code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # HATPRO
# The following example presents the nadir passive microwave radiometer HATPRO. The Humidity And Temperature PROfiler (HATPRO) replaced the MiRAC-P radiometer during the MOSAiC-ACA campaign.
#
# If you have questions or if you would like to use the data for a publication, please don't hesitate to get in contact with the dataset authors as stated in the dataset attributes `contact` or `author`.
#
# ## Data access
# * To analyse the data they first have to be loaded by importing the (AC)³airborne meta data catalogue. To do so the ac3airborne package has to be installed. More information on how to do that and about the catalog can be found [here](https://github.com/igmk/ac3airborne-intake#ac3airborne-intake-catalogue).
# ## Get data
import ac3airborne
cat = ac3airborne.get_intake_catalog()
list(cat.P5.HATPRO)
# ```{note}
# Have a look at the attributes of the xarray dataset `ds_hatpro` for all relevant information on the dataset, such as author, contact, or citation information.
# ```
ds_hatpro = cat['P5']['HATPRO']['MOSAiC-ACA_P5_RF11'].to_dask()
ds_hatpro
# The dataset includes brightness temperatures (`tb`) observed by HATPRO at the 22 GHz water vapor absorption line (22.24, 23.04, 23.84, 25.44, 26.24, 27.84 GHz), at the 31.3 GHz window frequency and the 56 GHz oxygen absorption line (51.26, 52.28, 53.86, 54.94, 56.66, 57.3, 58.0 GHz).
# ## Load Polar 5 flight phase information
# Polar 5 flights are divided into segments to easily access start and end times of flight patterns. For more information have a look at the respective [github](https://github.com/igmk/flight-phase-separation) repository.
#
# At first we want to load the flight segments of (AC)³airborne
meta = ac3airborne.get_flight_segments()
# The following command lists all flight segments into the dictionary `segments`
segments = {s.get("segment_id"): {**s, "flight_id": flight["flight_id"]}
for platform in meta.values()
for flight in platform.values()
for s in flight["segments"]
}
# In this example we want to look at a high-level segment during MOSAiC-ACA RF11
seg = segments["MOSAiC-ACA_P5_RF11_hl05"]
# Using the start and end times of the segment `MOSAiC-ACA_P5_RF11_hl05` stored in `seg`, we slice the HATPRO data to the selected flight section.
ds_hatpro_sel = ds_hatpro.sel(time=slice(seg["start"], seg["end"]))
# In polar regions, the surface type is helpful for the interpretation of airborne passive microwave observations, especially near the marginal sea ice zone, as generally a higher emissivity is expected over sea ice compared to open ocean. Therefore, we also load AMSR2 sea ice concentration data along the Polar 5 flight track, which is operationally derived by the University of Bremen.
ds_sea_ice = cat['P5']['AMSR2_SIC']['MOSAiC-ACA_P5_RF11'].to_dask().sel(
time=slice(seg["start"], seg["end"]))
# ## Plots
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from matplotlib import cm
import numpy as np
plt.style.use("../mplstyle/book")
# +
fig, (ax1, ax2, ax3) = plt.subplots(3, 1, sharex=True, gridspec_kw=dict(height_ratios=(1, 1, 0.1)))
kwargs = dict(s=10, linewidths=0)
colors = cm.get_cmap('viridis', 7).colors
for i in range(0, 7):
ax1.scatter(ds_hatpro_sel.time, ds_hatpro_sel.tb.sel(channel=i), label='%g GHz'%ds_hatpro_sel.frequency.sel(channel=i).item(), color=colors[i], **kwargs)
for i in range(7, 14):
ax2.scatter(ds_hatpro_sel.time, ds_hatpro_sel.tb.sel(channel=i), label='%g GHz'%ds_hatpro_sel.frequency.sel(channel=i).item(), color=colors[i-7], **kwargs)
ax1.legend(frameon=False, bbox_to_anchor=(1.05, 0.5), loc='center left')
ax2.legend(frameon=False, bbox_to_anchor=(1.05, 0.5), loc='center left')
ax1.set_ylabel('$T_b$ [K]')
ax2.set_ylabel('$T_b$ [K]')
# plot AMSR2 sea ice concentration
im = ax3.pcolormesh(np.array([ds_sea_ice.time, ds_sea_ice.time]),
np.array([0, 1]),
np.array([ds_sea_ice.sic]), cmap='Blues_r', vmin=0, vmax=100,
shading='auto')
cax = fig.add_axes([0.87, 0.085, 0.1, ax3.get_position().height])
fig.colorbar(im, cax=cax, orientation='horizontal', label='Sea ice [%]')
ax3.tick_params(axis='y', labelleft=False, left=False)
ax3.xaxis.set_major_formatter(mdates.DateFormatter('%H:%M'))
ax3.set_xlabel('Time (hh:mm) [UTC]')
plt.show()
# -
| how_to_ac3airborne/datasets/hatpro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Libraries
# +
#export
import torch
import torch.nn as nn
from fastai.vision.all import *
from fastai.data.all import *
from fastai.distributed import *
import pandas as pd
from pathlib import Path
import time
from datetime import date
from vidaug import augmentors as va
import cv2
import torchvision.transforms as T
from modules_th.video_block import *
from modules_th.inflator import *
from modules_th.triplet_loss import *
from modules_th.supcon_module import *
from modules_th.cus_cbs import *
#Pretrained models
from modules_th.pretrained_r2p1d50 import *
from resnetmodels.mean import get_mean_std
# -
# %config Completer.use_jedi = False
#export
def read_data():
items_path = '/mnt/data/adrianlopez/Datasets/kinetics700/kinetics_train.csv'
return pd.read_csv(items_path, index_col=0)
df = read_data()
df.head(3)
# +
# export
def sometimes(x):
return va.Sometimes(0.3,x)
def wrapVideo(x):
return Video(x)
# -
#export
def get_dsets(df, l=4, size=512,skip=2,n_views=1):
vid_pip = [ColReader('vid_files'),
createVideoForm(l=l,skip=skip, form='img'),
Resize(size, method=ResizeMethod.Pad),
# sometimes(va.HorizontalFlip()),
# va.GaussianBlur(1.),
sometimes(va.InvertColor()),
# va.RandomRotate(10),
wrapVideo]
lbl_pip = [ColReader('label'), Categorize()]
pip = [*([vid_pip]*n_views), lbl_pip]
# Splits
splits = ColSplitter('val')(df)
# Datasets and dataloaders
dsets = Datasets(df, pip, splits=splits)
return dsets, splits
df.head()
dsets,splits = get_dsets(df)
# %time dsets.train[0][0][1].show(), len(dsets.train[0][0])
for i, (vid, lbl) in enumerate(dsets.train[1000:1300]): print(i, end='\r')
# export
def get_dls(dsets,splits,df, n_el= 2, n_lbl = 2, shuffle_fn=UniformizedShuffle, normalize='kinetics'):
mean, std = get_mean_std(1,normalize)
dls = dsets.dataloaders(bs=n_el*n_lbl,
shuffle_train=True,
after_item=ToTensor(),
after_batch=[IntToFloatTensor(), Normalize.from_stats(*imagenet_stats)])
dls.valid.shuffle = True
if shuffle_fn is not None:
dls.train.shuffle_fn = UniformizedShuffle(df.label.iloc[splits[0]], n_el = n_el, n_lbl= n_lbl)
dls.valid.shuffle_fn = UniformizedShuffle(df.label.iloc[splits[1]], n_el = n_el, n_lbl= n_lbl)
return dls
True
dls = get_dls(dsets,splits,df, normalize='kinetics')
#export
def setup_log(learn,name, append=True):
# set up logs file
# now = datetime.now()
# time = now.strftimes("%d_%m")
logs_file = '/mnt/data/eugeniomarinelli/UCF_experiments/training_results/logs_kinetics_'+name+'.csv'
Logs_csv = CSVLogger(fname= logs_file, append=append)
learn.add_cb(Logs_csv)
#export
def get_learner(df,
pretrained_model='r2p1d50_K',
loss_name='CEL',
l=40, size=224, n_lbl =2, n_el=2, skip=20, embs_size=256,n_views=2,
normalize = 'kinetics'):
dsets,splits = get_dsets(df, l, size, skip, n_views)
dls = get_dls(dsets,splits,df, normalize=normalize)
if pretrained_model in inserted_models:
model = inserted_models[pretrained_model]
else: raise 'model not present in pretrained models'
body = create_body(model, cut=-2)
if loss_name == 'SCL+CEL':
Loss = SumLoss(SupConLoss,p='cos', alpha=1, n_views=n_views)
head = inflate(create_head(256, len(dls.vocab), lin_ftrs=[]))
model = MixedLossModel(body,head)
metrics = [supcon_accuracy, silh_score]
elif loss_name == 'SCL':
Loss= SupConLoss()
head = inflate(create_head(4096, embs_size, lin_ftrs=[]))
model = nn.Sequential(body,head)
metrics = [silh_score]
elif loss_name == 'CEL':
Loss = CEL()
head = inflate(create_head(4096, len(dls.vocab), lin_ftrs=[256]))
model = MixedLossModel(body,head)
metrics = [supcon_accuracy,silh_score]
elif loss_name == 'CEL_after_SCL':
Loss = CrossEntropyLossFlat()
saved_model = torch.load('/mnt/data/eugeniomarinelli/UCF_experiments/trained_models_cnn/models/r2p1d50_ucf101_SCL_tuned_15fr.pth')
model = nn.Sequential(saved_model,nn.Sequential(nn.ReLU(inplace=True),LinBnDrop(256, 101, p=0.5)))
metrics = [accuracy]
else:
raise 'Loss not implemented'
learn = Learner(dls,
model,
splitter=splitter ,
loss_func=Loss,
metrics=metrics)
if loss_name == 'SCL+CEL':
learn.add_cbs([ContrastiveCallback(n_views)])#,LossesRecorderCallback()])
elif loss_name == 'SCL':
learn.add_cb(ContrastiveCallback(n_views))
elif loss_name in ['CEL', 'CEL_after_SC']:
learn.add_cb(MultiViewsCallback(n_views))
time = date.today().strftime("_%d-%m")
setup_log(learn, str(pretrained_model)+'_'+loss_name+'_tuning_10_'+time, append=True)
return learn
learn = get_learner(df, 'r2p1d50_K', loss_name='CEL_after_SCL', l=2, size=64, n_lbl =2, n_el=2, skip=20, embs_size=256, n_views=2)
#learn.add_cb(CSVLogger(fname='data/eugeniomarinelli/UCF_experiments/trained_models_cnn/learners/LOGGER_PROVA' , append=False) )
from datetime import date
date.today().strftime("_%d-%m")
True
learn.fit_one_cycle(1)
inp = torch.randn(8,3,30,100,100).cuda()
model = learn.model.cuda()
model(inp).shape
# +
#export
def save_learner(learn, name):
prefix = '/mnt/data/eugeniomarinelli/'
try:
learn.export(prefix+'UCF_experiments/trained_models_cnn/learners/learner_kinetics_'+name)
except: print("learner export didn't work")
try:
torch.save(learn.model,prefix+'UCF_experiments/trained_models_cnn/models/model_kinetics_'+name+'.pth')
except: pass
torch.save(learn.model.state_dict(),prefix+'UCF_experiments/trained_models_cnn/models/state_dict_kinetics_'+name)
# -
# learn.export('/mnt/data/eugeniomarinelli/UCF_experiments/trained_models_cnn/learners/LEARNER_CANCELLARE')
save_learner(learn, 'CANCELLARE')
b = torch.load('/mnt/data/eugeniomarinelli/UCF_experiments/trained_models_cnn/learners/learner_ucf101_XXXXXX.pth')
saved_model_SCL = torch.load('/mnt/data/eugeniomarinelli/UCF_experiments/trained_models_cnn/models/r2p1d50_ucf101_SCL_tuned_15fr.pth')
# +
model = nn.Sequential(saved_model_SCL,nn.Sequential(nn.ReLU(inplace=True),LinBnDrop(256, 101, p=0.5)))
# -
from nbdev.export import *
notebook2script()
# +
#default_exp kinetics
# -
import cv2
files = df.vid_files
corrupted = L()
for file in files:
if os.path.exists(file):
stat = os.stat(file)
if stat.st_size < 50000:
corrupted.append(str(file))
else:
print(f"{file} Not found")
corrupted
# %time cap = cv2.VideoCapture('/mnt/data/adrianlopez/Datasets/kinetics700/Videos/Puppy pushes carts.mp4')
# %time duration = cap.get(cv2.CAP_PROP_FRAME_COUNT)
cap.release()
cap = cv2.VideoCapture('/mnt/data/adrianlopez/Datasets/kinetics700/Videos/Watering.mp4')
cap.set(cv2.CAP_PROP_POS_FRAMES, 0)
print(cap.get(cv2.CAP_PROP_FRAME_COUNT))
cap.release()
cap.get(cv2.CAP_PROP_FRAME_COUNT)
| KINETICS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## AlexNet in Keras
# #### We are goin to classify Oxford Flowers
# ###Set Seed
import numpy as np
np.random.seed(42)
# #### Load Dependencies
import keras
from keras.models import Sequential
from keras.layers import Dense,Dropout,Flatten,Conv2D,MaxPooling2D
from keras.layers.normalization import BatchNormalization
from keras.callbacks import TensorBoard
# #### Load and Preprocess Data
import tflearn.datasets.oxflower17 as oxflower17
X,y = oxflower17.load_data(one_hot=True)
# #### Design Neural Network
# +
model = Sequential()
model.add(Conv2D(96,kernel_size=(11,11),strides=(4,4),activation='relu',input_shape=(224,224,3)))
model.add(MaxPooling2D(pool_size=(3,3),strides=(2,2)))
model.add(BatchNormalization())
model.add(Conv2D(256,kernel_size=(5,5),strides=(1,1),activation='relu'))
model.add(MaxPooling2D(pool_size=(3,3),strides=(2,2)))
model.add(BatchNormalization())
model.add(Conv2D(256,kernel_size=(3,3),activation='relu'))
model.add(Conv2D(384,kernel_size=(3,3),activation='relu'))
model.add(Conv2D(384,kernel_size=(3,3),activation='relu'))
model.add(MaxPooling2D(pool_size=(3,3),strides=(2,2)))
model.add(BatchNormalization())
model.add(Flatten())
model.add(Dense(4096,activation='tanh'))
model.add(Dropout(0.5))
model.add(Dense(4096,activation='tanh'))
model.add(Dropout(0.5))
model.add(Dense(17,activation='relu'))
# -
model.summary()
# #### Configure Model
model.compile(loss='categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
# #### Configure TensorBoard
tensorbrd = TensorBoard('logs/alexnet')
# #### Train !!
model.fit(X,y,batch_size=64,epochs=1,verbose=1,validation_split=0.1,shuffle=True)
| TFLiveLessons/alexnet_in_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Strings
# + [markdown] slideshow={"slide_type": "slide"}
# ### Declaration
# + [markdown] slideshow={"slide_type": "slide"}
# #### Using Single quotes
# + slideshow={"slide_type": "fragment"}
a_str = 'Single'
# + [markdown] slideshow={"slide_type": "slide"}
# #### Using Double quotes
# + slideshow={"slide_type": "fragment"}
b_str = "Double"
# + [markdown] slideshow={"slide_type": "slide"}
# #### Using both single and double quotes
# + slideshow={"slide_type": "fragment"}
mix_str = "I'm fine"
# + slideshow={"slide_type": "fragment"}
print(mix_str)
# + slideshow={"slide_type": "slide"}
another_mix_str = 'Students said, "The mentor is really boring"'
# + slideshow={"slide_type": "fragment"}
print(another_mix_str)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Multi line strings
# + slideshow={"slide_type": "slide"}
really_long_str = """You can declare multi-line string with triple quotes.
Both single quotes and double quotes work fine.
Use them according to your requirement.
"""
# + slideshow={"slide_type": "fragment"}
print(really_long_str)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Single line long strings
# + slideshow={"slide_type": "slide"}
a_long_str = (
'This is a really long string that could not be expressed'
' in one single line.')
# + slideshow={"slide_type": "fragment"}
print(a_long_str)
# + [markdown] slideshow={"slide_type": "slide"}
# ### More about strings
# + slideshow={"slide_type": "slide"}
my_str = 'Python'
# -
type(my_str)
len(my_str)
my_str[-1]
my_str[0] = 's'
# + slideshow={"slide_type": "fragment"}
# type, len, index, mutable
# + slideshow={"slide_type": "slide"}
# + [markdown] slideshow={"slide_type": "slide"}
# ### String Formatting
# + slideshow={"slide_type": "fragment"}
action = 'Programming'
# + slideshow={"slide_type": "slide"}
lang = 'Python'
# + slideshow={"slide_type": "fragment"}
print('{} is a programming language'.format(lang))
# + slideshow={"slide_type": "fragment"}
name = 'foo'
age = 85
'{} is {} years old'.format(name, age)
# -
my_str = 'My string has a {}'
print(my_str)
# + slideshow={"slide_type": "slide"}
my_str.format(4)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Using `format` with keyword arguments
# -
name = 'foo'
age = 84
# + slideshow={"slide_type": "fragment"}
'{n} is {a} years old'.format(n=name, a=age)
# + slideshow={"slide_type": "fragment"}
# + slideshow={"slide_type": "slide"}
# + [markdown] slideshow={"slide_type": "slide"}
# #### Using `format` with both positional and keyword arguments
# + slideshow={"slide_type": "fragment"}
'{n} is {a} years old and lives on the planet {}'.format('earth', n=name, a=age)
# + slideshow={"slide_type": "slide"}
'{n} is {a} years old and lives on the planet {}'.format(n=name, a=age, 'earth')
# + [markdown] slideshow={"slide_type": "slide"}
# ### F strings
# + slideshow={"slide_type": "fragment"}
f'{name} is {age} years old'
# +
num = 30
divisor = 3
f'{num} when divided by {divisor} gives a remainder {num % divisor }'
# -
# + [markdown] slideshow={"slide_type": "slide"}
# #### Performing operations inside f-strings
# -
print(f'my name is {name}')
name
# + slideshow={"slide_type": "fragment"}
year = 1990
f'{name} was born in {year}. Their age is {2020 - year}.'
# + [markdown] slideshow={"slide_type": "slide"}
# ### Format Specifiers
# + slideshow={"slide_type": "fragment"}
country = 'India'
capital = 'New Delhi'
'%s is the capital of %s' % (capital, country)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Some Basic Argument Specifier
# + [markdown] slideshow={"slide_type": "fragment"}
# Specifier | Use|
# ------|-------------|
# `%s` | String (or any object with a string representation, like numbers).|
# `%d` | Integers.|
# `%f` | Floating point numbers.|
# `%.<number of digits>f` | Floating point numbers with a fixed amount of digits to the right of the dot.|
# + [markdown] slideshow={"slide_type": "slide"}
# #### Some more examples
# + slideshow={"slide_type": "fragment"}
num = 12
dec = 11.123656
print('%.3f' % (dec))
print('%d' % (num))
print('%f' % (num))
# + slideshow={"slide_type": "fragment"}
# same examples with f-strings
print(f'{dec:.3f}')
print(f'{num:d}')
print(f'{dec:f}')
# + [markdown] slideshow={"slide_type": "notes"}
# Example of a function to round off numbers with precision
# + [markdown] slideshow={"slide_type": "slide"}
# ### Problem
#
# Write a function that rounds off number with the given number of digits after decimal. When no value is passed for precision, it rounds off upto 2 decimal places.
#
# for example:
# ```python
# # my_fun is the function that you will write
# >>> my_fun(24.1345)
# 24.13
# >>> my_fun(24.13457, 4)
# 24.1346
# ```
# + slideshow={"slide_type": "slide"}
def round_off_num(num, precision=2):
return f'{num:.{precision}f}'
round_off_num(23.236, precision=2)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Basic Operations
# + [markdown] slideshow={"slide_type": "slide"}
# #### Slicing
# -
my_str = 'Python'
# characters at odd positions(not index)
my_str[::2]
my_str[::-1]
# + slideshow={"slide_type": "slide"}
# + [markdown] slideshow={"slide_type": "slide"}
# #### Case(Lower/Upper) Manipulations
# -
name = 'guido'
print(name.upper())
name = 'Guido'
name.lower()
name = 'guido'
name.capitalize()
# + slideshow={"slide_type": "slide"}
name = 'foo'
'{{{name}}}'.format(name=name)
# + [markdown] slideshow={"slide_type": "slide"}
# #### Checking for `substring`
# + slideshow={"slide_type": "fragment"}
name = 'guido'
'gu' in name.lower()
# -
'gud' in name.lower()
'G' in name.lower()
# + slideshow={"slide_type": "slide"}
# + slideshow={"slide_type": "slide"}
# + [markdown] slideshow={"slide_type": "slide"}
# #### Cleaning
# + slideshow={"slide_type": "slide"}
name = ' Guido is the author '
# removes the left hand whitespace
name.lstrip()
# removes the right hand whitespace
name.rstrip()
# removes the whitespace from both left and right
name.strip()
# + slideshow={"slide_type": "fragment"}
"AABAA".lstrip("A")
# + slideshow={"slide_type": "fragment"}
"ABBA".lstrip("AB") # both AB and BA are stripped
# + slideshow={"slide_type": "fragment"}
"ABCABBA".rstrip("AB")
# + slideshow={"slide_type": "fragment"}
"ABCABBA".lstrip("AB")
# + [markdown] slideshow={"slide_type": "slide"}
# #### Splitting
# + slideshow={"slide_type": "fragment"}
sentence = ' <NAME> is the author of Python '
# + slideshow={"slide_type": "slide"}
len(sentence.split())
# + slideshow={"slide_type": "slide"}
my_str = 'abcabcabcabc'
# + slideshow={"slide_type": "fragment"}
my_str.split('bc')
# + slideshow={"slide_type": "fragment"}
my_str.split('bc', 1)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Iterating through a string
# + slideshow={"slide_type": "fragment"}
my_str = 'Python'
for char in my_str:
print(char)
# + [markdown] slideshow={"slide_type": "slide"}
# ### `find` and `count` function
# + slideshow={"slide_type": "slide"}
my_str.find('p')
# + slideshow={"slide_type": "fragment"}
my_str.find('t')
# + slideshow={"slide_type": "fragment"}
my_str.count('t')
# + slideshow={"slide_type": "fragment"}
my_str.count('p')
# + slideshow={"slide_type": "slide"}
'abcabcabc'.count('b')
# + slideshow={"slide_type": "slide"}
'abcabc'.find('a')
# + [markdown] slideshow={"slide_type": "slide"}
# #### Problem
#
# Write a function that return the positions where a character(one letter string) is present inside a string as a list. If the character is not present even once, it should return 0.
#
# Example:
# ```python
# >>> find_indexes('abcabcabc', 'a')
# [1, 4, 7]
# ```
# > *Explanation*: The character `a` is not present at positions(it is **1 based not zero based**) 1, 4 and 7 inside the string.
#
# ```python
# >> find_indexes('abcabcabc', 'z')
# 0
# ```
# > *Explanation*: The character `z` is not present inside the string.
#
# + slideshow={"slide_type": "slide"}
# + [markdown] slideshow={"slide_type": "slide"}
# ### Some more useful functions
# + slideshow={"slide_type": "fragment"}
'Guido'.startswith('G')
# + slideshow={"slide_type": "fragment"}
'Guido'.endswith('do')
# + slideshow={"slide_type": "fragment"}
'abc' + 'abc'
# -
| chapter_09/strings_in_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import pandas as pd
import numpy as np
from nltk.tokenize import sent_tokenize
from nltk.metrics import edit_distance
from nltk import (word_tokenize, pos_tag)
import hunspell
import argparse
import sys
import csv
import os
import re
from tqdm import tqdm
import config as cf
import preprocess as pp
# +
def loadDictionary():
with open(dataPath + '%s_filtered.txt' % cf.WORD2VEC_FILE[0:-4], 'r') as f:
for line in f:
values = line.split()
word = pp.joinWord(values[:-300])
vector = np.array(values[-300:], dtype='float32')
dictionary[word] = vector
f.close()
def encodeReview(filePath, state):
index = list(dictionary.keys())
header = ['reviewid', 'sentence']
encoding = pd.DataFrame(columns = header)
with open(filePath) as json_data:
max = 0
j = 0
reviewIndex = []
for i, line in enumerate(tqdm(json_data)):
review = json.loads(line)
print(review)
# texts = review['text']
# reviewId = review['review_id']
# sentences = sent_tokenize(texts)
# for sentence in sentences:
# encoding.loc[j] = [reviewId, sentence]
# j = j + 1
# sentenceIndex = []
# for word in word_tokenize(sentence):
# try:
# idx = index.index(word)
# except ValueError:
# idx = 4859
# sentenceIndex.append(idx)
# if max < len(sentenceIndex):
# max = len(sentenceIndex)
# textIndex.append(sentenceIndex)
# print(max)
# np.save(dataPath + '%s' % state, np.array(textIndex))
# +
# hyperparameters
batch_iterations = 10000
batch_size = 32
full_iterations = 200
learning_rate = 0.001
reg_eta = 0.001
# dimensionalities
dim_lstm = 300
dim_word = 300
dim_aspect = 5
dim_sentence = 80
# setup utils object
isSample = True
u = utils.UTILS(batch_size, dim_sentence, dim_aspect, isSample)
# define tf placeholders
X = tf.placeholder(tf.int32, [None, dim_sentence])
y = tf.placeholder(tf.float32, [None, dim_aspect])
seqlen = tf.placeholder(tf.int32, [None])
# define tf variables
with tf.variable_scope('bilstm_vars'):
with tf.variable_scope('weights', reuse = tf.AUTO_REUSE):
lstm_w = tf.get_variable(
name = 'softmax_w',
shape = [dim_lstm * 2, dim_aspect],
initializer = tf.random_uniform_initializer(-0.003, 0.003),
regularizer = tf.contrib.layers.l2_regularizer(reg_eta)
)
with tf.variable_scope('biases', reuse = tf.AUTO_REUSE):
lstm_b = tf.get_variable(
name = 'softmax_b',
shape = [dim_aspect],
initializer = tf.random_uniform_initializer(-0.003, 0.003),
regularizer = tf.contrib.layers.l2_regularizer(reg_eta)
)
# define lstm model
def dynamic_lstm(inputs, seqlen):
inputs = tf.nn.dropout(inputs, keep_prob=1.0)
with tf.name_scope('bilstm_model'):
forward_lstm_cell = tf.contrib.rnn.LSTMCell(dim_lstm)
backward_lstm_cell = tf.contrib.rnn.LSTMCell(dim_lstm)
outputs, states = tf.nn.bidirectional_dynamic_rnn(
forward_lstm_cell,
backward_lstm_cell,
inputs = inputs,
sequence_length = seqlen,
dtype = tf.float32,
scope = 'bilstm'
)
forward_outputs, backward_outputs = outputs
backward_outputs = tf.reverse_sequence(backward_outputs, tf.cast(seqlen, tf.int64), seq_dim=1)
outputs = tf.concat([forward_outputs, backward_outputs], 2)
size = tf.shape(outputs)[0]
index = tf.range(0, size) * dim_sentence + seqlen - 1
output = tf.gather(tf.reshape(outputs, [-1, dim_lstm * 2]), index) # batch_size * n_hidden * 2
predict = tf.matmul(output, lstm_w) + lstm_b
return predict
# define operations
pred = dynamic_lstm(tf.nn.embedding_lookup(u.gloveDict, X), seqlen)
loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits = pred, labels = y))
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(loss)
correct = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# -
dictionary = {}
dataPath = cf.ROOT_PATH + cf.DATA_PATH
loadDictionary()
encodeReview(dataPath + 'AZ_reviews.json', 'AZ')
| data_analysis/yelp_preprocess/aspects/aspect analysis test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math as m
v=float(input("Digite um numero: "))
print('O valor digitado foi {} e a sua porção inteira é igual a {}'.format(v,m.trunc(v)))
| .ipynb_checkpoints/EX016 - Quebrando um Número -checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 283} cellView="form" id="EXwKpF9sh1wY" outputId="76a0503f-36d8-4392-887a-9345b367ec04"
#@markdown 載入MNIST資料
from matplotlib import pyplot as plt
from tensorflow.keras.datasets import mnist
import numpy as np
(x_train, _ ), (x_test, _ ) = mnist.load_data()
plt.imshow(x_train[0])
# + colab={"base_uri": "https://localhost:8080/"} cellView="form" id="rWwVGqJqiAkQ" outputId="8f6d8ec9-18c3-4a52-f1bd-6f2d0e070611"
#@markdown 資料前處理
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
x_train = x_train.reshape((len(x_train), np.prod(x_train.shape[1:])))
x_test = x_test.reshape((len(x_test), np.prod(x_test.shape[1:])))
print(x_train.shape)
print(x_test.shape)
# + colab={"base_uri": "https://localhost:8080/"} cellView="form" id="9WPPfuTmiD7N" outputId="bf7242b2-cc61-4e0b-d853-df457bc75333"
#@markdown 建立模型
from tensorflow.keras.layers import Input, Dense
from tensorflow.keras.models import Model, Sequential
# Input : 784 -> Hidden: 32 -> Output:784
# this is the size of our encoded representations
encoding_dim = 32 # 32 floats -> compression of factor 24.5, assuming the input is 784 floats
# this is our input placeholder
input_img = Input(shape=(784,))
# "encoded" is the encoded representation of the input
encoded = Dense(encoding_dim, activation='relu')(input_img)
# "decoded" is the lossy reconstruction of the input
decoded = Dense(784, activation='sigmoid')(encoded)
# this model maps an input to its reconstruction
autoencoder = Model(input_img, decoded)
autoencoder.summary()
# + colab={"base_uri": "https://localhost:8080/"} cellView="form" id="rQs6HA_9iRMj" outputId="ced3a6ed-a708-48e6-e6a7-2edb277b5632"
#@markdown 模型訓練
autoencoder.compile(optimizer='adam', loss='binary_crossentropy')
epoch_num = 5 #@param {type:"integer"}
autoencoder.fit(x_train, x_train,
epochs=5,
batch_size=256,
shuffle=True,
validation_data=(x_test, x_test))
# + colab={"base_uri": "https://localhost:8080/"} cellView="form" id="rzypZWDFiXgd" outputId="d160c124-cecc-4df5-d14e-f31a691688f3"
#@markdown 用 Encoder 編碼資料
encoder = Model(input_img, encoded)
encoded_imgs = encoder.predict(x_test)
print('Encoded DIM')
print(encoded_imgs.shape)
print('Latent Space')
print(encoded_imgs)
# + colab={"base_uri": "https://localhost:8080/"} cellView="form" id="hz0ZeMrqislo" outputId="29fc420f-987a-43f4-87a8-a75c99ddceb5"
#@markdown 用Decoder 還原資料
encoded_input = Input(shape=(encoding_dim,))
decoder_layer = autoencoder.layers[-1]
decoder = Model(encoded_input, decoder_layer(encoded_input))
decoded_imgs = decoder.predict(encoded_imgs)
print('Decoded DIM')
print(decoded_imgs.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 248} cellView="form" id="Kvx0eXdgjAwq" outputId="7e80cfdd-7b05-49c2-c9ca-d4318d44d81b"
#@markdown 顯現還原結果
import matplotlib.pyplot as plt
n = 10 # how many digits we will display
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# + id="lgaZINR-jx-U"
| Autoencoders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
error = """\
Traceback (most recent call last):
File "<string>", line 1, in <module>
File "/tmp/tmp9ghq3pig/lib/python3.8/site-packages/pymedphys/_streamlit/utilities/misc.py", line 7, in <module>
from . import config as _config
File "/tmp/tmp9ghq3pig/lib/python3.8/site-packages/pymedphys/_streamlit/utilities/config.py", line 27, in <module>
@st.cache
File "/tmp/tmp9ghq3pig/lib/python3.8/site-packages/pymedphys/_vendor/apipkg/__init__.py", line 238, in __getattribute__
return getattr(getmod(), name)
File "/tmp/tmp9ghq3pig/lib/python3.8/site-packages/pymedphys/_vendor/apipkg/__init__.py", line 216, in getmod
x = importobj(modpath, None)
File "/tmp/tmp9ghq3pig/lib/python3.8/site-packages/pymedphys/_vendor/apipkg/__init__.py", line 103, in importobj
module = __import__(modpath, None, None, ["__doc__"])
ModuleNotFoundError: No module named 'streamlit'
"""
import re
import os
import itertools
re.search('apipkg', error)
error_list = error.split('\n')
error_list
[False].index(True)
has_apipkg = ['apipkg' in item for item in error_list]
has_apipkg
i = has_apipkg.index(True)
error_list[i-2:i]
module = re.search("(pymedphys.*)\.py", error_list[i-2]).group(1).replace(os.sep, '.').replace('-', '_')
module
line = re.search("line (\d+),", error_list[i-2]).group(1)
line
# +
next(filter(lambda item: 'apipkg' in item, error_list))
| prototyping/maintenance/regex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#
# ___ _
# / _/______ ____ ____(_)__ _______
# / _/ __/ _ `/ _ \/ __/ (_-</ __/ _ \
#/_//_/ \_,_/_//_/\__/_/___/\__/\___/
# ___ _____(_)__ ___ ____ / /_(_)
# / _ `/ __/ (_-</ _ `/ _ \/ __/ /
# \_, /_/ /_/___/\_,_/_//_/\__/_/
#/___/
#
#Samee Lab @ Baylor College Of Medicine
#<EMAIL>
# + tags=[]
import psutil
import platform
from datetime import datetime
print("="*40, "System Information", "="*40)
uname = platform.uname()
print(f"System: {uname.system}")
print(f"Node Name: {uname.node}")
print(f"Release: {uname.release}")
print(f"Version: {uname.version}")
print(f"Machine: {uname.machine}")
# + tags=[]
#Load libraries
import matplotlib.pyplot as plt
import numpy as np
import random
import os
import sys
import pandas as pd
import warnings
import seaborn as sb
import sklearn
import tqdm
import scanpy as sc
import anndata
sc.logging.print_header()
# -
sys.path.append('../')
from STANN.models import STANN
import STANN.utils as utils
# + tags=[]
import tensorflow as tf
import tensorflow.keras as keras
print(f'tensorflow=={tf.__version__} \nkeras=={tf.keras.__version__}')
# -
#Reproducibility
seed = 10
np.random.seed(seed)
tf.random.set_seed(seed)
# + tags=[]
################LOAD DATA###################
# check to see which data
print("[INFO] loading training data...")
adata_train = sc.read_h5ad("../data/scrna.h5ad")
print("[INFO] loading predict data...")
adata_predict = sc.read_h5ad("../data/seqfish.h5ad")
# + tags=[]
model = STANN(act_fun='tanh',
first_dense=160,
second_dense=145.0,
learning_rate=0.01,input_dim=adata_train.X.shape[1],
output_dim=len(adata_train.obs.celltype.unique()))
model.summary()
# + tags=[]
X_train, Y_train, X_predict = utils.organize_data(adata_train=adata_train,
adata_predict=adata_predict)
# -
X_train_scaled , scaler_train = utils.min_max(X=X_train)
#Sample obs distribution
fig, ax = plt.subplots(figsize=(5,5))
sb.distplot(X_train_scaled[0],bins=200,ax=ax)
ax.set_xlim(-2,2)
ax.set(xlabel='Expression', ylabel='Frequency')
ax.set_title('w/ normalization')
plt.show()
X_predict_scaled , scaler_predict = utils.min_max(X=X_predict)
#Sample obs distribution
fig, ax = plt.subplots(figsize=(5,5))
sb.distplot(X_predict_scaled[0],bins=200,ax=ax)
ax.set_xlim(-2,2)
ax.set(xlabel='Expression', ylabel='Frequency')
ax.set_title('w/ normalization')
plt.show()
Y_train_dummy,Y_train_ohe,encoder = utils.label_encoder(Y_train=Y_train)
# + tags=[]
x_train, x_test, y_train, y_test = utils.get_train_test_split(X_train_scaled,
Y_train_ohe,
test_size=0.10,
random_state=40)
# -
class_weights = utils.get_class_weights(Y_train_ohe=y_train)
class_weights = {i : class_weights[i] for i in range(15)}
class_weights
#early stopping
es = tf.keras.callbacks.EarlyStopping(monitor='accuracy',
mode='min',
verbose=1,
patience=30)
# + tags=[]
history = model.fit(x_train,
y_train,
validation_data=(x_test, y_test),
epochs=30,
class_weight=class_weights,
callbacks=[es])
# -
fig, axs = plt.subplots(figsize=(5,5))
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label='val_loss')
plt.plot(history.history['accuracy'], label='accuracy')
plt.plot(history.history['val_accuracy'], label='val_accuracy')
plt.title('Training metrics')
plt.xlabel('Iterations')
plt.legend()
# + tags=[]
utils.print_metrics(model=model,
x_train=x_train,
y_train=y_train,
x_test=x_test,
y_test=y_test)
# + tags=[]
predictions = utils.make_predictions(model=model,
X_predict=X_predict_scaled,
encoder=encoder,
adata_predict=adata_predict,
probabilities=False,
save=False
)
predictions
# -
# ## Calculate discrepancy
old_predictions = pd.read_csv("../../notebooks/classifier/predictions_updated.csv",index_col="barcode")
old_predictions.head()
results = pd.DataFrame({"old":old_predictions.prediction,"new":predictions.STANN_predictions})
# +
results['diff'] = None
for index, row in results.iterrows():
if row.old == row.new:
results.at[index,'diff'] = True
else:
results.at[index,'diff'] = False
# -
results['diff'].value_counts()
old_predictions
| notebooks/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sample Similarity
#
# [](https://colab.research.google.com/github/ing-bank/probatus/blob/master/docs/tutorials/nb_sample_similarity.ipynb)
#
# The goal of a Resemblance Model is understanding how different two samples are from a multivariate perspective. For instance, if you suspect that your Out-Of-Time Test set may have a different distribution than the In-Time Train set, you can detect that using the Resemblance Model.
#
# <img src="../img/resemblance_model_schema.png" width="900">
#
# Having two samples `X1` and `X2` with the same set of features, one can analyse how well a model can recognize which dataset a randomly selected row comes from. The Resemblance model assigns label `0` to `X1` dataset, and label `1` to `X2`. Then, the data is shuffled and split into Train split (`X_train`, `y_train`) and Test split (`X_test`, `y_test`).
#
# The user provides a binary classifier that is then fitted on the Train split and evaluated on both Train and Test.
# Interpreting such model allows to understand, which features and interactions between them differ between these two samples.
#
# It is crucial that the model does not overfit or underfit, because interpretation of such model will lead to wrong conclusions. Therefore, you should try fitting the model with a couple different hyperparameter settings, and make sure that `Train AUC` is not significantly higher than `Test AUC`
#
# Once you have the final Resemblance Model, the `Test AUC` significantly above 0.5 indicates predictive power of the model, as well as the change in the distribution between `X1` and `X2`. The higher the `Test AUC`, the larger the difference between two datasets.
#
# Then you can use interpret the model, in order to understand the patterns that the model has learned.
# There are two classes in `probatus` that allow you to analyse, which features have changed between two samples:
#
# - **SHAPImportanceResemblance (Recommended)** - Trains a Resemblance model based on a **tree classifier**, then it uses SHAP library to analyse the differences in features between the two samples. The main advantage of using this method is its high speed, better understanding of the relations in the data and handling of categorical features and missing values.
#
# - **PermutationImportanceResemblance** - Trains a Resemblance model for **any provided classifier**, and uses Permutation Importance to analyse, which features the model relies on. It is significantly slower, and requires preprocessing of the data before training the resemblance model.
#
# ## Setup
# %%capture
# !pip install probatus
# +
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
import numpy as np
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import pandas as pd
# Prepare two samples
feature_names = ['f1', 'f2', 'f3', 'f4']
X1 = pd.DataFrame(make_classification(n_samples=1000, n_features=4, random_state=0)[0], columns=feature_names)
X2 = pd.DataFrame(make_classification(n_samples=1000, n_features=4, shift=0.5, random_state=0)[0], columns=feature_names)
# Prepare model
clf = RandomForestClassifier(n_estimators = 100, max_depth=2, random_state=0)
# -
# ## SHAP Importance Resemblance Model for Tree models
# Below you can see an example of how to use the model:
# +
from probatus.sample_similarity import SHAPImportanceResemblance
rm = SHAPImportanceResemblance(clf)
feature_importance, train_auc, test_auc = rm.fit_compute(X1, X2, column_names=feature_names, return_scores=True)
display(feature_importance)
# -
# By looking into the above results, one can conclude that the two samples significantly differ, since the Test AUC of the model is very high.
#
# The table shows the mean absolute shap values and mean shap values for the model's features:
#
# - **Mean Absolute SHAP Values** provide insights about overall **SHAP feature importance**.
#
# - **Mean SHAP Values** show in which direction on average the feature influences the prediction. Negative value indicates 0 class, and positive indicates 1 class.
#
# Below, the SHAP feature importance is plotted
ax = rm.plot()
# In order to get more insights of the change in underlying relations in the data, let's plot a dot summary plot.
ax = rm.plot(plot_type='dot')
# We can see that second sample have higher values in all the features.
# ## Permutation Importance Resemblance Model
#
# Below we show the example on how to use the PermutationImportanceResemblance
# +
from probatus.sample_similarity import PermutationImportanceResemblance
perm = PermutationImportanceResemblance(clf)
feature_importance, train_auc, test_auc = perm.fit_compute(X1, X2, column_names=feature_names, return_scores=True)
display(feature_importance)
# -
# Same as before, we can get more insights into the importance of the features. However, now we can also analyse standard deviation of the permutation importance. High std might indicate that permutation of this feature has a higher or lower impact only in part of the available samples, while low std, indicates a consistent effect.
ax = perm.plot()
# ## Visualize the difference in the most important feature
#
# We can also use the utils to provide more insights into the feature distribution difference in the two samples.
# +
from probatus.utils.plots import plot_distributions_of_feature
feature_distributions = [X1['f3'], X2['f3']]
plot_distributions_of_feature(feature_distributions, plot_perc_outliers_removed=0.01)
| docs/tutorials/nb_sample_similarity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 3. PyCBC Tutorial: Generating Waveforms and Matched Filtering
#
# We will be using the [PyCBC](http://github.com/ligo-cbc/pycbc) library, which is used to study gravitational-wave data, find astrophysical sources due to compact binary mergers, and study their parameters. These are some of the same tools that the LIGO and Virgo collaborations use to find gravitational waves in LIGO/Virgo data
#
# In this tutorial we will walk through how find a specific signal in LIGO data. We present how to generate the waveform of a gravitational-wave merger and matched filtering, which is optimal in the case of Gaussian noise and a known signal model. In reality our noise is not entirely Guassian, and in practice we use a variety of techniques to separate signals from noise in addition to the use of the matched filter.
#
# Additional [examples](http://pycbc.org/pycbc/latest/html/#library-examples-and-interactive-tutorials) and module level documentation are [here](http://pycbc.org/pycbc/latest/html/py-modindex.html)
# #### Getting the software environment setup
#
# PyCBC is installable through pip, but also relies on portions of the [LALSuite](https://git.ligo.org/lscsoft/lalsuite) c-library. A bundled version of this suitable for use with PyCBC is also available on Mac / Linux through pip. These can be installed as follows within the notebook.
import sys
# !{sys.executable} -m pip install pycbc lalsuite ligo-common --no-cache-dir
# ### Generate your first waveform !
#
# Here we'll generate the gravitational waveform using one of the available waveform approximants. These can be generated as a time series using [`get_td_waveform`](http://pycbc.org/pycbc/latest/html/pycbc.waveform.html#pycbc.waveform.waveform.get_td_waveform). There are some additional examples using this interface [here](http://pycbc.org/pycbc/latest/html/waveform.html). The key parameters are the masses of the binary (given in solar masses), the time between samples (in seconds), the starting gravitational-wave frequency (Hz) and the name of the approximant we'd like to generate. A variety of approximants are available that include different physical effects. A full review of the different models is outside of the scope of this tutorial.
#
# In this example, we've chosen to use the 'SEOBNRv4_opt' model. There are many others available as well with different methodologies and which include different physical effects. This is an implementation of the model introduced [in this paper](https://arxiv.org/pdf/1611.03703.pdf). It models the gravitational waveform of inspiralling and merging black holes, and includes the ability for each black hole to spin in the same direction as the orbit (aligned spin).
# +
# %matplotlib inline
from pycbc.waveform import get_td_waveform
import pylab
# The output of this function are the "plus" and "cross" polarizations of the gravitational-wave signal
# as viewed from the line of sight at a given source inclination (assumed face-on if not provided)
hp, hc = get_td_waveform(approximant="SEOBNRv4_opt",
mass1=10,
mass2=10,
delta_t=1.0/4096,
f_lower=30)
pylab.plot(hp.sample_times, hp, label='Plus Polarization')
pylab.plot(hp.sample_times, hc, label='Cross Polarization')
pylab.xlabel('Time (s)')
pylab.legend()
pylab.grid()
pylab.show()
# Zoom in near the merger time#
pylab.plot(hp.sample_times, hp, label='Plus Polarization')
pylab.plot(hp.sample_times, hc, label='Cross Polarization')
pylab.xlabel('Time (s)')
pylab.xlim(-.01, .01)
pylab.legend()
pylab.grid()
pylab.show()
# -
#
# We can see that in the this case, the two polariations differ only by the phase of the signal. This holds for systems where the orbital plane of the binary doesn't precess. In the zoom-in plot, we can see the merger itself and the ringdown that follows.
# ### How does the waveform change with the mass of the binary?
#
# Below you can see how the length of the waveform increases for lower mass binary mergers.
# Component mass of each binary component. We'll simplify here and assume that each
# component of the binary has the same mass. Again, units are in solar masses.
for m in [5, 10, 30, 100]:
hp, hc = get_td_waveform(approximant="SEOBNRv4_opt",
mass1=m,
mass2=m,
delta_t=1.0/4096,
f_lower=30)
pylab.plot(hp.sample_times, hp, label='$M_{\odot 1,2}=%s$' % m)
pylab.legend()
pylab.grid()
pylab.xlabel('Time (s)')
pylab.show()
# ### Changing the distance of the waveform
#
# The distance of the waveform is also selectable when you generate a waveform. The units used are Megaparsecs. Keep in mind that no redshift effects are taken into account here, so there is a simple linear relationship between distance and amplitude
for d in [100, 500, 1000]:
hp, hc = get_td_waveform(approximant="SEOBNRv4_opt",
mass1=10,
mass2=10,
delta_t=1.0/4096,
f_lower=30,
distance=d)
pylab.plot(hp.sample_times, hp, label='$Distance=%sMpc$' % d)
pylab.legend()
pylab.grid()
pylab.xlabel('Time (s)')
pylab.show()
# ### Looking for a specific signal in the data
#
# If you know what signal you are looking for in the data, then matched filtering is known to be the optimal method in Gaussian noise to extract the siganl. Even when the parameters of the signal are unkown, one can test for each set of parameters one is interesting in finding.
# #### preconditioning the data
#
# The purpose of this is to reduce the dynamic range of the data and supress low freqeuncy behavior which can introduce numerical artefacts. We may also wish to resample the data if high frequency content is not important.
# +
# As an example we use the GW150914 data
from pycbc.catalog import Merger
from pycbc.filter import resample_to_delta_t, highpass
merger = Merger("GW150914")
# Get the data from the Hanford detector
strain = merger.strain('H1')
# Remove the low frequency content and downsample the data to 2048Hz
strain = resample_to_delta_t(highpass(strain, 15.0), 1.0/2048)
pylab.plot(strain.sample_times, strain)
pylab.xlabel('Time (s)')
pylab.show()
# -
# #### filter wraparound
#
# Note the spike in the data at the boundaries. This is caused by the highpass and resampling stages filtering the data. When the filter is applied to the boundaries, it wraps around to the beginning of the data. Since the data itself has a discontinuity (i.e. it is not cyclic) the filter itself will ring off for a time up to the length of the filter.
#
# Even if a visible transient is not seen, we want to avoid filters that act on times which are not causally connect. To avoid this we trim the ends of the data sufficiently to ensure that they do not wraparound the input. We will enforce this requirement in all steps of our filtering.
# +
# Remove 2 seconds of data from both the beginning and end
conditioned = strain.crop(2, 2)
pylab.plot(conditioned.sample_times, conditioned)
pylab.xlabel('Time (s)')
pylab.show()
# -
# #### calculate the power spectral density
#
# Optimal matched filtering requires weighting the frequency components of the potential signal and data by the noise amplitude. We can view this as filtering the data with the time series equivelant of 1 / PSD. To ensure that we can control how much applying this filter to the data, we window the time domain equivelant of the PSD to a specific length. This has effect of losing some information about line behavior in the detector, however, since our signal span a large frequency range, and lines are narrow, this is a negligible effect.
# +
from pycbc.psd import interpolate, inverse_spectrum_truncation
# Estimate the power spectral density
# We use 4 second samles of our time series in Welch method.
psd = conditioned.psd(4)
# Now that we have the psd we need to interpolate it to match our data
# and then limit the filter length of 1 / PSD. After this, we can
# directly use this PSD to filter the data in a controlled manner
psd = interpolate(psd, conditioned.delta_f)
# 1/PSD will now act as a filter with an effective length of 4 seconds
# Since the data has been highpassed above 15 Hz, and will have low values
# below this we need to informat the function to not include frequencies
# below this frequency.
psd = inverse_spectrum_truncation(psd, int(4 * conditioned.sample_rate),
low_frequency_cutoff=15)
pylab.loglog(psd.sample_frequencies, psd)
pylab.ylabel('$Strain^2 / Hz$')
pylab.xlabel('Frequency (Hz)')
pylab.xlim(30, 1024)
# -
# #### make your signal model
#
# Conceptually, matched filtering involves laying the potential signal over your data and integrating (after weighting frequencies correctly). If there is a signal in the data that aligns with your 'template', you will get a large value when integrated over.
# +
# In this case we "know" what the signal parameters are. In a search
# we would grid over the parameters and calculate the SNR time series
# for each one
# We'll assume equal masses, which is within the posterior probability
# of GW150914.
m = 36 # Solar masses
hp, hc = get_td_waveform(approximant="SEOBNRv4_opt",
mass1=m,
mass2=m,
delta_t=conditioned.delta_t,
f_lower=20)
# We will resize the vector to match our data
hp.resize(len(conditioned))
# The waveform begins at the start of the vector, so if we want the
# SNR time series to correspond to the approximate merger location
# we need to shift the data so that the merger is approximately at the
# first bin of the data.
# This function rotates the vector by a fixed amount of time.
# It treats the data as if it were on a ring. Note that
# time stamps are *not* in general affected, but the true
# position in the vector is.
#
# By convention waveforms returned from `get_td_waveform` have their
# merger stamped with time zero, so we can use the start time to
# shift the merger into position
template = hp.cyclic_time_shift(hp.start_time)
pylab.plot(template)
pylab.show()
# -
# #### calculating the signal-to-noise time series
#
# In this section we will now calculate the signal-to-noise time series for our template. We'll take care to handle issues of filter corruption / wraparound by truncating the output time series. We need to account for both the length of the template and 1 / PSD.
# +
from pycbc.filter import matched_filter
import numpy
snr = matched_filter(template, conditioned,
psd=psd, low_frequency_cutoff=20)
# Remove time corrupted by the template filter and the psd filter
# We remove 4 seonds at the beginning and end for the PSD filtering
# And we remove 4 additional seconds at the beginning to account for
# the template length (this is somewhat generous for
# so short a template). A longer signal such as from a BNS, would
# require much more padding at the beginning of the vector.
snr = snr.crop(4 + 4, 4)
# Why am I taking an abs() here?
# The `matched_filter` function actually returns a 'complex' SNR.
# What that means is that the real portion correponds to the SNR
# associated with directly filtering the template with the data.
# The imaginary portion corresponds to filtering with a template that
# is 90 degrees out of phase. Since the phase of a signal may be
# anything, we choose to maximize over the phase of the signal.
pylab.figure(figsize=[10, 4])
pylab.plot(snr.sample_times, abs(snr))
pylab.ylabel('Signal-to-noise')
pylab.xlabel('Time (s)')
pylab.show()
peak = abs(snr).numpy().argmax()
snrp = snr[peak]
time = snr.sample_times[peak]
print("We found a signal at {}s with SNR {}".format(time,
abs(snrp)))
# -
# ### Aligning and Subtracting the Proposed Signal
#
# In the previous section we ound a peak in the signal-to-noise for a proposed binary black hole merger. We can use this SNR peak to align our proposal to the data, and to also subtract our proposal from the data.
# +
from pycbc.filter import sigma
# The time, amplitude, and phase of the SNR peak tell us how to align
# our proposed signal with the data.
# Shift the template to the peak time
dt = time - conditioned.start_time
aligned = template.cyclic_time_shift(dt)
# scale the template so that it would have SNR 1 in this data
aligned /= sigma(aligned, psd=psd, low_frequency_cutoff=20.0)
# Scale the template amplitude and phase to the peak value
aligned = (aligned.to_frequencyseries() * snrp).to_timeseries()
aligned.start_time = conditioned.start_time
# -
# #### Visualize the overlap between the signal and data
#
# To compare the data an signal on equal footing, and to concentrate on the frequency range that is important. We will whiten both the template and the data, and then bandpass both the data and template between 30-300 Hz. In this way, any signal that is in the data is transformed in the same way that the template is.
# +
# We do it this way so that we can whiten both the template and the data
white_data = (conditioned.to_frequencyseries() / psd**0.5).to_timeseries()
# apply a smoothing of the turnon of the template to avoid a transient
# from the sharp turn on in the waveform.
tapered = aligned.highpass_fir(30, 512, remove_corrupted=False)
white_template = (tapered.to_frequencyseries() / psd**0.5).to_timeseries()
white_data = white_data.highpass_fir(30., 512).lowpass_fir(300, 512)
white_template = white_template.highpass_fir(30, 512).lowpass_fir(300, 512)
# Select the time around the merger
white_data = white_data.time_slice(merger.time-.2, merger.time+.1)
white_template = white_template.time_slice(merger.time-.2, merger.time+.1)
pylab.figure(figsize=[15, 3])
pylab.plot(white_data.sample_times, white_data, label="Data")
pylab.plot(white_template.sample_times, white_template, label="Template")
pylab.legend()
pylab.show()
# -
# #### Subtracting the signal from the data
#
# Now that we've aligned the template we can simply subtract it. Let's see below how that looks in the time-frequency plots!
# +
subtracted = conditioned - aligned
# Plot the original data and the subtracted signal data
for data, title in [(conditioned, 'Original H1 Data'),
(subtracted, 'Signal Subtracted from H1 Data')]:
t, f, p = data.whiten(4, 4).qtransform(.001,
logfsteps=100,
qrange=(8, 8),
frange=(20, 512))
pylab.figure(figsize=[15, 3])
pylab.title(title)
pylab.pcolormesh(t, f, p**0.5, vmin=1, vmax=6)
pylab.yscale('log')
pylab.xlabel('Time (s)')
pylab.ylabel('Frequency (Hz)')
pylab.xlim(merger.time - 2, merger.time + 1)
pylab.show()
# -
# ## Challenge!
#
# Use the methods demonstrated above to see if you can calculate the SNR
# time series in the following data sets. What is the SNR of each signal?
# Which template matched which data?
#
# Information that may be useful:
#
# * Signals are all placed between 100 and 120 seconds into the frame file.
# * You may assume mass1 = mass2 (equal mass) and that each component mass is one of 22, 36, or 50.
# * Each file starts at gps time 0, and ends at gps time 128
# * The channel name in each file is "H1:TEST-STRAIN"
# +
# Download the challenge set files
from six.moves.urllib import request
def get_file(fname):
url = "https://github.com/ahnitz/odw-storage/raw/master/{}"
url = url.format(fname)
request.urlretrieve(url, fname)
print('Getting : {}'.format(url))
files = ['PyCBC_T2_0.gwf', 'PyCBC_T2_1.gwf', 'PyCBC_T2_2.gwf',
'PyCBC_T2_3.gwf', 'PyCBC_T2_4.gwf']
for fname in files:
get_file(fname)
| PyCBC-Tutorials-master/tutorial/.ipynb_checkpoints/3_WaveformMatchedFilter-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="0elixoydhII-"
# # **Otimização de Processos (COQ897)**
# # *Prof. <NAME>*
#
# $\
# $
#
# Primeira Lista de Exercícios - 2020
#
# $\
# $
#
# ***<NAME>***
#
# $\
# $
#
# 1) A engenheira <NAME> (!), responsável por um determinado processo químico, notou, ainda na fase de projeto da planta, a ocorrência da reação de hidrogenação do eteno:
#
# >$$C_{2}H_{4} + H_{2} \ {\rightleftarrows} \ C_{2}H_{6}$$
# >
# >$$n_{2} \qquad n_{3} \qquad n_{1}$$
#
#
# que para fins do processo em questão é indesejada. Querendo saber a quantidade de eteno que seria perdida no processo, Diana decidiu calcular o número de mols $n_{1}$ , $n_{2}$ e $n_{3}$ das espécies em equilíbrio, lembrando que, no equilíbrio, a energia de Gibbs total do sistema, $G_{t}$($n_{1}, n_{2}, n_{3}$) é mínima. Sabendo que as espécies atômicas se conservam, qual foi o problema de otimização formulado pela Eng. Diana?
# + [markdown] id="ZMCFr-wTvJ_G"
# $\
# $
#
# ## ***Solução:***
#
# O problema de otimização em questão é minimizar a função objetivo dada pela energia livre de Gibbs (G).
#
# Para qualquer processo infinitesimal em que a quantidade de espécies
# presente pode ser alterada pela transferência de espécies de/para uma fase ou por reação química, o diferencial da energia livre de Gibbs é dado por:
#
# >$dG= SdT + VdP + \sum\limits_{i=1}^{n} \ \mu_{i}dn_{i} \qquad (1)$
#
# Onde G, S, T e P são: a energia livre de Gibbs, a entropia, a temperatura e a
# pressão (total), respectivamente. A energia livre molal parcial da espécie i é $\mu_{i}$ (potencial químico),
# e $n_{i}$ é o número de moles da espécie i no sistema.
#
# Se for assumido que a temperatura e a pressão são mantidas constantes durante o processo, dT e dP desaparecem.
# Se agora fizermos alterações em $n_{i}$ de modo que $dn_{i} = dkn_{i}$, com as variações em $n_{i}$ na mesma proporção k; então, uma vez que G é uma quantidade extensiva, devemos ter $dG = dkG$. Isso implica que:
#
# >$G=\sum\limits_{i=1}^{n} \ \mu_{i}n_{i} \qquad (2)$
#
# A comparação das Equações (1) e (2) mostra que os potenciais químicos são quantidades intensivas, ou seja, não dependem da quantidade de cada espécie, pois se todos os $n_{i}$ são aumentados na mesma proporção com T e P constantes, $\mu_{i}$ deve permanecer inalterado para G aumentar na mesma taxa que $n_{i}$. Esta propriedade de invariância do $\mu_{i}$ é de extrema importância para restringir as formas possíveis que o $\mu_{i}$ pode assumir.
#
# A equação (2) expressa a energia livre de Gibbs em termos dos números molares $n_{i}$, que aparecem explícita e implicitamente (no $\mu_{i}$) no lado direito.
# A energia livre de Gibbs é mínima quando o sistema está em equilíbrio. O problema básico, então, torna-se o de encontrar aquele conjunto de $n_{i}$ que torna G um mínimo.
#
# Sendo $n_{i}^*$ o número de moles dos compostos em equilíbrio e M (3) o número de elementos presentes no sistema, e presumindo que o número inicial de moles de cada composto é conhecido:
#
#
# O problema consiste em (com T e P ctes):
#
# >$Minimizar \quad G= \sum\limits_{i=1}^{M = 3} \ (\mu_{i}^o + RT lnP +RT lnx_{i}) n_{i}
# \\
# G= RTlnP + [\sum\limits_{i} \ \mu_{i}^o + RT \sum\limits_{i} lnx_{i}] (n_{i})
# \\
# ,\ com \quad RTlnP=cte, \quad \mu_{i}^o = \sum\limits_{i} \ RTlnK_{x}, \quad x_{i} = \frac{n_{i}}{n} = \frac{n_{i}}{\sum{n_{i}}}
# \\
# ,\ sujeito \ ao \ balanço \ estequiométrico:
# \\
# \sum\limits_{i} \ a_{ik}n_{i} = b_{k}, \quad para \ cada \ um \ dos \ elementos \ k = 1 \ ...\ M(=3)
# \\
# e \ restrições \ de \ desigualdade:
# \\
# n_{i} \geq 0
# \\
# , \ com \quad n_{i} = x_{i}n.
# \\
# Para \ (2) + (3) \ {\rightleftarrows} \ (1), \quad K_{x} = (\frac{n_{1}}{n_{T}}) / (\frac{n_{2}}{n_{T}})(\frac{n_{3}}{n_{T}})
# $
#
#
#
# + [markdown] id="QHTvkhpvVhMp"
# $
# \\
# $
#
# 2) Dada a função objetivo $ S(x_{1}, x_{2}) = 7,5 x_{1}^2 + 12x_{2}^2 -3 x_{1}^2x_{2}^2 + 18x_{1} +11$, determine a
# localização e a natureza (mínimo, máximo ou sela) dos seus pontos estacionários.
# Esboce o gráfico da superfície da função objetivo em função de $x_{1}$ e $x_{2}$ e outro gráfico
# com 50 curvas de níveis, ambos contendo todos os pontos estacionários encontrados.
# Indique no segundo gráfico a localização dos pontos estacionários.
# + [markdown] id="Vvwi1lXT3jqz"
# $
# \\
# $
#
# ## ***Solução:***
#
# >$ S(x_{1}, x_{2}) = 7,5 x_{1}^2 + 12x_{2}^2 -3 x_{1}^2x_{2}^2 + 18x_{1} +11
# \\
# \nabla S(x_{1}, x_{2}) =
# \begin{pmatrix}
# 15x_{1} -6x_{1}x_{2}^2 + 18 \\
# 24x_{2} -6x_{1}^2x_{2}
# \end{pmatrix}
# \\
# Então, para \ encontrar \ o \ ponto \ ótimo \ x^*(x_{1}, x_{2}) \ em \ que \ \nabla S(x_{1}, x_{2}) = 0:
# $
# + colab={"base_uri": "https://localhost:8080/"} id="2fmoONUeWYnX" outputId="58a45807-e263-4743-fb45-9d2a8d1761a8"
import numpy as np
import scipy.integrate
import matplotlib.pyplot as plt
# %matplotlib inline
import scipy.optimize
# definindo o sistema de equações como uma função do Python
def func (x):
return [15.0*x[0] - (6.0*x[0])*(x[1])**2. + 18.0,
24.0*x[1] - (6.0*x[1])*(x[0])**2.]
# estimativa inicial
x0 = [0, 0]
# resolvendo
result = scipy.optimize.root(func, x0)
# imprimindo resultado
print(result)
print(result.x)
# + colab={"base_uri": "https://localhost:8080/"} id="9fLbAWwkCnO7" outputId="a11d9c82-11aa-49e1-d40c-eb91f70df6ea"
def func (x):
return [15.0*x[0] - (6.0*x[0])*(x[1])**2. + 18.0,
24.0*x[1] - (6.0*x[1])*(x[0])**2.]
# mudando a estimativa inicial
x0 = [-5, -5]
# resolvendo
result = scipy.optimize.root(func, x0)
# imprimindo resultado
print(result)
print(result.x)
# + colab={"base_uri": "https://localhost:8080/"} id="50nIE3FQMJJU" outputId="3af1a620-499b-4d3d-ac87-ac38b632899b"
def func (x):
return [15.0*x[0] - (6.0*x[0])*(x[1])**2. + 18.0,
24.0*x[1] - (6.0*x[1])*(x[0])**2.]
# mudando a estimativa inicial
x0 = [4, 1]
# resolvendo
result = scipy.optimize.root(func, x0)
# imprimindo resultado
print(result)
print(result.x)
# + colab={"base_uri": "https://localhost:8080/"} id="FIcgXveDsLsU" outputId="38b48e6a-2f1b-46e0-e5b1-9bbcd5b78c66"
def func (x):
return [15.0*x[0] - (6.0*x[0])*(x[1])**2. + 18.0,
24.0*x[1] - (6.0*x[1])*(x[0])**2.]
# mudando a estimativa inicial
x0 = [3, -3]
# resolvendo
result = scipy.optimize.root(func, x0)
# imprimindo resultado
print(result)
print(result.x)
# + colab={"base_uri": "https://localhost:8080/"} id="d6rNzv6Hs3XM" outputId="405d178c-b23e-4eae-94cc-daa28f5ddb10"
def func (x):
return [15.0*x[0] - (6.0*x[0])*(x[1])**2. + 18.0,
24.0*x[1] - (6.0*x[1])*(x[0])**2.]
# mudando a estimativa inicial
x0 = [-3, 3]
# resolvendo
result = scipy.optimize.root(func, x0)
# imprimindo resultado
print(result)
print(result.x)
# + [markdown] id="HiSKRinIDlWc"
# $
# \\
# $
#
# >$
# x^* =
# \begin{pmatrix}
# -1,2 \\
# 0,0
# \end{pmatrix},
# \begin{pmatrix}
# -2,0 \\
# -1,0
# \end{pmatrix},
# \begin{pmatrix}
# 2,0 \\
# 2,0
# \end{pmatrix}
# \begin{pmatrix}
# 2,0 \\
# -2,0
# \end{pmatrix}
# \begin{pmatrix}
# -2,0 \\
# 1,0
# \end{pmatrix}
# \\
#
# Calculando \ a \ matriz \ Hessiana:
# \\
# H(x) =
# \begin{pmatrix}
# 15 -6x_{2}^2 & \ -12x_{1}x_{2} \\
# -12x_{1}x_{2} & \ 24-6x_{1}^2
# \end{pmatrix}
# \\
# No \ ponto \ ótimo\ x^* =
# \begin{pmatrix}
# -1,2 \\
# 0,0
# \end{pmatrix}:
# \\
# H(x^*) =
# \begin{pmatrix}
# 15 & 0 \\
# 0 & 15,36
# \end{pmatrix}
# \\
# Logo,\ a \ matriz\ H(x^*)\ é \ positiva \ definida \ neste \ ponto, \ o \ que \ implica \ em \ x^* =
# \begin{pmatrix}
# -1,2 \\
# 0,0
# \end{pmatrix}\ ser \ um \ ponto \ de
# \\
# mínimo \ local.
# $
# + [markdown] id="mMWx0lKJM1U9"
# $
# \\
# $
#
# >$
#
# No \ ponto \ ótimo\ x^* =
# \begin{pmatrix}
# -2,0 \\
# -1,0
# \end{pmatrix}:
# \\
# H(x^*) =
# \begin{pmatrix}
# 9 & -24 \\
# -24 & 0
# \end{pmatrix}
# \\
# Como\ a \ matriz\ H(x^*)\ não \ é \ diagonal \ neste \ ponto, \ temos \ que \ calcular \ seus \ autovalores \ (\lambda):
#
# $
# + colab={"base_uri": "https://localhost:8080/"} id="cSIAvqsILSYV" outputId="156ef155-f544-4680-9c73-b6ed7a23cbae"
B = np.array([[9, -24],
[-24, 0]])
sigma = np.linalg.eigvals(B)
sigma
# + [markdown] id="NG2ErwmJOXYU"
# $
# \\
# $
# >$
#
# \lambda =
# \begin{pmatrix}
# 28,9\\
# -19,9
# \end{pmatrix}
# \\
# Logo,\ a \ matriz\ H(x^*)\ não \ é \ definida \ neste \ ponto, \ o \ que \ implica \ em \ x^* =
# \begin{pmatrix}
# -2,0 \\
# -1,0
# \end{pmatrix}\ ser \ um \ ponto \ de
# \\
# sela.
#
# $
# + [markdown] id="fBuDjR3XPYYc"
# $
# \\
# $
#
# >$
#
# No \ ponto \ ótimo\ x^* =
# \begin{pmatrix}
# 2,0 \\
# 2,0
# \end{pmatrix}:
# \\
# H(x^*) =
# \begin{pmatrix}
# -9 & -48 \\
# -48 & 0
# \end{pmatrix}
# \\
# Como\ a \ matriz\ H(x^*)\ não \ é \ diagonal \ neste \ ponto, \ temos \ que \ calcular \ seus \ autovalores \ (\lambda):
#
# $
# + colab={"base_uri": "https://localhost:8080/"} id="GwplO-BzPnkS" outputId="c3d2845b-49eb-4719-8f38-4e2abc62e967"
C = np.array([[-9, -48],
[-48, 0]])
sigma = np.linalg.eigvals(C)
sigma
# + [markdown] id="LqOUg4DQP0d5"
# $
# \\
# $
# >$
#
# \lambda =
# \begin{pmatrix}
# -52,7\\
# 43,7
# \end{pmatrix}
# \\
# Logo,\ a \ matriz\ H(x^*)\ não \ é \ definida \ neste \ ponto, \ o \ que \ implica \ em \ x^* =
# \begin{pmatrix}
# 2,0 \\
# 2,0
# \end{pmatrix}\ ser \ um \ ponto \ de
# \\
# sela.
#
# $
# + [markdown] id="VxcZQCGXt7er"
# $
# \\
# $
#
# >$
#
# No \ ponto \ ótimo\ x^* =
# \begin{pmatrix}
# 2,0 \\
# -2,0
# \end{pmatrix}:
# \\
# H(x^*) =
# \begin{pmatrix}
# -9 & 48 \\
# 48 & 0
# \end{pmatrix}
# \\
# Como\ a \ matriz\ H(x^*)\ não \ é \ diagonal \ neste \ ponto, \ temos \ que \ calcular \ seus \ autovalores \ (\lambda):
#
# $
# + colab={"base_uri": "https://localhost:8080/"} id="gEdHhiBqv1Mp" outputId="9b2b7215-684e-494c-fe1a-775971511a6e"
C = np.array([[-9, 48],
[48, 0]])
sigma = np.linalg.eigvals(C)
sigma
# + [markdown] id="lJYuMIInv7V4"
# $
# \\
# $
# >$
#
# \lambda =
# \begin{pmatrix}
# -52,7\\
# 43,7
# \end{pmatrix}
# \\
# Logo,\ a \ matriz\ H(x^*)\ não \ é \ definida \ neste \ ponto, \ o \ que \ implica \ em \ x^* =
# \begin{pmatrix}
# 2,0 \\
# -2,0
# \end{pmatrix}\ ser \ um \ ponto \ de
# \\
# sela.
#
# $
# + [markdown] id="SSsMVUZGwDcQ"
# $
# \\
# $
#
# >$
#
# No \ ponto \ ótimo\ x^* =
# \begin{pmatrix}
# -2,0 \\
# 1,0
# \end{pmatrix}:
# \\
# H(x^*) =
# \begin{pmatrix}
# 9 & 24 \\
# 24 & 0
# \end{pmatrix}
# \\
# Como\ a \ matriz\ H(x^*)\ não \ é \ diagonal \ neste \ ponto, \ temos \ que \ calcular \ seus \ autovalores \ (\lambda):
#
# $
# + colab={"base_uri": "https://localhost:8080/"} id="xofGjxurwYlQ" outputId="1f2f9032-c5f1-4e8d-ac95-d019cde8c109"
C = np.array([[9, 24],
[24, 0]])
sigma = np.linalg.eigvals(C)
sigma
# + [markdown] id="QJtXsXg2wmhB"
# $
# \\
# $
# >$
#
# \lambda =
# \begin{pmatrix}
# 28,9\\
# -19,,9
# \end{pmatrix}
# \\
# Logo,\ a \ matriz\ H(x^*)\ não \ é \ definida \ neste \ ponto, \ o \ que \ implica \ em \ x^* =
# \begin{pmatrix}
# -2,0 \\
# 1,0
# \end{pmatrix}\ ser \ um \ ponto \ de
# \\
# sela.
#
# $
# + [markdown] id="Vh2ELk6Rw8dr"
# $
# \\
# $
# ## ***Plotando a superfície e as curvas de níveis:***
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="m9j3siKKQw78" outputId="501ab8ae-8f47-4f00-a0e5-67ec01512f3a"
#Plot surface 3d:
from matplotlib import cm
x1 = np.linspace(-5., 5., 50)
x2 = np.linspace(-5., 5., 50)
X, Y = np.meshgrid(x1, x2)
Z = 7.5*X**2 + 12*Y**2 - 3*X**2*Y**2 + 18*X + 11
fig, ax = plt.subplots(subplot_kw={'projection': '3d'})
ax.plot_surface(X, Y, Z, cmap=cm.rainbow)
ax.set_xlabel('$x1$')
ax.set_ylabel('$x2$')
ax.set_zlabel('$L(x1,x2)$');
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="P-qVL-L5SVqW" outputId="c6c36c7c-f99f-4b34-9c0b-52d19185af23"
#Plot density - contour (with colorbar) - with stationary points:
plt.contourf(X, Y, Z, 50, cmap='RdGy')
plt.colorbar();
plt.scatter([-1.2, -2., 2., 2., -2.], [0, -1., 2., -2, 1.])
plt.annotate("(-1.2, 0)", (-1.2, 0))
plt.annotate("(-2, -1)", (-2., -1.))
plt.annotate("(2, 2)", (2., 2.))
plt.annotate("(2, -2)", (2., -2.))
plt.annotate("(-2, 1)", (-2., 1.))
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="L6x1LUo3TcPh" outputId="ca96ee99-ac62-4242-b54b-b0e50063b1bf"
#Calculando o valor de S(x1,x2) nos pontos estacionários:
Z = 7.5*X**2 + 12*Y**2 - 3*X**2*Y**2 + 18*X + 11
def f(x11, x22):
return 7.5*x11**2 + 12*x22**2 - 3*x11**2*x22**2 + 18*x11 + 11
result1 = f(-1.2, 0)
result2 = f(-2., -1.)
result3 = f(2., 2.)
print(result1, result2, result3)
# + colab={"base_uri": "https://localhost:8080/", "height": 314} id="QwdjBrMSTOyn" outputId="ab515301-b9e8-4513-f01a-ee24e16543c4"
#Plot density - contour (with labels):
Z = 7.5*X**2 + 12*Y**2 - 3*X**2*Y**2 + 18*X + 11
fig, ax = plt.subplots()
CS = ax.contour(X, Y, Z, [0.2,5.0,77.0], cmap='jet')
ax.clabel(CS, inline=1, fontsize=10)
ax.set_title('Countour with labels')
ax.set_xlabel('$x1$')
ax.set_ylabel('$x2$')
| Trabalhos/Trabalhos Grupo/lista1/1a_lista_torraca_gr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://seaborn.pydata.org/introduction.html
#
# * Seaborn is a library for making attractive and informative statistical graphics in Python. It is built on top of matplotlib and tightly integrated with the PyData stack, including support for numpy and pandas data structures and statistical routines from scipy and statsmodels.
#
# * Seaborn functions aim to make plots that are reasonably “production ready”
# +
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
import random
# %matplotlib inline
# +
df = pd.DataFrame()
df['x'] = random.sample(range(1, 100), 25)
df['y'] = random.sample(range(1, 100), 25)
print(df.info())
df.head()
# -
# ## Simple Plots:
#
# ## Scatter Plot
sns.lmplot('x', 'y', data=df, fit_reg=False)
# +
np.random.seed(19680801)
mu = 200
sigma = 25
n_bins = 50
x = np.random.normal(mu, sigma, size=100)
# -
# ## Plotting Distributions
sns.distplot(x)
# ## Nicer plots in seaborn:
#
# ## Scatter plots
# with sns.plotting_context(context='poster'):
with sns.plotting_context(context='notebook'):
pl = sns.lmplot(data=df, x='x', y='y', fit_reg=False, palette='subdued')
pl.fig.suptitle('Scatter Plot', fontsize=14, fontweight='bold')
pl.ax.set(xlabel='X axis data', ylabel='Y axis data');
# ## Plotting Distributions
# +
with sns.plotting_context(context='notebook'):
fig, ax = plt.subplots(figsize=(12, 8))
pl_dist = sns.distplot(x, bins=10, kde=False, rug=True, ax=ax)
sns.despine(fig=fig)
# -
with sns.plotting_context(context='notebook'):
fig, ax = plt.subplots(figsize=(12, 8))
pl_lm = sns.lmplot(data=df, x='x', y='y')
sns.despine(fig=fig)
print(type(pl_dist))
print(type(pl_lm))
# ## Data aware grids in Seaborn
#
# https://seaborn.pydata.org/tutorial/axis_grids.html
# +
tips = sns.load_dataset("tips")
print(tips.info())
tips.head()
# +
grid = sns.FacetGrid(tips, col="time", row='day', hue='smoker', sharey=True,
palette=dict(Yes="seagreen", No="gray"),
gridspec_kws={'width_ratios': [4, 2],
'height_ratios': [4, 2, 2, 2]})
#kwargs = {}
kwargs = {'alpha': .7, 'bins': 5}
grid.map(plt.hist, "total_bill", **kwargs)
grid.add_legend(title='FacetGrid Example')
# -
grid = sns.FacetGrid(tips, col='time', size=8)
grid.map(sns.distplot, "total_bill");
# The :class:`FacetGrid` is an object that links a Pandas DataFrame to
# a matplotlib figure with a particular structure.
assert isinstance(grid.fig, plt.Figure)
# https://stackoverflow.com/questions/48194193/seaborn-in-jupter-notebook-why-does-sns-despine-work-for-lmplot-but-not-regpl
# `ax` level functions and `fig` level functions.
# https://stackoverflow.com/questions/23969619/plotting-with-seaborn-using-the-matplotlib-object-oriented-interface
#
# ## PairGrid
#
# https://seaborn.pydata.org/tutorial/axis_grids.html
g = sns.PairGrid(tips)
g.map(plt.scatter);
g = sns.PairGrid(tips, hue='smoker')
g.map_diag(plt.hist)
g.map_offdiag(plt.scatter)
g.add_legend();
# ## Heatmap
sns.heatmap(df)
# ## Clustermap
sns.clustermap(df)
# ## Violin Plot
sns.violinplot(df)
| seaborn-plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis
# ---
# ### Research Questions:
# * What is the relationship between gender and disciplines and how does this relationship change over time?
# *
#
#
#
#
#
#
# ## RQ #1: The Proportion of Genders in the Olympics
# ---
# In my ELT I brushed over the relationship between gender and sport discipline and how this relationship changes over time. In this analysis I will be doing a deeper dive into the proportions of men and women in the Olympics thoughout the years, concluding by looking into where we have reached in our most recent data from 2008.
# Here we have the cleaned version of the data, I have removed a couple redundant variables to make it more concise. This is the dataframe that I will be transforming in order to further explore our data.
# + jupyter={"source_hidden": true}
import pandas as pd
import numpy as np
import seaborn as sns
from matplotlib import pyplot as plt
# + jupyter={"source_hidden": true}
df = pd.read_csv("../data/processed/data.csv")
df = df.copy().drop(['Country_Code', 'City', 'Athlete', 'Event_gender', 'Unnamed: 0'], axis=1)
df.head()
# + jupyter={"source_hidden": true}
df2 = df.groupby(["Year","Discipline"]).Gender.value_counts(normalize=True).mul(100).rename('Gender_pct').reset_index()
# + jupyter={"source_hidden": true}
gender_pct = pd.crosstab(df['Discipline'],df['Gender']).apply(lambda x: (x/x.sum())*100, axis=1)
gender_pct = gender_pct.sort_values(by=['Men'], ascending=True)
# + jupyter={"source_hidden": true}
gender_proportions2 = sns.lineplot(data=pd.crosstab(df['Year'],df['Gender']).apply(lambda x: (x/x.sum())*100, axis=1))
gender_proportions2.lines[1].set_linestyle("-")
gender_proportions2.legend().get_lines()[1].set_linestyle("-")
gender_proportions2.set_title('Gender Proportions Over Time')
gender_proportions2.axhline(50, ls=':', color='black')
# -
# Here is a plot of how the gender proportions have changed over time. You can see that the proportion of men is falling towards the 50% mark, while the proportion of women climbs up. Clearly, over time we are reaching an equilibrium, this is expecially clear if you observe around 2008, where the proportion is around 47% women. We are approaching an even split between the two. This chart shows valuable information, however, simply observing proportions could lead to a false understanding of the data. This chart could indicate a growth in female participation, but it could equally represent a fall in male participation. We are unable to assume from this graph that the number of women participants has increased, we must use the count of female and male athletes over time to draw any conclusions.
# + jupyter={"source_hidden": true}
gender_count = sns.lineplot(data=df.groupby(["Year", "Gender"])["Gender"].count().reset_index(name="Count"), x='Year', y='Count', hue='Gender')
gender_count.set_title('Number of Male/Female Participants over Time')
plt.savefig('gender_count.png', dpi=300)
# -
# This graph is similar to the other plot in that it shows how the gender balance has changed over time. However, this graph shows a count of male and female participants instead of a proportion. This gives another perspective on the situation, showing the growth of each gender instead of showing their growth in relation to one another. This graph displays a growth in both female and male participation, with the count of female athletes having a more significant change. This explains the change in proportions we observed in the previous graph. Let us now look at each individual discipline and how their gender proportions have changed over time.
# + jupyter={"source_hidden": true}
gender_proportions = gender_pct.plot(kind='barh', stacked='True', figsize=(8, 10), title='Gender Proportion per Discipline')
gender_proportions.axvline(50, ls=':', color='black')
plt.legend(loc="upper left")
# -
# This stacked barchart displays the proportions of men and women in each discipline, with a divider in the middle indicating the 50% mark. One of the main facts that is pulled from this visualization is that a majority of disciplines in the Olympics tend to lean towards having more male participants than female ones. However, there is also a significant group of disciplines with an equal split between genders. It is worth noting that this a cumulative representation of the data from 1976 to 2008, and not a representation of the current balance of men and women. We must dive into how this proportion has changed over time to have a better understanding of the progression of this data.
# + jupyter={"source_hidden": true}
genders2 = gender_pct.reset_index()
data2= df2[df2['Discipline'].isin(list(genders2.Discipline[10:30]))] ##taking the 20 middle disciplines from gender_proportions chart
data2 = data2[data2.Gender != 'Men'] ##dropping 'Men' to focus on data relating to 'Women'
# + jupyter={"source_hidden": true}
wmn_proportions = sns.lineplot(x='Year', y='Gender_pct', hue = 'Discipline', data=data2, ci=None)
wmn_proportions.set_title('Proportion of Women per Discipline Over Time')
plt.legend(loc="center right", bbox_to_anchor=(1.4, 0.5))
plt.savefig('wmn_proportions.png', dpi=300)
# -
# This last graph shows the how the proportion of women in each sport has changed over time. I decided to use the middle 20 disciplines from the Gender Proportions per Discipline chart because I felt that the disciplines closer to a 50% ratio would have the most interesting growth pattern in that they were less likely to be stagnant than the extremes (like wresling or synchronized swimming. You can see that for almost every discipline there is a steady growth towards the 50% mark. Some disciplines have more change than others, but it is clear that there is an overall shift towards equality over time.
# + jupyter={"source_hidden": true}
df2 = df2[df2.Year == 2008].drop(['Year'], axis = 1)
# + jupyter={"source_hidden": true}
df2 = pd.crosstab(df2['Discipline'],df2['Gender']).apply(lambda x: (x/x.sum())*100, axis=1)
df2 = df2.sort_values(by=['Men'], ascending=True)
# + jupyter={"source_hidden": true}
gender_proportions_2008 = df2.plot(kind='barh', stacked='True', figsize=(8, 10), title='Gender Proportion per Discipline (2008)')
gender_proportions_2008.axvline(50, ls=':', color='black')
plt.legend(loc="upper left")
# -
# Finally, here we have the gender proportions chart from 2008. In this chart it is clear to see that we have reached an equilibrium between the two genders for almost all disciplines. For each gender, there are three disciplines that exclusive to their gender. Looking back at the Gender Proportions per Discipline chart, it appears that these sports have been exclusive to each gender ever since they were originally introduced. You may be questioning why there was slight disproportion in 2008 shown in the first Gender Proportions Over Time graph. This can be explained by the difference between the count of male athletes that participated in their gender-exclusive disciplines compared to the count of women that competed in their exclusive sports. This is likely simply due to the number of events that are offered for each discipline. Without including the six gender-exclusive disciplines, a perfect equality has been reached.
# ### Conclusion
#
# We can conclude from all of the above visualizations that there certainly was a gender imbalance in the beginning years of the Olympics. However, as time has passed the ratio between male and female participants in the Olympics has leveled out to a near equilibrium.
#
| analysis/.ipynb_checkpoints/Analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/google/applied-machine-learning-intensive/blob/master/content/04_classification/08_video_processing_project/colab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="copyright"
# #### Copyright 2020 Google LLC.
# + colab={} colab_type="code" id="khlO4Bu21oZ4"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="AlzIlBsScJJ_"
# # Video Classification with Pre-Trained Models Project
# + [markdown] colab_type="text" id="nTirVS4FWaPx"
# In this project we will import a pre-existing model that recognizes objects and use the model to identify those objects in a video. We'll edit the video to draw boxes around the identified object, and then we'll reassemble the video so the boxes are shown around objects in the video.
# + [markdown] colab_type="text" id="YTVUYxPwcHhp"
# # Exercises
# + [markdown] colab_type="text" id="LdIOgOHP1ces"
# ## Exercise 1: Coding
# + [markdown] colab_type="text" id="jhTEOK1ZmqN8"
# You will process a video frame by frame, identify objects in each frame, and draw a bounding box with a label around each car in the video.
#
# Use the [SSD MobileNet V1 Coco](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/tf1_detection_zoo.md) (*ssd_mobilenet_v1_coco*) model. The video you'll process can be found [on Pixabay](https://pixabay.com/videos/cars-motorway-speed-motion-traffic-1900/). The 640x360 version of the video is smallest and easiest to handle, though any size should work since you must scale down the images for processing.
#
# Your program should:
#
# * Read in a video file (use the one in this colab if you want)
# * Load the TensorFlow model linked above
# * Loop over each frame of the video
# * Scale the frame down to a size the model expects
# * Feed the frame to the model
# * Loop over detections made by the model
# * If the detection score is above some threshold, draw a bounding box onto the frame and put a label in or near the box
# * Write the frame back to a new video
#
# Some tips:
#
# * Processing an entire video is slow, so consider truncating the video or skipping over frames during development. Skipping frames will make the video choppy. But you'll be able to see a wider variety of images than you would with a truncated video with all of the original frames in the clip.
# * The model expects a 300x300 image. You'll likely have to scale your frames to fit the model. When you get a bounding box, that box is relative to the scaled image. You'll need to scale the bounding box out to the original image size.
# * Don't start by trying to process the video. Instead, capture one frame and work with it until you are happy with your object detection, bounding boxes, and labels. Once you get those done, use the same logic on the other frames of the video.
# * The [Coco labels file](https://github.com/nightrome/cocostuff/blob/master/labels.txt) can be used to identify classified objects.
#
# + [markdown] colab_type="text" id="7XM35vYWSbim"
# ### **Student Solution**
# + colab={} colab_type="code" id="ivTzfzQN5jDk"
# Your code goes here
# + [markdown] colab_type="text" id="VEGDiC-IhcrM"
# ---
# + [markdown] colab_type="text" id="HniKdSXg0YHR"
# ## Exercise 2: Ethical Implications
# + [markdown] colab_type="text" id="W4FvC1Aa0ZT5"
# Even the most basic models have the potential to affect segments of the population in different ways. It is important to consider how your model might positively and negatively affect different types of users.
#
# In this section of the project, you will reflect on the positive and negative implications of your model. Frame the context of your model creation using this narrative:
#
# > The city of Seattle is attempting to reduce traffic congestion in its downtown area. As part of this project, they plan to allow each local driver one free trip to downtown Seattle per week. After that, the driver will have to pay a $50 toll for each extra day per week driven. As an early proof of concept for this project, your team is tasked with using machine learning to correctly identify automobiles on the road. The next phase of the project will involve detecting license plate numbers and then cross-referencing that data with RFID chips that should be mounted in all local drivers' cars.
# + [markdown] colab_type="text" id="lkyzwVQr0brd"
# ### **Student Solution**
# + [markdown] colab_type="text" id="gy4I2vG60ebd"
# **Positive Impact**
#
# Your model is trying to solve a problem. Think about who will benefit from that problem being solved and write a brief narrative about how the model will help.
# + [markdown] colab_type="text" id="k59MK1Ah0fWy"
# > *Hypothetical entities will benefit because...*
#
# + [markdown] colab_type="text" id="gzqkrLnk0hMU"
# **Negative Impact**
#
# Models rarely benefit everyone equally. Think about who might be negatively impacted by the predictions your model is making. This person(s) might not be directly using the model, but they might be impacted indirectly.
# + [markdown] colab_type="text" id="Hefa1JdP0kj3"
# > *Hypothetical entities will be negatively impacted because...*
# + [markdown] colab_type="text" id="Uax2HAzd0mHX"
# **Bias**
#
# Models can be biased for many reasons. The bias can come from the data used to build the model (e.g., sampling, data collection methods, available sources) and/or from the interpretation of the predictions generated by the model.
#
# Think of at least two ways bias might have been introduced to your model and explain both below.
# + [markdown] colab_type="text" id="6bJGm-qs0oQV"
# > *One source of bias in the model could be...*
#
# > *Another source of bias in the model could be...*
# + [markdown] colab_type="text" id="Ybb1zAkC0p2e"
# **Changing the Dataset to Mitigate Bias**
#
# Having bias in your dataset is one of the primary ways in which bias is introduced to a machine learning model. Look back at the input data you fed to your model. Think about how you might change something about the data to reduce bias in your model.
#
# What change or changes could you make to reduce the bias in your dataset? Consider the data you have, how and where it was collected, and what other sources of data might be used to reduce bias.
#
# Write a summary of changes that could be made to your input data.
# + [markdown] colab_type="text" id="UFsnF4_h08DD"
# > *Since the data has potential bias A we can adjust...*
# + [markdown] colab_type="text" id="ChEJbhXA02pW"
# **Changing the Model to Mitigate Bias**
#
# Is there any way to reduce bias by changing the model itself? This could include modifying algorithmic choices, tweaking hyperparameters, etc.
#
# Write a brief summary of changes you could make to help reduce bias in your model.
# + [markdown] colab_type="text" id="kEAhgO_U0p8Y"
# > *Since the model has potential bias A, we can adjust...*
# + [markdown] colab_type="text" id="rShB5BQv0wix"
# **Mitigating Bias Downstream**
#
# Models make predictions. Downstream processes make decisions. What processes and/or rules should be in place for people and systems interpreting and acting on the results of your model to reduce bias? Describe these rules and/or processes below.
# + [markdown] colab_type="text" id="C__BwBP-00HN"
# > *Since the predictions have potential bias A, we can adjust...*
# + [markdown] colab_type="text" id="1L_4RNXphYtI"
# ---
| content/04_classification/08_video_processing_project/colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="agK3X1at8tkL"
# ## Data Wrangling Step
#
# This step involves the cleaning, and formatting of data for the purpose of
# 1. Data Analysis.
# 2. Machine Learning Modelling.
# 3. Development of actionable insights.
#
#
# + [markdown] id="MGkHV45fehST"
# Data Should be downloaded and stored on the local machine or can be downloaded straight from the UCI repository depending on the preference.
#
# + id="63xMGYeusb4I"
BASE_FOLDER = "/content/drive/MyDrive/Colab_Notebooks/online_retail/"
# + id="rsCF3rE0xbpX"
# importing necessary libraries
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import plotly.graph_objects as go
import scipy as sp
import scipy.stats
# + id="N4lxDCUmxcm2"
# Import Retail data
retail_data = pd.read_csv("/content/drive/MyDrive/Colab_Notebooks/online_retail/online_retail_II.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="lFWsBMm9yYR2" outputId="cf066e0f-479b-45b7-cc4f-255ca730cb63"
retail_data.tail()
# + id="PQiCmSFFycQ1" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="2dc9ad90-1d7d-43bc-c278-e43526b9be68"
# To explore the data and give a descriptive statistics of the data
retail_data.describe()
# + [markdown] id="oBqh3w1agfhC"
# From the above descriptive statistics it can be observed that certain values are abnormal
#
# 1. For instance the Quantity and Price both have negative values.
#
# 2. The CustomerID does not uniquely represent all the transactions that were carried out.
#
# There are also other issues to be addressed.
#
# 3. Some descriptions are missing, this would also need to be sorted out as this will be useful for the analysis of the data.
# + [markdown] id="gi5vrwX4uU3f"
# # Data Cleaning tasks
# + colab={"base_uri": "https://localhost:8080/"} id="GrFPJLbCtIQN" outputId="05bcd73c-c26d-44da-f9d0-d1e20cffc2fb"
retail_data.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="bb3G4KtovxBa" outputId="8b8d630e-4986-4d3b-8d18-e480a213cf44"
print("Percentage of customers missing: ", round(retail_data['Customer ID'].isnull().sum() * 100 / len(retail_data),2),"%" )
# + [markdown] id="ygCgOQBiu7pj"
# From the look of things its the description and the customer ID that possess the missing data. This rows with this missing data makes up 22.77% of the data. It therefore important to devise a means to fill in this missing data intelligently.
# + [markdown] id="ZMF84e0h8H38"
# The average amount of orders made can be calculated using the invoice numbers and the total number of orders
# + colab={"base_uri": "https://localhost:8080/"} id="tXtH0C2G757b" outputId="5d2b181b-369c-4558-ada6-4b15b96b012f"
print("Number of orders in df1:", retail_data['Invoice'].nunique())
print("Avg. number of products per order in the dataset: ", round(retail_data['StockCode'].count() / retail_data['Invoice'].nunique(),2) )
# + [markdown] id="ZRKhGI9J8xrL"
# In the step to design or develop new customer IDs. This would ensure all customers can be uniquely identified.
# + id="4IEhxwIM8xQM"
retail_data["Null_CustomerID"] = [1 if i==True else 0 for i in retail_data["Customer ID"].isnull()]
# + [markdown] id="3FvnlR3P-xZG"
# It is safe to assume that creating new customer IDs will not affect the insights we hope to gain from the data. Reason being that the invoice number can be used to identify all unique orders, which is our purpose of analysis for now.
#
# New customer IDs can be assigned by using the invoice orders, however this would mean that customers for which their customer IDs were not captured are automatically seen as new and unique customers.
# + id="8LyNwMKA9XP7"
retail_data["New_ID"] = retail_data["Customer ID"]
retail_data.loc[retail_data["Null_CustomerID"] == 1, ["New_ID"]] = retail_data["Invoice"]
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="tLCrt2hEDvoo" outputId="aa47e6b9-f4f0-463f-a824-9928bff6a256"
retail_data[retail_data["Null_CustomerID"] == 1].head()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="jJgVz-BTD02H" outputId="c10796b7-edd2-4e7b-a375-4a9234c09c52"
retail_data[retail_data["Null_CustomerID"] == 0].head()
# + colab={"base_uri": "https://localhost:8080/"} id="TiVlWJNcECKT" outputId="b987f035-ea37-4c6c-c7c8-8036ce64d500"
retail_data.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="_tRfNRUkEJ-i" outputId="d66b8b75-e404-4676-8d77-25ccba43c855"
#drop Customer ID column since it isn't need anymore
retail_data = retail_data.drop('Customer ID', 1)
retail_data.head()
# + [markdown] id="5vdVsgL2R1i9"
# To ensure consistency the data types of the new ID is meant to be numeric, since cancelled orders were inclusive in the invoice column some of the rows must have affected the data type of the New ID,
#
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="ZzWGUpT5FLUb" outputId="d8e11a35-ac63-4945-e67d-28cafa88806d"
#Remove all non digits from column NewID
retail_data['New_ID'] = retail_data['New_ID'].astype(str).str.replace('\D+', '')
#Convert to integer
retail_data['New_ID'] = pd.to_numeric(retail_data['New_ID'])
#Check if the 'C' character was removed in the NewID column
retail_data[retail_data['New_ID']== 580605]
# + colab={"base_uri": "https://localhost:8080/"} id="jAdcIAI1TKVd" outputId="529f21ff-84ec-43d6-ef99-f92533a76d0c"
retail_data.info()
# + id="MzE4gCCrp--o" colab={"base_uri": "https://localhost:8080/"} outputId="66ad73f1-7a1f-4aba-d330-571a9586a00b"
retail_data.isnull().sum()
# + [markdown] id="ZNZKTUn6tcD9"
# From the above result it is evident that all the transactions now possess unique customer IDs.
#
# To aid the analysis of this data, other columns will also need be addressed. The columns would be addressed in this order:
#
# 1. Description column:
# * Ensure all items possess valid descriptions.
# * Ensure all transactions posses descriptions.
#
# 2. Invoice Date
# * Ensure Proper date-time formatting
#
# 3. StockCode
# * Ensure proper StockCode assignments.
# * Removing irrelevant StockCodes.
#
# 4. Country
# * Ensure unique values are assigned in the Country column.
#
# 5. Price
# * Remove outliers
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="sB9bkrSsxeXP" outputId="5b509308-c4a6-4669-cab5-cd74d2daee36"
# Dealing with the Description columns
# String formatting
#Strings
retail_data['Description'] = retail_data['Description'].str.replace('.','').str.upper().str.strip()
retail_data['Description'] = retail_data['Description'].replace('\s+',' ',regex = True)
# Assigning correct and missing descriptions by using the StockCode
for index,value in retail_data.StockCode[retail_data.Description.isna()==True].items():
if pd.notna(retail_data.Description[retail_data.StockCode == value]).sum() != 0:
retail_data.Description[index] = retail_data.Description[retail_data.StockCode == value].mode()[0]
else:
retail_data.drop(index = index, inplace = True)
retail_data['Description'] = retail_data['Description'].astype(str)
# + id="xmu5Ir0x1NtV"
# Dealing with the StockCode column
# String formatting
retail_data['StockCode'] = retail_data['StockCode'].str.upper()
# Dropping all stockcodes that contain only strings
code_types = list(map(lambda codes: any(char.isdigit() for char in codes), retail_data['StockCode']))
irrelevant_codes = [i for i,v in enumerate(code_types) if v == False]
retail_data.drop(irrelevant_codes , inplace = True)
# + colab={"base_uri": "https://localhost:8080/"} id="n-bBY_IiCpwz" outputId="f8c2837a-4294-40c9-d0ef-13f660a3e436"
retail_data.columns
# + colab={"base_uri": "https://localhost:8080/"} id="Fv3462ek_XAy" outputId="5f930d76-6161-4e82-a40a-d07518763ae3"
# Removing Outliers Based on Z-score
retail_data = retail_data[(np.abs(sp.stats.zscore(retail_data['Price']))<3) & (np.abs(sp.stats.zscore(retail_data['Quantity']))<5)]
# Dealing with missing & incorrect values in Price and Quantity columns
retail_data.drop(retail_data[(retail_data.Quantity>0) & (retail_data.Invoice.str.contains('C') == True)].index, inplace = True)
retail_data.drop(retail_data[(retail_data.Quantity<0) & (retail_data.Invoice.str.contains('C') == False)].index, inplace = True)
retail_data.drop(retail_data[retail_data.Description.str.contains('?',regex=False) == True].index, inplace = True)
retail_data.drop(retail_data[retail_data.Price == 0].index, inplace = True)
# + id="ExOroX_Zt5jY" colab={"base_uri": "https://localhost:8080/"} outputId="b4349b93-fcb3-4a2d-d508-8761b1ab593c"
# Dealing with the country column
# Customers with Different Countries
Customers = retail_data.groupby('New_ID')['Country'].unique()
Customers.loc[Customers.apply(lambda x:len(x)>1)]
# + id="MnVB1fUQB57q"
# Fixing duplicate CustomerIDs and Multiple Countries
for i,v in retail_data.groupby('New_ID')['Country'].unique().items():
if len(v)>1:
retail_data.Country[retail_data['New_ID'] == i] = retail_data.Country[retail_data['New_ID'] == i].mode()[0]
# + id="xFuGX353DBL1"
# Adding Desired Features
retail_data['Amount_Spent'] = retail_data['Quantity']*retail_data['Price'] # Total
# Formatting Date/Time
retail_data['InvoiceDate'] = pd.to_datetime(retail_data['InvoiceDate'], format = '%Y/%m/%d %H:%M')
retail_data['InvoiceMonth'] = retail_data['InvoiceDate'].apply(lambda x: x.strftime('%B'))
retail_data['Day of week'] = retail_data['InvoiceDate'].dt.day_name()
# + id="gMPquprQE8Iy"
# Exporting Processed Data
retail_data.to_csv(f'{BASE_FOLDER}cleaned_retail_data.csv', date_format = '%Y-%m-%d %H:%M', index = False)
| online_retail_data_wrangling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
from scipy import interpolate
from PIL import Image
import matplotlib.pyplot as plt
from astropy.visualization import (MinMaxInterval, SqrtStretch,
ImageNormalize, ZScaleInterval)
# %matplotlib notebook
# Resampling factor
factor = 4
# Open **png** image using PIL
img = Image.open('m31.png', 'r')
img.size
# Crop the image to a size based on the resampling factor
aux_dim = [(n // factor) * factor for n in img.size]
# For PIL the coordinate system starts at the left-top position
left, top, right, bottom = 0, 0, aux_dim[0], aux_dim[1]
img = img.crop(box=(left, top, right, bottom))
# +
fig, ax = plt.subplots(1, 4, figsize=(8, 3))
cmaps = ['Reds', 'Greens', 'Blues']
for i, axis in enumerate(ax.flatten()):
if i == 0:
axis.imshow(np.asarray(img))
else:
axis.imshow(np.asarray(img.split()[i - 1]), cmap=cmaps[i - 1])
axis.axis('off')
plt.subplots_adjust(left=0, bottom=0, top=1, right=1, wspace=0)
plt.savefig('resized_bands.png', dpi=300, format='png')
# -
# Now, do the subsampling to a lower resolution
dim_bin = [n // factor for n in aux_dim]
img_bin = img.resize(dim_bin, resample=Image.LANCZOS)
# Plotting the subsampled image
# +
fig, ax = plt.subplots(1, 4, figsize=(8, 3))
cmaps = ['Reds', 'Greens', 'Blues']
for i, axis in enumerate(ax.flatten()):
if i == 0:
axis.imshow(np.asarray(img_bin))
else:
axis.imshow(np.asarray(img_bin.split()[i - 1]), cmap=cmaps[i - 1])
axis.axis('off')
plt.subplots_adjust(left=0, bottom=0, top=1, right=1, wspace=0)
plt.savefig('subsampled_bands.png', dpi=300, format='png')
# -
# Define a finer grid, increasing the number of points using the sampling factor
N_dim = [factor * k for k in dim_bin]
xx, yy = np.meshgrid(np.linspace(0, dim_bin[1], N_dim[1]),
np.linspace(0, dim_bin[0], N_dim[0]),
sparse=False, indexing='ij')
# We need to also define the subsampled grid
xx_ini, yy_ini = np.mgrid[0:dim_bin[1], 0:dim_bin[0]]
points = np.vstack([yy_ini.ravel(), xx_ini.ravel()])
xx.shape, xx_ini.shape
# At this point we need to work separately with the red, green, and blue bands
values_r = np.asarray(img_bin.split()[0]).flatten()
values_g = np.asarray(img_bin.split()[1]).flatten()
values_b = np.asarray(img_bin.split()[2]).flatten()
values_r.shape, points.shape
# We then interpolate the subsampled grid into a finer, using the values pixel values for interpolation
interp_img = [interpolate.griddata(points.T, v, (yy, xx), method='cubic')
for v in [values_r, values_g, values_b]]
# Normalize byt the number of subpixels each pixel is divided
# interp_img /= np.power(factor, 2)
arr_rec = np.dstack([*interp_img])
arr_rec /= 255.
# +
arr_rec.shape
fig, ax = plt.subplots(1, 2, figsize=(8, 4))
ax[0].imshow(img)
ax[1].imshow(arr_rec)
ax[0].set_title('high res')
ax[1].set_title('reconstructed')
ax[0].axis('off')
ax[1].axis('off')
plt.tight_layout()
plt.savefig('reconstructed.png', dpi=300, format='png')
# +
fig, ax = plt.subplots(1, 3, figsize=(8, 4))
cmaps = ['Reds', 'Greens', 'Blues']
for j, axis in enumerate(ax.flatten()):
diff_img = np.asarray(img.split()[j]) - interp_img[j]
im_norm = ImageNormalize(diff_img,
interval=ZScaleInterval(),
stretch=SqrtStretch())
tmp = axis.imshow(diff_img, norm=im_norm, cmap=cmaps[j])
plt.colorbar(tmp, ax=axis, orientation='horizontal')
axis.axis('off')
plt.subplots_adjust(top=0.99, left=0.02, right=0.98, bottom=0)
plt.suptitle('high resolution minus reconstructed')
plt.savefig('difference_bands.png', dpi=300, format='png')
# -
| code/p03_grid_sampling/grid_resampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from keras.utils import np_utils
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Conv2D
from keras.layers import MaxPooling2D
from keras.layers import Dropout
from keras.layers import Flatten
import matplotlib.pyplot as plt
import time
from keras.datasets import cifar10
# +
#download the data
(trainX, trainy), (testX, testy) = cifar10.load_data()
#X_train and y_train will be used for training
X_train80, X_train, y_train80, y_train = train_test_split(trainX, trainy, test_size=0.2, random_state=42)
#For test and validation set
X_test, X_val, y_test, y_val = train_test_split(testX, testy, test_size=0.5, random_state=42)
# +
#plot the accuracy and loss
def train_val_plt(history, key1 , key2, name) :
plt.plot(history.history[key1])
plt.plot(history.history[key2])
plt.title(name)
plt.xlabel('epochs')
plt.ylabel('accuarcy')
X_range = np.arange(0, 5, 1)
plt.xticks(X_range)
plt.legend( ['train', 'validation'], loc = 'upper left')
plt.show()
def loss_plt(history, key1 , key2, name) :
plt.plot(history.history[key1])
plt.plot(history.history[key2])
plt.title(name)
plt.xlabel('epochs')
plt.ylabel('loss')
X_range = np.arange(0, 5, 1)
plt.xticks(X_range)
plt.legend( ['train', 'validation'], loc = 'upper left')
plt.show()
# -
# # Data Preprocessing
# +
# Data Preprocessing for MLP
X_train_normalization = X_train.reshape( len(X_train), 32*32*3 ).astype('float32')
X_train_normalization /= 255
X_train_normalization.shape
X_val_normalization = X_val.reshape( len(X_val), 32*32*3 ).astype('float32')
X_val_normalization /= 255
X_test_normalization = X_test.reshape( len(X_test), 32*32*3 ).astype('float32')
X_test_normalization /= 255
#one hot encoding for label
y_train_onehot = np_utils.to_categorical(y_train)
y_val_onehot = np_utils.to_categorical(y_val)
y_test_onehot = np_utils.to_categorical(y_test)
#DATA PROCESSING FOR CNN
train_CNN, val_CNN, test_CNN = X_train / 255.0, X_val / 255.0, X_test / 255.0
# -
# # MLP
#MLP
MLP = Sequential()
MLP.add(Dense(input_dim = 32*32*3, units = 512, activation = 'sigmoid'))
MLP.add(Dense(units = 512, activation = 'sigmoid'))
#output layer
MLP.add(Dense(units = 10, activation = 'softmax'))
MLP.summary()
#compile the model
MLP.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
#fit the model
history = MLP.fit(x = X_train_normalization,
y = y_train_onehot,
validation_data = (X_val_normalization, y_val_onehot),
epochs=5,
batch_size=32,
verbose=2)
train_val_plt( history, 'accuracy', 'val_accuracy','MLP acc')
loss_plt( history, 'loss', 'val_loss','MLP loss')
# +
# change MLP's the number of layer and the number of neurons as required in this problem
# #MLP
# MLP1 = Sequential()
# MLP1.add(Dense(input_dim = 32*32*3, units = 512, activation = 'sigmoid'))
# MLP1.add(Dense(units = 512, activation = 'sigmoid'))
# MLP1.add(Dense(units = 512, activation = 'sigmoid'))
# #output layer
# MLP1.add(Dense(units = 10, activation = 'softmax'))
# MLP1.summary()
# MLP1.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# history1 = MLP1.fit(x = X_train_normalization,
# y = y_train_onehot,
# validation_data = (X_val_normalization, y_val_onehot),
# epochs=5,
# batch_size=32,
# verbose=2)
# train_val_plt( history1, 'accuracy', 'val_accuracy','MLP acc')
#==============================================
# #MLP
# MLP2 = Sequential()
# MLP2.add(Dense(input_dim = 32*32*3, units = 512, activation = 'sigmoid'))
# MLP2.add(Dense(units = 256, activation = 'sigmoid'))
# #output layer
# MLP2.add(Dense(units = 10, activation = 'softmax'))
# MLP2.summary()
# MLP2.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# history2 = MLP2.fit(x = X_train_normalization,
# y = y_train_onehot,
# validation_data = (X_val_normalization, y_val_onehot),
# epochs=5,
# batch_size=32,
# verbose=2)
# train_val_plt( history2, 'accuracy', 'val_accuracy','MLP acc')
#==============================================
# #MLP
# MLP3 = Sequential()
# MLP3.add(Dense(input_dim = 32*32*3, units = 512, activation = 'sigmoid'))
# MLP3.add(Dense(units = 512, activation = 'sigmoid'))
# MLP3.add(Dense(units = 256, activation = 'sigmoid'))
# #output layer
# MLP3.add(Dense(units = 10, activation = 'softmax'))
# MLP3.summary()
# MLP3.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# history3 = MLP3.fit(x = X_train_normalization,
# y = y_train_onehot,
# validation_data = (X_val_normalization, y_val_onehot),
# epochs=5,
# batch_size=32,
# verbose=2)
# train_val_plt( history3, 'accuracy', 'val_accuracy','MLP acc')
#==============================================
# #MLP
# MLP4 = Sequential()
# MLP4.add(Dense(input_dim = 32*32*3, units = 512, activation = 'sigmoid'))
# MLP4.add(Dense(units = 512, activation = 'sigmoid'))
# MLP4.add(Dense(units = 256, activation = 'sigmoid'))
# MLP4.add(Dense(units = 256, activation = 'sigmoid'))
# #output layer
# MLP4.add(Dense(units = 10, activation = 'softmax'))
# MLP4.summary()
# MLP4.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# history4 = MLP4.fit(x = X_train_normalization,
# y = y_train_onehot,
# validation_data = (X_val_normalization, y_val_onehot),
# epochs=5,
# batch_size=32,
# verbose=2)
# train_val_plt( history4, 'accuracy', 'val_accuracy','MLP acc')
#==============================================
# #MLP
# MLP5 = Sequential()
# MLP5.add(Dense(input_dim = 32*32*3, units = 512, activation = 'sigmoid'))
# MLP5.add(Dense(units = 512, activation = 'sigmoid'))
# MLP5.add(Dense(units = 256, activation = 'sigmoid'))
# MLP5.add(Dense(units = 128, activation = 'sigmoid'))
# #output layer
# MLP5.add(Dense(units = 10, activation = 'softmax'))
# MLP5.summary()
# MLP5.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# history5 = MLP5.fit(x = X_train_normalization,
# y = y_train_onehot,
# validation_data = (X_val_normalization, y_val_onehot),
# epochs=5,
# batch_size=32,
# verbose=2)
# train_val_plt( history5, 'accuracy', 'val_accuracy','MLP acc')
# -
#evaluate the model
eval_MLP = MLP.evaluate(X_test_normalization, y_test_onehot)
print('Test_loss =', eval_MLP[0])
print('Test_accuracy =', eval_MLP[1])
# # CNN1
#CNN1
CNN1 = Sequential()
CNN1.add(Conv2D(64, (3, 3), padding = 'same', activation='relu', input_shape=(32, 32, 3)))
CNN1.add(Conv2D(64, (3, 3), padding = 'same', activation='relu'))
CNN1.add(Flatten())
CNN1.add(Dense(512, activation='sigmoid'))
CNN1.add(Dense(512, activation='sigmoid'))
CNN1.add(Dense(10,activation = 'softmax'))
CNN1.summary()
#compile the model
CNN1.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
#fit the model
start_time = time.time()
history_CNN1 = CNN1.fit(x = train_CNN,
y = y_train_onehot,
validation_data = (val_CNN, y_val_onehot),
epochs=5,
batch_size=32)
end_time = time.time()
print('Training_time: ' + str(end_time - start_time))
train_val_plt( history_CNN1, 'accuracy', 'val_accuracy','CNN1 acc')
loss_plt( history_CNN1, 'loss', 'val_loss','CNN1 loss')
#evaluate the model
eval_CNN1 = CNN1.evaluate(test_CNN, y_test_onehot)
print('Test_loss =', eval_CNN1[0])
print('Test_accuracy =', eval_CNN1[1])
# # CNN2
#CNN2
CNN2 = Sequential()
CNN2.add(Conv2D(64, (3, 3), padding = 'same', activation='relu', input_shape=(32, 32, 3)))
CNN2.add(MaxPooling2D((2, 2)))
CNN2.add(Conv2D(64, (3, 3), padding = 'same', activation='relu', input_shape=(32, 32, 3)))
CNN2.add(MaxPooling2D((2, 2)))
CNN2.add(Flatten())
CNN2.add(Dense(512, activation='sigmoid'))
CNN2.add(Dropout(0.2))
CNN2.add(Dense(512, activation='sigmoid'))
CNN2.add(Dropout(0.2))
CNN2.add(Dense(10,activation = 'softmax'))
CNN2.summary()
#compile the model
CNN2.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
#fit the model
start_time = time.time()
history_CNN2 = CNN2.fit(x = train_CNN,
y = y_train_onehot,
validation_data = (val_CNN, y_val_onehot),
epochs=5,
batch_size=32)
end_time = time.time()
print('Training_time: ' + str(end_time - start_time))
train_val_plt(history_CNN2, 'accuracy', 'val_accuracy','CNN2 acc')
loss_plt( history_CNN2, 'loss', 'val_loss','CNN2 loss')
#evaluate the model
eval_CNN2 = CNN2.evaluate(test_CNN, y_test_onehot)
print( 'Test_loss =', eval_CNN2[0] )
print( 'Test_accuracy =', eval_CNN2[1] )
| Problem1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/emenriquez/python-challenge-exercises/blob/master/Highest_Rank_Number_in_Array_(6).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8znr21lIZbRp" colab_type="text"
# # Highest Rank Number in Array
#
# Write a method `highestRank(arr)` (or `highest-rank` in clojure) which returns the number which is most frequent in the given input array (or `ISeq`). If there is a tie for most frequent number, return the largest number of which is most frequent.
#
# Example:
# ```
# arr = [12, 10, 8, 12, 7, 6, 4, 10, 12];
# highestRank(arr) //=> returns 12
#
# arr = [12, 10, 8, 12, 7, 6, 4, 10, 12, 10];
# highestRank(arr) //=> returns 12
#
# arr = [12, 10, 8, 8, 3, 3, 3, 3, 2, 4, 10, 12, 10];
# highestRank(arr) //=> returns 3
# ```
# + id="iibK-BEaZWA_" colab_type="code" colab={}
def highest_rank(arr):
# sort the set of values found in the array first by their frequency, then by their descending values
# return the highest ranking value sorted by these conditions
highestRank = max(set(arr), key= lambda x: (arr.count(x), x))
return highestRank
# + [markdown] id="SO1xCv-qfxZp" colab_type="text"
# ### Test Example
# + id="GHiYV3ZbaDQ-" colab_type="code" colab={}
example = [12, 10, 8, 12, 7, 6, 4, 10, 12, 10]
# + id="4tyY6SXxbFqx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8d48a67e-9f7d-4fd7-939e-24d26cfd434a"
highest_rank(example)
# + [markdown] id="XPb2WfQ2f8HV" colab_type="text"
# Completed in 11 Minutes
| Highest_Rank_Number_in_Array_(6).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Approximating square root iteratively
#
# There are a lot of useful functions that have no closed form solution, meaning we can't just do a computation and return the value. Instead, we need to use an iterative method to approximate the function value. It's an extremely useful pattern that will come in handy later in this program and in your professional life. We can use iterative computing to approximate sine (with Taylor series expansion), approximate square root (as we'll do in this lecture), or optimize a cost or error function (gradient descent in the introduction to machine learning course).
#
# The goal of this lab is to translate a recurrence relation, that yields a sequence of values, to Python. Instead of computing and returning a single value, we will look for **convergence of the series**. In other words, if we run the series out far enough, $x_{i+1}$ will be close to $x_i$ leaving $x_i$ as a very accurate approximation of square root. This will teach us the basics of iterative computing and prepare us for more complicated function optimization material needed later.
# ## Babylonian method
#
# To approximate square root, the idea is to pick an initial estimate, $x_0$, and then iterate with better and better estimates, $x_i$, using the ([Babylonian method](https://en.wikipedia.org/wiki/Methods_of_computing_square_roots#Babylonian_method)) recurrence relation:
#
# $x_{i+1} = \frac{1}{2}(x_i + \frac{n}{x_i})$
#
# There’s a great deal on the web you can read to learn more about why this process works but it relies on the average (midpoint) of $x_i$ and $n/x_i$ getting us closer to the square root of n. The cool thing is that the iteration converges quickly.
#
# Our goal is to write a function that takes a single number and returns its square root. What do we know about this function before even beginning to code? Well, we have a clear description of the problem per the recurrence relation, and we also have the function signature we want:
#
# ```python
# def sqrt(n):
# ```
#
# Because we are implementing a recurrence relation, we know that we will have a loop that computes $x_{i+1}$ from $x_{i}$.
# ### Convergence
#
# The terminating condition of the loop is when we have reached convergence or close to it. Convergence just means that $x_{i+1}$ is pretty close to $x_i$. Because we can never compare to real numbers for equality, we have to check for the difference being smaller than some precision like 0.00000001.
# ### Iterative method outline
#
# Just as we have an outline for a data science program, iterative methods all share the same basic outline. (I'm assuming here that $x_{i+1}$ depends only on a single previous value and that $i$ implicitly increments as the loop goes around.)
#
# *set $x_0$ to initial value*<br>
# *repeat:*<br>
# $x_{i+1} =$ function-giving-next-value$(x_i)$<br>
# *until $abs(x_{i+1} - x_i) \lt precision$<br>
# return $x_{i+1}$*<br>
#
# Because Python does not have a repeat-until loop, we fake it with an infinite loop containing a conditional that breaks us out upon convergence:
#
# *set $x_0$ to initial value*<br>
# *while True:*<br>
# $x_{i+1} =$ function-giving-next-value$(x_i)$<br>
# *if $abs(x_{i+1} - x_i) \lt precision$<br>
# return $x_{i+1}$*<br>
#
# That is a fairly straightforward implementation of the recurrence relation, but you will notice that we don't actually need to keep all previous $x_i$ around except for the new value and the previous value. Here is a Python implementation that tracks only two values and follows the infinite loop pattern:
def sqrt(n):
"compute square root of n"
PRECISION = 0.00000001 # stop iterating when we converge with this delta
x_0 = 1.0 # pick any old initial value
x_prev = x_0
while True: # Python doesn't have repeat-until loop so fake it
#print(x_prev)
x_new = 0.5 * (x_prev + n/x_prev)
if abs(x_new - x_prev) < PRECISION:
return x_new
x_prev = x_new # x_i+1 becomes x_i (previous value)
sqrt(100)
# To test our square root approximation, we can compare it to `math.sqrt()` and use numpy's `isclose` to do the comparison.
# +
import numpy as np
def check(n):
assert np.isclose(sqrt(n), np.sqrt(n))
def test_big():
check(125348)
def test_medium():
check(89.2342)
def test_100():
check(100)
def test_1():
check(1)
def test_0():
check(0)
test_big()
test_medium()
test_100()
test_1()
test_0()
# -
# ### Exercise
#
# Type in (don't cut/paste) the `sqrt(n)` function and test with, for example, `sqrt(125348.0)`. Make sure you get the right answer (354.045195) and then add print statements so that you can see the sequence of $x_{i}$ values. I get:
#
# ```
# 1.0
# 62674.5
# 31338.249992
# 15671.1249162
# 7839.56178812
# 3927.77547356
# 1979.84435152
# 1021.5781996
# 572.139273508
# 395.612894667
# 356.228988269
# 354.051888518
# 354.045194918
# 354.045194855
# ```
#
# Notice how quickly it converges!
# +
def sqrt_with_trace(n):
"compute square root of n"
PRECISION = 0.00000001 # stop iterating when we converge with this delta
x_0 = 1.0 # pick any old initial value
x_prev = x_0
while True: # Python doesn't have repeat-until loop so fake it
print(x_prev)
x_new = 0.5 * (x_prev + n/x_prev)
if abs(x_new - x_prev) < PRECISION:
return x_new
x_prev = x_new
sqrt_with_trace(125348.000000)
# -
# ### Exercise: Testing with pytest
# To run with the unit tester pytest, save the `sqrt()` function in a file called `sqrt.py` and then put the test code into a file called `test_sqrt.py`. That test file will also need to import the square root code so add
#
# ```
# from sqrt import *
# ```
#
# to the top of the test file. To run the tests from the command line, you can do the following:
#
# ```bash
# $ python test_sqrt.py
# $
# ```
#
# which prints nothing because there are no errors (hopefully).
#
# A better way is to use `pytest` which executes all functions that start with prefix `test_`:
#
# ```bash
# $ pip install pytest
# $ pytest -v test_sqrt.py
# ============================= test session starts ==============================
# platform darwin -- Python 3.8.8, pytest-6.2.3, py-1.10.0, pluggy-0.13.1 -- /Users/parrt/opt/anaconda3/bin/python
# cachedir: .pytest_cache
# rootdir: /private/tmp
# plugins: anyio-2.2.0
# collected 5 items
#
# test_sqrt.py::test_big PASSED [ 20%]
# test_sqrt.py::test_medium PASSED [ 40%]
# test_sqrt.py::test_100 PASSED [ 60%]
# test_sqrt.py::test_1 PASSED [ 80%]
# test_sqrt.py::test_0 PASSED [100%]
#
# ============================== 5 passed in 0.07s ===============================
# ```
#
# The other benefit is that we don't need the explicit calls so you can delete this part and it still works with pytest:
#
# ```python
# test_big()
# test_medium()
# test_100()
# test_1()
# test_0()
# ```
| labs/sqrt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="HlKnRpPMilEi" executionInfo={"status": "ok", "timestamp": 1618541498331, "user_tz": -480, "elapsed": 3905, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-BV-vQrnIvTY/AAAAAAAAAAI/AAAAAAAADxw/wSzRzhF8b6c/s64/photo.jpg", "userId": "01517089400226719298"}}
import os, requests
import numpy as np
from PIL import Image
import tensorflow as tf
from keras.preprocessing import image
# + colab={"base_uri": "https://localhost:8080/"} id="LR9au2crXZR-" executionInfo={"status": "ok", "timestamp": 1618541507035, "user_tz": -480, "elapsed": 12586, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-BV-vQrnIvTY/AAAAAAAAAAI/AAAAAAAADxw/wSzRzhF8b6c/s64/photo.jpg", "userId": "01517089400226719298"}} outputId="a3009bb0-5a48-4d67-fa8d-54e88eb5d5f5"
# !gdown --id 1sFV6E6PwX-ZNlDhdCvFA6TstK5wVqafE
# !unzip model_landmarks.zip
# + id="UyZS2saLin9w" executionInfo={"status": "ok", "timestamp": 1618541522520, "user_tz": -480, "elapsed": 28069, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-BV-vQrnIvTY/AAAAAAAAAAI/AAAAAAAADxw/wSzRzhF8b6c/s64/photo.jpg", "userId": "01517089400226719298"}}
model = tf.keras.models.load_model("inception_landmarks")
# + id="WX1OTM0je7AO" executionInfo={"status": "ok", "timestamp": 1618541522522, "user_tz": -480, "elapsed": 28069, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-BV-vQrnIvTY/AAAAAAAAAAI/AAAAAAAADxw/wSzRzhF8b6c/s64/photo.jpg", "userId": "01517089400226719298"}}
labels = ['Bugis', 'Chinatown', 'Clarke Quay', 'Esplanade', 'Flowerdome', 'Flyer', 'Hajilane', 'Marina Bay Sands', 'Merlion', 'Supertree', 'USS']
IMAGE_SIZE = (256, 256)
# + id="-mP44r-BiuSy" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1618405179582, "user_tz": -480, "elapsed": 1701, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-BV-vQrnIvTY/AAAAAAAAAAI/AAAAAAAADxw/wSzRzhF8b6c/s64/photo.jpg", "userId": "01517089400226719298"}} outputId="3963b807-29b5-4677-e550-09ae5da40580"
image_url = "https://www.rwsentosa.com/-/media/project/non-gaming/rwsentosa/attractions/universal-studios-singapore/others/uss-entrance-globe_1366x666.jpg" #@param {type:"string"}
try:
image_data = requests.get(image_url, stream=True).raw
except Exception as e:
print('Warning: Could not download image from %s' % image_url)
print('Error: %s' %e)
raise
try:
pil_image = Image.open(image_data)
except Exception as e:
print('Warning: Failed to parse image')
print('Error: %s' %e)
raise
try:
img = pil_image.convert('RGB').resize((IMAGE_SIZE))
except:
print('Warning: Failed to format image')
raise
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
classes = model.predict(x)
output = []
for i in range(len(classes[0])):
output.append((labels[i], classes[0][i]))
output.sort(key=lambda x:-x[1])
for i in range(len(output)):
print("%s: %s" % (output[i][0], output[i][1]))
# + id="ldwwZRzXalZK"
| Final_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SQLite
# Beispiel mit SQLite, ohne Server
# %load_ext sql
# %sql sqlite://
# + language="sql"
# CREATE TABLE writer (first_name, last_name, year_of_death);
# INSERT INTO writer VALUES ('William', 'Shakespeare', 1616);
# INSERT INTO writer VALUES ('Bertold', 'Brecht', 1956);
# -
# %sql select * from writer
# #Mit MySQL
# Voraussetzung, der MySQL Server läuft!
# %sql mysql://tester:test@localhost/test_db
# + language="sql"
# CREATE TABLE `test_db`.`writer` (`first_name` VARCHAR(30) NOT NULL,`last_name` VARCHAR(45) NULL,
# `year_of_death` DATETIME NULL, PRIMARY KEY (`first_name`));
# + language="sql"
# INSERT INTO writer VALUES ('William', 'Shakespeare', '1616-01-01 00:00');
# INSERT INTO writer VALUES ('Bertold', 'Brecht', '1956-01-01 00:00');
# -
# %sql select * from `test_db`.`writer`;
# %sql DROP TABLE `test_db`.`writer`
# #MongoDB
# Verbindung herstellen
# +
import pymongo
# Connection to Mongo DB
try:
conn=pymongo.MongoClient()
print ("Connected successfully!!!")
except (pymongo.errors.ConnectionFailure, e):
print ("Could not connect to MongoDB: %s" % e )
conn
# -
# Nun mal etwas rumprobieren:
db = conn.mydb
collection = db.my_collection
doc1 = {"first_name":"William","last_name":"Shakespeare","year_of_death":"1616"}
doc2 = {"first_name":"Bertold","last_name":"Brecht","year_of_death":"1956"}
collection.insert(doc1)
collection.insert(doc2)
conn.database_names()
db.collection_names()
collection.find_one()
list(collection.find())[:2]
collection.find({'year_of_death':'1956'}).count()
# # Redis
import redis
r = redis.StrictRedis(host='localhost', port=6379, db=0)
r.set('<NAME>', '1616')
r.set('<NAME>', '1956')
import pickle
class PicklePerson(object):
def __init__(self, first_name, last_name, year_of_death):
self.first_name = first_name
self.last_name = last_name
self.year_of_death = year_of_death
def __repr__(self):
return "Name: " + self.first_name + " " + self.last_name + \
"\n" + "Death: " + self.year_of_death
bob = pickle.dumps(PicklePerson("William","Shakespeare","1616"))
bert = pickle.dumps(PicklePerson("Bertold","Brecht","1956"))
r.set("bob", bob)
r.set("bert", bert)
pickle.loads(r.get("bob"))
# + active=""
# r.get('1616')
# -
print(r)
# #etcd
import etcd
client = etcd.Client(host='127.0.0.1', port=4001)
# Einen Key schreiben
client.write('/nodes/n1', 1)
client.write('/nodes/n2', 2)
# Den Key lesen
client.read('/nodes/n1').value
# +
directory = client.get("/nodes")
# loop through directory children
for result in directory.children:
print(result.key + ": " + result.value)
# -
# Ein paar Infos zum Cluster
client.machines
| examples_database/test_databases_python3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
class LRU_Cache(object):
def __init__(self, capacity):
# Initialize class variables
self.cache = {}
self.capacity = capacity
self.recent_key = None
def get(self, key):
# Retrieve item from provided key. Return -1 if nonexistent.
if key in self.cache.keys():
self.recent_key = key
return self.cache[key]
else:
return -1
def set(self, key, value):
# Set the value if the key is not present in the cache. If the cache is at capacity remove the oldest item.
if len(self.cache) < self.capacity:
self.cache[key] = value
else:
self.cache.pop(self.recent_key, None)
self.cache[key] = value
# +
our_cache = LRU_Cache(5)
our_cache.set(1, 1);
our_cache.set(2, 2);
our_cache.set(3, 3);
our_cache.set(4, 4);
print(our_cache.get(1)) # returns 1
print(our_cache.get(2)) # returns 2
print(our_cache.get(9)) # returns -1 because 9 is not present in the cache
our_cache.set(5, 5)
our_cache.set(6, 6)
print(our_cache.get(3)) # returns -1 because the cache reached it's capacity and 3 was the least recently used entry
# -
# Idealy, hashtable only takes O(1) complexity for search, insert and delete. Therefore, dictornary is used to implement this task.
| other_structure_&_algorithm/Least_Recent_Used_Cache.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to geospatial vector data in Python
# +
# %matplotlib inline
import pandas as pd
import geopandas
pd.options.display.max_rows = 10
# -
# ## Importing geospatial data
# Geospatial data is often available from specific GIS file formats or data stores, like ESRI shapefiles, GeoJSON files, geopackage files, PostGIS (PostgreSQL) database, ...
#
# We can use the GeoPandas library to read many of those GIS file formats (relying on the `fiona` library under the hood, which is an interface to GDAL/OGR), using the `geopandas.read_file` function.
#
# For example, let's start by reading a shapefile with all the countries of the world (adapted from http://www.naturalearthdata.com/downloads/110m-cultural-vectors/110m-admin-0-countries/, zip file is available in the `/data` directory), and inspect the data:
countries = geopandas.read_file("zip://./data/ne_110m_admin_0_countries.zip")
# or if the archive is unpacked:
# countries = geopandas.read_file("data/ne_110m_admin_0_countries/ne_110m_admin_0_countries.shp")
countries.head()
countries.plot()
# What can we observe:
#
# - Using `.head()` we can see the first rows of the dataset, just like we can do with Pandas.
# - There is a 'geometry' column and the different countries are represented as polygons
# - We can use the `.plot()` method to quickly get a *basic* visualization of the data
# ## What's a GeoDataFrame?
#
# We used the GeoPandas library to read in the geospatial data, and this returned us a `GeoDataFrame`:
type(countries)
# A GeoDataFrame contains a tabular, geospatial dataset:
#
# * It has a **'geometry' column** that holds the geometry information (or features in GeoJSON).
# * The other columns are the **attributes** (or properties in GeoJSON) that describe each of the geometries
#
# Such a `GeoDataFrame` is just like a pandas `DataFrame`, but with some additional functionality for working with geospatial data:
#
# * A `.geometry` attribute that always returns the column with the geometry information (returning a GeoSeries). The column name itself does not necessarily need to be 'geometry', but it will always be accessible as the `.geometry` attribute.
# * It has some extra methods for working with spatial data (area, distance, buffer, intersection, ...), which we will see in later notebooks
countries.geometry
type(countries.geometry)
countries.geometry.area
# **It's still a DataFrame**, so we have all the pandas functionality available to use on the geospatial dataset, and to do data manipulations with the attributes and geometry information together.
#
# For example, we can calculate average population number over all countries (by accessing the 'pop_est' column, and calling the `mean` method on it):
countries['pop_est'].mean()
# Or, we can use boolean filtering to select a subset of the dataframe based on a condition:
africa = countries[countries['continent'] == 'Africa']
africa.plot()
# ---
#
# **Exercise**: create a plot of South America
#
# <!--
# countries[countries['continent'] == 'South America'].plot()
# -->
# ---
countries.head()
# ---
#
# The rest of the tutorial is going to assume you already know some pandas basics, but we will try to give hints for that part for those that are not familiar.
# A few resources in case you want to learn more about pandas:
#
# - Pandas docs: https://pandas.pydata.org/pandas-docs/stable/10min.html
# - Other tutorials: chapter from pandas in https://jakevdp.github.io/PythonDataScienceHandbook/, https://github.com/jorisvandenbossche/pandas-tutorial, https://github.com/TomAugspurger/pandas-head-to-tail, ...
# <div class="alert alert-info" style="font-size:120%">
# <b>REMEMBER</b>: <br>
#
# <ul>
# <li>A `GeoDataFrame` allows to perform typical tabular data analysis together with spatial operations</li>
# <li>A `GeoDataFrame` (or *Feature Collection*) consists of:
# <ul>
# <li>**Geometries** or **features**: the spatial objects</li>
# <li>**Attributes** or **properties**: columns with information about each spatial object</li>
# </ul>
# </li>
# </ul>
# </div>
# ## Geometries: Points, Linestrings and Polygons
#
# Spatial **vector** data can consist of different types, and the 3 fundamental types are:
#
# * **Point** data: represents a single point in space.
# * **Line** data ("LineString"): represents a sequence of points that form a line.
# * **Polygon** data: represents a filled area.
#
# And each of them can also be combined in multi-part geometries (See https://shapely.readthedocs.io/en/stable/manual.html#geometric-objects for extensive overview).
# For the example we have seen up to now, the individual geometry objects are Polygons:
print(countries.geometry[2])
# Let's import some other datasets with different types of geometry objects.
#
# A dateset about cities in the world (adapted from http://www.naturalearthdata.com/downloads/110m-cultural-vectors/110m-populated-places/, zip file is available in the `/data` directory), consisting of Point data:
cities = geopandas.read_file("zip://./data/ne_110m_populated_places.zip")
print(cities.geometry[0])
# And a dataset of rivers in the world (from http://www.naturalearthdata.com/downloads/50m-physical-vectors/50m-rivers-lake-centerlines/, zip file is available in the `/data` directory) where each river is a (multi-)line:
rivers = geopandas.read_file("zip://./data/ne_50m_rivers_lake_centerlines.zip")
print(rivers.geometry[0])
# ### The `shapely` library
#
# The individual geometry objects are provided by the [`shapely`](https://shapely.readthedocs.io/en/stable/) library
type(countries.geometry[0])
# To construct one ourselves:
from shapely.geometry import Point, Polygon, LineString
p = Point(1, 1)
print(p)
polygon = Polygon([(1, 1), (2,2), (2, 1)])
# <div class="alert alert-info" style="font-size:120%">
# <b>REMEMBER</b>: <br><br>
#
# Single geometries are represented by `shapely` objects:
#
# <ul>
# <li>If you access a single geometry of a GeoDataFrame, you get a shapely geometry object</li>
# <li>Those objects have similar functionality as geopandas objects (GeoDataFrame/GeoSeries). For example:
# <ul>
# <li>`single_shapely_object.distance(other_point)` -> distance between two points</li>
# <li>`geodataframe.distance(other_point)` -> distance for each point in the geodataframe to the other point</li>
# </ul>
# </li>
# </ul>
# </div>
# ## Coordinate reference systems
#
# A **coordinate reference system (CRS)** determines how the two-dimensional (planar) coordinates of the geometry objects should be related to actual places on the (non-planar) earth.
#
# For a nice in-depth explanation, see https://docs.qgis.org/2.8/en/docs/gentle_gis_introduction/coordinate_reference_systems.html
# A GeoDataFrame or GeoSeries has a `.crs` attribute which holds (optionally) a description of the coordinate reference system of the geometries:
countries.crs
# For the `countries` dataframe, it indicates that it used the EPSG 4326 / WGS84 lon/lat reference system, which is one of the most used.
# It uses coordinates as latitude and longitude in degrees, as can you be seen from the x/y labels on the plot:
countries.plot()
# The `.crs` attribute is given as a dictionary. In this case, it only indicates the EPSG code, but it can also contain the full "proj4" string (in dictionary form).
#
# Under the hood, GeoPandas uses the `pyproj` / `proj4` libraries to deal with the re-projections.
#
# For more information, see also http://geopandas.readthedocs.io/en/latest/projections.html.
# ---
#
# There are sometimes good reasons you want to change the coordinate references system of your dataset, for example:
#
# - different sources with different crs -> need to convert to the same crs
# - distance-based operations -> if you a crs that has meter units (not degrees)
# - plotting in a certain crs (eg to preserve area)
#
# We can convert a GeoDataFrame to another reference system using the `to_crs` function.
#
# For example, let's convert the countries to the World Mercator projection (http://epsg.io/3395):
# remove Antartica, as the Mercator projection cannot deal with the poles
countries = countries[(countries['name'] != "Antarctica")]
countries_mercator = countries.to_crs(epsg=3395) # or .to_crs({'init': 'epsg:3395'})
countries_mercator.plot()
# Note the different scale of x and y.
# ---
#
# **Exercise**: project the countries to [Web Mercator](http://epsg.io/3857), the CRS used by Google Maps, OpenStreetMap and most web providers.
#
# <!--
# countries.to_crs(epsg=3857)
# -->
# ---
# ## Plotting our different layers together
ax = countries.plot(edgecolor='k', facecolor='none', figsize=(15, 10))
rivers.plot(ax=ax)
cities.plot(ax=ax, color='red')
ax.set(xlim=(-20, 60), ylim=(-40, 40))
# See the [04-more-on-visualization.ipynb](04-more-on-visualization.ipynb) notebook for more details on visualizing geospatial datasets.
# ---
#
# **Exercise**: replicate the figure above by coloring the countries in black and cities in yellow
#
# <!--
# ax = countries.plot(edgecolor='w', facecolor='k', figsize=(15, 10))
# rivers.plot(ax=ax)
# cities.plot(ax=ax, color='yellow')
# ax.set(xlim=(-20, 60), ylim=(-40, 40))
# -->
# ---
# ## A bit more on importing and creating GeoDataFrames
# ### Note on `fiona`
#
# Under the hood, GeoPandas uses the [Fiona library](http://toblerity.org/fiona/) (pythonic interface to GDAL/OGR) to read and write data. GeoPandas provides a more user-friendly wrapper, which is sufficient for most use cases. But sometimes you want more control, and in that case, to read a file with fiona you can do the following:
#
# +
import fiona
from shapely.geometry import shape
with fiona.drivers():
with fiona.open("data/ne_110m_admin_0_countries/ne_110m_admin_0_countries.shp") as collection:
for feature in collection:
# ... do something with geometry
geom = shape(feature['geometry'])
# ... do something with properties
print(feature['properties']['name'])
# -
# ### Constructing a GeoDataFrame manually
geopandas.GeoDataFrame({
'geometry': [Point(1, 1), Point(2, 2)],
'attribute1': [1, 2],
'attribute2': [0.1, 0.2]})
# ### Creating a GeoDataFrame from an existing dataframe
#
# For example, if you have lat/lon coordinates in two columns:
df = pd.DataFrame(
{'City': ['Buenos Aires', 'Brasilia', 'Santiago', 'Bogota', 'Caracas'],
'Country': ['Argentina', 'Brazil', 'Chile', 'Colombia', 'Venezuela'],
'Latitude': [-34.58, -15.78, -33.45, 4.60, 10.48],
'Longitude': [-58.66, -47.91, -70.66, -74.08, -66.86]})
df['Coordinates'] = list(zip(df.Longitude, df.Latitude))
df['Coordinates'] = df['Coordinates'].apply(Point)
gdf = geopandas.GeoDataFrame(df, geometry='Coordinates')
gdf
# See http://geopandas.readthedocs.io/en/latest/gallery/create_geopandas_from_pandas.html#sphx-glr-gallery-create-geopandas-from-pandas-py for full example
# ---
#
# **Exercise**: use [geojson.io](http://geojson.io) to mark five points, and create a `GeoDataFrame` with it. Note that coordinates will be expressed in longitude and latitude, so you'll have to set the CRS accordingly.
#
# <!--
# df = pd.DataFrame(
# {'Name': ['Hotel', 'Capitol', 'Barton Springs'],
# 'Latitude': [30.28195889019179, 30.274782936992608, 30.263728440902543],
# 'Longitude': [-97.74006128311157, -97.74038314819336, -97.77013421058655]})
# df['Coordinates'] = list(zip(df.Longitude, df.Latitude))
# df['Coordinates'] = df['Coordinates'].apply(Point)
# gdf = geopandas.GeoDataFrame(df, geometry='Coordinates', crs={'init': 'epsg:4326'})
# -->
# ---
| 01-introduction-geospatial-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="lJz6FDU1lRzc"
"""
You can run either this notebook locally (if you have all the dependencies and a GPU) or on Google Colab.
Instructions for setting up Colab are as follows:
1. Open a new Python 3 notebook.
2. Import this notebook from GitHub (File -> Upload Notebook -> "GITHUB" tab -> copy/paste GitHub URL)
3. Connect to an instance with a GPU (Runtime -> Change runtime type -> select "GPU" for hardware accelerator)
4. Run this cell to set up dependencies.
5. Restart the runtime (Runtime -> Restart Runtime) for any upgraded packages to take effect
"""
# If you're using Google Colab and not running locally, run this cell.
## Install dependencies
# !pip install wget
# !apt-get install sox libsndfile1 ffmpeg
# !pip install unidecode
# !pip install matplotlib>=3.3.2
## Install NeMo
BRANCH = 'v1.0.0'
# !python -m pip install git+https://github.com/NVIDIA/NeMo.git@$BRANCH#egg=nemo_toolkit[all]
## Grab the config we'll use in this example
# !mkdir configs
# !wget -P configs/ https://raw.githubusercontent.com/NVIDIA/NeMo/$BRANCH/examples/asr/conf/config.yaml
"""
Remember to restart the runtime for the kernel to pick up any upgraded packages (e.g. matplotlib)!
Alternatively, you can uncomment the exit() below to crash and restart the kernel, in the case
that you want to use the "Run All Cells" (or similar) option.
"""
# exit()
# + [markdown] id="v1Jk9etFlRzf"
# # Introduction to End-To-End Automatic Speech Recognition
#
# This notebook contains a basic tutorial of Automatic Speech Recognition (ASR) concepts, introduced with code snippets using the [NeMo framework](https://github.com/NVIDIA/NeMo).
# We will first introduce the basics of the main concepts behind speech recognition, then explore concrete examples of what the data looks like and walk through putting together a simple end-to-end ASR pipeline.
#
# We assume that you are familiar with general machine learning concepts and can follow Python code, and we'll be using the [AN4 dataset from CMU](http://www.speech.cs.cmu.edu/databases/an4/) (with processing using `sox`).
# + [markdown] id="YLln3U-IlRzg"
# ## Conceptual Overview: What is ASR?
#
# ASR, or **Automatic Speech Recognition**, refers to the problem of getting a program to automatically transcribe spoken language (speech-to-text). Our goal is usually to have a model that minimizes the **Word Error Rate (WER)** metric when transcribing speech input. In other words, given some audio file (e.g. a WAV file) containing speech, how do we transform this into the corresponding text with as few errors as possible?
#
# Traditional speech recognition takes a generative approach, modeling the full pipeline of how speech sounds are produced in order to evaluate a speech sample. We would start from a **language model** that encapsulates the most likely orderings of words that are generated (e.g. an n-gram model), to a **pronunciation model** for each word in that ordering (e.g. a pronunciation table), to an **acoustic model** that translates those pronunciations to audio waveforms (e.g. a Gaussian Mixture Model).
#
# Then, if we receive some spoken input, our goal would be to find the most likely sequence of text that would result in the given audio according to our generative pipeline of models. Overall, with traditional speech recognition, we try to model `Pr(audio|transcript)*Pr(transcript)`, and take the argmax of this over possible transcripts.
#
# Over time, neural nets advanced to the point where each component of the traditional speech recognition model could be replaced by a neural model that had better performance and that had a greater potential for generalization. For example, we could replace an n-gram model with a neural language model, and replace a pronunciation table with a neural pronunciation model, and so on. However, each of these neural models need to be trained individually on different tasks, and errors in any model in the pipeline could throw off the whole prediction.
#
# Thus, we can see the appeal of **end-to-end ASR architectures**: discriminative models that simply take an audio input and give a textual output, and in which all components of the architecture are trained together towards the same goal. The model's encoder would be akin to an acoustic model for extracting speech features, which can then be directly piped to a decoder which outputs text. If desired, we could integrate a language model that would improve our predictions, as well.
#
# And the entire end-to-end ASR model can be trained at once--a much easier pipeline to handle!
# + [markdown] id="0S5iZPMSlRzg"
# ### End-To-End ASR
#
# With an end-to-end model, we want to directly learn `Pr(transcript|audio)` in order to predict the transcripts from the original audio. Since we are dealing with sequential information--audio data over time that corresponds to a sequence of letters--RNNs are the obvious choice. But now we have a pressing problem to deal with: since our input sequence (number of audio timesteps) is not the same length as our desired output (transcript length), how do we match each time step from the audio data to the correct output characters?
#
# Earlier speech recognition approaches relied on **temporally-aligned data**, in which each segment of time in an audio file was matched up to a corresponding speech sound such as a phoneme or word. However, if we would like to have the flexibility to predict letter-by-letter to prevent OOV (out of vocabulary) issues, then each time step in the data would have to be labeled with the letter sound that the speaker is making at that point in the audio file. With that information, it seems like we should simply be able to try to predict the correct letter for each time step and then collapse the repeated letters (e.g. the prediction output `LLLAAAAPPTOOOPPPP` would become `LAPTOP`). It turns out that this idea has some problems: not only does alignment make the dataset incredibly labor-intensive to label, but also, what do we do with words like "book" that contain consecutive repeated letters? Simply squashing repeated letters together would not work in that case!
#
# 
#
# Modern end-to-end approaches get around this using methods that don't require manual alignment at all, so that the input-output pairs are really just the raw audio and the transcript--no extra data or labeling required. Let's briefly go over two popular approaches that allow us to do this, Connectionist Temporal Classification (CTC) and sequence-to-sequence models with attention.
#
# #### Connectionist Temporal Classification (CTC)
#
# In normal speech recognition prediction output, we would expect to have characters such as the letters from A through Z, numbers 0 through 9, spaces ("\_"), and so on. CTC introduces a new intermediate output token called the **blank token** ("-") that is useful for getting around the alignment issue.
#
# With CTC, we still predict one token per time segment of speech, but we use the blank token to figure out where we can and can't collapse the predictions. The appearance of a blank token helps separate repeating letters that should not be collapsed. For instance, with an audio snippet segmented into `T=11` time steps, we could get predictions that look like `BOO-OOO--KK`, which would then collapse to `"BO-O-K"`, and then we would remove the blank tokens to get our final output, `BOOK`.
#
# Now, we can predict one output token per time step, then collapse and clean to get sensible output without any fear of ambiguity from repeating letters! A simple way of getting predictions like this would be to apply a bidirectional RNN to the audio input, apply softmax over each time step's output, and then take the token with the highest probability. The method of always taking the best token at each time step is called **greedy decoding, or max decoding**.
#
# To calculate our loss for backprop, we would like to know the log probability of the model producing the correct transcript, `log(Pr(transcript|audio))`. We can get the log probability of a single intermediate output sequence (e.g. `BOO-OOO--KK`) by summing over the log probabilities we get from each token's softmax value, but note that the resulting sum is different from the log probability of the transcript itself (`BOOK`). This is because there are multiple possible output sequences of the same length that can be collapsed to get the same transcript (e.g. `BBO--OO-KKK` also results in `BOOK`), and so we need to **marginalize over every valid sequence of length `T` that collapses to the transcript**.
#
# Therefore, to get our transcript's log probability given our audio input, we must sum the log probabilities of every sequence of length `T` that collapses to the transcript (e.g. `log(Pr(output: "BOOK"|audio)) = log(Pr(BOO-OOO--KK|audio)) + log(Pr(BBO--OO-KKK|audio)) + ...`). In practice, we can use a dynamic programming approach to calculate this, accumulating our log probabilities over different "paths" through the softmax outputs at each time step.
#
# If you would like a more in-depth explanation of how CTC works, or how we can improve our results by using a modified beam search algorithm, feel free to check out the Further Reading section at the end of this notebook for more resources.
#
# #### Sequence-to-Sequence with Attention
#
# One problem with CTC is that predictions at different time steps are conditionally independent, which is an issue because the words in a continuous utterance tend to be related to each other in some sensible way. With this conditional independence assumption, we can't learn a language model that can represent such dependencies, though we can add a language model on top of the CTC output to mitigate this to some degree.
#
# A popular alternative is to use a sequence-to-sequence model with attention. A typical seq2seq model for ASR consists of some sort of **bidirectional RNN encoder** that consumes the audio sequence timestep-by-timestep, and where the outputs are then passed to an **attention-based decoder**. Each prediction from the decoder is based on attending to some parts of the entire encoded input, as well as the previously outputted tokens.
#
# The outputs of the decoder can be anything from word pieces to phonemes to letters, and since predictions are not directly tied to time steps of the input, we can just continue producing tokens one-by-one until an end token is given (or we reach a specified max output length). This way, we do not need to deal with audio alignment, and our predicted transcript is just the sequence of outputs given by our decoder.
#
# Now that we have an idea of what some popular end-to-end ASR models look like, let's take a look at the audio data we'll be working with for our example.
# + [markdown] id="38aYTCTIlRzh"
# ## Taking a Look at Our Data (AN4)
#
# The AN4 dataset, also known as the Alphanumeric dataset, was collected and published by Carnegie Mellon University. It consists of recordings of people spelling out addresses, names, telephone numbers, etc., one letter or number at a time, as well as their corresponding transcripts. We choose to use AN4 for this tutorial because it is relatively small, with 948 training and 130 test utterances, and so it trains quickly.
#
# Before we get started, let's download and prepare the dataset. The utterances are available as `.sph` files, so we will need to convert them to `.wav` for later processing. If you are not using Google Colab, please make sure you have [Sox](http://sox.sourceforge.net/) installed for this step--see the "Downloads" section of the linked Sox homepage. (If you are using Google Colab, Sox should have already been installed in the setup cell at the beginning.)
# + id="gAhsmi6HlRzh"
# This is where the an4/ directory will be placed.
# Change this if you don't want the data to be extracted in the current directory.
data_dir = '.'
# + id="Yb4fuUvWlRzk"
import glob
import os
import subprocess
import tarfile
import wget
# Download the dataset. This will take a few moments...
print("******")
if not os.path.exists(data_dir + '/an4_sphere.tar.gz'):
an4_url = 'http://www.speech.cs.cmu.edu/databases/an4/an4_sphere.tar.gz'
an4_path = wget.download(an4_url, data_dir)
print(f"Dataset downloaded at: {an4_path}")
else:
print("Tarfile already exists.")
an4_path = data_dir + '/an4_sphere.tar.gz'
if not os.path.exists(data_dir + '/an4/'):
# Untar and convert .sph to .wav (using sox)
tar = tarfile.open(an4_path)
tar.extractall(path=data_dir)
print("Converting .sph to .wav...")
sph_list = glob.glob(data_dir + '/an4/**/*.sph', recursive=True)
for sph_path in sph_list:
wav_path = sph_path[:-4] + '.wav'
cmd = ["sox", sph_path, wav_path]
subprocess.run(cmd)
print("Finished conversion.\n******")
# + [markdown] id="m_LFeM0elRzm"
# You should now have a folder called `an4` that contains `etc/an4_train.transcription`, `etc/an4_test.transcription`, audio files in `wav/an4_clstk` and `wav/an4test_clstk`, along with some other files we will not need.
#
# Now we can load and take a look at the data. As an example, file `cen2-mgah-b.wav` is a 2.6 second-long audio recording of a man saying the letters "G L E N N" one-by-one. To confirm this, we can listen to the file:
# + id="_M_bSs3MjQlz"
import librosa
import IPython.display as ipd
# Load and listen to the audio file
example_file = data_dir + '/an4/wav/an4_clstk/mgah/cen2-mgah-b.wav'
audio, sample_rate = librosa.load(example_file)
ipd.Audio(example_file, rate=sample_rate)
# + [markdown] id="qZyElgPVjQl5"
# In an ASR task, if this WAV file was our input, then "G L E N N" would be our desired output.
#
# Let's plot the waveform, which is simply a line plot of the sequence of values that we read from the file. This is a format of viewing audio that you are likely to be familiar with seeing in many audio editors and visualizers:
# + id="MqIAKkqelRzm"
# %matplotlib inline
import librosa.display
import matplotlib.pyplot as plt
# Plot our example audio file's waveform
plt.rcParams['figure.figsize'] = (15,7)
plt.title('Waveform of Audio Example')
plt.ylabel('Amplitude')
_ = librosa.display.waveplot(audio)
# + [markdown] id="Gg6RR_yolRzo"
# We can see the activity in the waveform that corresponds to each letter in the audio, as our speaker here enunciates quite clearly!
# You can kind of tell that each spoken letter has a different "shape," and it's interesting to note that last two blobs look relatively similar, which is expected because they are both the letter "N."
#
# ### Spectrograms and Mel Spectrograms
#
# However, since audio information is more useful in the context of frequencies of sound over time, we can get a better representation than this raw sequence of 57,330 values.
# We can apply a [Fourier Transform](https://en.wikipedia.org/wiki/Fourier_transform) on our audio signal to get something more useful: a **spectrogram**, which is a representation of the energy levels (i.e. amplitude, or "loudness") of each frequency (i.e. pitch) of the signal over the duration of the file.
# A spectrogram (which can be viewed as a heat map) is a good way of seeing how the *strengths of various frequencies in the audio vary over time*, and is obtained by breaking up the signal into smaller, usually overlapping chunks and performing a Short-Time Fourier Transform (STFT) on each.
#
# Let's examine what the spectrogram of our sample looks like.
# + id="oCFneEs1lRzp"
import numpy as np
# Get spectrogram using Librosa's Short-Time Fourier Transform (stft)
spec = np.abs(librosa.stft(audio))
spec_db = librosa.amplitude_to_db(spec, ref=np.max) # Decibels
# Use log scale to view frequencies
librosa.display.specshow(spec_db, y_axis='log', x_axis='time')
plt.colorbar()
plt.title('Audio Spectrogram');
# + [markdown] id="9OPc4tcalRzs"
# Again, we are able to see each letter being pronounced, and that the last two blobs that correspond to the "N"s are pretty similar-looking. But how do we interpret these shapes and colors? Just as in the waveform plot before, we see time passing on the x-axis (all 2.6s of audio). But now, the y-axis represents different frequencies (on a log scale), and *the color on the plot shows the strength of a frequency at a particular point in time*.
#
# We're still not done yet, as we can make one more potentially useful tweak: using the **Mel Spectrogram** instead of the normal spectrogram. This is simply a change in the frequency scale that we use from linear (or logarithmic) to the mel scale, which is "a perceptual scale of pitches judged by listeners to be equal in distance from one another" (from [Wikipedia](https://en.wikipedia.org/wiki/Mel_scale)).
#
# In other words, it's a transformation of the frequencies to be more aligned to what humans perceive; a change of +1000Hz from 2000Hz->3000Hz sounds like a larger difference to us than 9000Hz->10000Hz does, so the mel scale normalizes this such that equal distances sound like equal differences to the human ear. Intuitively, we use the mel spectrogram because in this case we are processing and transcribing human speech, such that transforming the scale to better match what we hear is a useful procedure.
# + id="7yQXVn-TlRzt"
# Plot the mel spectrogram of our sample
mel_spec = librosa.feature.melspectrogram(audio, sr=sample_rate)
mel_spec_db = librosa.power_to_db(mel_spec, ref=np.max)
librosa.display.specshow(
mel_spec_db, x_axis='time', y_axis='mel')
plt.colorbar()
plt.title('Mel Spectrogram');
# + [markdown] id="RSCyVizDlRz1"
# ## Convolutional ASR Models
#
# Let's take a look at the model that we will be building, and how we specify its parameters.
#
# ### The Jasper Model
#
# We will be training a small [Jasper (Just Another SPeech Recognizer) model](https://arxiv.org/abs/1904.03288) from scratch (e.g. initialized randomly).
# In brief, Jasper architectures consist of a repeated block structure that utilizes 1D convolutions.
# In a Jasper_KxR model, `R` sub-blocks (consisting of a 1D convolution, batch norm, ReLU, and dropout) are grouped into a single block, which is then repeated `K` times.
# We also have a one extra block at the beginning and a few more at the end that are invariant of `K` and `R`, and we use CTC loss.
#
# ### The QuartzNet Model
#
# The QuartzNet is better variant of Jasper with a key difference that it uses time-channel separable 1D convolutions. This allows it to dramatically reduce number of weights while keeping similar accuracy.
#
# A Jasper/QuartzNet models look like this (QuartzNet model is pictured):
#
# 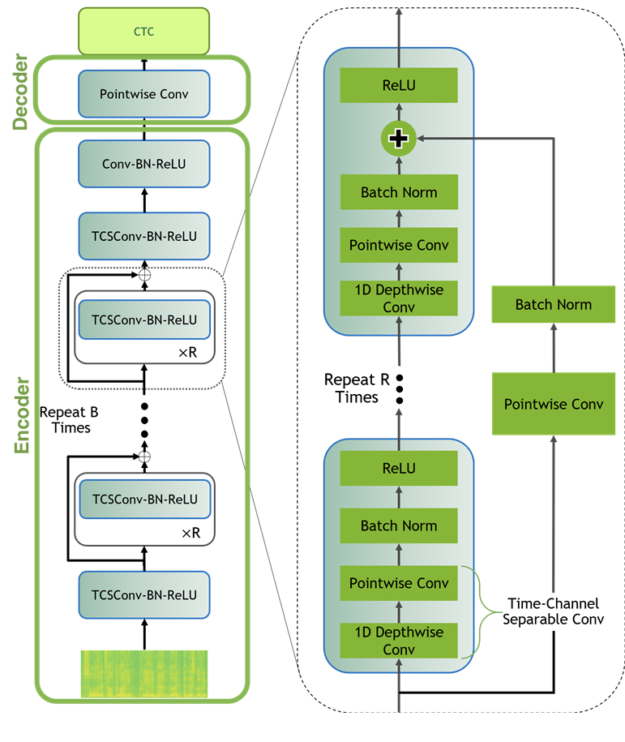
# + [markdown] id="gEpNci7slRzw"
# # Using NeMo for Automatic Speech Recognition
#
# Now that we have an idea of what ASR is and how the audio data looks like, we can start using NeMo to do some ASR!
#
# We'll be using the **Neural Modules (NeMo) toolkit** for this part, so if you haven't already, you should download and install NeMo and its dependencies. To do so, just follow the directions on the [GitHub page](https://github.com/NVIDIA/NeMo), or in the [documentation](https://docs.nvidia.com/deeplearning/nemo/user-guide/docs/en/v1.0.0/).
#
# NeMo lets us easily hook together the components (modules) of our model, such as the data layer, intermediate layers, and various losses, without worrying too much about implementation details of individual parts or connections between modules. NeMo also comes with complete models which only require your data and hyperparameters for training.
# + id="4_W0lhaQlRzx"
# NeMo's "core" package
import nemo
# NeMo's ASR collection - this collections contains complete ASR models and
# building blocks (modules) for ASR
import nemo.collections.asr as nemo_asr
# + [markdown] id="v_W8EbYktZE3"
# ## Using an Out-of-the-Box Model
#
# NeMo's ASR collection comes with many building blocks and even complete models that we can use for training and evaluation. Moreover, several models come with pre-trained weights. Let's instantiate a complete QuartzNet15x5 model.
# + id="KFZZpYult96G"
# This line will download pre-trained QuartzNet15x5 model from NVIDIA's NGC cloud and instantiate it for you
quartznet = nemo_asr.models.EncDecCTCModel.from_pretrained(model_name="QuartzNet15x5Base-En")
# + [markdown] id="KucxoFJhum0i"
# Next, we'll simply add paths to files we want to transcribe into the list and pass it to our model. Note that it will work for relatively short (<25 seconds) files.
# + id="3QCpR_93u1hp"
files = ['./an4/wav/an4_clstk/mgah/cen2-mgah-b.wav']
for fname, transcription in zip(files, quartznet.transcribe(paths2audio_files=files)):
print(f"Audio in {fname} was recognized as: {transcription}")
# + [markdown] id="ppUm_kuavm_f"
# That was easy! But there are plenty of scenarios where you would want to fine-tune the model on your own data or even train from scratch. For example, this out-of-the box model will obviously not work for Spanish and would likely perform poorly for telephone audio. So if you have collected your own data, you certainly should attempt to fine-tune or train on it!
# + [markdown] id="ABUDaC5Js7AW"
# ## Training from Scratch
#
# To train from scratch, you need to prepare your training data in the right format and specify your models architecture.
# + [markdown] id="RdNyw1b_zgtm"
# ### Creating Data Manifests
#
# The first thing we need to do now is to create manifests for our training and evaluation data, which will contain the metadata of our audio files. NeMo data sets take in a standardized manifest format where each line corresponds to one sample of audio, such that the number of lines in a manifest is equal to the number of samples that are represented by that manifest. A line must contain the path to an audio file, the corresponding transcript (or path to a transcript file), and the duration of the audio sample.
#
# Here's an example of what one line in a NeMo-compatible manifest might look like:
# ```
# {"audio_filepath": "path/to/audio.wav", "duration": 3.45, "text": "this is a nemo tutorial"}
# ```
#
# We can build our training and evaluation manifests using `an4/etc/an4_train.transcription` and `an4/etc/an4_test.transcription`, which have lines containing transcripts and their corresponding audio file IDs:
# ```
# ...
# <s> P I T T S B U R G H </s> (cen5-fash-b)
# <s> TWO SIX EIGHT FOUR FOUR ONE EIGHT </s> (cen7-fash-b)
# ...
# ```
# + id="lVB1sG1GlRzz"
# --- Building Manifest Files --- #
import json
# Function to build a manifest
def build_manifest(transcripts_path, manifest_path, wav_path):
with open(transcripts_path, 'r') as fin:
with open(manifest_path, 'w') as fout:
for line in fin:
# Lines look like this:
# <s> transcript </s> (fileID)
transcript = line[: line.find('(')-1].lower()
transcript = transcript.replace('<s>', '').replace('</s>', '')
transcript = transcript.strip()
file_id = line[line.find('(')+1 : -2] # e.g. "cen4-fash-b"
audio_path = os.path.join(
data_dir, wav_path,
file_id[file_id.find('-')+1 : file_id.rfind('-')],
file_id + '.wav')
duration = librosa.core.get_duration(filename=audio_path)
# Write the metadata to the manifest
metadata = {
"audio_filepath": audio_path,
"duration": duration,
"text": transcript
}
json.dump(metadata, fout)
fout.write('\n')
# Building Manifests
print("******")
train_transcripts = data_dir + '/an4/etc/an4_train.transcription'
train_manifest = data_dir + '/an4/train_manifest.json'
if not os.path.isfile(train_manifest):
build_manifest(train_transcripts, train_manifest, 'an4/wav/an4_clstk')
print("Training manifest created.")
test_transcripts = data_dir + '/an4/etc/an4_test.transcription'
test_manifest = data_dir + '/an4/test_manifest.json'
if not os.path.isfile(test_manifest):
build_manifest(test_transcripts, test_manifest, 'an4/wav/an4test_clstk')
print("Test manifest created.")
print("***Done***")
# + [markdown] id="W2fShQzRzo-M"
# ### Specifying Our Model with a YAML Config File
#
# For this tutorial, we'll build a *Jasper_4x1 model*, with `K=4` blocks of single (`R=1`) sub-blocks and a *greedy CTC decoder*, using the configuration found in `./configs/config.yaml`.
#
# If we open up this config file, we find model section which describes architecture of our model. A model contains an entry labeled `encoder`, with a field called `jasper` that contains a list with multiple entries. Each of the members in this list specifies one block in our model, and looks something like this:
# ```
# - filters: 128
# repeat: 1
# kernel: [11]
# stride: [2]
# dilation: [1]
# dropout: 0.2
# residual: false
# separable: true
# se: true
# se_context_size: -1
# ```
# The first member of the list corresponds to the first block in the Jasper architecture diagram, which appears regardless of `K` and `R`.
# Next, we have four entries that correspond to the `K=4` blocks, and each has `repeat: 1` since we are using `R=1`.
# These are followed by two more entries for the blocks that appear at the end of our Jasper model before the CTC loss.
#
# There are also some entries at the top of the file that specify how we will handle training (`train_ds`) and validation (`validation_ds`) data.
#
# Using a YAML config such as this is helpful for getting a quick and human-readable overview of what your architecture looks like, and allows you to swap out model and run configurations easily without needing to change your code.
# + id="PXVKBniMlRz5"
# --- Config Information ---#
try:
from ruamel.yaml import YAML
except ModuleNotFoundError:
from ruamel_yaml import YAML
config_path = './configs/config.yaml'
yaml = YAML(typ='safe')
with open(config_path) as f:
params = yaml.load(f)
print(params)
# + [markdown] id="wUmq3p2Aw_5N"
# ### Training with PyTorch Lightning
#
# NeMo models and modules can be used in any PyTorch code where torch.nn.Module is expected.
#
# However, NeMo's models are based on [PytorchLightning's](https://github.com/PyTorchLightning/pytorch-lightning) LightningModule and we recommend you use PytorchLightning for training and fine-tuning as it makes using mixed precision and distributed training very easy. So to start, let's create Trainer instance for training on GPU for 50 epochs
# + id="GUfR6tAK0k2u"
import pytorch_lightning as pl
trainer = pl.Trainer(gpus=1, max_epochs=50)
# + [markdown] id="IEn2RyvgxxvO"
# Next, we instantiate and ASR model based on our ``config.yaml`` file from the previous section.
# Note that this is a stage during which we also tell the model where our training and validation manifests are.
# + id="Cbf0fsMK09lk"
from omegaconf import DictConfig
params['model']['train_ds']['manifest_filepath'] = train_manifest
params['model']['validation_ds']['manifest_filepath'] = test_manifest
first_asr_model = nemo_asr.models.EncDecCTCModel(cfg=DictConfig(params['model']), trainer=trainer)
# + [markdown] id="hWtzwL5qXTYq"
# With that, we can start training with just one line!
# + id="inRJsnrz1psq"
# Start training!!!
trainer.fit(first_asr_model)
# + [markdown] id="jpYXX-GslR0E"
# There we go! We've put together a full training pipeline for the model and trained it for 50 epochs.
#
# If you'd like to save this model checkpoint for loading later (e.g. for fine-tuning, or for continuing training), you can simply call `first_asr_model.save_to(<checkpoint_path>)`. Then, to restore your weights, you can rebuild the model using the config (let's say you call it `first_asr_model_continued` this time) and call `first_asr_model_continued.restore_from(<checkpoint_path>)`.
#
# ### After Training: Monitoring Progress and Changing Hyperparameters
# We can now start Tensorboard to see how training went. Recall that WER stands for Word Error Rate and so the lower it is, the better.
# + id="n_0y3stSXDX_"
try:
from google import colab
COLAB_ENV = True
except (ImportError, ModuleNotFoundError):
COLAB_ENV = False
# Load the TensorBoard notebook extension
if COLAB_ENV:
# %load_ext tensorboard
# %tensorboard --logdir lightning_logs/
else:
print("To use tensorboard, please use this notebook in a Google Colab environment.")
# + [markdown] id="Z0h-BME7U8yb"
# We could improve this model by playing with hyperparameters. We can look at the current hyperparameters with the following:
# + id="7kdQbpohXnEd"
print(params['model']['optim'])
# + [markdown] id="sGZzRCvIW8kE"
# Let's say we wanted to change the learning rate. To do so, we can create a `new_opt` dict and set our desired learning rate, then call `<model>.setup_optimization()` with the new optimization parameters.
# + id="AbigFKUtYgvn"
import copy
new_opt = copy.deepcopy(params['model']['optim'])
new_opt['lr'] = 0.001
first_asr_model.setup_optimization(optim_config=DictConfig(new_opt))
# And then you can invoke trainer.fit(first_asr_model)
# + [markdown] id="D5Kwg8Cz-aaO"
# ## Inference
#
# Let's have a quick look at how one could run inference with NeMo's ASR model.
#
# First, ``EncDecCTCModel`` and its subclasses contain a handy ``transcribe`` method which can be used to simply obtain audio files' transcriptions. It also has batch_size argument to improve performance.
# + id="3FT0klSV268p"
print(first_asr_model.transcribe(paths2audio_files=['./an4/wav/an4_clstk/mgah/cen2-mgah-b.wav',
'./an4/wav/an4_clstk/fmjd/cen7-fmjd-b.wav',
'./an4/wav/an4_clstk/fmjd/cen8-fmjd-b.wav',
'./an4/wav/an4_clstk/fkai/cen8-fkai-b.wav'],
batch_size=4))
# + [markdown] id="6FiCfLX0D7py"
# Below is an example of a simple inference loop in pure PyTorch. It also shows how one can compute Word Error Rate (WER) metric between predictions and references.
# + id="7mP4r1Gx_Ilt"
# Bigger batch-size = bigger throughput
params['model']['validation_ds']['batch_size'] = 16
# Setup the test data loader and make sure the model is on GPU
first_asr_model.setup_test_data(test_data_config=params['model']['validation_ds'])
first_asr_model.cuda()
# We will be computing Word Error Rate (WER) metric between our hypothesis and predictions.
# WER is computed as numerator/denominator.
# We'll gather all the test batches' numerators and denominators.
wer_nums = []
wer_denoms = []
# Loop over all test batches.
# Iterating over the model's `test_dataloader` will give us:
# (audio_signal, audio_signal_length, transcript_tokens, transcript_length)
# See the AudioToCharDataset for more details.
for test_batch in first_asr_model.test_dataloader():
test_batch = [x.cuda() for x in test_batch]
targets = test_batch[2]
targets_lengths = test_batch[3]
log_probs, encoded_len, greedy_predictions = first_asr_model(
input_signal=test_batch[0], input_signal_length=test_batch[1]
)
# Notice the model has a helper object to compute WER
first_asr_model._wer.update(greedy_predictions, targets, targets_lengths)
_, wer_num, wer_denom = first_asr_model._wer.compute()
first_asr_model._wer.reset()
wer_nums.append(wer_num.detach().cpu().numpy())
wer_denoms.append(wer_denom.detach().cpu().numpy())
# Release tensors from GPU memory
del test_batch, log_probs, targets, targets_lengths, encoded_len, greedy_predictions
# We need to sum all numerators and denominators first. Then divide.
print(f"WER = {sum(wer_nums)/sum(wer_denoms)}")
# + [markdown] id="0kM9kBNOCptf"
# This WER is not particularly impressive and could be significantly improved. You could train longer (try 100 epochs) to get a better number. Check out the next section on how to improve it further.
# + [markdown] id="RBcJtg5ulR0H"
# ## Model Improvements
#
# You already have all you need to create your own ASR model in NeMo, but there are a few more tricks that you can employ if you so desire. In this section, we'll briefly cover a few possibilities for improving an ASR model.
#
# ### Data Augmentation
#
# There exist several ASR data augmentation methods that can increase the size of our training set.
#
# For example, we can perform augmentation on the spectrograms by zeroing out specific frequency segments ("frequency masking") or time segments ("time masking") as described by [SpecAugment](https://arxiv.org/abs/1904.08779), or zero out rectangles on the spectrogram as in [Cutout](https://arxiv.org/pdf/1708.04552.pdf). In NeMo, we can do all three of these by simply adding in a `SpectrogramAugmentation` neural module. (As of now, it does not perform the time warping from the SpecAugment paper.)
#
# Our toy model does not do spectrogram augmentation. But the real one we got from cloud does:
# + id="9glGogaPlR0H"
print(quartznet._cfg['spec_augment'])
# + [markdown] id="LdwdcA_a640R"
# If you want to enable SpecAugment in your model, make sure your .yaml config file contains 'model/spec_augment' section which looks like the one above.
# + [markdown] id="2f142kIQc1Z2"
# ### Transfer learning
#
# Transfer learning is an important machine learning technique that uses a model’s knowledge of one task to make it perform better on another. Fine-tuning is one of the techniques to perform transfer learning. It is an essential part of the recipe for many state-of-the-art results where a base model is first pretrained on a task with abundant training data and then fine-tuned on different tasks of interest where the training data is less abundant or even scarce.
#
# In ASR you might want to do fine-tuning in multiple scenarios, for example, when you want to improve your model's performance on a particular domain (medical, financial, etc.) or on accented speech. You can even transfer learn from one language to another! Check out [this paper](https://arxiv.org/abs/2005.04290) for examples.
#
# Transfer learning with NeMo is simple. Let's demonstrate how the model we got from the cloud could be fine-tuned on AN4 data. (NOTE: this is a toy example). And, while we are at it, we will change model's vocabulary, just to demonstrate how it's done.
# + id="hl320dsydWX0"
# Check what kind of vocabulary/alphabet the model has right now
print(quartznet.decoder.vocabulary)
# Let's add "!" symbol there. Note that you can (and should!) change the vocabulary
# entirely when fine-tuning using a different language.
quartznet.change_vocabulary(
new_vocabulary=[
' ', 'a', 'b', 'c', 'd', 'e', 'f', 'g', 'h', 'i', 'j', 'k', 'l', 'm', 'n',
'o', 'p', 'q', 'r', 's', 't', 'u', 'v', 'w', 'x', 'y', 'z', "'", "!"
]
)
# + [markdown] id="M7lvmiMSd3Aw"
# After this, our decoder has completely changed, but our encoder (which is where most of the weights are) remained intact. Let's fine tune-this model for 2 epochs on AN4 dataset. We will also use the smaller learning rate from ``new_opt` (see the "After Training" section)`.
# + id="_PZJIso-eDl-"
# Use the smaller learning rate we set before
quartznet.setup_optimization(optim_config=DictConfig(new_opt))
# Point to the data we'll use for fine-tuning as the training set
quartznet.setup_training_data(train_data_config=params['model']['train_ds'])
# Point to the new validation data for fine-tuning
quartznet.setup_validation_data(val_data_config=params['model']['validation_ds'])
# And now we can create a PyTorch Lightning trainer and call `fit` again.
trainer = pl.Trainer(gpus=[1], max_epochs=2)
trainer.fit(quartznet)
# + [markdown] id="VURa1NavlR0U"
# ### Fast Training
#
# Last but not least, we could simply speed up training our model! If you have the resources, you can speed up training by splitting the workload across multiple GPUs. Otherwise (or in addition), there's always mixed precision training, which allows you to increase your batch size.
#
# You can use [PyTorch Lightning's Trainer object](https://pytorch-lightning.readthedocs.io/en/latest/common/trainer.html?highlight=Trainer) to handle mixed-precision and distributed training for you. Below are some examples of flags you would pass to the `Trainer` to use these features:
#
# ```python
# # Mixed precision:
# trainer = pl.Trainer(amp_level='O1', precision=16)
#
# # Trainer with a distributed backend:
# trainer = pl.Trainer(gpus=2, num_nodes=2, accelerator='ddp')
#
# # Of course, you can combine these flags as well.
# ```
#
# Finally, have a look at [example scripts in NeMo repository](https://github.com/NVIDIA/NeMo/blob/main/examples/asr/speech_to_text.py) which can handle mixed precision and distributed training using command-line arguments.
# + [markdown] id="d1ym8QT3jQnj"
# ### Deployment
#
# Let's get back to our pre-trained model and see how easy it can be exported to an ONNX file
# in order to run it in an inference engine like TensorRT or ONNXRuntime.
# If you don't have one, let's install it:
# + id="I4WRcmakjQnj"
# !mkdir -p ort
# %cd ort
# !git clean -xfd
# !git clone --depth 1 --branch v1.5.2 https://github.com/microsoft/onnxruntime.git .
# !./build.sh --skip_tests --config Release --build_shared_lib --parallel --use_cuda --cuda_home /usr/local/cuda --cudnn_home /usr/lib/x86_64-linux-gnu --build_wheel
# !pip uninstall -y onnxruntime
# !pip uninstall -y onnxruntime-gpu
# !pip install --upgrade --force-reinstall ./build/Linux/Release/dist/onnxruntime*.whl
# %cd ..
# + [markdown] id="F9yO1BEbjQnm"
# Then run
# + id="HZnyWxPyjQnm"
import json
import os
import tempfile
import onnxruntime
import torch
import numpy as np
import nemo.collections.asr as nemo_asr
from nemo.collections.asr.data.audio_to_text import AudioToCharDataset
from nemo.collections.asr.metrics.wer import WER
def to_numpy(tensor):
return tensor.detach().cpu().numpy() if tensor.requires_grad else tensor.cpu().numpy()
def setup_transcribe_dataloader(cfg, vocabulary):
config = {
'manifest_filepath': os.path.join(cfg['temp_dir'], 'manifest.json'),
'sample_rate': 16000,
'labels': vocabulary,
'batch_size': min(cfg['batch_size'], len(cfg['paths2audio_files'])),
'trim_silence': True,
'shuffle': False,
}
dataset = AudioToCharDataset(
manifest_filepath=config['manifest_filepath'],
labels=config['labels'],
sample_rate=config['sample_rate'],
int_values=config.get('int_values', False),
augmentor=None,
max_duration=config.get('max_duration', None),
min_duration=config.get('min_duration', None),
max_utts=config.get('max_utts', 0),
blank_index=config.get('blank_index', -1),
unk_index=config.get('unk_index', -1),
normalize=config.get('normalize_transcripts', False),
trim=config.get('trim_silence', True),
parser=config.get('parser', 'en'),
)
return torch.utils.data.DataLoader(
dataset=dataset,
batch_size=config['batch_size'],
collate_fn=dataset.collate_fn,
drop_last=config.get('drop_last', False),
shuffle=False,
num_workers=config.get('num_workers', 0),
pin_memory=config.get('pin_memory', False),
)
quartznet = nemo_asr.models.EncDecCTCModel.from_pretrained(model_name="QuartzNet15x5Base-En")
quartznet.export('qn.onnx')
ort_session = onnxruntime.InferenceSession('qn.onnx')
with tempfile.TemporaryDirectory() as tmpdir:
with open(os.path.join(tmpdir, 'manifest.json'), 'w') as fp:
for audio_file in files:
entry = {'audio_filepath': audio_file, 'duration': 100000, 'text': 'nothing'}
fp.write(json.dumps(entry) + '\n')
config = {'paths2audio_files': files, 'batch_size': 4, 'temp_dir': tmpdir}
temporary_datalayer = setup_transcribe_dataloader(config, quartznet.decoder.vocabulary)
for test_batch in temporary_datalayer:
processed_signal, processed_signal_len = quartznet.preprocessor(
input_signal=test_batch[0].to(quartznet.device), length=test_batch[1].to(quartznet.device)
)
ort_inputs = {ort_session.get_inputs()[0].name: to_numpy(processed_signal),}
ologits = ort_session.run(None, ort_inputs)
alogits = np.asarray(ologits)
logits = torch.from_numpy(alogits[0])
greedy_predictions = logits.argmax(dim=-1, keepdim=False)
wer = WER(vocabulary=quartznet.decoder.vocabulary, batch_dim_index=0, use_cer=False, ctc_decode=True)
hypotheses = wer.ctc_decoder_predictions_tensor(greedy_predictions)
print(hypotheses)
break
# + [markdown] id="wteGqroafWg1"
# ## Under the Hood
#
# NeMo is open-source and we do all our model development in the open, so you can inspect our code if you wish.
#
# In particular, ``nemo_asr.model.EncDecCTCModel`` is an encoder-decoder model which is constructed using several ``Neural Modules`` taken from ``nemo_asr.modules.`` Here is what its forward pass looks like:
# ```python
# def forward(self, input_signal, input_signal_length):
# processed_signal, processed_signal_len = self.preprocessor(
# input_signal=input_signal, length=input_signal_length,
# )
# # Spec augment is not applied during evaluation/testing
# if self.spec_augmentation is not None and self.training:
# processed_signal = self.spec_augmentation(input_spec=processed_signal)
# encoded, encoded_len = self.encoder(audio_signal=processed_signal, length=processed_signal_len)
# log_probs = self.decoder(encoder_output=encoded)
# greedy_predictions = log_probs.argmax(dim=-1, keepdim=False)
# return log_probs, encoded_len, greedy_predictions
# ```
# Here:
#
# * ``self.preprocessor`` is an instance of ``nemo_asr.modules.AudioToMelSpectrogramPreprocessor``, which is a neural module that takes audio signal and converts it into a Mel-Spectrogram
# * ``self.spec_augmentation`` - is a neural module of type ```nemo_asr.modules.SpectrogramAugmentation``, which implements data augmentation.
# * ``self.encoder`` - is a convolutional Jasper/QuartzNet-like encoder of type ``nemo_asr.modules.ConvASREncoder``
# * ``self.decoder`` - is a ``nemo_asr.modules.ConvASRDecoder`` which simply projects into the target alphabet (vocabulary).
#
# Also, ``EncDecCTCModel`` uses the audio dataset class ``nemo_asr.data.AudioToCharDataset`` and CTC loss implemented in ``nemo_asr.losses.CTCLoss``.
#
# You can use these and other neural modules (or create new ones yourself!) to construct new ASR models.
# + [markdown] id="smzlvbhelR0U"
# # Further Reading/Watching:
#
# That's all for now! If you'd like to learn more about the topics covered in this tutorial, here are some resources that may interest you:
# - [Stanford Lecture on ASR](https://www.youtube.com/watch?v=3MjIkWxXigM)
# - ["An Intuitive Explanation of Connectionist Temporal Classification"](https://towardsdatascience.com/intuitively-understanding-connectionist-temporal-classification-3797e43a86c)
# - [Explanation of CTC with Prefix Beam Search](https://medium.com/corti-ai/ctc-networks-and-language-models-prefix-beam-search-explained-c11d1ee23306)
# - [Listen Attend and Spell Paper (seq2seq ASR model)](https://arxiv.org/abs/1508.01211)
# - [Explanation of the mel spectrogram in more depth](https://towardsdatascience.com/getting-to-know-the-mel-spectrogram-31bca3e2d9d0)
# - [Jasper Paper](https://arxiv.org/abs/1904.03288)
# - [QuartzNet paper](https://arxiv.org/abs/1910.10261)
# - [SpecAugment Paper](https://arxiv.org/abs/1904.08779)
# - [Explanation and visualization of SpecAugment](https://towardsdatascience.com/state-of-the-art-audio-data-augmentation-with-google-brains-specaugment-and-pytorch-d3d1a3ce291e)
# - [Cutout Paper](https://arxiv.org/pdf/1708.04552.pdf)
# - [Transfer Learning Blogpost](https://developer.nvidia.com/blog/jump-start-training-for-speech-recognition-models-with-nemo/)
# + id="V3ERGX86lR0V"
| tutorials/asr/01_ASR_with_NeMo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple example on using Instrumental Variables method for estimation
# +
import numpy as np
import pandas as pd
import patsy as ps
from statsmodels.sandbox.regression.gmm import IV2SLS
import os, sys
sys.path.append(os.path.abspath("../../../"))
from dowhy import CausalModel
# +
n_points = 1000
education_abilty = 1
education_voucher = 0.5
income_abilty = 2
income_education = 4
# confounder
ability = np.random.normal(0, 3, size=n_points)
# instrument
voucher = np.random.normal(2, 1, size=n_points)
# treatment
education = np.random.normal(5, 1, size=n_points) + education_abilty * ability +\
education_voucher * voucher
# outcome
income = np.random.normal(10, 3, size=n_points) +\
income_abilty * ability + income_education * education
# build dataset
data = np.stack([ability, education, income, voucher]).T
df = pd.DataFrame(data, columns = ['ability', 'education', 'income', 'voucher'])
# +
income_vec, endog = ps.dmatrices("income ~ education", data=df)
exog = ps.dmatrix("voucher", data=df)
m = IV2SLS(income_vec, endog, exog).fit()
m.summary()
# +
model=CausalModel(
data = df,
treatment='education',
outcome='income',
common_causes=['ability'],
instruments=['voucher']
)
identified_estimand = model.identify_effect()
estimate = model.estimate_effect(identified_estimand,
method_name="iv.instrumental_variable", test_significance=True
)
print(estimate)
| docs/source/example_notebooks/dowhy-simple-iv-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "-"}
# # Dynamic Tiling
#
# This notebook explores creating a series dynamicaly sized tiles out of a vector (rank-1 tensor).
#
# First, include some libraries
# +
# Run boilerplate code to set up environment
# %run ../prelude.py --style=tree --animation=movie
# -
# # Dynamic tiling class
#
# The following class is a generic class useful for tiling based on a computed criteria
#
class DynamicTiler():
""" DymamicTile
A callable class usable by the Fiber.prune() method to
create a tile based on a given lambda function that
decides if each element belongs in the tile
"""
def __init__(self, include_in_tile, max_tile_size=1):
""" __init__()
Initialize the class
Parameters
-----------
include_in_tile: function(coord, payload, cur_tile_size)
Function called to decide if each element should be in the tile
max_tile_size: integer
Maximum tile size
"""
self.include_in_tile = include_in_tile
self.max_tile_size = max_tile_size
self.cur_tile_size = 0
def __call__(self, n, c, p):
""" __call__()
Function callable by Fiber.prune() to decide what to do
with each element of a fiber. Either keep the element or
end the tile
Parameters:
n: integer
Position of element in tile
c: coordinate
Coordinate of element in tile
p: payload
Payload of elenent in tile
Return
------
Prune: Boolean | None
True - Include
False - Skip (never used)
None - End tile
"""
#
# Terminate the tile if it is already big enough
#
if self.cur_tile_size >= self.max_tile_size:
self.cur_tile_size = 0
return None
#
# Ask if we want another element in the tile
#
include = self.include_in_tile(c, p, self.cur_tile_size)
#
# Yep, add element to tile and increment the tile size
#
if include == True:
self.cur_tile_size += 1
return True
#
# End the tile
#
self.cur_tile_size = 0
return None
# ## Element selection function
#
# A function that randomly determines if an element of the original tensor is to be included in the current tile.
# +
import random
def random_tile(c, p, size):
""" random_tile
Decide if an element of a fiber should be included in
the current tile
Parameters
----------
c: coordinate
The coordinate of the element to consider for inclusion
p: payload
The payload of the element to conder for inclusion
size: integer
The number of elements already in the current tile
Return
-------
Include: Boolean
True means include this element; False means end tile
"""
#
# Tiles must have a least one element, but we
# get called in case this method has some internal
# state that gets updated
#
if size == 0:
return True
#
# Randomly decide if we want more elements
#
return random.random() > .2
# -
# ## Dynamically tile a tensor
# +
t = Tensor.fromRandom(["X"], [40], (0.6,), 10, seed=40)
t.setColor("green").setName("T")
displayTensor(t)
t_x1 = t.getRoot()
canvas = createCanvas(t)
tile_filter = DynamicTiler(random_tile, 4)
start_pos = 0
while start_pos < len(t_x1):
print(f"Tile starting at position: {start_pos}")
t_x0 = t_x1.prune(tile_filter, start_pos=start_pos)
start_pos = t_x1.getSavedPos()
# displayTensor(t_x0)
canvas.addFrame([(c,) for c,p in t_x0])
for c, p in t_x0:
print(f"c: {c}, p: {p}")
(count, distance) = t_x1.getSavedPosStats()
print("")
print(f"Average search distance: {distance/count:4.2}")
displayCanvas(canvas)
# -
# ## Testing area
#
# For running alternative algorithms
| notebooks/tiling/dynamic-tiling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit
# language: python
# name: python3
# ---
# ### 1. Import Libraries
import pandas as pd
from mazajak_api import predict
pd.set_option('display.max_columns', None)
pd.set_option('display.max_colwidth', None)
pd.set_option('display.max_rows', None)
pd.set_option('display.float_format', lambda x: '%.5f' % x)
# ### 2. Read the Data and Classify it
tweets = pd.read_csv('Tweets\\final_tweets_cleaned.csv')
tweets.columns
tweets = tweets.drop(['Unnamed: 0'], axis=1)
display(tweets.head())
display(tweets.tail())
# +
# Test run
tweet = 'كيفك'
print(predict(tweet))
tweets.shape[0]
# -
tweets.isna().sum()
tweets = tweets.dropna(subset=['Tweet'])
size = 50
chunks = [tweets[i:i+size].copy() for i in range(0, tweets.shape[0], size)]
no_chunks = len(chunks)
no_chunks
for i in range(0, no_chunks):
chunks[i]['Class'] = chunks[i]['Tweet'].map(predict)
print(f'__________ lap {i+1} of {no_chunks} __________')
# ### 3. Store the Classified Data
tweets = pd.concat(chunks)
display(tweets.head())
display(tweets.tail())
tweets = tweets.reset_index()
display(tweets.tail())
tweets.isna().sum()
tweets.to_csv('Tweets\\tweets_classified.csv')
| tweets-labeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import os
import numpy as np
import pickle
# %matplotlib inline
import seaborn as sb
sb.set()
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
X_train = pd.read_csv('C:/Users/<NAME>/Desktop/Prepared_Data/X_train.csv')
X_test = pd.read_csv('C:/Users/<NAME>/Desktop/Prepared_Data/X_test.csv')
Y_train = pd.read_csv('C:/Users/<NAME>/Desktop/Prepared_Data/Y_train.csv')
Y_test = pd.read_csv('C:/Users/<NAME>/Desktop/Prepared_Data/Y_test.csv')
# -
filename = 'C:\\Users\\<NAME>\\Desktop\\Model_V1.sav'
loaded_model = pickle.load(open(filename, 'rb'))
result = loaded_model.score(X_test, Y_test)
print(result*100)
# +
data = [[3, 800, 2, 0, 0, 0, 12, 10, 87.56826823, 1.542]]
predict = loaded_model.predict(data)
# -
predict[0]
# +
0, 24, 4, 0, 0, 9, 17, 15, 88.01383894, 0
data = [[0, 24, 4, 0, 0, 9, 17, 15, 88.01383894, 0]]
predict = loaded_model.predict(data)
predict[0]
# -
| Testing_V1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # After model.fit, before you deploy: Prototype with FastAPI in Jupyter!
#
# _By <NAME>, for JupyterCon 2020_
#
# You want to deploy your scikit-learn model. Now what? You can make an API for your model in Jupyter!
#
# You’ll learn [FastAPI](https://fastapi.tiangolo.com/), a Python web framework with automatic interactive docs. We’ll validate inputs with type hints, and convert to a dataframe, to make new predictions with your model. You’ll have a working API prototype, running from a notebook and ready to deploy!
#
# This talk is for people who feel comfortable in notebooks and can fit scikit-learn models. It’s about the technical process in-between developing your model and deploying it. Maybe you’ve never deployed an API before, or maybe you’ve tried Flask but you’re curious about FastAPI.
# ## Part 0, model.fit
# We'll use the [Palmer Penguins](https://github.com/allisonhorst/palmerpenguins) dataset. It's an alternative to [Iris](https://en.wikipedia.org/wiki/Iris_flower_data_set). Instead of using Iris flower measurements to predict one of three species, we'll use penguin measurements to predict one of three species.
# <img src="https://raw.githubusercontent.com/allisonhorst/palmerpenguins/master/man/figures/lter_penguins.png" width="50%" />
#
# Artwork by [@allison_horst](https://twitter.com/allison_horst)
# First, load and explore the data:
import seaborn as sns
penguins = sns.load_dataset('penguins')
sns.pairplot(data=penguins, hue='species')
# Looks like Adelie penguins have less bill length. Gentoo penguins have less bill depth, more flipper length, and more body mass.
#
# So we can classify the three species using two features: bill length and another numeric feature, such as bill depth.
# <img src="https://raw.githubusercontent.com/allisonhorst/palmerpenguins/master/man/figures/culmen_depth.png" width="50%" />
#
# Artwork by [@allison_horst](https://twitter.com/allison_horst)
# We'll select `bill_length_mm` and `bill_depth_mm` for our features, and `species` is our target. We'll use scikit-learn to fit a Logistic Regression model.
#
# Scikit-learn's implementation of Logistic Regression is regularized. We'll use cross-validation to automate the amount of regularization, after scaling the features. We can combine the scaler transformation and the model into a scikit-learn pipeline.
#
# We'll also use cross-validation to estimate how accurately the model generalizes.
# +
from sklearn.linear_model import LogisticRegressionCV
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import cross_val_score
from sklearn.pipeline import make_pipeline
features = ['bill_length_mm', 'bill_depth_mm']
target = 'species'
penguins.dropna(subset=features, inplace=True)
X = penguins[features]
y = penguins[target]
classifier = make_pipeline(
StandardScaler(),
LogisticRegressionCV()
)
classifier.fit(X, y)
scores = cross_val_score(classifier, X, y)
avg_acc = scores.mean() * 100
std_acc = scores.std() * 100
print(f'Cross-Validation Accuracy: {avg_acc:.0f}% +/- {2*std_acc:.0f}%')
# -
# So, our model seems to classify penguins nearly perfectly.
#
# Next, we'll deploy this model in a FastAPI app.
# Web apps aren't usually served from notebooks, especially temporary cloud notebooks like Binder. But it can be useful for rapid prototyping. Here's a helper function to make it possible:
def enable_cloud_notebook(port=8000):
"""
Enables you to run a FastAPI app from a cloud notebook.
Useful for rapid prototyping if you like notebooks!
Not needed when you develop in a local IDE or deploy "for real."
"""
# Prevent "RuntimeError: This event loop is already running"
import nest_asyncio
nest_asyncio.apply()
# Get a public URL to the localhost server
from pyngrok import ngrok
print('Public URL:', ngrok.connect(port=port))
# ## Part 1, random penguins, GET request
# Let's back up and begin with something like "Hello World." Before we make real predictions, we’ll make random guesses.
# +
import random
def random_penguin():
"""Return a random penguin species"""
return random.choice(['Adelie', 'Chinstrap', 'Gentoo'])
# -
# Run this function and you'll get random penguin species.
random_penguin()
# In the next cell, you'll see that we add a half-dozen lines of code to turn this function into a FastAPI app.
#
# These lines create a FastAPI app instance:
#
# ```python
# from fastapi import FastAPI
# app = FastAPI()
# ```
#
# This decorator tells FastAPI to call the function whenever the app receives a request to the `/` path using the HTTP GET method.
#
# ```python
# @app.get('/')
# def random_penguin():
# ...
# ```
#
# This line enables running FastAPI from a cloud notebook:
#
# ```python
# enable_cloud_notebook()
# ```
#
# These lines run the app with Uvicorn, the recommended web server for FastAPI:
#
# ```python
# import uvicorn
# uvicorn.run(app)
# ```
#
# The code below puts it all together. Run the cell. You'll see a "Public URL" that ends in "ngrok.io". Click the link to open it in a new tab. You'll see a random penguin species. Refresh the tab to get another random penguin.
# +
import random
from fastapi import FastAPI
import uvicorn
app = FastAPI()
@app.get('/')
def random_penguin():
"""Return a random penguin species"""
species = random.choice(['Adelie', 'Chinstrap', 'Gentoo'])
return species
enable_cloud_notebook()
uvicorn.run(app)
# -
# Every time you refresh you see it in the web logs above. The app is up on the public internet for anyone to access, but only while this cell in this notebook is running.
# In this notebook, stop the cell from running now.
# Next we'll add an app `title`, change the `docs_url` parameter, and change the path to `/random` for the `random_penguin` function.
#
# Run the cell and click the new Public URL.
# +
import random
from fastapi import FastAPI
import uvicorn
app = FastAPI(
title='🐧 Penguin predictor API',
docs_url='/'
)
@app.get('/random')
def random_penguin():
"""Return a random penguin species"""
species = random.choice(['Adelie', 'Chinstrap', 'Gentoo'])
return species
enable_cloud_notebook()
uvicorn.run(app)
# -
# Now you'll see automatically generated documentation. It's interactive too!
#
# Click on "/random", then the "Try It Out" button, then the "Execute" button. Scroll down to the "Server response." You'll see "Response body" with a penguin species, and "Code" with 200 which is a successful [status code](https://developer.mozilla.org/en-US/docs/Web/HTTP/Status).
# Or change the end of the URL to `/random` and you can use the API directly.
# You access your API with code like this, from another notebook or Python shell. Replace the url with your own dynamically generated ngrok URL.
#
# ```python
# import requests
# url = 'http://9571e5899f73.ngrok.io/random'
# response = requests.get(url)
# print(response.status_code, response.text)
# ```
# Then stop the cell above from running like before.
# ## Part 2, real predictions, POST request
# Okay, now let's work on adding our model to make real predictions.
# To make a prediction, we need penguin measurements, which we'll receive as [JSON](https://developer.mozilla.org/en-US/docs/Learn/JavaScript/Objects/JSON):
#
# > JavaScript Object Notation (JSON) is a standard text-based format for representing structured data based on JavaScript object syntax. It is commonly used for transmitting data in web applications (e.g., sending some data from the server to the client, so it can be displayed on a web page, or vice versa). You'll come across it quite often ... it can be used independently from JavaScript, and many programming environments feature the ability to read (parse) and generate JSON.
# JSON looks a lot like a Python dictionary, like this example:
gary_gentoo = {"bill_length_mm": 45, "bill_depth_mm": 15}
# How do we go from JSON / dictionary format to something our model can use?
#
# We need a Numpy array or a Pandas dataframe, with two columns (for our two features) and one row (for our one observation that we want to predict). We can make a dataframe from a list of dicts, like this:
import pandas as pd
pd.DataFrame([gary_gentoo])
# When we use this dataframe with our classifier's predict method, we get the correct result.
import pandas as pd
df = pd.DataFrame([gary_gentoo])
classifier.predict(df)
# The predict method returns a Numpy array with all our predictions. But we're just making a single prediction, so we want the "zeroeth" item from the array. Putting it all together, we could write a function like this:
def predict_species(penguin: dict):
"""Predict penguin species"""
df = pd.DataFrame([penguin])
species = classifier.predict(df)
return species[0]
predict_species(gary_gentoo)
# Here's another example.
amy_adelie = {"bill_length_mm": 35, "bill_depth_mm": 18}
predict_species(amy_adelie)
# We'll add the function to our FastAPI app using a decorator. The decorator tells FastAPI to call the function whenever the app receives a request to the `/predict` path using the [HTTP POST method](https://developer.mozilla.org/en-US/docs/Web/HTTP/Methods/POST). FastAPI will automatically parse the request body's JSON to a Python dict named `penguin`.
#
# ```python
# @app.post('/predict')
# def predict_species(penguin: dict):
# ...
# ```
#
# We'll also add a more descriptive `description` parameter to the app. Putting it all together:
# +
import random
from fastapi import FastAPI
import pandas as pd
import uvicorn
app = FastAPI(
title='🐧 Penguin predictor API',
description='Deploys a logistic regression model fit on the [Palmer Penguins](https://github.com/allisonhorst/palmerpenguins) dataset.',
docs_url='/'
)
@app.post('/predict')
def predict_species(penguin: dict):
"""Predict penguin species"""
df = pd.DataFrame([penguin])
species = classifier.predict(df)
return species[0]
@app.get('/random')
def random_penguin():
"""Return a random penguin species"""
species = random.choice(['Adelie', 'Chinstrap', 'Gentoo'])
return species
enable_cloud_notebook()
uvicorn.run(app)
# -
#
# Run the cell above, then try an example:
#
# - Click the "Try it out" button.
# - The "Request body" text field becomes editable. Copy-paste <NAME>'s measurements into the field: `{"bill_length_mm": 45, "bill_depth_mm": 15}`
# - Click the "Execute" button, then scroll down to the "Server response." You should see the species "Gentoo" correctly classified.
#
# Try another example:
#
# - Copy-paste <NAME>'s measurements into the "Request body" text field: `{"bill_length_mm": 35, "bill_depth_mm": 18}`
# - Click the "Execute" button. You should see the species "Adelie" correctly classified.
#
# But what happens if you change your "Request body" to something unexpected?
# - What if your input doesn't have exactly two keys, `bill_length_mm` and `bill_depth_mm`, in that order?
# - What if your input values are zero? Huge numbers? Negative numbers? Not a number?
#
# We aren't validating input yet. We just assume the API users give valid input. That's a dangerous assumption. When the inputs aren't valid, the app may respond with a Server Error instead of helpful warnings. Or worse, the app seems to work and returns a response, but because the inputs were flawed, the output is flawed too. "Garbage in, garbage out."
# Stop the cell above from running. Next we'll add data validation.
# ## Part 3, Data validation
# Look at the type annotation for the `predict_species` function's argument. The function accepts any `dict`.
#
# ```python
# @app.post('/predict')
# def predict_species(penguin: dict):
# ...
# ```
#
# We'll change this so the function expects an argument of type `Penguin`.
#
# ```python
# class Penguin:
# """Parse & validate penguin measurements"""
# ...
#
# @app.post('/predict')
# def predict_species(penguin: Penguin):
# ...
# ```
# We'll create a `Penguin` [data class](https://docs.python.org/3/library/dataclasses.html) with [type annotations](https://docs.python.org/3/library/typing.html) to define what attributes we expect our input to have. We'll use [Pydantic](https://pydantic-docs.helpmanual.io/), a data validation library integrated with FastAPI. It sounds complex, but it's just a few lines of code!
# +
from pydantic import BaseModel
class Penguin(BaseModel):
"""Parse & validate penguin measurements"""
bill_length_mm: float
bill_depth_mm: float
# -
# We can instantiate a penguin object like this:
Penguin(bill_length_mm=45, bill_depth_mm=15)
# Or like this, by unpacking our dictionary into parameters:
Penguin(**gary_gentoo)
# Now let's see what happens with missing input:
missing_input = {"bill_length_mm": 45}
Penguin(**missing_input)
# We automatically get a `ValidationError` with a helpful, descriptive error message. That's what we want in this situation!
# Next we'll try a misnamed input (`bill_depth` instead of `bill_depth_mm`)
wrong_name = {"bill_length_mm": 45, "bill_depth": 15}
Penguin(**wrong_name)
# Again, we get a `ValidationError`, which is want we want here.
# Let's try an input with the wrong type, such as a string instead of a number.
wrong_type = {"bill_length_mm": 45, "bill_depth_mm": "Hello Penguins!"}
Penguin(**wrong_type)
# We get a different `ValidationError` because the value is not a valid float.
# Let's try a different string:
convertable_type = {"bill_length_mm": 45, "bill_depth_mm": "15"}
Penguin(**convertable_type)
# This works because the string can be converted to a float.
# If we add an extra input ...
extra_input = {"bill_length_mm": 45, "bill_depth_mm": 15, "extra_feature": "will be ignored"}
Penguin(**extra_input)
# ... it will be ignored.
# If we flip the order of inputs ...
flipped_order = {"bill_depth_mm": 15, "bill_length_mm": 45}
Penguin(**flipped_order)
# ... they'll be flipped back.
# What about penguin measurements that are implausibly large or small? We can use "constrained floats" to catch this.
#
# We'll set constraints that each input must be greater than (`gt`) some minimum and less than (`lt`) some maximum.
from pydantic import confloat
help(confloat)
# First, let's look at the minimum and maximum measurements from our training data:
X.describe()
# Then, set some reasonable constraints:
# +
from pydantic import BaseModel, confloat
class Penguin(BaseModel):
"""Parse & validate penguin measurements"""
bill_length_mm: confloat(gt=32, lt=60)
bill_depth_mm: confloat(gt=13, lt=22)
# -
# Now when inputs are too large or small, we get a `ValidationError` with descriptive messages.
huge_penguin = {"bill_depth_mm": 1500, "bill_length_mm": 4500}
Penguin(**huge_penguin)
zero_penguin = {"bill_depth_mm": 0, "bill_length_mm": 0}
Penguin(**zero_penguin)
negative_penguin = {"bill_depth_mm": -45, "bill_length_mm": -15}
Penguin(**negative_penguin)
# One more thing. Let's add a helpful method to our class:
# +
from pydantic import BaseModel, confloat
class Penguin(BaseModel):
"""Parse & validate penguin measurements"""
bill_length_mm: confloat(gt=32, lt=60)
bill_depth_mm: confloat(gt=13, lt=22)
def to_df(self):
"""Convert to pandas dataframe with 1 row."""
return pd.DataFrame([dict(self)])
# -
# Now we can validate JSON input and convert it to a pandas dataframe with one line of code.
Penguin(**gary_gentoo).to_df()
# Let's put this all together in our FastAPI code.
#
# - Add the `Penguin` class.
# - Change the type annotation for the `predict_species` function argument. Instead of `dict`, the type is now `Penguin`.
# - When a POST request is made to the `/predict` path, then FastAPI will automatically validate and parse the request body's JSON into a `Penguin` object.
# - Use the penguin's `to_df` method to convert into a dataframe for our model.
# +
import random
from fastapi import FastAPI
import pandas as pd
from pydantic import BaseModel, confloat
import uvicorn
app = FastAPI(
title='🐧 Penguin predictor API',
description='Deploys a logistic regression model fit on the [Palmer Penguins](https://github.com/allisonhorst/palmerpenguins) dataset.',
docs_url='/'
)
class Penguin(BaseModel):
"""Parse & validate penguin measurements"""
bill_length_mm: confloat(gt=32, lt=60)
bill_depth_mm: confloat(gt=13, lt=22)
def to_df(self):
"""Convert to pandas dataframe with 1 row."""
return pd.DataFrame([dict(self)])
@app.post('/predict')
def predict_species(penguin: Penguin):
"""Predict penguin species from bill length & depth
Parameters
----------
bill_length_mm : float, greater than 32, less than 60
bill_depth_mm : float, greater than 13, less than 22
Returns
-------
str "Adelie", "Chinstrap", or "Gentoo"
"""
species = classifier.predict(penguin.to_df())
return species[0]
@app.get('/random')
def random_penguin():
"""Return a random penguin species"""
species = random.choice(['Adelie', 'Chinstrap', 'Gentoo'])
return species
enable_cloud_notebook()
uvicorn.run(app)
# -
# Test the app, then stop the cell from running.
# ## Part -1, Deploy
# Let's save the model so you can use it without retraining. This is sometimes called "pickling." See [scikit-learn docs on "model persistence."](https://scikit-learn.org/stable/modules/model_persistence.html)
from joblib import dump
dump(classifier, 'classifier.joblib', compress=True)
# Now even if we delete the object from memory ...
del classifier
# We can reload from our file ...
from joblib import load
classifier = load('classifier.joblib')
# ... and it's back, ready to use:
from sklearn import set_config
set_config(display='diagram')
classifier
# If you're using a cloud notebook, you can get a link to download the file using code like this:
from IPython.display import FileLink
FileLink('classifier.joblib')
# This last code cell has 3 changes from the previous iteration:
#
# - Loads the model with joblib
# - Adds image HTML tags in the app's description
# - Configures [CORS (Cross-Origin Resource Sharing)](https://fastapi.tiangolo.com/tutorial/cors/) so your API could be called by apps on different domains.
# +
import random
from fastapi import FastAPI
from fastapi.middleware.cors import CORSMiddleware
from joblib import load
import pandas as pd
from pydantic import BaseModel, confloat
import uvicorn
description = """
Deploys a logistic regression model fit on the [Palmer Penguins](https://github.com/allisonhorst/palmerpenguins) dataset.
<img src="https://raw.githubusercontent.com/allisonhorst/palmerpenguins/master/man/figures/lter_penguins.png" width="40%" /> <img src="https://raw.githubusercontent.com/allisonhorst/palmerpenguins/master/man/figures/culmen_depth.png" width="30%" />
Artwork by [@allison_horst](https://twitter.com/allison_horst)
"""
app = FastAPI(
title='🐧 Penguin predictor API',
description=description,
docs_url='/'
)
app.add_middleware(
CORSMiddleware,
allow_origins=['*'],
allow_methods=['*']
)
classifier = load('classifier.joblib')
class Penguin(BaseModel):
"""Parse & validate penguin measurements"""
bill_length_mm: confloat(gt=32, lt=60)
bill_depth_mm: confloat(gt=13, lt=22)
def to_df(self):
"""Convert to pandas dataframe with 1 row."""
return pd.DataFrame([dict(self)])
@app.post('/predict')
def predict_species(penguin: Penguin):
"""Predict penguin species from bill length & depth
Parameters
----------
bill_length_mm : float, greater than 32, less than 60
bill_depth_mm : float, greater than 13, less than 22
Returns
-------
str "Adelie", "Chinstrap", or "Gentoo"
"""
species = classifier.predict(penguin.to_df())
return species[0]
@app.get('/random')
def random_penguin():
"""Return a random penguin species"""
species = random.choice(['Adelie', 'Chinstrap', 'Gentoo'])
return species
enable_cloud_notebook()
uvicorn.run(app)
# -
# We've prototyped a complete working web app, running from a notebook. We're ready to deploy!
#
# Do you want to take this last step and go beyond the notebook? See the README in this repo for instructions how to deploy to Heroku, a popular cloud platform.
# ## Learn more
#
# Want to learn more about FastAPI? I recommend these links:
#
# - [Build a machine learning API from scratch](https://youtu.be/1zMQBe0l1bM) by <NAME>, FastAPI's creator
# - [calmcode.io — FastAPI videos](https://calmcode.io/fastapi/hello-world.html) by <NAME>
# - [FastAPI for Flask Users](https://amitness.com/2020/06/fastapi-vs-flask/) by <NAME>
# - [FastAPI official docs](https://fastapi.tiangolo.com/)
# - [testdriven.io — FastAPI blog posts](https://testdriven.io/blog/topics/fastapi/)
| notebook/Prototype with FastAPI in Jupyter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="0SIvr26kzpoH" colab_type="text"
# # A denoising baseline implementation of a residual dense network trained on CIFAR-100 with added Gaussian noise
# + [markdown] id="4-EsDDPszlIm" colab_type="text"
# ## Setup
# + [markdown] id="4Tu2wErUz2pY" colab_type="text"
# ### Imports
# + id="rtQppKyD0HzW" colab_type="code" colab={}
# !pip install -q --upgrade gsheet-keyring ipython-secrets comet_ml scikit-image imgaug torch torchvision tqdm
# + id="kHQpQfOVzdjx" colab_type="code" colab={}
from comet_ml import Experiment
import torch
import argparse
from torch import nn
from torch import optim
import torchvision.datasets as datasets
import torchvision.transforms as transforms
from ipython_secrets import get_secret
from torch.utils.data.dataset import Dataset
from torch.utils.data import DataLoader
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
sns.set()
from tqdm import tqdm as tqdm_base
def tqdm(*args, **kwargs):
if hasattr(tqdm_base, '_instances'):
for instance in list(tqdm_base._instances):
tqdm_base._decr_instances(instance)
return tqdm_base(*args, **kwargs)
# + id="vVlNa_bkWyVq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="60083ccd-2da2-4dc7-fadc-6bff9dcea13f" language="javascript"
# IPython.OutputArea.auto_scroll_threshold = 99999;
# + [markdown] id="SY5GL4qTz4ig" colab_type="text"
# ### Environment
# + id="yMy_U2sOz_HQ" colab_type="code" outputId="1e3d1567-0389-4931-a657-4a1f92951dc2" colab={"base_uri": "https://localhost:8080/", "height": 51}
PROJECT = "fastrino"
DEVICE = "cuda"
DATA_PATH = "./data"
COMET_ML_API_KEY = get_secret("comet-{}".format(PROJECT))
cifar100_train_data = datasets.CIFAR100(
root=DATA_PATH, train=True, download=True, transform=transforms.ToTensor()
)
cifar100_test_data = datasets.CIFAR100(
root=DATA_PATH, train=False, download=True, transform=transforms.ToTensor()
)
# + [markdown] id="kleeBt1G0dbz" colab_type="text"
# ### Utilities
# + id="goX6CbN20cdS" colab_type="code" colab={}
class NoisyCIFAR100(Dataset):
def __init__(self, original_data, noise_mean=0, noise_std=1):
self.original_data = original_data
self.noise_mean = noise_mean
self.noise_std = noise_std
def __len__(self):
return len(self.original_data)
def __getitem__(self, idx):
image, _ = self.original_data[idx]
noise = self.noise_mean + self.noise_std * torch.randn_like(
image
)
return image + noise, noise
def load_noisy_data(noise_mean=0, noise_std=1, batch_size=32):
train_data = NoisyCIFAR100(cifar100_train_data, noise_mean, noise_std)
test_data = NoisyCIFAR100(cifar100_test_data, noise_mean, noise_std)
train_loader = DataLoader(train_data, batch_size=batch_size, shuffle=True)
test_loader = DataLoader(test_data, batch_size=batch_size, shuffle=False)
return train_loader, test_loader
def get_arg_parser():
parser = argparse.ArgumentParser()
parser.add_argument("--num-channels", type=int, default=3)
parser.add_argument("--num-features", type=int, default=16)
parser.add_argument("--growth-rate", type=int, default=16)
parser.add_argument("--num-blocks", type=int, default=3)
parser.add_argument("--num-layers", type=int, default=5)
parser.add_argument("--kernel-size", type=int, default=3)
parser.add_argument("--lr", type=float, default=1e-4)
parser.add_argument("--batch-size", type=int, default=128)
parser.add_argument("--noise-mean", type=float, default=0)
parser.add_argument("--noise-std", type=int, default=0.1)
parser.add_argument("--num-epochs", type=int, default=5)
parser.add_argument("--seed", type=int, default=42)
return parser
def get_criterion():
return nn.L1Loss()
def get_optimizer(model, lr=0.001):
return optim.Adam(model.parameters(), lr)
def compute_psnr(prediction, target, max_pixel=255.0):
return (
10.0 * ((max_pixel ** 2) / ((prediction - target) ** 2).mean()).log10()
)
def train(noise_mean=0, noise_std=0.01, device=DEVICE):
parser = get_arg_parser()
args = parser.parse_args(args=[])
args.noise_mean = noise_mean
args.noise_std = noise_std
train_loader, test_loader = load_noisy_data(
args.noise_mean, args.noise_std, args.batch_size
)
experiment = Experiment(
api_key=COMET_ML_API_KEY,
project_name=PROJECT,
workspace=PROJECT,
auto_output_logging=None,
)
experiment.log_parameters(vars(args))
model = ResidualDenseNetwork(
args.num_channels,
args.num_features,
args.growth_rate,
args.num_blocks,
args.num_layers,
args.kernel_size,
).to(device)
criterion = get_criterion()
optimizer = get_optimizer(model, args.lr)
for epoch in tqdm(
range(args.num_epochs), desc="Epoch", unit="epochs"
):
with experiment.train():
model.train()
train_psnr = []
for image, noise in tqdm(
train_loader, desc="Train images", unit="images"
):
prediction = model(image.to(device))
loss = criterion(prediction, noise.to(device))
loss.backward()
optimizer.step()
optimizer.zero_grad()
current_psnr = compute_psnr(prediction, noise.to(device)).data.item()
train_psnr.append(current_psnr)
experiment.log_metric("psnr", current_psnr)
experiment.log_metric("loss", loss.data.item())
experiment.log_metric("mean_psnr", np.mean(train_psnr))
with experiment.test():
model.eval()
test_losses = []
test_psnr = []
for image, noise in test_loader:
prediction = model(image.to(device))
test_psnr.append(compute_psnr(prediction, noise.to(device)).data.item())
test_losses.append(
criterion(prediction, noise.to(device)).data.item()
)
test_psnr = np.mean(train_psnr)
test_loss = np.mean(test_losses)
experiment.log_metric("mean_psnr", np.mean(train_psnr))
experiment.log_metric("mean_loss", np.mean(test_losses))
return test_psnr, test_loss
# + [markdown] id="9lnFZPGMz63O" colab_type="text"
# ## Model
# + id="tEHBSWnRzWFE" colab_type="code" colab={}
# Adapted from the PyTorch implementation of Residual Dense Network
# for Image Super-Resolution by @yjn870,
# https://github.com/yjn870/RDN-pytorch
class DenseLayer(nn.Module):
def __init__(self, in_channels, out_channels, kernel_size=3):
super().__init__()
self.conv = nn.Conv2d(
in_channels,
out_channels,
kernel_size=kernel_size,
padding=(kernel_size - 1) // 2,
)
self.relu = nn.ReLU(inplace=True)
def forward(self, x):
return torch.cat([x, self.relu(self.conv(x))], 1)
class ResidualDenseBlock(nn.Module):
def __init__(self, in_channels, growth_rate, num_layers):
super().__init__()
self.dense_layers = nn.Sequential(
*[
DenseLayer(in_channels + growth_rate * layer_idx, growth_rate)
for layer_idx in range(num_layers)
]
)
self.local_feature_fusion = nn.Conv2d(
in_channels + growth_rate * num_layers, growth_rate, kernel_size=1
)
def forward(self, x):
return x + self.local_feature_fusion(self.dense_layers(x))
class ResidualDenseNetwork(nn.Module):
def __init__(
self,
num_channels,
num_features,
growth_rate,
num_blocks,
num_layers,
kernel_size=3,
):
super().__init__()
self.growth_rate = growth_rate
self.num_features = num_features
self.num_blocks = num_blocks
self.num_layers = num_layers
self.outer_shallow_features = nn.Conv2d(
num_channels,
num_features,
kernel_size=kernel_size,
padding=(kernel_size - 1) // 2,
)
self.inner_shallow_features = nn.Conv2d(
num_features,
num_features,
kernel_size=kernel_size,
padding=(kernel_size - 1) // 2,
)
self.residual_dense_blocks = nn.ModuleList(
[
ResidualDenseBlock(
self.num_features, self.growth_rate, self.num_layers
)
]
)
for _ in range(self.num_blocks - 1):
self.residual_dense_blocks.append(
ResidualDenseBlock(
self.growth_rate, self.growth_rate, self.num_layers
)
)
self.global_feature_fusion = nn.Sequential(
nn.Conv2d(
self.growth_rate * self.num_blocks,
self.num_features,
kernel_size=1,
),
nn.Conv2d(
self.num_features, self.num_features, kernel_size=3, padding=1,
),
)
self.output = nn.Conv2d(
self.num_features,
num_channels,
kernel_size=kernel_size,
padding=(kernel_size - 1) // 2,
)
def forward(self, x):
outer_shallow_features = self.outer_shallow_features(x)
x = self.inner_shallow_features(outer_shallow_features)
local_features = []
for block_idx in range(self.num_blocks):
x = self.residual_dense_blocks[block_idx](x)
local_features.append(x)
x = (
self.global_feature_fusion(torch.cat(local_features, 1))
+ outer_shallow_features
)
x = self.output(x)
return x
# + [markdown] id="0MAlE9_OeNwD" colab_type="text"
# ## Training
# + id="-OFs4Sc6EKjM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="089867f5-cf02-44ee-dac6-4d9157b3f6e3"
noise_mean = np.linspace(0.01, 0.5, 3)
noise_std = np.linspace(0.01, 0.5, 3)
index = pd.MultiIndex.from_product(
[noise_mean, noise_std],
names = ["noise_mean", "noise_std"]
)
results = pd.DataFrame(index = index).reset_index()
results["test_psnr"] = 0.0
results["test_loss"] = 0.0
results.head()
# + id="RB_P0vpBFL2T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="228da9cf-0474-4464-e206-e29f344e14e6"
for (idx, row) in results.iterrows():
noise_mean, noise_std, _, _ = row
test_psnr, test_loss = train(noise_mean, noise_std)
row[2] = test_psnr
row[3] = test_loss
# + id="hTtWSGNEHEsw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 328} outputId="4fcda5be-811c-414b-fc6e-e8c7e37b464c"
results
# + id="OIPyJDVmTN_T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 916} outputId="efd7b257-888a-4dd2-cf47-fe448d37ec1a"
plt.figure(dpi=120, figsize=(9, 9))
plt.subplot(2, 2, 1)
sns.scatterplot(x=results["noise_mean"], hue=results["noise_std"], y=results["test_loss"])
plt.subplot(2, 2, 2)
sns.scatterplot(x=results["noise_std"], hue=results["noise_mean"], y=results["test_loss"])
plt.subplot(2, 2, 3)
sns.scatterplot(x=results["noise_mean"], hue=results["noise_std"], y=results["test_psnr"])
plt.subplot(2, 2, 4)
sns.scatterplot(x=results["noise_std"], hue=results["noise_mean"], y=results["test_psnr"])
plt.show()
# + id="MVyXJCegUBe4" colab_type="code" colab={}
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import datetime as dt
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from viresclient import SwarmRequest
import chaosmagpy as cp # v0.1
# -
# # Get a day of model evaluations from VirES
# # 01/01/2017
# +
request = SwarmRequest()
request.set_collection("SW_OPER_MAGA_LR_1B")
request.set_products(measurements=["B_NEC", "F"],
models=[
"CHAOS-6-Core",
"CHAOS-6-Static",
"CHAOS-6-Combined",
"CHAOS-6-MMA-Primary",
"CHAOS-6-MMA-Secondary",
],
sampling_step="PT30S")
ds = request.get_between(
start_time=dt.datetime(2017,1,1),
end_time=dt.datetime(2017,1,2)
).as_xarray()
ds
# -
# # Functions to help use chaosmagpy
# Get the CHAOS model .mat file at http://www.spacecenter.dk/files/magnetic-models/CHAOS-6/
#
# `!wget http://www.spacecenter.dk/files/magnetic-models/CHAOS-6/CHAOS-6-x9.mat`
chaos_model = cp.load_CHAOS_matfile('CHAOS-6-x9.mat')
# +
def datetimeindex_to_mjd2000(dti):
"""Convert a pandas datetime index to MJD2000
Convert an array to a dti like this:
pd.core.indexes.datetimes.DatetimeIndex(np.array([dt.datetime(2017,1,1),]))
"""
if type(dti) is not pd.core.indexes.datetimes.DatetimeIndex:
raise Exception("Input should be a pandas DatetimeIndex")
return dti.to_julian_date() - 2400000.5 - 51544
def rtp2NEC(B_rtp):
"""Change an array from rtp to NEC frame
"""
if B_rtp.shape[1] != 3:
raise Exception("B_rtp is not the right shape")
return np.stack((-B_rtp[:, 1], B_rtp[:, 2], -B_rtp[:, 0])).T
def get_chaos_tdep_NEC(times_mjd2000, radius, theta, phi, nmax=20):
coeffs = chaos_model.synth_coeffs_tdep(
nmax=nmax,
time=times_mjd2000
)
B_radius, B_theta, B_phi = cp.model_utils.synth_values(
coeffs, radius, theta, phi, nmax=nmax, source='internal'
)
B_NEC = rtp2NEC(np.stack((B_radius, B_theta, B_phi)).T)
return B_NEC
def get_chaos_static_NEC(times_mjd2000, radius, theta, phi):
coeffs_static = chaos_model.synth_coeffs_static()
B_radius, B_theta, B_phi = cp.model_utils.synth_values(
coeffs_static, radius, theta, phi
)
B_NEC = rtp2NEC(np.stack((B_radius, B_theta, B_phi)).T)
return B_NEC
def get_chaos_mma_NEC(times_mjd2000=None, radius=None, theta=None, phi=None,
source=None):
B_radius = np.zeros(times_mjd2000.shape)
B_theta = np.zeros(times_mjd2000.shape)
B_phi = np.zeros(times_mjd2000.shape)
for synth_coeffs in (chaos_model.synth_coeffs_sm, chaos_model.synth_coeffs_gsm):
coeffs = synth_coeffs(times_mjd2000, source=source)
B_r, B_t, B_p = cp.model_utils.synth_values(
coeffs, radius, theta, phi, source=source
)
B_radius += B_r
B_theta += B_t
B_phi += B_p
return rtp2NEC(np.stack((B_radius, B_theta, B_phi)).T)
# -
# # Use chaosmagpy to evaluate CHAOS models and add them to the dataset
# +
times_mjd2000 = datetimeindex_to_mjd2000(ds.indexes["Timestamp"]).values
theta = 90 - ds["Latitude"]
phi = ds["Longitude"] % 360
radius = ds["Radius"] / 1e3
ds["B_NEC_CHAOS-6-Core-PY"] = (
("Timestamp","dim"),
get_chaos_tdep_NEC(times_mjd2000, radius, theta, phi)
)
ds["B_NEC_CHAOS-6-Static-PY"] = (
("Timestamp","dim"),
get_chaos_static_NEC(times_mjd2000, radius, theta, phi)
)
ds["B_NEC_CHAOS-6-MMA-Primary-PY"] = (
("Timestamp","dim"),
get_chaos_mma_NEC(times_mjd2000, radius, theta, phi, "external")
)
ds["B_NEC_CHAOS-6-MMA-Secondary-PY"] = (
("Timestamp","dim"),
get_chaos_mma_NEC(times_mjd2000, radius, theta, phi, "internal")
)
ds["B_NEC_CHAOS-6-MMA"] = ds["B_NEC_CHAOS-6-MMA-Primary"] + ds["B_NEC_CHAOS-6-MMA-Secondary"]
ds["B_NEC_CHAOS-6-MMA-PY"] = ds["B_NEC_CHAOS-6-MMA-Primary-PY"] + ds["B_NEC_CHAOS-6-MMA-Secondary-PY"]
ds
# -
# # Plot the difference between model values from chaosmagpy and from VirES
# +
plt.rcParams["font.size"] = 20
for mod in ("CHAOS-6-Core","CHAOS-6-Static", "CHAOS-6-MMA-Primary", "CHAOS-6-MMA-Secondary", "CHAOS-6-MMA"):
plt.figure(figsize=(20,5))
for i, label in enumerate(["dB$_N$", "dB$_E$", "dB$_C$"]):
(ds[f"B_NEC_{mod}-PY"] - ds[f"B_NEC_{mod}"])[:,i].plot(label=label, linewidth=1)
plt.title(mod)
plt.ylabel("Difference,\nchaosmagpy-VirES\n[nT]")
plt.legend(loc="lower right"); plt.grid();
| chaosmagpy_vs_vires.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import unittest
import os
try:
import cPickle as pickle
except:
import pickle
path = os.getcwd()
path = path[:-6]
sep = os.path.sep
# Load a sample file to use for testing
comps = pickle.load(open(path + '%ssample_data_files%sunittest_comparisons.pkl'
% (sep, sep), 'rb'))
import sys
from bokeh.plotting import output_notebook
lib_path = os.path.abspath(os.path.join('..'))
sys.path.append(lib_path)
import interactive_plots as ip
output_notebook()
sa_dict = comps[0]
p = ip.interact_with_plot_all_outputs(sa_dict, demo=True, manual=False)
class TestInteractWithPlotAllOutputs(unittest.TestCase):
def test_interact_with_plot_all_outputs_all_widgets_appear(self):
"""
Are all widgets appearing and are they in right order?
"""
l = p.widget.children
self.assertEqual(len(l), 6)
array_of_widget_names = []
for i in range(len(l)):
widgets = p.widget.children[i]
array_of_widget_names.append(str(widgets.class_own_traits.im_self)
.split('.')[-1].strip('\'>'))
self.assertEqual(array_of_widget_names, ['BoundedFloatText',
'IntText',
'Checkbox',
'Checkbox',
'Checkbox',
'SelectMultiple'])
def test_interact_with_plot_all_outputs_default_values(self):
"""
Are interactive widgets working properly and have proper
default values??
"""
self.assertEqual(p.widget.children[0].value, 0.01)
self.assertEqual(p.widget.children[1].value, 20.0)
self.assertEqual(p.widget.children[2].value, True)
self.assertEqual(p.widget.children[3].value, True)
self.assertEqual(p.widget.children[4].value, True)
def test_interact_with_plot_all_outputs_plots_the_glyphs(self):
"""
Are right number of glyphs showing up after calling interactive
ipywidgets?
"""
# This test confirms that the bokeh plots are generated
# as glyphs can only be seen if plots are generated
# when top 20 values are plotted number of glyphs = 12
figure = p.widget.result.doc.roots[0]
glyph_dict = figure.renderers
# total number of glyphs = 12 when default values are set
self.assertEquals(len(glyph_dict),12)
def test_interact_with_plot_all_outputs_error_bars_working(self):
"""
Are right number of glyphs showing up after calling interactive
ipywidgets?
"""
# when error bars are removed
# number of glyphs = 15
# q is the show error bars widget
q = p.widget.children[3]
q.value = False
# n is the number of tabs after 2 runs of interact_with__all_plots
# n = 4 after q.value is changed as it plots everything again
n = p.widget.result.doc.roots
glyph_dict_new = n[3].renderers
# since error bars are removed number of glyphs now become 9
self.assertEquals(len(glyph_dict_new),9)
def test_interact_with_plot_all_outputs_top_values_working(self):
"""
Are right number of glyphs showing up after calling interactive
ipywidgets
"""
# when error bars are removed
# number of glyphs = 15
# r is the show top widget for r = 1
# the graph is histogram and has 7 glyphs
r = p.widget.children[1]
r.value = 1
# n is the number of tabs after 3 runs of interact_with__all_plots
# n = 6 after q.value is changed as it plots everything again
n = p.widget.result.doc.roots
glyph_dict_new = n[5].renderers
# for histograms the number of glyphs now become 7
# when show top: value is 1
self.assertEquals(len(glyph_dict_new),7)
# Note other tests to be written to check interactivity of other widgets
suite = unittest.TestLoader().loadTestsFromTestCase(TestInteractWithPlotAllOutputs)
unittest.TextTestRunner(verbosity=20,stream=sys.stderr).run(suite)
# -
| savvy/tests/interactive_plots_unittests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
import random
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
import pandas as pd
import math
# %matplotlib inline
# + [markdown] deletable=true editable=true
# ## Gaussian distribution
# ### Generation sample by rolling dices many times
# + deletable=true editable=true
def rollDice():
return random.randint(1, 6)
def generate():
N = 10000
trials = {}
for i in range(N):
d1 = rollDice()
d2 = rollDice()
s = d1 + d2
trials[s] = trials.get(s, 0) + 1
return trials
# + deletable=true editable=true
data = generate()
plt.bar(list(data.keys()), list(data.values()))
plt.xticks(range(1, 12))
plt.show()
# + [markdown] deletable=true editable=true
# ## Cumulative Distribution
# + deletable=true editable=true
def calcCumulativeDist(data):
s = {}
for i, v in enumerate(data.values()):
s[i] = s.get(i - 1, 0) + v
return s
def takeSamples(rangeFrom, rangeTo, n):
return [random.randint(rangeFrom, rangeTo) for i in range(n)]
cumulativDist = calcCumulativeDist(data)
plt.plot(list(cumulativDist.keys()), list(cumulativDist.values()))
plt.show()
# -
# # Parameter Estimation
# We try to find mean and variance using given data points.
# + deletable=true editable=true
data = generate()
data
# -
# ### 1. Finding mean
# Mean is simply the average of data points. N is the number of data points.
# $$
# \mu = \frac{1}{N} \sum_{i}^{N} x_{i} \\
# $$
N = np.sum(list(data.values()))
m = sum(k*v for k, v in data.items()) / N
m
# ### 2. Finding variance
# To find the variance we sum the squared differences from the mean. Finally we take the average of the sum.
# $$
# \sigma^{2} = \frac{1}{N} \sum_{i}^{N}(x_{i} - \mu)^{2} \\
# $$
variance = sum(np.power(k - m, 2) for k, v in data.items() for i in range(v)) / N
variance
# # Gaussian (Normal) distribution
# Say $ X \in \mathbb{R} $ if `x` is distirbuted Gaussian with mean $\mu$, variance $\sigma^{2}$.
# $$
# X \sim \mathcal{N}(\mu,\,\sigma^{2})
# $$
#
# $$
# p(x, \mu, \sigma^{2}) = \frac{1}{\sqrt{2\pi } \ \sigma} exp(-\frac{(x-\mu)^{2}}{2\sigma^{2}})
# $$
def calc_p(x, m, variance):
return 1/ np.sqrt(2*np.pi * np.sqrt(variance)) * np.exp(- np.power(x-m, 2) / 2*variance)
# +
ps = dict((i, calc_p(i, m, variance)) for i in np.linspace(1, 12, 120))
plt.plot(list(ps.keys()), list(ps.values()))
plt.xticks(range(1, 13))
plt.show()
# -
dict((i, calc_p(i, m, variance)) for i in range(1, 13))
# ### Density Estimation & Cumulative Density Function
# What is the probability of rolling two dice whose sum is between 2 and 8? Cumulative distribution function is as follows:
# $$
# \phi(x, \mu, \sigma) = \frac{1}{2} \ (1 + erf( \frac{x-\mu}{\sigma \sqrt{2}} ) )
# $$
def cumulativeCDF(x, m, stdv):
return 0.5 * (1 + math.erf((x-m)/ (stdv * math.sqrt(2)) ))
print(cumulativeCDF(8, m, np.sqrt(variance)))
plt.plot(
np.linspace(1, 13, 130),
list(cumulativeCDF(i, m, np.sqrt(variance)) for i in np.linspace(1, 13, 130)))
plt.grid(True)
plt.show()
df = pd.read_csv("data/student/student-mat.csv", ";")
df.sample(5)
df.info()
df[["G1", "G3", "G2"]].plot.hist(stacked=True, bins=20,alpha=0.5)
def is_anomoly(g1, g2, g3, df):
g1_d = df["G1"].describe()
g2_d = df["G2"].describe()
g3_d = df["G3"].describe()
print(g1_d)
print(g2_d)
print(g3_d)
p1 = calc_p(g1, g1_d["mean"], np.power(g1_d["std"], 2))
p2 = calc_p(g2, g2_d["mean"], np.power(g2_d["std"], 2))
p3 = calc_p(g3, g3_d["mean"], np.power(g3_d["std"], 2))
print(p1)
print(p2)
print(p3)
print(np.log(p1)+np.log(p2)+np.log(p3))
is_anomoly(10.908861, 10.713924, 10.415190, df)
from pandas.tools.plotting import scatter_matrix
scatter_matrix(df[
["G1", "G2", "G3", "age", "Walc", "goout", "absences", "studytime","famsize"]], alpha=0.2, figsize=(12, 12), diagonal='kde')
plt.show()
| gaussian.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This package uses a Twitter POST api such as tweepy in order to schedule / send a series of threaded tweets that refer to one another so that they form a "chain".
#
# Here's an example of how to use it:
import tweepy
from threader import Threader
import os
tweepy.__version__
# ### Set up the Twitter API
#
# In order to post to Twitter from Python, you'll need access
# tokens for the Twitter API. To create these, check out
# https://developer.twitter.com/en/docs/basics/authentication/guides/single-user.
# +
# personal details
consumer_key = os.environ.get("TWITTER_API_KEY")
consumer_secret = os.environ.get("TWITTER_API_SECRET_KEY")
access_token = os.environ.get("TWITTER_ACCESS_TOKEN")
access_token_secret = os.environ.get("TWITTER_ACCESS_TOKEN_SECRET")
# authentication of consumer key and secret
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
# authentication of access token and secret
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth)
# -
# ### Construct our tweets
#
# Threader will automatically add "end" characters (if desired). You can check what the tweets will look like
# once they're posted before actually sending them to
# Twitter.
username = None
tweets = ["OK this should work now", "does it work?! is it threaded?!", "maybe........", "fingers crossed!"]
th = Threader(tweets, api, wait=1, user=username)
th
# Now let's send them off!
th.send_tweets()
# ### If your tweets are too long, Threader won't send them!
username = None
tweets = ["OK this should work now", "does it work?! is it threaded?!", "maybe........", "fingers crossed!"]
th = Threader(tweets, api, user=username, max_char=20)
th
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import xgboost as xgb
import lightgbm as lgb
import time
import datetime
import os
import sklearn
from sklearn import metrics
import xgboost as xgb
from xgboost import XGBClassifier
from sklearn.model_selection import GridSearchCV
from collections import OrderedDict
from datetime import datetime, timedelta
from sklearn.externals import joblib
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold, StratifiedKFold
from sklearn.metrics import roc_auc_score
import tqdm
from collections import Counter
import warnings
warnings.filterwarnings('ignore')
# ### Load data
data = joblib.load('feature_308.pkl')
data.shape
data.head()
data['标签'] = data['标签'].fillna(-1)
print(data['标签'].value_counts())
# +
# from sklearn.utils import shuffle
# data = shuffle(data)
# data = data.iloc[:10000,]
# train_data = data[data['标签']!=-1]
# test_data = data[data['标签']==-1].drop(columns = ['标签'])
# print('train_data:',train_data.shape,'\ntest_data:',test_data.shape)
# x = train_data.drop(columns = '标签')
# y = train_data['标签']
# print('train_x:',x.shape,'\ntrain_label:',y.shape)
# x_train, x_test, y_train, y_test = train_test_split(x, y, train_size=0.75, random_state=1024)
# +
from sklearn.utils import shuffle
data = shuffle(data)
data = data.iloc[:50000,]
train_data = data[data['标签']!=-1].drop(columns='申请编号',axis=1)
test_data = data[data['标签']==-1].drop(columns = ['申请编号','标签'])
print('train_data:',train_data.shape,'\ntest_data:',test_data.shape)
x = train_data.drop(columns = '标签')
y = train_data['标签']
print('train_x:',x.shape,'\ntrain_label:',y.shape)
x_train, x_test, y_train, y_test = train_test_split(x, y, train_size=0.75, random_state=1024)
# -
x_train.columns
# +
# def ks_min(preds,dtrain):
# #自定义损失函数
# labels = dtrain.get_label()
# fpr,tpr,thresholds= sklearn.metrics.roc_curve(labels,preds,pos_label=1)
# #ks_1 = 1-max(tpr - fpr)
# return '1-KS',1-abs(fpr - tpr).max()
# def xgb_auto_gs(x_train,y_train,x_test):
# '''
# x_train:训练用数据集 dataframe
# y_train:训练用标签 Series
# x_test:测试集
# return:
# result['best_params']:最优参数组
# result['y_test']:x_test预测结果
# '''
# params = OrderedDict()
# #调参范围(可调整,如需增加调整的参数,可继续在params组中添加)
# params['tree'] = {'max_depth': [3,5,7,9],'n_estimators': [500,1000,2000]}
# params['over_fitting'] = {'gamma':[0,0.1,0.3,0.5],'min_child_weight': [1,4,7,10,15]}
# params['variable'] = {'subsample':[0.6,0.7,0.8,0.9],'colsample_bytree':[0.6,0.7,0.8,0.9]}
# #默认参数(可调整)
# best_params = { 'learning_rate':0.1, #学习率 大:欠拟合;小 :过拟合
# 'n_estimators':500, # 树的个数--1000棵树建立xgboost 小:欠拟合;大 :过拟合
# 'max_depth':5, # 树的深度 小:欠拟合;大 :过拟合
# 'min_child_weight':7, # 叶子节点最小权重 大:欠拟合;小 :过拟合
# 'gamma':0.5, # 惩罚项中叶子结点个数前的参数 大:欠拟合;小 :过拟合
# 'subsample':0.7, # 随机选择%s样本建立决策树 小:欠拟合;大 :过拟合
# 'colsample_bytree':0.7, # 随机选择%s特征建立决策树 小:欠拟合;大 :过拟合
# 'objective':'binary:logistic','silent':1}
# #分批调试
# for i in params:
# print('-----searching params: %s-----' % params[i])
# xgbc_best = XGBClassifier(**best_params)
# gs = GridSearchCV(xgbc_best, params[i], n_jobs=-1, cv=5, verbose=1)
# gs.fit(x_train, y_train, eval_set=[(x_train, y_train)], eval_metric=ks_min, early_stopping_rounds=50,verbose=True)
# best_parami = gs.best_params_#最优参数
# print('-----best params: %s-----' % best_parami)
# for param_i in best_parami:
# best_params[param_i] = best_parami[param_i]#更新最优参数
# xgbc_best = XGBClassifier(**best_params)
# xgbc_best.fit(x_train, y_train, eval_set=[(x_train, y_train)], eval_metric=ks_min, early_stopping_rounds=50,verbose=True)
# y_pre = xgbc_best.predict_proba(x_test)
# y_pre = y_pre[:,1]
# result={'best_params':best_params,'y_test':y_pre}
# return result
# +
def ks_min(preds,dtrain):
#自定义损失函数
labels = dtrain.get_label()
fpr,tpr,thresholds= sklearn.metrics.roc_curve(labels,preds,pos_label=1)
#ks_1 = 1-max(tpr - fpr)
return '1-KS',1-abs(fpr - tpr).max()
def xgb_auto_gs(x_train,y_train,x_test):
'''
x_train:训练用数据集 dataframe
y_train:训练用标签 Series
x_test:测试集
return:
result['best_params']:最优参数组
result['y_test']:x_test预测结果
'''
# params = OrderedDict()
# #调参范围(可调整,如需增加调整的参数,可继续在params组中添加)
# params['tree'] = {'max_depth': [3,5,7,9],'n_estimators': [500,1000,2000]}
# params['over_fitting'] = {'gamma':[0,0.1,0.3,0.5],'min_child_weight': [1,4,7,10,15]}
# params['variable'] = {'subsample':[0.6,0.7,0.8,0.9],'colsample_bytree':[0.6,0.7,0.8,0.9]}
#默认参数(可调整)
best_params = { 'learning_rate':0.1, #学习率 大:欠拟合;小 :过拟合
'n_estimators':500, # 树的个数--1000棵树建立xgboost 小:欠拟合;大 :过拟合
'max_depth':5, # 树的深度 小:欠拟合;大 :过拟合
'min_child_weight':7, # 叶子节点最小权重 大:欠拟合;小 :过拟合
'gamma':0.5, # 惩罚项中叶子结点个数前的参数 大:欠拟合;小 :过拟合
'subsample':0.7, # 随机选择%s样本建立决策树 小:欠拟合;大 :过拟合
'colsample_bytree':0.7, # 随机选择%s特征建立决策树 小:欠拟合;大 :过拟合
'objective':'binary:logistic','silent':1}
# #分批调试
# for i in params:
# print('-----searching params: %s-----' % params[i])
# xgbc_best = XGBClassifier(**best_params)
# gs = GridSearchCV(xgbc_best, params[i], n_jobs=-1, cv=5, verbose=1)
# gs.fit(x_train, y_train, eval_set=[(x_train, y_train)], eval_metric=ks_min, early_stopping_rounds=50,verbose=True)
# best_parami = gs.best_params_#最优参数
# print('-----best params: %s-----' % best_parami)
# for param_i in best_parami:
# best_params[param_i] = best_parami[param_i]#更新最优参数
xgbc_best = XGBClassifier(**best_params)
xgbc_best.fit(x_train, y_train, eval_set=[(x_train, y_train)], eval_metric=ks_min,\
early_stopping_rounds=100,verbose=50)
y_pre = xgbc_best.predict_proba(x_test)
y_pre = y_pre[:,1]
result={'best_params':best_params,'y_test':y_pre}
return result
# -
xgb_auto_gs(x_train,y_train,x_test)
best_params = { 'learning_rate':0.1, #学习率 大:欠拟合;小 :过拟合
'n_estimators':500, # 树的个数--1000棵树建立xgboost 小:欠拟合;大 :过拟合
'max_depth':5, # 树的深度 小:欠拟合;大 :过拟合
'min_child_weight':7, # 叶子节点最小权重 大:欠拟合;小 :过拟合
'gamma':0.5, # 惩罚项中叶子结点个数前的参数 大:欠拟合;小 :过拟合
'subsample':0.7, # 随机选择%s样本建立决策树 小:欠拟合;大 :过拟合
'colsample_bytree':0.7, # 随机选择%s特征建立决策树 小:欠拟合;大 :过拟合
'objective':'binary:logistic','silent':1}
xgbc_best = XGBClassifier(**best_params)
xgbc_best.fit(x_train, y_train, eval_set=[(x_train, y_train)], eval_metric=ks_min,\
early_stopping_rounds=100,verbose=50)
y_pre = xgbc_best.predict_proba(x_test)
y_pre = y_pre[:,1]
fpr,tpr,thresholds= sklearn.metrics.roc_curve(y_test,y_pre,pos_label=1)
max(tpr - fpr)
y_pre = xgbc_best.predict_proba(x_train)
y_pre = y_pre[:,1]
fpr,tpr,thresholds= sklearn.metrics.roc_curve(y_train,y_pre,pos_label=1)
max(tpr - fpr)
pd.set_option('display.max_rows',400)
pd.DataFrame(data.columns)
data.iloc[:20,129:140]
def k_fold_xgb_train(feature_num):
feature_importance = joblib.load("feture_importance_data_B")
features = list(feature_importance["names"])[0:feature_num]
train_x_1 = train_x[features]
test_x_1 = test_x[features]
predictions_xgb = xgb_model(1, train_x_1.values, train_y.values, test_x_1.values, "xgb_model0927_1", "log1")
sub = pd.DataFrame(columns=["客户号", "违约概率"])
sub["申请编号"] = list(test_id)
sub["违约概率"] = list(predictions_lgb)
sub.to_csv("submit_xgb0927_1.csv", sep=",", index=False, header=0)
return predictions_xgb, sub
| XGB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution:
def getRow(self, rowIndex: int) -> [int]:
# Check the boundary
if rowIndex == 0:
return [1]
if rowIndex == 1:
return [1,1]
nums = [1,1]
for i in range(2, rowIndex+1):
nums.append(1)
for j in range(i-1, 0, -1):
nums[j] = nums[j-1] + nums[j]
return nums
s = Solution()
for i in range(8):
print(s.getRow(i))
| algorithms/119-Pascal's-Triangle-II.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mercari Price Suggestion
# This project seeks to solve the problem of building an algorithm for an electronic commerce company to suggest the right product prices based on the information provided by the sellers.
# ## LDA and Deep Learning
# * Latent Dirichlet allocation (LDA), a text mining technique, to discover topics in raw texts;
# * For each text, the LDA method calculates topic scores, which are combined with non-text features as the inputs for the DNN schematic
import numpy as np
import pandas as pd
import math
import matplotlib.pyplot as plt
# %matplotlib inline
# Root Mean Squared Logarithmic Error (RMSLE) as the evaluation metrics
def rmsle(y, y_pred):
"""
A function to calculate Root Mean Squared Logarithmic Error (RMSLE)
"""
assert len(y) == len(y_pred)
to_sum = [(math.log(y_pred[i] + 1) - math.log(y[i] + 1)) ** 2.0 for i,pred in enumerate(y_pred)]
return (sum(to_sum) * (1.0/len(y))) ** 0.5
#Source: https://www.kaggle.com/marknagelberg/rmsle-function
# Load the dataset
data = pd.read_table("train.tsv")
display(data.head(n=3))
print(data.shape)
# ## Data Preprocessing
# Show distribution of price
plt.hist(data['price'], normed=False, bins=10)
plt.ylabel('Number of Records');
plt.xlabel('Price')
# Log-transfer and normalize price
# +
from sklearn.preprocessing import LabelEncoder, MinMaxScaler, StandardScaler
data['target'] = np.log(data['price']+1)
target_scaler = MinMaxScaler(feature_range=(-1, 1))
data["target"] = target_scaler.fit_transform(data.target.values.reshape(-1,1))
plt.hist(data['target'], normed=False, bins=10)
plt.ylabel('Number of Records')
plt.xlabel('Target (Log-transformed and normalized Price)')
# -
print('Number of different brand names is {}.'.format(data['brand_name'].value_counts().size))
print('Number of different categories is {}.'.format(data['category_name'].value_counts().size))
# - Handle missing data;
# - Cut number of brand names and number of categories;
# - Transfer category_name, brand_name, and item_condition_id to categorical data.
# +
NUM_BRANDS = 1000
NUM_CATEGORIES = 1000
def handle_missing_inplace(dataset):
dataset['category_name'].fillna(value='missing', inplace=True)
dataset['brand_name'].fillna(value='missing', inplace=True)
dataset['item_description'].fillna(value='missing', inplace=True)
def cutting(dataset):
pop_brand = dataset['brand_name'].value_counts().loc[lambda x: x.index != 'missing'].index[:NUM_BRANDS]
dataset.loc[~dataset['brand_name'].isin(pop_brand), 'brand_name'] = 'missing'
pop_category = dataset['category_name'].value_counts().loc[lambda x: x.index != 'missing'].index[:NUM_CATEGORIES]
dataset.loc[~dataset['category_name'].isin(pop_category), 'category_name'] = 'missing'
def to_categorical(dataset):
dataset['category_name'] = dataset['category_name'].astype('category')
dataset['brand_name'] = dataset['brand_name'].astype('category')
dataset['item_condition_id'] = dataset['item_condition_id'].astype('category')
handle_missing_inplace(data)
print('Finished to handle missing')
cutting(data)
print('Finished to cut')
to_categorical(data)
print('Finished to convert categorical')
# -
# - Count vectorize category_name and brand_name.
# - Transfer item_condition_id and shipping to dummy variables.
# +
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.preprocessing import LabelBinarizer
cv = CountVectorizer()
X_category = cv.fit_transform(data['category_name'])
print('Finished count vectorize `category_name`')
#%%
lb = LabelBinarizer(sparse_output=True)
X_brand = lb.fit_transform(data['brand_name'])
print('Finished label binarize `brand_name`')
#%%
X_dummies = pd.get_dummies(data[['item_condition_id', 'shipping']],
sparse=True)
print('Finished to get dummies on `item_condition_id` and `shipping`')
# -
# ## PROCESS TEXT: RAW
# +
import nltk
#nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
import re
sw = set(stopwords.words('english')+
['eing','oing','aing','uing','aed','eed',
'would','always', 'food', 'good','great', 'us', 'b','like',
'amp', 'go','place']) # stopwords
def text_process(text,sw):
review_temp = text
review_temp = review_temp.strip() # remove \n at the end.
review_temp = re.sub('[^a-zA-Z]', ' ', review_temp)
review_temp = review_temp.split()
ps = PorterStemmer()
review_temp = [ps.stem(word) for word in review_temp
if not word in sw]
return review_temp
raw_text = np.hstack([data.item_description.str.lower(), data.name.str.lower()])
corpus = [text_process(text,sw) for text in raw_text]
# -
# Remove terms that appear less than 5 times.
from collections import defaultdict
frequency = defaultdict(int)
for text in corpus:
for token in text:
frequency[token] += 1
corpus = [[token for token in text if frequency[token] > 5]
for text in corpus]
# - Generate dictionary and term-document matrix.
# - They are saved in the **approach1_output** folder.
from gensim import corpora
dictionary = corpora.Dictionary(documents=corpus, prune_at=2000)
dictionary.save('approach1_output/dictionary.dict') # store the dictionary, for future reference
corpusVector = [dictionary.doc2bow(text) for text in corpus]
corpora.MmCorpus.serialize('approach1_output/corpusVector.mm', corpusVector)
print(corpusVector[:3])
print("Number of terms in dictionary is {}".format(len(dictionary)))
# Apply LDA method to extract 100 topics from the texts.
from gensim import models
lda = models.ldamodel.LdaModel(corpus=corpusVector, id2word=dictionary,
num_topics=100, update_every=1,
chunksize=2000, passes=1,
minimum_probability=0.0001,random_state=666)
# Show the first 5 topics.
lda.print_topics(5)
# Save the lda model.
lda.save('approach1_output/model.lda') # same for tfidf, lda, ...
f = open('approach1_output/lda_topics.csv','w')
topic_list = lda.print_topics(-1)
for topic in topic_list:
f.write(str(topic[0])+',' + ','.join(topic[1].replace('*','+').split('+')) + '\n')
f.close()
# - Create a double wrapper over the original corpus: bow->tfidf->fold-in-lsi
# - The LDA scores for each text are the topic distributions in this text.
# - The LDA scores for item_description and name are generated.
corpus_lda = lda[corpusVector]
lda_scores = pd.DataFrame([[topic[1] for topic in doc]
if doc != [] else [0,0]
for doc in corpus_lda])
print(lda_scores.shape)
nproducts = data.shape[0]
lda_item_description = lda_scores.iloc[:nproducts,]
lda_name = lda_scores.iloc[nproducts:,]
# Handle nan values in lda_item_description and lda_name.
lda_item_description[np.isnan(lda_item_description)] = 0
lda_name[np.isnan(lda_name)] = 0
# Combine variables into x (input) and y (output).
from scipy.sparse import csr_matrix, hstack
x = hstack((X_category, X_brand,X_dummies,lda_item_description,lda_name)).tocsr()
y = data["target"]
# Splite data.
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.1,
random_state=99)
X_dtrain, X_dvalid, y_dtrain, y_dvalid = train_test_split(X_train, y_train, test_size=0.01,
random_state=99)
print(X_dtrain.shape)
print(X_dvalid.shape)
# -
# # Deep Learning
# - KERAS DATA DEFINITION
# +
import keras
from keras.models import Sequential
from keras.layers import Dense,Dropout
from keras import backend as K
def rmsle_cust(y_true, y_pred):
first_log = K.log(K.clip(y_pred, K.epsilon(), None) + 1.)
second_log = K.log(K.clip(y_true, K.epsilon(), None) + 1.)
return K.sqrt(K.mean(K.square(first_log - second_log), axis=-1))
def get_model():
#params
dr_r = 0.1
model = Sequential()
model.add(Dense(units = 128, activation = 'relu', input_dim = X_dtrain.shape[1]))
model.add(Dropout(dr_r))
model.add(Dense(units = 64, activation = 'relu'))
model.add(Dropout(dr_r))
model.add(Dense(units = 1, activation = 'linear'))
model.compile(loss="mse", optimizer="rmsprop", metrics=["mae", rmsle_cust])
return model
model = get_model()
model.summary()
# -
# Fitting the model.
# +
BATCH_SIZE = 3000
epochs = 5
history = model.fit(X_dtrain, y_dtrain, epochs=epochs, batch_size=BATCH_SIZE
, validation_data=(X_dvalid, y_dvalid)
, verbose=1)
# -
# list all data in history
print(history.history.keys())
# summarize history for error
plt.plot(history.history['rmsle_cust'])
plt.plot(history.history['val_rmsle_cust'])
plt.title('model error')
plt.ylabel('rmsle')
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left')
#plt.show()
plt.savefig('error plot.png', dpi=300, bbox_inches='tight')
# Evaluate the model using validation data.
print(y_test[:6])
y_pred = model.predict(X_dvalid)
y_pred = target_scaler.inverse_transform(y_pred)
y_pred = np.expm1(y_pred)
y_true = target_scaler.inverse_transform(y_dvalid[:,np.newaxis])
y_true = np.expm1(y_true)
v_rmsle = rmsle(y_true, y_pred)
print(" RMSLE error on validation dataset: "+str(v_rmsle))
# Evaluate the model using test data.
y_pred = model.predict(X_test)
y_pred = target_scaler.inverse_transform(y_pred)
y_pred = np.expm1(y_pred)
y_true = target_scaler.inverse_transform(y_test[:,np.newaxis])
y_true = np.expm1(y_true)
v_rmsle = rmsle(y_true, y_pred)
print(" RMSLE error on test dataset: "+str(v_rmsle))
# Save model
from keras.models import model_from_yaml
# serialize model to YAML
model_yaml = model.to_yaml()
with open("approach1_output/model.yaml", "w") as yaml_file:
yaml_file.write(model_yaml)
# serialize weights to HDF5
model.save_weights("approach1_output/model.h5")
print("Saved model to disk")
| approach1_LDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Custom Adjustments
#
# The ParamTools adjustment format and logic can be augmented significantly. This is helpful for projects that need to support a pre-existing data format or require custom adjustment logic. Projects should customize their adjustments by writing their own `adjust` method and then calling the default `adjust` method from there:
#
# +
import paramtools
class Params(paramtools.Parameters):
def adjust(self, params_or_path, **kwargs):
params = self.read_params(params_or_path)
# ... custom logic here
# call default adjust method.
return super().adjust(params, **kwargs)
# -
# ## Example
#
# Some projects may find it convenient to use CSVs for their adjustment format. That's no problem for ParamTools as long as the CSV is converted to a JSON file or Python dictionary that meets the ParamTools criteria.
#
# +
import io
import os
import pandas as pd
import paramtools
class CSVParams(paramtools.Parameters):
defaults = {
"schema": {
"labels": {
"year": {
"type": "int",
"validators": {"range": {"min": 2000, "max": 2005}}
}
}
},
"a": {
"title": "A",
"description": "a param",
"type": "int",
"value": [
{"year": 2000, "value": 1},
{"year": 2001, "value": 2},
]
},
"b": {
"title": "B",
"description": "b param",
"type": "int",
"value": [
{"year": 2000, "value": 3},
{"year": 2001, "value": 4},
]
}
}
def adjust(self, params_or_path, **kwargs):
"""
A custom adjust method that converts CSV files to
ParamTools compliant Python dictionaries.
"""
if os.path.exists(params_or_path):
paramsdf = pd.read_csv(params_or_path, index_col="year")
else:
paramsdf = pd.read_csv(io.StringIO(params_or_path), index_col="year")
dfdict = paramsdf.to_dict()
params = {"a": [], "b": []}
for label in params:
for year, value in dfdict[label].items():
params[label] += [{"year": year, "value": value}]
# call adjust method on paramtools.Parameters which will
# call _adjust to actually do the update.
return super().adjust(params, **kwargs)
# -
# Now we create an example CSV file. To keep the example self-contained, the CSV is just a string, but this example works with CSV files, too. The values of "A" are updated to 5 in 2000 and 6 in 2001, and the values of "B" are updated to 6 in 2000 and 7 in 2001.
#
# +
# this could also be a path to a CSV file.
csv_string = """
year,a,b
2000,5,6\n
2001,6,7\n
"""
params = CSVParams()
params.adjust(csv_string)
# -
params.a
params.b
# Now, if we use `array_first` and [`label_to_extend`](/api/extend/), the params instance can be loaded into a Pandas
# DataFrame like this:
#
# +
csv_string = """
year,a,b
2000,5,6\n
2001,6,7\n
"""
params = CSVParams(array_first=True, label_to_extend="year")
params.adjust(csv_string)
params_df = pd.DataFrame.from_dict(params.to_dict())
params_df
# -
params_df["year"] = params.label_grid["year"]
params_df.set_index("year")
| docs/api/custom-adjust.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="K1spMRzajubL"
# 2021.1
#
#
# 免責事項:本教材(解説,およびプログラム)は教育目的に提供されるものです.本教材を利用したことによる直接あるいは間接的な損害に関して,著作者および著作者の所属組織等,如何なる個人・組織も一切の責任を負いません.本教材の利用は利用者個人の責任のもと行ってください.
# + id="5jJTLCi_Jfo2"
import numpy as np
import pandas as pd
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import GridSearchCV
import matplotlib.pyplot as plt
# + id="g-PyaAqLd46G"
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data', header=None) # web上のデータを読み込む
df.columns = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species'] # 列名を付ける
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="qjFWcfupeS-I" outputId="2d0d1318-874c-4861-a19a-9088cd10710e"
display(df)
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="cK5fqFPiel9u" outputId="d06e648c-1ad9-4582-c7f9-e5ea4d05f27b"
features = df.loc[:, 'sepal_length':'petal_width'] # 特徴量だけを取り出す
display(features)
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="0HCj3UUraDze" outputId="92284570-72d8-4e2e-fab9-72adaa673d68"
targets = df.loc[:, ['species']] # 正解カテゴリーを取り出す
display(targets)
# + id="jBgzDuUnfEcA"
# dataframeをndarrayに変換
#X = features.values
#y = targets.values
X = np.array(features)
y = np.array(targets)
y = y.ravel() # 配列の次元を落とす
# + id="idJy_yDsc3Pw"
scaler = StandardScaler() # scalerインスタンスの生成
X = scaler.fit_transform(X) # 特徴量データの標準化
# + id="B_5uBnbZfoqI"
X_train, X_test, y_train, y_test = train_test_split(X, y) # 訓練データと検証データへの分割
# + id="pPFbgQE_eT5g"
clf = MLPClassifier(max_iter=10000) # 多層ニューラルネットワークの生成
# + colab={"base_uri": "https://localhost:8080/"} id="gazP9-EYejgA" outputId="0784e63b-360e-49ba-f432-2c04350b3bd6"
clf.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="eq-MGClCgEdo" outputId="8a26b2d8-5720-4764-8ff3-0ff122ecdd68"
plt.plot(clf.loss_curve_)
plt.title("Loss curve") # グラフのタイトル
plt.xlabel("Iteration") # x軸は繰り返し数(エポック数)
plt.ylabel("Loss") # y軸は損失値
plt.grid() # グリッド線を描く
# + colab={"base_uri": "https://localhost:8080/"} id="q1rcSMLWgijX" outputId="4241a985-7849-4e38-bb63-8ac27bc0a2c1"
clf.predict(X_test) # 検証データに対する予測結果表示
# + colab={"base_uri": "https://localhost:8080/"} id="gPBMENIfg493" outputId="336c4d62-39db-4fb6-f293-19942bcc4c39"
c = clf.score(X_test, y_test)
print("正解率は{}%です.".format(c * 100)) # 検証データでの成績表示
# + id="QImgXAeNf9Qy"
| scikit_learnでニューラルネットワーク実習.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:analysis3-18.04]
# language: python
# name: conda-env-analysis3-18.04-py
# ---
# # When should I use `.load()`?
#
# The `.load()` function converts a Dask array to a Numpy array, reading it into memory
#
# If you've narrowed down your selection to a small number of grid points, but it covers a long timeseries (and so a large number of files) using `.load()` can speed things up, as Dask doesn't need to open up each file every time you do something with the array.
#
# Remember that the size of an array in bytes is normally `8 * array.size`
# # What other ways are there to use `.groupby()`?
#
# `.groupby()` is really handy in the time dimension. Some other ways to use it:
#
# * You can add a new dimension that categorises each point - the `regionmask` library is one way to do this
# * `.groupby_bins()` can bin a coordinate, either into a number of equal bins or using specified points
# # Filtering curvilinear Grids
#
# `.sel()` is less helpful on a curvilinear grid. You can however do a similar thing with `.where()`:
#
# ```
# data.where(10 <= data.lat < 20)
# ```
# # Libraries that build upon Xarray + Dask
#
#
# http://xarray.pydata.org/en/latest/faq.html#what-other-projects-leverage-xarray
#
# http://pangeo-data.org
| xarray+dask/3-q-and-a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this project, I will be analysing reviews that were given to Amazon Products. This dataset contains around 41000 reviews of various Amazon products such as Fire tablet, Fire stick. Along with it, there is also the rating given by the user and if the user recommends this product or not.
#
# **Aim**: Use the textual reviews of Amazon Products to predict ratings, while experimenting various parts of textual analysis.<br>
# I will be using the text column in the dataset to predict the rating given by the user, whether it is positive or negative. This is of course under the assumption that the review given by the user matches the rating given. Hopefully, the reviews here are not something like this one!<br>
#
# <img src="images/The-Contradictory.png" alt="Great review but one star?!" title="Contradictory review" /> <br>
# So, before I start, the idea is to change the *rating* from numbers to positive-negative or use the *doRecommend* flag and predict this rating using the reviews for each product.<br>
#
# **Dataset**: https://www.kaggle.com/datafiniti/consumer-reviews-of-amazon-products/kernels <br>
# So, let's start!
# ### Import all libraries that would be used
# +
# data analysis and wrangling
import pandas as pd
import numpy as np
import random as rnd
# visualization
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
import re
import string
import nltk
from wordcloud import WordCloud, STOPWORDS
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import confusion_matrix, roc_curve, auc, recall_score, precision_score, f1_score, accuracy_score
# -
reviews= pd.read_csv('1429_1.csv')
reviews.columns = ['id', 'name', 'asins', 'brand', 'categories', 'keys', 'manufacturer', 'date', 'dateAdded', 'dateSeen', 'didPurchase', 'doRecommend', 'id', 'numHelpful', 'rating', 'sourceURLs', 'text', 'title', 'userCity', 'userProvince', 'username']
reviews.head()
# Drop these columns as they would have no effect in the analysis later.
reviews.drop(['id', 'dateSeen', 'sourceURLs', 'userCity', 'userProvince', 'username'], axis = 1, inplace= True)
reviews[['rating', 'title', 'text' ]].isnull().sum()
# Drop the one review without the text as it has no use in the model
reviews_nna= reviews[pd.notnull(reviews['text'])]
reviews_nna[['rating', 'title', 'text' ]].isnull().sum()
# #### The top 5 most reviewed items in the dataset
# Looks like Fire tablet is way ahead of the rest here.
reviews_nna['name'].value_counts().nlargest(5)
# Create a new dataframe with the important columns. I am taking *asins* just to distinguish between the products.
sentiment_df= reviews_nna[[ 'asins', 'doRecommend', 'rating', 'title', 'text']]
sentiment_df.head()
# Create a new plot that shows the count of reviews across ratings and if the product is recommended by the reviwer.
sns.catplot(x= 'rating', col= 'doRecommend', data= sentiment_df, kind= 'count')
sentiment_df[sentiment_df['doRecommend']== False]['rating'].value_counts()
# **Interesting!** Looks like some people rate a product highly but do not recommend it.<br>
# Let's peek at the reviews that these products have been given.
pd.set_option('display.max_colwidth', -1)
sentiment_df[(sentiment_df['doRecommend']== False) & (sentiment_df['rating']== 5.0)][['doRecommend', 'rating', 'text']].head(10)
# So, taking a look at just 10 of these and you can tell that these seem like false flags. It seems that the reviews are indeed positive, so they match the ratings **BUT** the recommendation does not match that. <br>
# Now, let's look at the products recommended and their split by ratings.
sentiment_df[sentiment_df['doRecommend']== True]['rating'].value_counts()
# And now the text for these recommended products with bad ratings.
pd.set_option('display.max_colwidth', -1)
sentiment_df[(sentiment_df['doRecommend']== True) & (sentiment_df['rating']== 1.0)][['doRecommend', 'rating', 'text']].head(10)
# Well, this makes things a bit confusing. The texts here either dont match the *doRecommend* or they do not match the *rating*. <br>
# So, going just by this observation, it makes sense to use the *ratings* as a target variable instead of the *doRecommend* variable, since we would probably have a huge number of contradictory reviews. There will be a bunch of false flags, just like in the reviews above, but that is something we cannot really do anything about, unless I read through each one of the 34000 reviews and label them myself...
# ## Stemming <br>
# In Natural language writing, a word is written is several forms due to grammatical purposes. For example, the sentence- I am going to school can be written as I had gone to the school or I have been going to the school depending on the context. For processing in models, it makes sense to convert these types of sentences into a single format. This is where **stemming** and **lemmatization** come in. <br>
# Stemming and lemmatization is used to reduce a word to its root form or a common base form. So, *am*, *are*, *is* converts to *be*; *going*, *gone* converts to go and so on. The difference between stemming and lemmatization is the output of this conversion. Lemmatization reduces a word to its equivalent dictionary word, which might actually not be the case with stemming. *Amuse* stems to *amus* whereas its lemma is *amuse*.
#
# In this part, I check two types of stemming methods that are available- Porter and Snowball. So, passed a review to the functions created to generate stemmed equivalents and see the results.
# According to the outputs, the Snowball stemming seems to generate more readable output, while also cleaning up the texts( lower-case and removing special characters). So, I use the Snowball stemming for further use.
# +
from nltk import SnowballStemmer
from nltk import PorterStemmer
from nltk import sent_tokenize, word_tokenize
stopwords = nltk.corpus.stopwords.words('english')
ss = SnowballStemmer('english')
ps= PorterStemmer()
def sentencePorterStem(sentence):
token_words= word_tokenize(sentence)
stem_sentence=[]
for word in token_words:
if word not in stopwords:
stem_sentence.append(ps.stem(word))
stem_sentence.append(' ')
return ''.join(stem_sentence)
def sentenceSnowballStem(sentence):
token_words= word_tokenize(sentence)
stem_sentence=[]
for word in token_words:
if word not in stopwords:
stem_sentence.append(ss.stem(word))
stem_sentence.append(' ')
return ''.join(stem_sentence)
sen= str(sentiment_df['text'][3])
ps_sen= sentencePorterStem(sen)
ss_sen= sentenceSnowballStem(sen)
print(sen)
print('Porter Stem- '+ ps_sen)
print('Snowball Stem- '+ ss_sen)
# -
sentiment_df['text_stem']= sentiment_df['text'].apply(sentenceSnowballStem)
# ## Vectorization <br>
#
# Now, with the stemmed reviews, it is required to create vectorized versions of the text. Vectorization is the process of converting sentences into vector or array of numbers. <br>
# There are two methods we can go with, count vectorization and TF-IDF vectorization.
# - **Count Vectorization** <br>
# In the simplest of terms, it is creating an array whose values are the number of occurences of a word in a sentence. Consider the two sentences- I am a human and I enjoy doing human things (two very human-like sentences). <br>
# After stemming both sentences to a very minimal level, if I were to count the number of occurences of each word in the first sentence, I get something like this- <br>
# I- 1, be- 1, a- 1, human- 1.<br>
# Now, for the next sentence-<br>
# I-1, enjoy- 1, be- 1, human-1, things-1.<br>
# Now, creating a list of all words in two sentences- {I, be, enjoy, a, human, things}. <br>
# Creating an array where the words are the columns and the sentences are the rows, we get something like this-<br>
#
# array([[1, 1, 0, 0, 1, 0],<br>
#    [1, 1, 1, 0, 1, 1] ]
#
# This is the count vectorizer equivalent of the two sentences.
#
# - **TF-IDF Vectorization** <br>
# It is a combination of two concepts- TF (Term Frequency) and IDF (Inverse Document Frequency). <br>
# Term frequency is as the name suggests, the frequency of a word occuring in that document. So, it is the ratio of the number of occurences of the word in the document by the total number of words in the document.<br>
# Inverse document frequency is the log base 10 of the ratio of number of documents by the number of documents that the word appears in. The idea behind this is to find words that are more important compared to others.<br>
# <img src="images/IDF-Formula.png" alt="IDF formula" title="IDF formula" /> <br>
# So, if a word occurs in all the documents, it is not really an important word.<br>
# In this part, I generate the TF-IDF version of the whole review dataset to understand the method and see the output that is generated by it.
# Since TF-IDF and Count vectorization both use similar concepts and generate similar output format, I decided to check the TF-IDF version.
# +
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
sbstem_vectorizer= TfidfVectorizer()
textfeatures= sbstem_vectorizer.fit_transform(sentiment_df['text_stem'])
# -
# Check list of feature names generated by the version. The vocabulary dictionary has the terms which are mapped to the feature indices.
# +
from itertools import islice
def take(n, iterable):
"Return first n items of the iterable as a list"
return list(islice(iterable, n))
print(take(20, sbstem_vectorizer.vocabulary_.items()))
# -
pd.DataFrame(textfeatures.toarray()).head(15)
# Change the dataframe column names to the terms to make sense of the dataframe.
text_vect_df= pd.DataFrame(textfeatures.toarray(), columns= sbstem_vectorizer.vocabulary_)
text_vect_df.head(15)
#text_vect_df['product'].unique()
# ### Basic sentiment analysis using NLTK.<br>
# Using 3 sentences of my own, I use NLTK library to identify which sentence are positive, negative or neutral. This can be done by the 'compound' value in the polarity score.
# +
from nltk.sentiment.vader import SentimentIntensityAnalyzer
sia = SentimentIntensityAnalyzer()
pos_sen= 'I am very happy. I absolutely love it. Great job!'
print('positive - '+ str(sia.polarity_scores(pos_sen)))
neg_sen= 'It is so disgusting. I am very angry. I will murder him.'
print('negative - '+ str(sia.polarity_scores(neg_sen)))
neutral_sen= 'I am writing python in jupyter notebook.'
print('negative - '+ str(sia.polarity_scores(neutral_sen)))
# +
dataset=sentiment_df[['text_stem', 'rating']]
dataset['rating'] = dataset['rating'].apply(lambda x:'Positive' if x>=4 else 'Negative')
def SentimentCoeff(sentence):
score = sia.polarity_scores(sentence)
return score['compound']
dataset['sentiment_coeff']= dataset['text_stem'].apply(SentimentCoeff)
dataset.head()
# -
sns.boxenplot(x= 'rating', y= 'sentiment_coeff', data= dataset)
# So, after using the sentiment analyzer available in NLTK, the positive review seems more logical- higher sentiment coefficient for these reviews, whereas the sentiment coefficient seems more neutral for negative reviews. But, there is also the way categorization of these reviews is done may shed some light here. Reviews with 3 or 2 stars are now negative reviews, whereas the actual review might not be that negative.
# ### Preparing for model <br>
# Let's start with the data preparation for creating a logistic regression model. We will start by splitting the whole data into 2 parts for training and testing with 70-30 split.
# +
x= dataset['text_stem']
y= dataset['rating']
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.3, random_state=42)
# -
# Using TF-IDF and Count vector formats as inputs to the Logistic regression model to check which version provides better output, along with finding the optimal parameters using GridSearchCV.
# +
tfidf_vectorizer= TfidfVectorizer()
x_train_features_tfidf= tfidf_vectorizer.fit_transform(x_train)
x_test_features_tfidf= tfidf_vectorizer.transform(x_test)
count_vectorizer= CountVectorizer()
x_train_features_count= count_vectorizer.fit_transform(x_train)
x_test_features_count= count_vectorizer.transform(x_test)
# +
logreg_params={'penalty': ['l1', 'l2'], 'C': [0.01, 0.1, 1, 10, 100]}
grid_logreg= GridSearchCV(LogisticRegression(solver= 'liblinear'), logreg_params, cv=5)
grid_logreg.fit(x_train_features_count, y_train)
logreg= grid_logreg.best_estimator_
print('Best Penalty:', grid_logreg.best_estimator_.get_params()['penalty'])
print('Best C:', grid_logreg.best_estimator_.get_params()['C'])
c_value= grid_logreg.best_estimator_.get_params()['C']
pen_value= grid_logreg.best_estimator_.get_params()['penalty']
# -
logreg=LogisticRegression(C= c_value, penalty= pen_value, solver= 'liblinear')
logreg.fit(x_train_features_count, y_train)
y_pred= logreg.predict(x_test_features_count)
# ### F1 score and Confusion Matrix <br>
#
# In two-class classification algorithms, model evaluation is done by calculating metrics such as precision, recall and F1 score.
# Starting with the basics, in a two- class supervised classification algorithm, the data can be categorized in either the correct class or the incorrect class. If it is classified in the incorrect class, it would lead to an error. Depending upon the targeted and predicted class, it can be classified as a **Type-I error** or a **Type-II error**. <br>
# <img src="images/Precisionrecall.png" alt="PR" title="Precision- Recall basics" /> <br>
# The diagram gives a pretty good picture of the classification metrics. Type-I error is also known as False Positive, while Type-II error is also known as False Negative.<br>
# *Precision* is ratio of True Positive and actual results- that is True Positive and False Positive, while *Recall* is ratio of True Positive and predicted results- that is True Positive and False Negative. *Accuracy* is the average of True Positive and True Negative.<br>
# F1 Score is the harmonic mean of Precision and Recall. It calculates in such a way that both the metrics are given equal weightage, so there is no need to sacrifice one metric over the other.<br>
# Confusion matrix is a plot similar to the diagram above, with the counts in each predicted category.
print('Recall Score: {:.2f}'.format(recall_score(y_test, y_pred, pos_label= 'Positive')))
print('Precision Score: {:.2f}'.format(precision_score(y_test, y_pred, pos_label= 'Positive')))
print('F1 Score: {:.2f}'.format(f1_score(y_test, y_pred, pos_label= 'Positive')))
print('Accuracy Score: {:.2f}'.format(accuracy_score(y_test, y_pred)))
# Plotting confusion matrix of the predicted and actual values.
# +
cnf_matrix = confusion_matrix(y_test, y_pred)
df_cnf_matrix= pd.DataFrame(cnf_matrix)
sns.heatmap(df_cnf_matrix, annot=True, fmt='g', cmap="Blues")
plt.ylabel('True label')
plt.xlabel('Predicted label')
# +
logreg_params={'penalty': ['l1', 'l2'], 'C': [0.01, 0.1, 1, 10, 100]}
#Added solver saga to repress warnings- saga recommended by sklearn
grid_logreg= GridSearchCV(LogisticRegression(solver= 'liblinear'), logreg_params, cv=5)
grid_logreg.fit(x_train_features_tfidf, y_train)
logreg= grid_logreg.best_estimator_
print('Best Penalty:', grid_logreg.best_estimator_.get_params()['penalty'])
print('Best C:', grid_logreg.best_estimator_.get_params()['C'])
c_value_tf= grid_logreg.best_estimator_.get_params()['C']
pen_value_tf= grid_logreg.best_estimator_.get_params()['penalty']
# -
logreg_tf=LogisticRegression(C= c_value_tf, penalty= pen_value_tf, solver= 'liblinear')
logreg_tf.fit(x_train_features_tfidf, y_train)
y_pred_tf= logreg_tf.predict(x_test_features_tfidf)
print('Recall Score: {:.2f}'.format(recall_score(y_test, y_pred_tf, pos_label= 'Positive')))
print('Precision Score: {:.2f}'.format(precision_score(y_test, y_pred_tf, pos_label= 'Positive')))
print('F1 Score: {:.2f}'.format(f1_score(y_test, y_pred_tf, pos_label= 'Positive')))
print('Accuracy Score: {:.2f}'.format(accuracy_score(y_test, y_pred_tf)))
# +
cnf_matrix = confusion_matrix(y_test, y_pred_tf)
df_cnf_matrix= pd.DataFrame(cnf_matrix)
sns.heatmap(df_cnf_matrix, annot=True, fmt='g', cmap="Blues")
plt.ylabel('True label')
plt.xlabel('Predicted label')
# -
# Overall, it seems that the two versions generate similar results. The F1 scores are the same and they have similar scores for the rest of the parameters. So, for this dataset any vectorization of the text would work well.
# ## Final Words<br>
# So, we walked through various parts of a data science projects- data cleaning, exploration, data visualization, data manipulation for creating a model. We also looked at various techniques used for text analytics- vectorization and sentiment analysis.<br>
# Looking back, I would like to explore this dataset more using similar machine learning techniques, especially in the model part. There are several other classification algorithms such as Random Forest and Naive Bayes which I would like to read upon and possibly implement with the dataset to compare the different algorithms. There is also a possibility of including neural network systems which can be checked upon as well.<br>
| AmazonReview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring the IBL Data Pipeline
#
# Here we will introduce some useful DataJoint tools and concepts to help you explore the IBL data pipeline. Before proceeding make sure that you have installed the [IBL python environment](../../02_installation.md) and set up your [Datajoint credentials](../../dj_docs/dj_credentials.md)
#
#
# ## A few definitions
# First of all, let's define a few basic definitions:
#
# - Table - collection of rows and columns that contain data
# - Schema - a collection of related tables
# - Module - script where schema and associated tables are defined
# - Pipeline - collection of schemas
#
# Example nomenclature would be to say that we want to get data from the `Subject table` stored in the `ibl_subjects schema` which are together defined in the `subject module` in the `IBL pipeline`.
#
# In practice, `modules` are often referred to as schemas and so we would refer to the `subject module` as the `subject schema`
# Let's now proceed by importing Datajoint and some schemas from the IBL pipeline
import datajoint as dj
dj.config['display.limit'] = 5
from ibl_pipeline import reference, subject, action, acquisition, data, behavior
from ibl_pipeline.analyses import behavior as behavior_analyses
# ## 1. Browsing schemas
# The IBL pipeline contains a number of different schemas. To browse which schemas are available, we can use the `dj.list_schemas()` command,
dj.list_schemas()
#
# ### Major schemas include:
# Meta data from **Alyx**: `ibl_reference`, `ibl_subject`, `ibl_action`, `ibl_acquisition`, and `ibl_data`
# Imported data from **FlatIron**: `ibl_behavior` and `ibl_ephys`
# Computed analzyed results: `ibl_analyses_behavior`
#
# ## 2. Browsing tables in a schemas
# We can see what tables are defined in a schema using the `dj.Diagram` command. For example, to see the tables defined in the `subject` schema we can type,
# ```
# dj.Diagram()
# ```
# <div class="alert alert-info">
#
# Note
#
# For more information about the what the different colours and line types mean, please refer to this more comprehensive [tutorial](https://github.com/int-brain-lab/IBL-pipeline/blob/master/notebooks/notebooks_tutorial/201909_code_camp/1-Explore%20IBL%20data%20pipeline%20with%20DataJoint.ipynb)
# </div>
# We can also use the following code snippet to list the tables that are defined in a schema
# +
from datajoint.user_tables import UserTable
import inspect
def list_tables(schema):
for k in dir(schema):
t = getattr(schema, k)
if inspect.isclass(t) and issubclass(t, UserTable):
print(k)
# -
list_tables(subject)
# ## 3. Getting the detailed definition of a table
# To find out details about a table, we can use the `describe` method
subject.Subject.describe()
# ## 4. Browsing data in tables - queries
# Query all subjects
subject.Subject()
# ### Restriction `&`: filtering data
# #### Restriction: Query one subject
# restrict by string
subject.Subject & 'subject_nickname="ibl_witten_10"'
# restrict by dictionary
from uuid import UUID
subject.Subject & {'subject_uuid': UUID('00c60db3-74c3-4ee2-9df9-2c84acf84e92')}
# #### Restriction: Query all male subjects
subject.Subject & {'sex': 'm'}
# #### Restriction: Query subjects born after a date
subject.Subject & 'subject_birth_date > "2019-01-01"'
# #### Restriction: Query subjects within a range of dates
subject.Subject & 'subject_birth_date between "2019-01-01" and "2019-04-01"'
# #### Restriction: Query subjects on multiple attributes
subject.Subject & 'subject_birth_date > "2019-01-01"' & 'sex="M"'
# #### Restriction: Query subjects restricted by other tables
# subjects with Behavioural sessions
subject.Subject & acquisition.Session
# subjects without Behavioural sessions
subject.Subject - acquisition.Session
# ### Join `*`: gather information from different tables
subject.Subject * acquisition.Session
# ### Projection `.proj()`: focus on attributes of interest
subject.Subject.proj()
subject.Subject.proj('subject_birth_date', 'sex')
# #### rename attribute with ***proj()***
subject.Subject.proj('sex', dob='subject_birth_date')
# #### perform simple computations with ***proj***
# **Example 1: Get the date of a session**
sessions_with_date = acquisition.Session.proj(session_date='date(session_start_time)')
sessions_with_date
# **Example 2: Get the age of the animal at each session**
# First get the date of birth and the session date into the same query
q = subject.Subject * acquisition.Session
# Then compute the age
q_with_age = q.proj(age='datediff(session_start_time, subject_birth_date)')
q_with_age
# ### Aggregation `.aggr()`: simple computation of one table against another table
# **Example: How many sessions has each subject done so far?**
subject.Subject.aggr(acquisition.Session, 'subject_nickname', n='count(*)')
# ## 5. Fetching data
# ### Fetch all fields: `fetch()`
# fetch all data from a table
subjs = subject.Subject.fetch()
subjs[:5]
subjs['subject_uuid'][:5]
subjs['subject_birth_date'][:5]
# fetch as a list of dictionaries
subjs_dict = subject.Subject.fetch(as_dict=True)
subjs_dict[:5]
# fetch as pandas dataframe
subjs_df = subject.Subject.fetch(format='frame')
subjs_df[:5]
# fetch the primary key
pk = subject.Subject.fetch('KEY')
pk[:5]
# fetch specific attributes
dob, sex = subject.Subject.fetch('subject_birth_date', 'sex')
dob[:5]
info = subject.Subject.fetch('subject_birth_date', 'sex', as_dict=True)
info[:5]
# ### Fetch data from only one entry: `fetch1`
ibl_witten_10 = (subject.Subject & {'subject_nickname': 'ibl_witten_10'}).fetch1('KEY')
ibl_witten_10
IBL_10 = (subject.Subject & {'subject_nickname': 'IBL_10'}).fetch1()
IBL_10
| docs_gh_pages/notebooks/dj_basics/dj_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#Reading data
with tf.Session():
input = tf.placeholder(tf.float32)
classifier = ...
print(classifier.eval(feed_dict={input: my_python_preprocessing_fn()}))
# +
filename_queue = tf.train.string_input_producer(["file0.csv", "file1.csv"])
reader = tf.TextLineReader()
key, value = reader.read(filename_queue)
# Default values, in case of empty columns. Also specifies the type of the
# decoded result.
record_defaults = [[1], [1], [1], [1], [1]]
col1, col2, col3, col4, col5 = tf.decode_csv(
value, record_defaults=record_defaults)
features = tf.stack([col1, col2, col3, col4])
with tf.Session() as sess:
# Start populating the filename queue.
coord = tf.train.Coordinator()
threads = tf.train.start_queue_runners(coord=coord)
for i in range(1200):
# Retrieve a single instance:
example, label = sess.run([features, col5])
coord.request_stop()
coord.join(threads)
# +
def read_my_file_format(filename_queue):
reader = tf.SomeReader()
key, record_string = reader.read(filename_queue)
example, label = tf.some_decoder(record_string)
processed_example = some_processing(example)
return processed_example, label
def input_pipeline(filenames, batch_size, num_epochs=None):
filename_queue = tf.train.string_input_producer(
filenames, num_epochs=num_epochs, shuffle=True)
example, label = read_my_file_format(filename_queue)
# min_after_dequeue defines how big a buffer we will randomly sample
# from -- bigger means better shuffling but slower start up and more
# memory used.
# capacity must be larger than min_after_dequeue and the amount larger
# determines the maximum we will prefetch. Recommendation:
# min_after_dequeue + (num_threads + a small safety margin) * batch_size
min_after_dequeue = 10000
capacity = min_after_dequeue + 3 * batch_size
example_batch, label_batch = tf.train.shuffle_batch(
[example, label], batch_size=batch_size, capacity=capacity,
min_after_dequeue=min_after_dequeue)
return example_batch, label_batch
# +
#need more parallelism or shuffling of examples between files
def read_my_file_format(filename_queue):
# Same as above
def input_pipeline(filenames, batch_size, read_threads, num_epochs=None):
filename_queue = tf.train.string_input_producer(
filenames, num_epochs=num_epochs, shuffle=True)
example_list = [read_my_file_format(filename_queue)
for _ in range(read_threads)]
min_after_dequeue = 10000
capacity = min_after_dequeue + 3 * batch_size
example_batch, label_batch = tf.train.shuffle_batch_join(
example_list, batch_size=batch_size, capacity=capacity,
min_after_dequeue=min_after_dequeue)
return example_batch, label_batch
# +
# Create the graph, etc.
init_op = tf.global_variables_initializer()
# Create a session for running operations in the Graph.
sess = tf.Session()
# Initialize the variables (like the epoch counter).
sess.run(init_op)
# Start input enqueue threads.
coord = tf.train.Coordinator()
threads = tf.train.start_queue_runners(sess=sess, coord=coord)
try:
while not coord.should_stop():
# Run training steps or whatever
sess.run(train_op)
except tf.errors.OutOfRangeError:
print('Done training -- epoch limit reached')
finally:
# When done, ask the threads to stop.
coord.request_stop()
# Wait for threads to finish.
coord.join(threads)
sess.close()
# -
training_data = ...
training_labels = ...
with tf.Session():
input_data = tf.constant(training_data)
input_labels = tf.constant(training_labels)
...
training_data = ...
training_labels = ...
with tf.Session() as sess:
data_initializer = tf.placeholder(dtype=training_data.dtype,
shape=training_data.shape)
label_initializer = tf.placeholder(dtype=training_labels.dtype,
shape=training_labels.shape)
input_data = tf.Variable(data_initializer, trainable=False, collections=[])
input_labels = tf.Variable(label_initializer, trainable=False, collections=[])
...
sess.run(input_data.initializer,
feed_dict={data_initializer: training_data})
sess.run(input_labels.initializer,
feed_dict={label_initializer: training_labels})
| crackingcode/day7/cc_tf_day7_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
import pandas as pd
import time
url = "http://192.168.127.12" # cette url peut éventuellement changer !!
# le dataframe des publications
df = pd.read_csv('/Users/eric/Downloads/publis_lorraine_completes.csv')
# s'il y a un scientific field restant, on le supprime (il va etre recalculé )
if 'scientific_field' in df:
del df['scientific_field']
# le classifier n'a besoin que du titre et journal_name (pour le moment)
# le doi servira à recroiser les résultats in fine
publis_for_classif = df[['doi', 'title', 'journal_name']].to_dict(orient='records')
nb_publis = len(publis_for_classif)
print(f"sending {nb_publis} to classifier")
task_id = requests.post(f"{url}/classify", json={"type": "bso", "publications": publis_for_classif}).json()['data']['task_id']
print("classification", end=':')
for i in range(1000):
print(i, end='...')
r = requests.get(f"{url}/tasks/{task_id}").json()
if r.get('data', {}).get('task_status') == "started":
time.sleep(10)
elif r.get('data', {}).get('task_status') == "finished":
break
df_classification = pd.DataFrame(r['data']['task_result'])
df_classification['scientific_field'] = df_classification['bso_classification']
nb_results = len(df_classification)
print()
print(f"{nb_results} results back from classif")
# on recroise le dataframe original avec les résultats
df_final = df.merge(df_classification[['doi', 'scientific_field']], on='doi', how='left')
# -
df_final
| notebooks/TestBSOClassif.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from mpmath import *
mp.dps = 15; mp.pretty = True
from scipy.integrate import dblquad
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
#import plotly.plotly as py
import sys,os
#sys.path.append(os.path.dirname(__file__), '..','Lib')
sys.path.append(os.path.join('..','Lib'))
from stylelib.ase1_styles import ase1_sims_stl
plt.style.use(ase1_sims_stl)
a,b = 1., -100.
# Integral over gaussian function
y = quad(lambda x: exp(-a*(x-b)**2), linspace(-1100, 1000,100))
print y*y
# Integral over gaussian derivatives of known values, TEST
a,b = 1000., 36. # High 'a' turns integral into a delta function
y = quad(lambda x: x*exp(-a*(x-b)**2), linspace(-200, 100, 100))
print y*sqrt(a/pi)
# Integrals seem to always converge when the number of intervals ~= the coefficient of the exponent
# Double integrals of known values, TEST
k = 1000.0 # spring constant
f1 = lambda x1, x2: exp(-0.5*k*(x1**2 + x2**2 - 2*sqrt(1-(1/k))*x1*x2))
q = quad(f1, linspace(-10, 10, 12), linspace(-10, 10, 12))
#q = quad(f1, [-10, 10], [-10, 10])
print (q*sqrt(k))/(2.*pi)
# How does scipy's double quad method stack up to sympy? TEST
f1 = lambda x1, x2: np.exp(-0.5*k*(np.power(x1,2) + np.power(x2,2) - 2.*np.sqrt(1-(1/k))*x1*x2))
q, _= dblquad(f1, -10,10, lambda x2:-10, lambda x2:10, epsabs=0, epsrel=1.e-8)
#q = quad(f1, [-10, 10], [-10, 10])
print (q*sqrt(k))/(2.*np.pi)
# Force between parallel filaments of equal length, TESTED
k = 4.56 # spring constant
b = 1. # beta
yo = 1. # Horizontal separation
Dr = 10. # COM separation
ho = 0. # equilibrium length
c = 100. # Crosslinker affinity * fugacity
hL = 10. # Length of filaments
#fdr = lambda x1, x2, r:-1.*c*k*(x1 - x2 + r)*(1. - (ho/np.sqrt(np.power(x1-x2+r,2)+np.power(yo,2))))*np.exp(-.5*k*b*np.power(np.sqrt(np.power(x1-x2+r,2)+np.power(yo,2))-ho, 2))
fdr = lambda x1, x2, r:-1.*c*k*(x1 - x2 + r)*np.exp(-.5*k*b*np.power(np.power(x1-x2+r,2)+np.power(yo,2), 2))
#print fdr(0,0)
f, err = dblquad(fdr, -hL, hL, lambda x2:-hL, lambda x2:hL, args=[0.], epsabs=0, epsrel=1.e-13)
print f, err
# Scan over multiple values of Delta r
Dr_list = np.linspace(-22, 22, 100).tolist()
f_list = [dblquad(fdr, -hL, hL, lambda x2:-hL, lambda x2:hL, args = [r], epsabs=0, epsrel=1.e-13) for r in Dr_list]
f_arr = np.array(f_list)
# Graph scan over area
fig, ax = plt.subplots(figsize=(10,7))
ax.errorbar(np.array(Dr_list)*25., f_arr[:,0]*.1644*16, yerr=f_arr[:,1]*.1644)
#ax.set_xlim((-22,22))
ax.set_xlabel(r'Separation of MT centers $\Delta r$ (nm)')
ax.set_ylabel(r'Total crosslinker force $F_{\rm cl}$ (pN)')
plt.show()
# Partition function for parallel filaments as a function of delta r
Ndr = lambda x1, x2, r:c*np.exp(-.5*k*b*np.power(np.sqrt(np.power(x1-x2+r,2)+np.power(yo,2))-ho, 2))
N_list = [dblquad(Ndr, -hL, hL, lambda x2:-hL, lambda x2:hL, args = [r], epsabs=0, epsrel=1.e-13) for r in Dr_list]
N_arr = np.array(N_list)
# Graph number of crosslinkers based off partition function
fig, ax = plt.subplots(figsize=(10,7))
ax.errorbar(np.array(Dr_list)*25., N_arr[:,0], yerr=N_arr[:,1])
#ax.set_xlim((-22,22))
ax.set_xlabel(r'Separation of MT centers $\Delta r$ (nm)')
ax.set_ylabel(r'Total number of crosslinkers $N_{\rm cl}$')
plt.show()
# +
# Canonical force calculation
fig, ax = plt.subplots(figsize=(10,7))
ax.set_xlim((40,500))
ax.set_ylim((-4, 0))
ax.plot(np.array(Dr_list)*25., np.divide(f_arr[:,0],N_arr[:,0]))
ax.set_xlabel(r'Separation of MT COMs $\Delta r$ (nm)')
ax.set_ylabel(r'Total force from crosslinkers $F_{\rm cl}$ (pN)')
plt.show()
# -
# Crosslinkers with some equilibrium length
ho = 2.28
k = 31.25
Dr_list = np.linspace(-22, 22, 100).tolist()
fho_list = [dblquad(fdr, -hL, hL, lambda x2:-hL, lambda x2:hL, args = [r], epsabs=0, epsrel=1.e-13) for r in Dr_list]
fho_arr = np.array(f_list)
# Graph scan over area
fig, ax = plt.subplots(figsize=(10,7))
ax.errorbar(np.array(Dr_list)*25., fho_arr[:,0]*.1644, yerr=fho_arr[:,1]*.1644)
ax.errorbar(np.array(Dr_list)*25., f_arr[:,0]*.1644, yerr=f_arr[:,1]*.1644, c='r')
#ax.set_xlim((-22,22))
ax.set_xlabel(r'Separation of MT COMs $\Delta r$ (nm)')
ax.set_ylabel(r'Total crosslinker force $F_{\rm cl}$ (pN)')
plt.show()
fig, ax = plt.subplots(figsize=(10,7))
Fs = 6.08 # Crosslinker stall force
maxx = np.sqrt(np.power(Fs/k,2)-np.power(yo,2))
ax.errorbar(np.array(Dr_list)*25., -k*maxx*N_arr[:,0]*.1644, yerr=N_arr[:,1]*.1644)
#ax.set_xlim((-22,22))
ax.set_xlabel(r'Separation of MT centers $\Delta r$ (nm)')
ax.set_ylabel(r'Total motor force $F_{\rm cl}$ (pN)')
plt.show()
fig, ax = plt.subplots(figsize=(10,7))
Fs = 6.08 # Crosslinker stall force
maxx = np.sqrt(np.power(Fs/k,2)-np.power(yo,2))
ax.errorbar(N_arr[:,0], -k*maxx*N_arr[:,0]*.1644, yerr=N_arr[:,1]*.1644)
#ax.set_xlim((-22,22))
ax.set_xlabel(r'Average number of motors')
ax.set_ylabel(r'Total motor force $F_{\rm cl}$ (pN)')
plt.show()
| notebooks/integral_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <h1><center>Decision Trees</center></h1>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# In this lab exercise, you will learn a popular machine learning algorithm, Decision Tree. You will use this classification algorithm to build a model from historical data of patients, and their response to different medications. Then you use the trained decision tree to predict the class of a unknown patient, or to find a proper drug for a new patient.
# -
# <h1>Table of contents</h1>
#
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <ol>
# <li><a href="#about_dataset">About the dataset</a></li>
# <li><a href="#downloading_data">Downloading the Data</a></li>
# <li><a href="#pre-processing">Pre-processing</a></li>
# <li><a href="#setting_up_tree">Setting up the Decision Tree</a></li>
# <li><a href="#modeling">Modeling</a></li>
# <li><a href="#prediction">Prediction</a></li>
# <li><a href="#evaluation">Evaluation</a></li>
# <li><a href="#visualization">Visualization</a></li>
# </ol>
# </div>
# <br>
# <hr>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Import the Following Libraries:
# <ul>
# <li> <b>numpy (as np)</b> </li>
# <li> <b>pandas</b> </li>
# <li> <b>DecisionTreeClassifier</b> from <b>sklearn.tree</b> </li>
# </ul>
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
import numpy as np
import pandas as pd
from sklearn.tree import DecisionTreeClassifier
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <div id="about_dataset">
# <h2>About the dataset</h2>
# Imagine that you are a medical researcher compiling data for a study. You have collected data about a set of patients, all of whom suffered from the same illness. During their course of treatment, each patient responded to one of 5 medications, Drug A, Drug B, Drug c, Drug x and y.
# <br>
# <br>
# Part of your job is to build a model to find out which drug might be appropriate for a future patient with the same illness. The feature sets of this dataset are Age, Sex, Blood Pressure, and Cholesterol of patients, and the target is the drug that each patient responded to.
# <br>
# <br>
# It is a sample of binary classifier, and you can use the training part of the dataset
# to build a decision tree, and then use it to predict the class of a unknown patient, or to prescribe it to a new patient.
# </div>
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <div id="downloading_data">
# <h2>Downloading the Data</h2>
# To download the data, we will use !wget to download it from IBM Object Storage.
# </div>
# -
# !wget -O _datasets/drug200.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/drug200.csv
# __Did you know?__ When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
# now, read data using pandas dataframe:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
my_data = pd.read_csv("_datasets/drug200.csv", delimiter=",")
my_data[0:5]
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <div id="practice">
# <h3>Practice</h3>
# What is the size of data?
# </div>
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# write your code here
# -
# <div href="pre-processing">
# <h2>Pre-processing</h2>
# </div>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Using <b>my_data</b> as the Drug.csv data read by pandas, declare the following variables: <br>
#
# <ul>
# <li> <b> X </b> as the <b> Feature Matrix </b> (data of my_data) </li>
# <li> <b> y </b> as the <b> response vector (target) </b> </li>
# </ul>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Remove the column containing the target name since it doesn't contain numeric values.
# -
X = my_data[['Age', 'Sex', 'BP', 'Cholesterol', 'Na_to_K']].values
X[0:5]
# As you may figure out, some features in this dataset are categorical such as __Sex__ or __BP__. Unfortunately, Sklearn Decision Trees do not handle categorical variables. But still we can convert these features to numerical values. __pandas.get_dummies()__
# Convert categorical variable into dummy/indicator variables.
# +
from sklearn import preprocessing
le_sex = preprocessing.LabelEncoder()
le_sex.fit(['F','M'])
X[:,1] = le_sex.transform(X[:,1])
le_BP = preprocessing.LabelEncoder()
le_BP.fit([ 'LOW', 'NORMAL', 'HIGH'])
X[:,2] = le_BP.transform(X[:,2])
le_Chol = preprocessing.LabelEncoder()
le_Chol.fit([ 'NORMAL', 'HIGH'])
X[:,3] = le_Chol.transform(X[:,3])
X[0:5]
# -
# Now we can fill the target variable.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
y = my_data["Drug"]
y[0:5]
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <hr>
#
# <div id="setting_up_tree">
# <h2>Setting up the Decision Tree</h2>
# We will be using <b>train/test split</b> on our <b>decision tree</b>. Let's import <b>train_test_split</b> from <b>sklearn.cross_validation</b>.
# </div>
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
from sklearn.model_selection import train_test_split
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Now <b> train_test_split </b> will return 4 different parameters. We will name them:<br>
# X_trainset, X_testset, y_trainset, y_testset <br> <br>
# The <b> train_test_split </b> will need the parameters: <br>
# X, y, test_size=0.3, and random_state=3. <br> <br>
# The <b>X</b> and <b>y</b> are the arrays required before the split, the <b>test_size</b> represents the ratio of the testing dataset, and the <b>random_state</b> ensures that we obtain the same splits.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
X_trainset, X_testset, y_trainset, y_testset = train_test_split(X, y, test_size=0.3, random_state=3)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <h3>Practice</h3>
# Print the shape of X_trainset and y_trainset. Ensure that the dimensions match
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# your code
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Print the shape of X_testset and y_testset. Ensure that the dimensions match
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# your code
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <hr>
#
# <div id="modeling">
# <h2>Modeling</h2>
# We will first create an instance of the <b>DecisionTreeClassifier</b> called <b>drugTree</b>.<br>
# Inside of the classifier, specify <i> criterion="entropy" </i> so we can see the information gain of each node.
# </div>
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
drugTree = DecisionTreeClassifier(criterion="entropy", max_depth = 4)
drugTree # it shows the default parameters
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Next, we will fit the data with the training feature matrix <b> X_trainset </b> and training response vector <b> y_trainset </b>
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
drugTree.fit(X_trainset,y_trainset)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <hr>
#
# <div id="prediction">
# <h2>Prediction</h2>
# Let's make some <b>predictions</b> on the testing dataset and store it into a variable called <b>predTree</b>.
# </div>
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
predTree = drugTree.predict(X_testset)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# You can print out <b>predTree</b> and <b>y_testset</b> if you want to visually compare the prediction to the actual values.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
print (predTree [0:5])
print (y_testset [0:5])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <hr>
#
# <div id="evaluation">
# <h2>Evaluation</h2>
# Next, let's import <b>metrics</b> from sklearn and check the accuracy of our model.
# </div>
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
from sklearn import metrics
import matplotlib.pyplot as plt
print("DecisionTrees's Accuracy: ", metrics.accuracy_score(y_testset, predTree))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# __Accuracy classification score__ computes subset accuracy: the set of labels predicted for a sample must exactly match the corresponding set of labels in y_true.
#
# In multilabel classification, the function returns the subset accuracy. If the entire set of predicted labels for a sample strictly match with the true set of labels, then the subset accuracy is 1.0; otherwise it is 0.0.
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Practice
# Can you calculate the accuracy score without sklearn ?
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# your code here
# -
# <hr>
#
# <div id="visualization">
# <h2>Visualization</h2>
# Lets visualize the tree
# </div>
# +
# Notice: You might need to uncomment and install the pydotplus and graphviz libraries if you have not installed these before
# # !conda install -c conda-forge pydotplus -y
# # !conda install -c conda-forge python-graphviz -y
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
from sklearn.externals.six import StringIO
import pydotplus
import matplotlib.image as mpimg
from sklearn import tree
# %matplotlib inline
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
dot_data = StringIO()
filename = "drugtree.png"
featureNames = my_data.columns[0:5]
targetNames = my_data["Drug"].unique().tolist()
out=tree.export_graphviz(drugTree,feature_names=featureNames, out_file=dot_data, class_names= np.unique(y_trainset), filled=True, special_characters=True,rotate=False)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
graph.write_png(filename)
img = mpimg.imread(filename)
plt.figure(figsize=(100, 200))
plt.imshow(img,interpolation='nearest')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <h2>Want to learn more?</h2>
#
# IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: <a href="http://cocl.us/ML0101EN-SPSSModeler">SPSS Modeler</a>
#
# Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at <a href="https://cocl.us/ML0101EN_DSX">Watson Studio</a>
#
# <h3>Thanks for completing this lesson!</h3>
#
# <h4>Author: <a href="https://ca.linkedin.com/in/saeedaghabozorgi"><NAME></a></h4>
# <p><a href="https://ca.linkedin.com/in/saeedaghabozorgi"><NAME></a>, PhD is a Data Scientist in IBM with a track record of developing enterprise level applications that substantially increases clients’ ability to turn data into actionable knowledge. He is a researcher in data mining field and expert in developing advanced analytic methods like machine learning and statistical modelling on large datasets.</p>
#
# <hr>
#
# <p>Copyright © 2018 <a href="https://cocl.us/DX0108EN_CC">Cognitive Class</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.</p>
| 6 - Machine Learning With Python/3.2 Decision Trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pre-processing Texts
#
# This notebook is used to cleaned the text and put into chunks based on the model requirements.
#
# Current process:
# * Clean off top and bottom unnecessary words e.g. table of content
# * Chunk into 450 tokens (for BERT) - can be changed later depends on the model
# * Put into rows with Author name as csv
#
import nltk
import numpy as np
import random
import pandas as pd
import re
from collections import OrderedDict, defaultdict
nltk.download('punkt')
tokenizer = nltk.data.load('tokenizers/punkt/english.pickle')
# ## Support Functions
def split_by_chapter(text):
thisdict = defaultdict()
# ALERT - manual check if there is 3 digit chapters
# split by chapter with one or two digits number with newline
split_text = re.split("chapter (\d{1,2})+", text, flags=re.IGNORECASE)
i = 0
for s in split_text:
if i%2 ==0 or i==0:
Chapter = "chapter_" + str(int(i/2+1))
thisdict[Chapter] = s
i+=1
return thisdict
# ## Read Data
# ALERT - manual check your file location
f = open("Dataset/Original_Book/Mark/Pride_and_Prejudice_Jane_Austen.txt", "r", encoding="utf8")
book = f.read().replace('\n', ' ').replace('_', '')
# ### Clean top and botten unnecessary words
# ALERT - manual check where you want to start
#book = book.split('chapter 1 ')[1]
book = book.split('Chapter 1 ')[1]
# +
# ALERT - manual check where you want to end
#book = book.split('End of the Project Gutenberg EBook ')[0]
# -
# ### Clean words, space, newline
book = re.sub('([.,!?()""])', r' \1 ', book)
book_dict = split_by_chapter(book)
for key,item in book_dict.items():
## remove newline and space at beginning and end
book_dict[key] = re.sub(' +', ' ', item)
book_dict.keys()
# ALERT - manual check quickly to scan through
book_dict['chapter_18']
import csv
Author = "Jane_Austen"
Book = "Pride_and_Prejudice"
csvname = 'Dataset/Cleaned_CSV/Jane_Austen/'+Author+'_'+Book+'_chapter.csv'
with open(csvname, 'w') as csv_file:
fieldnames = ['chapter','text']
writer = csv.DictWriter(csv_file, fieldnames=fieldnames)
writer.writeheader()
writer = csv.writer(csv_file)
for key, value in book_dict.items():
writer.writerow([key, value])
# ### Chunk data into 128 tokens each
#
# BERT can hadle up to 512.
# ALERT - I change to 128 tokens, feel free to change your chunk size
# First check the chapter length
for key,item in book_dict.items():
if len(item.split()) > 128:
print(key, len(item.split()))
# +
# create two empty lists for df
chapterindex = []
text = []
for key,item in book_dict.items():
# wordcount starts from 0
wordcount = 0
# keep adding sentences until 128 tokens
chapter_chunk_text = ""
# sentences in each chapter
sentences = tokenizer.tokenize(item)
# loop through each sentence
for sent in sentences:
# check the word count per sentence
wv = len(sent.split())
# if adding this sentence makes it over 128 tokens
# ALERT - I change to 128 tokens, feel free to change your chunk size
if wordcount + wv >= 128:
# push row and clean cache
chapterindex.append(key)
text.append(chapter_chunk_text)
wordcount = 0
chapter_chunk_text = sent
# if not exceed, append the text and add wordcount
chapter_chunk_text += ' ' + sent
wordcount += wv
# once a chapter finished, push all rest text
chapterindex.append(key)
text.append(chapter_chunk_text)
# -
# create dataframe
book_df = pd.DataFrame(
{'chapter': chapterindex,
'text': text
})
# create total words count per row
book_df['totalwords'] = book_df['text'].str.split().str.len()
book_df.iloc[0]['text']
book_df.shape
book_df['totalwords'].sum()
# ALERT - depending on how well tokenizer.tokenize(item)
# split the sentence, some sentences can be VERY long because the
# function fails to identify the sentence end, which makes it possible
# to be over 300 token limit
book_df.describe()
book_df['author'] = 'Jane_Austen'
# ### Save as csv with author name
Author = "Jane_Austen"
Book = "Pride_and_Prejudice"
book_df.to_csv(Author+"_"+Book+".csv")
| Preprocessing/.ipynb_checkpoints/Pre-processing_byChapter-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="OdhkmJ0SJDb7" executionInfo={"status": "ok", "timestamp": 1645233031857, "user_tz": -540, "elapsed": 200, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}}
import numpy as np
import torch
import torch.optim as optim
import matplotlib.pyplot as plt
# + [markdown] id="zIt_Yt1XJcyo"
# ## **Objective function**
# $$
# f(x, y) = x^2 + y^2
# $$
# <br />
#
# ### with saddle point
# $$
# f(x, y) = x^2 - y^2
# $$
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="kndDwU7u7AKj" executionInfo={"status": "ok", "timestamp": 1645233043834, "user_tz": -540, "elapsed": 11580, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}} outputId="089e3050-fd86-476d-ed1c-6daaa3d47b04"
x = np.arange(-50, 50, 0.01)
y = np.arange(-50, 50, 0.01)
X, Y = np.meshgrid(x, y)
Z = X**2 + Y**2
fig = plt.figure()
ax = fig.add_subplot(projection='3d')
ax.plot_wireframe(X,Y,Z)
plt.show()
fig.savefig("/content/drive/MyDrive/log/images/fig1.png")
# + [markdown] id="XWROTio78NSC"
# # **SGD**
#
# if not use momentum
# $$
# w_t = w_{t-1} - \gamma \nabla_w f(w_{t-1})
# $$
#
# if use momentum
# $$
# \begin{align*}
# v_t &= \mu v_{t-1} + \nabla_wf(w_{t-1}) \\
# w_t &= w_{t-1} - \gamma v_t
# \end{align*}
# $$
#
# if use momentum and netrov
# $$
# \begin{align*}
# v_t &= \mu v_{t-1} + \nabla_wf(w_{t-1}) \\
# w_t &= w_{t-1} - \gamma (\nabla_wf(w_{t-1}) + \mu v_t)
# \end{align*}
# $$
# + colab={"base_uri": "https://localhost:8080/"} id="p1hckYfkJ8KK" executionInfo={"status": "ok", "timestamp": 1645233043836, "user_tz": -540, "elapsed": 11, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}} outputId="3fe0e030-7d21-4751-f162-6c1427658c3b"
x = torch.tensor(20.0, requires_grad = True)
y = torch.tensor(0.0, requires_grad = True)
op = optim.SGD([x, y], lr=0.01, momentum=0.9, nesterov=True)
print("op: \n", op)
def lr_function(mode, epoch):
if mode == "linear":
return -0.0002 * epoch + 0.01
iteration = 30
scheduler = optim.lr_scheduler.LambdaLR(op, lr_lambda = lambda epoch: 100 * lr_function(mode="linear", epoch=epoch))
lr = []
fx, fy = [], []
g_fx, g_fy = [], []
fx.append(x.item())
fy.append(y.item())
f_loss = []
for i in range(iteration):
op.zero_grad()
params = x**2 + y**2
params.backward(retain_graph=True)
g_fx.append(x.grad.item())
g_fy.append(y.grad.item())
op.step()
lr.append(op.param_groups[0]['lr'])
fx.append(x.item())
fy.append(y.item())
loss = params.item()
f_loss.append(loss)
scheduler.step()
# + [markdown] id="YaGpvXPBKSpx"
# # **Plot 2D**
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="fAInU_4Y6pdj" executionInfo={"status": "ok", "timestamp": 1645233056446, "user_tz": -540, "elapsed": 12617, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}} outputId="87378526-cfe0-49a3-e929-a6e9f5249587"
plt.plot(fx, fy, 'o-')
plt.contour(X, Y, Z)
plt.xlim(-50, 50)
plt.ylim(-50, 50)
plt.xlabel("x")
plt.ylabel("y")
plt.savefig("/content/drive/MyDrive/log/images/fig2.png")
plt.show()
# + [markdown] id="L1OvVit-V1Vg"
# # **Plot iteration vs gradient(objective function)**
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="WNwtCyxbVpoe" executionInfo={"status": "ok", "timestamp": 1645233057048, "user_tz": -540, "elapsed": 608, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}} outputId="d494f3a0-e263-42bc-c9d1-33cf30482dd5"
iteration = np.arange(0, 30, 1)
plt.plot(iteration, g_fx)
plt.xlabel("iteration")
plt.ylabel("gradient")
plt.title("gradient_x")
plt.savefig("/content/drive/MyDrive/log/images/fig3.png")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="tJ7FNVZJWMzY" executionInfo={"status": "ok", "timestamp": 1645233057049, "user_tz": -540, "elapsed": 11, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}} outputId="03f7f308-cd4d-4f30-b5f4-fbf73868c378"
plt.plot(iteration, g_fy)
plt.xlabel("iteration")
plt.ylabel("gradient")
plt.title("gradient_y")
plt.savefig("/content/drive/MyDrive/log/images/fig4.png")
plt.show()
# + [markdown] id="QsTVhhheKNOK"
# # **Plot iteration vs loss**
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="ckIt6pU9FsbE" executionInfo={"status": "ok", "timestamp": 1645233057050, "user_tz": -540, "elapsed": 9, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}} outputId="16442e01-1c76-4eac-ff6d-2bf0dcdf07b9"
iteration = np.arange(0, 30, 1)
plt.plot(iteration, f_loss)
plt.xlabel("iteration")
plt.ylabel("loss")
plt.savefig("/content/drive/MyDrive/log/images/fig5.png")
plt.show()
# + [markdown] id="0yUIyVzi6uC2"
# # **Plot learning rate vs gradient(SGD)**
# + id="6hQowodxRLbJ" executionInfo={"status": "ok", "timestamp": 1645233057461, "user_tz": -540, "elapsed": 419, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}}
grad_x = []
grad_x.append(-fx[0] / lr[0])
for i in range(1, len(iteration)):
grad_x.append(-(fx[i - 1] - fx[i]) / lr[i])
grad_y = []
grad_y.append(-fy[0] / lr[0])
for i in range(1, len(iteration)):
grad_y.append(-(fy[i - 1] - fy[i]) / lr[i])
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="m64FwI5_61Pb" executionInfo={"status": "ok", "timestamp": 1645233057462, "user_tz": -540, "elapsed": 17, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}} outputId="d878e646-b434-4247-e6b3-a5a7a89eda27"
plt.plot(iteration, grad_x)
plt.xlabel("iteration")
plt.ylabel("gradient")
plt.title("gradient_x")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="JMTmbxHR7FV6" executionInfo={"status": "ok", "timestamp": 1645233057463, "user_tz": -540, "elapsed": 17, "user": {"displayName": "ma saka", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07870361181562380675"}} outputId="16069666-e229-43f0-de21-fed1fdf32640"
plt.plot(iteration, grad_y)
plt.xlabel("iteration")
plt.ylabel("gradient")
plt.title("gradient_y")
plt.show()
| SGDonPytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center>
# <h1><b>Word Embedding Based Answer Evaluation System for Online Assessments (WebAES)</b></h1>
# <h3>A smart system to automate the process of answer evaluation in online assessments.</h3>
# <h5> LDA + BERT Model for WebAES</h5>
# +
# To perform text pre-processing
import string
# Natural Language Toolkit
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
# Set of stopwords in English
en_stopwords = set(stopwords.words('english'))
# To load text corpus and pre-trained LDA model
from gensim import corpora, models
import gensim.downloader as api
# To perform sentence encoding using BERT model
from sentence_transformers import SentenceTransformer
# To determine similarity between 2 vectors
from sklearn.metrics.pairwise import cosine_similarity
# +
# Load pre-trained and saved LDA model
lda_model = models.LdaModel.load('WebAES_LDA_Model.model')
# Load text8 corpus and convert to list of documents
text8_corpus = api.load('text8')
text8_corpus = [doc for doc in text8_corpus]
# List containing list of tokens from each document of text8 corpus
list_of_list_of_tokens = []
# For each document in the text8 corpus
for i in range(len(text8_corpus)):
# Remove stopwords from each document
text8_corpus[i] = [w for w in text8_corpus[i] if w not in en_stopwords]
# Add list of tokens for document to list of list of tokens
list_of_list_of_tokens.append(text8_corpus[i])
dictionary_LDA = corpora.Dictionary(list_of_list_of_tokens)
# -
# Function to perform text pre-processing operations
def preprocess(text):
# Remove punctuations
text = text.translate(str.maketrans('', '', string.punctuation)).lower()
# Remove stopwords
text = ' '.join([w for w in text.split() if not w.lower() in en_stopwords])
# Split text into list of tokens
text_tokens = text.split()
# Return list of tokens
return text_tokens
# Function to extract topic from text
def get_topic_prob(text_tokens):
max_prob_topic, max_prob = 0, 0
# Get topic probabilities for given text using LDA model
topic_probs = lda_model[dictionary_LDA.doc2bow(text_tokens)]
# Select topic with highest probability
for topic in topic_probs:
topic_index, topic_prob = topic[0], topic[1]
if topic_prob > max_prob:
max_prob = topic_prob
max_prob_topic = topic_index
# Return topic with max probability and probabilty of that topic
return max_prob_topic, max_prob
# Get document embeddings for list of documents
def get_bert_embeddings(docs):
# Load pre-trained BERT model
BERT_model = SentenceTransformer('bert-base-nli-mean-tokens')
# Encode documents using BERT model
doc_embeddings = BERT_model.encode(docs)
# Return document embeddings
return doc_embeddings
# Determine similarity between documents
def similarity(doc_embeddings):
# Similarity score based on cosine similarity measure
sim_score = cosine_similarity([doc_embeddings[0]], doc_embeddings[1:])[0][0]
# Return similarity score
return sim_score
# Function to automatically evaluate test
def evaluate(expected, response):
# Pre-rpocessing for expected answer
exp_ans_tokens = preprocess(expected)
# Get topic for expected answer
exp_ans_topic, exp_ans_topic_prob = get_topic_prob(exp_ans_tokens)
# Pre-processing for student's response
stu_ans_tokens = preprocess(response)
# Get topic for student's answer
stu_ans_topic, stu_ans_topic_prob = get_topic_prob(stu_ans_tokens)
# List of documents
docs = [expected, response]
# Get document embeddings for expected answer and student response
doc_embeddings = get_bert_embeddings(docs)
# Get similarity score based on documents embeddings
sim_score = similarity(doc_embeddings)
# Calculate marks iff topics match for expected answer and student's response
if stu_ans_topic==exp_ans_topic:
marks = (stu_ans_topic_prob/exp_ans_topic_prob)*sim_score*10
# If topics do not match, marks are 0
else:
marks = 0
# Return marks scored
return marks
# Function to start test
def take_test(question, max_marks, expected_answer):
# Get student's response
student_response = input('\n' + question + ' ({} marks)\n\n'.format(max_marks))
# Determine score for student's response
score = round(evaluate(expected_answer, student_response), 2)
# Display marks scored
print('\nYou have scored {} marks out of {}.'.format(score, max_marks))
# +
# Sample question
question = 'Give PEAS description for medical diagnosis system.'
# Maximum marks for question
max_marks = 10
# Answer expected by faculty
expected_answer = '''The performance measure for medical diagnosis system may include the number of patients healed by
correctly and accurately diagnosing diseases. For example, the performance measure may be the percentage of cases diagnosed
correctly by the system. The environment for a medical diagnosis system includes patients and their vital signs. This
environment is fully observable, dynamic and complete. The actuators include display screens and alert systems that send
feedback to doctors. Sensors include equipment including medical sensors as well as medical images.'''
# Start test and display result
take_test(question, max_marks, expected_answer)
| notebooks/WebAES-LDA-BERT-Evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# name: python3
# ---
# +
import os
from pathlib import Path
import shutil
import cv2
import numpy as np
from PIL import Image
from sklearn.model_selection import train_test_split
# -
# ## Transfer annotated data to cvat_annotation
#
# * Removes images with no annotations
# * Combines multiple annotation folders
# + tags=[]
data_dir = Path.cwd().parents[1] / "oidv6_data_converted_11classes"
for path in (data_dir / "images" / "train").glob("*_[0-9]*_none.jpg"):
os.remove(path)
os.remove(data_dir / "labels" / "train" / f"{path.stem}.txt")
# +
class_name = "tissue"
class_names = ["tissue", "wheelchair", "box", "table"] # tissue
# class_names = ["cart", "burner", "gas_cylinder", "wok", "box"] # cart
data_dirs = [
Path(r"C:\Users\Admin\Downloads") / sub_dir
for sub_dir in (
"task_label tissue_video_cut_1-2021_06_04_01_39_33-yolo 1.1",
)
]
output_dir = Path.cwd().parents[1] / "cvat_annotation" / class_name
data_split_dir = output_dir / "data_split"
train_dir = data_split_dir / "train"
test_dir = data_split_dir / "test"
train_images_dir = train_dir / "images"
train_labels_dir = train_dir / "labels"
test_images_dir = test_dir / "images"
test_labels_dir = test_dir / "labels"
output_dir.mkdir(parents=True, exist_ok=True)
train_images_dir.mkdir(parents=True, exist_ok=True)
train_labels_dir.mkdir(parents=True, exist_ok=True)
test_images_dir.mkdir(parents=True, exist_ok=True)
test_labels_dir.mkdir(parents=True, exist_ok=True)
# +
# Convert all to jpg
for data_dir in data_dirs:
for path in (data_dir / "obj_train_data").glob("*.png"):
im = Image.open(path)
im.convert("RGB").save(path.with_suffix(".jpg"), "JPEG")
os.remove(path)
count = 585
for data_dir in data_dirs:
for path in (data_dir / "obj_train_data").glob("*.jpg"):
# shutil.copy(path, output_dir / path.name)
# shutil.copy(path.with_suffix(".txt"), output_dir / f"{path.stem}.txt")
empty = False
with open(path.with_suffix(".txt"), "r") as infile:
empty = not infile.read()
if not empty:
shutil.copy(path, output_dir / f"{class_name}_scraped_{count}.jpg")
shutil.copy(path.with_suffix(".txt"), output_dir / f"{class_name}_scraped_{count}.txt")
count += 1
for path in output_dir.glob("*.jpg"):
im = Image.open(path)
try:
if im.is_animated:
im.convert("RGB").save(path, "JPEG")
except AttributeError:
pass
# -
for path in output_dir.glob("*.txt"):
with open(path, "r+") as file:
l = [x.split() for x in file.read().strip().splitlines()]
file.seek(0)
for line_parts in np.unique(l, axis=0):
file.write(f"{class_names[int(line_parts[0])]} {' '.join(line_parts[1:])}\n")
# Remove unannotated images
for path in output_dir.glob("*.jpg"):
empty = False
with open(output_dir / f"{path.stem}.txt", "r") as infile:
empty = not infile.read()
if empty:
os.remove(path)
os.remove(output_dir / f"{path.stem}.txt")
# Split data
image_list = list(output_dir.glob("*.jpg"))
image_train, image_test = train_test_split(image_list, test_size=0.3, random_state=1234)
# +
for path in image_train:
shutil.copy(path, train_images_dir / path.name)
shutil.copy(path.with_suffix(".txt"), train_labels_dir / f"{path.stem}.txt")
for path in image_test:
shutil.copy(path, test_images_dir / path.name)
shutil.copy(path.with_suffix(".txt"), test_labels_dir / f"{path.stem}.txt")
# -
for path in train_labels_dir.iterdir():
with open(path, "r+") as file:
l = [x.split() for x in file.read().strip().splitlines()]
file.seek(0)
for line_parts in np.unique(l, axis=0):
file.write(f"{class_names[int(line_parts[0])]} {' '.join(line_parts[1:])}\n")
for path in test_labels_dir.iterdir():
with open(path, "r+") as file:
l = [x.split() for x in file.read().strip().splitlines()]
file.seek(0)
for line_parts in np.unique(l, axis=0):
file.write(f"{class_names[int(line_parts[0])]} {' '.join(line_parts[1:])}\n")
# ## Transfer from syndata to oidv6_data_converted
#
# Single class
# +
class_name = "cart"
data_dir = Path.cwd().parent / "hawking_output_dir"
output_dir = Path.cwd().parents[1] / f"oidv6_data_converted_{class_name}"
output_image_dir = output_dir / "images"
output_label_dir = output_dir / "labels"
output_image_dir.mkdir(parents=True, exist_ok=True)
output_label_dir.mkdir(parents=True, exist_ok=True)
# -
subset = "train"
(output_image_dir / subset).mkdir(parents=True, exist_ok=True)
(output_label_dir / subset).mkdir(parents=True, exist_ok=True)
for path in (data_dir / "images").glob("*.jpg"):
path_stem = path.stem
shutil.copy(path, output_image_dir / subset / f"{class_name}_{path_stem}.jpg")
with open(data_dir / "annotations" / f"{path_stem.split('_')[0]}.txt", "r") as infile, open(
output_label_dir / subset / f"{class_name}_{path_stem}.txt", "w"
) as outfile:
l = [x.split() for x in infile.read().strip().splitlines()]
for line_parts in np.unique(l, axis=0):
outfile.write(f"0 {' '.join(line_parts[1:])}\n")
# + tags=[]
for path in (output_image_dir / "train").glob("*_none.jpg"):
os.remove(path)
os.remove(output_label_dir / "train" / f"{path.stem}.txt")
# -
# ## Transfer from cvat annotated to oidv6_data_converted
#
# Single class
# +
class_name = "cart"
rewrite = True
data_dir = Path.cwd().parents[1] / "cvat_annotation"
output_dir = Path.cwd().parents[1] / f"oidv6_data_converted_{class_name}"
output_image_dir = output_dir / "images"
output_label_dir = output_dir / "labels"
output_image_dir.mkdir(parents=True, exist_ok=True)
output_label_dir.mkdir(parents=True, exist_ok=True)
# -
for subset in ("train", "test"):
(output_image_dir / subset).mkdir(parents=True, exist_ok=True)
(output_label_dir / subset).mkdir(parents=True, exist_ok=True)
for path in (data_dir / class_name / "data_split" / subset / "images").glob("*.jpg"):
shutil.copy(path, output_image_dir / subset / path.name)
if rewrite:
with open(path.parents[1] / "labels" / f"{path.stem}.txt", "r") as infile, open(
output_label_dir / subset / f"{path.stem}.txt", "w"
) as outfile:
l = [x.split() for x in infile.read().strip().splitlines()]
for line_parts in np.unique(l, axis=0):
if line_parts[0] == class_name:
outfile.write(f"0 {' '.join(line_parts[1:])}\n")
else:
shutil.copy(
path.parents[1] / "labels" / f"{path.stem}.txt",
output_label_dir / subset / f"{path.stem}.txt",
)
# ## Transfer from cvat annotated to oidv6_data_converted
#
# Multiclass class
# +
class_name = "cart"
class_names = ["apple", "banana", "orange", "wheelchair", "wok", "box", "table", "tissue", "gas_cylinder", "burner", "cart"]
cvat_data_dir = Path.cwd().parents[1] / "cvat_annotation"
single_class_data_dir = Path.cwd().parents[1] / f"oidv6_data_converted_{class_name}"
output_dir = Path.cwd().parents[1] / f"oidv6_data_converted_{class_name}_multi"
output_image_dir = output_dir / "images"
output_label_dir = output_dir / "labels"
output_image_dir.mkdir(parents=True, exist_ok=True)
output_label_dir.mkdir(parents=True, exist_ok=True)
# -
for subset in ("train", "test"):
(output_image_dir / subset).mkdir(parents=True, exist_ok=True)
(output_label_dir / subset).mkdir(parents=True, exist_ok=True)
for path in (cvat_data_dir / class_name / "data_split" / subset / "images").glob("*.jpg"):
shutil.copy(path, output_image_dir / subset / path.name)
with open(path.parents[1] / "labels" / f"{path.stem}.txt", "r") as infile, open(
output_label_dir / subset / f"{path.stem}.txt", "w"
) as outfile:
l = [x.split() for x in infile.read().strip().splitlines()]
for line_parts in np.unique(l, axis=0):
if line_parts[0] in class_names:
outfile.write(f"{class_names.index(line_parts[0])} {' '.join(line_parts[1:])}\n")
# + tags=[]
subset = "train"
(output_image_dir / subset).mkdir(parents=True, exist_ok=True)
(output_label_dir / subset).mkdir(parents=True, exist_ok=True)
for path in (single_class_data_dir / "images" / subset).glob(f"{class_name}_[0-9]*.jpg"):
shutil.copy(path, output_image_dir / subset / path.name)
with open(single_class_data_dir / "labels" / subset / f"{path.stem}.txt", "r") as infile, open(
output_label_dir / subset / f"{path.stem}.txt", "w"
) as outfile:
l = [x.split() for x in infile.read().strip().splitlines()]
for line_parts in np.unique(l, axis=0):
outfile.write(f"{class_names.index(class_name)} {' '.join(line_parts[1:])}\n")
# -
# ## Transfer from single class oidv6_data_converted to multiclass oidv6_data_converted
# Includes relabeling
# +
class_name = "tissue"
classes =["apple", "banana", "orange", "wheelchair", "wok", "box", "table", "tissue", "gas_cylinder", "burner", "cart"]
data_dir = Path.cwd().parents[1] / f"oidv6_data_converted_{class_name}"
output_dir = Path.cwd().parents[1] / f"oidv6_data_converted_11classes"
# -
for subset in ("train", "test"):
for path in (data_dir / "images" / subset).glob("*.jpg"):
shutil.copy(path, output_dir / "images" / subset / path.name)
with open(data_dir / "labels" / subset / f"{path.stem}.txt", "r") as infile, open(
output_dir / "labels" / subset / f"{path.stem}.txt", "w"
) as outfile:
l = [x.split() for x in infile.read().strip().splitlines()]
for line_parts in np.unique(l, axis=0):
outfile.write(
f"{classes.index(class_name)} {' '.join(line_parts[1:])}\n"
)
| notebooks/transfer_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import collections
from collections import defaultdict
import sys
import json
import random
from jsmin import jsmin
from io import StringIO
import numpy as np
import copy
import importlib
from functools import partial
import math
import os
import compress_pickle
# script_n = os.path.basename(__file__).split('.')[0]
script_n = 'correlation_shares_syn_weight_210420_2share_shuffle_line'
sys.path.insert(0, '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc')
import my_plot
importlib.reload(my_plot)
from my_plot import MyPlotData
from weight_database import WeightDatabase
weightdb = WeightDatabase()
def weight_fn(syn):
z_len = syn['z_length'] - 40
major_axis_length = syn['major_axis_length'] * .9
diameter = max(z_len, major_axis_length)
diameter = int(diameter/40+.5)
diameter *= 40
return diameter
gzdb = '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/gen_db/pfs/gen_201224_setup01_syndb_threshold_10_coalesced.gz'
weightdb.load_syn_db(gzdb,
weight_fn=weight_fn)
mpd = MyPlotData()
hist = defaultdict(int)
weights_db = weightdb.get_weights()
n_pairs = 0
syn_weights = []
avg_data = []
mpd_data = MyPlotData()
areas = [[], []]
syn_weights = []
avg_data = []
mpd_data = MyPlotData()
hist_data = defaultdict(int)
for neuron, pc_weights in weights_db.items():
# print(n)
for pc, weights in pc_weights.items():
if len(weights) != 2:
continue
w1, w2 = weights
# w1 = sum(w1)/len(w1)
w1 = int(w1/40+.5)*40
# w2 = sum(w2)/len(w2)
w2 = int(w2/40+.5)*40
w1 /= 1000
w2 /= 1000
avg = (w1 + w2)/2
avg = int(avg/.04+.5)*.04
syn_weights.append(w1)
syn_weights.append(w2)
mpd.add_data_point(
w=avg,
x='Data',
)
avg_data.append(avg)
hist_data[avg] += 1
mpd_hist_data = MyPlotData()
for k in sorted(hist_data.keys()):
mpd_hist_data.add_data_point(
weight=k,
count=hist_data[k],
model='Data',
)
mpd_hist_data_cdf = mpd_hist_data.to_cdf('count')
# +
# import my_plot_data
# importlib.reload(my_plot_data)
# # from my_plot import MyPlotData
# mpd_shuffle = compute_shuffle_mpd()
# # mpd_shuffle.data
# mpd_shuffle_hist = mpd_shuffle.to_histogram(count_var='w')
def compute_shuffle_mpd():
random.shuffle(syn_weights)
# avg_shuffle = []
mpd_shuffle = MyPlotData()
hist = defaultdict(int)
i = 0
while i+1 <= len(syn_weights):
w1, w2 = syn_weights[i], syn_weights[i+1]
avg = (w1 + w2)/2
avg = int(avg/.04+.5)*.04
# avg_shuffle.append(avg)
hist[avg] += 1
i += 2
mpd_shuffle.add_data_point(
w=avg,
x='Shuffle',
)
mpd_hist_shuffle = MyPlotData()
for k in sorted(hist.keys()):
mpd_hist_shuffle.add_data_point(
weight=k,
count=hist[k],
model='Shuffle',
)
return mpd_hist_shuffle.to_cdf('count')
mpd_hist_shuffle = compute_shuffle_mpd()
mpd_hist_shuffles = MyPlotData()
for i in range(1000):
mpd_hist_shuffle = compute_shuffle_mpd()
mpd_hist_shuffles.append(mpd_hist_shuffle)
# -
mpd_hist_data_cdf.data
# +
mpd_plot_cdf = MyPlotData()
mpd_plot_cdf.append(mpd_hist_data_cdf)
mpd_plot_cdf.append(mpd_hist_shuffles)
importlib.reload(my_plot); my_plot.my_relplot(
mpd_plot_cdf,
x="weight",
y='count',
hue='model',
ci=99,
# save_filename=f'{script_n}_kde.svg',
show=True,
)
# +
importlib.reload(my_plot); my_plot.my_catplot(
mpd,
x="x",
y="w",
# save_filename=f'{script_n}_kde.svg',
show=True,
)
# +
importlib.reload(my_plot); my_plot.my_displot(
mpd,
x="w",
hue='x',
# y="w",
kind='ecdf',
# save_filename=f'{script_n}_kde.svg',
show=True,
)
# -
from scipy import stats
stats.ks_2samp(avg_data, avg_shuffle)
# +
import scipy.stats
scipy.stats.spearmanr(array_w1, array_w2)
#array_w1
# +
import scipy.stats
scipy.stats.pearsonr(array_w1, array_w2)
#array_w1
# +
# EM of a single pf with 2 synapses to a single PC, showing a pair of small and a big synapses
hist = []
n_pairs = 0
for neuron, pc_weights in weights_db.items():
# print(n)
for pc, weights in pc_weights.items():
if len(weights) != 2:
continue
w1, w2 = weights
w1 /= 1000
w2 /= 1000
hist.append((w1, w2, neuron, pc))
hist.sort(key=lambda x: (x[0], x[1]))
for e in hist: print(e)
# +
import compress_pickle
rawdb = compress_pickle.load(gzdb)
sid = 'pf_4495'
syns = rawdb[sid]
# print(syns)
print(sid)
for pc in syns:
pc_syns = syns[pc]
print(pc)
for syn in pc_syns:
print(syn)
print()
# print(f'{pc}: {syns[pc]}\n')
# +
# EM of a single pf with 2 synapses to a single PC, showing a pair of small and a big synapses
hist = []
n_pairs = 0
for neuron, pc_weights in weights_db.items():
# print(n)
for pc, weights in pc_weights.items():
if len(weights) < 5:
continue
hist.append((neuron, pc))
for e in hist: print(e)
# +
import compress_pickle
rawdb = compress_pickle.load(gzdb)
sid = 'pf_1511'
syns = rawdb[sid]
# print(syns)
print(sid)
for pc in syns:
pc_syns = syns[pc]
print(pc)
for syn in pc_syns:
print(syn)
print()
# print(f'{pc}: {syns[pc]}\n')
# +
#
hist = []
n_pairs = 0
for neuron, pc_weights in weights_db.items():
# print(n)
for pc, weights in pc_weights.items():
if len(weights) < 3:
continue
syns = rawdb[neuron]
pc_syns = syns[pc]
min_z, max_z = (9999, 0)
for s in pc_syns:
# print(s)
min_z = min(min_z, s['syn_loc'][2])
max_z = max(max_z, s['syn_loc'][2])
# print((min_z, max_z)); asdf
if max_z - min_z < 10:
# print((min_z, max_z)); asdf
hist.append((neuron, pc))
for e in hist: print(e)
# +
# finding a big synapse
hist = []
n_pairs = 0
for neuron, pc_weights in weights_db.items():
# print(n)
syns = rawdb[neuron]
for pc, weights in pc_weights.items():
for w in weights:
if w < .5:
continue
pc_syns = syns[pc]
for s in pc_syns:
if s['major_axis_length'] > 870:
print(s)
# +
# finding a small synapse
hist = []
n_pairs = 0
for neuron, pc_weights in weights_db.items():
# print(n)
syns = rawdb[neuron]
for pc, weights in pc_weights.items():
for w in weights:
if w > .5:
continue
pc_syns = syns[pc]
for s in pc_syns:
if s['major_axis_length'] > 50 and s['major_axis_length'] < 100:
print(s)
# +
# counting synapses
n_pairs = 0
n_syns = 0
for neuron, pc_weights in weights_db.items():
# print(n)
for pc, weights in pc_weights.items():
n_pairs += 1
n_syns += len(weights)
print(f'n={n_pairs} pairs')
print(f'n={n_syns} syns')
| analysis/pfs_pc_analysis/weight/correlation_syn_diameter_dual_210423_shuffle_line.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def navigate_to_baseurl(driver,url):
base_url = url_concat(url)
driver.get(base_url)
driver.implicitly_wait(10)
def url_concat(url):
urlstr= url
return urlstr
def init_web_driver():
options = webdriver.ChromeOptions()
options.add_argument("--disable-notifications");
options.add_argument("start-maximized")
options.add_experimental_option("excludeSwitches", ["enable-automation"])
options.add_experimental_option('useAutomationExtension', False)
driver=webdriver.Chrome(r"C:/Users/admin/OneDrive/Documents/Sindhu/Software/Selenium/Driver/chromedriver.exe",options=options)
return driver
# !pip install selenium
# +
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.common.action_chains import ActionChains
from selenium.webdriver.common.by import By
import csv
driver = init_web_driver()
url = "https://www.carvana.com/cars"
index=0
navigate_to_baseurl(driver,url)
input_model = driver.find_element_by_xpath("//*[@id='search-bar-container-wrapper']/div[2]/div/input")
input_model.send_keys('Forester')
#input_modelsend_keys(Keys.RETURN)
button = driver.find_element_by_xpath("//*[@id='search-bar-container-wrapper']/div[2]/button")
button.click()
pop_up_input = driver.find_element_by_xpath("//*[@id='experiment-sheet']")
pop_up_input.send_keys('123<EMAIL>')
pop_up_close = driver.find_element_by_xpath("//*[@id='email-capture']/div[2]/div/div/div[2]/button")
driver.implicitly_wait(10)
ActionChains(driver).move_to_element(pop_up_close).click(pop_up_close).perform()
#Navigating though the list of cars form the current page
#First i need to gte the list of all cars in teh current page
list_cars_page=driver.find_elements(by= 'class_name',value = "year-make")
for i in list_cars_page:
driver = webdriver.Chrome()
actionChains = ActionChains(driver)
actionChains.context_click(your_link).perform()
print(i.text)
# -
| WebScraping-Car.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KunalVaidya99/Image-Colorization-Using-Deep-Learning/blob/main/noGAN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="XqMHHrNKvOU1"
# %tensorflow_version 1.x
# + id="UqgQtKfYvdFK"
from google.colab import drive
drive.mount("/content/gdrive/")
# + id="mI79ymX6vd3y"
import tensorflow as tf
import keras
import keras.backend as K
from keras.layers import (Conv2D,Conv2DTranspose,UpSampling2D,MaxPooling2D,Input
,Activation,Concatenate,BatchNormalization,Flatten,Reshape
,Dense,RepeatVector)
from keras.optimizers import Adam
from keras.utils import plot_model
from keras.models import Model
import numpy as np
from keras.preprocessing.image import load_img
from PIL import Image
from skimage.color import gray2rgb
import os
import random
import matplotlib.pyplot as plt
from keras.callbacks import ModelCheckpoint
# + id="ZAzore6QvgiC"
class DataGenerator(keras.utils.Sequence):
def __init__(self,directory,filenames,batch_size=32,dim=(256,256),n_channels=3,shuffle=True):
self.dim=dim
self.filenames=filenames
self.batch_size=batch_size
self.n_channels=n_channels
self.shuffle=shuffle
self.directory=directory
self.on_epoch_end()
def __len__(self):
return int(np.floor(len(self.filenames)/self.batch_size))
def __getitem__(self,index):
batch_files = self.filenames[index*self.batch_size:(index+1)*self.batch_size]
X,Y,resized,gray = self.__data_generation(batch_files)
return X,Y,resized,gray
def on_epoch_end(self):
if self.shuffle == True :
np.random.shuffle(self.filenames)
def __data_generation(self,batch_files):
X = np.empty((self.batch_size,*self.dim,1))
Y = np.empty((self.batch_size,*self.dim,self.n_channels))
resized = np.empty((self.batch_size,224,224,3))
gray = np.empty((self.batch_size,224,224,3))
count =0
for files in batch_files:
image = Image.open(self.directory + "/" + files)
if (np.array(image).shape==(256,256,3)) :
g_image = image
img = image.resize((224,224))
resized[count] = np.array(img)/255
g = np.array(img.convert("L"))
gray[count] = gray2rgb(g)/255
Y[count,] = (np.array(image))/255
X[count,] = (np.expand_dims(np.array(g_image.convert("L")),axis=-1))/255
count = count + 1
return X,Y,resized,gray
# + id="Wft2e7kcwOWf"
directory = "/content/gdrive/My Drive/val_256"
filenames = os.listdir(directory)
random.shuffle(filenames)
split_size = 0.1
train_files = filenames[int(split_size*len(filenames)):len(filenames)]
test_files = filenames[:int(split_size*len(filenames))]
TrainGenerator = DataGenerator(directory,train_files)
TestGenerator = DataGenerator(directory,test_files)
img_shape = (256,256,1)
# + id="v7sTfJOrwhYn"
| noGAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Social Network Analysis
# ## AO3 User Bookmarks
#
# The following is an analysis of the network created by users bookmarking fics on AO3.
#
# We used the [Social Network Analysis with Python]("https://www.kirenz.com/post/2019-08-13-network_analysis/") article by <NAME> as a reference for our analysis.
#imports
import networkx as nx
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings; warnings.simplefilter('ignore')
import json
import pandas as pd
# First we must read in our data and format it so that we can make a graph of the data.
#read in data
topfic_bookmarks = pd.read_json('../ao3bot/top_fic_bookmarks.json')
topfic_bookmarks.head()
user_df = pd.read_json('../ao3bot/user_bookmarks_final.json')
user_df.head()
# This data is set up such that in `user_df` we have the `user` whose bookmarks we scraped, the `fic_info` of the bookmarks and the `fandoms` associated with those bookmarks. We also have the general stats of their bookmarks in `user_stats`.
#
# In `topfic_bookmarks` we have the information on the top fics in each fandom we have scraped and the users that have bookmarked it. This is the starting file that guiding the scraping for the `user_df` data.
# ### Common Bookmarks
#
# In this analysis we will be looking at the data in this way:
# - Nodes: Fic
# - Edges: A user has bookmarked this node (fic) and this node (fic)
#
# To do this we will need to wrangle the data into the correct format.
# We also want a list of nodes:
nodes = []
nodes_unformatted = []
tf_users = topfic_bookmarks["users"].tolist()
topfics = topfic_bookmarks["fic_info"].tolist()
users = user_df["user"].tolist()
count = 1
for fic in topfics:
for users in tf_users:
for user in users:
l = user_df.loc[user_df["user"] == user["user_name"]]["fic_info"]
if len(l) > 0:
for i in l:
for j in i:
for k in j:
if 'fic_link' in k.keys():
if k["fic_link"] not in nodes_unformatted:
nodes.append([count, k["fic_link"]])
nodes_unformatted.append(k["fic_link"])
count += 1
print(len(nodes))
# nodes_df = pd.DataFrame(nodes, columns=["Id","Label"])
# nodes_df.head()
nodes_df = pd.DataFrame(nodes, columns=["Id","Label"])
nodes_df.head()
nodes_df.to_csv("nodes_and_edges/common_fics_nodes_reformatted.csv", index = False)
nodes_df.loc[nodes_df.Label == '/works/31830625','Id'].tolist()[0]
edges = []
tf_users = topfic_bookmarks["users"].tolist()
topfics = topfic_bookmarks["fic_info"].tolist()
users = user_df["user"].tolist()
for fic in topfics:
for users in tf_users:
for user in users:
l = user_df.loc[user_df["user"] == user["user_name"]]["fic_info"]
if len(l) > 0:
for i in l:
for j in i:
for k in j:
if 'fic_link' in k.keys():
target = nodes_df.loc[nodes_df.Label == k["fic_link"],'Id'].tolist()[0]
src = nodes_df.loc[nodes_df.Label == fic["fic_link"],'Id'].tolist()[0]
edges.append([src, target, "Directed"])
print(len(edges))
# tf_users = topfic_bookmarks["users"].tolist()
# topfics = topfic_bookmarks["fic_info"].tolist()
# users = user_df["user"].tolist()
# fics = []
# for fic in topfics:
# for users in tf_users:
# for user in users:
# l = user_df.loc[user_df["user"] == user["user_name"]]["fic_info"]
# if len(l) > 0:
# for i in l:
# for j in i:
# for k in j:
# if 'fic_link' in k.keys():
# if fic["fic_link"] != k["fic_link"]:
# fics.append([fic["fic_link"], k["fic_link"]])
fics_df = pd.DataFrame(edges, columns = ["Source", "Target", "Type"])
fics_df
#we can go through and graph it here, but I am going to export and go to a different software for faster graphing
fics_df.to_csv("nodes_and_edges/common_fics_edges_reformatted.csv", index = False)
# fic1 = fics_df["ficA"].tolist()
# fic2 = fics_df["ficB"].tolist()
# i = 0;
# g = nx.Graph()
# while i < 10000:
# g.add_edge(fic1[i], fic2[i])
# i+=1
# print(nx.info(g))
# +
# pos = nx.spring_layout(g)
# betCent = nx.betweenness_centrality(g, normalized=True, endpoints=True)
# node_color = [20000.0 * g.degree(v) for v in g]
# node_size = [v * 10000 for v in betCent.values()]
# plt.figure(figsize=(10,10))
# nx.draw_networkx(g, pos=pos, with_labels=False,
# node_color=node_color,
# node_size=node_size )
# plt.axis('off');
# -
# ### Common Fandoms
# Lets look at the common fandoms based on bookmarking a top-fic.
fandom_nodes = []
nodes_unformatted = []
tf_users = topfic_bookmarks["users"].tolist()
topfics = topfic_bookmarks["fandom"].tolist()
users = user_df["user"].tolist()
count = 1
for fic in topfics:
for users in tf_users:
for user in users:
l = user_df.loc[user_df["user"] == user["user_name"]]["fandoms"]
if len(l) > 0:
for i in l:
for j in i:
for k in j:
if 'fandom_name' in k.keys():
if k["fandom_name"] not in nodes_unformatted:
fandom_nodes.append([count, k["fandom_name"]])
nodes_unformatted.append(k["fandom_name"])
count += 1
print(len(fandom_nodes))
# nodes_df = pd.DataFrame(nodes, columns=["Id","Label"])
# nodes_df.head()
# fandom_nodes = []
# for node in fandoms_df["fandomA"].tolist():
# if node not in fandom_nodes:
# fandom_nodes.append(node)
# for node in fandoms_df["fandomB"].tolist():
# if node not in fandom_nodes:
# fandom_nodes.append(node)
# fandom_nodes_df = pd.DataFrame(fandom_nodes, columns=["fandom_nodes"])
# fandom_nodes_df.head()
fandom_nodes_df = pd.DataFrame(fandom_nodes, columns=["Id", "Label"])
fandom_nodes_df.head()
#fandom_nodes_df.to_csv("nodes_and_edges/common_fandoms_nodes_reformatted.csv", index = False)
tf_users = topfic_bookmarks["users"].tolist()
topfics = topfic_bookmarks["fandom"].tolist()
users = user_df["user"].tolist()
fandom_egdes = []
for fandom in topfics:
for users in tf_users:
for user in users:
l = user_df.loc[user_df["user"] == user["user_name"]]["fandoms"]
if len(l) > 0:
for i in l:
for j in i:
for k in j:
if 'fandom_name' in k.keys():
target = fandom_nodes_df.loc[fandom_nodes_df.Label == k["fandom_name"],'Id'].tolist()
src = fandom_nodes_df.loc[fandom_nodes_df.Label == fandom,'Id'].tolist()
if len(src) > 0 and len(target) > 0:
src = src[0]
target = target[0]
fandom_egdes.append([src, target, "Directed"])
fandoms_df = pd.DataFrame(fandom_egdes, columns = ["Source", "Target", "Type"])
fandoms_df.to_csv("nodes_and_edges/common_fandoms_edges_reformatted.csv", index = False)
# +
#trying to get smaller graphs, lets look at My Hero Academia and Naruto
'''
My Hero Academia are both shounen anime/manga so lets see if there are any similarities within the fandoms people bookmark after they bookmarked one of their top-fics.
These are nodes 1 and 36
'''
#subset data to only edges with src = 1 or 36 then subset node list
believe_it = fandoms_df[fandoms_df["Source"].isin([1, 36])]
believe_it["Type"] = "Undirected"
print(believe_it.head())
nodes = []
nodes_unformatted = []
for idx, row in believe_it.iterrows():
temp = fandom_nodes_df.loc[fandom_nodes_df['Id'] == row["Target"]]["Label"].tolist()
if len(temp)>0:
if row["Target"] not in nodes_unformatted:
nodes.append([row["Target"], temp[0]])
nodes_unformatted.append(row["Target"])
print(len(nodes))
# -
quirks = pd.DataFrame(nodes, columns=["Id", "Label"])
quirks.head()
#quirks.to_csv("nodes_and_edges/mha_nar_nodes_reformatted.csv", index = False)
believe_it.to_csv("nodes_and_edges/mha_nar_edges_reformatted.csv", index = False)
# +
'''
Let's try looking at the most popular fandoms based on our analysis from before: Marvel, RPF, KPop
'''
#subset data to only edges with src = 1 or 36 then subset node list
df1 = fandoms_df[fandoms_df["Source"].isin([32, 733, 774])]
df1["Type"] = "Undirected"
df1
nodes = []
nodes_unformatted = []
for idx, row in df1.iterrows():
temp = fandom_nodes_df.loc[fandom_nodes_df['Id'] == row["Target"]]["Label"].tolist()
if len(temp)>0:
if row["Target"] not in nodes_unformatted:
nodes.append([row["Target"], temp[0]])
nodes_unformatted.append(row["Target"])
print(len(nodes))
df2 = pd.DataFrame(nodes, columns=["Id", "Label"])
df2.head()
df2.to_csv("nodes_and_edges/top3_nodes_reformatted.csv", index = False)
df1.to_csv("nodes_and_edges/top3_edges_reformatted.csv", index = False)
# +
'''
Let's try looking at the two less popular fandoms animes and see if it sticks in the genre or not. Attack on Titan and Miraculous Ladybug
'''
#subset data to only edges with src = 1 or 36 then subset node list
anime1 = fandoms_df[fandoms_df["Source"].isin([7, 90])]
anime1["Type"] = "Undirected"
#print(anime1)
nodes = []
nodes_unformatted = []
for idx, row in anime1.iterrows():
temp = fandom_nodes_df.loc[fandom_nodes_df['Id'] == row["Target"]]["Label"].tolist()
if len(temp)>0:
if row["Target"] not in nodes_unformatted:
nodes.append([row["Target"], temp[0]])
nodes_unformatted.append(row["Target"])
print(len(nodes))
anime2 = pd.DataFrame(nodes, columns=["Id", "Label"])
anime2.head()
anime2.to_csv("nodes_and_edges/anime3_nodes_reformatted.csv", index = False)
anime1.to_csv("nodes_and_edges/anime3_edges_reformatted.csv", index = False)
# +
'''
Let's try looking at the two less popular fandoms in the musical category and see if it sticks in the genre or not.
Hamilton - Miranda, Newsies - All Media Types, and Be More Chill - Iconis/Tracz
'''
#subset data to only edges with src = 1 or 36 then subset node list
musical1 = fandoms_df[fandoms_df["Source"].isin([152,1236, 815])]
musical1["Type"] = "Undirected"
#print(anime1)
nodes = []
nodes_unformatted = []
for idx, row in musical1.iterrows():
temp = fandom_nodes_df.loc[fandom_nodes_df['Id'] == row["Target"]]["Label"].tolist()
if len(temp)>0:
if row["Target"] not in nodes_unformatted:
nodes.append([row["Target"], temp[0]])
nodes_unformatted.append(row["Target"])
print(len(nodes))
musical2 = pd.DataFrame(nodes, columns=["Id", "Label"])
musical2.head()
musical2.to_csv("nodes_and_edges/musical3_nodes_reformatted.csv", index = False)
musical1.to_csv("nodes_and_edges/musical3_edges_reformatted.csv", index = False)
# -
| data_analysis/bookmarks_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Ubuntu Linux)
# language: python
# name: python3
# ---
# # Markdown in Jupyter Notebooks
# In order to be able to enter formatted Markdown text, select "Markdown" in the dropdown for the mode of the cell (it usually says "code").
# ## Quick reference
#
# ### Visual look
#
# * `**bold**`: **bold** text
# * `_italic_`: _italic_
# * `~~strike~~`: ~~strike~~
# * Headers: `# Main`, `## Subheader`, `### Subsub header`, ...
#
# Explore more by selecting words and clicking on the formatting buttons of this editor.
#
# ### Structure
#
# Lists:
# ```
# * item1
# * item2
# ```
#
# Enumeration:
# ```
# 1. item1
# 1. item2
# ```
#
# ### Special elements
#
# * Links: `[name of link](http://url.it.links/to)`
#
# ## Formuas
#
# It is also possible to embed formulas. They are formatted using [MathJax](https://www.mathjax.org/).
#
# Inline $x^2$ formula.
#
# \begin{equation}
# E = mc^2
# \end{equation}
#
# ## References
#
# * [Original description](https://daringfireball.net/projects/markdown/)
| markdown/markdown-in-jupyter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import nglview as nv
import pytraj as pt
traj = pt.datafiles.load_rna()
view = nv.show_pytraj(traj)
# view.clear_representations()
view
# -
cbs = view._ngl_displayed_callbacks[:]
[cb(view) for cb in cbs[1:]]
view._ngl_displayed_callbacks[-1](view)
view.center_view(component=0)
view.add_licorice(component=1)
ntraj = nv.PyTrajTrajectory(pt.datafiles.load_trpcage())
view.add_trajectory(ntraj, center_view=True, default_representation=False)
view._js_console()
view.add_cartoon(component=1)
view.add_cartoon(component=1)
s = nv.Structure()
t = nv.PyTrajTrajectory(traj)
t
t.id
class Dum:
def __init__(self):
self.trajlist = []
# +
dum = Dum()
dum.trajlist.append(t)
# -
t0 = dum.trajlist[0]
t0.id
id(t)
id(t0)
| nglview/tests/notebooks/add_trajectory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notebook para criar gráficos utilizados no exercício 2
# ## Bibliotecas utilizadas
# %matplotlib inline
##Bibliotecas importadas
# Biblioteca usada para abrir arquivos CSV
import csv
# Biblioteca mateḿática
import numpy as np
# Bibloteca para traçar gráficos
import matplotlib.pyplot as plt
#Biblioteca para mudar tamanho o gráfico apresentado
import matplotlib.cm as cm
import operator as op
import os
import math
# ## Gráficos que avaliam tamanho da cache L1
# +
dados = list(csv.reader(open('l1.csv','r')))
# Tamanhos da cache (KB)
sizes = [2,4,8,16,32]
# Listas da porcentagem de misses na cache de instruções e dados
# para cada tamanho
inst = []
data = []
for dado in dados:
# Nome do programa analisado
program = dado[0]
for i in range(1,6):
inst.append(float(dado[2*i-1]))
data.append(float(dado[2*i]))
x = np.arange(1, len(sizes) + 1)
##################### Gráfico para instruções
markerline, stemlines, baseline = plt.stem(x, inst)
plt.xticks(x, sizes)
# Bordas do gráfico
maxx = len(sizes)+1
maxy = max(inst)
maxy = maxy + maxy/50
miny = min(inst)
plt.ylim([miny,maxy])
plt.xlim([0,maxx])
# Estilo das linhas
plt.setp(stemlines, 'linestyle', '-', 'color', 'k', 'linewidth', 2, 'marker', 'h', 'markersize', 7)
# Legendas
plt.title('Misses % da L1 ICache - ' + program)
plt.xlabel('Tamanhos (KB)')
plt.ylabel('Miss %')
#Salva gŕafico, o mostra no notebook e fecha o plot
plt.savefig('graphs/l1i-' + program + '.png', dpi=300)
plt.show()
plt.close()
##################### Gráfico para dados
markerline, stemlines, baseline = plt.stem(x, data)
plt.xticks(x, sizes)
# Bordas do gráfico
maxx = len(sizes)+1
maxy = max(data)
maxy = maxy + maxy/50
miny = min(data)
plt.ylim([miny,maxy])
plt.xlim([0,maxx])
# Estilo das linhas
plt.setp(stemlines, 'linestyle', '-', 'color', 'k', 'linewidth', 2, 'marker', 'h', 'markersize', 7)
# Legendas
plt.title('Misses % da L1 DCache - ' + program)
plt.xlabel('Tamanhos (KB)')
plt.ylabel('Miss %')
#Salva gŕafico, o mostra no notebook e fecha o plot
plt.savefig('graphs/l1d-' + program + '.png', dpi=300)
plt.show()
plt.close()
# Reinicia listas
inst[:] = []
data[:] = []
# -
# ## Gráficos que avaliam tamanho dos blocos tendo o melhor tamanho das caches L1
# +
dados = list(csv.reader(open('block.csv','r')))
# Tamanhos do bloco (bits)
sizes = [2,4,8,16,32]
# Listas da porcentagem de misses na cache de instruções e dados
# para cada tamanho
inst = []
data = []
for dado in dados:
# Nome do programa analisado
program = dado[0]
for i in range(1,6):
inst.append(float(dado[2*i-1]))
data.append(float(dado[2*i]))
x = np.arange(1, len(sizes) + 1)
##################### Gráfico para instruções
markerline, stemlines, baseline = plt.stem(x, inst)
plt.xticks(x, sizes)
# Bordas do gráfico
maxx = len(sizes)+1
maxy = max(inst)
maxy = maxy + maxy/50
miny = min(inst)
plt.ylim([miny,maxy])
plt.xlim([0,maxx])
# Estilo das linhas
plt.setp(stemlines, 'linestyle', '-', 'color', 'k', 'linewidth', 2, 'marker', 'h', 'markersize', 7)
# Legendas
plt.title('Misses % da L1 ICache (32K) - ' + program)
plt.xlabel('Tamanho dos blocos (bits)')
plt.ylabel('Miss %')
#Salva gŕafico, o mostra no notebook e fecha o plot
plt.savefig('graphs/bi-' + program + '.png', dpi=300)
plt.show()
plt.close()
##################### Gráfico para dados
markerline, stemlines, baseline = plt.stem(x, data)
plt.xticks(x, sizes)
# Bordas do gráfico
maxx = len(sizes)+1
maxy = max(data)
maxy = maxy + maxy/50
miny = min(data)
plt.ylim([miny,maxy])
plt.xlim([0,maxx])
# Estilo das linhas
plt.setp(stemlines, 'linestyle', '-', 'color', 'k', 'linewidth', 2, 'marker', 'h', 'markersize', 7)
# Legendas
plt.title('Misses % da L1 DCache (32K) - ' + program)
plt.xlabel('Tamanho dos blocos (bits)')
plt.ylabel('Miss %')
#Salva gŕafico, o mostra no notebook e fecha o plot
plt.savefig('graphs/bd-' + program + '.png', dpi=300)
plt.show()
plt.close()
# Reinicia listas
inst[:] = []
data[:] = []
# -
# ## Gráficos que avaliam melhor associatividade
# +
dados = list(csv.reader(open('assoc.csv','r')))
# Associatividades
assoc = []
# Listas da porcentagem de misses na cache de instruções e dados
# para cada tamanho
inst = []
data = []
for dado in dados:
# Nome do programa analisado
program = dado[0]
for i in range(1,6):
inst.append(float(dado[2*i-1]))
data.append(float(dado[2*i]))
x = np.arange(1, len(assoc) + 1)
##################### Gráfico para instruções
markerline, stemlines, baseline = plt.stem(x, inst)
plt.xticks(x, assoc)
# Bordas do gráfico
maxx = len(assoc)+1
maxy = max(inst)
maxy = maxy + maxy/50
miny = min(inst)
plt.ylim([miny,maxy])
plt.xlim([0,maxx])
# Estilo das linhas
plt.setp(stemlines, 'linestyle', '-', 'color', 'k', 'linewidth', 2, 'marker', 'h', 'markersize', 7)
# Legendas
plt.title('Misses % da L1 ICache (32K) (32b block) - ' + program)
plt.xlabel('Associatividade')
plt.ylabel('Miss %')
#Salva gŕafico, o mostra no notebook e fecha o plot
plt.savefig('graphs/ai-' + program + '.png', dpi=300)
plt.show()
plt.close()
##################### Gráfico para dados
markerline, stemlines, baseline = plt.stem(x, data)
plt.xticks(x, assoc)
# Bordas do gráfico
maxx = len(assoc)+1
maxy = max(data)
maxy = maxy + maxy/50
miny = min(data)
plt.ylim([miny,maxy])
plt.xlim([0,maxx])
# Estilo das linhas
plt.setp(stemlines, 'linestyle', '-', 'color', 'k', 'linewidth', 2, 'marker', 'h', 'markersize', 7)
# Legendas
plt.title('Misses % da L1 DCache (32K) (32b) - ' + program)
plt.xlabel('Associatividade')
plt.ylabel('Miss %')
#Salva gŕafico, o mostra no notebook e fecha o plot
plt.savefig('graphs/ad-' + program + '.png', dpi=300)
plt.show()
plt.close()
# Reinicia listas
inst[:] = []
data[:] = []
| exercicio2/.ipynb_checkpoints/exercicio2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os
import matplotlib.pyplot as plt
# %matplotlib inline
from PIL import Image
from scipy.stats import truncnorm
import cv2
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import DataLoader
from torch.utils.data.sampler import SubsetRandomSampler
import tqdm
import time
import pdb
import argparse
import sys
from shapes_loader import *
from base_model import *
import tf_homo
from demo_superpoint import SuperPointFrontend
from homography import sample_homography
# +
# magicleap model
fe = SuperPointFrontend(weights_path='superpoint_v1.pth',
nms_dist=4,
conf_thresh=0.0001,
nn_thresh=10,
cuda=True)
# model
model = SuperPointNet().cuda()
# model = torch.load('./Fri Nov 9 15:54:21 2018/e_405_a_10.9042.model').cuda()
model.load_state_dict(torch.load('20181202_train/e_49_a_90.2644.dict'))
model.eval()
criterion = nn.Softmax(dim=1) #reduction='elementwise_sum')
W = 400
H = 300
clr_imgs = Image.open('../hpatches-dataset/img/images.png').resize((W*6,H),Image.ANTIALIAS)
# clr_imgs = Image.open('../tesrt/qc_left_br_023_1481830517.76.png').resize((W,H),Image.ANTIALIAS)
# clr_imgs_2 = Image.open('../tesrt/qc_left_br_030_1481830521.45.png').resize((W,H),Image.ANTIALIAS)
img1 = np.array(clr_imgs)
img1 = img1[:H,:W]
clr_img1 = Image.fromarray(img1)
img1 = np.array(clr_img1.convert('L'))
#img2 = np.array(clr_imgs_2)
img2 = np.array(clr_imgs)
img2 = img2[:H,W:2*W]
clr_img2 = Image.fromarray(img2)
img2 = np.array(clr_img2.convert('L'))
# +
# magic leap model
pt_1, desc_1, _ = fe.run(img1.astype('float32'))
pt_2, desc_2, _ = fe.run(img2.astype('float32'))
fig=plt.figure()
plt.imshow(clr_img1)
plt.plot(pt_1[0,:],pt_1[1,:],'xy')
fig=plt.figure()
plt.imshow(clr_img2)
plt.plot(pt_2[0,:],pt_2[1,:],'xy')
# -
probs_1, desc_1 = model(torch.from_numpy(img1).unsqueeze(0).unsqueeze(1).float().cuda())
probs_2, desc_2 = model(torch.from_numpy(img2).unsqueeze(0).unsqueeze(1).float().cuda())
# show results
threshold = 0.2
fig=plt.figure()
fig.add_subplot(1, 2, 1, title='Image 1')
plt.imshow(clr_img1)
ipt_sm_1 = criterion(probs_1)
ipt_sm_1 = ipt_sm_1[:,:-1,:,:]
#find the max entry and confidence
idx_conf_1, idx_locs_1 = ipt_sm_1.max(dim=1)
idx_mask_1 = idx_conf_1 > threshold
px = []
py = []
for x in range(probs_1.shape[2]):
for y in range(probs_1.shape[3]):
if idx_mask_1[0,x,y] == 1:
#location in the image
x_ = x*8 +(idx_locs_1[0,x,y]/8)
y_ = y*8 + (idx_locs_1[0,x,y]%8)
px.append(x_.item())
py.append(y_.item())
plt.plot(py,px,'xy')
# img 2
fig.add_subplot(1, 2, 2, title='Image 1')
plt.imshow(clr_img2)
ipt_sm_1 = criterion(probs_2)
ipt_sm_1 = ipt_sm_1[:,:-1,:,:]
#find the max entry and confidence
idx_conf_1, idx_locs_1 = ipt_sm_1.max(dim=1)
idx_mask_1 = idx_conf_1 > threshold
px = []
py = []
for x in range(probs_1.shape[2]):
for y in range(probs_1.shape[3]):
if idx_mask_1[0,x,y] == 1:
#location in the image
x_ = x*8 +(idx_locs_1[0,x,y]/8)
y_ = y*8 + (idx_locs_1[0,x,y]%8)
px.append(x_.item())
py.append(y_.item())
plt.plot(py,px,'xy')
# +
# see opencv
img_1_cv = cv2.imread('../tesrt/vc_left_015_1481830513.55.png')
img_2_cv = cv2.imread('../tesrt/vc_left_019_1481830515.64.png')
#img_2_cv = img_1_cv.copy()
# img_1_cv = img1
# img_2_cv = img2
# cv2.imwrite('p1.png',img_1_cv)
# cv2.imwrite('p2.png',img_2_cv)
# Initiate ORB detector
orb = cv2.ORB_create()
# find the keypoints and descriptors with SIFT
kp1, des1 = orb.detectAndCompute(img_1_cv, None)
kp2, des2 = orb.detectAndCompute(img_2_cv, None)
pts1 = np.array([kp1[idx].pt for idx in range(len(kp1))]).reshape(-1, 2)
pts2 = np.array([kp2[idx].pt for idx in range(len(kp2))]).reshape(-1, 2)
# plt.figure()
# plt.imshow(img_1_cv)
# plt.plot(pts1[:,0],pts1[:,1],'.r',markersize=2)
# plt.savefig('foo.png',dpi=300)
plt.figure()
plt.imshow(img_2_cv)
plt.plot(pts2[:,0],pts2[:,1],'.r',markersize=2)
plt.savefig('foo.png',dpi=300)
# # create BFMatcher object
# bf = cv2.BFMatcher(cv2.NORM_HAMMING, crossCheck=True)
# # Match descriptors.
# matches = bf.match(des1,des2)
# # Sort them in the order of their distance.
# matches = sorted(matches, key = lambda x:x.distance)
# print(len(matches))
# # Draw first 10 matches.
# img3 = cv2.drawMatches(img_1_cv,kp1,img_2_cv,kp2,matches[:30],None, flags=2)
# plt.imshow(img3),plt.show()
# -
pts1.shape
| generate_matches.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Adversarial Training
#
# This notebook shows the creation of an adversarial training methodology to harden a neural network against a digital FGSM attack.
#
# ### Assumptions
# - The dataset wrangling has already been completed (and is provided here)
# - The adversarial attack (FGSM) has already been completed
# - The outer loop of training has already been completed and we're only subclassing a single epoch
# - The plotting code has already been completed
#
# ### Components Recreated in Tutorial
# - Adversarial training constrained by a power ratio and a percentage of the dataset it alters each epoch.
#
# ### See Also
# The code in this tutorial is a stripped down version of the code in ``rfml.nn.train.adversarial`` that simplifies discussion. Further detail can be provided by directly browsing the source files.
# Install the library code
# !pip install git+https://github.com/brysef/rfml.git@1.0.1
# +
# Plotting Includes
import matplotlib.pyplot as plt
# External Includes
import numpy as np
import torch
from torch.autograd import Variable
from torch.nn import CrossEntropyLoss
from torch.optim import Adam
from torch.utils.data import DataLoader, TensorDataset
# Internal Includes
from rfml.attack import fgsm
from rfml.data import Dataset, Encoder
from rfml.data import build_dataset
from rfml.nbutils import plot_acc_vs_spr, plot_acc_vs_snr
from rfml.nn.eval import compute_accuracy, compute_accuracy_on_cross_sections
from rfml.nn.model import build_model, Model
from rfml.nn.train import StandardTrainingStrategy, PrintingTrainingListener
# -
# ### Configuration
gpu = True # Set to True to use a GPU for training
fig_dir = None # Set to a file path if you'd like to save the plots generated
data_path = None # Set to a file path if you've downloaded RML2016.10A locally
# ## Adversarial Training of a Model
# ### Load dataset and a DNN model
train, val, test, le = build_dataset("RML2016.10a", path=data_path)
# as_numpy returns x,y and x is shape BxCxIQxN
input_samples = val.as_numpy(le=le)[0].shape[3]
model = build_model(model_name="CNN", input_samples=input_samples, n_classes=len(le))
# ### Creating our own adversarial trainer
#
# One of the most effective, and simple, methodologies for hardening deep learning models against adversarial attacks is simply "showing" them what they are. A process known as adversarial training.
#
# Here, we recreate the adversarial training from [Kurakin et al.] that uses the FGSM attack from [Goodfellow et al.] to augment the training examples with adversarial examples. Note that this coupling of adversarial attack and training was found, in the context of computer vision, to produce robustness which was misleading because it was actually learning to obfuscate the gradient used as a "signal" to create the adversarial example and not necessarily becoming robust to the attack [Tramer et al.]. The adversarial training methodology was then extended to Ensemble Adversarial Training by [Tramer et al.], however, this notebook only demonstrates the adversarial training proposed in [Kurakin et al.] as it can be more easily self-contained into a notebook for demonstration.
#
# Also note that adversarial training was applied in the context of RF in [Kokalj-Filipovic and Miller].
#
# #### Goodfellow et al.
# <NAME>., <NAME>., and <NAME>. (2015). Explaining and harnessing adversarial examples. In Int. Conf. on Learning Representations.
#
# #### Kurakin et al.
#
# <NAME>., <NAME>., and <NAME>. (2016). Adversarial machine learning at scale.CoRR, abs/1611.01236.
#
# #### Tramer et. al
# <NAME>., <NAME>., <NAME>., <NAME>., and <NAME>. (2017). Ensemble adversarial training: Attacks and defenses.CoRR, abs/1705.07204.
#
# #### Kokalj-Filipovic and Miller
#
# <NAME>. and <NAME>. (2019). Adversarial examples in RF deep learning: Detection of the attack and its physical robustness.CoRR, abs/1902.06044.
class MyAdversarialTrainingStrategy(StandardTrainingStrategy):
def __init__(
self,
lr: float = 10e-4,
max_epochs: int = 50,
patience: int = 5,
batch_size: int = 512,
gpu: bool = True,
k: float = 0.05,
spr: float = 10.0,
):
super().__init__(
lr=lr,
max_epochs=max_epochs,
patience=patience,
batch_size=batch_size,
gpu=gpu,
)
self.k = k
self.spr = spr
# The exact value of the sps shouldn't actually matter. It's simply used for
# an intermediate scaling of the example before applying the adversarial
# perturbation with FGSM. This assumption that it shouldn't matter is based
# upon the expectation the model does the normalization as the first "layer"
# in its network.
self.sps = 8
def _train_one_epoch(
self, model: Model, data: DataLoader, loss_fn: CrossEntropyLoss, optimizer: Adam
) -> float:
total_loss = 0.0
# Switch the model mode so it remembers gradients, induces dropout, etc.
model.train()
for i, batch in enumerate(data):
x, y = batch
# Perform adversarial augmentation in the training loop using FGSM
x = self._adversarial_augmentation(x=x, y=y, model=model)
# Push data to GPU
if self.gpu:
x = Variable(x.cuda())
y = Variable(y.cuda())
else:
x = Variable(x)
y = Variable(y)
# Forward pass of prediction -- while some are adversarial
outputs = model(x)
# Zero out the parameter gradients, because they are cumulative,
# compute loss, compute gradients (backward), update weights
loss = loss_fn(outputs, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
total_loss += loss.item()
mean_loss = total_loss / (i + 1.0)
return mean_loss
def _adversarial_augmentation(
self, x: torch.Tensor, y: torch.Tensor, model: Model
) -> torch.Tensor:
# Rely on the fact that the DataLoader shuffles -- therefore can just take the
# first *n* examples and perform adversarial augmentation on it and it will be
# a random selection.
n_adversarial = int(self.k * x.shape[0])
if n_adversarial == 0:
return x
x[0:n_adversarial, ::] = fgsm(
x=x[0:n_adversarial, ::],
y=y[0:n_adversarial],
net=model,
spr=self.spr,
sps=self.sps,
)
return x
trainer = MyAdversarialTrainingStrategy(max_epochs=10,
patience=3,
gpu=gpu,
k=0.25,
spr=10)
trainer.register_listener(PrintingTrainingListener())
trainer(model=model, training=train, validation=val, le=le)
# ### Testing the Adversarial Trained Model on normal data
# This ensures that we haven't completely sacraficed our performance in the baseline case.
acc = compute_accuracy(model=model, data=test, le=le)
print("Overall Testing Accuracy: {:.4f}".format(acc))
# +
acc_vs_snr, snr = compute_accuracy_on_cross_sections(model=model,
data=test,
le=le,
column="SNR")
title = "Accuracy vs SNR of {model_name} on {dataset_name}".format(model_name="CNN", dataset_name="RML2016.10A")
fig = plot_acc_vs_snr(acc_vs_snr=acc_vs_snr, snr=snr, title=title)
if fig_dir is not None:
file_path = "{fig_dir}/hardened_acc_vs_snr.pdf"
print("Saving Figure -> {file_path}".format(file_path=file_path))
fig.savefig(file_path, format="pdf", transparent=True)
plt.show()
# -
# ### Verifying the Model has been Hardened
# Attempt Evading Signal Classification with Direct Access to the Classifier again to see if we have improved over our prior attack.
mask = test.df["SNR"] >= 18
dl = DataLoader(test.as_torch(le=le, mask=mask), shuffle=True, batch_size=512)
# +
# Ensure that the model is in "evaluation" mode
# -- Therefore batch normalization is not computed and dropout is not performed
# -- Note: This is the cause of a lot of bugs
model.eval()
acc_vs_spr = list()
sprs = list()
for spr in np.linspace(50.0, 0.0, num=26):
right = 0
total = 0
for x, y in dl:
adv_x = fgsm(x, y, spr=spr, input_size=input_samples, sps=8, net=model)
predictions = model.predict(adv_x)
right += (predictions == y).sum().item()
total += len(y)
acc = float(right) / total
acc_vs_spr.append(acc)
sprs.append(spr)
fig = plot_acc_vs_spr(acc_vs_spr=acc_vs_spr,
spr=sprs,
title="Performance of a Digital FGSM Attack after Hardening"
)
if fig_dir is not None:
file_path = "{fig_dir}/hardened_direct_access_fgsm.pdf".format(fig_dir=fig_dir)
print("Saving Figure -> {file_path}".format(file_path=file_path))
fig.savefig(file_path, format="pdf", transparent=True)
plt.show()
| notebooks/module6_empty.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''respy'': conda)'
# name: python3710jvsc74a57bd0b549dadab0c2edb1c58f223f7584f57e60f2cc8e65ef8392efb9b23cb30dad20
# ---
# # <NAME>oe or 10 minutes to respy
#
# This is a short introduction to respy for new users. As economists love Robinsonades [<sup>1</sup>](#fn1), we will showcase the implementation of a Robinson Crusoe economy as a discrete choice dynamic programming model. Throughout the notebook you find italic text which make you familiar with Robinson's story. We will first set the scene with a broad introduction and then turn to the precise model specification. We continue by simulating the model and analyze its comparative statics. We then extend the model and showcase the estimation of the model parameters.
#
# Just to be clear, don't misinterpret the fact that we explain **respy** using such a simplistic model. **respy** is not a toy and can just as well solve state-of-the-art structural models. It's just easier to explain **respy** in a situation where we don't have to explain a complicated model at the same time.
# +
# %matplotlib inline
import io
import matplotlib.pyplot as plt
import pandas as pd
import respy as rp
import yaml
import seaborn as sns
import numpy as np
from pathlib import Path
from time import time
plt.style.use("../_static/respy.mplstyle")
# -
# ## Introduction
#
# ---
# *After setting sail against his parents' wishes, being captured by pirates, escaping from them, building a plantation, and setting sail again to capture slaves in Africa, [<NAME>](https://en.wikipedia.org/wiki/Robinson_Crusoe) stranded on a small island. He is alone with one dog, two cats, and only some supplies. He goes fishing to make ends meet and if he is too tired he will relax in his hammock. But, he cannot relax to often as storing food is a difficult task on a tropical island.*
#
# ---
#
# In the discrete choice dynamic programming model, Robinson chooses every period $t = 0, \dots, T - 1$ to either go fishing, $a = 0$, or spend the day in the hammock, $a = 1$, to maximize his expected sum of discounted lifetime utility. The utility of a choice, $U(s_t, a_t)$, depends on the state $s_t$, which contains information on the individual's characteristics, and the chosen alternative $a_t$. For working alternatives like fishing utility consists of two components, a wage and a non-pecuniary component.
#
# $$
# U(s_t, a_t) = W(s_t, a_t) + N(s_t, a_t)
# $$
#
# For non-working alternatives like the hammock, $W(s_t, a_t) = 0$. The wage is defined as
#
# $$\begin{align}
# W(s_t, a_t) &= r_a \exp\{x^w_{at} \beta^w_a + \epsilon_{at}\}\\
# \ln(W(s_t, a_t)) &= \ln(r_a) + x^w_{at} \beta^w_a + \epsilon_{at}
# \end{align}$$
#
#
# where $r_a$ is normally a market rental price for the skill units generated in the exponential expression. Another interpretation is that $ln(r_a)$ is simply the constant in the skill units. The skill units are generated by two components. $x^w_{at}$ and $\beta^w_a$ are the choice- and time-dependent covariates and parameters related to the wage signaled by superscript $w$. $\epsilon_{at}$ is a choice-specific random shock from the shock vector $\epsilon_t \sim \mathcal{N}(0, \Sigma)$ for all choices. Shocks are usually correlated across choices in one period, but are independent across periods.
#
# The non-pecuniary rewards for working alternatives are a vector dot product of covariates $x_t^w$ and parameters $\beta^w$. The superscript $w$ signals that the components belong to working alternatives.
#
# $$
# N^w(s_t, a_t) = x_t^w\beta^w
# $$
#
# The non-pecuniary reward for non-working alternatives is very similar except that the shocks enter the equation additively. Superscript $n$ stands for non-pecuniary.
#
# $$
# N^n(s_t, a_t) = x_t^n\beta^n + \epsilon_{at}
# $$
#
# Along with the lower triangular elements of the shock variance-covariance matrix of $\epsilon_t$, the utility parameters $\beta_a^w$, $\beta_a^n$ and $r_a$ form the main parameters of the model.
#
# If Robinson chooses to go fishing, he gains one additional unit of experience in the next period. Experience starts at zero and goes over 1, 2, 3 up to $T - 1$.
#
# The general assumption imposed on Robinson is that he is forward-looking and maximizes the expected present value of utility over the remaining lifetime. W.l.o.g. $t = 0$ and let $V(s_0)$ define the value of the maximization which is achieved by a sequence of choices, $\{a_t\}^T_{t = 0}$, such that every action is in the choice set, $a_t \in C(s_t)$ and $s_{t + 1}$ adheres to the law of motion. Then, the expected present value of lifetime utility in state $s_0$ is
#
# $$
# V(s_0) = \text{E} \max_{\{a_t\}^T_{t = 0}} \left[
# \sum^T_{t = 0} \delta^t U(s_t, a_t) \, \Big|
# \, a_t \in C(s_t), s_{t+1} = m(s_t, a_t)
# \right]
# $$
#
# Note that the shocks in period $t = 0$ are not stochastic. Thus, one can extract the utility in the current period $U(s_0, a_0)$, also called the flow utility, and the discount factor $\delta$ from the expectation. Then, the formula can be rewritten so that the second term becomes the maximum over alternative-specific value functions at time $t = 1$ which are also called continuation values.
#
# $$\begin{align}
# V(s_0) &= \max_{a_0} \, U(s_0, a_0) + \delta \text{E} \max_{\{a_t\}^T_{t = 1}}
# \left[
# \sum^T_{t = 1} \delta^{t - 1} U(s_t, a_t) \, \Big|
# \, a_t \in C(s_t), s_{t + 1} = m(s_t, a_t)
# \right] \\
# &= \max_{a_0} \, U(s_0, a_0)
# + \delta \text{E} \max_{a_1} V_{a_1}(s_1)
# \end{align}$$
#
# The maximum over alternative-specific value functions can be rewritten as the value function of state $s_1$ or $V(s_1) = \max_{a_1} V_{a_1}(s_1)$ which yields the famous Bellman equation. Due to the recursive nature of the problem, the alternative-specific value functions are defined as
#
# $$\begin{equation}
# V_a(s_t) = \begin{cases}
# U(s_t, a_t) + \delta \text{E} V(s_{t+1}) & \text{if } t < T \\
# U(s_t, a_t) & \text{if } t = T
# \end{cases}
# \end{equation}$$
#
# The former equation shows that the shocks in period $t + 1$ are unknown to the individual in period $t$. Thus, utility must be maximized given the joint distribution of shocks in period $t + 1$ which is a maximization problem over a two-dimensional integral. Denote the non-stochastic part of a state as $s^-$. Then, Robinson maximizes
#
# $$\begin{equation}
# V(s_t) = \max_{a_t}\{
# U(s_t, a_t) + \delta \int_{\epsilon_{0, t + 1}} \dots \int_{\epsilon_{2, t + 1}}
# \max_{a_{t + 1}} V_{a_{t + 1}}(s^-_{t + 1}, \epsilon_{t + 1})
# f_\epsilon(\epsilon_{t + 1})
# d_{\epsilon_{0, t + 1}} \dots d_{\epsilon_{2, t + 1}}
# \}
# \end{equation}$$
# ## Specification
#
# How can we express the equations and parameters with **respy**? The following cell contains the code to write a `.csv` file which is the cornerstone of a model as it contains all parameters and some other specifications. With `io.StringIO` we can pretend it is an actual file on the filesystem and easily loaded with `pandas`.
params = """category,name,value
delta,delta,0.95
wage_fishing,exp_fishing,0.1
nonpec_fishing,constant,-1
nonpec_hammock,constant,2.5
nonpec_hammock,not_fishing_last_period,-1
shocks_sdcorr,sd_fishing,1
shocks_sdcorr,sd_hammock,1
shocks_sdcorr,corr_hammock_fishing,-0.2
lagged_choice_1_hammock,constant,1
"""
params = pd.read_csv(io.StringIO(params), index_col=["category", "name"])
params
# The DataFrame for the parameters contains a two-level MultiIndex to group parameters in categories. `name` should be uniquely assigned in each category or otherwise only the sum of identically named parameters is identified. `value` contains the value of the parameter. Note that we named Robinson's alternatives `"fishing"` and `"hammock"` and we have to use the names consistently. As long as you stick to lowercase letters separated by underscores, you can choose any name you want.
#
# The parameter specification contains following entries:
#
# - The first entry contains the discount factor of individuals.
# - The second category `"wage_fishing"` contains the parameters of the log wage equation for fishing. The group contains only one name called `"exp_fishing"` where `"exp_*"` is an identifier in the model for experience accumulated in a certain alternative. **respy** requires that you respect those identifiers of which there are not many and reference your alternatives consistently with the same name. If you stick to lowercase letters possibly separated by underscores, you are fine.
# - The third and fourth categories concern the non-pecuniary reward of fishing and relaxing in the hammock.
# - `"shocks_sdcorr"` groups the lower triangular of the variance-covariance matrix of shocks.
# - `"lagged_choice_1_hammock"` governs the distribution of previous choices at the begin of the model horizon.
#
# `params` is complemented with `options` which contains additional information. Here is short description:
#
# - `"n_periods"` defines the number of periods for which decision rules are computed.
# - `"_seed"`: Seeds are used in every model component to ensure reproducibility. You can use any seed you would like or even repeat the same seed number. Internally, we ensure that randomness is completely uncorrelated.
# - `"estimation_draws"` defines the number of draws used to simulate the choice probabilities with Monte Carlo simulation in the maximum likelihood estimation.
# - `"estimation_tau"` controls the temperature of the softmax function to avoid zero-valued choice probabilities.
# - `"interpolation_points"` controls how many states are used to approximate the value functions of others states in each period. `-1` turns the approximation off. The approximation is detailed in Keane and Wolpin (1994).
# - ``"simulation_agents"`` defines how many individuals are simulated.
# - ``"solution_draws"`` defines the number of draws used to simulate the expected value functions in the solution.
# - `"covariates"` is another dictionary where the key determines the covariate's name and the value is its definition. Here, we have to define what `"constant"` means.
options = """n_periods: 10
estimation_draws: 200
estimation_seed: 500
estimation_tau: 0.001
interpolation_points: -1
simulation_agents: 1_000
simulation_seed: 132
solution_draws: 500
solution_seed: 456
covariates:
constant: "1"
not_fishing_last_period: "lagged_choice_1 != 'fishing'"
"""
options = yaml.safe_load(options)
options
# ## Simulation
#
# We are now ready to simulate the model.
simulate = rp.get_simulate_func(params, options)
df = simulate(params)
df.head(15)
# We can inspect Robinson's decisions period by period.
# +
fig, ax = plt.subplots()
df.groupby("Period").Choice.value_counts(normalize=True).unstack().plot.bar(
stacked=True, ax=ax
)
plt.xticks(rotation="horizontal")
plt.legend(loc="lower center", bbox_to_anchor=(0.5,-0.275), ncol=2)
plt.show()
plt.close()
# -
# We can also analyze the persistence in decisions.
data = pd.crosstab(df.Lagged_Choice_1, df.Choice, normalize=True)
sns.heatmap(data, cmap="Blues", annot=True)
# ## Analysis
#
# We now study how Robinson's behavior changes as we increase the returns to experience. We do so by plotting the average level of final experience in the sample under the different parameterizations.
#
# This analysis of the comparative statics of the model is straightforward to implement. In models of educational choice, this type of analysis is often applied to evaluate the effect of alternative tuition policies on average educational attainment. See Keane & Wolpin (1997, 2001) for example. The basic structure of the analysis remains the same.
# +
# Specification of grid for evaluation
num_points = 15
grid_start = 0.0
grid_stop = 0.3
grid_points = np.linspace(grid_start, grid_stop, num_points)
rslts = list()
for value in grid_points:
params.loc["wage_fishing", "exp_fishing"] = value
df = simulate(params)
stat = df.groupby("Identifier")["Experience_Fishing"].max().mean()
rslts.append(stat)
# -
# We collected all results in `rslts` and are ready to create a basic visualization.
# +
fig, ax = plt.subplots()
ax.plot(grid_points, rslts)
ax.set_ylim([0, 10])
ax.set_xlabel("Return to experience")
ax.set_ylabel("Average final level of exerience")
plt.show()
plt.close()
# -
# In the absence of any returns to experience, Robinson still spends more than two periods fishing. This share then increases with the return. Starting at around 0.2, Robinson spends all his time fishing.
#
# ## Extension
#
# Let us make the model more interesting!
#
# ---
# *At some point Crusoe notices that a group of cannibals occasionally visits the island and celebrate one of their dark rituals. But then, a prisoner can escape and becomes Crusoe's new friend Friday whom he teaches English. In return Friday can share his knowledge once to help Robinson improve his fishing skills, but that is only possible after Robinson tried at least once to go fishing.*
#
# ---
#
# A common extension to structural models is to increase the choice set. Here, we want to add another choice called `"friday"` which affects the utility of fishing. The choice should be available once, starting with the third period, and only after Robinson has been fishing before.
#
# Note that, we load the example models with the function, `rp.get_example_model`. The command for the former model is `params, options, df = rp.get_example_model("robinson_crusoe_basic")`. You can use `with_data=False` to suppress the automatic simulation of a sample with this parameterization.
params, options = rp.get_example_model("robinson_crusoe_extended", with_data=False)
# At first, take a look at the parameterization. There is a new positive parameter called `"contemplation_with_friday"` which enters the wage equation of fishing. The choice `"friday"` itself has a negative constant utility term which models the effort costs of learning and the food penalty. The variance-covariance matrix is also adjusted.
params
# Turning to the `options`, we can see that the new covariate `"contemplation_with_friday"` is only affecting utility if Robinson is experienced in fishing and only for one interaction with friday. This naturally limits the interaction with Friday. The key `"negative_choice_set"` can be used to restrict the choice Friday to the third and following periods. The first key matches a choice. The value of the key can be a list of strings. If the string evaluates to `True`, a utility penalty ensures that individuals will never choose the corresponding states. There exist some states in the state space which will never be reached because choices are mutually exclusive or are affected by other restrictions. Filters under `"core_state_space_filters"` can be used to purge those states from the state space, reducing runtime and memory consumption.
options
# Now, let us simulate a sample of the new model.
simulate = rp.get_simulate_func(params, options)
df = simulate(params)
# +
fig, ax = plt.subplots()
df.groupby("Period").Choice.value_counts(normalize=True).unstack().plot.bar(
stacked=True, ax=ax, color=["C0", "C2", "C1"],
)
plt.xticks(rotation="horizontal")
plt.legend(loc="lower center", bbox_to_anchor=(0.5,-0.275), ncol=3)
plt.show()
plt.close()
# -
# ## Estimation
#
# For model calibration, **respy** supports estimation via maximum likelihood and the method of simulated moments. An appropriate criterion function function for both methods can be constructed in just a few steps.
#
# For optimization of the criterion, the use of external optimization libraries is requried. We recommend [estimagic](https://github.com/OpenSourceEconomics/estimagic), an open-source tool to estimate structural models and more. **estimagic** can be used for the optimization and standard error calculation of a criterion function produced by **respy**.
#
# Unlike other optimization libraries, ``estimagic`` does not optimize over a simple vector of parameters, but instead stores parameters in a ``pd.DataFrame``, which makes it easier to parse them into the quantities we need, store lower and upper bounds together with parameters and express constraints on the parameters.
#
# For ``estimagic``, we need to pass constraints on the parameters in a list containing dictionaries. Each dictionary is a constraint. A constraint includes two components: First, we need to tell ``estimagic`` which parameters we want to constrain. This is achieved by specifying an index location which will be passed to `df.loc`. Then, define the type of the constraint. Here, we only impose the constraint that the shock parameters have to be valid variances and correlations.
#
# *Note*: It is possible to utilize other optimization libraries but we recommend **estimagic** due to the reasons stated above.
from estimagic import maximize
# +
crit_func = rp.get_log_like_func(params, options, df)
crit_func(params)
constr = rp.get_parameter_constraints("robinson_crusoe")
# -
results = maximize(
criterion=crit_func,
params=params,
algorithm="scipy_lbfgsb",
algo_options={"stopping_max_criterion_evaluations": 3},
constraints=constr,
)
# Running the minimal working example shown above will start an optimization that is limited to three criterion evaluations (for the sake of this tutorial). **estimagic** will also produce a logging database called `logging.db` that stores information about the optimization. The package offers many useful options to set up a tractable estimation for your model.
#
# More information can be found in the **estimagic** documentation: https://estimagic.readthedocs.io/.
# ## Footnotes
#
# <span id="fn1"><sup>1</sup>
# One of the earliest references of Robinsonades in economics can be found in Marx (1867). In the 37th footnote, he mentions that even Ricardo used the theme before him.
# </span>
#
# ## References
#
# > <NAME>. (1957). Dynamic Programming. *Princeton University Press*, Princeton, NJ.
#
# > <NAME>. and <NAME>. (1997). [The Career Decisions of Young Men](https://doi.org/10.1086/262080). *Journal of Political Economy*, 105(3): 473-522.
#
# > <NAME>. (1867). Das Kapital, Bd. 1. *MEW*, Bd, 23, 405
| docs/tutorials/robinson_crusoe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# load the raw dataset
import numpy as np
import pandas as pd
from pandas import DataFrame
rawdf = pd.read_csv("unc.edu_PANCAN_IlluminaHiSeq_RNASeqV2.geneExp.tsv", sep="\t", index_col=0)
rawdf.head()
# -
# transpose raw dataset so row is patient samples and column is list of genes
processeddf = rawdf.transpose()
# load class label
dflabels = pd.read_csv('project_class_labels_original_10471.csv',index_col='Unnamed: 0')
# check to see if there is any feature (i.e. column) has all zero values so we will delete them
removedAllZeroColdf = processeddf.loc[:, (processeddf != 0).any(axis=0)]
removedAllZeroColdf.shape
# +
# data scaling
# method 1 : standardization
from sklearn.preprocessing import StandardScaler
stdscaler = StandardScaler()
stdscalerfit = stdscaler.fit_transform(removedAllZeroColdf)
stddf = DataFrame(stdscalerfit)
stddf.head()
# +
# Feature Selection - Tree Classifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.feature_selection import SelectFromModel
clf = ExtraTreesClassifier()
clf = clf.fit(stddf, dflabels.values.ravel())
clf.feature_importances_
model = SelectFromModel(clf, threshold = 0.000035, prefit=True)
reduced = model.transform(removedAllZeroColdf)
reduceddf = DataFrame(reduced)
reduceddf.shape
# +
# split 80% training set; 20% testing set
from sklearn.model_selection import train_test_split
trainData, testData, trainLabel, testLabel = train_test_split(reduceddf, dflabels, test_size=0.20)
# +
# distance wrighting function; k = 5
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_score
import pandas as pd
import time
clf_weighted = KNeighborsClassifier(n_neighbors=5,weights='distance')
start = time.time()
clf_weighted.fit(trainData, trainLabel.values.ravel())
end = time.time()
print("5 nearest neighbor (normal) training time: ", end - start)
pred = clf_weighted.predict(testData)
accuracy = accuracy_score(testLabel, pred)
print("Accuracy Score (distance weighted) k = 5:", accuracy)
# confusion matrix
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(testLabel, pred)
print(confusion_matrix)
# classficiation report
from sklearn.metrics import classification_report
print(classification_report(testLabel, pred))
# -
| project_exp_7_kNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create a NetCDF-CF file from a comma separated text file
import pandas as pd
import xarray as xr
# File that we will modify.
myfile = "./surf-irrad-test-201904.csv"
# Read ASCII file containing a timeseries.
mydata = pd.read_csv(myfile, header=0, infer_datetime_format=True, parse_dates=['datetime'],dayfirst=True)
mydata
mydata.plot(x='datetime', y=['QLI_01','QSI_01'])
mydata['QSI_01']
# Create a Xarray dataset from the Pandas data frame returned when reading the CSV file.
xrds = xr.Dataset.from_dataframe(mydata[['datetime','QLI_01','QSI_01']])
# Check content of dataset.
xrds
# Modify metadata for variable to make CF compliant. Meaning:
# - using standard_names
# - identifying units
# - identifying missing values
# - identifying cell_methods
# Rename datetime to time to be CF compliant.
xrds = xrds.rename_dims({'index':'time'})
xrds = xrds.rename({'datetime':'time'})
xrds
# Specify time as coordinate.
xrds = xrds.set_coords('time')
# Get rid of the numeric index as dimension.
xrds = xrds.reset_index('index', drop=True)
# Add global attributes according to ACDD.
xrds.attrs={
'Conventions':'CF-1.8',
'title':'This is a test file',
'abstract':'This is a test abstract, to be extended.',
'creator_name':'<NAME>',
'creator_email':'o.<EMAIL>',
}
xrds
# Add CF specific elements on variables.
xrds['QLI_01'].attrs = {
'standard_name':'surface_net_downward_longwave_flux',
'long_name':'longwave downward irradiance at the surface',
'units': 'Wm-2',
#'_FillValue': -999.0,
}
xrds['QSI_01'].attrs = {
'standard_name':'surface_net_downward_shortwave_flux',
'long_name':'surface downward irradiance at the surface',
'units': 'Wm-2',
#'_FillValue': -999.0,
}
xrds
# Ensure that data are written in the correct form to NetCDF. Data served through THREDDS Data Servers cannot have int64 datetime specifications, but int32. Also, encoding of missingvalues are done in this step along compression.
myencoding = {
'time': {
'dtype': 'int32',
},
'QLI_01': {
'_FillValue': -999.0,
'zlib': False,
},
'QSI_01': {
'_FillValue': -999.0,
'zlib': False,
}
}
xrds.to_netcdf('mytest.nc')
# + active=""
# xrds.to_netcdf('mytest.nc',encoding=myencoding)
| notebooks/xarray-create-netcdf-from-csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Reinforcement learning
# In this notebook, we give some example of reinforcement algorithm for a pendulum model. Beware that the implementation of the algorithms are ment to be simple, hence are not super efficient: don't use them as reference if you want to implement some dedicated RL for another problem
import numpy as np
from numpy.linalg import norm,inv,pinv,svd,eig
import matplotlib.pyplot as plt
import time
import random
# ## Environments
# We are going to work with an inverted pendulum with limited torque, that must swing to collect energy before raising up to the unstable equilibrium state. As the algorithms that we are going to explore are either working on discrate action-state spaces, or continuous ones, several versions of this environment are proposed. In general, they all work the same: get in random initial configuration with reset, display in meshcat with render, and run a simulation step with step(control). Examples:
from tp6.env_pendulum import EnvPendulum,EnvPendulumDiscrete,EnvPendulumHybrid,EnvPendulumSinCos
env = EnvPendulum(1,viewer='meshcat')
env.name = str(env.__class__)
env.u0 = np.zeros(env.nu)
env.jupyter_cell()
env.render()
for i in range(10):
env.step(env.u0)
env.render()
#
# We define here 4 main environments that you can similarly test:
#
# - EnvPendulum: state NX=2 continuous, control NU=1 continuous, Euler integration step with DT=1e-2 and high friction
# - EnvPendulumDiscrete: state NX=441 discrete, control NU=11 discrete, Euler step DT=0.5 low friction
# - EnvPendulumSinCos: state NX=3 with x=[cos,sin,vel], control NU=1 control, Euler step DT=1e-2, high friction
# - EnvPendulumHybrid: state NX=3 continuous with x=[cos,sin,vel], control NU=11 discrete, Euler step DT=0.5 low friction
#
# ## Value iteration
# For the first algorithm, we implement a Value iteration, which is an algorithm working on discrete states and discrete actions. As it is not very efficient, we must coarsly discretize the pendulum. Here is the implementation.
# +
# # %load tp6/qtable.py
'''
Example of Q-table learning with a simple discretized 1-pendulum environment.
-- concerge in 1k episods with pendulum(1)
-- Converge in 10k episods with cozmo model
'''
import matplotlib.pyplot as plt
import signal
import time
import numpy as np
### --- Random seed
RANDOM_SEED = 1188 #int((time.time()%10)*1000)
print("Seed = %d" % RANDOM_SEED)
np.random.seed(RANDOM_SEED)
### --- Environment
from tp6.env_pendulum import EnvPendulumDiscrete; Env = lambda : EnvPendulumDiscrete(1,viewer='meshcat')
env = Env()
### --- Hyper paramaters
NEPISODES = 400 # Number of training episodes
NSTEPS = 50 # Max episode length
LEARNING_RATE = 0.85 #
DECAY_RATE = 0.99 # Discount factor
Q = np.zeros([env.nx,env.nu]) # Q-table initialized to 0
def policy(s):
return np.argmax(Q[s,:])
def rendertrial(s0=None,maxiter=100):
'''Roll-out from random state using greedy policy.'''
s = env.reset(s0)
for i in range(maxiter):
a = np.argmax(Q[s,:])
s,r = env.step(a)
env.render()
signal.signal(signal.SIGTSTP, lambda x,y:rendertrial()) # Roll-out when CTRL-Z is pressed
h_rwd = [] # Learning history (for plot).
for episode in range(1,NEPISODES):
x = env.reset()
rsum = 0.0
for steps in range(NSTEPS):
u = np.argmax(Q[x,:] + np.random.randn(1,env.nu)/episode) # Greedy action with noise
x2,reward = env.step(u)
# Compute reference Q-value at state x respecting HJB
Qref = reward + DECAY_RATE*np.max(Q[x2,:])
# Update Q-Table to better fit HJB
Q[x,u] += LEARNING_RATE*(Qref-Q[x,u])
x = x2
rsum += reward
h_rwd.append(rsum)
if not episode%20:
print('Episode #%d done with average cost %.2f' % (episode,sum(h_rwd[-20:])/20))
print("Total rate of success: %.3f" % (sum(h_rwd)/NEPISODES))
rendertrial()
plt.plot( np.cumsum(h_rwd)/range(1,NEPISODES) )
plt.show()
# -
# After convergence, you can try the obtained policy using the method <rendertrial>.
env.jupyter_cell()
rendertrial(maxiter=NSTEPS)
# Let's display the optimal flow. As states are denoted by their indexes, we need to recover the 2d state from the index, with the following method:
def x2d(s):
return env.decode_x(s)
from tp6.flow import plotFlow
plotFlow(env,policy,x2d)
# ## Value iteration with a neural network
# Next, we marginally modifies the value iteration to store the Q function not as a table, but as a neural network. The main modification will be that the Belman contraction must now be achieve with a gradient descent ... and that is much less efficient. Let's see, on the same environment first.
#
# +
# # %load tp6/deeptable.py
'''
Example of Q-table learning with a simple discretized 1-pendulum environment using a linear Q network.
'''
import numpy as np
import random
import tensorflow as tf
import tensorflow.compat.v1 as tf1
import matplotlib.pyplot as plt
from tp6.env_pendulum import EnvPendulumDiscrete; Env = lambda : EnvPendulumDiscrete(1,viewer='meshcat')
import signal
import time
tf1.disable_eager_execution()
### --- Random seed
RANDOM_SEED = int((time.time()%10)*1000)
print("Seed = %d" % RANDOM_SEED)
np.random.seed(RANDOM_SEED)
### --- Hyper paramaters
NEPISODES = 2000 # Number of training episodes
NSTEPS = 50 # Max episode length
LEARNING_RATE = 0.1 # Step length in optimizer
DECAY_RATE = 0.99 # Discount factor
### --- Environment
env = Env()
NX = env.nx
NU = env.nu
### --- Q-value networks
class QValueNetwork:
def __init__(self):
x = tf1.placeholder(shape=[1,NX],dtype=tf.float32)
W = tf1.Variable(tf1.random_uniform([NX,NU],0,0.01,seed=100))
qvalue = tf1.matmul(x,W)
u = tf1.argmax(qvalue,1)
qref = tf1.placeholder(shape=[1,NU],dtype=tf.float32)
loss = tf1.reduce_sum(tf.square(qref - qvalue))
optim = tf1.train.GradientDescentOptimizer(LEARNING_RATE).minimize(loss)
self.x = x # Network input
self.qvalue = qvalue # Q-value as a function of x
self.u = u # Policy as a function of x
self.qref = qref # Reference Q-value at next step (to be set to l+Q o f)
self.optim = optim # Optimizer
### --- Tensor flow initialization
#tf.reset_default_graph()
qvalue = QValueNetwork()
sess = tf1.InteractiveSession()
tf1.global_variables_initializer().run()
def onehot(ix,n=NX):
'''Return a vector which is 0 everywhere except index <i> set to 1.'''
return np.array([[ (i==ix) for i in range(n) ],],np.float)
def disturb(u,i):
u += int(np.random.randn()*10/(i/50+10))
return np.clip(u,0,NU-1)
def rendertrial(maxiter=100):
x = env.reset()
for i in range(maxiter):
u = sess.run(qvalue.u,feed_dict={ qvalue.x:onehot(x) })
x,r = env.step(u)
env.render()
if r==1: print('Reward!'); break
signal.signal(signal.SIGTSTP, lambda x,y:rendertrial()) # Roll-out when CTRL-Z is pressed
### --- History of search
h_rwd = [] # Learning history (for plot).
### --- Training
for episode in range(1,NEPISODES):
x = env.reset()
rsum = 0.0
for step in range(NSTEPS-1):
u = sess.run(qvalue.u,feed_dict={ qvalue.x: onehot(x) })[0] # Greedy policy ...
u = disturb(u,episode) # ... with noise
x2,reward = env.step(u)
# Compute reference Q-value at state x respecting HJB
Q2 = sess.run(qvalue.qvalue,feed_dict={ qvalue.x: onehot(x2) })
Qref = sess.run(qvalue.qvalue,feed_dict={ qvalue.x: onehot(x ) })
Qref[0,u] = reward + DECAY_RATE*np.max(Q2)
# Update Q-table to better fit HJB
sess.run(qvalue.optim,feed_dict={ qvalue.x : onehot(x),
qvalue.qref : Qref })
rsum += reward
x = x2
if reward == 1: break
h_rwd.append(rsum)
if not episode%20: print('Episode #%d done with %d sucess' % (episode,sum(h_rwd[-20:])))
print("Total rate of success: %.3f" % (sum(h_rwd)/NEPISODES))
rendertrial()
plt.plot( np.cumsum(h_rwd)/range(1,NEPISODES) )
plt.show()
# -
# See? Each step is much more costly (partly due to the poor implementation, but a gradient step is certainly more costly than a table update), and much less informative: the algorithm is slower to converge.
# ## Q-learning
# The good point is that the algorithm scales well for more complex problem, and efficient Q-learning can be implemented on very large one (if you have enough CPUs). Let's take a look at the main one, the Deep-Q algorithm. The input of the neural network can be basically anything (discrete, continuous, an image, etc), the only limitation is that the control must remain discrete, and not too large. So let's take an hybrid version of the same pendulum, with continuous state space (cos,sin,velocity) and discrete control.
# +
# # %load tp6/qlearn.py
'''
Train a Q-value following a classical Q-learning algorithm (enforcing the
satisfaction of HJB method), using a noisy greedy exploration strategy.
The result of a training for a continuous pendulum (after 200 iterations)
are stored in qvalue.h5.
Reference:
Mnih, Volodymyr, et al. "Human-level control through deep reinforcement learning."
Nature 518.7540 (2015): 529.
'''
from tp6.env_pendulum import EnvPendulumHybrid; Env = lambda : EnvPendulumHybrid(1,viewer='meshcat')
from tp6.qnetwork import QNetwork
from collections import deque
import time
import signal
import matplotlib.pyplot as plt
import random
import numpy as np
import tensorflow as tf
### --- Random seed
RANDOM_SEED = int((time.time()%10)*1000)
print("Seed = %d" % RANDOM_SEED)
np .random.seed (RANDOM_SEED)
random.seed (RANDOM_SEED)
### --- Environment
env = Env()
### --- Hyper paramaters
NEPISODES = 1000 # Max training steps
NSTEPS = 60 # Max episode length
QVALUE_LEARNING_RATE = 0.001 # Base learning rate for the Q-value Network
DECAY_RATE = 0.99 # Discount factor
UPDATE_RATE = 0.01 # Homotopy rate to update the networks
REPLAY_SIZE = 10000 # Size of replay buffer
BATCH_SIZE = 64 # Number of points to be fed in stochastic gradient
NH1 = NH2 = 32 # Hidden layer size
### --- Replay memory
class ReplayItem:
def __init__(self,x,u,r,d,x2):
self.x = x
self.u = u
self.reward = r
self.done = d
self.x2 = x2
replayDeque = deque()
### --- Tensor flow initialization
qvalue = QNetwork(nx=env.nx,nu=env.nu,learning_rate=QVALUE_LEARNING_RATE)
qvalueTarget = QNetwork(name='target',nx=env.nx,nu=env.nu)
# Uncomment to load networks
#qvalue.load()
#qvalueTarget.load()
def rendertrial(maxiter=NSTEPS,verbose=True):
x = env.reset()
traj = [x.copy()]
rsum = 0.
for i in range(maxiter):
u = qvalue.policy(x)[0]
x, reward = env.step(u)
env.render()
time.sleep(1e-2)
rsum += reward
traj.append(x.copy())
if verbose: print('Lasted ',i,' timestep -- total reward:',rsum)
return np.array(traj)
signal.signal(signal.SIGTSTP, lambda x,y:rendertrial()) # Roll-out when CTRL-Z is pressed
### History of search
h_rwd = []
### --- Training
for episode in range(1,NEPISODES):
x = env.reset()
rsum = 0.0
for step in range(NSTEPS):
u = qvalue.policy(x, # Greedy policy ...
noise=1. / (1. + episode + step)) # ... with noise
x2,r = env.step(u)
done = False # Some environment may return information when task completed
replayDeque.append(ReplayItem(x,u,r,done,x2)) # Feed replay memory ...
if len(replayDeque)>REPLAY_SIZE: replayDeque.popleft() # ... with FIFO forgetting.
rsum += r
x = x2
if done: break
# Start optimizing networks when memory size > batch size.
if len(replayDeque) > BATCH_SIZE:
batch = random.sample(replayDeque,BATCH_SIZE) # Random batch from replay memory.
x_batch = np.vstack([ b.x for b in batch ])
u_batch = np.vstack([ b.u for b in batch ])
r_batch = np.array([ [b.reward] for b in batch ])
d_batch = np.array([ [b.done] for b in batch ])
x2_batch = np.vstack([ b.x2 for b in batch ])
# Compute Q(x,u) from target network
v_batch = qvalueTarget.value(x2_batch)
qref_batch = r_batch + (d_batch==False)*(DECAY_RATE*v_batch)
# Update qvalue to solve HJB constraint: q = r + q'
qvalue.trainer.train_on_batch([x_batch,u_batch],qref_batch)
# Update target networks by homotopy.
qvalueTarget.targetAssign(qvalue,UPDATE_RATE)
# \\\END_FOR step in range(NSTEPS)
# Display and logging (not mandatory).
print('Ep#{:3d}: lasted {:d} steps, reward={:3.0f}' .format(episode, step,rsum))
h_rwd.append(rsum)
if not (episode+1) % 200: rendertrial(30)
# \\\END_FOR episode in range(NEPISODES)
print("Average reward during trials: %.3f" % (sum(h_rwd)/NEPISODES))
rendertrial()
plt.plot( np.cumsum(h_rwd)/range(1,NEPISODES) )
plt.show()
# Uncomment to save networks
#qvalue.save()
# -
# ## Actor critic
# When the control space is discrete, the optimal policy is directly obtained by maximizing the Q value by an exhaustive search. This does not work for continuous space. In that case, a policy network must be trained in parallel, to greedily optimize the Q function. A famous algorithm for that, which is a near direct extension of Deep-Q, is the Deep Deterministic Policy Gradient (DDPG).
#
# Like Deep-Q, it optimizes the Q function to contract the Belman residual. To efficiently do that, it also uses minibatches to avoid the local collapses due to sample dependancy. In addition, it uses a smoothing of the gradient direction, using a so-called "target network", that can be understood as an ad-hoc trust region to avoid violent gradient steps. Finally, it greedily optimizes a policy network: one of the main trick of the paper is to compute the gradient direction of the policy network, which uses the jacobian of the value network. But in the following implementation, this is automatically computed by
# +
# # %load tp6/ddpg.py
'''
Deep actor-critic network,
From "Continuous control with deep reinforcement learning", by <NAME> al, arXiv:1509.02971
'''
from env_pendulum import EnvPendulumSinCos; Env = lambda : EnvPendulumSinCos(1,viewer='meshcat')
import gym
import tensorflow as tf
import tensorflow.keras as tfk
import numpy as np
import matplotlib.pyplot as plt
import time
import random
from collections import deque
import signal
#######################################################################################################33
#######################################################################################################33
#######################################################################################################33
### --- Random seed
RANDOM_SEED = 0 # int((time.time()%10)*1000)
print("Seed = %d" % RANDOM_SEED)
np .random.seed (RANDOM_SEED)
random.seed (RANDOM_SEED)
tf.random.set_seed (RANDOM_SEED)
### --- Hyper paramaters
NEPISODES = 1000 # Max training steps
NSTEPS = 200 # Max episode length
QVALUE_LEARNING_RATE = 0.001 # Base learning rate for the Q-value Network
POLICY_LEARNING_RATE = 0.0001 # Base learning rate for the policy network
DECAY_RATE = 0.99 # Discount factor
UPDATE_RATE = 0.01 # Homotopy rate to update the networks
REPLAY_SIZE = 10000 # Size of replay buffer
BATCH_SIZE = 64 # Number of points to be fed in stochastic gradient
NH1 = NH2 = 250 # Hidden layer size
EXPLORATION_NOISE = 0.2
### --- Environment
# problem = "Pendulum-v1"
# env = gym.make(problem)
# NX = env.observation_space.shape[0]
# NU = env.action_space.shape[0]
# UMAX = env.action_space.high[0]
# env.reset(seed=RANDOM_SEED)
# assert( env.action_space.low[0]==-UMAX)
env = Env() # Continuous pendulum
NX = env.nx # ... training converges with q,qdot with 2x more neurones.
NU = env.nu # Control is dim-1: joint torque
UMAX = env.umax[0] # Torque range
#######################################################################################################33
### NETWORKS ##########################################################################################33
#######################################################################################################33
class QValueNetwork:
'''
Neural representaion of the Quality function:
Q: x,y -> Q(x,u) \in R
'''
def __init__(self,nx,nu,nhiden1=32,nhiden2=256,learning_rate=None):
state_input = tfk.layers.Input(shape=(nx))
state_out = tfk.layers.Dense(nhiden1, activation="relu")(state_input)
state_out = tfk.layers.Dense(nhiden1, activation="relu")(state_out)
action_input = tfk.layers.Input(shape=(nu))
action_out = tfk.layers.Dense(nhiden1, activation="relu")(action_input)
concat = tfk.layers.Concatenate()([state_out, action_out])
out = tfk.layers.Dense(nhiden2, activation="relu")(concat)
out = tfk.layers.Dense(nhiden2, activation="relu")(out)
value_output = tfk.layers.Dense(1)(out)
self.model = tfk.Model([state_input, action_input], value_output)
@tf.function
def targetAssign(self,target,tau=UPDATE_RATE):
for (tar,cur) in zip(target.model.variables,self.model.variables):
tar.assign(cur * tau + tar * (1 - tau))
class PolicyNetwork:
'''
Neural representation of the policy function:
Pi: x -> u=Pi(x) \in R^nu
'''
def __init__(self,nx,nu,umax,nhiden=32,learning_rate=None):
random_init = tf.random_uniform_initializer(minval=-0.005, maxval=0.005)
state_input = tfk.layers.Input(shape=(nx,))
out = tfk.layers.Dense(nhiden, activation="relu")(state_input)
out = tfk.layers.Dense(nhiden, activation="relu")(out)
policy_output = tfk.layers.Dense(1, activation="tanh",
kernel_initializer=random_init)(out)*umax
self.model = tfk.Model(state_input, policy_output)
@tf.function
def targetAssign(self,target,tau=UPDATE_RATE):
for (tar,cur) in zip(target.model.variables,self.model.variables):
tar.assign(cur * tau + tar * (1 - tau))
def numpyPolicy(self,x,noise=None):
'''Eval the policy with numpy input-output (nx,)->(nu,).'''
x_tf = tf.expand_dims(tf.convert_to_tensor(x), 0)
u = np.squeeze(self.model(x_tf).numpy(),0)
if noise is not None:
u = np.clip( u+noise, -UMAX,UMAX)
return u
def __call__(self, x,**kwargs):
return self.numpyPolicy(x,**kwargs)
#######################################################################################################33
class OUNoise:
'''
Ornstein–Uhlenbeck processes are markov random walks with the nice property to eventually
converge to its mean.
We use it for adding some random search at the begining of the exploration.
'''
def __init__(self, mean, std_deviation, theta=0.15, dt=1e-2, y_initial=None,dtype=np.float32):
self.theta = theta
self.mean = mean.astype(dtype)
self.std_dev = std_deviation.astype(dtype)
self.dt = dt
self.dtype=dtype
self.reset(y_initial)
def __call__(self):
# Formula taken from https://www.wikipedia.org/wiki/Ornstein-Uhlenbeck_process.
noise = np.random.normal(size=self.mean.shape).astype(self.dtype)
self.y += \
self.theta * (self.mean - self.y) * self.dt \
+ self.std_dev * np.sqrt(self.dt) * noise
return self.y.copy()
def reset(self,y_initial = None):
self.y = y_initial.astype(self.dtype) if y_initial is not None else np.zeros_like(self.mean)
### --- Replay memory
class ReplayItem:
'''
Storage for the minibatch
'''
def __init__(self,x,u,r,d,x2):
self.x = x
self.u = u
self.reward = r
self.done = d
self.x2 = x2
#######################################################################################################33
quality = QValueNetwork(NX,NU,NH1,NH2)
qualityTarget = QValueNetwork(NX,NU,NH1,NH2)
quality.targetAssign(qualityTarget,1)
policy = PolicyNetwork(NX,NU,umax=UMAX,nhiden=NH2)
policyTarget = PolicyNetwork(NX,NU,umax=UMAX,nhiden=NH2)
policy.targetAssign(policyTarget,1)
replayDeque = deque()
ou_noise = OUNoise(mean=np.zeros(1), std_deviation=float(EXPLORATION_NOISE) * np.ones(1))
ou_noise.reset( np.array([ UMAX/2 ]) )
#######################################################################################################33
### MAIN ACTOR-CRITIC BLOCK
#######################################################################################################33
critic_optimizer = tfk.optimizers.Adam(QVALUE_LEARNING_RATE)
actor_optimizer = tfk.optimizers.Adam(POLICY_LEARNING_RATE)
@tf.function
def learn(state_batch, action_batch, reward_batch, next_state_batch):
'''
<learn> is isolated in a tf.function to make it more efficient.
@tf.function forces tensorflow to optimize the inner computation graph defined in this function.
'''
# Automatic differentiation of the critic loss, using tf.GradientTape
# The critic loss is the classical Q-learning loss:
# loss = || Q(x,u) - (reward + Q(xnext,Pi(xnexT)) ) ||**2
with tf.GradientTape() as tape:
target_actions = policyTarget.model(next_state_batch, training=True)
y = reward_batch + DECAY_RATE * qualityTarget.model(
[next_state_batch, target_actions], training=True
)
critic_value = quality.model([state_batch, action_batch], training=True)
critic_loss = tf.math.reduce_mean(tf.math.square(y - critic_value))
critic_grad = tape.gradient(critic_loss, quality.model.trainable_variables)
critic_optimizer.apply_gradients(
zip(critic_grad, quality.model.trainable_variables)
)
# Automatic differentiation of the actor loss, using tf.GradientTape
# The actor loss implements a greedy optimization on the quality function
# loss(u) = Q(x,u)
with tf.GradientTape() as tape:
actions = policy.model(state_batch, training=True)
critic_value = quality.model([state_batch, actions], training=True)
actor_loss = -tf.math.reduce_mean(critic_value)
actor_grad = tape.gradient(actor_loss, policy.model.trainable_variables)
actor_optimizer.apply_gradients(
zip(actor_grad, policy.model.trainable_variables)
)
#######################################################################################################33
#######################################################################################################33
#######################################################################################################33
def rendertrial(maxiter=NSTEPS,verbose=True):
'''
Display a roll-out from random start and optimal feedback.
Press ^Z to get a roll-out at training time.
'''
x = env.reset()
rsum = 0.
for i in range(maxiter):
u = policy(x)
x, reward = env.step(u)[:2]
env.render()
rsum += reward
if verbose: print('Lasted ',i,' timestep -- total reward:',rsum)
signal.signal(signal.SIGTSTP, lambda x,y:rendertrial()) # Roll-out when CTRL-Z is pressed
env.full.sleepAtDisplay=5e-3
# Logs
h_rewards = []
h_steps = []
# Takes about 4 min to train
for episode in range(NEPISODES):
prev_state = env.reset()
for step in range(NSTEPS):
# Uncomment this to see the Actor in action
# But not in a python notebook.
#env.render()
action = policy(prev_state, noise=ou_noise())
state, reward = env.step(action)[:2]
done=False
replayDeque.append(ReplayItem(prev_state, action, reward, done, state))
prev_state = state
if len(replayDeque) <= BATCH_SIZE: continue
####################################################################
# Sample a minibatch
batch = random.sample(replayDeque,BATCH_SIZE) # Random batch from replay memory.
state_batch = tf.convert_to_tensor([ b.x for b in batch ])
action_batch = tf.convert_to_tensor([ b.u for b in batch ])
reward_batch = tf.convert_to_tensor([ [ b.reward ] for b in batch ],dtype=np.float32)
done_batch = tf.convert_to_tensor([ b.done for b in batch ])
next_state_batch = tf.convert_to_tensor([ b.x2 for b in batch ])
####################################################################
# One gradient step for the minibatch
# Critic and actor gradients
learn(state_batch, action_batch, reward_batch, next_state_batch)
# Step smoothing using target networks
policy.targetAssign(policyTarget)
quality.targetAssign(qualityTarget)
if done: break # stop at episode end.
# Some prints and logs
episodic_reward = sum([ replayDeque[-i-1].reward for i in range(step+1) ])
h_rewards.append( episodic_reward )
h_steps.append(step+1)
print(f'Ep#{episode:3d}: lasted {step+1:d} steps, reward={episodic_reward:3.1f} ')
# avg_reward = np.mean(h_rewards[-40:])
# if episode==5 and RANDOM_SEED==0:
# assert( abs(avg_reward + 1423.0528188196286) < 1e-3 )
# if episode==0 and RANDOM_SEED==0:
# assert( abs(avg_reward + 1712.386325099637) < 1e-3 )
# Plotting graph
# Episodes versus Avg. Rewards
plt.plot(h_rewards)
plt.xlabel("Episode")
plt.ylabel("Epsiodic Reward")
plt.show()
#######################################################################################################33
#######################################################################################################33
#######################################################################################################33
# -
| 6_reinforcement_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importing the libraries
import numpy as np
import cv2
# ## Storing Image1 in a variable & displaying it
image1 = cv2.imread("image1.jpg")
cv2.imshow("1st Image", image1)
cv2.waitKey()
cv2.destroyAllWindows()
# ## Storing Image2 in a variable & displaying it
image2 = cv2.imread("image2.jpg")
cv2.imshow("2nd Image", image2)
cv2.waitKey()
cv2.destroyAllWindows()
# ## Cropping 1st Image & showing only the photo frame
image1_cropped = image1[60:410,190:610]
cv2.imshow("1st Cropped Image", image1_cropped)
cv2.waitKey()
cv2.destroyAllWindows()
# ## Cropping 2nd Image & showing only the photo frame
image2_cropped = image2[110:385,200:564]
cv2.imshow("2nd Cropped Image", image2_cropped)
cv2.waitKey()
cv2.destroyAllWindows()
# ## Resizing both cropped frames
image1_resized = cv2.resize(image1_cropped,(image2_cropped.shape[1],image2_cropped.shape[0]))
image2_resized = cv2.resize(image2_cropped,(image1_cropped.shape[1],image1_cropped.shape[0]))
# ## Swapping the photo frame & Showing 1st updated image
image1[60:410,190:610] = image2_resized
cv2.imshow("1st Image", image1)
cv2.waitKey()
cv2.destroyAllWindows()
# ## Swapping the photo frame & Showing 1st updated image
image2[110:385,200:564] = image1_resized
cv2.imshow("1st Image", image2)
cv2.waitKey()
cv2.destroyAllWindows()
# ## Saving both the new images
cv2.imwrite("image1_new.jpg", image1)
cv2.imwrite("image2_new.jpg", image2)
| Swapping Images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
'''
This is a baseline solution, original source code for all methods can be found at
https://github.com/konstmish/rr_prox_fed
'''
import matplotlib
import numpy as np
import psutil
import ray
import seaborn as sns
import matplotlib.pyplot as plt
import numpy.linalg as la
from matplotlib import rc, rcParams
from sklearn.datasets import load_svmlight_file, fetch_rcv1
from src.local_algorithms import LocalSgd, LocalShuffling
from src.datasets import get_dataset
from src.first_order import RestNest
from src.loss_functions import LogisticRegression
from src.utils import get_trace, relative_round
sns.set(style="whitegrid", context="talk", palette=sns.color_palette("bright"), color_codes=False)
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
matplotlib.rcParams['font.family'] = 'sans-serif'
matplotlib.rcParams['font.sans-serif'] = ['DejaVu Sans']
matplotlib.rcParams['mathtext.fontset'] = 'cm'
# Multiple plots in one figure: code below
# import matplotlib.backends.backend_pdf as bf
# <code for plt>
# pdf = bf.PdfPages("./output.pdf")
# for fig in xrange(1, plt.figure().number):
# pdf.savefig( fig )
# pdf.close()
# -
num_cpus = psutil.cpu_count(logical=False)
ray.init(num_cpus=num_cpus)
A, b = get_dataset('a1a', data_path='src/datasets/')
l1 = 0
loss = LogisticRegression(A, b, l1=l1, l2=0)
n, dim = A.shape
if n <= 20000 or dim <= 20000:
print('Computing the smoothness constant via SVD, it may take a few minutes...')
L = loss.smoothness
l2 = L / n
loss.l2 = l2
x0 = np.zeros(dim)
n_epoch = 100
batch_size = 8
it_max = (n_epoch * n) // batch_size
trace_len = 300
rest = RestNest(loss=loss, it_max=4000, doubling=True)
rest_tr = rest.run(x0=x0)
rest_tr.compute_loss_of_iterates()
rest.trace.plot_losses()
plt.yscale('log')
plt.savefig('RestNest.png', dpi=300, bbox_inches='tight')
it_local = 200
sgd_lr0 = 1 / loss.batch_smoothness(batch_size=batch_size) / it_local
sgd_decay_coef = l2 / 2
sgd_lr_max = sgd_lr0
sgd = LocalSgd(loss=loss, it_local=it_local, lr_max=sgd_lr_max, lr0=sgd_lr0, lr_decay_coef=sgd_decay_coef, it_start_decay=0,
it_max=n*100//(batch_size*it_local), n_seeds=1, batch_size=batch_size, trace_len=100, iid=True)
sgd.run(x0=x0)
sgd.trace.convert_its_to_epochs(batch_size=batch_size*it_local)
sgd.trace.compute_loss_of_iterates()
rr_lr0 = 1 / loss.batch_smoothness(batch_size=batch_size)
rr_decay_coef = l2 / 7
rr_lr_max = rr_lr0
rr = LocalShuffling(reshuffle=True, loss=loss, lr_max=rr_lr_max, lr0=rr_lr0, lr_decay_coef=rr_decay_coef, iid=True,
epoch_start_decay=0, it_max=100, n_seeds=1, batch_size=batch_size, trace_len=100)
rr.run(x0=x0)
rr.trace.compute_loss_of_iterates()
plt.figure(figsize=(9, 6))
sgd.trace.plot_losses(label='Local-SGD')
rr.trace.plot_losses(label='FedRR', marker='o')
plt.yscale('log')
plt.legend()
plt.savefig('fedVSsgd_a1a.png', dpi=300, bbox_inches='tight')
| code/run_and_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Simplification
from sympy import *
x, y, z = symbols('x y z')
init_printing()
# For each exercise, fill in the function according to its docstring.
# ## Polynomial/Rational Function Simplification
# In each exercise, apply specific simplification functions to get the desired result.
def polysimp1(expr):
"""
>>> polysimp1(cos(x)*sin(x) + cos(x))
(sin(x) + 1)*cos(x)
>>> polysimp1(cos(x)*sin(x) + cos(x) + 1)
(sin(x) + 1)*cos(x) + 1
"""
return collect(expr, cos(x))
polysimp1(cos(x)*sin(x) + cos(x))
polysimp1(cos(x)*sin(x) + cos(x) + 1)
def polysimp2(expr):
"""
>>> polysimp2((2*x + 1)/(x**2 + x))
1/(x + 1) + 1/x
>>> polysimp2((x**2 + 3*x + 1)/(x**3 + 2*x**2 + x))
1/(x**2 + 2*x + 1) + 1/x
"""
return apart(expr, x)
polysimp2((2*x + 1)/(x**2 + x))
polysimp2((x**2 + 3*x + 1)/(x**3 + 2*x**2 + x))
# ## Powers
# In each exercise, apply specific simplification functions to get the desired result.
def powersimp1(expr):
"""
>>> powersimp1(exp(x)*(exp(y) + 1))
exp(x) + exp(x + y)
"""
return powsimp(expand(expr))
powersimp1(exp(x)*(exp(y) + 1))
def powersimp2(expr):
"""
>>> powersimp2(2**x*x**x)
(2*x)**x
>>> powersimp2(x**x*x**x)
(x**2)**x
"""
return powsimp(expr, force=True)
powersimp2(2**x*x**x)
powersimp2(x**x*x**x)
def powersimp3(expr):
"""
>>> a, b, c = symbols('a b c')
>>> powersimp3((a**b)**c)
a**(b*c)
>>> powersimp3((a**b)**(c + 1))
a**(b*c + b)
"""
return powdenest(expand_power_exp(expr), force=True)
a, b, c = symbols('a b c')
expand_power_exp((a**b)**(c + 1))
powersimp3((a**b)**c)
powersimp3((a**b)**(c + 1))
# ## Logs
def logsimp1(expr):
"""
>>> a, b = symbols('a b', positive=True)
>>> logsimp1(log(x**y*a**b))
y*log(x) + log(a**b)
>>> logsimp1(log(x*y*a*b))
log(x) + log(y) + log(a*b)
"""
return logcombine(expand_log(expr, force=True))
a, b = symbols('a b', positive=True)
logsimp1(log(x**y*a**b))
logsimp1(log(x*y*a*b))
# ## Miscellaneous
def miscsimp1(expr):
"""
>>> miscsimp1(sin(x + y))
2*(-tan(x/2)**2 + 1)*tan(y/2)/((tan(x/2)**2 + 1)*(tan(y/2)**2 + 1)) + 2*(-tan(y/2)**2 + 1)*tan(x/2)/((tan(x/2)**2 + 1)*(tan(y/2)**2 + 1))
"""
return expand_trig(expr).rewrite(tan)
miscsimp1(sin(x + y))
def miscsimp2(expr):
"""
>>> miscsimp2(gamma(x + 4))
x**4*gamma(x) + 6*x**3*gamma(x) + 11*x**2*gamma(x) + 6*x*gamma(x)
"""
return expand(expr, func=True)
miscsimp2(gamma(x + 4))
# ## Continued Fractions
# If we do not cover this, see http://docs.sympy.org/latest/tutorial/simplification.html#example-continued-fractions
def list_to_frac(l):
expr = Integer(0)
for i in reversed(l[1:]):
expr += i
expr = 1/expr
return l[0] + expr
a0, a1, a2, a3, a4 = symbols('a0:5')
# Determine the list used to create the continued fraction $$\frac{a_{0} a_{1} a_{2} a_{3} a_{4} + a_{0} a_{1} a_{2} + a_{0} a_{3} a_{4} + a_{0} + a_{1} a_{2} a_{3} + a_{1} a_{3} a_{4} + a_{1} + a_{3}}{a_{0} a_{1} a_{2} a_{4} + a_{0} a_{4} + a_{1} a_{2} + a_{1} a_{4} + 1}.$$
def continued_frac():
"""
Determine the original list used to create the fraction.
Return the original list from this function.
>>> orig_frac = (a0*a1*a2*a3*a4 + a0*a1*a2 + a0*a3*a4 + a0 + a1*a2*a3 + a1*a3*a4 + a1 + a3)/(a0*a1*a2*a4 + a0*a4 + a1*a2 + a1*a4 + 1)
>>> pprint(orig_frac)
a₀⋅a₁⋅a₂⋅a₃⋅a₄ + a₀⋅a₁⋅a₂ + a₀⋅a₃⋅a₄ + a₀ + a₁⋅a₂⋅a₃ + a₁⋅a₃⋅a₄ + a₁ + a₃
─────────────────────────────────────────────────────────────────────────
a₀⋅a₁⋅a₂⋅a₄ + a₀⋅a₄ + a₁⋅a₂ + a₁⋅a₄ + 1
>>> cancel(list_to_frac(continued_frac())) == orig_frac
True
"""
return [a3, a4, a0, a2, a1]
orig_frac = (a0*a1*a2*a3*a4 + a0*a1*a2 + a0*a3*a4 + a0 + a1*a2*a3 + a1*a3*a4 + a1 + a3)/(a0*a1*a2*a4 + a0*a4 + a1*a2 + a1*a4 + 1)
orig_frac
cancel(list_to_frac(continued_frac())) == orig_frac
| tutorial_exercises/Advanced-Simplification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/VinceDeilord/CPEN21A-ECE-2-1/blob/main/Operations_and_Expressions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8vcYfFHx3szz"
# ##Boolean Opereator
# + colab={"base_uri": "https://localhost:8080/"} id="9WV6uNCX33go" outputId="fc82db06-660b-4ac0-c04f-9b240ceeeda1"
x=1
y=2
print(x>y)
print(10>11)
print(10==10)
# + id="34Csknux4w0Y"
#using bool
# + [markdown] id="5Xg3t0Zk5XGN"
# ###fuctions using return boolean
# + colab={"base_uri": "https://localhost:8080/"} id="Odn4-K1R5csu" outputId="56e507d8-3f0a-4498-8574-f78008ece59c"
def myfunction():return True
print(myfunction())
# + colab={"base_uri": "https://localhost:8080/"} id="J66wzaHM52SU" outputId="538c68df-158c-4259-9a41-a1163d6cbb68"
def yourfunction():return True
if yourfunction():
print("Yes!")
else:
print("No!")
# + [markdown] id="xr17uEtQ6mt8"
# ##Boolean Operator
# + colab={"base_uri": "https://localhost:8080/"} id="cu-IiGyA6fUw" outputId="53612b7a-82e7-439b-9a3e-29e25b2cc77a"
print (10>9)
a=6
b=7
print(a==b)
print(a!=a)
# + [markdown] id="1l-Jar5s8WjT"
# ###Arithmetic Operators
# + colab={"base_uri": "https://localhost:8080/"} id="RKnq9AFf8Pp4" outputId="1b988e97-c254-44cf-e6ac-8ac5a0221420"
print(10+5)
print(10-5)
print(10*5)
print(10/5)
print(10%5)
print(10//5)
print(10**5)
# + [markdown] id="seqFqie-9aYo"
# ##Bitwise Operators
# + colab={"base_uri": "https://localhost:8080/"} id="66uwptZy9fhN" outputId="da844ff6-4a68-4a69-830c-f1651573d7a6"
a=60 #0011 1100
b=13 #0000 1101
print(a&b)
print(a|b)
print(a^b)
print(~a)
print(a<<1) #0111 1000
print(a<<2) #1111 0000
print(b>>1)
print(b>>2)
# + [markdown] id="LkxwDWlBBwef"
# ##Python Assignment Operators
# + colab={"base_uri": "https://localhost:8080/"} id="M-IqaizUB5NG" outputId="2fe092da-d11a-4431-eca0-03641c29e70b"
a+=3 #Same As a = a + 3
#Same As a = 60 + 3, a = 63
print(a)
# + [markdown] id="DPzv9bTCCpgd"
# ##Logical Operators
# + colab={"base_uri": "https://localhost:8080/"} id="aSU2ZwZnCsm_" outputId="f1b5b360-39ec-46ee-b849-7eafa4e11cf3"
#and logical operator
a= True
b= False
print(a and b)
print(not(a and b))
print(a or B)
print(not(a or b))
# + colab={"base_uri": "https://localhost:8080/"} id="H7y72W-cDMe_" outputId="49d1c7d9-43da-4d3b-bd6d-8a237c27272e"
print(a is b)
print(a is not b)
| Operations_and_Expressions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.018006, "end_time": "2021-05-10T05:53:58.447237", "exception": false, "start_time": "2021-05-10T05:53:58.429231", "status": "completed"} tags=[]
# # Setup Environment
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 15.595292, "end_time": "2021-05-10T05:54:14.059422", "exception": false, "start_time": "2021-05-10T05:53:58.464130", "status": "completed"} tags=[]
import math, os, re, time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from imblearn.over_sampling import SMOTE
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import precision_score, recall_score, roc_auc_score, confusion_matrix, classification_report, roc_curve, auc
from sklearn.linear_model import LogisticRegressionCV
from sklearn.ensemble import RandomForestClassifier, StackingClassifier
from xgboost import XGBClassifier
from lightgbm import LGBMClassifier
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
import shap
from tqdm.notebook import tqdm
# %matplotlib inline
tqdm.pandas()
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# + [markdown] papermill={"duration": 0.017574, "end_time": "2021-05-10T05:54:14.094970", "exception": false, "start_time": "2021-05-10T05:54:14.077396", "status": "completed"} tags=[]
# # Load Data
# + papermill={"duration": 0.571368, "end_time": "2021-05-10T05:54:14.683474", "exception": false, "start_time": "2021-05-10T05:54:14.112106", "status": "completed"} tags=[]
train = pd.read_csv('/kaggle/input/GiveMeSomeCredit/cs-training.csv', index_col=0)
test = pd.read_csv('/kaggle/input/GiveMeSomeCredit/cs-test.csv', index_col=0)
renamed = {'SeriousDlqin2yrs':'default_occurred',
'RevolvingUtilizationOfUnsecuredLines':'utilization_of_unsecured_loc',
'NumberOfTime30-59DaysPastDueNotWorse':'30_to_59_days_pastdue_count',
'DebtRatio':'debt_ratio',
'MonthlyIncome':'monthly_income',
'NumberOfOpenCreditLinesAndLoans':'open_loc_and_loan_count',
'NumberOfTimes90DaysLate':'90_days_late_count',
'NumberRealEstateLoansOrLines':'re_loans_and_lines_count',
'NumberOfTime60-89DaysPastDueNotWorse':'60_to_89_days_pastdue_count',
'NumberOfDependents':'dependents'}
train.rename(columns=renamed, inplace=True)
test.rename(columns=renamed, inplace=True)
train
# + [markdown] papermill={"duration": 0.017572, "end_time": "2021-05-10T05:54:14.719036", "exception": false, "start_time": "2021-05-10T05:54:14.701464", "status": "completed"} tags=[]
# # Data Cleaning & Feature Engineering
# + [markdown] papermill={"duration": 0.017349, "end_time": "2021-05-10T05:54:14.754875", "exception": false, "start_time": "2021-05-10T05:54:14.737526", "status": "completed"} tags=[]
# ## Fill 'dependents' NaN values with 0
# + papermill={"duration": 0.129361, "end_time": "2021-05-10T05:54:14.902295", "exception": false, "start_time": "2021-05-10T05:54:14.772934", "status": "completed"} tags=[]
train['dependents'].fillna(value=0, inplace=True)
train['dependents'] = train['dependents'].apply(int)
test['dependents'].fillna(value=0, inplace=True)
test['dependents'] = test['dependents'].apply(int)
train
# + [markdown] papermill={"duration": 0.018119, "end_time": "2021-05-10T05:54:14.939531", "exception": false, "start_time": "2021-05-10T05:54:14.921412", "status": "completed"} tags=[]
# ## Binning and One-Hot Encoding Income Levels
# + papermill={"duration": 0.026277, "end_time": "2021-05-10T05:54:14.984233", "exception": false, "start_time": "2021-05-10T05:54:14.957956", "status": "completed"} tags=[]
def income_binned(data):
# Level 0 = NaN income values
if np.isnan(data):
return 0
# Level 1 = 0 to 20k
elif data <= 20000:
return 1
# Level 2 = 20k to 40k
elif data <= 40000:
return 2
# Level 3 = 40k to 60k
elif data <= 60000:
return 3
# Level 4 = 60k to 80k
elif data <= 80000:
return 4
# Level 5 = 80k to 100k
elif data <= 100000:
return 5
# Level 6 = Above 100k
else:
return 6
# + papermill={"duration": 0.541865, "end_time": "2021-05-10T05:54:15.546211", "exception": false, "start_time": "2021-05-10T05:54:15.004346", "status": "completed"} tags=[]
# Binning Income Levels
train['monthly_income_binned'] = train['monthly_income'].apply(income_binned)
test['monthly_income_binned'] = test['monthly_income'].apply(income_binned)
# Dropping original Income Levels
train.drop(columns=['monthly_income'], inplace=True)
test.drop(columns=['monthly_income'], inplace=True)
# Creating Income Level dummy variables
dummy_names = ['monthly_income_binned_' + str(i) for i in range(1,7)]
one_hot_encoded = pd.get_dummies(train['monthly_income_binned'], drop_first=True)
one_hot_encoded.columns = dummy_names
train = pd.concat([train, one_hot_encoded], axis=1)
one_hot_encoded = pd.get_dummies(test['monthly_income_binned'], drop_first=True)
one_hot_encoded.columns = dummy_names
test = pd.concat([test, one_hot_encoded], axis=1)
# Drop binned incomes after creating dummy variables
train.drop(columns=['monthly_income_binned'], inplace=True)
test.drop(columns=['monthly_income_binned'], inplace=True)
train
# + [markdown] papermill={"duration": 0.029082, "end_time": "2021-05-10T05:54:15.604201", "exception": false, "start_time": "2021-05-10T05:54:15.575119", "status": "completed"} tags=[]
# ## Aggregate Past Due Counts
# + papermill={"duration": 0.06587, "end_time": "2021-05-10T05:54:15.699332", "exception": false, "start_time": "2021-05-10T05:54:15.633462", "status": "completed"} tags=[]
train['above_30_days_pastdue_count'] = train['30_to_59_days_pastdue_count'] + train['60_to_89_days_pastdue_count'] + train['90_days_late_count']
train.drop(columns=['30_to_59_days_pastdue_count',
'60_to_89_days_pastdue_count',
'90_days_late_count'], inplace=True)
test['above_30_days_pastdue_count'] = test['30_to_59_days_pastdue_count'] + test['60_to_89_days_pastdue_count'] + test['90_days_late_count']
test.drop(columns=['30_to_59_days_pastdue_count',
'60_to_89_days_pastdue_count',
'90_days_late_count'], inplace=True)
train
# + papermill={"duration": 1.478567, "end_time": "2021-05-10T05:54:17.207887", "exception": false, "start_time": "2021-05-10T05:54:15.729320", "status": "completed"} tags=[]
# Correlation Matrix (Heatmap)
corr = train.corr(method='pearson')
plt.figure(figsize=(15, 15))
sns.heatmap(corr, annot=True, cmap="YlGnBu",cbar_kws={'label': 'Correlation'})
plt.tight_layout()
plt.xticks(rotation=45, horizontalalignment='right')
plt.yticks(rotation=0)
plt.show()
plt.clf()
# + [markdown] papermill={"duration": 0.034034, "end_time": "2021-05-10T05:54:17.276517", "exception": false, "start_time": "2021-05-10T05:54:17.242483", "status": "completed"} tags=[]
# # Resampling (for Imbalanced Dataset)
# + papermill={"duration": 0.073296, "end_time": "2021-05-10T05:54:17.383771", "exception": false, "start_time": "2021-05-10T05:54:17.310475", "status": "completed"} tags=[]
X_train = train.iloc[:,1:]
y_train = train.iloc[:,0:1]
X_test = test.iloc[:,1:]
y_test = test.iloc[:,0:1]
X_train
# + papermill={"duration": 0.052261, "end_time": "2021-05-10T05:54:17.472319", "exception": false, "start_time": "2021-05-10T05:54:17.420058", "status": "completed"} tags=[]
y_train
# + papermill={"duration": 0.176254, "end_time": "2021-05-10T05:54:17.686119", "exception": false, "start_time": "2021-05-10T05:54:17.509865", "status": "completed"} tags=[]
sns.countplot(x='default_occurred',data=y_train)
# + papermill={"duration": 1.025672, "end_time": "2021-05-10T05:54:18.737856", "exception": false, "start_time": "2021-05-10T05:54:17.712184", "status": "completed"} tags=[]
smote = SMOTE(sampling_strategy=30/70, random_state=42, n_jobs=-1)
X_train, y_train = smote.fit_resample(X_train, y_train)
# + papermill={"duration": 0.184018, "end_time": "2021-05-10T05:54:18.948116", "exception": false, "start_time": "2021-05-10T05:54:18.764098", "status": "completed"} tags=[]
sns.countplot(x='default_occurred',data=y_train)
# + [markdown] papermill={"duration": 0.026871, "end_time": "2021-05-10T05:54:19.002115", "exception": false, "start_time": "2021-05-10T05:54:18.975244", "status": "completed"} tags=[]
# # Feature Scaling
# + papermill={"duration": 0.063677, "end_time": "2021-05-10T05:54:19.092812", "exception": false, "start_time": "2021-05-10T05:54:19.029135", "status": "completed"} tags=[]
X_train = pd.concat([X_train.iloc[:,:6],
X_train.iloc[:,-1],
X_train.iloc[:,6:-1]],axis=1)
X_test = pd.concat([X_test.iloc[:,:6],
X_test.iloc[:,-1],
X_test.iloc[:,6:-1]],axis=1)
X_train
# + papermill={"duration": 0.058228, "end_time": "2021-05-10T05:54:19.179792", "exception": false, "start_time": "2021-05-10T05:54:19.121564", "status": "completed"} tags=[]
X_test
# + papermill={"duration": 0.119696, "end_time": "2021-05-10T05:54:19.327510", "exception": false, "start_time": "2021-05-10T05:54:19.207814", "status": "completed"} tags=[]
scaler = StandardScaler()
scale_vars = X_train.loc[:,:'above_30_days_pastdue_count'].columns
# Fit and Scale X_train
scaled = pd.DataFrame(scaler.fit_transform(X_train.loc[:,:'above_30_days_pastdue_count'].copy()), columns=scale_vars, index=X_train.index)
X_train = pd.concat([scaled,X_train.loc[:,'monthly_income_binned_1':]], axis=1)
# Scale X_var using the X_train fitted scaler (prevents data leakage)
scaled = pd.DataFrame(scaler.transform(X_test.loc[:,:'above_30_days_pastdue_count'].copy()), columns=scale_vars, index=X_test.index)
X_test = pd.concat([scaled,X_test.loc[:,'monthly_income_binned_1':]], axis=1)
X_train
# + papermill={"duration": 0.070952, "end_time": "2021-05-10T05:54:19.440137", "exception": false, "start_time": "2021-05-10T05:54:19.369185", "status": "completed"} tags=[]
X_test
# + [markdown] papermill={"duration": 0.043673, "end_time": "2021-05-10T05:54:19.526634", "exception": false, "start_time": "2021-05-10T05:54:19.482961", "status": "completed"} tags=[]
# # Final Modelling (Training & Prediction)
# + papermill={"duration": 9.762818, "end_time": "2021-05-10T05:54:29.332262", "exception": false, "start_time": "2021-05-10T05:54:19.569444", "status": "completed"} tags=[]
best_model = XGBClassifier(n_estimators=100,
learning_rate=0.05,
objective='binary:logistic',
max_depth=5,
subsample=0.9,
reg_alpha=0.01,
reg_lambda=0.5,
use_label_encoder=False,
n_jobs=-1,
random_state=42)
best_model.fit(X_train, y_train.values.ravel())
y_pred = best_model.predict(X_test)
y_proba = best_model.predict_proba(X_test)[:,1]
# + [markdown] papermill={"duration": 0.051331, "end_time": "2021-05-10T05:54:29.428851", "exception": false, "start_time": "2021-05-10T05:54:29.377520", "status": "completed"} tags=[]
# # Saving Output
# + papermill={"duration": 0.246066, "end_time": "2021-05-10T05:54:29.730422", "exception": false, "start_time": "2021-05-10T05:54:29.484356", "status": "completed"} tags=[]
ids = np.arange(1,101504)
submission = pd.DataFrame({'Id': ids,
'Probability': y_proba})
submission.to_csv("/kaggle/working/credit_score_submision.csv", index=False)
# + [markdown] papermill={"duration": 0.044239, "end_time": "2021-05-10T05:54:29.818761", "exception": false, "start_time": "2021-05-10T05:54:29.774522", "status": "completed"} tags=[]
# # Feature Importance (SHAP)
# + papermill={"duration": 38.740874, "end_time": "2021-05-10T05:55:08.604530", "exception": false, "start_time": "2021-05-10T05:54:29.863656", "status": "completed"} tags=[]
# DF, based on which importance is checked
X_importance = X_train
# Explain model predictions using shap library:
explainer = shap.TreeExplainer(best_model)
shap_values = explainer.shap_values(X_importance)
# + papermill={"duration": 17.133081, "end_time": "2021-05-10T05:55:25.782958", "exception": false, "start_time": "2021-05-10T05:55:08.649877", "status": "completed"} tags=[]
# Plot summary_plot
shap.summary_plot(shap_values, X_importance)
# + papermill={"duration": 0.281758, "end_time": "2021-05-10T05:55:26.111000", "exception": false, "start_time": "2021-05-10T05:55:25.829242", "status": "completed"} tags=[]
shap.summary_plot(shap_values, X_importance, plot_type='bar')
| Final Model Prediction, Feature Importance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from Py3DViewer import Trimesh, Quadmesh, Tetmesh, Hexmesh
from Py3DViewer import Viewer, laplacian_smoothing
from Py3DViewer import AABB, SpaceObject, Octree, NOctreeNode, NOctree, Skeleton, PointCloud
import numpy as np
import time
from numba.typed import List
# +
#Build of the octree
l=List()
t=Trimesh('data/cactus.obj')
for p in t.polys:
l.append(SpaceObject(np.array(t.vertices[p],dtype='float64')))
# %time o=Octree(10,8,l,t.vertices)
# -
#Search of a point in the octree
p = o.n.shapes[500].vertices[0]
# %time o.search_point(type(t).__name__,p)
# +
#Intersection of a ray with the octree
ori = np.array([p[0], p[1], p[2]+10],dtype='float64')
d = np.array([0, 0, -1],dtype='float64')
# %time o.intersects_ray(type(t).__name__,ori,d)
# +
#View of the octree
vertices,edges=AABB.get_all_vertices_and_edges(o.n.aabbs)
vertices=np.vstack(vertices)
edges=np.array(edges)
s = Skeleton(None,vertices,edges)
viewer = Viewer([s,t], mesh_color=np.array([214.,214.,214.]) ,width=1000, height=500)
viewer.show()
# -
mesh = [Trimesh('data/bunny_tris.obj'), Quadmesh('data/bunny_quad.obj')]
"""
After running this cell, you will notice that the quad bunny is quite larger than the tri bunny,
and is also oriented differently.
"""
viewer = Viewer(mesh, width=1000, height=300)
viewer.show()
# +
"""
Let's first scale the meshes to have a bounding box diagonal of 10, then update the viewer.
NOTE: The viewer update takes some time depending
on the quality of your connect, since the new geometry is streamed from the backend to your browser.
"""
mesh[0].vertices /= np.linalg.norm(mesh[0].bbox[1]-mesh[0].bbox[0])
mesh[0].vertices *= 10
mesh[1].vertices /= np.linalg.norm(mesh[1].bbox[1]-mesh[1].bbox[0])
mesh[1].vertices *=10
viewer.update_controls()
viewer.update()
# -
"""
Let's rotate the quad bunny by using the transform_rotation method
"""
mesh[1].transform_rotation(-90, 'x')
viewer.update()
"""
Let's now do the same with a "reactive" approach.
When using a reactive viewer, the user can simply act on the mesh data however they[-5.94054e-01 -5.72423e-04 -1.45624e+00] MAX CHILDREN: [0. 0.24394329 0. ] choose, and the viewer
is asynchronously updated (with a minimal, if any, overhead on the computation).
Again, consider that the redraws are strongly bound by the fact that the Python and Javascript backend are
de-coupled in this context, since Binder runs in the cloud. We highly suggest trying the library locally to
properly experience the reactive redraws.
"""
mesh2 = [Trimesh('data/tri_326064.obj')]
viewer2 = Viewer(mesh2, width=1000, height=500, reactive=True)
viewer2.show()
mesh2[0].vertices /= np.linalg.norm(mesh2[0].bbox[1]-mesh2[0].bbox[0])
mesh2[0].vertices *= 10
mesh2[1].vertices /= np.linalg.norm(mesh2[1].bbox[1]-mesh2[1].bbox[0])
mesh2[1].vertices *=10
viewer2.update_controls()
mesh2[1].transform_rotation(-90, 'x')
"""
Here's an example with the GUI, which only works with a single mesh per scene so far. We are working to support multiple meshes.
"""
mesh3 = Hexmesh('data/double_hinge_hex.mesh')
viewer3 = Viewer(t, width=1000, height=300, with_gui=True)
viewer3.show()
"""
Py3DViewer have texture support for surface meshes.
These are some examples.
"""
spot = Trimesh('data/spot.obj', texture='data/textures/spot_texture.png')
blub = Trimesh('data/blub.obj', texture='data/textures/blub_texture.png')
cube_tris = Trimesh('data/cube_tris_mtl.obj', mtl='data/cube.mtl')
cube_quad = Quadmesh('data/cube_quad_mtl.obj', mtl='data/cube.mtl')
spot.show()
blub.show()
cube_tris.show()
cube_quad.show()
| Playground.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.1
# language: julia
# name: julia-1.1
# ---
include("setup.jl")
# # Asymmetric Travelling Salesman Problem
#
# **Originally Contributed by**: <NAME> and <NAME>
#
# https://www.movable-type.co.uk/scripts/latlong.html
# ### Problem description
#
# $$
# \begin{array}{cl}
# \min_{x} \ \ \ &
# \sum_{i, j} d_{i, j} x_{i, j}\\
# s.t. &
# \sum_{j} x_{i, j} = 1, \ \ \ \forall i \in \Omega\\
# & \sum_{i} x_{i, j} = 1, \ \ \ \forall j \in \Omega\\
# & x_{k,k} = 0 \ \ \ \forall k \in \Omega\\
# & \sum_{i} \sum_{j \ne i} x_{i,j} \le |S| - 1, \forall S \subset \Omega, 2 \le |S| \le N-1\\
# & x_{i, j} \in \{0, 1\}, \ \ \ \forall (i,j) \in \Omega^2 , i \ne j\\
# & \Omega = \{1, \dots, N\}
# \end{array}
# $$
d = readdlm("./tsp/distances_quebec.txt")
c = readdlm("./tsp/cities_quebec.txt")
"""
all_subsets(x)
Compute all subsets of elements of vector `x`.
"""
function all_subsets(x::Vector{T}) where T
res = Vector{T}[[]] # Vector of vectors
for elem in x, j in eachindex(res)
push!(res, [res[j] ; elem])
end
return res
end
all_subsets([1:5;])
"""
solve_tsp(D, optimizer)
Compute a shortest TSP tour given then matrix distance `D`.
"""
function solve_tsp(D, optimizer)
# Number of cities
N = size(D, 1)
N == size(D, 2) || throw(DimensionMismatch()) # sanity check: `D` is square
N <= 16 || error("N cannot be larger than 16 for memory safety") # sanity check: `N` is not too large
# Instantiate a model
mip = Model(with_optimizer(optimizer))
# I. Create arc variables
@variable(mip, X[1:N, 1:N], Bin)
# II. Set objective
@objective(mip, Min, sum(X.*D)) # sum(D[i,j] * X[i,j] for i=1:N, j=1:N)
# III. Add constraints to the model
# III.1
# No city can be its own follower in the tour
for k in 1:N
@constraint(mip, X[k,k] == 0.0)
end
# III.2
# Each city has one predecessor and one successor
for i in 1:N
@constraint(mip, sum(X[i, j] for j in 1:N) == 1.0)
@constraint(mip, sum(X[j, i] for j in 1:N) == 1.0)
end
# III.3
# We only want a cycle of length N
tours = all_subsets([1:N;])
for st in tours
T = length(st)
# Sub-tour elimination constraints
if 2 ≤ T ≤ N-1
@constraint(mip, sum(X[st[k],st[k+1]] for k=1:T-1) + X[st[T],st[1]] ≤ T-1)
@constraint(mip, sum(X[st[k+1],st[k]] for k=1:T-1) + X[st[1],st[T]] ≤ T-1)
end
end
# Solve MIP model
optimize!(mip)
println("Optimal tour length is ", objective_value(mip))
# Return solution
return value.(X)
end
solve_tsp(d, Cbc.Optimizer)
sol = solve_tsp(d, GLPK.Optimizer)
include("tsp/RoadTrip.jl")
RoadTrip(9, c, sol)
# 
| julia/TSP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # New Angles from Right Range: Optimizing Car Sensor Positioning with D-Wave Hybrid Quantum Computers
# ### BMW Quantum Computing Challenge -- Round 2
# #### <NAME>, [Unified Database Management Systems](https://www2.helsinki.fi/en/researchgroups/unified-database-management-systems-udbms/people#section-63562)
# #### <NAME>, [Discovery Research Group](https://www2.helsinki.fi/en/researchgroups/computational-creativity-and-data-mining/people#section-102417)
# #### Department of Computer Science, University of Helsinki
# ## Introduction to the implementation
# We have been learning, testing and developing some basic quantum algorithms using D-waves own API but we are not yet very experienced with details related to these quantum computers.
#
# The structure of the code is simple: first we import the data, then we create the binary variables as described in the proposal. After that we construct the three objective functions. Finally, we send the total objective functions to D-wave's quantum computer which solves it. The final result is printed.
# ## Initializing parameters
# ### Importing D-wave packages
# In order to run the code here, you need to be able to successfully access D-wave's quantum cloud computing resources. You can see more info at https://cloud.dwavesys.com/. The reason is that, unfortunately, Amazon Bracket does not yet support D-wave's hybrid solvers.
# +
# %%time
import dimod
from dwave.system import LeapHybridSampler
import json
import csv
from itertools import combinations
import pprint
import os
import math
import warnings
warnings.filterwarnings("ignore")
import numpy as np
from matplotlib import pyplot
from mpl_toolkits.mplot3d import Axes3D
from mpl_toolkits import mplot3d
from sympy import *
notebook_path = os.path.abspath("main_3D_connecting_Dwave_Leap.ipynb")
# -
# ### Global parameters
# The parameters `enviroment_x`, `enviroment_y`, `enviroment_z` describe the dimensions of the environment. The environment is the space where the car is located in. Anything outside the environment is supposed not to be accessible by sensors. We took the values for these parameters from the data set `criticallity_grid` which BMW provided.
# +
# %%time
car_sample_accuracy = 500
angle_accuracy = 60
variables = dict()
all_variables = list()
criticallity_grid_abs_path = os.path.join(os.path.dirname(notebook_path), "sensor_position_data/criticallity_grid_0_5.csv")
xs, ys, zs = list(), list(), list()
with open(criticallity_grid_abs_path, newline='') as csvfile:
records = iter(csv.reader(csvfile, delimiter=','))
next(records)
for row in records:
xs.append(int(float(row[0])*100))
ys.append(int(float(row[1])*100))
zs.append(int(float(row[2])*100))
print("Corner points of the environment: ", max(xs), min(xs), max(ys), min(ys), max(zs), min(zs))
environment_x = abs(max(xs)) + abs(min(xs))
environment_y = abs(max(ys)) + abs(min(ys))
environment_z = abs(max(zs)) + abs(min(zs))
print("Environment x-coordinate: ", environment_x)
print("Environment y-coordinate: ", environment_y)
print("Environment z-coordinate: ", environment_z)
# -
# ### Importing sensors
# We created various sensors to demonstrate the code. It is difficult to say if these sensors have realistic values because we didn't have sensor examples in BMW data.
# +
# %%time
abs_sensors_file_path = os.path.join(os.path.dirname(notebook_path), "3d_example_data/3D_sensors_set_1.json")
sensor_types = {0: 'lidar', 1: 'radar', 2: 'camera', 3: 'ultrasound'}
f = open(abs_sensors_file_path)
sensor_root = json.load(f)
sensors = sensor_root["sensors"]
def get_sensor_price(sensor_id):
for sensor in sensors:
if sensor['id'] == sensor_id:
return sensor['price']
print(json.dumps(sensors, indent=4, sort_keys=True))
# -
# ### Initializing allowed sensor positions on car
# Here we utilize the data BMW provided. Compared to the first-round two-dimensional version, the car model in this second-round version is based on the allowed sensor positions data. In order to create variables, we need to sample some points from the surfaces. To sample points, we need some easier format for the surfaces than vector representation. Thus we calculate plane equations for the surfaces. The equation allows us to sample points when we are creating the variables.
# +
def unit_vector(vector):
return vector / np.linalg.norm(vector)
def test_vectors(v1, v2, a, b, c, cp):
if not np.allclose([1,1,0,0], [v1.dot(v1), v2.dot(v2), v1.dot(v2), v1.dot(v2)]):
print("Error in dot products!")
print(v1.dot(v2))
print(v1.dot(v2))
if not np.allclose([1,0,0], [cp.dot(cp), cp.dot(v1), cp.dot(v2)]):
print("Error in normal vector dot products!")
print(cp.dot(v1))
print(cp.dot(v2))
def plot_region(a, b, c, d):
x = np.linspace(-1,1,10)
y = np.linspace(-1,1,10)
X,Y = np.meshgrid(x,y)
if c != 0:
Z = (d - a*X - b*Y) / c
fig = plt.figure()
ax = fig.gca(projection='3d')
surf = ax.plot_surface(X, Y, Z)
def scatter_plot(points, save = False):
fig = pyplot.figure()
ax = Axes3D(fig)
x_vals = [c[0] for c in points]
y_vals = [c[1] for c in points]
z_vals = [c[2] for c in points]
#ax.scatter3D(x_vals, y_vals, z_vals, c=z_vals);
ax.scatter(x_vals, y_vals, z_vals)
#ax.set_xlabel('X Label')
#ax.set_ylabel('Y Label')
#ax.set_zlabel('Z Label')
pyplot.show()
def plane_equation(c1, c2, c3):
p1 = np.array([c1[0], c1[1], c1[2]])
p2 = np.array([c2[0], c2[1], c2[2]])
p3 = np.array([c3[0], c3[1], c3[2]])
# These two vectors are in the plane
v1 = unit_vector(p3 - p1)
v2 = unit_vector(p2 - p1)
# We modify the second vector so that it is orthogonal with the vector v1.
# This is important so that we get 90 degree angle between the vectors and we
# can use trigonometric functions.
v2 -= v1.dot(v2)*v1
# Cross product is a vector normal to the plane
# spanned by vectors v1 and v2
cp = unit_vector(np.cross(v1, v2))
cp = np.array([round(x, 7) for x in cp])
a, b, c = cp
# Testing vectors
# test_vectors(v1, v2, a, b, c, cp)
# This evaluates a * x3 + b * y3 + c * z3 which equals d
d = np.dot(cp, p3)
# print('Equation is {0}x + {1}y + {2}z = {3}'.format(a, b, c, d))
# plot_region(a, b, c, d)
return { 'span1': v1, 'span2': v2, 'normal_vector': cp, 'x': a, 'y': b, 'z': c, 'd': d }
# -
# In the following, we calculate the planes for the different surfaces of the car. Besides, we characterize each plane with its plane equation (or line).
# +
# %%time
abs_position_file_path = os.path.join(os.path.dirname(notebook_path), "sensor_position_data/allowed_sensor_positions.csv")
allowed_sensor_positions = dict()
with open(abs_position_file_path, newline='') as csvfile:
records = iter(csv.DictReader(csvfile, delimiter=';'))
next(records)
for row in records:
c1 = (int(row["x1"]), int(row["y1"]), int(row["z1"]))
c2 = (int(row["x2"]), int(row["y2"]), int(row["z2"]))
c3 = (int(row["x3"]), int(row["y3"]), int(row["z3"]))
c4 = (int(row["x4"]), int(row["y4"]), int(row["z4"]))
region = row["Region"].lower()
allowed_sensor_positions[region] = dict()
allowed_sensor_positions[region]["corners"] = [c1, c2, c3, c4]
allowed_sensors = [x.lower() for x in row["Allowed Sensors"].split(", ")]
allowed_sensor_positions[region]["allowed_sensors"] = allowed_sensors
equation = plane_equation(c1, c2, c3)
allowed_sensor_positions[region]["equation"] = equation
intervals = dict()
if equation["x"] == 0:
if c1[0] < c2[0]:
intervals["x"] = range(c1[0], c2[0], car_sample_accuracy)
elif c1[0] > c2[0]:
intervals["x"] = range(c2[0], c1[0], car_sample_accuracy)
elif c1[0] < c3[0]:
intervals["x"] = range(c1[0], c3[0], car_sample_accuracy)
elif c1[0] > c3[0]:
intervals["x"] = range(c3[0], c1[0], car_sample_accuracy)
if equation["y"] == 0:
if c1[1] < c2[1]:
intervals["y"] = range(c1[1], c2[1], car_sample_accuracy)
elif c1[1] > c2[1]:
intervals["y"] = range(c2[1], c1[1], car_sample_accuracy)
elif c1[1] < c3[1]:
intervals["y"] = range(c1[1], c3[1], car_sample_accuracy)
elif c1[1] > c3[1]:
intervals["y"] = range(c3[1], c1[1], car_sample_accuracy)
if equation["z"] == 0:
if c1[2] < c2[2]:
intervals["z"] = range(c1[2], c2[2], car_sample_accuracy)
elif c1[2] > c2[2]:
intervals["z"] = range(c2[2], c1[2], car_sample_accuracy)
elif c1[2] < c3[2]:
intervals["z"] = range(c1[2], c3[2], car_sample_accuracy)
elif c1[2] > c3[2]:
intervals["z"] = range(c3[2], c1[2], car_sample_accuracy)
allowed_sensor_positions[region]["fixed_intervals"] = intervals
#for_printing = allowed_sensor_positions[elem]
#pp = pprint.PrettyPrinter(width=41, compact=True)
for elem in allowed_sensor_positions:
print(elem, allowed_sensor_positions[elem])
print()
# -
# ### Initializing variables
# Compared to the first-round, variables are updated to be triples `(x, y, i)` where `x` and `y` are points in the 3D-space and `i` refers to the sensor's id. The point `x` belongs to some of the allowed sensor positions on the car's surface. For each `x`, points `y` are sampled from the environment so that the distance between `x` and `y` is range `R_i` for sensor `i`.
# We are dealing with different cases depending on how the car surface is positioned in the space. A bit simplifying we can say that one of the varibles, say `x`, runs over an interval. The values for the two other variables, `y` and `z`, we get from the plane equation when we substitute the value `x`. Now these triples belong to the car's surface.
# +
def sample_from_car_surface(corners, equation, fixed_intervals, car_sample_accuracy):
sample = list()
# This is the simple case that the plane is parallel with some of the three axis.
# Thus the parallel axis stays constant.
# For example, side mirror, back, trunk and sides are parallel to one of the axis
if len(fixed_intervals) == 2:
if 'x' in fixed_intervals and 'y' in fixed_intervals:
z = corners[0][2]
for x in fixed_intervals['x']:
for y in fixed_intervals['y']:
sample.append((x, y, z))
elif 'x' in fixed_intervals and 'z' in fixed_intervals:
y = corners[0][1]
for x in fixed_intervals['x']:
for z in fixed_intervals['z']:
sample.append((x, y, z))
elif 'y' in fixed_intervals and 'z' in fixed_intervals:
x = corners[0][0]
for y in fixed_intervals['y']:
for z in fixed_intervals['z']:
sample.append((x, y, z))
elif len(fixed_intervals) == 1:
if 'x' in fixed_intervals:
y, z = symbols('y z')
expr = equation['y']*y + equation['z']*z - equation['d']
y_interval = None
c1 = corners[0][1]
c2 = corners[1][1]
c3 = corners[2][1]
if c1 < c2:
y_interval = range(c1, c2, car_sample_accuracy)
elif c1 > c2:
y_interval = range(c2, c1, car_sample_accuracy)
elif c1 < c3:
y_interval = range(c1, c3, car_sample_accuracy)
elif c1 > c3:
y_interval = range(c3, c1, car_sample_accuracy)
for x in fixed_intervals['x']:
for y_var in y_interval:
y_expr = expr.subs(y, y_var)
z = math.floor(solve(y_expr)[0])
sample.append((x, y_var, z))
#print((x, y_var, z))
x += car_sample_accuracy
elif 'y' in fixed_intervals:
x, z = symbols('x z')
expr = equation['x']*x + equation['z']*z - equation['d']
x_interval = None
c1 = corners[0][0]
c2 = corners[1][0]
c3 = corners[2][0]
if c1 < c2:
x_interval = range(c1, c2, car_sample_accuracy)
elif c1 > c2:
x_interval = range(c2, c1, car_sample_accuracy)
elif c1 < c3:
x_interval = range(c1, c3, car_sample_accuracy)
elif c1 > c3:
x_interval = range(c3, c1, car_sample_accuracy)
for y in fixed_intervals['y']:
for x_var in x_interval:
x_expr = expr.subs(x, x_var)
z = math.floor(solve(x_expr)[0])
sample.append((x_var, y, z))
#print((x_var, y, z))
y += car_sample_accuracy
elif 'z' in fixed_intervals:
x, y = symbols('x y')
expr = equation['x']*x + equation['y']*y - equation['d']
x_interval = None
c1 = corners[0][0]
c2 = corners[1][0]
c3 = corners[2][0]
if c1 < c2:
x_interval = range(c1, c2, car_sample_accuracy)
elif c1 > c2:
x_interval = range(c2, c1, car_sample_accuracy)
elif c1 < c3:
x_interval = range(c1, c3, car_sample_accuracy)
elif c1 > c3:
x_interval = range(c3, c1, car_sample_accuracy)
for z in fixed_intervals['z']:
for x_var in x_interval:
x_expr = expr.subs(x, x_var)
y = math.floor(solve(x_expr)[0])
#print(y)
sample.append((x_var, y, z))
#print((x_var, y, z))
z += car_sample_accuracy
return sample
# -
# Works in terminal Python but not in the notebook version
def plot_sampled_points_from_car(all_variables):
fig = pyplot.figure()
ax = fig.add_subplot(111, projection='3d')
xs, ys, zs = list(), list(), list()
for var in all_variables:
point = var[1]
xs.append(point[0])
ys.append(point[1])
zs.append(point[2])
print(xs)
print()
print(ys)
print()
print(zs)
#ax.scatter(xs, ys, zs)
#ax.set_xlabel('X Label')
#ax.set_ylabel('Y Label')
#ax.set_zlabel('Z Label')
#pyplot.show()
# After we have sampled points from the car's surface, we sample points from the environment. We fix a point `x` on the car's surface and then pick a point `y` from the environment so that the distance between `x` and `y` is the range `R_i`. We let the angle between the car's surface and the vector `x - y` run over different values which produces multiple possible point pairs `(x, y)`. In the optimal situation we could sample points with some sufficiently small accuracy factor.
# +
# %%time
for sensor in sensors:
variables[sensor["id"]] = dict()
for pos_name in allowed_sensor_positions:
pos = allowed_sensor_positions[pos_name]
#print(sensor_types)
if sensor_types[sensor["type"]] in pos["allowed_sensors"]:
variables[sensor["id"]][pos_name] = list()
srange = sensor["view"]["range"]
corners = pos["corners"]
equation = pos["equation"]
fixed_intervals = pos["fixed_intervals"]
normal_vector = -equation['normal_vector']
spanning_vector_on_plane1 = equation['span1']
spanning_vector_on_plane2 = equation['span2']
car_sample = sample_from_car_surface(corners, equation, fixed_intervals, car_sample_accuracy)
#print(pos_name)
#print(car_sample)
#scatter_plot(car_sample)
for car_point in car_sample:
for angle_on_plane in range(0, 360, angle_accuracy):
for angle_for_normal in range(0, 90, angle_accuracy):
rad_angle_on_plane = math.radians(angle_on_plane)
rad_angle_for_normal = math.radians(angle_for_normal)
#print("Car point ", car_point)
#print("Normal vector", normal_vector)
#print("Spanning vector", spanning_vector_on_plane1)
#print("Spanning vector", spanning_vector_on_plane2)
# We start moving from the fixed point on the car's surface
point_in_environment = np.array([float(x) for x in car_point])
car_point_vector = np.array(car_point)
#print(rad_angle_on_plane)
#print(math.cos(rad_angle_on_plane))
#print("#11 ", srange*math.cos(rad_angle_on_plane)*spanning_vector_on_plane1)
#print("#12 ", srange*math.sin(rad_angle_on_plane)*spanning_vector_on_plane2)
#print("#13 ", srange*math.sin(rad_angle_for_normal)*normal_vector)
# Move along the plane to the direction of the first spanning vector
point_in_environment += srange*math.cos(rad_angle_on_plane)*math.cos(rad_angle_for_normal)*spanning_vector_on_plane1
#print("#1 ", point_in_environment)
# Move along the plane to the direction of the second spanning vector
point_in_environment += srange*math.sin(rad_angle_on_plane)*math.cos(rad_angle_for_normal)*spanning_vector_on_plane2
#print("#2 ", point_in_environment)
# Move to the orthonogal direction to the plane i.e. "upwards"
point_in_environment += srange*math.sin(rad_angle_for_normal)*normal_vector
#print("#3 ", point_in_environment)
#env_points.append(point_in_environment)
#scatter_plot(env_points)
#print("Distance defined in the sensor data is " + str(srange) + " and the distance between the sampled points is ", np.linalg.norm(car_point_vector.dot(point_in_environment)))
point_in_environment = [math.floor(x) for x in point_in_environment]
# For bugging purposes:
#print("Angles are ", angle_on_plane, angle_for_normal)
b_variable = (car_point, tuple(point_in_environment), sensor["id"])
variables[sensor["id"]][pos_name].append(b_variable)
all_variables.append(b_variable)
print("Number of variables: ", len(all_variables))
# -
# ## Constructing quadratic unconstrained binary optimization model
vartype = dimod.BINARY
main_bqm = dimod.BinaryQuadraticModel({}, {}, 0.0, vartype)
# The following functions enable us to append or initialize coefficients for the variables in the BQM. The distance function implements the ordinary Euclidean distance.
# +
def append_linear_safe(variable, value, linear_dict):
if variable in linear_dict.keys():
linear_dict[variable] = linear_dict[variable] + value
else:
linear_dict[variable] = value
def append_quadratic_safe(variable, value, quadratic_dict):
if variable in quadratic_dict.keys():
quadratic_dict[variable] = quadratic_dict[variable] + value
else:
quadratic_dict[variable] = value
def sensor_view_volume(sensor_id):
for sensor in sensors:
if sensor['id'] == sensor_id:
# Following code calculates volume of view of sensor
side1 = 2*sensor["view"]["range"]*float(math.tan(math.radians(sensor["view"]["angle"]["horizontal"])/2))
side2 = 2*sensor["view"]["range"]*float(math.tan(math.radians(sensor["view"]["angle"]["vertical"])/2))
return (1/3)*sensor["view"]["range"]*side1*side2
def print_current_qubo(number_of_linear_terms = 100, number_of_quadratic_terms = 100):
i = 0
for elem in main_bqm.linear:
print(elem, main_bqm.linear[elem])
i+=1
if i > number_of_linear_terms:
break
i = 0
for elem in main_bqm.quadratic:
print(elem, main_bqm.quadratic[elem])
i+=1
if i > number_of_quadratic_terms:
break
# -
# ### Constraint 1: selecting sufficiently sensors to cover the environment
# Every binary quadratic function which is part of the model contains four parameters: linear terms, quadratic terms, offset (constant) and variable type. Variable type is always BINARY since we are using QUBO. If we use Ising, we set variable type to be SPIN.
# +
# %%time
# Encoding constraint H1
E = environment_x*environment_y*environment_z
print("Total volume of the environment: ", E)
A1 = 1/math.pow(10, 12)
linear_h1 = {}
quadratic_h1 = {}
offset_h1 = float(math.pow(E, 2))
#print(offset_h1)
# Linear terms
for sensor_id in variables:
volume = sensor_view_volume(sensor_id)
print("Volume of sensor " + str(sensor_id) + ": ", volume)
coefficient = float(math.pow(volume, 2) - 2*E*volume)
for surface in variables[sensor_id]:
for var in variables[sensor_id][surface]:
append_linear_safe(var, coefficient, linear_h1)
# Quadratic terms
quadratic_terms = combinations(all_variables, 2)
for pair in quadratic_terms:
b1 = pair[0]
b2 = pair[1]
volume1 = sensor_view_volume(b1[2])
volume2 = sensor_view_volume(b2[2])
coefficient_quadratic = float(2*volume1*volume2)
append_quadratic_safe((b1, b2), coefficient_quadratic, quadratic_h1)
bqm_h1 = dimod.BinaryQuadraticModel(linear_h1, quadratic_h1, offset_h1, vartype)
bqm_h1.scale(A1)
main_bqm.update(bqm_h1)
# -
# The following code is for printing the linear and quadratic terms.
# +
#print_current_qubo(10, 10)
# -
# ### Constraint 2: optimizing overlap of sensor views
# One of the biggest problems in the current code is the following overlap function. It is just a rough estimate on how much two sensor views overlap.
# +
def sensor_inside_sensor(car_p1, env_p1, car_p2, env_p2):
if car_p2.all() == env_p2.all():
return True
if car_p1.all() == car_p2.all():
if (car_p2 - env_p2).all() != 0:
line = (car_p1 - env_p1)/(car_p2 - env_p2)
for i in range(len(line) - 1):
if line[i] != line[i+1]:
return False
else:
return True
return False
def overlap(b1, b2):
car_p1 = np.array([int(x) for x in b1[0]])
env_p1 = np.array([int(x) for x in b1[1]])
id1 = int(b1[2])
car_p2 = np.array([int(x) for x in b2[0]])
env_p2 = np.array([int(x) for x in b2[1]])
id2 = int(b2[2])
if sensor_inside_sensor(car_p1, env_p1, car_p2, env_p2):
return 1
cone_axis1 = env_p1 - car_p1
cone_axis2 = env_p2 - car_p2
# Angle between the axis
u_cone_axis1 = cone_axis1 / np.linalg.norm(cone_axis1)
u_cone_axis2 = cone_axis2 / np.linalg.norm(cone_axis2)
axis_angle = np.arccos(np.dot(u_cone_axis1, u_cone_axis2))
#print(math.degrees(axis_angle))
# Distance between the middle points of the vectors cone_axis1 and cone_axis2
mid_cone_axis1 = (np.linalg.norm(cone_axis1)/2)*u_cone_axis1
mid_cone_axis2 = (np.linalg.norm(cone_axis2)/2)*u_cone_axis2
mid_point_dist = np.linalg.norm(mid_cone_axis1 - mid_cone_axis2)
top_point_dist = np.linalg.norm(env_p1 - env_p2)
#print(mid_point_dist)
sensor1, sensor2 = None, None
for sensor in sensors:
if sensor['id'] == id1:
sensor1 = sensor
if sensor['id'] == id2:
sensor2 = sensor
# This part of the code is very heuristical and possibly works badly and ruins everything
side1 = np.linalg.norm(cone_axis1)
side2 = np.linalg.norm(cone_axis2)
top_half1_dist1 = math.tan(math.radians(sensor1["view"]["angle"]["horizontal"])/2)*side1
top_half1_dist2 = math.tan(math.radians(sensor1["view"]["angle"]["vertical"])/2)*side1
top_half2_dist1 = math.tan(math.radians(sensor2["view"]["angle"]["horizontal"])/2)*side2
top_half2_dist2 = math.tan(math.radians(sensor2["view"]["angle"]["vertical"])/2)*side2
#print("Distances on top: ", top_half1_dist1, top_half1_dist2, top_half2_dist1, top_half2_dist2)
mid_half1_dist1 = top_half1_dist1/2
mid_half1_dist2 = top_half1_dist2/2
mid_half2_dist1 = top_half2_dist1/2
mid_half2_dist2 = top_half2_dist2/2
#print("Distances in middle: ", mid_half1_dist1, mid_half1_dist2, mid_half2_dist1, mid_half2_dist2)
difference_top1 = top_half1_dist1 + top_half2_dist1 - top_point_dist
difference_top2 = top_half1_dist2 + top_half2_dist2 - top_point_dist
difference_mid1 = mid_half1_dist1 + mid_half2_dist1 - mid_point_dist
difference_mid2 = mid_half1_dist2 + mid_half2_dist2 - mid_point_dist
#print("Top differences: ", difference_top1, difference_top2)
#print("Middle differences: ", difference_mid1, difference_mid2)
top_divisor1 = max([top_half1_dist1, top_half2_dist1])
top_divisor2 = max([top_half1_dist2, top_half2_dist2])
top_divisor = top_divisor1 + top_divisor2
mid_divisor1 = max([mid_half1_dist1, mid_half2_dist1])
mid_divisor2 = max([mid_half1_dist2, mid_half2_dist2])
mid_divisor = mid_divisor1 + mid_divisor2
top_sum = 0
mid_sum = 0
if difference_top1 > 0:
top_sum += difference_top1
if difference_top2 > 0:
top_sum += difference_top2
if difference_mid1 > 0:
mid_sum += difference_mid1
if difference_mid2 > 0:
mid_sum += difference_mid2
overlap_result = (top_sum + mid_sum)/(top_divisor + mid_divisor)
#print("Final result: ", overlap_result)
if overlap_result > 1:
return 1/overlap_result
return overlap_result
# +
# %%time
# Encoding constraint H2
A2 = 1000
linear_h2 = {}
quadratic_h2 = {}
offset_h2 = 0
quadratic_terms = combinations(all_variables, 2)
for pair in quadratic_terms:
b1 = pair[0]
b2 = pair[1]
coefficient_quadratic = overlap(b1, b2)
append_quadratic_safe((b1, b2), coefficient_quadratic, quadratic_h2)
bqm_h2 = dimod.BinaryQuadraticModel(linear_h2, quadratic_h2, offset_h2, vartype)
bqm_h2.scale(A2)
main_bqm.update(bqm_h2)
# +
#print_current_qubo(50, 50)
# -
# ### Constraint 3: minimizing total price
# +
# %%time
# Encoding constraint H3
A3 = 1
linear_h3 = {}
quadratic_h3 = {}
offset_h3 = 0
for variable in all_variables:
sensor_id = variable[2]
price = get_sensor_price(sensor_id)
append_linear_safe(variable, price, linear_h3)
bqm_h3 = dimod.BinaryQuadraticModel(linear_h3, quadratic_h3, offset_h3, vartype)
bqm_h3.scale(A3)
main_bqm.update(bqm_h3)
# +
#print_current_qubo(50, 50)
# -
# ### Solve QUBO
# Unfortunately, LeapHybridSampler is not available in Amazon Bracket. That is why this code will not work in Bracket. On the other hand, we tried run the code using BracketDwaveSampler but the problem cannot be mapped to the circuits.
# +
# %%time
main_bqm.normalize()
sampler = LeapHybridSampler()
sampleset = sampler.sample(main_bqm)
sample = sampleset.first.sample
print(sampleset)
print()
# energy = sampleset.first.energy
print("Possible sensor positions in the space (point on car, point in environment, sensor id):")
i = 0
for varname, value in sample.items():
if value == 1:
i+=1
print(varname, value)
print(i)
| sensor_bmw/main_3D_connecting_Dwave_Leap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Doyle-Fuller-Newman Model (DFN)
# ## Dimensionless Model Equations
# The DFN comprises equations for charge and mass conservation in the solid the solid and electrolyte, and also prescribes behaviour for the electrochemical reactions occurring on the interface between the solid an electrolyte. For more information please see [[2]](#References) or other standard texts.
#
# Below we summarise the dimensionless form of the DFN, with all parameters give in the table at the end of this notebook. Here we use a roman subscript $\text{k} \in \text{n, s, p}$ is used to denote the regions negative electrode, separator, and positive electrode, respectively.
#
# The model equations for the DFN read:
#
# #### Charge conservation:
# \begin{align}
# \frac{\partial i_{\text{e,k}}}{\partial x} &= \begin{cases}
# j_{\text{k}}, \quad &\text{k} = \text{n, p}\\
# 0, \qquad &\text{k} = \text{s}
# \end{cases} , && \\
# \mathcal{C}_{\text{e}} i_{\text{e,k}} &= \epsilon_{\text{k}}^{\text{b}} \gamma_{\text{e}} \kappa_{\text{e}}(c_{\text{e,k}}) \left( - \frac{\partial \phi_{\text{e,k}}}{\partial x} + 2(1-t^+)\frac{\partial}{\partial x}\left(\log(c_{\text{e,k}})\right)\right), && \text{k} \in \text{n, s, p}, \\
# I-i_{\text{e,k}} &= - \sigma_{\text{k}} \frac{\partial \phi_{\text{e,k}}}{\partial x}, && \text{k} \in \text{n, s, p}.
# \end{align}
#
# #### Mass conservation:
# \begin{align}
# \mathcal{C}_{\text{e}} \epsilon_{\text{k}} \gamma_{\text{e}} \frac{\partial c_{\text{e,k}}}{\partial t} &= -\gamma_{\text{e}} \frac{\partial N_{\text{e,k}}}{\partial x} + \mathcal{C}_{\text{e}} \frac{\partial i_{\text{e,k}}}{\partial x}, && \text{k} \in \text{n, s, p},\\
# N_{\text{e,k}} &= -\epsilon_{\text{k}}^{\text{b}} D_{\text{e}}(c_{\text{e,k}}) \frac{\partial c_{\text{e,k}}}{\partial x} + \frac{\mathcal{C}_{\text{e}} t^+}{\gamma_{\text{e}}} i_{\text{e,k}},
# && \text{k} \in \text{n, s, p}, \\
# \mathcal{C}_{\text{k}} \frac{\partial c_{\text{s,k}}}{\partial t} &= -\frac{1}{r_{\text{k}}^2} \frac{\partial}{\partial r_{\text{k}}} \left(r_{\text{k}}^2 N_{\text{s,k}}\right), && \text{k} \in \text{n, p},\\
# N_{\text{s,k}} &= -D_{\text{s,k}}(c_{\text{s,k}}) \frac{\partial c_{\text{s,k}}}{\partial r_{\text{k}}}, && \text{k} \in \text{n, p}.
# \end{align}
#
#
# #### Electrochemical reactions:
# \begin{align}
# j_{\text{k}} &= 2 j_{\text{0,k}} \sinh\left(\frac{ \eta_{\text{k}}}{2} \right), && \text{k} \in \text{n, p}, \\
# j_{\text{0,k}} &= \frac{\gamma_{\text{k}}}{\mathcal{C}_{\text{r,k}}} c_{\text{s,k}}^{1/2} (1-c_{\text{s,k}})^{1/2}c_{\text{e,k}}^{1/2}\big|_{r_{\text{k}}=1}, && \text{k} \in \text{n, p}, \\
# \eta_{\text{k}} &= \phi_{\text{s,k}} - \phi_{\text{e,k}} - U_{\text{k}}(c_{\text{s,k}}\big|_{r_{\text{k}}=1}), && \text{k} \in \text{n, p}.
# \end{align}
#
#
# These are to be solved subject to the following boundary conditions:
#
# #### Current:
# \begin{gather}
# i_{\text{e,n}}\big|_{x=0} = 0, \quad i_{\text{e,p}}\big|_{x=1}=0, \\
# \phi_{\text{e,n}}\big|_{x=L_{\text{n}}} = \phi_{\text{e,s}}\big|_{x=L_{\text{n}}}, \quad i_{\text{e,n}}\big|_{x=L_{\text{n}}} = i_{\text{e,s}}\big\vert_{x=L_{\text{n}}} = I, \\
# \phi_{\text{e,s}}\big|_{x=1-L_{\text{p}}} = \phi_{\text{e,p}}\big|_{x=1-L_{\text{p}}}, \quad
# i_{\text{e,s}}\big|_{x=1-L_{\text{p}}} = i_{\text{e,p}}\big|_{x=1-L_{\text{p}}} = I.
# \end{gather}
#
# #### Concentration in the electrolyte:
# \begin{gather}
# N_{\text{e,n}}\big|_{x=0} = 0, \quad N_{\text{e,p}}\big|_{x=1}=0,\\
# c_{\text{e,n}}\big|_{x=L_{\text{n}}} = c_{\text{e,s}}|_{x=L_{\text{n}}}, \quad N_{\text{e,n}}\big|_{x=L_{\text{n}}}=N_{\text{e,s}}\big|_{x=L_{\text{n}}}, \\
# c_{\text{e,s}}|_{x=1-L_{\text{p}}}=c_{\text{e,p}}|_{x=1-L_{\text{p}}}, \quad N_{\text{e,s}}\big|_{x=1-L_{\text{p}}}=N_{\text{e,p}}\big|_{x=1-L_{\text{p}}}. &&
# \end{gather}
#
# #### Concentration in the electrode active material:
# \begin{gather}
# N_{\text{s,k}}\big|_{r_{\text{k}}=0} = 0, \quad \text{k} \in \text{n, p}, \quad \ \ - \frac{a_{R, \text{k}}\gamma_{\text{k}}}{\mathcal{C}_{\text{k}}} N_{\text{s,k}}\big|_{r_{\text{k}}=1} = j_{\text{k}}, \quad \text{k} \in \text{n, p}.
# \end{gather}
#
# #### Reference potential:
# $$\phi_{\text{s,cn}} = 0, \quad \boldsymbol{x} \in \partial \Omega_{\text{tab,n}}.$$
#
#
# And the initial conditions:
#
# \begin{align}
# &c_{\text{s,k}}(x,r,0) = c_{\text{s,k,0}}, \quad \phi_{\text{s,n}}(x,0) = 0, \quad \phi_{\text{s,p}}(x,0) = \phi_{\text{s,p,0}}, && \text{k} \in \text{n, p},\\
# &\phi_{\text{e,k}}(x,0) = \phi_{\text{e,0}}, \quad c_{\text{e,k}}(x,0) = 1, && \text{k} \in \text{n, s, p}.
# \end{align}
#
# ## Example solving DFN using PyBaMM
# Below we show how to solve the DFN model, using the default geometry, mesh, parameters, discretisation and solver provided with PyBaMM. For a more detailed example, see the notebook on the [SPM](https://github.com/pybamm-team/PyBaMM/blob/develop/examples/notebooks/models/SPM.ipynb).
#
# In order to show off all the different points at which the process of setting up and solving a model in PyBaMM can be customised we explicitly handle the stages of choosing a geometry, setting parameters, discretising the model and solving the model. However, it is often simpler in practice to use the `Simulation` class, which handles many of the stages automatically, as shown [here](../simulation-class.ipynb).
#
# First we need to import pybamm, along with numpy which we will use in this notebook.
# %pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import numpy as np
# We then load the DFN model and default geometry, and process them both using the default parameters.
# +
# load model
model = pybamm.lithium_ion.DFN()
# create geometry
geometry = model.default_geometry
# load parameter values and process model and geometry
param = model.default_parameter_values
param.process_model(model)
param.process_geometry(geometry)
# -
# The next step is to set the mesh and discretise the model. Again, we choose the default settings.
# +
# set mesh
mesh = pybamm.Mesh(geometry, model.default_submesh_types, model.default_var_pts)
# discretise model
disc = pybamm.Discretisation(mesh, model.default_spatial_methods)
disc.process_model(model);
# -
# The model is now ready to be solved. We select the default DAE solver for the DFN. Note that in order to successfully solve the system of DAEs we are required to give consistent initial conditions. This is handled automatically by PyBaMM during the solve operation.
# solve model
solver = model.default_solver
t_eval = np.linspace(0, 3600, 300) # time in seconds
solution = solver.solve(model, t_eval)
# To get a quick overview of the model outputs we can use the QuickPlot class, which plots a common set of useful outputs. The method `Quickplot.dynamic_plot` makes a slider widget.
quick_plot = pybamm.QuickPlot(solution, ["Positive electrode interfacial current density [A.m-2]"])
quick_plot.dynamic_plot();
# ## Dimensionless Parameters
# In the table below, we provide the dimensionless parameters in the DFN in terms of the dimensional parameters in mcmb2528_lif6-in-ecdmc_lico2_parameters_Dualfoil.csv. We use a superscript * to indicate dimensional quantities.
#
# | Parameter | Expression |Interpretation |
# |:--------------------------|:----------------------------------------|:------------------------------------------|
# | $L_{\text{k}}$ | $L_{\text{k}}^*/L^*$ | Ratio of region thickness to cell thickness|
# |$\sigma_{\text{k}}$ | $\sigma_{\text{k}}^*R^* T^*/(I^*F^*L^*)$| Dimensionless solid conductivity |
# |$\mathcal{C}_{\text{k}}$ | $\tau_{\text{k}}^*/\tau_{\text{d}}^*$ | Ratio of solid diffusion and discharge timescales |
# |$\mathcal{C}_{\text{e}}$ |$\tau_{\text{e}}^*/\tau_{\text{d}}^*$ |Ratio of electrolyte transport and discharge timescales|
# |$\mathcal{C}_{\text{r,k}}$ |$\tau_{\text{r,k}}^*/\tau_{\text{d}}^*$ |Ratio of reaction and discharge timescales|
# |$a_{R, \text{k}}$ |$a_{\text{k}}^* R_{\text{k}}^*$ | Product of particle radius and surface area to volume ratio|
# |$\gamma_{\text{k}}$ |$c_{\text{k,max}}^*/c_{\text{n,max}}^*$ |Ratio of maximum lithium concentrations in solid|
# |$\gamma_{\text{e}}$ |$c_{\text{e,typ}}^*/c_{\text{n,max}}^*$ |Ratio of maximum lithium concentration in the negative electrode solid and typical electrolyte concentration|
#
# Note that the dimensionless parameters $\epsilon_{\text{k}}$, $\text{b}$, and $t^+$ are already provided in the parameter file mcmb2528_lif6-in-ecdmc_lico2_parameters_Dualfoil.csv
# ## References
#
# The relevant papers for this notebook are:
pybamm.print_citations()
| examples/notebooks/models/DFN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CAMPUS PLACEMENT ANALYSIS
# ## Importing Project Dependencies
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
import os
# -
# ## Importing our data set
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
df = pd.read_csv('Placement_Data_Full_Class.csv')
# -
df.head()
# Now that we have our data, we will try and look for null values in our data. We will use the describe() method to find the mean of different columns and some other data that might help us in making our model.
df.isnull().sum()
# We only have null values in salary column, for now we will keep it and try and see whhat whether we need it as a feature or not. If needed, we will fill these null values.
df.info()
df.describe()
# We will try and visualise our data in order to find the right features to train our models on. We have a lot of columns, so let's try and compare with the variable that we need to predict i.e the status, whether a candidate is placed or not.
plt.style.use('dark_background')
plt.figure(figsize=(10,5))
sns.countplot('gender',data = df,palette = 'inferno')
plt.title("Distribution of Males and Females in our Data",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('ssc_b',data = df,palette = 'inferno')
plt.title("Distribution of the Boards the Students belong to in 10th",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('hsc_b',data = df,palette = 'inferno')
plt.title("Distribution of the Boards the Students belong to in 10th",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('hsc_s',data = df,palette = 'inferno')
plt.title("Distribution of the Streams that students chose in High school",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('degree_t',data = df,palette = 'inferno')
plt.title("Distribution of the Type of Degrees",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('workex',data = df,palette = 'inferno')
plt.title("Distribution of how many students have prior work experience",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('specialisation',data = df,palette = 'inferno')
plt.title("Distribution of the Types of Specialisation",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('status',data = df,palette = 'inferno')
plt.title("Distribution of the Placements",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('gender',data = df,palette = 'inferno',hue = 'status')
plt.title("Distribution of Placements in Males and Females",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('workex',data = df,palette = 'inferno',hue = 'status')
plt.title("Distribution of Placements in Males and Females",fontsize = 15)
plt.figure(figsize=(10,5))
sns.boxplot('ssc_b','ssc_p',data = df,palette = 'inferno')
plt.title("Relation between the Student's Boards and their score during their Secondary education",fontsize = 15)
plt.figure(figsize=(10,5))
sns.boxplot('status','ssc_p',data = df,palette = 'inferno')
plt.title("Relation between the Students that were placed and their score during their Secondary education",fontsize = 15)
plt.figure(figsize=(10,5))
sns.boxplot('hsc_b','hsc_p',data = df,palette = 'inferno')
plt.title("Relation between the Student's Boards and their score in High school",fontsize = 15)
plt.figure(figsize=(10,5))
sns.boxplot('status','hsc_p',data = df,palette = 'inferno')
plt.title("Relation between the Students that were placed and their score in High school",fontsize = 15)
plt.figure(figsize=(10,5))
sns.boxplot('status','mba_p',data = df,palette = 'inferno')
plt.title("Relation between the Students that were placed and their score during MBA",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('ssc_b',data = df,palette = 'inferno',hue = 'status')
plt.title("Relation between the Students that were placed and the boards that they were in during Secondary Education",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('hsc_b',data = df,palette = 'inferno',hue = 'status')
plt.title("Relation between the Students that were placed and the boards that they were in High School",fontsize = 15)
plt.figure(figsize=(10,5))
sns.boxplot('status','degree_p',data = df,palette = 'inferno')
plt.title("Relation between the Students that were placed and their degree percentage",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('hsc_s',data = df,palette = 'inferno',hue = 'status')
plt.title("Relation between the streams that students chose in highschool and their placement",fontsize = 15)
plt.figure(figsize=(10,5))
sns.countplot('degree_t',data = df,palette = 'inferno',hue = 'status')
plt.title("Relation between the degree types that students chose and their placement",fontsize = 15)
# +
#df.head()
# -
# Now that we know what each column looks like, we can start to select our features that we will feed our model. We will make a copy of the original data frame in order to be safe if anything goes wrong.
df1 = df.copy()
# We will drop the sl_no and salary column as they wont help us in any way. The salary also has null values so dropping it means we dint have to impute these values and it won't help us in predicting if a person gets placed or not.
df1.drop(['sl_no','salary'],axis = 1,inplace = True)
df1.head()
# We have values in the columns that are strings. In order to train our model, we need to convert these strings to numbers in order to feed it to the model.
df1['status']= df1['status'].map({'Placed':1,'Not Placed':0})
df1['workex']= df1['workex'].map({'Yes':1,'No':0})
df1['gender']= df1['gender'].map({'M':1,'F':0})
df1['hsc_b']= df1['hsc_b'].map({'Central':1,'Others':0})
df1['ssc_b']= df1['ssc_b'].map({'Central':1,'Others':0})
df1['degree_t']= df1['degree_t'].map({'Sci&Tech':0,'Comm&Mgmt':1,'Others':2})
df1['specialisation'] = df1['specialisation'].map({'Mkt&HR':1, 'Mkt&Fin':0})
df1['hsc_s']= df1['hsc_s'].map({'Commerce':0,'Science':1,'Arts':2})
df1.head()
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report, confusion_matrix ,accuracy_score,recall_score,precision_score
# We won't use all of these values. We will use the 'ssc_p','hsc_p','degree_p','workex','mba_p','etest_p','gender','degree_t' and 'specialisation' column as our features to predict whether a candidate will be placed or not.
X = df1[['ssc_p','hsc_p','degree_p','workex','mba_p','etest_p','gender','degree_t','specialisation']]
y = df1['status']
# Now that we have our features, we will split them and into training and testing sets and we will also scale down our features in order to have better results.
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size = 0.2,random_state = 101)
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# Our goal is to predict whether a candidate will be placed or not. This is a classification problem so we will use Logistic regression, KNN and SVM and compare how these perform individually.
# ## Logistic Regression
lr = LogisticRegression()
lr.fit(X_train,y_train)
pred = lr.predict(X_test)
print("Accuracy:",accuracy_score(y_test, pred)*100)
print("Precision:",precision_score(y_test, pred)*100)
print("Recall:",recall_score(y_test, pred)*100)
# Saving the model as an .pkl file
import pickle
pickle_out = open("LRModel.pkl", "wb")
pickle.dump(lr, pickle_out)
pickle_out.close()
user_input1 = [[67.00,91.00,58.00,0,58.80,55.00,1,0,1]]
user_pred1 = lr.predict(user_input1)
if user_pred1 == 1:
print("The Candidate will be Placed!")
else:
print("The Candidate won't be Placed :(")
# ## KNN
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train,y_train)
prediction = knn.predict(X_test)
print("Accuracy:",accuracy_score(y_test, prediction)*100)
print("Precision:",precision_score(y_test, prediction)*100)
print("Recall:",recall_score(y_test, prediction)*100)
user_input2 = [[67.00,91.00,58.00,0,58.80,55.00,1,0,1]]
user_pred2 = knn.predict(user_input2)
if user_pred1 == 1:
print("The Candidate will be Placed!")
else:
print("The Candidate won't be Placed :(")
# ## Optimizing KNN
error = []
for i in range(1,100):
knn = KNeighborsClassifier(n_neighbors= i)
knn.fit(X_train,y_train)
pred_i = knn.predict(X_test)
error.append(np.mean(pred_i != y_test))
plt.figure(figsize = (10,6))
plt.plot(range(1,100),error)
plt.title('K-values')
plt.xlabel('K')
plt.ylabel('Error')
plt.show()
knn1 = KNeighborsClassifier(n_neighbors=20)
knn1.fit(X_train,y_train)
prediction1 = knn1.predict(X_test)
print("Accuracy:",accuracy_score(y_test, prediction1)*100)
print("Precision:",precision_score(y_test, prediction1)*100)
print("Recall:",recall_score(y_test, prediction1)*100)
user_input3 = [[67.00,91.00,58.00,0,58.80,55.00,1,0,1]]
user_pred3 = knn1.predict(user_input1)
if user_pred3 == 1:
print("The Candidate will be Placed!")
else:
print("The Candidate won't be Placed :(")
# ## SVM
from sklearn.svm import SVC
svc = SVC()
svc.fit(X_train,y_train)
svc_pred = svc.predict(X_test)
print("Accuracy:",accuracy_score(y_test, svc_pred)*100)
print("Precision:",precision_score(y_test, svc_pred)*100)
print("Recall:",recall_score(y_test, svc_pred)*100)
# ## Optimizing SVM with Grid Search
from sklearn.model_selection import GridSearchCV
param_grid = {'C':[0.1,1,10,100,1000],'gamma':[1,0.1,0.01,0.001,0.0001]}
grid = GridSearchCV(SVC(),param_grid,verbose = 3)
grid.fit(X_train,y_train)
grid.best_estimator_
grid.best_params_
grid_pred = grid.predict(X_test)
print("Accuracy:",accuracy_score(y_test, grid_pred)*100)
print("Precision:",precision_score(y_test, grid_pred)*100)
print("Recall:",recall_score(y_test, grid_pred)*100)
user_input4 = [[56.00,52.00,52.00,0,59.43,66.00,1,0,1]]
user_pred4 = grid.predict(user_input4)
if user_pred4 == 1:
print("The Candidate will be Placed!")
else:
print("The Candidate won't be Placed :(")
# ## Random Forest:
from sklearn.ensemble import RandomForestClassifier
rfModel = RandomForestClassifier()
rfModel.fit(X_train, y_train)
forestPredict = rfModel.predict(X_test)
print("Accuracy:",accuracy_score(y_test, forestPredict)*100)
print("Precision:",precision_score(y_test, forestPredict)*100)
print("Recall:",recall_score(y_test, forestPredict)*100)
# ## Result
# ---
# The Result that we got aren't veru great at the moment. We were able to predict whether a person will be placed or not using the data in the table but the results aren't very great. It could be better by including some other feature or by reducing the number of features. This model still has some scope for improvement. But we could see that Logistic Regression performed better than both KNN and SVM. So atleast we have an idea which model would give us the better results.
| Campus-Recruitment/CampusPlacementAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:mysci]
# language: python
# name: conda-env-mysci-py
# ---
# # 使用 Debye 模型计算固体热容
# + [markdown] tags=[]
# 注意到粒子数 $N$ 尽管很大,但是有限,则振动模式只有 $3N$ 种。
# 即有:
# $$
# g(\epsilon) =
# \begin{cases}
# \frac{3V}{2\hbar^3\pi^2a^3}\epsilon^2,\epsilon \le \epsilon_0\\
# 0,\epsilon > \epsilon_0
# \end{cases}
# $$
# 其中 $\epsilon_0$ 满足:
# $$
# 3\cdot \frac{4}{3} \pi {\big( \frac{l\epsilon_0}{2\hbar\pi a} \big)}^3 = 3N \\
# \Leftrightarrow\epsilon_0^3 = \frac{6\hbar\pi^2 a^3}{V} N
# $$
# 从而将能级简并密度化为无量纲形式:
# $$
# g(\epsilon) = 9\frac N{\epsilon_0^3} \epsilon^2, \epsilon \le \epsilon_0
# $$
# 则有:
# $$
# n(\epsilon) = 9\frac N{\epsilon_0^3} \frac{\epsilon^2}{e^{\frac \epsilon T} -1}, \epsilon \le \epsilon_0
# $$
# 从而有内能:
# $$
# \begin{align*}
# U &= 9\frac N{\epsilon_0^3}\int_0^{\epsilon_0} \frac{\epsilon^3 d\epsilon}{e^{\frac \epsilon T} -1}\\
# &= 9\frac N{\epsilon_0^3} T^4 \int_0^{\frac{\epsilon_0} T} \frac{x^3 dx}{e^x -1}\\
# \end{align*}
# $$
# 做无量纲化:
# $$
# \begin{cases}
# N = 1\\
# \epsilon_0 = 1
# \end{cases}
# $$
# 则有:
# $$
# U = 9T^4\int_0^{\frac1 T} \frac{x^3 dx}{e^x -1}
# $$
# + [markdown] jupyter={"source_hidden": true} tags=[]
# 低温极限 $T\rightarrow 0$:
# $$
# U \rightarrow 9 T^4 \int_0^\infty \frac{x^3 dx}{e^x -1} = 54\zeta(4)\cdot T^4 \overset{recover}{\rightarrow} \frac{54\zeta(4)N}{\epsilon_0^3}\cdot T^4
# $$
# 高温极限 $T\rightarrow \infty$:
# $$
# U \rightarrow 9 T^4 \int_0^{\frac1T} x^2 dx = 3 T \overset{recover}{\rightarrow}3NT
# $$
# + tags=[]
import numpy as np
import matplotlib.pyplot as plt
import scipy.integrate as integrate
import scipy.special as special
plt.style.use("ggplot")
# -
f = lambda x : x**3/(np.exp(x) - 1)
gv = np.vectorize(lambda x: 9* x**4 * integrate.quad(f,0,1/x)[0])
gv_d = lambda x: (gv(x+1e-3)-gv(x-1e-3))/2e-3
gv_low = lambda x: 4*54*special.zeta(4)* x**4
# +
eps = 1e-2
T = np.linspace(eps,2,1000)
fig,ax = plt.subplots()
ax.plot(T,gv_d(T),label="precise")
#ax.plot(T,gv_low(T),label = "low temp lim")
ax.set_ylim(0-eps,3+2*eps)
ax.set_title("Heat Capacity for Solid")
ax.set_xlabel(r"Temperature $T$")
ax.set_ylabel(r"$C$")
ax.legend()
# +
T = np.linspace(eps,0.2,1000)
fig,ax = plt.subplots()
ax.plot(T,gv_d(T),label="precise")
ax.plot(T,gv_low(T),label = "low temp lim")
ax.set_ylim(0-eps,1.2+2*eps)
ax.set_title("Heat Capacity:Low Temperature Lim")
ax.set_xlabel(r"Temperature $T$")
ax.set_ylabel(r"$C$")
ax.legend()
# -
# # 使用一维振子链计算固体热容
# 是更加贴合实际的模型:
# * 自然的离散化,有限的自由度
| solid_heat_capacity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>REGIONE SARDEGNA
# </h1>
# Confronto dei dati relativi ai decessi registrati dall'ISTAT e i decessi causa COVID-19 registrati dalla Protezione Civile Italiana con i decessi previsti dal modello predittivo SARIMA.
# <h2>DECESSI MENSILI REGIONE SARDEGNA ISTAT</h2>
# Il DataFrame contiene i dati relativi ai decessi mensili della regione <b>Sardegna </b> dal <b>2015</b> al <b>30 settembre 2020</b>.
# +
import matplotlib.pyplot as plt
import pandas as pd
decessi_istat = pd.read_csv('../../csv/regioni/sardegna.csv')
decessi_istat.head()
# -
decessi_istat['DATA'] = pd.to_datetime(decessi_istat['DATA'])
decessi_istat.TOTALE = pd.to_numeric(decessi_istat.TOTALE)
# <h3>Recupero dei dati inerenti al periodo COVID-19</h3>
decessi_istat = decessi_istat[decessi_istat['DATA'] > '2020-02-29']
decessi_istat.head()
# <h3>Creazione serie storica dei decessi ISTAT</h3>
decessi_istat = decessi_istat.set_index('DATA')
decessi_istat = decessi_istat.TOTALE
decessi_istat
# <h2>DECESSI MENSILI REGIONE SARDEGNA CAUSATI DAL COVID</h2>
# Il DataFrame contine i dati forniti dalla Protezione Civile relativi ai decessi mensili della regione <b>Sardegna</b> da <b> marzo 2020</b> al <b>30 settembre 2020</b>.
covid = pd.read_csv('../../csv/regioni_covid/sardegna.csv')
covid.head()
covid['data'] = pd.to_datetime(covid['data'])
covid.deceduti = pd.to_numeric(covid.deceduti)
covid = covid.set_index('data')
covid.head()
# <h3>Creazione serie storica dei decessi COVID-19</h3>
covid = covid.deceduti
# <h2>PREDIZIONE DECESSI MENSILI REGIONE SECONDO MODELLO SARIMA</h2>
# Il DataFrame contiene i dati riguardanti i decessi mensili della regione <b>Sardegna</b> secondo la predizione del modello SARIMA applicato.
predictions = pd.read_csv('../../csv/pred/predictions_SARIMA_sardegna.csv')
predictions.head()
predictions.rename(columns={'Unnamed: 0': 'Data', 'predicted_mean':'Totale'}, inplace=True)
predictions.head()
predictions['Data'] = pd.to_datetime(predictions['Data'])
predictions.Totale = pd.to_numeric(predictions.Totale)
# <h3>Recupero dei dati inerenti al periodo COVID-19</h3>
predictions = predictions[predictions['Data'] > '2020-02-29']
predictions.head()
predictions = predictions.set_index('Data')
predictions.head()
# <h3>Creazione serie storica dei decessi secondo la predizione del modello</h3>
predictions = predictions.Totale
# <h1>INTERVALLI DI CONFIDENZA
# <h3>Limite massimo
upper = pd.read_csv('../../csv/upper/predictions_SARIMA_sardegna_upper.csv')
upper.head()
upper.rename(columns={'Unnamed: 0': 'Data', 'upper TOTALE':'Totale'}, inplace=True)
upper['Data'] = pd.to_datetime(upper['Data'])
upper.Totale = pd.to_numeric(upper.Totale)
upper.head()
upper = upper[upper['Data'] > '2020-02-29']
upper = upper.set_index('Data')
upper.head()
upper = upper.Totale
# <h3>Limite minimo
lower = pd.read_csv('../../csv/lower/predictions_SARIMA_sardegna_lower.csv')
lower.head()
lower.rename(columns={'Unnamed: 0': 'Data', 'lower TOTALE':'Totale'}, inplace=True)
lower['Data'] = pd.to_datetime(lower['Data'])
lower.Totale = pd.to_numeric(lower.Totale)
lower.head()
lower = lower[lower['Data'] > '2020-02-29']
lower = lower.set_index('Data')
lower.head()
lower = lower.Totale
# <h1> CONFRONTO DELLE SERIE STORICHE </h1>
# Di seguito il confronto grafico tra le serie storiche dei <b>decessi totali mensili</b>, dei <b>decessi causa COVID-19</b> e dei <b>decessi previsti dal modello SARIMA</b> della regione <b>Sardegna</b>.
# <br />
# I mesi di riferimento sono: <b>marzo</b>, <b>aprile</b>, <b>maggio</b>, <b>giugno</b>, <b>luglio</b>, <b>agosto</b> e <b>settembre</b>.
plt.figure(figsize=(15,4))
plt.title('SARDEGNA - Confronto decessi totali, decessi causa covid e decessi del modello predittivo', size=18)
plt.plot(covid, label='decessi causa covid')
plt.plot(decessi_istat, label='decessi totali')
plt.plot(predictions, label='predizione modello')
plt.legend(prop={'size': 12})
plt.show()
plt.figure(figsize=(15,4))
plt.title("SARDEGNA - Confronto decessi totali ISTAT con decessi previsti dal modello", size=18)
plt.plot(predictions, label='predizione modello')
plt.plot(upper, label='limite massimo')
plt.plot(lower, label='limite minimo')
plt.plot(decessi_istat, label='decessi totali')
plt.legend(prop={'size': 12})
plt.show()
# <h3>Calcolo dei decessi COVID-19 secondo il modello predittivo</h3>
# Differenza tra i decessi totali rilasciati dall'ISTAT e i decessi secondo la previsione del modello SARIMA.
# +
n = decessi_istat - predictions
n_upper = decessi_istat - lower
n_lower = decessi_istat - upper
plt.figure(figsize=(15,4))
plt.title("ABRUZZO - Confronto decessi accertati covid con decessi covid previsti dal modello", size=18)
plt.plot(covid, label='decessi covid accertati - Protezione Civile')
plt.plot(n, label='devessi covid previsti - modello SARIMA')
plt.plot(n_upper, label='limite massimo - modello SARIMA')
plt.plot(n_lower, label='limite minimo - modello SARIMA')
plt.legend(prop={'size': 12})
plt.show()
# -
# Gli <b>intervalli</b> corrispondono alla differenza tra i decessi totali forniti dall'ISTAT per i mesi di marzo, aprile, maggio e giugno 2020 e i valori degli <b>intervalli di confidenza</b> (intervallo superiore e intervallo inferiore) del modello predittivo SARIMA dei medesimi mesi.
d = decessi_istat.sum()
print("Decessi 2020:", d)
d_m = predictions.sum()
print("Decessi attesi dal modello 2020:", d_m)
d_lower = lower.sum()
print("Decessi attesi dal modello 2020 - livello mimino:", d_lower)
# <h3>Numero totale dei decessi accertati COVID-19 per la regione Sardegna
m = covid.sum()
print(int(m))
# <h3>Numero totale dei decessi COVID-19 previsti dal modello per la regione Sardegna</h3>
# <h4>Valore medio
total = n.sum()
print((total))
# <h4>Valore massimo
total_upper = n_upper.sum()
print((total_upper))
# <h4>Valore minimo
total_lower = n_lower.sum()
print(int(total_lower))
# <h3>Calcolo del numero dei decessi COVID-19 non registrati secondo il modello predittivo SARIMA della regione Sardegna</h3>
# <h4>Valore medio
x = decessi_istat - predictions - covid
x = x.sum()
print((x))
# <h4>Valore massimo
x_upper = decessi_istat - lower - covid
x_upper = x_upper.sum()
print((x_upper))
# <h4>Valore minimo
x_lower = decessi_istat - upper - covid
x_lower = x_lower.sum()
print(int(x_lower))
| Modulo 4 - Analisi per regioni/regioni/Sardegna/Confronto SARDEGNA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center> Compléments sur les listes, tuples, dictionnaires</center>
# ## 1. Compléments sur les listes.
# ### Listes en compréhension
# Voici un programme qui ajoute des éléments dans une liste façon successive:
L=[]
for i in range(20):
if i%2==0:
L.append(i)
print(L)
# Il existe une manière plus concise de procéder, la définition en compréhension :
# Une liste en compréhension
L=[i for i in range(20) if i%3==0]
print(L)
# #### Syntaxes
# Voici des types d'instructions possibles pour générer des listes en compréhension:
#
# * [*fonction de x* **for** *x in ensemble* ]
# * [*fonction de x* **if** *condition sur x* **else** *autre fonction de x* **for** *x in ensemble* ]
# * [*fonction de x* **for** *x in ensemble* **if** *condition sur x* ]
#
# #### Exemples:
# +
#1
L=[3*i for i in range(11)]
print(L)
#2
from random import *
n=10
Des=[randint(1,6) for i in range(n)]
print(Des)
#3
pileface=[choice(['P','F']) for i in range(20)]
print(pileface)
# -
# ### Exercice 1:
# En utilisant la syntaxe des listes en compréhension :
# 1. Générer les entiers naturels de 1 à 100.
# 2. Générer les multiples de 5 inférieurs ou égaux à 100.
# 3. Générer une liste des entiers naturels de 1 à 100 dans laquelle les multiples de 5 seront remplacées par le caractère `*`
# +
#1.
# +
#2.
# +
#3.
# -
# ### Exercice 2:
# En utilisant les fonctions `randint()` ou `choice()` du module `random` et la syntaxe des listes en compréhension:
# 1. Générer une liste de 20 entiers naturels aléatoires entre 0 et 255.
# 2. Générer 100 caractères au hasard parmi `a`,`b`,`c`
# +
#1.
from random import *
# +
#2.
# -
# ### Listes de listes
# Pour représenter des tableaux à double entrée(images, matrices...), on peut utiliser une liste de listes. On identifie ainsi un élément du tableau à l'aide de deux indexs.
#
# Exemple :
tableau=[['A','b','c','d'],['E','f','g'],['I','j','k','m'],['N','o','p','q']]
print(tableau[0][0])
# ### Exercice 3:
# 1. Quel est la lettre correpondant à `tableau[1][2]`?
# 2. Quelle instruction permet d'accéder à la lettre 'm' ?
# 3. Ajouter au tableau la ligne `['R','s','t','u']`.
# 4. Ajouter le caractère `h` à sa place.
# 5. Remplacer la caractère `N` par `n`.
# +
#1.
# +
#2.
# +
#3
# +
#4
# +
#5
# -
# ### Exercice 4:
# Générer en comprehension une liste de 10 listes contenant chacune 10 entiers binaires aléatoires(0 ou 1).
# +
# -
# ### Avant de poursuivre
# Nous avons étudié jusqu’ici deux types de données construits(composites) : les chaînes, qui sont composées de caractères, et les listes, qui sont composées d’éléments de n’importe quel type.
#
# Rappel sur une différence importante entre chaînes et listes : il n’est pas possible de changer les caractères au sein d’une chaîne existante, alors que l'on peut modifier les éléments d’une liste. En d’autres termes, les listes sont des séquences modifiables, alors que les chaînes de caractères sont des séquences non-modifiables.
#
# Exemples :
# Les listes sont modifiables
L=['a','b','d']
L[2]='c'
print(L)
# les chaînes ne sont pas modifiables
mot='abd'
mot[2]='c'
# Dans la suite de cette feuille deux nouveaux types de données : les tuples et les dictionnaires
# ## 2. tuples (ou p-uplets)
# Python propose un type de données appelé tuple, qui est assez semblable à une liste mais qui, comme les chaînes, n’est pas modifiable. Du point de vue de la syntaxe, un tuple est une collection d’éléments séparés par des virgules :
#Exécuter le code ci-dessous
tup1 = 1,2,3
tup2 = (6,7)
tup3 = 'abc','def'
tup4 = ('a',1,'b',2)
print(tup1,type(tup1))
print(tup2,type(tup2))
print(tup3,type(tup3))
print(tup4,type(tup4))
# A retenir :
# * De simples virgules suffisent à définir un tuple mais pour la lisibilité du code, il est préférable de l'enfermer dans des parenthèses.
# ### Opérations sur les tuples
#affectations
t= 7,8,9
a,b,c = t
print(a)
# opérateurs + et * : concanténation et répétition
t1= (3,2,1)
t2 = (6,5,4)
t3 = 2*t1+t2
print(t3)
# Accéder aux éléments
t4 = (2,4)
print(t4[0])
# longueur d'un tuple
print(len(t3))
# Parcours d'un tuple
t5 = (0,1,2,3,4,5,6,7,8,9)
for e in t5 :
print(e+1, end=' ')
# test in
b = 3 in (2,4,6)
print(b)
# les tuples ne sont pas des listes
t6 = (2,4,6,8)
t6.append(10)
#Ajouter un élément à un tuple
t7 = 1,2,3,4,5
t7 = t7 + (6,) # ne pas oublier la virgule et les parenthèses
print(t7)
# A retenir :
# * Les opérateurs de concaténation `+` et de multiplication `*` donnent les mêmes résultats que pour les chaînes et les listes.
# * On accède aux éléments d'un tuple comme avec les chaînes et les listes.
# * On peut déterminer la taille d’un tuple à l’aide de la fonction `len()`.
# * On peut le parcourir à l’aide d’une boucle `for`, utiliser l’instruction `in` pour savoir si un élément donné en fait partie, exactement comme pour une liste ou pour une chaîne.
# * Les tuples ne sont pas modifiables et on ne peut pas utiliser avec eux ni la méthode `.append()`ni l'instruction `del()`.
# * Les tuples sont moins souples que les listes mais c'est aussi leur force. On est sûr que leur contenu ne sera pas modifié par erreur. De plus ils sont beaucoup moins gourmands en ressources et sont éxécutés plus rapidement.
# ### Exercice 5 :
# Ecrire la fonction `reverse(t)` qui prend en paramètre un tuple de trois valeurs et qui renvoie un tuple contenant les 3 valeurs dans l'ordre inverse.
#
# Ainsi `reverse(1,2,3)` renvoie `(3,2,1)`
# +
#Réponse
# -
# ### Exercice 6 :
# Ecrire la fonction `initiales` qui prend en paramètre une chaîne de caractères de type `'NOM Prénom'` et qui renvoie un tuple contenant les initiales des noms et prénoms passés en argument.
#
# Ainsi `initiales('<NAME>')` doit renvoyer `('J','D')`.
# +
#Réponse:
# -
# ### Exercice 7 :
# En utilisant le résultat précédent, compléter la fonction `initiale_nom(noms,lettre)` qui prend en paramètres un tuple de noms et prénoms formatés comme précédemment ainsi qu'un caractère et qui renvoie un tuple contenant les chaînes avec la même initiale de nom.
#
# Ainsi avec le tuple ci-dessous, `initiale_nom(stars,'S')` doit renvoyer `('<NAME>','<NAME>', '<NAME>')`
# +
#Réponse
stars = ('<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>',
'<NAME>','<NAME>','<NAME>','<NAME>','<NAME>','<NAME>')
def initiale_nom(noms, lettre):
resultats=()
return resultats
#Appels
initiale_nom(stars,'S')
# -
# ### Exercice 8 :
# <img style='float:right;' src='fusions.png' width=200>
# On se place dans le contexte du codage des couleurs selon le système RGB, déjà vu dans une feuille précédente. Dans une image dite bitmap, chaque pixel contient une couleur composée de ses couches Rouge, Verte et Bleue, chacune de ces couches étant représentée par un entier compris entre $0$ et $255$
#
# Dans les logiciels de traitement d'images(Photoshop, Gimp,...), on travaille avec des calques d'images que l'on superpose et que l'on fusionne. En fonction des opérations mathématiques utilisées pour fusionner les calques, on obtient des rendus esthétiques différents:
# <img style='float:right;' src='http://www.info-3000.com/photoshop/modefusion02.jpg' width=400>
#
# Chacune des fonctions demandées correspondent à un mode de fusion de Photoshop , prennent en paramètres deux tuples de trois valeurs`pix1`et `pix2` correspondants aux deux pixels que l'on souhaite fusionner et renvoient un tuple contenant la couleur finale obtenue.
#
# Pour s'aider, voici une description des formules de certains modes de fusion( en milieu de page) :
# https://helpx.adobe.com/fr/after-effects/using/blending-modes-layer-styles.html
#
# On pourra tester avec les deux pixels `p1=(200,128,63)` et `p2=(125,205,50)`. Fonctions `min()` et `max()` autorisées !
p1 = (200,128,63)
p2 = (125,205,50)
# 1. `eclaircir(p1,p2)` renvoie `(200,205,63)`
# +
#1 : remplacer "to do" par les bonnes valeurs
def eclaircir(pix1, pix2):
r= "to do"
g= "to do"
b= "to do"
return (r,g,b)
eclaircir(p1,p2)
# -
# 2. `obscurcir(p1,p2)` renvoie `(125,128,50)`
# +
#2 :
def obscurcir(pix1, pix2):
r= "to do"
g= "to do"
b= "to do"
return (r,g,b)
obscurcir(p1,p2)
# -
# 3. `difference(p1,p2)` renvoie `(75,77,13)`
# +
#3 :
def difference(pix1,pix2):
r= "to do"
g= "to do"
b= "to do"
return (r,g,b)
difference(p1,p2)
# -
# 4. `addition(p1,p2)` renvoie `(255,255,113)`
# +
#4 :
def addition(pix1,pix2):
r= "to do"
g= "to do"
b= "to do"
return (r,g,b)
addition(p1,p2)
# -
# 5. `soustraction(p1,p2)` renvoie `(0,77,0)`
# +
#5 :
def soustraction(pix1,pix2):
r= "to do"
g= "to do"
b= "to do"
return (r,g,b)
soustraction(p1,p2)
# -
# 6. `produit(p1,p2)` renvoie `(98,103,12)`
# +
#6 :
def produit(pix1,pix2):
r= "to do"
g= "to do"
b= "to do"
return (r,g,b)
produit(p1,p2)
# -
# 7. `division(p1,p2)` renvoie `(159,255,202)`
# +
#7 :
def division(pix1,pix2):
r= "to do"
g= "to do"
b= "to do"
return (r,g,b)
division(p1,p2)
# -
# 8. `superposition(p1,p2)` renvoie `(226,230,100)`
# +
#8 :
def superposition(pix1,pix2):
r= "to do"
g= "to do"
b= "to do"
return (r,g,b)
superposition(p1,p2)
# -
# ## 3. Dictionnaires
# ### Définition et création
# Les types de données construits que nous avons abordés jusqu’à présent (chaînes, listes et tuples) sont tous des séquences, c’est-à-dire des suites ordonnées d’éléments. Dans une séquence, il est facile
# d’accéder à un élément quelconque à l’aide d’un index (un nombre entier), mais encore faut-il le connaître.
#
# Les dictionnaires constituent un autre type construit. Ils ressemblent aux
# listes( ils sont modifiables comme elles), mais ce ne sont pas des séquences.
# Les éléments que nous allons y enregistrer ne seront pas disposés dans un ordre immuable. En revanche, nous pourrons accéder à n’importe lequel d’entre eux à l’aide d’un index que l’on appellera une clé, laquelle pourra être alphabétique ou numérique.
#
#
# Comme dans une liste, les éléments mémorisés dans un dictionnaire peuvent être de n’importe quel type( valeurs numériques, chaînes, listes ou encore des dictionnaires, et même aussi des fonctions).
#
# Exemple :
#
# +
dico1={}
dico1['nom']='Paris-Orly Airport'
dico1['ville']='Paris'
dico1['pays']='France'
dico1['code']='ORY'
dico1['gps']=(48.7233333,2.3794444)
print(dico1)
print(dico1['code'])
print(dico1['gps'])
# -
# Remarques :
#
# - Des accolades délimitent un dictionnaire.
# - Les éléments d'un dictionnaire sont séparés par une virgule.
# - Chacun des éléments est une paire d'objets séparés par deux points : une clé et une valeur.
# - La valeur de la clé `'code'` est `'ORY'`.
# - La valeur de la clé `'gps'` est un tuple de deux flottants.
#
# ### Exercice 9 :
# Voici des données concernant l'aéroport international de Los Angeles :
#
# * __"Los Angeles International Airport","Los Angeles","United States","LAX",33.94250107,-118.4079971__
#
#
# En utilisant les mêmes clés que dans l'exemple précédent, créer le dictionnaire 'dico2' qui contiendra les données ci-dessus
# +
#Réponse
# -
# ### Méthodes
# +
#Afficher les clés
print(dico1.keys())
#Affichées les valeurs
print(dico1.values())
#Afficher les éléments
print(dico1.items())
# -
# ### Parcours
# On peut traiter par un parcours les éléments contenus dans un dictionnaire, mais attention :
# * Au cours de l'itération, __ce sont les clés__ qui servent d'index
# * L'ordre dans lequel le parcours s'effectue est imprévisible
#
# Exemple :
for element in dico1:
print(element)
# ### Exercice 10 :
# Modifier le programme ci-dessus pour obtenir l'affichage ci-dessous :
# +
# -
# ## 4. Exercices
# ### Exercice 11 :
# Lors d'une élection, entre 2 et 6 candidats se présentent. Il y a 100 votants, chacun glisse un bulletin avec le nom d'un candidat dans l'urne. Les lignes de code ci-dessous simulent cette expérience.
# +
from random import *
#liste des candidats potentiels
noms=['Alice','Bob','Charlie','Daniella','Eva','Fred']
#liste des candidats réels
candidats=list(set([choice(noms) for i in range(randint(2,len(noms)))]))
#nombre de votants
votants=100
#liste des bulletins dans l'urne
urne=[choice(candidats) for i in range(votants)]
print('Candidats : ',candidats)
#print("Contenu de l'urne :", urne)
# -
# 1. Vérifier que les candidats réels changent à chaque éxecution de la cellule ci-desssus ainsi que le contenu de l'urne.
# 2. Compléter la fonction `depouillement(urne)`. Elle prend en paramètre une liste (la liste des bulletins exprimés) et renvoie un dctionnaire. Les paires clés-valeurs sont respectivement constituées du noms d'un candidat réels et du nombre de bulletins exprimés en sa faveur. Par exemple, si la liste des candidats est `['Alice','Charlie,'Bob']`, le résultat pourra s'afficher sous la forme `{'Eva': 317, 'Charlie': 363, 'Alice': 320}`
# +
# Remplacer "pass" par les bonnes instructions
def depouillement(urne):
decompte={}
for bulletin in urne:
if bulletin in decompte:
pass
else:
pass
return decompte
depouillement(urne)
# -
# 3. Ecrire la fonction `vainqueur(election)` qui prend en paramètre un dictionnaire contenant le décompte d'une urne renvoyé par la fonction précédente et qui renvoie le nom du vainqueur.
# +
def vainqueur(election):
vainqueur=''
vainqueur(depouillement(urne))
# -
# ### Exercice 12
# <img style='float:center;' src='https://openflights.org/demo/openflights-routedb-2048.png' width=500>
#
# Sur le site https://openflights.org/data.html , on trouve des bases de données mondiales aéronautiques.Le fichier `airports.txt` présent dans le dossier de cette feuille contient des informations sur les aéroports.
#
#
# Chaque ligne de ce fichier est formatée comme l'exemple ci-dessous :
#
# `1989,"Bora Bora Airport","Bora Bora","French Polynesia","BOB","NTTB",-16.444400787353516,-151.75100708007812,10,-10,"U","Pacific/Tahiti","airport","OurAirports"`
#
# On souhaite extraire les informations suivantes pour chaque aéroport:
# * Sa référence unique, un entier.
# * Le nom de l'aéroport, une chaîne de caractères
# * La ville principale qu'il dessert, une chaîne de caractères
# * Le pays de cette ville,une chaîne de caractères
# * Le code IATA de l'aéroport composé de 3 lettres en majuscules
# * Ses coordonées gps (latitude puis longitude), un tuple de 2 flottants.
# **1. Compléter les champs ci-dessous pour l'aéroport cité en exemple:**
# * ref :
# * nom :
# * ville :
# * pays :
# * gps :
# **2. La fonction `data_extract` doit parcourir le fichier et extraire les données demandées qu'elle renvoie sous forme d'une liste de dictionnaires.**
#
# * Chaque élément de la liste est donc un dictionnaire qui correspond à un aéroport.
# * Les clés sont les noms des champs que l'on souhaite extraire et les valeurs sont celles associées à chaque aéroport.
#
# Recopier , modifier et compléter cette fonction pour qu'elle éxécute la tâche demandée :
# +
#2.
def data_extract(chemin):
fichier = open(chemin, "r",encoding='utf-8')
res = [] # Pour contenir le résultat qui sera une liste de dictionnaires
for ligne in fichier:
datas = ligne.strip().split(",") # une ligne du fichier
res.append(
{ "ref": int(datas[0]),
"nom": datas[1][1:-1],
"ville": "A compléter",
"pays": datas[3][1:-1],
"A compléter": datas[4][1:-1],
"gps" : "A compléter"
})
fichier.close()
return res
airports=data_extract('airports.txt')
#nombre d'aéroports référencés
print("A compléter")
#un aéroport au hasard
print(choice(airports))
# +
#2. Réponse
# -
# **3. A l'aide d'une liste en compréhension, récupérer la liste des villes françaises desservies par un aéroport**
#3. Réponse
country='France'
res=['A compléter']
# **4. Ecrire la fonction `infos(airports,ville)`.**
#
# Elle prend en paramètres la liste des dictionnaires de données extraites et une chaîne de caractères(le nom d'une ville). Elle renvoie les informations du ou des aéroports de cette ville dans une liste de dictionnaires:
# +
#4.
def infos(airports,ville):
res=[]
return res
print(infos(airports,'Nice'))
# -
# **5. Compléter les listes du code ci-dessous pour représenter les points de coordonnées gps de chacun des aéroports de la base de données(liste `X` des longitudes et liste `Y` des latitudes)**
# +
#5.
from matplotlib import pyplot #bibliothèque pour tracer des graphiques
X=[] #liste des longitudes
Y=[] #liste des latitudes
pyplot.figure(figsize = (14,7))
pyplot.xlim(-180, 180)
pyplot.ylim(-90, 90)
pyplot.plot(X,Y,'rx') # g=green, x est une croix
# -
# **6. Recopier et modifier le code précédent pour faire apparaître en rouge les aéroports situés en zone tropicale (c'est à dire dont la latitude est comprise entre $-23$ et $+23$)et en bleu les autres**
# +
#5.Réponse
# -
# # <center> FIN </center>
| listes_tuples_dictionnaires.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# metadata:
# interpreter:
# hash: aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# name: python3
# ---
# +
"""
Reverse the words in a string but maintain the relative order of delimiters.
For example:
input = "hello:world/here"
output = "here:world/hello"
"""
import re
def reverse(string, delimiters):
"""Split the words and delimiters into their respective lists.
Then reverse the list of words and merge the two lists together.
Complexity: O(n) time | O(n) space, where n == length of input string.
"""
words = re.split(f"[{delimiters}]+", string)
not_words = re.split(f"[^({delimiters})]+", string)
# remove last empty string if present
if words[-1] == '':
words = words[:-1]
# NOTE: we can in built reverse but there's always another way
# reversed_strings = list(reversed(words))
start = 0
end = len(words) - 1
while start < end:
words[start], words[end] = words[end], words[start]
start += 1
end -= 1
output = []
for index, delimiter in enumerate(not_words):
print(index)
output.append(delimiter)
try:
print(f"adding {words[index]}")
output.append(words[index])
except IndexError:
pass
return ''.join(output)
# -
reverse("hello:world/here", ":/")
| strings/delimiter_order.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hamiltonian Monte Carlo (HMC)
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from functools import partial
# ## Hamiltonian Monte Carlo (HMC)
#
# HMC uses an auxiliary variable corresponding to the momentum of particles in a potential energy well to generate proposal distributions that can make use of gradient information in the posterior distribution. For reversibility to be maintained, the total energy of the particle has to be conserved - hence we are interested in Hamiltonian systems. The main attraction of HMC is that it works much better than other methods when variables of interest are highly correlated. Because we have to solve problems involving momentum, we need to understand how to numerically solve differential equations in a way that is both accurate (i.e. second order) and preserves total energy (necessary for a Hamiltonian system).
#
# Example adapted from [MCMC: Hamiltonian Monte Carlo (a.k.a. Hybrid Monte Carlo)](https://theclevermachine.wordpress.com/2012/11/18/mcmc-hamiltonian-monte-carlo-a-k-a-hybrid-monte-carlo/)
# ### Hamiltonian systems
#
# In a Hamiltonian system, we consider particles with position $x$ and momentum (or velocity if we assume unit mass) $v$. The total energy of the system $H(x, v) = K(v) + U(x)$, where $K$ is the kinetic energy and $U$ is the potential energy, is conserved. Such a system satisfies the following Hamiltonian equations
#
# $$
# \begin{align}
# \frac{dx}{dt} &= & \frac{\delta H}{dv} \\
# \frac{dv}{dt} &= & -\frac{\delta H}{dx}
# \end{align}
# $$
#
# Since $K$ depends only on $v$ and $U$ depends only on $x$, we have
# $$
# \begin{align}
# \frac{dx}{dt} &= & \frac{\delta K}{dv} \\
# \frac{dv}{dt} &= & -\frac{\delta U}{dx}
# \end{align}
# $$
#
# #### Harmonic oscillator
#
# We will consider solving a classical Hamiltonian system - that of a undamped spring governed by the second order differential equation
#
# $$
# x'' + x = 0
# $$
#
# We convert this to two first order ODEs by using a dummy variable $x' = v$ to get
#
# $$
# \begin{align}
# x' &= v \\
# v' &= -x
# \end{align}
# $$
#
# From the Hamiltonian equations above, this is equivalent to a system with kinetic energy $K(v) = \frac{1}{2}v^2$ and potential energy $U(x) = \frac{1}{2}x^2$.
#
# Writing in matrix form,
#
# $$
# A = \pmatrix{ x' \\ v' } = \pmatrix{0 & 1 \\ -1 & 0} \pmatrix{x \\ v}
# $$
#
# and in general, for the state vector $x$,
#
# $$
# x' = Ax
# $$
#
# We note that $A$ is anti- or skew-symmetric ($A^T = -A$), and hence has purely imaginary eigenvalues. Solving $|A - \lambda I = 0$, we see that the eigenvalues and eigenvectors are $i, \pmatrix{1\\i}$ and $-i, \pmatrix{1\\-i}$. Since the eigenvalues are pure imaginary, we see that the solution for the initial conditions $(x,v) = (1, 0)$ is $x(t) = e^{it}$ and the orbit just goes around a circle with a period of $2\pi$, neither growing nor decaying. Another weay of seeing this is that the Hamiltonian $H(u, v)$ or sum of potential ($U(x)) = \frac{1}{2}x^2$) and kinetic energy ($K(v) = \frac{1}{2}v^2$) is constant, i.e. in vector form, $(x^T x) = \text{constant}$.
# ### Finite difference methods
#
# We want to find a finite difference approximation to $u' = Au$ that is **accurate** and **preserves total energy**. If total energy is not preserved, the orbit will either spiral in towards zero or outwards away from the unit circle. If the accuracy is poor, the orbit will not be close to its starting value after $t = 2\pi$. This gives us an easy way to visualize how good our numerical scheme is. We can also compare the numerical scheme to the Taylor series to evaluate its accuracy.
# #### Forward Euler
#
# The simplest finite difference scheme for integrating ODEs is the forward Euler
#
# $$
# \frac{u_{n+1} - u_n}{\Delta t} = A u_n
# $$
#
# Rearranging terms, we get
#
# $$
# u_{n+1} = u_n + \Delta t A u_n = \left( I + \Delta t A \right) u_n
# $$
#
# Since the eigenvalues of $A$ are $\pm i$, we see that the eigenvalues of the forward Euler matrix are $1 \pm i$. Since the absolute value of the eigenvalues is greater than 1, we expect **growing** solutions - i.e. the solution will spiral away from the unit circle.
import scipy.linalg as la
def f_euler(A, u, N):
orbit = np.zeros((N,2))
dt = 2*np.pi/N
for i in range(N):
u = u + dt * A @ u
orbit[i] = u
return orbit
A = np.array([[0,1],[-1,0]])
u = np.array([1.0,0.0])
N = 64
orbit = f_euler(A, u, N)
# ##### Accuracy
la.norm(np.array([1.0,0.0]) - orbit[-1])
# ##### Conservation of energy
plt.plot([p @ p for p in orbit])
pass
ax = plt.subplot(111)
plt.plot(orbit[:, 0], orbit[:,1], 'o')
ax.axis('square')
plt.axis([-1.5, 1.5, -1.5, 1.5])
pass
# ##### Accuracy and conservation of energy
#
# We can see that forward Euler is not very accurate and also does not preserve energy since the orbit spirals away from the unit circle.
# #### The trapezoidal method
#
# The trapezoidal method uses the following scheme
#
# $$
# \frac{u_{n+1} - u_n}{\Delta t} = \frac{1}{2} ( A u_{n+1} + A u_{n})
# $$
#
# This is an implicit scheme (because $u_{n+1}$ appears on the RHS) whose solution is
#
# $$
# u_{n+1} = \left(I - \frac{\Delta t}{2} A \right)^{-1} \left(I + \frac{\Delta t}{2} A \right) u_{n} = B u_n
# $$
#
# By inspection, we see that the eigenvalues are the complex conjugates of
#
# $$
# \frac{1 + \frac{\Delta t}{2} i}{1 - \frac{\Delta t}{2} i}
# $$
#
# whose absolute value is 1 - hence, energy is conserved. If we expand the matrix $B$ using the geometric series and compare with the Taylor expansion, we see that the trapezoidal method has local truncation error $O(h^3)$ and hence accuracy $O(h^2)$, where $h$ is the time step.
def trapezoidal(A, u, N):
p = len(u)
orbit = np.zeros((N,p))
dt = 2*np.pi/N
for i in range(N):
u = la.inv(np.eye(p) - dt/2 * A) @ (np.eye(p) + dt/2 * A) @ u
orbit[i] = u
return orbit
A = np.array([[0,1],[-1,0]])
u = np.array([1.0,0.0])
N = 64
orbit = trapezoidal(A, u, N)
# ##### Accuracy
la.norm(np.array([1.0,0.0]) - orbit[-1])
# ##### Conservation of energy
plt.plot([p @ p for p in orbit])
pass
ax = plt.subplot(111)
plt.plot(orbit[:, 0], orbit[:,1], 'o')
ax.axis('square')
plt.axis([-1.5, 1.5, -1.5, 1.5])
pass
# #### The leapfrog method
#
# The leapfrog method uses a second order difference to update $u_n$. The algorithm simplifies to the following explicit scheme:
#
# - First take one half-step for v
# - Then take a full step for u
# - Then take one final half step for v
#
# It performs almost as well as the trapezoidal method, with the advantage of being an explicit scheme and cheaper to calculate, so the leapfrog method is used in HMC.
def leapfrog(A, u, N):
orbit = np.zeros((N,2))
dt = 2*np.pi/N
for i in range(N):
u[1] = u[1] + dt/2 * A[1] @ u
u[0] = u[0] + dt * A[0] @ u
u[1] = u[1] + dt/2 * A[1] @ u
orbit[i] = u
return orbit
# ##### If we don't care about the intermediate steps, it is more efficient to just take 1/2 steps at the beginning and end
def leapfrog2(A, u, N):
dt = 2*np.pi/N
u[1] = u[1] + dt/2 * A[1] @ u
for i in range(N-1):
u[0] = u[0] + dt * A[0] @ u
u[1] = u[1] + dt * A[1] @ u
u[0] = u[0] + dt * A[0] @ u
u[1] = u[1] + dt/2 * A[1] @ u
return u
A = np.array([[0,1],[-1,0]])
u = np.array([1.0,0.0])
N = 64
orbit = leapfrog(A, u, N)
# ##### Accuracy
la.norm(np.array([1.0,0.0]) - orbit[-1])
# ##### Conservation of energy
#
# Note that unlike the trapezoidal scheme, energy is not perfectly conserved.
plt.plot([p @ p for p in orbit])
pass
ax = plt.subplot(111)
plt.plot(orbit[:, 0], orbit[:,1], 'o')
ax.axis('square')
plt.axis([-1.5, 1.5, -1.5, 1.5])
pass
# ### From Hamiltonians to probability distributions
#
# The physical analogy considers the negative log likelihood of the target distribution $p(x)$ to correspond to a potential energy well, with a collection of particles moving on the surface of the well. The state of each particle is given only by its position and momentum (or velocity if we assume unit mass for each particle). In a Hamiltonian system, the total energy $H(x, v) = U(x) + K(v)$ is conserved. From statistical mechanics, the probability of each state is related to the total energy of the system
#
# $$
# \begin{align}
# p(x, v) & \propto e^{-H(x, v)} \\
# &= e^{-U(x) - K(v)} \\
# &= e^{-P(x)}e^{-K(v)} \\
# & \propto p(x) \, p(v)
# \end{align}
# $$
#
# Since the joint distribution factorizes $p(x, v) = p(x)\, p(v)$, we can select an initial random $v$ for a particle, numerically integrate using a finite difference method such as the leapfrog and then use the updated $x^*$ as the new proposal. The acceptance ratio for the new $x^*$ is
#
# $$
# \frac{ e^{ -U(x^*)-K(v^*) }} { e^{-U(x)-K(v)} } = e^{U(x)-U(x^*)+K(x)-K(x^*)}
# $$
#
# If our finite difference scheme was exact, the acceptance ration would be 1 since energy is conserved with Hamiltonian dynamics. However, as we have seen, the leapfrog method does not conserve energy perfectly and an accept/reject step is still needed.
# #### Example of HMC
#
# We will explore how HMC works when the target distribution is bivariate normal centered at zero
#
# $$
# x \sim N(0, \Sigma)
# $$
#
# In practice of course, the target distribution will be the posterior distribution and depend on both data and distributional parameters.
#
# The potential energy or negative log likelihood is proportional to
# $$
# U(x) = \frac{x^T\Sigma^{-1} x}{2}
# $$
#
# The kinetic energy is given by
# $$
# K(v) = \frac{v^T v}{2}
# $$
#
# where the initial $v_0$ is chosen at random from the unit normal at each step.
#
# To find the time updates, we use the Hamiltonian equations and find the first derivatives of total energy with respect to $x$ and $v$
#
# $$
# \begin{align}
# x' &= \frac{\delta K}{\delta v} &= v \\
# v' &= -\frac{\delta U}{\delta x} &= -\Sigma^{-1} x \\
# \end{align}
# $$
#
# giving us the block matrix
#
# $$
# A = \pmatrix{0 & 1 \\ -\Sigma^{-1} & 0}
# $$
#
# By using the first derivatives, we are making use of the gradient information on the log posterior to guide the proposal distribution.
# ##### This is what the target distribution should look like
sigma = np.array([[1,0.8],[0.8,1]])
mu = np.zeros(2)
ys = np.random.multivariate_normal(mu, sigma, 1000)
sns.kdeplot(ys[:,0], ys[:,1])
plt.axis([-3.5,3.5,-3.5,3.5])
pass
# ##### This is the HMC posterior
def E(A, u0, v0, u, v):
"""Total energy."""
return (u0 @ tau @ u0 + v0 @ v0) - (u @ tau@u + v @ v)
def leapfrog(A, u, v, h, N):
"""Leapfrog finite difference scheme."""
v = v - h/2 * A @ u
for i in range(N-1):
u = u + h * v
v = v - h * A @ u
u = u + h * v
v = v - h/2 * A @ u
return u, v
# +
niter = 100
h = 0.01
N = 100
tau = la.inv(sigma)
orbit = np.zeros((niter+1, 2))
u = np.array([-3,3])
orbit[0] = u
for k in range(niter):
v0 = np.random.normal(0,1,2)
u, v = leapfrog(tau, u, v0, h, N)
# accept-reject
u0 = orbit[k]
a = np.exp(E(A, u0, v0, u, v))
r = np.random.rand()
if r < a:
orbit[k+1] = u
else:
orbit[k+1] = u0
# -
sns.kdeplot(orbit[:, 0], orbit[:, 1])
plt.plot(orbit[:,0], orbit[:,1], alpha=0.2)
plt.scatter(orbit[:1,0], orbit[:1,1], c='red', s=30)
plt.scatter(orbit[1:,0], orbit[1:,1], c=np.arange(niter)[::-1], cmap='Reds')
plt.axis([-3.5,3.5,-3.5,3.5])
pass
| notebooks/copies/lectures/T08F_HMC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS" tags=[]
# # LD workflow
#
# This workflow generate the LD matrixs base on the genotype for each gene and saved in both RDS and text format
#
# **FIXME: we may compute it with SuSiE and not use this pipeline. It will be removed once the SuSiE based pipeline is done**
#
# 
# + [markdown] kernel="SoS"
# ## LD computation for each gene
# This workflow takes in list of per gene plink trio and compute LD for them, one row of LD recipe will be generated to be merged later on
# + kernel="SoS"
[global]
# the output directory for generated files
parameter: wd = path
cwd = wd
# A string to identify your analysis run
parameter: name = f"{cwd:b}"
# For cluster jobs, number commands to run per job
parameter: job_size = 1
# Wall clock time expected
parameter: walltime = "5h"
# Memory expected
parameter: mem = "16G"
# Number of threads
parameter: numThreads = 20
# Software container option
parameter: container = 'statisticalgenetics/lmm:2.4'
# use this function to edit memory string for PLINK input
from sos.utils import expand_size
cwd = f"{cwd:a}"
import pandas as pd
parameter: genotype_list = path
geno_file_inv = pd.read_csv(genotype_list, sep = "\t")
genoFile = geno_file_inv["dir"].values.tolist()
# + kernel="SoS"
# Filter SNPs and select individuals
[LD_1]
# Window size
parameter: window = 50
# Shift window every 10 snps
parameter: shift = 10
parameter: r2 = 0.1
input: genoFile, group_by = 1
output: f'{cwd}/{name}_ld/{_input:bn}.ld',
ld_rds = f'{cwd}/{name}_ld/{_input:bn}.ld.rds'
task: trunk_workers = 1, walltime = walltime, mem = mem, cores = numThreads, tags = f'{step_name}_{_output:bn}'
bash: expand= "${ }", stderr = f'{_output[0]:n}.stderr', stdout = f'{_output[0]:n}.stdout',container = container
plink \
--bfile ${_input:n} \
--indep-pairwise ${window} ${shift} ${r2} \
--out ${_output[0]:n} \
--threads ${numThreads} \
--memory ${int(expand_size(mem) * 0.9)/1e6} \
--r square \
--allow-no-sex
R: expand = '${ }', stdout = f"{_output[0]:n}.stdout", stderr = f"{_output[0]:n}.stderr", container = container
name = read.table("${_input:n}.bim",sep = "\t")$V2
ld = as.matrix(data.table::fread("${_output[0]}"))
rownames(ld) <- colnames(ld) <- name
saveRDS(ld, "${_output[1]}")
# + kernel="SoS"
[LD_2]
import pandas as pd
input: group_by = "all"
output: f'{cwd}/{name}_LD_recipe'
df = pd.DataFrame({
"Theme" : name,
"ld_file_prefix" : f'{cwd}/{name}_ld/',
"ld_file_surfix" : ".ld.rds"
},index = [0])
df.to_csv(_output,sep = "\t",index = 0)
| pipeline/LD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1. Testing quality of predictions
# ==
#
# We now have a function that can predict the price for any living space we want to list as long as we know the number of people it can accommodate. The function we wrote represents a **machine learning model**, which means that it outputs a prediction based on the input to the model.
#
# A simple way to test the quality of your model is to:
#
# - split the dataset into 2 partitions:
# - the training set: contains the majority of the rows (75%)
# - the test set: contains the remaining minority of the rows (25%)
# - use the rows in the training set to predict the <span style="background-color: #F9EBEA; color:##C0392B">price</span> value for the rows in the test set
# - add new column named <span style="background-color: #F9EBEA; color:##C0392B">predicted_price</span> to the test set
# - compare the <span style="background-color: #F9EBEA; color:##C0392B">predicted_price</span> values with the actual <span style="background-color: #F9EBEA; color:##C0392B">price</span> values in the test set to see how accurate the predicted values were.
#
# This validation process, where we use the training set to make predictions and the test set to predict values for, is known as **train/test validation**. Whenever you're performing machine learning, you want to perform validation of some kind to ensure that your machine learning model can make good predictions on new data. While train/test validation isn't perfect, we'll use it to understand the validation process, to select an error metric, and then we'll dive into a more robust validation process later in this course.
#
# Let's modify the <span style="background-color: #F9EBEA; color:##C0392B">predicted_price</span> function to use only the rows in the training set, instead of the full dataset, to find the nearest neighbors, average the <span style="background-color: #F9EBEA; color:##C0392B">price</span> values for those rows, and return the predicted price value. Then, we'll use this function to predict the price for just the rows in the test set. Once we have the predicted price values, we can compare with the true price values and start to understand the model's effectiveness in the next screen.
#
# To start, we've gone ahead and assigned the first 75% of the rows in <span style="background-color: #F9EBEA; color:##C0392B">dc_listings</span> to <span style="background-color: #F9EBEA; color:##C0392B">train_df</span> and the last 25% of the rows to <span style="background-color: #F9EBEA; color:##C0392B">test_df</span>. Here's a diagram explaining the split:
#
# <img width="600" alt="creating a repo" src="https://drive.google.com/uc?export=view&id=11IctHIyFi18HxRsg9LpsOf4tVKqfqvRz">
#
# <br>
# <div class="alert alert-info">
# <b>Exercise Start.</b>
# </div>
#
# **Description**:
#
# 1. Within the <span style="background-color: #F9EBEA; color:##C0392B">predict_price</span> function, change the Dataframe that <span style="background-color: #F9EBEA; color:##C0392B">temp_df</span> is assigned to. Change it from <span style="background-color: #F9EBEA; color:##C0392B">dc_listings</span> to <span style="background-color: #F9EBEA; color:##C0392B">train_df</span>, so only the training set is used.
# 2. Use the Series method <span style="background-color: #F9EBEA; color:##C0392B">apply</span> to pass all of the values in the <span style="background-color: #F9EBEA; color:##C0392B">accommodates</span> column from <span style="background-color: #F9EBEA; color:##C0392B">test_df</span> through the <span style="background-color: #F9EBEA; color:##C0392B">predict_price</span> function.
# 3. Assign the resulting Series object to the <span style="background-color: #F9EBEA; color:##C0392B">predict_price</span> column in <span style="background-color: #F9EBEA; color:##C0392B">test_df</span>.
# +
# importing packages
import pandas as pd
import numpy as np
# import dataset
dc_listings = pd.read_csv("dc_airbnb.csv")
# cleaning & preparing
stripped_commas = dc_listings['price'].str.replace(',', '')
stripped_dollars = stripped_commas.str.replace('$', '')
dc_listings['price'] = stripped_dollars.astype('float')
# separte data into train and test (75%/25%)
train_df = dc_listings.iloc[0:2792]
test_df = dc_listings.iloc[2792:]
def predict_price(new_listing):
temp_df = dc_listings
temp_df['distance'] = temp_df['accommodates'].apply(lambda x: np.abs(x - new_listing))
temp_df = temp_df.sort_values('distance')
nearest_neighbor_prices = temp_df.iloc[0:5]['price']
predicted_price = nearest_neighbor_prices.mean()
return(predicted_price)
test_df['predict_price'] = test_df['accommodates'].apply(lambda x: predict_price(x))
# -
# 2. Error Metrics
# ==
#
# We now need a metric that quantifies how good the predictions were on the test set. This class of metrics is called an **error metric**. As the name suggests, an error metric quantifies how inaccurate our predictions were from the actual values. In our case, the error metric tells us how off our predicted price values were from the actual price values for the living spaces in the test dataset.
#
# We could start by calculating the difference between each predicted and actual value and then averaging these differences. This is referred to as **mean error** but isn't an effective error metric for most cases. Mean error treats a positive difference differently than a negative difference, but we're really interested in how far off the prediction is in either the positive or negative direction. If the true price was 200 dollars and the model predicted 210 or 190 it's off by 10 dollars either way.
#
# We can instead use the **mean absolute error**, where we compute the absolute value of each error before we average all the errors.
#
# $\displaystyle MAE = \frac{\left | actual_1 - predicted_1 \right | + \left | actual_2 - predicted_2 \right | + \
# \ldots + \left | actual_n - predicted_n \right | }{n}$
#
# <br>
# <div class="alert alert-info">
# <b>Exercise Start.</b>
# </div>
#
# **Description**:
#
# 1. Use <span style="background-color: #F9EBEA; color:##C0392B">numpy.absolute()</span> to calculate the mean absolute error between <span style="background-color: #F9EBEA; color:##C0392B">predicted_price</span> and <span style="background-color: #F9EBEA; color:##C0392B">price</span>.
# 2. Assign the MAE to <span style="background-color: #F9EBEA; color:##C0392B">mae</span>.
mae = np.absolute(test_df.price - test_df.predict_price).mean()
mae
# 3. Mean Squared Error
# ==
#
# For many prediction tasks, we want to penalize predicted values that are further away from the actual value much more than those that are closer to the actual value.
#
# We can instead take the mean of the squared error values, which is called the **mean squared error** or MSE for short. The MSE makes the gap between the predicted and actual values more clear. A prediction that's off by 100 dollars will have an error (of 10,000) that's 100 times more than a prediction that's off by only 10 dollars (which will have an error of 100).
#
# Here's the formula for MSE:
#
# $\displaystyle MSE = \frac{(actual_1 - predicted_1)^2 + (actual_2 - predicted_2)^2 + \
# \ldots + (actual_n - predicted_n)^2 }{n}$
#
# where **n** represents the number of rows in the test set. Let's calculate the MSE value for the predictions we made on the test set.
#
# <br>
# <div class="alert alert-info">
# <b>Exercise Start.</b>
# </div>
#
# **Description**:
#
# 1. Calculate the MSE value between the <span style="background-color: #F9EBEA; color:##C0392B">predicted_price</span> and <span style="background-color: #F9EBEA; color:##C0392B">price</span> columns and assign to <span style="background-color: #F9EBEA; color:##C0392B">mse</span>.
mse = (np.absolute(test_df.price - test_df.predict_price)**2).mean()
mse
# 4. Training another model
# ==
#
# The model we trained achieved a mean squared error of around **18646.5**. Is this a high or a low mean squared error value? What does this tell us about the quality of the predictions and the model? By itself, the mean squared error value for a single model isn't all that useful.
#
# The units of mean squared error in our case is dollars squared (not dollars), which makes it hard to reason about intuitively as well. We can, however, train another model and then compare the mean squared error values to see which model performs better on a relative basis. Recall that a low error metric means that the gap between the predicted list price and actual list price values is low while a high error metric means the gap is high.
#
# Let's train another model, this time using the <span style="background-color: #F9EBEA; color:##C0392B">bathrooms</span> column, and compare MSE values.
#
# <br>
# <div class="alert alert-info">
# <b>Exercise Start.</b>
# </div>
#
# **Description**:
#
# 1. Modify the <span style="background-color: #F9EBEA; color:##C0392B">predict_price</span> function below to use the <span style="background-color: #F9EBEA; color:##C0392B">bathrooms</span> column instead of the <span style="background-color: #F9EBEA; color:##C0392B">accommodates</span> column to make predictions.
# 2. Apply the function to <span style="background-color: #F9EBEA; color:##C0392B">test_df</span> and assign the resulting Series object containing the predicted price values to the <span style="background-color: #F9EBEA; color:##C0392B">predicted_price</span> column in <span style="background-color: #F9EBEA; color:##C0392B">test_df</span>.
# 3. Calculate the squared error between the price and <span style="background-color: #F9EBEA; color:##C0392B">predicted_price</span> columns in <span style="background-color: #F9EBEA; color:##C0392B">test_df</span> and assign the resulting Series object to the <span style="background-color: #F9EBEA; color:##C0392B">squared_error</span> column in <span style="background-color: #F9EBEA; color:##C0392B">test_df</span>.
# 4. Calculate the mean of the <span style="background-color: #F9EBEA; color:##C0392B">squared_error</span> column in <span style="background-color: #F9EBEA; color:##C0392B">test_df</span> and assign to <span style="background-color: #F9EBEA; color:##C0392B">mse</span>.
# 5. Use the <span style="background-color: #F9EBEA; color:##C0392B">print</span> function or the variables inspector to display the MSE value.
#
# +
train_df = dc_listings.iloc[0:2792]
test_df = dc_listings.iloc[2792:]
def predict_price(new_listing):
temp_df = train_df
temp_df['distance'] = temp_df['bathrooms'].apply(lambda x: np.abs(x - new_listing))
temp_df = temp_df.sort_values('distance')
nearest_neighbors_prices = temp_df.iloc[0:5]['price']
predicted_price = nearest_neighbors_prices.mean()
return(predicted_price)
test_df['predict_price'] = test_df['bathrooms'].apply(lambda x: predict_price(x))
test_df['squared_error'] = (np.absolute(test_df.price - test_df.predict_price)**2)
mse = test_df['squared_error'].mean()
print(mse)
# -
# 5. Root Mean Squared Error
# ==
#
# While comparing MSE values helps us identify which model performs better on a relative basis, it doesn't help us understand if the performance is good enough in general. This is because the units of the MSE metric are squared (in this case, dollars squared). An MSE value of 16377.5 dollars squared doesn't give us an intuitive sense of how far off the model's predictions are systematically off from the true price value in dollars.
#
# **Root mean squared error** is an error metric whose units are the base unit (in our case, dollars). RMSE for short, this error metric is calculated by taking the square root of the MSE value:
#
# $\displaystyle RMSE=\sqrt{MSE}$
#
# Since the RMSE value uses the same units as the target column, we can understand how far off in real dollars we can expect the model to perform. For example, if a model achieves an RMSE value of greater than 100, we can expect the predicted price value to be off by 100 dollars on average.
#
# Let's calculate the RMSE value of the model we trained using the <span style="background-color: #F9EBEA; color:##C0392B">bathrooms</span> column.
#
#
# <br>
# <div class="alert alert-info">
# <b>Exercise Start.</b>
# </div>
#
# **Description**:
#
# 1. Calculate the RMSE value of the model we trained using the <span style="background-color: #F9EBEA; color:##C0392B">bathrooms</span> column and assign it to **rmse**.
#
rmse = np.sqrt(mse)
print(rmse)
# 6. Comparing MAE and RMSE
# ==
#
# The model achieved an RMSE value of approximately **135.6**, which implies that we should expect for the model to be off by **135.6** dollars on average for the predicted price values. Given that most of the living spaces are listed at just a few hundred dollars, we need to reduce this error as much as possible to improve the model's usefulness.
#
# We discussed a few different error metrics we can use to understand a model's performance. As we mentioned earlier, these individual error metrics are helpeful for comparing models. To better understand a specific model, we can compare multiple error metrics for the same model. This requires a better understanding of the mathematical properties of the error metrics.
#
# If you look at the equation for MAE:
#
# $\displaystyle MAE = \frac{\left | actual_1 - predicted_1 \right | + \left | actual_2 - predicted_2 \right | + \
# \ldots + \left | actual_n - predicted_n \right | }{n}$
#
# you'll notice that a prediction that the individual errors (or differences between predicted and actual values) grow linearly. A prediction that's off by 10 dollars has a 10 times higher error than a prediction that's off by 1 dollar. If you look at the equation for RMSE, however:
#
# $\displaystyle RMSE = \sqrt{\frac{(actual_1 - predicted_1)^2 + (actual_2 - predicted_2)^2 + \
# \ldots + (actual_n - predicted_n)^2 }{n}}$
#
# you'll notice that each error is squared before the square root of the sum of all the errors is taken. This means that the individual errors grows quadratically and has a different effect on the final RMSE value.
#
# Let's look at an example using different data entirely. We've created 2 Series objects containing 2 sets of errors and assigned to <span style="background-color: #F9EBEA; color:##C0392B">errors_one</span> and <span style="background-color: #F9EBEA; color:##C0392B">errors_two</span>.
#
# <br>
# <div class="alert alert-info">
# <b>Exercise Start.</b>
# </div>
#
# **Description**:
#
# 1. Calculate the MAE for <span style="background-color: #F9EBEA; color:##C0392B">errors_one</span> and assign to <span style="background-color: #F9EBEA; color:##C0392B">mae_one</span>.
# 2. Calculate the RMSE for <span style="background-color: #F9EBEA; color:##C0392B">errors_one</span> and assign to <span style="background-color: #F9EBEA; color:##C0392B">rmse_one</span>.
# 3. Calculate the MAE for <span style="background-color: #F9EBEA; color:##C0392B">errors_two</span> and assign to <span style="background-color: #F9EBEA; color:##C0392B">mae_two</span>.
# 4. Calculate the RMSE for <span style="background-color: #F9EBEA; color:##C0392B">errors_two</span> and assign to <span style="background-color: #F9EBEA; color:##C0392B">rmse_two</span>.
# +
errors_one = pd.Series([5, 10, 5, 10, 5, 10, 5, 10, 5, 10, 5, 10, 5, 10, 5, 10, 5, 10])
errors_two = pd.Series([5, 10, 5, 10, 5, 10, 5, 10, 5, 10, 5, 10, 5, 10, 5, 10, 5, 1000])
mae_one = errors_one.mean()
rmse_one = np.sqrt((errors_one**2).mean())
mae_two = errors_two.mean()
rmse_two = np.sqrt((errors_two**2).mean())
print(mae_one)
print(rmse_one)
print(mae_two)
print(rmse_two)
# -
# 7. Next steps
# ==
#
# While the MAE (7.5) to RMSE (7.9056941504209481) ratio was about 1:1 for the first list of errors, the MAE (62.5) to RMSE (235.82302686548658) ratio was closer to 1:4 for the second list of errors. The only difference between the 2 sets of errors is the extreme 1000 value in errors_two instead of 10. When we're working with larger data sets, we can't inspect each value to understand if there's one or some outliers or if all of the errors are systematically higher. Looking at the ratio of MAE to RMSE can help us understand if there are large but infrequent errors. You can read more about comparing MAE and RMSE in [this wonderful post](https://medium.com/human-in-a-machine-world/mae-and-rmse-which-metric-is-better-e60ac3bde13d#.lyc8od1ix).
#
# In this mission, we learned how to test our machine learning models using basic cross validation and different metrics. In the next 2 missions, we'll explore how adding more features to the machine learning model and selecting a more optimal k value can help improve the model's performance.
| Lesson 13_Airbnb/Part II - Evaluating Model Performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Anaconda3]
# language: python
# name: conda-env-Anaconda3-py
# ---
from __future__ import print_function, division
from keras.datasets import fashion_mnist
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.interpolate import interp1d
import os
from keras.layers import Input, Dense, Reshape, Flatten, Dropout, multiply
from keras.layers import BatchNormalization, Activation, Embedding, ZeroPadding2D
from keras.layers.advanced_activations import LeakyReLU
from keras.layers.convolutional import UpSampling2D, Conv2D, MaxPooling2D
from keras.models import Sequential, Model
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint
from keras.utils import to_categorical
name = 'fashion_CNN'
if not os.path.exists("saved_model/"+name):
os.mkdir("saved_model/"+name)
if not os.path.exists("images/"+name):
os.mkdir("images/"+name)
# Download the dataset
(x_train, y_train), (x_test, y_test) = fashion_mnist.load_data()
print('x_train', x_train.shape,'y_train', y_train.shape)
print('x_test', x_test.shape,'y_test', y_test.shape)
input_classes = pd.Series(y_train).nunique()
input_classes
# Training Labels are evenly distributed
Train_label_count = pd.Series(y_train).value_counts()
Train_label_count
# Test Labels are evenly distributed
Test_label_count = pd.Series(y_test).value_counts()
Test_label_count
#label dictionary from documentation
label_dict = {0: 'tshirt',
1: 'trouser',
2: 'pullover',
3: 'dress',
4: 'coat',
5: 'sandal',
6: 'shirt',
7: 'sneaker',
8: 'bag',
9: 'boot'}
x_train[1].shape
#input dimensions
input_rows = x_train[1][0]
input_cols = x_train[1][1]
input_channels = 1
# plot images from the train dataset
for i in range(10):
# define subplot
a=plt.subplot(2, 5, 1 + i)
# turn off axis
plt.axis('off')
# plot raw pixel data
plt.imshow(x_train[i], cmap='gray_r')
a.set_title(y_train[i])
# plot images from the test dataset
for i in range(10):
# define subplot
a=plt.subplot(2, 5, 1 + i)
# turn off axis
plt.axis('off')
# plot raw pixel data
plt.imshow(x_test[i], cmap='gray_r')
a.set_title(y_test[i])
# +
x_train = x_train.astype('float32') / 255
x_test = x_test.astype('float32') / 255
# Further break training data into train / validation sets (# put 5000 into validation set and keep remaining 55,000 for train)
(x_train, x_valid) = x_train[5000:], x_train[:5000]
(y_train, y_valid) = y_train[5000:], y_train[:5000]
# Reshape input data from (28, 28) to (28, 28, 1)
w, h = 28, 28
x_train = x_train.reshape(x_train.shape[0], w, h, 1)
x_valid = x_valid.reshape(x_valid.shape[0], w, h, 1)
x_test = x_test.reshape(x_test.shape[0], w, h, 1)
# One-hot encode the labels
y_train = to_categorical(y_train, 10)
y_valid = to_categorical(y_valid, 10)
y_test = to_categorical(y_test, 10)
# Print training set shape
print("x_train shape:", x_train.shape, "y_train shape:", y_train.shape)
# Print the number of training, validation, and test datasets
print(x_train.shape[0], 'train set')
print(x_valid.shape[0], 'validation set')
print(x_test.shape[0], 'test set')
# -
model = Sequential()# Must define the input shape in the first layer of the neural network
model.add(Conv2D(filters=64, kernel_size=2, padding='same', activation='relu', input_shape=(28,28,1)))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.3))
model.add(Conv2D(filters=32, kernel_size=2, padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.3))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax'))# Take a look at the model summary
model.summary()
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
checkpointer = ModelCheckpoint(filepath="saved_model/"+name+'/model.weights.best.hdf5', verbose = 1, save_best_only=True)
model.fit(x_train,
y_train,
batch_size=64,
epochs=10,
validation_data=(x_valid, y_valid),
callbacks=[checkpointer])
# Evaluate the model on test set
score = model.evaluate(x_test, y_test, verbose=0)# Print test accuracy
print('\n', 'Test accuracy:', score[1])
# +
y_hat = model.predict(x_test)
# Plot a random sample of 10 test images, their predicted labels and ground truth
figure = plt.figure(figsize=(20, 8))
for i, index in enumerate(np.random.choice(x_test.shape[0], size=15, replace=False)):
ax = figure.add_subplot(3, 5, i + 1, xticks=[], yticks=[])
# Display each image
ax.imshow(np.squeeze(x_test[index]))
predict_index = np.argmax(y_hat[index])
true_index = np.argmax(y_test[index])
# Set the title for each image
ax.set_title("{} ({})".format(label_dict[predict_index],
label_dict[true_index]),
color=("green" if predict_index == true_index else "red"))
# -
| Predict_Fashion_using_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center>
# <b>CompEcon Toolbox:</b>
# <div style="font-size:175%;color:white; background-color: #0064b0;">DemQua08</div>
# <div style="font-size:250%;color:white; background-color: #0064b0;">Illustrates integration using Simpson's rule</div>
#
# <b><NAME>, PhD</b>
# <br><br>
#
# </center>
#
# This demo is based on the original Matlab demo accompanying the <a href="https://mitpress.mit.edu/books/applied-computational-economics-and-finance">Computational Economics and Finance</a> 2001 textbook by <NAME> and <NAME>.
#
#
# <i>Last updated: 2020-Sep-10</i>
# ## Initial tasks
if 'google.colab' in str(get_ipython()):
print("This notebook is running on Google Colab. Installing the compecon package.")
# !pip install compecon
from numpy import poly1d,polyfit, linspace, array
from compecon import qnwsimp, demo
import matplotlib.pyplot as plt
n = 1001
xmin, xmax = -1, 1
xwid = xmax-xmin
x = linspace(xmin, xmax, n)
f = poly1d([2.0, -1.0, 0.5, 5.0])
def fitquad(xi):
newcoef = polyfit(xi, f(xi), 2)
return poly1d(newcoef)
def plot_simp(n):
xi, wi = qnwsimp(n+1, xmin, xmax)
fig, ax = plt.subplots()
ax.plot(x, f(x), linewidth=3)
for k in range(n//2):
xii = xi[(2*k):(2*k+3)]
xiii = linspace(xii[0], xii[2], 125)
p = fitquad(xii)
ax.fill_between(xiii, p(xiii), alpha=0.35, color='LightSkyBlue')
ax.plot(xiii, p(xiii),color='Tab:Orange', linestyle='--')
plt.vlines(xi, 0, f(xi),'k', linestyle=':')
plt.hlines(0,xmin-0.1, xmax+0.1,'k',linewidth=2)
plt.xlim(xmin-0.1, xmax+0.1)
xtl = ['$x_{%d}$' % i for i in range(n+1)]
xtl[0] += '=a'
xtl[n] += '=b'
plt.xticks(xi, xtl)
plt.yticks([0],['0'])
plt.legend([r'$f(x)$', f'$\\tilde{{f}}_{n+1}(x)$'])
return fig
# +
def plot_simp(n):
xi, wi = qnwsimp(n+1, xmin, xmax)
fig, ax = plt.subplots()
ax.plot(x, f(x), linewidth=3)
for k in range(n//2):
xii = xi[(2*k):(2*k+3)]
xiii = linspace(xii[0], xii[2], 125)
p = fitquad(xii)
ax.fill_between(xiii, p(xiii), alpha=0.35, color='LightSkyBlue')
ax.plot(xiii, p(xiii), color='Tab:Orange', linestyle='--')
ax.vlines(xi, 0, f(xi), color='Tab:Orange', linestyle=':')
ax.axhline(0,color='k',linewidth=2)
xtl = [f'$x_{i}$' for i in range(n+1)]
xtl[0] += '=a'
xtl[n] += '=b'
ax.set(xlim=[xmin-0.1, xmax+0.1], xticks=xi, xticklabels=xtl,
yticks=[0], yticklabels=['0'])
plt.legend([r'$f(x)$', f'$\\tilde{{f}}_{n+1}(x)$'])
return fig
figs = [plot_simp(n) for n in [2, 4, 8]]
#demo.savefig(figs,name='demqua08')
| notebooks/qua/08 Illustrates integration using Simpson's rule.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Imports
import requests
import json
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import PIL
from IPython.display import Image
# ### API Server
#apiServer = "http://localhost:8080/api/v1"
apiServer = "http://x570-test.local:8080/api/v1"
# ### Input Image
image = "sample_facedetect.jpg"
Image(filename=image)
# ### Classify Request (remote)
# +
files = {'image': (image, open(image, 'rb') ) }
model = "densebox_320_320"
# make request
response = requests.post(apiServer + '/images/face-detect?model=' + model, files=files)
response.status_code
# -
# ### Inspect results:
# +
jsonResponse = json.loads(response.content)
for result in jsonResponse['data']['results']:
print("Pox: x={} y={} Size: w={} h={} Score: {}".format(
result['x'], result['y'], result['width'], result['height'], result['score']))
# Create figure and axes
fig, ax = plt.subplots()
# Display the image
ax.imshow(PIL.Image.open(image))
# Create a Rectangle patch
for result in jsonResponse['data']['results']:
ax.add_patch(patches.Rectangle(
(result['x'], result['y']), result['width'], result['height'], linewidth=2, edgecolor='r', facecolor='none'))
# Add the patch to the Axes
plt.show()
| Client/Python/Face-Detect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Function, Scope
def maxval(x,y):
if x>y:
return x
else:
return y
maxval(3,4)
def printName(firstname, lastname, reverse=True):
if reverse:
print(lastname + "," + firstname)
else:
print(firstname, lastname)
printName("Oiga","Puchmajerova", False)
printName("Oiga","Puchmajerova")
def f(x):
y = 1
x += y
print("x =", x)
return x
x,y =3,2
z = f(x)
print("z=", z)
print("y=", y)
print("x=", x)
def f(x):
def g():
x="abc"
print("x=", x)
def h():
z = x
print("z=", z)
x += 1
print("x=", x)
h()
g()
print("x=", x)
return g
x = 3
z = f(x)
print("x=", x)
print("z=", z)
z()
# ## Specification
#
# +
def findRoot(x, power, epsilon):
"""
Objective: Find Root
Arguments:
x:: Int or Float
epsilon:: Int or Float, epsilon >0
power:: Int , power >=1
Return:
y: Float
y satisfies y**power in [x-ϵ,x+ϵ].
If there does not exist such number, return null.
"""
if x < 0.0 and power%2 == 0: #negative number does not have root
return None
low = min(-1.0,x)
high = max(1.0,x)
ans =0.5*(low+high)
while abs(ans**power - x) >= epsilon:
if ans**power < x:
low = ans
else:
high = ans
ans = 0.5*(low+high)
return ans
# -
def testFindRoot():
"""
This function check whether findRoot function works well
"""
epsilon = 1e-3
for x in (0.25,-0.25,2,-2,8,-8):
for power in range(1,5):
print("Testing x =", str(x), "and power", power)
result = findRoot(x,power,epsilon)
if result == None:
print("No root")
else:
print("", result**power, "~=", x)
testFindRoot()
# ## Recursion
def factR(n):
"""
Argument:
n: Int, n >0
return;
n!
"""
if n == 1:
return n
else:
return n*factR(n-1)
factR(1)
factR(5)
def fib(n):
"""
Argument:
n: Int, n>0
"""
if n ==0 or n ==1:
return 1
else:
return fib(n-1) + fib(n-2)
def testfib(n):
for i in range(n+1):
print("fib of ", i , "=", fib(i))
testfib(5)
def isPalindrome(s):
"""
Argument:
s: strings
Return;
if x is Palindrome, True
else False
"""
def toChars(s):
#すべての文字を小文字にして文字列を作る
s = s.lower()
letters = ""
for c in s:
if c in "abcdefghijklmnopqrstuvwxyz":
letters += c
return letters
def isPal(s):
if len(s)<= 1:
return True
else:
#最初と最後の文字が同じかをチェック チェックしたら中心にうつる
return s[0]== s[-1] and isPal(s[1:-1])
return isPal(toChars(s))
x = "Abccba"
isPalindrome(x)
x = "sample"
isPalindrome(x)
# ## Global variable
def fib(n):
"""
Argument:
n: Int, n>0
"""
global numFibCalls
numFibCalls += 1
if n ==0 or n ==1:
return 1
else:
return fib(n-1) + fib(n-2)
def testfib(n):
for i in range(n+1):
global numFibCalls
numFibCalls = 0
print("fib of", i, "=", fib(i))
print("fib called", numFibCalls,"times.")
testfib(5)
# ## Module
# As programs are larger and larger, it is convenient to decompose the problem and save them.
#
# Using module is a good way to solve the problem.
import circle
pi = 3
print(pi)
print(circle.pi)
print(circle.area(3))
print(circle.circumference(3))
print(circle.sphereSurface(3))
# ## File IO
# +
nameHandle = open("kids", "w") # create file which name is kids
# "w" means we open the file by write mode
for i in range(2):
name = input("Enter name:")
nameHandle.write(name+ "\n") # \nは改行を示す writeメソッドはファイルに書き込むことを示す
nameHandle.close()#close file
# +
nameHandle = open("kids","r") #読みこみモードでファイルを開く
for line in nameHandle:
print(line[:-1]) # not to read \n
nameHandle.close()
# +
nameHandle = open("kids","a") # append mode
nameHandle.write("michael\n")
nameHandle.close()
nameHandle = open("kids","r") #読みこみモードでファイルを開く
for line in nameHandle:
print(line[:-1]) # not to read \n
nameHandle.close()
# -
| chap4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import os
import re
directory = os.getcwd()
print(directory)
os.chdir('D:\Machine learning concrete\Tensile_compressive_concrete')
directory = os.getcwd()
print(directory)
df = pd.read_csv("artifacts/initial_prepared_data/init_data.csv")
df.head()
# Calculating percentage of missing values in data set
## replacing "-" with np.nan i.e., null for all columns
print("--"*40)
for i in df.columns:
df[i]=df[i].replace('-',np.nan)
percent = (df[i]).isnull().sum()/len(df[i])*100
print(f"Number of Null value in {i} are {percent.round(1)} % ")
print("--"*40)
def nw_val(string):
regg=r'-?\d+\.?\d*'
lst = [float(s) for s in re.findall(regg,string)]
avg = sum(lst)/len(lst)
return avg
# +
# x = df[df['Stone powder content in Sand (%)'].str.contains("~")==True].index
# x[0]
# +
# (df['Stone powder content in Sand (%)'].iloc[x[0]])
# +
# df.at[x[0],'Stone powder content in Sand (%)'] = (nw_val(df['Stone powder content in Sand (%)'].iloc[x[0]]))
# -
df = df.apply(pd.to_numeric, errors='ignore')
num_col = df.describe().columns
# +
df = df.apply(pd.to_numeric, errors='ignore')
num_col = df.describe().columns
other_feature = []
for feature in df.columns:
if feature not in num_col:
other_feature.append(feature)
print(other_feature)
for i in other_feature:
df[i]=df[i].replace(regex=['※'], value='')
df[i]=df[i].replace(regex=["#"], value='')
df[i]=df[i].replace(regex=['\*'], value='')
# -
####There are some vales with range
####Those values are replaced by average of it
for i in other_feature:
x = df[df[i].str.contains("~")==True].index
for j in x:
# print(df[i].iloc[j])
df.at[j,i]=nw_val(df[i].iloc[j])
# print(df[i].iloc[j])
# +
df = df.apply(pd.to_numeric, errors='ignore')
num_col = df.describe().columns
other_feature = []
for feature in df.columns:
if feature not in num_col:
other_feature.append(feature)
print(other_feature)
# -
df['Slump(mm)'].unique()
df["Slump(mm)"] = df["Slump(mm)"].replace("<10",5)
# +
df = df.apply(pd.to_numeric, errors='ignore')
num_col = df.describe().columns
other_feature = []
for feature in df.columns:
if feature not in num_col:
other_feature.append(feature)
print(other_feature)
# -
df = df.drop(columns='Source')
directory = os.getcwd()
print(directory)
os.chdir('D:\\Machine learning concrete\\Tensile_compressive_concrete\\artifacts\\initial_prepared_data')
directory = os.getcwd()
print(directory)
df.to_csv("Updated_init_data.csv")
# +
# df[df['Stone powder content in Sand (%)'].str.contains("~")==True].index
# +
# df['Fineness modulus of sand '].unique()
# +
# mask = df['Stone powder content in Sand (%)'].str.contains("~")==True
# print(mask.unique())
# column_name = 'Stone powder content in Sand (%)'
# print(df.loc[mask,column_name])
# +
# (df['Stone powder content in Sand (%)'].str.contains("~")==True).sum()
# +
# a = df['Stone powder content in Sand (%)'].unique()
# sum([float(s) for s in re.findall(r'-?\d+\.?\d*','11.2~13.0')])/len([float(s) for s in re.findall(r'-?\d+\.?\d*','11.2~13.0')])
| artifacts/Research_env/2. EDA on raw data(init_data).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/prathak/2048/blob/master/TVP_VAR_DA_algorithm_variational.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="daRjgM78mhb6" colab_type="code" colab={}
pip install --upgrade tensorflow==2.2.0rc4
# + id="G8Ss36kImhcC" colab_type="code" colab={}
pip install --upgrade tfp-nightly
# + id="3hapJlXOmhcK" colab_type="code" colab={}
#import packages
import random
import os
import numpy as np
import tensorflow_probability as tfp
import tensorflow as tf
import matplotlib.pyplot as plt
import time
# #%qtconsole
def run_DGP(periods,dimensions):
##Data Generating Process######
np.random.seed(1337)# this is the only part of the whole code which is stochastic
n_dim = dimensions #INPUT # number of dimensions #
t=np.array(periods) #INPUT #500
#try out different scales
Q=np.array(0.001) #0.001
R=np.array(0.01) #0.01
Q_test = 0.001
R_test = 0.01
#generate white noise series
e1 = np.random.normal(0,1,[t, n_dim])
#e1 = e1*Q
e1 = e1*Q
e2 = np.random.normal(0,1,[t, n_dim])
e2 = e2*R
e3 = np.random.normal(0,1,[t, n_dim])
e3 = e3*Q_test
#generate containers
Beta = np.zeros(shape=[t, n_dim])
Y= np.zeros(shape=[t, n_dim])
#generate X as N-(3,1) process
X = np.random.normal(3,1,size=[t, n_dim])
#generate state variable beta as random walk process, and Y as a function of this.
for i in np.arange(0,n_dim):
for j in np.arange(1,t).reshape(-1):
Beta[j][i] = Beta[j-1][i] + e2[j][i]
Y[j][i] = X[j][i]*Beta[j][i] + e1[j][i] +e3[j][i] #+e4[j][n_dim-1]
X_raw = X
Y_raw = Y
return(X_raw, Y_raw, Beta)
# if __name__ == "__main__":
# X,Y,Beta = run_DGP(periods=180,dimensions=3)
# if 1:
# #Now check artifical series
# #Create subplot
# fig1, ax = plt.subplots(2, 2, figsize = (15, 10))
# ax[0, 0].plot(Beta)
# ax[0, 0].set_title(r'Betas', fontsize = 15)
# ax[1, 0].plot(Y[:,1:2])
# ax[1, 0].set_title(r'Example Y', fontsize = 15)
# ax[0, 1].plot(X[:,1:2])
# ax[0, 1].set_title(r'Example X', fontsize = 15)
# ax[1, 1].plot(X)
# ax[1, 1].set_title(r'All X', fontsize = 15)
# plt.tight_layout()
# + id="G3vZ1SmumhcM" colab_type="code" colab={}
# creating x = I ⊗ X, Xt = [y t−1 0 ,...,y t−l ,1]
def get_x(time, lag_length, K, Y, X, raw_x = True):
# t × K
X1 = np.zeros(shape=[time,K])
# t x q × qK
x = np.zeros(shape=[time,n_dim,K*n_dim])
for t in range(lag_length+1,time):
lag_sh = 0
if(raw):
X1[t,lag_sh:lag_sh+n_dim] = X[t]
else:
start = t-1 if lag_length == 0 else (t-lag_length+1)
lag_var = X[start:t+1,:].flatten()
lag_sh = lag_var.shape[0]
X1[t,:lag_sh] = lag_var
X1[t,-1] = 1
x[t] = np.kron(np.eye(n_dim),X1[t].T)
return x
# + id="fqwbWHgomhcQ" colab_type="code" colab={}
class TVP_variational_optimization:
def __init__(self, n_dim, K , window, Q_0, beta_b, t, x, Y):
# n_dim x n_dim
sigma_bar = np.identity(n_dim)*0.05
self.sigma = tf.convert_to_tensor(sigma_bar, dtype=tf.float32)
self.Q = tf.convert_to_tensor(Q_0, dtype=tf.float32)
self.beta_b = tf.convert_to_tensor(beta_b, dtype=tf.float32)
self.beta = tf.Variable(np.zeros((n_dim*K, 1)), dtype=tf.float32)
self.n_dim = n_dim
self.X = tf.convert_to_tensor(x, dtype=tf.float32)
self.Y = tf.convert_to_tensor(Y, dtype=tf.float32)
self.K = K
self.t = t
self.window = window
self.optimizer = tf.keras.optimizers.Adam(lr=0.01)
def cost(self):
beta_b_beta = (self.beta_b - self.beta)
beta_b_beta_transpose = tf.transpose(beta_b_beta)
beta_b_beta_Q_0 = tf.tensordot(beta_b_beta_transpose, tf.linalg.inv(self.Q),axes = 1)
J_b = tf.tensordot(beta_b_beta_Q_0,beta_b_beta,axes = 1)
total_residual_cost = tf.zeros([self.n_dim,self.n_dim])
for i in range(self.t, self.t + self.window):
Y_new = tf.reshape(self.Y[i],[n_dim,1])
residual = Y_new - (self.X[i] @ self.beta)
residual_t = tf.transpose(residual)
residula_sigma = tf.tensordot(residual_t, tf.linalg.inv(self.sigma),axes = 1)
J_0 = tf.tensordot(residula_sigma,residual,axes = 1)
total_residual_cost += J_0
return (J_b + total_residual_cost)
def optimize_w(self):
losses = tfp.math.minimize(self.cost,
num_steps=300,
optimizer=self.optimizer)
return self.beta
# print("optimized value is {} with loss {}".format(self.beta, losses[-1]))
# + id="1vd4JzsEmhcS" colab_type="code" colab={} outputId="abff0927-ed2a-418e-9278-a31753297512"
time = 150
n_dim = 3
raw = True
window = 5
lag_length = 0
K = n_dim + 1
X,Y,Beta = run_DGP(periods=time,dimensions=n_dim)
x = get_x(time,lag_length,K, Y, X,raw_x = True)
Q_0 = np.eye(n_dim*K)*1.01
beta_b = np.eye(n_dim*K,1)*0.06
b = np.zeros((time,n_dim))
qK = n_dim*K
beta = np.zeros([time,qK,1])
for t in range(0,time,window) :
if(t%15 == 0):
print("At time window : " + str(t))
optimizer = TVP_variational_optimization(n_dim, K, window, Q_0, beta_b, t, x, Y)
optim = optimizer.optimize_w()
beta[t] = optim.numpy()
for tau in range(0,window):
beta[t+tau] = beta[t]
beta_b = beta[t]
Q_0 = Q_0 + np.identity(K*n_dim)
for i in range(0,time):
a = int(beta[t,:,0].shape[0]/n_dim)
b[t] = np.diag(np.reshape(beta[t,:,0],(n_dim,a)))
# + id="FvcW6MfkmhcW" colab_type="code" colab={} outputId="03f9a2d2-fec5-4b6a-9873-f5fe223cb940"
plt.plot(beta[:,0,:],linestyle = '--')
plt.plot(Beta[:,0])
plt.tight_layout()
# + id="lgKJKCICmhca" colab_type="code" colab={}
| TVP_VAR_DA_algorithm_variational.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Corona Data Scrapping & visualization for India
# Coronavirus disease (COVID-19) is an infectious disease caused by a newly discovered coronavirus.
#
# Most people infected with the COVID-19 virus will experience mild to moderate respiratory illness and recover without requiring special treatment. Older people, and those with underlying medical problems like cardiovascular disease, diabetes, chronic respiratory disease, and cancer are more likely to develop serious illness.
#
# The best way to prevent and slow down transmission is be well informed about the COVID-19 virus, the disease it causes and how it spreads. Protect yourself and others from infection by washing your hands or using an alcohol based rub frequently and not touching your face.
# ## Import Libraries
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import requests
from bs4 import BeautifulSoup
import geopandas as gpd
from prettytable import PrettyTable
# -
# ## Let's Scap the Data
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
url = 'https://www.mohfw.gov.in/'# make a GET request to fetch the raw HTML content
web_content = requests.get(url).content# parse the html content
soup = BeautifulSoup(web_content, "html.parser")# remove any newlines and extra spaces from left and right
extract_contents = lambda row: [x.text.replace('\n', '') for x in row]# find all table rows and data cells within
stats = []
all_rows = soup.find_all('tr')
for row in all_rows:
stat = extract_contents(row.find_all('td')) # notice that the data that we require is now a list of length 5
if len(stat) == 5:
stats.append(stat)#now convert the data into a pandas dataframe for further processingnew_cols = ["Sr.No", "States/UT","Confirmed","Recovered","Deceased"]
new_cols = ["Sr.No", "States/UT","Confirmed","Recovered","Deceased"]
state_data = pd.DataFrame(data = stats, columns = new_cols)
state_data.head()
#03-05-2020
# -
state_data.shape
# ## scraped data columns are actually of ‘string’ datatype. We need to convert them into ‘int’ datatype.
state_data['Confirmed'] = state_data['Confirmed'].map(int)
state_data['Recovered'] = state_data['Recovered'].map(int)
state_data['Deceased'] = state_data['Deceased'].map(int)
# ## Now we will use PrettyTable
table = PrettyTable()
table.field_names = (new_cols)
for i in stats:
table.add_row(i)
table.add_row(["","Total",
sum(state_data['Confirmed']),
sum(state_data['Recovered']),
sum(state_data['Deceased'])])
print(table)
# ## Let' start Visualization
# +
sns.set_style("ticks")
# plt.rcParams['axes.facecolor'] = 'black'
plt.figure(figsize = (17,12))
plt.barh(state_data["States/UT"], state_data["Confirmed"].map(int),
align = 'center', color = 'green', edgecolor = 'blue')
plt.xlabel('Number of Confirmed cases', fontsize = 18)
plt.ylabel('States/UT', fontsize = 18)
plt.gca().invert_yaxis() #to maintain the order
plt.xticks(fontsize = 14)
plt.yticks(fontsize = 14)
plt.title('Total Confirmed Cases Statewise', fontsize = 20)
plt.show()
# -
# ## Donut chart for analysis of all India Cases
# +
# donut chart representing nationwide total confirmed, cured and deceased cases
group_size = [sum(state_data['Confirmed']),
sum(state_data['Recovered']),
sum(state_data['Deceased'])]
group_labels = ['Confirmed\n' + str(sum(state_data['Confirmed'])),
'Recovered\n' + str(sum(state_data['Recovered'])),
'Deceased\n' + str(sum(state_data['Deceased']))]
custom_colors = ['skyblue','yellowgreen','tomato']
plt.figure(figsize = (5,5))
plt.pie(group_size, labels = group_labels, colors = custom_colors)
central_circle = plt.Circle((0,0), 0.5, color = 'white')
fig = plt.gcf()
fig.gca().add_artist(central_circle)
plt.rc('font', size = 12)
plt.title('Nationwide total Confirmed, Recovered and Deceased Cases', fontsize = 16)
plt.show()
# -
# ## Let's plot Pie chart with Plotly to see a Confirmed cases ratio
# After seen analysis, we can say that recovered ratio is more than death ratio. So it is good point for us
# +
import plotly.express as px
fig = px.pie(state_data, values = 'Confirmed',names='States/UT', height=600)
fig.update_traces(textposition='inside', textinfo='percent+label')
fig.update_layout(
title_x = 0.5,
geo=dict(
showframe = False,
showcoastlines = False,
))
fig.show()
# -
# We can see Maharashtra have many number of cases
# ## Let's see for Recovered cases ratio
# +
fig = px.pie(state_data, values = 'Deceased',names='States/UT', height=600)
fig.update_traces(textposition='inside', textinfo='percent+label')
fig.update_layout(
title_x = 0.5,
geo=dict(
showframe = False,
showcoastlines = False,
))
fig.show()
# -
# ## Let's see TreeMap for Confirmed Cases
# +
fig = px.treemap(state_data, path=['States/UT'], values='Confirmed', height=600, width=800)
fig.update_layout(
title_x = 0.5,
geo=dict(
showframe = False,
showcoastlines = False,
))
fig.show()
# -
# ## Let's see for recovered cases
# +
fig = px.treemap(state_data, path=['States/UT'], values='Recovered', height=600, width=800)
fig.update_layout(
title_x = 0.5,
geo=dict(
showframe = False,
showcoastlines = False,
))
fig.show()
# -
# ## Let's see for Deceased class
# +
fig = px.treemap(state_data, path=['States/UT'], values='Deceased', height=600, width=800)
fig.update_layout(
title_x = 0.5,
geo=dict(
showframe = False,
showcoastlines = False,
))
fig.show()
# +
import geopandas as gpd
import pandas as pd
# read the state wise shapefile of India in a GeoDataFrame and preview it
map_data = gpd.read_file('/kaggle/input/india-states/Igismap/Indian_States.shp')
map_data.rename(columns = {'st_nm':'States/UT'}, inplace = True)
map_data.head()
# correct the name of states in the map dataframe
map_data['States/UT'] = map_data['States/UT'].str.replace('&', 'and')
map_data['States/UT'].replace('Arunanchal Pradesh', 'Arunachal Pradesh', inplace = True)
map_data['States/UT'].replace('Telangana', 'Telengana', inplace = True)
map_data['States/UT'].replace('NCT of Delhi', 'Delhi', inplace = True)
# merge both the dataframes - state_data and map_data
merged_data = pd.merge(map_data, state_data, how = 'left', on = 'States/UT')
merged_data.fillna(0, inplace = True)
merged_data.drop('Sr.No', axis = 1, inplace = True)
# create figure and axes for Matplotlib and set the title
fig, ax = plt.subplots(1, figsize=(20, 12))
ax.axis('off')
ax.set_title('Covid-19 Statewise Data - Confirmed Cases', fontdict = {'fontsize': '25', 'fontweight' : '3'})
# plot the figure
merged_data.plot(column = 'Confirmed', cmap='YlOrRd', linewidth=0.8, ax=ax, edgecolor='0.8', legend = True)
plt.show()
plt.savefig('in.png')
# -
# #### Reference:- https://towardsdatascience.com/tracking-corona-covid-19-spread-in-india-using-python-40ef8ffa7e31
| Corona Data India/mohfw corona dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GEOG-503 Lab 2
#
# ## Due date:
# Sep 12 (Wednesday) 11:59 pm
#
# ## What to submit:
#
# The Jupyter Notebook that contains solutions to the following questions. File name convention for submission: ** lastname_firstname_lab2.ipynb **
#
# It must be submitted to [myCourses](https://mycourses.binghamton.edu/)
#
# ## Questions
#
# Answer the questions or complete the tasks outlined in bold below, use the specific method described if applicable.
# ## Q1:
#
# Write a program which prompts the user for hours and rate per hour to compute the gross pay. Your program must have error handling capabilites (*try ... except ...*) when a user enters invalid numbers/characters.
#
# gross-pay = rate-per-hour * hours
# +
#This function need improvement.
#How to raise an exception that a string can't convert to number
def grossPay(rate, hours):
try:
if float(rate) <= 0:
raise Exception('''The 'rate' must be postive numbers''')
if float(hours) <= 0:
raise Exception('''The 'hours' must be postive numbers''')
gross = float(rate)*float(hours)
return gross
except ValueError:
print('''The 'rate' and 'hours' must be numbers''')
rate = input('Please enter the rate:')
hours = input('Please enter the hours:')
try:
print("The gross-pay is:" + str(grossPay(rate, hours)))
except Exception as err:
print('An exception happened:' + str(err))
# -
# ## Q2:
#
# Write a program which prompts the user for a Fahrenheit temperature, convert the temperature to Celsius, and print out the converted temperature. Your program must have error handling capabilites (*try ... except ...*) when a user enters invalid numbers/characters.
# +
def fahToCel(fahren): # The parameter needs to be numbers
celsius = (fahren - 32)*5/9
return celsius
intro = "This is a Function to covert Fahrenheit temperature to Celsius temperature"
print(intro.center(len(intro) + 10, '*'))
while True:
try:
fahrenheit = input('Please enter a Fahrenheit temperature:')
print('The Celsius temperature is:' + str(fahToCel(float(fahrenheit))) + '℃')
break
except ValueError:
print('The Fahrenheit temperature must be numbers, please enter again!')
# -
# ## Q3:
#
# Write a program that prompts a user for entering a grade (0 -100) and whether he/she has earned the extra credit (‘Yes’ or ‘No’). If ‘Yes’, add 3 extra credits to his/her grade, then print out the letter grade he/she should receive based on the grading scheme below.
#
# Letter Grade % of Points
#
# A 93.00 – 100%
#
# A- 90.00 – 92.99%
#
# B+ 87.00 – 89.99%
#
# B 82.00 – 86.99%
#
# B- 80.00 – 81.99%
#
# C+ 77.00 – 79.99%
#
# C 72.00 – 76.99%
#
# C- 70.00 – 71.99%
#
# D 60.00 – 69.99%
#
# F 0.00 – 59.99%
#
# +
def gradeIndentify(grades, extra): # grades must be number, extra must be string
extraPoints = 3
if extra.lower() == "yes":
grades = extraPoints + grades
if grades >= 93:
return "A"
elif grades >= 90:
return "A-"
elif grades >= 87:
return "B+"
elif grades >= 82:
return "B"
elif grades >= 80:
return "B-"
elif grades >= 77:
return "C+"
elif grades >= 72:
return "C"
elif grades >= 70:
return "C-"
elif grades >= 60:
return "D"
else:
return "F"
intro = "This is a function to calculate your grades"
print(intro.center(len(intro) + 30, '*'))
grades = input('''Please enter your grade(The grade must be numbers):''')
extra = input('''Please choose whether you earn the extra credit(YES/NO):''')
print('Your Letter grade is:' + gradeIndentify(float(grades), extra))
# -
# ## Q4:
#
# Write a program that repeatedly prompts a user for numbers until the user enters 'done'. Once 'done' is entered, print out the largest, smallest, sum, and average of the numbers. If the user enters anything other than a valid number put out an appropriate message and ignore the number.
#
intro = "This is a function to get the information of a series of numbers"
print(intro.center(len(intro) + 30, '*'))
nums = []
while True:
num = input('''Please enter a number(enter the 'done' to stop):''')
if num.lower() == 'done':
break
try:
num = float(num)
except ValueError:
print('You enter the wrong information, please enter a number again')
continue
nums.append(num)
print(nums)
info = {"largest number": max(nums), "smallest number": min(nums), "sum": sum(nums), "average": sum(nums)/len(nums)}
for k, v in info.items():
print('The {} of the numbers is:{}'.format(k, v))
# ## Q5:
#
# Write a program to find out the names listed in **names_1.csv** but not in **names_2.csv**. Save the results as a new csv named **name_not_found.csv**.
# +
import os, csv
intro = "This is a function to get the information in csv1 but not in csv2"
print(intro.center(len(intro) + 30, '*'))
wd = os.getcwd()
os.makedirs('generatedData', exist_ok=True) #generate a folder to store outcome
name1Path = os.path.join(wd, "data/names_1.csv")
name1File = open(name1Path)
name1Reader = csv.reader(name1File)
name2Path = os.path.join(wd, "data/names_2.csv")
name2File = open(name2Path)
name2Reader = csv.reader(name2File)
name1Rows = []
name2Rows = []
nameNewRows = []
for row in name1Reader:
name1Rows.append(row)
for row in name2Reader:
name2Rows.append(row)
for i in range(1, len(name1Rows)):
point = 0
for j in range(1, len(name2Rows)):
if name1Rows[i][-1] == name2Rows[j][-1]:
break
point += 1
if point < len(name2Rows) - 1:
continue
nameNewRows.append(name1Rows[i])
nameNewFile = open("generatedData/nameNew.csv", 'w', newline='')
nameWriter = csv.writer(nameNewFile)
for row in nameNewRows:
nameWriter.writerow(row)
nameNewFile.close()
if os.path.isfile("generatedData/nameNew.csv"):
print('The outcome has been generated, please check the generatedData folder in current directory.')
| GEOG503/labs/lab2/Xu_Jian_lab2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # Highly Performant TensorFlow Batch Inference on TFRecord Data Using the SageMaker CLI
#
# In this notebook, we'll show how to use SageMaker batch transform to get inferences on a large datasets. To do this, we'll use a TensorFlow Serving model to do batch inference on a large dataset of images encoded in TFRecord format, using the AWS command-line interface. We'll show how to use the new pre-processing and post-processing feature of the TensorFlow Serving container on Amazon SageMaker so that your TensorFlow model can make inferences directly on data in S3, and save post-processed inferences to S3.
#
# The dataset we'll be using is the [“Challenge 2018/2019"](https://github.com/cvdfoundation/open-images-dataset#download-the-open-images-challenge-28182019-test-set)” subset of the [Open Images V5 Dataset](https://storage.googleapis.com/openimages/web/index.html). This subset consists of 100,00 images in .jpg format, for a total of 10GB. For demonstration, the [model](https://github.com/tensorflow/models/tree/master/official/resnet#pre-trained-model) we'll be using is an image classification model based on the ResNet-50 architecture that has been trained on the ImageNet dataset, and which has been exported as a TensorFlow SavedModel.
#
# We will use this model to predict the class that each model belongs to. We'll write a pre- and post-processing script and package the script with our TensorFlow SavedModel, and demonstrate how to get inferences on large datasets with SageMaker batch transform quickly, efficiently, and at scale, on GPU-accelerated instances.
# ## Setup
#
# We'll begin with some necessary imports, and get an Amazon SageMaker session to help perform certain tasks, as well as an IAM role with the necessary permissions.
# +
import numpy as np
import os
import sagemaker
from sagemaker import get_execution_role
sagemaker_session = sagemaker.Session()
role = get_execution_role()
region = sagemaker_session.boto_region_name
bucket = sagemaker_session.default_bucket()
prefix = 'sagemaker/DEMO-tf-batch-inference-jpeg-images-python-sdk'
print('Region: {}'.format(region))
print('S3 URI: s3://{}/{}'.format(bucket, prefix))
print('Role: {}'.format(role))
# -
# ## Inspecting the SavedModel
# In order to make inferences, we'll have to preprocess our image data in S3 to match the serving signature of our TensorFlow SavedModel (https://www.tensorflow.org/guide/saved_model), which we can inspect using the saved_model_cli (https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/tools/saved_model_cli.py). This is the serving signature of the ResNet-50 v2 (NCHW, JPEG) (https://github.com/tensorflow/models/tree/master/official/resnet#pre-trained-model) model:
# !aws s3 cp s3://sagemaker-sample-data-{region}/batch-transform/open-images/model/resnet_v2_fp32_savedmodel_NCHW_jpg.tar.gz .
# !tar -zxf resnet_v2_fp32_savedmodel_NCHW_jpg.tar.gz
# !saved_model_cli show --dir resnet_v2_fp32_savedmodel_NCHW_jpg/1538687370/ --all
# The SageMaker TensorFlow Serving Container uses the model’s SignatureDef named serving_default , which is declared when the TensorFlow SavedModel is exported. This SignatureDef says that the model accepts a string of arbitrary length as input, and responds with classes and their probabilities. With our image classification model, the input string will be a base-64 encoded string representing a JPEG image, which our SavedModel will decode.
# ## Writing a pre- and post-processing script
#
# We will package up our SavedModel with a Python script named `inference.py`, which will pre-process input data going from S3 to our TensorFlow Serving model, and post-process output data before it is saved back to S3:
# !pygmentize code/inference.py
# Additionally, we add a `requirements.txt` file, which contains additional dependencies to install from the Python Package Index:
# !cat code/requirements.txt
# The input_handler intercepts inference requests, base-64 encodes the request body, and formats the request body to conform to TensorFlow Serving’s REST API (https://www.tensorflow.org/tfx/serving/api_rest). The return value of the input_handler function is used as the request body in the TensorFlow Serving request.
#
# Binary data must use key "b64", according to the TFS REST API (https://www.tensorflow.org/tfx/serving/api_rest#encoding_binary_values), and since our serving signature’s input tensor has the suffix "\_bytes", the encoded image data under key "b64" will be passed to the "image\_bytes" tensor. Some serving signatures may accept a tensor of floats or integers instead of a base-64 encoded string, but for binary data (including image data), it is recommended that your SavedModel accept a base-64 encoded string for binary data, since JSON representations of binary data can be large.
# Each incoming request originally contains a serialized JPEG image in its request body, and after passing through the input_handler, the request body contains the following, which our TensorFlow Serving accepts for inference:
#
# `{"instances": [{"b64":"[base-64 encoded JPEG image]"}]}`
# The first field in the return value of `output_handler` is what SageMaker Batch Transform will save to S3 as this example’s prediction. In this case, our `output_handler` passes the content on to S3 unmodified.
#
# Pre- and post-processing functions let you perform inference with TensorFlow Serving on any data format, not just images. To learn more about the `input_handler` and `output_handler`, consult the SageMaker TensorFlow Serving Container README (https://github.com/aws/sagemaker-tensorflow-serving-container/blob/master/README.md).
# ## Packaging a Model
# After writing a pre- and post-processing script, you’ll need to package your TensorFlow SavedModel along with your script into a `model.tar.gz` file, which we’ll upload to S3 for the SageMaker TensorFlow Serving Container to use. Let's package the SavedModel with the `inference.py` script and examine the expected format of the `model.tar.gz` file:
# !tar -cvzf model.tar.gz code --directory=resnet_v2_fp32_savedmodel_NCHW_jpg 1538687370
# `1538687370` refers to the model version number of the SavedModel, and this directory contains our SavedModel artifacts. The code directory contains our pre- and post-processing script, which must be named `inference.py`. I can also include an optional `requirements.txt` file, which is used to install dependencies with `pip` from the Python Package Index before the Transform Job starts, but we don’t need any additional dependencies in this case, so we don't include a requirements file.
# We will use this `model.tar.gz` when we create a SageMaker Model, which we will use to run Transform Jobs. To learn more about packaging a model, you can consult the SageMaker TensorFlow Serving Container [README](https://github.com/aws/sagemaker-tensorflow-serving-container/blob/master/README.md).
# ## Run a Batch Transform job
#
# Next, we'll run a Batch Transform job using our data processing script and GPU-based Amazon SageMaker Model. More specifically, we'll perform inference on a cluster of two instances, though we can choose more or fewer. The objects in the S3 path will be distributed between the instances.
# Before we create a Transform Job, let's inspect some of our input data. Here's an example, the first image in our dataset. We can inspect the format of each TFRecord file. The first record in the object named "train-00001-of-00100" refers to object "785877fb88018e89.jpg":
#
# <img src="sample_image/785877fb88018e89.jpg">
# The data in the input path consists of 100 TFRecord files, each with 1,000 JPEG images of varying sizes and shapes. Here is a subset:
# !aws s3 ls s3://sagemaker-sample-data-{region}/batch-transform/open-images/tfrecord/ --human-readable
# ### Creating a Model and Running a Transform Job
#
# The code below creates a SageMaker Model entity that will be used for Batch inference, and runs a Transform Job using that Model. The Model contains a reference to the TFS container, and the `model.tar.gz` containing our TensorFlow SavedModel and the pre- and post-processing `inference.py` script.
#
# After we create a SageMaker Model, we use it to run batch predictions using Batch Transform. We specify the input S3 data, content type of the input data, the output S3 data, and instance type and count.
#
# ### Performance
#
# For improved performance, we specify two additional parameters `max_concurrent_transforms` and `max_payload`, which control the maximum number of parallel requests that can be sent to each instance in a transform job at a time, and the maximum size of each request body.
#
# When performing inference on entire S3 objects that cannot be split by newline characters, such as images, it is recommended that you set `max_payload` to be slightly larger than the largest S3 object in your dataset, and that you experiment with the `max_concurrent_transforms` parameter in powers of two to find a value that maximizes throughput for your model. For example, we’ve set `max_concurrent_transforms` to 64 after experimenting with powers of two, and we set `max_payload` to 1, since the largest object in our S3 input is less than one megabyte.
#
# In addition to performance parameters, we specify AssembleWith to be “Line”, to instruct our Transform Job to assemble the individual predictions in each object by newline characters rather than concatenating them.
#
# Furthermore, we specify certain environment variables, which are passed to the TensorFlow Serving Container, and are used to enable request batching, a TensorFlow Serving feature that allows records from multiple requests be batched together. When carefully configured, this can improve throughput, especially with GPU-accelerated inference. You can learn more about the request batching environment variables in the [SageMaker TensorFlow Serving Container repository](https://github.com/aws/sagemaker-tensorflow-serving-container#enabling-batching).
# + magic_args="-s \"$bucket\" \"$prefix\" \"$role\" \"$region\"" language="bash"
# # For convenience, we pass in bucket, prefix, role, and region set in first Python set-up cell
#
# BUCKET=$1
# PREFIX=$2
# ROLE_ARN=$3
# REGION=$4
#
# timestamp() {
# date +%Y-%m-%d-%H-%M-%S
# }
#
# # Creating the SageMaker Model:
# MODEL_NAME="image-classification-tfs-$(timestamp)"
# MODEL_DATA_URL="s3://$BUCKET/$PREFIX/model/tfrecord/model.tar.gz"
#
# aws s3 cp model.tar.gz $MODEL_DATA_URL
#
# # This image is maintained at https://github.com/aws/sagemaker-tensorflow-serving-container
# TFS_VERSION="1.13"
# PROCESSOR_TYPE="gpu"
# IMAGE="520713654638.dkr.ecr.$REGION.amazonaws.com/sagemaker-tensorflow-serving:$TFS_VERSION-$PROCESSOR_TYPE"
#
# aws sagemaker create-model \
# --model-name $MODEL_NAME \
# --primary-container Image=$IMAGE,ModelDataUrl=$MODEL_DATA_URL \
# --execution-role-arn $ROLE_ARN
#
# # Creating the Transform Job:
# TRANSFORM_JOB_NAME="tfs-image-classification-job-$(timestamp)"
#
# # Specify where to get input data and where to get output data:
# TRANSFORM_S3_INPUT="s3://sagemaker-sample-data-$REGION/batch-transform/open-images/tfrecord"
# TRANSFORM_S3_OUTPUT="s3://$BUCKET/$PREFIX/output"
#
# # This configures Batch to split TFRecord files into individual records for each request
# # Other options for SPLIT_TYPE include "Line" to split by newline character, and "MultiRecord"
# # for BATCH_STRATEGY to include multiple records per request.
# # We choose "SingleRecord" so that our own pre-processing code doesn't have to manually split TFRecords.
# SPLIT_TYPE="TFRecord"
# BATCH_STRATEGY="SingleRecord"
#
# # Join outputs by newline characters. This will make JSONLines output, since each output is JSON.
# ASSEMBLE_WITH="Line"
#
# # The Data Source tells Batch to get all objects under the S3 prefix.
# TRANSFORM_INPUT_DATA_SOURCE={S3DataSource={S3DataType="S3Prefix",S3Uri=$TRANSFORM_S3_INPUT}}
# CONTENT_TYPE="application/x-tfexample"
# DATA_SOURCE=$TRANSFORM_INPUT_DATA_SOURCE,ContentType=$CONTENT_TYPE,SplitType=$SPLIT_TYPE
#
# # Specify resources used to transform the job
# INSTANCE_TYPE="ml.p3.2xlarge"
# INSTANCE_COUNT=2
#
# # Performance parameters. MaxPayloadInMB specifies how large each request body can be.
# # Our images happen to be less than 1MB, so we set MaxPayloadInMB to 1MB.
# # MaxConcurrentTransforms configures the number of concurrent requests made to the container at once.
# # The ideal number depends on the payload size, instance type, and model, so some experimentation
# # may be beneficial.
# MAX_PAYLOAD_IN_MB=1
# MAX_CONCURRENT_TRANSFORMS=64
# ENVIRONMENT=SAGEMAKER_TFS_ENABLE_BATCHING="true",SAGEMAKER_TFS_BATCH_TIMEOUT_MICROS="50000",SAGEMAKER_TFS_MAX_BATCH_SIZE="16"
#
# aws sagemaker create-transform-job \
# --model-name $MODEL_NAME \
# --transform-input DataSource=$DATA_SOURCE \
# --batch-strategy $BATCH_STRATEGY \
# --transform-output S3OutputPath=$TRANSFORM_S3_OUTPUT,AssembleWith=$ASSEMBLE_WITH \
# --transform-resources InstanceType=$INSTANCE_TYPE,InstanceCount=$INSTANCE_COUNT \
# --max-payload-in-mb $MAX_PAYLOAD_IN_MB \
# --max-concurrent-transforms $MAX_CONCURRENT_TRANSFORMS \
# --transform-job-name $TRANSFORM_JOB_NAME \
# --environment $ENVIRONMENT
#
# echo "Model name: $MODEL_NAME"
# echo "Transform job name: $TRANSFORM_JOB_NAME"
# echo "Transform job input path: $TRANSFORM_S3_INPUT"
# echo "Transform job output path: $TRANSFORM_S3_OUTPUT"
#
# # Wait for the transform job to finish.
# aws sagemaker wait transform-job-completed-or-stopped \
# --transform-job-name $TRANSFORM_JOB_NAME
#
# # Examine the output.
# aws s3 ls $TRANSFORM_S3_OUTPUT --human-readable
#
# # Copy an output example locally.
# aws s3 cp $TRANSFORM_S3_OUTPUT/train-00001-of-00100.out .
# -
# We see that after our transform job finishes, we find one S3 object in the output path for each object in the input path. This object contains the inferences from our model for that object, and has the same name as the corresponding input object, but with `.out` appended to it.
#
# Inspecting one of the output objects, we find the prediction from our TensorFlow Serving model. This is from the example image displayed above:
# !head -n 1 train-00001-of-00100.out
# ## Conclusion
#
# SageMaker batch transform can transform large datasets quickly and scalably. We used the SageMaker TensorFlow Serving Container to demonstrate how to quickly get inferences on a hundred thousand images using GPU-accelerated instances.
#
# The Amazon SageMaker TFS container supports CSV and JSON data out of the box. The pre- and post-processing feature of the container lets you run transform jobs on data of any format. The same container can be used for real-time inference as well using an Amazon SageMaker hosted model endpoint.
| sagemaker_batch_transform/tensorflow_open-images_tfrecord/tensorflow-serving-tfrecord.cli.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from typeguard import check_argument_types, check_return_type
def some_function(a: int, b: float, c: str, *args: str) -> bool:
retval = "Value"
assert check_argument_types()
assert check_return_type(retval)
return retval
# +
# # !pip install typeguard --upgrade
# -
some_function(10,1., "c")
import typeguard
typeguard.check_argument_types
| dev_nb/type_checking_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Step 1 - Prepare transcript2gene map
#
# ## Introduction
# The developmental forebrain dataset contain 16 samples, in our data preparation step, we used salmon to quantify transcript counts and the results were in 16 separate directories. The salmon quant results are in transcript level, however, for further DEG analysis, we would merge the result into gene level .
#
# ## Things I do below
# 1. I checked the content of one sample, to know the structure of the data
# 2. I checked the content of our GENCODE vm24 GTF file, the same gene annotation file used in salmon index. We will keep use this file throughout the analysis. In a separate notebook, I will display more way of manipulating and extract informations from the GTF.
# 3. I created a transcript to gene map csv file, which is needed in next notebook
# ## Load Package
import pandas as pd
import pathlib # deal with file paths
# ## Check out one sample first
#
# ### Load one sample quant table
# +
# use relative path to navigate to the salmon quants dir
# if this do not work, change to wherever you saved th salmon quants dirs.
data_dir = '../../../data/DevFB/quant/'
data_dir = pathlib.Path(data_dir)
data_dir # now data_dir is an Path obj, it has convenient functions to support path management
# +
# Let's check out this sample first
sample_id = 'forebrain_P0_1.quant'
sample_quant_path = data_dir / f'{sample_id}/quant.sf'
# +
quant_df = pd.read_csv(sample_quant_path, sep='\t',
index_col='Name')
# read_csv() can read tsv too, just use sep='\t'
# index_col tell the function which column can be row names.
# If index_col=None, the index will be default int index, you can try it yourself
print(quant_df.shape) # this table has (N_rows, N_columns)
quant_df.head()
# -
# ### Stop and read salmon's documentation
#
# Now that we have salmon's output table, we saw several columns. Although the name is kind of friendly, it's always good to fully understand what are they before we proceed.
#
# [Here](https://salmon.readthedocs.io/en/latest/file_formats.html) is salmon's documentation about their output format. I copied the column explaination here:
# - Name — This is the name of the target transcript provided in the input transcript database (FASTA file).
# - Length — This is the length of the target transcript in nucleotides.
# - EffectiveLength — This is the computed effective length of the target transcript. It takes into account all factors being modeled that will effect the probability of sampling fragments from this transcript, including the fragment length distribution and sequence-specific and gc-fragment bias (if they are being modeled).
# - TPM — This is salmon’s estimate of the relative abundance of this transcript in units of Transcripts Per Million (TPM). TPM is the recommended relative abundance measure to use for downstream analysis.
# - NumReads — This is salmon’s estimate of the number of reads mapping to each transcript that was quantified. It is an “estimate” insofar as it is the expected number of reads that have originated from each transcript given the structure of the uniquely mapping and multi-mapping reads and the relative abundance estimates for each transcript.
#
# Note: **TPM is an important term in RNA-seq**, everyone needs to know how it's calculated. If you don't, [check out this page and the video](https://www.rna-seqblog.com/rpkm-fpkm-and-tpm-clearly-explained/)
#
# ## Now let's focus on the index
#
# ### Important Note
# Salmon generate transcript-level quantification, the index of the above quant dataframe are transcript ids of mouse genes, we need to create a dict to annotate their corresponding gene ids. To do so, we need the gtf file we used in creating salmon index.
#
# ### More about IDs
#
# In database like GENCODE/ensembl or NCBI, gene, transcripts and protein all has different IDs.
#
# GENCODE and ensembl share the same ID pattern, which is also recommend to use in RNA-seq analysis for human and mouse. But you will also see NCBI RefSeq ID a lot.
#
# Use mouse ensembl/GENCODE as example:
# - ENSMUST... is transcript ID
# - ENSMUSG... is gene ID
#
# For human ensembl/GENCODE:
# - ENST... is transcript ID
# - ENSG... is gene ID
# ## Create transcript2gene dict using GTF file
# gtf table has some spetial formats, we need to set more parameters when read it in,
# here I just provide you the answer, you can search pandas.read_csv() documentation for more information
gtf = pd.read_csv(
'../../../data/ref/GENCODEvM24/gencode.vM24.annotation.gtf.gz',
comment='#',
sep='\t',
header=None,
names=[
'chrom', 'source', 'feature', 'start', 'end', 'na1', 'strand', 'na2',
'annotation'
])
gtf.head()
# gtf contain different type of features
gtf['feature'].value_counts()
# here we only need those transcript rows, so we do a filter here
# the transcript_gtf has 142552, its similar to our salmon quant table, which is a little bit less,
# probably some transcripts are unmappable by salmon.
transcript_gtf = gtf[gtf['feature'] == 'transcript'].copy()
print(transcript_gtf.shape)
transcript_gtf.head()
# ### Extract gene_id from annotation column
# the last column contains long annotation string, which you can see the gene_id is there
example_annotation = transcript_gtf.iloc[0, -1]
example_annotation
# +
# we need to extract the gene_id from this string. here is how I do this with a function
def extract_gene_id(annotation):
kv_pairs = annotation.split(';') # split into key-value pairs
for kv_pair in kv_pairs: # iterate the key-value pairs
kv_pair = kv_pair.strip(' ') # strip removed the space
if kv_pair.startswith('gene_id'):
_, gene_id = kv_pair.split(' ')
gene_id = gene_id.strip('"') # strip removed the ""
break
return gene_id
extract_gene_id(example_annotation)
# -
# now we need to apply this function on to each row of the transcript_gtf
# there are two way to do this, I first show you the fast (i.e. correct) way:
gene_ids = transcript_gtf['annotation'].apply(extract_gene_id)
# it take ~250ms in my computer
# search pandas.Dataframe.apply() see more about this function
gene_ids
# + deletable=false editable=false run_control={"frozen": true}
# # this is the slow way to achieve the same purpose,
# # I just want to demo that using correct pandas functions/methods can simplify your code,
# # and sometimes even make it faster
# # DO NOT USE THIS, USE THE ABOVE ONE
#
# # this is iterate the table and take the ann
# _gene_ids = []
# for annotation in transcript_gtf['annotation']:
# _gene_ids.append(extract_gene_id(annotation))
# _gene_ids = pd.Series(_gene_ids, index=transcript_gtf.index)
#
# _gene_ids
#
# # the result is the same, but here we need to write more
# -
# Now lets add gene_ids back to transcript_gtf
transcript_gtf['gene_ids'] = gene_ids
print(transcript_gtf.shape)
transcript_gtf.head()
# ### Extract transcript_id from annotation column
# +
## similarly we need to extract transcript_id from annotation too
# this time I will write a new function
def extract_transcript_id(annotation):
kv_pairs = annotation.split(';') # split into key-value pairs
for kv_pair in kv_pairs: # iterate the key-value pairs
kv_pair = kv_pair.strip(' ') # strip removed the space
if kv_pair.startswith('transcript_id'):
_, transcript_id = kv_pair.split(' ')
transcript_id = transcript_id.strip('"') # strip removed the ""
break
return transcript_id
extract_transcript_id(example_annotation)
# -
transcript_ids = transcript_gtf['annotation'].apply(extract_transcript_id)
transcript_gtf['transcript_id'] = transcript_ids
print(transcript_gtf.shape)
transcript_gtf.head()
# ### Save transcript2gene.csv
# subset the table give us the transcript to gene map needed in next notebook
tx2gene = transcript_gtf[['transcript_id', 'gene_ids']]
tx2gene
tx2gene.to_csv('tx2gene.csv', index=None)
# set index=None, which means do not export the index into csv, so the output file only has 2 columns
# !head tx2gene.csv
| analysis/DevFBProject/AggSalmon/Step 1 - Prepare transcript2gene map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Double 7's Portfolio
#
# (Optimize period and weighting scheme)
# +
import datetime
import matplotlib.pyplot as plt
import pandas as pd
from talib.abstract import *
import pinkfish as pf
import strategy
# Format price data.
pd.options.display.float_format = '{:0.2f}'.format
# %matplotlib inline
# -
# Set size of inline plots
'''note: rcParams can't be in same cell as import matplotlib
or %matplotlib inline
%matplotlib notebook: will lead to interactive plots embedded within
the notebook, you can zoom and resize the figure
%matplotlib inline: only draw static images in the notebook
'''
plt.rcParams["figure.figsize"] = (10, 7)
# Some global data
# +
# Symbol Lists
SP500_Sectors = ['XLB', 'XLE', 'XLF', 'XLI', 'XLK', 'XLP', 'XLU', 'XLV', 'XLY']
Mixed_Asset_Classes = ['IWB', 'SPY', 'VGK', 'IEV', 'EWJ', 'EPP', 'IEF', 'SHY', 'GLD']
FANG_Stocks = ['FB', 'AMZN', 'NFLX', 'GOOG']
Stocks_Bonds_Gold = ['SPY', 'QQQ', 'TLT', 'GLD']
# Pick one of the above
symbols = Stocks_Bonds_Gold
capital = 10000
start = datetime.datetime(1900, 1, 1)
end = datetime.datetime.now()
# -
options = {
'use_adj' : False,
'use_cache' : True,
'stop_loss_pct' : 0.15,
'margin' : 3,
'period' : 7,
'use_regime_filter' : True,
'use_vola_weight' : True
}
# Define Optimizations
# +
# pick one
optimize_period = True
optimize_use_vola_weight = False
optimize_stop_loss_pct = False
if optimize_period:
Xs = range(3, 18+1, 1)
Xs = [str(X) for X in Xs]
elif optimize_use_vola_weight:
Xs = (0, 1)
Xs = [str(X) for X in Xs]
elif optimize_stop_loss_pct:
Xs = range(5, 31, 1)
Xs = [str(X) for X in Xs]
# -
# Run Strategy
strategies = pd.Series(dtype=object)
for X in Xs:
print(X, end=" ")
if optimize_period:
options['period'] = int(X)
elif optimize_use_vola_weight:
options['use_vola_weight'] = False if X == '0' else True
elif optimize_stop_loss_pct:
options['stop_loss_pct'] = int(X)/100
strategies[X] = strategy.Strategy(symbols, capital, start, end, options)
strategies[X].run()
# Summarize results
# +
metrics = ('annual_return_rate',
'max_closed_out_drawdown',
'annualized_return_over_max_drawdown',
'best_month',
'worst_month',
'sharpe_ratio',
'sortino_ratio',
'monthly_std',
'pct_time_in_market',
'total_num_trades',
'pct_profitable_trades',
'avg_points')
df = pf.optimizer_summary(strategies, metrics)
df
# -
# Bar graphs
pf.optimizer_plot_bar_graph(df, 'annual_return_rate')
pf.optimizer_plot_bar_graph(df, 'sharpe_ratio')
pf.optimizer_plot_bar_graph(df, 'max_closed_out_drawdown')
# Run Benchmark
s = strategies[Xs[0]]
benchmark = pf.Benchmark('SPY', capital, s.start, s.end, use_adj=True)
benchmark.run()
# Equity curve
# +
if optimize_period: Y = '7'
elif optimize_use_vola_weight: Y = '1'
elif optimize_stop_loss_pct: Y = '15'
pf.plot_equity_curve(strategies[Y].dbal, benchmark=benchmark.dbal)
# +
labels = []
for strategy in strategies:
if optimize_period:
label = strategy.options['period']
elif optimize_use_vola_weight:
label = strategy.options['use_vola_weight']
elif optimize_stop_loss_pct:
label = strategy.options['stop_loss_pct']
labels.append(label)
pf.plot_equity_curves(strategies, labels)
# -
| examples/240.double-7s-portfolio/optimize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Setup
# +
# %matplotlib qt
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_gan as tfgan
import numpy as np
import os, sys
from tqdm.notebook import tqdm
from pathlib import Path
sys.path.append( os.path.abspath('..') )
import utils
# -
Path('MNIST').mkdir(exist_ok=True)
os.chdir('MNIST')
# +
(x_train, _), (x_test, _) = tf.keras.datasets.mnist.load_data()
data = np.concatenate((x_train, x_test))
data = (data.astype('float32') - 127.5) / 127.5 # normalize to [-1, 1]
data = np.expand_dims(data, axis=-1) # add channels dimension
assert data.shape == (70000, 28, 28, 1) # (batch, height, width, channel)
# -
# ## 1 Models
# ### 1.1 Architecture
def generator_model(latent_dims):
return tf.keras.Sequential([
tf.keras.layers.Dense(7*7*256, input_shape=(latent_dims,)),
tf.keras.layers.LeakyReLU(),
tf.keras.layers.Reshape((7, 7, 256)),
tf.keras.layers.Conv2D(128, kernel_size=5, strides=1, padding='same'),
tf.keras.layers.LeakyReLU(),
tf.keras.layers.UpSampling2D(size=2, interpolation='bilinear'),
tf.keras.layers.Conv2D(64, kernel_size=5, strides=1, padding='same'),
tf.keras.layers.LeakyReLU(),
tf.keras.layers.UpSampling2D(size=2, interpolation='bilinear'),
tf.keras.layers.Conv2D(1, kernel_size=5, strides=1, padding='same', activation='tanh')
])
def generator_model_bn(latent_dims):
return tf.keras.Sequential([
tf.keras.layers.Dense(7*7*256, input_shape=(latent_dims,)),
tf.keras.layers.LeakyReLU(),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Reshape((7, 7, 256)),
tf.keras.layers.Conv2D(128, kernel_size=5, strides=1, padding='same'),
tf.keras.layers.LeakyReLU(),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.UpSampling2D(size=2, interpolation='bilinear'),
tf.keras.layers.Conv2D(64, kernel_size=5, strides=1, padding='same'),
tf.keras.layers.LeakyReLU(),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.UpSampling2D(size=2, interpolation='bilinear'),
tf.keras.layers.Conv2D(1, kernel_size=5, strides=1, padding='same', activation='tanh')
])
def critic_model(constraint):
return tf.keras.Sequential([
tf.keras.layers.Conv2D(
64, kernel_size=5, strides=2, padding='same', input_shape=(28,28,1),
kernel_constraint=constraint, bias_constraint=constraint
),
tf.keras.layers.LeakyReLU(),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(
128, kernel_size=5, strides=2, padding='same',
kernel_constraint=constraint, bias_constraint=constraint
),
tf.keras.layers.LeakyReLU(),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(1, kernel_constraint=constraint, bias_constraint=constraint)
])
# ### 1.2 Losses
# For the Wasserstein metric, the loss for the generator (G) given the critic (f) is given by:
#
# $$
# -\mathbb{E}_{z \sim p(z)}\bigl\lbrack f\bigl(G(z)\bigr) \bigr\rbrack
# $$
def generator_loss(fake_output):
return -tf.reduce_mean(fake_output)
# The critic loss tries to maximize:
# $$
# \max_{\|f \|_{L} \leq 1} \mathbb{E}_{x \sim p_{data}} \bigl\lbrack f(x) \bigr\rbrack -
# \mathbb{E}_{z \sim p_{z}} \bigl\lbrack f\bigl(G(z)\bigr) \bigr\rbrack
# $$
#
# Which is equivalent to minimizing the negative of this value, as shown in the function bellow
def critic_loss(real_output, fake_output):
mu_real = tf.reduce_mean(real_output)
mu_fake = tf.reduce_mean(fake_output)
return mu_fake - mu_real
# ## 2 Training
# ### 2.1 Main functions
def critic_train_step(generator, critic, images, latent_dims):
noise = tf.random.normal([images.shape[0], latent_dims])
with tf.GradientTape() as crit_tape:
generated_imgs = generator(noise, training=True)
real_output = critic(images, training=True)
fake_output = critic(generated_imgs, training=True)
loss_C = critic_loss(real_output, fake_output)
grads_C = crit_tape.gradient(loss_C, critic.trainable_variables)
critic.optimizer.apply_gradients(zip(grads_C, critic.trainable_variables))
return loss_C
def generator_train_step(generator, critic, batch_size, latent_dims):
noise = tf.random.normal([batch_size, latent_dims])
with tf.GradientTape() as gen_tape:
generated_imgs = generator(noise, training=True)
fake_output = critic(generated_imgs, training=True)
loss_G = generator_loss(fake_output)
grads_G = gen_tape.gradient(loss_G, generator.trainable_variables)
generator.optimizer.apply_gradients(zip(grads_G, generator.trainable_variables))
return loss_G
def train(generator, critic, data, epochs, n_critic=1, batch_size=32, callbacks=None):
latent_dims = generator.input_shape[1]
dataset = tf.data.Dataset.from_tensor_slices(data).shuffle(data.shape[0]).batch(batch_size)
iterator = iter(dataset)
num_batches = 1 + (data.shape[0] - 1) // batch_size
batches_left = True
batch_count = 0
generator_step = tf.function(generator_train_step)
critic_step = tf.function(critic_train_step)
for epoch in tqdm(range(epochs)):
for c in callbacks:
c.on_epoch_begin(epoch=epoch + 1, generator=generator, discriminator=critic)
batch_pbar = tqdm(total=num_batches, leave=False)
while batches_left:
for i in range(n_critic):
if batch_count == num_batches:
batch_count = 0
batches_left = False
iterator = iter(dataset)
batch_count += 1
batch_pbar.update()
batch = iterator.get_next()
critic_step(generator, critic, batch, latent_dims)
generator_step(generator, critic, batch_size, latent_dims)
batches_left = True
batch_pbar.update(num_batches)
batch_pbar.close()
for c in callbacks:
c.on_epoch_end(epoch=epoch + 1, generator=generator, discriminator=critic)
# ### 2.2 Metrics classifier
# Loading the classifier that will be used to calculate the *Classifier Score* (CS) and *Fréchet Classifier Distance* (FCD). \
# The features of the real data are also precalculated to avoid doing that for each epoch.
classifier = tf.keras.models.load_model('../../Classifiers/mnist.h5')
feature_layer = classifier.get_layer('features')
logits_layer = classifier.get_layer('logits')
precalculated_features = utils.fn.calculate_features(classifier, feature_layer, data)
# ### 2.3 Hyperparameter Testing
# This class implements the *weight clipping* constraint used to guarantee the critic is a K-Lipschitz continuous function.
class ClipConstraint(tf.keras.constraints.Constraint):
def __init__(self, clip_value):
self.__clip_value = clip_value
def __call__(self, w):
return tf.clip_by_value(w, -self.__clip_value, self.__clip_value)
# These were the hyperparameters tested for the final document. Training all of them simultaneously may take a long time, consider commenting out some options to run the tests individually.
LATENT_DIMS = 24
BATCH_SIZE = 32
hparams_list = [
{'n_critic': 5, 'clip_value': 0.990, 'learning_rate': 5e-5, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 0.990, 'learning_rate': 1e-4, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 0.990, 'learning_rate': 5e-3, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 0.001, 'learning_rate': 1e-3, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 0.010, 'learning_rate': 1e-3, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 0.100, 'learning_rate': 1e-3, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 0.500, 'learning_rate': 1e-3, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 0.990, 'learning_rate': 1e-3, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 2.000, 'learning_rate': 1e-3, 'batch_norm': False},
{'n_critic': 1, 'clip_value': 0.990, 'learning_rate': 1e-3, 'batch_norm': False},
{'n_critic': 20, 'clip_value': 0.990, 'learning_rate': 1e-3, 'batch_norm': False},
{'n_critic': 5, 'clip_value': 0.990, 'learning_rate': 1e-4, 'batch_norm': True},
{'n_critic': 5, 'clip_value': 0.990, 'learning_rate': 5e-3, 'batch_norm': True},
{'n_critic': 5, 'clip_value': 0.100, 'learning_rate': 1e-3, 'batch_norm': True},
{'n_critic': 5, 'clip_value': 0.500, 'learning_rate': 1e-3, 'batch_norm': True},
{'n_critic': 5, 'clip_value': 0.990, 'learning_rate': 1e-3, 'batch_norm': True},
{'n_critic': 5, 'clip_value': 2.000, 'learning_rate': 1e-3, 'batch_norm': True},
{'n_critic': 1, 'clip_value': 0.990, 'learning_rate': 1e-3, 'batch_norm': True}
]
for hparams in hparams_list:
dirname = 'NCRIT{}_CLIP{}_LR{:.0e}{}'.format(
hparams['n_critic'],
hparams['clip_value'],
hparams['learning_rate'],
'_BN' if hparams['batch_norm'] else ''
)
Path(dirname).mkdir(exist_ok=True)
if hparams['batch_norm']:
generator = generator_model_bn(LATENT_DIMS)
else:
generator = generator_model(LATENT_DIMS)
# generator.optimizer = tf.keras.optimizers.Adam(hparams['learning_rate'], beta_1=0)
generator.optimizer = tf.keras.optimizers.RMSprop(hparams['learning_rate'])
constraint = ClipConstraint(hparams['clip_value'])
critic = critic_model(constraint)
# critic.optimizer = tf.keras.optimizers.Adam(hparams['learning_rate'], beta_1=0)
critic.optimizer = tf.keras.optimizers.RMSprop(hparams['learning_rate'])
## Callbacks
timer = utils.callback.TimerCallback()
save_samples = utils.callback.SaveSamplesCallback(
path_format=os.path.join(dirname, 'epoch-{}'),
inputs=tf.random.normal((10*10, LATENT_DIMS)),
n_cols=10,
savefig_kwargs={'bbox_inches': 'tight', 'pad_inches': 0, 'dpi': 192},
imshow_kwargs={'cmap': 'gray_r', 'vmin': -1, 'vmax': 1}
)
metrics = utils.callback.MetricsCallback(
generator=generator,
classifier=classifier,
latent_dims=LATENT_DIMS,
feature_layer=feature_layer,
logits_layer=logits_layer,
precalculated_features=precalculated_features,
save_after=5, save_to=os.path.join(dirname, 'best.h5')
)
## Train and save results
train(
generator, critic, data, epochs=30,
n_critic=hparams['n_critic'],
batch_size=BATCH_SIZE,
callbacks=[timer, save_samples, metrics]
)
metrics_obj = metrics.get_metrics()
metrics_obj['time'] = timer.get_time()
utils.fn.update_json_log(os.path.join(dirname, 'log.json'), metrics_obj)
generator.save(os.path.join(dirname, 'generator.h5'), overwrite=True, save_format='h5')
critic.save (os.path.join(dirname, 'critic.h5' ), overwrite=True, save_format='h5')
# \
# In windows the command bellow is used to turn down the machine after the training finishes, very useful if you wanna let the computer running while you go to sleep :)
# +
# # !shutdown /s /t 60
| WGAN/MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME>
#
# # NUMPY OPERATIONS
#
# Import Numpy Library
import numpy as np
import warnings
warnings.filterwarnings("ignore")
from IPython.display import Image
# # Numpy Array Creation
list1 = [10,20,30,40,50,60]
list1
# Display the type of an object
type(list1)
#Convert list to Numpy Array
arr1 = np.array(list1)
arr1
#Memory address of an array object
arr1.data
# Display type of an object
type(arr1)
#Datatype of array
arr1.dtype
# Convert Integer Array to FLOAT
arr1.astype(float)
# Generate evenly spaced numbers (space =1) between 0 to 10
np.arange(0,10)
# Generate numbers between 0 to 100 with a space of 10
np.arange(0,100,10)
# Generate numbers between 10 to 100 with a space of 10 in descending order
np.arange(100, 10, -10)
#Shape of Array
arr3 = np.arange(0,10)
arr3.shape
arr3
# Size of array
arr3.size
# Dimension
arr3.ndim
# Datatype of object
arr3.dtype
# Bytes consumed by one element of an array object
arr3.itemsize
# Bytes consumed by an array object
arr3.nbytes
# Length of array
len(arr3)
# Generate an array of zeros
np.zeros(10)
# Generate an array of ones with given shape
np.ones(10)
# Repeat 10 five times in an array
np.repeat(10,5)
# Repeat each element in array 'a' thrice
a= np.array([10,20,30])
np.repeat(a,3)
# Array of 10's
np.full(5,10)
# Generate array of Odd numbers
ar1 = np.arange(1,20)
ar1[ar1%2 ==1]
# Generate array of even numbers
ar1 = np.arange(1,20)
ar1[ar1%2 == 0]
# Generate evenly spaced 4 numbers between 10 to 20.
np.linspace(10,20,4)
# Generate evenly spaced 11 numbers between 10 to 20.
np.linspace(10,20,11)
# Create an array of random values
np.random.random(4)
# Generate an array of Random Integer numbers
np.random.randint(0,500,5)
# Generate an array of Random Integer numbers
np.random.randint(0,500,10)
# Using random.seed we can generate same number of Random numbers
np.random.seed(123)
np.random.randint(0,100,10)
# Using random.seed we can generate same number of Random numbers
np.random.seed(123)
np.random.randint(0,100,10)
# Using random.seed we can generate same number of Random numbers
np.random.seed(101)
np.random.randint(0,100,10)
# Using random.seed we can generate same number of Random numbers
np.random.seed(101)
np.random.randint(0,100,10)
# Generate array of Random float numbers
f1 = np.random.uniform(5,10, size=(10))
f1
# Extract Integer part
np.floor(f1)
# Truncate decimal part
np.trunc(f1)
# Convert Float Array to Integer array
f1.astype(int)
# Normal distribution (mean=0 and variance=1)
b2 =np.random.randn(10)
b2
arr1
# Enumerate for Numpy Arrays
for index, value in np.ndenumerate(arr1):
print(index, value)
# # Operations on an Array
arr2 = np.arange(1,20)
arr2
# Sum of all elements in an array
arr2.sum()
# Cumulative Sum
np.cumsum(arr2)
# Find Minimum number in an array
arr2.min()
# Find MAX number in an array
arr2.max()
# Find INDEX of Minimum number in an array
arr2.argmin()
# Find INDEX of MAX number in an array
arr2.argmax()
# Find mean of all numbers in an array
arr2.mean()
# Find median of all numbers present in arr2
np.median(arr2)
# Variance
np.var(arr2)
# Standard deviation
np.std(arr2)
# Calculating percentiles
np.percentile(arr2,70)
# 10th & 70th percentile
np.percentile(arr2,[10,70])
# # Operations on a 2D Array
A = np.array([[1,2,3,0] , [5,6,7,22] , [10 , 11 , 1 ,13] , [14,15,16,3]])
A
# SUM of all numbers in a 2D array
A.sum()
# MAX number in a 2D array
A.max()
# Minimum
A.min()
# Column wise mimimum value
np.amin(A, axis=0)
# Row wise mimimum value
np.amin(A, axis=1)
# Mean of all numbers in a 2D array
A.mean()
# Mean
np.mean(A)
# Median
np.median(A)
# 50 percentile = Median
np.percentile(A,50)
np.var(A)
np.std(A)
np.percentile(arr2,70)
# Enumerate for Numpy 2D Arrays
for index, value in np.ndenumerate(A):
print(index, value)
# # Reading elements of an array
a = np.array([7,5,3,9,0,2])
# Access first element of the array
a[0]
# Access all elements of Array except first one.
a[1:]
# Fetch 2nd , 3rd & 4th value from the Array
a[1:4]
# Get last element of the array
a[-1]
a[-3]
a[-6]
a[-3:-1]
# # Replace elements in array
ar = np.arange(1,20)
ar
# Replace EVEN numbers with ZERO
rep1 = np.where(ar % 2 == 0, 0 , ar)
print(rep1)
ar2 = np.array([10, 20 , 30 , 10 ,10 ,20, 20])
ar2
# Replace 10 with value 99
rep2 = np.where(ar2 == 10, 99 , ar2)
print(rep2)
p2 = np.arange(0,100,10)
p2
# Replace values at INDEX loc 0,3,5 with 33,55,99
np.put(p2, [0, 3 , 5], [33, 55, 99])
p2
# # Missing Values in an array
a = np.array([10 ,np.nan,20,30,60,np.nan,90,np.inf])
a
# Search for missing values and return as a boolean array
np.isnan(a)
# Index of missing values in an array
np.where(np.isnan(a))
# Replace all missing values with 99
a[np.isnan(a)] = 99
a
# Check if array has any NULL value
np.isnan(a).any()
A = np.array([[1,2,np.nan,4] , [np.nan,6,7,8] , [10 , np.nan , 12 ,13] , [14,15,16,17]])
A
# Search for missing values and return as a boolean array
np.isnan(A)
# Index of missing values in an array
np.where(np.isnan(A))
# # Stack Arrays Vertically
a = np.zeros(20).reshape(2,-1)
b = np.repeat(1, 20).reshape(2,-1)
a
b
np.vstack([a,b])
a1 = np.array([[1], [2], [3]])
b1 = np.array([[4], [5], [6]])
a1
b1
np.vstack([a1,b1])
# # Stack Arrays Horizontally
np.hstack([a,b])
np.hstack([a1,b1])
# +
### hstack & vstack
arr1 = np.array([[7,13,14],[18,10,17],[11,12,19]])
arr2= np.array([16,6,1])
arr3= np.array([[5,8,4,3]])
np.hstack((np.vstack((arr1,arr2)),np.transpose(arr3)))
# -
# # Common items between two Arrays
c1 = np.array([10,20,30,40,50,60])
c2 = np.array([12,20,33,40,55,60])
np.intersect1d(c1,c2)
# # Remove Common Elements
# +
# Remove common elements of C1 & C2 array from C1
np.setdiff1d(c1,c2)
# -
# # Process Elements on Conditions
# +
a = np.array([1,2,3,6,8])
b = np.array([10,2,30,60,8])
np.where(a == b) # returns the indices of elements in an input array where the given condition is satisfied.
# -
# Return an array where condition is satisfied
a[np.where(a == b)]
# Return all numbers betweeen 20 & 35
a1 = np.arange(0,60)
a1[np.where ((a1>20) & (a1<35))]
# Return all numbers betweeen 20 & 35 OR numbers divisible by 10
a1 = np.arange(0,60)
a1[np.where (((a1>20) & (a1<35)) | (a1 % 10 ==0)) ]
# Return all numbers betweeen 20 & 35 using np.logical_and
a1[np.where(np.logical_and(a1>20, a1<35))]
# # Check for elements in an Array using isin()
a = np.array([10,20,30,40,50,60,70])
a
# Check whether number 11 & 20 are present in an array
np.isin(a, [11,20])
#Display the matching numbers
a[np.isin(a,20)]
# Check whether number 33 is present in an array
np.isin(a, 33)
a[np.isin(a, 33)]
b = np.array([10,20,30,40,10,10,70,80,70,90])
b
# Check whether number 10 & 70 are present in an array
np.isin(b, [10,70])
# Display the indices where match occurred
np.where(np.isin(b, [10,70]))
# Display the matching values
b[np.where(np.isin(b, [10,70]))]
# Display the matching values
b[np.isin(b, [10,70])]
# # Reverse Array
a4 = np.arange(10,30)
a4
# Reverse the array
a4[::-1]
# Reverse the array
np.flip(a4)
a3 = np.array([[3,2,8,1] , [70,50,10,67] , [45,25,75,15] , [12,9,77,4]])
a3
# Reverse ROW positions
a3[::-1,]
# Reverse COLUMN positions
a3[:,::-1]
# Reverse both ROW & COLUMN positions
a3[::-1,::-1]
# # Sorting Array
a = np.array([10,5,2,22,12,92,17,33])
# Sort array in ascending order
np.sort(a)
a3 = np.array([[3,2,8,1] , [70,50,10,67] , [45,25,75,15]])
a3
# Sort along rows
np.sort(a3)
# Sort along rows
np.sort(a3,axis =1)
# Sort along columns
np.sort(a3,axis =0)
# Sort in descending order
b = np.sort(a)
b = b[::-1]
b
# Sort in descending order
c = np.sort(a)
np.flip(c)
# Sort in descending order
a[::-1].sort()
a
# # "N" Largest & Smallest Numbers in an Array
p = np.arange(0,50)
p
np.random.shuffle(p)
p
# Return "n" largest numbers in an Array
n = 4
p[np.argsort(p)[-nth:]]
# Return "n" largest numbers in an Array
p[np.argpartition(-p,n)[:n]]
# Return "n" smallest numbers in an Array
p[np.argsort(-p)[-n:]]
# Return "n" smallest numbers in an Array
p[np.argpartition(p,n)[:n]]
# # Repeating Sequences
a5 = [10,20,30]
a5
# Repeat whole array twice
np.tile(a5, 2)
# Repeat each element in an array thrice
np.repeat(a5, 3)
# # Compare Arrays
d1 = np.arange(0,10)
d1
d2 = np.arange(0,10)
d2
d3 = np.arange(10,20)
d3
d4 = d1[::-1]
d4
# Compare arrays using "allclose" function. If this function returns True then Arrays are equal
res1 = np.allclose(d1,d2)
res1
# Compare arrays using "allclose" function. If this function returns False then Arrays are not equal
res2 = np.allclose(d1,d3)
res2
# Compare arrays using "allclose" function.
res3 = np.allclose(d1,d4)
res3
# # Frequent Values in an Array
# unique numbers in an array
b = np.array([10,10,10,20,30,20,30,30,20,10,10,30,10])
np.unique(b)
# unique numbers in an array along with the count E.g value 10 occurred maximum times (5 times) in an array "b"
val , count = np.unique(b,return_counts=True)
val,count
# 10 is the most frequent value
np.bincount(b).argmax()
# # Read-Only Array
d5 = np.arange(10,100,10)
d5
# Make arrays immutable
d5.flags.writeable = False
d5[0] = 99
d5[2] = 11
# # Load & Save
# Load data from a text file using loadtext
p4 = np.loadtxt('sample.txt',
dtype = np.integer # Decides the datatype of resulting array
)
p4
# Load data from a text file using genfromtxt
p5 = np.genfromtxt('sample0.txt',dtype='str')
p5
# Accessing specific rows
p5[0]
# Accessing specific columns
p5[:,0]
p6 = np.genfromtxt('sample2.txt',
delimiter=' ',
dtype=None,
names=('Name', 'ID', 'Age')
)
p6
# Skip header using "skiprows" parameter
p6 = np.loadtxt('sample2.txt',
delimiter=' ',
dtype=[('Name', str, 50), ('ID', np.integer), ('Age', np.integer)],
skiprows=1
)
p6
# Return only first & third column using "usecols" parameter
np.loadtxt('sample.txt', delimiter =' ', usecols =(0, 2))
# Return only three rows using "max_rows" parameter
p6 = np.loadtxt('sample2.txt',
delimiter=' ',
dtype=[('Name', str, 50), ('ID', np.integer), ('Age', np.integer)],
skiprows=1,
max_rows = 3
)
p6
# Skip header using "skip_header" parameter
p6 = np.genfromtxt('sample2.txt',
delimiter=' ',
dtype=[('Name', str, 50), ('ID', np.integer), ('Age', np.float)],
names=('Name', 'ID', 'Age'),
skip_header=1
)
p6
p7 = np.arange(10,200,11)
p7
np.savetxt('test3.csv', p7, delimiter=',')
p8 = np.arange(0,121).reshape(11,11)
p8
np.save('test4.npy', p8)
p9 = np.load('test4.npy')
p9
np.save('numpyfile', p8)
p10 = np.load('numpyfile.npy')
p10
p11 = np.arange(0,1000000).reshape(1000,1000)
p11
# Save Numpy array to a compressed file
np.savez_compressed('test6.npz', p11)
# Save Numpy array to a npy file
np.save('test7.npy', p11)
# Compressed file size is much lesser than normal npy file
Image(filename='load_save.PNG')
# # Printing Options
# Display values upto 4 decimal place
np.set_printoptions(precision=4)
a = np.array([12.654398765 , 90.7864098354674])
a
# Display values upto 2 decimal place
np.set_printoptions(precision=2)
a = np.array([12.654398765 , 90.7864098354674])
a
# Array Summarization
np.set_printoptions(threshold=3)
np.arange(200)
# Reset Formatter
np.set_printoptions(precision=8,suppress=False, threshold=1000, formatter=None)
a = np.array([12.654398765 , 90.7864098354674])
a
np.arange(1,1100)
# Display all values
np.set_printoptions(threshold=np.inf)
np.arange(1,1100)
# # Vector Addition
v1 = np.array([1,2])
v2 = np.array([3,4])
v3 = v1+v2
v3 = np.add(v1,v2)
print('V3 =' ,v3)
# # Multiplication of vectors
a1 = [5 , 6 ,8]
a2 = [4, 7 , 9]
print(np.multiply(a1,a2))
# # Dot Product
# +
a1 = np.array([1,2,3])
a2 = np.array([4,5,6])
dotp = a1@a2
print(" Dot product - ",dotp)
dotp = np.dot(a1,a2)
print(" Dot product usign np.dot",dotp)
dotp = np.inner(a1,a2)
print(" Dot product usign np.inner", dotp)
dotp = sum(np.multiply(a1,a2))
print(" Dot product usign np.multiply & sum",dotp)
dotp = np.matmul(a1,a2)
print(" Dot product usign np.matmul",dotp)
dotp = 0
for i in range(len(a1)):
dotp = dotp + a1[i]*a2[i]
print(" Dot product usign for loop" , dotp)
# -
# # Length of Vector
v3 = np.array([1,2,3,4,5,6])
length = np.sqrt(np.dot(v3,v3))
length
v3 = np.array([1,2,3,4,5,6])
length = np.sqrt(sum(np.multiply(v3,v3)))
length
v3 = np.array([1,2,3,4,5,6])
length = np.sqrt(np.matmul(v3,v3))
length
# # Normalized Vector
#First Method
v1 = [2,3]
length_v1 = np.sqrt(np.dot(v1,v1))
norm_v1 = v1/length_v1
length_v1 , norm_v1
#Second Method
v1 = [2,3]
norm_v1 = v1/np.linalg.norm(v1)
norm_v1
# # Angle between vectors
#First Method
v1 = np.array([8,4])
v2 = np.array([-4,8])
ang = np.rad2deg(np.arccos( np.dot(v1,v2) / (np.linalg.norm(v1)*np.linalg.norm(v2))))
ang
#Second Method
v1 = np.array([4,3])
v2 = np.array([-3,4])
lengthV1 = np.sqrt(np.dot(v1,v1))
lengthV2 = np.sqrt(np.dot(v2,v2))
ang = np.rad2deg(np.arccos( np.dot(v1,v2) / (lengthV1 * lengthV2)))
print('Angle between Vectors - %s' %ang)
# # Inner & outer products
# +
v1 = np.array([1,2,3])
v2 = np.array([4,5,6])
np.inner(v1,v2)
print("\n Inner Product ==> \n", np.inner(v1,v2))
print("\n Outer Product ==> \n", np.outer(v1,v2))
# -
# # Vector Cross Product
v1 = np.array([1,2,3])
v2 = np.array([4,5,6])
print("\nVector Cross Product ==> \n", np.cross(v1,v2))
# # Matrix Creation
# Create a 4x4 matrix
A = np.array([[1,2,3,4] , [5,6,7,8] , [10 , 11 , 12 ,13] , [14,15,16,17]])
A
# Datatype of Matrix
A.dtype
B = np.array([[1.5,2.07,3,4] , [5,6,7,8] , [10 , 11 , 12 ,13] , [14,15,16,17]])
B
# Datatype of Matrix
B.dtype
# Shape of Matrix
A.shape
# Generate a 4x4 zero matrix
np.zeros((4,4))
#Shape of Matrix
z1 = np.zeros((4,4))
z1.shape
# Generate a 5x5 matrix filled with ones
np.ones((5,5))
# Return 10x10 matrix of random integer numbers between 0 to 500
np.random.randint(0,500, (10,10))
arr2
arr2.reshape(5,4)
mat1 = np.random.randint(0,1000,100).reshape(10,10)
mat1
mat1[0,0]
mat1[mat1 > 500]
# +
# Identity Matrix : https://en.wikipedia.org/wiki/Identity_matrix
I = np.eye(9)
I
# +
# Diagonal Matrix : https://en.wikipedia.org/wiki/Diagonal_matrix
D = np.diag([1,2,3,4,5,6,7,8])
D
# +
# Traingular Matrices (lower & Upper triangular matrix) : https://en.wikipedia.org/wiki/Triangular_matrix
M = np.random.randn(5,5)
U = np.triu(M)
L = np.tril(M)
print("lower triangular matrix - \n" , M)
print("\n")
print("lower triangular matrix - \n" , L)
print("\n")
print("Upper triangular matrix - \n" , U)
# -
# Generate a 5X5 matrix with a given fill value of 8
np.full((5,5) , 8)
# Generate 5X5 matrix of Random float numbers between 10 to 20
np.random.uniform(10,20, size=(5,5))
A
# Collapse Matrix into one dimension array
A.flatten()
# Collapse Matrix into one dimension array
A.ravel()
# # Reading elements of a Matrix
A
# Fetch first row of matrix
A[0,]
# Fetch first column of matrix
A[:,0]
# Fetch first element of the matrix
A[0,0]
A[1:3 , 1:3]
# # Reverse Rows / Columns of a Matrix
arr = np.arange(16).reshape(4,4)
arr
# Reverse rows
arr[::-1]
#Reverse Columns
arr[:, ::-1]
# # SWAP Rows & Columns
m1 = np.arange(0,16).reshape(4,4)
m1
# SWAP rows 0 & 1
m1[[0,1]] = m1[[1,0]]
m1
# SWAP rows 2 & 3
m1[[3,2]] = m1[[2,3]]
m1
m2 = np.arange(0,36).reshape(6,6)
m2
# Swap columns 0 & 1
m2[:,[0, 1]] = m2[:,[1, 0]]
m2
# Swap columns 2 & 3
m2[:,[2, 3]] = m2[:,[3, 2]]
m2
# # Concatenate Matrices
# Matrix Concatenation : https://docs.scipy.org/doc/numpy/reference/generated/numpy.concatenate.html
A = np.array([[1,2] , [3,4] ,[5,6]])
B = np.array([[1,1] , [1,1]])
C = np.concatenate((A,B))
C
# # Matrix Addition
# +
#********************************************************#
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
N = np.array([[1,1,1],[2,2,2],[3,3,3]])
print("\n First Matrix (M) ==> \n", M)
print("\n Second Matrix (N) ==> \n", N)
C = M+N
print("\n Matrix Addition (M+N) ==> \n", C)
# OR
C = np.add(M,N,dtype = np.float64)
print("\n Matrix Addition using np.add ==> \n", C)
#********************************************************#
# -
# # Matrix subtraction
# +
#********************************************************#
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
N = np.array([[1,1,1],[2,2,2],[3,3,3]])
print("\n First Matrix (M) ==> \n", M)
print("\n Second Matrix (N) ==> \n", N)
C = M-N
print("\n Matrix Subtraction (M-N) ==> \n", C)
# OR
C = np.subtract(M,N,dtype = np.float64)
print("\n Matrix Subtraction using np.subtract ==> \n", C)
#********************************************************#
# -
# # Matrices Scalar Multiplication
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
C = 10
print("\n Matrix (M) ==> \n", M)
print("\nMatrices Scalar Multiplication ==> \n", C*M)
# OR
print("\nMatrices Scalar Multiplication ==> \n", np.multiply(C,M))
# -
# # Transpose of a matrix
#
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
print("\n Matrix (M) ==> \n", M)
print("\nTranspose of M ==> \n", np.transpose(M))
# OR
print("\nTranspose of M ==> \n", M.T)
# -
# # Determinant of a matrix
#
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
print("\n Matrix (M) ==> \n", M)
print("\nDeterminant of M ==> ", np.linalg.det(M))
# -
# # Rank of a matrix
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
print("\n Matrix (M) ==> \n", M)
print("\nRank of M ==> ", np.linalg.matrix_rank(M))
# -
# # Trace of matrix
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
print("\n Matrix (M) ==> \n", M)
print("\nTrace of M ==> ", np.trace(M))
# -
# # Inverse of matrix A
#
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
print("\n Matrix (M) ==> \n", M)
print("\nInverse of M ==> \n", np.linalg.inv(M))
# -
# # Matrix Multiplication (pointwise multiplication)
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
N = np.array([[1,1,1],[2,2,2],[3,3,3]])
print("\n First Matrix (M) ==> \n", M)
print("\n Second Matrix (N) ==> \n", N)
print("\n Point-Wise Multiplication of M & N ==> \n", M*N)
# OR
print("\n Point-Wise Multiplication of M & N ==> \n", np.multiply(M,N))
# -
# # Matrix dot product
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
N = np.array([[1,1,1],[2,2,2],[3,3,3]])
print("\n First Matrix (M) ==> \n", M)
print("\n Second Matrix (N) ==> \n", N)
print("\n Matrix Dot Product ==> \n", M@N)
# OR
print("\n Matrix Dot Product using np.matmul ==> \n", np.matmul(M,N))
# OR
print("\n Matrix Dot Product using np.dot ==> \n", np.dot(M,N))
# -
# # Matrix Division
# +
M = np.array([[1,2,3],[4,-3,6],[7,8,0]])
N = np.array([[1,1,1],[2,2,2],[3,3,3]])
print("\n First Matrix (M) ==> \n", M)
print("\n Second Matrix (N) ==> \n", N)
print("\n Matrix Division (M/N) ==> \n", M/N)
# OR
print("\n Matrix Division (M/N) ==> \n", np.divide(M,N))
# -
# # Sum of all elements in a matrix
# +
N = np.array([[1,1,1],[2,2,2],[3,3,3]])
print("\n Matrix (N) ==> \n", N)
print ("Sum of all elements in a Matrix ==>")
print (np.sum(N))
# -
# # Column-Wise Addition
# +
N = np.array([[1,1,1],[2,2,2],[3,3,3]])
print("\n Matrix (N) ==> \n", N)
print ("Column-Wise summation ==> ")
print (np.sum(N,axis=0))
# -
# # Row-Wise Addition
# +
N = np.array([[1,1,1],[2,2,2],[3,3,3]])
print("\n Matrix (N) ==> \n", N)
print ("Row-Wise summation ==>")
print (np.sum(N,axis=1))
# -
# # Kronecker Product of matrices
#
M1 = np.array([[1,2,3] , [4,5,6]])
M1
M2 = np.array([[10,10,10],[10,10,10]])
M2
np.kron(M1,M2)
# # Matrix Powers
M1 = np.array([[1,2],[4,5]])
M1
# +
#Matrix to the power 3
M1@M1@M1
# +
#Matrix to the power 3
np.linalg.matrix_power(M1,3)
# -
# # Tensor
# +
# Create Tensor
T1 = np.array([
[[1,2,3], [4,5,6], [7,8,9]],
[[10,20,30], [40,50,60], [70,80,90]],
[[100,200,300], [400,500,600], [700,800,900]],
])
T1
# +
T2 = np.array([
[[0,0,0] , [0,0,0] , [0,0,0]],
[[1,1,1] , [1,1,1] , [1,1,1]],
[[2,2,2] , [2,2,2] , [2,2,2]]
])
T2
# -
# # Tensor Addition
A = T1+T2
A
np.add(T1,T2)
# # Tensor Subtraction
S = T1-T2
S
np.subtract(T1,T2)
# # Tensor Element-Wise Product
P = T1*T2
P
np.multiply(T1,T2)
# # Tensor Element-Wise Division
D = T1/T2
D
np.divide(T1,T2)
# # Tensor Dot Product
T1
T2
np.tensordot(T1,T2)
# # Solving Equations $$AX = B$$
A = np.array([[1,2,3] , [4,5,6] , [7,8,9]])
A
B = np.random.random((3,1))
B
# Ist Method
X = np.dot(np.linalg.inv(A) , B)
X
# 2nd Method
X = np.matmul(np.linalg.inv(A) , B)
X
# 3rd Method
X = np.linalg.inv(A)@B
X
# 4th Method
X = np.linalg.solve(A,B)
X
# # THANK YOU
| Machine Learning/Libraries/Introduction To Numpy/Introduction_to_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/zerotodeeplearning/ztdl-masterclasses/blob/master/solutions_do_not_open/Data_Manipulation_with_Pandas_solution.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="2bwH96hViwS7"
# ## Learn with us: www.zerotodeeplearning.com
#
# Copyright © 2021: Zero to Deep Learning ® Catalit LLC.
# + colab={} colab_type="code" id="bFidPKNdkVPg"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="DvoukA2tkGV4"
# # Data Manipulation with Pandas
# + [markdown] colab_type="text" id="a2c3dtIpqJQO"
# Pandas is an open source library providing high-performance, easy-to-use data structures and data analysis tools for the Python programming language.
#
# You can find it here: http://pandas.pydata.org/
#
# And the documentation can be found here: https://pandas.pydata.org/docs/getting_started/index.html
#
# In this notebook we review some of its functionality.
# + colab={} colab_type="code" id="oc7GxG8yKVJL"
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import seaborn as sns
# + colab={} colab_type="code" id="oEhGeaf9qJQR"
pd.__version__
# + colab={} colab_type="code" id="s0dZzGI9qJQT"
sns.__version__
# + [markdown] colab_type="text" id="BTun_fKVKQWD"
# ### Reading data from multiple sources
# + colab={} colab_type="code" id="8qGLbftIqJQW"
url = "https://raw.githubusercontent.com/zerotodeeplearning/ztdl-masterclasses/master/data/"
# + colab={} colab_type="code" id="PgND6MsjshTC"
#csv
df = pd.read_csv(url + "time_series_covid19_confirmed_global.csv")
# + colab={} colab_type="code" id="pM4zCZGDsr1E"
df.head()
# + colab={} colab_type="code" id="r7hot1pxsslm"
# json
df = pd.read_json(url + "gdp.json")
# + colab={} colab_type="code" id="QUx1qylXLAIa"
df.head()
# + colab={} colab_type="code" id="5bZPqdA-ME8f"
#html
dfl = pd.read_html(url + "FDIC_%20Failed%20Bank%20List.htm")
print(type(dfl))
print(len(dfl))
# + colab={} colab_type="code" id="KF3cFGEnMTH8"
dfl[0].head()
# + [markdown] colab_type="text" id="kEGpa0b0qJQp"
# ### Quick exploration
#
# Commands to quickly inspect the dataset
# + colab={} colab_type="code" id="wZF2KwA9qJQp"
df = pd.read_csv(url + "titanic-train.csv")
# + colab={} colab_type="code" id="mq1vBKwuqJQr"
df.head()
# + colab={} colab_type="code" id="zP6NgCkCqJQt"
df.info()
# + colab={} colab_type="code" id="Ido-qIQVqJQv"
df.describe()
# + [markdown] colab_type="text" id="0Yj83qd4qJQx"
# ### Plotting
# + colab={} colab_type="code" id="S9A6IPlyqJQy"
survived_counts = df['Survived'].value_counts()
survived_counts
# + colab={} colab_type="code" id="CMnPoowaqJQ0"
survived_counts.plot.bar(title='Dead / Survived');
# + colab={} colab_type="code" id="lOZXvtPiqJQ2"
survived_counts.plot.pie(
figsize=(5, 5),
explode=[0, 0.15],
labels=['Dead', 'Survived'],
autopct='%1.1f%%',
shadow=True,
startangle=90,
fontsize=16);
# + colab={} colab_type="code" id="LtFuP3icqJQ4"
df['Age']
# + colab={} colab_type="code" id="gRp96bgiqJQ6"
df['Age'].plot.hist(
bins=16,
range=(0, 80),
title='Passenger age distribution')
plt.xlabel("Age");
# + colab={} colab_type="code" id="VVzqVeOoqJQ7"
df['Age'].plot.hist(
bins=80,
range=(0, 80),
title='Passenger cumulative age distibution',
cumulative=True,
density=True)
plt.xlabel("Age")
plt.axhline(0.25, color='red')
plt.axhline(0.5, color='red')
plt.axhline(0.75, color='red');
# + [markdown] colab_type="text" id="6MqP-UvpqJQ9"
# ### Fancy plotting with seaborn
# + colab={} colab_type="code" id="ZdopJhU-qJQ9"
sns.pairplot(df[['Age', 'Pclass', 'Fare', 'SibSp', 'Survived']],
hue='Survived');
# + colab={} colab_type="code" id="yTLPL5EtqJQ_"
sns.jointplot(x='Age', y='Fare', data=df)
# + colab={} colab_type="code" id="zt6VlWTMqJRB"
corr = df.corr()
sns.heatmap(corr);
# + [markdown] colab_type="text" id="P-p-rTdXqJRD"
# ### Indexing
#
# Retrieving elements by row, by column or both. Try to understand each of the following statements
# + colab={} colab_type="code" id="SZA0jWpAqJRD"
df['Ticket']
# + colab={} colab_type="code" id="Z8Ioc4s6qJRF"
df[['Fare', 'Ticket']]
# + colab={} colab_type="code" id="mf6rCgfGqJRH"
df.iloc[3]
# + colab={} colab_type="code" id="a0rMAbcNqJRI"
df.iloc[3, 4]
# + colab={} colab_type="code" id="pbdwVHvYqJRK"
df.loc[0:4,'Ticket']
# + colab={} colab_type="code" id="EuGoeOJpqJRN"
df.loc[0:4, ['Fare', 'Ticket']]
# + [markdown] colab_type="text" id="ii8W8cl2qJRQ"
# ### Selections
#
# Retrieving part of the dataframe based on a condition. Try to understand each of the following statements.
# + colab={} colab_type="code" id="RLQ3S5xZqJRQ"
df[df.Age > 70]
# + colab={} colab_type="code" id="tgjoz4usqJRS"
df[(df['Age'] == 11) & (df['SibSp'] == 5)]
# + colab={} colab_type="code" id="1DfLF6-YqJRT"
df[(df.Age == 11) | (df.SibSp == 5)]
# + colab={} colab_type="code" id="8mRz-YggqJRV"
df.query('(Age == 11) & (SibSp == 5)')
# + colab={} colab_type="code" id="Lol7UCoBqJRX"
df.sort_values('Age', ascending = False).head()
# + [markdown] colab_type="text" id="S3wdVTv2qJRY"
# ### Distinct elements
# + colab={} colab_type="code" id="wnA1zSgiqJRZ"
df['Embarked'].unique()
# + [markdown] colab_type="text" id="9oT4fBe0qJRb"
# ### Group by
#
# Pandas supports many SQL-like operations like group by, order by and join. In pandas they are called:
# - groupby
# - sort_values
# - merge
#
# Try to make sense of each of the following expressions:
# + colab={} colab_type="code" id="sRXQst5kqJRb"
# Find average age of passengers that survived vs. died
df.groupby('Survived')['Age'].mean()
# + colab={} colab_type="code" id="ks6GFg-hqJRd"
df.sort_values('Age', ascending = False).head()
# + [markdown] colab_type="text" id="gaU7w-9ZqJRg"
# ### Join (merge)
# + colab={} colab_type="code" id="hoZG7hniqJRg"
df1 = df[['PassengerId', 'Survived']]
df2 = df[['PassengerId', 'Age']]
pd.merge(df1, df2, on='PassengerId').head()
# + [markdown] colab_type="text" id="bNC_ZcG8qJRh"
# ### Pivot Tables
#
# Pandas also supports Excel-like functionality like pivot tables.
# + colab={} colab_type="code" id="MRb0CyoqqJRi"
df.pivot_table(index='Pclass', columns='Survived', values='PassengerId', aggfunc='count')
# + colab={} colab_type="code" id="8edst5deqJRj"
df['Pclass'].value_counts()
# + [markdown] colab_type="text" id="RqyflXWPqJRl"
# ### Exercise 1:
#
# Try doing a few of these:
#
# - select passengers that survived
# - select passengers that embarked in port S
# - select male passengers
# - select passengers who paid less than 40.000 and were in third class
# - locate the name of passegner Id 674
# - calculate the average age of passengers using the function mean()
# - count the number of survived and the number of dead passengers
# - count the number of males and females
# - count the number of survived and dead per each gender
# - calculate average price paid by survived and dead people in each class
# + colab={} colab_type="code" id="9mZBuwBWqJRl" tags=["solution", "empty"]
df.query('Survived == 1').head()
# + colab={} colab_type="code" id="mhSysGxUqJRn" tags=["solution"]
df.query('Embarked == "S"').head()
# + colab={} colab_type="code" id="0dhNlVJWqJRo" tags=["solution"]
df[df['Sex'] == 'male'].head()
# + colab={} colab_type="code" id="6yJJNTwDqJRq" tags=["solution"]
df[(df.Fare < 40000) & (df.Pclass == 3)].head()
# + colab={} colab_type="code" id="4RXQyDqjqJRs" tags=["solution"]
df.query('PassengerId == 674')
# + colab={} colab_type="code" id="nXi6FxC0qJRt" tags=["solution"]
df['Age'].mean()
# + colab={} colab_type="code" id="-P0p7w50qJRw" tags=["solution"]
df['Survived'].value_counts()
# + colab={} colab_type="code" id="W77KgZ8jqJRx" tags=["solution"]
df['Sex'].value_counts()
# + colab={} colab_type="code" id="ITms2aB-qJR0" tags=["solution"]
df.pivot_table(index='Survived', columns='Sex', values='PassengerId', aggfunc='count')
# + colab={} colab_type="code" id="wHTRiTwnqJR1" tags=["solution"]
df.pivot_table(index='Survived', columns='Pclass', values='Fare', aggfunc='mean')
# + [markdown] colab_type="text" id="t11aXKg9qJR5"
# ### Time series data
# + colab={} colab_type="code" id="3uyILVmnqJR5"
df = pd.read_csv(url + "time_series_covid19_confirmed_global.csv")
# + colab={} colab_type="code" id="WEdrGMQVqJR7"
df.head()
# + [markdown] colab_type="text" id="D4jIe7TtqJR9"
# ### Dropping columns
# + colab={} colab_type="code" id="3yRzKqDcqJR9"
df1 = df.drop(['Lat', 'Long'], axis=1)
df1
# + [markdown] colab_type="text" id="WHTRRCE8qJSA"
# ### Groupby + Aggregation
# + colab={} colab_type="code" id="6h2IwmFTqJSA"
dfcountries = df1.groupby('Country/Region').sum().transpose()
dfcountries
# + [markdown] colab_type="text" id="23shWiZyqJSB"
# ### Data Time index
# + colab={} colab_type="code" id="yJouYnndqJSC"
dfcountries.index
# + colab={} colab_type="code" id="OXqX_7U-qJSE"
dfcountries.index = pd.to_datetime(dfcountries.index)
# + colab={} colab_type="code" id="WG6yJPu0qJSF"
dfcountries.index
# + colab={} colab_type="code" id="un5n4AX_qJSH"
dfcountries.head()
# + colab={} colab_type="code" id="1VyxSJzkqJSI"
series = dfcountries['Italy']
series[series > 100].plot(style='o-', logy=True, title='Italy')
plt.ylabel('Confirmed Cases');
# + colab={} colab_type="code" id="AEmAoHR4qJSJ"
country = 'Italy'
ts = dfcountries[country].diff()
ts = ts[ts>0]
fig, ax = plt.subplots(figsize=(10,6))
ax.bar(ts.index, ts.values)
ax.xaxis.set_major_locator(mdates.WeekdayLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%b %d'))
ax.set_title(f'{country}: New cases')
ax.set_ylabel('Count')
ax.set_xlabel('Date');
# + [markdown] colab_type="text" id="U67wuZ__qJSK"
# ### Exercise 2:
#
# - find the top 20 countries by number of cases. You will need to use selections and these methods:
# - `.tail()`
# - `.transpose`
# - `.sort_values`
#
# Your goal is to obtain a Pandas Series that looks like this:
# ```
# Country/Region
# US 188172
# Italy 105792
# Spain 95923
# China 82279
# Germany 71808
# France 52827
# ...
# ```
# - use a horizontal bar plot to plot them
# - bonus points if you add title and labels
# + colab={} colab_type="code" id="CbOIQblFqJSK" tags=["solution", "empty"]
top_countries = dfcountries.tail(1).transpose().iloc[:,0].sort_values(ascending=False).head(20)
top_countries
# + colab={} colab_type="code" id="Gbq5aQPlqJSL" tags=["solution"]
top_countries.sort_values().plot.barh(
title='Total cases',
figsize=(15, 10));
# + colab={} colab_type="code" id="E8oVKk_mqJSN"
| solutions_do_not_open/Data_Manipulation_with_Pandas_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Do a little TorchScript
# > lets see what's shakin'
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [jupyter]
# - image: images/chart-preview.png
#
import torch
torch.__version__
# +
# create a class that subclass torch.nn.Module
class MyCell(torch.nn.Module):
def __init__(self):
super(MyCell, self).__init__()
def forward(self, x, h):
new_h = torch.tanh(x + h)
return new_h, new_h
my_cell = MyCell()
x = torch.rand(3, 4)
h = torch.rand(3, 4)
print(my_cell(x, h))
# +
class MyCell(torch.nn.Module):
def __init__(self):
super(MyCell, self).__init__()
self.linear = torch.nn.Linear(4, 4)
def forward(self, x, h):
new_h = torch.tanh(self.linear(x) + h)
return new_h, new_h
my_cell = MyCell()
print(my_cell)
print(my_cell(x, h))
| _notebooks/2020-07-22-TorchScript-intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# VERYYYY VERYYYYY IMPORTANTTTTTTTTTTTTTTTTTTT
# %%writefile pythonfile.txt
HEY THIS IS A TRIAL TEXT FILE
myfile=open('pythonfile.txt')
pwd
myfile=open('pythonfile.txt')
myfile.read()
myfile
myfile.read()
myfile.seek(0)
myfile.read()
myfile.seek(0)
myfile.read()
myfile.seek(0)
contents=myfile.read()
contents
myfile.seek(0)
myfile.readlines()
myfile.readline()
myfile.seek(0)
myfile.readline()
myfile.seek(0)
myfile=open("pythonfile.txt")
myfile=open('pythonfile.txt')
with open('pythonfile.txt') as my_new_file:
contents = my_new_file.read()
contents
contents
contents
with open('pythonfile.txt',mode='r') as myfile:
contents=myfile.read()
$
# %%writefile myfile.txt
one on first
two on second
three on third
with open('myfile.txt',mode='r') as f:
print(f.read())
with open('myfile.txt',mode='a') as f:
f.write('\nFour On Fourth')
with open('myfile.txt',mode='r') as f:
print(f.read())
with open('asfuck.txt',mode='w') as f:
f.write('I MADE THIS FILE \n Dont TOUCH IT')
with open('asfuck.txt') as f:
print(f.read())
| trial scripts/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing data stored in Cloudant DB from Watson IoT Platform and Anomaly Detection by using IBM Watson Studio
# Install the spark-sql-cloudant package (library for reading data from Cloudant database using Spark SQL) in your IBM Watson Studio account by executing the following cell, and then restart the kernel. You will use **pixiedust** to import the required packages.
# !pip install --upgrade pixiedust
import pixiedust
# Use play-json version 2.5.9. Latest version is not supported at this time.
pixiedust.installPackage("com.typesafe.play:play-json_2.11:2.5.9")
# Get the matching version sql-cloudant library
pixiedust.installPackage("org.apache.bahir:spark-sql-cloudant_2.11:0")
# To fix PyJavaError
pixiedust.packageManager.uninstallPackage("org.apache.bahir:spark-sql-cloudant_2.11:2.2.1")
# **User input required** : Cloudant credentials.
#
# If you have a connection with Cloudant set up for this project, complete the following steps:
# 1. Import your Cloudant credentials by clicking on the following cell below to select it
# 2. Click **Find and Add Data**
# 3. Select the Connections tab and click on **Insert to code**. A dictionary called *credentials_1* is added to the cell that contains the Cloudant credentials. If the dictionary has another name, change it to *credentials_1*.
# 4. Run the cell.
#
# If you don’t have a connection with Cloudant set up, the credentials can be found on IBM Cloud dashboard by completing the following steps:
#
# 1. Go to your Cloudant service on IBM Cloud,
# 2. Select its Service Credentials section on the left
# 3. Click **View Credentials** to view the username and password.
# 4. Update the *username* and *password* variables with Cloudant’s username and password.
# +
# This empty cell will be populated with your Cloudant credentials if you follow the steps explained above.
# -
username = credentials_1["username"]
password = <PASSWORD>["password"]
host = username + '.cloudant.com'
# **User input required**: Cloudant database name.
#
# If you are not sure which database contains the data that you want to import, go to your Cloudant service on IBM Cloud and click **Launch** to display the database name. Update the *dbName* variable with the database name, for example
# *iotp_abcdef_default_2018-01-10*.
dbName = "DBName"
# Connect to the Cloudant database that is generated by WIoTP connector for historical data.
#
# The following code connects to Cloudant NoSQL DB and returns an RDD data frame for the stored IoT data. The line `option("jsonstore.rdd.partitions", 4)` is needed only if your Cloudant service plan is *lite* because this plan has an access quota of 5 requests per second.
# Spark might run parallel jobs that might lead to more than 5 requests being made in one second. If this happens, a "too many requests" error is raised. To resolve this error, decrease the value for the *jsonstore.rdd.partitions* option to 2. For paid service plans this line can be commented out.
#
cloudantdata=sqlContext.read.format("org.apache.bahir.cloudant").\
option("cloudant.host", host).\
option("cloudant.username", username).\
option("cloudant.password", password).\
option("view","_design/iotp/_view/by-date").\
option("jsonstore.rdd.partitions", 4).\
load(dbName)
# Observe the loaded data:
cloudantdata.show()
# All IoT data is located under the value column.
#
# Next, transform this hierarchical data frame into a flat one, and convert the timestamp from string type into a timestamp type.
# The function withColumn adds a column named 'ts' to the data frame, and calculates it's content based on timestamp column (string), by using the to_ts function that was defined.
# The cache() function of a data frame caches the data frame in memory, this is very useful when data is accessed repeatedly.
#
# The *deviceData* is a temporary view in the Spark Session and can be used for select statements if you want to write raw SQL.
#
import pandas as pd
from pyspark.sql import *
from pyspark.sql.functions import udf, col, asc, desc,to_date, unix_timestamp, weekofyear, countDistinct
from datetime import datetime
from pyspark.sql.types import DateType, TimestampType, IntegerType
# +
# This function converts the string cell into a timestamp type:
str_to_ts = udf (lambda d: datetime.strptime(d, "%Y-%m-%dT%H:%M:%S.%fZ"), TimestampType())
sparkDf = cloudantdata.selectExpr("value.deviceId as deviceId", "value.deviceType as deviceType", "value.eventType as eventType" , "value.timestamp as timestamp", "value.data.*","value.data.d.oa as oa","value.data.d.ob as ob","value.data.d.og as og")
sparkDf = sparkDf.withColumn('ts', str_to_ts(col('timestamp')))
sparkDf.cache()
sparkDf.createOrReplaceTempView("deviceData")
# show the resulting schema and data
sparkDf.printSchema()
spark.sql("SELECT * from deviceData").show(10)
# -
# ### Change the values of deviceId and deviceType for which you want to see visualizations.
deviceId = 'phone'
deviceType = 'simulator'
# Along with device movement data, you can also add acceleration data(ax,ay,az).
#
# ## Data visualization and comprehension
#
# ### Device Health
#
# In this section we will see how to learn about the population of IoT devices and answer questions such as:
# 1. How many reports each device type had?
# 2. What is the breakdown of the devices per device type?
# 3. How many reports have been sent by each device?
# 4. How many reports each event type had?
# 5. How many devices reported in a given time interval?
# %matplotlib inline
import matplotlib.pyplot as plt
import pprint
from collections import Counter
import numpy as np
from matplotlib import dates
# Use Spark to prepare the data for visualization, because Spark can support big data processing. When the data is ready for visualization, convert Spark data Frame into Pandas data Frame, because Pandas has good visualization support.
#
# #### How many reports each device type had?
#
# Setting the *deviceType* as index of the created Pandas data frame causes the bar plot to be aggregated by the deviceType. Call the plot function of the Pandas data frame.
# +
EperDtDF = spark.sql("SELECT ts,deviceType from deviceData").groupBy("deviceType").count()
EperDtDF.cache()
EperDtDF.show()
EperDtPanda = EperDtDF.toPandas().set_index('deviceType')
ax = EperDtPanda.plot(kind='bar',legend=False)
ax.set_xlabel("deviceType")
ax.set_ylabel("events count")
ax.set_title('count of events by deviceType')
# -
# #### What is the breakdown of the devices per device type?
# The bar chart is plotted in the same way as before, but now we will also show the pie chart of the data. Pandas data frame supports different plot types. Using pie generates a pie chart with percentage sizes of the pieces.
# To write the actual count of the devices, instead of percentages, use the autopct argument - multiply by the total amount of devices and divide by 100 to get the actual numbers.
# The total is calculated by using the *sum()* function of Pandas data frame, which sums up the device count of all the deviceTypes. The sum function returns a DataFrame, so use the *[0]* index to get only the value into the total.
# +
DperDtDF = spark.sql("SELECT deviceId,deviceType from deviceData").groupBy("deviceType").agg(countDistinct('deviceId'))
EperDtDF.cache()
DperDtDF.show()
# bar chart of deviceId by deviceType
EperDtPanda = DperDtDF.toPandas().set_index('deviceType')
ax = EperDtPanda.plot(kind='bar',legend=False)
ax.set_xlabel("deviceType")
ax.set_ylabel("devices count")
ax.set_title('count of deviceIds by deviceType')
# Pie chart of deviceId by deviceType
fig = plt.figure(figsize=(5,5))
ax = plt.subplot(111)
total = EperDtPanda.sum()[0]
ax = EperDtPanda.plot(kind='pie', ax=ax, figsize=(5,5), legend=False, shadow=True, subplots=True, autopct=lambda p: '{:.0f}'.format(p * total / 100))
plt.title("count of deviceIds by deviceType")
# -
# #### How many reports have been sent by each device?
# +
EperDdf = spark.sql("SELECT deviceId,ts from deviceData").groupBy("deviceId").count()####.sort()########
EperDtDF.cache()
EperDdf.show()
EperDPanda = EperDdf.toPandas().set_index('deviceId')
ax = EperDPanda.plot(kind='bar',legend=False)
ax.set_xlabel("deviceId")
ax.set_ylabel("events count")
ax.set_title('count of events by deviceId')
# -
# #### How many reports each event type had?
# +
EperEtDF = spark.sql("SELECT eventType,ts from deviceData").groupBy("eventType").count()
EperDtDF.cache()
EperEtDF.show()
EperEtPanda = EperEtDF.toPandas().set_index('eventType')
ax = EperEtPanda.plot(kind='bar',legend=False)
ax.set_xlabel("eventType")
ax.set_ylabel("events count")
ax.set_title('count of events by eventType')
# -
# #### How many devices reported in a given time interval?
#
# **User input required**: Replace the year, month and day in the following cell to specify `start` and `end` interval.
#
# For example:
#
# `start = datetime(2017, 7, 28, 0, 0, 0)
# end = datetime(2017, 7, 28, 23, 59, 59)`
#
# Make sure that the interval contains device events. You can narrow down the time interval as well. Then run the cell.
# +
# set the time interval of interest
dbDate = dbName.split("_")[3]
dbDate = dbDate.split("-")
# datetime(year, month, day, hours, minutes, seconds)
start = datetime(int(dbDate[0]), int(dbDate[1]), int(dbDate[2]), 0, 0, 0)
end = datetime(int(dbDate[0]), int(dbDate[1]), int(dbDate[2]), 23, 59, 59)
print(start)
print(end)
# -
# First we filter the data by a time interval, then group the resulting dataFrame by *deviceId*, and count the records for each *deviceId*.
# +
#filter by time interval
deviceMetaData = sparkDf.select('deviceId','deviceType','ts','timestamp','eventType').filter((col('ts')>=start) & (col('ts')<=end))
deviceMetaData.cache()
#deviceMetaData.show()
#how many devices reported in interval
byDevice = deviceMetaData.groupby(['deviceId']).count()
byDevice.cache()
print("Number of events by deviceId in the interval: ")
byDevice.show()
print("total number of devices reported in the interval:", byDevice.count())
# -
# Count of rows by time span for a specific device, using the filter function of Spark DataFrame:
byDevice.filter(byDevice["deviceId"]== deviceId).show() ##also show 5 with lowest counts
# Extract all the numeric columns for further analytics; only a subset of the numeric columns are selected for this demonstration:
# +
#find all numeric columns of the DataFrame
numericCols = [name_dt for name_dt in sparkDf.dtypes if (('double' in name_dt[1]) or ('int' in name_dt[1]) or ('long' in name_dt[1]))]
#numericCols is a list of pairs (columnName, dataType), here we select only the column name into the allkeys list
allkeys = [x[0] for x in numericCols]
print("all numeric columns", allkeys)
#select only 5 numeric columns for further detailed visualizations
keys = ['<KEY>']
print("selected 3 numeric columns", keys)
# -
# #### Device type sensor visualization
#
# Summary of sensor data that is reported by all devices of a device type, for example:
#
# 1. What is the Average/Min/Max of all reported sensor values?
# 2. Can I see a histogram of a sensor's output?
# 3. What is the correlation between two sensors?
#
# #### Average/Min/Max of all reported sensor values
# +
from pyspark.mllib.stat import Statistics
#showing visualization for device type
#show summary only for the selected 5 columns, for easier view, since we have too many columns to fit in a row
dfKeysType = sparkDf.select(*keys).filter(sparkDf["deviceType"]==deviceType)
dfKeysType.cache()
dfKeysType.describe().show()
# -
# #### Histogram of a device type sensor's output
#
# 1. Use Spark DataFrame to prepare the histogram for each specific sensor (key) (using rdd.flatMap)
# 2. Create Pandas DataFrame from the calculated histogram with 2 columns: "bin" and "frequency".
# 3. Plot the histogram using Pandas plot function.
for key in keys:
histogram = dfKeysType.select(key).rdd.flatMap(lambda x: x).histogram(11)
#print histogram
pandaDf = pd.DataFrame(list(zip(list(histogram)[0],list(histogram)[1])),columns=['bin','frequency']).set_index('bin')
ax =pandaDf.plot(kind='bar')
ax.set_ylabel("frequency")
ax.set_title('Histogram of ' + key + ' sensor output')
# #### Correlation between two sensors
#
# Correlation between two sensors can be plotted using Pandas plot with kind='scatter'. Remember that *dfKeysType* is a Spark DataFrame that includes only our selected 5 columns and is filtered by *deviceType*. You can also filter by *timestamp* to decrease the amount of data that is output.
# +
key1="oa"
key2="ob"
pandaDF = dfKeysType.toPandas()
ax = pandaDF.plot(kind='scatter', x=key1, y=key2, s=5, figsize=(7,7))
ax.set_title('Relationship between ' + key1 + ' and ' + key2 )
# -
# To view all the correlations of the selected 5 columns, together with a histogram on a diagonal, use the Pandas scatter_matrix function:
pd.plotting.scatter_matrix(pandaDF, figsize=(18,12))
plt.show()
# A correlation matrix can be plotted, using Pandas corr() function on the DataFrame:
# +
correlations = pandaDF.corr()
# plot correlation matrix
fig = plt.figure(figsize=(12,12))
ax = fig.add_subplot(111)
cax = ax.matshow(correlations, vmin=-1, vmax=1)
fig.colorbar(cax)
ticks = np.arange(0,5,1)
ax.set_xticks(ticks)
ax.set_yticks(ticks)
ax.set_xticklabels(keys)
ax.set_yticklabels(keys)
plt.show()
# -
# ## Sensor deep dive
#
# Sensor deep dive output is similar to the device type sensor visualization but the data is filtered by *deviceId*.
#
# #### Average/Min/Max of all reported sensor values by the device
# +
#showing visualization for specific deviceID set above
#show summary only for a selected group of columns, for easier view, since we have too many columns to fit in a row
dfKeysDev = sparkDf.select(*keys).filter(sparkDf["deviceId"]==deviceId)
dfKeysDev.cache()
dfKeysDev.describe().show()
# -
# A box plot is a method for graphically depicting groups of numerical data through their quartiles. The box extends from the lower to upper quartile values of the data, with a line at the median. The whiskers extend from the box to show the range of the data. Beyond the whiskers, data are considered outliers and are plotted as individual points.
#
# A box plot for each devices sensor, produced with the Pandas plot function with kind="box":
pandaDF = dfKeysDev.toPandas()
pandaDF.plot(kind='box', subplots=True, layout=(10,3), sharex=False, sharey=False, figsize=(25,60))
plt.show()
# #### Histogram of a device's sensor output
for key in keys:
try:
#The histogram is built with spark. Only the groupped by bins data will be converted to Pandas DataFrame
histogram = dfKeysDev.select(key).rdd.flatMap(lambda x: x).histogram(11)
#print histogram
pandaDf = pd.DataFrame(zip(list(histogram)[0],list(histogram)[1]),columns=['bin','frequency']).set_index('bin')
ax = pandaDf.plot(kind='bar')
ax.set_ylabel("frequency")
ax.set_title('Histogram of ' + key + ' sensor output')
except:
print("no values for sensor " + key + " for device " + deviceId + "\n")
# The histograms can also be built more easily with Pandas DataFrame, in case the dfKeysDev DataFrame is not too large. For the case of big data, spark is more scalable.
pandaDF = dfKeysDev.toPandas()
pandaDF.hist(layout=(3,3), sharex=False, figsize=(20,15))
# #### Density Plots
#
# Kernel density estimation (KDE) is a non-parametric way to estimate the probability density function of a random variable. Kernel density estimation is a fundamental data smoothing problem where inferences about the population are made, based on a finite data sample.
#
# **Note**: here we convert the data into Pandas DataFrame, after we filtered by deviceId and selected a subset of keys. In case this is still too much data for the Pandas DataFrame to handle, consider selecting fewer keys and filtering by time interval.
# +
pandaDF = dfKeysDev.toPandas()
ax = pandaDF.plot(kind='density', subplots=True, layout=(3,3), sharex=False, figsize=(20,15))
plt.show()
# -
# #### How a specific device sensor value changes over time
#
# Maximum, minimum, and average lines are shown on plots.
# +
from pyspark.sql.functions import mean, min, max
#showing visualization for specific deviceID set above
print(keys)
for key in keys:
df = spark.sql("SELECT deviceId, ts," + key +" from deviceData where deviceId='" + deviceId + "'").where(col(key).isNotNull())
df.cache()
if (df.count() > 0):
pandaDF = df.toPandas()
ax = pandaDF.plot(x='ts', y=key , legend=False, figsize=(15,9), ls='-', marker='o')
ax.xaxis.set_major_formatter(dates.DateFormatter('%d-%m-%Y %H:%M:%S'))
ax.set_title(key + ' over time')
ax.set_ylabel(key)
ax.grid(True)
# Draw lines to showcase the upper and lower threshold
ax.axhline(y=pandaDF[key].min(),c="red",linewidth=2,zorder=0)
ax.axhline(y=pandaDF[key].max(),c="red",linewidth=2,zorder=0)
ax.axhline(y=pandaDF[key].mean(),c="green",linewidth=2,zorder=0, ls='--')
ax.autoscale_view()
# -
# **Note**: Data can be aggregated by intervals, for example by seconds/minutes/hours in Spark and displayed as aggregated plots (average, minimum, maximum).
# +
from pyspark.sql.functions import window
from functools import reduce
#showing visualization for specific deviceID set above
for key in keys:
df = spark.sql("SELECT deviceId, ts," + key +" from deviceData where deviceId='" + deviceId + "'").where(col(key).isNotNull())
df = df.groupBy("deviceId", window("ts", "30 seconds")).agg(max(key), min(key), mean(key))
#df = df.groupBy("deviceId", window("ts", "1 minute")).agg(max(key), min(key), mean(key))
#df.groupBy("deviceId", window("ts", "5 minutes")).agg(max(key), min(key), mean(key))
#df.groupBy("deviceId", window("ts", "1 hour")).agg(max(key), min(key), mean(key))
#change automatic name of aggregated column
oldColumns = df.schema.names
newColumns = ["deviceId", "window", "max", "min", "avg"]
df = reduce(lambda df, idx: df.withColumnRenamed(oldColumns[idx], newColumns[idx]), range(len(oldColumns)), df)
win_to_ts = udf (lambda d: d.start, TimestampType())
df = df.withColumn('ts', win_to_ts(col('window')))
df = df.select('deviceId','ts',"max", "min", "avg")
df.cache()
if (df.count() > 0):
pandaDF = df.toPandas()
ax = pandaDF.plot(x='ts', y='min', legend=True, figsize=(15,9), ls='-', marker='o', c="red")
ax = pandaDF.plot(ax=ax, x='ts', y='max', legend=True, figsize=(15,9), ls='-', marker='o', c="red")
ax = pandaDF.plot(ax=ax, x='ts', y='avg', legend=True, figsize=(15,9), ls='-', marker='o', c="green")
#'S' secondly frequency, 'T' minutely frequency, 'H' hourly frequency
xtick = pd.date_range(start=pandaDF['ts'].min(), end=pandaDF['ts'].max(), freq='30S')
ax.set_xticks(xtick)
ax.xaxis_date()
ax.set_title(key + ' over time groupped by 30 sec')
ax.set_ylabel(key)
ax.autoscale_view()
# -
# For Big Data is also possible to show the plot around the extremum point. In the example below we show values spanned in 4 seconds around the maximum value. Note that the aggregation is done inside Spark and Pandas is used for visualization.
# +
from pyspark.sql.functions import mean, min, max
#showing visualization for specific deviceID
for key in keys:
df = spark.sql("SELECT deviceId, ts," + key +" from deviceData where deviceId='" + deviceId + "'").where(col(key).isNotNull())
df.cache()
if (df.count() > 0):
#find max and filter around them
max_value = df.select(max(key)).collect()[0][0]
maxts = df.filter(df[key] == max_value).rdd.map(lambda r: r['ts']).collect()[0]
startts = maxts - pd.DateOffset(seconds=4) #(minutes=2)#(days=15)
endts = maxts + pd.DateOffset(seconds=4)
df_max = df.filter((col('ts')>=startts) & (col('ts')<=endts))
pandaDF = df_max.toPandas()
ax = pandaDF.plot(x='ts', y=key , legend=False, figsize=(15,9), ls='-', marker='o')
ax.xaxis.set_major_formatter(dates.DateFormatter('%d-%m-%Y %H:%M:%S'))
ax.set_title(key + ' over time')
ax.set_ylabel(key)
ax.grid(True)
# Draw lines to showcase the upper and lower threshold
ax.axhline(y=pandaDF[key].min(),c="red",linewidth=2,zorder=0)
ax.axhline(y=pandaDF[key].max(),c="red",linewidth=2,zorder=0)
ax.axhline(y=pandaDF[key].mean(),c="green",linewidth=2,zorder=0, ls='--')
ax.autoscale_view()
# -
# #### Compare between the sensor values of devices over time
#
# The *dfKeysDev* DataFrame contains only keys columns, with no ts column, so we will create a new data frame that will also include the *ts*:
# +
#showing visualization for specific deviceID
columns = list(keys)
columns.append('ts')
df = sparkDf.select(*columns).filter(sparkDf["deviceId"]==deviceId)
pandaDF = df.toPandas().set_index('ts')
ax = pandaDF.plot(figsize=(15,9),ls='', marker='o')
ax.xaxis.set_major_formatter(dates.DateFormatter('%d-%m-%Y %H:%M:%S'))
ax.set_title(', '.join(keys) + ' over time')
ax.grid(True)
ax.autoscale_view()
# +
pandaDF = dfKeysDev.toPandas()
pd.plotting.scatter_matrix(pandaDF, figsize=(18,12))
plt.show()
# -
# ## Anomaly detection
#
#
# Anomaly detection will be performed using *z-score*. *z-score* is a standard score that indicates how many standard deviations an element is from the mean. A z-score can be calculated from the following formula:
# `z = (X - µ) / σ`
# where z is the *z-score*, *X* is the value of the element, *µ* is the population mean, and *σ* is the standard deviation.
#
# A higher *z-score* value represents a larger deviation from the mean value which can be interpreted as abnormal.
#
# We will calculate *z-score* for each selected column (sensor) of each device type, and plot only the sensors that have spikes. We define a spike in the following function spike(row), by reported value having z-score above 3 or below -3. Observe that the values for which the *z-score* is above 3 or below -3, are marked as abnormal events in the following graph.
#
# **Note**: The code triggers a number of Spark jobs and might take a few seconds to finish.
# +
# ignore warnings if any
import warnings
from pyspark.sql.functions import mean, min, max, mean, stddev
warnings.filterwarnings('ignore')
'''
This function detects the spike and dip by returning a non-zero value
when the z-score is above 3 (spike) and below -3(dip). Incase if you
want to capture the smaller spikes and dips, lower the zscore value from
3 to 2 in this function.
'''
upperThreshold = 3
lowerThreshold = -3
def spike(row):
if(row['zscore'] >=upperThreshold or row['zscore'] <=lowerThreshold):
return row[key]
else:
return 0
#get the list of available devices
deviceTypes = sparkDf.select("deviceType").groupBy("deviceType").count().rdd.map(lambda r: r[0]).collect()
#calculate for each device type and each key
for devt in deviceTypes:
for key in keys:
df = spark.sql("SELECT deviceType, ts," + key +" from deviceData where deviceType='" + devt + "'").where(col(key).isNotNull())
if (df.count() > 0):
pandaDF = df.toPandas().set_index("ts")
# calculate z-score and populate a new column
pandaDF['zscore'] = (pandaDF[key] - pandaDF[key].mean())/pandaDF[key].std(ddof=0)
#add new column - spike, and calculate its value based on the thresholds, usinf spike function, defined above
pandaDF['spike'] = pandaDF.apply(spike, axis=1)
#plot the chart, only if spikes were detected (not all values of "spike" are zero)
if (pandaDF['spike'].nunique() > 1):
# select rows that are required for plotting
plotDF = pandaDF[[key,'spike']]
#calculate the y minimum value
y_min = (pandaDF[key].max() - pandaDF[key].min()) / 10
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
ax.set_ylim(plotDF[key].min() - y_min, plotDF[key].max() + y_min)
x_filt = plotDF.index[plotDF.spike != 0]
plotDF['spikes'] = plotDF[key]
y_filt = plotDF.spikes[plotDF.spike != 0]
#Plot the raw data in blue colour
line1 = ax.plot(plotDF.index, plotDF[key], '-', color='blue', animated = True, linewidth=1, marker='o')
#plot the anomalies in red circle
line2 = ax.plot(x_filt, y_filt, 'ro', color='red', linewidth=2, animated = True)
#Fill the raw area
ax.fill_between(plotDF.index, (pandaDF[key].min() - y_min), plotDF[key], interpolate=True, color='blue',alpha=0.6)
# calculate the sensor value that is corresponding to z-score that defines a spike
valUpperThreshold = (pandaDF[key].std(ddof=0) * upperThreshold) + pandaDF[key].mean()
# calculate the sensor value that is corresponding to z-score that defines a dip
valLowerThreshold = (pandaDF[key].std(ddof=0) * lowerThreshold) + pandaDF[key].mean()
#plot the thresholds
ax.axhline(y=valUpperThreshold,c="red",linewidth=2,zorder=0,linestyle='dashed',label='Upper threshold')
ax.axhline(y=valLowerThreshold,c="red",linewidth=2,zorder=0,linestyle='dotted',label='Lower threshold')
# Label the axis
ax.set_xlabel("Sequence",fontsize=20)
ax.set_ylabel(key,fontsize=20)
ax.set_title("deviceType: " + devt + " sensor:" + key)
plt.tight_layout()
plt.legend()
print("Device Type: " + devt + ", sensor: " + key)
print("Upper treshould based on the z-score calculation: " , upperThreshold , ": " , valUpperThreshold)
print("Lower treshould based on the z-score calculation: ", lowerThreshold, ": " , valLowerThreshold)
plt.show()
# -
# The red marks indicate unexpected spikes where the z-score value is greater than 3 or less than -3. To detect lower spikes, modify the value to 2 or less. Similarly, if you want to detect only the higher spikes, increase the z-score value from 3 to 4 or more.
#
# For complete solution tutorial, refer [Gather, Visualize and Analyze IoT data](http://console.bluemix.net/docs/tutorials/gather-visualize-analyze-iot-data.html#gather-visualize-and-analyze-iot-data)
| anomaly-detection/Anomaly-detection-watson-studio-python3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import scipy
import scipy.misc
import scipy.ndimage
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import OneHotEncoder
from datetime import datetime
import resource
np.set_printoptions(suppress=True, precision=5)
# %matplotlib inline
# +
class Laptimer:
def __init__(self):
self.start = datetime.now()
self.lap = 0
def click(self, message):
td = datetime.now() - self.start
td = (td.days*86400000 + td.seconds*1000 + td.microseconds / 1000) / 1000
memory = resource.getrusage(resource.RUSAGE_SELF).ru_maxrss / (1024 ** 2)
print("[%d] %s, %.2fs, memory: %dmb" % (self.lap, message, td, memory))
self.start = datetime.now()
self.lap = self.lap + 1
return td
def reset(self):
self.__init__()
def __call__(self, message = None):
return self.click(message)
timer = Laptimer()
timer()
# +
def normalize_fetures(X):
return X * 0.98 / 255 + 0.01
def normalize_labels(y):
y = OneHotEncoder(sparse=False).fit_transform(y)
y[y == 0] = 0.01
y[y == 1] = 0.99
return y
# -
url = "https://raw.githubusercontent.com/makeyourownneuralnetwork/makeyourownneuralnetwork/master/mnist_dataset/mnist_train_100.csv"
train = pd.read_csv(url, header=None, dtype="float64")
train.sample(10)
X_train = normalize_fetures(train.iloc[:, 1:].values)
y_train = train.iloc[:, [0]].values.astype("int32")
y_train_ohe = normalize_labels(y_train)
# +
fig, _ = plt.subplots(5, 6, figsize = (15, 10))
for i, ax in enumerate(fig.axes):
ax.imshow(X_train[i].reshape(28, 28), cmap="Greys", interpolation="none")
ax.set_title("T: %d" % y_train[i])
plt.tight_layout()
# -
url = "https://raw.githubusercontent.com/makeyourownneuralnetwork/makeyourownneuralnetwork/master/mnist_dataset/mnist_test_10.csv"
test = pd.read_csv(url, header=None, dtype="float64")
test.sample(10)
X_test = normalize_fetures(test.iloc[:, 1:].values)
y_test = test.iloc[:, 0].values.astype("int32")
# # Neural Networks Classifier
#
# Author: <NAME>
# +
class NeuralNetwork:
def __init__(self, layers, learning_rate, random_state = None):
self.layers_ = layers
self.num_features = layers[0]
self.num_classes = layers[-1]
self.hidden = layers[1:-1]
self.learning_rate = learning_rate
if not random_state:
np.random.seed(random_state)
self.W_sets = []
for i in range(len(self.layers_) - 1):
n_prev = layers[i]
n_next = layers[i + 1]
m = np.random.normal(0.0, pow(n_next, -0.5), (n_next, n_prev))
self.W_sets.append(m)
def activation_function(self, z):
return 1 / (1 + np.exp(-z))
def fit(self, training, targets):
inputs0 = inputs = np.array(training, ndmin=2).T
assert inputs.shape[0] == self.num_features, \
"no of features {0}, it must be {1}".format(inputs.shape[0], self.num_features)
targets = np.array(targets, ndmin=2).T
assert targets.shape[0] == self.num_classes, \
"no of classes {0}, it must be {1}".format(targets.shape[0], self.num_classes)
outputs = []
for i in range(len(self.layers_) - 1):
W = self.W_sets[i]
inputs = self.activation_function(W.dot(inputs))
outputs.append(inputs)
errors = [None] * (len(self.layers_) - 1)
errors[-1] = targets - outputs[-1]
#print("Last layer", targets.shape, outputs[-1].shape, errors[-1].shape)
#print("Last layer", targets, outputs[-1])
#Back propagation
for i in range(len(self.layers_) - 1)[::-1]:
W = self.W_sets[i]
E = errors[i]
O = outputs[i]
I = outputs[i - 1] if i > 0 else inputs0
#print("i: ", i, ", E: ", E.shape, ", O:", O.shape, ", I: ", I.shape, ",W: ", W.shape)
W += self.learning_rate * (E * O * (1 - O)).dot(I.T)
if i > 0:
errors[i-1] = W.T.dot(E)
def predict(self, inputs, cls = False):
inputs = np.array(inputs, ndmin=2).T
assert inputs.shape[0] == self.num_features, \
"no of features {0}, it must be {1}".format(inputs.shape[0], self.num_features)
for i in range(len(self.layers_) - 1):
W = self.W_sets[i]
input_next = W.dot(inputs)
inputs = activated = self.activation_function(input_next)
return np.argmax(activated.T, axis=1) if cls else activated.T
def score(self, X_test, y_test):
y_test = np.array(y_test).flatten()
y_test_pred = nn.predict(X_test, cls=True)
return np.sum(y_test_pred == y_test) / y_test.shape[0]
# -
# # Run neural net classifier on small dataset
#
# ### Training set size: 100, testing set size 10
# +
nn = NeuralNetwork([784,100,10], 0.3, random_state=0)
for i in np.arange(X_train.shape[0]):
nn.fit(X_train[i], y_train_ohe[i])
nn.predict(X_train[2]), nn.predict(X_train[2], cls=True)
print("Testing accuracy: ", nn.score(X_test, y_test), ", training accuracy: ", nn.score(X_train, y_train))
#list(zip(y_test_pred, y_test))
# -
# # Load full MNIST dataset.
#
# ### Training set size 60,000 and test set size 10,000
#
# Original: http://yann.lecun.com/exdb/mnist/
#
# CSV version:
# training: https://pjreddie.com/media/files/mnist_train.csv
# testing: https://pjreddie.com/media/files/mnist_test.csv
# +
train = pd.read_csv("../data/MNIST/mnist_train.csv", header=None, dtype="float64")
X_train = normalize_fetures(train.iloc[:, 1:].values)
y_train = train.iloc[:, [0]].values.astype("int32")
y_train_ohe = normalize_labels(y_train)
print(y_train.shape, y_train_ohe.shape)
test = pd.read_csv("../data/MNIST/mnist_test.csv", header=None, dtype="float64")
X_test = normalize_fetures(test.iloc[:, 1:].values)
y_test = test.iloc[:, 0].values.astype("int32")
# -
# ## Runt the Neural Network classifier and measure performance
timer.reset()
nn = NeuralNetwork([784,100,10], 0.3, random_state=0)
for i in range(X_train.shape[0]):
nn.fit(X_train[i], y_train_ohe[i])
timer("training time")
accuracy = nn.score(X_test, y_test)
print("Testing accuracy: ", nn.score(X_test, y_test), ", Training accuracy: ", nn.score(X_train, y_train))
# # Effect of learning rate
# +
params = 10 ** - np.linspace(0.01, 2, 10)
scores_train = []
scores_test = []
timer.reset()
for p in params:
nn = NeuralNetwork([784,100,10], p, random_state = 0)
for i in range(X_train.shape[0]):
nn.fit(X_train[i], y_train_ohe[i])
scores_train.append(nn.score(X_train, y_train))
scores_test.append(nn.score(X_test, y_test))
timer()
plt.plot(params, scores_test, label = "Test score")
plt.plot(params, scores_train, label = "Training score")
plt.xlabel("Learning Rate")
plt.ylabel("Accuracy")
plt.legend()
plt.title("Effect of learning rate")
# -
print("Accuracy scores")
pd.DataFrame({"learning_rate": params, "train": scores_train, "test": scores_test})
# # Effect of Epochs
# +
epochs = np.arange(20)
learning_rate = 0.077
scores_train, scores_test = [], []
nn = NeuralNetwork([784,100,10], learning_rate, random_state = 0)
indices = np.arange(X_train.shape[0])
timer.reset()
for _ in epochs:
np.random.shuffle(indices)
for i in indices:
nn.fit(X_train[i], y_train_ohe[i])
scores_train.append(nn.score(X_train, y_train))
scores_test.append(nn.score(X_test, y_test))
timer("test score: %f, training score: %f" % (scores_test[-1], scores_train[-1]))
plt.plot(epochs, scores_test, label = "Test score")
plt.plot(epochs, scores_train, label = "Training score")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.legend(loc = "lower right")
plt.title("Effect of Epochs")
print("Accuracy scores")
pd.DataFrame({"epochs": epochs, "train": scores_train, "test": scores_test})
# -
# # Effect of size (num of nodes) of the single hidden layer
# +
num_layers = 50 * (np.arange(10) + 1)
learning_rate = 0.077
scores_train, scores_test = [], []
timer.reset()
for p in num_layers:
nn = NeuralNetwork([784, p,10], learning_rate, random_state = 0)
indices = np.arange(X_train.shape[0])
for i in indices:
nn.fit(X_train[i], y_train_ohe[i])
scores_train.append(nn.score(X_train, y_train))
scores_test.append(nn.score(X_test, y_test))
timer("size: %d, test score: %f, training score: %f" % (p, scores_test[-1], scores_train[-1]))
plt.plot(num_layers, scores_test, label = "Test score")
plt.plot(num_layers, scores_train, label = "Training score")
plt.xlabel("Hidden Layer Size")
plt.ylabel("Accuracy")
plt.legend(loc = "lower right")
plt.title("Effect of size (num of nodes) of the hidden layer")
print("Accuracy scores")
pd.DataFrame({"layer": num_layers, "train": scores_train, "test": scores_test})
# -
# # Effect of using multiple hidden layers
# +
num_layers = np.arange(5) + 1
learning_rate = 0.077
scores_train, scores_test = [], []
timer.reset()
for p in num_layers:
layers = [100] * p
layers.insert(0, 784)
layers.append(10)
nn = NeuralNetwork(layers, learning_rate, random_state = 0)
indices = np.arange(X_train.shape[0])
for i in indices:
nn.fit(X_train[i], y_train_ohe[i])
scores_train.append(nn.score(X_train, y_train))
scores_test.append(nn.score(X_test, y_test))
timer("size: %d, test score: %f, training score: %f" % (p, scores_test[-1], scores_train[-1]))
plt.plot(num_layers, scores_test, label = "Test score")
plt.plot(num_layers, scores_train, label = "Training score")
plt.xlabel("No of hidden layers")
plt.ylabel("Accuracy")
plt.legend(loc = "upper right")
plt.title("Effect of using multiple hidden layers, \nNodes per layer=100")
print("Accuracy scores")
pd.DataFrame({"layer": num_layers, "train": scores_train, "test": scores_test})
# -
# # Rotation
img = scipy.ndimage.interpolation.rotate(X_train[110].reshape(28, 28), -10, reshape=False)
print(img.shape)
plt.imshow(img, interpolation=None, cmap="Greys")
# +
epochs = np.arange(10)
learning_rate = 0.077
scores_train, scores_test = [], []
nn = NeuralNetwork([784,250,10], learning_rate, random_state = 0)
indices = np.arange(X_train.shape[0])
timer.reset()
for _ in epochs:
np.random.shuffle(indices)
for i in indices:
for rotation in [-10, 0, 10]:
img = scipy.ndimage.interpolation.rotate(X_train[i].reshape(28, 28), rotation, cval=0.01, order=1, reshape=False)
nn.fit(img.flatten(), y_train_ohe[i])
scores_train.append(nn.score(X_train, y_train))
scores_test.append(nn.score(X_test, y_test))
timer("test score: %f, training score: %f" % (scores_test[-1], scores_train[-1]))
plt.plot(epochs, scores_test, label = "Test score")
plt.plot(epochs, scores_train, label = "Training score")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.legend(loc = "lower right")
plt.title("Trained with rotation (+/- 10)\n Hidden Nodes: 250, LR: 0.077")
print("Accuracy scores")
pd.DataFrame({"epochs": epochs, "train": scores_train, "test": scores_test})
# -
# # Which charaters NN was most wrong about?
missed = y_test_pred != y_test
pd.Series(y_test[missed]).value_counts().plot(kind = "bar")
plt.title("No of mis classification by digit")
plt.ylabel("No of misclassification")
plt.xlabel("Digit")
fig, _ = plt.subplots(6, 4, figsize = (15, 10))
for i, ax in enumerate(fig.axes):
ax.imshow(X_test[missed][i].reshape(28, 28), interpolation="nearest", cmap="Greys")
ax.set_title("T: %d, P: %d" % (y_test[missed][i], y_test_pred[missed][i]))
plt.tight_layout()
img = scipy.ndimage.imread("/Users/abulbasar/Downloads/9-03.png", mode="L")
print("Original size:", img.shape)
img = normalize_fetures(scipy.misc.imresize(img, (28, 28)))
img = np.abs(img - 0.99)
plt.imshow(img, cmap="Greys", interpolation="none")
print("Predicted value: ", nn.predict(img.flatten(), cls=True))
| 01 Neural Network using Numpy.ipynb |