code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !wget http://cs.stanford.edu/~danqi/data/cnn.tar.gz
# !tar -xvzf cnn.tar.gz
import numpy as np
import matplotlib.pyplot as plt
keys = ['train', 'dev', 'test']
p, q, a = {}, {}, {}
for k in keys :
file = open('cnn/' + k + '.txt').read().strip().split('\n\n')
file = [x.split('\n') for x in file]
p[k] = [x[2] for x in file]
q[k] = [x[0] for x in file]
a[k] = [x[1] for x in file]
# +
entities = {}
for k in p :
entities[k] = []
for x in p[k] :
entities[k] += [y for y in x.split() if y.startswith('@entity')]
entities[k] = set(entities[k])
f = open('entity_list.txt', 'w')
f.write('\n'.join(list(entities['train'])))
f.close()
# -
def generate_possible_answers(p) :
possible_answers = []
for w in p.split() :
if w.startswith('@entity') :
possible_answers.append(w)
return ";".join(list(set(possible_answers)))
# +
import pandas as pd
df_paragraphs = []
df_questions = []
df_answers = []
df_possible_answers = []
df_exp_splits = []
for k in keys :
df_paragraphs += p[k]
df_questions += q[k]
df_answers += a[k]
df_possible_answers += [generate_possible_answers(x) for x in p[k]]
df_exp_splits += [k] * len(p[k])
df = {'paragraph' : df_paragraphs, 'question' : df_questions, 'answer' : df_answers,
'exp_split' : df_exp_splits, 'possible_answers' : df_possible_answers}
df = pd.DataFrame(df)
# -
df.to_csv('cnn_dataset.csv', index=False)
# %run "../preprocess_data_QA.py" --data_file cnn_dataset.csv --output_file ./vec_cnn.p --all_answers_file entity_list.txt \
# --word_vectors_type fasttext.simple.300d --min_df 8 --add_answers_to_vocab
| project/Transparency/preprocess/CNN/CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from matplotlib import pyplot as plt
x = range(2,26,2)
y = [15,13,14.5,17,20,25,26,26,24,22,18,15]
plt.plot(x,y)
plt.show()
# plt.figure() 可以设置图片大小:
# figure(figsize=(width,height),dpi=x) dpi 在图像模糊的时候可以传入dpi参数,使其清晰
x = range(2,26,2)
y = [15,13,14.5,17,20,25,26,26,24,22,18,15]
plt.figure(figsize=(20,8),dpi=70)
plt.plot(x,y)
plt.savefig('./sig_size.png') # 保存当前生成的图片,也可以保存为矢量图,放大不会有锯齿
plt.show()
# 修改x轴的数据的宽度
x = range(2,26,2)
y = [15,13,14.5,17,20,25,26,26,24,22,18,15]
plt.figure(figsize=(20,8),dpi=70)
plt.plot(x,y)
plt.xticks(range(1,25)) # x轴的取值范围变为1-24
plt.show()
# 修改x轴的数据的宽度
x = range(2,26,2)
y = [15,13,14.5,17,20,25,26,26,24,22,18,15]
plt.figure(figsize=(20,8),dpi=70)
plt.plot(x,y)
_xtick_lables = [i/2 for i in range(4,49)]
plt.xticks(_xtick_lables[::3]) # x轴的取值范围变为1-24
plt.show()
# y轴的刻度控制
x = range(2,26,2)
y = [15,13,14.5,17,20,25,26,26,24,22,18,15]
plt.figure(figsize=(20,7),dpi=70)
plt.plot(x,y)
plt.xticks(range(1,25)) # x轴的取值范围变为1-24
plt.yticks(range(min(y),max(y)+1))
plt.show()
# 如果列表a表示10点到12点每一分钟的气温,如何绘制折线图观察每分钟的气温变化。
import random
x = range(0,120)
y = [random.randint(20,35) for i in range(120)]
plt.figure(figsize=(20,8),dpi=80)
plt.plot(x,y)
plt.show()
plt.rc('font',family="Microsoft YaHei")
# 将横坐标设置为时间
import random
x = range(0,120)
y = [random.randint(15,20) for i in range(120)]
plt.figure(figsize=(20,8),dpi=80)
plt.plot(x,y)
_xtick_lables = ["10点{}分".format(i) for i in range(60)]
_xtick_lables += ["11点{}分".format(i) for i in range(60)]
plt.xticks(list(x)[::3],_xtick_lables[::3],rotation=45) # rotation 将lables旋转 参数是旋转的角度
plt.show()
# +
#添加描述信息:
x = range(0,120)
y = [random.randint(20,35) for i in range(120)]
plt.figure(figsize=(20,8),dpi=80)
plt.plot(x,y)
_xtick_lables = ["10点{}分".format(i) for i in range(60)]
_xtick_lables += ["11点{}分".format(i) for i in range(60)]
plt.xticks(list(x)[::3],_xtick_lables[::3],rotation=45) # rotation 将lables旋转 参数是旋转的角度
plt.xlabel('时间')
plt.ylabel('温度')
plt.title('10点到12点温度变化曲线')
plt.show()
# +
## 练习题
# 假设大家在30岁的时候,根据自己的实际情况,统计出来了从11岁到30岁每年交的女(男)朋友的数量如列表a,请绘制出该数据的折线图,以便分析自己每年交女(男)朋友的数量走势
# a = [1,0,1,1,2,4,3,2,3,4,4,5,6,5,4,3,3,1,1,1]
# 要求:
# y轴表示个数
# x轴表示岁数,比如11岁,12岁等
y = [1,0,1,1,2,4,3,2,3,4,4,5,6,5,4,3,3,1,1,1]
x = range(11,31)
plt.figure(figsize=(20,8),dpi=80)
plt.plot(x,y)
_xtick_lables = ["{}岁".format(i) for i in x]
plt.xticks(x,_xtick_lables)
plt.grid() # 设置网格
plt.show()
# -
# # 这是一个练习
# #### 练习
# 假设大家在30岁的时候,根据自己的实际情况,统计出来了你和你同桌各自从11岁到30岁每年交的女(男)朋友的数量如列表a和b,
# 请在一个图中绘制出该数据的折线图,以便比较自己和同桌20年间的差异,同时分析每年交女(男)朋友的数量走势
# a = [1,0,1,1,2,4,3,2,3,4,4,5,6,5,4,3,3,1,1,1]
# b = [1,0,3,1,2,2,3,3,2,1 ,2,1,1,1,1,1,1,1,1,1]
# 要求:
# y轴表示个数
# x轴表示岁数,比如11岁,12岁等
# +
y_1 = [1,0,1,1,2,4,3,2,3,4,4,5,6,5,4,3,3,1,1,1]
y_2 = [1,0,3,1,2,2,3,3,2,1 ,2,1,1,1,1,1,1,1,1,1]
x = range(11,31)
plt.figure(figsize=(20,8),dpi=80)
plt.plot(x,y_1,label='自己',color='r',linestyle='--',linewidth=1,alpha=0.5)
plt.plot(x,y_2,label='同桌')
_xtick_lables = ["{}岁".format(i) for i in x]
plt.xticks(x,_xtick_lables)
plt.legend(loc=2) # loc表示图例的位置,具体的参数要去看文件的源码
plt.grid(linestyle='--')
plt.show()
# 在画图形的线条的时候我们可以有一些参数进行设置
# color 线条的颜色,或者使用十六进制
# linestyle 线条的样式
# linewidth 线条宽度
# alpha 透明度
# -
# ## 绘制散点图
# 假设通过爬虫你获取到了北京2016年3,10月份每天白天的最高气温(分别位于列表a,b),那么此时如何寻找出气温和随时间(天)变化的某种规律?
#
# a = [11,17,16,11,12,11,12,6,6,7,8,9,12,15,14,17,18,21,16,17,20,14,15,15,15,19,21,22,22,22,23]
#
# b = [26,26,28,19,21,17,16,19,18,20,20,19,22,23,17,20,21,20,22,15,11,15,5,13,17,10,11,13,12,13,6]
#
# +
m_3 = [11,17,16,11,12,11,12,6,6,7,8,9,12,15,14,17,18,21,16,17,20,14,15,15,15,19,21,22,22,22,23]
m_10 = [26,26,28,19,21,17,16,19,18,20,20,19,22,23,17,20,21,20,22,15,11,15,5,13,17,10,11,13,12,13,6]
x_3 = range(1,32)
x_10 = range(51,82)
plt.figure(figsize=(20,8),dpi=80)
plt.scatter(x_3,m_3,label='3月份')
plt.scatter(x_10,m_10,label='10月份')
# 调整x轴的刻度
_x = list(x_3)+list(x_10)
_xtrick_label = ["3月{}日".format(i) for i in x_3] + ["10月{}日".format(i) for i in x_10]
plt.xticks(_x[::3],_xtrick_label[::3],rotation=45)
# 添加图例
plt.legend()
# 添加描述信息
plt.xlabel('时间')
plt.ylabel('温度')
plt.title('温度和时间的散点图')
plt.show()
# -
# ## 绘制条形图
#
# a = ["战狼2","速度与激情8","功夫瑜伽","西游伏妖篇","变形金刚5:最后的骑士","摔跤吧!爸爸","加勒比海盗5:死无对证","金刚:骷髅岛","极限特工:终极回归","生化危机6:终章","乘风破浪","神偷奶爸3","智取威虎山","大闹天竺","金刚狼3:殊死一战","蜘蛛侠:英雄归来","悟空传","银河护卫队2","情圣","新木乃伊",]
#
# b=[56.01,26.94,17.53,16.49,15.45,12.96,11.8,11.61,11.28,11.12,10.49,10.3,8.75,7.55,7.32,6.99,6.88,6.86,6.58,6.23]
#
# 单位:亿
# +
# 通过添加/n进行调整
a = ["战狼2","速度与激情8","功夫瑜伽","西游伏妖篇","变形金刚5:\n最后的骑士","摔跤吧!爸爸","加勒比海盗5:\n死无对证",
"金刚:骷髅岛","极限特工:\n终极回归","生化危机6:\n终章","乘风破浪","神偷奶爸3","智取威虎山","大闹天竺",
"金刚狼3:\n殊死一战","蜘蛛侠:\n英雄归来","悟空传","银河护卫队2","情圣","新木乃伊",]
b=[56.01,26.94,17.53,16.49,15.45,12.96,11.8,11.61,11.28,11.12,10.49,10.3,8.75,7.55,7.32,6.99,6.88,6.86,6.58,6.23]
plt.figure(figsize=(20,8),dpi=80)
plt.bar(range(len(a)),b,width=0.5)
plt.xticks(range(len(a)),a,rotation=45)
plt.show()
# +
# 将条形图的反转
a = ["战狼2","速度与激情8","功夫瑜伽","西游伏妖篇","变形金刚5:最后的骑士","摔跤吧!爸爸","加勒比海盗5:死无对证",
"金刚:骷髅岛","极限特工:终极回归","生化危机6:终章","乘风破浪","神偷奶爸3","智取威虎山","大闹天竺",
"金刚狼3:殊死一战","蜘蛛侠:英雄归来","悟空传","银河护卫队2","情圣","新木乃伊",]
b=[56.01,26.94,17.53,16.49,15.45,12.96,11.8,11.61,11.28,11.12,10.49,10.3,8.75,7.55,7.32,6.99,6.88,6.86,6.58,6.23]
plt.figure(figsize=(20,8),dpi=80)
plt.barh(range(len(a)),b,height=0.5,color='orange')
plt.yticks(range(len(a)),a)
plt.grid(alpha=0.4)
plt.xlabel('票房/忆')
plt.ylabel('电影名')
plt.title('票房与电影的条形统计图')
plt.show()
# -
# ## 绘制条形图
# 假设你知道了列表a中电影分别在2017-09-14(b_14), 2017-09-15(b_15), 2017-09-16(b_16)三天的票房,为了展示列表中电影本身的票房以及同其他电影的数据对比情况,应该如何更加直观的呈现该数据?
#
# a = ["猩球崛起3:终极之战","敦刻尔克","蜘蛛侠:英雄归来","战狼2"]
#
# b_16 = [15746,312,4497,319]
#
# b_15 = [12357,156,2045,168]
#
# b_14 = [2358,399,2358,362]
#
# +
a = ["猩球崛起3:终极之战","敦刻尔克","蜘蛛侠:英雄归来","战狼2"]
b_16 = [15746,312,4497,319]
b_15 = [12357,156,2045,168]
b_14 = [2358,399,2358,362]
x_14 = list(range(len(a)))
x_15 = [i+0.2 for i in x_14]
x_16 = [i+0.4 for i in x_14]
# 设置图像大小
plt.figure(figsize=(20,8),dpi=80)
plt.bar(range(len(a)),b_14,width=0.2,label='9月14日')
plt.bar(x_15,b_15,width=0.2,label='9月15日')
plt.bar(x_16,b_16,width=0.2,label='9月16日')
# 设置x轴的刻度
plt.xticks(x_15,a)
plt.legend()
plt.xlabel('电影名')
plt.ylabel('票房')
plt.title('票房与电影的条形统计图')
plt.show()
# -
# ## 绘制直方图
#
# 假设你获取了250部电影的时长(列表a中),希望统计出这些电影时长的分布状态(比如时长为100分钟到120分钟电影的数量,出现的频率)等信息,你应该如何呈现这些数据?
#
# a=[131, 98, 125, 131, 124, 139, 131, 117, 128, 108, 135, 138, 131, 102, 107, 114, 119, 128, 121, 142, 127, 130, 124, 101, 110, 116, 117, 110, 128, 128, 115, 99, 136, 126, 134, 95, 138, 117, 111,78, 132, 124, 113, 150, 110, 117, 86, 95, 144, 105, 126, 130,126, 130, 126, 116, 123, 106, 112, 138, 123, 86, 101, 99, 136,123, 117, 119, 105, 137, 123, 128, 125, 104, 109, 134, 125, 127,105, 120, 107, 129, 116, 108, 132, 103, 136, 118, 102, 120, 114,105, 115, 132, 145, 119, 121, 112, 139, 125, 138, 109, 132, 134,156, 106, 117, 127, 144, 139, 139, 119, 140, 83, 110, 102,123,107, 143, 115, 136, 118, 139, 123, 112, 118, 125, 109, 119, 133,112, 114, 122, 109, 106, 123, 116, 131, 127, 115, 118, 112, 135,115, 146, 137, 116, 103, 144, 83, 123, 111, 110, 111, 100, 154,136, 100, 118, 119, 133, 134, 106, 129, 126, 110, 111, 109, 141,120, 117, 106, 149, 122, 122, 110, 118, 127, 121, 114, 125, 126,114, 140, 103, 130, 141, 117, 106, 114, 121, 114, 133, 137, 92,121, 112, 146, 97, 137, 105, 98, 117, 112, 81, 97, 139, 113,134, 106, 144, 110, 137, 137, 111, 104, 117, 100, 111, 101, 110,105, 129, 137, 112, 120, 113, 133, 112, 83, 94, 146, 133, 101,131, 116, 111, 84, 137, 115, 122, 106, 144, 109, 123, 116, 111,111, 133, 150]
#
# +
a=[131, 98, 125, 131, 124, 139, 131, 117, 128, 108, 135, 138, 131, 102, 107, 114, 119, 128, 121, 142, 127, 130,
124, 101, 110, 116, 117, 110, 128, 128, 115, 99, 136, 126, 134, 95, 138, 117, 111,78, 132, 124, 113, 150,
110, 117, 86, 95, 144, 105, 126, 130,126, 130, 126, 116, 123, 106, 112, 138, 123, 86, 101, 99, 136,123,
117, 119, 105, 137, 123, 128, 125, 104, 109, 134, 125, 127,105, 120, 107, 129, 116, 108, 132, 103, 136, 118,
102, 120, 114,105, 115, 132, 145, 119, 121, 112, 139, 125, 138, 109, 132, 134,156, 106, 117, 127, 144, 139,
139, 119, 140, 83, 110, 102,123,107, 143, 115, 136, 118, 139, 123, 112, 118, 125, 109, 119, 133,112, 114,
122, 109, 106, 123, 116, 131, 127, 115, 118, 112, 135,115, 146, 137, 116, 103, 144, 83, 123, 111, 110, 111,
100, 154,136, 100, 118, 119, 133, 134, 106, 129, 126, 110, 111, 109, 141,120, 117, 106, 149, 122, 122, 110,
118, 127, 121, 114, 125, 126,114, 140, 103, 130, 141, 117, 106, 114, 121, 114, 133, 137, 92,121, 112, 146,
97, 137, 105, 98, 117, 112, 81, 97, 139, 113,134, 106, 144, 110, 137, 137, 111, 104, 117, 100, 111, 101,
110,105, 129, 137, 112, 120, 113, 133, 112, 83, 94, 146, 133, 101,131, 116, 111, 84, 137, 115, 122, 106,
144, 109, 123, 116, 111,111, 133, 150]
#计算组数
d = 3 #组距
num_bins = (max(a)-min(a))//d
# print(max(a),min(a),max(a)-min(a))
# print(num_bins)
#设置图形的大小
plt.figure(figsize=(20,8),dpi=80)
# density True 改为频率分布直方图
plt.hist(a,num_bins,density=True)
#设置x轴的刻度
plt.xticks(range(min(a),max(a)+d,d))
plt.grid()
plt.show()
# -
# ## 练习
#
# 在美国2004年人口普查发现有124 million的人在离家相对较远的地方工作。根据他们从家到上班地点所需要的时间,通过抽样统计(最后一列)出了下表的数据,这些数据能够绘制成直方图么?
#
# interval = [0,5,10,15,20,25,30,35,40,45,60,90]
#
# width = [5,5,5,5,5,5,5,5,5,15,30,60]
#
# quantity = [836,2737,3723,3926,3596,1438,3273,642,824,613,215,47]
#
# +
interval = [0,5,10,15,20,25,30,35,40,45,60,90]
width = [5,5,5,5,5,5,5,5,5,15,30,60]
quantity = [836,2737,3723,3926,3596,1438,3273,642,824,613,215,47]
print(len(interval),len(width),len(quantity))
#设置图形大小
plt.figure(figsize=(20,8),dpi=80)
plt.bar(interval,quantity,width=width)
#设置x轴的刻度
temp_d = [5]+ width[:-1]
_x = [i-temp_d[interval.index(i)]*0.5 for i in interval]
plt.xticks(_x,interval)
plt.grid(alpha=0.4)
plt.show()
# -
| matplotlob/first.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python395jvsc74a57bd09400c0b303c55aae944a7d117953411ee2c353c3ca48e8a9c758ac4a17625cdf
# ---
# ## Fonction de scraping des tirages
# # Cas de prédiction du Loto français
#Import des librairies utiles
from bs4 import BeautifulSoup
import time
import requests
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pickle
from sklearn.preprocessing import StandardScaler
from keras.models import Sequential
from keras.layers import LSTM, Dense, Bidirectional, TimeDistributed, RepeatVector, Flatten
from keras.callbacks import EarlyStopping
#fonction de scraping des tirages du loto
def scrap_loto_numbers():
my_list=[]
time.sleep(2)
loto_url = "http://loto.akroweb.fr/loto-historique-tirages/"
page = requests.get(loto_url)
soup = BeautifulSoup(page.text, 'html.parser')
body = soup.find('table')
tirage_line = body.find_all('tr')
for value in tirage_line:
my_dict = {}
res = value.text.split('\n')
my_dict['day']=res[2]
my_dict['month_year']=res[3]
for i,val in enumerate(res[5:10]):
my_dict['num'+str(i)]=int(val)
my_dict['chance']=int(res[10])
my_list.append(my_dict)
df=pd.DataFrame(my_list)
return df
# +
#A noter que plusieurs tirages se sont ajoutés dépuis le 21 : date de rédaction de l'article
# -
#sracping des tirages actuellement disponibles sur le site
df_tirage=scrap_loto_numbers()
df_tirage[['day','month_year','num0','num1','num2','num3','num4','chance']].head()
#suppression des tirages du super loto( A explorer later )
#df_tirage=df_tirage[(df_tirage['day']!='Vendredi') & (df_tirage['day']!='Mardi')]
# +
#df_tirage=df_tirage.tail(df_tirage.shape[0]-1)
# -
# ## commentaires:
# * le dernier tirage ici date du 07 décembre, ainsi afin de tester le modèle nous allons rétirer ce tirage du dataset dans la suite
# * Par contre on aurait évité de supprimer le tirage du 28 si on voulait prédire le prochain tirage ( celui du samedi 31)
#df_tirage=df_tirage.tail(df_tirage.shape[0])# suppression du dernier tirage/à éviter selon le cas
df_tirage.head()# le dernier tirage devient ici celui du 26
# ## Traitement des données
df = df_tirage.iloc[::-1]#inversion du dataframe pour placer le dernier tirage en dernière position
df = df[['num0', 'num1', 'num2', 'num3', 'num4', 'chance']]#sélection des numéros à traiter
df.tail()# notre tirage du 26 ici devient le dernier de notre dataset afin de pourvoir organiser les data par historique
# +
#fonction de vérification de nombres en dessous d'une certaine valeur pour les 5 premiers numéros, sauf celui de chance
def is_under(data, number):
return ((data['num0'] <= number).astype(int) +
(data['num1'] <= number).astype(int) +
(data['num2'] <= number).astype(int) +
(data['num3'] <= number).astype(int) +
(data['num4'] <= number).astype(int))
#fonction de vérification de nombres pairs pour les 5 premiers numéros sauf celui de chance
def is_pair(data):
return ((data['num0'].isin(pairs)).astype(int) +
(data['num1'].isin(pairs)).astype(int) +
(data['num2'].isin(pairs)).astype(int) +
(data['num3'].isin(pairs)).astype(int) +
(data['num4'].isin(pairs)).astype(int))
#fonction de vérification de nombres impairs pour les 5 premiers numéros sauf celui de chance
def is_impair(data):
return ((data['num0'].isin(impairs)).astype(int) +
(data['num1'].isin(impairs)).astype(int) +
(data['num2'].isin(impairs)).astype(int) +
(data['num3'].isin(impairs)).astype(int) +
(data['num4'].isin(impairs)).astype(int))
#fonction de vérification de nombres pairs pour le numéro de chance
def is_pair_etoile(data):
return (data['chance'].isin(pairs)).astype(int)
#fonction de vérification de nombres impairs pour le numéro de chance
def is_impair_etoile(data):
return (data['chance'].isin(impairs)).astype(int)
#liste de nombres pairs et impairs
pairs = [2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50]
impairs = [1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49]
#Fonction de calcul de la somme de la différence au carré des 5 premiers numéros, sauf celui de chance
def sum_diff(data):
return ((data['num1'] - data['num0'])**2 +
(data['num2'] - data['num1'])**2 +
(data['num3'] - data['num2'])**2 +
(data['num4'] - data['num3'])**2)
# Calcul de la fréquence de tirage de chaque numéro
freqs = []
for val in range(50):
count = ( (df['num0'] == val+1).sum() +
(df['num1'] == val+1).sum() +
(df['num2'] == val+1).sum() +
(df['num3'] == val+1).sum() +
(df['num4'] == val+1).sum() )
freqs.append(count)
ax = plt.gca() ; ax.invert_yaxis()
plt.gcf().set_size_inches(5, 4)
heatmap = plt.pcolor(np.reshape(np.array(freqs), (5, 10)), cmap=plt.cm.Blues)
def freq_val(data, column):
tab = data[column].values.tolist()
freqs = []
pos = 1
for e in tab:
freqs.append(tab[0:pos].count(e))
pos = pos + 1
return freqs
#df['sum'] = ((df.num0 + df.num1 + df.num2 + df.num3 + df.num4 + df.chance ) >185).astype(int)
# -
#ajout de la difference entre les numéros(A explorer ASAp)
#for i in range(4):
#print(i,i+1)
#df['diff_{}'.format(i)]=df['num{}'.format(i+1)]-df['num{}'.format(i)]
#application des fonctions sur le dataframe
df['freq_num0'] = freq_val(df, 'num0')
df['freq_num1'] = freq_val(df, 'num1')
df['freq_num2'] = freq_val(df, 'num2')
df['freq_num3'] = freq_val(df, 'num3')
df['freq_num4'] = freq_val(df, 'num4')
df['freq_chance'] = freq_val(df, 'chance')#calcul des frequences
df['sum_diff'] = sum_diff(df)#somme de la différence au carré entre chaque couple de numéros successifs dans le tirage
df['pair_chance'] = is_pair_etoile(df)
df['impair_chance'] = is_impair_etoile(df)
df['pair'] = is_pair(df)
df['impair'] = is_impair(df)#verification de nombre pair et impair
df['is_under_24'] = is_under(df, 24) # Les numeros en dessous de 24
df['is_under_40'] = is_under(df, 40)# Les numeros en dessous de 40
df.head(6)
# ## Modèle et fonction de formatage des données en entrée du LSTM
# +
#capture 3: fonction define model seulement
# +
# j'ai ici défini plusieurs modèles à tester mais pour l'intant je tavaille avec le lstm(fonction : define_model)
# j'ai ici défini window_length à 12 pour apprendre sur 1 mois de données
#Params du modèle
nb_label_feature=6
UNITS = 100
BATCHSIZE = 30
EPOCH = 1500
#ACTIVATION = "softmax"
OPTIMIZER ='adam' # rmsprop, adam, sgd
LOSS = 'mae'#'categorical_crossentropy' #mse
DROPOUT = 0.1
window_length =12 #12
number_of_features = df.shape[1]
#Architecture du modèle
def define_model(number_of_features,nb_label_feature):
#initialisation du rnn
model = Sequential()
#ajout de la premiere couche lstm
model.add(LSTM(UNITS, input_shape=(window_length, number_of_features), return_sequences=True))
model.add(LSTM(UNITS, dropout=0.1, return_sequences=False))
#ajout de la couche de sortie
model.add(Dense(nb_label_feature))
model.compile(loss=LOSS, optimizer=OPTIMIZER, metrics=['acc'])
return model
def define_bidirectionnel_model(number_of_features,nb_label_feature):
model = Sequential()
model.add(Bidirectional(LSTM(100, dropout=0.2, return_sequences=True), input_shape=(window_length, number_of_features)))
model.add(LSTM(50, return_sequences=True))
model.add(LSTM(100, dropout=0.1))
model.add(Dense(nb_label_feature))
model.compile(loss=LOSS, optimizer=OPTIMIZER, metrics=['acc'])
return model
def define_autoencoder_model(number_of_features,nb_label_feature):
model = Sequential()
model.add(LSTM(100, input_shape=(window_length, number_of_features), return_sequences=True))
model.add(LSTM(50, return_sequences=False))
model.add(RepeatVector(window_length))
model.add(LSTM(100, dropout=0.1, return_sequences=True))
model.add(LSTM(50, return_sequences=True))
model.add(TimeDistributed(Dense(number_of_features)))
model.add(Flatten())
model.add(Dense(nb_label_feature))
model.compile(loss=LOSS, optimizer=OPTIMIZER, metrics=['acc'])
return model
#model = define_model(number_of_features,nb_label_feature)
#model3 = define_autoencoder_model(number_of_features,nb_label_feature)
#model4 = define_bidirectionnel_model(number_of_features,nb_label_feature)
#Moniteur pour stoper le training
es = EarlyStopping(monitor='acc', mode='max', verbose=1, patience=100)
# -
# Fonction de formatage des données en entrée du LSTM
def create_lstm_dataset(df, window_length,nb_label_feature):
number_of_rows = df.shape[0] #taille du dataset number_of_features
number_of_features = df.shape[1]
scaler = StandardScaler().fit(df.values)
transformed_dataset = scaler.transform(df.values)
transformed_df = pd.DataFrame(data=transformed_dataset, index=df.index)
#tableau de tableau de taille(number_of_rows-window_length) et window_length ligne,number_of_features
#lstm:[nb total de row ,nb de ligne dans le passé, nb de colonne(feature)]
train = np.empty([number_of_rows-window_length, window_length, number_of_features], dtype=float)
label = np.empty([number_of_rows-window_length, nb_label_feature], dtype=float)
for i in range(0, number_of_rows-window_length):
train[i] = transformed_df.iloc[i:i+window_length, 0: number_of_features]
label[i] = transformed_df.iloc[i+window_length: i+window_length+1, 0:nb_label_feature]
#définition du modèle Lstm
model = define_model(number_of_features,nb_label_feature)
return train, label, model,scaler
# ## Training
#formatage des données
train, label,model,scaler1 = create_lstm_dataset(df, window_length,nb_label_feature)
print(train.shape)
print(label.shape)
# * On voit ici que notre dataset d'entrainement après formatage est constitué de 1911 vecteurs contenant chacun 12 tirages où chaque tirage contient 19 features calculés plus haut
#
# * Quant aux labels, on a bien 1911 vecteurs de 6 features soit les 6 numéros de chaque tirages
#
# * Ainsi à partir des 12 tirages précédent on éssaie de prédire le tirage suivant lors de l'entrainement
#Training
history=model.fit(train, label, batch_size=BATCHSIZE, epochs=EPOCH, verbose=2, callbacks=[es])
# ## Fonction de perte
# +
#capture 6
# -
plt.plot(history.history['loss'])
plt.legend(['train_loss'])
plt.show()
# ## Prédiction du tirage suivant le dernier tirage de notre dataset de train
#Prediction basée sur les 12 derniers tirages
last_twelve = df.tail(window_length) # on recupere les 12 derniers tirages
scaler = StandardScaler().fit(df.values)
scaled_to_predict = scaler.transform(last_twelve)
scaled_predicted_output_1 = model.predict(np.array([scaled_to_predict]))
#prediction
tom = df.tail(window_length).iloc[:,0:6] #
scaler = StandardScaler().fit(df.iloc[:,0:6])
scaled_to_predict = scaler.transform(tom)
print(scaler.inverse_transform(scaled_predicted_output_1).astype(int)[0])
| DEEP_LEARNING_ET_LE_HASARD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.3
# language: julia
# name: julia-1.0
# ---
# # Cascade decay of a Ne 1s-hole: Simulate the final ion distribution
# # **This nootebook is still under construction !!!**
using JAC
#
# The computation of the cascade tree alone does not provide much information which could be compared with experiment. To facilitate the prediction of cascades or the comparison with experiment, the amplitudes from the various decay pathways need to be combined properly in order to predict different spectra and observables. Such observables are
#
# (1)
# (2)
#
# What does it include: Let us have a look at the definition of such a `Cascade.Simulation`:
#
? Cascade.Simulation
# What is seen ... and which information needed to be provided for such a computation
| tutorials/72-simulate-cascade-neon-k-hole.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Feature selection using SelectFromModel and LassoCV
#
#
# Use SelectFromModel meta-transformer along with Lasso to select the best
# couple of features from the Boston dataset.
#
#
# +
# Author: <NAME> <<EMAIL>>
# License: BSD 3 clause
print(__doc__)
import matplotlib.pyplot as plt
import numpy as np
from sklearn.datasets import load_boston
from sklearn.feature_selection import SelectFromModel
from sklearn.linear_model import LassoCV
# Load the boston dataset.
boston = load_boston()
X, y = boston['data'], boston['target']
# We use the base estimator LassoCV since the L1 norm promotes sparsity of features.
clf = LassoCV()
# Set a minimum threshold of 0.25
sfm = SelectFromModel(clf, threshold=0.25)
sfm.fit(X, y)
n_features = sfm.transform(X).shape[1]
# Reset the threshold till the number of features equals two.
# Note that the attribute can be set directly instead of repeatedly
# fitting the metatransformer.
while n_features > 2:
sfm.threshold += 0.1
X_transform = sfm.transform(X)
n_features = X_transform.shape[1]
# Plot the selected two features from X.
plt.title(
"Features selected from Boston using SelectFromModel with "
"threshold %0.3f." % sfm.threshold)
feature1 = X_transform[:, 0]
feature2 = X_transform[:, 1]
plt.plot(feature1, feature2, 'r.')
plt.xlabel("Feature number 1")
plt.ylabel("Feature number 2")
plt.ylim([np.min(feature2), np.max(feature2)])
plt.show()
| Assignments/hw3/Failed_to_perform_with_dataset/HW3_feature_selection_from_Boston/.ipynb_checkpoints/plot_select_from_model_boston-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Wrangling COVID-19 Mini-Challenge
# ## Imports and Setup
# +
import os
import pandas as pd
import seaborn as sns
import plotly.io as pio
import pandas_profiling
import plotly.express as px
import plotly.offline as pyo
import matplotlib.pyplot as plt
import plotly.graph_objects as go
from glob import glob
from datetime import date
# with this renderer, the plots are also shown in the html file.
pio.renderers.default = "jupyterlab"
# -
# ## International COVID-data
# +
today = date.today()
dates = [date.strftime('%m-%d-%Y') for date in pd.date_range(start='2020-01-22', end=today)][:-1]
dates_as_date = pd.date_range(start='2020-01-22', end=today)[:-1]
# standardize column names for all entries
def rename_columns(column):
column_map = {
'Lat': 'Latitude',
'Long_': 'Longitude',
'Incidence_Rate': 'Incident_Rate'
}
if column in column_map:
return column_map[column]
return column.replace('/', '_').replace('-', '_').replace(' ', '_')
all_cases = []
for d in range(len(dates)):
path = "https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/{}.csv".format(dates[d])
data = pd.read_csv(path)
data.rename(columns=rename_columns, inplace=True)
data['Date'] = dates_as_date[d]
all_cases.append(data)
df = pd.concat(all_cases)
# +
# all_cases.Country_Region.unique()
# standardize country names for all entries
country_mapping = {
'MS Zaandam|Diamond Princess|Cruise Ship': 'Others', # move cruise ships to others
'Hong Kong.+': 'Hong Kong',
'Iran.+': 'Iran',
'.*Congo.*': 'Congo',
'Mainland China': 'China',
'.*Bahamas.*': 'The Bahamas',
'.*Gambia.*': 'The Gambia',
'Viet Nam': 'Vietnam',
'Taiwan\*': 'Taiwan',
'Cote d\'Ivoire': 'Ivory Coast',
'Cabo Verde': 'Cape Verde',
'Russian Federation': 'Russia',
' Azerbaijan': 'Azerbaijan',
'Holy See': 'Vatican City',
'Republic of Ireland': 'Ireland',
'Republic of Moldova': 'Moldova',
'Czechia': 'Czech Republic',
'Republic of Korea|Korea, South': 'South Korea',
'Timor-Leste': 'East Timor',
'Macao SAR|Macau': 'Macao',
'UK': 'United Kingdom',
'Jersey|Guernsey': 'Channel Islands',
'Dominican Republicn Republic|Dominica': 'Dominican Republic'
}
df['Country_Region'].replace(
to_replace=country_mapping.keys(),
value=country_mapping.values(),
regex=True,
inplace=True
)
# -
df.to_csv('worldwide.csv')
df = pd.read_csv('worldwide.csv')
# group data by country
daily_updates = df.groupby(['Country_Region', 'Date']).agg(
Confirmed = ('Confirmed','sum'),
Deaths = ('Deaths','sum'),
).reset_index()
# get changes in data
updates_per_country = daily_updates.groupby('Country_Region')
# diff takes difference to point in group before it
daily_updates['New_Confirmed'] = updates_per_country['Confirmed'].diff().fillna(0)
daily_updates['New_Deaths'] = updates_per_country['Deaths'].diff().fillna(0)
# reorder columns
worldwide_pretty = daily_updates.loc[:, ['Date', 'Country_Region', 'New_Confirmed', 'Confirmed', 'New_Deaths', 'Deaths']]
# ## Plotting International Data
# create df with one column per date
worldwide_all = worldwide_pretty.groupby('Date').agg(
Confirmed = ('Confirmed', 'sum'),
New_Confirmed = ('New_Confirmed', 'sum')
)
# create a moving average
worldwide_all['SMA_14'] = round(worldwide_all.loc[:,'New_Confirmed'].rolling(window=14).mean())
worldwide_all = worldwide_all.reset_index()
# +
plot = go.Figure()
plot.add_trace(
go.Bar(
name='Daily Meas',
x=worldwide_all["Date"],
y=worldwide_all["New_Confirmed"],
marker={
# size': 10,
'color': 'red',
'opacity': 1
},
)
)
plot.add_trace(
go.Scatter(
name='14 Day MA',
x=worldwide_all["Date"],
y=worldwide_all["SMA_14"],
mode='lines',
marker={
'size': 1,
'color': 'black',
'opacity': 0.7
},
line_width=1.5
)
)
plot.update_layout(
title_x=0,
title='New Corona Cases Worldwide',
margin={
'l': 25,
'b': 25,
't': 50,
'r': 20
},
height=400,
yaxis={
'color': '#748B99',
'showgrid': True,
'showline': True,
'gridwidth': 1,
'gridcolor': '#B6C3CC',
# 'rangemode': "tozero",
'range':
[0,
worldwide_all.New_Confirmed.max() * 1.05],
# 'ticksuffix': ' '
# 'tickprefix': 'id: '
},
xaxis={
'showgrid': False,
'color': '#748B99',
'showline': True,
'linecolor': '#B6C3CC',
},
paper_bgcolor='#FFFFFF',
plot_bgcolor='rgba(0,0,0,0)',
showlegend=False,
legend={
'yanchor': 'top',
'y': 0.99,
'xanchor': 'right',
'x': 0.99
})
plot.show()
# -
# create df with one column per date
df = worldwide_pretty[worldwide_pretty.Country_Region == 'Ireland']
# create a moving average
df['SMA_14'] = round(df.loc[:,'New_Confirmed'].rolling(window=14).mean())
df = df.reset_index()
# +
plot = go.Figure()
plot.add_trace(
go.Bar(
name='Daily Meas',
x=df["Date"],
y=df["New_Confirmed"],
marker={
# size': 10,
'color': 'red',
'opacity': 1
},
)
)
plot.add_trace(
go.Scatter(
name='14 Day MA',
x=df["Date"],
y=df["SMA_14"],
mode='lines',
marker={
'size': 1,
'color': 'black',
'opacity': 0.7
},
line_width=1.5
)
)
plot.update_layout(
title_x=0,
title='New Corona Cases in Ireland',
margin={
'l': 25,
'b': 25,
't': 50,
'r': 20
},
height=400,
yaxis={
'color': '#748B99',
'showgrid': True,
'showline': True,
'gridwidth': 1,
'gridcolor': '#B6C3CC',
# 'rangemode': "tozero",
'range':
[0,
df.New_Confirmed.max() * 1.05],
# 'ticksuffix': ' '
# 'tickprefix': 'id: '
},
xaxis={
'showgrid': False,
'color': '#748B99',
'showline': True,
'linecolor': '#B6C3CC',
},
paper_bgcolor='#FFFFFF',
plot_bgcolor='rgba(0,0,0,0)',
showlegend=False,
legend={
'yanchor': 'top',
'y': 0.99,
'xanchor': 'right',
'x': 0.99
})
plot.show()
# -
# ## Swiss COVID-data
# data import
path = "https://raw.githubusercontent.com/openZH/covid_19/master/COVID19_Fallzahlen_CH_total_v2.csv"
chData = pd.read_csv(path)
# +
# only use data from 1 june on
chData = chData[chData.date > '2020-06-01']
# remove Fürstentum Lichtenstein
chData = chData[chData.abbreviation_canton_and_fl != 'FL']
# calculate new cases and deaths
chGrouped = chData.groupby('abbreviation_canton_and_fl')
chData['new_cases'] = chGrouped['ncumul_conf'].diff().fillna(0)
chData['new_deaths'] = chGrouped['ncumul_deceased'].diff().fillna(0)
# calculate total cases and deaths
chData['total_cases'] = chGrouped['new_cases'].cumsum()
chData['total_deaths'] = chGrouped['new_deaths'].cumsum()
# remove obsolete columns
chData = chData[[
'date',
'abbreviation_canton_and_fl',
'new_cases',
'total_cases',
'new_deaths',
'total_deaths'
]]
# rename columns
chData = chData.rename(
{
'abbreviation_canton_and_fl': 'canton'
},
axis='columns'
)
# -
# ## Plotting Swiss Data
# create df with one column per date
df = chData.groupby('date').agg(
Confirmed = ('total_cases', 'sum'),
New_Confirmed = ('new_cases', 'sum')
)
# create a moving average
df['SMA_14'] = round(df.loc[:,'New_Confirmed'].rolling(window=14).mean())
df = df.reset_index()
# +
plot = go.Figure()
plot.add_trace(
go.Bar(
name='Daily Meas',
x=df["date"],
y=df["New_Confirmed"],
marker={
# size': 10,
'color': 'red',
'opacity': 1
},
)
)
plot.add_trace(
go.Scatter(
name='14 Day MA',
x=df["date"],
y=df["SMA_14"],
mode='lines',
marker={
'size': 1,
'color': 'black',
'opacity': 0.7
},
line_width=1.5
)
)
plot.update_layout(
title_x=0,
title='New Corona Cases Switzerland',
margin={
'l': 25,
'b': 25,
't': 50,
'r': 20
},
height=400,
yaxis={
'color': '#748B99',
'showgrid': True,
'showline': True,
'gridwidth': 1,
'gridcolor': '#B6C3CC',
# 'rangemode': "tozero",
'range':
[0,
df.New_Confirmed.max() * 1.05],
# 'ticksuffix': ' '
# 'tickprefix': 'id: '
},
xaxis={
'showgrid': False,
'color': '#748B99',
'showline': True,
'linecolor': '#B6C3CC',
},
paper_bgcolor='#FFFFFF',
plot_bgcolor='rgba(0,0,0,0)',
showlegend=False,
legend={
'yanchor': 'top',
'y': 0.99,
'xanchor': 'right',
'x': 0.99
})
plot.show()
# -
# create df with one column per date
df = chData[chData.canton == 'BE']
# create a moving average
df['SMA_14'] = round(df.loc[:,'new_cases'].rolling(window=14).mean())
df = df.reset_index()
# +
plot = go.Figure()
plot.add_trace(
go.Bar(
name='Daily Meas',
x=df["date"],
y=df["new_cases"],
marker={
# size': 10,
'color': 'red',
'opacity': 1
},
)
)
plot.add_trace(
go.Scatter(
name='14 Day MA',
x=df["date"],
y=df["SMA_14"],
mode='lines',
marker={
'size': 1,
'color': 'black',
'opacity': 0.7
},
line_width=1.5
)
)
plot.update_layout(
title_x=0,
title='New Corona Cases Bern',
margin={
'l': 25,
'b': 25,
't': 50,
'r': 20
},
height=400,
yaxis={
'color': '#748B99',
'showgrid': True,
'showline': True,
'gridwidth': 1,
'gridcolor': '#B6C3CC',
# 'rangemode': "tozero",
'range':
[0,
df.new_cases.max() * 1.05],
# 'ticksuffix': ' '
# 'tickprefix': 'id: '
},
xaxis={
'showgrid': False,
'color': '#748B99',
'showline': True,
'linecolor': '#B6C3CC',
},
paper_bgcolor='#FFFFFF',
plot_bgcolor='rgba(0,0,0,0)',
showlegend=False,
legend={
'yanchor': 'top',
'y': 0.99,
'xanchor': 'right',
'x': 0.99
})
plot.show()
# -
# ## Dataframes
worldwide_pretty.head(3)
chData.head(3)
# ## Exports and HTML-file creation
# export the dataframes
chData.to_csv('covid_ch.csv')
worldwide_pretty.to_csv('covid_international.csv')
# create a html file
os.system('jupyter nbconvert --to html data_pipeline.ipynb')
| data_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc" style="margin-top: 1em;"><ul class="toc-item"></ul></div>
# -
# # RNN in TensorFlow Keras - TimeSeries Data <a class="tocSkip">
# +
import math
import os
import numpy as np
np.random.seed(123)
print("NumPy:{}".format(np.__version__))
import pandas as pd
print("Pandas:{}".format(pd.__version__))
import sklearn as sk
from sklearn import preprocessing as skpp
print("sklearn:{}".format(sk.__version__))
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams.update({'font.size': 20,
'figure.figsize': [15,10]
}
)
print("Matplotlib:{}".format(mpl.__version__))
import tensorflow as tf
tf.set_random_seed(123)
print("TensorFlow:{}".format(tf.__version__))
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM, SimpleRNN, GRU
from keras.losses import mean_squared_error as k_mse
from keras.backend import sqrt as k_sqrt
import keras.backend as K
import keras
print("Keras:{}".format(keras.__version__))
# +
DATASETSLIB_HOME = '../datasetslib'
import sys
if not DATASETSLIB_HOME in sys.path:
sys.path.append(DATASETSLIB_HOME)
# %reload_ext autoreload
# %autoreload 2
import datasetslib
from datasetslib import util as dsu
datasetslib.datasets_root = os.path.join(os.path.expanduser('~'),'datasets')
# -
# # Read and pre-process the dataset
filepath = os.path.join(datasetslib.datasets_root,
'ts-data',
'international-airline-passengers-cleaned.csv'
)
dataframe = pd.read_csv(filepath,
usecols=[1],
header=0)
dataset = dataframe.values
dataset = dataset.astype(np.float32)
# normalize the dataset
scaler = skpp.MinMaxScaler(feature_range=(0, 1))
normalized_dataset = scaler.fit_transform(dataset)
# split into train and test sets
train,test=dsu.train_test_split(normalized_dataset,train_size=0.67)
n_x=1
X_train, Y_train, X_test, Y_test = dsu.mvts_to_xy(train,test,n_x=n_x,n_y=1)
# # Keras SimpleRNN for TimeSeries Data
# +
tf.reset_default_graph()
keras.backend.clear_session()
# reshape input to be [samples, time steps, features]
X_train = X_train.reshape(X_train.shape[0], X_train.shape[1],1)
X_test = X_test.reshape(X_test.shape[0], X_train.shape[1], 1)
# -
# create and fit the SimpleRNN model
model = Sequential()
model.add(SimpleRNN(units=4, input_shape=(X_train.shape[1], X_train.shape[2])))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.summary()
model.fit(X_train, Y_train, epochs=20, batch_size=1)
# +
# make predictions
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
# invert predictions
y_train_pred = scaler.inverse_transform(y_train_pred)
y_test_pred = scaler.inverse_transform(y_test_pred)
#invert originals
y_train_orig = scaler.inverse_transform(Y_train)
y_test_orig = scaler.inverse_transform(Y_test)
# calculate root mean squared error
trainScore = k_sqrt(k_mse(y_train_orig[:,0],
y_train_pred[:,0])
).eval(session=K.get_session())
print('Train Score: {0:.2f} RMSE'.format(trainScore))
testScore = k_sqrt(k_mse(y_test_orig[:,0],
y_test_pred[:,0])
).eval(session=K.get_session())
print('Test Score: {0:.2f} RMSE'.format(testScore))
# -
# shift train predictions for plotting
trainPredictPlot = np.empty_like(normalized_dataset)
trainPredictPlot[:, :] = np.nan
trainPredictPlot[n_x:len(y_train_pred)+n_x, :] = y_train_pred
# shift test predictions for plotting
testPredictPlot = np.empty_like(normalized_dataset)
testPredictPlot[:, :] = np.nan
testPredictPlot[len(y_train_pred)+(n_x*2):len(normalized_dataset),:]=y_test_pred
# plot baseline and predictions
plt.plot(scaler.inverse_transform(normalized_dataset),label='Original Data')
plt.plot(trainPredictPlot,label='y_train_pred')
plt.plot(testPredictPlot,label='y_test_pred')
plt.legend()
plt.xlabel('Timesteps')
plt.ylabel('Total Passengers')
plt.show()
# # Keras LSTM for TimeSeries Data
# +
tf.reset_default_graph()
keras.backend.clear_session()
# reshape input to be [samples, time steps, features]
X_train = X_train.reshape(X_train.shape[0], X_train.shape[1],1)
X_test = X_test.reshape(X_test.shape[0], X_train.shape[1], 1)
# create and fit the LSTM model
model = Sequential()
model.add(LSTM(units=4, input_shape=(X_train.shape[1], X_train.shape[2])))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.summary()
model.fit(X_train, Y_train, epochs=20, batch_size=1)
# make predictions
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
# invert predictions
y_train_pred = scaler.inverse_transform(y_train_pred)
y_test_pred = scaler.inverse_transform(y_test_pred)
# invert originals
y_train_orig = scaler.inverse_transform(Y_train)
y_test_orig = scaler.inverse_transform(Y_test)
# calculate root mean squared error
trainScore = k_sqrt(k_mse(y_train_orig[:,0],
y_train_pred[:,0])
).eval(session=K.get_session())
print('Train Score: {0:.2f} RMSE'.format(trainScore))
testScore = k_sqrt(k_mse(y_test_orig[:,0],
y_test_pred[:,0])
).eval(session=K.get_session())
print('Test Score: {0:.2f} RMSE'.format(testScore))
# shift train predictions for plotting
trainPredictPlot = np.empty_like(normalized_dataset)
trainPredictPlot[:, :] = np.nan
trainPredictPlot[n_x:len(y_train_pred)+n_x, :] = y_train_pred
# shift test predictions for plotting
testPredictPlot = np.empty_like(normalized_dataset)
testPredictPlot[:, :] = np.nan
testPredictPlot[len(y_train_pred)+(n_x*2):len(normalized_dataset), :] = y_test_pred
# plot baseline and predictions
plt.plot(scaler.inverse_transform(normalized_dataset),label='Original Data')
plt.plot(trainPredictPlot,label='y_train_pred')
plt.plot(testPredictPlot,label='y_test_pred')
plt.legend()
plt.xlabel('Timesteps')
plt.ylabel('Total Passengers')
plt.show()
# -
# # Keras GRU for TimeSeries Data
# +
tf.reset_default_graph()
keras.backend.clear_session()
# reshape input to be [samples, time steps, features]
X_train = X_train.reshape(X_train.shape[0], X_train.shape[1],1)
X_test = X_test.reshape(X_test.shape[0], X_train.shape[1], 1)
# create and fit the GRU Model
model = Sequential()
model.add(GRU(units=4, input_shape=(X_train.shape[1], X_train.shape[2])))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.summary()
model.fit(X_train, Y_train, epochs=20, batch_size=1)
# make predictions
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
# invert predictions
y_train_pred = scaler.inverse_transform(y_train_pred)
y_test_pred = scaler.inverse_transform(y_test_pred)
#invert originals
y_train_orig = scaler.inverse_transform(Y_train)
y_test_orig = scaler.inverse_transform(Y_test)
# calculate root mean squared error
trainScore = k_sqrt(k_mse(y_train_orig[:,0],
y_train_pred[:,0])
).eval(session=K.get_session())
print('Train Score: {0:.2f} RMSE'.format(trainScore))
testScore = k_sqrt(k_mse(y_test_orig[:,0],
y_test_pred[:,0])
).eval(session=K.get_session())
print('Test Score: {0:.2f} RMSE'.format(testScore))
# shift train predictions for plotting
trainPredictPlot = np.empty_like(normalized_dataset)
trainPredictPlot[:, :] = np.nan
trainPredictPlot[n_x:len(y_train_pred)+n_x, :] = y_train_pred
# shift test predictions for plotting
testPredictPlot = np.empty_like(normalized_dataset)
testPredictPlot[:, :] = np.nan
testPredictPlot[len(y_train_pred)+(n_x*2):len(normalized_dataset), :] = y_test_pred
# plot baseline and predictions
plt.plot(scaler.inverse_transform(normalized_dataset),label='Original Data')
plt.plot(trainPredictPlot,label='y_train_pred')
plt.plot(testPredictPlot,label='y_test_pred')
plt.legend()
plt.xlabel('Timesteps')
plt.ylabel('Total Passengers')
plt.show()
| Chapter07/ch-07b_RNN_TimeSeries_Keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="tmaHAc_KHG_5"
# # Feed-forward nets for image classification
# + [markdown] colab_type="text" id="zgNIWhzoHOIz"
# ## 1. The MNIST dataset
# + colab={} colab_type="code" id="dQiqJyO7E7Ek"
# #!pip install mnist
import mnist
train_imgs = mnist.train_images()
train_labels = mnist.train_labels()
test_imgs = mnist.test_images()
test_labels = mnist.test_labels()
# -
# ### Data standardization
#
# Rescale input values to have zero mean and standard deviation of one.
mean, std = train_imgs.mean(), train_imgs.std()
train_imgs = (train_imgs - mean) / std
test_imgs = (test_imgs - mean) / std
# ### See some inputs
# + colab={"base_uri": "https://localhost:8080/", "height": 99} colab_type="code" id="qeftJ_CpE7Eu" outputId="c7a9a956-0114-4975-90a0-8af5b2007c2b"
from typing import List
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 99} colab_type="code" id="qeftJ_CpE7Eu" outputId="c7a9a956-0114-4975-90a0-8af5b2007c2b"
idxs = np.random.randint(0, len(train_imgs), 15)
imgs = np.concatenate(tuple(train_imgs[idx,:,:] for idx in idxs), axis=1)
plt.imshow(imgs)
print("Labels:", train_labels[idxs])
# + [markdown] colab_type="text" id="ZgERUA07IuSr"
# ## 2. Building feed forward-networks
# -
# ### Layers
#
# Each layer will have three methods:
# - `forward` computes and returns ${\bf y}^{(l)} = f_l\left({\bf x}^{(l)}, {\bf w}^{(l)}\right)$
# - `backward` gets $\frac{\partial {\cal L}}{\partial {\bf y}^{(l)}}$, and stores $\frac{\partial {\cal L}}{\partial {\bf w}^{(l)}}$ internally, and returns $\frac{\partial {\cal L}}{\partial {\bf w}^{(l)}}$
# - `update` modifies parameters ${\bf w}^{(l)}$ using stored $\frac{\partial{\cal L}}{\partial{\bf w}}$
class Layer:
def forward(self, x: np.ndarray) -> np.ndarray:
raise NotImplementedError
def backward(self, x: np.ndarray, dy: np.ndarray) -> np.ndarray:
raise NotImplementedError
def update(self, *args, **kwargs):
pass # If a layer has no parameters, then this function does nothing
# ### The feed-forward netowork
class FeedForwardNetwork:
def __init__(self, layers: List[Layer]):
self.layers = layers
def forward(self, x: np.ndarray, train: bool = True) -> np.ndarray:
self._inputs = []
for layer in self.layers:
if train:
self._inputs.append(x)
x = layer.forward(x)
return x
def backward(self, dy:np.ndarray) -> np.ndarray:
# TODO <0> : Compute the backward phase
raise NotImplementedError
del self._inputs
def update(self, *args, **kwargs):
for layer in self.layers:
layer.update(*args, **kwargs)
# ### The linear layer
# + colab={} colab_type="code" id="S47ZsyKdE7FF"
class Linear(Layer):
def __init__(self, insize: int, outsize: int) -> None:
bound = np.sqrt(6. / insize)
self.weight = np.random.uniform(-bound, bound, (insize, outsize))
self.bias = np.zeros((outsize,))
self.dweight = np.zeros_like(self.weight)
self.dbias = np.zeros_like(self.bias)
def forward(self, x: np.ndarray) -> np.ndarray:
# TODO <1> : compute the output of a linear layer
raise NotImplementedError
def backward(self, x: np.ndarray, dy: np.ndarray) -> np.ndarray:
# TODO <2> : compute dweight, dbias and return dx
raise NotImplementedError
def update(self, mode='SGD', lr=0.001, mu=.9):
if mode == 'SGD':
self.weight -= lr * self.dweight
self.bias -= lr * self.dbias
elif mode == 'Nesterov':
# TODO <9> : compute the nesterov update (for Lab 2)
raise NotImplementedError
elif mode == 'Adam':
# TODO <10> : compute the Adam update (for Lab 2)
raise NotImplementedError
# -
# ### The Rectified Linear Unit
# $$y = \max\left(x, 0\right)$$
# + colab={} colab_type="code" id="QOR1DJiwE7FJ"
class ReLU(Layer):
def __init__(self) -> None:
pass
def forward(self, x: np.ndarray) -> np.ndarray:
# TODO <3> : Compute the output of a rectified linear unit
raise NotImplementedError
def backward(self, x: np.ndarray, dy: np.ndarray) -> np.ndarray:
# TODO <4> : Compute the gradient w.r.t. x
raise NotImplementedError
# + [markdown] colab_type="text" id="4NrWBTmbI9gW"
# ## 3. The loss function
#
# The negative log likelihood combines a softmax activation, and a cross-entropy cost.
# + colab={} colab_type="code" id="YDXiDEu8E7FW"
class NegativeLogLikelihood:
def __init__(self):
pass
def forward(self, y: np.ndarray, t: np.ndarray) -> float:
# TODO <5> : Compute the negative log likelihood
raise NotImplementedError
def backward(self, y: np.ndarray, t: np.ndarray) -> np.ndarray:
# TODO <6> : Compute dl/dy
raise NotImplementedError
# + [markdown] colab_type="text" id="Uz9qM5eHJLNw"
# ### Accuracy
# + colab={} colab_type="code" id="3nYfVCBSE7Fe"
def accuracy(y: np.ndarray, t: np.ndarray) -> float:
# TODO <7> : Compute accuracy
raise NotImplementedError
# + [markdown] colab_type="text" id="mIhtzd2gJQF2"
# ## 4. Training a neural network
# + colab={"base_uri": "https://localhost:8080/", "height": 237} colab_type="code" id="HTbmZv3YE7Fs" outputId="d6bb5b23-201b-4c2f-cf88-37f90bb5f6a6"
BATCH_SIZE = 128
HIDDEN_UNITS = 200
EPOCHS_NO = 50
optimize_args = {'mode': 'SGD', 'lr': .001}
net = FeedForwardNetwork([Linear(784, HIDDEN_UNITS),
ReLU(),
Linear(HIDDEN_UNITS, 10)])
nll = NegativeLogLikelihood()
for epoch in range(EPOCHS_NO):
for b_no, idx in enumerate(range(0, len(train_imgs), BATCH_SIZE)):
# 1. Prepare next batch
x = train_imgs[idx:idx + BATCH_SIZE,:,:].reshape(-1, 784)
t = train_labels[idx:idx + BATCH_SIZE]
# 2. Compute gradient
# TODO <8> : Compute gradient
raise NotImplementedError
# 3. Update network parameters
net.update(**optimize_args)
print(f'\rEpoch {epoch + 1:02d} '
f'| Batch {b_no:03d} '
f'| Train NLL: {loss:3.5f} '
f'| Train Accuracy: {accuracy(y, t):3.2f} ', end='')
y = net.forward(test_imgs.reshape(-1, 784), train=False)
test_nll = nll.forward(y, test_labels)
print(f' | Test NLL: {test_nll:3.5f} '
f' | Test Accuracy: {accuracy(y, test_labels):3.2f}')
# -
| labs/lab01/Feed-forward nets for image classification - Skel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="sAtovljrSzv8"
# # 데이터 준비
# + colab={"base_uri": "https://localhost:8080/"} id="Burr1ejRJr_Q" executionInfo={"status": "ok", "timestamp": 1647835633702, "user_tz": -540, "elapsed": 1899, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>xGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}} outputId="2282665b-ca3b-4ae0-c989-3c7268f277a5"
import os
import random
import numpy as np
import nltk
nltk.download("punkt")
from nltk.tokenize import word_tokenize
import torch
from torchtext.legacy.data import Field
from torchtext.legacy.data import TabularDataset
from torchtext.legacy.data import BucketIterator
from torchtext.legacy.data import Iterator
# + id="pbBR3MuyF7VY" executionInfo={"status": "ok", "timestamp": 1647835633703, "user_tz": -540, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
# Nondeterministic 한 작업 피하기
RANDOM_SEED = 2022
random.seed(RANDOM_SEED)
np.random.seed(RANDOM_SEED)
torch.manual_seed(RANDOM_SEED)
torch.backends.cudnn.deterministic = True # Deterministic 한 알고리즘만 사용하기
torch.backends.cudnn.benchmark = False # Cudnn benchmark 해제
torch.cuda.manual_seed_all(RANDOM_SEED) # if use multi-GPU
os.environ['PYTHONHASHSEED'] = str(RANDOM_SEED)
# + id="xjqPrRVSkzmt" executionInfo={"status": "ok", "timestamp": 1647835633703, "user_tz": -540, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
DATA_PATH = "/content/drive/Othercomputers/내 컴퓨터/Sat_english/data/processed"
# + [markdown] id="QsXwSJdPX5iN"
# 필드 정의
# + id="lfxKZjPrTBOz" executionInfo={"status": "ok", "timestamp": 1647835633704, "user_tz": -540, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
# 문장 필드
TEXT = Field(
sequential=True, # 문장 입력
use_vocab=True,
tokenize=word_tokenize, # nltk의 word_tokenize로 트큰화
lower=True, # 모두 소문자 처리
batch_first=True,
)
# 정답 필드
LABEL = Field(
sequential=False,
use_vocab=False,
batch_first=True,
)
# + [markdown] id="LValW3OmX-31"
# 데이터 불러오기
# + id="ggRQO7KgT4Bk" executionInfo={"status": "ok", "timestamp": 1647835636079, "user_tz": -540, "elapsed": 2379, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
# CoLA 데이터 = 사전 학습 데이터
cola_train_data, cola_valid_data, cola_test_data = TabularDataset.splits(
path=DATA_PATH,
train="cola_train.tsv",
validation="cola_valid.tsv",
test="cola_test.tsv",
format="tsv",
fields=[("text", TEXT), ("label", LABEL)],
skip_header=1, # column명이 있는 1열 생략
)
TEXT.build_vocab(cola_train_data, min_freq=2) # CoLA 데이터로 사전학습할 단어장 생성(2번 이상 나온 단어만)
# 수능 데이터 = 추가 학습 데이터
sat_train_data, sat_valid_data, sat_test_data = TabularDataset.splits(
path=DATA_PATH,
train="sat_train.tsv",
validation="sat_valid.tsv",
test="sat_test.tsv",
format="tsv",
fields=[("text", TEXT), ("label", LABEL)],
skip_header=1,
)
# + [markdown] id="4CdaR71bYBRP"
# DataLoader 정의
# + id="SEY6mJmCX1Fj" executionInfo={"status": "ok", "timestamp": 1647835636080, "user_tz": -540, "elapsed": 13, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>3=s64", "userId": "11886396836022920274"}}
# CoLA 데이터
cola_train_iterator, cola_valid_iterator, cola_test_iterator = BucketIterator.splits(
(cola_train_data, cola_valid_data, cola_test_data),
batch_size=32,
device=None,
sort=False,
)
# 수능 데이터
sat_train_iterator, sat_valid_iterator, sat_test_iterator = BucketIterator.splits(
(sat_train_data, sat_valid_data, sat_test_data),
batch_size=8,
device=None,
sort=False,
)
# + [markdown] id="niRzsFTqXpF-"
# # 네트워크 구성
# + id="723vR_q6X7o8" executionInfo={"status": "ok", "timestamp": 1647835636081, "user_tz": -540, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
import torch
import torch.nn as nn
# + id="iVAvvIEbX0IX" executionInfo={"status": "ok", "timestamp": 1647835636082, "user_tz": -540, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
class LSTM_Model(nn.Module):
def __init__(self, num_embeddings, embedding_dim, hidden_size, num_layers, pad_idx):
super().__init__()
# Embedding Layer
self.embed_layer = nn.Embedding(
num_embeddings=num_embeddings,
embedding_dim=embedding_dim,
padding_idx=pad_idx
)
# LSTM Layer
self.lstm_layer = nn.LSTM(
input_size=embedding_dim,
hidden_size=hidden_size,
num_layers=num_layers,
batch_first = True,
bidirectional=True, # 양방향 LSTM
dropout=0.5
)
# Fully-connetcted Layer
self.fc_layer1 = nn.Sequential(
nn.Linear(hidden_size * 2, hidden_size), # 양방향 LSTM의 출력은 입력의 2배
nn.Dropout(0.5),
nn.LeakyReLU() # f(x)=max(0.01x, x)로 dying ReLU 방지
)
self.fc_layer2 = nn.Sequential(
nn.Linear(hidden_size, 1)
)
def forward(self, x):
embed_x = self.embed_layer(x)
output, (_, _) = self.lstm_layer(embed_x) # hidden, cell state의 출력값 사용 안함
output = output[:, -1, :] # (batch_size, seq_length, 2*hidden_size) -> (batch_size, 2*hidden_size)
output = self.fc_layer1(output)
output = self.fc_layer2(output)
return output
# + [markdown] id="_GvTG58LuI9g"
# # 모델 학습 및 검증
# + [markdown] id="meeBGEVqwrVy"
# 파라미터 정의
# + id="Mqy27QLCutiw" executionInfo={"status": "ok", "timestamp": 1647835638989, "user_tz": -540, "elapsed": 2918, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
USE_CUDA = torch.cuda.is_available()
DEVICE = torch.device("cuda" if USE_CUDA else "cpu") # GPU 존재시 GPU 실행(CUDA)
PAD_IDX = TEXT.vocab.stoi[TEXT.pad_token] # 동일한 크기를 맞추기 위한 패딩문자를 숫자 식별자에 매칭 -> 숫자 식별자=index
lstm = LSTM_Model(
num_embeddings=len(TEXT.vocab),
embedding_dim=100,
hidden_size=200,
num_layers=4,
pad_idx=PAD_IDX
).to(DEVICE)
n_epochs = 20
learning_rate = 0.001
optimizer = torch.optim.Adam(lstm.parameters(), lr=learning_rate)
criterion = nn.BCEWithLogitsLoss() # Sigmoid + BCELoss
# + [markdown] id="jsONag9Zw1Xm"
# 훈련 데이터로 학습하여 모델화
# + id="RmCedQivuJNd" executionInfo={"status": "ok", "timestamp": 1647835638991, "user_tz": -540, "elapsed": 19, "user": {"displayName": "CaFe CoKe", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
def train(model, iterator, optimizer):
train_loss = 0
model.train() # 모델을 train모드로 설정(Dropout 적용)
for _, batch in enumerate(iterator):
optimizer.zero_grad() # optimizer 초기화(Gradient)
text = batch.text # 해당 Batch의 text 속성 불러오기
label = batch.label.type(torch.FloatTensor) # 해당 Batch의 label 속성 불러오기(32-bit float)
text = text.to(DEVICE)
label = label.to(DEVICE)
output = model(text).flatten() # output은 [batch_size, 1], label은 [batch_size]
loss = criterion(output, label)
loss.backward() # 역전파로 Gradient를 계산 후 파라미터에 할당
optimizer.step() # 파라미터 업데이트
train_loss += loss.item() # Loss 값 누적
# Loss 값을 Batch 값으로 나누어 미니 배치마다의 Loss 값의 평균을 구함
return train_loss/len(iterator)
# + [markdown] id="KITUxPSfuLjc"
# 모델 검증
# + id="rnvLNdwKuLyV" executionInfo={"status": "ok", "timestamp": 1647835638992, "user_tz": -540, "elapsed": 18, "user": {"displayName": "CaFe CoKe", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
def evaluate(model, iterator):
valid_loss = 0
model.eval() # 모델을 eval모드로 설정(Dropout 미적용)
with torch.no_grad(): # Gradient 계산 비활성화 (모델 평가에는 파라미터 업데이트 X)
for _, batch in enumerate(iterator):
text = batch.text
label = batch.label.type(torch.FloatTensor)
text = text.to(DEVICE)
label = label.to(DEVICE)
output = model(text).flatten()
loss = criterion(output, label)
valid_loss += loss.item()
return valid_loss/len(iterator)
# + [markdown] id="5wcFrq1uuFF_"
# CoLA 데이터 사전학습
# + id="_jdht0nnvx68" executionInfo={"status": "ok", "timestamp": 1647835638994, "user_tz": -540, "elapsed": 19, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
import time
# + id="pmwwn-YAwN8-" executionInfo={"status": "ok", "timestamp": 1647835638996, "user_tz": -540, "elapsed": 19, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
def epoch_time(start_time: int, end_time: int): # epoch 시간
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# + id="LwuMQY_pBY8s" executionInfo={"status": "ok", "timestamp": 1647835638997, "user_tz": -540, "elapsed": 19, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
loss_tr = []
loss_val = []
# + colab={"base_uri": "https://localhost:8080/"} id="2HsbUZ8cuIKZ" executionInfo={"status": "ok", "timestamp": 1647835723945, "user_tz": -540, "elapsed": 84966, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}} outputId="4807aaf4-7fb8-458e-af74-831ba8dbda4c"
for epoch in range(n_epochs):
start_time = time.time()
train_loss = train(lstm, cola_train_iterator, optimizer)
valid_loss = evaluate(lstm, cola_valid_iterator)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
print(f"Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s")
print(f"Train Loss: {train_loss:.5f}")
print(f" Val. Loss: {valid_loss:.5f}")
print('----------------------------------')
# overfitting 확인하기 위함
loss_tr.append(train_loss)
loss_val.append(valid_loss)
# + id="_8NlSZzrBsvS" executionInfo={"status": "ok", "timestamp": 1647835723946, "user_tz": -540, "elapsed": 33, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
import numpy as np
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 606} id="29atcZgNBo0p" executionInfo={"status": "ok", "timestamp": 1647835723947, "user_tz": -540, "elapsed": 32, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}} outputId="dce74ace-dcbc-4127-9b71-20f1c9e67e5f"
np1 = np.array(loss_tr)
np2 = np.array(loss_val)
plt.figure(figsize=(10, 10))
plt.xlabel('epoch')
plt.ylabel('loss')
plt.plot(np1, label='Loss of train')
plt.plot(np2, label='Loss of Validation')
plt.legend() # 라벨표시를 위한 범례
plt.show()
# + id="WAZEYOczWPK2" executionInfo={"status": "ok", "timestamp": 1647835723947, "user_tz": -540, "elapsed": 27, "user": {"displayName": "CaFe CoKe", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
from copy import deepcopy
# 사전학습 모델
before_tuning_lstm = deepcopy(lstm)
# + [markdown] id="NcygpOpUWq3J"
# 수능 데이터를 이용해 추가 학습 (Fine-Tune)
# + colab={"base_uri": "https://localhost:8080/"} id="OJVoGAfWWtJ2" executionInfo={"status": "ok", "timestamp": 1647835729668, "user_tz": -540, "elapsed": 5747, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}} outputId="fc0af1b1-883a-4506-dd05-108e7f7ad7dc"
loss_tr_tune = []
loss_val_tune = []
for epoch in range(n_epochs):
start_time = time.time()
train_loss = train(lstm, sat_train_iterator, optimizer)
valid_loss = evaluate(lstm, sat_valid_iterator)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
print(f"Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s")
print(f"\tTrain Loss: {train_loss:.5f}")
print(f"\t Val. Loss: {valid_loss:.5f}")
print('----------------------------------')
# overfitting 확인하기 위함
loss_tr_tune.append(train_loss)
loss_val_tune.append(valid_loss)
# + colab={"base_uri": "https://localhost:8080/", "height": 606} id="uxVTYJIHXejW" executionInfo={"status": "ok", "timestamp": 1647835729669, "user_tz": -540, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}} outputId="70ee1895-2c8f-4813-dfd5-2f331bd81af8"
np1 = np.array(loss_tr_tune)
np2 = np.array(loss_val_tune)
plt.figure(figsize=(10, 10))
plt.xlabel('epoch')
plt.ylabel('loss')
plt.plot(np1, label='Loss of train')
plt.plot(np2, label='Loss of Validation')
plt.legend() # 라벨표시를 위한 범례
plt.show()
# + [markdown] id="yG11HjHAgioo"
# # 모델 성능 테스트
# + id="9Kv_UUurgrWI" executionInfo={"status": "ok", "timestamp": 1647835729670, "user_tz": -540, "elapsed": 22, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
import dill
from sklearn.metrics import roc_curve, auc
# + id="ythUmHwBglMB" executionInfo={"status": "ok", "timestamp": 1647835730287, "user_tz": -540, "elapsed": 638, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
def test(model, iterator, device):
model.eval()
with torch.no_grad():
y_real = []
y_pred = []
for batch in iterator:
text = batch.text
label = batch.label.type(torch.FloatTensor)
text = text.to(device)
output = model(text).flatten().cpu() # roc_curve의 입력 형태는 ndarray의 형태
# 각 Batch의 예측값 list화
y_real += [label]
y_pred += [output]
y_real = torch.cat(y_real)
y_pred = torch.cat(y_pred)
fpr, tpr, _ = roc_curve(y_real, y_pred)
auroc = auc(fpr, tpr)
return auroc
# + colab={"base_uri": "https://localhost:8080/"} id="HSV_DAbGhLfq" executionInfo={"status": "ok", "timestamp": 1647835730287, "user_tz": -540, "elapsed": 12, "user": {"displayName": "CaFe CoKe", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}} outputId="ca495ebe-87b1-4817-89f4-3ffcad4103f6"
_ = before_tuning_lstm.cpu()
lstm_sat_test_auroc = test(before_tuning_lstm, sat_test_iterator, "cpu")
_ = lstm.cpu()
lstm_tuned_test_auroc = test(lstm, sat_test_iterator, "cpu")
print(f"Before fine-tuning SAT Dataset Test AUROC: {lstm_sat_test_auroc:.5f}")
print(f"After fine-tuning SAT Dataset Test AUROC: {lstm_tuned_test_auroc:.5f}")
# + id="pidf_OmGdOWP" executionInfo={"status": "ok", "timestamp": 1647835730287, "user_tz": -540, "elapsed": 9, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgKQbV_6mqL_VHCcbuNL4GHNUxGOJQBtuNsOFo3=s64", "userId": "11886396836022920274"}}
with open("before_tuning_model.dill", "wb") as f:
model = {
"TEXT": TEXT,
"LABEL": LABEL,
"classifier": before_tuning_lstm
}
dill.dump(model, f)
_ = lstm.cpu()
with open("after_tuning_model.dill", "wb") as f:
model = {
"TEXT": TEXT,
"LABEL": LABEL,
"classifier": lstm
}
dill.dump(model, f)
| Sat_pre_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import math
import matplotlib.pyplot as plt
import matplotlib
import seaborn as sns
from datetime import datetime, date, time
from scipy.interpolate import interp1d
import warnings
warnings.filterwarnings('ignore')
# # 1. LOADING THE PLOTS INFORMATION
# ## 1.1.1 AERODYNAMIC PROPERTIES
# +
#Defining the location of the data folder
DATA_FOLDER = 'Desktop/PLOTS_DATA_WIND_TURBINES/Aerodynamics_properties_blades/Data/'
#DU21-A17
#Definition of the data path
DATASET1 = DATA_FOLDER + "Drag DU21-A17_pi.csv"
DATASET2 = DATA_FOLDER + "Lift DU21-A17_pi.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_drag_DU21_A17 = pd.read_csv(DATASET1,sep =';',decimal=",")
df_lift_DU21_A17 = pd.read_csv(DATASET2,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_drag_DU21_A17.columns=["Angle_of_attack", "Cd"]
df_lift_DU21_A17.columns=["Angle_of_attack", "Cl"]
#DU25-A17
#Definition of the data path
DATASET1 = DATA_FOLDER + "Drag DU25-A17_pi.csv"
DATASET2 = DATA_FOLDER + "Lift DU25-A17_pi.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_drag_DU25_A17 = pd.read_csv(DATASET1,sep =';',decimal=",")
df_lift_DU25_A17 = pd.read_csv(DATASET2,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_drag_DU25_A17.columns=["Angle_of_attack", "Cd"]
df_lift_DU25_A17.columns=["Angle_of_attack", "Cl"]
#DU30-A17
#Definition of the data path
DATASET1 = DATA_FOLDER + "Drag DU30-A17_pi.csv"
DATASET2 = DATA_FOLDER + "Lift DU30-A17_pi.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_drag_DU30_A17 = pd.read_csv(DATASET1,sep =';',decimal=",")
df_lift_DU30_A17 = pd.read_csv(DATASET2,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_drag_DU30_A17.columns=["Angle_of_attack", "Cd"]
df_lift_DU30_A17.columns=["Angle_of_attack", "Cl"]
#DU35-A17
#Definition of the data path
DATASET1 = DATA_FOLDER + "Drag DU35-A17_pi.csv"
DATASET2 = DATA_FOLDER + "Lift DU35-A17_pi.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_drag_DU35_A17 = pd.read_csv(DATASET1,sep =';',decimal=",")
df_lift_DU35_A17 = pd.read_csv(DATASET2,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_drag_DU35_A17.columns=["Angle_of_attack", "Cd"]
df_lift_DU35_A17.columns=["Angle_of_attack", "Cl"]
#DU40-A17
#Definition of the data path
DATASET1 = DATA_FOLDER + "Drag DU40-A17_pi.csv"
DATASET2 = DATA_FOLDER + "Lift DU40-A17_pi.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_drag_DU40_A17 = pd.read_csv(DATASET1,sep =';',decimal=",")
df_lift_DU40_A17 = pd.read_csv(DATASET2,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_drag_DU40_A17.columns=["Angle_of_attack", "Cd"]
df_lift_DU40_A17.columns=["Angle_of_attack", "Cl"]
#NACA64-A17
#Definition of the data path
DATASET1 = DATA_FOLDER + "Drag NACA64-A17_pi.csv"
DATASET2 = DATA_FOLDER + "Lift NACA64-A17_pi.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_drag_NACA64_A17 = pd.read_csv(DATASET1,sep =';',decimal=",")
df_lift_NACA64_A17 = pd.read_csv(DATASET2,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_drag_NACA64_A17.columns=["Angle_of_attack", "Cd"]
df_lift_NACA64_A17.columns=["Angle_of_attack", "Cl"]
# -
# ## 1.1.2 AERODYNAMIC FORCES
# +
fx = np.load('fx.npy')
ft = np.load('ft.npy')
# t_total=800s
#500 time steps
#3 different blade turbines
#64 points in the aerodynamic calculation spanwise direction for 62.5 m
#1 turbine
# -
fx.shape
ft.shape
# Now the objective is extract the useful data only from one blade to reduce the multidimensional array. It will be more intuitively for the following steps have a less dimensions in the aerodynamic force applied in the blade.
fx_2D = fx[:,1,:,0]
ft_2D = ft[:,1,:,0]
# It is necessary translate the data from the 64 points in the span-wise direction into his correspondance radius position to be able to interpolate. The best way is create a data frame with value of aerodynamic force and the blade position. The main reason is because if the number of discretization points change in the structural program, be capable to interpolate and get a correct value.
#Creating set of data about the radius/blade position from aerodynamic force simulation.
N_aero = fx.shape[2]
radius_position_aero = np.zeros(N_aero)
R_aero = 63
delta_r_aero = R_aero/(N_aero-1)
r_aero = 0
for i in range(1,N_aero):
r_aero = r_aero + delta_r_aero
radius_position_aero[i] = r_aero
radius_position_aero
#Creating set of data about the time position from aerodynamic force simulation.
T_aero = fx.shape[0]
time_position_aero = np.zeros(T_aero)
total_t_aero = 800 #seconds
delta_t_aero = total_t_aero/(T_aero-1)
t_aero = 0
for i in range(1,T_aero):
t_aero = t_aero + delta_t_aero
time_position_aero[i] = t_aero
time_position_aero
#Creating dataframes to store the data from the aerodynamic simulation:
columns = ["time", "position"]
index_position = np.linspace(0, 499, num=500)
data_time = np.dstack((time_position_aero,index_position))
data_time = data_time[0,:,:]
df_time_aero = pd.DataFrame(data=data_time, columns=columns)
df_time_aero
# +
#Creation of the interpolation function for a determinated dataframe:
def interpolation(r,df):
xp = df.iloc[:,0]
fp = df.iloc[:,1]
interpolation = np.interp(r,xp,fp)
return interpolation
# -
#Create the function to find index postion of aerodynamic force stored:
def find_aero_index(t,df_time_aero):
time_index = interpolation(t,df_time_aero)
trunc_time_index = np.trunc(time_index)
trunc_time_index = int(trunc_time_index)
return time_index,trunc_time_index
#Creation of the interpolation function for a determinated time_step aerodynamic simulation:
def interpolation_time(time_index,trunc_time_index,force_2D):
fst = force_2D[trunc_time_index,:]
snd = force_2D[trunc_time_index+1,:]
linfit = interp1d([trunc_time_index,trunc_time_index+1], np.vstack([fst, snd]), axis=0)
return linfit(time_index)
F = interpolation_time(find_aero_index(3,df_time_aero)[0],find_aero_index(3,df_time_aero)[1],fx_2D)
F
#Creation a function that creates a dataframe with the time_step aerodynamic forces at each r:
def dataframe_creation(radius_position_aero,aero_force_dataset):
columns = ["spanwise_length", "Aerodynamic_force"]
data = np.dstack((radius_position_aero,aero_force_dataset))
data = data[0,:,:]
df_force = pd.DataFrame(data=data, columns=columns)
return df_force
H = dataframe_creation(radius_position_aero,F)
H
# ## 1.2 STRUCTURAL PROPERTIES
# +
#Defining the location of the data folder
DATA_FOLDER = 'Desktop/PLOTS_DATA_WIND_TURBINES/Structural_properties/Data/'
#BLADE DENSITY
#Definition of the data path
DATASET = DATA_FOLDER + "Blade_Density.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_blade_density = pd.read_csv(DATASET,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_blade_density.columns=["spanwise_length", "blade_density"]
#STRUCTURAL TWIST
#Definition of the data path
DATASET = DATA_FOLDER + "Structural_Twist.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_twist = pd.read_csv(DATASET,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_twist.columns=["spanwise_length", "twist"]
#FLAPWISE STIFFNESS
#Definition of the data path
DATASET = DATA_FOLDER + "Flapwise_stiffness.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_flapwise_stiffness = pd.read_csv(DATASET,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_flapwise_stiffness.columns=["spanwise_length", "flapwise_stiffness"]
#EDGEWISE STIFFNESS
#Definition of the data path
DATASET = DATA_FOLDER + "Edgewise_Stiffness.csv"
#Load of the compressed files in a specific way in function of the type of data file
df_edgewise_stiffness = pd.read_csv(DATASET,sep =';',decimal=",")
#Defining the columns names of the uploaded dataframe
df_edgewise_stiffness.columns=["spanwise_length", "edgewise_stiffness"]
# -
# # 2. DEFINING VARIABLES AND ARRAYS TO STORE DATA
# +
#Creating the 6 arrays neededs to store the infromation in each time step.
T = 16000 #Number of points that time is discretized
N = 60 #Number of points that the airfoil is discretized
#Dynamic velocity arrays:
V_0_new = np.zeros(N+2)
V_1_new = np.zeros(N+2)
V_0_old = np.zeros(N+2)
V_1_old = np.zeros(N+2)
#Moments arrays:
M_0_new = np.zeros(N+2)
M_1_new = np.zeros(N+2)
M_0_old = np.zeros(N+2)
M_1_old = np.zeros(N+2)
#Centrifual term arrays:
S_0_new = np.zeros(N+2)
S_1_new = np.zeros(N+2)
S_0_old = np.zeros(N+2)
S_1_old = np.zeros(N+2)
#Dynamic displacement arrays:
q_0_new = np.zeros(N+2)
q_1_new = np.zeros(N+2)
q_0_old = np.zeros(N+2)
q_1_old = np.zeros(N+2)
#Centrifugal force array:
N_force = np.zeros(N+1)
#Radial position in the blade array:
Position = np.zeros(N+2)
#Time position in the simulation array:
Time = np.zeros(T+1)
#Root bending moments array:
Root_M_0 = np.zeros(T+1)
Root_M_1 = np.zeros(T+1)
#Tip velocity of the displacements array:
Tip_V_0 = np.zeros(T+1)
Tip_V_1 = np.zeros(T+1)
#Tip displacements array:
Tip_q_0 = np.zeros(T+1)
Tip_q_1 = np.zeros(T+1)
# +
#Defining the different stiffness properties of the blade in a determinated radius r location:
def EI_00_f(EI_e,EI_f,twist):
return (EI_e - (EI_e-EI_f)*(math.cos(twist)**2))
def EI_11_f(EI_e,EI_f,twist):
return (EI_f + (EI_e-EI_f)*(math.cos(twist)**2))
def EI_01_f(EI_e,EI_f,twist):
return (math.sin(2*twist)*((EI_e-EI_f)/2))
def EI_10_f(EI_e,EI_f,twist):
return (math.sin(2*twist)*((EI_e-EI_f)/2))
# -
#Creation of a function that calculates the centrifugal term forces in the blade due to w:
def Centrifugal_force(N,w,df_blade_density):
N_force = 0
for k in range (N+1):
r = 1.5
rho = interpolation(r,df_blade_density)
R = 63
delta_r = R/(N-1)
N_force = N_force + rho*w*r*delta_r
r = r + delta_r
return N_force
Centrifugal_force(N,12.1,df_blade_density)
#Creation of a function that pass from rpm to rad/s:
def rpm_to_rads(w):
return (w*(2*math.pi)/60)
# # 3. MAIN PROGRAM
# +
#Define time and length steps:
total_time = 1.6
t = 0
w = rpm_to_rads(12.1) #rad/s (but visual value in rpm)
N_force = Centrifugal_force(N,w,df_blade_density)
phi_0 = math.radians(0) #rad/s initial angle for turbine 1 at t = 0s
phi_1 = math.radians(120) #rad/s initial angle for turbine 2 at t = 0s
phi_2 = math.radians(240) #rad/s initial angle for turbine 3 at t = 0s
g = 9.81 #gravity constant
#Program loop iterations
for j in range(T):
delta_t = total_time/T
r = 1.5
g_0 = 0.0
g_1 = g*math.cos(phi_1+w*t)
F_0_total = interpolation_time(find_aero_index(t,df_time_aero)[0],find_aero_index(t,df_time_aero)[1],fx_2D)
F_1_total = interpolation_time(find_aero_index(t,df_time_aero)[0],find_aero_index(t,df_time_aero)[1],ft_2D)
df_F_0 = dataframe_creation(radius_position_aero,F_0_total) #Creating dataframe aero_force direction 0 at actual time_step
df_F_1 = dataframe_creation(radius_position_aero,F_1_total) #Creating dataframe aero_force direction 1 at actual time_step
for i in range(1,N+1):
R = 63
delta_r = R/(N-1)
#First we need to define all the properties for the i position:
EI_e = (10**10)*interpolation(r,df_edgewise_stiffness)
EI_f = (10**10)*interpolation(r,df_flapwise_stiffness)
twist = interpolation(r,df_twist)
rho = interpolation(r,df_blade_density)
F_0 = interpolation(r,df_F_0) #Aerodynamic force direction 0
F_1 = interpolation(r,df_F_1) #Aerodynamic force direction 1
#Secondly calculate new values of data from the old step time:
V_0_new[i] = V_0_old[i] + delta_t*((-1/rho)*((M_0_old[i+1]-(2*M_0_old[i])+M_0_old[i-1])/delta_r**2)+((1/rho)*((S_0_old[i+1]-S_0_old[i])/delta_r))+(F_0/rho)+ g_0)
V_1_new[i] = V_1_old[i] + delta_t*((-1/rho)*((M_1_old[i+1]-(2*M_1_old[i])+M_1_old[i-1])/delta_r**2)+((1/rho)*((S_1_old[i+1]-S_1_old[i])/delta_r))+(F_1/rho)+ g_1)
#Boundary conditions at the ROOT position:
V_0_new[0] = 0
V_1_new[0] = 0
V_0_new[1] = 0
V_1_new[1] = 0
M_0_new[i] = M_0_old[i] + delta_t*((EI_00_f(EI_e,EI_f,twist)*((V_0_new[i+1]-(2*V_0_new[i])+V_0_new[i-1])/delta_r**2)+EI_01_f(EI_e,EI_f,twist)*((V_1_new[i+1]-(2*V_1_new[i])+V_1_new[i-1])/delta_r**2)))
M_1_new[i] = M_1_old[i] + delta_t*((EI_11_f(EI_e,EI_f,twist)*((V_1_new[i+1]-(2*V_1_new[i])+V_1_new[i-1])/delta_r**2)+EI_10_f(EI_e,EI_f,twist)*((V_0_new[i+1]-(2*V_0_new[i])+V_0_new[i-1])/delta_r**2)))
S_0_new[i] = S_0_old[i] + delta_t*(N_force*((V_0_new[i]-V_0_new[i-1])/delta_r))
S_1_new[i] = S_1_old[i] + delta_t*(N_force*((V_1_new[i]-V_1_new[i-1])/delta_r))
#Boundary conditions at the TIPS position:
M_0_new[N+1] = 0
M_1_new[N+1] = 0
M_0_new[N] = 0
M_1_new[N] = 0
q_0_new[i] = q_0_old[i] + delta_t*((V_0_old[i]))
q_1_new[i] = q_1_old[i] + delta_t*((V_1_old[i]))
#Position control:
Position[i] = r
r = r + delta_r
#Upload data to new time-step:
V_0_old = V_0_new
V_1_old = V_1_new
M_0_old = M_0_new
M_1_old = M_1_new
q_0_old = q_0_new
q_1_old = q_1_new
#Store data to plot:
Root_M_0[j] = M_0_new[1]
Root_M_1[j] = M_1_new[1]
Tip_V_0[j] = V_0_new[N]
Tip_V_1[j] = V_0_new[N]
Tip_q_0[j] = q_0_new[N]
Tip_q_1[j] = q_1_new[N]
#Time control:
Time[j] = t
t = t + delta_t
# -
Root_M_1
# # 4. RESULTS & PLOTS
# In this part, we will make different plots to be able to see in a better way the results obtained with the program developed.
#
# First we will need to create the new dataframes to be able to
# ## 4.1.1 TIP DISPLACEMENT OVER THE TIME
# +
#Creating the plot figure:
x = Time[:-1]
y = Tip_q_0[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Tip Displacement direction 0 [m]')
plt.xlabel('Time [s]')
plt.title('Tip Displacement direction 0 over the time');
# +
#Creating the plot figure:
x = Time[:-1]
y = Tip_q_1[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Tip Displacement direction 1 [m]')
plt.xlabel('Time [s]')
plt.title('Tip Displacement direction 1 over the time');
# -
# ## 4.1.2 DISPLACEMENT IN THE BLADE
# +
#Creating the plot figure:
x = Position[:-1]
y = q_0_new[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Displacement direction 0 [m]')
plt.xlabel('Blade radius [m]')
plt.title('Displacement along spanwise on direction 0');
# +
#Creating the plot figure:
x = Position[:-1]
#y = ((q_1_new[:-1]+(A_q/2))*math.exp(-beta*t)-(A_q/2))
y = q_1_new[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Displacement direction 1 [m]')
plt.xlabel('Blade radius [m]')
plt.title('Displacement along spanwise on direction 1');
# -
# ## 4.2.1 ROOT BENDING MOMENT OVER THE TIME
# +
#Creating the plot figure:
x = Time[:-1]
y = Root_M_0[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Root bending Moment 0 [N·m]')
plt.xlabel('Time [s]')
plt.title('Root bending Moment 0');
# +
#Creating the plot figure:
x = Time[:-1]
y = Root_M_1[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Root bending Moment 1 [N·m]')
plt.xlabel('Time [s]')
plt.title('Root bending Moment 1');
# -
# ## 4.2.2 BENDING MOMENT IN THE BLADE
# +
#Creating the plot figure:
x = Position[:-1]
y = M_0_new[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Bending Moment 0 [N·m]')
plt.xlabel('Blade radius [m]')
plt.title('Bending Moment 0 over the spanwise');
# +
#Creating the plot figure:
x = Position[:-1]
y = M_1_new[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Bending Moment 1 [N·m]')
plt.xlabel('Blade radius [m]')
plt.title('Bending Moment 1 over the spanwise');
# -
# ## 4.3.1 TIP VELOCITY DISPLACEMENT OVER THE TIME
# +
#Creating the plot figure:
x = Time[:-1]
y = Tip_V_0[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Tip Velocity direction 0 [m/s]')
plt.xlabel('Time [s]')
plt.title('Tip Velocity direction 0');
# +
#Creating the plot figure:
x = Time[:-1]
y = Tip_V_1[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Tip Velocity direction 1 [m/s]')
plt.xlabel('Time [s]')
plt.title('Tip Velocity direction 1');
# -
# ## 4.3.2 VELOCITY OF DISPLACEMENT IN THE BLADE
# +
#Creating the plot figure:
x = Position[:-1]
y = V_0_new[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Tip Velocity direction 0 [m/s]')
plt.xlabel('Blade radius [m]')
plt.title('Tip Velocity along spanwise on direction 0');
# +
#Creating the plot figure:
x = Position[:-1]
y = V_1_new[:-1]
plt.plot(x, y, color='blue', linewidth=2)
plt.ylabel('Tip Velocity direction 1 [m/s]')
plt.xlabel('Blade radius [m]')
plt.title('Tip Velocity along spanwise on direction 1');
# -
maxim(T,total_time,Root_M_1)
minim(T,total_time,Root_M_1)
amplitud(maxim(T,total_time,Root_M_1),minim(T,total_time,Root_M_1))
| REAL_BLADE_SIMULATION_60.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # The Current State of Nuclear Power
#
# In the context of world energy, where does nuclear power sit? It's producing about 18% of US electricity, but what about world electricity? What about world energy? Where is the world getting its energy, and what parts of the world are consuming what? How do different sources of energy and electricity compete and compare?
#
# ## Objectives
#
# After this lesson, you should be equipped to:
#
# - Identify the major sources of energy consumed worldwide and domestically
# - Differentiate between energy use and electricity use
# - Recognize the niche filled by nuclear power
# - Summarize the history of power from different sectors both worldwide and domestically
# - Map the geography of nuclear power production
# - Recognize the names of institutions for energy information and understand the data they collect
# - Distinguish among the major modes of energy production
#
# # Polls: Show What You Know
#
# Please go to [pollev.com/katyhuff](https://pollev.com/katyhuff).
from IPython.display import IFrame
IFrame("https://embed.polleverywhere.com/free_text_polls/CgIqfTSwUg2Ph7E?controls=none&short_poll=true", width="1000", height="700", frameBorder="0")
IFrame("https://www.polleverywhere.com/free_text_polls/4ZxqdRtjqMhDZ9A?preview=true", width=1000, height=700)
IFrame("https://embed.polleverywhere.com/free_text_polls/t6QSnqL7dORvP1d?preview=true", width=1000, height=700)
IFrame("https://www.polleverywhere.com/free_text_polls/NBjjz658zOJj6iR?preview=true", width=1000, height=700)
IFrame("https://www.polleverywhere.com/free_text_polls/Zm3pN2rMF2qFcDe?preview=true", width=1000, height=700)
# ## Energy Governance
# In the poll, you listed a lot of energy governance organiations and regulators. Here is a list with a few more:
#
# - DOE
# - NRC
# - EPA
# - state-level EPAs
# - IAEA
# - OECD-NEA
#
# ## Thinktanks, Nonprofits, Industry Coalitions, and Environmental Organizations
#
# Additionally, there is an enormous list of interest groups, thinktanks, and nonprofits that collect and share data like this. As with all sources of data, it is important to be aware of the funding model that generated the data.
#
# - Thinktanks & Policy Centers
# - [Belfer](http://belfercenter.ksg.harvard.edu/)
# - [Heritage](http://www.heritage.org/)
# - [RAND](http://www.rand.org/)
# - [Aspen](https://www.aspeninstitute.org/issues/energy-environment/)
# - [World Resources Institute](http://www.wri.org/)
# - [Breakthrough](http://thebreakthrough.org/)
# - [Third Way](http://www.thirdway.org/)
# - Industry Coalitions
# - [Nuclear Energy Institute (NEI)]()
# - [Nuclear Waste Strategy Coalition (NWSC)](http://thenwsc.org/)
# - [US Nuclear Infrastructure Council (NIC)](http://www.usnic.org/)
# - Environmental Organizations
# - [Greenpeace](http://www.greenpeace.org/usa/)
# - [Sierra Club](http://www.sierraclub.org/)
# - [UNEP](http://www.unep.org/)
# - [Environmental Progress](http://www.environmentalprogress.org/) (Action on Quad Cities and Clinton [coming up in October.](./save-the-nukes.pdf))
#
# # How to keep track?
#
# There is a lot to keep track of. There are some organizations for which keeping track of energy information is part of their energy governance missions.
#
# ## IEA
#
# The [International Energy Agency](https://iea.org) is an international organization focused on energy issues. It has 29 member countries that fund it collectively. It tackles international concerns related to energy and it was created in response to oil disruptions in the 70s. The a great resource for current information related to energy use and production worldwide. They produce an annual overview document called the World Energy Outlook. Alongside it, one can often gain access to a range of detail, from large quantities of raw data to slim primers on current energy statistics. A favorite is the [Key World Energy Statistics booklet](https://www.iea.org/publications/freepublications/publication/KeyWorld_Statistics_2015.pdf).
#
from IPython.display import IFrame
IFrame("https://www.iea.org/publications/freepublications/publication/KeyWorld_Statistics_2015.pdf", width=1000, height=1000)
# IEA also produces a Nuclear Technology Roadmap periodically, as an overview of the nuclear industry worldwide.
IFrame("http://www.iea.org/publications/freepublications/publication/Nuclear_RM_2015_FINAL_WEB_Sept_2015_V3.pdf", width=1000, height=1000)
# ## Journalism
#
# And, then, you have journalism. Lots is excellent. Some is less so. Follow the data.
IFrame("https://www.carbonbrief.org/mapped-the-worlds-nuclear-power-plants", width=1000, height=1000)
#
# At the national labs, some exceptional plots have been created to vizualize where our energy comes from and where it goes.
# 
# # EIA
#
# A lot of this information is collected by the Energy Information Administration (EIA), whose mission is steadfastly in the realm of keeping track of this data. So, the EIA is the place to go for domestic energy information.
#
#
# 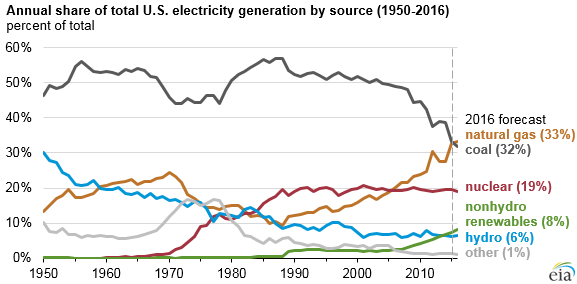
#
# ## Great Maps
# Let's explore this [EIA database](http://www.eia.gov/state/maps.cfm).
#
# ## Informative Grid Data
#
# The "grid" is also extremely important in the context of nuclear power market share. [Check out this great data from EIA about the grid.](http://www.eia.gov/beta/realtime_grid/#/summary/demand?end=20160824&start=20160724).
#
# ## Useful Nuclear Specific Data
#
# The open database can be queried directly for nuclear energy (the database code for nuclear is NUE).
# Import the IPython display module
from IPython.display import IFrame
IFrame('http:////www.eia.gov/opendata/embed/iframe.php?series_id=TOTAL.NUETPUS.A', width="100%", height=500)
IFrame("http://www.eia.gov/state/maps.cfm", width=1000, height=1000)
# ## NRC
# The Nuclear Regulatory Commission also has some useful, nuclear-specific data.
#
IFrame("http://www.nrc.gov/reactors/operating/map-power-reactors.html", width=1000, height=1000)
IFrame("https://upload.wikimedia.org/wikipedia/commons/6/6f/IL_PowerStations1.pdf", width=700, height=1000)
# ## Discussion Question: Why did the US not build new nuclear reactors for 30 years?
#
# ## Discussion Question: What worldwide impacts could result from China's new nuclear?
#
# ## Discussion Question: Why are Clinton and Quad Cities being shut down?
#
# ## Discussion Question: Why will New York state's reactors stay open?
#
# ## Resources
#
# NEI Wall Street Briefing [Video and Slides](http://www.nei.org/Issues-Policy/Economics/Financial-Analyst-Briefings/NEI-2016-Wall-Street-Briefing)
#
# Nuclear Innovation Bootcamp [closing talk by <NAME> (starts at 1:26:26)](https://www.youtube.com/watch?v=imvnBYa9Sb0&feature=youtu.be&t=5186).
| introduction/01-power-overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
# title: Iterators in Python
# toc: true
# ---
#
#
# One of the things about Python that I haven't fully appreciated are the use of iterators. I'll go over some iterators that are a part of base Python and then go over more sophisticated applications with the `itertools` package in another post. Like with many new concepts, I am grateful to be able to learn from other online sources which I acknowledge.
#
# +
# No packages necessary to import!
# -
# ## Basic iterators
#
# (I borrow shamelessly from [w3schools](https://www.w3schools.com/python/python_iterators.asp) since their page is so helpful.)
#
# It is easy to confuse an **iterable** object versus its **iterator** object. Once I made this distinction, it made it easier to understand how some methods worked. Examples of *iterable objects* are lists, dictionaries, and tuples--objects you've likely already used many times. You can get an *iterator* from these objects using the `iter()` method.
# Here's an example using a **tuple** as the iterable object.
mytuple = ("apple", "banana", "cherry")
mytupleit = iter(mytuple) # Generate it's iterator
print(mytupleit) # Note what the iterator object output when print() is called on it
# But the output of next() is a string
print(next(mytupleit))
# ...and it advances to the next item
print(next(mytupleit))
# ...and it advances to the next item
print(next(mytupleit))
# ...until the iterator is exhausted
print(next(mytupleit))
# This is a simple example but it illustrates some interesting behavior. Note that there's no explicit loop which you could do on the *iterable object* (the tuple). Instead we're outputting each element by making a `next` call on the *iterator object* (`myit`). Each `next` call "remembers" where it is in the iterable object. (Interestingly, under the hood, the `for` loop is actually creating an iterator object and using `next` method.)
#
# Here's an example using a **list** as the iterable object.
mylist = ["Winfield", "Gwynn", "Hoffman"]
mylistit = iter(mylist) # Generate it's iterator
# Let's be lazy and print it on one line
print(next(mylistit), next(mylistit), next(mylistit))
# If we want to be even more lazy, then we can also simply call `list` on the iterator to get back our original list (the iterable object). Remember the distinction between the iteratOR and the iterABLE objects!
# Regenerate the iterator
mylistit = iter(mylist)
# Get back the iterable object from the iterator
list(mylistit)
# Here's an example using a **string** as the iterable object.
mystring = "SAN"
mystringit = iter(mystring) # Generate it's iterator
print(next(mystringit), next(mystringit), next(mystringit))
# Here's an example using the **`range`** as the iterable object. Note that the range object itself is a generator-like object.
mynumbers = range(3)
mynumbersit = iter(mynumbers)
print(next(mynumbersit), next(mynumbersit), next(mynumbersit))
# ## Iterator operators `zip` and `map`
#
# Both `zip` and `map` are iterator operators that are built-in Python functions.
# Zip is commonly used to make tuples from two lists but the zip output itself is not a list
zip(mynumbers, mylist)
list(zip(mynumbers, mylist))
# From [this link](https://realpython.com/python-itertools/#what-is-itertools-and-why-should-you-use-it):
# "Under the hood, the zip() function works, in essence, by calling iter() on each of its arguments, then advancing each iterator returned by iter() with next() and aggregating the results into tuples. The iterator returned by zip() iterates over these tuples."
# Let's look at an example of the `map()` function before discussing how it works.
# Like when invoking zip, the map object itself is not a list
map(len, mylist)
list(map(len, mylist))
# What is going on here when `map()` is called? Underneath, an `iter()` object is being first called on `mylist`, advancing with `next()` then applying the first argument (`len()`) to the value returned by `next()` at each step.
# ## Acknowledgements
#
# Shout outs to the following:
#
# [w3schools](https://www.w3schools.com/python/python_iterators.asp)
#
# [RealPython](https://realpython.com/python-itertools/#what-is-itertools-and-why-should-you-use-it)
#
| _posts_drafts/2019-07-24-Python-iterators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example of confidence interval computation
# +
import numpy as np
import matplotlib.pyplot as plt
import zfit
from zfit.loss import ExtendedUnbinnedNLL
from zfit.minimize import Minuit
from hepstats.hypotests.calculators import FrequentistCalculator
from hepstats.hypotests import ConfidenceInterval
from hepstats.hypotests.parameters import POIarray
from utils import one_minus_cl_plot, pltdist, plotfitresult
# -
# ### Fit of a Gaussian signal over an exponential background:
# +
bounds = (0.1, 3.0)
# Data and signal
np.random.seed(0)
tau = -2.0
beta = -1/tau
data = np.random.exponential(beta, 300)
peak = np.random.normal(1.2, 0.1, 80)
data = np.concatenate((data,peak))
data = data[(data > bounds[0]) & (data < bounds[1])]
plt.hist(data, bins=100, histtype='step');
# -
obs = zfit.Space('x', limits=bounds)
mean = zfit.Parameter("mean", 1.2, 0.5, 2.0)
sigma = zfit.Parameter("sigma", 0.1, 0.02, 0.2)
lambda_ = zfit.Parameter("lambda",-2.0, -4.0, -1.0)
Nsig = zfit.Parameter("Nsig", 20., -20., len(data))
Nbkg = zfit.Parameter("Nbkg", len(data), 0., len(data)*1.1)
signal = zfit.pdf.Gauss(obs=obs, mu=mean, sigma=sigma).create_extended(Nsig)
background = zfit.pdf.Exponential(obs=obs, lambda_=lambda_).create_extended(Nbkg)
tot_model = zfit.pdf.SumPDF([signal, background])
# Create the negative log likelihood
data_ = zfit.data.Data.from_numpy(obs=obs, array=data)
nll = ExtendedUnbinnedNLL(model=tot_model, data=data_)
# Instantiate a minuit minimizer
minimizer = Minuit()
# minimisation of the loss function
minimum = minimizer.minimize(loss=nll)
print(minimum)
nbins = 80
pltdist(data, nbins, bounds)
plotfitresult(tot_model, bounds, nbins)
plt.xlabel("m [GeV/c$^2$]")
plt.ylabel("number of events")
# ### Confidence interval
#
# We want to compute the confidence interval of the mean of the Gaussian at 68% confidence level.
# instantation of the calculator
#calculator = FrequentistCalculator(nll, minimizer, ntoysnull=100)
calculator = FrequentistCalculator.from_yaml("toys/ci_freq_zfit_toys.yml", nll, minimizer, ntoysnull=2000)
calculator.bestfit = minimum #optionnal
# parameter of interest of the null hypothesis
poinull = POIarray(mean, np.linspace(1.15, 1.26, 50))
# instantation of the discovery test
ci = ConfidenceInterval(calculator, poinull)
ci.interval();
f = plt.figure(figsize=(9, 8))
one_minus_cl_plot(poinull.values, ci.pvalues())
plt.xlabel("mean")
calculator.to_yaml("toys/ci_freq_zfit_toys.yml")
| notebooks/hypotests/confidenceinterval_freq_zfit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Relatório de Análise VII
# ## Criando Agrupamentos
#importando a biblioteca
import pandas as pd
#importando o banco
dados = pd.read_csv('dados/aluguel_residencial.csv', sep= ';')
#visualizando os 10 primeiros
dados.head(10)
#calculando a média da coluna valor
dados['Valor'].mean()
#fazendo seleção para pegar os bairros para fazer media
bairros = ['Barra da Tijuca', 'Copacabana', 'Ipanema', 'Leblon', 'Botafogo', 'Flamengo', 'Tijuca']
selecao = dados['Bairro'].isin(bairros)
dados = dados[selecao]
#eleminando as duplicates
dados['Bairro'].drop_duplicates()
#criando o df apenas com bairro
grupo_bairro = dados.groupby('Bairro')
type(grupo_bairro)
#fazendo um laço para ver os grupos
for bairro, dados in grupo_bairro:
print(bairro)
for bairro, dados in grupo_bairro:
print('{} -> {}'.format(bairro, dados.Valor.mean()))
grupo_bairro['Valor'].mean()
grupo_bairro[['Valor', 'Condominio']].mean().round(2)
# # Estatistica descritivas
#selecionando a variável grupobairro e fazendo uma descrição
grupo_bairro['Valor'].describe().round(2)
#selecionando variável grupobairro e usando a função 'aggregate'
grupo_bairro['Valor'].aggregate(['min', 'max']).rename(columns= {'min': 'Mínimo', 'max': 'Máximo'})
# %matplotlib inline
#importando biblioteca de visualização
import matplotlib.pyplot as plt
#configurando o tamanho do gráfico
plt.rc('figure', figsize = (20, 10))
#criando gráfico para a média
fig = grupo_bairro['Valor'].mean().plot.bar(color = 'blue')
fig.set_ylabel('Valor do Aluguel')#colocando nome no eixo Y
fig.set_title('Valor Médio do Aluguel por Bairro', {'fontsize': 22})
#criando gráfico para o dvp
fig = grupo_bairro['Valor'].std().plot.bar(color = 'blue')
fig.set_ylabel('Valor do Aluguel')#colocando nome no eixo Y
fig.set_title('Valor Desvio Padrão do Aluguel por Bairro', {'fontsize': 22})
#criando gráfico para o max
fig = grupo_bairro['Valor'].max().plot.bar(color = 'blue')
fig.set_ylabel('Valor do Aluguel')#colocando nome no eixo Y
fig.set_title('Valor Máximo do Aluguel por Bairro', {'fontsize': 22})
#criando gráfico para o min
fig = grupo_bairro['Valor'].min().plot.bar(color = 'blue')
fig.set_ylabel('Valor do Aluguel')#colocando nome no eixo Y
fig.set_title('Valor Mínimo do Aluguel por Bairro', {'fontsize': 22})
#Valor por m²
fig = grupo_bairro['Valor por m²'].mean().plot.bar(color = 'blue')
fig.set_ylabel('Valor do Aluguel')#colocando nome no eixo Y
fig.set_title('Valor Médio do Aluguel por Bairro', {'fontsize': 22})
| 2. Python com Pandas Tratando e analisando dados/Projetos Python/Curso Pandas/Criando Agrupamentos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # A Framework for Comparing Graph Embeddings
#
import numpy as np
import pandas as pd
## for plotting only:
import igraph as ig
import umap
import matplotlib.pyplot as plt
# %matplotlib inline
# +
## Build igraph with colours
def buildGraph(edge_file, comm_file):
comm = pd.read_csv(comm_file, sep=r'\s+', header=None)[0].tolist()
E = pd.read_csv(edge_file, sep=r'\s+', header=None)
x = min(E.min())
E = np.array(E-x) ## make everything 0-based
n = len(comm)
E = np.array([x for x in E if x[0]<x[1]]) ## simplify
cl = ['magenta','grey','green','cyan','yellow','red','blue','tan','gold']
pal = ig.RainbowPalette(n=max(comm)+1)
v = [i for i in range(n)]
g = ig.Graph(vertex_attrs={"label":v}, edges=list(E), directed=False)
g['min']=x
g.vs["color"] = [pal.get(i) for i in comm]
g.vs['comm'] = comm
return g
## Read embedding from file in node2vec format
## For visualization: use UMAP if dim > 2
def embed2layout(fn):
D = pd.read_csv(fn, sep=' ', skiprows=1, header=None)
D = D.dropna(axis=1)
D = D.sort_values(by=0)
Y = np.array(D.iloc[:,1:])
if Y.shape[1]>=2:
Y = umap.UMAP().fit_transform(Y)
ly = []
for v in range(Y.shape[0]):
ly.append((Y[v][0],Y[v][1]))
return ly
# -
FN = "./Data/karate"
g = buildGraph(FN+'.edgelist',FN+'.community')
ly = g.layout_kamada_kawai()
ig.plot(g,layout=ly,bbox=(0,0,400,300), vertex_label_size=0, vertex_size=10 )
ly = embed2layout(FN+'.embedding.1')
ig.plot(g,layout=ly,bbox=(0,0,400,300), vertex_label_size=0, vertex_size=10 )
ly = embed2layout(FN+'.embedding.2')
ig.plot(g,layout=ly,bbox=(0,0,400,300), vertex_label_size=0, vertex_size=10 )
ly = embed2layout(FN+'.embedding.3')
ig.plot(g,layout=ly,bbox=(0,0,400,300), vertex_label_size=0, vertex_size=10 )
FN = "./Data/lfr15"
g = buildGraph(FN+'.edgelist',FN+'.community')
ly = g.layout_kamada_kawai()
ig.plot(g,layout=ly,bbox=(0,0,400,300), vertex_label_size=0, vertex_size=10 )
ly = embed2layout(FN+'.embedding.1')
ig.plot(g,layout=ly,bbox=(0,0,400,300), vertex_label_size=0, vertex_size=10 )
ly = embed2layout(FN+'.embedding.2')
ig.plot(g,layout=ly,bbox=(0,0,400,300), vertex_label_size=0, vertex_size=10 )
ly = embed2layout(FN+'.embedding.3')
ig.plot(g,layout=ly,bbox=(0,0,400,300), vertex_label_size=0, vertex_size=10 )
## running the C code to compare several embeddings
## ... make sure to compile GED.c first
import subprocess
FN = "./Data/lfr15"
for i in range(3):
x = './GED -g '+FN+'.edgelist'+' -c '+FN+'.ecg'+' -e '+FN+'.embedding.'+str(i+1)
s = subprocess.run(x, shell=True, stdout=subprocess.PIPE)
print(FN+'.embedding.'+str(i+1),':',s.stdout.decode())
| GraphEmbeddingDivergence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''arc_env'': conda)'
# language: python
# name: python37664bitarcenvconda83c4abf9215d4a698ce68e2a44e6e6bc
# ---
# # A Demo of using RDKitMol as intermediate to generate TS by ts_gen
#
# A demo to show how RDKitMol can connect RMG and GCN to help predict TS geometry. GCN requires a same atom ordering for the reactant and the product, which is seldomly accessible in practice. RDKitMol + RMG provides an opportunity to match reactant and product atom indexes according to RMG reaction family. <br>
#
# Some codes are compiled from https://github.com/PattanaikL/ts_gen and https://github.com/kspieks/ts_gen_v2.
#
# +
import os
import sys
import subprocess
# To add this RDMC into PYTHONPATH in case you doesn't do it
sys.path.append(os.path.dirname(os.path.abspath('')))
from rdmc.mol import RDKitMol
# import RMG dependencies
try:
from rdmc.external.rmg import (from_rdkit_mol,
load_rmg_database,
generate_product_complex,
mm)
# Load RMG database
database = load_rmg_database()
except (ImportError, ModuleNotFoundError):
print('You need to install RMG-Py first and run this IPYNB in rmg_env!')
# %load_ext autoreload
# %autoreload 2
# -
# ### 1. Input molecule information
# Perceive xyz and generate RMG molecule
#
# Example 1: Intra H migration
# +
reactant_xyz = """C -1.528265 0.117903 -0.48245
C -0.214051 0.632333 0.11045
C 0.185971 2.010727 -0.392941
O 0.428964 2.005838 -1.836634
O 1.53499 1.354342 -2.136876
H -1.470265 0.057863 -1.571456
H -1.761158 -0.879955 -0.103809
H -2.364396 0.775879 -0.226557
H -0.285989 0.690961 1.202293
H 0.605557 -0.056315 -0.113934
H -0.613001 2.746243 -0.275209
H 1.100271 2.372681 0.080302"""
product_xyz = """C 1.765475 -0.57351 -0.068971
H 1.474015 -1.391926 -0.715328
H 2.791718 -0.529486 0.272883
C 0.741534 0.368416 0.460793
C -0.510358 0.471107 -0.412585
O -1.168692 -0.776861 -0.612765
O -1.768685 -1.15259 0.660846
H 1.164505 1.37408 0.583524
H 0.417329 0.069625 1.470788
H -1.221189 1.194071 0.001131
H -0.254525 0.771835 -1.433299
H -1.297409 -1.977953 0.837367"""
# -
# Example 2: Intra_R_Add_Endocyclic
# +
reactant_xyz = """C -1.280629 1.685312 0.071717
C -0.442676 0.4472 -0.138756
C 0.649852 0.459775 -0.911627
C 1.664686 -0.612881 -1.217378
O 1.590475 -1.810904 -0.470776
C -0.908344 -0.766035 0.616935
O -0.479496 -0.70883 2.04303
O 0.804383 -0.936239 2.193929
H -1.330008 1.940487 1.13602
H -0.87426 2.544611 -0.46389
H -2.311393 1.527834 -0.265852
H 0.884957 1.398914 -1.412655
H 2.661334 -0.151824 -1.125202
H 1.56564 -0.901818 -2.270488
H 1.630132 -1.574551 0.469563
H -0.531309 -1.699031 0.2105
H -1.994785 -0.790993 0.711395"""
product_xyz = """C -1.515438 1.173583 -0.148858
C -0.776842 -0.102045 0.027824
C 0.680366 -0.300896 -0.240616
O 1.080339 -1.344575 0.660508
O -0.122211 -2.188293 0.768145
C -1.192654 -1.233281 0.917593
C -1.377606 -0.848982 2.395301
O -0.302953 -0.072705 2.896143
H -2.596401 1.013314 -0.200053
H -1.327563 1.859316 0.692798
H -1.211486 1.693094 -1.062486
H 0.888934 -0.598866 -1.280033
H 1.294351 0.57113 0.013413
H -2.08787 -1.759118 0.559676
H -1.514675 -1.774461 2.97179
H -2.282313 -0.243469 2.505554
H 0.511127 -0.541653 2.673033"""
# -
# Example3: ketoenol
# +
reactant_xyz = """O 0.898799 1.722422 0.70012
C 0.293754 -0.475947 -0.083092
C -1.182804 -0.101736 -0.000207
C 1.238805 0.627529 0.330521
H 0.527921 -1.348663 0.542462
H 0.58037 -0.777872 -1.100185
H -1.45745 0.17725 1.018899
H -1.813437 -0.937615 -0.310796
H -1.404454 0.753989 -0.640868
H 2.318497 0.360641 0.272256"""
product_xyz = """O 2.136128 0.058786 -0.999372
C -1.347448 0.039725 0.510465
C 0.116046 -0.220125 0.294405
C 0.810093 0.253091 -0.73937
H -1.530204 0.552623 1.461378
H -1.761309 0.662825 -0.286624
H -1.923334 -0.892154 0.536088
H 0.627132 -0.833978 1.035748
H 0.359144 0.869454 -1.510183
H 2.513751 -0.490247 -0.302535"""
# -
r_rdkitmol = RDKitMol.FromXYZ(reactant_xyz, backend='pybel', header=False)
p_rdkitmol = RDKitMol.FromXYZ(product_xyz, backend='pybel', header=False)
reactants = [from_rdkit_mol(r_rdkitmol.ToRWMol())]
products = [from_rdkit_mol(p_rdkitmol.ToRWMol())]
# ### [ALTERNATIVE] If you don't have xyzs
# +
reactant_smiles = 'CCC(=O)C'
product_smiles = 'CC=C(O)C'
r_rdkitmol = RDKitMol.FromSmiles(reactant_smiles)
p_rdkitmol = RDKitMol.FromSmiles(product_smiles)
r_rdkitmol.EmbedConformer()
p_rdkitmol.EmbedConformer()
reactants = [from_rdkit_mol(r_rdkitmol.ToRWMol())]
products = [from_rdkit_mol(p_rdkitmol.ToRWMol())]
# -
# ### 2. Check if this reaction matches RMG templates
# +
products_match = generate_product_complex(database,
reactants,
products)
# p_rmg is a product RDKitMol with the same atom order as the reactant
p_rmg = RDKitMol.FromRMGMol(products_match)
# -
# ### 4. Find structure match between RMG result and Original molecule
# +
#### 4.1
# -
# Find all possible atom mapping between the reactant and the product.
matches = p_rmg.GetSubstructMatches(p_rdkitmol, uniquify=False)
# Find the best atom mapping by RMSD. <br>
# Note, this can perform relatively poorly if the reactant and the product are in different stereotype (cis/trans). or most rotors are significantly different oriented. However, previous step (match according to RMG reaction) makes sure that all heavy atoms and reacting H atoms are consistent, so only H atoms that are more trivial are influenced by this.
# +
rmsds = []
# Make a copy of p_rdkitmol to preserve its original information
p_align = p_rdkitmol.Copy()
for i, match in enumerate(matches):
atom_map = [(ref, prb) for ref, prb in enumerate(match)]
# Align p_rdkitmol to r_rdkitmol
rmsd, reflect = p_align.GetBestAlign(refMol=r_rdkitmol,
atomMap=atom_map,
keepBestConformer=False)
rmsds.append((i, reflect, rmsd))
best = sorted(rmsds, key=lambda x: x[2])[0]
print('Match index: {0}, Reflect Conformation: {1}, RMSD: {2}'.format(*best))
# -
# Create a product molecule that has matched atom indexes and aligned xyz
# +
best_match = matches[best[0]]
p_align.AlignMol(refMol=r_rdkitmol,
atomMap=[(ref, prb) for ref, prb in enumerate(best_match)],
reflect=best[1])
new_order = [best_match.index(i) for i in range(len(best_match))]
p_rdkitmol_match = p_align.RenumberAtoms(new_order)
# -
# ### 5. View Molecules
# +
import py3Dmol
def show_mol(mol, view, grid):
from rdkit import Chem
mb = Chem.MolToMolBlock(mol)
view.removeAllModels(viewer=grid)
view.addModel(mb,'sdf', viewer=grid)
view.setStyle({'model':0},{'stick': {}}, viewer=grid)
view.zoomTo(viewer=grid)
return view
view = py3Dmol.view(width=960, height=500, linked=False, viewergrid=(1,3))
show_mol(r_rdkitmol.ToRWMol(), view, grid=(0, 0))
show_mol(p_rdkitmol_match.ToRWMol(), view, grid=(0, 1))
show_mol(p_rdkitmol.ToRWMol(), view, grid=(0, 2))
print('reactant matched product original product')
view.render()
# -
# ### 6. Export to SDF file and run ts_gen
r_rdkitmol.ToSDFFile('reactant.sdf')
p_rdkitmol_match.ToSDFFile('product.sdf')
# #### 6.1 TS Gen V2
TS_GEN_PYTHON = '~/Apps/anaconda3/envs/ts_gen_v2/bin/python3.7'
TS_GEN_DIR = '~/Apps/ts_gen_v2'
try:
subprocess.run(f'export PYTHONPATH=$PYTHONPATH:{TS_GEN_DIR};'
f'{TS_GEN_PYTHON} {TS_GEN_DIR}/inference.py '
f'--r_sdf_path reactant.sdf '
f'--p_sdf_path product.sdf '
f'--ts_xyz_path TS.xyz',
check=True,
shell=True)
except subprocess.CalledProcessError as e:
print(e)
else:
with open('TS.xyz', 'r') as f:
ts_xyz=f.read()
ts_rdkit = RDKitMol.FromXYZ(ts_xyz)
# ### 7. Visualize TS
# +
import py3Dmol
# Align the TS to make visualization more convenient
atom_map = [(i, i) for i in range(r_rdkitmol.GetNumAtoms())]
rmsd1 = ts_rdkit.AlignMol(r_rdkitmol,
atomMap=atom_map)
rmsd2 = ts_rdkit.AlignMol(r_rdkitmol,
atomMap=atom_map,
reflect=True)
if rmsd1 < rmsd2:
ts_rdkit.AlignMol(r_rdkitmol,
atomMap=atom_map,
reflect=True,
maxIters=1)
view = py3Dmol.view(width=960, height=500, linked=False, viewergrid=(1,3))
show_mol(r_rdkitmol.ToRWMol(), view, grid=(0, 0))
show_mol(ts_rdkit.ToRWMol(), view, grid=(0, 1))
show_mol(p_rdkitmol_match.ToRWMol(), view, grid=(0, 2))
print('reactant TS product')
view.render()
# -
| ipython/TS-GCN+RDMC_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/manashpratim/Algorithms-and-Data-Structures/blob/master/Graphs.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="yL2knVvzoeee" colab_type="text"
# # **Graphs Implementation (Adjacency List)**
# + id="UoeDBQ6D2Ioa" colab_type="code" colab={}
class Node(object):
def __init__(self,val,next=None):
self.val = val
self.next = next
# + id="ElAyJL0B3oT0" colab_type="code" colab={}
class LinkedList(object):
def __init__(self):
self.head = None
def isempty(self):
if not self.head:
return True
return False
def get_head(self):
return self.head
def insert_at_head(self,val):
node = Node(val)
if self.isempty():
self.head = node
else:
node.next = self.head
self.head = node
def insert_at_tail(self,val):
node = Node(val)
if self.isempty():
self.head = node
else:
curr = self.head
while curr.next:
curr = curr.next
curr.next = node
def delete_at_head(self):
if self.isempty():
print('List is empty!')
else:
temp = self.head
self.head = self.head.next
print(str(temp.val) + ' deleted!')
def delete(self,val):
if self.isempty():
print('List is empty!')
else:
curr = self.head
prev = None
while curr:
if curr.val == val:
if prev:
prev.next = curr.next
else:
self.head = curr.next
print(str(curr.val) + ' deleted!')
return
prev = curr
curr = curr.next
print(str(val)+' not in List!')
def length(self):
if self.isempty():
return 0
else:
l = 0
curr = self.head
while curr:
l = l + 1
curr = curr.next
return l
def printll(self):
if self.isempty():
print('List is empty')
else:
curr = self.head
while curr:
print(curr.val,end= '-> ')
curr = curr.next
if not curr:
print('None')
# + id="StHpTWouilmt" colab_type="code" colab={}
class Graph(object):
def __init__(self,vertices):
self.vertices = vertices
self.arr = []
for i in range(self.vertices):
self.arr.append(LinkedList())
def add_edge(self,source,des):
if source < self.vertices and des < self.vertices:
self.arr[source].insert_at_head(des)
#self.arr[des].insert_at_head(source) #Uncomment for undirected graph
def print_graph(self):
for i in range(self.vertices):
print('Vertex ',str(i)+': ',end=' ')
self.arr[i].printll()
# + id="cSLY9zHp6tTO" colab_type="code" colab={}
l = LinkedList()
# + id="WPAfFO5B60bc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e90804fb-b6a9-43ea-ea23-2cb3ade8a620"
l.insert_at_head(3)
l.insert_at_head(1)
l.insert_at_tail(4)
l.insert_at_head(6)
l.printll()
# + [markdown] id="oB7wvVk36KDZ" colab_type="text"
# ## **BFS**
# + id="DYip03gHve5N" colab_type="code" colab={}
def bfs_helper(graph,source):
q = []
q.append(source)
visited = set()
res = ''
while len(q)>0:
v = q.pop(0)
if v not in visited:
temp = graph.arr[v].get_head()
while temp:
q.append(temp.val)
temp = temp.next
res+= str(v)
visited.add(v)
return res,visited
def bfs(graph,source):
result,visited = bfs_helper(graph,source)
for i in range(graph.vertices):
if i not in visited:
res,_= bfs_helper(graph,i)
result+=res
return result
# + id="nGIBikdimqYa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="1b075744-5b1f-4f34-e78e-4cef1f5ea499"
g = Graph(6)
g.add_edge(0, 1)
g.add_edge(0, 2)
g.add_edge(1, 3)
g.add_edge(1, 4)
g.print_graph()
# + id="OoSB5YmMpYGQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="30277d47-5e19-421c-fee6-4dd455c1a5f5"
bfs(g,0)
# + [markdown] id="3VoA5yj26Oq8" colab_type="text"
# ## **DFS**
# + id="rJ2azf-i1ylB" colab_type="code" colab={}
def dfs_helper(graph,source):
s = []
s.append(source)
visited = set()
res = ''
while len(s)>0:
v = s.pop(0)
if v not in visited:
temp = graph.arr[v].get_head()
while temp:
s.insert(0,temp.val)
temp = temp.next
res = res + str(v)
visited.add(v)
return res,visited
def dfs(graph,source):
result,visited = dfs_helper(graph,source)
for i in range(graph.vertices):
if i not in visited:
res,_= dfs_helper(graph,i)
result+=res
return result
# + id="u3s_u2pA30d2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="c5866210-3c3f-416d-e506-efaa44836027"
g = Graph(7)
g.add_edge(1, 3)
g.add_edge(1, 2)
g.add_edge(2, 5)
g.add_edge(2, 4)
g.add_edge(3, 6)
g.print_graph()
# + id="p2GbY8Ex3nDU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a10c3983-e7c8-4b41-9e70-1068df1ad74b"
dfs(g,1)
# + [markdown] id="aXz-AO98_kqe" colab_type="text"
# ## **Detect Cycle**
# + id="6pBbbAbT7Fcz" colab_type="code" colab={}
def detect_helper(graph,source):
q = []
q.append(source)
visited = set()
while len(q)>0:
v = q.pop(0)
if v not in visited:
temp = graph.arr[v].get_head()
while temp:
q.append(temp.val)
temp = temp.next
visited.add(v)
else:
return True
return False
def detect_cycle(graph):
for i in range(graph.vertices):
if detect_helper(graph,i)
return True
return False
# + id="yQsq1pyD8fT6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="e298e1ae-b320-45de-d969-f41b8c524611"
g1 = Graph(3)
g1.add_edge(0, 1)
g1.add_edge(1, 2)
g1.add_edge(2, 0)
g1.print_graph()
# + id="vm2Bs7JU817A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e240b4da-e479-4ac0-8137-1e3dd8870181"
detect_cycle(g1)
# + id="4afUxXnU8tTv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="9a210fa7-c091-46ae-9134-f3e8c8561faa"
g2 = Graph(3)
g2.add_edge(0, 1)
g2.add_edge(1, 2)
g2.print_graph()
# + id="-zVzKFAm9baG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d2f0934e-96ef-483a-b5ea-fe217d7b97e8"
detect_cycle(g2)
# + id="xCOaA8iqB2Mo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="b4427eb9-a37e-4b33-a4c0-faf22135a68c"
g3 = Graph(4)
g3.add_edge(0, 1)
g3.add_edge(0, 2)
g3.add_edge(1, 2)
#g3.add_edge(2, 0)
g3.add_edge(2, 3)
g3.add_edge(3, 3)
g3.print_graph()
# + id="9whreoVwCFuB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8937d194-c4cb-4145-cf11-4d14118ad01b"
detect_cycle(g3)
# + id="0MHqn2ZiCaB6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="b0027805-7bf2-40ce-e817-76556e4b0620"
g4=Graph(6)
g4.add_edge(0,1)
g4.add_edge(1,2)
g4.add_edge(2,0)
g4.add_edge(3,4)
g4.add_edge(4,5)
g4.print_graph()
detect_cycle(g4)
# + [markdown] id="1fcn5cpexZTQ" colab_type="text"
# ## **Find Mother Vertex**
# + id="ozTT97bJbdVf" colab_type="code" colab={}
def mother_helper(graph,source,visited):
s = []
s.append(source)
visited[source] = True
res = ''
while len(s)>0:
v = s.pop(0)
temp = graph.arr[v].get_head()
while temp:
if not visited[temp.val]:
s.insert(0,temp.val)
visited[temp.val] =True
temp = temp.next
res = res + str(v)
return res,visited
def find_mother_vertex(graph):
for i in range(graph.vertices):
visited = [False]*graph.vertices
res,visited = mother_helper(graph,i,visited)
if sum(visited) == graph.vertices:
return i
return -1
# + id="KdcqgY33clVq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="cef83985-d9fd-4bc4-e37f-97cea52caf3d"
g=Graph(4)
g.add_edge(3,0)
g.add_edge(3,1)
g.add_edge(0,1)
g.add_edge(1,2)
g.print_graph()
find_mother_vertex(g)
# + id="kxnPzPPVexpJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="bb39d55d-c56f-4e0e-dfe3-95b9ea1cc60e"
g=Graph(3)
g.add_edge(0,1)
g.add_edge(1,2)
g.add_edge(2,0)
g.print_graph()
find_mother_vertex(g)
# + [markdown] id="61o4ka5p3ECp" colab_type="text"
# ## **Counting Number of Unique Edges in an Undirected Graph**
# + [markdown] id="KS53W-yU4GDj" colab_type="text"
# ### **Approach1**
# + id="bn7TDZt0xnP6" colab_type="code" colab={}
def edge_helper(graph,source,visited,unique):
s = []
s.append(source)
visited[source] = True
res = ''
while len(s)>0:
v = s.pop(0)
temp = graph.arr[v].get_head()
while temp:
if not visited[temp.val]:
s.insert(0,temp.val)
if (v,temp.val) not in unique and (temp.val,v) not in unique:
unique.add((v,temp.val))
visited[temp.val] =True
temp = temp.next
res = res + str(v)
return res,visited,unique
def count_edge1(graph):
unique = set()
for i in range(graph.vertices):
visited = [False]*graph.vertices
res,visited,unique = edge_helper(graph,i,visited,unique)
return len(unique)
# + id="LhPh_xTTydXB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="cba3dd2e-5ba4-4336-94f2-1a2edb2f1b30"
g=Graph(9)
g.add_edge(0,2)
g.add_edge(1,5)
g.add_edge(2,3)
g.add_edge(2,4)
g.add_edge(5,3)
g.add_edge(5,6)
g.add_edge(3,6)
g.add_edge(6,7)
g.add_edge(6,8)
g.add_edge(6,4)
g.add_edge(7,8)
g.print_graph()
count_edge1(g)
# + [markdown] id="mp9yX40h4J7e" colab_type="text"
# ### **Approach2**
# + id="JNjwQtvx2F_E" colab_type="code" colab={}
def count_edge2(graph):
count=0
for i in range(graph.vertices):
temp = graph.arr[i].get_head()
while temp:
temp=temp.next
count+=1
return count//2
# + id="jPrZHYRQ4A6p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="611b35c7-7779-4ed9-faf4-fd60fb6deb9c"
count_edge2(g)
# + [markdown] id="RrxMVw4XHLsL" colab_type="text"
# ## **Path between Two Vertices**
# + id="cPGKxpEZEeDv" colab_type="code" colab={}
def check_path(graph,source,destination):
s = []
s.append(source)
visited = set()
res = ''
while len(s)>0:
v = s.pop(0)
if v not in visited:
temp = graph.arr[v].get_head()
while temp:
if temp.val == destination:
return True
s.insert(0,temp.val)
temp = temp.next
res = res + str(v)
visited.add(v)
return False
# + id="0bN7OqewFVWH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="34c354d4-d168-438d-a47d-e4fe9a8ad523"
g=Graph(9)
g.add_edge(0,2)
g.add_edge(0,5)
g.add_edge(2,3)
g.add_edge(2,4)
g.add_edge(5,3)
g.add_edge(5,6)
g.add_edge(3,6)
g.add_edge(6,7)
g.add_edge(6,8)
g.add_edge(6,4)
g.add_edge(7,8)
g.print_graph()
# + id="aS3RW_xqFmeq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d1e7aee4-6269-45d5-a06d-e14eb196d070"
check_path(g,0,7)
# + [markdown] id="Lpv14pQQQLxQ" colab_type="text"
# ## **Remove Edges from a Graph**
# + id="r6pwCExQKYu8" colab_type="code" colab={}
def remove_edge(graph, source, dest):
if len(graph.arr) == 0:
return graph
if source>=len(graph.arr) or source<0:
return graph
if dest>=len(graph.arr) or dest<0:
return graph
graph.arr[source].delete(dest)
return graph
# + id="awgYuqzgLVYu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="8a001b24-07eb-4b02-f15c-c8380f9fe8c5"
g=Graph(5)
g.add_edge(0,2)
g.add_edge(0,1)
g.add_edge(1,3)
g.add_edge(2,4)
g.add_edge(4,0)
g.print_graph()
# + id="c2OhMsUvLjB4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="22aa62f1-95c3-48d0-956f-db2c12e772f6"
p = remove_edge(g, 2, 4)
# + id="TcSRNSfRMAPM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="1cf3be3b-06d1-466f-e63b-bc8bae0aae66"
p.print_graph()
# + id="_Z1p9MthReks" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="90e33e09-cc21-45c0-94ac-c15b53f0cb90"
g=Graph(6)
g.add_edge(0,2)
g.add_edge(0,1)
g.add_edge(0,3)
g.add_edge(3,5)
g.add_edge(5,4)
g.add_edge(2,4)
g.print_graph()
# + [markdown] id="wqxigXwsfpUz" colab_type="text"
# # **Tarjan's Algorithm**
# + [markdown] id="YIODgJNxuxuy" colab_type="text"
# ## **Connected Components**
# + id="RraCDQnRfLlV" colab_type="code" colab={}
from collections import defaultdict,deque
class Graph(object):
def __init__(self,vertices):
self.vertices = vertices
self.graph = defaultdict(list)
self.stack = deque()
self.stackmembers = [False]*vertices
self.discv = [-1]*vertices
self.low = [-1]*vertices
self.time = 0
def add_edge(self,u,v):
self.graph[u].append(v)
#self.graph[v].append(u) #undirected graph
def helper(self,u):
self.discv[u] = self.time
self.low[u] = self.time
self.time+=1
self.stack.appendleft(u)
self.stackmembers[u] = True
for v in self.graph[u]:
if self.discv[v] == -1:
self.helper(v)
self.low[u] = min(self.low[u],self.low[v])
elif self.stackmembers[v]:
self.low[u] = min(self.low[u],self.discv[v])
if self.low[u] == self.discv[u]:
w=-1
while w!=u and self.stack:
w = self.stack.popleft()
self.stackmembers[w] = False
print(w,end=' ')
print('\n')
def SCC(self):
for v in range(self.vertices):
if self.discv[v] == -1:
self.helper(v)
# + id="ySs1OmK1jUIY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="ca66da48-d7c3-421e-e240-d2bf435f35ac"
edges = [[0,1],[1,2],[2,0],[1,3]]
v = 4
g = Graph(v)
for i,edge in enumerate(edges):
g.add_edge(edge[0],edge[1])
g.SCC()
# + id="R4H5Sbl_iBTX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 117} outputId="605927b3-ac9f-43c7-99a5-2d997b007fbc"
g1 = Graph(5)
g1.add_edge(1, 0)
g1.add_edge(0, 2)
g1.add_edge(2, 1)
g1.add_edge(0, 3)
g1.add_edge(3, 4)
g1.SCC()
# + [markdown] id="qd6osayYu5ic" colab_type="text"
# ## **Bridges**
# + id="5j9EkBQZu4em" colab_type="code" colab={}
from collections import defaultdict,deque
class Graph(object):
def __init__(self,vertices):
self.vertices = vertices
self.graph = defaultdict(list)
self.parent = [-1]*vertices
self.visited = [False]*vertices
self.discv = [-1]*vertices
self.low = [-1]*vertices
self.time = 0
def add_edge(self,u,v):
self.graph[u].append(v)
self.graph[v].append(u) #undirected graph
def helper(self,u):
self.discv[u] = self.time
self.low[u] = self.time
self.time+=1
self.visited[u] = True
for v in self.graph[u]:
if not self.visited[v]:
self.parent[v] = u
self.helper(v)
self.low[u] = min(self.low[u],self.low[v])
if self.low[v]>self.discv[u]:
print('Bridge: '+str(u)+'->'+str(v))
elif self.parent[u] != v:
self.low[u] = min(self.low[u],self.discv[v])
def SCC(self):
for v in range(self.vertices):
if not self.visited[v]:
self.helper(v)
# + id="1S7fF54CwjQg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="06d97497-0938-4756-cb5c-3e36a0a38511"
edges = [[1,0],[2,0],[3,2],[4,2],[4,3],[3,0],[4,0]]
v = 5
g = Graph(v)
for i,edge in enumerate(edges):
g.add_edge(edge[0],edge[1])
g.SCC()
# + [markdown] id="GkZCu46S2Gqy" colab_type="text"
# ## **Articulation Points**
# + id="fw12LodW2DRz" colab_type="code" colab={}
from collections import defaultdict,deque
class Graph(object):
def __init__(self,vertices):
self.vertices = vertices
self.graph = defaultdict(list)
self.parent = [-1]*vertices
self.visited = [False]*vertices
self.discv = [-1]*vertices
self.low = [-1]*vertices
self.time = 0
self.art = []
def add_edge(self,u,v):
self.graph[u].append(v)
self.graph[v].append(u) #undirected graph
def helper(self,u):
self.discv[u] = self.time
self.low[u] = self.time
self.time+=1
children = 0
self.visited[u] = True
for v in self.graph[u]:
if not self.visited[v]:
self.parent[v] = u
children+=1
self.helper(v)
self.low[u] = min(self.low[u],self.low[v])
if self.parent[u]==-1 and children>1:
self.art.append(u)
if self.parent[u]!=-1 and self.low[v]>=self.discv[u]:
self.art.append(u)
elif self.parent[u] != v:
self.low[u] = min(self.low[u],self.discv[v])
def SCC(self):
for v in range(self.vertices):
if not self.visited[v]:
self.helper(v)
# + id="AKNv-tB83Bi0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="eb638e3e-8691-4dd8-e006-2619d6c18659"
edges = [[1,0],[2,0],[3,2],[4,2],[4,3],[3,0],[4,0]]
v = 5
g = Graph(v)
for i,edge in enumerate(edges):
g.add_edge(edge[0],edge[1])
g.SCC()
print('Articulation Points: ',g.art)
# + [markdown] id="MtGIFbUlTsKa" colab_type="text"
# ## **Graph Valid Tree**
# + id="WdNsGLtoTrk9" colab_type="code" colab={}
from collections import defaultdict
class Graph(object):
def __init__(self,vertices):
self.vertices = vertices
self.graph = defaultdict(list)
def add_edge(self,u,v):
self.graph[u].append(v)
self.graph[v].append(u)
# + id="2lb3UhoEUNSd" colab_type="code" colab={}
def dfs_detect_cycle(source,g,visited,parent):
visited[source] = True
for v in g.graph[source]:
if not visited[v]:
if dfs_detect_cycle(v,g,visited,source):
return True
else:
if v!=parent:
return True
return False
def detect_cycle(graph):
num_vertices = graph.vertices
visited = [False]*num_vertices
for v in range(num_vertices):
if not visited[v]:
if dfs_detect_cycle(v,graph,visited,-1):
return True
return False
def is_Tree(graph):
if detect_cycle(graph):
return False
vert = graph.vertices
for v in range(vert):
if not bfs(v,graph):
return False
return True
def bfs(source,graph):
visited = [False]*graph.vertices
from collections import deque
queue = deque()
queue.append(source)
while queue:
node = queue.popleft()
visited[node] = True
for v in graph.graph[node]:
if not visited[v]:
queue.append(v)
return sum(visited) == graph.vertices
# + id="sV4Jjs6wWMxs" colab_type="code" colab={}
g1 = Graph(4)
g1.add_edge(0,1)
g1.add_edge(0,2)
g1.add_edge(1,3)
g1.add_edge(2,3)
g2 = Graph(3)
g2.add_edge(0,1)
g2.add_edge(1,2)
g3 = Graph(5)
g3.add_edge(0,1)
g3.add_edge(0,2)
g3.add_edge(0,3)
g3.add_edge(1,4)
# + id="YZ3-BEzSpRAt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="ee24d512-7250-4d12-9790-b0b66c3c72d5"
print(detect_cycle(g3))
print(is_Tree(g3))
| Graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="NpJd3dlOCStH"
# <a href="https://colab.research.google.com/github/magenta/ddsp/blob/master/ddsp/colab/tutorials/1_synths_and_effects.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="hMqWDc_m6rUC"
#
# ##### Copyright 2020 Google LLC.
#
# Licensed under the Apache License, Version 2.0 (the "License");
#
#
#
#
# + colab={} colab_type="code" id="VNhgka4UKNjf"
# Copyright 2020 Google LLC. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] colab_type="text" id="ZFIqwYGbZ-df"
# # DDSP Synths and Effects
#
# This notebook demonstrates the use of several of the Synths and Effects Processors in the DDSP library. While the core functions are also directly accessible through `ddsp.core`, using Processors is the preferred API for end-2-end training.
#
# As demonstrated in the [0_processors.ipynb](colab/tutorials/0_processors.ipynb) tutorial, Processors contain the necessary nonlinearities and preprocessing in their `get_controls()` method to convert generic neural network outputs into valid processor controls, which are then converted to signal by `get_signal()`. The two methods are called in series by `__call__()`.
#
# While each processor is capable of a wide range of expression, we focus on simple examples here for clarity.
# + cellView="form" colab={} colab_type="code" id="jKDRJa6jztLT"
#@title Install and import dependencies
# %tensorflow_version 2.x
# !pip install -qU ddsp
# Ignore a bunch of deprecation warnings
import warnings
warnings.filterwarnings("ignore")
import ddsp
import ddsp.training
from ddsp.colab.colab_utils import (play, record, specplot, upload,
DEFAULT_SAMPLE_RATE)
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
sample_rate = DEFAULT_SAMPLE_RATE # 16000
# + [markdown] colab_type="text" id="6jXC-hm09dyl"
# # Synths
#
# Synthesizers, located in `ddsp.synths`, take network outputs and produce a signal (usually used as audio).
# + [markdown] colab_type="text" id="256dCv-T9xHi"
# ## Additive
# + [markdown] colab_type="text" id="5HxKR0UTGpyn"
# The additive synthesizer models a sound as a linear combination of harmonic sinusoids. Amplitude envelopes are generated with 50% overlapping hann windows. The final audio is cropped to `n_samples`.
#
# Inputs:
# * `amplitudes`: Amplitude envelope of the synthesizer output.
# * `harmonic_distribution`: Normalized amplitudes of each harmonic.
# * `frequencies`: Frequency in Hz of base oscillator.
# + colab={} colab_type="code" id="MQ-ZK2gI-df6"
n_frames = 1000
hop_size = 64
n_samples = n_frames * hop_size
# Amplitude [batch, n_frames, 1].
# Make amplitude linearly decay over time.
amps = np.linspace(1.0, -3.0, n_frames)
amps = amps[np.newaxis, :, np.newaxis]
# Harmonic Distribution [batch, n_frames, n_harmonics].
# Make harmonics decrease linearly with frequency.
n_harmonics = 20
harmonic_distribution = np.ones([n_frames, 1]) * np.linspace(1.0, -1.0, n_harmonics)[np.newaxis, :]
harmonic_distribution = harmonic_distribution[np.newaxis, :, :]
# Fundamental frequency in Hz [batch, n_frames, 1].
f0_hz = 440.0 * np.ones([1, n_frames, 1])
# + colab={} colab_type="code" id="1ZPz0Ej8-xKN"
# Create synthesizer object.
additive_synth = ddsp.synths.Additive(n_samples=n_samples,
scale_fn=ddsp.core.exp_sigmoid,
sample_rate=sample_rate)
# Generate some audio.
audio = additive_synth(amps, harmonic_distribution, f0_hz)
# Listen.
play(audio)
specplot(audio)
# + [markdown] colab_type="text" id="4HmOvGzs9zHB"
# ## Filtered Noise
#
# + [markdown] colab_type="text" id="6o8otCCoCRwe"
#
# The filtered noise synthesizer is a subtractive synthesizer that shapes white noise with a series of time-varying filter banks.
#
# Inputs:
# * `magnitudes`: Amplitude envelope of each filter bank (linearly spaced from 0Hz to the Nyquist frequency).
# + colab={} colab_type="code" id="eQSsBpCw_0f6"
n_frames = 250
n_frequencies = 1000
n_samples = 64000
# Bandpass filters, [n_batch, n_frames, n_frequencies].
magnitudes = [tf.sin(tf.linspace(0.0, w, n_frequencies)) for w in np.linspace(8.0, 80.0, n_frames)]
magnitudes = 0.5 * tf.stack(magnitudes)**4.0
magnitudes = magnitudes[tf.newaxis, :, :]
# + colab={} colab_type="code" id="Pz8vtRALBeCW"
# Create synthesizer object.
filtered_noise_synth = ddsp.synths.FilteredNoise(n_samples=n_samples,
scale_fn=None)
# Generate some audio.
audio = filtered_noise_synth(magnitudes)
# Listen.
play(audio)
specplot(audio)
# + [markdown] colab_type="text" id="d-K69x2790ad"
# ## Wavetable
# + [markdown] colab_type="text" id="xRWOXvZ5Gsek"
# The wavetable synthesizer generates audio through interpolative lookup from small chunks of waveforms (wavetables) provided by the network. In principle, it is very similar to the `Additive` synth, but with a parameterization in the waveform domain and generation using linear interpolation vs. cumulative summation of sinusoid phases.
#
# Inputs:
# * `amplitudes`: Amplitude envelope of the synthesizer output.
# * `wavetables`: A series of wavetables that are interpolated to cover n_samples.
# * `frequencies`: Frequency in Hz of base oscillator.
# + colab={} colab_type="code" id="zfYZ9P5yCjO8"
n_samples = 64000
n_wavetable = 2048
n_frames = 100
# Amplitude [batch, n_frames, 1].
amps = tf.linspace(0.5, 1e-3, n_frames)[tf.newaxis, :, tf.newaxis]
# Fundamental frequency in Hz [batch, n_frames, 1].
f0_hz = 110 * tf.linspace(1.5, 1, n_frames)[tf.newaxis, :, tf.newaxis]
# Wavetables [batch, n_frames, n_wavetable].
# Sin wave
wavetable_sin = tf.sin(tf.linspace(0.0, 2.0 * np.pi, n_wavetable))
wavetable_sin = wavetable_sin[tf.newaxis, tf.newaxis, :]
# Square wave
wavetable_square = tf.cast(wavetable_sin > 0.0, tf.float32) * 2.0 - 1.0
# Combine them and upsample to n_frames.
wavetables = tf.concat([wavetable_square, wavetable_sin], axis=1)
wavetables = ddsp.core.resample(wavetables, n_frames)
# + colab={} colab_type="code" id="3jOs_RsfCrd9"
# Create synthesizer object.
wavetable_synth = ddsp.synths.Wavetable(n_samples=n_samples,
sample_rate=sample_rate,
scale_fn=None)
# Generate some audio.
audio = wavetable_synth(amps, wavetables, f0_hz)
# Listen, notice the aliasing artifacts from linear interpolation.
play(audio)
specplot(audio)
# + [markdown] colab_type="text" id="C_lPZWTN92YJ"
# # Effects
#
# Effects, located in `ddsp.effects` are different in that they take network outputs to transform a given audio signal. Some effects, such as Reverb, optionally have trainable parameters of their own.
# + [markdown] colab_type="text" id="Lay_6Ldw93ZL"
# ## Reverb
#
# There are several types of reverberation processors in ddsp.
#
# * Reverb
# * ExpDecayReverb
# * FilteredNoiseReverb
#
# Unlike other processors, reverbs also have the option to treat the impulse response as a 'trainable' variable, and not require it from network outputs. This is helpful for instance if the room environment is the same for the whole dataset. To make the reverb trainable, just pass the kwarg `trainable=True` to the constructor
# + cellView="form" colab={} colab_type="code" id="tfNrv4MQXMW3"
#@markdown Record or Upload Audio
record_or_upload = "Record" #@param ["Record", "Upload (.mp3 or .wav)"]
record_seconds = 5#@param {type:"number", min:1, max:10, step:1}
if record_or_upload == "Record":
audio = record(seconds=record_seconds)
else:
# Load audio sample here (.mp3 or .wav3 file)
# Just use the first file.
filenames, audios = upload()
audio = audios[0]
# Add batch dimension
audio = audio[np.newaxis, :]
# Listen.
specplot(audio)
play(audio)
# + colab={} colab_type="code" id="xK951CM1XTGT"
# Let's just do a simple exponential decay reverb.
reverb = ddsp.effects.ExpDecayReverb(reverb_length=48000)
gain = [[-2.0]]
decay = [[2.0]]
# gain: Linear gain of impulse response. Scaled by self._gain_scale_fn.
# decay: Exponential decay coefficient. The final impulse response is
# exp(-(2 + exp(decay)) * time) where time goes from 0 to 1.0 over the
# reverb_length samples.
audio_out = reverb(audio, gain, decay)
# Listen.
specplot(audio_out)
play(audio_out)
# + colab={} colab_type="code" id="Skfxkn59VS01"
# Let's the filtered noise reverb can be quite expressive.
reverb = ddsp.effects.FilteredNoiseReverb(reverb_length=48000,
scale_fn=None)
# Rising gaussian filtered band pass.
n_frames = 1000
n_frequencies = 100
frequencies = np.linspace(0, sample_rate / 2.0, n_frequencies)
center_frequency = 4000.0 * np.linspace(0, 1.0, n_frames)
width = 500.0
gauss = lambda x, mu: 2.0 * np.pi * width**-2.0 * np.exp(- ((x - mu) / width)**2.0)
# Actually make the magnitudes.
magnitudes = np.array([gauss(frequencies, cf) for cf in center_frequency])
magnitudes = magnitudes[np.newaxis, ...]
magnitudes /= magnitudes.sum(axis=-1, keepdims=True) * 5
# Apply the reverb.
audio_out = reverb(audio, magnitudes)
# Listen.
specplot(audio_out)
play(audio_out)
plt.matshow(np.rot90(magnitudes[0]), aspect='auto')
plt.title('Impulse Response Frequency Response')
plt.xlabel('Time')
plt.ylabel('Frequency')
plt.xticks([])
_ = plt.yticks([])
# + [markdown] colab_type="text" id="Lc_md-cD99y6"
# ## FIR Filter
#
# Linear time-varying finite impulse response (LTV-FIR) filters are a broad class of filters that can vary over time.
# + cellView="form" colab={} colab_type="code" id="Gv5GQX5oeL-f"
#@markdown Record or Upload Audio
record_or_upload = "Record" #@param ["Record", "Upload (.mp3 or .wav)"]
record_seconds = 5#@param {type:"number", min:1, max:10, step:1}
if record_or_upload == "Record":
audio = record(seconds=record_seconds)
else:
# Load audio sample here (.mp3 or .wav3 file)
# Just use the first file.
filenames, audios = upload()
audio = audios[0]
# Add batch dimension
audio = audio[np.newaxis, :]
# Listen.
specplot(audio)
play(audio)
# + colab={} colab_type="code" id="YAJYBSUSfADI"
# Let's make an oscillating gaussian bandpass filter.
fir_filter = ddsp.effects.FIRFilter(scale_fn=None)
# Make up some oscillating gaussians.
n_seconds = audio.size / sample_rate
frame_rate = 100 # Hz
n_frames = int(n_seconds * frame_rate)
n_samples = int(n_frames * sample_rate / frame_rate)
audio_trimmed = audio[:, :n_samples]
n_frequencies = 1000
frequencies = np.linspace(0, sample_rate / 2.0, n_frequencies)
lfo_rate = 0.5 # Hz
n_cycles = n_seconds * lfo_rate
center_frequency = 1000 + 500 * np.sin(np.linspace(0, 2.0*np.pi*n_cycles, n_frames))
width = 500.0
gauss = lambda x, mu: 2.0 * np.pi * width**-2.0 * np.exp(- ((x - mu) / width)**2.0)
# Actually make the magnitudes.
magnitudes = np.array([gauss(frequencies, cf) for cf in center_frequency])
magnitudes = magnitudes[np.newaxis, ...]
magnitudes /= magnitudes.max(axis=-1, keepdims=True)
# Filter.
audio_out = fir_filter(audio_trimmed, magnitudes)
# Listen.
play(audio_out)
specplot(audio_out)
_ = plt.matshow(np.rot90(magnitudes[0]), aspect='auto')
plt.title('Frequency Response')
plt.xlabel('Time')
plt.ylabel('Frequency')
plt.xticks([])
_ = plt.yticks([])
# + [markdown] colab_type="text" id="mkjWUGpZ95Mr"
# ## ModDelay
#
# Variable length delay lines create an instantaneous pitch shift that can be useful in a variety of time modulation effects such as [vibrato](https://en.wikipedia.org/wiki/Vibrato), [chorus](https://en.wikipedia.org/wiki/Chorus_effect), and [flanging](https://en.wikipedia.org/wiki/Flanging).
# + cellView="form" colab={} colab_type="code" id="F_mihqZZx_s_"
#@markdown Record or Upload Audio
record_or_upload = "Record" #@param ["Record", "Upload (.mp3 or .wav)"]
record_seconds = 5#@param {type:"number", min:1, max:10, step:1}
if record_or_upload == "Record":
audio = record(seconds=record_seconds)
else:
# Load audio sample here (.mp3 or .wav3 file)
# Just use the first file.
filenames, audios = upload()
audio = audios[0]
# Add batch dimension
audio = audio[np.newaxis, :]
# Listen.
specplot(audio)
play(audio)
# + colab={} colab_type="code" id="KSEpt_DVEGbZ"
def sin_phase(mod_rate):
"""Helper function."""
n_samples = audio.size
n_seconds = n_samples / sample_rate
phase = tf.sin(tf.linspace(0.0, mod_rate * n_seconds * 2.0 * np.pi, n_samples))
return phase[tf.newaxis, :, tf.newaxis]
def modulate_audio(audio, center_ms, depth_ms, mod_rate):
mod_delay = ddsp.effects.ModDelay(center_ms=center_ms,
depth_ms=depth_ms,
gain_scale_fn=None,
phase_scale_fn=None)
phase = sin_phase(mod_rate) # Hz
gain = 1.0 * np.ones_like(audio)[..., np.newaxis]
audio_out = 0.5 * mod_delay(audio, gain, phase)
# Listen.
play(audio_out)
specplot(audio_out)
# Three different effects.
print('Flanger')
modulate_audio(audio, center_ms=0.75, depth_ms=0.75, mod_rate=0.25)
print('Chorus')
modulate_audio(audio, center_ms=25.0, depth_ms=1.0, mod_rate=2.0)
print('Vibrato')
modulate_audio(audio, center_ms=25.0, depth_ms=12.5, mod_rate=5.0)
| ddsp/colab/tutorials/1_synths_and_effects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
# This is a simple end to end example of how you can use SAS Viya for analysis
# The example follows these steps:
# 1. Importing the needed Python packages
# 1. Starting a CAS session on an already running CAS server
# 1. Load the needed CAS Action Sets
# 1. Loading data from the local file system to the CAS server
# 1. Explore the data
# 1. Impute missing values
# 1. Partition the data into training and validation partitions
# 1. Build a decision tree
# 1. Build a neural network
# 1. Build a decision forest
# 1. Build a gradient boost
# 1. Assess the models
# 1. Build ROC charts
# ## Set up and initialize
#
# Find doc for all the CAS actions [here](http://go.documentation.sas.com/?cdcId=vdmmlcdc&cdcVersion=8.11&docsetId=caspg&docsetTarget=titlepage.htm
# )
#
# ### Documentation Links:
# * [SAS® Viya™ 3.2: System Programming Guide](http://go.documentation.sas.com/?cdcId=vdmmlcdc&cdcVersion=8.11&docsetId=caspg&docsetTarget=titlepage.htm)
# * [Getting Started with SAS® Viya™ 3.2 for Python](http://go.documentation.sas.com/?cdcId=vdmmlcdc&cdcVersion=8.11&docsetId=caspg3&docsetTarget=titlepage.htm&locale=en)
#
# In this code we import the needed packages and we assign variables for the modeling details that will be used later in the analysis
# +
import os
import pandas as pd
import swat
import sys
from matplotlib import pyplot as plt
# %matplotlib inline
target = "bad"
class_inputs = ["reason", "job"]
class_vars = [target] + class_inputs
interval_inputs = ["im_clage", "clno", "im_debtinc", "loan", "mortdue", "value", "im_yoj", "im_ninq", "derog", "im_delinq"]
all_inputs = interval_inputs + class_inputs
indata = 'hmeq'
# -
# ## Start CAS session
#
# * Documentation to [Connect and Start a Session](http://go.documentation.sas.com/?cdcId=vdmmlcdc&cdcVersion=8.11&docsetId=caspg3&docsetTarget=home.htm&locale=en)
#
# In this code we assign values for the cashost, casport, and casauth values. These are then used to establish a CAS session named `sess`.
# +
cashost='localhost'
casport=5570
sess = swat.CAS(cashost, casport)
# Load the needed action sets for this example:
sess.loadactionset('datastep')
sess.loadactionset('datapreprocess')
sess.loadactionset('cardinality')
sess.loadactionset('sampling')
sess.loadactionset('regression')
sess.loadactionset('decisiontree')
sess.loadactionset('neuralnet')
sess.loadactionset('svm')
sess.loadactionset('astore')
sess.loadactionset('percentile')
# -
# ## Load data into CAS
#
#
indata = sess.CASTable('hmeq')
if not indata.tableexists().exists:
indata = sess.upload_file('http://support.sas.com/documentation/onlinedoc/viya/exampledatasets/hmeq.csv', casout=indata)
# ## Explore and Impute missing values
indata.summary()
# #### Explore data and plot missing values
# +
tbl_data_card = sess.CASTable('data_card', replace=True)
indata.cardinality.summarize(cardinality=tbl_data_card)
tbl_data_card = tbl_data_card.query('_NMISS_ > 0')
tbl_data_card.head()
# +
tbl_data_card['PERCENT_MISSING'] = (tbl_data_card['_NMISS_'] / tbl_data_card['_NOBS_']) * 100
ax = tbl_data_card[['_VARNAME_', 'PERCENT_MISSING']].to_frame().set_index('_VARNAME_').plot.bar(
title='Percentage of Missing Values', figsize=(15,7)
)
ax.set_ylabel('Percent Missing')
ax.set_xlabel('Variable Names');
# -
# #### Impute missing values
# +
hmeq_prepped = sess.CASTable('hmeq_prepped', replace=True)
indata.datapreprocess.transform(
casout=hmeq_prepped,
copyallvars=True,
outvarsnameglobalprefix='im',
requestpackages=[
{'impute': {'method': 'mean'}, 'inputs': ['clage']},
{'impute': {'method': 'median'}, 'inputs': ['delinq']},
{'impute': {'method': 'value', 'valuescontinuous': [2]}, 'inputs': ['ninq']},
{'impute': {'method': 'value', 'valuescontinuous': [35.0, 7, 2]}, 'inputs': ['debtinc', 'yoj']}
]
)
# -
# ## Partition data into Training and Validation
#
# The stratified action in the sampling actionset allows us to create two partition and observe the reponse rate of the target variable `bad` in both training and validation
# +
hmeq_part = sess.CASTable('hmeq_part', replace=True)
hmeq_prepped.groupby(target).sampling.stratified(
output=dict(casout=hmeq_part, copyvars='all'),
samppct=70,
partind=True
)
# -
# ## Decision Tree
#
# In this code block we do the following:
# 1. Train the decision tree using the variable listed we defined in the setup phase. We save the decision tree model `tree_model`. It is used in the subsequent step but it could just have easily been used a day, week, or month from now.
# 1. Score data using the `tree_model` that was created in the previous step
# 1. Run data step code on the scored output to prepare it for further analysis
# +
hmeq_part_1 = hmeq_part.query('_partind_ = 1')
tree_model = sess.CASTable('tree_model', replace=True)
scored_tree = sess.CASTable('_scored_tree', replace=True)
hmeq_part_1.decisiontree.dtreetrain(
inputs=all_inputs,
target='bad',
nominals=class_vars,
crit='gain',
prune=True,
varImp=True,
missing='useinsearch',
casout=tree_model
)
# Score
hmeq_part.decisiontree.dtreescore(
modeltable=tree_model,
casout=scored_tree,
copyvars=['bad', '_partind_']
)
# Create p_bad0 and p_bad1 as _dt_predp_ is the probability of event in _dt_predname_
scored_tree['p_bad1'] = scored_tree.eval("ifn( strip(_dt_predname_) = '1', _dt_predp_, 1-_dt_predp_ )")
scored_tree['p_bad0'] = scored_tree.eval("ifn( strip(_dt_predname_) = '0', 1-_dt_predp_, _dt_predp_ )")
# -
# ## Decision Forest
#
# In this code block we do the following:
# 1. Train the decision tree using the variable listed we defined in the setup phase. We save the decision tree model `forest_model`. It is used in the subsequent step but it could just have easily been used a day, week, or month from now.
# 1. Score data using the `forest_model` that was created in the previous step
# 1. Run data step code on the scored output to prepare it for further analysis
# +
forest_model = sess.CASTable('forest_model', replace=True)
scored_rf = sess.CASTable('_scored_rf', replace=True)
hmeq_part_1.decisiontree.foresttrain(
inputs=all_inputs,
nominals=class_vars,
target='bad',
ntree=50,
nbins=20,
leafsize=5,
maxlevel=21,
crit='gainratio',
varimp=True,
missing='useinsearch',
vote='prob',
casout=forest_model
)
# Score
hmeq_part.decisiontree.forestscore(
modeltable=forest_model,
casout=scored_rf,
copyvars=['bad', '_partind_'],
vote='prob'
)
# Create p_bad0 and p_bad1 as _rf_predp_ is the probability of event in _rf_predname_
scored_rf['p_bad1'] = scored_rf.eval("ifn( strip(_rf_predname_) = '1', _rf_predp_, 1-_rf_predp_ )")
scored_rf['p_bad0'] = scored_rf.eval("ifn( strip(_rf_predname_) = '0', 1-_rf_predp_, _rf_predp_ )")
# -
# ## Gradient Boosting Machine
#
#
# In this code block we do the following:
# 1. Train the decision tree using the variable listed we defined in the setup phase. We save the decision tree model `gb_model`. It is used in the subsequent step but it could just have easily been used a day, week, or month from now.
# 1. Score data using the `gb_model` that was created in the previous step
# 1. Run data step code on the scored output to prepare it for further analysis
# +
gb_model = sess.CASTable('gb_model', replace=True)
scored_gb = sess.CASTable('_scored_gb', replace=True)
hmeq_part_1.decisiontree.gbtreetrain(
inputs=all_inputs,
nominals=class_vars,
target=target,
ntree=10,
nbins=20,
maxlevel=6,
varimp=True,
missing='useinsearch',
casout=gb_model
)
# Score
hmeq_part.decisionTree.gbtreeScore(
modeltable=gb_model,
casout=scored_gb,
copyvars=[target, '_partind_']
)
# Create p_bad0 and p_bad1 as _gbt_predp_ is the probability of event in _gbt_predname_
scored_gb['p_bad1'] = scored_gb.eval("ifn( strip(_gbt_predname_) = '1', _gbt_predp_, 1-_gbt_predp_ )")
scored_gb['p_bad0'] = scored_gb.eval("ifn( strip(_gbt_predname_) = '0', 1-_gbt_predp_, _gbt_predp_ )")
# -
# ## Neural Network
#
# In this code block we do the following:
# 1. Train the decision tree using the variable listed we defined in the setup phase. We save the decision tree model `nnet_model`. It is used in the subsequent step but it could just have easily been used a day, week, or month from now.
# 1. Score data using the `nnet_model` that was created in the previous step
# 1. Run data step code on the scored output to prepare it for further analysis
# +
hmeq_part_0 = hmeq_part.query('_partind_ = 0')
nnet_model = sess.CASTable('nnet_model', replace=True)
scored_nn = sess.CASTable('_scored_nn', replace=True)
hmeq_part_1.neuralnet.anntrain(
validtable=hmeq_part_0,
inputs=all_inputs,
nominals=class_vars,
target="bad",
hiddens={9},
acts=['tanh'],
combs=['linear'],
targetact='softmax',
errorfunc='entropy',
std='midrange',
randdist='uniform',
scaleinit=1,
nloopts={
'optmlopt': {'maxiters': 250, 'fconv': 1e-10},
'lbfgsopt': {'numcorrections': 6},
'printopt': {'printlevel': 'printdetail'},
'validate': {'frequency': 1}
},
casout=nnet_model
)
# Score
hmeq_part.neuralnet.annscore(
modeltable=nnet_model,
casout=scored_nn,
copyvars=['bad', '_partind_']
)
# Create p_bad0 and p_bad1 as _nn_predp_ is the probability of event in _nn_predname_
scored_nn['p_bad1'] = scored_nn.eval("ifn( strip(_nn_predname_) = '1', _nn_predp_, 1-_nn_predp_ )")
scored_nn['p_bad0'] = scored_nn.eval("ifn( strip(_nn_predname_) = '0', 1-_nn_predp_, _nn_predp_ )")
# -
# ## Assess Models
# +
def assess_model(t):
return sess.percentile.assess(
table=t.query('_partind_ = 0'),
inputs=['p_bad1'],
response='bad',
event='1',
pvar=['p_bad0'],
pevent=['0']
)
tree_assess = assess_model(scored_tree)
tree_fitstat = tree_assess.FitStat
tree_rocinfo = tree_assess.ROCInfo
tree_liftinfo = tree_assess.LIFTInfo
rf_assess = assess_model(scored_rf)
rf_fitstat = rf_assess.FitStat
rf_rocinfo = rf_assess.ROCInfo
rf_liftinfo = rf_assess.LIFTInfo
gb_assess = assess_model(scored_gb)
gb_fitstat = gb_assess.FitStat
gb_rocinfo = gb_assess.ROCInfo
gb_liftinfo = gb_assess.LIFTInfo
nn_assess = assess_model(scored_nn)
nn_fitstat = nn_assess.FitStat
nn_rocinfo = nn_assess.ROCInfo
nn_liftinfo = nn_assess.LIFTInfo
# -
# ## Create ROC and Lift plots (using Validation data)
# #### Prepare assessment results for plotting
# +
# Add new variable to indicate type of model
tree_liftinfo['model'] = 'DecisionTree'
tree_rocinfo['model'] = 'DecisionTree'
rf_liftinfo['model'] = 'Forest'
rf_rocinfo['model'] = 'Forest'
gb_liftinfo['model'] = 'GradientBoosting'
gb_rocinfo['model'] = 'GradientBoosting'
nn_liftinfo['model'] = 'NeuralNetwork'
nn_rocinfo['model'] = 'NeuralNetwork'
# Concatenate data
all_liftinfo = pd.concat([rf_liftinfo, gb_liftinfo, nn_liftinfo, tree_liftinfo], ignore_index=True)
all_rocinfo = pd.concat([rf_rocinfo, gb_rocinfo, nn_rocinfo, tree_rocinfo], ignore_index=True)
# -
# ## Print AUC (Area Under the ROC Curve)
all_rocinfo[['model', 'C']].drop_duplicates(keep='first').sort_values(by='C', ascending=False)
# ## Draw ROC and Lift plots
# +
# Draw ROC charts
plt.figure(figsize=(15, 5))
for key, grp in all_rocinfo.groupby(['model']):
plt.plot(grp['FPR'], grp['Sensitivity'], label=key)
plt.plot([0,1], [0,1], 'k--')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.grid(True)
plt.legend(loc='best')
plt.title('ROC Curve')
plt.show()
# Draw lift charts
plt.figure(figsize=(15, 5))
for key, grp in all_liftinfo.groupby(['model']):
plt.plot(grp['Depth'], grp['CumLift'], label=key)
plt.xlabel('Depth')
plt.ylabel('Cumulative Lift')
plt.grid(True)
plt.legend(loc='best')
plt.title('Cumulative Lift Chart')
plt.show();
# -
# ## End CAS session
#
# This closes the CAS session freeing resources for others to leverage
# This is the same as sess.endsession(); sess.close();
sess.terminate()
| developerTrial/python/Basic+Predictive+Modeling+Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib widget
# +
import matplotlib as mpl
from matplotlib.gridspec import GridSpec
import pickle
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import scipy
from scipy import stats
# import seaborn as sns
import ipywidgets
from ipywidgets import widgets
# from IPython.display import display
import os
import mplcursors
# import re
import nmrProblem_28122020 as nmrProblem
import nmrglue as ng
import tempfile
import io
import ipysheet
from ipysheet import from_dataframe, to_dataframe
# -
df_dict = {'ppm': {'integral': '',
'symmetry': '',
'symmetry factor': '',
'J type': '',
'J Hz': '',
'C13 hyb': '',
'attached protons': '',
'ppm': '',
'H4': 1.19,
'H3': 2.27,
'H2': 2.4,
'H1': 4.15,
'C4': 19.0,
'C3': 49.0,
'C2': 68.0,
'C1': 184.0,
'hsqc': '',
'hmbc': '',
'cosy': ''},
'H1': {'integral': 1,
'symmetry': '',
'symmetry factor': '',
'J type': 'tq',
'J Hz': [6.0, 10.0],
'C13 hyb': 1,
'attached protons': '',
'ppm': 4.15,
'H4': 'o',
'H3': 'o',
'H2': 'o',
'H1': '',
'C4': '',
'C3': 'x',
'C2': 'o',
'C1': 'x',
'hsqc': ['C2'],
'hmbc': ['C1', 'C3'],
'cosy': ['H2', 'H3', 'H4']},
'H2': {'integral': 1,
'symmetry': '',
'symmetry factor': '',
'J type': 'dd',
'J Hz': [6.0, 3.0],
'C13 hyb': 2,
'attached protons': '',
'ppm': 2.4,
'H4': '',
'H3': 'o',
'H2': '',
'H1': 'o',
'C4': 'x',
'C3': 'o',
'C2': 'x',
'C1': 'x',
'hsqc': ['C3'],
'hmbc': ['C1', 'C2', 'C4'],
'cosy': ['H1', 'H3']},
'H3': {'integral': 1,
'symmetry': '',
'symmetry factor': '',
'J type': 'dd',
'J Hz': [6.0, 3.0],
'C13 hyb': 2,
'attached protons': '',
'ppm': 2.27,
'H4': '',
'H3': '',
'H2': 'o',
'H1': 'o',
'C4': 'x',
'C3': 'o',
'C2': 'x',
'C1': 'x',
'hsqc': ['C3'],
'hmbc': ['C1', 'C2', 'C4'],
'cosy': ['H1', 'H2']},
'H4': {'integral': 3,
'symmetry': '',
'symmetry factor': '',
'J type': 'd',
'J Hz': [10.0],
'C13 hyb': 3,
'attached protons': '',
'ppm': 1.19,
'H4': '',
'H3': '',
'H2': '',
'H1': 'o',
'C4': 'o',
'C3': 'x',
'C2': 'x',
'C1': '',
'hsqc': ['C4'],
'hmbc': ['C2', 'C3'],
'cosy': ['H1']},
'C1': {'integral': 1,
'symmetry': '',
'symmetry factor': '',
'J type': 's',
'J Hz': [0.0],
'C13 hyb': 0,
'attached protons': 0,
'ppm': 184.0,
'H4': '',
'H3': 'x',
'H2': 'x',
'H1': 'x',
'C4': '',
'C3': '',
'C2': '',
'C1': '',
'hsqc': [],
'hmbc': ['H1', 'H2', 'H3'],
'cosy': ''},
'C2': {'integral': 1,
'symmetry': '',
'symmetry factor': '',
'J type': 's',
'J Hz': [0.0],
'C13 hyb': 1,
'attached protons': 1,
'ppm': 68.0,
'H4': 'x',
'H3': 'x',
'H2': 'x',
'H1': 'o',
'C4': '',
'C3': '',
'C2': '',
'C1': '',
'hsqc': ['H1'],
'hmbc': ['H2', 'H3', 'H4'],
'cosy': ''},
'C3': {'integral': 1,
'symmetry': '',
'symmetry factor': '',
'J type': 's',
'J Hz': [0.0],
'C13 hyb': 2,
'attached protons': 2,
'ppm': 49.0,
'H4': 'x',
'H3': 'o',
'H2': 'o',
'H1': 'x',
'C4': '',
'C3': '',
'C2': '',
'C1': '',
'hsqc': ['H2', 'H3'],
'hmbc': ['H1', 'H4'],
'cosy': ''},
'C4': {'integral': 1,
'symmetry': '',
'symmetry factor': '',
'J type': 's',
'J Hz': [0.0],
'C13 hyb': 3,
'attached protons': 3,
'ppm': 19.0,
'H4': 'o',
'H3': 'x',
'H2': 'x',
'H1': '',
'C4': '',
'C3': '',
'C2': '',
'C1': '',
'hsqc': ['H4'],
'hmbc': ['H2', 'H3'],
'cosy': ''}}
from_dataframe(pd.DataFrame(df_dict))
df = pd.DataFrame(df_dict)
sheets= []
sheets.append(from_dataframe(df))
sheets[0]
sheets.append(from_dataframe(df[['H1','H2','H3','H4']]))
title = widgets.Label("DataFrame")
vbox = widgets.VBox([title, from_dataframe(df)])
vbox
vbox.children = [title, from_dataframe(df[['H1','H2','H3','H4']])]
to_dataframe(sheets[1])
to_dataframe(vbox.children[-1])
type(sheets[-1])
sheets[0].cells
for c in sheets[0].cells:
print(c)
| sheets_001.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// # Displaying output in a .NET notebook
// When writing C# in a .NET notebook, the C# scripting language is used, which you might be familiar with from using the C# Interactive window in Visual Studio. This dialect of C# allows you to end a code submission without a semicolon, which tells C# scripting to return the value of the expression.
var x = "Hello!";
display(x);
x
// When you end a cell with an expression evaluation like this, it is the return value of the cell. There can only be a single return value for a cell. If you add more code after a return value expression, you'll see a compile error.
//
// There are also several ways to display information without using the return value. The most intuitive one for many .NET users is to write to the console:
Console.Write("hello");
Console.Write(" ");
Console.WriteLine("world");
// But a more familiar API for many notebook users would be the `display` method.
display("hello");
display("world");
// Each call to `display` writes an additional display data value to the notebook.
// You can also update an existing displayed value by calling `Update` on the object returned by a call to `display`.
// +
var fruitOutput = display("Let's get some fruit!");
var basket = new [] {"apple", "orange", "coconut", "pear", "peach"};
foreach (var fruit in basket)
{
System.Threading.Thread.Sleep(1000);
fruitOutput.Update($"I have 1 {fruit}.");
}
System.Threading.Thread.Sleep(1000);
fruitOutput.Update(basket);
// -
// ---
// **_See also_**
// * [Formatters](Formatters.ipynb)
// * [HTML](HTML.ipynb)
| NotebookExamples/csharp/Docs/Displaying output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xarray as xr
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# #### DIC in the water column
ds = xr.open_dataset('data/base/kz145_scale10000/water.nc')
dic_df = ds['B_C_DIC'].to_dataframe()
dic_surface = dic_df.groupby('z').get_group(1.250)
dic = dic_surface.loc['2011-01-01':'2011-12-31']
ds = xr.open_dataset('data/with_wspeed_15/water.nc')
dic_df = ds['B_C_DIC'].to_dataframe()
dic_surface = dic_df.groupby('z').get_group(1.250)
dic_with_flux = dic_surface.loc['2011-01-01':'2011-12-31']
dic = dic.reset_index()
dic_with_flux = dic_with_flux.reset_index()
fig = plt.figure(figsize=(10, 2))
ax = fig.add_subplot(1, 1, 1)
ax.plot(dic['time'], dic['B_C_DIC'], linewidth=2,
label=r'with wind speed 5 m sec$^{-1}$')
ax.plot(dic_with_flux['time'], dic_with_flux['B_C_DIC'],
linewidth=2, label=r'with wind speed 15 m sec$^{-1}$')
ax.set_title("DIC concentrations")
ax.legend(loc='best');
dic['B_C_DIC'].values.mean() # no inflow
dic_with_flux['B_C_DIC'].values.mean() # with inflow
# #### TA in the water column
ds = xr.open_dataset('data/base/kz145_scale10000/water.nc')
alk_df = ds['B_C_Alk'].to_dataframe()
alk_surface = alk_df.groupby('z').get_group(1.250)
alk = alk_surface.loc['2011-01-01':'2011-12-31']
ds = xr.open_dataset('data/with_wspeed_15/water.nc')
alk_df = ds['B_C_Alk'].to_dataframe()
alk_surface = alk_df.groupby('z').get_group(1.250)
alk_with_flux = alk_surface.loc['2011-01-01':'2011-12-31']
alk = alk.reset_index()
alk_with_flux = alk_with_flux.reset_index()
fig = plt.figure(figsize=(10, 2))
ax = fig.add_subplot(1, 1, 1)
ax.plot(alk['time'], alk['B_C_Alk'], linewidth=2,
label=r'with wind speed 5 m sec$^{-1}$')
ax.plot(alk_with_flux['time'], alk_with_flux['B_C_Alk'],
linewidth=2, label=r'with wind speed 15 m sec$^{-1}$')
ax.set_title("TA")
ax.legend(loc='upper left');
alk['B_C_Alk'].values.mean() # no inflow
alk_with_flux['B_C_Alk'].values.mean() # with inflow
# #### The water column pCO$_2$
ds = xr.open_dataset('data/base/kz145_scale10000/water.nc')
co2_df = ds['B_C_pCO2'].to_dataframe()
co2_surface = co2_df.groupby('z').get_group(1.250)
co2_no = co2_surface.loc['2011-01-01':'2011-12-31']
ds = xr.open_dataset('data/with_wspeed_15/water.nc')
co2_df = ds['B_C_pCO2'].to_dataframe()
co2_surface = co2_df.groupby('z').get_group(1.250)
co2_with_flux = co2_surface.loc['2011-01-01':'2011-12-31']
co2_no = co2_no.reset_index()
co2_with_flux = co2_with_flux.reset_index()
co2_no['B_C_pCO2'] *= 1e6 # to convert to ppm
co2_with_flux['B_C_pCO2'] *= 1e6
atm_pCO2 = 390 # to be comparable with Thomas 2004
dco2_no = co2_no['B_C_pCO2']-atm_pCO2
dco2_with_flux = co2_with_flux['B_C_pCO2']-atm_pCO2
fig = plt.figure(figsize=(10, 3))
ax = fig.add_subplot(1, 1, 1)
ax.plot(co2_no['time'], dco2_no, linewidth=2,
label=r'with wind speed 5 m sec$^{-1}$')
ax.plot(co2_with_flux['time'], dco2_with_flux,
linewidth=2, label=r'with wind speed 15 m sec$^{-1}$')
ax.set_title('pCO$_2$ difference between seawater and atmosphere')
ax.legend(loc='best');
co2_no['B_C_pCO2'].mean() # no flux
co2_with_flux['B_C_pCO2'].mean() # with flux
dco2_no.mean()
dco2_with_flux.mean()
# #### Surface fluxes of CO$_2$
import numpy as np
import pandas as pd
ds = xr.open_dataset('data/base/kz145_scale10000/water.nc')
co2flux_df = ds['B_C_DIC _flux'].to_dataframe()
co2flux_surface = co2flux_df.groupby('z_faces').get_group(0)
co2flux_no = -co2flux_surface.loc['2011-01-01':'2011-12-31']
ds = xr.open_dataset('data/with_wspeed_15/water.nc')
co2flux_df = ds['B_C_DIC _flux'].to_dataframe()
co2flux_surface = co2flux_df.groupby('z_faces').get_group(0)
co2flux_alk = -co2flux_surface.loc['2011-01-01':'2011-12-31']
# *Calculate the monthly CO$_2$ flux*
year = (('2011-01-01','2011-01-31'), ('2011-02-01','2011-02-28'), ('2011-03-01','2011-03-31'),
('2011-04-01','2011-04-30'), ('2011-05-01','2011-05-31'), ('2011-06-01','2011-06-30'),
('2011-07-01','2011-07-31'), ('2011-08-01','2011-08-31'), ('2011-09-01','2011-09-30'),
('2011-10-01','2011-10-31'), ('2011-11-01','2011-11-30'), ('2011-12-01','2011-12-31'))
co2flux_no_year = []
co2flux_alk_year = []
for month in year:
co2flux_no_month = co2flux_no.loc[month[0]:month[1]]
co2flux_alk_month = co2flux_alk.loc[month[0]:month[1]]
co2flux_no_year.append(co2flux_no_month['B_C_DIC _flux'].mean())
co2flux_alk_year.append(co2flux_alk_month['B_C_DIC _flux'].mean())
co2flux_no_year = np.array(co2flux_no_year)
co2flux_alk_year = np.array(co2flux_alk_year)
year_days = np.array([31, 28, 31, 30, 31, 30, 31, 31, 30, 31, 30, 31])
co2flux_no_monthly = co2flux_no_year*year_days/1000
co2flux_alk_monthly = co2flux_alk_year*year_days/1000
dates = pd.date_range('2011-01-01', '2012-01-01', freq='M')
fig = plt.figure(figsize=(10, 3))
ax = fig.add_subplot(1, 1, 1)
ax.plot(dates, co2flux_no_monthly, linewidth=2,
label=r'with wind speed 5 m sec$^{-1}$')
ax.plot(dates, co2flux_alk_monthly,
linewidth=2, label=r'with wind speed 15 m sec$^{-1}$')
ax.set_title('Monthly air-sea CO$_2$ flux [mol m$^{-2}$ month$^{-1}$]; positive means inwards')
ax.legend(loc='best');
# *Show daily CO$_2$ flux*
co2flux_no = co2flux_no.reset_index()
co2flux_alk = co2flux_alk.reset_index()
fig = plt.figure(figsize=(10, 3))
ax = fig.add_subplot(1, 1, 1)
ax.plot(co2flux_no['time'], co2flux_no['B_C_DIC _flux'], linewidth=2,
label=r'with wind speed 5 m sec$^{-1}$')
ax.plot(co2flux_alk['time'], co2flux_alk['B_C_DIC _flux'],
linewidth=2, label=r'with wind speed 15 m sec$^{-1}$')
plt.title('Daily air-sea CO$_2$ flux [mmol m$^{-2}$ day$^{-1}$]; positive means inwards')
ax.legend(loc='best');
co2flux_no['B_C_DIC _flux'].values.sum()
co2flux_alk['B_C_DIC _flux'].values.sum() # CO2 [mmol m-2 year-1] seawater excreets
| s_10_different_wind_speeds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Process UniProt Data
#
# Jupyter Notebook to download and preprocess files to transform to BioLink RDF.
#
# ### Download files
#
# The download can be defined:
# * in this Jupyter Notebook using Python
# * as a Bash script in the `download/download.sh` file, and executed using `d2s download uniprot`
#
#
# +
import os
import glob
import requests
import functools
import shutil
import pandas as pd
# Use Pandas, load file in memory
def convert_tsv_to_csv(tsv_file):
csv_table=pd.read_table(tsv_file,sep='\t')
csv_table.to_csv(tsv_file[:-4] + '.csv',index=False)
# Variables and path for the dataset
dataset_id = 'uniprot'
dsri_flink_pod_id = 'flink-jobmanager-###'
input_folder = '/notebooks/workspace/input/' + dataset_id
mapping_folder = '/notebooks/datasets/' + dataset_id + '/mapping'
os.makedirs(input_folder, exist_ok=True)
# +
# Use input folder as working folder
os.chdir(input_folder)
files_to_download = [
'https://raw.githubusercontent.com/MaastrichtU-IDS/d2s-scripts-repository/master/resources/cohd-sample/concepts.tsv'
]
# Download each file and uncompress them if needed
# Use Bash because faster and more reliable than Python
for download_url in files_to_download:
os.system('wget -N ' + download_url)
os.system('find . -name "*.tar.gz" -exec tar -xzvf {} \;')
os.system('unzip -o \*.zip')
# Rename .txt to .tsv
listing = glob.glob('*.txt')
for filename in listing:
os.rename(filename, filename[:-4] + '.tsv')
## Convert TSV to CSV to be processed with the RMLStreamer
# use Pandas (load in memory)
convert_tsv_to_csv('concepts.tsv')
# Use Bash
# cmd_convert_csv = """sed -e 's/"/\\"/g' -e 's/\t/","/g' -e 's/^/"/' -e 's/$/"/' -e 's/\r//' concepts.tsv > concepts.csv"""
# os.system(cmd_convert_csv)
# -
# ## Process and load concepts
#
# We will use CWL workflows to integrate data with SPARQL queries. The structured data is first converted to a generic RDF based on the data structure, then mapped to BioLink using SPARQL. The SPARQL queries are defined in `.rq` files and can be [accessed on GitHub](https://github.com/MaastrichtU-IDS/d2s-project-template/tree/master/datasets/uniprot/mapping).
#
# Start the required services (here on our server, defined by the `-d trek` arg):
#
# ```bash
# d2s start tmp-virtuoso drill -d trek
# ```
#
# Run one of the following d2s command in the d2s-project folder:
#
# ```bash
# d2s run csv-virtuoso.cwl uniprot
# d2s run xml-virtuoso.cwl uniprot
# ```
#
# [HCLS metadata](https://www.w3.org/TR/hcls-dataset/) can be computed for the uniprot graph:
#
# ```bash
# d2s run compute-hcls-metadata.cwl uniprot
# ```
#
# ## Load the BioLink model
#
# Load the [BioLink model ontology as Turtle](https://github.com/biolink/biolink-model/blob/master/biolink-model.ttl) in the graph `https://w3id.org/biolink/biolink-model` in the triplestore
#
| archived-datasets/uniprot/process-uniprot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false
# # Assignment 1
# Welcome to the first programming assigment for the course. This assignments will help to familiarise you with qiskit while revisiting the topics discussed in this week's lectures.
#
# ### Submission Guidelines
# For final submission, and to ensure that you have no errors in your solution, please use the 'Restart Kernel and run all cells' option availble in the Kernel menu at the top of the page.
# To submit your solution, run the completed notebook and then copy the two strings at the bottom of the notebook (which will be generated from your answers) and paste them in the google form associated with the assignment. In addition to this, please also attach the solved notebook as a file using the 'Add or Create' option under the 'Your Work' heading on the assignment page.
# <div class="alert alert-block alert-info"><b>Instructions:</b> Only some of the cells in this notebook are editable. These are the only ones you need to edit to fill your submission. You can run all cells</div>
# + deletable=false editable=false
# %matplotlib inline
import hashlib
import numpy as np
import matplotlib.pyplot as plt
# Importing standard Qiskit libraries and configuring account
from qiskit import QuantumCircuit, execute
from qiskit.circuit import Parameter
from qiskit.providers.aer import QasmSimulator
from qiskit.visualization import *
# + [markdown] deletable=false editable=false
# ## Rotations on the Bloch sphere
# A general single-qubit state on the Bloch sphere is denoted by the statevector $$ |\psi\rangle = \cos{(\tfrac{\theta}{2})}|0\rangle + e^{i\phi}\sin{\tfrac{\theta}{2}}|1\rangle $$
# We have seen the phase-shift operation $R_{\phi}$ corresponding to rotation by some angle $\phi$ about the $z$-axis. The statevector on the Bloch sphere traces a horizontal circle (a line of latitude) by $\phi$ radians. This operation has the matrix representation
# $$
# R_{\phi} =
# \begin{pmatrix}
# 1 & 0 \\
# 0 & e^{i\phi}
# \end{pmatrix}
# $$
# This operation changes the relative phase of the statevector, hence the name.
# It stands to reason that there are operations which rotate a vector about other axes too. We have already seen that the phase-shift gate is a generalisation of the Pauli $Z$ gate. So it is reasonable to think that rotation about the $x$-axis and $y$-axis might be related to the Pauli $X$ and $Y$ operations. This is indeed the case. In general a rotation about a Pauli axis (these are synonymous with the $x$, $y$ and $z$ axes for our purposes) is represented by $$R_{P}(\theta) = \exp(-i\theta P/2) = \cos(\theta/2)I -i \sin(\theta/2)P$$
#
# For the purposes of this assigment
#
# $$
# R_x(\theta) =
# \begin{pmatrix}
# \cos(\theta/2) & -i\sin(\theta/2)\\
# -i\sin(\theta/2) & \cos(\theta/2)
# \end{pmatrix}
# $$
# $$
# R_y(\theta) =
# \begin{pmatrix}
# \cos(\theta/2) & - \sin(\theta/2)\\
# \sin(\theta/2) & \cos(\theta/2).
# \end{pmatrix}
# $$
# $$
# R_z(\phi) =
# \begin{pmatrix}
# e^{-i \phi/2} & 0 \\
# 0 & e^{i \phi/2}
# \end{pmatrix}
# $$
#
# Note that here we have used an equivalent as it is different to $R_{\phi}$ by a global phase $e^{-i \phi/2}$.
#
# If we start in the $|0\rangle$ state, as all quantum circuits in qiskit do, we can get to the general state $|\psi\rangle$ by first performing a rotation about the $y$-axis by an angle $\theta$, followed by a rotation about the $z$-axis by an angle $\phi$.
#
# $$ |\psi\rangle = R_z(\phi) R_y(\theta) |0\rangle$$
#
# You can find a summary of all the gates available in qiskit [here](https://qiskit.org/documentation/tutorials/circuits/3_summary_of_quantum_operations.html#Single-Qubit-Gates). For this assignment, you are only allowed to use these gates.
#
# ## Measurement in different bases
# Qiskit allows measurement only in the computational basis directly. However, we can also perform meaurements in other bases.
# For example, consider the Hadamard basis $\{|+\rangle, |-\rangle\}$. When we measure in this basis, we get an outcome that is one of the two basis vectors. This can be done by projecting a general state $|\psi\rangle$ onto each of the basis states $|+\rangle$ and $|-\rangle$. The probability of obtaining the $+$ outcome is $ P_{+}(|\psi\rangle) = |\langle +|\psi\rangle |^{2}$ and similarly for $-$. We note here that $|+\rangle = H|0\rangle$. Taking the adoint of this equation, we get $\langle +| = \langle 0|H^{\dagger} = \langle 0|H$ where we have used the fact that the Hadamard transformation is equal to it's adjoint (easily verified from the matrix representation). So we can write $ P_{+}(|\psi\rangle) = |\langle +|\psi\rangle |^{2} = |\langle 0| H\psi\rangle |^{2}$. It seems that the probability of measuring the $+$ outcome in the Hadamard basis is the same as the probability of measuring $0$ in the computational basis after applying the Hadamard transformation to the statevector $|\psi\rangle$. So, if we want to measure in the Hadamard basis, we need only to add a Hadamard gate to our qubit and measure in the computational basis.
#
# ## **Problem 1**
# Prepare the state $|i\rangle$ in a quantum circuit and measure it in the Hadamard basis $\{ |+\rangle, |-\rangle \}$
# Below we have provided you with some code to create a quantum circuit. Add appropriate gates to prepare the $|i\rangle$ state and then add gates and a measurement to measure it in the Hadamard basis.
# +
qc1 = QuantumCircuit(1)
# Insert gates below to create the state
# Insert the necessary gates to change to the Hadamard basis below and measure
# Do not change below this line
qc1.draw('mpl')
# + [markdown] deletable=false editable=false
# <div class="alert alert-block alert-info"><b>Instructions:</b>Once your circuit is ready, run the cell below to save your answer. You can change your answer by running these two cells in order again. </div>
# + deletable=false editable=false
basis_gates = ['id', 'x', 'y', 'z', 's', 't', 'sdg', 'tdg', 'h', 'p', 'sx' ,'r', 'rx', 'ry', 'rz', 'u', 'u1', 'u2', 'u3', 'barrier', 'measure']
assert list(qc1.count_ops()) != [], "Circuit cannot be empty"
assert set(qc1.count_ops().keys()).intersection(basis_gates) != set(), "Only basic gates are allowed"
job1 = execute(qc1, backend=QasmSimulator(), shots=1024, seed_simulator=0)
counts1 = job1.result().get_counts()
answer1 = hashlib.sha256(str(counts1).encode()).hexdigest()
plot_histogram(counts1)
# + [markdown] deletable=false editable=false
# ## Simulating polarisation of light on a quantum computer
# In this exercise, we will use quantum computing to simulate a toy model of photon polarisation.
#
# For a brief refresher on polarisation, and a demonstration of the experiment we will be simulating, watch the segment of the YouTube video below.
#
# Reference:
# <NAME>. "Polarization of Electromagnetic Waves" _YouTube_, Jun 20, 2019 https://www.youtube.com/watch?v=6N3bJ7Uxpp0
# + deletable=false editable=false
from IPython.display import YouTubeVideo, display
polariser_exp = YouTubeVideo('6N3bJ7Uxpp0', end=93, height=405, width=720, modestbranding=1)
display(polariser_exp)
# + [markdown] deletable=false editable=false
# To simulate the action of a polariser on plane-polarised light, we have to somehow map the problem onto the Bloch sphere. Here is one way to do it. As shown in the video, the light exiting the first polariser is plane-polarised, which means it is polarised in a plane at some angle to the horizontal and vertical axes. After that, the second polariser essentially _projects_ this onto the pass-axis orientation of the polariser, which can be seen as a kind of measurement. So in our quantum circuit, we will consider only this part of the experiment, and see what fraction of the plane polarised light is transmitted through the second polariser as a function of their relative orientation.
#
# To do this, we need to consider only plane-polarised light. Let us assume that the light exiting the first polarizer is horizontally polarised. We know that if we place the second polariser with it's pass-axis vertical, no light will be transmitted. So these two states are orthogonal. Let us map these two states to $|0\rangle$ and $|1\rangle$ respectively (since these are also orthogonal). However, note that while the angle between the two orthogonal polarisation states is $\pi/2$, the angle on the Bloch sphere between $|0\rangle$ and $|1\rangle$ is $\pi$. We know that any other orientation of linearly polarised light can be written as a superposition of these two basis vectors. Moreover, since we are only considering linearly polarised light, we know that the relative phase is $0$. So we can restrict ourselves to the great circle on the Bloch sphere passing through $|0\rangle$, $|+\rangle$, $|1\rangle$ and $|-\rangle$.
# ## **Problem 2**
# For this experiment, we will need to create a parameterised circuit, where the rotation angle is a parameter. Qiskit enables this using `Parameter()`. We will define a parameter $\beta$ which is the relative angle between the two polarisers. Given below is a quantum circuit where the qubit starts in the $|0\rangle$ state, the equivalent of horizontally plane polarised light. Change the measurement basis using a rotation gate with an angle of rotation in terms of the variable `beta` and measure in this basis. This basis change should correspond to rotating the second polariser by an angle $\beta$. We take $\beta$ to be positive in the counter-clockwise direction.
# +
beta = Parameter('β')
qc2 = QuantumCircuit(1)
# Enter your code below this line
# Do not change below this line
qc2.draw(output='mpl')
# + [markdown] deletable=false editable=false
# We will rotate the second polariser through an angle of $\pi$. This means that $0\le\beta\le\pi$. We can repeat this experiment for $\beta$ values in this range and count the number of times we measured a photon passing through the second polariser.
# <div class="alert alert-block alert-info"><b>Instructions:</b>When you have done that, run the cell below to perform the experiment</div>
#
# + deletable=false editable=false
beta_range = np.linspace(0, np.pi, 50)
num_shots = 1024
basis_gates = ['id', 'x', 'y', 'z', 's', 't', 'sdg', 'tdg', 'h', 'p', 'sx' ,'r', 'rx', 'ry', 'rz', 'u', 'u1', 'u2', 'u3', 'barrier', 'measure']
assert list(qc2.count_ops()) != [], "Circuit cannot be empty"
assert set(qc2.count_ops().keys()).intersection(basis_gates) != set(), "Only basic gates are allowed"
job2 = execute(qc2,
backend=QasmSimulator(),
shots = num_shots,
parameter_binds=[{beta: beta_val} for beta_val in beta_range],
seed_simulator=0) # For consistent results
counts2 = job2.result().get_counts()
# Calculating the probability of photons passing through
probabilities = list(map(lambda c: c.get('0', 0)/num_shots, counts2))
# + [markdown] deletable=false editable=false
# <div class="alert alert-block alert-info"><b>Instructions:</b>Refer to the lecture slides for the theoretical transmission probability. Fill in that expression in the function below.</div>
# -
def theoretical_prob(beta):
'''
Definition of theoretical transmission probability.
The expression for transmitted probability between two polarisers
with a relative angle `beta` given in radians
'''
# Fill in the correct expression for this probability and assign it to the variable tp below
# You may use numpy function like so: np.func_name()
tp =
return tp
# + [markdown] deletable=false editable=false
# <div class="alert alert-block alert-info"><b>Instructions:</b>When you have defined the expression for the transmission probability, run the cell below to save your answer. You can change your answer by running these two cells in order again. </div>
# + deletable=false editable=false
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
ax.plot(beta_range, probabilities, 'o', label='Experimental')
ax.plot(beta_range, theoretical_prob(beta_range), '-', label='Theoretical')
ax.set_xticks([i * np.pi / 4 for i in range(5)])
ax.set_xticklabels(['0', r'$\frac{\pi}{4}$', r'$\frac{\pi}{2}$', r'$\frac{3\pi}{4}$', r'$\pi$'], fontsize=14)
ax.set_xlabel('θ', fontsize=14)
ax.set_ylabel('Probability of Transmission', fontsize=14)
ax.legend(fontsize=14)
answer2 = hashlib.sha256(str(probabilities).encode()).hexdigest()
# + [markdown] deletable=false editable=false
# ## Final Submission
# <div class="alert alert-block alert-info"><b>Instructions:</b> Run the cell below. Copy the string of characters next to each 'Answer #:' below and paste it into the Google form for the assignment. You should run this cell again if you change your answers</div>
# + deletable=false editable=false
print(f'Answer 1: {answer1}')
print(f'Answer 2: {answer2}')
| assignments/assignment1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Federated Audio Classification tutorial with 🤗 Transformers
# !pip install "datasets==1.14" "transformers==4.11.3" "librosa" "torch" "ipywidgets" "numpy==1.21.5"
# # Connect to the Federation
# +
from openfl.interface.interactive_api.federation import Federation
client_id = "frontend"
director_node_fqdn = "localhost"
director_port = 50050
federation = Federation(
client_id=client_id,
director_node_fqdn=director_node_fqdn,
director_port=director_port,
tls=False,
)
# -
shard_registry = federation.get_shard_registry()
shard_registry
federation.target_shape
# ## Creating a FL experiment using Interactive API
from openfl.interface.interactive_api.experiment import (
DataInterface,
FLExperiment,
ModelInterface,
TaskInterface,
)
# ### Register dataset
import datasets
import numpy as np
import torch
from torch.utils.data import Dataset
from transformers import (
AutoFeatureExtractor,
AutoModelForAudioClassification,
Trainer,
TrainingArguments,
)
# +
model_checkpoint = "facebook/wav2vec2-base"
labels = [
"yes",
"no",
"up",
"down",
"left",
"right",
"on",
"off",
"stop",
"go",
"_silence_",
"_unknown_",
]
label2id, id2label = dict(), dict()
for i, label in enumerate(labels):
label2id[label] = str(i)
id2label[str(i)] = label
# +
feature_extractor = AutoFeatureExtractor.from_pretrained(model_checkpoint)
max_duration = 1.0
def preprocess_function(pre_processed_data):
audio_arrays = pre_processed_data
inputs = feature_extractor(
audio_arrays,
sampling_rate=feature_extractor.sampling_rate,
max_length=int(feature_extractor.sampling_rate * max_duration),
truncation=True,
)
return inputs
# + tags=[]
class SuperbShardDataset(Dataset):
def __init__(self, dataset):
self._dataset = dataset
def __getitem__(self, index):
x, y = self._dataset[index]
x = preprocess_function(x)
return {"input_values": x["input_values"][0], "labels": y}
def __len__(self):
return len(self._dataset)
class SuperbFedDataset(DataInterface):
def __init__(self, **kwargs):
super().__init__(**kwargs)
@property
def shard_descriptor(self):
return self._shard_descriptor
@shard_descriptor.setter
def shard_descriptor(self, shard_descriptor):
"""
Describe per-collaborator procedures for sharding.
This method will be called during a collaborator initialization.
Local shard_descriptor will be set by Envoy.
"""
self._shard_descriptor = shard_descriptor
self.train_set = SuperbShardDataset(
self._shard_descriptor.get_dataset("train"),
)
self.valid_set = SuperbShardDataset(
self._shard_descriptor.get_dataset("val"),
)
self.test_set = SuperbShardDataset(
self._shard_descriptor.get_dataset("test"),
)
def __getitem__(self, index):
return self.shard_descriptor[index]
def __len__(self):
return len(self.shard_descriptor)
def get_train_loader(self):
return self.train_set
def get_valid_loader(self):
return self.valid_set
def get_train_data_size(self):
return len(self.train_set)
def get_valid_data_size(self):
return len(self.valid_set)
# -
fed_dataset = SuperbFedDataset()
# ### Describe a model and optimizer
# +
"""
Download the pretrained model and fine-tune it. For classification we use the AutoModelForAudioClassification class.
"""
num_labels = len(id2label)
model = AutoModelForAudioClassification.from_pretrained(
model_checkpoint,
num_labels=num_labels,
label2id=label2id,
id2label=id2label,
)
# +
from transformers import AdamW
params_to_update = []
for param in model.parameters():
if param.requires_grad == True:
params_to_update.append(param)
optimizer = AdamW(params_to_update, lr=3e-5)
# -
# #### Register model
framework_adapter = (
"openfl.plugins.frameworks_adapters.pytorch_adapter.FrameworkAdapterPlugin"
)
MI = ModelInterface(
model=model, optimizer=optimizer, framework_plugin=framework_adapter
)
# ### Define and register FL tasks
batch_size = 16
args = TrainingArguments(
"finetuned_model",
save_strategy="epoch",
per_device_train_batch_size=batch_size,
per_device_eval_batch_size=batch_size,
num_train_epochs=1,
warmup_ratio=0.1,
logging_steps=10,
push_to_hub=False,
)
# +
from datasets import load_metric
metric = load_metric("accuracy")
def compute_metrics(eval_pred):
"""Computes accuracy on a batch of predictions"""
predictions = np.argmax(eval_pred.predictions, axis=1)
return metric.compute(predictions=predictions, references=eval_pred.label_ids)
# +
TI = TaskInterface()
import torch.nn as nn
import tqdm
@TI.register_fl_task(
model="model", data_loader="train_loader", device="device", optimizer="optimizer"
)
def train(model, train_loader, optimizer, device):
print(f"\n\n TASK TRAIN GOT DEVICE {device}\n\n")
trainer = Trainer(
model.to(device),
args,
train_dataset=train_loader,
tokenizer=feature_extractor,
optimizers=(optimizer, None),
compute_metrics=compute_metrics,
)
train_metrics = trainer.train()
return {"train_loss": train_metrics.metrics["train_loss"]}
@TI.register_fl_task(model="model", data_loader="val_loader", device="device")
def validate(model, val_loader, device):
print(f"\n\n TASK VALIDATE GOT DEVICE {device}\n\n")
trainer = Trainer(
model.to(device),
args,
eval_dataset=val_loader,
tokenizer=feature_extractor,
compute_metrics=compute_metrics,
)
eval_metrics = trainer.evaluate()
return {"eval_accuracy": eval_metrics["eval_accuracy"]}
# -
# ## Time to start a federated learning experiment
experiment_name = "HF_audio_test_experiment"
fl_experiment = FLExperiment(federation=federation, experiment_name=experiment_name)
fl_experiment.start(
model_provider=MI,
task_keeper=TI,
data_loader=fed_dataset,
rounds_to_train=2,
opt_treatment="CONTINUE_GLOBAL",
device_assignment_policy="CUDA_PREFERRED",
)
fl_experiment.stream_metrics()
| openfl-tutorials/interactive_api/PyTorch_Huggingface_transformers_SUPERB/workspace/PyTorch_Huggingface_transformers_SUPERB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''project_env'': venv)'
# name: python3
# ---
# Race-Bar Chart: Completed
# 2021-05-25 <br/>
# Mario Kart World Records <br/>
# https://github.com/rfordatascience/tidytuesday/blob/master/data/2021/2021-05-25/readme.md
# ### Learning Links
#
# https://towardsdatascience.com/creating-bar-chart-race-animation-with-python-cdb01144074e
#
import pandas as pd
df= pd.read_csv('https://raw.githubusercontent.com/rfordatascience/tidytuesday/master/data/2021/2021-05-25/records.csv')
df.head()
# +
# Skipping Cleaning
# -
df['date']= pd.to_datetime(df['date'])
df.groupby([df['date'].dt.year, df['date'].dt.month,df['player']]).agg({'time':min}).unstack()
a=df.groupby([df['date'].dt.year,'player'])['time'].min().unstack()
a.head()
import bar_chart_race as bcr
bcr.bar_chart_race(df=a, filename='a2.mp4')
bcr.bar_chart_race(df = a,
n_bars = 6,
sort='asc',
title='Premier League Clubs Points Since 1992',
filename = 'pl_clubs.mp4')
def bar_race(df, title):
bcr.bar_chart_race(
df=a,
filename=title+'.mp4',
orientation='h',
sort='asc',
n_bars=6,
fixed_order=False,
fixed_max=False,
steps_per_period=1,
period_label={'x': .99, 'y': .25, 'ha': 'right', 'va': 'center'},
period_fmt= None,
# period_summary_func=lambda v, r: {'x': .99, 'y': .18,
# 's': f'Fastest times: {v.nsmallest(6).min():,.0f}',
# 'ha': 'right', 'size': 8, 'family': 'Courier New'},
perpendicular_bar_func='median',
period_length=800,
figsize=(5, 8),
dpi=144,
cmap='dark12',
title=title,
title_size='',
bar_label_size=7,
tick_label_size=7,
shared_fontdict={'family' : 'Helvetica', 'color' : '.1'},
scale='linear',
writer='ffmpeg',
fig=None,
bar_kwargs={'alpha': .7},
filter_column_colors=True)
pass
bar_race(a,'Fastest time by player (Any Type)')
b= df[df["type"]=="Three Lap"].groupby(['date','player'])['time'].min().unstack()
bar_race(b,'Fastest time by player (Three Laps)')
c= df[df["type"]=="One Lap"].groupby(['date','player'])['time'].min().unstack()
bar_race(c,'Fastest time by player (One Lap)')
# ## This displays a race Bar chart of the top speed-runners over time.
| Tidy-Tuesday-Insights/mario.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Code to prepare data for use in an OWG
# ## Resize images and write to a seperate folder
import os
import shutil
from PIL import Image
import numpy as np
from distutils.dir_util import copy_tree
import matplotlib.pyplot as plt
import time
def estimate_sharpness(img):
"""
Estimate image sharpness
Input:
img - np.ndarray representing image
img read as skimage.io.imread( 'imagefilename.jpg' )
Returns:
s - sharpness estimates
adapted from https://stackoverflow.com/questions/6646371/detect-which-image-is-sharper
"""
gy, gx = np.gradient(img)
gnorm = np.sqrt(gx**2 + gy**2)
sharpness = np.average(gnorm)
return sharpness
# edited to remove the part where they convert to grayscale as my images already are grayscale. Removed contrast because I do not need it
# create new folder
newpath = "D:/Webcam Images/MI City/2017_imgprep"
#os.mkdir(newpath)
fromDirectory = "D:/Webcam Images/MI City/2017"
toDirectory = "D:/Webcam Images/MI City/2017_imgprep"
shutil.copy_tree(fromDirectory, toDirectory)
# create seperate folders for different views
view1path = "D:/Webcam Images/MI City/2017_imgprep/view1"
view2path = "D:/Webcam Images/MI City/2017_imgprep/view2"
os.mkdir(view1path)
os.mkdir(view2path)
# move images to correct folders and remove the view identifier
for filename in os.listdir(newpath):
if filename.endswith(".01.jpg"):
os.rename("D:/Webcam Images/MI City/2017_imgprep/{}".format(filename),"D:/Webcam Images/MI City/2017_imgprep/view1/{}".format(filename.replace(".01","")))
elif filename.endswith(".02.jpg"):
os.rename("D:/Webcam Images/MI City/2017_imgprep/{}".format(filename),"D:/Webcam Images/MI City/2017_imgprep/view2/{}".format(filename.replace(".02","")))
# +
# crop images to square, convert to grayscale, and reduce resolution to 128x128
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image
image = Image.open("C:/njc/src/mcyimgs/2019/20191011_1233.02.jpg")
bw = image.convert("L")
bwarray = np.array(bw)
print(len(np.shape(bwarray)))
arrayshape = np.index_exp[270:720, 830:1280]
small = bwarray[arrayshape]
print(np.shape(small))
plt.imshow(small, cmap = "gray")
print(np.shape(small))
# -
# ## Make practice code into functions
year = 2018
directory = "D:/Webcam Images/MI City"
newdir = directory+"/{}_imgprep".format(str(year))
print (newdir+"/view1")
# #### Note:
#
# ##### Functionality
# The function below is specifically for the analysis of view \#2 in MI City. This works with imagery from the old webcam location. It is designed for a proof of concept application of the OWG. This function will need to be adapted or rewritten to deal with contemporary datsets. Also, a record of the inputs to each version should be kept for research/repeatability purposes.
#
# ##### Next Steps
# The most important additon that is needed is a way to automate the creation of view folders within the new year_imgprep folder. This could be done by including a list of numbers as strings representing the camera view as an input, or using string parsing and appending the target string-numbers into a list within the function. The latter option is likely best as it requires no additonal inputs be added to the funciton
m = [1,2,3,4]
np.mean(m)
"OWG" == "OWG"
# +
y =[10]
for y in y:
x =5
three = [3]
for three in three:
print (x)
# -
# ## Function version 1.5
def prepowgimgs(year, directory, arrayshape, resolution, version):
'''Create a new directory of images and prepare them for further data filtering. version 1.5
Year is the target year of images,
directory is where the year files are stored,
resolution is the resolutiont the images will be reduced to,
version is the version that these inputs are associate with. A record of this should be kept by hand.
'''
#set counters
counter = 0
nightcounter = 0
blurcounter = 0
failcounter = 0
# create a new directory with the called year_imgprep
print("Creating folder")
foldername = "/{}_imgprep".format(str(year))
newdir = directory+foldername+"{}".format(version)
try:
shutil.rmtree(newdir)
print("Folder found: deleting contents")
os.mkdir(newdir)
except:
os.mkdir(newdir)
# copy files into new directory, this can take a while
print("Moving Files")
copy_tree(directory+"/{}".format(str(year)), newdir)
# seperate the views into different folders within the directory
print("Seperating Views")
view1path = newdir+"/view1"
view2path = newdir+"/view2"
os.mkdir(view1path)
os.mkdir(view2path)
for filename in os.listdir(newdir):
if filename.endswith(".01.jpg"):
os.rename(newdir+"/{}".format(filename), view1path+"/{}".format(filename.replace(".01","")))
elif filename.endswith(".02.jpg"):
os.rename(newdir+"/{}".format(filename), view2path+"/{}".format(filename.replace(".02","")))
# edit images from desired view into cropped lower resolution grayscale and store in new folder
print ("Editing images for analysis")
for filename in os.listdir(view2path):
if filename.endswith(".jpg"):
image = Image.open(view2path+"/{}".format(filename))
#convert image to grayscale
bw = image.convert("L")
# select target portion of image to make it square
bwarray = np.array(bw)
small = bwarray[arrayshape]
small_img = Image.fromarray(small)
# change image resoltuion
resize = (resolution, resolution)
smaller = small_img.resize(resize)
# Average image intensity
avgintensity = np.mean(plt.imread(smaller))
# Image Sharpness
sharpness, contrast = estimate_sharpness(plt.imread(smaller))
# if avg image intensity is below 40 (meaning it was taken at night) toss out image
if avgintensity < 40:
nightcounter = nightcounter + 1
# if image sharpness is less than 3.5 (the mean of sharpness for all images in 2017) toss out imaage
elif sharpness < 3.5:
blurcounter = blurcounter + 1
else:
# save images ready for interpolation and filtering as the same name sans underscore seperating date and time
try:
smaller = smaller.save(view2path+"/"+filename)
except IOError:
print("cannot create image for", filename)
failcounter = failcounter + 1
# remove the underscore in the filename
os.rename(view2path+"/"+filename, view2path+"/"+filename.replace("_",""))
print (failcounter, "files in folder could not be processed: IOError")
print(blurcounter, "images removed from training dataset due to insufficient sharpness score")
print(nightcounter, "images removed from training dataset due to inusfficeint intensity score")
print (nightcounter+blurcounter+failcounter, "total images removed from training dataset")
print (counter, "images moved to OWG training folder")
return print ("Completed.")
# ## Function version 2.0
def prepowgimgs(year, directory, arrayshape, resolution, trial, viewlist):
'''Create a new directory for images in each view and run QC and primary augmentation before interpolation and then model
training, validation, or testing. version 2.0
Year is the target year of images,
directory is where the year files are stored,
resolution is the resolutiont the images will be reduced to,
trial is the name of the trial that these inputs are associate with. A record of trial details should be kept by hand and stored
in C:/njc/src/SSF/OWG/OWG_records/trial.
viewlist is a list of the view numbers found in the target year folder
'''
start = time.time()
#set counters
counter = 0
nightcounter = 0
blurcounter = 0
failcounter = 0
# create a new directory with the called year_imgprep
foldername = "/{}_imgprep".format(str(year))
newdir = directory+foldername+"{}".format(trial)
try:
shutil.rmtree(newdir)
print("Folder found: deleting contents")
print("Creating folder")
os.mkdir(newdir)
except:
print("Creating folder")
os.mkdir(newdir)
# copy files into new directory, this can take a while
print("Moving Files")
copy_tree(directory+"/{}".format(str(year)), newdir)
# seperate the views into different folders within the directory
viewdirs = []
for view in viewlist:
print("Seperating View {}".format(view))
viewpath = newdir+"/view{}".format(view)
viewdirs.append(viewpath)
os.mkdir(viewpath)
for viewpath in viewdirs:
print ("Moving images to {}".format(viewpath))
# move images into views
for filename in os.listdir(newdir):
if filename[-5] == viewpath[-1]:
os.rename(newdir+"/{}".format(filename), viewpath+"/{}".format(filename.replace(filename[13:16],"")))
# edit images from desired view into cropped lower resolution grayscale and store in new folder
print ("Augmenting and filtering imgaes in {}".format(viewpath))
for filename in os.listdir(viewpath):
if filename.endswith(".jpg"):
image = Image.open(viewpath+"/{}".format(filename)).convert("L")
# select target portion of image to make it square
bwarray = np.array(image)
small = bwarray[arrayshape]
# change image resoltuion
img = Image.fromarray(small)
resize = (resolution, resolution)
smaller = img.resize(resize)
#Get QC metrics
sharpness = estimate_sharpness(np.asarray(smaller))
avgintensity = np.mean(np.asarray(smaller))
# perform QC
if sharpness < 3.5 or avgintensity < 85:
os.remove(viewpath+"/"+filename)
failcounter = failcounter + 1
else:
# save images to OWG directory
try:
counter = counter + 1
owgimage = smaller.save(viewpath+"/{}".format(filename))
except IOError:
print("cannot create image for", filename)
failcounter = failcounter + 1
# remove the underscore in the filename
os.rename(viewpath+"/"+filename, viewpath+"/"+filename.replace("_",""))
print (failcounter, "images removed for quality control")
print (counter, "images processed")
end = time.time()
return print ("Completed in {} minutes.".format((end - start)/60))
# #### Test function version 2.0
year = 2017
directory = "C:/njc/src/mcyimgs"
arrayshape = np.index_exp[100:700, 680:1280]
resolution = 512
trial = 'test'
viewlist = [1,2]
prepowgimgs(year, directory, arrayshape, resolution, trial, viewlist)
photo = "C:/njc/src/mcyimgs/2017_imgpreptest/view2/201712271234.jpg"
pic = plt.imread(photo)
print (estimate_sharpness(pic))
print(np.mean(pic))
x = plt.imread("C:/njc/src/mcyimgs/2017_imgpreptest/view2/201712270033.jpg")
y = np.asarray(x)
z = np.mean(y)
print (z)
# ## Move October images into a seperate folder for first analysis
# +
import os
from shutil import copyfile
directory = "D:/Webcam Images/MI City/2017_imgprep/view2/view2OWGimg"
newdir = directory+"/october2017"
os.mkdir(newdir)
for filename in os.listdir(directory):
if filename[4:6] == "10":
shutil.copyfile("D:/Webcam Images/MI City/2017_imgprep/view2/view2OWGimg"+"/"+filename, newdir+"/"+filename)
# -
# ## Create a new folder of training images at a higher resolution
year = 2017
directory = "C:/njc/src/mcyimgs"
arrayshape = np.index_exp[100:700, 680:1280]
resolution = 256
version = 1
prepmicityimgs(year, directory, arrayshape, resolution, version)
year = 2019
directory = "C:/njc/src/mcyimgs"
arrayshape = np.index_exp[270:720, 830:1280]
resolution = 256
version = "test"
prepmicityimgs(year, directory, arrayshape, resolution, version)
# ### Make a Fall Data Directory
# +
directory = "C:/njc/src/mcyimgs/2019_imgpreptest"
newdir = directory+"/view2fall"
os.mkdir(newdir)
for filename in os.listdir(directory+"/"+"view2"):
if filename[4:6] == "09" or filename[4:6] == "10":
shutil.copyfile(directory+"/"+"view2"+"/"+filename, newdir+"/"+filename)
# -
# ### Select out random images from a year to use as validation data
# csv's generated from functions in csv_interpolation notebook
import pandas as pd
df = pd.read_csv('C:/njc/src/SSF/OWG/mcyv22017trial3.csv', sep = ",", index_col=False)
df
# random sample ~30%
dfs = df.sample(n= int(.3*819))
dfs.reset_index(drop=True, inplace=True)
dfs
dfs.to_csv('C:/njc/src/SSF/OWG/mcyv22017(30trial3).csv', sep = ",")
# ### Drop rows of original dataframe if they have been randomly selected for validation
# this creates the csv file that will be used to train the OWG
# +
# load in the dataframes
total = pd.read_csv('C:/njc/src/SSF/OWG/mcyv22017trial3.csv', sep = ",", index_col=False)
validation = pd.read_csv('C:/njc/src/SSF/OWG/mcyv22017(30trial3).csv', sep = ",", index_col=False)
# if the imagename in the total dataframe exists in the validation dataframe do not include it in the train dataframe.
# the ~ inverts the boolean indexing, which we need to do because only True values will be kept and they start as False as they are not in the validation dataframe
train = total[~total.id.isin(validation.id)]
train
train.to_csv('C:/njc/src/SSF/OWG/mcyv22017trial3train.csv', sep = ",")
| OWG/Prepdata.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import os
import json
import seaborn as sns
# Detectron colors
_COLORS = np.array([
0.000, 0.447, 0.741,
0.850, 0.325, 0.098,
0.929, 0.694, 0.125,
0.494, 0.184, 0.556,
0.466, 0.674, 0.188
]).astype(np.float32).reshape((-1, 3))
# Directory where sweep summaries are stored
_DATA_DIR = '../data'
# -
def compute_norm_ws(cs, num_bins, c_range):
"""Computes normalized EDF weights."""
hist, edges = np.histogram(cs, bins=num_bins, range=c_range)
inds = np.digitize(cs, bins=edges) - 1
assert np.count_nonzero(hist) == num_bins
return 1 / hist[inds] / num_bins
def load_sweep(sweep_name):
"""Loads a sweep summary."""
summary_path = os.path.join(_DATA_DIR, '{}.json'.format(sweep_name))
with open(summary_path, 'r') as f:
sweep_summary = json.load(f)
return sweep_summary
# Source: http://kaiminghe.com/ilsvrc15/ilsvrc2015_deep_residual_learning_kaiminghe.pdf
point_ests = {
'ResNet': 3.57,
'VGG': 7.3,
'ZFNet': 11.7,
'AlexNet': 16.4
}
# Source: https://github.com/facebookresearch/ResNeXt
curve_ests_fs = {
'ResNet': [(8.0, 24), (16.0, 22.5), (23.0, 22.1)],
'ResNeXt': [(8.0, 22.1), (16.0, 21.1), (31.0, 20.4)]
}
sweeps = {
'ResNet-B': load_sweep('ResNet-B'),
'ResNeXt-B': load_sweep('ResNeXt-B')
}
# +
_MIN_P = 0.023
_MAX_P = 0.856
def is_valid_p(job):
return _MIN_P < job['params'] * 1e-6 and job['params'] * 1e-6 < _MAX_P
# +
print('Figure 1\n')
r, c = 1, 3
w, h = 3, 3
fig, axes = plt.subplots(
nrows=r, ncols=c,
figsize=(c * w, r * h),
gridspec_kw={'width_ratios':[w, w, w]}
)
title_font_size = 16
axis_font_size = 15
tick_font_size = 12
legend_font_size = 12.5
##########################################
# Point estimates
##########################################
xs = list(point_ests.keys())[::-1]
ys = list(point_ests.values())[::-1]
ax = axes[0]
ax = sns.barplot(x=xs, y=ys, palette=sns.color_palette("RdBu_r", 6, desat=1.0), ax=ax, alpha=1.0)
ax.set_title('(a) point estimates', fontsize=title_font_size)
ax.set_ylim([0, 17.5])
ax.grid(alpha=0.4)
ax.set_ylabel('error', fontsize=axis_font_size)
ax.tick_params(axis='x', labelsize=tick_font_size, rotation=-20)
##########################################
# Curve estimates
##########################################
ms = ['ResNet', 'ResNeXt']
ax = axes[1]
for i, m in enumerate(ms):
xs = [x for (x, y) in curve_ests_fs[m]]
ys = [y for (x, y) in curve_ests_fs[m]]
ax.plot(
xs, ys, label=m,
color=_COLORS[1 - i], alpha=0.8, linewidth=2.5,
marker='o', markersize=8
)
ax.grid(alpha=0.4)
ax.set_xlabel('complexity', fontsize=axis_font_size)
ax.set_ylabel('error', fontsize=axis_font_size)
ax.legend(loc='upper right', prop={'size' : legend_font_size})
ax.set_title('(b) curve estimates', fontsize=title_font_size)
ax.set_xlim([5, 35])
ax.set_ylim([20, 25])
ax.set_xticks([5, 10, 15, 20, 25, 30, 35], ['5.0', '10.0', '15.0', '20.0', '25.0', '30.0', '35.0'])
ax.set_yticks([20, 21, 22, 23, 24, 25], ['20.0', '21.0', '22.0', '23.0', '24.0', '25.0'])
##########################################
# Distribution estimates
##########################################
dss = ['ResNet-B', 'ResNeXt-B']
lbs = ['ResNet', 'ResNeXt']
ax = axes[2]
for i, ds in enumerate(dss):
errs = np.array([job['min_test_top1'] for job in sweeps[ds] if is_valid_p(job)])
ps = np.array([job['params'] * 1e-6 for job in sweeps[ds] if is_valid_p(job)])
inds = np.argsort(errs)
errs, ps = errs[inds], ps[inds]
ws = compute_norm_ws(ps, num_bins=40, c_range=(_MIN_P, _MAX_P))
assert np.isclose(np.sum(ws), 1.0)
ax.plot(
errs, np.cumsum(ws),
color=_COLORS[1 - i], linewidth=2.5, alpha=0.8, label=lbs[i]
)
ax.grid(alpha=0.4)
ax.set_title('(c) distribution estimates', fontsize=title_font_size)
ax.set_xlabel('error | complexity', fontsize=axis_font_size)
ax.set_ylabel('cumulative prob.', fontsize=axis_font_size)
ax.set_xlim([4.5, 12.5])
ax.legend(loc='lower right', prop={'size' : legend_font_size})
plt.tight_layout();
# -
| notebooks/teaser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
tf.enable_eager_execution()
import os
import time
import numpy as np
import glob
import matplotlib.pyplot as plt
import PIL
import imageio
from IPython import display
# %matplotlib inline
from softlearning.utils.tensorflow import nest
tfk = tf.keras
tfkl = tf.keras.layers
# !export CUDA_VISIBLE_DEVICES=1
# -
from softlearning.models.state_estimation import (
get_dumped_pkl_data
)
images_path = '/home/justinvyu/dev/softlearning-vice/goal_classifier/free_screw_state_estimator_data_invisible_claw/more_data.pkl'
images, _ = get_dumped_pkl_data(images_path)
images.shape
images = None
# +
image_shape = images.shape[1:]
num_images = images.shape[0]
# Shuffle images
np.random.shuffle(images)
# +
validation_split = 0.1
split_index = int(num_images * validation_split)
num_training_examples = 500000
_train_images = images[split_index:split_index + num_training_examples]
# _train_images = images[split_index:]
_test_images = images[:split_index]
train_images = _train_images
test_images = _test_images
# train_images = (_train_images / 255.).astype(np.float32)
# test_images = (_test_images / 255.).astype(np.float32)
# -
train_images = train_images[:200000]
train_images.shape, test_images.shape
train_images.shape, test_images.shape
# +
BATCH_SIZE = 128
def train_generator():
for image in train_images:
yield image
def test_generator():
for image in test_images:
yield image
train_dataset = tf.data.Dataset.from_generator(train_generator, tf.float32).batch(BATCH_SIZE)
test_dataset = tf.data.Dataset.from_generator(test_generator, tf.float32).batch(BATCH_SIZE)
# +
def preprocess(x):
"""Cast to float, normalize, and concatenate images along last axis."""
x = nest.map_structure(
lambda image: tf.image.convert_image_dtype(image, tf.float32), x)
x = nest.flatten(x)
x = tf.concat(x, axis=-1)
# x = (tf.image.convert_image_dtype(x, tf.float32) - 0.5) * 2.0
return x
class CVAE(tf.keras.Model):
def __init__(self, input_shape=(32, 32, 3), latent_dim=64):
super(CVAE, self).__init__()
self.latent_dim = latent_dim
self.encoder = tf.keras.Sequential([
tfkl.InputLayer(input_shape=input_shape),
tfkl.Lambda(preprocess),
tfkl.Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation=tfkl.LeakyReLU()),
tfkl.Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation=tfkl.LeakyReLU()),
tfkl.Conv2D(
filters=32, kernel_size=3, strides=(2, 2), activation=tfkl.LeakyReLU()),
tfkl.Flatten(),
# No activation
tfkl.Dense(latent_dim + latent_dim)])
self.decoder = tf.keras.Sequential([
tfkl.InputLayer(input_shape=(latent_dim,)),
tfkl.Dense(units=4*4*32, activation=tf.nn.relu),
tfkl.Reshape(target_shape=(4, 4, 32)),
tfkl.Conv2DTranspose(
filters=64,
kernel_size=3,
strides=(2, 2),
padding="SAME",
activation=tfkl.LeakyReLU()),
tfkl.Conv2DTranspose(
filters=64,
kernel_size=3,
strides=(2, 2),
padding="SAME",
activation=tfkl.LeakyReLU()),
tfkl.Conv2DTranspose(
filters=32,
kernel_size=3,
strides=(2, 2),
padding="SAME",
activation=tfkl.LeakyReLU()),
# No activation
tfkl.Conv2DTranspose(
filters=3, kernel_size=3, strides=(1, 1), padding="SAME")])
@tf.function
def sample(self, eps=None):
if eps is None:
eps = tf.random.normal(shape=(100, self.latent_dim))
return self.decode(eps, apply_sigmoid=True)
def encode(self, x):
mean, logvar = tf.split(self.encoder(x), num_or_size_splits=2, axis=1)
return mean, logvar
def reparameterize(self, mean, logvar):
eps = tf.random.normal(shape=mean.shape)
return eps * tf.exp(logvar * .5) + mean
def decode(self, z, apply_sigmoid=False):
logits = self.decoder(z)
if apply_sigmoid:
probs = tf.sigmoid(logits)
return probs
return logits
def __call__(self, x):
mean, logvar = self.encode(x)
z = self.reparameterize(mean, logvar)
x_reconstruct = self.decode(z, apply_sigmoid=True)
return x_reconstruct
# +
optimizer = tf.keras.optimizers.Adam(1e-4)
def log_normal_pdf(sample, mean, logvar, raxis=1):
log2pi = tf.math.log(2. * np.pi)
return tf.reduce_sum(
-.5 * ((sample - mean) ** 2. * tf.exp(-logvar) + logvar + log2pi),
axis=raxis)
@tf.function
def compute_loss(model, x):
mean, logvar = model.encode(x)
z = model.reparameterize(mean, logvar)
x_logit = model.decode(z)
cross_ent = tf.nn.sigmoid_cross_entropy_with_logits(
logits=x_logit, labels=x)
logpx_z = -tf.reduce_sum(cross_ent, axis=[1, 2, 3])
logpz = log_normal_pdf(z, 0., 0.)
logqz_x = log_normal_pdf(z, mean, logvar)
return -tf.reduce_mean(logpx_z + logpz - logqz_x)
@tf.function
def compute_apply_gradients(model, x, optimizer):
with tf.GradientTape() as tape:
loss = compute_loss(model, x)
gradients = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
# +
epochs = 500
latent_dim = 64
num_examples_to_generate = 16
# keeping the random vector constant for generation (prediction) so
# it will be easier to see the improvement.
random_vector_for_generation = tf.random.normal(
shape=[num_examples_to_generate, latent_dim])
model = CVAE(input_shape=image_shape, latent_dim=latent_dim)
# -
def generate_and_save_images(model, epoch, test_input):
predictions = model.sample(test_input)
# print(predictions, predictions.shape)
fig = plt.figure(figsize=(4,4))
# print(predictions[0, :, :, :])
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, :])
plt.axis('off')
plt.savefig('/home/justinvyu/dev/softlearning-vice/notebooks/vae_invisible_claw_images/image_at_epoch_{:04d}.png'.format(epoch))
plt.show()
# +
generate_and_save_images(model, 0, random_vector_for_generation)
# with tf.device('/GPU:1'):
for epoch in range(1, epochs + 1):
start_time = time.time()
for train_x in train_dataset:
compute_apply_gradients(model, train_x, optimizer)
end_time = time.time()
if epoch % 1 == 0:
loss = tf.keras.metrics.Mean()
for test_x in test_dataset:
loss(compute_loss(model, test_x))
elbo = -loss.result()
display.clear_output(wait=False)
print('Epoch: {}, Test set ELBO: {}, '
'time elapse for current epoch {}'.format(epoch,
elbo,
end_time - start_time))
generate_and_save_images(
model, epoch, random_vector_for_generation)
# +
encoder = tf.keras.Sequential(
[
tf.keras.layers.InputLayer(input_shape=(64, 64, 3)),
tf.keras.layers.Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation=tf.keras.layers.LeakyReLU()),
tf.keras.layers.Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation=tf.keras.layers.LeakyReLU()),
tf.keras.layers.Conv2D(
filters=32, kernel_size=3, strides=(2, 2), activation=tf.keras.layers.LeakyReLU()),
tf.keras.layers.Flatten(),
# No activation
tf.keras.layers.Dense(latent_dim + latent_dim),
]
)
decoder = tf.keras.Sequential(
[
tf.keras.layers.InputLayer(input_shape=(latent_dim,)),
tf.keras.layers.Dense(units=8*8*32, activation=tf.nn.relu),
tf.keras.layers.Reshape(target_shape=(8, 8, 32)),
tf.keras.layers.Conv2DTranspose(
filters=64,
kernel_size=3,
strides=(2, 2),
padding="SAME",
activation=tf.keras.layers.LeakyReLU()),
tf.keras.layers.Conv2DTranspose(
filters=64,
kernel_size=3,
strides=(2, 2),
padding="SAME",
activation=tf.keras.layers.LeakyReLU()),
tf.keras.layers.Conv2DTranspose(
filters=32,
kernel_size=3,
strides=(2, 2),
padding="SAME",
activation=tf.keras.layers.LeakyReLU()),
# No activation
tf.keras.layers.Conv2DTranspose(
filters=3, kernel_size=3, strides=(1, 1), padding="SAME"),
]
)
# -
encoder.summary()
decoder.summary()
model.encoder.save_weights('inference_weights.h5')
model.decoder.save_weights('generative_weights.h5')
model = CVAE(latent_dim)
model.encoder.load_weights('inference_weights.h5')
model.decoder.load_weights('generative_weights.h5')
test_image = images[1]
test_image = (test_image / 255.).astype(np.float32)
mean, logvar = model.encode(test_image[None, ...])
z = model.reparameterize(mean, logvar)
x_logit = model.decode(z, apply_sigmoid=True)
mean, logvar
z
x_logit
decoded = x_logit.numpy()
plt.imshow(decoded[0])
plt.show()
plt.imshow(test_image)
plt.show()
l2_loss = np.linalg.norm(decoded - test_image)
l2_loss
# +
anim_file = 'cvae.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('vae_images/image*.png')
filenames = sorted(filenames)
last = -1
for i,filename in enumerate(filenames):
frame = 2*(i**0.5)
if round(frame) > round(last):
last = frame
else:
continue
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
import IPython
if IPython.version_info >= (6,2,0,''):
display.Image(filename=anim_file)
# +
def plot_side_by_side(img1, img2, title1='', title2='', figsize=(4, 2)):
fig = plt.figure(figsize=figsize)
plt.subplot(1, 2, 1)
plt.title(title1)
plt.imshow(img1)
plt.axis('off')
plt.subplot(1, 2, 2)
plt.title(title2)
plt.imshow(img2)
plt.axis('off')
plt.show()
plot_side_by_side(test_image, decoded[0], 'Ground Truth', 'VAE Reconstruction')
# -
for image in images[10500:11000]:
image = (image / 255.).astype(np.float32)
reconstruction = model(image[None, ...])
plot_side_by_side(image, reconstruction[0], figsize=(2, 1))
| notebooks/vae/vae.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# In this post,we will talk about some of the most important papers that have been published over the last 5 years and discuss why they’re so important.We will go through different CNN Architectures (LeNet to DenseNet) showcasing the advancements in general network architecture that made these architectures top the ILSVRC results.
# # What is ImageNet
#
# [ImageNet](http://www.image-net.org/)
#
# ImageNet is formally a project aimed at (manually) labeling and categorizing images into almost 22,000 separate object categories for the purpose of computer vision research.
#
# However, when we hear the term “ImageNet” in the context of deep learning and Convolutional Neural Networks, we are likely referring to the ImageNet Large Scale Visual Recognition Challenge, or ILSVRC for short.
#
# The ImageNet project runs an annual software contest, the ImageNet Large Scale Visual Recognition Challenge (ILSVRC), where software programs compete to correctly classify and detect objects and scenes.
#
# The goal of this image classification challenge is to train a model that can correctly classify an input image into 1,000 separate object categories.
#
# Models are trained on ~1.2 million training images with another 50,000 images for validation and 100,000 images for testing.
#
# These 1,000 image categories represent object classes that we encounter in our day-to-day lives, such as species of dogs, cats, various household objects, vehicle types, and much more. You can find the full list of object categories in the ILSVRC challenge
#
# When it comes to image classification, the **ImageNet** challenge is the de facto benchmark for computer vision classification algorithms — and the leaderboard for this challenge has been dominated by Convolutional Neural Networks and deep learning techniques since 2012.
#
# # LeNet-5(1998)
#
# [Gradient Based Learning Applied to Document Recognition](http://yann.lecun.com/exdb/publis/pdf/lecun-01a.pdf)
#
# 1. A pioneering 7-level convolutional network by LeCun that classifies digits,
# 2. Found its application by several banks to recognise hand-written numbers on checks (cheques)
# 3. These numbers were digitized in 32x32 pixel greyscale which acted as an input images.
# 4. The ability to process higher resolution images requires larger and more convolutional layers, so this technique is constrained by the availability of computing resources.
#
# 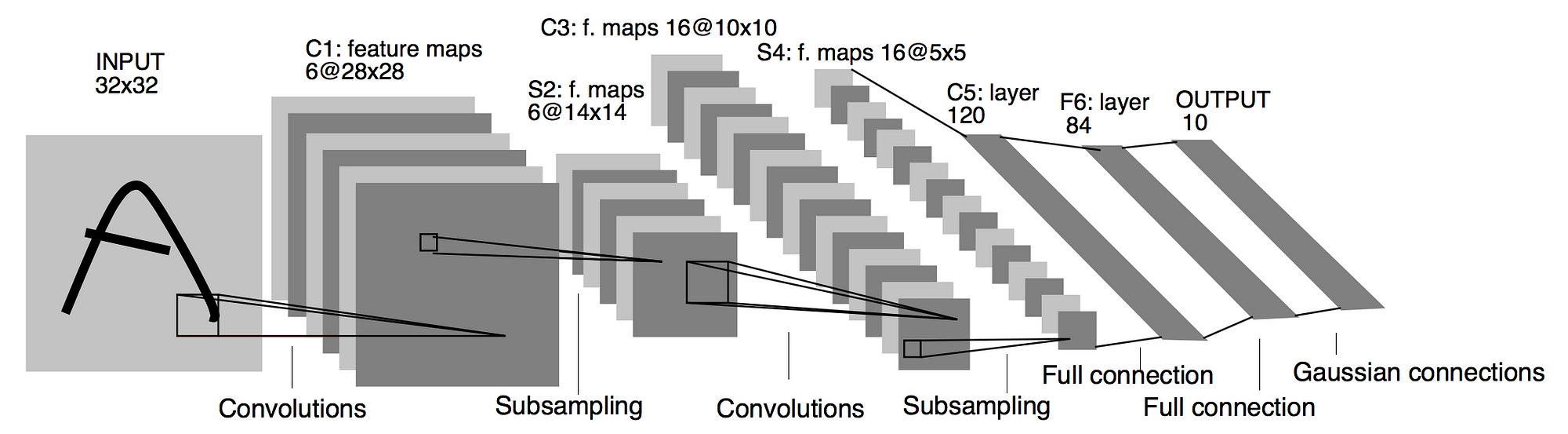
# # AlexNet(2012)
#
# [ImageNet Classification with Deep Convolutional Networks](https://papers.nips.cc/paper/4824-imagenet-classification-with-deep-convolutional-neural-networks.pdf)
#
# 1. One of the most influential publications in the field by <NAME>, <NAME>, and <NAME> that started the revolution of CNN in Computer Vision.This was the first time a model performed so well on a historically difficult ImageNet dataset.
# 2. The network consisted 11x11, 5x5,3x3, convolutions and made up of 5 conv layers, max-pooling layers, dropout layers, and 3 fully connected layers.
# 3. Used ReLU for the nonlinearity functions (Found to decrease training time as ReLUs are several times faster than the conventional tanh function) and used SGD with momentum for training.
# 4. Used data augmentation techniques that consisted of image translations, horizontal reflections, and patch extractions.
# 5. Implemented dropout layers in order to combat the problem of overfitting to the training data.
# 6. Trained the model using batch stochastic gradient descent, with specific values for momentum and weight decay.
# 7. AlexNet was trained for 6 days simultaneously on two Nvidia Geforce GTX 580 GPUs which is the reason for why their network is split into two pipelines.
# 8. AlexNet significantly outperformed all the prior competitors and won the challenge by reducing the top-5 error from 26% to 15.3%
# 
#
# # ZFNet(2013)
#
# [Visualizing and Understanding Convolutional Neural Networks](https://cs.nyu.edu/~fergus/papers/zeilerECCV2014.pdf)
# <br>
# This architecture was more of a fine tuning to the previous AlexNet structure by tweaking the hyper-parameters of AlexNet while maintaining the same structure but still developed some very keys ideas about improving performance.Few minor modifications done were the following:
# 1. AlexNet trained on 15 million images, while ZF Net trained on only 1.3 million images.
# 2. Instead of using 11x11 sized filters in the first layer (which is what AlexNet implemented), ZF Net used filters of size 7x7 and a decreased stride value. The reasoning behind this modification is that a smaller filter size in the first conv layer helps retain a lot of original pixel information in the input volume. A filtering of size 11x11 proved to be skipping a lot of relevant information, especially as this is the first conv layer.
# 3. As the network grows, we also see a rise in the number of filters used.
# 4. Used ReLUs for their activation functions, cross-entropy loss for the error function, and trained using batch stochastic gradient descent.
# 5. Trained on a GTX 580 GPU for twelve days.
# 6. Developed a visualization technique named **Deconvolutional Network**, which helps to examine different feature activations and their relation to the input space. Called **deconvnet** because it maps features to pixels (the opposite of what a convolutional layer does).
# 7. It achieved a top-5 error rate of 14.8%
# 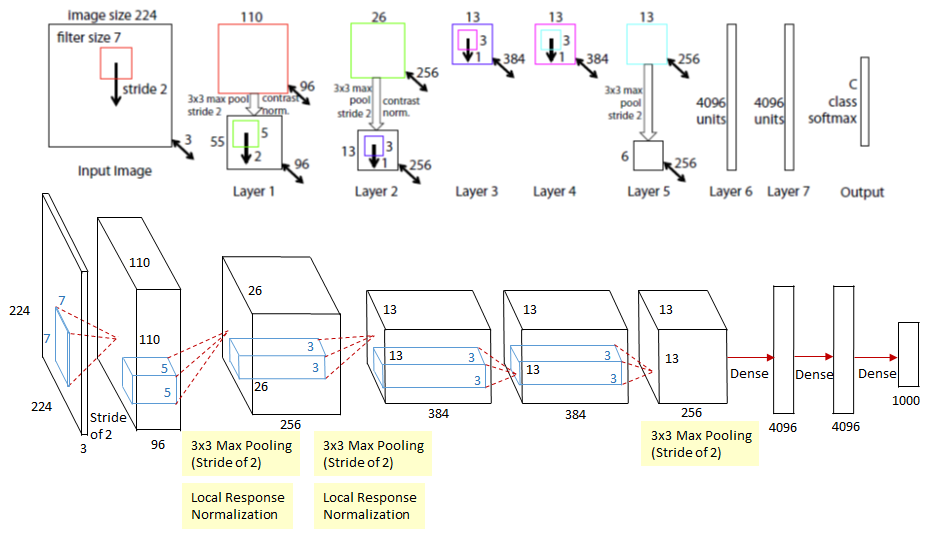
# # VggNet(2014)
#
# [VERY DEEP CONVOLUTIONAL NETWORKS FOR LARGE-SCALE IMAGE RECOGNITION](https://arxiv.org/pdf/1409.1556v6.pdf)
#
# This architecture is well konwn for **Simplicity and depth**.. VGGNet is very appealing because of its very uniform architecture.They proposed 6 different variations of VggNet however 16 layer with all 3x3 convolution produced the best result.
#
# Few things to note:
# 1. The use of only 3x3 sized filters is quite different from AlexNet’s 11x11 filters in the first layer and ZF Net’s 7x7 filters. The authors’ reasoning is that the combination of two 3x3 conv layers has an effective receptive field of 5x5. This in turn simulates a larger filter while keeping the benefits of smaller filter sizes. One of the benefits is a decrease in the number of parameters. Also, with two conv layers, we’re able to use two ReLU layers instead of one.
# 2. 3 conv layers back to back have an effective receptive field of 7x7.
# 3. As the spatial size of the input volumes at each layer decrease (result of the conv and pool layers), the depth of the volumes increase due to the increased number of filters as you go down the network.
# 4. Interesting to notice that the number of filters doubles after each maxpool layer. This reinforces the idea of shrinking spatial dimensions, but growing depth.
# 5. Worked well on both image classification and localization tasks. The authors used a form of localization as regression (see page 10 of the paper for all details).
# 6. Built model with the Caffe toolbox.
# 7. Used scale jittering as one data augmentation technique during training.
# 8. Used ReLU layers after each conv layer and trained with batch gradient descent.
# 9. Trained on 4 Nvidia Titan Black GPUs for two to three weeks.
# 10. It achieved a top-5 error rate of 7.3%
#
# 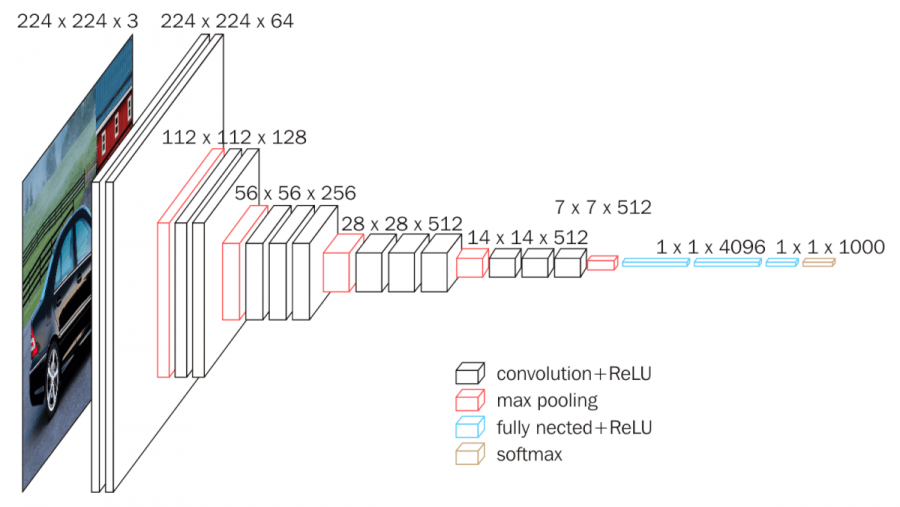
#
#
# 
# **In Standard ConvNet, input image goes through multiple convolution and obtain high-level features.**
# After Inception V1 ,the author proposed a number of upgrades which increased the accuracy and reduced the computational complexity.This lead to many new upgrades resulting in different versions of Inception Network :
# 1. Inception v2
# 2. Inception V3
# # Inception Network (GoogleNet)(2014)
# [Going Deeper with Convolutions](https://www.cv-foundation.org/openaccess/content_cvpr_2015/papers/Szegedy_Going_Deeper_With_2015_CVPR_paper.pdf)
#
# Prior to this, most popular CNNs just stacked convolution layers deeper and deeper, hoping to get better performance,however **Inception Network** was one of the first CNN architectures that really strayed from the general approach of simply stacking conv and pooling layers on top of each other in a sequential structure and came up with the **Inception Module**.The Inception network was complex. It used a lot of tricks to push performance; both in terms of speed and accuracy. Its constant evolution lead to the creation of several versions of the network. The popular versions are as follows:
#
# 1. Inception v1.
# 2. Inception v2 and Inception v3.
# 3. Inception v4 and Inception-ResNet.
# <br>
#
# Each version is an iterative improvement over the previous one.Let us go ahead and explore them one by one
# 
#
#
#
# ## Inception V1
# [Inception v1](https://arxiv.org/pdf/1409.4842v1.pdf)
#
# 
# **Problems this network tried to solve:**
# 1. **What is the right kernel size for convolution**
# <br>
# A larger kernel is preferred for information that is distributed more globally, and a smaller kernel is preferred for information that is distributed more locally.
# <br>
# **Ans-** Filters with multiple sizes.The network essentially would get a bit “wider” rather than “deeper”
# <br>
# <br>
# 3. **How to stack convolution which can be less computationally expensive**
# <BR>
# Stacking them naively computationally expensive.
# <br>
# **Ans-**Limit the number of input channels by adding an extra 1x1 convolution before the 3x3 and 5x5 convolutions
# <br>
# <br>
# 2. **How to avoid overfitting in a very deep network**
# <br>
# Very deep networks are prone to overfitting. It also hard to pass gradient updates through the entire network.
# <br>
# **Ans-**Introduce two auxiliary classifiers (The purple boxes in the image). They essentially applied softmax to the outputs of two of the inception modules, and computed an auxiliary loss over the same labels. The total loss function is a weighted sum of the auxiliary loss and the real loss.
#
# The total loss used by the inception net during training.
# <br>
# **total_loss = real_loss + 0.3 * aux_loss_1 + 0.3 * aux_loss_2**
# <br>
# <br>
#
# 
#
# **Points to note**
#
# 1. Used 9 Inception modules in the whole architecture, with over 100 layers in total! Now that is deep…
# 2. No use of fully connected layers! They use an average pool instead, to go from a 7x7x1024 volume to a 1x1x1024 volume. This saves a huge number of parameters.
# 3. Uses 12x fewer parameters than AlexNet.
# 4. Trained on “a few high-end GPUs within a week”.
# 5. It achieved a top-5 error rate of 6.67%
# ## Inception V2
# [Rethinking the Inception Architecture for Computer Vision](https://arxiv.org/pdf/1512.00567v3.pdf)
#
#
# Upgrades were targeted towards:
# 1. Reducing representational bottleneck by replacing 5x5 convolution to two 3x3 convolution operations which further improves computational speed
# <br>
# The intuition was that, neural networks perform better when convolutions didn’t alter the dimensions of the input drastically. Reducing the dimensions too much may cause loss of information, known as a **“representational bottleneck”**
# <br>
# 
# 2. Using smart factorization method where they factorize convolutions of filter size nxn to a combination of 1xn and nx1 convolutions.
# <br>
# For example, a 3x3 convolution is equivalent to first performing a 1x3 convolution, and then performing a 3x1 convolution on its output. They found this method to be 33% more cheaper than the single 3x3 convolution.
# 
#
# # ResNet(2015)
# [Deep Residual Learning for Image Recognition](https://arxiv.org/pdf/1512.03385.pdf)
# 
# **In ResNet, identity mapping is proposed to promote the gradient propagation. Element-wise addition is used. It can be viewed as algorithms with a state passed from one ResNet module to another one.**
#
# 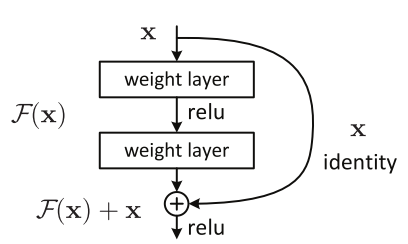
# 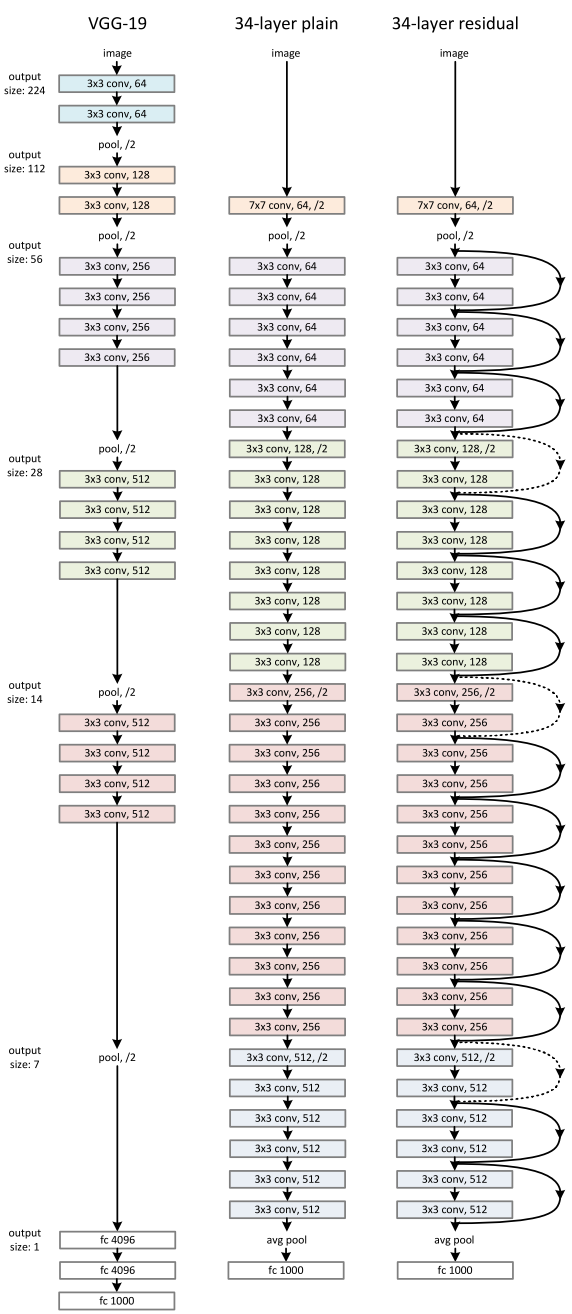
# # ResNet-Wide
# 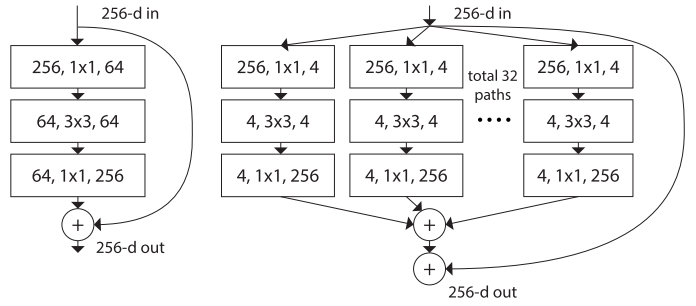
# left: a building block of [2], right: a building block of ResNeXt with cardinality = 32
# # DenseNet(2017)
#
# [Densely Connected Convolutional Networks](https://arxiv.org/pdf/1608.06993v3.pdf)
# <br>
# It is a logical extension to ResNet.
#
# **From the paper:**
# Recent work has shown that convolutional networks can be substantially deeper, more accurate, and efficient to train if they contain shorter connections between layers close to the input and those close to the output. In this paper, we embrace this observation and introduce the Dense Convolutional Network (DenseNet), which connects each layer to every other layer in a feed-forward fashion.
#
# **DenseNet Architecture**
# 
#
# Let us explore different componenets of the network
# <br>
# <br>
# **1. Dense Block**
# <br>
# Feature map sizes are the same within the dense block so that they can be concatenated together easily.
# 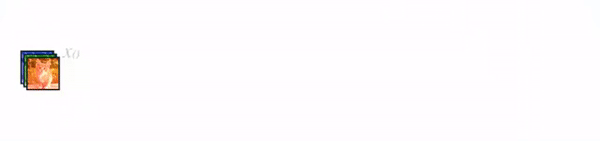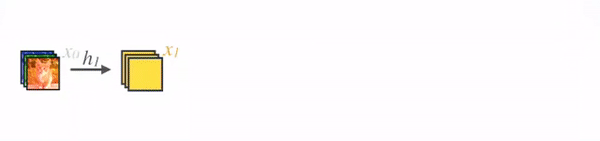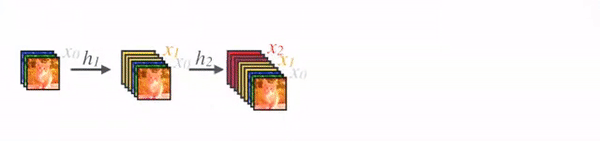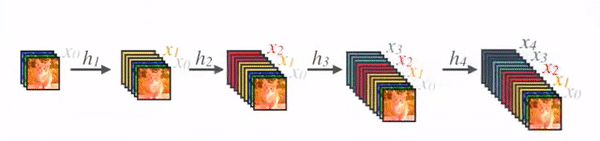
#
# **In DenseNet, each layer obtains additional inputs from all preceding layers and passes on its own feature-maps to all subsequent layers. Concatenation is used. Each layer is receiving a “collective knowledge” from all preceding layers.**
# 
#
# Since each layer receives feature maps from all preceding layers, network can be thinner and compact, i.e. number of channels can be fewer. The growth rate k is the additional number of channels for each layer.
#
# The paper proposed different ways to implement DenseNet with/without B/C by adding some variations in the Dense block to further reduce the complexity,size and to bring more compression in the architecture.
#
# 1. Dense Block (DenseNet)
# - Batch Norm (BN)
# - ReLU
# - 3×3 Convolution
# 2. Dense Block(DenseNet B)
# - Batch Norm (BN)
# - ReLU
# - 1×1 Convolution
# - Batch Norm (BN)
# - ReLU
# - 3×3 Convolution
# 3. Dense Block(DenseNet C)
# - If a dense block contains m feature-maps, The transition layer generate $\theta $ output feature maps, where $\theta \leq \theata \leq$ is referred to as the compression factor.
# - $\theta$=0.5 was used in the experiemnt which reduced the number of feature maps by 50%.
#
# 4. Dense Block(DenseNet BC)
# - Combination of Densenet B and Densenet C
# <br>
# **2. Trasition Layer**
# <br>
# The layers between two adjacent blocks are referred to as transition layers where the following operations are done to change feature-map sizes:
# - 1×1 Convolution
# - 2×2 Average pooling
#
#
# **Points to Note:**
# 1. it requires fewer parameters than traditional convolutional networks
# 2. Traditional convolutional networks with L layers have L connections — one between each layer and its subsequent layer — our network has L(L+1)/ 2 direct connections.
# 3. Improved flow of information and gradients throughout the network, which makes them easy to train
# 4. They alleviate the vanishing-gradient problem, strengthen feature propagation, encourage feature reuse, and substantially reduce the number of parameters.
# 5. Concatenating feature maps instead of summing learned by different layers increases variation in the input of subsequent layers and improves efficiency. This constitutes a major difference between DenseNets and ResNets.
# 6. It achieved a top-5 error rate of 6.66%
# # MobileNet
# ## Spatial Seperable Convolution
# 
#
# **Divides a kernel into two, smaller kernels**
#
# 
#
# **Instead of doing one convolution with 9 multiplications(parameters), we do two convolutions with 3 multiplications(parameters) each (6 in total) to achieve the same effect**
#
# 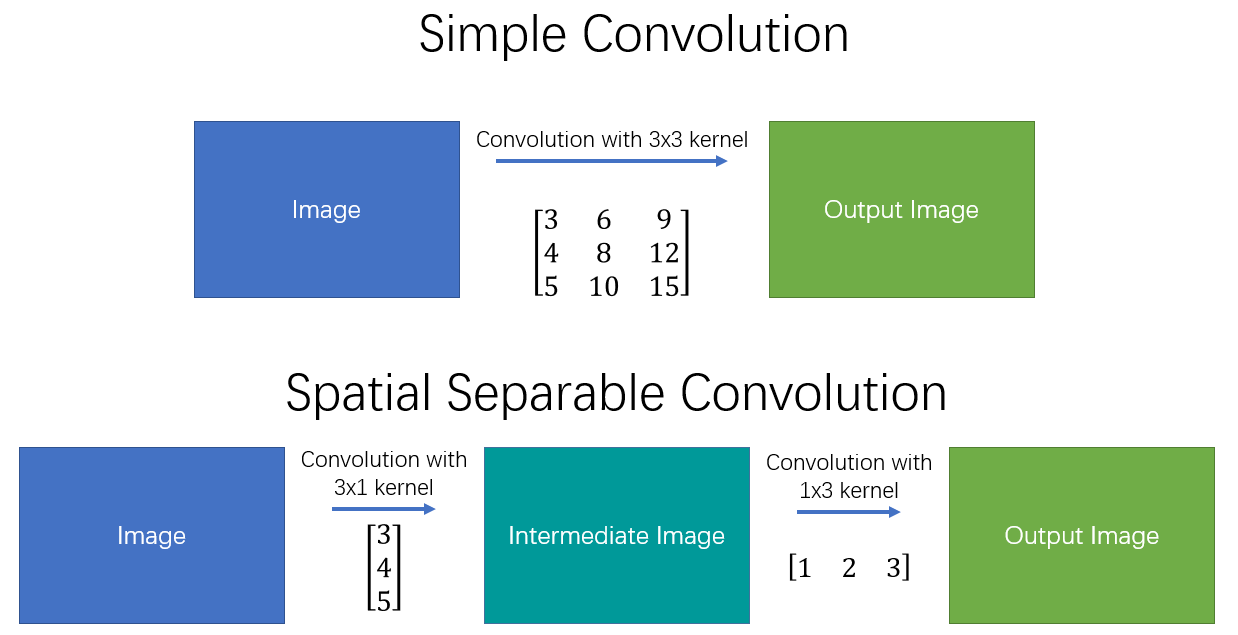
#
#
# **With less multiplications, computational complexity goes down, and the network is able to run faster.**
#
# This was used in an architecture called [Effnet](https://arxiv.org/pdf/1801.06434v1.pdf) showing promising results.
#
# The main issue with the spatial separable convolution is that not all kernels can be “separated” into two, smaller kernels. This becomes particularly bothersome during training, since of all the possible kernels the network could have adopted, it can only end up using one of the tiny portion that can be separated into two smaller kernels.
#
#
#
# ## Depthwise Convolution
# 
#
# Say we need to increase the number of channels from 16 to 32 using 3x3 kernel.
# <br>
#
# **Normal Convolution**
# <br>
# Total No of Parameters = 3 x 3 x 16 x 32 = 4608
#
# 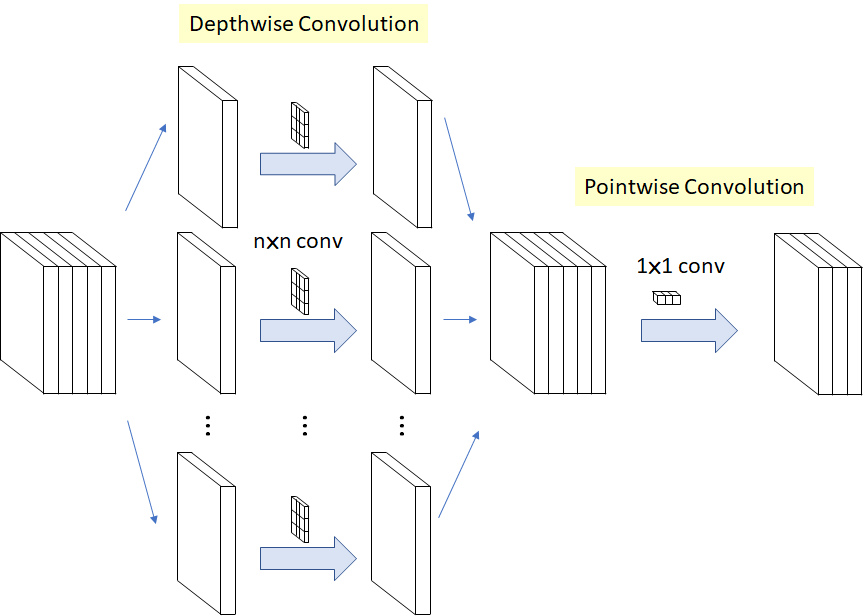
#
# **Depthwise Convolution**
#
# 1. DepthWise Convolution = 16 x [3 x 3 x 1]
# 2. PointWise Convolution = 32 x [1 x 1 x 16]
#
# Total Number of Parameters = 656
#
#
# **Mobile net uses depthwise seperable convolution to reduce the number of parameters**
# # References
#
# [Standford CS231n Lecture Notes](http://cs231n.stanford.edu/slides/2017/cs231n_2017_lecture9.pdf)
# <br>
# [The 9 Deep Learning Papers You Need To Know About](https://adeshpande3.github.io/adeshpande3.github.io/The-9-Deep-Learning-Papers-You-Need-To-Know-About.html)
# <br>
# [CNN Architectures](https://medium.com/@sidereal/cnns-architectures-lenet-alexnet-vgg-googlenet-resnet-and-more-666091488df5)
# <br>
# [Lets Keep It Simple](https://arxiv.org/pdf/1608.06037.pdf)
# <br>
# [CNN Architectures Keras](https://www.pyimagesearch.com/2017/03/20/imagenet-vggnet-resnet-inception-xception-keras/)
# <br>
# [Inception Versions](https://towardsdatascience.com/a-simple-guide-to-the-versions-of-the-inception-network-7fc52b863202)
# <br>
# [DenseNet Review](https://towardsdatascience.com/review-densenet-image-classification-b6631a8ef803)
# <br>
# [DenseNet](https://towardsdatascience.com/densenet-2810936aeebb)
# <br>
# [ResNet](http://teleported.in/posts/decoding-resnet-architecture/)
# <br>
# [ResNet Versions](https://towardsdatascience.com/an-overview-of-resnet-and-its-variants-5281e2f56035)
# <br>
# [Depthwise Convolution](https://towardsdatascience.com/a-basic-introduction-to-separable-convolutions-b99ec3102728)
| Phase - 1/Session4/4_Architectures in CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deploying a web service to Azure Kubernetes Service (AKS)
#
# In this notebook, we show the following steps for deploying a web service using AML:
#
# - Provision an AKS cluster (one time action)
# - Deploy the service
# - Test the web service
# +
import os
import matplotlib.pyplot as plt
import numpy as np
from testing_utilities import to_img, img_url_to_json, plot_predictions
import requests
import json
import subprocess
from azureml.core import Workspace
from azureml.core.compute import AksCompute, ComputeTarget
from azureml.core.webservice import Webservice, AksWebservice
from azureml.core.image import Image
from azureml.core.model import Model
import azureml.core
from dotenv import set_key, get_key, find_dotenv
# %load_ext autoreload
# %autoreload 2
print(azureml.core.VERSION)
# -
env_path = find_dotenv(raise_error_if_not_found=True)
# + tags=["parameters"]
aks_service_name = '<YOUR_AKS_SERVICE_NAME>' # aks_service_name ="my-aks-service-1"
aks_name = '<YOUR_AKS_NAME>' # aks_name = "my-aks-gpu1"
aks_location = '<YOUR_AKS_LOCATION>' # aks_location = "eastus"
# -
set_key(env_path, 'aks_service_name', aks_service_name)
set_key(env_path, 'aks_name', aks_name)
set_key(env_path, 'aks_location', aks_location)
# <a id='get_workspace'></a>
# ## Get workspace
# Load existing workspace from the config file info.
# +
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
# -
# <a id='provision_cluster'></a>
# ## Provision the AKS Cluster¶
# This is a one time setup. You can reuse this cluster for multiple deployments after it has been created. If you delete the cluster or the resource group that contains it, then you would have to recreate it. Let's first check if there are enough cores in the subscription for the cluster.
vm_dict = {
"NC": {
"size": "Standard_NC6",
"cores": 6
}
}
vm_family = "NC"
node_count = 1
requested_cores = node_count * vm_dict[vm_family]["cores"]
results = subprocess.run([
"az", "vm", "list-usage",
"--location", get_key(env_path, "aks_location"),
"--query", "[?contains(localName, '%s')].{max:limit, current:currentValue}" % (vm_family)
], stdout=subprocess.PIPE)
quota = json.loads(''.join(results.stdout.decode('utf-8')))
diff = int(quota[0]['max']) - int(quota[0]['current'])
if diff <= requested_cores:
print("Not enough cores of NC6 in region, asking for {} but have {}".format(requested_cores, diff))
raise Exception("Core Limit", "Note enough cores to satisfy request")
print("There are enough cores, you may continue...")
# +
#Provision AKS cluster with GPU machine
prov_config = AksCompute.provisioning_configuration(vm_size='Standard_NC6')
# Create the cluster
aks_target = ComputeTarget.create(workspace = ws,
name = aks_name,
provisioning_configuration = prov_config)
# -
# %%time
aks_target.wait_for_completion(show_output = True)
print(aks_target.provisioning_state)
print(aks_target.provisioning_errors)
# Optional step: Attach existing AKS cluster
#
# Modify and use below scripts if you have an existing cluster and want to use it as the aks_target. Note that you need to find out the ``cluster_name`` from Azure portal.
# +
# Attach an existing AKS cluster
#attach_config = AksCompute.attach_configuration(resource_group=ws.resource_group,
# cluster_name='my-aks-gpu1b579211166')
#aks_target = ComputeTarget.attach(ws, aks_name, attach_config)
#aks_target.wait_for_completion(True)
# +
# Execute following commands if you want to delete an AKS cluster
#aks_target = AksCompute(name=aks_name,workspace=ws)
#aks_target.delete()
# -
# <a id='deploy_ws'></a>
# ## Deploy web service to AKS¶
#Deploy web service to AKS
#Set the web service configuration (using customized configuration)
aks_config = AksWebservice.deploy_configuration(autoscale_enabled=False, num_replicas=1)
# get the image built in previous notebook
image_name = get_key(env_path, 'image_name')
image = ws.images[image_name]
# +
# %%time
aks_service = Webservice.deploy_from_image(workspace = ws,
name = aks_service_name,
image = image,
deployment_config = aks_config,
deployment_target = aks_target)
aks_service.wait_for_deployment(show_output = True)
print(aks_service.state)
# +
### debug
# aks_service.error
# aks_service.get_logs()
# Excute following commands if you want to delete a web service
# s = Webservice(ws, 'my-aks-service-1')
# s.delete()
# -
# <a id='test_ws'></a>
# ## Test Web Service¶
# We test the web sevice by passing data.
IMAGEURL = "https://upload.wikimedia.org/wikipedia/commons/thumb/6/68/Lynx_lynx_poing.jpg/220px-Lynx_lynx_poing.jpg"
plt.imshow(to_img(IMAGEURL))
jsonimg = img_url_to_json(IMAGEURL)
jsonimg[:100]
resp = aks_service.run(input_data = jsonimg)
print(resp)
# Having deplied web service succesfully, we can now move on to [Test Web app](05_TestWebApp.ipynb).
| Keras_Tensorflow/04_DeployOnAKS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="8cbb2142c6d6"
import matplotlib.pyplot as plt
import numpy as np
import networkx as nx
import torch
# %matplotlib inline
# + [markdown] id="14e0075735cc"
# # Decentralized Gradient under regular dynamic topology
#
#
# After introducing the average consensus over the dynamic one peer strategy of exponential graph, it is natural to apply that on the Adapt-With-Combine algorithm as well. It is simply just
#
# $$
# x_{k+1,i} = \sum_{j \in \mathcal{N}_i} w^{(k)}_{ij}x_{k,j} - \alpha \nabla f_i(x_{k,i})
# $$
#
# noting that we change from the static weights $w_{ij}$ to dynamic $w^{(k)}_{ij}$ and if step size $\alpha$ is zero or $f$ is constant, it regresses to average consensus problem. In order to achieve that in BlueFog, you need to specifiy the weights argument in the `neighbor_allreduce` function. Recall the full function signature is:
#
# ```
# bf.neighbor_allreduce(
# tensor: torch.Tensor,
# self_weight: Union[float, NoneType] = None,
# src_weights: Union[Dict[int, float], NoneType] = None,
# dst_weights: Union[List[int], Dict[int, float], NoneType] = None,
# enable_topo_check: bool = True,
# name: Union[str, NoneType] = None,
# ) -> torch.Tensor
# ```
# Different from the static case, you need to give different `self_weight`, `src_weights`, and `dst_weights` in each iteration.
#
# **Note:** the argument `src_weights`, which can be either the list of ranks sending to or the map of list of ranks to weights, is necessary for dynamic topology.
# + id="4944b03a02a5"
import ipyparallel as ipp
rc = ipp.Client(profile="bluefog")
# + id="447b4ef608a7"
# %%px
import torch
import bluefog.torch as bf
from bluefog.common import topology_util
bf.init()
print(f"Rank: {bf.rank()}, Size: {bf.size()}")
# + id="ec40d2b6f5e8"
# %%px
data_size = 100
seed = 1234
max_iters = 10
torch.random.manual_seed(seed * bf.rank())
x = torch.randn(data_size, dtype=torch.double)
x_bar = bf.allreduce(x, average=True)
mse = [torch.norm(x - x_bar, p=2) / torch.norm(x_bar, p=2)]
# + [markdown] id="bfb9ca16c601"
# Since the design of BlueFog is to sperate the communication functionality from the topology usage, you need to explicitly to create these arguments for the `neighbor_allreduce` function. It is time we combined the utility function `neighbor_allreduce` and `GetDynamicOnePeerSendRecvRanks` for dynamic topology usage.
#
# In this section, we only consider the [regular graph](https://en.wikipedia.org/wiki/Regular_graph) -- a regular graph is a graph where each vertex has the same number of neighbors; i.e. every vertex has the same degree or valency. It is easy to see that under $\tau$-regular graph, the `GetDynamicOnePeerSendRecvRanks` function is $\tau$-periodical function. Further, since `GetDynamicOnePeerSendRecvRanks` determine the send-to neighbor based on the relative difference of node index, every iteration, all nodes will have one destination (send-to) node and one source(receive-from) node only, which lead to $W^{(k)}$ is doubly stochastic (not necessary symmetric) at every iterations. This special property will gurantee that our consensus algorithm still converge unbiasedly. However, in general case, it doesn't hold. We will discuss that in the next subsection.
#
# For now, let's look at the Exponential Two graph, which is regular graph:
# + id="7f1ff0db0425"
# %%px
bf.set_topology(bf.ExponentialTwoGraph(bf.size()))
dynamic_neighbor_allreduce_gen = bf.GetDynamicOnePeerSendRecvRanks(
bf.load_topology(), bf.rank()
)
for ite in range(max_iters):
dst_neighbors, src_neighbors = next(dynamic_neighbor_allreduce_gen)
uniform_weight = 1 / (len(src_neighbors) + 1)
src_weights = {r: uniform_weight for r in src_neighbors}
self_weight = uniform_weight
x = bf.neighbor_allreduce(
x,
name="x",
self_weight=self_weight,
src_weights=src_weights,
dst_weights=dst_neighbors,
enable_topo_check=True,
)
mse.append(torch.norm(x - x_bar, p=2) / torch.norm(x_bar, p=2))
# + id="9ce252c80108"
mse = rc[0].pull("mse", block=True)
# + id="aa9bf7d840a5"
plt.semilogy(mse)
# + [markdown] id="5158ef1e2193"
# Above figure clearly illustrated the lemma in previous section -- the algorithm will reach the consensus under $\tau=$`log(bf.size())` step, which has been show in the lemma of previous subsection.
# + [markdown] id="71ebebf699fe"
# Next, we show the performance comparison between static and dynamic topology over linear cost function.
# + id="b9922908dec2"
num_data = 600
dimension = 20
noise_level = 0.1
X = np.random.randn(num_data, dimension)
x_o = np.random.randn(dimension, 1)
ns = noise_level * np.random.randn(num_data, 1)
y = X.dot(x_o) + ns
# We know the optimal solution in close form solution.
x_opt = np.linalg.inv(X.T.dot(X)).dot(X.T).dot(y)
# + [markdown] id="f311d7c5e868"
# For simplicity, we just use centralized solution to generate the data and distributed the partial data to all workers. Each worker should only contain `1/len(works)` number of all data.
# + id="c9b80f60722f"
num_workers = len(rc.ids)
assert (
num_data % num_workers == 0
), "Please adjust number of data so that it is the multiples of number of workers"
x_opt_torch = torch.from_numpy(x_opt)
for i in range(num_workers):
X_partial = torch.from_numpy(X[i::num_workers])
y_partial = torch.from_numpy(y[i::num_workers])
rc[i].push({"X": X_partial, "y": y_partial, "x_opt": x_opt_torch}, block=True)
# + id="3f36882cfdb1"
# %px print(X.shape, y.shape)
# + id="deca6cb9652d"
# %%px
max_iters = 100
mse_dynamic = []
mse_static = []
bf.set_topology(bf.ExponentialTwoGraph(bf.size()))
dynamic_neighbor_allreduce_gen = topology_util.GetDynamicOnePeerSendRecvRanks(
bf.load_topology(), bf.rank()
)
x_static = torch.randn(x_opt.shape, dtype=torch.double)
x_dynamic = torch.randn(x_opt.shape, dtype=torch.double)
step_size = 0.005
for ite in range(max_iters):
send_neighbors, recv_neighbors = next(dynamic_neighbor_allreduce_gen)
uniform_weight = 1 / (len(recv_neighbors) + 1)
neighbor_weights = {r: uniform_weight for r in recv_neighbors}
self_weight = uniform_weight
x_dynamic = x_dynamic - step_size * X.T.mm(X.mm(x_dynamic) - y)
x_dynamic = bf.neighbor_allreduce(
x_dynamic,
name="x_dynamic",
self_weight=self_weight,
src_weights=neighbor_weights,
dst_weights=send_neighbors,
enable_topo_check=True,
)
x_static = x_static - step_size * X.T.mm(X.mm(x_static) - y)
x_static = bf.neighbor_allreduce(x_static, name="x_static")
mse_dynamic.append(torch.norm(x_dynamic - x_opt, p=2) / torch.norm(x_opt, p=2))
mse_static.append(torch.norm(x_static - x_opt, p=2) / torch.norm(x_opt, p=2))
# + id="3d910b7e4103"
mse_dynamic, mse_static = rc[0].pull(["mse_dynamic", "mse_static"], block=True)
# + id="1af1b86a2d39"
plt.semilogy(mse_dynamic, label="dynamic topology")
plt.semilogy(mse_static, label="static topology")
plt.legend()
| Section 5/Sec-5.3-Average-consenus-over-regular-dyanmic-topology.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS109A Introduction to Data Science
#
# ## Standard Section 4: Regularization and Model Selection
#
# **Harvard University**<br/>
# **Fall 2019**<br/>
# **Instructors**: <NAME>, <NAME>, and <NAME><br/>
# **Section Leaders**: <NAME>, Abhimanyu (<NAME>, Robbert (<NAME><br/>
#
# <hr style='height:2px'>
#RUN THIS CELL
import requests
from IPython.core.display import HTML
styles = requests.get("http://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
# For this section, our goal is to get you familiarized with Regularization in Multiple Linear Regression and to start thinking about Model and Hyper-Parameter Selection.
#
# Specifically, we will:
#
# - Load in the King County House Price Dataset
# - Perform some basic EDA
# - Split the data up into a training, **validation**, and test set (we'll see why we need a validation set)
# - Scale the variables (by standardizing them) and seeing why we need to do this
# - Make our multiple & polynomial regression models (like we did in the previous section)
# - Learn what **regularization** is and how it can help
# - Understand **ridge** and **lasso** regression
# - Get an introduction to **cross-validation** using RidgeCV and LassoCV
# <img src="../fig/meme.png" width="400">
# +
# Data and Stats packages
import numpy as np
import pandas as pd
pd.set_option('max_columns', 200)
# Visualization packages
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
import warnings
warnings.filterwarnings("ignore")
# -
# # EDA: House Prices Data From Kaggle
#
# For our dataset, we'll be using the house price dataset from [King County, WA](https://en.wikipedia.org/wiki/King_County,_Washington). The dataset is from [Kaggle](https://www.kaggle.com/harlfoxem/housesalesprediction).
#
# The task is to build a regression model to **predict the price**, based on different attributes. First, let's do some EDA.
# Load the dataset
house_df = pd.read_csv('../data/kc_house_data.csv')
house_df = house_df.sample(frac=1, random_state=42)[0:4000]
print(house_df.shape)
print(house_df.dtypes)
house_df.head()
# Now let's check for null values and look at the datatypes within the dataset.
house_df.info()
house_df.describe()
# Let's choose a subset of columns here. **NOTE**: The way I'm selecting columns here is not principled and is just for convenience. In your homework assignments (and in real life), we expect you to choose columns more rigorously.
#
# 1. `bedrooms`
# 2. `bathrooms`
# 3. `sqft_living`
# 4. `sqft_lot`
# 5. `floors`
# 6. `sqft_above`
# 7. `sqft_basement`
# 8. `lat`
# 9. `long`
# 10. **`price`**: Our response variable
# +
cols_of_interest = ['bedrooms', 'bathrooms', 'sqft_living', 'sqft_lot', 'floors', 'sqft_above', 'sqft_basement',
'lat', 'long', 'price']
house_df = house_df[cols_of_interest]
# Convert house price to 1000s of dollars
house_df['price'] = house_df['price']/1000
# -
# Let's see how the response variable (`price`) is distributed
fig, ax = plt.subplots(figsize=(12,5))
ax.hist(house_df['price'], bins=100)
ax.set_title('Histogram of house price (in 1000s of dollars)');
# +
# This takes a bit of time but is worth it!!
# sns.pairplot(house_df);
# -
# ## Train-Validation-Test Split
#
# Up until this point, we have only had a train-test split. Why are we introducing a validation set? What's the point?
#
# This is the general idea:
#
# 1. **Training Set**: Data you have seen. You train different types of models with various different hyper-parameters and regularization parameters on this data.
#
#
# 2. **Validation Set**: Used to compare different models. We use this step to tune our hyper-parameters i.e. find the optimal set of hyper-parameters (such as $k$ for k-NN or our $\beta_i$ values or number of degrees of our polynomial for linear regression). Pick your best model here.
#
#
#
# 3. **Test Set**: Using the best model from the previous step, simply report the score e.g. R^2 score, MSE or any metric that you care about, of that model on your test set. **DON'T TUNE YOUR PARAMETERS HERE!**. Why, I hear you ask? Because we want to know how our model might do on data it hasn't seen before. We don't have access to this data (because it may not exist yet) but the test set, which we haven't seen or touched so far, is a good way to mimic this new data.
#
# Let's do 60% train, 20% validation, 20% test for this dataset.
# +
from sklearn.model_selection import train_test_split
# first split the data into a train-test split and don't touch the test set yet
train_df, test_df = train_test_split(house_df, test_size=0.2, random_state=42)
# next, split the training set into a train-validation split
# the test-size is 0.25 since we are splitting 80% of the data into 20% and 60% overall
train_df, val_df = train_test_split(train_df, test_size=0.25, random_state=42)
print('Train Set: {0:0.2f}%'.format(100*train_df.size/house_df.size))
print('Validation Set: {0:0.2f}%'.format(100*val_df.size/house_df.size))
print('Test Set: {0:0.2f}%'.format(100*test_df.size/house_df.size))
# -
# # Modeling
#
# In the [last section](https://github.com/Harvard-IACS/2019-CS109A/tree/master/content/sections/section3), we went over the mechanics of Multiple Linear Regression and created models that had interaction terms and polynomial terms. Specifically, we dealt with the following sorts of models.
#
# $$
# y = \beta_0 + \beta_1 x_1 + \beta_2 x_2 + \dots + \beta_M x_M
# $$
#
# Let's adopt a similar process here and get a few different models.
# ## Creating a Design Matrix
# From our model setup in the equation in the previous section, we obtain the following:
#
# $$
# Y = \begin{bmatrix}
# y_1 \\
# y_2 \\
# \vdots \\
# y_n
# \end{bmatrix}, \quad X = \begin{bmatrix}
# x_{1,1} & x_{1,2} & \dots & x_{1,M} \\
# x_{2,1} & x_{2,2} & \dots & x_{2,M} \\
# \vdots & \vdots & \ddots & \vdots \\
# x_{n,1} & x_{n,2} & \dots & x_{n,M} \\
# \end{bmatrix}, \quad \beta = \begin{bmatrix}
# \beta_1 \\
# \beta_2 \\
# \vdots \\
# \beta_M
# \end{bmatrix}, \quad \epsilon = \begin{bmatrix}
# \epsilon_1 \\
# \epsilon_2 \\
# \vdots \\
# \epsilon_n
# \end{bmatrix},
# $$
#
# $X$ is an n$\times$M matrix: this is our **design matrix**, $\beta$ is an M-dimensional vector (an M$\times$1 matrix), and $Y$ is an n-dimensional vector (an n$\times$1 matrix). In addition, we know that $\epsilon$ is an n-dimensional vector (an n$\times$1 matrix).
X = train_df[cols_of_interest]
y = train_df['price']
print(X.shape)
print(y.shape)
# ## Scaling our Design Matrix
#
# ### Warm-Up Exercise
#
# Warm-Up Exercise: for which of the following do the units of the predictors matter (e.g., trip length in minutes vs seconds; temperature in F or C)? A similar question would be: for which of these models do the magnitudes of values taken by different predictors matter?
#
# (We will go over Ridge and Lasso Regression in greater detail later)
#
# - k-NN (Nearest Neighbors regression)
# - Linear regression
# - Lasso regression
# - Ridge regression
#
# **Solutions**
#
# - kNN: **yes**. Scaling affects distance metric, which determines what "neighbor" means
# - Linear regression: **no**. Multiply predictor by $c$ -> divide coef by $c$.
# - Lasso: **yes**: If we divided coef by $c$, then corresponding penalty term is also divided by $c$.
# - Ridge: **yes**: Same as Lasso, except penalty divided by $c^2$.
#
# ### Standard Scaler (Standardization)
#
# [Here's](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html) the scikit-learn implementation of the standard scaler. What is it doing though? Hint: you may have seen this in STAT 110 or another statistics course multiple times.
#
# $$
# z = \frac{x-\mu}{\sigma}
# $$
#
# In the above setup:
#
# - $z$ is the standardized variable
# - $x$ is the variable before standardization
# - $\mu$ is the mean of the variable before standardization
# - $\sigma$ is the standard deviation of the variable before standardization
#
# Let's see an example of how this works:
# +
from sklearn.preprocessing import StandardScaler
x = house_df['sqft_living']
mu = x.mean()
sigma = x.std()
z = (x-mu)/sigma
# reshaping x to be a n by 1 matrix since that's how scikit learn likes data for standardization
x_reshaped = np.array(x).reshape(-1,1)
z_sklearn = StandardScaler().fit_transform(x_reshaped)
# Plotting the histogram of the variable before standardization
fig, ax = plt.subplots(nrows=1, ncols=3, figsize=(24,5))
ax = ax.ravel()
ax[0].hist(x, bins=100)
ax[0].set_title('Histogram of sqft_living before standardization')
ax[1].hist(z, bins=100)
ax[1].set_title('Manually standardizing sqft_living')
ax[2].hist(z_sklearn, bins=100)
ax[2].set_title('Standardizing sqft_living using scikit learn');
# making things a dataframe to check if they work
pd.DataFrame({'x': x, 'z_manual': z, 'z_sklearn': z_sklearn.flatten()}).describe()
# -
# ### Min-Max Scaler (Normalization)
#
# [Here's](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html) the scikit-learn implementation of the standard scaler. What is it doing though?
#
# $$
# x_{new} = \frac{x-x_{min}}{x_{max}-x_{min}}
# $$
#
# In the above setup:
#
# - $x_{new}$ is the normalized variable
# - $x$ is the variable before normalized
# - $x_{max}$ is the max value of the variable before normalization
# - $x_{min}$ is the min value of the variable before normalization
#
# Let's see an example of how this works:
# +
from sklearn.preprocessing import MinMaxScaler
x = house_df['sqft_living']
x_new = (x-x.min())/(x.max()-x.min())
# reshaping x to be a n by 1 matrix since that's how scikit learn likes data for normalization
x_reshaped = np.array(x).reshape(-1,1)
x_new_sklearn = MinMaxScaler().fit_transform(x_reshaped)
# Plotting the histogram of the variable before normalization
fig, ax = plt.subplots(nrows=1, ncols=3, figsize=(24,5))
ax = ax.ravel()
ax[0].hist(x, bins=100)
ax[0].set_title('Histogram of sqft_living before normalization')
ax[1].hist(x_new, bins=100)
ax[1].set_title('Manually normalizing sqft_living')
ax[2].hist(x_new_sklearn, bins=100)
ax[2].set_title('Normalizing sqft_living using scikit learn');
# making things a dataframe to check if they work
pd.DataFrame({'x': x, 'x_new_manual': x_new, 'x_new_sklearn': x_new_sklearn.flatten()}).describe()
# -
# **The million dollar question**
#
# Should I standardize or normalize my data? [This](https://medium.com/@rrfd/standardize-or-normalize-examples-in-python-e3f174b65dfc), [this](https://medium.com/@swethalakshmanan14/how-when-and-why-should-you-normalize-standardize-rescale-your-data-3f083def38ff) and [this](https://stackoverflow.com/questions/32108179/linear-regression-normalization-vs-standardization) are useful resources that I highly recommend. But in a nutshell, what they say is the following:
#
# **Pros of Normalization**
#
# 1. Normalization (which makes your data go from 0-1) is widely used in image processing and computer vision, where pixel intensities are non-negative and are typically scaled from a 0-255 scale to a 0-1 range for a lot of different algorithms.
# 2. Normalization is also very useful in neural networks (which we will see later in the course) as it leads to the algorithms converging faster.
# 3. Normalization is useful when your data does not have a discernible distribution and you are not making assumptions about your data's distribution.
#
# **Pros of Standardization**
#
# 1. Standardization maintains outliers (do you see why?) whereas normalization makes outliers less obvious. In applications where outliers are useful, standardization should be done.
# 2. Standardization is useful when you assume your data comes from a Gaussian distribution (or something that is approximately Gaussian).
#
# **Some General Advice**
#
# 1. We learn parameters for standardization ($\mu$ and $\sigma$) and for normalization ($x_{min}$ and $x_{max}$). Make sure these parameters are learned on the training set i.e use the training set parameters even when normalizing/standardizing the test set. In sklearn terms, fit your scaler on the training set and use the scaler to transform your test set and validation set (**don't re-fit your scaler on test set data!**).
# 2. The point of standardization and normalization is to make your variables take on a more manageable scale. You should ideally standardize or normalize all your variables at the same time.
# 3. Standardization and normalization is not always needed and is not an automatic thing you have to do on any data science homework!! Do so sparingly and try to justify why this is needed.
#
# **Interpreting Coefficients**
#
# A great quote from [here](https://stats.stackexchange.com/questions/29781/when-conducting-multiple-regression-when-should-you-center-your-predictor-varia)
#
# > [Standardization] makes it so the intercept term is interpreted as the expected value of 𝑌𝑖 when the predictor values are set to their means. Otherwise, the intercept is interpreted as the expected value of 𝑌𝑖 when the predictors are set to 0, which may not be a realistic or interpretable situation (e.g. what if the predictors were height and weight?)
#
# ### Standardizing our Design Matrix
# +
features = ['bedrooms', 'bathrooms', 'sqft_living', 'sqft_lot', 'floors', 'sqft_above', 'sqft_basement',
'lat', 'long']
X_train = train_df[features]
y_train = np.array(train_df['price']).reshape(-1,1)
X_val = val_df[features]
y_val = np.array(val_df['price']).reshape(-1,1)
X_test = test_df[features]
y_test = np.array(test_df['price']).reshape(-1,1)
scaler = StandardScaler().fit(X_train)
# This converts our matrices into numpy matrices
X_train_t = scaler.transform(X_train)
X_val_t = scaler.transform(X_val)
X_test_t = scaler.transform(X_test)
# Making the numpy matrices pandas dataframes
X_train_df = pd.DataFrame(X_train_t, columns=features)
X_val_df = pd.DataFrame(X_val_t, columns=features)
X_test_df = pd.DataFrame(X_test_t, columns=features)
display(X_train_df.describe())
display(X_val_df.describe())
display(X_test_df.describe())
# -
scaler = StandardScaler().fit(y_train)
y_train = scaler.transform(y_train)
y_val = scaler.transform(y_val)
y_test = scaler.transform(y_test)
# ## One-Degree Polynomial Model
# +
import statsmodels.api as sm
from statsmodels.regression.linear_model import OLS
model_1 = OLS(np.array(y_train).reshape(-1,1), sm.add_constant(X_train_df)).fit()
model_1.summary()
# -
# ## Two-Degree Polynomial Model
# +
def add_square_terms(df):
df = df.copy()
cols = df.columns.copy()
for col in cols:
df['{}^2'.format(col)] = df[col]**2
return df
X_train_df_2 = add_square_terms(X_train)
X_val_df_2 = add_square_terms(X_val)
# Standardizing our added coefficients
cols = X_train_df_2.columns
scaler = StandardScaler().fit(X_train_df_2)
X_train_df_2 = pd.DataFrame(scaler.transform(X_train_df_2), columns=cols)
X_val_df_2 = pd.DataFrame(scaler.transform(X_val_df_2), columns=cols)
print(X_train_df.shape, X_train_df_2.shape)
# Also check using the describe() function that the mean and standard deviations are the way we want them
X_train_df_2.head()
# -
model_2 = OLS(np.array(y_train).reshape(-1,1), sm.add_constant(X_train_df_2)).fit()
model_2.summary()
# ## Three-Degree Polynomial Model
# +
# generalizing our function from above
def add_square_and_cube_terms(df):
df = df.copy()
cols = df.columns.copy()
for col in cols:
df['{}^2'.format(col)] = df[col]**2
df['{}^3'.format(col)] = df[col]**3
return df
X_train_df_3 = add_square_and_cube_terms(X_train)
X_val_df_3 = add_square_and_cube_terms(X_val)
# Standardizing our added coefficients
cols = X_train_df_3.columns
scaler = StandardScaler().fit(X_train_df_3)
X_train_df_3 = p
X_val_df_3 = pd.DataFrame(scaler.transform(X_val_df_3), columns=cols)
print(X_train_df.shape, X_train_df_3.shape)
# Also check using the describe() function that the mean and standard deviations are the way we want them
X_train_df_3.head()
# -
model_3 = OLS(np.array(y_train).reshape(-1,1), sm.add_constant(X_train_df_3)).fit()
model_3.summary()
# ## N-Degree Polynomial Model
# +
# generalizing our function from above
def add_higher_order_polynomial_terms(df, N=7):
df = df.copy()
cols = df.columns.copy()
for col in cols:
for i in range(2, N+1):
df['{}^{}'.format(col, i)] = df[col]**i
return df
N = 8
X_train_df_N = add_higher_order_polynomial_terms(X_train,N)
X_val_df_N = add_higher_order_polynomial_terms(X_val,N)
# Standardizing our added coefficients
cols = X_train_df_N.columns
scaler = StandardScaler().fit(X_train_df_N)
X_train_df_N = pd.DataFrame(scaler.transform(X_train_df_N), columns=cols)
X_val_df_N = pd.DataFrame(scaler.transform(X_val_df_N), columns=cols)
print(X_train_df.shape, X_train_df_N.shape)
# Also check using the describe() function that the mean and standard deviations are the way we want them
X_train_df_N.head()
# -
model_N = OLS(np.array(y_train).reshape(-1,1), sm.add_constant(X_train_df_N)).fit()
model_N.summary()
# You can also create a model with interaction terms or any other higher order polynomial term of your choice.
# **Note:** Can you see how creating a function that takes in a dataframe and a degree and creates polynomial terms up until that degree can be useful? This is what we have you do in your homework!
# # Regularization
#
# ## What is Regularization and why should I care?
#
# When we have a lot of predictors, we need to worry about overfitting. Let's check this out:
# +
from sklearn.metrics import r2_score
x = [1,2,3,N]
models = [model_1, model_2, model_3, model_N]
X_trains = [X_train_df, X_train_df_2, X_train_df_3, X_train_df_N]
X_vals = [X_val_df, X_val_df_2, X_val_df_3, X_val_df_N]
r2_train = []
r2_val = []
for i,model in enumerate(models):
y_pred_tra = model.predict(sm.add_constant(X_trains[i]))
y_pred_val = model.predict(sm.add_constant(X_vals[i]))
r2_train.append(r2_score(y_train, y_pred_tra))
r2_val.append(r2_score(y_val, y_pred_val))
fig, ax = plt.subplots(figsize=(8,6))
ax.plot(x, r2_train, 'o-', label=r'Training $R^2$')
ax.plot(x, r2_val, 'o-', label=r'Validation $R^2$')
ax.set_xlabel('Number of degree of polynomial')
ax.set_ylabel(r'$R^2$ score')
ax.set_title(r'$R^2$ score vs polynomial degree')
ax.legend();
# -
# We notice a big difference between training and validation R^2 scores: seems like we are overfitting. **Introducing: regularization.**
# ## What about Multicollinearity?
# There's seemingly a lot of multicollinearity in the data. Take a look at this warning that we got when showing our summary for our polynomial models:
#
# <img src="../fig/warning.png" width="400">
#
# What is [multicollinearity](https://en.wikipedia.org/wiki/Multicollinearity)? Why do we have it in our dataset? Why is this a problem?
#
# Does regularization help solve the issue of multicollinearity?
# ## What does Regularization help with?
#
# We have some pretty large and extreme coefficient values in our most recent models. These coefficient values also have very high variance. We can also clearly see some overfitting to the training set. In order to reduce the coefficients of our parameters, we can introduce a penalty term that penalizes some of these extreme coefficient values. Specifically, regularization helps us:
#
# 1. Avoid overfitting. Reduce features that have weak predictive power.
# 2. Discourage the use of a model that is too complex
#
# <img src="../fig/overfit.png" width="600">
#
# ### Big Idea: Reduce Variance by Increasing Bias
#
# Image Source: [here](https://www.cse.wustl.edu/~m.neumann/sp2016/cse517/lecturenotes/lecturenote12.html)
#
# <img src="../fig/bias_variance.png" width="600">
#
# ## Ridge Regression
#
# Ridge Regression is one such form of regularization. In practice, the ridge estimator reduces the complexity of the model by shrinking the coefficients, but it doesn’t nullify them. We control the amount of regularization using a parameter $\lambda$. **NOTE**: sklearn's [ridge regression package](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Ridge.html) represents this $\lambda$ using a parameter alpha. In Ridge Regression, the penalty term is proportional to the L2-norm of the coefficients.
#
# <img src="../fig/ridge.png" width="400">
#
# ## Lasso Regression
#
# Lasso Regression is another form of regularization. Again, we control the amount of regularization using a parameter $\lambda$. **NOTE**: sklearn's [lasso regression package](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Lasso.html) represents this $\lambda$ using a parameter alpha. In Lasso Regression, the penalty term is proportional to the L1-norm of the coefficients.
#
# <img src="../fig/lasso.png" width="400">
#
# ### Some Differences between Ridge and Lasso Regression
#
# 1. Since Lasso regression tend to produce zero estimates for a number of model parameters - we say that Lasso solutions are **sparse** - we consider to be a method for variable selection.
# 2. In Ridge Regression, the penalty term is proportional to the L2-norm of the coefficients whereas in Lasso Regression, the penalty term is proportional to the L1-norm of the coefficients.
# 3. Ridge Regression has a closed form solution! Lasso Regression does not. We often have to solve this iteratively. In the sklearn package for Lasso regression, there is a parameter called `max_iter` that determines how many iterations we perform.
#
# ### Why Standardizing Variables was not a waste of time
#
# Lasso regression puts constraints on the size of the coefficients associated to each variable. However, this value will depend on the magnitude of each variable. It is therefore necessary to standardize the variables.
# ## Let's use Ridge and Lasso to regularize our degree N polynomial
# **Exercise**: Play around with different values of alpha. Notice the new $R^2$ value and also the range of values that the predictors take in the plot.
# +
from sklearn.linear_model import Ridge
# some values you can try out: 0.01, 0.1, 0.5, 1, 5, 10, 20, 40, 100, 200, 500, 1000, 10000
alpha = 100
ridge_model = Ridge(alpha=alpha).fit(X_train_df_N, y_train)
print('R squared score for our original OLS model: {}'.format(r2_val[-1]))
print('R squared score for Ridge with alpha={}: {}'.format(alpha, ridge_model.score(X_val_df_N,y_val)))
fig, ax = plt.subplots(figsize=(18,8), ncols=2)
ax = ax.ravel()
ax[0].hist(model_N.params, bins=10, alpha=0.5)
ax[0].set_title('Histogram of predictor values for Original model with N: {}'.format(N))
ax[0].set_xlabel('Predictor values')
ax[0].set_ylabel('Frequency')
ax[1].hist(ridge_model.coef_.flatten(), bins=20, alpha=0.5)
ax[1].set_title('Histogram of predictor values for Ridge Model with alpha: {}'.format(alpha))
ax[1].set_xlabel('Predictor values')
ax[1].set_ylabel('Frequency');
# +
from sklearn.linear_model import Lasso
# some values you can try out: 0.00001, 0.0001, 0.001, 0.01, 0.05, 0.1, 0.5, 1, 2, 5, 10, 20
alpha = 0.01
lasso_model = Lasso(alpha=alpha, max_iter = 1000).fit(X_train_df_N, y_train)
print('R squared score for our original OLS model: {}'.format(r2_val[-1]))
print('R squared score for Lasso with alpha={}: {}'.format(alpha, lasso_model.score(X_val_df_N,y_val)))
fig, ax = plt.subplots(figsize=(18,8), ncols=2)
ax = ax.ravel()
ax[0].hist(model_N.params, bins=10, alpha=0.5)
ax[0].set_title('Histogram of predictor values for Original model with N: {}'.format(N))
ax[0].set_xlabel('Predictor values')
ax[0].set_ylabel('Frequency')
ax[1].hist(lasso_model.coef_.flatten(), bins=20, alpha=0.5)
ax[1].set_title('Histogram of predictor values for Lasso Model with alpha: {}'.format(alpha))
ax[1].set_xlabel('Predictor values')
ax[1].set_ylabel('Frequency');
# -
# ## Model Selection and Cross-Validation
#
# Here's our current setup so far:
#
# <img src="../fig/train_val_test.png" width="400">
#
# So we try out 10,000 different models on our validation set and pick the one that's the best? No! **Since we could also be overfitting the validation set!**
#
# One solution to the problems raised by using a single validation set is to evaluate each model on multiple validation sets and average the validation performance. This is the essence of cross-validation!
#
# <img src="../fig/cross_val.png" width="700">
#
# Image source: [here](https://medium.com/@sebastiannorena/some-model-tuning-methods-bfef3e6544f0)
#
# Let's give this a try using [RidgeCV](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RidgeCV.html) and [LassoCV](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LassoCV.html):
# +
from sklearn.linear_model import RidgeCV
from sklearn.linear_model import LassoCV
alphas = (0.001, 0.01, 0.1, 10, 100, 1000, 10000)
# Let us do k-fold cross validation
k = 4
fitted_ridge = RidgeCV(alphas=alphas).fit(X_train_df_N, y_train)
fitted_lasso = LassoCV(alphas=alphas).fit(X_train_df_N, y_train)
print('R^2 score for our original OLS model: {}\n'.format(r2_val[-1]))
ridge_a = fitted_ridge.alpha_
print('Best alpha for ridge: {}'.format(ridge_a))
print('R^2 score for Ridge with alpha={}: {}\n'.format(ridge_a, fitted_ridge.score(X_val_df_N,y_val)))
lasso_a = fitted_lasso.alpha_
print('Best alpha for lasso: {}'.format(lasso_a))
print('R squared score for Lasso with alpha={}: {}'.format(lasso_a, fitted_lasso.score(X_val_df_N,y_val)))
# -
# We can also look at the coefficients of our CV models.
# **Final Step:** report the score on the test set for the model you have chosen to be the best.
# ----------------
# ### End of Standard Section
# ---------------
| content/sections/section4/notebook/cs109a_section_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 64-bit
# language: python
# name: python37364bit3d079e996ccd4349894ed7c686b41899
# ---
# +
import argparse
import configparser
import errno
import json
import os
import pprint
import time
import typing
from datetime import datetime, timedelta, timezone
import requests
# +
def get_conversations_history(
token: str,
channel: str,
latest: float = 0,
limit: int = 100):
Conversations_History_API_URL = "https://slack.com/api/conversations.history"
params = {
'token': token,
'channel': channel,
"limit": limit,
"latest": latest}
r = requests.get(Conversations_History_API_URL, params=params)
json_data = r.json()
if json_data.get('ok', False) is False:
print(json_data)
raise Exception('FAILED TO GET CONVERSATIONS HISTORY!')
else:
return json_data
def return_timestamp_movedbyday(diff: int = -1):
JST = timezone(timedelta(hours=+9), 'JST')
td_two_days_before = timedelta(days=diff)
today_at_midnight = datetime.now(JST).replace(
hour=0, minute=0, second=0, microsecond=0)
two_days_before = today_at_midnight + td_two_days_before
return two_days_before.timestamp()
def delete_message(token: str, channel: str, ts: float):
Chat_Delete_API_URL = "https://slack.com/api/chat.delete"
params = {'token': token, 'channel': channel, "ts": ts}
r = requests.get(Chat_Delete_API_URL, params=params)
json_data = r.json()
if json_data.get('ok', False) is False:
print('message の削除に失敗')
print(json_data)
# +
config_ini = configparser.ConfigParser()
config_ini_path = 'config.ini'
if not os.path.exists(config_ini_path):
raise FileNotFoundError(
errno.ENOENT, os.strerror(
errno.ENOENT), config_ini_path)
config_ini.read(config_ini_path, encoding='utf-8')
read_default = config_ini['DEFAULT']
Token = read_default.get('OAuth_Token')
CHANNEL_ID = read_default.get('CHANNEL_ID')
BOT_ID = read_default.get('BOT_ID')
# -
json_data = get_conversations_history(
Token, CHANNEL_ID, return_timestamp_movedbyday(-10))
# +
cnt = 0
for i in json_data["messages"]:
if 'bot_id' in i:
if i["bot_id"] == BOT_ID:
# delete_message(Token, CHANNEL_ID, i["ts"])
cnt += 1
pprint.pprint(i)
print(f"{cnt} msg del done")
# -
json_data
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # An overview of Gate Set Tomography
# The `pygsti` package provides multiple levels of abstraction over the core Gate Set Tomography (GST) algorithms. This tutorial will show you how to run Gate Set Tomography on some simulated (generated) data, hopefully giving you an overall sense of what it takes (and how easy it is!) to run GST. For more details and options for running GST, see the [GST circuits tutorial](../objects/advanced/GSTCircuitConstruction.ipynb) and the [tutorial covering the different protocols for running GST](GST-Protocols.ipynb).
#
# There are three basic steps to running protocols in pyGSTi:
#
# ## Step 1: create an experiment design
# The first step is creating an object that specifies what data (from the quantum processor) will be needed to perform GST, and how it should be taken. This is called an "experiment design" in pyGSTi.
#
# To run GST, we need the following three inputs:
# 1. a "**target model**" which describes the desired, or ideal, operations we want our experimental hardware to perform. In the example below, we use the target model from one of pyGSTi's build-in "model packs" (see the [tutorial on model packs](objects/advanced/ModelPacks.ipynb)) - which acts on a single qubit with the following operations:
# - three gates: the identity, and $\pi/2$ rotations around the $x$- and $y$-axes.
# - a single state preparation in the $|0\rangle$ state.
# - a 2-outcome measurement with the label "0" associated with measuring $|0\rangle$ and "1" with measuring $|1\rangle$.
#
# 2. a list of circuits tailored to the target model; essentially a list of what experiments we need to run. Using a standard model makes things especially straightforward here, since the building blocks, called *germ* and *fiducial* circuits, needed to make good GST circuits have already been computed (see the [tutorial on GST circuits](../objects/advanced/GSTCircuitConstruction.ipynb)). In the example below, the model pack also provides the necessary germ and fiducial lists, so that all that is needed is a list of "maximum lengths" describing how long (deep) the circuits should be.
#
# 3. data, in the form of experimental outcome counts, for each of the required sequences. In this example we'll generate "fake" or "simulated" data from a depolarized version of our ideal model. For more information about `DataSet` objects, see the [tutorial on DataSets](../objects/DataSet.ipynb).
#
# The first two inputs form an "experiment design", as they describe the experiment that must be performed on a quantum processor (usually running some prescribed set of circuits) in order to run the GST protocol. The third input - the data counts - is packaged with the experiment design to create a `ProtocolData`, or "data" object. As we will see later, a data object serves as the input to the GST protocol.
#
# **The cell below creates an experiment design for running standard GST on the 1-qubit quantum process described by the gates above using circuits whose depth is at most 32.**
# +
import pygsti
from pygsti.modelpacks import smq1Q_XYI
#Step 1: create an "experiment design" for doing GST on the std1Q_XYI gate set
target_model = smq1Q_XYI.target_model() # a Model object
prep_fiducials = smq1Q_XYI.prep_fiducials() # a list of Circuit objects
meas_fiducials = smq1Q_XYI.meas_fiducials() # a list of Circuit objects
germs = smq1Q_XYI.germs() # a list of Circuit objects
maxLengths = [1,2,4,8,16,32]
exp_design = pygsti.protocols.StandardGSTDesign(target_model, prep_fiducials, meas_fiducials,
germs, maxLengths)
# -
# **Pro tip:** the contents of the cell above (except the imports) could be replaced by the single line:
#
# ```exp_design = smq1Q_XYI.get_gst_experiment_design(max_max_length=32)```
#
#
# ## Step 2: collect data as specified by the experiment design
# Next, we just follow the instructions in the experiment design to collect data from the quantum processor (or the portion of the processor we're characterizing). In this example, we'll generate the data using a depolarizing noise model since we don't have a real quantum processor lying around. The call to `simulate_taking_data` should be replaced with the user filling out the empty "template" data set file with real data. Note also that we set `clobber_ok=True`; this is so the tutorial can be run multiple times without having to manually remove the dataset.txt file - we recommend you leave this set to False (the default) when using it in your own scripts.
def simulate_taking_data(data_template_filename):
"""Simulate taking 1-qubit data and filling the results into a template dataset.txt file"""
datagen_model = smq1Q_XYI.target_model().depolarize(op_noise=0.01, spam_noise=0.001)
pygsti.io.fill_in_empty_dataset_with_fake_data(datagen_model, data_template_filename, nSamples=1000, seed=1234)
# +
pygsti.io.write_empty_protocol_data(exp_design, '../tutorial_files/test_gst_dir', clobber_ok=True)
# -- fill in the dataset file in tutorial_files/test_gst_dir/data/dataset.txt --
simulate_taking_data("../tutorial_files/test_gst_dir/data/dataset.txt") # REPLACE with actual data-taking
data = pygsti.io.load_data_from_dir('../tutorial_files/test_gst_dir')
# -
# ## Step 3: Run the GST protocol and create a report
# Now we just instantiate a `StandardGST` protocol and `.run` it on our data object. This returns a results object that can be used to create a report.
# +
#run the GST protocol and create a report
gst_protocol = pygsti.protocols.StandardGST('TP,CPTP,Target')
results = gst_protocol.run(data)
report = pygsti.report.construct_standard_report(
results, title="GST Overview Tutorial Example Report", verbosity=2)
report.write_html("../tutorial_files/gettingStartedReport", verbosity=2)
# -
# You can now open the file [../tutorial_files/gettingStartedReport/main.html](../tutorial_files/gettingStartedReport/main.html) in your browser (Firefox works best) to view the report. **That's it! You've just run GST!**
#
# In the cell above, `results` is a `ModelEstimateResults` object, which is used to generate a HTML report. For more information see the [Results object tutorial](../objects/advanced/Results.ipynb) and [report generation tutorial](../reporting/ReportGeneration.ipynb).
| jupyter_notebooks/Tutorials/algorithms/GST-Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Gy2SN6o21sIM" colab_type="code" colab={}
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_wine
from IPython.display import display
# + id="2iCwUajM2Bvq" colab_type="code" colab={}
def train_validation_test_split(X, y,
train_size=0.7, val_size=0.1,
test_size=0.2, random_state=None,
shuffle=True):
"""
This function is a utility wrapper around the Scikit-Learn train_test_split that splits arrays or
matrices into train, validation, and test subsets.
Args:
X (Numpy array or DataFrame): This is the first param.
y (Numpy array or DataFrame): This is a second param.
train_size (float or int): Proportion of the dataset to include in the train split (0 to 1).
val_size (float or int): Proportion of the dataset to include in the validation split (0 to 1).
test_size (float or int): Proportion of the dataset to include in the test split (0 to 1).
random_state (int): Controls the shuffling applied to the data before applying the split for reproducibility.
shuffle (bool): Whether or not to shuffle the data before splitting
Returns:
Train, test, and validation dataframes for features (X) and target (y).
"""
X_train_val, X_test, y_train_val, y_test = train_test_split(
X, y, test_size=test_size, random_state=random_state, shuffle=shuffle)
X_train, X_val, y_train, y_val = train_test_split(
X_train_val, y_train_val, test_size=val_size / (train_size + val_size),
random_state=random_state, shuffle=shuffle)
return X_train, X_val, X_test, y_train, y_val, y_test
def print_split_summary(X_train, X_val, X_test):
print('######################## TRAINING DATA ########################')
print(f'X_train Shape: {X_train.shape}')
display(X_train.head())
display(X_train.describe(include='all').transpose())
print('')
print('######################## VALIDATION DATA ######################')
print(f'X_val Shape: {X_val.shape}')
display(X_val.head())
display(X_val.describe(include='all').transpose())
print('')
print('######################## TEST DATA ############################')
print(f'X_test Shape: {X_test.shape}')
display(X_test.head())
display(X_test.describe(include='all').transpose())
print('')
# + id="hSNND-_j13vz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="657a5622-69e0-4df0-b6c2-be7085cd8aa8"
raw_data = load_wine()
df = pd.DataFrame(data=raw_data['data'], columns=raw_data['feature_names'])
df['target'] = raw_data['target']
print(df.shape)
# + id="XpHaTPBF15st" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ea0c8e66-71f9-4fbd-c43e-6aaf42e7288e"
X_train, X_val, X_test, y_train, y_val, y_test = train_validation_test_split(
df[['alcohol', 'ash']], df['target'])
print(X_train.shape, X_val.shape, X_test.shape)
# + id="z2NqFCDs2Fhl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d583084d-8196-40da-9802-1528afd44e0c"
print_split_summary(X_train, X_val, X_test)
# + [markdown] id="zwzyhkqK4cRz" colab_type="text"
# ## Refactor Functions to Class
#
# + id="j7Lo7T4j2J24" colab_type="code" colab={}
class My_Data_Splitter():
def __init__(self, df, features, target):
self.df = df
self.features = features
self.target = target
self.X = df[features]
self.y = df[target]
def train_validation_test_split(self,
train_size=0.7, val_size=0.1,
test_size=0.2, random_state=None,
shuffle=True):
"""
This function is a utility wrapper around the Scikit-Learn train_test_split that splits arrays or
matrices into train, validation, and test subsets.
Args:
X (Numpy array or DataFrame): This is the first param.
y (Numpy array or DataFrame): This is a second param.
train_size (float or int): Proportion of the dataset to include in the train split (0 to 1).
val_size (float or int): Proportion of the dataset to include in the validation split (0 to 1).
test_size (float or int): Proportion of the dataset to include in the test split (0 to 1).
random_state (int): Controls the shuffling applied to the data before applying the split for reproducibility.
shuffle (bool): Whether or not to shuffle the data before splitting
Returns:
Train, test, and validation dataframes for features (X) and target (y).
"""
X_train_val, X_test, y_train_val, y_test = train_test_split(
self.X, self.y, test_size=test_size, random_state=random_state, shuffle=shuffle)
X_train, X_val, y_train, y_val = train_test_split(
X_train_val, y_train_val, test_size=val_size / (train_size + val_size),
random_state=random_state, shuffle=shuffle)
return X_train, X_val, X_test, y_train, y_val, y_test
def print_split_summary(self, X_train, X_val, X_test):
print('######################## TRAINING DATA ########################')
print(f'X_train Shape: {X_train.shape}')
display(X_train.head())
display(X_train.describe(include='all').transpose())
print('')
print('######################## VALIDATION DATA ######################')
print(f'X_val Shape: {X_val.shape}')
display(X_val.head())
display(X_val.describe(include='all').transpose())
print('')
print('######################## TEST DATA ############################')
print(f'X_test Shape: {X_test.shape}')
display(X_test.head())
display(X_test.describe(include='all').transpose())
print('')
# + id="jPAzNXrW41Lu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3d7add16-d421-4424-a818-a96cb5629500"
splitter = My_Data_Splitter(df=df, features=['alcohol', 'ash'], target='target')
X_train, X_val, X_test, y_train, y_val, y_test = splitter.train_validation_test_split()
splitter.print_split_summary(X_train, X_val, X_test)
# + id="JmRhbXYF5EMy" colab_type="code" colab={}
| notebook/Lambdata_DSPT6_Dev.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
from avgn.utils.paths import DATA_DIR, most_recent_subdirectory, ensure_dir
from sklearn.preprocessing import scale
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
from avgn.visualization.projections import scatter_spec
from avgn.utils.general import save_fig
from avgn.utils.paths import FIGURE_DIR, ensure_dir
datasets = list(DATA_DIR.glob('embeddings/*/*.pickle'))
datasets = pd.DataFrame([[i.parent.stem, i.stem, i] for i in datasets], columns = ['ds', 'indv', '_loc'])
datasets[:3]
ds_loc = datasets[datasets.ds == 'buckeye'].iloc[0]._loc
ds_loc
syllable_df = pd.read_pickle(ds_loc)
syllable_df[:3]
specs = np.stack(syllable_df['spectrogram'].values)
np.shape(specs)
embedding = np.vstack(syllable_df['umap'].values)
np.shape(embedding)
def remove_outliers(z, pct = 99.5):
""" GPU based UMAP algorithm produces some outliers that UMAP does not, but is much faster
this is a quick fix for that.
"""
_min = np.percentile(z, (100-pct), axis=0)
_max = np.percentile(z, pct, axis=0)
for col in range(np.shape(z)[1]):
mask = z[:,col] < _min[col]
z[mask,col] = _min[col]
mask = z[:,col] > _max[col]
z[mask,col] = _max[col]
return z
fig, ax = plt.subplots()
ax.scatter(embedding[:,0], embedding[:,1], s=1, color='k', alpha = 0.005)
ax.set_xlim([-8,8])
ax.set_ylim([-8,8])
# +
nex = -1
scatter_spec(
embedding[:nex],
specs[:nex],
column_size=8,
#x_range = [-4.5,4],
#y_range = [-4.5,5.5],
pal_color="hls",
color_points=False,
enlarge_points=0,
figsize=(10, 10),
range_pad = 0.2,
scatter_kwargs = {
'labels': syllable_df.labels.values[:nex],
'alpha':0.05,
's': 1,
'show_legend': False
},
matshow_kwargs = {
'cmap': plt.cm.Greys
},
line_kwargs = {
'lw':3,
'ls':"dashed",
'alpha':0.25,
},
draw_lines=True,
n_subset= 1000,
border_line_width = 3,
);
#save_fig(FIGURE_DIR / 'phonemes', dpi=300, save_jpg=True)
# -
| notebooks/11.0-umap-projection-fig/.ipynb_checkpoints/plot-hcassins-umap-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Calculating the Return of a Portfolio of Securities
# Load the data for a portfolio composed of 5 stocks - British Petroleum, Ford, Exxon, Lincoln, and Apple *(5_stocks_2000_2017.csv)*.
# ### Normalization to 100:
#
# $$
# \frac {P_t}{P_0} * 100
# $$
# Normalize to a hundred and plot the data on a graph (you can apply the .loc() or the .iloc() method).
# How would you interpret the behavior of the stocks? Just by looking at the chart, would you be able to create a portfolio that provides a solid return on investment?
# *****
# ### Calculating the Return of a Portfolio of Securities
# Obtain the simple return of the securities in the portfolio and store the results in a new table.
# First, assume you would like to create an equally-weighted portfolio. Create the array, naming it “weights”.
# Obtain the annual returns of each of the stocks and then calculate the dot product of these returns and the weights.
# Transform the result into a percentage form.
# Is the return of this portfolio satisfactory?
| Python for Finance - Code Files/67 Calculating the Return of a Portfolio of Securities/CSV/Python 3 CSV/Calculating the Return of a Portfolio of Securities - Exercise_CSV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# In this notebook we present an application that consists of the global *finite-element mesh* plot using the `pydiva2d` module.
# To make the work of the users easier, the files corresponding to the global mesh will be made available; the present test uses the mesh corresponding to the Mediterranean.
# + deletable=true editable=true
import os
import sys
import numpy as np
import pydiva2d
import logging
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
# -
pydiva2d.logger.handlers[1].setLevel(logging.DEBUG)
# + [markdown] deletable=true editable=true
# # Prepare the input
#
# Indicate the path and names of the 2 mesh files, and define a *figure* directory:
# + deletable=true editable=true
meshdir = '../data/Mesh/Global/'
meshfile = os.path.join(meshdir, 'fort.22')
meshtopofile = os.path.join(meshdir, 'fort.23')
figdir = './figures/GlobalMesh/'
# + [markdown] deletable=true editable=true
# Create a Mesh object and read the information from the files:
# + deletable=true editable=true
globalmesh = pydiva2d.Diva2DMesh()
globalmesh.read_from(meshfile, meshtopofile)
globalmesh.describe()
# + [markdown] deletable=true editable=true
# Create the figure directory if it doesn't exist yet:
# + deletable=true editable=true
if not(os.path.exists(figdir)):
os.makedirs(figdir)
# + [markdown] deletable=true editable=true
# # Create the figures
# ## Without Basemap
# + deletable=true editable=true
fig = plt.figure()
ax = plt.subplot(111)
globalmesh.add_to_plot(linewidth=.1)
plt.savefig(os.path.join(figdir, 'GlobalMesh.png'), dpi=300)
plt.close()
# + [markdown] deletable=true editable=true
# ## With Basemap
#
# We use the same commands as the previous cell, except that we create a Basemap instance.
# + deletable=true editable=true
from matplotlib import rcParams
rcParams['agg.path.chunksize'] = 10000
# -
m = Basemap(resolution='l',
projection='ortho',
lat_0=globalmesh.ynode.mean(),
lon_0=globalmesh.xnode.mean())
# + deletable=true editable=true
#m = Basemap(llcrnrlon=GlobalMesh.xnode.min(), llcrnrlat=GlobalMesh.ynode.min(),
# urcrnrlon=GlobalMesh.xnode.max(), urcrnrlat=GlobalMesh.ynode.max(),
# resolution = 'l', epsg=3857)
fig = plt.figure()
#ax = plt.subplot(111)
globalmesh.add_to_plot(m=m, linewidth=.1, color=(0.44, 0.55, .83))
#plt.savefig(os.path.join(figdir, 'GlobalMeshBasemap.png'), dpi=300)
plt.show()
plt.close()
# + [markdown] deletable=true editable=true
# # Generate an animation (if you have time)
# + [markdown] deletable=true editable=true
# Something you can try during coffee time, as it can take a while.<br>
# We repeat the previous cells in a loop in which we modify the central longitude in the `Basemap`.
# + deletable=true editable=true
lon_init = 0.
lon_end = 30.
lon_step = .25
# + deletable=true editable=true
for index, lonc in enumerate(np.arange(lon_init, lon_end, lon_step)):
m = Basemap(resolution='c', projection='ortho',lat_0=30., lon_0=lonc)
ax = plt.subplot(111)
globalmesh.add_to_plot(m, linewidth=.1, color=(0.44, 0.55, .83))
plt.savefig(os.path.join(figdir, "globalmesh{0}.png".format(str(index).zfill(4))),
dpi=300,
bbox='tight')
plt.close()
# -
| Notebooks/plot_global_mesh.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Gaussian Mixture Model for Density Estimation
# This notebook demonstrates Gaussian mixture models (GMMs) in 2D. We can see that the GMM can model quite complicated distributions, but in certain situations be unnecessarily parameterised.
#
# The approximation of a large GMM (i.e. with a large number of components) with a smaller one is a challenging task. This can be achieved using the well-known Expectation Maximisation algorithm for GMMs as applied to a _sample_ from such a model, but no analytic optimisation procedure is known. A poor-man's version might be the Variational Boosting algorithm (Miller, Foti, and Adams - ICML 2017) which greedily fits Gaussian components via KL minimisation, but this is reasonably expensive and by no means optimal.
#
# The below code generates a random Gaussian mixture, draws samples from it, and then fits a smaller GMM to this synthetic data.
import numpy as np
from sklearn.mixture import GaussianMixture
import matplotlib.pyplot as plt
def gaussian_2D_level_curve(mu, sigma, alpha=2, ncoods=100, plot=True):
# (<NAME> 2016)
# (Ported from my matlab utils / pyalexutil)
assert isinstance(mu, np.ndarray) and isinstance(sigma, np.ndarray), "mu/sigma must be numpy arrays."
assert mu.shape == (2, ), 'mu must be vector in R^2'
assert sigma.shape == (2, 2), 'sigma must be 2x2 array'
U, S, V = np.linalg.svd(sigma)
sd = np.sqrt(S)
coods = np.linspace(0, 2 * np.pi, ncoods)
coods = np.vstack((sd[0] * np.cos(coods), sd[1] * np.sin(coods))) * alpha
# project onto basis of ellipse
coods = (V @ coods).T
# add mean
coods += mu
if plot:
plt.plot(*coods.T)
return coods
def gen_mix_mvn(n_components=30, d=2, n_samples=400):
# Generate random multivariate Gaussians
pi = np.random.dirichlet([0.8]*n_components)
mu = np.random.multivariate_normal(np.zeros(d), np.eye(d)*25, size=(n_components))
mu = np.random.rand(n_components, d) * 10 - 5
sigma = np.zeros((n_components, d, d))
for n in range(n_components):
_tmpmat = np.random.rand(d,d)
Q, _junk = np.linalg.qr(_tmpmat)
lam = np.random.exponential(1, d)
sigma[n] = Q @ np.diag(lam) @ Q.T
# Draw samples
z = np.random.multinomial(n_samples, pi)
smps = np.zeros((n_samples, d))
indexes = np.stack((np.cumsum(np.concatenate(([0], z[:-1]))),
np.cumsum(z)), axis=1)
for ixs, n, m, s in zip(indexes, z, mu, sigma):
smps[slice(*ixs)] = np.random.multivariate_normal(m, s, size=n)
return smps, (pi, mu, sigma)
# %timeit -n10 gmm.fit(smps)
# +
n_components = 30 # number of Gaussians in original mixture
n_approx = 5 # number of Gaussians to approximate with
n_samples = 800 # number of samples to draw from mixture for visualisation.
# generate random Gaussian mixture
smps, pars = gen_mix_mvn(30, n_samples=n_samples)
f, axs = plt.subplots(2, 2)
f.set_size_inches(12,12)
# plot level curves of random Gaussian mixture
axs[0,0].set_title('Original Gaussian Mixture (alpha=weight)')
for pi, m, s in zip(*pars):
axs[0,0].plot(*gaussian_2D_level_curve(m, s, plot=False).T, alpha=pi/max(pars[0]))
# sample from this Gaussian mixture
axs[0,1].scatter(*smps.T, alpha=0.4)
axs[0,1].set_title('Sample from Original Density')
# fit a new Gaussian mixture with 5 components:
gmm = GaussianMixture(n_components=n_approx)
gmm.fit(smps)
axs[1,0].set_title('Approximated Gaussian Mixture (alpha=weight)')
maxw = max(gmm.weights_)
for pi, m, s in zip(gmm.weights_ ,gmm.means_, gmm.covariances_):
axs[1,0].plot(*gaussian_2D_level_curve(m, s, plot=False).T, alpha=pi/maxw)
# sample from this Gaussian mixture
smps_approx = gmm.sample(n_samples)[0]
axs[1,1].scatter(*smps_approx.T, alpha=0.4)
axs[1,1].set_title('Sample from Approximated Density');
# -
# #### ^^ Above
# [**<span style='color:blue'>Top left</span>**] The original (generated) Gaussian mixture. The 2 sigma level curves are drawn (note that this corresponds only to approx 63% of density in 2D) and their weight in the mixture denoted by the alpha value (transparency). Often a fairly large number of low weight components are present (this occurs e.g. in Gaussian Sum filtering).<br>
# [**<span style='color:blue'>Top right</span>**] A sample (default: 800) points are drawn from this mixture. The generative model is to choose the component proportionally to the weights, and then draw a sample from the relevant Gaussian. In a certain sense the sample appears less complicated than the original mixture.<br>
# [**<span style='color:blue'>Bottom left</span>**] The fitted GMM (using _sklearn_'s version of the EM algorithm). This is implemented reasonably well and converges quickly. However, this may take some time in high dimensions - it is an iterative co-ordinate descent algorithm and is particularly prone to slow convergence if boundaries between components are ill-defined.<br>
# [**<span style='color:blue'>Bottom right</span>**] A sample from the approximate GMM. This often appears superficially very similar to the original density and gives credence to the idea that small GMMs are capable of capturing the salient features of complicated densities.
# ### Comparison with standard Gaussian Sum collapse heuristics
# +
ixs_top4 = np.flip(np.argsort(pars[0]), axis=0)[:n_approx-1]
ixs_other = np.array(list(set(np.argsort(pars[0])) - set(ixs_top4)))
pi_approx = np.concatenate((pars[0][ixs_top4], [sum(pars[0][ixs_other])]))
mu_approx = pars[1][ixs_top4]
mu_other = np.dot(pars[0][ixs_other], pars[1][ixs_other])/pi_approx[-1]
sigma_approx = pars[2][ixs_top4]
sigma_other = -np.outer(mu_other, mu_other)
for i in ixs_other:
sigma_other += pars[0][i] * (pars[2][i] + np.outer(pars[1][i], pars[1][i]))
mu_approx = np.concatenate((mu_approx, mu_other[None,:]), axis=0)
sigma_approx = np.concatenate((sigma_approx, sigma_other[None,:]), axis=0)
# +
f, axs = plt.subplots(1, 2)
f.set_size_inches(12,6)
# plot level curves of random Gaussian mixture
axs[0].set_title('Heuristic Approximate Gaussian Mixture (alpha=weight)')
for pi, m, s in zip(pi_approx, mu_approx, sigma_approx):
axs[0].plot(*gaussian_2D_level_curve(m, s, plot=False).T, alpha=pi/max(pi_approx))
# sample from this Gaussian mixture
gmm.means_ = mu_approx; gmm.covariances_ = sigma_approx; gmm.weights_ = pi_approx
smps_approx = gmm.sample(n_samples)[0]
axs[1].scatter(*smps_approx.T, alpha=0.4)
axs[1].set_title('Sample from Heuristic Approximation');
# +
# np.exp?
# +
# KL divergence calculation
def calc_prob(x, pars):
n_dim = len(x)
n_mix = len(pars[0])
prob = 0
for m in range(n_mix):
sigma_inv = np.linalg.inv(pars[2][m])
_a = np.exp(-np.dot(x - pars[1][m], np.dot(sigma_inv, x - pars[1][m]))/2)
_a = _a / (np.sqrt(2*np.pi) * np.linalg.det(pars[2][m]))
prob += pars[0][m] * _a
return prob
def calc_kl_from_samples(sample, pars1, pars2):
kl = 0
for i in sample:
p = calc_prob(i, pars1)
q = calc_prob(i, pars2)
if q > 1e-10:
kl += p * (np.log(p) - np.log(q))
return kl
# +
approx_pars = (pi_approx, mu_approx, sigma_approx)
em_pars = gmm.weights_, gmm.means_, gmm.covariances_
approx_kl = calc_kl_from_samples(smps, pars, approx_pars)
em_kl = calc_kl_from_samples(smps, pars, em_pars)
# +
from IPython.display import Markdown as md
md(
"""
# Comparison
| dimension | Method | KL | Time |
| :---: | :---: | :---: | :---: |
| 2 | EM | {:.5} | {} |
|2 | Barber | {:.5} | {} |""".format(approx_kl, "20ms" , em_kl, 0.00001) )
| .ipynb_checkpoints/GMM-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tt
# language: python
# name: tt
# ---
# # Databases of measures
#
# This Notebook contains code to parse covid-19 measures from four different sources:
#
# 1. [Tracked Together](https://thecorrespondent.com/collection/track-ed-together). A database of covid-19 surveillance measures compiled by De Correspondent.
# 2. [CoronaNet](https://www.coronanet-project.org/download.html): (Cheng et al, 2020)
# 3. [CCCSL](https://github.com/amel-github/covid19-interventionmeasures): (Desvars-Larrive et al, 2020)
# 4. [Oxford Covid Policy Tracker](https://github.com/OxCGRT/covid-policy-tracker): (Hale et al, 2021)
import json
import pandas as pd
import numpy as np
import config
PATH = config.PATH_TRACKERS
# ## Tracked Together - De Correspondent
#
# There are three data dumps: tools (measures), methods (technologies used) and purposes (of the measures).
#
# We don't need all fields yet. These will do for now:
# * title - string
# * status - string
# * launch_date - date
# * involved_organizations - list
# * purposes - list
# * methods - list
# * location
# +
# Import data
tools = pd.read_json(PATH + 'tools.json')
#methods = pd.read_json('data/methods.json')
#purposes = pd.read_json('data/purposes.json')
# +
# Clean data
# TODO: Make more pythonic
tools['purpose'] = [[x['title'] for x in list_dict] for list_dict in tools['purposes']]
tools['purpose_id'] = [[x['_id'] for x in list_dict] for list_dict in tools['methods']]
tools['method'] = [[x['title'] for x in list_dict] for list_dict in tools['methods']]
tools['method_id'] = [[x['_id'] for x in list_dict] for list_dict in tools['methods']]
tools['method'] = tools['method'].apply(', '.join)
tools['method'] = tools['method'].apply(lambda x: ', '.join(sorted(x.split(', '))))
tools['purpose'] = tools['purpose'].apply(', '.join)
tools['purspose'] = tools['purpose'].apply(lambda x: ', '.join(sorted(x.split(', '))))
tools['country'] = [x.get('country') for x in tools['location']]
tools['country_code'] = [[x['iso_code'] for x in list_dict] for list_dict in tools['country']]
tools['country_code'] = tools['country_code'].apply(', '.join)
tools['organizations'] = [[x['name'] for x in list_dict] for list_dict in tools['involved_organisations']]
tools['organizations'] = tools['organizations'].apply(', '.join)
tools['organizations'] = tools['organizations'].apply(lambda x: ', '.join(sorted(x.split(', '))))
# +
# Extract date and add to new column
new_column = []
for i in tools.launch_date:
if i is None:
d = np.nan
new_column.append(d)
else:
d = i.get('date')
new_column.append(d)
tools['date'] = new_column
# +
# Trim columns
tools = tools[['id', 'title', 'status', 'date', 'purpose', 'method', 'country_code',
'description', 'organizations', 'target', 'link', 'enforcement_details', 'revision',
'involved_organisations', 'purposes', 'methods', 'location', 'launch_date']]
# +
purpose = 'Contact tracing'
ct = tools[tools['purpose'].apply(lambda x: purpose in x)]
#ct = ct[ct['status'] == 'launched']
ct.country_code.value_counts()
# -
len(tools)
# +
method = 'Hand washing'
f = tools[tools['method'].apply(lambda x: method in x)]
f.country_code.value_counts()
# -
tools.to_csv(PATH + 'tools.csv', index=None)
# ## CoronaNet
#
# Different datasets available:
# 1. [Country files](https://github.com/saudiwin/corona_tscs/tree/master/data/CoronaNet/data_country/coronanet_release?)
# 2. [Extended data set](https://github.com/saudiwin/corona_tscs/blob/master/data/CoronaNet/data_bulk/coronanet_release_allvars.csv.gz). This dataset contains ests from the [CoronaNet testing database](http://coronanet-project.org);
# Cases/deaths/recovered from the JHU [data repository](https://github.com/CSSEGISandData/COVID-19);
# Country-level covariates including GDP, V-DEM democracy scores, human rights indices, power-sharing indices, and press freedom indices from the [Niehaus World Economics and Politics Dataverse](https://niehaus.princeton.edu/news/world-economics-and-politics-dataverse)
# 3. [Core dataset](https://github.com/saudiwin/corona_tscs/blob/master/data/CoronaNet/data_bulk/coronanet_release.csv.gz)
cn = pd.read_csv(PATH + 'coronanet_release.csv')
len(cn)
cn.columns
# +
q = 'temperature sc'
df_cn = cn[cn['description'].str.contains(q, case=False, na=False)]
# -
df_cn.to_csv(PATH + 'temperature_screening.csv', index=None)
# ## CCCSL
#
# There is a glossary of codes [here](https://github.com/amel-github/covid19-interventionmeasures/blob/master/CCCSL_Glossary%20of%20codes.docx)
cc = pd.read_csv(PATH + 'CCCSL_database_version2.csv', encoding='cp1252')
cc.head()
q = 'temperature'
df = cc[cc['Measure_L1'].str.contains(q, case=False, na=False) | \
cc['Measure_L2'].str.contains(q, case=False, na=False) | \
cc['Measure_L3'].str.contains(q, case=False, na=False) | \
cc['Measure_L4'].str.contains(q, case=False, na=False) | \
cc['Comment'].str.contains(q, case=False, na=False)]
df.head()
# ## Oxford
#
# Codebook can be found [here](https://github.com/OxCGRT/covid-policy-tracker/blob/master/documentation/codebook.md).
ox = pd.read_csv(PATH + 'OxCGRT_latest_withnotes.csv', low_memory=False)
ox.head()
# +
# Create dicts from code book
c1 = {0.0: 'no measures',
1.0: 'recommend closing or all schools open with alterations resulting in significant differences compared to non-Covid-19 operations',
2.0: 'require closing (only some levels or categories, eg just high school, or just public schools)',
3.0: 'require closing all levels'
}
c_flag = {0.0: 'targeted',
1.0: 'general'
}
c2 = {0.0: 'no measures',
1.0: 'recommend closing (or recommend work from home)',
2.0: 'require cancelling',
3.0: 'require closing (or work from home) for all-but-essential workplaces (eg grocery stores, doctors)'
}
c3 = {0.0: 'no measures',
1.0: 'recommend cancelling',
2.0: 'require cancelling'
}
c4 = {0.0: 'no restrictions',
1.0: 'restrictions on very large gatherings (the limit is above 1000 people)',
2.0: 'restrictions on gatherings between 101-1000 people',
3.0: 'restrictions on gatherings between 11-100 people',
4.0: 'restrictions on gatherings of 10 people or less'
}
c5 = {0.0: 'no measures',
1.0: 'recommend closing (or significantly reduce volume/route/means of transport available)',
2.0: 'require closing (or prohibit most citizens from using it)'
}
c6 = {0.0: 'no measures',
1.0: 'recommend not leaving house',
2.0: 'require not leaving house with exceptions for daily exercise, grocery shopping, and "essential" trips',
3.0: 'require not leaving house with minimal exceptions (eg allowed to leave once a week, or only one person can leave at a time, etc)'
}
c7 = {0.0: 'no measures',
1.0: 'recommend not to travel between regions/cities',
2.0: 'internal movement restrictions in place'
}
c8 = {0.0: 'no restrictions',
1.0: 'screening arrivals',
2.0: 'quarantine arrivals from some or all regions',
3.0: 'ban arrivals from some regions',
4.0: 'ban on all regions or total border closure'
}
e1 = {0.0: 'no income support',
1.0: 'government is replacing less than 50% of lost salary (or if a flat sum, it is less than 50% median salary)',
2.0: 'government is replacing 50% or more of lost salary (or if a flat sum, it is greater than 50% median salary)'
}
e_flag = {0.0: 'formal sector workers only or informal sector workers only',
1.0: 'all workers'
}
e2 = {0.0: 'no debt/contract relief',
1.0: 'narrow relief, specific to one kind of contract',
2.0: 'broad debt/contract relief'
}
e3 = {0.0: 'no new spending that day'}
e4 = {0.0: 'no new spending that day'}
h_flag = {0.0: 'targeted',
1.0: 'general'
}
h1 = {0.0: 'no Covid-19 public information campaign',
1.0: 'public officials urging caution about Covid-19',
2.0: 'coordinated public information campaign (eg across traditional and social media)'
}
h2 = {0.0: 'no testing policy',
1.0: 'only those who both (a) have symptoms AND (b) meet specific criteria (eg key workers, admitted to hospital, came into contact with a known case, returned from overseas)',
2.0: 'testing of anyone showing Covid-19 symptoms',
3.0: 'open public testing (eg "drive through" testing available to asymptomatic people)'
}
h3 = {0.0: 'no contact tracing',
1.0: 'limited contact tracing; not done for all cases',
2.0: 'comprehensive contact tracing; done for all identified cases'
}
h4 = {0.0: 'no new spending that day'}
h5 = {0.0: 'no new spending that day'}
h6 = {0.0: 'No policy',
1.0: 'Recommended',
2.0: 'Required in some specified shared/public spaces outside the home with other people present, or some situations when social distancing not possible',
3.0: 'Required in all shared/public spaces outside the home with other people present or all situations when social distancing not possible',
4.0: 'Required outside the home at all times regardless of location or presence of other people'
}
h7 = {0.0: 'No availability',
1.0: 'Availability for ONE of following: key workers/ clinically vulnerable groups (non elderly) / elderly groups',
2.0: 'Availability for TWO of following: key workers/ clinically vulnerable groups (non elderly) / elderly groups',
3.0: 'Availability for ALL of following: key workers/ clinically vulnerable groups (non elderly) / elderly groups',
4.0: 'Availability for all three plus partial additional availability (select broad groups/ages)',
5.0: 'Universal availability'
}
h7_flag = {0.0: 'At cost to individual (or funded by NGO, insurance, or partially government funded)',
1.0: 'No or minimal cost to individual (government funded or subsidised)'
}
h8 = {0.0: 'no measures',
1.0: 'Recommended isolation, hygiene, and visitor restriction measures in LTCFs and/or elderly people to stay at home',
2.0: 'Narrow restrictions for isolation, hygiene in LTCFs, some limitations on external visitors and/or restrictions protecting elderly people at home',
3.0: 'Extensive restrictions for isolation and hygiene in LTCFs, all non-essential external visitors prohibited, and/or all elderly people required to stay at home and not leave the home with minimal exceptions, and receive no external visitors'
}
# +
# Replace code book values with strings. This could also be done later in te process.
ox['C1_School closing'] = ox['C1_School closing'].replace(c1)
ox['C1_Flag'] = ox['C1_Flag'].replace(c_flag)
ox['C2_Workplace closing'] = ox['C2_Workplace closing'].replace(c2)
ox['C2_Flag'] = ox['C2_Flag'].replace(c_flag)
ox['C3_Cancel public events'] = ox['C3_Cancel public events'].replace(c3)
ox['C3_Flag'] = ox['C3_Flag'].replace(c_flag)
ox['C4_Restrictions on gatherings'] = ox['C4_Restrictions on gatherings'].replace(c4)
ox['C4_Flag'] = ox['C4_Flag'].replace(c_flag)
ox['C5_Close public transport'] = ox['C5_Close public transport'].replace(c5)
ox['C5_Flag'] = ox['C5_Flag'].replace(c_flag)
ox['C6_Stay at home requirements'] = ox['C6_Stay at home requirements'].replace(c6)
ox['C6_Flag'] = ox['C6_Flag'].replace(c_flag)
ox['C7_Restrictions on internal movement'] = ox['C7_Restrictions on internal movement'].replace(c7)
ox['C7_Flag'] = ox['C7_Flag'].replace(c_flag)
ox['C8_International travel controls'] = ox['C8_International travel controls'].replace(c8)
ox['E1_Income support'] = ox['E1_Income support'].replace(e1)
ox['E1_Flag'] = ox['E1_Flag'].replace(e_flag)
ox['E2_Debt/contract relief'] = ox['E2_Debt/contract relief'].replace(e2)
ox['E3_Fiscal measures'] = ox['E3_Fiscal measures'].replace(e3)
ox['E4_International support'] = ox['E4_International support'].replace(e4)
ox['H1_Public information campaigns'] = ox['H1_Public information campaigns'].replace(h1)
ox['H1_Flag'] = ox['H1_Flag'].replace(h_flag)
ox['H2_Testing policy'] = ox['H2_Testing policy'].replace(h2)
ox['H3_Contact tracing'] = ox['H3_Contact tracing'].replace(h3)
ox['H4_Emergency investment in healthcare'] = ox['H4_Emergency investment in healthcare'].replace(h4)
ox['H5_Investment in vaccines'] = ox['H5_Investment in vaccines'].replace(h5)
ox['H6_Facial Coverings'] = ox['H6_Facial Coverings'].replace(h6)
ox['H6_Flag'] = ox['H6_Flag'].replace(h_flag)
ox['H7_Vaccination policy'] = ox['H7_Vaccination policy'].replace(h7)
ox['H7_Flag'] = ox['H7_Flag'].replace(h7_flag)
ox['H8_Protection of elderly people'] = ox['H8_Protection of elderly people'].replace(h8)
ox['H8_Flag'] = ox['H8_Flag'].replace(h_flag)
# -
def filter_df(query, df):
"""Searches in list of columns and
returns a filtered df"""
cols = [col for col in df.columns if 'Notes' in col]
mask = np.column_stack([df[col].str.contains(query, na=False, case=False) for col in cols])
df = df.loc[mask.any(axis=1)]
return df
filter_df('artificial', ox)
len(ox)
| notebooks/measure_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
requests.get('https://news.daum.net/economic#1')
path = 'https://news.daum.net/economic#1'
req = requests.get(path)
req.status_code
soup = BeautifulSoup(req.content, 'html.parser')
type(soup)
soup.select('div.box_timenews > ul.list_timenews > strong.tit_timenews > a[href].link_txt')
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img height="60px" src="http://astro.phys.wvu.edu/bhathome/images/NRPyPlusLogo.png" align="center" hspace="20px" vspace="5px">
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# ## The NRPy+ Tutorial: An Introduction to Python-Based Code Generation for Numerical Relativity... and Beyond!
#
# ### Lead author: [<NAME>](http://astro.phys.wvu.edu/zetienne/) $\leftarrow$ Please feel free to email comments, revisions, or errata!
#
# ***If you are unfamiliar with using Jupyter Notebooks, first review the official [Jupyter Notebook Basics Guide](https://nbviewer.jupyter.org/github/jupyter/notebook/blob/master/docs/source/examples/Notebook/Notebook%20Basics.ipynb).***
#
# ### PART 1: Basic Functionality of NRPy+, a First Application
# ##### NRPy+ Basics
# + [NRPy+: Introduction & Motivation](http://astro.phys.wvu.edu/bhathome/nrpy.html) (NRPy+ home page)
# + [Basic C Code Output, NRPy+'s Parameter Interface](Tutorial-Coutput__Parameter_Interface.ipynb)
# + [`cmdline_helper`: Multi-platform command-line helper functions](Tutorial-cmdline_helper.ipynb) (*Courtesy <NAME>*)
# + [Numerical Grids](Tutorial-Numerical_Grids.ipynb)
# + [Indexed Expressions (e.g., tensors, pseudotensors, etc.)](Tutorial-Indexed_Expressions.ipynb)
# + [Finite Difference Derivatives](Tutorial-Finite_Difference_Derivatives.ipynb)
# + Instructional notebook: [How NRPy+ Computes Finite Difference Derivative Coefficients](Tutorial-How_NRPy_Computes_Finite_Difference_Coeffs.ipynb)
# + **Start-to-Finish Example**: [Finite-Difference Playground: A Complete C Code for Validating NRPy+-Based Finite Differences](Tutorial-Start_to_Finish-Finite_Difference_Playground.ipynb)
# + Method of Lines for PDEs: Step PDEs forward in time using ODE methods
# + Solving ODEs using explicit Runge Kutta methods (*Courtesy <NAME>*)
# + [The Family of explicit Runge-Kutta methods and their Butcher tables](Tutorial-RK_Butcher_Table_Dictionary.ipynb)
# + [Validating Runge Kutta Butcher tables using truncated Taylor series](Tutorial-RK_Butcher_Table_Validation.ipynb)
# + [Generating C Code to implement Method of Lines timestepping with explicit Runge Kutta-like methods](Tutorial-Method_of_Lines-C_Code_Generation.ipynb) (*Courtesy <NAME>*)
# + Writing your own NRPy+ tutorial notebook
# + [The NRPy+ Tutorial Style Guide](Tutorial-Template_Style_Guide.ipynb) (*Courtesy <NAME>*)
# + [Adding Unit Tests](Tutorial-UnitTesting.ipynb) (*Courtesy <NAME>*)
#
# ### PART 2: Basic Physics Applications
# ##### Using NRPy+ to Numerically Solve PDEs
# + Application: [The Scalar **Wave Equation** in Cartesian Coordinates, with Plane-Wave Initial Data](Tutorial-ScalarWave.ipynb)
# + **Start-to-Finish Example**: [Numerically Solving the Scalar Wave Equation: A Complete C Code](Tutorial-Start_to_Finish-ScalarWave.ipynb)
# + Solving the Wave Equation with the <font color='green'>**Einstein Toolkit**</font> (*Courtesy <NAME>*)
# + [<font color='green'>**IDScalarWaveNRPy**</font>: Plane-wave initial data for the scalar wave equation](Tutorial-ETK_thorn-IDScalarWaveNRPy.ipynb)
# + [<font color='green'>**WaveToyNRPy**</font>: Solving the scalar wave equation, using the method of lines](Tutorial-ETK_thorn-WaveToyNRPy.ipynb)
#
# <html>
# <!-- + Application (<font color='red'>**in progress**</font>): [Two Formulations of **Maxwell's Equations** in Cartesian Coordinates](Tutorial-MaxwellCartesian.ipynb). (Formulations based on [Illustrating Stability Properties of Numerical Relativity in Electrodynamics](https://arxiv.org/abs/gr-qc/0201051) by Knapp, Walker, and Baumgarte.) (*Courtesy <NAME>*)
# + [<font color='green'>**IDMaxwellNRPy**</font>: An <font color='green'>**Einstein Toolkit**</font> initial data thorn for Maxwell's equations](Tutorial-ETK_thorn-IDMaxwellNRPy.ipynb)
# + [<font color='green'>**MaxwellEvol**</font>: Solving Maxwell's equations in the <font color='green'>**Einstein Toolkit**</font> using the method of lines](Tutorial-ETK_thorn-MaxwellEvol.ipynb)
# -->
# </html>
#
# ##### Diagnostic Notebooks: Gravitational Wave Extraction in Cartesian coordinates
# + Application: [All Weyl scalars and invariants in Cartesian Coordinates](Tutorial-WeylScalarsInvariants-Cartesian.ipynb) (*Courtesy <NAME>*)
# + [<font color='green'>**WeylScal4NRPy**</font>: An <font color='green'>**Einstein Toolkit**</font> Diagnostic Thorn](Tutorial-ETK_thorn-Weyl_Scalars_and_Spacetime_Invariants.ipynb) (*Courtesy <NAME>*)
#
#
# ##### Solving the Effective-One-Body Equations of Motion
#
# + Application: [SEOBNR: The Spinning-Effective-One-Body-Numerical-Relativity Hamiltonian](Tutorial-SEOBNR_Documentation.ipynb)
# + Solving the SEOBNR Hamiltonian equations of motion (<font color='red'>**in progress**</font>)
# + [Initial data: Setting the initial trajectory](in_progress/Tutorial-SEOBNR_Initial_Conditions.ipynb)
# + [SymPy-generated exact derivatives of the SEOBNR Hamiltonian](in_progress/Tutorial-Spinning_Effective_One_Body_Numerical_Relativity_Hamiltonian-Cartesian.ipynb)
#
# ### PART 3: Solving PDEs in Curvilinear Coordinate Systems
# + [Moving beyond Cartesian Grids: Reference Metrics](Tutorial-Reference_Metric.ipynb)
# + Application: [The Scalar Wave Equation in Curvilinear Coordinates, using a Reference Metric](Tutorial-ScalarWaveCurvilinear.ipynb)
# + **Start-to-Finish Example**: [Numerically Solving the Scalar Wave Equation in Curvilinear Coordinates: A Complete C Code](Tutorial-Start_to_Finish-ScalarWaveCurvilinear.ipynb)
# + **Start-to-Finish Example**: [Implementation of Curvilinear Boundary Conditions, Including for Tensorial Quantities](Tutorial-Start_to_Finish-Curvilinear_BCs.ipynb)
#
# ### PART 4: Numerical Relativity $-$ BSSN in Curvilinear Coordinates
#
# + [**Overview: Covariant BSSN formulation of general relativity in curvilinear coordinates**](Tutorial-BSSN_formulation.ipynb)
# + [Construction of useful BSSN quantities](Tutorial-BSSN_quantities.ipynb)
# + [BSSN time-evolution equations](Tutorial-BSSN_time_evolution-BSSN_RHSs.ipynb)
# + [Time-evolution equations for BSSN gauge quantities $\alpha$ and $\beta^i$](Tutorial-BSSN_time_evolution-BSSN_gauge_RHSs.ipynb)
# + [Hamiltonian and momentum constraint equations](Tutorial-BSSN_constraints.ipynb)
# + [Enforcing the conformal 3-metric $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint](Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.ipynb)
# + [Writing quantities of ADM formalism in terms of BSSN quantities](Tutorial-ADM_in_terms_of_BSSN.ipynb)
# + **Initial data notebooks**. Initial data are set in terms of standard [ADM formalism](https://en.wikipedia.org/wiki/ADM_formalism) spacetime quantities.
# + [Non-Spinning ("static trumpet") black hole initial data](Tutorial-ADM_Initial_Data-StaticTrumpet.ipynb) (*Courtesy <NAME> & <NAME>*)
# + [Spinning UIUC black hole initial data](Tutorial-ADM_Initial_Data-UIUC_BlackHole.ipynb) (*Courtesy <NAME> & <NAME>*)
# + [Spinning Shifted Kerr-Schild black hole initial data](Tutorial-ADM_Initial_Data-ShiftedKerrSchild.ipynb) (*Courtesy <NAME>*)
# + [Brill-Lindquist initial data: Two-black-holes released from rest](Tutorial-ADM_Initial_Data-Brill-Lindquist.ipynb)
# + [Black hole accretion disk initial data (Fishbone-Moncrief)](Tutorial-FishboneMoncriefID.ipynb)
# + [<font color='green'>**FishboneMoncriefID**</font>: Setting up Fishbone-Moncrief disk initial data within the <font color='green'>**Einstein Toolkit**</font>, using HydroBase variables as input](Tutorial-ETK_thorn-FishboneMoncriefID.ipynb)
# + [Neutron Star initial data: The Tolman-Oppenheimer-Volkoff (TOV) solution](Tutorial-ADM_Initial_Data-TOV.ipynb) (*Courtesy <NAME>*)
# + [Implementation of Single and Piecewise Polytropic EOSs](Tutorial-TOV-Piecewise_Polytrope_EOSs.ipynb) (*Courtesy <NAME>*)
# + **ADM-to-curvilinear-BSSN initial data conversion**
# + [**Exact** ADM Spherical/Cartesian to BSSN Curvilinear Initial Data Conversion](Tutorial-ADM_Initial_Data-Converting_Exact_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb) (Use this module for initial data conversion if the initial data are known *exactly*. The BSSN quantity $\lambda^i$ will be computed exactly using SymPy from given ADM quantities.)
# + [**Start-to-Finish *exact* initial data validation notebook**](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_Exact_Initial_Data.ipynb): Confirms all exact initial data types listed above satisfy Einstein's equations of general relativity. (*Courtesy <NAME> & <NAME>*)
# + [**Numerical** ADM Spherical/Cartesian to BSSN Curvilinear Initial Data Conversion](Tutorial-ADM_Initial_Data-Converting_Numerical_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb) (Use this module for initial data conversion if the initial data are provided by an initial data solver, and are thus known to roundoff error at best. The BSSN quantity $\lambda^i$ will be computed using finite-difference derivatives from given ADM quantities.)
# + [**Start-to-Finish *numerical* initial data validation notebook**: The TOV solution](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_TOV_initial_data.ipynb): Neutron star initial data, confirms numerical errors converge to zero at expected order (TOV initial data are generated via [the *numerical* solution of a system of ODEs](https://en.wikipedia.org/wiki/Tolman%E2%80%93Oppenheimer%E2%80%93Volkoff_equation), thus are known only numerically)
# + **Diagnostic curvilinear BSSN modules**
# + [The gravitational wave Weyl scalar $\psi_4$, in arbitrary curvilinear coordinates](Tutorial-Psi4.ipynb)
# + [Constructing the quasi-Kinnersley tetrad for $\psi_4$](Tutorial-Psi4_tetrads.ipynb)
# + [Start-to-Finish validation of above expressions in Cartesian coordinates, against Pat<NAME>son's Weyl Scalars & Invariants notebook](BSSN/Psi4Cartesianvalidation/Tutorial-Psi4-Cartesian_validation.ipynb)
# + [Spin-weighted spherical harmonics](Tutorial-SpinWeighted_Spherical_Harmonics.ipynb) (*Courtesy <NAME>*)
#
# + **Start-to-Finish curvilinear BSSN simulation examples**:
# + [<font color='purple'>**Colliding black holes!**</font>](Tutorial-Start_to_Finish-BSSNCurvilinear-Two_BHs_Collide.ipynb)
# + [The "Hydro without Hydro" test: evolving the spacetime fields of TOV star with $T^{\mu\nu}$ assumed static](Tutorial-Start_to_Finish-BSSNCurvilinear-Neutron_Star-Hydro_without_Hydro.ipynb) ***In progress***
#
# ### PART 5: Numerical Relativity $-$ General Relativistic Hydrodynamics (GRHD), Force-Free Electrodynamics (GRFFE), & Magnetohydrodynamics (GRMHD)
#
# + [The equations of general relativistic hydrodynamics (**GRHD**), in Cartesian coordinates](Tutorial-GRHD_Equations-Cartesian.ipynb)
# + [The equations of general relativistic, force-free electrodynamics (**GRFFE**), in Cartesian coordinates](Tutorial-GRFFE_Equations-Cartesian.ipynb)
# + [The equations of general relativistic magnetohydrodynamics (**GRMHD**), in Cartesian coordinates](Tutorial-GRMHD_Equations-Cartesian.ipynb)
# + [**`IllinoisGRMHD`** ](IllinoisGRMHD/doc/) with piecewise-polytrope equation of state support (*Courtesy <NAME>*): <font color='red'>***In progress***</font>
# + Diagnostic notebook: [<font color='green'>**sbPoynETNRPy**</font>: Evaluating $b^\mu$ and $S^i$ in the <font color='green'>**Einstein Toolkit**</font>, using HydroBase variables as input](Tutorial-ETK_thorn-u0_smallb_Poynting.ipynb)
# + Tutorial notebook: [Computing the 4-Velocity Time-Component $u^0$, the Magnetic Field Measured by a Comoving Observer $b^{\mu}$, and the Poynting Vector $S^i$](Tutorial-u0_smallb_Poynting-Cartesian.ipynb)
#
| NRPyPlus_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tf2_pcse)
# language: python
# name: conda_tf2_pcse
# ---
# +
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
# -
# ### Running PCSE/WOFOST with custom input data
data_dir = './pcse_examples/'
# + tags=[]
from pcse.fileinput import CABOFileReader
cropfile = os.path.join(data_dir, 'sug0601.crop')
cropdata = CABOFileReader(cropfile)
print(cropdata)
# + tags=[]
soilfile = os.path.join(data_dir, 'ec3.soil')
soildata = CABOFileReader(soilfile)
print(soildata)
# -
from pcse.util import WOFOST71SiteDataProvider
sitedata = WOFOST71SiteDataProvider(WAV=100, CO2=360)
print(sitedata)
from pcse.base import ParameterProvider
parameters = ParameterProvider(cropdata=cropdata, soildata=soildata, sitedata=sitedata)
# pcse.base_classes가 아니라 pcse.base
from pcse.fileinput import YAMLAgroManagementReader
agromanagement_file = os.path.join(data_dir, 'sugarbeet_calendar.agro')
agromanagement = YAMLAgroManagementReader(agromanagement_file)
print(agromanagement)
from pcse.db import NASAPowerWeatherDataProvider
wdp = NASAPowerWeatherDataProvider(latitude=52, longitude=5)
print(wdp)
# 시간이 오래 걸림
from pcse.models import Wofost71_WLP_FD
wofsim = Wofost71_WLP_FD(parameters, wdp, agromanagement)
wofsim.run_till_terminate()
output = wofsim.get_output()
len(output)
varnames = ["day", "DVS", "TAGP", "LAI", "TWSO"]
tmp = {}
for var in varnames:
tmp[var] = [t[var] for t in output]
day = tmp.pop("day")
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(10,8))
for var, ax in zip(["DVS", "TAGP", "LAI", "TWSO"], axes.flatten()):
ax.plot_date(day, tmp[var], 'b-')
ax.set_title(var)
fig.autofmt_xdate()
fig.savefig('./results/sugarbeet.png')
| 4_PCSE_custom_data_tuto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sklearn
import pickle
from PIL import Image, ImageOps
import numpy as np
import os
from keras.preprocessing import image
# +
images_val_list = []
aux = []
for root, dirs, files in os.walk("/home/exla24/TB2018/val/images", topdown=False):
for name in files:
aux.append(name)
aux.sort()
# -
for i in range(len(aux)):
img = image.load_img("/home/exla24/TB2018/val/images/" + aux[i], target_size=(224,224))
images_val_list.append(img)
# +
images_test_list = []
aux1 = []
for root, dirs, files in os.walk("/home/exla24/TB2018/test/images", topdown=False):
for name in files:
aux1.append(name)
aux1.sort()
# -
for i in range(len(aux1)):
img = image.load_img("/home/exla24/TB2018/test/images/" + aux1[i], target_size=(224,224))
images_test_list.append(img)
# +
images_train_list = []
aux2 = []
for root, dirs, files in os.walk("/home/exla24/TB2018/train/images", topdown=False):
for name in files:
aux2.append(name)
aux2.sort()
# -
for i in range(len(aux2)):
img = image.load_img("/home/exla24/TB2018/train/images/" + aux2[i], target_size=(224,224))
images_train_list.append(img)
# +
import pickle
with open('/home/jupyter/Pickles/Imagenes/images_train.pickle', 'wb') as train:
pickle.dump(images_train_list, train, protocol=pickle.HIGHEST_PROTOCOL)
with open('/home/jupyter/Pickles/Imagenes/images_validation.pickle', 'wb') as validation:
pickle.dump(images_val_list, validation, protocol=pickle.HIGHEST_PROTOCOL)
with open('/home/jupyter/Pickles/Imagenes/images_test.pickle', 'wb') as test:
pickle.dump(images_test_list, test, protocol=pickle.HIGHEST_PROTOCOL)
| Carga_de_Imagenes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The development in productivity and hourly wages in Denmark across industries and years.
# ## Introduction
# The development in productivity is the underlying condition for a natural increase in wages, and therefore a substantial contributor to the overall wealth in society. This statement is something the _The Productivity Commision_ and organizations from different sites of the political spectrum (ex. _Danish Metal Workers Union_ and _Cepos_) agrees with, even though the calculations of this is used to assist different claims.
#
# The trends in hourly wage and hourly productivity is said to follow each other in the long run. If the increment in wage is bigger than the increment in productivity, it will be too expensive for companies to produce, and this will lead to lower production and lower employment. Rising unemployment will lead to a lower increase in overall earnings, and will at last lead to better competitiveness and therefore productivity.
#
# "_If wages are persistently lagging behind productivity, workers do not receive their fair share of the produced wealth. This is not only deeply unjust but also economically detrimental, as growth remains behind its potential. Labour income remains the main source of income for households and private consumption makes up the largest part of aggregate demand._" - **<NAME>**
#
# In this assignment, we investigate the relationship between productivity and hourly wage in Denmark. We want to investigate the relationship when we compare different industries to each other and the total trends. Some industries may naturally have a close relationship, where other industries may have a bigger deviation between wage and productivity.
#
#
# We measure productivity and hourly wage as shown in the two following equations:
#
#
#
#
# \\[
# \begin{eqnarray*}
# Productivity_{ij} = \frac{Gross\:value\:added_{ij}}{Hours\:worked_{ij}}
# \\
# \\
# Hourly\: wage_{ij} = \frac{Compensation\:of\:employees_{ij}}{Hours\:worked\:by\:employees_{ij}}
# \end{eqnarray*}
# \\]
#
#
# where ${i}$ denotes years and ${j}$ denotes industries.
#
# Moving on, we start by importing all the modules, we need for this notebook.
# ## Data managemnet
# **Import of packages:** We import packages to calculate and manipulate data (_numpy_ and _pandas_) and packages to plot our results (_matplot_ and _seaborn_). Lastly, we import _pydst_ which enables the use of _Statistics Denmark_ from where we get our relevant data.
# +
#Import of packages
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import ipywidgets as widgets
import pandas as pd
import pandas_datareader as pdr
import pandas_datareader.data as web
import datetime
import pydst
dst = pydst.Dst(lang='en')
# -
# From statistics Denmark we load data which contains _the Gross value added, the compensation of employees, the hours worked in total_ and _the hours worked by the employees_ in the different industries. More specifically we load two datasets;
#
# 1. NABP10: A dataset containing the gross value added and compensation of employees
# 2. NABB10: A dataset containing the hours worked by employees and hours worked in total
#
# We need to merge the two datasets to one, but first, we have to check the specific datasets, their dimensions, renaming, convert strings to floats, etc.
#
# **Imports of data:**
# +
# a. gettign an orientation in data
dst.get_variables('NABP10') # command to se 'id'
dst.get_variables('NABP10').iloc[0]['values'] # command to se 'values' in 'TRANSAKT'
# b. loading data
gva10 = dst.get_data(table_id = 'NABP10', variables={'TRANSAKT':['B1GD','D1D'],'BRANCHE':['*'], 'PRISENHED':['V'], 'TID':['*']})
emp10 = dst.get_data(table_id = 'NABB10', variables={'SOCIO':['EMPH_DC','SALH_DC'],'BRANCHE':['*'], 'TID':['*']})
# c. checking the dimensions
print(f'Industries in gva10 = {len(gva10.BRANCHE.unique())}')
print(f'Industries in emp10 = {len(emp10.BRANCHE.unique())}')
print(f'Years in gva10 = {len(gva10.TID.unique())}')
print(f'Years in emp10 = {len(emp10.TID.unique())}')
# -
# From table "NABP10" we do only read data in nominal prices, which is due to the fact that "Compensation of employees" is not given in chained prices. Below we have a quick look a the imported data:
# a. quick look at data
#gva10.head()
emp10.head()
# **Renaming variables:**
# +
# a. define dictionaries to rename variables
columns_dict = {}
columns_dict['TRANSAKT'] = 'variable'
columns_dict['BRANCHE'] = 'industry'
columns_dict['PRISENHED'] = 'unit'
columns_dict['TID'] = 'year'
columns_dict['INDHOLD'] = 'value'
columns_dict['SOCIO'] = 'employment'
var_dict = {} # var is for variable
var_dict['B.1g Gross value added'] = 'GVA'
var_dict['D.1 Compensation of employees'] = 'ComEmp'
var_dict['Hours worked (1,000 hours)'] = 'Hours_TOT'
var_dict['Hours worked for employees (1,000 hours)'] = 'Hours_Emp'
unit_dict = {}
unit_dict['2010-prices, chained values'] = 'real'
unit_dict['Current prices'] = 'nominal'
industry_dict = {}
industry_dict['A Agriculture, forestry and fishing'] = 'Agriculture, forestry and fishing'
industry_dict['B Mining and quarrying'] = 'Mining and quarrying'
industry_dict['C Manufacturing'] = 'Manufacturing'
industry_dict['D_E Utility services'] = 'Utility services'
industry_dict['F Construction'] = 'Construction'
industry_dict['G_I Trade and transport etc.'] = 'Trade and transport etc.'
industry_dict['J Information and communication'] = 'Information and communication'
industry_dict['K Financial and insurance'] = 'Financial and insurance'
industry_dict['LA Real estate activities and renting of non-residential buildings'] = 'Real estate activities and renting of non-residential buildings'
industry_dict['LB Dwellings'] = 'Dwellings'
industry_dict['M_N Other business services'] = 'Other business services'
industry_dict['O_Q Public administration, education and health'] = 'Public administration, education and health'
industry_dict['R_S Arts, entertainment and other services'] = 'Arts, entertainment and other services'
# b. renaming
gva10.rename(columns=columns_dict,inplace=True)
emp10.rename(columns=columns_dict,inplace=True)
for key,value in var_dict.items():
gva10.variable.replace(key,value, inplace=True)
emp10.employment.replace(key,value, inplace=True)
for key,value in industry_dict.items():
gva10.industry.replace(key,value, inplace=True)
emp10.industry.replace(key,value, inplace=True)
for key,value in unit_dict.items():
gva10.unit.replace(key,value, inplace=True)
# c. redefine value from 'str' to 'float'
emp10.value = emp10.value.astype('float')
gva10['value'] = gva10['value'].apply(pd.to_numeric, errors='coerce')
#gva10.head()
emp10.tail()
# -
# **Split:**
#
# If we merge our dataset, we will get a long dataset in which our values would be in rows, and we would like to have them in columns. We are aware of the fact that there may be an operation converting a dataset from long to wide. But due to the lack of skills, we do it the simple way. We split our two datasets into four, and merge our relevant values together.
# +
# a. split dataframe emp10 in two - hours total and hours for employees.
H_TOT = emp10.loc[emp10.employment == ('Hours_TOT')]
H_TOT.rename(columns = {'value':'H_TOT'}, inplace=True)
H_Emp = emp10.loc[emp10.employment == ('Hours_Emp')]
H_Emp.rename(columns = {'value':'H_EMP'}, inplace=True)
# b. split dataframe gva10 in two - GVA and compensation for employees
GVA = gva10.loc[gva10.variable == ('GVA')]
GVA.rename(columns = {'value':'GVA'}, inplace=True)
ComEmp = gva10.loc[gva10.variable == ('ComEmp')]
ComEmp.rename(columns = {'value':'ComEmp'}, inplace=True)
#GVA.head()
H_TOT.head()
# +
# pass command if already ran once
# dropper 'employment' and 'variable'
H_TOT.drop(columns=['employment'], inplace = True)
H_Emp.drop(columns=['employment'], inplace = True)
GVA.drop(columns=['variable'], inplace = True)
ComEmp.drop(columns=['variable'], inplace = True)
# -
H_TOT.head()
# **Merge:**
#
# Finally we merge our data, sorted on year and industry. Doing it this way, we can calculate the productivity and hourly wage, in a way where we can compare the relationship over time in the different industries.
# +
# a. merged data
merged = pd.merge(GVA,ComEmp,how='left',on=['year','industry'])
merged1 = pd.merge(merged,H_TOT,how='left',on=['year','industry'])
df = pd.merge(merged1,H_Emp,how='left',on=['year','industry'])
df.drop(columns=['unit_y'], inplace = True)
# b. redefine value from 'str' to 'float'
df.GVA = df.GVA.astype('float')
df.ComEmp = df.ComEmp.astype('float')
df
# c. calculate the productivity and the hourly wage given the equations from the introduction.
# We multipy with 1000 since GVA and ComEmp is measured as millions while H_TOT and H_EMP is measured as thousand.
df['Productivity'] = df.GVA / df.H_TOT*1000
df['Hour_wage'] = df.ComEmp / df.H_EMP*1000
# d. checking the dimensions the
print(f'Years in main dataset "df_index" = {len(df.year.unique())}')
print(f'Industries in main dataset "df_index" = {len(df.industry.unique())}')
df.head()
# -
# We now have the dataset in the shape we want. We can see that the dimensions are the same as from the beginning - which implies that we did not lose any observations through the data manipulation.
# **Index:**
# We see that the values of the industries are very different in hourly wage and productivity. This makes sense since the wage-level and productivity in the industries differ. We want to compare the relationship between the industries over time - therefore we normalize the values. We normlize with respect to year 2010.
# +
# a. Create a copy
df_index = df.copy()
df_index.sort_values(['industry','year'], inplace=True)
# b. Normalizing by 2010
df_index['Pro_index'] = df_index.groupby(['industry'])['Productivity'].transform(lambda x: x/x.iloc[44]*100)
df_index['H_index'] = df_index.groupby(['industry'])['Hour_wage'].transform(lambda x: x/x.iloc[44]*100)
# c. Creat a diff-variable, which takes the different between the productivity and the hourly wage
df_index['Diff'] = df_index['Pro_index'] - df_index['H_index']
# d. view the index.
# df_index.tail(62) #following command show that the index start over again for each industry
df_index.tail(10)
# -
# ## Plots
#
# To follow up on the results, we plot the trends of productivity and hourly wage in the industries. First, we tried to plot all in one graph, but since the category of industries is quite big, "one graph to fit them all" will look messy. Based on that experience, we make use of ``Seaborn FacedGrid``, where we plot each industry in a separate graph.
# +
by_var = 'industry'
y_var = 'year'
g = sns.FacetGrid(df_index, col=by_var, hue=by_var, col_wrap=3, sharex=True, sharey=True, height=8, aspect=1)
g = (g.map(plt.plot, 'year', 'Pro_index', color='y')) # draw the upper line
g = (g.map(plt.plot, 'year', 'H_index', color='m')) # draw the upper line
g = g.set_titles("{col_name}") # # Control the title of each facet
g = g.set_ylabels('Index, base year = 2010 ')
print(f'The yellow graph plots the productivity.')
print(f'The magenta graph plots the hourly wage.')
# -
# As we see, trends of productivity and hourly wage in total seem to follow each other, where agriculture, mining, and utility services seem to bump a bit in productivity over the years. If we plot the difference of the productivity and hourly wage, we get a more simple look on the development. It becomes easier to evaluate if some industries has get paid to much or to little over the years. Remember that the diff-values is $Productivity\:Index - Hourly\:wage\:Index$. So if the trend is above zero, the increase in hourly wage is behind the increase in productivity - meaning that the employees should try to negotiate for a higher wage.
#
# Below, we illustrate the diff-values in a dropdown-graph. It is possible to make a comparison between an industry, chosen in the dropdown menu, with the development in the economy as a whole (Total).
# Set 'industry' as index in a new DataFrame
df_widget = df_index.copy()
df_widget = df_widget.set_index('industry')
df_widget = df_widget[['year','Diff']]
# +
# a. define the function which plot the graph
def _plot_timeseries(dataframe, industry):
fig = plt.figure(dpi=100)
ax = fig.add_subplot(1,1,1)
# comparisons with 'Total'
x_total = dataframe.loc['Total', 'year']
y_total = dataframe.loc['Total', 'Diff']
# unique industry
x = dataframe.loc[industry, 'year']
y = dataframe.loc[industry, 'Diff']
ax.set(xlabel='Years', ylabel='Diff', title='Comparisons of a unique industry with Total' )
ax.plot(x,y, label = industry)
ax.plot(x_total,y_total, label = 'Total')
ax.grid()
plt.legend(bbox_to_anchor=(0.5, -0.3), loc=8, ncol=4)
# b. define a function which easy fill-out the abow function and include the dropdown menu.
def plot_timeseries(dataframe):
widgets.interact(_plot_timeseries,
dataframe = widgets.fixed(dataframe),
industry = widgets.Dropdown(
#index=dataframe.loc[value],
description='Industry',
options= dataframe.index.unique(),
value='Manufacturing',
disabled=False)
);
plot_timeseries(df_lol)
# -
# Inspection of different plots; The hourly wage in the industries - 'Contruction' and 'Manufacturing', does not correspond with the increase in the productivity the past couple of years. Differently, the 'Agricultural, Fishing and forestry' sector, seems to have a volatile realtionship which may be due to the fact that these industries depends highly on the volatility in the seasons.
# In the industries - 'Information and communication' and 'Utility services' - does the productivity lags behind the wages.
# If one takes a look on the difference between productivity and hourly wage in _'the general goverment'_ sector, we see that it is nearly zero all time. This makes sence since it is hard to measure the gross value added, and because of the dynamic wage-process in this sector due to the "Reguleringsordningen".
#
#
# **Conclusion**
#
# We can therefore conclude, that the overall trends of productivity and hourly wage follows each other. Employees in the 'Manufacturing' and 'Construction' industries, should negotiate for a higher hourly wage, whereas the sector of 'Information and communication' should aim for a higher productivity.
| dataproject/datapro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## POS tagging using modified Viterbi
# ### Data Preparation
#Importing libraries
import nltk
import numpy as np
import pandas as pd
import pprint, time
import random
from sklearn.model_selection import train_test_split
from nltk.tokenize import word_tokenize
import math
import matplotlib.pyplot as plt
import seaborn as sns
# supress warnings
import warnings
warnings.filterwarnings('ignore')
# reading the Treebank tagged sentences
nltk_data = list(nltk.corpus.treebank.tagged_sents(tagset='universal'))
#taking 3 sents.
nltk_data[:3]
# converting the list of sent's to a list of (word, pos tag) tuples
tagged_words = [tup for sent in nltk_data for tup in sent]
print(len(tagged_words))
tagged_words[:10]
# +
#set the random seed
random.seed(1234)
#Divide the data into training and test sets
train_set, val_set = train_test_split(nltk_data,train_size=.95)
#Get the length of training and test sets
print(len(train_set))
print(len(val_set))
# -
#list of tagged words in train set
train_tag_words = [tup for sent in train_set for tup in sent]
#Get the length of the total tagged words in training set
len(train_tag_words)
#looking first 10 tokens in the train set
tokens = [pair[0] for pair in train_tag_words]
tokens[:10]
#the total unique words in the train set
V = set(tokens)
print(len(V))
#no. of pos tags in the train corpus
T = set([pair[1] for pair in train_tag_words])
print(len(T))
print(T)
# +
# 12 tags which is already a part of universal tagset.
# -
# ### Build the vanilla Viterbi based POS tagger
# computing P(w/t) and storing in T x V matrix
t = len(T)
v = len(V)
w_given_t = np.zeros((t, v))
# compute word given tag: Emission Probability
def word_given_tag(word, tag, train_bag = train_tag_words):
tag_list = [pair for pair in train_bag if pair[1]==tag]
count_tag = len(tag_list)
w_given_tag_list = [pair[0] for pair in tag_list if pair[0]==word]
count_w_given_tag = len(w_given_tag_list)
return (count_w_given_tag, count_tag)
#checking emission probabilities.
print("\n", "charges")
print(word_given_tag('charges', 'ADJ'))
print(word_given_tag('charges', 'VERB'))
print(word_given_tag('charges', 'NOUN'), "\n")
# +
# compute tag given tag: tag2(t2) given tag1 (t1), i.e. Transition Probability
def t2_given_t1(t2, t1, train_bag = train_tag_words):
tags = [pair[1] for pair in train_bag]
count_t1 = len([t for t in tags if t==t1])
count_t2_t1 = 0
for index in range(len(tags)-1):
if tags[index]==t1 and tags[index+1] == t2:
count_t2_t1 += 1
return (count_t2_t1, count_t1)
# -
#checking transition probabilities
print(t2_given_t1(t2='ADP', t1='ADJ'))
print(t2_given_t1('NOUN', 'ADJ'))
print(t2_given_t1('NOUN', 'DET'))
print(t2_given_t1('ADV', 'VERB'))
# +
# creating t x t transition matrix of tags
# each column is t2, each row is t1
# thus M(i, j) represents P(tj given ti)
tags_matrix = np.zeros((len(T), len(T)), dtype='float32')
for i, t1 in enumerate(list(T)):
for j, t2 in enumerate(list(T)):
tags_matrix[i, j] = t2_given_t1(t2, t1)[0]/t2_given_t1(t2, t1)[1]
# -
# convert the matrix to a df for better readability
tags_df = pd.DataFrame(tags_matrix, columns = list(T), index=list(T))
tags_df
tags_df.loc['.', :]
# heatmap of tags matrix
# T(i, j) means P(tag j given tag i)
plt.figure(figsize=(18, 12))
sns.heatmap(tags_df)
plt.show()
# frequent tags
# filter the df to get P(t2, t1) > 0.5
tags_frequent = tags_df[tags_df>0.5]
plt.figure(figsize=(18, 12))
sns.heatmap(tags_frequent)
plt.show()
# +
# from the heat it is clearly shown (NOUN and ADJ) and (NOUN and DET) are highly correlated.
# -
#lenght of train tag words
len(train_tag_words)
# +
# Viterbi Heuristic
def Viterbi(words, train_bag = train_tag_words):
state = []
T = list(set([pair[1] for pair in train_bag]))
for key, word in enumerate(words):
#initialise list of probability column for a given observation
p = []
for tag in T:
if key == 0:
transition_p = tags_df.loc['.', tag]
else:
transition_p = tags_df.loc[state[-1], tag]
# compute emission and state probabilities
emission_p = word_given_tag(words[key], tag)[0]/word_given_tag(words[key], tag)[1]
state_probability = emission_p * transition_p
p.append(state_probability)
pmax = max(p)
# getting state for which probability is maximum
state_max = T[p.index(pmax)]
state.append(state_max)
return list(zip(words, state))
# +
# Running on entire test dataset would take more than 3-4hrs.
# Let's test our Viterbi algorithm on a few sample sentences of test dataset
random.seed(1234)
# choose random 10 sents
rndom = [random.randint(1,len(val_set)) for x in range(10)]# change range to 5
# list of sents
test_run = [val_set[i] for i in rndom]
# list of tagged words
test_run_base = [tup for sent in test_run for tup in sent]
# list of untagged words
test_tag_words = [tup[0] for sent in test_run for tup in sent]
test_run
# -
# tagging the test sentences
start = time.time()
tagged_seq = Viterbi(test_tag_words)
end = time.time()
difference = end-start
print(difference)
# Get accuracy of model
#getting around 86% of accuracy with the default Viterbi algorithm.
check = [i for i, j in zip(tagged_seq, test_run_base) if i == j]
accuracy = len(check)/len(tagged_seq)
print(accuracy)
#check for incorrect tagged words
incorrect_tag_cases = [[test_run_base[i-1],j] for i, j in enumerate(zip(tagged_seq, test_run_base)) if j[0]!=j[1]]
incorrect_tag_cases
len(incorrect_tag_cases)
# ### Solve the problem of unknown words
#Read the text file using list
import codecs
test_line=[]
f = codecs.open("Test_sentences.txt", encoding='utf-8')
for line in f:
print(line)
test_line.append(line)
#getting sample test set
test_line
#Strip the special characters and empty strings in the list file
test_string =[sent.strip() for sent in test_line if sent.strip()!='']
test_string
# predictions on the test sentences
sample_pred_list =[]
for line in test_string:
sample_pred_list=sample_pred_list+list(Viterbi(word_tokenize(line)))
sample_pred_list
# +
#from above text sentence there are words that is incorrectly by POS tags.
# -
#the tokens in the test file
sample_words=[tokens for line in test_string for tokens in word_tokenize(line)]
#list of words which are present in test lines but not in the training corpus
words_not_in_corpus = list(set(sample_words) - set(tokens))
words_not_in_corpus
len(words_not_in_corpus)
#the tags predicted for unknown words
[tup for tup in sample_pred_list for word in words_not_in_corpus if tup[0]==word ]
#Let's count of all tags in the training set
from collections import Counter
tag_counts = Counter(pair[1] for pair in train_tag_words)
tag_counts
# the most common tags can in the training corpus
tag_counts.most_common(5)
#Lets count the incorrectly classfied words for each tag
incorrect_tag_counts = Counter(pair[0][1] for pair in incorrect_tag_cases)
incorrect_tag_counts
# Let check the percentage of verbs which are classifed as 'VERB' which end with 'ed'
#how many words with the tag 'VERB' (verb, past tense) ends with 'ed'
past_tense_verbs = [pair for pair in train_tag_words if pair[1]=='VERB']
ed_verbs = [pair for pair in past_tense_verbs if pair[0].endswith('ed')]
print(len(ed_verbs) / len(past_tense_verbs))
ed_verbs[:20]
# +
# around 20% of words which ends with 'ed' are verb.
# -
# Let check the percentage of verbs which are classifed as 'VERB' which end with 'ing'
#how many words with the tag 'VERB' ends with 'ing'
participle_verbs = [pair for pair in train_tag_words if pair[1]=='VERB']
ing_verbs = [pair for pair in participle_verbs if pair[0].endswith('ing')]
print(len(ing_verbs) / len(participle_verbs))
ing_verbs[:20]
# +
#around 10% of words which are ends with 'ing' are verbs.
# +
#Lets check the percentage of Adjective tags followed by NOUNS
# create a list of all tags (without the words)
tags = [pair[1] for pair in train_tag_words]
# create a list of Adj tags
adj_tags = [t for t in tags if t == 'ADJ']
# create a list of (ADJ, Noun) tags
adj_nn_tags = [(t, tags[index+1]) for index, t in enumerate(tags)
if t=='ADJ' and tags[index+1]=='NOUN']
print(len(adj_tags))
print(len(adj_nn_tags))
print(len(adj_nn_tags) / len(adj_tags))
# +
#around 70% of adj's are followed by noun's.
# +
#Lets check the percentage of Determinent tags followed by NOUN's
dt_tags = [t for t in tags if t == 'DET']
dt_nn_tags = [(t, tags[index+1]) for index, t in enumerate(tags)
if t=='DET' and tags[index+1]=='NOUN']
print(len(dt_tags))
print(len(dt_nn_tags))
print(len(dt_nn_tags) / len(dt_tags))
# +
#around 64% of det's followed by noun'd
# +
#Lets check the percentage of Adverbs tags followed by VERB's
md_tags = [t for t in tags if t == 'ADV']
md_vb_tags = [(t, tags[index+1]) for index, t in enumerate(tags)
if t=='ADV' and tags[index+1]=='VERB']
print(len(md_tags))
print(len(md_vb_tags))
print(len(md_vb_tags) / len(md_tags))
# +
#around 34% of adverb's are followed by verb's.
# -
# ### 1. Combining viterbi with n-gram tagging using BrillTagger
# +
# specify Rule-Based regex patterns for tagging
patterns = [
(r'.*ing$', 'VERB'), # gerund
(r'.*ed$', 'VERB'), # past tense verbs
(r'.*es$', 'VERB'), # singular present verbs
(r'.*ould$', 'VERB'), # modal verbs
(r'.*\'s$', 'NOUN'), # possessive nouns
(r'.*s$', 'NOUN'), # plural nouns
(r'^-?[0-9]+(.[0-9]+)?$', 'NUM'), # cardinal numbers
(r'(The|the|A|a|An|an)$', 'DET'), # articles or determinants
(r'.*able$', 'ADJ'), # adjectives
(r'.*ness$', 'NOUN'), # nouns formed from adjectives
(r'.*ly$', 'ADV'), # adverbs
(r'.*', 'NOUN') # nouns
]
# +
# rule based Unigram tagger
rule_based_tagger = nltk.RegexpTagger(patterns)
# lexicon backed up by the rule-based tagger
lexicon_tagger = nltk.UnigramTagger(train_set, backoff=rule_based_tagger)
#prediction on test set
lexicon_tagger.evaluate(val_set)
# +
# rule based Bigram tagger
#bigram_tagger = nltk.RegexpTagger(patterns)
# lexicon backed up by the rule-based tagger
lexicon_tagger2 = nltk.BigramTagger(train_set, backoff=lexicon_tagger)
#evaluation on test set
lexicon_tagger2.evaluate(val_set)
# +
# rule based Trigram tagger
#trigram_tagger = nltk.RegexpTagger(patterns)
# lexicon backed up by the rule-based tagger
lexicon_tagger3 = nltk.TrigramTagger(train_set, backoff=lexicon_tagger2)
#evaluation on test set
lexicon_tagger3.evaluate(val_set)
# -
#https://www.nltk.org/book/ch03.html
#http://www.nltk.org/book_1ed/ch05.html
#Trigram(t3) tagger backed by Bigram(t2) backed by Unigram(t1) which is backed by rule based tagger(t0)
t0 = nltk.RegexpTagger(patterns)
t1 = nltk.UnigramTagger(train_set, backoff=t0)
t2 = nltk.BigramTagger(train_set, backoff=t1)
t3 = nltk.TrigramTagger(train_set, backoff=t2)
# +
# if unknown word is the first word of a sentence, use unigram probability
# if unknown word is the second word of a sentence, use bigram probability
# if unknown word is present in any other position of a sentence, use trigram probability
# +
#BrillTagger Implementation
#https://docs.huihoo.com/nltk/0.9.5/api/nltk.tag.brill.BrillTaggerTrainer-class.html
#https://docs.huihoo.com/nltk/0.9.5/api/nltk.tag.api.TaggerI-class.html#tag
#https://stackoverflow.com/questions/32106090/nltk-brill-tagger-splitting-words
# -
# #### Evaluating tagging accuracy
# +
#https://streamhacker.com/2008/12/03/part-of-speech-tagging-with-nltk-part-3/
#https://www.nltk.org/_modules/nltk/tag/brill_trainer.html
#http://www.nltk.org/_modules/nltk/tag/brill_trainer_orig.html
#https://www.nltk.org/api/nltk.tag.html
#Import libraries for brill
import nltk.tag
from nltk.tag import brill
from nltk.tag.brill import *
from nltk.tbl.template import Template
from nltk.tag import BrillTaggerTrainer
#Clear templates created in earlier tests
Template._cleartemplates()
#Return 37 templates taken from the pos tagging task of the fntbl distribution
templates = fntbl37()
#Train the Brill model using trigram state(t3)
trainer = BrillTaggerTrainer(t3, templates)
brill_tagger = trainer.train(train_set, max_rules=100, min_score=3)
# -
# Brill tagging is a kind of transformation-based learning. The general idea is very simple: guess the tag of each word, then go back and fix the mistakes. In this way, a Brill tagger successively transforms a bad tagging of a text into a better one. As with n-gram tagging, this is a supervised learning method, since we need annotated training data to figure out whether the tagger's guess is a mistake or not. However, unlike n-gram tagging, it does not count observations but compiles a list of transformational correction rules.
# - The state probabilities for unknown words become zero and hence the Viterbi algorithm updates the unknown words by default with the first tag in the training set.
# Viterbi Heuristic
def Viterbi_updated(words, train_bag = train_tag_words):
state = []
T = list(set([pair[1] for pair in train_bag]))
for key, word in enumerate(words):
#initialise list of probability column for a given observation
p = []
for tag in T:
if key == 0:
transition_p = tags_df.loc['.', tag]
else:
transition_p = tags_df.loc[state[-1], tag]
# compute emission and state probabilities
emission_p = word_given_tag(words[key], tag)[0]/word_given_tag(words[key], tag)[1]
state_probability = emission_p * transition_p
p.append(state_probability)
pmax = max(p)
#Check if state probability is zero
if(pmax==0.0):
#If state probability is zero i.e if the word is unknown it updates the tag based on the brill tagger
state_max = brill_tagger.tag([word])[0][1]
else:
# getting state for which probability is maximum
state_max = T[p.index(pmax)]
state.append(state_max)
return list(zip(words, state))
# +
# validation_set
# Let's test our Viterbi algorithm on a few sample sentences of validation set
random.seed(1234)
# choose random 10 sents
rndom1 = [random.randint(1,len(val_set)) for x in range(10)]#change range to 5
# list of sents
test_run1 = [val_set[i] for i in rndom]
# list of tagged words
test_run_base1 = [tup for sent in test_run1 for tup in sent]
# list of untagged words
test_tag_words1 = [tup[0] for sent in test_run1 for tup in sent]
# -
# ### Compare the tagging accuracies of the modifications with the vanilla Viterbi algorithm
#checking the test sentences with the updated Viterbi algorithm
start = time.time()
updated_tagged_seq1 = Viterbi_updated(test_tag_words1)
end = time.time()
difference = end-start
#Print total time taken to train the algorithm
print(difference)
# Checking the accuracy of the updated Viterbi algorithm
check = [i for i, j in zip(updated_tagged_seq1, test_run_base1) if i == j]
accuracy = len(check)/len(updated_tagged_seq1)
accuracy
# ### List down cases which were incorrectly tagged by original POS tagger and got corrected by your modifications
#Let's print the test string with the original Viterbi algorithm
for line in test_string:
print(list(Viterbi(word_tokenize(line))))
#Let's print the test string with the updated Viterbi algorithm
for line in test_string:
print(list(Viterbi_updated(word_tokenize(line))))
# #### Results
# - Here Android, Google are unknown word incorrectly tagged in the original viterbi algorithm. In the updated viterbi algorithm they are correctly classified as 'NOUN'.
# - 2011, 2015 are unknown word, incorrectly tagged in the orginal viterbi algorithm. In the updated viterbi algorithm they are correctly classified as 'NUM' as cardinal numbers.
# - NASA, ICESAT-2 are unknown word incorrectly tagged in the orignal viterbi algorithm. In the updated viterbi algorithm they are correctly classified as 'NOUN'
# - Domineering and messages are unknown word incorrectly tagged in the original viterbi algorithm. In the updated viterbi algorithm they are correctly tagged as 'VERB'
# ### 2. Combine viterbi with rule based tagging
def rule_based_tagger(word):
# define regular expression patterns
patterns = [
(r'.*(ing|ed|es|ould)$', 'VERB'),
(r'.*\'s$', 'NOUN'),
(r'.*s$', 'NOUN'),
(r'^[A-Z]+.*$', 'NOUN'),
(r'^-?[0-9]+(.[0-9]+)?-?(.*)?$', 'NUM'),
(r'.*', 'NOUN') # nouns (default)
]
# using nltk.RegexpTagger
regexp_tagger = nltk.RegexpTagger(patterns)
regex_tag = regexp_tagger.tag([word])
return regex_tag[0][1]
def Viterbi_With_Regex(words, train_bag = train_tag_words):
state = []
T = list(set([pair[1] for pair in train_bag]))
for key, word in enumerate(words):
#initialise list of probability column for a given observation
p = []
for tag in T:
if key == 0:
transition_p = tags_df.loc['.', tag]
else:
try:
transition_p = tags_df.loc[state[-1], tag]
except:
transition_p = 0 # no previous state is found.
# compute emission and state probabilities
emission_p = word_given_tag(words[key], tag)[0]/word_given_tag(words[key], tag)[1]
state_probability = emission_p * transition_p
p.append(state_probability)
pmax = max(p)
if pmax == 0.0 :
#If state probability is zero i.e if the word is unknown it updates the tag based on the rule-based
state_max = rule_based_tagger(word)
else:
# getting state for which probability is maximum
state_max = T[p.index(pmax)]
state.append(state_max)
return list(zip(words, state))
# +
# validation_set
# Let's test our Viterbi algorithm on a few sample sentences of validation set
random.seed(1234)
# choose random 10 sents
rndom2 = [random.randint(1,len(val_set)) for x in range(10)]#change range to 5.
# list of sents
test_run2 = [val_set[i] for i in rndom]
# list of tagged words
test_run_base2 = [tup for sent in test_run2 for tup in sent]
# list of untagged words
test_tag_words2 = [tup[0] for sent in test_run2 for tup in sent]
# -
# tagging the test sentences with the updated Viterbi algorithm
start = time.time()
updated_tagged_seq2 = Viterbi_With_Regex(test_tag_words2)
end = time.time()
difference = end-start
#Print total time taken to train the algorithm
print(difference)
# ##### Accuracy for the updated Viterbi with rule-based algorithm
#Calculate the accuracy for the updated Viterbi with rule-based algorithm
check = [i for i, j in zip(updated_tagged_seq2, test_run_base2) if i == j]
accuracy = len(check)/len(updated_tagged_seq2)
accuracy
# #### Compare incorrectly tagged by original POS Tagger and got correctly by modify POS tagger using rule-based tagger
#Let's print the test string with the original Viterbi algorithm
for line in test_string:
print(list(Viterbi(word_tokenize(line))))
#checking updated regex algorithm on Sample test set
for line in test_string:
print(list(Viterbi_With_Regex(word_tokenize(line))))
# ### Results :
# - Android and Google are correctly updated as NOUN.
# - 2011, 2013, 2015 are correctly updated as NUM.
# - Message, Domineering are correctly updated as VERB.
| HMM+-based+POS+tagging-+Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="0259a7ce8120"
# # Vertex AI: Create, train, and deploy an AutoML text classification model
#
# ## Learning Objective
#
# In this notebook, you learn how to:
#
# * Create a dataset and import data
# * Train an AutoML model
# * Get and review evaluations for the model
# * Deploy a model to an endpoint
# * Get online predictions
# * Get batch predictions
#
# ## Introduction
#
# This notebook walks you through the major phases of building and using a text classification model on [Vertex AI](https://cloud.google.com/vertex-ai/docs/). In this notebook, you use the "Happy Moments" sample dataset to train a model. The resulting model classifies happy moments into categories that reflect the causes of happiness.
#
# Each learning objective will correspond to a __#TODO__ in the [student lab notebook](../labs/automl_text_classification.ipynb) -- try to complete that notebook first before reviewing this solution notebook.
#
# **Make sure to enable the Vertex AI, Cloud Storage, and Compute Engine APIs.**
# + [markdown] id="db52a0a61fca"
# ### Install additional packages
#
# This notebook uses the Python SDK for Vertex AI, which is contained in the `python-aiplatform` package. You must first install the package into your development environment.
# + id="b75757581291"
# Setup your dependencies
import os
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# Google Cloud Notebook requires dependencies to be installed with '--user'
USER_FLAG = ""
if IS_GOOGLE_CLOUD_NOTEBOOK:
USER_FLAG = "--user"
# Upgrade the specified package to the newest available version
# ! pip install {USER_FLAG} --upgrade google-cloud-aiplatform google-cloud-storage jsonlines
# -
# Please ignore any incompatibility warnings.
#
# **Restart** the kernel before proceeding further (On the Notebook menu - Kernel - Restart Kernel).
#
# + [markdown] id="WReHDGG5g0XY"
# ### Set your project ID
#
# Finally, you must initialize the client library before you can send requests to the Vertex AI service. With the Python SDK, you initialize the client library as shown in the following cell. This tutorial also uses the Cloud Storage Python library for accessing batch prediction results.
#
# Be sure to provide the ID for your Google Cloud project in the `project` variable. This notebook uses the `us-central1` region, although you can change it to another region.
#
# **If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
# + id="oM1iC_MfAts1"
# import necessary libraries
import os
from datetime import datetime
import jsonlines
from google.cloud import aiplatform, storage
from google.protobuf import json_format
PROJECT_ID = "qwiklabs-gcp-00-09d98f4803b0"
REGION = "us-central1"
# Get your Google Cloud project ID from gcloud
if not os.getenv("IS_TESTING"):
# shell_output=!gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID: ", PROJECT_ID)
aiplatform.init(project=PROJECT_ID, location=REGION)
# + [markdown] id="32c971919605"
# ## Create a dataset and import your data
#
# The notebook uses the 'Happy Moments' dataset for demonstration purposes. You can change it to another text classification dataset that [conforms to the data preparation requirements](https://cloud.google.com/vertex-ai/docs/datasets/prepare-text#classification).
#
# Using the Python SDK, you can create a dataset and import the dataset in one call to `TextDataset.create()`, as shown in the following cell.
#
# Creating and importing data is a long-running operation. This next step can take a while. The sample waits for the operation to complete, outputting statements as the operation progresses. The statements contain the full name of the dataset that you will use in the following section.
#
# **Note**: You can close the noteboook while you wait for this operation to complete.
# + id="6caf82e5e84e"
# TODO
# Use a timestamp to ensure unique resources
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
src_uris = "gs://cloud-ml-data/NL-classification/happiness.csv"
display_name = f"e2e-text-dataset-{TIMESTAMP}"
# + id="d35b8b6b94ae"
# TODO
# create a dataset and import the dataset
ds = aiplatform.TextDataset.create(
display_name=display_name,
gcs_source=src_uris,
import_schema_uri=aiplatform.schema.dataset.ioformat.text.single_label_classification,
sync=True,
)
# + [markdown] id="5b3cc427353a"
# ## Train your text classification model
#
# Once your dataset has finished importing data, you are ready to train your model. To do this, you first need the full resource name of your dataset, where the full name has the format `projects/[YOUR_PROJECT]/locations/us-central1/datasets/[YOUR_DATASET_ID]`. If you don't have the resource name handy, you can list all of the datasets in your project using `TextDataset.list()`.
#
# As shown in the following code block, you can pass in the display name of your dataset in the call to `list()` to filter the results.
#
# + id="52cf56f1c8a9"
# TODO
# list all of the datasets in your project
datasets = aiplatform.TextDataset.list(filter=f'display_name="{display_name}"')
print(datasets)
# + [markdown] id="58df3e02df82"
# When you create a new model, you need a reference to the `TextDataset` object that corresponds to your dataset. You can use the `ds` variable you created previously when you created the dataset or you can also list all of your datasets to get a reference to your dataset. Each item returned from `TextDataset.list()` is an instance of `TextDataset`.
#
# The following code block shows how to instantiate a `TextDataset` object using a dataset ID. Note that this code is intentionally verbose for demonstration purposes.
# + id="aa667203da03"
# Get the dataset ID if it's not available
dataset_id = "7829200088927830016"
if dataset_id == "7829200088927830016":
# Use the reference to the new dataset captured when we created it
dataset_id = ds.resource_name.split("/")[-1]
print(f"Dataset ID: {dataset_id}")
text_dataset = aiplatform.TextDataset(dataset_id)
# + [markdown] id="68f10356cab9"
# Now you can begin training your model. Training the model is a two part process:
#
# 1. **Define the training job.** You must provide a display name and the type of training you want when you define the training job.
# 2. **Run the training job.** When you run the training job, you need to supply a reference to the dataset to use for training. At this step, you can also configure the data split percentages.
#
# You do not need to specify [data splits](https://cloud.google.com/vertex-ai/docs/general/ml-use). The training job has a default setting of training 80%/ testing 10%/ validate 10% if you don't provide these values.
#
# To train your model, you call `AutoMLTextTrainingJob.run()` as shown in the following snippets. The method returns a reference to your new `Model` object.
#
# As with importing data into the dataset, training your model can take a substantial amount of time. The client library prints out operation status messages while the training pipeline operation processes. You must wait for the training process to complete before you can get the resource name and ID of your new model, which is required for model evaluation and model deployment.
#
# **Note**: You can close the notebook while you wait for the operation to complete.
# + id="0aa0f01805ea"
# Define the training job
training_job_display_name = f"e2e-text-training-job-{TIMESTAMP}"
# TODO
# constructs a AutoML Text Training Job
job = aiplatform.AutoMLTextTrainingJob(
display_name=training_job_display_name,
prediction_type="classification",
multi_label=False,
)
# + id="1ec60baf2c51"
model_display_name = f"e2e-text-classification-model-{TIMESTAMP}"
# TODO
# Run the training job
model = job.run(
dataset=text_dataset,
model_display_name=model_display_name,
training_fraction_split=0.7,
validation_fraction_split=0.2,
test_fraction_split=0.1,
sync=True,
)
# + [markdown] id="caaa3f32b12e"
# ## Get and review model evaluation scores
#
# After your model has finished training, you can review the evaluation scores for it.
#
# First, you need to get a reference to the new model. As with datasets, you can either use the reference to the `model` variable you created when deployed the model or you can list all of the models in your project. When listing your models, you can provide filter criteria to narrow down your search.
# + id="b0bb6be8621a"
# TODO
# list the aiplatform model
models = aiplatform.Model.list(filter=f'display_name="{model_display_name}"')
print(models)
# + [markdown] id="8481b6878ed2"
# Using the model name (in the format `projects/[PROJECT_NAME]/locations/us-central1/models/[MODEL_ID]`), you can get its model evaluations. To get model evaluations, you must use the underlying service client.
#
# Building a service client requires that you provide the name of the regionalized hostname used for your model. In this tutorial, the hostname is `us-central1-aiplatform.googleapis.com` because the model was created in the `us-central1` location.
# + id="a8443fc8861f"
# Get the ID of the model
model_name = "e2e-text-classification-model-20210824122127"
if model_name == "e2e-text-classification-model-20210824122127":
# Use the `resource_name` of the Model instance you created previously
model_name = model.resource_name
print(f"Model name: {model_name}")
# Get a reference to the Model Service client
client_options = {"api_endpoint": "us-central1-aiplatform.googleapis.com"}
model_service_client = aiplatform.gapic.ModelServiceClient(
client_options=client_options
)
# + [markdown] id="b8a788593609"
# Before you can view the model evaluation you must first list all of the evaluations for that model. Each model can have multiple evaluations, although a new model is likely to only have one.
# + id="fdcb045e29f2"
model_evaluations = model_service_client.list_model_evaluations(parent=model_name)
model_evaluation = list(model_evaluations)[0]
# + [markdown] id="cd7d3afae05c"
# Now that you have the model evaluation, you can look at your model's scores. If you have questions about what the scores mean, review the [public documentation](https://cloud.google.com/vertex-ai/docs/training/evaluating-automl-models#text).
#
# The results returned from the service are formatted as [`google.protobuf.Value`](https://googleapis.dev/python/protobuf/latest/google/protobuf/struct_pb2.html) objects. You can transform the return object as a `dict` for easier reading and parsing.
# + id="6eb9ccb0a0a0"
model_eval_dict = json_format.MessageToDict(model_evaluation._pb)
metrics = model_eval_dict["metrics"]
confidence_metrics = metrics["confidenceMetrics"]
print(f'Area under precision-recall curve (AuPRC): {metrics["auPrc"]}')
for confidence_scores in confidence_metrics:
metrics = confidence_scores.keys()
print("\n")
for metric in metrics:
print(f"\t{metric}: {confidence_scores[metric]}")
# + [markdown] id="b5dbe4dbaa60"
# ## Deploy your text classification model
#
# Once your model has completed training, you must deploy it to an _endpoint_ to get online predictions from it. When you deploy the model to an endpoint, a copy of the model is made on the endpoint with a new resource name and display name.
#
# You can deploy multiple models to the same endpoint and split traffic between the various models assigned to the endpoint. However, you must deploy one model at a time to the endpoint. To change the traffic split percentages, you must assign new values on your second (and subsequent) models each time you deploy a new model.
#
# The following code block demonstrates how to deploy a model. The code snippet relies on the Python SDK to create a new endpoint for deployment. The call to `model.deploy()` returns a reference to an `Endpoint` object--you need this reference for online predictions in the next section.
# + id="19bc4a55ccfe"
deployed_model_display_name = f"e2e-deployed-text-classification-model-{TIMESTAMP}"
# TODO
# deploy a model
endpoint = model.deploy(
deployed_model_display_name=deployed_model_display_name, sync=True
)
# + [markdown] id="531da446035b"
# In case you didn't record the name of the new endpoint, you can get a list of all your endpoints as you did before with datasets and models. For each endpoint, you can list the models deployed to that endpoint. To get a reference to the model that you just deployed, you can check the `display_name` of each model deployed to the endpoint against the model you're looking for.
# + id="f61fb44181b4"
endpoints = aiplatform.Endpoint.list()
endpoint_with_deployed_model = []
for endpoint_ in endpoints:
for model in endpoint_.list_models():
if model.display_name.find(deployed_model_display_name) == 0:
endpoint_with_deployed_model.append(endpoint_)
print(endpoint_with_deployed_model)
# + [markdown] id="351a6e8be3a5"
# ## Get online predictions from your model
#
# Now that you have your endpoint's resource name, you can get online predictions from the text classification model. To get the online prediction, you send a prediction request to your endpoint.
# + id="953b333fc0fc"
endpoint_name = "e2e-text-classification-model-20210824122127_endpoint"
if endpoint_name == "e2e-text-classification-model-20210824122127_endpoint":
endpoint_name = endpoint.resource_name
print(f"Endpoint name: {endpoint_name}")
endpoint = aiplatform.Endpoint(endpoint_name)
content = "I got a high score on my math final!"
# TODO
# send a prediction request to your endpoint
response = endpoint.predict(instances=[{"content": content}])
for prediction_ in response.predictions:
ids = prediction_["ids"]
display_names = prediction_["displayNames"]
confidence_scores = prediction_["confidences"]
for count, id in enumerate(ids):
print(f"Prediction ID: {id}")
print(f"Prediction display name: {display_names[count]}")
print(f"Prediction confidence score: {confidence_scores[count]}")
# + [markdown] id="f18811cd0477"
# ## Get batch predictions from your model
#
# You can get batch predictions from a text classification model without deploying it. You must first format all of your prediction instances (prediction input) in JSONL format and you must store the JSONL file in a Google Cloud Storage bucket. You must also provide a Google Cloud Storage bucket to hold your prediction output.
#
# To start, you must first create your predictions input file in JSONL format. Each line in the JSONL document needs to be formatted like so:
#
# ```
# { "content": "gs://sourcebucket/datasets/texts/source_text.txt", "mimeType": "text/plain"}
# ```
#
# The `content` field in the JSON structure must be a Google Cloud Storage URI to another document that contains the text input for prediction.
# [See the documentation for more information.](https://cloud.google.com/ai-platform-unified/docs/predictions/batch-predictions#text)
# + id="e4b838cbcd99"
instances = [
"We hiked through the woods and up the hill to the ice caves",
"My kitten is so cute",
]
input_file_name = "batch-prediction-input.jsonl"
# + [markdown] id="76ac422ab8dd"
# For batch prediction, you must supply the following:
#
# # + All of your prediction instances as individual files on Google Cloud Storage, as TXT files for your instances
# # + A JSONL file that lists the URIs of all your prediction instances
# # + A Google Cloud Storage bucket to hold the output from batch prediction
#
# For this tutorial, the following cells create a new Storage bucket, upload individual prediction instances as text files to the bucket, and then create the JSONL file with the URIs of your prediction instances.
# + id="1e0759fbb219"
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
BUCKET_NAME = "qwiklabs-gcp-00-09d98f4803b0"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "[your-bucket-name]":
BUCKET_NAME = f"automl-text-notebook-{TIMESTAMP}"
BUCKET_URI = f"gs://{BUCKET_NAME}"
# ! gsutil mb -l $REGION $BUCKET_URI
# + id="8b7cabbb86ad"
# Instantiate the Storage client and create the new bucket
storage = storage.Client()
bucket = storage.bucket(BUCKET_NAME)
# Iterate over the prediction instances, creating a new TXT file
# for each.
input_file_data = []
for count, instance in enumerate(instances):
instance_name = f"input_{count}.txt"
instance_file_uri = f"{BUCKET_URI}/{instance_name}"
# Add the data to store in the JSONL input file.
tmp_data = {"content": instance_file_uri, "mimeType": "text/plain"}
input_file_data.append(tmp_data)
# Create the new instance file
blob = bucket.blob(instance_name)
blob.upload_from_string(instance)
input_str = "\n".join([str(d) for d in input_file_data])
file_blob = bucket.blob(f"{input_file_name}")
file_blob.upload_from_string(input_str)
# + [markdown] id="31c262320610"
# Now that you have the bucket with the prediction instances ready, you can send a batch prediction request to Vertex AI. When you send a request to the service, you must provide the URI of your JSONL file and your output bucket, including the `gs://` protocols.
#
# With the Python SDK, you can create a batch prediction job by calling `Model.batch_predict()`.
# + id="f5ab2139d52d"
job_display_name = "e2e-text-classification-batch-prediction-job"
model = aiplatform.Model(model_name=model_name)
# TODO
# create a batch prediction job
batch_prediction_job = model.batch_predict(
job_display_name=job_display_name,
gcs_source=f"{BUCKET_URI}/{input_file_name}",
gcs_destination_prefix=f"{BUCKET_URI}/output",
sync=True,
)
batch_prediction_job_name = batch_prediction_job.resource_name
# + [markdown] id="11503f2e08a2"
# Once the batch prediction job completes, the Python SDK prints out the resource name of the batch prediction job in the format `projects/[PROJECT_ID]/locations/[LOCATION]/batchPredictionJobs/[BATCH_PREDICTION_JOB_ID]`. You can query the Vertex AI service for the status of the batch prediction job using its ID.
#
# The following code snippet demonstrates how to create an instance of the `BatchPredictionJob` class to review its status. Note that you need the full resource name printed out from the Python SDK for this snippet.
#
# + id="bf6e614723ed"
from google.cloud.aiplatform import jobs
batch_job = jobs.BatchPredictionJob(batch_prediction_job_name)
print(f"Batch prediction job state: {str(batch_job.state)}")
# + [markdown] id="1f9a12dadf6f"
# After the batch job has completed, you can view the results of the job in your output Storage bucket. You might want to first list all of the files in your output bucket to find the URI of the output file.
# + id="8ff1ec03205c"
BUCKET_OUTPUT = f"{BUCKET_URI}/output"
# ! gsutil ls -a $BUCKET_OUTPUT
# + [markdown] id="52f3f8af2e41"
# The output from the batch prediction job should be contained in a folder (or _prefix_) that includes the name of the batch prediction job plus a time stamp for when it was created.
#
# For example, if your batch prediction job name is `my-job` and your bucket name is `my-bucket`, the URI of the folder containing your output might look like the following:
#
# ```
# gs://my-bucket/output/prediction-my-job-2021-06-04T19:54:25.889262Z/
# ```
#
# To read the batch prediction results, you must download the file locally and open the file. The next cell copies all of the files in the `BUCKET_OUTPUT_FOLDER` into a local folder.
# + id="4bb16e040942"
RESULTS_DIRECTORY = "prediction_results"
RESULTS_DIRECTORY_FULL = f"{RESULTS_DIRECTORY}/output"
# Create missing directories
os.makedirs(RESULTS_DIRECTORY, exist_ok=True)
# Get the Cloud Storage paths for each result
# ! gsutil -m cp -r $BUCKET_OUTPUT $RESULTS_DIRECTORY
# Get most recently modified directory
latest_directory = max(
[
os.path.join(RESULTS_DIRECTORY_FULL, d)
for d in os.listdir(RESULTS_DIRECTORY_FULL)
],
key=os.path.getmtime,
)
print(f"Local results folder: {latest_directory}")
# + [markdown] id="f406e1e4d5ec"
# With all of the results files downloaded locally, you can open them and read the results. In this tutorial, you use the [`jsonlines`](https://jsonlines.readthedocs.io/en/latest/) library to read the output results.
#
# The following cell opens up the JSONL output file and then prints the predictions for each instance.
# + id="91d7f2a74a7c"
# Get downloaded results in directory
results_files = []
for dirpath, subdirs, files in os.walk(latest_directory):
for file in files:
if file.find("predictions") >= 0:
results_files.append(os.path.join(dirpath, file))
# Consolidate all the results into a list
results = []
for results_file in results_files:
# Open each result
with jsonlines.open(results_file) as reader:
for result in reader.iter(type=dict, skip_invalid=True):
instance = result["instance"]
prediction = result["prediction"]
print(f"\ninstance: {instance['content']}")
for key, output in prediction.items():
print(f"\n{key}: {output}")
# + [markdown] id="af3874f08502"
# ## Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# * Dataset
# * Training job
# * Model
# * Endpoint
# * Batch prediction
# * Batch prediction bucket
# + id="adce73b48b72"
if os.getenv("IS_TESTING"):
# ! gsutil rm -r $BUCKET_URI
batch_job.delete()
# `force` parameter ensures that models are undeployed before deletion
endpoint.delete(force=True)
model.delete()
text_dataset.delete()
# Training job
job.delete()
# + [markdown] id="fa6a8c434c79"
# ## Next Steps
#
# After completing this tutorial, see the following documentation pages to learn more about Vertex AI:
#
# * [Preparing text training data](https://cloud.google.com/vertex-ai/docs/datasets/prepare-text)
# * [Training an AutoML model using the API](https://cloud.google.com/vertex-ai/docs/training/automl-api#text)
# * [Evaluating AutoML models](https://cloud.google.com/vertex-ai/docs/training/evaluating-automl-models#text)
# * [Deploying a model using the Vertex AI API](https://cloud.google.com/vertex-ai/docs/predictions/deploy-model-api#aiplatform_create_endpoint_sample-python)
# * [Getting online predictions from AutoML models](https://cloud.google.com/vertex-ai/docs/predictions/deploy-model-api#aiplatform_create_endpoint_sample-python)
# * [Getting batch predictions](https://cloud.google.com/vertex-ai/docs/predictions/batch-predictions#text)
| courses/machine_learning/deepdive2/launching_into_ml/solutions/automl_text_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Домашнее задание
# ## Свойства матричных вычислений
import numpy as np
l = [[12, 5, 7, 8], [20, 3, 4, 2], [25, 3, 16, 1], [8, 6, 4, 10]]
A = np.array(l)
A
k = [[2, 8, 9, 4], [12, 24, 23, 42], [34, 15, 17, 4], [0, 1, 5, 32]]
B = np.array(k)
B
k = [[8, 4 , 7, 1], [4, 2, 1, 13], [7, 1, 5, 0], [7, 8, 0, 0]]
C = np.array(k)
C
# Транспонирование матрицы:
A.transpose()
A.T
# Дважды транспонированная матрица равна исходной матрице:
(A.T).T
# Транспонирование суммы матриц равно сумме транспонированных матриц:
(A+B).T
A.T+B.T
# Транспонирование произведения матриц равно произведению транспонированных
# матриц расставленных в обратном порядке:
(A.dot(B)).T
(B.T).dot(A.T)
# Транспонирование произведения матрицы на число равно произведению этого
# числа на транспонированную матрицу:
t = 3
(t * A).T
t * (A.T)
# Определители исходной и транспонированной матрицы совпадают:
format(np.linalg.det(A), '.9g')
format(np.linalg.det(A.T), '.9g')
# Умножение матрицы на число
#
2 * A
# Произведение единицы и любой заданной матрицы равно заданной матрице:
1 * A
# Произведение нуля и любой матрицы равно нулевой матрице, размерность которой
# равна исходной матрицы:
0 * A
# Произведение матрицы на сумму чисел равно сумме произведений матрицы на
# каждое из этих чисел:
(2 + 3) * A
2 * A + 3 * A
# Произведение матрицы на произведение двух чисел равно произведению второго
# числа и заданной матрицы, умноженному на первое число:
(2 * 3) * A
2 * (3 * A)
# Произведение суммы матриц на число равно сумме произведений этих матриц на
# заданное число:
2 * (A + B)
2 * A + 2 * B
# Сложение матриц
#
A + B
B + A
A + (B + C)
(A + B) + C
A + (-1)*A
# Диаграмма матричного умножения
#
A.dot(B)
# Ассоциативность умножения. Результат умножения матриц не зависит от порядка, в
# котором будет выполняться эта операция:
A.dot(B.dot(C))
(A.dot(B)).dot(C)
# Дистрибутивность умножения. Произведение матрицы на сумму матриц равно
# сумме произведений матриц:
A.dot(B+C)
A.dot(B)+A.dot(C)
# Умножение матриц в общем виде не коммутативно. Это означает, что для матриц не
# выполняется правило независимости произведения от перестановки множителей:
A.dot(B)
B.dot(A)
# Произведение заданной матрицы на единичную равно исходной матрице:
E = np.matrix('1 0 0 0; 0 1 0 0; 0 0 1 0; 0 0 0 1')
E.dot(A)
A.dot(E)
# Произведение заданной матрицы на нулевую матрицу равно нулевой матрице:
Z = np.matrix('0 0 0 0; 0 0 0 0; 0 0 0 0; 0 0 0 0')
Z.dot(A)
A.dot(Z)
# Определитель матрицы
np.linalg.det(A)
# Определитель матрицы остается неизменным при ее транспонировании:
round(np.linalg.det(A), 3)
round(np.linalg.det(A.T), 3)
# Если у матрицы есть строка или столбец, состоящие из нулей, то определитель
# такой матрицы равен нулю:
N = np.matrix('3 2 3; 0 0 0; 12 4 5')
np.linalg.det(N)
# При перестановке строк матрицы знак ее определителя меняется на
# противоположный:
N = np.matrix('3 2 3; 5 6 9; 12 4 5')
round(np.linalg.det(N), 3)
N = np.matrix('5 6 9; 3 2 3; 12 4 5')
round(np.linalg.det(N), 3)
# Если у матрицы есть две одинаковые строки, то ее определитель равен нулю:
N = np.matrix('3 2 3; 3 2 3; 12 4 5')
np.linalg.det(N)
# Если все элементы строки или столбца матрицы умножить на какое-то число, то и
# определитель будет умножен на это число:
N = np.matrix('3 5 3; 3 2 3; 12 4 5')
k = 2
M = N.copy()
M[2, :] = k * M[2, :]
print(M)
det_N = round(np.linalg.det(N), 3)
det_M = round(np.linalg.det(M), 3)
det_N * k
det_M
# Если все элементы строки или столбца можно представить как сумму двух
# слагаемых, то определитель такой матрицы равен сумме определителей двух соответствующих
# матриц:
C = A.copy()
C[1, :] += B[1, :]
print(C)
A
B
round(np.linalg.det(C), 3)
round(np.linalg.det(A), 3) + round(np.linalg.det(B), 3)
# Если к элементам одной строки прибавить элементы другой строки, умноженные на
# одно и тоже число, то определитель матрицы не изменится:
C = A.copy()
C[1, :] = C[1, :] + 2 * C[0, :]
A
C
round(np.linalg.det(A), 3)
round(np.linalg.det(C), 3)
# Если строка или столбец матрицы является линейной комбинацией других строк
# (столбцов), то определитель такой матрицы равен нулю:
A[1, :] = A[0, :] + 2 * A[2, :]
round(np.linalg.det(A), 3)
# Если матрица содержит пропорциональные строки, то ее определитель равен нулю:
A[1, :] = 2 * A[0, :]
A
round(np.linalg.det(A), 3)
# Обратная матрица
# Обратная матрица обратной матрицы есть исходная матрица:
N = np.matrix('6 2; -7 12')
np.linalg.inv(N)
# Обратная матрица транспонированной матрицы равна транспонированной матрице
# от обратной матрицы:
np.linalg.inv(N.T)
(np.linalg.inv(N)).T
# Обратная матрица произведения матриц равна произведению обратных матриц:
M = np.matrix('7. 2.; 6. 12.')
N = np.matrix('-1. 9.; 7. -8.')
np.linalg.inv(M.dot(N))
np.linalg.inv(N).dot(np.linalg.inv(M))
# ## Матричный метод
# Рассмотрим систему линейных уравнений:
# * 2x+3y+7z+6t=1
# * 3x+5y+3z+t=3
# * 5x+3y+z+3t=4
# * 3x+3y+z+6t=5
# Создадим матрицу
K = np.array([[2., 3., 7., 6.], [3., 5., 3., 1.], [5., 3., 1., 3.], [3., 3., 1., 6.]])
K
# Создадим вектор (правую часть системы)
L = np.array([[1.], [3.], [4.], [5.]])
L
# Для решения системы воспользуемся функцией numpy.linalg.solve
np.linalg.solve(K, L)
# ## Метод Крамера
# Найти решение СЛАУ при помощи метода Крамера.
# * 5x-3y=14
# * 2x+y=10
#
# Вычисляем определитель матрицы системы:
K = np.array([[5., 3.], [2., 1.]])
L = np.array([[14.], [10.]])
D = round(np.linalg.det(K), 3)
D
# Так как Δ≠0 , то по теореме Крамера система совместна и имеет единственное решение. Вычислим вспомогательные определители. Определитель Δ1 получим из определителя Δ заменой его первого столбца столбцом свободных коэффициентов. Будем иметь:
J = np.array([[14., 3], [10., 1.]])
D_1 = round(np.linalg.det(J), 3)
D_1
# Аналогично, определитель Δ2 получается из определителя матрицы системы Δ заменой второго столбца столбцом свободных коэффициентов:
J = np.array([[5., 14.], [2., 10]])
D_2 = round(np.linalg.det(J), 3)
D_2
# Тогда получаем, что
x_1 = D_1/D
print(x_1)
x_2 = D_2/D
print(x_2)
| Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Configure for local/external-to-Terra development
# + pycharm={"name": "#%%\n"}
import os
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Set Execution Directory
# + pycharm={"name": "#%%\n"}
# NOTEBOOK_EXECUTION_DIRECTORY="/Users/mbaumann/Repos/mbaumann-broad/"
# # %cd {NOTEBOOK_EXECUTION_DIRECTORY}
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Set Environment Variables
# + pycharm={"name": "#%%\n"}
# The processing performed by this Notebook requires grep with PCRE
# support, which the version of grep that comes with MacOS Monterey
# does not provide. Therefore, install it with homebrew and add to PATH:
# $ brew install grep
os.environ['PATH'] = f"/usr/local/opt/grep/libexec/gnubin:{os.environ['PATH']}"
# The processing performed by this Notebook requires sed "-z" option
# support, which the version of sed that comes with MacOS Monterey
# does not provide. Therefore, install it with homebrew and add to PATH:
# $ brew install gnu-sed
os.environ['PATH'] = f"/usr/local/opt/gnu-sed/libexec/gnubin:{os.environ['PATH']}"
# The processing performed by this Notebook requires xargs "-a" option
# support, which the version of sed that comes with MacOS Monterey
# does not provide. Therefore, install it with homebrew and add to PATH:
# $ brew install findutils
os.environ['PATH'] = f"/usr/local/opt/findutils/libexec/gnubin:{os.environ['PATH']}"
if os.environ.get('WORKSPACE_BUCKET') is None:
# Workspace bucket used by: `DRS and Signed URL Development - Dev`
# WORKSPACE_BUCKET="gs://fc-b14e50ee-ccbe-4ee9-9aa4-f4e4ff85bc03"
# Workspace bucket used by: `DRS and Signed URL Development - Alpha`
WORKSPACE_BUCKET = "gs://fc-26863db0-1fe6-463b-a05b-9f8c8cb33dac"
os.environ['WORKSPACE_BUCKET'] = WORKSPACE_BUCKET
if os.environ.get('WORKSPACE_NAMESPACE') is None:
WORKSPACE_NAMESPACE = "drs-billing-project"
os.environ['WORKSPACE_NAMESPACE'] = WORKSPACE_NAMESPACE
if os.environ.get('WORKSPACE_NAME') is None:
WORKSPACE_NAME = "DRS Data Access Scale Testing - Alpha"
os.environ['WORKSPACE_NAME'] = WORKSPACE_NAME
| terra_workflow_scale_test_tools/external_development.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import Counter
import os
import re
import sys
import time
from cltk.corpus.utils.formatter import assemble_phi5_works_filepaths
from cltk.corpus.utils.formatter import phi5_plaintext_cleanup
from cltk.tokenize.sentence import TokenizeSentence
from cltk.tag.pos import POSTag
from nltk.tokenize.punkt import PunktLanguageVars
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
import pandas as pd
# -
def works_texts_list(rm_punctuation, rm_periods):
fps = assemble_phi5_works_filepaths()
curly_comp = re.compile(r'{.+?}')
_list = []
for fp in fps:
with open(fp) as fo:
fr = fo.read()
text = phi5_plaintext_cleanup(fr, rm_punctuation, rm_periods)
text = curly_comp.sub('', text)
_list.append(text)
return _list
t0 = time.time()
text_list = works_texts_list(rm_punctuation=True, rm_periods=True)
print('Total texts', len(text_list))
print('Time to build list of texts: {}'.format(time.time() - t0))
# # Bag of words, indivudual word count
# bag of words/word count
def bow_csv():
t0 = time.time()
vectorizer = CountVectorizer(min_df=1)
column_names = ['wc_' + w for w in vectorizer.get_feature_names()]
term_document_matrix = vectorizer.fit_transform(text_list)
dataframe_bow = pd.DataFrame(term_document_matrix.toarray(), columns=column_names)
print('DF BOW shape', dataframe_bow.shape)
fp = os.path.expanduser('~/cltk_data/user_data/bow_latin.csv')
dataframe_bow.to_csv(fp)
print('Time to create BOW vectorizer and write csv: {}'.format(time.time() - t0))
# +
#bow_csv()
# -
# # tf-idf
# tf-idf
def tfidf_csv():
t0 = time.time()
vectorizer = TfidfVectorizer(min_df=1)
column_names = ['tfidf_' + w for w in vectorizer.get_feature_names()]
term_document_matrix = vectorizer.fit_transform(text_list)
dataframe_tfidf = pd.DataFrame(term_document_matrix.toarray(), columns=column_names)
print('DF tf-idf shape', dataframe_tfidf.shape)
fp = os.path.expanduser('~/cltk_data/user_data/tfidf_latin.csv')
dataframe_tfidf.to_csv(fp)
print('Time to create tf-idf vectorizer and write csv: {}'.format(time.time() - t0))
# +
#tfidf_csv()
# -
# # Character, simple word, and sentence counts
# +
# char count
# word count
# sentence
# word count lens
# +
def char_len():
"""Count char len in an input string (doc)."""
t0 = time.time()
char_len = {}
for i, doc in enumerate(text_list):
char_len[i] = pd.Series(len(doc), index=['char_len'])
df_char_len = pd.DataFrame(char_len).transpose()
fp = os.path.expanduser('~/cltk_data/user_data/char_len_latin.csv')
df_char_len.to_csv(fp)
print('Time to create doc len counts: {}'.format(time.time() - t0))
char_len()
# +
def word_count():
"""Count words in an input string (doc)."""
t0 = time.time()
p = PunktLanguageVars()
word_count = {}
for i, doc in enumerate(text_list):
wc_doc = len(p.word_tokenize(doc))
word_count[i] = pd.Series(wc_doc, index=['word_count'])
df_word_count = pd.DataFrame(word_count).transpose()
fp = os.path.expanduser('~/cltk_data/user_data/word_count_lens_latin.csv')
df_word_count.to_csv(fp)
print('Time to create doc word count: {}'.format(time.time() - t0))
word_count()
# -
text_list_no_cleanup = works_texts_list(rm_punctuation=False, rm_periods=False)
text_list_no_cleanup[1][:500]
# see how sent tokenizer works
s = ' ex scriptis eorum qui ueri arbitrantur . . . neque ipsi eos alii modi esse atque Amilcar dixit, ostendere possunt aliter. antequam Barcha perierat, alii rei causa in Africam missus . . . . . . tantum bellum suscitare conari aduersarios contra bellosum genus. qui cum is ita foedus icistis . . . . . . cum iure sine periculo bellum geri poteratur. qui intellegunt quae fiant, dissentiuntur. Legati quo missi sunt ueniunt, dedicant mandata. Saguntinorum Sempronius Lilybaeo celocem in Africam mittit u'
tokenizer = TokenizeSentence('latin')
sent_tokens = tokenizer.tokenize_sentences(s)
sent_tokens = [s for s in sent_tokens if len(s) > 1] # rm '.' sents
sent_tokens
# +
def sentence_count():
"""Count sentence in an input string (doc)."""
t0 = time.time()
tokenizer = TokenizeSentence('latin')
word_count = {}
for i, doc in enumerate(text_list_no_cleanup):
sent_tokens = tokenizer.tokenize_sentences(doc)
wc_doc = [s for s in sent_tokens if len(s) > 1]
word_count[i] = pd.Series(, index=['sentence_count'])
df_word_count = pd.DataFrame(word_count).transpose()
fp = os.path.expanduser('~/cltk_data/user_data/sentence_count_lens_latin.csv')
df_word_count.to_csv(fp)
print('Time to create doc word count: {}'.format(time.time() - t0))
sentence_count()
# +
def word_len_counts():
"""Count words lengths in an input string (doc)."""
t0 = time.time()
p = PunktLanguageVars()
word_counts = {}
for i, doc in enumerate(text_list_no_cleanup):
word_tokens = p.word_tokenize(doc)
list_of_counts = ['word_len_' + str(len(w)) for w in word_tokens]
counter_word_counts = Counter(list_of_counts)
word_counts[i] = pd.Series(counter_word_counts, index=counter_word_counts.keys())
df_word_count = pd.DataFrame(word_counts).transpose()
fp = os.path.expanduser('~/cltk_data/user_data/word_count_lens_latin.csv')
df_word_count.to_csv(fp)
print('Time to create doc word count: {}'.format(time.time() - t0))
word_len_counts()
# +
def sentence_word_count():
"""Count words lengths in an input string (doc)."""
t0 = time.time()
tokenizer_sent = TokenizeSentence('latin')
p = PunktLanguageVars()
word_counts = {}
for i, doc in enumerate(text_list_no_cleanup):
list_words_per_sentence = []
sent_tokens = tokenizer_sent.tokenize_sentences(doc)
sent_tokens = [s for s in sent_tokens if len(s) > 1]
for sent in sent_tokens:
word_tokens = p.word_tokenize(sent)
words_in_sent = len(word_tokens)
list_words_per_sentence.append(words_in_sent)
list_of_counts = ['words_in_sent_' + str(count) for count in list_words_per_sentence]
counter_word_counts_per_sents = Counter(list_of_counts)
word_counts[i] = pd.Series(counter_word_counts_per_sents,
index=counter_word_counts_per_sents.keys())
df_word_count_per_sent = pd.DataFrame(word_counts).transpose()
fp = os.path.expanduser('~/cltk_data/user_data/words_per_sent_latin.csv')
df_word_count_per_sent.to_csv(fp)
print('Time to create count of words per sentence: {}'.format(time.time() - t0))
sentence_word_count()
# +
def pos_counts(index_start=0, index_break=99):
"""Count part of speech input string (doc)."""
t0 = time.time()
tokenizer_sent = TokenizeSentence('latin')
pos_counts = {}
tagger = POSTag('latin')
for i, doc in enumerate(text_list_no_cleanup):
i += index_start
#if i % 1 == 0:
print('Processing doc #{}'.format(i))
pos_tags_list = []
sent_tokens = tokenizer_sent.tokenize_sentences(doc)
sent_tokens = [s for s in sent_tokens if len(s) > 1]
for sent in sent_tokens:
pos_tags = tagger.tag_tnt(sent.lower())
pos_tags = [t[1] for t in pos_tags]
pos_tags_list += pos_tags
pos_counts_counter = Counter(pos_tags_list)
pos_counts[i] = pd.Series(pos_counts_counter, index=pos_counts_counter.keys())
if i == index_break:
print('breaking …')
break
df_pos_counts = pd.DataFrame(pos_counts).transpose()
fp = os.path.expanduser('~/cltk_data/user_data/pos_counts_latin_{}.csv'.format(index_start))
df_pos_counts.to_csv(fp)
print('Time to create count of words per sentence: {}'.format(time.time() - t0))
pos_counts(index_start=0, index_break=99)
#pos_counts(index_start=100)
#pos_counts(index_start=200)
#pos_counts(index_start=300)
#pos_counts(index_start=400)
#pos_counts(index_start=500)
#pos_counts(index_start=600)
#pos_counts(index_start=700)
#pos_counts(index_start=800)
# -
| tf-idf/Latin feature tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Multiple Linear Regression
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# -
# Importing the dataset
dataset = pd.read_csv('50_Startups.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, 4].values
# Encoding categorical data
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
labelencoder = LabelEncoder()
X[:, 3] = labelencoder.fit_transform(X[:, 3])
onehotencoder = OneHotEncoder(categorical_features = [3])
X = onehotencoder.fit_transform(X).toarray()
# Avoiding the Dummy Variable Trap
X = X[:, 1:]
# Splitting the dataset into the Training set and Test set
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# Feature Scaling
"""from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
X_train = sc_X.fit_transform(X_train)
X_test = sc_X.transform(X_test)
sc_y = StandardScaler()
y_train = sc_y.fit_transform(y_train)"""
# Fitting Multiple Linear Regression to the Training set
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
# Predicting the Test set results
y_pred = regressor.predict(X_test)
print(y_pred)
print(y_test)
| Topics/4. Multiple Linear Regression/.ipynb_checkpoints/Multiple Linear Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Stack():
def __init__(self,stones):
self.stones = stones
def __str__(self):
return "There are {} stones left".format(len(self.stones)) # only need 'self.stones' (only had self.Stack.stones)
def still_has_stones(self):
return len(self.stones) != 0
#remove stone should be here, rather than in Player class
def remove_stone(self, drawn_stones): # missed 'drawn_stones' in the arguments declaration. if it was in player class, do i still need it?
for i in range(drawn_stones+1):
self.stones.pop() # i dont need Slack.stones since i am in Stack already ?
class Player():
def __init__(self,name):
self.name = name
def choice(self):
drawn_stones = int(input("{} Would you like to remove 1 or 2 stones?".format(self.name)))
#assert drawn_stones < 3, "invalid input, chose between 1 and 2"
return drawn_stones
# +
import random
Player1 = Player("Player1")
Player2 = Player("Player2")
list_of_stones = list(range(1,21)) # i did this - whic is ok - list_of_stones = Stack(list(range(1,21)))
stack = Stack(list_of_stones) # ok - missed that one - makes sense
while stack.still_has_stones():
print('-'*100)
Player1_Choice = Player1.choice()
if len(stack.stones) > Player1_Choice:
stack.stones.append(stack.remove_stone(Player1_Choice)) # missed that one - this one is crucial
# I did " Stack.append(remove_stone()) "
# ok we use choice = drawn stones
print(stack) # i would have thought print(Stack()) - ok then
else:
break
Player2_Choice = Player2.choice()
if len(stack.stones) > Player2_Choice:
stack.stones.append(stack.remove_stone(Player2_Choice))
print(stack)
else:
break
print("Game Over")
# -
| Exercises - Qasim/Python. Pandas, Viz/Nim Game - Resolved!.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jrhumberto/cd/blob/main/009_NLP_textblob.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-oin9e-pA9yt"
# # Glossário
# <ul>
# <li><a href="#introduction">Introduction</a></li>
# </ul>
# + [markdown] id="PuiJFQcWBaiB"
# # Introduction <a id='introduction'></a>
# + [markdown] id="WuNMpnSCRNNf"
# Fonte:
# - https://medium.com/@viniljf/criando-um-classificador-para-processamento-de-linguagem-natural-8dc27f3642a1
# - https://medium.com/@viniljf/criando-um-analisador-de-sentimentos-para-tweets-a53bae0c5147
# + colab={"base_uri": "https://localhost:8080/"} id="V4g3xFWwR8ra" outputId="e74560e6-8a25-4ce6-cfdc-dcb46f418928"
# !pip install textblob
# !python -m textblob.download_corpora
# + colab={"base_uri": "https://localhost:8080/"} id="EWOENcCkRJVD" outputId="55bdf30a-b91c-4ada-dbf4-2b1c543880f8"
from textblob.classifiers import NaiveBayesClassifier
from textblob import TextBlob
train_set = [
('Eu adoro comer hamburguer', 'positivo'),
('Este lugar é horrível', 'negativo'),
('Você é uma pessoa adorável', 'positivo'),
('Você é uma pessoa horrível', 'negativo'),
('A festa está ótima', 'positivo'),
('A festa está péssima', 'negativo'),
('Este lugar é maravilhoso', 'positivo'),
('Estou cansado disso.', 'negativo'),
('Eu te odeio', 'negativo'),
('Eu te adoro', 'positivo'),
('Eu te amo', 'positivo'),
('Você é incrível','positivo'),
('Eu estou com muita raiva','negativo'),
('Eu odeio essa linguagem','negativo'),
('Essa linguagem é fantátisca','positivo'),
('Essa linguagem é muito boa','positivo'),
('Que comida gostosa','positivo'),
('Que comida horrível','negativo'),
('Estou me sentindo ótimo','positivo'),
('Hoje eu estou péssimo','negativo')
]
test_set = [
('Ótima linguagem', 'positivo'),
('Péssima essa linguagem', 'negativo'),
('Você é horrível', 'negativo'),
('Comida gostosa!', 'positivo'),
('Que raiva!', 'negativo'),
('Ótima festa!', 'positivo'),
('Eu não odeio todo mundo','positivo')
]
cl = NaiveBayesClassifier(train_set)
accuracy = cl.accuracy(test_set)
frase = 'Eu não odeio todo mundo'
blob = TextBlob(frase,classifier=cl)
print('Esta frase é de caráter:{}'.format(blob.classify()))
print('Precisão da previsão:{}'.format(accuracy))
# + id="nlAT1YuhR6gG"
import tweepy
from textblob import TextBlob
consumer_key = 'yAL0H3XVN3TFmZtSXxONNAuLJ'
consumer_secret = '<KEY>'
access_token = '<KEY>'
access_token_secret = '<KEY>'
auth = tweepy.OAuthHandler(consumer_key,consumer_secret)
auth.set_access_token(access_token,access_token_secret)
api = tweepy.API(auth)
#print(api.me())
tweets = api.search('<NAME>')
for tweet in tweets:
frase = TextBlob(tweet.text)
if frase.detect_language() != 'en':
traducao = TextBlob(str(frase.translate(to='en')))
print('Tweet: {0} - Sentimento: {1}'.format(tweet.text, traducao.sentiment))
else:
print('Tweet: {0} - Sentimento: {1}'.format(tweet.text, frase.sentiment))
| 009_NLP_textblob.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MadMiner particle physics tutorial
#
# # Part 4a: Limit setting
#
# <NAME>, <NAME>, <NAME>, and <NAME> 2018-2019
# In part 4a of this tutorial we will use the networks trained in step 3a and 3b to calculate the expected limits on our theory parameters.
# ## 0. Preparations
# +
from __future__ import absolute_import, division, print_function, unicode_literals
import six
import logging
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
# %matplotlib inline
from madminer.limits import AsymptoticLimits
from madminer.sampling import SampleAugmenter
from madminer import sampling
from madminer.plotting import plot_histograms
# +
# MadMiner output
logging.basicConfig(
format='%(asctime)-5.5s %(name)-20.20s %(levelname)-7.7s %(message)s',
datefmt='%H:%M',
level=logging.INFO
)
# Output of all other modules (e.g. matplotlib)
for key in logging.Logger.manager.loggerDict:
if "madminer" not in key:
# print("Deactivating logging output for", key)
logging.getLogger(key).setLevel(logging.WARNING)
# -
# ## 1. Preparations
# In the end, what we care about are not plots of the log likelihood ratio, but limits on parameters. But at least under some asymptotic assumptions, these are directly related. MadMiner makes it easy to calculate p-values in the asymptotic limit with the `AsymptoticLimits` class in the `madminer.limits`:
# limits = AsymptoticLimits('data/lhe_data_shuffled.h5')
limits = AsymptoticLimits('/data_CMS/cms/cortinovis/smeftsim/data_sme_wb_100k/delphes_data_shuffled.h5')
# This class provids two high-level functions:
# - `AsymptoticLimits.observed_limits()` lets us calculate p-values on a parameter grid for some observed events, and
# - `AsymptoticLimits.expected_limits()` lets us calculate expected p-values on a parameter grid based on all data in the MadMiner file.
#
# First we have to define the parameter grid on which we evaluate the p-values.
grid_ranges = [(-1, 1.), (-1, 1.)]
#grid_ranges = [(-10, 10.), (-10, 10.)]
grid_resolutions = [25, 25]
# What luminosity (in inverse pb) are we talking about?
lumi = 3000.*1000.
# ## 2. Expected limits based on histogram
# First, as a baseline, let us calculate the expected limits based on a simple jet pT histogram. Right now, there are not a lot of option for this function; MadMiner even calculates the binning automatically. (We will add more functionality!)
#
# The keyword `include_xsec` determines whether we include information from the total rate or just use the shapes. Since we don't model backgrounds and systematics in this tutorial, the rate information is unrealistically large, so we leave it out here.
## create dictionaries to store results
p_values = {}
mle = {}
try:
theta_grid, p_values_expected_histo, best_fit_expected_histo, _, _, (histos, observed, observed_weights) = limits.expected_limits(
mode="histo",
hist_vars=["pt_j1"],
theta_true=[0.,0.],
grid_ranges=grid_ranges,
grid_resolutions=grid_resolutions,
luminosity=lumi,
include_xsec=False,
return_asimov=True,
)
p_values["Histogram"] = p_values_expected_histo
mle["Histogram"] = best_fit_expected_histo
except:
print("no Histogram")
# With `mode="rate"`, we could calculate limits based on only the rate -- but again, since the rate is extremely powerful when backgrounds and systematics are not taken into account, we don't do that in this tutorial.
# Let's visualize the likelihood estimated with these histograms:
# +
indices = [12 + i * 25 for i in [6,9,12,15,18]]
fig = plot_histograms(
histos=[histos[i] for i in indices],
observed=[observed[i] for i in indices],
observed_weights=observed_weights,
histo_labels=[r"$\theta_0 = {:.2f}$".format(theta_grid[i,0]) for i in indices],
xlabel="Jet $p_T$",
xrange=(0.,500.),
yrange=(0.,0.0125),
)
plt.show()
# -
# ## 3. Expected limits based on ratio estimators
# Next, `mode="ml"` allows us to calculate limits based on any `ParamterizedRatioEstimator` instance like the ALICES estimator trained above:
try:
theta_grid, p_values_expected_alices, best_fit_expected_alices, _, _, _ = limits.expected_limits(
mode="ml",
model_file='/data_CMS/cms/cortinovis/smeftsim/models_sme_wb_100k/alices',
theta_true=[0.,0.],
grid_ranges=grid_ranges,
grid_resolutions=grid_resolutions,
luminosity=lumi,
include_xsec=False,
)
p_values["ALICES"] = p_values_expected_alices
mle["ALICES"] = best_fit_expected_alices
except:
print("no ALICES")
# and the ALICES estimator which only used the observable `pt_j1`
'''try:
theta_grid, p_values_expected_alices_pt, best_fit_expected_alices_pt, _, _, _ = limits.expected_limits(
mode="ml",
model_file='dim6/models_tev/alices_pt',
theta_true=[0.,0.],
grid_ranges=grid_ranges,
grid_resolutions=grid_resolutions,
luminosity=lumi,
include_xsec=False,
)
p_values["ALICES_pt"] = p_values_expected_alices_pt
mle["ALICES_pt"] = best_fit_expected_alices_pt
except:
print("no ALICES_pt")'''
# ## 4. Expected limits based on score estimators
# To get p-values from a SALLY estimator, we have to use histograms of the estimated score:
try:
theta_grid, p_values_expected_sally, best_fit_expected_sally, _, _, (histos, observed, observed_weights) = limits.expected_limits(
mode="sally",
model_file='/data_CMS/cms/cortinovis/smeftsim/models_sme_wb_100k/sally',
theta_true=[0.,0.],
grid_ranges=grid_ranges,
grid_resolutions=grid_resolutions,
luminosity=lumi,
include_xsec=False,
return_asimov=True,
)
p_values["SALLY"] = p_values_expected_sally
mle["SALLY"] = best_fit_expected_sally
except:
print("no SALLY")
# Let's have a look at the underlying 2D histograms:
# +
indices = [12 + i * 25 for i in [0,6,12,18,24]]
fig = plot_histograms(
histos=[histos[i] for i in indices],
observed=observed[0,:100,:],
observed_weights=observed_weights[:100],
histo_labels=[r"$\theta_0 = {:.2f}$".format(theta_grid[i,0]) for i in indices],
xlabel=r'$\hat{t}_0(x)$',
ylabel=r'$\hat{t}_1(x)$',
xrange=(-3.,.5),
yrange=(-3.,3.),
log=True,
zrange=(1.e-3,1.),
markersize=10.
)
# -
# ## 5. Expected limits based on likelihood estimators
'''try:
theta_grid, p_values_expected_scandal, best_fit_expected_scandal, _, _, _ = limits.expected_limits(
mode="ml",
model_file='ewdim6/models_ew/scandal',
theta_true=[0.,0.],
grid_ranges=grid_ranges,
grid_resolutions=grid_resolutions,
luminosity=lumi,
include_xsec=False,
)
p_values["SCANDAL"] = p_values_expected_scandal
mle["SCANDAL"] = best_fit_expected_scandal
except:
print("no SCANDAL")'''
# ## 6. Toy signal
# In addition to these expected limits (based on the SM), let us inject a mock signal. We first generate the data:
sampler = SampleAugmenter('/data_CMS/cms/cortinovis/smeftsim/data_sme_wb_100k/delphes_data_shuffled.h5')
x_observed, _, _ = sampler.sample_test(
theta=sampling.morphing_point([0.5,0.5]),
n_samples=15,
folder=None,
filename=None,
)
# +
_, p_values_observed, best_fit_observed, _, _, _ = limits.observed_limits(
x_observed=x_observed,
mode="ml",
model_file='/data_CMS/cms/cortinovis/smeftsim/models_sme_wb_100k/alices',
grid_ranges=grid_ranges,
grid_resolutions=grid_resolutions,
luminosity=lumi,
include_xsec=False,
)
p_values["ALICES (observed)"] = p_values_observed
mle["ALICES (observed)"] = best_fit_observed
# -
# ## 7. Plot
# Let's plot the results:
# +
show = "SALLY"
bin_size = (grid_ranges[0][1] - grid_ranges[0][0])/(grid_resolutions[0] - 1)
edges = np.linspace(grid_ranges[0][0] - bin_size/2, grid_ranges[0][1] + bin_size/2, grid_resolutions[0] + 1)
centers = np.linspace(grid_ranges[0][0], grid_ranges[0][1], grid_resolutions[0])
fig = plt.figure(figsize=(6,5))
ax = plt.gca()
cmin, cmax = 1.e-2, 1.
pcm = ax.pcolormesh(
edges, edges, p_values[show].reshape((grid_resolutions[0], grid_resolutions[1])).T,
norm=matplotlib.colors.LogNorm(vmin=cmin, vmax=cmax),
cmap='Greys_r'
)
cbar = fig.colorbar(pcm, ax=ax, extend='both')
for i, (label, p_value) in enumerate(six.iteritems(p_values)):
plt.contour(
centers, centers, p_value.reshape((grid_resolutions[0], grid_resolutions[1])).T,
levels=[0.32],
linestyles='-', colors='C{}'.format(i)
)
plt.scatter(
theta_grid[mle[label]][0], theta_grid[mle[label]][1],
s=80., color='C{}'.format(i), marker='*',
label=label
)
plt.legend()
plt.xlabel(r'$\theta_0$')
plt.ylabel(r'$\theta_1$')
cbar.set_label('Expected p-value ({})'.format(show))
plt.tight_layout()
plt.show()
print('luminosity: ', lumi/1000, ' ifb')
# -
# ### Save results
# Let's save the results to later compare them to results obtained in other notebooks.
np.save("ewdim6/models_ew_02_100k/limits.npy", [p_values,mle])
| examples/tutorial_particle_physics/4a_limits_sme_wb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import discord
import config
import images
# -
class Client(discord.Client):
async def on_connect(self):
print(f"""
Client has successfully logged in as {client.user.name}#{client.user.discriminator}
Your discord ID is {client.user.id}
""")
async def on_message(self, message):
if message.author != client.user:
return
if message.content.count(config.prefix) >= 2:
emote_text = message.content
for image_name, image_value in images.emotes:
if config.prefix+image_name+config.prefix in emote_text:
emote_text = emote_text.replace(config.prefix+image_name+config.prefix,"\\"+image_value+"\\")
final_list = emote_text.split("\\")
if(len(final_list) > 0):
await message.delete()
for text in final_list:
if text != "":
await message.channel.send(text)
client = Client()
try:
client.run(config.token, bot = False)
except discord.LoginFailure:
print("ERROR: Client failed to log in. [Invalid token]")
except discord.HTTPException:
print("ERROR: Client failed to log in. [Unknown Error]")
| EmoteConvert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from shapely.geometry import box, Polygon
import folium
import requests
import json
import plotly.express as px
import pandas as pd
m = folium.Map(location=[40.28, -89.39])
# +
# playing around with folium, this is probably going to replace the google maps call in place on the app currently
url = (
'https://raw.githubusercontent.com/deldersveld/topojson/master/countries/us-states/IL-17-illinois-counties.json'
)
m = folium.Map(width = 500,
height = 500,
location=[40.28, -89.39],
tiles="cartodbpositron",
zoom_start=5,
)
folium.Marker(
location=[36.9815, -91.51135],
popup='reference',
).add_to(m)
folium.Circle(
radius=1115,
location=[40.28, -89.39],
popup="Chicago",
color="red",
fill=True,
fill_color='red'
).add_to(m)
# folium.GeoJson(url, name="geojson").add_to(m)
folium.TopoJson(
json.loads(requests.get(url).text),
"objects.cb_2015_illinois_county_20m",
name="topojson",
).add_to(m)
folium.LayerControl().add_to(m)
m
# -
m.save('m.html')
# +
# summoning and inspecting the topojson data
alpha = json.loads(requests.get(url).text)
# +
# coordinates for reference in understanding above data
# 41.148339, -89.638487
# 36.981528, -91.511353
# +
# inspecting the scale key under the transform key inside of the alpha dictionary
lat_ratio = ((41.148339 - 36.981528)/3344)
long_ratio = ((-89.638487 + 91.511353)/1061)
print(lat_ratio, 'lat ratio')
print(long_ratio, 'long ratio')
# +
# One degree of latitude equals approximately 364,000 feet (69 miles), one minute equals 6,068 feet (1.15 miles),
# and one-second equals 101 feet. One-degree of
# longitude equals 288,200 feet (54.6 miles), one minute equals 4,800 feet (0.91 mile),
# and one second equals 80 feet.
# +
# fitting polygons to shapely Polygon objects
alpha = json.loads(requests.get(url).text)
beta = alpha['arcs']
polys = []
county_names = []
for i, x in enumerate(alpha['objects']['cb_2015_illinois_county_20m']['geometries']):
poly_shape = []
for y in x['arcs'][0]:
if y<0:
temp = []
y = abs(y+1)
arc_ind = beta[y].copy()
ref = np.array(arc_ind[0])
temp.append(ref)
for j in range(1, len(arc_ind)):
ref = ref + np.array(arc_ind[j])
temp.append(ref)
for zed in reversed(temp):
poly_shape.append(zed)
else:
arc_ind = beta[y].copy()
ref = np.array(arc_ind[0])
poly_shape.append(ref)
for j in range(1, len(arc_ind)):
ref = ref + np.array(arc_ind[j])
poly_shape.append(ref)
shape_final = Polygon(poly_shape)
polys.append(shape_final)
county_names.append(x['properties']['NAME'])
# +
# take a look at marshall county!
polys[0]
# perfect
# -
for x in polys:
display(x)
# +
# testing to see that these distinct and seperate polygons are not intersecting:
marshall = polys[0]
cumberland = polys[1]
marshall.intersects(cumberland)
#False -- excellent!
# +
import math
# functions to determine points of circle
# This function gets just one pair of coordinates based on the angle theta
def get_circle_coord(theta, x_center, y_center, radius):
x = radius * math.cos(theta) + x_center
y = radius * math.sin(theta) + y_center
return [int(x),int(y)]
# This function gets all the pairs of coordinates
def get_all_circle_coords(x_center, y_center, radius, n_points):
thetas = [i/n_points * math.tau for i in range(n_points+1)]
circle_coords = [get_circle_coord(theta, x_center, y_center, radius) for theta in thetas]
return circle_coords
# Using the second function to generate all the pairs of coordinates.
# circle_coords = get_all_circle_coords(x_center = 5,
# y_center = 15,
# radius = 2.5,
# n_points = 5000)
# +
# testing above functions
get_all_circle_coords(0, 0, 50, 24)
# yep, thats a circle!
# +
# developing function to return intersections of circle radius
# enter radius in miles
coords = [41.015051, -89.307540]
def find_intersecting_counties(lat, long, radius):
"""Function to return intersecting counties"""
refs = [36.981528, -91.511353]
lat_ratio = ((41.148339 - 36.981528)/3344)
long_ratio = ((-89.638487 + 91.511353)/1061)
radius = radius/69
radius = int(radius/lat_ratio)
lat_pre = abs(refs[0] - lat)
long_pre = abs(refs[1] - long)
y_coord = int(long_pre/long_ratio)
x_coord = int(lat_pre/lat_ratio)
center = [y_coord, x_coord]
coords = get_all_circle_coords(center[0], center[1], radius, 60)
circle = Polygon(coords)
intersectors = []
for i, shape in enumerate(polys):
if shape.intersects(circle):
intersectors.append(i)
return intersectors, circle
# -
delta, beta = find_intersecting_counties(coords[0], coords[1], 10)
for x in delta:
print(county_names[x])
beta
for x in polys:
display(x)
df = pd.read_csv('<csv_here>')
df.head()
| test_app/Map_creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# LVQ for the Ionosphere Dataset
from random import seed
from random import randrange
from csv import reader
from math import sqrt
# Load a CSV file
def load_csv(filename):
dataset = list()
with open(filename, 'r') as file:
csv_reader = reader(file)
for row in csv_reader:
if not row:
continue
dataset.append(row)
return dataset
# Convert string column to float
def str_column_to_float(dataset, column):
for row in dataset:
row[column] = float(row[column].strip())
# Convert string column to integer
def str_column_to_int(dataset, column):
class_values = [row[column] for row in dataset]
unique = set(class_values)
lookup = dict()
for i, value in enumerate(unique):
lookup[value] = i
for row in dataset:
row[column] = lookup[row[column]]
return lookup
# Split a dataset into k folds
def cross_validation_split(dataset, n_folds):
dataset_split = list()
dataset_copy = list(dataset)
fold_size = int(len(dataset) / n_folds)
for i in range(n_folds):
fold = list()
while len(fold) < fold_size:
index = randrange(len(dataset_copy))
fold.append(dataset_copy.pop(index))
dataset_split.append(fold)
return dataset_split
# Calculate accuracy percentage
def accuracy_metric(actual, predicted):
correct = 0
for i in range(len(actual)):
if actual[i] == predicted[i]:
correct += 1
return correct / float(len(actual)) * 100.0
# Evaluate an algorithm using a cross validation split
def evaluate_algorithm(dataset, algorithm, n_folds, *args):
folds = cross_validation_split(dataset, n_folds)
scores = list()
for fold in folds:
train_set = list(folds)
train_set.remove(fold)
train_set = sum(train_set, [])
test_set = list()
for row in fold:
row_copy = list(row)
test_set.append(row_copy)
row_copy[-1] = None
predicted = algorithm(train_set, test_set, *args)
actual = [row[-1] for row in fold]
accuracy = accuracy_metric(actual, predicted)
scores.append(accuracy)
return scores
# calculate the Euclidean distance between two vectors
def euclidean_distance(row1, row2):
distance = 0.0
for i in range(len(row1)-1):
distance += (row1[i] - row2[i])**2
return sqrt(distance)
# Locate the best matching unit
def get_best_matching_unit(codebooks, test_row):
distances = list()
for codebook in codebooks:
dist = euclidean_distance(codebook, test_row)
distances.append((codebook, dist))
distances.sort(key=lambda tup: tup[1])
return distances[0][0]
# Make a prediction with codebook vectors
def predict(codebooks, test_row):
bmu = get_best_matching_unit(codebooks, test_row)
return bmu[-1]
# Create a random codebook vector
def random_codebook(train):
n_records = len(train)
n_features = len(train[0])
codebook = [train[randrange(n_records)][i] for i in range(n_features)]
return codebook
# Train a set of codebook vectors
def train_codebooks(train, n_codebooks, lrate, epochs):
codebooks = [random_codebook(train) for i in range(n_codebooks)]
for epoch in range(epochs):
rate = lrate * (1.0-(epoch/float(epochs)))
for row in train:
bmu = get_best_matching_unit(codebooks, row)
for i in range(len(row)-1):
error = row[i] - bmu[i]
if bmu[-1] == row[-1]:
bmu[i] += rate * error
else:
bmu[i] -= rate * error
return codebooks
# LVQ Algorithm
def learning_vector_quantization(train, test, n_codebooks, lrate, epochs):
codebooks = train_codebooks(train, n_codebooks, lrate, epochs)
predictions = list()
for row in test:
output = predict(codebooks, row)
predictions.append(output)
return(predictions)
# Test LVQ on Ionosphere dataset
seed(1)
# load and prepare data
filename = 'ionosphere.data.csv'
dataset = load_csv(filename)
for i in range(len(dataset[0])-1):
str_column_to_float(dataset, i)
# convert class column to integers
str_column_to_int(dataset, len(dataset[0])-1)
# evaluate algorithm
n_folds = 5
learn_rate = 0.3
n_epochs = 50
n_codebooks = 20
scores = evaluate_algorithm(dataset, learning_vector_quantization, n_folds, n_codebooks, learn_rate, n_epochs)
print('Scores: %s' % scores)
print('Mean Accuracy: %.3f%%' % (sum(scores)/float(len(scores))))
# -
| lvq/lvq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import glob2
location = 'D:\\FELion-Spectrum-Analyser\\testing\\_datas'
os.chdir(location)
png = glob2.glob('*.png')
png, type(png[0])
# +
from glob2 import glob as find
from os.path import join
find('*.mass')
# -
fname = 'pow'
location+"/OUT/{}.pdf".format(fname)
join(location, 'OUT', fname+'.pdf')
# +
file = '01_06_18-1.mass'
file1 = 'powerfile.pow'
string = '#SHOTS'
string1 = '#'
with open(file, 'r') as f:
info = f.readlines()
#for n, i in enumerate(info):
# print(n, i)
# +
import pandas as pd
df = pd.read_csv('massScan.mass')
# +
class add:
def __init__(self, x, y):
self.x = x
self.y = y
def result(self):
return self.x + self.y
def __repr__(self):
return '{}({}, {})'.format(add.__name__, self.x, self.y)
# +
import inspect
class A:
def a(self):
print("A.a()")
B().b()
class B:
def b(self):
print("B.b()")
stack = inspect.stack()
print(stack[1][0].f_locals["self"].__class__)
the_class = stack[1][0].f_locals["self"].__class__
the_method = stack[1][0].f_code.co_name
print(" I was called by {}.{}()".format(str(the_class), the_method))
A().a()
# +
from tkinter import *
master = Tk()
w = Scale(master, from_=0, to=100)
w.pack()
w = Scale(master, from_=0, to=200, orient=HORIZONTAL)
w.pack()
mainloop()
# -
import matplotlib.pyplot as plt
plt.axvline(0.4)
plt.axvline(0.3, color = 'C0')
from cycler import cycler
cycler(color='bgrcmyk'),
colors = []
for i in range(10):
c = 'C%i'%i
colors.append(c)
colors
# +
import tkinter as tk
class Example(tk.Frame):
def __init__(self, *args, **kwargs):
tk.Frame.__init__(self, *args, **kwargs)
self.l1 = tk.Label(self, text="Hover over me")
self.l2 = tk.Label(self, text="", width=40)
self.l1.pack(side="top")
self.l2.pack(side="top", fill="x")
self.l1.bind("<Enter>", self.on_enter)
self.l1.bind("<Leave>", self.on_leave)
def on_enter(self, event):
self.l2.configure(text="Hello world")
def on_leave(self, enter):
self.l2.configure(text="")
if __name__ == "__main__":
root = tk.Tk()
Example(root).pack(side="top", fill="both", expand="true")
root.mainloop()
# -
class P:
def __init__(self,x):
self.x = x
@property
def get(self):
return self.__x
@get.setter
def x(self, x):
if x < 0:
self.__x = 0
elif x > 1000:
self.__x = 1000
else:
self.__x = x
p1 = P(x = 0)
p1.x = 40
p1.x
| testing/archives/FELion_NormlineModifications.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Hypothesis testing with BayesDB
#
# ##### Prepared by <NAME>, PhD.
#
# ## Preamble
#
# This notebook is demonstrating hypothesis testing with BayesDB.
#
# **The demo is using early stage alpha version research software.** The demo is not
# intended to be used as is -- it won't scale to larger datasets and the
# interfaces described don't implement a reusable workflow.
# Instead it is intended to provide a snapshot of our current
# capability to solicit interest in BayesDB.
#
# One candidate beta version tailored to a scalable workflow will be able to deal
# with 100 000 rows and 1000 columns. A key question we will address in the research for this beta version is
# how dataset size, amount of computation, and accuracy of a synthetic data
# generator interact with predictive accuracy.
#
# ## BayesDB
#
# BayesDB is a probabilistic programming platform that provides built-in non-parametric Bayesian model discovery.
# BayesDB makes it easy for users without statistics training to search, clean, and model multivariate databases using an SQL-like language. In this notebook, we demonstrate how BayesDB can be used for hypothesis testing.
# The work is based on research results published here:
#
# * _Detecting dependencies in sparse, multivariate databases using probabilistic programming and non-parametric Bayes._ <NAME>.; and <NAME>. In Artificial Intelligence and Statistics 20 (AISTATS), volume 54, pages 632–641. PMLR, 2017 [(link)](http://proceedings.mlr.press/v54/saad17a/saad17a.pdf).
# * _Bayesian synthesis of probabilistic programs for automatic data modeling._ <NAME>.; <NAME>.; <NAME>.; <NAME>.; and <NAME>. Proceedings of the ACM on Programming Languages, 3(POPL): 37:1-37:29. January 2019 [(link)](https://dl.acm.org/citation.cfm?doid=3302515.3290350&fbclid=IwAR2GLpIo7YW2g_lSMpJDsgb2KZOIEX0dos5DZz2Vg1BcQBKbGTA3F5B_mmA).
# * _Crosscat: A fully Bayesian, nonparametric method for analyzing heterogeneous, high-dimensional data._ <NAME>.; <NAME>.; <NAME>.; <NAME>.; <NAME> <NAME>. Journal of Machine Learning Research, 17(138): 1-49. 2016. [(link)](http://jmlr.org/papers/volume17/11-392/11-392.pdf).
#
# ## Aim of this notebook
#
# The purpose of this notebook is twofold. First, we will provide a tutorial on
# probabilistic program synthesis for data modeling and analysis. Second, we show how the
# synthesized programs can be used for analysis.
# Both will be demoed using openly available RNA sequencing data (see the [notebook for
# download and pre-processing here](data-preprocessing.ipynb)). The notebook focuses on the
# on arabinose regulation and the operon of araC, the main gene involved in arabinose
# catabolism and transport.
#
# 
#
# ## Capabilities demonstrated in this notebook
#
# ### 1. Tutorial: Population assembly.
#
# The notion of a "population" is a central concept in BayesDB.
#
# ### 2. Tutorial: Probabilistic program synthesis.
#
# We synthesize generative models as probabilistic programs.
#
# ### 3. Tutorial: Inspecting synthesis results.
#
# We perform a quick, visual predictive check to assess whether our inference
# converged.
#
# ### 4. Simple hypothesis testing: BayesDB agrees with linear statistics.
#
# We confirm that one experimental treatment should not effect the other.
#
# ### 5. Simple hypothesis testing: BayesDB finds a dependency that linear statistics misses.
#
# We show a case where two genes are informative for each other because they
# live on the same [operon](https://ecocyc.org/gene?orgid=ECOLI&id=EG10054#tab=TU).
#
# ### 6. Causal hypothesis testing: BayesDB finds a evidence for a causal structure.
#
# We demonstrate testing of causal hypotheses.
#
# -------------
#
# #### Setting up the Jupyter environment
#
# The first step is to load the `jupyter_probcomp.magics` library, which provides BayesDB hooks for data exploration, plotting, querying, and analysis through this Jupyter notebook environment.
#
# It enables us to use three languages in the Jupyter cells below:
# 1. **BQL**. The Bayesian Query Language. This allows us to query a Bayesian database [(documentation)](http://probcomp.csail.mit.edu/dev/bayesdb/doc/bql.html).
# 2. **MML**. The Meta Modeling Language. This allows us to define the generative model that will model populations of interest.
#
#
# All of the cells below are by default python cells. We use the `%%` and `%`
# decorators to change the default behavior of jupyter.
# `%%bql` will introduce a whole cell of BQL code whereas `%bql`
# only marks the current line to be interpreted as BQL and the rest of the cell is
# python.
# %load_ext jupyter_probcomp.magics
# The cell allows plots from matplotlib and javascript to be shown inline.
# %matplotlib inline
# %vizgpm inline
# We load some helper functions for plotting.
run helper_functions.py
# #### Creating a BayesDB `.bdb` file on disk
#
# We next use the `%bayesdb` magic to create a `.bdb` file on disk named `bayesian_database.bdb`. This file will store all the data and models created in this session.
# %bayesdb bayesian-database.bdb
# ## 1. Tutorial: Population assembly
#
# #### Ingesting data from a `.csv` file into a BayesDB table
#
# We load the data we [previously created](data-munging.ipynb) from a csv file.Each column of the csv file is a variable, and each row is a record. We use the `CREATE TABLE` BQL query, with the pathname of the csv file, to convert the csv data into a database table named `data`.
# %bql CREATE TABLE "data" FROM 'rna-seq-araD-treatment.csv'
# #### Running basic queries on the table using BQL and SQL
#
# Now that the dataset has been loaded into at table, we can run standard SQL queries to explore the contents of the data.
# For example, we can select the first 5 records. Observe that each row in the
# table is a particular measurement, and each column is either a gene or an
# experimental treatment.
# %bql SELECT * FROM "data" LIMIT 5;
# Confirm the number of rows.
# %bql SELECT COUNT(*) AS "N rows in table" FROM "data";
# #### Creating a BayesDB population for the RNAseq data
#
# The notion of a "population" is a central concept in BayesDB. For a standard
# database table, such as `data`, each column is associated with a
# [data type](https://sqlite.org/datatype3.html), which in sqlite3 are
# `TEXT`, `REAL`, `INTEGER`, and `BLOB`. For a BayesDB population, each variable
# is associated with a _statistical data type_.
# In this tutorial, we will use the `NUMERICAL` and `NOMINAL` statistical data types (the latter for categorical random variables that take discrete values).
#
# We can use the `GUESS SCHEMA FOR <table>` command from the Metamodeling Language
# (MML) in BayesDB to guess the statistical data types of variables in the table.
#
# The guesses use heuristics based on the contents in the cells.
#
# %%mml
CREATE POPULATION FOR "data" WITH SCHEMA (
GUESS STATTYPES OF (*);
);
# %bql .interactive_pairplot --population=data SELECT "araC", "araB", "arabinose", "glucose" FROM data
# ## 2. Tutorial: Probabilistic program synthesis.
#
# Now that we have created the population, the next step is to analyze the data by
# synthezising probabilistic models as probabilistic programs which explain the data
# generating process. Probabilistic program synthesis for automated modeling in
# BayesDB is specified by declaring `GENERATOR` for a population (see
# [Bayesian synthesis of probabilistic programs for automatic data modeling](https://dl.acm.org/citation.cfm?doid=3302515.3290350&fbclid=IwAR2GLpIo7YW2g_lSMpJDsgb2KZOIEX0dos5DZz2Vg1BcQBKbGTA3F5B_mmA)
# ). The default generator in BayesDB is based on Cross-Categorization [(Crosscat)](http://jmlr.org/papers/v17/11-392.html). The CrossCat generator is a Bayesian factorial mixture model which learns a full joint distribution over all variables in the population, using a divide-and-conquer approach. We will explore CrossCat more in this notebook.
#
# For now we use MML to declare the a generator for the population. Note that that we have left the schema (there are not specific model commands or overrides), which will apply the built-in default model discovery strategies.
#
# %mml CREATE GENERATOR FOR "data";
# We turn BayesDB's multiprocessing on.
# %multiprocess on
# #### Bayesian inference
#
# After creating the generator, we now need to initialize `MODELS` for the
# generator. We can think of a `GENERATOR` as specifying a hypothesis space of
# explanations for the data generating process for the population, and each
# `MODEL` is a candidate hypothesis. We start by creating only 30 models, which
# are initialized __randomly__. You can also think of those model random
# initializations of 30 chains to run MCMC on (next cell).
# %mml INITIALIZE 30 MODELS IF NOT EXISTS FOR "data";
# We run Gibbs sampling for 5 minutes for each chain to reach an approximation of
# the posterior distribution that will allow us to draw synthetic data samples
# from.
#
# **Caveat:** For each chain a single available CPU is
# used. `%multiprocess on` tells BayesDB to run each available CPU in parallel. Thus,
# if we initialize 30 models/MCMC chains and run analysis for `5 MINUTES` but only have 6
# CPUs available, the cell will run for 25 minutes, 5 times it will run 6 models/MCMC chains
# in parallel for 6 minutes
# %mml ANALYZE "data" FOR 5 MINUTES (QUIET);
# Let's inspect the synthesized programs. **Caveat**: The following is stubbed. We synthesized the Metaprob programs with a pre-alpha prototype that we can't hand over yet. For this demo, we simply load previously
# synthesized files from disk.
n_models = 10 # For now, let's only export code for 10 out of the 30 models in the ensemble.
# bdb = %get_bdb
code = load_synthesized_code(bdb, n_models=n_models)
# Because there are 30 models in the ensemble, there is a lot of code. We print
# only the first model in the ensemble:
print code[125:4365] # Let's print one of the synthesized models.
# ## 3. Tutorial: Inspecting synthesis results.
#
#
# We use BQL's `SIMULATE` query to generate synthetic data from the synthezided programs. Note that we
# are simulating jointly from 6 columns -- if we wish to simulate the entire table we can
# simply mention all columns in the able in the query. We can also sample from conditional
# distributions using the phrase `GIVEN` in the query.
#
# We escape to python to simplify plotting below.
# table_simulate = %bql \
# SIMULATE \
# "araD", \
# "araA", \
# "araB", \
# "glucose", \
# "arabinose", \
# "araC" \
# FROM "data" \
# LIMIT 305
# We also save the observed data in a python dataframe.
# table_select = %bql \
# SELECT \
# "araD", \
# "araA", \
# "araB", \
# "glucose", \
# "arabinose", \
# "araC" \
# FROM "data"
# We'd like to use the two tables, `table_simulate` and `table_select` to asses
# how well BayesDB models the observed data. We do this by performing a simple, visual predictive check. We plot the generated data and check if what we synthetically generated agrees with the observed data.
columns = ["araA", "araB"]
plot_virtual_data(table_select[columns], table_simulate[columns]);
columns = ["arabinose", "araC"]
plot_virtual_data(table_select[columns], table_simulate[columns]);
columns = ["glucose", "araC"]
plot_virtual_data(table_select[columns], table_simulate[columns]);
columns = ["arabinose", "glucose"]
plot_virtual_data(table_select[columns], table_simulate[columns]);
# How does the code organize dependent columns in the data generator? Let's inspect
# the ensemble to find out. Dependent columns are grouped together and form a multivariate mixture model.
# For example, in the following code snippet from the first model in the ensemble,
# AraC and arabinose are grouped together in the code:
# ```
# (define cluster-id-for-araC-arabinose
# (categorical [0.599820 0.287668 0.031969 0.028648 0.051894]))
# (define [araC-mean araC-std]
# (cond
# (= cluster-id-for-araC-arabinose 0) [7.813161 0.634979]
# (= cluster-id-for-araC-arabinose 1) [9.185628 0.566170]
# (= cluster-id-for-araC-arabinose 2) [13.299314 0.743700]
# (= cluster-id-for-araC-arabinose 3) [9.725188 0.454566]
# (= cluster-id-for-araC-arabinose 4) [8.269303 0.674934]))
# (define araC
# (gaussian araC-mean araC-std))
#
# (define [arabinose-mean arabinose-std]
# (cond
# (= cluster-id-for-araC-arabinose 0) [0.078673 1.020788]
# (= cluster-id-for-araC-arabinose 1) [8.193512 0.864407]
# (= cluster-id-for-araC-arabinose 2) [16.518427 1.154188]
# (= cluster-id-for-araC-arabinose 3) [-0.151569 0.996753]
# (= cluster-id-for-araC-arabinose 4) [16.605226 0.732904]))
# (define arabinose
# (gaussian arabinose-mean arabinose-std))
# ```
# First, a cluster ID (corresponding to a single mixture component) is sampled
# ```
# (define cluster-id-for-araC-arabinose
# (categorical [0.599820 0.287668 0.031969 0.028648 0.051894]))
# ```
# The cluster ID determines the parameterization of the Gaussian distributions for
# araC and arabinose.
#
# We can inspect all the synthesized cluster IDs:
#
for line_of_code in code.split("\n"):
if 'define cluster-id' in line_of_code:
print line_of_code
# This information can be used to create a causal graphical model summarizing the
# data-generating process in every model of the ensemble.
model_index = 0
render_dependence(get_dependence_from_code(model_index, code))
# %mml .render_crosscat --subsample=50 data 0
model_index = 1
render_dependence(get_dependence_from_code(model_index, code))
# %mml .render_crosscat --subsample=50 data 1
model_index = 7
render_dependence(get_dependence_from_code(model_index, code))
# %mml .render_crosscat --subsample=50 data 7
# We can define a function to compute the fraction of times two columns end up
# dependent in the code.
def count_structural_dependence(col1, col2):
"""Check the synthesized code to check how often two columns are dependent."""
count = 0.
for line_of_code in code.split("\n"):
if 'define cluster-id' in line_of_code:
if col1 in line_of_code and col2 in line_of_code:
count+=1.
return count/n_models
# What is the probability given the code of 10 models that `glucose`
# and `araD` are dependent?
count_structural_dependence('glucose', 'araD')
# Note that so far, we have only generated code for 10 out of the 30 models in
# the ensemble.
n_models = 30
code = load_synthesized_code(bdb, n_models=n_models)
# Let's recompute the probability of dependence given code for all 30 models in the ensemble.
count_structural_dependence('glucose', 'araD')
# The BQL language provides a convenient user interface to write such a query to
# synthesized code.
# %%bql
ESTIMATE DEPENDENCE PROBABILITY OF
"glucose" WITH "araD" AS "Probability of dependence"
FROM "data"
LIMIT 1
# This allows a user to write and combine more interesting queries. For example,
# we can create a table for all pairwise, dependencies of columns in the data. This provides
# insights in predictive relationships. A cell (between 0 and 1) is the fraction of posterior CrossCat models in the ensemble where those
# two variables are detected to be probably dependent (i.e. they are in the same column grouping).m
# %%bql
CREATE TABLE "dependencies" AS
ESTIMATE
DEPENDENCE PROBABILITY AS "Probability of dependence"
FROM PAIRWISE VARIABLES OF data;
# We can show the results of this analysis for the column `glucose`.
# %bql .bar SELECT "name1" AS "column", "Probability of dependence"\
# FROM "dependencies" WHERE "name0" == 'glucose' ORDER BY "Probability of dependence" DESC
# The plot above corresponds to one row below.
# %bql .interactive_heatmap SELECT name0, name1, "Probability of dependence" FROM dependencies;
# #### Comparison to linear statistics
#
# Let us compare dependence probabilities from CrossCat to linear (Pearson r) correlation values, a very common technique for finding predictive relationships. We can compute the Pearson R (and its p-value) in BayesDB using the `CORRELATION` and `CORRELATION PVALUE` queries. The following cell creates a table named `correlations`, which contains the R and p-value for all pairs of variables.
# %%bql
CREATE TABLE "correlations" AS
ESTIMATE
CORRELATION AS "correlation",
CORRELATION PVALUE AS "pvalue"
FROM PAIRWISE VARIABLES OF "data"
# Plotting all the pairwise correlations reveals that linear analysis misses many
# predictive relationship that our query for `DEPENDENCE PROBABILITY` finds:
# %bql .interactive_heatmap SELECT name0, name1, "correlation" FROM "correlations"
# ## 4. Simple hypothesis testing: BayesDB agrees with linear statistics.
#
# Finally, we get to test hypotheses. In the following, we will confirm that
# BayesDB finds relationships in the data that we know should be there. For every
# hypothesis test, we compare BayesDB with a linear method (regression/correlation/partial
# correlation).
#
# We start with a simple case. One experimental treatment should not effect the
# other -- they should be statistically independent; which is confirmed by running
# a regression and BayesDB.
# #### Linear statistics
# data = %bql SELECT * FROM "data"
fig, ax = corr_plots('arabinose', 'glucose', data, xlims=[-3, 20])
# #### BayesDB
# %bql .density --xmax=0.6 SIMULATE MUTUAL INFORMATION OF "arabinose" WITH "glucose" USING 200 SAMPLES FROM MODELS OF "data"
# ## 5. Simple hypothesis testing: BayesDB finds a dependency that linear statistics misses.
#
# Next, we show a case where two genes are informative for each other. We know
# this is true because the two genes, `araC` and `araA`, live on the same
# [operon](https://ecocyc.org/gene?orgid=ECOLI&id=EG10054#tab=TU).
# However, the relationship is not well modelled by a line.
# Thus, regression does not detect it but BayesDB does.
# #### Linear statistics
fig, ax = corr_plots('araA', 'araC', data, xlims=[5, 14])
# #### BayesDB
# %bql .density --xmax=0.5 SIMULATE MUTUAL INFORMATION OF "araA" WITH "araC" USING 200 SAMPLES FROM MODELS OF "data"
# ## 6. Causal hypothesis testing: BayesDB finds a evidence for a causal structure.
#
# Sections 4. and 5. showed fairly simple examples of pairwise relationships. But
# BayesDB allows for testing for richer, causal hypotheses, too.
#
#
# The following diagram describes the causal ground truth between three variables in our
# data set. `araC` is a gene that is effected by both `Arabinose` and `Glucose`
# (see [this summary of `araC`)](https://ecocyc.org/gene?orgid=ECOLI&id=EG10054#).
# Arabinose and glucose are both experimental variables treatments. We can assume
# that their administration is independent (there is no edge between `Arabinose` and `Glucose`).
#
# 
#
#
# If we know that the causal structure (above) holds, then the following
# relationships are implied and should inferable through the data:
#
# * Arabinose$\,\;\not\!\perp\!\!\!\perp$ araC
# * Glucose $\;\;\;\not\!\perp\!\!\!\perp$ araC
# * Arabinose$\;\perp \!\!\! \perp$ Glucose"
# * Arabinose $\;\not\!\perp\!\!\!\perp$ Glucose | araC
#
# Where $\perp \!\!\! \perp$ and $\not\!\perp\!\!\!\perp$ means statistical independence and dependence.
# Note that conditioning on araC breaks independence between glucose and
# arabinose.
#
# Let's define those relationships and then see how we can investigate evidence
# for those relationships with (i) linear statistics and (ii) BayesDB.
relationships = [
'arabinose with araC',
'glucose with araC',
'arabinose with glucose',
'arabinose with glucose given araC'
]
# #### Linear statistics
# We compare against [partial correlation](https://en.wikipedia.org/wiki/Partial_correlation) for the case where we condition on `araC`.
# Linear correlation out of BQL
# corr_xz = %bql ESTIMATE CORRELATION OF "arabinose" WITH "araC" AS "r" BY "data"
# corr_yz = %bql ESTIMATE CORRELATION OF "glucose" WITH "araC" AS "r" BY "data"
# corr_xy = %bql ESTIMATE CORRELATION OF "arabinose" WITH "glucose" AS "r" BY "data"
# Partial correlation with python.
# df = %bql SELECT "arabinose", "glucose", "araC" FROM data
corr_xy_given_z = partial_correlation(df.values)
corr_estimates = [corr_xz, corr_yz, corr_xy, corr_xy_given_z]
# #### BayesDB
# mi_xz = %bql SIMULATE MUTUAL INFORMATION OF "arabinose" WITH "araC" USING 200 SAMPLES FROM MODELS OF "data"
# mi_yz = %bql SIMULATE MUTUAL INFORMATION OF "glucose" WITH "araC" USING 200 SAMPLES FROM MODELS OF "data"
# mi_xy = %bql SIMULATE MUTUAL INFORMATION OF "arabinose" WITH "glucose" USING 200 SAMPLES FROM MODELS OF "data"
# mi_xy_given_z = %bql SIMULATE MUTUAL INFORMATION OF "arabinose" WITH "glucose" GIVEN ("araC" = 13) \
# USING 200 SAMPLES FROM MODELS OF "data"
mi_estimates = [mi_xz, mi_yz, mi_xy, mi_xy_given_z]
# ### Comparison
for i, corr in enumerate(corr_estimates):
print 'Pearson r^2 of {}: {}'.format(relationships[i], corr.values[0,0])
for i, mi in enumerate(mi_estimates):
print 'Mutial information of {}: {}'.format(relationships[i], mi.mean()[0])
# The final value if shifting if one conditions on araC! Let's visualize the posterior
# distribution of mutual information.
plot_information_flow(mi_estimates, relationships)
| experiments/gene-expression-public/RNAseq-synthesis-and-hypothesis-testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
data = [0.5, None, None, 0.52, 0.54, None, None, 0.59, 0.6, None, 0.7]
s = pd.Series(data)
s
# ### Atribuir valores aos valores nulos
# +
# s.fillna(0)
# 0 0.50
# 1 0.00
# 2 0.00
# 3 0.52
# 4 0.54
# 5 0.00
# 6 0.00
# 7 0.59
# 8 0.60
# 9 0.00
# 10 0.70
# dtype: float64
# -
s.fillna(method = 'ffill')
# #### Esses métodos são muito válidos quanto estamos trabalhando com uma série de tempo, em que podemos encontrar dados nulos que podem ser preenchidos de forma rápida, tornando seu modelo executável.
s.fillna(method = 'bfill')
# #### Realiza a mesma análise das assinaturas em uma Series, mas dessa vez de baixo para cima, isto é, do último elemento da Series ao primeiro.
# #### Por meio de fillna(), coletaremos a média de todos os valores não-nulos e a usaremos como preenchimento. Para tanto, basta escrever s.mean().
s.fillna(s.mean())
s.fillna(method = 'ffill', limit = 1)
# Como resultado, veremos que a linha 1, originalmente nula, foi preenchida com o valor de 0, a linha acima.
# Em 2, o resultado permaneceu nulo, afinal colocamos um limite de interpolação de 1.
# #### Podemos, ainda, preencher as assinaturas nulas usando ora o valor da linha acima ora o valor da linha abaixo. Armazenaremos nossa Series em s1
s1 = s.fillna(method = 'ffill', limit = 1)
s1
s2 = s1.fillna(method = 'bfill', limit = 1)
s2
atletas = pd.DataFrame([['Marcos', 9.62], ['Pedro', None], ['João', 9.69],
['Beto', 9.72], ['Sandro', None], ['Denis', 9.69],
['Ary', None], ['Carlos', 9.74]],
columns = ['Corredor', 'Melhor Tempo'])
atletas
atletas.fillna(atletas['Melhor Tempo'].mean(), inplace=True)
atletas
| DS_02_Python_Pandas_Tratando_e_Analisando_Dados/extra_aula6_metodos_de_interpolacao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import all required functions and libraries
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import syft as sy
import copy
import numpy as np
import time
from torchvision import datasets, transforms
from torch.utils.data import DataLoader, Dataset
from ipynb.fs.full.FLDataset import load_dataset, getActualImages
from ipynb.fs.full.utils import averageModels
# +
# arguments class
class Arguments():
def __init__(self):
self.images = 60000
self.clients = 20
self.rounds = 2
self.epochs = 2
self.local_batches = 1
self.lr = 0.01
self.C = 0.8
self.drop_rate = 0.2
self.torch_seed = 0
self.log_interval = 500
self.iid = 'iid'
self.split_size = int(self.images / self.clients)
self.use_cuda = False
self.save_model = False
args = Arguments()
use_cuda = args.use_cuda and torch.cuda.isavailable()
device = torch.device("cuda" if use_cuda else "cpu")
kwargs = {'num_workers' : 1, 'pin_memory': True} if use_cuda else {}
# +
# hook
hook = sy.TorchHook(torch)
clients = []
for i in range(args.clients):
clients.append({"hook" : sy.VirtualWorker(hook, id="client{}".format(i+1))})
# +
global_train, global_test, train_group, test_group = load_dataset(args.clients, args.iid)
# -
for inx, client in enumerate(clients):
client['trainset'] = getActualImages(global_train, list(train_group[inx]), args.local_batches)
client['testset'] = getActualImages(global_test, list(test_group[inx]), args.local_batches)
client['samples'] = len(client['trainset']) / args.images
transform = transforms.Compose([transforms.ToTensor(), transforms.Normalize((0.1307,),(0.3081,))])
global_test_dataset = datasets.MNIST('./data', train=False, download=True, transform=transform)
global_test_loader = DataLoader(global_test_dataset, batch_size=args.local_batches, shuffle=True)
# +
# Network class
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 20, 5, 1)
self.conv2 = nn.Conv2d(20, 50, 5, 1)
self.fc1 = nn.Linear(4*4*50, 500)
self.fc2 = nn.Linear(500, 10)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2, 2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, 2, 2)
x = x.view(-1, 4*4*50)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x, dim=1)
# +
# ClientUpdate class
def ClientUpdate(args, device, client):
client['model'].train()
client['model'].send(client['hook'])
for epoch in range(1, args.epochs + 1):
for batch_idx, (data, target) in enumerate(client['trainset']):
data = data.send(client['hook'])
target = target.send(client['hook'])
data, target = data.to(device), target.to(device)
client['optim'].zero_grad()
output = client['model'](data)
loss = F.nll_loss(output, target)
loss.backward()
client['optim'].step()
if batch_idx % args.log_interval == 0:
loss = loss.get()
print('Model {} Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
client['hook'].id,
epoch, batch_idx * args.local_batches, len(client['trainset']) * args.local_batches,
100. * batch_idx / len(client['trainset']), loss))
client['model'].get()
# +
# test function
def test(args, model, device, test_loader, name):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
test_loss += F.nll_loss(output, target, reduction='sum').item() # sum up batch loss
pred = output.argmax(1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
print('\nTest set: Average loss for {} model: {:.4f}, Acccuracy: {}/{} ({:.0f}%)\n'.format(
name, test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
# +
# Federated
torch.manual_seed(args.torch_seed)
global_model = Net()
for client in clients:
torch.manual_seed(args.torch_seed)
client['model'] = Net().to(device)
client['optim'] = optim.SGD(client['model'].parameters(), lr=args.lr)
for fed_round in range(args.rounds):
m = int(max(args.C * args.clients, 1)) # here 1, so that at least 1 client will be chosen for a round
np.random.seed(fed_round)
selected_clients_inds = np.random.choice(range(len(clients)), m, replace=False)
selected_clients = [clients[i] for i in selected_clients_inds]
np.random.seed(fed_round)
active_clients_inds = np.random.choice(selected_clients_inds, int((1-args.drop_rate) * m), replace=False)
active_clients = [clients[i] for i in active_clients_inds]
# training
for client in active_clients:
ClientUpdate(args, device, client)
# testing the local client trained models
for client in active_clients:
test(args, client['model'], device, client['testset'], client['hook'].id)
# Averaging
global_model = averageModels(global_model, active_clients)
# Testing the average model to see the accuracy
test(args, global_model, device, global_test_loader, 'Global')
# Share the global model with the clients
for client in clients:
client['model'].load_state_dict(global_model.state_dict())
if (args.save_model):
torch.save(global_model.state_dict(), "FedAvg.pt")
# +
# weights of the global model
list(global_model.parameters())
# +
# 1st client's gradients
list(clients[0]['model'].parameters())[0].grad
| federated_techniques/FedAvg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # quant-econ Solutions: Infinite Horizon Dynamic Programming
# Solutions for http://quant-econ.net/py/optgrowth.html
# Our objective is to compute the policy functions in the figures from the lecture URL give above.
# %matplotlib inline
# First let's add a few imports
from __future__ import division # Not needed for Python 3.x
import numpy as np
import matplotlib.pyplot as plt
import quantecon as qe
# Now, we could import the functionality we need from `optgrowth_v0.py` or we could load that file here. A good software engineer would probably tell us to go for the first option, so as to avoid having the same code in two places. But sometimes it helps to break the rules. We want to break the rules here because it's nice to have the code all together in one place. So here's an excert from that file:
# +
from numpy import log
from scipy.optimize import fminbound
from scipy import interp
# Primitives and grid
alpha = 0.65
beta = 0.95
grid_max = 2
grid_size = 150
grid = np.linspace(1e-6, grid_max, grid_size)
# Exact solution
ab = alpha * beta
c1 = (log(1 - ab) + log(ab) * ab / (1 - ab)) / (1 - beta)
c2 = alpha / (1 - ab)
def v_star(k):
return c1 + c2 * log(k)
def bellman_operator(w):
"""
The approximate Bellman operator, which computes and returns the updated
value function Tw on the grid points.
* w is a flat NumPy array with len(w) = len(grid)
The vector w represents the value of the input function on the grid
points.
"""
# === Apply linear interpolation to w === #
Aw = lambda x: interp(x, grid, w)
# === set Tw[i] equal to max_c { log(c) + beta w(f(k_i) - c)} === #
Tw = np.empty(grid_size)
for i, k in enumerate(grid):
objective = lambda c: - log(c) - beta * Aw(k**alpha - c)
c_star = fminbound(objective, 1e-6, k**alpha)
Tw[i] = - objective(c_star)
return Tw
# -
# Now we need a function to get the greedy policy from an approximate value function.
def compute_greedy(w):
Aw = lambda x: interp(x, grid, w)
sigma = np.empty(grid_size)
for i, k in enumerate(grid):
objective = lambda c: - log(c) - beta * Aw(k**alpha - c)
sigma[i] = fminbound(objective, 1e-6, k**alpha)
return sigma
# ## Exercise 1
# +
alpha, beta = 0.65, 0.95
true_sigma = (1 - alpha * beta) * grid**alpha
fig, ax = plt.subplots(3, 1, figsize=(8, 10))
for i, n in enumerate((2, 4, 6)):
ax[i].set_ylim(0, 1)
ax[i].set_xlim(0, 2)
ax[i].set_yticks((0, 1))
ax[i].set_xticks((0, 2))
w = 5 * log(grid) - 25 # Initial condition
v_star = qe.compute_fixed_point(bellman_operator, w,
max_iter=n,
verbose=0)
sigma = compute_greedy(v_star)
ax[i].plot(grid, sigma, 'b-', lw=2, alpha=0.8, label='approximate optimal policy')
ax[i].plot(grid, true_sigma, 'k-', lw=2, alpha=0.8, label='true optimal policy')
ax[i].legend(loc='upper left')
ax[i].set_title('{} value function iterations'.format(n))
# -
# ## Exercise 2
# For this exercise we need to compute the optimal policy at a number of different parameter values. This is quite awkward to do given the way the code is written above. (One of the purposes of the present exercise is to highlight this fact.) In [a follow up lecture](http://quant-econ.net/py/optgrowth_2.html) we'll try to write code that can manage operating over different parameters in a nice way, using functions and classes. Here we'll just copy and paste from above. Note, however, that copying and pasting is almost always the worst solution (including this case too). The reason is that it violates the basic principle [do not repeat yourself](https://en.wikipedia.org/wiki/Don't_repeat_yourself).
#
# Bearing the above in mind, for now our (second best) strategy will be to copy the code for the Bellman operator and add in `beta` as an argument to the function.
def bellman_operator(w, beta=0.96):
# === Apply linear interpolation to w === #
Aw = lambda x: interp(x, grid, w)
# === set Tw[i] equal to max_c { log(c) + beta w(f(k_i) - c)} === #
Tw = np.empty(grid_size)
for i, k in enumerate(grid):
objective = lambda c: - log(c) - beta * Aw(k**alpha - c)
c_star = fminbound(objective, 1e-6, k**alpha)
Tw[i] = - objective(c_star)
return Tw
# Let's also do the same for `compute_greedy`
def compute_greedy(w, beta=0.96):
Aw = lambda x: interp(x, grid, w)
sigma = np.empty(grid_size)
for i, k in enumerate(grid):
objective = lambda c: - log(c) - beta * Aw(k**alpha - c)
sigma[i] = fminbound(objective, 1e-6, k**alpha)
return sigma
# Now let's compute the policies and the figures
# +
w = 5 * np.log(grid) - 25 # To be used as an initial condition
discount_factors = (0.9, 0.94, 0.98)
series_length = 25
fig, ax = plt.subplots(figsize=(8,5))
ax.set_xlabel("time")
ax.set_ylabel("capital")
ax.set_ylim(0.10, 0.30)
for beta in discount_factors:
w = 5 * log(grid) - 25 # Initial condition
v_star = qe.compute_fixed_point(bellman_operator,
w,
verbose=False,
beta=beta)
sigma = compute_greedy(v_star, beta)
# Compute the corresponding time series for capital
k = np.empty(series_length)
k[0] = 0.1
sigma_function = lambda x: interp(x, grid, sigma)
for t in range(1, series_length):
k[t] = k[t-1]**alpha - sigma_function(k[t-1])
ax.plot(k, 'o-', lw=2, alpha=0.75, label=r'$\beta = {}$'.format(beta))
ax.legend(loc='lower right')
plt.show()
# -
| optgrowth/optgrowth_solutions_py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 2
# Write a function to compute the roots of a mathematical equation of the form
# \begin{align}
# ax^{2} + bx + c = 0.
# \end{align}
# Your function should be sensitive enough to adapt to situations in which a user might accidentally set $a=0$, or $b=0$, or even $a=b=0$. For example, if $a=0, b\neq 0$, your function should print a warning and compute the roots of the resulting linear function. It is up to you on how to handle the function header: feel free to use default keyword arguments, variable positional arguments, variable keyword arguments, or something else as you see fit. Try to make it user friendly.
#
# Your function should return a tuple containing the roots of the provided equation.
#
# **Hint:** Quadratic equations can have complex roots of the form $r = a + ib$ where $i=\sqrt{-1}$ (Python uses the notation $j=\sqrt{-1}$). To deal with complex roots, you should import the `cmath` library and use `cmath.sqrt` when computing square roots. `cmath` will return a complex number for you. You could handle complex roots yourself if you want, but you might as well use available libraries to save some work.
# +
import cmath
def find_root(a,b,c):
if (a==0 and b==0 and c==0):
print("warning!\n x has infinite numbers")
return()
elif (a==0 and b==0 and c!=0):
print("error!\n no x")
return()
elif (a==0 and b!=0):
print("warning!\n x=",-c/b)
return(-c/b)
else:
x1=(-b+cmath.sqrt(b*b-4*a*c))/(2*a)
x2=(-b-cmath.sqrt(b*b-4*a*c))/(2*a)
print("x1=",x1)
print("x2=",x2)
return(x1,x2)
find_root(0,0,0)
# -
| lectures/L5/Exercise_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_keras)
# language: python
# name: conda_keras
# ---
# +
import csv
import numpy as np
from skimage.io import imread
from skimage.transform import resize
from keras.utils import np_utils
import keras
from keras.layers import Input, Dense, Dropout, Activation, Flatten, Conv2D,MaxPool2D,BatchNormalization
from keras.models import Sequential, Model
from keras.optimizers import Adam
import matplotlib.pyplot as plt
# %matplotlib inline
# +
#define a class to load yeast data
#keras code data
from keras.utils.data_utils import get_file
#data folderを作成して学習に必要な画像データをダウンロードします。
# #!mkdir data && cd data && wget http://kodu.ut.ee/~leopoldp/2016_DeepYeast/data/main.tar.gz -O - | tar -xz
# +
#ラベル情報を取得します
#train data
from skimage.io import imread
from skimage.transform import resize
import os
def load_data(train=True,valid=False,test=False):
'''
Codes adopted from keras https://github.com/keras-team/keras/blob/master/keras/utils/data_utils.py
Downloads and load files from if it not already in the cache..
by default, it will be saved to ~/.keras
'''
paths = ["data","trainlabel","validlabel","testlabel"]
data_path = get_file(paths[0],
origin="http://kodu.ut.ee/~leopoldp/2016_DeepYeast/data/main.tar.gz",
extract=True,
cache_subdir='deepyeast')
train_path = get_file(paths[1],origin="http://kodu.ut.ee/~leopoldp/2016_DeepYeast/code/reports/HOwt_train.txt",
cache_subdir='deepyeast')
val_path = get_file(paths[2],origin="http://kodu.ut.ee/~leopoldp/2016_DeepYeast/code/reports/HOwt_val.txt",
cache_subdir='deepyeast')
test_path = get_file(paths[3],origin="http://kodu.ut.ee/~leopoldp/2016_DeepYeast/code/reports/HOwt_test.txt",
cache_subdir='deepyeast')
label_name = ['Cell_periphery','Cytoplasm',
'endosome','ER','Golgi',
'Mitochondrion','Nuclear_Periphery',
'Nucleolus','Nucleus','Peroxisome',
'Spindle_pole','Vacuole']
print(data_path)
data_path, _ = os.path.split(data_path)
#画像データ格納用変数
X_train = []
X_valid = []
X_test = []
#ラベル格納用変数
y_train = []
y_valid = []
y_test = []
if train:
with open(train_path) as f:
reader = csv.reader(f)
for row in reader:
row = row[0].split(" ")
image_path = row[0]
image_path = os.path.join(data_path,image_path)
image = imread(image_path)
image = resize(image,(64,64)) #this will change the numpy range to 0 to 1
X_train.append(image)
label = row[1]
y_train.append(label)
X_train = np.array(X_train)
y_train = np.array(y_train)
y_train = np_utils.to_categorical(y_train)
if valid:
with open(valid_path) as f:
reader = csv.reader(f)
for row in reader:
row = row[0].split(" ")
image_path = row[0]
image_path = os.path.join(data_path,image_path)
image = imread(image_path)
image = resize(image,(64,64))
X_valid.append(image)
label = row[1]
y_valid.append(label)
X_valid = np.array(X_valid)
y_valid = np.array(y_valid)
y_valid = np_utils.to_categorical(y_valid)
if test:
with open(test_path) as f:
reader = csv.reader(f)
for row in reader:
row = row[0].split(" ")
image_path = row[0]
image_path = os.path.join(data_path,image_path)
image = imread(image_path)
image = resize(image,(64,64))
X_test.append(image)
label = row[1]
y_test.append(label)
X_test = np.array(X_test)
y_test = np.array(y_test)
y_test = np_utils.to_categorical(y_test)
count = sum([train,valid,test])
if count==0
return X_train, y_train, X_valid, y_valid, X_test, y_test
X_train, y_train, X_valid, y_valid, X_test, y_test = load_data()
# -
# !ls /home/dl-box/.keras/datasets/yeastdata
| examples/.ipynb_checkpoints/deepyeast-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from minio import Minio
# +
data_lake_server= '172.17.0.2:9000'
data_lake_server= '127.0.0.1:9000'
data_lake_login= 'miniouser'
data_lake_password= '<PASSWORD>'
client = Minio(
endpoint= data_lake_server,
access_key= data_lake_login,
secret_key= data_lake_password,
secure=False
)
# -
client.fget_object(
bucket_name= 'landing',
object_name= 'olist_order_items_dataset.csv',
file_path= 'tmp/olist_order_items_dataset.csv'
)
temp_df = pd.read_csv('tmp/olist_order_items_dataset.csv')
temp_df.to_parquet('tmp/olist_order_items_dataset.parquet')
client.fput_object(
bucket_name= 'processing',
object_name= 'olist_order_items_dataset.pa',
file_path= 'tmp/olist_order_items_dataset.parquet')
# +
import os
import glob
files_remove = glob.glob('tmp/*')
for f in files_remove:
os.remove(f)
# -
| notebooks/Criando ETL - Landing para Processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
# import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import torch
import torch.nn as nn
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms as T
import torchvision
import torch.nn.functional as F
from torch.autograd import Variable
from PIL import Image
import cv2
# import albumentations as A
import time
import os
from os.path import join
from tqdm.notebook import tqdm
# import segmentation_models_pytorch as smp
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# device="cpu"
print(torch.cuda.get_device_name(torch.cuda.current_device()))
TEST_SPLIT = 0.15
PIN_MEMORY = True if device == "cuda" else False
# +
# for windows vs linux reasons, otherwise it wont work '/' vs '\'
# just define the data folder for your system and the rest works
DATA_FOLDER = "/home/malika/Documents/Bonn_Stuff/DLRV/Project/dope-drone-desegmentation-dlrv/code"
IMAGE_PATH = join(DATA_FOLDER, 'data', 'images')
MASK_PATH = join(DATA_FOLDER, 'data', 'masks')
print(IMAGE_PATH)
print(MASK_PATH)
print(f"image folders exist: {os.path.isdir(IMAGE_PATH)}")
print(f"mask folders exist: {os.path.isdir(MASK_PATH)}")
# +
# define the number of channels in the input, number of classes,
# and number of levels in the U-Net model
NUM_CHANNELS = 3
NUM_CLASSES = 1
NUM_LEVELS = 3
# initialize learning rate, number of epochs to train for, and the
# batch size
INIT_LR = 0.001
NUM_EPOCHS = 100
BATCH_SIZE = 128
# define the input image dimensions
INPUT_IMAGE_WIDTH = 256
INPUT_IMAGE_HEIGHT = 256
# define threshold to filter weak predictions
THRESHOLD = 0.5
# define the path to the base output directory
BASE_OUTPUT = "output"
# define the path to the output serialized model, model training
# plot, and testing image paths
MODEL_PATH = os.path.join(BASE_OUTPUT, "unet_tgs_salt.pth")
PLOT_PATH = os.path.sep.join([BASE_OUTPUT, "plot.png"])
TEST_PATHS = os.path.sep.join([BASE_OUTPUT, "test_paths.txt"])
# +
# TODO into own file (dataset)
class SegmentationDataset(Dataset):
def __init__(self, imagePaths, maskPaths, transforms):
# store the image and mask filepaths, and augmentation
# transforms
self.imagePaths = imagePaths
self.maskPaths = maskPaths
self.transforms = transforms
def __len__(self):
# return the number of total samples contained in the dataset
return len(self.imagePaths)
def __getitem__(self, idx):
# grab the image path from the current index
imagePath = self.imagePaths[idx]
# load the image from disk, swap its channels from BGR to RGB,
# and read the associated mask from disk in grayscale mode
image = cv2.imread(imagePath)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
mask = cv2.imread(self.maskPaths[idx], 0)
# check to see if we are applying any transformations
if self.transforms is not None:
# apply the transformations to both image and its mask
image = self.transforms(image)
mask = self.transforms(mask)
# return a tuple of the image and its mask
return (image, mask)
# +
# TODO into own file (model)
from torch.nn import ConvTranspose2d
from torch.nn import Conv2d
from torch.nn import MaxPool2d
from torch.nn import Module
from torch.nn import ModuleList
from torch.nn import ReLU
from torchvision.transforms import CenterCrop
from torch.nn import functional as F
import torch
class Block(Module):
def __init__(self, inChannels, outChannels):
super().__init__()
# store the convolution and RELU layers
self.conv1 = Conv2d(inChannels, outChannels, 3)
self.relu = ReLU()
self.conv2 = Conv2d(outChannels, outChannels, 3)
def forward(self, x):
# apply CONV => RELU => CONV block to the inputs and return it
return self.conv2(self.relu(self.conv1(x)))
class Encoder(Module):
def __init__(self, channels=(3, 16, 32, 64)):
super().__init__()
# store the encoder blocks and maxpooling layer
self.encBlocks = ModuleList(
[Block(channels[i], channels[i + 1])
for i in range(len(channels) - 1)])
self.pool = MaxPool2d(2)
def forward(self, x):
# initialize an empty list to store the intermediate outputs
blockOutputs = []
# loop through the encoder blocks
for block in self.encBlocks:
# pass the inputs through the current encoder block, store
# the outputs, and then apply maxpooling on the output
x = block(x)
blockOutputs.append(x)
x = self.pool(x)
# return the list containing the intermediate outputs
return blockOutputs
class Decoder(Module):
def __init__(self, channels=(64, 32, 16)):
super().__init__()
# initialize the number of channels, upsampler blocks, and
# decoder blocks
self.channels = channels
self.upconvs = ModuleList(
[ConvTranspose2d(channels[i], channels[i + 1], 2, 2)
for i in range(len(channels) - 1)])
self.dec_blocks = ModuleList(
[Block(channels[i], channels[i + 1])
for i in range(len(channels) - 1)])
def forward(self, x, encFeatures):
# loop through the number of channels
for i in range(len(self.channels) - 1):
# pass the inputs through the upsampler blocks
x = self.upconvs[i](x)
# crop the current features from the encoder blocks,
# concatenate them with the current upsampled features,
# and pass the concatenated output through the current
# decoder block
encFeat = self.crop(encFeatures[i], x)
x = torch.cat([x, encFeat], dim=1)
x = self.dec_blocks[i](x)
# return the final decoder output
return x
def crop(self, encFeatures, x):
# grab the dimensions of the inputs, and crop the encoder
# features to match the dimensions
(_, _, H, W) = x.shape
encFeatures = CenterCrop([H, W])(encFeatures)
# return the cropped features
return encFeatures
class UNet(Module):
def __init__(self, encChannels=(3, 16, 32, 64),
decChannels=(64, 32, 16),
nbClasses=1, retainDim=True,
outSize=(INPUT_IMAGE_HEIGHT, INPUT_IMAGE_WIDTH)):
super().__init__()
# initialize the encoder and decoder
self.encoder = Encoder(encChannels)
self.decoder = Decoder(decChannels)
# initialize the regression head and store the class variables
self.head = Conv2d(decChannels[-1], nbClasses, 1)
self.retainDim = retainDim
self.outSize = outSize
def forward(self, x):
# grab the features from the encoder
encFeatures = self.encoder(x)
# pass the encoder features through decoder making sure that
# their dimensions are suited for concatenation
decFeatures = self.decoder(encFeatures[::-1][0],
encFeatures[::-1][1:])
# pass the decoder features through the regression head to
# obtain the segmentation mask
map = self.head(decFeatures)
# check to see if we are retaining the original output
# dimensions and if so, then resize the output to match them
if self.retainDim:
map = F.interpolate(map, self.outSize)
# return the segmentation map
return map
# +
# TODO into file MAIN
# USAGE
# python train.py
# import the necessary packages
# from pyimagesearch.dataset import SegmentationDataset
# from pyimagesearch.model import UNet
# from pyimagesearch import config
from torch.nn import BCEWithLogitsLoss
from torch.optim import Adam
from torch.utils.data import DataLoader
from sklearn.model_selection import train_test_split
from torchvision import transforms
from imutils import paths
from tqdm import tqdm
import matplotlib.pyplot as plt
import torch
import time
import os
# load the image and mask filepaths in a sorted manner
imagePaths = sorted(list(paths.list_images(IMAGE_PATH)))
maskPaths = sorted(list(paths.list_images(MASK_PATH)))
# partition the data into training and testing splits using 85% of
# the data for training and the remaining 15% for testing
split = train_test_split(imagePaths, maskPaths,
test_size=TEST_SPLIT, random_state=42)
# unpack the data split
(trainImages, testImages) = split[:2]
(trainMasks, testMasks) = split[2:]
# write the testing image paths to disk so that we can use then
# when evaluating/testing our model
print("[INFO] saving testing image paths...")
f = open(TEST_PATHS, "w")
f.write("\n".join(testImages))
f.close()
# define transformations
transforms = transforms.Compose([transforms.ToPILImage(),
transforms.Resize((INPUT_IMAGE_HEIGHT,
INPUT_IMAGE_WIDTH)),
transforms.ToTensor()])
# create the train and test datasets
trainDS = SegmentationDataset(imagePaths=trainImages, maskPaths=trainMasks,
transforms=transforms)
testDS = SegmentationDataset(imagePaths=testImages, maskPaths=testMasks,
transforms=transforms)
print(f"[INFO] found {len(trainDS)} examples in the training set...")
print(f"[INFO] found {len(testDS)} examples in the test set...")
# create the training and test data loaders
# trainLoader = DataLoader(trainDS, shuffle=True,
# batch_size=BATCH_SIZE, pin_memory=PIN_MEMORY,
# num_workers=os.cpu_count())
# testLoader = DataLoader(testDS, shuffle=False,
# batch_size=BATCH_SIZE, pin_memory=PIN_MEMORY,
# num_workers=os.cpu_count())
trainLoader = DataLoader(trainDS, shuffle=True,
batch_size=BATCH_SIZE, pin_memory=PIN_MEMORY)
testLoader = DataLoader(testDS, shuffle=False,
batch_size=BATCH_SIZE, pin_memory=PIN_MEMORY)
# initialize our UNet model
unet = UNet().to(device)
# initialize loss function and optimizer
lossFunc = BCEWithLogitsLoss()
opt = Adam(unet.parameters(), lr=INIT_LR)
# calculate steps per epoch for training and test set
trainSteps = len(trainDS) // BATCH_SIZE
testSteps = len(testDS) // BATCH_SIZE
# initialize a dictionary to store training history
H = {"train_loss": [], "test_loss": []}
# loop over epochs
print("[INFO] training the network...")
startTime = time.time()
for e in tqdm(range(NUM_EPOCHS)):
# set the model in training mode
unet.train()
# initialize the total training and validation loss
totalTrainLoss = 0
totalTestLoss = 0
# loop over the training set
for (i, (x, y)) in enumerate(trainLoader):
# send the input to the device
(x, y) = (x.to(device), y.to(device))
# perform a forward pass and calculate the training loss
pred = unet(x)
loss = lossFunc(pred, y)
# first, zero out any previously accumulated gradients, then
# perform backpropagation, and then update model parameters
opt.zero_grad()
loss.backward()
opt.step()
# add the loss to the total training loss so far
totalTrainLoss += loss
# switch off autograd
with torch.no_grad():
# set the model in evaluation mode
unet.eval()
# loop over the validation set
for (x, y) in testLoader:
# send the input to the device
(x, y) = (x.to(device), y.to(device))
# make the predictions and calculate the validation loss
pred = unet(x)
totalTestLoss += lossFunc(pred, y)
# calculate the average training and validation loss
avgTrainLoss = totalTrainLoss / trainSteps
avgTestLoss = totalTestLoss / testSteps
# update our training history
H["train_loss"].append(avgTrainLoss.cpu().detach().numpy())
H["test_loss"].append(avgTestLoss.cpu().detach().numpy())
# print the model training and validation information
print("[INFO] EPOCH: {}/{}".format(e + 1, NUM_EPOCHS))
print("Train loss: {:.6f}, Test loss: {:.4f}".format(
avgTrainLoss, avgTestLoss))
# display the total time needed to perform the training
endTime = time.time()
print("[INFO] total time taken to train the model: {:.2f}s".format(
endTime - startTime))
# -
# plot the training loss
plt.style.use("ggplot")
plt.figure()
plt.plot(H["train_loss"], label="train_loss")
plt.plot(H["test_loss"], label="test_loss")
plt.title("Training Loss on Dataset")
plt.xlabel("Epoch #")
plt.ylabel("Loss")
plt.legend(loc="lower left")
plt.savefig(PLOT_PATH)
# serialize the model to disk
torch.save(unet, MODEL_PATH)
# +
# USAGE
# python predict.py
# import the necessary packages
import matplotlib.pyplot as plt
import numpy as np
import torch
import cv2
import os
def prepare_plot(origImage, origMask, predMask):
# initialize our figure
figure, ax = plt.subplots(nrows=1, ncols=3, figsize=(10, 10))
# plot the original image, its mask, and the predicted mask
ax[0].imshow(origImage)
ax[1].imshow(origMask)
ax[2].imshow(predMask)
# set the titles of the subplots
ax[0].set_title("Image")
ax[1].set_title("Original Mask")
ax[2].set_title("Predicted Mask")
# set the layout of the figure and display it
figure.tight_layout()
figure.show()
def make_predictions(model, imagePath):
# set model to evaluation mode
model.eval()
# turn off gradient tracking
with torch.no_grad():
# load the image from disk, swap its color channels, cast it
# to float data type, and scale its pixel values
image = cv2.imread(imagePath)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = image.astype("float32") / 255.0
# resize the image and make a copy of it for visualization
image = cv2.resize(image, (128, 128))
orig = image.copy()
# find the filename and generate the path to ground truth
# mask
filename = imagePath.split(os.path.sep)[-1]
groundTruthPath = os.path.join(MASK_PATH, filename).replace("sat", "mask")
print(groundTruthPath)
# load the ground-truth segmentation mask in grayscale mode
# and resize it
gtMask = cv2.imread(groundTruthPath, 0)
gtMask = cv2.resize(gtMask, (INPUT_IMAGE_HEIGHT,
INPUT_IMAGE_HEIGHT))
# make the channel axis to be the leading one, add a batch
# dimension, create a PyTorch tensor, and flash it to the
# current device
image = np.transpose(image, (2, 0, 1))
image = np.expand_dims(image, 0)
image = torch.from_numpy(image).to(device)
# make the prediction, pass the results through the sigmoid
# function, and convert the result to a NumPy array
predMask = model(image).squeeze()
predMask = torch.sigmoid(predMask)
predMask = predMask.cpu().numpy()
# filter out the weak predictions and convert them to integers
predMask = (predMask > THRESHOLD) * 255
predMask = predMask.astype(np.uint8)
# prepare a plot for visualization
prepare_plot(orig, gtMask, predMask)
# load the image paths in our testing file and randomly select 10
# image paths
print("[INFO] loading up test image paths...")
imagePaths = open(TEST_PATHS).read().strip().split("\n")
imagePaths = np.random.choice(imagePaths, size=10)
# load our model from disk and flash it to the current device
print("[INFO] load up model...")
unet = torch.load(MODEL_PATH).to(device)
# iterate over the randomly selected test image paths
for path in imagePaths:
# make predictions and visualize the results
make_predictions(unet, path)
# -
| code/forest_segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Enplore Python training introduction and making of a dataframe point
#
# ### Why python
# [Why Python](http://www.data-analysis-in-python.org/why_python.html)
#
# [Why notebook](https://unidata.github.io/online-python-training/introduction.html)
#
# [Data Science Cookiecutter](http://drivendata.github.io/cookiecutter-data-science/)
# +
# debugging / printing values
csv_file_test_variable = 'mydata.csv'
print('Here is a good debugging or checking through the program for {}'.format(csv_file_test_variable))
# practical checking
one_more_variable = 1
print('What is one_more_variable {} lets also check for my csv_file {}'.format(csv_file_test_variable,one_more_variable))
# -
# #### Example of Kaggle data
# [Kaggle flight-delays](https://www.kaggle.com/usdot/flight-delays)
#
# Download the dataset at https://www.kaggle.com/usdot/flight-delays/data
# +
# read of the secbysec data
filepath = '../data/flights.csv.zip'
import collections
import gzip
import pandas as pd
flights = pd.read_csv(filepath,delimiter=',',compression='zip')
# -
# ##### Installation
# Windows - miniconda
# Conda - https://conda.io/miniconda.html
#
# ##### Preparation
# ```
# $ conda install numpy
# $ conda install ipython
# $ conda install jupyter
# ```
# ## Further Reading:
# [Idiomatic Python](https://www.amazon.com/Writing-Idiomatic-Python-Jeff-Knupp/dp/1482374811) - gentle introduction to good python syntax.
# Next step
# [Python for Data Analysis(https://www.amazon.com/Python-Data-Analysis-Wrangling-IPython/dp/1449319793)
| notebooks/Introduction to Reading file with python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Run many Batch Normalization experiments using Cloud using ML Engine
# +
# change these to try this notebook out
BUCKET = 'crawles-sandbox' # change this to your GCP bucket
PROJECT = 'crawles-sandbox' # change this to your GCP project
REGION = 'us-central1'
# Import os environment variables
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
# -
# Let’s test how Batch Normalization impacts models of varying depths. We can launch many experiments in parallel using Google Cloud ML Engine. We will fire off 14 jobs with varying hyperparameters:
#
# * With and without Batch Normalization
# * Varying model depths from 1 hidden layer to 7 hidden layers
#
# We use the [tf.estimator](https://www.tensorflow.org/api_docs/python/tf/estimator/Estimator) API to build a model and deploy it using [Cloud ML Engine](https://cloud.google.com/ml-engine/docs/technical-overview).
# !ls mnist_classifier/
# !ls mnist_classifier/trainer/
# + language="bash"
# submitMLEngineJob() {
# gcloud ml-engine jobs submit training $JOBNAME \
# --package-path=$(pwd)/mnist_classifier/trainer \
# --module-name trainer.task \
# --region $REGION \
# --staging-bucket=gs://$BUCKET \
# --scale-tier=BASIC \
# --runtime-version=1.4 \
# -- \
# --outdir $OUTDIR \
# --hidden_units $net \
# --num_steps 10 \
# $batchNorm
# }
#
# # submit for different layer sizes
# export PYTHONPATH=${PYTHONPATH}:${PWD}/mnist_classifier
# for batchNorm in '' '--use_batch_normalization'
# do
# net=''
# for layer in 500 400 300 200 100 50 25;
# do
# net=$net$layer
# netname=${net//,/_}${batchNorm/--use_batch_normalization/_bn}
# echo $netname
# JOBNAME=mnist$netname_$(date -u +%y%m%d_%H%M%S)
# OUTDIR=gs://${BUCKET}/mnist_models/mnist_model$netname/trained_model
# echo $OUTDIR $REGION $JOBNAME
# gsutil -m rm -rf $OUTDIR
# submitMLEngineJob
# net=$net,
# done
# done
# -
# Copyright 2018 Google Inc. All Rights Reserved.
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
| blogs/batch_normalization/estimator_batch_normalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import torch
from torch.autograd import Variable
import numpy as np
def rmse(y, y_hat):
"""
Compute root mean squared error
@param y: 입력 값
@param y_hat: 평균 값
return이 tensor로 되는데 결국 vector로 리턴되는 걸 의미
"""
return torch.sqrt(torch.mean((y - y_hat).pow(2).sum()))
def forward(x, e):
"""Forward pass for our function"""
return x.pow(e.repeat(x.size(0)))
# Let's define some settings
n = 100 # number of examples
learning_rate = 5e-6
loss_history = []
exp_history = []
# Model definition
x = Variable(torch.rand(n) * 10, requires_grad=False)
# Model parameter and it's true value
exp = Variable(torch.FloatTensor([2.0]), requires_grad=False)
exp_hat = Variable(torch.FloatTensor([4]), requires_grad=True)
y = forward(x, exp)
# + pycharm={"name": "#%%\n"}
# Training loop
for i in range(0, 200):
print("Iteration %d" % i)
# Compute current estimate
y_hat = forward(x, exp_hat)
# Calculate loss function
loss = rmse(y, y_hat)
# Do some recordings for plots
# PyTorch 버전 변경에 따라 Tensor object의 data는 아래와 같이 참조하는 것으로 변경
loss_history.append(loss.data)
exp_history.append(y_hat.data)
print(loss_history, exp_history)
# Compute gradients
loss.back11ward()
print("loss = %s" % loss.data)
print("exp = %s" % exp_hat.data)
# Update model parameters
exp_hat.data -= learning_rate * exp_hat.grad.data
exp_hat.grad.data.zero_()
# + pycharm={"name": "#%%\n"}
| study/pytorchExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Now You Code 3: Limerick Generator
#
# We will write code in this example to create the first two lines of a [Limerick](https://en.wikipedia.org/wiki/Limerick_(poetry)). We're going to keep it really simple, and ask for 4 inputs:
# - a woman's name
# - a place
# - an item
# - a material
#
# The place and material must rhyme. The Python program will then output the first two lines of the limerick, substituting the values you've entered.
#
# **Example 1:**
#
# INPUT:
#
# - Enter a woman's name: Jane
# - Enter a place: New York
# - Enter an item: hat
# - Enter a material, which rhymes with 'New York': cork.
#
# OUTPUT:
# I once knew Jane from New York. Her hat was constructed of cork.
#
# **Example 2:**
#
# INPUT:
#
# - Enter a woman's name: Agatha
# - Enter a place: my car
# - Enter an item: bike
# - Enter a material, which rhymes with 'my car': tar
#
# OUTPUT:
# I once knew Agatha from my car. Her bike was constructed of tar.
#
# ## Step 1: Problem Analysis
#
# Inputs:
#
# Outputs:
#
# Algorithm (Steps in Program):
# +
# Step 2: Write code here
woman = input("Enter a woman's name: ")
place = input("Enter a place: ")
item = input("Enter an item: ")
material = input("Enter a material, which rhymes with "+ place +":")
print("I once knew",woman,"from my",place,". Her",item,"was constructed of",material,".")
# -
# ## Step 3: Questions
#
# 1. What happens when neglect to follow the instructions and enter any inputs we desire? Does the code still run? Why?
# a. The code still runs because there are no specific rules besides entering an input for an output.
# 2. What type of error occurs when the program runs but does not handle bad input?
# a. Logical Error
# 3. Is there anything you can do in code to correct this type of error? Why or why not?
# a. Maybe add some rules or restricitions so the output must be specific.
# ## Reminder of Evaluation Criteria
#
# 1. What the problem attempted (analysis, code, and answered questions) ?
# 2. What the problem analysis thought out? (does the program match the plan?)
# 3. Does the code execute without syntax error?
# 4. Does the code solve the intended problem?
# 5. Is the code well written? (easy to understand, modular, and self-documenting, handles errors)
#
| content/lessons/02/Now-You-Code/NYC3-Limerick-Generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# TSG029 - Find dumps in the cluster
# ==================================
#
# Description
# -----------
#
# Look for coredumps and minidumps from processes like SQL Server or
# controller in a big data cluster.
#
# Steps
# -----
#
# ### Instantiate Kubernetes client
# + tags=["hide_input"]
# Instantiate the Python Kubernetes client into 'api' variable
import os
try:
from kubernetes import client, config
from kubernetes.stream import stream
if "KUBERNETES_SERVICE_PORT" in os.environ and "KUBERNETES_SERVICE_HOST" in os.environ:
config.load_incluster_config()
else:
try:
config.load_kube_config()
except:
display(Markdown(f'HINT: Use [TSG118 - Configure Kubernetes config](../repair/tsg118-configure-kube-config.ipynb) to resolve this issue.'))
raise
api = client.CoreV1Api()
print('Kubernetes client instantiated')
except ImportError:
from IPython.display import Markdown
display(Markdown(f'HINT: Use [SOP059 - Install Kubernetes Python module](../install/sop059-install-kubernetes-module.ipynb) to resolve this issue.'))
raise
# -
# ### Get the namespace for the big data cluster
#
# Get the namespace of the Big Data Cluster from the Kuberenetes API.
#
# **NOTE:**
#
# If there is more than one Big Data Cluster in the target Kubernetes
# cluster, then either:
#
# - set \[0\] to the correct value for the big data cluster.
# - set the environment variable AZDATA\_NAMESPACE, before starting
# Azure Data Studio.
# + tags=["hide_input"]
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = api.list_namespace(label_selector='MSSQL_CLUSTER').items[0].metadata.name
except IndexError:
from IPython.display import Markdown
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print('The kubernetes namespace for your big data cluster is: ' + namespace)
# -
# ### Get all relevant pods
# +
pod_list = api.list_namespaced_pod(namespace, label_selector='app in (compute-0, data-0, storage-0, master, controller, controldb)', field_selector='status.phase==Running')
pod_names = [pod.metadata.name for pod in pod_list.items]
print('Scanning pods: ' + ', '.join(pod_names))
command = 'find /var/opt /var/log | grep -E "core\\.sqlservr|core\\.controller|SQLD|\\.mdmp$|\\.dmp$|\\.gdmp$"'
all_dumps = ''
for name in pod_names:
print('Searching pod: ' + name)
container = 'mssql-server'
if 'control-' in name:
container = 'controller'
try:
dumps=stream(api.connect_get_namespaced_pod_exec, name, namespace, command=['/bin/sh', '-c', command], container=container, stderr=True, stdout=True)
except Exception as e:
print(f'Unable to connect to pod: {name} due to {str(e.__class__)}. Skipping dump check for this pod...')
else:
if dumps:
all_dumps += '*Pod: ' + name + '*\n'
all_dumps += dumps + '\n'
# -
# ### Validate
#
# Validate no dump files were found.
# +
if len(all_dumps) > 0:
raise SystemExit('FAIL - dump files found:\n' + all_dumps)
print('SUCCESS - no dump files were found.')
# -
print('Notebook execution complete.')
| Big-Data-Clusters/CU3/Public/content/diagnose/tsg029-find-dumps-in-the-cluster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Creating the Sentence Encoder Indices
# %cd ../../../../
from gamechangerml.src.search.sent_transformer.model import SentenceEncoder
encoder_model = "sentence-transformers/msmarco-distilbert-base-v2"
corpus_path = "./sample_corpus"
index_path = "./sample_index"
# +
# Calling the class
encoder = SentenceEncoder(
encoder_model = encoder_model
)
# Create the document indices
encoder.index_documents(
corpus_path = corpus_path,
index_path = index_path
)
# -
# ## Loading the Sentence Searcher Class
from gamechangerml.src.search.sent_transformer.model import SentenceSearcher
import pandas as pd
encoder_model = "sentence-transformers/msmarco-distilbert-base-v2"
sim_model = "valhalla/distilbart-mnli-12-3"
index_path = "./sample_index"
# Calling the search class
searcher = SentenceSearcher(
index_path = index_path,
encoder_model = encoder_model,
sim_model = sim_model
)
# Printing out the results
results = searcher.search("Under Secretary of Defense for Intelligence")
pd.set_option('display.max_colwidth', 0)
pd.DataFrame(results, columns = ["Score", "Paragraph ID", "Text"])
| gamechangerml/experimental/notebooks/sentence-transformer/Sample_Modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# +
import itertools
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from matplotlib import cm
import seaborn as sns
from scipy import stats
LABEL_ALIAS = {
"lit": "In literature?",
"ge": "Germanosilicate composition?",
"zeo": "Zeolite composition?",
"zty": "Zeotype composition?",
}
df = pd.read_csv("../data/binding.csv", index_col=0)
# +
# idxmin = df.groupby(['Zeolite', 'InchiKey']).idxmin()
# df.iloc[0]["Competition (SiO2)"]
# pd.unique(df["Zeolite"]) # -> 209 zeolites...
# pd.unique(df["SMILES"]) # -> 1194 smiles
import pdb
def construct_column_count_histogram(col: str, df: pd.DataFrame, raw=False, n_bins=20):
if raw:
histogram_by_count = df[col]
else:
col_vals = pd.unique(df[col])
histogram_by_count = {}
for val in col_vals:
count = len(df.loc[df[col] == val])
histogram_by_count[count] = histogram_by_count.get(count, 0) + 1
plt.hist(histogram_by_count, bins=n_bins, density=True)
# plt.ylabel("# unique " + col)
# plt.title("Histogram of " + col + " binned by templating energies")
# plt.xlabel("# templating energies")
plt.ylabel("Density ")
plt.title("Templating energies distribution")
plt.xlabel("Templating energies")
print(
"total cells: ",
sum([count * instances for count, instances in histogram_by_count.items()]),
)
# construct_column_count_histogram("Zeolite", df)
# construct_column_count_histogram("Ligand formula", df)
construct_column_count_histogram("Templating", df, raw=True)
# np.std(df['Templating'].dropna())
# np.mean(df['Templating'].dropna())
# np.median(df['Templating'].dropna())
# np.var(df['Templating'].dropna())
# +
import pdb
import os
import pathlib
def save_matrix(matrix, file_name):
file = os.path.abspath("")
pdb.set_trace()
dir_main = pathlib.Path(file).parent.absolute()
savepath = os.path.join(dir_main, file_name)
# if os.path.exists(savepath):
# overwrite = input(f"A file already exists at path {savepath}, do you want to overwrite? (Y/N): ")
matrix.to_pickle(savepath)
def create_skinny_matrix_nonbinding(df, num_zeolites=30, num_osdas=100):
# skinny_matrix = df.pivot_table(
# index=["SMILES", "Zeolite"],
# values=[
# "Templating",
# "Binding (SiO2)",
# "Binding (OSDA)",
# "Directivity (SiO2)",
# "Competition (SiO2)",
# "Competition (OSDA)",
# ],
# )
matrix = df.pivot(index="SMILES", columns="Zeolite", values="Templating")
matrix = matrix.fillna(0)
matrix[matrix != 0] = 1
matrix = matrix.iloc[:num_osdas, :num_zeolites]
matrix = matrix.reset_index()
melted_matrix = pd.melt(matrix, id_vars='SMILES', value_vars=list(matrix.columns[1:]))
melted_matrix_reindexed = melted_matrix.set_index(['SMILES', 'Zeolite'])
return melted_matrix_reindexed
matrix = df.pivot(index="SMILES", columns="Zeolite", values="Templating")
skinny_matrix_nonbinding = create_skinny_matrix_nonbinding(df)
save_matrix(skinny_matrix_nonbinding, 'zeoliteOSDANonbindingIndexedMatrix.pkl')
save_matrix(df, "completeZeoliteData.pkl")
# the values in this matrix that are nan are not actually missing.
matrix = matrix.fillna(30)
# matrix = matrix.fillna(0)
# matrix[matrix != 0] = 1
pdb.set_trace()
# pdb.set_trace()
save_matrix(matrix, "zeoliteNonbindingTensor.pkl")
print("total not na cells:", matrix.notna().sum().sum()) # 17587 what gives?
M = matrix.to_numpy()
def create_binary_mask(m, mask_nans=True, mask_zeros=False):
def cast_bool_to_int(m):
return np.array([1.0 if c else 0.0 for c in m])
def create_bool_mask(m, mask_nans, mask_zeros):
nan_mask = (
np.invert(np.isnan(m)) if mask_nans else np.full(m.shape, True, dtype=bool)
)
zero_mask = m == 1 if mask_zeros else np.full(m.shape, True, dtype=bool)
return np.logical_and(nan_mask, zero_mask,)
return np.array(
[cast_bool_to_int(r) for r in create_bool_mask(m, mask_nans, mask_zeros)]
)
def invert_binary_mask(m):
return np.logical_not(m).astype(int)
mask = create_binary_mask(M)
def plot_top_k_singular_values(var_explained, top_k=70):
fig = plt.figure()
ax = fig.add_axes([0, 0, 1, 1])
ax.bar(range(top_k), var_explained[0:top_k])
plt.xlabel("Singular Vector", fontsize=16)
plt.ylabel("Proportion of Variance", fontsize=16)
plt.tight_layout()
plt.savefig("svd_scree_plot.png", dpi=150, figsize=(8, 6))
plt.show()
M = np.nan_to_num(M)
U, s, V = np.linalg.svd(M)
var_explained = np.round(s ** 2 / np.sum(s ** 2), decimals=3)
plot_top_k_singular_values(var_explained)
def plot_matrix(M, mask, file_name, vmin=16, vmax=23):
fig, ax = plt.subplots()
cmap = mpl.cm.get_cmap()
cmap.set_bad(color="white")
if mask is not None:
inverted_mask = invert_binary_mask(mask)
masked_M = np.ma.masked_where(inverted_mask, M)
else:
masked_M = M
im = ax.imshow(masked_M, interpolation="nearest", cmap=cmap, vmin=vmin, vmax=vmax)
fig.colorbar(im)
fig.savefig(file_name + ".png", dpi=150)
def calc_MSE(M, M_star, mask):
total_error = 0.0
total_count = 0.0
for x, y in np.ndindex(mask.shape):
if mask[x, y] == 0:
continue
total_error += (M[x, y] - M_star[x, y]) ** 2
total_count += 1
return total_error / total_count
def reconstruct_M(U, s, V, num_components):
M_star = (
np.matrix(U[:, :num_components])
* np.diag(s[:num_components])
* np.matrix(V[:num_components, :])
)
return M_star
def run_SVD(M, mask, file_name, num_components):
M = np.nan_to_num(M)
U, s, V = np.linalg.svd(M)
M_star = reconstruct_M(U, s, V, num_components)
plot_matrix(M_star, mask, file_name)
mse_k1 = calc_MSE(M, M_star, mask)
print(file_name, ": ", mse_k1)
plot_matrix(M, mask, "matrix")
run_SVD(
M=M, mask=mask, file_name="reconstructed_matrix_k1", num_components=1,
)
run_SVD(
M=M, mask=mask, file_name="reconstructed_matrix_k5", num_components=5,
)
run_SVD(
M=M, mask=mask, file_name="reconstructed_matrix_k70", num_components=70,
)
run_SVD(
M=M, mask=mask, file_name="reconstructed_matrix_k209", num_components=209,
)
# +
# Divide M into train and test...
import random
random.seed(10)
train_ratio = 0.5
train_mask = np.zeros(mask.shape)
for x, y in np.ndindex(mask.shape):
if mask[x, y]:
train_mask[x, y] = 1 if random.random() <= train_ratio else 0
test_mask = mask - train_mask
print("total #? ", np.count_nonzero(mask))
print("train #? ", np.count_nonzero(train_mask))
print("test #? ", np.count_nonzero(test_mask))
# +
from matrix_completion import (
nuclear_norm_solve,
svt_solve,
) # nuclear_norm_solve# svt_solve, calc_unobserved_rmse
# taken from https://pypi.org/project/matrix-completion/
# which -a pip
# /opt/anaconda3/bin/pip install matrix-completion
# /opt/anaconda3/bin/pip install cvxpy
# okay let's try svt_solve first...
# M = np.nan_to_num(M)
M_hat = nuclear_norm_solve(M, train_mask)
total_mse = calc_MSE(M, M_hat, mask)
train_mse = calc_MSE(M, M_hat, train_mask)
test_mse = calc_MSE(M, M_hat, test_mask)
print("total mse? ", total_mse, " train_mse? ", train_mse, " test_mse? ", test_mse)
plot_matrix(M_hat, train_mask, "nuclear_norm_matrix_with_mask")
plot_matrix(M_hat, None, "nuclear_norm_matrix_without_mask")
# +
total_mse = calc_MSE(M, M_hat, mask)
train_mse = calc_MSE(M, M_hat, train_mask)
test_mse = calc_MSE(M, M_hat, test_mask)
print("total mse? ", total_mse, " train_mse? ", train_mse, " test_mse? ", test_mse)
def plot_matrix(M, mask, file_name, vmin=16, vmax=23):
fig, ax = plt.subplots()
cmap = mpl.cm.get_cmap()
cmap.set_bad(color="white")
if mask is not None:
inverted_mask = invert_binary_mask(mask)
masked_M = np.ma.masked_where(inverted_mask, M)
else:
masked_M = M
im = ax.imshow(masked_M, interpolation="nearest", cmap=cmap, vmin=vmin, vmax=vmax)
fig.colorbar(im)
fig.savefig(file_name + ".png", dpi=150)
plot_matrix(M_hat, train_mask, "nuclear_norm_matrix_with_mask", vmin=16, vmax=23)
plot_matrix(M_hat, None, "nuclear_norm_matrix_without_mask", vmin=16, vmax=23)
# +
literature_matrix = df.pivot(index="SMILES", columns="Zeolite", values="In literature?")
print("total not na cells:", literature_matrix.notna().sum().sum())
lit_M = literature_matrix.to_numpy()
lit_mask = create_binary_mask(lit_M, True, True)
# Total 2102 non zero lit cells... very sparse very sparse.
print("#non zero:", np.count_nonzero(np.nan_to_num(lit_M)))
concatted_M = np.concatenate((M, lit_M), axis=1)
concatted_mask = np.concatenate((mask, lit_mask), axis=1)
plot_matrix(concatted_M, concatted_mask, "lit_matrix")
concatted_M_hat = nuclear_norm_solve(concatted_M, concatted_mask)
plot_matrix(concatted_M_hat, concatted_mask, "lit_matrix_computed", vmin=0, vmax=1)
total_mse = calc_MSE(concatted_M, concatted_M_hat, concatted_mask)
print("total mse: ", total_mse)
test_concatted_mask = np.concatenate((np.zeros(mask.shape), lit_mask), axis=1)
test_concatted_mse = calc_MSE(concatted_M, concatted_M_hat, test_concatted_mask)
print("test_concatted_mse: ", test_concatted_mse)
| code/matrix_completion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="7q9s7AAxgDGY"
# importing the requisite libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score, mean_squared_error
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="UyR0EdRHm3lb"
# ### Modeling non-linear relationship using Polynomial Regression
# + id="8KNPRGw2VYJ6" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="f54a23d0-d3af-4dac-fcd1-1f4f6a2ef1c8"
# Model to predict marks given the number of courses taken and the time the student gives to study
# on a daily basis.
data = pd.read_csv("AR - Examples - 1.6.csv")
data.head()
# + id="nQQC0bwpYH0n" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="a9db6163-f0b9-4770-aac6-dc305a89c7cd"
# Plotting a scatter plot
sns.scatterplot( data = data , x = 'number_courses' , y = 'Marks')
# + id="H-YVPqtbYH0r" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="ae370dc2-6fc4-43e9-f676-4c0bf2dbc5dd"
# Plotting a scatter plot
sns.scatterplot( data = data , x = 'time_study' , y = 'Marks')
# + id="wyzsOLxvYH0y"
# Splitting the dataset into X and y
X = data[['number_courses' , 'time_study']]
y = np.array(data['Marks']).reshape(-1,1)
# + id="aGX4nAAPYH00" colab={"base_uri": "https://localhost:8080/"} outputId="24eaadb7-9aca-47e6-aab1-b6ef00f942da"
# Building the regression model
reg = LinearRegression()
reg.fit(X,y)
# + id="IqDN8e84YH02"
# Predictions on the basis of the model
y_pred = reg.predict(X)
#y_pred
# + id="pegMvW_DYH04" colab={"base_uri": "https://localhost:8080/"} outputId="ee61d305-9db9-493d-bf4f-156cbf429f87"
# Find the value of r squared
r2_score(y , y_pred)
# + id="ekn8Kn8ZYH1D" colab={"base_uri": "https://localhost:8080/"} outputId="fe9cd9bf-750e-4228-8637-668fc76d3b11"
# Calculate beta0 and beta1.
print(reg.intercept_)
print(reg.coef_)
# + id="cdlVIYH1YH1F" colab={"base_uri": "https://localhost:8080/"} outputId="fd2f3ccd-f4bd-4226-f6d9-fedebb168cbb"
# Metrics to give an overall sense of error in the model
rss = np.sum(np.square(y - y_pred))
print(rss)
mse = mean_squared_error(y, y_pred)
print(mse)
rmse = mse**0.5
print(rmse)
# + id="J9PaDywRYH1H"
# Residual analysis
y_res = y - y_pred
#y_res
# + id="TLShNOYsM737" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="d0065de5-2fab-471d-d8e0-350812eae860"
data['res'] = y_res
plt.scatter( y_pred , data['res'])
plt.axhline(y=0, color='r', linestyle=':')
plt.xlabel("Predictions")
plt.ylabel("Residual")
plt.show()
# + [markdown] id="-BwrFMy5ekn0"
# Checking which variable is non-linearly related to the response value
# + id="EAT8NM-KYH1K" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="2345b753-27ff-4278-f4ca-4248f75bc5ee"
data['res'] = y_res
plt.scatter( data['number_courses'] , data['res'])
plt.axhline(y=0, color='r', linestyle=':')
plt.xlabel("Number courses")
plt.ylabel("Residual")
plt.show()
# + id="5Y2GAw74Z2yf" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="2635eda9-ecc4-46b3-bde2-3cb2969ccf8d"
plt.scatter( data['time_study'] , data['res'])
plt.axhline(y=0, color='r', linestyle=':')
plt.xlabel("Time study")
plt.ylabel("Residual")
plt.show()
# + id="SR569GE7YH1M" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="8b2d7f51-6357-4cb7-d02f-3d21b26e43e0"
# Distribution of errors
p = sns.distplot(y_res,kde=True)
p = plt.title('Normality of error terms/residuals')
plt.xlabel("Residuals")
plt.show()
# + id="jLUTX2Ezekn2"
# + id="fw5zjyUrekn2"
# + id="af-Dc2Nrekn2"
# + id="KB3eSTsgekn2"
# + id="NjvX_GhRekn3"
# + id="XlLLghKmekn3"
# + id="hBqRB8gCekn3"
# + id="p2OJkPrIekn3"
# + id="Rj0jIVH0ekn4"
# + id="pBLhdxX8ekn4"
# + id="422KhyOLekn4"
# + [markdown] id="2sJeRTkqvtW5"
# ### Fitting Polynomial Regression Model
# + id="1Y7IkfB4aGhC"
# Transforming the time_study variable
data['time_study_squared'] = data['time_study']*data['time_study']
# + id="dYSB-UWCQ2Jq" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="5dc2b64d-3e13-4c41-c143-025edefa0bc7"
plt.scatter( data['time_study_squared'] , data['Marks'])
plt.xlabel("Time study squared")
plt.ylabel("Marks")
plt.show()
# + id="Asbt6moioop2"
# Splitting the dataset into X and y
X = data[['number_courses' , 'time_study', 'time_study_squared']]
y = np.array(data['Marks']).reshape(-1,1)
# + id="aRbQ3STYoop6" colab={"base_uri": "https://localhost:8080/"} outputId="8fe8ee84-2e70-432d-fbb3-d8eb4a7a35d8"
# Building the regression model
reg = LinearRegression()
reg.fit(X,y)
# + id="iwlR0PG6oop9"
# Predictions on the basis of the model
y_pred = reg.predict(X)
# y_pred
# + id="bUGyZkBSooqA" colab={"base_uri": "https://localhost:8080/"} outputId="0a4bbd46-b22a-4c25-a0b8-56c3e7806fec"
# Find the value of r squared
r2_score(y , y_pred)
# + id="HgAjmrchooqC" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="4d10e682-7961-426b-8647-c5fdb10680fb"
plt.scatter( X['number_courses'] , y , color = 'red')
plt.scatter( X['number_courses'] , y_pred , color = 'blue' )
plt.xlabel("Number Courses")
plt.ylabel("Marks")
plt.show()
# + id="yFdPbj-EooqE" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="6c813a80-b936-46fd-8f14-39f0fe6b42b2"
plt.scatter( X['time_study_squared'] , y , color = 'red')
plt.scatter( X['time_study_squared'] , y_pred , color = 'blue' )
plt.xlabel("Time Study Squared")
plt.ylabel("Marks")
plt.show()
# + id="nr6m2q7PooqG" colab={"base_uri": "https://localhost:8080/"} outputId="f15e09e1-d7e3-425b-81b8-1de51931a714"
# Calculate beta0 and beta1.
print(reg.intercept_)
print(reg.coef_)
# + id="AgaZ_6gbooqI" colab={"base_uri": "https://localhost:8080/"} outputId="38837445-9091-455d-f012-8ccc069e81cb"
#Residual Sum of Squares = Mean_Squared_Error * Total number of datapoints
rss = np.sum(np.square(y - y_pred))
print(rss)
mse = mean_squared_error(y, y_pred)
print(mse)
rmse = mse**0.5
print(rmse)
# + id="jHrr08xdooqK"
# Residual analysis
y_res = y - y_pred
#y_res
# + id="qYGD-wiXekn9" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="e9be2d48-1a4c-4753-c198-e4c4db91ae31"
data['res'] = y_res
plt.scatter( y_pred , data['res'])
plt.axhline(y=0, color='r', linestyle=':')
plt.xlabel("Predictions")
plt.ylabel("Residual")
plt.show()
# + id="YH1-x6xlooqN" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="ce231022-0936-4fcc-c8d5-8acc879d4ec0"
data['res'] = y_res
plt.scatter( data['number_courses'] , data['res'])
plt.axhline(y=0, color='r', linestyle=':')
plt.xlabel("Number courses")
plt.ylabel("Residual")
plt.show()
# + id="XcE_5tXRooqP" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="c033484e-733d-4a36-f83c-cb2b3a8ee0ee"
plt.scatter( data['time_study_squared'] , data['res'])
plt.axhline(y=0, color='r', linestyle=':')
plt.xlabel("Time study Squared")
plt.ylabel("Residual")
plt.show()
# + id="806WZJMvooqT" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="4ebbc0dc-53e5-4e7b-ab2e-83bfc00f5142"
# Distribution of errors
p = sns.distplot(y_res,kde=True)
p = plt.title('Normality of error terms/residuals')
plt.xlabel("Residuals")
plt.show()
# + id="arw-Zo8mSRV7"
# + [markdown] id="AB-7WEzzmvET"
# ### Modeling non-linear relationships using data transformation
# + [markdown] id="8k50BFpznpkw"
# Here, we need to make a model which predicts how much distance is covered by a truck in a given time unit after a break is applied.
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="OCXjKgllP0Lk" outputId="ec02c9c4-fc3a-43ec-9917-da4123ed0627"
dist = pd.read_csv(r"AR - Examples - 1.5.csv")
dist.head()
# + id="OVCCqjyvQRRt" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="bfdea22a-fb30-4c76-9349-0d7a6368d99f"
# Plotting a scatter plot
sns.scatterplot( data = dist , x ='time' , y='distance')
plt.show()
# + id="J7ixDMYcQ0ZI"
# Splitting the dataset into X and y
X = np.array(dist['time']).reshape(-1,1)
y = np.array(dist['distance']).reshape(-1,1)
# + id="4qSfoX2TQbDE"
# Building the regression model
model = LinearRegression()
# + id="7zQZyMkYQxZd" colab={"base_uri": "https://localhost:8080/"} outputId="01b15e7d-b906-40b4-dbad-5709699c3a2d"
model.fit(X, y)
# + id="QYPbYCKfRVck"
# Predictions on the basis of the model
y_pred2 = model.predict(X)
# y_pred2
# + id="YOHRRYgYRnZh" colab={"base_uri": "https://localhost:8080/"} outputId="e8e8d8ca-a9ff-4ee3-b54d-630040f8a6cd"
# Find the value of r squared
r2_score(y, y_pred2)
# + id="FenOPM6eeknm" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="ada81499-0571-4b3a-a6f6-43b0d61768e8"
plt.scatter( X , y , color = 'blue')
plt.plot(X , y_pred2 , color = 'red' , linewidth = 3)
plt.xlabel("time")
plt.ylabel("distance")
plt.show()
# + id="Ed_rLssSU_AN" colab={"base_uri": "https://localhost:8080/"} outputId="bd0699f3-fa00-4cc7-842f-f98a38261ebb"
# Calculate beta0 and beta1.
print(model.intercept_)
print(model.coef_)
# + id="SdZ4Pg2qU_A0" colab={"base_uri": "https://localhost:8080/"} outputId="467137b7-61b8-4966-8329-0f540cf0c924"
# Metrics to give an overall sense of error in the model
rss = np.sum(np.square(y - y_pred2))
print(rss)
mse = mean_squared_error(y, y_pred2)
print(mse)
rmse = mse**0.5
print(rmse)
# + id="yjkyLZthRyrz"
#residual
residual = y - y_pred2
# + id="yq1zygHBR7uT" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="9c9d2044-56b2-486d-e707-d00bf3732b11"
# Scatter plot of the predicted values on the x-axis and the residuals on the y-axis
plt.scatter( y_pred2 , residual)
plt.axhline(y=0, color='r', linestyle=':')
plt.xlabel("Predicted Distance (metres)")
plt.ylabel("Residual")
plt.show()
# + id="kWRnv8RmU_A7" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="415729b2-b9b2-469d-a196-8affc9f9a285"
# Distribution of errors
p = sns.distplot(residual,kde=True)
p = plt.title('Normality of error terms/residuals')
plt.xlabel("Residual")
plt.show()
# + id="iZydkRm_GZTg"
# As we can see that the residuals do not fulfill the conditions for linear regression, Lets see if we can make some changes so that the residuals are normally distributed.
dist['time (seconds)(log)'] = np.log(dist['time'])
# + id="L5pRvbyzP56h" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="69fc8270-8e6f-4a5e-e22b-085390cfd76a"
# Plotting a scatter plot
sns.scatterplot( data = dist , x ='time (seconds)(log)' , y='distance')
plt.show()
# + id="eGaZ6aH2GeR5"
# Splitting the dataset into X and y
X = np.array(dist['time (seconds)(log)']).reshape(-1,1)
y = np.array(dist['distance']).reshape(-1,1)
# + id="F4CkavBbGeli" colab={"base_uri": "https://localhost:8080/"} outputId="c9427656-9181-4e1f-a2d9-f0e91a3a7f91"
# Building the regression model
model = LinearRegression()
model.fit(X, y)
# + id="w0tcA9ZYGe7l"
# Predictions on the basis of the model
y_pred2 = model.predict(X)
# y_pred2
# + id="mi0YdcvhGfKv" colab={"base_uri": "https://localhost:8080/"} outputId="a8986948-a05b-4a48-fa95-508292cd21cf"
# Find the value of r squared
r2_score(y, y_pred2)
# + id="co43cHt_eknr" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="96800ceb-6ae6-42bc-808e-d84e963e37ba"
plt.scatter( X , y , color = 'blue')
plt.plot(X , y_pred2 , color = 'red' , linewidth = 3)
plt.xlabel("time (seconds)(log)")
plt.ylabel("distance")
plt.show()
# + id="F5_SnhXPGgGR" colab={"base_uri": "https://localhost:8080/"} outputId="d4ce7f34-47f7-49ff-cfec-c68c84fc661a"
# Calculate beta0 and beta1.
print(model.intercept_)
print(model.coef_)
# + id="Awrqfc0TGt9H" colab={"base_uri": "https://localhost:8080/"} outputId="03c829c9-2cd8-423b-a934-0a17ed3989a9"
# Metrics to give an overall sense of error in the model
rss = np.sum(np.square(y - y_pred2))
print(rss)
mse = mean_squared_error(y, y_pred2)
print(mse)
rmse = mse**0.5
print(rmse)
# + id="o-eMLCGCGfi_"
#residual
residual = y - y_pred2
# + id="YxL2MJuLGf1r" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="cf9ebac6-2932-487e-e76d-3c7194cdd338"
plt.scatter( y_pred2 , residual)
plt.axhline(y=0, color='r', linestyle=':')
plt.xlabel("Predicted Distance (metres)")
plt.ylabel("Residual")
plt.show()
# + id="Mo-yITcvGt2s" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="e3081136-c170-47e4-d099-ad31cb8a1fbe"
# Distribution of errors
p = sns.distplot(residual,kde=True)
p = plt.title('Normality of error terms/residuals')
plt.xlabel("Residual")
plt.show()
| 8. Machine Learning-2/1. Advanced Regression/2 .Polynomial_Regression and Data Transformation/.ipynb_checkpoints/Polynomial_Regression_and_Data_Transformation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Amazon SageMaker Processing jobs
#
# With Amazon SageMaker Processing jobs, you can leverage a simplified, managed experience to run data pre- or post-processing and model evaluation workloads on the Amazon SageMaker platform.
#
# A processing job downloads input from Amazon Simple Storage Service (Amazon S3), then uploads outputs to Amazon S3 during or after the processing job.
#
# <img src="Processing-1.jpg">
#
# This notebook shows how you can:
#
# 1. Run a processing job to run a scikit-learn script that cleans, pre-processes, performs feature engineering, and splits the input data into train and test sets.
# 2. Run a training job on the pre-processed training data to train a model
# 3. Run a processing job on the pre-processed test data to evaluate the trained model's performance
# 4. Use your own custom container to run processing jobs with your own Python libraries and dependencies.
#
# The dataset used here is the [Census-Income KDD Dataset](https://archive.ics.uci.edu/ml/datasets/Census-Income+%28KDD%29). You select features from this dataset, clean the data, and turn the data into features that the training algorithm can use to train a binary classification model, and split the data into train and test sets. The task is to predict whether rows representing census responders have an income greater than `$50,000`, or less than `$50,000`. The dataset is heavily class imbalanced, with most records being labeled as earning less than `$50,000`. After training a logistic regression model, you evaluate the model against a hold-out test dataset, and save the classification evaluation metrics, including precision, recall, and F1 score for each label, and accuracy and ROC AUC for the model.
# ## Data pre-processing and feature engineering
# To run the scikit-learn preprocessing script as a processing job, create a `SKLearnProcessor`, which lets you run scripts inside of processing jobs using the scikit-learn image provided.
# +
import boto3
import sagemaker
from sagemaker import get_execution_role
from sagemaker.sklearn.processing import SKLearnProcessor
region = boto3.session.Session().region_name
role = get_execution_role()
sklearn_processor = SKLearnProcessor(framework_version='0.20.0',
role=role,
instance_type='ml.m5.xlarge',
instance_count=1)
# -
# Before introducing the script you use for data cleaning, pre-processing, and feature engineering, inspect the first 20 rows of the dataset. The target is predicting the `income` category. The features from the dataset you select are `age`, `education`, `major industry code`, `class of worker`, `num persons worked for employer`, `capital gains`, `capital losses`, and `dividends from stocks`.
# +
import pandas as pd
input_data = 's3://sagemaker-sample-data-{}/processing/census/census-income.csv'.format(region)
df = pd.read_csv(input_data, nrows=10)
df.head(n=10)
# -
# This notebook cell writes a file `preprocessing.py`, which contains the pre-processing script. You can update the script, and rerun this cell to overwrite `preprocessing.py`. You run this as a processing job in the next cell. In this script, you
#
# * Remove duplicates and rows with conflicting data
# * transform the target `income` column into a column containing two labels.
# * transform the `age` and `num persons worked for employer` numerical columns into categorical features by binning them
# * scale the continuous `capital gains`, `capital losses`, and `dividends from stocks` so they're suitable for training
# * encode the `education`, `major industry code`, `class of worker` so they're suitable for training
# * split the data into training and test datasets, and saves the training features and labels and test features and labels.
#
# Our training script will use the pre-processed training features and labels to train a model, and our model evaluation script will use the trained model and pre-processed test features and labels to evaluate the model.
# +
# %%writefile preprocessing.py
import argparse
import os
import warnings
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, OneHotEncoder, LabelBinarizer, KBinsDiscretizer
from sklearn.preprocessing import PolynomialFeatures
from sklearn.compose import make_column_transformer
from sklearn.exceptions import DataConversionWarning
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
columns = ['age', 'education', 'major industry code', 'class of worker', 'num persons worked for employer',
'capital gains', 'capital losses', 'dividends from stocks', 'income']
class_labels = [' - 50000.', ' 50000+.']
def print_shape(df):
negative_examples, positive_examples = np.bincount(df['income'])
print('Data shape: {}, {} positive examples, {} negative examples'.format(df.shape, positive_examples, negative_examples))
if __name__=='__main__':
parser = argparse.ArgumentParser()
parser.add_argument('--train-test-split-ratio', type=float, default=0.3)
args, _ = parser.parse_known_args()
print('Received arguments {}'.format(args))
input_data_path = os.path.join('/opt/ml/processing/input', 'census-income.csv')
print('Reading input data from {}'.format(input_data_path))
df = pd.read_csv(input_data_path)
df = pd.DataFrame(data=df, columns=columns)
df.dropna(inplace=True)
df.drop_duplicates(inplace=True)
df.replace(class_labels, [0, 1], inplace=True)
negative_examples, positive_examples = np.bincount(df['income'])
print('Data after cleaning: {}, {} positive examples, {} negative examples'.format(df.shape, positive_examples, negative_examples))
split_ratio = args.train_test_split_ratio
print('Splitting data into train and test sets with ratio {}'.format(split_ratio))
X_train, X_test, y_train, y_test = train_test_split(df.drop('income', axis=1), df['income'], test_size=split_ratio, random_state=0)
preprocess = make_column_transformer(
(['age', 'num persons worked for employer'], KBinsDiscretizer(encode='onehot-dense', n_bins=10)),
(['capital gains', 'capital losses', 'dividends from stocks'], StandardScaler()),
(['education', 'major industry code', 'class of worker'], OneHotEncoder(sparse=False))
)
print('Running preprocessing and feature engineering transformations')
train_features = preprocess.fit_transform(X_train)
test_features = preprocess.transform(X_test)
print('Train data shape after preprocessing: {}'.format(train_features.shape))
print('Test data shape after preprocessing: {}'.format(test_features.shape))
train_features_output_path = os.path.join('/opt/ml/processing/train', 'train_features.csv')
train_labels_output_path = os.path.join('/opt/ml/processing/train', 'train_labels.csv')
test_features_output_path = os.path.join('/opt/ml/processing/test', 'test_features.csv')
test_labels_output_path = os.path.join('/opt/ml/processing/test', 'test_labels.csv')
print('Saving training features to {}'.format(train_features_output_path))
pd.DataFrame(train_features).to_csv(train_features_output_path, header=False, index=False)
print('Saving test features to {}'.format(test_features_output_path))
pd.DataFrame(test_features).to_csv(test_features_output_path, header=False, index=False)
print('Saving training labels to {}'.format(train_labels_output_path))
y_train.to_csv(train_labels_output_path, header=False, index=False)
print('Saving test labels to {}'.format(test_labels_output_path))
y_test.to_csv(test_labels_output_path, header=False, index=False)
# -
# Run this script as a processing job. Use the `SKLearnProcessor.run()` method. You give the `run()` method one `ProcessingInput` where the `source` is the census dataset in Amazon S3, and the `destination` is where the script reads this data from, in this case `/opt/ml/processing/input`. These local paths inside the processing container must begin with `/opt/ml/processing/`.
#
# Also give the `run()` method a `ProcessingOutput`, where the `source` is the path the script writes output data to. For outputs, the `destination` defaults to an S3 bucket that the Amazon SageMaker Python SDK creates for you, following the format `s3://sagemaker-<region>-<account_id>/<processing_job_name>/output/<output_name/`. You also give the ProcessingOutputs values for `output_name`, to make it easier to retrieve these output artifacts after the job is run.
#
# The `arguments` parameter in the `run()` method are command-line arguments in our `preprocessing.py` script.
# +
from sagemaker.processing import ProcessingInput, ProcessingOutput
sklearn_processor.run(code='preprocessing.py',
inputs=[ProcessingInput(
source=input_data,
destination='/opt/ml/processing/input')],
outputs=[ProcessingOutput(output_name='train_data',
source='/opt/ml/processing/train'),
ProcessingOutput(output_name='test_data',
source='/opt/ml/processing/test')],
arguments=['--train-test-split-ratio', '0.2']
)
preprocessing_job_description = sklearn_processor.jobs[-1].describe()
output_config = preprocessing_job_description['ProcessingOutputConfig']
for output in output_config['Outputs']:
if output['OutputName'] == 'train_data':
preprocessed_training_data = output['S3Output']['S3Uri']
if output['OutputName'] == 'test_data':
preprocessed_test_data = output['S3Output']['S3Uri']
# -
# Now inspect the output of the pre-processing job, which consists of the processed features.
training_features = pd.read_csv(preprocessed_training_data + '/train_features.csv', nrows=10)
print('Training features shape: {}'.format(training_features.shape))
training_features.head(n=10)
# ## Training using the pre-processed data
#
# We create a `SKLearn` instance, which we will use to run a training job using the training script `train.py`.
# +
from sagemaker.sklearn.estimator import SKLearn
sklearn = SKLearn(
entry_point='train.py',
framework_version='0.20.0',
instance_type="ml.m5.xlarge",
role=role)
# -
# The training script `train.py` trains a logistic regression model on the training data, and saves the model to the `/opt/ml/model` directory, which Amazon SageMaker tars and uploads into a `model.tar.gz` file into S3 at the end of the training job.
# +
# %%writefile train.py
import os
import pandas as pd
from sklearn.linear_model import LogisticRegression
from sklearn.externals import joblib
if __name__=="__main__":
training_data_directory = '/opt/ml/input/data/train'
train_features_data = os.path.join(training_data_directory, 'train_features.csv')
train_labels_data = os.path.join(training_data_directory, 'train_labels.csv')
print('Reading input data')
X_train = pd.read_csv(train_features_data, header=None)
y_train = pd.read_csv(train_labels_data, header=None)
model = LogisticRegression(class_weight='balanced', solver='lbfgs')
print('Training LR model')
model.fit(X_train, y_train)
model_output_directory = os.path.join('/opt/ml/model', "model.joblib")
print('Saving model to {}'.format(model_output_directory))
joblib.dump(model, model_output_directory)
# -
# Run the training job using `train.py` on the preprocessed training data.
sklearn.fit({'train': preprocessed_training_data})
training_job_description = sklearn.jobs[-1].describe()
model_data_s3_uri = '{}{}/{}'.format(
training_job_description['OutputDataConfig']['S3OutputPath'],
training_job_description['TrainingJobName'],
'output/model.tar.gz')
# ## Model Evaluation
#
# `evaluation.py` is the model evaluation script. Since the script also runs using scikit-learn as a dependency, run this using the `SKLearnProcessor` you created previously. This script takes the trained model and the test dataset as input, and produces a JSON file containing classification evaluation metrics, including precision, recall, and F1 score for each label, and accuracy and ROC AUC for the model.
#
# +
# %%writefile evaluation.py
import json
import os
import tarfile
import pandas as pd
from sklearn.externals import joblib
from sklearn.metrics import classification_report, roc_auc_score, accuracy_score
if __name__=="__main__":
model_path = os.path.join('/opt/ml/processing/model', 'model.tar.gz')
print('Extracting model from path: {}'.format(model_path))
with tarfile.open(model_path) as tar:
tar.extractall(path='.')
print('Loading model')
model = joblib.load('model.joblib')
print('Loading test input data')
test_features_data = os.path.join('/opt/ml/processing/test', 'test_features.csv')
test_labels_data = os.path.join('/opt/ml/processing/test', 'test_labels.csv')
X_test = pd.read_csv(test_features_data, header=None)
y_test = pd.read_csv(test_labels_data, header=None)
predictions = model.predict(X_test)
print('Creating classification evaluation report')
report_dict = classification_report(y_test, predictions, output_dict=True)
report_dict['accuracy'] = accuracy_score(y_test, predictions)
report_dict['roc_auc'] = roc_auc_score(y_test, predictions)
print('Classification report:\n{}'.format(report_dict))
evaluation_output_path = os.path.join('/opt/ml/processing/evaluation', 'evaluation.json')
print('Saving classification report to {}'.format(evaluation_output_path))
with open(evaluation_output_path, 'w') as f:
f.write(json.dumps(report_dict))
# +
import json
from sagemaker.s3 import S3Downloader
sklearn_processor.run(code='evaluation.py',
inputs=[ProcessingInput(
source=model_data_s3_uri,
destination='/opt/ml/processing/model'),
ProcessingInput(
source=preprocessed_test_data,
destination='/opt/ml/processing/test')],
outputs=[ProcessingOutput(output_name='evaluation',
source='/opt/ml/processing/evaluation')]
)
evaluation_job_description = sklearn_processor.jobs[-1].describe()
# -
# Now retrieve the file `evaluation.json` from Amazon S3, which contains the evaluation report.
# +
evaluation_output_config = evaluation_job_description['ProcessingOutputConfig']
for output in evaluation_output_config['Outputs']:
if output['OutputName'] == 'evaluation':
evaluation_s3_uri = output['S3Output']['S3Uri'] + '/evaluation.json'
break
evaluation_output = S3Downloader.read_file(evaluation_s3_uri)
evaluation_output_dict = json.loads(evaluation_output)
print(json.dumps(evaluation_output_dict, sort_keys=True, indent=4))
# -
# ## Running processing jobs with your own dependencies
#
# Above, you used a processing container that has scikit-learn installed, but you can run your own processing container in your processing job as well, and still provide a script to run within your processing container.
#
# Below, you walk through how to create a processing container, and how to use a `ScriptProcessor` to run your own code within a container. Create a scikit-learn container and run a processing job using the same `preprocessing.py` script you used above. You can provide your own dependencies inside this container to run your processing script with.
# !mkdir docker
# This is the Dockerfile to create the processing container. Install `pandas` and `scikit-learn` into it. You can install your own dependencies.
# +
# %%writefile docker/Dockerfile
FROM python:3.7-slim-buster
RUN pip3 install pandas==0.25.3 scikit-learn==0.21.3
ENV PYTHONUNBUFFERED=TRUE
ENTRYPOINT ["python3"]
# -
# This block of code builds the container using the `docker` command, creates an Amazon Elastic Container Registry (Amazon ECR) repository, and pushes the image to Amazon ECR.
# +
import boto3
account_id = boto3.client('sts').get_caller_identity().get('Account')
ecr_repository = 'sagemaker-processing-container'
tag = ':latest'
uri_suffix = 'amazonaws.com'
if region in ['cn-north-1', 'cn-northwest-1']:
uri_suffix = 'amazonaws.com.cn'
processing_repository_uri = '{}.dkr.ecr.{}.{}/{}'.format(account_id, region, uri_suffix, ecr_repository + tag)
# Create ECR repository and push docker image
# !docker build -t $ecr_repository docker
# !$(aws ecr get-login --region $region --registry-ids $account_id --no-include-email)
# !aws ecr create-repository --repository-name $ecr_repository
# !docker tag {ecr_repository + tag} $processing_repository_uri
# !docker push $processing_repository_uri
# -
# The `ScriptProcessor` class lets you run a command inside this container, which you can use to run your own script.
# +
from sagemaker.processing import ScriptProcessor
script_processor = ScriptProcessor(command=['python3'],
image_uri=processing_repository_uri,
role=role,
instance_count=1,
instance_type='ml.m5.xlarge')
# -
# Run the same `preprocessing.py` script you ran above, but now, this code is running inside of the Docker container you built in this notebook, not the scikit-learn image maintained by Amazon SageMaker. You can add the dependencies to the Docker image, and run your own pre-processing, feature-engineering, and model evaluation scripts inside of this container.
script_processor.run(code='preprocessing.py',
inputs=[ProcessingInput(
source=input_data,
destination='/opt/ml/processing/input')],
outputs=[ProcessingOutput(output_name='train_data',
source='/opt/ml/processing/train'),
ProcessingOutput(output_name='test_data',
source='/opt/ml/processing/test')],
arguments=['--train-test-split-ratio', '0.2']
)
script_processor_job_description = script_processor.jobs[-1].describe()
print(script_processor_job_description)
| sagemaker_processing/scikit_learn_data_processing_and_model_evaluation/scikit_learn_data_processing_and_model_evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # ChebLieNet: building graphs from sampled Lie groups
#
#
# In this tutorial, we introduce the notion of group manifold graph, a discretization of a Riemannian manifold. At the moment, four manifolds are available: the translation group $\mathbb{R}^2$, the roto-translation group $SE(2)$, the 3d rotation group $SO(3)$ and the 1-sphere $S(2)$.
#
# We define such a graph as following:
# - the vertices corresponds to **uniformly sampled** elements on the manifold,
# - the edges connects each vertex to its **K nearest neighbors**, w.r.t an **anisotropic riemannian distance**,
# - the edges' weights are computed by a **gaussian weight kernel** applied on the riemannian distance between vertices.
import torch
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# ## Create a graph manifold
from cheblienet.graphs.graphs import SE2GEGraph, SO3GEGraph, S2GEGraph, R2GEGraph, RandomSubGraph
r2_graph = R2GEGraph(
size=[28, 28, 1],
K=8,
sigmas=(1., 1., 1.),
path_to_graph="saved_graphs",
)
# +
eps = 0.1
se2_graph = SE2GEGraph(
size=[28, 28, 6],
K=16,
sigmas=(1., 1/eps**2, 2.048 / (28 ** 2)),
path_to_graph="saved_graphs"
)
# -
s2_graph = S2GEGraph(
size=[642, 1],
K=8,
sigmas=(1., 1., 1.),
path_to_graph="saved_graphs"
)
so3_graph = SO3GEGraph(
size=[642, 6],
K=32,
sigmas=(1., .1, 10/642),
path_to_graph="saved_graphs"
)
# ## Get informations
s2_graph.is_connected
s2_graph.is_undirected
s2_graph.manifold
s2_graph.num_vertices
s2_graph.num_edges # number of directed edges
s2_graph.vertex_index[:10]
s2_graph.vertex_attributes
s2_graph.vertex_beta[:10], s2_graph.vertex_gamma[:10]
s2_graph.edge_index[:10] # dim 0 is source, dim 1 is target
s2_graph.edge_weight[:10] # dim 0 is source, dim 1 is target
s2_graph.edge_sqdist[:10] # dim 0 is source, dim 1 is target
s2_graph.neighborhood(9) # neighbors index, edges' weights and squared riemannian distance
# ### Static visualization
# +
def plot_graph(graph, size):
M, L = size
fig = plt.figure(figsize=(5*L, 5))
X, Y, Z = graph.cartesian_pos()
for l in range(L):
ax = fig.add_subplot(1, L, l + 1, projection="3d")
ax.scatter(X[l*M:(l+1)*M], Y[l*M:(l+1)*M], Z[l*M:(l+1)*M], c="firebrick")
ax.axis("off")
fig.tight_layout()
def plot_graph_neighborhood(graph, index, size):
M, L = size
fig = plt.figure(figsize=(5, 5))
X, Y, Z = graph.cartesian_pos()
neighbors_indices, neighbors_weights, _ = graph.neighborhood(index)
weights = torch.zeros(graph.num_vertices)
weights[neighbors_indices] = neighbors_weights
for l in range(L):
ax = fig.add_subplot(L, 1, l + 1, projection="3d")
ax.scatter(X[l*M:(l+1)*M], Y[l*M:(l+1)*M], Z[l*M:(l+1)*M], c=weights[l*M:(l+1)*M], cmap=cm.PuRd)
ax.axis("off")
fig.tight_layout()
# -
plot_graph(s2_graph, [642, 1])
plot_graph_neighborhood(s2_graph, 406, [642, 1])
# ### Dynamic visualization
from cheblienet.graphs.viz import visualize_graph, visualize_graph_neighborhood, visualize_graph_signal
# +
eps = 0.1
xi = 6 / (28 ** 2)
se2_graph = SE2GEGraph(
size=[28, 28, 6],
K=32,
sigmas=(1., 1/eps, xi),
path_to_graph="saved_graphs"
)
# -
visualize_graph_neighborhood(se2_graph, 156)
so3_graph = SO3GEGraph(
size=[642, 6],
K=16,
sigmas=(1., .1, 10/642),
path_to_graph="saved_graphs"
)
visualize_graph(so3_graph)
signal = torch.rand(s2_graph.num_vertices)
visualize_graph_signal(s2_graph, signal)
# ## Random sub graph
random_subgraph = RandomSubGraph(s2_graph)
random_subgraph.num_vertices, random_subgraph.num_edges
random_subgraph.reinit()
random_subgraph.edges_sampling(0.9)
random_subgraph.num_vertices, random_subgraph.num_edges
random_subgraph.reinit()
random_subgraph.vertices_sampling(0.5)
random_subgraph.num_vertices, random_subgraph.num_edges
| notebooks/graph_manifold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# %pylab inline
# %load_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings('ignore')
warnings.simplefilter('ignore')
import os
import glob
from tqdm import tqdm, tqdm_notebook
import pandas as pd
import fitsne
from sklearn.model_selection import cross_val_score
from sklearn.metrics import confusion_matrix
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
from skimage.color import rgb2gray
from pyvirchow.io import WSIReader
from pyvirchow.io.operations import read_as_rgb
from pyvirchow.segmentation import poisson_deconvolve, perform_binary_cut, max_clustering
from pyvirchow.segmentation import collapse_labels, collapse_small_area, laplace_of_gaussian
from pyvirchow.segmentation import gmm_thresholding, label_nuclei, extract_features, summarize_region_properties
from pyvirchow.normalization import MacenkoNormalization
from pyvirchow.normalization import ReinhardNormalization
from pyvirchow.normalization import VahadaneNormalization
from pyvirchow.normalization import XuNormalization
from sklearn.decomposition import PCA, FastICA
from skimage.color import rgb2gray
from skimage.io import imread
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.manifold import TSNE
import umap
import seaborn as sns
sns.set_style('whitegrid')
sns.set_context('paper', font_scale=2)
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.datasets import load_wine
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn import linear_model
from sklearn.metrics import r2_score
from sklearn.linear_model import ElasticNet
from tpot import TPOTClassifier
import pickle
from multiprocessing import Pool
from sklearn.metrics import accuracy_score
import xgboost as xgb
from catboost import Pool, CatBoost
from catboost import CatBoostClassifier
from catboost import CatBoostRegressor
from sklearn.grid_search import GridSearchCV
scaler = StandardScaler()
# -
normal_patches_dir = '/Z/personal-folders/interns/saket/histopath_data/CAMELYON16_patches/normal_patches_test/level_0/'
tumor_patches_dir = '/Z/personal-folders/interns/saket/histopath_data/CAMELYON16_patches/tumor_patches_test/level_0/'
segmented_tsv_dir = '/Z/personal-folders/interns/saket/histopath_data/CAMELYON16_patches/normal_patches_test_segmented/level_0/'
# +
np.random.seed(42)
list_of_tumor_files = list(glob.glob('{}*.png'.format(tumor_patches_dir)))
list_of_normal_files = list(glob.glob('{}*.png'.format(normal_patches_dir)))
#list_of_tumor_files = list(np.random.choice(list_of_tumor_files, 20000))
#list_of_normal_files = list(np.random.choice(list_of_normal_files, 20000))
# -
def draw_nuclei(patch, local_max_search_radius=3, min_radius=5, max_radius=15, min_nucleus_area=100):
patch = read_as_rgb(patch)
label_nuclei(patch,
local_max_search_radius=local_max_search_radius,
min_radius=min_radius,
max_radius=max_radius,
min_nucleus_area=min_nucleus_area)
interact(draw_nuclei, patch=list_of_tumor_files+list_of_normal_files)
patch = read_as_rgb(list_of_tumor_files[0])
region_properties, fg_mask = label_nuclei(patch)
# +
features_df = []
labels = []
def process_sample(sample):
patch = read_as_rgb(sample)
region_properties, _ = label_nuclei(patch, draw=False)
summary = summarize_region_properties(region_properties,
patch)
return summary
#for sample in tqdm_notebook(list_of_normal_files):
with tqdm_notebook(total=len(list_of_tumor_files)) as pbar:
with Pool(processes=32) as p:
for i, summary in enumerate(p.imap_unordered(process_sample, list_of_tumor_files)):
pbar.update()
if summary is None:
print('Nothing found for {}'.format(sample))
continue
else:
labels.append('tumor')
features_df.append(summary)
pickle.dump(features_df, open('normal.pickle', 'wb'))
# -
with tqdm_notebook(total=len(list_of_normal_files)) as pbar:
with Pool(processes=32) as p:
for i, summary in enumerate(p.imap_unordered(process_sample, list_of_tumor_files)):
pbar.update()
if summary is None:
print('Nothing found for {}'.format(sample))
continue
else:
labels.append('normal')
features_df.append(summary)
pickle.dump(features_df, open('tumor.pickle', 'wb'))
#tfile = '/Z/personal-folders/interns/saket/histopath_data/CAMELYON16_patches/normal_patches_test/level_0/tumor_048_33856_186816_256.png'
y = np.array([1 if label=='normal' else 0 for label in labels])
f = pd.DataFrame(features_df)
X = f.values
X_scaled = scaler.fit(X).transform(X)
X_scaled.shape
# # PCA
#
# We start of with doing PCA/tSNE on the features.
#
# # UMAP
# +
embedding = umap.UMAP(n_neighbors=20,
min_dist=0.3,
metric='correlation').fit_transform(X_scaled)
fig = plt.figure(figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], label_matrix):
plt.scatter(embedding[y == i, 0], embedding[y == i, 1], color=color, alpha=.8, lw=lw,
label=target_name)
fig.tight_layout()
plt.title('UMAP')
# -
Y = fitsne.FItSNE(X_scaled.copy(order='C'))# max_iter=500)
# +
std_clf = make_pipeline(StandardScaler(), PCA(n_components=2))
std_clf.fit(X)
# + active=""
# """
# pca = PCA(n_components=2)
# X_r = pca.fit(X_scaled).transform(X_scaled)
#
#
# # Percentage of variance explained for each components
# print('explained variance ratio (first two components): %s'
# % str(pca.explained_variance_ratio_))
#
# fig = plt.figure(figsize=(10, 10))
# colors = ['navy', 'darkorange']
# lw = 0.2
# label_matrix = ['normal', 'tumor']
# for color, i, target_name in zip(colors, [0, 1], label_matrix):
# plt.scatter(X_r[y == i, 0], X_r[y == i, 1], color=color, alpha=.4, lw=lw,
# label=target_name)
# plt.legend(loc='best', shadow=False, scatterpoints=1)
# plt.title('PCA ')
# """
# +
colors = ['navy', 'darkorange']
lw = 0.2
label_matrix = ['normal', 'tumor']
fig = plt.figure(figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], label_matrix):
plt.scatter(Y[y == i, 0], Y[y == i, 1], color=color, alpha=.8, lw=lw,
label=target_name)
plt.legend(loc='best', shadow=False, scatterpoints=1)
fig.tight_layout()
plt.title('FIt-SNE')
# -
# # Random Forest
# +
RANDOM_STATE = 42
X_train, X_test, y_train, y_test = train_test_split(X_scaled, y,
test_size=0.30,
random_state=RANDOM_STATE)
rf = RandomForestClassifier(n_estimators=40)
print(clf.feature_importances_)
# -
# # LASSO
lasso = linear_model.Lasso(alpha = 0.1)
lasso.fit(X_train, y_train)
y_pred_lasso = lasso.predict(X_test)
r2_score_lasso = r2_score(y_test, y_pred_lasso)
r2_score_lasso
# +
alpha = 0.001
enet = ElasticNet(alpha=alpha, l1_ratio=0.7)
y_pred_enet = enet.fit(X_train, y_train).predict(X_test)
r2_score_enet = r2_score(y_test, y_pred_enet)
r2_score_enet
# -
pipeline_optimizer = TPOTClassifier(generations=5, population_size=20, cv=5,
random_state=42, verbosity=2)
pipeline_optimizer.fit(X_train, y_train)
print(pipeline_optimizer.score(X_test, y_test))
df = pd.DataFrame()
for f in list_of_normal_files:
uid = f.replace('.png', '.tsv').replace(os.path.dirname(f), '').replace('/', '')
temp_df = pd.read_table(os.path.join(segmented_tsv_dir, uid))
df = pd.concat([df, temp_df])
break
segmented_tsv_dir
df
# # Load df from files
import pandas as pd
normal_segmented_tsv_dir = '/Z/personal-folders/interns/saket/histopath_data/CAMELYON16_patches/normal_patches_test_segmented/level_0/'
tumor_segmented_tsv_dir = '/Z/personal-folders/interns/saket/histopath_data/CAMELYON16_patches/tumor_patches_test_segmented/level_0/'
# +
df = pd.DataFrame()
y = []
def load_df(path):
temp_df = pd.read_table(path)
if len(temp_df.index):
return temp_df
return None
normal_segmented_tsv = glob.glob(normal_segmented_tsv_dir+'/*.tsv')
tumor_segmented_tsv = glob.glob(tumor_segmented_tsv_dir+'/*.tsv')
np.random.seed(42)
normal_segmented_tsv_subsampled = np.random.choice(normal_segmented_tsv, 10000)
tumor_segmented_tsv_subsampled = np.random.choice(tumor_segmented_tsv, 10000)
with tqdm_notebook(total=len(normal_segmented_tsv_subsampled)) as pbar:
with Pool(processes=32) as p:
for i, temp_df in enumerate(p.imap_unordered(load_df, normal_segmented_tsv_subsampled)):
if temp_df is not None:
df = pd.concat((df, temp_df))
y.append(0)
pbar.update()
# -
with tqdm_notebook(total=len(tumor_segmented_tsv_subsampled)) as pbar:
with Pool(processes=32) as p:
for i, temp_df in enumerate(p.imap_unordered(load_df, tumor_segmented_tsv_subsampled)):
if temp_df is not None:
df = pd.concat((df, temp_df))
y.append(1)
pbar.update()
y = np.array(y)
#label = [0 for x in range(len(glob.glob(normal_segmented_tsv_dir+'/*.tsv')))]
#label += [1 for x in range(len(df.index)-len(label))]
label =y
len(label)
df.head()
df_with_label = df.copy()#.drop(columns=['0'])
df_with_label['label'] = np.nan
df_with_label['label'] = label
df_with_label.head()
# +
df_with_label = df_with_label.dropna()
label = df_with_label['label']
df_with_label = df_with_label.drop(columns=['label'])
df_with_label.to_csv('normal_tumor_segmented_df.tsv', sep='\t', index=False, header=True)
# -
matrix = df_with_label.as_matrix()
matrix.shape
X_scaled = scaler.fit(matrix).transform(matrix)
# +
RANDOM_STATE = 42
X_train, X_test, y_train, y_test = train_test_split(X_scaled, label,
test_size=0.30,
random_state=RANDOM_STATE)
# -
lasso = linear_model.Lasso(alpha = 0.1)
lasso.fit(X_train, y_train)
y_pred_lasso = lasso.predict(X_test)
r2_score_lasso = r2_score(y_test, y_pred_lasso)
r2_score_lasso
# +
alpha = 0.001
enet = ElasticNet(alpha=alpha, l1_ratio=0.7)
y_pred_enet = enet.fit(X_train, y_train).predict(X_test)
r2_score_enet = r2_score(y_test, y_pred_enet)
r2_score_enet
# -
pipeline_optimizer = TPOTClassifier(generations=5, population_size=20, cv=5,
random_state=42, verbosity=2)
pipeline_optimizer.fit(X_train, y_train)
print(pipeline_optimizer.score(X_test, y_test))
pipeline_optimizer.export('tpot_exported_pipeline_10ksamples.py')
# +
dtrain = xgb.DMatrix(X_train, label=y_train)
dtest = xgb.DMatrix(X_test, label=y_test)
param = {'max_depth':2, 'eta':1, 'silent':1, 'objective':'binary:logistic' }
num_round = 2
bst = xgb.train(param, dtrain, num_round)
# make prediction
preds = bst.predict(dtest)
# -
xgb.plot_importance(bst)
df.columns[4]
accuracy_score(y_test, [round(x) for x in preds])
model = CatBoostClassifier(iterations=2, depth=2, learning_rate=1,
loss_function='Logloss', logging_level='Verbose')
#train the model
model.fit(X_train, y_train, cat_features=[0,1])
# make the prediction using the resulting model
preds_class = model.predict(X_test)
preds_proba = model.predict_proba(X_test)
print("class = ", preds_class)
print("proba = ", preds_proba)
xgb.plot_tree(bst, num_trees=2)
# +
xgb_model = xgb.XGBClassifier()
optimization_dict = {'max_depth': [2,4,6],
'n_estimators': [50,100,200]}
model = GridSearchCV(xgb_model, optimization_dict,
scoring='accuracy', verbose=1)
model.fit(X_train, y_train)
print(model.best_score_)
print(model.best_params_)
# -
# # Test on test dataset
test_tumor_segmented_tsv_dir = '/Z/personal-folders/interns/saket/histopath_data/baidu_images/test_tumor_level0_segmented/level_0/'
test_normal_segmented_tsv_dir = '/Z/personal-folders/interns/saket/histopath_data/baidu_images/test_normal_level0_segmented/level_0/'
from multiprocessing import Pool
# +
test_df = pd.DataFrame()
test_y = []
test_normal_segmented_tsv = glob.glob(test_normal_segmented_tsv_dir+'/*.tsv')
test_tumor_segmented_tsv = glob.glob(test_tumor_segmented_tsv_dir+'/*.tsv')
np.random.seed(42)
test_normal_segmented_tsv_subsampled = test_normal_segmented_tsv
#np.random.choice(test_normal_segmented_tsv, 10000)
test_tumor_segmented_tsv_subsampled = test_tumor_segmented_tsv
#np.random.choice(tumor_segmented_tsv, 10000)
with tqdm_notebook(total=len(test_normal_segmented_tsv_subsampled)) as pbar:
with Pool(processes=32) as p:
for i, temp_df in enumerate(p.imap_unordered(load_df, test_normal_segmented_tsv_subsampled)):
if temp_df is not None:
test_df = pd.concat((test_df, temp_df))
test_y.append(0)
pbar.update()
# -
with tqdm_notebook(total=len(test_tumor_segmented_tsv_subsampled)) as pbar:
with Pool(processes=32) as p:
for i, temp_df in enumerate(p.imap_unordered(load_df, test_tumor_segmented_tsv_subsampled)):
if temp_df is not None:
test_df = pd.concat((test_df, temp_df))
test_y.append(1)
pbar.update()
test_df['label'] = test_y
test_df = test_df.dropna()
test_df.to_csv('./test_tumor_segmented_df.tsv', index=False, header=True, sep='\t')
label = test_df['label']
test_df_nolabel = test_df.copy()
test_df_nolabel = test_df_nolabel.drop(columns=['label'])
test_matrix = test_df_nolabel.as_matrix()
test_matrix.shape
matrix.shape
testX_scaled = scaler.fit(test_matrix).transform(test_matrix)
print(pipeline_optimizer.score(testX_scaled, label))
df.columns
| notebooks/06.Lasso.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
model_file_path = "/home/gsoykan/Desktop/comp541/comp541_term_project/results/imagenet-resnet-152-dag.mat"
cat_img_url = "https://nextjournal.com/data/QmXNbi2LE7u6yBdBXaQ9E2zGb48FELg3TxjrLiPKBmdvZc?filename=Qat.jpg&content-type=image/jpeg"
resnet50_model_file_path = "/userfiles/gsoykan20/resnet_pretrained/imagenet-resnet-50-dag.mat"
# +
# https://nextjournal.com/mpd/image-classification-with-knet
# -
import CUDA
using MAT, OffsetArrays, FFTViews, ArgParse, Images, Knet, ImageMagick, Printf
include("modular.resnet.jl")
atype = CUDA.functional() ? KnetArray{Float32} : Array{Float32}
Knet.atype() = atype
function get_params(params, atype)
len = length(params["value"])
ws, ms = [], []
for k = 1:len
name = params["name"][k]
value = convert(Array{Float32}, params["value"][k])
if endswith(name, "moments")
push!(ms, reshape(value[:,1], (1,1,size(value,1),1)))
push!(ms, reshape(value[:,2], (1,1,size(value,1),1)))
elseif startswith(name, "bn")
push!(ws, reshape(value, (1,1,length(value),1)))
elseif startswith(name, "fc") && endswith(name, "filter")
push!(ws, transpose(reshape(value,(size(value,3),size(value,4)))))
elseif startswith(name, "conv") && endswith(name, "bias")
push!(ws, reshape(value, (1,1,length(value),1)))
else
push!(ws, value)
end
end
map(wi->convert(atype, wi), ws),
map(mi->convert(atype, mi), ms)
end
# From vgg.jl
function data(img, averageImage)
if occursin("://",img)
@info "Downloading $img"
img = download(img)
end
a0 = load(img)
new_size = ntuple(i->div(size(a0,i)*224,minimum(size(a0))),2)
a1 = Images.imresize(a0, new_size)
i1 = div(size(a1,1)-224,2)
j1 = div(size(a1,2)-224,2)
b1 = a1[i1+1:i1+224,j1+1:j1+224]
c1 = permutedims(channelview(b1), (3,2,1))
d1 = convert(Array{Float32}, c1)
e1 = reshape(d1[:,:,1:3], (224,224,3,1))
f1 = (255 * e1 .- averageImage)
g1 = permutedims(f1, [2,1,3,4])
end
# +
# OLD IMPLEMENTATION
# Batch Normalization Layer
# works both for convolutional and fully connected layers
# mode, 0=>train, 1=>test
function batchnorm(w, x, ms; mode=1, epsilon=1e-5)
mu, sigma = nothing, nothing
if mode == 0
d = ndims(x) == 4 ? (1,2,4) : (2,)
s = prod(size(x,d...))
mu = sum(x,d) / s
x0 = x .- mu
x1 = x0 .* x0
sigma = sqrt(epsilon + (sum(x1, d)) / s)
elseif mode == 1
mu = popfirst!(ms)
sigma = popfirst!(ms)
end
# we need getval in backpropagation
push!(ms, AutoGrad.value(mu), AutoGrad.value(sigma))
xhat = (x.-mu) ./ sigma
return w[1] .* xhat .+ w[2]
end
function reslayerx0(w,x,ms; padding=0, stride=1, mode=1)
b = conv4(w[1],x; padding=padding, stride=stride)
bx = batchnorm(w[2:3],b,ms; mode=mode)
end
function reslayerx1(w,x,ms; padding=0, stride=1, mode=1)
relu.(reslayerx0(w,x,ms; padding=padding, stride=stride, mode=mode))
end
function reslayerx2(w,x,ms; pads=[0,1,0], strides=[1,1,1], mode=1)
ba = reslayerx1(w[1:3],x,ms; padding=pads[1], stride=strides[1], mode=mode)
bb = reslayerx1(w[4:6],ba,ms; padding=pads[2], stride=strides[2], mode=mode)
bc = reslayerx0(w[7:9],bb,ms; padding=pads[3], stride=strides[3], mode=mode)
end
function reslayerx3(w,x,ms; pads=[0,0,1,0], strides=[2,2,1,1], mode=1) # 12
a = reslayerx0(w[1:3],x,ms; stride=strides[1], padding=pads[1], mode=mode)
b = reslayerx2(w[4:12],x,ms; strides=strides[2:4], pads=pads[2:4], mode=mode)
relu.(a .+ b)
end
function reslayerx4(w,x,ms; pads=[0,1,0], strides=[1,1,1], mode=1)
relu.(x .+ reslayerx2(w,x,ms; pads=pads, strides=strides, mode=mode))
end
function reslayerx5(w,x,ms; strides=[2,2,1,1], mode=1)
x = reslayerx3(w[1:12],x,ms; strides=strides, mode=mode)
for k = 13:9:length(w)
x = reslayerx4(w[k:k+8],x,ms; mode=mode)
end
return x
end
# mode, 0=>train, 1=>test
function resnet152(w,x,ms; mode=1)
# layer 1
conv1 = reslayerx1(w[1:3],x,ms; padding=3, stride=2, mode=mode)
pool1 = pool(conv1; window=3, stride=2)
# layer 2,3,4,5
r2 = reslayerx5(w[4:33], pool1, ms; strides=[1,1,1,1], mode=mode)
r3 = reslayerx5(w[34:108], r2, ms; mode=mode)
r4 = reslayerx5(w[109:435], r3, ms; mode=mode)
r5 = reslayerx5(w[436:465], r4, ms; mode=mode)
# fully connected layer
pool5 = pool(r5; stride=1, window=7, mode=2)
fc1000 = w[466] * mat(pool5) .+ w[467]
end
# -
should_use_resnet_50 = true
o = Dict(
:atype => KnetArray{Float32},
:model => should_use_resnet_50 ? resnet50_model_file_path : model_file_path,
:image => cat_img_url,
:top => 10
)
@info "Reading $(o[:model])"
model = matread(abspath(o[:model]))
avgimg = model["meta"]["normalization"]["averageImage"]
avgimg = convert(Array{Float32}, avgimg)
description = model["meta"]["classes"]["description"]
w, ms = get_params(model["params"], o[:atype])
@info "Reading $(o[:image])"
img = data(o[:image], avgimg)
img = convert(o[:atype], img)
function predict(o)
@info "Reading $(o[:model])"
model = matread(abspath(o[:model]))
avgimg = model["meta"]["normalization"]["averageImage"]
avgimg = convert(Array{Float32}, avgimg)
description = model["meta"]["classes"]["description"]
w, ms = get_params(model["params"], o[:atype])
@info "Reading $(o[:image])"
img = data(o[:image], avgimg)
img = convert(o[:atype], img)
@info "Classifying."
#@time y1 = resnet152(w,img,ms)
modular_resnet152 = generate_resnet_from_weights(w, ms)
y1 = modular_resnet152(img)
return y1, description
end
#model = matread(abspath(o[:model]))
#w, ms = get_params(model["params"], o[:atype])
Knet.atype()
# +
# Batch Norm Fixing Codes
function init_model(;et=Float32)
# Use bnparams() to initialize gammas and betas
w = Any[
kaiming(et, 3, 3, 3, 16), bnparams(et, 16),
kaiming(et, 3, 3, 16, 32), bnparams(et, 32),
kaiming(et, 3, 3, 32, 64), bnparams(et, 64),
xavier(et, 100, 8 * 8 * 64), bnparams(et, 100),
xavier(et, 10, 100), zeros(et, 10, 1)
]
# Initialize a moments object for each batchnorm
m = Any[bnmoments() for i = 1:4]
w = map(Knet.array_type[], w)
return w, m
end
function conv_layer(w, m, x; maxpool=true)
o = conv4(w[1], x; padding=1)
o = batchnorm(o, m, w[2])
o = relu.(o)
if maxpool; o=pool(o); end
return o
end
moments = bnmoments()
params = bnparams(C)
...
### size(x) -> (H, W, C, N)
y = batchnorm(x, moments, params)
model = matread(abspath(o[:model]))
avgimg = model["meta"]["normalization"]["averageImage"]
avgimg = convert(Array{Float32}, avgimg)
description = model["meta"]["classes"]["description"]
w, ms = get_params(model["params"], o[:atype])
@info "Reading $(o[:image])"
img = data(o[:image], avgimg)
img = convert(o[:atype], img);
res_conv_0 = ResLayerX0(w[1:3], ms; padding=3, stride=2)
_wsize(y) = ((1 for _=1:ndims(y)-2)..., size(y)[end-1], 1)
_bnscale(param) = param[1:div(length(param), 2)]
_bnbias(param) = param[div(length(param), 2)+1:end]
_bnscale(w[2:3])
_bnbias(w[2:3])[begin]
o = conv4(w[1], img; padding=3, stride=2)
res_mean = popfirst!(ms)
res_variance = popfirst!(ms)
batch_ms = bnmoments(meaninit=res_mean, varinit=res_variance)
bnmoments()
f_res_mean = convert(Array{Float32}, res_mean)
f_res_variance = convert(Array{Float32}, res_variance)
f_batch_ms = bnmoments(mean=res_mean, var=res_variance)
function var_function(eltype, dims...)
return convert(eltype, f_res_variance)
end
function mean_function(eltype, dims...)
f_res_mean = convert(eltype, f_res_mean)
end
_wsize(o)
_bnscale(vcatted_ws)
#w2 = convert(Array{Float32}, w[2])
#w3 = convert(Array{Float32}, w[3])
w2 = w[2]
w3 = w[3]
vec_w2 = vec(w2)
vec_w3 = vec(w3)
vcatted_ws = vcat(vec_w2, vec_w3)
batchnorm(o, f_batch_ms, vcatted_ws)
batchnorm(o, bnmoments(), vcatted_ws)
res_conv_0(img)
# -
w, ms = get_params(model["params"], o[:atype]);
modular_resnet50 = generate_resnet50_from_weights(w, ms);
y_50 = modular_resnet50(img)
modular_resnet152 = generate_resnet_from_weights(w, ms)
y1 = modular_resnet152(img)
# +
function y_to_readable_output(raw_y)
z = vec(Array(raw_y))
s = sortperm(z,rev=true)
p = exp.(logp(z))
for ind in s[1:o[:top]]
print("$(description[ind]): $(@sprintf("%.2f",p[ind]*100))%\n")
end
end
# -
y_to_readable_output(y_50)
| resnet.example.ipynb |
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
All different except 0 Google CP Solver.
Decomposition of global constraint alldifferent_except_0.
From Global constraint catalogue:
http://www.emn.fr/x-info/sdemasse/gccat/Calldifferent_except_0.html
'''
Enforce all variables of the collection VARIABLES to take distinct
values, except those variables that are assigned to 0.
Example
(<5, 0, 1, 9, 0, 3>)
The alldifferent_except_0 constraint holds since all the values
(that are different from 0) 5, 1, 9 and 3 are distinct.
'''
Compare with the following models:
* Comet: http://hakank.org/comet/alldifferent_except_0.co
* ECLiPSe: http://hakank.org/eclipse/alldifferent_except_0.ecl
* Tailor/Essence': http://hakank.org/tailor/alldifferent_except_0.eprime
* Gecode: http://hakank.org/gecode/alldifferent_except_0.cpp
* Gecode/R: http://hakank.org/gecode_r/all_different_except_0.rb
* MiniZinc: http://hakank.org/minizinc/alldifferent_except_0.mzn
* SICStus_ http://hakank.org/sicstus/alldifferent_except_0.pl
* Choco: http://hakank.org/choco/AllDifferentExcept0_test.java
* JaCoP: http://hakank.org/JaCoP/AllDifferentExcept0_test.java
* Zinc: http://hakank.org/minizinc/alldifferent_except_0.zinc
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
from __future__ import print_function
from ortools.constraint_solver import pywrapcp
#
# Decomposition of alldifferent_except_0
# Thanks to Laurent Perron (Google) for
# suggestions of improvements.
#
def alldifferent_except_0(solver, a):
n = len(a)
for i in range(n):
for j in range(i):
solver.Add((a[i] != 0) * (a[j] != 0) <= (a[i] != a[j]))
# more compact version:
def alldifferent_except_0_b(solver, a):
n = len(a)
[
solver.Add((a[i] != 0) * (a[j] != 0) <= (a[i] != a[j]))
for i in range(n)
for j in range(i)
]
# Create the solver.
solver = pywrapcp.Solver("Alldifferent except 0")
# data
n = 7
# declare variables
x = [solver.IntVar(0, n - 1, "x%i" % i) for i in range(n)]
# Number of zeros.
z = solver.Sum([x[i] == 0 for i in range(n)]).VarWithName("z")
#
# constraints
#
alldifferent_except_0(solver, x)
# we require 2 0's
solver.Add(z == 2)
#
# solution and search
#
solution = solver.Assignment()
solution.Add([x[i] for i in range(n)])
solution.Add(z)
collector = solver.AllSolutionCollector(solution)
solver.Solve(
solver.Phase([x[i] for i in range(n)], solver.CHOOSE_FIRST_UNBOUND,
solver.ASSIGN_MIN_VALUE), [collector])
num_solutions = collector.SolutionCount()
for s in range(num_solutions):
print("x:", [collector.Value(s, x[i]) for i in range(n)])
print("z:", collector.Value(s, z))
print()
print("num_solutions:", num_solutions)
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
| examples/notebook/contrib/alldifferent_except_0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [source](../../api/alibi_detect.od.sr.rst)
# # Spectral Residual
# ## Overview
#
# The Spectral Residual outlier detector is based on the paper [Time-Series Anomaly Detection Service at Microsoft](https://arxiv.org/abs/1906.03821) and is suitable for **unsupervised online anomaly detection in univariate time series** data. The algorithm first computes the [Fourier Transform](https://en.wikipedia.org/wiki/Fourier_transform) of the original data. Then it computes the *spectral residual* of the log amplitude of the transformed signal before applying the Inverse Fourier Transform to map the sequence back from the frequency to the time domain. This sequence is called the *saliency map*. The anomaly score is then computed as the relative difference between the saliency map values and their moving averages. If the score is above a threshold, the value at a specific timestep is flagged as an outlier. For more details, please check out the [paper](https://arxiv.org/abs/1906.03821).
# ## Usage
#
# ### Initialize
#
# Parameters:
#
# * `threshold`: Threshold used to classify outliers. Relative saliency map distance from the moving average.
#
# * `window_amp`: Window used for the moving average in the *spectral residual* computation. The spectral residual is the difference between the log amplitude of the Fourier Transform and a convolution of the log amplitude over `window_amp`.
#
# * `window_local`: Window used for the moving average in the outlier score computation. The outlier score computes the relative difference between the saliency map and a moving average of the saliency map over `window_local` timesteps.
#
# * `padding_amp_method`:
# Padding method to be used prior to each convolution over log amplitude.
# Possible values: `constant` | `replicate` | `reflect`. Default value: `replicate`.
#
# - `constant` - padding with constant 0.
#
# - `replicate` - repeats the last/extreme value.
#
# - `reflect` - reflects the time series.
#
# * `padding_local_method`:
# Padding method to be used prior to each convolution over saliency map.
# Possible values: `constant` | `replicate` | `reflect`. Default value: `replicate`.
#
# - `constant` - padding with constant 0.
#
# - `replicate` - repeats the last/extreme value.
#
# - `reflect` - reflects the time series.
#
# * `padding_amp_side`:
# Whether to pad the amplitudes on both sides or only on one side.
# Possible values: `bilateral` | `left` | `right`.
#
# * `n_est_points`: Number of estimated points padded to the end of the sequence.
#
# * `n_grad_points`: Number of points used for the gradient estimation of the additional points padded to the end of the sequence. The paper sets this value to 5.
#
# Initialized outlier detector example:
#
# ```python
# from alibi_detect.od import SpectralResidual
#
# od = SpectralResidual(
# threshold=1.,
# window_amp=20,
# window_local=20,
# padding_amp_method='reflect',
# padding_local_method='reflect',
# padding_amp_side='bilateral',
# n_est_points=10,
# n_grad_points=5
# )
# ```
#
# It is often hard to find a good threshold value. If we have a time series containing both normal and outlier data and we know approximately the percentage of normal data in the time series, we can infer a suitable threshold:
#
# ```python
# od.infer_threshold(
# X,
# t=t, # array with timesteps, assumes dt=1 between observations if omitted
# threshold_perc=95
# )
# ```
# ### Detect
#
# We detect outliers by simply calling `predict` on a time series `X` to compute the outlier scores and flag the anomalies. We can also return the instance (timestep) level outlier score by setting `return_instance_score` to True.
#
# The prediction takes the form of a dictionary with `meta` and `data` keys. `meta` contains the detector's metadata while `data` is also a dictionary which contains the actual predictions stored in the following keys:
#
# * `is_outlier`: boolean whether instances are above the threshold and therefore outlier instances. The array is of shape *(timesteps,)*.
#
# * `instance_score`: contains instance level scores if `return_instance_score` equals True.
#
#
# ```python
# preds = od.predict(
# X,
# t=t, # array with timesteps, assumes dt=1 between observations if omitted
# return_instance_score=True
# )
# ```
# ## Examples
#
# [Time series outlier detection with Spectral Residuals on synthetic data](../../examples/od_sr_synth.ipynb)
| doc/source/od/methods/sr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# Shut up warnings
def warn(*args, **kwargs):
pass
import warnings
warnings.warn = warn
df = pd.read_csv("atrEuropa.csv",index_col="Last Name:")
# # For this, we'll only keep Keratoconus or normal patients.
df
df.fillna(value=0,inplace=True)
# # Removing static measures
df = df.drop(columns=["ARTh","SP A1"])
df = df[(df["Tipo"]=="Ceratocone")|(df["Tipo"]=="Normal")]
df["Tipo"] = df["Tipo"].apply(lambda x:1 if x=="Ceratocone" else 0)
seletores = ["Error","QS.1","Error IOP","QS IOP","Error Pachy","QS Pachy"]
dfSelect = df[seletores]
df = df.drop(columns=seletores)
# # TBI e CBI são vetores resposta de outros algoritmos preditivos, que usaremos como objetivos de igualar ou ultrapassar.
Outros = ["TBI","CBI"]
medidas = df[Outros]
from sklearn.metrics import accuracy_score,confusion_matrix,precision_score,roc_auc_score,recall_score
soloMetrics = accuracy_score,precision_score,roc_auc_score,recall_score
names = ["Accuracy","Precision","Roc_AUC","Recall"]
def scoreIt(predicted,real,intercept,metrics,metricNames,name):
mam = []
for metr in soloMetrics:
mam.append(metr(predicted.apply(lambda x:1 if x>intercept else 0),real))
return pd.DataFrame(mam,index=metricNames,columns = [name])
TBI = scoreIt(medidas["TBI"],df["Tipo"],0.5,soloMetrics,names,"TBI")
CBI = scoreIt(medidas["CBI"],df["Tipo"],0.5,soloMetrics,names,"CBI")
Old = pd.concat([TBI,CBI],axis=1)
Old
df = df.drop(columns=Outros)
df.head()
pedro = df[df.columns[:18]]
exams = df[df.columns[19:]]
exams["Tipo"] = df["Tipo"]
exams
from sklearn.model_selection import cross_validate
from sklearn.linear_model import LogisticRegression
from sklearn.neural_network import MLPClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
from sklearn.pipeline import Pipeline
from sklearn.feature_selection import RFECV
from sklearn.model_selection import StratifiedKFold
import sklearn
metrics = ["accuracy","f1","precision","recall","roc_auc"]
results = ['test_accuracy', 'test_f1', 'test_precision', 'test_recall', 'test_roc_auc']
models = [SVC(),LogisticRegression(),DecisionTreeClassifier()]
def tryIt(dataframe,models,metrics,results):
res = []
names = []
for model in models:
result = []
score = cross_validate(model,dataframe.drop(columns="Tipo"),dataframe["Tipo"],return_train_score=False,cv=10,scoring=metrics)
for k in results:
result.append([k,np.median(score[k])])
names.append(str(model.__class__).split(".")[-1].split("'")[0])
res.append(result)
return np.array(res),names
def showResults(res,names):
return pd.concat((pd.DataFrame(res[k,:,1:],index=res[0,:,0],columns=[names[k]+"-Median"]) for k in range(len(res))),axis=1).T
res,names = tryIt(exams,models,metrics,results)
showResults(res,names)
# # Multi-layer perceptron
# - We should scale the values from 0.1 to 0.9, literature says so.
from sklearn.preprocessing import MinMaxScaler
data = exams
scaler = MinMaxScaler()
scaler.fit(data)
ScaledData = pd.DataFrame(scaler.transform(data),columns=data.columns,index=data.index)
models.append(MLPClassifier(max_iter=300))
exams.columns
res,names = tryIt(ScaledData,models,metrics,results)
showResults(res,names)
# # Coeficientes da regressão logística
LR = LogisticRegression().fit(exams.drop(columns="Tipo"),exams["Tipo"])
pd.DataFrame(LR.coef_,columns=exams.drop(columns="Tipo").columns).T.sort_values(by=0,ascending=False)
# # Qual a importância dada aos atributos pela árvore de decisão?
tree = DecisionTreeClassifier().fit(exams.drop(columns="Tipo"),exams["Tipo"])
importance = pd.DataFrame(tree.feature_importances_,index=exams.drop(columns="Tipo").columns,columns=["Importance"])
importance.sort_values(by="Importance",ascending=False)
# # Now with some refined hyper-parametrization, grid search at first:
# - First runner up, Logistic Regression.
parameters = {
"penalty":["l2"],
"dual":[False],
"C":np.arange(0.5,10,0.5),
"solver":["newton-cg", "lbfgs", "liblinear", "sag", "saga"],
"max_iter":range(100,300,100)
}
from sklearn.model_selection import GridSearchCV
clf = GridSearchCV(LogisticRegression(),parameters,scoring=metrics,cv=10,refit="roc_auc",return_train_score=False,n_jobs=-1)
clf.fit(exams.drop(columns="Tipo"),exams["Tipo"]);
best = clf.best_estimator_
bestParams = clf.best_params_
clf = best
clf = LogisticRegression(C=7.75,solver="lbfgs")
rfecv = RFECV(clf,cv=StratifiedKFold(10),scoring="roc_auc")
rfecv.fit(exams.drop(columns="Tipo"),exams["Tipo"])
plt.figure()
plt.xlabel("Number of features selected")
plt.ylabel("Cross validation score (nb of correct classifications)")
plt.plot(range(1, len(rfecv.grid_scores_) + 1), rfecv.grid_scores_)
plt.show()
# # Analyzing Pedro's base
pedro.head()
clf = GridSearchCV(LogisticRegression(),parameters,scoring=metrics,cv=10,refit="roc_auc",return_train_score=False,n_jobs=-1)
clf.fit(pedro.drop(columns="Tipo"),pedro["Tipo"]);
best = clf.best_estimator_
bestParams = clf.best_params_
clf = best
clf = LogisticRegression(C=7.75,solver="lbfgs")
rfecv = RFECV(clf,cv=StratifiedKFold(10),scoring="roc_auc")
rfecv.fit(pedro.drop(columns="Tipo"),pedro["Tipo"])
plt.figure()
plt.xlabel("Number of features selected")
plt.ylabel("Cross validation score (nb of correct classifications)")
plt.plot(range(1, len(rfecv.grid_scores_) + 1), rfecv.grid_scores_)
plt.show()
# ### The difference in roc_auc score after a certain point is less than 0.5, we can choose fewer features and have roughly the same classfication power.
from sklearn.feature_selection import SelectKBest, mutual_info_classif, RFE
new = RFE(clf,n_features_to_select=10)
new.fit(exams.drop(columns="Tipo"),exams["Tipo"]);
# ## Selected features
list(exams.drop(columns="Tipo").columns[new.support_])
# ## Selected estimator
# - It is already fit to the reduced dataset, so we can save this and simply use it later
import pickle
with open("LogReg.pickle","wb") as f:
pickle.dump(new.estimator_,f)
# ## After reducing the dataset, we can save that and use it at the application
reduced = pd.DataFrame(new.transform(exams.drop(columns="Tipo")),columns=exams.drop(columns="Tipo").columns[new.support_],index=exams.index)
reduced["Tipo"] = exams["Tipo"]
reduced.head()
reduced.to_csv("reducedExams.csv")
| Data mining process.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Automated Transformations
#
# Automated transformations provide convenient handling of constrained
# continuous variables during inference by transforming them to an
# unconstrained space. Automated transformations are crucial for
# expanding the scope of algorithm classes such as gradient-based Monte
# Carlo and variational inference with reparameterization gradients.
#
# A webpage version of this tutorial is available at
# http://edwardlib.org/tutorials/automated-transformations.
#
# ## The Transform Primitive
#
# Automated transformations in Edward are enabled through the key
# primitive
# [`ed.transform`](http://edwardlib.org/api/ed/transform).
# It takes as input a (possibly constrained) continuous random variable
# $\mathbf{x}$, defaults to a choice of transformation $T$, and returns a
# [`TransformedDistribution`](http://edwardlib.org/api/ed/models/TransformedDistribution)
# $\mathbf{y}=T(\mathbf{x})$ with unconstrained support.
# An optional argument allows you to manually specify the transformation.
#
# The returned random variable $\mathbf{y}$'s density is the original
# random variable $\mathbf{x}$'s density adjusted by the determinant of
# the Jacobian of the inverse transformation (<NAME>, 2002),
#
# $$p(\mathbf{y}) = p(\mathbf{x})~|\mathrm{det}~J_{T^{-1}}(\mathbf{y}) |.$$
#
# Intuitively, the Jacobian describes how a transformation warps unit
# volumes across spaces. This matters for transformations of random
# variables, since probability density functions must always integrate
# to one.
#
# ## Automated Transformations in Inference
#
# To use automated transformations during inference, set the flag
# argument `auto_transform=True` in `inference.initialize`
# (or the all-encompassing method `inference.run`):
#
# ```python
# inference.initialize(auto_transform=True)
# ```
#
# By default, the flag is already set to `True`.
# With this flag, any key-value pair passed into inference's
# `latent_vars` with unequal support is transformed to the
# unconstrained space; no transformation is applied if already
# unconstrained. The algorithm is then run under
# `inference.latent_vars`, which explicitly stores the
# transformed latent variables and forgets the constrained ones.
#
# We illustrate automated transformations in a few inference examples.
# Imagine that the target distribution is a Gamma distribution.
# +
# %matplotlib inline
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import edward as ed
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
from edward.models import Empirical, Gamma, Normal
from tensorflow.contrib.distributions import bijectors
# -
x = Gamma(1.0, 2.0)
# This example is only used for illustration, but note this context of
# inference with latent variables of non-negative support occur
# frequently: for example, this appears when applying topic models with a deep exponential
# family where we might use a normal variational
# approximation to implicitly approximate latent variables with Gamma
# priors (in
# [`examples/deep_exponential_family.py`](https://github.com/blei-lab/edward/blob/master/examples/deep_exponential_family.py),
# we explicitly define a non-negative variational approximation).
#
# __Variational inference.__
# Consider a Normal variational approximation
# and use the algorithm [`ed.KLqp`](http://edwardlib.org/api/ed/KLqp).
# +
qx = Normal(loc=tf.Variable(tf.random_normal([])),
scale=tf.nn.softplus(tf.Variable(tf.random_normal([]))))
inference = ed.KLqp({x: qx})
inference.run()
# -
# The Gamma and Normal distribution have unequal support, so inference
# transforms both to the unconstrained space; normal is already
# unconstrained so only Gamma is transformed. `ed.KLqp` then
# optimizes with
# [reparameterization gradients](http://edwardlib.org/api/klqp).
# This means the Normal distribution's parameters are optimized to match
# the transformed (unconstrained) Gamma distribution.
#
# Oftentimes we'd like the approximation on the original (constrained)
# space. This was never needed for inference, so we must explicitly
# build it by first obtaining the target distribution's transformation
# and then inverting the transformation:
x_unconstrained = inference.transformations[x] # transformed prior
x_transform = x_unconstrained.bijector # transformed prior's transformation
qx_constrained = ed.transform(qx, bijectors.Invert(x_transform))
# The set of transformations is given by
# `inference.transformations`, which is a dictionary with keys
# given by any constrained latent variables and values given by their
# transformed distribution. We use the
# [`bijectors`](https://www.tensorflow.org/versions/master/api_docs/python/tf/distributions/bijectors)
# module in `tf.distributions` in order to handle invertible
# transformations.
# `qx_unconstrained` is a random variable distributed
# according to a inverse-transformed (constrained) normal distribution.
# For example, if the automated transformation from non-negative to
# reals is $\log$, then the constrained approximation is a LogNormal
# distribution; here, the default transformation is the inverse of
# $\textrm{softplus}$.
#
# We can visualize the densities of the distributions.
# The figure below shows that the inverse-transformed normal
# distribution has lighter tails than the Gamma but is overall a
# good fit.
plt.figure(figsize=(10, 6))
sns.distplot(x.sample(50000).eval(), hist=False, label='x')
sns.distplot(qx_constrained.sample(100000).eval(), hist=False, label='qx')
plt.show()
# __Gradient-based Monte Carlo.__
# Consider an Empirical approximation with 1000 samples
# and use the algorithm [`ed.HMC`](http://edwardlib.org/api/ed/HMC).
# +
qx = Empirical(params=tf.Variable(tf.random_normal([1000])))
inference = ed.HMC({x: qx})
inference.run(step_size=0.8)
# -
# Gamma and Empirical have unequal support so Gamma is transformed to
# the unconstrained space; by implementation, discrete delta
# distributions such as Empirical and PointMass are not transformed.
# `ed.HMC` then simulates
# Hamiltonian dynamics and writes the
# unconstrained samples to the empirical distribution.
#
# In order to obtain the approximation on the original (constrained)
# support, we again take the inverse of the target distribution's
# transformation.
x_unconstrained = inference.transformations[x] # transformed prior
x_transform = x_unconstrained.bijector # transformed prior's transformation
qx_constrained = Empirical(params=x_transform.inverse(qx.params))
# Unlike variational inference, we don't use `ed.transform` to
# obtain the constrained approximation, as it only applies to continuous
# distributions. Instead, we define a new Empirical distribution whose
# parameters (samples) are given by transforming all samples stored in
# the unconstrained approximation.
#
# We visualize the densities of the distributions.
# The figure below indicates that the samples accurately fit the Gamma
# distribution up to simulation error.
plt.figure(figsize=(10, 6))
sns.distplot(x.sample(50000).eval(), hist=False, label='x')
sns.distplot(qx_constrained.sample(100000).eval(), hist=False, label='qx')
plt.show()
# ## Acknowledgements & Remarks
#
# Automated transformations have largely been popularized by Stan
# for Hamiltonian Monte Carlo (Carpenter et al., 2016).
# This design is inspired by Stan's. However, a key distinction is that Edward
# provides users the ability to wield transformations and more flexibly
# manipulate results in both the original (constrained) and inferred
# (unconstrained) space.
#
# Automated transformations are also core to the algorithm automatic
# differentiation variational inference (Kucukelbir et al., 2017),
# which allows it to select a default variational family of normal
# distributions. However, note the automated transformation from
# non-negative to reals in Edward is not $\log$, which is used in Stan;
# rather, Edward uses $\textrm{softplus}$ which is more numerically
# stable (see also Kucukelbir et al. (2017, Fig. 9)).
#
# Finally, note that not all inference algorithms use or even need
# automated transformations.
# [`ed.Gibbs`](http://edwardlib.org/api/ed/Gibbs), moment
# matching with EP using Edward's conjugacy, and
# [`ed.KLqp`](http://edwardlib.org/api/ed/KLqp)
# with
# score function gradients all perform inference on the original latent
# variable space.
# Point estimation such as [`ed.MAP`](http://edwardlib.org/api/ed/MAP) also
# use the original latent variable space and only requires a
# constrained transformation on unconstrained free parameters.
# Model parameter estimation such as
# [`ed.GANInference`](http://edwardlib.org/api/ed/GANInference) do not even
# perform inference over latent variables.
| notebooks/automated_transformations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# +
# Dependencies and Setup
import pandas as pd
# File to Load
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame
purchase_data = pd.read_csv(file_to_load)
purchase_data.head()
# -
# ## Player Count
# * Display the total number of players
#
# Display the total number of players
player_count_df = pd.DataFrame({"Player Count": [len(purchase_data["SN"].unique())]})
player_count_df
# ## Purchasing Analysis (Total)
# + active=""
# * Run basic calculations to obtain number of unique items, average price, etc.
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
#
# +
# Basic calculations for analysis
unique_items = len(purchase_data["Item ID"].unique())
average_price = "${:,.2f}".format(purchase_data["Price"].mean())
total_purchases = len(purchase_data["Purchase ID"])
total_revenue = "${:,.2f}".format(purchase_data["Price"].sum())
# Create the summary data frame
analysis_df = pd.DataFrame({"Number of Unique Items": [unique_items],
"Average Price": average_price,
"Number of Purchases": total_purchases,
"Total Revenue": total_revenue})
# Display the summary data frame
analysis_df
# -
# ## Gender Demographics
# * Percentage and Count of Male Players
#
#
# * Percentage and Count of Female Players
#
#
# * Percentage and Count of Other / Non-Disclosed
#
#
#
# +
# Basic calculations to determine demographics
player_count = len(purchase_data["SN"].unique())
gender_demo = purchase_data.groupby("Gender")
gender_count = gender_demo.nunique()["SN"]
players_percentage = gender_count / player_count * 100
# Create the data frame and format
demographics_df = pd.DataFrame({"Total Count": gender_count, "Percentage of Players": players_percentage})
demographics_df.index.name = None
demographics_df.sort_values(["Total Count"], ascending = False).style.format({"Percentage of Players":"{:.2f}%"})
# -
#
# ## Purchasing Analysis (Gender)
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
#
#
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
# Basic calculations for analysis
purchase_count = gender_demo["Purchase ID"].count()
avg_price = gender_demo["Price"].mean()
avg_total = gender_demo["Price"].sum()
avg_per_person = avg_total/gender_count
# Create the summary data frame
purchasing_analysis_df = pd.DataFrame({"Purchase Count": purchase_count,
"Average Purchase Price": avg_price,
"Average Purchase Value": avg_total,
"Avg Purchase Total per Person": avg_per_person})
# Give displayed data cleaner formatting
purchasing_analysis_df.style.format({"Average Purchase Price":"${:.2f}",
"Average Purchase Value":"${:,.2f}",
"Avg Purchase Total per Person":"${:.2f}"})
# -
# ## Age Demographics
# * Establish bins for ages
#
#
# * Categorize the existing players using the age bins. Hint: use pd.cut()
#
#
# * Calculate the numbers and percentages by age group
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: round the percentage column to two decimal points
#
#
# * Display Age Demographics Table
#
# +
# Establish bins
bins = [0, 9, 14, 19, 24, 29, 34, 39, 50]
bin_names = ["<10", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", "40+"]
purchase_data["Age Group"] = pd.cut(purchase_data["Age"], bins, labels=bin_names)
# Calculations for data frame
age_group = purchase_data.groupby("Age Group")
total_age_count = age_group["SN"].nunique()
percentage_by_age = (total_age_count/player_count) * 100
# Create the summary data frame and format
age_demographics_df = pd.DataFrame({"Total Count": total_age_count, "Percentage of Players": percentage_by_age})
age_demographics_df.index.name = None
age_demographics_df.style.format({"Percentage of Players":"{:.2f}%"})
# -
# ## Purchasing Analysis (Age)
# * Bin the purchase_data data frame by age
#
#
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
# Calculations for data frame
purchases_by_age = age_group["Purchase ID"].count()
avg_purchase_price = age_group["Price"].mean()
total_purchase_value = age_group["Price"].sum()
avg_total_per_person = total_purchase_value/total_age_count
# Create the summary data frame
age_analysis_df = pd.DataFrame({"Purchase Count": purchases_by_age,
"Average Purchase Price": avg_purchase_price,
"Total Purchase Value":total_purchase_value,
"Average Purchase Total per Person": avg_total_per_person})
# Formatting for data frame
age_analysis_df.index.name = "Age Ranges"
age_analysis_df.style.format({"Average Purchase Price":"${:,.2f}",
"Total Purchase Value":"${:,.2f}",
"Average Purchase Total per Person":"${:,.2f}"})
# -
# ## Top Spenders
# * Run basic calculations to obtain the results in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the total purchase value column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
# Calculations for data frame
top_spenders = purchase_data.groupby("SN")
purchases_by_sn = top_spenders["Purchase ID"].count()
avg_purchases_by_sn = top_spenders["Price"].mean()
purchase_total_by_sn = top_spenders["Price"].sum()
# Create the summary data frame
top_spenders_df = pd.DataFrame({"Purchase Count": purchases_by_sn,
"Average Purchase Price": avg_purchases_by_sn,
"Total Purchase Value": purchase_total_by_sn})
# Formatting for data frame
format_top_spenders = top_spenders_df.sort_values(["Total Purchase Value"], ascending=False).head()
format_top_spenders.style.format({"Average Purchase Price":"${:.2f}",
"Total Purchase Value":"${:.2f}"})
# -
# ## Most Popular Items
# * Retrieve the Item ID, Item Name, and Item Price columns
#
#
# * Group by Item ID and Item Name. Perform calculations to obtain purchase count, average item price, and total purchase value
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the purchase count column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
# Retrieve the Item ID, Item Name, and Item Price columns
items = purchase_data[["Item ID", "Item Name", "Price"]]
# Group by Item ID and Item Name
item_data = items.groupby(["Item ID", "Item Name"])
# Calculations for data frame
item_purchases = item_data["Price"].count()
purchase_value = item_data["Price"].sum()
item_price = purchase_value/item_purchases
# Create the summary data frame
most_popular_df = pd.DataFrame({"Purchase Count": item_purchases,
"Item Price": item_price,
"Total Purchase Value": purchase_value})
# Formatting for data frame
most_popular_format = most_popular_df.sort_values(["Purchase Count"], ascending=False).head()
most_popular_format.style.format({"Item Price":"${:.2f}",
"Total Purchase Value":"${:.2f}"})
# -
# ## Most Profitable Items
# * Sort the above table by total purchase value in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the data frame
#
#
# +
# Re-format previous data frame for Total Purchase Value
most_profitable_format = most_popular_df.sort_values(["Total Purchase Value"], ascending=False).head()
most_profitable_format.style.format({"Item Price":"${:.2f}",
"Total Purchase Value":"${:.2f}"})
# -
| HeroesOfPymoli/.ipynb_checkpoints/HeroesOfPymoli_starter-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# # Preparation of databases before running SeroBA
# In order to use SeroBA for serotyping we must first download and prepare the necessary databases. Start by moving into the data directory:
cd data
# Now download the database from the GitHub repository:
svn checkout "https://github.com/sanger-pathog\
ens/seroba/trunk/database"
# **NOTE** if you are running a version of SeroBA older than v.0.1.3 the database is not packaged with the program and you will have to download it using the below command instead:
#
# seroba getPneumocat database_dir
# KMC is used by SeroBA to count k-mers and ARIBA is used to avoid the need for reads to be mapped to all reference sequences. Both of these require a database to be set up.
#
# To create a database for KMC and ARIBA run **createDBs**:
#
# seroba createDBs database/ kmer_size
#
# Where the options are:
#
# database The database directory which you just downloaded
# kmer_size The k-mer size you want to use for kmc. Recommended = 71
#
# SeroBA uses a default k-mer size of 71 for a read length of 250 bp. When deciding on a k-mer size, it is worth knowing that while a smaller k-mer size can keep the memory requirements low, it will also reduce the specificity. On the other hand, a larger k-mer size will require a larger amount of memory but will produce more unique k-mers and thus increase the specificity. What k-mer size to use also depends on the read length.
seroba createDBs database/ 71
# If you are working with SeroBA on the Sanger farm, the database with k-mer size 71 is already available centrally. This means you do not need to create the database for using SeroBA on the Sanger farm.
#
# However, for the sake of this tutorial, the above steps need to be compleated before you can continue with the tutorial.
#
# In the [next section](run_seroba.ipynb) we are going to run SeroBA to determine the serotype of one sample. You can also [return to the index](index.ipynb) or revisit the [previous section](serotyping.ipynb).
| SEROBA/db_setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Teste de Tukey
import numpy as np
from scipy.stats import f
from scipy.stats import f_oneway
grupo_a = np.array([165,152,143,140,155])
grupo_b = np.array([130,169,164,143,154])
grupo_c = np.array([163,158,154,149,156])
f.ppf(1-0.05, dfn=2, dfd=12)
_, p = f_oneway(grupo_a, grupo_b, grupo_c)
p
alpha = 0.05
if p < alpha:
print('Hipóteses nula rejeitada.')
else:
print('Hipóteses alternativa rejeitada.')
# #### Teste de Tukey
import pandas as pd
from statsmodels.stats.multicomp import MultiComparison
dados = {
'valores': [165,152,143,140,155,130,169,164,143,154,163,158,154,149,156],
'grupo': ['A','A','A','A','A','B','B','B','B','B','C','C','C','C','C']
}
dataset = pd.DataFrame(dados)
dataset.head(5)
comparar_grupo = MultiComparison(dataset['valores'], dataset['grupo'])
teste = comparar_grupo.tukeyhsd()
print(teste)
teste.plot_simultaneous();
dados2 = {
'valores': [70,90,80,50,20,130,169,164,143,154,163,158,154,149,156],
'grupo': ['A','A','A','A','A','B','B','B','B','B','C','C','C','C','C']
}
dataset2 = pd.DataFrame(dados2)
comparar_grupo2 = MultiComparison(dataset2['valores'], dataset2['grupo'])
teste2 = comparar_grupo2.tukeyhsd()
print(teste2)
teste2.plot_simultaneous();
| 8_intervalo_confianca/teste_ukey.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import networkx as nx
import numpy as np
import warnings
import datetime as dt
from scipy import stats
import scipy as sp
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rc('axes', axisbelow=True)
# -
import pandas as pd
import matplotlib
from matplotlib import gridspec
cases = pd.read_csv('../data/1581551273.37747wuhan_cases.csv')
cases = cases.rename(columns={cases.columns[0]: "date"})
provinces = cases.columns[1:] # ADDED
melted_cases = pd.melt(cases, id_vars='date',
value_vars=provinces, var_name='province',
value_name='cases')
full_data = melted_cases.copy() # ADDED
total_dates = np.unique(list(full_data.date))
total_cases_prov = {pr:sum(np.nan_to_num(list(full_data.loc[full_data.province==pr,].cases))) for pr in provinces}
final_cases_prov = dict(zip(list(full_data.loc[full_data.date==total_dates[-1],].province),
list(full_data.loc[full_data.date==total_dates[-1],].cases)))
sort_final_cases = {k: v for k, v in sorted(final_cases_prov.items(), key=lambda item: item[1])}
top_8 = {'Chongqing': 311.0,
'Anhui': 415.0,
'Jiangxi': 465.0,
'Henan': 583.0,
'Hunan': 589.0,
'Zhejiang': 706.0,
'Guangdong': 844.0,
'Hubei':6998.0}
# +
cols = dict(zip(list(top_8.keys()),plt.cm.Dark2(np.linspace(1,0,8))))
csum = 0
pad = 200
fig, ax = plt.subplots(1,1,dpi=150, figsize=(6,6))
num_success = 0
totals = 0
for i,pr in enumerate(list(top_8.keys())):
tot_cases_i = sum(np.nan_to_num(list(full_data.loc[full_data.province==pr,].cases)))
totals = totals + tot_cases_i
max_cases_i = top_8[pr]
xvals = total_dates[:-3]
slice_dat = full_data.loc[full_data.province==pr,]
slice_dat = slice_dat.loc[slice_dat.date<='2020-02-01',]
stdvs = np.array(list(slice_dat.cases))
yvals = np.array([csum+pad+np.nan_to_num(max_cases_i)]*len(xvals))#[:-3]
ax.plot(xvals, yvals-stdvs, alpha=0.7, color=cols[pr],zorder=20)
ax.plot(xvals, yvals+stdvs, alpha=0.7, color=cols[pr],zorder=20)
ax.fill_between(xvals, yvals-stdvs, yvals+stdvs, alpha=1.0, color='w',zorder=20)
ax.fill_between(xvals, yvals-stdvs, yvals+stdvs, alpha=0.7, color=cols[pr],zorder=20)
ax.text(min(xvals[~np.isnan(stdvs)]), yvals[0]+pad, pr,
color='#333333', fontsize=9)
ax.text(min(xvals[~np.isnan(stdvs)]), yvals[0]+pad, pr,
color=cols[pr],alpha=0.6, fontsize=9)
maxx = np.nan_to_num(yvals+stdvs)
csum = max(maxx)
num_success += 1
#ax.fill_between(xvals[43:46], 0, csum+150, alpha=1, color='w',zorder=1)
ax.fill_between(xvals[43:46], 0, csum+150, alpha=0.3, color='#999999',zorder=1, linewidth=0)
ax.fill_between(xvals[47:50], 0, csum+150, alpha=0.3, color='crimson',zorder=1, linewidth=0)
ax.set_xticks(total_dates[:-3][::4])
ax.set_ylim(150, csum+150)
ax.text(xvals[43], max(yvals+stdvs/1.25), 'Wuhan\nquarantine', horizontalalignment='right',
color='#333333', fontsize=10)
ax.text(xvals[47], max(yvals+stdvs/2.25), 'incubation\nperiod', horizontalalignment='right',
color='crimson', fontsize=9)
ax.set_xlim(total_dates[:-3][0], total_dates[:-3][-1])
ax.set_xticklabels([i[-5:] for i in list(total_dates[:-3][::4])], rotation=45, fontsize=7,y=0.01, rotation_mode ="anchor", horizontalalignment="right")
ax.set_yticks([])
ax.spines['top'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_visible(False)
ax.grid(linewidth=0.3, color='#999999', alpha=0.5)
ax.tick_params(axis='x', which='both', bottom=False,labelbottom=True)
plt.savefig('nCovCases1.png',dpi=425,bbox_inches='tight')
plt.savefig('nCovCases1.pdf',bbox_inches='tight')
plt.show()
# +
cols = dict(zip(list(top_8.keys()),plt.cm.Dark2(np.linspace(1,0,8))))
csum = 0
pad = 200
fig, ax = plt.subplots(1,1,dpi=150, figsize=(6,6))
num_success = 0
for i,pr in enumerate(list(top_8.keys())):
tot_cases_i = sum(np.nan_to_num(list(full_data.loc[full_data.province==pr,].cases)))
max_cases_i = top_8[pr]
xvals = total_dates[:-3]
slice_dat = full_data.loc[full_data.province==pr,]
slice_dat = slice_dat.loc[slice_dat.date<='2020-02-01',]
stdvs = np.array(list(slice_dat.cases))
yvals = np.array([csum+pad+np.nan_to_num(max_cases_i)]*len(xvals))#[:-3]
ax.plot(xvals, yvals-stdvs, alpha=0.7, color=cols[pr],zorder=20)
ax.plot(xvals, yvals+stdvs, alpha=0.7, color=cols[pr],zorder=20)
ax.fill_between(xvals, yvals-stdvs, yvals+stdvs, alpha=1.0, color='w',zorder=20)
ax.fill_between(xvals, yvals-stdvs, yvals+stdvs, alpha=0.7, color=cols[pr],zorder=20)
ax.text(min(xvals[~np.isnan(stdvs)]), yvals[0]+pad, pr+' (%i cumulative cases)'%max_cases_i,
color='#333333', fontsize=8)
ax.text(min(xvals[~np.isnan(stdvs)]), yvals[0]+pad, pr+' (%i cumulative cases)'%max_cases_i,
color=cols[pr],alpha=0.6, fontsize=8)
maxx = np.nan_to_num(yvals+stdvs)
csum = max(maxx)
num_success += 1
ax.fill_between(xvals[44:45], 0, csum+150, alpha=1, color='w',zorder=1)
ax.fill_between(xvals[44:45], 0, csum+150, alpha=0.3, color='#999999',zorder=1, linewidth=0)
ax.fill_between(xvals[48:50], 0, csum+150, alpha=0.3, color='crimson',zorder=1, linewidth=0)
ax.set_xticks(total_dates[:-3][::2])
ax.set_ylim(150, csum+150)
ax.text(xvals[44], max(yvals+stdvs/1.25), 'Wuhan\nquarantine', horizontalalignment='right',
color='#333333', fontsize=10)
ax.text(xvals[48], max(yvals+stdvs/2.25), 'incubation\nperiod', horizontalalignment='right',
color='crimson', fontsize=9)
ax.set_xlim(total_dates[:-3][0], total_dates[:-3][-1])
ax.set_xticklabels([i[-5:] for i in list(total_dates[:-3][::2])], rotation=0, fontsize=7,y=0.01)
ax.set_yticks([])
ax.spines['top'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_visible(False)
ax.grid(linewidth=0.3, color='#999999', alpha=0.5)
ax.tick_params(axis='x', which='both', bottom=False,labelbottom=True)
plt.savefig('nCovCases2.png',dpi=425,bbox_inches='tight')
plt.savefig('nCovCases2.pdf',bbox_inches='tight')
plt.show()
| code/Figure1a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# # Using SoS with spyder
# [spyder](https://pythonhosted.org/spyder/) is a Python GUI that works well with ipython, and therefore SoS if you configure it properly. Spyder should be readily available if you use Anaconda python, or you can install spyder using
#
# ```
# $ conda install spyder
# ```
# if you are using anaconda python distribution,
#
# ```
# $ pip install spyder
# ```
#
# for regular python, or according to instructions on the [spyder website](https://pythonhosted.org/spyder/) if you have a more complex Python environment.
#
# Because spyder does not support third-party kernels and does not yet support `.sos` files, sos provides a script to monkey-patch Spyder to let it accept files with `.sos` extension, and accept kernel `sos` as its default kernel. Before using Spyder with sos, you should execute command
!sos patch-spyder
# to patch spyder. After spyder is successfully patched, you can start spyder with a SoS kernel using command
#
# ```
# $ spyder --kernel sos
# ```
#
# Alternatively, you can connect spyder to an existing qtconsole with sos kernel by
#
# * Start a qtconsole with sos kernel using command `jupyter qtconsole --kernel sos`
# * Record connection file id from command line, or by running `%connect_info` from the console if you cannot find it.
# * Start `spyder`, select `consoles` -> `Connect an existing kernel`. Put connection id and connect.
#
# It is certainly possible to set up a remote Jupyter server and connect to a remote SoS kernel but this usage is beyond the scope of this tutorial.
#
# <font color="red">NOTE:</font> Monkey-patching a rapid-evolving project such as Spyder is difficult. Please [let us know](https://github.com/vatlab/SOS/issues) if the script fails to patch certain version of spyder or if Spyder does not work well after the patch.
# ### Open and edit files, the `%edit` magic
#
# You can open and edit files using Spyder's menu system. However, it can be more convenient to open files from within the SoS console. The SoS kernel provides a magic called `%edit` to do this. It supports string interpolation so you can run
#
# ```
# # %edit ${sdir}/myscript.sos
# ```
#
# if `sdir` is a variable pointing to your script directory. This magic also provides an option `--cd` that changes the current working directory to the script directory so
#
# ```
# # %edit --cd ${sdir}/myscript.sos
# ```
# would be equivalent to
#
# ```
# # %edit ${sdir}/myscript.sos
# # %cd ${sdir}
# ```
# ### Executing by line or by cell
#
# Whereas in batch SoS scripts statements are divided into **steps** using section headers (`[section: options]`), it uses cell magic `%cell` to define cells corresponding to cells of Jupyter notebooks. Basically, whenever you would like to define a new cell or separate an existing cell into two cells, you simple insert
#
# ```
# # %cell
# ```
#
# as the beginning of the new cell. Removing a cell or merging two existing cells can be easily achieved by removing a `%cell` line. Although a cell usually does not contain any step, it can contain multiple steps even multiple workflows. When you submit a line, a section (of multiple lines), or a cell to SoS, it execute the input as a complete workflow and returns the result of the last expression.
#
# Spyder provides shortcuts for executing current line, selection, or cell. It is highly configurable so you can define your own shortcuts for
#
# * run current line or selection (default to `F9`, which is awkward on my mac so I redefined it to **Ctrl-Enter**)
# * run current cell (**Cmd-Enter**)
# * execute current cell and move to next (**Shift-Enter**, the same as Jupter)
# ## Object inspector and Variable explorer
#
# Object inspector is useful for getting help message (e.g. definition of a function) of object. All you need to do is to navigate to the beginning of the work and press `Cmd-i` (Mac OSX).
#
# Variable explorer lists all variables in the SoS namespace. This saves your effort of using magic `%dict`, or `%preview vars` because all variables are right in the window and can be viewed easily.
| misc/spyder/Using_SoS_with_Spyder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
times = np.linspace(-0.1, 0.1, 300)
24 * 60 * (times[1] - times[0])
# +
from fleck import Star, generate_spots
from batman import TransitParams, TransitModel
import astropy.units as u
planet = TransitParams()
planet.per = 5 #88
planet.a = 15 #float(0.387*u.AU / u.R_sun)
planet.rp = 0.1
planet.w = 90
planet.ecc = 0
planet.inc = 90
planet.t0 = 0
planet.limb_dark = 'quadratic'
planet.u = [0.5079, 0.2239]
transit_model = TransitModel(planet, times).light_curve(planet)
# -
star = Star(spot_contrast=0.7, u_ld=planet.u, rotation_period=26)
koi_stdevs = np.load('../data/oot_scatter.npy')
# +
from tqdm import tqdm_notebook
n_transits = 10000
transits = []
residuals = []
spots_occulted = []
for i in range(n_transits):
b = np.random.rand()
planet.inc = np.degrees(np.arccos(b/planet.a))
spot_lons, spot_lats, spot_radii, inc_stellar = generate_spots(-90, 90, 0.04, 28, inclinations=90*u.deg)
lc, so = star.light_curve(spot_lons, spot_lats, spot_radii,
inc_stellar, planet=planet, times=times,
return_spots_occulted=True, fast=True)
lc += koi_stdevs[np.random.randint(0, len(koi_stdevs))] * np.random.randn(len(lc))[:, np.newaxis]
transits.append(lc)
lc = lc[:, 0] - transit_model + 1e-4 * np.random.randn(len(times))
fit = np.polyval(np.polyfit(times - times.mean(), lc, 3), times - times.mean())
residual = (lc - fit) / planet.rp**2
# transits.append(residuals)
spots_occulted.append(so)
residuals.append(residual)
print(spots_occulted[-1])
plt.plot(times, transits[-1])
# -
plt.plot(times, transits[np.random.randint(0, len(transits))])
np.save('data/simulated_transit_lcs.npy', np.hstack(transits).T)
np.save('data/simulated_spots_occulted.npy', spots_occulted)
np.count_nonzero(spots_occulted)/len(spots_occulted)
for transit in transits:
plt.plot(times, transit)
import os
os.system('say "done"')
| cnn/generate_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
#
# # `interp_sphgrid_MO_ETK`: An Einstein Toolkit Module for Interpolation to Spherical Grids
#
# ## Author: <NAME>
# ### Formatting improvements courtesy <NAME>
#
# ## This module is designed to interpolate arbitrary quantities on [Einstein Toolkit](https://einsteintoolkit.org/) Adaptive-Mesh Refinement (AMR) grids (using the [Carpet](https://carpetcode.org/) AMR infrastructure) to numerical grids with spherical sampling.
#
# **Notebook Status:** <font color='red'><b> In progress </b></font>
#
# **Validation Notes:** This module has not yet undergone validation testing.
#
# ## Introduction:
# Given some set of $N$ quantities $\mathbf{Q}=\{Q_0,Q_1,Q_2,...,Q_{N-2},Q_{N-1}\}$, this module performs the following for each $Q_i$:
#
# 1. Evaluate $Q_i$ at all gridpoints that are not ghost zones. Sometimes $Q_i$ is computed using finite difference derivatives, so this is necessary.
# 1. Call upon Carpet's interpolation and interprocessor synchronization functions to fill in $Q_i$ at all ghost zones, *except* at the outer boundary. We do not generally trust $Q_i$ at the outer boundary due to errors associated with the approximate outer boundary conditions.
# 1. At this point, $Q_i$ is set at all gridpoints except ghost zones at the outer boundary. Interpolate $Q_i$ to the spherical grids, **maintaining the Cartesian basis for all vectors and tensors**, and append the result to a file.
#
# This tutorial notebook takes a three-part structure. First, all the needed core Einstein Toolkit (ETK) C routines for interpolation are presented. Second, NRPy+ is used to output gridfunctions needed on the spherical grids. Third, the needed files for interfacing this module with the rest of the Einstein Toolkit (ccl files) are specified.
# <a id='toc'></a>
#
# # Table of Contents:
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#etkmodule): Setting up the Core C Code for the Einstein Toolkit Module
# 1. [Setp 1.a](#etk_interp): Low-Level Einstein Toolkit Interpolation Function
# 1. [Step 1.b](#sphericalgridnotes): Setting up the Spherical Grids
# 1. [Step 1.c](#fileformat): Outputting to File
# 1. [Step 1.d](#maininterpolator): The Main Interpolation Driver Function
# 1. [Step 2](#nrpy): Use NRPy+ C Output to Set All Output Gridfunctions
# 1. [Step 2.a](#nrpy_list_of_funcs_interp): Set up NRPy-based `list_of_functions_to_interpolate.h`
# 1. [Step 2.a.i](#nrpygrmhd): GRMHD quantities (***IN PROGRESS***)
# 1. [Step 2.a.ii](#nrpy4metric): Compute all 10 components of the 4-metric $g_{\mu\nu}$
# 1. [Step 2.a.iii](#nrpy4christoffels): Compute all 40 4-Christoffels $\Gamma^{\mu}_{\nu\delta}$
# 1. [Step 2.b](#nrpy_c_callingfunction): C code calling function for the NRPy+ C output
# 1. [Step 2.c](#nrpygetgfname): The `get_gf_name()` function
# 1. [Step 2.d](#nrpy_interp_counter): C Code for Initializing and incrementing `InterpCounter`
# 1. [Step 3](#cclfiles): Interfacing with the rest of the Einstein Toolkit; Setting up CCL files
# 1. [Step 3.a](#makecodedefn): `make.code.defn`
# 1. [Step 3.b](#interfaceccl): `interface.ccl`
# 1. [Step 3.c](#paramccl): `param.ccl`
# 1. [Step 3.d](#scheduleccl): `schedule.ccl`
# 1. [Step 4](#readingoutputfile): Python Script for Reading the Output File
# 1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='etkmodule'></a>
#
# # Step 1: Setting up the Core C Code for the Einstein Toolkit Module \[Back to [top](#toc)\]
# $$\label{etkmodule}$$
#
# First we set up the output directories for the ETK module:
# +
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
import shutil, os, sys, time # Standard Python modules for multiplatform OS-level functions, benchmarking
# Create C code output directory:
Ccodesdir = "interp_sphgrid_MO_ETK"
# First remove C code output directory and all subdirectories if they exist
# Courtesy https://stackoverflow.com/questions/303200/how-do-i-remove-delete-a-folder-that-is-not-empty
shutil.rmtree(Ccodesdir, ignore_errors=True)
# Then create a fresh directory
cmd.mkdir(Ccodesdir)
cmd.mkdir(os.path.join(Ccodesdir,"src/"))
# -
# <a id='etk_interp'></a>
#
# ## Step 1.a: Low-Level ETK Interpolation Function \[Back to [top](#toc)\]
# $$\label{etk_interp}$$
#
# We start by writing the low-level interpolation function **`Interpolate_to_sph_grid()`**, which to file.
#
# **`Interpolate_to_sph_grid()`** takes as input
# * **cctkGH**: Information about the underlying Cactus/Carpet grid hierarchy.
# * **interp_num_points**: Number of destination interpolation points
# * **point_x_temp, point_y_temp, point_z_temp**: Cartesian $(x,y,z)$ location for each of the **interp_num_points** interpolation points.
# * **input_array_names[1]**: List of input gridfunction names to interpolate. We will do this only one gridfunction at a time, for gridfunction $Q_i$, as described above.
#
# **`Interpolate_to_sph_grid()`** outputs:
# * **output_f[1]**: The gridfunction **input_array_names[1]** interpolated to the set of **interp_num_points** specified in the input.
# +
# %%writefile $Ccodesdir/src/Interpolate_to_sph_grid.h
void Interpolate_to_sph_grid(cGH *cctkGH,CCTK_INT interp_num_points, CCTK_INT interp_order,
CCTK_REAL *point_x_temp,CCTK_REAL *point_y_temp,CCTK_REAL *point_z_temp,
const CCTK_STRING input_array_names[1], CCTK_REAL *output_f[1]) {
DECLARE_CCTK_PARAMETERS;
CCTK_INT ierr;
const CCTK_INT NUM_INPUT_ARRAYS=1;
const CCTK_INT NUM_OUTPUT_ARRAYS=1;
CCTK_STRING coord_system = "cart3d";
// Set up handles
const CCTK_INT coord_system_handle = CCTK_CoordSystemHandle(coord_system);
if (coord_system_handle < 0) {
CCTK_VWarn(0, __LINE__, __FILE__, CCTK_THORNSTRING,
"can't get coordinate system handle for coordinate system \"%s\"!",
coord_system);
}
const CCTK_INT operator_handle = CCTK_InterpHandle(interpolator_name);
if (operator_handle < 0)
CCTK_VWarn(0, __LINE__, __FILE__, CCTK_THORNSTRING,
"couldn't find interpolator \"%s\"!",
interpolator_name);
char interp_order_string[10];
snprintf(interp_order_string, 10, "order=%d", interp_order);
CCTK_STRING interpolator_pars = interp_order_string;
CCTK_INT param_table_handle = Util_TableCreateFromString(interpolator_pars);
if (param_table_handle < 0) {
CCTK_VWarn(0, __LINE__, __FILE__, CCTK_THORNSTRING,
"bad interpolator parameter(s) \"%s\"!",
interpolator_pars);
}
CCTK_INT operand_indices[NUM_INPUT_ARRAYS]; //NUM_OUTPUT_ARRAYS + MAX_NUMBER_EXTRAS];
for(int i = 0 ; i < NUM_INPUT_ARRAYS ; i++) {
operand_indices[i] = i;
}
Util_TableSetIntArray(param_table_handle, NUM_OUTPUT_ARRAYS,
operand_indices, "operand_indices");
CCTK_INT operation_codes[NUM_INPUT_ARRAYS];
for(int i = 0 ; i < NUM_INPUT_ARRAYS ; i++) {
operation_codes[i] = 0;
}
Util_TableSetIntArray(param_table_handle, NUM_OUTPUT_ARRAYS,
operation_codes, "operation_codes");
const void* interp_coords[3]
= { (const void *) point_x_temp,
(const void *) point_y_temp,
(const void *) point_z_temp };
CCTK_INT input_array_indices[NUM_INPUT_ARRAYS];
for(int i = 0 ; i < NUM_INPUT_ARRAYS ; i++) {
input_array_indices[i] = CCTK_VarIndex(input_array_names[i]);
if(input_array_indices[i] < 0) {
CCTK_VWarn(0, __LINE__, __FILE__, CCTK_THORNSTRING,
"COULD NOT FIND VARIABLE '%s'.",
input_array_names[i]);
exit(1);
}
}
CCTK_INT output_array_types[NUM_OUTPUT_ARRAYS];
for(int i = 0 ; i < NUM_OUTPUT_ARRAYS ; i++) {
output_array_types[i] = CCTK_VARIABLE_REAL;
}
void * output_arrays[NUM_OUTPUT_ARRAYS]
= { (void *) output_f[0] };
// actual interpolation call
ierr = CCTK_InterpGridArrays(cctkGH,
3, // number of dimensions
operator_handle,
param_table_handle,
coord_system_handle,
interp_num_points,
CCTK_VARIABLE_REAL,
interp_coords,
NUM_INPUT_ARRAYS, // Number of input arrays
input_array_indices,
NUM_OUTPUT_ARRAYS, // Number of output arrays
output_array_types,
output_arrays);
if (ierr<0) {
CCTK_WARN(1,"interpolation screwed up");
Util_TableDestroy(param_table_handle);
exit(1);
}
ierr = Util_TableDestroy(param_table_handle);
if (ierr != 0) {
CCTK_WARN(1,"Could not destroy table");
exit(1);
}
}
# -
# <a id='sphericalgridnotes'></a>
#
# ## Step 1.b: Setting up the Spherical Grids \[Back to [top](#toc)\]
# $$\label{sphericalgridnotes}$$
#
# + By default, we set logarithmic radial coordinates: $r(x_{0,i}) = R_0 + e^{x_{0,i}}$, where
#
# + $x_{0,i} = x_{0, \mathrm{beg}} + \left(i+\frac{1}{2}\right) \Delta x_0$
# + $x_{0, {\mathrm{beg}}} = \log\left( R_{\mathrm{in}} - R_0 \right)$
# + $\Delta x_0 = \frac{1}{N_0}\log\left(\frac{R_\mathrm{out} - R_0}{R_\mathrm{in} - R_0}\right)$
#
#
# + As for the polar angle $\theta$, there are two options:
# + **Option 1**:
# $$ \theta(x_{1,j}) \, = \, \theta_c \, + \, \left( \pi - 2 \theta_c \right) x_{1,j} \, + \, \xi \, \sin\left(2 \pi x_{1,j} \right),$$
# where
# + $x_{1,j} = x_{1, \mathrm{beg}} + \left(j+\frac{1}{2}\right) \Delta x_1$
# + $\Delta x_1 = \frac{1}{N_1}$
#
# + **Option 2**:
# $$ \theta(x_{1,j}) = \frac{\pi}{2} \left[ 1 + \left(1-\xi \right) \left(2 x_{1,j} - 1 \right) + \left( \xi - \frac{2 \theta_c}{\pi} \right) \left( 2 x_{1,j} - 1 \right)^n \right],$$
# where
# + $n$ is odd
# + $x_{1,j} = x_{1, \mathrm{beg}} + \left(j+\frac{1}{2}\right) \Delta x_1$
# + $\Delta x_1 = \frac{1}{N_1}$
#
#
# + The azimuthal angle $\phi$ is uniform, so that $\phi(x_{2,k}) = x_{2,k}$:
#
# + $x_{2,k} \in [0,2\pi]$
# + $x_{2,k} = x_{2, \mathrm{beg}} + \left(k+\frac{1}{2}\right)\Delta x_{2}$
# + $\Delta x_{2} = \frac{ 2 \pi }{N_2}$
# +
# %%writefile $Ccodesdir/src/Set_up_interp_points_on_sph_grid.h
void sph_grid_Interpolate_many_pts__set_interp_pts(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
CCTK_REAL dx0 = log( (Rout - R0) / (Rin - R0) ) / ((CCTK_REAL)N0);
CCTK_REAL dx1 = 1.0 / ((CCTK_REAL)N1);
CCTK_REAL dx2 = 2.0*M_PI / ((CCTK_REAL)N2);
CCTK_REAL x0_beg = log( Rin - R0 );
CCTK_INT which_pt = 0;
for(CCTK_INT k=0;k<N2;k++) for(CCTK_INT j=0;j<N1;j++) for(CCTK_INT i=0;i<N0;i++) {
CCTK_REAL x0_i = x0_beg + ((CCTK_REAL)i + 0.5)*dx0;
CCTK_REAL rr = R0 + exp(x0_i);
CCTK_REAL x1_j = x1_beg + ((CCTK_REAL)j + 0.5)*dx1;
CCTK_REAL th = -1e300;
if(theta_option == 1) {
th = th_c + (M_PI - 2.0*th_c)*x1_j + xi*sin(2.0*M_PI*x1_j);
} else if (theta_option == 2) {
th = M_PI/2.0 * ( 1.0 + (1.0 - xi)*(2.0*x1_j - 1.0) + (xi - 2.0*th_c/M_PI)*pow(2.0*x1_j - 1.0 ,th_n) );
} else {
printf("Error: theta_option = %d NOT SUPPORTED.",theta_option);
exit(1);
}
CCTK_REAL x2_k = x2_beg + ((CCTK_REAL)k + 0.5)*dx2;
CCTK_REAL ph = x2_k;
points_x[which_pt] = rr*sin(th)*cos(ph);
points_y[which_pt] = rr*sin(th)*sin(ph);
points_z[which_pt] = rr*cos(th);
which_pt++;
}
}
# -
# <a id='fileformat'></a>
#
# ## Step 1.c: Outputting to File (File format notes) \[Back to [top](#toc)\]
# $$\label{fileformat}$$
#
# Since they take almost no space relative to the data chunks, we attach the entire metadata to each interpolated function that is output:
# +
# %%writefile $Ccodesdir/src/output_to_file.h
#include "define_NumInterpFunctions.h"
// output_to_file() starts order and InterpCounter both with the value 1
void output_to_file(CCTK_ARGUMENTS,char gf_name[100],int *order,CCTK_REAL *output_f[1]) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
char filename[100];
sprintf (filename, "%s/interp_sph_grids_MO.dat", out_dir);
FILE *file;
if(*InterpCounter == 1 && *order==1) {
file = fopen (filename,"w");
} else {
file = fopen (filename,"a+");
}
if (! file) {
CCTK_VWarn (1, __LINE__, __FILE__, CCTK_THORNSTRING,
"interp_sph_grid__ET_thorn: Cannot open output file '%s'", filename);
exit(1);
}
fwrite(gf_name, 100*sizeof(char), 1, file);
fwrite(order, sizeof(CCTK_INT), 1, file);
fwrite(&N0, sizeof(CCTK_INT), 1, file);
fwrite(&R0, sizeof(CCTK_REAL), 1, file);
fwrite(&Rin, sizeof(CCTK_REAL), 1, file);
fwrite(&Rout, sizeof(CCTK_REAL), 1, file);
fwrite(&N1, sizeof(CCTK_INT), 1, file);
fwrite(&x1_beg, sizeof(CCTK_REAL), 1, file);
fwrite(&theta_option, sizeof(CCTK_INT), 1, file);
fwrite(&th_c, sizeof(CCTK_REAL), 1, file);
fwrite(&xi, sizeof(CCTK_REAL), 1, file);
fwrite(&th_n, sizeof(CCTK_INT), 1, file);
fwrite(&N2, sizeof(CCTK_INT), 1, file);
fwrite(&x2_beg, sizeof(CCTK_REAL), 1, file);
CCTK_REAL magic_number = 1.130814081305130e-21;
fwrite(&magic_number, sizeof(CCTK_REAL), 1, file);
fwrite(&cctk_iteration, sizeof(CCTK_INT), 1, file);
fwrite(&cctk_time, sizeof(CCTK_REAL), 1, file);
for(CCTK_INT i=0;i<1;i++) {
fwrite(output_f[i], sizeof(CCTK_REAL)*N0*N1*N2, 1, file);
}
fclose(file);
}
# -
# <a id='maininterpolator'></a>
#
# ## Step 1.d: The Main Interpolation Driver Function \[Back to [top](#toc)\]
# $$\label{maininterpolator}$$
#
# The **`Interpolate_to_sph_grid_main_function()`** function calls the above functions as follows:
# 1. **`sph_grid_Interpolate_many_pts__set_interp_pts()`**: First set up the spherical grids
# 1. **`Interpolate_to_sph_grid()`**: Output
# +
# %%writefile $Ccodesdir/src/main_function.cc
// Include needed ETK & C library header files:
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <math.h>
// Needed for dealing with Cactus/ETK infrastructure
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
// Needed for low-level interpolation functions
#include "util_Table.h"
#include "util_String.h"
// Include locally-defined C++ functions:
#include "Set_up_interp_points_on_sph_grid.h"
#include "Interpolate_to_sph_grid.h"
#include "output_to_file.h"
#include "get_gf_name.h"
void Interpolate_to_sph_grid_main_function(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
// Perform interpolation only at iteration == interp_out_iteration:
if(cctk_iteration != interp_out_iteration) return;
// Set up spherically sampled interpolation grid arrays points_x,points_y,points_z:
sph_grid_Interpolate_many_pts__set_interp_pts(CCTK_PASS_CTOC);
// Set up output array:
CCTK_REAL *output_f[1];
output_f[0] = output_interped;
// The name of the input gridfunction is always "interp_sphgrid_MO_ETK::interped_gf":
const CCTK_STRING input_array_names[1] = { "interp_sphgrid_MO_ETK::interped_gf" };
// Perform interpolation!
for(int order=1; order <= 4; order *=2) {
char gf_name[100];
get_gf_name(*InterpCounter,gf_name);
printf("Interpolating\033[1m %s \033[0m... using interpolation order = %d\n",gf_name,order);
Interpolate_to_sph_grid(cctkGH, N0*N1*N2, order,
points_x,points_y,points_z, input_array_names, output_f);
if(CCTK_MyProc(cctkGH)==0) {
for(int i=0;i<N0*N1*N2;i++) {
if(output_f[0][i] > 1e20) {
printf("BAD POINT: %s %d %e %e %e %e\n",gf_name,i,points_x[i],points_y[i],points_z[i], output_f[0][i]);
}
}
output_to_file(CCTK_PASS_CTOC,gf_name,&order,output_f);
printf("Interpolate_to_sph_grid_main_function(): Just output to file at iteration %d\n",cctk_iteration);
} else {
printf("Interpolate_to_sph_grid_main_function(): Process !=0 waiting for file output at iteration %d\n",cctk_iteration);
}
}
}
# -
# <a id='nrpy'></a>
#
# # Step 2: Use NRPy+ C Output to Set All Output Gridfunctions \[Back to [top](#toc)\]
# $$ \label{nrpy}$$
# +
# Step 2: Import needed NRPy+ parameters
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import grid as gri # NRPy+: Functions having to do with numerical grids
import finite_difference as fin # NRPy+: Finite difference C code generation module
from outputC import lhrh # NRPy+: Core C code output module
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import NRPy_param_funcs as par # NRPy+: Parameter interface
import loop as lp # NRPy+: Generate C code loops
par.set_parval_from_str("grid::GridFuncMemAccess","ETK")
from collections import namedtuple
gf_interp = namedtuple('gf_interp', 'gf_description')
gf_interp_list = []
gf_interp_list.append(gf_interp("dummy -- used because this is a 1-offset array"))
interped_gf = gri.register_gridfunctions("AUX","interped_gf")
def interp_fileout(which_InterpCounter, expression, filename):
kernel = fin.FD_outputC("returnstring",lhrh(lhs=gri.gfaccess("out_gfs","interped_gf"),rhs=expression),"outCverbose=False")
output_type="a"
if which_InterpCounter == 1:
output_type="w"
with open(filename, output_type) as file:
file.write("if(*InterpCounter == "+str(which_InterpCounter)+") {\n")
file.write(lp.loop(["i2","i1","i0"],
["cctk_nghostzones[2]","cctk_nghostzones[1]","cctk_nghostzones[0]"],\
["cctk_lsh[2]-cctk_nghostzones[2]",
"cctk_lsh[1]-cctk_nghostzones[1]",
"cctk_lsh[0]-cctk_nghostzones[0]"],\
["1","1","1"],\
["#pragma omp parallel for","",""]," ",kernel))
file.write("}\n")
# If successful, return incremented which_InterpCounter:
return which_InterpCounter+1
# -
# <a id='nrpy_list_of_funcs_interp'></a>
#
# ## Step 2.a: Set up NRPy-based `list_of_functions_to_interpolate.h` \[Back to [top](#top)\]
# $$\label{nrpy_list_of_funcs_interp}$$
#
# First specify NRPy+ output file and initialize `which_InterpCounter`, which keeps track of the number of interpolated functions on the grid
# +
NRPyoutfilename = os.path.join(Ccodesdir,"src","list_of_functions_to_interpolate.h")
which_InterpCounter = 1
# -
# <a id='nrpygrmhd'></a>
#
# ### Step 2.a.i: GRMHD quantities (*IN PROGRESS; still working on adding vector potential*) \[Back to [top](#toc)\]
# $$\label{nrpygrmhd}$$
#
# These include
# * $\rho_b$, the baryonic density (i.e., the HydroBase variable $\verb|rho|$)
# * $P$, the total gas pressure (i.e., the HydroBase variable $\verb|press|$)
# * $\Gamma v_{(n)}^i$, the Valencia 3-velocity times the Lorentz factor (i.e., the HydroBase 3-gridfuntion $\verb|vel|$, multiplied by the Lorentz factor). This definition of velocity has the advantage that after interpolation, it will not violate $u^\mu u_\mu = -1$. In terms of the IllinoisGRMHD 3-velocity $v^i = u^i / u^0$, the Valencia 3-velocity is given by (Eq. 11 of [Etienne *et al*](https://arxiv.org/pdf/1501.07276.pdf)):
# $$
# v_{(n)}^i = \frac{1}{\alpha} \left(v^i + \beta^i\right).
# $$
# Further, $\Gamma = \alpha u^0$ is given by (as shown [here](Tutorial-u0_smallb_Poynting-Cartesian.ipynb)):
# $$
# \Gamma = \alpha u^0 = \sqrt{\frac{1}{1 - \gamma_{ij}v^i_{(n)}v^j_{(n)}}}.
# $$
# Therefore, $\Gamma v_{(n)}^i$ is given by
# $$
# \Gamma v_{(n)}^i = \frac{1}{\alpha} \left(v^i + \beta^i\right) \sqrt{\frac{1}{1 - \gamma_{ij}v^i_{(n)}v^j_{(n)}}}.
# $$
# * $A_i$, the *unstaggered* magnetic vector potential.
# * $B^i$, the *unstaggered* magnetic field vector (output only for validation purposes).
# +
# INPUT GRIDFUNCTIONS: The AUX or EVOL designation is *not* used in diagnostic modules.
gammaDD = ixp.register_gridfunctions_for_single_rank2("AUX","gammaDD", "sym01")
betaU = ixp.register_gridfunctions_for_single_rank1("AUX","betaU")
alpha = gri.register_gridfunctions("AUX","alpha")
DIM=3
gf_interp_list.append(gf_interp("IGM density primitive"))
rho_b = gri.register_gridfunctions("AUX","rho_b")
interp_expr = rho_b
which_InterpCounter = interp_fileout(which_InterpCounter,interp_expr,NRPyoutfilename)
gf_interp_list.append(gf_interp("IGM pressure primitive"))
P = gri.register_gridfunctions("AUX","P")
interp_expr = P
which_InterpCounter = interp_fileout(which_InterpCounter,interp_expr,NRPyoutfilename)
# -
# Next we implement:
# $$
# v_{(n)}^i = \frac{1}{\alpha} \left(v^i + \beta^i\right),
# $$
# and
# $$
# \Gamma v_{(n)}^i = \sqrt{\frac{1}{1 - \gamma_{ij}v^i_{(n)}v^j_{(n)}}} v_{(n)}^i.
# $$
# +
IGMvU = ixp.register_gridfunctions_for_single_rank1("AUX","IGMvU")
Valenciav = ixp.zerorank1()
for i in range(DIM):
Valenciav[i] = 1/alpha * (IGMvU[i] + betaU[i])
v_dot_v = sp.sympify(0)
for i in range(DIM):
for j in range(DIM):
v_dot_v += gammaDD[i][j]*Valenciav[i]*Valenciav[j]
Gamma_times_ValenciavU = ixp.zerorank1()
for i in range(DIM):
Gamma_times_ValenciavU[i] = sp.sqrt(1/(1 - v_dot_v))*Valenciav[i]
gf_interp_list.append(gf_interp("Lorentz factor, times Valencia vU"+str(i)))
interp_expr = Gamma_times_ValenciavU[i]
which_InterpCounter = interp_fileout(which_InterpCounter,interp_expr,NRPyoutfilename)
# For testing:
# gf_interp_list.append(gf_interp("Lorentz factor"))
# interp_expr = v_dot_v
# which_InterpCounter = interp_fileout(which_InterpCounter,interp_expr,NRPyoutfilename)
# for i in range(DIM):
# gf_interp_list.append(gf_interp("Valencia vU"+str(i)))
# interp_expr = Valenciav[i]
# which_InterpCounter = interp_fileout(which_InterpCounter,interp_expr,NRPyoutfilename)
BU = ixp.register_gridfunctions_for_single_rank1("AUX","BU")
for i in range(DIM):
gf_interp_list.append(gf_interp("IGM magnetic field component B"+str(i)))
interp_expr = BU[i]
which_InterpCounter = interp_fileout(which_InterpCounter,interp_expr,NRPyoutfilename)
# -
# <a id='nrpy4metric'></a>
#
# ### Step 2.a.ii: Compute all 10 components of the 4-metric $g_{\mu\nu}$ \[Back to [top](#toc)\]
# $$\label{nrpy4metric}$$
#
# We are given $\gamma_{ij}$, $\alpha$, and $\beta^i$ from ADMBase, and the 4-metric is given in terms of these quantities as
# $$
# g_{\mu\nu} = \begin{pmatrix}
# -\alpha^2 + \beta^k \beta_k & \beta_i \\
# \beta_j & \gamma_{ij}
# \end{pmatrix}.
# $$
# +
# Eq. 2.121 in B&S
betaD = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
betaD[i] += gammaDD[i][j]*betaU[j]
# Now compute the beta contraction.
beta2 = sp.sympify(0)
for i in range(DIM):
beta2 += betaU[i]*betaD[i]
# Eq. 2.122 in B&S
g4DD = ixp.zerorank2(DIM=4)
g4DD[0][0] = -alpha**2 + beta2
for i in range(DIM):
g4DD[i+1][0] = g4DD[0][i+1] = betaD[i]
for i in range(DIM):
for j in range(DIM):
g4DD[i+1][j+1] = gammaDD[i][j]
for mu in range(4):
for nu in range(mu,4):
gf_interp_list.append(gf_interp("4-metric component g4DD"+str(mu)+str(nu)))
interp_expr = g4DD[mu][nu]
which_InterpCounter = interp_fileout(which_InterpCounter,interp_expr,NRPyoutfilename)
# -
# <a id='nrpy4christoffels'></a>
#
# ### Step 2.a.iii: Compute all 40 4-Christoffels $\Gamma^{\mu}_{\nu\delta}$ \[Back to [top](#toc)\]
# $$\label{nrpy4christoffels}$$
#
# By definition,
# $$
# \Gamma^{\mu}_{\nu\delta} = \frac{1}{2} g^{\mu\eta} \left(g_{\eta\nu,\delta} + g_{\eta\delta,\nu} - g_{\nu\delta,\eta} \right)
# $$
#
# Recall that $g_{\mu\nu}$ is given from $\gamma_{ij}$, $\alpha$, and $\beta^i$ via
# $$
# g_{\mu\nu} = \begin{pmatrix}
# -\alpha^2 + \beta^k \beta_k & \beta_i \\
# \beta_j & \gamma_{ij}
# \end{pmatrix}.
# $$
#
# The derivatives $g_{\mu\nu,\eta}$ are then computed in terms of finite-difference derivatives of the input ADM gridfunctions $\gamma_{ij}$, $\alpha$, and $\beta^i$, **assuming that the 4-metric is static, so that $\partial_t g_{\mu\nu}=0$ for all $\mu$ and $\nu$**.
#
# To compute $g^{\mu\nu}$, we use the standard formula (Eq. 4.49 in [Gourgoulhon](https://arxiv.org/pdf/gr-qc/0703035.pdf)):
# $$
# g^{\mu\nu} = \begin{pmatrix}
# -\frac{1}{\alpha^2} & \frac{\beta^i}{\alpha^2} \\
# \frac{\beta^i}{\alpha^2} & \gamma^{ij} - \frac{\beta^i\beta^j}{\alpha^2}
# \end{pmatrix},
# $$
# where $\gamma^{ij}$ is given by the inverse of $\gamma_{ij}$.
# +
betaDdD = ixp.zerorank2()
gammaDD_dD = ixp.declarerank3("gammaDD_dD","sym01")
betaU_dD = ixp.declarerank2("betaU_dD","nosym")
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
# Recall that betaD[i] = gammaDD[i][j]*betaU[j] (Eq. 2.121 in B&S)
betaDdD[i][k] += gammaDD_dD[i][j][k]*betaU[j] + gammaDD[i][j]*betaU_dD[j][k]
# Eq. 2.122 in B&S
g4DDdD = ixp.zerorank3(DIM=4)
alpha_dD = ixp.declarerank1("alpha_dD")
for i in range(DIM):
# Recall that g4DD[0][0] = -alpha^2 + betaU[i]*betaD[i]
g4DDdD[0][0][i+1] += -2*alpha*alpha_dD[i]
for j in range(DIM):
g4DDdD[0][0][i+1] += betaU_dD[j][i]*betaD[j] + betaU[j]*betaDdD[j][i]
for i in range(DIM):
for j in range(DIM):
# Recall that g4DD[i][0] = g4DD[0][i] = betaD[i]
g4DDdD[i+1][0][j+1] = g4DDdD[0][i+1][j+1] = betaDdD[i][j]
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
# Recall that g4DD[i][j] = gammaDD[i][j]
g4DDdD[i+1][j+1][k+1] = gammaDD_dD[i][j][k]
gammaUU, dummyDET = ixp.symm_matrix_inverter3x3(gammaDD)
g4UU = ixp.zerorank2(DIM=4)
g4UU[0][0] = -1 / alpha**2
for i in range(DIM):
g4UU[0][i+1] = g4UU[i+1][0] = betaU[i]/alpha**2
for i in range(DIM):
for j in range(DIM):
g4UU[i+1][j+1] = gammaUU[i][j] - betaU[i]*betaU[j]/alpha**2
# -
# Again, we are to compute:
# $$
# \Gamma^{\mu}_{\nu\delta} = \frac{1}{2} g^{\mu\eta} \left(g_{\eta\nu,\delta} + g_{\eta\delta,\nu} - g_{\nu\delta,\eta} \right)
# $$
# +
Gamma4UDD = ixp.zerorank3(DIM=4)
for mu in range(4):
for nu in range(4):
for delta in range(4):
for eta in range(4):
Gamma4UDD[mu][nu][delta] += sp.Rational(1,2)*g4UU[mu][eta]*\
(g4DDdD[eta][nu][delta] + g4DDdD[eta][delta][nu] - g4DDdD[nu][delta][eta])
# Now output the 4-Christoffels to file:
for mu in range(4):
for nu in range(4):
for delta in range(nu,4):
gf_interp_list.append(gf_interp("4-Christoffel GammaUDD"+str(mu)+str(nu)+str(delta)))
interp_expr = Gamma4UDD[mu][nu][delta]
which_InterpCounter = interp_fileout(which_InterpCounter,interp_expr,NRPyoutfilename)
# -
# <a id='nrpy_c_callingfunction'></a>
#
# ## Step 2.b: C code calling function for the NRPy+ C output \[Back to [top](#toc)\]
# $$\label{nrpy_c_callingfunction}$$
#
# In the above blocks, we wrote and appended to a file `list_of_functions_to_interpolate.h`. Here we write the calling function for this C code.
# +
# %%writefile $Ccodesdir/src/construct_function_to_interpolate__store_to_interped_gf.cc
#include <stdio.h>
#include <stdlib.h>
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
// Set the gridfunction interped_gf, according to the interpolation counter variable interp_counter.
// For example, we might interpolate "IllinoisGRMHD::rho_b" if interp_counter==0. The following
// function takes care of these
void list_of_functions_to_interpolate(cGH *cctkGH,const CCTK_INT *cctk_lsh,const CCTK_INT *cctk_nghostzones,
const CCTK_REAL invdx0,const CCTK_REAL invdx1,const CCTK_REAL invdx2,
const CCTK_INT *InterpCounter,
const CCTK_REAL *rho_bGF,const CCTK_REAL *PGF,
const CCTK_REAL *IGMvU0GF,const CCTK_REAL *IGMvU1GF,const CCTK_REAL *IGMvU2GF,
const CCTK_REAL *BU0GF,const CCTK_REAL *BU1GF,const CCTK_REAL *BU2GF,
const CCTK_REAL *gammaDD00GF,const CCTK_REAL *gammaDD01GF,const CCTK_REAL *gammaDD02GF,
const CCTK_REAL *gammaDD11GF,const CCTK_REAL *gammaDD12GF,const CCTK_REAL *gammaDD22GF,
const CCTK_REAL *betaU0GF,const CCTK_REAL *betaU1GF,const CCTK_REAL *betaU2GF,
const CCTK_REAL *alphaGF, CCTK_REAL *interped_gfGF) {
#include "list_of_functions_to_interpolate.h"
}
void construct_function_to_interpolate__store_to_interped_gf(CCTK_ARGUMENTS) {
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
const CCTK_REAL invdx0 = 1.0 / CCTK_DELTA_SPACE(0);
const CCTK_REAL invdx1 = 1.0 / CCTK_DELTA_SPACE(1);
const CCTK_REAL invdx2 = 1.0 / CCTK_DELTA_SPACE(2);
list_of_functions_to_interpolate(cctkGH,cctk_lsh,cctk_nghostzones,invdx0,invdx1,invdx2,
InterpCounter,
rho_b,P,
vx,vy,vz,
Bx,By,Bz,
gxx,gxy,gxz,gyy,gyz,gzz,
betax,betay,betaz,alp, interped_gf);
// interped_gf will be interpolated across AMR boundaries, meaning that
// it must be prolongated. Only gridfunctions with 3 timelevels stored
// may be prolongated (provided time_interpolation_order is set to the
// usual value of 2). We should only call this interpolation routine
// at iterations in which all gridfunctions are on the same timelevel
// (usually a power of 2), which will ensure that the following
// "filling of the timelevels" is completely correct.
#pragma omp parallel for
for(int i=0;i<cctk_lsh[0]*cctk_lsh[1]*cctk_lsh[2];i++) {
interped_gf_p[i] = interped_gf[i];
interped_gf_p_p[i] = interped_gf[i];
}
}
# -
# <a id='nrpygetgfname'></a>
#
# ## Step 2.c: The `get_gf_name()` function \[Back to [top](#toc)\]
# $$\label{nrpygetgfname}$$
with open(os.path.join(Ccodesdir,"src","get_gf_name.h"), "w") as file:
file.write("void get_gf_name(const int InterpCounter,char gf_name[100]) {\n")
for i in range(1,which_InterpCounter):
file.write(" if(InterpCounter=="+str(i)+") { snprintf(gf_name,100,\""+gf_interp_list[i].gf_description+"\"); return; }\n")
file.write(" printf(\"Error. InterpCounter = %d unsupported. I should not be here.\\n\",InterpCounter); exit(1);\n")
file.write("}\n")
# <a id='nrpy_interp_counter'></a>
#
# ## Step 2.d: C Code for Initializing and incrementing "InterpCounter" \[Back to [top](#toc)\]
# $$\label{nrpy_interp_counter}$$
# The gridfunctions are interpolated one at a time based on the current value of the index quantity `InterpCounter`. Here we write the C code needed for initializing and incrementing this variable.
with open(os.path.join(Ccodesdir,"src","define_NumInterpFunctions.h"), "w") as file:
file.write("#define NumInterpFunctions "+str(which_InterpCounter)+"\n")
# +
# %%writefile $Ccodesdir/src/interp_counter.cc
#include <assert.h>
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <math.h>
#include <ctype.h>
#include "cctk.h"
#include "cctk_Arguments.h"
#include "cctk_Parameters.h"
#include "define_NumInterpFunctions.h"
void SphGrid_InitializeInterpCounterToZero(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
*InterpCounter = 0;
if(verbose==2) printf("interp_sphgrid_MO_ETK: Just set InterpCounter to %d\n",*InterpCounter);
}
void SphGrid_InitializeInterpCounter(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
if(cctk_iteration == interp_out_iteration) {
*InterpCounter = 1;
if(verbose==2) printf("interp_sphgrid_MO_ETK: Just set InterpCounter to %d ; ready to start looping over interpolated gridfunctions!\n",
*InterpCounter);
}
}
// This function increments InterpCounter if we are at the interp_out_iteration until
// it hits NumInterpFunctions. At this iteration, InterpCounter is set to zero, which
// exits the loop.
void SphGrid_IncrementInterpCounter(CCTK_ARGUMENTS)
{
DECLARE_CCTK_ARGUMENTS;
DECLARE_CCTK_PARAMETERS;
if(*InterpCounter == NumInterpFunctions-1) {
*InterpCounter = 0;
if(verbose==2) printf("interp_sphgrid_MO_ETK: Finished! Just zeroed InterpCounter.\n");
} else {
(*InterpCounter)++;
if(verbose==2) printf("interp_sphgrid_MO_ETK: Just incremented InterpCounter to %d of %d\n",*InterpCounter,NumInterpFunctions-1);
}
}
# -
# <a id='cclfiles'></a>
#
# # Step 3: Define how this module interacts and interfaces with the larger Einstein Toolkit infrastructure \[Back to [top](#toc)\]
# $$\label{cclfiles}$$
#
# Writing a module ("thorn") within the Einstein Toolkit requires that three "ccl" files be constructed, all in the root directory of the thorn:
#
# 1. `interface.ccl`: defines the gridfunction groups needed, and provides keywords denoting what this thorn provides and what it should inherit from other thorns.
# 1. `param.ccl`: specifies free parameters within the thorn.
# 1. `schedule.ccl`: allocates storage for gridfunctions, defines how the thorn's functions should be scheduled in a broader simulation, and specifies the regions of memory written to or read from gridfunctions.
# <a id='makecodedefn'></a>
#
# ## Step 3.a: `make.code.defn` \[Back to [top](#toc)\]
# $$\label{makecodedefn}$$
#
# Before writing the "ccl" files, we first add Einstein Toolkit's equivalent of a Makefile, the `make.code.defn` file:
# +
# %%writefile $Ccodesdir/src/make.code.defn
# Main make.code.defn file for thorn interp_sphgrid_MO_ETK
# Source files in this directory
SRCS = main_function.cc interp_counter.cc construct_function_to_interpolate__store_to_interped_gf.cc
# -
# <a id='interfaceccl'></a>
#
# ## Step 3.b: `interface.ccl` \[Back to [top](#toc)\]
# $$\label{interfaceccl}$$
#
# Let's now write `interface.ccl`. The [official Einstein Toolkit (Cactus) documentation](http://einsteintoolkit.org/usersguide/UsersGuide.html) defines what must/should be included in an `interface.ccl` file [**here**](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-178000D2.2).
# +
# %%writefile $Ccodesdir/interface.ccl
# With "implements", we give our thorn its unique name.
implements: interp_sphgrid_MO_ETK
# By "inheriting" other thorns, we tell the Toolkit that we
# will rely on variables/function that exist within those
# functions.
inherits: admbase IllinoisGRMHD Grid
# Tell the Toolkit that we want "interped_gf" and "InterpCounter"
# and invariants to NOT be visible to other thorns, by using
# the keyword "private". Note that declaring these
# gridfunctions here *does not* allocate memory for them;
# that is done by the schedule.ccl file.
private:
CCTK_REAL interpolation_gf type=GF timelevels=3 tags='Checkpoint="no"'
{
interped_gf
} "Gridfunction containing output from interpolation."
int InterpCounterVar type = SCALAR tags='checkpoint="no"'
{
InterpCounter
} "Counter that keeps track of which function we are interpolating."
CCTK_REAL interp_pointcoords_and_output_arrays TYPE=ARRAY DISTRIB=CONSTANT DIM=1 SIZE=N0*N1*N2 tags='checkpoint="no"'
{
points_x,points_y,points_z,
output_interped
}
# -
# <a id='paramccl'></a>
#
# ## Step 3.c: $\text{param.ccl}$ \[Back to [top](#toc)\]
# $$\label{paramccl}$$
#
# We will now write the file `param.ccl`. This file allows the listed parameters to be set at runtime. We also give allowed ranges and default values for each parameter. More information on this file's syntax can be found in the [official Einstein Toolkit documentation](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-183000D2.3).
# +
# %%writefile $Ccodesdir/param.ccl
# Output the interpolated data to the IO::out_dir directory:
shares: IO
USES STRING out_dir
restricted:
########################################
# BASIC THORN STEERING PARAMETERS
CCTK_INT interp_out_iteration "Which iteration to interpolate to spherical grids?" STEERABLE=ALWAYS
{
0:* :: ""
} 960000
## Interpolator information
CCTK_STRING interpolator_name "Which interpolator to use?" STEERABLE=ALWAYS
{
".+" :: "Any nonempty string; an unsupported value will throw an error."
} "Lagrange polynomial interpolation"
CCTK_INT verbose "Set verbosity level: 1=useful info; 2=moderately annoying (though useful for debugging)" STEERABLE=ALWAYS
{
0:2 :: "0 = no output; 1=useful info; 2=moderately annoying (though useful for debugging)"
} 2
########################################
# SPHERICAL COORDINATE SYSTEM PARAMETERS
CCTK_INT N0 "Number of points in r direction" STEERABLE=ALWAYS
{
0:* :: ""
} 96
CCTK_INT N1 "Number of points in theta direction" STEERABLE=ALWAYS
{
0:* :: ""
} 96
CCTK_INT N2 "Number of points in phi direction" STEERABLE=ALWAYS
{
0:* :: ""
} 96
##########
# Cartesian position of center of spherical grid (usually center of BH) -- CURRENTLY UNSUPPORTED!
CCTK_REAL x_center "x-position of center." STEERABLE=ALWAYS
{
0:* :: ""
} 0.0
CCTK_REAL y_center "y-position of center." STEERABLE=ALWAYS
{
0:* :: ""
} 0.0
CCTK_REAL z_center "z-position of center." STEERABLE=ALWAYS
{
0:* :: ""
} 0.0
##########
# Radial parameters:
CCTK_REAL R0 "Radial offset: r(x0) = R_0 + exp(x0). Probably should keep it set to zero." STEERABLE=ALWAYS
{
0:* :: ""
} 0.0
CCTK_REAL Rin "x0 offset: x0 = log(Rin-R0) + (i + 0.5)Dx0." STEERABLE=ALWAYS
{
0:* :: ""
} 1.08986052555408
CCTK_REAL Rout "Dx0 = log( (Rout-R0) / (Rin-R0) )/N0" STEERABLE=ALWAYS
{
0:* :: ""
} 80.0
##########
# Theta parameters:
CCTK_REAL x1_beg "x1 offset: x1 = x1_beg + (j + 0.5)Dx1. Probably should keep it set to zero." STEERABLE=ALWAYS
{
0:* :: ""
} 0.0
CCTK_INT theta_option "Which prescription for theta should be used? 1 or 2?" STEERABLE=ALWAYS
{
1:2 :: ""
} 1
CCTK_REAL th_c "theta_c: Angular cutout size for theta = 0 and pi" STEERABLE=ALWAYS
{
0:* :: ""
} 0.053407075111026485 # 0.017*pi
CCTK_REAL xi "Amplitude of nonlinear part of the theta distribution." STEERABLE=ALWAYS
{
0:* :: ""
} 0.25
CCTK_INT th_n "Power of nonlinear part of theta distribution. Only for theta_option=2" STEERABLE=ALWAYS
{
0:* :: ""
} 9
##########
# Phi parameters:
CCTK_REAL x2_beg "x2 offset: x2 = x2_beg + (k + 0.5)Dx2. Probably should keep it set to zero." STEERABLE=ALWAYS
{
0:* :: ""
} 0.0
########################################
# -
# <a id='scheduleccl'></a>
#
# ## Step 3.d: `schedule.ccl` \[Back to [top](#toc)\]
# $$\label{scheduleccl}$$
#
# Finally, we will write the file `schedule.ccl`; its official documentation is found [here](http://einsteintoolkit.org/usersguide/UsersGuidech12.html#x17-186000D2.4).
#
# This file declares storage for variables declared in the `interface.ccl` file and specifies when the various parts of the thorn will be run:
# +
# %%writefile $Ccodesdir/schedule.ccl
STORAGE: interpolation_gf[3]
STORAGE: InterpCounterVar
STORAGE: interp_pointcoords_and_output_arrays
#############################
SCHEDULE SphGrid_InitializeInterpCounterToZero AT CCTK_INITIAL
{
LANG: C
OPTIONS: GLOBAL
} "Initialize InterpCounter variable to zero"
SCHEDULE SphGrid_InitializeInterpCounterToZero AT CCTK_POST_RECOVER_VARIABLES
{
LANG: C
OPTIONS: GLOBAL
} "Initialize InterpCounter variable to zero"
SCHEDULE SphGrid_InitializeInterpCounter before SphGrid_InterpGroup AT CCTK_ANALYSIS
{
LANG: C
OPTIONS: GLOBAL
} "Initialize InterpCounter variable"
##################
SCHEDULE GROUP SphGrid_InterpGroup AT CCTK_ANALYSIS BEFORE CarpetLib_printtimestats BEFORE CarpetLib_printmemstats AFTER Convert_to_HydroBase WHILE interp_sphgrid_MO_ETK::InterpCounter
{
} "Perform all spherical interpolations. This group is only actually scheduled at cctk_iteration==interp_out_iteration."
SCHEDULE construct_function_to_interpolate__store_to_interped_gf in SphGrid_InterpGroup before DoSum
{
STORAGE: interpolation_gf[3],InterpCounterVar,interp_pointcoords_and_output_arrays
OPTIONS: GLOBAL,LOOP-LOCAL
SYNC: interpolation_gf
LANG: C
} "Construct the function to interpolate"
SCHEDULE Interpolate_to_sph_grid_main_function in SphGrid_InterpGroup after construct_function_to_interpolate__store_to_interped_gf
{
OPTIONS: GLOBAL
LANG: C
} "Perform interpolation and output result to file."
#######
SCHEDULE SphGrid_IncrementInterpCounter in SphGrid_InterpGroup after Interpolate_to_sph_grid_main_function
{
LANG: C
OPTIONS: GLOBAL
} "Increment InterpCounter variable, or set to zero once loop is complete."
##################
# -
# <a id='readingoutputfile'></a>
#
# # Step 4: Python Script for Reading the Output File \[Back to [top](#toc)\]
# $$\label{readingoutputfile}$$
#
# Here is a Python code for reading the output file generated by this thorn. It is based on a collection of Python scripts written by <NAME>, available [here](https://bitbucket.org/zach_etienne/nrpy/src/master/mhd_diagnostics/).
#
# After generating the output file `interp_sphgrid_MO_ETK.dat` using the Einstein Toolkit thorn above, this script will read in all the data. Processing can then be done by straightforward modification of this script. Save the script as "Interp_Sph_ReadIn.py", and run it using the command
#
# **`python Interp_Sph_ReadIn.py interp_sphgrid_MO_ETK.dat 58 outfile`**
#
# Currently the last parameter "outfile" is required but not used.
#
# ```python
# """
# interp_sphgrid_MO_ETK.dat File Reader. Compatible with Python 2.7+ and 3.6+ at least.
#
# <NAME>
#
# Based on Python scripts written by <NAME>:
# https://bitbucket.org/zach_etienne/nrpy/src/master/mhd_diagnostics/
#
# Find the latest version of this reader at the bottom of this Jupyter notebook:
# https://github.com/zachetienne/nrpytutorial/blob/master/Tutorial-ETK_thorn-Interpolation_to_Spherical_Grids.ipynb
#
# Usage instructions:
#
# From the command-line, run via:
# python Interp_Sph_ReadIn.py interp_sphgrid_MO_ETK.dat [number of gridfunctions (58 or so)] [outfile]
#
# Currently the last parameter "outfile" is required but not actually used.
# """
# import numpy as np
# import struct
# import sys
# import argparse
#
# parser = argparse.ArgumentParser(description='Read file.')
# parser.add_argument("datafile", help="main data file")
# parser.add_argument("number_of_gridfunctions", help="number of gridfunctions")
#
# parser.add_argument("outfileroot", help="root of output file names")
#
# args = parser.parse_args()
#
# datafile = args.datafile
# outfileroot = args.outfileroot
# number_of_gridfunctions = int(args.number_of_gridfunctions)
#
# print("reading from "+str(datafile))
#
# """
# read_char_array():
# Reads a character array of size="size"
# from a file (with file handle = "filehandle")
# and returns the character array as a proper
# Python string.
# """
# def read_char_array(filehandle,size):
# reached_end_of_string = False
# chartmp = struct.unpack(str(size)+'s', filehandle.read(size))[0]
#
# #https://docs.python.org/3/library/codecs.html#codecs.decode
# char_array_orig = chartmp.decode('utf-8',errors='ignore')
#
# char_array = ""
# for i in range(len(char_array_orig)):
# char = char_array_orig[i]
# # C strings end in '\0', which in Python-ese is '\x00'.
# # As characters read after the end of the string will
# # generally be gibberish, we no longer append
# # to the output string after '\0' is reached.
# if sys.version_info[0]==3 and bytes(char.encode('utf-8')) == b'\x00':
# reached_end_of_string = True
# elif sys.version_info[0]==2 and char == '\x00':
# reached_end_of_string = True
#
# if reached_end_of_string == False:
# char_array += char
# else:
# pass # Continue until we've read 'size' bytes
# return char_array
#
# """
# read_header()
# Reads the header from a file.
# """
# def read_header(filehandle):
# # This function makes extensive use of Python's struct.unpack
# # https://docs.python.org/3/library/struct.html
# # First store gridfunction name and interpolation order used:
# # fwrite(gf_name, 100*sizeof(char), 1, file);
# gf_name = read_char_array(filehandle,100)
# # fwrite(order, sizeof(CCTK_INT), 1, file);
# order = struct.unpack('i',filehandle.read(4))[0]
#
# # Then the radial grid parameters:
# # fwrite( & N0, sizeof(CCTK_INT), 1, file);
# N0 = struct.unpack('i',filehandle.read(4))[0]
# # fwrite( & R0, sizeof(CCTK_REAL), 1, file);
# R0 = struct.unpack('d',filehandle.read(8))[0]
# # fwrite( & Rin, sizeof(CCTK_REAL), 1, file);
# Rin = struct.unpack('d',filehandle.read(8))[0]
# # fwrite( & Rout, sizeof(CCTK_REAL), 1, file);
# Rout = struct.unpack('d',filehandle.read(8))[0]
#
# # Then the grid parameters related to the theta coordinate:
# # fwrite( & N1, sizeof(CCTK_INT), 1, file);
# N1 = struct.unpack('i', filehandle.read(4))[0]
# # fwrite( & x1_beg, sizeof(CCTK_REAL), 1, file);
# x1_beg = struct.unpack('d', filehandle.read(8))[0]
# # fwrite( & theta_option, sizeof(CCTK_INT), 1, file);
# theta_option = struct.unpack('i', filehandle.read(4))[0]
# # fwrite( & th_c, sizeof(CCTK_REAL), 1, file);
# th_c = struct.unpack('d', filehandle.read(8))[0]
# # fwrite( & xi, sizeof(CCTK_REAL), 1, file);
# xi = struct.unpack('d', filehandle.read(8))[0]
# # fwrite( & th_n, sizeof(CCTK_INT), 1, file);
# th_n = struct.unpack('i', filehandle.read(4))[0]
#
# # Then the grid parameters related to the phi coordinate:
# # fwrite( & N2, sizeof(CCTK_INT), 1, file);
# N2 = struct.unpack('i', filehandle.read(4))[0]
# # fwrite( & x2_beg, sizeof(CCTK_REAL), 1, file);
# x2_beg = struct.unpack('d', filehandle.read(8))[0]
#
# magic_number_check = 1.130814081305130e-21
# # fwrite( & magic_number, sizeof(CCTK_REAL), 1, file);
# magic_number = struct.unpack('d', filehandle.read(8))[0]
# if magic_number != magic_number_check:
# print("Error: Possible file corruption: Magic number mismatch. Found magic number = "+str(magic_number)+" . Expected "+str(magic_number_check))
# exit(1)
# # fwrite( & cctk_iteration, sizeof(CCTK_INT), 1, file);
# cctk_iteration = struct.unpack('i', filehandle.read(4))[0]
# # fwrite( & cctk_time, sizeof(CCTK_REAL), 1, file);
# cctk_time = struct.unpack('d', filehandle.read(8))[0]
#
# return gf_name,order,N0,R0,Rin,Rout,N1,x1_beg,theta_option,th_c,xi,th_n,N2,x2_beg,cctk_iteration,cctk_time
#
# # Now open the file and read all the data
# with open(datafile,"rb") as f:
# # Main loop over all gridfunctions
# for i in range(number_of_gridfunctions):
# # Data are output in chunks, one gridfunction at a time, with metadata
# # for each gridfunction stored at the top of each chunk
# # First read in the metadata:
# gf_name, order, N0, R0, Rin, Rout, N1, x1_beg, theta_option, th_c, xi, th_n, N2, x2_beg, cctk_iteration, cctk_time = read_header(f)
# print("\nReading gridfunction "+gf_name+", stored at interp order = "+str(order))
# data_chunk_size = N0*N1*N2*8 # 8 bytes per double-precision number
# # Next read in the full gridfunction data
# bytechunk = f.read(data_chunk_size)
# # Process the data using NumPy's frombuffer() function:
# # https://docs.scipy.org/doc/numpy/reference/generated/numpy.frombuffer.html
# buffer_res = np.frombuffer(bytechunk)
# # Reshape the data into a 3D NumPy array:
# # https://docs.scipy.org/doc/numpy/reference/generated/numpy.reshape.html
# this_data = buffer_res.reshape(N0,N1,N2)
#
# # Sanity check: Make sure the output in the "middle" of the grid looks reasonable.
# ii = int(N0/2)
# jj = int(N1/2)
# kk = int(N2/2)
# with open("output-gf"+str(i)+".txt","w") as file:
# for ii in range(N0):
# for kk in range(N2):
# r = ii*1.0/N0
# th = (jj*1.0)*np.pi/N1
# ph = (kk*1.0)*2.0*np.pi/N2
# xx = r*np.sin(th)*np.cos(ph)
# yy = r*np.sin(th)*np.sin(ph)
# zz = r*np.cos(th)
# file.write(str(xx)+" "+str(yy)+" "+str(zz)+" "+str(this_data[kk,jj,ii])+"\n")
#
# ```
# <a id='latex_pdf_output'></a>
#
# # Step 5: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ETK_thorn-Interpolation_to_Spherical_Grids_multi_order.pdf](Tutorial-ETK_thorn-Interpolation_to_Spherical_Grids_multi_order.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-ETK_thorn-Interpolation_to_Spherical_Grids_multi_order")
| Tutorial-ETK_thorn-Interpolation_to_Spherical_Grids_multi_order.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.1
# language: julia
# name: julia-1.4
# ---
# # A less basic backprop example
#
# The last notebook walked through an implementation of backpropagation for a small linear network with a single weight matrix and no bias. It ended with an example problem, *XOR*, that linear networks aren't able to solve. Here, we'll introduce a single hidden layer to this network to increase its expressivity and allow it solve the XOR problem.
#
# As part of this process, we'll work towards the implemention of a more generalizable backprop training loop.
#
# First, though, let's go over the problem again.
# ## XOR
#
# Exclusive OR, or XOR, is a logical operation on two inputs which evaluates to 1 if the inputs are different, and 0 if they are the same. Formally,
#
# $XOR(a, b) = \begin{cases}
# 1, & \text{if}~~~ a \neq b \\
# 0, & \text{otherwise}
# \end{cases}$
#
# This problem is not linearly separable, in that it is impossible to draw a straight line separating the inputs which evaluate to 1 from those which evaluate to 0. See the plot below.
# +
using Plots; pyplot()
X = [0 0; 1 1; 0 1; 1 0]
y = [0, 0, 1, 1]
scatter(X[:, 1], X[:, 2], color=[:blue, :blue, :red, :red], markersize=10, legend=false)
# -
# These four points $X$ and their corresponding XOR value $y$ will be our dataset.
#
# $X = \begin{pmatrix} 0 & 0 \\ 1 & 1 \\ 0 & 1 \\ 1 & 0 \end{pmatrix}$
#
# $y = \begin{pmatrix} 0 \\ 0 \\ 1 \\ 1 \end{pmatrix}$
#
# Now, we need to extend our previous linear model to include a hidden layer. Practically, this means two weight matrices, $W^1$ and $W^2$, where the superscript denotes the layer number. Like last time, we have two *features* as input. Rather than going from 2 features to 1 output node, however, we'll go from 2 features to 3 hidden nodes, and then from the 3 hidden nodes to 1 output node. For simplicity, both layers will use the sigmoid activation function.
#
# <br>
#
# ---
#
# *N.B. it is sufficient to have just 2 hidden nodes to solve this problem, but using 3 helps us keep the orientation of our matrices straight and is helpful for instruction.*
#
# ---
#
# <br>
#
# Putting this together, we have our model.
#
# $o^1 = \sigma(XW^1)$
#
# $o^2 = \sigma(o^1 W^2)$
#
# where
#
# $X \in \mathbb{R}^{4 \times 2}$ the input data
#
# $W^1 \in \mathbb{R}^{2 \times 3}$ the hidden layer weights
#
# $W^2 \in \mathbb{R}^{3 \times 1}$ the output weights
#
# $\sigma(z)= \frac{1}{1 + e^{-z}}$ the sigmoid activation function
#
#
# We define and initialize the weights and sigmoid function below.
# +
W1 = [0.1 0.1 0.1; 0.1 0.1 0.1]
W2 = [0.1, 0.1, 0.1]
function σ(z)
# Sigmoid activation function
return @. 1 / (1 + exp(-z))
end
# -
# ## The forward pass
#
# We start by computing the hidden layer output value, $o^1$.
#
# $z^1 = XW^1 =
# \begin{pmatrix} 0 & 0 \\ 1 & 1 \\ 0 & 1 \\ 1 & 0 \end{pmatrix}
# \begin{pmatrix} 0.1 & 0.1 & 0.1 \\ 0.1 & 0.1 & 0.1 \end{pmatrix}
# =
# \begin{pmatrix} 0 & 0 & 0 \\ 0.2 & 0.2 & 0.2 \\ 0.1 & 0.1 & 0.1 \\ 0.1 & 0.1 & 0.1 \end{pmatrix}$
#
# $o^1 = \sigma(z^1) = \begin{pmatrix}
# 0.5 & 0.5 & 0.5 \\
# 0.55 & 0.55 & 0.55 \\
# 0.52 & 0.52 & 0.52 \\
# 0.52 & 0.52 & 0.52
# \end{pmatrix}$
#
# Then, the network output $o^2$.
#
# $z^2 = o^1 W^2 =
# \begin{pmatrix} 0.5 & 0.5 & 0.5 \\0.55 & 0.55 & 0.55 \\ 0.52 & 0.52 & 0.52 \\ 0.52 & 0.52 & 0.52 \end{pmatrix}
# \begin{pmatrix} 0.1 \\ 0.1 \\ 0.1 \end{pmatrix}
# =
# \begin{pmatrix} 0.15 \\ 0.165 \\ 0.156 \\ 0.156 \end{pmatrix}
# $
#
# $o^2 = \sigma(z^2) =
# \begin{pmatrix} 0.537 \\ 0.541 \\ 0.539 \\ 0.539 \end{pmatrix}
# $
# +
z1 = X * W1
o1 = σ(z1)
z2 = o1 * W2
o2 = σ(z2)
println("z1: $z1")
println("o1: $o1")
println("z2: $z2")
println("o2: $o2")
# -
# We will now apply our decision function to the values of $o^2$ to get the predicted $y$ values $\hat{y}$ and compute the accuracy of the model. As a reminder, our decision function is a simple threshold
#
# $\hat{y} = \begin{cases}
# 1, & \text{if}~~~ o \ge 0.5 \\
# 0, & \text{otherwise}
# \end{cases}$
#
# The relevant functions and the output of our initial model are below.
# +
function decision(o)
return Int.(o .>= 0.5)
end
function accuracy(y, y_hat)
N = size(y, 1)
return sum(y .== y_hat) / N
end
y_hat = decision(o2)
acc = accuracy(y, y_hat)
println(y)
println(y_hat)
println("Accuracy: $acc")
# -
# We can also compute the loss of our initial model. Since this is still a binary classification problem, we'll use binary cross entropy again.
#
# $C = -\frac{1}{N} \sum_{i=1}^{N} y_i \times ln(o_i) + (1 - y_i) \times ln(1 - o_i)$
# +
function clip(a, lo=1e-15, hi=1 - 1e-15)
if a < lo
return lo
elseif a > hi
return hi
else
return a
end
end
function binary_crossentropy(y, o)
N = size(y, 1)
# We "clip" the predicted values to avoid
# domain errors in the log() function if our
# model happens to predict 0.0, and to prevent
# log returning 0.0 if the model predicts 1.0
o = clip.(o)
ces = @. (y * log(o)) + ((1-y) * log(1-o))
return (-1/N) * sum(ces)
end
loss = binary_crossentropy(y, o2)
println("Initial loss: $loss")
# -
# ## The backward pass
#
# Okay, the easy part is over. Now we have to compute the partial derivatives of each of the weights (from both weight matrices) with respect to the loss. We'll go through the equations at a high level first, so we can see how they fit together, then we'll get down to computing exactly what everything is.
#
# The partial wrt $W^2$ is analogous to the partial we computed for the linear model. That is,
#
# $\frac{\partial{C}}{\partial{W^2}} =
# \frac{\partial{C}}{\partial{o^2}}
# \frac{d{o^2}}{d{z^2}}
# \frac{\partial{z^2}}{\partial{W^2}}
# $
#
# The partial wrt $W^1$ is a bit more involved,
#
# $\frac{\partial{C}}{\partial{W^1}} =
# \frac{\partial{C}}{\partial{o^2}}
# \frac{d{o^2}}{d{z^2}}
# \frac{\partial{z^2}}{\partial{o^1}}
# \frac{d{o^1}}{d{z^1}}
# \frac{\partial{z^1}}{\partial{W^1}}
# $
#
# We can simplify this a bit, however, by noticing that the first two terms of $\frac{\partial{C}}{\partial{W^2}}$, namely $\frac{\partial{C}}{\partial{o^2}} \frac{d{o^2}}{d{z^2}}$, occur in both equations. We'll assign this operation the name $\delta^2$.
#
# $\delta^2 = \frac{\partial{C}}{\partial{o^2}} \frac{d{o^2}}{d{z^2}}$
#
# $\frac{\partial{C}}{\partial{W^2}} =
# \delta^2
# \frac{\partial{z^2}}{\partial{W^2}}
# $
#
# $\frac{\partial{C}}{\partial{W^1}} =
# \delta^2
# \frac{\partial{z^2}}{\partial{o^1}}
# \frac{d{o^1}}{d{z^1}}
# \frac{\partial{z^1}}{\partial{W^1}}
# $
#
# Before we go any further, let's look at the shapes. First for the forward pass:
#
# $X \in \mathbb{R}^{4 \times 2}$
#
# $W^1 \in \mathbb{R}^{2 \times 3}$
#
# $z^1 = XW^1 \in \mathbb{R}^{4 \times 3}$
#
# $o^1 = \sigma(z^1) \in \mathbb{R}^{4 \times 3}$
#
# $W^2 \in \mathbb{R}^{3 \times 1}$
#
# $z^2 = o^1W^2 \in \mathbb{R}^{4 \times 1}$
#
# $o^2 = \sigma(z^2) \in \mathbb{R}^{4 \times 1}$
#
# Then the backward pass:
#
# $\frac{\partial{C}}{\partial{o^2}} \in \mathbb{R}^{4 \times 1}$ (same shape as $o^2$)
#
# $\frac{d{o^2}}{d{z^2}} \in \mathbb{R}^{4 \times 1}$ (same shape as $z^2$)
#
# $\frac{\partial{z^2}}{\partial{W^2}} = o^1 \in \mathbb{R}^{4 \times 3}$
#
# $\frac{\partial{z^2}}{\partial{o^1}} = W^2 \in \mathbb{R}^{3 \times 1}$
#
# $\frac{d{o^1}}{d{z^1}} \in \mathbb{R}^{4 \times 3}$ (same shape as $z^1$)
#
# $\frac{\partial{z^1}}{\partial{W^1}} = X \in \mathbb{R}^{4 \times 2}$
#
# ### Output layer weights $W^2$
# ---
#
# $\frac{\partial{C}}{\partial{W^2}} =
# \frac{\partial{C}}{\partial{o^2}}
# \frac{d{o^2}}{d{z^2}}
# \frac{\partial{z^2}}{\partial{W^2}}
# $
#
# Now the shapes of the multiplications. First off, $\frac{\partial{C}}{\partial{o^2}}$ and $\frac{d{o^2}}{d{z^2}}$ have the same shape, so we want to compute their Hadamard product:
#
# $\delta^2 = \frac{\partial{C}}{\partial{o^2}} \frac{d{o^2}}{d{z^2}} = \mathbb{R}^{4 \times 1} \odot \mathbb{R}^{4 \times 1} = \mathbb{R}^{4 \times 1}$
#
# But $\frac{\partial{z^2}}{\partial{W^2}} = o^1 \in \mathbb{R}^{4 \times 3}$ so we'll need to transpose $\delta^2$ to get a compatible shape.
#
# $\frac{\partial{C}}{\partial{W^2}} = {\delta^2}^T \frac{\partial{z^2}}{\partial{W^2}} = \mathbb{R}^{1 \times 4} \mathbb{R}^{4 \times 3} = \mathbb{R}^{1 \times 3}$
#
# which transposed is $\mathbb{R}^{3 \times 1}$, so its compatible with $W^2$. This is encouraging, but let's make sure it makes sense by replacing the dimensions with variables indicating their meaning.
#
# $\frac{\partial{C}}{\partial{W^2}} = {\delta^2}^T \frac{\partial{z^2}}{\partial{W^2}} \in \mathbb{R}^{O \times N} \mathbb{R}^{N \times H} = \mathbb{R}^{O \times H}$
#
# $W^2 \in \mathbb{R}^{H ~\times~ O}$
#
# where $N$ is the number of examples, $H$ is the number of hidden nodes, and $O$ is the number of output nodes.
#
# Looking at the equations this way, we can see that $\frac{\partial{C}}{\partial{W^2}}$ connects the output nodes to the hidden nodes, and that transposing the result does indeed match up the proper dimensions to $W^2$ to allow us to do the weight updates for this layer. Great!
#
#
# ### Hidden layer weights $W^1$
# ---
#
# $\frac{\partial{C}}{\partial{W^1}} =
# \delta^2
# \frac{\partial{z^2}}{\partial{o^1}}
# \frac{d{o^1}}{d{z^1}}
# \frac{\partial{z^1}}{\partial{W^1}}
# $
#
# Next up, $\frac{\partial{C}}{\partial{W^1}}$. We know already that $\delta^2 \in \mathbb{R}^{4 \times 1}$ and $\frac{\partial{z^2}}{\partial{o^1}} = W^2 \in \mathbb{R}^{3 \times 1}$, so
#
# $\delta^2 \frac{\partial{z^2}}{\partial{o^1}} =
# \delta^2 {\frac{\partial{z^2}}{\partial{o^1}}}^T =
# \delta^2 {W^2}^T \in \mathbb{R}^{4 \times 1} \mathbb{R}^{1 \times 3} =
# \mathbb{R}^{4 \times 3}
# $
#
# $\frac{d{o^1}}{d{z^1}} \in \mathbb{R}^{4 \times 3}$ so we take the Hadamard product again.
#
# $\delta^2 {W^2}^T \odot \frac{d{o^1}}{d{z^1}} = \mathbb{R}^{4 \times 3} \mathbb{R}^{4 \times 3} = \mathbb{R}^{4 \times 3}$
#
# For concision's sake we'll define this as another $\delta$
#
# $\delta^1 = \delta^2 {W^2}^T \odot \frac{d{o^1}}{d{z^1}}$
#
# Finally, $\frac{\partial{z^1}}{\partial{W^1}} = X \in \mathbb{R}^{4 \times 1}$ so transposing the previous we get
#
# $\frac{\partial{C}}{\partial{W^1}} = {\delta^1}^T X = \mathbb{R}^{3 \times 4} \mathbb{R}^{4 \times 2} = \mathbb{R}^{3 \times 2}$
#
# Looking at the shapes as our variables again
#
# $\frac{\partial{C}}{\partial{W^1}} = {\delta^1}^T X = \mathbb{R}^{H \times N} \mathbb{R}^{N \times I} = \mathbb{R}^{H \times I}$ where $I$ is the number of input nodes.
#
# So transposing this we get $\mathbb{R}^{I \times H}$ which is the shape of $W^1$ and we're all set!
# +
function ∂C_∂o(y, o)
num = o .- y
denom = o .* (1 .- o)
return num ./ denom
end
function dσ_dz(z)
o = σ(z)
return @. o * (1.0 - o)
end
# +
function train(X, y, W1, W2, η=1, epochs=50, verbose=0)
log_at = epochs / 10
W1_t = copy(W1)
W2_t = copy(W2)
N = size(X, 1)
for t=1:epochs
# Forward pass
z1 = X * W1_t
o1 = σ(z1)
z2 = o1 * W2_t
o2 = σ(z2)
# Print the loss and training set accuracy every log_at epochs.
if t == 1 || t % log_at == 0
loss = binary_crossentropy(y, o2)
acc = accuracy(y, decision(o2))
println("Training step $t: Loss: $loss, Accuracy: $acc")
if verbose > 0
println("W1: $W1_t")
println("W2: $W2_t")
println("---")
end
end
# Backward pass
# Compute the gradients wrt W2 and W1
δ2 = ∂C_∂o(y, o2) .* dσ_dz(z2)
∂C_∂W2 = transpose(δ2) * o1
δ1 = (δ2 * transpose(W2_t)) .* dσ_dz(z1)
∂C_∂W1 = transpose(δ1) * X
# Update the weights given the gradients
W2_t = W2_t .- (η * transpose(∂C_∂W2))
W1_t = W1_t .- (η * transpose(∂C_∂W1))
end
return (W1_t, W2_t) # return the trained weights.
end
train(X, y, W1, W2, 0.1, 100)
# -
# ## Debugging
#
# What's going on here? We can see the loss is decreasing over the training steps, but incredibly slowly. Let's take advantage of our `verbose` option to get some more details. We'll also train for 1000 iterations just to make sure our model doesn't just need more time.
train(X, y, W1, W2, 0.1, 1000, 1)
# Even with 1000 iterations, the loss never decreases much and our weights don't seem to be diverging at all. To see why this might be happening, let's go back and compute the gradient wrt $W^2$ by hand.
#
# $\frac{\partial{C}}{\partial{W^2}} =
# \delta^2
# \frac{\partial{z^2}}{\partial{W^2}}
# $
#
# where $\delta^2 = \frac{\partial{C}}{\partial{o^2}} \frac{d{o^2}}{d{z^2}}$
#
# Incidentally, in the case of binary cross entropy loss with sigmoid activation, the equation for computing $\delta^2$ simplifies nicely.
#
# $\delta^2 = \frac{\partial{C}}{\partial{o^2}} \frac{d{o^2}}{d{z^2}} =
# \frac{o^2-y}{o^2(1-o^2)} \sigma(z^2)(1 - \sigma(z^2)) = o^2 - y$
#
# So
#
# $\frac{\partial{C}}{\partial{W^2}} = (o^2 - y)^T o^1 =
# \begin{pmatrix} .537 & .541 & .539 & .539 \end{pmatrix}
# \begin{pmatrix}
# 0.5 & 0.5 & 0.5 \\
# 0.55 & 0.55 & 0.55 \\
# 0.52 & 0.52 & 0.52 \\
# 0.52 & 0.52 & 0.52
# \end{pmatrix} =
# \begin{pmatrix} 1.13 & 1.13 & 1.13 \end{pmatrix}
# $
#
# Thus the equal updates for each weight seemed to be caused by the equal columns in $o^1$. More specifically, since $o^1 \in \mathbb{R}^{N \times H}$ where $H$ is the number of hidden layer nodes, this seems to be caused by the uniform initialization of the weight matrices.
#
# <br>
#
# This issue is discussed in section 8.4 of the book [*Deep Learning*](http://www.deeplearningbook.org/) by Goodfellow et al.
#
# > Perhaps the only property known with complete certainty is that the initial
# parameters need to “break symmetry” between different units. If two hidden
# units with the same activation function are connected to the same inputs, then
# these units must have different initial parameters. If they have the same initial
# parameters, then a deterministic learning algorithm applied to a deterministic cost
# and model will constantly update both of these units in the same way.
#
# That sounds like our problem! They go on to say shortly after,
#
# > The goal of having each unit compute a different function
# motivates random initialization of the parameters.
#
# So let's randomly initialize our weight matrices to see if that improves things.
import Random
Random.seed!(0)
W1_ = Random.rand(Float64, (2, 3))
W2_ = Random.rand(Float64, (3, 1))
W1_trained, W2_trained = train(X, y, W1_, W2_, 0.1, 1000)
# +
function predict(X, W1, W2)
z1 = X * W1
o1 = σ(z1)
z2 = o1 * W2
o2 = σ(z2)
return decision(o2)
end
predict([1 0], W1_trained, W2_trained)
# -
# And with that small tweak, we've solved the classic XOR problem!
#
# ---
#
# Continue on to the next notebook where we'll generalize the code we've implemented here so that we can build and train neural nets of arbitrary width and depth. This will involve writing some boilerplate code that builds the network as a directed graph of connected layers, a generalization of our training function to operate over this graph, as well as some additional code to let us work with larger datasets. Oh, and we'll finally add in the bias terms!
| notebooks/backprop_example2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Hh164S5d5wd_"
# ## Matrix multiplication from foundations
# + [markdown] id="-SkKioHH5weG"
# The *foundations* we'll assume throughout this course are:
#
# - Python
# - Python modules (non-DL)
# - pytorch indexable tensor, and tensor creation (including RNGs - random number generators)
# - fastai.datasets
# + [markdown] id="Y0UH7hId5weH"
# ## Check imports
# + id="Zq2NyCLM5-8T" colab={"base_uri": "https://localhost:8080/"} outputId="0cf556f9-547d-4140-8a2b-0f6ead1cb2f4"
from google.colab import drive
drive.mount('/content/gdrive')
# + id="hnAWcS__6H0B" colab={"base_uri": "https://localhost:8080/"} outputId="8318c69f-b2c0-4a60-f0a9-675a79205bfd"
# %cd /content/gdrive/MyDrive/ML/course-v3/nbs/dl2/
# + id="fRMgo_gu5weH"
# %load_ext autoreload
# %autoreload 2 #Reload all modules every time before executing the Python code typed.
# %matplotlib inline # renders the figure in a notebook (instead of displaying a dump of the figure object).
# + [markdown] id="W-I_4qnv5weI"
# [Jump_to lesson 8 video](https://course19.fast.ai/videos/?lesson=8&t=1850)
# + id="yaQkClza5weI"
#export
from exp.nb_00 import *
import operator
def test(a,b,cmp,cname=None):
if cname is None: cname=cmp.__name__
assert cmp(a,b),f"{cname}:\n{a}\n{b}" #The assert keyword is used when debugging code.The assert keyword lets you test if a condition in your code returns True, if not, the program will raise an AssertionError.
def test_eq(a,b): test(a,b,operator.eq,'==')
# + id="L3hRerlp5weJ"
test_eq(TEST,'test')
# + id="ObmMpXlD5weJ"
# To run tests in console:
# # ! python run_notebook.py 01_matmul.ipynb
# + [markdown] id="8GTbWAYz5weJ"
# ## Get data
# + [markdown] id="EtZXVzL45weJ"
# [Jump_to lesson 8 video](https://course19.fast.ai/videos/?lesson=8&t=2159)
# + id="hD6eyBh75weK"
#export
from pathlib import Path
from IPython.core.debugger import set_trace
from fastai import datasets
import pickle, gzip, math, torch, matplotlib as mpl
import matplotlib.pyplot as plt
from torch import tensor
MNIST_URL='http://deeplearning.net/data/mnist/mnist.pkl'
# + id="UH7I7n1t5weK"
#path = datasets.download_data(MNIST_URL, ext='.gz'); path
# + id="lSHFJugB5weL"
#with gzip.open(path, 'rb') as f:
# ((x_train, y_train), (x_valid, y_valid), _) = pickle.load(f, encoding='latin-1')
# + id="3DzZc5sT5weM" colab={"base_uri": "https://localhost:8080/"} outputId="a08f37e6-0a2e-4736-b827-f2eb1f11daac"
def get_data():
import os
import torchvision.datasets as datasets
root = '../data' #Root directory of dataset where MNIST/processed/training.pt and MNIST/processed/test.pt exist.
if not os.path.exists(root):
os.mkdir(root) #make directory for datasets.MNIST
train_set = datasets.MNIST(root=root, train=True, download=True)
test_set = datasets.MNIST(root=root, train=False, download=True)
x_train, x_valid = train_set.data.split([50000, 10000])
y_train, y_valid = train_set.targets.split([50000, 10000])
return (x_train.view(50000, -1) / 256.0), y_train.float(), (x_valid.view(10000, -1))/ 256.0, y_valid.float()
#Q:When you execute get_data() function, the content folder will look like as the following picture. Describe what the folder course-v3/nbs/data contain.
#A:The MNIST database (Modified National Institute of Standards
#and Technology database) is a large database of handwritten
#digits that is commonly used for training various image processing systems.
#The database is also widely used for training and testing in the field of machine learning.
#It contains 2 folders: processed and raw.
#The processed folder contains output. Right now we only have test.pt and training.pt
#The raw folder contains loaded database
#x_train,y_train,x_valid,y_valid = get_data()
#x_train,y_train,x_valid,y_valid = map(tensor, (x_train,y_train,x_valid,y_valid))
n,c = x_train.shape
x_train, x_train.shape, y_train, y_train.shape, y_train.min(), y_train.max()
# + id="kl-v-Gbm5weN"
assert n==y_train.shape[0]==50000
test_eq(c,28*28)
test_eq(y_train.min(),0)
test_eq(y_train.max(),9)
# + id="BanH20qk5weN"
mpl.rcParams['image.cmap'] = 'gray'
# + id="1uTTfw7K5weN"
img = x_train[0]
# + id="COPOuLuS5weO" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d25c3ea6-6782-46eb-9db3-8ba60aba7ddf"
img.view(28,28).type()
# + id="xTyr7u4N5weO" outputId="36a2a419-5fb9-4733-f878-c9712be7e38b" colab={"base_uri": "https://localhost:8080/", "height": 265}
plt.imshow(img.view((28,28)));
# + [markdown] id="v_4k3ELq5weO"
# ## Initial python model
# + [markdown] id="IUypptZs5weP"
# [Jump_to lesson 8 video](https://course19.fast.ai/videos/?lesson=8&t=2342)
# + [markdown] id="abxozlKa5weP"
# [Jump_to lesson 8 video](https://course19.fast.ai/videos/?lesson=8&t=2342)
# + id="GGmTsVvl5weQ"
weights = torch.randn(784,10)
# + id="qN-2KB-D5weQ"
bias = torch.zeros(10)
# + [markdown] id="iVursneS5weQ"
# #### Matrix multiplication
# + id="cmKVDe0_5weR"
def matmul(a,b):
ar,ac = a.shape # n_rows * n_cols
br,bc = b.shape
assert ac==br
c = torch.zeros(ar, bc)
for i in range(ar):
for j in range(bc):
for k in range(ac): # or br
c[i,j] += a[i,k] * b[k,j]
return c
# + id="WXnpCPkq5weS"
m1 = x_valid[:5]
m2 = weights
# + id="9Vv0psP35weS" outputId="8f446d6b-d366-47d8-a891-a1fd48e7e571"
m1.shape,m2.shape
# + id="uAPN4wgu5weT" outputId="91482cfc-fdbc-4d00-fb9b-06d2d0e4db44"
# %time t1=matmul(m1, m2)
# + id="5ldixQfR5weT" outputId="c41ac6ae-35ee-44f7-a311-d434360ba23a"
t1.shape
# + [markdown] id="xRKS4RkQ5weT"
# This is kinda slow - what if we could speed it up by 50,000 times? Let's try!
# + id="wofU5BU55weU" outputId="96e58c77-cc1a-42ae-b9ea-25abc2ec7a7c"
len(x_train)
# + [markdown] id="xy2NvoC15weU"
# #### Elementwise ops
# + [markdown] id="og6fSq5E5weU"
# Operators (+,-,\*,/,>,<,==) are usually element-wise.
#
# Examples of element-wise operations:
# + [markdown] id="2cC1xZa45weU"
# [Jump_to lesson 8 video](https://course19.fast.ai/videos/?lesson=8&t=2682)
# + id="ybp_ppAF5weV" outputId="02e2d932-5c6e-4f84-a1a6-bc4a434c0959"
a = tensor([10., 6, -4])
b = tensor([2., 8, 7])
a,b
# + id="hYcPlkUe5weV" outputId="50955ecb-c4f9-4f94-a28a-1fd1d162452f"
a + b
# + id="4EBtKsOf5weV" outputId="a55df0e5-225c-406e-dda7-814b75e84020"
(a < b).float().mean()
# + id="cM4YY1mv5weV" outputId="286d1fa7-8df1-48c6-f6af-a4ddc20d7521"
m = tensor([[1., 2, 3], [4,5,6], [7,8,9]]); m
# + [markdown] id="41VDP5V75weW"
# Frobenius norm:
#
# $$\| A \|_F = \left( \sum_{i,j=1}^n | a_{ij} |^2 \right)^{1/2}$$
#
# *Hint*: you don't normally need to write equations in LaTeX yourself, instead, you can click 'edit' in Wikipedia and copy the LaTeX from there (which is what I did for the above equation). Or on arxiv.org, click "Download: Other formats" in the top right, then "Download source"; rename the downloaded file to end in `.tgz` if it doesn't already, and you should find the source there, including the equations to copy and paste.
# + id="Ivtds24-5weW" outputId="1b502ea6-4448-4b18-fc6f-2f18d796ecb6"
(m*m).sum().sqrt()
# + [markdown] id="POpk3awE5weW"
# #### Elementwise matmul
# + id="mSAmtNDF5weW"
def matmul(a,b):
ar,ac = a.shape
br,bc = b.shape
assert ac==br
c = torch.zeros(ar, bc)
for i in range(ar):
for j in range(bc):
# Any trailing ",:" can be removed
c[i,j] = (a[i,:] * b[:,j]).sum()
return c
# + id="FGhLq5yg5weW" outputId="b838e70b-24db-4cdb-d556-9e7edd4ed1d1"
# %timeit -n 10 _=matmul(m1, m2)
# + id="dgyx_R7N5weX" outputId="ce591a57-42ea-4240-e8bd-cb6a78a3cf8e"
890.1/5
# + id="CYGCBooU5weX"
#export
def near(a,b): return torch.allclose(a, b, rtol=1e-3, atol=1e-5)
def test_near(a,b): test(a,b,near)
# + id="-MJeMgJT5weX"
test_near(t1,matmul(m1, m2))
# + [markdown] id="xO0Ddkvp5weX"
# ### Broadcasting
# + [markdown] id="-9bP_Oth5weY"
# The term **broadcasting** describes how arrays with different shapes are treated during arithmetic operations. The term broadcasting was first used by Numpy.
#
# From the [Numpy Documentation](https://docs.scipy.org/doc/numpy-1.10.0/user/basics.broadcasting.html):
#
# The term broadcasting describes how numpy treats arrays with
# different shapes during arithmetic operations. Subject to certain
# constraints, the smaller array is “broadcast” across the larger
# array so that they have compatible shapes. Broadcasting provides a
# means of vectorizing array operations so that looping occurs in C
# instead of Python. It does this without making needless copies of
# data and usually leads to efficient algorithm implementations.
#
# In addition to the efficiency of broadcasting, it allows developers to write less code, which typically leads to fewer errors.
#
# *This section was adapted from [Chapter 4](http://nbviewer.jupyter.org/github/fastai/numerical-linear-algebra/blob/master/nbs/4.%20Compressed%20Sensing%20of%20CT%20Scans%20with%20Robust%20Regression.ipynb#4.-Compressed-Sensing-of-CT-Scans-with-Robust-Regression) of the fast.ai [Computational Linear Algebra](https://github.com/fastai/numerical-linear-algebra) course.*
# + [markdown] id="KDUhs6cj5weY"
# [Jump_to lesson 8 video](https://course19.fast.ai/videos/?lesson=8&t=3110)
# + [markdown] id="L0ILLBBF5weY"
# #### Broadcasting with a scalar
# + id="V94VprX75weY" outputId="91d9b5f7-c6d5-4885-d493-afdcac64e38c"
a
# + id="mP5PzMGa5weZ" outputId="0174954a-7fc8-4afb-ca69-9eb21acf7560"
a > 0
# + [markdown] id="clrMNB8d5weZ"
# How are we able to do a > 0? 0 is being **broadcast** to have the same dimensions as a.
#
# For instance you can normalize our dataset by subtracting the mean (a scalar) from the entire data set (a matrix) and dividing by the standard deviation (another scalar), using broadcasting.
#
# Other examples of broadcasting with a scalar:
# + id="N_bUxzRS5weZ" outputId="fcecbcf9-0e6a-474a-aae6-d03168b3263d"
a + 1
# + id="tpuC3_gF5weZ" outputId="2f56c832-bae5-4c1c-d7d3-58353f872131"
m
# + id="TAd7motN5wea" outputId="69b6e699-b170-478c-ae80-aa467857e4e7"
2*m
# + [markdown] id="gL3PFFBP5wea"
# #### Broadcasting a vector to a matrix
# + [markdown] id="UAxMs4IC5wea"
# We can also broadcast a vector to a matrix:
# + id="tb0AqsW95web" outputId="7ad7399e-9d01-4289-ba8d-4cf21157e490"
c = tensor([10.,20,30]); c
# + id="QumR7M-H5web" outputId="9c70cbba-8ffe-4202-9ea1-5e37dbff31a8"
m
# + id="4aMty5W85web" outputId="b5292324-6b3d-4614-c068-2148d3c6f0e6"
m.shape,c.shape
# + id="eL3IAtR65wec" outputId="530058e5-542c-42df-a746-cd70d5573b0e"
m + c
# + id="76j5vREP5wec" outputId="64264775-507b-4e54-a48b-3667fa466528"
c + m
# + [markdown] id="tVBm5pQx5wec"
# We don't really copy the rows, but it looks as if we did. In fact, the rows are given a *stride* of 0.
# + id="3MR8nqNi5wec"
t = c.expand_as(m)
# + id="-DCyb4kw5wec" outputId="f716f3c7-bb02-46ce-a5d2-ba1932108614"
t
# + id="OajMYT225wec" outputId="48240d7c-afa9-49f1-d9cc-c470df918860"
m + t
# + id="3M9IkSoW5wed" outputId="265d8613-6efc-4265-9033-8c19a92b6e5e"
t.storage()
# + id="lPqGVMK-5wed" outputId="dbfbcddc-a8b0-4200-c772-43f5071736e1"
t.stride(), t.shape
# + [markdown] id="E-XH2eA85wed"
# You can index with the special value [None] or use `unsqueeze()` to convert a 1-dimensional array into a 2-dimensional array (although one of those dimensions has value 1).
# + id="wurEQisX5wed" outputId="2bca85ab-464b-417d-bf82-3b10e7b25f0d"
c.unsqueeze(0)
# + id="csnRc0DV5wee" outputId="733b86fb-cef5-409e-ec01-6b1b2d89ab07"
c.unsqueeze(1)
# + id="5n_1Tkri5wee" outputId="64497e00-9f37-4956-ee0e-472f6620ac6f"
m
# + id="xGpCvh5_5wee" outputId="84d5772c-1eed-4b88-fc73-e267e8d125c9"
c.shape, c.unsqueeze(0).shape,c.unsqueeze(1).shape
# + id="M0ZMR6FV5wee" outputId="c0a6f45b-3b0f-4851-e835-8de206e11fee"
c.shape, c[None].shape,c[:,None].shape
# + [markdown] id="rcq5MxOF5wef"
# You can always skip trailling ':'s. And '...' means '*all preceding dimensions*'
# + id="mYvMum7v5wef" outputId="3782871d-d586-4062-ac42-7615af80db11"
c[None].shape,c[...,None].shape
# + id="HZH1ZvRH5wef" outputId="389428fa-6795-40ea-b7b1-872d5f56773d"
c[:,None].expand_as(m)
# + id="ukoK_ki85wef" outputId="bd0018b6-35ae-46e4-afa8-9e5fbc0237e1"
m + c[:,None]
# + id="5CZBHXxE5weg" outputId="7ebbca41-6cd2-4b14-96ce-c6c5aeeca980"
c[:,None]
# + [markdown] id="vsczNMMo5weg"
# #### Matmul with broadcasting
# + id="QJZl6u0W5weg"
def matmul(a,b):
ar,ac = a.shape
br,bc = b.shape
assert ac==br
c = torch.zeros(ar, bc)
for i in range(ar):
# c[i,j] = (a[i,:] * b[:,j]).sum() # previous
c[i] = (a[i ].unsqueeze(-1) * b).sum(dim=0)
return c
# + id="wvHiLlLF5weg" outputId="5c2b2e4a-e836-4a88-cf00-d6b96535430c"
# %timeit -n 10 _=matmul(m1, m2)
# + id="imUqsLeP5weg" outputId="0f7f1c52-6cc8-4b88-883f-95cd300640bf"
885000/277
# + id="4Fb4fZ6M5weh"
test_near(t1, matmul(m1, m2))
# + [markdown] id="M_tJ013W5weh"
# #### Broadcasting Rules
# + id="6PV7joMn5weh" outputId="2ff1d0a5-8582-4ccf-9d9b-d037cf9b809e"
c[None,:]
# + id="m5PN6km65weh" outputId="8f40bd90-9bca-41b0-91dc-fb5057acdb3d"
c[None,:].shape
# + id="qroxBDg05wei" outputId="ebb6426c-ed56-4ad2-9877-b7b71708636e"
c[:,None]
# + id="_bK5GOse5wei" outputId="eb700ec6-b0a9-41a2-f510-a84b6e6f04df"
c[:,None].shape
# + id="8ixe-S5m5wei" outputId="cdc8c817-9621-4a96-a446-0c6a1c532c1f"
c[None,:] * c[:,None]
# + id="JcDDqVU55wei" outputId="e6595b33-ae96-4c9a-ea7d-2685176524cb"
c[None] > c[:,None]
# + [markdown] id="nLuOPfuw5wej"
# When operating on two arrays/tensors, Numpy/PyTorch compares their shapes element-wise. It starts with the **trailing dimensions**, and works its way forward. Two dimensions are **compatible** when
#
# - they are equal, or
# - one of them is 1, in which case that dimension is broadcasted to make it the same size
#
# Arrays do not need to have the same number of dimensions. For example, if you have a `256*256*3` array of RGB values, and you want to scale each color in the image by a different value, you can multiply the image by a one-dimensional array with 3 values. Lining up the sizes of the trailing axes of these arrays according to the broadcast rules, shows that they are compatible:
#
# Image (3d array): 256 x 256 x 3
# Scale (1d array): 3
# Result (3d array): 256 x 256 x 3
#
# The [numpy documentation](https://docs.scipy.org/doc/numpy-1.13.0/user/basics.broadcasting.html#general-broadcasting-rules) includes several examples of what dimensions can and can not be broadcast together.
# + [markdown] id="4a92SXMA5wej"
# ### Einstein summation
# + [markdown] id="CVt9KcaM5wej"
# Einstein summation (`einsum`) is a compact representation for combining products and sums in a general way. From the numpy docs:
#
# "The subscripts string is a comma-separated list of subscript labels, where each label refers to a dimension of the corresponding operand. Whenever a label is repeated it is summed, so `np.einsum('i,i', a, b)` is equivalent to `np.inner(a,b)`. If a label appears only once, it is not summed, so `np.einsum('i', a)` produces a view of a with no changes."
# + [markdown] id="T8974rRj5wek"
# [Jump_to lesson 8 video](https://course19.fast.ai/videos/?lesson=8&t=4280)
# + id="aBtCkNmV5wek"
# c[i,j] += a[i,k] * b[k,j]
# c[i,j] = (a[i,:] * b[:,j]).sum()
def matmul(a,b): return torch.einsum('ik,kj->ij', a, b)
# + id="NQBG02hR5wek" outputId="b3607bf8-10ab-45d0-af3d-7441bdee3473"
# %timeit -n 10 _=matmul(m1, m2)
# + id="by6rWbcB5wek" outputId="6166abd4-9f07-487c-d5f6-86f05f39ef4a"
885000/55
# + id="9UyeMjW05wel"
test_near(t1, matmul(m1, m2))
# + [markdown] id="JtrxnKQ85wel"
# ### pytorch op
# + [markdown] id="14oeF-uJ5wel"
# We can use pytorch's function or operator directly for matrix multiplication.
# + [markdown] id="KwOpKMKq5wel"
# [Jump_to lesson 8 video](https://course19.fast.ai/videos/?lesson=8&t=4702)
# + id="6AjNjY_p5wel" outputId="1c15be53-9bbf-4a21-ea55-7bd7bc4534ef"
# %timeit -n 10 t2 = m1.matmul(m2)
# + id="k1NUKDnA5wem" outputId="30fe4772-c91f-498a-936f-aba009b255a1"
# time comparison vs pure python:
885000/18
# + id="z5xTU1OA5wem"
t2 = m1@m2
# + id="8e_oGoqt5wem"
test_near(t1, t2)
# + id="exkkFzKI5wem" outputId="8cd367e4-c119-4146-f3e9-3589641228df"
m1.shape,m2.shape
# + [markdown] id="fIW4VwiE5wem"
# ## Export
# + id="nJZ7zBBF5wem" outputId="79ccd1fa-4cc2-4997-df68-97b3274a8355"
# !python notebook2script.py 01_matmul.ipynb
# + id="nTpWPv9R5wen"
| nbs/dl2/01_matmul.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recurrent Neural Network Example
#
# Build a recurrent neural network (LSTM) with TensorFlow.
#
# - Author: <NAME>
# - Project: https://github.com/aymericdamien/TensorFlow-Examples/
# ## RNN Overview
#
# <img src="http://colah.github.io/posts/2015-08-Understanding-LSTMs/img/RNN-unrolled.png" alt="nn" style="width: 600px;"/>
#
# References:
# - [Long Short Term Memory](http://deeplearning.cs.cmu.edu/pdfs/Hochreiter97_lstm.pdf), <NAME> & <NAME>, Neural Computation 9(8): 1735-1780, 1997.
#
# ## MNIST Dataset Overview
#
# This example is using MNIST handwritten digits. The dataset contains 60,000 examples for training and 10,000 examples for testing. The digits have been size-normalized and centered in a fixed-size image (28x28 pixels) with values from 0 to 1. For simplicity, each image has been flattened and converted to a 1-D numpy array of 784 features (28*28).
#
# 
#
# To classify images using a recurrent neural network, we consider every image row as a sequence of pixels. Because MNIST image shape is 28*28px, we will then handle 28 sequences of 28 timesteps for every sample.
#
# More info: http://yann.lecun.com/exdb/mnist/
# +
from __future__ import print_function
import tensorflow as tf
from tensorflow.contrib import rnn
# Import MNIST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
# +
# Training Parameters
learning_rate = 0.001
training_steps = 10000
batch_size = 128
display_step = 200
# Network Parameters
num_input = 28 # MNIST data input (img shape: 28*28)
timesteps = 28 # timesteps
num_hidden = 128 # hidden layer num of features
num_classes = 10 # MNIST total classes (0-9 digits)
# tf Graph input
X = tf.placeholder("float", [None, timesteps, num_input])
Y = tf.placeholder("float", [None, num_classes])
# -
# Define weights
weights = {
'out': tf.Variable(tf.random_normal([num_hidden, num_classes]))
}
biases = {
'out': tf.Variable(tf.random_normal([num_classes]))
}
def RNN(x, weights, biases):
# Prepare data shape to match `rnn` function requirements
# Current data input shape: (batch_size, timesteps, n_input)
# Required shape: 'timesteps' tensors list of shape (batch_size, n_input)
# Unstack to get a list of 'timesteps' tensors of shape (batch_size, n_input)
x = tf.unstack(x, timesteps, 1)
# Define a lstm cell with tensorflow
lstm_cell = rnn.BasicLSTMCell(num_hidden, forget_bias=1.0)
# Get lstm cell output
outputs, states = rnn.static_rnn(lstm_cell, x, dtype=tf.float32)
# Linear activation, using rnn inner loop last output
return tf.matmul(outputs[-1], weights['out']) + biases['out']
# +
logits = RNN(X, weights, biases)
prediction = tf.nn.softmax(logits)
# Define loss and optimizer
loss_op = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(
logits=logits, labels=Y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(loss_op)
# Evaluate model (with test logits, for dropout to be disabled)
correct_pred = tf.equal(tf.argmax(prediction, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# Initialize the variables (i.e. assign their default value)
init = tf.global_variables_initializer()
# -
# Start training
with tf.Session() as sess:
# Run the initializer
sess.run(init)
for step in range(1, training_steps+1):
batch_x, batch_y = mnist.train.next_batch(batch_size)
# Reshape data to get 28 seq of 28 elements
batch_x = batch_x.reshape((batch_size, timesteps, num_input))
# Run optimization op (backprop)
sess.run(train_op, feed_dict={X: batch_x, Y: batch_y})
if step % display_step == 0 or step == 1:
# Calculate batch loss and accuracy
loss, acc = sess.run([loss_op, accuracy], feed_dict={X: batch_x,
Y: batch_y})
print("Step " + str(step) + ", Minibatch Loss= " + \
"{:.4f}".format(loss) + ", Training Accuracy= " + \
"{:.3f}".format(acc))
print("Optimization Finished!")
# Calculate accuracy for 128 mnist test images
test_len = 128
test_data = mnist.test.images[:test_len].reshape((-1, timesteps, num_input))
test_label = mnist.test.labels[:test_len]
print("Testing Accuracy:", \
sess.run(accuracy, feed_dict={X: test_data, Y: test_label}))
Test Complete; Gopal
| tests/tf/recurrent_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Exploration and Visualization:
# - Univariable study of target and features (Continuous & Categorical features, separately)
# - Multivariate study of target and features
# - Testing the statistical assumptions: Normality, Homoscedasticity, etc.
# - Basic cleaning: Outliers, Missing data, Duplicate values
# - Chi-square test to examine dependency of target on categorical features (helpful for Feature Selection, if required)
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn
from sklearn import preprocessing
# %matplotlib inline
from scipy import stats
import warnings
warnings.filterwarnings("ignore")
# +
# Functions to detect & plot Outliers with different approaches:
def zscore_based_outliers(ys, threshold = 3):
mean_y = np.mean(ys)
stdev_y = np.std(ys)
z_scores = [(y - mean_y) / stdev_y for y in ys]
return np.abs(z_scores) > threshold
def mad_based_outlier(ys, thresh=3.5):
median = np.median(ys, axis=0)
mad=np.median(np.array([np.abs(y - median) for y in ys]))
modified_z_score=[0.6745 *(y - median) / mad for y in ys]
return np.abs(modified_z_score) > thresh
def iqr_based_outliers(ys):
quartile_1, quartile_3 = np.percentile(ys, [25, 75])
iqr = np.abs(quartile_3 - quartile_1)
lower_bound = quartile_1 - (iqr * 1.5)
upper_bound = quartile_3 + (iqr * 1.5)
return (ys > upper_bound) | (ys < lower_bound)
def plot_outliers(x):
fig, axes = plt.subplots(nrows=3)
fig.set_size_inches(6, 6)
for ax, func in zip(axes, [zscore_based_outliers, mad_based_outlier, iqr_based_outliers]):
sns.distplot(x, ax=ax, rug=True, hist=True)
outliers = x[func(x)]
ax.plot(outliers, np.zeros_like(outliers), 'ro', clip_on=False)
kwargs = dict(y=0.95, x=0.05, ha='left', va='top')
axes[0].set_title('Zscore-based Outliers', **kwargs)
axes[1].set_title('MAD-based Outliers', **kwargs)
axes[2].set_title('IQR-based Outliers', **kwargs)
fig.suptitle('Comparing Outlier Tests with n={}'.format(len(x)), size=14)
# +
df=pd.read_csv('C:/Users/rhash/Documents/Datasets/Loan prediction/train_loanPrediction.csv')
df.drop('Loan_ID', axis=1, inplace=True)
df.info()
# -
L_cat=['Gender', 'Married', 'Dependents', 'Education', 'Self_Employed', 'Credit_History', 'Property_Area', 'Loan_Status' ]
L_con=['ApplicantIncome', 'CoapplicantIncome', 'LoanAmount', 'Loan_Amount_Term']
# To detect and see the Missing Values:
sns.heatmap(df.isnull())
df.isnull().sum()
# +
df['Credit_History'].fillna(value=1, inplace=True)
df['Dependents'].fillna(value=str(0), inplace=True)
df['Self_Employed'].fillna(value='No', inplace=True)
df['Gender'].fillna(value='Male', inplace=True)
df['LoanAmount'].fillna(value=df['LoanAmount'].mean(), inplace=True)
# -
df.dropna(axis=0, inplace=True)
df.shape
# Encode text values to indexes(i.e. [1],[2],[3] for red,green,blue).
def encode_text_index(df, name):
le = preprocessing.LabelEncoder()
df[name] = le.fit_transform(df[name])
return le.classes_
for i in ['Gender', 'Married', 'Dependents', 'Education', 'Self_Employed', 'Property_Area', 'Loan_Status' ]:
encode_text_index(df, i)
df.head(3)
df.info()
# Imbalanced Data Set:
df["Loan_Status"].value_counts()
# Univariate analysis of Continuous Faetures: Statistical description (mean, std, skewness, Kurtosis) & Distribution plots
L=[]
for i in L_con:
print('_'*70 )
print('variable name: ', i, '\n')
print('Statistical description: \n', df[i].describe(), '\n', sep='')
if df[i].min()==0:
L.append(i)
print("Skewness = ", df[i].skew())
print("Kurtosis = ", df[i].kurt())
plot_outliers(np.array(df[i]))
plt.show()
# Multi-variable analysis of Continuous Features: Pairplot of all continuous features for different classes of target
sns.pairplot(pd.concat((df[L_con], df['Loan_Status']), axis=1 ), hue='Loan_Status')
# +
# Multivariable study: heatmap of correlation between continuous features
fig, ax = plt.subplots(figsize=(10,10))
sns.heatmap(df[L_con].corr(), annot=True, linewidths=1.5, ax=ax )
sns.clustermap(df[L_con].corr(), annot=True, linewidths=1.5 )
# -
# Multivariable analysis of Contineous Features:
for i in L_con:
print('_'*70 )
print('variable name: ', i)
S0=df[df['Loan_Status']==0][i]
S1=df[df['Loan_Status']==1][i]
t_test=stats.ttest_ind(S0, S1, equal_var = False)
print('z_statistic = ', round(t_test[0], 3))
print('p_value = ', round(t_test[1], 3), '\n')
if t_test[1]<=0.05:
print('This feature is significantly effective')
else:
print('This feature is NOT significantly effective')
fig = plt.figure(figsize=(9, 4))
ax1 = fig.add_subplot(121)
sns.barplot(x='Loan_Status', y=i, data=df)
ax2 = fig.add_subplot(122)
sns.boxplot( x="Loan_Status", y=i, data=df)
fig.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.25, wspace=0.7)
plt.show()
# To test the Statistical Assumptions on Continuous variables: We Check if our data meets the assumptions reuired by most mutivariate techniques _________
for i in L_con:
print('_'*70 )
print('variable name: ', i)
fig = plt.figure(figsize=(8, 6))
ax1 = fig.add_subplot(221)
ax1=sns.distplot(df[i], fit=stats.norm)
ax1.set_title('Before transformation:')
ax2 = fig.add_subplot(222)
res=stats.probplot(df[i], plot=ax2, rvalue=True)
b=0
if i in L:
b=0.1
ax3 = fig.add_subplot(223)
ax3=sns.distplot(stats.boxcox(b+df[i])[0], fit=stats.norm)
ax3.set_title('After "boxcox" transformation:')
ax4 = fig.add_subplot(224)
res=stats.probplot(stats.boxcox(b+df[i])[0], dist=stats.norm, plot=ax4, rvalue=True)
fig.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.4, wspace=0.3)
plt.show()
# +
# Multivariate analysis of Categorical Features: Value Counts and Success rate for different classes of a Categorical feature
for i in ['Gender', 'Married', 'Education', 'Dependents', 'Credit_History', 'Self_Employed', 'Property_Area']:
print('_'*70 )
print('variable name: ', i, '\n')
print('Value counts: \n', df[i].value_counts(), '\n', sep='')
p00=df[(df[i]==0) & (df['Loan_Status']==0)]['Loan_Status'].count()/df[df[i]==0]['Loan_Status'].count()
p01=df[(df[i]==0) & (df['Loan_Status']==1)]['Loan_Status'].count()/df[df[i]==0]['Loan_Status'].count()
p10=df[(df[i]==1) & (df['Loan_Status']==0)]['Loan_Status'].count()/df[df[i]==1]['Loan_Status'].count()
p11=df[(df[i]==1) & (df['Loan_Status']==1)]['Loan_Status'].count()/df[df[i]==1]['Loan_Status'].count()
print('Success rate for different values of this feature: \n', np.array([[p00, p01], [p10, p11]]))
sns.countplot(x=i, hue="Loan_Status", data=df[L_cat])
plt.show()
# +
F={}
for c in ['Gender', 'Married', 'Education', 'Dependents', 'Credit_History', 'Self_Employed', 'Property_Area']:
print('_'*70 )
print('_'*70 )
print('variable name: ', c, '\n')
c0=df[df['Loan_Status']==0][c].value_counts().sort_index().values
c1=df[df['Loan_Status']==1][c].value_counts().sort_index().values
obs = np.array([c0, c1])
g, p, dof, expctd = stats.chi2_contingency(obs)
F[c] = round(g,2)
print('Chi-square statistic= ', g)
print('p_value= ', p)
fig = plt.figure(figsize=(9, 4))
ax1 = fig.add_subplot(121)
sns.barplot(x='Loan_Status', y=c, data=df)
fig.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.25, wspace=0.7)
plt.show()
# +
# Sort and plot Categorical Features based on their Chi-square statistics (i.e. their dependency with Target):
# Helpful for Feature Selection
F_sorted=sorted(F,key=lambda i: F[i], reverse= True)
feature_df = pd.DataFrame([F[i] for i in F_sorted], index=[i for i in F_sorted]).reset_index()
feature_df.columns=['features', 'Chi-square test statistic']
fig, ax = plt.subplots(figsize=(18, 8))
sns.barplot(x='features', y='Chi-square test statistic', data=feature_df, color="blue", ax= ax)
plt.xticks(rotation=-45)
plt.show()
# -
| Projects in Python with Scikit-Learn- XGBoost- Pandas- Statsmodels- etc./Loan prediction (Data Exploration and Visualization) .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 2: Tides in the Puget Sound
# ## Learning Objectives
# I. Tidal Movement
# II. Tidal Cycle and Connection to Sea Surface Elevation
# ## Let's take a closer look at the movement of tides through the Strait of Juan de Fuca. We'll be using the tidal stations at Neah Bay, Port Angeles, and Port Townsend. Their tidal data and locations can be found at [NOAA Tides and Currents webpage.](https://tidesandcurrents.noaa.gov/map/)
# ## Below, we plotted the locations of the three tidal stations in the Strait of Juan de Fuca.
# ## From west to east: Neah Bay, Port Angeles, and Port Townsend.
import tydal.module2_utils as tide
import tydal.quiz2
stationmap = tide.add_station_maps()
stationmap
# ## As the tide moves through the Strait, it creates a change in the elevation of the water surface. Below we'll cycle through a tidal cycle and look at how the tide moves through the Strait. Use the slider to move through the time series and look how the measured tide at a station relates to the other stations, and its effect on the water elevation.
NeahBay = tide.load_Neah_Bay('Data/')
PortAngeles = tide.load_Port_Angeles('Data/')
PortTownsend = tide.load_Port_Townsend('Data/')
Tides = tide.create_tide_dataset(NeahBay,PortAngeles,PortTownsend)
# %matplotlib inline
tide.plot_tide_data(Tides,'2016-10-01','2016-10-02')
# ## Take a look at the time series for each station. It looks like a wave. In fact, the tide is a wave. That wave propogates through the Strait, starting at Neah Bay and travelling to Port Townsend. This is reflected in the elevation, as the peak elevation moves from one station to the following station.
# # # Module 2 Quiz
quiz2.quiz()
| Module2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 因子分析模型(241)
from IPython.display import Latex
from IPython.display import display, Math, Latex
print_latex = lambda latex_str: display(Latex(latex_str))
print('因子分析的基本概念:')
print_latex(r'因子分析模型: $X=\mu+\Lambda F+\epsilon$')
print_latex(r'其中$E(F)=0,\quad E(\epsilon)=0,\quad Cov(F)=I_m,\quad D(\epsilon)=Cov(\epsilon)=diag(\sigma_1^2,\cdots,\sigma_m^2),\quad Cov(F, \epsilon)=0$')
print_latex(r'原始变量$X$的协方差矩阵分解: $Cov(X)=\Lambda\Lambda^T+diag(\sigma_1^2,\cdots,\sigma_m^2)$')
print_latex(r'载荷因子$\alpha_{ij}$反映第$i$个变量和第$j$个公共因子的相关系数. 绝对值越大相关的密切程度越高.')
print_latex(r'变量$X_i$的共同度记为$h_i^2=\sum\limits_{j=1}^m\alpha_{ij}^2$, 又有$1=h_i^2+\sigma_i^2$, 故$h_i^2$越接近1, 因子分析效果越好')
print_latex(r'$\Lambda$中各列平方和$S_j=\sum\limits_{i=1}^p\alpha_{ij}^2$, 用于衡量$F_j$的相对重要性.')
print('\n'*3)
print('主成分分析法估计载荷因子:')
print_latex(r'设相关系数矩阵$R$的特征值和对应特征向量分别为: $\lambda_1\ge\lambda_2\ge\cdots\ge\lambda_p$和$\eta_1,\eta_2,\cdots,\eta_p$')
print_latex(r'设m<p, 则因子载荷矩阵$\Lambda=[\sqrt{\lambda_1}\eta_1,\sqrt{\lambda_2}\eta_2,\cdots,\sqrt{\lambda_m}\eta_m]$')
print_latex(r'特殊因子的方差用$R-\Lambda\Lambda^T$的对角元来估计. 即$\sigma_i^2=1-\sum\limits_{j=1}^m\alpha_{ij}^2$')
print_latex(r'因子载荷矩阵的估计方法: 1.主成分分析法(242页); ')
print_latex(r'通过因子旋转来直观的判断因子的实际意义')
print_latex(r'因子得分: 反过来把公共因子表示为原变量的线性组合.')
print_latex(r'因子得分函数: $F_j=c_j+\beta_{j1}X_1+\cdots+\beta_{jp}X_p,\ j=1,2,\cdots,m$')
print('\n'*3)
print('因子得分:')
print_latex(r'巴特莱特因子得分估计: $\hat{F}=(\Lambda^TD^{-1}\Lambda)^{-1}\Lambda^TD^{-1}(X-\mu)$')
print_latex(r'回归方法因子得分估计: $\hat{F}=(\hat{F}_{ij})_{n\times m}=X_0R^{-1}\Lambda$')
# +
import numpy as np
from sklearn.decomposition import PCA
import scipy
import sympy
import pandas as pd
R = np.array([[1.000, 0.577, 0.509, 0.387, 0.462],
[0.577, 1.000, 0.599, 0.389, 0.322],
[0.509, 0.599, 1.000, 0.436, 0.426],
[0.387, 0.389, 0.436, 1.000, 0.523],
[0.462, 0.322, 0.426, 0.523, 1.000]
])
# 列为单位特征向量. 由于R是对称阵, 因此都是正交的特征向量(下面一行可以验证这一点).
# print(np.array([[np.round(np.sum(eigvec[:,i]*eigvec[:,j])) for i in range(R.shape[0])] for j in range(R.shape[1])]))
eigval, eigvec = np.linalg.eig(R)
order = eigval.argsort()[::-1]
eigvec = np.array([eigvec[:, order[i]] for i in range(order.shape[0])]).T
eigval = np.sort(eigval)[::-1]
eigvec = eigvec*np.sign(np.sum(eigvec, axis=0))
# 因子载荷矩阵
Lambda = eigvec*np.sqrt(eigval)
print(eigval, Lambda, sep='\n')
# 信息贡献率
b = np.array([eigval[i]/eigval.sum() for i in range(eigval.shape[0])])
print(b)
# 累积贡献率
alpha = np.array([b[:i+1].sum() for i in range(b.shape[0])])
print(alpha)
m = 2
# 特殊因子方差
var_e = [1-Lambda[i, :m].sum() for i in range(Lambda.shape[0])]
print(var_e)
# +
from sklearn.datasets import load_digits
from sklearn.decomposition import FactorAnalysis
import numpy as np
X, _ = load_digits(return_X_y=True)
fa = FactorAnalysis(n_components=56, random_state=0)
X_transformed = fa.fit_transform(X)
print(X_transformed.shape)
print(fa.components_.shape)
print(fa.noise_variance_.shape)
print(fa.mean_.shape)
# 变换的公式满足下面这个:
print(np.round(fa.mean_ + np.matmul(X_transformed[0], fa.components_) + fa.noise_variance_, 0))
print(X[0])
| .ipynb_checkpoints/10.3 因子分析-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution:
def reverseStr(self, s: str, k: int) -> str:
n = len(s)
s = list(s)
p = 0
while p < n:
if p + k < n:
s[p:p+k] = s[p:p+k][::-1]
else:
s[p:] = s[p:][::-1]
p += 2*k
return ''.join(s)
s = Solution()
s.reverseStr("abcdefg", 2)
| algorithms/541-reverse-string-ii.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# +
import json
import os
import glob
import pprint
from tqdm import tqdm
from collections import Counter
import pandas as pd
import matplotlib.pyplot as plt
# -
df = pd.read_csv("datasets/verified.dat")
# df = df[df['friends_count'] <= 1000]
df.info()
df.head()
# +
# df[df['FriendsCount'] < 5000].info()
# -
# ### Dict structure
# uid:{
#
# uid: 123,
# name: '',
# friends_count: 0,
# followers_count: 0,
# listed_count: 0,
# statuses_count: 0,
# pf_fake: 0,
# pf_real: 0,
# gc_fake: 0,
# gc_real: 0,
# description: '',
# tweets: []
#
# }
ids = list(df['#ID'])
# +
users = {}
for i in ids:
users[i] = {
'uid': i,
'pf_fake': 0,
'pf_real': 0,
'gc_fake': 0,
'gc_real': 0,
'description': '',
'tweets': []
}
# -
len(users)
user_dict = users
dataset_path = "../FakeNewsNet/code/fakenewsnet_dataset"
# ## PolitiFact
# ### Fake Tweets & Retweets
# +
files = glob.iglob(f"../FakeNewsNet/code/fakenewsnet_dataset/politifact/fake/*/tweets/*.json")
# Expected number of files ~ 143000
for file in tqdm(files):
with open(file, encoding='utf-8', mode='r') as currentFile:
data = json.load(currentFile)
tweet = data['text']
des = data['user']['description']
user_id = data['user']['id']
if user_id in user_dict:
user_dict[user_id]['pf_fake']+=1
user_dict[user_id]['description'] = des
user_dict[user_id]['tweets'].append(tweet)
# +
files_rt = glob.iglob(f"../FakeNewsNet/code/fakenewsnet_dataset/politifact/fake/*/retweets/*.json")
# Expected number of files ~ 2400
for file in tqdm(files_rt):
with open(file, encoding='utf-8', mode='r') as currentFile:
data = json.load(currentFile)
for d in data['retweets']:
tweet = d['text']
des = d['user']['description']
user_id = d['user']['id']
if user_id in user_dict:
user_dict[user_id]['pf_fake']+=1
user_dict[user_id]['description'] = des
user_dict[user_id]['tweets'].append(tweet)
# -
# ### Real Tweets & Retweets
# +
files = glob.iglob(f"../FakeNewsNet/code/fakenewsnet_dataset/politifact/real/*/tweets/*.json")
# Expected number of files ~ 362000
for file in tqdm(files):
with open(file, encoding='utf-8', mode='r') as currentFile:
data = json.load(currentFile)
tweet = data['text']
des = data['user']['description']
user_id = data['user']['id']
if user_id in user_dict:
user_dict[user_id]['pf_real']+=1
user_dict[user_id]['description'] = des
user_dict[user_id]['tweets'].append(tweet)
# +
files_rt = glob.iglob(f"../FakeNewsNet/code/fakenewsnet_dataset/politifact/real/*/retweets/*.json")
# Expected number of files ~ 0
for file in tqdm(files_rt):
with open(file, encoding='utf-8', mode='r') as currentFile:
data = json.load(currentFile)
for d in data['retweets']:
tweet = d['text']
des = d['user']['description']
user_id = d['user']['id']
if user_id in user_dict:
user_dict[user_id]['pf_real']+=1
user_dict[user_id]['description'] = des
user_dict[user_id]['tweets'].append(tweet)
# -
# ## GossipCop
# ### Fake Tweets & Retweets
# +
files = glob.iglob(f"../FakeNewsNet/code/fakenewsnet_dataset/gossipcop/fake/*/tweets/*.json")
# Expected number of files ~ 524000
for file in tqdm(files):
with open(file, encoding='utf-8', mode='r') as currentFile:
data = json.load(currentFile)
tweet = data['text']
des = data['user']['description']
user_id = data['user']['id']
if user_id in user_dict:
user_dict[user_id]['gc_fake']+=1
user_dict[user_id]['description'] = des
user_dict[user_id]['tweets'].append(tweet)
# +
files_rt = glob.iglob(f"../FakeNewsNet/code/fakenewsnet_dataset/gossipcop/fake/*/retweets/*.json")
# Expected number of files ~ 0
for file in tqdm(files_rt):
with open(file, encoding='utf-8', mode='r') as currentFile:
data = json.load(currentFile)
for d in data['retweets']:
tweet = d['text']
des = d['user']['description']
user_id = d['user']['id']
if user_id in user_dict:
user_dict[user_id]['gc_fake']+=1
user_dict[user_id]['description'] = des
user_dict[user_id]['tweets'].append(tweet)
# -
# ### Real Tweets & Retweets
# +
files = glob.iglob(f"../FakeNewsNet/code/fakenewsnet_dataset/gossipcop/real/*/tweets/*.json")
# Expected number of files ~ 815000
for file in tqdm(files):
with open(file, encoding='utf-8', mode='r') as currentFile:
data = json.load(currentFile)
tweet = data['text']
des = data['user']['description']
user_id = data['user']['id']
if user_id in user_dict:
user_dict[user_id]['gc_real']+=1
user_dict[user_id]['description'] = des
user_dict[user_id]['tweets'].append(tweet)
# +
files = glob.iglob(f"../FakeNewsNet/code/fakenewsnet_dataset/gossipcop/real/*/retweets/*.json")
# Expected number of files ~
for file in tqdm(files):
with open(file, encoding='utf-8', mode='r') as currentFile:
data = json.load(currentFile)
for d in data['retweets']:
tweet = d['text']
des = d['user']['description']
user_id = d['user']['id']
if user_id in user_dict:
user_dict[user_id]['gc_real']+=1
user_dict[user_id]['description'] = des
user_dict[user_id]['tweets'].append(tweet)
# -
# ## Create Dataset
# +
list_of_lists = []
for user_id in user_dict:
gc_fake = user_dict[user_id]['gc_fake']
gc_real = user_dict[user_id]['gc_real']
pf_fake = user_dict[user_id]['pf_fake']
pf_real = user_dict[user_id]['pf_real']
des = user_dict[user_id]['description']
tweets = "^".join(user_dict[user_id]['tweets'])
list_of_lists.append([user_id, pf_fake, pf_real, gc_fake, gc_real, des, tweets])
# -
df_tw = pd.DataFrame(list_of_lists, columns=["uid", 'pf_fake','pf_real','gc_fake','gc_real', 'description', 'tweets'])
df_tw.head()
# Drop useless columns
df.drop(['Protected', 'CreatedAt', 'URL', 'ProfileImageURL', 'Location', 'Subject', 'Relation', 'Verified'], axis=1, inplace=True)
df.info()
df_tw.info()
df_feat = df.merge(df_tw, how='inner', left_on='#ID', right_on='uid')
df_feat.info()
df_feat.head()
# +
df_feat['total_fake'] = df_feat['pf_fake'] + df_feat['gc_fake']
df_feat['total_real'] = df_feat['pf_real'] + df_feat['gc_real']
df_feat['net_trust'] = df_feat['total_real'] - df_feat['total_fake']
df_feat['total_news'] = df_feat['total_real'] + df_feat['total_fake']
df_feat['fake_prob'] = df_feat['total_fake'] / df_feat['total_news']
df_feat['net_trust_norm'] = df_feat['net_trust']/df_feat['total_news']
# -
df_feat['fake'] = [1 if x >= 0.5 else 0 if x < 0.5 else 2 for x in df_feat['fake_prob']]
df_feat.info()
# +
# df_feat.to_csv('datasets/verified_features_300k.csv', index=False)
# +
# df_feat = pd.read_csv("datasets/verified_features_300k.csv")
# -
df_final = df_feat[
(df_feat['ScreenName'] != 'GossipCop') & (df_feat['ScreenName'] != 'PolitiFact')
# & (df_feat['net_trust_norm'] != 0)
& (df_feat['FriendsCount'] <= 10000)
& (df_feat['total_fake'] > 0)
& (df_feat['total_real'] > 0)
].sort_values(['total_news', 'net_trust_norm', 'net_trust', 'FollowersCount'], ascending=False)
df_final.info()
df_final['fake'] = [1 if x >= 0.5 else 0 if x < 0.5 else 2 for x in df_final['fake_prob']]
df_final.to_csv('datasets/verified_features_3k.csv', index=False)
# +
df_final.hist(column='fake_prob', bins=20)
df_final.hist(column='FriendsCount', bins=20)
df_final.hist(column='FollowersCount', bins=20)
plt.show()
# -
df_final["fake"].value_counts().plot.bar()
plt.show()
df_final["fake"].value_counts()
dat = pd.read_csv("datasets/verified.dat")
filtered = dat[dat['#ID'].isin(list(df_final['#ID']))]
filtered.info()
filtered.to_csv('datasets/verified_3k.dat', index = False)
| user_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
np.seterr(over='ignore')
import os
os.environ["TF_CPP_MIN_LOG_LEVEL"]="3"
from keras.datasets import mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
import matplotlib.pyplot as plt
w=10
h=10
fig=plt.figure(figsize=(8, 8))
columns = 10
rows = 10
for i in range(1, columns*rows +1):
img = x_test[i]
fig.add_subplot(rows, columns, i)
plt.imshow(img, cmap='gray')
plt.show()
print(x_train.shape)
print(y_train.shape)
print(x_test.shape)
print(y_test.shape)
# Changing dimensions to N x D = 28x28
# Number of samples you want in training data. 60,000 is max.
N = 50000
x_tr = []
for i in range(N):
x_tr.append(x_train[i,:].flatten())
x_te = []
for i in range(10000):
x_te.append(x_test[i,:].flatten())
print(len(x_tr),len(x_tr[0]))
print(len(x_te),len(x_te[0]))
class NN:
def __init__(self):
pass
# Memorize the training data
def train(self, X, y):
# X is 2D if size N x D = 28x28, so each row is an example
# y is 1D of size N
self.tr_x = X
self.tr_y = y
# For the test image:
# find nearest train image with minimum distance from the test image
# predict the label of nearest training image
def predict(self, x):
# x is of size D = 28x28 for which we want to predict the label
# returns the predicted label for the input x
min_idx = None
min_dist = 100000000
for test_sample in range(len(self.tr_x)):
dist = 0
for each_value in range(len(self.tr_x[0])):
dist += abs(float((self.tr_x[test_sample][each_value] - x[each_value])))
if dist < min_dist:
min_dist = dist
min_idx = test_sample
return self.tr_y[min_idx]
classifier = NN()
classifier.train(x_tr, y_train)
plt.figure(figsize=(2,2))
plt.imshow(x_test[3],cmap="gray")
plt.show()
print(classifier.predict(x_te[3]))
plt.figure(figsize=(2,2))
plt.imshow(x_test[2],cmap="gray")
plt.show()
print(classifier.predict(x_te[2]))
plt.figure(figsize=(2,2))
plt.imshow(x_test[33],cmap="gray")
plt.show()
print(classifier.predict(x_te[33]))
plt.figure(figsize=(2,2))
plt.imshow(x_test[59],cmap="gray")
plt.show()
print(classifier.predict(x_te[59]))
plt.figure(figsize=(2,2))
plt.imshow(x_test[36],cmap="gray")
plt.show()
print(classifier.predict(x_te[36]))
| NN_Self_Paced/MNIST_KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Sveučilište u Zagrebu
# Fakultet elektrotehnike i računarstva
#
# ## Strojno učenje 2019/2020
# http://www.fer.unizg.hr/predmet/su
# ------------------------------
#
# ### Laboratorijska vježba 1: Regresija
#
# *Verzija: 1.2
# Zadnji put ažurirano: 27. rujna 2019.*
#
# (c) 2015-2019 <NAME>, <NAME>
#
# Objavljeno: **30. rujna 2019.**
# Rok za predaju: **21. listopada 2019. u 07:00h**
#
# ------------------------------
# ### Upute
#
# Prva laboratorijska vježba sastoji se od deset zadataka. U nastavku slijedite upute navedene u ćelijama s tekstom. Rješavanje vježbe svodi se na **dopunjavanje ove bilježnice**: umetanja ćelije ili više njih **ispod** teksta zadatka, pisanja odgovarajućeg kôda te evaluiranja ćelija.
#
# Osigurajte da u potpunosti **razumijete** kôd koji ste napisali. Kod predaje vježbe, morate biti u stanju na zahtjev asistenta (ili demonstratora) preinačiti i ponovno evaluirati Vaš kôd. Nadalje, morate razumjeti teorijske osnove onoga što radite, u okvirima onoga što smo obradili na predavanju. Ispod nekih zadataka možete naći i pitanja koja služe kao smjernice za bolje razumijevanje gradiva (**nemojte pisati** odgovore na pitanja u bilježnicu). Stoga se nemojte ograničiti samo na to da riješite zadatak, nego slobodno eksperimentirajte. To upravo i jest svrha ovih vježbi.
#
# Vježbe trebate raditi **samostalno**. Možete se konzultirati s drugima o načelnom načinu rješavanja, ali u konačnici morate sami odraditi vježbu. U protivnome vježba nema smisla.
# Učitaj osnovne biblioteke...
import numpy as np
import sklearn
import matplotlib.pyplot as plt
# %pylab inline
# ## Zadatci
# ### 1. Jednostavna regresija
# Zadan je skup primjera $\mathcal{D}=\{(x^{(i)},y^{(i)})\}_{i=1}^4 = \{(0,4),(1,1),(2,2),(4,5)\}$. Primjere predstavite matrixom $\mathbf{X}$ dimenzija $N\times n$ (u ovom slučaju $4\times 1$) i vektorom oznaka $\textbf{y}$, dimenzija $N\times 1$ (u ovom slučaju $4\times 1$), na sljedeći način:
#
X = np.array([[0],[1],[2],[4]])
y = np.array([4,1,2,5])
def poly3(x):
return x**3+2*x**2
# ### (a)
#
# Proučite funkciju [`PolynomialFeatures`](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html) iz biblioteke `sklearn` i upotrijebite je za generiranje matrice dizajna $\mathbf{\Phi}$ koja ne koristi preslikavanje u prostor više dimenzije (samo će svakom primjeru biti dodane *dummy* jedinice; $m=n+1$).
#
from sklearn.preprocessing import PolynomialFeatures
# Vaš kôd ovdje
poly = PolynomialFeatures(1)
X_poly = poly.fit_transform(X)
print(X_poly)
# ### (b)
# Upoznajte se s modulom [`linalg`](http://docs.scipy.org/doc/numpy/reference/routines.linalg.html). Izračunajte težine $\mathbf{w}$ modela linearne regresije kao $\mathbf{w}=(\mathbf{\Phi}^\intercal\mathbf{\Phi})^{-1}\mathbf{\Phi}^\intercal\mathbf{y}$. Zatim se uvjerite da isti rezultat možete dobiti izračunom pseudoinverza $\mathbf{\Phi}^+$ matrice dizajna, tj. $\mathbf{w}=\mathbf{\Phi}^+\mathbf{y}$, korištenjem funkcije [`pinv`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.pinv.html).
from numpy import linalg
# Vaš kôd ovdje
invers = linalg.inv(np.matmul(X_poly.transpose(),X_poly))
pseudoinvers = np.matmul(invers,X_poly.transpose())
w = np.matmul(pseudoinvers,y)
w1 = np.matmul(linalg.pinv(X_poly),y)
print(w1)
# Radi jasnoće, u nastavku je vektor $\mathbf{x}$ s dodanom *dummy* jedinicom $x_0=1$ označen kao $\tilde{\mathbf{x}}$.
# ### (c)
# Prikažite primjere iz $\mathcal{D}$ i funkciju $h(\tilde{\mathbf{x}})=\mathbf{w}^\intercal\tilde{\mathbf{x}}$. Izračunajte pogrešku učenja prema izrazu $E(h|\mathcal{D})=\frac{1}{2}\sum_{i=1}^N(\tilde{\mathbf{y}}^{(i)} - h(\tilde{\mathbf{x}}))^2$. Možete koristiti funkciju srednje kvadratne pogreške [`mean_squared_error`]( http://scikit-learn.org/stable/modules/generated/sklearn.metrics.mean_squared_error.html) iz modula [`sklearn.metrics`](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.metrics).
#
# **Q:** Gore definirana funkcija pogreške $E(h|\mathcal{D})$ i funkcija srednje kvadratne pogreške nisu posve identične. U čemu je razlika? Koja je "realnija"?
# +
# Vaš kôd ovdje
from sklearn.metrics import mean_squared_error
import scipy as sp
def h(x):
return w[1]*x + w[0]
def squared_loss(y,hx):
return (y-hx)**2
def mean_square_error(X,y):
error = 0
if(len(X) != len(y)):
exit("Dimension error!")
for i in range(len(X)):
error = error + squared_loss(y[i],h(X[i]))
return error*0.5
def predicted(X):
predicted_values = list()
for i in range(len(X)):
predicted_values.append(h(X[i]))
return predicted_values
print(mean_square_error(X,y)[0])
print(mean_squared_error(y,predicted(X)))
xs = sp.linspace(0,4)
plt.scatter(X,y)
plt.plot(xs,h(xs),"r")
print(matmul(X_poly,w))
# -
# ### (d)
# Uvjerite se da za primjere iz $\mathcal{D}$ težine $\mathbf{w}$ ne možemo naći rješavanjem sustava $\mathbf{w}=\mathbf{\Phi}^{-1}\mathbf{y}$, već da nam doista treba pseudoinverz.
#
# **Q:** Zašto je to slučaj? Bi li se problem mogao riješiti preslikavanjem primjera u višu dimenziju? Ako da, bi li to uvijek funkcioniralo, neovisno o skupu primjera $\mathcal{D}$? Pokažite na primjeru.
# +
# <NAME>
#This piece of code is causing an error, because Desing matrix is not a square matrix!
#invers = linalg.inv(X_poly)
def phi(x):
return [1,x,x**2,x**3]
design_matrix = list()
for i in range(len(X)):
design_matrix.append(phi(X[i][0]))
invers = linalg.inv(design_matrix)
w = np.matmul(invers,y)
print(w)
# -
# ### (e)
# Proučite klasu [`LinearRegression`](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html) iz modula [`sklearn.linear_model`](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.linear_model). Uvjerite se da su težine koje izračunava ta funkcija (dostupne pomoću atributa `coef_` i `intercept_`) jednake onima koje ste izračunali gore. Izračunajte predikcije modela (metoda `predict`) i uvjerite se da je pogreška učenja identična onoj koju ste ranije izračunali.
from sklearn.linear_model import LinearRegression
# +
# Vaš kôd ovdje
model = LinearRegression()
model.fit(X_poly,y)
model.score(X_poly,y)
print("Lib w0: "+str(model.intercept_)+" w1: "+str(model.coef_[1]))
print("w0: "+str(w[0])+" w1: "+str(w[1]))
predicted_y = model.predict(X_poly)
print(mean_squared_error(y,predicted_y))
# -
# ### 2. Polinomijalna regresija i utjecaj šuma
# ### (a)
#
# Razmotrimo sada regresiju na većem broju primjera. Definirajte funkciju `make_labels(X, f, noise=0)` koja uzima matricu neoznačenih primjera $\mathbf{X}_{N\times n}$ te generira vektor njihovih oznaka $\mathbf{y}_{N\times 1}$. Oznake se generiraju kao $y^{(i)} = f(x^{(i)})+\mathcal{N}(0,\sigma^2)$, gdje je $f:\mathbb{R}^n\to\mathbb{R}$ stvarna funkcija koja je generirala podatke (koja nam je u stvarnosti nepoznata), a $\sigma$ je standardna devijacija Gaussovog šuma, definirana parametrom `noise`. Za generiranje šuma možete koristiti funkciju [`numpy.random.normal`](https://docs.scipy.org/doc/numpy-1.15.0/reference/generated/numpy.random.normal.html).
#
# Generirajte skup za učenje od $N=50$ primjera uniformno distribuiranih u intervalu $[-5,5]$ pomoću funkcije $f(x) = 5 + x -2 x^2 -5 x^3$ uz šum $\sigma=200$:
# +
from numpy.random import normal
def f(x):
return 10*x**3 + 2*x**2 + x + 1
def make_labels(X, f, noise=0) :
# Vaš kôd ovdje
y = list()
for i in range(len(X)):
y.append( f(X[i][0]) + normal(0,noise) )
return y
# -
def make_instances(x1, x2, N) :
return np.array([np.array([x]) for x in np.linspace(x1,x2,N)])
# Prikažite taj skup funkcijom [`scatter`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.scatter).
# Vaš kôd ovdje
X = make_instances(-5,5,50)
y = make_labels(X,f,500)
plt.scatter(X,y)
plt.plot(X,f(X),"r",label = "Function f(x)")
plt.legend()
# ### (b)
# Trenirajte model polinomijalne regresije stupnja $d=3$. Na istom grafikonu prikažite naučeni model $h(\mathbf{x})=\mathbf{w}^\intercal\tilde{\mathbf{x}}$ i primjere za učenje. Izračunajte pogrešku učenja modela.
# +
# Vaš kôd ovdje
from sklearn.linear_model import Ridge
from sklearn.preprocessing import PolynomialFeatures
X = make_instances(-5,5,50)
y = make_labels(X,f,700)
poly = PolynomialFeatures(2)
X_poly = poly.fit_transform(X)
model = Ridge(alpha = 100)
model.fit(X_poly,y)
error = mean_squared_error(y,h)
print(error)
#plt.scatter(X,y)
plt.plot(model.predict(X_poly),label = "Nauceni model regresije")
plt.plot(X,f(X),label = "Stvarna funkcija")
plt.legend()
# -
# ### 3. Odabir modela
# ### (a)
#
# Na skupu podataka iz zadatka 2 trenirajte pet modela linearne regresije $\mathcal{H}_d$ različite složenosti, gdje je $d$ stupanj polinoma, $d\in\{1,3,5,10,20\}$. Prikažite na istome grafikonu skup za učenje i funkcije $h_d(\mathbf{x})$ za svih pet modela (preporučujemo koristiti `plot` unutar `for` petlje). Izračunajte pogrešku učenja svakog od modela.
#
# **Q:** Koji model ima najmanju pogrešku učenja i zašto?
# +
# Vaš kôd ovdje
X = make_instances(-5,5,50)
y = make_labels(X,f,200)
ds = [1,3,5,10,20]
plt.figure(figsize=(15,10))
j = 0
for i in range(len(ds)):
poly = PolynomialFeatures(ds[i])
X_i = poly.fit_transform(X)
w = matmul(pinv(X_i),y)
h = matmul(X_i,w)
print(mean_squared_error(y,h))
j=j+1
plt.subplot(2,3,j)
plt.grid()
plt.plot(X,h,label = "d = "+str(ds[i]))
plt.plot(X,f(X),label = "f(x)")
plt.scatter(X,y,c = "r")
plt.legend()
# -
# ### (b)
# Razdvojite skup primjera iz zadatka 2 pomoću funkcije [`model_selection.train_test_split`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) na skup za učenja i skup za ispitivanje u omjeru 1:1. Prikažite na jednom grafikonu pogrešku učenja i ispitnu pogrešku za modele polinomijalne regresije $\mathcal{H}_d$, sa stupnjem polinoma $d$ u rasponu $d\in [1,2,\ldots,20]$. Budući da kvadratna pogreška brzo raste za veće stupnjeve polinoma, umjesto da iscrtate izravno iznose pogrešaka, iscrtajte njihove logaritme.
#
# **NB:** Podjela na skupa za učenje i skup za ispitivanje mora za svih pet modela biti identična.
#
# **Q:** Je li rezultat u skladu s očekivanjima? Koji biste model odabrali i zašto?
#
# **Q:** Pokrenite iscrtavanje više puta. U čemu je problem? Bi li problem bio jednako izražen kad bismo imali više primjera? Zašto?
from sklearn.model_selection import train_test_split
# +
# Vaš kôd ovdje
X = make_instances(-5,5,50)
y = make_labels(X,f,200)
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size = 0.5)
train_errors = list()
test_errors = list()
for i in range(20):
poly = PolynomialFeatures(i)
x_i_train = poly.fit_transform(X_train)
x_i_test = poly.fit_transform(X_test)
w = matmul(pinv(x_i_train),y_train)
h_train = matmul(x_i_train,w)
h_test = matmul(x_i_test,w)
train_errors.append(log(mean_squared_error(y_train,h_train)))
test_errors.append(log(mean_squared_error(y_test,h_test)))
plt.plot(train_errors,label = "Train errors")
plt.plot(test_errors,label = "Test errors")
plt.legend()
# -
# ### (c)
# Točnost modela ovisi o (1) njegovoj složenosti (stupanj $d$ polinoma), (2) broju primjera $N$, i (3) količini šuma. Kako biste to analizirali, nacrtajte grafikone pogrešaka kao u 3b, ali za sve kombinacija broja primjera $N\in\{100,200,1000\}$ i količine šuma $\sigma\in\{100,200,500\}$ (ukupno 9 grafikona). Upotrijebite funkciju [`subplots`](http://matplotlib.org/examples/pylab_examples/subplots_demo.html) kako biste pregledno posložili grafikone u tablicu $3\times 3$. Podatci se generiraju na isti način kao u zadatku 2.
#
# **NB:** Pobrinite se da svi grafikoni budu generirani nad usporedivim skupovima podataka, na sljedeći način. Generirajte najprije svih 1000 primjera, podijelite ih na skupove za učenje i skupove za ispitivanje (dva skupa od po 500 primjera). Zatim i od skupa za učenje i od skupa za ispitivanje načinite tri različite verzije, svaka s drugačijom količinom šuma (ukupno 2x3=6 verzija podataka). Kako bi simulirali veličinu skupa podataka, od tih dobivenih 6 skupova podataka uzorkujte trećinu, dvije trećine i sve podatke. Time ste dobili 18 skupova podataka -- skup za učenje i za testiranje za svaki od devet grafova.
# +
# <NAME>
plt.figure(figsize=(15,10))
Ns = [50,100,500]
noises = [100,200,500]
X = make_instances(-5,5,1000)
l=0
for i in noises:
y_i = make_labels(X,f,i)
for j in Ns:
X_train,X_test,y_train,y_test = train_test_split(X,y_i,test_size = 0.5)
l=l+1
X_i_train = X_train[:j]
X_i_test = X_test[:j]
y_i_train = y_train[:j]
y_i_test = y_test[:j]
train_errors = []
test_errors = []
for k in range(1,21):
poly = PolynomialFeatures(k)
X_train_poly = poly.fit_transform(X_i_train)
X_test_poly = poly.fit_transform(X_i_test)
w = matmul(pinv(X_train_poly),y_i_train)
h_train = matmul(X_train_poly,w)
h_test = matmul(X_test_poly,w)
train_errors.append(log(mean_squared_error(y_i_train,h_train)))
test_errors.append(log(mean_squared_error(y_i_test,h_test)))
plt.subplot(3,3,l)
plt.plot(train_errors, label = "Train")
plt.plot(test_errors, label = "Test")
plt.legend()
plt.title("N = "+str(j*2)+" "+" D = "+str(i))
# -
# ***Q:*** Jesu li rezultati očekivani? Obrazložite.
# ### 4. Regularizirana regresija
# ### (a)
#
# U gornjim eksperimentima nismo koristili **regularizaciju**. Vratimo se najprije na primjer iz zadatka 1. Na primjerima iz tog zadatka izračunajte težine $\mathbf{w}$ za polinomijalni regresijski model stupnja $d=3$ uz L2-regularizaciju (tzv. *ridge regression*), prema izrazu $\mathbf{w}=(\mathbf{\Phi}^\intercal\mathbf{\Phi}+\lambda\mathbf{I})^{-1}\mathbf{\Phi}^\intercal\mathbf{y}$. Napravite izračun težina za regularizacijske faktore $\lambda=0$, $\lambda=1$ i $\lambda=10$ te usporedite dobivene težine.
#
# **Q:** Kojih je dimenzija matrica koju treba invertirati?
#
# **Q:** Po čemu se razlikuju dobivene težine i je li ta razlika očekivana? Obrazložite.
# +
# Vaš kôd ovdje
X = np.array([[0],[1],[2],[4]])
y = np.array([4,1,2,5])
d = 3
poly = PolynomialFeatures(d)
X_poly = poly.fit_transform(X)
I_matrix = np.identity(d+1)
I_matrix[0][0] = 0
plt.scatter(X,y)
for gamma in [0,1,10]:
pseudoinvers = matmul(X_poly.T,X_poly)+gamma*I_matrix
w = matmul(matmul(inv(pseudoinvers),X_poly.T),y)
h = matmul(X_poly,w)
plt.plot(h,label = "Gamma = "+str(gamma))
plt.legend()
print(w)
# -
# ### (b)
# Proučite klasu [`Ridge`](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Ridge.html#sklearn.linear_model.Ridge) iz modula [`sklearn.linear_model`](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.linear_model), koja implementira L2-regularizirani regresijski model. Parametar $\alpha$ odgovara parametru $\lambda$. Primijenite model na istim primjerima kao u prethodnom zadatku i ispišite težine $\mathbf{w}$ (atributi `coef_` i `intercept_`).
#
# **Q:** Jesu li težine identične onima iz zadatka 4a? Ako nisu, objasnite zašto je to tako i kako biste to popravili.
from sklearn.linear_model import Ridge
# +
# Vaš kôd ovdje
for gamma in [0,1,10]:
model = Ridge(alpha = gamma)
model.fit(X_poly,y)
print([(model.intercept_,model.coef_[1],model.coef_[2],model.coef_[3])])
# -
# ### 5. Regularizirana polinomijalna regresija
# ### (a)
#
# Vratimo se na slučaj $N=50$ slučajno generiranih primjera iz zadatka 2. Trenirajte modele polinomijalne regresije $\mathcal{H}_{\lambda,d}$ za $\lambda\in\{0,100\}$ i $d\in\{2,10\}$ (ukupno četiri modela). Skicirajte pripadne funkcije $h(\mathbf{x})$ i primjere (na jednom grafikonu; preporučujemo koristiti `plot` unutar `for` petlje).
#
# **Q:** Jesu li rezultati očekivani? Obrazložite.
# +
# Vaš kôd ovdje
plt.figure(figsize=(15,10))
X = make_instances(-5,5,50)
y = make_labels(X,f,200)
ds = [2,10]
gammas = [0,100]
i = 0
for d in ds:
poly = PolynomialFeatures(d)
X_poly = poly.fit_transform(X)
I_matrix = np.identity(d+1)
I_matrix[0][0] = 0
for gamma in gammas:
pseudoinvers = matmul(X_poly.T,X_poly)+gamma*I_matrix
w = matmul(matmul(inv(pseudoinvers),X_poly.T),y)
h = matmul(X_poly,w)
i = i+1
plt.subplot(2,2,i)
plt.plot(X,h)
plt.scatter(X,y)
plt.title("Gamma = "+str(gamma)+" "+" Stupanj polinoma = "+str(d))
# -
# ### (b)
#
# Kao u zadataku 3b, razdvojite primjere na skup za učenje i skup za ispitivanje u omjeru 1:1. Prikažite krivulje logaritama pogreške učenja i ispitne pogreške u ovisnosti za model $\mathcal{H}_{d=10,\lambda}$, podešavajući faktor regularizacije $\lambda$ u rasponu $\lambda\in\{0,1,\dots,50\}$.
#
# **Q:** Kojoj strani na grafikonu odgovara područje prenaučenosti, a kojoj podnaučenosti? Zašto?
#
# **Q:** Koju biste vrijednosti za $\lambda$ izabrali na temelju ovih grafikona i zašto?
#
# +
# Vaš kôd ovdje
from sklearn.linear_model import Ridge
X = make_instances(-5,5,50)
y = make_labels(X,f,200)
gammas = [i for i in range(50)]
poly = PolynomialFeatures(10)
X_poly = poly.fit_transform(X)
X_train,X_test,y_train,y_test = train_test_split(X_poly,y,test_size = 0.5)
train_errors = []
test_errors = []
for gamma in gammas:
model = Ridge(alpha = gamma)
model.fit(X_train,y_train)
train_errors.append(log(mean_squared_error(y_train,model.predict(X_train))))
test_errors.append(log(mean_squared_error(y_test,model.predict(X_test))))
plt.plot(train_errors,label = "Train error")
plt.plot(test_errors,label = "Test error")
plt.legend()
# -
# ### 6. L1-regularizacija i L2-regularizacija
# Svrha regularizacije jest potiskivanje težina modela $\mathbf{w}$ prema nuli, kako bi model bio što jednostavniji. Složenost modela može se okarakterizirati normom pripadnog vektora težina $\mathbf{w}$, i to tipično L2-normom ili L1-normom. Za jednom trenirani model možemo izračunati i broj ne-nul značajki, ili L0-normu, pomoću sljedeće funkcije koja prima vektor težina $\mathbf{w}$:
def nonzeroes(coef, tol=1e-2):
return len(coef) - len(coef[np.isclose(0, coef, atol=tol)])
# ### (a)
#
# Za ovaj zadatak upotrijebite skup za učenje i skup za testiranje iz zadatka 3b. Trenirajte modele **L2-regularizirane** polinomijalne regresije stupnja $d=10$, mijenjajući hiperparametar $\lambda$ u rasponu $\{1,2,\dots,100\}$. Za svaki od treniranih modela izračunajte L{0,1,2}-norme vektora težina $\mathbf{w}$ te ih prikažite kao funkciju od $\lambda$. Pripazite što točno šaljete u funkciju za izračun normi.
#
# **Q:** Objasnite oblik obiju krivulja. Hoće li krivulja za $\|\mathbf{w}\|_2$ doseći nulu? Zašto? Je li to problem? Zašto?
#
# **Q:** Za $\lambda=100$, koliki je postotak težina modela jednak nuli, odnosno koliko je model rijedak?
# +
# Vaš kôd ovdje
from sklearn.linear_model import Ridge
gammas = [i for i in range(100)]
X = make_instances(-5,5,50)
y = make_labels(X,f,200)
poly = PolynomialFeatures(10)
X_poly = poly.fit_transform(X)
l2_norm = list()
l1_norm = list()
l0_norm = list()
for gamma in gammas:
model = Ridge(alpha = gamma)
model.fit(X_poly,y)
w = model.coef_
l2_norm.append(norm(w,ord = 2))
l1_norm.append(norm(w,ord = 1))
l0_norm.append(nonzeroes(w))
plt.figure(figsize(15,5))
plt.grid()
plt.plot(gammas,l2_norm,label = "L2 - norm ")
plt.plot(gammas, l1_norm, label = "L1 - norm ")
plt.plot(gammas, l0_norm, label = "L0 - norm ")
plt.legend()
# -
# ### (b)
# Glavna prednost L1-regularizirane regresije (ili *LASSO regression*) nad L2-regulariziranom regresijom jest u tome što L1-regularizirana regresija rezultira **rijetkim modelima** (engl. *sparse models*), odnosno modelima kod kojih su mnoge težine pritegnute na nulu. Pokažite da je to doista tako, ponovivši gornji eksperiment s **L1-regulariziranom** regresijom, implementiranom u klasi [`Lasso`](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Lasso.html) u modulu [`sklearn.linear_model`](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.linear_model). Zanemarite upozorenja.
# +
# Vaš kôd ovdje
from sklearn.linear_model import Lasso
gammas = [i for i in range(100)]
X = make_instances(-5,5,50)
y = make_labels(X,f,200)
poly = PolynomialFeatures(10)
X_poly = poly.fit_transform(X)
l2_norm = list()
l1_norm = list()
l0_norm = list()
for gamma in gammas:
model = Lasso(alpha = gamma+1)
model.fit(X_poly,y)
w = model.coef_
l2_norm.append(norm(w,ord = 2))
l1_norm.append(norm(w,ord = 1))
l0_norm.append(nonzeroes(w))
plt.figure(figsize(15,5))
plt.grid()
plt.plot(gammas,l2_norm,label = "L2 - norm ")
plt.plot(gammas, l1_norm, label = "L1 - norm ")
plt.plot(gammas, l0_norm, label = "L0 - norm ")
plt.legend()
# -
# ### 7. Značajke različitih skala
# Često se u praksi možemo susreti sa podatcima u kojima sve značajke nisu jednakih magnituda. Primjer jednog takvog skupa je regresijski skup podataka `grades` u kojem se predviđa prosjek ocjena studenta na studiju (1--5) na temelju dvije značajke: bodova na prijamnom ispitu (1--3000) i prosjeka ocjena u srednjoj školi. Prosjek ocjena na studiju izračunat je kao težinska suma ove dvije značajke uz dodani šum.
#
# Koristite sljedeći kôd kako biste generirali ovaj skup podataka.
# +
n_data_points = 500
np.random.seed(69)
# Generiraj podatke o bodovima na prijamnom ispitu koristeći normalnu razdiobu i ograniči ih na interval [1, 3000].
exam_score = np.random.normal(loc=1500.0, scale = 500.0, size = n_data_points)
exam_score = np.round(exam_score)
exam_score[exam_score > 3000] = 3000
exam_score[exam_score < 0] = 0
# Generiraj podatke o ocjenama iz srednje škole koristeći normalnu razdiobu i ograniči ih na interval [1, 5].
grade_in_highschool = np.random.normal(loc=3, scale = 2.0, size = n_data_points)
grade_in_highschool[grade_in_highschool > 5] = 5
grade_in_highschool[grade_in_highschool < 1] = 1
# Matrica dizajna.
grades_X = np.array([exam_score,grade_in_highschool]).T
# Završno, generiraj izlazne vrijednosti.
rand_noise = np.random.normal(loc=0.0, scale = 0.5, size = n_data_points)
exam_influence = 0.9
grades_y = ((exam_score / 3000.0) * (exam_influence) + (grade_in_highschool / 5.0) \
* (1.0 - exam_influence)) * 5.0 + rand_noise
grades_y[grades_y < 1] = 1
grades_y[grades_y > 5] = 5
# -
# **a)** Iscrtajte ovisnost ciljne vrijednosti (y-os) o prvoj i o drugoj značajki (x-os). Iscrtajte dva odvojena grafa.
# +
# Vaš kôd ovdje
plt.figure(figsize = (15,10))
plt.subplot(1,2,1)
plt.scatter(grades_X[:,0],grades_y)
plt.title("Ovisnost prosjecne ocjene o broju bodova na prijamnom")
plt.subplot(1,2,2)
plt.scatter(grades_X[:,1],grades_y)
plt.title("Ovisnost prosjecne ocjene o broju ocjeni u srednjoj")
# -
# **b)** Naučite model L2-regularizirane regresije ($\lambda = 0.01$), na podacima `grades_X` i `grades_y`:
# +
# Vaš kôd ovdje
from sklearn.linear_model import Ridge
model = Ridge(alpha = 0.01)
model.fit(grades_X,grades_y)
print(model.coef_)
print(model.intercept_)
plt.subplot(1,2,1)
plt.scatter(grades_X[:,0],model.predict(grades_X))
plt.subplot(1,2,2)
plt.scatter(grades_X[:,1],model.predict(grades_X))
# -
# Sada ponovite gornji eksperiment, ali prvo skalirajte podatke `grades_X` i `grades_y` i spremite ih u varijable `grades_X_fixed` i `grades_y_fixed`. Za tu svrhu, koristite [`StandardScaler`](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html).
from sklearn.preprocessing import StandardScaler
# +
# Vaš kôd ovdje
from sklearn.linear_model import Ridge
scaler = StandardScaler()
grades_X_fixed = scaler.fit_transform(grades_X)
grades_y_fixed = scaler.fit_transform(grades_y.reshape(-1,1))
model = Ridge(alpha = 0.01)
model.fit(grades_X_fixed,grades_y_fixed)
print(model.coef_)
print(model.intercept_)
plt.subplot(1,2,1)
plt.scatter(grades_X_fixed[:,0],model.predict(grades_X_fixed))
plt.subplot(1,2,2)
plt.scatter(grades_X_fixed[:,1],model.predict(grades_X_fixed))
# -
# **Q:** Gledajući grafikone iz podzadatka (a), koja značajka bi trebala imati veću magnitudu, odnosno važnost pri predikciji prosjeka na studiju? Odgovaraju li težine Vašoj intuiciji? Objasnite.
# ### 8. Multikolinearnost i kondicija matrice
# **a)** Izradite skup podataka `grades_X_fixed_colinear` tako što ćete u skupu `grades_X_fixed` iz
# zadatka *7b* duplicirati zadnji stupac (ocjenu iz srednje škole). Time smo efektivno uveli savršenu multikolinearnost.
# Vaš kôd ovdje
extra_column = grades_X_fixed[:,1]
extra_column = extra_column.reshape(-1,1)
grades_X_fixed_colinear = np.append(grades_X_fixed,extra_column,axis=1)
print(grades_X_fixed_colinear)
# Ponovno, naučite na ovom skupu L2-regularizirani model regresije ($\lambda = 0.01$).
# Vaš kôd ovdje
from sklearn.linear_model import Ridge
model = Ridge(alpha = 0.01)
model.fit(grades_X_fixed_colinear,grades_y_fixed)
print(model.coef_,model.intercept_)
# **Q:** Usporedite iznose težina s onima koje ste dobili u zadatku *7b*. Što se dogodilo?
# **b)** Slučajno uzorkujte 50% elemenata iz skupa `grades_X_fixed_colinear` i naučite dva modela L2-regularizirane regresije, jedan s $\lambda=0.01$ i jedan s $\lambda=1000$). Ponovite ovaj pokus 10 puta (svaki put s drugim podskupom od 50% elemenata). Za svaki model, ispišite dobiveni vektor težina u svih 10 ponavljanja te ispišite standardnu devijaciju vrijednosti svake od težina (ukupno šest standardnih devijacija, svaka dobivena nad 10 vrijednosti).
# +
# Vaš kôd ovdje
from sklearn.linear_model import Ridge
gammas = [0.01,1000]
for gamma in gammas:
model = Ridge(alpha = gamma)
ws = []
for i in range(1,11):
grades_dataset_1,grades_dataset_2,grades_y_1,grades_y_2 = train_test_split(grades_X_fixed_colinear,grades_y,test_size = 0.5)
model.fit(grades_dataset_1,grades_y_1)
print(model.coef_)
ws.append(model.coef_)
print("")
w1 = list()
w2 = list()
w3 = list()
for j in range(10):
w1.append(ws[j][0])
w2.append(ws[j][1])
w3.append(ws[j][2])
print("Standard deviation w1: "+str(std(w1)))
print("Standard deviation w2: "+str(std(w2)))
print("Standard deviation w3: "+str(std(w3)))
print("")
# -
# **Q:** Kako regularizacija utječe na stabilnost težina?
# **Q:** Jesu li koeficijenti jednakih magnituda kao u prethodnom pokusu? Objasnite zašto.
# **c)** Koristeći [`numpy.linalg.cond`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.linalg.cond.html) izračunajte kondicijski broj matrice $\mathbf{\Phi}^\intercal\mathbf{\Phi}+\lambda\mathbf{I}$, gdje je $\mathbf{\Phi}$ matrica dizajna (`grades_X_fixed_colinear`). Ponovite i za $\lambda=0.01$ i za $\lambda=10$.
#
# +
# Vaš kôd ovdje
pseudoinvers = matmul(grades_X_fixed_colinear.T,grades_X_fixed_colinear)+0.01*diag([0,1,1])
conditional_number = cond(pseudoinvers)
print(conditional_number)
pseudoinvers = matmul(grades_X_fixed_colinear.T,grades_X_fixed_colinear)+10*diag([0,1,1])
conditional_number = cond(pseudoinvers)
print(conditional_number)
# -
# **Q:** Kako regularizacija utječe na kondicijski broj matrice $\mathbf{\Phi}^\intercal\mathbf{\Phi}+\lambda\mathbf{I}$?
X = np.array([[0.25],[0.5],[1],[1.5],[2]])
y = np.array([0.707,1,0,-1,0])
poly = PolynomialFeatures(4)
X_poly = poly.fit_transform(X)
print(X_poly)
clf = Ridge(alpha = 1)
clf.fit(X_poly,y)
plt.plot(X,clf.predict(X_poly))
plt.scatter(X,y)
# +
def fa(x):
return x**3
def make_labels(X, f, noise=0) :
# Vaš kôd ovdje
y = list()
for i in range(len(X)):
y.append( f(X[i]) + normal(0,noise) )
return y
plt.figure(figsize=(15,10))
points = sp.linspace(-10,10)
y = make_labels(points,fa,500)
plt.plot(points,fa(points))
plt.scatter(points,y)
X = make_instances(-10,10,50)
clf = Ridge(alpha = 1000)
poly = PolynomialFeatures(5)
X_poly = poly.fit_transform(X)
clf.fit(X_poly,y)
plt.plot(X,clf.predict(X_poly))
# +
from sklearn.linear_model import LinearRegression
X = np.array([[1,-3,1],[1,-3,3],[1,1,2],[1,2,1],[1,1,-2],[1,2,-3]])
y0 = np.array([1,1,0,0,0,0])
y1 = np.array([0,0,1,1,0,0])
y2 = np.array([0,0,0,0,1,1])
clf = LinearRegression()
clf.fit(X,y0)
print(clf.coef_)
print(clf.intercept_)
# -
X = np.array([])
| SU-2019-LAB01-<0036500216>.ipynb |