code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Approximate-Nearest-Neighborhood-Search-with-Navigable-Small-World" data-toc-modified-id="Approximate-Nearest-Neighborhood-Search-with-Navigable-Small-World-1"><span class="toc-item-num">1 </span>Approximate Nearest Neighborhood Search with Navigable Small World</a></span><ul class="toc-item"><li><span><a href="#Data-Preparation-and-Model" data-toc-modified-id="Data-Preparation-and-Model-1.1"><span class="toc-item-num">1.1 </span>Data Preparation and Model</a></span></li><li><span><a href="#Navigable-Small-World" data-toc-modified-id="Navigable-Small-World-1.2"><span class="toc-item-num">1.2 </span>Navigable Small World</a></span></li><li><span><a href="#Hnswlib" data-toc-modified-id="Hnswlib-1.3"><span class="toc-item-num">1.3 </span>Hnswlib</a></span></li></ul></li><li><span><a href="#Reference" data-toc-modified-id="Reference-2"><span class="toc-item-num">2 </span>Reference</a></span></li></ul></div>
# +
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir(os.path.join('..', '..', 'notebook_format'))
from formats import load_style
load_style(plot_style=False)
# +
os.chdir(path)
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
# 4. magic to enable retina (high resolution) plots
# https://gist.github.com/minrk/3301035
# %matplotlib inline
# %load_ext watermark
# %load_ext autoreload
# %autoreload 2
# %config InlineBackend.figure_format='retina'
import time
import fasttext
import numpy as np
import pandas as pd
# prevent scientific notations
pd.set_option('display.float_format', lambda x: '%.3f' % x)
# %watermark -a 'Ethen' -d -t -v -p numpy,pandas,fasttext,scipy
# -
# # Approximate Nearest Neighborhood Search with Navigable Small World
# Performing nearest neighborhood search on embeddings has become a crucial process in many applications, such as similar image/text search. The [ann benchmark](https://github.com/erikbern/ann-benchmarks) contains benchmark on various approximate nearest neighborhood search algorithms/libraries and in this document, we'll take a look at one of them, **Navigable Small World Graph**.
# ## Data Preparation and Model
# For the embedding, we'll be training a fasttext multi-label text classification model ourselves, and using the output embedding for this example. The fasttext library has already been introduced in another post, hence we won't be going over it in detail. The readers can also swap out the data preparation and model section with the embedding of their liking.
# +
# download the data and un-tar it under the 'data' folder
# -P or --directory-prefix specifies which directory to download the data to
# !wget https://dl.fbaipublicfiles.com/fasttext/data/cooking.stackexchange.tar.gz -P data
# -C specifies the target directory to extract an archive to
# !tar xvzf data/cooking.stackexchange.tar.gz -C data
# -
# !head -n 3 data/cooking.stackexchange.txt
# +
# train/test split
import os
from fasttext_module.split import train_test_split_file
from fasttext_module.utils import prepend_file_name
data_dir = 'data'
test_size = 0.2
input_path = os.path.join(data_dir, 'cooking.stackexchange.txt')
input_path_train = prepend_file_name(input_path, 'train')
input_path_test = prepend_file_name(input_path, 'test')
random_state = 1234
encoding = 'utf-8'
train_test_split_file(input_path, input_path_train, input_path_test,
test_size, random_state, encoding)
print('train path: ', input_path_train)
print('test path: ', input_path_test)
# +
# train the fasttext model
fasttext_params = {
'input': input_path_train,
'lr': 0.1,
'lrUpdateRate': 1000,
'thread': 8,
'epoch': 15,
'wordNgrams': 1,
'dim': 80,
'loss': 'ova'
}
model = fasttext.train_supervised(**fasttext_params)
print('vocab size: ', len(model.words))
print('label size: ', len(model.labels))
print('example vocab: ', model.words[:5])
print('example label: ', model.labels[:5])
# -
# model.get_input_matrix().shape
print('output matrix shape: ', model.get_output_matrix().shape)
model.get_output_matrix()
# Given the output matrix, we would like to compute each of its nearest neighbors using the compressed vectors.
#
# For those that are more interested in using some other embeddings, replace the `index_factors` with the embedding, and `query_factors` with a random element from that set of embeddings, and the rest of the document should still function properly.
# +
# we'll get one of the labels to find its nearest neighbors
label_id = 0
print(model.labels[label_id])
index_factors = model.get_output_matrix()
query_factors = model.get_output_matrix()[label_id]
query_factors.shape
# -
# ## Navigable Small World
# We'll start off by formally defining the problem. k-nearest neighbor search is a problem where given a query object $q$ we need to find the $k$ closest objects from a fixed set of objects $O \in D$, where $D$ is the set of all possible objects at hand.
#
# The idea behind navigable small world is to use a graph data structure $G(V, E)$ to represent these objects $O$, where every object $o_i$ is represented by a vertex/node $v_i$. The navigable small world graph structure is constructed by sequential addition of all elements. For every new element, we find the set of its closest neighbors using a variant of the greedy search algorithm, upon doing so, we'll then introduce a bidirectional connection between that set of neighbors and the incoming element.
#
# Upon building the graph, searching for the closest objects to $q$ is very similar to adding objects to the graph. i.e. It involves traversing through the graph to find the closest vertices/nodes using the same variant of greedy search algorithm that's used when constructing the graph.
#
# Another thing worth noting is that determining closest neighbors is dependent on a distance function. As the algorithm doesn't make any strong assumption about the data, it can be used on any distance function of our likings. Here we'll be using the cosine distance as an illustration.
class Node:
"""
Node for a navigable small world graph.
Parameters
----------
idx : int
For uniquely identifying a node.
value : 1d np.ndarray
To access the embedding associated with this node.
neighborhood : set
For storing adjacent nodes.
References
----------
https://book.pythontips.com/en/latest/__slots__magic.html
https://hynek.me/articles/hashes-and-equality/
"""
__slots__ = ['idx', 'value', 'neighborhood']
def __init__(self, idx, value):
self.idx = idx
self.value = value
self.neighborhood = set()
def __hash__(self):
return hash(self.idx)
def __eq__(self, other):
return (
self.__class__ == other.__class__ and
self.idx == other.idx
)
# +
from scipy.spatial import distance
def build_nsw_graph(index_factors, k):
n_nodes = index_factors.shape[0]
graph = []
for i, value in enumerate(index_factors):
node = Node(i, value)
graph.append(node)
for node in graph:
query_factor = node.value.reshape(1, -1)
# note that the following implementation is not the actual procedure that's
# used to find the k closest neighbors, we're just implementing a quick version,
# will come back to this later
# https://codereview.stackexchange.com/questions/55717/efficient-numpy-cosine-distance-calculation
# the smaller the cosine distance the more similar, thus the most
# similar item will be the first element after performing argsort
# since argsort by default sorts in ascending order
dist = distance.cdist(index_factors, query_factor, metric='cosine').ravel()
neighbors_indices = np.argsort(dist)[:k].tolist()
# insert bi-directional connection
node.neighborhood.update(neighbors_indices)
for i in neighbors_indices:
graph[i].neighborhood.add(node.idx)
return graph
# +
k = 10
graph = build_nsw_graph(index_factors, k)
graph[0].neighborhood
# -
# In the original paper, the author used the term "friends" of vertices that share an edge, and "friend list" of vertex $v_i$ for the list of vertices that share a common with the vertex $v_i$.
#
# We'll now introduce the variant of greedy search that the algorithm uses. The pseudocode looks like the following:
#
# ```
# greedy_search(q: object, v_entry_point: object):
# v_curr = v_entry_point
# d_min = dist_func(q, v_current)
# v_next = None
#
# for v_friend in v_curr.get_friends():
# d_friend = dist_func(q, v_friend)
# if d_friend < d_min:
# d_min = d_friend
# v_next = v_friend
#
# if v_next is None:
# return v_curr
# else:
# return greedy_search(q, v_next)
# ```
#
# Where starting from some entry point (chosen at random at the beginning), the greedy search algorithm computes a distance from the input query to each of the current entry point's friend vertices. If the distance between the query and the friend vertex is smaller than the current ones, then the greedy search algorithm will move to the vertex and repeats the process until it can't find a friend vertex that is closer to the query than the current vertex.
#
# This approach can of course lead to local minimum, i.e. the closest vertex/object determined by this greedy search algorithm is not the actual true closest element to the incoming query. Hence, the idea to extend this is to pick a series of entry point, denoted by `m` in the pseudocode below and return the best results from all those greedy searches. With each additional search, the chances of not finding the true nearest neighbors should decrease exponentially.
#
# The key idea behind the knn search is given a random entry point, it iterates on vertices closest to the query that we've never previously visited. And the algorithm keeps greedily exploring the neighborhood until the $k$ nearest elements can't be improved upon. Then this process repeats for the next random entry point.
#
# ```
# knn_search(q: object, m: int, k: int):
# queue[object] candidates, temp_result, result
# set[object] visited_set
#
# for i in range(m):
# put random entry point in candidates
# temp_result = None
#
# repeat:
# get element c closet from candidate to q
# remove c from candidates
#
# if c is further than the k-th element from result:
# break repeat
#
# for every element e from friends of c:
# if e is not visited_set:
# add e to visited_set, candidates, temp_result
#
#
# add objects from temp_result to result
#
# return best k elements from result
#
#
# ```
#
# We'll be using the [`heapq`](https://docs.python.org/3/library/heapq.html) module as our priority queue.
# +
import heapq
import random
from typing import List, Tuple
def nsw_knn_search(
graph: List[Node],
query: np.ndarray,
k: int=5,
m: int=50) -> Tuple[List[Tuple[float, int]], float]:
"""
Performs knn search using the navigable small world graph.
Parameters
----------
graph :
Navigable small world graph from build_nsw_graph.
query : 1d np.ndarray
Query embedding that we wish to find the nearest neighbors.
k : int
Number of nearest neighbors returned.
m : int
The recall set will be chosen from m different entry points.
Returns
-------
The list of nearest neighbors (distance, index) tuple.
and the average number of hops that was made during the search.
"""
result_queue = []
visited_set = set()
hops = 0
for _ in range(m):
# random entry point from all possible candidates
entry_node = random.randint(0, len(graph) - 1)
entry_dist = distance.cosine(query, graph[entry_node].value)
candidate_queue = []
heapq.heappush(candidate_queue, (entry_dist, entry_node))
temp_result_queue = []
while candidate_queue:
candidate_dist, candidate_idx = heapq.heappop(candidate_queue)
if len(result_queue) >= k:
# if candidate is further than the k-th element from the result,
# then we would break the repeat loop
current_k_dist, current_k_idx = heapq.nsmallest(k, result_queue)[-1]
if candidate_dist > current_k_dist:
break
for friend_node in graph[candidate_idx].neighborhood:
if friend_node not in visited_set:
visited_set.add(friend_node)
friend_dist = distance.cosine(query, graph[friend_node].value)
heapq.heappush(candidate_queue, (friend_dist, friend_node))
heapq.heappush(temp_result_queue, (friend_dist, friend_node))
hops += 1
result_queue = list(heapq.merge(result_queue, temp_result_queue))
return heapq.nsmallest(k, result_queue), hops / m
# -
results = nsw_knn_search(graph, query_factors, k=5)
results
# Now that we've implemented the knn search algorithm, we can go back and modify the graph building function and use it to implement the actual way of building the navigable small world graph.
def build_nsw_graph(index_factors: np.ndarray, k: int) -> List[Node]:
n_nodes = index_factors.shape[0]
graph = []
for i, value in enumerate(index_factors):
node = Node(i, value)
if i > k:
neighbors, hops = nsw_knn_search(graph, node.value, k)
neighbors_indices = [node_idx for _, node_idx in neighbors]
else:
neighbors_indices = list(range(i))
# insert bi-directional connection
node.neighborhood.update(neighbors_indices)
for i in neighbors_indices:
graph[i].neighborhood.add(node.idx)
graph.append(node)
return graph
# +
k = 10
index_factors = model.get_output_matrix()
graph = build_nsw_graph(index_factors, k)
graph[0].neighborhood
# -
results = nsw_knn_search(graph, query_factors, k=5)
results
# ## Hnswlib
# We can check the results with a more robust variant of the algorithm, [**Hierarchical Navigable Small World (HNSW)**](https://arxiv.org/abs/1603.09320) provided by [hnswlib](https://github.com/nmslib/hnswlib). The idea is very similar to the skip list data structure, except we now replace the link list with nagivable small world graphs. Although we never formally introduce the hierarchical variant, but hopefully the major parameters of the algorithm should look familiar.
#
# - `ef`: The algorithm searches for the `ef` closest neighbors to the inserted element $q$, this was set to $k$ in the original navigable small world paper. The `ef` closest neighbors then becomes the candidate/recall set for inserting the bidirectional edges during insertion/construction phase (which is termed `ef_construction`) or after done construction, the candidate/recall set for finding the actual top k closest elements to the input query object.
# - `M`: After choosing the `ef_construction` objects, only the `M` closest ones will we create the edges between the enter point and those nodes. i.e. it controls the number of bi-directional links.
#
# The actual process of constructing HNSW and doing knn search is a bit more involved compared to vanilla navigable small world. We won't be getting into all the gory details in this post.
# +
import hnswlib
def build_hnsw(factors, space, ef_construction, M):
# Declaring index
max_elements, dim = factors.shape
hnsw = hnswlib.Index(space, dim) # possible options for space are l2, cosine or ip
# Initing index - the maximum number of elements should be known beforehand
hnsw.init_index(max_elements, M, ef_construction)
# Element insertion (can be called several times)
hnsw.add_items(factors)
return hnsw
# +
space = 'cosine'
ef_construction = 200
M = 24
start = time.time()
hnsw = build_hnsw(index_factors, space, ef_construction, M)
build_time = time.time() - start
build_time
# +
k = 5
# Controlling the recall by setting ef, should always be > k
hnsw.set_ef(70)
# retrieve the top-n search neighbors
labels, distances = hnsw.knn_query(query_factors, k=k)
print(labels)
# -
# find the nearest neighbors and "translate" it to the original labels
[model.labels[label] for label in labels[0]]
# Based on the [ann benchmark](https://github.com/erikbern/ann-benchmarks), Hierarchical Navigable Small World (HNSW) stood out as one of the top performing approximate nearest neighborhood algorithms at the time of writing this document. Here, we introduced the vanilla variant of that algorithm, Navigable Small World and also matched the result with a more robust implementation from the open sourced library hnswlib.
# # Reference
# - [Github: Hnswlib - fast approximate nearest neighbor search](https://github.com/nmslib/hnswlib)
# - [Github: Navigable Small World Graphs For Approximate Nearest Neighbors In Rust](https://github.com/dkohlsdorf/NSWG)
# - [Paper: <NAME>, <NAME>, <NAME>, <NAME> - Approximate nearest neighbor algorithm based on navigable small world graphs (2014)](https://publications.hse.ru/mirror/pubs/share/folder/x5p6h7thif/direct/128296059)
# - [Paper: <NAME>, <NAME> - Efficient and robust approximate nearest neighbor search using Hierarchical Navigable Small World graphs (2016)](https://arxiv.org/abs/1603.09320)
| deep_learning/multi_label/nsw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
greetings = "<NAME>"
print(greetings)
# ### Libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# ### DataSet
df = pd.read_excel("Canada.xlsx", sheet_name = "Canada by Citizenship", skiprows = range(20), skipfooter = 2)
df.head()
# dimensions
df.shape
# Clean up the dataset to remove columns that are not informative to us for visualization (eg. Type, AREA, REG)
df.drop(["AREA", "REG", "DEV", "Type", "Coverage"], axis = 1, inplace = True)
df.head()
# Rename some of the columns so that they make sense.
df.rename(columns = {"OdName": "Country", "AreaName": "Continent", "RegName": "Region"}, inplace = True)
df.head()
# data types
df.dtypes
# Labels (int to str)
df.columns = list(map(str, df.columns))
df.columns
# Set the country name as index - useful for quickly looking up countries using .loc method.
df.set_index('Country', inplace=True)
df.head(5)
# Add total Immiogrants column
df["Total"] = df.sum(axis = 1)
# List of years (strings)
years = list(map(str, range(1980, 2014)))
years
# ### Visualizing Data using Matplotlib
# +
# Top 5 immigrant countries
df.sort_values(['Total'], ascending=False, axis=0, inplace=True)
# get the top 5 entries
df_top5 = df.head()
# transpose the dataframe
df_top5 = df_top5[years].transpose()
df_top5.head()
# +
# Plot
df_top5.plot(kind = "area", alpha = 0.25, stacked = False, figsize = (10, 6))
plt.title('Immigration Trend of Top 5 Countries')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
# -
# option 2: preferred option with more flexibility
ax = df_top5.plot(kind = "area", stacked = False, alpha = 0.25, figsize = (10, 6))
ax.set_title("Immigration trend of top 5 Countries")
ax.set_ylabel("Number of Immigrants")
ax.set_xlabel("Years")
plt.show()
# **stacked area plot of the 5 countries that contributed the least to immigration to Canada from 1980 to 2013**
df.sort_values(["Total"], ascending = True, axis = 0, inplace = True)
df_low5 = df.head()
df_low5 = df_low5[years].T
df_low5.head()
# Scripting Layer
df_low5.plot(kind = "area", stacked = True, alpha = 0.45, figsize = (10, 6))
plt.title("Immigration Trend of least 5 Countries")
plt.xlabel("Years")
plt.ylabel("Number of Immigrants")
plt.show()
# Artist Layer
df_low5.plot(kind = "area", stacked = False, alpha = 0.55)
ax.set_title("Immigration Trend of least 5 countries")
ax.set_xlabel("Years")
ax.set_ylabel("Number of Immigrants")
plt.show()
# ### Histograms
# lets quickly view 2013 data
df[["2013"]].head()
# np.histogram returns 2 values
count, bin_edges = np.histogram(df["2013"])
print(count) # frequency
print(bin_edges) # bin ranges
# +
# bin_edges is a list of bin intervels
count, bin_edges = np.histogram(df["2013"])
df["2013"].plot(kind = "hist", figsize = (10, 6), xticks = bin_edges)
plt.title('Histogram of Immigration from 195 countries in 2013') # add a title to the histogram
plt.ylabel('Number of Countries') # add y-label
plt.xlabel('Number of Immigrants') # add x-label
plt.show()
# -
# **What is the immigration distribution for Denmark, Norway, and Sweden for years 1980 - 2013?**
# let's quickly view the dataset
df.loc[["Denmark", "Norway", "Sweden"], years]
# transpose DataFrame
df_t = df.loc[["Denmark", "Norway", "Sweden"], years].T
df_t.head()
# +
count, bin_edges = np.histogram(df_t, 15)
xmin = bin_edges[0] - 10 # first bin value is 31.0, adding buffer of 10 for aesthetic purposes
xmax = bin_edges[-1] + 10 # last bin value is 308.0, adding buffer of 10 for aesthetic purposes
# stacked Histogram
df_t.plot(kind='hist',
figsize=(10, 6),
bins=15,
xticks=bin_edges,
color=['coral', 'darkslateblue', 'mediumseagreen'],
stacked=True,
xlim=(xmin, xmax)
)
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
# -
# **immigration distribution for Greece, Albania, and Bulgaria for years 1980 - 2013**
df_gab = df.loc[["Greece", "Albania", "Bulgaria"], years]
df_T = df_gab.T
df_T.head()
# +
count, bin_edges = np.histogram(df_T, 15)
# Staked Histogram
df_T.plot(kind = "hist",
figsize = (10, 6),
alpha = 0.35,
bins = 15,
xticks = bin_edges,
color = ["coral", "darkslateblue", "mediumseagreen"])
plt.title("Histogram of Immigration from Denmark")
plt.ylabel("Number of Years")
plt.xlabel("Number of Immigrants")
plt.show()
# -
# ### Bar Charts (Dataframe)
# **Vertical Plot: number of Icelandic immigrants (country = 'Iceland') to Canada from year 1980 to 2013**
# step 1: get the data
iceland = df.loc["Iceland", years]
iceland.head()
# +
iceland.plot(kind='bar', figsize=(10, 6), rot=90)
plt.xlabel('Year')
plt.ylabel('Number of Immigrants')
plt.title('Icelandic Immigrants to Canada from 1980 to 2013')
# Annotate arrow
plt.annotate('', # s: str. will leave it blank for no text
xy=(32, 70), # place head of the arrow at point (year 2012 , pop 70)
xytext=(28, 20), # place base of the arrow at point (year 2008 , pop 20)
xycoords='data', # will use the coordinate system of the object being annotated
arrowprops=dict(arrowstyle='->', connectionstyle='arc3', color='blue', lw=2)
)
# Annotate Text
plt.annotate('2008 - 2011 Financial Crisis', # text to display
xy=(28, 30), # start the text at at point (year 2008 , pop 30)
rotation=72.5, # based on trial and error to match the arrow
va='bottom', # want the text to be vertically 'bottom' aligned
ha='left', # want the text to be horizontally 'left' algned.
)
plt.show()
# -
# **Horizontal Plot**
# Get the data pertaining to the top 15 countries.
df.sort_values(["Total"], ascending = False, axis = 0, inplace = True)
top_15 = df.head(15)
df_top_15 = top_15[years]
top_T = df_top_15.T
top_T
# Plot data
top_T.plot(kind='barh', figsize=(12, 12), color='steelblue')
plt.xlabel('Number of Immigrants')
plt.title('Top 15 Conuntries Contributing to the Immigration to Canada between 1980 - 2013')
plt.show()
df_pakistan = df[years].T
x = df_pakistan[["Pakistan"]]
print(x.idxmax(axis = 0))
print(x.max())
| Jupiter Book/Area Plots, Histograms, and Bar Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:4.2.0]
# language: python
# name: conda-env-4.2.0-py
# ---
# + deletable=true editable=true
# #!/usr/bin/env python
# Fitting of cmb measurements
from __future__ import division, print_function
import numpy as np
import matplotlib.pyplot as plt
#Load data set
Angle, Power = np.loadtxt('group4.txt', skiprows=3, comments='#', unpack=True)
Pcold = -23.240 #mdB +- 0.002
Phot = -20.460 #mdB +- 0.002
Thot = 5.6+273.16 #K +-0.05?
Tcold = 77.355 #k
Z_Angle = Angle-90
Phot2 = 10**(Phot/10)/1000 # From db to watt
Pcold2 = 10**(Pcold/10)/1000 # From db to watt
Power_watt = 10**(Power/10)/1000
#constants
Delta_nu = 1*10**9
T_R = 145.93
k = 1.3806*10**(-23)
Tau_zen = 0.01
#Calculated values
y = (10**(Phot/10))/(10**(Pcold/10)) # y factor
print('y =',y)
Trcvr = (Thot-(Tcold*y))/(y-1)
print('Trcvr =',Trcvr, 'K')
Gain = Phot2 / ((Thot+Trcvr)*k*Delta_nu)
print(Gain,'Gain in watts')
Gain_Dbm = 10*np.log10(1000*Gain) #From watt to db
print(Gain_Dbm, 'gain in Dbm')
Tsys = Power_watt/(k*Delta_nu*Gain)
#Gain_c = Pcold / ((Tcold+Trcvr)*k*delta_nu)
#print(Gain_c)
#Ptot = 10**((Power-30)/10) #Watts
fig = plt.figure()
frame1 = fig.add_subplot(1,1,1)
frame1.set_title("System temperature as a function of Zenith temperature")
frame1.set_xlabel('# Angle (deg)')
frame1.set_ylabel('Power (dBm)')
frame1.plot(Z_Angle,Tsys, c='r', label='Measurements')
frame1.axhline(Trcvr, c='b', label='Receiver Temperature')
leg = frame1.legend()
plt.show()
#"""
#Gain Directivity = 4*pi / d ohm)
#Fit guassian, Find Gain of the horn
#"""
# + deletable=true editable=true
p[mW]= 10**([dBmW/10])
# + deletable=true editable=true
| Kapteyn Radio Telescope.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="9RAVFL-lNGTo"
# This note provides basic idea to check the normality and understand the statistics of the return data. It can help you to link to our lectures on the lognormal and normal properties of stock price and returns.
# + [markdown] colab_type="text" id="puU6_VcLx58s"
# Load libraries
# + colab={} colab_type="code" id="ugmYbsrax58x" persistent_id="e5c76e7b-25f6-4b2b-a8f0-2a6adead2dd6"
import numpy as np # array operations
import pandas as pd # dataframe
import scipy.stats as scs
import statsmodels.api as sm
from pylab import plt
plt.style.use('ggplot')
# put all plots in the notebook itself
# %matplotlib inline
# -
#
# # 1. A stock
# + [markdown] colab_type="text" id="R6Qoyoz-Egwj"
# Download some stock data from the Yahoo Finance as we did in the first tutor.
# + colab={"base_uri": "https://localhost:8080/", "height": 225} colab_type="code" id="hZ1BNJWdDfCG" outputId="28e0fdbd-7cc7-4871-fb6d-634e3d8674e9"
from pandas_datareader import data
TSM = data.DataReader("TSM", start='2010-1-1', end='2019-12-31', data_source='yahoo')
TSM.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 600} colab_type="code" id="mDurKzyWEy0Z" outputId="27b24ff7-6b3d-46de-ba27-23d845cd59f4"
TSM['Close'].plot(figsize=(16, 10), grid=True, title='TSMC Close Price')
# + colab={"base_uri": "https://localhost:8080/", "height": 431} colab_type="code" id="UB25QrAeJvkS" outputId="f067edb5-7d5c-475c-b2d4-9f6f21095c53"
TSM['Return'] = np.log(TSM['Close'] / TSM['Close'].shift(1))
TSM
# + colab={"base_uri": "https://localhost:8080/", "height": 659} colab_type="code" id="crIuTdaRJ5RC" outputId="26455d9f-8b77-4557-e434-a1529564cf65"
TSM[['Close', 'Return']].hist(bins=50, figsize=(16, 10))
# + colab={"base_uri": "https://localhost:8080/", "height": 284} colab_type="code" id="FZ2Hy_z9KVE1" outputId="e2489b91-a119-4b57-d2b0-6df7c5e7034d"
TSM[['Close', 'Return']].describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="Nz39nwnbKdTs" outputId="91a787a2-96ae-4ecb-c3fb-386654e3f240"
scs.describe(TSM['Return'].dropna()) # to see skewness & kurtosis
# + [markdown] colab_type="text" id="gIVcdpnHLzoQ"
# Another good skim for the normality data is to check the Q-Q plot:
#
# + colab={"base_uri": "https://localhost:8080/", "height": 299} colab_type="code" id="bNjcJTFkL40W" outputId="9f70b60c-962e-46d0-e1ea-e2be3db3945d"
sm.qqplot(TSM['Return'].dropna(), line='s')
plt.grid(True)
plt.xlabel('theoretical quantiles')
plt.ylabel('sample quantiles')
# + [markdown] colab_type="text" id="rjozDA87MXJS"
# Lastly, we do the normality test. `scipy` gives us several functions to do the test:
#
# - [`skewtest`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.skewtest.html#scipy.stats.skewtest)
# - [`kurtosistest`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.kurtosistest.html#scipy.stats.kurtosistest)
# - [`normaltest`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.normaltest.html#scipy.stats.normaltest)
#
# + colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" id="8b9m7Gp3MZ5u" outputId="0ec6f8a8-7e68-40e4-fb18-77118a762fac"
def normality_tests(arr):
''' Tests for normality distribution of given data set.
Parameters
==========
array: ndarray
object to generate statistics on
'''
print("Skew of data set %14.3f" % scs.skew(arr))
print("Skew test p-value %14.3f" % scs.skewtest(arr)[1])
print("Kurt of data set %14.3f" % scs.kurtosis(arr))
print("Kurt test p-value %14.3f" % scs.kurtosistest(arr)[1])
print("Norm test p-value %14.3f" % scs.normaltest(arr)[1])
normality_tests(TSM['Return'].dropna())
# -
# Now, please read the results and conclude.
# # 2. The market index
#
# While a stock return may has some bias due to firm-specific risk, we may find normality from the market returns, which is the value-weighted returns from many stocks. Let's take the market returns from Fama-French database.
Factors5 = data.DataReader('F-F_Research_Data_5_Factors_2x3','famafrench', start='1925-01-01')
Factors5[0].head()
Factors5[0]['Mkt'] = Factors5[0]['Mkt-RF'] + Factors5[0]['RF'] # get market returns from excess returns
Factors5[0][['Mkt', 'Mkt-RF']].hist(bins=30, figsize=(16, 10))
# another way
import seaborn as sns
sns.distplot(Factors5[0][['Mkt']], hist=True, kde=True,
bins=int(180/5), color = 'darkblue',
hist_kws={'edgecolor':'black'},
kde_kws={'linewidth': 4})
sns.distplot(Factors5[0][['Mkt-RF']], hist=True, kde=True,
bins=int(180/5), color = 'darkblue',
hist_kws={'edgecolor':'black'},
kde_kws={'linewidth': 4})
# Let's check some statistics. We'll rely on the [`scipy.stats`](https://docs.scipy.org/doc/scipy/reference/stats.html) package.
Mkt = Factors5[0]['Mkt']
Mkt.describe()
scs.describe(Mkt)
scs.scoreatpercentile(Mkt, 5), scs.scoreatpercentile(Mkt, 95)
normality_tests(Mkt)
| 07_priceReturnDist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Israel Mapping Data
#
# For now just a pile of code fragments
# +
# get places data from gov mapping agency
import xml.etree.cElementTree as et
from urllib.request import urlopen
xml = et.fromstring(urlopen("https://www.mapi.gov.il/ProfessionalInfo/Documents/dataGov/CITY.xml").read())
# +
import pandas as pd
# covert into a dataframe
dfcols = ['name', 'code', 'ita_x', 'ita_y']
df_cities = pd.DataFrame(columns=dfcols)
for elem in xml.iter('Record'):
values = elem.find("Values").findall("Value");
x = float(values[1].find("X").text)
y = float(values[1].find("Y").text)
code = int(values[3].text);
name = values[8].text;
df_cities = df_cities.append(pd.Series([name, code, x, y], index=dfcols), ignore_index=True)
df_cities
# -
cities = [3000, 4000, 5000, 6000, 7000, 8000]
df_cities.loc[df_cities['code'].isin(cities)]
# +
from ipyleaflet import Map, Marker
# example for setting up a map
center = (32.046874,34.7455205)
m = Map(center=center, zoom=8)
marker = Marker(location=center, draggable=False)
m.add_layer(marker);
m
# -
from geopy.geocoders import Nominatim
geolocator = Nominatim(user_agent="dkddj")
location = geolocator.geocode("ירושלים")
print((location.latitude, location.longitude))
# +
def gps(name):
location = Nominatim(user_agent="dkddj").geocode("ירושלים")
return (location.latitude, location.longitude)
gps("ירושלים")
# +
#df2 = df['gps'] = df.apply(lambda x: gps(x['שם ישוב']), axis=1)
# -
| notebooks/israel-mapping-xml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="BKrIssRxE0Zf" colab={"base_uri": "https://localhost:8080/"} outputId="f7785d98-311d-46c3-dae4-ee3529dc052c"
import pandas as pd
pd.set_option('display.max_rows', 100)
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.express as px
# %load_ext rpy2.ipython
# + [markdown] id="QU-Unp2zNl5M"
# #**Introduction**
#
# In this tutorial, I will walk through a modeling example of how to use Python and R in the same notebook.
#
# The data set we are using is from Duke Univeristy on the growth of trees within Duke Forest. We want to infer what variables lead to tree growth from the data set.
# + [markdown] id="f2d2ruNiL9w5"
# # **Exploratory Data Analysis**
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="5pOHN0_BFD95" outputId="43c5c7dd-d939-4ceb-ccf3-54b160ac0f0a"
#import the data into a pandas DataFrame
df = pd.read_excel('trees.xlsx')
df
# + id="0Z3dkCJSxCgz" colab={"base_uri": "https://localhost:8080/"} outputId="4502ea14-656a-4b8a-b181-2c7a72accf5d"
#number of distinct trees
df['ID'].nunique()
# + id="wt2Le155GCsk" colab={"base_uri": "https://localhost:8080/"} outputId="1d60e329-4850-42da-ffbf-319c28ef3087"
#look at the value collected by year
df['yr'].value_counts().sort_index()
# + id="E9SHnsvMRXUh" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="69c799c3-1a57-4c41-e4c4-51aabde4bfa0"
#calculate the growth rates for the trees. Note that it will be recalculated further down in the notebook
df.sort_values(by = ['yr', 'ID'])
df['cm_growth'] = df.groupby(['ID'])['cm'].pct_change()
df
# + id="fjpamJsWHQoL" colab={"base_uri": "https://localhost:8080/", "height": 604} outputId="a51cffe8-61b2-4a19-ffab-c8c11fced386"
#correlation heatmap
plt.rcParams.update({'font.size':15})
plt.rcParams['figure.figsize'] = 15,10
corr_matrix = df.corr()
fig1 = sns.heatmap(corr_matrix, annot = True, cmap = 'Blues')
# + id="5yy1Ex_ccfXI" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ff414cac-77c2-4966-9973-a0992d235077"
#pairplot
fig2 = sns.pairplot(df);
# + id="C9pjrdJLa-pT" colab={"base_uri": "https://localhost:8080/", "height": 636} outputId="24cdfbc5-df83-4a85-ca2e-7f795c5acc8e"
plt.hist(x = df['cm'], lw = 2.5, edgecolor = 'purple', bins = 15)
plt.xticks(np.arange(0,round(df['cm'].max() + 1), 5))
plt.xlabel('Centimeters')
plt.ylabel('Observations')
plt.title('Trees by Diameter in Centimeters');
# + id="9cv0bPCVa-tS" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="c4ca4661-2ec6-4d03-a919-eaa4e79b49f0"
#categorize the trees by size
conditions = [(df['cm'] <= 2.54), (df['cm'] > 2.54) & (df['cm'] <= 12.45), (df['cm'] > 12.45) & (df['cm'] <= 22.6),(df['cm'] > 22.6)]
values = ['Seedlings', 'Saplings', 'Poletimbers', 'Sawtimbers']
df['Tree_Size_Category'] = np.select(conditions, values)
df
# + id="Af6dlwgJa-yz" colab={"base_uri": "https://localhost:8080/", "height": 615} outputId="47258183-26e7-48bd-a6f3-7d266659e0ce"
px.scatter(data_frame = df, x = 'summerpdsi', y = 'cm', color = 'Tree_Size_Category', trendline = 'ols')
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="eO9m1Ff-KEK2" outputId="39aed193-1857-42f6-efa3-9598f0d78daa"
px.scatter(data_frame = df, x = 'summerpdsi', y = 'cm_growth', color = 'Tree_Size_Category', trendline = 'ols')
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="rUOg_g2XKEO3" outputId="1a034b99-12e9-4c3a-f32c-3031900ea114"
px.scatter(data_frame = df, x = 'wintertemp', y = 'cm', color = 'Tree_Size_Category', trendline = 'ols')
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="pOPEgjJxKET0" outputId="b05e3060-1966-4c30-df75-9914ecc829de"
px.scatter(data_frame = df, x = 'wintertemp', y = 'cm_growth', color = 'Tree_Size_Category', trendline = 'ols')
# + colab={"base_uri": "https://localhost:8080/", "height": 990} id="6EpYhBl0BK1x" outputId="37456ebd-325b-4a20-c5e1-bdb54c75613f"
#take the average size of each group of trees by year
df_tree_categories = df.groupby(['Tree_Size_Category', 'yr']).mean()
df_tree_categories.drop(columns = ['ID', 'cm_growth'], inplace = True)
df_tree_categories.reset_index(inplace = True)
#growth rate annualized
df_tree_categories['cm_growth'] = df_tree_categories.groupby(['Tree_Size_Category'])['cm'].pct_change()
df_tree_categories['yr_difference'] = df_tree_categories.groupby(['Tree_Size_Category'])['yr'].diff()
df_tree_categories['cm_growth'] = df_tree_categories['cm_growth']/df_tree_categories['yr_difference']
df_tree_categories
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="2AucLmnpBK5B" outputId="6dcaf455-c064-46ac-8662-17239fe0fd40"
fig3 = px.scatter(data_frame = df_tree_categories, x = 'summerpdsi', y = 'cm', color = 'Tree_Size_Category', trendline = 'ols')
fig3
# + colab={"base_uri": "https://localhost:8080/", "height": 617} id="1NtS4UIOBK7l" outputId="7d8bc9c9-b24e-4803-f95f-93840e9fc195"
results = px.get_trendline_results(fig3)
print(results)
results.query("Tree_Size_Category == 'Seedlings'").px_fit_results.iloc[0].summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="y0MC4e4cBK-F" outputId="1bcaa690-f108-4828-cdb7-9f869275d57d"
fig4 = px.scatter(data_frame = df, x = 'summerpdsi', y = 'cm', color = 'Tree_Size_Category', trendline = 'ols')
fig4
# + colab={"base_uri": "https://localhost:8080/", "height": 544} id="nGY4Q98jBLAW" outputId="01614937-81ae-4124-b349-7d77900a9772"
results = px.get_trendline_results(fig4)
print(results)
results.query("Tree_Size_Category == 'Seedlings'").px_fit_results.iloc[0].summary()
# + id="xZW3-GWqBLDb"
df_cm = df_tree_categories.drop(columns = ['cm_growth', 'yr_difference'])
df_cm_growth = df_tree_categories.drop(columns = ['cm', 'yr_difference'])
df_cm_growth.dropna(inplace = True)
# + id="D2BqqGijBLF1"
df_cm.to_csv('df_cm.csv', index = False)
df_cm_growth.to_csv('df_cm_growth.csv', index = False)
# + [markdown] id="nXCyFq-IACRf"
# # **Modeling with R**
# + colab={"base_uri": "https://localhost:8080/"} id="w_lp9wv5eEYi" outputId="c0608316-b3e1-426a-c81d-ccf3962eb8bb" language="R"
# #install packages
# devtools::install_github("dustinfife/flexplot")
# install.packages("Metrics")
#
# #read in packages
# library(flexplot)
# library(lme4)
# library(Metrics)
# + colab={"base_uri": "https://localhost:8080/"} id="lTmmyhSXg4sY" outputId="8609ace0-1f0c-49ee-dc33-74ae82d4f299" language="R"
# #you can call R objects in different cells
# #load the csv files from pandas into an R DataFrame
# df_cm <- read.csv('df_cm.csv')
# df_cm_growth <- read.csv('df_cm_growth.csv')
# df_cm
# + colab={"base_uri": "https://localhost:8080/"} id="GROSJ7k8g4u5" outputId="9a814227-0640-4046-eb75-a7a00f95d205" language="R"
# #isolate the explanatory variables
# explanatory_variables <- names(df_cm)
# explanatory_variables <- explanatory_variables[-c(1,3)]
# explanatory_variables
# + colab={"base_uri": "https://localhost:8080/"} id="eBnOzPhVBMqK" outputId="6c74b1ba-af32-482e-83bb-e1714db8e064" language="R"
# #code from Stack Overflow, see references section
# n <- length(explanatory_variables)
#
# id <- unlist(
# lapply(1:n,
# function(i)combn(1:n,i,simplify=FALSE)
# )
# ,recursive=FALSE)
#
# Formulas <- sapply(id,function(i)
# paste("cm~",paste(explanatory_variables[i],collapse="+"))
# )
#
# #model out exhaustively all possible multiple linear regression models for the explanatory variables
# linear_models = lapply(Formulas, function(i)
# lm(as.formula(i),data = df_cm))
#
# linear_model_summaries = lapply(linear_models, summary)
#
# linear_model_summaries
# + colab={"base_uri": "https://localhost:8080/"} id="fp-NmNNfkaYa" outputId="834c526a-44c3-46aa-df35-22b8f9ca794a" language="R"
# Formulas <- sapply(id,function(i)
# paste("cm~",paste(explanatory_variables[i],collapse="+"),'+(1|Tree_Size_Category)')
# )
#
# #model out exhaustively all possible mixed effects models
# mixed_models = lapply(Formulas,function(i)
# lmer(as.formula(i),data = df_cm))
#
# mixed_model_summaries = lapply(mixed_models, summary)
#
# mixed_model_summaries
# + colab={"base_uri": "https://localhost:8080/"} id="EeJfT7nKkabZ" outputId="c3e29681-1855-4db5-8619-51094a466f84" language="R"
# Formulas_cm_growth <- sapply(id,function(i)
# paste("cm_growth~",paste(explanatory_variables[i],collapse="+"),'+(1|Tree_Size_Category)')
# )
#
# #model out exhaustively all possible mixed effects models for cm growth list
# mixed_models_cm_growth = lapply(Formulas_cm_growth,function(i)
# lmer(as.formula(i),data = df_cm_growth))
#
# mixed_model_summaries = lapply(mixed_models_cm_growth, summary)
#
# mixed_models_cm_growth
# + colab={"base_uri": "https://localhost:8080/"} id="sOk6cgpykads" outputId="9b47d3e0-483b-485c-f954-4b156b0f0f24" language="R"
# #BIC is used for model selection, the lower the value the better the model score
# lapply(linear_models, BIC)
# + colab={"base_uri": "https://localhost:8080/"} id="IQV1UQlRkagL" outputId="3be4da6b-be40-4c8d-c806-8e8a07e24e96" language="R"
# lapply(mixed_models, BIC)
# + colab={"base_uri": "https://localhost:8080/"} id="78Aqatt8kaiv" outputId="98dde346-e7a0-404b-b934-a5b58f09be19" language="R"
# #intraclass correlation shows how much variability is caused by clustering
# lapply(mixed_models, icc)
# + colab={"base_uri": "https://localhost:8080/"} id="ZlKbVZKGkalU" outputId="0541fc9b-0af7-43c3-cfd6-2a729d99a407" language="R"
# lapply(mixed_models_cm_growth, icc)
# + colab={"base_uri": "https://localhost:8080/"} id="NvbRcD6JkaoM" outputId="f4156ebe-5e48-468e-87da-edaa08fc0b09" language="R"
# #focus on the 3 best mixed models based on the Bayesian Information Criterion (BIC)
# #models 2, 3 and 9
# print(mse(df_cm$cm, predict(mixed_models[[2]], df_cm)))
# print(mse(df_cm$cm, predict(mixed_models[[3]], df_cm)))
# print(mse(df_cm$cm, predict(mixed_models[[9]], df_cm)))
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="YhovGpsYkaql" outputId="6ed33e39-da76-43f8-8932-bc4b61172888" language="R"
# #model 9 explains the data the best out of the rest of the models
# plot(predict(mixed_models[[9]]), df_cm$cm, xlab = 'Predicted', ylab = 'Actual', main = 'Mixed Effects Model\ncm ~ annualprecp + wintertemp + (1|Tree_Size_Category)')
# abline(a = 0, b = 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 497} id="inxYPpa8katb" outputId="0d300a41-9e94-46f7-d454-d7580d76dd52" language="R"
# plot(predict(linear_models[[15]]), df_cm$cm, xlab = 'Predicted', ylab = 'Actual', main = 'Multiple Linear Regerssion Model\ncm~yr+annualprecp+wintertemp+summerpdsi+(1|Tree Categories)')
# abline(a = 0, b = 1)
# + [markdown] id="ovKJbKDhq7jk"
# # **Conclusion**
# + [markdown] id="1pf4TMmYrATY"
# ## **Model**
# $\large Avg \ \ CM_{imt} = \underbrace{\beta_0 + \beta_1AnnualPrec_{it} + \beta_2WinterTemp_{it}}_\text{Fixed Effects} + \underbrace{TC_{0mt}}_\text{Random Effect} + \epsilon_{imt}$
#
# <br/>
# <br/>
#
# - $Avg \ \ CM_{imt} = \text{Average diameter in centimeters of group $m$ and observation $i$ at year $t$, where the range is: }[0,\infty)]$
#
# - $Avg \ \ CM_{it} = \text{cumulative precipitation within a year, where the domain is: }[\text{22 inches}, \text{140 inches})]$
#
# - $WinterTemp_{it} = \text{Temperature during January, February and March, where the domain is: }[-37 \ Celsius, 43 \ Celsius)$
#
# - $TC_{0mt} = \text{Tree classification, the random effect, where the groups are: (saplings, seedlings, poletimbers, sawtimbers]}$
#
# + [markdown] id="rwMaN53ErAZu"
# ## **Model Assumptions**
# The mixed effects model has one random effect (tree size category), with the remaining explanatory variables being fixed effects. Tree size categories were chosen as the random effect because of the varying growth rates among the groups. Because seedlings grow at a faster pace than poletimbers, they can not be grouped together because that would skew the results. The random effect is a way to deal with the tree size parameter. For the fixed effects, the following assumptions were made: the residuals are IID, large outliers are unlikely and there is no significant multicollinearity.
# + [markdown] id="wVe_I3y0rAhj"
# ## **Results**
#
# The highlighted model performed the best based on BIC and Mean Squared Error metrics. The model does a good job in explaining the average diameter of the tree groups, however a model to explain the growth of individual trees was not found.
#
# <br/>
#
# Analyzing the explanatory fixed effect variables, more rain tends to lead to a larger diameter measurement for a tree, while warmer winters tended to lead to a smaller diameter measurement. The result from the winter temperature variable seems counterintuitive, however, professors from UW-Madison noted that if winter temperatures from January to March are unseasonably warm bringing an early bloom and then April turns cold, this can kill trees (Ackerman & Martin, 2017)
#
# <br/>
#
# Futhermore, the data set is small (only 30 observations). More data needs to be collected in order to test the model to verify that it can infer the average diameter measurement of tree groups. Additional data may not be available from Duke University, so data sets from national parks or other university forests should be considered.
#
# <br/>
#
# What can be infered is that grouping trees by their category size is helpful when modeling the diameter of the trees. The data clearly show clustering and this is further quantified with the high intraclass correlations seen across the mixed effects models.
# + [markdown] id="jFOZJ0AmHRFo"
# # **References**
#
# - Data retrieved from [Duke Department of Statistical Science](https://stat.duke.edu/datasets/diameter-measurement
# )
# + [markdown] id="SxVBQ31qq0hM"
# ## **Dendrology (tree science) References**
#
# - **[Do trees need cold weather?](https://wxguys.ssec.wisc.edu/2017/10/02/trees-need-cold/) by <NAME> and <NAME>**
#
# - **[Periodic Mapped-plot Design Inventory Terminology](https://www.fs.fed.us/rm/ogden/publications/reference/periodic_terminology_final.pdf) (tree size classification) by the USDA Forest Service**
# + [markdown] id="aFBJcUHFq1D2"
# ## **R References**
#
# - **[Mixed Models with R](https://m-clark.github.io/mixed-models-with-R/random_intercepts.html) by <NAME>**
#
# - **[Building and Comparing Mixed Models in R](https://www.youtube.com/watch?v=Wtk5iZ65XHk) by Quant Psych**
#
# - **[Use = or <- for assignment?](https://blog.revolutionanalytics.com/2008/12/use-equals-or-arrow-for-assignment.html) by <NAME>**
#
# - **[Stack Overflow Post 1](https://stackoverflow.com/questions/5300595/automatically-create-formulas-for-all-possible-linear-models) on Looping through Explanatory Variables in R**
#
# - **[Stack Overflow Post 2](https://stackoverflow.com/questions/15901224/what-is-difference-between-dataframe-and-list-in-r) on R Lists and DataFrames**
| Google Colab Tutorials/Exploratory_Data_Analysis_&_Modeling_with_Python_and_R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example 4b: Fermionic single impurity model
#
# ## Example of the Fermionic HEOM solver
#
# Here we model a single fermion coupled to two electronic leads or reservoirs (e.g., this can describe a single quantum dot, a molecular transistor, etc). Note that in this implementation we primarily follow the definitions used by <NAME> in his Dissertation https://opus4.kobv.de/opus4-fau/files/10984/DissertationChristianSchinabeck.pdf and related publications.
#
#
# Notation:
# $K=L/R$ refers to left or right leads.
#
# $\sigma=\pm$ refers to input/output
#
#
# We choose a Lorentzian spectral density for the leads, with a peak at the chemical potential. The latter simplifies a little the notation required for the correlation functions, but can be relaxed if neccessary.
#
# $$J(\omega) = \frac{\Gamma W^2}{((\omega-\mu_K)^2 +W^2 )}$$
#
#
# Fermi distribution is
#
# $$f_F (x) = (\exp(x) + 1)^{-1}$$
#
# gives correlation functions
#
# $$C^{\sigma}_K(t) = \frac{1}{2\pi} \int_{-\infty}^{\infty} d\omega e^{\sigma i \omega t} \Gamma_K(\omega) f_F[\sigma\beta(\omega - \mu)]$$
#
#
# As with the Bosonic case we can treat these with Matsubara, Pade, or fitting approaches.
#
# The Pade decomposition approximates the Fermi distubition as
#
# $$f_F(x) \approx f_F^{\mathrm{approx}}(x) = \frac{1}{2} - \sum_l^{l_{max}} \frac{2k_l x}{x^2 + \epsilon_l^2}$$
#
# $k_l$ and $\epsilon_l$ are co-efficients defined in J. Chem Phys 133,10106
#
# Evaluating the integral for the correlation functions gives,
#
#
# $$C_K^{\sigma}(t) \approx \sum_{l=0}^{l_{max}} \eta_K^{\sigma_l} e^{-\gamma_{K,\sigma,l}t}$$
#
# where
#
# $$\eta_{K,0} = \frac{\Gamma_KW_K}{2} f_F^{approx}(i\beta_K W)$$
#
# $$\gamma_{K,\sigma,0} = W_K - \sigma i\mu_K$$
#
# $$\eta_{K,l\neq 0} = -i\cdot \frac{k_m}{\beta_K} \cdot \frac{\Gamma_K W_K^2}{-\frac{\epsilon^2_m}{\beta_K^2} + W_K^2}$$
#
# $$\gamma_{K,\sigma,l\neq 0}= \frac{\epsilon_m}{\beta_K} - \sigma i \mu_K$$
#
#
#
from qutip import *
# %pylab inline
# %load_ext autoreload
# %autoreload 2
# +
from bofinfast.heom import FermionicHEOMSolver
import time
from scipy.integrate import quad
# -
def deltafun(j,k):
if j==k:
return 1.
else:
return 0.
# +
from qutip.states import enr_state_dictionaries
def get_aux_matrices(full, level, N_baths, Nk, N_cut, shape, dims):
"""
Extracts the auxiliary matrices at a particular level
from the full hierarchy ADOs.
Parameters
----------
full: ndarray
A 2D array of the time evolution of the ADOs.
level: int
The level of the hierarchy to get the ADOs.
N_cut: int
The hierarchy cutoff.
k: int
The total number of exponentials used in each bath (assumed equal).
N_baths: int
The number of baths.
shape : int
the size of the ''system'' hilbert space
dims : list
the dimensions of the system hilbert space
"""
#Note: Max N_cut is Nk*N_baths
nstates, state2idx, idx2state = enr_state_dictionaries([2]*(Nk*N_baths) ,N_cut)#_heom_state_dictionaries([Nc + 1]*(Nk), Nc)
aux_indices = []
aux_heom_indices = []
for stateid in state2idx:
if np.sum(stateid) == level:
aux_indices.append(state2idx[stateid])
aux_heom_indices.append(stateid)
full = np.array(full)
aux = []
for i in aux_indices:
qlist = [Qobj(full[k, i, :].reshape(shape, shape).T,dims=dims) for k in range(len(full))]
aux.append(qlist)
return aux, aux_heom_indices, idx2state
# +
#Define parameters and plot lead spectra
Gamma = 0.01 #coupling strength
W=1. #cut-off
T = 0.025851991 #temperature
beta = 1./T
theta = 2. #Bias
mu_l = theta/2.
mu_r = -theta/2.
w_list = np.linspace(-2,2,100)
def Gamma_L_w(w):
return Gamma*W**2/((w-mu_l)**2 + W**2)
def Gamma_R_w(w):
return Gamma*W**2/((w-mu_r)**2 + W**2)
def f(x):
kB=1.
return 1/(exp(x)+1.)
def f2(x):
return 0.5
# +
fig, ax1 = plt.subplots(figsize=(12, 7))
gam_list_in = [Gamma_L_w(w)*f(beta*(w-mu_l)) for w in w_list]
ax1.plot(w_list,gam_list_in, "b--", linewidth=3, label= r"S_L(w) input (absorption)")
ax1.set_xlabel("w")
ax1.set_ylabel(r"$S(\omega)$")
ax1.legend()
gam_list_out = [Gamma_L_w(w)*f(-beta*(w-mu_l)) for w in w_list]
spec = [Gamma_L_w(w) for w in w_list]
ax1.plot(w_list,gam_list_out, "r--", linewidth=3, label= r"S_L(w) output (emission)")
gam_list_in = [Gamma_R_w(w)*f(beta*(w-mu_r)) for w in w_list]
ax1.plot(w_list,gam_list_in, "b", linewidth=3, label= r"S_R(w) input (absorption)")
gam_list_out = [Gamma_R_w(w)*f(-beta*(w-mu_r)) for w in w_list]
spec = [Gamma_R_w(w) for w in w_list]
ax1.plot(w_list,gam_list_out, "r", linewidth=3, label= r"S_R(w) output (emission)")
ax1.set_xlabel("w")
ax1.set_ylabel(r"$n$")
ax1.legend()
# +
#Pade decompositon: construct correlation parameters
tlist = np.linspace(0,10,200)
#Pade cut-off
lmax =10
w_list = np.linspace(-2,2,100)
def Gamma_L_w(w):
return Gamma*W**2/((w-mu_l)**2 + W**2)
def Gamma_w(w, mu):
return Gamma*W**2/((w-mu)**2 + W**2)
def f(x):
kB=1.
return 1/(exp(x)+1.)
Alpha =np.zeros((2*lmax,2*lmax))
for j in range(2*lmax):
for k in range(2*lmax):
Alpha[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)-1)*(2*(k+1)-1))
eigvalsA=eigvalsh(Alpha)
eps = []
for val in eigvalsA[0:lmax]:
#print(-2/val)
eps.append(-2/val)
AlphaP =np.zeros((2*lmax-1,2*lmax-1))
for j in range(2*lmax-1):
for k in range(2*lmax-1):
AlphaP[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)+1)*(2*(k+1)+1))
#AlphaP[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+2)-1)*(2*(k+2)-1))
eigvalsAP=eigvalsh(AlphaP)
chi = []
for val in eigvalsAP[0:lmax-1]:
#print(-2/val)
chi.append(-2/val)
eta_list = [0.5*lmax*(2*(lmax + 1) - 1)*(
np.prod([chi[k]**2 - eps[j]**2 for k in range(lmax - 1)])/
np.prod([eps[k]**2 - eps[j]**2 +deltafun(j,k) for k in range(lmax)]))
for j in range(lmax)]
kappa = [0]+eta_list
print(kappa)
epsilon = [0]+eps
print(epsilon)
def f_approx(x):
f = 0.5
for l in range(1,lmax+1):
f= f - 2*kappa[l]*x/(x**2+epsilon[l]**2)
return f
def f(x):
kB=1.
return 1/(exp(x)+1.)
def C(tlist,sigma,mu):
eta_list = []
gamma_list =[]
eta_0 = 0.5*Gamma*W*f_approx(1.0j*beta*W)
gamma_0 = W - sigma*1.0j*mu
eta_list.append(eta_0)
gamma_list.append(gamma_0)
if lmax>0:
for l in range(1,lmax+1):
eta_list.append(-1.0j*(kappa[l]/beta)*Gamma*W**2/(-(epsilon[l]**2/beta**2)+W**2))
gamma_list.append(epsilon[l]/beta - sigma*1.0j*mu)
c_tot = []
for t in tlist:
c_tot.append(sum([eta_list[l]*exp(-gamma_list[l]*t) for l in range(lmax+1)]))
return c_tot, eta_list, gamma_list
def c_t_L_num(t,sigma,mu):
integrand = lambda w: (1/(2*pi))*exp(sigma*1.0j*w*t)*Gamma_w(w,mu)*f(sigma*beta*(w-mu))
def real_func(x):
return real(integrand(x))
def imag_func(x):
return imag(integrand(x))
#return quad(integrand,-np.inf,np.inf)[0]
# This bounds must be increased if W is increased
# But this integration is quite unstable for large frequencies.
a= -50
b= 50
real_integral = quad(real_func, a, b)
imag_integral = quad(imag_func, a, b)
return real_integral[0] + 1.0j * imag_integral[0]
cppL,etapL,gampL = C(tlist,1.0,mu_l)
cpmL,etamL,gammL = C(tlist,-1.0,mu_l)
cppR,etapR,gampR = C(tlist,1.0,mu_r)
cpmR,etamR,gammR = C(tlist,-1.0,mu_r)
c_num =[c_t_L_num(t,-1.0,mu_r) for t in tlist]
fig, ax1 = plt.subplots(figsize=(12, 7))
ax1.plot(tlist,real(c_num), color="b", linewidth=3, label= r"C num")
ax1.plot(tlist,real(cpmR), "r--", linewidth=3, label= r"C pade")
pos = 1
ax1.set_xlabel("t")
ax1.set_ylabel(r"$C$")
ax1.legend()
fig, ax1 = plt.subplots(figsize=(12, 7))
ax1.plot(tlist,imag(c_num), color="b", linewidth=3, label= r"C num")
ax1.plot(tlist,imag(cpmR), "r--", linewidth=3, label= r"C pade")
pos = 0
# +
#heom simulation with above params (Pade)
options = Options(nsteps=15000, store_states=True, rtol=1e-14, atol=1e-14)
#Single fermion.
d1 = destroy(2)
#Site energy
e1 = 1.
H0 = e1*d1.dag()*d1
#There are two leads, but we seperate the interaction into two terms, labelled with \sigma=\pm
#such that there are 4 interaction operators (See paper)
Qops = [d1.dag(),d1,d1.dag(),d1]
Kk=lmax+1
Ncc = 2 #For a single impurity we converge with Ncc = 2
#Note here that the functionality differs from the bosonic case. Here we send lists of lists, were each sub-list
#refers to one of the two coupling terms for each bath (the notation here refers to eta|sigma|L/R)
start = time.time()
eta_list = [etapR,etamR,etapL,etamL]
gamma_list = [gampR,gammR,gampL,gammL]
Qops = [d1.dag(),d1,d1.dag(),d1]
resultHEOM2 = FermionicHEOMSolver(H0, Qops, eta_list, gamma_list, Ncc,options=options)
end = time.time()
print("new c code", end - start)
# +
rho_0 = basis(2,0)*basis(2,0).dag()
start = time.time()
rhossHP2,fullssP2=resultHEOM2.steady_state()
end = time.time()
print(end - start)
# -
rho_0 = basis(2,0)*basis(2,0).dag()
tlist = np.linspace(0,100,1000)
out1P2=resultHEOM2.run(rho_0,tlist)
# +
# Plot the results
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(8,8))
axes.plot(tlist, expect(out1P2.states,rho_0), 'r--', linewidth=2, label="P11 ")
axes.set_xlabel(r't', fontsize=28)
axes.legend(loc=0, fontsize=12)
# +
#We can perform the same calculation using Matsubara decomposition
tlist = np.linspace(0,10,100)
lmax = 10
kappa = [0.]
kappa.extend([1. for l in range(1,lmax+1)])
epsilon = [0]
epsilon.extend([(2*l-1)*pi for l in range(1,lmax+1)])
def f_approx(x):
f = 0.5
for l in range(1,lmax+1):
f= f - 2*kappa[l]*x/(x**2+epsilon[l]**2)
return f
def C(tlist,sigma,mu):
eta_list = []
gamma_list =[]
#l = 0
eta_0 = 0.5*Gamma*W*f(1.0j*beta*W)
gamma_0 = W - sigma*1.0j*mu
eta_list.append(eta_0)
gamma_list.append(gamma_0)
if lmax>0:
for l in range(1,lmax+1):
eta_list.append(-1.0j*(kappa[l]/beta)*Gamma*W**2/(-(epsilon[l]**2/beta**2)+W**2))
gamma_list.append(epsilon[l]/beta - sigma*1.0j*mu)
c_tot = []
for t in tlist:
c_tot.append(sum([eta_list[l]*exp(-gamma_list[l]*t) for l in range(lmax+1)]))
return c_tot, eta_list, gamma_list
def c_t_L_num(t,sigma,mu):
integrand = lambda w: (1/(2*pi))*exp(sigma*1.0j*w*t)*Gamma_w(w,mu)*f(sigma*beta*(w-mu))
def real_func(x):
return real(integrand(x))
def imag_func(x):
return imag(integrand(x))
a = -50
b = 50
real_integral = quad(real_func, a, b)
imag_integral = quad(imag_func, a, b)
return real_integral[0] + 1.0j * imag_integral[0]
cppL,etapL,gampL = C(tlist,1.0,mu_l)
cpmL,etamL,gammL = C(tlist,-1.0,mu_l)
cppR,etapR,gampR = C(tlist,1.0,mu_r)
cpmR,etamR,gammR = C(tlist,-1.0,mu_r)
c_num =[c_t_L_num(t,1.0,mu_l) for t in tlist]
fig, ax1 = plt.subplots(figsize=(12, 7))
ax1.plot(tlist,real(cppL), color="b", linewidth=3, label= r"C Matsubara")
ax1.plot(tlist,real(c_num), "r--", linewidth=3, label= r"C num")
ax1.set_xlabel("t")
ax1.set_ylabel(r"$Re[C]$")
ax1.legend()
fig, ax1 = plt.subplots(figsize=(12, 7))
#print(gam_list)
ax1.plot(tlist,imag(cppL), color="b", linewidth=3, label= r"C Matsubara")
ax1.plot(tlist,imag(c_num), "r--", linewidth=3, label= r"C num")
ax1.set_xlabel("t")
ax1.set_ylabel(r"$Im[C]$")
# +
#heom simu on above params (Matsubara)
d1 = destroy(2)
e1 = 1.
H0 = e1*d1.dag()*d1
Qops = [d1.dag(),d1,d1.dag(),d1]
rho_0 = basis(2,0)*basis(2,0).dag()
Kk=lmax+1
Ncc = 2
tlist = np.linspace(0,100,1000)
eta_list = [etapR,etamR,etapL,etamL]
gamma_list = [gampR,gammR,gampL,gammL]
start = time.time()
resultHEOM2 = FermionicHEOMSolver(H0, Qops, eta_list, gamma_list, Ncc)
end = time.time()
print("C code", end - start)
# +
out1M2 = resultHEOM2.run(rho_0,tlist)
# +
start = time.time()
rhossHM2,fullssM2 = resultHEOM2.steady_state()
end = time.time()
print(end - start)
# +
# Plot the results
fig, axes = plt.subplots(1, 1, sharex=True, figsize=(8,8))
axes.plot(tlist, expect(out1P2.states,rho_0), 'r--', linewidth=2, label="P11 Pade ")
axes.plot(tlist, expect(out1M2.states,rho_0), 'b--', linewidth=2, label="P12 Mats C")
axes.set_ylabel(r"$\rho_{11}$")
axes.set_xlabel(r't', fontsize=28)
axes.legend(loc=0, fontsize=12)
# +
#One advantage of this simple model is the current is analytically solvable, so we can check convergence of the result
def CurrFunc():
def lamshift(w,mu):
return (w-mu)*Gamma_w(w,mu)/(2*W)
integrand = lambda w: ((2/(pi))*Gamma_w(w,mu_l)*Gamma_w(w,mu_r)*(f(beta*(w-mu_l))-f(beta*(w-mu_r))) /
((Gamma_w(w,mu_l)+Gamma_w(w,mu_r))**2 +4*(w-e1 - lamshift(w,mu_l)-lamshift(w,mu_r))**2))
def real_func(x):
return real(integrand(x))
def imag_func(x):
return imag(integrand(x))
#in principle the bounds should be checked if parameters are changed
a= -2
b=2
real_integral = quad(real_func, a, b)
imag_integral = quad(imag_func, a, b)
return real_integral[0] + 1.0j * imag_integral[0]
curr_ana = CurrFunc()
print(curr_ana)
# +
#we can extract the current from the auxiliary ADOs calculated in the steady state
aux_1_list_list=[]
aux1_indices_list=[]
aux_2_list_list=[]
aux2_indices_list=[]
K = Kk
shape = H0.shape[0]
dims = H0.dims
aux_1_list, aux1_indices, idx2state = get_aux_matrices([fullssP2], 1, 4, K, Ncc, shape, dims)
aux_2_list, aux2_indices, idx2state = get_aux_matrices([fullssP2], 2, 4, K, Ncc, shape, dims)
d1 = destroy(2) #Kk to 2*Kk
currP = -1.0j * (((sum([(d1*aux_1_list[gg][0]).tr() for gg in range(Kk,2*Kk)]))) - ((sum([(d1.dag()*aux_1_list[gg][0]).tr() for gg in range(Kk)]))))
# +
aux_1_list_list=[]
aux1_indices_list=[]
aux_2_list_list=[]
aux2_indices_list=[]
K = Kk
shape = H0.shape[0]
dims = H0.dims
aux_1_list, aux1_indices, idx2state = get_aux_matrices([fullssM2], 1, 4, K, Ncc, shape, dims)
aux_2_list, aux2_indices, idx2state = get_aux_matrices([fullssM2], 2, 4, K, Ncc, shape, dims)
d1 = destroy(2) #Kk to 2*Kk
currM = -1.0j * (((sum([(d1*aux_1_list[gg][0]).tr() for gg in range(Kk,2*Kk)]))) - ((sum([(d1.dag()*aux_1_list[gg][0]).tr() for gg in range(Kk)]))))
# +
print("Pade current", -currP)
print("Matsubara current", -currM)
print("Analytical curernt", curr_ana)
# +
start=time.time()
currPlist = []
curranalist = []
theta_list = linspace(-4,4,100)
for theta in theta_list:
mu_l = theta/2.
mu_r = -theta/2.
#Pade cut-off
lmax = 10
Alpha =np.zeros((2*lmax,2*lmax))
for j in range(2*lmax):
for k in range(2*lmax):
Alpha[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)-1)*(2*(k+1)-1))
eigvalsA=eigvalsh(Alpha)
eps = []
for val in eigvalsA[0:lmax]:
#print(-2/val)
eps.append(-2/val)
AlphaP =np.zeros((2*lmax-1,2*lmax-1))
for j in range(2*lmax-1):
for k in range(2*lmax-1):
AlphaP[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)+1)*(2*(k+1)+1))
#AlphaP[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+2)-1)*(2*(k+2)-1))
eigvalsAP=eigvalsh(AlphaP)
chi = []
for val in eigvalsAP[0:lmax-1]:
#print(-2/val)
chi.append(-2/val)
eta_list = [0.5*lmax*(2*(lmax + 1) - 1)*(
np.prod([chi[k]**2 - eps[j]**2 for k in range(lmax - 1)])/
np.prod([eps[k]**2 - eps[j]**2 +deltafun(j,k) for k in range(lmax)]))
for j in range(lmax)]
kappa = [0]+eta_list
epsilon = [0]+eps
def f_approx(x):
f = 0.5
for l in range(1,lmax+1):
f= f - 2*kappa[l]*x/(x**2+epsilon[l]**2)
return f
def f(x):
kB=1.
return 1/(exp(x)+1.)
def C(sigma,mu):
eta_list = []
gamma_list =[]
eta_0 = 0.5*Gamma*W*f_approx(1.0j*beta*W)
gamma_0 = W - sigma*1.0j*mu
eta_list.append(eta_0)
gamma_list.append(gamma_0)
if lmax>0:
for l in range(1,lmax+1):
eta_list.append(-1.0j*(kappa[l]/beta)*Gamma*W**2/(-(epsilon[l]**2/beta**2)+W**2))
gamma_list.append(epsilon[l]/beta - sigma*1.0j*mu)
return eta_list, gamma_list
etapL,gampL = C(1.0,mu_l)
etamL,gammL = C(-1.0,mu_l)
etapR,gampR = C(1.0,mu_r)
etamR,gammR = C(-1.0,mu_r)
#heom simu on above params (Matsubara)
d1 = destroy(2)
e1 = .3
H0 = e1*d1.dag()*d1
Qops = [d1.dag(),d1,d1.dag(),d1]
rho_0 = basis(2,0)*basis(2,0).dag()
Kk=lmax+1
Ncc = 2
tlist = np.linspace(0,100,1000)
eta_list = [etapR,etamR,etapL,etamL]
gamma_list = [gampR,gammR,gampL,gammL]
resultHEOM = FermionicHEOMSolver(H0, Qops, eta_list, gamma_list, Ncc)
rho_0 = basis(2,0)*basis(2,0).dag()
rhossHP,fullssP=resultHEOM.steady_state()
#we can extract the current from the auxiliary ADOs calculated in the steady state
aux_1_list_list=[]
aux1_indices_list=[]
aux_2_list_list=[]
aux2_indices_list=[]
K = Kk
shape = H0.shape[0]
dims = H0.dims
aux_1_list, aux1_indices, idx2state = get_aux_matrices([fullssP], 1, 4, K, Ncc, shape, dims)
aux_2_list, aux2_indices, idx2state = get_aux_matrices([fullssP], 2, 4, K, Ncc, shape, dims)
d1 = destroy(2) #Kk to 2*Kk
currP = -1.0j * (((sum([(d1*aux_1_list[gg][0]).tr() for gg in range(Kk,2*Kk)]))) - ((sum([(d1.dag()*aux_1_list[gg][0]).tr() for gg in range(Kk)]))))
curr_ana = CurrFunc()
currPlist.append(currP)
curranalist.append(curr_ana)
end=time.time()
print("run time", end-start)
# -
matplotlib.rcParams['figure.figsize'] = (7, 5)
matplotlib.rcParams['axes.titlesize'] = 25
matplotlib.rcParams['axes.labelsize'] = 30
matplotlib.rcParams['xtick.labelsize'] = 28
matplotlib.rcParams['ytick.labelsize'] = 28
matplotlib.rcParams['legend.fontsize'] = 28
matplotlib.rcParams['axes.grid'] = False
matplotlib.rcParams['savefig.bbox'] = 'tight'
matplotlib.rcParams['lines.markersize'] = 5
matplotlib.rcParams['font.family'] = 'STIXgeneral'
matplotlib.rcParams['mathtext.fontset'] = 'stix'
matplotlib.rcParams["font.serif"] = "STIX"
matplotlib.rcParams['text.usetex'] = False
# +
fig, ax1 = plt.subplots(figsize=(12,7))
ax1.plot(theta_list,2.434e-4*1e6*array(curranalist), color="black", linewidth=3, label= r"Analytical")
ax1.plot(theta_list,-2.434e-4*1e6*array(currPlist), 'r--', linewidth=3, label= r"HEOM $l_{\mathrm{max}}=10$, $n_{\mathrm{max}}=2$")
ax1.locator_params(axis='y', nbins=4)
ax1.locator_params(axis='x', nbins=4)
axes.set_xticks([-2.5,0.,2.5])
axes.set_xticklabels([-2.5,0,2.5])
ax1.set_xlabel(r"Bias voltage $\Delta \mu$ ($V$)",fontsize=28)
ax1.set_ylabel(r"Current ($\mu A$)",fontsize=28)
ax1.legend(fontsize=25)
fig.savefig("figImpurity.pdf")
# +
#We can also generate the above data using the MKL solver in the steady-state method
#This tends to be quicker on very large examples. Here it converges to the correct result, but can
#sometimes fail to converge, or give incorrect results.
currPlist = []
curranalist = []
theta_list = linspace(-4,4,100)
start = time.time()
for theta in theta_list:
mu_l = theta/2.
mu_r = -theta/2.
#Pade cut-off
lmax = 10
Alpha =np.zeros((2*lmax,2*lmax))
for j in range(2*lmax):
for k in range(2*lmax):
Alpha[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)-1)*(2*(k+1)-1))
eigvalsA=eigvalsh(Alpha)
eps = []
for val in eigvalsA[0:lmax]:
#print(-2/val)
eps.append(-2/val)
AlphaP =np.zeros((2*lmax-1,2*lmax-1))
for j in range(2*lmax-1):
for k in range(2*lmax-1):
AlphaP[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+1)+1)*(2*(k+1)+1))
#AlphaP[j][k] = (deltafun(j,k+1)+deltafun(j,k-1))/sqrt((2*(j+2)-1)*(2*(k+2)-1))
eigvalsAP=eigvalsh(AlphaP)
chi = []
for val in eigvalsAP[0:lmax-1]:
#print(-2/val)
chi.append(-2/val)
eta_list = [0.5*lmax*(2*(lmax + 1) - 1)*(
np.prod([chi[k]**2 - eps[j]**2 for k in range(lmax - 1)])/
np.prod([eps[k]**2 - eps[j]**2 +deltafun(j,k) for k in range(lmax)]))
for j in range(lmax)]
kappa = [0]+eta_list
epsilon = [0]+eps
def f_approx(x):
f = 0.5
for l in range(1,lmax+1):
f= f - 2*kappa[l]*x/(x**2+epsilon[l]**2)
return f
def f(x):
kB=1.
return 1/(exp(x)+1.)
def C(sigma,mu):
eta_list = []
gamma_list =[]
eta_0 = 0.5*Gamma*W*f_approx(1.0j*beta*W)
gamma_0 = W - sigma*1.0j*mu
eta_list.append(eta_0)
gamma_list.append(gamma_0)
if lmax>0:
for l in range(1,lmax+1):
eta_list.append(-1.0j*(kappa[l]/beta)*Gamma*W**2/(-(epsilon[l]**2/beta**2)+W**2))
gamma_list.append(epsilon[l]/beta - sigma*1.0j*mu)
return eta_list, gamma_list
etapL,gampL = C(1.0,mu_l)
etamL,gammL = C(-1.0,mu_l)
etapR,gampR = C(1.0,mu_r)
etamR,gammR = C(-1.0,mu_r)
#heom simu on above params (Matsubara)
d1 = destroy(2)
e1 = .3
H0 = e1*d1.dag()*d1
Qops = [d1.dag(),d1,d1.dag(),d1]
rho_0 = basis(2,0)*basis(2,0).dag()
Kk=lmax+1
Ncc = 2
tlist = np.linspace(0,100,1000)
eta_list = [etapR,etamR,etapL,etamL]
gamma_list = [gampR,gammR,gampL,gammL]
resultHEOM = FermionicHEOMSolver(H0, Qops, eta_list, gamma_list, Ncc)
rho_0 = basis(2,0)*basis(2,0).dag()
rhossHP,fullssP=resultHEOM.steady_state(use_mkl=True)
#we can extract the current from the auxiliary ADOs calculated in the steady state
aux_1_list_list=[]
aux1_indices_list=[]
aux_2_list_list=[]
aux2_indices_list=[]
K = Kk
shape = H0.shape[0]
dims = H0.dims
aux_1_list, aux1_indices, idx2state = get_aux_matrices([fullssP], 1, 4, K, Ncc, shape, dims)
aux_2_list, aux2_indices, idx2state = get_aux_matrices([fullssP], 2, 4, K, Ncc, shape, dims)
d1 = destroy(2) #Kk to 2*Kk
currP = -1.0j * (((sum([(d1*aux_1_list[gg][0]).tr() for gg in range(Kk,2*Kk)]))) - ((sum([(d1.dag()*aux_1_list[gg][0]).tr() for gg in range(Kk)]))))
curr_ana = CurrFunc()
currPlist.append(currP)
curranalist.append(curr_ana)
end=time.time()
print("run time", end-start)
# +
fig, ax1 = plt.subplots(figsize=(12,7))
ax1.plot(theta_list,2.434e-4*1e6*array(curranalist), color="black", linewidth=3, label= r"Analytical")
ax1.plot(theta_list,-2.434e-4*1e6*array(currPlist), 'r--', linewidth=3, label= r"HEOM $l_{\mathrm{max}}=10$, $n_{\mathrm{max}}=2$")
ax1.locator_params(axis='y', nbins=4)
ax1.locator_params(axis='x', nbins=4)
axes.set_xticks([-2.5,0.,2.5])
axes.set_xticklabels([-2.5,0,2.5])
ax1.set_xlabel(r"bias voltage $\Delta \mu$ ($V$)",fontsize=28)
ax1.set_ylabel(r"Current ($\mu A$)",fontsize=28)
ax1.legend(fontsize=25)
# -
| examples/example-4b-fermions-single-impurity-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Library Fine
# +
# #!/bin/python3
import sys
def libraryFine(d1, m1, y1, d2, m2, y2):
if y1 < y2:
return 0
elif y1 > y2:
return (y1 - y2) * 10000
elif m1 < m2:
return 0
elif m1 > m2:
return (m1 - m2) * 500
elif d1 > d2:
return (d1 - d2) * 15
else:
return 0
if __name__ == "__main__":
d1, m1, y1 = input().strip().split(' ')
d1, m1, y1 = [int(d1), int(m1), int(y1)]
d2, m2, y2 = input().strip().split(' ')
d2, m2, y2 = [int(d2), int(m2), int(y2)]
result = libraryFine(d1, m1, y1, d2, m2, y2)
print(result)
| Algorithms/Library Fine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# I want to create a data-driven model to better predict the winners of the current year.
# The model i want to create uses a region-based Elo rating. <br/>
#
# We use a genetic algorithm to learn an elo model where the variable are:
# - one K constant (positive) for the maximum points gainable/losable per win/lose
# - a number of C(Region) costant based on the number of regions. Used in the calculation of the Expected Score.<br/>
#
# Precisely, the Expected Score for A is equal to E(A) = Q(A)/(Q(A)+Q(B)). <br/>
# The classic Q(A) function is Q(A) = 10 ^ (R(A)/400), where R(A) represent the Elo rating of the team A. <br/>
# In our case, instead, it is Q(A;Region) = 10^(R(A)/C(Region)).<br/>
# The Rating is upgraded using the following formula:<br/>
# R(A)' = R(A) + K ( S(A) - E(A) ), where the newly introduced S(A) is the score of A. <br/>
# This mode is used to calculate the most probable results for the Worlds Main Event. Group and Finals <br/>
#
# I will add another variable called "Elo Degradation" D . This will automatically reduce (or improve, for experimentation) of the elo before the process of upgrading it.<br/>
# R(A)' = (R(A)) * D + K ( S(A) - E(A) )<br/>
# Starting Elo is 0, this makes for an easier implementation of both Elo scaling and conversion methods.
import mwclient
site = mwclient.Site('lol.gamepedia.com', path='/')
# I obtain all of the data I need to create the various worlds dataset
from tqdm.notebook import tqdm
leagues=['GPL','IWCQ','WCS','LCS','LEC','LCK','LPL','CBLOL','LCL','LJL','LLA','OPL','PCS','VCS','TCL','LMS','LST','NA LCS','EU LCS','LLN','CLS']
results = []
for league in tqdm(leagues):
off=0
while(True):
response = site.api('cargoquery',
offset=str(off),
limit="500",
tables = "Tournaments=T, Leagues=L, MatchSchedule=M",
fields="L.League_Short=League, T.Name=Tournament, M.DateTime_UTC=Date, M.Team1Final=Team1, M.Team2Final=Team2, M.Team1Score, M.Team2Score",
where = 'L.League_Short = "'+league+'"',
join_on = "T.League = L.League, T.OverviewPage=M.OverviewPage")
results += response["cargoquery"]
off=off+500
if(len(response["cargoquery"])<500): break
off=0
while(True):
response = site.api('cargoquery',
offset=str(off),
limit="500",
tables = "Tournaments=T, Leagues=L, MatchSchedule=M",
fields="T.League, T.Name=Tournament, M.DateTime_UTC=Date, M.Team1Final=Team1, M.Team2Final=Team2, M.Team1Score, M.Team2Score",
where = 'T.League = "League of Legends SEA Tour"',
join_on = "T.League = L.League, T.OverviewPage=M.OverviewPage")
results += response["cargoquery"]
off=off+500
if(len(response["cargoquery"])<500): break
import pandas as pd
MatchData = pd.DataFrame ( [ a['title'] for a in results ] )
MatchData = MatchData.replace("",float("NaN")).dropna(axis='index', how='any')
# Make "date" easier to process
MatchData.sort_values(by="Date",inplace=True)
MatchData = MatchData.astype({'Team1Score':'int32','Team2Score':'int32'})
MatchData["Date"] = pd.to_datetime(MatchData["Date"], format='%Y-%m-%d %H:%M:%S')
MatchData = MatchData.drop("Date__precision", axis=1)
MatchData.League[MatchData.League=='League of Legends SEA Tour']="LST"
# Now we need to take the data we need for the learning process. Precisely we will use the matches from the end of the previous World Championship to the end of the analyzed year World Play-Ins for the learning process.
LearningDates = {
2020:{'PreviousWCSEnd':'2019-11-11 00:00:00','PlayInEnd':'2020-10-01 00:00:00'},
2019:{'PreviousWCSEnd':'2018-11-04 00:00:00','PlayInEnd':'2019-10-09 00:00:00'},
2018:{'PreviousWCSEnd':'2017-11-05 00:00:00','PlayInEnd':'2018-10-08 00:00:00'},
2017:{'PreviousWCSEnd':'2016-10-30 00:00:00','PlayInEnd':'2017-09-30 00:00:00'}
}
PerYearMatchData=dict()
for date in LearningDates:
PerYearMatchData[date] = MatchData[
(MatchData.Date>LearningDates[date]['PreviousWCSEnd'])&
(MatchData.Date<LearningDates[date]['PlayInEnd'])]
PerYearMatchData[2017]
# The leagues appearing in the dataset are
MatchData["League"].unique()
PerYearMatchData[2020]["League"].unique()
# We will use the first League a team appears in to decide the "League" it is part of.
# For the compatibility from 2017 to 2020 World Championships we need to alias many of those leagues like this:<br/>
# "LCS" <- "NA LCLS"<br/>
# "LEC" <- "EU LCS"<br/>
# "LCK"<br/>
# "LPL"<br/>
# "CBLOL"<br/>
# "LCL"<br/>
# "LJL"<br/>
# "LLA" <- "LLN" and "CLS"<br/>
# "OPL"<br/>
# "PCS" <- "VCS"(was with GPL during 2017), "LMS", "LST" and "GPL"
# "TCL"
#
# +
alias = {
'GPL': 'PCS',
'NA LCS': 'LCS',
'EU LCS': 'LEC',
'LPL': 'LPL',
'TCL': 'PCS',
'LJL': 'LJL',
'CBLOL': 'CBLOL',
'LMS': 'PCS',
'OPL': 'OPL',
'CLS': 'LLA',
'LCK': 'LCK',
'LCL': 'LCL',
'LLN': 'LLA',
'VCS': 'PCS',
'LEC': 'LEC',
'LLA': 'LLA',
'LCS': 'LCS',
'PCS': 'PCS',
'LST': 'PCS'
}
regions = list(set([alias[name] for name in alias]))
# -
regions_dict = {regions[i]:i for i in range(len(regions))}
regions_indexes = {region:regions_dict[alias[region]] for region in alias}
# We should create the Worlds dataset to be used at the end of the calculations to verify our Region-based elo models. Before i didn't take the "Group" column, which can be used to infer the results of the various groups.
WorldsData=dict()
GroupsData=dict()
for year in [2017,2018,2019,2020]:
response = site.api('cargoquery',
limit="max",
tables = "MatchSchedule=M",
fields="M.Team1Final=Team1, M.Team2Final=Team2, M.Team1Score, M.Team2Score, M.Tab",
where = 'M.OverviewPage = "'+str(year)+' Season World Championship/Main Event"')
WorldsData[year] = pd.DataFrame ( [ a['title'] for a in response['cargoquery'] ] )
response = site.api('cargoquery',
limit="max",
tables = "TournamentGroups=T",
fields="T.Team, T.GroupName",
where = 'T.OverviewPage = "'+str(year)+' Season World Championship/Main Event"')
GroupsData[year] = pd.DataFrame ( [ a['title'] for a in response['cargoquery'] ] )
GroupsData[2019]
def WhoWon(row):
if row["Team1Score"]>row["Team2Score"]: return row["Team1"]
else: return row["Team2"]
FinalPlayOff = dict() # Will contain the matches for the final play-off
for year in [2017,2018,2019,2020]:
GroupsData[year]["Wins"] = 0
FinalPlayOff[year] = pd.DataFrame([], columns=WorldsData[year].columns)
for index,row in WorldsData[year].iterrows():
if (row.Tab not in ['Quarterfinals', 'Semifinals', 'Finals']):
winner_row = GroupsData[year].Team == WhoWon(row)
GroupsData[year].Wins[winner_row] += 1
else:
FinalPlayOff[year] = FinalPlayOff[year].append(row)
FinalPlayOff[year]["Winner"] = FinalPlayOff[year].apply(WhoWon,axis=1)
import numpy as np
class EloRating:
def __init__(self,K,C,D,starting_elo=0):
self.K = max(K,1) # Maximum Elo variation
self.C = C # Regional Elo correction constant
self.D = D # Elo degradation constant
self.start = starting_elo
def RealElo(self, Rating, Region):
return Rating/self.C[Region]*400
def E(self,RatingA, RatingB, RegionA, RegionB):
return 1/(1+10**(RatingB/self.C[RegionB]-RatingA/self.C[RegionA]))
def updateRating(self, RatingA, RatingB, RegionA, RegionB, ScoreA):
E_A = self.E(RatingA, RatingB, RegionA, RegionB)
return self.D * (RatingA - self.start) + self.K * (ScoreA - E_A)
# +
import copy
class FloatArrayGene:
def __init__(self, distributions):
self.dists = distributions
self.genes = np.array([dist['fun'](*dist['args']) for dist in distributions])
class Mutation:
def __init__(self):
pass
def mutate(self, gene, **kwargs):
return gene
class MirrorMutation(Mutation):
def __init__(self, mid): #mid is the middle point
self.midPoint = mid
def mutate(self, gene, mp=0.5, di=2): #mp is mutation probability, di a distribution index
select = np.random.random(len(gene.genes))<mp
new_gene = copy.deepcopy(gene)
new_gene.genes[select] = self.midPoint[select] - di*np.random.random()*( gene.genes[select] - self.midPoint[select])
return new_gene
class Crossover:
def __init__(self):
pass
def mutate(self,geneA, geneB, **kwargs):
return geneA
class GeneFlipCrossover(Crossover):
def __init__(self):
pass
def cross(self, geneA, geneB, cxp=0.5):
select = np.random.random(len(geneA.genes))<cxp
new_gene_A = copy.deepcopy(geneA)
new_gene_A.genes[select] = geneB.genes[select]
return new_gene_A
# -
import numpy as np
# Known K are 26,30(Chess), 258(NFL)
elo_gene_distr = [{'fun':np.random.uniform, 'args': [1,500]}]
# Known C is 400. Let's move from it, but stay around it
elo_gene_distr += [{'fun':np.random.normal, 'args': [400,80]}]*len(regions)
# Known D is 1. Let's move from it, but stay around it
elo_gene_distr += [{'fun':np.random.normal, 'args': [1,0.1]}]
# I'm creating the function to evalueate an Elo Model now
# +
from collections import defaultdict
def Score(row):
return row["Team1Score"]/(row["Team1Score"]+row["Team2Score"])
def WhoWonElo(row):
if row["Team1Elo"]>row["Team2Elo"]: return row["Team1"]
else: return row["Team2"]
class EvaluateElo:
def __init__ (self, matchData, groupData, finalPlayOff, region_indexes):
self.matches = matchData.sort_values(by="Date")
self.matches["NormScore"] = self.matches.apply(Score,axis=1)
self.groups = groupData
self.playoff = finalPlayOff
self.region_indexes = region_indexes
def evaluate(self, EloModel):
self.Rating = dict()
self.League = dict()
#Calculate ELO ratings
for index, row in self.matches.iterrows():
if(row.Team1 not in self.League):
self.League[row.Team1] = row.League
self.Rating[row.Team1] = 0
if(row.Team2 not in self.League):
self.League[row.Team2] = row.League
self.Rating[row.Team2] = 0
self.Rating[row.Team1] = EloModel.updateRating(self.Rating[row.Team1],
self.Rating[row.Team2],
self.region_indexes[self.League[row.Team1]],
self.region_indexes[self.League[row.Team2]],
row["NormScore"])
self.Rating[row.Team2] = EloModel.updateRating(self.Rating[row.Team2],
self.Rating[row.Team1],
self.region_indexes[self.League[row.Team2]],
self.region_indexes[self.League[row.Team1]],
1-row["NormScore"])
score = 0
#Scoring - Group Stage
Groups = self.groups["GroupName"].unique()
RealElo = lambda x: EloModel.RealElo(self.Rating[x],self.region_indexes[self.League[x]])
self.groups["Elo"]=self.groups["Team"].apply(RealElo)
for group in Groups:
analyzedGroup = self.groups[self.groups.GroupName==group].sort_values(by="Wins", ascending=False)
real = analyzedGroup.Team.to_numpy()
expected = analyzedGroup.sort_values(by="Elo", ascending=False).Team.to_numpy()
score += np.sum(( real == expected )*[3,3,2,2]) + 2*len(np.intersect1d(real[:2],expected[:2]))
#Number of people who passed
pointValue=5
self.playoff["Team1Elo"] = self.playoff["Team1"].apply(RealElo)
self.playoff["Team2Elo"] = self.playoff["Team2"].apply(RealElo)
self.playoff["RealWinner"] = self.playoff.apply(WhoWon,axis=1)
self.playoff["ExpectedWinner"] = self.playoff.apply(WhoWonElo,axis=1)
value = {"Quarterfinals":5,"Semifinals":10,"Finals":20}
self.playoff["Value"] = self.playoff["Tab"].apply(lambda x: value[x])
score+=self.playoff["Value"][self.playoff["RealWinner"]==self.playoff["ExpectedWinner"]].sum()
return score
# -
class LoLEloFitnessEvaluation:
def __init__ (self, matchData, groupData, finalPlayOff, region_indexes, years):
self.raters = [ EvaluateElo(matchData[year],
groupData[year],
finalPlayOff[year],
region_indexes) for year in years]
def evaluate(self, individual):
eloModel = EloRating(individual.genes[0], individual.genes[1:-1], individual.genes[-1])
return np.sum([rater.evaluate(eloModel) for rater in self.raters])
# Now the the evaluation function is present, we can proceed to write the code for the genetic algorithm.
# +
class EliteSelection:
def __init__(self):
pass
def select(self, fitnesses, n):
return np.argsort(fitnesses)[::-1][0:n]
class RouletteWheelSelection:
def __init__(self):
pass
def select(self, fitnesses, n):
weights = np.divide (fitnesses,np.sum(fitnesses))
return np.random.choice(np.arange(0,len(fitnesses)),size=n, p=weights)
class AdaptiveGeneticAlgorithm:
def __init__ (self, PopulationGene, SurvivingSelection, ParentsSelection, Mutation, Crossover, FitnessEvaluator):
self.PopulationGene = PopulationGene
self.SurvivingSelection = SurvivingSelection
self.ParentsSelection = ParentsSelection
self.Mutation = Mutation
self.Crossover = Crossover
self.Fitness = FitnessEvaluator
def evolve(self, total_population, generations=10, adaptiveMutationCrossover=True, startingMutationProb = 0.5, startingCrossoverProb = 1.0):
#Initialize individuals
individuals = np.array([self.PopulationGene() for i in range(total_population)])
mutation = startingMutationProb
crossover = startingCrossoverProb
#Calculate fitnesses
print("evaluating Starting Pop fitness")
fitnesses = np.array([self.Fitness.evaluate(individual) for individual in tqdm(individuals)])
order = np.argsort(fitnesses)[::-1]
fitnesses = fitnesses[order]
individuals = individuals[order]
self.best_individual = individuals[0]
self.best_fitness = fitnesses[0]
fitness_historical = np.array(self.best_fitness)
n_gen=0
while(n_gen<generations):
print("Generation "+str(n_gen)+" Best Fitness "+str(self.best_fitness))
new_individuals_mutation = int(total_population*mutation)
new_individuals_crossover = int(total_population*crossover)
new_individuals = np.zeros(new_individuals_crossover+new_individuals_mutation, dtype=np.object)
parents = self.ParentsSelection.select(fitnesses, new_individuals_mutation+2*new_individuals_crossover)
#Create individuals using Mutation
new_individuals[0:new_individuals_mutation] = [
self.Mutation.mutate(individuals[parents[i]]) for i in range(new_individuals_mutation)]
#Create individuals using Crossover
new_individuals[new_individuals_mutation:] = [
self.Crossover.cross(individuals[parents[new_individuals_mutation+2*i]],
individuals[parents[new_individuals_mutation+2*i+1]]) for i in range(new_individuals_crossover)]
#Calculate fitnesses
print("evaluating New Pop fitness")
new_fitnesses = np.array([self.Fitness.evaluate(individual) for individual in tqdm(new_individuals)])
#Select survivors
individuals = np.append(individuals, new_individuals)
fitnesses = np.append(fitnesses, new_fitnesses)
select = self.SurvivingSelection.select(fitnesses, total_population)
fitnesses = fitnesses[select]
individuals = individuals[select]
if(fitnesses[0]>self.best_fitness):
self.best_individual = individuals[0]
self.best_fitness = fitnesses[0]
fitness_historical=np.append(fitness_historical, self.best_fitness)
if(adaptiveMutationCrossover):
non_stagnation = self.best_fitness/np.mean(fitness_historical[::-1][0:5])
mutation = startingMutationProb + non_stagnation*(startingCrossoverProb-startingMutationProb)
crossover = startingCrossoverProb - non_stagnation*(startingCrossoverProb-startingMutationProb)
n_gen+=1
# -
aga = AdaptiveGeneticAlgorithm(
lambda : FloatArrayGene(elo_gene_distr),
EliteSelection(), #Selecting
RouletteWheelSelection(),
MirrorMutation(np.array([50]+[400]*len(regions)+[1])),
GeneFlipCrossover(),
LoLEloFitnessEvaluation(PerYearMatchData, GroupsData, FinalPlayOff, regions_indexes, [2017,2018])
)
aga.evolve(20, generations=20)
individual=aga.best_individual
Evaluate = LoLEloFitnessEvaluation(PerYearMatchData, GroupsData, FinalPlayOff, regions_indexes, [2017,2018])
Evaluate.evaluate(individual)
Evaluate = LoLEloFitnessEvaluation(PerYearMatchData, GroupsData, FinalPlayOff, regions_indexes, [2019])
Evaluate.evaluate(individual)
# It's ok for this part to fail :)
Evaluate = LoLEloFitnessEvaluation(PerYearMatchData, GroupsData, FinalPlayOff, regions_indexes, [2020])
Evaluate.evaluate(individual)
# The important part for now is having the following table, to decide the various winners
GroupsData[2020].sort_values(by=["GroupName","New_Elo"], ascending=False)
eloModel = EloRating(individual.genes[0], individual.genes[1:-1], individual.genes[-1])
eloModel.D
GroupsData[2019].sort_values(by=["GroupName","Elo"], ascending=False)
GroupsData[2020].sort_values(by=["GroupName","Elo"], ascending=False)
{region: eloModel.C[regions_indexes[region]] for region in regions_indexes}
def addStandings(df):
df["Rank"] = 0
for group in df["GroupName"]:
ranking = df[df["GroupName"] == group].sort_values(by=["Elo"], ascending=False).index
df["Rank"].loc[ranking]=np.arange(1,5)
return df.sort_values(by=["GroupName","Rank"])
addStandings(GroupsData[2020])
addStandings(GroupsData[2019])
| Dataset-Scrape-And-Analysis.ipynb |
# # Decision tree for regression
#
# In this notebook, we present how decision trees are working in regression
# problems. We show differences with the decision trees previously presented in
# a classification setting.
#
# First, we load the penguins dataset specifically for solving a regression
# problem.
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Note</p>
# <p class="last">If you want a deeper overview regarding this dataset, you can refer to the
# Appendix - Datasets description section at the end of this MOOC.</p>
# </div>
# +
import pandas as pd
penguins = pd.read_csv("../datasets/penguins_regression.csv")
data_columns = ["Flipper Length (mm)"]
target_column = "Body Mass (g)"
data_train, target_train = penguins[data_columns], penguins[target_column]
# -
# To illustrate how decision trees are predicting in a regression setting, we
# will create a synthetic dataset containing all possible flipper length from
# the minimum to the maximum of the original data.
# +
import numpy as np
data_test = pd.DataFrame(np.arange(data_train[data_columns[0]].min(),
data_train[data_columns[0]].max()),
columns=data_columns)
# +
import matplotlib.pyplot as plt
import seaborn as sns
sns.scatterplot(data=penguins, x="Flipper Length (mm)", y="Body Mass (g)",
color="black", alpha=0.5)
_ = plt.title("Illustration of the regression dataset used")
# -
# We will first illustrate the difference between a linear model and a decision
# tree.
# +
from sklearn.linear_model import LinearRegression
linear_model = LinearRegression()
linear_model.fit(data_train, target_train)
target_predicted = linear_model.predict(data_test)
# -
sns.scatterplot(data=penguins, x="Flipper Length (mm)", y="Body Mass (g)",
color="black", alpha=0.5)
plt.plot(data_test, target_predicted, label="Linear regression")
plt.legend()
_ = plt.title("Prediction function using a LinearRegression")
# On the plot above, we see that a non-regularized `LinearRegression` is able
# to fit the data. A feature of this model is that all new predictions
# will be on the line.
ax = sns.scatterplot(data=penguins, x="Flipper Length (mm)", y="Body Mass (g)",
color="black", alpha=0.5)
plt.plot(data_test, target_predicted, label="Linear regression",
linestyle="--")
plt.scatter(data_test[::3], target_predicted[::3], label="Test predictions",
color="tab:orange")
plt.legend()
_ = plt.title("Prediction function using a LinearRegression")
# Contrary to linear models, decision trees are non-parametric models:
# they do not make assumptions about the way data is distributed.
# This will affect the prediction scheme. Repeating the above experiment
# will highlight the differences.
# +
from sklearn.tree import DecisionTreeRegressor
tree = DecisionTreeRegressor(max_depth=1)
tree.fit(data_train, target_train)
target_predicted = tree.predict(data_test)
# -
sns.scatterplot(data=penguins, x="Flipper Length (mm)", y="Body Mass (g)",
color="black", alpha=0.5)
plt.plot(data_test, target_predicted, label="Decision tree")
plt.legend()
_ = plt.title("Prediction function using a DecisionTreeRegressor")
# We see that the decision tree model does not have an *a priori* distribution
# for the data and we do not end-up with a straight line to regress flipper
# length and body mass.
#
# Instead, we observe that the predictions of the tree are piecewise constant.
# Indeed, our feature space was split into two partitions. Let's check the
# tree structure to see what was the threshold found during the training.
# +
from sklearn.tree import plot_tree
_, ax = plt.subplots(figsize=(8, 6))
_ = plot_tree(tree, feature_names=data_columns, ax=ax)
# -
# The threshold for our feature (flipper length) is 206.5 mm. The predicted
# values on each side of the split are two constants: 3683.50 g and 5023.62 g.
# These values corresponds to the mean values of the training samples in each
# partition.
#
# In classification, we saw that increasing the depth of the tree allowed us to
# get more complex decision boundaries.
# Let's check the effect of increasing the depth in a regression setting:
tree = DecisionTreeRegressor(max_depth=3)
tree.fit(data_train, target_train)
target_predicted = tree.predict(data_test)
sns.scatterplot(data=penguins, x="Flipper Length (mm)", y="Body Mass (g)",
color="black", alpha=0.5)
plt.plot(data_test, target_predicted, label="Decision tree")
plt.legend()
_ = plt.title("Prediction function using a DecisionTreeRegressor")
# Increasing the depth of the tree will increase the number of partition and
# thus the number of constant values that the tree is capable of predicting.
#
# In this notebook, we highlighted the differences in behavior of a decision
# tree used in a classification problem in contrast to a regression problem.
| notebooks/trees_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="l5mSViZt7HqP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="72c1c4b3-9354-4b85-dc7f-fb39c893777d"
import pandas as pd
import numpy as np
import os
import datetime
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, MaxPool2D, Dense, Flatten, Dropout
from tensorflow.keras.utils import to_categorical
# %load_ext tensorboard
import matplotlib.pyplot as plt
from skimage import color, exposure
from sklearn.metrics import accuracy_score
# + id="hxBlUs8t7Vc6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fabc321f-c10e-4a72-82b1-ad2d94e59efd"
# cd '/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_three/dw_matrix_road_sign'
# + id="_8QGLE-G7YxK" colab_type="code" colab={}
train = pd.read_pickle('data/train.p')
test = pd.read_pickle('data/test.p')
x_train, y_train = train['features'], train['labels']
x_test, y_test = test['features'], test['labels']
# + id="ITub0Vt17ss2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7001f955-8017-4d91-a64b-56aa8e548004"
y_train
# + id="Ogj4MP0j7zNM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a600b28a-8fc5-4635-9e5d-6a159a6d3c3f"
len(np.unique(y_train))
# + id="qt45MBX975HL" colab_type="code" colab={}
if y_train.ndim == 1: y_train = to_categorical(y_train)
if y_test.ndim == 1: y_test = to_categorical(y_test)
# + id="HfGp0AU578iQ" colab_type="code" colab={}
input_shape = x_train.shape[1:]
num_classes = y_train.shape[1]
# + id="vXbYVpeG7_E0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="0d2f9fcb-2d3f-4287-948d-15f48207a309"
model = Sequential([
Conv2D(filters=64, kernel_size=(3, 3), activation='relu', input_shape=input_shape),
Flatten(),
Dense(num_classes, activation='softmax'),
])
#model.summary()
model.compile(loss='categorical_crossentropy', optimizer='Adam', metrics=['accuracy'])
model.fit(x_train, y_train)
# + id="0wwrxNnd8C2J" colab_type="code" colab={}
def get_cnn_v1(input_shape, num_classes):
return Sequential([
Conv2D(filters=64, kernel_size=(3, 3), activation='relu', input_shape=input_shape),
Flatten(),
Dense(num_classes, activation='softmax'),
])
def train_model(model, x_train, y_train, params_fit={}):
model.compile(loss='categorical_crossentropy', optimizer='Adam', metrics=['accuracy'])
logdir = os.path.join("logs", datetime.datetime.now().strftime("%Y%m%d-%H%M%S"))
tensorboard_callback = tf.keras.callbacks.TensorBoard(logdir, histogram_freq=1)
model.fit(
x_train,
y_train,
batch_size=params_fit.get('batch_size', 128),
epochs=params_fit.get('epochs', 5),
verbose=params_fit.get('verbose', 1),
validation_data=params_fit.get('validation_data', (x_train, y_train)),
callbacks=[tensorboard_callback]
)
return model
# + id="KJKReT1i8KmS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="c92467cc-95cf-4061-9a7e-aa0d2d85f649"
model = get_cnn_v1(input_shape, num_classes)
model_trained = train_model(model, x_train, y_train)
# + id="FVhts8xG8tnJ" colab_type="code" colab={}
def predict(model_trained, x_test, y_test, scoring=accuracy_score):
y_test_norm = y_pred = np.argmax(y_test, axis=1)
y_pred_prob = model_trained.predict(x_test)
y_pred = np.argmax(y_pred_prob, axis=1)
return scoring(y_test_norm, y_pred)
# + id="Col0xI1PDaJx" colab_type="code" colab={}
def train_and_predict(model):
model_trained = train_model(model, x_train, y_train)
return predict(model_trained, x_test, y_test)
# + id="L92b1XJr8Ofl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="e286614a-dc7d-4dd2-8189-c8a593976184"
def get_cnn_v2(input_shape, num_classes):
return Sequential([
Conv2D(filters=32, kernel_size=(3, 3), activation='relu', input_shape=input_shape),
MaxPool2D(),
Dropout(0.3),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Flatten(),
Dense(1024, activation='relu'),
Dropout(0.3),
Dense(num_classes, activation='softmax'),
])
train_and_predict(get_cnn_v2(input_shape, num_classes))
# + id="3SnHH6sTCKva" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="9028535e-3e0f-4a6d-e309-7bfaf937ac77"
def get_cnn_v3(input_shape, num_classes):
return Sequential([
Conv2D(filters=32, kernel_size=(3, 3), activation='relu', input_shape=input_shape),
Conv2D(filters=32, kernel_size=(3, 3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu'),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Flatten(),
Dense(1024, activation='relu'),
Dropout(0.3),
Dense(num_classes, activation='softmax'),
])
train_and_predict(get_cnn_v3(input_shape, num_classes))
# + id="-2j-9rV9Famg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="a92e339c-14db-42b9-9835-7c151d627aba"
def get_cnn_v4(input_shape, num_classes):
return Sequential([
Conv2D(filters=32, kernel_size=(3, 3), activation='relu', input_shape=input_shape),
Conv2D(filters=32, kernel_size=(3, 3), activation='relu', padding='same'),
MaxPool2D(),
Dropout(0.3),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu', padding='same'),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu', padding='same'),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Flatten(),
Dense(1024, activation='relu'),
Dropout(0.3),
Dense(num_classes, activation='softmax'),
])
train_and_predict(get_cnn_v4(input_shape, num_classes))
# + id="FiomOQyIGYkv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="0da42547-1996-4e9d-b3ed-a113877f07c5"
def get_cnn_v5(input_shape, num_classes):
return Sequential([
Conv2D(filters=32, kernel_size=(3, 3), activation='relu', input_shape=input_shape),
Conv2D(filters=32, kernel_size=(3, 3), activation='relu', padding='same'),
MaxPool2D(),
Dropout(0.3),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu', padding='same'),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu', padding='same'),
Conv2D(filters=64, kernel_size=(3, 3), activation='relu'),
MaxPool2D(),
Dropout(0.3),
Flatten(),
Dense(1024, activation='relu'),
Dropout(0.3),
Dense(1024, activation='relu'),
Dropout(0.3),
Dense(num_classes, activation='softmax'),
])
train_and_predict(get_cnn_v5(input_shape, num_classes))
# + id="6EeWrVaSHY2T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dff85b54-00ff-42ef-e549-4a6b89f25640"
x_train_gray = color.rgb2gray(x_train).reshape(-1, 32, 32, 1)
x_test_gray = color.rgb2gray(x_test).reshape(-1, 32, 32, 1)
# + id="uF5Ph_P0G4YI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="a87d5dba-d74f-4a43-b264-d05b235a7b37"
plt.imshow( color.rgb2gray(x_train[0]), cmap=plt.get_cmap('gray'))
# + id="DNMQx7Mi-nSf" colab_type="code" colab={}
df = pd.read_csv('data/signnames.csv')
labels_dict = df.to_dict()['b']
# + id="LClkRF58-RFC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="33714e6b-d3e4-4cd1-d6b0-137045b31be9"
labels_dict [ np.argmax( y_pred_prob[400] ) ]
# + id="8A9qes85-YMI" colab_type="code" colab={}
# + id="rYcOx8-N9ZJx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="08343bad-ace9-4a31-c29a-7c52e9668933"
plt.imshow(x_test[400])
# + id="_a52e3Kh9eXW" colab_type="code" colab={}
# + id="vSPL4hFR-AqL" colab_type="code" colab={}
# + id="sopi0WZI9zCl" colab_type="code" colab={}
| day4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from time import time
from os import getenv
import pandas as pd
# + [markdown] pycharm={"name": "#%% md\n"}
# # Setting up AWS Credentials
# S3 buckets requires an access key and an access secret, and since nobody wants to
# include a key and a secret in a notebook, we used `python-dotenv` to keep this
# in a `.env` file together with the scripts. If the `.env` file is configured correctly
# the next cell will load that into the current notebook kernel and create the necessary
# parameter dictionary for pandas.
# + pycharm={"name": "#%%\n"}
region_name = 'us-east-1'
aws_key = getenv('AWS_ACCESS_KEY_ID')
aws_secret = getenv('AWS_SECRET_ACCESS_KEY')
s3_path = 's3://geophysics-on-cloud/poseidon/wells/poseidon_geoml_training_wells.json.gz'
s3_options = {
'client_kwargs': {
'aws_access_key_id': aws_key,
'aws_secret_access_key': aws_secret,
'region_name': region_name,
}
}
# -
aws_key, aws_secret
# + [markdown] pycharm={"name": "#%% md\n"}
# # Reading Cloud Wells into Pandas DataFrame
# With the location and configuration above, now we can use the built in `pandas.read_json()`
# function to download and deserialize the `JSON` file into a `pandas.DataFrame()`.
# The file gets downloaded in a few seconds. Parsing the `JSON` takes longer.
# + pycharm={"name": "#%%\n"}
start_time = time()
well_data = pd.read_json(
path_or_buf=s3_path,
compression='gzip',
storage_options=s3_options,
)
well_data.set_index(['well_id', 'twt'], inplace=True)
print(f"Completed read in {time() - start_time} seconds")
# + [markdown] pycharm={"name": "#%% md\n"}
# # Data Description
# ### Raw Data
# + pycharm={"name": "#%%\n"}
well_data
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Statistics
# + pycharm={"name": "#%%\n"}
well_data.describe()
# + [markdown] pycharm={"name": "#%% md\n"}
# # Plotting Wells
# + pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
for well in well_data.index.levels[0]:
fig, ax = plt.subplots(1, 3, sharey='all')
curves = well_data.loc[well][['dtc', 'dts', 'rhob']]
fig.suptitle(well)
for idx, curve in enumerate(curves.columns):
ax[idx].plot(curves[curve], curves[curve].index)
ax[idx].set_ylabel(curves[curve].index.name)
ax[idx].set_xlabel(curves[curve].name)
ax[0].invert_yaxis()
# + pycharm={"name": "#%%\n"}
import seaborn as sns
sns.pairplot(well_data[['dtc', 'dts', 'rhob']])
# + pycharm={"name": "#%%\n"}
| seismic_inversion_2021/notebooks/03_read_cloud_wells.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 用Keras Tuner实现超参数搜索
# 安装keras-tuner:
# ```shell
# pip install -q -U keras-tuner
# ```
# +
import sys
import os
import IPython
import tensorflow as tf
from tensorflow import keras
import kerastuner as kt
print("python version:", sys.version_info)
for module in tf, keras, kt:
print(module.__name__, "version:", module.__version__)
# -
# ### 1. 加载和准备数据集
(img_train, label_train), (img_test, label_test) = keras.datasets.fashion_mnist.load_data()
img_train = img_train.astype("float32") / 255.0
img_test = img_test.astype("float32") / 255.0
# ### 2. 定义模型
# 当构建用于超调整的模型时,除了模型体系结构之外,您还定义了超参数搜索空间。 您为超调优设置的模型称为超模型。
#
# 您可以通过两种方法定义超模型:
# * 通过使用模型构建器功能
# * 通过子类化Keras Tuner API的HyperModel类
#
# 您还可以使用两个预定义的HyperModel类-HyperXception和HyperResNet用于计算机视觉应用程序。
#
# 在本教程中,您将使用模型构建器功能来定义图像分类模型。 模型构建器函数返回已编译的模型,并使用您内联定义的超参数对模型进行超调。
def model_builder(hp):
model = keras.Sequential()
model.add(keras.layers.Flatten(input_shape=(28, 28)))
hp_units = hp.Int("units", min_value=32, max_value=512, step=32)
model.add(keras.layers.Dense(units=hp_units, activation="relu"))
model.add(keras.layers.Dense(10))
hp_learning_rate=hp.Choice("learning_rate", values=[1e-2, 1e-3, 1e-4])
model.compile(optimizer=keras.optimizers.Adam(learning_rate=hp_learning_rate),
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=["accuracy"])
return model
# ### 3.tuner搜索超参数
# 实例化调谐器以执行超调谐。 Keras调谐器有四个可用的调谐器-RandomSearch,Hyperband,BayesianOptimization和Sklearn。 在本教程中,您将使用Hyperband调谐器。
#
# 要实例化Hyperband调谐器,必须指定超模型,要优化的目标以及要训练的最大时期数(max_epochs)。
tuner = kt.Hyperband(
model_builder,
objective="val_accuracy",
max_epochs=10,
factor=3,
directory="my_dir",
project_name="intro_to_kt",
)
# Hyperband 调整算法使用自适应资源分配和提前停止来快速收敛到高性能模型上。 这是使用运动冠军风格的支架完成的。
# 该算法在几个 epoch 内训练了大量模型,并且仅将性能最高的一半模型进行到下一轮。 Hyperband 通过计算 $1 + log_{factor}(max\_epochs)$ 并将其四舍五入到最接近的整数来确定要在括号中训练的模型的数量。
#
#
# 在运行超参数搜索之前,定义一个回调以在每个训练步骤结束时清除训练输出。
class ClearTrainingOutput(keras.callbacks.Callback):
def on_train_end(*args, **kwargs):
IPython.display.clear_output(wait=True)
# 运行超参数搜索。 除了上面的回调外,搜索方法的参数与 keras.model.fit 所使用的参数相同。
# +
tuner.search(img_train, label_train, epochs = 10, validation_data = (img_test, label_test), callbacks = [ClearTrainingOutput()])
# Get the optimal hyperparameters
best_hps = tuner.get_best_hyperparameters(num_trials = 1)[0]
print(f"""
The hyperparameter search is complete. The optimal number of units in the first densely-connected
layer is {best_hps.get('units')} and the optimal learning rate for the optimizer
is {best_hps.get('learning_rate')}.
""")
# -
# 要完成本教程,请使用搜索中的最佳超参数重新训练模型。
model = tuner.hypermodel.build(best_hps)
model.fit(img_train, label_train, epochs=10, validation_data=(img_test, label_test))
# my_dir / intro_to_kt目录包含在超参数搜索期间运行的每个试验(模型配置)的详细日志和checkpoints。
# 如果重新运行超参数搜索,Keras Tuner将使用这些日志中的现有状态来继续搜索。 要禁用此行为,请在实例化调谐器时传递一个额外的overwrite = True参数。
| Chapter01-ML-basics-with-Keras/07-keras-tuner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.3
# language: julia
# name: julia-0.6
# ---
# # Load MEDLINE
# The MEDLINE loader process in BioMedQuery saves the MEDLINE baseline files to a
# MySQL database and saves the raw (xml.gz) and parsed (csv) files to a ```medline```
# directory that will be created in the provided ```output_dir```.
#
# **WARNING:** There are 900+ medline files each with approximately 30,000 articles.
# This process will take hours to run for the full baseline load.
#
# The baseline files can be found [here](ftp://ftp.ncbi.nlm.nih.gov/pubmed/baseline/).
# ### Set Up
# The database and tables must already be created before loading the medline files.
# This process is set up for parallel processing. To take advantage of this, workers
# can be added before loading the BioMedQuery package using the ```addprocs``` function.
using BioMedQuery.DBUtils
using BioMedQuery.PubMed
using BioMedQuery.Processes
# BioMedQuery has utility functions to create the database and tables. *Note: creating
# the tables using this function will drop any tables that already exist in the target
# database.*
const conn = DBUtils.init_mysql_database("127.0.0.1","root","","test_db", true);
PubMed.create_tables!(conn);
# ### Load a Test File
# As the full medline load is a large operation, it is recommended that a test run
# be completed first.
@time Processes.load_medline!(conn, pwd(), test=true)
# Review the output of this run in MySQL to make sure that it ran as expected.
# Additionally, the sample raw and parsed file should be in the new ```medline```
# directory in the current directory.
# ### Performing a Full Load
# To run a full load, use the same code as above, but do not pass the test variable.
# It is also possible to break up the load by passing which files to start and stop at -
# simply pass ```start_file=n``` and ```end_file=p```.
#
# After loading, it is recommended you add indexes to the tables, the ```add_mysql_keys!```
# function can be used to add a standard set of indexes.
add_mysql_keys!(conn)
# *This notebook was generated using [Literate.jl](https://github.com/fredrikekre/Literate.jl).*
| docs/src/notebooks/5_load_medline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# __<NAME> & <NAME>__
# __Imports__
import numpy as np
import pandas as pd
from pyspark.sql import SparkSession
from google_drive_downloader import GoogleDriveDownloader as gdd
from pyspark.ml.feature import RegexTokenizer, StopWordsRemover, CountVectorizer
from pyspark.ml.classification import LogisticRegression
import nltk
from nltk.corpus import stopwords
import string
from pyspark.ml import Pipeline
from pyspark.ml.feature import OneHotEncoder, StringIndexer, VectorAssembler
from pyspark.sql.functions import col
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
# __Download File from link given in Canvas__
#
# This will be stored into your local...Do not add into git, file is too large to be pushed onto git master branch
#
# So, we download locally
gdd.download_file_from_google_drive(file_id='0B04GJPshIjmPRnZManQwWEdTZjg',
dest_path='/Users/mwoo/Downloads/trainingandtestdata.zip',
unzip=True)
# +
# gdd.download_file_from_google_drive(file_id='0B04GJPshIjmPRnZManQwWEdTZjg',
# dest_path='/Users/swapnilbasu/Downloads/trainingandtestdata.zip',
# unzip=True)
# -
# __Create spark session object (Data Processing)__
spark=SparkSession.builder.appName('classification_tweet').getOrCreate()
# __Load in data__
training_data = spark.read.csv("/Users/mwoo/Downloads/training.1600000.processed.noemoticon.csv",header=False)
# __Renaming columns__
training_data.columns
training_data = training_data.toDF("target",'id','date','query','user_name','text')
training_data.columns
# __Exploratory__
training_data.describe()
# __Selecting the target value and text__
#
# These are the values that we believe are important in predicting whether a tweet was negative or positive
df = training_data.select('text','target')
df.show(5)
df.printSchema()
# We can see below that its an even split between positive and negative tweets
#
# 0: negative
# 4: positive
df.groupBy("target").count().orderBy(col("count").desc()).show()
# __Model Pipeline__
# __Regular Expression Tokenizer__
#
# Seperates the texts into words
regexTokenizer = RegexTokenizer(inputCol="text", outputCol="words", pattern="\\W")
# __Stop Words Download from NLTK__
#
# Downloads Stop words
nltk.download('stopwords')
# __Stop Words Remover__
#
# Remove unnecessary words
sp = set(string.punctuation)
stop_words = set(stopwords.words('english'))
extra_words = {"http","https","amp","rt","t","c","the"}
for i in extra_words:
stop_words.add(i)
stop_words = list(stop_words)
stopwordsRemover = StopWordsRemover(inputCol="words", outputCol="filtered").setStopWords(stop_words)
# __Bag of words count__
#
# This is a type of feature engineering
countVectors = CountVectorizer(inputCol="filtered", outputCol="features", vocabSize=10000, minDF=5)
# __StringIndexer__
#
# Indexing target values
label_stringIdx = StringIndexer(inputCol = "target", outputCol = "label")
pipeline = Pipeline(stages=[regexTokenizer, stopwordsRemover, countVectors, label_stringIdx])
pipelineFit = pipeline.fit(df)
dataset = pipelineFit.transform(df)
dataset.show(5)
# __Selecting data from the previous dataframe__
dataset = dataset.select('text','features','label')
# __Set seed for reproducibility__
#
# This set is use for testing purposes 70/30 split
(trainingData, testData) = dataset.randomSplit([0.7, 0.3], seed = 100)
print("Training Dataset Count: " + str(trainingData.count()))
print("Test Dataset Count: " + str(testData.count()))
# This will be used to fully train our classifcation model
model_df = dataset.select('features','label')
print("Full Training Dataset Count: " + str(model_df.count()))
# __Testing our model through the split data above__
lr = LogisticRegression(maxIter=20, regParam=0.3, elasticNetParam=0).fit(trainingData)
predictions = lr.transform(testData)
predictions.filter(predictions['prediction'] == 0) \
.select("text","probability","label","prediction") \
.orderBy("probability", ascending=False) \
.show(n = 10, truncate = 30)
predictions = lr.transform(testData)
predictions.show(10)
predictions.printSchema()
from pyspark.ml.evaluation import BinaryClassificationEvaluator
bce = BinaryClassificationEvaluator()
print("Test Area Under ROC: " + str(bce.evaluate(predictions, {bce.metricName: "areaUnderROC"})))
df_selection = predictions.select("label",'prediction').toPandas()
from sklearn.metrics import classification_report
true = np.array(df_selection['label'])
pred = np.array(df_selection['prediction'])
print(classification_report(true,pred))
# __We can say that this model is able to distinguish whether a tweet is positive or negative is fair__
# __Retrain our model using the fully dataset__
lr = LogisticRegression(maxIter=20, regParam=0.3, elasticNetParam=0).fit(model_df)
# __Twitter Authentication__
# +
import tweepy
from tweepy import OAuthHandler
from tweepy import Stream
ACCESS_TOKEN = "1458842253779161088-QFeO6udaAdHR4VARxaDza1w4LUlooE"
ACCESS_TOKEN_SECRET = "<KEY>"
API_KEY = "<KEY>"
API_KEY_SECRET = "<KEY>"
auth = tweepy.OAuthHandler(API_KEY, API_KEY_SECRET)
auth.set_access_token(ACCESS_TOKEN, ACCESS_TOKEN_SECRET)
api = tweepy.API(auth)
# -
# __Twitter Streaming Tweets__
#
# We can set the limit of tweet samples to 100 and where the streaming only streams tweets that are in english
# +
tweet_list = list()
# Subclass Stream to print IDs of Tweets received
class IDPrinter(tweepy.Stream):
def on_status(self, status):
tweet_list.append(status.text)
#print(tweet_list)
#print(status.text)
if len(tweet_list) == 100:
Stream.disconnect(self)
# Initialize instance of the subclass
printer = IDPrinter(
API_KEY, API_KEY_SECRET,
ACCESS_TOKEN, ACCESS_TOKEN_SECRET
)
printer.sample(languages=['en'])
# -
# __Create new dataframe from tweet stream__
df_2 = pd.DataFrame(np.array(tweet_list))
df_2.columns = ['text']
df_2 = spark.createDataFrame(df_2)
df_2.show()
# __Transforms the data received from stream__
dataset_1 = pipelineFit.transform(df_2)
dataset_1.show()
# __Selecting text and Features__
dataset_test = dataset_1.select('text','features')
# __Having the pre-trained model predict on tweets__
model_predictions = lr.transform(dataset_test)
model_predictions.show()
# __Storing Results and Text in Database__
# Create connection to mongoDB, database, and table
import pymongo
myclient = pymongo.MongoClient("mongodb://localhost:27017/")
mydb = myclient["my_database"]
mycol = mydb["predictions"]
# Converting spark dataframe to pandas dataframe
mp_df = model_predictions.toPandas()
mp_df.head()
# __Inserting into MongoDB__
for index, row in mp_df.iterrows():
#print(row['text'], row['prediction'])
mydict = { "text": row['text'], "prediction": row['prediction'] }
mycol.insert_one(mydict)
# __Definitions__
#
# 0.0 means that the tweet was negative
#
# 1.0 means that the tweet was positive
# __Retrieving Positive Tweets__
myquery = { "prediction": 1.0 }
mydoc = mycol.find(myquery)
for x in mydoc:
print(x)
# __Retrieving Negative Tweets__
myquery = { "prediction": 0.0 }
mydoc = mycol.find(myquery)
for x in mydoc:
print(x)
| Spark_Twitter_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:wildfires] *
# language: python
# name: conda-env-wildfires-py
# ---
# ## Setup
from specific import *
# ### Get shifted data
(
endog_data,
exog_data,
master_mask,
filled_datasets,
masked_datasets,
land_mask,
) = get_offset_data()
client = get_client()
client
# ### Define the training and test data
# +
@data_split_cache
def get_split_data():
X_train, X_test, y_train, y_test = train_test_split(
exog_data, endog_data, random_state=1, shuffle=True, test_size=0.3
)
return X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = get_split_data()
# -
# ### Specific model training without grid seach
rf = get_model(X_train, y_train)
| analyses/seasonality_paper_2/no_temporal_shifts/model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Let's see if we detected anything this time?
# +
import numpy as np
import matplotlib.pyplot as plt
import astropy.io.fits as fits
import os
import glob
from astropy.table import Table
from astropy.io import ascii
import astropy.units as u
import astropy.constants as const
#matplotlib set up
# %matplotlib inline
from matplotlib import rcParams
rcParams["figure.figsize"] = (14, 5)
rcParams["font.size"] = 20
# -
path = '/media/david/5tb_storage1/muscles/ltt1445/hst/redo_data/'
x1ds = glob.glob(path+'*x1d.fits')
print(x1ds)
sx1s = glob.glob(path+'sx1.fits')
print(sx1s)
# No sx1 - no sign of any flux in the ccd raw images. Huh? There should at least be something at longer wavelengths?
for x in x1ds:
hdul = fits.open(x)
print(hdul[0].header['OPT_ELEM'])
data = hdul[1].data[0]
hdul.close()
#plt.step(data['WAVELENGTH'][data['DQ']==0], data['FLUX'][data['DQ']==0])
plt.step(data['WAVELENGTH'], data['DQ'])
plt.show()
hdul[1].header
# DQ filters out everything! No Mg ii lines- check raw data...
# Mucking around with the Swift data to do a gif...
spath = '/media/david/5tb_storage1/muscles/ltt1445/swift/'
sobs = glob.glob(spath+'00*')
sobs
from matplotlib.colors import LogNorm
from pylab import cm
for sob in sobs:
dpath = glob.glob('{}/uvot/event/clean*'.format(sob))[0]
data = fits.getdata(dpath, 1)
plt.figure(figsize=(5, 5))
NBINS = (500,500)
img_zero_mpl = plt.hist2d(data['RAWX'], data['RAWY'], NBINS, cmap='gray', norm=LogNorm())
lims = [1000, 1300]
plt.xlim(lims)
plt.ylim(lims)
#cbar = plt.colorbar(ticks=[1.0,3.0,6.0])
# cbar.ax.set_yticklabels(['1','3','6'])
# plt.xlabel('x')
#plt.ylabel('y')
# plt.hist2d(data['RAWX'], data['RAWY'], bins=100)
plt.show()
# +
from astropy.time import Time
from astropy.coordinates import SkyCoord, Distance
c = SkyCoord(ra='03h01m51.39s', dec='-16d35m36.1s', distance = Distance(parallax=0.1455e3*u.mas), frame='icrs', pm_ra_cosdec = -369.2*u.mas/u.yr, pm_dec= -268.3*u.mas/u.yr, obstime=Time(2015.05, format='decimalyear'))
c
# -
c2 = c.apply_space_motion(new_obstime=Time('2020-06-17 10:13:11'))
c2.ra.hms
c2.dec.dms
1/0.1455
c3 = SkyCoord(ra=45.46247805566359*u.degree, dec=-16.594495614724412*u.degree, distance = Distance(parallax=0.1455e3*u.mas), frame='icrs', pm_ra_cosdec = -369.2*u.mas/u.yr, pm_dec= -268.3*u.mas/u.yr, obstime=Time(2015.05, format='decimalyear'))
c4 = c3.apply_space_motion(new_obstime=Time('2020-06-17 10:13:11'))
c4.ra.hms
c4.dec.dms
c3.ra.hms
c3.dec.dms
c5 = c.apply_space_motion(new_obstime=Time('2003-06-18 23:52:38'))
print(c5.ra.hms, c5.dec.hms)
hdul = fits.open(path +'oe83h3010_flt.fits')
# +
#hdul[1].header
# -
lc100 = fits.getdata('/media/david/5tb_storage1/muscles/ltt1445/swift/00012837002/uvot/event/ltt1445A_1800slc_002.fits', 1)
#plt.errorbar(lc100['TIME'][2:], lc100['AB_MAG'][2:], lc100['AB_MAG_ERR'][2:], ls='none', marker='o')
mask = ((lc100['AB_MAG']/lc100['AB_MAG_ERR']) > 10)
lc100 = lc100[mask]
plt.errorbar(lc100['TIME'], lc100['AB_MAG'], lc100['AB_MAG_ERR'], ls='none', marker='o')
plt.ylim(0, 30)
len(lc100['AB_MAG'])
print (lc100['AB_MAG'], lc100['AB_MAG_ERR'])
mf = np.median(lc100['AB_MAG'])
me = mf*np.sum((lc100['AB_MAG_ERR']/lc100['AB_MAG'])**2)**0.5
print(mf, me)
mf = np.median(lc100['AB_FLUX_AA'])
me = mf*np.sum((lc100['AB_FLUX_AA_ERR']/lc100['AB_FLUX_AA'])**2)**0.5
print(mf, me)
mf = np.mean(lc100['MAG'])
me = mf*np.sum((lc100['MAG_ERR']/lc100['MAG'])**2)**0.5
print(mf, me)
mf = np.mean(lc100['FLUX_AA'])
me = mf*np.sum((lc100['FLUX_AA_ERR']/lc100['FLUX_AA'])**2)**0.5
print(mf, me)
lc100.names
(614067120.916038-614048228.21956)/60
| ltt1445a_redo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Data preparation
import numpy as np
np.random.seed(42)
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_breast_cancer
# +
dataset = load_breast_cancer()
x = dataset.data
y = dataset.target
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.2)
print(f"x_train shape: {x_train.shape} x_test.shape: {x_test.shape}")
# -
# #### Grid Search
from sklearn.model_selection import GridSearchCV
# +
# Params for KNN: n_neighbors and weights
parameters = {
'n_neighbors': [3, 5, 7, 9],
'weights': ['uniform', 'distance']
}
clf = KNeighborsClassifier()
grid_cv = GridSearchCV(clf, parameters, cv=10)
grid_cv.fit(x_train, y_train)
# -
print("GridSearch Keys:")
for key in grid_cv.cv_results_.keys():
print(f"\t{key}")
print(f"GridSearch Params: {grid_cv.cv_results_['params']}")
# +
print(f"Best parameters set found on development set: {grid_cv.best_params_}\n")
means = grid_cv.cv_results_['mean_test_score']
stds = grid_cv.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, grid_cv.cv_results_['params']):
print(f"{mean:.3f} (+/-{2*std:.3f}) for {params}")
# -
# #### Best Found model
clf = KNeighborsClassifier(n_neighbors=3, weights="distance")
clf.fit(x_train, y_train)
score = clf.score(x_test, y_test)
print(f"Accuracy: {score}")
| Chapter8_MetricsAndEvaluation/GridSearch/GridSearch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Notebook 3b - Breakdown Soil Type
#
# In this notebook, we test out breaking down the soil-type features using domain knowledge and their descriptions
# Global variables for testing changes to this notebook quickly
RANDOM_SEED = 0
NUM_FOLDS = 12
# +
import numpy as np
import pandas as pd
import time
import pyarrow
import gc
# Model evaluation
from functools import partial
from sklearn.base import clone
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import StratifiedKFold, train_test_split
from sklearn.metrics import accuracy_score, recall_score
from sklearn.inspection import partial_dependence, permutation_importance
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import AdaBoostClassifier, BaggingClassifier, ExtraTreesClassifier, RandomForestClassifier
# Plotting
import matplotlib
import seaborn as sns
from matplotlib import pyplot as plt
# Hide warnings
import warnings
warnings.filterwarnings('ignore')
# -
# # Prepare Data
# Encode soil type
def categorical_encoding(input_df):
data = input_df.copy()
data['Soil_Type'] = 0
soil_features = list()
for i in range(1,41):
data['Soil_Type'] += i*data[f'Soil_Type{i}']
soil_features.append(f'Soil_Type{i}')
nonsoil_features = [x for x in data.columns if x not in soil_features]
return data[nonsoil_features]
# +
# %%time
# Load original data
original = categorical_encoding(pd.read_feather('../data/original.feather'))
# Label Encode
old_encoder = LabelEncoder()
original["Cover_Type"] = old_encoder.fit_transform(original["Cover_Type"])
y_train = original['Cover_Type'].iloc[:15119]
y_test = original['Cover_Type'].iloc[15119:]
# Get feature columns
features = [x for x in original.columns if x not in ['Id','Cover_Type']]
# Data structures for summary scores
bagging_scores = list()
extratrees_scores = list()
adaboost_scores = list()
random_scores = list()
# -
# # Scoring Function
def train_original(sklearn_model, processing = None):
# Original Training/Test Split
X_temp = original[features].iloc[:15119]
X_test = original[features].iloc[15119:]
y_temp = original['Cover_Type'].iloc[:15119]
y_test = original['Cover_Type'].iloc[15119:]
# Feature Engineering
if processing:
X_temp = processing(X_temp)
X_test = processing(X_test)
# Store the out-of-fold predictions
test_preds = np.zeros((X_test.shape[0],7))
oof_preds = np.zeros((X_temp.shape[0],))
scores, times = np.zeros(NUM_FOLDS), np.zeros(NUM_FOLDS)
# Stratified k-fold cross-validation
skf = StratifiedKFold(n_splits = NUM_FOLDS, shuffle = True, random_state = RANDOM_SEED)
for fold, (train_idx, valid_idx) in enumerate(skf.split(X_temp,y_temp)):
# Training and Validation Sets
X_train, X_valid = X_temp.iloc[train_idx], X_temp.iloc[valid_idx]
y_train, y_valid = y_temp.iloc[train_idx], y_temp.iloc[valid_idx]
# Create model
start = time.time()
model = clone(sklearn_model)
model.fit(X_train, y_train)
# validation and test predictions
valid_preds = np.ravel(model.predict(X_valid))
oof_preds[valid_idx] = valid_preds
test_preds += model.predict_proba(X_test)
# Save scores and times
scores[fold] = accuracy_score(y_valid, valid_preds)
end = time.time()
times[fold] = end-start
time.sleep(0.5)
test_preds = np.argmax(test_preds, axis = 1)
test_score = accuracy_score(y_test, test_preds)
print('\n'+model.__class__.__name__)
print("Train Accuracy:", round(scores.mean(), 5))
print('Test Accuracy:', round(test_score, 5))
print(f'Training Time: {round(times.sum(), 2)}s')
return scores.mean(), oof_preds, test_score
# # Models
#
# We use the following 4 models from the scikit-learn library:
#
# 1. AdaBoost
# 2. ExtraTrees
# 3. Bagging
# 4. Random Forest
# +
# AdaBoost Classifier
adaboost = AdaBoostClassifier(
base_estimator = DecisionTreeClassifier(
splitter = 'random',
random_state = RANDOM_SEED,
),
random_state = RANDOM_SEED,
)
# ExtraTrees Classifier
extratrees = ExtraTreesClassifier(
n_jobs = -1,
random_state = RANDOM_SEED,
max_features = None,
)
# Bagging Classifier
bagging = BaggingClassifier(
base_estimator = DecisionTreeClassifier(
splitter = 'random',
random_state = RANDOM_SEED,
),
n_jobs = -1,
random_state = RANDOM_SEED
)
# Random Forest Classifier
randomforest = RandomForestClassifier(
n_jobs = -1,
random_state = RANDOM_SEED,
)
# -
# # Baselines
# +
# AdaBoost
cv_score, oof_preds, test_score = train_original(adaboost)
adaboost_scores.append((
'Baseline', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# ExtraTrees
cv_score, oof_preds, test_score = train_original(extratrees)
extratrees_scores.append((
'Baseline', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Bagging
cv_score, oof_preds, test_score = train_original(bagging)
bagging_scores.append((
'Baseline', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Random Forest
cv_score, oof_preds, test_score = train_original(randomforest)
random_scores.append((
'Baseline', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# -
# # Soil Type Features
#
# We test the following features based off of the soil type descriptions from the original data:
#
# 1. Climatic Zone (Ordinal)
# 2. Geologic Zone (Nominal)
# 3. Surface Cover (Ordinal)
# 4. Rock Size (Ordinal)
# ## Climatic Zone (Ordinal)
#
# We create a feature based on the climatic zone of the soil, which has a natural ordering:
#
# 1. lower montane dry
# 2. lower montane
# 3. montane dry
# 4. montane
# 5. montane dry and montane
# 6. montane and subalpine
# 7. subalpine
# 8. alpine
#
# However, the ordering of the soil type labels roughly follows the ordering of their respectively climatic zones, so there's a chance this feature won't be particularly informative.
def climatic_zone_original(input_df):
code = {
1:2702,2:2703,3:2704,4:2705,5:2706,6:2717,7:3501,8:3502,9:4201,
10:4703,11:4704,12:4744,13:4758,14:5101,15:5151,16:6101,17:6102,
18:6731,19:7101,20:7102,21:7103,22:7201,23:7202,24:7700,25:7701,
26:7702,27:7709,28:7710,29:7745,30:7746,31:7755,32:7756,33:7757,
34:7790,35:8703,36:8707,37:8708,38:8771,39:8772,40:8776
}
df = input_df.copy()
df['Climatic_Zone'] = input_df['Soil_Type'].apply(lambda x: int(str(code[x])[0]))
return df
# +
# AdaBoost
cv_score, oof_preds, test_score = train_original(adaboost, climatic_zone_original)
adaboost_scores.append((
'Climatic_Zone', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Extra Trees
cv_score, oof_preds, test_score = train_original(extratrees, climatic_zone_original)
extratrees_scores.append((
'Climatic_Zone', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Bagging
cv_score, oof_preds, test_score = train_original(bagging, climatic_zone_original)
bagging_scores.append((
'Climatic_Zone', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Random Forest
cv_score, oof_preds, test_score = train_original(randomforest, climatic_zone_original)
random_scores.append((
'Climatic_Zone', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# -
# ## Geologic Zones (Nominal)
#
# This is another feature which is based on the soil type codes, but is not ordered like climatic zone.
#
# 1. alluvium
# 2. glacial
# 3. shale
# 4. sandstone
# 5. mixed sedimentary
# 6. unspecified in the USFS ELU Survey
# 7. igneous and metamorphic
# 8. volcanic
def geologic_zone_original(input_df):
code = {
1:2702,2:2703,3:2704,4:2705,5:2706,6:2717,7:3501,8:3502,9:4201,
10:4703,11:4704,12:4744,13:4758,14:5101,15:5151,16:6101,17:6102,
18:6731,19:7101,20:7102,21:7103,22:7201,23:7202,24:7700,25:7701,
26:7702,27:7709,28:7710,29:7745,30:7746,31:7755,32:7756,33:7757,
34:7790,35:8703,36:8707,37:8708,38:8771,39:8772,40:8776
}
df = input_df.copy()
df['Geologic_Zone'] = input_df['Soil_Type'].apply(lambda x: int(str(code[x])[1]))
return df
# +
# AdaBoost
cv_score, oof_preds, test_score = train_original(adaboost, geologic_zone_original)
adaboost_scores.append((
'Geologic_Zone', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Extra Trees
cv_score, oof_preds, test_score = train_original(extratrees, geologic_zone_original)
extratrees_scores.append((
'Geologic_Zone', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Bagging
cv_score, oof_preds, test_score = train_original(bagging, geologic_zone_original)
bagging_scores.append((
'Geologic_Zone', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Random Forest
cv_score, oof_preds, test_score = train_original(randomforest, geologic_zone_original)
random_scores.append((
'Geologic_Zone', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# -
# ## Surface Cover (Ordinal)
#
# According to the [USDA reference](https://www.nrcs.usda.gov/wps/portal/nrcs/detail/soils/ref/?cid=nrcs142p2_054253#surface_fragments) on soil profiling:
#
# 1. **(Stony/Bouldery)** — Stones or boulders cover 0.01 to less than 0.1 percent of the surface. The smallest stones are at least 8 meters apart; the smallest boulders are at least 20 meters apart (fig. 3-9).
#
# 2. **(Very Stony/Very Bouldery)** — Stones or boulders cover 0.1 to less than 3 percent of the surface. The smallest stones are not less than 1 meter apart; the smallest boulders are not less than 3 meters apart (fig. 3-10).
#
# 3. **(Extremely Stony/Extremely Bouldery)** — Stones or boulders cover 3 to less than 15 percent of the surface. The smallest stones are as little as 0.5 meter apart; the smallest boulders are as little as 1 meter apart (fig. 3-11).
#
# 4. **(Rubbly)** — Stones or boulders cover 15 to less than 50 percent of the surface. The smallest stones are as little as 0.3 meter apart; the smallest boulders are as little as 0.5 meter apart. In most places it is possible to step from stone to stone or jump from boulder to boulder without touching the soil (fig. 3-12).
#
# 5. **(Very Rubbly)** — Stones or boulders appear to be nearly continuous and cover 50 percent or more of the surface. The smallest stones are less than 0.03 meter apart; the smallest boulders are less than 0.05 meter apart. Classifiable soil is among the rock fragments, and plant growth is possible (fig. 3-13).
def surface_cover_original(input_df):
# Group IDs
no_desc = [7,8,14,15,16,17,19,20,21,23,35]
stony = [6,12]
very_stony = [2,9,18,26]
extremely_stony = [1,22,24,25,27,28,29,30,31,32,33,34,36,37,38,39,40]
rubbly = [3,4,5,10,11,13]
# Create dictionary
surface_cover = {i:0 for i in no_desc}
surface_cover.update({i:1 for i in stony})
surface_cover.update({i:2 for i in very_stony})
surface_cover.update({i:3 for i in extremely_stony})
surface_cover.update({i:4 for i in rubbly})
# Create Feature
df = input_df.copy()
df['Surface_Cover'] = input_df['Soil_Type'].apply(lambda x: surface_cover[x])
return df
# +
# AdaBoost
cv_score, oof_preds, test_score = train_original(adaboost, surface_cover_original)
adaboost_scores.append((
'Surface_Cover', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Extra Trees
cv_score, oof_preds, test_score = train_original(extratrees, surface_cover_original)
extratrees_scores.append((
'Surface_Cover', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Bagging
cv_score, oof_preds, test_score = train_original(bagging, surface_cover_original)
bagging_scores.append((
'Surface_Cover', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Random Forest
cv_score, oof_preds, test_score = train_original(randomforest, surface_cover_original)
random_scores.append((
'Surface_Cover', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# -
# ## Rock Size (Ordinal)
def rock_size_original(input_df):
# Group IDs
no_desc = [7,8,14,15,16,17,19,20,21,23,35]
stones = [1,2,6,9,12,18,24,25,26,27,28,29,30,31,32,33,34,36,37,38,39,40]
boulders = [22]
rubble = [3,4,5,10,11,13]
# Create dictionary
rock_size = {i:0 for i in no_desc}
rock_size.update({i:1 for i in stones})
rock_size.update({i:2 for i in boulders})
rock_size.update({i:3 for i in rubble})
df = input_df.copy()
df['Rock_Size'] = input_df['Soil_Type'].apply(lambda x: rock_size[x])
return df
# +
# AdaBoost
cv_score, oof_preds, test_score = train_original(adaboost, rock_size_original)
adaboost_scores.append((
'Rock_Size', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Extra Trees
cv_score, oof_preds, test_score = train_original(extratrees, rock_size_original)
extratrees_scores.append((
'Rock_Size', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Bagging
cv_score, oof_preds, test_score = train_original(bagging, rock_size_original)
bagging_scores.append((
'Rock_Size', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# Random Forest
cv_score, oof_preds, test_score = train_original(randomforest, rock_size_original)
random_scores.append((
'Rock_Size', cv_score, test_score,
*recall_score(y_train, oof_preds, average = None)
))
# -
# # Summary
#
# All of these features seem promising, so we won't rule any out just yet.
# AdaBoost
pd.DataFrame.from_records(
data = adaboost_scores,
columns = ['features','cv_score','holdout','recall_0', 'recall_1','recall_2','recall_3','recall_4','recall_5','recall_6']
).sort_values('cv_score')
# Extra Trees Classifier
pd.DataFrame.from_records(
data = extratrees_scores,
columns = ['features','cv_score','holdout','recall_0', 'recall_1','recall_2','recall_3','recall_4','recall_5','recall_6']
).sort_values('cv_score')
# Bagging Classifier
pd.DataFrame.from_records(
data = bagging_scores,
columns = ['features','cv_score','holdout','recall_0', 'recall_1','recall_2','recall_3','recall_4','recall_5','recall_6']
).sort_values('cv_score')
# Random Forest
pd.DataFrame.from_records(
data = random_scores,
columns = ['features','cv_score','holdout','recall_0', 'recall_1','recall_2','recall_3','recall_4','recall_5','recall_6']
).sort_values('cv_score')
| notebooks/Notebook 3a - Soil Feature Engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # While statements and arrays
# ## Legend
# <div class = "alert alert-block alert-info">
# In <b> blue</b>, the <b> instructions </b> and <b> goals </b> are highlighted.
# </div>
# <div class = "alert alert-block alert-success">
# In <b> green</b>, the <b> information </b> is highlighted.
# </div>
# <div class = "alert alert-block alert-warning">
# In <b> yellow</b>, the <b> exercises </b> are highlighted.
# </div>
# <div class = "alert alert-block alert-danger">
# In <b> red</b>, the <b> error </b> and <b> alert messages </b> are highlighted.
# </div>
# ## Instructions
# <div class = "alert alert-block alert-info">
# In <b> red</b>, the <b> error </b> and <b> alert messages </b> are highlighted.
# Click "Run" on each cell to go through the code in each cell. <br> <br> This will take you through the cell and print out the results.
# If you wish to see all the outputs at once in the whole notebook, just click Cell and then Run All. In case the cell keeps running and does not stop, go to the respective cell, press <b> Kernel</b>, then <b> Interrupt</b>.
# </div>
# ## Goals
# <div class = "alert alert-block alert-info">
# After this workshop, the student should get more familiar with the following topics: <br>
# <ul>
# <li> printing basic statements and commands in Jupyter Notebook</li>
# <li> performing basic arithmetic calculations in Python</li>
# <li> improving an existent model of the code</li>
# <li> recognizing and checking variable types in Python </li>
# <li> using the <b> if </b> and <b> for </b> statements for basic operations
# <li> working with <b> characters </b> and <b> strings </b>
# </ul>
#
# <b> These objectives are in agreement with the Advanced Higher Scottish Curriculum for high-school students. </b> <br> <br> <b> Note: </b> For most of the workshop, the student will be given some coding examples. In some cases, the student will have to code himself/herself. The coding part is optional, but highly recommended to approach. <br> <br> <b> Note: </b> The game at the end of the notebook is completely optional material. You can either only play it, or analyze how it is designed (and even come up with suggestions). However, the coding there is a bit more advanced... after a couple of weeks, you will be equipped with more tools for designing small games.
#
# </div>
# ## Explore
# ### Conditional while statement...
# <div class = "alert alert-block alert-success">
# Suppose that, in the case where we do not know how many times we would like to print the statement. All we would like is the following: <b> while </b> a condition is not reached yet, we keep <b> executing </b> the instruction. This is the main role of the <b> while </b> instruction.
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> Investigate the following code below. <b> Predict </b> the ouptut, then <b> Run </b> the cell.
# </div>
i = 0
while(i < 10):
print("Welcome to this programming session!")
i += 1
# <div class = "alert alert-block alert-success">
# The <b> while </b> instruction plays indeed a similar role to the <b> for </b> statement. <b> However</b>, there are differences. This time, we do not instruct the computer <b> how many </b> steps we want to perform. The boundary for the above program is set by the condition: $ i == 10 $
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Investigate: </b> What happens in the following case?
# </div>
i = 0
while(i < 10):
print("Welcome to this programming session!")
# <div class = "alert alert-block alert-danger">
# We have skipped the crucial statement: $ i += 1 $. Hence, $ i == 0 $, and the statement $ 0 < 10 $ holds <b> forever</b>. It is of <b> utmost importance </b> to establish from the beginning condition to advance in the while statement, in order to avoid getting stuck. <b> Note: </b> This is done automatically when in a loop.
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> let us <b> analyze </b> this phenomenon in greater detail, by <b> debugging </b> the code.
# </div>
# <div class = "alert alert-block alert-success">
# <b> Exercise: </b> <b> Debugging </b> involves an <b> in-depth </b> analyss of the code. At each step, you will tell the computer what step to perform and what to print out. This is the most <b> accurate </b> way to check how the computer performs the algorithm. Let us see together how <b> debugging </b> is done in Jupyter Notebooks. Firstly, there is a specific library which needs to be imported
# </div>
import pdb
# <div class = "alert alert-block alert-success">
# Afterwards, a command line <b> breakpoint </b> is added. That is where you will have to play with all the variables. You will need in particular the following commands:
#
# <ul>
# <li> <b> n </b> is for going to te next command line</li>
# <li> <b> p </b> is for printing out the value(example: p b)</li>
# <li> <b> w </b> is for finding out your location</li>
# <li> <b> q </b> is for quitting the debugger</li>
# </ul>
# </div>
# +
my_sum = 0
i = 0
while(i<10):
breakpoint()
my_sum += i
my_sum
# -
# <div class = "alert alert-block alert-danger">
# You should figure out that our conditional variable $ i $ does <b> not </b> change. This is why we never get out from the loop.
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> <b> Add </b> the condition of instruction exit (<b> Hint: </b> you only need one line of code)
# </div>
# Write your own code here
i = 0
sum = 0
while(i < 10):
sum += i
print("The sum of the first " + "10 " + "elements is: " + str(sum))
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> <b> Compare </b> this method of calculating sum of first $ N $ consecutive elements with the for-method. Afterwards, use <b> while </b> instruction to calculate the product of the first 10 elements
# </div>
# +
# Write your own code here
# -
# ### ...And Arrays
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> <b> Investigate </b> the code below, which tells you how to find the greatest value of an element in an array.
# </div>
# <div class = "alert alert-block alert-success">
# <b> Approach: </b>
# <ul>
# <li> <b> Initiate </b> a dummy element with very small value (in our problem, it will be a <b> negative </b> number </li>
# <li> <b> Go </b> through the array. For <b> each </b> element greater than our test variable, the dummy number takes the value of the element in question
# <li> <b> Print </b> the test element </li>
# </ul>
# </div>
# +
my_trial_array = [23,1,54,23,65,75,234,0] # This is my test array created (Notice how the elements can repeat themselves)
maxx = -1 # Test variable
for i in range(len(my_trial_array)):
if(maxx < my_trial_array[i]):
maxx = my_trial_array[i]
print("The maximum element of the test array is: " + str(maxx))
# -
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> <b> Debug </b> the code, to make sure you understand what happens.
# </div>
# +
import pdb
my_trial_array = [23,1,54,23,65,75,234,0] # This is my test array created (Notice how the elements can repeat themselves)
maxx = -1 # Test variable
for i in range(len(my_trial_array)):
breakpoint()
if(maxx < my_trial_array[i]):
maxx = my_trial_array[i]
print("The maximum element of the test array is: " + str(maxx))
# -
# <div class = "alert alert-block alert-success">
# Just as arrays can be created for integers, they can be created for characters as well. They are called <b> strings. </b> Similarly, all the characters (data type) are stored in such a string. The elements are, once again, easily accessible.
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Investigate: </b> How is a string declared? Can you see the similarity to an array declaration?
# </div>
my_first_string = "I have a good day!"
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> Do you remember how to access the elements of an array? The procedure is the same for accessing characters of a string. Access the fifth element:
# +
# Write your own code here
# -
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> <b> Investigate </b> the ways in which you can access the last element of a string:
# </div>
# +
# First method
length_str = len(my_first_string)
print("The last element of a string is: " + my_first_string[length_str - 1])
# Second method
print("The last element of a string is: " + my_first_string[-1])
# -
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> <b> Access </b> the second last element of a string:
# </div>
# +
# Write your own code here
# -
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> <b> Inspect </b> the following line:
# </div>
print("All the elements apart from the last character of the string are: " + str(my_first_string[:-1]))
# <div class = "alert alert-block alert-success">
# In this case, only the <b> last </b> element of the string is not printed out.
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Investigate </b> the following code lines below and <b> comment </b> with your peers what happens in each case:
# </div>
for i in range(len(my_first_string)):
print(my_first_string[i], end = '*')
for i in range(len(my_first_string)):
if(i % 2 == 0):
print(my_first_string[i], end = '')
# <div class = "alert alert-block alert-success">
# The <b> spacebar </b> is also counted as a character in the <b> string</b>.
# ### Optional material
# <div class = "alert alert-block alert-warning">
# <b> Let's play a game: </b> We use all the above tricks for creating the game below. It is not required for you to know by heart how to do the exercises below...You just got started with all these notions, and you need practice to get more used to it...Hopefully, however, by the end of the year, you will get more acquainted with the little game below.
# </div>
# +
ok = 1
A = input("Introduce your initial word here: ")
B = []
while(ok == 1):
B = A
A = input("Introduce the word here: ")
if(A[0] != B[len(B) - 1]):
ok = 0
print("Game over! Try again!")
# -
# Let us make it a bit harder, shall we? Your next word should have its first <b> two </b> characters the same as the last one. Are you up to it?
# +
ok = 1
A = input("Introduce your initial word here: ")
B = []
while(ok == 1):
B = A
A = input("Introduce the word here: ")
if( (A[0] != B[len(B) - 2]) and (A[1] != B[len(B) - 1]) ):
ok = 0
print("Game over! Try again!")
# -
# ### Take-away
# <div class = "alert alert-block alert-success">
# This is it for today, and well done for managing to go through the material!! <br> <br> After this session, you should be more familiar with how simple sentences, numbers and conditional statements can be printed in Python. Moreover, ponder a bit on the for instruction, as it is heavily used in programming. Also, feel free to work more on this notebook using any commands you would like. <br> <br>
# <b> Note: </b> Always keep a back-up of the notebook, in case the original one is altered. <br><br>
# For today's session, this should be enough! See you later!!
# </div>
print("Bye bye! :D")
# 
| GeneralExemplars/Coding Activities for Schools/Advanced Higher/while_arrays_AdvHigher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="G40l9lO2ObLr"
# <center><img src="./images/logo_fmkn.png" width=300 style="display: inline-block;"></center>
#
# ## Домашнее задание №2, теоретическое. Логические методы
# -
# Решите предложенные задачи. Каждая задача должна быть подробно обоснована, задачи без
# обоснования не засчитываются. Решения пишутся в свободной форме, однако так, чтобы проверяющий смог разобраться. Если проверяющий не смог разобраться в решении какой-нибудь задачи, то
# она автоматически не засчитывается. Если используются какие-либо внешние источники, их нужно обязательно указывать.
# + [markdown] id="C86UTDuUlQBk"
# ### Задача 1 (1 балл). Кроссвалидация, LOO, k-fold.
#
# Объясните, стоит ли использовать оценку `leave-one-out-CV` или `k-fold-CV` с небольшим $k$ в случае, когда:
#
# * обучающая выборка содержит очень малое количество объектов;
#
# Стоит использовать `leave-one-out`, так как основной его недостаток --- долгое вычисление на больших выборках.
# * обучающая выборка содержит очень большое количество объектов.
#
# Стоит использовать `k-fold-CV`, так как быстрее всего посчитает.
# -
# **OK**
# + [markdown] id="ZX6sPkky9jS6"
# ### Задача 2 (1 балл). Логистическая регрессия, вывод функции потерь.
#
# Рассмотрим выборку объектов $X = \{x_1,...,x_\ell\}$ и их целевых меток $Y = \{y_1,...,y_\ell\}$, где $y_i ∈ \{0, 1\}$. Предположим, что мы хотим обучить линейный классификатор:
#
# $$Q(w, X^\ell) = \sum\limits_{i=1}^\ell\mathcal{L}(y_i\left<w,x_i\right>) \rightarrow \min\limits_w$$
#
# где $w$ – веса линейной модели, $\mathcal{L}(y, z)$ – некоторая гладкая функция потерь.
#
# Так как решается задача двухклассовой классификации, то будем обучать классификатор предсказывать вероятности принадлежности объекта классу $1$, то есть решать задачу логистической регрессии. Для измерения качества такого классификатора обычно используют правдоподобие $P(Y|X)$ целевых меток $Y$ при заданных объектах $X$ в соответствии с предсказанными распределениями $p$ — чем выше правдоподобие, тем точнее классификатор. Для удобства с вычислительной точки зрения обычно используется отрицательный логарифм правдоподобия, также называемый LogLoss (Logarithmic Loss). Будем считать, что пары объект-ответ $(x_i,y_i)$ независимы между собой для разных $i$.
#
# + [markdown] id="_UX9noDn-jcv"
# #### 1. (0.5 балла)
# Покажите что:
#
# $$\text{LogLoss} = -\text{LogLikelihood} = - \log(P(Y|X)) = - \sum\limits_{i=1}^\ell (y_i \log \tilde y_i + (1-y_i) \log (1-\tilde y_i))$$
#
# ##### Решение:
# -
#
#
#
# \begin{align}P(Y|X) &= \prod_{i = 1}^n P(y_i|X) = \prod_{i = 1}^n \begin{cases} \tilde{y}_i, & y_i = 1\\ 1 - \tilde{y}_i, & y_i = 0 \end{cases}=\\
# &=\prod_{i=1}^n \widetilde y_i^{y_i} (1-\widetilde y_i)^{1-y_i}.
# \end{align}
# После взятия логарифма равенство становится очевидным.
# **OK**
# + [markdown] id="sAZJvhZVA5-7"
# #### 2. (0.5 балла)
# Для того, чтобы классификатор возвращал числа из отрезка $[0,1]$, положите
# $$
# p(y_i=1|x_i) = \sigma \left ( \left< w, x_i \right> \right) =
# \frac{1}{1 + \exp{\left(- \left< w, x_i \right> \right)}};
# $$
#
# сигмоидная функция монотонно возрастает, поэтому чем больше скалярное призведение, тем большая вероятность положительного класса будет предсказана объекту.
#
# Подставьте трансформированный ответ линейной модели в логарифм правдоподобия. К какой функции потерь мы пришли? (Обратите внимание, что функция обычно записывается для классов $\{-1, 1\}$).
# -
# **Выходил к доске**, ответ
#
# $$\text{LogLoss}=-\sum \log(1+\exp(y_i\langle w, x\rangle)$$
# + [markdown] id="lG4d_zxADKDF"
# ### Задача 3 (1.5 балла). Логистическая регрессия, решение оптимизационной задачи.
# + [markdown] id="lA5GvcRcDY52"
# #### 1. (0.8 балла)
#
# Докажите, что в случае линейно разделимой выборки не существует вектора параметров (весов), который бы максимизировал правдоподобие вероятностной модели логистической регрессии в задаче двухклассовой классификации.
# -
# ##### Решение.
#
# Если $w$ максимум правдоподобия, то для любого $ i $
# $$P(y_i|X)>0.5.$$
# Вероятность у нас вычисляется по формуле
# $$P(y_i|X)=\sigma(y_i \langle w, x_i\rangle).$$
# Раз вероятность больше 0.5, то аргумент у сигмы положительный. Так что домножая вектор весов на число, большее 1, аргумент станет больше для любого $ i $, увеличивая сигму, а значит и правдоподобие.
# + [markdown] id="3VBN5jiVE32G"
# #### 2. (0.4 балла)
#
# Предложите, как можно модифицировать вероятностную модель, чтобы оптимум достигался.
# -
# **Решение:** добавим условие: вектор весов по модулю равен 1. Тогда если $w$ --- вектор ортогональный разделяющей плоскости, то существует единственный коэффициент $ с > 0$ такой, что $cw$ можно использовать как вектор весов, так что оптимум будет достигаться в нем.
# + [markdown] id="7C4UiKK3HV94"
# #### 3. (0.3 балла)
# Выпишите формулы пересчета значений параметров при оптимизации методом градиентного спуска для обычной модели логистической регрессии и предложенной модификации.
#
# Для 1:
#
# $$w\mapsto w + \frac 1t \sum_{i=1}^l y_i\sigma(-y_i\langle w, x_i\rangle) \cdot x_i.$$
#
# Для 2:
#
#
# $$ w\mapsto \frac{w + \frac 1t 2\sum_{i=1}^l y_i\sigma(a + \langle w, x_i\rangle) \cdot x_i}{||w + \frac 1t 2\sum_{i=1}^l y_i\sigma(a + \langle w, x_i\rangle) \cdot x_i||}.$$
# + [markdown] id="NtrnMSYXKHRS"
# ### Задача 4 (1 балл). Мультиномиальная регрессия
#
# В случае многоклассовой классификации логистическую регрессию можно обобщить: пусть для каждого класса $k$ есть свой вектор весов $w_k$. Тогда вероятность принадлежности классу $k$ запишем следующим образом:
#
# $$
# P(y=k | x, W) = \frac{e^{\langle w_k, x\rangle}}{\sum\limits_{j=1}^K e^{\langle w_j, x\rangle}}
# $$
#
# Тогда оптимизируемая функция примет вид:
#
# $$
# \mathcal{L}_{sm}(W) = -\sum_{i=1}^N \sum_{k=1}^K [y_i=k]\ln P(y_i=k | x_i, W),~\text{где}~[y_i=k]=\begin{cases}
# 1, y_i=k,\\
# 0, \text{иначе}
# \end{cases}
# $$
#
# Пусть количество классов $K=2$. Для простоты положим, что выборка линейно неразделима.
# + [markdown] id="p_Rd-dJWKm8r"
# #### 1. (0.5 балла)
# Единственно ли решение задачи? Почему?
#
# Если $w_1,w_2$ --- решения, то $w_1+w,w_2+w$ так же решения для любого вектора $w$, ибо $P(y-k|x,W)$ не поменяется, ибо числитель и каждое слагаемое в знаменателе домножатся на $ e^{\langle w, x \rangle} $.
# + [markdown] id="TXe4eM0JL0xL"
# #### 2. (0.5 балла)
# Покажите, что предсказанные распределения вероятностей на классах в случае логистической и мультиномиальной регрессий будут совпадать.
# -
# Очевидно из определения правдоподобия, что $P(y=k|x, w_1, w_2) = P(y=k|x, 0, w_2-w_1)$.
#
# Следовательно, $ \mathcal{L}_{sm}(w_1,w_2) = \mathcal{L}_{sm}(0,w_2 - w_1)$.
# Более того, несложно видеть, что
# $$ \mathcal{L}_{sm}(0,w_2 - w_1) = LogLoss(w_2-w_1) $$, так что на самом деле можно считать, что в мультиномиальной регрессии один параметр, в котором значение правдоподобия совпадает с правдоподобией логистической регрессии, иными словами оптимизируется одна и та же функция.
# **OK**
# + [markdown] id="W2a3lm4KMFGl"
# ### Задача 5 (1.5 балла) Решающие деревья, константное предсказание, функции потерь.
#
# Допустим, при построении решающего дерева в некоторый лист попало $N$ объектов $x_1, ... , x_N$ с метками $y_1, ... , y_N$.
# Предсказание в каждом листе дерева - константа. Найдите, какое значение $\tilde y$
# должен предсказывать этот лист для минимизации следующих функций потерь:
# + [markdown] id="pBcO-ZNpMSYY"
# #### 1. (0.5 балла)
# Mean Squared Error (средний квадрат ошибки) для задачи регрессии:
# \begin{equation}Q=\frac{1}{N}\sum_{i=1}^N (y_i - \tilde y)^2;\end{equation}
#
# $\tilde y = \frac 1N \sum_{i=1}^N y_i$, так как известно, что для любой случайной величины $ X $ минимальное значение функции $\mathbb{E}(X-t)^2$ достигается в точке $t = \mathbb{E}X$ и значение является дисперсией.
#
# **ОК**
# + [markdown] id="xrKaxpXmNaMf"
# #### 2. (0.5 балла)
# Mean Absolute Error (средний модуль отклонения) для задачи регрессии:
# \begin{equation}Q=\frac{1}{N}\sum_{i=1}^N |y_i - \tilde y|.\end{equation}
# -
# Рассмотрим $y_k \leq \tilde y = y_k + x \leq y_{k+1}$. Тогда
# $$ Q(\tilde y) = Q(y_k) + xk - x(N-k) = Q(y_k) + x(2k-N). $$
# Ошибка убывает с ростом $x$ при $k<\frac N2$ и растет при больших $k$. Отмечу, что если $N$ четно, то при $k=N/2$ величина $x$ не влияет на ошибку. Таким образом $\tilde y$ это просто напросто медиана.
# **ОК**
# + [markdown] id="TnKGc3keOgF4"
# #### 3. (0.5 балла)
# $\text{LogLoss}$ (логарифмические потери) для задачи классификации:
# $$Q=-\frac{1}{N}\sum_{i=1}^N \left(y_i\log\tilde y+(1-y_i)\log(1-\tilde y)\right),
# \quad \tilde y\in[0,1], \quad y_i \in \{0,1\}.$$
# -
# $$0=Q' = -\frac 1N \sum_{i=1}^n \frac{y_i}{\tilde y} - \frac{1-y_i}{1-\tilde y}=-\sum_{i=1}^n\frac{\bar y - \tilde y}{\tilde y (1-\tilde y)},$$
# где $ \bar y = \frac 1N \sum_{i=1}^n y_i. $ Тогда ясно, что минимум достигается в выборочном среднем.
# **ОК**
# + [markdown] id="cSmKQAILP0T0"
# ### Задача 6 (1 балл). Решающие деревья, функции потерь, impurity functions.
# $$
# \Phi(U) - \frac{|U_1|}{|U|}\Phi(U_1) - \frac{|U_2|}{|U|}\Phi(U_2) \to \max
# $$
# таким выражением в лекции задается критерий, по которому происходит ветвление вершины решающего дерева. Давайте разберемся подробнее.
#
# Impurity function $\Phi(U)$ («функция неопределенности» или «функция нечистоты») используется для того, чтобы измерить степень неоднородности целевых меток $y_1,\dots, y_\ell$ для множества объектов $U$ размера $\ell$. Например, при обучении решающего дерева в текущем листе выбирается такое разбиение множества объектов $U$ на два непересекающихся множества $U_1$ и $U_2$, чтобы impurity function $\Phi(U)$ исходного множества $U$ как можно сильнее превосходила нормированную impurity function в новых листьях $\frac{|U_1|}{|U|}\Phi(U_1) + \frac{|U_2|}{|U|}\Phi(U_2)$. Отсюда и получается, что нужно выбрать разбиение, решающее задачу
# $$
# \Phi(U) - \frac{|U_1|}{|U|}\Phi(U_1) - \frac{|U_2|}{|U|}\Phi(U_2) \to \max.
# $$
# Полученную разность называют Gain (выигрыш), и она показывает, на сколько удалось уменьшить «неопределенность» от разбиения листа два новых.
#
# В соответствии с одним из возможных определений, impurity function — это значение функционала ошибки $Q = \frac{1}{\ell}\sum\limits_{i=1}^\ell \mathcal{L}(y_i, \tilde{y})$ в листе с множеством объектов $U$ при константном предсказании $\tilde{y}$, оптимальном для $Q$ (см. задачу 7):
# $$
# \Phi(U) = \frac{1}{\ell}\sum\limits_{i=1}^\ell \mathcal L (y_i, \tilde y).
# $$
# Понятно, что каждому критерию разбиения соответствует своя impurity function $\Phi(U)$, а в основе каждой $\Phi(U)$ лежит некоторая функция потерь. Давайте разберемся, откуда берутся различные критерии разбиения.
# + [markdown] id="7ZNypaDVQDYZ"
# #### 1. (0.5 балла)
# Покажите, что для квадратичных потерь $\mathcal L (y_i, \tilde y) = (y_i - \tilde y)^2$ в задаче регрессии $y_i \in \mathbb{R}$ impurity function $\Phi(U)$ равна выборочной дисперсии целевых меток объектов, попавших в лист дерева.
# -
# Как было показано в 5.1, при квадратичных потерях $\mathcal{L}(y_i,i)=(y_i-y)^2$ $\tilde y = \bar y$ --- выборочному среднему. Таким образом,
# $$ \Phi(U) = \frac 1l \sum_{i=1}^l \mathcal L (y_i,\tilde y) = \frac 1l \sum_{i=1}^l (y_i-\bar y)^2 = S(y),$$
# она же выборочная дисперсия.
#
# **ОК**
# + [markdown] id="MeWp8iIxR96A"
# #### 2. (0.5 балла)
# Покажите, что для функции потерь $\text{Logloss}$ $\mathcal L (y_i, \tilde y) =-y_i\log(\tilde y) - (1-y_i)\log(1 - \tilde y)$ в задаче классификации $y_i \in \{0,1\}$ impurity function $\Phi(U)$ соответствует энтропийному критерию разбиения.
# -
# Энтропийный критерий разбиения
# $$\Phi(U) = -q\log_2 q - (1-q) \log_2 (1-q),$$
# где $q = P(y=1|U)$, иными словами доля 1 среди значений $ y $ при данных значениях $U$. Однако $ \bar y $ так же является таковым, ибо это количество значений 1 деленное на количество значений. Осталось заметить, что
# $$\Phi(U) = \text{LogLoss} \mathcal L (y_i, \tilde y) =[\text{см 5.3}]= \mathcal L (y_i, \bar y) = \mathcal L (y_i, q).$$
# **ОК**
# + [markdown] id="KvAw2PMSTrkw"
# ### Задача 7 (1 балл). Решающие деревья, индекс Джини
#
# Пусть имеется построенное решающее дерево для задачи многоклассовой классификации. Рассмотрим лист дерева с номером $m$ и объекты $R_m$, попавшие в него. Обозначим за $p_{mk}$ долю объектов $k$-го класса в листе $m$. *Индексом Джини* этого листа называется величина
# $$\sum_{k = 1}^{K} p_{mk} (1 - p_{mk}),$$
# где $K$ — общее количество классов. Индекс Джини обычно служит мерой того, насколько хорошо в данном листе выделен какой-то один класс (см. impurity function в предыдущей задаче).
# + [markdown] id="39o2PqSqT9U-"
# #### 1. (0.5 балла)
# Поставим в соответствие листу $m$ алгоритм классификации $a(x)$, который предсказывает класс случайно, причем класс $k$ выбирается с вероятностью $p_{mk}$. Покажите, что матожидание частоты ошибок этого алгоритма на объектах из $R_m$ равно индексу Джини.
# -
# Частота ошибок --- это вероятность не угадать класс, которая вычисляется следующим образом:
# $$\sum_{i=1}^K P(y_{predicted} = i, y_{actual}\ne i) = \sum_{i=1}^K p_{mk} \cdot (1-p_{mk}),$$
# где события в левой части несовместны, оттого и сумма ($y_{predicted}$ не может быть одновременно 1 и 2, например), а события внутри P для любого i независимы, ибо наше предсказание не зависит от реального значения(иначе не понятно что такое случайное предсказание), следовательно вероятности перемножаются.
# + [markdown] id="_2_WiDXWU5Mu"
# #### 2. (0.5 балла)
# *Дисперсией класса $k$* назовем дисперсию выборки $\{ [y_i = k]:\ x_i \in R_m$\},
# где $y_i$ - класс объекта $x_i$, $[f]$ — индикатор истинности выражения $f$, равный 1 если $f$ верно, и нулю в противном случае, а $R_m$ — множество объектов в листе.
# Покажите, что сумма дисперсий всех классов в заданном листе равна его индексу Джини.
# -
# Дисперсия класса $k$ является дисперсией Бернуллевской выборки $\{ [y_i = k]:\ x_i \in R_m\}$, где доля 1 равна $p_{mk}$. Известный факт, что дисперсия у Бернуллевской величины равна $p_{mk}(1-p_{mk})$. Далее берется сумма всех дисперсий, что натурально является индексом Джини.
# **Ок**
# + [markdown] id="-6-ynDEVYCF4"
# ### Задача 8 (2 балла). Бинарные решающие деревья, MSE
#
# Предложите алгоритм построения *оптимального* бинарного решающего дерева для задачи регрессии на $\ell$ объектах в $n$-мерном пространстве с асимптотической сложностью $O(n \ell \log \ell)$. В качестве предикатов нужно рассматривать пороговые правила (наиболее распространенный случай на практике). Для простоты можно считать, что получающееся дерево близко к сбалансированному (т.е. его глубина имеет порядок $O(\log \ell)$) и в качестве функции ошибки используется Mean Squared Error (MSE):
# $$Q=\frac{1}{\ell}\sum_{i=1}^\ell (y_i - \tilde y_i)^2$$
# Под оптимальностью в данной задаче подразумевается, что в каждом узле дерева делается оптимальное с точки зрения MSE разбиение на два поддерева.
#
# -
# На каждом уровне узлы имеют суммарный размер $\leq l$, так что если каждый узел будет $v$ будет обрабатываться за $O(|v|n)$ действий, то алгоритм будет работать за $O(depth \cdot nl )=O( l\log nl)$, что и требуется. Алгоритм будет следующий:
# * Перебираем признаки.
# * В каждом перебираем пороги.
# * Заранее считаем суммы на префиксах и суффиксах (если их делить на индекс префикса +1, то получится $\tilde y_i$ при сплите на соответствующем пороге).
# * Для первого порога честно считаем изменение MSE.
# * Для каждого следующего используя массив префиксных и суффиксных сумм можно за O(1) посчитать изменение MSE:
# $$ Q = \frac 1{l+1}\sum_{i=1}^{l+1}(y_i-\tilde y_i)^2 =[\delta = \tilde y_i - \tilde y_{i,prev}]= \frac 1{l+1}\sum_{i=1}^{l+1}(y_i-\tilde y_{i,prev} - \delta)^2 = $$
# $$ =\frac l{l+1}Q_{prev} -\frac 1{l+1}\sum_{i=1}^l 2\delta (y_i-\tilde y_{i,prev}) - \delta^2+\frac 1{l+1}(y_{l+1}-\tilde y_{l+1})^2.$$
# Учитывая предподсчет префиксов и суффиксов, за $|v|$ смогли найти наилучший сплит по признаку $k$ для любого $k=1,\dots,n$
| hws/HW2/HW2_logic_methods_theory_Sviridov_Oleg_4bSwMOr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import scipy.io
from scipy import interpolate
import numpy as np
def d3_scale(dat, out_range=(-1, 1), in_range=None):
if in_range == None:
domain = [np.min(dat, axis=0), np.max(dat, axis=0)]
else:
domain = in_range
def interp(x):
return out_range[0] * (1.0 - x) + out_range[1] * x
def uninterp(x):
b = 0
if (domain[1] - domain[0]) != 0:
b = domain[1] - domain[0]
else:
b = 1.0 / domain[1]
return (x - domain[0]) / b
return interp(uninterp(dat))
# load the patterns from matlab
pattern_file_names = [
"nnRawExaStride",
"nnRawSlowWalk",
"nnRawWalk",
"nnRawRunJog",
"nnRawCartWheel",
"nnRawWaltz",
"nnRawCrawl",
"nnRawStandup",
"nnRawGetdown",
"nnRawSitting",
"nnRawGetSeated",
"nnRawStandupFromStool",
"nnRawBox1",
"nnRawBox2",
"nnRawBox3",
]
output_dims = 61
pattern_num = 2
pattern_file_names = pattern_file_names[:2]
# -
min_maxs = np.zeros((output_dims, 2))
min_maxs[0,:].shape
# +
min_maxs = np.zeros((output_dims, 2))
min_maxs[:, 0] = np.inf
min_maxs[:, 1] = -np.inf
raw_dats = []
# get the actual maximum and minimums for each dimension
for nm in pattern_file_names:
name = nm[5:]
raw_dats.append(scipy.io.loadmat("section2.3_demoMotionCapture/nnData/%s.mat" %(nm))["nnRawData"+name].T)
for o_i in range(output_dims):
assert raw_dats[-1][o_i].shape != (61,)
min_val = np.min(raw_dats[-1][o_i])
if min_val < min_maxs[o_i, 0]:
min_maxs[o_i, 0] = min_val
max_val = np.max(raw_dats[-1][o_i])
if max_val > min_maxs[o_i, 1]:
min_maxs[o_i, 1] = max_val
# -
assert np.all(min_maxs[:, 0] != np.inf)
assert np.all(min_maxs[:, 1] != -np.inf)
# +
function_list = []
for n_i, nm in enumerate(pattern_file_names):
# make each pattern values normalised between -1, 1
# and temporally squash them between -1 and 1 too
function_list.append([])
raw_dat = raw_dats[n_i]
xv = np.linspace(-1, 1, raw_dat.shape[1])
assert raw_dat.shape[0] == output_dims
normed_data = np.zeros_like(raw_dat)
for o_i in range(output_dims):
assert min_maxs[o_i][0] <= np.min(raw_dat[o_i, :])
assert min_maxs[o_i][1] >= np.max(raw_dat[o_i, :])
normed_data[o_i, :] = d3_scale(raw_dat[o_i, :], in_range=min_maxs[o_i])
assert np.max(normed_data) <= 1.0
assert np.min(normed_data) >= -1.0
function_list[-1].append(interpolate.interp1d(xv, normed_data[o_i, :]))
# -
import matplotlib.pyplot as plt
# %matplotlib inline
print(raw_dats[0][0].shape)
print(min_maxs[0])
plt.plot(raw_dats[0][0])
#plt.plot(raw_dats[1][0])
plt.savefig("good")
# +
# can approx? yes!
raw_dat = raw_dats[0][0]
x_max = raw_dat.shape[0]
xv = np.linspace(0, x_max, x_max)
plt.scatter(xv, raw_dat)
x_new = np.linspace(-np.pi, np.pi, x_max)
f_approx = interpolate.interp1d(x_new, d3_scale(raw_dat))
plt.plot(d3_scale(f_approx(x_new), in_range=(-1, 1), out_range=min_maxs[0]))
# +
xv = np.linspace(-1, 1, raw_dat.shape[0])
plt.plot(function_list[0][0](xv))
# -
af = scipy.io.loadmat("section2.3_demoMotionCapture/nnData/%s.mat" %(pattern_file_names[0]))
tmp = af['nnRawDataExaStride'].T
tmp[0].shape
pat_out = scipy.io.loadmat("pattern_out.mat")
reg_out = pat_out["reg_out"]
#ideal_out = pat_out["ideal_out"]
compr = pat_out["compressed"]
#print(ideal_out.T.shape)
print(np.min(reg_out))
print(np.max(reg_out))
print(reg_out.T.shape)
print(np.min(reg_out))
print(np.max(reg_out))
# +
import nengo
plt_val = reg_out[:, 0][975:]
#plt.plot(plt_val)
plt.plot(nengo.Lowpass(0.01).filt(plt_val, dt=0.001))
# -
plt.plot(ideal_out[0])
plt.plot(compr[0])
plt.plot(compr[0][::-1])
plt.plot(ideal_out[0][::-1][9:-10])
plt.plot(raw_dats[0][0])
plt.plot(compr[1]*10-0.1)
plt.plot(d3_scale(compr[1], in_range=(np.min(compr[1]), np.max(compr[1])), out_range=(np.min(raw_dats[0][1]), np.max(raw_dats[0][1]))))
plt.plot(raw_dats[0][1][::-1])
plt.plot(compr[2]-1)
plt.plot(raw_dats[0][2])
plt.plot(compr[3]-1)
plt.plot(raw_dats[0][3])
# it's basically zero, so whatevs
plt.plot(compr[4]-1)
#plt.plot(raw_dats[0][4])
plt.plot(compr[5]-1)
plt.plot(raw_dats[0][5])
fin_out = scipy.io.loadmat("final_pattern.mat")["final_out"]
plt.plot(fin_out[0][:315][::-1])
plt.plot(raw_dats[0][0])
# This looks important for comparison:
#http://docs.scipy.org/doc/scipy-0.16.1/reference/generated/scipy.signal.coherence.html
# +
from scipy import signal
f, Cx = signal.coherence(fin_out[0][:315][::-1], raw_dats[0][0], fs=10e3)
# -
plt.plot(f, Cx)
# check that shifting does what you expect
corr = signal.correlate(fin_out[0][200:515][::-1], raw_dats[0][0])
plt.plot(corr)
# +
from numpy.linalg import norm
all_max = np.max([np.max(fin_out[0][:315][::-1]), np.max(raw_dats[0][0])])
all_min = np.min([np.min(fin_out[0][:315][::-1]), np.min(raw_dats[0][0])])
corr = signal.correlate(
d3_scale(fin_out[0][:315][::-1], in_range=(all_min, all_max)),
d3_scale(raw_dats[0][0], in_range=(all_min, all_max))
)
plt.plot(corr)
# -
plt.plot(d3_scale(fin_out[0][:315][::-1], in_range=(all_min, all_max)))
plt.plot(d3_scale(raw_dats[0][0], in_range=(all_min, all_max)))
# +
ps1 = np.abs(np.fft.fft(fin_out[0][:315][::-1]))**2
ps2 = np.abs(np.fft.fft(raw_dats[0][0]))**2
# try shifting and scaling
plt.plot(ps1[1:10])
plt.plot(ps2[1:10])
print(ps1[0])
print(ps2[0])
| func_test-python2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Imports
from PW_explorer.load_worlds import load_worlds
from PW_explorer.run_clingo import run_clingo
from DLV_Input_Parser.dlv_rules_parser import parse_dlv_rules
# %load_ext PWE_NB_Extension
# ### Implementation
# We know that the symmetry degree is defined as the number of automorphism groups of the incidence graph of a query. Below we define an encoding that finds all the automorphisms of a given graph.
# ##### Automorphism Encoding
# +
# %%clingo --donot-display_input -lci automorphisms --donot-run
% e(X,Y) :- e(Y,X). --> only if undirected
gnode(X) :- e(X,_).
gnode(X) :- e(_,X).
vmap(X,Y) ; vout(X,Y) :- gnode(X), gnode(Y).
:- vmap(X1,Y1), vmap(X2,Y2), e(X1,X2), not e(Y1,Y2).
:- vmap(X1,Y1), vmap(X2,Y2), not e(X1,X2), e(Y1,Y2).
% used1(X) :- vmap(X,_).
% :- gnode(X), not used1(X).
% :- vmap(X,Y),vmap(X,Z),Y!=Z.
% :- vmap(Y,X),vmap(Z,X),Y!=Z.
:- gnode(X), #count {Y: vmap(X,Y)} != 1.
:- gnode(X), #count {Y: vmap(Y,X)} != 1.
% #show vmap/2.
#show.
# -
symm_degree_rules = str(automorphisms).split('\n')
# We can construct a incidence graph of query Q using the function below.
def get_incidence_graph_edge_facts(rule):
listener = parse_dlv_rules(rule, print_parse_tree=False)
edges = []
for rule in listener.rules:
head_atoms, tail_atoms = rule[0], rule[1]
atom_count = 0
for head in head_atoms+tail_atoms:
atom_node = '"{}_{}_{}"'.format(head.rel_name, head.rel_arity, atom_count)
edges.extend([('"{}"'.format(v), atom_node) for v in head.vars])
atom_count += 1
edge_facts = []
for e in edges:
edge_facts.append("e({},{}).".format(*e))
return edge_facts
# We now combine these two to find the symmetry degree.
def get_symm_degree(rule):
edge_facts = get_incidence_graph_edge_facts(rule)
# print(edge_facts)
asp_out, _ = run_clingo(symm_degree_rules+edge_facts)
_, _, pw_objs = load_worlds(asp_out, silent=True)
return len(pw_objs)
# #### Testing
# tri/0
get_symm_degree('tri :- e(X,Y), e(Y,Z), e(Z,X).')
# tri/1
get_symm_degree('tri(X) :- e(X,Y), e(Y,Z), e(Z,X).')
# tri/2
get_symm_degree('tri(X,Y) :- e(X,Y), e(Y,Z), e(Z,X).')
# tri/3
get_symm_degree('tri(X,Y,Z) :- e(X,Y), e(Y,Z), e(Z,X).')
# thop/0
get_symm_degree('thop :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# thop/1
get_symm_degree('thop(X) :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# thop/2
get_symm_degree('thop(X,Y) :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# thop/3
get_symm_degree('thop(X,Y,Z1) :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
# thop/4
get_symm_degree('thop(X,Y,Z1,Z2) :- hop(X,Z1), hop(Z1,Z2), hop(Z2,Y).')
| Symmetry-Degree-of-a-Query.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="t5DCDUTabzzY"
# ISB-CGC Community Notebooks
# Check out more notebooks at our [Community Notebooks Repository](https://github.com/isb-cgc/Community-Notebooks)!
#
# -
# Title: How to use Kallisto to quantify genes in 10X scRNA-seq
#
# Author: <NAME>
#
# Created: 2019-08-07
#
# Purpose: Demonstrate how to use 10X fastq files and produce the gene quantification matrix
#
# Notes:
# In this notebook, we're going to use the 10X genomics fastq files that we generated earlier, to quantify gene expression per cell using Kallisto and Bustools.
# It is assumed that this notebook is running INSIDE THE CLOUD! By starting up a Jupyter notebook, you are already authenticated, can read and write to cloud storage (buckets) for free, and data transfers are super fast. To start up a notebook, log into your Google Cloud Console, use the main 'hamburger' menu to find the 'AI platform' near the bottom. Select Notebooks and you'll have an interface to start either an R or Python notebook.
# ## Resources:
# Bustools paper:
# https://www.ncbi.nlm.nih.gov/pubmed/31073610
# https://www.kallistobus.tools/getting_started_explained.html
# https://github.com/BUStools/BUS_notebooks_python/blob/master/dataset-notebooks/10x_hgmm_6k_v2chem_python/10x_hgmm_6k_v2chem.ipynb
# https://pachterlab.github.io/kallisto/starting
# cd /home/jupyter/
# + [markdown] colab_type="text" id="E7qiNoHdb9vh"
#
#
# + [markdown] colab_type="text" id="pJ2wnujWwlFb"
# ## Software install
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="w5UxLaCSwm9_" outputId="1ccaa519-e295-4f61-999a-fef686950c94"
# !git clone https://github.com/pachterlab/kallisto.git
# -
# cd kallisto/
# ls -lha
# + colab={"base_uri": "https://localhost:8080/", "height": 850} colab_type="code" id="1MAgAT2Kw7GS" outputId="e877a752-aea9-44bd-e9b5-cb7bd927c0cd"
# !sudo apt --yes install autoconf cmake
# + colab={} colab_type="code" id="5qUPqfMrxbih"
# !mkdir build
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="nN3bP7X2xtFC" outputId="2acf0f2d-99a7-4db9-c5d4-c14ac07bf402"
# cd build
# + colab={"base_uri": "https://localhost:8080/", "height": 2009} colab_type="code" id="9zMImaArxA_h" outputId="9a2843a9-0862-4697-9396-e951d4ce3df3"
# !sudo cmake ..
# !sudo make
# !sudo make install
# + colab={"base_uri": "https://localhost:8080/", "height": 323} colab_type="code" id="kZRc9jLTxsig" outputId="7fe3990b-6c91-4df5-a0b9-98c5077b759b"
# !kallisto
# -
# cd ../..
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="hDAdMXUoymbG" outputId="e8347ef1-01d4-421d-8af7-2f70409666ca"
# !git clone https://github.com/BUStools/bustools.git
# -
# we need the devel version due to a bug that stopped compilation ...
# !git checkout devel
# !git status
# cd bustools/
# + colab={} colab_type="code" id="ZyTY539G0Pf3"
# !mkdir build
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="pqRkGsdA1afB" outputId="519845e6-45db-4fdd-8273-7026be71acb2"
# cd build
# + colab={"base_uri": "https://localhost:8080/", "height": 765} colab_type="code" id="_95UWvCb0UFK" outputId="25594773-fd95-4b27-bf95-5dce80db3a3b"
# !sudo cmake ..
# !sudo make
# !sudo make install
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="BjD5mz5B0pxu" outputId="c26a4c88-480a-4b4f-9d9c-8a348a837bbc"
# cd ../..
# + colab={"base_uri": "https://localhost:8080/", "height": 255} colab_type="code" id="Dwhu4Z5c0k1k" outputId="3c05bf3d-11a5-4523-bdf7-013036470a4a"
# !bustools
# + colab_type="text" id="Q5qi1JRAuobI"
# + [markdown] colab_type="text" id="tU_SHwE5utez"
# ## Reference Gathering
# + colab={} colab_type="code" id="qchHsWTnsrVM"
# mkdir kallisto_bustools_getting_started/; cd kallisto_bustools_getting_started/
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="zypqbJ8JsraW" outputId="73957f0d-cf91-45e7-f6b4-3df790455a15"
# !wget ftp://ftp.ensembl.org/pub/release-96/fasta/homo_sapiens/cdna/Homo_sapiens.GRCh38.cdna.all.fa.gz
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="Zno4t5dUsrda" outputId="ab9f1835-7563-41ad-fa6f-09a50acf3ecb"
# !wget ftp://ftp.ensembl.org/pub/release-96/gtf/homo_sapiens/Homo_sapiens.GRCh38.96.gtf.gz
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="i6ogJW9isrgM" outputId="c29204e8-5254-47de-f186-5edf4e6862db"
# + [markdown] colab_type="text" id="Y_0E9g3Du1k5"
# ## Barcode whitelist
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="RSp31-dSuiCu" outputId="4a908e89-4819-40d9-f645-59570f7401ae"
# Version 3 chemistry
# !wget https://github.com/BUStools/getting_started/releases/download/species_mixing/10xv3_whitelist.txt
# -
# Version 2 chemistry
# !wget https://github.com/bustools/getting_started/releases/download/getting_started/10xv2_whitelist.txt
# + [markdown] colab_type="text" id="5Cd9ggS9u-au"
# ## Gene map utility
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="I1AwPwzbv3xP" outputId="07d6eaf6-0ba4-449d-e6db-f7a858164de4"
# !wget https://raw.githubusercontent.com/BUStools/BUS_notebooks_python/master/utils/transcript2gene.py
# -
# !gunzip Homo_sapiens.GRCh38.96.gtf.gz
# + colab={} colab_type="code" id="9F9t9InPuiO5"
# !python transcript2gene.py --use_version < Homo_sapiens.GRCh38.96.gtf > transcripts_to_genes.txt
# -
# !head transcripts_to_genes.txt
# + [markdown] colab_type="text" id="JRrZ9P-k1397"
# ## Data
# -
# mkdir data
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="h2LG71Vl2olI" outputId="64e6d28a-81cf-48a5-94ed-9f1bdcd9e404"
# !gsutil -m cp gs://your-bucket/bamtofastq_S1_* data
# -
# mkdir output
# cd /home/jupyter
# ls -lha data
# + [markdown] colab_type="text" id="krdOGVfL3qVU"
# ## Indexing
# + colab={"base_uri": "https://localhost:8080/", "height": 255} colab_type="code" id="7uwhAtw33hlL" outputId="3b012501-f71f-4127-be94-a6dd0a99c5c4"
# !kallisto index -i Homo_sapiens.GRCh38.cdna.all.idx -k 31 Homo_sapiens.GRCh38.cdna.all.fa.gz
# + [markdown] colab_type="text" id="yHqoOdsB6D5H"
# ## Kallisto
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="APhJDd4e6CFH" outputId="3f2ad505-4d5e-4c16-b89f-04c17e66bdb6"
# !kallisto bus -i Homo_sapiens.GRCh38.cdna.all.idx -o output -x 10xv3 -t 8 \
# data/bamtofastq_S1_L005_R1_001.fastq.gz data/bamtofastq_S1_L005_R2_001.fastq.gz \
# data/bamtofastq_S1_L005_R1_002.fastq.gz data/bamtofastq_S1_L005_R2_002.fastq.gz \
# data/bamtofastq_S1_L005_R1_003.fastq.gz data/bamtofastq_S1_L005_R2_003.fastq.gz \
# data/bamtofastq_S1_L005_R1_004.fastq.gz data/bamtofastq_S1_L005_R2_004.fastq.gz \
# data/bamtofastq_S1_L005_R1_005.fastq.gz data/bamtofastq_S1_L005_R2_005.fastq.gz \
# data/bamtofastq_S1_L005_R1_006.fastq.gz data/bamtofastq_S1_L005_R2_006.fastq.gz \
# data/bamtofastq_S1_L005_R1_007.fastq.gz data/bamtofastq_S1_L005_R2_007.fastq.gz
# -
# ## Bustools
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="wMj9XJy6CwSk" outputId="09a57b2c-d93a-439f-df83-f9e3e8bec0da"
# cd /home/jupyter/output/
# + colab={} colab_type="code" id="6xRptPRHDHRw"
# !mkdir genecount;
# !mkdir tmp;
# !mkdir eqcount
# -
# !bustools correct -w ../10xv3_whitelist.txt -o output.correct.bus output.bus
# !bustools sort -t 8 -o output.correct.sort.bus output.correct.bus
# !bustools text -o output.correct.sort.txt output.correct.sort.bus
# !bustools count -o eqcount/output -g ../transcripts_to_genes.txt -e matrix.ec -t transcripts.txt output.correct.sort.bus
# !bustools count -o genecount/output -g ../transcripts_to_genes.txt -e matrix.ec -t transcripts.txt --genecounts output.correct.sort.bus
# !gzip output.bus
# !gzip output.correct.bus
# ## Copyting out results
# cd /home/jupyter
# !gsutil -m cp -r output gs://my-output-bucket/my-results
| Notebooks/How_to_use_Kallisto_on_scRNAseq_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from functools import reduce
import operator
import math
from cospar import reader, F, colour
from everest.window import Canvas, DataChannel
# %matplotlib inline
# -
conds = (
F('inputs/aspect') == 1,
F('inputs/f') == 1,
F('inputs/tauRef') <= 1e6,
F('inputs/tauRef') >= 1e5,
F('inputs/temperatureField') == '_built_peaskauslu-thoesfthuec',
)
cut = reader[reduce(operator.__and__, conds)]
# datas = reader[cut : ('tauRef', 't', 'Nu')]
# datas = {k : v for k, v in datas.items() if len(v[1])}
datas = sorted(reader[cut : ('inputs/tauRef', 'outputs/t', 'outputs/Nu')].values())
# +
canvas = Canvas(size = (12, 6), colour = 'white', fill = 'black')
ax = canvas.make_ax()
for tau, t, Nu in datas:
ax.line(t, Nu, c = colour(math.log10(tau), 5, 6))
ax.props.edges.x.lims = (0., 0.3)
ax.props.edges.x.label.text = 'Dimensionless time'
ax.props.edges.y.label.text = 'Nusselt number'
ax.props.title.text = 'Nusselt number profiles for varying yield strength'
ax.props.legend.set_handles_labels(
[h for h, *_ in ax.collections],
[str(round(math.log10(tau), 2)) for tau, *_ in datas],
)
ax.props.legend.title.text = 'tauRef (10^n)'
ax.props.legend.kwargs.update(
loc = 'upper right',
ncol = 2,
frameon = True,
bbox_to_anchor = (1.22, 1),
)
canvas.show()
| analysis/cospar/working_020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import plotly.graph_objects as go
import plotly.offline as po
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
import matplotlib.pyplot as plt
import dash
import plotly.express as px
import random
import plotly.figure_factory as ff
# # Loading Datasets
pokemon = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/pokemon_updated.csv")
pokemon.head(10)
stdperf = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/studentp.csv")
stdperf.head(10)
corona = pd.read_csv('C:/Users/DELL/Documents/GitHub/Public/COVID-19/covid/data/countries-aggregated.csv' ,
index_col='Date' , parse_dates=True)
corona.head(10)
spotify = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/spotify.csv" ,
index_col="Date")
spotify.head(10)
housing = pd.read_csv('C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/housing.csv')
housing.tail()
insurance = pd.read_csv('C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/insurance.csv')
insurance.head(10)
employment = pd.read_excel("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/unemployment.xlsx")
employment.head(10)
helpdesk = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/helpdesk.csv")
helpdesk.head(10)
fish= pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/Fish.csv")
fish.head(10)
exercise = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/exercise.csv")
exercise.head(10)
suicide = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/suicide.csv")
suicide.head(10)
canada = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/canada.csv")
canada.head()
canada.columns
canada.drop(columns=['AREA' , 'DEV', 'DevName' , 'REG', 'Type', 'Coverage' , 'AreaName', 'RegName' ], inplace=True)
canada.head()
canada.rename(columns={'OdName':'Country'} , inplace=True)
canada.set_index(canada.Country,inplace=True)
canada.head()
canada2 = canada.copy()
canada2.head()
canada.index.name=None
canada.head()
del canada['Country']
canada.head()
canada = canada.transpose()
canada.head()
| Data Visualization/Plotly/1. Load Datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.examples.tutorials.mnist import input_data
import tensorflow as tf
# %matplotlib inline
# -
input_data
mnist = input_data.read_data_sets('MNIST_data', one_hot=True)
mnist
mnist.train.images.shape
mnist.test.images.shape
mnist.train.labels.sum(axis=0).tolist()
mnist.train.images[0].shape
np.sqrt(784)
plt.imshow(mnist.train.images[100].reshape(28, 28))
sess = tf.InteractiveSession()
sess
x = tf.placeholder(tf.float32, shape=[None, 784])
y_ = tf.placeholder(tf.float32, shape=[None, 10])
W = tf.Variable(tf.zeros([784,10]))
b = tf.Variable(tf.zeros([10]))
sess.run(tf.global_variables_initializer())
y = tf.matmul(x,W) + b
cross_entropy = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=y_, logits=y))
train_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)
train_step
for _ in range(1000):
batch = mnist.train.next_batch(100)
train_step.run(feed_dict={x: batch[0], y_: batch[1]})
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
accuracy.eval(feed_dict={x: mnist.test.images, y_: mnist.test.labels})
| tf-tutorial/mnist/tf-mnist-tutorial-multinomial-logistic-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from matplotlib import rcParams
rcParams['figure.figsize'] = 11.7,8.27 # figure size in inches
pd.options.mode.chained_assignment = None # default='warn'
pd.set_option('display.max_colwidth', None)
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 30)
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# %config Completer.use_jedi = False
import joblib
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder, StandardScaler
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import *
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from lightgbm import LGBMClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import StackingClassifier
from sklearn.ensemble import VotingClassifier
from IPython.display import display
# -
df_daybin_cat = pd.read_csv('CSV/df_merged_4_cat.csv')
# # Build models
# ## Generate result and store in dict
# * Set weight to prevent bias, 4 to CKD (1), 1 to non-CKD (0)
# * Implement GridSearchCV to find the best parameters
# + tags=[]
def train_model(model, parameters, algoname, X_train, X_test, y_train, preds, months_name):
print('========================================')
print('Training %s ' % algoname)
print()
scores = ['f1']
# retain for loop in case want to try different scores
for score in scores:
print("# Tuning hyper-parameters for %s" % score)
print()
pipe = Pipeline(steps=[('model', model)])
search = GridSearchCV(pipe, parameters, n_jobs=-1)
search.fit(X_train, y_train)
print("Best parameter (CV score=%0.3f):" % search.best_score_)
print()
print("Best parameters set found on development set:")
print()
print(search.best_params_)
print()
print("Grid scores on development set:")
print()
means = search.cv_results_['mean_test_score']
stds = search.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, search.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
# Obtained best model
optimized_model = search.best_estimator_
# Save model
joblib.dump(optimized_model, 'models/' + months_name + '/'+ algoname +'_cat.joblib')
# Get predictions
y_pred = optimized_model.predict(X_test)
y_pred_proba = optimized_model.predict_proba(X_test)[:,1] # only get probability for 1
preds[algoname] = {
'pred': y_pred,
'pred_proba': y_pred_proba
}
return preds
# -
# For ensemble
def get_predictions(model, preds, X_test, algoname):
y_pred = model.predict(X_test)
y_pred_proba = model.predict_proba(X_test)[:,1] # only get probability for 1
preds[algoname] = {
'pred': y_pred,
'pred_proba': y_pred_proba
}
return preds
# +
# Logistic regression
model_lr = LogisticRegression(random_state=0, max_iter=10000, class_weight={0: 1, 1: 4})
parameters_lr = {'model__C': [1e-05, 1e-04, 1e-03, 1e-02, 1e-01, 1, 10, 100,1000, 1e4, 1e5, 1e6]}
# Decision tree
model_dt = DecisionTreeClassifier(random_state=0, class_weight={0: 1, 1: 4}, min_samples_leaf = 30)
parameters_dt = {'model__max_depth': np.linspace(10, 100, 10)}
# lightGBM
model_lgbm = LGBMClassifier(n_estimators=1000, objective='binary',scale_pos_weight=4)
parameters_lgbm = {'model__learning_rate': [1e-05, 1e-04, 1e-03, 1e-02, 1e-01, 1, 10, 100]}
# Random forest
model_rf = RandomForestClassifier(class_weight={0: 1, 1: 4}, n_estimators=800, n_jobs=-1, verbose=1)
parameters_rf = {'model__max_depth': list(range(1,31))}
# -
# ## 0-179 days (6 months)
# +
df_6 = df_daybin_cat[df_daybin_cat['days_bin']==1]
X = df_6[['sbp', 'dbp', 'creatinine', 'glucose', 'ldl', 'hgb',
'atenolol', 'atorvastatin', 'bisoprolol', 'canagliflozin', 'carvedilol',
'dapagliflozin', 'irbesartan', 'labetalol', 'losartan', 'lovastatin',
'metformin', 'metoprolol', 'nebivolol', 'olmesartan', 'pitavastatin',
'pravastatin', 'propranolol', 'rosuvastatin', 'simvastatin',
'telmisartan', 'valsartan', 'race', 'gender', 'age']]
y = df_6['Stage_Progress']
# Split train-test data
from sklearn.model_selection import train_test_split
X_train_6, X_test_6, y_train_6, y_test_6 = train_test_split(X, y, test_size=0.2, shuffle=True, stratify=y, random_state=0)
preds_6 = {'y': y_test_6}
preds_6 = train_model(model_lr, parameters_lr, 'LogisticRegression', X_train_6, X_test_6, y_train_6, preds_6, '6m')
preds_6 = train_model(model_dt, parameters_dt, 'DecisionTreeClassifier', X_train_6, X_test_6, y_train_6, preds_6, '6m')
preds_6 = train_model(model_lgbm, parameters_lgbm, 'LGBMClassifier', X_train_6, X_test_6, y_train_6, preds_6, '6m')
preds_6 = train_model(model_rf, parameters_rf, 'RandomForestClassifier', X_train_6, X_test_6, y_train_6, preds_6, '6m')
joblib.dump(preds_6, 'predictions/predictions_6m_cat.joblib')
# + tags=[]
folder_name = '6m'
# Load models
clf1 = joblib.load('models/'+ folder_name +'/LogisticRegression_cat.joblib')
clf2 = joblib.load('models/'+ folder_name +'/RandomForestClassifier_cat.joblib')
clf3 = joblib.load('models/'+ folder_name +'/DecisionTreeClassifier_cat.joblib')
clf4 = joblib.load('models/'+ folder_name +'/LGBMClassifier_cat.joblib')
# Build ensemble
eclf = VotingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], voting='soft').fit(X_train_6, y_train_6)
# Stacked
model_stack1 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=LogisticRegression(), cv=5).fit(X_train_6, y_train_6)
model_stack2 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=RandomForestClassifier(), cv=5).fit(X_train_6, y_train_6)
model_stack3 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=LGBMClassifier(), cv=5).fit(X_train_6, y_train_6)
# Get predictions
preds_6_ensemble = {'y': y_test_6}
preds_6_ensemble = get_predictions(eclf, preds_6_ensemble, X_test_6, 'VotingClassifier')
preds_6_ensemble = get_predictions(model_stack1, preds_6_ensemble, X_test_6, 'StackingClassifier_logistic')
preds_6_ensemble = get_predictions(model_stack2, preds_6_ensemble,X_test_6, 'StackingClassifier_rf')
preds_6_ensemble = get_predictions(model_stack3, preds_6_ensemble,X_test_6, 'StackingClassifier_lgbm')
joblib.dump(preds_6_ensemble, 'predictions/preds_6_ensemble_cat.joblib')
# -
# # 180 - 359 days (12 months)
# +
df_12 = df_daybin_cat[df_daybin_cat['days_bin']==2]
X = df_12[['sbp', 'dbp', 'creatinine', 'glucose', 'ldl', 'hgb',
'atenolol', 'atorvastatin', 'bisoprolol', 'canagliflozin', 'carvedilol',
'dapagliflozin', 'irbesartan', 'labetalol', 'losartan', 'lovastatin',
'metformin', 'metoprolol', 'nebivolol', 'olmesartan', 'pitavastatin',
'pravastatin', 'propranolol', 'rosuvastatin', 'simvastatin',
'telmisartan', 'valsartan', 'race', 'gender', 'age']]
y = df_12['Stage_Progress']
# Split train-test data
from sklearn.model_selection import train_test_split
X_train_12, X_test_12, y_train_12, y_test_12 = train_test_split(X, y, test_size=0.2, shuffle=True, stratify=y, random_state=0)
preds_12 = {'y': y_test_12}
preds_12 = train_model(model_lr, parameters_lr, 'LogisticRegression', X_train_12, X_test_12, y_train_12, preds_12, '12m')
preds_12 = train_model(model_dt, parameters_dt, 'DecisionTreeClassifier', X_train_12, X_test_12, y_train_12, preds_12, '12m')
preds_12 = train_model(model_lgbm, parameters_lgbm, 'LGBMClassifier', X_train_12, X_test_12, y_train_12, preds_12, '12m')
preds_12 = train_model(model_rf, parameters_rf, 'RandomForestClassifier', X_train_12, X_test_12, y_train_12, preds_12, '12m')
joblib.dump(preds_12, 'predictions/predictions_12m_cat.joblib')
# + tags=[]
folder_name = '12m'
# Load models
clf1 = joblib.load('models/'+ folder_name +'/LogisticRegression_cat.joblib')
clf2 = joblib.load('models/'+ folder_name +'/RandomForestClassifier_cat.joblib')
clf3 = joblib.load('models/'+ folder_name +'/DecisionTreeClassifier_cat.joblib')
clf4 = joblib.load('models/'+ folder_name +'/LGBMClassifier_cat.joblib')
# Build ensemble
eclf = VotingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], voting='soft').fit(X_train_12, y_train_12)
# Stacked
model_stack1 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=LogisticRegression(), cv=5).fit(X_train_12, y_train_12)
model_stack2 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=RandomForestClassifier(), cv=5).fit(X_train_12, y_train_12)
model_stack3 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=LGBMClassifier(), cv=5).fit(X_train_12, y_train_12)
# Get predictions
preds_12_ensemble = {'y': y_test_12}
preds_12_ensemble = get_predictions(eclf, preds_12_ensemble, X_test_12, 'VotingClassifier')
preds_12_ensemble = get_predictions(model_stack1, preds_12_ensemble, X_test_12, 'StackingClassifier_logistic')
preds_12_ensemble = get_predictions(model_stack2, preds_12_ensemble,X_test_12, 'StackingClassifier_rf')
preds_12_ensemble = get_predictions(model_stack3, preds_12_ensemble,X_test_12, 'StackingClassifier_lgbm')
joblib.dump(preds_12_ensemble, 'predictions/preds_12_ensemble_cat.joblib')
# -
# # 180 - 359 days (12 months)
# +
df_18 = df_daybin_cat[df_daybin_cat['days_bin']==3]
X = df_18[['sbp', 'dbp', 'creatinine', 'glucose', 'ldl', 'hgb',
'atenolol', 'atorvastatin', 'bisoprolol', 'canagliflozin', 'carvedilol',
'dapagliflozin', 'irbesartan', 'labetalol', 'losartan', 'lovastatin',
'metformin', 'metoprolol', 'nebivolol', 'olmesartan', 'pitavastatin',
'pravastatin', 'propranolol', 'rosuvastatin', 'simvastatin',
'telmisartan', 'valsartan', 'race', 'gender', 'age']]
y = df_18['Stage_Progress']
# Split train-test data
from sklearn.model_selection import train_test_split
X_train_18, X_test_18, y_train_18, y_test_18 = train_test_split(X, y, test_size=0.2, shuffle=True, stratify=y, random_state=0)
preds_18 = {'y': y_test_18}
preds_18 = train_model(model_lr, parameters_lr, 'LogisticRegression', X_train_18, X_test_18, y_train_18, preds_18, '18m')
preds_18 = train_model(model_dt, parameters_dt, 'DecisionTreeClassifier', X_train_18, X_test_18, y_train_18, preds_18, '18m')
preds_18 = train_model(model_lgbm, parameters_lgbm, 'LGBMClassifier', X_train_18, X_test_18, y_train_18, preds_18, '18m')
preds_18 = train_model(model_rf, parameters_rf, 'RandomForestClassifier', X_train_18, X_test_18, y_train_18, preds_18, '18m')
joblib.dump(preds_18, 'predictions/predictions_18m_cat.joblib')
# + tags=[]
folder_name = '18m'
# Load models
clf1 = joblib.load('models/'+ folder_name +'/LogisticRegression_cat.joblib')
clf2 = joblib.load('models/'+ folder_name +'/RandomForestClassifier_cat.joblib')
clf3 = joblib.load('models/'+ folder_name +'/DecisionTreeClassifier_cat.joblib')
clf4 = joblib.load('models/'+ folder_name +'/LGBMClassifier_cat.joblib')
# Build ensemble
eclf = VotingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], voting='soft').fit(X_train_18, y_train_18)
# Stacked
model_stack1 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=LogisticRegression(), cv=5).fit(X_train_18, y_train_18)
model_stack2 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=RandomForestClassifier(), cv=5).fit(X_train_18, y_train_18)
model_stack3 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=LGBMClassifier(), cv=5).fit(X_train_18, y_train_18)
# Get predictions
preds_18_ensemble = {'y': y_test_18}
preds_18_ensemble = get_predictions(eclf, preds_18_ensemble, X_test_18, 'VotingClassifier')
preds_18_ensemble = get_predictions(model_stack1, preds_18_ensemble, X_test_18, 'StackingClassifier_logistic')
preds_18_ensemble = get_predictions(model_stack2, preds_18_ensemble,X_test_18, 'StackingClassifier_rf')
preds_18_ensemble = get_predictions(model_stack3, preds_18_ensemble,X_test_18, 'StackingClassifier_lgbm')
joblib.dump(preds_18_ensemble, 'predictions/preds_18_ensemble_cat.joblib')
# -
# # 540 - 719 days (24 months)
# +
df_24 = df_daybin_cat[df_daybin_cat['days_bin']==4]
X = df_24[['sbp', 'dbp', 'creatinine', 'glucose', 'ldl', 'hgb',
'atenolol', 'atorvastatin', 'bisoprolol', 'canagliflozin', 'carvedilol',
'dapagliflozin', 'irbesartan', 'labetalol', 'losartan', 'lovastatin',
'metformin', 'metoprolol', 'nebivolol', 'olmesartan', 'pitavastatin',
'pravastatin', 'propranolol', 'rosuvastatin', 'simvastatin',
'telmisartan', 'valsartan', 'race', 'gender', 'age']]
y = df_24['Stage_Progress']
# Split train-test data
from sklearn.model_selection import train_test_split
X_train_24, X_test_24, y_train_24, y_test_24 = train_test_split(X, y, test_size=0.2, shuffle=True, stratify=y, random_state=0)
preds_24 = {'y': y_test_24}
preds_24 = train_model(model_lr, parameters_lr, 'LogisticRegression', X_train_24, X_test_24, y_train_24, preds_24, '24m')
preds_24 = train_model(model_dt, parameters_dt, 'DecisionTreeClassifier', X_train_24, X_test_24, y_train_24, preds_24, '24m')
preds_24 = train_model(model_lgbm, parameters_lgbm, 'LGBMClassifier', X_train_24, X_test_24, y_train_24, preds_24, '24m')
preds_24 = train_model(model_rf, parameters_rf, 'RandomForestClassifier', X_train_24, X_test_24, y_train_24, preds_24, '24m')
joblib.dump(preds_24, 'predictions/predictions_24m_cat.joblib')
# + tags=[]
folder_name = '24m'
# Load models
clf1 = joblib.load('models/'+ folder_name +'/LogisticRegression_cat.joblib')
clf2 = joblib.load('models/'+ folder_name +'/RandomForestClassifier_cat.joblib')
clf3 = joblib.load('models/'+ folder_name +'/DecisionTreeClassifier_cat.joblib')
clf4 = joblib.load('models/'+ folder_name +'/LGBMClassifier_cat.joblib')
# Build ensemble
eclf = VotingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], voting='soft').fit(X_train_24, y_train_24)
# Stacked
model_stack1 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=LogisticRegression(), cv=5).fit(X_train_24, y_train_24)
model_stack2 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=RandomForestClassifier(), cv=5).fit(X_train_24, y_train_24)
model_stack3 = StackingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('dt', clf3), ('lgbm', clf4)], final_estimator=LGBMClassifier(), cv=5).fit(X_train_24, y_train_24)
# Get predictions
preds_24_ensemble = {'y': y_test_24}
preds_24_ensemble = get_predictions(eclf, preds_24_ensemble, X_test_24, 'VotingClassifier')
preds_24_ensemble = get_predictions(model_stack1, preds_24_ensemble, X_test_24, 'StackingClassifier_logistic')
preds_24_ensemble = get_predictions(model_stack2, preds_24_ensemble,X_test_24, 'StackingClassifier_rf')
preds_24_ensemble = get_predictions(model_stack3, preds_24_ensemble,X_test_24, 'StackingClassifier_lgbm')
joblib.dump(preds_24_ensemble, 'predictions/preds_24_ensemble_cat.joblib')
# -
# # Compare models
# +
def specificity(y_valid, y_pred):
tn, fp, fn, tp = confusion_matrix(y_valid, y_pred).ravel()
specificity = tn / (tn+fp)
return specificity
# Performance of a dichotomous diagnostic test
def youden_index(y, pred_proba):
fpr, tpr, thresh = roc_curve(y, pred_proba)
optimal_idx = np.argmax(tpr - fpr)
optimal_threshold = thresh[optimal_idx]
return optimal_threshold
# Threshold used
def get_threshold_table(preds):
thresh = [[algo, val['threshold']] for algo, val in preds.items() if algo != 'y']
return pd.DataFrame(thresh, columns=['Algo', "Threshold"])
# -
def compute_matrics(predications, days_bin):
# Calculate probability thresholds
for algo, preds in predications.items():
if algo == 'y':
continue
threshold = youden_index(predications['y'], preds['pred_proba'])
preds['threshold'] = threshold
preds['pred'] = preds['pred_proba'] >= threshold
# Get threshold to categorize as 1 (CKD)
threshold = pd.concat([get_threshold_table(preds) for preds in [predications]])
pd.set_option('display.float_format', lambda x: '%.5f' % x)
display(threshold)
# Compute matrics
metrics_list = []
y_truth = predications['y']
for algo, preds in predications.items():
if algo == 'y':
continue
y_pred = preds['pred']
y_pred_proba = preds['pred_proba']
acc = accuracy_score(y_truth, y_pred)
f1 = f1_score(y_truth, y_pred)
prec = precision_score(y_truth, y_pred)
rec = recall_score(y_truth, y_pred)
spec = specificity(y_truth, y_pred)
roc = roc_auc_score(y_truth, y_pred_proba)
row = [algo, acc, f1, prec, rec, spec, roc]
metrics_list.append(row)
cols = ['Algorithm', 'Accuracy', 'F1', 'Precision', 'Recall', 'Specificiy', 'ROC_AUC']
metrics = pd.DataFrame(metrics_list, columns=cols)
display(metrics.sort_values('ROC_AUC'))
# Plot ROC
fig = plt.figure(figsize=(6,6))
lw = 2
rocs = []
for algo, model in predications.items():
if algo == 'y':
continue
fpr, tpr, thresh = roc_curve(predications['y'], predications[algo]['pred_proba'])
auc = roc_auc_score(predications['y'], predications[algo]['pred_proba'])
auc = round(auc, 3)
rocs.append((algo, auc, [fpr, tpr]))
# sort legend by AUC
rocs = sorted(rocs, key=lambda x: x[1], reverse=True)
for algo, auc, [fpr, tpr] in rocs:
plt.plot(fpr,tpr, label=algo+f' ({auc})', lw=lw)
# chance line
plt.plot([0, 1], [0, 1], color='navy', label='chance line (0.5)', linestyle='--', lw=lw)
plt.xlabel('1 - Specificity')
plt.ylabel('Sensitivity')
# plt.title('ROC curve for 6 months', size=12)
plt.text(0.7,0.3, days_bin, fontsize=12)
plt.grid(linewidth=1.2, color='lightgray')
plt.legend(loc='lower right')
plt.savefig('figures/' + days_bin + '_daybin_cat.jpg')
compute_matrics(joblib.load('predictions/predictions_6m_cat.joblib'), '0-179 days_cat')
compute_matrics(joblib.load('predictions/preds_6_ensemble_cat.joblib'), '0-179 days_EC')
compute_matrics(joblib.load('predictions/predictions_12m_cat.joblib'), '180-359 days_cat')
compute_matrics(joblib.load('predictions/preds_12_ensemble_cat.joblib'), '180-359 days_EC')
compute_matrics(joblib.load('predictions/predictions_18m_cat.joblib'), '360-539 days_cat')
compute_matrics(joblib.load('predictions/preds_18_ensemble_cat.joblib'), '360-539 days_EC')
compute_matrics(joblib.load('predictions/predictions_24m_cat.joblib'), '540-719 days_cat')
compute_matrics(joblib.load('predictions/preds_24_ensemble_cat.joblib'), '540-719 days_EC')
| .ipynb_checkpoints/Modelling_daybin_cat-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring New Alloy Systems with Pymatgen
# #### Author: <NAME>-Robinson
# #### Version: July 29, 2020
#
# <img src="assets/goals.png" style="max-width:50%">
# ## Outline
# 1. Select a test-case system
# * 1.1 Exercise: `Structure` and `MPRester` refresher
# * 1.2 Lesson: add oxidation states to a `Structure`
# 2. Select an alloy partner
# * 2.1 Lesson: find possible dopants
# * 2.2 Exercise: find the best alloy partner (A = ?) for A<sub>x</sub>Zn<sub>1-x</sub>S
# * 2.3 Lesson: explore phase diagrams
# 3. Transform to make a new Cu<sub>x</sub>Zn<sub>1-x</sub>S alloy
# * 3.1 Lesson: structure transformation
# * 3.2 Exercise: try your own transformation on CuZnS<sub>2</sub>
# 4. Calculate new properties
# * 4.1 Lesson: volume prediction and XRD plot
# * 4.2 Exercise: try this on your CuZnS<sub>2</sub> structure
# 5. Test your skills
# * 5.1 Exercise: compare relaxed DFT structures to estimates
# * 5.2 Lesson: add computed entries to phase diagram
# * 5.3 Next steps
# ## 1. Select a test-case system
# ***In this notebook we will focus on cubic zinc-blende ZnS, a wide band gap (transparent) semiconductor. In my PhD research I study p-type transparent semiconductors, so I will pose the question: how can we use ZnS as a starting point to create a p-type transparent semiconductor, and how can pymatgen help with this?***
# Import the `MPRester` API:
# + jupyter={"outputs_hidden": false}
# -
# The Materials ID (mp-id) of zinc-blende ZnS is mp-10695, see https://materialsproject.org/materials/mp-10695/.
# + jupyter={"outputs_hidden": false}
# -
# ### 1.1 Exercise: `Structure` and `MPRester` refresher
# #### Get the structure
# + jupyter={"outputs_hidden": false}
# +
#### if you're having problems with your internet or API key:
# from monty.serialization import loadfn
# ZnS_structure = loadfn("assets/ZnS_structure.json")
# -
# #### Get space group information
# + jupyter={"outputs_hidden": false}
# -
# If you want to, try it out on our web app [here](https://materialsproject.org/#apps/xtaltoolkit/%7B%22input%22%3A0%2C%22materialIDs%22%3A%22mp-10695%22%7D).
#
# - Click "Draw atoms outside unit cell bonded to atoms within unit cell"
# - Play around with it!
# <img src="assets/ZnS_F-43m_MP.png" style="max-width:50%">
# ### 1.2 Lesson: add oxidation states to a `Structure`
# Pymatgen has a simple transformation to estimate the likely oxidation state of each specie in stoichiometric compounds using a bond-valence analysis approach. This information is needed to compare ionic radii and assess substitutional dopant probability. You can also enter the oxidation states manually if you'd prefer.
# + jupyter={"outputs_hidden": false}
# -
# Initialize this transformation:
# + jupyter={"outputs_hidden": false}
# + jupyter={"outputs_hidden": false}
# -
# ## 2. Select an alloy partner
# ### 2.1 Lesson: find possible dopants
# ***Scientific question: Which p-type dopants are most likely to sit at substitutional sites in ZnS?***
# Pymatgen has a machine-learned method for estimating the probability that one ion will substitute for another ([Hautier et al. 2011](https://doi.org/10.1021/ic102031h)), and reports the results ranked in order of probability. Note the input structure has to be "decorated" with oxidation states for this method to work.
# + jupyter={"outputs_hidden": false}
# + jupyter={"outputs_hidden": false}
# -
# Here are some options to dope ZnS p-type:
# + jupyter={"outputs_hidden": false}
# -
# We can see this returns a list of dictionaries:
# To make this easier to read we can use the `pandas` package:
# + jupyter={"outputs_hidden": false}
# -
# ### 2.2 Exercise: find the best alloy partner (A = ?) for A<sub>x</sub>Zn<sub>1-x</sub>S
# ***Scientific question: is a p-type zinc-blende A<sub>x</sub>Zn<sub>1-x</sub>S alloy possible?***
# Let's see if zinc-blende binaries exist for these ternaries, and how far off the hull they sit.
# #### Find dopants
# First, find a list of possible cation dopant elements:
# + jupyter={"outputs_hidden": false}
# I've pre-written this code block for convenience
# all it does is take the possible dopants list given previously, takes the cations, and makes a list of their elements
possible_cation_dopants = []
for x in p_dopants:
specie = x["dopant_species"]
if specie.oxi_state > 0:
possible_cation_dopants.append(str(specie.element))
# + jupyter={"outputs_hidden": false}
print(possible_cation_dopants)
# -
# Next, let's query the `MPRester` to make a table of all of the binary compounds with a space group `"F-43m"` that contain sulfur and one of these `possible_cation_dopants`. Note that the query criteria are listed on the [mapidoc](https://github.com/materialsproject/mapidoc/tree/master/materials).
# #### Query for end-point structure
# +
# the query criteria
criteria = {
_____ : { # criteria for elements
"$all": ["S"], # we require S is present
"$in": ______ # possible dopants
},
"nelements": _____, # desired number of elements
"spacegroup.symbol": _____ # desired spacegroup symbol
}
# -
# We want to return five properties: the spacegroup (`spacegroup.symbol`, which I've filled out), `task_id`, as well as energy above hull, formula, and whether the compound is experimental or theoretical. Refer to [mapidoc](https://github.com/materialsproject/mapidoc/tree/master/materials).
# +
# the properties we want to return
properties = [
_____,
_____,
_____,
_____,
"spacegroup.symbol"
]
# + jupyter={"outputs_hidden": false}
# type your query in here:
# +
#### if you're having problems with your internet or API key, or want to check your results
# from monty.serialization import loadfn
# query = loadfn("assets/alloy_partner_query.json")
# -
# Tabulate results as a `DataFrame`
# + jupyter={"outputs_hidden": false}
pd.DataFrame(_____)
# -
# Which alloy partner compound seems most reasonable? Thus, which cation should we pick? _____
# #### Retrieve end-point structure
# To proceed, we have to retrieve the `Structure` for your selected alloy partner:
# + jupyter={"outputs_hidden": false}
# -
# Yep! We’re not done, but this is a good starting point for dopants to investigate with further defect calculations. This can be accomplished using workflows from packages like [PyCDT (Broberg et al. 2018)](https://doi.org/10.1016/j.cpc.2018.01.004) which integrate with `pymatgen`'s defect capabilities.
# ### 2.3 Lesson: explore phase diagrams
# ***Scientific question: what does Cu-Zn-S phase space look like?***
# There are many built-in tools to explore phase diagrams in `pymatgen`. To build a phase diagram, you must define a set of `ComputedEntries` with compositions, formation energies, corrections, and other calculation details.
# + jupyter={"outputs_hidden": false}
# -
# We can import entries in this system using the `MPRester`. This gives a list of all of the `ComputedEntries` on the database:
# + jupyter={"outputs_hidden": false}
# +
#### if you're having problems with your internet or API key:
# from monty.serialization import loadfn
# entries = loadfn("assets/Cu-Zn-S_entries.json")
# + jupyter={"outputs_hidden": false}
# -
# #### Conventional phase diagram
# + jupyter={"outputs_hidden": false}
# -
# #### Contour phase diagram
# #### Binary phase diagram
# Let's zoom in on the tie-line between ZnS and CuS, which is where we are interested in alloying.
# #### Mapping out chemical potential of cations
# This may be a useful tool to think about tuning chemical potential for synthesis.
# There are a lot of different types of phase diagrams (see the [`pymatgen.analysis.phase_diagram` module](https://pymatgen.org/pymatgen.analysis.phase_diagram.html)).
# Our key takeaway is that in MP, the Cu-Zn-S ternary space is EMPTY!! So let's fill it in...
# ## 3. Transform to make a new Cu<sub>x</sub>Zn<sub>1-x</sub>S alloy
# ### 3.1 Lesson: structure transformation
# #### Substitute your dopant to create a disordered structure
# Now, so let's substitute 1/4 of the Zn<sup>2+</sup> with Cu<sup>+</sup> ions (note: we will be ignoring charge compensation here, but this is important to take into account in real calculations!). That is, let's set substitutional fraction `x = 1/4` in Cu<sub>x</sub>Zn<sub>1-x</sub>S. Doing so using `Structure.replace_species()` will create a ***disordered structure object***.
# + jupyter={"outputs_hidden": false}
# -
# We can print the integer formula of this composition:
# Let's rename this structure with its chemical formula to avoid confusion later on:
# + jupyter={"outputs_hidden": false}
# -
# Here's a screenshot of the CuZn<sub>3</sub>S<sub>4</sub> disordered structure, where each cation site has partial occupancy of a Zn and Cu atom.
# <img src="assets/CuZn3S4_disordered.png" style="max-width:50%">
# #### Transform structure
# Though disorder may indeed be more representative of a real crystal structure, we need to convert this to an ordered structure to perform DFT calculations. This is because DFT can only perform simulations on whole atoms, not fractional atoms!
#
# Pymatgen supports a variety of structural "transformations" (a list of supported transformations is available [here](https://pymatgen.org/pymatgen.transformations.html)). Here are three methods from the `pymatgen.transformations.advanced_transformations` module to take a disordered structure, and order it:
#
# 1. `OrderDisorderStructureTransformation`: a highly simplified method to create an ordered supercell ranked by Ewald sums.
# 2. `EnumerateStructureTransformation`: a method to order a disordered structure that requires [the `enumlib` code](https://github.com/msg-byu/enumlib) to also be installed.
# 3. `SQSTransformation`: a method that requires the [`ATAT` code (Van de Walle et al. 2013)](https://doi.org/10.1016/j.calphad.2013.06.006) to be installed that creates a special quasirandom structure (SQS) from a structure with partial occupancies.
# For this demo, we'll be focusing on the most simple transformation: `OrderDisorderStructureTransformation`
# + jupyter={"outputs_hidden": false}
# + jupyter={"outputs_hidden": false}
# -
# We have to be careful though!! If we just apply this transformation, it doesn't fail, but it returns a structure where all the Cu<sup>+</sup> is gone! `OrderDisorderedStructureTransformation` will round up or down if the cell is not large enough to account for `x`. Thus, we need to first make a supercell and then apply the transformation.
# #### Make a supercell
# With this transformation, we have to first create a disordered ***supercell*** to transform into. A supercell is just a structure that is scaled by a matrix so that it repeats several times. Here, the supercell must be large enough such that the composition in question can be achieved.
# Let's scale the structure by 8x. I like to use the `numpy` package to construct scaling matrices here (a 4x supercell would be sufficient for `x = 1/4`, but this leaves room to try e.g. `x = 1/8`):
# We can see that this would scale the cell's volume by 8, but to verify:
# For convenience, you can also simply use `scaling_matrix = 2` if you're scaling the same in all directions, or `scaling_matrix = [2, 2, 2]`. These are the same in practice.
# + jupyter={"outputs_hidden": false}
# + jupyter={"outputs_hidden": false}
# + jupyter={"outputs_hidden": false}
# + jupyter={"outputs_hidden": false}
# -
# This is a list of ten ordered structures ranked by ***Ewald sum*** (dict key `"energy"`). Note that this does NOT correlate with the lowest energy structure! Let's just use the first entry for our example:
# + jupyter={"outputs_hidden": false}
# + jupyter={"outputs_hidden": false}
# -
# If you want to download this file:
# + jupyter={"outputs_hidden": false}
# -
# BOOM! Now we have an alloy structure!! To view this structure you can upload your "CuZn3S4_ordered_structure.cif" file on [Crystal Toolkit](https://materialsproject.org/#apps/xtaltoolkit).
# <img src="assets/Zn3CuS4_P-43m_estimate.png" style="max-width:50%">
# ### 3.2 Exercise: try your own transformation on CuZnS<sub>2</sub>
# Set a new composition, `x = 1/2` (simpler fractions are easier in DFT calculations because supercells can be smaller!). This will yield a structure with composition CuZnS<sub>2</sub>.
x_CuZnS2 =
CuZnS2_disordered = ZnS_structure_oxi.copy()
CuZnS2_disordered._____(
{
"Zn2+": {
_____: _____,
_____: _____
}
}
)
# Reminder: for more complex fractions (e.g. `x = 1/16`), supercells need to be scaled accordingly!
scaling_matrix = _____
CuZnS2_disordered_supercell =
CuZnS2_ordered_structures = odst.apply_transformation(_____,
return_ranked_list = _____)
# Pick one:
CuZnS2_ordered_structure = _____
print(CuZnS2_ordered_structure)
# Check that this is the composition you expect:
# And check the space group:
# Is it the same as ZnS?
# ## 4. Calculate new properties
# ### 4.1 Lesson: volume prediction and XRD plot
# So far we just have a really rough guess of an alloy structure, and the lattice parameters are still equal to those of ZnS. We can estimate the new volume $V_{x-guess}$ after the substitution using Vegard's Law (assuming zero bowing).
# $V_{x-estimate} = V_{scaling } \times [ V_{CuS}(x) + V_{ZnS}(1-x) ] $
# $V_{CuZn_3S_4-estimate} = 8 \times [ V_{CuS}(0.25) + V_{ZnS}(0.75) ] $
# + jupyter={"outputs_hidden": false}
# -
# This is better but still wrong, and does not take into account any structural distortions. Note that there are some other methods on pymatgen to guess structure volume (see `pymatgen.analysis.structure_prediction.volume_predictor`), but in my experience Vegard's law is usually just as helpful. Your next step would be to relax this new structure using DFT or another method (see below).
# #### Calculate XRD, compare to original structure
# Now we can compare this structure to our original ZnS and CuS structure to, for example, see how the ***X-ray diffraction (XRD)*** patterns are expected to shift as `x` increases in Cu<sub>x</sub>Zn<sub>1-x</sub>S:
# Initialize the `XRDCalculator` with the conventional Cu-K$\alpha$ wavelength (note: Cu here has nothing to do with the Cu we're adding to the structure):
# + jupyter={"outputs_hidden": false}
# -
# + jupyter={"outputs_hidden": false}
# -
# You can see how the $2\theta$ peaks shift slightly to the right with addition of Cu!
# + [markdown] jupyter={"outputs_hidden": false}
# ### 4.2 Exercise: try this on your CuZnS<sub>2</sub> structure
# -
# Guess the structure volume using Vegard's Law, and correct for this:
# $V_{x-estimate} = V_{scaling } \times [ V_{CuS}(?) + V_{ZnS}(?) ] $
x_CuZnS2
scaling_volume
CuZnS2_structure_estimate = CuZnS2_ordered_structure.copy()
CuZnS2_structure_estimate.scale_lattice(_____
# Print the new structure
print(CuZnS2_structure_estimate)
# Add this structure to the series of XRD plots to compare XRD for `x = 0, 0.25, 0.5, 1`:
structures = [
ZnS_structure,
_____,
_____,
_____
]
xrd_plots = xrd.plot_structures(_____
# ## 5. Test your skills
# This is the wee beginning of making an alloy. Here are some follow-up steps:
# <img src="assets/next_steps.png" style="max-width:60%">
# I constructed similar alloys to those that we just explored, at `x = 1/4` and `x = 1/2`, and relaxed them with DFT. We'll explore my results here:
# ### 5.1 Exercise: compare relaxed DFT structures to estimates
from pymatgen import Structure
# These are my output .cif files from one of our DFT workflows. See `fireworks` and `atomate` packages for details [here](https://atomate.org/atomate.vasp.fireworks.html).
CuZn3S4_relaxed = Structure.from_file("assets/Zn3CuS4_Amm2.cif")
CuZnS2_relaxed = Structure.from_file("assets/ZnCuS2_R3m.cif")
# #### How do these space groups compare to our estimates?
# Are they higher or lower in symmetry? After relaxation, both structures are in a different space group (lower symmetry) than the alloys we just made. This is likely due to structural distortions.
# #### Add in DFT structure to XRD
# Replace the alloy structures in the previous XRD exercise with the two relaxed alloy structures, again comparing XRD for `x = 0, 0.25, 0.5, 1`:
structures = [
ZnS_structure,
_____,
_____,
CuS_structure
]
xrd_plots = xrd.plot_structures(_____
# Peak splittings are now present in the diffraction patterns, and the shift to higher $2\theta$ is not as significant.
# ### 5.2 Lesson: add computed entries to phase diagram
# ***Scientific question: are these new phases stable?***
# To assess the stability of these new phases, let's look at JSON files containing `ComputedEntry` data:
# These entries were created by relaxing the above structure using one of our DFT workflows. An "entry" is mainly just a composition and an energy, so can be created manually, without performing a calculation, or even from experimental data
# We can add these two entries to `entries`, our set of `ComputedEntry` data from MP in the Cu-Zn-S phase space:
# #### Conventional phase diagram
# We see our two new phases show up here! How does the energy landscape change?
# #### Contour phase diagram
# Compare to the phase diagram before new phases were added:
# #### Binary phase diagram
# Because these phases break the binary hull, this shows that there is likely a stable phase of CuZnS<sub>2</sub>, though this may not be the lowest energy phase. Zn<sub>3</sub>CuS<sub>4</sub> is highly metastable.
# ### 5.3 Next steps
# Here are some follow-up calculations you can do to explore ternary space:
# * ***Structure Prediction***: To explore other possibilities of polymorphs in Cu-Zn-S ternary space, one could perform a "structure prediction" in this chemical space. You can use the Materials Project website's [Structure Prediction app](https://materialsproject.org/#apps/structurepredictor). There are also methods in `pymatgen.analysis.structure_prediction` to do this.
# * ***DFT Calculations***: You can submit your structure(s) to MP by loading it with the [Crystal Toolkit app](https://materialsproject.org/#apps/crystaltoolkit) and clicking "Submit to Materials Project". You can use `fireworks` and `atomate` to submit DFT calculations yourself to relax the structure, refine it, calculate band structure, and other properties.
# * ***Local Environment Analysis***: With your relaxed structures, you can upload your structure to the [Crystal Toolkit app](https://materialsproject.org/#apps/crystaltoolkit) and look at "Analyze: Local Environments" to compare coordination environments. You can also explore the features in `pymatgen.analysis.local_env`.
# * ***Substrate Matcher***: If you want to grow this alloy structure epitaxially, you can explore different substrates to use on the website using methods in `pymatgen.analysis.substrate_analyzer module`.
# ### Thank you! Feel free to reach me at <EMAIL> if you have any questions about this lesson.
| workshop/lessons/06_new_systems/06 - New Systems - blank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="CazISR8X_HUG"
#
# + [markdown] id="WnJB6v7IIz6p"
# #<NAME>eb#
# #AI-IoT NTI and ML A-Z courses#
#
# ```
#
#
# + [markdown] id="-duevGecJAMp"
#
# + [markdown] id="pOyqYHTk_Q57"
# ## Importing the libraries
# + id="T_YHJjnD_Tja"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] id="vgC61-ah_WIz"
# ## Importing the dataset
#
#
#
# + id="UrxyEKGn_ez7" colab={"base_uri": "https://localhost:8080/"} outputId="879c324c-7622-487f-bcad-6cc8229e497e"
dataset = pd.read_csv('50_Startups.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
print(X)
# + [markdown] id="VadrvE7s_lS9"
# ## Encoding categorical data
# + id="wV3fD1mbAvsh"
# we need to change the Sate column from string values to on-hot encoded integer values
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
#the State column has now an index of 3
ct = ColumnTransformer(transformers=[('encoder', OneHotEncoder(), [3])], remainder='passthrough')
X = np.array(ct.fit_transform(X))
# + id="4ym3HdYeCGYG" colab={"base_uri": "https://localhost:8080/"} outputId="ac20651a-881b-4086-cb9c-72773036d14d"
print(X)
# + [markdown] id="WemVnqgeA70k"
# ## Splitting the dataset into the Training set and Test set
# + id="Kb_v_ae-A-20"
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# + [markdown] id="k-McZVsQBINc"
# ## Training the Multiple Linear Regression model on the Training set
# + id="ywPjx0L1BMiD" colab={"base_uri": "https://localhost:8080/"} outputId="aa8e005d-dfc7-4a4b-f43e-b6ad0f9ef335"
#Note: we dont need to do the backelimination ourselves because its already took care of in the sklearn linearregression class
#the class also cares for the dummy variabbles trap itself
from sklearn.linear_model import LinearRegression
#create a regressor object from the Linearregression class
regressor = LinearRegression()
regressor.fit(x_train, y_train)
# + [markdown] id="xNkXL1YQBiBT"
# ## Predicting the Test set results
# + id="TQKmwvtdBkyb" colab={"base_uri": "https://localhost:8080/"} outputId="6228f39d-8ef3-4526-e123-760b93f64b51"
y_pred = regressor.predict(x_test)
np.set_printoptions(precision = 2 )
#in this line, we are trying to compare the predicted and actual values in an array of 2 columns so we can visualize them.
#the
print(np.concatenate((y_pred.reshape(len(y_pred),1), y_test.reshape(len(y_test),1)), 1))
# + [markdown] id="mckKvnr8Gd_E"
#
| ML/Regression/Multiple Linear regression/Copy_of_multiple_linear_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Study session 11 - classes
# ### BIOINF 575 - Fall 2020
#
# SOLUTION
# ### Resources - classes and object oriented programming
#
# https://docs.python.org/3/tutorial/classes.html
# https://www.python-course.eu/python3_properties.php
# https://www.tutorialspoint.com/python3/python_classes_objects.htm
# https://www.w3schools.com/python/python_classes.asp
# https://www.geeksforgeeks.org/python-classes-and-objects/
# https://www.datacamp.com/community/tutorials/property-getters-setters#property
#
# ____
#
# #### Below is the Pileup class to provide functionality for a pileup object that stores information regarding alignment at a specific genomic position.
#
# +
# Pileup uses Counter so we need to import before we can create an object of that type
# The definition will run with no error bu we will not be able to use it without importing Counter
from collections import Counter
# +
# Pileup Object
class Pileup:
"""
Contains a counter of nulceotides aligned at a genomic position and
computes depth and consensus for a genomic position
Attributes:
counts (Counter): counter of all nucleotides aligned at the genomic position
depth (int): total number of reads aligned at the genomic position
consensus (str): most common nucleotide
"""
# method called when we initialize an object of type Pileup
# p = Pileup() or p = Pileup(Counter("AAA")) will call this method
def __init__(self, counts = None):
self.counts = counts
if self.counts == None:
self.counts = Counter()
self.depth = sum(self.counts.values())
if self.depth == 0:
self.__consensus = "" # using __ at the beginning of the attribute name makes it inaccessible outside of the class definition
else:
self.__consensus = self.counts.most_common()[0][0]
# see more examples in the last link in RESOURCES
# property (the @property above the function makes it so)
# should be used without () when called for an object e.g. p.consensus
# getter that allows us to display the value of a hidden attribute outside of the class definition
@property
def consensus(self):
"""
Get the consensus nucleotide for the pileup
"""
return self.__consensus
# method that implements what the string representation of a Pileup object is
# for a Pileup object p this method is called when print(p) or str(p) are used
def __str__(self):
return f"Pileup({self.counts})"
# method that implements what the string representation of a Pileup object is
# and that allows you to create another object idential to the one you are calling it for
# it is called when you use the name of the object to see its value
# typing
# it is called when print(p) or str(p) are used
def __repr__(self):
return f"Pileup({self.counts})"
# used to implement the addition operator
# if p1 and p2 are Pileup objects this is called when we compute p1+p2
# self will be assigned p1 and p will be assigned p2 in the __add__ method
def __add__(self, p):
c = self.counts.copy() # make a copy otherwise we are updating p1 when we do p1+p2
c.update(p.counts)
return Pileup(c)
#the first argument of every method in a class (self) will be assigned the value of the object we call it from
# e.g if variable p contains a pileup object the statement p.update("AAA") will call this method below and assign p to self and "AAA" to seq
def update(self, seq):
"""
Update the counts depth and consensus for the pileup
Given a sequence of nucleotides to add to the pileup
"""
self.counts.update(seq)
self.depth = sum(self.counts.values())
self.__consensus = self.counts.most_common()[0][0]
# -
# ### We copied and ran all or part of the below code for each exercise to test we did not break something while trying to implement the exercise requirement
p = Pileup()
p
# hidden attribute only accesible in the class definition
p.__consensus
# +
#dir(p)
# -
# Nothing in Python is truly private; internally
p._Pileup__consensus = 3
p.consensus
p.counts
p.depth
# we have only implementerd a getter (a way to retrieve the value)
# a setter (a way to set the value) was not implemented
# we want this to be changed only internally when the counts change
p.consensus = "T"
# we update the counts but the consensus and depth stay the same
# we are going to make this a property so we can also update the depth and consensus when we assign a value to the counts
# see the class defintion below
p.counts = Counter("ACGTTTT")
p.counts
p.depth
p.consensus
p.depth = 40
p.depth
p.update("ACGTTGGGG")
p.counts
p.consensus
p.depth
# ### <font color = "red">Exercise</font>
# #### - Make the Pileup attribute depth read only
# #### - Test the change
# +
# Pileup Object
class Pileup:
"""
Contains a counter of nulceotides aligned at a genomic position and
computes depth and consensus for a genomic position
Attributes:
counts (Counter): counter of all nucleotides aligned at the genomic position
depth (int): total number of reads aligned at the genomic position
consensus (str): most common nucleotide
"""
def __init__(self, counts = None):
self.__counts = counts
if self.__counts == None:
self.__counts = Counter()
self.__depth = sum(self.__counts.values())
if self.__depth == 0:
self.__consensus = ""
else:
self.__consensus = self.__counts.most_common()[0][0]
@property # getter
def consensus(self):
"""
Get the consensus nucleotide for the pileup
"""
return self.__consensus
@property # getter
def depth(self):
"""
Get the depth for the pileup - number of reads that aliged at that position
"""
return self.__depth
@property # getter
def counts(self):
"""
Get the counts for the pileup - frequencies of the nucleotides that aligned at that position
"""
return self.__counts
@counts.setter # property setter used when we assign a value to the property: p.counts = Counter("AACG")
def counts(self, counts_value):
self.__counts = counts_value
self.__depth = sum(self.__counts.values())
self.__consensus = self.__counts.most_common()[0][0]
def __str__(self):
return f"Pileup({self.__counts})"
def __repr__(self):
return f"Pileup({self.__counts})"
def __add__(self, p):
c = self.__counts.copy()
c.update(p.counts)
return Pileup(c)
def update(self, seq):
"""
Update the counts, depth and consensus nucleotide for the pileup
given a sequence of nucleotides to add to the pileup
"""
self.__counts.update(seq)
self.__depth = sum(self.__counts.values())
self.__consensus = self.__counts.most_common()[0][0]
# -
# #### We also updated the counts to a property in the class definition above to be able to update the consensus and depth when we set up the counts
p = Pileup()
p
p.consensus
p.counts
p.depth
p.consensus = "T"
p.counts = Counter("ACGTTTT")
p.counts
p.depth
p.consensus
# +
# depth is now read only as intended
# it can only be computed using the counts and gets updated every time the counts change
p.depth = 40
# -
p.depth
p.update("ACGTTGGGG")
p.counts
p.consensus
p.depth
# ### <font color = "red">Exercise</font>
# #### - Implement a dunder method so that we can apply the `len()` function to the pileup object
# #### - Test the change
# we did not tell our class what to do when we apply the function len to our object
# we can do that by implementing the __len__ method see that in the class definition below
len(p)
# +
# Pileup Object
class Pileup:
"""
Contains a counter of nulceotides aligned at a genomic position and
computes depth and consensus for a genomic position
Attributes:
counts (Counter): counter of all nucleotides aligned at the genomic position
depth (int): total number of reads aligned at the genomic position
consensus (str): most common nucleotide
"""
def __init__(self, counts = None):
self.__counts = counts
if self.__counts == None:
self.__counts = Counter()
self.__depth = sum(self.__counts.values())
if self.__depth == 0:
self.__consensus = ""
else:
self.__consensus = self.__counts.most_common()[0][0]
@property # getter
def consensus(self):
"""
Get the consensus nucleotide for the pileup
"""
return self.__consensus
# @set.consensus # property setter
# def consensus(self, cons):
# self.__consensus = consensus
@property # getter
def depth(self):
"""
Get the depth for the pileup - number of reads that aliged at that position
"""
return self.__depth
@property # getter
def counts(self):
"""
Get the counts for the pileup - frequencies of the nucleotides that aligned at that position
"""
return self.__counts
@counts.setter # property setter
def counts(self, counts_value):
self.__counts = counts_value
self.__depth = sum(self.__counts.values())
self.__consensus = self.__counts.most_common()[0][0]
def __str__(self):
return f"Pileup({self.__counts})"
def __repr__(self):
return f"Pileup({self.__counts})"
def __add__(self, p):
c = self.__counts.copy()
c.update(p.counts)
return Pileup(c)
# called when the len function is applied to a Pileup object
# e.g. len(p)
def __len__(self):
return self.__depth
# this is method to test that the first argument in the method is self
def test(arg1, x):
print(x)
def update(self, seq):
"""
Update the counts, depth and consensus nucleotide for the pileup
given a sequence of nucleotides to add to the pileup
"""
self.__counts.update(seq)
self.__depth = sum(self.__counts.values())
self.__consensus = self.__counts.most_common()[0][0]
# -
p = Pileup()
p
p.consensus
p.counts
p.depth
p.consensus = "T"
p.counts = Counter("ACGTTTT")
p.counts
p.depth
p.consensus
p.depth = 40
p.depth
p.update("ACGTTGGGG")
p.counts
p.consensus
p.depth
len(p)
len(5)
# +
# from the class we will apply the test method
## this is method to test that the first argument in the method is self
# def test(arg1, x):
# print(x)
# The test method only has one argument
# the first argument (arg1) is not counted
# since it is automatically assigned the value of the variable that is being called from in this case p
p.test()
# -
p.test(5)
p.test(p)
# +
# The test method only has one argument
# the first argument (arg1) is not counted
# since it is automatically assigned the value of the variable that is being called from
p.test(3,4)
# -
# ### <font color = "red">Exercise</font>
# #### - Create a list of 10 Pileup objects
# #### - Update the pileup objects with the corresponding sequences fom the following list.
seq_list = ["CCCCATTTG","CATTTAG","GGGATC","AACTGA", "GCCCTAA", "CCCCATTTG", "AAAAAC","TTTTTTG","GGGGAT", "TTTTA"]
# +
pileup_list = [Pileup(Counter(seq_list[i])) for i in range(10)]
# -
pileup_list
# creating a list of empty pileups
pileup_list = [Pileup() for i in range(10)]
pileup_list
# +
# creating a list of pileups based on the sequence in seq_list
for i in range(10):
pileup_list[i].update(seq_list[i])
# -
pileup_list
pileup_list = [Pileup() for i in range(10)]
pileup_list
list(map(Pileup.update, pileup_list, seq_list))
pileup_list
# ### <font color = "red">Exercise</font>
# #### - Implement a method that compares the pileup it is called for with another given pileup and returns a tuple with the consensus for the two pileups and the ratio of the consensus frequencies available for the consensus in the counts attribute.
# #### - Test the change
# +
# Pileup Object
class Pileup:
"""
Contains a counter of nulceotides aligned at a genomic position and
computes depth and consensus for a genomic position
Attributes:
counts (Counter): counter of all nucleotides aligned at the genomic position
depth (int): total number of reads aligned at the genomic position
consensus (str): most common nucleotide
"""
def __init__(self, counts = None):
self.__counts = counts
if self.__counts == None:
self.__counts = Counter()
self.__depth = sum(self.__counts.values())
if self.__depth == 0:
self.__consensus = ""
else:
self.__consensus = self.__counts.most_common()[0][0]
@property # getter
def consensus(self):
"""
Get the consensus nucleotide for the pileup
"""
return self.__consensus
# @set.consensus # property setter
# def consensus(self, cons):
# self.__consensus = consensus
@property # getter
def depth(self):
"""
Get the depth for the pileup - number of reads that aliged at that position
"""
return self.__depth
@property # getter
def counts(self):
"""
Get the counts for the pileup - frequencies of the nucleotides that aligned at that position
"""
return self.__counts
@counts.setter # property setter
def counts(self, counts_value):
self.__counts = counts_value
self.__depth = sum(self.__counts.values())
self.__consensus = self.__counts.most_common()[0][0]
def __str__(self):
return f"Pileup({self.__counts})"
def __repr__(self):
return f"Pileup({self.__counts})"
def __add__(self, p):
c = self.__counts.copy()
c.update(p.counts)
return Pileup(c)
def __len__(self):
return self.__depth
def update(self, seq):
"""
Update the counts, depth and consensus for the pileup
given a sequence of nucleotides to add to the pileup
"""
self.__counts.update(seq)
self.__depth = sum(self.__counts.values())
self.__consensus = self.__counts.most_common()[0][0]
def compare_pileup(self, p):
"""
Creates a tuple with three elements:
- consensus of the pileup we are calling this method for
- consensus of the pileup p given as an argumet
- ration of the frequencies of the consensus in the counts for each of the pileups
"""
tuple_res = ()
f1 = self.__counts[self.__consensus]
f2 = p.counts[p.consensus]
try:
tuple_res = (self.__consensus, p.consensus, f1/f2)
except ZeroDivisionError:
tuple_res = (self.__consensus, p.consensus, 0)
return tuple_res
# -
3/0
p = Pileup()
p
p.consensus
p.counts
p.depth
p.consensus = "T"
p.counts = Counter("ACGTTTT")
p.counts
p.depth
p.consensus
p.depth = 40
p.depth
p.update("ACGTTGGGG")
p.counts
p.consensus
p.depth
len(p)
len(5)
p1 = Pileup(Counter("AACTTTGAAAAA"))
p1
p.compare_pileup(p1)
p.compare_pileup(Pileup())
pileup_list = [Pileup(Counter(seq_list[i])) for i in range(10)]
pileup_list
pileup_list[0].compare_pileup(pileup_list[1])
# ### An alternative is to compare the pileups and compute the tuple (variant) in a function outside of the class
# +
# Pileup Object
class Pileup:
"""
Contains a counter of nulceotides aligned at a genomic position and
computes depth and consensus for a genomic position
Attributes:
counts (Counter): counter of all nucleotides aligned at the genomic position
depth (int): total number of reads aligned at the genomic position
consensus (str): most common nucleotide
"""
def __init__(self, counts = None):
self.__counts = counts
if self.__counts == None:
self.__counts = Counter()
self.__depth = sum(self.__counts.values())
if self.__depth == 0:
self.__consensus = ""
else:
self.__consensus = self.__counts.most_common()[0][0]
@property # getter
def consensus(self):
"""
Get the consensus nucleotide for the pileup
"""
return self.__consensus
# @set.consensus # property setter
# def consensus(self, cons):
# self.__consensus = consensus
@property # getter
def depth(self):
"""
Get the depth for the pileup - number of reads that aliged at that position
"""
return self.__depth
@property # getter
def counts(self):
"""
Get the counts for the pileup - frequencies of the nucleotides that aligned at that position
"""
return self.__counts
@counts.setter # property setter
def counts(self, counts_value):
self.__counts = counts_value
self.__depth = sum(self.__counts.values())
self.__consensus = self.__counts.most_common()[0][0]
def __str__(self):
return f"Pileup({self.__counts})"
def __repr__(self):
return f"Pileup({self.__counts})"
def __add__(self, p):
c = self.__counts.copy()
c.update(p.counts)
return Pileup(c)
def __len__(self):
return self.__depth
def test(self, x):
print(x)
def update(self, seq):
"""
Update the counts, depth and consensus for the pileup
given a sequence of nucleotides to add to the pileup
"""
self.__counts.update(seq)
self.__depth = sum(self.__counts.values())
self.__consensus = self.__counts.most_common()[0][0]
# -
def create_variant(pn,pt):
"""
Creates a tuple with three elements:
- consensus of the pileup we are calling this method for
- consensus of the pileup p given as an argumet
- ration of the frequencies of the consensus in the counts for each of the pileups
"""
tuple_res = ()
f1 = pn.counts[pn.consensus]
f2 = pt.counts[pt.consensus]
try:
tuple_res = (pn.consensus, pt.consensus, f1/f2)
except ZeroDivisionError:
tuple_res = (pn.consensus, pt.consensus, 0)
return tuple_res
p1 = Pileup(Counter("CCCGT"))
create_variant(p1, Pileup())
p2 = Pileup(Counter("GGGGTACGGGG"))
create_variant(p2, p1)
# ## Unrelated extra example
# ### Passing arguments to a function by value or by reference
# +
## By reference - using the variable name for a list, set, dict ...
# -
def test(d):
d["A"] = 2120
d1 = {"A":4, "B":5}
d1
test(d1)
d1
# +
## By value - unpacking the vaariable values using * or ** for a list, set, dict ...
# -
def test(**d):
x = d["A"]
x = x + 5
d["A"] = 4000
test(**d1)
d1
| study_sessions/study_session11_classes_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: py35-paddle1.2.0
# ---
# +
import requests
import random
import json
from hashlib import md5
import time
import threading
mutex = threading.Lock()
# Set your own appid/appkey.
appid = '20211020000977904' #your personel appid which you can apply for free from baidu
appkey = '<KEY>' #your personel appkey
endpoint = 'http://api.fanyi.baidu.com'
path = '/api/trans/vip/translate'
url = endpoint + path
headers = {'Content-Type': 'application/x-www-form-urlencoded'}
last_send_time = 0
# Generate salt and sign
def make_md5(s, encoding='utf-8'):
return md5(s.encode(encoding)).hexdigest()
#auto to target
def translate2auto(query_txt, from_lang = "auto", to_lang = "zh"):
global last_send_time;
mutex.acquire()
cur_time = time.time() * 1000
diff = cur_time - last_send_time
# print("diff: ", diff)
if diff < 1000:
time.sleep((1000 - diff) / 1000.0)
salt = random.randint(32768, 65536)
print(query_txt)
sign = make_md5(appid + query_txt + str(salt) + appkey)
payload = {'appid': appid, 'q': query_txt, 'from': from_lang, 'to': to_lang, 'salt': salt, 'sign': sign}
# Send request
r = requests.post(url, params=payload, headers=headers)
result = r.json()
# Show response
print(result)
last_send_time = time.time()*1000
# print("last_send_time: ", last_send_time)
mutex.release()
return result["from"], result["trans_result"][0]["dst"]
# -
print(translate2auto("Guten Tag!", "auto", "en"))
print(translate2auto("study hard", "auto", "en"))
print(translate2auto("everything will be ok", "auto", "zh"))
| Systemcode/imental-Chatbot/Baidu_AISTUDIO_BML_Codelab/baidu_translate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Auto Visualization using pandas_visual_analysis
# This Code template is for the Exploratory Data Analysis using the PandasVisualAnalysis. It is a package provided by python for interactive visual analysis in a Jupyter notebook. It generates an interactive visual analysis widget to analyze pandas Data Frame.
# ### Required Packages
#installation
# !pip install pandas-visual-analysis
import pandas as pd
from pandas_visual_analysis import VisualAnalysis
import warnings
warnings.filterwarnings("ignore")
# #### Dataset
#
#
# Filepath of CSV file
#filepath
file_path="https://raw.githubusercontent.com/Thilakraj1998/Datasets_general/main/Iris.csv"
df=pd.read_csv(file_path)
df.head()
# ### Visualization
#
# pandas_visual_analysis provides VisualAnalysis module which generates an interactive visual analysis widget to analyze a pandas DataFrame in Jupyter notebooks. It can display various different types of graphs with support for linked-brushing in interactive widgets. This allows data exploration and cognition to be simple, even with complex multivariate datasets.
#
# #### Arguments
#
# * **data (Union[DataFrame, DataSource])** – A pandas.DataFrame object or a DataSource.
#
# * **layout (Union[str, List[List[str]]])** – Layout specification name or explicit definition of widget names in rows. Those columns have to include all columns of the DataFrame which have type object, str, bool or category. This means it can only add columns which do not have the aforementioned types. Defaults to ‘default’.
#
# * **categorical_columns (Optional[List[str]])** – If given, specifies which columns are to be interpreted as categorical. Defaults to None.
#
# * **row_height (Union[int, List[int]])** – Height in pixels each row should have. If given an integer, each row has the height specified by that value, if given a list of integers, each value in the list specifies the height of the corresponding row. Defaults to 400.
#
# * **sample (Union[float, int, None])** – Int or float value specifying if the DataFrame should be sub-sampled. When an int is given, the DataFrame will be limited to that number of rows given by the value. When a float is given, the DataFrame will include the fraction of rows given by the value. Defaults to None.
#
# * **select_color (Union[str, Tuple[int, int, int]])** – RGB tuple or hex color specifying the color display selected data points. Values in the tuple have to be between 0 and 255 inclusive or a hex string that converts to such RGB values. Defaults to ‘#323EEC’.
#
# * **deselect_color (Union[str, Tuple[int, int, int]])** – RGB tuple or hex color specifying the color display deselected data points. Values in the tuple have to be between 0 and 255 inclusive or a hex string that converts to such RGB values. Defaults to ‘#8A8C93’.
#
# * **alpha (float)** – Opacity of data points when applicable ranging from 0.0 to 1.0 inclusive. Defaults to 0.75.
#
# * **seed (Optional[int])** – Random seed used for sampling the data. Values can be any integer between 0 and 2**32 - 1 inclusive or None. Defaults to None.
VisualAnalysis(df)
# #### Creator: <NAME> , Github: [Profile](https://github.com/Thilakraj1998)
#
| EDA/Auto Visualization/PandasVisualAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NLP Exercises
#
# We have five exercises in this section. The exercises are:
# 1. Build your own tokenizer, where you need to implement two functions to implement a tokenizer based on regular expression.
# 2. Get tags from Trump speech.
# 3. Get the nouns in the last 10 sentences from Trump's speech and find the nouns divided by sentencens. Use SpaCy.
# 4. Build your own Bag Of Words implementation using tokenizer created before.
# 5. Build a 5-gram model and clean up the results.
# ## Exercise 1. Build your own tokenizer
#
# Build two different tokenizers:
# - ``tokenize_sentence``: function tokenizing text into sentences,
# - ``tokenize_word``: function tokenizing text into words.
# +
from typing import List
import re
def tokenize_words(text: str) -> list:
"""Tokenize text into words using regex.
Parameters
----------
text: str
Text to be tokenized
Returns
-------
List[str]
List containing words tokenized from text
"""
return re.sub("[^a-zA-Z0-9]", " ", text.lower()).split()
def tokenize_sentence(text: str) -> list:
"""Tokenize text into words using regex.
Parameters
----------
text: str
Text to be tokenized
Returns
-------
List[str]
List containing words tokenized from text
"""
tokens = []
return re.split(r"(?<=[?.!])\s(?=[:oA-Z])", text)
text = "Here we go again. I was supposed to add this text later. \
Well, it's 10.p.m. here, and I'm actually having fun making this course. :o\
I hope you are getting along fine with this presentation, I really did try. \
And one last sentence, just so you can test you tokenizers better."
print("Tokenized sentences:")
print(tokenize_sentence(text))
print("Tokenized words:")
print(tokenize_words(text))
# -
# ## Exercise 2. Get tags from Trump speech using NLTK
#
# You should use the ``trump.txt`` file, read it and find the tags for each word. Use NLTK for it.
# +
from nltk import word_tokenize, pos_tag
with open("../datasets/trump.txt", "r",encoding="utf-8") as file:
trump = file.read()
words = word_tokenize(trump)
print(pos_tag(words))
# fill the gap and imports
# -
# ## Exercise 3. Get the nouns in the last 10 sentences from Trump's speech and find the nouns divided by sentencens. Use SpaCy.
#
# Please use Python list features to get the last 10 sentences and display nouns from it.
# +
import spacy
with open("../datasets/trump.txt", "r",encoding='utf-8') as file:
trump = file.read()
nlp = spacy.load("en_core_web_sm")
doc = nlp(trump)
lastTen = list(doc.sents)[-10:]
for sen in lastTen:
for token in sen:
if token.pos_ == "NOUN":
print(token)
### here comes your code
# -
# ## Exercise 4. Build your own Bag Of Words implementation using tokenizer created before
#
# You need to implement following methods:
#
# - ``fit_transform`` - gets a list of strings and returns matrix with it's BoW representation
# - ``get_features_names`` - returns list of words corresponding to columns in BoW
# +
import numpy as np
import spacy
import pandas as pd
class BagOfWords:
"""Basic BoW implementation."""
__nlp = spacy.load("en_core_web_sm")
__bow_list = []
# your code goes maybe also here
def fit_transform(self, corpus: list):
"""Transform list of strings into BoW array.
Parameters
----------
corpus: List[str]
Corpus of texts to be transforrmed
Returns
-------
np.array
Matrix representation of BoW
"""
wordset = []
for sentence in corpus:
wordset = np.union1d(wordset, tokenize_words(sentence))
self.__bow_list = list(wordset)
bow = []
for i, sentence in enumerate(corpus):
bow.append(dict.fromkeys(wordset, 0))
for word in tokenize_words(sentence):
bow[i][word] = sentence.count(word)
return pd.DataFrame(bow)
def get_feature_names(self) -> list:
"""Return words corresponding to columns of matrix.
Returns
-------
List[str]
Words being transformed by fit function
"""
return self.__bow_list
corpus = [
'Bag Of Words is based on counting',
'words occurences throughout multiple documents.',
'This is the third document.',
'As you can see most of the words occur only once.',
'This gives us a pretty sparse matrix, see below. Really, see below',
]
vectorizer = BagOfWords()
X = vectorizer.fit_transform(corpus)
print(X)
vectorizer.get_feature_names()
len(vectorizer.get_feature_names())
# -
# ## Exercise 5. Build a 5-gram model and clean up the results.
#
# There are three tasks to do:
# 1. Use 5-gram model instead of 3.
# 2. Change to capital letter each first letter of a sentence.
# 3. Remove the whitespace between the last word in a sentence and . ! or ?.
#
# Hint: for 2. and 3. implement a function called ``clean_generated()`` that takes the generated text and fix both issues at once. It could be easier to fix the text after it's generated rather then doing some changes in the while loop.
# +
from nltk.book import *
wall_street = text7.tokens
import re
tokens = wall_street
def cleanup():
compiled_pattern = re.compile("^[a-zA-Z0-9.!?]")
clean = list(filter(compiled_pattern.match,tokens))
return clean
tokens = cleanup()
def build_ngrams():
ngrams = []
for i in range(len(tokens)-N+1):
ngrams.append(tokens[i:i+N])
return ngrams
def ngram_freqs(ngrams):
counts = {}
for ngram in ngrams:
token_seq = SEP.join(ngram[:-1])
last_token = ngram[-1]
if token_seq not in counts:
counts[token_seq] = {}
if last_token not in counts[token_seq]:
counts[token_seq][last_token] = 0
counts[token_seq][last_token] += 1
return counts
def next_word(text, N, counts):
token_seq = SEP.join(text.split()[-(N-1):])
choices = counts[token_seq].items()
total = sum(weight for choice, weight in choices)
r = random.uniform(0, total)
upto = 0
for choice, weight in choices:
upto += weight;
if upto > r: return choice
assert False # should not reach here
# +
import re
def clean_generated(text):
text = text.replace(text[0], text[0].upper(), 1)
text = re.sub("(?<=\s[.!?])\s(\w)", lambda x: x.group(0).upper(), text)
text = re.sub("\s[.!?]", lambda x: x.group(0)[1], text)
return text
import random
N=5 # fix it for other value of N
SEP=" "
sentence_count=5
ngrams = build_ngrams()
start_seq="are assigned lunch partners"
counts = ngram_freqs(ngrams)
if start_seq is None:
start_seq = random.choice(list(counts.keys()))
generated = start_seq.lower();
sentences = 0
while sentences < sentence_count:
generated += SEP + next_word(generated, N, counts)
sentences += 1 if generated.endswith(('.','!', '?')) else 0
generated = clean_generated(generated)
# put your code here:
print(generated)
| ML1/nlp/106_NLP_Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
# read image
img = cv2.imread('male.jpg')
#convert to grayscale
gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
#apply haar cascade
haar = cv2.CascadeClassifier('haarcascade_frontalface_default.xml')
faces = haar.detectMultiScale(gray,1.3,5)
print(faces)
cv2.rectangle(img,(80,120),(154+250,94+350),(0,255,0),5)
plt.imshow(img)
cv2.imshow('object_detect',img)
cv2.waitKey(0)
cv2.destroyAllWindows()
#crop
face_crop = img[120:94+350,80:154+250]
plt.imshow(face_crop)
cv2.imwrite('male_01.png',face_crop)
| OpenCV_Object_Detection/04 - Object Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bootstrap Examples
# _This setup code is required to run in an IPython notebook_
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn
seaborn.set_style("darkgrid")
plt.rc("figure", figsize=(16, 6))
plt.rc("savefig", dpi=90)
plt.rc("font", family="sans-serif")
plt.rc("font", size=14)
# -
# ## Sharpe Ratio
# The Sharpe Ratio is an important measure of return per unit of risk. The example shows how to estimate the variance of the Sharpe Ratio and how to construct confidence intervals for the Sharpe Ratio using a long series of U.S. equity data.
# +
import arch.data.frenchdata
import numpy as np
import pandas as pd
ff = arch.data.frenchdata.load()
# -
# The data set contains the Fama-French factors, including the excess market return.
excess_market = ff.iloc[:, 0]
print(ff.describe())
# The next step is to construct a function that computes the Sharpe Ratio. This function also return the annualized mean and annualized standard deviation which will allow the covariance matrix of these parameters to be estimated using the bootstrap.
def sharpe_ratio(x):
mu, sigma = 12 * x.mean(), np.sqrt(12 * x.var())
values = np.array([mu, sigma, mu / sigma]).squeeze()
index = ["mu", "sigma", "SR"]
return pd.Series(values, index=index)
# The function can be called directly on the data to show full sample estimates.
params = sharpe_ratio(excess_market)
params
# ## Reproducibility
#
# All bootstraps accept the keyword argument `seed` which can contain a NumPy `Generator` or `RandomState` or an `int`. When using an `int`, the argument is passed `np.random.default_rng` to create the core generator. This allows the same pseudo random values to be used across multiple runs.
# ### _Warning_
#
# _The bootstrap chosen must be appropriate for the data. Squared returns are serially correlated, and so a time-series bootstrap is required._
# Bootstraps are initialized with any bootstrap specific parameters and the data to be used in the bootstrap. Here the `12` is the average window length in the Stationary Bootstrap, and the next input is the data to be bootstrapped.
# +
from arch.bootstrap import StationaryBootstrap
# Initialize with entropy from random.org
entropy = [877788388, 418255226, 989657335, 69307515]
seed = np.random.default_rng(entropy)
bs = StationaryBootstrap(12, excess_market, seed=seed)
results = bs.apply(sharpe_ratio, 2500)
SR = pd.DataFrame(results[:, -1:], columns=["SR"])
fig = SR.hist(bins=40)
# -
cov = bs.cov(sharpe_ratio, 1000)
cov = pd.DataFrame(cov, index=params.index, columns=params.index)
print(cov)
se = pd.Series(np.sqrt(np.diag(cov)), index=params.index)
se.name = "Std Errors"
print("\n")
print(se)
ci = bs.conf_int(sharpe_ratio, 1000, method="basic")
ci = pd.DataFrame(ci, index=["Lower", "Upper"], columns=params.index)
print(ci)
# Alternative confidence intervals can be computed using a variety of methods. Setting `reuse=True` allows the previous bootstrap results to be used when constructing confidence intervals using alternative methods.
ci = bs.conf_int(sharpe_ratio, 1000, method="percentile", reuse=True)
ci = pd.DataFrame(ci, index=["Lower", "Upper"], columns=params.index)
print(ci)
# ### Optimal Block Length Estimation
#
# The function `optimal_block_length` can be used to estimate the optimal block lengths for the Stationary and Circular bootstraps. Here we use the squared market return since the Sharpe ratio depends on the mean and the variance, and the autocorrelation in the squares is stronger than in the returns.
# +
from arch.bootstrap import optimal_block_length
opt = optimal_block_length(excess_market**2)
print(opt)
# -
# We can repeat the analysis above using the estimated optimal block length. Here we see that the extremes appear to be slightly more extreme.
# +
# Reinitialize using the same entropy
rs = np.random.default_rng(entropy)
bs = StationaryBootstrap(opt.loc["Mkt-RF", "stationary"], excess_market, seed=seed)
results = bs.apply(sharpe_ratio, 2500)
SR = pd.DataFrame(results[:, -1:], columns=["SR"])
fig = SR.hist(bins=40)
# -
# ## Probit (statsmodels)
# The second example makes use of a Probit model from statsmodels. The demo data is university admissions data which contains a binary variable for being admitted, GRE score, GPA score and quartile rank. This data is downloaded from the internet and imported using pandas.
# +
import arch.data.binary
binary = arch.data.binary.load()
binary = binary.dropna()
print(binary.describe())
# -
# ### Fitting the model directly
# The first steps are to build the regressor and the dependent variable arrays. Then, using these arrays, the model can be estimated by calling `fit`
# +
import statsmodels.api as sm
endog = binary[["admit"]]
exog = binary[["gre", "gpa"]]
const = pd.Series(np.ones(exog.shape[0]), index=endog.index)
const.name = "Const"
exog = pd.DataFrame([const, exog.gre, exog.gpa]).T
# Estimate the model
mod = sm.Probit(endog, exog)
fit = mod.fit(disp=0)
params = fit.params
print(params)
# -
# ### The wrapper function
# Most models in statsmodels are implemented as classes, require an explicit call to `fit` and return a class containing parameter estimates and other quantities. These classes cannot be directly used with the bootstrap methods. However, a simple wrapper can be written that takes the data as the only inputs and returns parameters estimated using a Statsmodel model.
def probit_wrap(endog, exog):
return sm.Probit(endog, exog).fit(disp=0).params
# A call to this function should return the same parameter values.
probit_wrap(endog, exog)
# The wrapper can be directly used to estimate the parameter covariance or to construct confidence intervals.
# +
from arch.bootstrap import IIDBootstrap
bs = IIDBootstrap(endog=endog, exog=exog)
cov = bs.cov(probit_wrap, 1000)
cov = pd.DataFrame(cov, index=exog.columns, columns=exog.columns)
print(cov)
# -
se = pd.Series(np.sqrt(np.diag(cov)), index=exog.columns)
print(se)
print("T-stats")
print(params / se)
ci = bs.conf_int(probit_wrap, 1000, method="basic")
ci = pd.DataFrame(ci, index=["Lower", "Upper"], columns=exog.columns)
print(ci)
# ### Speeding things up
# Starting values can be provided to `fit` which can save time finding starting values. Since the bootstrap parameter estimates should be close to the original sample estimates, the full sample estimated parameters are reasonable starting values. These can be passed using the `extra_kwargs` dictionary to a modified wrapper that will accept a keyword argument containing starting values.
def probit_wrap_start_params(endog, exog, start_params=None):
return sm.Probit(endog, exog).fit(start_params=start_params, disp=0).params
bs.reset() # Reset to original state for comparability
cov = bs.cov(
probit_wrap_start_params, 1000, extra_kwargs={"start_params": params.values}
)
cov = pd.DataFrame(cov, index=exog.columns, columns=exog.columns)
print(cov)
# ## Bootstrapping Uneven Length Samples
# Independent samples of uneven length are common in experiment settings, e.g., A/B testing of a website. The `IIDBootstrap` allows for arbitrary dependence within an observation index and so cannot be naturally applied to these data sets. The `IndependentSamplesBootstrap` allows datasets with variables of different lengths to be sampled by exploiting the independence of the values to separately bootstrap each component. Below is an example showing how a confidence interval can be constructed for the difference in means of two groups.
# +
from arch.bootstrap import IndependentSamplesBootstrap
def mean_diff(x, y):
return x.mean() - y.mean()
rs = np.random.RandomState(0)
treatment = 0.2 + rs.standard_normal(200)
control = rs.standard_normal(800)
bs = IndependentSamplesBootstrap(treatment, control, seed=seed)
print(bs.conf_int(mean_diff, method="studentized"))
| examples/bootstrap_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Food Group Nutrients Analysis - RightFood_HealthyFood
# ## Project Goal
# The goal of this project is to check the amount of calories and other nutrient present in different categories of food. From the finding, we will be able understand the quality of food that we are consuming every day. It will help each and every individual who are diet conscious or people who are trying to figure out the quality of food they are consuming. After the analysis it will be easy for a person to pick the right food over the counter.
# ## Summary of Data
# The dataset used for analysis contains 40 major food nutrients like calories, fat, vitamins, minerals etc. And it also contains 25 different categories of food products like Beef Products, Vegetable Products, Baked Products etc. Analysis is done on wide range of data present.
# #### Data Source : https://www.kaggle.com/haithemhermessi/usda-national-nutrient-database/download
# ## Data Visualization
# +
#Importing libraries
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
# %matplotlib inline
from mpl_toolkits.mplot3d import Axes3D
import seaborn as sns
import sklearn
# -
#Importing Dataset
df = pd.read_csv("train.csv")
#Reading the dataset
df
# ## Checking and cleaning the dataset
#Checking the shape of the data
df.shape
df.info()
df.head()
df.describe()
# ### Data Examination and cleaning it according to our requirement
#
# This dataset is a part of **Methods and Application of Food Composition Laboratory** which mission is to identify critical food composition needs for researchers, policymakers, food producers etc.
#
# reference:https://www.kaggle.com/haithemhermessi/usda-national-nutrient-database?select=train.csv
#
# In the above data set there are 40 different nutritional values. My goal of the project is to use the widely known food nutritient values to identify the food category. Thus, I have consider only 13 out of the 40 nutrients which I feel is know by a common man. The nutrients I have considered are as follows:
# 1.Energy_kcal
# 2.Protein_g
# 3.Fat_g
# 4.Carb_g
# 5.Sugar_g
# 6.Fiber_g
# 7.VitA_mcg
# 8.VitB6_mg
# 9.VitB12_mcg
# 10.VitE_mg
# based on the above nutrient values I have done my analysis. I have used iloc to pick only the ones I have mentioned.
#
# The dataset contains 6894 entries which is sustainially good for my SVM model.
#Choosing the columns which is easily understood by a common man
df=df.iloc[:,1:14]
df.columns
#Dropping Descrip column as it is was telling the ingridents present in the FoodGroup but not the nutrition in that
df=df.drop('Descrip',axis=1)
df
#Checking for any Null values
df.isnull().sum()
# Visualizing the data and analyzing the amount of nutrients present
plt.rcParams["font.family"] = "Times New Roman"
plt.rcParams.update({'font.size': 12})
df.hist(bins=10, figsize=(20, 15))
plt.grid(axis='y', alpha=0.75)
plt.xlabel('Value',fontsize=15)
plt.ylabel('Frequency',fontsize=15)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.ylabel('Frequency',fontsize=15)
plt.title('Normal Distribution Histogram',fontsize=15)
plt.show()
# **The above graphs shows different amount of nutrient present in the dataset.**
#Checking how many different kinds of FoodGroups are present
df['FoodGroup'].nunique()
# ## Training the datasets
# Assigning the nutrients to X in order to identify the foodgroup
X = df.drop('FoodGroup',axis=1).values
X
# Choosing food groups to be y
y = df['FoodGroup'].values
y
X.shape
y.shape
fig=plt.figure(figsize=(10,5))
plt.hist(y, bins=50)
plt.title('Food Nutrition')
plt.xlabel('FoodGroup')
plt.ylabel('count')
plt.xticks(rotation='vertical')
plt.show()
#Checking different engergy levels of each category of nutrients
for column in df:
q=df['FoodGroup'].values
if column!='FoodGroup':
p=df[column].values
plt.scatter(q,p, s=10)
plt.xlabel('Classification', fontsize=12)
plt.xticks(rotation='vertical')
plt.ylabel(column, fontsize=10)
plt.show()
# Applying Correlation to dataset
corr = df.corr()
# +
# plotting the Heatmap for the correlation
fig, ax = plt.subplots(figsize=(8,5))
sns.heatmap(corr,annot=True,ax=ax)
plt.title('Correlation Matrix for cleaned dataset', fontsize =20)
# -
# Counting the number of products in each Food Group
category_count=pd.DataFrame()
category_count['count']=df['FoodGroup'].value_counts()
category_count['count']
# ## Graph for the Target Column
# Visualizing the FoodGroups
fig, ax=plt.subplots(figsize=(10,5))
sns.barplot(x=category_count.index, y=category_count['count'],ax=ax)
ax.set_ylabel('count',fontsize=15)
ax.set_xlabel('label',fontsize=15)
ax.tick_params(labelsize=15)
ax.set_xticklabels(ax.get_xticklabels(), rotation=90)
# ## Dataset created for testing
#
# After processing the dataset according to our requirements and then analyzing it, we understand that 'Beef Products' have more counts when compared to any other Food category where as the 'Spices and Herbs' have the least count. The data is now ready for application of Machine learning models. The dataset is now 6894 rows × 12 columns which is should be good to predict accurate values.
#
# After creating this dataset we are now ready to go to further analysis
| Notebooks/RightFood_HealthyLife_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup
# +
# Set threshold for when motion is detected in a pixel
thres = 0.1
path = '../data_copies/01_PreprocessedData/01_BudgieFemale_green1/00_Baseline_night/Video/'
# -
import skimage.color
import imageio
import numpy as np
import csv
import os
# # motion_detect function
def motion_detect(file, thres=0.1):
'''
file = path/file.avi to motion detect
thres = proportion (from 0 to 1) of amount of change for a pixel to be counted as motion
'''
output_filename = file[:-4] + '_motion_detect_thres' + str(thres) + '.csv'
# Open writable CSV file
with open(output_filename, 'w') as csvfile:
# CSV writer object
writer = csv.writer(csvfile, quoting=csv.QUOTE_NONNUMERIC)
# header row
writer.writerow(["timestamp (ms)", "Frame #", "Motion (# pixels)"])
# Create a reader object for the video
vid = imageio.get_reader(file, 'ffmpeg')
# Get frames per second
fps = vid.get_meta_data()['fps']
# Iterate through each frame
nums = range(vid.get_length() - 2)
for num in nums:
# Current frame
image = vid.get_data(num)
# Next frame
image_next = vid.get_next_data()
# Transform to greyscale
image_grey = skimage.color.rgb2gray(image)
image_next_grey = skimage.color.rgb2gray(image_next)
# Calculate motion
motion = np.abs(image_next_grey - image_grey)
moved = motion>thres
nPixels_moved = moved.sum()
timestamp = 1000* (num+1)/fps
writer.writerow([timestamp, num+1, nPixels_moved])
# # Run on all video files (set file type)
for filename in os.listdir(path):
if ".mov" in filename.lower(): # .mov files only for Bird 1, else .avi
print(filename)
file = path + filename
motion_detect(file, thres)
| Figure 09d-e - Motion detect all video files in a folder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to Render a Metal Design into Ansys
# ## Prerequisite
# You need to have Ansys installed locally (any version) - at the time of this notebook, Ansys is only supported in Windows.
#
# ## 1. Perform the necessary imports and create a design in Metal
# %load_ext autoreload
# %autoreload 2
import qiskit_metal as metal
from qiskit_metal import designs, MetalGUI
from qiskit_metal import Dict, Headings
from qiskit_metal.renderers.renderer_ansys.ansys_renderer import QAnsysRenderer
QAnsysRenderer.default_options
design = designs.DesignPlanar()
# +
gui = MetalGUI(design)
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
# -
design.variables['cpw_width'] = '15 um'
design.variables['cpw_gap'] = '9 um'
# ### In this notebook, the design consists of 4 qubits and 4 CPWs
# +
# Allow running the same cell here multiple times to overwrite changes
design.overwrite_enabled = True
## Custom options for all the transmons
options = dict(
# Some options we want to modify from the defaults
# (see below for defaults)
pad_width = '425 um',
pocket_height = '650um',
# Adding 4 connectors (see below for defaults)
connection_pads=dict(
a = dict(loc_W=+1,loc_H=-1, pad_width='200um'),
b = dict(loc_W=-1,loc_H=+1, pad_height='30um'),
c = dict(loc_W=-1,loc_H=-1, pad_height='50um')
)
)
## Create 4 transmons
q1 = TransmonPocket(design, 'Q1', options = dict(
pos_x='+2.42251mm', pos_y='+0.0mm', **options))
q2 = TransmonPocket(design, 'Q2', options = dict(
pos_x='+0.0mm', pos_y='-0.95mm', orientation = '270', **options))
q3 = TransmonPocket(design, 'Q3', options = dict(
pos_x='-2.42251mm', pos_y='+0.0mm', orientation = '180', **options))
q4 = TransmonPocket(design, 'Q4', options = dict(
pos_x='+0.0mm', pos_y='+0.95mm', orientation = '90', **options))
from qiskit_metal.qlibrary.interconnects.meandered import RouteMeander
RouteMeander.get_template_options(design)
options = Dict(
lead=Dict(
start_straight='0.2mm',
end_straight='0.2mm'),
trace_gap='9um',
trace_width='15um')
def connect(component_name: str, component1: str, pin1: str, component2: str, pin2: str,
length: str, asymmetry='0 um', flip=False, fillet='90um'):
"""Connect two pins with a CPW."""
myoptions = Dict(
fillet=fillet,
pin_inputs=Dict(
start_pin=Dict(
component=component1,
pin=pin1),
end_pin=Dict(
component=component2,
pin=pin2)),
total_length=length)
myoptions.update(options)
myoptions.meander.asymmetry = asymmetry
myoptions.meander.lead_direction_inverted = 'true' if flip else 'false'
return RouteMeander(design, component_name, myoptions)
asym = 140
cpw1 = connect('cpw1', 'Q1', 'c', 'Q2', 'b', '5.6 mm', f'+{asym}um')
cpw2 = connect('cpw2', 'Q3', 'b', 'Q2', 'c', '5.7 mm', f'-{asym}um', flip=True)
cpw3 = connect('cpw3', 'Q3', 'c', 'Q4', 'b', '5.6 mm', f'+{asym}um')
cpw4 = connect('cpw4', 'Q1', 'b', 'Q4', 'c', '5.7 mm', f'-{asym}um', flip=True)
gui.rebuild()
gui.autoscale()
# -
# ## 2. Render into Ansys HFSS
# #### 2.1 Create a Metal renderer for Ansys HFSS
fourq_hfss = design.renderers.hfss
# #### 2.2 Setup an Ansys project
# To setup the project **manually**, follow these instructions:
# 1. Launch `ANSYS Electronics Desktop yyyy Rx` (from your Windows Start menu).
# 2. Create a new Ansys project by clicking on the `New` icon at the top left. (or open an existing project)
#
# Alternatively, you can **automatically** set up the project by executing the following two cells. Make sure to wait after executing the first cell for Ansys to completely open. Only then execute the second cell.
#
# Note about Ansys version: open_ansys() will look by default for the 2020 R2 version of Ansys. You can easily reroute it to your Ansys of choice by providing the name of the environment variable that contains the path (path_env) or the path itself (path)
fourq_hfss.open_ansys() # this opens Ansys 2020 R2 if present
# fourq_hfss.open_ansys(path_var='ANSYSEM_ROOT211')
# fourq_hfss.open_ansys(path='C:\Program Files\AnsysEM\AnsysEM20.2\Win64')
# fourq_hfss.open_ansys(path='../../../Program Files/AnsysEM/AnsysEM20.2/Win64')
# NOTE: A new project should have automatically opened with the execution of the cell above. If not, uncomment the cell below and execute it. You can also load an existing project by passing the project information to the `connect_ansys()`.
# +
# fourq_hfss.new_ansys_project()
# -
# #### 2.3 Connect the Metal renderer with the Ansys project
# Open either a new or existing design based on default options
fourq_hfss.connect_ansys()
# fourq_hfss.connect_ansys('C:\\project_path\\', 'Project1') # Example of opening a saved project
# You can optionally indicate with the optional parameters whether you intend to open and use a previously saved project. <br>
# Make sure that the saved project contains at least one design, or the method will produce a warning
# #### 2.4 Setup an Ansys HFSS design
# You can either create a new design or select and use an old one.
#
# To **create** a new design **manually**, go to the Ansys GUI and follow these instructions:
# 1. Select the project from the leftmost menu in the Ansys GUI.
# 2. Go into the menu `Project` and select `Insert HFSS Design`.
# 3. Change the HFSS design to either eigenmode or modal by right-clicking on the HFSSdesign1 that just got created inside your project (left panel) and then selecting: `Solution Type...`.
#
# To **create** a new eigenmode design **automatically**, execute the following cell<br>
# The design will be added to the project that was active when the command `fourq_hfss.connect_ansys()` was executed.
# Note: If a design named `HFSSTransmonQubit` already exists in the project, a new design will be created, with the name suffixed with an incremental integer: `HFSSTransmonQubit1`, `HFSSTransmonQubit2`, etc.
fourq_hfss.add_eigenmode_design("HFSSMetalEigenmode")
# To **create** a new driven modal design **automatically**, execute the following cell instead
fourq_hfss.add_drivenmodal_design("HFSSMetalDrivenModal")
# To **select** an existing design, means to activate an Ansys design and connect to it. You can follow one of three ways:
# * re-running the `fourq_hfss.connect_ansys(*with parameters*)`, this time specifying which design to connect to (see section 2.3)
# * manually activating the design from the Ansys GUI. You will find the list of designs in the leftmost panel, and you can activate them with a double click. After this, re-run the `fourq_hfss.connect_ansys()` *without parameters*.
# * use methods: activate_eigenmode_design() or activate_drivenmodal_design(). If the design name exists, it will be added, but no integer will be added to the suffix. If the design name does not exist, then it will be added to the project.
#
# To **add or select** a setup for the active design, you can use activate_drivenmodal_setup() or activate_eigenmode_setup(). If the setup exists, the QRenderer will reference the setup, otherwise, will make a new setup with the name give. If no name given, the default name of "Setup" will be used.
fourq_hfss.activate_eigenmode_design("HFSSMetalEigenmode")
fourq_hfss.activate_eigenmode_setup('SetupNEW')
fourq_hfss.activate_eigenmode_setup()
fourq_hfss.activate_eigenmode_setup('Setup')
fourq_hfss.activate_drivenmodal_design("HFSSMetalDrivenModalNEW")
fourq_hfss.activate_drivenmodal_setup('SetupNEW')
fourq_hfss.activate_drivenmodal_setup('Setup')
# #### 2.5 Render some component from the Metal design
#
# Find below several rendering examples. You can choose to only execute one of them if you are just browsing this notebook.
#
# Notice how we explicitly clear the design before re-rendering. Indeed `render_design()` only adds shapes to the Ansys design. Re-rendering the same shapes will cause violations.
fourq_hfss.render_design([], []) # entire Metal design.
fourq_hfss.clean_active_design()
fourq_hfss.render_design(['Q1'], [('Q1', 'b'), ('Q1', 'c')]) # single qubit with 2 endcaps.
fourq_hfss.clean_active_design()
fourq_hfss.render_design(['Q1', 'cpw1', 'Q2'], [('Q1', 'b'), ('Q2', 'c')]) # 2 qubits and 2 endcaps, one per qubit.
# For Driven-Modal analysis, we can also add terminations. In the example below we render 1 qubit with 1 endcap and 1 port with a 70 Ohm termination.
fourq_hfss.clean_active_design()
fourq_hfss.render_design(['Q2'], [('Q2', 'a')], [('Q2', 'b', '70')])
# In the previous examples, rendering area dimenstions is determined by the size of the selected geometries, with some buffer.
#
# For a more accurate control of the chip size, you need to disable the buffering as below. This will use `design._chips['main']['size']` to determine the rendering area dimensions.
fourq_hfss.clean_active_design()
fourq_hfss.render_design([], [], box_plus_buffer=False)
# You can also modify the chip size directly by updating `design._chips['main']['size']`. Example below.
#
# NOTE: we purposfully make the chip size smaller than the size of the geometry. This will cause a warning to show which will need to be fixed by the user intending to conduct a valid analysis.
fourq_hfss.clean_active_design()
design._chips['main']['size']['size_x'] = '4mm'
fourq_hfss.render_design([], [], box_plus_buffer=False)
# Return back to original size, for the remainder of the notebook
design._chips['main']['size']['size_x'] = '6mm'
# **Finally** disconnect the Metal renderer from the Ansys session.
#
# NOTE: This is needed every time before re-connecting. If you do not disconnect explicitly, you might not be able to close the Ansys GUI later.
fourq_hfss.disconnect_ansys()
# ## 3. Render into Ansys Q3D
# #### 3.1 Create a Metal renderer for Ansys Q3D
fourq_q3d = design.renderers.q3d
# #### 3.2 Setup an Ansys project
# Skip this session if you have already executed cells in 2.2, and you left Ansys open. Indeed, it does not matter which renderer has opened Ansys or created the project. It will suffice to connect the fourq_q3d renderer to the Ansys session (section 3.3).
#
# The following cell will open a new session of Ansys with a brand new project
fourq_q3d.open_ansys() # this opens Ansys 2020 R2 if present
# #### 3.3 Connect the Metal renderer with the Ansys project
fourq_q3d.connect_ansys()
# fourq_q3d.connect_ansys('C:\\project_path\\', 'Project1') # Example of opening a saved project
# You can optionally indicate with the method in the above cell whether you intend to open and use a previously saved project. <br>
# Make sure that the saved project contains at least one design, or the method will produce a warning.
# #### 3.4 Setup an Ansys Q3D design
# You can either create a new design or select and use an old one.
#
# To **create** a new design **manually**, go to the Ansys GUI and follow these instructions:
# 1. Select the project from the leftmost menu in the Ansys GUI.
# 2. Go into the menu `Project` and select `Insert Q3D Extractor Design`.
#
# To **create** a new Q3D design **automatically**, execute the following cell.<br>
# Note: If a design named `Q3dMetalDesign` already exists in the project, a new design will be created, with the name suffixed with an incremental integer: `Q3dMetalDesign1`, `Q3dMetalDesign2`, etc.
fourq_q3d.add_q3d_design("Q3dMetalDesign")
# To **select** an existing design, means to activate an Ansys design and connect to it. You can follow one of three ways:
# * re-running the `fourq_q3d.connect_ansys(*with parameters*)`, this time specifying which design to connect to (see section 2.3)
# * manually activating the design from the Ansys GUI. You will find the list of designs in the leftmost panel, and you can activate them with a double click. After this, re-run the `fourq_q3d.connect_ansys()` *without parameters*.
# * use method: activate_q3d_design(). If the design name exists, it will be added, but no integer will be added to the suffix. If the design name does not exist, then it will be added to the project.
#
# To **add or select** a setup for the active design, you can use activate_q3d_setup(). If the setup exists, the QRenderer will reference the setup, otherwise, will make a new setup with the name give. If no name given, the default name of "Setup" will be used.
# The same command will make new design for Q3dMetalDesignNEW since it is not in the project,
# but will just make design Q3dMetalDesign active.
fourq_q3d.activate_q3d_design("Q3dMetalDesignNEW")
fourq_q3d.activate_q3d_design("Q3dMetalDesign")
fourq_q3d.activate_q3d_setup("SetupNEW")
fourq_q3d.activate_q3d_setup()
fourq_q3d.activate_q3d_setup("Setup")
# #### 3.5 Render some component from the Metal design
#
# Find below several rendering examples. You can choose to only execute one of them if you are just browsing this notebook.
#
# Notice how we explicitly clear the design before re-rendering. Indeed `render_design()` only adds shapes to the Ansys design. Re-rendering the same shapes will cause violations.
fourq_q3d.render_design([], []) # entire Metal design.
fourq_q3d.clean_active_design()
fourq_q3d.render_design(['Q1'], [('Q1', 'b'), ('Q1', 'c')]) # single qubit with 2 endcaps.
fourq_q3d.clean_active_design()
fourq_q3d.render_design(['Q1', 'cpw1', 'Q2'], [('Q1', 'b'), ('Q2', 'c')]) # 2 qubits and 2 endcaps, one per qubit.
# In the previous examples, rendering area dimenstions is determined by the size of the selected geometries, with some padding.
#
# For a more accurate control of the chip size, you need to disable the buffering as below. This will use `design._chips['main']['size']` to determine the rendering area dimensions.
fourq_q3d.clean_active_design()
fourq_q3d.render_design([], [], box_plus_buffer=False)
# You can also modify the chip size directly by updating `design._chips['main']['size']`. Example below:
fourq_q3d.clean_active_design()
design._chips['main']['size']['size_y'] = '4mm'
fourq_q3d.render_design([], [], box_plus_buffer=False)
# Return back to original size, for the remainder of the notebook
design._chips['main']['size']['size_y'] = '6mm'
# **Finally** disconnect the Metal renderer from the Ansys session. You will not be able to close Ansys without executing this.
fourq_q3d.disconnect_ansys()
# ## References - Miscellaneous pyEPR/Ansys commands
# The following commands are for reference only to better understand how the backend code works. They're not meant to be run directly in this notebook as part of the tutorial.
# import pyEPR as epr
#
# Connect to Ansys directly from notebook:
#
# pinfo = epr.ProjectInfo(project_path = None,
# project_name = None,
# design_name = None)
# modeler = pinfo.design.modeler
#
# Access methods within HfssDesign class in pyEPR:
#
# epr.ansys.HfssDesign.create_dm_setup
# epr.ansys.HfssDesign.create_q3d_setup
#
# Get project and design names:
#
# pinfo.project_name
# design._design.GetName()
#
# Filter qgeometry table:
#
# full_table = design.qgeometry.tables['poly']
# mask = full_table['subtract'] == False
# table = full_table[mask]
#
# Draw centered rectangles:
#
# bigsquare = modeler.draw_rect_center([0, 0, 0], x_size=8, y_size=8, name='bigsquare')
# topright = modeler.draw_rect_center([2, 2, 0], x_size=2, y_size=2, name='topright')
#
# Subtracting shapes:
#
# modeler.subtract('bigsquare', ['topright'])
#
# Draw centered box:
#
# modeler.draw_box_center([0, 0, 0], [1, 2, 3])
#
# Draw closed polygon:
#
# trianglepts = [[-1, 5, 0], [1, 5, 0], [0, 7, 0]]
# modeler.draw_polyline(trianglepts, closed=True)
#
# Draw polyline:
#
# smallpts = [[2.85, 0, 0], [3.15, 0, 0]]
# modeler.draw_polyline(smallpts, closed=False)
#
# Sweep one polyline with another:
#
# modeler._sweep_along_path('Polyline8', 'Polyline7')
| tutorials/5 All QRenderers/5.1 Ansys/Ansys QRenderer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # RNA World Hypothesis
#
# RNA is a simpler cousin of DNA. As you may know, RNA is widely thought to be the first self replicating life-form to arise perhaps around 4 billion years ago. One of the strongest arguments for this theory is that RNA is able to carry information in its nucleotides like DNA, and like protein, it is able to adopt higher order structures to catalyze reactions, such as self replication. So it is likely, and there is growing evidence that this is the case, that the first form of replicating life was RNA. And because of this dual property of RNA as a hereditary information vessel as well as a structural/functional element we can use RNA molecules to build very nice population models.
#
# So in this section, we'll be walking you through building genetic populations, simulating their evolution, and using statistics and other mathematical tools for understanding key properties of populations.
# +
from IPython.display import HTML
# Youtube
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/K1xnYFCZ9Yg" frameborder="0" allow="accelerometer; autoplay; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>')
# +
#HTML('<iframe width="784" height="441" src="https://scaleofuniverse.com/" /iframe>')
# + [markdown] slideshow={"slide_type": "-"}
# ## Population Evolution in *an* RNA World
#
# In order to study the evolution of a population, we first need a model of a population. And even before that, we need to define what we mean by *population*. Populations can be defined on many levels and with many diffferent criteria. For our purposes, we will simply say that a population is a set of individuals sharing a common environment. And because this is population *genetics* we can think of individuals as entities comprising of specific genes or chromosomes.
#
# So where do we get a population from? As you may have discussed in previous workshops, there are very large datasets containing sequencing information from different populations. So we could download one of these datasets and perform some analysis on it. But I find this can be dry and tedious. So why download data when we can simply create our own?
#
# In this section we're going to be creating and studying our own "artificial" populations to illustrate some important population genetics concepts and methodologies. Not only will this help you learn population genetics, but you will get a lot more programming practice than if we were to simply parse data files and go from there.
#
# More specifically, we're going to build our own RNA world.
#
# ### Building an RNA population
#
# As we saw earlier, RNA has the nice property of posessing a strong mapping between information carrying (sequence) and function (structure). This is analogous to what is known in evolutionary terms as a genotype and a phenotype. With these properties, we have everything we need to model a population, and simulate its evolution.
#
# #### RNA sequence-structure
#
# We can think of the genotype as a sequence $s$ consisting of letters/nucleotides from the alphabet $\{U,A,C,G\}$. The corresponding phenotype $\omega$ is the secondary structure of $s$ which can be thought of as a pairing between nucleotides in the primary sequence that give rise to a 2D architecture. Because it has been shown that the function of many biomolecules, including RNA, is driven by structure this gives us a good proxy for phenotype.
#
# Below is an example of what an RNA secondary structure, or pairing, looks like.
# + slideshow={"slide_type": "-"}
### 1
from IPython.display import Image
#This will load an image of an RNA secondary structure
Image(url='https://viennarna.github.io/forgi/_images/1y26_ss.png')
# +
from IPython.display import HTML
# Youtube
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/JQByjprj_mA" frameborder="0" allow="accelerometer; autoplay; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>')
#import matplotlib.pyplot as plt
#import forgi.visual.mplotlib as fvm
#import forgi
#cg = forgi.load_rna("1y26.fx", allow_many=False)
#fvm.plot_rna(cg, text_kwargs={"fontweight":"black"}, lighten=0.7,
# backbone_kwargs={"linewidth":3})
#plt.show()
# -
# As you can see, unpaired positions are forming loop-like structures, and paired positions are forming stem-like structures. It is this spatial arrangement of nucleotides that drives RNA's function. Therefore, another sequence that adopts a similar shape, is likely to behave in a similar manner. Another thing to notice is that, although in reality this is often not the case, in general we only allow pairs between $\{C,G\}$ and $\{A, U\}$ nucleotides, most modern approaches allow for non-canonical pairings and you will find some examples of this in the above structure.
#
# *How do we go from a sequence to a structure?*
#
# So a secondary structure is just a list of pairings between positions. How do we get the optimal pairing?
#
# The algorithm we're going to be using in our simulations is known as the Nussinov Algorithm. The Nussinov algorithm is one of the first and simplest attempts at predicting RNA structure. Because bonds tend to stabilize RNA, the algorithm tries to maximize the number of pairs in the structure and return that as its solution. Current approaches achieve more accurate solutions by using energy models based one experimental values to then obtain a structure that minimizes free energy. But since we're not really concerned with the accuracy of our predictions, Nussinov is a good entry point. Furthermore, the main algorithmic concepts are the same between Nussinov and state of the art RNA structure prediction algorithms. I implemented the algorithm in a separate file called `fold.py` that we can import and use its functions. I'm not going to go into detail here on how the algorithm works because it is beyond the scope of this workshop but there is a bonus exercise at the end if you're curious.
#
# You can predict a secondary structure by calling `nussinov()` with a sequence string and it will return a tuple in the form `(structure, pairs)`.
# + slideshow={"slide_type": "-"}
### 2
import numpy as np
from fold import nussinov #Codes by <NAME> (https://github.com/cgoliver)
sequence_to_fold = "ACCCGAUGUUAUAUAUACCU"
struc = nussinov(sequence_to_fold)
print(">test") #creates the structure of a .fx file for "forna"
print(sequence_to_fold)
print(struc[0])
#Check the molecule at: http://rna.tbi.univie.ac.at/forna/
# Paste the text below in the webpage and see the structure
#>test
#ACCCGAUGUUAUAUAUACCU
#(...(..(((....).))))
# -
# You will see a funny dot-bracket string in the output. This is a representation of the structure of an RNA. Quite simply, a matching parir of parentheses (open and close) correspond to the nucleotides at those positions being paried. Whereas, a dot means that that position is unpaired in the structure. Feel free to play around with the input sequence to get a better understanding of the notation.
#
# If you want to visually check the sequence, go to: [forna](http://rna.tbi.univie.ac.at/forna/forna.html) and copy the text above with the sequence and its structure in the **Add Molecule** button. The webpage is embedded below.
#
# So that's enough about RNA structure prediction. Let's move on to building our populations.
HTML('<iframe width="784" height="441" src="http://rna.tbi.univie.ac.at/forna/forna.html" /iframe>')
print(np.random.choice(5, 3, p=[0.1, 0, 0.6, 0.3, 0],replace=True))
# + [markdown] slideshow={"slide_type": "-"}
# ### Fitness of a sequence: Target Structure
#
# Now that we have a good way of getting a phenotype (secondary structure), we need a way to evaluate the fitness of that phenotype. If we think in real life terms, fitness is the ability of a genotype to replicate into the next generation. If you have a gene carrying a mutation that causes some kind of disease, your fitness is decreased and you have a lower chance of contributing offspring to the next generation. On a molecular level the same concept applies. A molecule needs to accomplish a certain function, i.e. bind to some other molecule or send some kind of signal. And as we've seen before, the most important factor that determines how well it can carry out this function is its structure. So we can imagine that a certain structure, we can call this a 'target' structure, is required in order to accomplish a certain function. So a sequence that folds correctly to a target structure is seen as having a greater fitness than one that does not. Since we've encoded structures as simple dot-bracket strings, we can easily compare structures and thus evaluate the fitness between a given structure and the target, or 'correct' structure.
#
# There are many ways to compare structures $w_{1}$ and $w_{2}$, but we're going to use one of the simplest ways, which is base-pair distance. This is just the number of pairs in $w_{1}$ that are not in $w_{2}$. Again, this is beyond the scope of this workshop so I'll just give you the code for it and if you would like to know more you can ask me.
# +
### 3
#ss_to_bp() and bp_distance() by <NAME>.
def ss_to_bp(ss):
bps = set()
l = []
for i, x in enumerate(ss):
if x == '(':
l.append(i)
elif x == ')':
bps.add((l.pop(), i))
return bps
def bp_distance(w1, w2):
"""
return base pair distance between structures w1 and w1.
w1 and w1 are lists of tuples representing pairing indices.
"""
return len(set(w1).symmetric_difference(set(w2)))
#let's fold two sequences
w1 = nussinov("CCAAAAGG")
w2 = nussinov("ACAAAAGA")
print(w1)
print(w2)
#give the list of pairs to bp_distance and see what the distance is.
print(bp_distance(w1[-1], w2[-1]))
# -
# ## Defining a cell
#
# The cell we will define here is a simple organism with two copies of an RNA gene, each with its own structure. This simple definition of a cell will help us create populations to play around in our evolutionary reactor.
# +
### 4
class Cell:
def __init__(self, seq_1, struc_1, seq_2, struc_2):
self.sequence_1 = seq_1
self.sequence_2 = seq_2
self.structure_1 = struc_1
self.structure_2 = struc_2
#for now just try initializing a Cell with made up sequences and structures
cell = Cell("AACCCCUU", "((.....))", "GGAAAACA", "(....).")
print(cell.sequence_1, cell.structure_2, cell.sequence_1, cell.structure_2)
# -
# ## Populations of Cells
#
# Now we've defined a 'Cell'. Since a population is a collection of individuals our populations will naturally consist of **lists** of 'Cell' objects, each with their own sequences. Here we initialize all the Cells with random sequences and add them to the 'population' list.
# +
### 5
import random
def populate(target, pop_size):
'''Creates a population of cells (pop_size) with a number of random RNA nucleotides (AUCG)
matching the length of the target structure'''
population = []
for i in range(pop_size):
#get a random sequence to start with
sequence = "".join([random.choice("AUCG") for _ in range(len(target))])
#use nussinov to get the secondary structure for the sequence
structure = nussinov(sequence)
#add a new Cell object to the population list
new_cell = Cell(sequence, structure, sequence, structure)
new_cell.id = i
new_cell.parent = i
population.append(new_cell)
return population
# -
# Try creating a new population and printing the first 10 sequences and structures (in dot-bracket) on the first chromosome!
# +
### 6
target = "(((....)))"
pop = populate(target, pop_size=300)
for p in pop[:10]:
print(p.id, p.sequence_1, p.structure_1[0], p.sequence_2, p.structure_2[0])
for p in pop[-10:]:#for p in pop[:10]:#for p in pop[-10:]:#for p in pop[:10]:
print(p.id, p.sequence_1, p.structure_1[0], p.sequence_2, p.structure_2[0])
# -
# ## The Fitness of a Cell
#
#
# Once we are able to create populations of cells, we need a way of asssessing their individual fitness. In our model, a *Cell* is an object that contains two sequences of RNA, something analogous to having two copies of a gene in each chromosome.
#
# We could simply loop through each *Cell* in the population and check the base pair distance to the target structure we defined. However, this approach of using base-pair distance is not the best for defining fitness. There are two reasons for this:
#
# 1. We want fitness to represent a *probability* that a cell will reproduce (pass its genes to the next generation), and base pair distance is, in our case, an integer value.
# 2. We want this probability to be a *relative* measure of fitness. That is, we want the fitness to be proportional to how good a cell is with respect to all others in the population. This touches on an important principle in evolution where we only need to be 'better' than the rest of the population (the competition) and not good in some absolute measure. For instance, if you and I are being chased by a predator. In order to survive, I only need to be faster than you, and not necessarily some absolute level of fitness.
#
# In order to get a probability (number between 0 and 1) we use the following equation to define the fitness of a structure $\omega$ on a target structure $T$:
#
# $$P(\omega, T) = N^{-1} exp(\frac{-\beta \texttt{dist}(\omega, T)}{\texttt{len}(\omega)})$$
#
# $$N = \sum_{i \in Pop}{P(\omega_i, T})$$
#
# Here, the $N$ is what gives us the 'relative' measure because we divide the fitness of the Cell by the sum of the fitness of every other Cell.
#
# Let's take a quick look at how this function behaves if we plot different base pair distance values.
#
# What is the effect of the parameter $\beta$? Try plotting the same function but with different values of $\beta$.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import math
#import seaborn as sns
target_length = 50
beta = -3
plt.plot([math.exp(beta * (bp_dist / float(target_length))) for bp_dist in range(target_length)])
plt.xlabel("Base pair distance to target structure")
plt.ylabel("P(w, T)")
# -
# As you can see, it's a very simple function that evaluates to 1 (highest fitness) if the base pair distance is 0, and decreases as the structures get further and further away from the target. I didn't include the $N$ in the plotting as it will be a bit more annoying to compute, but it is simply a scaling factor so the shape and main idea won't be different.
#
# Now we can use this function to get a fitness value for each Cell in our population.
# +
### 7
def compute_fitness(population, target, beta=-3):
"""
Assigns a fitness and bp_distance value to each cell in the population.
"""
#store the fitness values of each cell
tot = []
#iterate through each cell
for cell in population:
#calculate the bp_distance of each chromosome using the cell's structure
bp_distance_1 = bp_distance(cell.structure_1[-1], ss_to_bp(target))
bp_distance_2 = bp_distance(cell.structure_2[-1], ss_to_bp(target))
#use the bp_distances and the above fitness equation to calculate the fitness of each chromosome
fitness_1 = math.exp((beta * bp_distance_1 / float(len(cell.sequence_1))))
fitness_2 = math.exp((beta * bp_distance_2 / float(len(cell.sequence_2))))
#get the fitness of the whole cell by multiplying the fitnesses of each chromosome
cell.fitness = fitness_1 * fitness_2
#store the bp_distance of each chromosome.
cell.bp_distance_1 = bp_distance_1
cell.bp_distance_2 = bp_distance_2
#add the cell's fitness value to the list of all fitness values (used for normalization later)
tot.append(cell.fitness)
#normalization factor is sum of all fitness values in population
norm = np.sum(tot)
#divide all fitness values by the normalization factor.
for cell in population:
cell.fitness = cell.fitness / norm
return None
compute_fitness(pop, target)
for cell in pop[:10]:
print(cell.fitness, cell.bp_distance_1, cell.bp_distance_2)
# -
# ## Introducing diversity: Mutations
#
# Evolution would go nowhere without random mutations. While mutations are technically just random errors in the copying of genetic material, they are essential in the process of evolution. This is because they introduce novel diversity to populatons, which with a low frequency can be beneficial. And when a beneficial mutation arises (i.e. a mutation that increases fitness, or replication probability) it quickly takes over the population and the populatioin as a whole has a higher fitness.
#
# Implementing mutations in our model will be quite straightforward. Since mutations happen at the genotype/sequence level, we simply have to iterate through our strings of nucleotides (sequences) and randomly introduce changes.
# +
def mutate(sequence, mutation_rate=0.001):
"""Takes a sequence and mutates bases with probability mutation_rate"""
#start an empty string to store the mutated sequence
new_sequence = ""
#boolean storing whether or not the sequence got mutated
mutated = False
#go through every bp in the sequence
for bp in sequence:
#generate a random number between 0 and 1
r = random.random()
#if r is below mutation rate, introduce a mutation
if r < mutation_rate:
#add a randomly sampled nucleotide to the new sequence
new_sequence = new_sequence + random.choice("aucg")
mutated = True
else:
#if the mutation condition did not get met, copy the current bp to the new sequence
new_sequence = new_sequence + bp
return (new_sequence, mutated)
sequence_to_mutate = 'AAAAGGAGUGUGUAUGU'
print(sequence_to_mutate)
print(mutate(sequence_to_mutate, mutation_rate=0.5))
# -
# ## Selection
#
# The final process in this evolution model is selection. Once you have populations with a diverse range of fitnesses, we need to select the fittest individuals and let them replicate and contribute offspring to the next generation. In real populations this is just the process of reproduction. If you're fit enough you will be likely to reproduce more than another individual who is not as well suited to the environment.
#
# In order to represent this process in our model, we will use the fitness values that we assigned to each Cell earlier and use that to select replicating Cells. This is equivalent to sampling from a population with the sampling being weighted by the fitness of each Cell. Thankfully, `numpy.random.choice` comes to the rescue here. Once we have sampled enough Cells to build our next generation, we introduce mutations and compute the fitness values of the new generation.
# +
def selection(population, target, mutation_rate=0.001, beta=-3):
"""
Returns a new population with offspring of the input population
"""
#select the sequences that will be 'parents' and contribute to the next generation
parents = np.random.choice(population, len(population), p=[rna.fitness for rna in population], replace=True)
#build the next generation using the parents list
next_generation = []
for i, p in enumerate(parents):
new_cell = Cell(p.sequence_1, p.structure_1, p.sequence_2, p.structure_2)
new_cell.id = i
new_cell.parent = p.id
next_generation.append(new_cell)
#introduce mutations in next_generation sequeneces and re-fold when a mutation occurs
for rna in next_generation:
mutated_sequence_1, mutated_1 = mutate(rna.sequence_1, mutation_rate=mutation_rate)
mutated_sequence_2, mutated_2 = mutate(rna.sequence_2, mutation_rate=mutation_rate)
if mutated_1:
rna.sequence_1 = mutated_sequence_1
rna.structure_1 = nussinov(mutated_sequence_1)
if mutated_2:
rna.sequence_2 = mutated_sequence_2
rna.structure_2 = nussinov(mutated_sequence_2)
else:
continue
#update fitness values for the new generation
compute_fitness(next_generation, target, beta=beta)
return next_generation
next_gen = selection(pop, target)
for cell in next_gen[:10]:
print(cell.sequence_1)
# -
# ## Gathering information on our populations
#
# Here we simply store some statistics (in a dictionary) on the population at each generation such as the average base pair distance and the average fitness of the populations. No coding to do here, it's not a very interesting function but feel free to give it a look.
def record_stats(pop, population_stats):
"""
Takes a population list and a dictionary and updates it with stats on the population.
"""
generation_bp_distance_1 = [rna.bp_distance_1 for rna in pop]
generation_bp_distance_2 = [rna.bp_distance_2 for rna in pop]
mean_bp_distance_1 = np.mean(generation_bp_distance_1)
mean_bp_distance_2 = np.mean(generation_bp_distance_2)
mean_fitness = np.mean([rna.fitness for rna in pop])
population_stats.setdefault('mean_bp_distance_1', []).append(mean_bp_distance_1)
population_stats.setdefault('mean_bp_distance_2', []).append(mean_bp_distance_2)
population_stats.setdefault('mean_fitness', []).append(mean_fitness)
return None
# ## And finally.... evolution
#
# We can put all the above parts together in a simple function that does the following:
#
# 1. start a new population and compute its fitness
# 2. repeat the following for the desired number of generations:
# 1. record statistics on population
# 2. perform selection+mutation
# 3. store new population
#
# And that's it! We have an evolutionary reactor!
def evolve(target, generations=20, pop_size=100, mutation_rate=0.001, beta=-2):
"""
Takes target structure and sets up initial population, performs selection and iterates for desired generations.
"""
#store list of all populations throughotu generations [[cells from generation 1], [cells from gen. 2]...]
populations = []
#start a dictionary that will hold some stats on the populations.
population_stats = {}
#get a starting population
initial_population = populate(target, pop_size=pop_size)
#compute fitness of initial population
compute_fitness(initial_population, target)
#set current_generation to initial population.
current_generation = initial_population
#iterate the selection process over the desired number of generations
for i in range(generations):
#let's get some stats on the structures in the populations
record_stats(current_generation, population_stats)
#add the current generation to our list of populations.
populations.append(current_generation)
#select the next generation
new_gen = selection(current_generation, target, mutation_rate=mutation_rate, beta=beta)
#set current generation to be the generation we just obtained.
current_generation = new_gen
return (populations, population_stats)
# Try a run of the `evolve()` function.
# +
target = "(((....)))"
pops, pops_stats = evolve(target, generations=20, pop_size=300, mutation_rate=0.001, beta=-2)
#Print the first 10 sequences of the population
for p in pop[:10]:
print(p.id, p.sequence_1, p.structure_1[0], p.sequence_2, p.structure_2[0])
#Print the last 10 sequences of the population
for p in pop[-10:]:
print(p.id, p.sequence_1, p.structure_1[0], p.sequence_2, p.structure_2[0])
# -
# Let's see if it actually worked by plotting the average base pair distance as a function of generations for both genes in each cell. We should expect a gradual decrease as the populations get closer to the target structure.
# +
def evo_plot(pops_stats):
"""
Plot base pair distance for each chromosome over generations.
"""
plt.figure('Mean base pair Distance',figsize=(10,5))
for m in ['mean_bp_distance_1', 'mean_bp_distance_2']:
plt.plot(pops_stats[m], label=m)
plt.legend()
plt.xlabel("Generations")
plt.ylabel("Mean Base Pair Distance")
evo_plot(pops_stats)
# -
# Let's see the structure of random cells from each generation. Run the code below and copy the output in the RNA folding webpage. Compare the base-pair distance plot with the structures.
#
# Questions:
# - Do you notice some simmilarities from a particular generation onwards? Compare your observations to the plot with the Mean Base Pair Distance.
# - What could trigger another evolutionary jump?
#
# 
#
from IPython.display import HTML
HTML('<iframe width="784" height="441" src="http://rna.tbi.univie.ac.at/forna/forna.html" /iframe>')
#Select a random RNA sequence from each generation to check its folding structure
from random import randrange
#print(randrange(999))
generations=20
pop_size=300
#Print some random cells from each generation
#pops[generation][cell in that generation].{quality to retrieve}
for gen in range(0,generations):
cid=randrange(pop_size)
print('>Gen'+np.str(gen+1)+'_Cell_'+np.str(pops[gen][cid].id)+'')
print(pops[gen][2].sequence_1)
print(''+np.str(pops[gen][2].structure_1[0])+'\n')
# You should see a nice drop in base pair distance! Another way of visualizing this is by plotting a histogram of the base pair distance of all Cells in the initial population versus the final population.
def bp_distance_distributions(pops):
"""
Plots histograms of base pair distance in initial and final populations.
"""
#plot bp_distance_1 for rnas in first population
g = sns.distplot([rna.bp_distance_1 for rna in pops[0]], label='initial population')
#plot bp_distance_1 for rnas in first population
g = sns.distplot([rna.bp_distance_1 for rna in pops[-1]], label='final population')
g.set(xlabel='Mean Base Pair Distance')
g.legend()
bp_distance_distributions(pops)
# ## Introducing mating to the model
#
# The populations we generated evolved asexually. This means that individuals do not mate or exchange genetic information. So to make our simulation a bit more interesting let's let the Cells mate. This is going to require a few small changes in the `selection()` function. Previously, when we selected sequences to go into the next generation we just let them provide one offspring which was a copy of itself and introduced mutations. Now instead of choosing one Cell at a time, we will randomly choose two 'parents' that will mate. When they mate, each parent will contribute one of its chromosomes to the child. We'll repeat this process until we have filled the next generation.
# +
def selection_with_mating(population, target, mutation_rate=0.001, beta=-2):
next_generation = []
counter = 0
while len(next_generation) < len(population):
#select two parents based on their fitness
parents_pair = np.random.choice(population, 2, p=[rna.fitness for rna in population], replace=False)
#take the sequence and structure from the first parent's first chromosome and give it to the child
child_chrom_1 = (parents_pair[0].sequence_1, parents_pair[0].structure_1)
#do the same for the child's second chromosome and the second parent.
child_chrom_2 = (parents_pair[1].sequence_2, parents_pair[1].structure_2)
#initialize the new child Cell with the new chromosomes.
child_cell = Cell(child_chrom_1[0], child_chrom_1[1], child_chrom_2[0], child_chrom_2[1])
#give the child and id and store who its parents are
child_cell.id = counter
child_cell.parent_1 = parents_pair[0].id
child_cell.parent_2 = parents_pair[1].id
#add the child to the new generation
next_generation.append(child_cell)
counter = counter + 1
#introduce mutations in next_generation sequeneces and re-fold when a mutation occurs (same as before)
for rna in next_generation:
mutated_sequence_1, mutated_1 = mutate(rna.sequence_1, mutation_rate=mutation_rate)
mutated_sequence_2, mutated_2 = mutate(rna.sequence_2, mutation_rate=mutation_rate)
if mutated_1:
rna.sequence_1 = mutated_sequence_1
rna.structure_1 = nussinov(mutated_sequence_1)
if mutated_2:
rna.sequence_2 = mutated_sequence_2
rna.structure_2 = nussinov(mutated_sequence_2)
else:
continue
#update fitness values for the new generation
compute_fitness(next_generation, target, beta=beta)
return next_generation
#run a small test to make sure it works
next_gen = selection_with_mating(pop, target)
for cell in next_gen[:10]:
print(cell.sequence_1)
# -
# Now we just have to update our `evolution()` function to call the new `selection_with_mating()` function.
def evolve_with_mating(target, generations=10, pop_size=100, mutation_rate=0.001, beta=-2):
populations = []
population_stats = {}
initial_population = populate(target, pop_size=pop_size)
compute_fitness(initial_population, target)
current_generation = initial_population
#iterate the selection process over the desired number of generations
for i in range(generations):
#let's get some stats on the structures in the populations
record_stats(current_generation, population_stats)
#add the current generation to our list of populations.
populations.append(current_generation)
#select the next generation, but this time with mutations
new_gen = selection_with_mating(current_generation, target, mutation_rate=mutation_rate, beta=beta)
current_generation = new_gen
return (populations, population_stats)
# Try out the new evolution model!
# +
pops_mating, pops_stats_mating = evolve_with_mating("(((....)))", generations=20, pop_size=1000, beta=-2)
evo_plot(pops_stats_mating)
# -
#Select a random RNA sequence from each generation to check its folding structure
from random import randrange
#print(randrange(999))
generations=20
pop_size=1000
#Print some random cells from each generation
#pops[generation][cell in that generation].{quality to retrieve}
for gen in range(0,generations):
cid=randrange(pop_size)
print('>Gen'+np.str(gen+1)+'_Cell_'+np.str(pops_mating[gen][cid].id)+'')
print(pops_mating[gen][2].sequence_1)
print(''+np.str(pops_mating[gen][2].structure_1[0])+'\n')
# # Acknowledgements
#
# The computational codes of this notebook were originally created by [<NAME>](https://github.com/cgoliver), and adapted by <NAME> for ASTRO200.
| content/07/2/.ipynb_checkpoints/RNA World-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import pickle
import os
import pandas
import pynumdiff
import scipy.fftpack
from IPython.display import display,SVG
import figurefirst
fifi = figurefirst
import copy
# -
import run_pareto_plot
# +
def get_gamma(dt, freq, timeseries_length=None):
log_gamma = -1.59*np.log(freq) -0.71*np.log(dt) - 5.5
return np.exp(log_gamma)
if 0:
try:
if dt == 0.1:
log_g = np.log(freq)*(-1.5) -4
if dt == 0.01:
log_g = np.log(freq)*(-1.5) -1.8
if dt == 0.001:
log_g = np.log(freq)*(-1.5) -0.5
return np.exp(log_g)
except:
log_gs = []
for f in freq:
if dt == 0.1:
log_gs.append(np.log(freq)*(-1.5) -4)
if dt == 0.01:
log_gs.append(np.log(freq)*(-1.5) -1.8)
if dt == 0.001:
log_gs.append(np.log(freq)*(-1.5) -0.5)
return np.exp(np.array(log_gs))
# -
def load_data(fname):
f = open(fname, 'rb')
data = pickle.load(f)
f.close()
return data
# +
def get_goldilocks_gamma(data):
rmses = data['rmses_gamma']
errcorrs = data['errcorrs_gamma']
threshold = 0.25
while len(np.where(errcorrs<threshold)[0]) < 1:
threshold += 0.05
idx_errcorr_okay = np.where(errcorrs<threshold)
idx_opt = idx_errcorr_okay[0][np.argmin(rmses[idx_errcorr_okay])]
opt_rmse = rmses[idx_opt]
opt_errcorr = errcorrs[idx_opt]
return data['metadata']['gammas'][idx_opt], opt_rmse, opt_errcorr, idx_opt
def plot_direct_goldilocks_gamma(ax, data, color):
goldgamma, opt_rmse, opt_errcorr, idx_opt = get_goldilocks_gamma(data)
print('goldilocks gamma: ', goldgamma)
print('goldilocks rmse and errcorr: ', opt_rmse, opt_errcorr)
print('opt params: ', data['metadata']['params'][idx_opt])
ax.plot(opt_rmse, opt_errcorr, '*', color=color, markersize=20, markeredgecolor='black')
#ax.set_xlim(1e-1, 1e2)
# -
def plot_xdots(ax, data, x, dxdt_truth, t, color):
idx_best = np.argmin(data['rmses'])
params = run_pareto_plot.get_params_for_method('savgoldiff', 'linear_model')
print('best params: ', params[idx_best])
x_hat, xdot_hat = pynumdiff.linear_model.savgoldiff(x, dt, params[idx_best])
ax.plot(t, xdot_hat, color='gray', linewidth=4, zorder=-10, alpha=0.5)
goldgamma, _, _, goldidx = get_goldilocks_gamma(data)
params = data['metadata']['params']
print('goldilocks gamma params: ', params[goldidx])
x_hat, xdot_hat = pynumdiff.linear_model.savgoldiff(x, dt, params[goldidx])
ax.plot(t, xdot_hat, color=color, linewidth=1, zorder=-5)
ax.plot(t, dxdt_truth, '--', color='black', zorder=-1, linewidth=0.5)
ax.set_rasterization_zorder(0)
def plot_example(example, x, x_truth, dxdt_truth, t, color, data, xlimits, ylimits, xticks, yticks):
layout = fifi.svg_to_axes.FigureLayout(figure_layout, autogenlayers=True,
make_mplfigures=True, hide_layers=[])
ax_pareto = layout.axes[(example, 'pareto')]
ax_pos = layout.axes[(example, 'sine')]
ax_vel = layout.axes[(example, 'vel')]
ax_pos.plot(t, x, '.', color='blue', zorder=-10, markersize=2)
ax_pos.set_rasterization_zorder(0)
ax_pos.plot(t, x_truth, '--', color='black', linewidth=0.5)
plot_xdots(ax_vel, data, x, dxdt_truth, t, color)
ax_pareto.plot((data['rmses']), data['errcorrs'], '.', color='gray', zorder=-10, markersize=2)
try:
ax_pareto.set_rasterization_zorder(0)
except:
print('could not rasterize')
ax_pareto.plot((data['rmses_gamma']), data['errcorrs_gamma'], '.', color=color, zorder=1, markersize=2)
ax_pareto.plot((data['rmses_gamma']), data['errcorrs_gamma'], '-', color=color)
plot_direct_goldilocks_gamma(ax_pareto, data, color)
ax_pos.set_xlim(xlimits['pos'][0], xlimits['pos'][-1])
ax_pos.set_ylim(ylimits['pos'][0], ylimits['pos'][-1])
ax_vel.set_xlim(xlimits['vel'][0], xlimits['vel'][-1])
ax_vel.set_ylim(ylimits['vel'][0], ylimits['vel'][-1])
if example == 'freq_1':
fifi.mpl_functions.adjust_spines(ax_pos, ['left'],
xticks=xticks['pos'],
yticks=yticks['pos'],
tick_length=2.5,
spine_locations={'left': 4, 'bottom': 4})
fifi.mpl_functions.adjust_spines(ax_vel, ['left', 'bottom'],
xticks=xticks['vel'],
yticks=yticks['vel'],
tick_length=2.5,
spine_locations={'left': 4, 'bottom': 4})
else:
fifi.mpl_functions.adjust_spines(ax_pos, ['left'],
xticks=xticks['pos'],
yticks=yticks['pos'],
tick_length=2.5,
spine_locations={'left': 4, 'bottom': 4})
fifi.mpl_functions.adjust_spines(ax_vel, ['left', 'bottom'],
xticks=xticks['vel'],
yticks=yticks['vel'],
tick_length=2.5,
spine_locations={'left': 4, 'bottom': 4})
exp = int(np.log10(yticks['vel'][-1]))
ax_vel.set_yticklabels(['$-10^{'+str(exp)+'}$', '$0$', '$10^{'+str(exp)+'}$'])
ax_pareto.set_xscale('log')
ax_pareto.minorticks_off()
ax_pareto.set_xlim(xlimits['pareto'][0], xlimits['pareto'][1])
ax_pareto.set_ylim(ylimits['pareto'][0], ylimits['pareto'][1])
if example == 'freq_1':
fifi.mpl_functions.adjust_spines(ax_pareto, ['left', 'bottom'],
xticks=xticks['pareto'],
yticks=yticks['pareto'],
tick_length=2.5,
spine_locations={'left': 4, 'bottom': 4})
else:
fifi.mpl_functions.adjust_spines(ax_pareto, ['bottom'],
xticks=xticks['pareto'],
tick_length=2.5,
spine_locations={'bottom': 4})
fifi.mpl_functions.set_fontsize(ax_pareto, 6)
layout.append_figure_to_layer(layout.figures[example], example, cleartarget=True)
layout.write_svg(figure_layout)
figure_layout = 'fig_2.svg'
# +
# define problem
example = 'freq_2'
dt = 0.01
noise = 0.5
timeseries_length = 4
problem = 'sine'
freq = 1
if timeseries_length < np.pi/freq:
raise ValueError()
if dt > 1/freq/2.:
raise ValueError()
read_existing = True
simdt = 0.0001
color = 'dodgerblue'
# define method
method_parent = 'linear_model'
method = 'savgoldiff'
# define limits
xlimits = {'pos': [0,4],
'vel': [0,4],
'pareto': [5e-1, 1e1]}
ylimits = {'pos': [-0.2,2.2],
'vel': [-10, 10],
'pareto': [-.1, 1.1]}
xticks = { 'pos': [0,2,4],
'vel': [0,2,4],
'pareto': [5e-1, 1e0, 1e1]}
yticks = { 'pos': [0, 1, 2],
'vel': [-10, 0, 10],
'pareto': [0, 1]}
r = pynumdiff.utils.simulate.sine(timeseries_length=timeseries_length,
noise_parameters=[0, noise],
dt=dt,
frequencies=[freq])
x, x_truth, dxdt_truth, _ = r
t = np.arange(0, timeseries_length, dt)
print('done simulating')
padding = 'auto'
fname = run_pareto_plot.run_pareto_analysis_on_specific_sine(noise, dt, timeseries_length, problem, freq, method, method_parent, simdt=simdt, read_existing=read_existing, num_gammas=40, padding=padding)
print(fname)
data = load_data(fname)
plot_example(example, x, x_truth, dxdt_truth, t, color, data, xlimits, ylimits, xticks, yticks) #0.0001_0.1_0.01_4_1
freq_1_gg, opt_rmse, opt_errcorr, opt_idx = get_goldilocks_gamma(data)
freq_1 = copy.copy(freq)
freq_1_color = copy.copy(color)
print('Better RMSE than % randos: ' + str(len(np.where( (opt_rmse<data['rmses']) )[0]) / len(data['rmses']) * 100) + '%')
print('Better Err Corr than % randos: ' + str(len(np.where( (opt_errcorr<data['errcorrs']) )[0]) / len(data['errcorrs']) * 100) + '%')
# +
# define problem
example = 'freq_3'
dt = 0.001
noise = 0.1
timeseries_length = 0.5
problem = 'sine'
freq = 10
if timeseries_length < np.pi/freq:
raise ValueError()
if dt > 1/freq/2.:
raise ValueError()
read_existing = True
simdt = 0.0001
color = 'forestgreen'
# define method
method_parent = 'linear_model'
method = 'savgoldiff'
# define limits
xlimits = {'pos': [0,0.5],
'vel': [0,0.5],
'pareto': [0, 1e2]}
ylimits = {'pos': [-0.2,2.2],
'vel': [-100, 100],
'pareto': [-.1, 1.1]}
xticks = { 'pos': [0,.25, 0.5],
'vel': [0,.25, 0.5],
'pareto': [1e0, 1e1, 1e2]}
yticks = { 'pos': [0, 1, 2],
'vel': [-100, 0, 100],
'pareto': [0, 1]}
r = pynumdiff.utils.simulate.sine(timeseries_length=timeseries_length,
noise_parameters=[0, noise],
dt=dt,
frequencies=[freq])
x, x_truth, dxdt_truth, _ = r
t = np.arange(0, timeseries_length, dt)
print('done simulating')
padding = 'auto'
fname = run_pareto_plot.run_pareto_analysis_on_specific_sine(noise, dt, timeseries_length, problem, freq, method, method_parent, simdt=simdt, read_existing=read_existing, num_gammas=40, padding=padding)
print(fname)
data = load_data(fname)
plot_example(example, x, x_truth, dxdt_truth, t, color, data, xlimits, ylimits, xticks, yticks)
freq_2_gg, opt_rmse, opt_errcorr, opt_idx = get_goldilocks_gamma(data)
freq_2 = copy.copy(freq)
freq_2_color = copy.copy(color)
print('Better RMSE than % randos: ' + str(len(np.where( (opt_rmse<data['rmses']) )[0]) / len(data['rmses']) * 100) + '%')
print('Better Err Corr than % randos: ' + str(len(np.where( (opt_errcorr<data['errcorrs']) )[0]) / len(data['errcorrs']) * 100) + '%')
# +
# define problem
example = 'freq_1'
dt = 0.1
noise = 0.1
timeseries_length = 100
problem = 'sine'
freq = 0.01
if timeseries_length < 1/freq:
raise ValueError()
if dt > 1/freq/2.:
raise ValueError()
read_existing = True
simdt = 0.0001
color = 'darkorchid'
# define method
method_parent = 'linear_model'
method = 'savgoldiff'
# define limits
xlimits = {'pos': [0,100],
'vel': [0,100],
'pareto': [1e-3, 1e1]}
ylimits = {'pos': [-0.2,2.2],
'vel': [-.1, .1],
'pareto': [-.1, 1.1]}
xticks = { 'pos': [0, 50, 100],
'vel': [0, 50, 100],
'pareto': [1e-3, 1e-1, 1e1]}
yticks = { 'pos': [0, 1, 2],
'vel': [-0.1, 0, 0.1],
'pareto': [0, 1]}
r = pynumdiff.utils.simulate.sine(timeseries_length=timeseries_length,
noise_parameters=[0, noise],
dt=dt,
frequencies=[freq])
x, x_truth, dxdt_truth, _ = r
t = np.arange(0, timeseries_length, dt)
print('done simulating')
padding = 'auto'
fname = run_pareto_plot.run_pareto_analysis_on_specific_sine(noise, dt, timeseries_length, problem, freq, method, method_parent, simdt=simdt, read_existing=read_existing, num_gammas=40, padding=padding)
print(fname)
data = load_data(fname)
plot_example(example, x, x_truth, dxdt_truth, t, color, data, xlimits, ylimits, xticks, yticks)
freq_3_gg, opt_rmse, opt_errcorr, opt_idx = get_goldilocks_gamma(data)
freq_3 = copy.copy(freq)
freq_3_color = copy.copy(color)
print('Better RMSE than % randos: ' + str(len(np.where( (opt_rmse<data['rmses']) )[0]) / len(data['rmses']) * 100) + '%')
print('Better Err Corr than % randos: ' + str(len(np.where( (opt_errcorr<data['errcorrs']) )[0]) / len(data['errcorrs']) * 100) + '%')
# +
# define problem
example = 'freq_4'
noise = 0.1
dt = 0.001
timeseries_length = 0.05
problem = 'sine'
freq = 100
if timeseries_length < np.pi/freq:
raise ValueError()
if dt > 1/freq/2.:
raise ValueError()
read_existing = True
simdt = 0.0001
color = 'peru'
# define method
method_parent = 'linear_model'
method = 'savgoldiff'
# define method
xlimits = {'pos': [0,0.05],
'vel': [0,0.05],
'pareto': [1e1, 1e3]}
ylimits = {'pos': [-0.2,2.2],
'vel': [-1000, 1000],
'pareto': [-.1, 1.1]}
xticks = { 'pos': [0, 0.025, 0.05],
'vel': [0, 0.025, 0.05],
'pareto': [1e1, 1e2, 1e3]}
yticks = { 'pos': [0, 1, 2],
'vel': [-1000, 0, 1000],
'pareto': [0, 1]}
r = pynumdiff.utils.simulate.sine(timeseries_length=timeseries_length,
noise_parameters=[0, noise],
dt=dt,
frequencies=[freq])
x, x_truth, dxdt_truth, _ = r
t = np.arange(0, timeseries_length, dt)
print('done simulating')
padding = 'auto'
fname = run_pareto_plot.run_pareto_analysis_on_specific_sine(noise, dt, timeseries_length, problem, freq, method, method_parent, simdt=simdt, read_existing=read_existing, num_gammas=40, padding=padding)
print(fname)
data = load_data(fname)
plot_example(example, x, x_truth, dxdt_truth, t, color, data, xlimits, ylimits, xticks, yticks)
freq_4_gg, opt_rmse, opt_errcorr, opt_idx = get_goldilocks_gamma(data)
freq_4 = copy.copy(freq)
freq_4_color = copy.copy(color)
print('Better RMSE than % randos: ' + str(len(np.where( (opt_rmse<data['rmses']) )[0]) / len(data['rmses']) * 100) + '%')
print('Better Err Corr than % randos: ' + str(len(np.where( (opt_errcorr<data['errcorrs']) )[0]) / len(data['errcorrs']) * 100) + '%')
# +
# make freq plots
# -
def get_filenames(path, contains, does_not_contain=['~', '.pyc']):
cmd = 'ls ' + '"' + path + '"'
ls = os.popen(cmd).read()
all_filelist = ls.split('\n')
try:
all_filelist.remove('')
except:
pass
filelist = []
for i, filename in enumerate(all_filelist):
if contains in filename:
fileok = True
for nc in does_not_contain:
if nc in filename:
fileok = False
if fileok:
filelist.append( os.path.join(path, filename) )
return filelist
def get_freq_dt_noise_for_files(dirname, method, method_parent):
filenames = get_filenames(dirname, method)
freqs = []
dt = []
noises = []
fnames = []
paddings = []
timeseries_length = []
goldgammas = []
for fname in filenames:
data = load_data(fname)
if method == data['metadata']['method']:
if method_parent == data['metadata']['method_parent']:
try:
freqs.append(data['metadata']['freq'])
except:
freqs.append(None)
dt.append(data['metadata']['dt'])
noises.append(data['metadata']['noise'])
fnames.append(fname)
paddings.append(data['metadata']['padding'])
timeseries_length.append(data['metadata']['timeseries_length'])
goldgammas.append(get_goldilocks_gamma(data)[0])
df = pandas.DataFrame({'freq': freqs,
'dt': dt,
'noise': noises,
'fname': fnames,
'padding': paddings,
'timeseries_length': timeseries_length,
'goldgammas': goldgammas})
return df
dirname = 'pareto_sine_freq_data_varpadding'
method = 'savgoldiff'
method_parent = 'linear_model'
df = get_freq_dt_noise_for_files(dirname, method, method_parent)
df.timeseries_length.unique()
def plot_gamma_vs_freq(ax, df, color, marker):
dfq = df[df.timeseries_length >= 1/df.freq]
dfq = dfq[dfq.dt <= 1/dfq.freq/2.]
print(len(dfq))
ax.plot(dfq.freq.values + np.random.uniform(-.5, 0.5, len(dfq.freq.values))*np.abs(dfq.freq.values),
dfq.goldgammas.values + np.random.uniform(-0.5, 0.5, len(dfq.freq.values))*np.abs(dfq.goldgammas.values),
marker, color=color)
dfq = df[df.timeseries_length < 1/df.freq]
ax.plot(dfq.freq.values + np.random.uniform(-.5, 0.5, len(dfq.freq.values))*np.abs(dfq.freq.values),
dfq.goldgammas.values + np.random.uniform(-0.5, 0.5, len(dfq.freq.values))*np.abs(dfq.goldgammas.values),
'+', color=color)
#dfq = dfq[dfq.dt > 1/dfq.freq/2.]
#ax.plot(dfq.freq.values + np.random.uniform(-.5, 0.5, len(dfq.freq.values))*np.abs(dfq.freq.values),
# dfq.goldgammas.values + np.random.uniform(-0.5, 0.5, len(dfq.freq.values))*np.abs(dfq.goldgammas.values),
# '+', color=color)
# +
layout = fifi.svg_to_axes.FigureLayout(figure_layout, autogenlayers=True,
make_mplfigures=True, hide_layers=[])
ax4 = layout.axes[('gamma_vs_freq_4', 'gamma_vs_freq')]
df_dt1 = df.query('dt == 0.1')
plot_gamma_vs_freq(ax4, df_dt1, "coral", "^")
#plot_gamma_vs_freq(ax4, df_dt1[df_dt1.timeseries_length >= 1/df_dt1.freq], "coral", "s")
df_dt01 = df.query('dt == 0.01')
plot_gamma_vs_freq(ax4, df_dt01, "orangered", "^")
#plot_gamma_vs_freq(ax4, df_dt01[df_dt01.timeseries_length >= 1/df_dt01.freq], "orangered", "s")
df_dt001 = df.query('dt == 0.001')
plot_gamma_vs_freq(ax4, df_dt001, "firebrick", "^")
#plot_gamma_vs_freq(ax4, df_dt001[df_dt001.timeseries_length >= 1/df_dt001.freq], "firebrick", "s")
# empirical relationship
freqs = np.logspace(-4, 2)
gg = get_gamma(0.1, freqs)
gg = np.exp(-0.71*np.log(0.1) -1.59*np.log(freqs) -5.1)
ax4.plot(freqs, gg, color='coral')
freqs = np.logspace(-4, 2)
gg = get_gamma(0.01, freqs)
gg = np.exp(-0.71*np.log(0.01) -1.59*np.log(freqs) -5.1)
ax4.plot(freqs, gg, color='orangered')
freqs = np.logspace(-4, 2)
gg = get_gamma(0.001, freqs)
gg = np.exp(-0.71*np.log(0.001) -1.59*np.log(freqs) -5.1)
ax4.plot(freqs, gg, color='firebrick')
# plot stars
try:
ax4.plot(freq_1, freq_1_gg, '*', color=freq_1_color, markersize=15, markeredgecolor='black')
ax4.plot(freq_2, freq_2_gg, '*', color=freq_2_color, markersize=15, markeredgecolor='black')
ax4.plot(freq_3, freq_3_gg, '*', color=freq_3_color, markersize=15, markeredgecolor='black')
ax4.plot(freq_4, freq_4_gg, '*', color=freq_4_color, markersize=15, markeredgecolor='black')
except:
pass
for ax in [ax4]:
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_ylim(5e-6,5e4)
ax.set_xlim(5e-5,5e2)
ax.minorticks_off()
fifi.mpl_functions.adjust_spines(ax, ['left', 'bottom'],
tick_length=2.5,
xticks = [1e-4, 1e-3, 1e-2, 1e-1, 1e0, 1e1, 1e2],
yticks = [1e-5, 1e-4, 1e-3, 1e-2, 1e-1, 1e-0, 1e1, 1e2, 1e3, 1e4],
spine_locations={'left': 4, 'bottom': 4})
fifi.mpl_functions.set_fontsize(ax, 6)
layout.append_figure_to_layer(layout.figures['gamma_vs_freq_4'], 'gamma_vs_freq_4', cleartarget=True)
layout.write_svg(figure_layout)
# -
def get_correlation(df):
dfq = df[df.timeseries_length >= 1/df.freq]
dfq = dfq[dfq.dt <= 1/dfq.freq/2.]
return scipy.stats.linregress(np.log(dfq.freq), np.log(dfq.goldgammas) )
get_correlation(df)
df_dt01 = df.query('dt == 0.1')
get_correlation(df_dt01)
df_dt01 = df.query('dt == 0.01')
get_correlation(df_dt01)
df_dt01 = df.query('dt == 0.001 ')
get_correlation(df_dt01)
df.columns
import statsmodels.formula.api as smf
def show_ols(df, formula):
dfq = df[df.timeseries_length >= 1/df.freq]
dfq = dfq[dfq.dt <= 1/dfq.freq/2.]
#dfq = dfq[dfq.dt == 0.001]
logdfq_dict = {}
for col in dfq.columns:
if col == 'padding' or col == 'fname':
logdfq_dict[col] = dfq[col]
else:
logdfq_dict[col] = np.log(dfq[col])
logdfq = pandas.DataFrame(logdfq_dict)
est = smf.ols(formula=formula, data=logdfq).fit()
return est
formula = 'goldgammas ~ freq + dt + noise + timeseries_length'
est = show_ols(df, formula)
est.summary2()
formula = 'goldgammas ~ freq + dt'
est = show_ols(df, formula)
est.summary2()
# # Try all combinations for statsmodel
import statsmodels.formula.api as smf
def show_ols(df, formula):
dfq = df[df.timeseries_length >= 1/df.freq]
dfq = dfq[dfq.dt <= 1/dfq.freq/2.]
#dfq = dfq[dfq.dt == 0.001]
log_cols = []
logdfq_dict = {}
for col in dfq.columns:
if col == 'padding' or col == 'fname':
logdfq_dict[col] = dfq[col]
else:
if np.random.random() < 0.3:
log_cols.append(col)
logdfq_dict[col] = np.log(dfq[col])
else:
logdfq_dict[col] = dfq[col]
logdfq = pandas.DataFrame(logdfq_dict)
est = smf.ols(formula=formula, data=logdfq).fit()
return est, log_cols
formula = 'goldgammas ~ freq + dt + noise + timeseries_length'
est, log_cols = show_ols(df, formula)
print(est.rsquared_adj)
rsqs = []
logs = []
for i in range(1000):
est, log_cols = show_ols(df, formula)
rsqs.append(est.rsquared_adj)
logs.append(log_cols)
logs[np.argmax(rsqs)]
| notebooks/paper_figures/make_fig_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Modern House Premium
#
# [The Modern House](https://www.themodernhouse.com) is a niche real estate agent that lists only architecturally unique homes for sale. The listings are tastefully presented with professional photos and introduction into the architecture history of the building. As such, the website claims that they are able to achieve a 12% premium in selling prices compared with other real estate agencies.
#
# <img src="dataloft.png" alt="claim" width="300" height="200"/>
#
# I attempt to verify this claim through a webscraping exercise. To do this, I scrape Modern House's current listing prices, address and number of bedrooms from their website. I then use the Zoopla API to download price information of other house for sale around 0.5 mile radius of the Modern House listings to get a sense of average selling price of the neighbourhood. I then compare to see if the Modern House listings do actually have a premium price listing compared to its neighbours.
#
# +
#Importing all the packages that will be used
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore') #Ignore package warnings
import os
#Webscraping tools
from urllib.request import urlopen
from bs4 import BeautifulSoup
from selenium import webdriver
import re
from zoopla import Zoopla
import time
#Data analysis
import pandas as pd
import numpy as np
#Visualisations
import seaborn as sns
import matplotlib.pyplot as plt
import folium
import geopandas
from geopandas import GeoDataFrame
from shapely.geometry import Point
import fiona
import matplotlib.pyplot as plt
pd.options.display.max_rows = 10
# -
# ## Data collection
#
# ### 1. Webscraping The Modern House website using Selenium & Beautiful Soup
#
# The modern house url ends with all the houses on sale for a specific number of bedrooms. Ideally, I would need the per square meter prices of a property. However, this information is only available in the form of floor plan which is in a PDF image. The closest approximation to the size would be number of bedrooms. My strategy is to create functions to scroll through the website and collect data. This is then repeated for all the pages for different number of bedrooms.
# +
#Webscraping url - The Modern House
##Build a function to scroll down the pages to the end and extract page source using Chrome
def scrollExtractChrome(url):
#Using the chrome driver
chrome_path = os.getcwd() + '/chromedriver'
browser = webdriver.Chrome(chrome_path)
# Tell Selenium to get the URL you're interested in.
browser.get(url)
# Selenium script to scroll to the bottom, wait 3 seconds for the next batch of data to load, then continue scrolling. It will continue to do this until the page stops loading new data.
lenOfPage = browser.execute_script("window.scrollTo(0, document.body.scrollHeight);var lenOfPage=document.body.scrollHeight;return lenOfPage;")
match=False
while(match==False):
lastCount = lenOfPage
time.sleep(3)
lenOfPage = browser.execute_script("window.scrollTo(0, document.body.scrollHeight);var lenOfPage=document.body.scrollHeight;return lenOfPage;")
if lastCount==lenOfPage:
match=True
# Now that the page is fully scrolled, grab the source code.
return browser.page_source
##Define a function to extract data from the page source using a specific regex pattern
def extractFromXML(page,tag,tag_class,pattern):
#Create BeautifulSoup object
soup = BeautifulSoup(page,'lxml')
#Filter all the entries
rows= soup.find_all(tag,attrs={'class':tag_class})
#Use the regex pattern to extract data needed
attributes=[re.findall(pattern, i.prettify()) for i in rows]
#Flatten out row list for easy import into pandas
return [item for sublist in attributes for item in sublist]
# +
#Define parameters for the functions made above
url = "https://www.themodernhouse.com/sales-list/homes/all/"
pattern_location = "<h3 class=\"listing-name\">\\n\s+(?P<name>.*)\\n\s+<br/>\\n\s+(?P<postcode>.*)\\n</h3>\\n"
pattern_price = "<div class=\"listing-price\">\\n\s+(?P<price>£.*)\\n\s+<br/>\\n\s+(?P<hold>.*)\\n</div>"
#Compile all the information downloaded into a dataframe
df = pd.DataFrame()
for i in range(1,6):
link = url + str(i)
#Extract data using the defined functions:
page = scrollExtractChrome(link)
location_extract = extractFromXML(page,'h3', 'listing-name', pattern_location)
price_extract = extractFromXML(page,'div', 'listing-price', pattern_price)
#Join two datasets together and import to pandas
data = [a+b for a,b in zip(location_extract,price_extract)]
labels = ['address','postcode','price','hold']
df_part = pd.DataFrame.from_records(data, columns=labels)
df_part['bedrooms'] = i
df = df.append(df_part)
# -
df = df.reset_index(drop=True)
df.head(20)
# ### Data cleaning for the Modern House price data
#
# The main problems are:
# 1. Changing price data to numeric
# 2. Parts of the address data is within the postcode column: Need to split out the address into building, block, street, postcode and area
df.info()
# +
#Data cleaning
#change price to numeric
df['price'] = pd.to_numeric(df['price'].replace('[\D]','',regex=True))
#separate out postcode column with further details of address
postcode_split1=df['postcode'].str.split(',',expand=True)
postcode_split1.head(20)
#The problem is that address information isn't uniform across all listings. Some have building names and some don't.
#Require another function to push last non-blank information to the last column.
# +
def pushLastColumn(df):
n = df.shape
#Find rows that already have values in all columns or no values at all: we can ignore
fixed = [i for i,v in enumerate(df.notnull().all(axis=1)) if v == True]
empty = [i for i,v in enumerate(df.isnull().all(axis=1)) if v == True]
exceptions = fixed + empty
#Find all the position where the figures should move
i = np.where(df.notnull())
#Those in the exception rows doesn't need to move
boolean_filter= [False if x in exceptions else True for x in i[0]]
#Move the last value in each row
i_last = pd.Index(i[0]).duplicated(keep='last')
dat_loc = [not i for i in i_last]
dat_loc = np.logical_and(dat_loc, boolean_filter)
#Get the iloc locations of all the figures that should move
fromloc = list(zip(i[0][dat_loc],i[1][dat_loc]))
#Find the location of all the NaN
j = np.where(df.isnull())
#Find the location where the NaN should be filled in last columns
boolean_filter= [False if x in exceptions else True for x in j[0]]
j_last = pd.Index(j[0]).duplicated(keep='last')
fill_loc = [not i for i in j_last]
fill_loc = np.logical_and(fill_loc, boolean_filter)
toloc = list(zip(j[0][fill_loc],j[1][fill_loc]))
#update dataframe by shifting cell positions
l = len(fromloc)
for x in range(l):
df.iloc[toloc[x][0],toloc[x][1]] = df.iloc[fromloc[x][0],fromloc[x][1]]
df.iloc[fromloc[x][0],fromloc[x][1]] = np.nan
return df
# -
postcode_split2 = pushLastColumn(postcode_split1)
postcode_split2.rename(columns = {0:'building', 1:'street', 2:'area'}, inplace=True)
postcode_split2.head(20)
#We still have a third part of address not contained in the initial postcode
address_parts = pd.concat([df.loc[:,'address'],postcode_split2.drop(['area'], axis=1)], axis=1)
address_parts.head(20)
#We want to push and collect all the first parts to the streets column
address_parts = pushLastColumn(address_parts)
address_parts.rename(columns={'address':'building','building':'block'}, inplace=True)
address_parts.head(20)
#Further split postcode into area and actual postcode
area_postcode=pd.DataFrame()
area_postcode[['area','postcode']]=postcode_split2['area'].str.strip().str.split(' ', expand = True)
area_postcode.head()
#Combining all the different parts of the address and original data
data = pd.concat([address_parts,area_postcode,df.drop(['postcode','address'],axis=1)], axis=1)
data.head()
data.to_csv('modernhousedata.csv')
# ### Visual exploration of the modern home dataset
#Create a category dummy
data['london_dummy'] = ['Inside London' if i == 'London' else 'Outside London' for i in data['area']]
# +
#A quick visualisation to ensure that the price data makes sense --
#The larger the number of bedrooms, the higher the price
g = sns.FacetGrid(data, col='london_dummy')
g.map(sns.regplot, 'bedrooms','price')
#Can see that there is a much steeper line for london which is expected.
# -
# ## Data collection
#
# ### 2. Download neighbourhood price data from Zoopla API
# +
#Get data from Zoopla API
from zoopla import Zoopla
import time
zoopla = Zoopla(api_key='')
def getdata(address,town):
try:
search = zoopla.property_listings({
'radius':'0.5',
'listing_status': 'sale',
'area': address,
'town': town
})
return [(address,town,i.latitude,i.longitude,i.price,i.num_bedrooms, i.agent_name) for i in search.listing]
except Exception as e:
return []
# +
#Number of calls needed
#The limit is 100 calls per 60minutes => Each call interval should be around 36seconds
zoopla_data = []
for i in list(zip(data.loc[:,'street'], data.loc[:,'area'])):
x=getdata(i[0],i[1]) #get data from zoopla
x=[item+(i[0],)+(i[1],) for item in x ] #append search street and area
#time.sleep(36)
zoopla_data = zoopla_data + x
# -
cols = ['address','town','latitude','longitude','price','bedrooms','agent','search_street','search_area']
zoopla_df = pd.DataFrame(zoopla_data, columns=cols)
#zoopla_df.to_csv('zoopla_df.csv')
df = pd.read_csv('zoopla_df.csv')
df.head()
df.info()
# ### Data cleaning for Zoopla data set
#Remove first unnamed column
longdata = df.drop(list(df)[0],axis=1)
#Remove any listing from The Modern House
filter_mh = longdata.loc[:,'agent'].str.contains('The Modern House')
neighbourhood = longdata[[not i for i in filter_mh]]
neighbourhood = neighbourhood.reset_index(drop=True)
neighbourhood.head()
# ### Exploratory visualisation of Zoopla data
#Turn CSV into a Point geomatry (Point GeoDataFrame) with WSG84 Coordinate Reference System
geometry = [Point(xy) for xy in zip(df.longitude, df.latitude)]
crs = {'init': 'epsg:4326'}
houses = GeoDataFrame(df, crs=crs, geometry=geometry)
#Edit first column name
houses.columns.values[0] = "objectid"
houses.head()
#Create two subsets of the data: one for ModernHouse houses and one for all other agents:
mh_filter = houses.loc[:,'agent'].str.contains('The Modern House')
reverse = [not i for i in mh_filter]
modernhouse = houses[mh_filter].reset_index(drop=True)
otherhouse = houses[reverse].reset_index(drop=True)
# +
#Create a list of locations for both datasets:
locationlistMH = modernhouse[['latitude', 'longitude']].values.tolist()
locationlistOthers = otherhouse[['latitude', 'longitude']].values.tolist()
# +
from folium.plugins import MarkerCluster
#Create the basemap
map1 = folium.Map(location=[51.5, -0.10], tiles='CartoDB dark_matter', zoom_start=7)
#Create a cluster of Modern House points in Red
for point in range(0, len(locationlistMH)):
folium.Marker(locationlistMH[point], popup=modernhouse['agent'][point],
icon=folium.Icon(color='red', prefix='fa', icon='home')).add_to(map1)
#Create a cluster of Other house points in Green
for point in range(0, len(locationlistOthers)):
folium.Marker(locationlistOthers[point], popup=otherhouse['agent'][point],
icon=folium.Icon(color='green', prefix='fa', icon='home')).add_to(map1)
#Plot the map
map1
# -
# ## Data analysis - Finding the price premium
#Get the mean house price for each street and type of bedroom
#Compare this with The Modern House's prices
g=pd.DataFrame(neighbourhood.groupby(['search_street','bedrooms']).price.mean(), columns = ['price'])
combinedata = pd.merge(data, g, how = 'inner', left_on=['street','bedrooms'], right_index=True).reset_index(drop=True)
combinedata.head()
#Plot an inital comparison between Modern House price and Zoopla prices
ax = combinedata.set_index('street')[['price_x','price_y']].plot(kind='bar', \
title ="Comparison", figsize=(15, 10), legend=True, fontsize=12, use_index=True)
ax.set_xlabel("Addresses", fontsize=12)
ax.set_ylabel("Prices", fontsize=12)
ax.legend(('Modern Home Prices','Zoopla prices'))
plt.show()
#filter out differences higher than 50% as this is likely to be driven by differences in square meters
combinedata['price_difference'] = combinedata['price_x'] - combinedata['price_y']
combinedata['lower_price'] = combinedata[['price_x','price_y']].min(axis=1)
combinedata2 = combinedata[(combinedata['price_difference'].abs()/combinedata['lower_price'])<0.5].reset_index(drop=True)
combinedata2
#I get a really small sample to make the right comparison
# +
#Plotting the filtered list to compare
ax = combinedata2.set_index('street')[['price_x','price_y']].plot(kind='bar', \
title ="Comparison", figsize=(15, 10), legend=True, fontsize=12, use_index=True)
ax.set_xlabel("Addresses", fontsize=12)
ax.set_ylabel("Prices", fontsize=12)
ax.legend(('Modern Home Prices','Zoopla prices'))
plt.show()
#No reason to say that Modern House Prices have a premium over its neighbours
# -
#Have a sense of the premium over Zoopla prices
combinedata2['premium']=combinedata2['price_difference']/combinedata2['price_y']
combinedata2['premium'].describe() #Doesn't seem to have any premium
#Perform a formal statistical test for a paired list
from scipy import stats
stats.wilcoxon(combinedata2['price_difference'])
#Do not reject that there is a price difference
# ## Conclusion and results
#
# - The main limitations of my approach is from the Zoopla API which is ill-maintained. The API key expires after a day of usage and hence not alot of data could be collected. Future work could include direct scraping from the Zoopla website itself.
#
# - The other main limitation is that my result tries to compare like-for-like for number of bedrooms. However, there is quite a large variation regarding actual floor size of the houses even within two bedroom flats. A better analysis requires us to calculate per square foot price of houses.
#
# **From the limited dataset for comparison, I found that there isn't enough evidence to say that Modern House is able to charge a premium on its listings.**
| ModernHouseScraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as img
import cv2
import math
import imageio
from matplotlib import pyplot
# # Step 1: Generation of mask
def calculate_filter_size (sigma, T):
sHalf = round(np.sqrt(-np.log(T)*2*sigma**2))
filter_size = 2*sHalf+1
n = np.linspace(int(-sHalf),int(sHalf),int(filter_size))
return (n)
def calculate_gradient(sigma,n):
[Y,X] = np.meshgrid(n, n)
Gx = np.round((-X/(sigma**2))*np.exp(-(X**2+Y**2)/(2*sigma**2))*255)
Gy = np.round((-Y/(sigma**2))*np.exp(-(X**2+Y**2)/(2*sigma**2))*255)
return(Gx,Gy)
sigma = 1
T = 0.3
filter_size = calculate_filter_size (sigma, T)
Gx, Gy = calculate_gradient(0.5,filter_size)
Gx
Gy
# # Step 2: Applying Masks to Images
def applying_mask(img_g, Gx, Gy):
r1,c1 = img_g.shape
r2,c2 = Gx.shape
fx = np.zeros([r1-2,c1-2])
fy = np.zeros([r1-2,c1-2])
offset = np.floor(r2//2)
for i in range(int(r1-2*offset)):
i = i + offset
for j in range(int(c1-2*offset)):
j = j + offset
A = img_g[int((i-offset)):int(i+offset)+1, int(j-offset):int(j+offset)+1]
fx[int(i-offset),int(j-offset)] = sum(sum(A*Gx))
fy[int(i-offset),int(j-offset)] = sum(sum(A*Gy))
return (fx, fy)
img_g = cv2.imread('Dataset/img_1.jpg', cv2.IMREAD_GRAYSCALE)
fx,fy = applying_mask(img_g, Gx, Gy)
# +
fig, axs = plt.subplots(1,3, figsize=(15, 10))
axs[0].imshow(img_g, cmap = 'gray')
axs[0].title.set_text("Input Image")
axs[0].axis('off')
axs[1].imshow(fx, cmap = 'gray')
axs[1].title.set_text("fx")
axs[1].axis('off')
axs[2].imshow(fy, cmap = 'gray')
axs[2].title.set_text("fy")
axs[2].axis('off')
# -
imageio.imwrite('Results/img_1.jpg', (img_g).astype(np.uint8))
imageio.imwrite('Results/img_1_fx_[{}].jpg'.format(sigma), (fx))
imageio.imwrite('Results/img_1_fy_[{}].jpg'.format(sigma), (fy))
# # Step 3: Compute gradient magnitude
def gradient_magnitude(fx, fy):
m = np.sqrt(fx**2+fy**2)
max_val = np.max(m)
M=np.round((m/max_val)*255)
return(M)
M = gradient_magnitude(fx, fy)
# +
fig, axs = plt.subplots(1,2, figsize=(15, 10))
axs[0].imshow(img_g, cmap = 'gray')
axs[0].title.set_text("Input Image")
axs[0].axis('off')
axs[1].imshow(M, cmap = 'gray')
axs[1].title.set_text("Gradient Magnitude (M)")
axs[1].axis('off')
# -
imageio.imwrite('Results/img_1_magnitude_[{}].jpg'.format(sigma), (M))
# # Step 4: Compute gradient Direction
def gradient_direction(fx, fy):
r,c = fx.shape
theta = np.zeros([r, c])
for i in range(r):
for j in range(c):
theta[i,j] = math.degrees(math.atan2(fy[i,j],fx[i,j]))
theta = theta + 180
return (theta)
theta=gradient_direction(fx, fy)
theta[M < 10] = -1
fig = plt.figure(111, figsize=(10,10))
plt.imshow(theta)
plt.show()
# # Step 5: Non-Maxima Suppression
def quantizing_theta(theta):
r,c = theta.shape
new_theta = np.zeros([r, c])
for i in range(r):
for j in range(c):
if ((0 <= theta[i,j])&(theta[i,j] < 22.5)):
new_theta[i,j] = 0
if ((157.5 <= theta[i,j])&(theta[i,j] < 202.5)):
new_theta[i,j] = 0
if ((337.5 <= theta[i,j])&(theta[i,j] < 360)):
new_theta[i,j] = 0
if ((22.5 <= theta[i,j])&(theta[i,j] < 67.5)):
new_theta[i,j] = 1
if ((202.5 <= theta[i,j])&(theta[i,j] < 247.5)):
new_theta[i,j] = 1
if ((67.5 <= theta[i,j])&(theta[i,j] < 112.5)):
new_theta[i,j] = 2
if ((247.5 <= theta[i,j])&(theta[i,j] < 292.5)):
new_theta[i,j] = 2
if ((112.5 <= theta[i,j])&(theta[i,j] < 157.5)):
new_theta[i,j] = 3
if ((292.5 <= theta[i,j])&(theta[i,j] < 337.5)):
new_theta[i,j] = 3
return(new_theta)
theta = quantizing_theta(theta)
fig = plt.figure(111, figsize=(10,10))
plt.imshow(theta)
plt.show()
imageio.imwrite('Results/img_1_quantized_[{}].jpg'.format(sigma), (theta))
def non_maximum_supression(M, theta):
new_M = np.pad(M, [1, 1], mode='constant')
r,c = M.shape
NMS_Image = np.zeros([r,c])
for i in range(r):
for j in range(c):
if (theta[i,j] == 0):
Compare_Val = max(new_M[(i+1),(j+1)-1], new_M[(i+1),(j+1)+1])
elif (theta[i,j] == 1):
Compare_Val = max(new_M[(i+1)-1,(j+1)-1], new_M[(i+1)+1,(j+1)+1])
elif (theta[i,j] == 2):
Compare_Val = max(new_M[(i+1)-1,(j+1)], new_M[(i+1)+1,(j+1)])
elif (theta[i,j] == 3):
Compare_Val = max(new_M[(i+1)+1,(j+1)-1],new_M[(i+1)-1,(j+1)+1])
if (M[i,j] > Compare_Val):
NMS_Image[i,j] = M[i,j]
NMS_Image = (NMS_Image/np.max(NMS_Image))*255
return(NMS_Image)
new_M = non_maximum_supression(M, theta)
# +
fig, axs = plt.subplots(1,3, figsize=(15, 10))
axs[0].imshow(img_g, cmap = 'gray')
axs[0].title.set_text("Input Image")
axs[0].axis('off')
axs[1].imshow(M, cmap = 'gray')
axs[1].title.set_text("Gradient Magnitude (M)")
axs[1].axis('off')
axs[2].imshow(new_M, cmap = 'gray')
axs[2].title.set_text("Gradient Magnitude with NMS")
axs[2].axis('off')
# -
fig = plt.figure(111, figsize=(10,10))
plt.imshow(new_M, cmap="gray")
plt.show()
imageio.imwrite('Results/img_1_finalresult_[{}].jpg'.format(sigma), (new_M))
# # Step 6: Hysteresis Thresholding
def hysteresis_thresholding(new_M, Th, Tl):
r,c = new_M.shape
new_M[0,:] =0
new_M[r-1,:]=0
new_M[:,0] =0
new_M[:,c-1]=0
new_M[new_M >= Th] = 255
new_M[new_M < Tl] = 0
Edge = np.zeros([r,c])
for i in range(r-1):
i=i+1
for j in range(c-1):
j=j+1
if(M[i,j] >= Th):
(Edge[i-1:i+1,j-1:j+1])[M[i-1:i+1,j-1:j+1] >= Tl] = 255
return (Edge)
T=hysteresis_thresholding(new_M, 70, 0)
fig = plt.figure(111, figsize=(10,10))
plt.imshow(T, cmap="gray")
plt.show()
imageio.imwrite('Results/img_1_hysteresis_[{}]_[70]_[0].jpg'.format(sigma), (T))
| Source_Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ##### iris 데이터
from sklearn.datasets import load_iris
iris = load_iris()
print(iris.DESCR)
df = pd.DataFrame(iris.data, columns=iris.feature_names)
sy = pd.Series(iris.target, dtype='category')
sy.cat.rename_categories(iris.target_names)
df['species'] = sy
df.tail()
sns.pairplot(df, hue='species')
plt.show()
# ##### 뉴스 그룹 텍스트 데이터
from sklearn.datasets import fetch_20newsgroups
newsgroups = fetch_20newsgroups(subset='all')
print(newsgroups.description)
print(newsgroups.keys())
from pprint import pprint
pprint(list(newsgroups.target_names))
| machinelearning/classification/basic/make_classification_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Vocabulary building on Semantic Scholar
#
# The semantic scholar dataset is our largest dataset of purely computer science text. We would like to build an overall vocabulary based on this dataset. The data is already cleaned using the cleaning3 script.
#
# - Remove stopwords (remember that text is lemmatised and a lemmatised list must be passed!)
# - Find a representative list of 1-3grams
#
# In order to filter, we would like to take only words that occur a certain number of times. However, since there is more data in more recent years, it is unfair to set a min document frequency over the entire dataset.
#
# In our previous work:
#
# **For each year:**
# - remove stopwords
# - use countvectorizer to create a vocabulary of ngrams (1-5), using min_df of 6. The idea of this is that if a term is important, at some point it will appear in more than 6 papers in a year. This acts to limit the size of the vocabulary.
# - We then attempted to remove redundant terms (more important when using large ngrams). This prevents too much term overlap.
#
# However:
# - We are playing with a very large amount of data
# - We have not yet tried to remove german abstracts
#
# What we could do this time:
# - remove german abstracts
# - remove stopwords
# - use countvectorizer to create a vocab of 1-5grams using min df of 6
# - vectorize the entire dataset and apply a second min df of 20. We cannot reasonably work out trends which occur less than 20 times in a dataset of 2 million abstracts.
# +
from langdetect import detect
import json
from collections import defaultdict
import numpy as np
import boto3
import re
import unicodedata
from nltk.stem import WordNetLemmatizer
from sklearn.feature_extraction.text import CountVectorizer
from tqdm import tqdm
import random
import pickle
import sys
import time
import csv
sys.path.append("../../tools")
import my_stopwords3
from sys import getsizeof
stop = my_stopwords3.get_stopwords()
# -
# # Language detection
# ### How long does it take to detect german language abstracts?
# +
langs = defaultdict(list)
with open("../../Data/semantic_scholar_cleaned/2000.txt", "r") as f:
for line in tqdm(f, desc='Lines in file processed...'):
langs[detect(line[0:1000])].append(line[0:200])
# -
# It would take about 7 hours to run this on the entire dataset on this PC
# ### How good is the detection?
#
for key in langs.keys():
print(key, len(langs[key]))
langs['de']
langs['es']
del(langs)
# **Thoughts: it's pretty good**
#
# We'd be doing this to remove about 1% of abstracts. Unfortunately I think it will be necessary. Let's try again with a smaller sample of text:
# +
langs = defaultdict(list)
with open("../../Data/semantic_scholar_cleaned/2000.txt", "r") as f:
for line in tqdm(f, desc='Lines in file processed...'):
langs[detect(line[0:200])].append(line[0:200])
# -
for key in langs.keys():
print(key, len(langs[key]))
langs['fr']
# **Thoughts: the reduction in time taken isn't sufficient to justify a major cut in the amount of available text**
#
# ## Go through all abstracts and remove the ones which are not in english language
for year in range(1980, 2000):
t0 = time.time()
to_write = []
with open("D:/CDT/Data/semantic_scholar_cleaned/"+str(year)+".txt", "r") as f:
for line in tqdm(f, desc='Lines in file processed...', mininterval=10):
if detect(line[0:1000]) == 'en':
to_write.append(line)
with open("../../Data/semantic_scholar_cleaned_langdetect/"+str(year)+".txt", "a") as f:
for line in to_write:
f.write(line)
del to_write
print(year, time.time()-t0)
# # Build a vocabulary
#
# - lemmatise stopwords
# - for each year, remove stopwords, then build a vocabulary.
# - I want the overall vocabulary to be stored in a dictionary such that the count is the total document count of the word. This should be updated each year.
#
# +
vectorizer = CountVectorizer(strip_accents='unicode',
ngram_range=(1,5),
stop_words=stop,
min_df=6,
max_df = 1000000
)
t0 = time.time()
with open("../../Data/semantic_scholar_cleaned/1980.txt", "r") as f:
documents = f.readlines()
documents = [d.strip() for d in documents]
vector = vectorizer.fit_transform(documents)
print(time.time()-t0)
# -
# set all elements that are >1 in the vector to 1. This is done so we can calculate document frequency of terms.
print(np.sum(vector))
vector[vector>1] = 1
print(np.sum(vector))
np.asarray(np.sum(vector, axis=0))[0]
len(vectorizer.vocabulary_)
# +
vocabulary = defaultdict(int)
vector[vector>1] = 1
document_frequency = np.asarray(np.sum(vector, axis=0))[0]
for term in vectorizer.vocabulary_:
vocabulary[term] += document_frequency[vectorizer.vocabulary_[term]]
del vector
del vectorizer
# -
# ## Now build the vocabulary for real...
# +
vocabulary = defaultdict(int)
for year in range(2020, 1999, -1):
t0 = time.time()
with open("E:/Data/semantic_scholar_cleaned_langdetect/"+str(year)+".txt", "r") as f:
documents = f.readlines()
documents = [d.strip() for d in documents]
vectorizer = CountVectorizer(strip_accents='unicode',
ngram_range=(1,4),
stop_words=stop,
min_df=10,
max_df = 1000000
)
vector = vectorizer.fit_transform(documents)
del documents
vector[vector>1] = 1
document_frequency = np.asarray(np.sum(vector, axis=0))[0]
for term in vectorizer.vocabulary_:
vocabulary[term] += document_frequency[vectorizer.vocabulary_[term]]
del vector
del vectorizer
del document_frequency
pickle.dump(vocabulary, open("interim_vocabulary.p", "wb"))
print(year, len(vocabulary.keys()), time.time()-t0)
# -
list(vocabulary.keys())
pickle.dump(list(vocabulary.keys()), open("vocabulary.p", "wb"))
len(set(list(vocabulary.keys())))
# The issue with this approach is that it uses a massive amount of space, and my poor computer cannot cope. The smallest file, 2000.txt, contains 60,000 documents. We can't really realistically find terms occuring in less than 0.2% per year. Therefore, set cut off at 30 documents in a single year at their peak.
#
# We also don't need to hold the entire matrix, since we are only trying to build a vocabulary.
#
# In order to do this in manageable chunks, how about vectorizing 10,000 at a time?
# +
chunk = 10000
min_yearly_df = 20
vocabulary2 = defaultdict(int)
t0 = time.time()
interim_vocabulary = defaultdict(int)
with open("../../Data/semantic_scholar_cleaned_langdetect/2000.txt", "r") as f:
documents = f.readlines()
documents = [d.strip() for d in documents]
random.shuffle(documents)
# Go through the documents in chunks, creating a vocabulary
for i in tqdm(range(int(np.ceil(len(documents)/chunk))-1), desc='Chunks processed'):
vectorizer = CountVectorizer(strip_accents='unicode',
ngram_range=(1,4),
stop_words=stop,
min_df=2,
max_df = chunk
)
vector = vectorizer.fit_transform(documents[chunk*i:chunk*(i+1)])
vector[vector>1] = 1
document_frequency = np.asarray(np.sum(vector, axis=0))[0]
del vector
for term in vectorizer.vocabulary_:
interim_vocabulary[term] += document_frequency[vectorizer.vocabulary_[term]]
del document_frequency
del vectorizer
# Now do for the last remaining documents
vectorizer = CountVectorizer(strip_accents='unicode',
ngram_range=(1,4),
stop_words=stop,
min_df=1,
max_df = chunk
)
vector = vectorizer.fit_transform(documents[chunk*(i+1):chunk*(i+2)])
vector[vector>1] = 1
document_frequency = np.asarray(np.sum(vector, axis=0))[0]
del vector
for term in vectorizer.vocabulary_:
interim_vocabulary[term] += document_frequency[vectorizer.vocabulary_[term]]
del document_frequency
del vectorizer
del documents
for term in interim_vocabulary.keys():
if interim_vocabulary[term] >= min_yearly_df:
vocabulary2[term] += interim_vocabulary[term]
print(len(vocabulary2), time.time()-t0)
# +
chunk = 10000
min_yearly_df = 30
vocabulary = defaultdict(int)
t0 = time.time()
interim_vocabulary = defaultdict(int)
with open("../../Data/semantic_scholar_cleaned_langdetect/2000.txt", "r") as f:
documents = f.readlines()
documents = [d.strip() for d in documents]
vectorizer = CountVectorizer(strip_accents='unicode',
ngram_range=(1,4),
stop_words=stop,
min_df=30,
max_df = 100000
)
vector = vectorizer.fit_transform(documents)
vector[vector>1] = 1
document_frequency = np.asarray(np.sum(vector, axis=0))[0]
for term in vectorizer.vocabulary_:
vocabulary[term] += document_frequency[vectorizer.vocabulary_[term]]
print(len(vocabulary), time.time()-t0)
# -
del vector
del document_frequency
del vectorizer
del documents
| 2_vectorization/Semantic scholar vocabulary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/daanishrasheed/DS-Unit-2-Applied-Modeling/blob/master/module2/assignment_applied_modeling_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="gDvAbg9kLZYz" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 3, Module 3*
#
# ---
#
#
# # Applied Modeling, Module 3
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your work.
#
# - [ ] Continue to iterate on your project: data cleaning, exploration, feature engineering, modeling.
# - [ ] Make at least 1 partial dependence plot to explain your model.
# - [ ] Share at least 1 visualization on Slack.
#
# (If you have not yet completed an initial model yet for your portfolio project, then do today's assignment using your Tanzania Waterpumps model.)
#
# ## Stretch Goals
# - [ ] Make multiple PDPs with 1 feature in isolation.
# - [ ] Make multiple PDPs with 2 features in interaction.
# - [ ] Use Plotly to make a 3D PDP.
# - [ ] Make PDPs with categorical feature(s). Use Ordinal Encoder, outside of a pipeline, to encode your data first. If there is a natural ordering, then take the time to encode it that way, instead of random integers. Then use the encoded data with pdpbox. Get readable category names on your plot, instead of integer category codes.
#
# ## Links
# - [<NAME>: Interpretable Machine Learning — Partial Dependence Plots](https://christophm.github.io/interpretable-ml-book/pdp.html) + [animated explanation](https://twitter.com/ChristophMolnar/status/1066398522608635904)
# - [Kaggle / <NAME>: Machine Learning Explainability — Partial Dependence Plots](https://www.kaggle.com/dansbecker/partial-plots)
# - [Plotly: 3D PDP example](https://plot.ly/scikit-learn/plot-partial-dependence/#partial-dependence-of-house-value-on-median-age-and-average-occupancy)
# + id="yabrei1wUg9Y" colab_type="code" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# !pip install eli5
# !pip install pdpbox
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="wpLdx44r5Izj" colab_type="code" outputId="c25ebd2d-a777-45e3-e329-263a60b26616" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 75}
from google.colab import files
uploaded = files.upload()
# + id="JpB9oB6v-CuX" colab_type="code" outputId="a4d337cf-f581-4049-dbbf-508305646a18" colab={"base_uri": "https://localhost:8080/", "height": 430}
import pandas as pd
play = pd.read_csv('pbp-2018.csv', engine='python')
print(play.shape)
play.head()
# + id="teYQEejMId8P" colab_type="code" colab={}
play = play.drop(play[play.Down == 0].index)
# + id="W7IzYpxaIqWi" colab_type="code" outputId="0a25e434-ad22-4a15-f213-d478d0e1c6eb" colab={"base_uri": "https://localhost:8080/", "height": 638}
play.head()
# + id="bdyl7hAbfKtF" colab_type="code" colab={}
df = play.drop(columns=['Unnamed: 10', 'Unnamed: 12', 'Unnamed: 16', 'Unnamed: 17', 'Challenger'])
# + id="FizrD7n4f_dR" colab_type="code" outputId="19cd6cad-5d16-49c6-d0ff-841ab1d4ecd0" colab={"base_uri": "https://localhost:8080/", "height": 551}
df.tail(5)
# + id="wFc6oq7ChS-Q" colab_type="code" colab={}
df1 = df.drop(df[(df.IsRush == 1) & (df.IsPass == 1)].index)
# + id="RPNqBKIBn68l" colab_type="code" outputId="f19ed0d8-c27d-4127-93aa-06965b1712f4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df1.head(10)
# + id="XUP9JiZIqODZ" colab_type="code" colab={}
df = df1.drop(df1[df1.PlayType == 'QB KNEEL'].index)
# + id="TbIoj4SQqa7W" colab_type="code" outputId="66300048-6e5d-4ec8-fa73-93b88b2ef009" colab={"base_uri": "https://localhost:8080/", "height": 621}
df.head()
# + id="wEE-5pOe2_Cp" colab_type="code" outputId="8bc9acf2-8a01-4ab5-e004-b88294b7f5dc" colab={"base_uri": "https://localhost:8080/", "height": 225}
df['PlayType'].value_counts()
# + id="nJYbyRr43_dA" colab_type="code" outputId="fcdf77d2-8b09-4878-8fd3-dbdb65eb3e15" colab={"base_uri": "https://localhost:8080/", "height": 156}
df['Formation'].value_counts()
# + id="ZiOhpo9Z4LuE" colab_type="code" colab={}
df = df.drop(df[df.Formation == 'PUNT'].index)
df = df.drop(df[df.PlayType == 'PUNT'].index)
df = df.drop(df[df.Formation == 'FIELD GOAL'].index)
df = df.drop(df[df.PlayType == 'FIELD GOAL'].index)
# + id="7CvxGGrB4sR-" colab_type="code" outputId="2673e6c8-3d9e-4ae6-bda7-f1130c6ac4ad" colab={"base_uri": "https://localhost:8080/", "height": 621}
df.head()
# + id="8Z61bjHQ5Xcu" colab_type="code" outputId="2e8fe13f-2e8a-42bc-fa44-8bf4025cd472" colab={"base_uri": "https://localhost:8080/", "height": 191}
df['PlayType'].value_counts()
# + id="nAiLOp8pGm5b" colab_type="code" colab={}
df = df.drop(df[df.PlayType == 'NO PLAY'].index)
df = df.drop(df[df.PlayType == 'FUMBLES'].index)
df = df.drop(df[df.PlayType == 'CLOCK STOP'].index)
df = df.drop(df[df.PlayType == 'EXCEPTION'].index)
df = df.drop(df[df.PlayType == 'PENALTY'].index)
# + id="bMcuZefJIYGC" colab_type="code" colab={}
df = df.replace('SACK', 'PASS')
df = df.replace('SCRAMBLE', 'RUSH')
# + id="Dc4KfSFVEtV2" colab_type="code" outputId="d1a153a0-25ac-427d-ad96-3becba3a6035" colab={"base_uri": "https://localhost:8080/", "height": 381}
df['Formation'].head(20)
# + id="0qSJrWiHGOAW" colab_type="code" outputId="f4a2a1d4-de34-4259-ad07-273554c003de" colab={"base_uri": "https://localhost:8080/", "height": 728}
df.iloc[19582]
# + id="BkxNrz5DIn5j" colab_type="code" outputId="7c3e7a5f-b026-4950-aa6d-d05b116a85d4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df.head(13345)
# + id="Gc4iAJbHJ-Og" colab_type="code" colab={}
values = {'PlayType':'RUSH'}
df = df.fillna(value=values)
# + id="vGewC1QTFloY" colab_type="code" colab={}
df['Formation play'] = df[['Formation', 'PlayType']].apply(lambda x: ' '.join(x), axis=1)
# + id="Bu_JncYQY8M0" colab_type="code" colab={}
df = df.drop(df[df.Formation == 'WILDCAT'].index)
# + id="lPstw6N4Ks7b" colab_type="code" outputId="95e37a3c-3971-4b3d-8654-2be07f489d94" colab={"base_uri": "https://localhost:8080/", "height": 638}
df.head()
# + id="FGejJraEIXvN" colab_type="code" colab={}
df['GameDate']= pd.to_datetime(df['GameDate'])
# + id="W29IvnugHBVI" colab_type="code" colab={}
test = df[((df.GameDate.dt.month == 12)) & ((df.GameDate.dt.day ==22) | (df.GameDate.dt.day == 23) | (df.GameDate.dt.day == 24) | (df.GameDate.dt.day == 30))]
# + id="oYoSpPJ5XPcT" colab_type="code" outputId="4a853283-5480-4287-cc3c-64d4cd95b8e9" colab={"base_uri": "https://localhost:8080/", "height": 673}
test.head()
# + id="yXxEp5nGImZN" colab_type="code" colab={}
train = df[(df.GameDate.dt.month == 9) | (df.GameDate.dt.month == 10) | (((df.GameDate.dt.month == 11)) & ((df.GameDate.dt.day == 5) | (df.GameDate.dt.day == 4) | (df.GameDate.dt.day == 1) | (df.GameDate.dt.day == 11) | (df.GameDate.dt.day == 12) | (df.GameDate.dt.day == 8)))]
# + id="ZOxplRgcXVDV" colab_type="code" outputId="6f45cfb3-7fcf-4af6-a6e7-0b704107ade8" colab={"base_uri": "https://localhost:8080/", "height": 638}
train.head()
# + id="8Q91KoEqJyuB" colab_type="code" colab={}
val = df[(((df.GameDate.dt.month == 11)) & ((df.GameDate.dt.day == 18) | (df.GameDate.dt.day == 22) | (df.GameDate.dt.day == 29) | (df.GameDate.dt.day == 25) | (df.GameDate.dt.day == 15) | (df.GameDate.dt.day == 19) | (df.GameDate.dt.day == 26)) | ((df.GameDate.dt.month == 12)) & ((df.GameDate.dt.day == 9) | (df.GameDate.dt.day == 16) | (df.GameDate.dt.day == 2) | (df.GameDate.dt.day == 3) | (df.GameDate.dt.day == 10) | (df.GameDate.dt.day == 17) | (df.GameDate.dt.day == 15) | (df.GameDate.dt.day == 13) | (df.GameDate.dt.day == 6)))]
# + id="719CikBVXcIZ" colab_type="code" outputId="50855706-b2a4-4f46-c6ad-692bf34bb5f5" colab={"base_uri": "https://localhost:8080/", "height": 603}
val.head()
# + id="J0Lxt1YTEY33" colab_type="code" colab={}
target = 'Formation play'
features = ['Quarter', 'Minute', 'Second', 'OffenseTeam', 'DefenseTeam', 'Down', 'ToGo', 'YardLine', 'Formation play']
# + id="kaJ9ifa0717k" colab_type="code" colab={}
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test[features]
y_test = test[target]
# + id="iR_Y50TmWFeS" colab_type="code" outputId="31dcfaeb-c44c-4c2f-c780-e01a9c1da1c1" colab={"base_uri": "https://localhost:8080/", "height": 202}
X_val.head()
# + id="f66MSmVqSF2M" colab_type="code" outputId="0dcb0bb4-6800-495b-a513-3d11247062c6" colab={"base_uri": "https://localhost:8080/", "height": 69}
# %%time
from sklearn.ensemble import RandomForestClassifier
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
pipeline = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(strategy='median'),
RandomForestClassifier(n_estimators=100, random_state=4, n_jobs=-1)
)
# Fit on train, score on val
pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_train)
print('Validation Accuracy', pipeline.score(X_val, y_val))
# + id="IkJmemgFYroG" colab_type="code" outputId="29272fd4-7321-461c-fa7c-9c1b4c87fba1" colab={"base_uri": "https://localhost:8080/", "height": 780}
from xgboost import XGBClassifier
pipeline = make_pipeline(
ce.OrdinalEncoder(),
XGBClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
pipeline.fit(X_train, y_train)
# + id="iN35nr0jYLt2" colab_type="code" outputId="c672490e-9906-43e0-81d7-b433f0a1bc1d" colab={"base_uri": "https://localhost:8080/", "height": 1000}
encoder = ce.OrdinalEncoder()
X_train_encoded = encoder.fit_transform(X_train)
X_val_encoded = encoder.transform(X_val)
model = XGBClassifier(
n_estimators=1000, # <= 1000 trees, depends on early stopping
max_depth=7, # try deeper trees because of high cardinality categoricals
learning_rate=0.5, # try higher learning rate
n_jobs=-1
)
eval_set = [(X_train_encoded, y_train),
(X_val_encoded, y_val)]
model.fit(X_train_encoded, y_train,
eval_set=eval_set,
eval_metric='merror',
early_stopping_rounds=50)
# + id="v5XY-FsEMeWu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="3a9182db-2973-4b0c-9f66-560690fc89d6"
feature = 'Formation play'
X_val[feature].head()
# + id="aX0FTUnvOy_p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="f2c9c16a-cc71-4a6f-d98a-9542cbfb5fbd"
X_val[feature].value_counts()
# + id="rmLrCWVjPGbc" colab_type="code" colab={}
import numpy as np
X_val_permuted = X_val.copy()
X_val_permuted[feature] = np.random.permutation(X_val[feature])
# + id="AbtFAw7KPRCs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="f2faccf9-5220-4f30-dcf6-a0f60e2e29c7"
X_val_permuted[feature].head()
| module2/assignment_applied_modeling_2.ipynb |
/ -*- coding: utf-8 -*-
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ + [markdown] cell_id="00000-8fe37c28-fbd7-47c9-bad4-b2527b2de557" deepnote_cell_type="markdown" tags=[]
/ # Medidas de tendencia central
/
/ ### Media = $$ \frac{1}{N} \sum_{i=1}^N x_i $$
/
/ ### Mediana(impar) = $$ x_{(n+1)/2}^{\text{ordered}} $$
/
/ ### Mediana(par) = $$ \frac{x_{n/2}^{\text{ordered}} + x_{n/2+1}^{\text{ordered}}}{2} $$
/
/ ### Moda = $$ x_k $$ donde $$ \text{Freq}(x_k) = \max{(\text{Freq}(x_i))} $$
/
/ continuamos con el dataset https://www.kaggle.com/lepchenkov/usedcarscatalog
/ + cell_id="00001-4949ec10-a4b1-46d5-b4a7-fcaf895688fd" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=261 execution_start=1622484690689 output_cleared=false source_hash="deaac73e" tags=[]
import pandas as pd
df = pd.read_csv('cars.csv')
df.head()
/ + [markdown] cell_id="00003-5bab5b0d-41f4-4fa7-833b-7f9bf950b644" deepnote_cell_type="markdown" tags=[]
/ inspeccionemos el atributo de `price_usd` **(variable numérica continua)** de los autos listados en el dataset:
/ + cell_id="00002-e401d9fc-ec69-4479-b714-b0ae09329840" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=13 execution_start=1622484690950 output_cleared=false source_hash="bc174504" tags=[]
df['price_usd'].mean()
/ + cell_id="00004-76438628-17f7-42fd-b624-30a9d1940c9c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1622484690960 output_cleared=false source_hash="b7a97061" tags=[]
df['price_usd'].median()
/ + cell_id="00005-40382fd8-491d-487e-958f-cf27c670a3f1" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=828 execution_start=1622484690970 output_cleared=false source_hash="71a18e28" tags=[]
df['price_usd'].plot.hist(bins=20)
/ + [markdown] cell_id="00007-52fff311-495a-4d27-bdb2-642ccf7322eb" deepnote_cell_type="markdown" tags=[]
/ resulta más interesante analizar los precios por marcas:
/
/ * **pro tip:** usar seaborn: https://seaborn.pydata.org/tutorial/distributions.html
/ + cell_id="00008-ebc08b64-affc-4f34-a032-6d4358179ba5" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=27500 execution_start=1622484691838 output_cleared=false source_hash="d5a94b2e" tags=[]
import seaborn as sns
sns.displot(df, x = 'price_usd', hue = 'manufacturer_name')
/ + cell_id="00008-1970d9f6-50df-4a73-843a-c67bbd8d552b" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1769 execution_start=1622484719382 source_hash="f543d35b" tags=[]
sns.displot(df, x="price_usd", hue="engine_type")
/ + [markdown] cell_id="00010-76395294-c383-42da-b492-bf0b4d2ca4f2" deepnote_cell_type="markdown" tags=[]
/ el histograma anterior es muy dificil de analizar, ¿donde están los autos eléctricos?
/ + cell_id="00008-0bb11463-1e55-46b0-baee-2e646d43796d" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1844 execution_start=1622484721157 output_cleared=false source_hash="3a3ac3c2" tags=[]
sns.displot(df, x='price_usd', hue = 'engine_type', multiple='stack')
/ + cell_id="00009-2c3bdab8-0952-43cc-8dc6-a7bcef8753bf" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=119 execution_start=1622484723014 output_cleared=false source_hash="4c516938" tags=[]
df.groupby('engine_type').count()
/ + [markdown] cell_id="00013-70002ad3-5431-42ce-8104-5d7d7c2f5dd1" deepnote_cell_type="markdown" tags=[]
/ **RETO:** Inspeccionemos precios de una marca y modelo particular !
/
/
/ + cell_id="00009-4ced9f5a-0590-443d-8740-4052581ee182" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=649 execution_start=1622484723144 output_cleared=false source_hash="ec032a66" tags=[]
Q7_df = df[(df['manufacturer_name']=='Audi') & (df['model_name']=='Q7')]
sns.histplot(Q7_df, x='price_usd', hue = 'year_produced')
/ + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
/ <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=ae1bc9ae-edaf-4bc7-b716-9d8df4e551fd' target="_blank">
/ <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODBweCIgaGVpZ2h0PSI4MHB4IiB2aWV3Qm94PSIwIDAgODAgODAiIHZlcnNpb249IjEuMSIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayI+CiAgICA8IS0tIEdlbmVyYXRvcjogU2tldGNoIDU0LjEgKDc2NDkwKSAtIGh0dHBzOi8vc2tldGNoYXBwLmNvbSAtLT4KICAgIDx0aXRsZT5Hcm91cCAzPC90aXRsZT4KICAgIDxkZXNjPkNyZWF0ZWQgd2l0aCBTa2V0Y2guPC9kZXNjPgogICAgPGcgaWQ9IkxhbmRpbmciIHN0cm9rZT0ibm9uZSIgc3Ryb2tlLXdpZHRoPSIxIiBmaWxsPSJub25lIiBmaWxsLXJ1bGU9ImV2ZW5vZGQiPgogICAgICAgIDxnIGlkPSJBcnRib2FyZCIgdHJhbnNmb3JtPSJ0cmFuc2xhdGUoLTEyMzUuMDAwMDAwLCAtNzkuMDAwMDAwKSI+CiAgICAgICAgICAgIDxnIGlkPSJHcm91cC0zIiB0cmFuc2Zvcm09InRyYW5zbGF0ZSgxMjM1LjAwMDAwMCwgNzkuMDAwMDAwKSI+CiAgICAgICAgICAgICAgICA8cG9seWdvbiBpZD0iUGF0aC0yMCIgZmlsbD0iIzAyNjVCNCIgcG9pbnRzPSIyLjM3NjIzNzYyIDgwIDM4LjA0NzY2NjcgODAgNTcuODIxNzgyMiA3My44MDU3NTkyIDU3LjgyMTc4MjIgMzIuNzU5MjczOSAzOS4xNDAyMjc4IDMxLjY4MzE2ODMiPjwvcG9seWdvbj4KICAgICAgICAgICAgICAgIDxwYXRoIGQ9Ik0zNS4wMDc3MTgsODAgQzQyLjkwNjIwMDcsNzYuNDU0OTM1OCA0Ny41NjQ5MTY3LDcxLjU0MjI2NzEgNDguOTgzODY2LDY1LjI2MTk5MzkgQzUxLjExMjI4OTksNTUuODQxNTg0MiA0MS42NzcxNzk1LDQ5LjIxMjIyODQgMjUuNjIzOTg0Niw0OS4yMTIyMjg0IEMyNS40ODQ5Mjg5LDQ5LjEyNjg0NDggMjkuODI2MTI5Niw0My4yODM4MjQ4IDM4LjY0NzU4NjksMzEuNjgzMTY4MyBMNzIuODcxMjg3MSwzMi41NTQ0MjUgTDY1LjI4MDk3Myw2Ny42NzYzNDIxIEw1MS4xMTIyODk5LDc3LjM3NjE0NCBMMzUuMDA3NzE4LDgwIFoiIGlkPSJQYXRoLTIyIiBmaWxsPSIjMDAyODY4Ij48L3BhdGg+CiAgICAgICAgICAgICAgICA8cGF0aCBkPSJNMCwzNy43MzA0NDA1IEwyNy4xMTQ1MzcsMC4yNTcxMTE0MzYgQzYyLjM3MTUxMjMsLTEuOTkwNzE3MDEgODAsMTAuNTAwMzkyNyA4MCwzNy43MzA0NDA1IEM4MCw2NC45NjA0ODgyIDY0Ljc3NjUwMzgsNzkuMDUwMzQxNCAzNC4zMjk1MTEzLDgwIEM0Ny4wNTUzNDg5LDc3LjU2NzA4MDggNTMuNDE4MjY3Nyw3MC4zMTM2MTAzIDUzLjQxODI2NzcsNTguMjM5NTg4NSBDNTMuNDE4MjY3Nyw0MC4xMjg1NTU3IDM2LjMwMzk1NDQsMzcuNzMwNDQwNSAyNS4yMjc0MTcsMzcuNzMwNDQwNSBDMTcuODQzMDU4NiwzNy43MzA0NDA1IDkuNDMzOTE5NjYsMzcuNzMwNDQwNSAwLDM3LjczMDQ0MDUgWiIgaWQ9IlBhdGgtMTkiIGZpbGw9IiMzNzkzRUYiPjwvcGF0aD4KICAgICAgICAgICAgPC9nPgogICAgICAgIDwvZz4KICAgIDwvZz4KPC9zdmc+' > </img>
/ Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| descriptive-statistics/descriptive-statistics-course/notebooks/[clase-07]medidas-central.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import syft as sy
# # Part 1: Join the Duet Server the Data Owner connected to
#
duet = sy.join_duet(loopback=True)
# ### <img src="https://github.com/OpenMined/design-assets/raw/master/logos/OM/mark-primary-light.png" alt="he-black-box" width="100"/> Checkpoint 0 : Now STOP and run the Data Owner notebook until Checkpoint 1.
# # Part 2: Search for Available Data
#
# The data scientist can check the list of searchable data in Data Owner's duet store
duet.store.pandas
# +
# Data Scientist finds that there are Heights and Weights of a group of people. There are some analysis he/she can do with them together.
heights_ptr = duet.store[0]
weights_ptr = duet.store[1]
# heights_ptr is a reference to the height dataset remotely available on data owner's server
print(heights_ptr)
# weights_ptr is a reference to the weight dataset remotely available on data owner's server
print(weights_ptr)
# -
# ## Calculate BMI (Body Mass Index) and weight status
#
# Using the heights and weights pointers of the people of Group A, calculate their BMI and get a pointer to their individual BMI. From the BMI pointers, you can check if a person is normal-weight, overweight or obese, without knowing their actual heights and weights and even BMI values.
#
# BMI from 19 to 24 - Normal
# BMI from 25 to 29 - Overweight
# BMI from 30 to 39 - Obese
#
# BMI = [weight (kg) / (height (cm)^2)] x 10,000
# Hint: run duet.torch and find the required operators
# One amazing thing about pointers is that from a pointer to a list of items, we can get the pointers to each item in the list. As example, here we have weights_ptr pointing to the weight-list, but from that we can also get the pointer to each weight and perform computation on each of them without even the knowing the value! Below code will show you how to access the pointers to each weight and height from the list pointer.
for i in range(6):
print("Pointer to Weight of person", i + 1, weights_ptr[i])
print("Pointer to Height of person", i + 1, heights_ptr[i])
# +
def BMI_calculator(w_ptr, h_ptr):
bmi_ptr = 0
##TODO
"Write your code here for calculating bmi_ptr"
###
return bmi_ptr
def weight_status(w_ptr, h_ptr):
status = None
bmi_ptr = BMI_calculator(w_ptr, h_ptr)
##TODO
"""Write your code here.
Possible values for status:
Normal,
Overweight,
Obese,
Out of range
"""""
###
return status
# -
for i in range(0, 6):
bmi_ptr = BMI_calculator(weights_ptr[i], heights_ptr[i])
# +
statuses = []
for i in range(0, 6):
status = weight_status(weights_ptr[i], heights_ptr[i])
print("Weight of Person", i + 1, "is", status)
statuses.append(status)
assert statuses == ["Normal", "Overweight", "Obese", "Normal", "Overweight", "Normal"]
# -
| Foundations_of_Private_Computation/Federated_Learning/duet_basics/exercise/Exercise_Duet_Basics_Data_Scientist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="jrI6q7RmWQam"
# <table align="center">
# <td align="center"><a target="_blank" href="http://introtodeeplearning.com">
# <img src="http://introtodeeplearning.com/images/colab/mit.png" style="padding-bottom:5px;" />
# Visit MIT Deep Learning</a></td>
# <td align="center"><a target="_blank" href="https://colab.research.google.com/github/aamini/introtodeeplearning/blob/master/lab3/RL.ipynb">
# <img src="http://introtodeeplearning.com/images/colab/colab.png?v2.0" style="padding-bottom:5px;" />Run in Google Colab</a></td>
# <td align="center"><a target="_blank" href="https://github.com/aamini/introtodeeplearning/blob/master/lab3/RL.ipynb">
# <img src="http://introtodeeplearning.com/images/colab/github.png" height="70px" style="padding-bottom:5px;" />View Source on GitHub</a></td>
# </table>
#
# # Copyright Information
# + colab={} colab_type="code" id="wkd375upWYok"
# Copyright 2020 MIT 6.S191 Introduction to Deep Learning. All Rights Reserved.
#
# Licensed under the MIT License. You may not use this file except in compliance
# with the License. Use and/or modification of this code outside of 6.S191 must
# reference:
#
# © MIT 6.S191: Introduction to Deep Learning
# http://introtodeeplearning.com
#
# + [markdown] colab_type="text" id="WoXYKhfZMHiw"
# # Laboratory 3: Reinforcement Learning
#
# Reinforcement learning (RL) is a subset of machine learning which poses learning problems as interactions between agents and environments. It often assumes agents have no prior knowledge of a world, so they must learn to navigate environments by optimizing a reward function. Within an environment, an agent can take certain actions and receive feedback, in the form of positive or negative rewards, with respect to their decision. As such, an agent's feedback loop is somewhat akin to the idea of "trial and error", or the manner in which a child might learn to distinguish between "good" and "bad" actions.
#
# In practical terms, our RL agent will interact with the environment by taking an action at each timestep, receiving a corresponding reward, and updating its state according to what it has "learned".
#
# 
#
# While the ultimate goal of reinforcement learning is to teach agents to act in the real, physical world, games provide a convenient proving ground for developing RL algorithms and agents. Games have some properties that make them particularly well suited for RL:
#
# 1. In many cases, games have perfectly describable environments. For example, all rules of chess can be formally written and programmed into a chess game simulator;
# 2. Games are massively parallelizable. Since they do not require running in the real world, simultaneous environments can be run on large data clusters;
# 3. Simpler scenarios in games enable fast prototyping. This speeds up the development of algorithms that could eventually run in the real-world; and
# 4. ... Games are fun!
#
# In previous labs, we have explored both supervised (with LSTMs, CNNs) and unsupervised / semi-supervised (with VAEs) learning tasks. Reinforcement learning is fundamentally different, in that we are training a deep learning algorithm to govern the actions of our RL agent, that is trying, within its environment, to find the optimal way to achieve a goal. The goal of training an RL agent is to determine the best next step to take to earn the greatest final payoff or return. In this lab, we focus on building a reinforcement learning algorithm to master two different environments with varying complexity.
#
# 1. **Cartpole**: Balance a pole, protruding from a cart, in an upright position by only moving the base left or right. Environment with a low-dimensional observation space.
# 2. [**Pong**](https://en.wikipedia.org/wiki/Pong): Beat your competitors (whether other AI or humans!) at the game of Pong. Environment with a high-dimensional observation space -- learning directly from raw pixels.
#
# Let's get started! First we'll import TensorFlow, the course package, and some dependencies.
#
# + colab={} colab_type="code" id="EvdePP-VyVWp"
# # !apt-get install -y xvfb python-opengl x11-utils > /dev/null 2>&1
# # !pip install gym pyvirtualdisplay scikit-video > /dev/null 2>&1
# # %tensorflow_version 2.x
import tensorflow as tf
import numpy as np
import base64, io, time, gym
import IPython, functools
import matplotlib.pyplot as plt
from tqdm import tqdm
# # !pip install mitdeeplearning
import mitdeeplearning as mdl
# + [markdown] colab_type="text" id="zmrHSiXKTXTY"
# Before we dive in, let's take a step back and outline our approach, which is generally applicable to reinforcement learning problems in general:
#
# 1. **Initialize our environment and our agent**: here we will describe the different observations and actions the agent can make in the environemnt.
# 2. **Define our agent's memory**: this will enable the agent to remember its past actions, observations, and rewards.
# 3. **Define a reward function**: describes the reward associated with an action or sequence of actions.
# 4. **Define the learning algorithm**: this will be used to reinforce the agent's good behaviors and discourage bad behaviors.
#
# + [markdown] colab_type="text" id="UT7YL8KBJIIc"
# # Part 1: Cartpole
#
# ## 3.1 Define the Cartpole environment and agent
#
# ### Environment
#
# In order to model the environment for both the Cartpole and Pong tasks, we'll be using a toolkit developed by OpenAI called [OpenAI Gym](https://gym.openai.com/). It provides several pre-defined environments for training and testing reinforcement learning agents, including those for classic physics control tasks, Atari video games, and robotic simulations. To access the Cartpole environment, we can use `env = gym.make("CartPole-v0")`, which we gained access to when we imported the `gym` package. We can instantiate different [environments](https://gym.openai.com/envs/#classic_control) by passing the enivronment name to the `make` function.
#
# One issue we might experience when developing RL algorithms is that many aspects of the learning process are inherently random: initializing game states, changes in the environment, and the agent's actions. As such, it can be helpful to set a initial "seed" for the environment to ensure some level of reproducibility. Much like you might use `numpy.random.seed`, we can call the comparable function in gym, `seed`, with our defined environment to ensure the environment's random variables are initialized the same each time.
# + colab={} colab_type="code" id="quv9SC0iIYFm"
### Instantiate the Cartpole environment ###
env = gym.make("CartPole-v0")
env.seed(1)
# + [markdown] colab_type="text" id="mhEITUcKK455"
# In Cartpole, a pole is attached by an un-actuated joint to a cart, which moves along a frictionless track. The pole starts upright, and the goal is to prevent it from falling over. The system is controlled by applying a force of +1 or -1 to the cart. A reward of +1 is provided for every timestep that the pole remains upright. The episode ends when the pole is more than 15 degrees from vertical, or the cart moves more than 2.4 units from the center of the track. A visual summary of the cartpole environment is depicted below:
#
# <img width="400px" src="https://danielpiedrahita.files.wordpress.com/2017/02/cart-pole.png"></img>
#
# Given this setup for the environment and the objective of the game, we can think about: 1) what observations help define the environment's state; 2) what actions the agent can take.
#
# First, let's consider the observation space. In this Cartpole environment our observations are:
#
# 1. Cart position
# 2. Cart velocity
# 3. Pole angle
# 4. Pole rotation rate
#
# We can confirm the size of the space by querying the environment's observation space:
#
# + colab={} colab_type="code" id="UVJaEcbdIX82"
n_observations = env.observation_space
print("Environment has observation space =", n_observations)
# + [markdown] colab_type="text" id="ZibGgjrALgPM"
# Second, we consider the action space. At every time step, the agent can move either right or left. Again we can confirm the size of the action space by querying the environment:
# + colab={} colab_type="code" id="qc9SIPxBIXrm"
n_actions = env.action_space.n
print("Number of possible actions that the agent can choose from =", n_actions)
# + [markdown] colab_type="text" id="pPfHME8aRKkb"
# ### Cartpole agent
#
# Now that we have instantiated the environment and understood the dimensionality of the observation and action spaces, we are ready to define our agent. In deep reinforcement learning, a deep neural network defines the agent. This network will take as input an observation of the environment and output the probability of taking each of the possible actions. Since Cartpole is defined by a low-dimensional observation space, a simple feed-forward neural network should work well for our agent. We will define this using the `Sequential` API.
#
# + colab={} colab_type="code" id="W-o_XK4oQ4eu"
### Define the Cartpole agent ###
# Defines a feed-forward neural network
def create_cartpole_model():
model = tf.keras.models.Sequential([
# First Dense layer
tf.keras.layers.Dense(units=32, activation='relu'),
# TODO: Define the last Dense layer, which will provide the network's output.
# Think about the space the agent needs to act in!
tf.keras.layers.Dense(n_actions)
])
return model
cartpole_model = create_cartpole_model()
# + [markdown] colab_type="text" id="d5D5NSIYS2IW"
# Now that we have defined the core network architecture, we will define an *action function* that executes a forward pass through the network, given a set of observations, and samples from the output. This sampling from the output probabilities will be used to select the next action for the agent.
#
# **Critically, this action function is totally general -- we will use this function for both Cartpole and Pong, and it is applicable to other RL tasks, as well!**
# + colab={} colab_type="code" id="E_vVZRr8Q4R_"
### Define the agent's action function ###
# Function that takes observations as input, executes a forward pass through model,
# and outputs a sampled action.
# Arguments:
# model: the network that defines our agent
# observation: observation which is fed as input to the model
# Returns:
# action: choice of agent action
def choose_action(model, observation):
# add batch dimension to the observation
observation = np.expand_dims(observation, axis=0)
'''TODO: feed the observations through the model to predict the log probabilities of each possible action.'''
logits = model.predict(observation)
# pass the log probabilities through a softmax to compute true probabilities
prob_weights = tf.nn.softmax(logits).numpy()
'''TODO: randomly sample from the prob_weights to pick an action.
Hint: carefully consider the dimensionality of the input probabilities (vector) and the output action (scalar)'''
action = np.random.choice(n_actions, size=1, p=prob_weights[0])[0]
return action
# + [markdown] colab_type="text" id="_tR9uAWcTnkr"
# ## 3.2 Define the agent's memory
#
# Now that we have instantiated the environment and defined the agent network architecture and action function, we are ready to move on to the next step in our RL workflow:
# 1. **Initialize our environment and our agent**: here we will describe the different observations and actions the agent can make in the environemnt.
# 2. **Define our agent's memory**: this will enable the agent to remember its past actions, observations, and rewards.
# 3. **Define the learning algorithm**: this will be used to reinforce the agent's good behaviors and discourage bad behaviors.
#
# In reinforcement learning, training occurs alongside the agent's acting in the environment; an *episode* refers to a sequence of actions that ends in some terminal state, such as the pole falling down or the cart crashing. The agent will need to remember all of its observations and actions, such that once an episode ends, it can learn to "reinforce" the good actions and punish the undesirable actions via training. Our first step is to define a simple memory buffer that contains the agent's observations, actions, and received rewards from a given episode.
#
# **Once again, note the modularity of this memory buffer -- it can and will be applied to other RL tasks as well!**
# + colab={} colab_type="code" id="8MM6JwXVQ4JG"
### Agent Memory ###
class Memory:
def __init__(self):
self.clear()
# Resets/restarts the memory buffer
def clear(self):
self.observations = []
self.actions = []
self.rewards = []
# Add observations, actions, rewards to memory
def add_to_memory(self, new_observation, new_action, new_reward):
self.observations.append(new_observation)
self.actions.append(new_action)
# TODO: your update code here
self.rewards.append(new_reward)
# TODO: your update code here
memory = Memory()
# + [markdown] colab_type="text" id="D<KEY>"
# ## 3.3 Reward function
#
# We're almost ready to begin the learning algorithm for our agent! The next step is to compute the rewards of our agent as it acts in the environment. Since we (and the agent) is uncertain about if and when the game or task will end (i.e., when the pole will fall), it is useful to emphasize getting rewards **now** rather than later in the future -- this is the idea of discounting. This is a similar concept to discounting money in the case of interest. ecall from lecture, we use reward discount to give more preference at getting rewards now rather than later in the future. The idea of discounting rewards is similar to discounting money in the case of interest.
#
# To compute the expected cumulative reward, known as the **return**, at a given timestep in a learning episode, we sum the discounted rewards expected at that time step $t$, within a learning episode, and projecting into the future. We define the return (cumulative reward) at a time step $t$, $R_{t}$ as:
#
# >$R_{t}=\sum_{k=0}^\infty\gamma^kr_{t+k}$
#
# where $0 < \gamma < 1$ is the discount factor and $r_{t}$ is the reward at time step $t$, and the index $k$ increments projection into the future within a single learning episode. Intuitively, you can think of this function as depreciating any rewards received at later time steps, which will force the agent prioritize getting rewards now. Since we can't extend episodes to infinity, in practice the computation will be limited to the number of timesteps in an episode -- after that the reward is assumed to be zero.
#
# Take note of the form of this sum -- we'll have to be clever about how we implement this function. Specifically, we'll need to initialize an array of zeros, with length of the number of time steps, and fill it with the real discounted reward values as we loop through the rewards from the episode, which will have been saved in the agents memory. What we ultimately care about is which actions are better relative to other actions taken in that episode -- so, we'll normalize our computed rewards, using the mean and standard deviation of the rewards across the learning episode.
#
# + colab={} colab_type="code" id="5_Q2OFYtQ32X"
### Reward function ###
# Helper function that normalizes an np.array x
def normalize(x):
x -= np.mean(x)
x /= np.std(x)
return x.astype(np.float32)
# Compute normalized, discounted, cumulative rewards (i.e., return)
# Arguments:
# rewards: reward at timesteps in episode
# gamma: discounting factor
# Returns:
# normalized discounted reward
def discount_rewards(rewards, gamma=0.95):
discounted_rewards = np.zeros_like(rewards)
R = 0
for t in reversed(range(0, len(rewards))):
# update the total discounted reward
R = R * gamma + rewards[t]
discounted_rewards[t] = R
return normalize(discounted_rewards)
# + [markdown] colab_type="text" id="QzbY-mjGYcmt"
# ## 3.4 Learning algorithm
#
# Now we can start to define the learing algorithm which will be used to reinforce good behaviors of the agent and discourage bad behaviours. In this lab, we will focus on *policy gradient* methods which aim to **maximize** the likelihood of actions that result in large rewards. Equivalently, this means that we want to **minimize** the negative likelihood of these same actions. We achieve this by simply **scaling** the probabilities by their associated rewards -- effectively amplifying the likelihood of actions that resujlt in large rewards.
#
# Since the log function is monotonically increasing, this means that minimizing **negative likelihood** is equivalent to minimizing **negative log-likelihood**. Recall that we can easily compute the negative log-likelihood of a discrete action by evaluting its [softmax cross entropy](https://www.tensorflow.org/api_docs/python/tf/nn/sparse_softmax_cross_entropy_with_logits). Like in supervised learning, we can use stochastic gradient descent methods to achieve the desired minimization.
#
# Let's begin by defining the loss function.
# + colab={} colab_type="code" id="fsgZ3IDCY_Zn"
### Loss function ###
# Arguments:
# logits: network's predictions for actions to take
# actions: the actions the agent took in an episode
# rewards: the rewards the agent received in an episode
# Returns:
# loss
def compute_loss(logits, actions, rewards):
neg_logprob = tf.nn.sparse_softmax_cross_entropy_with_logits(logits=logits, labels=actions)
loss = tf.reduce_mean(neg_logprob * rewards)
return loss
# + [markdown] colab_type="text" id="Rr5vQ9fqbPpp"
# Now let's use the loss function to define a training step of our learning algorithm:
# + colab={} colab_type="code" id="_50ada7nbZ7L"
### Training step (forward and backpropagation) ###
def train_step(model, optimizer, observations, actions, discounted_rewards):
with tf.GradientTape() as tape:
# Forward propagate through the agent network
logits = model(observations)
loss = compute_loss(logits, actions, discounted_rewards)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# + [markdown] colab_type="text" id="XsjKXh6BcgjR"
# ## 3.5 Run cartpole!
#
# Having had no prior knowledge of the environment, the agent will begin to learn how to balance the pole on the cart based only on the feedback received from the environment! Having defined how our agent can move, how it takes in new observations, and how it updates its state, we'll see how it gradually learns a policy of actions to optimize balancing the pole as long as possible. To do this, we'll track how the rewards evolve as a function of training -- how should the rewards change as training progresses?
# + colab={} colab_type="code" id="XmOzc2rrcn8Q"
### Cartpole training! ###
# Learning rate and optimizer
learning_rate = 1e-3
optimizer = tf.keras.optimizers.Adam(learning_rate)
# instantiate cartpole agent
cartpole_model = create_cartpole_model()
# to track our progress
smoothed_reward = mdl.util.LossHistory(smoothing_factor=0.9)
plotter = mdl.util.PeriodicPlotter(sec=2, xlabel='Iterations', ylabel='Rewards')
if hasattr(tqdm, '_instances'): tqdm._instances.clear() # clear if it exists
for i_episode in range(500):
plotter.plot(smoothed_reward.get())
# Restart the environment
observation = env.reset()
memory.clear()
while True:
# using our observation, choose an action and take it in the environment
action = choose_action(cartpole_model, observation)
next_observation, reward, done, info = env.step(action)
# add to memory
memory.add_to_memory(observation, action, reward)
# is the episode over? did you crash or do so well that you're done?
if done:
# determine total reward and keep a record of this
total_reward = sum(memory.rewards)
smoothed_reward.append(total_reward)
# initiate training - remember we don't know anything about how the
# agent is doing until it has crashed!
train_step(cartpole_model, optimizer,
observations=np.vstack(memory.observations),
actions=np.array(memory.actions),
discounted_rewards = discount_rewards(memory.rewards))
# reset the memory
memory.clear()
break
# update our observatons
observation = next_observation
# + [markdown] colab_type="text" id="mkcUtGF1VE-K"
# To get a sense of how our agent did, we can save a video of the trained model working on balancing the pole. Realize that this is a brand new environment that the agent has not seen before!
#
# Let's display the saved video to watch how our agent did!
#
# + colab={} colab_type="code" id="PAYBkv6Zbk0J"
saved_cartpole = mdl.lab3.save_video_of_model(cartpole_model, "CartPole-v0")
mdl.lab3.play_video(saved_cartpole)
# + [markdown] colab_type="text" id="CSbVNDpaVb3_"
# How does the agent perform? Could you train it for shorter amounts of time and still perform well? Do you think that training longer would help even more?
# + [markdown] colab_type="text" id="Eu6Mqxc720ST"
# #Part 2: Pong
#
# In Cartpole, we dealt with an environment that was static -- in other words, it didn't change over time. What happens if our environment is dynamic and unpredictable? Well that's exactly the case in [Pong](https://en.wikipedia.org/wiki/Pong), since part of the environment is the opposing player. We don't know how our opponent will act or react to our actions, so the complexity of our problem increases. It also becomes much more interesting, since we can compete to beat our opponent. RL provides a powerful framework for training AI systems with the ability to handle and interact with dynamic, unpredictable environments. In this part of the lab, we'll use the tools and workflow we explored in Part 1 to build an RL agent capable of playing the game of Pong.
#
# + [markdown] colab_type="text" id="srZ4YE29isuA"
# ## 3.6 Define and inspect the Pong environment
#
# As with Cartpole, we'll instantiate the Pong environment in the OpenAI gym, using a seed of 1.
# + colab={} colab_type="code" id="lbYHLr66i15n"
env = gym.make("Pong-v0", frameskip=5)
env.seed(1); # for reproducibility
# + [markdown] colab_type="text" id="52uZ2Xhyi-MW"
# Let's next consider the observation space for the Pong environment. Instead of four physical descriptors of the cart-pole setup, in the case of Pong our observations are the individual video frames (i.e., images) that depict the state of the board. Thus, the observations are 210x160 RGB images (arrays of shape (210,160,3)).
#
# We can again confirm the size of the observation space by query:
# + colab={} colab_type="code" id="0yX4GWvxjnHS"
print("Environment has observation space =", env.observation_space)
# + [markdown] colab_type="text" id="uuEC2TdSjx9D"
# In Pong, at every time step, the agent (which controls the paddle) has six actions to choose from: no-op (no operation), move right, move left, fire, fire right, and fire left. Let's confirm the size of the action space by querying the environment:
# + colab={} colab_type="code" id="Iuy9oPc1kag3"
n_actions = env.action_space.n
print("Number of possible actions that the agent can choose from =", n_actions)
# + [markdown] colab_type="text" id="9-fghDRigUE5"
# ## 3.7 Define the Pong agent
#
# As before, we'll use a neural network to define our agent. What network architecture do you think would be especially well suited to this game? Since our observations are now in the form of images, we'll add convolutional layers to the network to increase the learning capacity of our network.
# + colab={} colab_type="code" id="IJiqbFYpgYRH"
### Define the Pong agent ###
# Functionally define layers for convenience
# All convolutional layers will have ReLu activation
Conv2D = functools.partial(tf.keras.layers.Conv2D, padding='same', activation='relu')
Flatten = tf.keras.layers.Flatten
Dense = tf.keras.layers.Dense
# Defines a CNN for the Pong agent
def create_pong_model():
model = tf.keras.models.Sequential([
# Convolutional layers
# First, 16 7x7 filters with 4x4 stride
Conv2D(filters=16, kernel_size=7, strides=4),
# TODO: define convolutional layers with 32 5x5 filters and 2x2 stride
Conv2D(filters=32, kernel_size=5, strides=2),
# TODO: define convolutional layers with 48 3x3 filters and 2x2 stride
Conv2D(filters=48, kernel_size=3, strides=2),
Flatten(),
# Fully connected layer and output
Dense(units=64, activation='relu'),
# TODO: define the output dimension of the last Dense layer.
# Pay attention to the space the agent needs to act in
Dense(n_actions)
])
return model
pong_model = create_pong_model()
# + [markdown] colab_type="text" id="yaeZ067olFiJ"
# Since we've already defined the action function, `choose_action(model, observation)`, we don't need to define it again. Instead, we'll be able to reuse it later on by passing in our new model we've just created, `pong_model`. This is awesome because our action function provides a modular and generalizable method for all sorts of RL agents!
# + [markdown] colab_type="text" id="l0RvqOVkmc2r"
# ## 3.8 Pong-specific functions
#
# In Part 1 (Cartpole), we implemented some key functions and classes to build and train our RL agent -- `choose_action(model, observation)` and the `Memory` class, for example. However, in getting ready to apply these to a new game like Pong, we might need to make some slight modifications.
#
# Namely, we need to think about what happens when a game ends. In Pong, we know a game has ended if the reward is +1 (we won!) or -1 (we lost unfortunately). Otherwise, we expect the reward at a timestep to be zero -- the players (or agents) are just playing eachother. So, after a game ends, we will need to reset the reward to zero when a game ends. This will result in a modified reward function.
# + colab={} colab_type="code" id="iEZG2o50luLu"
### Pong reward function ###
# Compute normalized, discounted rewards for Pong (i.e., return)
# Arguments:
# rewards: reward at timesteps in episode
# gamma: discounting factor. Note increase to 0.99 -- rate of depreciation will be slower.
# Returns:
# normalized discounted reward
def discount_rewards(rewards, gamma=0.99):
discounted_rewards = np.zeros_like(rewards)
R = 0
for t in reversed(range(0, len(rewards))):
# NEW: Reset the sum if the reward is not 0 (the game has ended!)
if rewards[t] != 0:
R = 0
# update the total discounted reward as before
R = R * gamma + rewards[t]
discounted_rewards[t] = R
return normalize(discounted_rewards)
# + [markdown] colab_type="text" id="HopLpb4IoOqA"
# Additionally, we have to consider the nature of the observations in the Pong environment, and how they will be fed into our network. Our observations in this case are images. Before we input an image into our network, we'll do a bit of pre-processing to crop and scale, clean up the background colors to a single color, and set the important game elements to a single color. Let's use this function to visualize what an observation might look like before and after pre-processing.
# + colab={} colab_type="code" id="no5IIYtFm8pI"
observation = env.reset()
for i in range(30):
observation, _,_,_ = env.step(0)
observation_pp = mdl.lab3.preprocess_pong(observation)
f = plt.figure(figsize=(10,3))
ax = f.add_subplot(121)
ax2 = f.add_subplot(122)
ax.imshow(observation); ax.grid(False);
ax2.imshow(np.squeeze(observation_pp)); ax2.grid(False); plt.title('Preprocessed Observation');
# + [markdown] colab_type="text" id="bYwIWC-Cz8F2"
# What do you notice? How might these changes be important for training our RL algorithm?
# + [markdown] colab_type="text" id="mRqcaDQ1pm3x"
# ## 3.9 Training Pong
#
# We're now all set up to start training our RL algorithm and agent for the game of Pong! We've already defined our loss function with `compute_loss`, which employs policy gradient learning, as well as our backpropagation step with `train_step` which is beautiful! We will use these functions to execute training the Pong agent. Let's walk through the training block.
#
# In Pong, rather than feeding our network one image at a time, it can actually improve performance to input the difference between two consecutive observations, which really gives us information about the movement between frames -- how the game is changing. We'll first pre-process the raw observation, `x`, and then we'll compute the difference with the image frame we saw one timestep before.
#
# This observation change will be forward propagated through our Pong agent, the CNN network model, which will then predict the next action to take based on this observation. The raw reward will be computed, and the observation, action, and reward will be recorded into memory. This will continue until a training episode, i.e., a game, ends.
#
# Then, we will compute the discounted rewards, and use this information to execute a training step. Memory will be cleared, and we will do it all over again!
#
# Let's run the code block to train our Pong agent. Note that completing training will take quite a bit of time (estimated at least a couple of hours). We will again visualize the evolution of the total reward as a function of training to get a sense of how the agent is learning.
# + colab={} colab_type="code" id="xCwyQQrPnkZG"
### Training Pong ###
# Hyperparameters
learning_rate=1e-4
MAX_ITERS = 10000 # increase the maximum number of episodes, since Pong is more complex!
# Model and optimizer
pong_model = create_pong_model()
optimizer = tf.keras.optimizers.Adam(learning_rate)
# plotting
smoothed_reward = mdl.util.LossHistory(smoothing_factor=0.9)
plotter = mdl.util.PeriodicPlotter(sec=5, xlabel='Iterations', ylabel='Rewards')
memory = Memory()
for i_episode in range(MAX_ITERS):
plotter.plot(smoothed_reward.get())
# Restart the environment
observation = env.reset()
previous_frame = mdl.lab3.preprocess_pong(observation)
while True:
# Pre-process image
current_frame = mdl.lab3.preprocess_pong(observation)
'''TODO: determine the observation change
Hint: this is the difference between the past two frames'''
obs_change = current_frame - previous_frame
'''TODO: choose an action for the pong model, using the frame difference, and evaluate'''
action = choose_action(pong_model, obs_change)
# Take the chosen action
next_observation, reward, done, info = env.step(action)
'''TODO: save the observed frame difference, the action that was taken, and the resulting reward!'''
memory.add_to_memory(obs_change, action, reward)
# is the episode over? did you crash or do so well that you're done?
if done:
# determine total reward and keep a record of this
total_reward = sum(memory.rewards)
smoothed_reward.append( total_reward )
# begin training
train_step(pong_model,
optimizer,
observations = np.stack(memory.observations, 0),
actions = np.array(memory.actions),
discounted_rewards = discount_rewards(memory.rewards))
memory.clear()
break
observation = next_observation
previous_frame = current_frame
# + [markdown] colab_type="text" id="8LiEY5Y_ts-Z"
# Finally we can put our trained agent to the test! It will play in a newly instantiated Pong environment against the "computer", a base AI system for Pong. Your agent plays as the green paddle. Let's watch the match instant replay!
# + colab={} colab_type="code" id="TvHXbkL0tR6M"
saved_pong = mdl.lab3.save_video_of_model(
pong_model, "Pong-v0", obs_diff=True,
pp_fn=mdl.lab3.preprocess_pong)
mdl.lab3.play_video(saved_pong)
# + [markdown] colab_type="text" id="TIlwIgBP3Js6"
# ## 3.10 Conclusion
#
# That's it! Congratulations on training two RL agents and putting them to the test! We encourage you to consider the following:
#
# * How does the agent perform?
# * Could you train it for shorter amounts of time and still perform well?
# * Do you think that training longer would help even more?
# * How does the complexity of Pong relative to Cartpole alter the rate at which the agent learns and its performance?
# * What are some things you could change about the agent or the learning process to potentially improve performance?
#
# If you want to go further, try to optimize your model to achieve the best performance! **[Email us](mailto:<EMAIL>) a copy of your notebook with the Pong training executed AND a saved video of your Pong agent competing! We'll give out prizes to the best performers!**
| lab3/RL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.svm import SVC
from sklearn import metrics
from mlxtend.plotting import plot_decision_regions
from sklearn import preprocessing
from sklearn.linear_model import LogisticRegression
from ast import literal_eval
import warnings
import numpy as np
from collections import OrderedDict
from lob_data_utils import lob, db_result, model
from lob_data_utils.svm_calculation import lob_svm
import os
sns.set_style('whitegrid')
warnings.filterwarnings('ignore')
# -
data_length = 10000
stocks = ['9064', '9061', '9265']
def convert_scores(df, column):
scores = []
for i, row in df.iterrows():
try:
scores.append(np.mean(row[column]))
except:
scores.append(np.mean(np.array(literal_eval(row[column])).astype(np.float64)))
return scores
scores_columns = ['f1', 'kappa', 'matthews', 'precision', 'recall', 'roc_auc', 'train_f1', 'train_kappa',
'train_matthews', 'train_precision', 'train_recall', 'train_roc_auc']
dfs = {}
dfs_test = {}
dfs_reg = {}
dfs_reg_test = {}
data_dir='../gaussian_filter/data_gdf'
for stock in stocks:
r = 0.1
s = 0.1
gdf_filename = 'gdf_{}_len{}_r{}_s{}_K50'.format(stock, data_length, r, s)
reg_filename = '{}'.format(stock)
print(gdf_filename)
dfs[stock], dfs_test[stock] = lob.load_prepared_data(
gdf_filename, data_dir=data_dir, cv=False, length=data_length)
dfs_reg[stock], dfs_reg_test[stock] = lob.load_prepared_data(
reg_filename, data_dir='../gaussian_filter/data', cv=False, length=data_length)
df = dfs[stock]
feature_columns = ['gdf_24', 'gdf_25']
res = {}
for C in [1, 10, 100, 1000, 1100]:
for g in [0.1, 1, 2, 5]:
clf = SVC(kernel='rbf', C=C, gamma=g)
scores = model.validate_model(clf, df[feature_columns], df['mid_price_indicator'])
df_score = pd.DataFrame(scores)
res['C{}-g{}'.format(C, g)] = np.mean(df_score['matthews'].values)
res
df = dfs[stock]
feature_columns = ['gdf_16', 'gdf_33']
res = {}
for C in [1000, 1100, 1500, 10000]:
for g in [0.20, 0.25, 0.30]:
clf = SVC(kernel='rbf', C=1000, gamma=g)
scores = model.validate_model(clf, df[feature_columns], df['mid_price_indicator'])
df_score = pd.DataFrame(scores)
res['C{}-g{}'.format(C, g)] = np.mean(df_score['matthews'].values)
res
cols = feature_columns + ['mid_price_indicator'] + ['gdf_24', 'gdf_25', 'gdf_26']
print(cols)
sns.heatmap(df[cols].corr(), annot=True)
# +
from sklearn.decomposition import PCA
X = df[[c for c in df.columns if 'gdf' in c]]
pca = PCA(n_components=3)
pca.fit(X)
X_transf = pca.transform(X)
res = {}
for C in [1000, 1100, 1500, 10000]:
for g in [0.20, 0.25, 0.30]:
clf = SVC(kernel='rbf', C=1000, gamma=g)
scores = model.validate_model(clf, X_transf, df['mid_price_indicator'])
df_score = pd.DataFrame(scores)
res['C{}-g{}'.format(C, g)] = np.mean(df_score['matthews'].values)
res
# -
dfs_reg[stock].head()
plt.figure(figsize=(24,4))
plt.plot(dfs[stock].iloc[2000][['gdf_{}'.format(i) for i in range(0, 50)]])
for stock in stocks:
dfs[stock]['queue_imbalance'] = dfs_reg[stock]['queue_imbalance']
dfs[stock]['prev_queue_imbalance'] = dfs[stock]['queue_imbalance'].shift()
dfs[stock].dropna(inplace=True)
dfs_test[stock]['queue_imbalance'] = dfs_reg_test[stock]['queue_imbalance']
dfs_test[stock]['prev_queue_imbalance'] = dfs_test[stock]['queue_imbalance'].shift()
dfs_test[stock].dropna(inplace=True)
cols = feature_columns + ['mid_price_indicator'] + ['gdf_24', 'gdf_25', 'gdf_26'] + ['queue_imbalance', 'prev_queue_imbalance']
print(cols)
plt.figure(figsize=(8,8))
sns.heatmap(dfs[stock][cols].corr(), annot=True)
feature_columns_dict = {
'gdf_24-26_que_prev': ['gdf_24', 'gdf_25', 'queue_imbalance', 'prev_queue_imbalance'],
'que': ['queue_imbalance'],
'que_prev': ['queue_imbalance', 'prev_queue_imbalance'],
'gdf_24-26_que': ['gdf_24', 'gdf_25', 'queue_imbalance'],
'gdf_23-27_que': ['gdf_23', 'gdf_24', 'gdf_25', 'gdf_26', 'queue_imbalance'],
'gdf_23-27': ['gdf_23', 'gdf_24', 'gdf_25', 'gdf_26'],
'gdf_24_26': ['gdf_24', 'gdf_25'],
'pca_gdf': ['gdf_{}'.format(i) for i in range(0, 50)],
'pca_gdf_que': ['gdf_{}'.format(i) for i in range(0, 50)] + ['queue_imbalance'],
'pca_gdf_que_prev': ['gdf_{}'.format(i) for i in range(0, 50)] + ['queue_imbalance', 'prev_queue_imbalance'],
'gdf_20_30': ['gdf_{}'.format(i) for i in range(20, 30)],
'gdf_20_30_que': ['gdf_{}'.format(i) for i in range(20, 30)] + ['queue_imbalance'],
'gdf_20_30_que_prev': ['gdf_{}'.format(i) for i in range(20, 30)] + ['queue_imbalance', 'prev_queue_imbalance']
}
res = []
for stock in stocks:
df = dfs[stock]
df_test = dfs_test[stock]
print(stock)
for k, v in feature_columns_dict.items():
C = 1
gamma = 1
train_x = df[v]
if 'pca' in k:
X = df[v]
for i in range(2, 10):
pca = PCA(n_components=i)
pca.fit(X)
train_x = pca.transform(X)
clf = SVC(kernel='sigmoid')
scores = model.validate_model(clf, train_x, df['mid_price_indicator'])
df_score = pd.DataFrame(scores)
res.append({'matthews': np.mean(df_score['matthews'].values),
'stock': stock,
'C': C,
'gamma': gamma,
'features': k + str(i)
})
continue
clf = SVC(kernel='rbf', C=C, gamma=gamma)
scores = model.validate_model(clf, train_x, df['mid_price_indicator'])
#scores_test = model.test_model(clf, df_test[feature_columns], df_test['mid_price_indicator'])
df_score = pd.DataFrame(scores)
res.append({'matthews': np.mean(df_score['matthews'].values),
#'matthews_test': scores_test['test_matthews'],
'stock': stock,
'C': C,
'gamma': gamma,
'features': k
})
df_results = pd.DataFrame(res)
df_results.sort_values(by='matthews', ascending=False).groupby('stock').head(1)
## let's train SVM!
stock = '9265'
df = dfs[stock]
res = []
for C in [0.001, 0.01, 0.1, 1, 10, 100, 1000]:
for g in [0.001, 0.01, 0.1, 1, 10, 100, 1000]:
for coef0 in [0.01, 0.1, 1, 10, 100]:
clf = SVC(kernel='sigmoid', C=C, gamma=g, coef0=coef0)
X = df[feature_columns_dict['pca_gdf_que']]
pca = PCA(n_components=3)
pca.fit(X)
train_x = pca.transform(X)
scores = model.validate_model(clf, train_x, df['mid_price_indicator'])
df_score = pd.DataFrame(scores)
res.append({'matthews': np.mean(df_score['matthews'].values),
'stock': stock,
'C': C,
'coef0': coef0,
'gamma': g,
'features': feature_columns
})
df_results_9265 = pd.DataFrame(res)
df_results_9265.sort_values(by='matthews', ascending=False)
## let's train SVM!
stock = '9061'
df = dfs[stock]
res = []
for C in [0.001, 0.01, 0.1, 1, 10, 100, 1000]:
for g in [0.001, 0.01, 0.1, 1, 10, 100, 1000]:
for coef0 in [0.01, 0.1, 1, 10, 100]:
clf = SVC(kernel='sigmoid', C=C, gamma=g, coef0=coef0)
X = df[feature_columns_dict['pca_gdf_que']]
pca = PCA(n_components=4)
pca.fit(X)
train_x = pca.transform(X)
scores = model.validate_model(clf, train_x, df['mid_price_indicator'])
df_score = pd.DataFrame(scores)
res.append({'matthews': np.mean(df_score['matthews'].values),
'stock': stock,
'C': C,
'coef0': coef0,
'gamma': g,
'features': feature_columns
})
df_results_9061 = pd.DataFrame(res)
df_results_9061.sort_values(by='matthews', ascending=False)
stock = '9064'
df = dfs[stock]
res = []
for C in [0.001, 0.01, 0.1, 1, 10, 100, 1000]:
for g in [0.001, 0.01, 0.1, 1, 10, 100, 1000]:
for coef0 in [0.01, 0.1, 1, 10, 100]:
clf = SVC(kernel='sigmoid', C=C, gamma=g, coef0=coef0)
X = df[feature_columns_dict['pca_gdf_que_prev']]
pca = PCA(n_components=4)
pca.fit(X)
train_x = pca.transform(X)
scores = model.validate_model(clf, train_x, df['mid_price_indicator'])
df_score = pd.DataFrame(scores)
res.append({'matthews': np.mean(df_score['matthews'].values),
'stock': stock,
'C': C,
'coef0': coef0,
'gamma': g,
'features': feature_columns
})
df_results_9064 = pd.DataFrame(res)
df_results_9064.sort_values(by='matthews', ascending=False)
df_results_9061.to_csv('res_svm_training/res_overview_all_svm_gdf-small_r_s-sigmoid_kernel_{}.csv'.format(9061))
df_results_9064.to_csv('res_svm_training/res_overview_all_svm_gdf-small_r_s-sigmoid_kernel_{}.csv'.format(9064))
df_results_9265.to_csv('res_svm_training/res_overview_all_svm_gdf-small_r_s-sigmoid_kernel_{}.csv'.format(9265))
df_results.to_csv('res_svm_training/res_features_overview_all_svm_gdf-small_r_s-sigmoid_kernel.csv')
| overview_val10/overview_junk/overview_all_svm_gdf-small_r_s-sigmoid_kernel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Optimizing the SVM Classifier
#
# Machine learning models are parameterized so that their behavior can be tuned for a given problem. Models can have many parameters and finding the best combination of parameters can be treated as a search problem. In this notebook, I aim to tune parameters of the SVM Classification model using scikit-learn.
#
# #### Load Libraries and Data
# +
# %matplotlib inline
import matplotlib.pyplot as plt
#Load libraries for data processing
import pandas as pd #data processing, CSV file I/O (e.g. pd.read_csv)
import numpy as np
from scipy.stats import norm
## Supervised learning.
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import make_pipeline
from sklearn.metrics import confusion_matrix
from sklearn import metrics, preprocessing
from sklearn.metrics import classification_report
from sklearn.feature_selection import SelectKBest, f_regression
# visualization
import seaborn as sns
plt.style.use('fivethirtyeight')
sns.set_style("white")
plt.rcParams['figure.figsize'] = (8,4)
#plt.rcParams['axes.titlesize'] = 'large'
# -
# #### Build a predictive model and evaluate with 5-cross validation using support vector classifies (ref NB4) for details
#
# +
data = pd.read_csv('data/clean-data.csv', index_col=False)
data.drop('Unnamed: 0',axis=1, inplace=True)
#Assign predictors to a variable of ndarray (matrix) type
array = data.values
X = array[:,1:31]
y = array[:,0]
#transform the class labels from their original string representation (M and B) into integers
le = LabelEncoder()
y = le.fit_transform(y)
# Normalize the data (center around 0 and scale to remove the variance).
scaler =StandardScaler()
Xs = scaler.fit_transform(X)
from sklearn.decomposition import PCA
# feature extraction
pca = PCA(n_components=10)
fit = pca.fit(Xs)
X_pca = pca.transform(Xs)
# 5. Divide records in training and testing sets.
X_train, X_test, y_train, y_test = train_test_split(X_pca, y, test_size=0.3, random_state=2, stratify=y)
# 6. Create an SVM classifier and train it on 70% of the data set.
clf = SVC(probability=True)
clf.fit(X_train, y_train)
#7. Analyze accuracy of predictions on 30% of the holdout test sample.
classifier_score = clf.score(X_test, y_test)
print '\nThe classifier accuracy score is {:03.2f}\n'.format(classifier_score)
clf2 = make_pipeline(SelectKBest(f_regression, k=3),SVC(probability=True))
scores = cross_val_score(clf2, X_pca, y, cv=3)
# Get average of 5-fold cross-validation score using an SVC estimator.
n_folds = 5
cv_error = np.average(cross_val_score(SVC(), X_pca, y, cv=n_folds))
print '\nThe {}-fold cross-validation accuracy score for this classifier is {:.2f}\n'.format(n_folds, cv_error)
y_pred = clf.fit(X_train, y_train).predict(X_test)
cm = metrics.confusion_matrix(y_test, y_pred)
print(classification_report(y_test, y_pred ))
fig, ax = plt.subplots(figsize=(5, 5))
ax.matshow(cm, cmap=plt.cm.Reds, alpha=0.3)
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(x=j, y=i,
s=cm[i, j],
va='center', ha='center')
plt.xlabel('Predicted Values', )
plt.ylabel('Actual Values')
plt.show()
# -
# ## Importance of optimizing a classifier
#
# We can tune two key parameters of the SVM algorithm:
# * the value of C (how much to relax the margin)
# * and the type of kernel.
#
# The default for SVM (the SVC class) is to use the Radial Basis Function (RBF) kernel with a C value set to 1.0. Like with KNN, we will perform a grid search using 10-fold cross validation with a standardized copy of the training dataset. We will try a number of simpler kernel types and C values with less bias and more bias (less than and more than 1.0 respectively).
#
# Python scikit-learn provides two simple methods for algorithm parameter tuning:
# * Grid Search Parameter Tuning.
# * Random Search Parameter Tuning.
# +
# Train classifiers.
kernel_values = [ 'linear' , 'poly' , 'rbf' , 'sigmoid' ]
param_grid = {'C': np.logspace(-3, 2, 6), 'gamma': np.logspace(-3, 2, 6),'kernel': kernel_values}
grid = GridSearchCV(SVC(), param_grid=param_grid, cv=5)
grid.fit(X_train, y_train)
# -
print("The best parameters are %s with a score of %0.2f"
% (grid.best_params_, grid.best_score_))
grid.best_estimator_.probability = True
clf = grid.best_estimator_
# +
y_pred = clf.fit(X_train, y_train).predict(X_test)
cm = metrics.confusion_matrix(y_test, y_pred)
#print(cm)
print(classification_report(y_test, y_pred ))
fig, ax = plt.subplots(figsize=(5, 5))
ax.matshow(cm, cmap=plt.cm.Reds, alpha=0.3)
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(x=j, y=i,
s=cm[i, j],
va='center', ha='center')
plt.xlabel('Predicted Values', )
plt.ylabel('Actual Values')
plt.show()
# -
# ### Decision boundaries of different classifiers
# Let's see the decision boundaries produced by the linear, Gaussian and polynomial classifiers.
# +
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn import svm, datasets
def decision_plot(X_train, y_train, n_neighbors, weights):
h = .02 # step size in the mesh
Xtrain = X_train[:, :2] # we only take the first two features.
#================================================================
# Create color maps
#================================================================
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
#================================================================
# we create an instance of SVM and fit out data.
# We do not scale ourdata since we want to plot the support vectors
#================================================================
C = 1.0 # SVM regularization parameter
svm = SVC(kernel='linear', random_state=0, gamma=0.1, C=C).fit(Xtrain, y_train)
rbf_svc = SVC(kernel='rbf', gamma=0.7, C=C).fit(Xtrain, y_train)
poly_svc = SVC(kernel='poly', degree=3, C=C).fit(Xtrain, y_train)
# +
# %matplotlib inline
plt.rcParams['figure.figsize'] = (15, 9)
plt.rcParams['axes.titlesize'] = 'large'
# create a mesh to plot in
x_min, x_max = Xtrain[:, 0].min() - 1, Xtrain[:, 0].max() + 1
y_min, y_max = Xtrain[:, 1].min() - 1, Xtrain[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.1),
np.arange(y_min, y_max, 0.1))
# title for the plots
titles = ['SVC with linear kernel',
'SVC with RBF kernel',
'SVC with polynomial (degree 3) kernel']
# +
for i, clf in enumerate((svm, rbf_svc, poly_svc)):
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
plt.subplot(2, 2, i + 1)
plt.subplots_adjust(wspace=0.4, hspace=0.4)
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.coolwarm, alpha=0.8)
# Plot also the training points
plt.scatter(Xtrain[:, 0], Xtrain[:, 1], c=y_train, cmap=plt.cm.coolwarm)
plt.xlabel('radius_mean')
plt.ylabel('texture_mean')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.xticks(())
plt.yticks(())
plt.title(titles[i])
plt.show()
# -
# ## Conclusion
#
# This work demonstrates the modelling of breast cancer as classification task using Support Vector Machine
#
# The SVM performs better when the dataset is standardized so that all attributes have a mean value of zero and a standard deviation of one. We can calculate this from the entire training dataset and apply the same transform to the input attributes from the validation dataset.
# Next Task:
# 1. Summary and conclusion of findings
# 2. Compare with other classification methods
# * Decision trees with tree.DecisionTreeClassifier();
# * K-nearest neighbors with neighbors.KNeighborsClassifier();
# * Random forests with ensemble.RandomForestClassifier();
# * Perceptron (both gradient and stochastic gradient) with mlxtend.classifier.Perceptron; and
# * Multilayer perceptron network (both gradient and stochastic gradient) with mlxtend.classifier.MultiLayerPerceptron.
#
| NB5 OptimizingSVMClassifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false run_control={"frozen": true}
# #### Sources
# - 鏈接在此: [link](https://docs.python.org/3/library/secrets.html)
# - 注意: **新于 Python 3.7.0 出現的函數 不會在此測試**.
# -
before_using_it = '''
The 'secrets' module is used for
generating cryptographically strong random numbers
which is suitable for managing data such as pwd, account auth etc.
In particularly, it should be
used in preference to the default pseudo-random generator in 'random',
it's designed for modelling and simulation, not security or cryptography.
----- In short, you SHOULD USE 'secrets' module. ----
'''
# +
import random
import secrets
# use these by creating an instance
rsr = random.SystemRandom()
ssr = secrets.SystemRandom()
# the same? I didn't check each one of them
len(rsr.__dir__()), len(ssr.__dir__())
# -
# #### random numbers
# +
options = [
'Base64',
'Descriptor',
'BlockChain',
'Cryptography',
]
secrets.choice(options)
secrets.randbelow(30) # an random int in [0, n)
secrets.randbits(1) # an int with k random bits
secrets.randbits(120)
# -
# #### generating tokens
# +
aim = '''
The secure tokens is suitable for
applications such password resets, hard-to-guess urls, and similar.
'''
tip = '''
The default 'nbytes' is ant integer.
For now, the 32 bytes is the default.
Of course, you can specify your own token length.
It was used to make tokens have sufficient randomness (against attack).
And note:
the default is subject to change at any time,
including during maintenance releases. (XD)
'''
# nbytes=32
tb1 = secrets.token_bytes() # byte string
tb2 = secrets.token_bytes(20)
# nbytes=64
th1 = secrets.token_hex() # text string in hexadecimal
th2 = secrets.token_hex(20)
# nbytes=43
tu1 = secrets.token_urlsafe() # random URL-safe text string
tu2 = secrets.token_urlsafe(9)
# default
tb1 # token_bytes()
th1 # token_hex()
tu1 # token_urlsafe()
# compare!
secrets.compare_digest(tu1,tu1)
# -
# #### recipes and best practices
# +
from string import ascii_letters, digits
from secrets import choice
a_basic_password = '''
generate an 12-char alphanumeric password
'''
alphab = ascii_letters + digits
passwd = ''.join(choice(alphab) for i in range(12))
# Btw, applications should NOT
# store pwds in a Recoverable format, whether plain text or encrypted.
# They should be Salted and Hashed
# using a cryptographically-strong one-way(irreversible) hash function.
passwd
# +
create_a_password = '''
Generate a ten-char alphanumeric pwd
with at least one lowercase char,
at lease one uppercase char,
and at lease three digits.
'''
alphab = ascii_letters + digits
while True:
password = ''.join(choice(alphab) for i in range(10))
if ( any( c.islower() for c in password )
and any( c.isupper() for c in password )
and sum( c.isdigit() for c in password ) >= 3
):
break
password
# +
# Generate a hard-to-guess temporary
# URL containing a security token
# suitable for password recovery applications
url = 'https://example.com/reset=' + secrets.token_urlsafe(10)
url
| cryptographic_services/secrets_offidoc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="cb6oxCjrNaIR"
# # Spectrally Normalized Generative Adversarial Networks (SN-GAN)
#
# *Please note that this is an optional notebook, meant to introduce more advanced concepts if you're up for a challenge, so don't worry if you don't completely follow!*
#
# **Goals**
#
# In this notebook, you'll learn about and implement **spectral normalization**, a weight normalization technique to stabilize the training of the discriminator, as proposed in [Spectral Normalization for Generative Adversarial Networks](https://arxiv.org/abs/1802.05957) (Miyato et al. 2018).
#
# **Background**
#
# As its name suggests, SN-GAN normalizes the weight matrices in the discriminator by their corresponding [spectral norm](https://calculus.subwiki.org/wiki/Spectral_norm#:~:text=The%20spectral%20norm%20of%20a,where%20denotes%20the%20Euclidean%20norm.), which helps control the **Lipschitz constant** of the discriminator. As you have learned with WGAN, [Lipschitz continuity](https://en.wikipedia.org/wiki/Lipschitz_continuity) is important in ensuring the boundedness of the optimal discriminator. In the WGAN case, this makes it so that the underlying W-loss function for the discriminator (or more precisely, the critic) is valid.
#
# As a result, spectral normalization helps improve stability and avoid vanishing gradient problems, such as mode collapse.
# + [markdown] id="Q2W91zMySHNM"
# ## Spectral Norm
#
# Notationally, the spectral norm of a matrix $W$ is typically represented as $\sigma(W)$. For neural network purposes, this $W$ matrix represents a weight matrix in one of the network's layers. The spectral norm of a matrix is the matrix's largest singular value, which can be obtained via singular value decomposition (SVD).
#
# **A Quick Refresher on SVD**
#
# SVD is a generalization of [eigendecomposition](https://en.wikipedia.org/wiki/Eigendecomposition_of_a_matrix) and is used to factorize a matrix as $W = U\Sigma V^\top$, where $U, V$ are orthogonal matrices and $\Sigma$ is a matrix of singular values on its diagonal. Note that $\Sigma$ doesn't have to be square.
#
# \begin{align*}
# \Sigma = \begin{bmatrix}\sigma_1 & & \\ & \sigma_2 \\ & & \ddots \\ & & & \sigma_n\end{bmatrix}
# \end{align*}
#
# where $\sigma_1$ and $\sigma_n$ are the largest and smallest singular values, respectively. Intuitively, larger values correspond to larger amounts of stretching a matrix can apply to another vector. Following this notation, $\sigma(W) = \sigma_1$.
#
# **Applying SVD to Spectral Normalization**
#
# To spectrally normalize the weight matrix, you divide every value in the matrix by its spectral norm. As a result, a spectrally normalized matrix $\overline{W}_{SN}$ can be expressed as
#
# \begin{align*}
# \overline{W}_{SN} = \dfrac{W}{\sigma(W)},
# \end{align*}
#
# In practice, computing the SVD of $W$ is expensive, so the authors of the SN-GAN paper do something very neat. They instead approximate the left and right singular vectors, $\tilde{u}$ and $\tilde{v}$ respectively, through power iteration such that $\sigma(W) \approx \tilde{u}^\top W\tilde{v}$.
#
# Starting from randomly initialization, $\tilde{u}$ and $\tilde{v}$ are updated according to
#
# \begin{align*}
# \tilde{u} &:= \dfrac{W^\top\tilde{u}}{||W^\top\tilde{u}||_2} \\
# \tilde{v} &:= \dfrac{W\tilde{v}}{||W\tilde{v}||_2}
# \end{align*}
#
# In practice, one round of iteration is sufficient to "achieve satisfactory performance" as per the authors.
#
# Don't worry if you don't completely follow this! The algorithm is conveniently implemented as `torch.nn.utils.spectral_norm` in PyTorch, so as long as you get the general gist of how it might be useful and when to use it, then you're all set.
#
# + [markdown] id="yCmphbpAU0Uq"
# ## A Bit of History on Spectral Normalization
#
# This isn't the first time that spectral norm has been proposed in the context of deep learning models. There's a paper called [Spectral Norm Regularization for Improving the Generalizability of Deep Learning](https://arxiv.org/abs/1705.10941) (Yoshida et al. 2017) that proposes **spectral norm regularization**, which they showed to improve the generalizability of models by adding extra loss terms onto the loss function (just as L2 regularization and gradient penalty do!). These extra loss terms specifically penalize the spectral norm of the weights. You can think of this as *data-independent* regularization because the gradient with respect to $W$ isn't a function of the minibatch.
#
# **Spectral normalization**, on the other hand, sets the spectral norm of the weight matrices to 1 -- it's a much harder constraint than adding a loss term, which is a form of "soft" regularization. As the authors show in the paper, you can think of spectral normalization as *data-dependent* regularization, since the gradient with respect to $W$ is dependent on the mini-batch statistics (shown in Section 2.1 of the [main paper](https://arxiv.org/pdf/1802.05957.pdf)). Spectral normalization essentially prevents the transformation of each layer
# from becoming to sensitive in one direction and mitigates exploding gradients.
# + [markdown] id="4ig1Vfl5dUR8"
# ## DCGAN with Spectral Normalization
#
# In rest of this notebook, you will walk through how to apply spectral normalization to DCGAN as an example, using your earlier DCGAN implementation. You can always add spectral normalization to your other models too.
#
# Here, you start with the same setup and helper function, as you've seen before.
# + id="m5lJAm2DaY_e"
# Some setup
import torch
from torch import nn
from tqdm.auto import tqdm
from torchvision import transforms
from torchvision.datasets import MNIST
from torchvision.utils import make_grid
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
torch.manual_seed(0) # Set for our testing purposes, please do not change!
'''
Function for visualizing images: Given a tensor of images, number of images, and
size per image, plots and prints the images in an uniform grid.
'''
def show_tensor_images(image_tensor, num_images=25, size=(1, 28, 28)):
image_tensor = (image_tensor + 1) / 2
image_unflat = image_tensor.detach().cpu()
image_grid = make_grid(image_unflat[:num_images], nrow=5)
plt.imshow(image_grid.permute(1, 2, 0).squeeze())
plt.show()
# + [markdown] id="80LI0O_BeH99"
# ### DCGAN Generator
#
# Since spectral normalization is only applied to the matrices in the discriminator, the generator implementation is the same as the original.
# + id="CIRaryyqeLnv"
class Generator(nn.Module):
'''
Generator Class
Values:
z_dim: the dimension of the noise vector, a scalar
im_chan: the number of channels of the output image, a scalar
MNIST is black-and-white, so that's our default
hidden_dim: the inner dimension, a scalar
'''
def __init__(self, z_dim=10, im_chan=1, hidden_dim=64):
super(Generator, self).__init__()
self.z_dim = z_dim
# Build the neural network
self.gen = nn.Sequential(
self.make_gen_block(z_dim, hidden_dim * 4),
self.make_gen_block(hidden_dim * 4, hidden_dim * 2, kernel_size=4, stride=1),
self.make_gen_block(hidden_dim * 2, hidden_dim),
self.make_gen_block(hidden_dim, im_chan, kernel_size=4, final_layer=True),
)
def make_gen_block(self, input_channels, output_channels, kernel_size=3, stride=2, final_layer=False):
'''
Function to return a sequence of operations corresponding to a generator block of the DCGAN,
corresponding to a transposed convolution, a batchnorm (except for in the last layer), and an activation
Parameters:
input_channels: how many channels the input feature representation has
output_channels: how many channels the output feature representation should have
kernel_size: the size of each convolutional filter, equivalent to (kernel_size, kernel_size)
stride: the stride of the convolution
final_layer: whether we're on the final layer (affects activation and batchnorm)
'''
# Build the neural block
if not final_layer:
return nn.Sequential(
nn.ConvTranspose2d(input_channels, output_channels, kernel_size, stride),
nn.BatchNorm2d(output_channels),
nn.ReLU(),
)
else: # Final Layer
return nn.Sequential(
nn.ConvTranspose2d(input_channels, output_channels, kernel_size, stride),
nn.Tanh(),
)
def unsqueeze_noise(self, noise):
'''
Function for completing a forward pass of the Generator: Given a noise vector,
returns a copy of that noise with width and height = 1 and channels = z_dim.
Parameters:
noise: a noise tensor with dimensions (batch_size, z_dim)
'''
return noise.view(len(noise), self.z_dim, 1, 1)
def forward(self, noise):
'''
Function for completing a forward pass of the Generator: Given a noise vector,
returns a generated image.
Parameters:
noise: a noise tensor with dimensions (batch_size, z_dim)
'''
x = self.unsqueeze_noise(noise)
return self.gen(x)
def get_noise(n_samples, z_dim, device='cpu'):
'''
Function for creating a noise vector: Given the dimensions (n_samples, z_dim)
creates a tensor of that shape filled with random numbers from the normal distribution.
Parameters:
n_samples: the number of samples in the batch, a scalar
z_dim: the dimension of the noise vector, a scalar
device: the device type
'''
return torch.randn(n_samples, z_dim, device=device)
# + [markdown] id="DPqEoOBRemUn"
# ### DCGAN Discriminator
#
# For the discriminator, you can wrap each `nn.Conv2d` with `nn.utils.spectral_norm`. In the backend, this introduces parameters for $\tilde{u}$ and $\tilde{v}$ in addition to $W$ so that the $W_{SN}$ can be computed as $\tilde{u}^\top W\tilde{v}$ in runtime.
#
# Pytorch also provides a `nn.utils.remove_spectral_norm` function, which collapses the 3 separate parameters into a single explicit $\overline{W}_{SN} := \tilde{u}^\top W\tilde{v}$. You should only apply this to your convolutional layers during inference to improve runtime speed.
#
# It is important note that spectral norm does not eliminate the need for batch norm. Spectral norm affects the weights of each layer, while batch norm affects the activations of each layer. You can see both in a discriminator architecture, but you can also see just one of them. Hope this is something you have fun experimenting with!
# + id="WJu0YF5TgMho"
class Discriminator(nn.Module):
'''
Discriminator Class
Values:
im_chan: the number of channels of the output image, a scalar
MNIST is black-and-white (1 channel), so that's our default.
hidden_dim: the inner dimension, a scalar
'''
def __init__(self, im_chan=1, hidden_dim=16):
super(Discriminator, self).__init__()
self.disc = nn.Sequential(
self.make_disc_block(im_chan, hidden_dim),
self.make_disc_block(hidden_dim, hidden_dim * 2),
self.make_disc_block(hidden_dim * 2, 1, final_layer=True),
)
def make_disc_block(self, input_channels, output_channels, kernel_size=4, stride=2, final_layer=False):
'''
Function to return a sequence of operations corresponding to a discriminator block of the DCGAN,
corresponding to a convolution, a batchnorm (except for in the last layer), and an activation
Parameters:
input_channels: how many channels the input feature representation has
output_channels: how many channels the output feature representation should have
kernel_size: the size of each convolutional filter, equivalent to (kernel_size, kernel_size)
stride: the stride of the convolution
final_layer: whether we're on the final layer (affects activation and batchnorm)
'''
# Build the neural block
if not final_layer:
return nn.Sequential(
nn.utils.spectral_norm(nn.Conv2d(input_channels, output_channels, kernel_size, stride)),
nn.BatchNorm2d(output_channels),
nn.LeakyReLU(0.2),
)
else: # Final Layer
return nn.Sequential(
nn.utils.spectral_norm(nn.Conv2d(input_channels, output_channels, kernel_size, stride)),
)
def forward(self, image):
'''
Function for completing a forward pass of the Discriminator: Given an image tensor,
returns a 1-dimension tensor representing fake/real.
Parameters:
image: a flattened image tensor with dimension (im_dim)
'''
disc_pred = self.disc(image)
return disc_pred.view(len(disc_pred), -1)
# + [markdown] id="tFMgTfTSgosg"
# ### Training SN-DCGAN
#
# You can now put everything together and train a spectrally normalized DCGAN! Here are all your parameters for initialization and optimization.
# + id="LDn5w93Ej33c"
criterion = nn.BCEWithLogitsLoss()
n_epochs = 50
z_dim = 64
display_step = 500
batch_size = 128
# A learning rate of 0.0002 works well on DCGAN
lr = 0.0002
# These parameters control the optimizer's momentum, which you can read more about here:
# https://distill.pub/2017/momentum/ but you don’t need to worry about it for this course
beta_1 = 0.5
beta_2 = 0.999
device = 'cuda'
# We tranform our image values to be between -1 and 1 (the range of the tanh activation)
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
dataloader = DataLoader(
MNIST(".", download=True, transform=transform),
batch_size=batch_size,
shuffle=True)
# + [markdown] id="bNk2_TZ0j-2n"
# Now, initialize the generator, the discriminator, and the optimizers.
# + id="PhSNFuTfkFnw"
gen = Generator(z_dim).to(device)
gen_opt = torch.optim.Adam(gen.parameters(), lr=lr, betas=(beta_1, beta_2))
disc = Discriminator().to(device)
disc_opt = torch.optim.Adam(disc.parameters(), lr=lr, betas=(beta_1, beta_2))
# We initialize the weights to the normal distribution
# with mean 0 and standard deviation 0.02
def weights_init(m):
if isinstance(m, nn.Conv2d) or isinstance(m, nn.ConvTranspose2d):
torch.nn.init.normal_(m.weight, 0.0, 0.02)
if isinstance(m, nn.BatchNorm2d):
torch.nn.init.normal_(m.weight, 0.0, 0.02)
torch.nn.init.constant_(m.bias, 0)
gen = gen.apply(weights_init)
disc = disc.apply(weights_init)
# + [markdown] id="AVzM2OdjkP29"
# Finally, train the whole thing! And babysit those outputs :)
# -
def get_disc_loss(gen, disc, criterion, real, num_images, z_dim, device):
'''
Return the loss of the discriminator given inputs.
Parameters:
gen: the generator model, which returns an image given z-dimensional noise
disc: the discriminator model, which returns a single-dimensional prediction of real/fake
criterion: the loss function, which should be used to compare
the discriminator's predictions to the ground truth reality of the images
(e.g. fake = 0, real = 1)
real: a batch of real images
num_images: the number of images the generator should produce,
which is also the length of the real images
z_dim: the dimension of the noise vector, a scalar
device: the device type
Returns:
disc_loss: a torch scalar loss value for the current batch
'''
# These are the steps you will need to complete:
# 1) Create noise vectors and generate a batch (num_images) of fake images.
# Make sure to pass the device argument to the noise.
# 2) Get the discriminator's prediction of the fake image
# and calculate the loss. Don't forget to detach the generator!
# (Remember the loss function you set earlier -- criterion. You need a
# 'ground truth' tensor in order to calculate the loss.
# For example, a ground truth tensor for a fake image is all zeros.)
# 3) Get the discriminator's prediction of the real image and calculate the loss.
# 4) Calculate the discriminator's loss by averaging the real and fake loss
# and set it to disc_loss.
# Note: Please do not use concatenation in your solution. The tests are being updated to
# support this, but for now, average the two losses as described in step (4).
# *Important*: You should NOT write your own loss function here - use criterion(pred, true)!
#### START CODE HERE ####
noise = get_noise(num_images, z_dim, device)
image_gen = gen(noise).detach()
fake_loss = criterion(disc(image_gen), torch.zeros(num_images, 1, device=device))
real_loss = criterion(disc(real), torch.ones(num_images, 1, device=device))
disc_loss = (fake_loss + real_loss) / 2
#### END CODE HERE ####
return disc_loss
def get_gen_loss(gen, disc, criterion, num_images, z_dim, device):
'''
Return the loss of the generator given inputs.
Parameters:
gen: the generator model, which returns an image given z-dimensional noise
disc: the discriminator model, which returns a single-dimensional prediction of real/fake
criterion: the loss function, which should be used to compare
the discriminator's predictions to the ground truth reality of the images
(e.g. fake = 0, real = 1)
num_images: the number of images the generator should produce,
which is also the length of the real images
z_dim: the dimension of the noise vector, a scalar
device: the device type
Returns:
gen_loss: a torch scalar loss value for the current batch
'''
# These are the steps you will need to complete:
# 1) Create noise vectors and generate a batch of fake images.
# Remember to pass the device argument to the get_noise function.
# 2) Get the discriminator's prediction of the fake image.
# 3) Calculate the generator's loss. Remember the generator wants
# the discriminator to think that its fake images are real
# *Important*: You should NOT write your own loss function here - use criterion(pred, true)!
#### START CODE HERE ####
noise = get_noise(num_images, z_dim, device)
image_gen = gen(noise)
gen_loss = criterion(disc(image_gen), torch.ones(num_images, 1, device=device))
#### END CODE HERE ####
return gen_loss
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["238087a0e5814cffb2169acb635d5e9e", "3fbe6494be0f494490caadf2199827ea", "<KEY>", "b9c9e9672ddd4bab8dfe405c35f75b15", "<KEY>", "<KEY>", "d604ae4c97a243139340887c863ef308", "<KEY>", "<KEY>", "9992f227848a431b885d2194171d9463", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "e04fda10ade4408d9065546f530a4c5c", "92e1ade8fed4437fb7b3a7f7fbb3a37c", "cec486129ada469c926229245b9bb073", "09bf49e4f6334c58a2f00195872affea", "<KEY>", "<KEY>", "<KEY>", "f884a4669c9b48499be6ba967ed73e44", "<KEY>", "141d1ddc43a54728ab36edac1ea48774", "1ec3c866af53444ea256b749f3a4761b", "18a1c610eeed4f74ac4d129122539cf0", "<KEY>", "<KEY>", "207e47be04904355b3eed7eede1e0d69", "<KEY>", "0e2190b598144d5784aba86e438a4fac", "<KEY>", "<KEY>", "d66981ab056546ea9e08e7221a2685a7", "<KEY>", "<KEY>", "a06283273fed4f7780f55597e5c5d941", "<KEY>", "16b9525432b347ed9bb0407d6a8f322a", "f0d3492c67914550adfbd2a411adfab0", "<KEY>", "947719603245458bb89a08da5ad286c1", "addaeb08c420490d9ea015bda50df402", "93116d6bd5604c32818482efa78d1a6e", "47cbcf61349b481e8ade6b2778315eec", "<KEY>", "af2732ad2bef4e58b1469416cd359d14", "98ce755862dd4fa7b043c25e97b1e9dc", "7d1dc1ddd9b244a19f0b50b5ad9576c8", "ec43115276df4713854e9c5977a5ee79", "ac89168f03f443fcae5fac8ecdb70a29", "<KEY>", "<KEY>", "0c60733eaf00411e896bf75f2ddf784a", "<KEY>", "84b440081a8f475eae4a0d3a9e3674b8", "dd117022733a4c2f8e2bb86e9012a0d3", "<KEY>", "<KEY>", "2e206bd228634745888688535ad7caba", "<KEY>", "<KEY>", "228c14a120064d2c841d84b0e5f7b5af", "<KEY>", "<KEY>", "<KEY>", "031d7f6655934900993e9dd7b93a95f2", "e26fefc113654db3a7556d3745912312", "77a828a1e5d0420cab2ed96feeb0acaf", "e7851c1ae436430695fb3dd6edd9462d", "e340d4556e8841dc893d4d695db9c344", "<KEY>", "<KEY>", "<KEY>", "468b6e0928764f168dd5d143d33ea15d", "<KEY>", "71977107589647a68916052ff7ad73b2", "688a0e91f2354d50aab8967431e4e240", "<KEY>", "53ccd63ceeb04e2eb1ce81f959687004", "efe1a7de3e8b46ef8639b413cb79ac0e", "<KEY>", "699be3961bb644b78d1722d177dc44ec", "33bac321805840519f92672f2d04ce94", "6467ddf71e37425d86af1cc2f0d3061c", "<KEY>", "11e1cd4a278c4dd19d4e98a128ca78cf", "<KEY>", "<KEY>", "<KEY>", "6dc395e2aee640a58a00c6c11a05400d", "<KEY>", "f48f2c336e6e45a5aba5ec4bb1e52f40", "d5db35e2a59f465cb01d595ceae25015", "<KEY>", "653bbe437d6d47f89cfe98303a1bb212", "<KEY>", "ac6dbd1e81cc4b37af3050b3838ea406", "<KEY>", "<KEY>", "00f6de36ceb94d2eaf79b25a98988646", "<KEY>", "b80b982ab0ae436bad7a7f8a58676190", "6824c41e3b294193af3e26ca81cd8714", "412a5d5aeea448ccadd7ee1200106727", "<KEY>", "fec23f62fbef4643ba558bead12a7b88", "<KEY>", "ff42a5ac96a84477877bf611583ba4d8", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "f7627905af4e4e948058c92ec169f373", "6c23ab2d04454dc8a1d3f14ad683d5f0", "16894ec52ede4682a7b4ea1d25627879", "<KEY>", "<KEY>", "8dd3f05340e0492a918a6681495daf5c", "<KEY>", "<KEY>", "9b5551e934af4c718432bedcebe02060", "<KEY>", "1d8b18fd6ea24f37928044ad66299d0e", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "8ca0162f880f4c068ab7fa4e54031eab", "6a76ea00dcbe4e17975d2021ee963b93", "<KEY>", "415859dbdff341a39d50424788f44cc5", "<KEY>", "7af5abed1a5d40ddb32d36f770172d27", "2444be8f6d86495c97a0675bdeb62480", "<KEY>", "7d6050f596834f6182c8102769974025", "<KEY>", "2bc8a51693424a3497df60d99c3779a1", "b83e3e3f4f67475fa24900b6e5d78467", "<KEY>", "<KEY>", "d5e7d0e561b044798af6dc272f63c473", "<KEY>", "<KEY>", "<KEY>", "e991e9ec89e04a4e9bab709740e72684", "76d01c4ea9a14170b45ef8cb95ed9990", "<KEY>", "<KEY>", "<KEY>", "2ef8795705694b9dad1cee771562d54a", "fce36e62a85643e29f8795fe5b46b329", "<KEY>", "9b09438538654515a67a93c9cd64e31d", "<KEY>", "<KEY>", "a5190be18d7f4c808be8105602234228", "<KEY>", "f08a3c55919e4ab8904fafcc5d908b3c", "<KEY>", "<KEY>", "48e51261d810468eb201322e12b7481c", "<KEY>", "<KEY>", "0adfafe8ea7c4248b27bed3462f6f3ad", "<KEY>", "2d26b36d1e634ce9894db9940e55fad6", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "00f5ded52d0541efa2721ad6753222c6", "7eae05ee7f75426a934724d15dc10ef5", "c7a733b3ac624a8b85e25c9907a1df16", "<KEY>", "85531d9a53944adaa80b1fa0cafd2a0f", "2b1ace36028d48d3b599694d9db0469b", "5f3498a8d3774137b28ca2c1d4636303", "ed77d6d50b1747e7a8d9882df81c2fc4", "<KEY>", "b8102e1b16164825ad00597835166965", "ed8ba436d4be44a995e09cac481481c1", "<KEY>", "<KEY>", "1827f6903f92482e98b2ac2dc4aa4124", "eca50345fad048f79eb2bf21050ed736", "74a9644d5d2b4c6eb7f3eed1a23dedaa", "<KEY>", "<KEY>", "8ad7f48572664140b060c36e145f6902", "<KEY>", "<KEY>", "94f916dc9c7d4e2c997c253142aadb02", "8de390c272db43a09c989ca85bc91662", "1c6c7a1c52744c809948217480d0e48e", "db771d2155264020be2ecce293202a24", "25d14c1076584b989bc616a65cee5ff4", "be6d8957a68147e2a501f825754a820b", "424ed16c9f1447d8a6e91837d4249217", "94e7d67985ef474b9f84621d3e7a104d", "<KEY>", "02108a5f0a944f658226106d1f7ad3a6", "be78816556334746ae257e5c51606f5e", "<KEY>", "<KEY>", "fd0d96fbee9d4fd69f41125c3f9d200e", "<KEY>", "00fee5cbe3ab4d0096e87f1e706f3611", "ad51bc4e1d174a2aa0eaa617cced8f35", "d3bbf15ffaac4627b000e50443e7c603", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "43c0ee831c334683a5508124b4a79776", "<KEY>", "<KEY>", "89f86896c43f476e9d78883a1ddc5daf", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "672339438a5043ca9820789ea93df6d0", "9a3a51a2aba5425abe0860fa16c7a373", "2cfc8e940ec9489c88ec98844fac4b7e", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "31255ec3a59540e284d3db79c858de95", "<KEY>", "<KEY>", "ada9407958bb4a2799919349715dd06d", "<KEY>", "8b353878ed644ec5879fb37a89c464f2", "<KEY>", "f07dabb98afa4a3781302c5d007e1f50", "379906bcaa61469cacc29e585449a1af", "5cd9fa9dea284afea657fe9f4ec29baa", "<KEY>", "143198a917c4453c90528e103963be6d", "3e0ac7389f04428b94d0fca63a678cee", "7614db82dcea48658ce78aabca03a5f8", "934a7ed269fd4271bc080ab7dcce3e8c", "cedc73924a1b4ccc948c737dd904e48a", "21f2379348554becbba7755c25142441", "0a8d13e6f7bf48289fd684cc5a44297d", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "4f238aeef1d04d69b04c308035e1ddb4", "<KEY>", "<KEY>", "<KEY>", "62ca5189c8044d58b7dad2e6ad64906e", "e67f7a12282a4145a55271381d7330ba", "aa1d1a449447467bac98d3ea19b135be", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "ac9a92acc3d74e6694eec8e43494796d", "<KEY>", "37d1e9b9faec462eac1e17282454fff3", "385ada0779af43f3b8ec1baaf3098e9e", "<KEY>", "<KEY>", "63f103e7495745ebb03f467bea8316d3", "23ff9d45666f4057bd902af18ae81ca3", "30fa5b07cce94523ad220bcc68ea7bf3", "<KEY>", "f0ba21035f9346f3aed9d86cd0888392", "f54cecdc77d84c0c9c2a1027dd1a4674", "<KEY>", "e66673f4bde34c648f97941e5ab91a8a", "224c6e41d5974fa6bcc193f4c1e569d2", "5b82721583064be5872e08834027d9c3", "<KEY>", "<KEY>", "<KEY>", "46c8eb28e62f440295c37513c56fe723", "<KEY>", "<KEY>", "<KEY>", "12a666b22e4040cab499d93a28c5d820", "<KEY>", "fa61552203c145e28f8ef7b2a38329e3", "<KEY>", "<KEY>", "925e1e2b80be466daf67b7065e628854", "6ced3efe5af14ff09ba55059d9a969c6", "2fa58df3cc8d472985a35c4329171a36", "92e01766583a40debe4bc8f875a052c5", "<KEY>", "3597a0c6c4da42acaa4f2043257fdca4", "1c824641932248f088f64ef88837ac38", "<KEY>", "<KEY>", "<KEY>", "6b58e74cd5b04462a1089b2769167e5c", "fd81f78bd593448a8b0da5ab9efeb027", "fd8ff8853f204c4eb03e842a0368f4d7", "<KEY>", "2c0a730df08845d88d27295d8045306b", "0cf74f8fe48240c19059ca346975bbb2", "<KEY>", "<KEY>", "<KEY>", "c570beab0d244daf8c76c093537e89ec", "<KEY>", "<KEY>", "060b7e8d67324071bdd53c721c5c4dc7", "0d5e19eff67841d8add20f81c812bac2", "<KEY>", "<KEY>", "<KEY>", "f40501ef89d14642aba345ae629dc84a", "<KEY>", "32657ac3046a446eaa367b70d50975ad", "20f7f473658e4daeb3a1e27efedd763c", "a9babc50c1c24fd1a6424dec8a466b53", "<KEY>", "04d63eee83f04437a022552384efe7ae", "e6bd9d58e65343a9978ab2ed02472bec", "682f652c9d9d44539a5538a2cc87c8a8", "1fe2ab9ee4ce49a3bfbc74ff211ced22", "f73ffa901b3540079a512ca0e714bbfa", "3ff4124686724226a2ec3fef05cc14c3", "<KEY>", "e216d762a1d241c18bd6eb729ec6dc43", "5fac53d079124a7099daa6d23d62e9f4", "<KEY>", "<KEY>", "9134ea0f84db43b8bce64a4c51739558", "<KEY>", "<KEY>", "3a8d87084efe4a3d8a6b1ddffa6a6b76", "b17bd3fa5de84bfe8a664c3e04df21e7", "6c8f0064a2d543ca9d32d2a14b04321d", "<KEY>", "55d73e07138d4ca8b2e47e999f5db6ac", "<KEY>", "a5cafbb1ebe542969c09a186182e9e16", "95691287ec644d479c31abd7aba9c996", "<KEY>", "41009632b931487ca0c86a6c08c4dcec", "<KEY>", "<KEY>", "2ff6a15c4d1e41e4a069ff1d838ca965", "<KEY>", "20a80dbc4e8e4194993e25087ae2822b", "4320ed22b17a47c0971a79a3f79c859c", "<KEY>", "<KEY>", "9f62e7d1a81e4a138024626c145805d8", "<KEY>", "105a663c3607478a9c278a8d5d8c28a9", "<KEY>", "<KEY>", "27143846c3a041ddb8ec86cb9009ce17", "e976534a7dcf44d29a7b313760124e10", "f579d01fe220410887462f24600be6b2", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "ee27065011884360a7e6288a418c3b3f", "<KEY>", "<KEY>", "5296a1f496734b6c9432e18d7e5c192c", "c8eb4157396e4fbdb6e10dee56f9c421", "f95498b91e584888b1773c316c359686", "<KEY>", "5164159d8b3b4c39a66352692316ca26", "<KEY>", "dbd0deb4d1d343b2ae662cb6e8147d05", "8c2844defe8f43b5aee26b8c3797e22d", "fe921fc89c7e4985bc30a8f46ec4ced7", "32ed845b84494aa08d1cb38ee5020c5a", "061328e026764148a98199d6b9a67623", "<KEY>", "<KEY>", "<KEY>", "d375bfd19c7e4fc9ab7bf281ee5517ed", "36cf1e01ba2b411399ee05461b4e6770", "02ff312f25ed4ec28590a595e84b0538", "95a81ff9af63457795f03e8bd5b649d0", "ee19cc8a48e843229e4bfaf47f25ca78", "<KEY>", "<KEY>", "<KEY>", "b5f059feb96c4af0b8b187493e7c37e8"]} id="1AZCWsaSkSkd" outputId="0e03c8ad-1edb-44c4-d0c4-ec16d6beab64"
cur_step = 0
disc_losses = []
gen_losses = []
for epoch in range(n_epochs):
# Dataloader returns the batches
for real, _ in tqdm(dataloader):
cur_batch_size = len(real)
real = real.to(device)
## Update discriminator ##
# Zero out the gradients before backpropagation
disc_opt.zero_grad()
# Calculate discriminator loss
disc_loss = get_disc_loss(gen, disc, criterion, real, cur_batch_size, z_dim, device)
# Update gradients
disc_loss.backward(retain_graph=True)
# Update optimizer
disc_opt.step()
## Update generator ##
gen_opt.zero_grad()
gen_loss = get_gen_loss(gen, disc, criterion, cur_batch_size, z_dim, device)
gen_loss.backward()
gen_opt.step()
# Keep track of the average discriminator loss
disc_losses += [disc_loss.item()]
# Keep track of the average generator loss
gen_losses += [gen_loss.item()]
## Visualization code ##
if cur_step % display_step == 0 and cur_step > 0:
mean_disc_loss = sum(disc_losses[-display_step:]) / display_step
mean_gen_loss = sum(gen_losses[-display_step:]) / display_step
print(f"Step {cur_step}: discriminator loss: {mean_disc_loss}, Generator loss: {mean_gen_loss}")
fake_noise = get_noise(cur_batch_size, z_dim, device=device)
fake = gen(fake_noise)
show_tensor_images(fake)
show_tensor_images(real)
step_bins = 20
num_examples = (len(gen_losses) // step_bins) * step_bins
plt.plot(
range(num_examples // step_bins),
torch.Tensor(disc_losses[:num_examples]).view(-1, step_bins).mean(1),
label="Disc Loss"
)
plt.plot(
range(num_examples // step_bins),
torch.Tensor(gen_losses[:num_examples]).view(-1, step_bins).mean(1),
label="Gen Loss"
)
plt.legend()
plt.show()
cur_step += 1
| 06-SNGAN-MNIST-PyTorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The aim of this project is to use a simple dataset that reflects the type of population health analytics that we will want to do in Discover-NOW.
#
# The project will seek to answer three questions:
# 1. Does age play a role in stroke incidence?
# 2. Is hypertension more likely to lead to strokes?
# 3. Can we predict who is likely to have a stroke based on their characteristics in the dataset?
# + pycharm={"name": "#%%\n"}
#Import all the required packages
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split, cross_validate
from sklearn.metrics import r2_score, mean_squared_error, accuracy_score, plot_confusion_matrix
import sklearn.metrics as metrics
from pywaffle import Waffle
from sklearn.svm import SVC
import missingno as msno
import scipy.stats as stats
import seaborn as sns
# %matplotlib inline
## read in data
df = pd.read_csv('C:/Users/mchisambi/GoogleDrive/udadatasci/stroke-data.csv')
# + [markdown] pycharm={"name": "#%% md\n"}
# Data wrangling steps:
# 1. Review missing data. BMI is the only category that has missing data.
# + pycharm={"name": "#%%\n"}
#df.info() #great for missing data
color = ['#3891a6','#3891a6','#3891a6','#3891a6','#3891a6','#3891a6','#3891a6','#3891a6','#3891a6','#3891a6','#3891a6','#29335c']
fig, ax = plt.subplots(figsize = (12,4), dpi = 70)
fig.patch.set_facecolor('#F8F8F8')
ax.set_facecolor('#F8F8F8')
msno.bar(df, sort = 'descending',
color = color,
ax = ax, fontsize =8,
labels = 'off',filter = 'top')
ax.text(-1,1.35,'Visualization of null values in the data set',{'font': 'Helvetica', 'Size': 24, 'color':'black'},alpha = 0.9)
ax.text(-1,1.2,'Overall there are 5110 datapoints are present in the dataset. Only "bmi" feature has null values.',{'font': 'Helvetica', 'Size': 12, 'color':'black'}, alpha = 0.7)
ax.text(-1,1.15, 'Source: https://www.kaggle.com/bhuvanchennoju/data-stroytelling-auc-focus-on-strokes',{'font': 'Helvetica', 'Size': 10, 'color':'black'}, alpha = 0.7)
ax.set_xticklabels(ax.get_xticklabels(),rotation = 90,
ha = 'center', **{'font': 'Helvetica', 'Size': 10,'weight':'normal','color':'#512b58'}, alpha = 1)
ax.set_yticklabels('')
ax.spines['bottom'].set_visible(True)
# fig.show()
# + [markdown] pycharm={"name": "#%% md\n"}
# 2. An alternative visualisation is using the matrix function
# + pycharm={"name": "#%%\n"}
fig, ax = plt.subplots(figsize = (12,4), dpi = 70)
fig.patch.set_facecolor('#f6f5f5')
ax.set_facecolor('#f6f5f5')
msno.matrix(df, sort='ascending',
sparkline=False, ax = ax,
fontsize=8, labels='off',
filter="top", color=(56/255,145/255,166/255))
ax.set_title('Use of missingno matrix',{'font': 'Serif', 'Size': 24, 'color':'black'},alpha = 0.75)
ax.spines['bottom'].set_visible(True)
# fig.show()
# + [markdown] pycharm={"name": "#%% md\n"}
# 3. Replace the missing BMI values with the mean
# + pycharm={"name": "#%%\n"}
#subset
df['bmi2'] = df['bmi'].fillna(df['bmi'].mean())
df = df.drop(['bmi'], axis=1)
# msno.matrix(df, sort='ascending',
# sparkline=False, ax = ax,
# fontsize=8, labels='off',
# filter="top", color=(0.1,0.1,0.5))
#
# fig.show()
fig, ax = plt.subplots(figsize = (12,4), dpi = 70)
fig.patch.set_facecolor('#f6f5f5')
ax.set_facecolor('#f6f5f5')
msno.matrix(df, sort='ascending',
sparkline=False, ax = ax,
fontsize=8, labels='off',
filter="top", color=(56/255,145/255,166/255))
ax.set_title('Use of missingno matrix',{'font': 'Serif', 'Size': 24, 'color':'black'},alpha = 0.75)
ax.spines['bottom'].set_visible(True)
# fig.show()
# -
# 3. Review data, initially continuous variables
#
# + pycharm={"name": "#%%\n"}
df.hist()
round (df.describe(exclude = 'object'), 2)
# -
# 4. Then, categorical variables
# + pycharm={"name": "#%%\n"}
round (df.describe(exclude = ['float', 'int64']),2)
# + [markdown] pycharm={"name": "#%% md\n"}
# 5. Create bins for some of the continuous values
# + pycharm={"name": "#%%\n"}
df['bmi_cat'] = pd.cut(df['bmi2'], bins = [0, 19, 25,30,10000], labels = ['Underweight', 'Ideal', 'Overweight', 'Obesity'])
df['age_cat'] = pd.cut(df['age'], bins = [0,20,40,60,80,200], labels = ['0-20','21-40','41-60','61-80','80+' ])
df['glucose_cat'] = pd.cut(df['avg_glucose_level'], bins = [0,90,160,230,500], labels = ['Low', 'Normal', 'High', 'Very High'])
print(df)
# + [markdown] pycharm={"name": "#%% md\n"}
# 6. Establish colour palette for consistency.
# + pycharm={"name": "#%%\n"}
colors = ['#3891a6','#29335c', '#E4572E', '#def2c8', '#9f87af', '#918450']
palette = sns.color_palette(palette = colors)
customPalette = sns.set_palette(sns.color_palette(colors))
fig = plt.figure(figsize = (12,6), dpi = 60)
sns.palplot(palette, size =2.5)
plt.text(-0.75,-0.75,'Color Palette for this Visualization', {'font':'serif', 'size':25, 'weight':'bold'})
plt.text(-0.75,-0.64,'The same colors will be used for throughout this notebook.', {'font':'serif', 'size':18, 'weight':'normal'}, alpha = 0.8)
plt.show()
# -
# ****DATA WRANGLING COMPLETE****
# ~~~~~~~
# Now to address the three questions:
# 1. Does age play a role in stroke incidence?
# 2. Is hypertension more likely to lead to strokes?
# 3. Can we predict who is likely to have a stroke based on their characteristics in the dataset?
#
# ~~~~~~~
# **QUESTION 1: Does age play a role in stroke incidence?**
# 1a) Review the overall coelations in the dataset.
# + pycharm={"name": "#%%\n"}
fig = plt.figure(figsize = (8,6), dpi = 60)
fig.patch.set_facecolor('#f6f5f5')
sns.heatmap(df.corr(), annot=True,fmt='.2f',robust=True, cmap=colors, linecolor='black')
fig.text(0.25,1,'Age is positively associated with several conditions',{'font':'Serif', 'weight':'bold','ha':'center', 'color': 'black', 'size':20})
# + [markdown] pycharm={"name": "#%% md\n"}
# 1b) Age is clearly positively associated with several other continuous variables (bmi, stroke, avg_glucose_level, heart_disease, hypertension)
# 1c) The positive association with stroke can be visualised as follows:
# + pycharm={"name": "#%%\n"}
#plot
fig = plt.figure(figsize = (12,6), dpi = 60)
gs = fig.add_gridspec(10,24)
gs.update(wspace = 1, hspace = 0.05)
fig.patch.set_facecolor('#f6f5f5')
sns.kdeplot(data = df[df['stroke'] == 0], x = 'age', shade = True, alpha = 1, color = colors[0] )
sns.kdeplot(data = df[df['stroke'] == 1], x = 'age', shade = True, alpha = 0.8, color = colors[1])
ax.set_xlabel('Age of a person', fontdict = {'font':'Serif', 'color': 'black', 'weight':'bold','size': 16})
ax.text(-17,0.0525,'Age-Stroke Distribution - How serious is it?', {'font':'Serif', 'weight':'bold','color': 'black', 'size':24}, alpha= 0.9)
ax.text(-17,0.043,'From stoke Distribution it is clear that aged people are \nhaving significant number of strokes.', {'font':'Serif', 'color': 'black', 'size':14})
ax.text(100,0.043, 'Stroke ', {'font': 'Serif','weight':'bold','Size': '16','weight':'bold','style':'normal', 'color':'#fe346e'})
ax.text(117,0.043, '|', {'color':'black' , 'size':'16', 'weight': 'bold'})
ax.text(120,0.043, 'Healthy', {'font': 'Serif','weight':'bold', 'Size': '16','style':'normal', 'weight':'bold','color':'#512b58'})
fig.text(0.25,1,'The number of strokes increases with age',{'font':'Serif', 'weight':'bold','color': 'black', 'ha': 'center', 'size':20})
# + pycharm={"name": "#%%\n"}
plt.figure(figsize = (12,6), dpi = 60)
sns.displot(df, x='age_cat', hue='stroke', palette=customPalette)
# -
# **QUESTION 2: Is hypertension more likely to lead to strokes?**
#
# 2a) A simple plot of the poportion of patients who have stroke based on their hypertension status.
# + pycharm={"name": "#%%\n"}
sns.factorplot(data = df, x = 'hypertension', y='stroke', kind='bar')
# sns.factorplot(data = df, x = 'heart_disease', y='stroke', kind='bar')
# + [markdown] pycharm={"name": "#%% md\n"}
# 2b) Odds Ratio estimate of stroke if a patient has hypertension is positive and statistically significant.
# + pycharm={"name": "#%%\n"}
table = pd.crosstab(df['hypertension'], df['stroke'])
oddsratio, oddsratio_confint = stats.fisher_exact(table)
print(table)
print("OddsR: ", oddsratio, "p-Value:", oddsratio_confint)
# + [markdown] pycharm={"name": "#%% md\n"}
# **QUESTION 3: Can I predict whether someone will have a stroke or not using certain variables?**
#
# For this question, I will be using support vector classification, as we are trying to predict a categorical variable.
#
# 3a) Create new dataset with categorical data dummies
# + pycharm={"name": "#%%\n"}
#dummy_df = pd.get_dummies(df)
df_drop = df.drop(['id', 'bmi_cat', 'age_cat', 'glucose_cat'], axis=1)
df_pred = pd.get_dummies(df_drop)
#create a subset
print(df.columns)
print(df_drop.columns)
print(df_pred.columns)
# + [markdown] pycharm={"name": "#%% md\n"}
# 3b) Create training and testing dataset
# + pycharm={"name": "#%%\n"}
print(df_pred.columns)
X = df_pred.drop(['stroke'], axis=1)
y = df_pred['stroke']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.25, random_state=42)
# + pycharm={"name": "#%%\n"}
SVM_Model = SVC(gamma='auto')
SVM_Model.fit(X_train,y_train)
predicted = SVM_Model.predict(X_test)
# + [markdown] pycharm={"name": "#%% md\n"}
# 3d) Evaluate the model
# + pycharm={"name": "#%%\n"}
metrics.accuracy_score(y_test, predicted)
metrics.confusion_matrix(y_test, predicted)
plot_confusion_matrix(SVM_Model, X_test, y_test, cmap='Blues', normalize='pred')
| strokePrediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ML Basics
# **What is linear regression?**
#
# In simple terms, linear regression is a method of finding the best straight line fitting to the given data, i.e. finding the best linear relationship between the independent and dependent variables.
# In technical terms, linear regression is a machine learning algorithm that finds the best linear-fit relationship on any given data, between independent and dependent variables. It is mostly done by the Sum of Squared Residuals Method.
# **State the assumptions in a linear regression model.**
# 1. The assumption about the form of the model:
# - It is assumed that there is a linear relationship between the dependent and independent variables. It is known as the ‘linearity assumption’.
#
# 2. Assumptions about the residuals:
# - Normality assumption:
# - It is assumed that the error terms, ε(i), are normally distributed.
# - Zero mean assumption:
# - It is assumed that the residuals have a mean value of zero.
# - Constant variance assumption:
# - It is assumed that the residual terms have the same (but unknown) variance, $\sigma^2$
# - This assumption is also known as the assumption of homogeneity or homoscedasticity.
# - Independent error assumption:
# - It is assumed that the residual terms are independent of each other, i.e. their pair-wise covariance is zero.
# - If the residuals are not normally distributed, their randomness is lost, which implies that the model is not able to explain the relation in the data.
# - If the expectation(mean) of residuals, E(ε(i)), is zero, the expectations of the target variable and the model become the same, which is one of the targets of the model.
# The residuals (also known as error terms) should be independent. This means that there is no correlation between the residuals and the predicted values, or among the residuals themselves. If some correlation is present, it implies that there is some relation that the regression model is not able to identify.
#
# 3. Assumptions about the estimators:
# - The independent variables are measured without error.
# - The independent variables are linearly independent of each other, i.e. there is no multicollinearity in the data.
# - If the independent variables are not linearly independent of each other, the uniqueness of the least squares solution (or normal equation solution) is lost.
#
# **What is the use of regularisation? Explain L1 and L2 regularisations.**
#
# Regularisation is a technique that is used to tackle the problem of overfitting of the model. When a very complex model is implemented on the training data, it overfits. At times, the simple model might not be able to generalise the data and the complex model overfits. To address this problem, regularisation is used.
# Regularisation is nothing but adding the coefficient terms (betas) to the cost function so that the terms are penalised and are small in magnitude. This essentially helps in capturing the trends in the data and at the same time prevents overfitting by not letting the model become too complex.
#
# - L1 or LASSO regularisation: Here, the absolute values of the coefficients are added to the cost function. This can be seen in the following equation; the highlighted part corresponds to the L1 or LASSO regularisation. This regularisation technique gives sparse results, which lead to feature selection as well.
# 
# - L2 or Ridge regularisation: Here, the squares of the coefficients are added to the cost function. This can be seen in the following equation, where the highlighted part corresponds to the L2 or Ridge regularisation.
#
# 
#
# Inroducing a penalty to the sum of the weights means that the model has to “distribute” its weights optimally, so naturally most of this “resource” will go to the simple features that explain most of the variance, with complex features getting small or zero weights.
#
# https://towardsdatascience.com/intuitions-on-l1-and-l2-regularisation-235f2db4c261
# **How to choose the value of the regularisation parameter (λ)?**
#
# Selecting the regularisation parameter is a tricky business. If the value of λ is too high, it will lead to extremely small values of the regression coefficient β, which will lead to the model underfitting (high bias – low variance). On the other hand, if the value of λ is 0 (very small), the model will tend to overfit the training data (low bias – high variance).
# There is no proper way to select the value of λ. What you can do is have a sub-sample of data and run the algorithm multiple times on different sets. Here, the person has to decide how much variance can be tolerated. Once the user is satisfied with the variance, that value of λ can be chosen for the full dataset.
# One thing to be noted is that the value of λ selected here was optimal for that subset, not for the entire training data.
| notebooks/MLBasics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Survey EDA based on care giver
#
# 
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt
import seaborn as sns
from collections import Counter
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# +
df = pd.read_csv('./Datasets/Input1/survey.csv')
# -
df.head()
# ## Question 1# What is your initial conclusion looking at the table above?
df.shape #Rows and columns
df.info()
df.groupby("Age").size()
# # Question2 : What does negative age signify?
# # Ravi codes for data cleansing as age is below 0
df['Age'] = pd.to_numeric(df['Age'],errors='coerce') #If ‘coerce’, then invalid parsing will be set as NaN
def age_process(age):
if age>=0 and age<=100:
return age
else:
return np.nan
df['Age'] = df['Age'].apply(age_process)
fig,ax = plt.subplots(figsize=(8,6))
sns.distplot(df['Age'].dropna(),ax=ax,kde=False,color='#ffa726')
plt.title('Age Distribution')
plt.ylabel('Freq')
# # Question 3# What do you interpret from the histogram above?
# most_common Return a list of the n most common elements and their counts from the most common to the least.
#If n is omitted or None, most_common() returns all elements in the counter.
#Elements with equal counts are ordered arbitrarily:
country_count = Counter(df['Country'].dropna().tolist()).most_common(10)
country_count
country_count = Counter(df['Country'].dropna().tolist()).most_common(10)
country_idx = [country[0] for country in country_count] # Define X axis
country_val = [country[1] for country in country_count] # Define Y axis
fig,ax = plt.subplots(figsize=(8,6))
sns.barplot(x = country_idx,y=country_val ,ax =ax)
plt.title('Top ten country')
plt.xlabel('Country')
plt.ylabel('Count')
ticks = plt.setp(ax.get_xticklabels(),rotation=90)
df['Timestamp'] = pd.to_datetime(df['Timestamp'],format='%Y-%m-%d')
df['Year'] = df['Timestamp'].apply(lambda x:x.year)
sns.countplot(df['treatment'])
plt.title('Treatement Distribution')
df['Age_Group'] = pd.cut(df['Age'].dropna(),
[0,18,25,35,45,99],
labels=['<18','18-24','25-34','35-44','45+'])
fig,ax = plt.subplots(figsize=(8,6))
sns.countplot(data=df,x = 'Age_Group',hue= 'family_history',ax=ax)
plt.title('Age vs family_history')
fig,ax =plt.subplots(figsize=(8,6))
sns.countplot(data = df,x = 'Age_Group', hue='treatment')
plt.title('Age Group vs Treatment')
fig,ax =plt.subplots(figsize=(8,6))
sns.countplot(df['work_interfere'].dropna(),ax=ax)
plt.title('Work interfere Distribution')
plt.ylabel('Count')
fig,ax = plt.subplots(figsize=(8,6))
total = df['no_employees'].dropna().shape[0] * 9.0
employee_count = Counter(df['no_employees'].dropna().tolist())
for key,val in employee_count.items():
employee_count[key] = employee_count[key] / total
employee_group = np.asarray(list(employee_count.keys()))
employee_val = np.asarray(list(employee_count.values()))
sns.barplot(x = employee_group , y = employee_val)
plt.title('employee group ratio')
plt.ylabel('ratio')
plt.xlabel('employee group')
fig,ax = plt.subplots(figsize=(8,6))
sns.countplot(data = df,x = 'no_employees', hue ='tech_company',ax=ax )
ticks = plt.setp(ax.get_xticklabels(),rotation=45)
plt.title('no_employee vs tech_company')
fig,ax = plt.subplots(figsize=(8,6))
sns.countplot(data = df,x = 'no_employees', hue ='remote_work',ax=ax )
ticks = plt.setp(ax.get_xticklabels(),rotation=45)
plt.title('no_employee vs remote_work')
| 2 Survey EDA based on caregiver centered experience domain V1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Partial Least Squares Regression (PLSR) on Near Infrared Spectroscopy (NIR) data and octane data
# This notebook illustrates how to use the **hoggorm** package to carry out partial least squares regression (PLSR) on multivariate data. Furthermore, we will learn how to visualise the results of the PLSR using the **hoggormPlot** package.
# ---
# ### Import packages and prepare data
# First import **hoggorm** for analysis of the data and **hoggormPlot** for plotting of the analysis results. We'll also import **pandas** such that we can read the data into a data frame. **numpy** is needed for checking dimensions of the data.
import hoggorm as ho
import hoggormplot as hop
import pandas as pd
import numpy as np
# Next, load the data that we are going to analyse using **hoggorm**. After the data has been loaded into the pandas data frame, we'll display it in the notebook.
# Load fluorescence data
X_df = pd.read_csv('gasoline_NIR.txt', header=None, sep='\s+')
X_df
# Load response data, that is octane measurements
y_df = pd.read_csv('gasoline_octane.txt', header=None, sep='\s+')
y_df
# The ``nipalsPLS1`` class in hoggorm accepts only **numpy** arrays with numerical values and not pandas data frames. Therefore, the pandas data frames holding the imported data need to be "taken apart" into three parts:
# * two numpy array holding the numeric values
# * two Python list holding variable (column) names
# * two Python list holding object (row) names.
#
# The numpy arrays with values will be used as input for the ``nipalsPLS2`` class for analysis. The Python lists holding the variable and row names will be used later in the plotting function from the **hoggormPlot** package when visualising the results of the analysis. Below is the code needed to access both data, variable names and object names.
# +
# Get the values from the data frame
X = X_df.values
y = y_df.values
# Get the variable or columns names
X_varNames = list(X_df.columns)
y_varNames = list(y_df.columns)
# Get the object or row names
X_objNames = list(X_df.index)
y_objNames = list(y_df.index)
# -
# ---
# ### Apply PLSR to our data
# Now, let's run PLSR on the data using the ``nipalsPLS1`` class, since we have a univariate response. The documentation provides a [description of the input parameters](https://hoggorm.readthedocs.io/en/latest/plsr.html). Using input paramter ``arrX`` and ``vecy`` we define which numpy array we would like to analyse. ``vecy`` is what typically is considered to be the response vector, while the measurements are typically defined as ``arrX``. By setting input parameter ``Xstand=False`` we make sure that the variables are only mean centered, not scaled to unit variance, if this is what you want. This is the default setting and actually doesn't need to expressed explicitly. Setting paramter ``cvType=["loo"]`` we make sure that we compute the PLS2 model using full cross validation. ``"loo"`` means "Leave One Out". By setting paramter ``numpComp=10`` we ask for four components to be computed.
model = ho.nipalsPLS1(arrX=X, Xstand=False,
vecy=y,
cvType=["loo"],
numComp=10)
# That's it, the PLS2 model has been computed. Now we would like to inspect the results by visualising them. We can do this using plotting functions of the separate [**hoggormPlot** package](https://hoggormplot.readthedocs.io/en/latest/). If we wish to plot the results for component 1 and component 2, we can do this by setting the input argument ``comp=[1, 2]``. The input argument ``plots=[1, 6]`` lets the user define which plots are to be plotted. If this list for example contains value ``1``, the function will generate the scores plot for the model. If the list contains value ``6`` the explained variance plot for y will be plotted. The hoggormPlot documentation provides a [description of input paramters](https://hoggormplot.readthedocs.io/en/latest/mainPlot.html).
hop.plot(model, comp=[1, 2],
plots=[1, 6],
objNames=X_objNames,
XvarNames=X_varNames,
YvarNames=y_varNames)
# Plots can also be called separately.
# Plot cumulative explained variance (both calibrated and validated) using a specific function for that.
hop.explainedVariance(model)
# Plot cumulative validated explained variance in X.
hop.explainedVariance(model, which=['X'])
hop.scores(model)
# Plot X loadings in line plot
hop.loadings(model, weights=True, line=True)
# Plot regression coefficients
hop.coefficients(model, comp=[3])
# ---
# ### Accessing numerical results
# Now that we have visualised the PLSR results, we may also want to access the numerical results. Below are some examples. For a complete list of accessible results, please see this part of the documentation.
# +
# Get X scores and store in numpy array
X_scores = model.X_scores()
# Get scores and store in pandas dataframe with row and column names
X_scores_df = pd.DataFrame(model.X_scores())
X_scores_df.index = X_objNames
X_scores_df.columns = ['Comp {0}'.format(x+1) for x in range(model.X_scores().shape[1])]
X_scores_df
# -
help(ho.nipalsPLS1.X_scores)
# Dimension of the X_scores
np.shape(model.X_scores())
# We see that the numpy array holds the scores for all countries and OECD (35 in total) for four components as required when computing the PCA model.
# +
# Get X loadings and store in numpy array
X_loadings = model.X_loadings()
# Get X loadings and store in pandas dataframe with row and column names
X_loadings_df = pd.DataFrame(model.X_loadings())
X_loadings_df.index = X_varNames
X_loadings_df.columns = ['Comp {0}'.format(x+1) for x in range(model.X_loadings().shape[1])]
X_loadings_df
# -
help(ho.nipalsPLS1.X_loadings)
np.shape(model.X_loadings())
# Here we see that the array holds the loadings for the 10 variables in the data across four components.
# +
# Get Y loadings and store in numpy array
Y_loadings = model.Y_loadings()
# Get Y loadings and store in pandas dataframe with row and column names
Y_loadings_df = pd.DataFrame(model.Y_loadings())
Y_loadings_df.index = y_varNames
Y_loadings_df.columns = ['Comp {0}'.format(x+1) for x in range(model.Y_loadings().shape[1])]
Y_loadings_df
# +
# Get X correlation loadings and store in numpy array
X_corrloadings = model.X_corrLoadings()
# Get X correlation loadings and store in pandas dataframe with row and column names
X_corrloadings_df = pd.DataFrame(model.X_corrLoadings())
X_corrloadings_df.index = X_varNames
X_corrloadings_df.columns = ['Comp {0}'.format(x+1) for x in range(model.X_corrLoadings().shape[1])]
X_corrloadings_df
# -
help(ho.nipalsPLS1.X_corrLoadings)
# +
# Get Y loadings and store in numpy array
Y_corrloadings = model.X_corrLoadings()
# Get Y loadings and store in pandas dataframe with row and column names
Y_corrloadings_df = pd.DataFrame(model.Y_corrLoadings())
Y_corrloadings_df.index = y_varNames
Y_corrloadings_df.columns = ['Comp {0}'.format(x+1) for x in range(model.Y_corrLoadings().shape[1])]
Y_corrloadings_df
# -
help(ho.nipalsPLS1.Y_corrLoadings)
# +
# Get calibrated explained variance of each component in X
X_calExplVar = model.X_calExplVar()
# Get calibrated explained variance in X and store in pandas dataframe with row and column names
X_calExplVar_df = pd.DataFrame(model.X_calExplVar())
X_calExplVar_df.columns = ['calibrated explained variance in X']
X_calExplVar_df.index = ['Comp {0}'.format(x+1) for x in range(model.X_loadings().shape[1])]
X_calExplVar_df
# -
help(ho.nipalsPLS1.X_calExplVar)
# +
# Get calibrated explained variance of each component in Y
Y_calExplVar = model.Y_calExplVar()
# Get calibrated explained variance in Y and store in pandas dataframe with row and column names
Y_calExplVar_df = pd.DataFrame(model.Y_calExplVar())
Y_calExplVar_df.columns = ['calibrated explained variance in Y']
Y_calExplVar_df.index = ['Comp {0}'.format(x+1) for x in range(model.Y_loadings().shape[1])]
Y_calExplVar_df
# -
help(ho.nipalsPLS1.Y_calExplVar)
# +
# Get cumulative calibrated explained variance in X
X_cumCalExplVar = model.X_cumCalExplVar()
# Get cumulative calibrated explained variance in X and store in pandas dataframe with row and column names
X_cumCalExplVar_df = pd.DataFrame(model.X_cumCalExplVar())
X_cumCalExplVar_df.columns = ['cumulative calibrated explained variance in X']
X_cumCalExplVar_df.index = ['Comp {0}'.format(x) for x in range(model.X_loadings().shape[1] + 1)]
X_cumCalExplVar_df
# -
help(ho.nipalsPLS1.X_cumCalExplVar)
# +
# Get cumulative calibrated explained variance in Y
Y_cumCalExplVar = model.Y_cumCalExplVar()
# Get cumulative calibrated explained variance in Y and store in pandas dataframe with row and column names
Y_cumCalExplVar_df = pd.DataFrame(model.Y_cumCalExplVar())
Y_cumCalExplVar_df.columns = ['cumulative calibrated explained variance in Y']
Y_cumCalExplVar_df.index = ['Comp {0}'.format(x) for x in range(model.Y_loadings().shape[1] + 1)]
Y_cumCalExplVar_df
# -
help(ho.nipalsPLS1.Y_cumCalExplVar)
# +
# Get cumulative calibrated explained variance for each variable in X
X_cumCalExplVar_ind = model.X_cumCalExplVar_indVar()
# Get cumulative calibrated explained variance for each variable in X and store in pandas dataframe with row and column names
X_cumCalExplVar_ind_df = pd.DataFrame(model.X_cumCalExplVar_indVar())
X_cumCalExplVar_ind_df.columns = X_varNames
X_cumCalExplVar_ind_df.index = ['Comp {0}'.format(x) for x in range(model.X_loadings().shape[1] + 1)]
X_cumCalExplVar_ind_df
# -
help(ho.nipalsPLS1.X_cumCalExplVar_indVar)
# +
# Get calibrated predicted Y for a given number of components
# Predicted Y from calibration using 1 component
Y_from_1_component = model.Y_predCal()[1]
# Predicted Y from calibration using 1 component stored in pandas data frame with row and columns names
Y_from_1_component_df = pd.DataFrame(model.Y_predCal()[1])
Y_from_1_component_df.index = y_objNames
Y_from_1_component_df.columns = y_varNames
Y_from_1_component_df
# +
# Get calibrated predicted Y for a given number of components
# Predicted Y from calibration using 4 component
Y_from_4_component = model.Y_predCal()[4]
# Predicted Y from calibration using 1 component stored in pandas data frame with row and columns names
Y_from_4_component_df = pd.DataFrame(model.Y_predCal()[4])
Y_from_4_component_df.index = y_objNames
Y_from_4_component_df.columns = y_varNames
Y_from_4_component_df
# -
help(ho.nipalsPLS1.X_predCal)
# +
# Get validated explained variance of each component X
X_valExplVar = model.X_valExplVar()
# Get calibrated explained variance in X and store in pandas dataframe with row and column names
X_valExplVar_df = pd.DataFrame(model.X_valExplVar())
X_valExplVar_df.columns = ['validated explained variance in X']
X_valExplVar_df.index = ['Comp {0}'.format(x+1) for x in range(model.X_loadings().shape[1])]
X_valExplVar_df
# -
help(ho.nipalsPLS1.X_valExplVar)
# +
# Get validated explained variance of each component Y
Y_valExplVar = model.Y_valExplVar()
# Get calibrated explained variance in X and store in pandas dataframe with row and column names
Y_valExplVar_df = pd.DataFrame(model.Y_valExplVar())
Y_valExplVar_df.columns = ['validated explained variance in Y']
Y_valExplVar_df.index = ['Comp {0}'.format(x+1) for x in range(model.Y_loadings().shape[1])]
Y_valExplVar_df
# -
help(ho.nipalsPLS1.Y_valExplVar)
# +
# Get cumulative validated explained variance in X
X_cumValExplVar = model.X_cumValExplVar()
# Get cumulative validated explained variance in X and store in pandas dataframe with row and column names
X_cumValExplVar_df = pd.DataFrame(model.X_cumValExplVar())
X_cumValExplVar_df.columns = ['cumulative validated explained variance in X']
X_cumValExplVar_df.index = ['Comp {0}'.format(x) for x in range(model.X_loadings().shape[1] + 1)]
X_cumValExplVar_df
# -
help(ho.nipalsPLS1.X_cumValExplVar)
# +
# Get cumulative validated explained variance in Y
Y_cumValExplVar = model.Y_cumValExplVar()
# Get cumulative validated explained variance in Y and store in pandas dataframe with row and column names
Y_cumValExplVar_df = pd.DataFrame(model.Y_cumValExplVar())
Y_cumValExplVar_df.columns = ['cumulative validated explained variance in Y']
Y_cumValExplVar_df.index = ['Comp {0}'.format(x) for x in range(model.Y_loadings().shape[1] + 1)]
Y_cumValExplVar_df
# -
help(ho.nipalsPLS1.Y_cumValExplVar)
help(ho.nipalsPLS1.X_cumValExplVar_indVar)
# +
# Get validated predicted Y for a given number of components
# Predicted Y from validation using 1 component
Y_from_1_component_val = model.Y_predVal()[1]
# Predicted Y from calibration using 1 component stored in pandas data frame with row and columns names
Y_from_1_component_val_df = pd.DataFrame(model.Y_predVal()[1])
Y_from_1_component_val_df.index = y_objNames
Y_from_1_component_val_df.columns = y_varNames
Y_from_1_component_val_df
# +
# Get validated predicted Y for a given number of components
# Predicted Y from validation using 3 components
Y_from_3_component_val = model.Y_predVal()[3]
# Predicted Y from calibration using 3 components stored in pandas data frame with row and columns names
Y_from_3_component_val_df = pd.DataFrame(model.Y_predVal()[3])
Y_from_3_component_val_df.index = y_objNames
Y_from_3_component_val_df.columns = y_varNames
Y_from_3_component_val_df
# -
help(ho.nipalsPLS1.Y_predVal)
# +
# Get predicted scores for new measurements (objects) of X
# First pretend that we acquired new X data by using part of the existing data and overlaying some noise
import numpy.random as npr
new_X = X[0:4, :] + npr.rand(4, np.shape(X)[1])
np.shape(X)
# Now insert the new data into the existing model and compute scores for two components (numComp=2)
pred_X_scores = model.X_scores_predict(new_X, numComp=2)
# Same as above, but results stored in a pandas dataframe with row names and column names
pred_X_scores_df = pd.DataFrame(model.X_scores_predict(new_X, numComp=2))
pred_X_scores_df.columns = ['Comp {0}'.format(x+1) for x in range(2)]
pred_X_scores_df.index = ['new object {0}'.format(x+1) for x in range(np.shape(new_X)[0])]
pred_X_scores_df
# -
help(ho.nipalsPLS1.X_scores_predict)
# +
# Predict Y from new X data
pred_Y = model.Y_predict(new_X, numComp=2)
# Predict Y from nex X data and store results in a pandas dataframe with row names and column names
pred_Y_df = pd.DataFrame(model.Y_predict(new_X, numComp=2))
pred_Y_df.columns = y_varNames
pred_Y_df.index = ['new object {0}'.format(x+1) for x in range(np.shape(new_X)[0])]
pred_Y_df
# -
| examples/PLSR/.ipynb_checkpoints/PLSR_on_NIR_and_octane_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="w_X--pBaZ3sk"
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
from numpy import linalg as LA
import copy
import torch.optim as optim
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
# + id="giKZEoj0a1KY" colab={"base_uri": "https://localhost:8080/", "height": 519, "referenced_widgets": ["db70d475bbf646219e3bfecae346ea47", "1f48635feb3643a4b056199045a8d137", "b68407b11239445393d10e6e77aa2566", "3580e609be294424955f3b9b3101cbc0", "b16d80b0945d4029ba91e60efa3da6a2", "ab3145656fe24ccd8ae7ef4b0aba839d", "<KEY>", "c0007729ebed49afb1dac65fd88916b5", "35b00be205994418ac5a8f8377fc93a1", "103410d1055d42b1a9805d53d6801f19", "6d472ba346c2421c96f45adb50d62f37", "8adac2a7648946b8b6b53cb5d21ed1c2", "c0a8708a79a249e590799d70747be715", "8b6e122a2cd343a98055b4c0762e345b", "0ba9f37f8e8146f295baca5569ebf3ce", "992133747be24a1587e49dd87536ddc8", "a35ac633cc9d450290eca20bbbe17b45", "f021ba6014c04ae58540d6cb91061731", "<KEY>", "258ce28ad44d4291a5b20d91c3fed92b", "<KEY>", "<KEY>", "0d49888efbf945c3afb61499ee390283", "363de7d73d0842caa085f64cac8c497d", "798ccff58f7d4d8f9192e13002bb0ed4", "<KEY>", "<KEY>", "995a9ddec8c845019e6398d70e50c215", "e8716b7f438746a59fbe485a3a373aec", "8c0a10aa8da24d5b87b7e602e313b3b5", "1086932ec0414f7caedf15a7f3175b4a", "f97998ea36f04f66935a8946be65e690"]} outputId="aa37fa3a-00e8-4fc3-9619-3766372fedc7"
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5), (0.5))])
trainset = torchvision.datasets.MNIST(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.MNIST(root='./data', train=False, download=True, transform=transform)
# + id="GbqCx5Bca2Eh" colab={"base_uri": "https://localhost:8080/"} outputId="9648a180-3bf4-4146-eaef-2433477c11d2"
classes = ('zero','one','two','three','four','five','six','seven','eight','nine')
foreground_classes = {'zero','one'}
fg_used = '01'
fg1, fg2 = 0,1
all_classes = {'zero','one','two','three','four','five','six','seven','eight','nine'}
background_classes = all_classes - foreground_classes
background_classes
# + id="AqOQPp7nd64x"
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle = False)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle = False)
# + id="ACeFPtf1eCEe"
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(6000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
# + id="YHShWR-Oh8Lf"
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
plt.imshow(np.reshape(npimg, (28,28)))
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="SjRlj-MPidSx" outputId="26d729d7-efca-40ee-bba0-6d564396ef24"
foreground_data.shape, foreground_label.shape, background_data.shape, background_label.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="SaEMipIH9L5S" outputId="19c23cb9-26f7-4fb3-d2ad-41ac210ec13e"
imshow(foreground_data[0])
# + id="kW1PGzUL7kXA"
mean_bg = torch.mean(background_data, dim=0, keepdims= True)
std_bg, _ = torch.max(background_data, dim=0, keepdims= True)
# + colab={"base_uri": "https://localhost:8080/"} id="s8FwUW-l7kUm" outputId="f52915e6-0b06-485f-c5d6-7afc5c96e210"
mean_bg.shape, std_bg.shape
# + id="-aM-NeCP7uo2"
foreground_data = (foreground_data - mean_bg) / std_bg
background_data = (background_data - mean_bg) / torch.abs(std_bg)
# + colab={"base_uri": "https://localhost:8080/"} id="qLaRFBGZ7umW" outputId="e3fa6fb6-a93d-4a2f-df62-63c4d90fe952"
foreground_data.shape, foreground_label.shape, background_data.shape, background_label.shape
# + colab={"base_uri": "https://localhost:8080/"} id="HAedI_5W7uje" outputId="40c7b00f-a48a-473d-ad82-e567e24f194e"
torch.sum(torch.isnan(foreground_data)), torch.sum(torch.isnan(background_data))
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="MnDBTsPm7kRw" outputId="2752527c-6b24-4ba7-fbf6-44a141d05907"
imshow(foreground_data[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="oLp8rsaQ7-VR" outputId="59ec99f6-7099-4c54-905d-d16df4348923"
imshow(background_data[0])
# + id="d-UCgJT9eHT4"
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])
j+=1
else:
image_list.append(foreground_data[fg_idx])
label = foreground_label[fg_idx] - fg1 # minus fg1 because our fore ground classes are fg1,fg2,fg3 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
# + id="HES-uRlLeVh8"
desired_num = 20000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
list_set_labels = []
for i in range(desired_num):
set_idx = set()
np.random.seed(i)
bg_idx = np.random.randint(0,47335,8)
set_idx = set(background_label[bg_idx].tolist())
fg_idx = np.random.randint(0,12665)
set_idx.add(foreground_label[fg_idx].item())
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
list_set_labels.append(set_idx)
# + id="FJeVQuFVj9Yy" colab={"base_uri": "https://localhost:8080/"} outputId="95f79674-6d2b-40cd-bd37-34a5272ddbb8"
print(len(mosaic_list_of_images) , len(mosaic_label), len(mosaic_list_of_images[0:10000]))
print(len(fore_idx))
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="Me2gS4_k8Z_L" outputId="cceeab40-177a-4cea-8880-e2fa56df25a7"
imshow(mosaic_list_of_images[0][2])
# + colab={"base_uri": "https://localhost:8080/"} id="b6unmYWy8Luh" outputId="9b3dc49a-bd28-453a-84df-9cd0c75f2859"
mosaic_list_of_images = torch.stack(mosaic_list_of_images)
mosaic_list_of_images.shape
# + id="7qFphyDh8MYW"
mean_train = torch.mean(mosaic_list_of_images[0:10000], dim=0, keepdims= True)
# std_train, _ = torch.max(mosaic_list_of_images_train[0:10000], dim=0, keepdims= True)
mosaic_list_of_images = (mosaic_list_of_images - mean_train) # / torch.abs(std_train
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="yiYPKvlY8a2q" outputId="43ae25f9-54f0-4690-905c-def20b69d008"
imshow(mosaic_list_of_images[0][2])
# + colab={"base_uri": "https://localhost:8080/"} id="0T9wERWz8dFm" outputId="ac0c6b22-5840-4c9e-eadf-273a4d88ef8f"
torch.sum(torch.isnan(mosaic_list_of_images))
# + id="goLIiZQsemTJ"
def create_avg_image_from_mosaic_dataset(mosaic_dataset,labels,foreground_index,dataset_number):
"""
mosaic_dataset : mosaic_dataset contains 9 images 32 x 32 each as 1 data point
labels : mosaic_dataset labels
foreground_index : contains list of indexes where foreground image is present so that using this we can take weighted average
dataset_number : will help us to tell what ratio of foreground image to be taken. for eg: if it is "j" then fg_image_ratio = j/9 , bg_image_ratio = (9-j)/8*9
"""
avg_image_dataset = []
for i in range(len(mosaic_dataset)):
img = torch.zeros([ 28,28], dtype=torch.float64)
for j in range(9):
if j == foreground_index[i]:
img = img + mosaic_dataset[i][j]*dataset_number/9
else :
img = img + mosaic_dataset[i][j]*(9-dataset_number)/(8*9)
avg_image_dataset.append(img)
return avg_image_dataset , labels , foreground_index
# + id="Rl8FVV0Fi5UK"
avg_image_dataset_1 , labels_1, fg_index_1 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0)
avg_image_dataset_2 , labels_2, fg_index_2 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.001)
avg_image_dataset_3 , labels_3, fg_index_3 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.002)
avg_image_dataset_4 , labels_4, fg_index_4 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.004)
avg_image_dataset_5 , labels_5, fg_index_5 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.008)
avg_image_dataset_6 , labels_6, fg_index_6 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.016)
avg_image_dataset_7 , labels_7, fg_index_7 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.032)
avg_image_dataset_8 , labels_8, fg_index_8 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.064)
avg_image_dataset_9 , labels_9, fg_index_9 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.128)
avg_image_dataset_10 , labels_10, fg_index_10 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0.256)
test_dataset , labels , fg_index = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[10000:20000], mosaic_label[10000:20000], fore_idx[10000:20000] , 9)
# + id="8ZAX2z2ijLYX"
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
#self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] #, self.fore_idx[idx]
# + id="TZWIbWTcjZTC"
batch = 512
# training_data = avg_image_dataset_5 #just change this and training_label to desired dataset for training
# training_label = labels_5
traindata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
trainloader_1 = DataLoader( traindata_1 , batch_size= batch ,shuffle=True)
traindata_2 = MosaicDataset(avg_image_dataset_2, labels_2 )
trainloader_2 = DataLoader( traindata_2 , batch_size= batch ,shuffle=True)
traindata_3 = MosaicDataset(avg_image_dataset_3, labels_3 )
trainloader_3 = DataLoader( traindata_3 , batch_size= batch ,shuffle=True)
traindata_4 = MosaicDataset(avg_image_dataset_4, labels_4 )
trainloader_4 = DataLoader( traindata_4 , batch_size= batch ,shuffle=True)
traindata_5 = MosaicDataset(avg_image_dataset_5, labels_5 )
trainloader_5 = DataLoader( traindata_5 , batch_size= batch ,shuffle=True)
traindata_6 = MosaicDataset(avg_image_dataset_6, labels_6 )
trainloader_6 = DataLoader( traindata_6 , batch_size= batch ,shuffle=True)
traindata_7 = MosaicDataset(avg_image_dataset_7, labels_7 )
trainloader_7 = DataLoader( traindata_7 , batch_size= batch ,shuffle=True)
traindata_8 = MosaicDataset(avg_image_dataset_8, labels_8 )
trainloader_8 = DataLoader( traindata_8 , batch_size= batch ,shuffle=True)
traindata_9 = MosaicDataset(avg_image_dataset_9, labels_9 )
trainloader_9 = DataLoader( traindata_9 , batch_size= batch ,shuffle=True)
traindata_10 = MosaicDataset(avg_image_dataset_10, labels_10 )
trainloader_10 = DataLoader( traindata_10 , batch_size= batch ,shuffle=True)
# + id="rCaLHr5Wvxfc"
testdata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
testloader_1 = DataLoader( testdata_1 , batch_size= batch ,shuffle=False)
testdata_2 = MosaicDataset(avg_image_dataset_2, labels_2 )
testloader_2 = DataLoader( testdata_2 , batch_size= batch ,shuffle=False)
testdata_3 = MosaicDataset(avg_image_dataset_3, labels_3 )
testloader_3 = DataLoader( testdata_3 , batch_size= batch ,shuffle=False)
testdata_4 = MosaicDataset(avg_image_dataset_4, labels_4 )
testloader_4 = DataLoader( testdata_4 , batch_size= batch ,shuffle=False)
testdata_5 = MosaicDataset(avg_image_dataset_5, labels_5 )
testloader_5 = DataLoader( testdata_5 , batch_size= batch ,shuffle=False)
testdata_6 = MosaicDataset(avg_image_dataset_6, labels_6 )
testloader_6 = DataLoader( testdata_6 , batch_size= batch ,shuffle=False)
testdata_7 = MosaicDataset(avg_image_dataset_7, labels_7 )
testloader_7 = DataLoader( testdata_7 , batch_size= batch ,shuffle=False)
testdata_8 = MosaicDataset(avg_image_dataset_8, labels_8 )
testloader_8 = DataLoader( testdata_8 , batch_size= batch ,shuffle=False)
testdata_9 = MosaicDataset(avg_image_dataset_9, labels_9 )
testloader_9 = DataLoader( testdata_9 , batch_size= batch ,shuffle=False)
testdata_10 = MosaicDataset(avg_image_dataset_10, labels_10 )
testloader_10 = DataLoader( testdata_10 , batch_size= batch ,shuffle=False)
# + id="QTmJHboqv5Ga"
testdata_11 = MosaicDataset(test_dataset, labels )
testloader_11 = DataLoader( testdata_11 , batch_size= batch ,shuffle=False)
# + id="Nej7sxdrmz1b"
class Module2(nn.Module):
def __init__(self):
super(Module2, self).__init__()
# self.conv1 = nn.Conv2d(1, 6, 5)
# self.pool = nn.MaxPool2d(2, 2)
# self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(28*28, 2)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.zeros_(self.fc1.bias)
# self.fc2 = nn.Linear(64, 3)
# self.fc3 = nn.Linear(64, 10)
# self.fc4 = nn.Linear(10,3)
def forward(self,z):
# y1 = self.pool(F.relu(self.conv1(z)))
# y1 = self.pool(F.relu(self.conv2(y1)))
y1 = z.view(-1, 28*28)
y1 = (self.fc1(y1))
# y1 = (self.fc2(y1))
# y1 = F.relu(self.fc3(y1))
# y1 = self.fc4(y1)
return y1
# + id="5JKIMetphOYV"
def calculate_loss(dataloader,model,criter):
model.eval()
r_loss = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = model(inputs)
loss = criter(outputs, labels)
r_loss += loss.item()
return r_loss/i
# + id="Hiif2MO5nAis"
def test_all(number, testloader,net):
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
pred = np.concatenate(pred, axis = 0)
out = np.concatenate(out, axis = 0)
print("pred: ", pred, np.unique(pred))
print("out: ", out, np.unique(pred))
print("correct: ", correct," total: ", total, "sum: ", sum(pred==out))
print('Accuracy of the network on the 10000 test dataset %d: %d %%' % (number , 100 * correct / total))
# + id="ut2fHl-lnDpL"
def train_all(trainloader, ds_number, testloader_list):
print("--"*40)
print("training on data set ", ds_number)
torch.manual_seed(12)
net = Module2().double()
net = net.to("cuda")
criterion_net = nn.CrossEntropyLoss()
optimizer_net = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
acti = []
loss_curi = []
epochs = 1000
running_loss = calculate_loss(trainloader,net,criterion_net)
loss_curi.append(running_loss)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
net.train()
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_net.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion_net(outputs, labels)
# print statistics
running_loss += loss.item()
loss.backward()
optimizer_net.step()
running_loss = calculate_loss(trainloader,net,criterion_net)
if(epoch%200 == 0):
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.005:
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
break
print('Finished Training')
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 train images: %d %%' % ( 100 * correct / total))
for i, j in enumerate(testloader_list):
test_all(i+1, j,net)
print("--"*40)
return loss_curi
# + colab={"base_uri": "https://localhost:8080/"} id="5zjRI237IyAW" outputId="a53433f2-fb9b-41ef-c45b-30f032fc0cfc"
torch.manual_seed(12)
net = Module2().double()
net = net.to("cuda")
test_all(11, testloader_11 ,net)
# + id="shv3ygDijMAg"
train_loss_all=[]
testloader_list= [ testloader_1, testloader_2, testloader_3, testloader_4, testloader_5, testloader_6,
testloader_7, testloader_8, testloader_9, testloader_10, testloader_11]
# + id="kPIoOsmrnLyX" colab={"base_uri": "https://localhost:8080/"} outputId="2a7d95bf-5077-4be0-a066-d33544474591"
train_loss_all.append(train_all(trainloader_1, 1, testloader_list))
train_loss_all.append(train_all(trainloader_2, 2, testloader_list))
train_loss_all.append(train_all(trainloader_3, 3, testloader_list))
train_loss_all.append(train_all(trainloader_4, 4, testloader_list))
train_loss_all.append(train_all(trainloader_5, 5, testloader_list))
train_loss_all.append(train_all(trainloader_6, 6, testloader_list))
train_loss_all.append(train_all(trainloader_7, 7, testloader_list))
train_loss_all.append(train_all(trainloader_8, 8, testloader_list))
train_loss_all.append(train_all(trainloader_9, 9, testloader_list))
train_loss_all.append(train_all(trainloader_10, 10, testloader_list))
# + id="DlDXudp5naB3" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="69a0085b-df06-40d3-a252-36a5c4ebb477"
# %matplotlib inline
fig = plt.figure()
for i,j in enumerate(train_loss_all):
plt.plot(j,label ="dataset "+str(i+1))
plt.xlabel("Epochs")
plt.ylabel("Training_loss")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
fig.savefig("Figure.pdf")
# + id="Fe45SA2hiUTN"
| CODS_COMAD/CIN/Mnist_10k_linear_alpha_0_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
experiment_name = "uci_all"
results_dir = './results/' + experiment_name + '/'
# +
import seaborn as sns
sns.set_theme(style="ticks")
sns.set_context("poster")
import matplotlib.pyplot as plt
import pandas as pd
import pickle
import os
import numpy as np
import pandas
import runs.config_experiments_uci_all as run
experiment_list = run.config_experiments(results_dir, create_json=False)
# +
if not os.path.isfile('./results/datasets/UCI/datasets_info.csv'):
import json
datasets = []
for data_set in range(20,66):
with open('/vast/robustness/configs_datasets/' + str(data_set) + '.json') as config_file:
config = json.load(config_file)
tmpX = np.shape(np.genfromtxt('/vast/robustness/datasets/UCI/imp_' + config["name_file"]
+ '_' + "trainX.csv", delimiter=','))
config["num_examples"] = tmpX[0]
config["num_features"] = tmpX[1]
datasets.append(config)
df = pd.DataFrame(datasets)
df.to_csv('./results/datasets/UCI/datasets_info.csv')
else:
df = pd.read_csv('./results/datasets/UCI/datasets_info.csv')
name_attacks = ["l1_pgd_norm", "l1_fgm_norm","l2_pgd_norm", "l2_fgm_norm", "linf_pgd","l2_pgd", "linf_fgsm", "l2_fgm"]#
# -
df[(df.num_features>50) & (df.num_examples > 500)]
df.to_csv('dataset.csv',index=False)
# +
list_entries = []
for net in ['OneLayer', 'ThreeLayer']:
for dataset_id in list(df.dataset_id):
for idx, exp in enumerate(experiment_list):
if exp['data_set'] == dataset_id:
break
for attack in name_attacks:
file_name = results_dir + experiment_list[idx]['model_name'] + '/results/acc_' + 'val' + '_' + attack + '.pkl'
if not os.path.isfile(file_name):
print("Missing!! " + file_name)
continue
with open(file_name, 'rb') as f:
tmp = pickle.load(f)
for cv_epsilon in list(tmp.keys()):
# Hash table of parameters
parameters = {"epsilon": {}, "backbone": {}, "initial_learning_rate": {},
"robust_training": {}, "type_robust": {}, "epsilon_pgd_training":{}}
to_exclude = []
experiment_list_tmp = [element for i, element in enumerate(experiment_list) if i not in to_exclude]
for exp in experiment_list_tmp:
if not exp['data_set'] == dataset_id:
continue
for kk in parameters.keys():
if exp[kk] in parameters[kk]:
parameters[kk][exp[kk]].append(int(exp["model_name"]))
else:
parameters[kk][exp[kk]] = [int(exp["model_name"])]
# For all methods, do cross-val and create an entry of the results
backbones = [net, net + '+pgd']
for backbone in backbones:
for robust_training in [True, False]:
if robust_training:
type_robust_trainings = ['l1','linf', "certificate", "grad"]
else:
type_robust_trainings = ['none']
for type_robust in type_robust_trainings:
if (backbone == 'Madry' and robust_training == True) or \
(backbone == 'CNN+clipping' and robust_training == False):
continue
if robust_training==False:
ids = list(set(parameters["backbone"][backbone]) &
set(parameters["robust_training"][False]))
else:
ids = list(set(parameters["backbone"][backbone]) &
set(parameters["robust_training"][True])&
set(parameters["type_robust"][type_robust]))
if backbone == net + '+pgd' and robust_training == True:
continue
if ids == []:
continue
#print(ids)
# Cross-validation among learning rates and epsilons:
best_acc = -1
best_id = ids[0]
for id in ids:
file_name = results_dir + experiment_list[id]['model_name'] + '/results/acc_' + 'val' + '_' + attack + '.pkl'
if not os.path.isfile(file_name):
print("Missing!! " + file_name)
continue
with open(file_name, 'rb') as f:
tmp = pickle.load(f)
acc = tmp[cv_epsilon]
if acc>best_acc:
best_id = id
best_acc = acc
if best_acc == -1:
continue
if (robust_training == False) & (backbone==net):
name_legend = 'vanilla'
elif backbone== net + '+pgd':
name_legend = 'pgd'
else:
if type_robust=='certificate':
name_legend = 'RUB'
elif type_robust=='linf':
name_legend = 'aRUB_Linf'
elif type_robust=='grad':
name_legend = 'grad'
else:
name_legend = 'aRUB_L1'
entry = {"dataset": dataset_id,
"net": net,
"learning_rate": experiment_list[best_id]['initial_learning_rate'],
"robust_training": name_legend,
"epsilon": experiment_list[best_id]['epsilon'],
"epsilon_pgd_training": experiment_list[best_id]['epsilon_pgd_training']}
dataset = "test"
entry["attack"] = attack
entry["experiment_id"] = best_id
with open(results_dir + experiment_list[best_id]['model_name'] + '/results/acc_' + dataset + '_' +
attack + '.pkl', 'rb') as f:
tmp = pickle.load(f)
entry["test_epsilon"] = cv_epsilon
entry["accuracy"] = 100*tmp[cv_epsilon]
with open(results_dir + experiment_list[best_id]['model_name'] +
'/results/training_time.pkl', 'rb') as f:
tmp = pickle.load(f)
entry["images_per_second"] = np.mean(tmp)
entry["std_images_per_second"] = np.std(tmp)
list_entries.append(entry.copy())
df_results = pd.DataFrame.from_dict(list_entries)
# + pycharm={"name": "#%%\n"}
df_results.to_csv('uci_norm.csv', index=False)
| notebooks/UCI.ipynb |
-- ---
-- jupyter:
-- jupytext:
-- text_representation:
-- extension: .hs
-- format_name: light
-- format_version: '1.5'
-- jupytext_version: 1.14.4
-- kernelspec:
-- display_name: Haskell
-- language: haskell
-- name: haskell
-- ---
-- +
{-# LANGUAGE LambdaCase #-}
import Control.Monad.Trans.State.Strict
import qualified Data.IntMap.Strict as M
import qualified Data.Vector as V
import Data.Foldable (for_)
import Data.Functor (void)
-- -
graph = M.fromList
[ (1, [2])
, (2, [3])
, (3, [1])
, (4, [2, 3, 5])
, (5, [4, 6])
, (6, [7, 3])
, (7, [6])
, (8, [5, 7, 8])
]
-- +
data TState = TState
{ index :: Int
, stack :: [Int]
, stackSet :: V.Vector Bool
, stackTop :: Int
, indices :: V.Vector (Maybe Int)
, lowlinks :: V.Vector (Maybe Int)
, sccs :: [[Int]]
} deriving (Eq, Show)
data Event
= IncrementIndex
| InsertIndex Int (Maybe Int)
| InsertLowlink Int (Maybe Int)
| Push Int
| Pop
| OutputSCC [Int]
deriving (Eq, Show)
data EState = EState
{ current :: TState
, history :: [Event]
} deriving (Eq, Show)
whenM :: Monad m => m Bool -> m () -> m ()
whenM boolM block = boolM >>= \b -> if b then block else return ()
insert :: V.Vector a -> Int -> a -> V.Vector a
insert vector index value = vector V.// [(index, value)]
apply :: Event -> TState -> TState
apply event state = case event of
IncrementIndex -> state { index = index state + 1 }
InsertIndex k v -> state { indices = insert (indices state) k v }
InsertLowlink k v -> state { lowlinks = insert (lowlinks state) k v }
Push i -> state
{ stack = i : (stack state)
, stackSet = insert (stackSet state) i True
}
Pop -> let i = head (stack state) in state
{ stack = tail (stack state)
, stackSet = insert (stackSet state) i False
, stackTop = i
}
OutputSCC scc -> state { sccs = sccs state ++ [scc] }
initial = TState 0 [] falses undefined nothings nothings []
where
size = (fst $ M.findMax graph) + 1
falses = V.replicate size False
nothings = V.replicate size Nothing
emit :: Event -> State EState ()
emit event = void . modify' $ \s -> s
{ current = apply event (current s)
, history = event : (history s)
}
incrementIndex = emit IncrementIndex
insertIndex k v = emit $ InsertIndex k v
insertLowlink k v = emit $ InsertLowlink k v
push i = emit $ Push i
pop = emit Pop
outputSCC scc = emit $ OutputSCC scc
query accessor i = (V.! i) . accessor <$> gets current
-- +
tarjan :: M.IntMap [Int] -> EState
tarjan graph = flip execState (EState initial []) $
for_ (M.keys graph) $ \v ->
whenM ((Nothing==) <$> query indices v) $
strongConnect v graph
strongConnect :: Int -> M.IntMap [Int] -> State EState ()
strongConnect v graph = do
incrementIndex
i <- index <$> gets current
insertIndex v (Just i)
insertLowlink v (Just i)
push v
for_ (graph M.! v) $ \w ->
query indices w >>= \case
Nothing -> do
strongConnect w graph
insertLowlink v =<< (min <$> query lowlinks v <*> query lowlinks w)
Just{} -> whenM (query stackSet w) $
insertLowlink v =<< (min <$> query lowlinks v <*> query indices w)
whenM ((==) <$> query lowlinks v <*> query indices v) $
outputSCC =<< addSCC v []
addSCC :: Int -> [Int] -> State EState [Int]
addSCC v scc = do
pop
w <- stackTop <$> gets current
if w == v
then return (w:scc)
else addSCC v (w:scc)
-- -
complete = tarjan graph
sccs . current $ complete
void $ traverse print (reverse . history $ complete)
| tarjan/TarjansSCC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
"""
Author: <NAME>
This file is under MIT License.
Link: https://github.com/shaimaa-elbaklish/funcMinimization/blob/main/LICENSE.md
"""
# + id="PYJiN7auQkew"
import numpy as np
import plotly.graph_objects as go
# -
# ## Benchmark Multimodal Functions Available
# | Function | Dimension | Bounds | Optimal Function Value |
# | -------- | --------- | ------ | ---------------------- |
# | $$ f_{1} = 4x_1^2 - 2.1x_1^4 + \frac{1}{3}x_1^6 + x_1 x_2 - 4x_2^2 + 4 x_2^4 $$ | 2 | [-5, 5] | -1.0316 |
# | $$ f_{2} = (x_2 - \frac{5.1}{4\pi^2}x_1^2 + \frac{5}{\pi}x_1 -6)^2 +10(1 - \frac{1}{8\pi})\cos{x_1} + 10 $$ | 2 | [-5, 5] | 0.398 |
# | $$ f_{3} = -\sum_{i=1}^{4} c_i exp(-\sum_{j=1}^{3} a_{ij}(x_j - p_{ij})^2) $$ | 3 | [1, 3] | -3.86 |
# | $$ f_{4} = -\sum_{i=1}^{4} c_i exp(-\sum_{j=1}^{6} a_{ij}(x_j - p_{ij})^2) $$ | 6 | [0, 1] | -3.32 |
# | $$ f_{5} = -\sum_{i=1}^{7} [(X - a_i)(X - a_i)^T + c_i]^{-1} $$ | 4 | [0, 10] | -10.4028 |
class Function:
def __init__(self, x = None, n = 2, lb = np.array([-5, -5]), ub = np.array([5, 5])):
self.n_x = n
if x is not None:
assert(x.shape[0] == self.n_x)
self.fvalue = self.getFValue(x)
self.x = x
self.fvalue = None
assert(lb.shape[0] == self.n_x)
self.lb = lb
assert(ub.shape[0] == self.n_x)
self.ub = ub
def setBenchmarkFunction(self, f_name = "f1"):
benchmarks = {
"f1": [2, np.array([-5, -5]), np.array([5, 5])],
"f2": [2, np.array([-5, -5]), np.array([5, 5])],
"f3": [3, 1*np.ones(shape=(3,)), 3*np.ones(shape=(3,))],
"f4": [6, np.zeros(shape=(6,)), 1*np.ones(shape=(6,))],
"f5": [4, np.zeros(shape=(4,)), 10*np.ones(shape=(4,))]
}
self.benchmark_selected = f_name
[self.n_x, self.lb, self.ub] = benchmarks.get(f_name, benchmarks.get("f1"))
def isFeasible(self, x):
return np.all(x >= self.lb) and np.all(x <= self.ub)
def getFValue(self, x):
if self.benchmark_selected is None:
func_value = 4*x[0]**2 - 2.1*x[0]**4 + (x[0]**6)/3 + x[0]*x[1] - 4*x[1]**2 + 4*x[1]**4
return func_value
benchmarks_coeffs = {
"f3": {"a": np.array([[3, 10, 30], [0.1, 10, 35], [3, 10, 30], [0.1, 10, 35]]),
"c": np.array([1, 1.2, 3, 3.2]),
"p": np.array([[0.3689, 0.117, 0.2673], [0.4699, 0.4387, 0.747], [0.1091, 0.8732, 0.5547], [0.03815, 0.5743, 0.8828]])},
"f4": {"a": np.array([[10, 3, 17, 3.5, 1.7, 8], [0.05, 10, 17, 0.1, 8, 14], [3, 3.5, 1.7, 10, 17, 8], [17, 8, 0.05, 10, 0.1, 14]]),
"c": np.array([1, 1.2, 3, 3.2]),
"p": np.array([[0.1312, 0.1696, 0.5569, 0.0124, 0.8283, 0.5886], [0.2329, 0.4135, 0.8307, 0.3736, 0.1004, 0.9991], [0.2348, 0.1415, 0.3522, 0.2883, 0.3047, 0.6650], [0.4047, 0.8828, 0.8732, 0.5743, 0.1091, 0.0381]])},
"f5": {"a": np.array([[4, 4, 4, 4], [1, 1, 1, 1], [8, 8, 8, 8], [6, 6, 6, 6], [3, 7, 3, 7], [2, 9, 2, 9], [5, 5, 3, 3], [8, 1, 8, 1], [6, 2, 6, 2], [7, 3.6, 7, 3.6]]),
"c": np.array([0.1, 0.2, 0.2, 0.4, 0.4, 0.6, 0.3, 0.7, 0.5, 0.5])}
}
benchmarks = {
"f1": lambda z: 4*z[0]**2 - 2.1*z[0]**4 + (z[0]**6)/3 + z[0]*z[1] - 4*z[1]**2 + 4*z[1]**4,
"f2": lambda z: (z[1] - (5.1/(4*np.pi**2))*z[0]**2 + (5/np.pi)*z[0] -6)**2 + 10*(1 - (1/(8*np.pi)))*np.cos(z[0]) + 10,
"f3": lambda z: -np.sum(benchmarks_coeffs["f3"]["c"] * np.exp(-np.sum(list(map(lambda ai, pi: ai*(z - pi)**2, benchmarks_coeffs["f3"]["a"], benchmarks_coeffs["f3"]["p"])), axis=1))),
"f4": lambda z: -np.sum(benchmarks_coeffs["f4"]["c"] * np.exp(-np.sum(list(map(lambda ai, pi: ai*(z - pi)**2, benchmarks_coeffs["f4"]["a"], benchmarks_coeffs["f4"]["p"])), axis=1))),
"f5": lambda z: -np.sum(list(map(lambda ai, ci: 1/((z - ai) @ (z - ai).T + ci), benchmarks_coeffs["f5"]["a"], benchmarks_coeffs["f5"]["c"])))
}
func_value = benchmarks.get(self.benchmark_selected)(x)
return func_value
def initRandomSoln(self):
self.x = np.random.rand(self.n_x) * (self.ub - self.lb) + self.lb
assert(self.isFeasible(self.x))
self.fvalue = self.getFValue(self.x)
def getNeighbourSoln(self):
r = np.random.rand(self.n_x)
x_new = self.x + r * (self.ub - self.x) + (1 - r) * (self.lb - self.x)
assert(self.isFeasible(x_new))
return x_new
class SimulatedAnnealing:
def __init__(self, problem, max_iter = 100, init_temp = 100, final_temp = 1e-03, iter_per_temp = 5, cooling_schedule = "linear", beta = 5, alpha = 0.9):
self.problem = problem
self.max_iter = max_iter
self.init_temp = init_temp
self.final_temp = final_temp
self.iter_per_temp = iter_per_temp
self.cooling_schedule = cooling_schedule
self.beta = min(beta, (init_temp - final_temp)/max_iter)
self.alpha = alpha
self.curr_temp = None
self.sols = None
self.fvalues = None
self.best_sol = None
self.best_fvalue = None
def cool_down_temp(self, curr_iter):
schedules = {
"linear": self.curr_temp - self.beta,
"geometric": self.curr_temp * self.alpha,
"exponential": self.curr_temp / (1 + self.beta*self.curr_temp),
"hybrid": (curr_iter*self.curr_temp/(curr_iter+1)) if curr_iter <= 0.5*self.max_iter else (self.init_temp*(self.alpha**curr_iter))
}
return schedules.get(self.cooling_schedule, schedules.get("linear"))
def perform_algorithm(self):
self.problem.initRandomSoln()
self.curr_temp = self.init_temp
self.sols = [self.problem.x]
self.fvalues = [self.problem.fvalue]
self.best_sol = self.problem.x
self.best_fvalue = self.problem.fvalue
for iter in range(self.max_iter):
for _ in range(self.iter_per_temp):
sol_neighbour = self.problem.getNeighbourSoln()
fvalue_neighbour = self.problem.getFValue(sol_neighbour)
if fvalue_neighbour < self.fvalues[-1]:
# neighbour is better and is accpeted
self.sols.append(sol_neighbour)
self.fvalues.append(fvalue_neighbour)
else:
p = np.exp(-(fvalue_neighbour - self.fvalues[-1]) / self.curr_temp)
if np.random.rand() < p:
# neighbour is worse and is accepted according to probability
self.sols.append(sol_neighbour)
self.fvalues.append(fvalue_neighbour)
else:
# neighbour is worse and is rejected
self.sols.append(self.sols[-1])
self.fvalues.append(self.fvalues[-1])
# update best solution reached so far
if self.fvalues[-1] < self.best_fvalue:
self.best_sol = self.sols[-1]
self.best_fvalue = self.fvalues[-1]
# update temperature
self.curr_temp = self.cool_down_temp(iter)
def visualize(self):
# convergence plot
fig1 = go.Figure(data=go.Scatter(x=np.arange(0, self.max_iter*self.iter_per_temp), y=self.fvalues, mode="lines"))
fig1.update_layout(
title="Convergence Plot",
xaxis_title="Iteration Number",
yaxis_title="Objective Function Value"
)
fig1.show()
problem = Function()
problem.setBenchmarkFunction(f_name="f4")
SA = SimulatedAnnealing(problem, max_iter=500, init_temp=100, final_temp=1e-03, iter_per_temp=10, cooling_schedule="geometric", alpha=0.95)
SA.perform_algorithm()
print(SA.best_sol)
print(SA.best_fvalue)
SA.visualize()
| SA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# # Query Marius Output for WN18 #
# This notebook shows the steps of doing inferencing with Marius output for dataset *WN18*. The example inference we used here has a source node name of *__wisconsin_NN_2* and a relation type of *_instance_hypernym*.
#
# Note: the training over the dataset *WN18* must be done before running this script. Use the following commands to perform the training in Marius root directory:
#
#
# marius_preprocess ./output_dir --dataset wn18 -gc CPU --training.num_epochs=10
# marius_train ./outut_dir/wn18_cpu.ini
# ### Import function ###
import marius_infer as mi
# ### Get embedding of nodes and relations ###
node_embeddings, node_emb_dict = mi.tensor_from_file("node")
relation_embeddings, rel_emb_dict = mi.tensor_from_file("rel")
# ### Obtain the embedding vectors for given node and relation in our inference example ###
# +
src_node = "__wisconsin_NN_2"
relation = "_instance_hypernym"
src_emb = mi.lookup_embedding("node", src_node, node_emb_dict)
rel_emb = mi.lookup_embedding("rel", relation, rel_emb_dict)
# -
# ### Get top 10 nodes and similarity scores that are inferenced based on given node and relation ###
scores, topk = mi.infer_topk_nodes(3, src_emb, rel_emb, node_embeddings)
topk
scores
# +
#__scandinavia_NN_2, _member_meronym, __kingdom_of_denmark_NN_1
src_node = "__scandinavia_NN_2"
relation = "_member_meronym"
src_emb = mi.lookup_embedding("node", src_node, node_emb_dict)
rel_emb = mi.lookup_embedding("rel", relation, rel_emb_dict)
scores, topk = mi.infer_topk_nodes(3, src_emb, rel_emb, node_embeddings)
topk
# +
#__kobenhavn_NN_1, _instance_hypernym, __national_capital_NN_1
src_node = "__kobenhavn_NN_1"
relation = "_instance_hypernym"
src_emb = mi.lookup_embedding("node", src_node, node_emb_dict)
rel_emb = mi.lookup_embedding("rel", relation, rel_emb_dict)
scores, topk = mi.infer_topk_nodes(3, src_emb, rel_emb, node_embeddings)
topk
| examples/inference/wn18/query_deom.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''tf21'': conda)'
# name: python_defaultSpec_1599484198269
# ---
# +
# from preprocessing_dev import edge_detection
# -
import numpy as np
from data_loader import load_npy
import matplotlib.pyplot as plt
from pathlib import Path
import cv2
DATASET_PATH = Path("C:/Users/josep/Documents/work/crate_classifier_dev/outputs/backup/images_3")
images = load_npy(DATASET_PATH / "images_3_part_1/dataset_images_aug.npy")
annots = load_npy(DATASET_PATH / "images_3_part_1/dataset_annots_aug.npy")
index = 7
sample_img = images[index,:,:,3:6]
plt.imshow(sample_img/255)
plt.show()
# + tags=[]
#img_ch = np.expand_dims(sample_img[:,:,0], axis =-1)
img_ch = np.uint8(images[:,:,0])
# + tags=[]
plt.figure(figsize=(12,12))
j = 0
edges_arr = []
for i in range(9):
plt.subplot(5,4,j+1)
img_ch = np.uint8(images[index,:,:,i])
plt.imshow(img_ch)
plt.colorbar()
plt.subplot(5,4,j+2)
edges = cv2.Canny(img_ch, 44, 89) # sigma 0.33
edges_arr.append(edges)
plt.imshow(edges)
plt.colorbar()
j += 2
plt.subplot(5,4,j+1)
sample_img = images[index,:,:,3:6]
plt.imshow(sample_img/255)
edges_arr = np.asarray(edges_arr)
summed = edges_arr.sum(axis=0)/(255*9) # combining
summed = np.where(summed<0.4, 0, summed) # clipping
plt.subplot(5,4,j+2)
plt.imshow(summed)
plt.colorbar()
plt.show()
# + tags=[]
edges_arr = np.asarray(edges_arr)
summed = edges_arr.sum(axis=0)/(255*9) # combining
summed = np.where(summed<0.4, 0, 1) # clipping
# -
product = cv2.bitwise_and(sample_img, sample_img, mask= np.uint8(summed))
scaling = 0.8
denom = scaling*255 + 255
res = (product*scaling + sample_img)/denom
# + tags=[]
plt.imshow(res)
plt.colorbar()
plt.show()
# -
| dev/dev_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
all_base_matrices = np.concatenate((pod.matrices, sym_nap_hex.matrices_hex), axis=0)[:, :-1, :-1]
all_labels = np.concatenate((pod.labels, sym_nap_hex.labels_hex))
trans_amount = len(pod.matrices)
empty_arr = np.zeros((4,4))
full_list = np.zeros(len(sym_nap.all_matrix_translations) * len(pod.matrices) *16, dtype=int).reshape(-1, 4, 4)
full_list_labels = [""] * len(full_list)
# +
ID = np.eye(4)
for i in range(len(sym_nap.all_matrix_translations)):
for j in range(len(pod.matrices)):
combination = pod.matrices[j] + sym_nap.all_matrix_translations[i] - ID
place_in_array = trans_amount*i + j
full_list[place_in_array] = combination
name_m = pod.labels[j]
name_t = sym_nap.all_labels_trans[i].lstrip("t")
full_list_labels[place_in_array] = f'_matrix_{name_m}{name_t} = matrices_dict["{name_m}"] + _translation_{name_t} #{place_in_array}'
print(len(full_list))
for mat in full_list_labels:
print(mat, end='\n')
# -
mydict_mats = {matrix.tostring(): label for matrix, label in zip(pod.matrices, pod.labels)}
mydict_trans = {tr.tostring(): label for tr, label in zip(sym_nap.all_translations, sym_nap.all_labels_trans)}
# +
from collections import deque
from itertools import combinations, product
import podstawa as pod
import symetrie_napisane as sym_nap
# import symetrie_napisane_hex as sym_nap_hex
import symetrie_generowane as sym_gen
import numpy as np
def mult(M1, M2):
M = M1 @ M2
mask = M[:-1, -1] < 0
M[:-1, -1][mask] += 12
mask = M[:-1, -1] >= 12
M[:-1, -1][mask] -= 12
mask0 = M == 0
M[mask0] -= 1
M[mask0] += 1
return M
def inverse(mat):
new_mat = mat.copy()
rot = new_mat[:-1, :-1]
rot = rot.T
trans = mat[:-1, -1] * -1
mask_trans = trans < 0
trans[mask_trans] += 12
new_mat[:-1, :-1] = rot
new_mat[:-1, -1] = trans
return new_mat
def mat2name(mat, mapper):
mat2 = inverse(mat)
str_temp = mat.tostring()
str_temp2 = mat2.tostring()
val = mapper.get(str_temp)
val2 = mapper.get(str_temp2)
return val if val else val2
all_matrices_with_missing = sym_gen.all_missing_matrices + sym_nap.all_matrices
all_matrices_with_missing_labels = sym_gen.all_missing_matrices_labels + sym_nap.names
import pandas as pd
def translate_matrix_to_name():
all_hashed_M = {M.tostring(): name for M, name in zip(all_matrices_with_missing, all_matrices_with_missing_labels)}
l = np.zeros(len(all_hashed_M.keys()) * 16, dtype=int).reshape(-1, 4, 4)
l_labels = [""] * len(l)
for index, mat in enumerate(all_hashed_M.keys()):
l[index] = np.frombuffer(mat, dtype=int).reshape(4,4)
size = len(l)
multi_table = np.empty((size, size), dtype=object)
for ix in range(size):
for iy in range(size):
matrix = M(l[ix], l[iy])
result = mat2name(matrix, all_hashed_M)
multi_table[ix, iy] = result
return multi_table, all_hashed_M
result, labels_from_function = translate_matrix_to_name()
data = pd.DataFrame(result)
data.columns = labels_from_function.values()
data.index = labels_from_function.values()
# data
class Queue:
def __init__(self, *names):
self.queue = deque([(el1, el2) for el1, el2 in product(names, repeat=2)])
self.size = len(self.queue)
def is_not_empty(self):
return self.size != 0
def enqueue(self, item):
self.queue.append(item)
self.size += 1
def add_combinations(self, addition):
for name in self.uniques:
self.enqueue((name, addition))
self.enqueue((addition, name))
self.enqueue((addition, addition))
def dequeue(self):
self.size -= 1
return self.queue.popleft()
def __iter__(self):
return self
def __next__(self):
if self.is_not_empty():
return self.dequeue()
else:
raise StopIteration
class mycontainer:
def __init__(self, table, *names):
self.tablica = table
self.queue = Queue(names)
self.uniques = set(names)
self.origin = names
# self.calc()
def calc(self):
for name1, name2 in self.queue:
wynik = self.tablica[name1][name2]
if wynik not in self.uniques:
self.expand(wynik)
def expand(self, item):
self.queue.add_combinations(item)
self.uniques.add(item)
def __repr__(self):
return str(self.queue)
def __contains__(self, item):
return item in self.uniques
class mycontainer:
def __init__(self, table, *names):
self.tablica = table
self.queue = deque([(el1, el2) for el1, el2 in product(names, repeat=2)])
self.size = len(self.queue)
self.uniques = set(names)
self.origin = names
# self.calc()
def calc(self):
for name1, name2 in self.queue:
wynik = self.tablica[name1][name2]
if wynik not in kolejka:
kolejka.expand(wynik)
def is_not_empty(self):
return self.size != 0
def expand(self, item):
self.add_combinations(item)
self.uniques.add(item)
def add_combinations(self, addition):
for name in self.uniques:
self.enqueue((name, addition))
self.enqueue((addition, name))
self.enqueue((addition, addition))
def __repr__(self):
return str(self.queue)
def enqueue(self, item):
self.queue.append(item)
self.size += 1
def dequeue(self):
self.size -= 1
return self.queue.popleft()
def __iter__(self):
return self
def __next__(self):
if self.is_not_empty():
return self.dequeue()
else:
raise StopIteration
def __contains__(self, item):
return item in self.uniques
from time import time
class Grupa:
def __init__(self, konstruktory, all_syms):
self.konstruktory = konstruktory
self.all_syms = set(all_syms)
def __repr__(self):
return ' '.join((str(self.konstruktory), str(self.all_syms)))
def __eq__(self, other):
return self.all_syms == other.all_syms
def __hash__(self):
return hash(tuple(sorted(self.all_syms)))
@classmethod
def from_queue(cls, queue):
return cls(queue.origin, queue.uniques)
class Timer:
def __init__(self):
self.start = time()
self.mid = time()
self.prev = 0
def __call__(self, num):
if num == self.prev:
starter = round((time() - self.start) / 60, 1)
mider = round(time() - self.mid, 1)
self.mid = time()
print(self.prev, starter, mider)
self.prev += 1
zespol = [set() for _ in range(500)]
mytimer = Timer()
# kolejak osobno kurde
for num1, num2 in combinations(range(len(data.columns)), 2):
mytimer(num1)
name1, name2 = [data.columns[num1], data.columns[num2]]
tabliczka = mycontainer(data, name1, name2)
tabliczka.calc()
num = len(tabliczka.uniques)
zespol[num].add(Grupa.from_queue(tabliczka))
# print(num, zespol)
break
| krysztalki/workDir/proba/tabliczka mnozenia/reduce_cell_tabliczka_mnozenia-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import binascii
from PIL import Image
from IPython.display import display
file = os.path.join(os.getcwd(), 'noname')
def map_to_exe(file):
if not os.path.exists(file):
return
img = Image.open(file, 'r')
print('map — noname png')
display(img)
pixels = img.load()
data = img.info['comment']
print('png comment: {}...'.format(data[:12]))
coords = [(int(data[i:i+3]), int(data[i+3:i+6])) for i in range(0, len(data), 6)]
print('coordinates of pixels with yellow shade: {}...'.format(coords[:4]))
hex_data = ''
for x, y in coords:
dc = pixels[x, y][2]
hx = hex(dc)[2:]
if len(hx) < 2:
hx = '0' + hx
hex_data += hx
bin_data = binascii.unhexlify(hex_data.encode())
print('hex data: {}...'.format(hex_data[:15]))
print('bin data: {}...'.format(bin_data[:15]))
pe = os.path.join(os.getcwd(), os.path.splitext(file)[0] + '.exe')
with open(pe, 'wb') as f:
f.write(bin_data)
print('saved to', pe)
map_to_exe(file)
| 6.sem/reversecup2019/2.map/map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python(base)
# language: python
# name: base
# ---
allcountries = []
# !ls Custom
# +
import plotly.graph_objects as go
import time
from PIL import Image
from Custom.GlobalData import *
LASTFILE="COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv"
STAT =LASTFILE[57:-4]
DataIn = open(LASTFILE).readlines()
SEARCH = "Germany"
X=GlobalData(SEARCH)
x=X.split(",")
population=x[4]
print(SEARCH,"Population:", population)
cnt = 0
CNTS=0
counts=[]
for line in DataIn:
if cnt==0:print(line)
cnt=cnt+1
line=line.lstrip(",")
#if SEARCH in line:print(line)
if SEARCH in line:
print(line)
# -
import plotly.graph_objects as go
import time
from PIL import Image
from Custom.GlobalData import *
DATA=[]
counts=[]
cnt=0
LASTFILE="COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv"
DataIn = open(LASTFILE).readlines()
for line in DataIn:
cnt=cnt+1
if cnt>1:
line=line.lstrip(",")
line=line.rstrip("\n")
line="0,0"+line.split("0,0",1)[-1]
entry=line.split(",")
print(entry)
# +
import plotly.graph_objects as go
import time
from PIL import Image
from Custom.GlobalData import *
LASTFILE="COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv"
STAT =LASTFILE[57:-4]
DataIn = open(LASTFILE).readlines()
#SEARCH = input("SEARCH: ")
#SEARCH = "Brazil"
SEARCH = "Spain"
#SEARCH = "Philippine"
#SEARCH = "Ecuador"
#SEARCH = "Germany"
#SEARCH = "Japan"
#SEARCH = "US"
X=GlobalData(SEARCH)
x=X.split(",")
population=x[4]
print(SEARCH,"Population:", population)
cnt = 0
CNTS=0
counts=[]
for line in DataIn:
if cnt==0:print(line)
cnt=cnt+1
line=line.lstrip(",")
#if SEARCH in line:print(line)
if SEARCH in line:
line="0,0"+line.split("0,0",1)[-1]
print
entry=line.split(",")
for num in entry:
counts.append(int(num))
print(SEARCH," counts/population",int(counts[-1])/int(population))
allcountries.append([SEARCH," counts/population",int(counts[-1])/int(population)])
# +
import plotly.graph_objects as go
import time
from PIL import Image
from Custom.GlobalData import *
LASTFILE="COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv"
STAT =LASTFILE[57:-4]
DataIn = open(LASTFILE).readlines()
#SEARCH = input("SEARCH: ")
#SEARCH = "Brazil"
SEARCH = "Spain"
#SEARCH = "Philippine"
#SEARCH = "Ecuador"
#SEARCH = "Germany"
#SEARCH = "Japan"
#SEARCH = "US"
X=GlobalData(SEARCH)
x=X.split(",")
population=x[4]
print(SEARCH,"Population:", population)
cnt = 0
CNTS=0
counts=[]
for line in DataIn:
if cnt==0:print(line)
cnt=cnt+1
line=line.lstrip(",")
#if SEARCH in line:print(line)
if SEARCH in line:
line="0,0"+line.split("0,0",1)[-1]
print
entry=line.split(",")
for num in entry:
counts.append(int(num))
print(SEARCH," counts/population",int(counts[-1])/int(population))
allcountries.append([SEARCH," counts/population",int(counts[-1])/int(population)])
# +
import plotly.graph_objects as go
import time
from PIL import Image
from Custom.GlobalData import *
LASTFILE="COVID-19/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv"
STAT =LASTFILE[57:-4]
DataIn = open(LASTFILE).readlines()
#SEARCH = input("SEARCH: ")
#SEARCH = "Brazil"
SEARCH = "Spain"
#SEARCH = "Philippine"
#SEARCH = "Ecuador"
#SEARCH = "Germany"
#SEARCH = "Japan"
#SEARCH = "US"
X=GlobalData(SEARCH)
x=X.split(",")
population=x[4]
print(SEARCH,"Population:", population)
cnt = 0
CNTS=0
counts=[]
for line in DataIn:
if cnt==0:print(line)
cnt=cnt+1
line=line.lstrip(",")
#if SEARCH in line:print(line)
if SEARCH in line:
line="0,0"+line.split("0,0",1)[-1]
print
entry=line.split(",")
for num in entry:
counts.append(int(num))
print(SEARCH," counts/population",int(counts[-1])/int(population))
allcountries.append([SEARCH," counts/population",int(counts[-1])/int(population)])
filename0 = time.strftime("images/"+SEARCH+"_"+STAT+"Deaths_%Y%m%d%H%M%S.png")
fig = go.Figure()
fig.add_trace(go.Scatter(y=counts))
fig.add_trace(go.Bar(y=counts))
fig.update_layout(title = SEARCH+' CONDID-19 Deaths')
fig.show()
IncreasePerDay=[]
All = (len(counts))
for x in range(0,All):
try:
Sum = (counts[x+1]-counts[x])
print(Sum, end = " ")
IncreasePerDay.append(Sum)
except:
pass
filename1 = time.strftime("images/"+SEARCH+"_"+STAT+"IncreasePerDay"+"_%Y%m%d%H%M%S.png")
fig = go.Figure()
fig.add_trace(go.Scatter(y=IncreasePerDay))
fig.add_trace(go.Bar(y=IncreasePerDay))
fig.update_layout(title = SEARCH+' Increase Each day CONDID-19 Cases')
fig.show()
# -
from Custom.GlobalData import *
data=str(DATA[4]).split(",")
print(data[3])
# +
# %%writefile GlobalData.py
'''
Usage:
To get Zambia Population-2019
from GlobalData import *
X=GlobalData("Zambia")
x=X.split(",")
print(x[4])
To get Zambia Population Change
from GlobalData import *
X=GlobalData("Zambia")
x=X.split(",")
print(x[5])
from GlobalData import *
print(DATA[4])
>>> ['American Samoa,Oceania,Polynesia,55465,55312,−0.28%']
from GlobalData import *
data=str(DATA[4]).split(",")
print(data[3])
>>> 55465
'''
DATA=[['Country,UN Continental,Unstatistical,Population-2018,Population-2019,Population Change'],
['Afghanistan,Asia,Southern Asia,37171921,38041754,+2.34%'],
['Albania,Europe,Southern Europe,2882740,2880917,−0.06%'],
['Algeria,Africa,Northern Africa,42228408,43053054,+1.95%'],
['American Samoa,Oceania,Polynesia,55465,55312,−0.28%'],
['Andorra,Europe,Southern Europe,77006,77142,+0.18%'],
['Angola,Africa,Middle Africa,30809787,31825295,+3.30%'],
['Anguilla,Americas,Caribbean,14731,14869,+0.94%'],
['Antigua and Barbuda,Americas,Caribbean,96286,97118,+0.86%'],
['Argentina,Americas,South America,44361150,44780677,+0.95%'],
['Armenia,Asia,Western Asia,2951745,2957731,+0.20%'],
['Aruba,Americas,Caribbean,105845,106314,+0.44%'],
['Australia,Oceania,Australia and New Zealand,24898152,25203198,+1.23%'],
['Austria,Europe,Western Europe,8891388,8955102,+0.72%'],
['Azerbaijan,Asia,Western Asia,9949537,10047718,+0.99%'],
['Bahamas,Americas,Caribbean,385637,389482,+1.00%'],
['Bahrain,Asia,Western Asia,1569446,1641172,+4.57%'],
['Bangladesh,Asia,Southern Asia,161376708,163046161,+1.03%'],
['Barbados,Americas,Caribbean,286641,287025,+0.13%'],
['Belarus,Europe,Eastern Europe,9452617,9452411,0.00%'],
['Belgium,Europe,Western Europe,11482178,11539328,+0.50%'],
['Belize,Americas,Central America,383071,390353,+1.90%'],
['Benin,Africa,Western Africa,11485044,11801151,+2.75%'],
['Bermuda,Americas,Northern America,62756,62506,−0.40%'],
['Bhutan,Asia,Southern Asia,754388,763092,+1.15%'],
['Bolivia,Americas,South America,11353142,11513100,+1.41%'],
['Bosnia and Herzegovina,Europe,Southern Europe,3323925,3301000,−0.69%'],
['Botswana,Africa,Southern Africa,2254068,2303697,+2.20%'],
['Brazil,Americas,South America,209469323,211049527,+0.75%'],
['British Virgin Islands,Americas,Caribbean,29802,30030,+0.77%'],
['Brunei,Asia,South-eastern Asia,428963,433285,+1.01%'],
['Bulgaria,Europe,Eastern Europe,7051608,7000119,−0.73%'],
['Burkina Faso,Africa,Western Africa,19751466,20321378,+2.89%'],
['Burundi,Africa,Eastern Africa,10524117,10864245,+3.23%'],
['Cambodia,Asia,South-eastern Asia,16249792,16486542,+1.46%'],
['Cameroon,Africa,Middle Africa,25216267,25876380,+2.62%'],
['Canada,Americas,Northern America,37074562,37411047,+0.91%'],
['Cape Verde,Africa,Western Africa,543767,549935,+1.13%'],
['Caribbean Netherlands[t],Americas,Caribbean,25711,25979,+1.04%'],
['Cayman Islands,Americas,Caribbean,64174,64948,+1.21%'],
['Central African Republic,Africa,Middle Africa,4666368,4745185,+1.69%'],
['Chad,Africa,Middle Africa,15477729,15946876,+3.03%'],
['Chile,Americas,South America,18729160,18952038,+1.19%'],
['China[a],Asia,Eastern Asia,1427647786,1433783686,+0.43%'],
['Colombia,Americas,South America,49661048,50339443,+1.37%'],
['Comoros,Africa,Eastern Africa,832322,850886,+2.23%'],
['Congo,Africa,Middle Africa,5244359,5380508,+2.60%'],
['Cook Islands,Oceania,Polynesia,17518,17548,+0.17%'],
['Costa Rica,Americas,Central America,4999441,5047561,+0.96%'],
['Croatia,Europe,Southern Europe,4156405,4130304,−0.63%'],
['Cuba,Americas,Caribbean,11338134,11333483,−0.04%'],
['Curaçao,Americas,Caribbean,162752,163424,+0.41%'],
['Cyprus,Asia,Western Asia,1170125,1179551,+0.81%'],
['Czech Republic,Europe,Eastern Europe,10665677,10689209,+0.22%'],
['Denmark,Europe,Northern Europe,5752126,5771876,+0.34%'],
['Djibouti,Africa,Eastern Africa,958923,973560,+1.53%'],
['Dominica,Americas,Caribbean,71625,71808,+0.26%'],
['Dominican Republic,Americas,Caribbean,10627141,10738958,+1.05%'],
['DR Congo,Africa,Middle Africa,84068091,86790567,+3.24%'],
['East Timor,Asia,South-eastern Asia,1267974,1293119,+1.98%'],
['Ecuador,Americas,South America,17084358,17373662,+1.69%'],
['Egypt,Africa,Northern Africa,98423598,100388073,+2.00%'],
['El Salvador,Americas,Central America,6420746,6453553,+0.51%'],
['Equatorial Guinea,Africa,Middle Africa,1308975,1355986,+3.59%'],
['Eritrea,Africa,Eastern Africa,3452786,3497117,+1.28%'],
['Estonia,Europe,Northern Europe,1322920,1325648,+0.21%'],
['Eswatini,Africa,Southern Africa,1136281,1148130,+1.04%'],
['Ethiopia,Africa,Eastern Africa,109224414,112078730,+2.61%'],
['F.S. Micronesia,Oceania,Micronesia,112640,113815,+1.04%'],
['Falkland Islands,Americas,South America,3234,3377,+4.42%'],
['Faroe Islands,Europe,Northern Europe,48497,48678,+0.37%'],
['Fiji,Oceania,Melanesia,883483,889953,+0.73%'],
['Finland,Europe,Northern Europe,5522576,5532156,+0.17%'],
['France,Europe,Western Europe,64990511,65129728,+0.21%'],
['French Guiana,Americas,South America,275713,282731,+2.55%'],
['French Polynesia,Oceania,Polynesia,277679,279287,+0.58%'],
['Gabon,Africa,Middle Africa,2119275,2172579,+2.52%'],
['Gambia,Africa,Western Africa,2280094,2347706,+2.97%'],
['Georgia,Asia,Western Asia,4002942,3996765,−0.15%'],
['Germany,Europe,Western Europe,83124418,83517045,+0.47%'],
['Ghana,Africa,Western Africa,28206728,28833629,+2.22%'],
['Gibraltar,Europe,Southern Europe,33718,33701,−0.05%'],
['United Kingdom,Europe,Northern Europe,67141684,67530172,+0.58%'],
['Greece,Europe,Southern Europe,10522246,10473455,−0.46%'],
['Greenland,Americas,Northern America,56564,56672,+0.19%'],
['Grenada,Americas,Caribbean,111454,112003,+0.49%'],
['Guadeloupe[s],Americas,Caribbean,446928,447905,+0.22%'],
['Guam,Oceania,Micronesia,165768,167294,+0.92%'],
['Guatemala,Americas,Central America,17247849,17581472,+1.93%'],
['Guernsey and Jersey,Europe,Northern Europe,170499,172259,+1.03%'],
['Guinea,Africa,Western Africa,12414293,12771246,+2.88%'],
['Guinea-Bissau,Africa,Western Africa,1874303,1920922,+2.49%'],
['Guyana,Americas,South America,779006,782766,+0.48%'],
['Haiti,Americas,Caribbean,11123178,11263770,+1.26%'],
['Honduras,Americas,Central America,9587522,9746117,+1.65%'],
['Hong Kong,Asia,Eastern Asia,7371730,7436154,+0.87%'],
['Hungary,Europe,Eastern Europe,9707499,9684679,−0.24%'],
['Iceland,Europe,Northern Europe,336713,339031,+0.69%'],
['India,Asia,Southern Asia,1352642280,1366417754,+1.02%'],
['Indonesia,Asia,South-eastern Asia,267670543,270625568,+1.10%'],
['Iran,Asia,Southern Asia,81800188,82913906,+1.36%'],
['Iraq,Asia,Western Asia,38433600,39309783,+2.28%'],
['Ireland,Europe,Northern Europe,4818690,4882495,+1.32%'],
['Isle of Man,Europe,Northern Europe,84077,84584,+0.60%'],
['Israel,Asia,Western Asia,8381516,8519377,+1.64%'],
['Italy,Europe,Southern Europe,60627291,60550075,−0.13%'],
['Ivory Coast,Africa,Western Africa,25069230,25716544,+2.58%'],
['Jamaica,Americas,Caribbean,2934847,2948279,+0.46%'],
['Japan,Asia,Eastern Asia,127202192,126860301,−0.27%'],
['Jordan,Asia,Western Asia,9965318,10101694,+1.37%'],
['Kazakhstan,Asia,Central Asia,18319618,18551427,+1.27%'],
['Kenya,Africa,Eastern Africa,51392565,52573973,+2.30%'],
['Kiribati,Oceania,Micronesia,115847,117606,+1.52%'],
['Kuwait,Asia,Western Asia,4137312,4207083,+1.69%'],
['Kyrgyzstan,Asia,Central Asia,6304030,6415850,+1.77%'],
['Laos,Asia,South-eastern Asia,7061507,7169455,+1.53%'],
['Latvia,Europe,Northern Europe,1928459,1906743,−1.13%'],
['Lebanon,Asia,Western Asia,6859408,6855713,−0.05%'],
['Lesotho,Africa,Southern Africa,2108328,2125268,+0.80%'],
['Liberia,Africa,Western Africa,4818973,4937374,+2.46%'],
['Libya,Africa,Northern Africa,6678559,6777452,+1.48%'],
['Liechtenstein,Europe,Western Europe,37910,38019,+0.29%'],
['Lithuania,Europe,Northern Europe,2801264,2759627,−1.49%'],
['Luxembourg,Europe,Western Europe,604245,615729,+1.90%'],
['Macau,Asia,Eastern Asia,631636,640445,+1.39%'],
['Madagascar,Africa,Eastern Africa,26262313,26969307,+2.69%'],
['Malawi,Africa,Eastern Africa,18143217,18628747,+2.68%'],
['Malaysia,Asia,South-eastern Asia,31528033,31949777,+1.34%'],
['Maldives,Asia,Southern Asia,515696,530953,+2.96%'],
['Mali,Africa,Western Africa,19077749,19658031,+3.04%'],
['Malta,Europe,Southern Europe,439248,440372,+0.26%'],
['Marshall Islands,Oceania,Micronesia,58413,58791,+0.65%'],
['Martinique,Americas,Caribbean,375673,375554,−0.03%'],
['Mauritania,Africa,Western Africa,4403313,4525696,+2.78%'],
['Mauritius,Africa,Eastern Africa,1189265,1198575,+0.78%'],
['Mayotte,Africa,Eastern Africa,259531,266150,+2.55%'],
['Mexico,Americas,Central America,126190788,127575529,+1.10%'],
['Moldova,Europe,Eastern Europe,4051944,4043263,−0.21%'],
['Monaco,Europe,Western Europe,38682,38964,+0.73%'],
['Mongolia,Asia,Eastern Asia,3170216,3225167,+1.73%'],
['Montenegro,Europe,Southern Europe,627809,627987,+0.03%'],
['Montserrat,Americas,Caribbean,4993,4989,−0.08%'],
['Morocco,Africa,Northern Africa,36029093,36471769,+1.23%'],
['Mozambique,Africa,Eastern Africa,29496004,30366036,+2.95%'],
['Myanmar,Asia,South-eastern Asia,53708320,54045420,+0.63%'],
['Namibia,Africa,Southern Africa,2448301,2494530,+1.89%'],
['Nauru,Oceania,Micronesia,10670,10756,+0.81%'],
['Nepal,Asia,Southern Asia,28095714,28608710,+1.83%'],
['Netherlands,Europe,Western Europe,17059560,17097130,+0.22%'],
['New Caledonia,Oceania,Melanesia,279993,282750,+0.98%'],
['New Zealand,Oceania,Australia and New Zealand,4743131,4783063,+0.84%'],
['Nicaragua,Americas,Central America,6465501,6545502,+1.24%'],
['Niger,Africa,Western Africa,22442822,23310715,+3.87%'],
['Nigeria,Africa,Western Africa,195874683,200963599,+2.60%'],
['Niue,Oceania,Polynesia,1620,1615,−0.31%'],
['North Korea,Asia,Eastern Asia,25549604,25666161,+0.46%'],
['North Macedonia,Europe,Southern Europe,2082957,2083459,+0.02%'],
['Northern Mariana Islands,Oceania,Micronesia,56882,56188,−1.22%'],
['Norway,Europe,Northern Europe,5337962,5378857,+0.77%'],
['Oman,Asia,Western Asia,4829473,4974986,+3.01%'],
['Pakistan,Asia,Southern Asia,212228286,216565318,+2.04%'],
['Palau,Oceania,Micronesia,17907,18008,+0.56%'],
['Palestine[n],Asia,Western Asia,4862979,4981420,+2.44%'],
['Panama,Americas,Central America,4176869,4246439,+1.67%'],
['Papua New Guinea,Oceania,Melanesia,8606323,8776109,+1.97%'],
['Paraguay,Americas,South America,6956066,7044636,+1.27%'],
['Peru,Americas,South America,31989260,32510453,+1.63%'],
['Philippines,Asia,South-eastern Asia,106651394,108116615,+1.37%'],
['Poland,Europe,Eastern Europe,37921592,37887768,−0.09%'],
['Portugal,Europe,Southern Europe,10256193,10226187,−0.29%'],
['Puerto Rico,Americas,Caribbean,3039596,2933408,−3.49%'],
['Qatar,Asia,Western Asia,2781682,2832067,+1.81%'],
['Réunion,Africa,Eastern Africa,882526,888927,+0.73%'],
['Romania,Europe,Eastern Europe,19506114,19364557,−0.73%'],
['Russia,Europe,Eastern Europe,145734038,145872256,+0.09%'],
['Rwanda,Africa,Eastern Africa,12301970,12626950,+2.64%'],
['Saint Helena Ascension and Tristan da Cunha,Africa,Western Africa,6035,6059,+0.40%'],
['Saint Kitts and Nevis,Americas,Caribbean,52441,52823,+0.73%'],
['Saint Lucia,Americas,Caribbean,181889,182790,+0.50%'],
['Saint Pierre and Miquelon,Americas,Northern America,5849,5822,−0.46%'],
['Saint Vincent and the Grenadines,Americas,Caribbean,110211,110589,+0.34%'],
['Samoa,Oceania,Polynesia,196129,197097,+0.49%'],
['San Marino,Europe,Southern Europe,33785,33860,+0.22%'],
['São Tomé and Príncipe,Africa,Middle Africa,211028,215056,+1.91%'],
['Saudi Arabia,Asia,Western Asia,33702756,34268528,+1.68%'],
['Senegal,Africa,Western Africa,15854323,16296364,+2.79%'],
['Serbia[k],Europe,Southern Europe,8802754,8772235,−0.35%'],
['Seychelles,Africa,Eastern Africa,97096,97739,+0.66%'],
['Sierra Leone,Africa,Western Africa,7650150,7813215,+2.13%'],
['Singapore,Asia,South-eastern Asia,5757499,5804337,+0.81%'],
['Sint Maarten,Americas,Caribbean,41940,42388,+1.07%'],
['Slovakia,Europe,Eastern Europe,5453014,5457013,+0.07%'],
['Slovenia,Europe,Southern Europe,2077837,2078654,+0.04%'],
['Solomon Islands,Oceania,Melanesia,652857,669823,+2.60%'],
['Somalia,Africa,Eastern Africa,15008226,15442905,+2.90%'],
['South Africa,Africa,Southern Africa,57792518,58558270,+1.33%'],
['South Korea,Asia,Eastern Asia,51171706,51225308,+0.10%'],
['South Sudan,Africa,Eastern Africa,10975927,11062113,+0.79%'],
['Spain[d],Europe,Southern Europe,46692858,46736776,+0.09%'],
['Sri Lanka,Asia,Southern Asia,21228763,21323733,+0.45%'],
['Sudan,Africa,Northern Africa,41801533,42813238,+2.42%'],
['Suriname,Americas,South America,575990,581372,+0.93%'],
['Sweden,Europe,Northern Europe,9971638,10036379,+0.65%'],
['Switzerland,Europe,Western Europe,8525611,8591365,+0.77%'],
['Syria,Asia,Western Asia,16945057,17070135,+0.74%'],
['Taiwan,Asia,Eastern Asia,23726460,23773876,+0.20%'],
['Tajikistan,Asia,Central Asia,9100835,9321018,+2.42%'],
['Tanzania[c],Africa,Eastern Africa,56313438,58005463,+3.00%'],
['Thailand,Asia,South-eastern Asia,68863514,69037513,+0.25%'],
['Togo,Africa,Western Africa,7889093,8082366,+2.45%'],
['Tokelau,Oceania,Polynesia,1319,1340,+1.59%'],
['Tonga,Oceania,Polynesia,110589,110940,+0.32%'],
['Trinidad and Tobago,Americas,Caribbean,1389843,1394973,+0.37%'],
['Tunisia,Africa,Northern Africa,11565201,11694719,+1.12%'],
['Turkey,Asia,Western Asia,82340088,83429615,+1.32%'],
['Turkmenistan,Asia,Central Asia,5850901,5942089,+1.56%'],
['Turks and Caicos Islands,Americas,Caribbean,37665,38191,+1.40%'],
['Tuvalu,Oceania,Polynesia,11508,11646,+1.20%'],
['U.S. Virgin Islands,Americas,Caribbean,104680,104578,−0.10%'],
['Uganda,Africa,Eastern Africa,42729036,44269594,+3.61%'],
['Ukraine,Europe,Eastern Europe,44246156,43993638,−0.57%'],
['United Arab Emirates,Asia,Western Asia,9630959,9770529,+1.45%'],
['United States,Americas,Northern America,327096265,329064917,+0.60%'],
['US,Americas,Northern America,327096265,329064917,+0.60%'],
['Uruguay,Americas,South America,3449285,3461734,+0.36%'],
['Uzbekistan,Asia,Central Asia,32476244,32981716,+1.56%'],
['Vanuatu,Oceania,Melanesia,292680,299882,+2.46%'],
['Vatican City,Europe,Southern Europe,801,799,−0.25%'],
['Venezuela,Americas,South America,28887118,28515829,−1.29%'],
['Vietnam,Asia,South-eastern Asia,95545962,96462106,+0.96%'],
['Wallis and Futuna,Oceania,Polynesia,11661,11432,−1.96%'],
['Western Sahara,Africa,Northern Africa,567402,582463,+2.65%'],
['Yemen,Asia,Western Asia,28498683,29161922,+2.33%'],
['Zambia,Africa,Eastern Africa,17351708,17861030,+2.94%'],
['Zimbabwe,Africa,Eastern Africa,14438802,14645468,+1.43%']]
HELP="""help,Usage:
To get Zambia Population-2019
from GlobalData import *
X=GlobalData("Zambia")
x=X.split(",")
print(x[4])
\n
To get Zambia Population Change
from GlobalData import *
X=GlobalData("Zambia")
x=X.split(",")
print(x[5])
\n
from GlobalData import *
print(DATA[4])
>>> [\'American Samoa,Oceania,Polynesia,55465,55312,−0.28%\']
\n
from GlobalData import *
data=str(DATA[4]).split(",")
print(data[3])
>>> 55465']]"""
def GlobalData(country,HELP=HELP):
if country=="HELP":
print(HELP)
for i in range(0,len(DATA)):
if country in DATA[i][0]:
res=str(DATA[i])
return res
# -
from GlobalData import *
X=GlobalData("Zambia")
x=X.split(",")
print(x[4])
from GlobalData import *
GlobalData("Turkey")
GlobalData('HELP')
| .ipynb_checkpoints/021-plotly.graph-Deaths-by-Country-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Itinerary
#
# ## Intro
#
# * [Download larger files](00%20Get%20Training%20and%20Graph%20Data.ipynb)
# * [Quick slides: TensorFlow Introduction](https://docs.google.com/presentation/d/1ccO_czXmGSWKQJMWV63nsblFCNgGxyex2wxc9hwPiIM/edit?usp=sharing)
#
# ## Basics
#
# * [TensorFlow API Fundamentals](01%20TensorFlow%20Fundamentals.ipynb)
# * [Doing simple linear regression with TensorFlow](02%20Linear%20Regression.ipynb)
# * [Adding in TensorBoard](03%20Linear%20TensorBoard.ipynb)
# * [Feedforward neural network](04%20Feedforward%20Network.ipynb)
#
# ## Convolutional Neural Networks, Cats, and Dogs
#
# * [What are convolutions?](05%20Convolutional%20Networks.ipynb)
# * [Loading in pre-built models: Inception](06%20Inception%20Viewer.ipynb)
# * [Introduce concept of transfer learning using prebuilt Inception example](07%20Inception%20Retraining.ipynb)
# * [Loading in pre-trained variables "the real way" with AlexNet](08%20Create%20Base%20AlexNet.ipynb)
# * [Transfer learning by hand with AlexNet](09%20AlexNet%20Transfer%20Learning.ipynb)
# * [Feature extraction and Nearest Neighbor image retrieval with AlexNet](10%20AlexNet%20Image%20Retrieval.ipynb)
| Itinerary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:stable]
# language: python
# name: conda-env-stable-py
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Reset-notebook-here-to-reload-pandas" data-toc-modified-id="Reset-notebook-here-to-reload-pandas-1"><span class="toc-item-num">1 </span>Reset notebook here to reload pandas</a></div>
# -
# Smallest example:
# +
import pandas as pd
from pathlib import Path
def create_and_save():
df = pd.DataFrame({'a':[0,1], 'b':[1,2]})
p = Path(".") / 'test.csv'
df.to_hdf(p, 'df')
# + run_control={"frozen": false, "read_only": false}
# !conda install pandas=0.19 --yes
# -
create_and_save()
# !conda update pandas --yes
# # Reset notebook here to reload pandas
create_and_save()
| notebooks/pandas regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
"""
Single galaxy simulations: determine the shear dependence on PSF higher moment errors.
"""
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import scipy
import galsim
from IPython.display import clear_output
from astropy.io import fits
from matplotlib.colors import LogNorm
from numpy import mgrid, sum
import scipy.linalg as alg
import scipy.stats as stats
from galsim.zernike import Zernike
import matplotlib
# +
import sys
sys.path.append('../psfhome')
from homesm import *
from metasm import *
from moments import *
from HOMExShapeletPair import *
# +
SMALL_SIZE = 8
MEDIUM_SIZE = 10
BIGGER_SIZE = 14
plt.rc('font', size=BIGGER_SIZE) # controls default text sizes
plt.rc('axes', titlesize=BIGGER_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=BIGGER_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=BIGGER_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=BIGGER_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=MEDIUM_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
# -
def do_tests(tests,j,test_m, test_c,n):
testsresult=[]
for i in range(len(tests)):
test = HOMExShapeletPair(*tests[i][:-1],**tests[i][-1])
test.setup_shapelet_psf(test_m[i],test_c[i],n)
results = test.get_results(metacal = False)
testsresult.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(tests)*100)+"%")
return testsresult
def do_tests_speed(tests,j,test_m, test_c,n):
testsresult=[]
for i in range(len(tests)):
test = HOMExShapeletPair(*tests[i][:-1],**tests[i][-1])
if i!=0:
test.speed_setup_shapelet_psf(test_m[i],test_c[i],n,psf_light, psf_model_light, dm)
else:
test.setup_shapelet_psf(test_m[i],test_c[i],n)
psf_light = test.psf_light
psf_model_light = test.psf_model_light
dm = test.dm
results = test.get_results(metacal = False)
testsresult.append(results)
#clear_output()
#print ("Finished "+str(float((i+1))/len(tests)*100)+"%")
return testsresult
def e2(e1,e):
return np.sqrt(e**2 - e1**2)
test1 = HOMExShapeletPair("gaussian", 3.0, 0.2, 0.2, 0.01, 0.01, "gaussian", 2.0)
m = np.zeros(12)
c = np.zeros(12)
c[9]+=0.001
test1.setup_shapelet_psf(m,c,4)
pqlist = test1.sxm.get_pq_full(6)[3:]
# +
test2_init = [("gaussian" ,0.85 ,0.28,0.28,1e-8,1e-8,"gaussian" ,1.2,{'subtract_intersection':True}) for i in range(21)
]
test2_m = np.zeros(shape = (22,21,25))
test2_c = np.zeros(shape = (22,21,25))
for index in range(22):
for i in range(21):
test2_c[index][i][index+3]+=-0.01 + 0.001*i
# -
test2result = []
for i in range(len(test2_m)):
print( "Start tests for moment"+ str(i+4))
test2result.append(do_tests(test2_init,i,test2_m[i],test2_c[i],6))
#print test2result
# %store test2result
# +
pqlist = test1.sxm.get_pq_full(6)[3:]
fig = plt.figure(figsize = (21,12))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.5, hspace=0.5)
param1_dir = {}
param2_dir = {}
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
ax = plt.subplot(4,7,1+7*(n-3)+p)
dm = np.array([t['dm'][j+3] for t in test2result[j]])
dg1 = np.array([t["abs_bias"][0] for t in test2result[j]])
dg2 = np.array([t["abs_bias"][1] for t in test2result[j]])
params1= np.polyfit(dm,dg1,2)
params2= np.polyfit(dm,dg2,2)
# print params1
plt.plot(dm,dg1,label='g1')
plt.plot(dm,dg2,label='g2')
dg1_project = params1[2] + dm*params1[1] + dm**2*params1[0]
dg2_project = params2[2] + dm*params2[1] + dm**2*params2[0]
plt.plot(dm,dg1_project)
plt.plot(dm,dg2_project)
param1_dir[(p,q)] = params1
param2_dir[(p,q)] = params2
#print test4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
plt.ticklabel_format(axis='y',style='sci',scilimits=(0,3))
plt.xlabel(r"$\Delta m_{p,q}$")
plt.ylabel(r'$\Delta g_i$')
plt.title(str((p,q)))
#plt.show()
plt.legend()
#fig.colorbar(axes)
# +
import pickle
with open("../plots2/pickle/shear_response.pkl","wb") as f:
pickle.dump([pqlist,test2result],f)
# +
import pickle
with open("../plots2/pickle/shear_response.pkl","rb") as f:
pqlist,test2result = pickle.load(f)
# +
import pickle
f = open("../notebook/data/params1.pkl","wb")
pickle.dump(param1_dir,f)
f.close()
f = open("../notebook/data/params2.pkl","wb")
pickle.dump(param2_dir,f)
f.close()
# +
HSC_moment_bias = np.load('data/mean_residual.npy')
# +
#gal_size = 0.17 arcsec, psf_size = 0.24 arcsec, pixel_size = 0.2 arcsec
test3_init = [("gaussian" ,0.85 ,0.28,0.28,0.001+0.001*i,0.001+0.001*i,"gaussian" ,1.2 ,{'subtract_intersection':True}) for i in range(10)
]
# test3_init = [("gaussian" ,3.98 ,0.28,0.28,0.001+0.001*i,0.001+0.001*i,"gaussian" ,2.4 ,{'subtract_intersection':True}) for i in range(10)
# ]
test3_m = np.zeros(shape = (22,10,25))
test3_c = np.zeros(shape = (22,10,25))
for index in range(22):
for i in range(10):
test3_c[index][i][index+3]+=HSC_moment_bias[index+3]
#test3_c[index][i][index+3]+=0.005
# -
test3result = []
for i in range(len(test3_m)):
print( "Start tests for moment"+ str(i+4))
test3result.append(do_tests_speed(test3_init,i,test3_m[i],test3_c[i],6))
# %store test3result
# +
import pickle
with open("../plots2/pickle/add_and_mul.pkl","wb") as f:
pickle.dump([pqlist,test3result,test3_init],f)
# +
pqlist = test1.sxm.get_pq_full(6)[3:]
fig = plt.figure(figsize = (21,12))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.5, hspace=0.5)
g1_m = [];g1_c = [];g2_m = [];g2_c = []
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
ax = plt.subplot(4,7,1+7*(n-3)+p)
g1 = np.array([param[4] for param in test3_init])
g2 = np.array([param[5] for param in test3_init])
dg1 = np.array([t["abs_bias"][0] for t in test3result[j]])
dg2 = np.array([t["abs_bias"][1] for t in test3result[j]])
params1= np.polyfit(g1,dg1,1)
params2= np.polyfit(g2,dg2,1)
g1_m.append(params1[0]);g1_c.append(params1[1]);g2_m.append(params2[0]);g2_c.append(params2[1])
#print params1,params2
dg1_project = params1[1] + g1*params1[0]
dg2_project = params2[1] + g2*params2[0]
plt.plot(g1,dg1,'.',label='g1')
plt.plot(g2,dg2,'.',label='g2')
plt.plot(g1,dg1_project)
plt.plot(g2,dg2_project)
print(str((p,q)), (params1[0]/0.005, params2[0]/0.005), (params1[1]/0.005, params2[1]/0.005))
plt.ticklabel_format(axis='y',style='sci',scilimits=(0,3))
#print test4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
plt.xlabel(r"$g_1$")
plt.ylabel(r'${\Delta g_i}$')
plt.title(str((p,q)))
#plt.show()
plt.legend()
#fig.colorbar(axes)
# +
test4_init = [("gaussian" ,3.98 ,-0.2+i*0.04,e2(-0.2+i*0.04, 0.28),1e-8,1e-8,"gaussian" ,2.4,{'subtract_intersection':True}) for i in range(5)
]
test4_m = np.zeros(shape = (22,5,25))
test4_c = np.zeros(shape = (22,5,25))
for index in range(22):
for i in range(5):
test4_c[index][i][index+3]+=0.005
# -
test4result = []
for i in range(len(test4_m)):
print "Start tests for moment"+ str(i+4)
test4result.append(do_tests(test4_init,i,test4_m[i],test4_c[i],6))
# %store test4result
print test4result[1]
# +
pqlist = test1.sxm.get_pq_full(6)[3:]
fig = plt.figure(figsize = (21,12))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.5, hspace=0.5)
param1_dir = {}
param2_dir = {}
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
ax = plt.subplot(4,7,1+7*(n-3)+p)
e1 = np.array([t['e1'] for t in test4result[j]])
dg1 = np.array([t["abs_bias"][0] for t in test4result[j]])
dg2 = np.array([t["abs_bias"][1] for t in test4result[j]])
plt.plot(e1,dg1,label='g1')
plt.plot(e1,dg2,label='g2')
#print te bvst4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
plt.ticklabel_format(axis='y',style='sci',scilimits=(0,3))
plt.xlabel(r"$ e_1$")
plt.ylabel(r'$\Delta g_i$')
plt.title(str((p,q)))
#plt.show()
plt.legend()
#fig.colorbar(axes)
# -
# $g(x) = (1+m_1(x)+m_2(x)+\dots)g_{true}(x)$
#
# $<g(x) g(x+\theta)> = <((1+m_1(x)+m_2(x)+\dots)g_{true}(x))((1+m_1(x+\theta)+m_2(x+\theta)+\dots)g_{true}(x+\theta))>$
# $= <g_{true}(x)g_{true}(x+\theta)> + <(m_1(x)+m_2(x)+\dots)g_{true}(x) g_{true}(x+\theta)> + <(m_1(x+\theta)+m_2(x+\theta)+\dots)g_{true}(x)g_{true}(x+\theta)>$
# $= \xi_{true} + 2 \sum_i < m_i(x)> \xi_{true}$
# +
nob = 50
label_list = []
pqlist = test1.sxm.get_pq_full(6)
for i in range(nob):
if i < 25:
i_pre = 't'
else:
i_pre = 'r'
label1 = i_pre+str(pqlist[i%25][0])+str(pqlist[i%25][1])
label_list.append(label1)
fig, ax = plt.subplots(1,1,figsize=(8, 6))
ax.plot(np.arange(3,25),g1_m,'o',label = 'm1')
ax.plot(np.arange(3,25),g2_m,'o',label = 'm2')
ax.axvspan(7, 11, color='r', alpha=0.2, lw=0)
ax.axvspan(18, 24, color='r', alpha=0.2, lw=0)
ax.set_xticks(np.arange(3,25,1))
ax.set_xticklabels(label_list[28:], rotation='vertical', fontsize=14)
plt.grid()
plt.legend()
plt.ylabel("Multiplicative Bias")
print( "m1 = " + str(np.sum(g1_m)))
print( "m2 = " + str(np.sum(g2_m)))
# +
fig, ax = plt.subplots(1,1,figsize=(8, 6))
#ax.plot(np.arange(3,25),g1_c,'o',label = 'c1')
ax.plot(np.arange(3,25),g2_c,'o',label = 'c2')
ax.axvspan(7, 11, color='r', alpha=0.2, lw=0)
ax.axvspan(18, 24, color='r', alpha=0.2, lw=0)
ax.set_xticks(np.arange(3,25,1))
ax.set_xticklabels(label_list[28:], rotation='vertical', fontsize=14)
plt.grid()
plt.legend()
plt.ylabel("Additive Bias")
plt.yscale('symlog', linthresh = 0.00001)
print( "c1 = " + str(np.sum(g1_c)))
print( "c2 = " + str(np.sum(g2_c)))
# +
g = [param[4] for param in test3_init]
dg1 = np.array([t["abs_bias"][0] for t in test3result[4]])+2*np.array([t["abs_bias"][0] for t in test3result[6]])+np.array([t["abs_bias"][0] for t in test3result[8]])
dg2 = np.array([t["abs_bias"][1] for t in test3result[4]])+2*np.array([t["abs_bias"][1] for t in test3result[6]])+np.array([t["abs_bias"][1] for t in test3result[8]])
plt.plot(g,dg1)
plt.plot(g,dg2)
# +
#change coma1
#d(trefoil1) = du^3 - 3 d(uv^2) = 0
#d(coma1) = 0.04
test51_init = [("gaussian" ,3.0 ,0.28,0.28,0.001+0.001*i,0.001+0.001*i,"gaussian" ,2.0,{'subtract_intersection':True}) for i in range(10)
]
test51_m = np.zeros(shape = (10,12))
test51_c = np.zeros(shape = (10,12))
for i in range(test51_c.shape[0]):
test51_c[i][6]+=0.03
test51_c[i][4]+=0.01
test51result = []
for i in range(len(test51_m)):
test = HOMExShapeletPair(*test51_init[i][:-1],**test51_init[i][-1])
test.setup_shapelet_psf(test51_m[i],test51_c[i],4)
results = test.get_results()
test51result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test51_m)*100)+"%")
# +
#change coma2
#d(trefoil2) = 3d(u^2 v) - d(v^3) = 0
#d(coma1) = 0.04
test52_init = [("gaussian" ,3.0 ,0.28,0.28,0.001+0.001*i,0.001+0.001*i,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test52_m = np.zeros(shape = (10,12))
test52_c = np.zeros(shape = (10,12))
for i in range(test52_c.shape[0]):
test52_c[i][5]+=0.01
test52_c[i][3]+=0.03
test52result = []
for i in range(len(test52_m)):
test = HOMExShapeletPair(*test52_init[i][:-1],**test52_init[i][-1])
test.setup_shapelet_psf(test52_m[i],test52_c[i],4)
results = test.get_results()
test52result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test52_m)*100)+"%")
# +
#change trefoil1
#d(coma1) = du^3 + d(uv^2) = 0
#d(coma1) = 0.04
test53_init = [("gaussian" ,3.0 ,0.28,0.28,0.001+0.001*i,0.001+0.001*i,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test53_m = np.zeros(shape = (10,12))
test53_c = np.zeros(shape = (10,12))
for i in range(test53_c.shape[0]):
test53_c[i][6]+=0.01
test53_c[i][4]-=0.01
test53result = []
for i in range(len(test53_m)):
test = HOMExShapeletPair(*test53_init[i][:-1],**test53_init[i][-1])
test.setup_shapelet_psf(test53_m[i],test53_c[i],4)
results = test.get_results()
test53result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test53_m)*100)+"%")
# +
#change trefoil2
#d(coma2) = d(u^2 v) - d(v^3) = 0
#d(coma1) = 0.04
test54_init = [("gaussian" ,3.0 ,0.28,0.28,0.001+0.001*i,0.001+0.001*i,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test54_m = np.zeros(shape = (10,12))
test54_c = np.zeros(shape = (10,12))
for i in range(test54_c.shape[0]):
test54_c[i][5]+=0.01
test54_c[i][3]-=0.01
test54result = []
for i in range(len(test54_m)):
test = HOMExShapeletPair(*test54_init[i][:-1],**test54_init[i][-1])
test.setup_shapelet_psf(test54_m[i],test54_c[i],4)
results = test.get_results()
test54result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test54_m)*100)+"%")
# -
plt.plot([param[4] for param in test51_init],np.array([t["abs_bias"][0] for t in test51result]),label='coma1', color = 'blue')
plt.plot([param[5] for param in test51_init],np.array([t["abs_bias"][1] for t in test51result]),'-.', color = 'blue')
plt.plot([param[4] for param in test52_init],np.array([t["abs_bias"][0] for t in test52result]),label='coma2', color = 'orange')
plt.plot([param[5] for param in test52_init],np.array([t["abs_bias"][1] for t in test52result]),'-.', color = 'orange')
plt.plot([param[4] for param in test53_init],np.array([t["abs_bias"][0] for t in test53result]),label='trefoil1', color = 'green')
plt.plot([param[5] for param in test53_init],np.array([t["abs_bias"][1] for t in test53result]),'-.',color = 'green')
plt.plot([param[4] for param in test54_init],np.array([t["abs_bias"][0] for t in test54result]),label='trefoil2',color = 'purple')
plt.plot([param[5] for param in test54_init],np.array([t["abs_bias"][1] for t in test54result]),'-.',color = 'purple')
plt.xlabel(r'$g$')
plt.ylabel(r'$\Delta g$')
plt.title('g1 solid, g2 dashed')
plt.legend()
# +
test61_init = [("gaussian" ,4.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0+0.5*i,{'subtract_intersection':True}) for i in range(20)
]
test61_m = np.zeros(shape = (20,12))
test61_c = np.zeros(shape = (20,12))
for i in range(test61_c.shape[0]):
test61_c[i][6]+=0.03
test61_c[i][4]+=0.01
test61result = []
for i in range(len(test61_m)):
test = HOMExShapeletPair(*test61_init[i][:-1],**test61_init[i][-1])
test.setup_shapelet_psf(test61_m[i],test61_c[i],4)
results = test.get_results()
test61result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test61_m)*100)+"%")
# +
test62_init = [("gaussian" ,4.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0+0.5*i,{'subtract_intersection':True}) for i in range(20)
]
test62_m = np.zeros(shape = (20,12))
test62_c = np.zeros(shape = (20,12))
for i in range(test62_c.shape[0]):
test62_c[i][5]+=0.01
test62_c[i][3]+=0.03
test62result = []
for i in range(len(test62_m)):
test = HOMExShapeletPair(*test62_init[i][:-1],**test62_init[i][-1])
test.setup_shapelet_psf(test62_m[i],test62_c[i],4)
results = test.get_results()
test62result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test62_m)*100)+"%")
# +
test63_init = [("gaussian" ,4.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0+0.5*i,{'subtract_intersection':True}) for i in range(20)
]
test63_m = np.zeros(shape = (20,12))
test63_c = np.zeros(shape = (20,12))
for i in range(test63_c.shape[0]):
test63_c[i][6]+=0.01
test63_c[i][4]-=0.01
test63result = []
for i in range(len(test63_m)):
test = HOMExShapeletPair(*test63_init[i][:-1],**test63_init[i][-1])
test.setup_shapelet_psf(test63_m[i],test63_c[i],4)
results = test.get_results()
test63result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test63_m)*100)+"%")
# +
test64_init = [("gaussian" ,4.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0+0.5*i,{'subtract_intersection':True}) for i in range(20)
]
test64_m = np.zeros(shape = (20,12))
test64_c = np.zeros(shape = (20,12))
for i in range(test64_c.shape[0]):
test64_c[i][5]+=0.01
test64_c[i][3]-=0.01
test64result = []
for i in range(len(test64_m)):
test = HOMExShapeletPair(*test64_init[i][:-1],**test64_init[i][-1])
test.setup_shapelet_psf(test64_m[i],test64_c[i],4)
results = test.get_results()
test64result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test64_m)*100)+"%")
# +
plt.figure(figsize = (8,6))
plt.plot([t['gal_sigma']/t['psf_sigma'] for t in test61result],np.array([ t["abs_bias"][0]/0.02 for t in test61result]),label = 'coma1',color = 'blue')
plt.plot([t['gal_sigma']/t['psf_sigma'] for t in test61result],np.array([ t["abs_bias"][1]/0.02 for t in test61result]),'-.',color = 'blue')
plt.plot([t['gal_sigma']/t['psf_sigma'] for t in test62result],np.array([ t["abs_bias"][0]/0.02 for t in test62result]),label = 'coma2',color = 'orange')
plt.plot([t['gal_sigma']/t['psf_sigma'] for t in test62result],np.array([ t["abs_bias"][1]/0.02 for t in test62result]),'-.',color = 'orange')
plt.plot([t['gal_sigma']/t['psf_sigma'] for t in test63result],np.array([ t["abs_bias"][0]/0.02 for t in test63result]),label = 'trefoil1',color = 'green')
plt.plot([t['gal_sigma']/t['psf_sigma'] for t in test63result],np.array([ t["abs_bias"][1]/0.02 for t in test63result]),'-.',color = 'green')
plt.plot([t['gal_sigma']/t['psf_sigma'] for t in test64result],np.array([ t["abs_bias"][0]/0.02 for t in test64result]),label = 'trefoil2',color = 'purple')
plt.plot([t['gal_sigma']/t['psf_sigma'] for t in test64result],np.array([ t["abs_bias"][1]/0.02 for t in test64result]),'-.',color = 'purple')
plt.xlabel(r"$\sigma_{galaxy}/\sigma_{PSF}$")
plt.ylabel(r'$\frac{\delta g}{\delta_{moment}}$')
plt.title('Gaussian galaxy & Gaussian PSF')
plt.legend()
# +
test71_init = ("gaussian" ,4.0 ,0.28,0.28,0.001,0.001,"gaussian" ,4.0,{'subtract_intersection':True})
test71_m = np.zeros(shape = (1,12))
test71_c = np.zeros(shape = (1,12))
test71_c[0][6]+=0.03
test71_c[0][4]+=0.03
test = HOMExShapeletPair(*test71_init[:-1],**test71_init[-1])
test.setup_shapelet_psf(test71_m[0],test71_c[0],4)
truth = test.psf_light
model = test.psf_model_light
residual = model.drawImage(scale = 1.0, nx = 100, ny = 100).array - truth.drawImage(scale = 1.0, nx = 100, ny = 100).array
# +
fig,ax = plt.subplots(figsize = (3,3))
ax.set_xticks([])
ax.set_yticks([])
plt.title('coma 1 residual')
plt.imshow(residual, vmin = -0.001, vmax = 0.001)
#plt.colorbar()
plt.show()
fig,ax = plt.subplots(figsize = (3,3))
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(model.drawImage(scale = 1.0, nx = 100, ny = 100).array, cmap=plt.cm.BuPu)
plt.title('coma 1')
#plt.colorbar()
plt.show()
# +
test72_init = ("gaussian" ,4.0 ,0.28,0.28,0.001,0.001,"gaussian" ,4.0,{'subtract_intersection':True})
test72_m = np.zeros(shape = (1,12))
test72_c = np.zeros(shape = (1,12))
test72_c[0][3]+=0.03
test72_c[0][5]+=0.03
test = HOMExShapeletPair(*test72_init[:-1],**test72_init[-1])
test.setup_shapelet_psf(test72_m[0],test72_c[0],4)
truth = test.psf_light
model = test.psf_model_light
residual = model.drawImage(scale = 1.0, nx = 100, ny = 100).array - truth.drawImage(scale = 1.0, nx = 100, ny = 100).array
# +
fig,ax = plt.subplots(figsize = (3,3))
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(residual, vmin = -0.001, vmax = 0.001)
plt.title('coma 2 residual')
plt.show()
fig,ax = plt.subplots(figsize = (3,3))
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(model.drawImage(scale = 1.0, nx = 100, ny = 100).array, cmap=plt.cm.BuPu)
plt.title('coma 2')
#plt.colorbar()
plt.show()
# +
test73_init = ("gaussian" ,4.0 ,0.28,0.28,0.001,0.001,"gaussian" ,4.0,{'subtract_intersection':True})
test73_m = np.zeros(shape = (1,12))
test73_c = np.zeros(shape = (1,12))
test73_c[0][6]+=0.02
test73_c[0][4]-=0.06
test = HOMExShapeletPair(*test73_init[:-1],**test73_init[-1])
test.setup_shapelet_psf(test73_m[0],test73_c[0],4)
truth = test.psf_light
model = test.psf_model_light
residual = model.drawImage(scale = 1.0, nx = 100, ny = 100).array - truth.drawImage(scale = 1.0, nx = 100, ny = 100).array
# +
fig,ax = plt.subplots(figsize = (3,3))
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(residual, vmin = -0.001, vmax = 0.001)
plt.title('trefoil 1 residual')
plt.show()
fig,ax = plt.subplots(figsize = (3,3))
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(model.drawImage(scale = 1.0, nx = 100, ny = 100).array, cmap=plt.cm.BuPu)
plt.title('trefoil 2')
#plt.colorbar()
plt.show()
# +
test74_init = ("gaussian" ,4.0 ,0.28,0.28,0.001,0.001,"gaussian" ,4.0,{'subtract_intersection':True})
test74_m = np.zeros(shape = (1,12))
test74_c = np.zeros(shape = (1,12))
test74_c[0][6]-=0.02
test74_c[0][5]+=0.06
test = HOMExShapeletPair(*test74_init[:-1],**test74_init[-1])
test.setup_shapelet_psf(test74_m[0],test74_c[0],4)
truth = test.psf_light
model = test.psf_model_light
residual = model.drawImage(scale = 1.0, nx = 100, ny = 100).array - truth.drawImage(scale = 1.0, nx = 100, ny = 100).array
plt.imshow(residual)
plt.title('trefoil 2')
plt.colorbar()
plt.show()
plt.imshow(model.drawImage(scale = 1.0, nx = 100, ny = 100).array)
plt.title('trefoil 2')
#plt.colorbar()
plt.show()
# +
test81_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test81_m = np.zeros(shape = (10,12))
test81_c = np.zeros(shape = (10,12))
for i in range(test81_c.shape[0]):
test81_c[i][4]+=0.001*i
test81result = []
for i in range(len(test81_m)):
test = HOMExShapeletPair(*test81_init[i][:-1],**test81_init[i][-1])
test.setup_shapelet_psf(test81_m[i],test81_c[i],4)
results = test.get_results()
test81result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test81_m)*100)+"%")
# +
test82_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test82_m = np.zeros(shape = (10,12))
test82_c = np.zeros(shape = (10,12))
for i in range(test82_c.shape[0]):
test82_c[i][6]+=0.001*i
test82result = []
for i in range(len(test81_m)):
test = HOMExShapeletPair(*test82_init[i][:-1],**test82_init[i][-1])
test.setup_shapelet_psf(test82_m[i],test82_c[i],4)
results = test.get_results()
test82result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test82_m)*100)+"%")
# +
test83_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test83_m = np.zeros(shape = (10,12))
test83_c = np.zeros(shape = (10,12))
for i in range(test83_c.shape[0]):
test83_c[i][4]+=0.001*i
test83_c[i][6]+=0.001*i
test83result = []
for i in range(len(test83_m)):
test = HOMExShapeletPair(*test83_init[i][:-1],**test83_init[i][-1])
test.setup_shapelet_psf(test83_m[i],test83_c[i],4)
results = test.get_results()
test83result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test83_m)*100)+"%")
# +
dm1 = [t['dm'][4] for t in test81result]
dm2 = [t['dm'][6] for t in test82result]
dmtot = np.array(dm1)+np.array(dm2)
dshear1 = [t['abs_bias'][0] for t in test81result]
dshear2 = [t['abs_bias'][0] for t in test82result]
dsheartot = np.array(dshear1)+np.array(dshear2)
plt.plot(dmtot, dsheartot, label = 'dg('+r"$dm_{1,2}$"+') + dg('+r"$dm_{3,0}$"+')')
plt.plot(dmtot, [t['abs_bias'][0] for t in test83result],label = 'dg('+r"$dm_{1,2}$"+' + '+r"$dm_{3,0}$"+')')
plt.ylabel(r'$\Delta g_1$')
plt.xlabel(r'$\Delta dm_{1,2} + dm_{3,0} $')
plt.legend()
# +
test91_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test91_m = np.zeros(shape = (10,12))
test91_c = np.zeros(shape = (10,12))
for i in range(test91_c.shape[0]):
test91_c[i][7]+=0.001*i
test91result = []
for i in range(len(test91_m)):
test = HOMExShapeletPair(*test91_init[i][:-1],**test91_init[i][-1])
test.setup_shapelet_psf(test91_m[i],test91_c[i],4)
results = test.get_results()
test91result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test91_m)*100)+"%")
# +
test92_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test92_m = np.zeros(shape = (10,12))
test92_c = np.zeros(shape = (10,12))
for i in range(test92_c.shape[0]):
test92_c[i][8]+=0.001*i
test92result = []
for i in range(len(test92_m)):
test = HOMExShapeletPair(*test92_init[i][:-1],**test92_init[i][-1])
test.setup_shapelet_psf(test92_m[i],test92_c[i],4)
results = test.get_results()
test92result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test92_m)*100)+"%")
# +
test93_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test93_m = np.zeros(shape = (10,12))
test93_c = np.zeros(shape = (10,12))
for i in range(test93_c.shape[0]):
test93_c[i][7]+=0.001*i
test93_c[i][8]+=0.001*i
test93result = []
for i in range(len(test93_m)):
test = HOMExShapeletPair(*test93_init[i][:-1],**test93_init[i][-1])
test.setup_shapelet_psf(test93_m[i],test93_c[i],4)
results = test.get_results()
test93result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test93_m)*100)+"%")
# +
dm1 = [t['dm'][7] for t in test91result]
dm2 = [t['dm'][8] for t in test92result]
dmtot = np.array(dm1)+np.array(dm2)
dshear1 = [t['abs_bias'][0] for t in test91result]
dshear2 = [t['abs_bias'][0] for t in test92result]
dsheartot = np.array(dshear1)+np.array(dshear2)
plt.plot(dmtot, dsheartot, label = 'dg('+r"$dm_{4,0}$"+') + dg('+r"$dm_{3,1}$"+')')
plt.plot(dmtot, [t['abs_bias'][0] for t in test93result],label = 'dg('+r"$dm_{4,0}$"+' + '+r"$dm_{3,1}$"+')')
plt.ylabel(r'$\Delta g_1$')
plt.xlabel(r'$\Delta dm_{4,0} + dm_{3,1} $')
plt.legend()
# +
test101_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test101_m = np.zeros(shape = (10,12))
test101_c = np.zeros(shape = (10,12))
for i in range(test101_c.shape[0]):
test101_c[i][3]+=0.003*i
test101result = []
for i in range(len(test101_m)):
test = HOMExShapeletPair(*test101_init[i][:-1],**test101_init[i][-1])
test.setup_shapelet_psf(test101_m[i],test101_c[i],4)
results = test.get_results()
test101result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test101_m)*100)+"%")
# +
test102_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test102_m = np.zeros(shape = (10,12))
test102_c = np.zeros(shape = (10,12))
for i in range(test102_c.shape[0]):
test102_c[i][8]+=0.001*i
test102result = []
for i in range(len(test102_m)):
test = HOMExShapeletPair(*test102_init[i][:-1],**test102_init[i][-1])
test.setup_shapelet_psf(test102_m[i],test102_c[i],4)
results = test.get_results()
test102result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test102_m)*100)+"%")
# +
test103_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test103_m = np.zeros(shape = (10,12))
test103_c = np.zeros(shape = (10,12))
for i in range(test103_c.shape[0]):
test103_c[i][8]+=0.001*i
test101_c[i][3]+=0.003*i
test103result = []
for i in range(len(test103_m)):
test = HOMExShapeletPair(*test103_init[i][:-1],**test103_init[i][-1])
test.setup_shapelet_psf(test103_m[i],test103_c[i],4)
results = test.get_results()
test103result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test103_m)*100)+"%")
# +
dm1 = [t['dm'][3] for t in test101result]
dm2 = [t['dm'][8] for t in test102result]
dmtot = np.array(dm1)+np.array(dm2)
dshear1 = [t['abs_bias'][0] for t in test101result]
dshear2 = [t['abs_bias'][0] for t in test102result]
dsheartot = np.array(dshear1)+np.array(dshear2)
plt.plot(dmtot, dsheartot, label = 'dg('+r"$dm_{3,1}$"+') + dg('+r"$dm_{3,0}$"+')')
plt.plot(dmtot, [t['abs_bias'][0] for t in test103result],label = 'dg('+r"$dm_{3,1}$"+' + '+r"$dm_{3,0}$"+')')
plt.ylabel(r'$\Delta g_1$')
plt.xlabel(r'$\Delta dm_{1,2} + dm_{3,0} $')
plt.legend()
# +
test111_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test111_m = np.zeros(shape = (10,25))
test111_c = np.zeros(shape = (10,25))
for i in range(test111_c.shape[0]):
test111_c[i][13]+=0.001*i
test111result = []
for i in range(len(test111_m)):
test = HOMExShapeletPair(*test111_init[i][:-1],**test111_init[i][-1])
test.setup_shapelet_psf(test111_m[i],test111_c[i],6)
results = test.get_results()
test111result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test111_m)*100)+"%")
# +
test112_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test112_m = np.zeros(shape = (10,25))
test112_c = np.zeros(shape = (10,25))
for i in range(test112_c.shape[0]):
test112_c[i][19]+=0.001*i
test112result = []
for i in range(len(test112_m)):
test = HOMExShapeletPair(*test112_init[i][:-1],**test112_init[i][-1])
test.setup_shapelet_psf(test112_m[i],test112_c[i],6)
results = test.get_results()
test112result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test112_m)*100)+"%")
# +
test113_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test113_m = np.zeros(shape = (10,25))
test113_c = np.zeros(shape = (10,25))
for i in range(test113_c.shape[0]):
test113_c[i][19]+=0.001*i
test113_c[i][13]+=0.001*i
test113result = []
for i in range(len(test113_m)):
test = HOMExShapeletPair(*test113_init[i][:-1],**test113_init[i][-1])
test.setup_shapelet_psf(test113_m[i],test113_c[i],6)
results = test.get_results()
test113result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test113_m)*100)+"%")
# +
dm1 = [t['dm'][13] for t in test111result]
dm2 = [t['dm'][19] for t in test112result]
dmtot = np.array(dm1)+np.array(dm2)
dshear1 = [t['abs_bias'][0] for t in test111result]
dshear2 = [t['abs_bias'][0] for t in test112result]
dsheartot = np.array(dshear1)+np.array(dshear2)
plt.plot(dm1, dshear1)
plt.xlabel(r'$\Delta dm_{4,1} $')
plt.ylabel(r'$\Delta g_1$')
plt.show()
plt.plot(dm2, dshear2)
plt.xlabel(r'$\Delta dm_{5,1} $')
plt.ylabel(r'$\Delta g_1$')
plt.show()
plt.plot(dmtot, dsheartot, label = 'dg('+r"$dm_{4,1}$"+') + dg('+r"$dm_{5,1}$"+')')
plt.plot(dmtot, [t['abs_bias'][0] for t in test113result],label = 'dg('+r"$dm_{4,1}$"+' + '+r"$dm_{5,1}$"+')')
plt.ylabel(r'$\Delta g_1$')
plt.xlabel(r'$\Delta dm_{4,1} + dm_{5,1} $')
plt.legend()
plt.show()
# +
test121_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test121_m = np.zeros(shape = (10,25))
test121_c = np.zeros(shape = (10,25))
for i in range(test121_c.shape[0]):
test111_c[i][3]+=0.001*i
test121result = []
for i in range(len(test121_m)):
test = HOMExShapeletPair(*test121_init[i][:-1],**test121_init[i][-1])
test.setup_shapelet_psf(test121_m[i],test121_c[i],6)
results = test.get_results()
test121result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test121_m)*100)+"%")
# +
test123_init = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(10)
]
test123_m = np.zeros(shape = (10,25))
test123_c = np.zeros(shape = (10,25))
for i in range(test113_c.shape[0]):
test123_c[i][19]+=0.001*i
test123_c[i][3]+=0.001*i
test123result = []
for i in range(len(test123_m)):
test = HOMExShapeletPair(*test123_init[i][:-1],**test123_init[i][-1])
test.setup_shapelet_psf(test123_m[i],test123_c[i],6)
results = test.get_results()
test123result.append(results)
clear_output()
print ("Finished "+str(float((i+1))/len(test123_m)*100)+"%")
# +
dm1 = [t['dm'][3] for t in test121result]
dm2 = [t['dm'][19] for t in test112result]
dmtot = np.array(dm1)+np.array(dm2)
dshear1 = [t['abs_bias'][0] for t in test121result]
dshear2 = [t['abs_bias'][0] for t in test112result]
dsheartot = np.array(dshear1)+np.array(dshear2)
plt.plot(dm1, dshear1)
plt.xlabel(r'$\Delta dm_{3,0} $')
plt.ylabel(r'$\Delta g_1$')
plt.show()
plt.plot(dm2, dshear2)
plt.xlabel(r'$\Delta dm_{5,1} $')
plt.ylabel(r'$\Delta g_1$')
plt.show()
plt.plot(dmtot, dsheartot, label = 'dg('+r"$dm_{4,1}$"+') + dg('+r"$dm_{5,1}$"+')')
plt.plot(dmtot, [t['abs_bias'][0] for t in test113result],label = 'dg('+r"$dm_{4,1}$"+' + '+r"$dm_{5,1}$"+')')
plt.ylabel(r'$\Delta g_1$')
plt.xlabel(r'$\Delta dm_{4,1} + dm_{5,1} $')
plt.legend()
plt.show()
# +
test13_init = [("gaussian" ,0.5+0.1*i ,0.1,e2(0.1,0.28),1e-8,1e-8,"gaussian" , 1.5 ,{'subtract_intersection':True}) for i in range(40)
]
test13_m = np.zeros(shape = (22,40,25))
test13_c = np.zeros(shape = (22,40,25))
for index in range(22):
for i in range(40):
test13_c[index][i][index+3]+=0.005
# -
test13result = []
for i in range(len(test13_m)):
print( "Start tests for moment"+ str(i+4))
test13result.append(do_tests_speed(test13_init,i,test13_m[i],test13_c[i],6))
from scipy import interpolate
# +
import pickle
with open("../plots2/pickle/add_size_ratio.pkl","wb") as f:
pickle.dump([pqlist,test13result,test13_init, test131result, test131_init],f)
# +
spine_list1 = []
spine_list2 = []
pq4nersc = []
pqlist = test1.sxm.get_pq_full(6)[3:]
fig = plt.figure(figsize = (21,12))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.5, hspace=0.5)
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
ax = plt.subplot(4,7,1+7*(n-3)+p)
size_ratio = np.array([t['gal_sigma']/t['psf_sigma'] for t in test13result[j]])
dg1 = np.array([t["abs_bias"][0] for t in test13result[j]])/0.005
dg2 = np.array([t["abs_bias"][1] for t in test13result[j]])/0.005
plt.plot(size_ratio,dg1,label='g1')
plt.plot(size_ratio,dg2,label='g2')
#print test4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
plt.ticklabel_format(axis='y',style='sci',scilimits=(0,3))
spine_list1.append(dg1)
spine_list2.append(dg2)
pq4nersc.append([p,q])
plt.xlabel(r"$\sigma_{galaxy}/\sigma_{PSF}$")
plt.ylabel(r'$\delta g_i / \delta m_{p,q}$')
plt.title(str((p,q)))
#plt.show()
plt.legend()
#fig.colorbar(axes)
# -
np.save('Results/size_ratio.npy',size_ratio)
np.save('Results/dg1.npy',np.array(spine_list1))
np.save('Results/dg2.npy',np.array(spine_list2))
np.save('Results/pq4nersc.npy', np.array(pq4nersc))
# +
test131_init = [("sersic" ,0.5+0.1*i ,0.1,e2(0.1,0.28),1e-8,1e-8,"gaussian" ,1.5 ,{'subtract_intersection':True,'sersicn':3.0}) for i in range(40)
]
test131_m = np.zeros(shape = (22,40,25))
test131_c = np.zeros(shape = (22,40,25))
for index in range(22):
for i in range(40):
test131_c[index][i][index+3]+=0.005
# -
test131result = []
for i in range(len(test131_m)):
print( "Start tests for moment"+ str(i+4))
test131result.append(do_tests_speed(test131_init,i,test131_m[i],test131_c[i],6))
# +
test132_init = [("sersic" ,1.0+0.2*i ,0.1,e2(0.1,0.28),1e-8,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':0.5}) for i in range(40)
]
test132_m = np.zeros(shape = (22,40,25))
test132_c = np.zeros(shape = (22,40,25))
for index in range(22):
for i in range(40):
test132_c[index][i][index+3]+=0.005
# -
test132result = []
for i in range(len(test132_m)):
print "Start tests for moment"+ str(i+4)
test132result.append(do_tests_speed(test132_init,i,test132_m[i],test132_c[i],6))
# %store test13result
# %store test131result
# %store test132result
# %store -r test13result
# %store -r test131result
# %store -r test132result
y_range = {}
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
if n not in y_range.keys():
y_range[n] = [0,0]
#print min(min(np.array([t["abs_bias"][0] for t in test13result[j]])/0.005),y_range[n][0])
y_range[n][0] = min(min(np.array([t["abs_bias"][0] for t in test13result[j]])/0.005)*1.1,y_range[n][0])
y_range[n][0] = min(min(np.array([t["abs_bias"][1] for t in test13result[j]])/0.005)*1.1,y_range[n][0])
y_range[n][0] = min(min(np.array([t["abs_bias"][0] for t in test131result[j]])/0.005)*1.1,y_range[n][0])
y_range[n][0] = min(min(np.array([t["abs_bias"][1] for t in test131result[j]])/0.005)*1.1,y_range[n][0])
y_range[n][1] = max(max(np.array([t["abs_bias"][0] for t in test13result[j]])/0.005)*1.1,y_range[n][1])
y_range[n][1] = max(max(np.array([t["abs_bias"][1] for t in test13result[j]])/0.005)*1.1,y_range[n][1])
y_range[n][1] = max(max(np.array([t["abs_bias"][0] for t in test131result[j]])/0.005)*1.1,y_range[n][1])
y_range[n][1] = max(max(np.array([t["abs_bias"][1] for t in test131result[j]])/0.005)*1.1,y_range[n][1])
print y_range
# +
pqlist = test1.sxm.get_pq_full(6)[3:]
fig = plt.figure(figsize = (21,12))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.0, hspace=0.0)
# f, axes = plt.subplots(4, 7, sharex='col', sharey='row', figsize=(21,12))
# f.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.0, hspace=0.0)
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
#print n
ax = plt.subplot(4,7,1+7*(n-3)+p)
plt.plot(np.array([t['gal_hlr']/t['psf_hlr'] for t in test13result[j]]),np.array([t["abs_bias"][0] for t in test13result[j]])/0.005,color = 'blue')
plt.plot(np.array([t['gal_hlr']/t['psf_hlr'] for t in test13result[j]]),np.array([t["abs_bias"][1] for t in test13result[j]])/0.005,color = 'orange')
plt.plot(np.array([t['gal_hlr']/t['psf_hlr'] for t in test131result[j]]),np.array([t["abs_bias"][0] for t in test131result[j]])/0.005,'--',color = 'blue')
plt.plot(np.array([t['gal_hlr']/t['psf_hlr'] for t in test131result[j]]),np.array([t["abs_bias"][1] for t in test131result[j]])/0.005,'--',color = 'orange')
# plt.plot(np.array([t['gal_hlr']/t['psf_hlr'] for t in test132result[j]]),np.array([t["abs_bias"][0] for t in test131result[j]])/0.005,'.-',color = 'blue')
# plt.plot(np.array([t['gal_hlr']/t['psf_hlr'] for t in test132result[j]]),np.array([t["abs_bias"][1] for t in test131result[j]])/0.005,'.-',color = 'orange')
#print test4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
ax.tick_params(
axis='x', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
bottom=True, # ticks along the bottom edge are off
top=True, # ticks along the top edge are off
labelbottom=False)
ax.tick_params(
axis='y', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
left=True, # ticks along the bottom edge are off
right=False, # ticks along the top edge are off
labelleft=False)
#ax.tick_params(axis="y",direction="in")
if j in list(range(15,22)):
plt.xlabel(r"$R_h^{galaxy}/R_h^{PSF}$")
ax.tick_params(
axis='x', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
bottom=True, # ticks along the bottom edge are off
top=True, # ticks along the top edge are off
labelbottom=True)
if j in [0,4,9,15]:
plt.ylabel(r'$\delta g_i / \delta m_{p,q}$')
plt.ticklabel_format(axis='y',style='scientific',scilimits=(0,3))
ax.tick_params(
axis='y', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
left=True, # ticks along the bottom edge are off
right=False, # ticks along the top edge are off
labelleft=True)
plt.ylim(y_range[n])
plt.title(str((p,q)),y = 0.8)
#plt.show()
#plt.legend([])
plt.subplot(4,7,7,frame_on = False)
plt.plot([0],[0],color = 'blue',label = r'Gaussian $g_1$')
plt.plot([0],[0],color = 'orange',label = r'Gaussian $g_2$')
plt.plot([0],[0],'--',color = 'blue',label = r'Sersic n = 3.0 $g_1$')
plt.plot([0],[0],'--',color = 'orange',label = r'Sersic n = 3.0 $g_2$')
plt.axis('off')
plt.legend(fontsize = 'large',frameon = False)
#fig.colorbar(axes)
# -
print np.array(pq4nersc)
# +
def linearity_check(m1, dm1, m2, dm2, config, n_max = 6):
vector_length = (n_max +1 + 3) * (n_max - 1) / 2
test1_m = np.zeros(shape = (1,vector_length))
test1_c = np.zeros(shape = (1,vector_length))
test1_c[0][m1]+=dm1
test1 = HOMExShapeletPair(*config[0][:-1],**config[0][-1])
test1.setup_shapelet_psf(test1_m[0],test1_c[0],n_max)
results1 = test1.get_results()
test2_m = np.zeros(shape = (1,vector_length))
test2_c = np.zeros(shape = (1,vector_length))
test2_c[0][m2]+=dm2
test2 = HOMExShapeletPair(*config[0][:-1],**config[0][-1])
test2.setup_shapelet_psf(test2_m[0],test2_c[0],n_max)
results2 = test2.get_results()
test3_m = np.zeros(shape = (1,vector_length))
test3_c = np.zeros(shape = (1,vector_length))
test3_c[0][m1]+=dm1
test3_c[0][m2]+=dm2
test3 = HOMExShapeletPair(*config[0][:-1],**config[0][-1])
test3.setup_shapelet_psf(test3_m[0],test3_c[0],n_max)
results3 = test3.get_results()
dshear1 = results1['abs_bias'][0]
dshear2 = results2['abs_bias'][0]
#print dshear1, dshear2
linear_results = dshear1 + dshear2
auto_results = results3['abs_bias'][0]
#print results3['actual_dm']
#print linear_results, auto_results
error_over_minor = abs(linear_results - auto_results) / min(np.abs(dshear1) , np.abs(dshear2) )
error_over_sum = abs(linear_results - auto_results) / (np.abs(dshear1) + np.abs(dshear2))
return error_over_minor, error_over_sum
# +
config = [("gaussian" ,3.0 ,0.28,0.28,0.001,0.001,"gaussian" ,2.0 ,{'subtract_intersection':True}) for i in range(1)]
error_over_minor_matrix = np.zeros(shape = (12,12))
error_over_sum_matrix = np.zeros(shape = (12,12))
for i in range(12):
for j in range(i,12):
print i,j
eom, eos = linearity_check(i,0.001,j,0.001,config,4)
error_over_minor_matrix[i][j] = eom
error_over_sum_matrix[i][j] = eos
# +
n_max = 4
dg_scale = []
for i in range(12):
print i
vector_length = (n_max +1 + 3) * (n_max - 1) / 2
test1_m = np.zeros(shape = (1,vector_length))
test1_c = np.zeros(shape = (1,vector_length))
test1_c[0][i]+=0.001
test1 = HOMExShapeletPair(*config[0][:-1],**config[0][-1])
test1.setup_shapelet_psf(test1_m[0],test1_c[0],n_max)
results1 = test1.get_results()
dg_scale.append(np.abs(results1['abs_bias'][0]))
# +
pqlist = test1.sxm.get_pq_full(4)
label_list = []
for i in range(12):
label_list.append("m"+str(pqlist[i][0])+str(pqlist[i][1]))
fig, ax = plt.subplots(1,1,figsize=(8, 8))
mappable = ax.imshow(error_over_minor_matrix, cmap = 'Blues',vmin = -0.0, vmax = 0.5)
# Set number of ticks for x-axis
# Set ticks labels for x-axis
ax.set_xticks(np.arange(0,12,1))
ax.set_yticks(np.arange(0,12,1))
ax.set_xticklabels(label_list, rotation='vertical', fontsize=14)
ax.set_yticklabels(label_list, rotation='horizontal', fontsize=14)
plt.colorbar(mappable, ax = ax, label = r"$ \frac{dg(dm_1) + dg_2(dm_2) - dg(dm_1+dm_2)}{min(dg(dm_1), dg(dm_2))}$")
plt.title(r"$ \frac{dg(dm_1) + dg_2(dm_2) - dg(dm_1+dm_2)}{min(dg(dm_1), dg(dm_2))}$")
# +
pqlist = test1.sxm.get_pq_full(4)
label_list = []
for i in range(12):
label_list.append("m"+str(pqlist[i][0])+str(pqlist[i][1]))
fig, ax = plt.subplots(1,1,figsize=(8, 8))
mappable = ax.imshow(error_over_sum_matrix, cmap = 'Blues',vmin = -0.0, vmax = 1.0)
# Set number of ticks for x-axis
# Set ticks labels for x-axis
ax.set_xticks(np.arange(0,12,1))
ax.set_yticks(np.arange(0,12,1))
ax.set_xticklabels(label_list, rotation='vertical', fontsize=14)
ax.set_yticklabels(label_list, rotation='horizontal', fontsize=14)
plt.colorbar(mappable, ax = ax, label = r"$ \frac{dg(dm_1) + dg_2(dm_2) - dg(dm_1+dm_2)}{dg(dm_1) + dg(dm_2)}$")
plt.title(r"$ \frac{dg(dm_1) + dg_2(dm_2) - dg(dm_1+dm_2)}{dg(dm_1) + dg(dm_2)}$")
plt.show()
fig, ax = plt.subplots(1,1,figsize=(6, 4))
mappable = plt.plot(np.arange(0,12), dg_scale,'+')
plt.yscale('log')
ax.axvspan(3, 6, color='r', alpha=0.2, lw=0)
ax.set_xticks(np.arange(0,12,1))
ax.set_xticklabels(label_list, rotation='vertical', fontsize=14)
plt.ylabel('dg1')
plt.show()
# -
print pqlist
pq_for_m = [4,6,8,15,17,19,21]
# +
test14_init = [("gaussian" ,1.0+0.2*i ,0.1,0.26,1e-8,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True}) for i in range(40)
]
test14_m = np.zeros(shape = (7,40,25))
test14_c = np.zeros(shape = (7,40,25))
for index in range(7):
for i in range(40):
test14_c[index][i][pq_for_m[index]+3]+=0.005
# -
test14result = []
for i in range(len(test14_m)):
print "Start tests for moment"+ str(pq_for_m[i]+4)
test14result.append(do_tests(test14_init,i,test14_m[i],test14_c[i],6))
# +
test141_init = [("gaussian" ,1.0+0.2*i ,0.1,0.26,0.01,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True}) for i in range(40)
]
test141_m = np.zeros(shape = (7,40,25))
test141_c = np.zeros(shape = (7,40,25))
for index in range(7):
for i in range(40):
test141_c[index][i][pq_for_m[index]+3]+=0.005
# -
test141result = []
for i in range(len(test141_m)):
print "Start tests for moment"+ str(pq_for_m[i]+4)
test141result.append(do_tests(test141_init,i,test141_m[i],test141_c[i],6))
# +
test142_init = [("gaussian" ,1.0+0.2*i ,0.1,0.26,1e-8,0.01,"gaussian" ,3.0 ,{'subtract_intersection':True}) for i in range(40)
]
test142_m = np.zeros(shape = (7,40,25))
test142_c = np.zeros(shape = (7,40,25))
for index in range(7):
for i in range(40):
test142_c[index][i][pq_for_m[index]+3]+=0.005
# -
test142result = []
for i in range(len(test142_m)):
print "Start tests for moment"+ str(pq_for_m[i]+4)
test142result.append(do_tests(test142_init,i,test142_m[i],test142_c[i],6))
print test14result[0][0]
# +
size_ratio = np.zeros(shape = (40))
m1_size = np.zeros(shape = (7,40))
m2_size = np.zeros(shape = (7,40))
for i in range(40):
size_ratio[i] = test14result[0][i]['gal_sigma']/test14result[0][i]['psf_sigma']
for j in range(7):
m1_size[j][i] = (test141result[j][i]['abs_bias'][0] - test14result[j][i]['abs_bias'][0])/0.01/0.005
m2_size[j][i] = (test142result[j][i]['abs_bias'][1] - test14result[j][i]['abs_bias'][1])/0.01/0.005
# -
print m1_size.shape
np.save('data/multiplicative_size_ratio',size_ratio)
np.save('data/m1_size_ratio',m1_size)
np.save('data/m2_size_ratio',m2_size)
# +
fig = plt.figure(figsize = (21,4))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.5, hspace=0.5)
for j in range(7):
p,q = pqlist[pq_for_m[j]][0],pqlist[pq_for_m[j]][1]
n = p+q
ax = plt.subplot(1,7,j+1)
m1 = m1_size[j]
m2 = m2_size[j]
plt.plot(size_ratio,m1,label='g1')
plt.plot(size_ratio,m2,label='g2')
#print test4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
plt.ticklabel_format(axis='y',style='sci',scilimits=(0,3))
plt.xlabel(r"$\sigma_{galaxy}/\sigma_{PSF}$")
plt.ylabel(r'$ m / B[ \mathbf{m}_{p,q}]$')
plt.title(str((p,q)))
#plt.show()
plt.legend()
#fig.colorbar(axes)
# +
test15_init = [("sersic" ,1.0+0.2*i ,0.1,0.26,1e-8,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':0.5}) for i in range(40)
]+[("sersic" ,1.0+0.2*i ,0.1,0.26,0.01,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':0.5}) for i in range(40)
]+[("sersic" ,1.0+0.2*i ,0.1,0.26,1e-8,0.01,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':0.5}) for i in range(40)
]
test15_m = np.zeros(shape = (7,120,25))
test15_c = np.zeros(shape = (7,120,25))
for index in range(7):
for i in range(120):
test15_c[index][i][pq_for_m[index]+3]+=0.005
# -
test15result = []
for i in range(len(test15_m)):
print "Start tests for moment"+ str(pq_for_m[i]+4)
test15result.append(do_tests(test15_init,i,test15_m[i],test15_c[i],6))
# +
test151_init = [("sersic" ,1.0+0.2*i ,0.1,0.26,1e-8,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':3.0}) for i in range(40)
]+[("sersic" ,1.0+0.2*i ,0.1,0.26,0.01,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':3.0}) for i in range(40)
]+[("sersic" ,1.0+0.2*i ,0.1,0.26,1e-8,0.01,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':3.0}) for i in range(40)
]
test151_m = np.zeros(shape = (7,120,25))
test151_c = np.zeros(shape = (7,120,25))
for index in range(7):
for i in range(120):
test151_c[index][i][pq_for_m[index]+3]+=0.005
# -
test151result = []
for i in range(len(test151_m)):
print "Start tests for moment"+ str(pq_for_m[i]+4)
test151result.append(do_tests(test151_init,i,test151_m[i],test151_c[i],6))
# +
size_ratio_gau = np.zeros(shape = (40))
m1_size_gau = np.zeros(shape = (7,40))
m2_size_gau = np.zeros(shape = (7,40))
for i in range(40):
size_ratio_gau[i] = test15result[0][i]['gal_hlr']/test15result[0][i]['psf_hlr']
for j in range(7):
m1_size_gau[j][i] = (test15result[j][i+40]['abs_bias'][0] - test15result[j][i]['abs_bias'][0])/0.01/0.005
m2_size_gau[j][i] = (test15result[j][i+80]['abs_bias'][1] - test15result[j][i]['abs_bias'][1])/0.01/0.005
# +
size_ratio_ser = np.zeros(shape = (40))
m1_size_ser = np.zeros(shape = (7,40))
m2_size_ser = np.zeros(shape = (7,40))
for i in range(40):
size_ratio_ser[i] = test151result[0][i]['gal_hlr']/test151result[0][i]['psf_hlr']
for j in range(7):
m1_size_ser[j][i] = (test151result[j][i+40]['abs_bias'][0] - test151result[j][i]['abs_bias'][0])/0.01/0.005
m2_size_ser[j][i] = (test151result[j][i+80]['abs_bias'][1] - test151result[j][i]['abs_bias'][1])/0.01/0.005
# -
y_range_15 = {}
for j in range(7):
p,q = pqlist[pq_for_m[j]][0],pqlist[pq_for_m[j]][1]
n = p+q
if n not in y_range_15.keys():
y_range_15[n] = [0,0]
#print min(min(np.array([t["abs_bias"][0] for t in test13result[j]])/0.005),y_range[n][0])
y_range_15[n][0] = min(min(m1_size_gau[j]*1.1),y_range_15[n][0])
y_range_15[n][0] = min(min(m2_size_gau[j]*1.1),y_range_15[n][0])
y_range_15[n][0] = min(min(m1_size_ser[j]*1.1),y_range_15[n][0])
y_range_15[n][0] = min(min(m2_size_ser[j]*1.1),y_range_15[n][0])
y_range_15[n][1] = max(max(m1_size_gau[j]*1.1),y_range_15[n][1])
y_range_15[n][1] = max(max(m2_size_gau[j]*1.1),y_range_15[n][1])
y_range_15[n][1] = max(max(m1_size_ser[j]*1.1),y_range_15[n][1])
y_range_15[n][1] = max(max(m2_size_ser[j]*1.1),y_range_15[n][1])
print y_range_15
# +
fig = plt.figure(figsize = (12,6))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0, hspace=0)
for j in range(7):
p,q = pqlist[pq_for_m[j]][0],pqlist[pq_for_m[j]][1]
n = p+q
position = 1+j
if j>2: position = 2+j
ax = plt.subplot(2,4,position)
plt.plot(size_ratio_gau,m1_size_gau[j],'--',color = 'blue',label = r'Gaussian $g_1$')
plt.plot(size_ratio_gau,m2_size_gau[j],'--',color = 'orange',label = r'Gaussian $g_2$')
plt.plot(size_ratio_ser,m1_size_ser[j],color = 'blue',label = r'Sersic n=3 $g_1$')
plt.plot(size_ratio_ser,m2_size_ser[j],color = 'orange',label = r'Sersic n=3 $g_2$')
#print test4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
plt.ticklabel_format(axis='y',style='sci',scilimits=(0,3))
ax.tick_params(
axis='x', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
bottom=True, # ticks along the bottom edge are off
top=True, # ticks along the top edge are off
labelbottom=False)
ax.tick_params(
axis='y', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
left=True, # ticks along the bottom edge are off
right=False, # ticks along the top edge are off
labelleft=False)
#ax.tick_params(axis="y",direction="in")
if j in list(range(3,7)):
plt.xlabel(r"$\sigma_{galaxy}/\sigma_{PSF}$")
ax.tick_params(
axis='x', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
bottom=True, # ticks along the bottom edge are off
top=True, # ticks along the top edge are off
labelbottom=True)
if j in [0,3]:
plt.ylabel(r'$ m / B[ \mathbf{m}_{p,q}]$')
plt.ticklabel_format(axis='y',style='scientific',scilimits=(0,3))
ax.tick_params(
axis='y', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
left=True, # ticks along the bottom edge are off
right=False, # ticks along the top edge are off
labelleft=True)
plt.ylim(y_range_15[n])
#plt.xlabel(r"$\sigma_{galaxy}/\sigma_{PSF}$")
#plt.ylabel(r'$ m / B[ \mathbf{m}_{p,q}]$')
plt.title(str((p,q)),y = 0.8)
#plt.show()
#plt.legend()
plt.subplot(2,4,4,frame_on = False)
plt.plot([0],[0],'--',color = 'blue',label = r'Gaussian $g_1$')
plt.plot([0],[0],'--',color = 'orange',label = r'Gaussian $g_2$')
plt.plot([0],[0],color = 'blue',label = r'Sersic n = 3.0 $g_1$')
plt.plot([0],[0],color = 'orange',label = r'Sersic n = 3.0 $g_2$')
plt.axis('off')
plt.legend(fontsize = 'medium',frameon = False)
# -
psf = galsim.Gaussian(sigma = 1.0)
image = psf.drawImage(scale = 0.1,method = 'no_pixel')
FWHM = psf.calculateFWHM()
# +
test17_init = [("gaussian" ,0.5+0.1*i ,0.1,0.26,1e-8,1e-8,"gaussian" ,1.5 ,{'subtract_intersection':True}) for i in range(40)
]+[("gaussian" ,0.5+0.1*i ,0.1,0.26,0.01,1e-8,"gaussian" ,1.5 ,{'subtract_intersection':True}) for i in range(40)
]+[("gaussian" ,0.5+0.1*i ,0.1,0.26,1e-8,0.01,"gaussian" ,1.5 ,{'subtract_intersection':True}) for i in range(40)
]
test17_m = np.zeros(shape = (22,120,25))
test17_c = np.zeros(shape = (22,120,25))
for index in range(22):
for i in range(120):
test17_c[index][i][index+3]+=0.005
# -
test17result = []
for i in range(len(test17_m)):
print( "Start tests for moment"+ str(i+4))
test17result.append(do_tests_speed(test17_init,i,test17_m[i],test17_c[i],6))
# +
size_ratio_gau = np.zeros(shape = (40))
m1_size_gau = np.zeros(shape = (22,40))
m2_size_gau = np.zeros(shape = (22,40))
for i in range(40):
size_ratio_gau[i] = test17result[0][i]['gal_hlr']/test17result[0][i]['psf_hlr']
for j in range(22):
m1_size_gau[j][i] = (test17result[j][i+40]['abs_bias'][0] - test17result[j][i]['abs_bias'][0])/0.01/0.005
m2_size_gau[j][i] = (test17result[j][i+80]['abs_bias'][1] - test17result[j][i]['abs_bias'][1])/0.01/0.005
# +
size_ratio_ser = np.zeros(shape = (40))
m1_size_ser = np.zeros(shape = (22,40))
m2_size_ser = np.zeros(shape = (22,40))
for i in range(40):
size_ratio_ser[i] = test171result[0][i]['gal_hlr']/test171result[0][i]['psf_hlr']
for j in range(22):
m1_size_ser[j][i] = (test171result[j][i+40]['abs_bias'][0] - test171result[j][i]['abs_bias'][0])/0.01/0.005
m2_size_ser[j][i] = (test171result[j][i+80]['abs_bias'][1] - test171result[j][i]['abs_bias'][1])/0.01/0.005
# -
y_range_15 = {}
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
if n not in y_range_15.keys():
y_range_15[n] = [0,0]
#print min(min(np.array([t["abs_bias"][0] for t in test13result[j]])/0.005),y_range[n][0])
y_range_15[n][0] = min(min(m1_size_gau[j]*1.1),y_range_15[n][0])
y_range_15[n][0] = min(min(m2_size_gau[j]*1.1),y_range_15[n][0])
y_range_15[n][0] = min(min(m1_size_ser[j]*1.1),y_range_15[n][0])
y_range_15[n][0] = min(min(m2_size_ser[j]*1.1),y_range_15[n][0])
y_range_15[n][1] = max(max(m1_size_gau[j]*1.1),y_range_15[n][1])
y_range_15[n][1] = max(max(m2_size_gau[j]*1.1),y_range_15[n][1])
y_range_15[n][1] = max(max(m1_size_ser[j]*1.1),y_range_15[n][1])
y_range_15[n][1] = max(max(m2_size_ser[j]*1.1),y_range_15[n][1])
with open("../plots2/pickle/mul_size_ratio.pkl","wb") as f:
pickle.dump([pqlist,test17result,test171result ],f)
with open('../plots2/pickle/mul_size_ratio.pkl','rb') as f:
pqlist,test17result,test171result = pickle.load(f)
# +
pqlist = test1.sxm.get_pq_full(6)[3:]
fig = plt.figure(figsize = (21,12))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.0, hspace=0.0)
# f, axes = plt.subplots(4, 7, sharex='col', sharey='row', figsize=(21,12))
# f.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.0, hspace=0.0)
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
#print n
ax = plt.subplot(4,7,1+7*(n-3)+p)
plt.plot(size_ratio_gau,m1_size_gau[j],color = 'blue',label = r'Gaussian $g_1$')
plt.plot(size_ratio_gau,m2_size_gau[j],color = 'orange',label = r'Gaussian $g_2$')
plt.plot(size_ratio_ser,m1_size_ser[j],'--',color = 'blue',label = r'Gaussian $g_1$')
plt.plot(size_ratio_ser,m2_size_ser[j],'--',color = 'orange',label = r'Gaussian $g_2$')
# plt.plot(np.array([t['gal_hlr']/t['psf_hlr'] for t in test132result[j]]),np.array([t["abs_bias"][0] for t in test131result[j]])/0.005,'.-',color = 'blue')
# plt.plot(np.array([t['gal_hlr']/t['psf_hlr'] for t in test132result[j]]),np.array([t["abs_bias"][1] for t in test131result[j]])/0.005,'.-',color = 'orange')
#print test4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
ax.tick_params(
axis='x', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
bottom=True, # ticks along the bottom edge are off
top=True, # ticks along the top edge are off
labelbottom=False)
ax.tick_params(
axis='y', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
left=True, # ticks along the bottom edge are off
right=False, # ticks along the top edge are off
labelleft=False)
#ax.tick_params(axis="y",direction="in")
if j in list(range(15,22)):
plt.xlabel(r"$R_h^{galaxy}/R_h^{PSF}$")
ax.tick_params(
axis='x', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
bottom=True, # ticks along the bottom edge are off
top=True, # ticks along the top edge are off
labelbottom=True)
if j in [0,4,9,15]:
plt.ylabel(r'$\delta g_i / \delta m_{p,q}$')
plt.ticklabel_format(axis='y',style='scientific',scilimits=(0,3))
ax.tick_params(
axis='y', # changes apply to the x-axis
direction = 'in',
which='both', # both major and minor ticks are affected
left=True, # ticks along the bottom edge are off
right=False, # ticks along the top edge are off
labelleft=True)
plt.ylim(y_range_15[n])
plt.title(str((p,q)),y = 0.8)
#plt.show()
#plt.legend([])
plt.subplot(4,7,7,frame_on = False)
plt.plot([0],[0],color = 'blue',label = r'Gaussian $g_1$')
plt.plot([0],[0],color = 'orange',label = r'Gaussian $g_2$')
plt.plot([0],[0],'--',color = 'blue',label = r'Sersic n = 3.0 $g_1$')
plt.plot([0],[0],'--',color = 'orange',label = r'Sersic n = 3.0 $g_2$')
plt.axis('off')
plt.legend(fontsize = 'large',frameon = False)
#fig.colorbar(axes)
# +
test171_init = [("sersic" ,1.0+0.2*i ,0.1,0.26,1e-8,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':3.0}) for i in range(40)
]+[("sersic" ,1.0+0.2*i ,0.1,0.26,0.01,1e-8,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':3.0}) for i in range(40)
]+[("sersic" ,1.0+0.2*i ,0.1,0.26,1e-8,0.01,"gaussian" ,3.0 ,{'subtract_intersection':True,'sersicn':3.0}) for i in range(40)
]
test171_m = np.zeros(shape = (22,120,25))
test171_c = np.zeros(shape = (22,120,25))
for index in range(22):
for i in range(120):
test171_c[index][i][index+3]+=0.005
# -
test171result = []
for i in range(len(test171_m)):
print( "Start tests for moment"+ str(i+4))
test171result.append(do_tests_speed(test171_init,i,test171_m[i],test171_c[i],6))
size_ratio_cosmos = np.load('data/size_ratio_array.npy')
size_ratio_cosmos = size_ratio_cosmos[size_ratio_cosmos<2.9]
print(size_ratio_gau)
HSC_moment_bias = np.load('data/mean_residual.npy')
# +
from scipy import interpolate
g1_m = []; g2_m = []
for i in range(22):
# this_f1 = interpolate.LinearNDInterpolator(x, dg1[i])
# this_f2 = interpolate.LinearNDInterpolator(x, dg2[i])
this_f1 = interpolate.interp1d(size_ratio_gau, m1_size_gau[i])
m1 = this_f1(size_ratio_cosmos)
g1_m.append(np.mean(m1) * HSC_moment_bias[i+3])
this_f2 = interpolate.interp1d(size_ratio_gau, m2_size_gau[i])
m2 = this_f2(size_ratio_cosmos)
g2_m.append(np.mean(m2) * HSC_moment_bias[i+3] )
# +
nob = 50
label_list = []
pqlist = test1.sxm.get_pq_full(6)
for i in range(nob):
if i < 25:
i_pre = 't'
else:
i_pre = 'r'
label1 = i_pre+str(pqlist[i%25][0])+str(pqlist[i%25][1])
label_list.append(label1)
fig, ax = plt.subplots(1,1,figsize=(8, 6))
ax.plot(np.arange(3,25),g1_m,'o',label = 'm1')
ax.plot(np.arange(3,25),g2_m,'o',label = 'm2')
ax.axvspan(6.5, 11.5, color='r', alpha=0.2, lw=0)
ax.axvspan(17.5, 24.5, color='r', alpha=0.2, lw=0)
ax.set_xticks(np.arange(3,25,1))
ax.set_xticklabels(label_list[28:], rotation='vertical', fontsize=14)
plt.grid()
plt.legend()
plt.ylabel("Multiplicative Bias")
print( "m1 = " + str(np.sum(g1_m)))
print( "m2 = " + str(np.sum(g2_m)))
# -
import pickle
with open("../plots2/pickle/mul_prelim.pkl","wb") as f:
pickle.dump([g1_m,g2_m,label_list ],f)
psf = galsim.Gaussian(sigma = 1.5)
image = psf.drawImage(scale = 1.0, method = 'no_pixel')
print(image.calculateFWHM()*0.2)
# +
pixel_size = [0.1,0.15, 0.2,0.25, 0.3]
#gal_size = 0.17 arcsec, psf_size = 0.24 arcsec, pixel_size = 0.2 arcsec
test18_init = [("gaussian" ,0.5/this_pixel ,0.28,0.28,1e-8,1e-8,"gaussian" ,0.3/this_pixel ,{'subtract_intersection':True}) for this_pixel in pixel_size
]
test18_m = np.zeros(shape = (22,5,25))
test18_c = np.zeros(shape = (22,5,25))
for index in range(22):
for i in range(5):
#test3_c[index][i][index+3]+=HSC_moment_bias[index+3]
test18_c[index][i][index+3]+=0.01
# -
test18result = []
for i in range(len(test18_m)):
print( "Start tests for moment"+ str(i+4))
test18result.append(do_tests(test18_init,i,test18_m[i],test18_c[i],6))
# +
pqlist = test1.sxm.get_pq_full(6)[3:]
fig = plt.figure(figsize = (21,12))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.5, hspace=0.5)
for j in range(22):
p,q = pqlist[j][0],pqlist[j][1]
n = p+q
ax = plt.subplot(4,7,1+7*(n-3)+p)
dg1 = np.array([t["abs_bias"][0] for t in test18result[j]])
dg2 = np.array([t["abs_bias"][1] for t in test18result[j]])
plt.plot(pixel_size,dg1,'.',label='g1')
plt.plot(pixel_size,dg2,'.',label='g2')
plt.ticklabel_format(axis='y',style='sci',scilimits=(0,3))
#print test4_gaussian_results[j][0]['psf_bvec'][:15]/test4_gaussian_results[j][0]['psf_bvec'][0]
plt.xlabel(r"pixel size (arcsec)")
plt.ylabel(r'${\Delta g_i}$')
plt.title(str((p,q)))
#plt.show()
plt.legend()
#fig.colorbar(axes)
# -
| notebooks/Single-simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7OQ5XB1xdtRD"
# ### VGG16 Image Captioning.
# + [markdown] id="AugizpuxeBpn"
# ### Imports
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="yx0XyYcXdmLZ" outputId="a91951f1-1091-4d8c-9d98-d0a90df0e069"
from matplotlib import pyplot as plt
import tensorflow as tf
import numpy as np
import os, time, sys
from PIL import Image
from tensorflow.keras import backend as K
from tensorflow import keras
from tensorflow.keras.applications import VGG16
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
tf.__version__
# + [markdown] id="pdxTWLC6eDfx"
# ### Mounting the drive
#
# We are going to load the data and their annotations from google drive. So we need to mount the drive.
# + colab={"base_uri": "https://localhost:8080/"} id="HtRrwb7idsMp" outputId="d10d5865-bde6-43e3-cfc5-ff6adc0d4ee2"
from google.colab import drive
drive.mount("/content/drive")
# + colab={"base_uri": "https://localhost:8080/"} id="0-w0pop7dsKC" outputId="733a9aa8-1802-4d7a-f6ce-04ed6dad8a19"
base_path = "/content/drive/My Drive/image-captioning/coco"
os.listdir(base_path)
# + [markdown] id="1nWxmOe1eTCI"
# ### Extracting zip files
# + colab={"base_uri": "https://localhost:8080/"} id="gQ7m_UFGdsGi" outputId="9037fbc5-0dcd-44f0-a195-63e20eaa7432"
import zipfile
# annotations
with zipfile.ZipFile(os.path.join(base_path, 'annotations_trainval2017.zip'), "r") as z:
z.extractall(base_path)
# annotations
with zipfile.ZipFile(os.path.join(base_path, 'val2017.zip'), "r") as z:
z.extractall(base_path)
print("Done")
# + id="-LQdPKrUdsCo"
import json
# + id="4ajIhIBqdr68"
def load_records():
path = os.path.join(base_path,
"annotations", "captions_val2017.json")
with open(path, "r", encoding="utf-8") as file:
data_raw = json.load(file)
images = data_raw['images']
annotations = data_raw['annotations']
records = dict()
for image in images:
record = dict()
record["filename"] = image["file_name"]
record["id"] = image["id"]
record["captions"] = list()
records[image["id"]] = record
for annotation in annotations:
record = records[annotation['image_id']]
record['captions'].append(annotation.get("caption"))
records_list = [(key, record['filename'], record['captions'])
for key, record in sorted(records.items())]
ids, filenames, captions = zip(*records_list)
return ids, filenames, captions
ids, filenames, captions = load_records()
# + colab={"base_uri": "https://localhost:8080/"} id="uUUyAh8Edr2Z" outputId="1c625ec8-c337-4718-a389-5128d07d312f"
filenames[:2], ids[:2], captions[:2]
# + [markdown] id="eQNF3YGdenF9"
# ### Get names and captions
# + id="9uNWcpu_drwB"
_, image_names, captions = load_records()
# + [markdown] id="8Vu-Lhenesrh"
# ### Counting examples
# + colab={"base_uri": "https://localhost:8080/"} id="pOJT9V-ddrsC" outputId="bc899215-c4a9-4d3b-dd0c-0abbd9ccfb55"
num_images = len(image_names)
num_images
# + [markdown] id="WOlxcva4ezEQ"
# As you can see we only have ``5000`` images in this dataset, you can use the train set which contains a lot of training examples.
# + [markdown] id="DCBuyUbLe4Zw"
# ### Helper functions
#
# We are going to define helper functions that will do the following:
#
# 1. loading images
# 2. display images
# + id="Tuu1bozSdrpD"
def load_image(path, size=(224, 224)): # vgg16 size
image = Image.open(path)
if not size is None:
image = image.resize(size=size, resample=Image.LANCZOS)
image = np.array(image, dtype=np.float32)
image = image/255.0
# gray scale image to a 3-dim RGB array
if (len(image.shape) == 2):
image = np.repeat(image[:, :, np.newaxis], 3, axis=2)
return image
# + id="Cynrg-W8drli"
def show_image(index):
assert index < num_images - 1, f"recieved index {index} for index range [0, {num_images}]"
# path to images
img_dirs = os.path.join(base_path, "val2017")
file_name = image_names[index]
caps = captions[index]
for caption in caps:
print(caption)
image = load_image(os.path.join(img_dirs, file_name))
plt.imshow(image)
plt.show()
# + [markdown] id="QYthJHVafRCw"
# ### Displaying sample examples.
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="NsxYVGzJdriw" outputId="568ed7c1-5fb9-4c93-9d27-81bef3b01b46"
show_image(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="6lQ8i1AUdrfM" outputId="af9b512f-fa26-4c83-cf79-3c9e7ea50111"
show_image(0)
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="8Vvz5RWSdrb4" outputId="ba6f4e65-532c-49d6-9268-d515896d21ed"
show_image(500)
# + [markdown] id="zMxuJEnmfzOy"
# ### Closer look to the dataset.
#
# Our dataset contains an image that has more than one caption. We want to create a function that takes an index of an image and generates the same image with a single caption in the caption list.
# + id="QrcOhqKUdrY1"
def expand_dataset(index):
img_dirs = os.path.join(base_path, "val2017")
file_name = image_names[index]
caps = captions[index]
image = load_image(os.path.join(img_dirs, file_name))
images_to_captions = []
for caption in caps:
images_to_captions.append([image, caption])
return images_to_captions
# + colab={"base_uri": "https://localhost:8080/"} id="0aLY2rmhnXER" outputId="bff23b1e-c2f8-4061-d1f7-75ddd532a365"
sample = expand_dataset(0)
len(sample)
# + id="LDSprl6Bnhth"
def display_sample(index):
img, cap = sample[index]
print(cap)
plt.imshow(img)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 555} id="GiiZNLT6nhmA" outputId="d9b7228e-88eb-48ba-8845-e240dcf8c96a"
for i in range(2):
display_sample(i)
# + [markdown] id="GZys8lIGoFN7"
# ### Expanding the dataset.
#
# We want to pair each image with it's caption.
# + id="jQ6XVW0HoHch"
images_captions = []
for i in range(int(num_images * .7)):
images_captions.extend(expand_dataset(i))
# + colab={"base_uri": "https://localhost:8080/"} id="bCq016GeoHaJ" outputId="9d0f706a-db96-450a-b92e-4722f4be5d36"
len(images_captions)
# + [markdown] id="7clCdFZVoyZp"
# ### Randomly shuffling the images to captions pairs.
# + id="FtCkQyL3osH6"
import random
# + id="YrzmjILdpCMB"
random.shuffle(images_captions)
# + id="Le-pONNypM2x"
def display_sample(index):
img, cap = images_captions[index]
print(cap)
plt.imshow(img)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 555} id="8RnU39ohpGIE" outputId="358e922e-4855-4158-b15c-ec051e1284d7"
for i in range(2):
display_sample(i)
# + [markdown] id="CbP8-9i8rb57"
# ### Text preprocessing
#
# We are going to use the Tokenizer from keras to preprocess the text into vectors of integers.
# + id="c9aAsa-frRPC"
captions = []
for _, cap in images_captions:
captions.append(f"<start> {cap} <end>")
# + colab={"base_uri": "https://localhost:8080/"} id="tZgfkSizrRJ3" outputId="8c25d8e3-ed6d-4822-828c-2c11dfba06a3"
captions[:2]
# + id="7X7PYAsfrRG6"
num_words = 10_000
# + id="obHMn3jorRDK"
tokenizer = tf.keras.preprocessing.text.Tokenizer(num_words=num_words,
oov_token="<unk>",
)
tokenizer.fit_on_texts(captions)
tokenizer.word_index['<pad>'] = 0
tokenizer.index_word[0] = '<pad>'
# + [markdown] id="I31P8c_qtDCJ"
# ### Converting text to integers
# + id="Ig8qRAFxrRAs"
token_captions = tokenizer.texts_to_sequences(captions)
# + colab={"base_uri": "https://localhost:8080/"} id="8yyhujQ_rQ8v" outputId="9ef39410-ec0d-44aa-9ade-b2d5e98afe4e"
token_captions[:2]
# + [markdown] id="7PL-37TYtGkK"
# ### Padding the sequences
# + id="6Zivce0wrQ5K"
token_captions_padded = pad_sequences(
token_captions, maxlen=20,
padding='post', truncating='post'
)
# + colab={"base_uri": "https://localhost:8080/"} id="QqlZzZbTrPy6" outputId="5845e64f-9338-482b-80ca-5632b49e739f"
token_captions_padded[:2]
# + [markdown] id="KJAhTNgIuAoN"
# > Now we need to map captions tokens with their respective images vector.
#
#
# Our image should be preprocessed
# + id="zMm7EI6vqwRv"
images = []
for image, _ in images_captions:
images.append(image)
# + id="6ADip6owqwKU"
assert len(images) == len(token_captions_padded), "images and captions must have the same length"
# + [markdown] id="1YAT8tktyKKA"
# ### Creating the dataset.
#
# We are going to create the dataset using the `tf.data.Dataset.from_tensor_slice`.
# + id="OsIJFy7OqwGg"
dataset = tf.data.Dataset.from_tensor_slices(
(images, token_captions_padded)
)
# + id="CMDN3H6bqwCX"
BATCH_SIZE = 64
BUFFER_SIZE = 1000
dataset = dataset.shuffle(BUFFER_SIZE
).batch(BATCH_SIZE
).prefetch(
buffer_size=tf.data.AUTOTUNE)
# + [markdown] id="FnGcdL5ayjtj"
# ### Building the model.
#
# We are going to build our end to end
# + id="9fqVYnQSqv-e"
# + id="pBWx4lFCqv30"
# + id="6H3ZY-pTqvy7"
# + [markdown] id="QB_1GJqyhwTB"
# ### The VGG16 model
#
# We are going to use transfare learnig (fine tunning) to us the VGG16 model to perform image classification.
#
# > If ``include_top=True`` then the whole VGG16 model is downloaded which is about 528 MB. If ``include_top=False`` then only the convolutional part of the VGG16 model is downloaded which is just ``57 MB``.
#
# We are going to download the whole model.
# + colab={"base_uri": "https://localhost:8080/"} id="0XNvVR3RdrVt" outputId="c717b5b2-e389-405b-d897-be22d39619b0"
image_model = VGG16(include_top=True, weights='imagenet')
image_model.summary()
# + [markdown] id="rvnyYPKhiTQI"
# We are going to use all the layers of the models up from the top up to ``fc2`` layer. we are not interested in the ``prediction`` layer.
# + colab={"base_uri": "https://localhost:8080/"} id="nrmLTiExdrS8" outputId="1cd4ede9-9864-468b-a387-04edce06faa0"
transfare_layer = image_model.get_layer("fc2")
transfare_layer
# + [markdown] id="qE01EvGAifZF"
# We called it the "transfer-layer" because we will transfer its output to another model that creates the image captions.
#
# To do this, first we need to create a new model which has the same input as the original ``VGG16`` model but outputs the transfer-values from the ``fc2`` layer.
# + id="_UsIzecjdrPY"
image_model_transfare = keras.Model(
inputs=image_model.input, outputs = transfare_layer.output,
name= "image_model_transfare"
)
image_model_transfare.summary()
# + [markdown] id="2goSYCc6isvA"
# ### Getting the size of the image.
#
# We are now going to get the shape of the image that `VGG16` accept as follows
# + id="QKgrT7t4drMc"
image_size = K.int_shape(image_model.input)[1:3]
image_size
# + [markdown] id="1_ocT4OTjL6r"
# For each input image, the new model will output a vector of transfer-values with the following length.
# + id="SIgaVjn0z7Xh"
class BahdanauAttention(keras.Model):
def __init__(self, units):
super(BahdanauAttention, self).__init__()
self.W1 = keras.layers.Dense(units)
self.W2 = keras.layers.Dense(units)
self.V = keras.layers.Dense(1)
def call(self, features, hidden):
# features(CNN_encoder output) shape == (batch_size, 64, embedding_dim)
# hidden shape == (batch_size, hidden_size)
# hidden_with_time_axis shape == (batch_size, 1, hidden_size)
hidden_with_time_axis = tf.expand_dims(hidden, 1)
# attention_hidden_layer shape == (batch_size, 64, units)
attention_hidden_layer = (tf.nn.tanh(self.W1(features) +
self.W2(hidden_with_time_axis)))
# score shape == (batch_size, 64, 1)
# This gives you an unnormalized score for each image feature.
score = self.V(attention_hidden_layer)
# attention_weights shape == (batch_size, 64, 1)
attention_weights = tf.nn.softmax(score, axis=1)
# context_vector shape after sum == (batch_size, hidden_size)
context_vector = attention_weights * features
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
# + id="wBHjgJWvdrJY"
class CNN_Encoder(keras.Model):
def __init__(self, embedding_dim, vgg16):
super(CNN_Encoder, self).__init__()
self.fc = keras.layers.Dense(embedding_dim)
self.vgg16 = vgg16
def call(self, x):
x = self.vgg16(x)
x = self.fc(x)
x = tf.nn.relu(x)
return x
# + id="jGuq6cH6drGA"
class RNN_Decoder(keras.Model):
def __init__(self, embedding_dim, units, vocab_size):
super(RNN_Decoder, self).__init__()
self.units = units
self.embedding = keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = keras.layers.GRU(self.units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc1 = keras.layers.Dense(self.units)
self.fc2 = keras.layers.Dense(vocab_size)
self.attention = BahdanauAttention(self.units)
def call(self, x, features, hidden):
# defining attention as a separate model
context_vector, attention_weights = self.attention(features, hidden)
# x shape after passing through embedding == (batch_size, 1, embedding_dim)
x = self.embedding(x)
# x shape after concatenation == (batch_size, 1, embedding_dim + hidden_size)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# passing the concatenated vector to the GRU
output, state = self.gru(x)
# shape == (batch_size, max_length, hidden_size)
x = self.fc1(output)
# x shape == (batch_size * max_length, hidden_size)
x = tf.reshape(x, (-1, x.shape[2]))
# output shape == (batch_size * max_length, vocab)
x = self.fc2(x)
return x, state, attention_weights
def reset_state(self, batch_size):
return tf.zeros((batch_size, self.units))
# + id="MG5ScxIddrCl"
embedding_dim = 100
encoder = CNN_Encoder(embedding_dim)
# + id="ye2n9diSdq8v"
encoder = CNN_Encoder(embedding_dim)
# + id="uDiq89pYdq5M"
# + id="yFPFx-hTdq2o"
# + id="5GnjCZ5Jdqzh"
# + id="fxD74Bkedqwa"
| 18_Image_Captioning/03_Image_Caption_Splits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
data_set= pd.read_csv('profit_data.csv')
data_set
data_set.iloc[0] # first row of data frame (<NAME>) - Note a Series data type output.
data_set.iloc[-1] # last row of data frame (<NAME>)
# Columns:
data_set.iloc[:,2] # first column of data frame (RD)
data_set.iloc[:,-1] # last column of data frame (Profit)
| ML2/linear_regression/Untitled.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ + cell_id="61b59a28-1fd4-4692-8385-1b48336f7075" deepnote_cell_height=99.46875 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1653428092652 source_hash="b01a95aa" tags=[]
#!pip install -U pandasql
#!pip install statsmodels
/ + cell_id="00001-006bee7e-c7fc-4d42-9110-3333d1514c68" deepnote_cell_height=207.453125 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=6216 execution_start=1653344102532 source_hash="ee12eb6e" tags=[]
import pandas as pd
pd.set_option('max_columns', None)
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
#from pandasql import sqldf
import plotly.express as px
/ + cell_id="00002-a38e98ca-f9d5-4cc1-a610-b761b11d5a55" deepnote_cell_height=360.859375 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=103 execution_start=1653428622814 owner_user_id="dbcfdc03-cb41-42df-88d8-3fffbb8930b7" source_hash="45c4a4d" tags=[]
# Bring in the pickled files
# if any of the pickle files cannot be reached, please update the directories
# to the download loaction of the zip then navigate to data/Races_Combined
allraces21 = pd.read_pickle("/work/Milestone_I/data/Races_Combined/allraces_21.pkl")
allraces22 = pd.read_pickle("/work/Milestone_I/data/Races_Combined/allraces_22.pkl")
sameraces21 = pd.read_pickle("/work/Milestone_I/data/Races_Combined/sameraces_21.pkl")
sameraces22 = pd.read_pickle("/work/Milestone_I/data/Races_Combined/sameraces_22.pkl")
sameraces21_22 = pd.read_pickle("/work/Milestone_I/data/Races_Combined/sameraces_21_22.pkl")
allraces21_22 = pd.concat([allraces21, allraces22], ignore_index=True)
/ + cell_id="00003-850e964e-550b-418d-b6d5-190f0d5a69af" deepnote_cell_height=81.453125 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=20 execution_start=1653344108866 source_hash="c796b328" tags=[]
#allraces21_22.columns
/ + [markdown] cell_id="00004-67331db1-904e-4e6a-b1fc-73a0e919034f" deepnote_cell_height=52.859375 deepnote_cell_type="markdown" tags=[]
/ This notebook is used to look at the Green Passes - comparing 2021 to 2022...
/ + cell_id="00005-95239218-a997-4496-92d6-8a4572afca8f" deepnote_cell_height=171.453125 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=53 execution_start=1653344108898 source_hash="f358f12d" tags=[]
# Starting with just plain passing year over year
passdiff_yoy = allraces21_22.groupby(['Season'])[['Pass Differential']].mean().reset_index()
grnpass_yoy = allraces21_22.groupby(['Season'])[['Green Passes']].mean().reset_index()
#grnpassed_yoy = allraces21_22.groupby(['Season'])[['Green Passed']].mean().reset_index()
qltypass_yoy = allraces21_22.groupby(['Season'])[['Quality Passes']].mean().reset_index()
pctqltypass_yoy = allraces21_22.groupby(['Season'])[['% Quality Passes']].mean().reset_index()
/ + cell_id="00006-2f7b31f8-7b9e-487b-862f-304bd7511787" deepnote_cell_height=81.453125 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1653344108958 source_hash="9e1bc834" tags=[]
#all_passing_df = pd.merge(passing_yoy, passdiff_yoy, on="Season")
/ + cell_id="00007-e19b829a-fcec-4079-997c-50bb71dae7c9" deepnote_cell_height=117.453125 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=35 execution_start=1653344108981 owner_user_id="45152886-e80a-4457-8e84-9d99a262197d" source_hash="779bf94d" tags=[]
from functools import reduce
data_frames = [passdiff_yoy, grnpass_yoy, qltypass_yoy, pctqltypass_yoy]
all_passing_df = reduce(lambda left,right: pd.merge(left,right,on=['Season'], how='outer'), data_frames)
/ + cell_id="00008-43535123-5b43-4b87-b53c-8edb6363c594" deepnote_cell_height=272.90625 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=339 execution_start=1653344109023 source_hash="c5d1a95" tags=[]
all_passing_df
/ + [markdown] cell_id="00009-5aa53f89-ad77-453e-b46f-fd6758ee3f23" deepnote_cell_height=75.265625 deepnote_cell_type="markdown" tags=[]
/ The amount of difference between Green Passes in 2021 vs 2022 looks like it could be significant. Next lets do a ttest on this and maybe a correlation to find out what those tell us...
/ + cell_id="00010-938cd3e5-0f50-4507-8e15-ec55bd969e91" deepnote_cell_height=226.890625 deepnote_cell_type="code" deepnote_output_heights=[21.171875] deepnote_to_be_reexecuted=false execution_millis=203 execution_start=1653344109159 source_hash="dea45e7c" tags=[]
import scipy.stats as stats
gpdata_21 = allraces21['Green Passes'].to_list()
gpdata_22 = allraces22['Green Passes'].to_list()
#perform two sample t-test with equal variances
stats.ttest_ind(a=gpdata_21, b=gpdata_22, equal_var=True)
/ + [markdown] cell_id="00011-59e3171a-c3e0-47e0-8d22-4bcf5aa67974" deepnote_cell_height=52.859375 deepnote_cell_type="markdown" tags=[]
/ This shows that these two samples are statistically significant. We'll make a plot of this as a possibility for our slides...
/ + cell_id="00012-6c9eb2cc-65e5-4b01-861e-fbff77ee5e37" deepnote_cell_height=208.875 deepnote_cell_type="code" deepnote_output_heights=[21.171875] deepnote_to_be_reexecuted=false execution_millis=198 execution_start=1653344109165 source_hash="7096729a" tags=[]
# Curious about pass differential - lets do the same thing.
gpdata_21 = allraces21['Pass Differential'].to_list()
gpdata_22 = allraces22['Pass Differential'].to_list()
#perform two sample t-test with equal variances
stats.ttest_ind(a=gpdata_21, b=gpdata_22, equal_var=True)
/ + [markdown] cell_id="00013-030ba713-4e64-4c0d-917b-8ca0cc210302" deepnote_cell_height=52.859375 deepnote_cell_type="markdown" tags=[]
/ Pass differential is not much different. The amount of passing is but the difference between drivers is not.
/ + [markdown] cell_id="00014-1b8edd75-13ac-4c57-ab6c-dfa5c9698173" deepnote_cell_height=52.859375 deepnote_cell_type="markdown" tags=[]
/ Same thing except for just the races that are the same between 2021 and 2022...
/ + cell_id="00015-210834ce-f3af-42b7-92ff-a0f74ae494a3" deepnote_cell_height=416.90625 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=172 execution_start=1653344109191 source_hash="a6357337" tags=[]
# Starting with just plain passing year over year
passdiff_yoy = sameraces21_22.groupby(['Season'])[['Pass Differential']].mean().reset_index()
grnpass_yoy = sameraces21_22.groupby(['Season'])[['Green Passes']].mean().reset_index()
#grnpassed_yoy = sameraces21_22.groupby(['Season'])[['Green Passed']].mean().reset_index()
qltypass_yoy = sameraces21_22.groupby(['Season'])[['Quality Passes']].mean().reset_index()
pctqltypass_yoy = sameraces21_22.groupby(['Season'])[['% Quality Passes']].mean().reset_index()
data_frames = [passdiff_yoy, grnpass_yoy, qltypass_yoy, pctqltypass_yoy]
sr_all_passing_df = reduce(lambda left,right: pd.merge(left,right,on=['Season'], how='outer'), data_frames)
sr_all_passing_df
/ + [markdown] cell_id="00016-eefcd40f-d697-447a-af48-9ec0d77f3750" deepnote_cell_height=52.859375 deepnote_cell_type="markdown" tags=[]
/ Lets do the same ttest on just the races that are the same...
/ + cell_id="00017-8dd589c4-0d2b-46f9-85cc-022be4441b42" deepnote_cell_height=190.890625 deepnote_cell_type="code" deepnote_output_heights=[21.171875] deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1653344109417 source_hash="81f0fdc" tags=[]
gpdata_21 = sameraces21['Green Passes'].to_list()
gpdata_22 = sameraces22['Green Passes'].to_list()
#perform two sample t-test with equal variances
stats.ttest_ind(a=gpdata_21, b=gpdata_22, equal_var=True)
/ + [markdown] cell_id="00018-71a2d632-09ab-4347-a94f-8314332310fa" deepnote_cell_height=52.859375 deepnote_cell_type="markdown" tags=[]
/ The pvalue changed very slightly. Still significant.
/ + cell_id="00019-338bf03c-985a-475f-9a73-23e56c198235" deepnote_cell_height=190.890625 deepnote_cell_type="code" deepnote_output_heights=[21.171875] deepnote_to_be_reexecuted=false execution_millis=11 execution_start=1653344109419 source_hash="7308027d" tags=[]
gpdata_21 = sameraces21['Pass Differential'].to_list()
gpdata_22 = sameraces22['Pass Differential'].to_list()
#perform two sample t-test with equal variances
stats.ttest_ind(a=gpdata_21, b=gpdata_22, equal_var=True)
/ + [markdown] cell_id="00020-d1e7de8c-3299-4e08-be79-576269a510a5" deepnote_cell_height=52.859375 deepnote_cell_type="markdown" tags=[]
/ Same outcome as all races. Pass differential is very close to the same.
/ + cell_id="00021-81ee9314-14c2-4803-bb3c-10fbecf7ef48" deepnote_cell_height=135.46875 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=83189986 execution_start=1653344109462 source_hash="c539360a" tags=[]
#Going to start a plot here...
plt.style.use('_mpl-gallery')
/ + cell_id="00022-8db948ab-c8e5-4c71-af91-0eca165df08e" deepnote_cell_height=171.453125 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=11 execution_start=1653344109463 source_hash="3175eaaa" tags=[]
grnpass_std = sameraces21_22.groupby(['Season'])[['Green Passes']].std().reset_index()
grnpass_std.rename(columns={"Green Passes": "Green Passes Std Dev"}, inplace = True)
grnpass_avg = sameraces21_22.groupby(['Season'])[['Green Passes']].mean().reset_index()
grnpass_avg.rename(columns={"Green Passes": "Green Passes Mean"}, inplace = True)
grnpass_df = pd.merge(grnpass_std, grnpass_avg, on="Season")
/ + cell_id="00023-1352020f-258c-4911-b8b3-2638198189ca" deepnote_cell_height=272.90625 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=25 execution_start=1653344109478 source_hash="b5ef737e" tags=[]
grnpass_df
/ + cell_id="00024-892838c0-5023-4c06-9d58-7c6e4e67d924" deepnote_cell_height=225.46875 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1653344109551 source_hash="e89ce2ba" tags=[]
# Changing 'Same Races - Race #' out for 'Track'
grnpass_list21 = sameraces21.groupby(['Track'])['Green Passes'].sum().tolist()
grnpass_list22 = sameraces22.groupby(['Track'])['Green Passes'].sum().tolist()
# racenum_list21 = sameraces21.groupby(['Same Races - Race #'])['Same Races - Race #'].unique().tolist()
# racenum_list22 = sameraces22.groupby(['Same Races - Race #'])['Same Races - Race #'].unique().tolist()
# racenum_list21 = sameraces21['Same Races - Race #'].unique().tolist()
# racenum_list22 = sameraces22['Same Races - Race #'].unique().tolist()
racenum_list = np.arange(1,12,1)
/ + cell_id="00025-5624a0f4-1d79-4979-9b56-c0119949c43a" deepnote_cell_height=99.46875 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=43 execution_start=1653344109551 source_hash="49750c5f" tags=[]
track_list21 = sameraces21.groupby(['Track'])['Track'].min()
track_list21 = track_list21.to_list()
/ + cell_id="00026-9a9706c5-0962-4e80-974e-03fedd1d929e" deepnote_cell_height=670.671875 deepnote_cell_type="code" deepnote_output_heights=[356.984375] deepnote_to_be_reexecuted=false execution_millis=724 execution_start=1653344109595 source_hash="f61168d6" tags=[]
#import plotly.graph_objects as go
fig, axes = plt.subplots(1, figsize=(5,5))
plt.sca(axes)
plt.plot(racenum_list,grnpass_list21,'#FFD93D')
plt.plot(racenum_list,grnpass_list22,'#FF0F27')
plt.legend(['2021','2022'], loc='upper left')
# plt.grid() not sure I love this with a grid line or not
plt.title('Green Passes: 2021 vs 2022',fontweight="bold")
#fig.update_layout(title={'text': "Green Passes: 2021 vs 2022"})
# plt.show()
plt.savefig('green_pass_21_22.png') # saving the chart
/ + allow_embed=false cell_id="00027-00227c20-65cd-4582-b7c2-8fe50fba9d71" deepnote_cell_height=817.6875 deepnote_cell_type="code" deepnote_output_heights=[413.984375] deepnote_to_be_reexecuted=false execution_millis=1114 execution_start=1653344110314 source_hash="d6acc579" tags=[]
#import plotly.graph_objects as go
from matplotlib import rc,rcParams
fig, axes = plt.subplots(1, figsize=(5,5))
plt.sca(axes)
plt.plot(track_list21,grnpass_list21,'#FFD93D')
plt.plot(track_list21,grnpass_list22,'#FF0F27')
plt.legend(['2021','2022'], loc='upper left')
axes.set_ylabel('Total Green Passes', fontweight='bold')
axes.set_xlabel('Tracks', fontweight='bold')
plt.grid(False)
plt.xticks(rotation = 45)
# plt.grid() not sure I love this with a grid line or not
plt.title('Green Passes: 2021 vs 2022',fontweight="bold", fontsize=20)
#fig.update_layout(title={'text': "Green Passes: 2021 vs 2022"})
# plt.show()
plt.savefig('green_pass_21_22.png') # saving the chart
/ + cell_id="00028-54f32608-9112-45f6-a0cb-7c2c8017b8e4" deepnote_cell_height=707.6875 deepnote_cell_type="code" deepnote_output_heights=[303.984375] deepnote_to_be_reexecuted=false execution_millis=1123 execution_start=1653344111486 source_hash="49a7b146" tags=[]
#same information as above except as a bar
from matplotlib import rc,rcParams
fig = plt.figure(figsize=(5,3))
ax = fig.add_axes([0,0,1,1])
ax.bar(track_list21,grnpass_list21,color='#FFD93D',width = 0.4, align = 'edge')
ax.bar(track_list21,grnpass_list22,color='#FF0F27',width = 0.25, align = 'center')
plt.legend(['2021','2022'], loc='upper left')
axes.set_ylabel('Total Green Passes', fontweight='bold')
axes.set_xlabel('Tracks', fontweight='bold')
plt.grid(False)
plt.xticks(rotation = 45)
# plt.grid() not sure I love this with a grid line or not
plt.title('Green Passes: 2021 vs 2022',fontweight="bold", fontsize=20)
#fig.update_layout(title={'text': "Green Passes: 2021 vs 2022"})
# plt.show()
plt.savefig('green_pass_21_22_bar.png', bbox_inches='tight') # saving the chart
/ + cell_id="00029-97d29b24-516a-497c-8c56-ba522bc02ba1" deepnote_cell_height=844.6875 deepnote_cell_type="code" deepnote_output_heights=[332.984375] deepnote_to_be_reexecuted=false execution_millis=881 execution_start=1653344112615 source_hash="290374bd" tags=[]
fig, (ax1, ax2) = plt.subplots(2, figsize=(10,5))
plt.sca(ax1)
plt.plot(racenum_list,grnpass_list21, 'o')
#obtain m (slope) and b(intercept) of linear regression line
m, b = np.polyfit(racenum_list,grnpass_list21, 1)
#add linear regression line to scatterplot
plt.plot(racenum_list, m*racenum_list+b,'#FFD93D',linewidth=5,alpha=0.5)
plt.title("2021 Green Pass Trend",fontweight="bold")
plt.sca(ax2)
plt.plot(racenum_list,grnpass_list22, 'o')
#obtain m (slope) and b(intercept) of linear regression line
m, b = np.polyfit(racenum_list,grnpass_list22, 1)
#add linear regression line to scatterplot
plt.plot(racenum_list, m*racenum_list+b,'#FF0F27',linewidth=5,alpha=0.5)
plt.title("2022 Green Pass Trend",fontweight="bold")
#adding some spacing to the subplots so the titles don't overlap
plt.subplots_adjust(left=0.1,bottom=0.1,right=0.9,top=0.9,wspace=0.4,hspace=0.4)
# plt.show()
plt.savefig('green_pass_trends.png') # saving the chart
/ + [markdown] cell_id="00030-ff9f16d2-7eff-4efc-8e7e-86cc4fb4fae3" deepnote_cell_height=75.265625 deepnote_cell_type="markdown" tags=[]
/ Going to re-do the regression lines using the race # instead of the tracks to see how passing progressed throughout the season instead of putting them in order of tracks...
/ + cell_id="00031-413cbd2a-62a8-41bf-8b5d-039afebb732f" deepnote_cell_height=135.46875 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1653344113480 source_hash="d11ce993" tags=[]
# Using 'Same Races - Race #' instead of 'Track' - this will put the races in order
grnpass_list21 = sameraces21.groupby(['Same Races - Race #'])['Green Passes'].sum().tolist()
grnpass_list22 = sameraces22.groupby(['Same Races - Race #'])['Green Passes'].sum().tolist()
racenum_list = np.arange(1,12,1)
/ + cell_id="00032-56a96fd3-df58-468c-b957-b174f319d8cf" deepnote_cell_height=118.875 deepnote_cell_type="code" deepnote_output_heights=[21.171875] deepnote_to_be_reexecuted=false execution_millis=20 execution_start=1653344113491 source_hash="bbe2b45b" tags=[]
grnpass_list21
/ + cell_id="00033-19d82c90-911d-493d-bb69-78f134048fe5" deepnote_cell_height=118.875 deepnote_cell_type="code" deepnote_output_heights=[21.171875] deepnote_to_be_reexecuted=false execution_millis=26 execution_start=1653344113511 source_hash="fe5f4351" tags=[]
grnpass_list22
/ + [markdown] cell_id="00034-c9572b26-84a4-4159-8bd9-82bca6044aea" deepnote_cell_height=52.859375 deepnote_cell_type="markdown" tags=[]
/ It might be good to have the Track Names along the x-axis...
/ + cell_id="00035-e8bde5a4-2fee-424c-aabe-4f87cce1b449" deepnote_cell_height=99.46875 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=23 execution_start=1653344113536 source_hash="da965992" tags=[]
track_list21 = sameraces21.groupby(['Same Races - Race #'])['Track'].min()
track_list22 = sameraces22.groupby(['Same Races - Race #'])['Track'].min()
/ + cell_id="00036-f93ce5f6-8168-47d8-9489-72b37a7d4c83" deepnote_cell_height=99.46875 deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1653344113576 source_hash="cc719a12" tags=[]
track_list21 = track_list21.to_list()
track_list22 = track_list22.to_list()
/ + cell_id="00037-b5c5308b-a062-4e79-90af-b09269827507" deepnote_cell_height=1266.71875 deepnote_cell_type="code" deepnote_output_heights=[611] deepnote_to_be_reexecuted=false execution_millis=1740 execution_start=1653344113597 source_hash="4f361a1e" tags=[]
fig, (ax1, ax2) = plt.subplots(2, figsize=(10,10))
plt.sca(ax1)
plt.plot(racenum_list,grnpass_list21, 'o')
#obtain m (slope) and b(intercept) of linear regression line
m, b = np.polyfit(racenum_list,grnpass_list21, 1)
#add linear regression line to scatterplot
plt.plot(track_list21, m*racenum_list+b,'#FFD93D',linewidth=5,alpha=0.5)
ax1.set_ylabel('Total Green Passes')
ax1.set_xlabel('Races In Season Order')
plt.grid(False)
plt.xticks(rotation = 45)
plt.title("2021 Green Pass Trend",fontweight="bold", fontsize=20)
plt.sca(ax2)
plt.plot(racenum_list,grnpass_list22, 'o')
#obtain m (slope) and b(intercept) of linear regression line
m, b = np.polyfit(racenum_list,grnpass_list22, 1)
#add linear regression line to scatterplot
plt.plot(track_list22, m*racenum_list+b,'#FF0F27',linewidth=5,alpha=0.5)
ax2.set_ylabel('Total Green Passes')
ax2.set_xlabel('Races In Season Order')
plt.grid(False)
plt.xticks(rotation = 45)
plt.title("2022 Green Pass Trend",fontweight="bold", fontsize=20)
#adding some spacing to the subplots so the titles don't overlap
plt.subplots_adjust(left=0.1,bottom=0.1,right=0.9,top=0.9,wspace=0.4,hspace=0.4)
# plt.show()
plt.savefig('green_pass_trends.png') # saving the chart
/ + [markdown] cell_id="00038-527fbda6-05cd-4415-9173-d75fc3738acd" deepnote_cell_height=46.484375 deepnote_cell_type="markdown" tags=[]
/
/ + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
/ <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=9ef4eb23-f38f-47e8-bfb0-51d8616b0dee' target="_blank">
/ <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODBweCIgaGVpZ2h0PSI4MHB4IiB2aWV3Qm94PSIwIDAgODAgODAiIHZlcnNpb249IjEuMSIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayI+CiAgICA8IS0tIEdlbmVyYXRvcjogU2tldGNoIDU0LjEgKDc2NDkwKSAtIGh0dHBzOi8vc2tldGNoYXBwLmNvbSAtLT4KICAgIDx0aXRsZT5Hcm91cCAzPC90aXRsZT4KICAgIDxkZXNjPkNyZWF0ZWQgd2l0aCBTa2V0Y2guPC9kZXNjPgogICAgPGcgaWQ9IkxhbmRpbmciIHN0cm9rZT0ibm9uZSIgc3Ryb2tlLXdpZHRoPSIxIiBmaWxsPSJub25lIiBmaWxsLXJ1bGU9ImV2ZW5vZGQiPgogICAgICAgIDxnIGlkPSJBcnRib2FyZCIgdHJhbnNmb3JtPSJ0cmFuc2xhdGUoLTEyMzUuMDAwMDAwLCAtNzkuMDAwMDAwKSI+CiAgICAgICAgICAgIDxnIGlkPSJHcm91cC0zIiB0cmFuc2Zvcm09InRyYW5zbGF0ZSgxMjM1LjAwMDAwMCwgNzkuMDAwMDAwKSI+CiAgICAgICAgICAgICAgICA8cG9seWdvbiBpZD0iUGF0aC0yMCIgZmlsbD0iIzAyNjVCNCIgcG9pbnRzPSIyLjM3NjIzNzYyIDgwIDM4LjA0NzY2NjcgODAgNTcuODIxNzgyMiA3My44MDU3NTkyIDU3LjgyMTc4MjIgMzIuNzU5MjczOSAzOS4xNDAyMjc4IDMxLjY4MzE2ODMiPjwvcG9seWdvbj4KICAgICAgICAgICAgICAgIDxwYXRoIGQ9Ik0zNS4wMDc3MTgsODAgQzQyLjkwNjIwMDcsNzYuNDU0OTM1OCA0Ny41NjQ5MTY3LDcxLjU0MjI2NzEgNDguOTgzODY2LDY1LjI2MTk5MzkgQzUxLjExMjI4OTksNTUuODQxNTg0MiA0MS42NzcxNzk1LDQ5LjIxMjIyODQgMjUuNjIzOTg0Niw0OS4yMTIyMjg0IEMyNS40ODQ5Mjg5LDQ5LjEyNjg0NDggMjkuODI2MTI5Niw0My4yODM4MjQ4IDM4LjY0NzU4NjksMzEuNjgzMTY4MyBMNzIuODcxMjg3MSwzMi41NTQ0MjUgTDY1LjI4MDk3Myw2Ny42NzYzNDIxIEw1MS4xMTIyODk5LDc3LjM3NjE0NCBMMzUuMDA3NzE4LDgwIFoiIGlkPSJQYXRoLTIyIiBmaWxsPSIjMDAyODY4Ij48L3BhdGg+CiAgICAgICAgICAgICAgICA8cGF0aCBkPSJNMCwzNy43MzA0NDA1IEwyNy4xMTQ1MzcsMC4yNTcxMTE0MzYgQzYyLjM3MTUxMjMsLTEuOTkwNzE3MDEgODAsMTAuNTAwMzkyNyA4MCwzNy43MzA0NDA1IEM4MCw2NC45NjA0ODgyIDY0Ljc3NjUwMzgsNzkuMDUwMzQxNCAzNC4zMjk1MTEzLDgwIEM0Ny4wNTUzNDg5LDc3LjU2NzA4MDggNTMuNDE4MjY3Nyw3MC4zMTM2MTAzIDUzLjQxODI2NzcsNTguMjM5NTg4NSBDNTMuNDE4MjY3Nyw0MC4xMjg1NTU3IDM2LjMwMzk1NDQsMzcuNzMwNDQwNSAyNS4yMjc0MTcsMzcuNzMwNDQwNSBDMTcuODQzMDU4NiwzNy43MzA0NDA1IDkuNDMzOTE5NjYsMzcuNzMwNDQwNSAwLDM3LjczMDQ0MDUgWiIgaWQ9IlBhdGgtMTkiIGZpbGw9IiMzNzkzRUYiPjwvcGF0aD4KICAgICAgICAgICAgPC9nPgogICAgICAgIDwvZz4KICAgIDwvZz4KPC9zdmc+' > </img>
/ Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| AnalysisData/Step03-PassingAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Intro to using Pandas - with healthcare for all data
# NumPy is a Python library used for working with arrays.
#
# It also has functions for working in domain of linear algebra, fourier transform, and matrices.
#
# NumPy was created in 2005 by <NAME>. It is an open source project and you can use it freely.
#
# NumPy stands for Numerical Python.
#step 1 import your libraries / packages - try running, if not installed use conda / pip to install
# eg conda install -c anaconda numpy or pip install numpy
import numpy as np
# +
#conda install -c anaconda numpy
# -
#should be included with anaconda installation
import pandas as pd
# +
#conda install -c anaconda pandas
# -
import seaborn as sns
# ### get data
#bring in the file and convert to a pandas dataframe
file1 = pd.read_csv('file1.csv')
#look at the top rows of the first file using df.head()
file1.head()
file1.shape
#use describe to review whats in the columns and some basic descriptive statistics
file1.describe(include = "all")
#bring in the next file
file2 = pd.read_csv('file2.txt',sep='\t')
# note the use of separators necessary for txt files, not for csv files
#look at the top rows of the second file using df.head()
file2.head()
#look at the shape of the second file using df.shape
file2.shape
# +
#read in and review the head/ shape of the remaining two excel files - this time using pd.read_excel, as file3 and file4
file3=pd.read_excel('file3.xlsx')
file3.head()
# -
file3.shape
file4=pd.read_excel('file4.xlsx')
file4.head()
file4.shape
# ### merge the data frames
#
# after reviewing the column headers for a match we will combine data sources 1 and 2
#lets check the column names for file 1 and 2 using df.columns
file3.columns
file4.columns
file1.columns
file2.columns
# +
# pull out the column names from file1 as a variable (
column_names=file1.columns
column_names
# +
#this command will set those column names in the target data frame
data=pd.DataFrame(columns=column_names)
# -
#use head() to review what we have
data.head()
# next lets concatenate our new target data frame with our first file1
data=pd.concat([data,file1],axis=0)
#use head() to review what we have
data.head()
# +
# same again, concat the file 2 into the data df.
data=pd.concat([data,file2],axis=0)
#hint : be careful not to run this more than once or you will have to clear some output!
# -
#check the shape to ensure you have the correct no of rows
data.shape
# +
#before bringing in the other dfs lets confirm the data columns will line up for all files
data.columns
#if they dont - what can be done?
# +
# if happy to proceed, lets concat in file 3 using the same method as before and review the head()
data=pd.concat([data,file3],axis=0)
# +
#lets concat in file 4 using the same method as before and review the head()
data=pd.concat([data,file4],axis=0)
data.shape
# -
# hint: we are doing this step by step- there are faster methods, to concat all files at once
#
# we could use something like:
# data = pd.concat([data,file2,file3, file4], axis=0)
#
#check the shape to ensure you have the correct no of rows and columns
# ### Standardizing header names
# Some standards are:
# use lower case
# if headers have spaces, replace them with underscores
#lets look at one column header (using the index position)
data.columns[1]
#how many columns do we have?
len(data.columns)
#we want to make all the columns into lower case
cols = []
for i in range(len(data.columns)):
cols.append(data.columns[i].lower())
cols
# +
#reset the columns from our function
data.columns = cols
#hint: we could also have used a STR function data.columns=data.columns.str.lower()
# +
#use head() to check the df
data.head()
# -
# ### examining the columns and looking for empty cells
#check the data types of all columns
data.dtypes
#lets look at how many NA values we have in each column
data.info()
# +
#we can easily create a df of missing values using isna()
missingdata=data.isna()
missingdata.head()
# +
#what about showing the % nulls - heres one technique
#the above data frame is a boolean ie 1s and 0s - so if we add them up we can see a count of actual values
missingdata.sum()/len(data)
#what does this look like from maths?
#Summing up all the values in a column and then dividing by the total number is the mean.
#this is the same as missingdata.mean()
# -
#to summarise this in one line of code and round the values
data.isna().mean().round(4) *100
# +
# lets assume we wanted to drop a single column due to poor coverage across all data sets: domain
data = data.drop(['domain'],axis=1)
# -
#what we are left with
data.columns
# +
#this time lets drop a few columns in one hit by choosing which we want to keep - also rearranging the columns
data=data[['controln', 'state', 'gender', 'hv1', 'hvp1',
'pobc1', 'pobc2','avggift', 'target_d','ic1','ic2','ic3','ic4','ic5']]
# -
data.head()
#lets rename a few of the columns to sensible names
data = data.rename(columns={ 'controln':'id','hv1':'median_home_val', 'ic1':'median_household_income'})
#review the data using head()
data.head()
file1.head()
# ### heres one i wish i had dun earlier
from IPython.display import Image
Image("face_palm.jpeg")
data.info()
# +
# - I noticed earlier when running data.info() that I had forgotten to reset the index
#after merging all the data frames! oops ... Lets do that now ....
data.reset_index(drop=True, inplace=True)
# -
data.tail()
# ### filtering and subsetting
# +
#method 1 focus on just male donors in florida
data[(data["state"]=="FL") & (data["gender"]=='M')]
# +
#alternative method :
data.query('gender=="M" & state=="FL" ')
# -
# last option
data.loc[data.gender == "M"]
# quick question - are we picking up all males in our data set?
data['gender'].value_counts()
data['gender'].unique()
# +
#challenge1: view a filtered subset of the data where the average gift is over 10 dollars
data[(data['avggift']>10)]
# +
#challenge 2: view a filtered subset of the data where the gender is M, state is florida and avg gift size is more than 10 dollars
data.query('gender=="M" & state=="FL" & avggift > 10')
# +
#before we apply any lasting filter to our data frame lets create a new version which we can play with
# so that the original data frame wont be affected
tempdata=data.copy()
# +
#use head(), .columns or shape() to review the tempdata
tempdata.head()
# -
#create a filtered dataframe from our M gender subset
filtered =data[data['gender']=='M']
#use shape for your filtered df
filtered.shape
#use head to review the top rows of the filtered df
filtered.head()
#notice we need to reset the index
filtered.reset_index(drop=True, inplace=True)
filtered.head()
#use the index to display the first ten rows
filtered[0:10]
# use the index to display 3 columns, first ten rows
filtered[['gender', 'ic2', 'ic3']][23:29]
#use the index to display rows with index number 1 and 2 (remember an index starts at 0)
filtered[1:3]
#use loc to return a selected row
filtered.loc[326]
# +
#use iloc to return first 10 rows and the first 4 columns
filtered.iloc[1:10,0:4]
# -
# loc v iloc
# loc is label based
# iloc is integer based only
# https://www.analyticsvidhya.com/blog/2020/02/loc-iloc-pandas/
# +
#sian typo! should refer to id not controln
# tip : its possible to set the index from any column eg CONTROLN:id
filtered2=filtered.reset_index().set_index('id')
filtered2.head()
#in which case using iloc and loc would give very different results
# -
filtered2.sort_index()
# +
#when reviewing subsets it is often smart to reconfigure how many rows you will see max
pd.set_option('display.max_rows', 20)
# -
filtered.loc[:218]
# +
#sian typo! should say filtered2 not filtered 3
filtered.iloc[:218]
# -
# ### data cleaning steps (1) data type change
#reverting back to our temp data copy
# lets look at the data types. any data type worth changing?
tempdata.dtypes
#focus on float data types
tempdata.select_dtypes('float64')
#simple change from float to int type would drop the decimals
tempdata['avggift'] = tempdata['avggift'].astype('int')
#focus on the object data types
tempdata.select_dtypes('object')
# although tempting to force an object into a float using as type :
#
# tempdata['median_home_val'] = tempdata['median_home_val'].astype('float', errors='ignore')
#
# we know some values of the data are strings and this could produce an error
# +
#lets have a look at all the numeric data
tempdata._get_numeric_data()
# +
#do a data type change for column and replace non float values with NaN
tempdata['median_home_val'] = pd.to_numeric(tempdata['median_home_val'], errors='coerce')
# -
# do the same for median household income, ic3 and ic5
tempdata['median_household_income'] = pd.to_numeric(tempdata['median_household_income'], errors='coerce')
tempdata['ic3'] = pd.to_numeric(tempdata['ic3'], errors='coerce')
tempdata['ic5'] = pd.to_numeric(tempdata['ic5'], errors='coerce')
tempdata.dtypes
# +
tempdata.to_csv('tempdataoutput.csv', index = False)
# -
# ### data cleaning steps (2) duplicates
#lets rename our tempdata df into cleandata
cleandata =tempdata
# +
#drop all dupes using df.drop_duplicates()- remember to replace the dataframe with the de duped df
cleandata=cleandata.drop_duplicates()
#we can also do a de dupe on specified columns such as:
#cleandata = cleandata.drop_duplicates(subset=['state','gender', 'ic2', 'ic3'])
# -
#review using shape
cleandata.shape
cleandata.head()
cleandata.reset_index(drop=True, inplace=True)
# ### data cleaning steps (3) null values
# review where are we starting from - use info()
cleandata.info()
#for a more detailed snapshot, we can calculate the % nulls per column as a new df
nulls_df = pd.DataFrame(round(cleandata.isna().sum()/len(cleandata),4)*100)
nulls_df
# +
#tidy the nulls_df by resetting the index and renaming the columns as 'header_name' & 'percent_nulls'
nulls_df=nulls_df.reset_index()
nulls_df = nulls_df.rename(columns={'index':'header_name',0:'percent_nulls'})
nulls_df
# +
#nulls_df.reset_index(inplace=True)
# -
# #### categorical data nulls
#we could potentially drop columns we see as having high null % or drop columns according to a null% rule
columns_drop = nulls_df[nulls_df['percent_nulls']>3]['header_name'] #3% threshold, arbitrary value
print(columns_drop.values)
#example dropping the gender column on a new dataframe cleandata1- just an example, we wont do this with our df:
cleandata1 = cleandata.drop(['gender'], axis=1))
# +
#what if i were to drop just those rows with empty gender ? how many rows would this effect?
cleandata[cleandata['gender'].isna()==True]
# +
#alternative - replace the nulls with a sensible value. one option is use the most common value
#step 1 find the most common gender value in the data set with the value counts()
cleandata['gender'].value_counts()
# -
#step 2 fillna with most freq value using fillna
cleandata['gender'] = cleandata['gender'].fillna('F')
#run the nulls % data frame creation steps again to see what your cleandata data frame looks like now
# +
# options for handling null values in categorical variables
# Ignore observation
# Replace by most frequent value
# Replace using an algorithm like KNN using the neighbours.
# Predict the observation using a multiclass predictor.
# Treat missing data as just another category
# -
# #### numerical data nulls
#run nulls_df if you need a reminder of how many nulls we have
cleandata.info()
#if some columns contain relatively few null values in our data we can filter the null rows away (drop them).
cleandata = cleandata[cleandata['ic2'].isna()==False]
# +
#do the same for ic4 and ic5
cleandata = cleandata[cleandata['ic5'].isna()==False]
# -
np.mean(cleandata['median_home_val'])
#with columns that have more nulls we will use an impute method, for example using the mean value
mean_median_home_value = np.mean(cleandata['median_home_val'])
#use fillna to complete the step
cleandata['median_home_val'] = cleandata['median_home_val'].fillna(mean_median_home_value)
# +
#options for null values in Numerical columns:
# Ignore these observations
# Replace with general average
# Replace with similar type of averages
# Build model to predict missing values
# -
#output to csv file
cleandata.to_csv('checkpoint2.csv')
# ### data cleaning steps (4) too many unique (categorical) values
#how many unique values do we have in the gender column?
cleandata['gender'].unique()
# +
#introducing lamba and map to solve our gender issue
#A lambda function is a small anonymous function.
#A lambda function can take any number of arguments, but can only have one expression.
# -
# #### some simple maths examples
y = lambda x: x+2
#what would we get with
y(200)
addition = lambda x,y : x+y
addition(1,3)
square = lambda x: x*x
square(24)
lst = [1,2,3,4,5,6,7,8,9,10]
new_list = []
for item in lst:
new_list.append(square(item))
new_list
#we could also have used list comprehension
new_list = [square(item) for item in lst]
new_list
# +
#apply lambda from above to square only the even numbers in our lst
new_list = []
for item in lst:
if item %2==0:
new_list.append(square(item))
new_list
# -
# https://www.w3schools.com/python/python_lambda.asp
# +
# map function
# The map() function executes a specified function for each item in an iterable. The item is sent to the function as a parameter.
def myfunc(n):
return len(n)
x = map(myfunc, ('apple', 'banana', 'cherry'))
print(list(x))
# -
#heres a simple map/lambda combination to make all gender labels upper case
cleandata['gender'] = list(map(lambda x: x.upper(), cleandata['gender']))
cleandata['gender'].unique()
# +
#lambda is good for quick jobs like lower/upper STR functions that you dont need much transparency over
#for more complicated tasks, create a function to match the unique values to M or F using IF logic
def clean(x):
if x in ['M', 'MALE']:
return 'Male'
elif x.startswith('F'):
return 'Female'
else:
return 'U'
# +
#apply the above function with map to create a smaller set of gender labels in our cleandata
#- hint the function clean should be in place of lambda as shown in the last map
cleandata['gender']= list(map(clean,cleandata['gender']))
#syntax structure : df['column']=list(map(functionname, df['column']))
# +
#use value_counts() or df['column'].unique() to confirm the effect of what you just did
# -
# #### OPTION - create buckets to solve too many unique values
#
# this depends on the business case!
#create buckets on the ic2 data based on cutting the data range into 4 sections.
ic2_labels = ['Low', 'Moderate', 'High', 'Very High']
cleandata['ic2_'] = pd.cut(cleandata['ic2'],4, labels=ic2_labels)
# +
#lets check what the bins look like
pd.cut(cleandata['ic2'],4)
# +
#alternatively using quantiles to cut the data could be helpful
#pd.qcut = “Quantile-based discretization function.”
#This basically means that qcut tries to divide up the underlying data into equal sized bins.
#The function defines the bins using percentiles based on the distribution of the data, not the actual numeric edges of the bins
# -
#lets look at another candidate column for this with describe
cleandata['ic3'].describe()
# +
#and with value_counts() can we get an idea of how unique each value is
# +
#cut up ic3 into 4 quantiles - let Python work out the rest
pd.qcut(cleandata['ic3'], q=4)
# +
#A common use case is to store the bin results back in the original dataframe for future analysis.
#For this example, we will create 4 bins (aka quartiles) and store the results back in the original dataframe:
cleandata['quantile_ic3'] = pd.qcut(cleandata['ic3'], q=4)
cleandata.head()
# +
#use value_counts on the new quantile ic3 field to see how the split
cleandata['quantile_ic3'].value_counts()
# -
#a clear disadvantage - not super easy to interpret to the end user and requires relabelling at some point.
# But for transparency lets leave it as is - for more information on this: https://pbpython.com/pandas-qcut-cut.html
# ### finally - lets export what we have so far to a csv
# Exporting this processed cleandata to a csv
cleandata.to_csv('merged_clean_ver1.csv', index = False)
| class_1/pandas intro 2 with annotation-students-wip.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dictionary example
dict = {1:"Maryam","Lastname":"sheifu","age":25}
# prints the value where key value is 1
print(dict[1])
#prints the value where key value is "lastname"
print(dict["Lastname"])
# prints the vale where key value is "age"
print(dict["age"])
# -
| PYTHON PRACTICE/PYTHON-DATA-TYPE-Dictionary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Generating Specific Handwritten Digit Using CGAN
# We just learned how CGAN works and the architecture of CGAN. To strengthen our understanding, now we will learn how to implement CGAN in TensorFlow for generating an image of specific handwritten digit.
# ## Import libraries
#
# First, we will import all the necessary libraries:
# +
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
tf.logging.set_verbosity(tf.logging.ERROR)
tf.reset_default_graph()
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython import display
# -
# ### Read the Dataset
#
# Load the MNIST dataset:
data = input_data.read_data_sets("data/mnist",one_hot=True)
# ## Defining Generator
#
# Generator $G$ takes the noise $z$ and also the conditional variable $c$ as an input and returns an image. We define the generator as a simple 2 layer feed forward network.
def generator(z, c,reuse=False):
with tf.variable_scope('generator', reuse=reuse):
#initialize weights
w_init = tf.contrib.layers.xavier_initializer()
#concatenate noize z and conditional variable c to form an input
inputs = tf.concat([z, c], 1)
#define the first layer with relu activation
dense1 = tf.layers.dense(inputs, 128, kernel_initializer=w_init)
relu1 = tf.nn.relu(dense1)
#define the second layer and compute the output using the tanh activation function
logits = tf.layers.dense(relu1, 784, kernel_initializer=w_init)
output = tf.nn.tanh(logits)
return output
# ## Defining Discriminator
#
# We know that discriminator $D$ returns the probability. i.e it will tell us the probability of the given image being real. Along with the input image $x$, it also takes the conditional varaible $c$ as an input. We define the discriminator also as a simple 2 layer feed forward network.
def discriminator(x, c, reuse=False):
with tf.variable_scope('discriminator', reuse=reuse):
#initialize weights
w_init = tf.contrib.layers.xavier_initializer()
#concatenate noize z and conditional variable c to form an input
inputs = tf.concat([x, c], 1)
#define the first layer with the relu activation
dense1 = tf.layers.dense(inputs, 128, kernel_initializer=w_init)
relu1 = tf.nn.relu(dense1)
#define the second layer and compute the output using sigmoid activation function
logits = tf.layers.dense(relu1, 1, kernel_initializer=w_init)
output = tf.nn.sigmoid(logits)
return logits
# ## Define the input placeholders
#
#
# Now, we define the placeholder for the input $x$, conditional variable $c$ and the noise $z$:
x = tf.placeholder(tf.float32, shape=(None, 784))
c = tf.placeholder(tf.float32, shape=(None, 10))
z = tf.placeholder(tf.float32, shape=(None, 100))
# ## Start the GAN!
# First we feed the noise z to the generator and it will output the fake image. i.e $ fake \; x = G(z|c) $
fake_x = generator(z, c)
# Now, we feed the real image $x$ along with conditional variable $c$ to the discriminator $D(x|c)$ and get the probabillty of being real:
D_logits_real = discriminator(x,c)
# Similarly, we feed the fake image fake_x and conditional variable $c$ to the discirminator $D(z|c)$ and get the probabillty of it being real:
D_logits_fake = discriminator(fake_x, c, reuse=True)
# ## Computing Loss Function
#
# Now, we will see, how to compute the loss function. It is essentially same as vanilla GAN except that we add a condtional variable.
#
#
#
# ### Discriminator Loss
#
# Discriminator loss is given as,
#
# ${L ^{D} = - \mathbb{E}_{x \sim p_{r}(x)}[\log D(x|c)] - \mathbb{E}_{z \sim p_{z}(z)}[\log (1-D(G(z|c))]} $
#
#
# <br>
#
#
# First we will implement the first term i.e $\mathbb{E}_{x \sim p_{r}(x)}[\log D(x|c)]$
D_loss_real = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=D_logits_real,
labels=tf.ones_like(D_logits_real)))
# Now the second term, $\mathbb{E}_{z \sim p_{z}(z)}[\log (1-D(G(z|c))]$
D_loss_fake = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=D_logits_fake,
labels=tf.zeros_like(D_logits_fake)))
# The final loss can be written as:
D_loss = D_loss_real + D_loss_fake
# ### Generator Loss
#
# Generator loss is given as:
#
# ${L^{G}= - \mathbb{E}_{z \sim p_{z}(z)}[\log (D(G(z|c)))] } $
G_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=D_logits_fake,
labels=tf.ones_like(D_logits_fake)))
# ## Optimizing the Loss
#
#
# Now we need to optimize our generator and discriminator. So, we collect the parameters of the discriminator and generator as $\theta_D$ and $\theta_G$ respectively.
training_vars = tf.trainable_variables()
theta_D = [var for var in training_vars if var.name.startswith('discriminator')]
theta_G = [var for var in training_vars if var.name.startswith('generator')]
# Optimize the loss using adam optimizer:
D_optimizer = tf.train.AdamOptimizer(0.001, beta1=0.5).minimize(D_loss, var_list=theta_D)
G_optimizer = tf.train.AdamOptimizer(0.001, beta1=0.5).minimize(G_loss, var_list=theta_G)
# ## Start the Training
#
#
# Start the TensorFlow session and initialize variables:
#
session = tf.InteractiveSession()
tf.global_variables_initializer().run()
# Define the batch size, number of epochs and number of classes:
batch_size = 128
num_epochs = 5000
num_classes = 10
# Define the images and labels:
images = data.train.images
labels = data.train.labels
# ## Generate the Handwritten Digit 7
# We set the digit to generate as 7:
label_to_generate = 7
onehot = np.eye(10)
for epoch in range(num_epochs):
for i in range(len(images) // batch_size):
#sample images
batch_image = images[i * batch_size:(i + 1) * batch_size]
#sample the condition that is, digit we want to generate
batch_c = labels[i * batch_size:(i + 1) * batch_size]
#sample noise
batch_noise = np.random.normal(0, 1, (batch_size, 100))
#train the generator
generator_loss, _ = session.run([D_loss, D_optimizer], {x: batch_image, c: batch_c, z: batch_noise})
#train the discriminator
discriminator_loss, _ = session.run([G_loss, G_optimizer], {x: batch_image, c: batch_c, z: batch_noise})
#sample noise
noise = np.random.rand(1,100)
#select specific digit
gen_label = np.array([[label_to_generate]]).reshape(-1)
#convert the selected digit
one_hot_targets = np.eye(num_classes)[gen_label]
#Feed the noise and one hot encoded condition to the generator and generate the fake image
_fake_x = session.run(fake_x, {z: noise, c: one_hot_targets})
_fake_x = _fake_x.reshape(28,28)
print("Epoch: {},Discriminator Loss:{}, Generator Loss: {}".format(epoch,discriminator_loss,generator_loss))
#plot the generated image
display.clear_output(wait=True)
plt.imshow(_fake_x)
plt.show()
# Thus, with CGAN we can generate a speific image that we want to generate. In the next section, we will learn about InfoGAN which is the unsupervised version of CGAN.
| Chapter09/9.02 Generating Specific Handwritten Digit Using CGAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import gmaps
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import csv
gmaps.configure(api_key="API") # Your Google API key, personal information
results=[]
# put locations_top_100_cites.csv in the same dic
with open("locations_top_100_cites_3.csv") as csvfile: # data of locations of top 100 cites
reader = csv.reader(csvfile, quoting=csv.QUOTE_NONNUMERIC) # change contents to floats
next(reader, None) # skip the headers
for row in reader: # each row is a list
results.append(row)
# change the list to array
data=np.array(results)
locations=[[row[0], row[1]] for row in data]
weights=lat=[row[2] for row in data]
#print(locations)
fig=gmaps.figure(layout={
'width': '700px',
'height': '500px',
'padding': '3px',
'border': '1px solid black'
})
fig.add_layer(gmaps.heatmap_layer(locations,weights,point_radius=12))
fig
# -
| Heatmap_of _top_100_cites_in_Vermont_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was used to generate the inputs and expected outputs for fir_test
import sys
sys.path.append('..')
import numpy as np
import plotly.graph_objects as go
from rtmha.filter import FirFilter
from rtmha.elevenband import elevenband_taps_min
def plot_res(x,y, res):
ms = np.linspace(0,10,len(x))
fig = go.Figure()
fig.add_trace(go.Scatter(x=ms, y=y, name='input'))
fig.add_trace(go.Scatter(x=ms, y=res, name='output'))
fig.update_layout(xaxis_title='milliseconds',
yaxis_title='Amplitude',
template='plotly_dark')
fig.show()
# this is the band 0 filter, so the rate is downsampled to 1/16
sample_rate = 32000
down_rate=sample_rate / 16
nyq_rate=down_rate / 2
def generate_sine_waves(freq_list, duration=1, sample_rate=32000):
"""Generates a signal with multiple sine waves
Args:
freq_list: List of frequencies
duration: signal length in seconds (default 1)
sample_rate: sample rate in Hz (default 32000)
Returns:
(t, y): t is time. y is value in range [-1,1]
"""
x = np.linspace(0, duration, int(sample_rate * duration), endpoint=False)
y = np.zeros(len(x))
for freq in freq_list:
frequencies = x * freq
y += np.sin((2 * np.pi) * frequencies)
y /= len(freq_list) # normalize
return x, y
x, y = generate_sine_waves([200, 4000], duration=32*10/down_rate, sample_rate=down_rate)
f = FirFilter(elevenband_taps_min[0], len(y))
res = f.filter(y)
plot_res(x,y,res)
from scipy.signal import lfilter
out = lfilter(elevenband_taps_min[0], 1.0, y)
plot_res(x, y, out)
np.allclose(out, res)
f = FirFilter(elevenband_taps_min[10], len(y))
res = f.filter(y)
plot_res(x,y,res)
inp = np.zeros(128).astype('float32')
inp[1]=1
f = FirFilter(elevenband_taps_min[0], 128)
res = f.filter(inp)
plot_res(inp, inp,res)
# %%timeit
res = f.filter(inp)
# %%timeit
out = lfilter(elevenband_taps_min[0], 1.0, inp)
out = lfilter(elevenband_taps_min[0], 1.0, inp)
plot_res(inp, inp, out)
f = FirFilter(elevenband_taps_min[10], len(inp))
res = f.filter(inp)
plot_res(inp, inp,res)
down_rate = 32000
x, y = generate_sine_waves([8000], duration=32*4/down_rate, sample_rate=down_rate)
f = FirFilter(elevenband_taps_min[10], len(y))
res = f.filter(y)
plot_res(y, y,res)
| Sources/libosp/python/test/Fir_Filter-MakeTestData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import re
def remove_html(text):
cleanr = re.compile('<.*?>')
cleantext = re.sub(cleanr, '', text)
return cleantext
import nltk.tokenize as tk
def sentence_tokenize(text):
sentences = tk.sent_tokenize(text)
return len(sentences), sentences
def word_tokenize(text):
words = tk.word_tokenize(text)
return len(words), words
# +
# Call on concatenation of body and title
from collections import namedtuple
tokenized_row = namedtuple('tokenized_row', 'sent_count sentences word_count words')
def convert_row(text):
text = remove_html(text)
sent_count, sentences = sentence_tokenize(text)
word_count, words = word_tokenize(text)
return tokenized_row(sent_count, sentences, word_count, words)
def build_dict(dataframe):
token_dict = {}
body_words = []
title_words = []
for i in range(len(dataframe.index.values)):
index = dataframe.index.values[i]
title = convert_row(dataframe['Title'].values[i])
title_words = title_words + title.words
body = convert_row(dataframe['Body'].values[i])
body_words = body_words + body.words
token_dict[index] = (title, body)
return token_dict, title_words, body_words
# -
import pickle
import pandas
import os
from sklearn.feature_extraction.text import CountVectorizer
filenames = ['combined_train_test.p', 'r_train_so_test.p', 'so_train_r_test.p',
'so_alone.p', 'reddit_alone.p']
for filename in filenames:
directory_name = filename.split('.p')[0]
if not os.path.isdir(directory_name):
os.mkdir(directory_name)
with open(filename, 'rb') as pfile:
train, test = pickle.load(pfile)
body_vectorizer = CountVectorizer(stop_words='english', max_features = 2**12)
title_vectorizer = CountVectorizer(stop_words='english', max_features = 2**12)
train_token_dict, train_title_words, train_body_words = build_dict(train)
test_token_dict, test_title_words, test_body_words = build_dict(test)
body_vectorizer.fit((train_body_words + test_body_words))
title_vectorizer.fit((train_title_words + test_title_words))
with open(directory_name + "/tokenized_dict.p", 'wb') as pfile:
pickle.dump((train_token_dict, test_token_dict), pfile)
with open(directory_name + "/body_vectorizer.p", 'wb') as pfile:
pickle.dump(body_vectorizer, pfile)
with open(directory_name + "/title_vectorizer.p", 'wb') as pfile:
pickle.dump(title_vectorizer, pfile)
| Tokenize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import os
import numpy as np
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.decomposition import PCA
from sklearn.metrics import log_loss
from sklearn.pipeline import Pipeline
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.model_selection import train_test_split, cross_val_predict, cross_validate, cross_val_score
# reading data files and storing them in a dataframe
df_train_features = pd.read_csv('D:/Fundamental of AI/lish-moa/train_features.csv')
df_test_features = pd.read_csv('D:/Fundamental of AI/lish-moa/test_features.csv')
df_train_target_nonscored = pd.read_csv('D:/Fundamental of AI/lish-moa/train_targets_nonscored.csv')
df_train_target_scored = pd.read_csv('D:/Fundamental of AI/lish-moa/train_targets_scored.csv')
# Feature Extraction
ctl_vehicle_ids = (df_train_features['cp_type'] == 'ctl_vehicle')
cp_cat_cols = ['cp_type', 'cp_time', 'cp_dose']
# select all indices when 'cp_type' is 'ctl_vehicle'
ctl_vehicle_cols = (df_train_features['cp_type'] == 'ctl_vehicle')
# take a copy of all our training sig_ids for reference
train_sig_ids = df_train_features['sig_id'].copy()
# drop cp_type column since we no longer need it
X = df_train_features.drop(['sig_id', 'cp_type'], axis=1).copy()
X = X.loc[~ctl_vehicle_ids].copy()
y = df_train_target_scored.drop('sig_id', axis=1).copy()
y = y.loc[~ctl_vehicle_ids].copy()
# Data pre processing
class data_pre_processor(BaseEstimator, TransformerMixin):
# data prep processing and data loading into the function
def __init__(self, remove_ctl_vehicles=True, std_features=True, encode_cat=True,
cp_cat_cols=['cp_time', 'cp_dose']):
self.remove_ctl_vehicles = remove_ctl_vehicles
self.std_features = std_features
self.cp_cat_cols = cp_cat_cols
self.encode_cat = encode_cat
def fit(self, X, y=None):
return self
def transform(self, X):
temp_df = self._remove_features(X).copy()
if self.std_features:
pass
if self.encode_cat:
temp_df = pd.concat([pd.get_dummies(temp_df.cp_dose, prefix='cp_dose'),
temp_df.drop('cp_dose', axis=1)], axis=1)
temp_df = pd.concat([pd.get_dummies(temp_df.cp_time, prefix='cp_time'),
temp_df.drop('cp_time', axis=1)], axis=1)
return temp_df
def _remove_features(self, dataframe):
if self.remove_ctl_vehicles:
temp_df = dataframe.drop(['sig_id', 'cp_type'], axis=1)
else:
temp_df = dataframe.drop('sig_id', axis=1)
return temp_df
def _standardise_features(self, dataframe):
pass
# features extraction continued
train_sig_ids = df_train_features['sig_id'].copy()
# select all indices when 'cp_type' is 'ctl_vehicle'
train_ctl_vehicle_ids = (df_train_features['cp_type'] == 'ctl_vehicle')
# initialization of data processing class for transforming dataset
data_pre_processor = data_pre_processor()
# dropping low correlated features and which are not required and encode of categorical features from feature and target variables
X = data_pre_processor.fit_transform(df_train_features)
y = df_train_target_scored.drop('sig_id', axis=1).copy()
# dropping the rows where cp_type is ctl_vehicle from training features and target sets
X = X.loc[~train_ctl_vehicle_ids].copy()
y = df_train_target_scored.drop('sig_id', axis=1).copy()
y = y.loc[~train_ctl_vehicle_ids].copy()
# standardisation of numerical columns
std_scaler = StandardScaler()
num_cols = [x for x in X.columns.values if not x.startswith(('cp_time', 'cp_dose'))]
X_std = X.copy()
X_std[num_cols] = std_scaler.fit_transform(X.loc[:, num_cols])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
# Model Building, loss calcuation and prediction of values
lin_reg = LinearRegression()
lin_reg = LinearRegression()
lr_val_preds = cross_val_predict(lin_reg, X_std, y, cv=5)
lr_log_loss = log_loss(np.ravel(y), np.ravel(lr_val_preds))
print(f"Log loss for Linear Regression Model: {lr_log_loss:.5f}\n")
# +
# principle component analysis and calculating loss for each iteration
n_range = [1, 2, 5, 10, 25, 50, 100, 150, 200, 250]
log_losses = []
lin_reg = LinearRegression()
for n in n_range:
pca = PCA(n_components=n)
lr_model = Pipeline(steps=[('pca', pca), ('linear regression', lin_reg)])
# evaluate of model using cross validation
lr_val_preds = cross_val_predict(lr_model, X_std, y, cv=5)
# we need to flatten both arrays before computing log loss
lr_log_loss = log_loss(np.ravel(y), np.ravel(lr_val_preds))
print(f"Log loss for Linear Regression with PCA (n={n}): {lr_log_loss:.5f}\n")
log_losses.append(lr_log_loss)
# -
# Performance of model visualization using seaborn library
import matplotlib.pyplot as plt
plt.figure(figsize=(14,5))
sns.lineplot(x=n_range, y=log_losses)
plt.ylabel("Average Logarithmic Loss", weight='bold')
plt.xlabel("Principle component analysis components count", weight='bold')
plt.grid()
plt.show()
# using model to make predictions on test data set
# filtering sig ids for splitting feature and target variables
test_sig_ids = df_test_features['sig_id'].copy()
# filtering the rows in test dataset where 'cp_type' is 'ctl_vehicle'
test_dataset_ctl_vehicle_ids = (df_test_features['cp_type'] == 'ctl_vehicle')
X_test = data_pre_processor.transform(df_test_features)
# Sclaing of test dataset
test_cols = [x for x in X_test.columns.values if not x.startswith(('cp_time', 'cp_dose'))]
X_test_standardization = X_test.copy()
X_test_standardization[test_cols] = std_scaler.transform(X_test.loc[:, test_cols])
# fitting model on test dataset
lin_reg = LinearRegression()
pca = PCA(n_components=5)
lr_model = Pipeline(steps=[('pca', pca), ('linear regression', lin_reg)])
# %time lr_model.fit(X_std, y)
# final predicted values on test dataset
test_prediction = lr_model.predict(X_test_standardization)
test_prediction[test_sig_ids[test_dataset_ctl_vehicle_ids].index.values]
#plotting qq plot
import statsmodels.api as sm
sm.qqplot(test_prediction,fit=True,line='45')
plt.show()
import pandas as pd
import numpy as np
from sklearn.model_selection import KFold
from sklearn.multioutput import MultiOutputClassifier
from sklearn.linear_model import LogisticRegression
from joblib import dump, load
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import log_loss
from datetime import date
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
# +
# random seed value
xseed = 43
# number of folds for cv
nfolds = 5
# number of components from PCA decomposition
nof_comp = 250
model_name = 'lr'
X_train = pd.read_csv('D:/Fundamental of AI/lish-moa/train_features.csv')
X_test = pd.read_csv('D:/Fundamental of AI/lish-moa/test_features.csv')
y_train = pd.read_csv('D:/Fundamental of AI/lish-moa/train_targets_scored.csv')
# +
# data pre processing
print(set(X_train['cp_time']), set(X_test['cp_time']) )
# cp_time processing
X_train['cp_time_24'] = (X_train['cp_time'] == 24) + 0
X_train['cp_time_48'] = (X_train['cp_time'] == 48) + 0
X_test['cp_time_24'] = (X_test['cp_time'] == 24) + 0
X_test['cp_time_48'] = (X_test['cp_time'] == 48) + 0
X_train.drop('cp_time', axis = 1, inplace = True)
X_test.drop('cp_time', axis = 1, inplace = True)
# cp_dose processing
print(set(X_train['cp_dose']), set(X_test['cp_dose']) )
X_train['cp_dose_D1'] = (X_train['cp_dose'] == 'D1') + 0
X_test['cp_dose_D1'] = (X_test['cp_dose'] == 'D1') + 0
X_train.drop('cp_dose', axis = 1, inplace = True)
X_test.drop('cp_dose', axis = 1, inplace = True)
# cp_type processing
X_train['cp_type_control'] = (X_train['cp_type'] == 'ctl_vehicle') + 0
X_test['cp_type_control'] = (X_test['cp_type'] == 'ctl_vehicle') + 0
X_train.drop('cp_type', axis = 1, inplace = True)
X_test.drop('cp_type', axis = 1, inplace = True)
# +
# prepare split for the k fold
kf = KFold(n_splits = nfolds)
# separation of train and test
id_train = X_train['sig_id']
id_test = X_test['sig_id']
y_train.drop('sig_id', axis = 1, inplace = True)
X_train.drop('sig_id', axis = 1, inplace = True)
X_test.drop('sig_id', axis = 1, inplace = True)
# storage matrices for test predictions
prval = np.zeros(y_train.shape)
prfull = np.zeros((X_test.shape[0], y_train.shape[1]))
# +
# base model defining
pca = PCA(n_components = nof_comp)
logistic = LogisticRegression(max_iter=10000, tol=0.1, C = 0.5)
base_model = Pipeline(steps=[('pca', pca), ('logistic', logistic)])
# a pipeline can be fed into MultiOutputClassifier just like a regular estimator would
mo_base = MultiOutputClassifier(base_model, n_jobs=-1)
# -
for (ff, (id0, id1)) in enumerate(kf.split(X_train)):
x0, x1 = X_train.loc[id0], X_train.loc[id1]
y0, y1 = np.array(y_train.loc[id0]), np.array(y_train.loc[id1])
check_for_empty_cols = np.where(y0.sum(axis = 0) == 0)[0]
if len(check_for_empty_cols):
y0[0,check_for_empty_cols] = 1
# fit model
mo_base.fit(x0,y0)
# generate the prediction
vpred = mo_base.predict_proba(x1)
fpred = mo_base.predict_proba(X_test)
for ii in range(0,y_train.shape[1]):
prval[id1,ii] = vpred[ii][:,1]
prfull[:,ii] += fpred[ii][:,1]/nfolds
# +
prval = pd.DataFrame(prval); prval.columns = y_train.columns
prval['sig_id'] = id_train
prfull = pd.DataFrame(prfull); prfull.columns = y_train.columns
prfull['sig_id'] = id_test
# -
metrics = []
for _target in y_train.columns:
metrics.append(log_loss(y_train.loc[:, _target], prval.loc[:, _target]))
print(f'OOF Metric: {np.round(np.mean(metrics),4)}')
#plotting qq plot for logistic
import statsmodels.api as sm
sm.qqplot(prfull,fit=True,line='45')
plt.show()
| EAI 6000 - FoAI/Discussions & Assignments/Week 2 - Funadamentals of AI Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Off-policy Ensemble
# +
#load infowave data
import pickle, random
import numpy as np
y_sparse = pickle.load(open('all_question_article_ratings.pkl','rb'))
y = np.zeros((996,321))
for i,j in y_sparse:
y[i,j] = 1
X = pickle.load(open('question_tfidf_bow.pickle','rb'))
X = X.todense()
# -
from sklearn.linear_model import LogisticRegression
K_learners = 50
# +
st_seed = 0
end_seed = 300
st_exploration = 0
end_exploration = 800
st_test = 800
end_test = 996
explore_list = random.sample(range(996),end_exploration)
seed_list = random.sample(explore_list,end_seed)
test_list = np.setdiff1d(range(996), explore_list)
# rearrange the data
X =np.vstack((X[explore_list,:], X[test_list,:]))
y= np.vstack((y[explore_list,:], y[test_list,:]))
# +
Xseed = X[st_seed:end_seed, :]
Xexplore_sample = X[st_exploration:end_exploration, :]
yexplore_sample = y[st_exploration:end_exploration, :]
Xtest = X[st_test:end_test, :]
ytest = y[st_test:end_test, :]
nchoices = y.shape[1]
actions_ensemble = np.zeros([K_learners, end_exploration-st_exploration], dtype=int)
rewards_ensemble = np.zeros([K_learners, end_exploration-st_exploration])
prob_actions_ensemble = np.zeros([K_learners, end_exploration-st_exploration])
# Xseed=X[seed_list,:]
# Xexplore_sample=X[explore_list,:]
# Xtest=X[test_list,:]
# nchoices=y.shape[1]
# actions_ensemble = np.zeros([K_learners, len(explore_list)], dtype=int)
# rewards_ensemble = np.zeros([K_learners, len(explore_list)])
# prob_actions_ensemble = np.zeros([K_learners, len(explore_list)])
# -
for k in range(K_learners):
sample_index = np.random.randint(st_seed, end_seed, size=3000)
Xseed_sample = Xseed[sample_index, :]
explorer = LogisticRegression(solver="lbfgs", max_iter=1500, multi_class='auto')
explorer.fit(Xseed_sample, np.argmax(y[sample_index], axis=1))
actions_ensemble[k] = explorer.predict(Xexplore_sample)
rewards_ensemble[k] = y[st_exploration:end_exploration, :]\
[np.arange(end_exploration - st_exploration), actions_ensemble[k]]
ix_internal_actions = {j:i for i,j in enumerate(explorer.classes_)}
ix_internal_actions = [ix_internal_actions[i] for i in actions_ensemble[k]]
ix_internal_actions = np.array(ix_internal_actions)
prob_actions_ensemble[k] = explorer.predict_proba(Xexplore_sample)[np.arange(Xexplore_sample.shape[0]),
ix_internal_actions]
from contextualbandits.offpolicy import OffsetTree
from sklearn.linear_model import LogisticRegression
reward_ensemble = np.empty([K_learners, ])
# +
for k in range(K_learners):
new_policy = OffsetTree(base_algorithm=LogisticRegression(solver="lbfgs", max_iter=15000), nchoices=y.shape[1])
new_policy.fit(X=Xexplore_sample, a=actions_ensemble[k], r=rewards_ensemble[k], p=prob_actions_ensemble[k])
mean_reward_ot = np.mean(y[st_test:end_test, :][np.arange(end_test - st_test), new_policy.predict(Xtest)])
reward_ensemble[k] = mean_reward_ot
# -
from collections import Counter
# +
ensemble_predict = np.zeros([K_learners, 996-end_exploration])
vote = np.zeros([end_test-st_test, ], dtype = int)
for k in range(K_learners):
new_policy = OffsetTree(base_algorithm=LogisticRegression(solver="lbfgs", max_iter=1500), nchoices=y.shape[1])
new_policy.fit(X=Xexplore_sample, a=actions_ensemble[k], r=rewards_ensemble[k], p=prob_actions_ensemble[k])
ensemble_predict[k] = new_policy.predict(Xtest)
for i in range(end_test-st_test):
c = Counter(ensemble_predict[:, i])
mode_count = max(c.values())
mode = [key for key, count in c.items() if count == mode_count]
vote[i] = mode[0]
mean_reward_ot_ensemble = np.mean(y[st_test:end_test, :][np.arange(end_test - st_test), vote])
print("Test set mean reward - Offset Tree Ensemble technique: ", mean_reward_ot_ensemble)
# -
from sklearn.metrics import accuracy_score
yt = np.argmax(ytest, axis=1)
yr = np.argmax(yexplore_sample, axis=1)
accuracy_score(yt,vote)
# # Off-policy Methods Comparison
# +
from sklearn.linear_model import LogisticRegression
# separating the covariates data for each case
Xseed = X[st_seed:end_seed, :]
Xexplore_sample = X[st_exploration:end_exploration, :]
Xtest = X[st_test:end_test, :]
nchoices = y.shape[1]
# now constructing an exploration policy as explained above, with fully-labeled data
explorer = LogisticRegression(solver="lbfgs", max_iter=15000, multi_class='auto')
explorer.fit(Xseed, np.argmax(y[st_seed:end_seed], axis=1))
# letting the exploration policy choose actions for the new policy input
actions_explore_sample = explorer.predict(Xexplore_sample)
rewards_explore_sample = y[st_exploration:end_exploration, :]\
[np.arange(end_exploration - st_exploration), actions_explore_sample]
# extracting the probabilities it estimated
ix_internal_actions = {j:i for i,j in enumerate(explorer.classes_)}
ix_internal_actions = [ix_internal_actions[i] for i in actions_explore_sample]
ix_internal_actions = np.array(ix_internal_actions)
prob_actions_explore = explorer.predict_proba(Xexplore_sample)[np.arange(Xexplore_sample.shape[0]),
ix_internal_actions]
# +
from contextualbandits.online import SeparateClassifiers
from sklearn.linear_model import LogisticRegression
new_policy = SeparateClassifiers(base_algorithm=LogisticRegression(solver="lbfgs", max_iter=15000),
nchoices=y.shape[1], beta_prior=None, smoothing=None)
new_policy.fit(X=Xexplore_sample, a=actions_explore_sample, r=rewards_explore_sample)
mean_reward_naive = np.mean(y[st_test:end_test, :]\
[np.arange(end_test - st_test), new_policy.predict(Xtest)])
print("Test set mean reward - Separate Classifiers: ", mean_reward_naive)
yacc_naive=accuracy_score(new_policy.predict(Xtest), yt)
yacc_naive=accuracy_score(new_policy.predict(Xtest), yt)
yacc_naive_train = accuracy_score(new_policy.predict(Xexplore_sample), yr)
print("Test set accuarcy - Separate Classifiers: ", yacc_naive)
print("Train set accuarcy - Separate Classifiers: ", yacc_naive_train)
# +
from contextualbandits.online import SeparateClassifiers
from sklearn.linear_model import LogisticRegression
new_policy = SeparateClassifiers(base_algorithm=LogisticRegression(solver="lbfgs", max_iter=15000),
nchoices=y.shape[1], beta_prior="auto")
new_policy.fit(X=Xexplore_sample, a=actions_explore_sample, r=rewards_explore_sample)
mean_reward_beta = np.mean(y[st_test:end_test, :]\
[np.arange(end_test - st_test), new_policy.predict(Xtest)])
print("Test set mean reward - Separate Classifiers + Prior: ", mean_reward_beta)
yacc_beta=accuracy_score(new_policy.predict(Xtest), yt)
yacc_beta_train = accuracy_score(new_policy.predict(Xexplore_sample), yr)
print("Test set accuarcy - Separate Classifiers + Prior: ", yacc_beta)
print("Train set accuarcy - Separate Classifiers + Prior: ", yacc_beta_train)
# +
from contextualbandits.online import SeparateClassifiers
from sklearn.linear_model import LogisticRegression
new_policy = SeparateClassifiers(base_algorithm=LogisticRegression(solver="lbfgs", max_iter=15000),
nchoices=y.shape[1], beta_prior=None, smoothing = (1,2))
new_policy.fit(X=Xexplore_sample, a=actions_explore_sample, r=rewards_explore_sample)
mean_reward_sm = np.mean(y[st_test:end_test, :]\
[np.arange(end_test - st_test), new_policy.predict(Xtest)])
print("Test set mean reward - Separate Classifiers + Smoothing: ", mean_reward_sm)
yacc_sm=accuracy_score(new_policy.predict(Xtest), yt)
yacc_sm_train = accuracy_score(new_policy.predict(Xexplore_sample), yr)
print("Test set accuarcy - Separate Classifiers + Smoothing: ", yacc_sm)
print("Train set accuarcy - Separate Classifiers + Smoothing: ", yacc_sm_train)
# +
from contextualbandits.offpolicy import OffsetTree
from sklearn.linear_model import LogisticRegression
new_policy = OffsetTree(base_algorithm=LogisticRegression(solver="lbfgs", max_iter=15000), nchoices=y.shape[1])
new_policy.fit(X=Xexplore_sample, a=actions_explore_sample, r=rewards_explore_sample, p=prob_actions_explore)
mean_reward_ot = np.mean(y[st_test:end_test, :][np.arange(end_test - st_test), new_policy.predict(Xtest)])
print("Test set mean reward - Offset Tree technique: ", mean_reward_ot)
yacc_ot=accuracy_score(new_policy.predict(Xtest), yt)
yacc_ot_train = accuracy_score(new_policy.predict(Xexplore_sample), yr)
print("Test set accuarcy - Offset Tree technique: ", yacc_ot)
print("Train set accuarcy - Offset Tree technique: ", yacc_ot_train)
# +
from contextualbandits.offpolicy import DoublyRobustEstimator
from sklearn.linear_model import LogisticRegression, Ridge
new_policy = DoublyRobustEstimator(base_algorithm = Ridge(),
reward_estimator = LogisticRegression(solver="lbfgs", max_iter=15000),
nchoices = y.shape[1],
method = 'rovr', beta_prior = None, smoothing = None)
new_policy.fit(X=Xexplore_sample, a=actions_explore_sample, r=rewards_explore_sample, p=prob_actions_explore)
mean_reward_dr = np.mean(y[st_test:end_test, :][np.arange(end_test - st_test), new_policy.predict(Xtest)])
print("Test set mean reward - Doubly-Robust Estimator: ", mean_reward_dr)
yt = np.argmax(ytest, axis=1)
yacc_dr=accuracy_score(new_policy.predict(Xtest), yt)
yacc_dr_train=accuracy_score(new_policy.predict(Xexplore_sample), yr)
print("Test set accuarcy - Doubly-Robust Estimator: ", yacc_dr)
print("Train set accuarcy - Doubly-Robust Estimator: ", yacc_dr_train)
# -
new_policy = DoublyRobustEstimator(base_algorithm = Ridge(),
reward_estimator = LogisticRegression(solver="lbfgs", max_iter=15000),
nchoices = y.shape[1],
method = 'rovr', beta_prior = "auto", smoothing = None)
new_policy.fit(X=Xexplore_sample, a=actions_explore_sample, r=rewards_explore_sample, p=prob_actions_explore)
mean_reward_dr_prior = np.mean(y[st_test:end_test, :][np.arange(end_test - st_test), new_policy.predict(Xtest)])
print("Test set mean reward - Doubly-Robust Estimator + Prior: ", mean_reward_dr_prior)
yt = np.argmax(ytest, axis=1)
yacc_dr_prior=accuracy_score(new_policy.predict(Xtest), yt)
yacc_dr_prior_train=accuracy_score(new_policy.predict(Xexplore_sample), yr)
print("Test set accuarcy - Doubly-Robust Estimator + Prior: ", yacc_dr_prior)
print("Train set accuarcy - Doubly-Robust Estimator + Prior: ", yacc_dr_prior_train)
new_policy = DoublyRobustEstimator(base_algorithm = Ridge(),
reward_estimator = LogisticRegression(solver="lbfgs", max_iter=15000),
nchoices = y.shape[1],
method = 'rovr', beta_prior = None, smoothing = (1, 2))
new_policy.fit(X=Xexplore_sample, a=actions_explore_sample, r=rewards_explore_sample, p=prob_actions_explore)
mean_reward_dr_sm = np.mean(y[st_test:end_test, :][np.arange(end_test - st_test), new_policy.predict(Xtest)])
print("Test set mean reward - Doubly-Robust Estimator + Smoothing: ", mean_reward_dr_sm)
yacc_dr_sm=accuracy_score(new_policy.predict(Xtest), yt)
yacc_dr_sm_train=accuracy_score(new_policy.predict(Xexplore_sample), yr)
print("Test set accuarcy - Doubly-Robust Estimator + Smoothing: ", yacc_dr_sm)
print("Train set accuarcy - Doubly-Robust Estimator + Smoothing: ", yacc_dr_sm_train)
# +
import matplotlib.pyplot as plt, pandas as pd
import seaborn as sns
from pylab import rcParams
# %matplotlib inline
results = pd.DataFrame({
'Off-policy Learning Method' : ['Naive', 'Naive + Prior', 'Naive + Smoothing', 'Doubly-Robust',
'Doubly-Robust + Prior', 'Doubly-Robust + Smoothing', 'Offset Tree', 'Offset Tree + Ensemble'],
'Test set mean reward' : [mean_reward_naive, mean_reward_beta, mean_reward_sm, mean_reward_dr,
mean_reward_dr_prior, mean_reward_dr_sm, mean_reward_ot, mean_reward_ot_ensemble]
})
sns.set(font_scale = 1.1)
rcParams['figure.figsize'] = 22, 9
ax = sns.barplot(x = "Off-policy Learning Method", y="Test set mean reward", data=results)
sns.set(font_scale=2.5)
plt.xlabel("Off-policy Learning Method", fontsize = 20)
plt.ylabel("Test set mean reward", fontsize = 20)
for bar in ax.patches:
if bar.get_height() > 0.4:
bar.set_color('teal')
plt.title('Off-policy Learning on Bibtex Dataset\n(Base Classifier is Logistic Regression)')
plt.show()
# -
fig = ax.get_figure()
fig.savefig("Off_policy_MAB.png")
| .ipynb_checkpoints/Off_policy_Ensemble-checkpoint.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Aerospike Java Client – Reading and Updating Lists
// *Last updated: June 22, 2021*
//
// This notebook demonstrates Java Aerospike CRUD operations (Create, Read, Update, Delete) for lists of data, focusing on server-side **read** and **update** operations, including **sort**.
//
// This [Jupyter Notebook](https://jupyter-notebook.readthedocs.io/en/stable/notebook.html) requires the Aerospike Database running locally with Java kernel and Aerospike Java Client. To create a Docker container that satisfies the requirements and holds a copy of these notebooks, visit the [Aerospike Notebooks Repo](https://github.com/aerospike-examples/interactive-notebooks).
// + [markdown] heading_collapsed=true
// # Notebook Setup
//
// Run these first to initialize Jupyter, download the Java Client, and make sure the Aerospike Database is running.
// + [markdown] hidden=true
// ## Import Jupyter Java Integration
//
// Make it easier to work with Java in Jupyter.
// + hidden=true
import io.github.spencerpark.ijava.IJava;
import io.github.spencerpark.jupyter.kernel.magic.common.Shell;
IJava.getKernelInstance().getMagics().registerMagics(Shell.class);
// + [markdown] hidden=true
// ## Start Aerospike
//
// Ensure Aerospike Database is running locally.
// + hidden=true
// %sh asd
// + [markdown] hidden=true
// ## Download the Aerospike Java Client
//
// Ask Maven to download and install the project object model (POM) of the Aerospike Java Client.
// + hidden=true
// %%loadFromPOM
<dependencies>
<dependency>
<groupId>com.aerospike</groupId>
<artifactId>aerospike-client</artifactId>
<version>5.0.0</version>
</dependency>
</dependencies>
// + [markdown] hidden=true
// ## Start the Aerospike Java Client and Connect
//
// Create an instance of the Aerospike Java Client, and connect to the demo cluster.
//
// The default cluster location for the Docker container is *localhost* port *3000*. If your cluster is not running on your local machine, modify *localhost* and *3000* to the values for your Aerospike cluster.
// + hidden=true
import com.aerospike.client.AerospikeClient;
AerospikeClient client = new AerospikeClient("localhost", 3000);
System.out.println("Initialized the client and connected to the cluster.");
// -
// # CREATING Lists in Aerospike
// ## Create and Print List Data
//
// Create and print a String list and an Integer List.
// +
import java.util.ArrayList;
import java.util.Arrays;
import java.util.List;
import java.util.Map;
ArrayList<String> listStr = new ArrayList<String>();
listStr.add("Annette");
listStr.add("Redwood");
listStr.add("Aquamarine");
listStr.add("Pineapple");
System.out.println("String List: " + listStr);
ArrayList<Integer> listInt = new ArrayList<Integer>();
listInt.add(81);
listInt.add(3);
listInt.add(27);
listInt.add(9);
listInt.add(27);
listInt.add(1);
System.out.println("Integer List: " + listInt);
// -
// ## Insert the Lists into Aerospike
// ### Create a Key Object
//
// A **Key** uniquely identifies a specific **Record** in your Aerospike server or cluster. Each key must have a **Namespace** and optionally a **Set** name.
//
// * In Aerospike, a **Namespace** is like a relational database's tablespace.
// * A **Set** is like a relational database table in Aerospike.
// * A **Record** is like a row in a relational database table.
//
// The namespace *test* is configured on your Aerospike server or cluster. The rest can be defined and modified by Aerospike Java Client Code.
//
// For additional information on the [Aerospike Data Model](https://www.aerospike.com/docs/architecture/data-model.html), go [here](https://www.aerospike.com/docs/architecture/data-model.html).
// +
import com.aerospike.client.Key;
String listSet = "listset1";
String listNamespace = "test";
Integer theKey = 0;
Key key = new Key(listNamespace, listSet, theKey);
System.out.println("Key created." );
// -
// ### Create a Bin Object for Each List
//
// A **Bin** is a data field in an Aerospike record.
// +
import com.aerospike.client.Bin;
String listStrBinName = "liststrbin";
String listIntBinName = "listintbin";
Bin bin1 = new Bin(listStrBinName, listStr);
Bin bin2 = new Bin(listIntBinName, listInt);
System.out.println( "Created " + bin1 + " and " + bin2 + ".");
// -
// ### Create a Policy Object for Record Insertion
//
// A **Policy** tells Aerospike the intent of a database operation.
//
// For more information on policies, go [here](https://www.aerospike.com/docs/guide/policies.html).
// +
import com.aerospike.client.policy.ClientPolicy;
ClientPolicy clientPolicy = new ClientPolicy();
System.out.println("Created a client policy.");
// -
// ### Put the List Data into Aerospike
client.put(clientPolicy.writePolicyDefault, key, bin1, bin2);
System.out.println("Key: " + theKey + ", " + listStrBinName + ": " + listStr + ", " + listIntBinName + ": " + listInt);
// # READING Lists Elements From the Server
//
// Now that the lists are in Aerospike, the client can return full or partial lists from **bin** contents. No data is modified by these ops.
// ## Get the Record
//
// A record can be retrieved using the **key**, **namespace**, and **set** name.
//
// In the output:
// * **gen** is the generation number, the number of record writes.
// * **exp** is the expiration counter for the record.
//
// For more information on [both generation number and expiration](https://www.aerospike.com/docs/guide/FAQ.html), see the [Aerospike FAQ](https://www.aerospike.com/docs/guide/FAQ.html).
// +
import com.aerospike.client.Record;
Key key = new Key(listNamespace, listSet, theKey);
Record record = client.get(null, key);
System.out.println(record);
// -
// ## Get String Elements By Index and Rank
//
// The Aerospike API contains the operations to get list elements using index and rank.
// ### Get the Last String
//
// Aerospike provides operations to read list element(s) by **index**. As a convenience, the client returns the specified value as the contents of the bin.
//
// Aerospike operations allow indexing forward from the beginning of the list using zero-based numbering. Negative numbers index backwards from the end of a list.
//
// For more examples of indexes, go [here](https://www.aerospike.com/apidocs/java/com/aerospike/client/cdt/ListOperation.html).
// +
import com.aerospike.client.Operation;
import com.aerospike.client.Value;
import com.aerospike.client.cdt.ListOperation;
int last = -1;
Key key = new Key(listNamespace, listSet, theKey);
Record record = client.get(null, key);
Record lastString = client.operate(null, key,
ListOperation.get(listStrBinName, last)
);
System.out.println("The string list: " + record.getValue(listStrBinName));
System.out.println("The last string: " + lastString.getValue(listStrBinName));
// -
// ### Get the Highest Rank Item
//
// Aerospike provides operations to read list item(s) by **Rank**. The API methods contain options prescribing what type of data to return from an operation.
//
// For information on list ranking, go [here](https://en.wikipedia.org/wiki/List_ranking).
//
// For the list of return type options, go [here](https://www.aerospike.com/apidocs/java/com/aerospike/client/cdt/ListReturnType.html).
// +
import com.aerospike.client.cdt.ListReturnType;
int highestRank = -1;
Key key = new Key(listNamespace, listSet, theKey);
Record record = client.get(null, key);
Record highestRankString = client.operate(null, key,
ListOperation.getByRank(listStrBinName, highestRank, ListReturnType.VALUE)
);
System.out.println("The string list: " + record.getValue(listStrBinName));
System.out.println("The highest rank string: " + highestRankString.getValue(listStrBinName));
// -
// ## Get Integer Elements By Value Range and Rank Range
//
// Read integer values from the Aerospike Server or Cluster using value range or rank range.
// ### Get Integers Between 3 and 27
//
// In addition to reading list elements by rank and index, Aerospike operations can return a **Range** of elements by value.
// * The lower bound of a range is included.
// * The upper bound of a range is excluded.
//
// For more examples of ranges, go [here](https://www.aerospike.com/apidocs/java/com/aerospike/client/cdt/ListOperation.html).
// +
int lowerBound = 3;
int upperBound = 27;
Key key = new Key(listNamespace, listSet, theKey);
Record record = client.get(null, key);
Record between3And27 = client.operate(null, key,
ListOperation.getByValueRange(listIntBinName, Value.get(lowerBound), Value.get(upperBound),
ListReturnType.VALUE)
);
System.out.println("The integer list: " + record.getValue(listIntBinName));
System.out.println("The integers between " + lowerBound + " and " + upperBound + ": "
+ between3And27.getValue(listIntBinName));
// -
// ### Get the 2nd and 3rd Ranked Integers
//
// Aerospike provides operations to return a range of elements by rank. Rank is zero-based.
// +
int secondRank = 1;
int rangeRankSize = 2;
Key key = new Key(listNamespace, listSet, theKey);
Record record = client.get(null, key);
Record rank1And2 = client.operate(client.writePolicyDefault, key,
ListOperation.getByRankRange(listIntBinName, secondRank, rangeRankSize, ListReturnType.VALUE)
);
System.out.println("The integer list: " + record.getValue(listIntBinName));
System.out.println("The 2nd and 3rd ranked integers: " + rank1And2.getValue(listIntBinName));
// -
// # UPDATING Lists on the Aerospike Server
//
// Aerospike's [list operations](https://www.aerospike.com/apidocs/java/com/aerospike/client/cdt/ListOperation.html) can also modify data in the Aerospike Database.
// ## Update the String List in Aerospike
// ### Insert a Fish into the Second and Second-from-last Position
//
// Aerospike's list insert operation inserts before the item at an index, and increases the index of the item at the index and all subsequent items in the list.
// +
String Fish = "Koi";
int secondPosition = 1;
int beforeLastPosition = -1;
Key key = new Key(listNamespace, listSet, theKey);
Record origRecord = client.get(null, key);
System.out.println("Before – " + origRecord.getValue(listStrBinName));
origRecord = client.operate(client.writePolicyDefault, key,
ListOperation.insert(listStrBinName, beforeLastPosition, Value.get(Fish)),
ListOperation.insert(listStrBinName, secondPosition, Value.get(Fish))
);
Record finalRecord = client.get(null, key);
System.out.println(" After – " + finalRecord.getValue(listStrBinName));
// -
// ### Remove By Index from the String List
// +
int firstPosition = 0;
Key key = new Key(listNamespace, listSet, theKey);
Record origRecord = client.get(null, key);
System.out.println("Before – " + origRecord.getValue(listStrBinName));
origRecord = client.operate(client.writePolicyDefault, key,
ListOperation.remove(listStrBinName, firstPosition)
);
Record finalRecord = client.get(null, key);
System.out.println(" After – " + finalRecord.getValue(listStrBinName));
// -
// ## Update the Integer List in Aerospike
// ### Append 17 to the List
// +
int seventeen = 17;
Key key = new Key(listNamespace, listSet, theKey);
Record origRecord = client.get(null, key);
System.out.println("Before – " + origRecord.getValue(listIntBinName));
origRecord = client.operate(client.writePolicyDefault, key,
ListOperation.append(listIntBinName, Value.get(seventeen))
);
Record finalRecord = client.get(null, key);
System.out.println(" After – " + finalRecord.getValue(listIntBinName));
// -
// ### Increment the 4th Integer by 111
//
// Indexes into lists start at 0.
// +
int incNum = 111;
int incIndex = 3;
Key key = new Key(listNamespace, listSet, theKey);
Record origRecord = client.get(null, key);
System.out.println("Before – " + origRecord.getValue(listIntBinName) );
origRecord = client.operate(client.writePolicyDefault, key,
ListOperation.increment(listIntBinName, incIndex, Value.get(incNum))
);
Record finalRecord = client.get(null, key);
System.out.println(" After – " + finalRecord.getValue(listIntBinName) );
// -
// ## Sorting the Lists in the Aerospike Java Client
//
// Aerospike also provides both:
// 1. An operation to **sort** lists in the client and optionally remove duplicates.
// 2. An operation to store list data in **order**.
//
// For information on maintaining list data in order, go [here](https://www.aerospike.com/apidocs/java/com/aerospike/client/cdt/ListOrder.html).
// ### Sort the String and Drop Duplicates
//
// For information on the flags specifying whether to remove duplicates, go [here](https://www.aerospike.com/apidocs/java/com/aerospike/client/cdt/ListSortFlags.html).
// +
import com.aerospike.client.cdt.ListSortFlags;
Key key = new Key(listNamespace, listSet, theKey);
Record origRecord = client.get(null, key);
System.out.println("Unsorted – " + origRecord.getValue(listStrBinName));
origRecord = client.operate(client.writePolicyDefault, key,
ListOperation.sort(listStrBinName, ListSortFlags.DROP_DUPLICATES)
);
Record finalRecord = client.get(null, key);
System.out.println(" Sorted – " + finalRecord.getValue(listStrBinName));
// -
// ### Sort the Integer List and Keep Duplicates
// +
Key key = new Key(listNamespace, listSet, theKey);
Record origRecord = client.get(null, key);
System.out.println("Unsorted – " + origRecord.getValue(listIntBinName));
origRecord = client.operate(client.writePolicyDefault, key,
ListOperation.sort(listIntBinName, ListSortFlags.DEFAULT)
);
Record finalRecord = client.get(null, key);
System.out.println(" Sorted – " + finalRecord.getValue(listIntBinName));
// + [markdown] heading_collapsed=true
// # Notebook Cleanup
// + [markdown] hidden=true
// ## Truncate the Set
// Truncate the set from the Aerospike Database.
// + hidden=true
import com.aerospike.client.policy.InfoPolicy;
InfoPolicy infoPolicy = new InfoPolicy();
client.truncate(infoPolicy, listNamespace, listSet, null);
System.out.println("Set truncated.");
// + [markdown] hidden=true
// ## Close the Connection to Aerospike
// + hidden=true
client.close();
System.out.println("Server connection closed.");
// + [markdown] heading_collapsed=true
// # Code Summary
//
// Here is a collection of all of the non-Jupyter code from this tutorial.
// + [markdown] hidden=true
// ## Overview
//
// 1. Import Java Libraries.
// 2. Import Aerospike Client Libraries.
// 3. Start the Aerospike Client.
// 4. Create Test Data.
// 5. Put Record into Aerospike.
// 6. Get Data from Aerospike.
// 1. Get the Record.
// 2. Get the Last String and Highest Rank.
// 3. Get Integers Between 3 and 27.
// 4. Get 2 Integers By Rank Starting with the Second Rank Item.
// 7. Update the Record in Aerospike
// 1. Add Koi twice to the String List.
// 2. Remove the Name from the String List.
// 3. Append 17 to the Integer List.
// 4. Increment the 4th Integer by 111.
// 5. Sort the Strings and Drop Duplicates.
// 6. Sort the Integers and Keep Duplicates.
// 8. Truncate the Set.
// 9. Close the Client Connections.
// + hidden=true
// Import Java Libraries.
import java.util.ArrayList;
import java.util.Arrays;
import java.util.List;
import java.util.Map;
// Import Aerospike Client Libraries.
import com.aerospike.client.AerospikeClient;
import com.aerospike.client.Key;
import com.aerospike.client.Bin;
import com.aerospike.client.policy.ClientPolicy;
import com.aerospike.client.Record;
import com.aerospike.client.Operation;
import com.aerospike.client.Value;
import com.aerospike.client.cdt.ListOperation;
import com.aerospike.client.cdt.ListReturnType;
import com.aerospike.client.cdt.ListSortFlags;
import com.aerospike.client.policy.InfoPolicy;
// Start the Aerospike Client.
AerospikeClient client = new AerospikeClient("localhost", 3000);
System.out.println("Initialized the client and connected to the cluster.");
// Create Test Data.
ArrayList<String> listStr = new ArrayList<String>();
listStr.add("Annette");
listStr.add("Redwood");
listStr.add("Aquamarine");
listStr.add("Pineapple");
System.out.println("Created String List: " + listStr);
ArrayList<Integer> listInt = new ArrayList<Integer>();
listInt.add(81);
listInt.add(3);
listInt.add(27);
listInt.add(9);
listInt.add(27);
listInt.add(1);
System.out.println("Created Integer List: " + listInt);
// Put Record into Aerospike.
String listSet = "listset1";
String listNamespace = "test";
String listStrBinName = "liststrbin";
String listIntBinName = "listintbin";
ClientPolicy clientPolicy = new ClientPolicy();
InfoPolicy infoPolicy = new InfoPolicy();
Integer theKey = 0;
Key key = new Key(listNamespace, listSet, theKey);
Bin bin1 = new Bin(listStrBinName, listStr);
Bin bin2 = new Bin(listIntBinName, listInt);
client.put(clientPolicy.writePolicyDefault, key, bin1, bin2);
System.out.println("Inserted Key: " + theKey + ", " + listStrBinName + ": " + listStr + ", " + listIntBinName + ": " + listInt);
// Get Data from Aerospike.
// 1. Get Record.
// 2. Get the Last String and Highest Rank.
// 3. Get Integers Between 3 and 27.
// 4. Get 2 Integers By Rank Starting with the Second Rank Item.
int last = -1;
int highestRank = -1;
int lowerBound = 3;
int upperBound = 27;
int secondRank = 1;
int rangeRankSize = 2;
Key key = new Key(listNamespace, listSet, theKey);
Record record = client.get(null, key);
Record postOp = client.operate(client.writePolicyDefault, key,
ListOperation.get(listStrBinName, last),
ListOperation.getByRank(listStrBinName, highestRank, ListReturnType.VALUE),
ListOperation.getByValueRange(listIntBinName, Value.get(lowerBound), Value.get(upperBound),
ListReturnType.VALUE),
ListOperation.getByRankRange(listIntBinName, secondRank, rangeRankSize, ListReturnType.VALUE)
);
List<?> returnStr = postOp.getList(listStrBinName);
List<?> returnIntList = postOp.getList(listIntBinName);
System.out.println("Read the Full Record From Aerospike:" + record);
System.out.println("The last string: " + returnStr.get(0));
System.out.println("The highest rank string: " + returnStr.get(1));
System.out.println("The integers between " + lowerBound + " and " + upperBound + ": "
+ returnIntList.get(0));
System.out.println("The 2nd and 3rd ranked integers: " + returnIntList.get(1));
// Update the Record in Aerospike
// 1. Add Koi twice to the String List.
// 2. Remove the Name from the String List.
// 3. Append 17 to the Integer List.
// 4. Increment the 4th Integer by 111.
// 5. Sort the Strings and Drop Duplicates.
// 6. Sort the Integers and Keep Duplicates.
String Fish = "Koi";
int fishIndex0 = 1;
int fishIndex1 = -1;
int nameIndex = 0;
int seventeen = 17;
int incNum = 111;
int incIndex = 3;
origRecord = client.operate(client.writePolicyDefault, key,
ListOperation.insert(listStrBinName, fishIndex0, Value.get(Fish)),
ListOperation.insert(listStrBinName, fishIndex1, Value.get(Fish)),
ListOperation.remove(listStrBinName, nameIndex),
ListOperation.append(listIntBinName, Value.get(seventeen)),
ListOperation.increment(listIntBinName, incIndex, Value.get(incNum)),
ListOperation.sort(listStrBinName, ListSortFlags.DROP_DUPLICATES),
ListOperation.sort(listIntBinName, ListSortFlags.DEFAULT)
);
List<?> opStrResults = origRecord.getList(listStrBinName);
List<?> opIntResults = origRecord.getList(listIntBinName);
Record finalRecord = client.get(null, key);
System.out.println("Inserted " + Fish + "; " + listStrBinName + " size is now: " + opStrResults.get(0));
System.out.println("Inserted " + Fish + "; " + listStrBinName + " size is now: " + opStrResults.get(1));
System.out.println("Removed item at index " + nameIndex + "; removed " + opStrResults.get(2) + " item");
System.out.println("Appended " + seventeen + ", " + listIntBinName + " size is now " + opIntResults.get(0));
System.out.println("Incremented item at index " + incIndex + " by " + incNum + "; new value is: " + opIntResults.get(1));
System.out.println("Sorted both lists and removed duplicates in " + listStrBinName);
System.out.println("After Record Edits – " + finalRecord);
// Truncate the Set.
client.truncate(infoPolicy, listNamespace, listSet, null);
System.out.println("Set truncated.");
// Close the Client Connections.
client.close();
System.out.println("Closed client connections.");
// -
// # Takeaway – Aerospike Does Lists
//
// Aerospike and its Java Client are up to the task of working with your list data. APIs contain rich operations to read and update list data using index, value, and rank.
// # What's Next?
//
//
//
//
// ## Next Steps
//
// Have questions? Don't hesitate to reach out if you have additional questions about working with lists at https://discuss.aerospike.com/.
//
// Want to learn about modeling using Lists? See, [Modeling Using Lists](java-modeling_using_lists.ipynb).
//
//
// Want to check out other Java notebooks?
// 1. [Hello, World](hello_world.ipynb)
// 2. [Reading and Updating Maps](java-working_with_maps.ipynb)
// 3. [Aerospike Query and UDF](query_udf.ipynb)
//
// Are you running this from Binder? [Download the Aerospike Notebook Repo](https://github.com/aerospike-examples/interactive-notebooks) and work with Aerospike Database and Jupyter locally using a Docker container.
// ## Additional Resources
//
// * Want to get started with Java? [Download](https://www.aerospike.com/download/client/) or [install](https://github.com/aerospike/aerospike-client-java) the Aerospike Java Client.
// * What other ways can we work with Lists? Take a look at [Aerospike's List Operations](https://www.aerospike.com/apidocs/java/com/aerospike/client/cdt/ListOperation.html).
// * What are Namespaces, Sets, and Bins? Check out the [Aerospike Data Model](https://www.aerospike.com/docs/architecture/data-model.html).
// * How robust is the Aerospike Database? Browse the [Aerospike Database Architecture](https://www.aerospike.com/docs/architecture/index.html).
| notebooks/java/java-working_with_lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID" # see issue #152
os.environ["CUDA_VISIBLE_DEVICES"]="0"
res_folders=os.listdir('../../results/')
import tensorflow as tf
import keras
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
config.gpu_options.visible_device_list = str(0)# str(hvd.local_rank())
keras.backend.set_session(tf.Session(config=config))
verbose=1
init=tf.global_variables_initializer() #initialize_all_variables()
sess=tf.Session()
sess.run(init)
model_folder='/home/mara/multitask_adversarial/results/STANDARD_FCONTRAST_2409/'
CONCEPT=['full_contrast']
import keras
keras.__version__
from sklearn.metrics import accuracy_score
'../../doc/data_shuffle.csv'
new_folder=model_folder
# +
"""
RUN SEQUENTIAL
python hvd_train_unc.py SEED EXPERIMENT_TYPE
"""
BATCH_SIZE=32
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
import warnings
warnings.filterwarnings("ignore")
import logging
logging.getLogger('tensorflow').disabled = True
from keras import *
import setproctitle
SERVER_NAME = 'ultrafast'
import time
import sys
import shutil
sys.path.append('../../lib/TASK_2_UC1/')
from models import *
from util import otsu_thresholding
from extract_xml import *
from functions import *
sys.path.append('../../lib/')
from mlta import *
import math
import keras.callbacks as callbacks
from keras.callbacks import Callback
import tensorflow as tf
cam16 = hd.File('/home/mara/adversarialMICCAI/data/ultrafast/cam16_500/patches.h5py', 'r', libver='latest', swmr=True)
all500 = hd.File('/home/mara/adversarialMICCAI/data/ultrafast/all500/patches.h5py', 'r', libver='latest', swmr=True)
extra17 = hd.File('/home/mara/adversarialMICCAI/data/ultrafast/extra17/patches.h5py', 'r', libver='latest', swmr=True)
tumor_extra17=hd.File('/home/mara/adversarialMICCAI/data/ultrafast/1129-1155/patches.h5py', 'r', libver='latest', swmr=True)
test2 = hd.File('/home/mara/adversarialMICCAI/data/ultrafast/test_data2/patches.h5py', 'r', libver='latest', swmr=True)
pannuke= hd.File('/home/mara/adversarialMICCAI/data/ultrafast/pannuke/patches_fix.h5py', 'r', libver='latest', swmr=True)
global data
data={'cam16':cam16,'all500':all500,'extra17':extra17, 'tumor_extra17':tumor_extra17, 'test_data2': test2, 'pannuke':pannuke}
global concept_db
concept_db = hd.File('../../data/normalized_cmeasures/concept_values_def.h5py','r', libver='latest', swmr=True)
# DATA SPLIT CSVs
train_csv=open('/mnt/nas2/results/IntermediateResults/Camelyon/train_shuffle.csv', 'r') # How is the encoding of .csv files ?
val_csv=open('/mnt/nas2/results/IntermediateResults/Camelyon/val_shuffle.csv', 'r')
test_csv=open('/mnt/nas2/results/IntermediateResults/Camelyon/test_shuffle.csv', 'r')
train_list=train_csv.readlines()
val_list=val_csv.readlines()
test_list=test_csv.readlines()
test2_csv = open('/mnt/nas2/results/IntermediateResults/Camelyon/test2_shuffle.csv', 'r')
test2_list=test2_csv.readlines()
test2_csv.close()
train_csv.close()
val_csv.close()
test_csv.close()
data_csv=open('../../doc/data_shuffle.csv', 'r')
data_list=data_csv.readlines()
data_csv.close()
# STAIN NORMALIZATION
def get_normalizer(patch, save_folder=''):
normalizer = ReinhardNormalizer()
normalizer.fit(patch)
np.save('{}/normalizer'.format(save_folder),normalizer)
np.save('{}/normalizing_patch'.format(save_folder), patch)
#print('Normalisers saved to disk.')
return normalizer
def normalize_patch(patch, normalizer):
return np.float64(normalizer.transform(np.uint8(patch)))
# LOAD DATA NORMALIZER
global normalizer
db_name, entry_path, patch_no = get_keys(data_list[0])
normalization_reference_patch = data[db_name][entry_path][patch_no]
normalizer = get_normalizer(normalization_reference_patch, save_folder=new_folder)
"""
Batch generators:
They load a patch list: a list of file names and paths.
They use the list to create a batch of 32 samples.
"""
# Retrieve Concept Measures
def get_concept_measure(db_name, entry_path, patch_no, measure_type=''):
if measure_type=='domain':
return get_domain(db_name, entry_path)
path=db_name+'/'+entry_path+'/'+str(patch_no)+'/'+measure_type.strip(' ')
try:
cm=concept_db[path][0]
return cm
except:
print("[ERR]: {}, {}, {}, {} with path {}".format(db_name, entry_path, patch_no, measure_type, path))
return 1.
# BATCH GENERATORS
class DataGenerator(keras.utils.Sequence):
def __init__(self, patch_list, concept=CONCEPT, batch_size=32, shuffle=True, data_type=0):
self.batch_size=batch_size
self.patch_list=patch_list
self.shuffle=shuffle
self.concept = concept
self.data_type=data_type
#print 'data type:', data_type
self.on_epoch_end()
def __len__(self):
return int(np.floor(len(self.patch_list)/self.batch_size))
def __getitem__(self, index):
#import pdb; pdb.set_trace()
indexes=self.indexes[index*self.batch_size:(index+1)*self.batch_size]
patch_list_temp=[self.patch_list[k] for k in indexes]
self.patch_list_temp=patch_list_temp
return self.__data_generation(self)
def on_epoch_end(self):
self.indexes = np.arange(len(self.patch_list))
if self.shuffle == True:
np.random.shuffle(self.indexes)
def __data_generation(self, patch_list_temp):
patch_list_temp=self.patch_list_temp
batch_x=np.zeros((len(patch_list_temp), 224,224,3))
batch_y=np.zeros(len(patch_list_temp))
i=0
for line in patch_list_temp:
db_name, entry_path, patch_no = get_keys(line)
patch=data[db_name][entry_path][patch_no]
patch=normalize_patch(patch, normalizer)
patch=keras.applications.inception_v3.preprocess_input(patch)
label = get_class(line, entry_path)
if self.data_type!=0:
label=get_test_label(entry_path)
batch_x[i]=patch
batch_y[i]=label
i+=1
generator_output=[batch_y]
for c in self.concept:
batch_concept_values=np.zeros(len(patch_list_temp))
i=0
for line in patch_list_temp:
db_name, entry_path, patch_no = get_keys(line)
batch_concept_values[i]=get_concept_measure(db_name, entry_path, patch_no, measure_type=c)
i+=1
if c=='domain':
batch_concept_values=keras.utils.to_categorical(batch_concept_values, num_classes=7)
generator_output.append(batch_concept_values)
return batch_x, generator_output # batch_x, np.asarray(batch_y), generator_output[1], generator_output[2]
"""
Credits for the Gradient Reversal Layer
https://github.com/michetonu/gradient_reversal_keras_tf/blob/master/flipGradientTF.py
"""
def reverse_gradient(X, hp_lambda):
'''Flips the sign of the incoming gradient during training.'''
hp_lambda = hp_lambda
try:
reverse_gradient.num_calls += 1
except AttributeError:
reverse_gradient.num_calls = 1
grad_name = "GradientReversal%d" % reverse_gradient.num_calls
@tf.RegisterGradient(grad_name)
def _flip_gradients(op, grad):
grad = tf.negative(grad)
#grad = tf.Print(grad, [grad], 'grad')
final_val = grad * hp_lambda
#final_val =tf.Print(final_val, [final_val], 'final_val')
return [final_val]
g = keras.backend.get_session().graph
with g.gradient_override_map({'Identity': grad_name}):
y = tf.identity(X)
return y
class Hp_lambda():
def __init__(self,in_val):
self.value=in_val
def update(self,new_val):
self.value=new_val
def get_hyperparameter_lambda(self):
#val = tf.Print(self.value,[self.value],'hplambda: ')
return tf.Variable(self.value, name='hp_lambda')
#return lmb
class GradientReversal(Layer):
'''Flip the sign of gradient during training.'''
def __init__(self, hp_lambda, **kwargs):
super(GradientReversal, self).__init__(**kwargs)
self.supports_masking = False
self.hp_lambda = Hp_lambda(hp_lambda)
#self.hp_lambda = tf.Variable(hp_lambda, name='hp_lambda')
#self.hp_lambda = tf.Variable(hp_lambda, name='hp_lambda')
def build(self, input_shape):
self.trainable_weights = []
return
def call(self, x, mask=None):
#tf.Print(self.hp_lambda, [self.hp_lambda],'self.hp_lambda: ')
#with tf.Session() as sess: print(self.hp_lambda.eval())
lmb=self.hp_lambda.get_hyperparameter_lambda()
return reverse_gradient(x, lmb)
def get_output_shape_for(self, input_shape):
return input_shape
def get_config(self):
config = {"name": self.__class__.__name__,
'hp_lambda': keras.backend.get_value(self.hp_lambda)}
base_config = super(GradientReversal, self).get_config()
return dict(list(base_config.items()) + list(config.items()))
"""
Building guidable model
"""
def get_baseline_model(hp_lambda=0., c_list=[]):
base_model = keras.applications.inception_v3.InceptionV3(include_top=False, weights='imagenet', input_shape=(224,224,3))
layers_list=['conv2d_92', 'conv2d_93', 'conv2d_88', 'conv2d_89', 'conv2d_86']
#layers_list=[]
for i in range(len(base_model.layers[:])):
layer=base_model.layers[i]
if layer.name in layers_list:
print layer.name
layer.trainable=True
else:
layer.trainable = False
feature_output=base_model.layers[-1].output
gap_layer_output = keras.layers.GlobalAveragePooling2D()(feature_output)
feature_output = Dense(2048, activation='relu', name='finetuned_features1',kernel_regularizer=keras.regularizers.l2(0.01))(gap_layer_output)
feature_output = keras.layers.Dropout(0.8, noise_shape=None, seed=None)(feature_output)
feature_output = Dense(512, activation='relu', name='finetuned_features2',kernel_regularizer=keras.regularizers.l2(0.01))(feature_output)
feature_output = keras.layers.Dropout(0.8, noise_shape=None, seed=None)(feature_output)
feature_output = Dense(256, activation='relu', name='finetuned_features3',kernel_regularizer=keras.regularizers.l2(0.01))(feature_output)
feature_output = keras.layers.Dropout(0.8, noise_shape=None, seed=None)(feature_output)
finetuning = Dense(1,name='predictions')(feature_output)
if 'domain' in CONCEPT:
grl_layer=GradientReversal(hp_lambda=hp_lambda)
feature_output_grl = grl_layer(feature_output)
domain_adversarial = keras.layers.Dense(7, activation = keras.layers.Activation('softmax'), name='domain_adversarial')(feature_output_grl)
output_nodes=[finetuning, domain_adversarial]
else:
output_nodes=[finetuning]
for c in c_list:
if c!='domain':
concept_layer= keras.layers.Dense(1, activation = keras.layers.Activation('linear'), name='extra_{}'.format(c.strip(' ')))(feature_output)
output_nodes.append(concept_layer)
model = Model(input=base_model.input, output=output_nodes)
if 'domain' in CONCEPT:
model.grl_layer=grl_layer
return model
"""
LOSS FUNCTIONS
"""
def keras_mse(y_true, y_pred):
return tf.reduce_mean(tf.keras.losses.mean_squared_error(y_true, y_pred))
def bbce(y_true, y_pred):
# we use zero weights to set the loss to zero for unlabeled data
verbose=0
zero= tf.constant(-1, dtype=tf.float32)
where = tf.not_equal(y_true, zero)
where = tf.reshape(where, [-1])
indices=tf.where(where) #indices where the item of y_true is NOT -1
indices = tf.reshape(indices, [-1])
sliced_y_true = tf.nn.embedding_lookup(y_true, indices)
sliced_y_pred = tf.nn.embedding_lookup(y_pred, indices)
n1 = tf.shape(indices)[0] #number of train images in batch
batch_size = tf.shape(y_true)[0]
n2 = batch_size - n1 #number of test images in batch
sliced_y_true = tf.reshape(sliced_y_true, [n1, -1])
n1_ = tf.cast(n1, tf.float32)
n2_ = tf.cast(n2, tf.float32)
multiplier = (n1_+ n2_) / n1_
zero_class = tf.constant(0, dtype=tf.float32)
where_class_is_zero=tf.cast(tf.reduce_sum(tf.cast(tf.equal(sliced_y_true, zero_class), dtype=tf.float32)), dtype=tf.float32)
if verbose:
where_class_is_zero=tf.Print(where_class_is_zero,[where_class_is_zero],'where_class_is_zero: ')
class_weight_zero = tf.cast(tf.divide(n1_, 2. * tf.cast(where_class_is_zero, dtype=tf.float32)+0.001), dtype=tf.float32)
if verbose:
class_weight_zero=tf.Print(class_weight_zero,[class_weight_zero],'class_weight_zero: ')
one_class = tf.constant(1, dtype=tf.float32)
where_class_is_one=tf.cast(tf.reduce_sum(tf.cast(tf.equal(sliced_y_true, one_class), dtype=tf.float32)), dtype=tf.float32)
if verbose:
where_class_is_one=tf.Print(where_class_is_one,[where_class_is_one],'where_class_is_one: ')
n1_=tf.Print(n1_,[n1_],'n1_: ')
class_weight_one = tf.cast(tf.divide(n1_, 2. * tf.cast(where_class_is_one,dtype=tf.float32)+0.001), dtype=tf.float32)
class_weight_zero = tf.constant(23477.0/(23477.0+123820.0), dtype=tf.float32)
class_weight_one = tf.constant(123820.0/(23477.0+123820.0), dtype=tf.float32)
A = tf.ones(tf.shape(sliced_y_true), dtype=tf.float32) - sliced_y_true
A = tf.scalar_mul(class_weight_zero, A)
B = tf.scalar_mul(class_weight_one, sliced_y_true)
class_weight_vector=A+B
ce = tf.nn.sigmoid_cross_entropy_with_logits(labels=sliced_y_true,logits=sliced_y_pred)
ce = tf.multiply(class_weight_vector,ce)
return tf.reduce_mean(ce)
from keras.initializers import Constant
global domain_weight
global main_task_weight
"""
EVALUATION FUNCTIONs
"""
def accuracy_domain(y_true,y_pred):
#y_p_r=
y_p_r = np.asarray([np.argmax(y_pred[i,:]) for i in range(len(y_pred[:,0]))])
#y_p_r=np.round(y_pred)
y_true = np.asarray([np.argmax(y_true[i,:]) for i in range(len(y_true[:,0]))])
acc = np.equal(y_p_r, y_true)**1.
acc = np.mean(np.float32(acc))
return acc
def my_sigmoid(x):
return 1 / (1 + np.exp(-x))
def my_accuracy_np(y_true, y_pred):
sliced_y_pred = my_sigmoid(y_pred)
y_pred_rounded = np.round(sliced_y_pred)
acc = np.equal(y_pred_rounded, y_true)**1.
acc = np.mean(np.float32(acc))
return acc
def r_square_np(y_true, y_pred):
SS_res = np.sum(np.square(y_true - y_pred))
SS_tot = np.sum(np.square(y_true - np.mean(y_true)))
r2_mine=( 1 - SS_res/(SS_tot + keras.backend.epsilon()) )
return ( 1 - SS_res/(SS_tot + keras.backend.epsilon()) )
global report_val_acc
global report_val_r2
global report_val_mse
report_val_acc=[]
report_val_r2=[]
report_val_mse=[]
# LOG FILE
global log_file
"""
DATA GENERATORS CREATION
"""
train_generator=DataGenerator(data_list, concept=CONCEPT, batch_size=BATCH_SIZE, data_type=0)
train_generator2=DataGenerator(data_list, concept=CONCEPT, batch_size=BATCH_SIZE, data_type=1)#get_batch_data(data_list, batch_size=BATCH_SIZE)
val_generator=DataGenerator(val_list, concept=CONCEPT, batch_size=BATCH_SIZE, data_type=1) #get_test_batch(val_list, batch_size=BATCH_SIZE)
val_generator2= DataGenerator(val_list, concept=CONCEPT, batch_size=BATCH_SIZE, data_type=1) #get_test_batch(val_list, batch_size=BATCH_SIZE)
test_generator= DataGenerator(test_list, concept=CONCEPT, batch_size=BATCH_SIZE, data_type=1) #get_test_batch(test_list, batch_size=BATCH_SIZE)
print('data generators created')
"""
Get trainable model with Baseline Loss weighting
"""
def compile_base_model(baseline_model, opt, concept_list=CONCEPT, loss=None, metrics=None):
losses = [bbce, 'categorical_crossentropy']
loss_weights=[1.,1.]
for i in range(1,len(concept_list)):
losses.append('mean_squared_error')
loss_weights.append(1.)
model.compile(optimizer=opt,
loss=losses,
loss_weights=loss_weights,
metrics=[my_acc_f, r_square])
return
verbose=True
def custom_train_model(baseline_model,
epochs= 1,
lr=1e-4, verbose=True,
train_generator=train_generator,
val_generator=val_generator,
grl_layer=None,
):
opt = keras.optimizers.SGD(lr=lr, momentum=0.9, nesterov=True)
compile_base_model(baseline_model,opt,loss=None,metrics=None)
history = baseline_model.fit_generator(train_generator,
steps_per_epoch= len(data_list) // (BATCH_SIZE ),
callbacks=callbacks,
epochs=epochs,
verbose=verbose,
workers=4,
use_multiprocessing=False,
validation_data= val_generator,
validation_steps= len(val_list)//BATCH_SIZE)
return baseline_model
# -
def get_bootstrap_sample(data, n_samples=2):
sample_=[data[i] for i in np.random.choice(len(data),n_samples)]
#sample_=[data[i] for i in range(len(data))]
return sample_
# +
# END Callbacks
#
def keras_mse(y_true, y_pred):
return tf.reduce_mean(tf.keras.losses.mean_squared_error(y_true, y_pred))
#return tf.keras.losses.mean_squared_error(y_true, y_pred)
def bbce(y_true, y_pred):
# we use zero weights to set the loss to zero for unlabeled data
verbose=0
zero= tf.constant(-1, dtype=tf.float32)
where = tf.not_equal(y_true, zero)
where = tf.reshape(where, [-1])
indices=tf.where(where) #indices where the item of y_true is NOT -1
indices = tf.reshape(indices, [-1])
sliced_y_true = tf.nn.embedding_lookup(y_true, indices)
sliced_y_pred = tf.nn.embedding_lookup(y_pred, indices)
n1 = tf.shape(indices)[0] #number of train images in batch
batch_size = tf.shape(y_true)[0]
n2 = batch_size - n1 #number of test images in batch
sliced_y_true = tf.reshape(sliced_y_true, [n1, -1])
n1_ = tf.cast(n1, tf.float32)
n2_ = tf.cast(n2, tf.float32)
multiplier = (n1_+ n2_) / n1_
zero_class = tf.constant(0, dtype=tf.float32)
where_class_is_zero=tf.cast(tf.reduce_sum(tf.cast(tf.equal(sliced_y_true, zero_class), dtype=tf.float32)), dtype=tf.float32)
if verbose:
where_class_is_zero=tf.Print(where_class_is_zero,[where_class_is_zero],'where_class_is_zero: ')
class_weight_zero = tf.cast(tf.divide(n1_, 2. * tf.cast(where_class_is_zero, dtype=tf.float32)+0.001), dtype=tf.float32)
if verbose:
class_weight_zero=tf.Print(class_weight_zero,[class_weight_zero],'class_weight_zero: ')
one_class = tf.constant(1, dtype=tf.float32)
where_class_is_one=tf.cast(tf.reduce_sum(tf.cast(tf.equal(sliced_y_true, one_class), dtype=tf.float32)), dtype=tf.float32)
if verbose:
where_class_is_one=tf.Print(where_class_is_one,[where_class_is_one],'where_class_is_one: ')
n1_=tf.Print(n1_,[n1_],'n1_: ')
class_weight_one = tf.cast(tf.divide(n1_, 2. * tf.cast(where_class_is_one,dtype=tf.float32)+0.001), dtype=tf.float32)
class_weight_zero = tf.constant(23477.0/(23477.0+123820.0), dtype=tf.float32)
class_weight_one = tf.constant(123820.0/(23477.0+123820.0), dtype=tf.float32)
A = tf.ones(tf.shape(sliced_y_true), dtype=tf.float32) - sliced_y_true
A = tf.scalar_mul(class_weight_zero, A)
B = tf.scalar_mul(class_weight_one, sliced_y_true)
class_weight_vector=A+B
ce = tf.nn.sigmoid_cross_entropy_with_logits(labels=sliced_y_true,logits=sliced_y_pred)
ce = tf.multiply(class_weight_vector,ce)
return tf.reduce_mean(ce)
# Custom loss layer
from keras.initializers import Constant
class CustomMultiLossLayer(Layer):
def __init__(self, nb_outputs=2, **kwargs):
self.nb_outputs = nb_outputs
self.is_placeholder = True
super(CustomMultiLossLayer, self).__init__(**kwargs)
def build(self, input_shape=None):
self.log_vars = []
for i in range(self.nb_outputs):
self.log_vars += [self.add_weight(name='log_var' + str(i), shape=(1,), initializer=Constant(0.), trainable=True)]
super(CustomMultiLossLayer, self).build(input_shape)
def multi_loss(self, ys_true, ys_pred):
#print len(ys_true)
assert len(ys_true) == self.nb_outputs and len(ys_pred) == self.nb_outputs
loss = 0
i=0
for y_true, y_pred, log_var in zip(ys_true, ys_pred, self.log_vars):
precision =K.exp(-log_var[0]) ###MODIFICATION HERE
if i==0:
pred_loss = bbce(y_true, y_pred)
term = precision*pred_loss + 0.5 * log_var[0]
#term=tf.Print(term, [term], 'bbce: ')
else:
pred_loss = keras_mse(y_true, y_pred)
#pred_loss=tf.Print(pred_loss, [pred_loss], 'MSE: ')
term = 0.5 * precision * pred_loss + 0.5 * log_var[0]
#term=tf.Print(term, [term], 'MSE: ')
loss+=term
term = 0.
i+=1
return K.mean(loss)
def call(self, inputs):
ys_true = inputs[:self.nb_outputs]
ys_pred = inputs[self.nb_outputs:]
loss = self.multi_loss(ys_true, ys_pred)
self.add_loss(loss, inputs=inputs)
# We won't actually use the output.
return K.concatenate(inputs, -1)
def get_trainable_model(baseline_model):
inp = keras.layers.Input(shape=(224,224,3,), name='inp')
y1_pred, y2_pred = baseline_model(inp)
y1_true=keras.layers.Input(shape=(1,),name='y1_true')
y2_true=keras.layers.Input(shape=(1,),name='y2_true')
out = CustomMultiLossLayer(nb_outputs=2)([y1_true, y2_true, y1_pred, y2_pred])
return Model(input=[inp, y1_true, y2_true], output=out)
# -
keras.backend.clear_session()
model=get_baseline_model(hp_lambda=1., c_list=CONCEPT)
model.summary()
model.load_weights(model_folder+'/best_model.h5')
model.summary()
BATCH_SIZE = 32
from sklearn.metrics import auc
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve, auc
def standard_evaluate(input_,pred_, save_file=None, c_list=CONCEPT):
#import pdb; pdb.set_trace()
y_true = input_[0]
min_idx=0
if 'domain' in c_list:
domain_true = input_[1]
min_idx=1
true_extra_concepts={}
if len(c_list)>min_idx:
for i in range(min_idx, len(c_list)):
true_extra_concepts[i]=input_[min_idx+i]
y_pred = pred_[0]
if 'domain' in c_list:
domain_pred = pred_[1]
min_idx=1
pred_extra_concepts={}
if len(c_list)>min_idx:
for i in range(min_idx, len(c_list)):
pred_extra_concepts[i]=pred_[min_idx+i]
val_acc = my_accuracy_np(y_true, y_pred)
if 'domain' in c_list:
val_acc_d = accuracy_domain(domain_true, domain_pred)
val_r2={}
val_mse={}
if len(c_list)>1:
for i in range(1, len(c_list)):
val_r2[i] = r_square_np(true_extra_concepts[i], pred_extra_concepts[i])
val_mse[i] = compute_mse(true_extra_concepts[i], pred_extra_concepts[i])
extra_string=''
if len(c_list)>1:
for i in range(1, len(c_list)):
extra_string=extra_string+" {}: r2 {}, mse {}; ".format(i, val_r2[i], val_mse[i])
#print("Acc: {}, Acc domain: {}\n".format(val_acc, val_acc_d)+extra_string)
save_file=None
if save_file is not None:
if 'domain' in c_list:
save_file.write("Val acc: {}, acc_domain: {}\n".format(val_acc, val_acc_d)+extra_string)
else:
save_file.write("Val acc: {}, \n".format(val_acc)+extra_string)
if 'domain' in c_list:
return y_true, domain_true, true_extra_concepts, y_pred, domain_pred, pred_extra_concepts
else:
return y_true, true_extra_concepts, y_pred, pred_extra_concepts
def compute_mse(labels, predictions):
errors = labels - predictions
sum_squared_errors = np.sum(np.asarray([pow(errors[i],2) for i in range(len(errors))]))
mse = sum_squared_errors / len(labels)
return mse
def evaluate_model(d_list, model, batch_size=BATCH_SIZE, test_type='', save_file=None):
batch_size=32
t_gen=DataGenerator(d_list, concept=CONCEPT, batch_size=BATCH_SIZE, data_type=0)
steps=len(d_list)//batch_size
initial_lr = 1e-4
opt = keras.optimizers.SGD(lr=initial_lr, momentum=0.9, nesterov=True)
compile_model(t_m,opt,loss=None,metrics=None)
callbacks = []
y_true=np.zeros(len(d_list))
y_pred=np.zeros((len(d_list),1))
N=0
all_true_domain=[]
all_pred_domain=[]
all_true_extra_cm={}#[]#np.zeros(len(d_list))
all_pred_extra_cm={}#[]#np.zeros(len(d_list))
batch_counter=0
while N<len(d_list):
#print N
input_,labels = t_gen.__getitem__(batch_counter)
pred_ = t_m.predict(input_)
#import pdb; pdb.set_trace()
if 'domain' in CONCEPT:
y_true_batch, d_true, true_ec, y_pred_batch, d_pred, pred_ec = standard_evaluate(labels,pred_)
else:
y_true_batch, true_ec, y_pred_batch, pred_ec = standard_evaluate(labels,pred_)
#maybe some import pdb here
y_true[N:N+len(y_true_batch)]=y_true_batch.reshape(len(y_true_batch))
y_pred[N:N+len(y_pred_batch)]=y_pred_batch.reshape(len(y_pred_batch),1)
if 'domain' in CONCEPT:
all_true_domain.append(d_true)
all_pred_domain.append(d_pred)
for extra_concept in true_ec.keys():
try:
all_true_extra_cm[extra_concept].append(true_ec[extra_concept])
except:
all_true_extra_cm[extra_concept]=[]
all_true_extra_cm[extra_concept].append(true_ec[extra_concept])
for extra_concept in pred_ec.keys():
try:
all_pred_extra_cm[extra_concept].append(pred_ec[extra_concept])
except:
all_pred_extra_cm[extra_concept]=[]
all_pred_extra_cm[extra_concept].append(pred_ec[extra_concept])
N+=len(y_pred_batch)
batch_counter+=1
#import pdb; pdb.set_trace()
y_true=y_true.reshape((len(d_list),1))
#y_pred=y_pred.reshape((len(d_list),1))
acc = my_accuracy(y_true, y_pred).eval(session=tf.Session())
sliced_y_pred = tf.sigmoid(y_pred)
y_pred_rounded = K.round(sliced_y_pred)
acc_sc = accuracy_score(y_pred_rounded.eval(session=tf.Session()), y_true)
print('accuracy: ', acc_sc)
y_pred = sliced_y_pred.eval(session=tf.Session())
#sliced_y_pred = tf.sigmoid(y_pred)
#y_pred_rounded = K.round(sliced_y_pred)
auc_score=sklearn.metrics.roc_auc_score(y_true,sliced_y_pred.eval(session=tf.Session()))
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(1):
fpr[i], tpr[i], _ = roc_curve(y_true[:, i], y_pred[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(y_true.ravel(), y_pred.ravel())
roc_auc["micro"] = auc_score
plt.figure()
lw = 2
plt.plot(fpr[0], tpr[0], color='darkorange',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc[0])
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic example: {}'.format(roc_auc[0]))
plt.legend(loc="lower right")
plt.show()
if test_type is not None:
auc_record = open('{}/auc_{}.txt'.format(model_folder,test_type), 'w')
auc_record.write('{}'.format(roc_auc[0]))
print('saved in {}/auc_{}.txt'.format(model_folder,test_type))
auc_record.close()
return all_true_domain, all_true_extra_cm, all_pred_domain, all_pred_extra_cm
# ## Performance on Main Task
# ##### How do we do on the patch classification?
#keras.backend.get_session().run(tf.global_variables_initializer())
model.load_weights(model_folder+'/best_model.h5')
t_m = model
#all_true_domain, all_true_extra_cm, all_pred_domain, all_pred_extra_cm=evaluate_model(train_list[:1000],t_m, test_type='train')
#keras.backend.get_session().run(tf.global_variables_initializer())
#t_m.load_weights(model_folder+'/best_model.h5')
t_m=model
all_true_domain_i, all_true_extra_cm_i, all_pred_domain_i, all_pred_extra_cm_i=evaluate_model(test_list,t_m, test_type='internal')
# +
#keras.backend.get_session().run(tf.global_variables_initializer())
#t_m.load_weights(model_folder+'/best_model.h5')
all_true_domain_e, all_true_extra_cm_e, all_pred_domain_e, all_pred_extra_cm_e=evaluate_model(test2_list,t_m, test_type='external')
# -
import pbd; pdb.set_trace()
aucs_i=[]
for i in range(100):
test_list_b=get_bootstrap_sample(test_list, n_samples=len(test_list))
all_cm_i, all_p_cm_i, roc_auc=evaluate_model(test_list_b,t_m, test_type='internal',save_file=None)
aucs_i.append(roc_auc)
print "AUC avg (std): {} ({})".format(np.mean(aucs_i), np.std(aucs_i))
aucs_e=[]
for i in range(100):
test2_list_b=get_bootstrap_sample(test2_list, n_samples=len(test2_list))
all_cm_e, all_p_cm_e, roc_auc=evaluate_model(test2_list_b,t_m, test_type='external')
aucs_e.append(roc_auc)
print "AUC avg (std): {} ({})".format(np.mean(aucs_e), np.std(aucs_e))
# # Performance on Auxiliary tasks
# ## Are we learning the concepts?
def compute_rsquared(labels, predictions):
errors = labels - predictions
sum_squared_errors = np.sum(np.asarray([pow(errors[i],2) for i in range(len(errors))]))
# total sum of squares, TTS
average_y = np.mean(labels)
total_errors = labels - average_y
total_sum_squares = np.sum(np.asarray([pow(total_errors[i],2) for i in range(len(total_errors))]))
#rsquared is 1-RSS/TTS
rss_over_tts = sum_squared_errors/total_sum_squares
rsquared = 1-rss_over_tts
return rsquared
def compute_mse(labels, predictions):
errors = labels - predictions
sum_squared_errors = np.sum(np.asarray([pow(errors[i],2) for i in range(len(errors))]))
mse = sum_squared_errors / len(labels)
return mse
r2_i = compute_rsquared(all_cm_i, all_p_cm_i)
mse_i = compute_mse(all_cm_i, all_p_cm_i)
print 'Internal: ', r2_i, mse_i
r2_e = compute_rsquared(all_cm_e, all_p_cm_e)
mse_e = compute_mse(all_cm_e, all_p_cm_e)
print 'External: ', r2_e, mse_e
test_type='internal'
auc_record = open('{}/concept_metrics_{}.txt'.format(model_folder,test_type), 'w')
auc_record.write('{}, {}'.format(r2_i, mse_i))
auc_record.close()
test_type='external'
auc_record = open('{}/concept_metrics_{}.txt'.format(model_folder,test_type), 'w')
auc_record.write('{}, {}'.format(r2_e, mse_e))
auc_record.close()
r2_t = compute_rsquared(all_cm_t, all_p_cm_t)
mse_t = compute_mse(all_cm_t, all_p_cm_t)
print 'Train: ', r2_t, mse_t
val_r2=np.load('{}/val_r2_log.npy'.format(model_folder))
plt.plot(val_r2)
history=np.load('{}/training_log.npy'.format(model_folder), allow_pickle=True).item()
plt.plot(history['loss'])
plt.plot(history['val_loss'])
f=open('{}/val_by_epoch.txt'.format(model_folder), 'r')
f_l=f.readlines()
val_acc=[]
val_r2=[]
val_mse=[]
for line in f_l:
acc=line.split('Val acc: ')[1].split(', r2')[0]
val_acc.append(acc)
r2=line.split(', r2:')[1].split(', mse:')[0]
mse=line.split(', mse:')[1].split('\n')[0]
val_r2.append(r2)
val_mse.append(mse)
plt.plot(np.asarray(val_acc, dtype=np.float32))
plt.plot(np.asarray(val_r2, dtype=np.float32))
plt.plot(np.asarray(val_mse, dtype=np.float32))
| notebooks/test/standard_multitask_adversarial_bootstrap_test-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Flask Survival Guide
#
# [Flask](https://flask.palletsprojects.com/en/1.1.x/) is a micro web framework powered by Python. It’s API is fairly small, making it easy to learn and simple to use.
#
# But don’t let this fool you, as it’s powerful enough to support enterprise-level applications handling large amounts of traffic.
#
# You can start small with an app contained entirely in one file, then slowly scale up to multiple files and folders in a well-structured manner as your site becomes more and more complex.
#
# It’s an excellent framework to start with, which you’ll be learning in true “Real Python-style”: with hands-on, practical examples that are interesting and fun.
#
# Flask is based on the Werkzeug [WSGI](https://www.fullstackpython.com/wsgi-servers.html) toolkit and [Jinja2](https://jinja.palletsprojects.com/en/2.11.x/) template engine.
#
# In order follow easily this tutorial there are some prerequisites:
#
# * Good knowledge of Linux **command line**.
# * Experience with **python**.
# * Hands-on **[HTML](https://www.tutorialspoint.com/html/index.htm)**.
#
# Some system requirements:
#
# * Python 2.7.x or up (we will use 3.6.x or up)
# * pip
# * Virtual environment
#
#
#
#
| Bonus_resources/deployment/1.Flask/1.Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Histograms
#
# **Teammates: <NAME>, <NAME>, <NAME>**
#
# ### What is a Histogram?
# A histogram is a visual representation of the frequencies of elements in certain ranges for a variable in a given dataset. It consists of bars whose height give the frequency of the data. The y-axis of a histogram represents the frequency, and the x-axis represents the type of data.
#
# Histograms are used to plot numerical data, for example, grades on an English test. In the histogram below, a bar represents the number of students with a grade that falls within the range of the bin. For example, there are aproximately 12-13 students with test scores between 40 and 50 points.
#
# 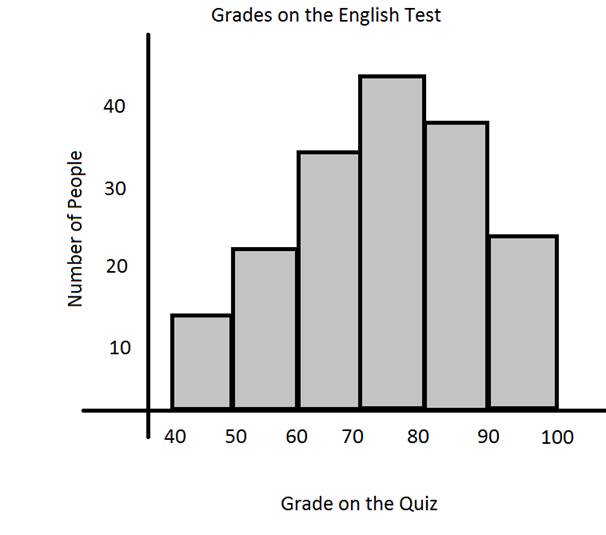
#
#
# ### Key Characteristics
# * Counts frequency of variable
# * Equally spaced bins
# * Visual representation of quantitative / numerical data
#
# ### Purposes of Histograms
# * Easily identify and analyze distribution of variable
# * Identify center, shape, and spread
#
# ### Deficiencies of Histograms
# * Unable to identify singular data points due to aggregated nature
# * Does not provide specific statistics such as the mean, median, or standard deviation
# * Histograms can provide different interpretations for the same dataset depending on how many bins are used
#
# ### Common Mistakes - Bar Charts
# A bar chart and a histogram look very similar to one another. There are a few major differences between the two visualizations
# * Visually, the bars in a histogram are touching each other. The bars in a bar chart have space between them.
# * A histogram is used to graph quantitative data while a bar chart is used for categorical data.
# * Another easy way to distinguish them would be the x axis. Since a histogram is for quantitative data, the x-axis will be a range of numbers. On the other hand, the x-axis of a bar chart will most likely look like a list of words or some other form of categories.
# ## Histograms in Python
#
# ##### There are many libraries that allow for plotting histograms in Python. Three of the most common include Matplotlib, Seaborn and Pandas
#
# Before we can plot in any library, we need a set of data
import numpy as np
#create a set of 100 normally distributed data points, with mean 60 and standard deviation of 10
temps = np.random.normal(60, 10, 100)
# #### Matplotlib
# +
import matplotlib.pyplot as plt
plt.hist(temps)
plt.title("Temperatures")
plt.xlabel("Temperature")
plt.ylabel("Frequency")
plt.show()
# -
# #### Seaborn
import seaborn as sns
#notice that you can use matplotlib to set the titles still
sns.histplot(temps)
plt.title("Temperatures")
plt.xlabel("Temperature")
plt.ylabel("Frequency")
plt.show()
# #### Pandas
#
# In pandas, you must first make a dataframe with the data set. A histogram can then be generated from your dataframe.
import pandas as pd
#once again, use matplotlib to set axis labels and titles
temp = pd.DataFrame(temps)
temp.hist()
plt.title("Temperatures")
plt.xlabel("Temperature")
plt.ylabel("Frequency")
plt.show()
# ### Bins
# The default for all of the methods above is to make a histogram with 10 bins if there is enough data. If you want to change the number of bins, you can do so by the following:
# #### Matplotlib
plt.hist(temps, bins = 6)
plt.title("Temperatures")
plt.xlabel("Temperature")
plt.ylabel("Frequency")
plt.show()
# #### Seaborn
sns.histplot(temps, bins = 8)
plt.title("Temperatures")
plt.xlabel("Temperature")
plt.ylabel("Frequency")
plt.show()
# #### Pandas
temp.hist(bins = 4)
plt.title("Temperatures")
plt.xlabel("Temperature")
plt.ylabel("Frequency")
plt.show()
# ## Exercises
# ### True/False
# 1. One can easily identify what the exact mean and standard deviation are for the dataset based solely on the histogram.
# Answer: False
# 2. A histogram's bins must all be the same size.
# Answer: True
# 3. The shape of a histogram will always look the same if the data is the same.
# Answer: False
# ### Plotting
# Plot the following data using each library. Set 15 bins for matplotlib, 8 bins for seaborn, and 4 for pandas.
#
# Make the title be "Distribution of Salaries" with a y-axis of "Frequency" and an x-axis called "Salary (in $1000)"
l1 = np.random.normal(20, 15, 50)
l2 = np.random.normal(80, 15, 50)
sal = np.concatenate([l1, l2])
# #### Matplotlib
plt.hist(sal, bins=15)
plt.title("Distribution of Salaries")
plt.xlabel("Salary (in $1000)")
plt.ylabel("Frequency")
plt.show()
# #### Seaborn
sns.histplot(sal, bins=8)
plt.title("Distribution of Salaries")
plt.xlabel("Salary (in $1000)")
plt.ylabel("Frequency")
plt.show()
# #### Pandas
sal2 = pd.DataFrame(sal)
sal2.hist(bins=4)
plt.title("Distribution of Salaries")
plt.xlabel("Salary (in $1000)")
plt.ylabel("Frequency")
plt.show()
# What did you notice changed as the number of bins changed? What do you think is the best number of bins to use for this data of the provided options?
# As the bin number decreased, the data became harder to differentiate from each other. The best number of bins for this data out of the 3 options is 8.
# ## Notes
# This gave a good idea of how to use histograms, however it might've been nicer if I could've tested my knowledge along the way instead of all at the end. Also, saying that the default is 10 bins "if there is enough data" seems rather vague and unhelpful. Lastly, using xticks() could've made the histograms better by clearly defining the boundaries of the bins.
| VisualizationLessons/histograms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
from nbdev.showdoc import *
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from htools import add_docstring
# +
# Used in notebook but not needed in package.
from collections import defaultdict
import matplotlib.pyplot as plt
import pandas as pd
from pathlib import Path
import spacy
from htools import assert_raises, InvalidArgumentError
import pandas_htools
# -
FPATH = Path('../data/warbreaker.txt')
with open(FPATH, 'r') as f:
text = f.read()
len(text)
len(text)
c2i = {k: i for i, k in enumerate(sorted(set(text.lower())))}
i2c = list(c2i.keys())
print(c2i)
print(i2c)
nlp = spacy.load('en_core_web_sm', disable=['ner', 'tagger', 'parser'])
def tokenize_one(text):
return [t.text for t in nlp(text)]
def tokenize(texts):
with multiprocessing.Pool() as p:
tokens = p.map(tokenize_one, texts)
return tokens
tokens = tokenize_one(text)
len(tokens)
# ## Issues
#
# - currently assuming all words len >= 4
# - haven't used any padding, so inputs are all different lengths
# - haven't used padding, so outputs are all different lengths
# - character encode? word encode? figure out how to handle
# +
class CharJumbleDS(Dataset):
def __init__(self, tokens, c2i, window=3):
# TO DO: For now, start by assuming all words have len >= 4. Fix later.
self.tokens = [t for t in tokens if len(t) >= 4]
self.c2i = c2i
self.i2c = list(c2i.keys())
self.window = window
self.mid_i = window // 2
def __getitem__(self, idx):
chunk = self.tokens[idx:idx+self.window]
label = self.encode(' '.join(chunk)) # Only needed for seq2seq approach in v3
mid = chunk[self.mid_i]
mid_len = len(mid)
order = np.random.permutation(mid_len - 2) + 1
chunk[self.mid_i] = mid[0] + ''.join(mid[i] for i in order) + mid[-1]
# This version returns the order that was used to permute the original indices.
# Maybe less intuitive but simpler - can always do the conversion in some
# prediction wrapper that doesn't add computation during training.
# return chunk, [0] + list(order) + [mid_len-1]
# This version returns the order to map from the permuted indices to the original
# indices. Intuitive but adds computation and hard-to-read logic.
# return (chunk,
# [0]
# + [k for k, v in sorted(dict(enumerate(order, 1)).items(),key=lambda x: x[1])]
# + [mid_len-1])
# V3: just return whole seq of char indices as input and output.
# Prob more computationally expensive (seq2seq vs multiclass classification)
return self.encode(' '.join(chunk)), label
def encode(self, word_str):
return [self.c2i[char] for char in word_str.lower()]
def decode(self, idx):
return ''.join(self.i2c[i] for i in idx)
def __len__(self):
return len(self.tokens)
def __repr__(self):
return f'CharJumbleDS(len={len(self)})'
# -
ds = CharJumbleDS(tokens, c2i, 4)
ds
for i in range(50):
x, y = ds[i]
print(x)
print(y)
print(ds.decode(x))
print(ds.decode(y))
print()
| notebooks/scratch_self_supervised_text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="mQFbageR7eSe"
# # 3. Plotting for Exploratory data analysis (EDA)
# + [markdown] colab_type="text" id="IYm45v1y7eSj"
# ## Iris Flower dataset
# + [markdown] colab_type="text" id="iW8l_61b7eSk"
# Toy Dataset: Iris Dataset:
# * A simple dataset to learn the basics.
# * 3 flowers of Iris species.
# * 1936 by <NAME>.
# * Petal and Sepal
# * Objective: Classify a new flower as belonging to one of the 3 classes given the 4 features.
# * Importance of domain knowledge.
# * Why use petal and sepal dimensions as features?
# * Why do we not use 'color' as a feature?
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 425} colab_type="code" executionInfo={"elapsed": 1598, "status": "error", "timestamp": 1560498105276, "user": {"displayName": "Applied AI Course", "photoUrl": "https://lh3.googleusercontent.com/-EsJzSyawCkQ/AAAAAAAAAAI/AAAAAAAAEIk/w9ORR2FfvaE/s64/photo.jpg", "userId": "06629147635963609455"}, "user_tz": -330} id="p2oKsGVW7eSm" outputId="1d72c0c2-016b-4c18-8307-1c6f92fb04e9"
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
'''downlaod iris.csv from https://raw.githubusercontent.com/uiuc-cse/data-fa14/gh-pages/data/iris.csv'''
#Load Iris.csv into a pandas dataFrame.
iris = pd.read_csv("iris.csv")
# + colab={} colab_type="code" id="qs8rT4Up7eSz" outputId="376f286f-2e9e-4df1-d145-aba2dc4b5c3c"
# (Q) how many data-points and features?
print (iris.shape)
# + colab={} colab_type="code" id="3RT2Gdlq7eTC" outputId="9e162731-b25e-4169-97eb-a66b9c6df076"
#(Q) What are the column names in our dataset?
print (iris.columns)
# + colab={} colab_type="code" id="4ARdqlVj7eTH" outputId="c8b7c898-35af-4328-e91e-393d5ce361a1"
#(Q) How many data points for each class are present?
#(or) How many flowers for each species are present?
iris["species"].value_counts()
# balanced-dataset vs imbalanced datasets
#Iris is a balanced dataset as the number of data points for every class is 50.
# + [markdown] colab_type="text" id="uExhLJ1-7eTT"
# # (3.2) 2-D Scatter Plot
# + colab={} colab_type="code" id="m8fu7kKy7eTU" outputId="201c4f80-4071-4e22-db53-402547ccb725"
#2-D scatter plot:
#ALWAYS understand the axis: labels and scale.
iris.plot(kind='scatter', x='sepal_length', y='sepal_width') ;
plt.show()
#cannot make much sense out it.
#What if we color the points by thier class-label/flower-type.
# + colab={} colab_type="code" id="fCu1FPvO7eTc" outputId="f6c71a2f-0947-421c-a50a-221df66f4f8c"
# 2-D Scatter plot with color-coding for each flower type/class.
# Here 'sns' corresponds to seaborn.
sns.set_style("whitegrid");
sns.FacetGrid(iris, hue="species", size=4) \
.map(plt.scatter, "sepal_length", "sepal_width") \
.add_legend();
plt.show();
# Notice that the blue points can be easily seperated
# from red and green by drawing a line.
# But red and green data points cannot be easily seperated.
# Can we draw multiple 2-D scatter plots for each combination of features?
# How many cobinations exist? 4C2 = 6.
# + [markdown] colab_type="text" id="2L5YF3Tj7eTl"
# **Observation(s):**
# 1. Using sepal_length and sepal_width features, we can distinguish Setosa flowers from others.
# 2. Seperating Versicolor from Viginica is much harder as they have considerable overlap.
# + [markdown] colab_type="text" id="TdiZ9lLL7eTn"
# ## 3D Scatter plot
#
# https://plot.ly/pandas/3d-scatter-plots/
#
# Needs a lot to mouse interaction to interpret data.
#
# What about 4-D, 5-D or n-D scatter plot?
# + [markdown] colab_type="text" id="ikdQY1fR7eTp"
# # (3.3) Pair-plot
# + colab={} colab_type="code" id="TkuKNKA37eTs" outputId="b0b81e40-40d7-44eb-f446-d5509c9c7d94"
# pairwise scatter plot: Pair-Plot
# Dis-advantages:
##Can be used when number of features are high.
##Cannot visualize higher dimensional patterns in 3-D and 4-D.
#Only possible to view 2D patterns.
plt.close();
sns.set_style("whitegrid");
sns.pairplot(iris, hue="species", size=3);
plt.show()
# NOTE: the diagnol elements are PDFs for each feature. PDFs are expalined below.
# + [markdown] colab_type="text" id="aJAPMfBU7eT1"
# **Observations**
# 1. petal_length and petal_width are the most useful features to identify various flower types.
# 2. While Setosa can be easily identified (linearly seperable), Virnica and Versicolor have some overlap (almost linearly seperable).
# 3. We can find "lines" and "if-else" conditions to build a simple model to classify the flower types.
# + [markdown] colab_type="text" id="LWsvwUkL7eT4"
# # (3.4) Histogram, PDF, CDF
# + colab={} colab_type="code" id="wUvH2M817eT6" outputId="1f577f0d-475c-4710-b6d3-f744f558fa02"
# What about 1-D scatter plot using just one feature?
#1-D scatter plot of petal-length
import numpy as np
iris_setosa = iris.loc[iris["species"] == "setosa"];
iris_virginica = iris.loc[iris["species"] == "virginica"];
iris_versicolor = iris.loc[iris["species"] == "versicolor"];
#print(iris_setosa["petal_length"])
plt.plot(iris_setosa["petal_length"], np.zeros_like(iris_setosa['petal_length']), 'o')
plt.plot(iris_versicolor["petal_length"], np.zeros_like(iris_versicolor['petal_length']), 'o')
plt.plot(iris_virginica["petal_length"], np.zeros_like(iris_virginica['petal_length']), 'o')
plt.show()
#Disadvantages of 1-D scatter plot: Very hard to make sense as points
#are overlapping a lot.
#Are there better ways of visualizing 1-D scatter plots?
# + colab={} colab_type="code" id="gjZTt3WS7eUD" outputId="f71f78ef-2d20-408e-f187-707e5b31b3e1"
sns.FacetGrid(iris, hue="species", size=5) \
.map(sns.distplot, "petal_length") \
.add_legend();
plt.show();
# + colab={} colab_type="code" id="zHGF-B3h7eUK" outputId="9f613f86-35a8-47f9-94e3-3fd74a25588f"
sns.FacetGrid(iris, hue="species", size=5) \
.map(sns.distplot, "petal_width") \
.add_legend();
plt.show();
# + colab={} colab_type="code" id="eKMrbu917eUU" outputId="8d2132d0-66ae-484d-a70f-2ba2574a9989"
sns.FacetGrid(iris, hue="species", size=5) \
.map(sns.distplot, "sepal_length") \
.add_legend();
plt.show();
# + colab={} colab_type="code" id="RyusT9e47eUb" outputId="0cf8dda7-ece7-4fcc-f9a6-7306c0525ffa"
sns.FacetGrid(iris, hue="species", size=5) \
.map(sns.distplot, "sepal_width") \
.add_legend();
plt.show();
# + colab={} colab_type="code" id="sUTbOpd67eUg"
# Histograms and Probability Density Functions (PDF) using KDE
# How to compute PDFs using counts/frequencies of data points in each window.
# How window width effects the PDF plot.
# Interpreting a PDF:
## why is it called a density plot?
## Why is it called a probability plot?
## for each value of petal_length, what does the value on y-axis mean?
# Notice that we can write a simple if..else condition as if(petal_length) < 2.5 then flower type is setosa.
# Using just one feature, we can build a simple "model" suing if..else... statements.
# Disadv of PDF: Can we say what percentage of versicolor points have a petal_length of less than 5?
# Do some of these plots look like a bell-curve you studied in under-grad?
# Gaussian/Normal distribution.
# What is "normal" about normal distribution?
# e.g: Hieghts of male students in a class.
# One of the most frequent distributions in nature.
# + colab={} colab_type="code" id="5w3mEOUR7eUk" outputId="bc3aeb10-a564-449e-8ca7-9b60d268d01b"
# Need for Cumulative Distribution Function (CDF)
# We can visually see what percentage of versicolor flowers have a
# petal_length of less than 5?
# How to construct a CDF?
# How to read a CDF?
#Plot CDF of petal_length
counts, bin_edges = np.histogram(iris_setosa['petal_length'], bins=10,
density = True)
pdf = counts/(sum(counts))
print(pdf);
print(bin_edges);
cdf = np.cumsum(pdf)
plt.plot(bin_edges[1:],pdf);
plt.plot(bin_edges[1:], cdf)
counts, bin_edges = np.histogram(iris_setosa['petal_length'], bins=20,
density = True)
pdf = counts/(sum(counts))
plt.plot(bin_edges[1:],pdf);
plt.show();
# + colab={} colab_type="code" id="KDX4yFj17eUq" outputId="5926d33d-e5fd-48eb-be13-43b5166ab05e"
# Need for Cumulative Distribution Function (CDF)
# We can visually see what percentage of versicolor flowers have a
# petal_length of less than 1.6?
# How to construct a CDF?
# How to read a CDF?
#Plot CDF of petal_length
counts, bin_edges = np.histogram(iris_setosa['petal_length'], bins=10,
density = True)
pdf = counts/(sum(counts))
print(pdf);
print(bin_edges)
#compute CDF
cdf = np.cumsum(pdf)
plt.plot(bin_edges[1:],pdf)
plt.plot(bin_edges[1:], cdf)
plt.show();
# + colab={} colab_type="code" id="TjHpJqSz7eUw" outputId="fd91266a-0ad3-4ef2-8877-0797528ba053"
# Plots of CDF of petal_length for various types of flowers.
# Misclassification error if you use petal_length only.
counts, bin_edges = np.histogram(iris_setosa['petal_length'], bins=10,
density = True)
pdf = counts/(sum(counts))
print(pdf);
print(bin_edges)
cdf = np.cumsum(pdf)
plt.plot(bin_edges[1:],pdf)
plt.plot(bin_edges[1:], cdf)
# virginica
counts, bin_edges = np.histogram(iris_virginica['petal_length'], bins=10,
density = True)
pdf = counts/(sum(counts))
print(pdf);
print(bin_edges)
cdf = np.cumsum(pdf)
plt.plot(bin_edges[1:],pdf)
plt.plot(bin_edges[1:], cdf)
#versicolor
counts, bin_edges = np.histogram(iris_versicolor['petal_length'], bins=10,
density = True)
pdf = counts/(sum(counts))
print(pdf);
print(bin_edges)
cdf = np.cumsum(pdf)
plt.plot(bin_edges[1:],pdf)
plt.plot(bin_edges[1:], cdf)
plt.show();
# + [markdown] colab_type="text" id="1JykhrwO7eUz"
# # (3.5) Mean, Variance and Std-dev
# + colab={} colab_type="code" id="7rhG9mB17eU0" outputId="d6383c6f-1007-4876-9907-ef4dc0a58982"
#Mean, Variance, Std-deviation,
print("Means:")
print(np.mean(iris_setosa["petal_length"]))
#Mean with an outlier.
print(np.mean(np.append(iris_setosa["petal_length"],50)));
print(np.mean(iris_virginica["petal_length"]))
print(np.mean(iris_versicolor["petal_length"]))
print("\nStd-dev:");
print(np.std(iris_setosa["petal_length"]))
print(np.std(iris_virginica["petal_length"]))
print(np.std(iris_versicolor["petal_length"]))
# + [markdown] colab_type="text" id="abmP92Sn7eU4"
# # (3.6) Median, Percentile, Quantile, IQR, MAD
# + colab={} colab_type="code" id="cICgORTF7eU5" outputId="bf2a36d9-e954-4ce1-ce5e-69c8999a743b"
#Median, Quantiles, Percentiles, IQR.
print("\nMedians:")
print(np.median(iris_setosa["petal_length"]))
#Median with an outlier
print(np.median(np.append(iris_setosa["petal_length"],50)));
print(np.median(iris_virginica["petal_length"]))
print(np.median(iris_versicolor["petal_length"]))
print("\nQuantiles:")
print(np.percentile(iris_setosa["petal_length"],np.arange(0, 100, 25)))
print(np.percentile(iris_virginica["petal_length"],np.arange(0, 100, 25)))
print(np.percentile(iris_versicolor["petal_length"], np.arange(0, 100, 25)))
print("\n90th Percentiles:")
print(np.percentile(iris_setosa["petal_length"],90))
print(np.percentile(iris_virginica["petal_length"],90))
print(np.percentile(iris_versicolor["petal_length"], 90))
from statsmodels import robust
print ("\nMedian Absolute Deviation")
print(robust.mad(iris_setosa["petal_length"]))
print(robust.mad(iris_virginica["petal_length"]))
print(robust.mad(iris_versicolor["petal_length"]))
# + [markdown] colab_type="text" id="6OHiqoR-7eU9"
# # (3.7) Box plot and Whiskers
# + colab={} colab_type="code" id="s4ZG6dZw7eU_" outputId="e71ac9d1-fb27-4825-e75e-21d8fbc3f072"
#Box-plot with whiskers: another method of visualizing the 1-D scatter plot more intuitivey.
# The Concept of median, percentile, quantile.
# How to draw the box in the box-plot?
# How to draw whiskers: [no standard way] Could use min and max or use other complex statistical techniques.
# IQR like idea.
#NOTE: IN the plot below, a technique call inter-quartile range is used in plotting the whiskers.
#Whiskers in the plot below donot correposnd to the min and max values.
#Box-plot can be visualized as a PDF on the side-ways.
sns.boxplot(x='species',y='petal_length', data=iris)
plt.show()
# + [markdown] colab_type="text" id="3S8dI16V7eVC"
# # (3.8) Violin plots
# + colab={} colab_type="code" id="ha1SwMC47eVE" outputId="629aa3d3-15b9-473b-bdd8-95d115bb0634"
# A violin plot combines the benefits of the previous two plots
#and simplifies them
# Denser regions of the data are fatter, and sparser ones thinner
#in a violin plot
sns.violinplot(x="species", y="petal_length", data=iris, size=8)
plt.show()
# + [markdown] colab_type="text" id="axQROeiL7eVK"
# # (3.9) Summarizing plots in english
# * Exaplain your findings/conclusions in plain english
# * Never forget your objective (the probelm you are solving) . Perform all of your EDA aligned with your objectives.
#
# # (3.10) Multivariate probability density, contour plot.
# + colab={} colab_type="code" id="C9HUB0R37eVQ"
#2D Density plot, contors-plot
sns.jointplot(x="petal_length", y="petal_width", data=iris_setosa, kind="kde");
plt.show();
# + colab={} colab_type="code" id="Sy-AO9SZ7eVj"
| Day 13,14 - EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import warnings
warnings.simplefilter(action='ignore')
# -
df = pd.read_csv('data.csv')
df.head()
df.isna().sum()
df.dropna(inplace=True)
df = df[~(df['price'] == 'Price on Request')]
df1 = pd.DataFrame()
df1['Type'] = ''
df1['Type'] = df['location'].apply(lambda x: 'House' if('house' in x.lower()) else x)
df1['Type'] = df1['Type'].apply(lambda x: 'Apartment' if('apartment' in x.lower()) else x)
df1['Type'] = df1['Type'].apply(lambda x: 'Villa' if('villa' in x.lower()) else x)
df1['Type'] = df1['Type'].apply(lambda x: 'Apartment' if('independent' in x.lower()) else x)
df1['Type'].unique()
df1['Location'] = [x.split('in')[1].strip() for x in df['location']]
df1['Beds'] = [int(x.split('B')[0].strip()) for x in df['beds']]
# +
df1.rename(columns={'price':'Price', 'area':'Area','beds':'Beds','location':'Location'},inplace=True)
for i,x in enumerate(df['price']):
#print(type(x))
if x != 'Price on Request':
if 'Cr' in x:
df1['Price'][i] = x.split(' Cr')[0]
elif 'Lac' in x:
df1['Price'][i] = x.split(' Lac')[0]
else:
df1['Price'][i] = x
else:
df1['Price'][i] = x
# -
df1 = df1[~(df1['Price'] == '₹ 25,000\n₹ 18/sq.ft.')]
df1 = df1[~(df1['Price'] == '₹ 30,000\n₹ 1/sq.ft.')]
df1['Price'] = df1['Price'].apply(lambda x : x.strip(' Lac\n') if('Lac\n' in x) else x)
df1['Price'] = df1['Price'].apply(lambda x : x.strip(' Cr'))
df1['Price'].unique()
df = df[~(df['price'] == '₹ 25,000\n₹ 18/sq.ft.')]
df = df[~(df['price'] == '₹ 30,000\n₹ 1/sq.ft.')]
df1.Price.unique()
# +
for i,x in enumerate(df1['Price']):
if '-' in x:
print('1')
df1['Price'][i] = float(x.split('-')[0]),float(x.split('-')[1])
else:
df1['Price'][i] = float(x)
# -
df1.Price
for i,x in enumerate(df1['Price']):
if type(x) == tuple:
df1['Price'][i] = np.mean(x)
elif type(x) == str:
df1['Price'][i] = np.float(x)
else:
df1['Price'][i] = np.mean(x)
df1['Price'] = df1['Price'].apply(lambda x : float(x) if(type(x) is str) else(x))
df1['Price'] = df1['Price'].apply(lambda x : round(x*100,2) if(x < 10) else(x))
df1['Price'].describe()
df1['Area'] = [x.split('sq')[0] for x in df['area']]
for i,x in enumerate(df1['Area']):
#print(x)
if '-' in x:
temp1 = x.strip().split('-')[0].replace(',','')
temp2 = x.strip().split('-')[1].replace(',','')
temp1 = np.float(temp1)
temp2 = np.float(temp2)
avg = np.mean((temp1,temp2))
df1['Area'][i] = avg
else:
temp = x.strip().replace(',','')
df1['Area'][i] = np.float(temp)
df1[df1['Area'] == '3,230 ']['Area'].index
df1.Area[528] = 3320
df1[df1['Area'] == '600 ']['Area'].index
df1.Area[529] = 600
df1.describe()
type(df1['Area'][3])
df1['Area'].describe()
df1['Beds'].describe()
df1[df1.Price < 200].hist()
df1.shape
df1.Area.unique()
df1 = df1[~(df1.Area == 120000)]
df1[(df1.Area == 12000)]
df1['PricePerSqft'] = df1['Price']/df1['Area']
df1.drop('Price_per_sqft',1,inplace=True)
df1.head()
def remove_pps_ourliers(df):
df_out = pd.DataFrame()
for key, subdf in df.groupby('Location'):
avg = np.mean(subdf['PricePerSqft'])
stdev = np.std(subdf['PricePerSqft'])
reduced_df = subdf[(subdf['PricePerSqft'] > (avg-stdev)) & (subdf.PricePerSqft < (avg+stdev)) ]
df_out = pd.concat([df_out,reduced_df],ignore_index=True)
return df_out
df2 = remove_pps_ourliers(df1)
df2.shape
df2.Price.hist(bins=25)
df2.describe()
df1.describe()
df2.to_csv('data_2_merge.csv',index=False)
df1['Type'].size
| Dataset/data_99acres/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import smtplib
from email.message import EmailMessage
from string import Template
from pathlib import Path
html = Template(Path('index.html').read_text())
email = EmailMessage()
email['from'] = 'Sandeep'
email['to'] = '<EMAIL>'
email['subject'] = 'You have worn a Million Dollars!!!'
email.set_content(html.substitute({'name': 'TinTin'}), 'html')
with smtplib.SMTP(host='smtp.gmail.com', port=587) as smtp:
smtp.ehlo()
smtp.starttls()
smtp.set_debuglevel(True)
smtp.login('<EMAIL>', 'abcdefgh1jkl')
smtp.send_message(email)
| NoteBooks/mail_html.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Automatic sleep staging
#
# This notebook demonstrates how to perform automatic sleep staging of polysomnography data in YASA. For more details, make sure to read the [eLife publication](https://elifesciences.org/articles/70092).
#
# Please install the latest version of YASA first with: `pip install --upgrade yasa`.
import mne
import yasa
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# ## Data loading
#
# The automatic sleep staging function requires that the data are loaded using the [MNE Python package](https://mne.tools/stable/index.html). For instance, if your polysomnography data are stored in the standard European Data Fomat (.edf), you can use [this function](https://mne.tools/stable/generated/mne.io.read_raw_edf.html?highlight=read_raw_edf#mne.io.read_raw_edf) to load the data into Python.
# Let's load a nap recording, directly as an MNE Raw object
raw = mne.io.read_raw_fif('sub-02_mne_raw.fif', preload=True, verbose=False)
print('The channels are:', raw.ch_names)
print('The sampling frequency is:', raw.info['sfreq'])
raw
# Let's now load the human-scored hypnogram, where each value represents a 30-sec epoch.
hypno = np.loadtxt('sub-02_hypno_30s.txt', dtype=str)
hypno
# ## Sleep staging
#
# Automatic sleep stages classification can be done since YASA 0.4.0 using the [SleepStaging](https://raphaelvallat.com/yasa/build/html/generated/yasa.SleepStaging.html#yasa.SleepStaging) class. Make sure to read the [documentation](https://raphaelvallat.com/yasa/build/html/generated/yasa.SleepStaging.html#yasa.SleepStaging), which explains how the algorithm works.
# We first need to specify the channel names and, optionally, the age and sex of the participant
# - "raw" is the name of the variable containing the polysomnography data loaded with MNE.
# - "eeg_name" is the name of the EEG channel, preferentially a central derivation (e.g. C4-M1). This is always required to run the sleep staging algorithm.
# - "eog_name" is the name of the EOG channel (e.g. LOC-M1). This is optional.
# - "eog_name" is the name of the EOG channel (e.g. EMG1-EMG3). This is optional.
# - "metadata" is a dictionary containing the age and sex of the participant. This is optional.
sls = yasa.SleepStaging(raw, eeg_name="C4", eog_name="EOG1", emg_name="EMG1", metadata=dict(age=21, male=False))
# Getting the predicted sleep stages is now as easy as:
y_pred = sls.predict()
y_pred
# What is the accuracy of the prediction, compared to the human scoring
accuracy = (hypno == y_pred).sum() / y_pred.size
print("The overall agreement is %.3f" % accuracy)
# **Stage probabilities and confidence of the algorithm at each epoch**
# What are the predicted probabilities of each sleep stage at each epoch?
sls.predict_proba()
# Plot the predicted probabilities
sls.plot_predict_proba();
# From the probabilities, we can extract a confidence level (ranging from 0 to 1) for each epoch.
confidence = sls.predict_proba().max(1)
confidence
# **Exporting to a CSV file**
# +
# Let's first create a dataframe with the predicted stages and confidence
df_pred = pd.DataFrame({'Stage': y_pred, 'Confidence': confidence})
df_pred.head(6)
# Now export to a CSV file
# df_pred.to_csv("my_hypno.csv")
# -
# **Applying the detection using only a single EEG derivation**
# Using just an EEG channel (= no EOG or EMG)
y_pred = yasa.SleepStaging(raw, eeg_name="C4").predict()
y_pred
| notebooks/14_automatic_sleep_staging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # cGAN Generate Synthetic Data for German Dataset
# CTGAN model is based on the GAN-based Deep Learning data synthesizer
# +
from implementation_functions import *
import six
import sys
sys.modules['sklearn.externals.six'] = six
import mlrose
import pandas as pd
import numpy as np
from prince import FAMD #Factor analysis of mixed data
from aif360.metrics import BinaryLabelDatasetMetric
from sklearn.model_selection import train_test_split
from sklearn.metrics import silhouette_score
import matplotlib.pyplot as plt
import skfuzzy as fuzz
# -
data_name = "german"
dataset_orig, privileged_groups, unprivileged_groups = aif_data(data_name, False)
sens_attr = ['age', 'sex']
decision_label = 'credit'
fav_l = 1
unfav_l = 0
# +
orig_df, num_list, cat_list = preprocess(dataset_orig, sens_attr, decision_label)
# The list of sub-group sizes in the dataset (to monitor the dist. of sub-groups)
orig_df['sub_labels'].value_counts()
# -
import time
start_time = time.time()
print("--- %s seconds ---" % (time.time() - start_time))
# Train-test split WITH stratification
X = orig_df.loc[:, orig_df.columns != decision_label]
y = orig_df.loc[:, orig_df.columns == decision_label].values
X_train, X_test, y_train, y_txest = train_test_split(X, y, test_size=0.30,
shuffle=True,
stratify=X['sub_labels'])
X_train_new = X_train.drop(['age', 'sex', 'sub_labels'], axis=1)
# +
#X_train_new['class_labels'] = y_train
# -
# # Here we start the GAN work
from sdv.tabular import CTGAN
model = CTGAN()
from sdv.tabular import CTGAN
model = CTGAN()
start_time = time.time()
model.fit(X_train)
print("--- %s seconds ---" % (time.time() - start_time))
# + active=""
# model.save('my_fariness_German_V1.pkl')
# -
loaded = CTGAN.load('my_fariness_German_V1.pkl')
# Generating X_train, the following cell shows
# +
available_rows = {}
for row_count in range(8):
available_rows[row_count] = X_train["sub_labels"].value_counts()[row_count]
target_rows = max(available_rows.values())
max_label = max(available_rows, key=available_rows.get)
print(target_rows)
print(max_label)
# -
main_df = pd.DataFrame()
# +
for key, value in available_rows.items():
if int(key) != int(max_label):
conditions = {
"sub_labels" : int(key),
}
needed_rows = target_rows - value
main_df = pd.concat([main_df, loaded.sample(needed_rows, conditions=conditions)])
print(len(main_df.index))
# -
main_df
# + active=""
# concatanate the y_test as a new column with the name 'class_labels' to main_df
# -
print(type(main_df))
main_df.to_csv('german_synth.csv', index = False, header=True)
# not to use any of the sensetive attributes ...trying to be blind by not using any of these obvious attributes ... we delete the sent att .. but still there is bias. Deleting them, will not remove the bias, but as a first step we are tryring to delete the traces of bias by deleteing sent and the techq
# +
# Keep the subgroup labels to append them back later
keep_sub_l = X_train['sub_labels']
# Required drops for the GERMAN dataset (THIS DF CREATION IS A MUST)
X_train_new = X_train.drop(['age', 'sex', 'sub_labels'], axis=1)
# Get the idx of categ and numeric columns again due to the column drops above
num_list, cat_list = type_lists(X_train_new)
# -
# Type the desired classifier to train the classification models with model obj
clf = GradientBoostingClassifier()
baseline_stats, cm, ratio_table, preds = baseline_metrics(clf, X_train, X_test,
y_train, y_test, sens_attr,
fav_l, unfav_l)
X_train_new
# + active=""
# # Dimensionality reduction for big datasets with FAMD
# X_train_new['sub_labels'] = keep_sub_l
#
# famd = FAMD(n_components=2, random_state = 42)
# famd.fit(X_train_new.drop('sub_labels', axis=1))
# X_train_reduc = famd.transform(X_train_new)
# #plotting the reduced dimensions
# ax = famd.plot_row_coordinates(X_train_new,
# color_labels=['sub-labels {}'.format(t) for t in X_train_new['sub_labels']])
# # X_train_red = famd.partial_row_coordinates(X_train_new)
# # famd.explained_inertia_
# # ax = famd.plot_partial_row_coordinates(X_train_new,
# # color_labels=['sub-labels {}'.format(t) for t in X_train_new['sub_labels']])
#
# # Delete the subgroup label column again if dimensionality reduction is used
# X_train_new = X_train_new.drop(['sub_labels'], axis=1)
# -
clf = RandomForestClassifier()
baseline_stats, cm, ratio_table, preds = baseline_metrics(clf, X_train, X_test,
y_train, y_test, sens_attr,
fav_l, unfav_l)
# -str2: clusters membership-->each sample for cluster..
# -str3: all of the clusters, each trained clusters * pp
# + active=""
# print(baseline_stats)
# -
print(type(main_df))
print(type(X_test))
print(type(clf))
print(ratio_table)
test_sublabels = X_test['sub_labels']
X_test_n = X_test.drop(['age', 'sex','sub_labels'], axis=1)
num_list, cat_list = type_lists(X_test_n)
# Predicting the test sets based on strategy 1
X_test_pred1 = predict_whole_set(clf, main_df, X_test_n)
# print(dict_german["month"])
metrics_table1, cm1, ratio_t1 = metrics_calculate(X_test, X_test_pred1, y_test,
sens_attr, fav_l, unfav_l)
metrics_table2, cm2, ratio_t2 = metrics_calculate(X_test, X_test_pred2, y_test,
sens_attr, fav_l, unfav_l)
metrics_table3, cm3, ratio_t3 = metrics_calculate(X_test, X_test_pred3, y_test,
sens_attr, fav_l, unfav_l)
#outputs from strategy 1
print(metrics_table1)
print("Confusion Matrix:", cm1)
print(ratio_t1)
#outputs from strategy 2
print(metrics_table2)
print("Confusion Matrix:", cm2)
print(ratio_t2)
#outputs from strategy 3
print(metrics_table3)
print("Confusion Matrix:", cm3)
print(ratio_t3)
| cGAN-German_october26.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Exercise
#
#
# Download the [gene expression cancer RNA-Seq Data Set](https://archive.ics.uci.edu/ml/datasets/gene+expression+cancer+RNA-Seq) from the UCI Machine Learning Repository. This dataset contains gene expressions of patients having different types of tumor: BRCA, KIRC, COAD, LUAD and PRAD.
#
# 1. Perform at least two different types of clustering. Check the clustering module in [scikit-learn](https://scikit-learn.org/stable/modules/clustering.html) and/or [scipy](https://docs.scipy.org/doc/scipy/reference/cluster.html). Do some preprocessing if necessary (check attributes, check for NaN values, perform dimension reduction, etc).
# 2. Evaluate clustering performance using the [adjusted Rand score.](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.adjusted_rand_score.html#sklearn.metrics.adjusted_rand_score) Try to achieve a clustering with Rand score higher than 0.985.
# 3. Try some manifold learning methods for projecting the data to 2 dimensional space for visualization. Read the documentation for [Multidimensional scaling](https://scikit-learn.org/stable/modules/manifold.html#multidimensional-scaling) and [Stochastic neighbourhood embedding](https://scikit-learn.org/stable/modules/manifold.html#t-sne) and try them. You may try other methods as well.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from sklearn.metrics import adjusted_rand_score
# add more imports if necessary
# from sklearn.cluster import ...
# from scipy.cluster import ...
| homework/week04_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tipos de dados
#
# Dados comuns do nosso dia-a-dia podem ser classificados em 4 tipos:
#
# - Inteiros: números sem casa decimal: 10, 5, -4
# /
# - Numéricos -
# / \
# / - Reais: números com casa decima: 3.14, 1075.18, 0.0
# - Dados -
# \ - Lógicos: True ou False
# \ /
# - Não numéricos -
# \
# - Texto (cadeia de caracteres): "Python", "<NAME>", "34567-890"
#
# O Python possui tipos primitivos para cada um desses tipos de dados. São eles o int, float, bool e o str.
#
# # Operadores aritméticos
#
#
# | Operador | Significado | Exemplo |
# |:---|:---|:---|
# | + | Adição | a = 3 + 2 |
# | - | Subtração | b = 7 - a |
# | * | Multiplicação | c = a * b |
# | / | Divisão | d = 10/3 |
# | // | Quociente | e = 10//3 |
# | % | Resto | f = 10%3 |
# | ** | Exponenciação | g = 2 ** 3 |
# | ** | Radiciação | h = 16 ** (1/2) |
#
# ## Precedência de Operadores
#
# 1. Parênteses
# 2. Potenciação
# 3. Multiplicação e divisão
# 4. Soma e substração
# Adição
3 + 2
# Subtração
7-5
# +
# Multiplicação
# -
2 * 3
3 * 3.5
4 * 5.0
# Divisão
9 / 3
10 / 3
9 / 3.0
# Divisão - calculando o coeficiente
5 // 2
5.0 // 2
4 // 1.2
# Divisão - calculando o resto
5 % 2
9 % 2.5
# Potenciação: (1) Exponenciação e (2) Radiciação
# (1) Exponenciação
3 ** 2
2 ** 10
# (2) Radiciação
# Raiz quadrada de 9
9 ** (1/2)
# Raiz quadrada de 16 (elevando a 0.5 ao invés da fração 1/2)
16 ** 0.5
# Raiz cúbica de 27
27 ** (1/3)
# # Exercício de fixação
#
# Converter as equações em expressões aritméticas válidas no Python.
#
# Eq1.
#
# $ 2+3*4/5-6 $
2 + 3 * 4 / 5 - 6
# Agora a mesma equação acima, mas alterando a precedência para que as operações de soma e subtração sejam executadas antes das outras. _Dica: use parênteses._
(2 + 3) * 4 / (5 - 6)
# Eq2. $ \sqrt{\frac{(2-3)^{2}}{4}}$
((2-3)**2/4)**(1/2)
# # Variáveis e o comando de atribuição
#
# ## Conceito de variável
# O que é uma [variável](https://pt.wikipedia.org/wiki/Variável_\(programação\) "Conceito de variável no Wikipedia")? De forma bem simples e objetiva, uma **variável** é um espaço na memória (*memória RAM*) do seu computador, utilizada por um programa para armazenar dados.
#
#
# ## Identificadores
#
# Uma variável precisa necessariamente de um nome, o qual chamamos de **identificador**. Em Python, um identificador é formado apenas por letras (_de A a Z, maiúsculas ou minúsculas, sem acentuação_), dígitos (_0 a 9_) e o _underscore_ ( \_ ). Um identificador deve começar **obrigatoriamente** com uma letra ou *underscore*, e não pode conter espaços, caracteres especiais (@, !, \*, -, &, ˜, etc) ou caracteres acentuados (é, à, ã, ç, í, ö, ü, û, etc..).
#
# Portanto, os caracteres permitidos em identificadores são:
#
# `ABCDEFGHIJKLMNOPQRSTUVWXYZabcdefghijklmnopqrstuvwxyz_0123456789`
#
# ### Identificadores válidos e inválidos
# Portanto podemos considerar os seguintes identificadores como nomes válidos para variáveis:
#
# * nome
# * Salario
# * NUMERO10
# * i23ER21
# * \_data\_de\_nascimento\_
#
# Já os identificadores abaixo não são nomes válidos para variáveis:
# * nome do aluno (*identificadores não podem conter espaços*)
# * Salário (*contém uma letra acentuada*)
# * 10NUMERO (*começa com dígito*)
# * !23ER21 (_possui um caractere especial **!**_)
# * data-de-nascimento (_possui um caractere especial **-**_)
#
# ### Identificadores significativos e não significativos
#
# Além de serem válidos, como nos exemplos acima, os **identificadores** precisam ter nomes significativos. Ou seja, ao ver um identificador, o programador deve conseguir perceber que dado está armazenado na variável.
#
# Os identificadores abaixo, embora sejam válidos, não tem nomes significativos:
#
# * i23ER1
# * _123
# * BATMAN
# * T
#
# Embora todos os identificadores acima sejam válidos, não são significativos. Que informação será armazenada nas variáveis _i23ER21_, _\_123_, _BATMAN_ ou _T_? É difícil descobrir.
#
# Os identificadores abaixo são válidos e tem nomes significativos:
#
# * salario
# * idade
# * nome
# * nota
# * complementoenderecoparacorrespondencia
#
# Uma observação importante: para que um nome seja significativo, ele não precisa ser tão longo quanto _complementoenderecoparacorrespondencia_. Por exemplo, o que a variável `dtnasc` armazena? Se você pensou em uma data de nascimento, você acertou. Não precisamos de identificadores longos como *data\_de\_nascimento* para criar identificadores significativos. O ideal é utilizar identificadores mais curtos e fáceis de memorizar e utilizar como _dtnasc_.
#
# Por fim, uma última informação. Python faz diferenciação entre letras maiúsculas e minúsculas em identificadores. Portanto, os identificadores _nota, Nota e NOTA_ representam três variáveis diferentes.
#
# ## Criando variáveis
#
# Variáveis em Python não são **declaradas** como em outras linguagens *(ex: Pascal, C, C++, Java, C#, etc...)* mas **criadas** a partir da atribuição de um valor, uma variável, uma expressão ou um objeto. O comando de atribuição, representado pelo sinal de igualdade (**=**), é o responsável por atribuir (ou armazenar) um valor em uma variável.
#
# Vejamos agora alguns exemplos:
idade = 47
nome = '<NAME>'
pi = 3.14159
aprovado = True
# # Verificando tipos de dados e variáveis
#
# ## type()
#
# O Python fornece o método **type()** que retorna o tipo de dados e variáveis. Seu uso é muito simples, basta colocar um objeto, o nome de uma variável ou um valor entre parênteses.
type(idade)
type(2)
type(nome)
type('python')
type(pi)
type(aprovado)
# ## isinstance()
#
# Além do método **type()** que retorna o tipo de um objeto, Python fornece o método **isinstance()** que permite verificar se um objeto ou variável (primeiro parâmetro da função) é uma instância de uma classe ou é de um determinado tipo.
isinstance(5, int)
isinstance(5, str)
isinstance(5, float)
isinstance('Python', str)
isinstance('Python', int)
# # Imprimindo valores e variáveis
#
# Python fornece o método print para impressão de valores e variáveis.
#
# Algumas células acima, nós criamos as variáveis _nome, idade, pi_ e _aprovado_ e usamos o comando de atribuição para armazenar alguns valores. Que tal imprimir esse valores?
print('nome')
# Perceba que ao executar o comando *print('nome')*, a palavra **nome** foi impressa.
print(nome)
# Já nesse exemplo, ao executar o comando *print(nome)*, o resultado apresentado foi o nome `<NAME>`.
#
# A diferença é que no primeiro exemplo pedimos para o Python imprimir o **texto** `'nome'`, enquanto que no segundo pedimos ao Python para imprimir o conteúdo da variável `nome`. A diferença está no apóstrofo (ou, como alguns costumam chamar, aspas simples). A presença de apóstrofos indica um texto, enquanto que a falta deles indica que se deseja imprimir uma variável.
print(idade)
print(pi)
print(aprovado)
# Quando usamos o método print com uma variável entre parênteses, ele imprime o conteúdo dessa variável, ou seja, o valor que está armazenado dentro dela.
#
# Nós podemos montar mensagens um pouco mais sofisticadas juntando texto e variáveis em um único comando print(). Para isso, vamos usar o que chamamos de concatenação, ou seja, vamos juntar texto e variáveis com o símbolo +. Veja os exemplos abaixo.
print('Nome: '+nome)
# Podemos também usar a vírgula ao invés do símbolo +
print('Nome:',nome)
# A forma mais utilizada na versão 3 do Python, entretanto, é usar o método format.
print('Meu nome é {} e tenho {} anos'.format(nome, idade))
# # Transformando equações matemáticas em expressões aritméticas
#
# ### Cálculo da área e perímetro do retângulo
#
# $area = base \times altura$
#
# $ perímetro = 2 \times (base + altura)$
base = 7
altura = 8
area = base * altura
perimetro = 2 * (base + altura)
print(area)
# ### Cálculo da área e comprimento da circunferência
#
# $ area = \pi \times raio^{2} $
#
# $ comprimento = 2 \times \pi * raio $
raio = 2
area = 3.14159 * raio ** 2
comprimento = 2 * 3.14159 * raio
print('Para o raio {}, a área = {} e o comprimento = {}'.format(raio, area, comprimento))
# ### Cálculo da distância euclidiana entre 2 pontos no plano cartesiano
#
# $ distância = \sqrt{(x_{1}-x_{2})^{2} + (y_{1}-y_{2})^{2}} $
# Calculando a distância entre os pontos (3,4) e (5,6)
x1 = 3
y1 = 4
x2 = 5
y2 = 6
dist = ((x1-x2)**2 + (y1-y2)**2)**(1/2)
print('A distância entre os pontos({},{}) e ({},{}) = {}.'.format(x1, y1, x2, y2, dist))
# ### Cálculo do delta e das raízes de uma equação do segundo grau
#
# $ \Delta = b^{2} - 4ac $
#
# $ x = \frac{-b \pm \sqrt{\Delta}}{2a} $
# Calculando o Delta e raízes x1 e x2 para a equação y = x^2 - 5x + 6
a = 1
b = -5
c = 6
delta = b**2 - 4*a*c
x1 = (-b+delta**(1/2))/(2*a)
x2 = (-b-delta**(1/2))/(2*a)
print('Delta = {}, x1 = {} e x2 = {}'.format(delta, x1, x2))
# ### Cálculo da área do setor circular (fatia de um círculo)
#
# $ S = \frac{\alpha \pi r^{2}}{360} $
# Para um círculo de raio 15, calcular a área de uma fatia com angulo = 45 graus (um pizza)
# Isso equivale a calcular a área de uma das 8 fatias de uma pizza com 30 cm de diâmetro
a = 45
r = 15
s = (a * 3.14159 * r ** 2)/360
print('Num círculo de raio {}cm, o setor circular de ângulo {} terá área = {} cm^2.'.format(r,a,s))
| Tipos de dados no Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring Raw Data
import sys
sys.path.append("..")
from src.prepare_data import load_dataset
# +
data_dir = "../data/external"
prefixes = ['train', 'test', 'valid']
target, source = [], []
for prefix in prefixes:
t, s = load_dataset(data_dir, prefix)
target.extend(t)
source.extend(s)
output = [s + t for s, t in zip(source, target)]
# -
source[0].strip().replace("[ WP ]", "[WP]")
"<|startoftext|> " + target[0].strip() + " <|endoftext|>"
source[0].strip().replace("[ WP ]", "[WP]") + "<|startoftext|> " + target[0].strip() + " <|endoftext|>"
# # Text Distribution
# +
import seaborn as sns
output_len = [len(o.split(" ")) for o in output]
sns.distplot([o for o in output_len if o < 3000], kde=False)
# -
import numpy as np
ind = np.where(np.array(output_len) < 400)[0]
print(len(ind))
short_outputs = [output[i] for i in ind]
# +
import random
random.choice(short_outputs)
# -
| notebooks/raw_data_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import the required libraries: Pandas, Numpy, Matplotlib and Seaborn
import pandas as pd
import numpy as np
import seaborn as sns # For mathematical calculations
import matplotlib.pyplot as plt # For plotting graphs
from datetime import datetime # To access datetime
from pandas import Series # To work on series
# %matplotlib inline
import warnings # To ignore the warnings
warnings.filterwarnings("ignore")
# set seed for reproducibility
np.random.seed(0)
# +
# fuzz is used to compare TWO strings
from fuzzywuzzy import fuzz
# process is used to compare a string to MULTIPLE other strings
from fuzzywuzzy import process
import chardet
# +
# Load and read the data
df=pd.read_csv("PakistanSuicideAttacks Ver 6 (10-October-2017).csv",encoding = "ISO-8859-1")
df.head(3)
# -
# Conduct preliminary text pre-processing
#
# Here I'm interested in cleaning up the "" Column to make sure there's no data entry inconsistencies in it.
# we will go through and check each row by hand and correct inconsistencies where we find them
#
# +
# Get all the unique values in the 'City' column
City= df["City"].unique()
City
# +
# Sort the City column rows alphabetically and then take a closer look
City.sort()
City
# -
#
# There are problems due to inconsistent data entry: 'Lahore' and 'Lahore ', for example, or 'Lakki Marwat' and 'Lakki marwat'.
# Fixing the text data entry inconsistencies
#
# Make everything into lowercase in the dataset.
#
# Remove the white spaces at the beginnign and the end of the cells.
#
# This will remove the inconsistencies in capitalizations and trailing white spaces in our City Column text data.
# +
# Convert the rows in the City Column to lower case from the dataset
# Set the values in the dataset df and column city to lower case
df["City"]= df['City'].str.lower()
df["City"].sample(4)
# +
# Remove the trailing white spaces in the City Column
df["City"] =df['City'].str.strip()
df["City"].sample(4)
# -
# Lets tackle more difficult Inconsistencies
# Use fuzzy matching to correct inconsistent data entry
#
# Still in the City Column, lets have a lok and see if there is more data cleaning that we need to do
# +
# Get all the unique values in the 'City' column
City = df['City'].unique()
City
# +
# Sort the City 2 Alphabetically and then take a closer look
City.sort()
City
# -
#
#
# There are some remaining inconsistencies: 'd. i khan' and 'd.i khan' should probably be the same.
# (I looked it up and 'd.g khan' is a seperate city, so I shouldn't combine those.)
#
# Using Fuzzywuzzy package to help identify which string are closest to each other.
#
#
#
# Fuzzy matching: The process of automatically finding text strings that are very similar to the target string. In general, a string is considered "closer" to another one the fewer characters you'd need to change if you were transforming one string into another. So "apple" and "snapple" are two changes away from each other (add "s" and "n") while "in" and "on" and one change away (rplace "i" with "o"). You won't always be able to rely on fuzzy matching 100%, but it will usually end up saving you at least a little time.
# Fuzzywuzzy returns a ratio given two strings. The closer the ratio is to 100, the smaller the edit distance between the two strings. Here, we're going to get the ten strings from our list of cities that have the closest distance to "d.i khan".
# +
# Get the top 10 closest matches to "d.i khan"
Matches = fuzzywuzzy.process.extract("d.i khan", Cities, limit=10, scorer=fuzzywuzzy.fuzz.token_sort_ratio)
Matches
# -
# We can see that two of the items in the cities are very close to "d.i khan": "d. i khan" and "d.i khan". We can also see the "d.g khan", which is a seperate city, has a ratio of 88. Since we don't want to replace "d.g khan" with "d.i khan", let's replace all rows in our City column that have a ratio of > 90 with "d. i khan".
#
# To do this, I'm going to write a function. (It's a good idea to write a general purpose function you can reuse if you think you might have to do a specific task more than once or twice. This keeps you from having to copy and paste code too often, which saves time and can help prevent mistakes.)
# function to replace rows in the provided column of the provided dataframe
# that match the provided string above the provided ratio with the provided string
def replace_matches_in_column(df, column, string_to_match, min_ratio = 90):
# get a list of unique strings
strings = df[column].unique()
# get the top 10 closest matches to our input string
matches = fuzzywuzzy.process.extract(string_to_match, strings,
limit=10, scorer=fuzzywuzzy.fuzz.token_sort_ratio)
# only get matches with a ratio > 90
close_matches = [match[0] for match in matches if match[1] >= min_ratio]
# get the rows of all the close matches in our dataframe
rows_with_matches = df[column].isin(close_matches)
# replace all rows with close matches with the input matches
df.loc[rows_with_matches, column] = string_to_match
# let us know the function's done
print("All done!")
# use the function we just wrote to replace close matches to "d.i khan" with "d.i khan"
replace_matches_in_column(df=df, column='City', string_to_match="d.i khan")
# And now let's can check the unique values in our City column again and make sure we've tidied up d.i khan correctly.
# +
# Get all the unique values in the 'City' column
City = df['City'].unique()
# sort them alphabetically and then take a closer look
City.sort()
City
# -
# Excellent! Now we only have "d.i khan" in our dataframe and we didn't have to change anything by hand.
| Data Cleaning Challenges/Data Cleaning Inconsistent Data Entry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="jKplW26sWhxM" outputId="f58ce85d-b7dd-4857-f3f4-26ddb952f067"
# !pip install stable-baselines3
# + id="g95_87S4T5__"
import gym
from gym import error, spaces, utils
from gym.utils import seeding
import numpy as np
from tqdm import tqdm
import matplotlib.pyplot as plt
# + id="aMvcF0WIUHaO"
class ForagingReplenishingPatches(gym.Env):
def __init__(self, block_type=1, manual_play=False):
self.reset_flag = False
self.action_space = spaces.Discrete(8)
self.observation_space = spaces.Discrete(8)
self.block_type = block_type
self.HARVEST_ACTION_ID = 8
if self.block_type == 1:
self.rewards = np.asarray([0, 70, 70, 0, 70, 0, 70, 0])
elif self.block_type == 2:
self.rewards = np.asarray([0, 0, 70, 70, 0, 70, 0, 70])
elif self.block_type == 3:
self.rewards = np.asarray([70, 0, 0, 70, 70, 0, 70, 0])
self.rewarding_sites = np.arange(8)[self.rewards > 0]
self.current_state = 0
self.time_elapsed = 1.307
self.farmer_reward = 0
self.init_env_variables()
if manual_play:
self.init_foraging_img()
self.manual_play()
def replenish_rewards(self):
if self.block_type == 1:
replenish_rates = np.asarray([0, 4, 4, 0, 4, 0, 4, 0])
elif self.block_type == 2:
replenish_rates = np.asarray([0, 0, 8, 2, 0, 5, 0, 8])
elif self.block_type == 3:
replenish_rates = np.asarray([2, 0, 0, 4, 8, 0, 16, 0])
replenish_rates[self.current_state] = 0
self.rewards += replenish_rates
self.rewards = np.clip(self.rewards, 0, 200)
def step(self, action):
self.time_elapsed += self.time_dist[str(self.current_state) + "to" + str(action)]
self.current_state = action
if self.time_elapsed >= 300:
self.reset_flag = True
return (self.current_state, 0 , self.reset_flag, {})
self.time_elapsed += 1
reward_old = self.farmer_reward
if self.current_state in self.rewarding_sites:
self.replenish_rewards()
self.farmer_reward += self.rewards[self.current_state] * 0.90
self.rewards[self.current_state] = (self.rewards[self.current_state] * 0.9)
if self.time_elapsed >= 300:
self.reset_flag = True
return (self.current_state, self.farmer_reward - reward_old, self.reset_flag, {})
def reset(self):
self.reset_flag = False
if self.block_type == 1:
self.rewards = np.asarray([0, 70, 70, 0, 70, 0, 70, 0])
elif self.block_type == 2:
self.rewards = np.asarray([0, 0, 70, 70, 0, 70, 0, 70])
elif self.block_type == 3:
self.rewards = np.asarray([70, 0, 0, 70, 70, 0, 70, 0])
self.rewarding_sites = np.arange(8)[self.rewards > 0]
self.current_state = 0
self.time_elapsed = 2
self.farmer_reward = 0
return self.current_state
def render(self, mode="human"):
print("Current State:", self.current_state, "Current Total Reward:", self.farmer_reward)
def close(self):
cv2.destroyAllWindows()
return None
def init_env_variables(self, first_point_angle=0):
a = 1 / (2 * np.sin(np.pi / 8)) # fix a (radius) for unit side octagon
self.octagon_points = np.asarray(
[
(
a * np.sin(first_point_angle + n * np.pi / 4),
a * np.cos(first_point_angle + n * np.pi / 4),
)
for n in range(8)
]
)
self.time_dist = {}
for i in range(8):
for j in range(8):
dist = np.linalg.norm(self.octagon_points[i] - self.octagon_points[j])
self.time_dist.update({str(i) + "to" + str(j): dist})
# + id="S3jayMHhWfPu"
from stable_baselines3.common.env_checker import check_env
env = ForagingReplenishingPatches(block_type=3)
check_env(env, warn=True)
# + id="mTqLTrc2XKZ5"
env.reset()
for i in range(300):
action = np.random.randint(8)
state, reward, done, _ = env.step(action)
print(action, state, reward, done)
if done:
break
# + colab={"base_uri": "https://localhost:8080/"} id="bNYWOQr-XX_L" outputId="1fae237e-e418-4f9c-f419-653ec6ec96f0"
from stable_baselines3 import PPO
env.reset()
model = PPO("MlpPolicy", env, verbose=1)
model.learn(total_timesteps=10**5)
# + colab={"base_uri": "https://localhost:8080/"} id="1KxVyexdXksl" outputId="56435eaf-e05e-4366-c3bd-c4b9deaa4ab5"
obs = env.reset()
while True:
action, _state = model.predict(obs, deterministic=False)
obs, reward, done, info = env.step(action)
env.render()
if done:
obs = env.reset()
break
# + colab={"base_uri": "https://localhost:8080/"} id="baRQxgqJYbcN" outputId="cfc191d8-0330-41ff-abed-d6986df4d5b0"
# save
# %cd /content/drive/MyDrive/Sem 5/CS698
# !mkdir saved_models
# %cd saved_models
model.save("PPOmlpPolicy")
# + colab={"base_uri": "https://localhost:8080/"} id="faCYLBmgZyo6" outputId="1fd376ae-54e7-4e92-e42c-50d3b235b13d"
print(model.policy)
# + colab={"base_uri": "https://localhost:8080/"} id="ni0eMPojaMUJ" outputId="d9064099-bc55-49ee-b478-173069be0b2b"
# %cd /content/drive/MyDrive/Sem 5/CS698
# !mkdir ppo_forager_tensorboard
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="pne4UBvZZ2zZ" outputId="fa28cb93-8701-4f5a-9153-747c4757cb56"
from stable_baselines3 import PPO
env.reset()
model = PPO("MlpPolicy", env, verbose=1, tensorboard_log="./ppo_forager_tensorboard/")
model.learn(total_timesteps=10**6)
# + colab={"base_uri": "https://localhost:8080/"} id="hZCiC8xrjLHd" outputId="fa52e24c-6c98-4ea8-9d3e-311bb439a7b5"
# save
# %cd /content/drive/MyDrive/Sem 5/CS698
# !mkdir saved_models
# %cd saved_models
model.save("PPOmlpPolicy1M")
# + colab={"base_uri": "https://localhost:8080/"} id="0QetMvOVadLA" outputId="d0de53e1-06bc-4dbe-d84a-2b62a98a2beb"
# !tensorboard --logdir ./ppo_forager_tensorboard/
| PROJECT/CS698R-Project-Foraging-in-Replenishing-Patches-7-main/notebooks/DRL/MDP_Foraging_PPO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Deployment sample (under construction)
# * Setup Workspace
# * Register a trained model
# * Convert the model to DLC for camera
# * Build IoT Image
# * Create Hub and Prepare for device and deploy
# * Deploy Model to Camera

# For prod
# !source activate py36 && pip install azureml-core azureml-contrib-iot azure-mgmt-containerregistry azure-cli
# !source activate py36 && az extension add --name azure-cli-iot-ext
import os
print(os.__file__)
# +
# Check core SDK version number
import azureml.core as azcore
print("SDK version:", azcore.VERSION)
# -
# ### Create a Workspace
# #### Change this cell from markdown to code and run this if you need to create a workspace
# #### Update the values for your workspace below
# ws=Workspace.create(subscription_id="replace-with-subscription-id",
# resource_group="your-resource-group",
# name="your-workspace-name",
# location="eastus2")
#
# ws.write_config()
# +
# Initialize existing Workspace
# reading config from ./aml_config/config.json
# update this file with your subscription ID, AML workspace and resource group
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
# -
# set the location for retrained model
# MUST include va-snpe-library_config.json file
retrained_model_location = '<path to the location of your trained model>'
# 
# +
# Register the model by uploading the model file and label file
from azureml.core.model import Model
model = Model.register(model_path = retrained_model_location,
model_name = "name_of_your_model",
tags = {"data": "Imagenet", "model": "object_classification", "type": "imagenet"},
description = "Description_of_your_model",
workspace = ws)
# -
# 
# +
from azureml.contrib.iot.model_converters import SnpeConverter
# submit a compile request to convert the model to SNPE compatible DLC file
compile_request = SnpeConverter.convert_tf_model(
ws,
source_model=model,
input_node="input",
input_dims="1,224,224,3",
outputs_nodes = ["final_result"],
allow_unconsumed_nodes = True)
print(compile_request._operation_id)
# -
# wait for the request to complete
compile_request.wait_for_completion(show_output=True)
# Get converted model
converted_model = compile_request.result
print(converted_model.name, converted_model.url, converted_model.version, converted_model.id, converted_model.created_time)
# set the image name. must be lower case alphanumeric characters, '-' or '.'
image_name = "name_your_image"+str(model.version)
print("image name:", image_name)
# +
from azureml.core.image import Image
from azureml.contrib.iot import IotContainerImage
# create the IoT container image for the camera - Yocto/ARM32.
# includes scripts to access the VAM engine to send camer frames for inferecning
print ('We will create an image for you now ...')
image_config = IotContainerImage.image_configuration(
architecture="arm32v7",
execution_script="main.py",
dependencies=["camera.py","iot.py","ipcprovider.py","utility.py", "frame_iterators.py"],
docker_file="Dockerfile",
tags = ["mobilenet"],
description = "MobileNet model retrained soda cans")
image = Image.create(name = image_name,
models = [converted_model],
image_config = image_config,
workspace = ws)
image.wait_for_creation(show_output = True)
# -
# 
# +
# Parameter list to configure the IoT Hub and the IoT Edge device
# Pick a name for what you want to call the module you deploy to the camera
module_name = "name of the AI module"++
# Resource group in Azure
resource_group_name= ws.resource_group
iot_rg=resource_group_name
# Azure region where your services will be provisioned
iot_location="location"
# Azure IoT Hub name
iot_hub_name="name of IoT Hub"
# Pick a name for your camera
iot_device_id="name of IoT Edge Device"
# Pick a name for the deployment configuration
iot_deployment_id="name for the deployment ID"
# -
# Getting your container details
container_reg = ws.get_details()["containerRegistry"]
reg_name=container_reg.split("/")[-1]
container_url = "\"" + image.image_location + "\","
subscription_id = ws.subscription_id
print('{}'.format(image.image_location), "<-- this is the URI configured in the IoT Hub for the device")
print('{}'.format(reg_name))
print('{}'.format(subscription_id))
from azure.mgmt.containerregistry import ContainerRegistryManagementClient
from azure.mgmt import containerregistry
client = ContainerRegistryManagementClient(ws._auth,subscription_id)
result= client.registries.list_credentials(resource_group_name, reg_name, custom_headers=None, raw=False)
username = result.username
password = result.passwords[0].value
# ### Deployment file
# This is the deployment.json file that you will use to deploy your model. Please see the other sample notebooks on using this file to deploy the new model you created.
file = open('./deployment-template.json')
contents = file.read()
contents = contents.replace('__MODULE_NAME', module_name)
contents = contents.replace('__REGISTRY_NAME', reg_name)
contents = contents.replace('__REGISTRY_USER_NAME', username)
contents = contents.replace('__REGISTRY_PASSWORD', password)
contents = contents.replace('__REGISTRY_IMAGE_LOCATION', image.image_location)
with open('./deployment.json', 'wt', encoding='utf-8') as output_file:
output_file.write(contents)
# !az login
# !az account set --subscription "<subscription>"
print("Pushing deployment to IoT Edge device")
print ("Set the deployement")
# !az iot edge set-modules --device-id $iot_device_id --hub-name $iot_hub_name --content deployment.json
# +
# THE END
# now check the IoT Edge device in Azure portal to verify
# the URI is updated and will be pushed to the device when it comes online.
| machine-learning-notebooks/04-Deploy-Trained-Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
# # Python for Finance (2nd ed.)
#
# **Mastering Data-Driven Finance**
#
# © Dr. <NAME> | The Python Quants GmbH
#
# <img src="http://hilpisch.com/images/py4fi_2nd_shadow.png" width="300px" align="left">
# # Appendix: BSM Option Class
from bsm_option_class import *
# + uuid="6fbac054-7632-4b1a-905a-1e7617ab1353"
o = bsm_call_option(100., 105., 1.0, 0.05, 0.2)
type(o)
# + uuid="29db884c-ae9d-44fc-ac14-ac7f42842d58"
value = o.value()
value
# + uuid="ad80b6ae-ba85-4e77-bc51-d539888e3ad1"
o.vega()
# + uuid="498adc88-13d0-4440-adbd-d4214e9529dc"
o.imp_vol(C0=value)
# + uuid="a3bd9b68-82cc-45e5-8c29-407a98ad1aec"
import numpy as np
maturities = np.linspace(0.05, 2.0, 20)
strikes = np.linspace(80, 120, 20)
T, K = np.meshgrid(strikes, maturities)
C = np.zeros_like(K)
V = np.zeros_like(C)
for t in enumerate(maturities):
for k in enumerate(strikes):
o.T = t[1]
o.K = k[1]
C[t[0], k[0]] = o.value()
V[t[0], k[0]] = o.vega()
# + uuid="4beed5a6-ed44-4fd3-aaf0-6bc4be2b1c83"
from pylab import cm, mpl, plt
from mpl_toolkits.mplot3d import Axes3D
mpl.rcParams['font.family'] = 'serif'
# %config InlineBackend.figure_format = 'svg'
# + uuid="aae2d996-fa8e-469d-8ffa-f0a5684160a7"
fig, ax = plt.subplots(subplot_kw={"projection": "3d"}, figsize=(12, 7))
surf = ax.plot_surface(T, K, C, rstride=1, cstride=1,
cmap=cm.coolwarm, linewidth=0.5, antialiased=True)
ax.set_xlabel('strike')
ax.set_ylabel('maturity')
ax.set_zlabel('European call option value')
fig.colorbar(surf, shrink=0.5, aspect=5);
# plt.savefig('../../images/b_bsm/bsm_01.png');
# + uuid="a0fc3738-180b-4be7-9c31-460b3d015194"
fig, ax = plt.subplots(subplot_kw={"projection": "3d"}, figsize=(12, 7))
surf = ax.plot_surface(T, K, V, rstride=1, cstride=1,
cmap=cm.coolwarm, linewidth=0.5, antialiased=True)
ax.set_xlabel('strike')
ax.set_ylabel('maturity')
ax.set_zlabel('Vega of European call option')
fig.colorbar(surf, shrink=0.5, aspect=5);
# plt.savefig('../../images/b_bsm/bsm_02.png');
# -
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
#
# <a href="http://tpq.io" target="_blank">http://tpq.io</a> | <a href="http://twitter.com/dyjh" target="_blank">@dyjh</a> | <a href="mailto:<EMAIL>"><EMAIL></a>
| code/b_bsm/b_bsm_option_class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# - Abstract - Train a classification model on a dataset of my choice
# - Introduction - I went to UCI ML Repo and discovered the mushroom dataset. I was told in my culinary education that gill color is best indictator that a mushroom is edible.
# - Very specific and clear research questions - How accurate is gill color to poisonous?
# - Brief EDA(Exploratory Data Analysis)
# - Data Cleaning
# - Feature Engineering
# - Modeling
# - Results and discussion
# - Conclusion and summary
# - Limitations and later work.
# - References and contributions
# https://archive.ics.uci.edu/ml/machine-learning-databases/mushroom/agaricus-lepiota.data
# -
# !pip install python-utils
import numpy as np
import pandas as pd
import warnings
warnings.simplefilter("ignore")
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
col_Names=['class','cap-shape','cap-surface','cap-color','bruises?','odor','gill-attachment','gill-spacing','gill-size','gill-color','stalk-shape' ,'stalk-root','stalk-surface-above-ring','stalk-surface-below-ring','stalk-color-above-ring','stalk-color-below-ring','veil-type','veil-color','ring-number','ring-type','spore-print-color','population','habitat']
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/mushroom/agaricus-lepiota.data"
df = pd.read_table(url , sep=",", names=col_Names)
df.shape
df.describe()
#Let's pick a random sample of the dataset of mushrooms
df_sample = df.loc[np.random.choice(df.index, 8000, False)]
#Get all unique cap-colors
df_sample['ring-type'].unique()
# +
#Get 'gill-color' Series
ringType= df_sample['ring-type']
#Get the total number of mushrooms for each unique gill color.
ringType.value_counts()
# -
X=df.drop('class', axis=1) #Predictors
y=df['class'] #Response
X.head()
from sklearn.preprocessing import LabelEncoder
Encoder_X = LabelEncoder()
for col in X.columns:
X[col] = Encoder_X.fit_transform(X[col])
Encoder_y=LabelEncoder()
y = Encoder_y.fit_transform(y)
X.head()
# Poisonous = 1
# Edible = 0
X.hist(bins=20, figsize=(20,15))
save_fig("attribute_histogram_plots")
plt.show()
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X,y)
print(lr.intercept_, lr.coef_)
X=pd.get_dummies(X,columns=X.columns,drop_first=True)
X.head()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1, stratify =y)
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
sc.fit(X_train)
X_train = sc.transform(X_train)
X_test = sc.transform(X_test)
X_train.shape, X_test.shape
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(penalty = 'none', random_state=1)
logreg.fit(X_train, y_train)
score = logreg.score(X,y)
score
predictions = logreg.predict(X)
# +
from sklearn import metrics
cm = metrics.confusion_matrix(y, predictions)
# -
plt.figure(figsize=(9,9))
sns.heatmap(cm, annot=True, fmt=".3f", linewidths=.5, square = True, cmap = 'Pastel1');
plt.ylabel('Actual label');
plt.xlabel('Predicted label');
all_sample_title = 'Accuracy Score: {0}'.format(score)
plt.title(all_sample_title, size = 15);
plt.savefig('toy_Digits_ConfusionSeabornCodementor.png')
plt.show()
# +
sns.relplot(x="odor", y="gill-color",
col="class", aspect=1,
kind="scatter", data=df)
plt.tight_layout()
# +
sns.relplot(x="odor", y="cap-color",
col="class", aspect=.7,
kind="scatter", data=df)
plt.tight_layout()
# +
sns.relplot(x="gill-color", y="cap-color",
col="class", aspect=.7,
kind="scatter", data=df)
plt.tight_layout()
# -
| Technical Notebook/Mid_Term_Mushroom_Hist_LogReg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# -*- coding: utf-8 -*-
###
# Demonstration of MR reconstruction with CCP PET-MR Software
#
# This demonstration shows how to reconstruct MR images on a coil-by-coil basis
# and how to combine the image information from the different receiver coils
#
# This demo is a 'script', i.e. intended to be run step by step in a
# Python IDE such as spyder. It is organised in 'cells'. spyder displays these
# cells nicely and allows you to run each cell on its own.
#
# First version: 27th of May 2017
# Updated 10th of April 2019
# Author: <NAME>, <NAME>
#
## CCP PETMR Synergistic Image Reconstruction Framework (SIRF).
## Copyright 2015 - 2017 Rutherford Appleton Laboratory STFC.
## Copyright 2015 - 2017 University College London.
## Copyright 2015 - 2017 Physikalisch-Technische Bundesanstalt.
##
## This is software developed for the Collaborative Computational
## Project in Positron Emission Tomography and Magnetic Resonance imaging
## (http://www.ccppetmr.ac.uk/).
##
## Licensed under the Apache License, Version 2.0 (the "License");
## you may not use this file except in compliance with the License.
## You may obtain a copy of the License at
## http://www.apache.org/licenses/LICENSE-2.0
## Unless required by applicable law or agreed to in writing, software
## distributed under the License is distributed on an "AS IS" BASIS,
## WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
## See the License for the specific language governing permissions and
## limitations under the License.
# %matplotlib notebook
# -
# ## Coil Combination Methods
# ### Goals of this notebook:
# - Explore ways to combine acquisition data from multiple receiver coils.
# +
__version__ = '0.1.0'
# import engine module
import sirf.Gadgetron as pMR
from sirf.Utilities import examples_data_path
# import further modules
import os, numpy
import matplotlib.pyplot as plt
#%% GO TO MR FOLDER
os.chdir( examples_data_path('MR'))
# -
# ### Coil Combination
# #### Goals of this notebook:
# - Explore ways to combine acquisition data from multiple receiver coils.
# ### Multi Receiver Channel Imaging
# In principle one does not need multiple coils placed on a patient or phantom in an MR exam. Every scanner has the so called body coil which is able to receive the MRI signal and one can reconstruct an image from that.
#
# However, nobody uses the body coil images any more but instead employs what is called __receiver coils__, aka "coils", or somethimes refered to as __phased arrays__. When talking about data recorded by different coils these are also referred to as __channels__ (in the literature it always says something of the kind: "_... data were acquired using a 32-channel cardiac coil..._").
#
# This has several advantages, one being that the receiver can be placed very close to the signal source. Others we will discuss in the following!
#
# __Important:__ each of this coils also adds a global complex phase onto the recorded signal (hence the name __phased__ arrays!)
#
#
# #### Naming Convention
# The following expressions are usually used synonymously:
# - coil sensitivity profile
# - coil sensitivity maps (CSM)
# - coil maps
# +
#%% LOAD AND PREPROCESS RAW MR DATA
filename = 'ptb_resolutionphantom_fully_ismrmrd.h5'
acq_data = pMR.AcquisitionData(filename)
preprocessed_data = pMR.preprocess_acquisition_data(acq_data)
preprocessed_data.sort()
# +
#%% RETRIEVE K-SPACE DATA
k_array = preprocessed_data.as_array()
print('Size of k-space %dx%dx%d' % k_array.shape)
# +
#%% PLOT K-SPACE DATA
k_array = k_array / numpy.max(abs(k_array[:]))
num_channels = k_array.shape[1]
fig = plt.figure()
plt.set_cmap('gray')
for c in range( num_channels ):
ax = fig.add_subplot(2,num_channels/2,c+1)
ax.imshow(abs(k_array[:,c,:]), vmin=0, vmax=0.05)
ax.set_title('Coil '+str(c+1))
ax.axis('off')
plt.tight_layout()
# +
#%% APPLY INVERSE FFT TO EACH COIL AND VIEW IMAGES
image_array = numpy.zeros(k_array.shape, numpy.complex128)
for c in range(k_array.shape[1]):
image_array[:,c,:] = numpy.fft.fftshift(numpy.fft.ifft2(numpy.fft.ifftshift(k_array[:,c,:])))
image_array = image_array/image_array.max()
fig = plt.figure()
plt.set_cmap('gray')
for c in range(image_array.shape[1]):
ax = fig.add_subplot(2,num_channels/2,c+1)
ax.imshow(abs(image_array[:,c,:]), vmin=0, vmax=0.4)
ax.set_title('Coil '+str(c+1))
ax.axis('off')
plt.tight_layout()
# -
# ### Question:
# - What differences appear in the individual channel reconstructions compared to the combined image we saw in the last notebook?
#
# ### Sum of Square (SOS) Coil Combination
#
# As you can see the individual receiver channels have a spatially varying intensity due to the coil sensitivity profiles. This information needs to be combined.
#
# When you have a set of independently reconstructed images for each channel $f_c$ where $c \in \{1, \dots N_c \}$ labels the individual coil channels.
#
# One way to combine the signal from all coil channels is to use a sum-of-squares approach:
#
# $$
# f_{sos} = \sqrt{ \sum_c \bigl{|} \, f_c \bigr{|}^2 }
# $$
# +
#%% COMBINE COIL IMAGES USING SOS
#image_array_sos = numpy.sqrt(abs(numpy.sum(image_array,1)))
image_array_sos = numpy.sqrt(numpy.sum(numpy.square(numpy.abs(image_array)),1))
image_array_sos = image_array_sos/image_array_sos.max()
fig = plt.figure()
plt.set_cmap('gray')
plt.imshow(image_array_sos, vmin=0, vmax=0.7)
plt.title('Combined image using sum-of-squares')
# -
# ### Question:
#
# - Apart from fancy parallel imaging techniques, why is it even useful to have more than one receiver channel?
# - What could be a possible disadvantage of this coil combination approach?
# - Why is SOS preferable to simply summing them without squaring: $\;f_{combined} = \sum_c f_c$
# ### Why Coil Sensitivity Maps?
# There are several reasons why one would need to compute the CSMs, e.g.:
# 1. I want to do parallel imaging (i.e. use spatial encoding provided by CSMs)
# 2. I want to improve my SNR.
#
#
# ##### Weighted Sum (WS) Coil Combination
# Instead of squaring each channel before adding them up there is another method of combining coils, by weighting each image with it's corresponding CSM.
# If the spatial distribution of the coil sensitivity $C_c$ for each coil $c$ is known then combining images as:
#
# $$
# f_{ws} = \frac{1}{{\sum_{c'}{\bigl{|} C_{c'} \bigr{|}^2}}}{\sum_c C_c^* \cdot f_c}
# $$
#
# yields an optimal signal-to-noise ratio (SNR).
# Note also, that this way of combining channels does not destroy the phase information.
#
# __However,__ for each coil one needs to either
# - estimate the coil map $C_c$ from the data itself.
# - measure them separately (clinically not feasible).
#
#
# ### Computing Coil Sensitivities
# The blunt approach is to compute the CSMs using the __SRSS__ (Square Root of the Sum of Squares) approach:
#
# $$
# C^{SRSS}_c = \frac{f_c}{f_{sos}}
# $$
#
# and to apply some smoothing to the data.
#
# As you can imagine there will be no big SNR difference between $f_{ws}$ and $f_{sos}$ using these coil maps. __We didn't put in any additional effort!__ This works well if the SOS image is homogenous.
#
# __This seems a bit pointless!__ True, combining your images this way will __not give you a gigantic SNR gain__, but you __still get a CSM which you can use for parallel imaging__. And you can generate an coil-combined image __without losing phase information__ (because we smooth the CSMs!)!
#
# This all works well when the SOS image is good to begin with. Otherwise, there are more "sophisticated" ways to estimate the coilmaps, e.g. methods named _Walsh_ or _Inati_ which lie beyond of the scope of this workshop.
# __SIRF already provides this functionality__. For the task of estimating coil sensitivities from the acquisition data, in `pMR` there is a class called `CoilSensitivityData`.
#
# +
#%% CALCULATE COIL SENSITIVITIES
csm = pMR.CoilSensitivityData()
#help(csm)
csm.smoothness = 50
csm.calculate(preprocessed_data)
# +
# Cell plotting the Coilmaps
csm_array = numpy.squeeze(csm.as_array(0))
# csm_array has orientation [coil, im_x, im_y]
csm_array = csm_array.transpose([1,0,2])
fig = plt.figure()
plt.set_cmap('jet')
for c in range(num_channels):
ax = fig.add_subplot(2,num_channels/2,c+1)
ax.imshow(abs(csm_array[:,c,:]))
ax.set_title('Coil '+str(c+1))
ax.axis('off')
# -
# ### Question
# Please answer the following questions:
# - Why is there noise in some regions of the coilmaps and outside the object?
# - Is this noise in the coilmap going to have a strong negative impact on the image quality in this region of the combined image?
# - In which organ in the human anatomy would you expect a coilmap to look similarly noisy?
#
#
# +
#%% COMBINE COIL IMAGES USING WEIGHTED SUM
image_array_ws = numpy.sum(numpy.multiply(image_array, numpy.conj(csm_array)),1)
image_array_ws = abs(numpy.divide(image_array_ws, numpy.sum(numpy.multiply(csm_array, numpy.conj(csm_array)),1)))
image_array_ws = image_array_ws/image_array_ws.max()
diff_img_arr = abs(image_array_sos-image_array_ws)
diff_img_arr = diff_img_arr/diff_img_arr.max()
fig = plt.figure(figsize=[12, 4])
plt.set_cmap('gray')
ax = fig.add_subplot(1,3,1)
ax.imshow(image_array_sos, vmin=0, vmax=0.7)
ax.set_title('Sum-of-squares (SOS)')
ax.axis('off')
ax = fig.add_subplot(1,3,2)
ax.imshow(image_array_ws, vmin=0, vmax=0.7)
ax.set_title('Weighted sum (WS)')
ax.axis('off')
ax = fig.add_subplot(1,3,3)
ax.imshow(diff_img_arr, vmin=-0, vmax=0.1)
ax.set_title('SOS - WS')
ax.axis('off')
# -
# ### Image Quality Assessment
# In low-signal regions you can see from the difference image, a weighted coil combination will give you an improved SNR.
# This dataset was acquired with a head-coil so which is very well-matched, and no flow artifacts appear so the difference is not huge.
# ### Question: What's "The mysterious next step?":
# 1. Reconstructing individual channels, add them up and get one image out.
# 2. Reconstructing individual channels, weight them by their coil maps, add them up and get one image out.
# 3. Do the mysterious next step, and get one image out.
#
# ### Recap Coil Combination
#
# In this exercise we learned:
# - how to combine multichannel image reconstructions
# - how to compute a simple coil sensitivity from our data.
# - how to employ SIRF to execute this computation for us.
#
#
#
plt.close('all')
| notebooks/MR/c_coil_combination.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This module focuses on Basic Visualization Tools
# +
UserPath= "/home/cerbero/"
ImportPath = UserPath+"Documents/IBM DV0101EN/01/"
ExportPath = UserPath+"Documents/IBM DV0101EN/02/"
import pandas as pd
# -
df_canada = pd.read_excel(ImportPath+"Canada Immigration Data.xlsx")
df_canada.describe()
df_canada.head()
df_canada.shape
df_01 = df_canada.sort_values(["Total"], ascending = False, axis = 0)
df_01 = df_01.set_index("Country")
df_01.head()
df_02 = df_01.head()
df_02 = df_02.drop(['Continent','Region','DevName','Total'], axis=1)
df_02 = df_02.transpose()
df_02
import matplotlib as mpl
import matplotlib.pyplot as plt
# +
# %matplotlib inline
df_02.plot(kind="area", alpha=0.8, stacked=True, figsize=(14,8))
#stacked=False makes plot translucent, doesn't work here
#The unstacked plot has a default transparency (alpha value) at 0.5.
#We can modify this value by passing in the alpha parameter
plt.title("Immigration to Canada - Top 5 Trend")
plt.ylabel("People")
plt.xlabel("Years")
# plt.show() #works without this line. Not sure what it is for...
# the show() above messes up with the savefig function
plt.savefig(ExportPath+"Immigration to Canada - Top 5 Trend.png")
# -
years = list(map(int, range(1980, 2014)))
print(years)
# +
# get the 4 countries with the least contribution
# Lab wanted to use 5 countries, but it does not look good - Vietnam is much larger
# Will use the Artist layer to show an alternative way to plot
df_least4 = df_canada.tail(4)
df_least4 = df_least4[years].transpose()
df_least4.head()
ax = df_least4.plot(kind='area', alpha=0.55, stacked=True, figsize=(14, 8)) #object-oriented
ax.set_title('Immigration Trend of 4 Countries with Least Contribution to Immigration') #notice the setters
ax.set_ylabel('Number of Immigrants')
ax.set_xlabel('Years')
plt.savefig(ExportPath+"Immigration to Canada - Bottom 4 Trend.png")
# -
# You can use an Axes instance of your current plot and store it in a variable (eg. ax). You can add more elements by calling methods with a little change in syntax (by adding "set_" to the previous methods). For example, use ax.set_title() instead of plt.title() to add title, or ax.set_xlabel() instead of plt.xlabel() to add label to the x-axis.
#
# This option sometimes is more transparent and flexible to use for advanced plots (in particular when having multiple plots, as you will see later).
df_least4 #Western Sahara is there, but barely
#might as well drop the index. Not very useful.
df_canada = df_canada.set_index("Country")
df_canada
# +
df_canada[2013].plot(kind="hist", figsize=(20,10))
plt.title("Histogram of Immigration to Canada from 195 countries in 2013")
plt.ylabel("Number of Countries")
plt.xlabel("Partitions of Number of Immigrants")
plt.savefig(ExportPath+"Histogram of 195 countries.png")
# -
# The histogram above is not very effective. Letś fix it.
import numpy as np
# +
count, bin_edges = np.histogram(df_canada[2013]) #this numpy function gives nicer bin-edges; count is unused
# default is 10 bins
df_canada[2013].plot(kind = "hist", xticks = bin_edges, figsize=(14,8)) #xticks now is bin_edges
# plt.grid(zorder = 5)
# plt.bar(bin_edges, 195, zorder = 0) #z-order did not work well
plt.grid()
plt.title("Histogram of Immigration to Canada from 195 countries in 2013")
plt.ylabel("Number of Countries")
plt.xlabel("Partitions of Number of Immigrants")
plt.savefig(ExportPath+"Improved Histogram of 195 countries.png")
# -
print(count, bin_edges) #two arrays are returned, naturally
df_canada.loc[['Denmark', 'Norway', 'Sweden'], years]
df_canada.loc[['Denmark', 'Norway', 'Sweden'], years].plot.hist()
#need to transpose
df_t = df_canada.loc[['Denmark', 'Norway', 'Sweden'], years].transpose()
df_t.head()
# +
df_t.plot(kind='hist', figsize=(10, 6))
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
# +
#still does not look good
# let's get the x-tick values
count, bin_edges = np.histogram(df_t, 15)
# un-stacked histogram
df_t.plot(kind ='hist',
figsize=(10, 6),
bins=15,
alpha=0.6,
xticks=bin_edges,
color=['coral', 'darkslateblue', 'mediumseagreen']
)
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
# +
# Tip: For a full listing of colors available in Matplotlib, run the following code in your python shell:
import matplotlib
for name, hex in matplotlib.colors.cnames.items():
print(name, hex)
# -
# Finally, the bar chart
df_iceland = df_canada.loc["Iceland", years]
df_iceland.head(), df_iceland.tail()
# +
df_iceland.plot(kind="bar", figsize =(14,8))
plt.title("Immigration from Iceland to Canada over the Years")
plt.xlabel("Years")
plt.ylabel("Number of Immigrants")
plt.savefig(ExportPath+"Immigration from Iceland to Canada over the Years - Bar Chart.png")
# +
df_iceland.plot(kind='bar', figsize=(10, 6), rot=90)
plt.xlabel('Year')
plt.ylabel('Number of Immigrants')
plt.title('Icelandic Immigrants to Canada from 1980 to 2013')
# Annotate arrow
plt.annotate('', # s: str. will leave it blank for no text
xy=(32, 70), # place head of the arrow at point (year 2012 , pop 70)
xytext=(28, 20), # place base of the arrow at point (year 2008 , pop 20)
xycoords='data', # will use the coordinate system of the object being annotated
arrowprops=dict(arrowstyle='->', connectionstyle='arc3', color='blue', lw=2)
)
# Annotate Text
plt.annotate('2008 - 2011 Financial Crisis', # text to display
xy=(28, 30), # start the text at at point (year 2008 , pop 30)
rotation=72.5, # based on trial and error to match the arrow
va='bottom', # want the text to be vertically 'bottom' aligned
ha='left', # want the text to be horizontally 'left' algned.
)
plt.savefig(ExportPath+"Immigration from Iceland to Canada over the Years - Bar Chart with Arrow.png")
# +
#Using horizontal bars
# sort dataframe on 'Total' column (descending)
df_top15 = df_canada.sort_values(by='Total', ascending=True)
# get top 15 countries
df_top15 = df_top15['Total'].tail(15)
df_top15
# +
# generate plot
df_top15.plot(kind='barh', figsize=(12, 12), color='steelblue') #kind="barh" is key here
plt.xlabel('Number of Immigrants')
plt.title('Top 15 Countries in Immigration to Canada between 1980 - 2013')
# annotate value labels to each country
for index, value in enumerate(df_top15):
label = format(int(value), ',') # format int with commas
# place text at the end of bar (subtracting 47000 from x, and 0.1 from y to make it fit within the bar)
plt.annotate(label, xy=(value - 47000, index - 0.10), color='white')
plt.savefig(ExportPath+"Top 15 Countries in Immigration to Canada between 1980 - 2013.png")
# -
df_canada.to_excel(ExportPath+"Canada Immigration Data.xlsx")
df_iceland.to_excel(ExportPath+"Immigration from Iceland to Canada.xlsx")
| 02 Second Module.ipynb |