.TFTd_model #hidden_layer_size
TFTTransformertestvalbatch_size = max(128,TFTTransformerbatch_size) #maxibatch_size
TFThidden_layer_size = TFTd_model
number_LSTMnodes = TFTd_model
LSTMactivationvalue = 'tanh'
LSTMrecurrent_activation = 'sigmoid'
LSTMdropout1 = 0.0
LSTMrecurrent_dropout1 = 0.0
TFTLSTMEncoderInitialMLP = 0
TFTLSTMDecoderInitialMLP = 0
TFTLSTMEncoderrecurrent_dropout1 = LSTMrecurrent_dropout1
TFTLSTMDecoderrecurrent_dropout1 = LSTMrecurrent_dropout1
TFTLSTMEncoderdropout1 = LSTMdropout1
TFTLSTMDecoderdropout1 = LSTMdropout1
TFTLSTMEncoderrecurrent_activation = LSTMrecurrent_activation
TFTLSTMDecoderrecurrent_activation = LSTMrecurrent_activation
TFTLSTMEncoderactivationvalue = LSTMactivationvalue
TFTLSTMDecoderactivationvalue = LSTMactivationvalue
TFTLSTMEncoderSecondLayer = True
TFTLSTMDecoderSecondLayer = True
TFTLSTMEncoderThirdLayer = False
TFTLSTMDecoderThirdLayer = False
TFTLSTMEncoderFinalMLP = 0
TFTLSTMDecoderFinalMLP = 0
TFTnum_heads = 4
TFTnum_heads = config.TFTnum_heads #num_heads
TFTnum_AttentionLayers = 2
TFTnum_AttentionLayers = config.TFTnum_AttentionLayers #num_stacks | stack_size
# For default TFT
TFTuseCUDALSTM = True
TFTdefaultLSTM = False
if TFTdefaultLSTM:
TFTuseCUDALSTM = True
TFTLSTMEncoderFinalMLP = 0
TFTLSTMDecoderFinalMLP = 0
TFTLSTMEncoderrecurrent_dropout1 = 0.0
TFTLSTMDecoderrecurrent_dropout1 = 0.0
TFTLSTMEncoderdropout1 = 0.0
TFTLSTMDecoderdropout1 = 0.0
TFTLSTMEncoderSecondLayer = False
TFTLSTMDecoderSecondLayer = False
TFTFutures = 0
TFTFutures = 1 + LengthFutures
if TFTFutures == 0:
printexit('No TFT Futures defined')
TFTSingleQuantity = True
TFTLossFlag = 11
HuberLosscut = 0.01
if TFTSingleQuantity:
TFTQuantiles =[1.0]
TFTQuantilenames = ['MSE']
TFTPrimaryQuantileIndex = 0
else:
TFTQuantiles = [0.1,0.5,0.9]
TFTQuantilenames = ['p10','p50','p90']
TFTPrimaryQuantileIndex = 1
if TFTLossFlag == 11:
TFTQuantilenames = ['MAE']
if TFTLossFlag == 12:
TFTQuantilenames = ['Huber']
TFTfixed_params = {
'total_time_steps': Tseq + TFTFutures,
'num_encoder_steps': Tseq,
'num_epochs': TFTTransformerepochs,
#'early_stopping_patience': 60,
'early_stopping_patience': config.early_stopping_patience, #early_stopping_patience
'multiprocessing_workers': 12,
'optimizer': 'adam',
'lossflag': TFTLossFlag,
'HuberLosscut': HuberLosscut,
'AnalysisOnly': DLAnalysisOnly,
'inputRunName': DLinputRunName,
'Restorefromcheckpoint': DLRestorefromcheckpoint,
'inputCheckpointpostfix': DLinputCheckpointpostfix,
'maxibatch_size': TFTTransformertestvalbatch_size,
'TFTuseCUDALSTM':TFTuseCUDALSTM,
'TFTdefaultLSTM':TFTdefaultLSTM,
}
TFTmodel_params = {
'dropout_rate': TFTdropout_rate,
'hidden_layer_size': TFTd_model,
#'learning_rate': 0.0000005,
'learning_rate': config.learning_rate, #learning_rate
'minibatch_size': TFTTransformerbatch_size,
#'max_gradient_norm': 0.01,
'max_gradient_norm': config.max_gradient_norm, #max_gradient_norm
'num_heads': TFTnum_heads,
'stack_size': TFTnum_AttentionLayers,
}
TFTSymbolicWindows = False
TFTFinalGatingOption = 1
TFTMultivariate = True
TFTuse_testing_mode = False
# + [markdown] id="RBMkPML6MXY7" pycharm={"name": "#%% md\n"}
# ###Base Formatter
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="hWusLYsSMdX9" outputId="2dd58e48-d596-4787-fd21-8aa5c4928cdc" pycharm={"name": "#%%\n"}
class GenericDataFormatter(abc.ABC):
"""Abstract base class for all data formatters.
User can implement the abstract methods below to perform dataset-specific
manipulations.
"""
@abc.abstractmethod
def set_scalers(self, df):
"""Calibrates scalers using the data supplied."""
raise NotImplementedError()
@abc.abstractmethod
def transform_inputs(self, df):
"""Performs feature transformation."""
raise NotImplementedError()
@abc.abstractmethod
def format_predictions(self, df):
"""Reverts any normalisation to give predictions in original scale."""
raise NotImplementedError()
@abc.abstractmethod
def split_data(self, df):
"""Performs the default train, validation and test splits."""
raise NotImplementedError()
@property
@abc.abstractmethod
def _column_definition(self):
"""Defines order, input type and data type of each column."""
raise NotImplementedError()
@abc.abstractmethod
def get_fixed_params(self):
"""Defines the fixed parameters used by the model for training.
Requires the following keys:
'total_time_steps': Defines the total number of time steps used by TFT
'num_encoder_steps': Determines length of LSTM encoder (i.e. history)
'num_epochs': Maximum number of epochs for training
'early_stopping_patience': Early stopping param for keras
'multiprocessing_workers': # of cpus for data processing
Returns:
A dictionary of fixed parameters, e.g.:
fixed_params = {
'total_time_steps': 252 + 5,
'num_encoder_steps': 252,
'num_epochs': 100,
'early_stopping_patience': 5,
'multiprocessing_workers': 5,
}
"""
raise NotImplementedError
# Shared functions across data-formatters
@property
def num_classes_per_cat_input(self):
"""Returns number of categories per relevant input.
This is seqeuently required for keras embedding layers.
"""
return self._num_classes_per_cat_input
def get_num_samples_for_calibration(self):
"""Gets the default number of training and validation samples.
Use to sub-sample the data for network calibration and a value of -1 uses
all available samples.
Returns:
Tuple of (training samples, validation samples)
"""
return -1, -1
def get_column_definition(self):
""""Returns formatted column definition in order expected by the TFT."""
column_definition = self._column_definition
# Sanity checks first.
# Ensure only one ID and time column exist
def _check_single_column(input_type):
length = len([tup for tup in column_definition if tup[2] == input_type])
if length != 1:
raise ValueError(f'Illegal number of inputs ({length}) of type {input_type}')
_check_single_column(InputTypes.ID)
_check_single_column(InputTypes.TIME)
identifier = [tup for tup in column_definition if tup[2] == InputTypes.ID]
time = [tup for tup in column_definition if tup[2] == InputTypes.TIME]
real_inputs = [
tup for tup in column_definition if tup[1] == DataTypes.REAL_VALUED and
tup[2] not in {InputTypes.ID, InputTypes.TIME}
]
categorical_inputs = [
tup for tup in column_definition if tup[1] == DataTypes.CATEGORICAL and
tup[2] not in {InputTypes.ID, InputTypes.TIME}
]
return identifier + time + real_inputs + categorical_inputs
# XXX Looks important in reordering
def _get_input_columns(self):
"""Returns names of all input columns."""
return [
tup[0]
for tup in self.get_column_definition()
if tup[2] not in {InputTypes.ID, InputTypes.TIME}
]
def _get_tft_input_indices(self):
"""Returns the relevant indexes and input sizes required by TFT."""
# Functions
def _extract_tuples_from_data_type(data_type, defn):
return [
tup for tup in defn if tup[1] == data_type and
tup[2] not in {InputTypes.ID, InputTypes.TIME}
]
def _get_locations(input_types, defn):
return [i for i, tup in enumerate(defn) if tup[2] in input_types]
# Start extraction
column_definition = [
tup for tup in self.get_column_definition()
if tup[2] not in {InputTypes.ID, InputTypes.TIME}
]
categorical_inputs = _extract_tuples_from_data_type(DataTypes.CATEGORICAL,
column_definition)
real_inputs = _extract_tuples_from_data_type(DataTypes.REAL_VALUED,
column_definition)
locations = {
'input_size':
len(self._get_input_columns()),
'output_size':
len(_get_locations({InputTypes.TARGET}, column_definition)),
'category_counts':
self.num_classes_per_cat_input,
'input_obs_loc':
_get_locations({InputTypes.TARGET}, column_definition),
'static_input_loc':
_get_locations({InputTypes.STATIC_INPUT}, column_definition),
'known_regular_inputs':
_get_locations({InputTypes.STATIC_INPUT, InputTypes.KNOWN_INPUT},
real_inputs),
'known_categorical_inputs':
_get_locations({InputTypes.STATIC_INPUT, InputTypes.KNOWN_INPUT},
categorical_inputs),
}
return locations
def get_experiment_params(self):
"""Returns fixed model parameters for experiments."""
required_keys = [
'total_time_steps', 'num_encoder_steps', 'num_epochs',
'early_stopping_patience', 'multiprocessing_workers'
]
fixed_params = self.get_fixed_params()
for k in required_keys:
if k not in fixed_params:
raise ValueError(f'Field {k} missing from fixed parameter definitions!')
fixed_params['column_definition'] = self.get_column_definition()
fixed_params.update(self._get_tft_input_indices())
return fixed_params
# + [markdown] id="X-k-se9TA9M2" pycharm={"name": "#%% md\n"}
# ###TFT FFFFWNPF Formatter
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="rRZ5qaEdBKzm" outputId="1acb4f4e-ef4d-4b8d-ef71-591bc3c30ed3" pycharm={"name": "#%%\n"}
# Custom formatting functions for FFFFWNPF datasets.
#GenericDataFormatter = data_formatters.base.GenericDataFormatter
#DataTypes = data_formatters.base.DataTypes
#InputTypes = data_formatters.base.InputTypes
class FFFFWNPFFormatter(GenericDataFormatter):
"""
Defines and formats data for the Covid April 21 dataset.
Attributes:
column_definition: Defines input and data type of column used in the
experiment.
identifiers: Entity identifiers used in experiments.
"""
_column_definition = TFTcolumn_definition
def __init__(self):
"""Initialises formatter."""
self.identifiers = None
self._real_scalers = None
self._cat_scalers = None
self._target_scaler = None
self._num_classes_per_cat_input = []
self._time_steps = self.get_fixed_params()['total_time_steps']
def split_data(self, df, valid_boundary=-1, test_boundary=-1):
"""Splits data frame into training-validation-test data frames.
This also calibrates scaling object, and transforms data for each split.
Args:
df: Source data frame to split.
valid_boundary: Starting time for validation data
test_boundary: Starting time for test data
Returns:
Tuple of transformed (train, valid, test) data.
"""
print('Formatting train-valid-test splits.')
if LocationBasedValidation:
index = df['TrainingSet']
train = df[index == True]
index = df['ValidationSet']
valid = df[index == True]
index = train['Time from Start']
train = train[index<(Num_Time-0.5)]
index = valid['Time from Start']
valid = valid[index<(Num_Time-0.5)]
if test_boundary == -1:
test = df
# train.drop('TrainingSet', axis=1, inplace=True)
# train.drop('ValidationSet', axis=1, inplace=True)
# valid.drop('TrainingSet', axis=1, inplace=True)
# valid.drop('ValidationSet', axis=1, inplace=True)
else:
index = df['Time from Start']
train = df[index<(Num_Time-0.5)]
valid = df[index<(Num_Time-0.5)]
if test_boundary == -1:
test = df
if valid_boundary > 0:
train = df.loc[index < valid_boundary]
if test_boundary > 0:
valid = df.loc[(index >= valid_boundary - 7) & (index < test_boundary)]
else:
valid = df.loc[(index >= valid_boundary - 7)]
if test_boundary > 0:
test = df.loc[index >= test_boundary - 7]
self.set_scalers(train)
Trainshape = train.shape
Traincols = train.columns
print(' Train Shape ' + str(Trainshape))
print(Traincols)
Validshape = valid.shape
Validcols = valid.columns
print(' Validation Shape ' + str(Validshape))
print(Validcols)
if test_boundary >= -1:
return (self.transform_inputs(data) for data in [train, valid, test])
else:
return [train, valid]
def set_scalers(self, df):
"""Calibrates scalers using the data supplied.
Args:
df: Data to use to calibrate scalers.
"""
print('Setting scalers with training data...')
column_definitions = self.get_column_definition()
# print(column_definitions)
# print(InputTypes.TARGET)
id_column = myTFTTools.utilsget_single_col_by_input_type(InputTypes.ID,
column_definitions, TFTMultivariate)
target_column = myTFTTools.utilsget_single_col_by_input_type(InputTypes.TARGET,
column_definitions, TFTMultivariate)
# Format real scalers
real_inputs = myTFTTools.extract_cols_from_data_type(
DataTypes.REAL_VALUED, column_definitions,
{InputTypes.ID, InputTypes.TIME})
# Initialise scaler caches
self._real_scalers = {}
self._target_scaler = {}
identifiers = []
for identifier, sliced in df.groupby(id_column):
data = sliced[real_inputs].values
if TFTMultivariate == True:
targets = sliced[target_column].values
else:
targets = sliced[target_column].values
# self._real_scalers[identifier] = sklearn.preprocessing.StandardScaler().fit(data)
# self._target_scaler[identifier] = sklearn.preprocessing.StandardScaler().fit(targets)
identifiers.append(identifier)
# Format categorical scalers
categorical_inputs = myTFTTools.extract_cols_from_data_type(
DataTypes.CATEGORICAL, column_definitions,
{InputTypes.ID, InputTypes.TIME})
categorical_scalers = {}
num_classes = []
# Set categorical scaler outputs
self._cat_scalers = categorical_scalers
self._num_classes_per_cat_input = num_classes
# Extract identifiers in case required
self.identifiers = identifiers
def transform_inputs(self, df):
"""Performs feature transformations.
This includes both feature engineering, preprocessing and normalisation.
Args:
df: Data frame to transform.
Returns:
Transformed data frame.
"""
return df
def format_predictions(self, predictions):
"""Reverts any normalisation to give predictions in original scale.
Args:
predictions: Dataframe of model predictions.
Returns:
Data frame of unnormalised predictions.
"""
return predictions
# Default params
def get_fixed_params(self):
"""Returns fixed model parameters for experiments."""
fixed_params = TFTfixed_params
return fixed_params
def get_default_model_params(self):
"""Returns default optimised model parameters."""
model_params = TFTmodel_params
return model_params
def get_num_samples_for_calibration(self):
"""Gets the default number of training and validation samples.
Use to sub-sample the data for network calibration and a value of -1 uses
all available samples.
Returns:
Tuple of (training samples, validation samples)
"""
numtrain = TFTdfTotalshape[0]
numvalid = TFTdfTotalshape[0]
return numtrain, numvalid
# + [markdown] id="yywSY1y0_XIE" pycharm={"name": "#%% md\n"}
# ###Set TFT Parameter Dictionary
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="CTG5j-nVIDSX" outputId="de452b5b-36b7-4805-be71-fadc90ccc5c4" pycharm={"name": "#%%\n"}
def setTFTparameters(data_formatter):
# Sets up default params
fixed_params = data_formatter.get_experiment_params()
params = data_formatter.get_default_model_params()
params["model_folder"] = TFTmodel_folder
params['optimizer'] = Transformeroptimizer
fixed_params["quantiles"] = TFTQuantiles
fixed_params["quantilenames"] = TFTQuantilenames
fixed_params["quantileindex"] = TFTPrimaryQuantileIndex
fixed_params["TFTLSTMEncoderFinalMLP"] = TFTLSTMEncoderFinalMLP
fixed_params["TFTLSTMDecoderFinalMLP"] = TFTLSTMDecoderFinalMLP
fixed_params["TFTLSTMEncoderrecurrent_dropout1"] = TFTLSTMEncoderrecurrent_dropout1
fixed_params["TFTLSTMDecoderrecurrent_dropout1"] = TFTLSTMDecoderrecurrent_dropout1
fixed_params["TFTLSTMEncoderdropout1"] = TFTLSTMEncoderdropout1
fixed_params["TFTLSTMDecoderdropout1"] = TFTLSTMDecoderdropout1
fixed_params["TFTLSTMEncoderSecondLayer"] = TFTLSTMEncoderSecondLayer
fixed_params["TFTLSTMDecoderSecondLayer"] = TFTLSTMDecoderSecondLayer
fixed_params["TFTLSTMEncoderThirdLayer"] = TFTLSTMEncoderThirdLayer
fixed_params["TFTLSTMDecoderThirdLayer"] = TFTLSTMDecoderThirdLayer
fixed_params["TFTLSTMEncoderrecurrent_activation"] = TFTLSTMEncoderrecurrent_activation
fixed_params["TFTLSTMDecoderrecurrent_activation"] = TFTLSTMDecoderrecurrent_activation
fixed_params["TFTLSTMEncoderactivationvalue"] = TFTLSTMEncoderactivationvalue
fixed_params["TFTLSTMDecoderactivationvalue"] = TFTLSTMDecoderactivationvalue
fixed_params["TFTLSTMEncoderInitialMLP"] = TFTLSTMEncoderInitialMLP
fixed_params["TFTLSTMDecoderInitialMLP"] = TFTLSTMDecoderInitialMLP
fixed_params['number_LSTMnodes'] = number_LSTMnodes
fixed_params["TFTOption1"] = 1
fixed_params["TFTOption2"] = 0
fixed_params['TFTMultivariate'] = TFTMultivariate
fixed_params['TFTFinalGatingOption'] = TFTFinalGatingOption
fixed_params['TFTSymbolicWindows'] = TFTSymbolicWindows
fixed_params['name'] = 'TemporalFusionTransformer'
fixed_params['nameFFF'] = TFTexperimentname
fixed_params['runname'] = TFTRunName
fixed_params['runcomment'] = TFTRunComment
fixed_params['data_formatter'] = data_formatter
fixed_params['Validation'] = LocationBasedValidation
# Parameter overrides for testing only! Small sizes used to speed up script.
if TFTuse_testing_mode:
fixed_params["num_epochs"] = 1
params["hidden_layer_size"] = 5
# train_samples, valid_samples = 100, 10 is applied later
# Load all parameters -- fixed and model
for k in fixed_params:
params[k] = fixed_params[k]
return params
# + [markdown] id="g95F8DIAGR2e" pycharm={"name": "#%% md\n"}
# ###TFTTools
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="FwzZRu1RGZlp" outputId="637c14fc-af96-432e-ad28-cd907184b3fb" pycharm={"name": "#%%\n"}
class TFTTools(object):
def __init__(self, params, **kwargs):
# Args: params: Parameters to define TFT
self.name = params['name']
self.experimentname = params['nameFFF']
self.runname = params['runname']
self.runcomment = params['runcomment']
self.data_formatter = params['data_formatter']
self.lossflag = params['lossflag']
self.HuberLosscut = params['HuberLosscut']
self.optimizer = params['optimizer']
self.validation = params['Validation']
self.AnalysisOnly = params['AnalysisOnly']
self.Restorefromcheckpoint = params['Restorefromcheckpoint']
self.inputRunName = params['inputRunName']
self.inputCheckpointpostfix = params['inputCheckpointpostfix']
# Data parameters
self.time_steps = int(params['total_time_steps'])
self.input_size = int(params['input_size'])
self.output_size = int(params['output_size'])
self.category_counts = json.loads(str(params['category_counts']))
self.n_multiprocessing_workers = int(params['multiprocessing_workers'])
# Relevant indices for TFT
self._input_obs_loc = json.loads(str(params['input_obs_loc']))
self._static_input_loc = json.loads(str(params['static_input_loc']))
self._known_regular_input_idx = json.loads(
str(params['known_regular_inputs']))
self._known_categorical_input_idx = json.loads(
str(params['known_categorical_inputs']))
self.column_definition = params['column_definition']
# Network params
# self.quantiles = [0.1, 0.5, 0.9]
self.quantiles = params['quantiles']
self.NumberQuantiles = len(self.quantiles)
self.Quantilenames = params['quantilenames']
self.PrimaryQuantileIndex = int(params['quantileindex'])
self.useMSE = False
if self.NumberQuantiles == 1 and self.Quantilenames[0] == 'MSE':
self.useMSE = True
self.TFTOption1 = params['TFTOption1']
self.TFTOption2 = params['TFTOption2']
self.TFTMultivariate = params['TFTMultivariate']
self.TFTuseCUDALSTM = params['TFTuseCUDALSTM']
self.TFTdefaultLSTM = params['TFTdefaultLSTM']
self.number_LSTMnodes = params['number_LSTMnodes']
self.TFTLSTMEncoderInitialMLP = params["TFTLSTMEncoderInitialMLP"]
self.TFTLSTMDecoderInitialMLP = params["TFTLSTMDecoderInitialMLP"]
self.TFTLSTMEncoderFinalMLP = params['TFTLSTMEncoderFinalMLP']
self.TFTLSTMDecoderFinalMLP = params['TFTLSTMDecoderFinalMLP']
self.TFTLSTMEncoderrecurrent_dropout1 = params["TFTLSTMEncoderrecurrent_dropout1"]
self.TFTLSTMDecoderrecurrent_dropout1 = params["TFTLSTMDecoderrecurrent_dropout1"]
self.TFTLSTMEncoderdropout1 = params["TFTLSTMEncoderdropout1"]
self.TFTLSTMDecoderdropout1 = params["TFTLSTMDecoderdropout1"]
self.TFTLSTMEncoderrecurrent_activation = params["TFTLSTMEncoderrecurrent_activation"]
self.TFTLSTMDecoderrecurrent_activation = params["TFTLSTMDecoderrecurrent_activation"]
self.TFTLSTMEncoderactivationvalue = params["TFTLSTMEncoderactivationvalue"]
self.TFTLSTMDecoderactivationvalue = params["TFTLSTMDecoderactivationvalue"]
self.TFTLSTMEncoderSecondLayer = params["TFTLSTMEncoderSecondLayer"]
self.TFTLSTMDecoderSecondLayer = params["TFTLSTMDecoderSecondLayer"]
self.TFTLSTMEncoderThirdLayer = params["TFTLSTMEncoderThirdLayer"]
self.TFTLSTMDecoderThirdLayer = params["TFTLSTMDecoderThirdLayer"]
self.TFTFinalGatingOption = params['TFTFinalGatingOption']
self.TFTSymbolicWindows = params['TFTSymbolicWindows']
self.FinalLoopSize = 1
if (self.output_size == 1) and (self.NumberQuantiles == 1):
self.TFTFinalGatingOption = 0
if self.TFTFinalGatingOption > 0:
self.TFTLSTMFinalMLP = 0
self.FinalLoopSize = self.output_size * self.NumberQuantiles
# HYPER PARAMETERS
self.hidden_layer_size = int(params['hidden_layer_size']) # PARAMETER TFTd_model search for them in code
self.dropout_rate = float(params['dropout_rate']) # PARAMETER TFTdropout_rate
self.max_gradient_norm = float(params['max_gradient_norm']) # PARAMETER max_gradient_norm
self.learning_rate = float(params['learning_rate']) # PARAMETER learning_rate
self.minibatch_size = int(params['minibatch_size']) # PARAMETER TFTTransformerbatch_size
self.maxibatch_size = int(params['maxibatch_size']) # PARAMETER TFTTransformertestvalbatch_size = max(128, TFTTransformerbatch_size)
self.num_epochs = int(params['num_epochs']) # PARAMETER TFTTransformerepochs
self.early_stopping_patience = int(params['early_stopping_patience']) # PARAMETER early_stopping_patience????
self.num_encoder_steps = int(params['num_encoder_steps']) # PARAMETER Tseq (fixed by the problem, may not be useful)
self.num_stacks = int(params['stack_size']) # PARAMETER TFTnum_AttentionLayers ???
self.num_heads = int(params['num_heads']) # PARAMETER TFTnum_heads +++++
# Serialisation options
# XXX
# self._temp_folder = os.path.join(params['model_folder'], 'tmp')
# self.reset_temp_folder()
# Extra components to store Tensorflow nodes for attention computations
# XXX
# self._input_placeholder = None
# self._attention_components = None
# self._prediction_parts = None
self.TFTSeq = 0
self.TFTNloc = 0
self.UniqueLocations = []
def utilsget_single_col_by_input_type(self, input_type, column_definition, TFTMultivariate):
"""Returns name of single or multiple column.
Args:
input_type: Input type of column to extract
column_definition: Column definition list for experiment
"""
columnname = [tup[0] for tup in column_definition if tup[2] == input_type]
# allow multiple targets
if TFTMultivariate and (input_type == 0):
return columnname
else:
if len(columnname) != 1:
printexit(f'Invalid number of columns for Type {input_type}')
return columnname[0]
def _get_single_col_by_type(self, input_type):
return self.utilsget_single_col_by_input_type(input_type, self.column_definition, self.TFTMultivariate)
def extract_cols_from_data_type(self, data_type, column_definition,
excluded_input_types):
"""Extracts the names of columns that correspond to a define data_type.
Args:
data_type: DataType of columns to extract.
column_definition: Column definition to use.
excluded_input_types: Set of input types to exclude
Returns:
List of names for columns with data type specified.
"""
return [
tup[0]
for tup in column_definition
if tup[1] == data_type and tup[2] not in excluded_input_types
]
# Quantile Loss functions.
def tensorflow_quantile_loss(self, y, y_pred, quantile):
"""Computes quantile loss for tensorflow.
Standard quantile loss as defined in the "Training Procedure" section of
the main TFT paper
Args:
y: Targets
y_pred: Predictions
quantile: Quantile to use for loss calculations (between 0 & 1)
Returns:
Tensor for quantile loss.
"""
# Checks quantile
if quantile < 0 or quantile > 1:
printexit(f'Illegal quantile value={quantile}! Values should be between 0 and 1.')
prediction_underflow = y - y_pred
q_loss = quantile * tf.maximum(prediction_underflow, 0.) + (
1. - quantile) * tf.maximum(-prediction_underflow, 0.)
return tf.reduce_sum(q_loss, axis=-1)
def PrintTitle(self, extrawords):
current_time = timenow()
line = self.name + ' ' + self.experimentname + ' ' + self.runname + ' ' + self.runcomment
beginwords = ''
if extrawords != '':
beginwords = extrawords + ' '
print(wraptotext(startbold + startred + beginwords + current_time + ' ' + line + resetfonts))
ram_gb = StopWatch.get_sysinfo()["mem.available"]
print(f'Your runtime has {ram_gb} gigabytes of available RAM\n')
# + [markdown] id="5zKOTiH3smSk" pycharm={"name": "#%% md\n"}
# ### Setup Classic TFT
# + colab={"base_uri": "https://localhost:8080/", "height": 1250} id="-i1xIk-Gz3Cl" outputId="16623bbc-4cba-4d25-fd13-e95882b6562b" pycharm={"name": "#%%\n"}
'''
%cd "/content/gdrive/MyDrive/Colab Datasets/TFToriginal/"
%ls
%cd TFTCode/
TFTexperimentname= "FFFFWNPF"
output_folder = "../TFTData" # Please don't change this path
Rootmodel_folder = os.path.join(output_folder, 'saved_models', TFTexperimentname)
TFTmodel_folder = os.path.join(Rootmodel_folder, "fixed" + RunName)
'''
TFTexperimentname= "FFFFWNPF"
TFTmodel_folder="Notused"
TFTRunName = RunName
TFTRunComment = RunComment
if TFTexperimentname == 'FFFFWNPF':
formatter = FFFFWNPFFormatter()
# Save data frames
# TFTdfTotalSpec.to_csv('TFTdfTotalSpec.csv')
# TFTdfTotal.to_csv('TFTdfTotal.csv')
else:
import expt_settings.configs
ExperimentConfig = expt_settings.configs.ExperimentConfig
config = ExperimentConfig(name, output_folder)
formatter = config.make_data_formatter()
TFTparams = setTFTparameters(formatter)
myTFTTools = TFTTools(TFTparams)
myTFTTools.PrintTitle('Start TFT')
for k in TFTparams:
print('# {} = {}'.format(k, TFTparams[k]))
# + [markdown] id="M2F6ZPgybDtJ" pycharm={"name": "#%% md\n"}
# ###Read TFT Data
# + colab={"base_uri": "https://localhost:8080/", "height": 795} id="CJNPrs26bL7N" outputId="edad3de3-6132-46ee-ce97-0a80811d7b4a" pycharm={"name": "#%%\n"}
class TFTDataCache(object):
"""Caches data for the TFT.
This is a class and has no instances so uses cls not self
It just sets and uses a dictionary to record batched data locations"""
_data_cache = {}
@classmethod
def update(cls, data, key):
"""Updates cached data.
Args:
data: Source to update
key: Key to dictionary location
"""
cls._data_cache[key] = data
@classmethod
def get(cls, key):
"""Returns data stored at key location."""
return cls._data_cache[key]
@classmethod
def contains(cls, key):
"""Retuns boolean indicating whether key is present in cache."""
return key in cls._data_cache
class TFTdatasetup(object):
def __init__(self, **kwargs):
super(TFTdatasetup, self).__init__(**kwargs)
self.TFTNloc = 0
# XXX TFTNloc bad
if myTFTTools.TFTSymbolicWindows:
# Set up Symbolic maps allowing location order to differ (due to possible sorting in TFT)
id_col = myTFTTools._get_single_col_by_type(InputTypes.ID)
time_col = myTFTTools._get_single_col_by_type(InputTypes.TIME)
target_col = myTFTTools._get_single_col_by_type(InputTypes.TARGET)
input_cols = [
tup[0]
for tup in myTFTTools.column_definition
if tup[2] not in {InputTypes.ID, InputTypes.TIME}
]
self.UniqueLocations = TFTdfTotal[id_col].unique()
self.TFTNloc = len(self.UniqueLocations)
self.LocationLookup ={}
for i,locationname in enumerate(self.UniqueLocations):
self.LocationLookup[locationname] = i # maps name to TFT master location number
self.TFTnum_entries = 0 # Number of time values per location
for identifier, df in TFTdfTotal.groupby(id_col):
localnum_entries = len(df)
if self.TFTnum_entries == 0:
self.TFTnum_entries = localnum_entries
else:
if self.TFTnum_entries != localnum_entries:
printexit('Incorrect length in time for ' + identifier + ' ' + str(localnum_entries))
self.Lookupinputs = np.zeros((self.TFTNloc, self.TFTnum_entries, myTFTTools.input_size))
for identifier, df in TFTdfTotal.groupby(id_col):
location = self.LocationLookup[identifier]
self.Lookupinputs[location,:,:] = df[input_cols].to_numpy(dtype=np.float32,copy=True)
def __call__(self, data, Dataset_key, num_samples=-1):
"""Batches Dataset for training, Validation.
Testing not Batched
Args:
data: Data to batch
Dataset_key: Key used for cache
num_samples: Maximum number of samples to extract (-1 to use all data)
"""
max_samples = num_samples
if max_samples < 0:
max_samples = data.shape[0]
sampleddata = self._sampled_data(data, Dataset_key, max_samples=max_samples)
TFTDataCache.update(sampleddata, Dataset_key)
print(f'Cached data "{Dataset_key}" updated')
return sampleddata
def _sampled_data(self, data, Dataset_key, max_samples):
"""Samples segments into a compatible format.
Args:
data: Sources data to sample and batch
max_samples: Maximum number of samples in batch
Returns:
Dictionary of batched data with the maximum samples specified.
"""
if (max_samples < 1) and (max_samples != -1):
raise ValueError(f'Illegal number of samples specified! samples={max_samples}')
id_col = myTFTTools._get_single_col_by_type(InputTypes.ID)
time_col = myTFTTools._get_single_col_by_type(InputTypes.TIME)
#data.sort_values(by=[id_col, time_col], inplace=True) # gives warning message
print('Getting legal sampling locations.')
StopWatch.start("legal sampling location")
valid_sampling_locations = []
split_data_map = {}
self.TFTSeq = 0
for identifier, df in data.groupby(id_col):
self.TFTnum_entries = len(df)
self.TFTSeq = max(self.TFTSeq, self.TFTnum_entries-myTFTTools.time_steps+1)
if self.TFTnum_entries >= myTFTTools.time_steps:
valid_sampling_locations += [
(identifier, myTFTTools.time_steps + i)
for i in range(self.TFTnum_entries - myTFTTools.time_steps + 1)
]
split_data_map[identifier] = df
print(Dataset_key + ' max samples ' + str(max_samples) + ' actual ' + str(len(valid_sampling_locations)))
actual_samples = min(max_samples, len(valid_sampling_locations))
if 0 < max_samples < len(valid_sampling_locations):
print(f'Extracting {max_samples} samples...')
ranges = [
valid_sampling_locations[i] for i in np.random.choice(
len(valid_sampling_locations), max_samples, replace=False)
]
else:
print('Max samples={} exceeds # available segments={}'.format(
max_samples, len(valid_sampling_locations)))
ranges = valid_sampling_locations
id_col = myTFTTools._get_single_col_by_type(InputTypes.ID)
time_col = myTFTTools._get_single_col_by_type(InputTypes.TIME)
target_col = myTFTTools._get_single_col_by_type(InputTypes.TARGET)
input_cols = [
tup[0]
for tup in myTFTTools.column_definition
if tup[2] not in {InputTypes.ID, InputTypes.TIME}
]
if myTFTTools.TFTSymbolicWindows:
inputs = np.zeros((actual_samples), dtype = np.int32)
outputs = np.zeros((actual_samples, myTFTTools.time_steps, myTFTTools.output_size))
time = np.empty((actual_samples, myTFTTools.time_steps, 1), dtype=object)
identifiers = np.empty((actual_samples, myTFTTools.time_steps, 1), dtype=object)
oldlocationnumber = -1
storedlocation = np.zeros(self.TFTNloc, dtype = np.int32)
for i, tup in enumerate(ranges):
identifier, start_idx = tup
newlocationnumber = self.LocationLookup[identifier]
if newlocationnumber != oldlocationnumber:
oldlocationnumber = newlocationnumber
if storedlocation[newlocationnumber] == 0:
storedlocation[newlocationnumber] = 1
sliced = split_data_map[identifier].iloc[start_idx -
myTFTTools.time_steps:start_idx]
# inputs[i, :, :] = sliced[input_cols]
inputs[i] = np.left_shift(start_idx,16) + newlocationnumber
# Sequence runs from start_idx - myTFTTools.time_steps to start_idx i.e. start_idx is label of FINAL time step in position start_idx - 1
if myTFTTools.TFTMultivariate:
outputs[i, :, :] = sliced[target_col]
else:
outputs[i, :, :] = sliced[[target_col]]
time[i, :, 0] = sliced[time_col]
identifiers[i, :, 0] = sliced[id_col]
inputs = inputs.reshape(-1,1,1)
sampled_data = {
'inputs': inputs,
'outputs': outputs[:, myTFTTools.num_encoder_steps:, :],
'active_entries': np.ones_like(outputs[:, self.num_encoder_steps:, :]),
'time': time,
'identifier': identifiers
}
else:
inputs = np.zeros((actual_samples, myTFTTools.time_steps, myTFTTools.input_size), dtype=np.float32)
outputs = np.zeros((actual_samples, myTFTTools.time_steps, myTFTTools.output_size), dtype=np.float32)
time = np.empty((actual_samples, myTFTTools.time_steps, 1), dtype=object)
identifiers = np.empty((actual_samples, myTFTTools.time_steps, 1), dtype=object)
for i, tup in enumerate(ranges):
identifier, start_idx = tup
sliced = split_data_map[identifier].iloc[start_idx -
myTFTTools.time_steps:start_idx]
inputs[i, :, :] = sliced[input_cols]
if myTFTTools.TFTMultivariate:
outputs[i, :, :] = sliced[target_col]
else:
outputs[i, :, :] = sliced[[target_col]]
time[i, :, 0] = sliced[time_col]
identifiers[i, :, 0] = sliced[id_col]
sampled_data = {
'inputs': inputs,
'outputs': outputs[:, myTFTTools.num_encoder_steps:, :],
'active_entries': np.ones_like(outputs[:, myTFTTools.num_encoder_steps:, :], dtype=np.float32),
'time': time,
'identifier': identifiers
}
StopWatch.stop("legal sampling location")
return sampled_data
def dothedatasetup():
myTFTTools.PrintTitle("Loading & splitting data...")
if myTFTTools.experimentname == 'FFFFWNPF':
raw_data = TFTdfTotal
else:
printexit('Currently only FFFWNPF supported')
# raw_data = pd.read_csv(TFTdfTotal, index_col=0)
# XXX don't use test Could simplify
train, valid, test = myTFTTools.data_formatter.split_data(raw_data, test_boundary = -1)
train_samples, valid_samples = myTFTTools.data_formatter.get_num_samples_for_calibration()
test_samples = -1
if TFTuse_testing_mode:
train_samples, valid_samples,test_samples = 100, 10, 100
myTFTReader = TFTdatasetup()
train_data = myTFTReader(train, "train", num_samples=train_samples)
val_data = None
if valid_samples > 0:
val_data = myTFTReader(valid, "valid", num_samples=valid_samples)
test_data = myTFTReader(test, "test", num_samples=test_samples)
return train_data, val_data, test_data
StopWatch.start("data head setup")
TFTtrain_datacollection, TFTval_datacollection, TFTtest_datacollection = dothedatasetup()
StopWatch.stop("data head setup")
TFToutput_map = None # holder for final output
# + pycharm={"name": "#%%\n"}
# + [markdown] id="4YcUwbvg0hsU" pycharm={"name": "#%% md\n"}
# ##Predict TFT
# + pycharm={"name": "#%%\n"}
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="n8URv0ll0hsU" outputId="7eab50fa-aaca-442b-d02d-1af58c0a4a0e" pycharm={"name": "#%%\n"}
class TFTSaveandInterpret:
def __init__(self, currentTFTmodel, currentoutput_map, ReshapedPredictionsTOT):
# output_map is a dictionary pointing to dataframes
# output_map["targets"]) targets are called outputs on input
# output_map["p10"] is p10 quantile forecast
# output_map["p50"] is p10 quantile forecast
# output_map["p90"] is p10 quantile forecast
# Labelled by last real time in sequence (t-1) which starts at time Tseq-1 going up to Num_Time-1
# order of Dataframe columns is 'forecast_time', 'identifier',
#'t+0-Obs0', 't+0-Obs1', 't+1-Obs0', 't+1-Obs1', 't+2-Obs0', 't+2-Obs1', 't+3-Obs0', 't+3-Obs1',
#'t+4-Obs0', 't+4-Obs1', 't+5-Obs0', 't+5-Obs1', 't+6-Obs0', 't+6-Obs1', 't+7-Obs0', 't+7-Obs1',
#'t+8-Obs0', 't+8-Obs1', 't+9-Obs0', 't+9-Obs1', 't+10-Obs0', 't+10-Obs1', 't+11-Obs0', 't+11-Obs1',
#'t+12-Obs0', 't+12-Obs1', 't+13-Obs0', 't+13-Obs1', 't+14-Obs0', 't+14-Obs1''
# First time is FFFFWNPF Sequence # + Tseq-1
# Rows of data frame are ilocation*(Num_Seq+1) + FFFFWNPF Sequence #
# ilocation runs from 0 ... Nloc-1 in same order in both TFT and FFFFWNPF
self.ScaledProperty = -1
self.Scaled = False
self.savedcolumn = []
self.currentoutput_map = currentoutput_map
self.currentTFTmodel = currentTFTmodel
Sizes = self.currentoutput_map[TFTQuantilenames[TFTPrimaryQuantileIndex]].shape
self.Numx = Sizes[0]
self.Numy = Sizes[1]
self.Num_Seq1 = 1 + Num_Seq
self.MaxTFTSeq = self.Num_Seq1-1
expectednumx = self.Num_Seq1*Nloc
if expectednumx != self.Numx:
printexit(' Wrong sizes of TFT compared to FFFFWNPF ' + str(expectednumx) + ' ' + str(self.Numx))
self.ReshapedPredictionsTOT = ReshapedPredictionsTOT
return
def setFFFFmapping(self):
self.FFFFWNPFresults = np.zeros((self.Numx, NpredperseqTOT,3), dtype=np.float32)
mapFFFFtoTFT = np.empty(Nloc, dtype = np.int32)
TFTLoc = self.currentoutput_map[TFTQuantilenames[TFTPrimaryQuantileIndex]]['identifier'].unique()
FFFFWNPFLocLookup = {}
for i,locname in enumerate(FFFFWNPFUniqueLabel):
FFFFWNPFLocLookup[locname] = i
TFTLocLookup = {}
for i,locname in enumerate(TFTLoc):
TFTLocLookup[locname] = i
if FFFFWNPFLocLookup[locname] is None:
printexit('Missing TFT Location '+locname)
for i,locname in enumerate(FFFFWNPFUniqueLabel):
j = TFTLocLookup[locname]
if j is None:
printexit('Missing FFFFWNPF Location '+ locname)
mapFFFFtoTFT[i] = j
indexposition = np.empty(NpredperseqTOT, dtype=int)
output_mapcolumns = self.currentoutput_map[TFTQuantilenames[TFTPrimaryQuantileIndex]].columns
numcols = len(output_mapcolumns)
for ipred in range(0, NpredperseqTOT):
predstatus = PredictionTFTAction[ipred]
if predstatus > 0:
indexposition[ipred]= -1
continue
label = PredictionTFTnamemapping[ipred]
if label == ' ':
indexposition[ipred]=ipred
else:
findpos = -1
for i in range(0,numcols):
if label == output_mapcolumns[i]:
findpos = i
if findpos < 0:
printexit('Missing Output ' +str(ipred) + ' ' +label)
indexposition[ipred] = findpos
for iquantile in range(0,myTFTTools.NumberQuantiles):
for ilocation in range(0,Nloc):
for seqnumber in range(0,self.Num_Seq1):
for ipred in range(0,NpredperseqTOT):
predstatus = PredictionTFTAction[ipred]
if predstatus > 0:
continue
label = PredictionTFTnamemapping[ipred]
if label == ' ': # NOT calculated by TFT
if seqnumber >= Num_Seq:
value = 0.0
else:
value = self.ReshapedPredictionsTOT[ilocation, seqnumber, ipred]
else:
ActualTFTSeq = seqnumber
if ActualTFTSeq <= self.MaxTFTSeq:
ipos = indexposition[ipred]
dfindex = self.Num_Seq1*mapFFFFtoTFT[ilocation] + ActualTFTSeq
value = self.currentoutput_map[TFTQuantilenames[iquantile]].iloc[dfindex,ipos]
else:
dfindex = self.Num_Seq1*mapFFFFtoTFT[ilocation] + self.MaxTFTSeq
ifuture = int(ipred/FFFFWNPFNumberTargets)
jfuture = ActualTFTSeq - self.MaxTFTSeq + ifuture
if jfuture <= LengthFutures:
jpred = ipred + (jfuture-ifuture)*FFFFWNPFNumberTargets
value = self.currentoutput_map[TFTQuantilenames[iquantile]].iloc[dfindex,indexposition[jpred]]
else:
value = 0.0
FFFFdfindex = self.Num_Seq1*ilocation + seqnumber
self.FFFFWNPFresults[FFFFdfindex,ipred,iquantile] = value
# Set Calculated Quantities as previous ipred loop has set base values
for ipred in range(0,NpredperseqTOT):
predstatus = PredictionTFTAction[ipred]
if predstatus <= 0:
continue
Basedonprediction = CalculatedPredmaptoRaw[ipred]
predaveragevaluespointer = PredictionAverageValuesPointer[Basedonprediction]
rootflag = QuantityTakeroot[predaveragevaluespointer]
rawdata = np.empty(PredictionCalcLength[ipred],dtype =np.float32)
ActualTFTSeq = seqnumber
if ActualTFTSeq <= self.MaxTFTSeq:
for ifuture in range(0,PredictionCalcLength[ipred]):
if ifuture == 0:
kpred = Basedonprediction
else:
jfuture = NumpredbasicperTime + NumpredFuturedperTime*(ifuture-1)
kpred = jfuture + FuturedPointer[Basedonprediction]
if predstatus == 3:
newvalue = self.ReshapedPredictionsTOT[ilocation, ActualTFTSeq, kpred]/ QuantityStatistics[predaveragevaluespointer,2] + QuantityStatistics[predaveragevaluespointer,0]
else:
kpos = indexposition[kpred]
dfindex = self.Num_Seq1*mapFFFFtoTFT[ilocation] + ActualTFTSeq
newvalue = self.currentoutput_map[TFTQuantilenames[iquantile]].iloc[dfindex,kpos] / QuantityStatistics[predaveragevaluespointer,2] + QuantityStatistics[predaveragevaluespointer,0]
if rootflag == 2:
newvalue = newvalue**2
if rootflag == 3:
newvalue = newvalue**3
rawdata[ifuture] = newvalue
# Form collective quantity
if predstatus == 1:
value = rawdata.sum()
elif predstatus >= 2:
value = log_energy(rawdata, sumaxis=0)
else:
value = 0.0
value = SetTakeroot(value,QuantityTakeroot[ipred])
actualpredaveragevaluespointer = PredictionAverageValuesPointer[ipred]
value = (value-QuantityStatistics[actualpredaveragevaluespointer,0])*QuantityStatistics[actualpredaveragevaluespointer,2]
else: # Sequence out of range
value = 0.0
FFFFdfindex = self.Num_Seq1*ilocation + seqnumber
self.FFFFWNPFresults[FFFFdfindex,ipred,iquantile] = value
return
# Default returns the median (50% quantile)
def __call__(self, InputVector, Time= None, training = False, Quantile = None):
lenvector = InputVector.shape[0]
result = np.empty((lenvector,NpredperseqTOT), dtype=np.float32)
if Quantile is None:
Quantile = TFTPrimaryQuantileIndex
for ivector in range(0,lenvector):
dfindex = self.Num_Seq1*InputVector[ivector,0] + InputVector[ivector,1]
result[ivector,:] = self.FFFFWNPFresults[dfindex, :, Quantile]
return result
def CheckProperty(self, iprop):
# Return true if property defined for TFT
# set ScaledProperty to be column to be changed
if (iprop < 0) or (iprop >= NpropperseqTOT):
return False
jprop = TFTPropertyChoice[iprop]
if jprop >= 0:
return True
return False
def SetupProperty(self, iprop):
if self.Scaled:
self.ResetProperty()
if (iprop < 0) or (iprop >= NpropperseqTOT):
return False
jprop = TFTPropertyChoice[iprop]
if jprop >= 0:
self.ScaledProperty = jprop
self.savedcolumn = TFTdfTotal.iloc[:,jprop].copy()
return True
return False
def ScaleProperty(self, ScalingFactor):
jprop = self.ScaledProperty
TFTdfTotal.iloc[:,jprop] = ScalingFactor*self.savedcolumn
self.Scaled = True
return
def ResetProperty(self):
jprop = self.ScaledProperty
if jprop >= 0:
TFTdfTotal.iloc[:,jprop] = self.savedcolumn
self.Scaled = False
self.ScaledProperty = -1
return
# XXX Check MakeMapping
def MakeMapping(self):
best_params = TFTopt_manager.get_best_params()
TFTmodelnew = ModelClass(best_params, TFTdfTotal = TFTdfTotal, use_cudnn=use_tensorflow_with_gpu)
TFTmodelnew.load(TFTopt_manager.hyperparam_folder)
self.currentoutput_map = TFTmodelnew.predict(TFTdfTotal, return_targets=False)
self.setFFFFmapping()
return
# + [markdown] id="O3gAe0WaTYCS" pycharm={"name": "#%% md\n"}
# ###Visualize TFT
#
# Called from finalizeDL
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="8i_tRpVRaQ4x" outputId="ae6d73a1-9831-4e05-be28-cf6d9eed44a5" pycharm={"name": "#%%\n"}
def VisualizeTFT(TFTmodel, output_map):
MyFFFFWNPFLink = TFTSaveandInterpret(TFTmodel, output_map, ReshapedPredictionsTOT)
MyFFFFWNPFLink.setFFFFmapping()
modelflag = 2
FitPredictions = DLprediction(ReshapedSequencesTOT, RawInputPredictionsTOT, MyFFFFWNPFLink, modelflag, LabelFit ='TFT')
# Input Predictions RawInputPredictionsTOT for DLPrediction are ordered Sequence #, Location but
# Input Predictions ReshapedPredictionsTOT for TFTSaveandInterpret are ordered Location, Sequence#
# Note TFT maximum Sequence # is one larger than FFFFWNPF
# + [markdown] id="6GDZmm4c6Wmn" pycharm={"name": "#%% md\n"}
# ##TFT Routines
# + [markdown] id="yZXP6vYo4oGu" pycharm={"name": "#%% md\n"}
# ### GLUplusskip: Gated Linear unit plus add and norm with Skip
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="GSTW9UXV4wng" outputId="5866ba3a-add5-40c0-ad65-23ab088a39ec" pycharm={"name": "#%%\n"}
# GLU with time distribution optional
# Dropout on input dropout_rate
# Linear layer with hidden_layer_size and activation
# Linear layer with hidden_layer_size and sigmoid
# Follow with an add and norm
class GLUplusskip(tf.keras.layers.Layer):
def __init__(self, hidden_layer_size,
dropout_rate=None,
use_time_distributed=True,
activation=None,
GLUname = 'Default',
**kwargs):
"""Applies a Gated Linear Unit (GLU) to an input.
Follow with an add and norm
Args:
hidden_layer_size: Dimension of GLU
dropout_rate: Dropout rate to apply if any
use_time_distributed: Whether to apply across time (index 1)
activation: Activation function to apply to the linear feature transform if necessary
Returns:
Tuple of tensors for: (GLU output, gate)
"""
super(GLUplusskip, self).__init__(**kwargs)
self.Gatehidden_layer_size = hidden_layer_size
self.Gatedropout_rate = dropout_rate
self.Gateuse_time_distributed = use_time_distributed
self.Gateactivation = activation
if self.Gatedropout_rate is not None:
n1 = 'GLUSkip' + 'dropout' + GLUname
self.FirstDropout = tf.keras.layers.Dropout(self.Gatedropout_rate, name = n1)
n3 = 'GLUSkip' + 'DenseAct1' + GLUname
n5 = 'GLUSkip' + 'DenseAct2' + GLUname
if self.Gateuse_time_distributed:
n2 = 'GLUSkip' + 'TD1' + GLUname
self.Gateactivation_layer = tf.keras.layers.TimeDistributed(tf.keras.layers.Dense(self.Gatehidden_layer_size, activation=self.Gateactivation, name=n3), name=n2)
n4 = 'GLUSkip' + 'TD2' + GLUname
self.Gategated_layer = tf.keras.layers.TimeDistributed(tf.keras.layers.Dense(self.Gatehidden_layer_size, activation='sigmoid', name=n5), name=n4)
else:
self.Gateactivation_layer = tf.keras.layers.Dense(self.Gatehidden_layer_size, activation=self.Gateactivation, name=n3)
self.Gategated_layer = tf.keras.layers.Dense(self.Gatehidden_layer_size, activation='sigmoid', name=n5)
n6 = 'GLUSkip' + 'Mul' + GLUname
self.GateMultiply = tf.keras.layers.Multiply(name = n6)
n7 = 'GLUSkip'+ 'Add' + GLUname
n8 = 'GLUSkip' + 'Norm' + GLUname
self.GateAdd = tf.keras.layers.Add(name = n7)
self.GateNormalization = tf.keras.layers.LayerNormalization(name = n8)
#<EMAIL>
def call(self, Gateinput, Skipinput, training=None):
# Args:
# Gateinput: Input to gating layer
# Skipinput: Input to add and norm
if self.Gatedropout_rate is not None:
x = self.FirstDropout(Gateinput)
else:
x = Gateinput
activation_layer = self.Gateactivation_layer(x)
gated_layer = self.Gategated_layer(x)
# Formal end of GLU
GLUoutput = self.GateMultiply([activation_layer, gated_layer])
# Applies skip connection followed by layer normalisation to get GluSkip.
GLUSkipoutput = self.GateAdd([Skipinput,GLUoutput])
GLUSkipoutput = self.GateNormalization(GLUSkipoutput)
return GLUSkipoutput,gated_layer
# + [markdown] id="i7efbEn8kBlQ" pycharm={"name": "#%% md\n"}
# ###Linear Layer (Dense)
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="BeuDkoxDkG0C" outputId="bd0d2e70-1ed6-4a32-a2b5-54a38d607cf1" pycharm={"name": "#%%\n"}
# Layer utility functions.
# Single layer size activation with bias and time distribution optional
def TFTlinear_layer(size,
activation=None,
use_time_distributed=False,
use_bias=True,
LLname = 'Default'):
"""Returns simple Keras linear layer.
Args:
size: Output size
activation: Activation function to apply if required
use_time_distributed: Whether to apply layer across time
use_bias: Whether bias should be included in layer
"""
n1 = 'LL'+'Dense'+LLname
linear = tf.keras.layers.Dense(size, activation=activation, use_bias=use_bias,name=n1)
if use_time_distributed:
n2 = 'LL'+'TD'+LLname
linear = tf.keras.layers.TimeDistributed(linear,name=n2)
return linear
# + [markdown] id="v3-kq7hAvJpw" pycharm={"name": "#%% md\n"}
# ###Apply MLP
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="70vYILKAvM2M" outputId="5371e8c4-ccbe-4b66-d56e-bce37dd65602" pycharm={"name": "#%%\n"}
class apply_mlp(tf.keras.layers.Layer):
def __init__(self, hidden_layer_size, output_size, output_activation=None, hidden_activation='tanh', use_time_distributed=False, MLPname='Default', **kwargs):
"""Applies simple feed-forward network to an input.
Args:
hidden_layer_size: Hidden state size
output_size: Output size of MLP
output_activation: Activation function to apply on output
hidden_activation: Activation function to apply on input
use_time_distributed: Whether to apply across time
Returns:
Tensor for MLP outputs.
"""
super(apply_mlp, self).__init__(**kwargs)
self.MLPhidden_layer_size = hidden_layer_size
self.MLPoutput_size = output_size
self.MLPoutput_activation = output_activation
self.MLPhidden_activation = hidden_activation
self.MLPuse_time_distributed = use_time_distributed
n1 = 'MLPDense1' + MLPname
n2 = 'MLPDense2' + MLPname
if self.MLPuse_time_distributed:
n3 = 'MLPTD1' + MLPname
n4 = 'MLPTD2' + MLPname
MLPFirstLayer = tf.keras.layers.TimeDistributed(
tf.keras.layers.Dense(self.MLPhidden_layer_size, activation=self.MLPhidden_activation, name = n1), name = n3)
MLPSecondLayer = tf.keras.layers.TimeDistributed(
tf.keras.layers.Dense(self.MLPoutput_size, activation=self.MLPoutput_activation, name = n2),name = n4)
else:
MLPFirstLayer = tf.keras.layers.Dense(self.MLPhidden_layer_size, activation=self.MLPhidden_activation, name = n1)
MLPSecondLayer = tf.keras.layers.Dense(self.MLPoutput_size, activation=self.MLPoutput_activation, name = n2)
#EAGER@tf.function
def call(self, inputs):
# inputs: MLP inputs
hidden = MLPFirstLayer(inputs)
return MLPSecondLayer(hidden)
# + [markdown] id="lBZx6JdpQgn5" pycharm={"name": "#%% md\n"}
# ###GRN Gated Residual Network
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="ZSlsALLnQsaQ" outputId="eebfb89f-ad92-4602-fca0-6830a7c1de23" pycharm={"name": "#%%\n"}
# GRN Gated Residual Network
class GRN(tf.keras.layers.Layer):
def __init__(self, hidden_layer_size, output_size=None, dropout_rate=None,
use_additionalcontext = False, use_time_distributed=True, GRNname='Default', **kwargs):
"""Applies the gated residual network (GRN) as defined in paper.
Args:
hidden_layer_size: Internal state size
output_size: Size of output layer
dropout_rate: Dropout rate if dropout is applied
use_time_distributed: Whether to apply network across time dimension
Returns:
Tuple of tensors for: (GRN output, GLU gate)
"""
super(GRN, self).__init__(**kwargs)
self.GRNhidden_layer_size = hidden_layer_size
self.GRNoutput_size = output_size
if self.GRNoutput_size is None:
self.GRNusedoutput_size = self.GRNhidden_layer_size
else:
self.GRNusedoutput_size = self.GRNoutput_size
self.GRNdropout_rate = dropout_rate
self.GRNuse_time_distributed = use_time_distributed
self.use_additionalcontext = use_additionalcontext
if self.GRNoutput_size is not None:
n1 = 'GRN'+'Dense4' + GRNname
if self.GRNuse_time_distributed:
n2 = 'GRN'+'TD4' + GRNname
self.GRNDense4 = tf.keras.layers.TimeDistributed(tf.keras.layers.Dense(self.GRNusedoutput_size,name=n1),name=n2)
else:
self.GRNDense4 = tf.keras.layers.Dense(self.GRNusedoutput_size,name=n1)
n3 = 'GRNDense1' + GRNname
self.GRNDense1 = TFTlinear_layer(
self.GRNhidden_layer_size,
activation=None,
use_time_distributed=self.GRNuse_time_distributed,
LLname=n3)
if self.use_additionalcontext:
n4 = 'GRNDense2' + GRNname
self.GRNDense2= TFTlinear_layer(
self.GRNhidden_layer_size,
activation=None,
use_time_distributed=self.GRNuse_time_distributed,
use_bias=False,
LLname=n4)
n5 = 'GRNAct' + GRNname
self.GRNActivation = tf.keras.layers.Activation('elu',name=n5)
n6 = 'GRNDense3' + GRNname
self.GRNDense3 = TFTlinear_layer(
self.GRNhidden_layer_size,
activation=None,
use_time_distributed=self.GRNuse_time_distributed,
LLname =n6)
n7 = 'GRNGLU' + GRNname
self.GRNGLUplusskip = GLUplusskip(hidden_layer_size = self.GRNusedoutput_size, dropout_rate=self.GRNdropout_rate,
use_time_distributed= self.GRNuse_time_distributed, GLUname=n7)
#<EMAIL>
def call(self, x, additional_context=None, return_gate=False, training=None):
"""Args:
x: Network inputs
additional_context: Additional context vector to use if relevant
return_gate: Whether to return GLU gate for diagnostic purposes
"""
# Setup skip connection of given size
if self.GRNoutput_size is None:
skip = x
else:
skip = self.GRNDense4(x)
# Apply feedforward network
hidden = self.GRNDense1(x)
if additional_context is not None:
if not self.use_additionalcontext:
printexit('Inconsistent context in GRN')
hidden = hidden + self.GRNDense2(additional_context)
else:
if self.use_additionalcontext:
printexit('Inconsistent context in GRN')
hidden = self.GRNActivation(hidden)
hidden = self.GRNDense3(hidden)
gating_layer, gate = self.GRNGLUplusskip(hidden,skip)
if return_gate:
return gating_layer, gate
else:
return gating_layer
# + [markdown] id="HrQ7fndaXFfO" pycharm={"name": "#%% md\n"}
# ###Process Static Variables
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="jdAOB1s2XJ8K" outputId="1f61899d-484f-424b-f88e-a865361f5dc6" pycharm={"name": "#%%\n"}
# Process Static inputs in TFT Style
# TFTScaledStaticInputs[Location,0...NumTrueStaticVariables]
class ProcessStaticInput(tf.keras.layers.Layer):
def __init__(self, hidden_layer_size, dropout_rate, num_staticproperties, **kwargs):
super(ProcessStaticInput, self).__init__(**kwargs)
self.hidden_layer_size = hidden_layer_size
self.num_staticproperties = num_staticproperties
self.dropout_rate = dropout_rate
n4 = 'ProcStaticFlat'
self.Flatten = tf.keras.layers.Flatten(name=n4)
n5 = 'ProcStaticG1'
n7 = 'ProcStaticSoftmax'
n8 = 'ProcStaticMul'
self.StaticInputGRN1 = GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate,
output_size=self.num_staticproperties, use_time_distributed=False, GRNname=n5)
self.StaticInputGRN2 = []
for i in range(0,self.num_staticproperties):
n6 = 'ProcStaticG2-'+str(i)
self.StaticInputGRN2.append(GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate,
use_time_distributed=False, GRNname = n6))
self.StaticInputsoftmax = tf.keras.layers.Activation('softmax', name= n7)
self.StaticMultiply = tf.keras.layers.Multiply(name = n8)
#<EMAIL>
def call(self, static_inputs, training=None):
# Embed Static Inputs
num_static = static_inputs.shape[1]
if num_static != self.num_staticproperties:
printexit('Incorrect number of static variables')
if num_static == 0:
return None, None
# static_inputs is [Batch, Static variable, TFTd_model] converted to
# flatten is [Batch, Static variable*TFTd_model]
flatten = self.Flatten(static_inputs)
# Nonlinear transformation with gated residual network.
mlp_outputs = self.StaticInputGRN1(flatten)
sparse_weights = self.StaticInputsoftmax(mlp_outputs)
sparse_weights = tf.expand_dims(sparse_weights, axis=-1)
trans_emb_list = []
for i in range(num_static):
e = self.StaticInputGRN2[i](static_inputs[:,i:i+1,:])
trans_emb_list.append(e)
transformed_embedding = tf.concat(trans_emb_list, axis=1)
combined = self.StaticMultiply([sparse_weights, transformed_embedding])
static_encoder = tf.math.reduce_sum(combined, axis=1)
return static_encoder, sparse_weights
# + [markdown] id="w-CC_kwDcT55" pycharm={"name": "#%% md\n"}
# ###Process Dynamic Variables
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="L79o4ClUcbLo" outputId="774f4299-ea46-4c1d-bb30-ebe57a0fb75c" pycharm={"name": "#%%\n"}
# Process Initial Dynamic inputs in TFT Style
# ScaledDynamicInputs[Location, time_steps,0...NumDynamicVariables]
class ProcessDynamicInput(tf.keras.layers.Layer):
def __init__(self, hidden_layer_size, dropout_rate, NumDynamicVariables, PDIname='Default', **kwargs):
super(ProcessDynamicInput, self).__init__(**kwargs)
self.hidden_layer_size = hidden_layer_size
self.NumDynamicVariables = NumDynamicVariables
self.dropout_rate = dropout_rate
n6 = PDIname + 'ProcDynG1'
n8 = PDIname + 'ProcDynSoftmax'
n9 = PDIname + 'ProcDynMul'
self.DynamicVariablesGRN1 = GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate,
output_size=self.NumDynamicVariables, use_additionalcontext = True, use_time_distributed=True, GRNname = n6)
self.DynamicVariablesGRN2 = []
for i in range(0,self.NumDynamicVariables):
n7 = PDIname + 'ProcDynG2-'+str(i)
self.DynamicVariablesGRN2.append(GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate,
use_additionalcontext = False, use_time_distributed=True, name = n7))
self.DynamicVariablessoftmax = tf.keras.layers.Activation('softmax', name = n8)
self.DynamicVariablesMultiply = tf.keras.layers.Multiply(name = n9)
#<EMAIL>
def call(self, dynamic_variables, static_context_variable_selection=None, training=None):
# Add time window index to static context
if static_context_variable_selection is None:
self.expanded_static_context = None
else:
self.expanded_static_context = tf.expand_dims(static_context_variable_selection, axis=1)
# Test Dynamic Variables
num_dynamic = dynamic_variables.shape[-1]
if num_dynamic != self.NumDynamicVariables:
printexit('Incorrect number of Dynamic Inputs ' + str(num_dynamic) + ' ' + str(self.NumDynamicVariables))
if num_dynamic == 0:
return None, None, None
# dynamic_variables is [Batch, Time window index, Dynamic variable, TFTd_model] converted to
# flatten is [Batch, Time window index, Dynamic variable,*TFTd_model]
_,time_steps,embedding_dimension,num_inputs = dynamic_variables.get_shape().as_list()
flatten = tf.reshape(dynamic_variables, [-1,time_steps,embedding_dimension * num_inputs])
# Nonlinear transformation with gated residual network.
mlp_outputs, static_gate = self.DynamicVariablesGRN1(flatten, additional_context=self.expanded_static_context, return_gate=True)
sparse_weights = self.DynamicVariablessoftmax(mlp_outputs)
sparse_weights = tf.expand_dims(sparse_weights, axis=2)
trans_emb_list = []
for i in range(num_dynamic):
e = self.DynamicVariablesGRN2[i](dynamic_variables[Ellipsis,i], additional_context=None)
trans_emb_list.append(e)
transformed_embedding = tf.stack(trans_emb_list, axis=-1)
combined = self.DynamicVariablesMultiply([sparse_weights, transformed_embedding])
temporal_ctx = tf.math.reduce_sum(combined, axis=-1)
return temporal_ctx, sparse_weights, static_gate
# + [markdown] id="r4IlhggIYhVZ" pycharm={"name": "#%% md\n"}
# ###TFT LSTM
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="td9kgEmYoG3F" outputId="a1acf827-efc5-4f18-8dd3-685b93342626" pycharm={"name": "#%%\n"}
class TFTLSTMLayer(tf.keras.Model):
# Class for TFT Encoder multiple layer LSTM with possible FCN at start and end
# All parameters defined externally
def __init__(self, TFTLSTMSecondLayer, TFTLSTMThirdLayer,
TFTLSTMInitialMLP, TFTLSTMFinalMLP,
TFTnumber_LSTMnodes, TFTLSTMd_model,
TFTLSTMactivationvalue, TFTLSTMrecurrent_activation,
TFTLSTMdropout1, TFTLSTMrecurrent_dropout1,
TFTreturn_state, LSTMname='Default', **kwargs):
super(TFTLSTMLayer, self).__init__(**kwargs)
self.TFTLSTMSecondLayer = TFTLSTMSecondLayer
self.TFTLSTMThirdLayer = TFTLSTMThirdLayer
self.TFTLSTMInitialMLP = TFTLSTMInitialMLP
self.TFTLSTMFinalMLP = TFTLSTMFinalMLP
self.TFTLSTMd_model = TFTLSTMd_model
self.TFTnumber_LSTMnodes = TFTnumber_LSTMnodes
self.TFTLSTMactivationvalue = TFTLSTMactivationvalue
self.TFTLSTMdropout1 = TFTLSTMdropout1
self.TFTLSTMrecurrent_dropout1 = TFTLSTMrecurrent_dropout1
self.TFTLSTMrecurrent_activation = TFTLSTMrecurrent_activation
self.TFTLSTMreturn_state = TFTreturn_state
self.first_return_state = self.TFTLSTMreturn_state
if self.TFTLSTMSecondLayer:
self.first_return_state = True
self.second_return_state = self.TFTLSTMreturn_state
if self.TFTLSTMThirdLayer:
self.second_return_state = True
self.third_return_state = self.TFTLSTMreturn_state
if self.TFTLSTMInitialMLP > 0:
n1= LSTMname +'LSTMDense1'
self.dense_1 = tf.keras.layers.Dense(self.TFTLSTMInitialMLP, activation=self.TFTLSTMactivationvalue, name =n1)
n2= LSTMname +'LSTMLayer1'
if myTFTTools.TFTuseCUDALSTM:
self.LSTM_1 = tf.compat.v1.keras.layers.CuDNNLSTM(
self.TFTnumber_LSTMnodes,
return_sequences=True,
return_state=self.first_return_state,
stateful=False, name=n2)
else:
self.LSTM_1 =tf.keras.layers.LSTM(self.TFTnumber_LSTMnodes, recurrent_dropout= self.TFTLSTMrecurrent_dropout1, dropout = self.TFTLSTMdropout1,
return_state = self.first_return_state, activation= self.TFTLSTMactivationvalue , return_sequences=True,
recurrent_activation= self.TFTLSTMrecurrent_activation, name=n2)
if self.TFTLSTMSecondLayer:
n3= LSTMname +'LSTMLayer2'
if myTFTTools.TFTuseCUDALSTM:
self.LSTM_2 = tf.compat.v1.keras.layers.CuDNNLSTM(
self.TFTnumber_LSTMnodes,
return_sequences=True,
return_state=self.second_return_state,
stateful=False, name=n3)
else:
self.LSTM_2 =tf.keras.layers.LSTM(self.TFTnumber_LSTMnodes, recurrent_dropout= self.TFTLSTMrecurrent_dropout1, dropout = self.TFTLSTMdropout1,
return_state = self.second_return_state, activation= self.TFTLSTMactivationvalue , return_sequences=True,
recurrent_activation= self.TFTLSTMrecurrent_activation, name=n3)
if self.TFTLSTMThirdLayer:
n4= LSTMname +'LSTMLayer3'
if myTFTTools.TFTuseCUDALSTM:
self.LSTM_3 = tf.compat.v1.keras.layers.CuDNNLSTM(
self.TFTnumber_LSTMnodes,
return_sequences=True,
return_state=self.third_return_state,
stateful=False, name=n4)
else:
self.LSTM_3 =tf.keras.layers.LSTM(self.TFTnumber_LSTMnodes, recurrent_dropout= self.TFTLSTMrecurrent_dropout1, dropout = self.TFTLSTMdropout1,
return_state = self.third_return_state, activation= self.TFTLSTMactivationvalue ,
return_sequences=True, recurrent_activation= self.TFTLSTMrecurrent_activation, name=n4)
if self.TFTLSTMFinalMLP > 0:
n5= LSTMname +'LSTMDense2'
n6= LSTMname +'LSTMDense3'
self.dense_2 = tf.keras.layers.Dense(self.TFTLSTMFinalMLP, activation=self.TFTLSTMactivationvalue, name=n5)
self.dense_f = tf.keras.layers.Dense(self.TFTLSTMd_model, name= n6)
#<EMAIL>
def call(self, inputs, initial_state = None, training=None):
if initial_state is None:
printexit(' Missing context in LSTM ALL')
if initial_state[0] is None:
printexit(' Missing context in LSTM h')
if initial_state[1] is None:
printexit(' Missing context in LSTM c')
returnstate_h = None
returnstate_c = None
if self.TFTLSTMInitialMLP > 0:
Runningdata = self.dense_1(inputs)
else:
Runningdata = inputs
if self.first_return_state:
Runningdata, returnstate_h, returnstate_c = self.LSTM_1(inputs, training=training, initial_state=initial_state)
if returnstate_h is None:
printexit('Missing context in LSTM returnstate_h')
if returnstate_c is None:
printexit('Missing context in LSTM returnstate_c')
else:
Runningdata = self.LSTM_1(inputs, training=training, initial_state=initial_state)
if self.TFTLSTMSecondLayer:
initial_statehc2 = None
if self.first_return_state:
initial_statehc2 = [returnstate_h, returnstate_c]
if self.second_return_state:
Runningdata, returnstate_h, returnstate_c = self.LSTM_2(Runningdata, training=training, initial_state=initial_statehc2)
if returnstate_h is None:
printexit('Missing context in LSTM returnstate_h2')
if returnstate_c is None:
printexit('Missing context in LSTM returnstate_c2')
else:
Runningdata = self.LSTM_2(Runningdata, training=training, initial_state=initial_statehc2)
if self.TFTLSTMThirdLayer:
initial_statehc3 = None
if self.first_return_state:
initial_statehc3 = [returnstate_h, returnstate_c]
if self.third_return_state:
Runningdata, returnstate_h, returnstate_c = self.LSTM_3(Runningdata, training=training, initial_state=initial_statehc3)
else:
Runningdata = self.LSTM_3(Runningdata, training=training, initial_state=initial_statehc3)
if self.TFTLSTMFinalMLP > 0:
Runningdata = self.dense_2(Runningdata)
Outputdata = self.dense_f(Runningdata)
else:
Outputdata = Runningdata
if self.TFTLSTMreturn_state:
return Outputdata, returnstate_h, returnstate_c
else:
return Outputdata
def build_graph(self, shapes):
input = tf.keras.layers.Input(shape=shapes, name="Input")
return tf.keras.models.Model(inputs=[input], outputs=[self.call(input)])
# + [markdown] id="0re4iqkmCuv4" pycharm={"name": "#%% md\n"}
# ###TFT Multihead Temporal Attention
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="r01Yst0VJgbv" outputId="b9b3c62a-f511-49f2-e3d6-4a0d8d7598a3" pycharm={"name": "#%%\n"}
# Attention Components.
#<EMAIL>
def TFTget_decoder_mask(self_attn_inputs):
"""Returns causal mask to apply for self-attention layer.
Args:
self_attn_inputs: Inputs to self attention layer to determine mask shape
"""
len_s = tf.shape(self_attn_inputs)[1]
bs = tf.shape(self_attn_inputs)[:1]
mask = tf.math.cumsum(tf.eye(len_s, batch_shape=bs), 1)
return mask
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="p3PNlQ5eJYq-" outputId="758bf5b2-8a3c-49ff-f361-37d0bdbf8ed9" pycharm={"name": "#%%\n"}
class TFTScaledDotProductAttention(tf.keras.Model):
"""Defines scaled dot product attention layer for TFT
Attributes:
dropout: Dropout rate to use
activation: Normalisation function for scaled dot product attention (e.g.
softmax by default)
"""
def __init__(self, attn_dropout=0.0, SPDAname='Default', **kwargs):
super(TFTScaledDotProductAttention, self).__init__(**kwargs)
n1 = SPDAname + 'SPDADropout'
n2 = SPDAname + 'SPDASoftmax'
n3 = SPDAname + 'SPDAAdd'
self.dropoutlayer = tf.keras.layers.Dropout(attn_dropout, name= n1)
self.activationlayer = tf.keras.layers.Activation('softmax', name= n2)
self.addlayer = tf.keras.layers.Add(name=n3)
#<EMAIL>
def call(self, q, k, v, mask):
"""Applies scaled dot product attention.
Args:
q: Queries
k: Keys
v: Values
mask: Masking if required -- sets softmax to very large value
Returns:
Tuple of (layer outputs, attention weights)
"""
temper = tf.sqrt(tf.cast(tf.shape(k)[-1], dtype='float32'))
attn = tf.keras.layers.Lambda(lambda x: tf.keras.backend.batch_dot(x[0], x[1], axes=[2, 2]) / temper)(
[q, k]) # shape=(batch, q, k)
if mask is not None:
mmask = tf.keras.layers.Lambda(lambda x: (-1e+9) * (1. - tf.cast(x, 'float32')))( mask) # setting to infinity
attn = self.addlayer([attn, mmask])
attn = self.activationlayer(attn)
attn = self.dropoutlayer(attn)
output = tf.keras.layers.Lambda(lambda x: tf.keras.backend.batch_dot(x[0], x[1]))([attn, v])
return output, attn
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="ZpkCmJw3C715" outputId="60804b2f-62f0-4aa0-d7b3-482eb101cf65" pycharm={"name": "#%%\n"}
class TFTInterpretableMultiHeadAttention(tf.keras.Model):
"""Defines interpretable multi-head attention layer for time only.
Attributes:
n_head: Number of heads
d_k: Key/query dimensionality per head
d_v: Value dimensionality
dropout: Dropout rate to apply
qs_layers: List of queries across heads
ks_layers: List of keys across heads
vs_layers: List of values across heads
attention: Scaled dot product attention layer
w_o: Output weight matrix to project internal state to the original TFT
state size
"""
#<EMAIL>
def __init__(self, n_head, d_model, dropout, MHAname ='Default', **kwargs):
super(TFTInterpretableMultiHeadAttention, self).__init__(**kwargs)
"""Initialises layer.
Args:
n_head: Number of heads
d_model: TFT state dimensionality
dropout: Dropout discard rate
"""
self.n_head = n_head
self.d_k = self.d_v = d_model // n_head
self.d_model = d_model
self.dropout = dropout
self.qs_layers = []
self.ks_layers = []
self.vs_layers = []
# Use same value layer to facilitate interp
n3= MHAname + 'MHAV'
vs_layer = tf.keras.layers.Dense(self.d_v, use_bias=False,name= n3)
self.Dropoutlayer1 =[]
for i_head in range(n_head):
n1= MHAname + 'MHAQ' + str(i)
n2= MHAname + 'MHAK' + str(i)
self.qs_layers.append(tf.keras.layers.Dense(self.d_k, use_bias=False, name = n1))
self.ks_layers.append(tf.keras.layers.Dense(self.d_k, use_bias=False, name = n2))
self.vs_layers.append(vs_layer) # use same vs_layer
n4= MHAname + 'Dropout1-' + str(i)
self.Dropoutlayer1.append(tf.keras.layers.Dropout(self.dropout, name = n4))
self.attention = TFTScaledDotProductAttention(SPDAname = MHAname)
n5= MHAname + 'Dropout2'
n6= MHAname + 'w_olayer'
self.Dropoutlayer2 = tf.keras.layers.Dropout(self.dropout, name = n5)
self.w_olayer = tf.keras.layers.Dense(d_model, use_bias=False, name = n6)
#EAGER@tf.<EMAIL>
def call(self, q, k, v, mask=None):
"""Applies interpretable multihead attention.
Using T to denote the number of past + future time steps fed into the transformer.
Args:
q: Query tensor of shape=(?, T, d_model)
k: Key of shape=(?, T, d_model)
v: Values of shape=(?, T, d_model)
mask: Masking if required with shape=(?, T, T)
Returns:
Tuple of (layer outputs, attention weights)
"""
heads = []
attns = []
for i in range(self.n_head):
qs = self.qs_layers[i](q)
ks = self.ks_layers[i](k)
vs = self.vs_layers[i](v)
head, attn = self.attention(qs, ks, vs, mask)
head_dropout = self.Dropoutlayer1[i](head)
heads.append(head_dropout)
attns.append(attn)
head = tf.stack(heads) if self.n_head > 1 else heads[0]
attn = tf.stack(attns)
outputs = tf.math.reduce_mean(head, axis=0) if self.n_head > 1 else head
outputs = self.w_olayer(outputs)
outputs = self.Dropoutlayer2(outputs) # output dropout
return outputs, attn
# + [markdown] id="f04YOJfnF0Eb" pycharm={"name": "#%% md\n"}
# ###TFTFullNetwork
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="Nd09IS9VF58i" outputId="0bde69c8-8f2e-4caa-dfe0-6dc144117054" pycharm={"name": "#%%\n"}
class TFTFullNetwork(tf.keras.Model):
def __init__(self, **kwargs):
super(TFTFullNetwork, self).__init__(**kwargs)
# XXX check TFTSeq TFTNloc UniqueLocations
self.TFTSeq = 0
self.TFTNloc = 0
self.UniqueLocations = []
self.hidden_layer_size = myTFTTools.hidden_layer_size
self.dropout_rate = myTFTTools.dropout_rate
self.num_heads = myTFTTools.num_heads
# New parameters in this TFT version
self.num_static = len(myTFTTools._static_input_loc)
self.num_categorical_variables = len(myTFTTools.category_counts)
self.NumDynamicHistoryVariables = myTFTTools.input_size - self.num_static # Note Future (targets) are also in history
self.num_regular_variables = myTFTTools.input_size - self.num_categorical_variables
self.NumDynamicFutureVariables = 0
for i in myTFTTools._known_regular_input_idx:
if i not in myTFTTools._static_input_loc:
self.NumDynamicFutureVariables += 1
for i in myTFTTools._known_categorical_input_idx:
if i + self.num_regular_variables not in myTFTTools._static_input_loc:
self.NumDynamicFutureVariables += 1
# Embed Categorical Variables
self.CatVariablesembeddings = []
for i in range(0,self.num_categorical_variables):
numcat = self.category_counts[i]
n1 = 'CatEmbed-'+str(i)
n2 = n1 + 'Input ' + str(numcat)
n3 = n1 + 'Map'
n1 = n1 +'Seq'
embedding = tf.keras.Sequential([
tf.keras.layers.InputLayer([myTFTTools.time_steps],name=n2),
tf.keras.layers.Embedding(
numcat,
self.hidden_layer_size,
input_length=myTFTTools.time_steps,
dtype=tf.float32,name=n3)
],name=n1)
self.CatVariablesembeddings.append(embedding)
# Embed Static Variables
numstatic = 0
self.StaticInitialembeddings = []
for i in range(self.num_regular_variables):
if i in myTFTTools._static_input_loc:
n1 = 'StaticRegEmbed-'+str(numstatic)
embedding = tf.keras.layers.Dense(self.hidden_layer_size, name=n1)
self.StaticInitialembeddings.append(embedding)
numstatic += 1
# Embed Targets _input_obs_loc - also included as part of Observed inputs
self.convert_obs_inputs = []
num_obs_inputs = 0
for i in myTFTTools._input_obs_loc:
n1 = 'OBSINPEmbed-Dense-'+str(num_obs_inputs)
n2 = 'OBSINPEmbed-Time-'+str(num_obs_inputs)
embedding = tf.keras.layers.TimeDistributed(tf.keras.layers.Dense(self.hidden_layer_size,name=n1), name=n2)
num_obs_inputs += 1
self.convert_obs_inputs.append(embedding)
# Embed unknown_inputs which are elsewhere called observed inputs
self.convert_unknown_inputs = []
num_unknown_inputs = 0
for i in range(self.num_regular_variables):
if i not in myTFTTools._known_regular_input_idx and i not in myTFTTools._input_obs_loc:
n1 = 'UNKINPEmbed-Dense-'+str(num_unknown_inputs)
n2 = 'UNKINPEmbed-Time-'+str(num_unknown_inputs)
embedding = tf.keras.layers.TimeDistributed(tf.keras.layers.Dense(self.hidden_layer_size,name=n1), name=n2)
num_unknown_inputs += 1
self.convert_unknown_inputs.append(embedding)
# Embed Known Inputs
self.convert_known_regular_inputs = []
num_known_regular_inputs = 0
for i in myTFTTools._known_regular_input_idx:
if i not in myTFTTools._static_input_loc:
n1 = 'KnownINPEmbed-Dense-'+str(num_known_regular_inputs)
n2 = 'KnownINPEmbed-Time-'+str(num_known_regular_inputs)
embedding = tf.keras.layers.TimeDistributed(tf.keras.layers.Dense(self.hidden_layer_size,name=n1), name=n2)
num_known_regular_inputs += 1
self.convert_known_regular_inputs.append(embedding)
# Select Input Static Variables
self.ControlProcessStaticInput = ProcessStaticInput(self.hidden_layer_size,self.dropout_rate, self.num_static)
self.StaticGRN1 = GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate, use_time_distributed=False, GRNname = 'Control1')
self.StaticGRN2 = GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate, use_time_distributed=False, GRNname = 'Control2')
self.StaticGRN3 = GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate, use_time_distributed=False, GRNname = 'Control3')
self.StaticGRN4 = GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate, use_time_distributed=False, GRNname = 'Control4')
# Select Input Dynamic Variables
self.ControlProcessDynamicInput1 = ProcessDynamicInput(self.hidden_layer_size, self.dropout_rate,
self.NumDynamicHistoryVariables, PDIname='Control1')
if myTFTTools.TFTdefaultLSTM:
self.TFTLSTMEncoder = tf.compat.v1.keras.layers.CuDNNLSTM(
self.hidden_layer_size,
return_sequences=True,
return_state=True,
stateful=False,
)
self.TFTLSTMDecoder = tf.compat.v1.keras.layers.CuDNNLSTM(
self.hidden_layer_size,
return_sequences=True,
return_state=False,
stateful=False,
)
else:
self.TFTLSTMEncoder = TFTLSTMLayer( myTFTTools.TFTLSTMEncoderSecondLayer, myTFTTools.TFTLSTMEncoderThirdLayer,
myTFTTools.TFTLSTMEncoderInitialMLP, myTFTTools.TFTLSTMEncoderFinalMLP,
myTFTTools.number_LSTMnodes, self.hidden_layer_size,
myTFTTools.TFTLSTMEncoderactivationvalue, myTFTTools.TFTLSTMEncoderrecurrent_activation,
myTFTTools.TFTLSTMEncoderdropout1, myTFTTools.TFTLSTMEncoderrecurrent_dropout1, TFTreturn_state = True, LSTMname='ControlEncoder')
self.TFTLSTMDecoder = TFTLSTMLayer(myTFTTools.TFTLSTMDecoderSecondLayer, myTFTTools.TFTLSTMDecoderThirdLayer,
myTFTTools.TFTLSTMDecoderInitialMLP, myTFTTools.TFTLSTMDecoderFinalMLP,
myTFTTools.number_LSTMnodes, self.hidden_layer_size,
myTFTTools.TFTLSTMDecoderactivationvalue, myTFTTools.TFTLSTMDecoderrecurrent_activation,
myTFTTools.TFTLSTMDecoderdropout1, myTFTTools.TFTLSTMDecoderrecurrent_dropout1, TFTreturn_state = False, LSTMname='ControlDecoder')
self.TFTFullLSTMGLUplusskip = GLUplusskip(self.hidden_layer_size, self.dropout_rate, activation=None,
use_time_distributed=True, GLUname='ControlLSTM')
self.TemporalGRN5 = GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate, use_additionalcontext = True,
use_time_distributed=True, GRNname = 'Control5')
self.ControlProcessDynamicInput2 = ProcessDynamicInput(self.hidden_layer_size, self.dropout_rate,
self.NumDynamicFutureVariables, PDIname='Control2')
# Decoder self attention
self.TFTself_attn_layer = TFTInterpretableMultiHeadAttention(
self.num_heads, self.hidden_layer_size, self.dropout_rate)
# Set up for final prediction
self.FinalGLUplusskip2 = []
self.FinalGLUplusskip3 = []
self.FinalGRN6 = []
for FinalGatingLoop in range(0, myTFTTools.FinalLoopSize):
self.FinalGLUplusskip2.append(GLUplusskip(self.hidden_layer_size, self.dropout_rate, activation=None,
use_time_distributed=True, GLUname='ControlFinal2-'+str(FinalGatingLoop)))
self.FinalGLUplusskip3.append(GLUplusskip(self.hidden_layer_size, self.dropout_rate, activation=None,
use_time_distributed=True, GLUname='ControlFinal3-'+str(FinalGatingLoop)))
self.FinalGRN6.append(GRN(self.hidden_layer_size, dropout_rate=self.dropout_rate, use_time_distributed=True, GRNname = 'Control6-'+str(FinalGatingLoop)))
# Final Processing
if myTFTTools.TFTLSTMFinalMLP > 0:
self.FinalApplyMLP = apply_mlp(myTFTTools.TFTLSTMFinalMLP, output_size = myTFTTools.output_size * myTFTTools.NumberQuantiles,
output_activation = None, hidden_activation = 'selu',
use_time_distributed = True, MLPname='Predict')
else:
if myTFTTools.FinalLoopSize == 1:
n1 = 'FinalTD'
n2 = 'FinalDense'
self.FinalLayer = tf.keras.layers.TimeDistributed(
tf.keras.layers.Dense(myTFTTools.output_size * myTFTTools.NumberQuantiles, name = n2), name =n1)
else:
self.FinalStack =[]
localloopsize = myTFTTools.output_size * myTFTTools.NumberQuantiles
for localloop in range(0,localloopsize):
self.FinalStack.append(tf.keras.layers.Dense(1))
# Called with each batch as input
#<EMAIL>
def call(self, all_inputs, ignoredtime, ignoredidentifiers, training=None):
# ignoredtime, ignoredidentifiers not used
time_steps = myTFTTools.time_steps
combined_input_size = myTFTTools.input_size
encoder_steps = myTFTTools.num_encoder_steps
# Sanity checks on inputs
for InputIndex in myTFTTools._known_regular_input_idx:
if InputIndex in myTFTTools._input_obs_loc:
printexit('Observation cannot be known a priori!' + str(InputIndex))
for InputIndex in myTFTTools._input_obs_loc:
if InputIndex in myTFTTools._static_input_loc:
printexit('Observation cannot be static!' + str(InputIndex))
Sizefrominputs = all_inputs.get_shape().as_list()[-1]
if Sizefrominputs != myTFTTools.input_size:
raise printexit(f'Illegal number of inputs! Inputs observed={Sizefrominputs}, expected={myTFTTools.input_size}')
regular_inputs, categorical_inputs = all_inputs[:, :, :self.num_regular_variables], all_inputs[:, :, self.num_regular_variables:]
# Embed categories of all categorical variables -- static and Dynamic
# categorical variables MUST be at end and reordering done in preprocessing (definition of train valid test)
# XXX add reordering
categoricalembedded_inputs = []
for i in range(0,self.num_categorical_variables):
categoricalembedded_inputs.append( CatVariablesembeddings[i](categorical_inputs[Ellipsis, i]) )
# Complete Static Variables -- whether categorical or regular -- they are essentially thought of as known inputs
if myTFTTools._static_input_loc:
static_inputs = []
numstatic = 0
for i in range(self.num_regular_variables):
if i in myTFTTools._static_input_loc:
static_inputs.append(self.StaticInitialembeddings[numstatic](regular_inputs[:, 0, i:i + 1]) )
numstatic += 1
static_inputs = static_inputs + [self.categoricalembedded_inputs[i][:, 0, :]
for i in range(self.num_categorical_variables)
if i + self.num_regular_variables in myTFTTools._static_input_loc]
static_inputs = tf.stack(static_inputs, axis=1)
else:
static_inputs = None
# Targets misleadingly labelled obs_inputs. They are used as targets to predict and as observed inputs
obs_inputs = []
num_obs_inputs = 0
for i in myTFTTools._input_obs_loc:
e = self.convert_obs_inputs[num_obs_inputs](regular_inputs[Ellipsis, i:i + 1])
num_obs_inputs += 1
obs_inputs.append(e)
obs_inputs = tf.stack(obs_inputs, axis=-1)
# Categorical Unknown inputs. Unknown + Target is complete Observed InputCategory
categorical_unknown_inputs = []
for i in range(self.num_categorical_variables):
if i not in myTFTTools._known_categorical_input_idx and i + self.num_regular_variables not in myTFTTools._input_obs_loc:
e = self.categoricalembedded_inputs[i]
categorical_unknown_inputs.append(e)
# Regular Unknown inputs
unknown_inputs = []
num_unknown_inputs = 0
for i in range(self.num_regular_variables):
if i not in myTFTTools._known_regular_input_idx and i not in myTFTTools._input_obs_loc:
e = self.convert_unknown_inputs[num_unknown_inputs](regular_inputs[Ellipsis, i:i + 1])
num_unknown_inputs += 1
unknown_inputs.append(e)
# Add in categorical_unknown_inputs into unknown_inputs
if unknown_inputs + categorical_unknown_inputs:
unknown_inputs = tf.stack(unknown_inputs + categorical_unknown_inputs, axis=-1)
else:
unknown_inputs = None
# A priori known inputs
known_regular_inputs = []
num_known_regular_inputs = 0
for i in myTFTTools._known_regular_input_idx:
if i not in myTFTTools._static_input_loc:
e = self.convert_known_regular_inputs[num_known_regular_inputs](regular_inputs[Ellipsis, i:i + 1])
num_known_regular_inputs += 1
known_regular_inputs.append(e)
known_categorical_inputs = []
for i in myTFTTools._known_categorical_input_idx:
if i + self.num_regular_variables not in myTFTTools._static_input_loc:
e = categoricalembedded_inputs[i]
known_categorical_inputs.append(e)
known_combined_layer = tf.stack(known_regular_inputs + known_categorical_inputs, axis=-1)
# Now we know unknown_inputs, known_combined_layer, obs_inputs, static_inputs
# Identify known and observed historical_inputs.
if unknown_inputs is not None:
historical_inputs = tf.concat([
unknown_inputs[:, :encoder_steps, :],
known_combined_layer[:, :encoder_steps, :],
obs_inputs[:, :encoder_steps, :]
], axis=-1)
else:
historical_inputs = tf.concat([
known_combined_layer[:, :encoder_steps, :],
obs_inputs[:, :encoder_steps, :]
], axis=-1)
# Identify known future inputs.
future_inputs = known_combined_layer[:, encoder_steps:, :]
# Process Static Variables
static_encoder, static_weights = self.ControlProcessStaticInput(static_inputs)
static_context_variable_selection = self.StaticGRN1(static_encoder)
static_context_enrichment = self.StaticGRN2(static_encoder)
static_context_state_h = self.StaticGRN3(static_encoder)
static_context_state_c = self.StaticGRN4(static_encoder)
# End set up of static variables
historical_features, historical_flags, _ = self.ControlProcessDynamicInput1(historical_inputs,
static_context_variable_selection = static_context_variable_selection)
history_lstm, state_h, state_c = self.TFTLSTMEncoder(historical_features, initial_state = [static_context_state_h, static_context_state_c])
input_embeddings = historical_features
lstm_layer = history_lstm
future_features, future_flags, _ = self.ControlProcessDynamicInput2(future_inputs, static_context_variable_selection = static_context_variable_selection)
future_lstm = self.TFTLSTMDecoder(future_features, initial_state= [state_h, state_c])
input_embeddings = tf.concat([historical_features, future_features], axis=1)
lstm_layer = tf.concat([history_lstm, future_lstm], axis=1)
temporal_feature_layer, _ = self.TFTFullLSTMGLUplusskip(lstm_layer, input_embeddings)
expanded_static_context = tf.expand_dims(static_context_enrichment, axis=1) # Add fake time axis
enriched = self.TemporalGRN5(temporal_feature_layer, additional_context=expanded_static_context, return_gate=False)
# Calculate attention
# mask does not use "time" as implicit in order of entries in window
mask = TFTget_decoder_mask(enriched)
x, self_att = self.TFTself_attn_layer(enriched, enriched, enriched, mask=mask)
if myTFTTools.FinalLoopSize > 1:
StackLayers = []
for FinalGatingLoop in range(0, myTFTTools.FinalLoopSize):
x, _ = self.FinalGLUplusskip2[FinalGatingLoop](x,enriched)
# Nonlinear processing on outputs
decoder = self.FinalGRN6[FinalGatingLoop](x)
# Final skip connection
transformer_layer, _ = self.FinalGLUplusskip3[FinalGatingLoop](decoder, temporal_feature_layer)
if myTFTTools.FinalLoopSize > 1:
StackLayers.append(transformer_layer)
# End Loop over FinalGatingLoop
if myTFTTools.FinalLoopSize > 1:
transformer_layer = tf.stack(StackLayers, axis=-1)
# Attention components for explainability IGNORED
attention_components = {
# Temporal attention weights
'decoder_self_attn': self_att,
# Static variable selection weights
'static_flags': static_weights[Ellipsis, 0],
# Variable selection weights of past inputs
'historical_flags': historical_flags[Ellipsis, 0, :],
# Variable selection weights of future inputs
'future_flags': future_flags[Ellipsis, 0, :]
}
self._attention_components = attention_components
# Original split procerssing here and did
# return transformer_layer, all_inputs, attention_components
if myTFTTools.TFTLSTMFinalMLP > 0:
outputs = self.FinalApplyMLP(transformer_layer[Ellipsis, encoder_steps:, :])
else:
if myTFTTools.FinalLoopSize == 1:
outputs = self.FinalLayer(transformer_layer[Ellipsis, encoder_steps:, :])
else:
outputstack =[]
localloopsize = myTFTTools.output_size * myTFTTools.NumberQuantiles
for localloop in range(0,localloopsize):
localoutput = self.FinalStack[localloop](transformer_layer[Ellipsis, encoder_steps:, :, localloop])
outputstack.append(localoutput)
outputs = tf.stack(outputstack, axis=-2)
outputs = tf.squeeze(outputs, axis=-1)
return outputs
# + [markdown] id="ckwZHK12xwCY" pycharm={"name": "#%% md\n"}
# ##TFT Run & Output
# + [markdown] id="23a1qMuO_yVT" pycharm={"name": "#%% md\n"}
# ### General Utilities
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="awPqzc4H_yVh" outputId="38fb0abf-ec98-46d7-839b-611faf76d5d5" pycharm={"name": "#%%\n"}
def get_model_summary(model):
stream = io.StringIO()
model.summary(print_fn=lambda x: stream.write(x + '\n'))
summary_string = stream.getvalue()
stream.close()
return summary_string
def setDLinput(Spacetime = True):
# Initial data is Flatten([Num_Seq][Nloc]) [Tseq] with values [Nprop-Sel + Nforcing + Add(ExPosEnc-Selin)] starting with RawInputSequencesTOT
# Predictions are Flatten([Num_Seq] [Nloc]) [Predvals=Npred+ExPosEnc-Selout] [Predtimes = Forecast-time range] starting with RawInputPredictionsTOT
# No assumptions as to type of variables here
if SymbolicWindows:
X_predict = SymbolicInputSequencesTOT.reshape(OuterBatchDimension,1,1)
else:
X_predict = RawInputSequencesTOT.reshape(OuterBatchDimension,Tseq,NpropperseqTOT)
y_predict = RawInputPredictionsTOT.reshape(OuterBatchDimension,NpredperseqTOT)
if Spacetime:
SpacetimeforMask_predict = SpacetimeforMask.reshape(OuterBatchDimension,1,1).copy()
return X_predict, y_predict, SpacetimeforMask_predict
return X_predict, y_predict
def setSeparateDLinput(model, Spacetime = False):
# Initial data is Flatten([Num_Seq][Nloc]) [Tseq] with values [Nprop-Sel + Nforcing + Add(ExPosEnc-Selin)] starting with RawInputSequencesTOT
# Predictions are Flatten([Num_Seq] [Nloc]) [Predvals=Npred+ExPosEnc-Selout] [Predtimes = Forecast-time range] starting with RawInputPredictionsTOT
# No assumptions as to type of variables here
# model = 0 LSTM =1 transformer
if model == 0:
Spacetime = False
X_val = None
y_val = None
Spacetime_val = None
Spacetime_train = None
if SymbolicWindows:
InputSequences = np.empty([Num_Seq, TrainingNloc], dtype = np.int32)
for iloc in range(0,TrainingNloc):
InputSequences[:,iloc] = SymbolicInputSequencesTOT[:,ListofTrainingLocs[iloc]]
if model == 0:
X_train = InputSequences.reshape(Num_Seq*TrainingNloc,1,1)
else:
X_train = InputSequences
if Spacetime:
Spacetime_train = X_train.copy()
if LocationValidationFraction > 0.001:
UsedValidationNloc = ValidationNloc
if FullSetValidation:
UsedValidationNloc = Nloc
ValInputSequences = np.empty([Num_Seq, UsedValidationNloc], dtype = np.int32)
if FullSetValidation:
for iloc in range(0,Nloc):
ValInputSequences[:,iloc] = SymbolicInputSequencesTOT[:,iloc]
else:
for iloc in range(0,ValidationNloc):
ValInputSequences[:,iloc] = SymbolicInputSequencesTOT[:,ListofValidationLocs[iloc]]
if model == 0:
X_val = ValInputSequences.reshape(Num_Seq * UsedValidationNloc,1,1)
else:
X_val = ValInputSequences
if Spacetime:
Spacetime_val = X_val.copy()
else: # Symbolic Windows false Calculate Training
InputSequences = np.empty([Num_Seq, TrainingNloc,Tseq,NpropperseqTOT], dtype = np.float32)
for iloc in range(0,TrainingNloc):
InputSequences[:,iloc,:,:] = RawInputSequencesTOT[:,ListofTrainingLocs[iloc],:,:]
if model == 0:
X_train = InputSequences.reshape(Num_Seq*TrainingNloc,Tseq,NpropperseqTOT)
else:
X_train = InputSequences
if Spacetime:
Spacetime_train = np.empty([Num_Seq, TrainingNloc], dtype = np.int32)
for iloc in range(0,TrainingNloc):
Spacetime_train[:,iloc] = SpacetimeforMask[:,ListofTrainingLocs[iloc]]
if LocationValidationFraction > 0.001: # Symbolic Windows false Calculate Validation
UsedValidationNloc = ValidationNloc
if FullSetValidation:
UsedValidationNloc = Nloc
ValInputSequences = np.empty([Num_Seq, UsedValidationNloc,Tseq,NpropperseqTOT], dtype = np.float32)
if FullSetValidation:
for iloc in range(0,Nloc):
ValInputSequences[:,iloc,:,:] = RawInputSequencesTOT[:,iloc,:,:]
else:
for iloc in range(0,ValidationNloc):
ValInputSequences[:,iloc,:,:] = RawInputSequencesTOT[:,ListofValidationLocs[iloc],:,:]
if model == 0:
X_val = ValInputSequences.reshape(Num_Seq * UsedValidationNloc,Tseq,NpropperseqTOT)
else:
X_val = ValInputSequences
if Spacetime:
Spacetime_val = np.empty([Num_Seq, UsedValidationNloc], dtype = np.int32)
if FullSetValidation:
for iloc in range(0,Nloc):
Spacetime_val[:,iloc] = SpacetimeforMask[:,iloc]
else:
for iloc in range(0,ValidationNloc):
Spacetime_val[:,iloc] = SpacetimeforMask[:,ListofValidationLocs[iloc]]
# Calculate training predictions
InputPredictions = np.empty([Num_Seq, TrainingNloc,NpredperseqTOT], dtype = np.float32)
for iloc in range(0,TrainingNloc):
InputPredictions[:,iloc,:] = RawInputPredictionsTOT[:,ListofTrainingLocs[iloc],:]
if model == 0:
y_train = InputPredictions.reshape(OuterBatchDimension,NpredperseqTOT)
else:
y_train = InputPredictions
# Calculate validation predictions
if LocationValidationFraction > 0.001:
ValInputPredictions = np.empty([Num_Seq, UsedValidationNloc,NpredperseqTOT], dtype = np.float32)
if FullSetValidation:
for iloc in range(0,Nloc):
ValInputPredictions[:,iloc,:] = RawInputPredictionsTOT[:,iloc,:]
else:
for iloc in range(0,ValidationNloc):
ValInputPredictions[:,iloc,:] = RawInputPredictionsTOT[:,ListofValidationLocs[iloc],:]
if model == 0:
y_val = ValInputPredictions.reshape(Num_Seq * ValidationNloc,NpredperseqTOT)
else:
y_val = ValInputPredictions
if Spacetime:
return X_train, y_train, Spacetime_train, X_val, y_val, Spacetime_val
else:
return X_train, y_train,X_val,y_val
def InitializeDLforTimeSeries(message,processindex,y_predict):
if processindex == 0:
current_time = timenow()
line = (startbold + current_time + ' ' + message + resetfonts + " Window Size " + str(Tseq) +
" Number of samples over time that sequence starts at and location:" +str(OuterBatchDimension) +
" Number input features per sequence:" + str(NpropperseqTOT) +
" Number of predicted outputs per sequence:" + str(NpredperseqTOT) +
" Batch_size:" + str(LSTMbatch_size) +
" n_nodes:" + str(number_LSTMnodes) +
" epochs:" + str(TFTTransformerepochs))
print(wraptotext(line))
checkNaN(y_predict)
# + [markdown] id="42UqTW0xDoKr" pycharm={"name": "#%% md\n"}
# ### Tensorflow Monitor
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="rIocfMfZBjfa" outputId="ffdbd7c6-9640-405c-90ac-754d2ac72fa5" pycharm={"name": "#%%\n"}
class TensorFlowTrainingMonitor:
def __init__(self):
# These OPERATIONAL variables control saving of best fits
self.lastsavedepoch = -1 # Epoch number where last saved fit done
self.BestLossValueSaved = NaN # Training Loss value of last saved fit
self.BestValLossValueSaved = NaN # Validation Loss value of last saved fit
self.Numsuccess = 0 # count little successes up to SuccessLimit
self.Numfailed = 0
self.LastLossValue = NaN # Loss on previous epoch
self.MinLossValue = NaN # Saved minimum loss value
self.LastValLossValue = NaN # Validation Loss on previous epoch
self.MinValLossValue = NaN # validation loss value at last save
self.BestLossSaved = False # Boolean to indicate that best Loss value saved
self.saveMinLosspath = '' # Checkpoint path for saved network
self.epochcount = 0
self.NumberTimesSaved = 0 # Number of Checkpointing steps for Best Loss
self.NumberTimesRestored = 0 # Number of Checkpointing Restores
self.LittleJumpdifference = NaN
self.LittleValJumpdifference = NaN
self.AccumulateSuccesses = 0
self.AccumulateFailures = np.zeros(5, dtype=int)
self.RestoreReasons = np.zeros(8, dtype = int)
self.NameofFailures = ['Success','Train Only Failed','Val Only Failed','Both Failed', 'NaN']
self.NameofRestoreReasons = ['Both Big Jump', 'Both Little Jump','Train Big Jump', 'Train Little Jump','Val Big Jump','Val Little Jump',' Failure Limit', ' NaN']
# End OPERATIONAL Control set up for best fit checkpointing
# These are parameters user can set
self.UseBestAvailableLoss = True
self.LittleJump = 2.0 # Multiplier for checking jump compared to recent changes
self.ValLittleJump = 2.0 # Multiplier for checking jump compared to recent changes
self.startepochs = -1 # Ignore this number of epochs to let system get started
self.SuccessLimit = 20 # Don't keep saving. Wait for this number of (little) successes
self.FailureLimit = 10 # Number of failures before restore
self.BadJumpfraction = 0.2 # This fractional jump will trigger attempt to go back to saved value
self.ValBadJumpfraction = 0.2 # This fractional jump will trigger attempt to go back to saved value
self.ValidationFraction = 0.0 # Must be used validation fraction
DownplayValidationIncrease = True
# End parameters user can set
self.checkpoint = None
self.CHECKPOINTDIR = ''
self.RunName = ''
self.train_epoch = 0.0
self.val_epoch = 0.0
tfepochstep = None
recordtrainloss =[]
recordvalloss = []
def SetControlParms(self, UseBestAvailableLoss = None, LittleJump = None, startepochs = None, ValLittleJump = None,
ValBadJumpfraction = None, SuccessLimit = None, FailureLimit = None, BadJumpfraction = None, DownplayValidationIncrease=True):
if UseBestAvailableLoss is not None:
self.UseBestAvailableLoss = UseBestAvailableLoss
if LittleJump is not None:
self.LittleJump = LittleJump
if ValLittleJump is not None:
self.ValLittleJump = ValLittleJump
if startepochs is not None:
self.startepochs = startepochs
if SuccessLimit is not None:
self.SuccessLimit = SuccessLimit
if FailureLimit is not None:
self.FailureLimit = FailureLimit
if BadJumpfraction is not None:
self.BadJumpfraction = BadJumpfraction
if ValBadJumpfraction is not None:
self.ValBadJumpfraction = ValBadJumpfraction
if DownplayValidationIncrease:
self.ValBadJumpfraction = 200.0
self.ValLittleJump = 2000.0
elif ValLittleJump is None:
self.ValLittleJump = 2.0
elif ValBadJumpfraction is None:
self.ValBadJumpfraction = 0.2
def SetCheckpointParms(self,checkpointObject,CHECKPOINTDIR,RunName = '',Restoredcheckpoint= False, Restored_path = '',
ValidationFraction = 0.0, SavedTrainLoss = NaN, SavedValLoss = NaN):
self.ValidationFraction = ValidationFraction
self.checkpoint = checkpointObject
self.CHECKPOINTDIR = CHECKPOINTDIR
self.RunName = RunName
if Restoredcheckpoint:
self.BestLossSaved = True
self.saveMinLosspath = Restored_path # Checkpoint path for saved network
self.LastLossValue = SavedTrainLoss
self.LastValLossValue = SavedValLoss
self.BestLossValueSaved = SavedTrainLoss
self.BestValLossValueSaved = SavedValLoss
self.lastsavedepoch = self.epochcount
self.MinLossValue = SavedTrainLoss
self.MinValLossValue = SavedValLoss
def EpochEvaluate(self, epochcount,train_epoch, val_epoch, tfepochstep, recordtrainloss, recordvalloss):
FalseReturn = 0
TrueReturn = 1
self.epochcount = epochcount
self.train_epoch = train_epoch
self.val_epoch = val_epoch
self.tfepochstep = tfepochstep
self.recordtrainloss = recordtrainloss
self.recordvalloss = recordvalloss
Needtorestore = False
Failreason = 5 # nonsense
LossChange = 0.0
ValLossChange = 0.0
if np.math.isnan(self.train_epoch) or np.math.isnan(self.val_epoch):
Restoreflag = 7
self.RestoreReasons[Restoreflag] += 1
Needtorestore = True
Failreason = 4
self.AccumulateFailures[Failreason] += 1
print(str(self.epochcount) + ' NAN Seen Reason ' + str(Failreason) + ' #succ ' + str(self.Numsuccess) + ' #fail ' + str(self.Numfailed) + ' ' + str(round(self.train_epoch,6)) + ' ' + str(round(self.val_epoch,6)), flush=True)
return TrueReturn, self.train_epoch, self.val_epoch
if self.epochcount <= self.startepochs:
return FalseReturn, self.train_epoch, self.val_epoch
if not np.math.isnan(self.LastLossValue):
LossChange = self.train_epoch - self.LastLossValue
if self.ValidationFraction > 0.001:
ValLossChange = self.val_epoch - self.LastValLossValue
if LossChange <= 0:
if self.ValidationFraction > 0.001:
# Quick Fix
self.Numsuccess +=1
self.AccumulateSuccesses += 1
if ValLossChange <= 0:
Failreason = 0
else:
Failreason = 2
else:
self.Numsuccess +=1
self.AccumulateSuccesses += 1
Failreason = 0
else:
Failreason = 1
if self.ValidationFraction > 0.001:
if ValLossChange > 0:
Failreason = 3
if Failreason > 0:
self.Numfailed += 1
self.AccumulateFailures[Failreason] += 1
if (not np.math.isnan(self.LastLossValue)) and (Failreason > 0):
print(str(self.epochcount) + ' Reason ' + str(Failreason) + ' #succ ' + str(self.Numsuccess) + ' #fail ' + str(self.Numfailed) + ' ' + str(round(self.train_epoch,6))
+ ' ' + str(round(self.LastLossValue,6)) + ' '+ str(round(self.val_epoch,6))+ ' ' + str(round(self.LastValLossValue,6)), flush=True)
self.LastLossValue = self.train_epoch
self.LastValLossValue = self.val_epoch
StoreMinLoss = False
if not np.math.isnan(self.MinLossValue):
# if (self.train_epoch < self.MinLossValue) and (self.val_epoch <= self.MinValLossValue):
if self.train_epoch < self.MinLossValue:
if self.Numsuccess >= self.SuccessLimit:
StoreMinLoss = True
else:
StoreMinLoss = True
if StoreMinLoss:
self.Numsuccess = 0
extrastuff = ''
extrastuff_val = ' '
if not np.math.isnan(self.MinLossValue):
extrastuff = ' Previous ' + str(round(self.MinLossValue,7))
self.LittleJumpdifference = self.MinLossValue - self.train_epoch
if self.ValidationFraction > 0.001:
if not np.math.isnan(self.MinValLossValue):
extrastuff_val = ' Previous ' + str(round(self.MinValLossValue,7))
LittleValJumpdifference = max(self.MinValLossValue - self.val_epoch, self.LittleJumpdifference)
self.saveMinLosspath = self.checkpoint.save(file_prefix=self.CHECKPOINTDIR + self.RunName +'MinLoss')
if not self.BestLossSaved:
print('\nInitial Checkpoint at ' + self.saveMinLosspath + ' from ' + self.CHECKPOINTDIR)
self.MinLossValue = self.train_epoch
self.MinValLossValue = self.val_epoch
if self.ValidationFraction > 0.001:
extrastuff_val = ' Val Loss ' + str(round(self.val_epoch,7)) + extrastuff_val
print(' Epoch ' + str(self.epochcount) + ' Loss ' + str(round(self.train_epoch,7)) + extrastuff + extrastuff_val+ ' Failed ' + str(self.Numfailed), flush = True)
self.Numfailed = 0
self.BestLossSaved = True
self.BestLossValueSaved = self.train_epoch
self.BestValLossValueSaved = self.val_epoch
self.lastsavedepoch = self.epochcount
self.NumberTimesSaved += 1
return FalseReturn, self.train_epoch, self.val_epoch
RestoreTrainflag = -1
Trainrestore = False
if LossChange > 0.0:
if LossChange > self.BadJumpfraction * self.train_epoch:
Trainrestore = True
RestoreTrainflag = 0
if not np.math.isnan(self.LittleJumpdifference):
if LossChange > self.LittleJumpdifference * self.LittleJump:
Trainrestore = True
if RestoreTrainflag < 0:
RestoreTrainflag = 1
if self.BestLossSaved:
if self.train_epoch < self.MinLossValue:
Trainrestore = False
RestoreTrainflag = -1
RestoreValflag = -1
Valrestore = False
if ValLossChange > 0.0:
if ValLossChange > self.ValBadJumpfraction * self.val_epoch:
Valrestore = True
RestoreValflag = 0
if not np.math.isnan(self.LittleValJumpdifference):
if ValLossChange > self.LittleValJumpdifference * self.ValLittleJump:
Valrestore = True
if RestoreValflag < 0:
RestoreValflag = 1
if self.BestLossSaved:
if self.val_epoch < self.MinValLossValue:
Valrestore = False
RestoreValflag = -1
Restoreflag = -1
if Trainrestore and Valrestore:
Needtorestore = True
if RestoreTrainflag == 0:
Restoreflag = 0
else:
Restoreflag = 1
elif Trainrestore:
Needtorestore = True
Restoreflag = RestoreTrainflag + 2
elif Valrestore:
Needtorestore = True
Restoreflag = RestoreValflag + 4
if (self.Numfailed >= self.FailureLimit) and (Restoreflag == -1):
Restoreflag = 6
Needtorestore = True
if Restoreflag >= 0:
self.RestoreReasons[Restoreflag] += 1
if Needtorestore and (not self.BestLossSaved):
print('bad Jump ' + str(round(LossChange,7)) + ' Epoch ' + str(self.epochcount) + ' But nothing saved')
return FalseReturn, self.train_epoch, self.val_epoch
if Needtorestore:
return TrueReturn, self.train_epoch, self.val_epoch
else:
return FalseReturn, self.train_epoch, self.val_epoch
def RestoreBestFit(self):
if self.BestLossSaved:
self.checkpoint.tfrecordvalloss = tf.Variable([], shape =tf.TensorShape(None), trainable = False)
self.checkpoint.tfrecordtrainloss = tf.Variable([], shape =tf.TensorShape(None), trainable = False)
self.checkpoint.restore(save_path=self.saveMinLosspath).expect_partial()
self.tfepochstep = self.checkpoint.tfepochstep
self.recordvalloss = self.checkpoint.tfrecordvalloss.numpy().tolist()
self.recordtrainloss = self.checkpoint.tfrecordtrainloss.numpy().tolist()
trainlen = len(self.recordtrainloss)
self.Numsuccess = 0
extrastuff = ''
if self.ValidationFraction > 0.001:
vallen =len(self.recordvalloss)
if vallen > 0:
extrastuff = ' Replaced Val Loss ' + str(round(self.recordvalloss[vallen-1],7))+ ' bad val ' + str(round(self.val_epoch,7))
else:
extrastuff = ' No previous Validation Loss'
print(str(self.epochcount) + ' Failed ' + str(self.Numfailed) + ' Restored Epoch ' + str(trainlen-1) + ' Replaced Loss ' + str(round(self.recordtrainloss[trainlen-1],7))
+ ' bad ' + str(round(self.train_epoch,7)) + extrastuff + ' Checkpoint at ' + self.saveMinLosspath)
self.train_epoch = self.recordtrainloss[trainlen-1]
self.Numfailed = 0
self.LastLossValue = self.train_epoch
self.NumberTimesRestored += 1
if self.ValidationFraction > 0.001:
vallen = len(self.recordvalloss)
if vallen > 0:
self.val_epoch = self.recordvalloss[vallen-1]
else:
self.val_epoch = 0.0
return self.tfepochstep, self.recordtrainloss, self.recordvalloss, self.train_epoch, self.val_epoch
def PrintEndofFit(self, Numberofepochs):
print(startbold + 'Number of Saves ' + str(self.NumberTimesSaved) + ' Number of Restores ' + str(self.NumberTimesRestored))
print('Epochs Requested ' + str(Numberofepochs) + ' Actually Stored ' + str(len(self.recordtrainloss)) + ' ' + str(self.tfepochstep.numpy())
+ ' Successes ' +str(self.AccumulateSuccesses) + resetfonts)
trainlen = len(self.recordtrainloss)
train_epoch1 = self.recordtrainloss[trainlen-1]
lineforval = ''
if self.ValidationFraction > 0.001:
lineforval = ' Last val '+ str(round(self.val_epoch,7))
print(startbold + 'Last loss '+ str(round(self.train_epoch,7)) + ' Last loss in History ' + str(round(train_epoch1,7))+ ' Best Saved Loss '
+ str(round(self.BestLossValueSaved,7)) + lineforval + resetfonts)
print(startbold + startred +"\nFailure Reasons" + resetfonts)
for ireason in range(0,len(self.AccumulateFailures)):
print('Optimization Failure ' + str(ireason) + ' ' + self.NameofFailures[ireason] + ' ' + str(self.AccumulateFailures[ireason]))
print(startbold + startred +"\nRestore Reasons" + resetfonts)
for ireason in range(0,len(self.RestoreReasons)):
print('Backup to earlier fit ' + str(ireason) + ' ' + self.NameofRestoreReasons[ireason] + ' ' + str(self.RestoreReasons[ireason]))
def BestPossibleFit(self): # Use Best Saved if appropriate
if self.UseBestAvailableLoss:
if self.BestLossSaved:
if self.BestLossValueSaved < self.train_epoch:
self.checkpoint.tfrecordvalloss = tf.Variable([], shape =tf.TensorShape(None), trainable = False)
self.checkpoint.tfrecordtrainloss = tf.Variable([], shape =tf.TensorShape(None), trainable = False)
self.checkpoint.restore(save_path=self.saveMinLosspath).expect_partial()
self.tfepochstep = self.checkpoint.tfepochstep
self.recordvalloss = self.checkpoint.tfrecordvalloss.numpy().tolist()
self.recordtrainloss = self.checkpoint.tfrecordtrainloss.numpy().tolist()
trainlen = len(self.recordtrainloss)
Oldtraining = self.train_epoch
self.train_epoch = self.recordtrainloss[trainlen-1]
extrainfo = ''
if self.ValidationFraction > 0.001:
vallen = len(self.recordvalloss)
if vallen > 0:
extrainfo = '\nVal Loss ' + str(round(self.recordvalloss[vallen-1],7)) + ' old Val ' + str(round(self.val_epoch,7))
self.val_epoch = self.recordvalloss[vallen-1]
else:
self.val_epoch = 0.0
extrainfo = '\n no previous validation loss'
print(startpurple+ startbold + 'Switch to Best Saved Value. Restored Epoch ' + str(trainlen-1)
+ '\nNew Loss ' + str(round(self.recordtrainloss[trainlen-1],7)) + ' old ' + str(round(Oldtraining,7))
+ extrainfo + '\nCheckpoint at ' + self.saveMinLosspath + resetfonts)
else:
print(startpurple+ startbold + '\nFinal fit is best: train ' + str(round(self.train_epoch,7)) + ' Val Loss ' + str(round(self.val_epoch,7)) + resetfonts)
return self.tfepochstep, self.recordtrainloss, self.recordvalloss, self.train_epoch, self.val_epoch
# + [markdown] id="bV-u9bfz6OIL" pycharm={"name": "#%% md\n"}
# ###TFT Output
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="01KS6sEG6VSw" outputId="d2f5f3df-dc85-46bd-a7f1-56f33bcb0ac5" pycharm={"name": "#%%\n"}
def TFTTestpredict(custommodel,datacollection):
"""Computes predictions for a given input dataset.
Args:
df: Input dataframe
return_targets: Whether to also return outputs aligned with predictions to
faciliate evaluation
Returns:
Input dataframe or tuple of (input dataframe, algined output dataframe).
"""
inputs = datacollection['inputs']
time = datacollection['time']
identifier = datacollection['identifier']
outputs = datacollection['outputs']
combined = None
OuterBatchDimension = inputs.shape[0]
batchsize = myTFTTools.maxibatch_size
numberoftestbatches = math.ceil(OuterBatchDimension/batchsize)
count1 = 0
for countbatches in range(0,numberoftestbatches):
count2 = min(OuterBatchDimension, count1+batchsize)
if count2 <= count1:
continue
samples = np.arange(count1,count2)
count1 += batchsize
X_test = inputs[samples,Ellipsis]
time_test = []
id_test =[]
Numinbatch = X_test.shape[0]
if myTFTTools.TFTSymbolicWindows:
X_test = X_test.numpy()
X_test = np.reshape(X_test,Numinbatch)
iseqarray = np.right_shift(X_test,16)
ilocarray = np.bitwise_and(X_test, 0b1111111111111111)
X_testFull = list()
for iloc in range(0,Numinbatch):
X_testFull.append(ReshapedSequencesTOT[ilocarray[iloc],iseqarray[iloc]:iseqarray[iloc]+Tseq])
X_test = np.array(X_testFull)
batchprediction = custommodel(X_test, time_test, id_test, training=False).numpy()
if combined is None:
combined = batchprediction
else:
combined = np.concatenate((combined, batchprediction),axis=0)
def format_outputs(prediction):
"""Returns formatted dataframes for prediction."""
reshapedprediction = prediction.reshape(prediction.shape[0], -1)
flat_prediction = pd.DataFrame(
reshapedprediction[:, :],
columns=[
't+{}-Obs{}'.format(i, j)
for i in range(myTFTTools.time_steps - myTFTTools.num_encoder_steps)
for j in range(0, myTFTTools.output_size)
])
cols = list(flat_prediction.columns)
flat_prediction['forecast_time'] = time[:,
myTFTTools.num_encoder_steps - 1, 0]
flat_prediction['identifier'] = identifier[:, 0, 0]
# Arrange in order
return flat_prediction[['forecast_time', 'identifier'] + cols]
# Extract predictions for each quantile into different entries
process_map = {
qname:
combined[Ellipsis, i * myTFTTools.output_size:(i + 1) * myTFTTools.output_size]
for i, qname in enumerate(myTFTTools.Quantilenames)
}
process_map['targets'] = outputs
return {k: format_outputs(process_map[k]) for k in process_map}
# Simple Plot of Loss from history
def finalizeTFTDL(ActualModel, recordtrainloss, recordvalloss, validationfrac, test_datacollection, modelflag, LabelFit =''):
# Ouput Loss v Epoch
histlen = len(recordtrainloss)
trainloss = recordtrainloss[histlen-1]
plt.rcParams["figure.figsize"] = [8,6]
plt.plot(recordtrainloss)
if (validationfrac > 0.001) and len(recordvalloss) > 0:
valloss = recordvalloss[histlen-1]
plt.plot(recordvalloss)
else:
valloss = 0.0
current_time = timenow()
print(startbold + startred + current_time + ' ' + RunName + ' finalizeDL ' + RunComment +resetfonts)
plt.title(LabelFit + ' ' + RunName+' model loss ' + str(round(trainloss,7)) + ' Val ' + str(round(valloss,7)))
plt.ylabel('loss')
plt.xlabel('epoch')
plt.yscale("log")
plt.grid(True)
plt.legend(['train', 'val'], loc='upper left')
plt.show()
# Setup TFT
if modelflag == 2:
global SkipDL2F, IncreaseNloc_sample, DecreaseNloc_sample
SkipDL2F = True
IncreaseNloc_sample = 1
DecreaseNloc_sample = 1
TFToutput_map = TFTTestpredict(ActualModel,test_datacollection)
VisualizeTFT(ActualModel, TFToutput_map)
else:
printexit("unsupported model " +str(modelflag))
# + [markdown] id="ysYaHvNAuxFe" pycharm={"name": "#%% md\n"}
# ###TFTcustommodel
#
# Control Full TFT Network
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="F_kBmZ0xu3eW" outputId="80ab63fb-c99e-4734-a921-b701ef2581df" pycharm={"name": "#%%\n"}
class TFTcustommodel(tf.keras.Model):
def __init__(self, **kwargs):
super(TFTcustommodel, self).__init__(**kwargs)
self.myTFTFullNetwork = TFTFullNetwork()
def compile(self, optimizer, loss):
super(TFTcustommodel, self).compile()
if optimizer == 'adam':
self.optimizer = tf.keras.optimizers.Adam(learning_rate=myTFTTools.learning_rate)
else:
self.optimizer = tf.keras.optimizers.get(optimizer)
Dictopt = self.optimizer.get_config()
print(startbold+startred + 'Optimizer ' + resetfonts, Dictopt)
if loss == 'MSE' or loss =='mse':
self.loss_object = tf.keras.losses.MeanSquaredError()
elif loss == 'MAE' or loss =='mae':
self.loss_object = tf.keras.losses.MeanAbsoluteError()
else:
self.loss_object = loss
self.loss_tracker = tf.keras.metrics.Mean(name="loss")
self.loss_tracker.reset_states()
self.val_tracker = tf.keras.metrics.Mean(name="val")
self.val_tracker.reset_states()
return
def resetmetrics(self):
self.loss_tracker.reset_states()
self.val_tracker.reset_states()
return
def build_graph(self, shapes):
input = tf.keras.layers.Input(shape=shapes, name="Input")
return tf.keras.models.Model(inputs=[input], outputs=[self.call(input)])
@tf.function
def train_step(self, data):
if len(data) == 5:
X_train, y_train, sw_train, time_train, id_train = data
else:
X_train, y_train = data
sw_train = []
time_train = []
id_train = []
with tf.GradientTape() as tape:
predictions = self(X_train, time_train, id_train, training=True)
# loss = self.loss_object(y_train, predictions, sw_train)
loss = self.loss_object(y_train, predictions)
gradients = tape.gradient(loss, self.trainable_variables)
self.optimizer.apply_gradients(zip(gradients, self.trainable_variables))
self.loss_tracker.update_state(loss)
return {"loss": self.loss_tracker.result()}
@tf.function
def test_step(self, data):
if len(data) == 5:
X_val, y_val, sw_val, time_val, id_val = data
else:
X_val, y_val = data
sw_val = []
time_train = []
id_train = []
predictions = self(X_val, time_val, id_val, training=False)
# loss = self.loss_object(y_val, predictions, sw_val)
loss = self.loss_object(y_val, predictions)
self.val_tracker.update_state(loss)
return {"val_loss": self.val_tracker.result()}
#@tf.function
def call(self, inputs, time, identifier, training=None):
predictions = self.myTFTFullNetwork(inputs, time, identifier, training=training)
return predictions
# + [markdown] id="Jfb6ttCt8EHI" pycharm={"name": "#%% md\n"}
# ### TFT Overall Batch Training
#
# * TIME not set explicitly
# * Weights allowed or not
# * Assumes TFTFullNetwork is full Network
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="VqBnNKee8RYN" outputId="93fa03c2-3c94-4d4c-9700-71e1b8e90b5d" pycharm={"name": "#%%\n"}
def RunTFTCustomVersion():
myTFTTools.PrintTitle("Start Tensorflow")
TIME_start("RunTFTCustomVersion init")
global AnyOldValidation
UseClassweights = False
usecustomfit = True
AnyOldValidation = myTFTTools.validation
garbagecollectcall = 0
# XXX InitializeDLforTimeSeries setSeparateDLinput NOT USED
tf.keras.backend.set_floatx('float32')
# tf.compat.v1.disable_eager_execution()
myTFTcustommodel = TFTcustommodel(name ='myTFTcustommodel')
lossobject = 'MSE'
if myTFTTools.lossflag == 8:
lossobject = custom_lossGCF1
if myTFTTools.lossflag == 11:
lossobject = 'MAE'
if myTFTTools.lossflag == 12:
lossobject = tf.keras.losses.Huber(delta=myTFTTools.HuberLosscut)
myTFTcustommodel.compile(loss= lossobject, optimizer= myTFTTools.optimizer)
recordtrainloss = []
recordvalloss = []
tfrecordtrainloss = tf.Variable([], shape =tf.TensorShape(None), trainable = False)
tfrecordvalloss = tf.Variable([], shape =tf.TensorShape(None), trainable = False)
tfepochstep = tf.Variable(0, trainable = False)
TIME_stop("RunTFTCustomVersion init")
# Set up checkpoints to read or write
mycheckpoint = tf.train.Checkpoint(optimizer=myTFTcustommodel.optimizer,
model=myTFTcustommodel, tfepochstep=tf.Variable(0),
tfrecordtrainloss=tfrecordtrainloss,tfrecordvalloss=tfrecordvalloss)
TIME_start("RunTFTCustomVersion restore")
# This restores back up
if Restorefromcheckpoint:
save_path = inputCHECKPOINTDIR + inputRunName + inputCheckpointpostfix
mycheckpoint.restore(save_path=save_path).expect_partial()
tfepochstep = mycheckpoint.tfepochstep
recordvalloss = mycheckpoint.tfrecordvalloss.numpy().tolist()
recordtrainloss = mycheckpoint.tfrecordtrainloss.numpy().tolist()
trainlen = len(recordtrainloss)
extrainfo = ''
vallen = len(recordvalloss)
SavedTrainLoss = recordtrainloss[trainlen-1]
SavedValLoss = 0.0
if vallen > 0:
extrainfo = ' Val Loss ' + str(round(recordvalloss[vallen-1],7))
SavedValLoss = recordvalloss[vallen-1]
print(startbold + 'Network restored from ' + save_path + '\nLoss ' + str(round(recordtrainloss[trainlen-1],7))
+ extrainfo + ' Epochs ' + str(tfepochstep.numpy()) + resetfonts )
TFTTrainingMonitor.SetCheckpointParms(mycheckpoint,CHECKPOINTDIR,RunName = RunName,Restoredcheckpoint= True,
Restored_path = save_path, ValidationFraction = AnyOldValidation, SavedTrainLoss = SavedTrainLoss,
SavedValLoss =SavedValLoss)
else:
TFTTrainingMonitor.SetCheckpointParms(mycheckpoint,CHECKPOINTDIR,RunName = RunName,Restoredcheckpoint= False,
ValidationFraction = AnyOldValidation)
TIME_stop("RunTFTCustomVersion restore")
TIME_start("RunTFTCustomVersion analysis")
# This just does analysis
if AnalysisOnly:
if OutputNetworkPictures:
outputpicture1 = APPLDIR +'/Outputs/Model_' +RunName + '1.png'
outputpicture2 = APPLDIR +'/Outputs/Model_' +RunName + '2.png'
# TODO: also save as pdf if possible
tf.keras.utils.plot_model(myTFTcustommodel.build_graph([Tseq,NpropperseqTOT]),
show_shapes=True,
to_file = outputpicture1,
show_dtype=True,
expand_nested=True)
tf.keras.utils.plot_model(myTFTcustommodel.myTFTFullNetwork.build_graph([Tseq,NpropperseqTOT]),
show_shapes=True,
to_file = outputpicture2,
show_dtype=True,
expand_nested=True)
if myTFTTools.TFTSymbolicWindows:
finalizeTFTDL(myTFTcustommodel,recordtrainloss,recordvalloss,AnyOldValidation,TFTtest_datacollection,2, LabelFit = 'Custom TFT Fit')
else:
finalizeTFTDL(myTFTcustommodel,recordtrainloss,recordvalloss,AnyOldValidation,TFTtest_datacollection,2, LabelFit = 'Custom TFT Fit')
return
TIME_stop("RunTFTCustomVersion analysis")
TIME_start("RunTFTCustomVersion train")
# Initialize progress bars
epochsize = len(TFTtrain_datacollection["inputs"])
if AnyOldValidation > 0.001:
epochsize += len(TFTval_datacollection["inputs"])
pbar = notebook.trange(myTFTTools.num_epochs, desc='Training loop', unit ='epoch')
bbar = notebook.trange(epochsize, desc='Batch loop', unit = 'sample')
train_epoch = 0.0 # Training Loss this epoch
val_epoch = 0.0 # Validation Loss this epoch
Ctime1 = 0.0
Ctime2 = 0.0
Ctime3 = 0.0
GarbageCollect = True
# train_dataset = tf.data.Dataset.from_tensor_slices((TFTtrain_datacollection['inputs'],TFTtrain_datacollection['outputs'],TFTtrain_datacollection['active_entries']))
# val_dataset = tf.data.Dataset.from_tensor_slices((TFTval_datacollection['inputs'],TFTval_datacollection['outputs'],TFTval_datacollection['active_entries']))
OuterTrainBatchDimension = TFTtrain_datacollection['inputs'].shape[0]
OuterValBatchDimension = TFTval_datacollection['inputs'].shape[0]
print('Samples to batch Train ' + str(OuterTrainBatchDimension) + ' Val ' + str(OuterValBatchDimension))
# train_dataset = train_dataset.shuffle(buffer_size = OuterBatchDimension, reshuffle_each_iteration=True).batch(myTFTTools.minibatch_size)
# val_dataset = val_dataset.batch(myTFTTools.maxibatch_size)
np.random.seed(int.from_bytes(os.urandom(4), byteorder='little'))
trainbatchsize = myTFTTools.minibatch_size
valbatchsize = myTFTTools.maxibatch_size
numberoftrainbatches = math.ceil(OuterTrainBatchDimension/trainbatchsize)
numberofvalbatches = math.ceil(OuterValBatchDimension/valbatchsize)
for e in pbar:
myTFTcustommodel.resetmetrics()
train_lossoverbatch=[]
val_lossoverbatch=[]
if batchperepoch:
qbar = notebook.trange(epochsize, desc='Batch loop epoch ' +str(e))
# for batch, (X_train, y_train, sw_train) in enumerate(train_dataset.take(-1))
trainingorder = np.arange(0, OuterTrainBatchDimension)
np.random.shuffle(trainingorder)
count1 = 0
for countbatches in range(0,numberoftrainbatches):
count2 = min(OuterTrainBatchDimension, count1+trainbatchsize)
if count2 <= count1:
continue
samples = trainingorder[count1:count2]
count1 += trainbatchsize
X_train = TFTtrain_datacollection['inputs'][samples,Ellipsis]
y_train = TFTtrain_datacollection['outputs'][samples,Ellipsis]
sw_train = []
time_train = []
id_train = []
Numinbatch = X_train.shape[0]
# myTFTTools.TFTSymbolicWindows X_train is indexed by Batch index, 1(replace by Window), 1 (replace by properties)
if myTFTTools.TFTSymbolicWindows:
StopWatch.start('label1')
X_train = X_train.numpy()
X_train = np.reshape(X_train,Numinbatch)
iseqarray = np.right_shift(X_train,16)
ilocarray = np.bitwise_and(X_train, 0b1111111111111111)
StopWatch.stop('label1')
Ctime1 += StopWatch.get('label1', digits=4)
StopWatch.start('label3')
X_train_withSeq = list()
for iloc in range(0,Numinbatch):
X_train_withSeq.append(ReshapedSequencesTOT[ilocarray[iloc],iseqarray[iloc]:iseqarray[iloc]+Tseq])
# X_train_withSeq=[ReshapedSequencesTOT[ilocarray[iloc],iseqarray[iloc]:iseqarray[iloc]+Tseq] for iloc in range(0,Numinbatch)]
StopWatch.stop('label3')
Ctime3 += StopWatch.get('label3', digits=5)
StopWatch.start('label2')
loss = myTFTcustommodel.train_step((np.array(X_train_withSeq), y_train, sw_train, time_train,id_train))
StopWatch.stop('label2')
Ctime2 += StopWatch.get('label2', digits=4)
else:
loss = myTFTcustommodel.train_step((X_train, y_train, sw_train, time_train, id_train))
GarbageCollect = False
if GarbageCollect:
if myTFTTools.TFTSymbolicWindows:
X_train_withSeq = None
X_train = None
y_train = None
sw_train = None
time_train = None
id_train = None
if garbagecollectcall > GarbageCollectionLimit:
garbagecollectcall = 0
gc.collect()
garbagecollectcall += 1
localloss = loss["loss"].numpy()
train_lossoverbatch.append(localloss)
if batchperepoch:
qbar.update(LSTMbatch_size)
qbar.set_postfix(Loss = localloss, Epoch = e)
bbar.update(Numinbatch)
bbar.set_postfix(Loss = localloss, Epoch = e)
# End Training step for one batch
# Start Validation
if AnyOldValidation:
count1 = 0
for countbatches in range(0,numberofvalbatches):
count2 = min(OuterValBatchDimension, count1+valbatchsize)
if count2 <= count1:
continue
samples = np.arange(count1,count2)
count1 += valbatchsize
X_val = TFTval_datacollection['inputs'][samples,Ellipsis]
y_val = TFTval_datacollection['outputs'][samples,Ellipsis]
sw_val = []
# for batch, (X_val, y_val, sw_val) in enumerate(val_dataset.take(-1)):
time_val = []
id_val =[]
Numinbatch = X_val.shape[0]
# myTFTTools.TFTSymbolicWindows X_val is indexed by Batch index, 1(replace by Window), 1 (replace by properties)
if myTFTTools.TFTSymbolicWindows:
StopWatch.start('label1')
X_val = X_val.numpy()
X_val = np.reshape(X_val,Numinbatch)
iseqarray = np.right_shift(X_val,16)
ilocarray = np.bitwise_and(X_val, 0b1111111111111111)
StopWatch.stop('label1')
Ctime1 += StopWatch.get('label1', digits=4)
StopWatch.start('label3')
X_valFull = list()
for iloc in range(0,Numinbatch):
X_valFull.append(ReshapedSequencesTOT[ilocarray[iloc],iseqarray[iloc]:iseqarray[iloc]+Tseq])
StopWatch.stop('label3')
Ctime3 += StopWatch.get('label3', digits=5)
StopWatch.start('label2')
loss = myTFTcustommodel.test_step((np.array(X_valFull), y_val, sw_val, time_val, id_val))
StopWatch.stop('label2')
Ctime2 += StopWatch.get('label2', digits=4)
else:
loss = myTFTcustommodel.test_step((X_val, y_val, sw_val, time_val, id_val))
localval = loss["val_loss"].numpy()
val_lossoverbatch.append(localval)
bbar.update(Numinbatch)
bbar.set_postfix(Val_loss = localval, Epoch = e)
# End Batch
train_epoch = train_lossoverbatch[-1]
recordtrainloss.append(train_epoch)
mycheckpoint.tfrecordtrainloss = tf.Variable(recordtrainloss)
'''
line = 'Train ' + str(round(np.mean(train_lossoverbatch),5)) + ' '
count = 0
for x in train_lossoverbatch:
if count%100 == 0:
line = line + str(count) +':' + str(round(x,5)) + ' '
count += 1
print(wraptotext(line,size=180))
'''
val_epoch = 0.0
if AnyOldValidation > 0.001:
val_epoch = val_lossoverbatch[-1]
recordvalloss.append(val_epoch)
mycheckpoint.tfrecordvalloss = tf.Variable(recordvalloss)
'''
line = 'Val ' + str(round(np.mean(val_lossoverbatch),5)) + ' '
count = 0
for x in val_lossoverbatch:
if count%100 == 0:
line = line + str(count) +':' + str(round(x,5)) + ' '
count += 1
print(wraptotext(line,size=180))
'''
pbar.set_postfix(Loss = train_epoch, Val = val_epoch)
bbar.reset()
tfepochstep = tfepochstep + 1
mycheckpoint.tfepochstep.assign(tfepochstep)
# Decide on best fit
MonitorResult, train_epoch, val_epoch = TFTTrainingMonitor.EpochEvaluate(e,train_epoch, val_epoch,
tfepochstep, recordtrainloss, recordvalloss)
if MonitorResult==1:
tfepochstep, recordtrainloss, recordvalloss, train_epoch, val_epoch = TFTTrainingMonitor.RestoreBestFit() # Restore Best Fit
else:
continue
# *********************** End of Epoch Loop
TIME_stop("RunTFTCustomVersion train")
# Print Fit details
print(startbold + 'Times ' + str(round(Ctime1,5)) + ' ' + str(round(Ctime3,5)) + ' TF ' + str(round(Ctime2,5)) + resetfonts)
TFTTrainingMonitor.PrintEndofFit(TFTTransformerepochs)
# Set Best Possible Fit
TIME_start("RunTFTCustomVersion bestfit")
TIME_start("RunTFTCustomVersion bestfit FTTrainingMonitor")
tfepochstep, recordtrainloss, recordvalloss, train_epoch, val_epoch = TFTTrainingMonitor.BestPossibleFit()
TIME_stop("RunTFTCustomVersion bestfit FTTrainingMonitor")
if Checkpointfinalstate:
TIME_start("RunTFTCustomVersion bestfit Checkpointfinalstate")
savepath = mycheckpoint.save(file_prefix=CHECKPOINTDIR + RunName)
print('Checkpoint at ' + savepath + ' from ' + CHECKPOINTDIR)
TIME_stop("RunTFTCustomVersion bestfit Checkpointfinalstate")
trainlen = len(recordtrainloss)
extrainfo = ''
if AnyOldValidation > 0.001:
vallen = len(recordvalloss)
extrainfo = ' Val Epoch ' + str(vallen-1) + ' Val Loss ' + str(round(recordvalloss[vallen-1],7))
print('Train Epoch ' + str(trainlen-1) + ' Train Loss ' + str(round(recordtrainloss[trainlen-1],7)) + extrainfo)
#
TIME_start("RunTFTCustomVersion bestfit summary")
myTFTcustommodel.summary()
TIME_stop("RunTFTCustomVersion bestfit summary")
TIME_start("RunTFTCustomVersion bestfit network summary")
print('\nmyTFTcustommodel.myTFTFullNetwork **************************************')
myTFTcustommodel.myTFTFullNetwork.summary()
TIME_stop("RunTFTCustomVersion bestfit network summary")
print('\nmyTFTcustommodel.myTFTFullNetwork.TFTLSTMEncoder **************************************')
if not myTFTTools.TFTdefaultLSTM:
TIME_start("RunTFTCustomVersion bestfit TFTLSTMEncoder summary")
myTFTcustommodel.myTFTFullNetwork.TFTLSTMEncoder.summary()
TIME_stop("RunTFTCustomVersion bestfit TFTLSTMEncoder summary")
print('\nmyTFTcustommodel.myTFTFullNetwork.TFTLSTMDecoder **************************************')
TIME_start("RunTFTCustomVersion bestfit TFTLSTMDecoder summary")
myTFTcustommodel.myTFTFullNetwork.TFTLSTMEncoder.summary()
# TODO: Gregor thinks it shoudl be: myTFTcustommodel.myTFTFullNetwork.TFTLSTMDecoder.summary()
TIME_stop("RunTFTCustomVersion bestfit TFTLSTMDecoder summary")
print('\nmyTFTcustommodel.myTFTFullNetwork.TFTself_attn_layer **************************************')
TIME_start("RunTFTCustomVersion bestfit Network attn layer summary")
myTFTcustommodel.myTFTFullNetwork.TFTself_attn_layer.summary()
TIME_stop("RunTFTCustomVersion bestfit Network attn layer summary")
TIME_start("RunTFTCustomVersion bestfit Network attn layer attention summary")
myTFTcustommodel.myTFTFullNetwork.TFTself_attn_layer.attention.summary()
TIME_stop("RunTFTCustomVersion bestfit Network attn layer attention summary")
if OutputNetworkPictures:
outputpicture1 = APPLDIR +'/Outputs/Model_' +RunName + '1.png'
outputpicture2 = APPLDIR +'/Outputs/Model_' +RunName + '2.png'
# ALso save as PDF if possible
TIME_start("RunTFTCustomVersion bestfit Model build graph")
tf.keras.utils.plot_model(myTFTcustommodel.build_graph([Tseq,NpropperseqTOT]),
show_shapes=True, to_file = outputpicture1,
show_dtype=True,
expand_nested=True)
TIME_stop("RunTFTCustomVersion bestfit Model build graph")
TIME_start("RunTFTCustomVersion bestfit Network build graph")
tf.keras.utils.plot_model(myTFTcustommodel.myTFTFullNetwork.build_graph([Tseq,NpropperseqTOT]),
show_shapes=True, to_file = outputpicture2,
show_dtype=True,
expand_nested=True)
TIME_stop("RunTFTCustomVersion bestfit Network build graph")
TIME_start("RunTFTCustomVersion bestfit finalize")
if myTFTTools.TFTSymbolicWindows:
finalizeTFTDL(myTFTcustommodel,recordtrainloss,recordvalloss,AnyOldValidation,TFTtest_datacollection,2, LabelFit = 'Custom TFT Fit')
else:
finalizeTFTDL(myTFTcustommodel,recordtrainloss,recordvalloss,AnyOldValidation,TFTtest_datacollection,2, LabelFit = 'Custom TFT Fit')
TIME_stop("RunTFTCustomVersion bestfit finalize")
TIME_stop("RunTFTCustomVersion bestfit")
return
# + [markdown] pycharm={"name": "#%% md\n"}
#
# + [markdown] id="YdakV_4cz3Ck" pycharm={"name": "#%% md\n"}
#
# ###Run TFT
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="dD_cDFla_yV2" outputId="887d249d-5386-4069-e99b-9a59d9c8d45d" pycharm={"name": "#%%\n"}
# Run TFT Only
TIME_start("RunTFTCustomVersion tft only")
AnalysisOnly = myTFTTools.AnalysisOnly
Dumpoutkeyplotsaspics = True
Restorefromcheckpoint = myTFTTools.Restorefromcheckpoint
Checkpointfinalstate = True
if AnalysisOnly:
Restorefromcheckpoint = True
Checkpointfinalstate = False
if Restorefromcheckpoint:
inputCHECKPOINTDIR = CHECKPOINTDIR
inputRunName = myTFTTools.inputRunName
inputCheckpointpostfix = myTFTTools.inputCheckpointpostfix
inputCHECKPOINTDIR = APPLDIR + "/checkpoints/" + inputRunName + "dir/"
batchperepoch = False # if True output a batch bar for each epoch
GlobalSpacetime = False
IncreaseNloc_sample = 1
DecreaseNloc_sample = 1
SkipDL2F = True
FullSetValidation = False
TFTTrainingMonitor = TensorFlowTrainingMonitor()
TFTTrainingMonitor.SetControlParms(SuccessLimit = 1,FailureLimit = 2)
TIME_stop("RunTFTCustomVersion tft only")
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="DZIby3dM_yV5" outputId="8b81e9c1-c714-4733-d7ab-4fb26597ee38" pycharm={"name": "#%%\n"}
def PrintLSTMandBasicStuff(model):
myTFTTools.PrintTitle('Start TFT Deep Learning')
if myTFTTools.TFTSymbolicWindows:
print(startbold + startred + 'Symbolic Windows used to save space'+resetfonts)
else:
print(startbold + startred + 'Symbolic Windows NOT used'+resetfonts)
print('Training Locations ' + str(TrainingNloc) + ' Validation Locations ' + str(ValidationNloc) +
' Sequences ' + str(Num_Seq))
if LocationBasedValidation:
print(startbold + startred + " Location Based Validation with fraction " + str(LocationValidationFraction)+resetfonts)
if RestartLocationBasedValidation:
print(startbold + startred + " Using Validation set saved in " + RestartRunName+resetfonts)
print('\nAre futures predicted ' + str(UseFutures) + ' Custom Loss Pointer ' + str(CustomLoss) + ' Class weights used ' + str(UseClassweights))
print('\nProperties per sequence ' + str(NpropperseqTOT))
print('\n' + startbold +startpurple + 'Properties ' + resetfonts)
labelline = 'Name '
for propval in range (0,7):
labelline += QuantityStatisticsNames[propval] + ' '
print('\n' + startbold + labelline + resetfonts)
for iprop in range(0,NpropperseqTOT):
line = startbold + startpurple + str(iprop) + ' ' + InputPropertyNames[PropertyNameIndex[iprop]] + resetfonts
jprop = PropertyAverageValuesPointer[iprop]
line += ' Root ' + str(QuantityTakeroot[jprop])
for proppredval in range (0,7):
line += ' ' + str(round(QuantityStatistics[jprop,proppredval],3))
print(line)
print('\nPredictions per sequence ' + str(NpredperseqTOT))
print('\n' + startbold +startpurple + 'Predictions ' + resetfonts)
print('\n' + startbold + labelline + resetfonts)
for ipred in range(0,NpredperseqTOT):
line = startbold + startpurple + str(ipred) + ' ' + Predictionname[ipred] + ' wgt ' + str(round(Predictionwgt[ipred],3)) + resetfonts + ' '
jpred = PredictionAverageValuesPointer[ipred]
line += ' Root ' + str(QuantityTakeroot[jpred])
for proppredval in range (0,7):
line += ' ' + str(round(QuantityStatistics[jpred,proppredval],3))
print(line)
print('\n')
myTFTTools.PrintTitle('Start TFT Deep Learning')
for k in TFTparams:
print('# {} = {}'.format(k, TFTparams[k]))
# + colab={"base_uri": "https://localhost:8080/", "height": 20000, "referenced_widgets": ["0634f0a96f5d406cae0d78a1b8f24346", "f28ff11c09ab4843bfaacf725b997f95", "d03038f77d084d1b9b0e4f42b2ebb622", "9b04396d9eb2420e9259b8573eef3b70", "<KEY>", "<KEY>", "0a1d9b55698544a581539f34595b7755", "ce660d6f798d472199837af0be65be0a", "4c5a70fcb0e64de5892f41aabba46b81", "bcee1a2685974293af383558e636e347", "decf3f3a60c3460694dab0bd74a621a5"]} id="GeEXX-lSuuhq" outputId="570bbe2e-0bd0-49ab-a7e8-373f01c10a4d" pycharm={"name": "#%%\n"}
TIME_start("RunTFTCustomVersion print")
runtype = ''
if Restorefromcheckpoint:
runtype = 'Restarted '
myTFTTools.PrintTitle(runtype)
PrintLSTMandBasicStuff(2)
TIME_stop("RunTFTCustomVersion print")
TIME_start("RunTFTCustomVersion A")
RunTFTCustomVersion()
myTFTTools.PrintTitle('TFT run completed')
TIME_stop("RunTFTCustomVersion A")
# + pycharm={"name": "#%%\n"}
StopWatch.stop("total")
StopWatch.benchmark()
# + pycharm={"name": "#%%\n"}
StopWatch.benchmark(sysinfo=False, attributes="short")
if in_rivanna:
print("Partition is " + str(os.getenv('SLURM_JOB_PARTITION')))
print("Job ID is " + str(os.getenv("SLURM_JOB_ID")))
# + pycharm={"name": "#%%\n"}
sys.exit(0)
# + [markdown] id="w7sWeoGSNREO" pycharm={"name": "#%% md\n"}
# #End modified TFT
| benchmarks/earthquake/mar2022/FFFFWNPFEARTHQ_newTFTv29-gregor-parameters-pre-may2022.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Naive Bayes
# ## Bayes Theorem
#
# Bayes theorem is one of the earliest inference algorithm. Bayes Theorem calculates the probability of a certain event happening (e.g. a message being spam) based on the joint probabilistic distributions of certain other events (e.g. the appearance of certain words in a message).
#
# \begin{equation*}
# P(A|B) = \frac{P(A)P(B|A)}{P(B)}
# \end{equation*}
#
# Example: Brenda and Alex are employees at the office. We know that:
# * Alex works mostly in the office, 3 times a week
# * Brenda mainly travels, so she comes to the office once a week
# NOTE: a week is made of 5 working days
#
# We observe someone running through the office wearing a red sweather, but we're unable to recognise who she/he is. So, we make a guess. Based on the above data, we calculate the single probabilities of the single events:
#
# \begin{equation*}
# P(A) = \frac{3}{5}=0.60 \;\;\; P(B) = \frac{1}{5}=0.20
# \end{equation*}
#
# We have now to normalize these probability, since we do now that someone was wearing a red sweather. Therefore:
#
# \begin{equation*}
# P(A) = \frac{P(A)}{P(A)+P(B)} \;\;\; P(B) = \frac{P(B)}{P(A)+P(B)}\\\\
# P(A) = \frac{\frac{3}{5}}{\frac{3}{5}+\frac{1}{5}}=0.75 \;\;\; P(B) = \frac{\frac{1}{5}}{\frac{3}{5}+\frac{1}{5}}=0.25
# \end{equation*}
#
# Now, we introduce more knowledge:
# * Alex wears a red sweather 2 days a week
# * Brenda wears a sweather 3 days a week
#
#
# <img src=".\images\4.01_bayes-example.png" style="width: 600px;"/>
#
# Therefore, the probability of having seen Alex wearing a red sweather is:
#
# \begin{equation*}
# P(A|R) = \frac{P(A)P(R|A)}{P(R)} \\
# P(A|R) = \frac{P(A)P(R|A)}{P(A)P(R|A)+P( \neg A)P(R|\neg A)}\\
# P(A|R) = \frac{0.75 \cdot 0.40}{0.75 \cdot 0.40 + 0.25 \cdot 0.60} = 66.7\%
# \end{equation*}
# ## Naive Bayes
#
# Spam email classifier can be built as a Naive Bayes Classifier. We check the words of an email against a sample of sentences we have.
#
# <img src=".\images\4.02_spam-example_01.png" style="width: 400px;"/>
#
# So, given an email that contains the word `easy`, the probability that is spam is $1/3$, while given an email that contains the word `money`, the proabbility that is spam is $2/3$ (total sentences in spam cat = 3).
#
# #### Example
#
# Suppose you have a bag with three standard 6-sided dice with face values [1,2,3,4,5,6] and two non-standard 6-sided dice with face values [2,3,3,4,4,5]. Someone draws a
# die from the bag, rolls it, and announces it was a 3. What is the probability that the die that was rolled was a standard die?
#
# \begin{equation*}
# P(std) = 3/5 \\
# P(\neg std) = 2/5 \\
# \\
# P(std|'3') = \frac{P(std) \cdot P('3'|std)}{P(std) \cdot P('3'|std) + P(\neg std) \cdot P('3'|\neg std)} = \frac{3/5 \cdot 1/6}{3/5 \cdot 1/6+2/5 \cdot 1/3} = \frac{1/10}{1/10+2/15} = \frac{3}{7} = 43\%
# \end{equation*}
# +
# Import our libraries
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
# Read in our dataset
df = pd.read_csv('data\smsspamcollection/SMSSpamCollection',
sep='\t',
header=None,
names=['label', 'sms_message'])
# Fix our response value
df['label'] = df.label.map({'ham':0, 'spam':1})
# Split our dataset into training and testing data
X_train, X_test, y_train, y_test = train_test_split(df['sms_message'],
df['label'],
random_state=1)
# Instantiate the CountVectorizer method
count_vector = CountVectorizer()
# Fit the training data and then return the matrix
training_data = count_vector.fit_transform(X_train)
# Transform testing data and return the matrix. Note we are not fitting the testing data into the CountVectorizer()
testing_data = count_vector.transform(X_test)
# Instantiate our model
naive_bayes = MultinomialNB()
# Fit our model to the training data
naive_bayes.fit(training_data, y_train)
# Predict on the test data
predictions = naive_bayes.predict(testing_data)
# Score our model
print('Accuracy score: ', format(accuracy_score(y_test, predictions)))
print('Precision score: ', format(precision_score(y_test, predictions)))
print('Recall score: ', format(recall_score(y_test, predictions)))
print('F1 score: ', format(f1_score(y_test, predictions)))
| course/1_supervised_learning/04 Naive Bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# # VCFTools.jl
#
# VCFTools.jl provides some Julia utilities for handling the VCF files.
# dispay Julia version info
versioninfo()
# ## Example VCF file
#
# Current folder contains an example VCF file for demonstation.
;ls -l test.vcf.gz
# Load the VCF file and display the first 35 lines
# +
using VCFTools
fh = openvcf("test.vcf.gz", "r")
for l in 1:35
println(readline(fh))
end
close(fh)
# -
# As in typical VCF files, it has a bunch of meta-information lines, one header line, and then one line for each each marker. In this VCF, genetic data has fields GT (genotype), DS (dosage), and GL (genotype likelihood).
# ## Summary statistics
#
# * Number of records (markers) in a VCF file.
records = nrecords("test.vcf.gz")
# * Number of samples (individuals) in a VCF file.
samples = nsamples("test.vcf.gz")
# * `gtstats` function calculates genotype statistics for each marker with GT field.
@time records, samples, lines, missings_by_sample, missings_by_record,
maf_by_record, minorallele_by_record = gtstats("test.vcf.gz");
# number of markers
records
# number of samples (individuals)
samples
# number of markers with GT field
lines
# number of missing genotypes in each sample (individual)
missings_by_sample'
# number of missing genotypes in each marker with GT field
missings_by_record'
# minor allele frequency of each marker with GT field
maf_by_record'
# minor allele of each marker (with GT field): true (REF) or false (ALT)
minorallele_by_record'
# The optional second argument of `gtstats` function specifies an output file or IO stream for genotype statistics per marker. Each line has fields:
# - 1-8: VCF fixed fields (CHROM, POS, ID, REF, ALT, QUAL, FILT, INFO)
# - 9: Missing genotype count
# - 10: Missing genotype frequency
# - 11: ALT allele count
# - 12: ALT allele frequency
# - 13: Minor allele count (REF allele vs ALT alleles)
# - 14: Minor allele frequency (REF allele vs ALT alleles)
# - 15: HWE P-value (REF allele vs ALT alleles)
# write genotype statistics in file gtstats.out.txt
@time gtstats("test.vcf.gz", "gtstats.out.txt");
# The output file can be read as a `DataFrame` for further analysis.
# +
using CSV
gstat = CSV.read("gtstats.out.txt";
header = [:chr, :pos, :id, :ref, :alt, :qual, :filt, :info, :missings, :missfreq, :nalt, :altfreq, :nminor, :maf, :hwe],
delim = '\t',
)
# -
# ## Filter
#
# Sometimes we wish to subset entire VCF files, such as filtering out certain samples or records (SNPs). This is achieved via the filter function:
# filtering by specifying indices to keep
record_mask = 1:records # keep all records (SNPs)
sample_mask = 2:(samples - 1) # keep all but first and last sample (individual)
@time VCFTools.filter("test.vcf.gz", record_mask, sample_mask,
des="filtered.test.vcf.gz")
# One can also supply bitvectors as masks:
record_mask = trues(records)
sample_mask = trues(samples)
record_mask[1] = record_mask[end] = false
@time VCFTools.filter("test.vcf.gz", record_mask, sample_mask,
des="filtered.test.vcf.gz")
# ## Convert
#
# Convert GT data in VCF file `test.vcf.gz` to a `Matrix{Union{Missing, Int8}}`. Here `as_minorallele = false` indicates that `VCFTools.jl` will copy the `0`s and `1`s of the file directly into `A`, without checking if ALT or REF is the minor allele.
@time A = convert_gt(Int8, "test.vcf.gz"; as_minorallele = false,
model = :additive, impute = false, center = false, scale = false)
# Convert GT data in VCF file `test.vcf.gz` to a numeric array. This checks which of `ALT/REF` is the minor allele, imputes the missing genotypes according to allele frequency, centers the dosages around 2MAF, and scales the dosages by `sqrt(2MAF*(1-MAF))`.
@time A = convert_gt(Float64, "test.vcf.gz"; as_minorallele = true,
model = :additive, impute = true, center = true, scale = true)
# ## Extract data marker-by-maker or window-by-window
#
# Large VCF files easily generate numeric arrays that cannot fit into computer memory. Many analyses only need to loop over markers or sets of markers. Previous functions for importing genotypes/haplotypes/dosages have equivalent functions to achieve this:
#
# + `copy_gt!` loops over genotypes
# + `copy_ht!` loops over haplotypes
# + `copy_ds!` loops over dosages
#
# For example, to loop over all genotype markers in the VCF file `test.vcf.gz`:
# +
using GeneticVariation
# initialize VCF reader
people, snps = nsamples("test.vcf.gz"), nrecords("test.vcf.gz")
reader = VCF.Reader(openvcf("test.vcf.gz"))
# pre-allocate vector for marker data
g = zeros(Union{Missing, Float64}, people)
for j = 1:snps
copy_gt!(g, reader; model = :additive, impute = true, center = true, scale = true)
# do statistical anlaysis
end
close(reader)
# -
# To loop over markers in windows of size 25:
# initialize VCF reader
people, snps = nsamples("test.vcf.gz"), nrecords("test.vcf.gz")
reader = VCF.Reader(openvcf("test.vcf.gz"))
# pre-allocate matrix for marker data
windowsize = 25
g = zeros(Union{Missing, Float64}, people, windowsize)
nwindows = ceil(Int, snps / windowsize)
for j = 1:nwindows
copy_gt!(g, reader; model = :additive,
impute = true, center = true, scale = true)
# do statistical anlaysis
end
close(reader)
| LangeSymposium-ProgrammingWorkshop-20202022-master/LangeSymposium-ProgrammingWorkshop-20202022-master/02-vcftools/vcftools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Latexify Matplotlib
# > Towards amazing plots in research papers!
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [visualisation]
# Every time I would prepare a matplotlib graph for a paper, I would iteratively adjust the figure size, the font size, scaling in LaTeX. This turns to be a tedious process. Fortunately, when along with [Jack](http://www.jack-kelly.com/) and [Oliver](http://www.oliverparson.co.uk/), I was writing our [nilmtk paper](http://arxiv.org/pdf/1404.3878v1.pdf), Jack demonstrated a function to "latexify" plots. This function would take care of font sizes and scaling, so that in one go one could generate a plot and stick in LaTeX. In this post I'll illustrate this technique to save all that iterative effort and make plots look nicer.
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import matplotlib
from math import sqrt
SPINE_COLOR = 'gray'
# The following is the `latexify` function. It allows you to create 2 column or 1 column figures. You may also wish to alter the `height` or `width` of the figure. The default settings are good for most cases.
# You may also change the parameters such as labelsize and fontsize based on your classfile. For this post, I'll use the following [ACM classfile](http://www.acm.org/sigs/publications/proceedings-templates).
# +
def latexify(fig_width=None, fig_height=None, columns=1):
"""Set up matplotlib's RC params for LaTeX plotting.
Call this before plotting a figure.
Parameters
----------
fig_width : float, optional, inches
fig_height : float, optional, inches
columns : {1, 2}
"""
# code adapted from http://www.scipy.org/Cookbook/Matplotlib/LaTeX_Examples
# Width and max height in inches for IEEE journals taken from
# computer.org/cms/Computer.org/Journal%20templates/transactions_art_guide.pdf
assert(columns in [1,2])
if fig_width is None:
fig_width = 3.39 if columns==1 else 6.9 # width in inches
if fig_height is None:
golden_mean = (sqrt(5)-1.0)/2.0 # Aesthetic ratio
fig_height = fig_width*golden_mean # height in inches
MAX_HEIGHT_INCHES = 8.0
if fig_height > MAX_HEIGHT_INCHES:
print("WARNING: fig_height too large:" + fig_height +
"so will reduce to" + MAX_HEIGHT_INCHES + "inches.")
fig_height = MAX_HEIGHT_INCHES
params = {'backend': 'ps',
'text.latex.preamble': [r'\usepackage{gensymb}'],
'axes.labelsize': 8, # fontsize for x and y labels (was 10)
'axes.titlesize': 8,
'font.size': 8, # was 10
'legend.fontsize': 8, # was 10
'xtick.labelsize': 8,
'ytick.labelsize': 8,
'text.usetex': True,
'figure.figsize': [fig_width,fig_height],
'font.family': 'serif'
}
matplotlib.rcParams.update(params)
def format_axes(ax):
for spine in ['top', 'right']:
ax.spines[spine].set_visible(False)
for spine in ['left', 'bottom']:
ax.spines[spine].set_color(SPINE_COLOR)
ax.spines[spine].set_linewidth(0.5)
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
for axis in [ax.xaxis, ax.yaxis]:
axis.set_tick_params(direction='out', color=SPINE_COLOR)
return ax
# -
# %matplotlib inline
# Let us create a dummy data frame
df = pd.DataFrame(np.random.randn(10,2))
df.columns = ['Column 1', 'Column 2']
ax = df.plot()
ax.set_xlabel("X label")
ax.set_ylabel("Y label")
ax.set_title("Title")
plt.tight_layout()
plt.savefig("image1.pdf")
# Now, let us call the `latexify` function to alter matplotlib parameters suited to our LaTeX classfile.
latexify()
ax = df.plot()
ax.set_xlabel("X label")
ax.set_ylabel("Y label")
ax.set_title("Title")
plt.tight_layout()
format_axes(ax)
plt.savefig("image2.pdf")
# Let us have a quick look at our latex source file. I have scaled down the plot generated by default matploltib settings by 50%. The next plot which is generated using `latexified` settings doesn't need any scaling.
# ! cat 1.tex
# Finally, let us look at the "png" version of our generated pdf.
# 
# Clearly, the LaTeXified version is likely to save you a lot of figure tweaking! You don't need to play with different scaling settings. Nor, do you have to play with font sizes and ofcourse not with a combination of these two which can be pretty hard.
| _notebooks/2014-06-02-latexify.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import random
import pandas as pd
import numpy as np
train_ship_segmentations_df = pd.read_csv(os.path.join("./datasets/train_val/train_ship_segmentations.csv"))
# +
msk = np.random.rand(len(train_ship_segmentations_df)) < 0.8
train = train_ship_segmentations_df[msk]
test = train_ship_segmentations_df[~msk]
print("Total", train_ship_segmentations_df.shape)
print("Train",len(train))
print("Validation",len(test))
# -
# Move train set
for index, row in train.iterrows():
image_id = row["ImageId"]
old_path = "./datasets/train_val/{}".format(image_id)
new_path = "./datasets/train/{}".format(image_id)
if os.path.isfile(old_path):
os.rename(old_path, new_path)
# Move val set
for index, row in test.iterrows():
image_id = row["ImageId"]
old_path = "./datasets/train_val/{}".format(image_id)
new_path = "./datasets/val/{}".format(image_id)
if os.path.isfile(old_path):
os.rename(old_path, new_path)
# +
# Count files
path, dirs, files = next(os.walk("./datasets/train_val"))
file_count = len(files)
print(file_count)
path, dirs, files = next(os.walk("./datasets/train"))
file_count = len(files)
print(file_count)
path, dirs, files = next(os.walk("./datasets/val"))
file_count = len(files)
print(file_count)
# +
# Save new csv files
train = train.dropna()
test = test.dropna()
train.to_csv("./datasets/train/train_ship_segmentations.csv", index=False)
test.to_csv("./datasets/val/val_ship_segmentations.csv", index=False)
# +
# Verify new vsc files
train_segs = pd.read_csv(os.path.join("./datasets/train/train_ship_segmentations.csv"))
val_segs = pd.read_csv(os.path.join("./datasets/val/val_ship_segmentations.csv"))
print("train_segs", train_segs.shape)
print("val_segs", val_segs.shape)
# -
train_segs.head()
| samples/ship/Generate Train Val Sets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Contact Codes
#
# Trying to predict the likelihood of enrollment based solely on previous contacts with the college.
# These contacts include things such as:
# * campus visits
# * meeting at a college fair
# * talking with a recruiter at their high school
# +
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
import matplotlib.pyplot as plt
import matplotlib.cm as cm
plt.style.use('fivethirtyeight')
import warnings
warnings.filterwarnings('ignore')
import sys
sys.path.insert(0, '../src/visualization/')
import visualize as vis
from tqdm import tqdm_notebook
# -
# Load Contact Codes Data Sets and the Application Data Set.
#
# The Contact Codes Data Set was altered such that all date-like attributes are recorded solely by the total days since 0 A.D. Both files are included in the analysis.
# Fields of Interest:
# * Admission_application_date
# * Admission_date
# * Enrolled?
# * Unique_student_ID
# +
codes_dates = pd.read_csv('../data/processed/Qry_Contacts_xtab.csv')
codes_days = pd.read_csv('../data/processed/Qry_Contacts_xtab_days.csv')
df = pd.read_csv(
'../data/processed/CriticalPath_Data_EM_Confidential_lessNoise.csv').drop(columns='Unnamed: 0')[[
"Unique_student_ID","Enrolled","Admission_application_date","WeightatAcpt","TotalWeight"
]]
df['Admission_application_date'] = pd.to_datetime(df['Admission_application_date'])
months_to_days = {
1:0,
2:31,
3:59,
4:90,
5:120,
6:151,
7:181,
8:212,
9:243,
10:273,
11:304,
12:334
}
df['Admission_application_date_asDays'] = df['Admission_application_date'].dt.day + df['Admission_application_date'].dt.month.map(months_to_days) + df['Admission_application_date'].dt.year*365
# -
# Take only the students that are in the `CriticalPath_Data` file.
codes_days = codes_days.where(
df.Unique_student_ID.isin(codes_days.Unique_student_ID)).dropna(subset=['Unique_student_ID'])
# Merge the files together such that whether or not a student enrolls is linkable to all of the contact codes.
# +
data = pd.merge(codes_days,df,how='left',on="Unique_student_ID")
for col in codes_days.columns.values:
if col!='Unique_student_ID':
data[col] = data[col][data[col] < data['Admission_application_date_asDays']]
# -
# ## Create a shadow matrix of the data.
shadow_matrix = ~data.isna().drop(columns=["Unique_student_ID","Admission_application_date","Enrolled","WeightatAcpt","Admission_application_date_asDays","TotalWeight"]).astype(int)+2
# Ned had previously said that the contact code weights in the WeightAtAccpt column were determined by the first letter of the contact code.
#
# Therefore, take the sum of interactions for each letter, and use these columns in place of the original contact codes.
for letter in "ABCDEFGHIJKLMNOPQRSTVUWXYZ":
filter_col = [col for col in shadow_matrix if col.startswith(letter)]
if len(filter_col)>0:
shadow_matrix[letter] = shadow_matrix[filter_col].T.sum().T
shadow_matrix = shadow_matrix.drop(columns=filter_col)
# Some contact codes do not even appear and should be discounted. Only look at the fields with greater than 1000 students with interactions.
shadow_matrix = shadow_matrix[shadow_matrix.columns[shadow_matrix[shadow_matrix>0].count()>1000]]
# +
# pd.merge(shadow_matrix,data['Unique_student_ID'],left_index=True,right_index=True).to_csv('../data/processed/contact_codes_at_application.csv')
# -
# ## PCA
# +
# scaling the data before PCA
from sklearn.preprocessing import scale
scaled = pd.DataFrame(scale(shadow_matrix),columns=shadow_matrix.columns.values)
# implementing PCA
from sklearn.decomposition import PCA
pca = PCA(n_components=6).fit(scaled)
pca_samples = pca.transform(scaled)
# -
pca_results = vis.pca_results(scaled, pca)
pca_results.cumsum()
# ## Scree Plot
#Explained variance
vis.plot_explained_variance_ratio(pca);
# # Biplot
vis.biplot(shadow_matrix, scaled, pca);
# ### Linear Regression: Target of WeightAtAcpt
# +
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import Imputer
y = data['WeightatAcpt']
X = shadow_matrix[~y.isna()]
y = y[~y.isna()]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
# -
from sklearn.linear_model import LinearRegression, LogisticRegression
reg = LinearRegression()
reg.fit(X_train, y_train)
# # Feature Importance
# +
feature_importance = pd.Series(reg.coef_)
feature_importance.index = shadow_matrix.columns.values
feature_importance.plot(kind='barh');
plt.xlabel("Feature Weight")
plt.ylabel("Feature");
# +
reg = vis.residual_error(X_train, X_test, y_train, y_test)
plt.ylim(-3000,3000);
plt.xlim(1500);
plt.xticks(rotation=70);
MAPE = 1/len(y) * abs(y-reg.predict(X.values))/y
print("The Mean Absolute Percentage Error is: %.2f percent" % (MAPE.sum()*100))
# -
# ### How good of a predictor of Enrollment is Weight???
#
# First plot the distributions against each other.
# +
f = plt.figure(figsize=(12,6))
f.add_subplot(1,2,1)
sns.stripplot(data=data,x="Enrolled",y="WeightatAcpt",size=1);
plt.ylim(800,3000)
f.add_subplot(1,2,2)
sns.boxplot(data=data,x="Enrolled",y="WeightatAcpt",showfliers=False);
plt.ylim(800,3000);
# -
from sklearn.ensemble import AdaBoostClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.datasets import make_gaussian_quantiles
# +
y = data['Enrolled'].fillna(False).astype(int)
X = shadow_matrix[~y.isna()]
y = y.values
X = np.array(data['WeightatAcpt'].fillna(0)).reshape(-1,1)
class_names = 'AB'
plot_colors = 'br'
# Create and fit an AdaBoosted decision tree
bdt = AdaBoostClassifier(DecisionTreeClassifier(max_depth=1),
algorithm="SAMME",
n_estimators=200)
bdt.fit(X, y)
# Plot the two-class decision scores
twoclass_output = bdt.decision_function(X)
plot_range = (twoclass_output.min(), twoclass_output.max())
plt.subplots(figsize=(10,6))
status = ['Unlikely to Enroll','Likely to Enroll']
for i, n, c in zip(range(2), class_names, plot_colors):
plt.hist(twoclass_output[y == i],
bins=20,
range=plot_range,
facecolor=c,
label=f'{status[i]}',
alpha=.5,
edgecolor='k',
density=True)
x1, x2, y1, y2 = plt.axis()
plt.axis((x1, x2, y1, y2 * 1.2))
plt.legend(loc='upper right')
plt.ylabel('Samples')
plt.xlabel('Score')
plt.title('Decision Scores')
plt.tight_layout()
plt.xlim(-0.25,-0.05)
plt.xticks([]);
# plt.subplots_adjust(wspace=0.5)
| models/5.0-st-contact_codes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# This makes a first contrast curve for Altair
# created 2018 Sept. 20 by E.S.
# -
# ## SECTION TO INITIALIZE
# +
# import stuff
import urllib
import numpy as np
import matplotlib.pyplot as plt
import PynPoint
from PynPoint import Pypeline
from PynPoint.IOmodules.Hdf5Reading import Hdf5ReadingModule
from PynPoint.IOmodules.FitsWriting import FitsWritingModule
from PynPoint.IOmodules.FitsReading import FitsReadingModule
from PynPoint.IOmodules.TextReading import ParangReadingModule
from PynPoint.ProcessingModules import PSFpreparationModule, \
PcaPsfSubtractionModule, \
ContrastCurveModule, \
FluxAndPosition, \
StackingAndSubsampling
from PynPoint.ProcessingModules.StackingAndSubsampling import DerotateAndStackModule
from PynPoint.ProcessingModules.FluxAndPosition import FakePlanetModule
#from PynPoint.Util import AnalysisTools
# +
# define workspaces and initialize Pypeline
stem = "/home/../../media/unasemaje/Elements/lbti_data_reduction/180507_fizeau_altair/pynpoint_testing/"
working_place = stem+"pynpoint_experimentation_altair/working_place/"
input_place = stem+"/pynpoint_experimentation_altair/input_place/"
output_place = stem+"/pynpoint_experimentation_altair/output_place/"
pipeline = Pypeline(working_place_in=working_place,
input_place_in=input_place,
output_place_in=output_place)
# now a *.ini file has been generated (this includes the PIXSCALE), if no pre-existing one was there
# +
## IF THE *INI FILE WAS NEWLY GENERATED, EDIT THE *INI FILE TO SET
## PIXSCALE = 0.0107
# -
# ## SECTION TO READ IN DATA
# +
# read in science FITS files
# (should have PARANG=0 in all headers, so as to keep the PSF in the same original orientation on the array;
# we need to wait to correct for PARANG later)
read_science = FitsReadingModule(name_in="read_science",
input_dir=None,
image_tag="science",
check=True)
pipeline.add_module(read_science)
# +
# confirm PIXSCALE change
pixscale_config = pipeline.get_attribute("config", "PIXSCALE")
print("Plate scale for /config/ is "+str(pixscale_config))
#pixscale_sci = pipeline.get_attribute("science", "PIXSCALE")
#print("Plate scale for /science/ is "+str(pixscale_sci))
# +
# read in PSF reference FITS files (i.e., unsaturated frames)
# (these do not have any PARANG in the header, but it probably doesn't matter)
read_ref_psf = FitsReadingModule(name_in="read_ref_psf",
input_dir=input_place+'ref_psf/',
image_tag="ref_psf",
check=True)
pipeline.add_module(read_ref_psf)
# -
# ## SECTION TO PROCESS DATA
# +
## PIPELINE CHANNEL 1: IMAGE RESIDUALS
# generate PCA basis from unsaturated frames and
# do PCA PSF subtraction of the saturated frames
'''
pca_pca_subt = PcaPsfSubtractionModule(pca_numbers=(5, ),
name_in="pca",
images_in_tag="science",
reference_in_tag="ref_psf",
res_mean_tag="mean_residuals",
res_median_tag="median_residuals",
res_arr_out_tag="all_resids",
res_rot_mean_clip_tag="resid_rot",
basis_out_tag="pca_components",
subtract_mean=True,
verbose=True)
pipeline.add_module(pca_pca_subt)
'''
# note:
# images_in_tag: science images
# reference_in_tag: reference images, which COULD be the science images
# +
## PIPELINE CHANNEL 2: MAKE CONTRAST CURVE
# make a contrast curve
# (N.b. this does not separately require PcaPsfSubtractionModule)
'''
cent_size: mask radius
'''
# N.b. scale the reference PSF by 3.28 to match Altair's amplitude
contrast_curve = ContrastCurveModule(name_in="contrast_curve",
image_in_tag="science",
psf_in_tag="ref_psf",
contrast_out_tag="contrast_landscape",
pca_out_tag="pca_resids",
pca_number=20,
psf_scaling=3.28,
separation=(0.05, 0.45, 0.05),
angle=(0.0, 360.0, 60.0),
magnitude=(7.5, 1.0),
cent_size=None)
pipeline.add_module(contrast_curve)
# +
# write out PCA residuals
write_pca_resids = FitsWritingModule(file_name="second_pass_pca_resids.fits",
name_in="write_pca_resids",
output_dir=output_place,
data_tag="pca_resids")
pipeline.add_module(write_pca_resids)
# -
pipeline.run()
# +
#####################################################################################################################
# +
# extract contrast curve data
# -
contrast_curve_results = pipeline.get_data("contrast_landscape")
print(contrast_curve_results)
# +
# make plots
f, (ax1, ax2) = plt.subplots(1, 2)
ax1.plot(contrast_curve_results[:,0],contrast_curve_results[:,1])
ax1.set_title('Azim. averaged contrast limit')
ax1.set_xlabel('Radius (asec)')
ax1.set_ylabel('Radius (asec)')
ax1.invert_yaxis()
ax2.set_ylabel('del_mag')
ax2.plot(contrast_curve_results[:,0],contrast_curve_results[:,3])
ax2.set_title('Threshold of FPF')
ax2.set_xlabel('Radius (asec)')
plt.tight_layout()
f.savefig('test.png')
# -
| pynpoint_testing_altair_contrast_curve_take2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Churn Project
# --------------------------------------------------------------------
#
# _note_: This notebook was adapted from a number external sources:
# * **[Churn Prediction and Prevention in Python](https://towardsdatascience.com/churn-prediction-and-prevention-in-python-2d454e5fd9a5)**
# * **[Telecom Customer Churn Prediction](https://www.kaggle.com/pavanraj159/telecom-customer-churn-prediction*)**
#
# #### **notebook how-to's**
# * Create and test a custom `data_clean` function
# * Examine data using a serverless (containerized) `describe` function
# * Train a number of machine learning algorithms
# * Tune hyperparameters
# * Create an automated ML pipeline from various library functions
# * Run and track the pipeline results and artifacts
# ## a custom data cleaning function
#
# nuclio: ignore
import nuclio
# +
import os
import json
import pandas as pd
import numpy as np
from collections import defaultdict
from cloudpickle import dumps, dump, load
from sklearn.preprocessing import (OneHotEncoder,
LabelEncoder)
from mlrun.execution import MLClientCtx
from mlrun.datastore import DataItem
def data_clean(
context:MLClientCtx,
src: DataItem,
file_ext: str = "csv",
models_dest: str = "models/encoders",
cleaned_key: str = "cleaned-data",
encoded_key: str = "encoded-data"
):
"""process a raw churn data file
Data has 3 states here: `raw`, `cleaned` and `encoded`
* `raw` kept by default, the pipeline begins with a raw data artifact
* `cleaned` kept for charts, presentations
* `encoded` is input for a cross validation and training function
steps (not necessarily in correct order, some parallel)
* column name maps
* deal with nans and other types of missings/junk
* label encode binary and ordinal category columns
* create category ranges from numerical columns
And finally,
* test
Why we don't one-hot-encode here? One hot encoding isn't a necessary
step for all algorithms. It can also generate a very large feature
matrix that doesn't need to be serialized (even if sparse).
So we leave one-hot-encoding for the training step.
What about scaling numerical columns? Same as why we don't one hot
encode here. Do we scale before train-test split? IMHO, no. Scaling
before splitting introduces a type of data leakage. In addition,
many estimators are completely immune to the monotonic transformations
implied by scaling, so why waste the cycles?
TODO:
* parallelize where possible
* more abstraction (more parameters, chain sklearn transformers)
* convert to marketplace function
:param context: the function execution context
:param src: an artifact or file path
:param file_ext: file type for artifacts
:param models_dest: label encoders and other preprocessing steps
should be saved together with other pipeline
models
:param cleaned_key: key of cleaned data table in artifact store
:param encoded_key: key of encoded data table in artifact store
"""
df = src.as_df()
# drop columns
drop_cols_list = ["customerID", "TotalCharges"]
df.drop(drop_cols_list, axis=1, inplace=True)
# header transformations
old_cols = df.columns
rename_cols_map = {
"SeniorCitizen" : "senior",
"Partner" : "partner",
"Dependents" : "deps",
"Churn" : "labels"
}
df.rename(rename_cols_map, axis=1, inplace=True)
# add drop column to logs:
for col in drop_cols_list:
rename_cols_map.update({col: "_DROPPED_"})
# log the op
tp = os.path.join(models_dest, "preproc-column_map.json")
context.log_artifact("preproc-column_map.json",
body=json.dumps(rename_cols_map),
local_path=tp)
# VALUE transformations
# clean
# truncate reply to "No"
df = df.applymap(lambda x: "No" if str(x).startswith("No ") else x)
# encode numerical type as category bins (ordinal)
bins = [0, 12, 24, 36, 48, 60, np.inf]
labels = [0, 1, 2, 3, 4, 5]
tenure = df.tenure.copy(deep=True)
df["tenure_map"] = pd.cut(df.tenure, bins, labels=False)
tenure_map = dict(zip(bins, labels))
# save this transformation
tp = os.path.join(models_dest, "preproc-numcat_map.json")
context.log_artifact("preproc-numcat_map.json",
body=bytes(json.dumps(tenure_map).encode("utf-8")),
local_path=tp)
context.log_dataset(cleaned_key, df=df, format=file_ext, index=False)
# label encoding - generate model for each column saved in dict
# some of these columns may be hot encoded in the training step
fix_cols = ["gender", "partner", "deps", "OnlineSecurity",
"OnlineBackup", "DeviceProtection", "TechSupport",
"StreamingTV", "StreamingMovies", "PhoneService",
"MultipleLines", "PaperlessBilling", "InternetService",
"Contract", "PaymentMethod", "labels"]
d = defaultdict(LabelEncoder)
df[fix_cols] = df[fix_cols].apply(lambda x: d[x.name].fit_transform(x.astype(str)))
context.log_dataset(encoded_key, df=df, format=file_ext, index=False)
model_bin = dumps(d)
context.log_model("model",
body=model_bin,
artifact_path=os.path.join(context.artifact_path,
models_dest),
model_file="model.pkl")
# would be nice to have a check here on the integrity of all done
# raw->clean->encoded->clean->raw
# +
# nuclio: end-code
# -
# ## Create a project to host our functions, jobs and artifacts
#
# Projects are used to package multiple functions, workflows, and artifacts. We usually store project code and definitions in a Git archive.
#
# The following code creates a new project in a local dir and initialize git tracking on that
# +
import mlrun
project_name = "churn-project"
project_dir = "./project"
mlrun.set_environment('http://mlrun-api:8080', artifact_path='./data', project=project_name)
churn_proj = mlrun.new_project(project_name, project_dir, init_git=True)
# -
#
# ### register an input artifact/dataset in the store
# Log the raw data file as an artifact, enabeling us to refer to it by name (`store:///raw-data`) and ensuring we keep a record of the source data used
# +
DATA_URL = "https://raw.githubusercontent.com/mlrun/demos/master/customer-churn-prediction/WA_Fn-UseC_-Telco-Customer-Churn.csv"
churn_proj.log_artifact("raw-data", target_path=DATA_URL)
# -
# <a id="test-locally"></a>
# ### Run the data generator function locally
#
# The functions above can be tested locally. Parameters, inputs, and outputs can be specified in the API or the `Task` object.<br>
# when using `run_local()` the function inputs and outputs are automatically recorded by MLRun experiment and data tracking DB.
#
# In each run we can specify the function, inputs, parameters/hyper-parameters, etc... For more details, see the [mlrun_basics notebook](mlrun_basics.ipynb).
# run the function locally
cleaner = mlrun.run_local(
name="data_clean",
handler=data_clean,
inputs={"src": "store:///raw-data"},
params={"file_ext" : "csv",
"apply_tenure_map": False})
# ## Create a Fully Automated ML Pipeline
# Are we using GPU's?
GPUS = False
# #### Convert our local code to a distributed serverless function object
# +
clean_data_fn_params = {
"name" : "clean_data",
"kind" : "job",
"image" : "mlrun/ml-models-gpu" if GPUS else "mlrun/ml-models",
"description" : "clean and encode raw data",
"categories" : ["data-prep"],
"labels" : {"author": "yasha", "framework": "xgboost"}
}
clean_func = mlrun.code_to_function(**clean_data_fn_params)
churn_proj.set_function(clean_func)
# -
# #### Add more functions to our project to be used in our pipeline (from the functions hub/marketplace)
#
# AutoML training (classifier), Model validation (test_classifier), Real-time model server, and Model REST API Tester
# +
churn_proj.set_function("hub://describe", "describe")
churn_proj.set_function("hub://xgb_trainer", "classify")
churn_proj.set_function("hub://xgb_test", "xgbtest")
churn_proj.set_function("hub://coxph_trainer", "survive")
churn_proj.set_function("hub://coxph_test", "coxtest")
churn_proj.set_function("hub://churn_server", "server")
# -
# #### Define and save a pipeline
#
# The following workflow definition will be written into a file, it describes a Kubeflow execution graph (DAG)<br>
# and how functions and data are connected to form an end to end pipeline.
#
# Check the code below to see how functions objects are initialized and used (by name) inside the workflow.<br>
# The `workflow.py` file has two parts, initialize the function objects and define pipeline dsl (connect the function inputs and outputs).
# +
# %%writefile project/workflow.py
from kfp import dsl
from mlrun import mount_v3io
funcs = {}
GPUS = False
# init functions is used to configure function resources and local settings
def init_functions(functions: dict, project=None, secrets=None):
for f in functions.values():
f.apply(mount_v3io())
functions["server"].set_env("INFERENCE_STREAM", "users/admin/artifacts/customer-churn-prediction/model_stream")
@dsl.pipeline(
name="Demo training pipeline",
description="Shows how to use mlrun."
)
def kfpipeline():
# encode the data
clean = funcs["clean-data"].as_step(
name="clean-data",
handler="data_clean",
params={"file_ext": "csv",
"models_dest": "models/encoders"},
inputs={"src": "store:///raw-data"}, # use an artifact from the feature store
outputs=["cleaned-data",
"encoded-data"])
# analyze our dataset
describe = funcs["describe"].as_step(
name="summary",
params={"label_column" : "labels"},
inputs={"table": clean.outputs["encoded-data"]})
# train with hyper-paremeters
xgb = funcs["classify"].as_step(
name="current-state",
handler="train_model",
params={"sample" : -1,
"label_column" : "labels",
"model_type" : "classifier",
# xgb class initializers (tuning candidates):
"CLASS_tree_method" : "gpu_hist" if GPUS else "hist",
"CLASS_objective" : "binary:logistic",
"CLASS_n_estimators" : 50,
"CLASS_max_depth" : 5,
"CLASS_learning_rate" : 0.15,
"CLASS_colsample_bylevel" : 0.7,
"CLASS_colsample_bytree" : 0.8,
"CLASS_gamma" : 1.0,
"CLASS_max_delta_step" : 3,
"CLASS_min_child_weight" : 1.0,
"CLASS_reg_lambda" : 10.0,
"CLASS_scale_pos_weight" : 1,
"FIT_verbose" : 0,
"CLASS_subsample" : 0.9,
"CLASS_booster" : "gbtree",
"CLASS_random_state" : 1,
# encoding:
"encode_cols" : {"InternetService": "ISP",
"Contract" : "Contract",
"PaymentMethod" : "Payment"},
# outputs
"models_dest" : "models",
"plots_dest" : "plots",
"file_ext" : "csv"
},
inputs={"dataset" : clean.outputs["encoded-data"]},
outputs=["model", "test_set"])
cox = funcs["survive"].as_step(
name="survival-curves",
params={"sample" : -1,
"event_column" : "labels",
"strata_cols" : ['InternetService', 'StreamingMovies',
'StreamingTV', 'PhoneService'],
"encode_cols" : {"Contract" : "Contract",
"PaymentMethod" : "Payment"},
# outputs
"models_dest" : "models/cox",
"plots_dest" : "plots",
"file_ext" : "csv"
},
inputs={"dataset" : clean.outputs["encoded-data"]},
outputs=["cx-model", "tenured-test-set"])
test_xgb = funcs["xgbtest"].as_step(
name="test-classifier",
params={"label_column": "labels",
"plots_dest" : "customer-churn-prediction/test/xgb"},
inputs={"models_path" : xgb.outputs["model"],
"test_set" : xgb.outputs["test_set"]})
test_cox = funcs["coxtest"].as_step(
name="test-regressor",
params={"label_column": "labels",
"plots_dest" : "customer-churn-prediction/test/cox"},
inputs={"models_path" : cox.outputs["cx-model"],
"test_set" : cox.outputs["tenured-test-set"]})
# deploy our model as a serverless function
deploy_xgb = funcs["server"].deploy_step(
models={"churn_server_v1": xgb.outputs["model"]})
deploy_xgb.after(cox)
# -
# register the workflow file as "main"
churn_proj.set_workflow("main", "workflow.py", embed=True)
# Save the project definitions to a file (project.yaml), it is recommended to commit all changes to a Git repo.
churn_proj.save()
# +
#print(churn_proj.to_yaml())
# -
# <a id="run-pipeline"></a>
# ## Run a pipeline workflow
# use the `run` method to execute a workflow, you can provide alternative arguments and specify the default target for workflow artifacts.<br>
# The workflow ID is returned and can be used to track the progress or you can use the hyperlinks
#
# > Note: The same command can be issued through CLI commands:<br>
# `mlrun project my-proj/ -r main -p "v3io:///users/admin/mlrun/kfp/{{workflow.uid}}/"`
#
# The dirty flag allow us to run a project with uncommited changes (when the notebook is in the same git dir it will always be dirty)
artifact_path = os.path.join(mlrun.mlconf.artifact_path, "pipeline/{{workflow.uid}}")
run_id = churn_proj.run(
"main",
arguments={},
artifact_path=artifact_path,
dirty=True, watch=True)
# **[back to top](#top)**
| customer-churn-prediction/churn-project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''tf'': conda)'
# language: python
# name: python3
# ---
import pandas as pd
import pprint
from collections import defaultdict
# +
df_report = pd.read_csv('experiments_results.csv', sep='|', encoding='utf-8')
pd.options.display.float_format = '{:.4f}'.format
df_report.dropna(axis=0, inplace=True)
cols_max = ['ms_jaccard2', 'ms_jaccard3', 'ms_jaccard4', 'ms_jaccard5', 'forward_bleu2', 'backward_bleu2', 'ha_bleu2', 'forward_bleu3', 'backward_bleu3', 'ha_bleu3', 'forward_bleu4', 'backward_bleu4', 'ha_bleu4', 'forward_bleu5', 'backward_bleu5', 'ha_bleu5', 'rouge-1', 'rouge-2', 'rouge-l', 'bertscore_f1_l11', 'bertscore_f1_l12']
cols_min = ['tfidf_distance', 'fbd_1-6', 'fbd_7-12', 'self_bleu2', 'self_bleu3', 'self_bleu4', 'self_bleu5']
best = defaultdict(list)
for col in cols_max:
best[df_report.iloc[df_report[col].argmax()][0]].append(col)
for col in cols_min:
best[df_report.iloc[df_report[col].argmin()][0]].append(col)
pprint.pprint(best)
| analyse_results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NLP Basics: Exploring the dataset
# ### Read in text data
# +
import pandas as pd
fullCorpus = pd.read_csv('SMSSpamCollection.tsv', sep='\t', header=None)
fullCorpus.columns = ['label', 'body_test']
print(fullCorpus)
# -
# ### Explore the dataset
# +
# What is the shape of the dataset?
# +
# How many spam/ham are there?
# +
# How much missing data is there?
| nlp_with_python_for_ml/Exercise Files/Ch01/01_04/Start/.ipynb_checkpoints/01_04-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # OpTaliX to CodeV to RayOptics / Rayopt
# At first install the python package "ray-optics" (conda install rayoptics --channel conda-forge).
#
# initialization of ray-optics
from rayoptics.environment import *
import os
cwd = os.getcwd()
filename = os.path.join(cwd, os.path.join('OpTaliX','AFL12-15.seq'))
root_pth = Path(rayoptics.__file__).resolve().parent
opm = open_model(filename)
sm = opm['seq_model']
osp = opm['optical_spec']
pm = opm['parax_model']
em = opm['ele_model']
pt = opm['part_tree']
ar = opm['analysis_results']
sm.list_model()
| _build/html/_sources/OpTaliX_CodeV_RayOptics.ipynb |
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''codeforecon'': conda)'
# name: python3
# ---
# # Intro to Mathematics with Code
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Set seed for reproducibility
np.random.seed(10)
# Set max rows displayed for readability
pd.set_option('display.max_rows', 6)
# Plot settings
plt.style.use('plot_style.txt')
# -
# In this chapter, you'll learn about doing mathematics with code, including solving equations both in the abstract and numerically.
#
# This chapter uses the **numpy**, **scipy**, and **sympy** packages. If you're running this code, you may need to install these packages using, for example, `pip install packagename` on your computer's command line. (If you're not sure what a command line or terminal is, take a quick look at the basics of coding chapter.)
#
# ## Symbolic mathematics
from myst_nb import glue
import sympy
a = 8
glue('sqrt', 2*np.sqrt(a))
glue('symsqrt', sympy.sqrt(a))
# When using computers to do mathematics, we're most often performing numerical computations such as $\sqrt{8} = ${glue:}`sqrt`. Although we have the answer, it's only useful for the one special case. Symbolic mathematics allows us to use coding to solve equations in the general case, which can often be more illuminating. As an example, if we evaluate this in symbolic mathematics we get $\sqrt{8} = ${glue:}`symsqrt`.
#
# The Python package for symbolic mathemtics is [**sympy**](https://www.sympy.org/en/index.html), which provides some features of a computer algebra system.
#
# To define *symbolic* variables, we use sympy's symbols function. For ease, we'll import the entire sympy library into the namespace by using `from sympy import *`.
from sympy import *
x, t, α, β = symbols(r'x t \alpha \beta')
# ```{note}
# The leading 'r' in some strings tells Python to treat the string literally so that backslashes are not treated as instructions--otherwise, combinations like `\n` would begin a newline.
# ```
#
# Having created these symbolic variables, we can refer to and see them just like normal variables--though they're not very interesting *because* they are just symbols (for now):
α
# Things get much more interesting when we start to do maths on them. Let's see some integration, for example, say we want to evaluate
Integral(log(x), x)
# (note that the symbols are printed as latex equations) we simply call
integrate(log(x), x)
# We can differentiate too:
diff(sin(x)*exp(x), x)
# and even take limits!
limit(sin(x)/x, x, 0)
# It is also possible to solve equations using **sympy**. The solve function tries to find the roots of $f(x)$ and has syntax `solve(f(x)=0, x)`. Here's an example:
solve(x*5 - 2, x)
# There are also solvers for differential equations (`dsolve`), continued fractions, simplifications, and more.
#
# Another really important thing to know about symbolic mathematics is that you can 'cash in' at any time by substituting in an actual value. For example,
expr = 1 - 2*sin(x)**2
expr.subs(x, np.pi/2)
# But you don't have to substitute in a real value; you can just as well substitute in a different symbolic variable:
expr = 1 - 2*sin(x)**2
simplify(expr.subs(x, t/2))
# I snuck in a simplify here too!
#
# ### Symbolic mathematics for economics
#
# The library does a lot, so let's focus on a few features that are likely to be useful for economics in particular.
#
# #### Series expansion
#
# The first is performing **Taylor series expansions**. These come up all the time in macroeconomic modelling, where models are frequently log-linearised. Let's see an example of a couple of expansions together:
# +
expr = log(sin(α))
expr.series(α, 0, 4)
# -
# This is a 3rd order expansion around $\alpha=0$.
#
# #### Symbolic linear algebra
#
# The support for **matrices** can also come in handy for economic applications. Here's a matrix,
M = Matrix([[1, 0, x], [α, -t, 3], [4, β, 2]])
M
# and its determinant:
M.det()
# I can hardly go to a talk in economics that involves matrices that doesn't see those matrices get diagonalised: there's a function for that too.
P, D = Matrix([[1, 0], [α, -t]]).diagonalize()
D
# #### Lagrangians
#
# Function optimisation using Lagrangians is about as prevalent in economics as any bit of maths: let's see how it's done symbolically.
#
# We're going to find the minimum over x, y of the function $f(x,y)$, subject to $g(x,y)=0$, where $f(x,y) = 4xy - 2x^2 + y^2$ and $g(x,y) = 3x+y-5$.
#
# First we need to specify the problem, and the Lagrangian for it, in code
# +
x, y, λ = symbols(r'x y \lambda', real=True)
f = 4*x*y - 2*x**2 + y**2
g = 3*x+y-5
ℒ = f - λ*g
ℒ
# -
# The Karush-Kuhn-Tucker (KKT) conditions tell us whether any solutions we find will be optimal. Simply, the constaint is that a solution vector is a saddle point of the Lagrangian, $\nabla \mathcal{L} = 0$. Let's solve this.
gradL = [diff(ℒ, c) for c in [x, y]]
KKT_eqns = gradL + [g]
KKT_eqns = gradL + [g]
glue('kkt_0', KKT_eqns[0])
glue('kkt_1', KKT_eqns[1])
glue('kkt_2', KKT_eqns[2])
# This gives 3 equations from the KKT conditions: {glue:}`kkt_0`, {glue:}`kkt_1`, and {glue:}`kkt_2`. (The symbolic manipulation is now over: we solved for the conditions in terms of algebra--now we're looking for real values.) Now we look for the values of $x, y$ that minimise $f$ given that $g=0$ by solving these equations over $x$, $y$, and $\lambda$.
stationary_pts = solve(KKT_eqns, [x, y, λ], dict=True)
stationary_pts
# Now, we can substitute these in to find the (first--and in this case only) point that minimises our function:
stationary_pts[0][x], stationary_pts[0][y], f.subs(stationary_pts[0])
# #### Exporting to latex
#
# To turn any equation, for example `diff(sin(x)*exp(x), x)`, into latex and export it to a file that can be included in a paper, use
#
# ```python
# eqn_to_export = latex(diff(sin(x)*exp(x), x), mode='equation')
# open('latex_equation.tex', 'w').write(eqn_to_export)
# ```
#
# which creates a file called 'latex_equation.tex' that has a single line in it: '\begin{equation}\int \log{\left(x \right)}\, dx\end{equation}'. There are a range of options for exporting to latex, `mode='equation*'` produces an unnumbered equation, 'inline' produces an inline equation, and so on. To include these in your latex paper, use '\input{latex_equation.tex}'.
#
# ### Why coding symbolic mathematics is useful
#
# 1. Accuracy--using a computer to solve the equations means you're less likely to make a mistake. At the very least, it's a useful check on your by-hand working.
#
# 2. Consistency--by making your code export the equations you're solving to your write-up, you can ensure that the equations are consistent across both *and* you only have to type them once.
#
# ## Numerical Mathematics
#
# For much of the time, you'll be dealing with numbers rather than symbols. The workhorses of numerical mathematics are the two packages **numpy** and **scipy**. Both have excellent documentation, where you can find out more. In this section, we'll look at how to use them in some standard mathematical operations that arise in economics.
#
# The most basic object is an array, which can be defined as follows:
import numpy as np
a = np.array([0, 1, 2, 3], dtype='int64')
a
# Arrays are very memory efficient and fast objects that you should use in preference to lists for any heavy duty numerical operation.
#
# To demonstrate this, let's do a time race between lists and arrays for squaring all elements of an array:
#
# Lists:
a_list = range(1000)
# %timeit [i**2 for i in a_list]
# Arrays:
a = np.arange(1000)
# %timeit a**2
# Using arrays was *two orders of magnitude** faster! Okay, so we should use arrays for numerical works. How do we make them? You can specify an array explicitly as we did above to create a vector. This manual approach works for other dimensions too:
mat = np.array([[0, 1, 2], [3, 4, 5], [6, 7, 8]])
mat
# To find out about matrix properties, we use `.shape`
mat.shape
# We already saw how `np.arange(start, stop, step)` produces a vector; `np.linspace(start, stop, number)` produces a vector of length `number` by equally dividing the space between `start` and `stop`.
#
# Three really useful arrays are `np.ones(shape)`, for example,
np.ones((3, 3))
# `np.diag` for diagnoal arrays,
np.diag(np.array([1, 2, 3, 4]))
# and `np.zeros` for empty arrays:
np.zeros((2, 2))
# Random numbers are supplied by `np.random.rand()` for a uniform distribution in [0, 1], and `np.random.randn()` for numbers drawn from a standard normal distribution.
#
# You can, of course, specify a function to create an array:
c = np.fromfunction(lambda i, j: i**2+j**2, (4, 5))
c
# To access values in an array, you can use all of the by-position slicing methods that you've seen already in data analysis and with lists. The figure gives an example of some common slicing operations:
#
# 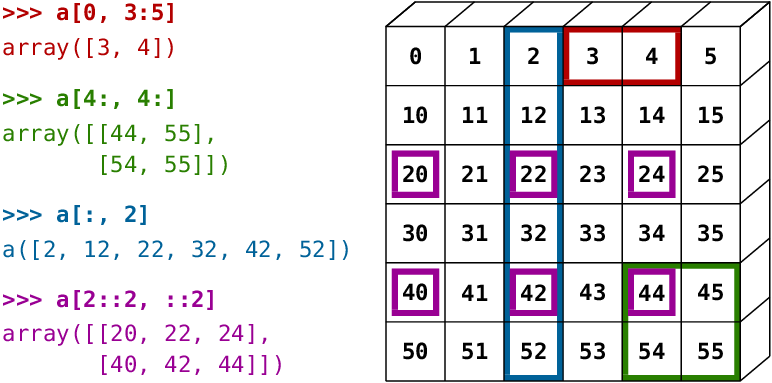
#
# Arrays can also be sliced and diced based on boolean indexing, just like a dataframe.
#
# For example, using the array defined above, we can create a boolean array of true and false values from a condition such as `c > 6` and use that to only access some elements of an array (it doesn't have to be the same array, though it usually is):
c[c > 6]
# As with dataframes, arrays can be combined. The main command to remember is `np.concatenate`, which has an `axis` keyword option.
x = np.eye(3)
np.concatenate([x, x], axis=0)
# Splitting is performed with `np.split(array, splits, axis=)`, for example
np.split(x, [3], axis=0)
# Aggregation operations are very similar to those found in dataframes: `x.sum(i)` to sum across the $i$th dimension of the array; similarly for standard deviation, and so on.
#
# As with dataframes, you can (and often should) specify the datatype of an array when you create it by passing a `dtype=` keyword, eg `c = np.array([1, 2, 3], dtype=float)`. To find out the data type of an array that already exists, use `c.dtype`.
#
# Finally, numpy does a lot of smart broadcasting of arrays. Broadcasting is what means that summing two arrays gives you a third array that has elements that are each the sum of the relevant elements in the two original arrays. Put another way, it's what causes `x + y = z` (for arrays x and y with the same shape) to result in an array z for which $z_{ij} = x_{ij} + y_{ij}$.
#
# Summing two arrays of the same shape is a pretty obvious example, but it also applies to cases that are *not* completely matched. For example, multiplication by a scalar is broadcast across all elements of an array:
x = np.ones(shape=(3, 3))
x*3
# Similarly, numpy functions are broadcast across elements of an array:
np.exp(x)
# ### Numeric linear algebra
#
# The transpose of an array `x` is given by `x.T`.
#
# Matrix multiplation is performed using the `@` operator. Here we perform $ M_{il} = \sum_{k} x_{ik} * (x^T)_{kl}$, where $x^T$ is the transpose of $x$.
x @ x.T
# To multiply two arrays element wise, ie to do $ M_{ij} = x_{ij} y_{ij}$, it's the usual multiplication operator `*`.
#
# Inverting matrices:
a = np.random.randint(9, size=(3, 3), dtype='int')
b = a @ np.linalg.inv(a)
b
# Computing the trace:
b.trace()
# Determinant:
np.linalg.det(a)
# Computing a Cholesky decomposition, i.e. finding lower triangular matrix $C$ such that $C C' = \Sigma$ for $\Sigma$ a 2-dimensional positive definite matrix.
# +
Σ = np.array([[4, 1], [1, 3]])
c = np.linalg.cholesky(Σ)
c @ c.T - Σ
# -
# #### Solving systems of linear equations
#
# Say we have a system of equations, $4x + 3y + 2z = 25$,
# $-2x + 2y + 3z = -10$, and $3x -5y + 2z = -4$. We can solve these three equations for the three unknowns, x, y, and z, using the `solve` method. First, remember that this equation can be written in matrix form as
#
# $$
# M\cdot \vec{x} = \vec{c}
# $$
#
# We can solve this by multiplying by the matrix inverse of $M$:
#
# $$
# M^{-1} M \cdot \vec{x} = I \cdot \vec{x} = M^{-1} \cdot \vec{c}
# $$
#
# which could be called by running `x = la.inv(M).dot(c)`. There's a convenience function in **numpy** called solve that does the same thing: here it finds the real values of the vector $\vec{x}$.
M = np.array([[4, 3, 2], [-2, 2, 3], [3, -5, 2]])
c = np.array([25, -10, -4])
np.linalg.solve(M, c)
# Finally, eigenvalues and eigenvectors can be found from:
import scipy.linalg as la
eigvals, eigvecs = la.eig(M)
eigvals
# ### Interpolation
#
# This section draws on the **scipy** documentation. There are built-in **pandas** methods for interpolation in dataframes, but **scipy** also has a range of functions for this including for univariate data `interp1d`, multidimensional interpolation on a grid `interpn`, `griddata` for unstructured data. Let's see a simple example with interpolation between a regular grid of integers.
import matplotlib.pyplot as plt
from scipy import interpolate
x = np.arange(0, 10)
y = np.exp(-x/3.0)
f = interpolate.interp1d(x, y, kind='cubic')
# Create a finer grid to interpolation function f
xnew = np.arange(0, 9, 0.1)
ynew = f(xnew)
plt.plot(x, y, 'o', xnew, ynew, '-')
plt.show()
# What about unstructured data? Let's create a Cobb-Douglas function on a detailed grid but then only retain a random set of the established points.
# +
from scipy.interpolate import griddata
def cobb_doug(x, y):
alpha = 0.8
return x**(alpha)*y**(alpha-1)
# Take some random points of the Cobb-Douglas function
points = np.random.rand(1000, 2)
values = cobb_doug(points[:,0], points[:,1])
# Create a grid
grid_x, grid_y = np.mgrid[0.01:1:200j, 0.01:1:200j]
# Interpolate the points we have onto the grid
interp_data = griddata(points, values, (grid_x, grid_y), method='cubic')
# Plot results
fig, axes = plt.subplots(1, 2)
# Plot function & scatter of random points
axes[0].imshow(cobb_doug(grid_x, grid_y).T,
extent=(0, 1, 0, 1),
origin='lower', cmap='plasma_r',
vmin=0, vmax=1)
axes[0].plot(points[:, 0], points[:, 1], 'r.', ms=1.2)
axes[0].set_title('Original + points')
# Interpolation of random points
axes[1].imshow(interp_data.T, extent=(0, 1, 0, 1),
origin='lower', cmap='plasma_r',
vmin=0, vmax=1)
axes[1].set_title('Cubic interpolation');
# -
# ### Optimisation
#
# **scipy** has functions for minimising scalar functions, minimising multivariate functions with complex surfaces, and root-finding. Let's see an example of finding the minimum of a scalar function.
# +
from scipy import optimize
def f(x):
return x**2 + 10*np.sin(x) - 1.2
result = optimize.minimize(f, x0=0)
result
# -
# The result of the optimisation is in the 'x' attribute of `result`. Let's see this:
x = np.arange(-10, 10, 0.1)
fig, ax = plt.subplots()
ax.plot(x, f(x))
ax.scatter(result.x, f(result.x), s=150, color='k')
ax.set_xlabel('x')
ax.set_ylabel('f(x)', rotation=90)
plt.show()
# In higher dimensions, the minimisation works in much the same way, with the same function `optimize.minimize`. There are a LOT of minimisation options that you can pass to the `method=` keyword; the default is intelligently chosen from BFGS, L-BFGS-B, or SLSQP, depending upon whether you supply constraints or bounds.
#
# Root finding, aka solving equations of the form $f(x)=0$, is also catered for by **scipy**, through `optimize.root`. It works in much the same way as `optimizer.minimize`.
#
# In both of these cases, be warned that multiple roots and multiple minima can be hard to detect, and you may need to carefully specify the bounds or the starting positions in order to find the root you're looking for. Also, both of these methods can accept the Jacobian of the function you're working with as an argument, which is likely to improve performance with some solvers.
#
# ### Numerical Integration
#
# **scipy** provides routines to numerically evaluate integrals in `scipy.integrate`, which you can find the documentation for [here](https://docs.scipy.org/doc/scipy/reference/integrate.html). Let's see an example using the 'vanilla' integration method, `quad`, to solve a known function between given (numerical) limits:
#
# $$
# \displaystyle\int_0^{\pi} \sin(x) d x
# $$
from scipy.integrate import quad
res, err = quad(np.sin, 0, np.pi)
res
# What if we just have data samples? In that case, there are several routines that perform purely numerical integration:
# +
from scipy.integrate import simps
x = np.arange(0, 10)
f_of_x = np.arange(0, 10)
simps(f_of_x, x) - 9**2/2
# -
# Even with just 10 evenly spaced points, the composite Simpson’s rule integration given by `simps` is able to accurately find the answer as $\left( x^2/2\right) |_{0}^{9}$.
#
# ## Advanced: Composable Function Transformations
#
# In recent years, there have been great developments in the ability of Python to easily carry out numerical 'composable function transformations'. What this means is that, if you can dream up an arbitrary numerical operations -- including differentiation, linear algebra, and optimisation -- you can write code that will execute it quickly and automatically on CPUs, GPUs, or TPUs as you like.
#
# Here we'll look at one library that does this, **jax**, developed by Google {cite}`jax2018github`. It can automatically differentiate native Python and **numpy** functions, including when they are in loops, branches, or subject to recursion, and it can take derivatives of derivatives of derivatives. It supports reverse-mode differentiation (a.k.a. backpropagation) via grad as well as forward-mode differentiation, and the two can be composed arbitrarily to any order.
#
# To do these at speed, it uses just-in-time compilation. If you don't know what that is, don't worry: the details aren't important. It's just a way of getting close to C++ or Fortran speeds while still being able to write code in *much* more user friendly Python!
#
# ### Auto-differentiation
#
# Let's see an example of auto-differentiation an arbitrary function. We'll write the definition of $\tanh(x)$ as a function and evaluate it. Because we already imported a (symbolic) `tanh` function from Sympy above, we'll call the function below `tanh_num`.
# ```python
# from jax import grad
# import jax.numpy as jnp
#
# def tanh_num(θ): # Define a function
# y = jnp.exp(-2.0 * θ)
# return (1.0 - y) / (1.0 + y)
#
# grad_tanh = grad(tanh_num) # Obtain its gradient function
# grad_tanh(1.0) # Evaluate it at x = 1.0
# ```
#
# ```bash
# DeviceArray(0.4199743, dtype=float32)
# ```
# You can differentiate to any order using grad:
# ```python
# grad(grad(grad(tanh_num)))(1.0)
# ```
# ```bash
# DeviceArray(0.6216266, dtype=float32)
# ```
# Let's check this using symbolic mathematics:
θ = Symbol(r'\theta')
triple_deriv = diff(diff(diff(tanh(θ), θ)))
triple_deriv
symp_est = triple_deriv.subs(θ, 1.)
glue('symp_est', f'{symp_est:.3f}')
# If we evaluate this at $\theta=1$, we get {glue:}`symp_est`. This was a simple example that had a (relatively) simple mathematical expression. But imagine if we had lots of branches (eg if, else statements), and/or a really complicated function: **jax**'s grad would still work. It's designed for really complex derivatives of the kind encountered in machine learning.
#
# ### Just-in-time compilation
#
# The other nice feature of **jax** is the ability to do just-in-time (JIT) compilation. Because they do not compile their code into machine-code before running, high-level languages like Python and R are not as fast as the same code written in C++ or Fortran (the benefit is that it takes you less time to write the code in the first place). Much of the time, there are pre-composed functions that call C++ under the hood to do these things--but only for those operations that people have already taken the time to code up in a lower level language. JIT compilation offers a compromise: you can code more or less as you like in the high-level language but it will be compiled just-in-time to give you a speed-up!
#
# **jax** is certainly not the only Python package that does this, and if you're not doing anything like differentiating or propagating, **numba** is a more mature alternative. But here we'll see the time difference for JIT compilation on an otherwise slow operation: element wise multiplication and addition.
# ```python
# from jax import jit
#
# def slow_f(x):
# """Slow, element-wise function"""
# return x * x + x * 2.0
#
# x = jnp.ones((5000, 5000))
# fast_f = jit(slow_f)
# ```
# Now let's see how fast the 'slow' version goes:
# ```python
# # %timeit -n15 -r3 slow_f(x)
# ```
# ```bash
# 60.1 ms ± 3.67 ms per loop (mean ± std. dev. of 3 runs, 15 loops each)
# ```
# what about with the JIT compilation?
# ```python
# # %timeit -n15 -r3 fast_f(x)
# ```
# ```bash
# 17.7 ms ± 434 µs per loop (mean ± std. dev. of 3 runs, 15 loops each)
# ```
# This short introduction has barely scratched the surface of **jax** and what you can do with it. For more, see the [official documentation](https://jax.readthedocs.io/en/latest/).
#
# ## Set theory
#
# Set theory is a surprisingly useful tool in research (and invaluable in spatial analysis). Here are some really useful bits of set theory inspired by examples in {cite}`sheppard2012introduction`.
#
# Sets are first class citizens in Python in the same way that lists are. We can define and view a set like this:
x = set(['<NAME>', '<NAME>',
'<NAME>', '<NAME>',
'<NAME>', '<NAME>'])
x
# Notice that a couple of entries appeared twice in the list but only once in the set: that's because a set contains only unique elements. Let's define a second set in order to demonstrate some of the operations we can perform on sets.
y = set(['<NAME>', '<NAME>',
'<NAME>', '<NAME>',
'<NAME>', '<NAME>'])
y
from myst_nb import glue
inters = x.intersection(y)
differ = x.difference(y)
union = x.union(y)
glue("inters", inters)
glue("differ", differ)
glue("union", union)
# Now we have two sets we can look at to demonstrate some of the basic functions you can call on the set object type. `x.intersection(y)` gives, in this example, {glue:}`inters`, `x.difference(y)` gives {glue:}`differ`, and `x.union(y)` gives {glue:}`union`.
#
# **numpy** also has functions that use set theory. `np.unique` returns only the unique entries of an input array or list:
np.unique(['Lovelace', 'Hopper', 'Alexander', 'Hopper', 45, 27, 45])
# We can also ask which of a second set is a repeat of a first:
x = np.arange(10)
y = np.arange(5, 10)
np.in1d(x, y)
# And we have the numpy equivalents of intersection, `np.intersect1d(x, y)`, difference, `np.setdiff1d(x, y)`, and union, `np.union1d(x, y)`. Additionally, there is the exclusive-or (that I like to call 'xor'). This effectively returns the two arrays with their union removed:
a = np.array([1, 2, 3, 2, 4])
b = np.array([2, 3, 5, 7, 5])
np.setxor1d(a,b)
# ## Review
#
# In this chapter, you should have:
#
# - ✅ seen how to use symbolic algebra with code, including Lagrangrians and linear algebra;
# - ✅ seen how to code numerical mathematics, including linear algebra and optimisation; and
# - ✅ found out about using set theory via the `set` object type and set-oriented functions.
| maths-maths-coding.ipynb |