code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:miniconda3]
# language: python
# name: conda-env-miniconda3-py
# ---
# # Question 3: What factors influence the income of an Airbnb apartment in Berlin?
# Now that we know how much money one can make with Airbnb apartments in Berlin, I'd like to answer another question. The third question around the Airbnb Berlin data set is about the __factors that influence a listing's income__. Knowing these things in advance allows Airbnb apartment owners to __make the most out of their renting offers__.
#
#
# Similar to the previous two, the analysis is divided into the following __five steps__:
# * 1 - Load libraries
# * 2 - Import & understand data
# * 3 - Pre-process data
# * 4 - Model data
# * 5 - Analyze data & evaluate results
# ## 1 - Load Libraries
# +
import numpy as np
import pandas as pd
import math
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
# ## 2 - Import & Understand Data
# +
# Import listings data (Source: http://insideairbnb.com/get-the-data.html)
df_listings = pd.read_csv('../data/listings_berlin.csv')
# Preview data
print(df_listings.shape)
pd.options.display.max_columns = None
df_listings.head(1)
# -
# ### 2.1 - Overview
# The data set holds many columns which could potentially have an influence on the monthly income of an apartment. Even though one may guess what might be important or not it's important to __confirm or reject__ a subjective assessment with __objective results__. In contrast to the previous two analyses - in which I limited the number of columns to use to a small subset - I'm going to look at a __much larger subset of variables__ now.
# ## 3 - Pre-Process Data
# I want to be able to __combine previous and future results__. Therefore I will apply __similar pre-processing steps__ as used when answering "__Question 1__" and "__Question 2__". Nonetheless, additional pre-processing steps are included in the following section as well.
# ### 3.1 - Property Types
# Removing listings with property types other than "Apartment".
# +
# Check distriutions of property types
property_count = df_listings["property_type"].value_counts().sum()
# Keep only apartments in analysis (~ 90% of listings)
n_all = df_listings.shape[0]
df_listings = df_listings[df_listings["property_type"] == "Apartment"]
print("{} records were removed from the Airbnb Berlin listings data frame.".format(n_all - df_listings.shape[0]))
# -
# ### 3.2 - Missing Values
# Once again I will check the data set for missing values.
# Define columns to use in analysis
custom_cols = ['accommodates','bathrooms','bedrooms','beds','cleaning_fee','extra_people','guests_included',
'host_is_superhost','host_response_rate','instant_bookable','monthly_price','minimum_nights',
'neighbourhood_group_cleansed','number_of_reviews','price','review_scores_rating','reviews_per_month',
'weekly_price']
# Check for missing values
df_listings[custom_cols].isnull().sum()
# Now that I'd like to work with more columns it's important to think about the __meaning of missing values__ in each column. I will __keep the missing values for all price columns__ as the corresponding listings simply don't offer different rates. Both "__cleaning_fee__" and "__reviews_per_month__" are going to be __imputed with 0s__, as cleaning fees on the one hand are optinal extra charges and reviews on the other hand are not available thus can bet set to zero. The __superhost flag__ will also be set to __zero__. Last but not least I will run a __mean imputation on "host_response_rate" and "review_scores_rating"__ assuming that listings with missing values perform at least as good as average.
#
# Some of the columns require __additional re-engineering__ steps before their missing values can be imputed. These steps will be included where necessary.
# Imputing missing values for "reviews_per_month"
df_listings["reviews_per_month"] = df_listings["reviews_per_month"].fillna(0)
# Imputing missing values for "host_is_superhost"
df_listings["host_is_superhost"] = df_listings["host_is_superhost"].astype(str).apply(lambda x: 1 if x == "t" else 0)
# Imputing missing values for "review_scores_rating"
df_listings["review_scores_rating"] = df_listings["review_scores_rating"].fillna(df_listings["review_scores_rating"].mean())
# Imputing missing values for "cleaning_fee"
df_listings["cleaning_fee"] = df_listings["cleaning_fee"].fillna("$0.00")
# Imputing missing values for "host_response_rate"
df_listings["host_response_rate"] = df_listings["host_response_rate"].astype(str)
df_listings["host_response_rate"] = df_listings["host_response_rate"].apply(lambda x: x.replace('%','').replace(',',''))
df_listings["host_response_rate"] = df_listings["host_response_rate"].astype(float)
df_listings["host_response_rate"] = df_listings["host_response_rate"].fillna(df_listings["host_response_rate"].mean())
# Let's check if everything looks as expected.
# Check again for missing values
df_listings[custom_cols].isnull().sum()
# ### 3.3 - Neighbourhoods
# Only small adjusted to the values of the "neighbourhood_groups" column thus completely optional (as in the previous notebook).
# Adjust values of "neighbourhood_group_cleansed"
df_listings["neighbourhood_group_cleansed"] = df_listings["neighbourhood_group_cleansed"].astype(str).str.replace("Charlottenburg-Wilm.", "Charlottenburg-Wilmersdorf")
df_listings["neighbourhood_group_cleansed"] = df_listings["neighbourhood_group_cleansed"].astype(str).str.replace(" ", "")
# Also, I'd like to know __how many listings__ there are in each neighbourhood and if that may have an influence on the actual income. Let's add two more colums to the data set.
# Add listing counts per neighbourhood (group)
df_listings["listings_in_hood"] = df_listings["id"].groupby(df_listings["neighbourhood_cleansed"]).transform("size")
df_listings["listings_in_hood_group"] = df_listings["id"].groupby(df_listings["neighbourhood_group_cleansed"]).transform("size")
# ### 3.4 - Bookings Per Month
# An income estimation requires as to know the number of bookings a listing receives. The calculation happens according to the way I did it when answering "__Question 1__" and "__Question 2__". Please refere to the previous notebook(s) for more details.
# Define adjusted proxy for number of bookings per months
df_listings["max_bookings_per_month"] = (30.42 / df_listings["minimum_nights"]).fillna(0)
df_listings["bookings_per_month"] = df_listings[["max_bookings_per_month","reviews_per_month"]].min(axis=1)
df_listings = df_listings.drop("max_bookings_per_month", axis=1)
# I'm also going to add a column showing the __number of bookings per neighbourhood and neighbourhood group__ as a form of competition assessment. We can even add something like the __number bookings per listing__ by using the previously defined columns for listing counts.
# Add booking counts per neighbourhood (group)
df_listings["bookings_in_hood"] = df_listings["bookings_per_month"].groupby(df_listings["neighbourhood_cleansed"]).transform("sum")
df_listings["bookings_in_hood_group"] = df_listings["bookings_per_month"].groupby(df_listings["neighbourhood_group_cleansed"]).transform("sum")
# Add booking counts per neighbourhood (group)
df_listings["bookings_per_listing_in_hood"] = df_listings["bookings_in_hood"] / df_listings["listings_in_hood"]
df_listings["bookings_per_listing_in_hood_group"] = df_listings["bookings_in_hood_group"] / df_listings["listings_in_hood_group"]
# ### 3.5 Accomodates
# Check if accomodates can be less than guests included
(df_listings["accommodates"] < df_listings["guests_included"]).sum()
# Apparently, __95 listings offer space for more people than the apartment can actually accomodate__. I'm going to adjust these cases so that the number of guests included doesn't exceed the number of people an apartment can accomodate.
# +
# Adjust values
df_listings["guests_included"] = np.where(df_listings["accommodates"] < df_listings["guests_included"],
df_listings["accommodates"], df_listings["guests_included"])
# Check if adjustment was successful
(df_listings["accommodates"] < df_listings["guests_included"]).sum()
# -
# ### 3.6 - Price
# Apparently, all __columns related to price information__ are formatted as objects with a "$" sign in them, I need to convert them to strings in order to remove it. Finally, I can easily convert them to __float values__.
# +
# Convert price columns used in calculation to float
price_cols = ["price","weekly_price","monthly_price","cleaning_fee","extra_people"]
for col in price_cols:
df_listings[col] = df_listings[col].astype(str)
df_listings[col] = df_listings[col].apply(lambda x: x.replace('$','').replace(',',''))
df_listings[col] = df_listings[col].astype(float)
# -
# I'm also going to introducde an __adjusted version of "price"__ which includes additinal costs. It's going to represent the __price per person__.
# Create "price_per_person" column (based on total number of possible guests)
df_listings["price_per_person"] = ((
df_listings["price"] + df_listings["cleaning_fee"] +
df_listings["extra_people"] * (df_listings["accommodates"] - df_listings["guests_included"]))
/
df_listings["guests_included"])
# Summary statistics
df_listings["price_per_person"].describe()
# As seen in the previous notebook, the summary statistics already indicate that prices of Airbnb listings may have a couple of __outliers__. Again, I'm going to deal with this later.
# ### 3.7 - Instant Bookings
# This column simply needs a dtype conversion from object to integer so it can be used in later modeling steps.
# Re-engineering of "instant_bookable"
df_listings["instant_bookable"] = df_listings["instant_bookable"].astype(str).apply(lambda x: 1 if x == "t" else 0)
# ## 4 - Model Data
# Finally, I'm going to __model the actual income__ of an Airbnb listing. This step is simply a copy of the one used when working on "__Question 2__" (please go back there for further details).
# +
# Calculate each listing's income per booking
booking_income = []
for (p,wp,mp,n) in zip(df_listings["price"], df_listings["weekly_price"], df_listings["monthly_price"], df_listings["minimum_nights"]):
price = 0.0
if n > 29:
if not math.isnan(mp):
months = n / 30.0
price += mp * math.floor(months)
if not math.isnan(wp):
weeks = ((months - math.floor(months)) * 30.0) / 7.0
days = (weeks - math.floor(weeks)) * 7.0
price += (wp * math.floor(weeks)) + (n * days)
else:
days = (months - math.floor(months)) * 30.0
price += p * days
elif not math.isnan(wp):
weeks = n / 7.0
days = (weeks - math.floor(weeks)) * 7.0
price += (wp * math.floor(weeks)) + (p * days)
else:
price += p * n
elif n > 6:
if not math.isnan(wp):
weeks = n / 7.0
days = (weeks - math.floor(weeks)) * 7.0
price += (wp * math.floor(weeks)) + (p * days)
else:
price += p * n
else:
price += p * n
booking_income.append(price)
# Add to data frame and check summary statistics
df_listings["income_per_booking"] = booking_income
df_listings["income_per_booking"].describe()
# -
# Let's __transform__ the income per booking column by multiplying it with the __number of bookings per month__. We then get the new column "__income_per_month__".
# Convert the price per booking to a price per months
df_listings["income_per_month"] = df_listings["income_per_booking"] * df_listings["bookings_per_month"]
df_listings["income_per_month"].describe()
# ## 5 - Analyze Data & Evaluate Results
# Before starting off with the analysis I will again remove outliers in "income_per_month" from the data set.
# Remove upper 5% from data set
n_all = df_listings.shape[0]
df_listings = df_listings[df_listings["income_per_month"] <= df_listings["income_per_month"].quantile(.95)]
print("{} records were removed from the Airbnb Berlin listings data frame.".format(n_all - df_listings.shape[0]))
# I've decided to focus my analysis on __correlations between a listings income per month and other columns__ in the data set. It's important to note that the results will __not necessarily reveal causations__. Nonetheless, correlations can be a good __starting point__ in order to understand what things to look for to make the most out one's listing on Airbnb.
# Define columns to use in analysis
custom_cols = ['income_per_month','accommodates','bathrooms','bedrooms','beds','bookings_per_month','bookings_in_hood',
'bookings_in_hood_group','bookings_per_listing_in_hood','bookings_per_listing_in_hood_group',
'cleaning_fee','extra_people','guests_included','host_is_superhost','host_response_rate','instant_bookable',
'listings_in_hood','listings_in_hood_group','minimum_nights','number_of_reviews','price','price_per_person',
'review_scores_rating','reviews_per_month']
# +
# Get correlations between columns
corrs = df_listings[custom_cols].dropna(axis=0).corr()#.sort_values("income_per_month", ascending=False)
# Plot correlations
plt.subplots(figsize=(20,10))
sns.set(font_scale=1)
corr_plot = sns.heatmap(corrs, yticklabels=custom_cols, xticklabels=custom_cols,
cmap="OrRd", cbar=True, annot=True, square=True, fmt=".1f")
corr_plot.set_title("Heatmap of Correlations Between Subset of Columns in Airbnb Berlin Data Set", fontsize=14);
plt.tight_layout()
# Save plot
corr_fig = corr_plot.get_figure()
corr_fig.savefig('../results/correlations.png')
# -
# The heatmap above shows the correlation between the pre-defined subset of columns from the listings data set. The __primary focus__ of the analysis is on the heatmap's __first column__ which shows all correlations with a listing's income per month. Let's evaluate the __observations__:
#
# * Correlations are __mainly positive__, despite a small negative correlation with the number of listings in the neighbourhood
# * The __strongest__ positive correlations (0.6) can be observed between the __number of bookings/reviews per month__. Both columns are strongly related to each other as the number of bookings per month was based on the number of reviews per month. However, this observation __doesn't really help__ us to gain more information due to the relationship between number of bookings and income __by definition__.
# * A __moderate__ positive correlation with __number of reviews__ can as well be explained by its connections to the number of reviews per month.
# * A __weak__ positive correlation with the __superhost__ flag indicates that being a good host in general leads to a higher income for a listing (according to Aribnb a host has to meet certain criteria which indicate a higher quality to become a superhost)
# * A __weak__ positive correlations with the __number of people__ allowed in the apartment as well as the __number of beds__ finally indicates that __larger apartments earn more money__.
#
# To summarize, the correlations __don't reveal any suprising findings__. Nonetheless, people who are interested in renting out apartments in Berlin could potentially __benefit from offering larger apartments__ as well as following Airbnb guidelines to __become a superhost__ (please see: https://www.airbnb.de/superhost).
| notebooks/q3_airbnb_income_features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c2e61c32-1a5d-4666-8526-5e85367bd21d", "showTitle": false, "title": ""}
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "1dde5164-5373-4eab-a4de-ba6e8fd0f07e", "showTitle": false, "title": ""}
# ## Run BERT-Large training workload
#
# In the beginning, data preprocessing will take some minutes. Once the preprocessing is done, the training workload can output throughput performance number in real time. It takes around five hours to complete the entire training process on Standard_F32s_v2 instance. The precise elapsed time depends on instance type and whether is Intel-optimized TensorFlow.
#
#
# **Note:** ***If you click "Stop Execution" for running training/inference cells, and then run training/inference again immediately. You may see lower performance number, because another training/inference is still on-going.***
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "96f083b7-e3a3-4cdf-8474-7f304b641fd7", "showTitle": false, "title": ""}
import os
import subprocess
from pathlib import Path
def run_training():
training = '/tmp/training.sh'
with open(training, 'w') as f:
f.write("""#!/bin/bash
# BERT-Large Training
# Install necessary package
sudo apt-get update
sudo apt-get install zip -y
sudo apt-get -y install git
sudo apt-get install -y libblacs-mpi-dev
sudo apt-get install -y numactl
# Remove old materials if exist
rm -rf /TF/
mkdir /TF/
# Create ckpt directory
mkdir -p /TF/BERT-Large-output/
# Download IntelAI benchmark
cd /TF/
wget https://github.com/IntelAI/models/archive/refs/tags/v1.8.1.zip
unzip v1.8.1.zip
cores_per_socket=$(lscpu | awk '/^Core\(s\) per socket/{ print $4 }')
numa_nodes=$(lscpu | awk '/^NUMA node\(s\)/{ print $3 }')
export SQUAD_DIR=/dbfs/home/TF/bert-large/SQuAD-1.1
export BERT_LARGE_MODEL=/dbfs/home/TF/bert-large/wwm_uncased_L-24_H-1024_A-16
export BERT_LARGE_OUTPUT=/TF/BERT-Large-output/
export PYTHONPATH=$PYTHONPATH:.
function run_training_without_numabind() {
python launch_benchmark.py \
--model-name=bert_large \
--precision=fp32 \
--mode=training \
--framework=tensorflow \
--batch-size=4 \
--benchmark-only \
--data-location=$BERT_LARGE_MODEL \
-- train-option=SQuAD DEBIAN_FRONTEND=noninteractive config_file=$BERT_LARGE_MODEL/bert_config.json init_checkpoint=$BERT_LARGE_MODEL/bert_model.ckpt vocab_file=$BERT_LARGE_MODEL/vocab.txt train_file=$SQUAD_DIR/train-v1.1.json predict_file=$SQUAD_DIR/dev-v1.1.json do-train=True learning-rate=1.5e-5 max-seq-length=384 do_predict=True warmup-steps=0 num_train_epochs=0.1 doc_stride=128 do_lower_case=False experimental-gelu=False mpi_workers_sync_gradients=True
}
function run_training_with_numabind() {
intra_thread=`expr $cores_per_socket - 2`
python launch_benchmark.py \
--model-name=bert_large \
--precision=fp32 \
--mode=training \
--framework=tensorflow \
--batch-size=4 \
--mpi_num_processes=$numa_nodes \
--num-intra-threads=$intra_thread \
--num-inter-threads=1 \
--benchmark-only \
--data-location=$BERT_LARGE_MODEL \
-- train-option=SQuAD DEBIAN_FRONTEND=noninteractive config_file=$BERT_LARGE_MODEL/bert_config.json init_checkpoint=$BERT_LARGE_MODEL/bert_model.ckpt vocab_file=$BERT_LARGE_MODEL/vocab.txt train_file=$SQUAD_DIR/train-v1.1.json predict_file=$SQUAD_DIR/dev-v1.1.json do-train=True learning-rate=1.5e-5 max-seq-length=384 do_predict=True warmup-steps=0 num_train_epochs=0.1 doc_stride=128 do_lower_case=False experimental-gelu=False mpi_workers_sync_gradients=True
}
# Launch Benchmark for training
cd /TF/models-1.8.1/benchmarks/
if [ "$numa_nodes" = "1" ];then
run_training_without_numabind
else
run_training_with_numabind
fi """)
os.chmod(training, 555)
p = subprocess.Popen([training], stdin=None, stdout=subprocess.PIPE, stderr=subprocess.PIPE)
directory_to_second_numa_info = Path("/sys/devices/system/node/node1")
if directory_to_second_numa_info.exists():
# 2 NUMA nodes
for line in iter(p.stdout.readline, ''):
if b"Reading package lists..." in line or b"answer: [UNK] 1848" in line:
print("\t\t\t\t Preparing data ......", end='\r')
if b"INFO:tensorflow:examples/sec" in line:
print("\t\t\t\t Training started, current real-time throughput (examples/sec) : " + str(float(str(line).strip("\\n'").split(' ')[1])*2), end='\r')
if line == b'' and p.poll() != None:
break
else:
# 1 NUMA node
for line in iter(p.stdout.readline, ''):
if b"Reading package lists..." in line or b"answer: [UNK] 1848" in line:
print("\t\t\t\t Preparing data ......", end='\r')
if b"INFO:tensorflow:examples/sec" in line:
print("\t\t\t\t Training started, current real-time throughput (examples/sec) : " + str(line).strip("\\n'").split(' ')[1], end='\r')
if line == b'' and p.poll() != None:
break
p.stdout.close()
run_training()
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c35a944c-41a1-44bc-be3a-146859d6a0df", "showTitle": false, "title": ""}
# ## Check whether is Intel-optimized TensorFlow
#
# This is a simple auxiliary script tool to check whether the installed TensorFlow is Intel-optimized TensorFlow. "Ture" represents Intel-optimized TensorFlow.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c3ba1baf-4329-4b05-8071-12ccec717f16", "showTitle": false, "title": ""}
# Print version, and check whether is intel-optimized
import tensorflow
print("tensorflow version: " + tensorflow.__version__)
from packaging import version
if (version.parse("2.5.0") <= version.parse(tensorflow.__version__)):
from tensorflow.python.util import _pywrap_util_port
print( _pywrap_util_port.IsMklEnabled())
else:
from tensorflow.python import _pywrap_util_port
print(_pywrap_util_port.IsMklEnabled())
| integrations/ml/databricks/benchmark/benchmark_tensorflow_bertlarge_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import urllib.request
import json
with open('/home/ej/github/naver_api_info.txt', 'r') as f:
client_id = str(f.readline().replace('\n',''))
client_secret = str(f.readline().replace('\n',''))
encText = urllib.parse.quote("Google")
encDisp = urllib.parse.quote("100")
url = "https://openapi.naver.com/v1/search/news?query={}&display={}".format(encText, encDisp)
# url = "https://openapi.naver.com/v1/search/blog.xml?query=" + encText # xml 결과
request = urllib.request.Request(url)
request.add_header("X-Naver-Client-Id", client_id)
request.add_header("X-Naver-Client-Secret", client_secret)
response = urllib.request.urlopen(request)
rescode = response.getcode()
if(rescode == 200):
response_body = response.read()
else:
print("Error Code:" + rescode)
# -
data = json.loads(response_body.decode('utf-8'))
data['display']
dic = data['items']
dic[0]
for each_news in dic:
print(each_news['link'])
| workspace/naver_api_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# metadata:
# interpreter:
# hash: cf85b414d3663472de89104473c842eaab37d7b845999caf56a47ccda76ea2f8
# name: python3
# ---
# # Student Alcohol Consumption
# ### Introduction:
#
# This time you will download a dataset from the UCI.
#
# ### Step 1. Import the necessary libraries
import pandas as pd
import numpy as np
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/04_Apply/Students_Alcohol_Consumption/student-mat.csv).
url = 'https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/04_Apply/Students_Alcohol_Consumption/student-mat.csv'
# ### Step 3. Assign it to a variable called df.
# +
alco_consmp = pd.read_csv(url, sep=',')
alco_consmp.head(5)
# -
# ### Step 4. For the purpose of this exercise slice the dataframe from 'school' until the 'guardian' column
alco_consmp_df = alco_consmp.loc[:5,'school':'guardian']
alco_consmp_df
# ### Step 5. Create a lambda function that will capitalize strings.
cap_fun = lambda elemento: elemento.capitalize()
alco_consmp_df[['Mjob', 'Fjob']].applymap(cap_fun)
# ### Step 6. Capitalize both Mjob and Fjob
# +
for e in ['Mjob', 'Fjob']:
alco_consmp_df[e] = alco_consmp_df[e].apply(cap_word)
alco_consmp_df
# -
# ### Step 7. Print the last elements of the data set.
alco_consmp_df.tail(5)
# ### Step 8. Did you notice the original dataframe is still lowercase? Why is that? Fix it and capitalize Mjob and Fjob.
# +
# I don't see that it is still in lowercase letters Looool
# -
# ### Step 9. Create a function called majority that returns a boolean value to a new column called legal_drinker (Consider majority as older than 17 years old)
# +
majority = lambda x: True if x > 17 else False
alco_consmp_df['legal_drinker'] = alco_consmp_df['age'].apply(majority)
alco_consmp_df
# -
# ### Step 10. Multiply every number of the dataset by 10.
# ##### I know this makes no sense, don't forget it is just an exercise
# +
lista_ints = []
for item in alco_consmp_df.loc[:,'school':'legal_drinker']:
if alco_consmp_df[item].dtype == alco_consmp_df['age'].dtype:
lista_ints.append(item)
alco_consmp_df[lista_ints] = alco_consmp_df[lista_ints].apply(lambda x: x*10)
alco_consmp_df
# -
| 2_Ejercicios/Primera_parte/Entregables/2. Ejercios_Mod1/2020_12_18/4_Apply/Students_Alcohol_Consumption/Student_Alc_Cons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # https://raw.githubusercontent.com/eco32i/biodata/master/data/gradtimes.txt
# # Graduation time by training program
# <p>COB vs NEURO (see slide)</p>
print("Hellow")
print(1+1)
x = 'asdf' + 'wer'
print(x)
x = [3,4,'asdf']
print x[-1]
f = open('../data/gradtimes.txt','r')
print(f.readline())
# +
f = open('../data/gradtimes.txt','r')
numStudents = 0
for line in f:
numStudents = numStudents + 1
print(numStudents)
# +
f = open('/Users/justinjee/Desktop/neuro.txt','r')
allTimes = []
print(len(allTimes))
for line in f:
(program, time) = line.split()
allTimes.append(float(time))
print(len(allTimes))
# -
a ='5'
b = '8'
print(a+b)
print(int(a)+float(b))
print(sum(allTimes)/len(allTimes))
# +
f = open('/Users/justinjee/Desktop/nonneuro.txt','r')
allTimes = []
for line in f:
(program, time) = line.split()
allTimes.append(float(time))
print(sum(allTimes)/len(allTimes))
# -
import random
r = random.sample(allTimes,84)
print sum(r)/len(r)
# +
n = 0
for i in range(100000):
r = random.sample(allTimes,84)
if sum(r)/len(r)>=5.93:
n+=1
print n/100000.0
# -
# # standard deviation vs. standard deviation of the mean
# +
def mean(a):
return sum(a)/len(a)
a = [1,2,3]
print mean(a)
def std(a):
er = 0
for i in a:
er+= (mean(a)-i)**2
return (float(er)/len(a))**0.5
print std(a)
# -
r = random.sample(allTimes,84)
print(mean(r))
print(std(r))
print(std(allTimes))
# +
means = []
for i in range(10):
r = random.sample(allTimes,84)
means.append(mean(r))
print std(allTimes)
print std(means) #standard error of the mean!
# -
print std(allTimes)/(84)**0.5 #theoretical standard error of the mean
| sessions/Stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
# +
# CrossEntropyError
def cross_entropy_error(y, t):
delta=1e-7
return -np.sum(t*np.log(y+delta))
# -
class simpleNet:
def __init__(self):
self.W = np.random.randn(2, 3)
def predict(self, x):
return x @ self.W
def softmax(self, x):
c = np.max(x)
exp_x = np.exp(x - c)
sum_exp_x = np.sum(exp_x)
return exp_x / sum_exp_x
def loss(self, x, t):
z = self.predict(x)
y = self.softmax(z)
loss = cross_entropy_error(y, t)
return loss
net = simpleNet()
net.W
x = np.array([0.6, 0.9])
p = net.predict(x)
p
np.argmax(p)
t = np.array([0, 0, 1])
net.loss(x, t)
| ch4/simpleNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1-minute introduction to Jupyter ##
#
# A Jupyter notebook consists of cells. Each cell contains either text or code.
#
# A text cell will not have any text to the left of the cell. A code cell has `In [ ]:` to the left of the cell.
#
# If the cell contains code, you can edit it. Press <kbd>Enter</kbd> to edit the selected cell. While editing the code, press <kbd>Enter</kbd> to create a new line, or <kbd>Shift</kbd>+<kbd>Enter</kbd> to run the code. If you are not editing the code, select a cell and press <kbd>Ctrl</kbd>+<kbd>Enter</kbd> to run the code.
# Before you turn this problem in, make sure everything runs as expected. First, **restart the kernel** (in the menubar, select Kernel$\rightarrow$Restart) and then **run all cells** (in the menubar, select Cell$\rightarrow$Run All).
#
# Make sure you fill in any place that says `YOUR CODE HERE` or "YOUR ANSWER HERE", as well as your name and collaborators below:
NAME = ""
COLLABORATORS = ""
# ---
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "d7581d01cccef86c931a03276638654e", "grade": false, "grade_id": "cell-a965c0c70630d0c7", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # Assignment 10: Advanced modular programming, testing
#
# In this assignment, you should write your code in a **readable** way, and **modularise** chunks of code that would otherwise be repeated.
#
# Modularising a section of program code means to reorganise it in a way that allows it to be reused in another part of the code easily, usually in the form of a function.
#
# Your function definitions should have appropriate **docstrings**.
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "d40de432d74890ebf5f204493b78d5d5", "grade": false, "grade_id": "cell-1725022ebfb43d85", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Part 1: Testing a Sudoku solver
#
# A 3×3 sudoku solver receives the following generated input:
# -
puzzle = [[None, 9, None],
[3, None, 7],
[None, 1, None],
]
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "97e82441e3163279b8b3b4dfe12fe4da", "grade": false, "grade_id": "cell-9729b16d2ccecc36", "locked": true, "schema_version": 3, "solution": false, "task": false}
# It is called to solve the puzzle, and it returns the following output:
# -
solution = [[4, 9, 2],
[3, 5, 7],
[8, 1, 6],
]
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "7bb68360f1a54c094d9ecc0e80e9ead4", "grade": false, "grade_id": "cell-a29b9923da204a58", "locked": true, "schema_version": 3, "solution": false, "task": false}
# **Task:** Write `assert` statements to _verify_ that the solution is correct.
#
# A correct solution should have all its rows, columns, and diagonals sum to 15.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "a58f1419d002aedf4f9e049f1182ed5c", "grade": true, "grade_id": "cell-e03c120171de668c", "locked": false, "points": 3, "schema_version": 3, "solution": true, "task": false}
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "9d757d211009f3f0f24ae8ee2368e79b", "grade": false, "grade_id": "cell-02630fc3daa783fd", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Part 2: Setting a `debug` flag for `print`ing
#
# Now that you have learned how to use keyword arguments, let’s try using them for something potentially useful: `print()` statements for debugging.
#
# When you don’t need them, you are afraid to remove them, because re-inserting them again is so annoying. Is there a better alternative to commenting out multiple `print()` statements, then uncommenting them again?
#
# Yes! Let’s use keyword arguments to determine whether the function should print debug statements or not.
#
# A sample `text_numeral()` function is provided in the code cell below.
#
# ### Task: Add keyword arguments
#
# Edit the code so that the `print()` statements are executed only when the following function call is made:
#
# >>> text_numeral(367, debug=True)
# prev_key: 0
# this_key: 1
# prev_key: 1
# [...] <-- represents truncated output
# selected key: 7
# output: three hundred and sixty-seven
# remainder: 0
# 'three hundred and sixty-seven' <-- this is the output value (str type)
#
# but not when it is called without the `debug` keyword argument:
#
# >>> text_numeral(367)
# 'three hundred and sixty-seven' <-- this is the output value (str type)
#
# **Hint:** The keyword arguments are passed to the function as a dictionary. you can use the dict’s `.get()` method to retrieve the keyword argument, and set it to a fallback value if it is not provided.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "99044a50152e7f6cb8204e8557b8b4f3", "grade": false, "grade_id": "cell-06d02db7c32d0c73", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Edit this code to print() only when debug=True is passed to the function as a keyword argument
def text_numeral(num:int, **kwargs):
'''
Takes in a number below 1000 and converts it to text form.
Usage:
text_numeral(num)
'''
if num >= 1000:
raise ValueError('input must be less than 1000.')
numeral = {1: 'one', 2: 'two', 3: 'three', 4: 'four', 5: 'five',
6: 'six', 7: 'seven', 8: 'eight', 9: 'nine', 10: 'ten',
11: 'eleven', 12: 'twelve', 13: 'thirteen', 14: 'fourteen', 15: 'fifteen',
16: 'sixteen', 17: 'seventeen', 18: 'eighteen', 19: 'nineteen', 20: 'twenty',
30: 'thirty',
40: 'forty',
50: 'fifty',
60: 'sixty',
70: 'seventy',
80: 'eighty',
90: 'ninety',
}
output = ''
while num > 0:
# Select largest key that is still smaller than num
# Since we need to check that this_key is greater than num,
# we need to store prev_key and prev_word since they are the
# actual key and value we want.
key, word = 100, '<PASSWORD>'
prev_key, prev_word = 0, None
for this_key, this_word in numeral.items():
print(f'prev_key: {prev_key}')
print(f'this_key: {this_key}')
if prev_key <= num < this_key:
print(f'match: {prev_key} <= {num} < {this_key}')
key, word = prev_key, prev_word
prev_key, prev_word = this_key, this_word
print(f'selected key: {key}')
# Build up the output string
if key == 100:
hundred_count, num = divmod(num, key)
output += f'{numeral[hundred_count]} {word}'
if num > 0:
output += ' and '
else:
output += word
num -= key
if num > 0:
output += '-'
print(f'output: {output}')
print(f'remainder: {num}')
if num == 0:
return output
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "0917033406f15ccde3ba54f268eac95e", "grade": true, "grade_id": "cell-0d323b2b11660e53", "locked": true, "points": 2, "schema_version": 3, "solution": false, "task": false}
# Autograding test for no debug kwarg
# If your code works, this cell should produce no output when run
import sys
from io import StringIO
capture = StringIO()
temp = sys.stdout
sys.stdout = capture
result = text_numeral(367)
sys.stdout = temp
assert capture.getvalue().strip() == '', 'Function printed output when it should be silent'
assert result == 'three hundred and sixty-seven', f'output from function \'{result}\' is incorrect'
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "7a801a7e1776706c270917bf6ce8b668", "grade": true, "grade_id": "cell-1c657f7fe577e4be", "locked": true, "points": 2, "schema_version": 3, "solution": false, "task": false}
# Autograding test for debug=True kwarg
# If your code works, this cell should produce no output when run
import sys
from io import StringIO
capture = StringIO()
temp = sys.stdout
sys.stdout = capture
result = text_numeral(367, debug=True)
sys.stdout = temp
for each in ('selected key: 7',
'output: three hundred and sixty-seven',
'remainder: 0',
):
assert each in capture.getvalue().strip(), f'expected print statement not found: {each}'
assert result == 'three hundred and sixty-seven', f'output from function \'{result}\' is incorrect'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "36250801a8bedd36d630469d88db7dc6", "grade": false, "grade_id": "cell-c531f9728731d658", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Part 2: Write a `try-except` structure
#
# The Open Library API (application programming interface) offers programmers a way to search for books and retrieve book information in their program. The following functions are provided below:
#
# 1. `search()` takes in a `searchterm` (`str`) argument and returns search results from the Open Library API
# 2. `save_to_csv()` takes in `list_of_dict` (`list`) and `filepath` (`str`) positional arguments, and writes the title, author name, and first year of publication of the results to a CSV file located in `filepath`.
# +
# Run this cell to use the functions below
# Do not modify the functions
from openlibrary import search, save_to_csv
# -
help(search)
help(save_to_csv)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "f4a8e8281038d4de3b9e6ff138886398", "grade": false, "grade_id": "cell-72d1267cba41c5cc", "locked": true, "schema_version": 3, "solution": false, "task": false}
# A developer is trying to use these two functions to save search results to a CSV file in _his_ function `search_and_save()` (in the code cell below). He wants the search results to be saved into a folder representing the current date, to keep the files organised. But he keeps encountering a `FileNotFoundError`.
#
# This `FileNotFoundError` happens because the directory `'20200420'` does not exist yet. He needs to call `os.mkdir(folder)` to create the folder. But `os.mkdir()` will raise a `FileExistsError` if the directory already exists.
#
# **Task:** Modify the code in `search_and_save()` below by adding `try-except` statements so that these errors are handled successfully.
#
# **Hint:** You may have to write nested `try-except` statements to handle further Exceptions that occur in the `except`, `else`, or `finally` statements
# + deletable=false nbgrader={"cell_type": "code", "checksum": "7bb49c93deef30f91eabd0964958c7d5", "grade": false, "grade_id": "cell-89d8ab25ade8cd31", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Modify the code in this cell with try-except-else statements
# To take appropriate actions when exceptions occur
# And write the CSV file successfully
import os
from datetime import date
def search_and_save(searchterm):
'''
Search for results from Open Library API for searchterm,
and print the results to a file in a folder.
Folder is named by date, YYYYMMDD format.
File path: ./<YYYYMMDD>/<searchterm>.csv
'''
result = search(searchterm)
folder = date.today().strftime('%Y%m%d')
filepath = f'./{folder}/{searchterm}.csv'
save_to_csv(result['docs'], filepath)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "c0722f74c4e0c201996dd40ef680cee4", "grade": true, "grade_id": "cell-77b553ad9dd7a273", "locked": true, "points": 4, "schema_version": 3, "solution": false, "task": false}
# AUTOGRADING: Test for file creation and correct data
import os, shutil
from datetime import date
import unittest
from pathlib import Path
def readfile(filepath):
if os.path.exists(filepath):
with open(filepath, 'r') as f:
data = [row.strip().split(',') for row in f.readlines()]
return data.pop(0), data
def remove_notexist_raise(filepath):
try:
assert os.path.exists(filepath), f'File not found: {filepath}'
except Exception:
raise
else:
shutil.rmtree(Path(filepath).parent)
class testCheck(unittest.TestCase):
def testNotRaiseError(self):
searchterm = '50 Shades of Grey'
try:
search_and_save(searchterm)
except Exception as e:
print(f'{e.__class__} Test failed, exception raised: {e}')
finally:
folder = date.today().strftime('%Y%m%d')
filepath_to_check = f'./{folder}/{searchterm}.csv'
if os.path.exists(filepath_to_check):
shutil.rmtree(f'./{folder}')
def testFileCreated(self):
searchterm = '<NAME>'
folder = date.today().strftime('%Y%m%d')
filepath = f'./{folder}/{searchterm}.csv'
try:
search_and_save(searchterm)
except Exception as e:
print(f'{e.__class__} Test failed, exception raised: {e}')
else:
assert os.path.isfile(filepath), f'File not found: {filepath}'
finally:
remove_notexist_raise(filepath)
def testDataHeader(self):
searchterm = '<NAME>'
folder = date.today().strftime('%Y%m%d')
filepath = f'./{folder}/{searchterm}.csv'
try:
search_and_save(searchterm)
except Exception as e:
print(f'{e.__class__} Test failed, exception raised: {e}')
else:
header, data = readfile(filepath)
assert header == ['title', 'author_name', 'first_publish_year'], \
f'Wrong header ({header} != [\'title\', \'author_name\', \'first_publish_year\'])'
finally:
remove_notexist_raise(filepath)
def testDataFormat(self):
searchterm = '<NAME>'
folder = date.today().strftime('%Y%m%d')
filepath = f'./{folder}/{searchterm}.csv'
try:
search_and_save(searchterm)
except Exception as e:
print(f'{e.__class__} Test failed, exception raised: {e}')
else:
header, data = readfile(filepath)
for row in data:
assert len(row) == 3, f'Wrong number of elements in row ({len(row)}), expected 3\n' + \
f'Row data: {row}'
finally:
remove_notexist_raise(filepath)
# unittest.main looks at sys.argv and first parameter is what started IPython or Jupyter
# exit=False prevents unittest.main from shutting down the kernel process
result = unittest.main(argv=['ignored'], exit=False)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "6261f5013b0bc976c1bc84c3ac2e5879", "grade": false, "grade_id": "cell-ad1672e66728167f", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # Feedback and suggestions
#
# Any feedback or suggestions for this assignment?
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "8b2c5aa73b1a793329d6b92780994acd", "grade": true, "grade_id": "cell-63f8f1cb7ff65bb1", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false}
# YOUR ANSWER HERE
| Assignment 10/assignment_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="jxK1_8f1dvrc"
# <div>
# <img src="https://drive.google.com/uc?export=view&id=1vK33e_EqaHgBHcbRV_m38hx6IkG0blK_" width="350"/>
# </div>
#
# #**Artificial Intelligence - MSc**
# ET5003 - MACHINE LEARNING APPLICATIONS
#
# ###Instructor: <NAME>
# ###ET5003_Etivity-1
# + id="LqXD_IwUQuBF" cellView="form"
#@title Current Date
Today = '2021-08-22' #@param {type:"date"}
# + id="uzDKau31OjVO"
#@markdown ---
#@markdown ### Enter your details here:
Student_ID = "19187289" #@param {type:"string"}
Student_full_name = "<NAME>" #@param {type:"string"}
#@markdown ---
# + id="r39xGZckTpKx"
#@title Notebook information
Notebook_type = 'Etivity' #@param ["Example", "Lab", "Practice", "Etivity", "Assignment", "Exam"]
Version = 'Draft' #@param ["Draft", "Final"] {type:"raw"}
Submission = False #@param {type:"boolean"}
# + [markdown] id="80m304lUefG4"
# ## MNIST dataset
# + [markdown] id="Bs8mHGcidHSa"
#
#
# The MNIST database is a dataset of handwritten digits that has been and is extensively used in machine learning. There are $10$ classes, each image is $28\times28$ pixels and, therefore, each input is $x_i\in\mathbb{R}^{784}$.
# + [markdown] id="ailycCq5epj2"
# ## Task
# + [markdown] id="a-yNAxhUemjM"
# You have to extend the code to manage any arbitrary number of classes, in other words you have to implement a general-recipe multinomial logistic classifier and Bayesian multinomial logistic classifier.
#
# You must then select 3 digits at random and perform the following task.
#
# 1. Your goal is to use Bayesian multinomial logistic regression (as in the road-sign notebook) to solve this classification problem.
#
# 2. You can downsize the training dataset (e.g., 40% training and 60%testing) if the computation of the posterior takes too much time in your computer.
#
# 3. Use the posterior uncertainty to detect the instances (digits) in the test set that are hard to classify and remove them from the test-set.
#
# 4. Then you need to compute again the accuracy of the general-recipe logistic regression on the remaining (non-difficult) instances and comment on the result.
#
# 5. In practice, the task is to use uncertainty estimation to detect the difficult instances in the test-set. This is equivalent to refuse to classify all high-uncertainty instances or, in other words, when we are uncertain we say "I don't know" and we do not return any class. In this way, you will learn how uncertainty can be used to make safer decisions, by detecting the instances that are difficult to classify.
#
# + [markdown] id="nMRKRTQZe5fW"
# ## Libraries
# + id="IxusAui7AX_f"
# Suppressing Warnings:
import warnings
warnings.filterwarnings("ignore")
# + id="MQOfGMQpdHSb"
# Import libraries
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
import scipy.optimize as optimize
from scipy.special import erf
import pandas as pd
import numpy as np
import seaborn as sns
from tqdm import tqdm
from skimage.io import imread, imshow
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
import arviz as az
from scipy.io import loadmat
import pymc3 as pm
import random
from IPython.display import HTML
import pickle
import theano as tt
import cv2
from sklearn.utils import shuffle
from skimage.color import rgb2gray
from matplotlib import pyplot
# + id="P5-qMSjpAQ-9"
# Setting a seed:
np.random.seed(123)
# + [markdown] id="r4hSuwkUfVQb"
# ## Dataset
# + [markdown] id="w99Pc66YdHSd"
# ### Loading the MNIST dataset
# + colab={"base_uri": "https://localhost:8080/"} id="CYFWAbXVzynp" outputId="b833c4c2-d02a-4939-f038-de5d902f7fab"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="g4rCnS4vdHSd" outputId="ed720e55-3441-4ba7-8f20-b6ca34b941f3"
# Path, copy the path from your Drive
path = '/content/drive/MyDrive/Colab Notebooks/Data/mnist/'
# MNIST Data
train_data = path + 'mnist_train.csv'
test_data = path + 'mnist_test.csv'
print('train_data',train_data)
# train data
df_train = pd.read_csv(train_data)
X_train_original = df_train.drop("label",axis=1).values
y_train = df_train.label.values
print(X_train_original.shape)
# test data
df_test = pd.read_csv(test_data)
X_test_original = df_test.drop("label",axis=1).values
y_test = df_test.label.values
print(X_test.shape)
# + id="sILcmvTDsn8m"
# + colab={"base_uri": "https://localhost:8080/"} id="O2ubJ_WoAqBh" outputId="e78ceffb-42df-4aaa-9d6f-ebcd3204dcd1"
# Normalizing the inputs by making it float between 0.0 and 1.0:
print("X_train", X_train)
X_train_normal = X_train_original/255
print("X_train_normal", X_train)
X_test = X_test_original/255
# Printing the new input range of values:
minv = np.min(X_train)
maxv = np.max(X_train)
print(minv,maxv)
# + [markdown] id="SR6HpkWndHSe"
# ### Description of Data:
# + colab={"base_uri": "https://localhost:8080/"} id="sibN1Vv1dHSf" outputId="88785c62-da8d-4d02-fd11-0eb7c70da05c"
# Get the number of examples
n_train, n_test = len(X_train), len(X_test)
# get the shape of a digit image
image_shape = X_train.shape[1]
# unique classes/labels in the training dataset.
print("y_train", y_train)
# Let's see what classes exist in my training set
distinct_classes = set(y_train)
print("distinct_classes", distinct_classes)
# Get the number of classes
n_classes = len(distinct_classes)
print("Number of Training examples =", n_train)
print("Number of Test examples =", n_test)
print("Image input shape =", image_shape)
print("Number of classes =", n_classes)
# + [markdown] id="6HQDSvrRKZF6"
# ### Class Distribution:
# + id="XG8GdlpBKdCt" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="33eb9f62-37c9-4e5c-b4fd-87ca23167ae2"
fig, ax = plt.subplots()
ind = np.arange(n_classes)
n, bins, patches = ax.hist(y_train, n_classes)
ax.set_xlabel('classes')
ax.set_ylabel('counts')
ax.set_title(r'Histogram of Digit images')
plt.show()
# + [markdown] id="EyLWw3nsLCtk"
# ## Downsampling
# + [markdown] id="2U1lFEwhLKBf"
# ### Randomly selecting 3 of the 10 Digit Classes
# + id="0EeRZZWdLRPT" colab={"base_uri": "https://localhost:8080/"} outputId="13ce95ea-020a-4373-b6cd-86833198a516"
# We select the number of Classes we want:
n_selected_classes = 3
# Empty list to append the random digit classes we select:
classes = []
# We select 3 digits at random and make sure they are unique:
while len(classes) < n_selected_classes:
# Randomly draw a digit from 0-9:
num2choose = np.random.randint(0,10)
# Append the digit if it's not already in our list of classes:
if num2choose not in classes:
classes.append(num2choose)
print('classes', classes)
# Sorting the Classes smallest to largest
classes.sort()
# print classes selected
print('classes', classes)
# + id="2M8R5NqKMB_M" colab={"base_uri": "https://localhost:8080/"} outputId="3bef8bb9-aa30-4a3f-ca59-1acffb8e142b"
# The number of instances we'll keep for each of our 3 digits:
inst_class = 1000
# Loop to randomly sample the instances for each digit:
inputs, labels = [], []
for r in classes:
imgs = X_train_normal[np.where(y_train==r)[0],:]
print("imgs.shape", imgs.shape[0])
# randonly shuffle and then select instance classes from the set
_input = imgs[np.random.permutation(imgs.shape[0]),:][0:inst_class,:]
print("_input", _input)
inputs.append(_input)
_labels = np.ones(inst_class)*r
print("_labels",_labels)
labels.append(_labels)
# Shaping inputs and labels in the right format
X_train = np.vstack(inputs).astype(np.float64)
y_train = np.hstack(labels)
# + [markdown] id="_6-YHrQQMicy"
# New Classes Distribution
# + id="RA300COaMxWm" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="5e1d5519-1db7-4289-a3ea-1eea3c42fee5"
# new histogram
fig, ax = plt.subplots()
ind = np.arange(n_classes)
n, bins, patches = ax.hist(y_train, n_classes)
ax.set_xlabel('classes')
ax.set_ylabel('counts')
ax.set_title(r'Histogram of Digit images')
plt.show()
# + id="eFgP4xugMvJm"
# plot digits
def plot_digits(instances, images_per_row=5, **options):
size = 28
images_per_row = min(len(instances), images_per_row)
print('instances length', len(instances))
images = [instance.reshape(size,size) for instance in instances]
n_rows = (len(instances) - 1) // images_per_row + 1
row_images = []
n_empty = n_rows * images_per_row - len(instances)
images.append(np.zeros((size, size * n_empty)))
for row in range(n_rows):
rimages = images[row * images_per_row : (row + 1) * images_per_row]
row_images.append(np.concatenate(rimages, axis=1))
image = np.concatenate(row_images, axis=0)
plt.imshow(image, cmap='gist_yarg', **options)
plt.axis("off")
# + colab={"base_uri": "https://localhost:8080/", "height": 537} id="PY23H3OdNjgf" outputId="d8201117-b4c7-46ed-b81a-002804537166"
# plot 98 example images
plt.figure(figsize=(9,9))
example_images = X_train[2:100]
plot_digits(example_images, images_per_row=10)
plt.show()
# + id="zeEG-LGOM4fJ" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="dd6cb584-2dea-4ebf-98ac-c443f004a45a"
# Show a few instances from each Digit:
plt.figure(figsize=(8,8))
# Selecting a few label indices from each of the 3 classes to show:
label_indices = 2
# Plotting 'original' image
plot_digits(X_train[label_indices:100],images_per_row=9)
plt.title("Original", fontsize=14)
# + [markdown] id="FsAOnOcNNG_V"
# ### Splitting the Training data into both Training and Validation Sets:
#
# - Although this is the Training set, we can still set aside some samples (for instance 20%) of the 1,500 instances we have for Model Validation purposes.
#
#
# - With that Validation Set, we can then select the amount of Uncertainty we are happy with from our Model to use out of sample on other unseen data.
#
#
# - We can then test out how well our decision performs on the Test Set that we put aside earlier.
# + id="YdsmyVAtPXNn"
### Split tha dataset in training and validation sets
# choose the fraction of your validation data from the training set
w = 0.20
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=w, random_state=0)
# Shuffling the training instaces around to randomize the order of inputs to the model:
X_train, y_train = shuffle(X_train, y_train)
# + id="qXwJwP0iPxhi" colab={"base_uri": "https://localhost:8080/"} outputId="86c46598-f8ad-4e2f-835d-7990352ef153"
# print shape of your validation and training set
# Shape of the validation and training set
validation_set_shape = X_val.shape
training_set_shape = X_train.shape
print("Training Set Shape =", training_set_shape)
print("Validation Set Shape =", validation_set_shape)
# + [markdown] id="aOroY1QoP8DY"
# ### Encoding the Class labels for the Probabilistic ML Model:
#
# This is an example:
#
# - **[1,0,0]** for first digit
# - **[0,1,0]** for second digit
# - **[0,0,1]** for third digit
# + [markdown] id="rjUaqWTqQIcp"
# ### General-Recipe ML
# + id="QzgdivxfQNv5" colab={"base_uri": "https://localhost:8080/"} outputId="2290926d-f540-4491-9854-69883dbb34f0"
# model
model_log = LogisticRegression(random_state=0, max_iter=2000, C=100, solver='lbfgs', multi_class='multinomial').fit(X_train, y_train)
# Classification:
y_pred_log = model_log.predict(X_val)
y_pred_logi_prob = model_log.predict_proba(X_val)
# Maybe taking the maximum probability
# in any of the classes for each observation
prob_classmax = np.max(y_pred_logi_prob,axis=1)
# Computing the Accuracy:
accuracy_score(y_pred_log, y_val)
# + [markdown] id="3uQG6JsOQxH5"
# ### Probabilistic Multinomial Logistic Regression:
# + id="W3jzczJzRAtT" colab={"base_uri": "https://localhost:8080/"} outputId="5a3bca0f-3945-4a6a-8422-4f0a21d94ff7"
np.sort(prob_classmax)
# + colab={"base_uri": "https://localhost:8080/"} id="7y6zsz5BWrf4" outputId="55564d10-9fa3-43c7-b037-ec99cdea09c6"
# probability of general-recipe logistic regression in wrong instances
prob_classmax[y_pred_log!=y_val]
# + colab={"base_uri": "https://localhost:8080/"} id="doyYTm7AW-W5" outputId="e4f65375-8e89-4ecd-f1ca-d97c2e00b8f1"
prob_classmax[y_pred_log!=y_val]
# + [markdown] id="irlmUNw7Q5YL"
# The Multinomial Logistic Regression has some parameters:
#
# - $\alpha$, which is the intercept term:
#
# - $\beta$, which is a vector of coefficients which give a weighting to the importance of each input feature:
#
# + id="1o7mbKWmRhz5"
import sklearn.preprocessing
## We use LabelBinarizer to transfor classes into counts
# neg_label=0, pos_label=1
y_2_bin = sklearn.preprocessing.LabelBinarizer().fit_transform(y_train.reshape(-1,1))
nf = X_train.shape[1]
# number of classes
nc = len(classes)
# floatX = float32
floatX = tt.config.floatX
init_b = np.random.randn(nf, nc-1).astype(floatX)
init_a = np.random.randn(nc-1).astype(floatX)
with pm.Model() as multi_logistic:
# Prior
β = pm.Normal('beta', 0, sigma=100, shape=(nf, nc-1), testval=init_b)
α = pm.Normal('alpha', 0, sigma=100, shape=(nc-1,), testval=init_a)
# we need to consider nc-1 features because the model is not identifiable
# the softmax turns a vector into a probability that sums up to one
# therefore we add zeros to go back to dimension nc
# so that softmax returns a vector of dimension nc
β1 = tt.tensor.concatenate([np.zeros((nf,1)),β ],axis=1)
α1 = tt.tensor.concatenate([[0],α ],)
# Likelihood
mu = pm.math.matrix_dot(X_train,β1) + α1
# It doesn't work if the problem is binary
p = tt.tensor.nnet.nnet.softmax(mu)
observed = pm.Multinomial('likelihood', p=p, n=1, observed=y_2_bin)
# + id="aj6Uzc05Rhtr" colab={"base_uri": "https://localhost:8080/"} outputId="a2e9659c-f7b0-4cce-8baf-0b3587a44c6f"
y_2_bin
# + id="2MFH4gwlRhrB" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="7f0fba28-6564-4157-feca-c59f9724d8eb"
with multi_logistic:
#approx = pm.fit(300000, method='advi') # takes longer
approx = pm.fit(3000, method='advi')
# + id="WNmJvYc4Rho7" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="d9b7dd5f-6e2c-4e86-b9b3-aca69c567839"
plt.plot(approx.hist)
# + id="XXh5GXJsRhmr"
dd = 300
posterior = approx.sample(draws=dd)
# + colab={"base_uri": "https://localhost:8080/", "height": 244} id="st_Jcd4hbJR6" outputId="9c6a9e79-a0ba-4f98-a20a-74fa07dba1d6"
## The softmax function transforms each element of a collection by computing the exponential
# of each element divided by the sum of the exponentials of all the elements.
from scipy.special import softmax
#select an image in the test set
i = 10
#i = random.randint(0, dd)
#select a sample in the posterior
s = 100
#s = random.randint(0, dd)
beta = np.hstack([np.zeros((nf,1)), posterior['beta'][s,:] ])
alpha = np.hstack([[0], posterior['alpha'][s,:] ])
image = X_val[i,:].reshape(28,28)
plt.figure(figsize=(2,2))
plt.imshow(image,cmap="Greys_r")
np.set_printoptions(suppress=True)
print("test image #" + str(i))
print("posterior sample #" + str(s))
print("true class=", y_val[i])
print("classes: " + str(classes))
print("estimated prob=",softmax((np.array([X_val[i,:].dot(beta) + alpha])))[0,:])
# + id="JqA6iCOgbRP_"
# Bayesian prediction
# return the class that has the highest posterior probability
y_pred_Bayesian=[]
for i in range(X_val.shape[0]):
val=np.zeros((1,len(classes)))
for s in range(posterior['beta'].shape[0]):
beta = np.hstack([np.zeros((nf,1)), posterior['beta'][s,:] ])
alpha = np.hstack([[0], posterior['alpha'][s,:] ])
val = val + softmax((np.array([X_val[i,:].dot(beta) + alpha])))
mean_probability = val/posterior['beta'].shape[0]
y_pred_Bayesian.append( np.argmax(mean_probability))
# + colab={"base_uri": "https://localhost:8080/"} id="-NtQ5LGRbVQ2" outputId="48a2fa9b-d212-4dfc-d329-778ca6c2d0ff"
print(y_pred_Bayesian)
# + colab={"base_uri": "https://localhost:8080/"} id="AZrukzIpbZYo" outputId="29c79dc4-001a-4597-f588-f6f04c54bd86"
# recall the classes we are using
print(classes)
# + colab={"base_uri": "https://localhost:8080/"} id="I48TaTcUbZl5" outputId="b59a57a9-3690-425a-d1c1-b599ed2edffc"
# prediction array (using classes)
nn = 10 # just an example
np.array(classes)[y_pred_Bayesian[0:nn]]
# + colab={"base_uri": "https://localhost:8080/"} id="Vve2qdxPbZqk" outputId="b51e5ec0-4824-48a9-bf45-3dce47d3317d"
# using validation: y_val
print("Accuracy=", accuracy_score(np.array(classes)[y_pred_Bayesian], y_val))
# + id="uIMzQskzbZt5"
y_predB=[]
for i in range(X_val.shape[0]):
#print(i)
val=[]
for s in range(posterior['beta'].shape[0]):
beta = np.hstack([np.zeros((nf,1)), posterior['beta'][s,:] ])
alpha = np.hstack([[0], posterior['alpha'][s,:] ])
val.append(softmax((np.array([X_val[i,:].dot(beta) + alpha])))[0,:])
#mean probability
valmean = np.mean(val,axis=0)
#class with maximum mean probability
classmax = np.argmax(valmean)
#ranks
ranks = np.array(val.copy())
ranks = ranks *0 #init
colmax = np.argmax(np.array(val),axis=1)
ranks[np.arange(0,len(colmax)),colmax]=1
y_predB.append( [classmax, valmean[classmax], np.std(ranks,axis=0)[classmax]])
y_predB= np.array(y_predB)
# + colab={"base_uri": "https://localhost:8080/"} id="iLb5u5GnbngG" outputId="81c889d4-0f11-41d6-de6c-44cce46b2ddb"
# prediction array
mm = 10
y_predB[0:mm,:]
# + colab={"base_uri": "https://localhost:8080/"} id="dovEEoTObnjZ" outputId="64a840fb-85b6-423e-e62a-526934f9b634"
#sorting in descending order
difficult = np.argsort(-y_predB[:,2])
y_predB[difficult[0:mm],:]
# + colab={"base_uri": "https://localhost:8080/"} id="17GQnD56bnmY" outputId="01f7f626-304c-4d2e-808d-41f72b555604"
#probability of general-recipe logistic regression in wrong instances
prob_classmax[y_pred_log != y_val]
# + colab={"base_uri": "https://localhost:8080/"} id="_6sigSj3bnpA" outputId="fed25eb7-185a-45a5-8753-b26c05e70d54"
y_predB[y_pred_log != y_val,:]
# + colab={"base_uri": "https://localhost:8080/"} id="Bf32hHGybnsG" outputId="e07a20cc-8caf-45d6-e8bd-22e320783b9d"
## Difficult & easy instances
easy = np.argsort(y_predB[:,2])
print("Accuracy in easy instances =", accuracy_score(y_pred_log[easy[0:100]], y_val[easy[0:100]]))
difficult = np.argsort(-y_predB[:,2])
print("Accuracy in difficult instances =", accuracy_score(y_pred_log[difficult[0:100]], y_val[difficult[0:100]]))
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="JTFBbrD6bnu6" outputId="8a1c8855-119b-47fe-8fcb-20c1faa51d0c"
# show 10 random 'easy' images
fig, axs = plt.subplots(2,5, figsize=(15, 6))
fig.subplots_adjust(hspace = .2, wspace=.001)
axs = axs.ravel()
for i in range(10):
index = easy[i]
image = X_val[index,:].reshape(28,28)
axs[i].axis('off')
axs[i].imshow(image,cmap="Greys_r")
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="DCxOA9b8bZww" outputId="dc10c942-e4c7-40a5-f0cd-81384e7016bd"
# show 10 random 'difficult' images
fig, axs = plt.subplots(2,5, figsize=(15, 6))
fig.subplots_adjust(hspace = .2, wspace=.001)
axs = axs.ravel()
for i in range(10):
index = difficult[i]
image = X_val[index,:].reshape(28,28)
axs[i].axis('off')
axs[i].imshow(image,cmap="Greys_r")
# + id="6RkezZMdbZ0W"
# + [markdown] id="NcshsLOGRPrk"
# ## Summary
# + [markdown] id="XTc4pYKGRR60"
# Populate this section with all of your findings and comments fron the discussion with your peers.
| Week-1/Etivity_1_MarkDevine_19187289.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HurleyJames/GoogleColabExercise/blob/master/Using_Word2Vec_Embeddings.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="JtYLDuNmwnnv" colab_type="text"
# # News Category Classification using Word2Vec embeddings
#
# University of Leeds
#
# COMP5623 Artificial Intelligence
#
# ---
#
# We will use two Python libraries:
#
# 1. **sklearn** a machine learning library
#
# 2. **Gensim** is a library for unsupervised topic modeling and natural language processing, using modern statistical machine learning
#
# ---
# + [markdown] id="ElMkuzUm7sOR" colab_type="text"
#
# # 1. Dataset preparation - 20 Newsgroups
#
# We will use sklearn to download **20 Newsgroups** (http://qwone.com/~jason/20Newsgroups/), a public available dataset of approximately 20,000 newsgroup posts, partitioned across 20 different newsgroups. We will only load 3 categories for this example.
# + id="OAFHvAKcwyH5" colab_type="code" colab={}
categories = [
'comp.graphics',
'sci.med',
'rec.sport.baseball'
]
# + [markdown] id="OAFCFVY_yiCD" colab_type="text"
# Load the subset of the 20 Newsgroups dataset.
# + id="sD84pKuGwxKF" colab_type="code" colab={}
from sklearn.datasets import fetch_20newsgroups
train_set = fetch_20newsgroups(
subset='train',
categories=categories,
shuffle=True,
)
# + [markdown] id="jlWIPMyey2DF" colab_type="text"
# Look at some sample data.
# + id="zWYaiPTKy5sR" colab_type="code" colab={}
train_set.target_names
# + id="7nLorBMe5iJC" colab_type="code" colab={}
print("Total number of news articles:", len(train_set.data))
# + id="gciDCHgE5tyj" colab_type="code" colab={}
print("\n".join(train_set.data[0].split("\n")[:10]))
# + id="IiQMJBoS55J4" colab_type="code" colab={}
print(train_set.target_names[train_set.target[0]])
# + [markdown] id="Nm2WQXeq7wee" colab_type="text"
# # 2. Training a Word2Vec model
# + [markdown] id="wqlR8EQVyxlo" colab_type="text"
# Now we will train a Word2Vec model which we will use to map each word in each news article to a feature representation. While the best models are trained on very large amounts of data, due to resources, we will use a model trained on this small corpus.
#
# First we pre-process the data into lists of words.
# + id="l7LUiNMJyjmQ" colab_type="code" colab={}
parsed_train_data = []
for article in train_set.data:
parsed_train_data.append(article.replace('\n',' ').split())
# + [markdown] id="WSLO2Un5-6rN" colab_type="text"
# Train the model.
# + id="6AnxiOTQzxbe" colab_type="code" colab={}
from gensim.models import Word2Vec
feature_length = 50
word2vec_model = Word2Vec(
sentences=parsed_train_data,
window=5,
sg=1, # Use skip-gram
size=feature_length
)
# + [markdown] id="GtoED80r_VNq" colab_type="text"
# How good is it? Sanity check...
# + id="GVfIoTa0_ZsK" colab_type="code" colab={}
word2vec_model.most_similar("disease", topn=5)
# + id="4DAo_PMyz6em" colab_type="code" colab={}
word2vec_model.most_similar("baseball", topn=5)
# + [markdown] id="XktqzfRhAEtf" colab_type="text"
# What is the feature representation for one word?
# + id="LRznixyfAH8X" colab_type="code" colab={}
word2vec_model["baseball"]
# + [markdown] id="x9axD6V5_3Bh" colab_type="text"
# # 3. Performing classification on news articles using feature embeddings
#
# Finally, we will represent each news article as a block of word features, and perform classification on the embedded representations.
#
# As an over-simplification of the problem (for the purposes of illustration), we will choose the N first words to represent an article so that all our article sizes are fixed.
# + id="P9WJeqvnE_Ph" colab_type="code" colab={}
article_size = 13 * feature_length
# + id="YWYY3zHXIUkt" colab_type="code" colab={}
import numpy as np
def embed_article(article, cutoff):
# Save the feature representation for each word in the article
embedded_article = []
for word in article:
try:
embedded_article.append(word2vec_model[word])
except(KeyError): # Ignore words not in the model vocabulary
pass
return np.array(embedded_article).flatten()[:cutoff]
# + id="bvjf7Ow1AzJs" colab_type="code" colab={}
embedded_train_set = []
for article in parsed_train_data:
embedded_train_set.append(embed_article(article, article_size))
# + [markdown] id="1w5UV_V6B2Xw" colab_type="text"
# Now we can try training a simple linear classifier to classify the news articles into their categories.
# + id="wN4G1K_bBS_n" colab_type="code" colab={}
from sklearn.linear_model import SGDClassifier
# + id="5ZWDSn8ZB0Kn" colab_type="code" colab={}
linear_classifier = SGDClassifier()
linear_classifier.fit(
embedded_train_set,
train_set.target,
)
# + [markdown] id="Mim8hOp5Gy5R" colab_type="text"
# Can it predict on new articles?
# + id="G5MmRfMDEMhs" colab_type="code" colab={}
new_articles = [
# Article one - from latest NVidia news
"Nvidia RTX 2080 Ti Cyberpunk 2077 GPU is revealed – but you can’t buy one." +
"77 limited edition graphics cards are to be given away in a competition" +
"The mysterious Cyberpunk 2077-themed GPU Nvidia recently teased has been revealed," +
"and the reality of the graphics card may be a touch disappointing for some folks, in that" +
"it isn’t a new model – and you won’t be able to buy one." +
"The card is simply a GeForce RTX 2080 Ti (and appears to be exactly the same model," +
"and shroud design) decked out with the Cyberpunk 2077 colors and logo, which admittedly" +
"looks pretty cool, but isn’t the GeForce RTX 2080 Ti Super" +
"AMD confirms ‘Nvidia killer’ graphics card will be out in 2020",
# Article two - from Health -> Oncology
"Breast cancer test could predict chances of disease return 20 years later, study shows" +
"Molecular nature of a woman’s breast cancer determines how their disease could progress," +
"not just for the first five years, but also later,' says researcher" +
"A new test could identify breast cancers that are likely to return more than 20 years later" +
"development that might herald an era of personalised medicine." +
"The way a patient’s cancer will progress can be determined by categorising molecular and" +
"genetic markers of breast tumours into 11 subtypes, University of Cambridge researchers found." +
"Following around 2,000 women over 20 years, the team funded by the Cancer Research charity found" +
"some women with initially aggressive cancers had a low chance of tumours returning after five years."
]
# Parse and embed
parsed_na = [a.replace('\n',' ').split() for a in new_articles]
embedded_new_articles = [embed_article(a, article_size) for a in parsed_na]
# + id="4hudi7IQIAIB" colab_type="code" colab={}
predicted = linear_classifier.predict(embedded_new_articles)
# + id="wUIjj0DrKEfQ" colab_type="code" colab={}
for i, category in enumerate(predicted):
print("New article", i, " predicted cateogry =>", train_set.target_names[category])
# + id="U5XJkagZKcCr" colab_type="code" colab={}
| Using_Word2Vec_Embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bus
#
# This bus has a passenger entry and exit control system to monitor the number of occupants it carries and thus detect when there is too high a capacity.
#
# At each stop the entry and exit of passengers is represented by a tuple consisting of two integer numbers.
# ```
# bus_stop = (in, out)
# ```
# The succession of stops is represented by a list of these tuples.
# ```
# stops = [(in1, out1), (in2, out2), (in3, out3), (in4, out4)]
# ```
#
# ## Goals:
# * lists, tuples
# * while/for loops
# * minimum, maximum, length
# * average, standard deviation
#
# ## Tasks
# 1. Calculate the number of stops.
# 2. Assign to a variable a list whose elements are the number of passengers at each stop (in-out),
# 3. Find the maximum occupation of the bus.
# 4. Calculate the average occupation. And the standard deviation.
#
# +
# variables
# +
# 1. Calculate the number of stops.
# +
# 2. Assign a variable a list whose elements are the number of passengers in each stop:
# Each item depends on the previous item in the list + in - out.
# +
# 3. Find the maximum occupation of the bus.
# +
# 4. Calculate the average occupation. And the standard deviation.
# -
| bus/bus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numpy and Pandas
import numpy as np
from numpy import random
# ## 1. Numpy random
# `np.random.choice` generates a random sample from a given 1-D array
np.random.choice(a=['a', 'b', 'c', 'd', 'e', 'f'], size=60, replace=True)
np.random.choice(a=['a', 'b', 'c', 'd'], p=[0.1, 0.1, 0.7, 0.1], size=100)
np.random.choice(a=np.arange(1, 101), size=100, replace=False)
# similar to (note that this changes the array in place):
x = np.arange(1, 101)
np.random.shuffle(x)
x
# to reorder x (note that this does not change the array in place)
x = np.sort(x)
x
np.flip(x)
# 20 numbers between 0 and 10
np.random.randint(0, 10, size=20)
# We can also sample from distributions:
np.random.uniform(low=0, high=10, size=20)
np.random.normal(loc=0, scale=1, size=10)
np.random.normal(loc=[-10, 0, 10], scale=1)
x = np.random.binomial(n=1, p=.8, size=50)
x
np.sum(x)/50
# ## 2. Tiling and repeating arrays
letters = np.array(['A', 'B', 'C'])
colors = np.array(['red', 'blue', 'grey'])
colors = np.repeat(colors, repeats=3)
colors
letters = np.tile(letters, reps=3)
letters
np.repeat(np.tile(letters, reps=2), repeats=3)
np.tile(np.repeat(letters, repeats=2), reps=3)
colors + letters
letters[0] + colors[0]
np.char.add(letters, colors)
# ## 3. Creating a pandas dataframe
import pandas as pd
df = pd.DataFrame({
'letters': letters,
'colors': colors,
'stimuli': np.char.add(np.char.add(letters, "_"), colors)
})
df
df['letters']
df.loc[:,'letters']
df.loc[6,:]
df.loc[0,'letters']
df['trial'] = np.arange(1, 10)
df['participant'] = 1
df
df.shape
df.shape[0]
df['block'] = 1
df
df.loc[(df.trial > 3) & (df.trial < 7), 'block'] = 2
df.loc[df.trial > 6, 'block'] = 3
df
df['block'] = np.repeat(np.arange(1, 4), 3)
# #### Filling-in an empty dataframe
df = pd.DataFrame([]) # create an empty dataframe
df
df.shape
df = df.append({'rt': 2, 'choice': 'l', 'trial': 1, 'reward': 34}, ignore_index=True) # record some responses
df
df = df.append({'rt': 2.1, 'choice': 'k', 'trial': 2, 'reward': 36}, ignore_index=True) # record some responses
df
responses = pd.Series([2, 'l', 3, 20], index=['rt', 'choice', 'trial', 'reward'])
responses
df = df.append(responses, ignore_index=True)
df
# #### Save the dataframe to file
df.to_csv('data.csv') # save to file
# #### Shuffling a dataframe
df = pd.DataFrame({'image_number': np.arange(1, 11), 'difficulty': np.repeat(['easy','hard'], 5)})
df
df.sample(frac=1) # shaffle across all trials
df = df.sample(frac=1).reset_index(drop=True)
df
df.groupby(['difficulty']).mean()
df.groupby(['difficulty']).apply(lambda x: x.sample(frac=1)).reset_index(drop=True) # shaffle within condition
# ## Exercises
#
# 1. Create a dataframe called `stimuli`. The first column should be called `participant`, the second column `trial`, and the third one `condition`. There should be 10 trials per condition, and 3 conditions for each participant, and 4 participants. Therefore, in total the dataframe should have 4X3X10=120 rows, and 3 columns. The participant numbers should go from 1 to 4, the conditions should be labelled `['low', 'med', 'high']`, and the trials should be numbered from 1 to 30. The order of the condition should be randomized within participant.
# 2. Add a column called `probability`, which is equal to .1 when the condition is `low`, .5 when it's `med`, and .9 when it's 'high'.
# 3. Create a new dataframe called `data`, that is a copy of the `stimuli` dataframe (using the `.copy()` attribute, as in `new_dataframe = old_dataframe.copy()`). Add to `data` a column called `choice`, by generating data from a binomial distribution with parameters `n=1` and the probability that is in the probability column of each trial.
#
# **Send me your solutions** before the end of this Friday (at <EMAIL>).
#
# **NOTE**: Do not send me a notebook but export it to a Python script first: `File> Export notebook as… > Export as executable script`
# ## Solutions
import pandas as pd
import numpy as np
# +
stimuli = pd.DataFrame({
'participant': np.repeat(np.arange(1, 5), 3*10),
'condition': np.tile(np.repeat(['low', 'med', 'high'], 10), 4)
})
stimuli = stimuli.groupby(['participant']).apply(lambda x: x.sample(frac=1)).reset_index(drop=True) # shaffle within participants
stimuli['trial'] = np.tile(np.arange(1, 31), 4)
# -
stimuli.head()
# to check that there are exactly 10 trials per condition:
stimuli.groupby(['participant', 'condition']).count()
# +
stimuli['probability'] = .1
stimuli.loc[stimuli.condition == 'med', 'probability'] = .5
stimuli.loc[stimuli.condition == 'high', 'probability'] = .9
# -
stimuli.head()
# +
data = stimuli.copy()
data['choice'] = np.random.binomial(n=1, p=data.probability)
# -
# to check % success per particiapant and condition:
data.groupby(['participant', 'condition']).mean()
# or just per condition:
data.groupby(['condition']).mean()
data.describe()
data.groupby('participant').describe().loc[1]
data.groupby('participant').describe().loc[:,"trial"]
data.loc[50:100,:]
| notebooks/wpa_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="nsnFjhKKVUXB"
# # **ANALYSIS OF FINANCIAL INCLUSION IN EAST AFRICA BETWEEN 2016 TO 2018**
# + [markdown] id="6yL2ylFeUAlM"
# ##DEFINING QUESTION
# + [markdown] id="jzHACIUKN83L"
# The research problem is to figure out how we can predict which individuals are most likely to have or use a bank account.
# + [markdown] id="v07RZUoMN-wm"
# ### METRIC FOR SUCCESS
# + [markdown] id="yvN-0d9YQpGx"
# My solution procedure will be to help provide an indication of the state of financial inclusion in Kenya, Rwanda, Tanzania, and Uganda, while providing insights into some of the key demographic factors that might drive individuals financial outcomes.
# + [markdown] id="QEfzN_YiRu7j"
# ###THE CONTEXT
# + [markdown] id="D41HPsDsRvrC"
# Financial Inclusion remains one of the main obstacles to economic and human development in Africa. For example, across Kenya, Rwanda, Tanzania, and Uganda only 9.1 million adults (or 13.9% of the adult population) have access to or use a commercial bank account.
#
# Traditionally, access to bank accounts has been regarded as an indicator of financial inclusion. Despite the proliferation of mobile money in Africa and the growth of innovative fintech solutions, banks still play a pivotal role in facilitating access to financial services. Access to bank accounts enables households to save and facilitate payments while also helping businesses build up their credit-worthiness and improve their access to other financial services. Therefore, access to bank accounts is an essential contributor to long-term economic growth.
# + [markdown] id="ipV9WMzbRwQL"
# ### EXPERIMENTAL DESIGN TAKEN
# + [markdown] id="ri-2AQc5Rwx9"
# The procedure taken is:
# 1. Definition of the question
# 2. Reading and checking of the data
# 3. External data source validation
# 4. Cleaning of the dataset
# 5. Exploratory analysis
#
# + [markdown] id="ksm0_fccRxQd"
# ### DATA RELEVANCE
# + [markdown] id="9O7CJiITRx0F"
#
# Data to be used contains demographic information and what financial services are used by individuals in East Africa. The data is extracted from various Finsscope surveys and is ranging from 2016 to 2018. The data files include:
#
# Variable Definitions: http://bit.ly/VariableDefinitions
#
# Dataset: http://bit.ly/FinancialDataset
#
# FinAccess Kenya 2018: https://fsdkenya.org/publication/finaccess2019/
#
# Finscope Rwanda 2016: http://www.statistics.gov.rw/publication/finscope-rwanda-2016
#
# Finscope Tanzania 2017: http://www.fsdt.or.tz/finscope/
#
# Finscope Uganda 2018: http://fsduganda.or.ug/finscope-2018-survey-report/
#
# This data is relevant in this project since it provides important insights that will help in solving the research question.
# + [markdown] id="Ln1gbS1IUBLX"
# ## LOADING LIBRARIES
# + id="g-ry44zPXUQF" executionInfo={"status": "ok", "timestamp": 1631267040924, "user_tz": -180, "elapsed": 22, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
# importing libraries
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# + [markdown] id="LPRqZvg-T6YR"
# ## READING AND CHECKING DATA
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="s19c5Dj25Ilc" executionInfo={"status": "ok", "timestamp": 1631267040928, "user_tz": -180, "elapsed": 25, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="b4b93ef6-93f2-4ecd-c237-621d25941367"
# loading and viewing variable definitions dataset
url = "http://bit.ly/VariableDefinitions"
vb_df = pd.read_csv(url)
vb_df
# + colab={"base_uri": "https://localhost:8080/", "height": 656} id="N_gE8pNh5lQB" executionInfo={"status": "ok", "timestamp": 1631267041371, "user_tz": -180, "elapsed": 456, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="dbf1e1e0-2215-42c7-e23f-6a401c787277"
# loading and viewing financial dataset
url2 = "http://bit.ly/FinancialDataset"
fds = pd.read_csv(url2)
fds
# + colab={"base_uri": "https://localhost:8080/"} id="x3-US5OdBItZ" executionInfo={"status": "ok", "timestamp": 1631267041372, "user_tz": -180, "elapsed": 15, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="4c00e9fc-e70e-402e-84f8-9c8abc260345"
fds.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 330} id="FAzsrWIP5_Gl" executionInfo={"status": "ok", "timestamp": 1631267041374, "user_tz": -180, "elapsed": 16, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="51b605ea-ff32-4ee9-f1f9-35d41fa13f6a"
fds.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 313} id="-Y7PX5IM-V_S" executionInfo={"status": "ok", "timestamp": 1631267042114, "user_tz": -180, "elapsed": 753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="d9146ce7-d641-4836-e407-4aad061a93b6"
fds.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="LblxUCa8OLbq" executionInfo={"status": "ok", "timestamp": 1631267042116, "user_tz": -180, "elapsed": 57, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="954b15d7-d6e8-4853-d7cc-abe08eba7dba"
fds.dtypes
# + colab={"base_uri": "https://localhost:8080/"} id="o__5QbPFC9uJ" executionInfo={"status": "ok", "timestamp": 1631267042117, "user_tz": -180, "elapsed": 55, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="f70773f0-5042-4487-ded0-6730cd85ec03"
fds.columns
# + colab={"base_uri": "https://localhost:8080/"} id="48lYQ5ta-YXI" executionInfo={"status": "ok", "timestamp": 1631267042118, "user_tz": -180, "elapsed": 54, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="2ee43682-2909-414b-b4cd-ff45a82714f3"
fds.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 290} id="OZGVvqzwAd5V" executionInfo={"status": "ok", "timestamp": 1631267042119, "user_tz": -180, "elapsed": 53, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="43bf9401-1c80-4785-a91b-e8d479593cf5"
fds.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 202} id="sGfG3pR-IGEA" executionInfo={"status": "ok", "timestamp": 1631267042120, "user_tz": -180, "elapsed": 52, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="f9b145fb-4882-4dbb-b075-441325a98009"
fds.describe(include=object)
# + colab={"base_uri": "https://localhost:8080/"} id="WCz84IrXETzo" executionInfo={"status": "ok", "timestamp": 1631267042121, "user_tz": -180, "elapsed": 51, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="2ff8bc08-f070-44ee-c85a-6176cda8a9ae"
len(fds)
# + colab={"base_uri": "https://localhost:8080/"} id="7nKGQjbXpWQx" executionInfo={"status": "ok", "timestamp": 1631267042122, "user_tz": -180, "elapsed": 50, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="b5886309-2f22-42dc-8a54-a6e3063db32e"
fds.nunique()
# + colab={"base_uri": "https://localhost:8080/"} id="MTD2ATlaqCqR" executionInfo={"status": "ok", "timestamp": 1631267042124, "user_tz": -180, "elapsed": 50, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="6ed87ee9-a149-4828-f331-2844debfe60d"
fds.count()
# + [markdown] id="cDYzVN0iVKRP"
# ## EXTERNAL DATA SOURCE VALIDATION
# + [markdown] id="vHxekg75VPFt"
#
# FinAccess Kenya 2018: https://fsdkenya.org/publication/finaccess2019/
#
# Finscope Rwanda 2016: http://www.statistics.gov.rw/publication/finscope-rwanda-2016
#
# Finscope Tanzania 2017: http://www.fsdt.or.tz/finscope/
#
# Finscope Uganda 2018: http://fsduganda.or.ug/finscope-2018-survey-report/
# + [markdown] id="mpv0DchiXbDK"
# ## CLEANING THE DATASET
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="JOK4Ci275GOq" executionInfo={"status": "ok", "timestamp": 1631267042128, "user_tz": -180, "elapsed": 51, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="2c0e837f-fcc3-4eea-faa2-f2ebcc0821dd"
fds.head(2)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="79Zw3sa7XrI4" executionInfo={"status": "ok", "timestamp": 1631267042129, "user_tz": -180, "elapsed": 50, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="6283ef1a-b39e-4152-c6ac-c58d8aae4f7d"
# CHECKING FOR OUTLIERS IN YEAR COLUMN
sns.boxplot(x=fds['year'])
# + colab={"base_uri": "https://localhost:8080/"} id="RwJQ02xPwnYC" executionInfo={"status": "ok", "timestamp": 1631267042130, "user_tz": -180, "elapsed": 47, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="cf40e27a-8aa4-4021-fdab-1d0c3c92843a"
fds.shape
# + colab={"base_uri": "https://localhost:8080/"} id="vmg96X1xxPxR" executionInfo={"status": "ok", "timestamp": 1631267042131, "user_tz": -180, "elapsed": 46, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="4db03a0b-9561-4fef-9769-4832ff62d7be"
# dropping year column outliers
fds1= fds[fds['year']<2020]
fds1.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="eGNOSVV9v5sA" executionInfo={"status": "ok", "timestamp": 1631267042597, "user_tz": -180, "elapsed": 509, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="2c0fbd98-5902-4d71-f5a7-a6a3b772cd06"
# CHECKING FOR OUTLIERS IN HOUSEHOLD SIZE COLUMN
sns.boxplot(x=fds1['household_size'])
# + colab={"base_uri": "https://localhost:8080/"} id="ZQeJvfISFxGw" executionInfo={"status": "ok", "timestamp": 1631267042598, "user_tz": -180, "elapsed": 23, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="005d44db-78b1-4dcc-a340-cef65242aabf"
# dropping household size outliers
fds2 =fds1[fds1['household_size']<10]
fds2.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="YCB9rkZ4HVtN" executionInfo={"status": "ok", "timestamp": 1631267042599, "user_tz": -180, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="439de2d2-f785-4794-8d51-863f840874b2"
# CHECKING FOR OUTLIERS IN AGE OF RESPONDENT
sns.boxplot(x=fds2['Respondent Age'])
# + colab={"base_uri": "https://localhost:8080/"} id="l-tjbRDtHrNr" executionInfo={"status": "ok", "timestamp": 1631267042600, "user_tz": -180, "elapsed": 18, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="3ecce50c-dc0b-458e-eaa8-ff9a20e26407"
# dropping age of respondent outliers
fds3 = fds2[fds2['Respondent Age']<82]
fds3.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 611} id="kLXxqDI9IFMh" executionInfo={"status": "ok", "timestamp": 1631267043351, "user_tz": -180, "elapsed": 767, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="5727bd91-bbe7-43b1-df80-90f4a025b4b7"
# plotting the final boxplots
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2,2, figsize=(10, 7))
fig.suptitle('Boxplots')
sns.boxplot(fds3['Respondent Age'], ax=ax1)
sns.boxplot(fds3['year'], ax=ax2)
sns.boxplot(fds3['household_size'], ax=ax3)
plt.show()
# the outliers have finally been droppped
# + colab={"base_uri": "https://localhost:8080/"} id="lMTfjYpAdPCJ" executionInfo={"status": "ok", "timestamp": 1631267043352, "user_tz": -180, "elapsed": 54, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="e815c87a-7b9a-4558-bd23-dcd138f05032"
# CHECKING FOR NULLL OR MISSING DATA
fds3.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="9c9bqq1wSpaL" executionInfo={"status": "ok", "timestamp": 1631267043353, "user_tz": -180, "elapsed": 51, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="a02a88c2-d339-4b86-d47e-11ec4386dc5d"
# dropping nulls
fds4 = fds3.dropna()
fds4.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="AsLwrXk_OZYe" executionInfo={"status": "ok", "timestamp": 1631269130820, "user_tz": -180, "elapsed": 631, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="7354bf54-3e8c-4280-c131-da181514bb4c"
# dropping duplicates
fds4.drop_duplicates().head(2)
# + id="7WhITuWt8RiW" colab={"base_uri": "https://localhost:8080/", "height": 177} executionInfo={"status": "ok", "timestamp": 1631267043355, "user_tz": -180, "elapsed": 49, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="0ecedcd6-9e9e-472e-9847-1cf0a4aeb081"
# changing column names and columns to lowercase
#for columns in fds.columns:
#fds1[columns] = fds[columns].astype(str).str.lower()
#fds1
#fds1.rename(columns=str.lower)
# renaming columns
fds5 = fds4.rename(columns={'Type of Location':'location_type', 'Has a Bank account' : 'bank account','Cell Phone Access':'cellphone_access', 'Respondent Age': 'age_of_respondent',
'The relathip with head': 'relationship_with_head', 'Level of Educuation' : 'education_level', 'Type of Job': 'job_type'})
fds5.head(2)
# + colab={"base_uri": "https://localhost:8080/"} id="DuZ-syPhcMlz" executionInfo={"status": "ok", "timestamp": 1631267043356, "user_tz": -180, "elapsed": 48, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="d7d01d0c-3848-4e23-ccfc-e47cbec20361"
fds5.shape
# + colab={"base_uri": "https://localhost:8080/"} id="wcqV5eX7cQ4R" executionInfo={"status": "ok", "timestamp": 1631267043357, "user_tz": -180, "elapsed": 45, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="49be20bb-958e-4f2c-cdec-b38d4ac130b0"
fds5.size
# + colab={"base_uri": "https://localhost:8080/"} id="yPyNTRs8a3Mi" executionInfo={"status": "ok", "timestamp": 1631267043357, "user_tz": -180, "elapsed": 43, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="ba804a66-f79b-4824-94f6-1adff4429609"
fds5.nunique()
# + colab={"base_uri": "https://localhost:8080/"} id="brnAu1Qod-4H" executionInfo={"status": "ok", "timestamp": 1631267043358, "user_tz": -180, "elapsed": 42, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="32240dca-e7ec-4472-eae6-2cce38189927"
fds5['country'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="-Sqzrj3Rv5ob" executionInfo={"status": "ok", "timestamp": 1631267043359, "user_tz": -180, "elapsed": 40, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="93dc9d3d-0dee-4c7a-f55b-f24a56ca602c"
fds5['year'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="UaMvQ6PTwoKm" executionInfo={"status": "ok", "timestamp": 1631267043360, "user_tz": -180, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="c24e87aa-8666-43eb-bdfd-0e2750772cba"
fds5['bank account'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="jNHcgixFxAfG" executionInfo={"status": "ok", "timestamp": 1631267043361, "user_tz": -180, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="a8715d78-1ab5-42e4-9ab1-d88ed4b3a395"
fds5['location_type'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="YnkohQOOxS9D" executionInfo={"status": "ok", "timestamp": 1631267043368, "user_tz": -180, "elapsed": 45, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="4a9e02d5-8c3e-4e59-bc51-c7db19639a73"
fds5['cellphone_access'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="oDZwzaM_xa5r" executionInfo={"status": "ok", "timestamp": 1631267043369, "user_tz": -180, "elapsed": 44, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="4caf1904-4352-405a-a105-cdf3ded6c491"
fds5['education_level'].unique()
# + id="tN_TTwSAxlYk" executionInfo={"status": "ok", "timestamp": 1631267043370, "user_tz": -180, "elapsed": 43, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
fds5.drop(fds5.loc[fds5['education_level'] ==6].index, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="4id2SVAR3zlh" executionInfo={"status": "ok", "timestamp": 1631267043371, "user_tz": -180, "elapsed": 43, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="b4a66a55-81ef-4cba-cce8-1aad94fa3f4b"
fds5['education_level'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="aSV1aqMK2gPL" executionInfo={"status": "ok", "timestamp": 1631267043372, "user_tz": -180, "elapsed": 41, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="d6f32001-3a3f-4de5-9523-ee1b19265d4e"
fds5['gender_of_respondent'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="Zm8fphme3F-G" executionInfo={"status": "ok", "timestamp": 1631267043372, "user_tz": -180, "elapsed": 40, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="0a387804-cbd1-4524-dc98-865d8d3a587c"
fds5['household_size'].unique()
# + id="QqmU8wil3NZP" executionInfo={"status": "ok", "timestamp": 1631267043373, "user_tz": -180, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
fds5.drop(fds5.loc[fds5['household_size'] == 0].index, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="9A55HITo3rIF" executionInfo={"status": "ok", "timestamp": 1631267043374, "user_tz": -180, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="358dccc2-b269-4001-dd96-fc283f8cf4cf"
fds5['household_size'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="ryPMBZWt4iOC" executionInfo={"status": "ok", "timestamp": 1631267043374, "user_tz": -180, "elapsed": 38, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="6d995367-cc39-4c2c-d714-6af6dedaaec5"
fds5['job_type'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="Cr5fGUew4o36" executionInfo={"status": "ok", "timestamp": 1631267043375, "user_tz": -180, "elapsed": 37, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="36788eff-3e51-49a7-dad1-824fcd431d94"
fds5['relationship_with_head'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="qLkeu1j84vAV" executionInfo={"status": "ok", "timestamp": 1631267044023, "user_tz": -180, "elapsed": 683, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="4d92cf7c-7cb5-465e-d21b-f79daa7190ad"
fds5['age_of_respondent'].unique()
# + [markdown] id="PHGb1Y5GSz04"
# ## **EXPLORATORY ANALYSIS**
# + [markdown] id="PmmqqH16lmoT"
# ### 1.UNIVARIATE ANALYSIS
# + [markdown] id="kF-0sWdUm3ay"
# #### a. NUMERICAL VARIABLES
# + [markdown] id="OtPcPlxU8ghu"
# ##### MODE
#
# + id="T_39juuiC9C3" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1631267044023, "user_tz": -180, "elapsed": 30, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="5cb8e41d-385a-4b25-8f31-231bef4b8bb0"
fds5['year'].mode()
# + colab={"base_uri": "https://localhost:8080/"} id="e5NfmGVc5bAk" executionInfo={"status": "ok", "timestamp": 1631267044024, "user_tz": -180, "elapsed": 26, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="3cf54377-ae4d-4073-fae8-977b861ba81d"
fds5['household_size'].mode()
# + colab={"base_uri": "https://localhost:8080/"} id="opm9e7jJ6B2o" executionInfo={"status": "ok", "timestamp": 1631267044024, "user_tz": -180, "elapsed": 22, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="9550a611-809f-45a0-b103-316c39218a9f"
fds5['age_of_respondent'].mode()
# + [markdown] id="56jHFfRN_u_0"
# ##### MEAN
# + colab={"base_uri": "https://localhost:8080/"} id="pDhraQie_T_w" executionInfo={"status": "ok", "timestamp": 1631267044025, "user_tz": -180, "elapsed": 19, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="313ab974-9209-4dd5-a015-2979ab23cee5"
fds5['age_of_respondent'].mean()
# + colab={"base_uri": "https://localhost:8080/"} id="Jai227Y4_flv" executionInfo={"status": "ok", "timestamp": 1631267044105, "user_tz": -180, "elapsed": 94, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="af409a6b-379e-429c-8ab3-190fea47ebd1"
fds5['household_size'].mean()
# + colab={"base_uri": "https://localhost:8080/"} id="vPovgtuxCLaN" executionInfo={"status": "ok", "timestamp": 1631267045924, "user_tz": -180, "elapsed": 1906, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="e51be046-9066-4dca-a555-17d98f427d89"
fds5.mean()
# + [markdown] id="tZfc8v1O_4U7"
# ##### MEDIAN
# + colab={"base_uri": "https://localhost:8080/"} id="lA1ySTGA_8pG" executionInfo={"status": "ok", "timestamp": 1631267045929, "user_tz": -180, "elapsed": 85, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="8da3f52a-2507-4742-9546-e8b4c2a7676a"
fds5['age_of_respondent'].median()
# + colab={"base_uri": "https://localhost:8080/"} id="9h5rbRskADlD" executionInfo={"status": "ok", "timestamp": 1631267045930, "user_tz": -180, "elapsed": 78, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="b6f4a031-76ed-4e61-86cb-70bde062fded"
fds5['household_size'].median()
# + colab={"base_uri": "https://localhost:8080/"} id="MmBZGL9RCSf_" executionInfo={"status": "ok", "timestamp": 1631267045931, "user_tz": -180, "elapsed": 75, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="2d76f0eb-de04-4996-fc81-987240e7d8f2"
fds5.median()
# + [markdown] id="9TafiDIWBOTI"
# ##### RANGE
# + colab={"base_uri": "https://localhost:8080/"} id="x4DYmR9-AIIy" executionInfo={"status": "ok", "timestamp": 1631267045933, "user_tz": -180, "elapsed": 71, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="4b550dd8-94bf-459a-914c-43662b9ec1b4"
a = fds5['age_of_respondent'].max()
b = fds5['age_of_respondent'].min()
c = a-b
print('The range of the age for the respondents is', c)
# + colab={"base_uri": "https://localhost:8080/"} id="v3ZGZZyhDNaI" executionInfo={"status": "ok", "timestamp": 1631267045933, "user_tz": -180, "elapsed": 67, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="b9134fe8-a3ea-48af-e18d-2bbfc35493ba"
d = fds5['household_size'].max()
e = fds5['household_size'].min()
f = d-e
print('The range of the household_sizes is', f)
# + [markdown] id="3LvtGLAhEqFc"
# ##### QUANTILE AND INTERQUANTILE
# + colab={"base_uri": "https://localhost:8080/", "height": 138} id="OOxCpxQUDxAp" executionInfo={"status": "ok", "timestamp": 1631267045935, "user_tz": -180, "elapsed": 64, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="2ee6ede0-267c-4d65-b53f-811822ae8b6a"
fds5.quantile([0.25,0.5,0.75])
# + colab={"base_uri": "https://localhost:8080/"} id="auJ_xbDgFWOx" executionInfo={"status": "ok", "timestamp": 1631267045936, "user_tz": -180, "elapsed": 63, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="9964e9f7-9286-4273-d3f8-364d8eb6b7be"
# FINDING THE INTERQUANTILE RANGE = IQR
Q3 = fds5['age_of_respondent'].quantile(0.75)
Q2 = fds5['age_of_respondent'].quantile(0.25)
IQR= Q3-Q2
print('The IQR for the respondents age is', IQR)
# + colab={"base_uri": "https://localhost:8080/"} id="wPQuhTFRJ7XO" executionInfo={"status": "ok", "timestamp": 1631267045939, "user_tz": -180, "elapsed": 62, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="fd32e3b5-68ae-4167-c59f-c8507fb981b8"
q3 = fds5['household_size'].quantile(0.75)
q2 = fds5['household_size'].quantile(0.25)
iqr = q3-q2
print('The IQR for household sizes is', iqr)
# + id="xliV4OsPKxCG" executionInfo={"status": "ok", "timestamp": 1631267045944, "user_tz": -180, "elapsed": 61, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
# + [markdown] id="_WTAxkG7Ld3V"
# ##### STANDARD DEVIATION
# + colab={"base_uri": "https://localhost:8080/"} id="v0MG6cGeLkx4" executionInfo={"status": "ok", "timestamp": 1631267045945, "user_tz": -180, "elapsed": 61, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="26cf5ce9-caba-490e-e2ed-93c9794d3119"
fds5.std()
# + [markdown] id="C8bw6LjdL2Cn"
# ##### VARIANCE
# + colab={"base_uri": "https://localhost:8080/"} id="ICsG_FnxLoy9" executionInfo={"status": "ok", "timestamp": 1631267045946, "user_tz": -180, "elapsed": 59, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="dbde30df-e13a-43a8-ec4d-8470eea44662"
fds5.var()
# + [markdown] id="cAjETCHLMAuZ"
# ##### KURTOSIS
# + colab={"base_uri": "https://localhost:8080/"} id="XIEnkrzwL797" executionInfo={"status": "ok", "timestamp": 1631267045947, "user_tz": -180, "elapsed": 56, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="1969596d-4249-43d7-9c8c-40049643f828"
fds5.kurt()
# + [markdown] id="T3QcI3NNMMrD"
# ##### SKEWNESS
# + colab={"base_uri": "https://localhost:8080/"} id="fWpy-SFWMKST" executionInfo={"status": "ok", "timestamp": 1631267045948, "user_tz": -180, "elapsed": 52, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="c6dbc738-b0bb-4f69-b961-9073e0595581"
fds5.skew()
# + id="pNHsUSqBMT7u" executionInfo={"status": "ok", "timestamp": 1631267045951, "user_tz": -180, "elapsed": 48, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
# + [markdown] id="h0c8tGbiMm-N"
# #### b. CATEGORICAL
# + [markdown] id="WT1bgKHiM2-f"
# ##### MODE
#
# + colab={"base_uri": "https://localhost:8080/", "height": 131} id="kLEOUeJvMXbA" executionInfo={"status": "ok", "timestamp": 1631267045952, "user_tz": -180, "elapsed": 49, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="a02ad260-b473-41e4-a16e-5fdedbb6855d"
fds5.mode().head(1)
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="9HFuDRhFMYxF" executionInfo={"status": "ok", "timestamp": 1631267046774, "user_tz": -180, "elapsed": 870, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="cae94e6c-ebe1-4700-ef42-9b85948c7160"
fds5['age_of_respondent'].plot(kind="hist")
plt.xlabel('ages of respondents')
plt.ylabel('frequency')
plt.title(' Frequency of the ages of the respondents')
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="rStHF7fMaUJE" executionInfo={"status": "ok", "timestamp": 1631267046776, "user_tz": -180, "elapsed": 48, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="827cfcc7-1e9f-4b75-ac5d-dfa6d4109c0c"
country=fds5['country'].value_counts()
print(country)
# Plotting the pie chart
colors=['pink','white','cyan','yellow']
country.plot(kind='pie',colors=colors,autopct='%1.1f%%',shadow=True,startangle=90)
plt.title('Distribution of the respondents by country')
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="_-KlFxtPb-HF" executionInfo={"status": "ok", "timestamp": 1631267046777, "user_tz": -180, "elapsed": 36, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="8f30450c-3fc8-4361-d628-a0d765f0d4c2"
bank =fds5['bank account'].value_counts()
# Plotting the pie chart
colors=['plum', 'aqua']
bank.plot(kind='pie',colors=colors,autopct='%1.1f%%',shadow=True,startangle=90)
plt.title('availability of bank accounts')
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="BQTansfEdYZE" executionInfo={"status": "ok", "timestamp": 1631267048101, "user_tz": -180, "elapsed": 1348, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="de8bf9d5-15d4-4e8e-d741-0e8659b1a763"
location=fds5['location_type'].value_counts()
# Plotting the pie chart
colors=['aquamarine','beige']
location.plot(kind='pie',colors=colors,autopct='%1.3f%%',shadow=True,startangle=00)
plt.title('Distribution of the respondents according to location')
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="38VGxmSaeaYl" executionInfo={"status": "ok", "timestamp": 1631267048102, "user_tz": -180, "elapsed": 95, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="e390df4d-1ba5-4c71-f698-8e3da8856266"
celly =fds5['cellphone_access'].value_counts()
# Plotting the pie chart
colors=['plum','lavender']
celly.plot(kind='pie',colors=colors,autopct='%1.1f%%',shadow=True,startangle=0)
plt.title('cellphone access for the respondents')
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="u6WAIBFGh-3J" executionInfo={"status": "ok", "timestamp": 1631267048102, "user_tz": -180, "elapsed": 88, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="b5382ea8-cc17-4a7c-c137-73ac534971d6"
gen =fds5['gender_of_respondent'].value_counts()
# Plotting the pie chart
colors=['red','lavender']
gen.plot(kind='pie',colors=colors,autopct='%1.1f%%',shadow=True,startangle=0)
plt.title('gender distribution')
# + [markdown] id="ySG4FrdgH5Ut"
# ##### CONCLUSION AND RECOMMENDATION
# + [markdown] id="xk6nvmbOIccM"
# Most of the data was collected in Rwanda.
# Most of the data was collected in Rural areas.
# Most of those who were interviewed were women.
# Most of the population has mobile phones.
# There were several outliers.
#
# Since 75% of the population has phones, phones should be used as the main channel for information and awareness of bank accessories.
#
# + [markdown] id="c_2cuP-gkSjN"
# ### 2. BIVARIATE ANALYSIS
# + colab={"base_uri": "https://localhost:8080/", "height": 333} id="nr8wZpmrkaLO" executionInfo={"status": "ok", "timestamp": 1631267048103, "user_tz": -180, "elapsed": 85, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="821809e3-19fe-40d9-c95d-0423c859962a"
fds5.head()
# + id="LoPLzxKzkbiF" executionInfo={"status": "ok", "timestamp": 1631267048103, "user_tz": -180, "elapsed": 83, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
#@title Since i am predicting the likelihood of the respondents using the bank,I shall be comparing all variables against the bank account column.
# + [markdown] id="lsMIevQgqS_j"
# ##### NUMERICAL VS NUMERICAL
# + colab={"base_uri": "https://localhost:8080/", "height": 550} id="YhC3TPRjkbfR" executionInfo={"status": "ok", "timestamp": 1631267051728, "user_tz": -180, "elapsed": 3708, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="3d61efd3-1686-4207-b6d9-b488d7931acb"
sns.pairplot(fds5)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 270} id="be_MGZV8ka_1" executionInfo={"status": "ok", "timestamp": 1631268302022, "user_tz": -180, "elapsed": 1031, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="8207b7c4-2780-43dd-d8c3-a25211258497"
# pearson correlation of numerical variables
sns.heatmap(fds5.corr(),annot=True)
plt.show()
# possible weak correlation
# + colab={"base_uri": "https://localhost:8080/", "height": 138} id="cdVUJ5Dt5dsT" executionInfo={"status": "ok", "timestamp": 1631267051737, "user_tz": -180, "elapsed": 153, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="908dcfb8-2666-4fab-f7cb-f542bb920bda"
fds5.corr()
# + [markdown] id="nmlKOhut5GRZ"
# ##### CATEGORICAL VS CATEGORICAL
# + colab={"base_uri": "https://localhost:8080/", "height": 426} id="JjzEu7y0ka2L" executionInfo={"status": "ok", "timestamp": 1631267051740, "user_tz": -180, "elapsed": 155, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="ce3ff902-a936-458b-f2d3-7723cedcc0cd"
# Grouping bank usage by country
country1 = fds5.groupby('country')['bank account'].value_counts(normalize=True).unstack()
colors= ['lightpink', 'skyblue']
country1.plot(kind='bar', figsize=(8, 6), color=colors, stacked=True)
plt.title('Bank usage by country', fontsize=15, y=1.015)
plt.xlabel('country', fontsize=14, labelpad=15)
plt.xticks(rotation = 360)
plt.ylabel('Bank usage by country', fontsize=14, labelpad=15)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 433} id="cnGmBoZekaps" executionInfo={"status": "ok", "timestamp": 1631267051742, "user_tz": -180, "elapsed": 156, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="ac9b9c6d-0c03-4663-f3cd-c9150d8d03db"
# Bank usage by gender
gender1 = fds5.groupby('gender_of_respondent')['bank account'].value_counts(normalize=True).unstack()
colors= ['lightpink', 'skyblue']
gender1.plot(kind='bar', figsize=(8, 6), color=colors, stacked=True)
plt.title('Bank usage by gender', fontsize=17, y=1.015)
plt.xlabel('gender', fontsize=17, labelpad=17)
plt.xticks(rotation = 360)
plt.ylabel('Bank usage by gender', fontsize=17, labelpad=17)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 433} id="tzRNKPrp5OaB" executionInfo={"status": "ok", "timestamp": 1631267052548, "user_tz": -180, "elapsed": 960, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="415cafb7-70bd-4f22-d6da-bb6a85ff8a8f"
# Bank usage depending on level of education
ed2 = fds5.groupby('education_level')['bank account'].value_counts(normalize=True).unstack()
colors= ['cyan', 'darkcyan']
ed2.plot(kind='barh', figsize=(8, 6), color=colors, stacked=True)
plt.title('Bank usage by level of education', fontsize=17, y=1.015)
plt.xlabel('frequency', fontsize=17, labelpad=17)
plt.xticks(rotation = 360)
plt.ylabel('level of education', fontsize=17, labelpad=17)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="3ALM8pBnBjlE" executionInfo={"status": "ok", "timestamp": 1631267052552, "user_tz": -180, "elapsed": 63, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="b1636411-6b80-43f5-a68b-7c18f08bca91"
ms = fds5.groupby('marital_status')['bank account'].value_counts(normalize=True).unstack()
colors= ['coral', 'orange']
ms.plot(kind='barh', figsize=(8, 6), color=colors, stacked=True)
plt.title('Bank usage by marital status', fontsize=17, y=1.015)
plt.xlabel('frequency', fontsize=17, labelpad=17)
plt.xticks(rotation = 360)
plt.ylabel('marital status', fontsize=17, labelpad=17)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="HfGxVMY9uvZM" executionInfo={"status": "ok", "timestamp": 1631267053551, "user_tz": -180, "elapsed": 1032, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="bda5f006-0eaf-4099-bcb5-f4d06a99632b"
gj = fds5.groupby('gender_of_respondent')['job_type'].value_counts(normalize=True).unstack()
#colors= ['coral', 'orange']
gj.plot(kind='bar', figsize=(8, 6), stacked=True)
plt.title('job type by gender', fontsize=17, y=1.015)
plt.xlabel('gender_of_respondent', fontsize=17, labelpad=17)
plt.xticks(rotation = 360)
plt.ylabel('job type', fontsize=17, labelpad=17)
# + [markdown] id="3Kk6-V-MtOqG"
# ##### NUMERICAL VS CATEGORICAL
# + [markdown] id="Espg6Z80t6rF"
# ##### IMPLEMENTING AND CHALLENGING SOLUTION
# + id="nxK1gH19D63M" executionInfo={"status": "ok", "timestamp": 1631267053561, "user_tz": -180, "elapsed": 197, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
# + [markdown] id="UTawlVDSJetr"
# Most of those interviewed do not have bank accounts of which 80% is the uneducated.
# Most of the population that participated is married,followed by single/never married.
# Most of the population has primary school education level.
# Most of the population is involved in farming followed by self employment.
# Bank usage has more males than females.
#
#
# More channeling needs to be done in Kenya as it has the least bank users.
# + [markdown] id="867BDuRtyGPW"
# ###3. MULTIVARIATE ANALYSIS
# + id="FvgMmrOqyOLG" executionInfo={"status": "ok", "timestamp": 1631267053569, "user_tz": -180, "elapsed": 204, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
# Multivariate analysis - This is a statistical analysis that involves observation and analysis of more than one statistical outcome variable at a time
# + id="4SqFKBCHyN9C" executionInfo={"status": "ok", "timestamp": 1631267053576, "user_tz": -180, "elapsed": 209, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
# LETS MAKE A COPY
fds_new = fds5.copy()
# + colab={"base_uri": "https://localhost:8080/"} id="qQ2FK3_KyNu4" executionInfo={"status": "ok", "timestamp": 1631267053579, "user_tz": -180, "elapsed": 211, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="8a77f2e7-65fc-4b10-c404-554da9ba4473"
fds_new.columns
# + colab={"base_uri": "https://localhost:8080/"} id="TgM5AtwS7SlN" executionInfo={"status": "ok", "timestamp": 1631267053588, "user_tz": -180, "elapsed": 196, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="8c16eab4-5405-4f62-80e5-d5d6b196dd93"
fds_new.dtypes
# + id="e-1vZGM97g3L" executionInfo={"status": "ok", "timestamp": 1631267053699, "user_tz": -180, "elapsed": 294, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
# IMPORTING THE LABEL ENCODER
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
# encoding categorial values
fds_new['country']=le.fit_transform(fds_new['country'].astype(str))
fds_new['location_type']=le.fit_transform(fds_new['location_type'].astype(str))
fds_new['cellphone_access']=le.fit_transform(fds_new['cellphone_access'].astype(str))
fds_new['gender_of_respondent']=le.fit_transform(fds_new['gender_of_respondent'].astype(str))
fds_new['relationship_with_head']=le.fit_transform(fds_new['relationship_with_head'].astype(str))
fds_new['marital_status']=le.fit_transform(fds_new['marital_status'].astype(str))
fds_new['education_level']=le.fit_transform(fds_new['education_level'].astype(str))
fds_new['job_type']=le.fit_transform(fds_new['job_type'].astype(str))
# + colab={"base_uri": "https://localhost:8080/", "height": 236} id="mFLsS-6i99Nv" executionInfo={"status": "ok", "timestamp": 1631267053700, "user_tz": -180, "elapsed": 294, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="3647b21f-f73a-4363-d677-39f76e811165"
fds_new.sample(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 108} id="cx20LnKgB7C-" executionInfo={"status": "ok", "timestamp": 1631267053701, "user_tz": -180, "elapsed": 293, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="dd010b02-f5d3-4539-a5b3-278415676ddb"
# dropping unnecessary columns
fds_new.drop(['age_of_respondent','uniqueid','year'], axis=1).head(2)
# + [markdown] id="gXRH7FT4FkBf"
# ##### FACTOR ANALYSIS
# + colab={"base_uri": "https://localhost:8080/"} id="QmrgjffF-g30" executionInfo={"status": "ok", "timestamp": 1631267055850, "user_tz": -180, "elapsed": 2439, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="75ee2a58-78f7-4e38-fcbe-3608ddf8e887"
# Installing factor analyzer
# !pip install factor_analyzer==0.2.3
from factor_analyzer.factor_analyzer import calculate_bartlett_sphericity
chi_square_value,p_value=calculate_bartlett_sphericity(fds_new)
chi_square_value, p_value
# In Bartlett ’s test, the p-value is 0. The test was statistically significant,
# indicating that the observed correlation matrix is not an identity matrix.
# + id="FZlChFJ0BTFG" executionInfo={"status": "ok", "timestamp": 1631267055851, "user_tz": -180, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}}
# Value of KMO less than 0.6 is considered inadequate.
#
#from factor_analyzer.factor_analyzer import calculate_kmo
#kmo_all,kmo_model=calculate_kmo(fds_new)
#calculate_kmo(fds_new)
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="tDB8-MZ7EWqW" executionInfo={"status": "ok", "timestamp": 1631267057598, "user_tz": -180, "elapsed": 1753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="f8cd9dc6-a5fb-4bb7-f610-62b9816a607c"
# Choosing the Number of Factors
from factor_analyzer.factor_analyzer import FactorAnalyzer
# Creating factor analysis object and perform factor analysis
fa = FactorAnalyzer()
fa.analyze(fds_new, 10, rotation=None)
# Checking the Eigenvalues
ev, v = fa.get_eigenvalues()
ev
# We choose the factors that are > 1.
# so we choose 4 factors only
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="HBHWpyNaENd2" executionInfo={"status": "ok", "timestamp": 1631267251777, "user_tz": -180, "elapsed": 2685, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="87e399c3-0925-4a14-8f93-7d49ab25b639"
# PERFOMING FACTOR ANALYSIS FOR 4 FACTORS
fa = FactorAnalyzer()
fa.analyze(fds_new, 4, rotation="varimax")
fa.loadings
# + colab={"base_uri": "https://localhost:8080/", "height": 138} id="YL8lrzVtBzxy" executionInfo={"status": "ok", "timestamp": 1631267060226, "user_tz": -180, "elapsed": 151, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgZPXtM0j7W1i2xcmvtNcYKC8542e6-0SEw5qTn=s64", "userId": "02466132087222530859"}} outputId="91417beb-6ceb-4a29-d051-450986115778"
# GETTING VARIANCE FOR THE FACTORS
fa.get_factor_variance()
# + [markdown] id="zV4OazzOHEug"
# ##### CONCLUSION
#
# + [markdown] id="7AHTmgrYL9at"
# The reduction method used was factor analysis.
# Four factors had an eigen value greater than 1.
#
# + [markdown] id="DSuM4ftdMnkh"
# ##### CHALLENGING SOLUTION
# + [markdown] id="LfB1u26DMuY0"
# There is room for modification because there are other methodologies that could be used for analysis.
# There is also an assumption that this data is accurate to the real world.
| Moringa_Data_Science_Core_W2_Independent_Project_2021_09_Moreen_Mugambi_Python_Notebook_ipyn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
N, K=map(int, input().split())
x_max=100000
arrived=[False for i in range(x_max+1)]
arrived[N]=True
time=[0 for i in range(x_max+1)]
start_mark=0
end_mark=0
location=[0 for i in range(x_max+1)]
location[0]=N
STOP=False
while end_mark<=x_max:
start=location[start_mark]
start_mark+=1
for i in range(3):
if i==0: next_loc=start+1
if i==1: next_loc=start-1
if i==2: next_loc=start*2
if next_loc>=0 and next_loc<=x_max and arrived[next_loc]==False:
end_mark+=1
location[end_mark]=next_loc
arrived[next_loc]=True
time[next_loc]=time[start]+1
if next_loc==K:
print(time[next_loc])
STOP=True
if STOP:
break
| OpenJudge/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import Counter
from collections import defaultdict
import functools
from IPython.display import set_matplotlib_formats
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import skimage
from skimage import io
# %matplotlib inline
set_matplotlib_formats('svg')
plt.rcParams["patch.force_edgecolor"] = True
# -
df = pd.read_csv("../data/interim/texts.csv")
df.head()
print("Median amount of symbols in text: %.0f" % df.text.str.len().median())
print("Mean amount of symbols in text: %.2f" % df.text.str.len().mean())
print("Min amount of symbols in text: %.0f" % df.text.str.len().min())
print("Max amount of symbols in text: %.0f" % df.text.str.len().max())
n_bins = int(1 + 3.222 * np.log(len(df)))
sns.distplot(df.text.str.len(), kde=False, bins=n_bins)
plt.xlabel("Amount of characters in text")
plt.title("Texts' char-level length distribution");
texts = df.text.to_list()
# Посмотрим количество уникальных символов.
print("N unique symbols: %.0f" % len(functools.reduce(lambda x, y: set(x) | set(y), texts, set())))
# Посмотрим на частотность употребления символов.
char_counts = Counter()
for text in texts:
char_counts += Counter(text)
# Посмотрим на примеры, где употребляются самые редкие символы.
# +
threshold = 3
symbols = dict(filter(lambda x: x[1] < 10, char_counts.items())).keys()
def if_contains(string, symbols=None):
if symbols is None:
symbols = []
return any(c in symbols for c in string)
df[df.text.apply(lambda x: if_contains(x, symbols=symbols))]
# -
img = io.imread("../data/train/images/202_18_19.jpg")
io.imshow(img)
io.show()
# Выведем топ-10 самых частотных символов.
symbol, frequency = list(zip(*sorted(char_counts.items(), key=lambda x: x[1], reverse=True)[:15]))
dataframe = pd.DataFrame({"symbol": symbol, "frequency": frequency})
sns.barplot(y="symbol", x="frequency", data=dataframe, orient="h")
plt.title("15 the most frequent symbols");
# И 10 самых редких.
symbol, frequency = list(zip(*sorted(char_counts.items(), key=lambda x: x[1])[:15]))
dataframe = pd.DataFrame({"symbol": symbol, "frequency": frequency})
sns.barplot(y="symbol", x="frequency", data=dataframe, orient="h")
plt.title("5 the rarest symbols");
# ## Словарные статистики
lengths = df.text.str.split().apply(len)
print("Median amount of tokens in text: %.0f" % lengths.median())
print("Mean amount of tokens in text: %.2f" % lengths.mean())
print("Min amount of tokens in text: %.0f" % lengths.min())
print("Max amount of tokens in text: %.0f" % lengths.max())
n_bins = int(1 + 3.222 * np.log(len(df)))
sns.distplot(lengths, kde=False, bins=n_bins)
plt.xlabel("Amount of tokens in text")
plt.title("Texts' token-level length distribution");
word_counts = Counter()
for text in texts:
word_counts += Counter(text.split())
print("Number of unique tokens: %.0f" % len(word_counts))
word, frequency = list(zip(*sorted(word_counts.items(), key=lambda x: x[1], reverse=True)[:15]))
dataframe = pd.DataFrame({"word": word, "frequency": frequency})
sns.barplot(y="word", x="frequency", data=dataframe, orient="h")
plt.title("15 the most frequent tokens");
word, frequency = list(zip(*sorted(word_counts.items(), key=lambda x: x[1])[:15]))
dataframe = pd.DataFrame({"word": word, "frequency": frequency})
sns.barplot(y="word", x="frequency", data=dataframe, orient="h")
plt.title("15 the rarest tokens");
| notebooks/eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (pytorch_src2)
# language: python
# name: pytorch_src2
# ---
# +
import torch
import sys
import os
os.chdir("../../")
# sys.path.append("../../scripts/")
# sys.path.append("../../protein/")
# -
from scripts import proteinnet2pytorch as p2p
import prody as pr
from structure_utils import angle_list_to_sin_cos, seq_to_onehot
p, header = pr.parsePDB("research/notebooks/2ldj.pdb", chain="A", header=True)
seq = header["A"].sequence
dihedrals, coords, sequence = p2p.get_seq_and_masked_coords_and_angles(p, seq)
dihedrals = angle_list_to_sin_cos([dihedrals])[0]
sequence = seq_to_onehot(sequence)
import numpy as np
np.isnan(dihedrals).all(axis=-1).any()
dataset = {"train": {"seq": [sequence],
"ang": [dihedrals],
"crd": [coords],
"ids": ["2ldj"]},
"valid": {"seq": [sequence],
"ang": [dihedrals],
"crd": [coords],
"ids": ["2ldj"]},
"test": {"seq": [sequence],
"ang": [dihedrals],
"crd": [coords],
"ids": ["2ldj"]},
"settings": {"max_len": len(sequence)}
}
torch.save(dataset, "trp_cage.pt")
| research/notebooks/190929 OpenMM Loss Starter Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Attention Scoring Functions
# :label:`sec_attention-scoring-functions`
#
# In :numref:`sec_nadaraya-watson`,
# we used a Gaussian kernel to model
# interactions between queries and keys.
# Treating the exponent of the Gaussian kernel
# in :eqref:`eq_nadaraya-watson-gaussian`
# as an *attention scoring function* (or *scoring function* for short),
# the results of this function were
# essentially fed into
# a softmax operation.
# As a result,
# we obtained
# a probability distribution (attention weights)
# over values that are paired with keys.
# In the end,
# the output of the attention pooling
# is simply a weighted sum of the values
# based on these attention weights.
#
# At a high level,
# we can use the above algorithm
# to instantiate the framework of attention mechanisms
# in :numref:`fig_qkv`.
# Denoting an attention scoring function by $a$,
# :numref:`fig_attention_output`
# illustrates how the output of attention pooling
# can be computed as a weighted sum of values.
# Since attention weights are
# a probability distribution,
# the weighted sum is essentially
# a weighted average.
#
# 
# :label:`fig_attention_output`
#
#
#
# Mathematically,
# suppose that we have
# a query $\mathbf{q} \in \mathbb{R}^q$
# and $m$ key-value pairs $(\mathbf{k}_1, \mathbf{v}_1), \ldots, (\mathbf{k}_m, \mathbf{v}_m)$, where any $\mathbf{k}_i \in \mathbb{R}^k$ and any $\mathbf{v}_i \in \mathbb{R}^v$.
# The attention pooling $f$
# is instantiated as a weighted sum of the values:
#
# $$f(\mathbf{q}, (\mathbf{k}_1, \mathbf{v}_1), \ldots, (\mathbf{k}_m, \mathbf{v}_m)) = \sum_{i=1}^m \alpha(\mathbf{q}, \mathbf{k}_i) \mathbf{v}_i \in \mathbb{R}^v,$$
# :eqlabel:`eq_attn-pooling`
#
# where
# the attention weight (scalar) for the query $\mathbf{q}$
# and key $\mathbf{k}_i$
# is computed by
# the softmax operation of
# an attention scoring function $a$ that maps two vectors to a scalar:
#
# $$\alpha(\mathbf{q}, \mathbf{k}_i) = \mathrm{softmax}(a(\mathbf{q}, \mathbf{k}_i)) = \frac{\exp(a(\mathbf{q}, \mathbf{k}_i))}{\sum_{j=1}^m \exp(a(\mathbf{q}, \mathbf{k}_j))} \in \mathbb{R}.$$
# :eqlabel:`eq_attn-scoring-alpha`
#
# As we can see,
# different choices of the attention scoring function $a$
# lead to different behaviors of attention pooling.
# In this section,
# we introduce two popular scoring functions
# that we will use to develop more
# sophisticated attention mechanisms later.
#
# + origin_pos=1 tab=["mxnet"]
import math
from mxnet import np, npx
from mxnet.gluon import nn
from d2l import mxnet as d2l
npx.set_np()
# + [markdown] origin_pos=3
# ## Masked Softmax Operation
#
# As we just mentioned,
# a softmax operation is used to
# output a probability distribution as attention weights.
# In some cases,
# not all the values should be fed into attention pooling.
# For instance,
# for efficient minibatch processing in :numref:`sec_machine_translation`,
# some text sequences are padded with
# special tokens that do not carry meaning.
# To get an attention pooling
# over
# only meaningful tokens as values,
# we can specify a valid sequence length (in number of tokens)
# to filter out those beyond this specified range
# when computing softmax.
# In this way,
# we can implement such a *masked softmax operation*
# in the following `masked_softmax` function,
# where any value beyond the valid length
# is masked as zero.
#
# + origin_pos=4 tab=["mxnet"]
#@save
def masked_softmax(X, valid_lens):
"""Perform softmax operation by masking elements on the last axis."""
# `X`: 3D tensor, `valid_lens`: 1D or 2D tensor
if valid_lens is None:
return npx.softmax(X)
else:
shape = X.shape
if valid_lens.ndim == 1:
valid_lens = valid_lens.repeat(shape[1])
else:
valid_lens = valid_lens.reshape(-1)
# On the last axis, replace masked elements with a very large negative
# value, whose exponentiation outputs 0
X = npx.sequence_mask(X.reshape(-1, shape[-1]), valid_lens, True,
value=-1e6, axis=1)
return npx.softmax(X).reshape(shape)
# + [markdown] origin_pos=6
# To demonstrate how this function works,
# consider a minibatch of two $2 \times 4$ matrix examples,
# where the valid lengths for these two examples
# are two and three, respectively.
# As a result of the masked softmax operation,
# values beyond the valid lengths
# are all masked as zero.
#
# + origin_pos=7 tab=["mxnet"]
masked_softmax(np.random.uniform(size=(2, 2, 4)), np.array([2, 3]))
# + [markdown] origin_pos=9
# Similarly, we can also
# use a two-dimensional tensor
# to specify valid lengths
# for every row in each matrix example.
#
# + origin_pos=10 tab=["mxnet"]
masked_softmax(np.random.uniform(size=(2, 2, 4)), np.array([[1, 3], [2, 4]]))
# + [markdown] origin_pos=12
# ## Additive Attention
# :label:`subsec_additive-attention`
#
# In general,
# when queries and keys are vectors of different lengths,
# we can use additive attention
# as the scoring function.
# Given a query $\mathbf{q} \in \mathbb{R}^q$
# and a key $\mathbf{k} \in \mathbb{R}^k$,
# the *additive attention* scoring function
#
# $$a(\mathbf q, \mathbf k) = \mathbf w_v^\top \text{tanh}(\mathbf W_q\mathbf q + \mathbf W_k \mathbf k) \in \mathbb{R},$$
# :eqlabel:`eq_additive-attn`
#
# where
# learnable parameters
# $\mathbf W_q\in\mathbb R^{h\times q}$, $\mathbf W_k\in\mathbb R^{h\times k}$, and $\mathbf w_v\in\mathbb R^{h}$.
# Equivalent to :eqref:`eq_additive-attn`,
# the query and the key are concatenated
# and fed into an MLP with a single hidden layer
# whose number of hidden units is $h$, a hyperparameter.
# By using $\tanh$ as the activation function and disabling
# bias terms,
# we implement additive attention in the following.
#
# + origin_pos=13 tab=["mxnet"]
#@save
class AdditiveAttention(nn.Block):
"""Additive attention."""
def __init__(self, num_hiddens, dropout, **kwargs):
super(AdditiveAttention, self).__init__(**kwargs)
# Use `flatten=False` to only transform the last axis so that the
# shapes for the other axes are kept the same
self.W_k = nn.Dense(num_hiddens, use_bias=False, flatten=False)
self.W_q = nn.Dense(num_hiddens, use_bias=False, flatten=False)
self.w_v = nn.Dense(1, use_bias=False, flatten=False)
self.dropout = nn.Dropout(dropout)
def forward(self, queries, keys, values, valid_lens):
queries, keys = self.W_q(queries), self.W_k(keys)
# After dimension expansion, shape of `queries`: (`batch_size`, no. of
# queries, 1, `num_hiddens`) and shape of `keys`: (`batch_size`, 1,
# no. of key-value pairs, `num_hiddens`). Sum them up with
# broadcasting
features = np.expand_dims(queries, axis=2) + np.expand_dims(
keys, axis=1)
features = np.tanh(features)
# There is only one output of `self.w_v`, so we remove the last
# one-dimensional entry from the shape. Shape of `scores`:
# (`batch_size`, no. of queries, no. of key-value pairs)
scores = np.squeeze(self.w_v(features), axis=-1)
self.attention_weights = masked_softmax(scores, valid_lens)
# Shape of `values`: (`batch_size`, no. of key-value pairs, value
# dimension)
return npx.batch_dot(self.dropout(self.attention_weights), values)
# + [markdown] origin_pos=15
# Let us demonstrate the above `AdditiveAttention` class
# with a toy example,
# where shapes (batch size, number of steps or sequence length in tokens, feature size)
# of queries, keys, and values
# are ($2$, $1$, $20$), ($2$, $10$, $2$),
# and ($2$, $10$, $4$), respectively.
# The attention pooling output
# has a shape of (batch size, number of steps for queries, feature size for values).
#
# + origin_pos=16 tab=["mxnet"]
queries, keys = np.random.normal(0, 1, (2, 1, 20)), np.ones((2, 10, 2))
# The two value matrices in the `values` minibatch are identical
values = np.arange(40).reshape(1, 10, 4).repeat(2, axis=0)
valid_lens = np.array([2, 6])
attention = AdditiveAttention(num_hiddens=8, dropout=0.1)
attention.initialize()
attention(queries, keys, values, valid_lens)
# + [markdown] origin_pos=18
# Although additive attention contains learnable parameters,
# since every key is the same in this example,
# the attention weights are uniform,
# determined by the specified valid lengths.
#
# + origin_pos=19 tab=["mxnet"]
d2l.show_heatmaps(attention.attention_weights.reshape((1, 1, 2, 10)),
xlabel='Keys', ylabel='Queries')
# + [markdown] origin_pos=20
# ## Scaled Dot-Product Attention
#
# A more computationally efficient
# design for the scoring function can be
# simply dot product.
# However,
# the dot product operation
# requires that both the query and the key
# have the same vector length, say $d$.
# Assume that
# all the elements of the query and the key
# are independent random variables
# with zero mean and unit variance.
# The dot product of
# both vectors has zero mean and a variance of $d$.
# To ensure that the variance of the dot product
# still remains one regardless of vector length,
# the *scaled dot-product attention* scoring function
#
#
# $$a(\mathbf q, \mathbf k) = \mathbf{q}^\top \mathbf{k} /\sqrt{d}$$
#
# divides the dot product by $\sqrt{d}$.
# In practice,
# we often think in minibatches
# for efficiency,
# such as computing attention
# for
# $n$ queries and $m$ key-value pairs,
# where queries and keys are of length $d$
# and values are of length $v$.
# The scaled dot-product attention
# of queries $\mathbf Q\in\mathbb R^{n\times d}$,
# keys $\mathbf K\in\mathbb R^{m\times d}$,
# and values $\mathbf V\in\mathbb R^{m\times v}$
# is
#
#
# $$ \mathrm{softmax}\left(\frac{\mathbf Q \mathbf K^\top }{\sqrt{d}}\right) \mathbf V \in \mathbb{R}^{n\times v}.$$
# :eqlabel:`eq_softmax_QK_V`
#
# In the following implementation of the scaled dot product attention, we use dropout for model regularization.
#
# + origin_pos=21 tab=["mxnet"]
#@save
class DotProductAttention(nn.Block):
"""Scaled dot product attention."""
def __init__(self, dropout, **kwargs):
super(DotProductAttention, self).__init__(**kwargs)
self.dropout = nn.Dropout(dropout)
# Shape of `queries`: (`batch_size`, no. of queries, `d`)
# Shape of `keys`: (`batch_size`, no. of key-value pairs, `d`)
# Shape of `values`: (`batch_size`, no. of key-value pairs, value
# dimension)
# Shape of `valid_lens`: (`batch_size`,) or (`batch_size`, no. of queries)
def forward(self, queries, keys, values, valid_lens=None):
d = queries.shape[-1]
# Set `transpose_b=True` to swap the last two dimensions of `keys`
scores = npx.batch_dot(queries, keys, transpose_b=True) / math.sqrt(d)
self.attention_weights = masked_softmax(scores, valid_lens)
return npx.batch_dot(self.dropout(self.attention_weights), values)
# + [markdown] origin_pos=23
# To demonstrate the above `DotProductAttention` class,
# we use the same keys, values, and valid lengths from the earlier toy example
# for additive attention.
# For the dot product operation,
# we make the feature size of queries
# the same as that of keys.
#
# + origin_pos=24 tab=["mxnet"]
queries = np.random.normal(0, 1, (2, 1, 2))
attention = DotProductAttention(dropout=0.5)
attention.initialize()
attention(queries, keys, values, valid_lens)
# + [markdown] origin_pos=26
# Same as in the additive attention demonstration,
# since `keys` contains the same element
# that cannot be differentiated by any query,
# uniform attention weights are obtained.
#
# + origin_pos=27 tab=["mxnet"]
d2l.show_heatmaps(attention.attention_weights.reshape((1, 1, 2, 10)),
xlabel='Keys', ylabel='Queries')
# + [markdown] origin_pos=28
# ## Summary
#
# * We can compute the output of attention pooling as a weighted average of values, where different choices of the attention scoring function lead to different behaviors of attention pooling.
# * When queries and keys are vectors of different lengths, we can use the additive attention scoring function. When they are the same, the scaled dot-product attention scoring function is more computationally efficient.
#
#
#
# ## Exercises
#
# 1. Modify keys in the toy example and visualize attention weights. Do additive attention and scaled dot-product attention still output the same attention weights? Why or why not?
# 1. Using matrix multiplications only, can you design a new scoring function for queries and keys with different vector lengths?
# 1. When queries and keys have the same vector length, is vector summation a better design than dot product for the scoring function? Why or why not?
#
# + [markdown] origin_pos=29 tab=["mxnet"]
# [Discussions](https://discuss.d2l.ai/t/346)
#
| scripts/d21-en/mxnet/chapter_attention-mechanisms/attention-scoring-functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 헤드리스 크롬
#
# 구글 인기영화 - 할인 중인 영화
# ### User-Agent 설정하기
# User-Agent 설정 없을 시, header 정보에 HeadlessChrome 이 들어가게 되어 해당 서버에서 접근을 막을 수 있어
# 아래와 같이 User-Agent를 설정해준다
import time
from selenium import webdriver
from selenium.webdriver.chrome.options import Options
options = webdriver.ChromeOptions()
options.headless = True
options.add_argument('--headless')
options.add_argument('--no-sandbox')
options.add_argument('--disable-dev-shm-usage')
options.add_argument("window-size=1920x1080") # 가상화면 크기
options.add_argument("user-agent=Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/84.0.4147.135 Safari/537.36")
browser = webdriver.Chrome(options=options)
#browser.maximize_window() # 최대화
url = "https://www.whatismybrowser.com/detect/what-is-my-user-agent"
browser.get(url)
detected_value = browser.find_element_by_id("detected_value")
print (detected_value.text)
# 브라우저 종료
browser.quit()
| shared/notebooks/2_web-scraping-I/.ipynb_checkpoints/21_docker_headless_selenium_scraping_user_agent-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
print("Hello there")
tax = 12.5/100
price= 100
price*tax
price+_
round(_,2)
"hello there"
"what's up?? he said - \" he is not going to any kind of ceremony anymore\" "
'isn\'t is a good decision?'
print(r'C:dir\files\downloads\newfile\cs50.org')
print("""\
\
\
hello there i am doing great.
can you tell me what to do next?
i am really confused nowadays :(
""")
4*'im'+'um'
'py'+'thon'
print('Hello there i am talking about a funny issue. please leave it dear btw.'
'and again no need to think about it.'
'its fishy and clumsy hehe')
prefix ='py'
prefix+'thon'
word ='python'
word[-3]
word[2:-2]
word[:42]
word[42:]
'J'+word[1:]
s='Jsfhkajsfbhdfg oanfdnakjh nfbaskjbfk'
len(s)
square=['name','age','hfakjf','aoppp','subject']
print(square)
len(square)
# +
#fibonacci series
a,b=0,2
while a<100:
print(a,end=" ")
a,b=b,a+b
# +
#factorial numbers
a=int(input("Enter any number to have the factorial of it : "))
fact=1
while a>0:
fact = fact*a
a=a-1
print('factorial is :',fact)
# -
i =256*256
print("The value of i is : ",i)
# +
a=int(input("Please enter any number between 0 to 10 : "))
if a<0:
print("This is less than zero , so it's not valid")
elif a>10:
print("This is less than zero , so it's not valid")
else :
print("You have entered a valid number which is : ",a)
# -
w=["name",'ajhf','ajf','asjhfdk']
for i in w[:]:
print(i,end=" ")
for i in range(len(w)):
print(i,w[i])
print(range(10))
list(range(10))
for n in range(2,10):
for x in range(2,n):
if n%x == 0:
print(n,' equals',x,'*',n//x)
break
else :
print(n,' is a prime number')
# +
even =[]
odd =[]
for i in range(2,30):
if i%2==0 :
print('Found a even number : ',i)
even.append(i)
else :
print("odd number : ",i)
odd.append(i)
print("even is : ",even)
print("Odd is : ",odd)
# +
def okk(prompt, retry =5, reminder = "try again later !"):
while True:
ok = input(prompt)
if ok in ('ye','yes','y'):
return True
if ok in ('n','nope','na','no'):
return False
retry = retry - 1
if retry <= 0:
print("your chance is over")
else :
raise ValueError('Invalid user response')
print(reminder)
okk('enter y or no')
# -
def ask_ok(prompt, retries=4, reminder='Please try again!'):
while True:
ok = input(prompt)
if ok in ('y', 'ye', 'yes'):
return True
if ok in ('n', 'no', 'nop', 'nope'):
return False
retries = retries - 1
if retries < 0:
raise ValueError('invalid user response')
print(reminder)
ask_ok("y or n")
# +
a=6
def n(a,l=[]):
l.append(a)
return l
n(a)
a=5
n(a)
# +
a=5
def n(arg=a):
print(a)
a=10
n()
a=12
n(a)
# -
dict={'name':'nam','age':'22'}
print(dict)
# +
a=[3,10]
list(range(*a)) # * is very important here
# -
pairs = [(1, 'one'), (2, 'two'), (3, 'three'), (4, 'four')]
pairs.sort(key=lambda pair : pair[1])
print(pairs)
list=[1,2,3,4,5,6,7,8]
print(list)
list.append(99)
print(list)
list.extend("y")
print(list)
list.insert(3,"mm")
print(list)
list.remove("mm")
print(list)
list.pop()
print(list)
list.clear()
print(list)
list=[1, 2, 3, 4, 5, 6, 7, 8, 99, 99, 'n', 'a', 'y', 'e', 'm', 'n', 'a', 'y', 'y']
list.count('y')
list.reverse()
print(list)
a=list.copy()
print(a)
list.reverse()
print(list)
print(list)
list=[1,34,5,6,78,89,88]
print(list)
list.insert(2,'nick sessman')
print(list)
list.pop(5)
print(list)
list.index(88)
# +
list=['y', 'y', 'a', 'n', 'm', 'e', 'y', 'a', 'n']
list.sort()
print(list)
# -
l=[1, 34, 5, 6, 78, 89, 88]
l.sort()
print(l)
from collections import deque
queue = deque(["eric",'adam','john','rick','peter'])
queue.append('obama')
print(queue)
queue.popleft()
print(queue)
squares = [x**2 for x in range(10)]
print(squares)
[(x,y) for x in [1,2,3,4,5] for y in [6,5,4] if x!=y]
# +
from math import pi
[str(round(pi,i)) for i in range(6)]
# +
matrix = [
[1, 2, 3, 4], [5, 6, 7, 8],
[9, 10, 11, 12],
]
[[row[i] for row in matrix] for i in range(4)]
# -
l=[1,2,3,4,5,6,7,88,9]
print(l)
del l[0]
print(l)
v = ([1, 2, 3], [3, 2, 1])
print(v)
s="atrinfdsfjb", #trailing comma is the difference between tuple
print(len(s))
print(s)
t='s','d','f','g'
print(t)
basket = {'apple', 'orange', 'apple', 'pear', 'orange', 'banana'}
print(basket)
s=(1,2,3,4,5,4,3)
print(s)
b=set("ahsjfdk")
print(b)
a={3,4,5,6,7}
print(a)
a=sorted(set('namd',))
print(a)
# +
#set comprehension just like list comprehension
s={x**2 for x in range(10)}
print(sorted(s))
# -
a=dict((('naem','nayem'),('age',22),('op','programmer')))
print(a)
dict([('sape', 4139), ('guido', 4127), ('jack', 4098)])
{x: x**2 for x in range(10)}
>>> dict(sape=4139, guido=4127, jack=4098)
>>> for i, v in enumerate(['tic', 'tac', 'toe']):
print(i,v, end=" | ")
questions = ['name', 'quest', 'favorite color']
answers = ['lancelot', 'the holy grail', 'blue']
for q, a in zip(questions, answers):
print("What is your {}? it is {}".format(q,a))
print("my name is %s and i am {} years old and my fav number is : {} {}".format('nayem',22,99.99))
for i in reversed(range(1,100,5)):
print(i,end=" ")
basket = ['apple', 'orange', 'apple', 'pear', 'orange', 'banana']
for f in sorted(set(basket)):
print(f)
# +
import math
raw_data = [56.2, float('NaN'), 51.7, 55.3, 52.5, float('NaN'), 47.8]
filtered_data = []
for value in raw_data:
if not math.isnan(value):
filtered_data.append(value)
print(filtered_data)
# -
r= [56.2,float("NaN"), 51.7, 55.3, 52.5, float('NaN'), 47.8]
print(r)
type(float('nan'))
a=float('Nan')
print(a)
s ,s1,s2 = 'Novelman','Novel',"nayemnovel"
print(s,s1,s2)
nn = s or s1 or s2
print(nn)
nn = s1 and s2 and s
print(nn)
(1, 2, 3, 4) < (1, 2, 4)
'ABC' < 'C' < 'Pascal' < 'Python'
(1, 2, 3, 4) < (1, 2, 4)
(1, 2) < (1, 2, -1)
(1, 2, 3) == (1.0, 2.0, 3.0)
(1, 2, ('aa', 'ab')) < (1, 2, ('abc', 'a'), 4)
(1, 2, 883) < (1, 2, 66)
'ABC' < 'C' < 'Pascal' < 'Python'
#input and output
year =2015
event ="referendum"
print(f"results of the {year} is {event}")
yes =7437
no=3582
p = yes/(yes+no)
print("the percentage is {:2.4} where yes votes are : {:5}".format(p,yes))
s="hello world"
print(s)
str(s)
repr(s)
str(2/9)
repr(2/9)
x = 10 * 3.25
y = 200 * 200
s = 'The value of x is '+ repr(x) +' and y is ' + repr(y)
print(s)
import math
print(f'The value of pi is approximately {math.pi:.10f}.')
# Other modifiers can be used to convert the value before it is formatted. '!a' applies ascii(), '!s' applies str(), and '!r' applies repr():
#
#
animals = 'eels'
>>> print(f'My hovercraft is full of {animals}.')
>>> print(f'My hovercraft is full of {animals!r}.')
for x in range(1,10):
print("{0:4d} {1:5d} {2:20d}".format(x, x*x, x*x*x))
for x in range(1, 11):
print(repr(x).rjust(2), repr(x*x).rjust(3))
'12'.zfill(5)
'-3.14'.zfill(7)
>>> '3.14159265359'.zfill(5)
import math
>>> print('The value of pi is approximately %5.3f.' % math.pi)
while True:
try:
a=str(input("Enter any number : "))
print(a)
break
except ValueError:
print("Opps ! that was not a valid number....try again ! ")
| tutorial_pyhton_org.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="5UyFJYXOx-WH" slideshow={"slide_type": "slide"}
# # Syllabus (https://bit.ly/intro_python_00)
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 1 - Intro (https://bit.ly/intro_python_01)
# * What's a program?
# * Hello World!
# * Basic Input and Output
# * Interpreted vs. compiled languages
# * Ways to run Python (notebooks, via the interactive console, via an editor (PyCharm))
# * The debug loop
# * Basic input and output
# * Comments
# * Basic types
# * integers (ints)
# * strings
# * floating point numbers (floats)
# * ZyBook: Chapter 1
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 2 - Variables and Expressions (https://bit.ly/intro_python_02)
# * Objects
# * Variables
# * Assignment is not equals
# * Copying References
# * Variable Names
# * Expressions
# * Statements
# * Operators
# * Abbreviated assignment
# * Logical operators
# * Order of operations
# * ZyBook: Chapter 2
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 3 - More Types (https://bit.ly/intro_python_03)
# * More strings
# * Type Conversions
# * Automatic Type Conversion
# * More Input
# * List basics
# * Dictionary Basics
# * ZyBook: Chapter 3
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 4 - Conditionals and Branching (https://bit.ly/intro_python_04)
# * If statements
# * Else
# * Elif
# * Pass
# * First looping control flow:
# * While loops
# * ZyBook: Chapter 4
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 5 - Loops (https://bit.ly/intro_python_05)
# * More control flow:
# * For loops and list iteration
# * Nested loops
# * Break statement
# * Continue statement
# * Look at some complex examples of control flow that use control statements to get more comfortable
# * ZyBook: Chapter 5
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 6 - Functions (https://bit.ly/intro_python_06)
# * Function definitions and function calls
# * Return values are optional
# * Docstrings
# * Return can be used for control flow
# * None and the default return value
# * Return can often sub for break
# * Functions can call other functions - the stack
# * ZyBook: Chapter 6
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 7 - Functions Continued (https://bit.ly/intro_python_07)
# * Functions continued:
# * Functions, namespaces and score
# * Optional functional arguments and other tricks
# * A few common mistakes about functions
# * Some examples
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 8 - Strings (https://bit.ly/intro_python_08)
# * Strings:
# * String operators
# * Length function
# * Slicing
# * Immutability
# * String comparison
# * For loops
# * In operator
# * Convenience functions
# * Find
# * Split
# * Join
# * etc.
# * Format method
# * Formatting numbers
# * ZyBook: Chapter 7
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 9 - Tuples, Lists and Dictionaries (https://bit.ly/intro_python_09)
# * Tuples (recap many things introduced with respect to strings)
# * Lists
# * List comprehensions
# * More Dictionaries
# * Sets
# * ZyBook: Chapter 8
# + [markdown] slideshow={"slide_type": "slide"}
# # Midterm in class (everything in lectures 1 - 9)
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 10 - Modules (https://bit.ly/intro_python_10)
# * Finishing up sequences:
# * Iterators vs. lists
# * Generators and the yield keyword
# * Modules:
# * Some useful modules
# * Hierarchical namespaces
# * Making your own modules
# * The main() function
# * Parsing user arguments with the argparse module
# * PEP 8 (v. briefly)
# * A revisit to debugging, now that we're writing longer programs:
# * Looking at different error types (syntax, runtime, logical)
# * ZyBook: Chapter 11
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 11 - Files (https://bit.ly/intro_python_11)
# * Writing Files
# * Reading files
# * With keyword
# * For loops on file handles
# * Binary files
# * Large worked example (Fasta file processing)
# * Directories
# * Web file retrieval
# * ZyBook: Chapter 12
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 12 - Classes and Objects (https://bit.ly/intro_python_12)
# * The basics of Python classes and objects:
# * Classes and Objects
# * The \__init__ constructor method
# * Object membership: Dot notation and classes
# * Everything is an object in Python!
# * Methods: adding functions to class and the self argument
# * Object vs. class variables
# * Objects Mutability
# * Is vs. ==
# * ZyBook: Chapter 9
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 13 - Objects and Polymorphism (https://bit.ly/intro_python_13)
# * Python objects:
# * Polymorphism
# * Polymorphism by reimplementing basic object functionality:
# * Redefining the \__str__ method
# * How Python implements ''==" for objects
# * General operator overloading, including comparison operators
# * Copying objects
# * No Zybooks!
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 14 - Inheritance (https://bit.ly/intro_python_14)
# * Inheritance
# * Object composition
# * Inheritance vs. composition: aka: 'Is a' vs. 'has a'.
# * An Example of OOP: Implementing the card game Old Maid
# * ZyBook: Chapter 13
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 15 - More Functions and Recursion (https://bit.ly/intro_python_15)
# * Recursion
# * Consolidate: Look at some more complex examples of recursive control flow that use functions and control statements to get more comfortable
# * Lambda functions
# * Functions as arguments
# * ZyBook: Chapter 14
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 16 - Exceptions and Unit Testing (https://bit.ly/intro_python_16)
# * Unit-testing with unitest
# * Exceptions and error handling
# * ZyBook: Chapter 10
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 17 - Search Algorithms (https://bit.ly/intro_python_17)
# * A couple of simple list algorithms:
# * Linear search
# * Binary search
# * A very brief introduction to runtime analysis and Big O Notation
# * Word counting with dictionaries
# * Zybook Chapter 16
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 18 - Intro to data science (https://bit.ly/intro_python_18)
# Worked example intro to Python data science
# * NumPy
# * MatPlotLib (to make plots)
# * Pandas
# * Sklearn
# * Zybook Chapter 15
# + [markdown] slideshow={"slide_type": "slide"}
# # Revision for final (everything!)
| lecture_notebooks/L-- Syllabus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plot
# 07.20: 目标是寻找plot绘制的mark不会变形的秘密
# 将一组点描绘在一个平面直角坐标系中最简单的办法就是每一个点的对应位置做一个记号。
# + jupyter={"source_hidden": true}
"""
Plot y versus x as lines and/or markers.
Call signatures::
plot([x], y, [fmt], *, data=None, **kwargs)
plot([x], y, [fmt], [x2], y2, [fmt2], ..., **kwargs)
The coordinates of the points or line nodes are given by *x*, *y*.
The optional parameter *fmt* is a convenient way for defining basic
formatting like color, marker and linestyle. It's a shortcut string
notation described in the *Notes* section below.
>>> plot(x, y) # plot x and y using default line style and color
>>> plot(x, y, 'bo') # plot x and y using blue circle markers
>>> plot(y) # plot y using x as index array 0..N-1
>>> plot(y, 'r+') # ditto, but with red plusses
You can use `.Line2D` properties as keyword arguments for more
control on the appearance. Line properties and *fmt* can be mixed.
The following two calls yield identical results:
>>> plot(x, y, 'go--', linewidth=2, markersize=12)
>>> plot(x, y, color='green', marker='o', linestyle='dashed',
... linewidth=2, markersize=12)
When conflicting with *fmt*, keyword arguments take precedence.
**Plotting labelled data**
There's a convenient way for plotting objects with labelled data (i.e.
data that can be accessed by index ``obj['y']``). Instead of giving
the data in *x* and *y*, you can provide the object in the *data*
parameter and just give the labels for *x* and *y*::
>>> plot('xlabel', 'ylabel', data=obj)
All indexable objects are supported. This could e.g. be a `dict`, a
`pandas.DataFrame` or a structured numpy array.
**Plotting multiple sets of data**
There are various ways to plot multiple sets of data.
- The most straight forward way is just to call `plot` multiple times.
Example:
>>> plot(x1, y1, 'bo')
>>> plot(x2, y2, 'go')
- Alternatively, if your data is already a 2d array, you can pass it
directly to *x*, *y*. A separate data set will be drawn for every
column.
Example: an array ``a`` where the first column represents the *x*
values and the other columns are the *y* columns::
>>> plot(a[0], a[1:])
- The third way is to specify multiple sets of *[x]*, *y*, *[fmt]*
groups::
>>> plot(x1, y1, 'g^', x2, y2, 'g-')
In this case, any additional keyword argument applies to all
datasets. Also this syntax cannot be combined with the *data*
parameter.
By default, each line is assigned a different style specified by a
'style cycle'. The *fmt* and line property parameters are only
necessary if you want explicit deviations from these defaults.
Alternatively, you can also change the style cycle using
:rc:`axes.prop_cycle`.
Parameters
----------
x, y : array-like or scalar
The horizontal / vertical coordinates of the data points.
*x* values are optional and default to ``range(len(y))``.
Commonly, these parameters are 1D arrays.
They can also be scalars, or two-dimensional (in that case, the
columns represent separate data sets).
These arguments cannot be passed as keywords.
fmt : str, optional
A format string, e.g. 'ro' for red circles. See the *Notes*
section for a full description of the format strings.
Format strings are just an abbreviation for quickly setting
basic line properties. All of these and more can also be
controlled by keyword arguments.
This argument cannot be passed as keyword.
data : indexable object, optional
An object with labelled data. If given, provide the label names to
plot in *x* and *y*.
.. note::
Technically there's a slight ambiguity in calls where the
second label is a valid *fmt*. ``plot('n', 'o', data=obj)``
could be ``plt(x, y)`` or ``plt(y, fmt)``. In such cases,
the former interpretation is chosen, but a warning is issued.
You may suppress the warning by adding an empty format string
``plot('n', 'o', '', data=obj)``.
Returns
-------
list of `.Line2D`
A list of lines representing the plotted data.
Other Parameters
----------------
scalex, scaley : bool, default: True
These parameters determine if the view limits are adapted to the
data limits. The values are passed on to `autoscale_view`.
**kwargs : `.Line2D` properties, optional
*kwargs* are used to specify properties like a line label (for
auto legends), linewidth, antialiasing, marker face color.
Example::
>>> plot([1, 2, 3], [1, 2, 3], 'go-', label='line 1', linewidth=2)
>>> plot([1, 2, 3], [1, 4, 9], 'rs', label='line 2')
If you make multiple lines with one plot call, the kwargs
apply to all those lines.
Here is a list of available `.Line2D` properties:
%(_Line2D_docstr)s
See Also
--------
scatter : XY scatter plot with markers of varying size and/or color (
sometimes also called bubble chart).
Notes
-----
**Format Strings**
A format string consists of a part for color, marker and line::
fmt = '[marker][line][color]'
Each of them is optional. If not provided, the value from the style
cycle is used. Exception: If ``line`` is given, but no ``marker``,
the data will be a line without markers.
Other combinations such as ``[color][marker][line]`` are also
supported, but note that their parsing may be ambiguous.
**Markers**
============= ===============================
character description
============= ===============================
``'.'`` point marker
``','`` pixel marker
``'o'`` circle marker
``'v'`` triangle_down marker
``'^'`` triangle_up marker
``'<'`` triangle_left marker
``'>'`` triangle_right marker
``'1'`` tri_down marker
``'2'`` tri_up marker
``'3'`` tri_left marker
``'4'`` tri_right marker
``'s'`` square marker
``'p'`` pentagon marker
``'*'`` star marker
``'h'`` hexagon1 marker
``'H'`` hexagon2 marker
``'+'`` plus marker
``'x'`` x marker
``'D'`` diamond marker
``'d'`` thin_diamond marker
``'|'`` vline marker
``'_'`` hline marker
============= ===============================
**Line Styles**
============= ===============================
character description
============= ===============================
``'-'`` solid line style
``'--'`` dashed line style
``'-.'`` dash-dot line style
``':'`` dotted line style
============= ===============================
Example format strings::
'b' # blue markers with default shape
'or' # red circles
'-g' # green solid line
'--' # dashed line with default color
'^k:' # black triangle_up markers connected by a dotted line
**Colors**
The supported color abbreviations are the single letter codes
============= ===============================
character color
============= ===============================
``'b'`` blue
``'g'`` green
``'r'`` red
``'c'`` cyan
``'m'`` magenta
``'y'`` yellow
``'k'`` black
``'w'`` white
============= ===============================
and the ``'CN'`` colors that index into the default property cycle.
If the color is the only part of the format string, you can
additionally use any `matplotlib.colors` spec, e.g. full names
(``'green'``) or hex strings (``'#008000'``).
"""
# -
self._get_lines = _process_plot_var_args(self)
# Uses a custom implementation of data-kwarg handling in
# _process_plot_var_args.
@docstring.dedent_interpd
def plot(self, *args, scalex=True, scaley=True, data=None, **kwargs):
kwargs = cbook.normalize_kwargs(kwargs, mlines.Line2D)
lines = [*self._get_lines(*args, data=data, **kwargs)]
for line in lines:
self.add_line(line)
self._request_autoscale_view(scalex=scalex, scaley=scaley)
return lines
| mpl_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import requests
page = requests.get('http://scores.sify.com/?ref=deskcric')
soup = BeautifulSoup(page.content,'html.parser')
match_name=soup.find_all('a',{'class':'scroll'})[0].string
match_name
match_location=soup.select_one('div.scoresbox-center span').string
match_location
# finding team 1 name
live_match_link=soup.select_one('div.scoresbox-center h2 a').attrs.get('href')
live_match_link
page_curr_match = requests.get(live_match_link)
soup_live = BeautifulSoup(page_curr_match.content,'html.parser')
teams_info=soup_live.find_all('div',{'class':'team-live'})[0].find('h1').string
teams_info
soup_live.find_all('div',{'class':'team-live'})[0]
items=soup_live.find_all('div',{'class':'team-live'},recursive=True)[0]
# you need to extract vanue here
soup_live.find_all('div',{'class':'team-live'},recursive=True)[0]
for i in items:
print(i)
print('--------------------')
bat_team_info=soup_live.select_one('div.team-live-venue h2').string
bat_team_info
# +
bat_team_name=''
bat_team_run=''
for i in bat_team_info:
if i is not ' ':
bat_team_name = bat_team_name + i
else:
break
bat_team_name
# -
bat_team_run=''
for i in range(0,len(bat_team_info)):
if bat_team_info[i] is ' ':
for j in range(i+1,len(bat_team_info)):
bat_team_run = bat_team_run + bat_team_info[j]
bat_team_run
# run rat and needs that run to win the match
try:
target_split=soup_live.find_all('div',{'class':'team-live-venue'})[0].find('p').string.split('|')
target=target_split[1]
print(target)
except:
print('Match yet to start')
# finding the team1 members
team1_finder=soup_live.find_all('div',{'id':'team1'})[0].find_all('ul')
team1_finder
# +
#team1_finder[0].find_all('li')
# -
team1_squad=[]
for i in team1_finder[0].find_all('li'):
team1_squad.append(i.find_all('a')[0].string)
team1_squad
# finding the team2 members
team2_finder=soup_live.find_all('div',{'id':'team2'})[0].find_all('ul')
team2_finder
team2_squad=[]
for i in team2_finder[0].find_all('li'):
team2_squad.append(i.find_all('a')[0].string)
team2_squad
over=soup.find_all('h3',{'id':'batteamnameruns1'})[0].text.split('(')[1].split(')')[0]
# batsman info
batsman1=soup.find_all('h4',{'id':'batsmanbowlerdetails1'})[0].find_all('p')[0].find('a').string
batsman1
a=soup.find_all('h4',{'id':'batsmanbowlerdetails1'})[0].find_all('p')[0]
try:
batsman2=str(soup.find_all('h4',{'id':'batsmanbowlerdetails1'})[0].find_all('p')[0]).split('|')[1].split('>')[1].split('<')[0]
except:
batsman2='wait for match to start'
soup.find_all('div',{'class':'scoresbox-center'})[0].find_all('h2')[0]
| sify_scrap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Case study: K-means
#
# K-means is a simple algorithm to cluster data – that is to identify groups of similar objects based only on their properties. The algorithm is best-illustrated by the following graph:
#
# <img src="../images/kmeans_illustration.png" alt="drawing" width=800 >
#
#
#
# ## Loading data
#
# We first need to load sample data
# +
import numpy as np
import matplotlib.pylab as plt
data = np.loadtxt('../data/kmeans_data.csv')
data.shape
# -
# To visualise the data we can use the scatter function from matplotlib package:
plt.scatter(data[:, 0], data[:, 1], s=40)
# ## Initialisation
#
# In the first step of the algorithm we need to initialise the centers of the clusters. We will initialise them randomly but consistently with the mean and standard deviation of the data:
K = 3
centroids = np.random.randn(K, 2)
# To center the cluster centroids on the data it’s better to normalise to the mean and standard deviation of the data:
centroids = centroids * np.std(data, 0)
centroids = centroids + np.mean(data, 0)
# Let’s now plot the data and the random cluster centers on the same figure:
plt.scatter(data[:, 0], data[:, 1], s=40)
plt.scatter(centroids[:, 0], centroids[:, 1], c=np.arange(3), s=100)
# We now need to assign each point to the closest cluster center. <br> First, we will calculate the Euclidean distance of each point to each of the centers. <br>
#
# For this we can use the [broadcasting](broadcasting.ipynb).
#
# The term broadcasting describes how numpy treats arrays with different shapes during arithmetic operations. Subject to certain constraints, the smaller array is “broadcast” across the larger array so that they have compatible shapes.
data.shape
centroids.shape
data[:, np.newaxis, :].shape
deltas = data[:, np.newaxis, :] - centroids
deltas.shape
distances = np.sqrt(np.sum((deltas) ** 2, 2))
# For each data point we find the center with minimum distance. We can use the argmin method with the axis argument:
closest = distances.argmin(1)
# Now we plot the centroids and data points with the color-code reflecting cluster membership:
plt.scatter(data[:, 0], data[:, 1], s=40, c=closest)
plt.scatter(centroids[:, 0], centroids[:, 1], c=np.arange(3), s=100)
# ## $\color{green}{\text{Excercise}}$ Calculate new cluster centers
# Given the array of cluster assignments closest calculate the center coordinates of the first cluster (index 0). <br> __Note__: To calculate new centers of the clusters, we average all points belonging to that cluster. We can use a boolean mask
# To repeat it for all clusters we can use a for loop or list comprehension. Since the number of clusters is usually much smaller than the number of data points, this for loop won’t affect the performance of our algorithm:
centroids = np.array([data[closest == i, :].mean(0) for i in range(3)])
# Lets check the positions of new centers and assignment of points to clusters.
plt.scatter(data[:, 0], data[:, 1], s=40, c=closest)
plt.scatter(centroids[:, 0], centroids[:, 1], c=np.arange(3), s=100)
# ## Iterations
# Now we can repeat the steps of assigning point to clusters and updating the cluster centers iteratively and watch the progress of the algorithm:
for iteration in range(5):
# assign points to clusters
deltas = data[:, np.newaxis, :] - centroids
distances = np.sqrt(np.sum((deltas) ** 2, 2))
closest = distances.argmin(1)
# calculate new centroids
centroids = np.array([data[closest == i, :].mean(0) for i in range(3)])
# ## Single cluster?
# Note that sometimes the algorithm can produce degenerate results – all of the points will be assigned to a single cluster (or final number of clusters will be less than K). This is one of drawbacks of K-means with random initialisations. A possible solution is to repeat the algorithm with other initialisations and find the best cluster assignment, but better solutions exist.
# ## Putting it all together
#
# Our final script will look as the following:
# +
import numpy as np
import matplotlib.pyplot as plt
data = np.loadtxt('../data/kmeans_data.csv')
# randomly initalize the centroids
K = 3
centroids = np.random.randn(K, 2)
centroids = centroids * np.std(data, 0)
centroids = centroids + np.mean(data, 0)
for iteration in range(5):
# assign points to clusters
deltas = data[:, np.newaxis, :] - centroids
distances = np.sqrt(np.sum((deltas) ** 2, 2))
closest = distances.argmin(1)
# calculate new centroids
centroids = np.array([data[closest == i, :].mean(0) for i in range(K)])
# plot
plt.scatter(data[:, 0], data[:, 1], s=40, c=closest)
plt.scatter(centroids[:, 0], centroids[:, 1], c=np.arange(K), s=100)
# -
# ## $\color{green}{\text{Excercise}}$ Stopping criterion
# After each iteration test whether any point changes their cluster membership. Stop the algorithm if convergence was reached i.e. clusters do not change after the re-assignment step.
# ## $\color{green}{\text{Excercise}}$ Choice of K
#
# Check whether the algorithm works for any K. Try using K > 3. What happens then?
# ## $\color{green}{\text{Excercise}}$ Memory or speed
#
# Replace the assignment and calculation of new clusters with a for loop. Which implementation would be preferable for small (few observations and dimensions) and which for large datasets (large number of observations and dimensions).
# [Previous: Stacking](stacking.ipynb) <br>[Next: Savez() and load()](savez.ipynb)
| Day_1_Scientific_Python/numpys/k_means.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#This code creates list of all files within same directory and
#deletes every other file, starting with the second file (index=1)
# +
import os
import glob
import numpy as np
filelist=glob.glob('B*****.txt')
number=np.arange(1,len(filelist)+1)
even_check=number[0::2]
print(even_check)
for i, f in enumerate(filelist):
for i in even_check:
#print(filelist[i])
os.remove(filelist[i])
# -
| PIV_PROCESSING_FINAL/1surface/deletion_even.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Advance Data Science Innovation - Assignment 1 Second Submission
#
# ### NBA Career Prediction - Predicting 5-Year Career Longevity for NBA Rookies
#
# **Group 1:** <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# **Pre-requisites:**
# - Create a github account (https://github.com/join)
# - Install git (https://git-scm.com/book/en/v2/Getting-Started-Installing-Git)
# - Install Docker (https://docs.docker.com/get-docker/)
#
# The steps are:
# 1. Launch Docker image
# 2. Load and prepare data
# 3. Model training and evaluation
# - neural net
# - logistic regression
# 4. Xgboost training and evaluation
# 5. Push changes
#
# ## 1. Load Python packages and libraries
# Only run this step once per Docker session - run then RESTART the Kernal & run this code again
# !pip install imbalanced-learn
# +
# required python libraries
import pandas as pd
import numpy as np
from joblib import dump
import seaborn as sns
import matplotlib.pyplot as plt
import time
# scikit-learn models and functions
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import mean_absolute_error as mae
from sklearn.metrics import accuracy_score
from sklearn.feature_selection import SelectFromModel
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import f_regression
from sklearn.feature_selection import mutual_info_regression
# Logistic Regression Models
from sklearn.preprocessing import StandardScaler, LabelEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
from sklearn.metrics import plot_confusion_matrix
import warnings
warnings.filterwarnings('ignore')
# -
# ## 2. Prepare data for model training
#
# **[2.1]** Import training & final test (kaggle) data
#
# **[2.2]** Copy data for transformation for modelling steps
# import training & final test data
df_train = pd.read_csv('../data/raw/train.csv')
df_test = pd.read_csv('../data/raw/test.csv')
# +
# Create a copy of df and save it into a variable called df_cleaned
df_cleaned = df_train.copy()
df_clean_test = df_test.copy()
# Drop columns 'Id'
drop_cols = ['Id']
# We need to drop the Id column as this is irrelevant for modelling
df_cleaned.drop(drop_cols, axis=1, inplace=True)
df_clean_test.drop(drop_cols, axis=1, inplace=True)
# Remove leading and trailing space from the column names
df_cleaned.columns = df_cleaned.columns.str.strip()
df_clean_test.columns = df_clean_test.columns.str.strip()
# Extract the column TARGET_5Yrs and save it into variable called target
target = df_cleaned.pop('TARGET_5Yrs')
print('df_cleaned row,cols ',df_cleaned.shape)
print('df_clean_test rows,cols',df_clean_test.shape,'\n')
# we will need labels later for plotting results
labels = df_cleaned.columns.tolist()
print(labels)
# -
# **[2.3]** Function to select required features only
#
# **[2.4]** Create first train/test data with ALL features remaining for Baseline testing & evaluation
# +
# only keep selected features / cols
def select_features(df_train, df_test, keep_cols=['']):
# take copies of the Cleaned data up to now - so we can rerun from here different feature selections
df_clean_new = df_train.copy()
df_clean_new_test = df_test.copy()
return df_clean_new[keep_cols], df_clean_new_test[keep_cols]
# ALL Features for baseline testing for training models
df_clean_R1, df_clean_R1_test = select_features(df_cleaned, df_clean_test,keep_cols=labels)
# check still the same number of rows & columns as original cleaned training data
print('df_clean_R1 row,cols ',df_clean_R1.shape)
# -
# ### Standardising, Resampling Imbalance Data
#
# **[2.5]** Create function to prepare training data for modelling
# - apply Sci-kit Learn `StandardScaler` as the data contains negative values
# - allow for resampling techniques to treat imbalanced target data `SMOTE`, `ADASYN` are the current techniques
#
# (i) the Synthetic Minority Oversampling Technique (SMOTE) Sampling <br>
# (ii) the Adaptive Synthetic (ADASYN) Sampling <br>
# https://imbalanced-learn.org/stable/over_sampling.html#smote-adasyn
# +
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.model_selection import train_test_split
from imblearn.over_sampling import SMOTE, ADASYN
# set defaults
random_state=100
scaler = StandardScaler()
# Prepare training data for modelling - Standardising, Resampling for imbalanced data
def Prep_Model_Data(df_clean, target, scaler, resample='', random_state=8):
# rescale eg: scaler = StandardScaler()
df_clean = scaler.fit_transform(df_clean)
# df_clean_test = scaler.fit_transform(df_clean_test)
#---------------------------------------------------------------------------------------
# Split randomly the dataset with random_state=8 into 2 different sets: data (80%) and test (20%)
X_data, X_test, y_data, y_test = train_test_split(df_clean, target, test_size=0.2, random_state=random_state)
# Split the remaining data (80%) randomly with random_state=8 into 2 different sets: training (80%) and validation (20%)
X_train, X_val, y_train, y_val = train_test_split(X_data, y_data, test_size=0.2, random_state=random_state)
#---------------------------------------------------------------------------------------
# RESAMPLE if indicated
# Original Training data
if resample=='NO RESAMPLE' or resample=='':
X_train_res, y_train_res = X_train, y_train
X_val_res, y_val_res = X_val, y_val
X_test_res, y_test_res = X_test, y_test
print('NO Resample - ALL Features')
# SMOTE - treat Imbalanced Target data after splitting Training & Test & Validation data
if resample=='SMOTE':
X_train_res, y_train_res = SMOTE(random_state=random_state).fit_resample(X_train, y_train)
X_val_res, y_val_res = SMOTE(random_state=random_state).fit_resample(X_val, y_val)
X_test_res, y_test_res = SMOTE(random_state=random_state).fit_resample(X_test, y_test)
# ADASYN - treat Imbalanced Target data after splitting Training & Test & Validation data
if resample=='ADASYN':
X_train_res, y_train_res = ADASYN(random_state=random_state).fit_resample(X_train, y_train)
X_val_res, y_val_res = ADASYN(random_state=random_state).fit_resample(X_val, y_val)
X_test_res, y_test_res = ADASYN(random_state=random_state).fit_resample(X_test, y_test)
return X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res
# -
# ## 3. Model Training and Evaluation
#
# **[3.1]** Function to perform **Classification** training and basic evaluation
# +
from sklearn.metrics import roc_auc_score, make_scorer
def Train_Classifier(X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res, classifier, features='all', resample='resample'):
t_start = time.process_time()
# fit model
classify = classifier.fit(X_train_res, y_train_res)
t_end = time.process_time()
t_diff = t_end - t_start
#--------------------------------------------------------------------------
# Compare accuracy of the given test data
# baseline target average
y_mode = y_train_res.mode()
if resample=='NO RESAMPLE' or resample=='':
y_base_res = np.full((len(y_train_res), 1), y_mode)
else:
y_base_res = np.full((len(y_train_res), 1), y_mode[0])
# for debugging during dev/test
if 1==2:
print('mode',y_train_res.mode(), '\ttype:',type(y_train_res.mode()))
print('len(X_train)',len(X_train_res))
print('len(y_train)',len(y_train_res))
print('y_base_res',y_base_res)
print(f"Compare accuracy between data sets - testing {features} features ")
print(f"Classifier - {classify}")
print(f"Imbalanced data fix - {resample}")
print("Baseline: ",accuracy_score(y_train_res, y_base_res))
print("Train data: ",classifier.score(X_train_res, y_train_res))
print("Validation: ",classifier.score(X_val_res, y_val_res))
#--------------------------------------------------------------------------
# final test (unseen) data AUC score
pred_prob = classifier.predict_proba(X_test_res)
auc_score = roc_auc_score(y_test_res, pred_prob[:,1])
print('Test data auc:',auc_score,'\n')
return classify, X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res
# -
# **[3.2]** Functions to evaluate model accuracy and performance metrics
#
# - AUC - area under the curve https://stats.stackexchange.com/questions/132777/what-does-auc-stand-for-and-what-is-it
# +
import matplotlib.pyplot as plt
from sklearn.metrics import roc_curve, roc_auc_score, plot_confusion_matrix
from sklearn.metrics import make_scorer
font_size=14
plt.rcParams['font.size'] = font_size
def Plot_Confusion_Matrix(classifier, X, y, title):
print(f'Confusion matrix - {title}')
plot_confusion_matrix(classifier, X, y, cmap=plt.cm.Blues, normalize='true')
plt.show();
def Model_Perform_Evaluate(classifier, X_test, y_test, show_roc, class_label):
# predict probabilities
pred_prob = classifier.predict_proba(X_test)
if show_roc==True:
# roc curve for models
fpr, tpr, thresh = roc_curve(y_test, pred_prob[:,1], pos_label=1)
Plot_ROC_Curve(pred_prob, fpr, tpr, class_label)
return pred_prob, class_label
def Plot_ROC_Curve(pred_prob, fpr, tpr, model_name):
plt.style.use('seaborn')
# roc curve for models
# fpr, tpr, thresh = roc_curve(y_test, pred_prob[:,1], pos_label=1)
# plot roc curves
plt.plot(fpr, tpr, linestyle='--',color='orange', label=model_name)
# plt.plot(p_fpr, p_tpr, linestyle='--', color='blue')
# title
plt.title(f'ROC curve - {model_name}', fontsize=font_size+2)
# x label
plt.xlabel('False Positive Rate', fontsize=font_size)
# y label
plt.ylabel('True Positive rate', fontsize=font_size)
plt.legend(loc='best', fontsize=(font_size-2))
plt.savefig('ROC',dpi=300)
plt.show();
# -
# **[3.3]** Save the model predictions on the Kaggle test data to CSV, then submit online to the Kaggle competition
#
# https://www.kaggle.com/c/uts-advdsi-22-02-nba-career-prediction/leaderboard
# +
#save the final "test" prediction probabilities for Kaggle
def Save_Kaggle_Results(classifier, scaler, df_clean_test, save_csv=False, csv_name=''):
df_final_test = scaler.fit_transform(df_clean_test)
y_final_preds = classifier.predict_proba(df_final_test)
# combine df_test Id column with prediction probabilities column (cover to dataframe first)
frames = [df_test.iloc[:,0], pd.DataFrame(y_final_preds[:,1])]
result = pd.concat(frames, axis=1)
result.columns = ['Id','tmp']
result['TARGET_5Yrs'] = [round(num, 2) for num in result['tmp']]
result.drop(['tmp'], axis=1, inplace=True)
# print(len(df_final_test),len(y_final_preds))
# print(result.shape)
#--------------------------------------------------------------------------
# Save the final predictions for submission to Kaggle
if save_csv==True:
result.to_csv('../data/processed/cazmayhem_{c}.csv'.format(c=csv_name), index=False)
print('kaggle results saved ../data/processed/cazmayhem_{c}.csv'.format(c=csv_name))
return csv_name
# -
# **[3.4]** Initialise Classification models & parameters
#
# 1. Neural Network MLPClassifier()
# 2. LogisticRegression()
#
# These 2 models performed the best in Test 1.
# +
from sklearn.neural_network import MLPClassifier
from sklearn.linear_model import LogisticRegression
# MLPClassifier parameters
activation='logistic'
solver='adam' # sgd, adam (default)
alpha=0.01 # 0.0001 default
max_iter=300
batch_size=100
random_state=100
# Classification Models defined - MLPClassifier()
clf_NN_MLP=MLPClassifier(activation=activation, solver=solver, alpha=alpha, batch_size=batch_size, random_state=random_state)
# Classification Models defined - LogisticRegression()
clf_LR=LogisticRegression(C=109.85411419875572, class_weight=None, dual=False,
fit_intercept=True, intercept_scaling=1, max_iter=100, n_jobs=None, penalty='l2',
tol=0.0001, verbose=0, warm_start=False, random_state=random_state)
# -
# #### [3.5] TEST 1 - Neural Net (MLPClassifier) ALL Features with SMOTE resampling
#
# Previous testing has been performed - these tests performed best for each model type
# +
# Test 1 - Neural Net (MLPClassifier) Resampling Imbalance Data - ALL Original Features
random_state = 100
features='ALL'
resample='SMOTE'
model_name=f'Neural Net (MLPClassifier) {resample}'
# labels = keep all features
top_feat=labels
classifier=clf_NN_MLP
# create train/test data from selected important features only
df_clean_mod, df_clean_mod_test = select_features(df_cleaned, df_clean_test, top_feat)
# Prepare data for model training
X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Prep_Model_Data(df_clean_mod, target, scaler, resample, random_state)
# Train the Classification model
classify, X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Train_Classifier(X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res,
classifier, features, resample)
# Confusion Matrix Training data
Plot_Confusion_Matrix(classify, X_train_res, y_train_res, 'training data')
# Confusion Matrix Testing data
Plot_Confusion_Matrix(classify, X_test_res, y_test_res, 'test data')
# Model predictions & roc_curve
pred_prob_1, model_name_1 = Model_Perform_Evaluate(classify, X_test_res, y_test_res, False, model_name)
# save results for Kaggle comp
Save_Kaggle_Results(classify, scaler, df_clean_mod_test, False, 'nn_mlp_smote')
# -
# #### [3.6] TEST 2 - Logistic Regression ALL Features with SMOTE resampling
# +
# Test 2 - Logistic Regression Resampling Imbalance Data - ALL Original Features
random_state = 100
features='ALL'
resample='SMOTE'
model_name=f'Logistic Regression {resample}'
csv_name='lr_smote'
# labels = keep all features
top_feat=labels
classifier=clf_LR
# intiailse training data for this test
df_clean_mod, df_clean_mod_test = select_features(df_cleaned, df_clean_test, top_feat)
# X_train, y_train, X_val, y_val, X_test, y_test = Prep_Model_Data(df_clean_R1, df_clean_R1_test, target, scaler, classifier, features, model_name, random_state)
X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Prep_Model_Data(df_clean_mod, target, scaler, resample, random_state)
# run the final Train_Classifier()
classify, X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Train_Classifier(X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res,
classifier, features, resample)
# Confusion Matrix Training data
Plot_Confusion_Matrix(classify, X_train_res, y_train_res, 'training data')
# Confusion Matrix Testing data
Plot_Confusion_Matrix(classify, X_test_res, y_test_res, 'test data')
# Plot ROC Curve
pred_prob_2, model_name_2 = Model_Perform_Evaluate(classify, X_test_res, y_test_res, False, model_name)
# save results for Kaggle comp
Save_Kaggle_Results(classify, scaler, df_clean_mod_test, False, 'lr_smote')
# -
# #### [3.7] TEST 3 - Neural Net (MLPClassifier) Reduced Features with SMOTE resampling
#
# - remove the %-age columns as they are a reflection of "made" / "attempts" for the various features
# +
# Test 3 - Neural Net (MLPClassifier) Resampling Imbalance Data - Reduced Features
random_state = 100
features='reduced'
resample='SMOTE'
model_name=f'Neural Net (MLPClassifier) {resample}'
csv_name='mlp_smote_feat_2'
# remove %-age columns
top_feat=['GP', 'MIN', 'PTS', 'FGM', 'FGA', '3P Made', '3PA', 'FTM', 'FTA', 'OREB', 'DREB', 'REB', 'AST', 'STL', 'BLK', 'TOV']
classifier=clf_NN_MLP
# intiailse training data for this test
df_clean_mod, df_clean_mod_test = select_features(df_cleaned, df_clean_test, top_feat)
# Prepare data for model training
X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Prep_Model_Data(df_clean_mod, target, scaler, resample, random_state)
# Train the Classification model
classify, X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Train_Classifier(X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res,
classifier, features, resample)
# Confusion Matrix Training data
Plot_Confusion_Matrix(classify, X_train_res, y_train_res, 'training data')
# Confusion Matrix Testing data
Plot_Confusion_Matrix(classify, X_test_res, y_test_res, 'test data')
# Model predictions & roc_curve
Model_Perform_Evaluate(classify, X_test_res, y_test_res, False, model_name)
# save results for Kaggle comp
Save_Kaggle_Results(classify, scaler, df_clean_mod_test, False, csv_name)
# -
# #### [3.8] TEST 4 - Logistic Regression Reduced Features with SMOTE resampling
#
# - remove the %-age columns as they are a reflection of "made" / "attempts" for the various features
# +
# Test 4 - Logistic Regression Resampling Imbalance Data - Reduced Features
random_state = 100
features='reduced'
resample='SMOTE'
model_name=f'Logistic Regression {resample}'
csv_name='lr_smote_feat_2'
# remove %-age columns
top_feat=['GP', 'MIN', 'PTS', 'FGM', 'FGA', '3P Made', '3PA', 'FTM', 'FTA', 'OREB', 'DREB', 'REB', 'AST', 'STL', 'BLK', 'TOV']
classifier=clf_LR
# intiailse training data for this test
df_clean_mod, df_clean_mod_test = select_features(df_cleaned, df_clean_test, top_feat)
# Prepare data for model training
X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Prep_Model_Data(df_clean_mod, target, scaler, resample, random_state)
# Train the Classification model
classify, X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Train_Classifier(X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res,
classifier, features, resample)
# Confusion Matrix Training data
Plot_Confusion_Matrix(classify, X_train_res, y_train_res, 'training data')
# Confusion Matrix Testing data
Plot_Confusion_Matrix(classify, X_test_res, y_test_res, 'test data')
# Model predictions & roc_curve
Model_Perform_Evaluate(classify, X_test_res, y_test_res, False, model_name)
# save results for Kaggle comp
Save_Kaggle_Results(classify, scaler, df_clean_mod_test, False, csv_name)
# -
# ## 4. Perform Gradient Boosting model training
#
# **[4.1]** `Autoreload` extensions to refresh functions defined externally
#
# **[4.2]** Reload `Autoreload` extension if already loaded
#
# **[4.3]** Function to perform **Train_XGBoost** training and basic evaluation
# %load_ext autoreload
# %autoreload 2
# %reload_ext autoreload
# +
# Import the xgboost package as xgb
import xgboost as xgb
from src.models.performance import print_class_perf
def Train_XGBoost(xgboost, X_train, y_train, X_val, y_val, X_test, y_test):
# Fit the model with the prepared data
xgboost.fit(X_train, y_train)
print('\n',xgboost.fit(X_train, y_train),'\n')
# Import dump from joblib and save the fitted model into the folder models as a file called xgboost_default
from joblib import dump
dump(xgboost, '../models/xgboost_default.joblib')
# Save the predictions from this model for the training and validation sets into 2 variables called y_train_preds and y_val_preds
y_train_preds = xgboost.predict(X_train)
y_val_preds = xgboost.predict(X_val)
y_test_preds = xgboost.predict(X_test)
# prediction probabilities for AUC
y_train_pred_prob = xgboost.predict_proba(X_train)
y_val_pred_prob = xgboost.predict_proba(X_val)
y_test_pred_prob = xgboost.predict_proba(X_test)
print_class_perf(y_preds=y_train_preds, y_actuals=y_train, y_pred_prob=y_train_pred_prob[:,1], set_name='Training', average='weighted')
print_class_perf(y_preds=y_val_preds, y_actuals=y_val, y_pred_prob=y_val_pred_prob[:,1], set_name='Validation', average='weighted')
print_class_perf(y_preds=y_test_preds, y_actuals=y_test, y_pred_prob=y_test_pred_prob[:,1], set_name='Test', average='weighted')
return xgboost, xgboost
# -
# **[4.4]** Function to identify **feature importance** for xgboost trained data
#
# - return the top N features by importance value
def xgb_feature_important(clf_xgb, df_clean, max_num_features=5):
# Extract the feature importance from the trained xgboost model
feat_imp = clf_xgb.get_booster().get_score(importance_type="gain")
# Using a dictionary comprehension, create a dictionary called mapper with keys
# in the format of f0 (0 is the index of the first column) and values as column names
mapper = {'f{0}'.format(i): v for i, v in enumerate(df_clean.columns)}
# Using a dictionary comprehension, create a dictionary called mapping with keys
# as the column names from mapper and values as the corresponding feature importance
mapping = {mapper[k]: v for k, v in feat_imp.items()}
# Plot the partial dependence plot using plot_importance for the top 5 features
xgb.plot_importance(mapping, max_num_features=max_num_features)
# extract top 5 features - but this will be an array
n_feat = max_num_features
top_feat = pd.DataFrame(sorted(mapping.items(), key=lambda x: x[1], reverse=True)).iloc[:n_feat,:1].values
# Convert elements in a numpy array to string
top_feat = [",".join(item) for item in top_feat.astype(str)]
print('\n')
print('Top features list',top_feat)
return top_feat
# **[4.5]** **TEST 5** - XGBoost training with default parameters
# +
# Test 5 - XGBoost Resampling Imbalance Data - ALL Original Features
random_state = 100
features='ALL'
resample='NO RESAMPLE'
model_name=f'XGBoost {resample}'
top_feat=labels
# intiailse training data for this test
df_clean_mod, df_clean_mod_test = select_features(df_cleaned, df_clean_test, top_feat)
# instantiate the XGBClassifier class into a variable called xgboost1
xgboost1 = xgb.XGBClassifier(learning_rate=0.01, random_state=random_state)
X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Prep_Model_Data(df_clean_mod, target, scaler, resample, random_state)
Train_XGBoost(xgboost1, X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res)
# -
# **[4.6]** Perform hyperparameter tuning with Hyperopt
#
# - input parameters: hyperparameter seacrh space (`space`)
# - logics: train a xgboost model with the search space and calculate the average accuracy score for cross validation with 10 folds
# - output parameters: dictionary with the loss score and STATUS_OK
# +
# Import Trials, STATUS_OK, tpe, hp, fmin from hyperopt package
from hyperopt import Trials, STATUS_OK, tpe, hp, fmin
# Define the search space for xgboost hyperparameters
space = {
'max_depth' : hp.choice('max_depth', range(5, 20, 1)),
'learning_rate' : hp.quniform('learning_rate', 0.01, 0.5, 0.05),
'min_child_weight' : hp.quniform('min_child_weight', 1, 10, 1),
'subsample' : hp.quniform('subsample', 0.1, 1, 0.05),
'colsample_bytree' : hp.quniform('colsample_bytree', 0.1, 1.0, 0.05)
}
# Define a function called objective
def objective(space):
from sklearn.model_selection import cross_val_score
xgboost = xgb.XGBClassifier(
max_depth = int(space['max_depth']),
learning_rate = space['learning_rate'],
min_child_weight = space['min_child_weight'],
subsample = space['subsample'],
colsample_bytree = space['colsample_bytree'],
random_state=random_state
)
acc = cross_val_score(xgboost, X_train_res, y_train_res, cv=10, scoring="accuracy").mean()
return{'loss': 1-acc, 'status': STATUS_OK }
# -
# **[4.7]** Launch Hyperopt search and save the result in a variable called `best`
# +
# Launch Hyperopt search and save the result in a variable called `best`
best = fmin(
fn=objective,
space=space,
algo=tpe.suggest,
max_evals=10
)
# Print the best set of hyperparameters
print("\nBest: ", best)
# -
# **[4.7]** Instantiate a XGBClassifier with best set of hyperparameters
#
# **[4.8]** Fit the model with the prepared data
# +
# Instantiate a XGBClassifier with best set of hyperparameters
xgboost2 = xgb.XGBClassifier(
max_depth = best['max_depth'],
learning_rate = best['learning_rate'],
min_child_weight = best['min_child_weight'],
subsample = best['subsample'],
colsample_bytree = best['colsample_bytree'],
random_state=random_state
)
# Fit the model with the prepared data
xgboost2.fit(X_train_res, y_train_res)
# +
# Display the accuracy and f1 scores of this baseline model on the training and validation sets
print_class_perf(y_preds=xgboost2.predict(X_train_res), y_actuals=y_train_res, y_pred_prob=xgboost2.predict_proba(X_train_res)[:,1], set_name='Training', average='weighted')
print_class_perf(y_preds=xgboost2.predict(X_val_res), y_actuals=y_val_res, y_pred_prob=xgboost2.predict_proba(X_val_res)[:,1], set_name='Validation', average='weighted')
print_class_perf(y_preds=xgboost2.predict(X_test_res), y_actuals=y_test_res, y_pred_prob=xgboost2.predict_proba(X_test_res)[:,1], set_name='Test', average='weighted')
# Test Accuracy Score
# Test: 0.6852169333512617
# -
# **[4.9]** **TEST 6** Perform XGBoost training using SMOTE resampling for imbalanced target data
#
# - using best performing parameters from Hyperbot
# +
# Test 6 - XGBoost Hyperparameter Tuned - Resampling Imbalance Data - ALL Original Features
features='ALL'
resample='SMOTE'
scaler = StandardScaler()
max_num_features=6
# labels = ALL original features
top_feat=labels
# select required features for this tesm
df_clean_mod, df_clean_mod_test = select_features(df_cleaned, df_clean_test, top_feat)
# intiailse training data for this test
X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Prep_Model_Data(df_clean_mod, target, scaler, resample, random_state)
# use xgboost2 with best parameters from Hyperbot
classify, clf_xgb_1 = Train_XGBoost(xgboost2, X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res)
# New Test AUC Score
# Test: 0.9286273886499112
# Show XGB feature importance
top_feat = xgb_feature_important(classify, df_clean_mod, max_num_features)
# -
# **[4.10]** **TEST 7** Repeat previous step with only top N features identified as most important
#
# - still using xgboost2
# +
# TEST 7 XGBoost repeat previous step with only top N features identified as most important
features='Top 5 Important'
resample='SMOTE'
model_name=f'XGBoost {resample}'
csv_name='xgboost_smote_top_feat'
# features identified previous step
top_feat=top_feat
scaler = StandardScaler()
# select required features for this tesm
df_clean_mod, df_clean_mod_test = select_features(df_cleaned, df_clean_test, top_feat)
# intiailse training data for this test
X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res = Prep_Model_Data(df_clean_mod, target, scaler, resample, random_state)
# quick check we have the same features selected in training data
print('df_clean_mod rows,copls',df_clean_mod.shape)
print('X_train_res rows,copls ',X_train_res.shape)
# Train the model using xgboost2 after hyperopt
classify, clf_xgb_2 = Train_XGBoost(xgboost2, X_train_res, y_train_res, X_val_res, y_val_res, X_test_res, y_test_res)
# Confusion Matrix Training data
Plot_Confusion_Matrix(classify, X_train_res, y_train_res, 'training data')
# Confusion Matrix Testing data
Plot_Confusion_Matrix(classify, X_test_res, y_test_res, 'test data')
# Plot ROC Curve - save pret_prob & model_name for combined ROC graph
pred_prob_3, model_name_3 = Model_Perform_Evaluate(classify, X_test_res, y_test_res, True, model_name)
# save results for Kaggle comp - XGBoost with hyperopt using SMOTE for imbalanced target - correct model this time
best_mod_xgb = Save_Kaggle_Results(classify, scaler, df_clean_mod_test, True, csv_name)
# Show XGB feature importance - not required this step
# xgb_feature_important(classify, df_clean_mod, max_num_features=5)
# Test AUC Score
# Test: 0.9336216847478108
# +
def Save_Kaggle_Results_test(classifier, scaler, df_clean_test, save_csv=False, csv_name=''):
df_final_test = scaler.fit_transform(df_clean_test)
y_final_preds = classifier.predict_proba(df_final_test)
# combine df_test Id column with prediction probabilities column (cover to dataframe first)
frames = [df_test.iloc[:,0], pd.DataFrame(y_final_preds[:,1])]
result = pd.concat(frames, axis=1)
result.columns = ['Id','tmp']
result['TARGET_5Yrs'] = [round(num, 2) for num in result['tmp']]
result.drop(['tmp'], axis=1, inplace=True)
# print(len(df_final_test),len(y_final_preds))
# print(result.shape)
#--------------------------------------------------------------------------
# Save the final predictions for submission to Kaggle
if save_csv==True:
result.to_csv('../data/processed/cazmayhem_{c}.csv'.format(c=csv_name), index=False)
print('kaggle results saved ../data/processed/cazmayhem_{c}.csv'.format(c=csv_name))
return csv_name
df_clean_mod_test.shape
# +
#=========================================================================
# COMBINE ROC CURVE GRAPH
#=========================================================================
# roc curve for models
fpr1, tpr1, thresh1 = roc_curve(y_test_res, pred_prob_1[:,1], pos_label=1)
fpr2, tpr2, thresh2 = roc_curve(y_test_res, pred_prob_2[:,1], pos_label=1)
fpr3, tpr3, thresh3 = roc_curve(y_test_res, pred_prob_3[:,1], pos_label=1)
# roc curve for tpr = fpr
random_probs = [0 for i in range(len(y_test_res))]
p_fpr, p_tpr, _ = roc_curve(y_test_res, random_probs, pos_label=1)
# chart size
plt.rcParams["figure.figsize"] = (9,7)
# plot roc curves
plt.plot(fpr1, tpr1, linestyle='--',color='orange', label=model_name_1)
plt.plot(fpr2, tpr2, linestyle='--',color='green', label=model_name_2)
plt.plot(fpr3, tpr3, linestyle='--',color='cyan', label=model_name_3)
plt.plot(p_fpr, p_tpr, linestyle='--', color='blue', label='TPR = FPR')
# title
plt.title(f'ROC curve combined models', fontsize=font_size+2)
# x label
plt.xlabel('False Positive Rate', fontsize=font_size)
# y label
plt.ylabel('True Positive rate', fontsize=font_size)
plt.legend(loc='best', fontsize=(font_size-2))
plt.savefig('ROC',dpi=300)
plt.show();
# -
# **[4.11]** Save the best fitted model into the folder 'models'
#
# +
#--------------------------------------------------------------------------
# Save the best fitted model into the folder 'models', named for each classifier
dump(clf_xgb_2, '../models/{c}.joblib'.format(c=best_mod_xgb))
# -
# ## 5. Push changes to GitHub
"""
# Add you changes to git staging area
# Create the snapshot of your repository and add a description
# Push your snapshot to Github
git add .
git commit -m "commit 4 xgboost hyperopt"
git push https://*********@github.com/CazMayhem/adv_dsi_AT1.git
# Check out to the master branch
# Pull the latest updates
git checkout master
git pull https://*********@github.com/CazMayhem/adv_dsi_AT1.git
# Check out to the data_prep branch
# Merge the master branch and push your changes
git checkout xgboost_hyperopt
git merge master
git push https://*********@github.com/CazMayhem/adv_dsi_AT1.git
"""
| notebooks/paipa_carol-90014679-week2_xgboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Training SentimentDetector Model in Italian language
# ### A brief explaination about `SentimentDetector` annotator in Spark NLP:
#
# Scores a sentence for a sentiment<br>
# **Type:** sentiment<br>
# **Requires:** Document, Token<br>
#
# **Functions:**<br>
# * setSentimentCol(colname): Column with sentiment analysis row's result for training. If not set, external sources need to be set instead.<br>
# * setPositiveSource(path, tokenPattern, readAs, options): Path to file or folder with positive sentiment text, with tokenPattern the regex pattern to match tokens in source. readAs either LINE_BY_LINE or as SPARK_DATASET. If latter is set, options is passed to reader <br>
# * setNegativeSource(path, tokenPattern, readAs, options): Path to file or folder with positive sentiment text, with tokenPattern the regex pattern to match tokens in source. readAs either LINE_BY_LINE or as SPARK_DATASET. If latter is set, options is passed to reader <br>
# * setPruneCorpus(true): when training on small data you may want to disable this to not cut off unfrequent words
# <br>
#
# **Input:** File or folder of text files of positive and negative data<br>
# **Example:**<br>
# ```python
# sentiment_detector = SentimentDetector() \
# .setInputCols(["lemma", "sentence"]) \
# .setOutputCol("sentiment")
# ```
# Let's import required libraries including `SQL` and `ML` from Spark and some annotators from Spark NLP
#Spark ML and SQL
from pyspark.ml import Pipeline, PipelineModel
from pyspark.sql.functions import array_contains
from pyspark.sql import SparkSession
from pyspark.sql.types import StructType, StructField, IntegerType, StringType
#Spark NLP
import sparknlp
from sparknlp.annotator import *
from sparknlp.common import RegexRule
from sparknlp.base import DocumentAssembler, Finisher
# ### Let's create a Spark Session for our app
# +
spark = sparknlp.start()
print("Spark NLP version: ", sparknlp.version())
print("Apache Spark version: ", spark.version)
# -
# ! wget -N https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/resources/it/lemma/dxc.technology/lemma_italian.txt -P /tmp
# ! wget -N https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/resources/it/sentiment/dxc.technology/sentiment_italian.txt -P /tmp
# ### Now we are going to create a Spark NLP Pipeline by using Spark ML Pipeline natively
# +
document_assembler = DocumentAssembler() \
.setInputCol("text")
sentence_detector = SentenceDetector() \
.setInputCols(["document"]) \
.setOutputCol("sentence")
tokenizer = Tokenizer() \
.setInputCols(["sentence"]) \
.setOutputCol("token")
normalizer = Normalizer() \
.setInputCols(["token"]) \
.setOutputCol("normal")
lemmatizer = Lemmatizer() \
.setInputCols(["normal"]) \
.setOutputCol("lemma") \
.setDictionary(
path = "/tmp/lemma_italian.txt",
read_as = "LINE_BY_LINE",
key_delimiter = "\\s+",
value_delimiter = "->"
)
sentiment_detector = SentimentDetector() \
.setInputCols(["lemma", "sentence"]) \
.setOutputCol("sentiment_score") \
.setDictionary(
path = "/tmp/sentiment_italian.txt",
read_as = "LINE_BY_LINE",
delimiter = ","
)
pipeline = Pipeline(stages=[document_assembler, sentence_detector, tokenizer, normalizer, lemmatizer, sentiment_detector])
# -
# Now that we have our Spark NLP Pipeline, we can go ahead with training it by using `fit()`. Since we are using an external dataset to train our `Lemmatizer` and `SentimentDetector` models we don't need to pass any DataFrame with real data. We are going to create an empty DataFrame to just trigger the training.
# Let's see how good our model does when it comes to prediction. We are going to create a DataFrame with Italian text for testing purposes and use `transform()` to predict.
# +
# Let's create a DataFrame with Italian text for testing our Spark NLP Pipeline
dfTest = spark.createDataFrame(["Finchè non avevo la linea ADSL di fastweb potevo entrare nel router e configurare quelle pochissime cose configurabili (es. nome dei device), da ieri che ho avuto la linea niente è più configurabile...",
"L'uomo è insoddisfatto del prodotto.",
"La coppia contenta si abbraccia sulla spiaggia."], StringType()).toDF("text")
# Of course you can select multiple columns at the same time however, this way we see each annotator without truncating their results
pipeline.fit(dfTest).transform(dfTest).select("token.result").show(truncate=False)
pipeline.fit(dfTest).transform(dfTest).select("normal.result").show(truncate=False)
pipeline.fit(dfTest).transform(dfTest).select("lemma.result").show(truncate=False)
pipeline.fit(dfTest).transform(dfTest).select("sentiment_score").show(truncate=False)
# Print the schema of the Pipeline
pipeline.fit(dfTest).transform(dfTest).printSchema()
# -
# ### Credits
# We would like to thank `DXC.Technology` for sharing their Italian datasets and models with Spark NLP community. The datasets are used to train `Lemmatizer` and `SentimentDetector` Models.
| jupyter/training/italian/Train-SentimentDetector-Italian.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python для анализа данных
#
# *<NAME>, НИУ ВШЭ*
#
# дополнения: *<NAME>, НИУ ВШЭ*
# Посмотрим на другие примеры использования `selenium`.
#
# **Пример.** Зайдем на сайт книжного магазина и найдем все книги про Python. Загрузим библиотеку, веб-драйвер и откроем страницу в браузере через Python.
# +
from selenium import webdriver
from webdriver_manager.chrome import ChromeDriverManager
br = webdriver.Chrome(ChromeDriverManager().install())
# открываем страницу в Chrome в автоматическом режиме
br.get("http://www.biblio-globus.ru/")
# -
# Найдем с помощью CSS Selector'а (*SelectorGadget*) поле для ввода названия книги или автора.
field = br.find_element_by_css_selector("input")
# Сохраним запрос:
author = "Python" # переменная author - условность
# Введем запрос в поле для поиска (`.send_keys`) и подождем чуть-чуть:
field.send_keys(author)
br.implicitly_wait(2) # подождем пару секунд
# Теперь найдем кнопку для поиска (значок *лупа* рядом со строкой поиска) через CSS Selector:
# #search_submit
submit = br.find_element_by_css_selector("#search_submit")
# Кликнем на нее:
submit.click()
# Сохраним первую страницу с результатами в переменную `page1`.
page1 = br.page_source
page1
# Теперь обработаем эту страницу через `BeautifulSoup`:
from bs4 import BeautifulSoup
soup1 = BeautifulSoup(page1, 'lxml')
# Найдем все названия книг на этой странице. По исходному коду можно увидеть, что они имеют тэг `a` с атрибутом `class`, равным `name`:
soup1.find_all('a', {'class':'name'})
# С помощью списковых включений выберем из ссылок с тэгом `<a>` текст (так мы уже делали, и не раз).
a = []
for tag in soup1.find_all('a', {'class':'name'}):
a.append(tag.text)
books1 = [b.text for b in soup1.find_all('a', {'class':'name'})]
books1
# Теперь аналогичным образом сгрузим информацию об авторах:
authors1 = [a.text for a in soup1.find_all('div', {'class': 'author'})]
# Сгрузим расположение:
place1 = [p.text for p in soup1.find_all('div', {'class':'placement'})]
place1
# И, конечно, цену:
price1 = [p.text for p in soup1.find_all('div',
{'class':'title_data price'})]
price1
# Осталось пройтись по всем страницам, которые были выданы в результате поиска. Для примера перейдем на страницу 2 и на этом остановимся.
next_p = br.find_element_by_css_selector('.next_page')
next_p.click()
# Проделаем то же самое, что и с первой страницей. По-хорошему нужно написать функцию, которая будет искать на странице названия книг, их расположение и цену. Но оставим это в качестве задания читателю :)
page2 = br.page_source
soup2 = BeautifulSoup(page2, 'lxml')
books2 = [b.text for b in soup2.find_all('a', {'class':'name'})]
author2 = [a.text for a in soup2.find_all('div', {'class': 'author'})]
place2 = [p.text for p in soup2.find_all('div', {'class':'placement'})]
price2 = [p.text for p in soup2.find_all('div', {'class':'title_data price'})]
# Расширим списки результатов с первой страницы данными, полученными со второй страницы, используя метод `.extend()`.
books1.extend(books2)
authors1.extend(books2)
place1.extend(place2)
price1.extend(price2)
# Осталось импортировать библиотеку `pandas` и создать датафрейм.
import pandas as pd
# Для разнообразия создадим датафрейм не из списка списков, а из словаря. Ключами словаря будут названия столбцов в таблице, а значениями – списки с сохраненной информацией (названия книг, цены и проч.).
df = pd.DataFrame({'book': books1, 'author': authors1,
'placement': place1, 'price': price1})
df.head()
# Давайте приведем столбец с ценой к числовому типу. Уберем слова *Цена* и *руб*, а потом сконвертируем строки в числа с плавающей точкой. Напишем функцию `get_price()`,
float(df['price'][0].split()[1].replace(',', '.'))
def get_price(price):
book_price = price.split(' ')[1] # разобьем строку по пробелу и возьмем второй элемент
book_price = book_price.replace(',', '.') # заменим запятую на точку
price_num = float(book_price) # сконвертируем в float
return price_num
import re
def preis(x):
return float('.'.join(re.findall(r'\d+',x)))
# проверка
get_price(df.price[0])
preis(df.price[0])
# Всё отлично работает! Применим функцию к столбцу *price* и создадим новый столбец *nprice*.
df['nprice'] = df.price.apply(preis)
df.head()
df.shape
df.nprice.hist();
# Теперь можем расположить книги по цене в порядке возрастания:
df.sort_values('nprice', ascending=False)
# И сохраним всю таблицу в csv-файл:
df.to_csv("books.csv", index=False)
# ## Задание
#
# * http://www.biblio-globus.ru/
# * Нажать на кнопку "Книги"
# * Выбрать любую категорию
# * Собрать книги с 2 страниц
from bs4 import BeautifulSoup
from webdriver_manager.chrome import ChromeDriverManager
driver = webdriver.Chrome(ChromeDriverManager().install())
# +
driver.get("http://www.biblio-globus.ru/")
driver.find_element_by_xpath('//*[@id="TableMRight"]/tbody/tr/td/table/tbody/tr[2]/td[2]/a').click()
driver.find_element_by_xpath('/html/body/table/tbody/tr[2]/td[2]/div/div[2]/div[1]/div/ul/li/ul/li[3]/ul/li[5]/a').click()
page = driver.page_source
# +
soup = BeautifulSoup(page, 'html')
books_on_page = soup.find_all('div', {'class':'details_1'})
books = []
authors = [a.text for a in books_on_page.find_all('div', {'class':'author'})[:2]]
# -
len(books_on_page)
| week5/2020_DPO_10_3_selenium-books.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Falta fazer
# ## Relatório
# ## Documentação
# import das bibliotecas
import os
import pandas as pd
import random
import time
from greedy_filter import *
from math import inf
import numpy as np
import matplotlib.pyplot as plt
from sklearn import linear_model
# definicao das constantes
PATH_EXCEL = os.getcwd() + '/excel_files/'
# Estruturas que irao armazenar os dataframes com o histórico de preços das ações.
# 1. dict_excels: dicionario onde a chave é o nome da ação e o valor da chave é o dataframe
# 2. filenames: lista com o nome dos arquivos
# 3. excels: lista com os dataframes
dict_excels = {}
filenames = []
excels = []
for filename in os.listdir(PATH_EXCEL):
filenames.append(filename[:filename.find('.')])
excels.append(pd.read_excel(PATH_EXCEL + filename).sort_values(by=['Exchange Date']).reset_index(drop=True))
dict_excels[filename[:filename.find('.')]] = (pd.read_excel(PATH_EXCEL + filename).sort_values(by=['Exchange Date']).reset_index(drop=True))
filenames[6]
# ## Modelagem do problema
# ### Algoritmo Genético 1 - Escolha dos Parâmetros das Regras de Filtro
# 1. Gene: cada um dos parâmetros do filtro
# 1. x - porcentagem de variação: porcentagem acima/abaixo da ultima subida ou queda
# 1. varia de 0 a 1
# 2. varia de 0.01 a 0.10
# 2. h - hold days: após um sinal de compra/venda esperamos por h dias
# 1. varia de 1 até o número de dias entre a data atual e a data onde acabam as informacoes
# 2. varia de 1 até 30
# 3. d - delay days: após receber um sinal ignora-se os próximos d dias
# 1. varia de 1 até o número de dias entre a data atual e a data onde acabam as informacoes
# 2. varia de 1 até 30
# 4. p - previous days: olha-se p dias para tomar uma decisão
# 1. varia de 0 até o número de dias transcorridos até o momento
# 2. varia de 30 até o número máximo de dias
# 2. Cromossomo: conjunto de parâmetros
# 1. Estrutura usada: lista com os parâmetros → [x, h, d, p]
# 3. População: conjunto de cromossomos
# 1. Tamanho da População:
# 4. Mutação:
# 1. taxa de mutação:
# 5. Crossover:
# 1. taxa de crossover:
# 6. Critério de Parada:
# 7. Seleção:
# 1. Fitness: cálculo do lucro obtido por cada cromossomo
# 2. Técnica de Seleção:
# 1. Técnica 1:
# 2. Técnica 2:
# Funções para realizar o crossover entre dois cromossomos
# +
def crossover(chromosome1, chromosome2):
"""
Funcao para realizar o crossover entre dois cromossomos
:param: chromosome1 - cromossomo pai
:param: chromosome2 - cromossomo mae
:return: new_chromosome1 - primeiro filho gerado no crossover
:return: new_chromosome2 - segundo filho gerado no crossover
"""
# escolhe um gene aleatorio para realizar o crossover
rand = random.randint(1,len(chromosome1) - 1)
# gera os dois novos cromossomos
new_chromosome1 = chromosome1[:rand] + chromosome2[rand:]
new_chromosome2 = chromosome2[:rand] + chromosome1[rand:]
return new_chromosome1, new_chromosome2
def crossover2(chromosome1, chromosome2):
"""
Funcao para realizar o crossover entre dois cromossomos com 2 pontos de corte
:param: chromosome1 - cromossomo pai
:param: chromosome2 - cromossomo mae
:return: new_chromosome1 - primeiro filho gerado no crossover
:return: new_chromosome2 - segundo filho gerado no crossover
"""
# escolhe um gene aleatorio para realizar o crossover
rand = random.randint(1,len(chromosome1) - 2)
rand2 = random.randint(rand+1, len(chromosome1) - 1)
# gera os dois novos cromossomos
new_chromosome1 = chromosome1[:rand] + chromosome2[rand:rand2] + chromosome1[rand2:]
new_chromosome2 = chromosome2[:rand] + chromosome1[rand:rand2] + chromosome2[rand2:]
return new_chromosome1, new_chromosome2
# -
# Funções para realizar a mutação de um cromossomo
# +
def mutation(chromossome):
"""
Funcao para realizar a mutacao de um dado cromossomo
:param: chromossome - cromossomo que ira passar pelo processo de mutacao
:param: today_index - indice da tabela referente a data atual da acao
:param: last_day_index - indice da tabela referente a ultima data da qual se tem informacao da acao
:return: new_chromossome - novo cromossomo apos o processo de mutacao
"""
# faz a copia do cromossomo original
new_chromossome = chromossome
# escolhe, aleatoriamente, um gene para ser alterado
gene_initial_position = random.randint(0,len(chromossome) - 1)
gene_final_position = random.randint(gene_initial_position, len(chromossome) - 1)
# modifica o conjunto de genes escolhidos seguindo, como unica regra, os valores que eles podem assumir
for i in range(gene_initial_position, gene_final_position + 1):
if i == 0:
new_chromossome[0] = random.uniform(0.01,0.10)
elif i == 1:
new_chromossome[1] = random.randint(1,30)
elif i == 2:
new_chromossome[2] = random.randint(1,30)
elif i == 3:
new_chromossome[3] = random.randint(30,500)
else:
raise Exception('Gene inexistente no cromossomo!')
return new_chromossome
def mutation_v2(chromossome):
"""
Funcao para realizar a mutacao de um dado cromossomo
:param: chromossome - cromossomo que ira passar pelo processo de mutacao
:return: new_chromossome - novo cromossomo apos o processo de mutacao
"""
# faz a copia do cromossomo original
new_chromossome = chromossome
# escolhe, aleatoriamente, um gene para ser alterado
gene_position = random.randint(0,len(chromossome) - 1)
# modifica o gene escolhido seguindo, como unica regra, os valores que ele pode assumir
if gene_position == 0:
new_chromossome[0] = random.uniform(0.01,0.10)
elif gene_position == 1:
new_chromossome[1] = random.randint(1,30)
elif gene_position == 2:
new_chromossome[2] = random.randint(1,30)
elif gene_position == 3:
new_chromossome[3] = random.randint(30,500)
else:
raise Exception('Gene inexistente no cromossomo!')
return new_chromossome
# -
# Função para realizar a inicialização randômica de uma população
def create_population(population_size):
"""
Funcao para criar uma populacao randomica de cromossomos
:param: population_size - tamanho da populacao que sera criada
:return: population - nova populacao
"""
population = []
for i in range(0, population_size):
population.append([random.uniform(0,0.1), random.randint(1,30), random.randint(1,30), random.randint(1,500)])
return population
def fitness(np_array, chromossomes, budget):
"""
Funcao para calcular o lucro de cada um dos cromossomos de uma dada populacao
:param: np_array - conjunto de dados de determinada acao
:param: chromossomes - lista de cromossomos que sera avaliada
:param: budget - dinheiro inicial do problema
:return: fit_chromossomes - matriz com os cromossomos e o lucro(em porcentagem) obtido por eles
"""
fit_chromossomes = []
for chromossome in chromossomes:
money = greedy_filter_rule(np_array, chromossome, budget)
fit_chromossomes.append([chromossome, (money-budget)/budget])
return fit_chromossomes
def selection(stock_value, list_chromossomes, budget, cut_size):
"""
:param: stock_value -
:param: list_chromossomes -
:param: budget -
:param: cut_size -
:return: new_generation -
:return: fitness_array -
"""
fitness_array = fitness(stock_value, list_chromossomes, budget)
fitness_array.sort(key=lambda x: x[1], reverse = True)
new_generation = []
for i in range (0,cut_size):
new_generation.append(fitness_array[i][0])
return new_generation, fitness_array[:cut_size]
def roulette_selection(stock_value, list_chromossomes, budget, cut_size):
"""
:param: stock_value -
:param: list_chromossomes -
:param: budget -
:param: cut_size -
:return: new_population -
:return: cut_size -
"""
fitness_array = fitness(stock_value, list_chromossomes, budget)
# fitness_array = linear_normalization(fitness_array)
adds_skills = 0
for fit in fitness_array:
adds_skills = adds_skills + fit[1]
new_population = []
for i in range(cut_size):
r = random.uniform(0, adds_skills)
temp_soma = 0
for fit in fitness_array:
temp_soma = temp_soma + fit[1]
if temp_soma >= r:
new_population.append(fit[0])
break
return new_population, fitness(stock_value, new_population, budget)
def stop_criterion(old_population, new_population, limit_to_converge):
"""
:param: old_population - populacao ao iniciar a iteracao
:param: new_population - populacao ao fim da iteracao
:param: limit_to_converge - limiar abaixo do qual iremos considerar que ambas as pop convergem
:return: True se for para parar, False c.c.
"""
soma_old = 0
for x in old_population:
soma_old = soma_old + x[1]
soma_new = 0
for x in new_population:
soma_new = soma_new + x[1]
media_old = soma_old / len(old_population)
media_new = soma_new / len(new_population)
if abs(media_new - media_old) < limit_to_converge:
return True
else:
return False
def generate_children(old_generation, crossover_function, crossover_rate):
"""
:param: old_generation -
:param: crossover_function -
:param: crossover_rate -
:return: children -
"""
size_generation = len(old_generation)
number_to_crossover = int(size_generation * crossover_rate)
random.shuffle(old_generation)
children = []
for i in range (number_to_crossover):
for j in range (i+1, number_to_crossover):
new_chromossome1, new_chromossome2 = crossover_function(old_generation[i], old_generation[j])
children.append(new_chromossome1)
children.append(new_chromossome2)
return children
def mutation_chromossome(chromossomes, mutation_function, mutation_rate):
"""
:param: chromossomes -
:param: mutation_function -
:param: mutation_rate -
:return: chromossomes -
"""
number_chromossomes_to_mutate = int(len(chromossomes) * mutation_rate)
random.shuffle(chromossomes)
for i in range (0,number_chromossomes_to_mutate):
chromossomes[i] = mutation_function(chromossomes[i])
return chromossomes
def evolutionary_strategy1(stock_values, first_population, budget, crossover_function, delta, mutation_rate, crossover_rate, min_iteration_converge):
"""
:param: stock_values -
:param: first_population -
:param: budget -
:param: crossover_function -
:param: delta -
:return: old_population -
"""
flag = False
iteration = 0
old_population = first_population
while (not flag):
fitness_old_population = fitness(stock_values, old_population, budget)
children = generate_children(first_population, crossover_function, crossover_rate)
parents_and_children = old_population + children
chromossomes_parents_children_mutated = mutation_chromossome(parents_and_children, mutation_v2, mutation_rate)
new_population, fitness_new_population = selection(stock_values, chromossomes_parents_children_mutated, budget, len(old_population))
flag = (stop_criterion(fitness_old_population, fitness_new_population, delta) and iteration > min_iteration_converge)
iteration = iteration + 1
old_population = new_population
return old_population
def linear_normalization(fitness_population, increment=20):
pop_size = len(fitness_population)
fitness_population = sorted(fitness_population, key=lambda x: x[1], reverse = False)
min_value = 1
max_value = 201
normalized_fitness = []
for i in range(pop_size):
temp_fit = min_value + ((max_value - min_value)/(pop_size - 1)) * (i)
normalized_fitness.append([fitness_population[i][0],temp_fit])
return normalized_fitness
# +
def evolutionary_strategy2(stock_values, first_population, budget, crossover_function, mutation_func,delta, mutation_rate, crossover_rate, min_iteration_converge):
"""
:param: stock_values -
:param: first_population -
:param: budget -
:param: crossover_function -
:param: mutation_func -
:param: delta -
:return: old_population -
"""
flag = False
iteration = 0
old_population = first_population
while (not flag):
# print("While: ", iteration)
fitness_old_population = fitness(stock_values, old_population, budget)
children = generate_children(first_population, crossover_function, crossover_rate)
parents_and_children = old_population + children
chromossomes_parents_children_mutated = mutation_chromossome(parents_and_children, mutation_func, mutation_rate)
chromossomes_parents_children_mutated = random.sample(chromossomes_parents_children_mutated, len(chromossomes_parents_children_mutated))
new_population, fitness_new_population = roulette_selection(stock_values, chromossomes_parents_children_mutated, budget, len(old_population))
flag = (stop_criterion(fitness_old_population, fitness_new_population, delta) and iteration > min_iteration_converge) or iteration > 100
iteration = iteration + 1
old_population = new_population
return old_population
# -
# # Let's play
# ## Primeiro conjunto de testes - Defuzzi: Risk - centroid Pct - LOM
# ### Variaveis que serão usadas para a primeira população
# Para as ações da Alphabet serão testados:
# 1. Dados Fixos:
# 1. Budget → 10.000,00
# 2. Delta para considerar convergência → 0.01
# 2. Dados variáveis:
# 1. População:
# 1. Para uma população de 100 e outra de 1000, testamos:
# 1. Duas funções de Crossover:
# 1. Taxa de Crossover → 0.3
# 2. Taxa de Crossover → 0.8
# 2. Duas funções de Mutação:
# 1. Taxa de Mutação → 0.3
# 2. Taxa de Mutação → 0.8
#
# #### Primeira iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.01
# 4. Crossover → crossover
# 1. Taxa → 0.3
# 5. Mutação → mutation
# 1. Taxa → 0.3
# 6. Melhor Resultado:
# [[0.029757806673384436, 16, 10, 339], 1.6809029999999998]
# 7. Pior Resultado:
# [[0.0903877154523101, 24, 3, 304], 0.0]
# 8. Média:
# 0.3572531
# 9. Desvio:
# 0.409838862518003
# Dados fixos que serão usados para todas as iterações de 100 cromossomos
population_len = 10
budget = 10000
delta_to_converge = 0.01
min_iteration_converget = 3
np_array = excels[6].values # Irá utilizar a primeira lista de valores da bolsa
chromossomes = create_population(population_len)
# Aplicação da estratégia evolucionária
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_1 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# #### Resultados da Primeira Iteração
tempo_1 = round(time.time() - begin_time, 2)
tempo_1
# best_chromossomes_1 = best_chromossomes_temp
best_chromossomes_1 = fitness(np_array, best_chromossomes_1, budget)
best_chromossomes_1
best_chromossomes_1 = sorted(best_chromossomes_1, key=lambda x: x[1], reverse = False)
best_chromossomes_1
# Melhor cromossomo
best_1 = best_chromossomes_1[-1]
best_1
# Pior cromossomo
worst_1 = best_chromossomes_1[0]
worst_1
# Valor médio dos fitness
mean_1 = 0
for x in best_chromossomes_1: mean_1 = mean_1 + x[1]
mean_1 = mean_1/len(best_chromossomes_1)
mean_1
std_1 = 0
for x in best_chromossomes_1: std_1 = std_1 + (x[1] - mean_1) ** 2
std_1 = (std_1 / (len(best_chromossomes_1) - 1)) ** (1/2)
std_1
# #### Segunda iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.8
# 5. Mutação → mutation
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.8
begin_time = time.time()
best_chromossomes_2 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da segunda iteração
tempo_2 = 153.58
tempo_2
best_chromossomes_2 = fitness(np_array, best_chromossomes_2, budget)
best_chromossomes_2 = sorted(best_chromossomes_2, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_2 = best_chromossomes_2[-1]
best_2
# Pior cromossomo
worst_2 = best_chromossomes_2[0]
worst_2
# Valor médio dos fitness
mean_2 = 0
for x in best_chromossomes_2: mean_2 = mean_2 + x[1]
mean_2 = mean_2/len(best_chromossomes_2)
mean_2
std_2 = 0
for x in best_chromossomes_2: std_2 = std_2 + (x[1] - mean_2) ** 2
std_2 = (std_2 / (len(best_chromossomes_2) - 1)) ** (1/2)
std_2
# #### Terceira iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.3
# 5. Mutação → mutation
# 1. Taxa → 0.8
# Dados variáveis
mutation_rate = 0.8
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_3 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da terceira iteração
tempo_3 = round(time.time() - begin_time, 2)
tempo_9
best_chromossomes_3 = fitness(np_array, best_chromossomes_3, budget)
best_chromossomes_3 = sorted(best_chromossomes_3, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_3 = best_chromossomes_3[-1]
best_3
# Pior cromossomo
worst_3 = best_chromossomes_3[0]
worst_3
# Valor médio dos fitness
mean_3 = 0
for x in best_chromossomes_3: mean_3 = mean_3 + x[1]
mean_3 = mean_3/len(best_chromossomes_3)
mean_3
std_3 = 0
for x in best_chromossomes_3: std_3 = std_3 + (x[1] - mean_3) ** 2
std_3 = (std_3 / (len(best_chromossomes_3) - 1)) ** (1/2)
std_3
# #### Quarta iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover2
# 1. Taxa → 0.3
# 5. Mutação → mutation
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_4 = evolutionary_strategy2(np_array, chromossomes, budget, crossover2, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
tempo_4 = round(time.time() - begin_time, 2)
best_chromossomes_4 = fitness(np_array, best_chromossomes_4, budget)
best_chromossomes_4 = sorted(best_chromossomes_4, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_4 = best_chromossomes_4[-1]
best_4
# Pior cromossomo
worst_4 = best_chromossomes_4[0]
worst_4
# Valor médio dos fitness
mean_4 = 0
for x in best_chromossomes_4: mean_4 = mean_4 + x[1]
mean_4 = mean_4/len(best_chromossomes_4)
mean_4
std_4 = 0
for x in best_chromossomes_4: std_4 = std_4 + (x[1] - mean_4) ** 2
std_4 = (std_4 / (len(best_chromossomes_4) - 1)) ** (1/2)
std_4
# #### Quinta iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover2
# 1. Taxa → 0.8
# 5. Mutação → mutation
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.8
begin_time = time.time()
best_chromossomes_5 = evolutionary_strategy2(np_array, chromossomes, budget, crossover2, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da quinta iteração
tempo_5 = round(time.time() - begin_time, 2)
best_chromossomes_5 = fitness(np_array, best_chromossomes_5, budget)
best_chromossomes_5 = sorted(best_chromossomes_5, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_5 = best_chromossomes_5[-1]
best_5
# Pior cromossomo
worst_5 = best_chromossomes_5[0]
worst_5
# Valor médio dos fitness
mean_5 = 0
for x in best_chromossomes_5: mean_5 = mean_5 + x[1]
mean_5 = mean_5/len(best_chromossomes_5)
mean_5
std_5 = 0
for x in best_chromossomes_5: std_5 = std_5 + (x[1] - mean_5) ** 2
std_5 = (std_5 / (len(best_chromossomes_5) - 1)) ** (1/2)
std_5
# #### Sexta iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover2
# 1. Taxa → 0.3
# 5. Mutação → mutation
# 1. Taxa → 0.8
# Dados variáveis
mutation_rate = 0.8
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_6 = evolutionary_strategy2(np_array, chromossomes, budget, crossover2, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da sexta iteração
tempo_6 = round(time.time() - begin_time, 2)
best_chromossomes_6 = fitness(np_array, best_chromossomes_6, budget)
best_chromossomes_6 = sorted(best_chromossomes_6, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_6 = best_chromossomes_6[-1]
best_6
# Pior cromossomo
worst_6 = best_chromossomes_6[0]
worst_6
# Valor médio dos fitness
mean_6 = 0
for x in best_chromossomes_6: mean_6 = mean_6 + x[1]
mean_6 = mean_6/len(best_chromossomes_6)
mean_6
std_6 = 0
for x in best_chromossomes_6: std_6 = std_6 + (x[1] - mean_6) ** 2
std_6 = (std_6 / (len(best_chromossomes_6) - 1)) ** (1/2)
std_6
# #### Sétima iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.3
# 5. Mutação → mutation_v2
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_7 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation_v2, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da sétima iteração
tempo_7 = round(time.time() - begin_time, 2)
best_chromossomes_7 = fitness(np_array, best_chromossomes_7, budget)
best_chromossomes_7 = sorted(best_chromossomes_7, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_7 = best_chromossomes_7[-1]
best_7
# Pior cromossomo
worst_7 = best_chromossomes_7[0]
worst_7
# Valor médio dos fitness
mean_7 = 0
for x in best_chromossomes_7: mean_7 = mean_7 + x[1]
mean_7 = mean_7/len(best_chromossomes_7)
mean_7
std_7 = 0
for x in best_chromossomes_7: std_7 = std_7 + (x[1] - mean_7) ** 2
std_7 = (std_7 / (len(best_chromossomes_7) - 1)) ** (1/2)
std_7
# #### Oitava iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.8
# 5. Mutação → mutation_v2
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.8
begin_time = time.time()
best_chromossomes_8 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation_v2, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da oitava iteração
tempo_8 = round(time.time() - begin_time, 2)
best_chromossomes_8 = fitness(np_array, best_chromossomes_8, budget)
best_chromossomes_8 = sorted(best_chromossomes_8, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_8 = best_chromossomes_8[-1]
best_8
# Pior cromossomo
worst_8 = best_chromossomes_8[0]
worst_8
# Valor médio dos fitness
mean_8 = 0
for x in best_chromossomes_8: mean_8 = mean_8 + x[1]
mean_8 = mean_8/len(best_chromossomes_8)
mean_8
std_8 = 0
for x in best_chromossomes_8: std_8 = std_8 + (x[1] - mean_8) ** 2
std_8 = (std_8 / (len(best_chromossomes_8) - 8)) ** (1/2)
std_8
# #### Nona iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.3
# 5. Mutação → mutation_v2
# 1. Taxa → 0.8
# Dados variáveis
mutation_rate = 0.8
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_9 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation_v2, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da nona iteração
tempo_9 = round(time.time() - begin_time, 2)
best_chromossomes_9 = fitness(np_array, best_chromossomes_9, budget)
best_chromossomes_9 = sorted(best_chromossomes_9, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_9 = best_chromossomes_9[-1]
best_9
# Pior cromossomo
worst_9 = best_chromossomes_9[0]
worst_9
# Valor médio dos fitness
mean_9 = 0
for x in best_chromossomes_9: mean_9 = mean_9 + x[1]
mean_9 = mean_9/len(best_chromossomes_9)
mean_9
std_9 = 0
for x in best_chromossomes_9: std_9 = std_9 + (x[1] - mean_9) ** 2
std_9 = (std_9 / (len(best_chromossomes_9) - 1)) ** (1/2)
std_9
# ## Segundo conjunto de testes - Defuzzi: Risk/Pct - Centroid
# ### Variaveis que serão usadas para a primeira população
# Para as ações da Alphabet serão testados:
# 1. Dados Fixos:
# 1. Budget → 10.000,00
# 2. Delta para considerar convergência → 0.01
# 2. Dados variáveis:
# 1. População:
# 1. Para uma população de 100 e outra de 1000, testamos:
# 1. Duas funções de Crossover:
# 1. Taxa de Crossover → 0.3
# 2. Taxa de Crossover → 0.8
# 2. Duas funções de Mutação:
# 1. Taxa de Mutação → 0.3
# 2. Taxa de Mutação → 0.8
#
# #### Primeira iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.01
# 4. Crossover → crossover
# 1. Taxa → 0.3
# 5. Mutação → mutation
# 1. Taxa → 0.3
# 6. Melhor Resultado:
# [[0.029757806673384436, 16, 10, 339], 1.6809029999999998]
# 7. Pior Resultado:
# [[0.0903877154523101, 24, 3, 304], 0.0]
# 8. Média:
# 0.3572531
# 9. Desvio:
# 0.409838862518003
# Dados fixos que serão usados para todas as iterações de 100 cromossomos
population_len = 10
budget = 10000
delta_to_converge = 0.01
min_iteration_converget = 3
np_array = excels[6].values # Irá utilizar a primeira lista de valores da bolsa
chromossomes = create_population(population_len)
# Aplicação da estratégia evolucionária
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_1 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# #### Resultados da Primeira Iteração
tempo_1 = 68.92
tempo_1
# best_chromossomes_1 = best_chromossomes_temp
best_chromossomes_1 = fitness(np_array, best_chromossomes_1, budget)
best_chromossomes_1
best_chromossomes_1 = sorted(best_chromossomes_1, key=lambda x: x[1], reverse = False)
best_chromossomes_1
# Melhor cromossomo
best_1 = best_chromossomes_1[-1]
best_1
# Pior cromossomo
worst_1 = best_chromossomes_1[0]
worst_1
# Valor médio dos fitness
mean_1 = 0
for x in best_chromossomes_1: mean_1 = mean_1 + x[1]
mean_1 = mean_1/len(best_chromossomes_1)
mean_1
std_1 = 0
for x in best_chromossomes_1: std_1 = std_1 + (x[1] - mean_1) ** 2
std_1 = (std_1 / (len(best_chromossomes_1) - 1)) ** (1/2)
std_1
# #### Segunda iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.8
# 5. Mutação → mutation
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.8
begin_time = time.time()
best_chromossomes_2 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da segunda iteração
tempo_2 = 228.94
tempo_2
best_chromossomes_2 = fitness(np_array, best_chromossomes_2, budget)
best_chromossomes_2 = sorted(best_chromossomes_2, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_2 = best_chromossomes_2[-1]
best_2
# Pior cromossomo
worst_2 = best_chromossomes_2[0]
worst_2
# Valor médio dos fitness
mean_2 = 0
for x in best_chromossomes_2: mean_2 = mean_2 + x[1]
mean_2 = mean_2/len(best_chromossomes_2)
mean_2
std_2 = 0
for x in best_chromossomes_2: std_2 = std_2 + (x[1] - mean_2) ** 2
std_2 = (std_2 / (len(best_chromossomes_2) - 1)) ** (1/2)
std_2
# #### Terceira iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.3
# 5. Mutação → mutation
# 1. Taxa → 0.8
# Dados variáveis
mutation_rate = 0.8
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_3 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da terceira iteração
tempo_3 = 67.53
best_chromossomes_3 = fitness(np_array, best_chromossomes_3, budget)
best_chromossomes_3 = sorted(best_chromossomes_3, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_3 = best_chromossomes_3[-1]
best_3
# Pior cromossomo
worst_3 = best_chromossomes_3[0]
worst_3
# Valor médio dos fitness
mean_3 = 0
for x in best_chromossomes_3: mean_3 = mean_3 + x[1]
mean_3 = mean_3/len(best_chromossomes_3)
mean_3
std_3 = 0
for x in best_chromossomes_3: std_3 = std_3 + (x[1] - mean_3) ** 2
std_3 = (std_3 / (len(best_chromossomes_3) - 1)) ** (1/2)
std_3
# #### Quarta iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover2
# 1. Taxa → 0.3
# 5. Mutação → mutation
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_4 = evolutionary_strategy2(np_array, chromossomes, budget, crossover2, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
tempo_4 = 62.34
best_chromossomes_4 = fitness(np_array, best_chromossomes_4, budget)
best_chromossomes_4 = sorted(best_chromossomes_4, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_4 = best_chromossomes_4[-1]
best_4
# Pior cromossomo
worst_4 = best_chromossomes_4[0]
worst_4
# Valor médio dos fitness
mean_4 = 0
for x in best_chromossomes_4: mean_4 = mean_4 + x[1]
mean_4 = mean_4/len(best_chromossomes_4)
mean_4
std_4 = 0
for x in best_chromossomes_4: std_4 = std_4 + (x[1] - mean_4) ** 2
std_4 = (std_4 / (len(best_chromossomes_4) - 1)) ** (1/2)
std_4
# #### Quinta iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover2
# 1. Taxa → 0.8
# 5. Mutação → mutation
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.8
begin_time = time.time()
best_chromossomes_5 = evolutionary_strategy2(np_array, chromossomes, budget, crossover2, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da quinta iteração
tempo_5 = 444.98
best_chromossomes_5 = fitness(np_array, best_chromossomes_5, budget)
best_chromossomes_5 = sorted(best_chromossomes_5, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_5 = best_chromossomes_5[-1]
best_5
# Pior cromossomo
worst_5 = best_chromossomes_5[0]
worst_5
# Valor médio dos fitness
mean_5 = 0
for x in best_chromossomes_5: mean_5 = mean_5 + x[1]
mean_5 = mean_5/len(best_chromossomes_5)
mean_5
std_5 = 0
for x in best_chromossomes_5: std_5 = std_5 + (x[1] - mean_5) ** 2
std_5 = (std_5 / (len(best_chromossomes_5) - 1)) ** (1/2)
std_5
# #### Sexta iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover2
# 1. Taxa → 0.3
# 5. Mutação → mutation
# 1. Taxa → 0.8
# Dados variáveis
mutation_rate = 0.8
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_6 = evolutionary_strategy2(np_array, chromossomes, budget, crossover2, mutation, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da sexta iteração
tempo_6 = 136.27
best_chromossomes_6 = fitness(np_array, best_chromossomes_6, budget)
best_chromossomes_6 = sorted(best_chromossomes_6, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_6 = best_chromossomes_6[-1]
best_6
# Pior cromossomo
worst_6 = best_chromossomes_6[0]
worst_6
# Valor médio dos fitness
mean_6 = 0
for x in best_chromossomes_6: mean_6 = mean_6 + x[1]
mean_6 = mean_6/len(best_chromossomes_6)
mean_6
std_6 = 0
for x in best_chromossomes_6: std_6 = std_6 + (x[1] - mean_6) ** 2
std_6 = (std_6 / (len(best_chromossomes_6) - 1)) ** (1/2)
std_6
# #### Sétima iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.3
# 5. Mutação → mutation_v2
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_7 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation_v2, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da sétima iteração
tempo_7 = 147.09
best_chromossomes_7 = fitness(np_array, best_chromossomes_7, budget)
best_chromossomes_7 = sorted(best_chromossomes_7, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_7 = best_chromossomes_7[-1]
best_7
# Pior cromossomo
worst_7 = best_chromossomes_7[0]
worst_7
# Valor médio dos fitness
mean_7 = 0
for x in best_chromossomes_7: mean_7 = mean_7 + x[1]
mean_7 = mean_7/len(best_chromossomes_7)
mean_7
std_7 = 0
for x in best_chromossomes_7: std_7 = std_7 + (x[1] - mean_7) ** 2
std_7 = (std_7 / (len(best_chromossomes_7) - 1)) ** (1/2)
std_7
# #### Oitava iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.8
# 5. Mutação → mutation_v2
# 1. Taxa → 0.3
# Dados variáveis
mutation_rate = 0.3
crossover_rate = 0.8
begin_time = time.time()
best_chromossomes_8 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation_v2, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da oitava iteração
tempo_8 = 330.69
best_chromossomes_8 = fitness(np_array, best_chromossomes_8, budget)
best_chromossomes_8 = sorted(best_chromossomes_8, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_8 = best_chromossomes_8[-1]
best_8
# Pior cromossomo
worst_8 = best_chromossomes_8[0]
worst_8
# Valor médio dos fitness
mean_8 = 0
for x in best_chromossomes_8: mean_8 = mean_8 + x[1]
mean_8 = mean_8/len(best_chromossomes_8)
mean_8
std_8 = 0
for x in best_chromossomes_8: std_8 = std_8 + (x[1] - mean_8) ** 2
std_8 = (std_8 / (len(best_chromossomes_8) - 8)) ** (1/2)
std_8
# #### Nona iteração:
# 1. População → 100
# 2. Budget → 10.000
# 3. Delta → 0.001
# 4. Crossover → crossover
# 1. Taxa → 0.3
# 5. Mutação → mutation_v2
# 1. Taxa → 0.8
# Dados variáveis
mutation_rate = 0.8
crossover_rate = 0.3
begin_time = time.time()
best_chromossomes_9 = evolutionary_strategy2(np_array, chromossomes, budget, crossover, mutation_v2, delta_to_converge, mutation_rate, crossover_rate, min_iteration_converget)
print ("Tempo para convergir: ", round(time.time() - begin_time, 2), "segundos")
# Resultado da nona iteração
tempo_9 = 109.46
best_chromossomes_9 = fitness(np_array, best_chromossomes_9, budget)
best_chromossomes_9 = sorted(best_chromossomes_9, key=lambda x: x[1], reverse = False)
# Melhor cromossomo
best_9 = best_chromossomes_9[-1]
best_9
# Pior cromossomo
worst_9 = best_chromossomes_9[0]
worst_9
# Valor médio dos fitness
mean_9 = 0
for x in best_chromossomes_9: mean_9 = mean_9 + x[1]
mean_9 = mean_9/len(best_chromossomes_9)
mean_9
std_9 = 0
for x in best_chromossomes_9: std_9 = std_9 + (x[1] - mean_9) ** 2
std_9 = (std_9 / (len(best_chromossomes_9) - 1)) ** (1/2)
std_9
| projeto3/genetic_algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/semishen/ML100Days/blob/master/Day_013_HW.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="emuqIErQ_KMw" colab_type="text"
# # 常用的 DataFrame 操作
# * merge / transform
# * subset
# * groupby
# + [markdown] id="5sLfbMUE_KMx" colab_type="text"
# # [作業目標]
# - 練習填入對應的欄位資料或公式, 完成題目的要求
# + [markdown] id="BGN8kZw-_KMy" colab_type="text"
# # [作業重點]
# - 填入適當的輸入資料, 讓後面的程式顯示題目要求的結果 (Hint: 填入對應區間或欄位即可, In[4]~In[6], Out[4]~In[6])
# - 填入z轉換的計算方式, 完成轉換後的數值 (Hint: 參照標準化公式, In[7])
# + id="9qZc05w4_KMy" colab_type="code" colab={}
# Import 需要的套件
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + id="0apRsvGv_KM5" colab_type="code" outputId="34047db8-4064-446d-8220-ecd78988cd31" colab={"base_uri": "https://localhost:8080/", "height": 321}
app_train = pd.read_csv('drive/My Drive/Colab Notebooks/ML100Days/data/application_train.csv')
app_train.head()
# + [markdown] id="9T4jf9IW_KM8" colab_type="text"
# ## 作業
# 1. 請將 app_train 中的 CNT_CHILDREN 依照下列規則分為四組,並將其結果在原本的 dataframe 命名為 CNT_CHILDREN_GROUP
# * 0 個小孩
# * 有 1 - 2 個小孩
# * 有 3 - 5 個小孩
# * 有超過 5 個小孩
#
# 2. 請根據 CNT_CHILDREN_GROUP 以及 TARGET,列出各組的平均 AMT_INCOME_TOTAL,並繪製 baxplot
# 3. 請根據 CNT_CHILDREN_GROUP 以及 TARGET,對 AMT_INCOME_TOTAL 計算 [Z 轉換](https://en.wikipedia.org/wiki/Standard_score) 後的分數
# + id="Kodv-gQ-___W" colab_type="code" outputId="41287664-864d-4823-c07b-8b1a068498b9" colab={"base_uri": "https://localhost:8080/", "height": 323}
df_CNT_CHILDREN = app_train['CNT_CHILDREN']
print(df_CNT_CHILDREN.max())
df_CNT_CHILDREN.value_counts()
# + id="CO4ZQzmx_KM9" colab_type="code" outputId="c49ef019-57a3-494a-952e-ee98ba1099c9" colab={"base_uri": "https://localhost:8080/", "height": 107}
#1
cut_rule = [0, 0.9, 2.9, 5.9, app_train['CNT_CHILDREN'].max() ]
app_train['CNT_CHILDREN_GROUP'] = pd.cut(app_train['CNT_CHILDREN'].values, cut_rule, include_lowest=True)
app_train['CNT_CHILDREN_GROUP'].value_counts()
# + id="q2_1rJoV_KM_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="6c341680-58ce-4387-8aad-d1445fe1fe39"
#2-1
grp = ['CNT_CHILDREN_GROUP', 'TARGET']
grouped_df = app_train.groupby(grp)['AMT_INCOME_TOTAL']
grouped_df.mean()
# + id="Qzc5YEBW_KNB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 727} outputId="6e0bd8a3-1d89-4c96-f7f4-0bf48ba56c47"
#2-2
plt_column = 'AMT_INCOME_TOTAL'
plt_by = grp
app_train.boxplot(column=plt_column, by = plt_by, showfliers = False, figsize=(12,12), grid=False)
plt.suptitle('')
plt.show()
# + id="XksTIyFz_KNE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="a0dcc6a2-6dcb-4548-f27d-25726c249f90"
#3
app_train['AMT_INCOME_TOTAL_Z_BY_CHILDREN_GRP-TARGET'] = grouped_df.apply(lambda x: (x-np.mean(x))/np.std(x))
app_train[['AMT_INCOME_TOTAL','AMT_INCOME_TOTAL_Z_BY_CHILDREN_GRP-TARGET']].head()
| Day_013_HW.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
#
# <h2 id="Rough-volatility-with-Python">Rough volatility with Python<a class="anchor-link" href="#Rough-volatility-with-Python">¶</a></h2><p>
# </p><p><NAME></p>
# <p>For Python Quants, New York, Friday May 6, 2016</p>
# <h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2014/07/BaruchLogo2.png" width="160"/></h3><h3><img align="right" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2014/07/MFElogo.png" width="100"/></h3>
#
#
# <h3 id="Acknowledgements">Acknowledgements<a class="anchor-link" href="#Acknowledgements">¶</a></h3><p>The code in this iPython notebook used to be in R. I am very grateful to <NAME> and <NAME> for translating my R-code to Python.</p>
# <p>For slideshow functionality I use RISE by Damián Avila.</p>
# $$
# \newcommand{\beas}{\begin{eqnarray*}}
# \newcommand{\eeas}{\end{eqnarray*}}
# \newcommand{\bea}{\begin{eqnarray}}
# \newcommand{\eea}{\end{eqnarray}}
# \newcommand{\ben}{\begin{enumerate}}
# \newcommand{\een}{\end{enumerate}}
# \newcommand{\bi}{\begin{itemize}}
# \newcommand{\ei}{\end{itemize}}
# \newcommand{\beq}{\begin{equation}}
# \newcommand{\eeq}{\end{equation}}
# \newcommand{\bv}{\begin{verbatim}}
# \newcommand{\ev}{\end{verbatim}}
# \newcommand{\E}{\mathbb{E}}
# \newcommand{\R}{\mathbb{R}}
# \newcommand{\mP}{\mathbb{P}}
# \newcommand{\mQ}{\mathbb{Q}}
# \newcommand{\sigl}{\sigma_L}
# \newcommand{\BS}{\rm BS}
# \newcommand{\vix}{\text{VIX}}
# \newcommand{\p}{\partial}
# \newcommand{\var}{{\rm var}}
# \newcommand{\cov}{{\rm cov}}
# \newcommand{\mt}{\mathbf{t}}
# \newcommand{\mS}{\mathbf{S}}
# \newcommand{\tC}{\widetilde{C}}
# \newcommand{\hC}{\widehat{C}}
# \newcommand{\cE}{\mathcal{E}}
# \newcommand{\tH}{\widetilde{H}}
# \newcommand{\cD}{\mathcal{D}}
# \newcommand{\cM}{\mathcal{M}}
# \newcommand{\cS}{\mathcal{S}}
# \newcommand{\cR}{\mathcal{R}}
# \newcommand{\cF}{\mathcal{F}}
# \newcommand{\cV}{\mathcal{V}}
# \newcommand{\cG}{\mathcal{G}}
# \newcommand{\cv}{\mathcal{v}}
# \newcommand{\cg}{\mathcal{g}}
# \newcommand{\cL}{\mathcal{L}}
# \newcommand{\cO}{\mathcal{O}}
# \newcommand{\dt}{\Delta t}
# \newcommand{\tr}{{\rm tr}}
# \newcommand{\sgn}{\mathrm{sign}}
# \newcommand{\ee}[1]{{\mathbb{E}\left[{#1}\right]}}
# \newcommand{\eef}[1]{{\mathbb{E}\left[\left.{#1}\right|\cF_t\right]}}
# \newcommand{\eefm}[2]{{\mathbb{E}^{#2}\left[\left.{#1}\right|\cF_t\right]}}
# \newcommand{\angl}[1]{{\langle{#1}\rangle}}
# $$
#
#
# <h3 id="Outline-of-presentation">Outline of presentation<a class="anchor-link" href="#Outline-of-presentation">¶</a></h3><ul>
# <li><p>The time series of historical volatility</p>
# <ul>
# <li>Scaling properties</li>
# </ul>
# </li>
# <li><p>The RFSV model</p>
# </li>
# <li><p>Pricing under rough volatility</p>
# </li>
# <li><p>Forecasting realized variance</p>
# </li>
# <li><p>The time series of variance swaps</p>
# </li>
# <li><p>Relating historical and implied</p>
# </li>
# </ul>
#
#
# <h3 id="The-time-series-of-realized-variance">The time series of realized variance<a class="anchor-link" href="#The-time-series-of-realized-variance">¶</a></h3><ul>
# <li><p>Assuming an underlying variance process $v_s$, integrated variance $
# \frac 1 \delta \,\int_t^{t+\delta}\,v_s\,ds$ may (in principle) be estimated arbitrarily accurately given enough price data.</p>
# <ul>
# <li>In practice, market microstructure noise makes estimation harder at very high frequency.</li>
# <li>Sophisticated estimators of integrated variance have been developed to adjust for market microstructure noise. See Gatheral and Oomen <sup class="reference" id="cite_ref-GO"><a href="#cite_note-GO"><span>[</span>6<span>]</span></a></sup> (for example) for details of these.</li>
# </ul>
# </li>
# </ul>
#
#
# <ul>
# <li><p>The Oxford-Man Institute of Quantitative Finance makes historical realized variance (RV) estimates freely available at <a href="http://realized.oxford-man.ox.ac.uk">http://realized.oxford-man.ox.ac.uk</a>. These estimates are updated daily.</p>
# <ul>
# <li>Each day, for 21 different indices, all trades and quotes are used to estimate realized (or integrated) variance over the trading day from open to close.</li>
# </ul>
# </li>
# </ul>
# <ul>
# <li>Using daily RV estimates as proxies for instantaneous variance, we may investigate the time series properties of $v_t$ empirically.</li>
# </ul>
#
#
# <p>First load all necessary Python libraries.</p>
#
# + active=""
#
# import warnings; warnings.simplefilter('ignore')
# import datetime as dt
# import numpy as np
# import matplotlib.pyplot as plt
# import matplotlib.mlab as mlab
# from matplotlib.mlab import stineman_interp
# import pandas as pd
# import pandas.io.data as web
# import requests
# import zipfile as zi
# import StringIO as sio
# from sklearn import datasets, linear_model
# import scipy.special as scsp
# import statsmodels.api as sm
# import math
# import seaborn as sns; sns.set()
# %matplotlib inline
#
#
# -
#
# <p>Then update and save the latest Oxford-Man data.</p>
#
# + active=""
#
# url = 'http://realized.oxford-man.ox.ac.uk/media/1366/'
# url += 'oxfordmanrealizedvolatilityindices.zip'
# data = requests.get(url, stream=True).content
# z = zi.ZipFile(sio.StringIO(data))
# z.extractall()
#
#
# -
#
# <p>There are many different estimates of realized variance, all of them very similar. We will use the realized kernel estimates denoted by ".rk".</p>
#
# + active=""
#
# df = pd.read_csv('OxfordManRealizedVolatilityIndices.csv', index_col=0, header=2 )
# rv1 = pd.DataFrame(index=df.index)
# for col in df.columns:
# if col[-3:] == '.rk':
# rv1[col] = df[col]
# rv1.index = [dt.datetime.strptime(str(date), "%Y%m%d") for date in rv1.index.values]
#
#
# -
print("Hello world")
#
# <p>Let's plot SPX realized variance.</p>
#
# +
spx = pd.DataFrame(rv1['SPX2.rk'])
spx.plot(color='red', grid=True, title='SPX realized variance',
figsize=(16, 9), ylim=(0,0.003));
# -
#
# <p>Figure 1: Oxford-Man KRV estimates of SPX realized variance from January 2000 to the current date.</p>
#
# +
spx.head()
# +
spx.tail()
# -
#
# <p>We can get SPX data from Yahoo using the <code>DataReader</code> function:</p>
#
# +
SPX = web.DataReader(name = '^GSPC',data_source = 'yahoo', start='2000-01-01')
SPX = SPX['Adj Close']
SPX.plot(title='SPX',figsize=(14, 8));
# -
#
# <h3 id="The-smoothness-of-the-volatility-process">The smoothness of the volatility process<a class="anchor-link" href="#The-smoothness-of-the-volatility-process">¶</a></h3><p>For $q\geq 0$, we define the $q$th sample moment of differences of log-volatility at a given lag $\Delta$.($\angl{\cdot}$ denotes the sample average):</p>
# $$
# m(q,\Delta)=\angl{\left|\log \sigma_{t+\Delta} -\log \sigma_{t} \right|^q}
# $$<p>For example</p>
# $$
# m(2,\Delta)=\angl{\left(\log \sigma_{t+\Delta} -\log \sigma_{t} \right)^2}
# $$<p>is just the sample variance of differences in log-volatility at the lag $\Delta$.</p>
#
#
# <h3 id="Scaling-of-$m(q,\Delta)$-with-lag-$\Delta$">Scaling of $m(q,\Delta)$ with lag $\Delta$<a class="anchor-link" href="#Scaling-of-$m(q,\Delta)$-with-lag-$\Delta$">¶</a></h3>
#
# +
spx['sqrt']= np.sqrt(spx['SPX2.rk'])
spx['log_sqrt'] = np.log(spx['sqrt'])
def del_Raw(q, x):
return [np.mean(np.abs(spx['log_sqrt'] - spx['log_sqrt'].shift(lag)) ** q)
for lag in x]
# +
plt.figure(figsize=(8, 8))
plt.xlabel('$log(\Delta)$')
plt.ylabel('$log\ m(q.\Delta)$')
plt.ylim=(-3, -.5)
zeta_q = list()
qVec = np.array([.5, 1, 1.5, 2, 3])
x = np.arange(1, 100)
for q in qVec:
plt.plot(np.log(x), np.log(del_Raw(q, x)), 'o')
model = np.polyfit(np.log(x), np.log(del_Raw(q, x)), 1)
plt.plot(np.log(x), np.log(x) * model[0] + model[1])
zeta_q.append(model[0])
print zeta_q
# -
#
# <p>Figure 2: $\log m(q,\Delta)$ as a function of $\log \Delta$, SPX.</p>
#
#
# <h3 id="Monofractal-scaling-result">Monofractal scaling result<a class="anchor-link" href="#Monofractal-scaling-result">¶</a></h3><ul>
# <li>From the above log-log plot, we see that for each $q$, $m(q,\Delta) \propto \Delta ^{\zeta_q}$.</li>
# </ul>
# <ul>
# <li>How does $\zeta_q$ scale with $q$?</li>
# </ul>
#
#
# <h3 id="Scaling-of-$\zeta_q$-with-$q$">Scaling of $\zeta_q$ with $q$<a class="anchor-link" href="#Scaling-of-$\zeta_q$-with-$q$">¶</a></h3>
#
# +
plt.figure(figsize=(8,8))
plt.xlabel('q')
plt.ylabel('$\zeta_{q}$')
plt.plot(qVec, zeta_q, 'or')
line = np.polyfit(qVec[:4], zeta_q[:4],1)
plt.plot(qVec, line[0] * qVec + line[1])
h_est= line[0]
print(h_est)
# -
#
# <p>Figure 3: Scaling of $\zeta_q$ with $q$.</p>
#
#
# <p>We find the monofractal scaling relationship</p>
# $$
# \zeta_q = q\,H
# $$<p>with $H \approx 0.13$.</p>
# <ul>
# <li>Note however that $H$ does vary over time, in a narrow range.</li>
# </ul>
# <ul>
# <li>Note also that our estimate of $H$ is biased high because we proxied instantaneous variance $v_t$ with its average over each day $\frac 1T\,\int_0^T\,v_t\,dt$, where $T$ is one trading day.</li>
# </ul>
#
#
# <h3 id="Estimated-$H$-for-all-indices">Estimated $H$ for all indices<a class="anchor-link" href="#Estimated-$H$-for-all-indices">¶</a></h3><p>We now repeat this analysis for all 21 indices in the Oxford-Man dataset.</p>
#
# +
def dlsig2(sic, x, pr=False):
if pr:
a= np.array([(sig-sig.shift(lag)).dropna() for lag in x])
a=a ** 2
print a.info()
return [np.mean((sig-sig.shift(lag)).dropna() ** 2) for lag in x]
# +
h = list()
nu = list()
for col in rv1.columns:
sig = rv1[col]
sig = np.log(np.sqrt(sig))
sig = sig.dropna()
model = np.polyfit(np.log(x), np.log(dlsig2(sig, x)), 1)
nu.append(np.sqrt(np.exp(model[1])))
h.append(model[0]/2.)
OxfordH = pd.DataFrame({'names':rv1.columns, 'h_est': h, 'nu_est': nu})
# +
OxfordH
# -
#
# <h3 id="Distributions-of-$(\log-\sigma_{t+\Delta}-\log-\sigma_t)$-for-various-lags-$\Delta$">Distributions of $(\log \sigma_{t+\Delta}-\log \sigma_t)$ for various lags $\Delta$<a class="anchor-link" href="#Distributions-of-$(\log-\sigma_{t+\Delta}-\log-\sigma_t)$-for-various-lags-$\Delta$">¶</a></h3><p>Having established these beautiful scaling results for the moments, how do the histograms look?</p>
#
# +
def plotScaling(j, scaleFactor):
col_name = rv1.columns[j]
v = rv1[col_name]
x = np.arange(1,101)
def xDel(x, lag):
return x-x.shift(lag)
def sdl(lag):
return (xDel(np.log(v), lag)).std()
sd1 = (xDel(np.log(v), 1)).std()
h = OxfordH['h_est'][j]
f, ax = plt.subplots(2,2,sharex=False, sharey=False, figsize=(10, 10))
for i_0 in range(0, 2):
for i_1 in range(0, 2):
la = scaleFactor ** (i_1*1+i_0*2)
hist_val = xDel(np.log(v), la).dropna()
std = hist_val.std()
mean = hist_val.mean()
ax[i_0][i_1].set_title('Lag = %s Days' %la)
n, bins, patches = ax[i_0][i_1].hist(hist_val.values, bins=100,
normed=1, facecolor='green',alpha=0.2)
ax[i_0][i_1].plot(bins, mlab.normpdf(bins,mean,std), "r")
ax[i_0][i_1].plot(bins, mlab.normpdf(bins,0,sd1 * la ** h), "b--")
hist_val.plot(kind='density', ax=ax[i_0][i_1])
# +
plotScaling(1,5)
# -
#
# <p>Figure 4: Histograms of $(\log \sigma_{t+\Delta}-\log \sigma_t)$ for various lags $\Delta$; normal fit in red; $\Delta=1$ normal fit scaled by $\Delta^{0.14}$ in blue.</p>
#
#
# <h3 id="Universality?">Universality?<a class="anchor-link" href="#Universality?">¶</a></h3><ul>
# <li><span>[Gatheral, <NAME> Rosenbaum]<sup class="reference" id="cite_ref-GJR"><a href="#cite_note-GJR"><span>[</span>5<span>]</span></a></sup> compute daily realized variance estimates over one hour windows for DAX and Bund futures contracts, finding similar scaling relationships.</a></sup></span></li>
# </ul>
# <ul>
# <li><p>We have also checked that Gold and Crude Oil futures scale similarly.</p>
# <ul>
# <li>Although the increments $(\log \sigma_{t+\Delta}-\log \sigma_t)$ seem to be fatter tailed than Gaussian. </li>
# </ul>
# </li>
# </ul>
#
#
# <h3 id="A-natural-model-of-realized-volatility">A natural model of realized volatility<a class="anchor-link" href="#A-natural-model-of-realized-volatility">¶</a></h3><ul>
# <li><p>As noted originally by <span>[Andersen et al.]<sup class="reference" id="cite_ref-ABDE"><a href="#cite_note-ABDE"><span>[</span>1<span>]</span></a></sup>, distributions of differences in the log of realized volatility are close to Gaussian.</a></sup></span></p>
# <ul>
# <li>This motivates us to model $\sigma_t$ as a lognormal random variable.</li>
# </ul>
# </li>
# </ul>
# <ul>
# <li>Moreover, the scaling property of variance of RV differences suggests the model:</li>
# </ul>
# <p><a name="eq:dataDriven"></a>(1)
# $$
# \log \sigma_{t+\Delta} - \log \sigma_t =\nu\,\left( W^H_{t+\Delta}-W^H_t\right)
# $$</p>
# <p>where $W^H$ is fractional Brownian motion.</p>
#
#
# <ul>
# <li>In <span>[<NAME> and Rosenbaum]<sup class="reference" id="cite_ref-GJR"><a href="#cite_note-GJR"><span>[</span>5<span>]</span></a></sup>, we refer to a stationary version of </a><a href="#eq:dataDriven">(1)</a> as the RFSV (for Rough Fractional Stochastic Volatility) model.</sup></span></li>
# </ul>
#
#
# <h3 id="Fractional-Brownian-motion-(fBm)">Fractional Brownian motion (fBm)<a class="anchor-link" href="#Fractional-Brownian-motion-(fBm)">¶</a></h3><ul>
# <li><p><em>Fractional Brownian motion</em> (fBm) $\{W^H_t; t \in \mathbb{R}\}$ is the unique Gaussian process with mean zero and autocovariance function
# $$
# \ee{ W^H_t\,W^H_s } = \frac12\,\left\{ |t|^{2\,H}+|s|^{2\,H}-|t-s|^{2\,H} \right\}
# $$
# where $H \in (0,1)$ is called the <em>Hurst index</em> or parameter.</p>
# <ul>
# <li><p>In particular, when $H=1/2$, fBm is just Brownian motion.</p>
# </li>
# <li><p>If $H>1/2$, increments are positively correlated.% so the process is trending.</p>
# </li>
# <li>If $H<1/2$, increments are negatively correlated.% so the process is reverting.</li>
# </ul>
# </li>
# </ul>
#
#
# <h3 id="Representations-of-fBm">Representations of fBm<a class="anchor-link" href="#Representations-of-fBm">¶</a></h3><p>There are infinitely many possible representations of fBm in terms of Brownian motion. For example, with $\gamma = \frac 12 - H$,</p>
# <blockquote><div style="background-color:#add8e6; color:#FFFFFF; font-style: normal; "><h4>
# <NAME></h4>
# </div>
# <div style="background-color:#E8E8E8; color:#000000; font-style: normal; ">
# <br/>
#
# $$
# W^H_t ={C_H}\,\left\{\int_{-\infty}^t \,\frac{dW_s}{(t-s)^\gamma} - \int_{-\infty}^0 \,\frac{dW_s}{(-s)^\gamma}\right\}.
# $$
# <br/>
# </div>
# </blockquote>
#
#
# <p>The choice</p>
# $$
# C_H = \sqrt{ \frac{2\,H\,\Gamma(3/2-H)}{\Gamma(H+1/2)\,\Gamma(2-2\,H)}}
# $$<p>ensures that</p>
# $$
# \ee{W^H_t\,W^H_s }= \frac{1}{2}\,\left\{t^{2 H} + s^{2 H} - |t-s|^{2 H}\right\}.
# $$
#
#
# <h3 id="Does-simulated-RSFV-data-look-real?">Does simulated RSFV data look real?<a class="anchor-link" href="#Does-simulated-RSFV-data-look-real?">¶</a></h3>
#
#
# <h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2015/02/vol2.png" width="900"/></h3>
#
#
# <p>Figure 8: Volatility of SPX (above) and of the RFSV model (below).</p>
#
#
# <h3 id="Remarks-on-the-comparison">Remarks on the comparison<a class="anchor-link" href="#Remarks-on-the-comparison">¶</a></h3><ul>
# <li><p>The simulated and actual graphs look very alike.</p>
# </li>
# <li><p>Persistent periods of high volatility alternate with low volatility periods.</p>
# </li>
# <li><p>$H \sim 0.1$ generates very rough looking sample paths (compared with $H=1/2$ for Brownian motion).</p>
# </li>
# <li><p>Hence <em>rough volatility</em>.</p>
# </li>
# </ul>
#
#
# <ul>
# <li><p>On closer inspection, we observe fractal-type behavior.</p>
# </li>
# <li><p>The graph of volatility over a small time period looks like the same graph over a much longer time period.</p>
# </li>
# <li><p>This feature of volatility has been investigated both empirically and theoretically in, for example, <span>[Bacry and Muzy]<sup class="reference" id="cite_ref-BacryMuzy"><a href="#cite_note-BacryMuzy"><span>[</span>3<span>]</span></a></sup>
# .</a></sup></span></p>
# </li>
# <li><p>In particular, their Multifractal Random Walk (MRW) is related to a limiting case of the RSFV model as $H \to 0$.</p>
# </li>
# </ul>
#
#
# <h3 id="Pricing-under-rough-volatility">Pricing under rough volatility<a class="anchor-link" href="#Pricing-under-rough-volatility">¶</a></h3><p>The foregoing behavior suggest the following model (see <span>[Bayer et al.]<sup class="reference" id="cite_ref-BayerFriz"><a href="#cite_note-BayerFriz"><span>[</span>2<span>]</span></a></sup> for volatility under the real (or historical or physical) measure $\mP$:</a></sup></span></p>
# $$
# \log \sigma_t =\nu\,W^H_t.
# $$<p>Let $\gamma=\frac{1}{2}-H$. We choose the Mandelbrot-Van Ness representation of fractional Brownian motion $W^H$ as follows:</p>
# $$
# W^H_t ={C_H}\,\left\{\int_{-\infty}^t \,\frac{dW^{\mP}_s}{(t-s)^\gamma} - \int_{-\infty}^0 \,\frac{dW^{\mP}_s}{(-s)^\gamma}\right\}.
# $$
#
#
# <p>Then</p>
# $$
# \begin{eqnarray}
# &&\log v_u - \log v_t \nonumber\\
# &=&\nu\,C_H\,\left\{ \int_t^u\,\frac{1}{(u-s)^\gamma}\,d{W}^{\mP}_s +\int_{-\infty}^t\,\left[ \frac{1}{(u-s)^\gamma}-\frac{1}{(t-s)^\gamma} \right]\,d{W}^{\mP}_s\right\}\nonumber\\
# &=:& 2\,\nu\,C_H\,\left[M_t(u)+ Z_t(u)\right].
# \end{eqnarray}
# $$<ul>
# <li><p>Note that $\eefm{M_t(u)}{\mP}=0$ and $Z_t(u)$ is $\cF_t$-measurable.</p>
# <ul>
# <li>To price options, it would seem that we would need to know $\cF_t$, the entire history of the Brownian motion $W_s$ for $s<t>
# </t></li></ul>
# </li>
# </ul>
#
#
# <h3 id="Pricing-under-$\mP$">Pricing under $\mP$<a class="anchor-link" href="#Pricing-under-$\mP$">¶</a></h3><p>Let</p>
# $$
# \tilde W^{\mP}_t(u) := \sqrt{2\,H}\,\int_t^u\,\frac{dW^{\mP}_s}{(u-s)^\gamma}
# $$<p>With
# $\eta := 2\,\nu\,C_H/\sqrt{2\,H}$ we have $2\,\nu\,C_H\, M_t(u)
# = \eta\, \tilde W^{\mP}_t(u)$ so denoting the stochastic exponential by $\cE(\cdot)$, we may write</p>
# $$
# \begin{eqnarray}
# v_u &=& v_t \exp\left\{ \eta \tilde W^{\mP}_t(u) +
# 2\,\nu\,C_H\,
# Z_t(u) \right\}\nonumber\\
# &=& \eefm{v_u}{\mP}\,\cE \left(\eta\,\tilde W^{\mP}_t(u) \right).
# %\label{eq:rBergomiP}
# \end{eqnarray}
# $$<p></p>
#
#
# <ul>
# <li>The conditional distribution of $v_u$ depends on $\cF_t$ only through the variance forecasts $\eefm{v_u}{\mP}$, </li>
# </ul>
# <ul>
# <li>To price options, one does not need to know $\cF_t$, the entire history of the Brownian motion $W_s^{\mP}$ for $s<t>
# </t></li></ul>
#
#
# <h3 id="Pricing-under-$\mQ$">Pricing under $\mQ$<a class="anchor-link" href="#Pricing-under-$\mQ$">¶</a></h3><p>Our model under $\mP$ reads:</p>
# <p><a name="eq:Pmodel"></a>(2)
# $$
# v_u =\eefm{v_u}{\mP}\,\cE\left(\eta\,\tilde W^{\mP}_t(u) \right).
# %\label{eq:Pmodel}
# $$</p>
# <p>Consider some general change of measure</p>
# $$
# dW^{\mP}_s = dW^{\mQ}_s + \lambda_s\,ds,
# %\label{eq:dQdP}
# $$<p>where $\{ \lambda_s: s > t \}$ has a natural interpretation as the price of volatility risk.</p>
#
#
# <p>We may then rewrite <a href="#eq:Pmodel">(2)</a> as</p>
# $$
# v_u
# = \eefm{v_u}{\mP}\,\cE\left(\eta\,\tilde W^{\mQ}_t(u) \right)
# \exp \left\{ \eta\,\sqrt{2\,H}\, \int_t^u\,\frac{\lambda_s}{(u-s)^\gamma}\,ds\right\}.
# %\label{eq:explicitBergomiQ1}
# $$<ul>
# <li><p>Although the conditional distribution of $v_u$ under $\mP$ is lognormal, it will not be lognormal in general under $\mQ$.</p>
# <ul>
# <li>The upward sloping smile in VIX options means $\lambda_s$ cannot be deterministic in this picture.</li>
# </ul>
# </li>
# </ul>
#
#
# <h3 id="The-rough-Bergomi-(rBergomi)-model">The rough Bergomi (rBergomi) model<a class="anchor-link" href="#The-rough-Bergomi-(rBergomi)-model">¶</a></h3><p>Let's nevertheless consider the simplest change of measure</p>
# $$
# d{W}^{\mP}_s = d{W}^{\mQ}_s + \lambda(s)\,ds,
# $$<p>where $\lambda(s)$ is a deterministic function of $s$. Then from <a href="#eq:Pmodel">(2)</a>, we would have</p>
# $$
# \begin{eqnarray}
# v_u
# &=& \eefm{v_u}{\mP}\,\cE\left(\eta\,\tilde W^{\mQ}_t(u) \right)
# \exp \left\{ \eta\,\sqrt{2\,H}\, \int_t^u\,\frac{1}{(u-s)^\gamma}\,\lambda(s)\,ds\right\}\nonumber\\
# &=& \xi_t(u) \,\cE\left(\eta\,\tilde W^{\mQ}_t(u) \right)%\label{eq:explicitBergomiQ}
# \end{eqnarray}
# $$<p>where the forward variances $\xi_t(u) = \eefm{v_u}{\mQ}$ are (at least in principle) tradable and observed in the market.</p>
#
#
# <ul>
# <li><p>$\xi_t(u)$ is the product of two terms:</p>
# <ul>
# <li>$ \eefm{v_u}{\mP}$ which depends on the historical path $\{W_s, s<t brownian="" motion="" of="" the="">
# <li>a term which depends on the price of risk $\lambda(s)$.</li>
# </t></li></ul>
# </li>
# </ul>
#
#
# <h3 id="Features-of-the-rough-Bergomi-model">Features of the rough Bergomi model<a class="anchor-link" href="#Features-of-the-rough-Bergomi-model">¶</a></h3><ul>
# <li>The rBergomi model is a non-Markovian generalization of the Bergomi model:
# $$
# \eef{v_u}\neq \E[v_u|v_t].
# $$<ul>
# <li>The rBergomi model is Markovian in the (infinite-dimensional) state vector $\eefm{v_u}{\mQ}=\xi_t(u)$.</li>
# </ul>
# </li>
# </ul>
# <ul>
# <li>We have achieved our aim from Session 1 of replacing the exponential kernels in the Bergomi model with a power-law kernel. </li>
# </ul>
# <ul>
# <li>We may therefore expect that the rBergomi model will generate a realistic term structure of ATM volatility skew.</li>
# </ul>
#
#
# <h3 id="Re-interpretation-of-the-conventional-Bergomi-model">Re-interpretation of the conventional Bergomi model<a class="anchor-link" href="#Re-interpretation-of-the-conventional-Bergomi-model">¶</a></h3><ul>
# <li><p>A conventional $n$-factor Bergomi model is not self-consistent for an arbitrary choice of the initial forward variance curve $\xi_t(u)$.</p>
# <ul>
# <li>$\xi_t(u)=\eef{v_u}$ should be consistent with the assumed dynamics.</li>
# </ul>
# </li>
# </ul>
#
#
# <ul>
# <li><p>Viewed from the perspective of the fractional Bergomi model however:</p>
# <ul>
# <li>The initial curve $\xi_t(u)$ reflects the history $\{W_s; s<t brownian="" driving="" motion="" of="" the="" time="" to="" up="">
# <li>The exponential kernels in the exponent of the conventional Bergomi model approximate more realistic power-law kernels.</li>
# </t></li></ul>
# </li>
# </ul>
# <ul>
# <li>The conventional two-factor Bergomi model is then justified in practice as a tractable Markovian engineering approximation to a more realistic fractional Bergomi model.</li>
# </ul>
#
#
# <h3 id="The-stock-price-process">The stock price process<a class="anchor-link" href="#The-stock-price-process">¶</a></h3><ul>
# <li>The observed anticorrelation between price moves and volatility moves may be modeled naturally by anticorrelating the Brownian motion $W$ that drives the volatility process with the Brownian motion driving the price process. </li>
# </ul>
# <ul>
# <li>Thus
# $$
# \frac{dS_t}{S_t}=\sqrt{v_t}\,dZ_t
# $$
# with
# $$
# dZ_t = \rho\,dW_t + \sqrt{1-\rho^2}\,dW^\perp_t
# $$
# where $\rho$ is the correlation between volatility moves and price moves.</li>
# </ul>
#
#
# <h3 id="Simulation-of-the--rBergomi-model">Simulation of the rBergomi model<a class="anchor-link" href="#Simulation-of-the--rBergomi-model">¶</a></h3><p>We simulate the rBergomi model as follows:</p>
# <ul>
# <li>Construct the joint covariance matrix for the Volterra process $\tilde
# W$ and the Brownian motion $Z$ and compute its Cholesky decomposition.</li>
# </ul>
# <ul>
# <li>For each time, generate iid normal random vectors {and
# multiply them by the lower-triangular matrix obtained by the Cholesky
# decomposition} to get a $m \times 2\,n$ matrix of paths of $\tilde W$ and $Z$ with the correct joint marginals.</li>
# </ul>
#
#
# <ul>
# <li>With these paths held in memory, we may evaluate the expectation under $\mQ$ of any payoff of interest.</li>
# </ul>
# <ul>
# <li><p>This procedure is very slow!</p>
# <ul>
# <li>Speeding up the simulation is work in progress.</li>
# </ul>
# </li>
# </ul>
#
#
# <h3 id="Guessing-rBergomi-model-parameters">Guessing rBergomi model parameters<a class="anchor-link" href="#Guessing-rBergomi-model-parameters">¶</a></h3><ul>
# <li>The rBergomi model has only three parameters: $H$, $\eta$ and $\rho$.</li>
# </ul>
# <ul>
# <li>If we had a fast simulation, we could just iterate on these parameters to find the best fit to observed option prices. But we don't.</li>
# </ul>
#
#
# <ul>
# <li><p>However, the model parameters $H$, $\eta$ and $\rho$ have very direct interpretations:</p>
# <ul>
# <li><p>$H$ controls the decay of ATM skew $\psi(\tau)$ for very short expirations.</p>
# </li>
# <li><p>The product $\rho\,\eta$ sets the level of the ATM skew for longer expirations.</p>
# <ul>
# <li>Keeping $\rho\,\eta$ constant but decreasing $\rho$ (so as to make it more negative) pushes the minimum of each smile towards higher strikes. </li>
# </ul>
# </li>
# </ul>
# </li>
# </ul>
#
#
# <ul>
# <li>So we can guess parameters in practice.</li>
# </ul>
# <ul>
# <li>As we will see, even without proper calibration (<em>i.e.</em> just guessing parameters), rBergomi model fits to the volatility surface are amazingly good.</li>
# </ul>
#
#
# <h3 id="SPX-smiles-in-the-rBergomi-model">SPX smiles in the rBergomi model<a class="anchor-link" href="#SPX-smiles-in-the-rBergomi-model">¶</a></h3><ul>
# <li><p>In Figures 9 and 10, we show how well a rBergomi model simulation with guessed parameters fits the SPX option market as of February 4, 2010, a day when the ATM volatility term structure happened to be pretty flat.</p>
# <ul>
# <li>rBergomi parameters were: $H=0.07$, $\eta=1.9$, $\rho=-0.9$.</li>
# </ul>
# </li>
# </ul>
# <ul>
# <li>Only three parameters to get a very good fit to the whole SPX volatility surface!</li>
# </ul>
#
#
# <h3 id="rBergomi-fits-to-SPX-smiles-as-of-04-Feb-2010">rBergomi fits to SPX smiles as of 04-Feb-2010<a class="anchor-link" href="#rBergomi-fits-to-SPX-smiles-as-of-04-Feb-2010">¶</a></h3><h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2015/02/spxSmiles20140204-07.png" width="900"/></h3>
#
#
# <p>Figure 9: Red and blue points represent bid and offer SPX implied volatilities; orange smiles are from the rBergomi simulation.</p>
#
#
# <h3 id="Shortest-dated-smile-as-of-February-4,-2010">Shortest dated smile as of February 4, 2010<a class="anchor-link" href="#Shortest-dated-smile-as-of-February-4,-2010">¶</a></h3><h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2015/02/spxSmiles20140204-07-1.png" width="900"/></h3>
#
#
# <p>Figure 10: Red and blue points represent bid and offer SPX implied volatilities; orange smile is from the rBergomi simulation.</p>
#
#
# <h3 id="ATM-volatilities-and-skews">ATM volatilities and skews<a class="anchor-link" href="#ATM-volatilities-and-skews">¶</a></h3><p>In Figures 11 and 12, we see just how well the rBergomi model can match empirical skews and vols. Recall also that the parameters we used are just guesses!</p>
#
#
# <h3 id="Term-structure-of-ATM-skew-as-of-February-4,-2010">Term structure of ATM skew as of February 4, 2010<a class="anchor-link" href="#Term-structure-of-ATM-skew-as-of-February-4,-2010">¶</a></h3><h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2015/02/atmSkew20100204.png" width="900"/></h3>
#
#
# <p>Figure 11: Blue points are empirical skews; the red line is from the rBergomi simulation.</p>
#
#
# <h3 id="Term-structure-of-ATM-vol-as-of-February-4,-2010">Term structure of ATM vol as of February 4, 2010<a class="anchor-link" href="#Term-structure-of-ATM-vol-as-of-February-4,-2010">¶</a></h3><h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2015/02/atmVols20100204.png" width="900"/></h3>
#
#
# <p>Figure 12: Blue points are empirical ATM volatilities; the red line is from the rBergomi simulation.</p>
#
#
# <h3 id="Another-date">Another date<a class="anchor-link" href="#Another-date">¶</a></h3><ul>
# <li>Now we take a look at another date: August 14, 2013, two days before the last expiration date in our dataset.</li>
# </ul>
# <ul>
# <li>Options set at the open of August 16, 2013 so only one trading day left.</li>
# </ul>
# <ul>
# <li>Note in particular that the extreme short-dated smile is well reproduced by the rBergomi model.</li>
# </ul>
# <ul>
# <li>There is no need to add jumps!</li>
# </ul>
#
#
# <h3 id="SPX-smiles-as-of-August-14,-2013">SPX smiles as of August 14, 2013<a class="anchor-link" href="#SPX-smiles-as-of-August-14,-2013">¶</a></h3><h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2015/02/spxSmiles20130814-05.png" width="900"/></h3>
#
#
# <p>Figure 13: Red and blue points represent bid and offer SPX implied volatilities; orange smiles are from the rBergomi simulation.</p>
#
#
# <h3 id="The-forecast-formula">The forecast formula<a class="anchor-link" href="#The-forecast-formula">¶</a></h3><ul>
# <li>In the RFSV model <a href="#eq:dataDriven">(1)</a>, $\log \sigma_t \approx \nu\,W^H_t+C$ for some constant $C$.</li>
# </ul>
# <ul>
# <li><span>[Nuzman and Poor]<sup class="reference" id="cite_ref-NuzmanPoor"><a href="#cite_note-NuzmanPoor"><span>[</span>7<span>]</span></a></sup> show that $W^H_{t+\Delta}$ is conditionally Gaussian with conditional expectation</a></sup></span></li>
# </ul>
# $$\E[W^H_{t+\Delta}|\cF_t]=\frac{\cos(H\pi)}{\pi} \Delta^{H+1/2} \int_{-\infty}^t \frac{W^H_s}{(t-s+\Delta)(t-s)^{H+1/2}} ds
# $$<p></p>
# <p>and conditional variance</p>
# $$
# \text{Var}[W^H_{t+\Delta}|\cF_t]=c\,\Delta^{2H}.
# $$<p>where $$
# c = \frac{\Gamma(3/2-H)}{\Gamma(H+1/2)\,\Gamma(2-2H)}.
# $$</p>
#
#
# <h3 id="The-forecast-formula">The forecast formula<a class="anchor-link" href="#The-forecast-formula">¶</a></h3><p>Thus, we obtain</p>
# <blockquote><div style="background-color:#add8e6; color:#FFFFFF; font-style: normal; "><h4>
# Variance forecast formula</h4>
# </div>
# <div style="background-color:#E8E8E8; color:#000000; font-style: normal; ">
# <br/>
# <a name="eq:vForecast"></a>(3)
# $$
# \eefm{v_{t+\Delta}}{\mP}=\exp\left\{\eefm{\log(v_{t+\Delta})}{\mP}+2\, c\,\nu^2\Delta^{2\,H}\right\}
# %\label{eq:vForecast}
# $$
#
# <br/>
# </div>
# </blockquote><p>where</p>
# $$
# \beas
# &&\eefm{\log v_{t+\Delta}}{\mP}\\
# &&= \frac{\cos(H\pi)}{\pi} \Delta^{H+1/2} \int_{-\infty}^t \frac{\log v_s}{(t-s+\Delta)(t-s)^{H+1/2}} ds.
# \eeas
# $$
#
#
# <h3 id="Implement-variance-forecast-in-Python">Implement variance forecast in Python<a class="anchor-link" href="#Implement-variance-forecast-in-Python">¶</a></h3>
#
# +
def c_tilde(h):
return scsp.gamma(3. / 2. - h) / scsp.gamma(h + 1. / 2.) * scsp.gamma(2. - 2. * h)
def forecast_XTS(rvdata, h, date, nLags, delta, nu):
i = np.arange(nLags)
cf = 1./((i + 1. / 2.) ** (h + 1. / 2.) * (i + 1. / 2. + delta))
ldata = rvdata.truncate(after=date)
l = len(ldata)
ldata = np.log(ldata.iloc[l - nLags:])
ldata['cf'] = np.fliplr([cf])[0]
# print ldata
ldata = ldata.dropna()
fcst = (ldata.iloc[:, 0] * ldata['cf']).sum() / sum(ldata['cf'])
return math.exp(fcst + 2 * nu ** 2 * c_tilde(h) * delta ** (2 * h))
# -
#
# <h3 id="SPX-actual-vs-forecast-variance">SPX actual vs forecast variance<a class="anchor-link" href="#SPX-actual-vs-forecast-variance">¶</a></h3>
#
# +
rvdata = pd.DataFrame(rv1['SPX2.rk'])
nu = OxfordH['nu_est'][0] # Vol of vol estimate for SPX
h = OxfordH['h_est'][0]
n = len(rvdata)
delta = 1
nLags = 500
dates = rvdata.iloc[nLags:n-delta].index
rv_predict = [forecast_XTS(rvdata, h=h, date=d, nLags=nLags,
delta=delta, nu=nu) for d in dates]
rv_actual = rvdata.iloc[nLags+delta:n].values
# -
#
# <h3 id="Scatter-plot-of-delta-days-ahead-predictions">Scatter plot of delta days ahead predictions<a class="anchor-link" href="#Scatter-plot-of-delta-days-ahead-predictions">¶</a></h3>
#
# +
plt.figure(figsize=(8, 8))
plt.plot(rv_predict, rv_actual, 'bo');
# -
#
# <p>Figure 14: Actual vols vs predicted vols.</p>
#
#
# <h3 id="Superimpose-actual-and-predicted-vols">Superimpose actual and predicted vols<a class="anchor-link" href="#Superimpose-actual-and-predicted-vols">¶</a></h3>
#
# +
plt.figure(figsize=(11, 6))
vol_actual = np.sqrt(np.multiply(rv_actual,252))
vol_predict = np.sqrt(np.multiply(rv_predict,252))
plt.plot(vol_actual, "b")
plt.plot(vol_predict, "r");
# -
#
# <p>Figure 15: Actual volatilities in blue; predicted vols in red.</p>
#
#
# <h3 id="Forecasting-the-variance-swap-curve">Forecasting the variance swap curve<a class="anchor-link" href="#Forecasting-the-variance-swap-curve">¶</a></h3><p>Finally, we forecast the whole variance swap curve using the variance forecasting formula <a href="#eq:vForecast">(3)</a>.</p>
#
# +
def xi(date, tt, nu,h, tscale): # dt=(u-t) is in units of years
rvdata = pd.DataFrame(rv1['SPX2.rk'])
return [ forecast_XTS(rvdata,h=h,date=date,nLags=500,delta=dt*tscale,nu=nu) for dt in tt]
nu = OxfordH["nu_est"][0]
h = OxfordH["h_est"][0]
def varSwapCurve(date, bigT, nSteps, nu, h, tscale, onFactor):
# Make vector of fwd variances
tt = [ float(i) * (bigT) / nSteps for i in range(nSteps+1)]
delta_t = tt[1]
xicurve = xi(date, tt, nu, h, tscale)
xicurve_mid = (np.array(xicurve[0:nSteps]) + np.array(xicurve[1:nSteps+1])) / 2
xicurve_int = np.cumsum(xicurve_mid) * delta_t
varcurve1 = np.divide(xicurve_int, np.array(tt[1:]))
varcurve = np.array([xicurve[0],]+list(varcurve1))
varcurve = varcurve * onFactor * tscale # onFactor is to compensate for overnight moves
res = pd.DataFrame({"texp":np.array(tt), "vsQuote":np.sqrt(varcurve)})
return(res)
# +
def varSwapForecast(date,tau,nu,h,tscale,onFactor):
vsc = varSwapCurve(date, bigT=2.5, nSteps=100, nu=nu, h=h,
tscale=tscale, onFactor=onFactor) # Creates the whole curve
x = vsc['texp']
y = vsc['vsQuote']
res = stineman_interp(tau,x,y,None)
return(res)
# Test the function
tau = (.25,.5,1,2)
date = dt.datetime(2008,9,8)
varSwapForecast(date,tau,nu=nu,h=h,tscale=252,onFactor=1)
# -
#
# <h3 id="'Constructing-a-time-series-of-variance-swap-curves">'Constructing a time series of variance swap curves<a class="anchor-link" href="#'Constructing-a-time-series-of-variance-swap-curves">¶</a></h3><p>For each of 2,658 days from Jan 27, 2003 to August 31, 2013:</p>
# <ul>
# <li>We compute proxy variance swaps from closing prices of SPX options sourced from OptionMetrics (www.optionmetrics.com) via WRDS.</li>
# </ul>
# <ul>
# <li>We form the forecasts $\eefm{v_u}{\mP}$ using <a href="#eq:vForecast">(3)</a> with 500 lags of SPX RV data sourced from The Oxford-Man Institute of Quantitative Finance (<a href="http://realized.oxford-man.ox.ac.uk">http://realized.oxford-man.ox.ac.uk</a>).</li>
# </ul>
#
#
# <ul>
# <li>We note that the actual variance swap curve is a factor (of roughly 1.4) higher than the forecast, which we may attribute to a combination of overnight movements of the index and the price of volatility risk.</li>
# </ul>
# <ul>
# <li>Forecasts must therefore be rescaled to obtain close-to-close realized variance forecasts.</li>
# </ul>
#
#
# <h3 id="3-month-forecast-vs-actual-variance-swaps">3-month forecast vs actual variance swaps<a class="anchor-link" href="#3-month-forecast-vs-actual-variance-swaps">¶</a></h3><h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2015/02/vsfa3m.png" width="900"/></h3>
#
#
# <p>Figure 16: Actual (proxy) 3-month variance swap quotes in blue vs forecast in red (with no scaling factor).</p>
#
#
# <h3 id="Ratio-of-actual-to-forecast">Ratio of actual to forecast<a class="anchor-link" href="#Ratio-of-actual-to-forecast">¶</a></h3><h3><img align="left" src="http://mfe.baruch.cuny.edu/wp-content/uploads/2015/02/3mratio.png" width="900"/></h3>
#
#
# <p>Figure 17: The ratio between 3-month actual variance swap quotes and 3-month forecasts.</p>
#
#
# <h3 id="The-Lehman-weekend">The Lehman weekend<a class="anchor-link" href="#The-Lehman-weekend">¶</a></h3><ul>
# <li><p>Empirically, it seems that the variance curve is a simple scaling factor times the forecast, but that this scaling factor is time-varying.</p>
# <ul>
# <li>We can think of this factor as having two multiplicative components: the overnight factor, and the price of volatility risk.</li>
# </ul>
# </li>
# </ul>
# <ul>
# <li>Recall that as of the close on Friday September 12, 2008, it was widely believed that Lehman Brothers would be rescued over the weekend. By Monday morning, we knew that Lehman had failed. </li>
# </ul>
#
#
# <ul>
# <li>In Figure 18, we see that variance swap curves just before and just after the collapse of Lehman are just rescaled versions of the RFSV forecast curves.</li>
# </ul>
#
#
# <h3 id="We-need-variance-swap-estimates-for-12-Sep-2008-and-15-Sep-2008">We need variance swap estimates for 12-Sep-2008 and 15-Sep-2008<a class="anchor-link" href="#We-need-variance-swap-estimates-for-12-Sep-2008-and-15-Sep-2008">¶</a></h3><p>We proxy these by taking SVI fits for the two dates and computing the log-strips.</p>
#
# +
varSwaps12 =(
0.2872021, 0.2754535, 0.2601864, 0.2544684, 0.2513854, 0.2515314,
0.2508418, 0.2520099, 0.2502763, 0.2503309, 0.2580933, 0.2588361,
0.2565093)
texp12 = (
0.01916496, 0.04654346, 0.09582478, 0.19164956, 0.26830938, 0.29842574,
0.51745380, 0.54483231, 0.76659822, 0.79397673, 1.26488706, 1.76317591,
2.26146475)
varSwaps15 = (
0.4410505, 0.3485560, 0.3083603, 0.2944378, 0.2756881, 0.2747838,
0.2682212, 0.2679770, 0.2668113, 0.2706713, 0.2729533, 0.2689598,
0.2733176)
texp15 = (
0.01095140, 0.03832991, 0.08761123, 0.18343600, 0.26009582, 0.29021218,
0.50924025, 0.53661875, 0.75838467, 0.78576318, 1.25667351, 1.75496235,
2.25325120)
# -
#
# <h3 id="Actual-vs-predicted-over-the-Lehman-weekend">Actual vs predicted over the Lehman weekend<a class="anchor-link" href="#Actual-vs-predicted-over-the-Lehman-weekend">¶</a></h3>
#
# +
nu = OxfordH['nu_est'][0]
h = OxfordH['h_est'][0]
date1 = dt.datetime(2008, 9, 12)
date2 = dt.datetime(2008, 9, 15)
# Variance curve fV model forecasts
tau1000 = [ float(i) * 2.5 / 1000. for i in range(1,1001)]
vs1 = varSwapForecast(date1, tau1000, nu=nu,h=h, tscale=252, onFactor=1.29)
vs2 = varSwapForecast(date2, tau1000, nu=nu,h=h, tscale=252, onFactor=1.29)
# +
plt.figure(figsize=(11, 6))
plt.plot(texp12, varSwaps12, "r")
plt.plot(texp15, varSwaps15, "b")
plt.plot(tau1000, vs1, "r--")
plt.plot(tau1000, vs2, "b--");
# -
#
# <p>Figure 18: SPX variance swap curves as of September 12, 2008 (red) and September 15, 2008 (blue). The dashed curves are RFSV model forecasts rescaled by the 3-month ratio ($1.29$) as of the Friday close.</p>
#
#
# <h3 id="Remarks">Remarks<a class="anchor-link" href="#Remarks">¶</a></h3><p>We note that</p>
# <ul>
# <li>The actual variance swaps curves are very close to the forecast curves, up to a scaling factor.</li>
# </ul>
# <ul>
# <li>We are able to explain the change in the variance swap curve with only one extra observation: daily variance over the trading day on Monday 15-Sep-2008. </li>
# </ul>
# <ul>
# <li>The SPX options market appears to be backward-looking in a very sophisticated way.</li>
# </ul>
#
#
# <h3 id="The-Flash-Crash">The Flash Crash<a class="anchor-link" href="#The-Flash-Crash">¶</a></h3><ul>
# <li>The so-called Flash Crash of Thursday May 6, 2010 caused intraday realized variance to be much higher than normal. </li>
# </ul>
# <ul>
# <li>In Figure 19, we plot the actual variance swap curves as of the Wednesday and Friday market closes together with forecast curves rescaled by the 3-month ratio as of the close on Wednesday May 5 (which was $2.52$). </li>
# </ul>
# <ul>
# <li>We see that the actual variance curve as of the close on Friday is consistent with a forecast from the time series of realized variance that <em>includes</em> the anomalous price action of Thursday May 6. </li>
# </ul>
#
#
# <h3 id="Variance-swap-estimates">Variance swap estimates<a class="anchor-link" href="#Variance-swap-estimates">¶</a></h3><p>We again proxy variance swaps for 05-May-2010, 07-May-2010 and 10-May-2010 by taking SVI fits (see <span>[Gatheral and Jacquier]<sup class="reference" id="cite_ref-GatheralJacquierSSVI"><a href="#cite_note-GatheralJacquierSSVI"><span>[</span>4<span>]</span></a></sup> ) for the three dates and computing the log-strips.</a></sup></span></p>
#
# +
varSwaps5 = (
0.4250369, 0.2552473, 0.2492892, 0.2564899, 0.2612677, 0.2659618, 0.2705928, 0.2761203,
0.2828139, 0.2841165, 0.2884955, 0.2895839, 0.2927817, 0.2992602, 0.3116500)
texp5 = (
0.002737851, 0.043805613, 0.120465435, 0.150581793, 0.197125257, 0.292950034,
0.369609856, 0.402464066, 0.618754278, 0.654346338, 0.867898700, 0.900752909,
1.117043121, 1.615331964, 2.631074606)
varSwaps7 = (
0.5469727, 0.4641713, 0.3963352, 0.3888213, 0.3762354, 0.3666858, 0.3615814, 0.3627013,
0.3563324, 0.3573946, 0.3495730, 0.3533829, 0.3521515, 0.3506186, 0.3594066)
texp7 = (
0.01642710, 0.03832991, 0.11498973, 0.14510609, 0.19164956, 0.28747433, 0.36413415,
0.39698836, 0.61327858, 0.64887064, 0.86242300, 0.89527721, 1.11156742, 1.60985626,
2.62559890)
varSwaps10 = (
0.3718439, 0.3023223, 0.2844810, 0.2869835, 0.2886912, 0.2905637, 0.2957070, 0.2960737,
0.3005086, 0.3031188, 0.3058492, 0.3065815, 0.3072041, 0.3122905, 0.3299425)
texp10 = (
0.008213552, 0.030116359, 0.106776181, 0.136892539, 0.183436003, 0.279260780,
0.355920602, 0.388774812, 0.605065024, 0.640657084, 0.854209446, 0.887063655,
1.103353867, 1.601642710, 2.617385352)
# +
date1 = dt.datetime(2010, 5, 5)
date2 = dt.datetime(2010, 5, 7)
vsf5 = varSwapCurve(date1, bigT=2.5, nSteps=100, nu=nu, h=h, tscale=252, onFactor=2.52)
vsf7 = varSwapCurve(date2, bigT=2.5, nSteps=100, nu=nu, h=h, tscale=252, onFactor=2.52)
# +
plt.figure(figsize=(11, 6))
plt.plot(texp5, varSwaps5, "r", label='May 5')
plt.plot(texp7, varSwaps7, "g", label='May 7')
plt.legend()
plt.xlabel("Time to maturity")
plt.ylabel("Variance swap quote")
plt.plot(vsf5['texp'], vsf5['vsQuote'], "r--")
plt.plot(vsf7['texp'], vsf7['vsQuote'], "g--");
# -
#
# <p>Figure 19: SPX variance swap curves as of May 5, 2010 (red) and May 7, 2010 (green). The dashed curves are RFSV model forecasts rescaled by the 3-month ratio ($2.52$) as of the close on Wednesday May 5. The curve as of the close on May 7 is consistent with the forecast <strong>including</strong> the crazy moves on May 6.</p>
#
#
# <h3 id="The-weekend-after-the-Flash-Crash">The weekend after the Flash Crash<a class="anchor-link" href="#The-weekend-after-the-Flash-Crash">¶</a></h3><p>Now we plot forecast and actual variance swap curves as of the close on Friday May 7 and Monday May 10.</p>
#
# +
date1 = dt.datetime(2010,5,7)
date2 = dt.datetime(2010,5,10)
vsf7 = varSwapCurve(date1, bigT=2.5, nSteps=100, nu=nu, h=h, tscale=252, onFactor=2.52)
vsf10 = varSwapCurve(date2, bigT=2.5, nSteps=100, nu=nu, h=h, tscale=252, onFactor=2.52)
# +
plt.figure(figsize=(11, 6))
plt.plot(texp7, varSwaps7, "g", label='May 7')
plt.plot(texp10, varSwaps10, "m", label='May 10')
plt.legend()
plt.xlabel("Time to maturity")
plt.ylabel("Variance swap quote")
plt.plot(vsf7['texp'], vsf7['vsQuote'], "g--")
plt.plot(vsf10['texp'], vsf10['vsQuote'], "m--");
# -
#
# <p>Figure 20: The May 10 actual curve is inconsistent with a forecast that includes the Flash Crash.</p>
#
#
# <p>Now let's see what happens if we exclude the Flash Crash from the time series used to generate the variance curve forecast.</p>
#
# +
plt.figure(figsize=(11, 6))
ax = plt.subplot(111)
rvdata_p = rvdata.drop((dt.datetime(2010, 5, 6)), axis=0)
rvdata.loc["2010-05-04":"2010-05-10"].plot(ax=ax, legend=False)
rvdata_p.loc["2010-05-04":"2010-05-10"].plot(ax=ax, legend=False);
# -
#
# <p>Figure 21: <code>rvdata_p</code> has the May 6 realized variance datapoint eliminated (green line). Notice the crazy realized variance estimate for May 6!</p>
# <p>We need a new variance curve forecast function that uses the new time series.</p>
#
# +
def xip(date, tt, nu,h, tscale): # dt=(u-t) is in units of years
rvdata = pd.DataFrame(rv1['SPX2.rk'])
rvdata_p = rvdata.drop((dt.datetime(2010, 5, 6)), axis=0)
return [ forecast_XTS(rvdata_p, h=h, date=date,nLags=500,
delta=delta_t * tscale, nu=nu) for delta_t in tt]
nu = OxfordH["nu_est"][0]
h = OxfordH["h_est"][0]
def varSwapCurve_p(date, bigT, nSteps, nu, h, tscale, onFactor):
# Make vector of fwd variances
tt = [ float(i) * (bigT) / nSteps for i in range(nSteps+1)]
delta_t = tt[1]
xicurve = xip(date, tt, nu, h, tscale)
xicurve_mid = (np.array(xicurve[0:nSteps]) + np.array(xicurve[1:nSteps + 1])) / 2
xicurve_int = np.cumsum(xicurve_mid) * delta_t
varcurve1 = np.divide(xicurve_int, np.array(tt[1:]))
varcurve = np.array([xicurve[0],]+list(varcurve1))
varcurve = varcurve * onFactor * tscale # onFactor is to compensate for overnight moves
res = pd.DataFrame({"texp":np.array(tt), "vsQuote":np.sqrt(varcurve)})
return(res)
def varSwapForecast_p(date, tau, nu, h, tscale, onFactor):
vsc = varSwapCurve_p(date, bigT=2.5, nSteps=100, nu=nu, h=h,
tscale=tscale, onFactor=onFactor) # Creates the whole curve
x = vsc['texp']
y = vsc['vsQuote']
res = stineman_interp(tau, x, y, None)
return(res)
# Test the function
tau = (.25, .5 ,1, 2)
date = dt.datetime(2010, 5, 10)
varSwapForecast_p(date, tau, nu=nu, h=h, tscale=252, onFactor=1. / (1 - .35))
# -
#
# <p>Finally, we compare our new forecast curves with the actuals.</p>
#
# +
date1 = dt.datetime(2010, 5, 7)
date2 = dt.datetime(2010, 5, 10)
vsf7 = varSwapCurve(date1, bigT=2.5, nSteps=100, nu=nu, h=h, tscale=252, onFactor=2.52)
vsf10p = varSwapCurve_p(date2, bigT=2.5, nSteps=100, nu=nu, h=h, tscale=252, onFactor=2.52)
# +
plt.figure(figsize=(11, 6))
plt.plot(texp7, varSwaps7, "g", label='May 7')
plt.plot(texp10, varSwaps10, "m", label='May 10')
plt.legend()
plt.xlabel("Time to maturity")
plt.ylabel("Variance swap quote")
plt.plot(vsf7['texp'], vsf7['vsQuote'], "g--")
plt.plot(vsf10p['texp'], vsf10p['vsQuote'], "m--");
# -
#
# <p>Figure 22: The May 10 actual curve is consistent with a forecast that excludes the Flash Crash.</p>
#
#
# <h3 id="Resetting-of-expectations-over-the-weekend">Resetting of expectations over the weekend<a class="anchor-link" href="#Resetting-of-expectations-over-the-weekend">¶</a></h3><ul>
# <li>In Figures 20 and 22, we see that the actual variance swap curve on Monday, May 10 is consistent with a forecast that excludes the Flash Crash.</li>
# </ul>
# <ul>
# <li>Volatility traders realized that the Flash Crash should not influence future realized variance projections.</li>
# </ul>
#
#
# <h3 id="Summary">Summary<a class="anchor-link" href="#Summary">¶</a></h3><ul>
# <li><p>We uncovered a remarkable monofractal scaling relationship in historical volatility.</p>
# <ul>
# <li>A corollary is that volatility is not a long memory process, as widely believed.</li>
# </ul>
# </li>
# </ul>
# <ul>
# <li>This leads to a natural non-Markovian stochastic volatility model under $\mP$.</li>
# </ul>
# <ul>
# <li>The simplest specification of $\frac{d\mQ}{d\mP}$ gives a non-Markovian generalization of the Bergomi model.</li>
# </ul>
#
#
# <ul>
# <li>The history of the Brownian motion $\lbrace W_s, s<t curve="" encoded="" for="" forward="" in="" is="" market.="" observed="" pricing="" required="" the="" variance="" which="">
# </t></li></ul>
# <ul>
# <li>This model fits the observed volatility surface surprisingly well with very few parameters.</li>
# </ul>
# <ul>
# <li>For perhaps the first time, we have a simple consistent model of historical and implied volatility.</li>
# </ul>
#
#
# <h2 id="References">References<a class="anchor-link" href="#References">¶</a></h2><p><br/></p>
# <div class="reflist" style="list-style-type: decimal;">
# <ol>
# <li id="cite_note-ABDE"><span class="mw-cite-backlink"><b><a href="#cite_ref-ABDE">^</a></b></span>
# <NAME>, <NAME>, <NAME>, and <NAME>, The distribution of realized stock return volatility, *Journal of Financial Economics* **61**(1) 43-76 (2001).
# </li>
# <li id="cite_note-BayerFriz"><span class="mw-cite-backlink"><b><a href="#cite_ref-BayerFriz">^</a></b></span>
# <NAME>, <NAME> and <NAME>, Pricing under rough volatility, *Quantitative Finance* forthcoming, available at http://papers.ssrn.com/sol3/papers.cfm?abstract_id=2554754, (2015).
# </li>
# <li id="cite_note-BacryMuzy"><span class="mw-cite-backlink"><b><a href="#cite_ref-BacryMuzy">^</a></b></span>
# <NAME> and <NAME>, Log-infinitely divisible multifractal processes,
# *Communications in Mathematical Physics* **236**(3) 449-475 (2003).</li>
# <li id="cite_note-GatheralJacquierSSVI"><span class="mw-cite-backlink"><b><a href="#cite_ref-GatheralJacquierSSVI">^</a></b></span> <NAME> and <NAME>, Arbitrage-free SVI volatility
# surfaces, <span>*Quantitative Finance*</span> <span>**14**</span>(1) 59-71 (2014).</li>
# <li id="cite_note-GJR"><span class="mw-cite-backlink"><b><a href="#cite_ref-GJR">^</a></b></span> <NAME>, <NAME> and <NAME>, Volatility is rough, available at http://papers.ssrn.com/sol3/papers.cfm?abstract_id=2509457, (2014).</li>
# <li id="cite_note-GO"><span class="mw-cite-backlink"><b><a href="#cite_ref-GO">^</a></b></span> <NAME> and <NAME>, Zero-intelligence realized variance estimation, *Finance and Stochastics* **14**(2) 249-283 (2010).</li>
# <li id="cite_note-NuzmanPoor"><span class="mw-cite-backlink"><b><a href="#cite_ref-NuzmanPoor">^</a></b></span> <NAME> and <NAME>, Linear estimation of self-similar processes via Lamperti’s transformation, *Journal of Applied Probability* **37**(2) 429-452 (2000).</li>
# </ol>
# </div>
#
| src/.ipynb_checkpoints/RoughVolatilityFromPythonToJulia-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Готовим данные
import pandas as pd
df = pd.read_csv('02_titanic_train.csv')
df.head()
df.info()
df = df.drop(['Cabin', 'Name', 'Sex', 'Ticket', 'Embarked'], axis=1)
df = df[~df['Age'].isnull()]
df.shape
df.info()
y = df['Survived']
df = df.drop(['Survived'], axis=1)
# ## А где же машинное обучение?
from sklearn.tree import DecisionTreeClassifier
dtc = DecisionTreeClassifier()
dtc.fit(df, y)
predict = dtc.predict(df)
df.head()
predict
# Но как сравнить? Как понять хорошо мы отработали или не очень?
from sklearn.metrics import accuracy_score
accuracy_score(y, predict)
# 
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df,
y,
test_size=0.33,
random_state=42)
from sklearn.tree import DecisionTreeClassifier
dtc = DecisionTreeClassifier()
dtc.fit(X_train, y_train)
predict = dtc.predict(X_test)
accuracy_score(y_test, predict)
predict
# 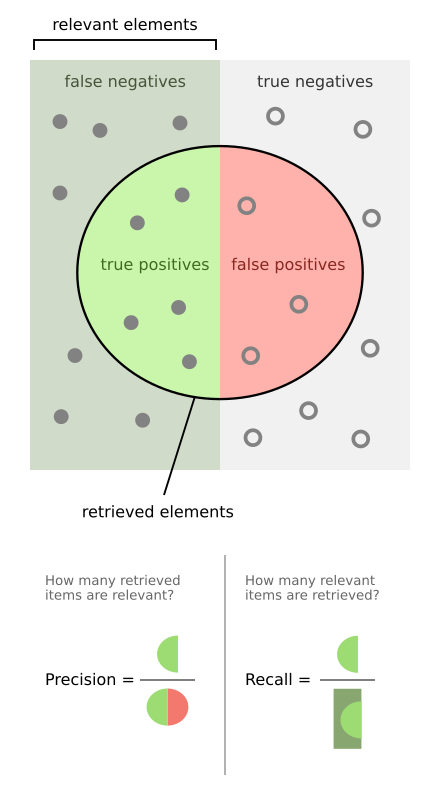
from sklearn.metrics import precision_score, recall_score
precision_score(y_test, predict)
recall_score(y_test, predict)
# +
## Так ли хорошо работает модель?
# -
from sklearn.cross_validation import train_test_split
# +
from sklearn.metrics import accuracy_score, f1_score
print('precision', precision_score(y_test, predict))
print('recall', recall_score(y_test, predict))
print('accuracy', accuracy_score(y_test, predict))
print('f1', f1_score(y_test, predict))
# +
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier()
rfc.fit(X_train, y_train)
predict = rfc.predict(X_test)
print('precision', precision_score(y_test, predict))
print('recall', recall_score(y_test, predict))
print('accuracy', accuracy_score(y_test, predict))
# -
rfc = RandomForestClassifier()
rfc.fit(X_train, y_train)
predict = rfc.predict_proba(X_test)
predict
predict[:,1]
from sklearn.metrics import precision_recall_curve
precision, recall, tresholds = precision_recall_curve(y_test, predict[:,1])
# %matplotlib inline
from matplotlib import pyplot as plt
from sklearn.metrics import roc_auc_score, roc_curve
fpr, tpr, thresholds = roc_curve(y_test, predict[:,1])
plt.figure(figsize=(5, 5))
plt.plot(fpr, tpr)
plt.ylabel('tpr')
plt.xlabel('fpr')
plt.grid(True)
plt.title('ROC curve')
plt.xlim((-0.01, 1.01))
plt.ylim((-0.01, 1.01))
# %matplotlib inline
from matplotlib import pyplot as plt
from sklearn.metrics import roc_auc_score, roc_curve
fpr, tpr, thresholds = roc_curve(y_test, predict[:,1])
plt.figure(figsize=(5, 5))
plt.plot(precision, recall)
plt.ylabel('recall')
plt.xlabel('precision')
plt.grid(True)
plt.title('PR curve')
plt.xlim((-0.01, 1.01))
plt.ylim((-0.01, 1.01))
precision_recall_curve(y_test, predict[:,1])roc_auc_score(y_test, predict[:,1])
# 
from sklearn.cross_validation import cross_val_score
rfc = RandomForestClassifier()
cross_val_score(
rfc,
df,
y,
scoring='roc_auc',
cv=10
)
# ## Вернемся к данным
df = pd.read_csv('02_titanic_train.csv')
df.head()
df.info()
df = df.drop(['Cabin', 'Name', 'Ticket', 'PassengerId'], axis=1)
df.head()
df = pd.get_dummies(df, columns=['Sex', 'Embarked'])
df.head()
df.info()
df.Age[df.Age.isnull()] = df.Age.mean()
df.info()
dtc = DecisionTreeClassifier()
cross_val_score(
dtc,
df,
df['Survived'],
scoring='roc_auc',
cv=10
)
| hw02/lesson_02/.ipynb_checkpoints/02_titanic-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
# +
#! ! pip install FPDF
# -
from fpdf import FPDF
from PIL import Image
# # deve renomear as imagens com mesmo nome
# # selecione tudo usando 'crtl + a', selecione renomear e digite um nome para padronizar todos os arquivos
# # exemplo para declarar o caminho do arquivos \\
imagem='C:\\Users\\dougl\\Desktop\\projetos\python\\fotos-20210624T144130Z-001\\sapato\\'
class Imagem ():
def localizar_imagens (nome_da_imagem,local_das_imagens, numero_de_imagens):
listaImagens = []
listaPages= []
for i in range(1,numero_de_imagens + 1):
#i.toString()
# Abra a imagem | Coloque o local da imagem!
imagem = Image.open(local_das_imagens + nome_da_imagem + str(i) + ').jpg')
listaImagens.append(imagem)
caminho = local_das_imagens + nome_da_imagem + str(i) + ').jpg'
listaPages.append(caminho)
return imagem , caminho
class Pdf():
def fabricar_PDF (batizar_pdf,imagem,caminho):
pdfFileName = 'catalogo de sapatos'
dir = ''
if (dir):
dir += "/"
cover = imagem
width, height = cover.size
pdf = FPDF(unit = "pt", format = [width, height])
for page in caminho:
pdf.add_page()
pdf.image(page, 0, 0)
return pdf.output(dir + pdfFileName + ".pdf", "F")
imagem=Imagem.localizar_imagens('sapatos',local_das_imagens , 147)
Pdf.fabricar_PDF('CATALOGO', imagem[0], imagem[1])
| ESTUDO DE POO/ESTUDO-POO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Implementation of Softmax Regression from Scratch
#
# :label:`chapter_softmax_scratch`
#
#
# Just as we implemented linear regression from scratch,
# we believe that multiclass logistic (softmax) regression
# is similarly fundamental and you ought to know
# the gory details of how to implement it from scratch.
# As with linear regression, after doing things by hand
# we will breeze through an implementation in Gluon for comparison.
# To begin, let's import our packages
# (only `autograd`, `nd` are needed here
# because we will be doing the heavy lifting ourselves.)
# + attributes={"classes": [], "id": "", "n": "1"}
import d2l
from mxnet import autograd, nd, gluon
from IPython import display
# -
# We will work with the Fashion-MNIST dataset just introduced,
# cuing up an iterator with batch size 256.
# + attributes={"classes": [], "id": "", "n": "2"}
batch_size = 256
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size)
# -
# ## Initialize Model Parameters
#
# Just as in linear regression, we represent each example as a vector.
# Since each example is a $28 \times 28$ image,
# we can flatten each example, treating them as $784$ dimensional vectors.
# In the future, we'll talk about more sophisticated strategies
# for exploiting the spatial structure in images,
# but for now we treat each pixel location as just another feature.
#
# Recall that in softmax regression,
# we have as many outputs as there are categories.
# Because our dataset has $10$ categories,
# our network will have an output dimension of $10$.
# Consequently, our weights will constitute a $784 \times 10$ matrix
# and the biases will constitute a $1 \times 10$ vector.
# As with linear regression, we will initialize our weights $W$
# with Gaussian noise and our biases to take the initial value $0$.
# + attributes={"classes": [], "id": "", "n": "3"}
num_inputs = 784
num_outputs = 10
W = nd.random.normal(scale=0.01, shape=(num_inputs, num_outputs))
b = nd.zeros(num_outputs)
# -
# Recall that we need to *attach gradients* to the model parameters.
# More literally, we are allocating memory for future gradients to be stored
# and notifiying MXNet that we want gradients to be calculated with respect to these parameters in the first place.
# + attributes={"classes": [], "id": "", "n": "4"}
W.attach_grad()
b.attach_grad()
# -
# ## The Softmax
#
# Before implementing the softmax regression model,
# let's briefly review how operators such as `sum` work
# along specific dimensions in an NDArray.
# Given a matrix `X` we can sum over all elements (default) or only
# over elements in the same column (`axis=0`) or the same row (`axis=1`).
# Note that if `X` is an array with shape `(2, 3)`
# and we sum over the columns (`X.sum(axis=0`),
# the result will be a (1D) vector with shape `(3,)`.
# If we want to keep the number of axes in the original array
# (resulting in a 2D array with shape `(1,3)`),
# rather than collapsing out the dimension that we summed over
# we can specify `keepdims=True` when invoking `sum`.
# + attributes={"classes": [], "id": "", "n": "5"}
X = nd.array([[1, 2, 3], [4, 5, 6]])
X.sum(axis=0, keepdims=True), X.sum(axis=1, keepdims=True)
# -
# We are now ready to implement the softmax function.
# Recall that softmax consists of two steps:
# First, we exponentiate each term (using `exp`).
# Then, we sum over each row (we have one row per example in the batch)
# to get the normalization constants for each example.
# Finally, we divide each row by its normalization constant,
# ensuring that the result sums to $1$.
# Before looking at the code, let's recall
# what this looks expressed as an equation:
#
# $$
# \mathrm{softmax}(\mathbf{X})_{ij} = \frac{\exp(X_{ij})}{\sum_k \exp(X_{ik})}
# $$
#
# The denominator, or normalization constant,
# is also sometimes called the partition function
# (and its logarithm the log-partition function).
# The origins of that name are in [statistical physics](https://en.wikipedia.org/wiki/Partition_function_(statistical_mechanics))
# where a related equation models the distribution
# over an ensemble of particles).
# + attributes={"classes": [], "id": "", "n": "6"}
def softmax(X):
X_exp = X.exp()
partition = X_exp.sum(axis=1, keepdims=True)
return X_exp / partition # The broadcast mechanism is applied here
# -
# As you can see, for any random input, we turn each element into a non-negative number. Moreover, each row sums up to 1, as is required for a probability.
# Note that while this looks correct mathematically,
# we were a bit sloppy in our implementation
# because failed to take precautions against numerical overflow or underflow
# due to large (or very small) elements of the matrix,
# as we did in
# :numref:`chapter_naive_bayes`.
# + attributes={"classes": [], "id": "", "n": "7"}
X = nd.random.normal(shape=(2, 5))
X_prob = softmax(X)
X_prob, X_prob.sum(axis=1)
# -
# ## The Model
#
# Now that we have defined the softmax operation,
# we can implement the softmax regression model.
# The below code defines the forward pass through the network.
# Note that we flatten each original image in the batch
# into a vector with length `num_inputs` with the `reshape` function
# before passing the data through our model.
# + attributes={"classes": [], "id": "", "n": "8"}
def net(X):
return softmax(nd.dot(X.reshape((-1, num_inputs)), W) + b)
# -
# ## The Loss Function
#
# Next, we need to implement the cross entropy loss function,
# introduced in :numref:`chapter_softmax`.
# This may be the most common loss function
# in all of deep learning because, at the moment,
# classification problems far outnumber regression problems.
#
#
# Recall that cross entropy takes the negative log likelihood
# of the predicted probability assigned to the true label $-\log p(y|x)$.
# Rather than iterating over the predictions with a Python `for` loop
# (which tends to be inefficient), we can use the `pick` function
# which allows us to select the appropriate terms
# from the matrix of softmax entries easily.
# Below, we illustrate the `pick` function on a toy example,
# with 3 categories and 2 examples.
# + attributes={"classes": [], "id": "", "n": "9"}
y_hat = nd.array([[0.1, 0.3, 0.6], [0.3, 0.2, 0.5]])
y = nd.array([0, 2], dtype='int32')
nd.pick(y_hat, y)
# -
# Now we can implement the cross-entropy loss function efficiently
# with just one line of code.
# + attributes={"classes": [], "id": "", "n": "10"}
def cross_entropy(y_hat, y):
return - nd.pick(y_hat, y).log()
# -
# ## Classification Accuracy
#
# Given the predicted probability distribution `y_hat`,
# we typically choose the class with highest predicted probability
# whenever we must output a *hard* prediction. Indeed, many applications require that we make a choice. Gmail must catetegorize an email into Primary, Social, Updates, or Forums. It might estimate probabilities internally, but at the end of the day it has to choose one among the categories.
#
# When predictions are consistent with the actual category `y`, they are coorect. The classification accuracy is the fraction of all predictions that are correct. Although we cannot optimize accuracy directly (it is not differentiable), it's often the performance metric that we care most about, and we will nearly always report it when training classifiers.
#
# To compute accuracy we do the following:
# First, we execute `y_hat.argmax(axis=1)`
# to gather the predicted classes
# (given by the indices for the largest entires each row).
# The result has the same shape as the variable `y`.
# Now we just need to check how frequently the two match.
# Since the equality operator `==` is datatype-sensitive
# (e.g. an `int` and a `float32` are never equal),
# we also need to convert both to the same type (we pick `float32`).
# The result is an NDArray containing entries of 0 (false) and 1 (true).
# Taking the mean yields the desired result.
# + attributes={"classes": [], "id": "", "n": "11"}
# Save to the d2l package.
def accuracy(y_hat, y):
return (y_hat.argmax(axis=1) == y.astype('float32')).sum().asscalar()
# -
# We will continue to use the variables `y_hat` and `y`
# defined in the `pick` function,
# as the predicted probability distribution and label, respectively.
# We can see that the first example's prediction category is 2
# (the largest element of the row is 0.6 with an index of 2),
# which is inconsistent with the actual label, 0.
# The second example's prediction category is 2
# (the largest element of the row is 0.5 with an index of 2),
# which is consistent with the actual label, 2.
# Therefore, the classification accuracy rate for these two examples is 0.5.
# + attributes={"classes": [], "id": "", "n": "12"}
accuracy(y_hat, y) / len(y)
# -
# Similarly, we can evaluate the accuracy for model `net` on the data set
# (accessed via `data_iter`).
# + attributes={"classes": [], "id": "", "n": "13"}
# Save to the d2l package.
def evaluate_accuracy(net, data_iter):
metric = Accumulator(2) # num_corrected_examples, num_examples
for X, y in data_iter:
y = y.astype('float32')
metric.add(accuracy(net(X), y), y.size)
return metric[0] / metric[1]
# -
# Here `Accumulator` is a utility class to accumulated sum over multiple numbers.
# Save to the d2l package.
class Accumulator(object):
"""Sum a list of numbers over time"""
def __init__(self, n):
self.data = [0.0] * n
def add(self, *args):
self.data = [a+b for a, b in zip(self.data, args)]
def reset(self):
self.data = [0] * len(self.data)
def __getitem__(self, i):
return self.data[i]
# Because we initialized the `net` model with random weights,
# the accuracy of this model should be close to random guessing,
# i.e. 0.1 for 10 classes.
# + attributes={"classes": [], "id": "", "n": "14"}
evaluate_accuracy(net, test_iter)
# -
# ## Model Training
#
# The training loop for softmax regression should look strikingly familiar
# if you read through our implementation
# of linear regression in :numref:`chapter_linear_scratch`. Here we refactor the implementation to make it reusable. First, we define a function to train for one data epoch. Note that `updater` is general function to update the model parameters, which accepts the batch size as an argument. It can be either a wrapper of `d2l.sgd` or a Gluon trainer.
# + attributes={"classes": [], "id": "", "n": "15"}
# Save to the d2l package.
def train_epoch_ch3(net, train_iter, loss, updater):
metric = Accumulator(3) # train_loss_sum, train_acc_sum, num_examples
if isinstance(updater, gluon.Trainer):
updater = updater.step
for X, y in train_iter:
# compute gradients and update parameters
with autograd.record():
y_hat = net(X)
l = loss(y_hat, y)
l.backward()
updater(X.shape[0])
metric.add(l.sum().asscalar(), accuracy(y_hat, y), y.size)
# Return training loss and training accuracy
return metric[0]/metric[2], metric[1]/metric[2]
# -
# Before showing the implementation of the training function, we define a utility class that draw data in animation. Again, it aims to simplify the codes in later chapters.
# + attributes={"classes": [], "id": "", "n": "16"}
# Save to the d2l package.
class Animator(object):
def __init__(self, xlabel=None, ylabel=None, legend=[], xlim=None,
ylim=None, xscale='linear', yscale='linear', fmts=None,
nrows=1, ncols=1, figsize=(3.5, 2.5)):
"""Incrementally plot multiple lines."""
d2l.use_svg_display()
self.fig, self.axes = d2l.plt.subplots(nrows, ncols, figsize=figsize)
if nrows * ncols == 1: self.axes = [self.axes,]
# use a lambda to capture arguments
self.config_axes = lambda : d2l.set_axes(
self.axes[0], xlabel, ylabel, xlim, ylim, xscale, yscale, legend)
self.X, self.Y, self.fmts = None, None, fmts
def add(self, x, y):
"""Add multiple data points into the figure."""
if not hasattr(y, "__len__"): y = [y]
n = len(y)
if not hasattr(x, "__len__"): x = [x] * n
if not self.X: self.X = [[] for _ in range(n)]
if not self.Y: self.Y = [[] for _ in range(n)]
if not self.fmts: self.fmts = ['-'] * n
for i, (a, b) in enumerate(zip(x, y)):
if a is not None and b is not None:
self.X[i].append(a)
self.Y[i].append(b)
self.axes[0].cla()
for x, y, fmt in zip(self.X, self.Y, self.fmts):
self.axes[0].plot(x, y, fmt)
self.config_axes()
display.display(self.fig)
display.clear_output(wait=True)
# -
# The training function then runs multiple epochs and visualize the training progress.
# + attributes={"classes": [], "id": "", "n": "17"}
# Save to the d2l package.
def train_ch3(net, train_iter, test_iter, loss, num_epochs, updater):
trains, test_accs = [], []
animator = Animator(xlabel='epoch', xlim=[1, num_epochs],
ylim=[0.3, 0.9],
legend=['train loss', 'train acc', 'test acc'])
for epoch in range(num_epochs):
train_metrics = train_epoch_ch3(net, train_iter, loss, updater)
test_acc = evaluate_accuracy(net, test_iter)
animator.add(epoch+1, train_metrics+(test_acc,))
# -
# Again, we use the mini-batch stochastic gradient descent
# to optimize the loss function of the model.
# Note that the number of epochs (`num_epochs`),
# and learning rate (`lr`) are both adjustable hyper-parameters.
# By changing their values, we may be able to increase the classification accuracy of the model. In practice we'll want to split our data three ways
# into training, validation, and test data, using the validation data to choose the best values of our hyperparameters.
# + attributes={"classes": [], "id": "", "n": "18"}
num_epochs, lr = 10, 0.1
updater = lambda batch_size: d2l.sgd([W, b], lr, batch_size)
train_ch3(net, train_iter, test_iter, cross_entropy, num_epochs, updater)
# -
# ## Prediction
#
# Now that training is complete, our model is ready to classify some images.
# Given a series of images, we will compare their actual labels
# (first line of text output) and the model predictions
# (second line of text output).
# + attributes={"classes": [], "id": "", "n": "19"}
# Save to the d2l package.
def predict_ch3(net, test_iter, n=6):
for X, y in test_iter:
break
trues = d2l.get_fashion_mnist_labels(y.asnumpy())
preds = d2l.get_fashion_mnist_labels(net(X).argmax(axis=1).asnumpy())
titles = [true+'\n'+ pred for true, pred in zip(trues, preds)]
d2l.show_images(X[0:n].reshape((n,28,28)), 1, n, titles=titles[0:n])
predict_ch3(net, test_iter)
# -
# ## Summary
#
# With softmax regression, we can train models for multi-category classification. The training loop is very similar to that in linear regression: retrieve and read data, define models and loss functions,
# then train models using optimization algorithms. As you'll soon find out, most common deep learning models have similar training procedures.
#
# ## Exercises
#
# 1. In this section, we directly implemented the softmax function based on the mathematical definition of the softmax operation. What problems might this cause (hint - try to calculate the size of $\exp(50)$)?
# 1. The function `cross_entropy` in this section is implemented according to the definition of the cross-entropy loss function. What could be the problem with this implementation (hint - consider the domain of the logarithm)?
# 1. What solutions you can think of to fix the two problems above?
# 1. Is it always a good idea to return the most likely label. E.g. would you do this for medical diagnosis?
# 1. Assume that we want to use softmax regression to predict the next word based on some features. What are some problems that might arise from a large vocabulary?
#
# ## Scan the QR Code to [Discuss](https://discuss.mxnet.io/t/2336)
#
# 
| d2l-en/chapter_linear-networks/softmax-regression-scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Algorithms and Data Structures #
# [Toward Data science article 1](https://towardsdatascience.com/a-data-scientists-guide-to-data-structures-algorithms-1176395015a0)
#
# [Toward Data science article 2](https://towardsdatascience.com/a-data-scientists-guide-to-data-structures-algorithms-part-2-6bc27066f3fe)
#
# 
# ## I. Search ##
# ### 1. Search ###
# - $O(n)$
# ### 2. Binary search
# - $Big O = O(\log{n})$
# ## II. Sort
# ### 1.Selection Sort
# - go through the list to find the laregst/smallert item, and add to a second list
# - go through the original list again to find the next largest/least, add
# - repeat until finish the whole list
# - $Big \mathcal{O} = O(n^2)$ as for each element, need to run through the whole list, thus $n \times n$
# ### 2. Quick Sort
# - get a pivot from current list (normally random selected), and partition elements smaller than it to the left, elements larger than it to the right
# - for each of the half list, treat it as anew list and rearrange using the same way
# - Quicksort is unique because its speed is dependent on the pivot selection. At worst, it can take O(n²) time, which is as slow as selection sort. However, if the pivot is always some random element in the list, quicksort runs in O(n log n) time on average.
# - $Big \mathcal{O} = O(n\log{n})$
#
# 
# ### 3. Merge Sort
# - break down the list into individual elements, and sort every pair
# - sort two pairs together, and get the pairs grow until the whole lists are sorted
# - Mergesort runs on $O(n \log n)$ time because the entire list is halved ($O(\log n)$) and this is done for n items.
#
# 
# ## III. Abstract Data Structures (ADT)##
# ### 1. Stack ###
# - first in, last out
# - push/pop
# ### 2. Queue ###
# - first in, first out (like waiting in the line of being seated at restaurants)
# - push/pop
# ### 3. Graph (network)
# - node and edges
# - directed graph(edges with arrows => one-way direction of the connection)
# - undirected graph(edges without arrows, thus two-way connections)
# - could have multiple egdes between nodes
# ### 4. Tree (hierarchy)
# - special type of graph, with "hierarchical" structure
# - with only one root, which is also the origin
# - only one path allowed between two nodes
#
#
# ## IIII. Concrete Data Structures (CDT) ##
# ### Array###
# An array is comprised of a linear collection of items (known as elements), stored contiguously in memory. Any given element can be accessed using a numerical index which points to the element’s location in the array. In Python native, this is implemented with the list type.
#
# ### Hash Tables###
# In a hash table, unique keys are mapped to values. In Python native, a dictionary is the implementation of a hash table.
# 
# note: learn BFS/DFS/Binary search tree
| Algorithm and Data Structure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import models
import matplotlib.pyplot as plt
import numpy as np
import xlrd
from sklearn.preprocessing import MinMaxScaler
from sklearn.svm import SVC
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import roc_curve, auc
import itertools
from sklearn.metrics import confusion_matrix
# load data
## Pathology for train
data = xlrd.open_workbook('Pathology.xlsx').sheets()[0]
X_tra = np.zeros((111, 1))
X_tra[:, 0] = data.col_values(3) # X0 = Integrated backscatter
#X_tra[:, 1] = data.col_values(4) # X1 = Q factor (HHT)
#X_tra[:, 2] = data.col_values(6) # X2 = Homogeneity factor
### scaling to 0~1
Scaler = MinMaxScaler(copy=True, feature_range=(0, 1)).fit(X_tra)
X_train = Scaler.transform(X_tra)
### label
nrows = data.nrows
y_train = []
for i in range(nrows): # y = 'T' or 'F' from Pathology data
if data.cell(i,1).value < 10:
y_train.append(0)
elif data.cell(i,1).value >= 10:
y_train.append(1)
y_train = np.ravel(y_train) # Return a contiguous flattened array
class_names = np.ravel(['negative', 'positive'])
## New Pathology for test
data = xlrd.open_workbook('NPathology.xlsx').sheets()[0]
X_te = np.zeros((74, 1))
X_te[:, 0] = data.col_values(3) # X0 = Integrated backscatter
#X_te[:, 1] = data.col_values(4) # X1 = Q factor (HHT)
#X_te[:, 2] = data.col_values(6) # X2 = Homogeneity factor
### scaling to 0~1
X_test = Scaler.transform(X_te)
### label
nrows = data.nrows
y_test = []
for i in range(nrows): # y = 'T' or 'F' from Pathology data
if data.cell(i,1).value < 10:
y_test.append(0)
elif data.cell(i,1).value >= 10:
y_test.append(1)
y_test = np.ravel(y_test) # Return a contiguous flattened array
# training & validation
## set CV
cv = StratifiedKFold(10)
## set parameters
C_l = list(2**(i/100) for i in range(-300, 125, 25))
C_r = list(2**(i/100) for i in range(1100, 1525, 25))
C_p = list(2**(i/100) for i in range(-500, -75, 25))
gamma_r = list(2**(i/100) for i in range(-500, -75, 25))
gamma_p = list(2**(i/100) for i in range(100, 525, 25))
## SVC_linear
linear_prarmeters = {'kernel': ['linear'], 'C': C_l}
clf_linear = GridSearchCV(SVC(), linear_prarmeters, cv=cv, n_jobs=3).fit(X_train, y_train)
print('The best parameters of linear are {0}\n with a validation score of {1:.2%}'
.format(clf_linear.best_params_, clf_linear.best_score_))
## SVC_rbf
rbf_prarmeters = {'kernel': ['rbf'], 'C': C_r, 'gamma': gamma_r}
clf_rbf = GridSearchCV(SVC(), rbf_prarmeters, cv=cv, n_jobs=3).fit(X_train, y_train)
print('The best parameters of rbf are {0}\n with a validation score of {1:.2%}'
.format(clf_rbf.best_params_, clf_rbf.best_score_))
## SVC_poly
poly_prarmeters = {'kernel': ['poly'], 'C': C_p, 'gamma': gamma_p, 'degree': [1, 2, 3]}
clf_poly = GridSearchCV(SVC(), poly_prarmeters, cv=cv, n_jobs=3).fit(X_train, y_train)
print('The best parameters of poly are {0}\n with a validation score of {1:.2%}\n'
.format(clf_poly.best_params_, clf_poly.best_score_))
# test
## SVC_linear
test_predict_linear = clf_linear.predict(X_test)
test_score_linear = clf_linear.score(X_test, y_test)
print('test accuracy of linear = {0:.2%}'.format(test_score_linear))
dec_linear = clf_linear.decision_function(X_test)
## SVC_rbf
test_predict_rbf = clf_rbf.predict(X_test)
test_score_rbf = clf_rbf.score(X_test, y_test)
print('test accuracy of rbf = {0:.2%}'.format(test_score_rbf))
dec_rbf = clf_rbf.decision_function(X_test)
## SVC_poly
test_predict_poly = clf_poly.predict(X_test)
test_score_poly = clf_poly.score(X_test, y_test)
print('test accuracy of poly = {0:.2%}\n'.format(test_score_poly))
dec_poly = clf_poly.decision_function(X_test)
## Confusion matrix
### define confusion matrix
def plot_confusion_matrix(cm, classes, normalize=False, title='Confusion matrix', cmap=plt.cm.Blues):
global i
print('{0}'.format(kernel[i]))
print('Sensitivity = {0:.2%}'.format(cm[1,1]/(cm[1,0]+cm[1,1])))
print('Specificity = {0:.2%}'.format(cm[0,0]/(cm[0,0]+cm[0,1])))
print('LR+ = {0:.2f}'.format((cm[1,1]/(cm[1,0]+cm[1,1]))/(cm[0,1]/(cm[0,0]+cm[0,1]))))
print('LR- = {0:.2f}'.format((cm[1,0]/(cm[1,0]+cm[1,1]))/(cm[0,0]/(cm[0,0]+cm[0,1]))))
print('PPV = {0:.2%}'.format(cm[1,1]/(cm[0,1]+cm[1,1])))
print('NPV = {0:.2%}'.format(cm[0,0]/(cm[0,0]+cm[1,0])))
print('Accuracy = {0:.2%}\n'.format((cm[0,0]+cm[1,1])/(cm[0,0]+cm[0,1]+cm[1,0]+cm[1,1])))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment='center',
color='white' if cm[i, j] > thresh else 'black')
plt.tight_layout()
plt.title(title, size = 20)
plt.ylabel('True label', size = 15)
plt.xlabel('Predicted label', size = 15)
kernel = ['linear', 'rbf', 'poly']
pre_kernel = [test_predict_linear, test_predict_rbf, test_predict_poly] # predict of each kernel
for i in range(0,3):
## Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, pre_kernel[i])
np.set_printoptions(precision=4)
## Plot non-normalized confusion matrix
plt.figure(figsize=(6,6))
plot_confusion_matrix(cnf_matrix, classes=class_names,
title='{0} Confusion matrix'.format(kernel[i]))
plt.show()
## ROC curve
### compute ROC and AUC
fpr = dict()
tpr = dict()
thresholds = dict()
t_score = dict()
roc_auc = dict()
dec = [dec_linear, dec_rbf, dec_poly] # direction of each kernel
for i in range(0,3):
fpr[i], tpr[i], thresholds[i] = roc_curve(y_test, dec[i])
roc_auc[i] = auc(fpr[i], tpr[i])
t_score[i] = tpr[i] - fpr[i]
for j in range(len(t_score[i])):
if t_score[i][j] == max(t_score[i]):
s = j
break
print('{0} Cut-off value = {1:.4f}'.format(kernel[i], thresholds[i][s]))
print('{0} AUROC = {1:.4f}'.format(kernel[i], roc_auc[i]))
### plot
plt.figure(figsize=(10,10))
lw=2
colors = ['red', 'green', 'blue']
linestyles = ['-', '-.', ':']
for i, color, linestyle in zip(range(0,3), colors, linestyles):
plt.plot(fpr[i], tpr[i], color=color, linestyle=linestyle, lw=lw,
label='ROC curve of {0} (auc = {1:.4f})'.format(kernel[i], roc_auc[i]))
plt.plot([0, 1], [0, 1], color='orange', lw=lw, linestyle='--')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate (1-specificity)', size = 20)
plt.ylabel('True Positive Rate (sensitivity)', size = 20)
plt.title('Receiver operating characteristic', size = 25)
plt.legend(loc='lower right')
plt.show()
# -
| SVM/10Fold for Pathology _ NPathology/'1 Integrated backscatter (IB).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="yWWwioF0h_u9" colab={"base_uri": "https://localhost:8080/"} outputId="2b61c21f-51bc-4c26-f720-8a6f1e86931a"
# !pip install transformers datasets jsonlines folium==0.2.1
# + [markdown] id="9Ey8W8E-iCTi"
# # <h1><center><b>Enkóderek</b></h1></center>
# + [markdown] id="sL1uU0qhkB18"
# ## **Tokenizálás**
#
# * Karakteralapú: túl hosszúak a szekvenciák
# * Szóalapú: túl nagy a szótár
# * Megoldás: **subword**
# * Algoritmusok: Wordpiece, Byte Pair Encoding, Sentencepiece stb.
#
# <br/>
#
# <center><img src="https://drive.google.com/uc?export=view&id=1x-RgrQW3CaOu_HItCYGPQ0Wgs0a7CcFZ"/><p>Forrás: Devlin et al. (2019) </P></center>
# + colab={"base_uri": "https://localhost:8080/", "height": 162, "referenced_widgets": ["8da95453faa146e799be629dba5957cc", "58a2933bfa5a4f909e50439e25ff0f5e", "d010bb3257fd4e3695b56ef670546503", "2741599eaf314411b1ad24ee30bf3142", "<KEY>", "<KEY>", "5607792a246344f8be9ebea8f3433273", "8377d06b2a3d4802827279034d2dd8f3", "a8cd0b6209694c65b028eafa727caa58", "96180f30f9e64b6ebc50bd9024ab6e4d", "<KEY>", "<KEY>", "685c4140b67a4cd2a81c30ce8a3c9b87", "<KEY>", "1ed1ec23a3d342bc99af8a1a719715cd", "<KEY>", "<KEY>", "b93e6c26b5ea4d489e8e56f63d543023", "e2a82de88a3d43de9b4f8f6c40a3d9ed", "30618d1db8ce45f6aadba2fc438283ba", "c6555794cbbe4e188466ade9554eac74", "<KEY>", "<KEY>", "<KEY>", "6ca710913bab4a3ebb38d56f0792e05b", "<KEY>", "58e67070e103469ab1abc756d4a0caa0", "e853624c86384aafb6eed21ed6b87d09", "<KEY>", "cb19b5d814e74d4aabba886fea225930", "f95372435d2e4c2185a0a962c15d4e4c", "<KEY>", "46e6f183e69c4018abf3ea060482c35d", "<KEY>", "<KEY>", "e1b9e5f0487d4f18b500a685091127b0", "<KEY>", "<KEY>", "<KEY>", "30061ae3ca784736a5a1df1a86587434", "66483ba8959d4fa287e518d3862015bb", "2174a07de107458ab0d0563fab2e6b9b", "42f8fa04c7c340608274eb36b67f3a87", "764ba6b6f689477bb58aea2c5c68ce27"]} id="-FMDLstJmGDC" outputId="0f826562-f8d4-4500-f08f-8df98b29bfcf"
from transformers import BertTokenizer
hu_tokenizer = BertTokenizer.from_pretrained("SZTAKI-HLT/hubert-base-cc")
hu_tokenizer.save_vocabulary("hubert_vocab.txt")
# + colab={"base_uri": "https://localhost:8080/"} id="I_ALCKv_ocVu" outputId="f7cfea53-0493-4c95-c3b3-3c0a4b068beb"
# !shuf -n 30 hubert_vocab.txt
# + [markdown] id="XBZ-icZQjqJt"
# ## **Transfer learning az NLP-ben: előnyök és hátrányok**
#
# A _transfer learning_ egy tanítási stratégia, mely a következő lépéseket feltételezi:
#
# * Egy modell paramétereinek véletlenszerű inicializálása
# * Előtanítás: a modell tanítása gradiensereszkedéssel egy olyan feladaton, melyre nagymennyiségű tanítóanyag érhető el
# * Finomhangolás: az előtanított modell továbbtanítása egy célfeladaton
#
# **Előnyök:**
# * Az előtanított modell paraméterei több finomhangoláshoz is felhasználhatóak
# * A finomhangolás kevesebb tanítóanyagot igényel, mint a célfeladaton történő közvetlen (előtanítás nélküli) tanítás
# * A finomhangolt modell jól teljesít azon a célfeladaton, amelyen tanult a finomhangolás során
#
# **Hátrányok**:
# * Az előtanítás sok időt és energiát vesz igénybe
# * Az előtanítás eredményessége nehezen mérhető
# * Az előtanított modell még nem képes kezelni a legtöbb célfeladatot
# * A finomhangolás célfeladattól függő tervezést és adatokat igényel
# * Egy finomhangolt modell csak egy adott célfeladatot képes ellátni
# + [markdown] id="covxy-dSeUow"
# <center><img src="https://drive.google.com/uc?export=view&id=19Wl3YTCMv2B9ZdNieC3vjD0WETJ3892q"/><p>Forrás: Face Recognition Using Transfer Learning </P></center>
# + [markdown] id="qMZrWNMESGdL"
# ## **A BERT modellek: az előtanítás**
#
# * Tanítóanyag előkészítése
# * Tokenizálás
# * Tanítás
# + [markdown] id="yrLReDrZjhJH"
# <center><img src="https://drive.google.com/uc?export=view&id=1wyeMHbbwm4DtbsfEz7VURrkHBYtV-nne"/><p>Forrás: Devlin et al. (2019) </P></center>
#
# + colab={"base_uri": "https://localhost:8080/"} id="SE5N_3ubq4lV" outputId="68462d5d-4371-450c-e9e1-525a5ce8300a"
# !wget "https://nessie.ilab.sztaki.hu/~ndavid/Webcorpus2_text/2017_2018_0001.txt.gz"
# !gzip -d "2017_2018_0001.txt.gz"
# + id="mPoSIAzSrhOs"
# !head -n 40 "2017_2018_0001.txt"
# + id="3VLp6o-_yHFO" colab={"base_uri": "https://localhost:8080/"} outputId="4bbccaba-aa4e-4d1e-e88d-0e224c363861"
# !paste - - -d$"\t" <2017_2018_0001.txt >pairs.txt
# !wc -l pairs.txt
# + id="NhPza_QfzWBz"
from typing import Iterable, List, Tuple, Sequence, Dict, Any, Optional
from random import choices, shuffle
from itertools import tee, islice
import jsonlines
# + id="8hJnxwshznb-"
def get_nsp_pairs(
lines: Iterable[str],
num_lines: int,
sep: str = "\t"
) -> Tuple[List[str], List[str]]:
nopair_indices = set(choices(
range(num_lines), k=num_lines // 2))
pairs = []
nopairs = []
for i, sents in enumerate(map(lambda x: x.strip().split(sep), lines)):
if len(sents) == 2:
if "" in sents:
nopairs.append("".join(sents))
elif i in nopair_indices:
for sent in sents:
nopairs.append(sent)
else:
pairs.append(tuple(sents))
shuffle(nopairs)
nopairs = list(zip(*(islice(it, i, None, 2)
for i, it in enumerate(tee(nopairs, 2)))))
return pairs, nopairs
def write_dataset(
pair_sents: List[Tuple[str, str]],
nopair_sents: List[Tuple[str, str]],
out_path: str
) -> None:
field1, field2 = ("sentence1", "sentence2")
label_field = "next_sentence_label"
with jsonlines.open(out_path, "w") as f:
for data_source, label in zip((pair_sents, nopair_sents), (1, 0)):
for sent1, sent2 in data_source:
f.write({field1: sent1, field2: sent2, label_field: label})
# + id="a3IRG5tRtzMf"
with open("pairs.txt", "r", encoding="utf-8") as pairs_f:
pair_sents, nopair_sents = get_nsp_pairs(pairs_f, 44610)
write_dataset(pair_sents, nopair_sents, out_path="dataset.jsonl")
del pair_sents, nopair_sents
# + id="twDH3xqLuphh"
# !shuf dataset.jsonl >dataset_shuf.jsonl
# !head -n 10 dataset_shuf.jsonl
# + id="BarDEqYkxk3d"
from transformers import (
BatchEncoding,
BertForPreTraining,
TrainingArguments,
Trainer,
DataCollatorForLanguageModeling,
BertConfig
)
from datasets import Dataset, load_dataset
def tokenize_dataset(
dataset: Dataset,
tokenizer: BertTokenizer,
text_col_names: Sequence[str],
batch_size: int,
max_seq_length: Optional[int] = None,
label_col_name: Optional[str] = None
) -> Dataset:
training_keys = list(tokenizer("Dummy", "input").keys())
if label_col_name is not None:
training_keys.append(label_col_name)
if max_seq_length is not None:
tokenizer.model_max_length = max_seq_length
def tok_func(example: Dict[str, Any]) -> BatchEncoding:
return tokenizer(
*(example[text_col_name] for text_col_name in text_col_names),
padding=True,
truncation=True,
verbose=False
)
dataset = dataset.map(tok_func, batched=True, batch_size=batch_size)
dataset.set_format("torch", columns=training_keys)
return dataset
def pre_train_bert(
dataset: Dataset,
tokenizer: BertTokenizer,
label_name: str,
output_dir: str,
batch_size: int,
num_epochs: int,
learning_rate: float
) -> None:
model = BertForPreTraining(
config=BertConfig(vocab_size=tokenizer.vocab_size))
data_collator = DataCollatorForLanguageModeling(
tokenizer=tokenizer,
mlm=True,
mlm_probability=0.15
)
training_args = TrainingArguments(
output_dir=output_dir,
overwrite_output_dir=True,
learning_rate=learning_rate,
label_names=[label_name, "labels"],
per_device_train_batch_size=batch_size,
num_train_epochs=num_epochs
)
trainer = Trainer(
model=model,
train_dataset=dataset,
args=training_args,
data_collator=data_collator,
)
trainer.train()
trainer.save_model(output_dir)
# + id="UtpVtwydy6br" colab={"base_uri": "https://localhost:8080/"} outputId="47b64c28-de40-4b25-fc8a-c7b3f3e097a0"
dataset = load_dataset("json", split="train", data_files=["dataset_shuf.jsonl"])
BATCH_SIZE = 8
NSP_LABEL = "next_sentence_label"
MAX_SEQ_LENGTH = 128
dataset = tokenize_dataset(
dataset=dataset,
tokenizer=hu_tokenizer,
text_col_names=("sentence1", "sentence2"),
batch_size=BATCH_SIZE,
max_seq_length=MAX_SEQ_LENGTH,
label_col_name=NSP_LABEL
)
# + id="9a4N6kAm1qlW" colab={"base_uri": "https://localhost:8080/"} outputId="e30fa8ea-dce4-4ad7-e0c5-5daab4113b30"
# !rm -r sample_data && mkdir checkpoints
# + colab={"base_uri": "https://localhost:8080/"} id="f_sttB_A6nld" outputId="107cf27c-0157-4cc2-deea-409f5e9b0629"
print(dataset[0])
# + id="pDAmstJg5GZX"
pre_train_bert(
dataset=dataset,
tokenizer=hu_tokenizer,
label_name=NSP_LABEL,
output_dir="checkpoints",
batch_size=BATCH_SIZE,
num_epochs=1,
learning_rate=1e-7
)
# + [markdown] id="SV7cwrKRSo-5"
# ## **Finomhangolás szentimentanalízisre**
# + id="wh64KALJFEt6"
import torch
from torch.utils.data import DataLoader
from transformers import BertForSequenceClassification
from tqdm import tqdm
def fine_tune_bert(
model: BertForSequenceClassification,
data_loader: DataLoader,
num_epochs: int,
learning_rate: float = 1e-7,
logging_freq: int = 20
) -> BertForSequenceClassification:
"""Original source: https://huggingface.co/docs/datasets/quickstart"""
device = torch.device("cuda:0") if torch.cuda.is_available() \
else torch.device("cpu")
model.to(device)
model.train()
optimizer = torch.optim.AdamW(params=model.parameters(), lr=learning_rate)
for epoch in range(num_epochs):
for i, batch in enumerate(tqdm(data_loader)):
batch = {k: v.to(device) for k, v in batch.items()}
outputs = model(**batch)
loss = outputs[0]
loss.backward()
optimizer.step()
optimizer.zero_grad()
if i % logging_freq == 0:
print(f"Loss at step {i}, epoch {epoch}: {loss}")
return model
# + colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["ddcdec3c3e95401eb94c45da68d308ee", "cb6f206697124cb1acc6cff56a064bb8", "aa3db625eb414727a562eb2dc8c8b5c2", "47c84b06d3674c23b0f6ba913cd2060d", "b5287db89b8247ceba1d6887d075dcf7", "71fbf74805514c39ae28bef773b2aaae", "ca2dda33a0b74e16b4977179bca6452d", "b7e7b54bf3744ef18b3e3960d96f0013", "db8701d3f0b8469a97e1f41c4f890117", "30d67c3c76364a6185fec81f0da8b2b8", "34be657edad34ef6928147eec9525811"]} id="6VlsUkm8HJBR" outputId="715c8cac-9040-401b-c66d-9076c52088be"
tokenizer = BertTokenizer.from_pretrained("bert-base-uncased")
dataset = load_dataset("glue", "sst2", split="train")
dataset = tokenize_dataset(
dataset=dataset,
tokenizer=tokenizer,
text_col_names=("sentence",),
batch_size=BATCH_SIZE,
max_seq_length=MAX_SEQ_LENGTH,
label_col_name="label"
)
dataset = dataset.rename_column("label", "labels")
dataset = dataset.remove_columns(["sentence", "idx"])
# + colab={"base_uri": "https://localhost:8080/"} id="NtcGY-RvJJJQ" outputId="d93e9f2d-1502-4e81-c8af-01c9f3334248"
print(dataset[0])
# + id="q-pb06rcJRLY"
data_loader = DataLoader(dataset, batch_size=BATCH_SIZE)
model = BertForSequenceClassification.from_pretrained("bert-base-uncased")
_ = fine_tune_bert(model, data_loader, num_epochs=2)
# + [markdown] id="2U5vDreSpIkT"
# ## **Sentence-BERT tanítása NLI dataseten**
#
# * Mondatbeágyazások tanítása sziámi modellekkel
# * Jól tanítható NLI-n (Natural Language Inference)
# * [Dokumentáció, tananyagok](https://www.sbert.net/examples/training/nli/README.html)
#
# <br/>
#
# <center><img src="https://drive.google.com/uc?export=view&id=1-C8H-ExrRKcJfJQp1WaAOtCu7s58futj"/><p>Forrás: Reimers and Gurevych (2019)</p></center>
#
# + [markdown] id="rLA4DzLceGM5"
# ## **Cikkek, linkek**
#
# Tokenizálás:
# * Schuster et al. (2012): _Japanese and Korean Voice Search_ [Link](https://www.semanticscholar.org/paper/Japanese-and-Korean-voice-search-Schuster-Nakajima/ed6262b569c0a62c51d941228c54f34e563af022)
# * Sennrich et al. (2015): _Neural Machine Translation of Rare Words with Subword Units_ [Link](https://arxiv.org/abs/1508.07909)
# * Kudo and Richardson (2018): _SentencePiece: A simple and language independent subword tokenizer and detokenizer for Neural Text Processing_ [Link](https://arxiv.org/abs/1808.06226)
# * _Summary of the tokenizers_ (Hugging Face) [Link](https://huggingface.co/docs/transformers/tokenizer_summary)
#
# Transfer learning:
# * _Face Recognition Using Transfer Learning_ (Medium, 2020) [Link](https://medium.com/@agarwallashivam/face-recognition-using-transfer-learning-b8e36e6a15f1)
#
# BERT:
# * Devlin et al. (2019): _BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding_ [Link](https://arxiv.org/abs/1810.04805)
#
# Sentence-BERT:
# * Reimers and Gurevych (2019): _Sentence-BERT: Sentence Embeddings using Siamese BERT-Networks_ [Link](https://arxiv.org/abs/1810.04805)
#
# Mondatbeágyazások kiértékelése:
# * Perone et al. (2018): _Evaluation of sentence embeddings in downstream and linguistic probing tasks_ [Link](https://arxiv.org/abs/1806.06259)
| materials/lesson2/Encoders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="6x1ypzczQCwy"
# # TFX Pipeline Tutorial using Penguin dataset
#
# ## Learning objectives
# 1. Prepare example data.
# 2. Create a pipeline.
# 3. Run the pipeline.
# + [markdown] id="_VuwrlnvQJ5k"
# ## Introduction
# In this notebook, you will create and run a TFX pipeline
# for a simple classification model.
# The pipeline will consist of three essential TFX components: ExampleGen,
# Trainer and Pusher. The pipeline includes the most minimal ML workflow like
# importing data, training a model and exporting the trained model.
#
# Please see
# [Understanding TFX Pipelines](https://www.tensorflow.org/tfx/guide/understanding_tfx_pipelines)
# to learn more about various concepts in TFX.
#
# Each learning objective will correspond to a __#TODO__ in the [student lab notebook](../labs/penguin_simple.ipynb) -- try to complete that notebook first before reviewing this solution notebook.
# + [markdown] id="Fmgi8ZvQkScg"
# ## Set Up
# You first need to install the TFX Python package and download
# the dataset which you will use for our model.
#
# ### Upgrade Pip
#
# To avoid upgrading Pip in a system when running locally,
# check to make sure that you are running in Colab.
# Local systems can of course be upgraded separately.
# + colab={"base_uri": "https://localhost:8080/"} id="as4OTe2ukSqm" outputId="6660a20a-3e5c-42a5-9d12-55392589885a"
try:
import colab
# !pip install --upgrade pip
except:
pass
# + [markdown] id="MZOYTt1RW4TK"
# ### Install TFX
#
# + colab={"base_uri": "https://localhost:8080/"} id="iyQtljP-qPHY" outputId="55a57bee-74d4-4112-8885-a2b5ec5e74fb"
# !pip install -U tfx
# + [markdown] id="EwT0nov5QO1M"
# **Note:** Restart the kernel before proceeding further (On the Notebook menu, click **Kernel** > **Restart Kernel**).
# + [markdown] id="BDnPgN8UJtzN"
# Check the TensorFlow and TFX versions.
# + colab={"base_uri": "https://localhost:8080/"} id="6jh7vKSRqPHb" outputId="07bb86f7-2628-4fb7-a92e-3aff49d20130"
import tensorflow as tf
print('TensorFlow version: {}'.format(tf.__version__))
from tfx import v1 as tfx
print('TFX version: {}'.format(tfx.__version__))
# + [markdown] id="aDtLdSkvqPHe"
# ### Set up variables
#
# There are some variables used to define a pipeline. You can customize these
# variables as you want. By default all output from the pipeline will be
# generated under the current directory.
# + id="EcUseqJaE2XN"
import os
PIPELINE_NAME = "penguin-simple"
# Output directory to store artifacts generated from the pipeline.
PIPELINE_ROOT = os.path.join('pipelines', PIPELINE_NAME)
# Path to a SQLite DB file to use as an MLMD storage.
METADATA_PATH = os.path.join('metadata', PIPELINE_NAME, 'metadata.db')
# Output directory where created models from the pipeline will be exported.
SERVING_MODEL_DIR = os.path.join('serving_model', PIPELINE_NAME)
from absl import logging
logging.set_verbosity(logging.INFO) # Set default logging level.
# + [markdown] id="8F2SRwRLSYGa"
# ### Prepare example data
# You will download the example dataset for use in our TFX pipeline. The dataset we
# are using is
# [Palmer Penguins dataset](https://allisonhorst.github.io/palmerpenguins/articles/intro.html)
# which is also used in other
# [TFX examples](https://github.com/tensorflow/tfx/tree/master/tfx/examples/penguin).
#
# There are four numeric features in this dataset:
#
# - culmen_length_mm
# - culmen_depth_mm
# - flipper_length_mm
# - body_mass_g
#
# All features were already normalized to have range [0,1]. you will build a
# classification model which predicts the `species` of penguins.
# + [markdown] id="11J7XiCq6AFP"
# Because TFX ExampleGen reads inputs from a directory, you need to create a
# directory and copy dataset to it.
# + colab={"base_uri": "https://localhost:8080/"} id="4fxMs6u86acP" outputId="ab0fae39-fee3-42d4-e49f-5004ce50bde5"
import urllib.request
import tempfile
DATA_ROOT = tempfile.mkdtemp(prefix='tfx-data') # Create a temporary directory.
_data_url = 'https://raw.githubusercontent.com/tensorflow/tfx/master/tfx/examples/penguin/data/labelled/penguins_processed.csv'
_data_filepath = os.path.join(DATA_ROOT, "data.csv")
urllib.request.urlretrieve(_data_url, _data_filepath)
# + [markdown] id="ASpoNmxKSQjI"
# Take a quick look at the CSV file.
# + colab={"base_uri": "https://localhost:8080/"} id="-eSz28UDSnlG" outputId="06eb0734-9d39-4dd8-deba-25394984dde5"
# TODO 1
# Review the contents of the CSV file
# !head {_data_filepath}
# + [markdown] id="OTtQNq1DdVvG"
# You should be able to see five values. `species` is one of 0, 1 or 2, and all
# other features should have values between 0 and 1.
# + [markdown] id="nH6gizcpSwWV"
# ## Create a pipeline
#
# TFX pipelines are defined using Python APIs. You will define a pipeline which
# consists of following three components.
# - CsvExampleGen: Reads in data files and convert them to TFX internal format
# for further processing. There are multiple
# [ExampleGen](https://www.tensorflow.org/tfx/guide/examplegen)s for various
# formats. In this tutorial, you will use CsvExampleGen which takes CSV file input.
# - Trainer: Trains an ML model.
# [Trainer component](https://www.tensorflow.org/tfx/guide/trainer) requires a
# model definition code from users. You can use TensorFlow APIs to specify how to
# train a model and save it in a _saved_model_ format.
# - Pusher: Copies the trained model outside of the TFX pipeline.
# [Pusher component](https://www.tensorflow.org/tfx/guide/pusher) can be thought
# of an deployment process of the trained ML model.
#
# Before actually define the pipeline, you need to write a model code for the
# Trainer component first.
# + [markdown] id="lOjDv93eS5xV"
# ### Write model training code
#
# You will create a simple DNN model for classification using TensorFlow Keras
# API. This model training code will be saved to a separate file.
#
# In this tutorial you will use
# [Generic Trainer](https://www.tensorflow.org/tfx/guide/trainer#generic_trainer)
# of TFX which support Keras-based models. You need to write a Python file
# containing `run_fn` function, which is the entrypoint for the `Trainer`
# component.
# + id="aES7Hv5QTDK3"
_trainer_module_file = 'penguin_trainer.py'
# + colab={"base_uri": "https://localhost:8080/"} id="Gnc67uQNTDfW" outputId="bc9cebbb-645e-465f-cd67-d84ac5310c8c"
# %%writefile {_trainer_module_file}
from typing import List
from absl import logging
import tensorflow as tf
from tensorflow import keras
from tensorflow_transform.tf_metadata import schema_utils
from tfx import v1 as tfx
from tfx_bsl.public import tfxio
from tensorflow_metadata.proto.v0 import schema_pb2
_FEATURE_KEYS = [
'culmen_length_mm', 'culmen_depth_mm', 'flipper_length_mm', 'body_mass_g'
]
_LABEL_KEY = 'species'
_TRAIN_BATCH_SIZE = 20
_EVAL_BATCH_SIZE = 10
# Since we're not generating or creating a schema, you will instead create
# a feature spec. Since there are a fairly small number of features this is
# manageable for this dataset.
_FEATURE_SPEC = {
**{
feature: tf.io.FixedLenFeature(shape=[1], dtype=tf.float32)
for feature in _FEATURE_KEYS
},
_LABEL_KEY: tf.io.FixedLenFeature(shape=[1], dtype=tf.int64)
}
def _input_fn(file_pattern: List[str],
data_accessor: tfx.components.DataAccessor,
schema: schema_pb2.Schema,
batch_size: int = 200) -> tf.data.Dataset:
"""Generates features and label for training.
Args:
file_pattern: List of paths or patterns of input tfrecord files.
data_accessor: DataAccessor for converting input to RecordBatch.
schema: schema of the input data.
batch_size: representing the number of consecutive elements of returned
dataset to combine in a single batch
Returns:
A dataset that contains (features, indices) tuple where features is a
dictionary of Tensors, and indices is a single Tensor of label indices.
"""
return data_accessor.tf_dataset_factory(
file_pattern,
tfxio.TensorFlowDatasetOptions(
batch_size=batch_size, label_key=_LABEL_KEY),
schema=schema).repeat()
def _build_keras_model() -> tf.keras.Model:
"""Creates a DNN Keras model for classifying penguin data.
Returns:
A Keras Model.
"""
# The model below is built with Functional API, please refer to
# https://www.tensorflow.org/guide/keras/overview for all API options.
inputs = [keras.layers.Input(shape=(1,), name=f) for f in _FEATURE_KEYS]
d = keras.layers.concatenate(inputs)
for _ in range(2):
d = keras.layers.Dense(8, activation='relu')(d)
outputs = keras.layers.Dense(3)(d)
model = keras.Model(inputs=inputs, outputs=outputs)
model.compile(
optimizer=keras.optimizers.Adam(1e-2),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[keras.metrics.SparseCategoricalAccuracy()])
model.summary(print_fn=logging.info)
return model
# TFX Trainer will call this function.
def run_fn(fn_args: tfx.components.FnArgs):
"""Train the model based on given args.
Args:
fn_args: Holds args used to train the model as name/value pairs.
"""
# This schema is usually either an output of SchemaGen or a manually-curated
# version provided by pipeline author. A schema can also derived from TFT
# graph if a Transform component is used. In the case when either is missing,
# `schema_from_feature_spec` could be used to generate schema from very simple
# feature_spec, but the schema returned would be very primitive.
schema = schema_utils.schema_from_feature_spec(_FEATURE_SPEC)
train_dataset = _input_fn(
fn_args.train_files,
fn_args.data_accessor,
schema,
batch_size=_TRAIN_BATCH_SIZE)
eval_dataset = _input_fn(
fn_args.eval_files,
fn_args.data_accessor,
schema,
batch_size=_EVAL_BATCH_SIZE)
model = _build_keras_model()
model.fit(
train_dataset,
steps_per_epoch=fn_args.train_steps,
validation_data=eval_dataset,
validation_steps=fn_args.eval_steps)
# The result of the training should be saved in `fn_args.serving_model_dir`
# directory.
model.save(fn_args.serving_model_dir, save_format='tf')
# + [markdown] id="blaw0rs-emEf"
# Now you have completed all preparation steps to build a TFX pipeline.
# + [markdown] id="w3OkNz3gTLwM"
# ### Write a pipeline definition
#
# You define a function to create a TFX pipeline. A `Pipeline` object
# represents a TFX pipeline which can be run using one of pipeline
# orchestration systems that TFX supports.
#
# + id="M49yYVNBTPd4"
# TODO 2
def _create_pipeline(pipeline_name: str, pipeline_root: str, data_root: str,
module_file: str, serving_model_dir: str,
metadata_path: str) -> tfx.dsl.Pipeline:
"""Creates a three component penguin pipeline with TFX."""
# Brings data into the pipeline.
example_gen = tfx.components.CsvExampleGen(input_base=data_root)
# Uses user-provided Python function that trains a model.
trainer = tfx.components.Trainer(
module_file=module_file,
examples=example_gen.outputs['examples'],
train_args=tfx.proto.TrainArgs(num_steps=100),
eval_args=tfx.proto.EvalArgs(num_steps=5))
# Pushes the model to a filesystem destination.
pusher = tfx.components.Pusher(
model=trainer.outputs['model'],
push_destination=tfx.proto.PushDestination(
filesystem=tfx.proto.PushDestination.Filesystem(
base_directory=serving_model_dir)))
# Following three components will be included in the pipeline.
components = [
example_gen,
trainer,
pusher,
]
return tfx.dsl.Pipeline(
pipeline_name=pipeline_name,
pipeline_root=pipeline_root,
metadata_connection_config=tfx.orchestration.metadata
.sqlite_metadata_connection_config(metadata_path),
components=components)
# + [markdown] id="mJbq07THU2GV"
# ## Run the pipeline
#
# TFX supports multiple orchestrators to run pipelines.
# In this tutorial you will use `LocalDagRunner` which is included in the TFX
# Python package and runs pipelines on local environment.
# You often call TFX pipelines "DAGs" which stands for directed acyclic graph.
#
# `LocalDagRunner` provides fast iterations for developemnt and debugging.
# TFX also supports other orchestrators including Kubeflow Pipelines and Apache
# Airflow which are suitable for production use cases.
#
# See
# [TFX on Cloud AI Platform Pipelines](https://www.tensorflow.org/tfx/tutorials/tfx/cloud-ai-platform-pipelines)
# or
# [TFX Airflow Tutorial](https://www.tensorflow.org/tfx/tutorials/tfx/airflow_workshop)
# to learn more about other orchestration systems.
# + [markdown] id="7mp0AkmrPdUb"
# Now you create a `LocalDagRunner` and pass a `Pipeline` object created from the
# function you already defined.
#
# The pipeline runs directly and you can see logs for the progress of the pipeline including ML model training.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="fAtfOZTYWJu-" outputId="126e4374-09c7-422b-b52d-48b906ba1aa8"
tfx.orchestration.LocalDagRunner().run(
_create_pipeline(
pipeline_name=PIPELINE_NAME,
pipeline_root=PIPELINE_ROOT,
data_root=DATA_ROOT,
module_file=_trainer_module_file,
serving_model_dir=SERVING_MODEL_DIR,
metadata_path=METADATA_PATH))
# + [markdown] id="ppERq0Mj6xvW"
# You should see "INFO:absl:Component Pusher is finished." at the end of the
# logs if the pipeline finished successfully. Because `Pusher` component is the
# last component of the pipeline.
#
# The pusher component pushes the trained model to the `SERVING_MODEL_DIR` which
# is the `serving_model/penguin-simple` directory if you did not change the
# variables in the previous steps. You can see the result from the file browser
# in the left-side panel in Colab, or using the following command:
# + colab={"base_uri": "https://localhost:8080/"} id="NTHROkqX6yHx" outputId="3946c94a-af5f-40ef-aa69-cf9b0d991a71"
# TODO 3
# List files in created model directory.
# !find {SERVING_MODEL_DIR}
# + [markdown] id="08R8qvweThRf"
# ## Next steps
#
# You can find more resources on https://www.tensorflow.org/tfx/tutorials.
#
# Please see
# [Understanding TFX Pipelines](https://www.tensorflow.org/tfx/guide/understanding_tfx_pipelines)
# to learn more about various concepts in TFX.
#
| courses/machine_learning/deepdive2/tensorflow_extended/solutions/penguin_simple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tabula import read_pdf
import os
import pandas as pd
tables = read_pdf('Descolamento Aço + Killing + Passadeira 15d.is_tens.pdf',
pages='all', encoding='ISO-8859-1', multiple_tables=True)
cp1=[]
cp2=[]
cp3=[]
cp4=[]
cp5=[]
nome=[]
dirname = 'descolamentos'
for file_name in os.listdir(dirname):
if file_name.endswith('.pdf'):
file_path = os.path.join(dirname, file_name)
try:
tables = read_pdf(file_path, pages='all', encoding='ISO-8859-1', multiple_tables=True)
nome.append(file_name)
cp1.append(tables[1].loc[5][6])
cp2.append(tables[1].loc[6][6])
cp3.append(tables[1].loc[7][6])
cp4.append(tables[1].loc[8][6])
cp5.append(tables[1].loc[9][6])
except:
print(file_name)
d = {'nome':nome,'cp1':cp1,'cp2':cp2,'cp3':cp3,'cp4':cp4,'cp5':cp5}
df = pd.DataFrame(d)
df.head(8)
df["nome"] = df["nome"].str.split('Descolamento ', expand = True)[1]
new = df["nome"].str.split("+", n=2, expand = True)
df['Base']= new[0]
df['Cola']= new[1]
df['Substrato']= new[2]
df.drop(columns =["nome"], inplace = True)
df["Substrato"] = df["Substrato"].str.split('72h', expand = True)
df.to_csv('desc.csv', sep=';', encoding='ISO-8859-1')
df
cp1=[]
cp2=[]
cp3=[]
cp4=[]
cp5=[]
nome=[]
dirname = 'cisalhamentos'
for file_name in os.listdir(dirname):
if file_name.endswith('.pdf'):
file_path = os.path.join(dirname, file_name)
try:
tables = read_pdf(file_path, pages='all', encoding='ISO-8859-1', multiple_tables=True)
nome.append(file_name)
cp1.append(tables[1].loc[5][4])
cp2.append(tables[1].loc[6][4])
cp3.append(tables[1].loc[7][4])
cp4.append(tables[1].loc[8][4])
cp5.append(tables[1].loc[9][4])
except:
print(file_name)
d = {'nome':nome,'cp1':cp1,'cp2':cp2,'cp3':cp3,'cp4':cp4,'cp5':cp5}
df = pd.DataFrame(d)
df.head(8)
df["nome"] = df["nome"].str.split('Cisalhamento ', expand = True)[1]
new = df["nome"].str.split("+", n=2, expand = True)
df['Base']= new[0]
df['Cola']= new[1]
df['Substrato']= new[2]
df.drop(columns =["nome"], inplace = True)
df["Substrato"] = df["Substrato"].str.split('72h', expand = True)
df.to_csv('cisal.csv', sep=';', encoding='ISO-8859-1')
df
| ANOVA/Leitura.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [[source]](../api/alibi.explainers.anchor_tabular.rst)
# # Anchors
# ## Overview
# The anchor algorithm is based on the [Anchors: High-Precision Model-Agnostic Explanations](https://homes.cs.washington.edu/~marcotcr/aaai18.pdf) paper by Ribeiro et al. and builds on the open source [code](https://github.com/marcotcr/anchor) from the paper's first author.
#
# The algorithm provides model-agnostic (*black box*) and human interpretable explanations suitable for classification models applied to images, text and tabular data. The idea behind anchors is to explain the behaviour of complex models with high-precision rules called *anchors*. These anchors are locally sufficient conditions to ensure a certain prediction with a high degree of confidence.
#
# Anchors address a key shortcoming of local explanation methods like [LIME](https://arxiv.org/abs/1602.04938) which proxy the local behaviour of the model in a linear way. It is however unclear to what extent the explanation holds up in the region around the instance to be explained, since both the model and data can exhibit non-linear behaviour in the neighborhood of the instance. This approach can easily lead to overconfidence in the explanation and misleading conclusions on unseen but similar instances. The anchor algorithm tackles this issue by incorporating coverage, the region where the explanation applies, into the optimization problem. A simple example from sentiment classification illustrates this (Figure 1). Dependent on the sentence, the occurrence of the word *not* is interpreted as positive or negative for the sentiment by LIME. It is clear that the explanation using *not* is very local. Anchors however aim to maximize the coverage, and require *not* to occur together with *good* or *bad* to ensure respectively negative or positive sentiment.
#
#
# 
#
# Ribeiro et al., *Anchors: High-Precision Model-Agnostic Explanations*, 2018
# As highlighted by the above example, an anchor explanation consists of *if-then rules*, called the anchors, which sufficiently guarantee the explanation locally and try to maximize the area for which the explanation holds. This means that as long as the anchor holds, the prediction should remain the same regardless of the values of the features not present in the anchor. Going back to the sentiment example: as long as *not good* is present, the sentiment is negative, regardless of the other words in the movie review.
# ### Text
# For text classification, an interpretable anchor consists of the words that need to be present to ensure a prediction, regardless of the other words in the input. The words that are not present in a candidate anchor can be sampled in 2 ways:
#
# * Replace word token by UNK token.
#
# * Replace word token by sampled token from a corpus with the same POS tag and probability proportional to the similarity in the embedding space. By sampling similar words, we keep more context than simply using the UNK token.
# ### Tabular Data
# Anchors are also suitable for tabular data with both categorical and continuous features. The continuous features are discretized into quantiles (e.g. deciles), so they become more interpretable. The features in a candidate anchor are kept constant (same category or bin for discretized features) while we sample the other features from a training set. As a result, anchors for tabular data need access to training data. Let's illustrate this with an example. Say we want to predict whether a person makes less or more than £50,000 per year based on the person's characteristics including age (continuous variable) and marital status (categorical variable). The following would then be a potential anchor: Hugo makes more than £50,000 because he is married and his age is between 35 and 45 years.
# ### Images
# Similar to LIME, images are first segmented into superpixels, maintaining local image structure. The interpretable representation then consists of the presence or absence of each superpixel in the anchor. It is crucial to generate meaningful superpixels in order to arrive at interpretable explanations. The algorithm supports a number of standard image segmentation algorithms ([felzenszwalb, slic and quickshift](https://scikit-image.org/docs/dev/auto_examples/segmentation/plot_segmentations.html#sphx-glr-auto-examples-segmentation-plot-segmentations-py)) and allows the user to provide a custom segmentation function.
#
# The superpixels not present in a candidate anchor can be masked in 2 ways:
#
# * Take the average value of that superpixel.
#
# * Use the pixel values of a superimposed picture over the masked superpixels.
# 
#
# Ribeiro et al., *Anchors: High-Precision Model-Agnostic Explanations*, 2018
# ### Efficiently Computing Anchors
# The anchor needs to return the same prediction as the original instance with a minimal confidence of e.g. 95%. If multiple candidate anchors satisfy this constraint, we go with the anchor that has the largest coverage. Because the number of potential anchors is exponential in the feature space, we need a faster approximate solution.
#
# The anchors are constructed bottom-up in combination with [beam search](https://en.wikipedia.org/wiki/Beam_search). We start with an empty rule or anchor, and incrementally add an *if-then* rule in each iteration until the minimal confidence constraint is satisfied. If multiple valid anchors are found, the one with the largest coverage is returned.
#
# In order to select the best candidate anchors for the beam width efficiently during each iteration, we formulate the problem as a [pure exploration multi-armed bandit](https://www.cse.iitb.ac.in/~shivaram/papers/kk_colt_2013.pdf) problem. This limits the number of model prediction calls which can be a computational bottleneck.
#
# For more details, we refer the reader to the original [paper](https://homes.cs.washington.edu/~marcotcr/aaai18.pdf).
# ## Usage
# While each data type has specific requirements to initialize the explainer and return explanations, the underlying algorithm to construct the anchors is the same.
#
# In order to efficiently generate anchors, the following hyperparameters need to be set to sensible values when calling the `explain` method:
#
# * `threshold`: the previously discussed minimal confidence level. `threshold` defines the minimum fraction of samples for a candidate anchor that need to lead to the same prediction as the original instance. A higher value gives more confidence in the anchor, but also leads to more computation time. The default value is 0.95.
#
# * `tau`: determines when we assume convergence for the multi-armed bandit. A bigger value for `tau` means faster convergence but also looser anchor conditions. By default equal to 0.15.
#
# * `beam_size`: the size of the beam width. A bigger beam width can lead to a better overall anchor at the expense of more computation time.
#
# * `batch_size`: the batch size used for sampling. A bigger batch size gives more confidence in the anchor, again at the expense of computation time since it involves more model prediction calls. The default value is 100.
#
# * `coverage_samples`: number of samples used to compute the coverage of the anchor. By default set to 10000.
# ### Text
# #### Initialization
# Since the explainer works on black box models, only access to a predict function is needed. The model below is a simple logistic regression trained on movie reviews with negative or positive sentiment and pre-processed with a CountVectorizer:
#
# ```python
# predict_fn = lambda x: clf.predict(vectorizer.transform(x))
# ```
#
# If we choose to sample similar words from a corpus, we first need to load a spaCy model:
#
# ```python
# import spacy
# from alibi.utils.download import spacy_model
#
# model = 'en_core_web_md'
# spacy_model(model=model)
# nlp = spacy.load(model)
# ```
#
# We can now initialize our explainer:
#
# ```python
# explainer = AnchorText(nlp, predict_fn)
# ```
# #### Explanation
# Let's define the instance we want to explain and verify that the sentiment prediction on the original instance is positive:
#
# ```python
# text = 'This is a good book .'
# class_names = ['negative', 'positive']
# pred = class_names[predict_fn([text])[0]]
# ```
# Now we can explain the instance:
#
# ```python
# explanation = explainer.explain(text, threshold=0.95, use_proba=True, use_unk=False)
# ```
#
# We set the confidence `threshold` at 95%. `use_proba` equals True means that we will sample from the corpus proportional to the word similarity, while `use_unk` False implies that we are not replacing words outside the anchor with UNK tokens. `use_proba` False and `use_unk` True would mean that we are simply replacing the words that are not in the candidate anchor with the UNK tokens.
# The `explain` method returns a dictionary that contains *key: value* pairs for:
#
# * *names*: the words in the anchor.
#
# * *precision*: the fraction of times the sampled instances where the anchor holds yields the same prediction as the original instance. The precision will always be $\geq$ `threshold` for a valid anchor.
#
# * *coverage*: the coverage of the anchor over a sampled part of the training set.
#
# Under the *raw* key, the dictionary also contains example instances where the anchor holds and the prediction is the same as on the original instance, as well as examples where the anchor holds but the prediction changed to give the user a sense of where the anchor fails. *raw* also stores information on the *names*, *precision* and *coverage* of partial anchors. This allows the user to track the improvement in for instance the *precision* as more features (words in the case of text) are added to the anchor.
# ### Tabular Data
# #### Initialization and fit
# To initialize the explainer, we provide a predict function, a list with the feature names to make the anchors easy to understand as well as an optional mapping from the encoded categorical features to a description of the category. An example for `categorical_names` would be *category_map = {0: list('married', 'divorced'), 3: list('high school diploma', 'master's degree')}*. Each key in *category_map* refers to the column index in the input for the relevant categorical variable, while the values are lists with the options for each categorical variable. To make it easy, we provide a utility function `gen_category_map` to generate this map automatically from a Pandas dataframe:
#
# ```python
# from alibi.utils.data import gen_category_map
# category_map = gen_category_map(df)
# ```
#
# Then initialize the explainer:
# ```python
# predict_fn = lambda x: clf.predict(preprocessor.transform(x))
# explainer = AnchorTabular(predict_fn, feature_names, categorical_names=category_map)
# ```
# Tabular data requires a fit step to map the ordinal features into quantiles and therefore needs access to a representative set of the training data. `disc_perc` is a list with percentiles used for binning:
#
# ```python
# explainer.fit(X_train, disc_perc=[25, 50, 75])
# ```
# #### Explanation
# Let's check the prediction of the model on the original instance and explain:
#
# ```python
# class_names = ['<=50K', '>50K']
# pred = class_names[explainer.predict_fn(X)[0]]
# explanation = explainer.explain(X, threshold=0.95)
# ```
#
# The returned explanation dictionary contains the same *key: value* pairs as the text explainer, so you could explain a prediction as follows:
#
# ```
# Prediction: <=50K
# Anchor: Marital Status = Never-Married AND Relationship = Own-child
# Precision: 1.00
# Coverage: 0.13
# ```
# ### Images
# #### Initialization
# Besides the predict function, we also need to specify either a built in or custom superpixel segmentation function. The built in methods are [felzenszwalb](https://scikit-image.org/docs/dev/api/skimage.segmentation.html#skimage.segmentation.felzenszwalb), [slic](https://scikit-image.org/docs/dev/api/skimage.segmentation.html#skimage.segmentation.slic) and [quickshift](https://scikit-image.org/docs/dev/api/skimage.segmentation.html#skimage.segmentation.quickshift). It is important to create sensible superpixels in order to speed up convergence and generate interpretable explanations. Tuning the hyperparameters of the segmentation method is recommended.
#
# ```python
# explainer = AnchorImage(predict_fn, image_shape, segmentation_fn='slic',
# segmentation_kwargs={'n_segments': 15, 'compactness': 20, 'sigma': .5},
# images_background=None)
# ```
#
# Example of superpixels generated for the Persian cat picture using the *slic* method:
#
# 
# 
#
# The following function would be an example of a custom segmentation function dividing the image into rectangles.
#
#
# ```python
# def superpixel(image, size=(4, 7)):
# segments = np.zeros([image.shape[0], image.shape[1]])
# row_idx, col_idx = np.where(segments == 0)
# for i, j in zip(row_idx, col_idx):
# segments[i, j] = int((image.shape[1]/size[1]) * (i//size[0]) + j//size[1])
# return segments
# ```
#
# The `images_background` parameter allows the user to provide images used to superimpose on the masked superpixels, not present in the candidate anchor, instead of taking the average value of the masked superpixel. The superimposed images need to have the same shape as the explained instance.
# #### Explanation
# We can then explain the instance in the usual way:
#
# ```python
# explanation = explainer.explain(image, p_sample=.5)
# ```
#
# `p_sample` determines the fraction of superpixels that are either changed to the average superpixel value or that are superimposed.
#
# The explanation dictionary again contains information about the anchor's *precision*, *coverage* and examples where the anchor does or does not hold. On top of that, it also contains a masked image with only the anchor superpixels visible under the *anchor* key (see image below) as well as the image's superpixels under *segments*.
#
# 
# ## Examples
# ### Image
# [Anchor explanations for ImageNet](../examples/anchor_image_imagenet.nblink)
#
# [Anchor explanations for fashion MNIST](../examples/anchor_image_fashion_mnist.nblink)
# ### Tabular Data
# [Anchor explanations on the Iris dataset](../examples/anchor_tabular_iris.nblink)
#
# [Anchor explanations for income prediction](../examples/anchor_tabular_adult.nblink)
# ### Text
# [Anchor explanations for movie sentiment](../examples/anchor_text_movie.nblink)
| doc/source/methods/Anchors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# Image Deconvolution (ADMM Plug-and-Play Priors w/ BM4D)
# =======================================================
#
# This example demonstrates the use of class
# [admm.ADMM](../_autosummary/scico.optimize.rst#scico.optimize.ADMM) to
# solve a 3D image deconvolution problem using the Plug-and-Play Priors
# framework <cite data-cite="venkatakrishnan-2013-plugandplay2"/>, using BM4D
# <cite data-cite="maggioni-2012-nonlocal"/> as a denoiser.
#
# The 3D deconvolution problem is the task of recovering a 3D image
# that has been convolved with a 3D kernel and corrupted by noise.
# +
import numpy as np
import jax
import scico.numpy as snp
from scico import functional, linop, loss, metric, plot, random
from scico.examples import create_3D_foam_phantom, downsample_volume, tile_volume_slices
from scico.optimize.admm import ADMM, LinearSubproblemSolver
from scico.util import device_info
plot.config_notebook_plotting()
# -
# Create a ground truth image.
np.random.seed(1234)
N = 128 # phantom size
Nx, Ny, Nz = N, N, N // 4
upsamp = 2
x_gt_hires = create_3D_foam_phantom((upsamp * Nz, upsamp * Ny, upsamp * Nx), N_sphere=100)
x_gt = downsample_volume(x_gt_hires, upsamp)
x_gt = jax.device_put(x_gt) # convert to jax array, push to GPU
# Set up forward operator and test signal consisting of blurred signal with
# additive Gaussian noise.
# +
n = 5 # convolution kernel size
σ = 20.0 / 255 # noise level
psf = snp.ones((n, n, n)) / (n**3)
A = linop.Convolve(h=psf, input_shape=x_gt.shape)
Ax = A(x_gt) # blurred image
noise, key = random.randn(Ax.shape)
y = Ax + σ * noise
# -
# Set up ADMM solver.
# +
f = loss.SquaredL2Loss(y=y, A=A)
C = linop.Identity(x_gt.shape)
λ = 40.0 / 255 # BM4D regularization strength
g = λ * functional.BM4D()
ρ = 1.0 # ADMM penalty parameter
maxiter = 10 # number of ADMM iterations
solver = ADMM(
f=f,
g_list=[g],
C_list=[C],
rho_list=[ρ],
x0=A.T @ y,
maxiter=maxiter,
subproblem_solver=LinearSubproblemSolver(cg_kwargs={"tol": 1e-3, "maxiter": 100}),
itstat_options={"display": True},
)
# -
# Run the solver.
print(f"Solving on {device_info()}\n")
x = solver.solve()
x = snp.clip(x, 0, 1)
hist = solver.itstat_object.history(transpose=True)
# Show slices of the recovered 3D volume.
show_id = Nz // 2
fig, ax = plot.subplots(nrows=1, ncols=3, figsize=(15, 5))
plot.imview(tile_volume_slices(x_gt), title="Ground truth", fig=fig, ax=ax[0])
nc = n // 2
yc = y[nc:-nc, nc:-nc, nc:-nc]
yc = snp.clip(yc, 0, 1)
plot.imview(
tile_volume_slices(yc),
title="Slices of blurred, noisy volume: %.2f (dB)" % metric.psnr(x_gt, yc),
fig=fig,
ax=ax[1],
)
plot.imview(
tile_volume_slices(x),
title="Slices of deconvolved volume: %.2f (dB)" % metric.psnr(x_gt, x),
fig=fig,
ax=ax[2],
)
fig.show()
# Plot convergence statistics.
plot.plot(
snp.vstack((hist.Prml_Rsdl, hist.Dual_Rsdl)).T,
ptyp="semilogy",
title="Residuals",
xlbl="Iteration",
lgnd=("Primal", "Dual"),
)
| notebooks/deconv_ppp_bm4d_admm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# ### Egeria Hands-On Lab
# # Welcome to the Automated Curation Lab
#
# **NOTE - this lab is still under construction and should not be used**
#
# ## Introduction
#
# Egeria is an open source project that provides open standards and implementation libraries to connect tools, catalogs and platforms together so they can share information about data and technology (called metadata).
#
# In the [Building a Data Catalog](building-a-data-catalog.ipynb) lab, <NAME> and <NAME>
# manually catalogued the weekly measurement files for the Drop Foot clinical trial.
#
# In this hands-on lab you will get a chance to work with Egeria's governance servers to
# automate this onboarding process.
#
# ## The scenario
#
# [Coco Pharmaceuticals](https://opengovernance.odpi.org/coco-pharmaceuticals/)
# is conducting a clinical trial with two hospitals: Oak Dene Hospital and Old Market Hospital.
# Each week the two hospitals send Coco Pharmaceuticals a set of measurements from the patients
# involved in the trial. These measurements are located in a CSV file that the hospital sends through
# secure file transfer to a folder in Coco Pharmaceutical's landing area.
#
# These files need to be copied into the data lake and catalogued so that they are only visible to the
# staff involved in the clinical trial. It is also important that the lineage of these files is
# maintained so the source of the data can be traced. This process is shown in Figure 1.
#
# 
# > **Figure 1:** Clinical trial weekly measurements onboarding process
#
# <NAME> and Erin Overview are responsible for this onboarding process.
# 
#
# They have defined a list of requirements for the process:
#
# * Files must be in the landing area for a minimum amount of time.
# * As a new file is received, it needs to be catalogued, including:
# * Description
# * Connection details to enable the data scientists to access the contents
# * Column details
# * Governance zones defining the files' visibility
# * Owner
# * Origin
# * A file is not accessible by any of the data lake users until the cataloguing process is complete.
# * They must record lineage of each measurements file so they know which hospital it came from.
#
# They have been [manually cataloguing the measurements files](building-a-data-catalog.ipynb) for
# the first few weeks to prove the approach but now it is time to automate the process since:
# * This clinical trial is planned to run for two years.
# * There is a expected to be a ramp up of other clinical trials running simultaneously and the
# file onboarding workload would soon become overwhelming if they continued with the manual approach.
#
# They plan to use an
# [Integration Daemon](https://egeria.odpi.org/open-metadata-implementation/admin-services/docs/concepts/integration-daemon.html)
# called **exchangeDL01** to capture the technical metadata of the files.
# Then the
# [Engine Host](https://egeria.odpi.org/open-metadata-implementation/admin-services/docs/concepts/engine-host.html)
# server called **governDL01** will manage the move of the file into the data lake,
# the augmentation of the metadata properties of the files and the creation of the lineage.
#
# This lab sets up the automated onbourding process using 6 phases as shown in Figure 2:
#
# 
# > **Figure 2:** All six phases
#
# 1. Bring files in from "outside" using the move/copy file governance action service
# 2. Create a template used to catalog files in the landing areas
# 3. Trigger a governance action process when a file appears in a landing folder
# 4. Move the file into the data lake folder using a governance action process
# 5. Create a template used to catalog files in the data lake folder
# 6. Enrich the metadata using governance action services
#
#
# ## Setting up
#
# Coco Pharmaceuticals make widespread use of Egeria for tracking and managing their data and related assets.
# Figure 3 below shows their servers and the Open Metadata and Governance (OMAG) Server Platforms that are hosting them.
#
# 
# > **Figure 3:** Coco Pharmaceuticals' OMAG Server Platforms
#
# The code below checks that the platforms are running. It checks that the servers are configured and whether they are running on the platform. If a server is configured, but not running, it will start it.
#
# Look for the second "Done." message that is displayed just after the governance servers have started. It may take up to a minute to start up all of the servers. If `cocoMDS2` seems to be very slow starting, check that Apache Kafka is running. The metadata servers will wait for Kafka during start up if they are connected to a cohort. When they are waiting in this manner, they periodically output a message on the console.
# +
# Start up the metadata servers and the view server
# %run ../common/environment-check.ipynb
print("Start up the Governance Servers")
activatePlatform(dataLakePlatformName, dataLakePlatformURL, [governDL01Name, exchangeDL01Name, monitorGov01Name])
print("Done. ")
# -
# ----
# You should see that both the metadata servers `cocoMDS1` and `cocoMDS2` along with the integration daemon `exchangeDL01` and the engine host server `governDL01` have started. These are the servers that will be used in this lab and they are connected together
# as shown in figure 4. The `cocoMDS1` metadata server is where the new files are catalogued, whereas the definitions used
# to drive the governance processes come from `cocoMDS2`.
#
# 
# > **Figure 4:** Governance servers for automated curation
#
# If any of the platforms are not running, follow [this link to set up and run the platform](https://egeria.odpi.org/open-metadata-resources/open-metadata-labs/). If any server is reporting that it is not configured then
# run the steps in the [Server Configuration](../egeria-server-config.ipynb) lab to configure
# the servers. Then re-run the previous step to ensure all of the servers are started.
# ## Review the status of the integration daemon
#
# At this point, even though both `exchangeDL01` and `governDL01` are running, there is still work to set up the full data onboarding pipeline. Lets start with the integration daemon that catalogs the files as they appear in a folder.
#
# The command below queries the status of the integration daemon called `exchangeDL01`.
# +
getIntegrationDaemonStatus(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, petersUserId)
# -
# ----
# Notice that the Files Integrator Open Metadata Integration Service (OMIS) is running three integation connectors and they are all failing because the directories (folders) that they are supposed to be monitoring do not exist. As the data onboarding pipeline is set up in this lab, the directories will get created and we will be able to restart the connectors to get them working.
# +
OakDeneLandingAreaConnectorName = 'OakDeneLandingAreaFilesMonitor'
OldMarketLandingAreaConnectorName = 'OldMarketLandingAreaFilesMonitor'
DataLakeDirectoryConnectorName = 'DropFootClinicalTrialResultsFolderMonitor'
# -
# Below are the names of the directories that the three integration connectors are monitoring. The integration connector monitoring the files in the data lake directory is maintaining the last update time for the [DataFolder](https://egeria.odpi.org/open-metadata-publication/website/modelling-technology/) asset that represents the collection of measurements received from all hospitals for the clinical trial.
# +
landingAreaDirectory = fileSystemRoot + '/landing-area/hospitals'
dataLakeDirectory = fileSystemRoot + '/data-lake/research/clinical-trials/drop-foot/weekly-measurements'
OakDeneLandingDirectory = landingAreaDirectory + '/oak-dene/clinical-trials/drop-foot'
OldMarketLandingDirectory = landingAreaDirectory + '/old-market/clinical-trials/drop-foot'
# -
# The files that will provisioned into these directories are located in the sample data that is in the Egeria distribution.
# +
OakDeneSourceFolder = egeriaSampleDataRoot + '/sample-data/oak-dene-drop-foot-weekly-measurements'
OldMarketSourceFolder = egeriaSampleDataRoot + '/sample-data/old-market-drop-foot-weekly-measurements'
# -
#
# At this time there are no assets catalogued for either the landing area or the weekly measurements directory.
#
# ----
# +
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*landing.*")
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*drop-foot/weekly-measurements.*")
# -
# ----
# ## Setting up the file transfer into the landing area
#
# The directories (folders) that the integration connectors are configured to monitor could be created using a file system command.
# In this lab, however, the creation of a landing area folder will occur when the first file is received from the corresponding hospital.
# This is simply to avoid needing to provide this notebook with access to the file system.
#
# We are going to use a
# [provisioning governance action service](https://egeria.odpi.org/open-metadata-implementation/frameworks/governance-action-framework/docs/provisioning-governance-service.html) called
# [Move/Copy File Governance Action Service](https://egeria.odpi.org/open-metadata-publication/website/connector-catalog/move-copy-file-provisioning-governance-action-service.html)
# to simulate the file transfer from a hospital to its folder in the landing zone.
# This service runs in a governance engine that is supported by the
# [Governance Action Open Metadata Engine Service (OMES)](https://egeria.odpi.org/open-metadata-implementation/engine-services/governance-action/).
#
# It is possible to validate and understand the function of the governance action service
# through an API call to the Governance Action OMES running an engine host server such as `governDL01`.
# This API call is designed to support tools and others services configuring governance action services.
#
# Governance action services are implemented as a connector, which means they are initialized with a
# [connection](https://egeria.odpi.org/open-metadata-implementation/frameworks/open-connector-framework/docs/concepts/connection.html) object.
# This includes configuration properties that can be used to control its behavior.
# `Move/Copy File` is highly configurable.
#
# ----
# +
moveCopyFileProviderClassName = "org.odpi.openmetadata.adapters.connectors.governanceactions.provisioning.MoveCopyFileGovernanceActionProvider"
print("Move/Copy File Governance Action Service:")
validateGovernanceActionEngineConnector(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId, moveCopyFileProviderClassName)
# -
# ----
#
# To simulate the secure file transfer from the hospital to the landing area, we will configure
# `Move/Copy File`
# so it expects that the name of the source file to copy will be supplied when it is called (rather than using the catalog entry for the
# source file) and it does not create lineage.
#
# ----
# +
ftpGovernanceServiceName = "ftp-governance-action-service"
ftpGovernanceServiceDisplayName = "FTP Governance Action Service"
ftpGovernanceServiceDescription = "Simulates FTP from an external party."
ftpGovernanceServiceProviderClassName = moveCopyFileProviderClassName
ftpGovernanceServiceConfigurationProperties = {
"noLineage" : ""
}
ftpGovernanceServiceRequestType = "copy-file"
# -
# ----
#
# This governance action service
# runs in a governance engine supported by the Governance Action Open Metadata Engine Services (OMES).
# Figure 5 shows the three governance engines configured in the `governDL01` engine host server during the
# [Configuring Egeria Servers Lab](../egeria-server-config.ipynb).
#
# 
# > **Figure 5:** Governance Engines for governDL01
#
# The command below queries the status of each governance engine running in `governDL01`.
# The governance action services that will support the onboarding of files for clinical trials will run in the `AssetGovernance`
# governance engine. The other two governance engines are the subject of the [Improving Data Quality](../administration-labs/open-discovery-config.ipynb) lab.
#
# ----
# +
printGovernanceEngineStatuses(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId)
# -
# ----
# The status code `ASSIGNED` means that the governance engine was listed in Engine Host's configuration
# document - ie the governance engine was assigned to this server - but Engine Host has not been
# able to retrieve the configuration for the governance engine from the governance metadata server (`cocoMDS2`).
#
# The next step in the lab is to add configuration for the governance engine to `cocoMDS2` until the
# `AssetGovernance` governance engine is running. The first step is to add a GovernanceEngine definition
# to the metadata server for `AssetGovernance`.
#
# ----
# +
assetGovernanceEngineName = "AssetGovernance"
assetGovernanceEngineDisplayName = "Asset Governance Action Engine"
assetGovernanceEngineDescription = "Monitors, validates and enriches metadata relating to assets."
assetGovernanceEngineGUID = createGovernanceEngine(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
"GovernanceActionEngine",
assetGovernanceEngineName,
assetGovernanceEngineDisplayName,
assetGovernanceEngineDescription)
if (assetGovernanceEngineGUID):
print (" ")
print ("The guid for the " + assetGovernanceEngineName + " governance engine is: " + assetGovernanceEngineGUID)
print (" ")
print ("Done. ")
# -
# ----
#
# Now the governance engine is defined, its definition is sent to `governDL01` in an event
# and the engine's status moves to `CONFIGURING`.
# Notice more descriptive information is returned with the status.
#
# ----
# +
waitForConfiguringGovernanceEngine(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId, assetGovernanceEngineName)
printGovernanceEngineStatuses(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId)
# -
# ----
#
# Now we add the description of `Move/Copy File` to `cocoMDS2`. There are two parts to
# registering a governance action service. The first is to create a GovernanceService definition that identifies
# the implementation of the governance action service and the second part registers this
# GovernanceService definition with the GovernanceEngine definition.
# This registration maps one or more of the governance engine's request types, along with the default request parameters to
# the GovernanceService definition. This mapping is used to translate a request to the governance engine into an
# invocation of a governance action service.
#
# Figure 6 shows the structure of the resulting definitions for a governance engine. A governance action service may be
# registered with multiple governance engines, using the same or different request types.
#
# 
# > **Figure 6:** Structure of the governance services within a governance engine
#
# This next code issues the calls to create a definition of the GovernanceService and add it to the AssetGovernance
# GovernanceEngine definition.
#
# ----
# +
governanceServiceGUID = createGovernanceService(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
"GovernanceActionService",
ftpGovernanceServiceName,
ftpGovernanceServiceDisplayName,
ftpGovernanceServiceDescription,
ftpGovernanceServiceProviderClassName,
ftpGovernanceServiceConfigurationProperties)
if (governanceServiceGUID):
print (" ")
print ("The guid for the " + ftpGovernanceServiceName + " governance service is: " + governanceServiceGUID)
registerGovernanceServiceWithEngine(cocoMDS1Name,
cocoMDS1PlatformName,
cocoMDS1PlatformURL,
petersUserId,
assetGovernanceEngineGUID,
governanceServiceGUID,
ftpGovernanceServiceRequestType)
print ("Service registered as: " + ftpGovernanceServiceRequestType)
print (" ")
# -
# ----
#
# When at least one governance service is registered with the governance engine, the status of the governance engine in
# `governDL01` moves to `RUNNING` and it is possible to see the list of supported request types for the governance engine.
#
# ----
# +
waitForRunningGovernanceEngine(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId, assetGovernanceEngineName)
printGovernanceEngineStatuses(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId)
# -
# ----
#
# With the governance action service defined in a running governance engine, all that remains is to call it.
#
# Any defined governance action service can be called directly through the Governance Engine OMAS,
# which in this lab is running in the `cocoMDS2` metadata server. This call results in an event being sent to all
# engine hosts running the named governance engine. The first one that claims it gets to run it. In this lab
# there is only one engine host and so the request will run on `governDL01`.
#
# This governance action does not take long to run so you should soon see its status as completed.
#
# ----
# +
def addFileToLandingArea(sourceFolder, destinationFolder, weekNumber, qualifiedName):
sourceFileName = sourceFolder + "/" + "week" + weekNumber + ".csv"
requestParameters = {
"sourceFile" : sourceFileName,
"destinationFolder" : destinationFolder
}
governanceActionGUID = None
governanceActionGUID = initiateGovernanceAction(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
assetGovernanceEngineName,
qualifiedName,
ftpGovernanceServiceRequestType,
requestParameters)
waitForRunningGovernanceAction(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, governanceActionGUID)
printGovernanceAction(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, governanceActionGUID)
addFileToLandingArea(OakDeneSourceFolder, OakDeneLandingDirectory, "1", "FTP Oak Dene Week 1")
# -
# ----
# We now have the ability to provision new files into either landing area as shown in Figure 7.
#
# 
# > **Figure 7:** Phase 1: Arrival of new files
#
# ----
# Restarting the oak dene integration connector in the `exchangeDL01` integration daemon will cause it to
# test again to see if the directory it is monitoring exists.
#
# ----
# +
restartIntegrationConnector(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, petersUserId, "files-integrator", OakDeneLandingAreaConnectorName)
# -
# ----
#
# If we check the status of the integration connectors again, you can see this connector is now running.
#
# ----
# +
getIntegrationDaemonStatus(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, petersUserId)
# -
# ----
#
# The connector will begin cataloguing any files placed in the directory. We should be able to see the `week1.csv` file in the catalog along with catalog entries for each of the nested directories it sits in. If no assets are shown wait a short while and run the cell again and they will appear.
#
# ----
# +
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*landing.*")
# -
# ----
#
# The call below captures the full path name of the landing area directory by extracting it from the catalogued asset for the this directory.
# This will be used when configuring a later part of the onboarding process.
#
# ----
# +
landingAreaDirectoryName = assetOwnerFindAssetQualifiedName(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*" + landingAreaDirectory)
print(landingAreaDirectoryName)
# -
# ----
# The action of the integration connector has been to catalog the new file and the folders above it.
# The catalog definition is minimal, consisting of just what can be gleaned from the file system.
#
# It is possible to provide the integration connector with a template metadata element to copy when it
# is cataloguing files. This template can include classifications such as zones, origin, confidentiality and attachments such as
# connections, schemas and lineage mappings. (There is more information on templates in the
# [Egeria website](https://egeria.odpi.org/open-metadata-publication/website/cataloging-assets/templated-cataloging.html)).
#
# Templates are used to set up values for the assets created by the connector that are always the same.
# In this example, all of the files that this connector encounters are from the Oak Dene Hospital so we can use a template to set up the
# file's origin and the connection information needed to access the file.
#
# Including this type of detail in the asset means that no-one has to remember that this landing area folder was used by
# the Oak Dene Hospital and it simplifies the downstream cataloguing.
# Other values that are useful to set up in a template are any licenses for the file, schema information, zones, known lineage to this
# directory, plus other classifications. Information about the types of information that can be attached to an
# asset are available on the [Egeria website](https://egeria.odpi.org/open-metadata-publication/website/cataloging-assets/asset-catalog-contents.html).
#
#
# ## Working with templates
#
# If you know the class name of an integration connector's provider, it is possible to check if the connector is of the right type for an integration service. This function also returns full details of the connector type, which often includes descriptive information as well as the configuration properties that it supports.
#
# The connectors configured in the Files Integrator OMIS are shown in figure 8:
#
# 
# > **Figure 8:** exchangeDL01 with its connectors
#
# The class names of these integration connectors' providers can be seen in the connection object embedded in the error message displayed with the connectors' status.
#
# The commands below request that the Files Integrator OMIS service validate and return the connector type for each of these connectors.
# +
dataFilesMonitorProviderClassName = "org.odpi.openmetadata.adapters.connectors.integration.basicfiles.DataFilesMonitorIntegrationProvider"
dataFolderMonitorProviderClassName = "org.odpi.openmetadata.adapters.connectors.integration.basicfiles.DataFolderMonitorIntegrationProvider"
print("Data Files Monitor Integration Connector Type:")
validateIntegrationConnector(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, "files-integrator", petersUserId, dataFilesMonitorProviderClassName)
print("")
print("Data Folder Monitor Integration Connector Type:")
validateIntegrationConnector(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, "files-integrator", petersUserId, dataFolderMonitorProviderClassName)
# -
# ----
# Both connectors supports the `templateQualifiedName` and the `allowCatalogDelete` configuration properties. If you are curious
# about their meaning, review the definitions in the connector catalog:
#
# * [Data Files Monitor Integration Connector](https://egeria.odpi.org/open-metadata-publication/website/connector-catalog/data-files-monitor-integration-connector.html)
# * [Data Folder Monitor Integration Connector](https://egeria.odpi.org/open-metadata-publication/website/connector-catalog/data-folder-monitor-integration-connector.html)
#
# We are going to set the `templateQualifiedName` with the qualified name of an asset that has the origin set up for the each of the appropriate originating hospitals and membership of the default governance zone of "quarantine".
# This zone means that the asset is still being set up and it is not visible to the data lake users.
#
# ----
# The commands below create the template assets for each of the hospitals. In addition, we will create a third template for the asset created to
# represent the file when it is stored in the data lake folder. It will set up <NAME> as the owner of the files and marks the transition
# of ownership from the data lake oeprations team to the clinical trials team. It also has the zone membership set to default
# governance zone of "quarantine".
#
# This third template will be incorporated into the flow when we configure the governance action service that moves the file from the landing area to the data lake.
# +
newAssetZone = "quarantine"
t1AssetName = "Oak Dene Template"
t1QualifiedName = "template:clinical-trials:drop-foot:weekly-measurements:oak-dene"
t1DisplayName = "Drop Foot Clinical Trial Measurements Template Asset: Oak Dene"
t1Description = "Template asset for Oak Dene Hospital."
t1Contact = "<NAME>"
t1Dept = "Drop Foot Research Centre"
t1Org = "Oak Dene Hospital"
t2AssetName = "Old Market Template"
t2QualifiedName = "template:clinical-trials:drop-foot:weekly-measurements:old-market"
t2DisplayName = "Drop Foot Clinical Trial Measurements Template Asset: Old Market"
t2Description = "Template asset for Old Market Hospital."
t2Contact = "TBD"
t2Dept = "DFRG1F6"
t2Org = "Old Market Hospital"
template1guids = assetOwnerCreateCSVAsset(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, t1DisplayName, t1Description, t1QualifiedName)
template1guid = getLastGUID(template1guids)
addOrigin(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, t1AssetName, template1guid, t1Contact, t1Dept, t1Org)
addZones(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, petersUserId, t1AssetName, template1guid, [newAssetZone])
template2guids = assetOwnerCreateCSVAsset(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, t2DisplayName, t2Description, t2QualifiedName)
template2guid = getLastGUID(template2guids)
addOrigin(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, t2AssetName, template2guid, t2Contact, t2Dept, t2Org)
addZones(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, petersUserId, t2AssetName, template2guid, [newAssetZone])
t3AssetName = "Data Lake Measurements Template"
t3QualifiedName = "template:clinical-trials:drop-foot:weekly-measurements:data-lake"
t3DisplayName = "Drop Foot Clinical Trial Measurements Template Asset: Data Lake"
t3Description = "Template asset for Data Lake destination folder."
t3Owner = "tanyatidie"
t3OwnerType = "USER_ID"
template3guids = assetOwnerCreateCSVAsset(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, t3DisplayName, t3Description, t3QualifiedName)
template3guid = getLastGUID(template3guids)
addOwner(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, petersUserId, t3AssetName, template3guid, t3Owner, t3OwnerType)
addZones(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, petersUserId, t3AssetName, template3guid, [newAssetZone])
print ("\n\nNewTemplate Assets:")
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*Template.*")
# -
# ----
#
# The next commands reconfigure the integration connectors that are monitoring for new files in the landing area to use the templates ...
#
# ---
# +
t1ConnectorName = "OakDeneLandingAreaFilesMonitor"
t1ConfigurationProperties = {
"templateQualifiedName" : t1QualifiedName
}
t2ConnectorName = "OldMarketLandingAreaFilesMonitor"
t2ConfigurationProperties = {
"templateQualifiedName" : t2QualifiedName
}
updateConnectorConfigurationProperties(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, "files-integrator", petersUserId, t1ConnectorName, t1ConfigurationProperties)
getIntegrationConnectorConfigProperties(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, "files-integrator", petersUserId, t1ConnectorName)
updateConnectorConfigurationProperties(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, "files-integrator", petersUserId, t2ConnectorName, t2ConfigurationProperties)
getIntegrationConnectorConfigProperties(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, "files-integrator", petersUserId, t2ConnectorName)
# -
# ----
#
# Now lets add the file for week 2 into the landing area.
#
# ----
# +
addFileToLandingArea(OakDeneSourceFolder, OakDeneLandingDirectory, "2", "FTP Oak Dene Week 2")
# -
# ----
#
# The newly catalogued file will show that the origin and zones are set up.
#
# ----
# +
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*landing-area/hospitals/oak-dene/clinical-trials/drop-foot/week.*")
# -
# ----
# We are now cataloguing files arriving in the landing folders and their origin is set up - and so phase 2 shown in figure 9 is complete.
#
#
# 
# > **Figure 9:** Phase 2: Creating assets for newly arrived files with a template
#
#
# At this point, the integration daemon supports the onboarding of data from either hospital.
# The next step is to detect the creation of each file's catalogued asset to drive the process that moves the file into the data lake directory,
# and creates a catalog asset for its new location, along with lineage to show the onboarding path of the file.
#
# ## Listening for new assets
#
# The watchdog governance action services are responsible for listening for events that indicate specific types of activity and then acting on it.
# They can initiate
# a [governance action](https://egeria.odpi.org/open-metadata-implementation/access-services/governance-engine/docs/concepts/governance-action.html),
# a [governance action process](https://egeria.odpi.org/open-metadata-implementation/access-services/governance-engine/docs/concepts/governance-action-process.html) or
# an [incident report](https://egeria.odpi.org/open-metadata-implementation/access-services/governance-engine/docs/concepts/incident-report.html).
#
# For this onboarding process, we are going to set up a watchdog governance action service called `GenericFolderWatchdog` to
# listen for assets that are catalogued and linked to the landing area directory.
#
# Below is the definition of this governance action service.
#
# ----
# +
genericFolderWatchdogProviderClassName = "org.odpi.openmetadata.adapters.connectors.governanceactions.watchdog.GenericFolderWatchdogGovernanceActionProvider"
print("Generic Folder Watchdog Governance Action Service:")
validateGovernanceActionEngineConnector(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId, genericFolderWatchdogProviderClassName)
# -
# ----
#
# The calls below defines the governance action service and registers it with the `AssetGovernance` Governance Engine.
#
# ----
# +
watchdogGovernanceServiceName = "new-measurements-watchdog-governance-action-service"
watchdogGovernanceServiceDisplayName = "New Measurements Watchdog Governance Action Service"
watchdogGovernanceServiceDescription = "Initiates a governance action process when a new weekly measurements file arrives."
watchdogGovernanceServiceProviderClassName = genericFolderWatchdogProviderClassName
watchdogGovernanceServiceConfigurationProperties = {}
watchdogGovernanceServiceRequestType = "watch-nested-in-folder"
watchdogGovernanceServiceProcessToCall = "governance-action-process:clinical-trials:drop-foot:weekly-measurements:onboarding"
governanceServiceGUID = createGovernanceService(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
"GovernanceActionService",
watchdogGovernanceServiceName,
watchdogGovernanceServiceDisplayName,
watchdogGovernanceServiceDescription,
watchdogGovernanceServiceProviderClassName,
watchdogGovernanceServiceConfigurationProperties)
if (governanceServiceGUID):
print (" ")
print ("The guid for the " + watchdogGovernanceServiceDisplayName + " governance service is: " + governanceServiceGUID)
registerGovernanceServiceWithEngine(cocoMDS1Name,
cocoMDS1PlatformName,
cocoMDS1PlatformURL,
petersUserId,
assetGovernanceEngineGUID,
governanceServiceGUID,
watchdogGovernanceServiceRequestType)
print ("Service registered as: " + watchdogGovernanceServiceRequestType)
print (" ")
# -
# ----
#
# Now lets start it running in the Engine Host server `governDL01` by initiating a governance action.
#
# ----
# +
requestParameters = {
"interestingTypeName" : "DataFile",
"folderName" : landingAreaDirectoryName,
"actionTargetName" : "sourceFile",
"newElementProcessName" : watchdogGovernanceServiceProcessToCall
}
qualifiedName = "Listen for newly catalogued CSV Files in directory " + landingAreaDirectory
governanceActionGUID = None
governanceActionGUID = initiateGovernanceAction(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
assetGovernanceEngineName,
qualifiedName,
watchdogGovernanceServiceRequestType,
requestParameters)
# -
# ----
#
# With the watchdog in place, any new files added to the metadata repository will trigger a governance action process called `governance-action-process:clinical-trials:drop-foot:weekly-measurements:onboarding`.
#
# 
# > **Figure 10:** Phase 3: Triggering provisioning of the file in to the data lake
#
# ## Creating a governance action process for new assets
#
# A governance action process defines a flow of calls to governance action services
# that are linked together by the guards that they produce when they run.
# The guards produced by one governance action service invocation determine which governance action service runs next.
#
# A governance action process is represented by a metadata element of type [GovernanceActionProcess](https://egeria.odpi.org/open-metadata-publication/website/open-metadata-types/0462-Governance-Action-Types.html).
# This element gives the process a unique name and the anchor point to connect it into the governance action definitions.
#
# ----
# +
qualifiedName = watchdogGovernanceServiceProcessToCall
displayName = "Drop Foot Onboard Weekly Measurement Files"
description = "Ensures that new weekly drop foot measurement files from the hospitals are correctly catalogued in the data lake."
technicalName = "DFOBWKLY01"
technicalDescription = """ This process performs the follow function:
1) The physical file is moved to the data lake and renamed
2) A new asset is created for the new file
3) Lineage is created between the orginal file asset and the new file asset
4) The owner and origin are assigned
5) The governance zones are assigned to make the new asset visible to the research team."""
governanceActionProcessGUID = createGovernanceActionProcess(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, qualifiedName, displayName, description, technicalName, technicalDescription)
# -
# ----
#
# Added to the governance action process are the definitions of which type of governance actions should run when the process is initiated. These definitions are represented by
# [GovernanceActionType](https://egeria.odpi.org/open-metadata-publication/website/open-metadata-types/0462-Governance-Action-Types.html) metadata elements. They are linked together to define which governance action is started depending on the
# guards produced by the previous governance action.
#
# Each GovernanceActionType links to a GovernanceActionService configured in the GovernanceEngine.
#
# The first step in the process is to move the file from the landing area to the data lake folder.
#
# 
# > **Figure 11:** Phase 4: Move the file into the data lake
#
# For this we use a second instance of the `Move/Copy File` Governance Action Service.
# This instance will be set up to move the file, produce lineage and change the resulting filename so that the files are sequenced according to their arrival. For example:
# * DropFoot_000001.csv
# * DropFoot_000002.csv
#
# This aids the time-based loading of the files into a database by ensuring any corrections to the readings are applied in the
# correct order if a hospital sends a correction file at a later date.
#
# ----
# +
prGovernanceServiceName = "provision-weekly-measurements-governance-action-service"
prGovernanceServiceDisplayName = "File Provisioning Governance Action Service"
prGovernanceServiceDescription = "Provisions weekly measurement files from the landing area to the data lake."
prGovernanceServiceProviderClassName = moveCopyFileProviderClassName
prGovernanceServiceConfigurationProperties = {
"targetFileNamePattern" : "DropFoot_{1, number,000000}.csv",
"destinationFolder" : dataLakeDirectory
}
prGovernanceServiceRequestType = "move-file"
governanceServiceGUID = createGovernanceService(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
"GovernanceActionService",
prGovernanceServiceName,
prGovernanceServiceDisplayName,
prGovernanceServiceDescription,
prGovernanceServiceProviderClassName,
prGovernanceServiceConfigurationProperties)
if (governanceServiceGUID):
print (" ")
print ("The guid for the " + prGovernanceServiceName + " governance service is: " + governanceServiceGUID)
registerGovernanceServiceWithEngine(cocoMDS1Name,
cocoMDS1PlatformName,
cocoMDS1PlatformURL,
petersUserId,
assetGovernanceEngineGUID,
governanceServiceGUID,
prGovernanceServiceRequestType)
print ("Service registered as: " + prGovernanceServiceRequestType)
print (" ")
# -
# ----
#
# This governance action service is linked into the process via the following governance action type.
# Notice that the template created for cataloguing the files as they move into the data lake directory is set up in the `destinationFileTemplateQualifiedName` requestParameter as required for phase 5.
#
# 
# > **Figure 12:** Phase 5: Set up the template for newly arrived files in the data lake
#
# ----
# +
prGovernanceServiceType = "provision-weekly-measurements-governance-action-type"
prSupportedGuards = ["provisioning-complete", "provisioning-failed"]
prGovernanceServiceRequestParameters = {
"destinationFileTemplateQualifiedName" : t3QualifiedName,
}
prActionTypeGUID = None
prActionTypeGUID = createGovernanceActionType(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, assetGovernanceEngineGUID, prGovernanceServiceType, prSupportedGuards, prGovernanceServiceRequestType, prGovernanceServiceRequestParameters)
if prActionTypeGUID:
setupFirstActionType(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, governanceActionProcessGUID, prActionTypeGUID, False)
# -
# ----
#
# Now that the files are in the correct folder with the appropiate file name and a new catalog entry has been created, it is time to
# add the last touches to the catalog entry before it is published to the data lake users.
# This is phase 6 of the onboarding process.
#
# 
# > **Figure 12:** Phase 6: Extend the governance action process to enrich the asset
#
# The template used to create the catalog was able to set up the owner of the file because it is the same for each file that is moved into this directory.
# However, the origin is variable, depending on which hospital send the original file.
# To set the origin, we are going to use the
# [Origin Seeker Remediation Governance Action Service](https://egeria.odpi.org/open-metadata-publication/website/connector-catalog/origin-seeker-remediation-governance-action-service.html)
# that is able to navigate back through the lineage relationship created by the `Move/Copy File` governance action service to pick up the origin
# from the landing area asset.
#
# Below is the definition of `OriginSeeker`.
#
# ----
# +
originSeekerProviderClassName = "org.odpi.openmetadata.adapters.connectors.governanceactions.remediation.OriginSeekerGovernanceActionProvider"
print("Origin Seeker Remediation Governance Action Service:")
validateGovernanceActionEngineConnector(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId, originSeekerProviderClassName)
# -
# ----
#
# The calls below add `OriginSeeker` to the `AssetGovernance` governance engine, creates a governance action type and links it to the
# governance action type for `Move/Copy File`.
#
# ----
# +
osGovernanceServiceName = "origin-seeker-governance-action-service"
osGovernanceServiceDisplayName = "Locate and Set Origin Governance Action Service"
osGovernanceServiceDescription = "Navigates back through the lineage relationships to locate the origin classification(s) from the source(s) and sets it on the requested asset if the origin is unique."
osGovernanceServiceProviderClassName = originSeekerProviderClassName
osGovernanceServiceConfigurationProperties = {}
osGovernanceServiceRequestType = "seek-origin"
governanceServiceGUID = createGovernanceService(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
"GovernanceActionService",
osGovernanceServiceName,
osGovernanceServiceDisplayName,
osGovernanceServiceDescription,
osGovernanceServiceProviderClassName,
osGovernanceServiceConfigurationProperties)
if (governanceServiceGUID):
print (" ")
print ("The guid for the " + osGovernanceServiceName + " governance service is: " + governanceServiceGUID)
registerGovernanceServiceWithEngine(cocoMDS1Name,
cocoMDS1PlatformName,
cocoMDS1PlatformURL,
petersUserId,
assetGovernanceEngineGUID,
governanceServiceGUID,
osGovernanceServiceRequestType)
print ("Service registered as: " + osGovernanceServiceRequestType)
print (" ")
osPreviousGuard = "provisioning-complete"
osGovernanceServiceType = "origin-seeker-measurements-governance-action-type"
osSupportedGuards = [ "origin-assigned", "origin-already-assigned", "multiple-origins-detected", "no-origins-detected", "no-targets-detected", "multiple-targets-detected", "origin-seeking-failed"]
osGovernanceServiceRequestParameters = {
"destinationFileTemplateQualifiedName" : t3QualifiedName
}
osActionTypeGUID = createGovernanceActionType(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, assetGovernanceEngineGUID, osGovernanceServiceType, osSupportedGuards, osGovernanceServiceRequestType, osGovernanceServiceRequestParameters)
setupNextActionType(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, prActionTypeGUID, osActionTypeGUID, osPreviousGuard, True, True)
# -
# ----
#
# Once the origin is set, the asset is completely defined and it is ready to be consumed by the data lake users.
# The last governance action service called `ZonePublisher` sets the zone membership for the asset so that it becomes visible to the
# data lake users.
#
# Below is the definition of `ZonePublisher`.
#
# ----
# +
zonePublisherProviderClassName = "org.odpi.openmetadata.adapters.connectors.governanceactions.remediation.ZonePublisherGovernanceActionProvider"
print("Zone Publisher Remediation Governance Action Service:")
validateGovernanceActionEngineConnector(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId, zonePublisherProviderClassName)
# -
# ----
#
# The calls below add `ZonePublisher` to the `AssetGovernance` governance engine, creates a governance action type and links it to the
# governance action type for `OriginSeeker`.
#
# ----
# +
zpGovernanceServiceName = "zone-publisher-governance-action-service"
zpGovernanceServiceDisplayName = "Update Asset's Zone Membership Governance Action Service"
zpGovernanceServiceDescription = "Set up the zone membership for one or more assets supplied as action targets."
zpGovernanceServiceProviderClassName = zonePublisherProviderClassName
zpGovernanceServiceConfigurationProperties = {
"publishZones" : "data-lake,clinical-trials"
}
zpGovernanceServiceRequestType = "set-zone-membership"
governanceServiceGUID = createGovernanceService(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
"GovernanceActionService",
zpGovernanceServiceName,
zpGovernanceServiceDisplayName,
zpGovernanceServiceDescription,
zpGovernanceServiceProviderClassName,
zpGovernanceServiceConfigurationProperties)
if (governanceServiceGUID):
print (" ")
print ("The guid for the " + zpGovernanceServiceName + " governance service is: " + governanceServiceGUID)
registerGovernanceServiceWithEngine(cocoMDS1Name,
cocoMDS1PlatformName,
cocoMDS1PlatformURL,
petersUserId,
assetGovernanceEngineGUID,
governanceServiceGUID,
zpGovernanceServiceRequestType)
print ("Service registered as: " + zpGovernanceServiceRequestType)
print (" ")
zpPreviousGuard = "origin-assigned"
zpGovernanceServiceType = "zone-publisher-measurements-governance-action-type"
zpSupportedGuards = [ "zone-assigned","no-zones-detected","no-targets-detected", "zone-publishing-failed"]
zpGovernanceServiceRequestParameters = {
"destinationFileTemplateQualifiedName" : t3QualifiedName
}
zpActionTypeGUID = createGovernanceActionType(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, assetGovernanceEngineGUID, zpGovernanceServiceType, zpSupportedGuards, zpGovernanceServiceRequestType, zpGovernanceServiceRequestParameters)
setupNextActionType(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, osActionTypeGUID, zpActionTypeGUID, zpPreviousGuard, True, True)
# -
# ----
#
# We can check that these services are now registered with the `AssetGovernance` governance engine.
#
# ----
# +
printGovernanceEngineStatuses(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId)
# -
# ----
#
# Lets provision week 3 to see if the process is triggered and the file is moved to the data lake folder.
#
# ----
# +
addFileToLandingArea(OakDeneSourceFolder, OakDeneLandingDirectory, "3", "FTP Oak Dene Week 3")
# -
# ----
#
# The file is created in the data lake folder with all of the correct attributes.
#
# ----
# +
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*clinical-trials/drop-foot/weekly-measurements.*")
# -
# ----
#
# The file in the landing area is gone.
#
# ----
# +
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*landing-area/hospitals/oak-dene/clinical-trials/drop-foot/.*")
# -
# ----
# ## Running the complete onboarding pipeline
#
# The finale of this lab is to show the complete pipeline running. To do this we will use the files from both Oak Dene Hospital and the Old Market Hospital. First it is necessary to delete the Oak Dene files used in the experiments when setting up the pipeline. This can be done with a `Move/Copy File` governance action service configured to delete named files.
# +
dlGovernanceServiceName = "delete-files-governance-action-service"
dlGovernanceServiceDisplayName = "File Delete Governance Action Service"
dlGovernanceServiceDescription = "Removes weekly measurement files from the landing area."
dlGovernanceServiceProviderClassName = moveCopyFileProviderClassName
dlGovernanceServiceConfigurationProperties = {
"noLineage" : ""
}
dlGovernanceServiceRequestType = "delete-file"
governanceServiceGUID = createGovernanceService(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
"GovernanceActionService",
dlGovernanceServiceName,
dlGovernanceServiceDisplayName,
dlGovernanceServiceDescription,
dlGovernanceServiceProviderClassName,
dlGovernanceServiceConfigurationProperties)
if (governanceServiceGUID):
print (" ")
print ("The guid for the " + dlGovernanceServiceName + " governance service is: " + governanceServiceGUID)
registerGovernanceServiceWithEngine(cocoMDS1Name,
cocoMDS1PlatformName,
cocoMDS1PlatformURL,
petersUserId,
assetGovernanceEngineGUID,
governanceServiceGUID,
dlGovernanceServiceRequestType)
print ("Service registered as: " + dlGovernanceServiceRequestType)
print (" ")
# -
# ----
# The call below removes the three files.
#
# ----
# +
def removeFileFromLandingArea(sourceFolder, weekNumber, qualifiedName):
sourceFileName = sourceFolder + "/" + "week" + weekNumber + ".csv"
requestParameters = {
"sourceFile" : sourceFileName
}
governanceActionGUID = None
governanceActionGUID = initiateGovernanceAction(cocoMDS2Name,
cocoMDS2PlatformName,
cocoMDS2PlatformURL,
erinsUserId,
assetGovernanceEngineName,
qualifiedName,
dlGovernanceServiceRequestType,
requestParameters)
waitForRunningGovernanceAction(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, governanceActionGUID)
printGovernanceAction(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, erinsUserId, governanceActionGUID)
removeFileFromLandingArea(OakDeneLandingDirectory, "1", "Remove Oak Dene Week 1")
removeFileFromLandingArea(OakDeneLandingDirectory, "2", "Remove Oak Dene Week 2")
removeFileFromLandingArea(OakDeneLandingDirectory, "3", "Remove Oak Dene Week 3")
# -
# ----
#
# The code below uses the `Move/Copy File` governance action service to add the first five weeks files from each hospital into their landing area.
#
# ----
# +
addFileToLandingArea(OldMarketSourceFolder, OldMarketLandingDirectory, "1", "FTP Old Market Hospital Week 1")
addFileToLandingArea(OakDeneSourceFolder, OakDeneLandingDirectory, "1", "FTP Oak Dene Hospital Week 1")
addFileToLandingArea(OldMarketSourceFolder, OldMarketLandingDirectory, "2", "FTP Old Market Hospital Week 2")
addFileToLandingArea(OakDeneSourceFolder, OakDeneLandingDirectory, "2", "FTP Oak Dene Hospital Week 2")
addFileToLandingArea(OldMarketSourceFolder, OldMarketLandingDirectory, "3", "FTP Old Market Hospital Week 3")
addFileToLandingArea(OakDeneSourceFolder, OakDeneLandingDirectory, "3", "FTP Oak Dene Hospital Week 3")
addFileToLandingArea(OldMarketSourceFolder, OldMarketLandingDirectory, "4", "FTP Old Market Hospital Week 4")
addFileToLandingArea(OakDeneSourceFolder, OakDeneLandingDirectory, "4", "FTP Oak Dene Hospital Week 4")
addFileToLandingArea(OldMarketSourceFolder, OldMarketLandingDirectory, "5", "FTP Old Market Hospital Week 5")
addFileToLandingArea(OakDeneSourceFolder, OakDeneLandingDirectory, "5", "FTP Oak Dene Hospital Week 5")
# -
# ----
#
# At this point, nothing has been received from Old Market hospital, so its landing folder is not defined.
# This means its integration connector is in FAILED status. The code below restarts all of the integration connnectors in `exchangeDL01`.
#
# ----
# +
restartIntegrationConnector(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, petersUserId, "files-integrator", OldMarketLandingAreaConnectorName)
restartIntegrationConnector(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, petersUserId, "files-integrator", OakDeneLandingAreaConnectorName)
restartIntegrationConnector(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, petersUserId, "files-integrator", DataLakeDirectoryConnectorName)
# -
# ----
#
# When we check the status of the integration daemon, all connectors are running.
#
# ----
# +
getIntegrationDaemonStatus(exchangeDL01Name, exchangeDL01PlatformName, exchangeDL01PlatformURL, petersUserId)
# -
# ----
#
# This is the final status of the `AssetGovernance` governance engine.
#
# ----
# +
printGovernanceEngineStatuses(governDL01Name, governDL01PlatformName, governDL01PlatformURL, petersUserId)
# -
# ----
#
# From <NAME>'s perspective, he can see that all of the files have been created in the data lake folder and the landing area is empty ...
#
# ----
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*clinical-trials/drop-foot/weekly-measurements.*")
# +
oakDeneLandingAreaFullPath = "/" + landingAreaDirectoryName + "/oak-dene/clinical-trials/drop-foot"
os.listdir (oakDeneLandingAreaFullPath)
# -
# ----
# +
oldMarketLandingAreaFullPath = "/" + landingAreaDirectoryName + "/old-market/clinical-trials/drop-foot"
os.listdir (oldMarketLandingAreaFullPath)
# -
monitorGovernanceActions(cocoMDS2Name, cocoMDS2PlatformName, cocoMDS2PlatformURL, petersUserId)
# ----
assetOwnerPrintAssets(cocoMDS1Name, cocoMDS1PlatformName, cocoMDS1PlatformURL, petersUserId, ".*landing-.*")
# ----
#
# <img src="https://raw.githubusercontent.com/odpi/data-governance/master/docs/coco-pharmaceuticals/personas/callie-quartile.png" style="float:right">
#
# Finally we check that <NAME>, a data scientist working in the research team, is able to see the files from `cocoMDS3`:
#
#
# +
assetConsumerPrintAssets(cocoMDS3Name, cocoMDS3PlatformName, cocoMDS3PlatformURL, calliesUserId, ".*clinical-trials/drop-foot/weekly-measurements.*")
# -
# ----
# ## Where to next
#
# * [Improving Data Quality Lab](improving-data-quality-lab.ipynb) - follow Peter as he makes use of automated metadata discovery to detect quality errors in the
# measurement files from the hospitals.
# * [Understanding an Asset](understanding-an-asset.ipynb) - work with
#
# ----
| open-metadata-resources/open-metadata-labs/asset-management-labs/automated-curation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W0D3_LinearAlgebra/W0D3_Outro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>[](https://kaggle.com/kernels/welcome?src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W0D3_LinearAlgebra/W0D3_Outro.ipynb)
# + [markdown] pycharm={"name": "#%% md\n"}
# # Outro
#
# + [markdown] colab_type="text"
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Daily survey
#
# Don't forget to complete your reflections and content check in the daily survey! Please be patient after logging in as there is
# a small delay before you will be redirected to the survey.
#
# <a href="https://portal.neuromatchacademy.org/api/redirect/to/104eecd7-c1c3-4d82-8795-93ec4fb61d68"><img src="../static/button.png" alt="button link to survey" style="width:410px"></a>
#
| tutorials/W0D5_Statistics/W0D5_Outro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1) Implementation of Naive Bayes
import numpy as np
def fit(x_train , y_train):
result = {}
result["total_data"] = len(y_train)
class_values = set(y_train) ## get the distinct class values
num_features = x_train.shape[1]
for current_class in class_values:
result[current_class] = {}
current_class_rows = (y_train == current_class) ## Gives a true false array
x_train_current = x_train[current_class_rows]
y_train_current = y_train[current_class_rows] ## only those row which has true
result[current_class]["total_count"] = len(y_train_current)
for i in range(1 , num_features + 1):
result[current_class][i] = {};
all_possible_values = set(x_train[: , i - 1])
for current_values in all_possible_values:
result[current_class][i][current_values] =
(x_train_current[: , i - 1] == current_values).sum()
return result
## probability lead to very small number and we could not able to store
## that nuch small number therefore we will going to cal. log probility
def probability(dict , x ,current_class):
output = np.log(dict[current_class]["total_count"]) - np.log(dict["total_data"])
num_features = len(dict[current_class].keys()) - 1
for i in range(1 , num_features + 1):
xi = x[i - 1]
count_current_class_with_xi = dict[current_class][i][xi] + 1
count_current_class = dict[current_class]["total_count"]
+ len(dict[current_class][i].keys())
current_xi_probability = np.log(count_current_class_with_xi)
- np.log(count_current_class)
output += current_xi_probability
return output
def predictSinglePoint(dict , x):
## for each classes calculate bayes probability
## class whose prob is the smaller will be our output
classes = dict.keys()
best_p = -1000
best_class = -1
first_run = True
for current_class in classes:
if(current_class == 'total_data'):
continue
p_current_class = probability(dict , x , current_class)
if (first_run or p_current_class > best_p):
best_p = p_current_class
best_class = current_class
first_run = False
return best_class
def predict(dict, x_test):
y_pred = []
for x in x_test:
x_class = predictSinglePoint(dict, x)
y_pred.append(x_class)
return y_pred
# # 2) Testing of Iris Datasets on our Implemented Naive Byes
from sklearn import datasets
iris = datasets.load_iris()
x = iris.data
y = iris.target
def makeLabelled(column):
second_limit = column.mean()
first_limit = 0.5 * second_limit
third_limit = 1.5 * second_limit
for i in range(len(column)):
if column[i] < first_limit:
column[i] = 0
elif column[i] < second_limit:
column[i] = 1
elif column[i] < third_limit:
column[i] = 2
else:
column[i] = 3;
return column
## change continous data into labelled data
for i in range(0 , x.shape[-1]):
x[ : , i] = makeLabelled(x[ : , i])
from sklearn import model_selection
x_train, x_test, y_train, y_test =
model_selection.train_test_split(x, y, test_size = 0.25, random_state = 0)
dict = fit(x_train, y_train)
y_pred = predict(dict, x_test)
from sklearn.metrics import classification_report, confusion_matrix
print(classification_report(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# # 3) Naive Bayes for Continous Datasets
from sklearn.naive_bayes import GaussianNB
clf = GaussianNB()
clf.fit(x_train , y_train)
y_pred = clf.predict(x_test)
print(classification_report(y_test , y_pred))
print(confusion_matrix(y_test , y_pred))
# # 4) Multinomial Naive Bayes on iris datasets
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB()
clf.fit(x_train , y_train)
y_pred = clf.predict(x_test)
print(classification_report(y_test , y_pred))
print(confusion_matrix(y_test , y_pred))
| 15. Naive Bayes ..... [abhishek201202].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from matplotlib import pyplot as plot
from sklearn.cross_validation import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler
dataset = pd.read_csv('datasets/salaries.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, 1].values
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=1/3, random_state=0
)
# Although our data is not feature scaled we can use the LinearRegression.
# It will do feature scaling before fitting the training set.
regressor = LinearRegression()
regressor = regressor.fit(X_train, y_train)
# we then use the testing set to get the predicted values
y_prediction = regressor.predict(X_test)
# then we plot the oberserved points and the predicted values
plot.title('Salary vs. Experience (Trainig set)')
plot.xlabel('Years of experience')
plot.ylabel('Yearly salary')
plot.scatter(X_train, y_train, color='red')
plot.scatter(X_test, y_test, color='purple')
plot.plot(X_train, regressor.predict(X_train), color='blue')
plot.show()
| Part 2 - Regression/simple-linear-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Quality assurance with MNE report
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's say we want to analyze 100 subjects.
#
# How do we do quality assurancy in a scalable manner?
# + [markdown] slideshow={"slide_type": "subslide"}
# ## MNE report
# + slideshow={"slide_type": "-"}
from mne.report import Report
# + [markdown] slideshow={"slide_type": "fragment"}
# A report contains:
# * Figures
# * Images
# * Custom HTML
# * Sliders
# + [markdown] slideshow={"slide_type": "subslide"}
# First, let us generate some figures for the report.
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
import mne
from mne.datasets import sample
data_path = sample.data_path()
raw_fname = data_path + '/MEG/sample/sample_audvis_raw.fif'
raw = mne.io.read_raw_fif(raw_fname, preload=True)
# + [markdown] slideshow={"slide_type": "subslide"}
# Now let's pretend this data came from 3 different subjects
# + slideshow={"slide_type": "-"}
raw1 = raw.copy().crop(0, 20)
raw2 = raw.copy().crop(20, 40)
raw3 = raw.copy().crop(40, 60)
raw1.save(data_path + '/MEG/sample/sub-01_raw.fif', overwrite=True)
raw2.save(data_path + '/MEG/sample/sub-02_raw.fif', overwrite=True)
raw3.save(data_path + '/MEG/sample/sub-03_raw.fif', overwrite=True)
# + [markdown] slideshow={"slide_type": "subslide"}
# Now, we can have a function to go from raw to evoked
# + slideshow={"slide_type": "-"}
event_id = {'Auditory/Left': 3, 'Auditory/Right': 4}
def raw_to_evoked(raw_fname, tmin=-0.1, tmax=0.5):
raw = mne.io.read_raw_fif(data_path + '/MEG/sample/' + raw_fname, preload=True)
fig1 = raw.plot();
raw.filter(0, 40.)
events = mne.find_events(raw, stim_channel='STI 014')
epochs = mne.Epochs(raw, events, event_id, tmin, tmax)
fig2 = epochs.plot();
evoked_l = epochs['Left'].average();
fig3 = evoked_l.plot_topomap()
fig4 = evoked_l.plot();
return [fig1, fig2, fig3, fig4]
# + [markdown] slideshow={"slide_type": "subslide"}
# Now, we can get all the figure handles:
# + slideshow={"slide_type": "-"}
# %%capture
figs = raw_to_evoked('sub-01_raw.fif')
# + [markdown] slideshow={"slide_type": "subslide"}
# Now comes the actual report
# + slideshow={"slide_type": "-"}
rep = Report()
# + slideshow={"slide_type": "subslide"}
captions = ['Raw', 'Epochs', 'Topomap', 'Butterfly']
rep.add_figs_to_section(figs, captions=captions)
rep.save('report_raw_to_evoked.html')
# + [markdown] slideshow={"slide_type": "fragment"}
# The report can be found [here](report_raw_to_evoked.html)
# + [markdown] slideshow={"slide_type": "subslide"}
# We can go even more fancy. Let's try to process all the three subjects.
# + slideshow={"slide_type": "-"}
# %%capture
rep = Report()
for idx, r in enumerate(['sub-01_raw.fif', 'sub-02_raw.fif', 'sub-03_raw.fif']):
figs = raw_to_evoked(r)
rep.add_figs_to_section(figs, captions=captions, section='Subject %02d' % idx)
rep.save('report_raw_to_evoked.html', overwrite=True)
# + [markdown] slideshow={"slide_type": "skip"}
# There are tabs for each subject!
# + [markdown] slideshow={"slide_type": "slide"}
# Parallel processing
# -------------------
# + slideshow={"slide_type": "-"}
def raw_to_evoked(raw_fname, tmin=-0.1, tmax=0.5):
raw = mne.io.read_raw_fif(data_path + '/MEG/sample/' + raw_fname, preload=True)
raw.filter(0, 40.)
events = mne.find_events(raw, stim_channel='STI 014')
epochs = mne.Epochs(raw, events, event_id, tmin, tmax)
evoked_l = epochs['Left'].average();
from mne.parallel import parallel_func
fnames = ['sub-01_raw.fif', 'sub-02_raw.fif', 'sub-03_raw.fif']
parallel, myfunc, _ = parallel_func(raw_to_evoked, n_jobs=3)
parallel(myfunc(fname) for fname in fnames);
# + [markdown] slideshow={"slide_type": "slide"}
# BEM sliders
# -----------
# + [markdown] slideshow={"slide_type": "-"}
# What else can you do? You can inspect quality of the BEM with sliders.
# + slideshow={"slide_type": "fragment"}
subjects_dir = data_path + '/subjects'
rep = Report()
rep.add_bem_to_section(subject='sample', subjects_dir=subjects_dir, decim=36)
rep.save('report_bem.html')
# + [markdown] slideshow={"slide_type": "fragment"}
# Check out the report [here](report_bem.html)
# + [markdown] slideshow={"slide_type": "slide"}
# Custom HTML
# --------------------
# + [markdown] slideshow={"slide_type": "skip"}
# We can even add custom htmls. For example, we can say:
# + slideshow={"slide_type": "fragment"}
html = """
<table class="table table-hover">
<tr>
<th>Meas time range</th>
<th>Sampling freq</th>
</tr>
<tr>
<td> %0.2f to %0.2f </td>
<td> %0.2f </td>
</tr>
</table>
"""
# + slideshow={"slide_type": "subslide"}
rep.add_htmls_to_section(html % (raw.times[0], raw.times[-1], raw.info['sfreq']), captions='Info table')
rep.save('report_bem.html', overwrite=True)
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is the [report](report_bem.html).
# + [markdown] slideshow={"slide_type": "slide"}
# Custom sliders
# --------------
# + [markdown] slideshow={"slide_type": "skip"}
# And we can make our own sliders
# + slideshow={"slide_type": "fragment"}
import matplotlib.pyplot as plt
fname = data_path + '/MEG/sample/sample_audvis-ave.fif'
evoked = mne.read_evokeds(fname, condition='Left Auditory',
baseline=(None, 0), verbose=False)
rep = Report()
figs = list()
times = evoked.times[::4]
for time in times:
figs.append(evoked.plot_topomap(time, vmin=-300, vmax=300,
res=100, show=False))
plt.close(figs[-1])
rep.add_slider_to_section(figs, times, 'Evoked Response')
rep.save('report_slider.html')
# -
# To learn more about quality assurance, check out [this paper](https://www.biorxiv.org/content/biorxiv/early/2017/12/28/240044.full.pdf)
# + [markdown] slideshow={"slide_type": "slide"}
# Exercise
# --------
#
# 1) Can you think of creative ways to use the report for your own analysis?
# + [markdown] slideshow={"slide_type": "subslide"}
# <details>
# <summary>
# Here are some ideas:<br/><br/>
#
# </summary>
# - sections with subject names instead of preprocessing step<br/>
# - custom html/javascript to get quality labels<br/>
# - sliders to browse through the raw data<br/>
# </details>
| 2019_04_Brown/preprocessing/mne-report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from bs4 import BeautifulSoup as bs
import os
import requests
from splinter import Browser
from webdriver_manager.chrome import ChromeDriverManager
import pymongo
conn = 'mongodb://localhost:27017'
client = pymongo.MongoClient(conn)
db = client.MarsDB
collection = db.Mars
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
news_url = 'https://redplanetscience.com/'
browser.visit(news_url)
response = requests.get(news_url)
news_html = browser.html
soup = bs(news_html, 'html.parser')
#News Title
news_title = soup.find('div', class_='content_title').text
print(news_title)
#News Paragraph
news_p = soup.find('div', class_='article_teaser_body').text
print(news_p)
post = {
'title': news_title,
'paragraph': news_p,
'url': news_html
}
collection.insert_one(post)
#Photo
url='https://spaceimages-mars.com'
browser.visit(url)
browser.links.find_by_partial_text("FULL IMAGE").click()
pic_html = browser.html
soup = bs(pic_html,'html.parser')
pic_url = soup.find('img', class_='fancybox-image')['src']
print(url)
print(pic_url)
featured_image_url = url + pic_url
featured_image_url
post1= {'featured_image_url': featured_image_url}
collection.insert_one(post1)
#Facts
facts_url = 'https://galaxyfacts-mars.com'
browser.visit(facts_url)
facts_html = browser.html
soup = bs(facts_html, 'html.parser')
#mars_facts = soup.find('table', class_ = 'tbody')
mars_table = pd.read_html(facts_html)
mars_table
mars_df = mars_table[0]
mars_df
mars_df.columns = ["Categories", "Mars", "Earth"]
mars_df
marshtml = (mars_df.to_html())
print(marshtml)
mars_df.to_csv('../marsData.csv',index=False)
#Hemisphere
hemi_url = "https://marshemispheres.com/"
browser.visit(hemi_url)
hemi_html = browser.html
soup = bs(hemi_html, "html.parser")
hemi = soup.find_all("div", class_="item")
#hemi
hemi_info = []
for result in hemi:
h_title = result.find("h3").text
hemi_img = result.find("a", class_="itemLink product-item")["href"]
browser.visit(hemi_url + hemi_img)
image_html = browser.html
web_info = bs(image_html, "html.parser")
img_url = hemi_url + web_info.find("img", class_="wide-image")['src']
hemi_info.append({"title":h_title, "img_url":img_url})
browser.links.find_by_partial_text('Back').click()
print(h_title)
print(img_url)
post2 = {
'title': h_title,
'img_url': img_url
}
collection.insert_one(post2)
| Missions_to_Mars/Mission_to_Mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Important Announcements
# ### Rescheduled Class
# You should all have received an email concerning the 2 rescheduled class.
# <br>Rescheduled classes will be held on:
# - 06/26 (June 26, Tuesday ), period 1, room CALL 23
# - 06/29 (June 29, Friday ), period 5, room CALL 23
#
# You may attend either class.
# <br>The content will be identical.
#
# __THERE IS A CLASS ON JUNE 25, MONDAY AS USUAL__
# + [markdown] slideshow={"slide_type": "slide"}
# ### Rescheduled Class
# All classes on June 18, Monday are cancelled for University's Foundation Day.
#
# <br>The rescheduled class will be on June 20, period 5, Wednesday, room CALL 23.
#
# <br>(Normal Wednesday classes have been rescheduled)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exam
# A take-home exam with a time limit of 1 week will be issued on July 16 *instead of* the previously sheduled exam on July 23.
#
# The deadline for the take-home exam will be July 23.
#
# Please submit your exam solution by email.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Please download the new class notes.
# ### Step 1 : Navigate to the directory where your files are stored.
# Open a terminal.
# <br>Using `cd`, navigate to *inside* the ILAS_Python_for_engineers folder on your computer.
# ### Step 3 : Update the course notes by downloading the changes
# In the terminal type:
#
# >`git add -A
# git commit -m "commit"
# git fetch upstream
# git merge -X theirs upstream/master`
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Working with Data
#
# <br> <a href='#Pandas'>Pandas</a>
# <br> <a href='#Installation'>Installation</a>
# <br> <a href='#ImportingData'>Importing Data</a>
# <br>    <a href='#ConvertingPythonObject'>Converting a Python Object</a>
# <br>    <a href='#ImportingDataExternalFile'>Importing Data from an External File</a>
# <br>       <a href='#AssigningNamesUnlabelledData'>Assigning Names to Unlabelled Data</a>
# <br>       <a href='#RenamingColumns'>Renaming Columns</a>
# <br>       <a href='#RenamingRows'>Renaming Rows</a>
# <br>       <a href='#RenamingRowHeader'>Renaming Row Header</a>
# <br>       <a href='#ImportOptions'>Import Options</a>
# <br> <a href='#ViewingInspectingData'>ViewingInspectingData</a>
# <br> <a href='#SelectingData'>Selecting Data</a>
# <br> <a href='#Indexing'>Indexing</a>
# <br> <a href='#NumpyFunctionsDataFrames'>Numpy Functions for Data Frames</a>
# <br> <a href='#BooleanDataFrameIndexing'>Boolean Data Frame Indexing</a>
# <br> <a href='#ChangingValues'>Changing Values</a>
# <br> <a href='#ApplyingFunctionWholeSeries'>Applying a Function to a Whole Series</a>
# <br> <a href='#SortingDataFrames'>Sorting Data Frames</a>
# <br> <a href='#DataCleaning'>Data Cleaning</a>
# <br> <a href='#Iterating'>Iterating</a>
# <br> <a href='#PlottingDataFrames'>Plotting Data Frames</a>
# <br> <a href='#AddingRemovingJoining'>Adding, Removing and Joining</a>
#    <a href='#AddingColumnDataFrame'>Adding a Column to a Data Frame</a>
# <br>    <a href='#InsertingColumnDataFrame'>Inserting a Column to a Data Frame</a>
# <br>    <a href='#AddingRowDataFrame'>Adding a Row to a Data Frame</a>
# <br> <a href='#Resampling'>Resampling</a>
# <br> <a href='#SavingDataFrame'>Saving a Data Frame</a>
# <br> <a href='#OtherWaysSaveProgramOutput'>Other Ways to Save Program Output</a>
# <br> <a href='#ReadingWritingTextFiles'>Reading and Writing Text Files</a>
# <br>    <a href='#WritingFiles'>Writing Files</a>
# <br>    <a href='#ReadingFiles'>Reading Files</a>
# <br>    <a href='#AppendingFiles'>Appending Files</a>
# <br>    <a href='#AutomaticallyClosingFileswith'>Automatically Closing Files using `with`</a>
# <br>    <a href='#ReadingEditingFiles'>Reading and Editing Files</a>
# <br>    <a href='#BinaryFilesPicklingData'>Binary Files ("Pickling" Data)</a>
# <br>       <a href='#PicklingPandasDataFrame'>Pickling a Pandas Data Frame</a>
# <br>    <a href='#SavingDataNumpy'>Saving Data with Numpy</a>
# <br> <a href='#Summary'>Summary</a>
# <br> <a href='#TestYourselfExercises'>Test-Yourself Exercises</a>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Lesson Goal
#
# 1. Represent data in a way that is easier to manipulate.
# 1. Save data from your programs to output files.
#
# ### Fundamental programming concepts
# - Computational methods for handling medium-large data sets
# - Storing the output data from your programs
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pandas
# <a id='Pandas'></a>
# Pandas stands for “Python Data Analysis Library”.
#
# <img src="img/panda.jpg" alt="Drawing" style="width: 300px;"/>
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# Pandas takes data (e.g. .csv, .dat or .txt files) and creates a Python object with rows and columns.
#
# This object is called a data frame and looks similar to table in a statistical software (e.g. Excel).
#
# For real data, can be easier to work with in comparison to working with lists and/or numpy arrays.
#
# This seminar will give an introduction to some widely used functions of the `pandas` package.
# + [markdown] slideshow={"slide_type": "slide"}
#
#
# More information can be found in the documentation at http://pandas.pydata.org/.
#
#
# + slideshow={"slide_type": "slide"}
import pandas as pd
# + [markdown] slideshow={"slide_type": "slide"}
# Pandas is installed automatically when you install anaconda.
#
# If Pandas is not installed on your computer, follow the installation instructions below.
#
# (This procedure can be used to install any Python package.
# <br>You can repeat this process should you wish to use functions from a package that you do not already have installed.)
# + [markdown] slideshow={"slide_type": "subslide"}
#
# To install Pandas...
#
# ##### On Windows
#
# 1. Open the Anaconda Prompt from the terminal.
# <p align="center">
# <img src="img/anaconda_prompt.png" alt="Drawing" style="width: 300px;"/>
# </p>
#
# 1. The window that opens will look like the command line. In the window type the following code then press 'Enter':
# >`conda install -c anaconda pip`
#
# 1. When the installation completes type the following code then press 'Enter':
# >`pip install pandas`
#
#
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### On mac
#
# 1. Open a terminal.
#
# 1. Type the following code then press 'Enter':
# >`conda install -c anaconda pip`
#
# 1. When the installation completes type the following code then press 'Enter':
# >`pip install pandas`
# + [markdown] slideshow={"slide_type": "subslide"}
# To check the installation has worked type:
# >`import pandas`
#
# in Spyder or Jupyter notebook and run the code. If no error is generated you have installed pandas successfully.
# + [markdown] slideshow={"slide_type": "slide"}
# We will also use numpy and matplotlib in this class.
# -
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# ## Importing Data
# <a id='ImportingData'></a>
# .csv files contain comma separated values (although the values can be separated by other *delimiters*.).
#
# Real data is often stored in .csv files because dedicated software is not needed to read the file.
#
# This is the type of file is typically viewed in a program like Excel.
# <br>In this case, the commas determine the seperation of values into individual cells.
#
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Recap : Loading data as a Numpy array.
#
# Previously we learnt to use the function `numpy.loadtxt` to load data from a file.
# + slideshow={"slide_type": "-"}
np.loadtxt('sample_data/sample_student_data.txt', delimiter="\t", dtype=str)
# + [markdown] slideshow={"slide_type": "slide"}
# A limitation of this is that we sometimes need to specify the:
# - delimiter type
# - data type
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# If mixed data types are included (e.g. sex (string), weight (float)), we need to manipulate numeric data for use in mathematical operations.
#
#
# -
np.loadtxt('sample_data/sample_student_data.txt', dtype=float, skiprows=9, usecols=(3,4))
# + [markdown] slideshow={"slide_type": "slide"}
# `np.loadtxt` provides basic functionality.
#
# `pandas` loads data into a `DataFrame`.
#
# A `DataFrame` is like a Numpy array, with additional features for data analysis...
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# To create a Pandas data frame you either:
#
# - Convert a Python object (list, dictionary or Numpy array) to a Pandas data frame
# - Import the contents of a file (usually .csv)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Convert a Python Object
# <a id='ConvertPythonObject'></a>
# The basic command is:
#
# pd.DataFrame()
# +
# generate random data
data = np.random.randint(0, high=10, size=(4, 4))
print(data)
print()
print(pd.DataFrame(data))
# + [markdown] slideshow={"slide_type": "slide"}
# The data is automatically assigned:
# - an `index` to labelthe rows.
# - `headers` to label the columns
#
# We can specify our own label names for rows and columns...
# +
indices = ["Student 1", "Student 2", "Student 3", "Student 4"]
headers = ["score1", "score2", "score3", "score4"]
data_frame = pd.DataFrame(data, index = indices, columns = headers)
print(data)
print()
print(data_frame)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Importing Data from an External File
# <a id='ImportingDataExternalFile'></a>
# The basic command is:
#
# pd.read_csv()
# + [markdown] slideshow={"slide_type": "slide"}
# An advantages of Pandas vs just using NumPy is that Pandas allows you to have columns with different data types.
#
# DataFrames also accomodate `NaN` (not a number) values.
# + [markdown] slideshow={"slide_type": "slide"}
# The following example uses Pandas `read_csv` function to load the same student data we imported with `np.loadtxt`.
# -
students = pd.read_csv('sample_data/sample_student_data.csv')
print(students)
# + [markdown] slideshow={"slide_type": "slide"}
# Data Frames can be viewed more clearly using the function `display`.
# -
from IPython.display import display
# +
display(students[:4]) # Use regular indexing to display first 4 entries
display(students.head(4)) # Display first 4 entries
display(students.tail()) # Display last 4 entries
# + [markdown] slideshow={"slide_type": "slide"}
# We can use similar commands to import data from other sources e.g. data pubilshed remotely on the internet.
# +
url = 'https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv'
chipo = pd.read_csv(url, sep = '\t')
display(chipo.head())
# + slideshow={"slide_type": "slide"}
# Location of file
Location = 'sample_data/noheader_noindex.xlsx'
# Parse a specific sheet
df = pd.read_excel(Location)
display(df.head())
# + [markdown] slideshow={"slide_type": "slide"}
# The first row is automatically used as the columns headings.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Assigning Names to Unlabelled Data.
# <a id='AssigningNamesUnlabelledData'></a>
# Sometimes your data doesn't have column or row names.
#
# `pandas` will try to create them.
#
# `sample_data/noHeader_noIndex.csv` contains the data:
#
# ```Python
# 65.056, 1.053, 2.105, 3.158, 4.211
# 74.452, 48.348, 68.733, 59.796, 54.123
# ```
#
# + slideshow={"slide_type": "slide"}
unlabelled = pd.read_csv('sample_data/noHeader_noIndex.csv')
display(unlabelled)
print(unlabelled.keys())
# + [markdown] slideshow={"slide_type": "slide"}
# Therefore, if there are no headers, in the imported file, they should either be omitted...
# +
unlabelled = pd.read_csv('sample_data/noHeader_noIndex.csv', header=None)
display(unlabelled)
# + [markdown] slideshow={"slide_type": "-"}
# ... or assigned by the user:
# +
headers = ["X","Y"]
unlabelled_vert = pd.read_csv('sample_data/noHeader_noIndex_vert.csv', names=headers)
display(unlabelled_vert)
# + [markdown] slideshow={"slide_type": "slide"}
# Note, this can also be used to *replace* existing column names on import as we will see later...
# + [markdown] slideshow={"slide_type": "slide"}
# ### Import Options
# <a id='ImportOptions'></a>
# The read_csv function has a large number of keyword arguments:
# http://pandas.pydata.org/pandas-docs/version/0.23/generated/pandas.read_csv.html
#
#
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# Here are two examples.
# -
# `skiprows`: rows to omit
students = pd.read_csv('sample_data/sample_student_data.csv', skiprows=[1, 4])
# + [markdown] slideshow={"slide_type": "subslide"}
# `index_col`: the names in any column can be used as the index used to select a row.
# +
# DOB as index
students = pd.read_csv('sample_data/sample_student_data.csv', skiprows=[1], index_col=2)
# Student (JW- code) as index
students = pd.read_csv('sample_data/sample_student_data.csv', skiprows=[1], index_col=0)
display(students[:4])
# + [markdown] slideshow={"slide_type": "subslide"}
# To return column and index names:
# -
print(students.columns)
print(students.index)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Renaming Columns
# <a id='RenamingColumns'></a>
# Often, the columns names of imported data are not very convenient (long, including spaces, etc.).
#
# In the `students` DataFrame, the columns have convenient names.
#
# Lets pretend they don't and change them for demonstration purposes.
# + slideshow={"slide_type": "slide"}
# Create a new DataFrame with new names.
capitalised = students.rename(columns={'Height': 'HEIGHT',
'Weight': 'WEIGHT'}, inplace=False)
display(students.head())
display(capitalised.head())
# + slideshow={"slide_type": "slide"}
# Change the names of the original DataFrame inplace.
students.rename(columns={'Height': 'HEIGHT',
'Weight': 'WEIGHT'}, inplace=True)
display(students.head())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Renaming Rows
# <a id='RenamingRows'></a>
# Use `index` in place of `columns` to rename rows.
# +
nums = students.rename(index={'JW-1': '1',
'JW-10': '10'}, inplace=False)
display(nums.head())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Renaming Row Header
# <a id='RenamingRowHeader'></a>
# +
students.index.names = ['STUDENTS']
display(students.head())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Viewing and Inspecting Data
# <a id='ViewingInspectingData'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# You can print generic summary statistics for numerical columns:
# -
students.describe()
# + [markdown] slideshow={"slide_type": "slide"}
# ...or use functions to obtain specific statistics:
# -
print(students.mean()) # mean of all columns
print(students.corr()) # correlation between columns
print(students.count()) # number of non-null values in each column
print(students.max()) # highest value in each column
print(students.min()) # lowest value in each column
print(students.median()) # median of each column
print(students.std()) # standard deviation of each colum
# + [markdown] slideshow={"slide_type": "slide"}
# Functions such as these can also be applied to selected columns of the data set.
#
# To do this we need to select which columns we want to apply the function to.
#
# Pandas makes selecting data easier compared to a list or array.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Selecting Data
# <a id='SelectingData'></a>
# Columns can be selected using their *keywords*.
#
# (This is similar to a dictionary).
# +
students = pd.read_csv('sample_data/sample_student_data.csv', skiprows=[1], index_col=0)
print(students['Weight'].head())
print()
print(students['Weight'][:5])
# + slideshow={"slide_type": "slide"}
print(students[['Weight', 'DOB']][:4])
# + [markdown] slideshow={"slide_type": "slide"}
# Alternative method for single columns:
# -
print(students.Weight.head())
# +
print(students['Weight'].head())
print(students[:5].Weight)
# + [markdown] slideshow={"slide_type": "slide"}
# The selected data is returned as a new data frame.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Indexing
# <a id='Indexing'></a>
# A DataFrame may be indexed into like an array.
#
# #### .iloc syntax
#
# The `.iloc` (*index location*) indexing syntax is used.
# + slideshow={"slide_type": "slide"}
print(students.iloc[0, 1]) # row 0, col 1 : sex of student JW-1
print(students.iloc[1, 2]) # row 1, col 2 : DOB of student JW-2
print(students.iloc[2:4, 3]) # row 2-4, col 3 : Height of students JW-3 to JW-4
# print(students.iloc[3, 0:3]) # row 3, col 0-3 : Sex, DOB, Height of student JW-4
# print(students.iloc[0]) # row 0, all cols : all data for student JW-1
# print(students.iloc[:, 3]) # col 3, all rows, all Height data (first 5 entries)
# + [markdown] slideshow={"slide_type": "slide"}
# #### .loc syntax
# Alternatively, values in a DataFrame can be indexed by keyword.
#
# The `.loc` indexing syntax is used.
#
# - This is *much* more explicit.
# - Individual rows/columns can be selected but ranges cannot be specified
# + slideshow={"slide_type": "slide"}
print(students.loc['JW-1', 'Sex']) # Single data point
# print(students.loc['JW-2', ['DOB', 'Height']) # One row, two columns
# print(students.loc[['JW-3', 'JW-12'], 'Height']) # Two columns, one row
print(students.loc['JW-1']) # One row, all columns
print(students.loc[:,'Height']) # All rows, one column
# + [markdown] slideshow={"slide_type": "slide"}
# We can now apply the functions as before, but to the selected data only.
# + slideshow={"slide_type": "slide"}
# dot syntax : 1 column
print(students.Weight.mean()) # mean of all weight
# + slideshow={"slide_type": "slide"}
# loc syntax : individual rows/columns
print(students.loc[:, ['sex', 'BP']].count()) # number of non-null sex and BP values
print(students.loc[['JW-3', 'JW-5', 'JW-10'], 'Height'].mean()) # mean Height, students JW-3, JW-5, JW-10
print(students.loc[:, ['Height', 'Weight']].corr()) # correlation between Height and Weight values
# + slideshow={"slide_type": "slide"}
# # iloc syntax : ranges
print(students.iloc[2:6, 2:4].max()) # highest value of Height and Weight, students JW-3 to JW-5
print(students.iloc[:, 2:4].min()) # lowest value of all Height and Weight
print(students.iloc[:, 2].median()) # median of all Height
print(students.iloc[:, 3].std()) # standard deviation of all Weight
print(students.iloc[:, 2].sum()) # sum of all Weights
# + [markdown] slideshow={"slide_type": "slide"}
# ### Time Series Data
# <a id='TimeSeriesData'></a>
# For time series data, `pandas` can be used to read csv files where one of the columns includes date-time data.
#
# Indicating the column contains date-time values prompts `pandas` to try to convert that column to *datetime objects*.
#
# Datetime objects are very convenient as specifics of the datetime object may be accessed with dot syntax (e.g. `.year` returns the year, `.month` returns the month, etc.)
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# In the DataFrame `students`, DOB appears in the column with index 2 <br>(regular indexing, not .iloc or .loc indexing) <br>so we use the argument `parse_dates=[2]`
# + slideshow={"slide_type": "-"}
students = pd.read_csv('sample_data/sample_student_data.csv',
skiprows=[1],
index_col=0,
parse_dates=[2])
# + [markdown] slideshow={"slide_type": "slide"}
# Time series data may also contain the time in addition to the date.
#
# This is autonmatically included in a datetime object.
# + slideshow={"slide_type": "-"}
print(students.loc['JW-2', 'DOB'], end='\n\n')
print(students.loc['JW-2', 'DOB'].year, end='\n\n')
print(students.loc['JW-2', 'DOB'].month, end='\n\n')
print(students.loc['JW-2', 'DOB'].day, end='\n\n')
# + [markdown] slideshow={"slide_type": "slide"}
# Time data may be accessed using:
# - `.hour`
# - `.minute`
# - `.time()`
# + [markdown] slideshow={"slide_type": "slide"}
# ### Numpy Functions for Data Frames
# <a id='NumpyFunctionsDataFrames'></a>
# In general, many numpy fuctions can be applied to data frames in the same way as to numpy arrays.
#
# Functions can also be accessed with the *dot* syntax:
# > `dataframe.function()`
#
# where `dataframe_name` and `function` are replaced with the actual data frame and function names.
# + [markdown] slideshow={"slide_type": "slide"}
# Jupyter notebook has a convenient command to show all available functions.
#
# 1.Type the name of the data frame followed by a dot e.g.
#
# >`students.`
#
# 1. Press "Tab" to see all the functions that are available (there are many).
# + [markdown] slideshow={"slide_type": "slide"}
# Example: The Numpy function `argmax` returns the index of the greatest value.
#
# This can then be used to find the value of another parameter at that index.
# +
print('Maximum height amongst students', students.Height.max())
print('Tallest student:', students.Height.argmax())
print('Sex of tallest student:', students.Sex[students.Height.argmax()])
# + [markdown] slideshow={"slide_type": "slide"}
# Example: Applying a Numpy function with two columns as input arguments.
# + slideshow={"slide_type": "-"}
BMI = np.divide(students['Height']**2, students['Weight'])
print(BMI)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Boolean Data Frame Indexing
# <a id='BooleanDataFrameIndexing'></a>
# Using the same principle as the `argmax` example, we can use *conditionals* to select items that meet a certain criteria.
# + slideshow={"slide_type": "slide"}
girls = students.index[students.Sex == 'F']
taller_than = students.index[students.Height > 1.6]
girls_taller_than = students.index[((students.Sex == 'F') * (students.Height > 1.6))]
print('All female students:', girls.tolist())
print('All students taller than 1.6m:', taller_than.tolist())
print('All female students taller than 1.6m:', girls_taller_than.tolist())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Changing Values
# <a id='ChangingValues'></a>
# We can overwrite the values of a data frame.
#
# __Example : Applying a conversion factor.__
#
# Convert Heights, given in m, to units of cm
#
#
# +
print(students.head())
students.Height *= 100
print()
print(students.head())
# + [markdown] slideshow={"slide_type": "slide"}
# We can select (and change) values within a DataFrame based on a condition.
#
# __Example__
# Set all Weight above 80 to 80.
#
# Row index : `students.Weight > 80`
#
# Column index : `Weight`
# +
print(students.head())
students.loc[students.Weight > 80, 'Weight'] = 80
print()
print(students.head())
# + [markdown] slideshow={"slide_type": "slide"}
# __Example__
# Set the Height of all students with Weight = 80, to 160
#
# Row index : `students.Weight == 80`
#
# Column index : `Height`
#
# +
print(students.head())
students.loc[students.Weight == 80, 'Height'] = 160
print()
print(students.head())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Applying a Function to a Whole Series
# <a id='ApplyingFunctionWholeSeries'></a>
# You can apply a function that takes a single value as input, to a whole Data Frame series, using the keyword `apply`.
#
#
# The column or index name is used to indicte which series to apply the function to.
# + [markdown] slideshow={"slide_type": "slide"}
# Example: apply the function $x^2$ to every value ($x$) in the column __Weight__.
# +
def square(x):
return x**2
students.Weight = students.Weight.apply(square)
print(students.head())
# + [markdown] slideshow={"slide_type": "slide"}
# This can also be expressed as a Lambda function
# -
students.Weight = students.Weight.apply(lambda x : x**2)
print(students.head())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Sorting Data Frames
# <a id='SortingDataFrames'></a>
# -
# DataFrames may be sorted with the `.sort_values` function.
#
# The argument `by` determines which column the entries should be ordered by.
# + slideshow={"slide_type": "slide"}
print('Students in height order:')
display(students.sort_values(by='Height'))
# + [markdown] slideshow={"slide_type": "slide"}
# The function has several default/keyword arguments.
#
# `inplace=True` replaces the original DataFrame with the new sorted DataFrame.
# <br>When `inplace=False` (default) a new DataFrame is returned (which you can store in a separate variable).
#
# `ascending=True` (default) sorts the values in ascending order.
# <br>`ascending=False` sorts the values in descending order.
# + slideshow={"slide_type": "slide"}
print('Students in height order:')
print(students.sort_values(by='Height', ascending=False).head())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data Cleaning
# <a id='DataCleaning'></a>
# Data cleaning is a very important step in data analysis.
#
# For example, we always as missing values can appear in the data set without generating an error, we must check to identify null or `NaN` values.
#
# The .csv file, 'sample_data/data_with_holes.csv' has some empty cells.
# +
header = ['Item 1', 'Item 2', 'Item 3', 'Item 4']
df = pd.read_csv('sample_data/data_with_holes.csv', names=header)
print(df)
# + slideshow={"slide_type": "slide"}
print(df.isnull()) # checks for null/missing values, and returns a boolean array
print()
print(df.isnull().sum()) # number of null/missing in each column
print()
print(df.notnull()) # checks for NOT null/missing values
# + [markdown] slideshow={"slide_type": "slide"}
# Missing values can then be:
# - removed (whole column or row), `df.dropna()`
# - filled, `df.fillna()`
# + slideshow={"slide_type": "slide"}
print(pd.read_csv('sample_data/data_with_holes.csv', names=header))
print()
# # remove rows
# print(pd.read_csv('sample_data/data_with_holes.csv', names=header).dropna())
# # remove columns
# print(pd.read_csv('sample_data/data_with_holes.csv', names=header).dropna(axis=1))
# # fill gaps with 0
# print(pd.read_csv('sample_data/data_with_holes.csv', names=header).fillna(0))
# # fill gaps with mean of others values in column
# df = pd.read_csv('sample_data/data_with_holes.csv', names=header)
# print(df.fillna(df.mean))
# + [markdown] slideshow={"slide_type": "slide"}
# Duplicates can be removed from the data set
# +
df = pd.read_csv('sample_data/data_with_holes.csv', names=header)
df_filtered = df.drop_duplicates('Item 4')
print(df)
print()
print(df_filtered)
# + [markdown] slideshow={"slide_type": "slide"}
# Optionally, you can specify:
# - which value to keep (default keeps first)
# - to apply the change to the current data frame
# +
df = pd.read_csv('sample_data/data_with_holes.csv', names=header)
df.drop_duplicates('Item 4', keep='last', inplace=True)
print(df)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Iterating
# <a id='Iterating'></a>
# To iterate through each index (row) in the data frame, there is a method called `iterrows`.
#
#
# -
for index, row in students.iterrows():
print(index, row["DOB"])
# + [markdown] slideshow={"slide_type": "slide"}
# ## Plotting Data Frames
# <a id='PlottingDataFrames'></a>
# You can plot the column or row of a DataFrame with `matplotlib` functions, as we have studied.
# -
students = pd.read_csv('sample_data/sample_student_data.csv', skiprows=[1], index_col=[0])
plt.plot(students.Weight, students.Height, 'o')
# + slideshow={"slide_type": "slide"}
unlabelled = pd.read_csv('sample_data/noHeader_noIndex.csv', header=None)
display(unlabelled)
# + [markdown] slideshow={"slide_type": "slide"}
# Where there are no headers or row names, `iloc` indexing can be used as normal.
# -
# Plot row 1 (y axis) against row 0 (x axis)
plt.plot(unlabelled.iloc[0], unlabelled.iloc[1])
# + [markdown] slideshow={"slide_type": "slide"}
# Where there are no headers or row names, `loc` indexing can uses numerical indices in place of strings (i.e. same as `iloc`):
# +
print(unlabelled.keys())
plt.plot(unlabelled.loc[0], unlabelled.loc[1])
# + [markdown] slideshow={"slide_type": "subslide"}
# To plot all columns against each other:
# +
#import data
headers = ["X","Y"]
unlabelled_vert = pd.read_csv('sample_data/noHeader_noIndex_vert.csv', names=headers)
display(unlabelled_vert)
#plot
unlabelled_vert.plot()
# + [markdown] slideshow={"slide_type": "slide"}
# `Pandas` also has its own plotting functions (based on `matplotlib`, using the *dot* syntax).
#
# Axes names and legend are generated automatically.
#
# The documentation can be found here:
# https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.plot.html
#
# Below are some examples...
# -
unlabelled_vert.plot(kind="bar")
# + slideshow={"slide_type": "subslide"}
unlabelled_vert.plot(kind="barh")
# + slideshow={"slide_type": "subslide"}
students.plot('Height', 'Weight', kind="scatter")
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Adding, Removing and Joining
# <a id='AddingRemovingJoining'></a>
# ### Adding a Column to a Data Frame
# <a id='AddingColumnDataFrame'></a>
# A column may be added to a Data Frame by specifying the keyword and values of the new column in the same way as a new entry is added to a dictionary.
# +
# Add a column called BloodGroup
students["BloodGroup"] = ['B', 'A', 'O', 'B', 'O',
'AB', 'O', 'B', 'A', 'B',
'A', 'A', 'O', 'O', 'A',
'O', 'O', 'A', 'O', 'B', 'B', 'B']
print(students.head())
# Remove the column
del students["BloodGroup"]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Inserting a Column to a Data Frame
# <a id='InsertingColumnDataFrame'></a>
# A column may be inserted at a specific position.
# +
# Add a column called BMI, calculated from two other columns
students.insert(loc=4, column='BMI', value=(students.Height**2 / students.Weight))
print(students.head())
# Remove the column
del students['BMI']
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Adding a Row to a Data Frame
# <a id='AddingRowDataFrame'></a>
# -
# The `.loc` indexing syntax can be used to add a row to a data frame
students.loc['JW-8'] = ['M', '17/04/1996', 1.69, 55.0 , '121/82'] # adding a row
students.loc['JW-13'] = ['M', '17/04/1996', 1.69, 55.0 , '121/82']
print(students.tail())
# + [markdown] slideshow={"slide_type": "subslide"}
# Single or multiple rows can be removed from a data frame.
#
# The keyword argument `inplace` applies the changes to the original data frame.
# + slideshow={"slide_type": "-"}
students = pd.read_csv('sample_data/sample_student_data.csv', skiprows=[1], index_col=0)
display(students.head())
students.drop(index=['JW-1', 'JW-2', 'JW-4'], columns=['DOB'], inplace=True)
display(students.head())
# + [markdown] slideshow={"slide_type": "subslide"}
# Note that if your indexing systen refers to a continuous series of numbers, removing entries may leave holes in the sequence.
#
# It can be convenient to re-assign the indexes once entries have been removed.
# + slideshow={"slide_type": "-"}
# column headings
headers = ["X","Y"]
# import data
df = pd.read_csv('sample_data/noHeader_noIndex_vert.csv', names=headers)
display(df)
# remove two rows
df.drop([1, 3], inplace=True)
display(df)
# re-assign index
df = df.reset_index(drop=True)
display(df)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Resampling
# <a id='Resampling'></a>
# Downsampling refers to going from high frequency to low frequency e.g. daily data to monthly data.
#
# Upsampling refers to going from low frequency to high frequency e.g. monthly data to daily data.
#
# In this example, monthly totals are computed from daily values (downsampling):
# + [markdown] slideshow={"slide_type": "slide"}
# The file `rotterdam_rainfall_2012.txt` contains the default ranifall data for Rotterdam for 2012.
#
# Import the data, using the dates as the index (the column labeled `YYYYMMDD`).
#
#
# + slideshow={"slide_type": "slide"}
rain = pd.read_csv('sample_data/rotterdam_rainfall_2012.txt',
skiprows=9, # skip 9 rows of header data
parse_dates=['YYYYMMDD'], # create datetime object
index_col='YYYYMMDD', # choose the index
skipinitialspace=True) # skips spaces after the delimiter
display(rain.head())
rain.RH[rain.RH < 0] = 0 # remove negative values
display(rain.head())
rain.RH = rain.RH * 0.1 # convert 0.1mm/day to mm/day
display(rain.head())
# + [markdown] slideshow={"slide_type": "slide"}
# `resample` is a method. We apply it to column rain.RH
#
# __Function arguments:__
# - desired sampling frequency <br> `'A'` (annually or yearly), `'M'` (monthly), `'W'` (weekly), `'D'` (daily), `'H'` (hourly). <br>There are many other options: see http://pandas.pydata.org/pandas-docs/version/0.12.0/timeseries.html.
#
# <br>
# - `kind`<br>Tell `pandas` which point in the original scale to assign the computed value to. <br>e.g. last day of the period, first day, entire period ...
#
# <br>
# - Specify how to resample by adding a numpy function e.g. `np.mean` (the default), `np.sum`, `np.min`, `np.max`, etc.
# + slideshow={"slide_type": "slide"}
monthlyrain = rain.RH.resample('M', # Monthly sampling
kind='period' # Assign downsampled value to whole period
).sum() # Sum all values to obtain new value
print(monthlyrain)
# + [markdown] slideshow={"slide_type": "slide"}
# Lastly make a bar graph using `pandas`:
# + slideshow={"slide_type": "-"}
monthlyrain.plot(kind='bar')
plt.ylabel('mm/month')
plt.xlabel('month');
# + [markdown] slideshow={"slide_type": "slide"}
# ### Saving a Data Frame
# <a id='SavingDataFrame'></a>
# The `to_csv` method can be used to save a `DataFrame`.
#
# In theory, this command can be used to save the data as *any* filetype.
#
# Only files that can be opened as unformatted text seperated by a text delimiter (default = , comma) will be able to be read.
#
# Function arguments may be used to format the output file.
#
# As an example, let's store the new, resampled data set, `monthlyrain` to a .csv, .txt, and .xls file:
# + slideshow={"slide_type": "slide"}
monthlyrain.to_csv('sample_data/rotterdam_monthly_rainfall_2012.csv')
monthlyrain.to_csv('sample_data/rotterdam_monthly_rainfall_2012.txt')
monthlyrain.to_excel('sample_data/rotterdam_monthly_rainfall_2012.xls')
# + [markdown] slideshow={"slide_type": "slide"}
# The full list of optional function arguments can be found here: https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.to_csv.html
#
# Five useful optional function arguments as examples:
# - `header` : new headers can be assigend as a list of strings OR the header can be omitted using
# - `float_format` : the number of decimal places
# - `sep`: delimter (default = ",")
# - `mode` : the Python mode specifier (default = `w`)
# `header=False`.
# - `index` : write row names (default = True)
# + slideshow={"slide_type": "-"}
monthlyrain.to_csv('sample_data/rotterdam_monthly_rain_formatted.csv',
header=False,
float_format="%.3f",
sep="\t",
mode="a",
index=False)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Other Ways to Save Program Output
# <a id='OtherWaysSaveProgramOutput'></a>
# Pandas data frames are very useful for handling data that can be represented as a spreadsheet.
#
# However, we need a more generic way to save output data
#
# Examples:
# - saving a block of text
# - saving a small collection of values
# + [markdown] slideshow={"slide_type": "slide"}
# Python has a collection of easy to use functions for reading and writing files:
# - `open()`
# - `read()`
# - `write()`
# - `close()`
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# Before a file can be read or written to, it must be opened using the `open()` function.
#
# __Function arguments:__
# 1. Path to the file (filename and location)
# 2. The *mode specifier* with which to open the file:
#
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# <br>
# - `r`: open an existing file to read
#
# <br>
# - `w`: open an existing file to write to. <br>If no file exists: creates a new file.<br>If file exists : over-writes previous contents.
#
# <br>
# - `a`: open an existing file to write to. <br>If no file exists: creates a new file.<br>If file exists : appends text to end of file.
# + [markdown] slideshow={"slide_type": "slide"}
# - `r+`: open a text file to read from __or__ write to. <br>File must already exist.<br>If file exists : over-writes previous contents.
#
# <br>
# - `w+`: open a text file to read from __or__ write to.<br>If no file exists: creates a new file.<br>If file exists : over-writes previous contents.
#
# <br>
# - `a+` : open a text file to read from __or__ write to.<br>If no file exists: creates a new file.<br>If file exists : appends text to end of file.
#
# + [markdown] slideshow={"slide_type": "slide"}
# Once the file is open, it creates a *file object*.
#
# We can interrogate the file for various details including whether it is open or closed:
#
# - `name`: Name of the opened file
# - `mode`: Mode specifier with which the file was opened
# - `closed`: Boolean True or False
# - `readable( )`: Read permission, Boolean True or False
# - `writable( )`: Write permission, Boolean True or False
# + [markdown] slideshow={"slide_type": "slide"}
# __Example__
#
# Once a file has been opened it's contents can be read, overwritten, or added to, depnding on the mode specifier used to open it.
#
# We are going to open an example file as if we were going to *write* some data to it.
#
# The mode specifier is `w`.
# + slideshow={"slide_type": "-"}
file = open("sample_data/my_file.txt", "w" )
# + [markdown] slideshow={"slide_type": "slide"}
# Once the file is open, we can *interrogate it for information.
# + slideshow={"slide_type": "-"}
# Interrrogate the file:
print( "File Name:", file.name )
print( "Open Mode:", file.mode )
print( "Readable:", file.readable())
print( "Writable:", file.writable())
# + [markdown] slideshow={"slide_type": "slide"}
# An open file must then always be closed again by calling the close() method.
#
# Let's first write a function to show us if a file is open or closed.
# -
# Write a function to determine the file's status:
def get_status( f ):
if ( f.closed != False ) :
return "Closed"
else :
return "Open"
# + [markdown] slideshow={"slide_type": "slide"}
# ##### Why do we need to close a file?
# 1. Python does not automatically close the file. Leaving it open means that you risk overwriting information.
# 1. Closing the file saves any changes to it.
# 1. Depending on your computer's operating system, you may not be able to open a file simultaneously for reading and writing. <br>__Example:__ If a program attempts to open a file that is already open (has not been closed), an error may be generated.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# Let's try out our function.
# + slideshow={"slide_type": "-"}
print( "File Status:" , get_status(file))
file.close()
print( "File Status:" , get_status(file))
# + [markdown] slideshow={"slide_type": "slide"}
# __Try it yourself__
#
# 1. Save the previous 4 code cells contents to a file, `interrogate_file.py`.
#
# 1. Correct the following line to specify the path to `my_file.txt`, *relative* to the location of `interrogate.py`:
# >`file = open("sample_data/my_file.txt’", "w" )`
#
# 1. Run `interrogate_file.py` from the terminal to see the file get created and display information about `my_file.txt`.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Reading and Writing Text Files
# <a id='ReadingWritingTextFiles'></a>
# ### Writing Files
# <a id='WritingFiles'></a>
# Data can be written to a files in a number of ways.
#
# This time, let's actually write some data to the file.
# + [markdown] slideshow={"slide_type": "slide"}
# Let's first create a string to write to a text file:
#
# The inclusion of `\n` anywhere in the string creates a line break.
# + slideshow={"slide_type": "-"}
poem = 'I never saw a man who looked\nWith such a wistful eye\n'
print(poem)
# + [markdown] slideshow={"slide_type": "slide"}
# A string my be *concatenated* (stuck together end to end) using the `+=` arithmetic operator.
#
# This allows us to build the string over the course of the program.
# + slideshow={"slide_type": "-"}
poem += "Upon that little tent of blue\n"
poem += "Which prisoners call the sky\n"
# + [markdown] slideshow={"slide_type": "slide"}
# Create an object to __write__ the string to a file:
# -
file = open("sample_data/poem.txt", "w" )
# + [markdown] slideshow={"slide_type": "slide"}
# Write the string to the file, then close it.
# +
file.write(poem)
file.close()
# + [markdown] slideshow={"slide_type": "slide"}
# You can open the file using a text editor to confirm it's contents.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Reading and Editing Files
# We will now study some more functions to read and edit a file that has been opened using the `read` file specifier.
# - `read()`
# - `seek()`
# - `tell()`
# - `truncate()`
# + [markdown] slideshow={"slide_type": "slide"}
# ##### `read()`
# We studied read earlier in today's class.
#
# By default `read()` will read the entire contents of the file from the beginning (index 0) to the end (index -1).
#
# An optional integer argument can be used to specify *how many characters to read*.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ##### `seek()`
# The position within a file:
# - from which to read or...
# - at which to write
#
# can be selected using `seek()`.
#
# An optional integer argument can be used to specify *how many characters to offset* the start position from the start of the file.
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ##### `tell()`
# The current position within a file can be returned an integer location index at any time using `tell()`.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ##### `truncate()`
# Deletes all characters after the current position.
# + [markdown] slideshow={"slide_type": "slide"}
# __Example :__`read`, `seek()`, `tell()` and `truncate()`.
#
# Let's first write some data to a file:
# -
with open("sample_data/twister.txt" , "w" ) as file :
file.write("How can a clam cram in a clean cream can?")
# + [markdown] slideshow={"slide_type": "slide"}
# Now we are going to read and edit the data.
#
# We first:
# - open the file using a mode specifier to read *and* write
# - print the current position in the file:
# +
file = open("sample_data/twister.txt" , "r+")
print("Position In File Now:", file.tell())
# + [markdown] slideshow={"slide_type": "slide"}
# The position changes, once the file has been read.
# -
# reading the file moves the position
print(file.read())
print("Position In File Now:", file.tell())
# + [markdown] slideshow={"slide_type": "slide"}
# Let's move to a specific position, 31 characters into the file:
#
# *(This looks for a specific __position__. In next week's class we will learn to look for a specific __word__.)*
# -
# seek
file.seek( 31 )
print("Position In File Now:", file.tell())
# + [markdown] slideshow={"slide_type": "slide"}
# Position 31 is just before the words `' cream can?'`.
#
# We can see this by reading the file at this point.
# -
print(file.read())
# + [markdown] slideshow={"slide_type": "slide"}
# We are going to replace:
#
# 'cream can?'
#
# ...with:
#
# 'caravan?'
# + [markdown] slideshow={"slide_type": "slide"}
# We are now at the end of the file.
#
# Let's go back to position 31 and write `'caravan'`.
# -
# write starting at new position
file.seek( 31 )
file.write("caravan?")
# + [markdown] slideshow={"slide_type": "slide"}
# What does the whole file look like now?
#
# We can go back to position 0 and read the file to check:
# -
# return to position 0 and print the file contents
file.seek( 0 )
print(file.read())
# + [markdown] slideshow={"slide_type": "slide"}
# `'cream can?'` has been *overwritten* with `'caravan?'`.
#
# However, `'cream can?'` contains more letters than `'caravan?'` so the additional letters are note overwritten.
# + [markdown] slideshow={"slide_type": "slide"}
# We can use `truncate` to remove any traling charcters:
# -
print(len('caravan?'))
file.seek( 31 + len('caravan?'))
# truncate deletes all characters from current position onwards
file.truncate()
# + [markdown] slideshow={"slide_type": "slide"}
# What does the whole file look like now?
#
# We can go back to position 0 and read the file to check:
# -
# return to position 0 and print the file contents
file.seek( 0 )
print(file.read())
# + [markdown] slideshow={"slide_type": "slide"}
# Finally, remember to close the file:
# -
file.close()
# + [markdown] slideshow={"slide_type": "slide"}
# The full example, using `with`:
# +
# write file
with open("sample_data/twister.txt" , "w" ) as file :
file.write("How can a clam cram in a clean cream can?")
# read and edit
with open("sample_data/twister.txt" , "r+" ) as file :
# tell
print("Position In File Now:", file.tell())
# reading the file moves the position
print(file.read())
print("Position In File Now:", file.tell())
# seek
file.seek( 31 )
print("Position In File Now:", file.tell())
# write starting at new position
file.write("caravan?")
# truncate deletes all characters from current position onwards
file.truncate()
# move back to start poistion
file.seek( 0 )
print("Position In File Now:", file.tell())
# print updated string
print(file.read())
# + [markdown] slideshow={"slide_type": "slide"}
# Alternatively, if we know the word to replace we can use the function `replace()`:
# -
with open("sample_data/twister.txt" , "w+" ) as file :
# write contents
file.write("How can a clam cram in a clean cream can?")
# go to start of file
file.seek( 0 )
# create a new string with the words replaced
data = file.read().replace("cream can", "caravan")
# go to start of file
file.seek( 0 )
# over-write the old string
file.write(data)
file.truncate()
# go to start of file
file.seek( 0 )
print(file.read())
# + [markdown] slideshow={"slide_type": "slide"}
# A maximum file sixe can be specified when using truncate:
# -
with open("sample_data/twister.txt" , "w+" ) as file :
# write contents
file.write("How can a clam cram in a clean cream can?")
# set maximum length of string
file.truncate(10)
# go to start of file
file.seek( 0 )
print(file.read())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Binary Files ("Pickling" Data)
# <a id='BinaryFilesPicklingData'></a>
# String data can easily be stored in text files, as shown.
#
# Other data types (e.g. numbers, lists, dictionaries) can also be stored in text files but require conversion to strings first.
#
# Restoring that stored data to their original data type, if read into a Python file, requires the opposite conversion.
# + [markdown] slideshow={"slide_type": "slide"}
# Unless the storage file needs to be human-readable for some reason, it is more efficient to use `pickle` to store data as a machine-readable binary file.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### Advantages of pickling data.
#
# A binary file is essentially a computer file that __stored__ in a non-plain text file format(e.g. .py, .csv, .txt.....etc).
#
# A character stream is used to represent (typically non-text) information.
#
# <img src="img/binary-file-in-text-editor.png" alt="Drawing" style="width: 300px;"/>
# + [markdown] slideshow={"slide_type": "slide"}
# This character stream contains all the information necessary to reconstruct the object in another (python) script.
#
# Many binary file formats contain parts that can be interpreted as text:
#
# __Example:__ A computer document file may contain:
# - un-formatted text
# - formatting information
#
#
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# __Serialising__
# <br>Translating data structures or object state into a format that can be stored or transmitted and reconstructed later.
#
# The serialised character stream contains all the information necessary to reconstruct the object in another (python) script / computer environment.
# + [markdown] slideshow={"slide_type": "slide"}
# Pickling a python object serialises the object:
# - converting it to a character stream
# - allowing it to be stored at a smaller size
# + [markdown] slideshow={"slide_type": "slide"}
#
#
# Read (`r`), write (`w`) and append (`a`) can be used to produce binary files simply by using the mode specifiers:
# - `rb` : read
# - `wb` : write
# - `ab` : append
# + [markdown] slideshow={"slide_type": "slide"}
# “Pickling” a data object stores a string representation of the object.
#
# The string representation can later be “unpickled” to its former state.
# + [markdown] slideshow={"slide_type": "slide"}
# The following example shows how to pickle data.
# + [markdown] slideshow={"slide_type": "slide"}
# Import the `pickle` package:
# -
import pickle
# + [markdown] slideshow={"slide_type": "slide"}
# Create sample data:
# +
data = [ "Numbers" ,
[1, 2, 3, 4, 5] ]
print(data)
# + [markdown] slideshow={"slide_type": "slide"}
# In the `with` block:
# - Use the function `pickled.dump()`
# + slideshow={"slide_type": "-"}
with open( "sample_data/pickled.dat" , "wb" ) as file:
pickle.dump( data, file )
# + [markdown] slideshow={"slide_type": "slide"}
# In the `with` block:
# - Use the function `pickled.load()`
# +
with open( "sample_data/pickled.dat" , "rb" ) as file:
data = pickle.load(file)
print(f"recovered data: \n{data[0]}, type = {type(data[0])} \n{data[1]}, type = {type(data[1])} ")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Pickling a Pandas Data Frame
# <a id='PicklingPandasDataFrame'></a>
# `pandas DataFrame`s have a built-in method, `to_pickle`, that can be used to pickle the data that they contain.
#
# To demonstrate, let's use the `pandas`, `read_csv` function to read in the sample student data again:
# +
students = pd.read_csv('sample_data/sample_student_data.csv',
skiprows=[1],
index_col=0)
print(students.keys())
# + [markdown] slideshow={"slide_type": "slide"}
# The cell below shows how to pickle (save) and read back the `students DataFrame`.
# -
students.to_pickle("sample_data/pickled_student_data.dat")
# + slideshow={"slide_type": "slide"}
from pandas import read_pickle
read_back = read_pickle("sample_data/pickled_student_data.dat")
print(read_back)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Saving Data with Numpy
# <a id='SavingDataNumpy'></a>
# `numpy` provides a few options for saving data that has the form of a Numpy array only.
#
# Example array:
# +
# This is a numpy array
A = np.zeros(4)
# This is not a numpy array
# A = ["kit", "kat"]
# + [markdown] slideshow={"slide_type": "slide"}
# ##### `numpy.save()`
#
# You can use the argument `fix_imports` (default = `False`).
# <br>If `True` this will make sure your data is pickled in a way that is readable in Python 2.
#
# The file is stored as a binary file with extension `.npy`:
# - you do not need to specify a filetype.
# - this can only be opened using Python (numpy).
#
# The file is generally smaller than standard pickling (112 bytes vs 187 bytes) for this example.
# + slideshow={"slide_type": "-"}
np.save("sample_data/np_save", A)
B = np.load("sample_data/np_save.npy")
print(B)
# + [markdown] slideshow={"slide_type": "slide"}
# ##### `numpy.dump()`
# Numpy arrays also have a method:
# - concise
# - file type that is accessible outside of numpy
# - larger (this example 240 bytes)
# +
A.dump("sample_data/np_dump.dat")
B = np.load("sample_data/np_dump.dat")
print(B)
# + [markdown] slideshow={"slide_type": "slide"}
# ##### `numpy.savetxt`
# Lastly, to save a numpy array containing *only numerical values*:
# - in human readable form.
# - at small size (this examples 100 bytes)
#
# +
# np.savetxt("sample_data/np_savetxt.txt", A)
np.savetxt("sample_data/np_savetxt.txt", A)
B = np.loadtxt("sample_data/np_savetxt.txt")
print(B)
# + [markdown] slideshow={"slide_type": "slide"}
# # Summary
# <a id='Summary'></a>
# - Data from external files (e.g. .csv files) can be imported using the standard Python library.
# - A file object has open(), read(), write(), and close() methods for working with files, and features that describe the file properties
# - The open() method must specify a file name string argument and a file mode string argument, such as ’r’ to read the file.
# - Position in a file, at which to read or write, can be specified with the seek() method and reported by the tell() method the Python with keyword groups file operational statements within a block and automatically closes an open file.
# - Pandas DataFrames can be used to store and manipulate data for anaysis.
# - The process of “pickling” objects stores a string representation of an object that can later be “unpickled” to its former state.
# - A pickle object’s dump() method requires arguments to specify an object for conversion and a le name in which to store data.
# - Stored object data can be retrieved by specifying the le name in which it is stored to the pickle object’s load() method.
#
# *(Adapted from) McGrath 2016*
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='TestYourselfExercises'></a>
# # Test-Yourself Exercises
#
# Compete the Test-Youself exercises below.
#
# Save your answers as .py files and email them to:
# <br><EMAIL>
# + [markdown] slideshow={"slide_type": "slide"}
# ### Test-Yourself Excercise: Sorting Data Frames.
#
# 1. Import the necessary libraries
#
# 1. Import the dataset from https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv to a Pandas data frame.
#
# 1. Print the names of all the columns
#
# 1. Remove all duplicate items keeping only the first to appear in the data frame.
#
# 1. Remove all items without a choice_description from the data set.
#
# 1. Cast all prices as float data. <br>*Hint: Currently each price is a __string__ of which the __last 4 characters__ are a floating point number*.
#
# 1. What is the price of each item? Generate a *new data frame* with only two columns item_name and item_priceipo.
#
# 1. Sort this data frame by price in descending order.
#
# 1. Save the new data frame as a .csv file.
# +
# Example Solution
# 1. Import the necessary libraries
import pandas as pd
# 2. Import the dataset
url = 'https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv'
chipo = pd.read_csv(url, sep = '\t')
# 3. Print the names of all the columns
print(chipo.columns)
# 4. Remove all duplicate items
chipo.drop_duplicates('item_name', inplace=True)
# 5. Remove all items without a choice_description
chipo.dropna()
# 6. Cast all prices as float data.
chipo.item_price = chipo.item_price.apply(lambda x : float(x[1:-1]))
# 7. Generate a new data frame with only two columns item_name and item_priceipo
price_per_item = chipo[['item_name', 'item_price']]
# 8. Sort this data frame by price in descending order.
price_per_item = price_per_item.sort_values(by = "item_price", ascending = False)
# 9. Save the new data frame as a .csv file.
price_per_item.to_csv('sample_data/price_per_item.csv')
# -
# + [markdown] slideshow={"slide_type": "slide"}
# ### Test-Yourself Excercise: Appending text files.
#
# Find out the origin of the quote in `dot.txt`.
#
# Write Python code directly into the terminal to append the file `dot.txt` with a citation.
# -
# Example Solution
file = open("sample_data/dot.txt", "a" )
file.write('\n<NAME>, speech at Cornell University, October 13, 1994')
file.close()
| 06_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Global Fishing Effort
import time
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
import matplotlib
from matplotlib import colors,colorbar
import matplotlib
# %matplotlib inline
import csv
import math
from math import radians, cos, sin, asin, sqrt
# from scipy import stats
import matplotlib.dates as mdates
import argparse
import googleapiclient
from googleapiclient.discovery import build
from googleapiclient.errors import HttpError
from oauth2client.client import GoogleCredentials
credentials = GoogleCredentials.get_application_default()
# Construct the service object for interacting with the BigQuery API.
bigquery_service = build('bigquery', 'v2', credentials=credentials)
# create a bounding box:
max_lat = 90
min_lat = -90
max_lon = 180
min_lon = -180
cellsize = .5
def Query(q):
query_request = bigquery_service.jobs()
query_data = {
'query': (q)
}
query_response = query_request.query(
projectId='world-fishing-827',
body=query_data).execute()
rows = []
for row in query_response['rows']:
s = [item['v'] for item in row['f']]
rows.append(s)
return rows
# +
q = '''
select count(distinct mmsi)
FROM
[tilesets.pipeline_2015_08_24_08_19_01]
WHERE
latitude > '''+str(min_lat)+'''
AND latitude <'''+str(max_lat)+'''
AND longitude > '''+str(min_lon)+'''
AND longitude < '''+str(max_lon)+'''
AND weight >=.5'''
number_of_mmsi = Query(q)
# -
print "Number of unique MMSI:",number_of_mmsi[0][0]#int(number_of_mmsi[0]['f'][0]['v'])
# # 66,245 MMSI were fishing between January 2014 to July 2015
# # Map the Fishing Effort in This Region
# +
# Now Group by lat, lon
q = '''
SELECT
lat,
lon,
count(*) fishing_days,
avg(distance_to_shore) distance_to_shore,
first(eez) eez
from
(SELECT
mmsi,
date(timestamp) date,
integer(first(latitude)*2) lat,
integer(first(longitude)*2) lon,
first(FLOAT(JSON_EXTRACT(extra,"$.distance_to_shore"))) distance_to_shore,
first(JSON_EXTRACT(extra,"$.eez")) EEZ,
FROM
[tilesets.pipeline_2015_08_24_08_19_01]
WHERE
latitude > -90
AND latitude <90
AND longitude > -180
AND longitude < 180
AND weight >=.5
and mmsi != 220364000 //this one is bad
and mmsi not in(412437961,412437962,412420502,412420503,412420576,412420574,412420789,412420871,
900025357,900025393,413322650,414203586,412211196,412440255,412440256,412440257,412440258,412440259,
412440261,150200162,412440077,412440078,412420805,412420421,412440377,412425706,412447093) // these
// are mmsi that were spoofing according to Bjorn
and JSON_EXTRACT(extra,"$.distance_to_shore") is not Null
group by mmsi, date)
group by lat, lon
'''
fishing_grid = Query(q)
# +
cellsize = .5
one_over_cellsize = 2
num_lats = (max_lat-min_lat)*one_over_cellsize
num_lons = (max_lon-min_lon)*one_over_cellsize
grid = np.zeros(shape=(num_lats,num_lons))
for row in fishing_grid:
lat = int(row[0])
lon = int(row[1])
lat_index = lat-min_lat*one_over_cellsize
lon_index = lon-min_lon*one_over_cellsize
grid[lat_index][lon_index] = int(row[2])
# +
plt.rcParams["figure.figsize"] = [12,7]
cutoff = 0 # 4 degress away from the pole
firstlat = 90-cutoff
lastlat = -90+cutoff
firstlon = -180
lastlon = 180
scale = cellsize
one_over_cellsize = 2
fishing_days_truncated = grid[one_over_cellsize*cutoff:(180*one_over_cellsize)-cutoff*one_over_cellsize][:]
numlats = int((firstlat-lastlat)*one_over_cellsize+.5)
numlons = int((lastlon-firstlon)*one_over_cellsize+.5)
lat_boxes = np.linspace(lastlat,firstlat,num=numlats,endpoint=False)
lon_boxes = np.linspace(firstlon,lastlon,num=numlons,endpoint=False)
fig = plt.figure()
m = Basemap(llcrnrlat=lastlat, urcrnrlat=firstlat,
llcrnrlon=lastlon, urcrnrlon=firstlon, lat_ts=0, projection='robin',resolution="h", lon_0=0)
m.drawmapboundary(fill_color='#111111')
# m.drawcoastlines(linewidth=.2)
m.fillcontinents('#111111',lake_color='#111111')#, lake_color, ax, zorder, alpha)
x = np.linspace(-180, 180, 360*one_over_cellsize)
y = np.linspace(lastlat, firstlat, (firstlat-lastlat)*one_over_cellsize)
x, y = np.meshgrid(x, y)
converted_x, converted_y = m(x, y)
from matplotlib import colors,colorbar
maximum = 100000
minimum = 1
norm = colors.LogNorm(vmin=minimum, vmax=maximum)
# norm = colors.Normalize(vmin=0, vmax=1000)
m.pcolormesh(converted_x, converted_y, fishing_days_truncated, norm=norm, vmin=minimum, vmax=maximum, cmap = plt.get_cmap('viridis'))
t = "Fishing Days, January 2014 to July 2015"
plt.title(t, color = "#ffffff", fontsize=18)
ax = fig.add_axes([0.2, 0.1, 0.4, 0.02]) #x coordinate ,
norm = colors.LogNorm(vmin=minimum, vmax=maximum)
# norm = colors.Normalize(vmin=0, vmax=1000)
lvls = np.logspace(np.log10(minimum),np.log10(maximum),num=8)
cb = colorbar.ColorbarBase(ax,norm = norm, orientation='horizontal', ticks=lvls, cmap = plt.get_cmap('viridis'))
the_labels = []
for l in lvls:
if l>=1:
l = int(l)
the_labels.append(l)
#cb.ax.set_xticklabels(["0" ,round(m3**.5,1), m3, round(m3**1.5,1), m3*m3,round(m3**2.5,1), str(round(m3**3,1))+"+"], fontsize=10)
cb.ax.set_xticklabels(the_labels, fontsize=10, color = "#ffffff")
cb.set_label('Number of Fishing Days, January 2014 to July 2015',labelpad=-40, y=0.45, color = "#ffffff")
ax.text(1.7, -0.5, 'Data Source: Orbcomm\nMap by Global Fishing Watch',
verticalalignment='bottom', horizontalalignment='right',
transform=ax.transAxes,
color='#ffffff', fontsize=6)
plt.savefig("fishing_days_2014-2015.png",bbox_inches='tight',dpi=300,transparent=True,pad_inches=.1, facecolor="#000000")
plt.show()
# -
grid = np.array(grid)
# +
avg_by_lat = np.sum(grid, axis=1)
lat = [(i-180)/2. for i in range(180*2)]
ind = np.arange(len(lat)) # the x locations for the groups
width = 1 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, avg_by_lat, width, color='b')
ax.set_ylabel('Number of Fishing Days')
ax.set_xlabel('Latitude')
ax.set_title('Fishing Days versus Latitude')
plt.xticks(np.arange(0,360*2,30*2))
ax.set_xticklabels((-90,-60,-30,0,30,60,90))
plt.xlim([0,360])
plt.show()
# +
avg_by_lon = np.sum(grid, axis=0)
lon = [(i-360)/2. for i in range(360*2)]
ind = np.arange(len(lon)) # the x locations for the groups
width = 1 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, avg_by_lon, width, color='b')
ax.set_ylabel('Number of Fishing Days')
ax.set_xlabel('Longitude')
ax.set_title('Fishing Days versus Longitude')
plt.xticks(np.arange(0,360*2+1,30*2))
ax.set_xticklabels((i for i in range(-180, 181, 30)))
plt.xlim([0,720])
plt.show()
# -
fishings = []
distance_to_shore = []
for r in fishing_grid:
fishings.append(int(r[2]))
distance_to_shore.append(float(r[3]))
distance_to_shore = np.array(distance_to_shore)
fishings = np.array(fishings)
distance_to_shore.max()
distances = np.zeros(int(distance_to_shore.max()/50)+1)
for d,f in zip(distance_to_shore,fishings):
distances[int(d/50)] += f
ind = np.arange(len(distances)) # the x locations for the groups
width = 1 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, distances, width, color='b')
ax.set_ylabel('Number of Fishing Days')
ax.set_xlabel('Distance from Shore')
ax.set_title('Fishing Days by Distance from Shore, km')
plt.xticks(np.arange(0,51,10))
ax.set_xticklabels((i*50 for i in range(0,51,10)))
plt.show()
#log log!
x = [i*50+25 for i in range(len(distances))]
y = distances
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.set_yscale('log')
ax.set_xscale('log')
plt.scatter(x, y, alpha=.8)#, color = color)
plt.title("Fishing Days by Distance from Shore")
plt.xlabel('Distance from Shore, km')
plt.ylabel('Number of Fishing Days')
plt.show()
# # Fishing effort in the High Seas versus EEZs
in_eezs=0
high_seas=0
for r in fishing_grid:
if r[4]:
in_eezs+=int(r[2])
else:
high_seas+=int(r[2])
print int(100*high_seas/float(in_eezs+high_seas)), "percent of fishing days are in the high seas"
# # Only 5 percent of the fishing days are in the high seas!
#
# # Okay, now make histogram of lat and lon for high seas only
# +
cellsize = .5
one_over_cellsize = 2
num_lats = (max_lat-min_lat)*one_over_cellsize
num_lons = (max_lon-min_lon)*one_over_cellsize
grid_highseas = np.zeros(shape=(num_lats,num_lons))
for row in fishing_grid:
lat = int(row[0])
lon = int(row[1])
if not row[4]:
lat_index = lat-min_lat*one_over_cellsize
lon_index = lon-min_lon*one_over_cellsize
grid_highseas[lat_index][lon_index] = int(row[2])
# +
avg_by_lat = np.sum(grid_highseas, axis=1)
lat = [(i-180)/2. for i in range(180*2)]
ind = np.arange(len(lat)) # the x locations for the groups
width = 1 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, avg_by_lat, width, color='b')
ax.set_ylabel('Number of Fishing Days')
ax.set_xlabel('Latitude')
ax.set_title('Fishing Days versus Latitude, High Seas')
plt.xticks(np.arange(0,360*2,30*2))
ax.set_xticklabels((-90,-60,-30,0,30,60,90))
plt.xlim([0,360])
plt.show()
# -
avg_by_lon = np.sum(grid_highseas, axis=0)
lon = [(i-360)/2. for i in range(360*2)]
ind = np.arange(len(lon)) # the x locations for the groups
width = 1 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, avg_by_lon, width, color='b')
ax.set_ylabel('Number of Fishing Days')
ax.set_xlabel('Longitude')
ax.set_title('Fishing Days versus Longitude, High Seas')
plt.xticks(np.arange(0,360*2+1,30*2))
ax.set_xticklabels((i for i in range(-180, 181, 30)))
plt.xlim([0,720])
plt.show()
| initial_maps/Global_Fishing_Effort_v0.ipynb |
# ---
# jupytext:
# text_representation:
# extension: .md
# format_name: myst
# format_version: '0.10'
# jupytext_version: 1.5.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numerical methods
#
# My (now finished) PhD student <NAME> has been looking at numerical methods to solve
# saddle point systems arising from trace constraints coupling 2D and 1D domains,
# or 3D and 1D domains {cite}`kuchta2016,kuchta2016preconditioning,kuchta15`.
#
# We have also been studying the singular Neumann problem of linear elasticity
# {cite}`kuchta2016singular`.
# Four different formulations of the problem have been analyzed and mesh independent preconditioners established for the resulting linear systems within the framework of operator preconditioning. We have proposed a preconditioner for the (singular) mixed formulation of linear elasticity, that is robust with respect to the material parameters. Using an orthonormal basis of the space of rigid motions, discrete projection operators have been derived and employed in a modification to the conjugate gradients method to ensure optimal error convergence of the solution.
#
# With colleagues at the Extreme Computing Research Center (ECRC), King Abdullah University of Science and Technology (KAUST), we have been using [spectralDNS](https://github.com/spectralDNS) to investigate time integration of Fourier pseudospectral Direct Numerical Simulations {cite}`ketcheson2020`. We investigate the use of higher‐order Runge‐Kutta pairs and automatic step size control based on local error estimation. We find that the fifth‐order accurate Runge‐Kutta pair of Bogacki and Shampine gives much greater accuracy at a significantly reduced computational cost.
# ## References
#
# ```{bibliography} ../../references.bib
# :filter: docname in docnames
# ```
| _build/jupyter_execute/content/research/numerical.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="rkkKCW0qWL_0" colab_type="code" colab={}
# #!pip install datadotworld
# #!pip install datadotworld[pandas]
# + id="7wHUcbk0ZAtm" colab_type="code" colab={}
# #!dw configure
# + id="KyhIOt-TX78c" colab_type="code" colab={}
from google.colab import drive
import pandas as pd
import numpy as np
import datadotworld as dw
# + id="KC-d4hMQZWLW" colab_type="code" colab={}
#drive.mount("/content/drive")
# + id="IFSMJ4K2Zvdr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="53755af6-4ec4-48ed-f7ba-cbe6bb83a35e" executionInfo={"status": "ok", "timestamp": 1581544659464, "user_tz": -60, "elapsed": 610, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
# cd "drive/My Drive/Colab Notebooks/dw_matrix"
# + id="x_h18r6zZytk" colab_type="code" colab={}
# !mkdir data
# + id="Xo8wadE-aLwc" colab_type="code" colab={}
# !echo 'data' > .gitignore
# + id="SQjZFQMfauX_" colab_type="code" colab={}
# !git add .gitignore
# + id="nV94fQq2dd1C" colab_type="code" colab={}
data = dw.load_dataset('datafiniti/mens-shoe-prices')
# + id="Q-vuY8h6eKTT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7e415b68-bc7b-48f3-ac14-ae062705043a" executionInfo={"status": "ok", "timestamp": 1581545733108, "user_tz": -60, "elapsed": 2091, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
data
# + id="6eNw48ZZeLI3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="7fad52d9-015e-4680-c1e9-f8cda51dca3e" executionInfo={"status": "ok", "timestamp": 1581545833915, "user_tz": -60, "elapsed": 1962, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
df = data.dataframes['7004_1']
df.shape
# + id="fdo09rI2eo7e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 598} outputId="a826966a-115a-4856-f264-275aaed0f0fa" executionInfo={"status": "ok", "timestamp": 1581545901560, "user_tz": -60, "elapsed": 718, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
df.sample(5)
# + id="XaTCuSvaezVq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="c349d16f-eac1-477a-ff78-8eb0c3c1b925" executionInfo={"status": "ok", "timestamp": 1581545936876, "user_tz": -60, "elapsed": 681, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
df.columns
# + id="TlsgclnBgVAq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="40470174-97d5-4b42-bdff-6bb5ae10056d" executionInfo={"status": "ok", "timestamp": 1581546331817, "user_tz": -60, "elapsed": 673, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
df.prices_currency.unique()
# + id="EFoSV2cGgh1O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="f607f924-47d1-4955-cb39-950911f8e6b3" executionInfo={"status": "ok", "timestamp": 1581546413278, "user_tz": -60, "elapsed": 705, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
df.prices_currency.value_counts(normalize=True)
# + id="EfFOFRuAg3mf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aabc2bea-312d-43bb-b892-28b0bf4c6ca4" executionInfo={"status": "ok", "timestamp": 1581546624157, "user_tz": -60, "elapsed": 690, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
df_usd = df[ df.prices_currency == 'USD' ].copy()
df_usd.shape
# + id="kk8yggw_h1Be" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="d7d164a3-170a-4db2-fdf8-2498c800591d" executionInfo={"status": "ok", "timestamp": 1581546901043, "user_tz": -60, "elapsed": 664, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
df_usd['prices_amountmin'] = df_usd.prices_amountmin.astype(np.float)
df_usd['prices_amountmin'].hist()
# + id="u3PblTa3ijOR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a972f3c1-3463-4fa2-e5e3-8e26a0c391ea" executionInfo={"status": "ok", "timestamp": 1581547069445, "user_tz": -60, "elapsed": 666, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
filter_max = np.percentile(df_usd['prices_amountmin'],99)
filter_max
# + id="SA_yrL3jjAYU" colab_type="code" colab={}
df_usd_filter = df_usd[df_usd['prices_amountmin'] < filter_max]
# + id="9lYH5aAdjnbn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="0ffa03ed-68ba-4c8f-ad0e-00c8c6a96762" executionInfo={"status": "ok", "timestamp": 1581547263003, "user_tz": -60, "elapsed": 922, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
df_usd_filter.prices_amountmin.hist(bins=100)
# + id="EBKpDzDgqdYo" colab_type="code" colab={}
# + id="wy20dRfJkhW1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e139e7ff-39ef-47bd-fed3-c6aac84f9c9b" executionInfo={"status": "ok", "timestamp": 1581547400727, "user_tz": -60, "elapsed": 1882, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
# ls
# + id="zhlQDxKukiZh" colab_type="code" colab={}
df.to_csv('data/shoes_prices.csv', index=False)
# + id="6A6hsRZGlK2b" colab_type="code" colab={}
# !git add matrix_one/day3.ipynb
# + id="T4bbAAPtl7OM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="ca2b0cba-b799-4a8a-feb2-92661bbfb04a" executionInfo={"status": "ok", "timestamp": 1581547953558, "user_tz": -60, "elapsed": 2962, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
# !git commit -m "Read Men's Shoe Prices dataset from data.world"
# + id="nTZan4AemXta" colab_type="code" colab={}
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "Eeyore"
# + id="idRnKDzEnxhF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="37d16948-1395-4333-ccf1-061392f705d1" executionInfo={"status": "ok", "timestamp": 1581548280634, "user_tz": -60, "elapsed": 5644, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
# !git push -u origin master
# + id="V5iqau55qemu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="6971a909-9a48-481a-8530-fe7e16a868e7" executionInfo={"status": "ok", "timestamp": 1581549699497, "user_tz": -60, "elapsed": 2047, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
# !git reset --hard<1d04b1facc4945060e2ee2ef33334ebdcb1252f6>
# + id="kzSZUX9ItWwL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="a4d824ef-76c6-412a-8cbb-816cb70aca73" executionInfo={"status": "ok", "timestamp": 1581549737889, "user_tz": -60, "elapsed": 2272, "user": {"displayName": "Eeyore", "photoUrl": "", "userId": "04930882641739340690"}}
# !git push -u origin master --force
| matrix_one/day3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 17. Making Complex Decisions
# **17.1** \[mdp-model-exercise\]For the $4\times 3$ world shown in
# Figure [sequential-decision-world-figure](#/), calculate
# which squares can be reached from (1,1) by the action sequence
# $[{Up},{Up},{Right},{Right},{Right}]$ and with what
# probabilities. Explain how this computation is related to the prediction
# task (see Section [general-filtering-section](#/)) for a
# hidden Markov model.
#
# **17.2** \[mdp-model-exercise\]For the $4\times 3$ world shown in
# Figure [sequential-decision-world-figure](#/), calculate
# which squares can be reached from (1,1) by the action sequence
# $[{Right},{Right},{Right},{Up},{Up}]$ and with what
# probabilities. Explain how this computation is related to the prediction
# task (see Section [general-filtering-section](#/)) for a
# hidden Markov model.
#
# **17.3** Select a specific member of the set of policies that are optimal for
# $R(s)>0$ as shown in
# Figure [sequential-decision-policies-figure](#/)(b), and
# calculate the fraction of time the agent spends in each state, in the
# limit, if the policy is executed forever. (*Hint*:
# Construct the state-to-state transition probability matrix corresponding
# to the policy and see
# Exercise [markov-convergence-exercise](#/).)
#
# **17.4** \[nonseparable-exercise\]Suppose that we define the utility of a state
# sequence to be the *maximum* reward obtained in any state
# in the sequence. Show that this utility function does not result in
# stationary preferences between state sequences. Is it still possible to
# define a utility function on states such that MEU decision making gives
# optimal behavior?
#
# **17.5** Can any finite search problem be translated exactly into a Markov
# decision problem such that an optimal solution of the latter is also an
# optimal solution of the former? If so, explain *precisely*
# how to translate the problem and how to translate the solution back; if
# not, explain *precisely* why not (i.e., give a
# counterexample).
#
# **17.6** \[reward-equivalence-exercise\] Sometimes MDPs are formulated with a
# reward function $R(s,a)$ that depends on the action taken or with a
# reward function $R(s,a,s')$ that also depends on the outcome state.
#
# 1. Write the Bellman equations for these formulations.
#
# 2. Show how an MDP with reward function $R(s,a,s')$ can be transformed
# into a different MDP with reward function $R(s,a)$, such that
# optimal policies in the new MDP correspond exactly to optimal
# policies in the original MDP.
#
# 3. Now do the same to convert MDPs with $R(s,a)$ into MDPs with $R(s)$.
#
# **17.7** \[threshold-cost-exercise\]For the environment shown in
# Figure [sequential-decision-world-figure](#/), find all the
# threshold values for $R(s)$ such that the optimal policy changes when
# the threshold is crossed. You will need a way to calculate the optimal
# policy and its value for fixed $R(s)$. (*Hint*: Prove that
# the value of any fixed policy varies linearly with $R(s)$.)
#
# **17.8** \[vi-contraction-exercise\]
# Equation ([vi-contraction-equation](#/)) on
# page [vi-contraction-equation](#/) states that the Bellman operator is a contraction.
#
# 1. Show that, for any functions $f$ and $g$,
# $$|\max_a f(a) - \max_a g(a)| \leq \max_a |f(a) - g(a)|\ .$$
#
# 2. Write out an expression for $|(B\,U_i - B\,U'_i)(s)|$ and then apply
# the result from (a) to complete the proof that the Bellman operator
# is a contraction.
#
# **17.9** This exercise considers two-player MDPs that correspond to zero-sum,
# turn-taking games like those in
# Chapter [game-playing-chapter](#/). Let the players be $A$
# and $B$, and let $R(s)$ be the reward for player $A$ in state $s$. (The
# reward for $B$ is always equal and opposite.)
#
# 1. Let $U_A(s)$ be the utility of state $s$ when it is $A$’s turn to
# move in $s$, and let $U_B(s)$ be the utility of state $s$ when it is
# $B$’s turn to move in $s$. All rewards and utilities are calculated
# from $A$’s point of view (just as in a minimax game tree). Write
# down Bellman equations defining $U_A(s)$ and $U_B(s)$.
#
# 2. Explain how to do two-player value iteration with these equations,
# and define a suitable termination criterion.
#
# 3. Consider the game described in
# Figure [line-game4-figure](#/) on page [line-game4-figure](#/).
# Draw the state space (rather than the game tree), showing the moves
# by $A$ as solid lines and moves by $B$ as dashed lines. Mark each
# state with $R(s)$. You will find it helpful to arrange the states
# $(s_A,s_B)$ on a two-dimensional grid, using $s_A$ and $s_B$ as
# “coordinates.”
#
# 4. Now apply two-player value iteration to solve this game, and derive
# the optimal policy.
#
# <center>
# <b id="grid-mdp-figure">Figure [grid-mdp-figure]</b> (a) $3 \times 3$ world for Exercise [3x3-mdp-exercise](#/). The reward for each state is indicated. The upper right square is a terminal state. (b) $101 \times 3$ world for Exercise [101x3-mdp-exercise](#/) (omitting 93 identical columns in the middle).
# The start state has reward 0.
# </center>
#
# 
#
# **17.10** \[3x3-mdp-exercise\] Consider the $3 \times 3$ world shown in
# Figure [grid-mdp-figure](#grid-mdp-figure)(a). The transition model is the
# same as in the $4\times 3$
# Figure [sequential-decision-world-figure](#/): 80% of the
# time the agent goes in the direction it selects; the rest of the time it
# moves at right angles to the intended direction.
#
# Implement value iteration for this world for each value of $r$ below.
# Use discounted rewards with a discount factor of 0.99. Show the policy
# obtained in each case. Explain intuitively why the value of $r$ leads to
# each policy.
#
# 1. $r = -100$
#
# 2. $r = -3$
#
# 3. $r = 0$
#
# 4. $r = +3$
#
# **17.11** \[101x3-mdp-exercise\] Consider the $101 \times 3$ world shown in
# Figure [grid-mdp-figure](#grid-mdp-figure)(b). In the start state the agent
# has a choice of two deterministic actions, *Up* or
# *Down*, but in the other states the agent has one
# deterministic action, *Right*. Assuming a discounted reward
# function, for what values of the discount $\gamma$ should the agent
# choose *Up* and for which *Down*? Compute the
# utility of each action as a function of $\gamma$. (Note that this simple
# example actually reflects many real-world situations in which one must
# weigh the value of an immediate action versus the potential continual
# long-term consequences, such as choosing to dump pollutants into a
# lake.)
#
# **17.12** Consider an undiscounted MDP having three states, (1, 2, 3), with
# rewards $-1$, $-2$, $0$, respectively. State 3 is a terminal state. In
# states 1 and 2 there are two possible actions: $a$ and $b$. The
# transition model is as follows:
#
# - In state 1, action $a$ moves the agent to state 2 with probability
# 0.8 and makes the agent stay put with probability 0.2.
#
# - In state 2, action $a$ moves the agent to state 1 with probability
# 0.8 and makes the agent stay put with probability 0.2.
#
# - In either state 1 or state 2, action $b$ moves the agent to state 3
# with probability 0.1 and makes the agent stay put with
# probability 0.9.
#
# Answer the following questions:
#
# 1. What can be determined *qualitatively* about the
# optimal policy in states 1 and 2?
#
# 2. Apply policy iteration, showing each step in full, to determine the
# optimal policy and the values of states 1 and 2. Assume that the
# initial policy has action $b$ in both states.
#
# 3. What happens to policy iteration if the initial policy has action
# $a$ in both states? Does discounting help? Does the optimal policy
# depend on the discount factor?
#
# **17.13** Consider the $4\times 3$ world shown in
# Figure [sequential-decision-world-figure](#/).
#
# 1. Implement an environment simulator for this environment, such that
# the specific geography of the environment is easily altered. Some
# code for doing this is already in the online code repository.
#
# 2. Create an agent that uses policy iteration, and measure its
# performance in the environment simulator from various
# starting states. Perform several experiments from each starting
# state, and compare the average total reward received per run with
# the utility of the state, as determined by your algorithm.
#
# 3. Experiment with increasing the size of the environment. How does the
# run time for policy iteration vary with the size of the environment?
#
# **17.14** \[policy-loss-exercise\]How can the value determination algorithm be
# used to calculate the expected loss experienced by an agent using a
# given set of utility estimates ${U}$ and an estimated
# model ${P}$, compared with an agent using correct values?
#
# **17.15** \[4x3-pomdp-exercise\] Let the initial belief state $b_0$ for the
# $4\times 3$ POMDP on page [4x3-pomdp-page](#/) be the uniform distribution
# over the nonterminal states, i.e.,
# $< \frac{1}{9},\frac{1}{9},\frac{1}{9},\frac{1}{9},\frac{1}{9},\frac{1}{9},\frac{1}{9},\frac{1}{9},\frac{1}{9},0,0 >$.
# Calculate the exact belief state $b_1$ after the agent moves and its
# sensor reports 1 adjacent wall. Also calculate $b_2$ assuming that the
# same thing happens again.
#
# **17.16** What is the time complexity of $d$ steps of POMDP value iteration for a
# sensorless environment?
#
# **17.17** \[2state-pomdp-exercise\] Consider a version of the two-state POMDP on
# page [2state-pomdp-page](#/) in which the sensor is 90% reliable in state 0 but
# provides no information in state 1 (that is, it reports 0 or 1 with
# equal probability). Analyze, either qualitatively or quantitatively, the
# utility function and the optimal policy for this problem.
#
# **17.18** \[dominant-equilibrium-exercise\]Show that a dominant strategy
# equilibrium is a Nash equilibrium, but not vice versa.
#
# **17.19** In the children’s game of rock–paper–scissors each player reveals at the
# same time a choice of rock, paper, or scissors. Paper wraps rock, rock
# blunts scissors, and scissors cut paper. In the extended version
# rock–paper–scissors–fire–water, fire beats rock, paper, and scissors;
# rock, paper, and scissors beat water; and water beats fire. Write out
# the payoff matrix and find a mixed-strategy solution to this game.
#
# **17.20** Solve the game of *three*-finger Morra.
#
# **17.21** In the *Prisoner’s Dilemma*, consider the case where after
# each round, Alice and Bob have probability $X$ meeting again. Suppose
# both players choose the perpetual punishment strategy (where each will
# choose ${refuse}$ unless the other player has ever played
# ${testify}$). Assume neither player has played ${testify}$ thus far.
# What is the expected future total payoff for choosing to ${testify}$
# versus ${refuse}$ when $X = .2$? How about when $X = .05$? For what
# value of $X$ is the expected future total payoff the same whether one
# chooses to ${testify}$ or ${refuse}$ in the current round?
#
# **17.22** The following payoff matrix, from @Blinder:1983 by way of @Bernstein:1996, shows a game between
# politicians and the Federal Reserve.
#
# | | Fed: contract | Fed: do nothing | Fed: expand |
# | --- | --- | --- | --- |
# | **Pol: contract** | $F=7, P=1$ | $F=9,P=4$ | $F=6,P=6$ |
# | **Pol: do nothing** | $F=8, P=2$ | $F=5,P=5$ | $F=4,P=9$ |
# | **Pol: expand** | $F=3, P=3$ | $F=2,P=7$ | $F=1,P=8$ |
#
# Politicians can expand or contract fiscal policy, while the Fed can
# expand or contract monetary policy. (And of course either side can
# choose to do nothing.) Each side also has preferences for who should do
# what—neither side wants to look like the bad guys. The payoffs shown are
# simply the rank orderings: 9 for first choice through 1 for last choice.
# Find the Nash equilibrium of the game in pure strategies. Is this a
# Pareto-optimal solution? You might wish to analyze the policies of
# recent administrations in this light.
#
# **17.23** A Dutch auction is similar in an English auction, but rather than
# starting the bidding at a low price and increasing, in a Dutch auction
# the seller starts at a high price and gradually lowers the price until
# some buyer is willing to accept that price. (If multiple bidders accept
# the price, one is arbitrarily chosen as the winner.) More formally, the
# seller begins with a price $p$ and gradually lowers $p$ by increments of
# $d$ until at least one buyer accepts the price. Assuming all bidders act
# rationally, is it true that for arbitrarily small $d$, a Dutch auction
# will always result in the bidder with the highest value for the item
# obtaining the item? If so, show mathematically why. If not, explain how
# it may be possible for the bidder with highest value for the item not to
# obtain it.
#
# **17.24** Imagine an auction mechanism that is just like an ascending-bid auction,
# except that at the end, the winning bidder, the one who bid $b_{max}$,
# pays only $b_{max}/2$ rather than $b_{max}$. Assuming all agents are
# rational, what is the expected revenue to the auctioneer for this
# mechanism, compared with a standard ascending-bid auction?
#
# **17.25** Teams in the National Hockey League historically received 2 points for
# winning a game and 0 for losing. If the game is tied, an overtime period
# is played; if nobody wins in overtime, the game is a tie and each team
# gets 1 point. But league officials felt that teams were playing too
# conservatively in overtime (to avoid a loss), and it would be more
# exciting if overtime produced a winner. So in 1999 the officials
# experimented in mechanism design: the rules were changed, giving a team
# that loses in overtime 1 point, not 0. It is still 2 points for a win
# and 1 for a tie.
#
# 1. Was hockey a zero-sum game before the rule change? After?
#
# 2. Suppose that at a certain time $t$ in a game, the home team has
# probability $p$ of winning in regulation time, probability $0.78-p$
# of losing, and probability 0.22 of going into overtime, where they
# have probability $q$ of winning, $.9-q$ of losing, and .1 of tying.
# Give equations for the expected value for the home and
# visiting teams.
#
# 3. Imagine that it were legal and ethical for the two teams to enter
# into a pact where they agree that they will skate to a tie in
# regulation time, and then both try in earnest to win in overtime.
# Under what conditions, in terms of $p$ and $q$, would it be rational
# for both teams to agree to this pact?
#
# 4. @Longley+Sankaran:2005 report that since the rule change, the percentage of games with a
# winner in overtime went up 18.2%, as desired, but the percentage of
# overtime games also went up 3.6%. What does that suggest about
# possible collusion or conservative play after the rule change?
#
| Jupyter notebook/complex-decisions-exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Estimating a continuous variable from a near-infrared spectra
#
# In this example data set we aim to predict a continous variable. This is the case for instance in the food industry, where one wants to predict nutrient content.
#
#
# ## Data exploration
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
df=pd.read_csv("./data/nir.csv", header=None, index_col=0, sep=";")
df.head(10)
df.columns = ['target']+[i for i in range(1,681)]
from ipywidgets import interact
ymin=df.iloc[:,1:].min().min()
ymax=df.iloc[:,1:].max().max()
def choose_spectra(val):
tol = 0.5
ix=np.where((df.iloc[:,0]>=val-tol) & (df.iloc[:,0]<=val+tol) )[0][0]
plt.figure(figsize=(5,5))
plt.plot(df.iloc[ix,1:])
plt.ylim(ymin,ymax)
plt.show()
minval=df.iloc[:,0].min()
maxval=df.iloc[:,0].max()
interact(choose_spectra, val=(minval,maxval));
# ## Baseline model
#
# We will use **PLS** as a baseline model (http://www.eigenvector.com/Docs/Wise_pls_properties.pdf)
X_train=df.iloc[:,1:].values
y_train=df.iloc[:,0].values
from sklearn.cross_decomposition import PLSRegression
from sklearn.model_selection import GridSearchCV
pls = PLSRegression() # intuition: some decomposition that knows about target
gs=GridSearchCV(pls, {'n_components':[5,10,15,20,25]})
gs.fit(X_train,y_train)
gs.best_estimator_
gs.best_score_
import seaborn as sns
X_train_r = gs.best_estimator_.transform(X_train)
plt.figure(figsize=(10,7))
sns.scatterplot(X_train_r[:,0], X_train_r[:,1], hue=y_train);
from sklearn.decomposition import PCA
pca = PCA()
X_pca= pca.fit_transform(X_train)
plt.figure(figsize=(10,7))
sns.scatterplot(X_pca[:,0], X_pca[:,1], hue=y_train);
y_train_clf = np.where(y_train>50, 1, 0)
y_train_clf
X_train_clf, _ = pls.fit_transform(X_train, y_train_clf)
plt.figure(figsize=(10,7))
sns.scatterplot(X_pca[:,0], X_pca[:,1], hue=y_train_clf);
plt.figure(figsize=(10,7))
sns.scatterplot(X_train_clf[:,0], X_train_clf[:,1], hue=y_train_clf);
X_train_clf.shape
| 03 Regression of NIR spectra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ## 1. Import Libraries
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from xgboost import XGBClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
# hide warnings
import warnings
warnings.filterwarnings('ignore')
# -
# ## 2. Load Dataset
# +
#from sklearn.datasets import load_iris
# create bunch object containing iris dataset and its attributes.
iris = load_iris()
# -
# if we want to know what sort of detail is provided with this dataset
iris.keys()
# the info will tell us more
print (iris.DESCR)
# convert data from a Scikit-learn bunch object to a pandas DataFrame
df_iris = pd.DataFrame(iris['data'], columns=iris['feature_names'])
df_iris['target'] = iris['target']
# ## 3. Summarize the Dataset
# peek at the data
df_iris.head(10)
# dimensions of the dataset
df_iris.shape
# data types
df_iris.dtypes
# statistical summary
pd.set_option('precision', 3)
df_iris.describe()
# all of the numerical values have the same scale (centimeters) and similar ranges between 0 and 8 centimeters
# class distribution
df_iris.groupby('target').size()
#df_iris.target.value_counts()
# each class has the same number of instances (50 or 33% of the dataset).
# pairwise Pearson correlations
df_iris.corr()
# skew for each attribute
df_iris.drop(columns='target').skew() #positive (right) or negative (left) skew
# ## 4. Data Visualization
# ### a. Univariate Plots
# +
#import matplotlib.pyplot as plt
#import seaborn as sns
# box and whisker plots
df_iris.drop(columns='target').plot(kind='box', subplots=True, layout=(2,2), sharex=False, sharey=False, figsize=(10,10))
plt.show()
#sns.boxplot(data=df_iris.drop(columns='target'))
# -
# histograms
df_iris.drop(columns='target').hist(figsize=(10,10))
plt.show()
# density
df_iris.drop(columns='target').plot(kind='density', subplots=True, layout=(2,2), sharex=False, sharey=False, figsize=(10,10))
plt.show()
# It looks like perhaps two of the input variables have a Gaussian distribution.
# ### b. Multivariate Plots
# scatter plot
sns.pairplot(df_iris.drop(columns='target'))
# Note the diagonal grouping of some pairs of attributes. This suggests a high correlation and a predictable relationship.
# different labels in different color
sns.pairplot(df_iris, hue='target')
# heatmap - correlation
sns.heatmap(df_iris.corr(),vmin=-1, vmax=1, annot=True)
# ## 5. Split the data
# +
#from sklearn.model_selection import train_test_split
X = df_iris.iloc[:,:-1].values
y = df_iris.iloc[:,-1].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42,stratify=y)
# -
# ## 6. Evaluate Algorithms
# +
#from sklearn.linear_model import LogisticRegression
#from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
#from sklearn.tree import DecisionTreeClassifier
#from sklearn.neighbors import KNeighborsClassifier
#from sklearn.naive_bayes import GaussianNB
#from sklearn.svm import SVC
#from sklearn.model_selection import KFold
#from sklearn.model_selection import cross_val_score
# Spot-Check Algorithms
models = []
models.append(('LOG', LogisticRegression()))
models.append(('LDA', LinearDiscriminantAnalysis()))
models.append(('KNN', KNeighborsClassifier()))
models.append(('CART', DecisionTreeClassifier()))
models.append(('NB', GaussianNB()))
models.append(('SVM', SVC()))
# evaluate each model in turn
results = []
names = []
for name, model in models:
kfold = KFold(n_splits=10, random_state=42)
cv_results = cross_val_score(model, X_train, y_train, cv=kfold, scoring ='accuracy')
results.append(cv_results)
names.append(name)
msg = "%s: %f (+/-%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
# -
# KNN has the largest estimated accuracy score and may be worth further study.
# Compare Algorithms
fig = plt.figure()
fig.suptitle('Algorithm Comparison')
ax = fig.add_subplot(111)
plt.boxplot(results)
ax.set_xticklabels(names)
plt.show()
# We can see that the box and whisker plots are squashed at the top of the range, with many samples achieving approx. 100% accuracy.
# ## 7. Improve Results
# ### Tuning
# Result for KNN very good so we don't tune models.
# ### Ensemble Methods
# +
#from xgboost import XGBClassifier
#from sklearn.ensemble import AdaBoostClassifier
#from sklearn.ensemble import RandomForestClassifier
#from sklearn.ensemble import ExtraTreesClassifier
# Spot-Check Ensemble Algorithms
ensembles = []
ensembles.append(('XGB', XGBClassifier()))
ensembles.append(('ADA', AdaBoostClassifier()))
ensembles.append(('RF', RandomForestClassifier()))
ensembles.append(('ET', ExtraTreesClassifier()))
# evaluate each model in turn
results = []
names = []
for name, model in ensembles:
kfold = KFold(n_splits=10, random_state=42)
cv_results = cross_val_score(model, X_train, y_train, cv=kfold, scoring ='accuracy')
results.append(cv_results)
names.append(name)
msg = "%s: %f (%f)" % (name, cv_results.mean(), cv_results.std())
print(msg)
# -
# Compare Ensemble Algorithms
fig = plt.figure()
fig.suptitle('Algorithm Comparison')
ax = fig.add_subplot(111)
plt.boxplot(results)
ax.set_xticklabels(names)
plt.show()
# Results are not good as for KNN.
# ## 8. Make Predictions
# +
#from sklearn.metrics import classification_report
#from sklearn.metrics import confusion_matrix
#from sklearn.metrics import accuracy_score
knn=KNeighborsClassifier()
knn.fit(X_train, y_train)
predictions = knn.predict(X_test)
print(accuracy_score(y_test, predictions))
print(confusion_matrix(y_test, predictions))
print(classification_report(y_test, predictions))
# -
# We can see that for KNN the accuracy is 100%. The confusion matrix provides an indication of the none errors made. Finally the classification report provides a breakdown of each class by precision, recall, f1-score and support showing excellent results.
| BestClassificationAlgorithm_IrisDataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tools for NLP
#
# There are lots of feature transformations that need to be done on text data to get it to a point that machine learning algorithms can understand. Luckily, Spark has placed the most important ones in convienent Feature Transformer calls.
#
# Let's go over them before jumping into the project.
# +
import findspark
findspark.init('/home/pushya/spark-2.1.0-bin-hadoop2.7')
import pyspark
from pyspark.sql import SparkSession
# -
spark = SparkSession.builder.appName('nlp').getOrCreate()
# ## Tokenizer
# <p><a href="http://en.wikipedia.org/wiki/Lexical_analysis#Tokenization">Tokenization</a> is the process of taking text (such as a sentence) and breaking it into individual terms (usually words). A simple <a href="api/scala/index.html#org.apache.spark.ml.feature.Tokenizer">Tokenizer</a> class provides this functionality. The example below shows how to split sentences into sequences of words.</p>
#
# <p><a href="api/scala/index.html#org.apache.spark.ml.feature.RegexTokenizer">RegexTokenizer</a> allows more
# advanced tokenization based on regular expression (regex) matching.
# By default, the parameter “pattern” (regex, default: <code>"\\s+"</code>) is used as delimiters to split the input text.
# Alternatively, users can set parameter “gaps” to false indicating the regex “pattern” denotes
# “tokens” rather than splitting gaps, and find all matching occurrences as the tokenization result.</p>
from pyspark.ml.feature import Tokenizer, RegexTokenizer
from pyspark.sql.functions import col, udf
from pyspark.sql.types import IntegerType
sentenceDataFrame = spark.createDataFrame([
(0, "Hi I heard about Spark"),
(1, "I wish Java could use case classes"),
(2, "Logistic,regression,models,are,neat")
], ["id", "sentence"])
sentenceDataFrame.show(truncate=False)
# +
tokenizer = Tokenizer(inputCol="sentence", outputCol="words")
regexTokenizer = RegexTokenizer(inputCol="sentence", outputCol="words", pattern="\\W")
# alternatively, pattern="\\w+", gaps(False)
countTokens = udf(lambda words: len(words), IntegerType())
tokenized = tokenizer.transform(sentenceDataFrame)
tokenized.select("sentence", "words")\
.withColumn("tokens", countTokens(col("words"))).show(truncate=False)
regexTokenized = regexTokenizer.transform(sentenceDataFrame)
regexTokenized.select("sentence", "words") \
.withColumn("tokens", countTokens(col("words"))).show(truncate=False)
# -
#
# ## Stop Words Removal
#
# <p><a href="https://en.wikipedia.org/wiki/Stop_words">Stop words</a> are words which
# should be excluded from the input, typically because the words appear
# frequently and don’t carry as much meaning.</p>
#
# <p><code>StopWordsRemover</code> takes as input a sequence of strings (e.g. the output
# of a <a href="ml-features.html#tokenizer">Tokenizer</a>) and drops all the stop
# words from the input sequences. The list of stopwords is specified by
# the <code>stopWords</code> parameter. Default stop words for some languages are accessible
# by calling <code>StopWordsRemover.loadDefaultStopWords(language)</code>, for which available
# options are “danish”, “dutch”, “english”, “finnish”, “french”, “german”, “hungarian”,
# “italian”, “norwegian”, “portuguese”, “russian”, “spanish”, “swedish” and “turkish”.
# A boolean parameter <code>caseSensitive</code> indicates if the matches should be case sensitive
# (false by default).</p>
# +
from pyspark.ml.feature import StopWordsRemover
sentenceData = spark.createDataFrame([
(0, ["I", "saw", "the", "red", "balloon"]),
(1, ["Mary", "had", "a", "little", "lamb"])
], ["id", "raw"])
remover = StopWordsRemover(inputCol="raw", outputCol="filtered")
remover.transform(sentenceData).show(truncate=False)
# -
# ## n-grams
#
# An n-gram is a sequence of nn tokens (typically words) for some integer nn. The NGram class can be used to transform input features into nn-grams.
#
# <p><code>NGram</code> takes as input a sequence of strings (e.g. the output of a <a href="ml-features.html#tokenizer">Tokenizer</a>). The parameter <code>n</code> is used to determine the number of terms in each $n$-gram. The output will consist of a sequence of $n$-grams where each $n$-gram is represented by a space-delimited string of $n$ consecutive words. If the input sequence contains fewer than <code>n</code> strings, no output is produced.</p>
#
# +
from pyspark.ml.feature import NGram
wordDataFrame = spark.createDataFrame([
(0, ["Hi", "I", "heard", "about", "Spark"]),
(1, ["I", "wish", "Java", "could", "use", "case", "classes"]),
(2, ["Logistic", "regression", "models", "are", "neat"])
], ["id", "words"])
ngram = NGram(n=2, inputCol="words", outputCol="ngrams")
ngramDataFrame = ngram.transform(wordDataFrame)
ngramDataFrame.select("ngrams").show(truncate=False)
# -
# _______
# # Feature Extractors
# _______
# <h2 id="tf-idf">TF-IDF</h2>
#
# <p><a href="http://en.wikipedia.org/wiki/Tf%E2%80%93idf">Term frequency-inverse document frequency (TF-IDF)</a>
# is a feature vectorization method widely used in text mining to reflect the importance of a term
# to a document in the corpus. Denote a term by <code>$t$</code>, a document by d , and the corpus by D.
# Term frequency <code>$TF(t, d)$</code> is the number of times that term <code>$t$</code> appears in document <code>$d$</code>, while
# document frequency <code>$DF(t, D)$</code> is the number of documents that contains term <code>$t$</code>. If we only use
# term frequency to measure the importance, it is very easy to over-emphasize terms that appear very
# often but carry little information about the document, e.g. “a”, “the”, and “of”. If a term appears
# very often across the corpus, it means it doesn’t carry special information about a particular document.
# Inverse document frequency is a numerical measure of how much information a term provides:
#
# $$ IDF(t, D) = \log \frac{|D| + 1}{DF(t, D) + 1} $$
#
# where |D| is the total number of documents in the corpus. Since logarithm is used, if a term
# appears in all documents, its IDF value becomes 0. Note that a smoothing term is applied to avoid
# dividing by zero for terms outside the corpus. The TF-IDF measure is simply the product of TF and IDF:
# $$ TFIDF(t, d, D) = TF(t, d) \cdot IDF(t, D). $$
#
# +
from pyspark.ml.feature import HashingTF, IDF, Tokenizer
sentenceData = spark.createDataFrame([
(0.0, "Hi I heard about Spark"),
(0.0, "I wish Java could use case classes"),
(1.0, "Logistic regression models are neat")
], ["label", "sentence"])
sentenceData.show()
# -
tokenizer = Tokenizer(inputCol="sentence", outputCol="words")
wordsData = tokenizer.transform(sentenceData)
wordsData.show(truncate=False)
# +
hashingTF = HashingTF(inputCol="words", outputCol="rawFeatures", numFeatures=20)
featurizedData = hashingTF.transform(wordsData)
# alternatively, CountVectorizer can also be used to get term frequency vectors
idf = IDF(inputCol="rawFeatures", outputCol="features")
idfModel = idf.fit(featurizedData)
rescaledData = idfModel.transform(featurizedData)
rescaledData.select("label", "features").show(truncate =False)
# -
# ## CountVectorizer
# CountVectorizer and CountVectorizerModel aim to help convert a collection of text documents to vectors of token counts. When an a-priori dictionary is not available, CountVectorizer can be used as an Estimator to extract the vocabulary, and generates a CountVectorizerModel. The model produces sparse representations for the documents over the vocabulary, which can then be passed to other algorithms like LDA.
#
# During the fitting process, CountVectorizer will select the top vocabSize words ordered by term frequency across the corpus. An optional parameter minDF also affects the fitting process by specifying the minimum number (or fraction if < 1.0) of documents a term must appear in to be included in the vocabulary. Another optional binary toggle parameter controls the output vector. If set to true all nonzero counts are set to 1. This is especially useful for discrete probabilistic models that model binary, rather than integer, counts.
# +
from pyspark.ml.feature import CountVectorizer
# Input data: Each row is a bag of words with a ID.
df = spark.createDataFrame([
(0, "a b c".split(" ")),
(1, "a b b c a".split(" "))
], ["id", "words"])
# fit a CountVectorizerModel from the corpus.
cv = CountVectorizer(inputCol="words", outputCol="features", vocabSize=3, minDF=2.0)
model = cv.fit(df)
result = model.transform(df)
result.show(truncate=False)
# -
| Tools_for_NLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#STS-ECG GitHub Issue 39 "Create MGH AVR features & labels CSV #39"
import pandas as pd
#Isolate subset of mgh-all-features-labels.csv using MRNs in mgh-avr.csv.
mghallfeatureslabels = pd.read_csv("mgh-all-features-labels.csv")
mghallfeatureslabels['medrecn_surgdt'] = mghallfeatureslabels.medrecn.astype(str) + "_" + mghallfeatureslabels.surgdt
mghecgreadpcsupranodalrhythms = pd.read_csv("mgh-ecg-read-pc-supranodal-rhythms.csv")
mghecgreadpcsupranodalrhythms['ecg_patientid_clean_ecg_datetime'] = mghecgreadpcsupranodalrhythms.partners_ecg_patientid_clean.astype(str) + "_" + mghecgreadpcsupranodalrhythms.partners_ecg_datetime.astype(str)
#To find AFib labels in the 3-6 mo post-op range, cross-reference a) MRNs of patients with an ECG in 3-6 mo
#post-op range (dropbox:/sts-afib/results/sts-crossref-3-to-6-mo-postop/list_1+_in_all_windows.csv) against
#MRNs of AFib labels: (dropbox:/sts-data/mgh-ecg-read-pc-supranodal-rhythms.csv). Description of label CSV here.
stscrossref3to6mopostop = pd.read_csv("list_1+_in_all_windows.csv")
# -
stscrossref3to6mopostop
# +
stscrossref3to6mopostop['surgdt_str'] = stscrossref3to6mopostop.surgdt.astype(str)
stscrossref3to6mopostop['surgdt_str'] = stscrossref3to6mopostop['surgdt_str'].str[:10]
stscrossref3to6mopostop['medrecn_surgdt'] = stscrossref3to6mopostop.medrecn.astype(str) + "_" + stscrossref3to6mopostop.surgdt_str
stscrossref3to6mopostop['ecg_patientid_clean_ecg_datetime'] = stscrossref3to6mopostop.partners_ecg_patientid_clean.astype(str) + "_" + stscrossref3to6mopostop.partners_ecg_datetime.astype(str)
stscrossref3to6mopostop['ecg_patientid_clean_ecg_datetime'] = stscrossref3to6mopostop['ecg_patientid_clean_ecg_datetime'].str.replace(' ','T')
stscrossref3to6mopostop = stscrossref3to6mopostop.merge(mghecgreadpcsupranodalrhythms, on='ecg_patientid_clean_ecg_datetime', how='left')
sts_afib_272 = stscrossref3to6mopostop.groupby(['medrecn_surgdt']).sum()
sts_afib_272.loc[sts_afib_272['partners_ecg_read_pc_afib'] == 0, 'long-term-afib'] = 0
sts_afib_272.loc[sts_afib_272['partners_ecg_read_pc_afib'] > 0, 'long-term-afib'] = 1
#For each patient (MRN) with an ECG in the 3-6 mo post-op window, you should be able to find the AFib label
#for that (MRN, ECG_datetime) tuple.
sts_afib_272 = sts_afib_272.loc[:, 'long-term-afib']
sts_afib_272 = sts_afib_272.reset_index()
sts_afib_272.to_csv('sts_afib_272.csv', index = None, header=True)
# -
mgh_avr_features_labels = mghavr.merge(sts_afib_272, on='medrecn_surgdt', how='left')
mgh_avr_features_labels = mgh_avr_features_labels.dropna(subset=['long-term-afib'])
mgh_avr_features_labels = mgh_avr_features_labels.drop('medrecn_surgdt', axis=1)
mgh_avr_features_labels['long-term-afib'] = mgh_avr_features_labels['long-term-afib'].astype(int)
mgh_avr_features_labels = mgh_avr_features_labels[['medrecn', 'surgdt', 'mtopd', 'cnstrokp', 'crenfail', 'cpvntlng',
'deepsterninf', 'reop', 'anymorbidity', 'los', 'slos', 'llos', 'long-term-afib', 'opcab',
'opvalve', 'hdef', 'perfustm', 'xclamptm', 'creatlst', 'wbc',
'platelets', 'hct', 'heightcm', 'weightkg', 'gender', 'age', 'diabetes',
'dialysis', 'hypertn', 'cva', 'infendty', 'chrlungd', 'immsupp', 'pvd',
'cvd', 'prcvint', 'prcab', 'prvalve', 'chf', 'carshock', 'resusc',
'classnyh', 'medinotr', 'medster', 'numdisv', 'vdstena', 'vdstenm',
'vdinsufa', 'vdinsufm', 'vdinsuft', 'status', 'incidenc', 'pocpci',
'pocpciin', 'medgp', 'cvdtia', 'cvdpcarsurg', 'medadp5days',
'ethnicity', 'racecaucasian', 'raceblack', 'raceasian', 'racenativeam',
'raceothernativepacific']]
mgh_avr_features_labels.to_csv('mgh-avr-features-labels.csv', index = False, header=True)
| notebooks/create-mgh-all-features-labels-csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.model_selection import cross_val_score
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv), data manipulation as in SQL
import matplotlib.pyplot as plt # this is used for the plot the graph
import seaborn as sns # used for plot interactive graph. I like it most for plot
from sklearn.linear_model import LogisticRegression # to apply the Logistic regression
from sklearn.model_selection import train_test_split # to split the data into two parts
from sklearn import metrics # for the check the error and accuracy of the model
from sklearn.tree import DecisionTreeClassifier
# Any results you write to the current directory are saved as output.
# dont worry about the error if its not working then insteda of model_selection we can use cross_validation
data = pd.read_csv("data.csv",header=0)
# here header 0 means the 0 th row is our coloumn name
# have a look at the data
print(data.head(6))# as u can see our data have imported and having 33 columns
# head is used for to see top 5 by default I used 2 so it will print 6 rows
# now lets look at the type of data we have. We can use
data.info()
# now we can drop this column Unnamed: 32
data.drop("Unnamed: 32",axis=1,inplace=True) # in this process this will change in our data itself
# if you want to save your old data then you can use below code
# data1=data.drop("Unnamed:32",axis=1)
# here axis 1 means we are droping the column
# here you can check the column has been droped
data.columns # this gives the column name which are persent in our data no Unnamed: 32 is not now there
# like this we also don't want the Id column for our analysis
data.drop("id",axis=1,inplace=True)
# As I said above the data can be divided into three parts.lets divide the features according to their category
# now as you know our diagnosis column is a object type so we can map it to integer value
data['diagnosis']=data['diagnosis'].map({'M':1,'B':0})
data.describe() # this will describe the all statistical function of our data
# lets get the frequency of cancer stages
sns.countplot(data['diagnosis'],label="Count")
plt.show()
corr = data.corr() # .corr is used for find corelation
plt.figure(figsize=(14,14))
sns.heatmap(corr, cbar = True, square = True,
cmap= 'coolwarm')
plt.show()
#Based on corrplot let's select some features for the model ( decision is made in order to remove collinearity)
prediction_var = ['texture_mean','perimeter_mean','smoothness_mean','compactness_mean','symmetry_mean']
# now these are the variables which will use for prediction
#now split our data into train and test
train, test = train_test_split(data, test_size = 0.3)# in this our main data is splitted into train and test
# we can check their dimension
print(train.shape)
print(test.shape)
train_X = train[prediction_var]# taking the training data input
train_y=train.diagnosis# This is output of our training data
# same we have to do for test
test_X= test[prediction_var] # taking test data inputs
test_y =test.diagnosis #output value of test dat
logistic = LogisticRegression()
logistic.fit(train_X,train_y)
temp=logistic.predict(test_X)
print(metrics.accuracy_score(temp,test_y)) # to check the accuracy
clf = DecisionTreeClassifier(random_state=0)
cross_val_score(clf, train_X, train_y, cv=10)
clf.fit(train_X,train_y, sample_weight=None, check_input=True, X_idx_sorted=None)
clf.get_params(deep=True)
clf.predict(test_X, check_input=True)
clf.predict_log_proba(test_X)
clf.predict(test_X,check_input=True)
print(clf.score(test_X,test_y, sample_weight=None))
| Cls4-Supervised Learning - 1/Code 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Dictionaries
# ## The Python Dictionary
# Python supports a container type called a dictionary.
# This is also known as an "associative array", "map" or "hash" in other languages.
# In a list, we use a number to look up an element:
names = "<NAME>".split(" ")
names[1]
# In a dictionary, we look up an element using **another object of our choice**:
me = {"name": "James", "age": 39, "Jobs": ["Programmer", "Teacher"]}
me
me["Jobs"]
me["age"]
type(me)
# ### Keys and Values
# The things we can use to look up with are called **keys**:
me.keys()
# The things we can look up are called **values**:
me.values()
# When we test for containment on a `dict` we test on the **keys**:
"Jobs" in me
"James" in me
"James" in me.values()
# ### Immutable Keys Only
# The way in which dictionaries work is one of the coolest things in computer science:
# the "hash table". The details of this are beyond the scope of this course, but we will consider some aspects in the section on performance programming.
#
# One consequence of this implementation is that you can only use **immutable** things as keys.
good_match = {
("Lamb", "Mint"): True,
("Bacon", "Chocolate"): False
}
# but:
# + tags=["raises-exception"]
illegal = {
["Lamb", "Mint"]: True,
["Bacon", "Chocolate"]: False
}
# -
# Remember -- square brackets denote lists, round brackets denote `tuple`s.
# ### No guarantee of order
#
# Another consequence of the way dictionaries work is that there's no guaranteed order among the
# elements:
#
#
#
my_dict = {"0": 0, "1": 1, "2": 2, "3": 3, "4": 4}
print(my_dict)
print(my_dict.values())
# ## Sets
# A set is a `list` which cannot contain the same element twice.
# We make one by calling `set()` on any sequence, e.g. a list or string.
name = "<NAME>"
unique_letters = set(name)
unique_letters
# Or by defining a literal like a dictionary, but without the colons:
primes_below_ten = {2, 3, 5, 7}
type(unique_letters)
type(primes_below_ten)
unique_letters
# This will be easier to read if we turn the set of letters back into a string, with `join`:
"".join(unique_letters)
# A set has no particular order, but is really useful for checking or storing **unique** values.
# Set operations work as in mathematics:
x = set("Hello")
y = set("Goodbye")
x & y # Intersection
x | y # Union
y - x # y intersection with complement of x: letters in Goodbye but not in Hello
# Your programs will be faster and more readable if you use the appropriate container type for your data's meaning.
# Always use a set for lists which can't in principle contain the same data twice, always use a dictionary for anything
# which feels like a mapping from keys to values.
| module01_introduction_to_python/01_06_dictionaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Dictionaries in Python</h1>
# <p><strong>Welcome!</strong> This notebook will teach you about the dictionaries in the Python Programming Language. By the end of this lab, you'll know the basics dictionary operations in Python, including what it is, and the operations on it.</p>
# <h2 id="Dic">Dictionaries</h2>
# <h3 id="content">What are Dictionaries?</h3>
# A dictionary consists of keys and values. It is helpful to compare a dictionary to a list. Instead of the numerical indexes such as a list, dictionaries have keys. These keys are the keys that are used to access values within a dictionary.
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/DictsList.png" width="650" />
# An example of a Dictionary <code>Dict</code>:
# +
# Create the dictionary
Dict = {"key1": 1, "key2": "2", "key3": [3, 3, 3], "key4": (4, 4, 4), ('key5'): 5, (0, 1): 6}
Dict
# -
# The keys can be strings:
# +
# Access to the value by the key
Dict["key1"]
# -
# Keys can also be any immutable object such as a tuple:
# +
# Access to the value by the key
Dict[(0, 1)]
# -
# Each key is separated from its value by a colon "<code>:</code>". Commas separate the items, and the whole dictionary is enclosed in curly braces. An empty dictionary without any items is written with just two curly braces, like this "<code>{}</code>".
# +
# Create a sample dictionary
release_year_dict = {"Thriller": "1982", "Back in Black": "1980", \
"The Dark Side of the Moon": "1973", "The Bodyguard": "1992", \
"Bat Out of Hell": "1977", "Their Greatest Hits (1971-1975)": "1976", \
"Saturday Night Fever": "1977", "Rumours": "1977"}
release_year_dict
# -
# In summary, like a list, a dictionary holds a sequence of elements. Each element is represented by a key and its corresponding value. Dictionaries are created with two curly braces containing keys and values separated by a colon. For every key, there can only be one single value, however, multiple keys can hold the same value. Keys can only be strings, numbers, or tuples, but values can be any data type.
# It is helpful to visualize the dictionary as a table, as in the following image. The first column represents the keys, the second column represents the values.
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/DictsStructure.png" width="650" />
# <h3 id="key">Keys</h3>
# You can retrieve the values based on the names:
# +
# Get value by keys
release_year_dict['Thriller']
# -
# This corresponds to:
#
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/DictsKeyOne.png" width="500" />
# Similarly for <b>The Bodyguard</b>
# +
# Get value by key
release_year_dict['The Bodyguard']
# -
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/DictsKeyTwo.png" width="500" />
# Now let you retrieve the keys of the dictionary using the method <code>release_year_dict()</code>:
# +
# Get all the keys in dictionary
release_year_dict.keys()
# -
# You can retrieve the values using the method <code>values()</code>:
# +
# Get all the values in dictionary
release_year_dict.values()
# -
# We can add an entry:
# +
# Append value with key into dictionary
release_year_dict['Graduation'] = '2007'
release_year_dict
# -
# We can delete an entry:
# +
# Delete entries by key
del(release_year_dict['Thriller'])
del(release_year_dict['Graduation'])
release_year_dict
# -
# We can verify if an element is in the dictionary:
# +
# Verify the key is in the dictionary
'The Bodyguard' in release_year_dict
# -
# <hr>
# <h2 id="quiz">Quiz on Dictionaries</h2>
# <b>You will need this dictionary for the next two questions:</b>
# +
# Question sample dictionary
soundtrack_dic = {"The Bodyguard":"1992", "Saturday Night Fever":"1977"}
soundtrack_dic
# -
# a) In the dictionary <code>soundtrack_dict</code> what are the keys ?
# +
# Write your code below and press Shift+Enter to execute
# -
# Double-click __here__ for the solution.
#
# <!-- Your answer is below:
# soundtrack_dic.keys() # The Keys "The Bodyguard" and "Saturday Night Fever"
# -->
# b) In the dictionary <code>soundtrack_dict</code> what are the values ?
# +
# Write your code below and press Shift+Enter to execute
# -
# Double-click __here__ for the solution.
#
# <!-- Your answer is below:
# soundtrack_dic.values() # The values are "1992" and "1977"
# -->
# <hr>
# <b>You will need this dictionary for the following questions:</b>
# The Albums <b>Back in Black</b>, <b>The Bodyguard</b> and <b>Thriller</b> have the following music recording sales in millions 50, 50 and 65 respectively:
# a) Create a dictionary <code>album_sales_dict</code> where the keys are the album name and the sales in millions are the values.
# +
# Write your code below and press Shift+Enter to execute
# -
# Double-click __here__ for the solution.
#
# <!-- Your answer is below:
# album_sales_dict = {"The Bodyguard":50, "Back in Black":50, "Thriller":65}
# -->
# b) Use the dictionary to find the total sales of <b>Thriller</b>:
# +
# Write your code below and press Shift+Enter to execute
# -
# Double-click __here__ for the solution.
#
# <!-- Your answer is below:
# album_sales_dict["Thriller"]
# -->
# c) Find the names of the albums from the dictionary using the method <code>keys</code>:
# +
# Write your code below and press Shift+Enter to execute
# -
# Double-click __here__ for the solution.
#
# <!-- Your answer is below:
# album_sales_dict.keys()
# -->
# d) Find the names of the recording sales from the dictionary using the method <code>values</code>:
# +
# Write your code below and press Shift+Enter to execute
# -
# Double-click __here__ for the solution.
#
# <!-- Your answer is below:
# album_sales_dict.values()
# -->
| Python-Dictionaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Before We Start!
#
# 1. Github repository for these notebooks: **github.com/ericmjl/Network-Analysis-Made-Simple/**
# 1. Please clone the repository if you'd like to do the hands-on coding activities.
# 1. Some of the coding activities are going to be hard! Be ready to discuss the problem with your fellow Pythonistas.
# 1. If you are using legacy Python, you may wish to pair up.
# + [markdown] slideshow={"slide_type": "subslide"}
# # Quiz!
#
# In the list comprehension:
#
# [s for s in my_fav_things if s[‘name’] == ‘raindrops on roses’]
#
# What are a plausible data structure for `s` and `my_fav_things`?
# + slideshow={"slide_type": "fragment"}
my_fav_things = []
my_fav_things.append({'name': 'raindrops on roses', 'line': 1})
my_fav_things.append({'name': 'whiskers on kittens', 'line': 1})
my_fav_things.append({'name': 'bright copper kettles', 'line': 2})
[s for s in my_fav_things if s['name'] == 'raindrops on roses']
# + [markdown] slideshow={"slide_type": "subslide"}
# # Prerequisites
#
# Use the `check_env.py` script provided in the repository to determine whether you need to install any new dependencies. You may do so while we quickly go through some background information.
# + [markdown] slideshow={"slide_type": "slide"}
# # Network Basics
#
# *All your relational problems are belong to networks.*
#
# Networks, a.k.a. graphs, are an immensely useful modelling tool to model complex relational problems.
#
# Networks are comprised of two main entities:
#
# - Nodes: commonly represented as circles. In the academic literature, nodes are also known as "vertices".
# - Edges: commonly represented as lines between circles
# + [markdown] slideshow={"slide_type": "subslide"}
# Edges denote relationships between the nodes.
#
# > The heart of a graph lies in its edges, not in its nodes.
# > (<NAME>, Harvard School of Public Health)
# + [markdown] slideshow={"slide_type": "fragment"}
# In a network, if two nodes are joined together by an edge, then they are neighbors of one another.
#
# There are generally two types of networks - directed and undirected. In undirected networks, edges do not have a directionality associated with them. In directed networks, they do.
# + [markdown] slideshow={"slide_type": "slide"}
# # Examples of Networks
#
# 1. Facebook's network: Individuals are nodes, edges are drawn between individuals who are FB friends with one another. **undirected network**.
# 2. Air traffic network: Airports are nodes, flights between airports are the edges. **directed network**.
#
# Can you think of any others?
#
# The key questions here are as follows. How do we...:
#
# 1. model a problem as a network?
# 2. extract useful information from a network?
# + [markdown] slideshow={"slide_type": "slide"}
# # Take-Homes
#
# It is my hope that when you leave this tutorial, practically, you will be equipped to:
#
# - Use NetworkX to construct graphs in the Jupyter environment.
# - Visualize network data using node-link diagrams, heat maps, Circos plots and Hive plots.
# - Write basic algorithms to find structures and paths in a graph.
# - Compute network statistics.
# + [markdown] slideshow={"slide_type": "subslide"}
# # Take-Homes (cont'd)
#
# From a broader perspective, I hope you will be able to:
#
# - Think in terms of "interactions" between entities, and not just think about the entities themselves.
# - Think through statistical problems in network analysis.
# + [markdown] slideshow={"slide_type": "skip"}
# # Credits
#
# Much of this work is inspired by Prof. <NAME> (Olin College of Engineering) and Prof. <NAME> (Harvard School of Public Health).
#
# Statistics methods are inspired by Dr. <NAME>, UW.
#
# Hive and Circos Plots' original inventor is <NAME> of the BC Genome Sciences Center.
#
# Circos plots were implemented with help from <NAME> (MIT).
#
# Many thanks to the PyCon Rehearsal class for providing feedback on the material prior to PyCon 2015.
#
# Thank you all who attended actual iterations of this tutorial, at
#
# - PyCon 2015 (Montreal)
# - Data Science for Social Good (Boston)
# - PyData NYC 2015 (New York City)
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# # The Data
# In this tutorial, we will go through a synthetic social network comprising of 30 individuals, to illustrate some of the basic concepts when constructing and analyzing networks. I will use this data set for the entirety of this tutorial.
# -
| 1. Preface.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A brief intro to pydeck
#
# pydeck is made for visualizing data points in 2D or 3D maps. Specifically, it handles
#
# - rendering large (>1M points) data sets, like LIDAR point clouds or GPS pings
# - large-scale updates to data points, like plotting points with motion
# - making beautiful maps
#
# Under the hood, it's powered by the [deck.gl](https://github.com/uber/deck.gl/) JavaScript framework.
#
# pydeck is strongest when used in tandem with [Pandas](https://pandas.pydata.org/) but doesn't have to be.
#
# Please note that **these demo notebooks are best when executed cell-by-cell**, so ideally clone this repo.
import pydeck as pdk
# # There are three steps for most pydeck visualizations
#
# We'll walk through pydeck using a visualization of vehicle accident data in the United Kingdom.
#
# ## 1. Choose your data
#
# Here, we'll use the history of accident data throughout the United Kingdom. This data set presents the location of every latitude and longitude of car accidents in the UK in 2014 ([source](https://data.gov.uk/dataset/053a6529-6c8c-42ac-ae1e-455b2708e535/road-traffic-accidents)).
# +
import pandas as pd
UK_ACCIDENTS_DATA = 'https://raw.githubusercontent.com/uber-common/deck.gl-data/master/examples/3d-heatmap/heatmap-data.csv'
pd.read_csv(UK_ACCIDENTS_DATA).head()
# -
# ## 2. Configure the visualization: Choose your layer(s) and viewport
#
# pydeck's **`Layer`** object takes two positional and many keyword arguments:
#
# - First, a string specifying the layer type, with our example below using `'HexagonLayer'`
# - Next, a data URL–below you'll see the `UK_ACCIDENTS_DATA` that we set above, but we could alternately pass a data frame or list of dictionaries
# - Finally, keywords representing that layer's attributes–in our example, this would include `elevation_scale`, `elevation_range`, `extruded`, `coverage`. `pickable=True` also allows us to add a tooltip that appears on hover.
#
# ```python
# layer = pdk.Layer(
# 'HexagonLayer',
# UK_ACCIDENTS_DATA,
# get_position='[lng,lat]',
# elevation_scale=50,
# pickable=True,
# auto_highlight=True,
# elevation_range=[0, 3000],
# extruded=True,
# coverage=1)
# ```
#
# There is of course an entire catalog of layers which you're welcome to check out within the [deck.gl documentation](https://deck.gl/#/documentation/deckgl-api-reference/layers/overview).
#
# ### Configure your viewport
#
# We also have to specifiy a **`ViewState`** object.
#
# The **`ViewState`** object specifies a camera angle relative to the map data. If you don't want to manually specify it, the function **`pydeck.data_utils.compute_viewport`** can take your data and automatically zoom to it.
#
# pydeck also provides some controls, most of which should be familiar from map applications throughout the web. By default, you can hold out and drag to rotate the map.
# +
layer = pdk.Layer(
'HexagonLayer',
UK_ACCIDENTS_DATA,
get_position='[lng,lat]',
auto_highlight=True,
elevation_scale=50,
pickable=True,
elevation_range=[0, 3000],
extruded=True,
coverage=1)
# Set the viewport location
view_state = pdk.ViewState(
longitude=-1.415,
latitude=52.2323,
zoom=6,
min_zoom=5,
max_zoom=15,
pitch=40.5,
bearing=-27.36)
# Combined all of it and render a viewport
r = pdk.Deck(layers=[layer], initial_view_state=view_state)
r.show()
# -
# ## Render an update to the visualization
#
# Execute the cell below and look at the map in the cell above–you'll notice a seamless rendered update on the map
# +
layer.elevation_range = [0, 10000]
r.update()
# -
# ## Support updates over time
#
# We can combine any Python function with our work here, of course. Execute the cell below to update our map above over time.
import time
r.show()
for i in range(0, 10000, 1000):
layer.elevation_range = [0, i]
r.update()
time.sleep(0.1)
# # pydeck without Jupyter
#
# If you prefer not to use Jupyter or you'd like to export your map to a separate file, you can also write out maps to HTML locally using the `.to_html` function.
#
# (Note that if you're executing this example on mybinder.org, it won't render, since write access in the binder environment is restricted.)
r.to_html('deck-iframe.html')
| bindings/python/pydeck/examples/01 - Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## <small>
# Copyright (c) 2017-21 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# </small>
#
#
#
# # Deep Learning: A Visual Approach
# ## by <NAME>, https://glassner.com
# ### Order: https://nostarch.com/deep-learning-visual-approach
# ### GitHub: https://github.com/blueberrymusic
# ------
#
# ### What's in this notebook
#
# This notebook is provided as a “behind-the-scenes” look at code used to make some of the figures in this chapter. It is cleaned up a bit from the original code that I hacked together, and is only lightly commented. I wrote the code to be easy to interpret and understand, even for those who are new to Python. I tried never to be clever or even more efficient at the cost of being harder to understand. The code is in Python3, using the versions of libraries as of April 2021.
#
# This notebook may contain additional code to create models and images not in the book. That material is included here to demonstrate additional techniques.
#
# Note that I've included the output cells in this saved notebook, but Jupyter doesn't save the variables or data that were used to generate them. To recreate any cell's output, evaluate all the cells from the start up to that cell. A convenient way to experiment is to first choose "Restart & Run All" from the Kernel menu, so that everything's been defined and is up to date. Then you can experiment using the variables, data, functions, and other stuff defined in this notebook.
# ## Chapter 10: Data Preparation, Notebook 1
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import math
# +
# Make a File_Helper for saving and loading files.
save_files = False
import os, sys, inspect
current_dir = os.path.dirname(os.path.abspath(inspect.getfile(inspect.currentframe())))
sys.path.insert(0, os.path.dirname(current_dir)) # path to parent dir
from DLBasics_Utilities import File_Helper
file_helper = File_Helper(save_files)
# +
# The following cells are intended to be executed
# serially, since they build new data as they go
# based on previous data held in global variables.
# +
# Points for the electric guitar dataset. I wrote a little program in Processing that
# let me click, click, click and it saved the XY locations to a text file, which I
# just copied in here.
points_list = np.array([
[ 0.14343707, 0.8876861 ], [ 0.086603515, 0.84032476 ], [ 0.1014885, 0.9093369 ],
[ 0.21244925, 0.77943164 ], [ 0.2503383, 0.91745603 ], [ 0.24763194, 0.8362652 ],
[ 0.21244925, 0.87280107 ], [ 0.13396482, 0.84573746 ], [ 0.16508795, 0.95805144 ],
[ 0.17185386, 0.9066306 ], [ 0.1217862, 0.93098783 ], [ 0.07577808, 0.94316643 ],
[ 0.054127198, 0.8958051 ], [ 0.24763194, 0.744249 ], [ 0.26928282, 0.76860625 ],
[ 0.29499322, 0.80784845 ], [ 0.25845736, 0.79566985 ], [ 0.20838971, 0.8362652 ],
[ 0.27198917, 0.8579161 ], [ 0.30175912, 0.84573746 ], [ 0.3044655, 0.8755074 ],
[ 0.27604872, 0.8849797 ], [ 0.27469555, 0.9201624 ], [ 0.21650879, 0.9188092 ],
[ 0.21921515, 0.95128554 ], [ 0.25169146, 0.94993234 ], [ 0.07848444, 0.864682 ],
[ 0.18132612, 0.83491206 ], [ 0.1894452, 0.8890392 ], [ 0.1894452, 0.9729364 ],
[ 0.15155616, 0.93775374 ], [ 0.09336942, 0.96346414 ], [ 0.12990528, 0.96481735 ],
[ 0.113667116, 0.8849797 ], [ 0.09336942, 0.93098783 ], [ 0.046008117, 0.9255751 ],
[ 0.031123139, 0.87415427 ], [ 0.03924222, 0.8254398 ], [ 0.0608931, 0.8173207 ],
[ 0.086603515, 0.78890395 ], [ 0.119079836, 0.78213805 ], [ 0.15832205, 0.80108255 ],
[ 0.17185386, 0.7699594 ], [ 0.23139377, 0.76184034 ], [ 0.22868742, 0.8037889 ],
[ 0.23139377, 0.85926926 ], [ 0.27198917, 0.8294993 ], [ 0.24221921, 0.8903924 ],
[ 0.19485791, 0.932341 ], [ 0.135318, 0.9079838 ], [ 0.113667116, 0.8579161 ],
[ 0.05277402, 0.84709066 ], [ 0.09607578, 0.8186739 ], [ 0.07713126, 0.8958051 ],
[ 0.16102841, 0.86738837 ], [ 0.19350474, 0.80108255 ], [ 0.13937753, 0.8159675 ],
[ 0.062246278, 0.78078485 ], [ 0.0879567, 0.7523681 ], [ 0.1217862, 0.75372124 ],
[ 0.24086604, 0.6738836 ], [ 0.25169146, 0.6414073 ], [ 0.21380244, 0.7198917 ],
[ 0.22868742, 0.744249 ], [ 0.19891746, 0.74830854 ], [ 0.18673883, 0.7239513 ],
[ 0.16508795, 0.729364 ], [ 0.124492556, 0.7185386 ], [ 0.1488498, 0.744249 ],
[ 0.10554804, 0.7388363 ], [ 0.108254395, 0.7023004 ], [ 0.15155616, 0.7036536 ],
[ 0.19350474, 0.6955345 ], [ 0.22868742, 0.6941813 ], [ 0.2300406, 0.65223277 ],
[ 0.20974289, 0.6752368 ], [ 0.19756427, 0.6481732 ], [ 0.1894452, 0.6738836 ],
[ 0.15696888, 0.6738836 ], [ 0.135318, 0.6752368 ], [ 0.16779432, 0.64952636 ],
[ 0.13802436, 0.65629226 ], [ 0.10554804, 0.6752368 ], [ 0.10554804, 0.64952636 ],
[ 0.08389716, 0.65629226 ], [ 0.07171854, 0.63734776 ], [ 0.0879567, 0.6265223 ],
[ 0.1217862, 0.6332882 ], [ 0.16102841, 0.6332882 ], [ 0.18673883, 0.62516916 ],
[ 0.16914749, 0.61163735 ], [ 0.16102841, 0.58998644 ], [ 0.14073072, 0.60622466 ],
[ 0.11096076, 0.6075778 ], [ 0.13396482, 0.5872801 ], [ 0.10013532, 0.58998644 ],
[ 0.07577808, 0.60487145 ], [ 0.15155616, 0.57239515 ], [ 0.16644114, 0.5615697 ],
[ 0.15832205, 0.53721243 ], [ 0.15696888, 0.511502 ], [ 0.16238159, 0.48173207 ],
[ 0.1556157, 0.47090665 ], [ 0.16102841, 0.44384304 ], [ 0.15290934, 0.4316644 ],
[ 0.15967524, 0.40324762 ], [ 0.15832205, 0.37077132 ], [ 0.15426251, 0.35453317 ],
[ 0.15967524, 0.3301759 ], [ 0.15696888, 0.2976996 ], [ 0.15696888, 0.26792964 ],
[ 0.16238159, 0.23951286 ], [ 0.15832205, 0.21380244 ], [ 0.15832205, 0.1826793 ],
[ 0.16238159, 0.14479026 ], [ 0.15155616, 0.119079836 ], [ 0.1759134, 0.1150203 ],
[ 0.15696888, 0.097428955 ], [ 0.17861976, 0.08930988 ], [ 0.17050068, 0.066305816 ],
[ 0.15426251, 0.07713126 ], [ 0.1556157, 0.05006766 ], [ 0.17726658, 0.05006766 ],
[ 0.17320703, 0.03247632 ], [ 0.15426251, 0.028416779 ], [ 0.17861976, 0.5223275 ],
[ 0.18132612, 0.4993234 ], [ 0.17861976, 0.47225982 ], [ 0.17861976, 0.44384304 ],
[ 0.17861976, 0.4127199 ], [ 0.17726658, 0.39106902 ], [ 0.1759134, 0.3653586 ],
[ 0.17456022, 0.33423546 ], [ 0.1759134, 0.30040595 ], [ 0.17456022, 0.27334234 ],
[ 0.17726658, 0.25304466 ], [ 0.17456022, 0.22192152 ], [ 0.17456022, 0.19485791 ],
[ 0.17185386, 0.1759134 ], [ 0.1759134, 0.1556157 ], [ 0.135318, 0.53315294 ],
[ 0.13802436, 0.5142084 ], [ 0.13802436, 0.4871448 ], [ 0.13802436, 0.46414074 ],
[ 0.13667117, 0.43572396 ], [ 0.13667117, 0.41001353 ], [ 0.13937753, 0.38159674 ],
[ 0.13937753, 0.36129904 ], [ 0.13802436, 0.32611638 ], [ 0.1420839, 0.29364005 ],
[ 0.14073072, 0.2638701 ], [ 0.14073072, 0.2368065 ], [ 0.14343707, 0.21786198 ],
[ 0.14343707, 0.1894452 ], [ 0.14479026, 0.1556157 ], [ 0.14479026, 0.13802436 ],
[ 0.1826793, 0.13396482 ], [ 0.19215156, 0.11772666 ], [ 0.135318, 0.123139374 ],
[ 0.19079837, 0.0879567 ], [ 0.13125846, 0.0947226 ], [ 0.19485791, 0.056833558 ],
[ 0.135318, 0.05818674 ], [ 0.20027064, 0.01894452 ], [ 0.18673883, 0.013531799 ],
[ 0.17050068, 0.00947226 ], [ 0.15155616, 0.013531799 ], [ 0.13396482, 0.014884979 ],
[ 0.13396482, 0.028416779 ], [ 0.21244925, 0.102841675 ], [ 0.1962111, 0.102841675 ],
[ 0.21515562, 0.07307172 ], [ 0.19891746, 0.0744249 ], [ 0.21650879, 0.040595397 ],
[ 0.20162381, 0.040595397 ], [ 0.13261163, 0.04194858 ], [ 0.113667116, 0.043301757 ],
[ 0.13125846, 0.07848444 ], [ 0.113667116, 0.07713126 ], [ 0.1285521, 0.108254395 ],
[ 0.11231394, 0.108254395 ], [ 0.3044655, 0.7767253 ], [ 0.2909337, 0.759134 ],
[ 0.27604872, 0.74154264 ], [ 0.25981057, 0.7239513 ], [ 0.2571042, 0.699594 ],
[ 0.25981057, 0.67658997 ], [ 0.270636, 0.65223277 ], [ 0.2774019, 0.6278755 ],
[ 0.2774019, 0.608931 ], [ 0.26251692, 0.60622466 ], [ 0.24898511, 0.6197564 ],
[ 0.22598106, 0.62381595 ], [ 0.20838971, 0.61705005 ], [ 0.19485791, 0.59810555 ],
[ 0.18809202, 0.57645464 ], [ 0.18673883, 0.5534506 ], [ 0.3112314, 0.7983762 ],
[ 0.32205683, 0.8227334 ], [ 0.32746956, 0.849797 ], [ 0.32476318, 0.8782138 ],
[ 0.31393775, 0.9079838 ], [ 0.30175912, 0.93098783 ], [ 0.28552097, 0.95534503 ],
[ 0.26251692, 0.9729364 ], [ 0.23815967, 0.98240864 ], [ 0.21380244, 0.9864682 ],
[ 0.19215156, 0.9918809 ], [ 0.16373478, 0.9945873 ], [ 0.13802436, 0.9918809 ],
[ 0.11231394, 0.9864682 ], [ 0.086603515, 0.97699594 ], [ 0.0608931, 0.9661705 ],
[ 0.04194858, 0.94993234 ], [ 0.027063599, 0.93640053 ], [ 0.01623816, 0.9147497 ],
[ 0.00947226, 0.8930988 ], [ 0.0067658997, 0.86738837 ], [ 0.0067658997, 0.8376184 ],
[ 0.01759134, 0.8105548 ], [ 0.029769959, 0.7848444 ], [ 0.0473613, 0.7631935 ],
[ 0.062246278, 0.7428958 ], [ 0.07307172, 0.7253045 ], [ 0.0744249, 0.6968877 ],
[ 0.06901218, 0.6738836 ], [ 0.05548038, 0.64546686 ], [ 0.048714478, 0.62516916 ],
[ 0.05818674, 0.59675235 ], [ 0.081190795, 0.5751015 ], [ 0.1014885, 0.56427604 ],
[ 0.13125846, 0.5602165 ],
])
# +
# Rotate the points by the given angle and return a new list
def rotate_points(pts, theta):
p2 = pts.copy()
for i in range(pts.shape[0]):
xrot = (pts[i][0]*math.cos(theta)) + (pts[i][1]*math.sin(theta))
yrot = (-pts[i][0]*math.sin(theta)) + (pts[i][1]*math.cos(theta))
p2[i][0] = xrot
p2[i][1] = yrot
return p2
# +
# set generic controls for a nice plot
def setup_plot(pts):
pmin = 1.2 * np.min(pts[:,0:2])
pmax = 1.2 * np.max(pts[:,0:2])
plt.xlim(pmin, pmax)
plt.ylim(pmin, pmax)
plt.gca().set_aspect('equal', adjustable='box')
# -
# The variable names in the following cells are prefixed by what's been done to their data:
#
# - S=sorted
# - C=colored
# - R=rotated
# - N=normalized
# - V=variance adjusted
# +
# draw the guitar with colored points
Spts = np.array(sorted(points_list, key=lambda x: x[1], reverse=False))
SCpts = np.array([[Spts[i][0], Spts[i][1], i] for i in range(len(Spts))])
theta = math.pi * 2 * -60/180.0
# rotate the points a bit so the guitar is tilted
SCRpts = rotate_points(SCpts, theta)
plt.scatter(SCRpts[:,0], SCRpts[:,1], c=SCRpts[:,2], cmap='jet', edgecolor='black')
setup_plot(SCRpts)
plt.title("Input points")
file_helper.save_figure('eguitar-input')
plt.show()
# -
# N=normalized
xmin = np.min(SCRpts[:,0])
xmax = np.max(SCRpts[:,0])
ymin = np.min(SCRpts[:,1])
ymax = np.max(SCRpts[:,1])
SCRNpts = SCRpts.copy()
for i in range(SCRpts.shape[0]):
SCRNpts[i][0] = np.interp(SCRpts[i][0], [xmin, xmax], [-1,1])
SCRNpts[i][1] = np.interp(SCRpts[i][1], [ymin, ymax], [-1,1])
plt.clf()
plt.scatter(SCRNpts[:,0], SCRNpts[:,1], c=SCRNpts[:,2], cmap='jet', edgecolor='black')
setup_plot(SCRNpts)
plt.title("normalized")
file_helper.save_figure('eguitar-normalized')
plt.show()
# +
# V=variance (and mean) adjusted
# We'll do this explicitly to show the steps
# It would be shorter (and probably faster) to use Numpy
# compute the means and center the data
mean_scrn_x = (1/len(SCRNpts))*np.sum(SCRNpts[:,0])
mean_scrn_y = (1/len(SCRNpts))*np.sum(SCRNpts[:,1])
Mpts = []
for i in range(len(SCRNpts)):
pt = SCRNpts[i]
vx = pt[0]-mean_scrn_x
vy = pt[1]-mean_scrn_y
Mpts.append([vx, vy, pt[2]])
Mpts = np.array(Mpts)
# compute the variances and scale each feature
sigma_x = math.sqrt((1/len(Mpts))*np.sum(Mpts[:,0]**2))
sigma_y = math.sqrt((1/len(Mpts))*np.sum(Mpts[:,1]**2))
SCRNVpts = []
for i in range(len(Mpts)):
pt = Mpts[i]
vx = pt[0]/sigma_x
vy = pt[1]/sigma_y
SCRNVpts.append([vx, vy, pt[2]])
SCRNVpts = np.array(SCRNVpts)
plt.clf()
plt.scatter(SCRNVpts[:,0], SCRNVpts[:,1], c=SCRNVpts[:,2], cmap='jet', edgecolor='black')
setup_plot(SCRNVpts)
plt.title("mean and variance adjusted")
file_helper.save_figure('eguitar-mean-variance')
plt.show()
# -
# project the adjusted points to the y=0 line
plt.clf()
plt.scatter(SCRNVpts[:,0], SCRNVpts[:,1], c=SCRNVpts[:,2], cmap='jet', alpha=0.3, edgecolor='black')
for i in range(SCRNVpts.shape[0]):
if np.random.uniform() < .25:
xyc = SCRNVpts[i]
plt.plot([xyc[0], xyc[0]], [xyc[1], 0], lw=1, color='black')
plt.plot([np.min(SCRNVpts[:,0]), np.max(SCRNVpts[:,0])], [0,0], lw=2, color='black')
setup_plot(SCRNVpts)
plt.title("projecting points to y=0")
file_helper.save_figure('eguitar-project-y0')
plt.show()
# show the projected points
plt.clf()
plt.scatter(SCRNVpts[:,0], SCRNVpts.shape[0]*[0], c=SCRNVpts[:,2], lw=.3, cmap='jet', edgecolor='black')
setup_plot(SCRNVpts)
plt.title("projected to y=0")
file_helper.save_figure('eguitar-projected-to-y0')
plt.show()
# projected onto the line we found by PCA
plt.clf()
ptheta = math.pi * 2 * 29/180.0
dx = math.cos(ptheta)
dy = math.sin(ptheta)
plt.scatter(SCRNVpts[:,0], SCRNVpts[:,1], c=SCRNVpts[:,2], cmap='jet', alpha=0.3, edgecolor='black')
SCRNVPpts = SCRNVpts.copy()
plt.plot([-10*dx, 10*dx], [-10*dy, 10*dy], lw=2, color='black')
for i in range(SCRNVpts.shape[0]):
xyc = SCRNVpts[i]
plen = math.sqrt((xyc[0]*xyc[0])+(xyc[1]*xyc[1]))
normp = [xyc[0]/plen, xyc[1]/plen]
dot = (dx * xyc[0])+(dy * xyc[1])
SCRNVPpts[i][0] = dx*dot
SCRNVPpts[i][1] = dy*dot
for i in range(SCRNVpts.shape[0]):
if np.random.uniform() < .25:
plt.plot([SCRNVpts[i][0], SCRNVPpts[i][0]],
[SCRNVpts[i][1], SCRNVPpts[i][1]], lw=1.5, color='black')
setup_plot(SCRNVPpts)
plt.title("projecting points")
file_helper.save_figure('eguitar-projected-tilted')
plt.show()
# show the line we found by PCA
plt.clf()
ptheta = math.pi * 2 * 29/180.0
dx = math.cos(ptheta)
dy = math.sin(ptheta)
plt.scatter(SCRNVpts[:,0], SCRNVpts[:,1], c=SCRNVpts[:,2], cmap='jet', alpha=0.5, edgecolor='black')
plt.plot([-10*dx, 10*dx], [-10*dy, 10*dy], lw=2, color='black')
setup_plot(SCRNVPpts)
plt.title("line of max variance")
file_helper.save_figure('eguitar-max-variance')
plt.show()
# show the projected points
plt.clf()
plt.scatter(SCRNVPpts[:,0], SCRNVPpts[:,1], c=SCRNVPpts[:,2], lw=.3, cmap='jet', edgecolor='black')
setup_plot(SCRNVPpts)
plt.title("projected points")
file_helper.save_figure('eguitar-rot-line')
plt.show()
plt.clf()
line_pts = rotate_points(SCRNVPpts, ptheta)
plt.scatter(line_pts[:,0], line_pts[:,1], c=line_pts[:,2], lw=.3, cmap='jet', edgecolor='black')
setup_plot(line_pts)
plt.title("rotated projected points")
file_helper.save_figure('eguitar-unrot-line')
plt.show()
# +
# Show our made-up data for #cars on the road as a function of temperature
np.random.seed(42)
num_points = 100
temp_lo = -30
temp_hi = 40
cars_lo = 20
cars_hi = 80
temps = np.random.uniform(low=temp_lo, high=temp_hi, size=num_points)
cars = np.zeros(num_points)
for i in range(num_points):
sint = math.sin(np.interp(temps[i], [temp_lo, temp_hi], [0, math.pi/2]))
cars[i] = np.interp(sint, [0,1], [cars_lo, cars_hi]) + np.random.uniform(-5, 5)
plt.scatter(temps, cars)
plt.xlabel('Midnight temperature in degrees Celsius')
plt.ylabel('Cars on the road')
plt.title("Starting Data")
file_helper.save_figure('raw-traffic-data')
plt.show()
# +
# Normalize the car data
ntemps = np.interp(temps, [np.min(temps), np.max(temps)], [0,1])
ncars = np.interp(cars, [np.min(cars), np.max(cars)], [0,1])
plt.scatter(ntemps, ncars)
plt.xlabel('Normalized temperature')
plt.ylabel('Normalized cars')
plt.title("Normalized Data")
file_helper.save_figure('scaled-traffic-data')
plt.show()
# -
| Notebooks/Chapter10-DataPreparation/Chapter10-DataPreparation-1-Miscellaneous.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize OK
from client.api.notebook import Notebook
ok = Notebook('lab01.ok')
# + [markdown] deletable=false
# # Lab 1: Expressions
#
# Welcome to Data 8: *Foundations of Data Science*! Each week you will complete a lab assignment like this one. You can't learn technical subjects without hands-on practice, so labs are an important part of the course.
#
# Before we get started, there are some administrative details.
#
# The weekly lab session has two components: questions and discussion (not using the computer) about recent material, and a lab assignment (like this one!) that develops skills with computational and inferential concepts. These lab assignments are a required part of the course and will be released on Monday mornings.
#
# Lab sessions are not webcast. The set of questions covered in lab will be posted; for the related discussion, please attend the session. There are two ways to each full credit the weekly lab:
#
# 1. Attend your own assigned lab section and make progress substantial enough for your work to be checked off by course staff. Before you leave, you need to **submit the lab and have a staff member check you off**. Even if you finish the lab notebook before your lab, **you must attend at least the discussion portion of the lab to be checked off.** Note that your submitted work need not be complete in order to receive full credit if you were checked off.
#
#
# 2. Complete the lab on your own and submit the completed lab by Wednesday morning at 8:59 a.m. If you choose this route, you must finish the entire lab and pass all autograder tests to receive credit. Because missing lab means missing group discussion of important course concepts, we recommend that you don't use this option except in weeks when you are physically unable to come to lab. If you have finished your lab early, you can still attend and participate in the discussion.
#
# For Lab 1, however, you are **required** to attend lab and be checked off. For all other labs you can follow one of the two methods listed above.
#
# Collaborating on labs is more than okay -- it's encouraged! You should rarely remain stuck for more than a few minutes on questions in labs, so ask a neighbor or an instructor for help. (Explaining things is beneficial, too -- the best way to solidify your knowledge of a subject is to explain it.) Please don't just share answers, though.
#
# You can read more of the [course policies](http://data8.org/sp20/policies.html) on the [course website](http://data8.org/sp20/).
#
# #### Today's lab
#
# In today's lab, you'll learn how to:
#
# 1. navigate Jupyter notebooks (like this one);
# 2. write and evaluate some basic *expressions* in Python, the computer language of the course;
# 3. call *functions* to use code other people have written; and
# 4. break down Python code into smaller parts to understand it.
#
# This lab covers parts of [Chapter 3](http://www.inferentialthinking.com/chapters/03/programming-in-python.html) of the online textbook. You should read the examples in the book, but not right now. Instead, let's get started!
# -
# # 1. Jupyter notebooks
# This webpage is called a Jupyter notebook. A notebook is a place to write programs and view their results, and also to write text.
#
# ## 1.1. Text cells
# In a notebook, each rectangle containing text or code is called a *cell*.
#
# Text cells (like this one) can be edited by double-clicking on them. They're written in a simple format called [Markdown](http://daringfireball.net/projects/markdown/syntax) to add formatting and section headings. You don't need to learn Markdown, but you might want to.
#
# After you edit a text cell, click the "run cell" button at the top that looks like ▶| or hold down `shift` + `return` to confirm any changes. (Try not to delete the instructions of the lab.)
# + [markdown] deletable=false
# **Question 1.1.1.** This paragraph is in its own text cell. Try editing it so that this sentence is the last sentence in the paragraph, and then click the "run cell" ▶| button or hold down `shift` + `return`. This sentence, for example, should be deleted. So should this one.
# -
# ## 1.2. Code cells
# Other cells contain code in the Python 3 language. Running a code cell will execute all of the code it contains.
#
# To run the code in a code cell, first click on that cell to activate it. It'll be highlighted with a little green or blue rectangle. Next, either press ▶| or hold down `shift` + `return`.
#
# Try running this cell:
print("Hello, World!")
# And this one:
print("\N{WAVING HAND SIGN}, \N{EARTH GLOBE ASIA-AUSTRALIA}!")
# The fundamental building block of Python code is an expression. Cells can contain multiple lines with multiple expressions. When you run a cell, the lines of code are executed in the order in which they appear. Every `print` expression prints a line. Run the next cell and notice the order of the output.
# + deletable=false
print("First this line is printed,")
print("and then this one.")
# -
# **Question 1.2.1.** Change the cell above so that it prints out:
#
# First this line,
# then the whole 🌏,
# and then this one.
#
# *Hint:* If you're stuck on the Earth symbol for more than a few minutes, try talking to a neighbor or a staff member. That's a good idea for any lab problem.
# ## 1.3. Writing Jupyter notebooks
# You can use Jupyter notebooks for your own projects or documents. When you make your own notebook, you'll need to create your own cells for text and code.
#
# To add a cell, click the + button in the menu bar. It'll start out as a text cell. You can change it to a code cell by clicking inside it so it's highlighted, clicking the drop-down box next to the restart (⟳) button in the menu bar, and choosing "Code".
#
# **Question 1.3.1.** Add a code cell below this one. Write code in it that prints out:
#
# A whole new cell! ♪🌏♪
#
# (That musical note symbol is like the Earth symbol. Its long-form name is `\N{EIGHTH NOTE}`.)
#
# Run your cell to verify that it works.
# ## 1.4. Errors
# Python is a language, and like natural human languages, it has rules. It differs from natural language in two important ways:
# 1. The rules are *simple*. You can learn most of them in a few weeks and gain reasonable proficiency with the language in a semester.
# 2. The rules are *rigid*. If you're proficient in a natural language, you can understand a non-proficient speaker, glossing over small mistakes. A computer running Python code is not smart enough to do that.
#
# Whenever you write code, you'll make mistakes. When you run a code cell that has errors, Python will sometimes produce error messages to tell you what you did wrong.
#
# Errors are okay; even experienced programmers make many errors. When you make an error, you just have to find the source of the problem, fix it, and move on.
#
# We have made an error in the next cell. Run it and see what happens.
# + deletable=false
print("This line is missing something."
# + for_assignment_type="solution"
print("This line is missing something.")
# -
# **Note:** In the toolbar, there is the option to click `Cell > Run All`, which will run all the code cells in this notebook in order. However, the notebook stops running code cells if it hits an error, like the one in the cell above.
# You should see something like this (minus our annotations):
#
# <img src="error.jpg"/>
#
# The last line of the error output attempts to tell you what went wrong. The *syntax* of a language is its structure, and this `SyntaxError` tells you that you have created an illegal structure. "`EOF`" means "end of file," so the message is saying Python expected you to write something more (in this case, a right parenthesis) before finishing the cell.
#
# There's a lot of terminology in programming languages, but you don't need to know it all in order to program effectively. If you see a cryptic message like this, you can often get by without deciphering it. (Of course, if you're frustrated, ask a neighbor or a staff member for help.)
#
# Try to fix the code above so that you can run the cell and see the intended message instead of an error.
# ## 1.5. The Kernel
# The kernel is a program that executes the code inside your notebook and outputs the results. In the top right of your window, you can see a circle that indicates the status of your kernel. If the circle is empty (⚪), the kernel is idle and ready to execute code. If the circle is filled in (⚫), the kernel is busy running some code.
#
# Next to every code cell, you'll see some text that says `In [...]`. Before you run the cell, you'll see `In [ ]`. When the cell is running, you'll see `In [*]`. If you see an asterisk (\*) next to a cell that doesn't go away, it's likely that the code inside the cell is taking too long to run, and it might be a good time to interrupt the kernel (discussed below). When a cell is finished running, you'll see a number inside the brackets, like so: `In [1]`. The number corresponds to the order in which you run the cells; so, the first cell you run will show a 1 when it's finished running, the second will show a 2, and so on.
#
# You may run into problems where your kernel is stuck for an excessive amount of time, your notebook is very slow and unresponsive, or your kernel loses its connection. If this happens, try the following steps:
# 1. At the top of your screen, click **Kernel**, then **Interrupt**.
# 2. If that doesn't help, click **Kernel**, then **Restart**. If you do this, you will have to run your code cells from the start of your notebook up until where you paused your work.
# 3. If that doesn't help, restart your server. First, save your work by clicking **File** at the top left of your screen, then **Save and Checkpoint**. Next, click **Control Panel** at the top right. Choose **Stop My Server** to shut it down, then **Start My Server** to start it back up. Then, navigate back to the notebook you were working on. You'll still have to run your code cells again.
# ## 1.6. Submitting your work
# All assignments in the course will be distributed as notebooks like this one, and you will submit your work from the notebook. We will use a system called OK that checks your work and helps you submit. At the top of each assignment, you'll see a cell like the one below that prompts you to identify yourself. Run it and follow the instructions. Please use your <EMAIL> address when logging in.
# + deletable=false
# Don't change this cell; just run it.
# The result will give you directions about how to log in to the submission system, called OK.
# Once you're logged in, you can run this cell again,
# but it won't ask you who you are because it remembers you.
# However, you will need to log in once per assignment.
# When you log-in please hit return (not shift + return) after typing in your email
from client.api.notebook import Notebook
ok = Notebook('lab01.ok')
# -
# When you finish an assignment, you need to submit it by running the submit command below. It's fine to submit multiple times. OK will only try to grade your final submission for each assignment. Don't forget to submit your lab assignment at the end of section, even if you haven't finished everything.
# + deletable=false
_ = ok.submit()
# -
# # 2. Numbers
#
# Quantitative information arises everywhere in data science. In addition to representing commands to print out lines, expressions can represent numbers and methods of combining numbers. The expression `3.2500` evaluates to the number 3.25. (Run the cell and see.)
3.2500
# Notice that we didn't have to `print`. When you run a notebook cell, if the last line has a value, then Jupyter helpfully prints out that value for you. However, it won't print out prior lines automatically.
print(2)
3
4
# Above, you should see that 4 is the value of the last expression, 2 is printed, but 3 is lost forever because it was neither printed nor last.
#
# You don't want to print everything all the time anyway. But if you feel sorry for 3, change the cell above to print it.
# ## 2.1. Arithmetic
# The line in the next cell subtracts. Its value is what you'd expect. Run it.
3.25 - 1.5
# Many basic arithmetic operations are built into Python. The textbook section on [Expressions](http://www.inferentialthinking.com/chapters/03/1/expressions.html) describes all the arithmetic operators used in the course. The common operator that differs from typical math notation is `**`, which raises one number to the power of the other. So, `2**3` stands for $2^3$ and evaluates to 8.
#
# The order of operations is the same as what you learned in elementary school, and Python also has parentheses. For example, compare the outputs of the cells below. The second cell uses parentheses for a happy new year!
3+6*5-6*3**2*2**3/4*7
4+(6*5-(6*3))**2*((2**3)/4*7)
# In standard math notation, the first expression is
#
# $$3 + 6 \times 5 - 6 \times 3^2 \times \frac{2^3}{4} \times 7,$$
#
# while the second expression is
#
# $$3 + (6 \times 5 - (6 \times 3))^2 \times (\frac{(2^3)}{4} \times 7).$$
#
# **Question 2.1.1.** Write a Python expression in this next cell that's equal to $5 \times (3 \frac{10}{11}) - 50 \frac{1}{3} + 2^{.5 \times 22} - \frac{7}{33} + 3$. That's five times three and ten elevenths, minus fifty and a third, plus two to the power of half twenty-two, minus seven thirty-thirds plus three. By "$3 \frac{10}{11}$" we mean $3+\frac{10}{11}$, not $3 \times \frac{10}{11}$.
#
# Replace the ellipses (`...`) with your expression. Try to use parentheses only when necessary.
#
# *Hint:* The correct output should start with a familiar number.
# + deletable=false
...
# -
# # 3. Names
# In natural language, we have terminology that lets us quickly reference very complicated concepts. We don't say, "That's a large mammal with brown fur and sharp teeth!" Instead, we just say, "Bear!"
#
# In Python, we do this with *assignment statements*. An assignment statement has a name on the left side of an `=` sign and an expression to be evaluated on the right.
ten = 3 * 2 + 4
# When you run that cell, Python first computes the value of the expression on the right-hand side, `3 * 2 + 4`, which is the number 10. Then it assigns that value to the name `ten`. At that point, the code in the cell is done running.
#
# After you run that cell, the value 10 is bound to the name `ten`:
ten
# The statement `ten = 3 * 2 + 4` is not asserting that `ten` is already equal to `3 * 2 + 4`, as we might expect by analogy with math notation. Rather, that line of code changes what `ten` means; it now refers to the value 10, whereas before it meant nothing at all.
#
# If the designers of Python had been ruthlessly pedantic, they might have made us write
#
# define the name ten to hereafter have the value of 3 * 2 + 4
#
# instead. You will probably appreciate the brevity of "`=`"! But keep in mind that this is the real meaning.
#
# **Question 3.1.** Try writing code that uses a name (like `eleven`) that hasn't been assigned to anything. You'll see an error!
...
# A common pattern in Jupyter notebooks is to assign a value to a name and then immediately evaluate the name in the last line in the cell so that the value is displayed as output.
close_to_pi = 355/113
close_to_pi
# Another common pattern is that a series of lines in a single cell will build up a complex computation in stages, naming the intermediate results.
semimonthly_salary = 841.25
monthly_salary = 2 * semimonthly_salary
number_of_months_in_a_year = 12
yearly_salary = number_of_months_in_a_year * monthly_salary
yearly_salary
# Names in Python can have letters (upper- and lower-case letters are both okay and count as different letters), underscores, and numbers. The first character can't be a number (otherwise a name might look like a number). And names can't contain spaces, since spaces are used to separate pieces of code from each other.
#
# Other than those rules, what you name something doesn't matter *to Python*. For example, this cell does the same thing as the above cell, except everything has a different name:
a = 841.25
b = 2 * a
c = 12
d = c * b
d
# **However**, names are very important for making your code *readable* to yourself and others. The cell above is shorter, but it's totally useless without an explanation of what it does.
# ## 3.1. Checking your code
# Now that you know how to name things, you can start using the built-in *tests* to check whether your work is correct. Sometimes, there are multiple tests for a single question, and passing all of them is required to receive credit for the question. Please don't change the contents of the test cells.
#
# Go ahead and attempt Question 3.2. Running the cell directly after it will test whether you have assigned `seconds_in_a_decade` correctly in Question 3.2. If you haven't, this test will tell you the correct answer. Resist the urge to just copy it, and instead try to adjust your expression. (Sometimes the tests will give hints about what went wrong...)
# + [markdown] deletable=false editable=false
# **Question 3.1.2.** Assign the name `seconds_in_a_decade` to the number of seconds between midnight January 1, 2010 and midnight January 1, 2020. Note that there are two leap years in this span of a decade. A non-leap year has 365 days and a leap year has 366 days.
#
# *Hint:* If you're stuck, the next section shows you how to get hints.
#
# <!--
# BEGIN QUESTION
# name: q3_1_2
# -->
# + deletable=false
# Change the next line
# so that it computes the number of seconds in a decade
# and assigns that number the name, seconds_in_a_decade.
seconds_in_a_decade = ...
# We've put this line in this cell
# so that it will print the value you've given to seconds_in_a_decade when you run it.
# You don't need to change this.
seconds_in_a_decade
# + deletable=false editable=false
ok.grade("q3_1_2");
# -
# # 3.2. Comments
# You may have noticed these lines in the cell in which you answered Question 3.2:
#
# # Change the next line
# # so that it computes the number of seconds in a decade
# # and assigns that number the name, seconds_in_a_decade.
#
# This is called a *comment*. It doesn't make anything happen in Python; Python ignores anything on a line after a `#`. Instead, it's there to communicate something about the code to you, the human reader. Comments are extremely useful.
#
#
# <img src="http://imgs.xkcd.com/comics/future_self.png">
# Source: http://imgs.xkcd.com/comics/future_self.png
# ## 3.3. Application: A physics experiment
#
# On the Apollo 15 mission to the Moon, astronaut <NAME> famously replicated Galileo's physics experiment in which he showed that gravity accelerates objects of different mass at the same rate. Because there is no air resistance for a falling object on the surface of the Moon, even two objects with very different masses and densities should fall at the same rate. <NAME> compared a feather and a hammer.
#
# You can run the following cell to watch a video of the experiment.
# +
from IPython.display import YouTubeVideo
# The original URL is:
# https://www.youtube.com/watch?v=U7db6ZeLR5s
YouTubeVideo("U7db6ZeLR5s")
# -
# Here's the transcript of the video:
#
# **167:22:06 Scott**: Well, in my left hand, I have a feather; in my right hand, a hammer. And I guess one of the reasons we got here today was because of a gentleman named Galileo, a long time ago, who made a rather significant discovery about falling objects in gravity fields. And we thought where would be a better place to confirm his findings than on the Moon. And so we thought we'd try it here for you. The feather happens to be, appropriately, a falcon feather for our Falcon. And I'll drop the two of them here and, hopefully, they'll hit the ground at the same time.
#
# **167:22:43 Scott**: How about that!
#
# **167:22:45 Allen**: How about that! (Applause in Houston)
#
# **167:22:46 Scott**: Which proves that Mr. Galileo was correct in his findings.
# **Newton's Law.** Using this footage, we can also attempt to confirm another famous bit of physics: Newton's law of universal gravitation. Newton's laws predict that any object dropped near the surface of the Moon should fall
#
# $$\frac{1}{2} G \frac{M}{R^2} t^2 \text{ meters}$$
#
# after $t$ seconds, where $G$ is a universal constant, $M$ is the moon's mass in kilograms, and $R$ is the moon's radius in meters. So if we know $G$, $M$, and $R$, then Newton's laws let us predict how far an object will fall over any amount of time.
#
# To verify the accuracy of this law, we will calculate the difference between the predicted distance the hammer drops and the actual distance. (If they are different, it might be because Newton's laws are wrong, or because our measurements are imprecise, or because there are other factors affecting the hammer for which we haven't accounted.)
#
# Someone studied the video and estimated that the hammer was dropped 113 cm from the surface. Counting frames in the video, the hammer falls for 1.2 seconds (36 frames).
# + [markdown] deletable=false editable=false
# **Question 3.3.1.** Complete the code in the next cell to fill in the data from the experiment.
#
# *Hint:* No computation required; just fill in data from the paragraph above.
#
# <!--
# BEGIN QUESTION
# name: q3_3_1
# -->
# + deletable=false
# t, the duration of the fall in the experiment, in seconds.
# Fill this in.
time = ...
# The estimated distance the hammer actually fell, in meters.
# Fill this in.
estimated_distance_m = ...
# + deletable=false editable=false
ok.grade("q3_3_1");
# + [markdown] deletable=false editable=false
# **Question 3.3.2.** Now, complete the code in the next cell to compute the difference between the predicted and estimated distances (in meters) that the hammer fell in this experiment.
#
# This just means translating the formula above ($\frac{1}{2}G\frac{M}{R^2}t^2$) into Python code. You'll have to replace each variable in the math formula with the name we gave that number in Python code.
#
# *Hint:* Try to use variables you've already defined in question 3.3.1
#
# <!--
# BEGIN QUESTION
# name: q3_3_2
# -->
# + deletable=false
# First, we've written down the values of the 3 universal constants
# that show up in Newton's formula.
# G, the universal constant measuring the strength of gravity.
gravity_constant = 6.674 * 10**-11
# M, the moon's mass, in kilograms.
moon_mass_kg = 7.34767309 * 10**22
# R, the radius of the moon, in meters.
moon_radius_m = 1.737 * 10**6
# The distance the hammer should have fallen
# over the duration of the fall, in meters,
# according to Newton's law of gravity.
# The text above describes the formula
# for this distance given by Newton's law.
# **YOU FILL THIS PART IN.**
predicted_distance_m = ...
# Here we've computed the difference
# between the predicted fall distance and the distance we actually measured.
# If you've filled in the above code, this should just work.
difference = predicted_distance_m - estimated_distance_m
difference
# + deletable=false editable=false
ok.grade("q3_3_2");
# -
# ## 4. Calling functions
#
# The most common way to combine or manipulate values in Python is by calling functions. Python comes with many built-in functions that perform common operations.
#
# For example, the `abs` function takes a single number as its argument and returns the absolute value of that number. Run the next two cells and see if you understand the output.
abs(5)
abs(-5)
# + [markdown] deletable=false editable=false
# ## 4.1. Application: Computing walking distances
# Chunhua is on the corner of 7th Avenue and 42nd Street in Midtown Manhattan, and she wants to know far she'd have to walk to get to Gramercy School on the corner of 10th Avenue and 34th Street.
#
# She can't cut across blocks diagonally, since there are buildings in the way. She has to walk along the sidewalks. Using the map below, she sees she'd have to walk 3 avenues (long blocks) and 8 streets (short blocks). In terms of the given numbers, she computed 3 as the difference between 7 and 10, *in absolute value*, and 8 similarly.
#
# Chunhua also knows that blocks in Manhattan are all about 80m by 274m (avenues are farther apart than streets). So in total, she'd have to walk $(80 \times |42 - 34| + 274 \times |7 - 10|)$ meters to get to the park.
#
# <img src="map.jpg"/>
#
# **Question 4.1.1.** Fill in the line `num_avenues_away = ...` in the next cell so that the cell calculates the distance Chunhua must walk and gives it the name `manhattan_distance`. Everything else has been filled in for you. **Use the `abs` function.** Also, be sure to run the test cell afterward to test your code.
#
# <!--
# BEGIN QUESTION
# name: q4_1_1
# -->
# + deletable=false
# Here's the number of streets away:
num_streets_away = abs(42-34)
# Compute the number of avenues away in a similar way:
num_avenues_away = ...
street_length_m = 80
avenue_length_m = 274
# Now we compute the total distance Chunhua must walk.
manhattan_distance = street_length_m*num_streets_away + avenue_length_m*num_avenues_away
# We've included this line so that you see the distance you've computed
# when you run this cell.
# You don't need to change it, but you can if you want.
manhattan_distance
# + deletable=false editable=false
ok.grade("q4_1_1");
# -
# ##### Multiple arguments
# Some functions take multiple arguments, separated by commas. For example, the built-in `max` function returns the maximum argument passed to it.
max(2, -3, 4, -5)
# # 5. Understanding nested expressions
# Function calls and arithmetic expressions can themselves contain expressions. You saw an example in the last question:
#
# abs(42-34)
#
# has 2 number expressions in a subtraction expression in a function call expression. And you probably wrote something like `abs(7-10)` to compute `num_avenues_away`.
#
# Nested expressions can turn into complicated-looking code. However, the way in which complicated expressions break down is very regular.
#
# Suppose we are interested in heights that are very unusual. We'll say that a height is unusual to the extent that it's far away on the number line from the average human height. [An estimate](http://press.endocrine.org/doi/full/10.1210/jcem.86.9.7875?ck=nck&) of the average adult human height (averaging, we hope, over all humans on Earth today) is 1.688 meters.
#
# So if Kayla is 1.21 meters tall, then her height is $|1.21 - 1.688|$, or $.478$, meters away from the average. Here's a picture of that:
#
# <img src="numberline_0.png">
#
# And here's how we'd write that in one line of Python code:
abs(1.21 - 1.688)
# What's going on here? `abs` takes just one argument, so the stuff inside the parentheses is all part of that *single argument*. Specifically, the argument is the value of the expression `1.21 - 1.688`. The value of that expression is `-.478`. That value is the argument to `abs`. The absolute value of that is `.478`, so `.478` is the value of the full expression `abs(1.21 - 1.688)`.
#
# Picture simplifying the expression in several steps:
#
# 1. `abs(1.21 - 1.688)`
# 2. `abs(-.478)`
# 3. `.478`
#
# In fact, that's basically what Python does to compute the value of the expression.
# + [markdown] deletable=false editable=false
# **Question 5.1.** Say that Paola's height is 1.76 meters. In the next cell, use `abs` to compute the absolute value of the difference between Paola's height and the average human height. Give that value the name `paola_distance_from_average_m`.
#
# <img src="numberline_1.png">
#
# <!--
# BEGIN QUESTION
# name: q51
# -->
# + deletable=false
# Replace the ... with an expression
# to compute the absolute value
# of the difference between Paola's height (1.76m) and the average human height.
paola_distance_from_average_m = ...
# Again, we've written this here
# so that the distance you compute will get printed
# when you run this cell.
paola_distance_from_average_m
# + deletable=false editable=false
ok.grade("q51");
# -
# ## 5.1. More nesting
# Now say that we want to compute the more unusual of the two heights. We'll use the function `max`, which (again) takes two numbers as arguments and returns the larger of the two arguments. Combining that with the `abs` function, we can compute the larger distance from average among the two heights:
# +
# Just read and run this cell.
kayla_height_m = 1.21
paola_height_m = 1.76
average_adult_height_m = 1.688
# The larger distance from the average human height, among the two heights:
larger_distance_m = max(abs(kayla_height_m - average_adult_height_m), abs(paola_height_m - average_adult_height_m))
# Print out our results in a nice readable format:
print("The larger distance from the average height among these two people is", larger_distance_m, "meters.")
# -
# The line where `larger_distance_m` is computed looks complicated, but we can break it down into simpler components just like we did before.
#
# The basic recipe is to repeatedly simplify small parts of the expression:
# * **Basic expressions:** Start with expressions whose values we know, like names or numbers.
# - Examples: `paola_height_m` or `5`.
# * **Find the next simplest group of expressions:** Look for basic expressions that are directly connected to each other. This can be by arithmetic or as arguments to a function call.
# - Example: `kayla_height_m - average_adult_height_m`.
# * **Evaluate that group:** Evaluate the arithmetic expression or function call. Use the value computed to replace the group of expressions.
# - Example: `kayla_height_m - average_adult_height_m` becomes `-.478`.
# * **Repeat:** Continue this process, using the value of the previously-evaluated expression as a new basic expression. Stop when we've evaluated the entire expression.
# - Example: `abs(-.478)` becomes `.478`, and `max(.478, .072)` becomes `.478`.
#
# You can run the next cell to see a slideshow of that process.
from IPython.display import IFrame
IFrame('https://docs.google.com/presentation/d/e/2PACX-1vTiIUOa9tP4pHPesrI8p2TCp8WCOJtTb3usOacQFPfkEfvQMmX-JYEW3OnBoTmQEJWAHdBP6Mvp053G/embed?start=false&loop=false&delayms=3000', 800, 600)
# + [markdown] deletable=false editable=false
# Ok, your turn.
#
# **Question 5.1.1.** Given the heights of players from the Golden State Warriors, write an expression that computes the smallest difference between any of the three heights. Your expression shouldn't have any numbers in it, only function calls and the names `klay`, `steph`, and `dangelo`. Give the value of your expression the name `min_height_difference`.
#
# <!--
# BEGIN QUESTION
# name: q5_1_1
# -->
# + deletable=false
# The three players' heights, in meters:
klay = 2.01 # <NAME> is 6'7"
steph = 1.91 # <NAME> is 6'3"
dangelo = 1.95 # <NAME> is 6'5"
# We'd like to look at all 3 pairs of heights,
# compute the absolute difference between each pair,
# and then find the smallest of those 3 absolute differences.
# This is left to you!
# If you're stuck, try computing the value for each step of the process
# (like the difference between Klay's heigh and Steph's height)
# on a separate line and giving it a name (like klay_steph_height_diff)6
min_height_difference = ...
# + deletable=false editable=false
ok.grade("q5_1_1");
# -
# You're done with Lab 1! Be sure to run the tests and verify that they all pass, then choose **Save and Checkpoint** from the **File** menu, then **run the final cell** to submit your work. If you submit multiple times, your last submission will be counted.
# + deletable=false
# For your convenience, you can run this cell to run all the tests at once!
import os
_ = [ok.grade(q[:-3]) for q in os.listdir("tests") if q.startswith('q')]
# -
# **Important.** Before you leave lab, run this final cell to submit your work. Now, get checked off by flagging down a staff member.
# + deletable=false
_ = ok.submit()
| materials/sp20/lab/lab01/lab01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pte
# language: python
# name: pte
# ---
# + [markdown] origin_pos=0
# # 多层感知机的从零开始实现
# :label:`sec_mlp_scratch`
#
# 我们已经在数学上描述了多层感知机(MLP),现在让我们尝试自己实现一个多层感知机。为了与我们之前使用softmax回归( :numref:`sec_softmax_scratch` )获得的结果进行比较,我们将继续使用Fashion-MNIST图像分类数据集( :numref:`sec_fashion_mnist`)。
#
# + origin_pos=2 tab=["pytorch"]
import paddle
from paddle import nn
from d2l import torch as d2l
import pd2l
# + origin_pos=4 tab=["pytorch"]
batch_size = 256
train_iter, test_iter = pd2l.load_data_fashion_mnist(batch_size)
# + [markdown] origin_pos=5
# ## 初始化模型参数
#
# 回想一下,Fashion-MNIST中的每个图像由$28 \times 28 = 784$个灰度像素值组成。所有图像共分为10个类别。忽略像素之间的空间结构,我们可以将每个图像视为具有784个输入特征和10个类的简单分类数据集。首先,我们将[**实现一个具有单隐藏层的多层感知机,它包含256个隐藏单元**]。注意,我们可以将这两个量都视为超参数。通常,我们选择2的若干次幂作为层的宽度。因为内存在硬件中的分配和寻址方式,这么做往往可以在计算上更高效。
#
# 我们用几个张量来表示我们的参数。注意,对于每一层我们都要记录一个权重矩阵和一个偏置向量。跟以前一样,我们要为这些参数的损失的梯度分配内存。
#
# + origin_pos=7 tab=["pytorch"]
num_inputs, num_outputs, num_hiddens = 784, 10, 256
W1 = paddle.randn([num_inputs, num_hiddens]) * 0.01
W1.stop_gradient = False
b1 = paddle.zeros([num_hiddens])
b1.stop_gradient = False
W2 = paddle.randn([num_hiddens, num_outputs]) * 0.01
W2.stop_gradient = False
b2 = paddle.zeros([num_outputs])
b2.stop_gradient = False
params = [W1, b1, W2, b2]
# + [markdown] origin_pos=9
# ## 激活函数
#
# 为了确保我们知道一切是如何工作的,我们将使用最大值函数自己[**实现ReLU激活函数**],而不是直接调用内置的`relu`函数。
#
# + origin_pos=11 tab=["pytorch"]
def relu(X):
a = paddle.zeros_like(X)
return paddle.max(X, a)
# + [markdown] origin_pos=13
# ## 模型
#
# 因为我们忽略了空间结构,所以我们使用`reshape`将每个二维图像转换为一个长度为`num_inputs`的向量。我们只需几行代码就可以(**实现我们的模型**)。
#
# + origin_pos=15 tab=["pytorch"]
def net(X):
X = X.reshape((-1, num_inputs))
H = relu(X@W1 + b1) # 这里“@”代表矩阵乘法
return (H@W2 + b2)
# + [markdown] origin_pos=17
# ## 损失函数
#
# 为了确保数值稳定性,同时由于我们已经从零实现过softmax函数( :numref:`sec_softmax_scratch` ),因此在这里我们直接使用高级API中的内置函数来计算softmax和交叉熵损失。回想一下我们之前在 :numref:`subsec_softmax-implementation-revisited` 中对这些复杂问题的讨论。我们鼓励感兴趣的读者查看损失函数的源代码,以加深对实现细节的了解。
#
# + origin_pos=19 tab=["pytorch"]
loss = nn.CrossEntropyLoss()
# + [markdown] origin_pos=21
# ## 训练
#
# 幸运的是,[**多层感知机的训练过程与softmax回归的训练过程完全相同**]。可以直接调用`d2l`包的`train_ch3`函数(参见 :numref:`sec_softmax_scratch` ),将迭代周期数设置为10,并将学习率设置为0.1.
#
# + origin_pos=23 tab=["pytorch"]
num_epochs, lr = 10, 0.1
updater = paddle.optimizer.SGD(learning_rate=lr, parameters=params)
pd2l.train_ch3(net, train_iter, test_iter, loss, num_epochs, updater)
# + [markdown] origin_pos=25
# 为了对学习到的模型进行评估,我们将[**在一些测试数据上应用这个模型**]。
#
# + origin_pos=26 tab=["pytorch"]
d2l.predict_ch3(net, test_iter)
# + [markdown] origin_pos=27
# ## 小结
#
# * 我们看到即使手动实现一个简单的多层感知机也是很容易的。
# * 然而,如果有大量的层,从零开始实现多层感知机会变得很麻烦(例如,要命名和记录模型的参数)。
#
# ## 练习
#
# 1. 在所有其他参数保持不变的情况下,更改超参数`num_hiddens`的值,并查看此超参数的变化对结果有何影响。确定此超参数的最佳值。
# 1. 尝试添加更多的隐藏层,并查看它对结果有何影响。
# 1. 改变学习速率会如何影响结果?保持模型结构和其他超参数(包括迭代周期数)不变,学习率设置为多少会带来最好的结果?
# 1. 通过对所有超参数(学习率、迭代周期数、隐藏层数、每层的隐藏单元数)进行联合优化,可以得到的最佳结果是什么?
# 1. 描述为什么涉及多个超参数更具挑战性。
# 1. 如果要构建多个超参数的搜索方法,你能想到的最聪明的策略是什么?
#
# + [markdown] origin_pos=29 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1804)
#
| Dive-into-DL-paddlepaddle/docs/4_multilayer-perceptrons/4.2mlp-scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Udacity Self-Driving Car Engineer Nanodegree Program
# ## *Vehicle Detection Project*
#
# [](http://www.udacity.com/drive)
# <img src="output_images/project_video_out_pipeline.png" width="480" alt="Combined Image" />
#
# ---
#
# The goals / steps of this project are the following:
#
# * Perform a Histogram of Oriented Gradients (HOG) feature extraction on a labeled training set of images and train a classifier Linear SVM classifier
# * Optionally, you can also apply a color transform and append binned color features, as well as histograms of color, to your HOG feature vector.
# * Note: for those first two steps don't forget to normalize your features and randomize a selection for training and testing.
# * Implement a sliding-window technique and use your trained classifier to search for vehicles in images.
# * Run your pipeline on a video stream (start with the test_video.mp4 and later implement on full project_video.mp4) and create a heat map of recurring detections frame by frame to reject outliers and follow detected vehicles.
# * Estimate a bounding box for vehicles detected.
#
# [//]: # (Image References)
# [image1]: ./output_images/data_visualization.png
# [image2]: ./output_images/HOG_Car_and_NonCar1.png
# [image3]: ./output_images/HOG_Car_and_NonCar2.png
# [image4]: ./output_images/bounding_box_78.png
# [image5]: ./output_images/bounding_box_50.png
# [image6]: ./output_images/bounding_box_38.png
# [image7]: ./output_images/bounding_box_24.png
# [image8]: ./output_images/bounding_box_combined_window.png
# [image9]: ./output_images/heat_map.png
# [image9a]: ./output_images/heat_map_threshold.png
# [image10]: ./output_images/heat_map_sci_py.png
# [image10a]: ./output_images/final_detect.png
# [image11]: ./output_images/TestImagePipeline.png
# [image12]: ./output_images/test_video_out_pipeline.png
# [image13]: ./output_images/project_video_out_pipeline.png
# [image14]: ./output_images/bounding_box.png
# [video1]: ./test_video_out.mp4
# [video2]: ./test_video_out_2.mp4
# [video3]: ./project_video_out.mp4
#
#
# ## [Rubric](https://review.udacity.com/#!/rubrics/513/view) Points
# ### Here I will consider the rubric points individually and describe how I addressed each point in my implementation.
#
# ---
# ### Writeup / README
#
# #### 1. Provide a Writeup / README that includes all the rubric points and how you addressed each one.
#
# In this project, computer vision methods are used. It is not the only technique — deep learning could be used instead. The advantage of CV is that we can analyze, extract features in each step. Deep learning, in contrast, is more like a black box. The code is contained in Vehicle_Detection.ipynb
#
# ### Histogram of Oriented Gradients (HOG)
#
# #### 1. Explain how (and identify where in your code) you extracted HOG features from the training images.
#
#
# Algorithm needs to learn to detect a car in picture. The best way is to train the algorithm with lot of images, labeled "cars" and "non cars". I started by reading in all the `vehicle` and `non-vehicle` images. There are 8792 car images and 8968 non car Images which are 64*64 pixels.
# Here is an random example of the `vehicle` and `non-vehicle` classes.
#
# ![alt text][image1]
#
# HOG stands for “Histogram of Oriented Gradients”. Basically, it divides an image in several pieces. For each piece, it calculates the gradient of variation in a given number of orientations. I explored different color spaces and different `skimage.hog()` parameters (`orientations`, `pixels_per_cell`, and `cells_per_block`). I grabbed random images from each of the two classes and displayed them to get a feel for what the `skimage.hog()` output looks like.
#
# The less the number of pixels per cells (and other parameters), more general the data. The high the number of pixels per cells (and other parameters) more specific. By playing with the parameters, I found that orientations above 7, and 8 pixels per cell are enough to identify a car.
#
# Here is an example using the `YCrCb` color space and HOG parameters of `orientations=8`, `pixels_per_cell=(8, 8)` and `cells_per_block=(2, 2)`:
#
# The code for extracting HOG features from an image is defined by the method `get_hog_features`. The figure below shows a comparison of a car image and its associated histogram of oriented gradients, as well as the same for a non-car image.
#
# ![alt text][image2]
#
# The method `extract_features` in the section titled "Method to Extract HOG Features from an Array of Car and Non-Car Images" accepts a list of image paths and HOG parameters (as well as one of a variety of destination color spaces, to which the input image is converted), and produces a flattened array of HOG features for each image in the list.
#
# Next, in the section titled "Extract Features for Input Datasets and Combine, Define Labels Vector, Shuffle and Split," I define parameters for HOG feature extraction and extract features for the entire dataset. These feature sets are combined and a label vector is defined (`1` for cars, `0` for non-cars). The features and labels are then shuffled and split into training and test sets in preparation to be fed to a linear support vector machine (SVM) classifier. The table below documents the twenty-five different parameter combinations that I explored.
#
# | Configuration Label | Colorspace | Orientations | Pixels Per Cell | Cells Per Block | HOG Channel | Extract Time |
# | :-----------------: | :--------: | :----------: | :-------------: | :-------------: | :---------: | ------------:|
# | 1 | RGB | 9 | 8 | 2 | ALL | 71.16 |
# | 2 | HSV | 9 | 8 | 2 | 1 | 43.74 |
# | 3 | HSV | 9 | 8 | 2 | 2 | 36.35 |
# | 4 | LUV | 9 | 8 | 2 | 0 | 37.42 |
# | 5 | LUV | 9 | 8 | 2 | 1 | 38.34 |
# | 6 | HLS | 9 | 8 | 2 | 0 | 37.42 |
# | 7 | HLS | 9 | 8 | 2 | 1 | 42.04 |
# | 8 | YUV | 9 | 8 | 2 | 0 | 35.86 |
# | 9 | YCrCb | 9 | 8 | 2 | 1 | 38.32 |
# | 10 | YCrCb | 9 | 8 | 2 | 2 | 38.99 |
# | 11 | HSV | 9 | 8 | 2 | ALL | 79.72 |
# | 12 | LUV | 9 | 8 | 2 | ALL | 78.57 |
# | 13 | HLS | 9 | 8 | 2 | ALL | 81.37 |
# | 14 | YUV | 9 | 8 | 2 | ALL | 81.82 |
# | 15 | YCrCb | 9 | 8 | 2 | ALL | 79.05 |
# | 16 | YUV | 9 | 8 | 1 | 0 | 44.04 |
# | 17 | YUV | 9 | 8 | 3 | 0 | 37.74 |
# | 18 | YUV | 6 | 8 | 2 | 0 | 37.12 |
# | 19 | YUV | 12 | 8 | 2 | 0 | 40.11 |
# | 20 | YUV | 11 | 8 | 2 | 0 | 38.01 |
# | 21 | YUV | 11 | 16 | 2 | 0 | 30.21 |
# | 22 | YUV | 11 | 12 | 2 | 0 | 30.33 |
# | 23 | YUV | 11 | 4 | 2 | 0 | 69.08 |
# | 24 | YUV | 7 | 16 | 2 | ALL | 53.18 |
# | __25__ | __YUV__| __11__ | __16__ | __2__ |__ALL__ | __55.20 __|
#
# The final parameters chosen were those labeled __Configuration 25__ in the table above: `YUV colorspace`, `orientations=11`, `pixels per cell = 16` , `cells per block = 2` , and `hog_channel = ALL` of the colorspace.
#
# With these parameters -
# __57.15 Seconds to extract HOG features...
# Using: 11 orientations 16 pixels per cell and 2 cells per block
# Feature vector length: 1188__
#
# #### 2. Explain how you settled on your final choice of HOG parameters.
#
# The next step is to train a classifier. It receives the cars / non-cars data transformed with HOG detector, and returns if the sample is or is not a car.
# I settled on my final choice of HOG parameters based upon the performance of the SVM classifier produced using them. It gave a Test accuracy of __97.8%__. I considered not only the accuracy with which the classifier made predictions on the test dataset, but also the speed at which the classifier is able to make predictions. I also tried with different type of Classifiers such as __NuSVC__, __SVC__ and __LinearSVC__. There is a balance to be struck between accuracy and speed of the classifier, and my strategy was to bias toward speed first, and achieve as close to real-time predictions as possible, and then pursue accuracy if the detection pipeline were not to perform satisfactorily.
#
# 269.09 Seconds to train __NuSVC__...
# Test Accuracy of NuSVC = 0.9451
# SVC predicts: [ 0. 0. 1. 0. 0. 0. 0. 1. 1. 0.]
# For these 10 labels: [ 0. 0. 1. 0. 0. 0. 0. 1. 1. 0.]
# 0.10513 Seconds to predict 10 labels with NuSVC
#
# 264.76 Seconds to train __SVC__...
# Test Accuracy of SVC = 0.8953
# SVC predicts: [ 0. 0. 1. 0. 0. 0. 0. 1. 1. 0.]
# For these 10 labels: [ 0. 0. 1. 0. 0. 0. 0. 1. 1. 0.]
# 0.18816 Seconds to predict 10 labels with SVC
#
# 1.08 Seconds to train __LinearSVC__...
# Test Accuracy of SVC = __0.978__
# SVC predicts: [ 0. 0. 1. 0. 0. 1. 0. 1. 1. 0.]
# For these 10 labels: [ 0. 0. 1. 0. 0. 0. 0. 1. 1. 0.]
# 0.002 Seconds to predict 10 labels with LinearSVC
#
# The classifier performance of each of the configurations from the table above are summarized in the table below:
#
# | Configuration (above) | Classifier | Accuracy | Train Time |
# | :-------------------: | :--------: | -------: | ---------: |
# | 1 | Linear SVC | 97.52 | 18.21 |
# | 2 | Linear SVC | 91.92 | 5.33 |
# | 3 | Linear SVC | 96.09 | 4.56 |
# | 4 | Linear SVC | 95.72 | 4.33 |
# | 5 | Linear SVC | 94.51 | 4.51 |
# | 6 | Linear SVC | 92.34 | 4.97 |
# | 7 | Linear SVC | 95.81 | 4.04 |
# | 8 | Linear SVC | 96.28 | 5.04 |
# | 9 | Linear SVC | 94.88 | 4.69 |
# | 10 | Linear SVC | 93.78 | 4.59 |
# | 11 | Linear SVC | 98.31 | 16.03 |
# | 12 | Linear SVC | 97.52 | 14.67 |
# | 13 | Linear SVC | 98.42 | 13.46 |
# | 14 | Linear SVC | 98.40 | 15.68 |
# | 15 | Linear SVC | 98.06 | 13.86 |
# | 16 | Linear SVC | 94.76 | 5.11 |
# | 17 | Linear SVC | 96.11 | 6.71 |
# | 18 | Linear SVC | 95.81 | 3.79 |
# | 19 | Linear SVC | 95.95 | 4.84 |
# | 20 | Linear SVC | 96.59 | 5.46 |
# | 21 | Linear SVC | 95.16 | 0.59 |
# | 22 | Linear SVC | 94.85 | 1.27 |
# | 23 | Linear SVC | 95.92 | 20.39 |
# | 24 | Linear SVC | 97.61 | 1.23 |
# | __25__ | __Linear SVC__ | __97.8__ | __1.08__|
#
#
#
# #### 3. Describe how (and identify where in your code) you trained a classifier using your selected HOG features (and color features if you used them).
#
# In the section titled "Train a Classifier" I trained a linear SVM with the default classifier parameters and using HOG features alone (I did not use spatial intensity or channel intensity histogram features) and was able to achieve a test accuracy of __97.8%__.
#
#
# ### Sliding Window Search
#
# #### 1. Describe how (and identify where in your code) you implemented a sliding window search. How did you decide what scales to search and how much to overlap windows?
#
# I adapted the method `find_cars` from the lesson materials. The method combines HOG feature extraction with a sliding window search, but rather than perform feature extraction on each window individually which can be time consuming, the HOG features are extracted for the entire image (or a selected portion of it) and then these full-image features are subsampled according to the size of the window and then fed to the classifier. The method performs the classifier prediction on the HOG features for each window region and returns a list of rectangle objects corresponding to the windows that generated a positive ("car") prediction.
#
# The image below shows the first attempt at using `find_cars` on one of the test images, using a single window size:
#
# ![alt text][image14]
#
# I explored several configurations of window sizes and positions, with various overlaps in the X and Y directions. The following four images show the configurations of all search windows in the final implementation, for small (1x), medium (1.5x, 2x), and large (3x) windows:
#
# ![alt text][image4]
# ![alt text][image5]
# ![alt text][image6]
# ![alt text][image7]
#
# The final algorithm calls `find_cars` for each window scale and the rectangles returned from each method call are aggregated. In previous implementations smaller (0.5) scales were explored but found to return too many false positives, and originally the window overlap was set to 50% in both X and Y directions, but an overlap of 75% in the Y direction (yet still 50% in the X direction) produced more redundant true positive detections, which were preferable given the heatmap strategy described below. Additionally, only an appropriate vertical range of the image is considered for each window size (e.g. smaller range for smaller scales) to reduce the chance for false positives in areas where cars at that scale are unlikely to appear.
#
# The image below shows the rectangles returned by `find_cars` drawn onto one of the test images in the final implementation. Notice that there are several positive predictions on each of the near-field cars.
#
# ![alt text][image8]
#
# Because a true positive is typically accompanied by several positive detections, while false positives are typically accompanied by only one or two detections, a combined heatmap and threshold is used to differentiate the two. The `add_heat` function increments the pixel value (referred to as "heat") of an all-black image the size of the original image at the location of each detection rectangle. Areas encompassed by more overlapping rectangles are assigned higher levels of heat. The following image is the resulting heatmap from the detections in the image above:
#
# ![alt text][image9]
#
# A threshold is applied to the heatmap (in this example, with a value of 1), setting all pixels that don't exceed the threshold to zero. The result is below:
#
# ![alt text][image9a]
#
# The `scipy.ndimage.measurements.label()` function collects spatially contiguous areas of the heatmap and assigns each a label:
#
# ![alt text][image10]
#
# And the final detection area is set to the extremities of each identified label:
#
# ![alt text][image10a]
#
# #### 2. Show some examples of test images to demonstrate how your pipeline is working. What did you do to optimize the performance of your classifier?
#
# The results of passing all of the project test images through the above pipeline are displayed in the images below:
#
# ![alt text][image11]
#
# The final implementation performs very well, identifying the near-field vehicles in each of the images with no false positives.
#
# The first implementation did not perform as well, so I began by optimizing the SVM classifier. The original classifier used HOG features from the YUV channel only, and achieved a test accuracy of 96.28%. Using all three YUV channels increased the accuracy to 98.40%, but also tripled the execution time. However, changing the `pixels_per_cell` parameter from 8 to 16 produced a roughly ten-fold increase in execution speed with minimal cost to accuracy (97.80%).
#
# Other optimization techniques included changes to window sizing and overlap as described above, and lowering the heatmap threshold to improve accuracy of the detection (higher threshold values tended to underestimate the size of the vehicle).
#
# ---
#
# ### Video Implementation
#
# #### 1. Provide a link to your final video output. Your pipeline should perform reasonably well on the entire project video (somewhat wobbly or unstable bounding boxes are ok as long as you are identifying the vehicles most of the time with minimal false positives.)
# Here's a link to my video result:
# __./project_video_out.mp4__
#
#
# #### 2. Describe how (and identify where in your code) you implemented some kind of filter for false positives and some method for combining overlapping bounding boxes.
#
#
# The code for processing frames of video is contained in the cell titled "Pipeline for Processing Video Frames" and is identical to the code for processing a single image described above, with the exception of storing the detections (returned by `find_cars`) from the previous 15 frames of video using the `prev_rects` parameter from a class called `Vehicle_Detect`. Rather than performing the heatmap/threshold/label steps for the current frame's detections, the detections for the past 15 frames are combined and added to the heatmap and the threshold for the heatmap is set to `1 + len(det.prev_rects)//2` (one more than half the number of rectangle sets contained in the history) - this value was found to perform best empirically (rather than using a single scalar, or the full number of rectangle sets in the history).
#
# ---
#
# ### Discussion
#
# #### 1. Briefly discuss any problems / issues you faced in your implementation of this project. Where will your pipeline likely fail? What could you do to make it more robust?
#
# The problems that I faced while implementing this project were mainly concerned with detection accuracy. Balancing the accuracy of the classifier with execution speed was crucial. Example, Scanning 190 windows using a classifier that achieves ~98% accuracy should result in around 4 misidentified windows per frame. Of course, integrating detections from previous frames mitigates the effect of the misclassifications, but it also introduces another problem: vehicles that significantly change position from one frame to the next (e.g. oncoming traffic) will tend to escape being labeled. Producing a very high accuracy classifier and maximizing window overlap might improve the per-frame accuracy to the point that integrating detections from previous frames is unnecessary (and oncoming traffic is correctly labeled), but it would also be far from real-time without massive processing power.
#
# The pipeline is probably most likely to fail in cases where vehicles (or the HOG features thereof) don't resemble those in the training dataset, but lighting and environmental conditions might also play a role (e.g. a white car against a white background) .As stated above, oncoming cars are an issue, as well as distant cars (as mentioned earlier, smaller window scales tended to produce more false positives, but they also did not often correctly label the smaller, distant cars).
#
# I believe better approach would be to combine a very high accuracy classifier with high overlap in the search windows. The execution cost could be offset with more intelligent tracking strategies, such as:
# - determine vehicle location and speed to predict its location in subsequent frames
# - use a convolutional neural network, to preclude the sliding window search altogether
| .ipynb_checkpoints/writeup-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import px4tools
import pandas
import pylab as pl
# %matplotlib inline
pl.rcParams['figure.figsize'] = (15,5)
pl.rcParams['lines.linewidth'] = 3
data = px4tools.project_lat_lon(px4tools.process_data(pandas.read_csv('logs/15-10-24-16_46_42-jgoppert.csv')))
px4tools.alt_analysis(data);
px4tools.pos_analysis(data[data.STAT_MainState == 3]);
px4tools.pos_analysis(data[data.STAT_MainState == 2]);
data.EST2_P0.plot()
data.EST2_P1.plot()
data.EST2_P2.plot()
data.EST2_P3.plot()
data.EST2_P4.plot()
data.EST2_P5.plot()
pl.legend()
data1 = data
data1.EST0_s6.plot(label='b_x')
data1.EST0_s7.plot(label='b_y')
data1.EST0_s8.plot(label='b_z')
pl.legend(loc='best', ncol=3)
pl.title('bias estimates')
pl.grid()
data1.EST2_P6.plot(label='b_x')
data1.EST2_P7.plot(label='b_y')
data1.EST2_P8.plot(label='b_z')
pl.grid()
pl.title('bias variances')
pl.legend(loc='best', ncol=3)
px4tools.pos_analysis(data);
px4tools.plot_velocity_loops(data[data.STAT_MainState==3])
px4tools.plot_position_loops(data[data.STAT_MainState==2])
px4tools.plot_attitude_rate_loops(data)
px4tools.plot_attitude_loops(data)
# +
data1 = data[74:92]
#pl.quiver()
gps_x = px4tools.new_sample(data1.GPS_X)
gps_y = px4tools.new_sample(data1.GPS_Y)
p_x = px4tools.new_sample(data1.LPOS_X)
p_y = px4tools.new_sample(data1.LPOS_Y)
pl.plot(p_y, p_x, label='trajectory', alpha=0.5)
pl.plot(gps_y, gps_x, 'x', label='GPS')
pl.quiver(p_y, p_x, gps_y - p_y, gps_x - p_x, scale_units='xy', scale=1, units='xy', angles='xy', label='$\Delta$')
pl.legend()
pl.grid()
#pl.legend(loc='best')
# -
data1 = []
f_gps_delay = lambda dt, t1, t2, data: pl.norm(data.GPS_X[t1:t2] - data.LPOS_X[t1-dt: t2-dt])
f_gps_delay(0, 70, 80, data)
pl.plot(data.LPOS_Dist.shift(100));
pl.plot(data.LPOS_Dist.shift(100));
def f_delay(series1, series2, dt):
d = series2.shift(dt)
pl.interp(pl.array(series1.index, dtype=float), pl.array(series1.index, dtype=float) + 0.1,
pl.array(series1, dtype=float))
dx = series1.shift(dt) - series2.shift(0)
dx_data = pl.array(dx[pl.isfinite(dx)])
return pl.norm(dx_data)
pl.interp(pl.array(data.index, dtype=float), pl.array(data.index, dtype=float) + 0.1,
pl.array(data.LPOS_X, dtype=float))
f_delay(data.LPOS_X, data.GPS_X, 1)
import scipy.optimize
scipy.optimize.fmin(lambda dt: f_delay(data.LPOS_X, data.GPS_X, dt), 0);
pl.plot(dx[pl.isfinite(dx)])
pl.plot(data.EST0_s0)
pl.plot(data.EST0_s1)
pl.plot(data.EST0_s2)
pl.grid()
| 15-10-24.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../../taxonomy-enrichment/baselines/ruwordnet')
sys.path.append('../../taxonomy-enrichment/baselines')
import re
import numpy as np
import pandas as pd
import fasttext
from ruwordnet_reader import RuWordnet
from sklearn.metrics.pairwise import cosine_similarity
from tqdm import tqdm
import xml.etree.ElementTree as ET
ruwordnet = RuWordnet(db_path="../../dialogue2020_shared_task_hypernyms/dataset/ruwordnet.db", ruwordnet_path=None)
public_test = []
with open('../../dialogue2020_shared_task_hypernyms/dataset/public/verbs_public_no_labels.tsv', 'r', encoding="utf-8") as f:
for line in f:
line = line.rstrip()
public_test.append(line)
private_test = []
with open('../../dialogue2020_shared_task_hypernyms/dataset/private/verbs_private_no_labels.tsv', 'r', encoding="utf-8") as f:
for line in f:
line = line.rstrip()
private_test.append(line)
public_test[:3], private_test[:3]
nouns = {}
nouns_list = []
for sense_id, synset_id, text in ruwordnet.get_all_senses():
if synset_id.endswith("V"):
ltext = text.lower()
if ltext not in nouns:
nouns_list.append(ltext)
nouns.setdefault(ltext, []).append(synset_id)
len(nouns), len(nouns_list)
synset2words = {}
for sense_id, synset_id, text in ruwordnet.get_all_senses():
if synset_id.endswith("V"):
synset2words.setdefault(synset_id, []).append(text.lower())
len(synset2words)
list(nouns.items())[:3]
df_test = pd.DataFrame(data={'word': public_test + private_test})
df_test.shape
df_test['private'] = [1 if x in private_test else 0 for x in df_test['word']]
df_test['private'].value_counts()
df_test.head()
wiktionarydump = "ruwiktionary-20200120-pages-articles-multistream.xml"
title2doc = {}
doc = {}
fields = {
"timestamp": "timestamp",
"title": "title",
"text": "text",
"redirect title": "redirect_title",
}
cnt = 0
for _, elem in tqdm(ET.iterparse(wiktionarydump, events=("end",))):
prefix, has_namespace, postfix = elem.tag.partition('}')
tag = postfix if postfix else prefix
if tag in fields:
doc[fields[tag]] = elem.text
if tag == "page":
elem.clear()
cnt += 1
title2doc[doc["title"]] = doc
doc = {}
# longest article by lowercased word
ltitle2doc = {}
for x in title2doc.keys():
if x.lower() in ltitle2doc:
if len(title2doc[x]['text']) > len(ltitle2doc[x.lower()]['text']):
ltitle2doc[x.lower()] = title2doc[x]
else:
ltitle2doc[x.lower()] = title2doc[x]
ltitle_list = list(ltitle2doc.keys())
# longest article by lowercased word
ltitle2docs = {}
for x in title2doc.keys():
ltitle2docs.setdefault(x.lower(), []).append(title2doc[x])
ltitle_list = list(ltitle2docs.keys())
df_test['wikt_in'] = [1 if x.lower() in ltitle2doc else 0 for x in df_test['word']]
print(df_test[df_test['private']==0]['wikt_in'].value_counts())
print(df_test[df_test['private']==1]['wikt_in'].value_counts())
ftmodel = fasttext.load_model("../../dialogue2020_shared_task_hypernyms/baselines/models/cc.ru.300.bin")
ftwords_list = ftmodel.get_words()
ftwords = set(ftwords_list)
lword2word = {word.lower(): word for word in ftwords_list}
len(lword2word), len(ftwords_list)
df_test['ft_in'] = [1 if x.lower() in lword2word else 0 for x in df_test['word']]
print(df_test[df_test['private']==0]['ft_in'].value_counts())
print(df_test[df_test['private']==1]['ft_in'].value_counts())
nouns_vectors = np.zeros((len(nouns_list), ftmodel.get_dimension()))
for i, word in enumerate(tqdm(nouns_list)):
nouns_vectors[i] = ftmodel.get_sentence_vector(word)
ltitle_vectors = np.zeros((len(ltitle_list), ftmodel.get_dimension()))
for i, word in enumerate(tqdm(ltitle_list)):
ltitle_vectors[i] = ftmodel.get_sentence_vector(word)
def get_top_k_similar(vectors, vector, k=1):
res = []
dots = np.dot(vectors, vector)
for i in range(k):
idx = np.argmax(dots)
res.append(idx)
dots[idx] = 0
return res
i = 1
lword = public_test[i].lower()
idxs = get_top_k_similar(ltitle_vectors, ftmodel.get_sentence_vector(lword), k=5)
for idx in idxs:
print(lword, ltitle_list[idx])
df_test['wn_top10'] = [
[nouns_list[x] for x in get_top_k_similar(nouns_vectors, ftmodel.get_sentence_vector(word.lower()), k=10)]
for word in tqdm(df_test['word'])
]
df_train = pd.DataFrame(data={'word': [x[0].upper() for x in df_test['wn_top10']]})
# df_train = pd.DataFrame(data={'word': []})
df_train.shape
# skip self
df_train['wn_top10'] = [
[nouns_list[x] for x in get_top_k_similar(nouns_vectors, ftmodel.get_sentence_vector(word.lower()), k=11) if nouns_list[x] != word.lower()]
for word in tqdm(df_train['word'])
]
for df in [
df_test,
df_train
]:
df['wikt_top10'] = [
[ltitle_list[x] for x in get_top_k_similar(ltitle_vectors, ftmodel.get_sentence_vector(word.lower()), k=10)]
for word in tqdm(df['word'])
]
# +
def clean_markup(text):
return text.replace("[[", "").replace("]]", "").replace("{{aslinks|", "")
def parse_item(text):
items = []
if text.startswith("# ") and len(line) > 2:
items.extend([
clean_markup(x).replace("?", "").replace(";", "").replace("'", "").strip()
for x in re.split(',|;', text[2:]) if x not in {'-', '?', '—', ''}
])
return items
def parse_translation(trans):
res = {}
for line in trans.split('\n'):
if line.startswith('|'):
l, r = line.split('=')
res[l[1:]] = r.replace('[[', '').replace(']]', '')
return res
def parse_wiktionary(text):
res = {'hypernym': [], 'synonym': [], 'meaning': []}
h1 = ""
texts = []
for line in text.split("\n"):
if line.startswith("= ") and line.endswith(" ="):
h1 = line
if h1 == '= {{-ru-}} =':
texts.append(line)
text = "\n".join(texts)
for par in text.split("\n\n"):
for h, f in [('==== Гиперонимы ====', 'hypernym'), ('==== Синонимы ====', 'synonym')]:
if h in par:
res[f] = [w for line in par.split("\n") for w in parse_item(line)]
for h, f in [('==== Значение ====', 'meaning')]:
if h in par:
res[f] = [clean_markup(line[2:]) for line in par.split("\n") if line.startswith('# ') and len(line) > 2]
if '=== Перевод ===' in par:
res['translation'] = par.replace('=== Перевод ===\n', '')
return res
# -
for df in [df_test, df_train]:
df['wikt_hypernyms_text'] = [
parse_wiktionary(ltitle2doc[word.lower()]['text'])['hypernym'] if word.lower() in ltitle2doc else []
for word in tqdm(df['word'])
]
for df in [df_test, df_train]:
df['wikt_top1_hypernyms_text'] = [
parse_wiktionary(ltitle2doc[words[0].lower()]['text'])['hypernym']
for words in tqdm(df['wikt_top10'])
]
sum([len(x) for x in df_test['wikt_top1_hypernyms_text']]), sum([len(x) for x in df_train['wikt_top1_hypernyms_text']])
for df in [df_test, df_train]:
res = []
for words in tqdm(df['wikt_top10']):
res_el = []
for doc in ltitle2docs[words[0].lower()]:
res_el.extend(parse_wiktionary(doc['text'])['hypernym'])
res.append(res_el)
df['wikt_top1_hypernyms_text_docs'] = res
sum([len(x) for x in df_test['wikt_top1_hypernyms_text_docs']]), sum([len(x) for x in df_train['wikt_top1_hypernyms_text_docs']])
for df in [df_test, df_train]:
for i in range(10):
df['wn_top%d_hypernyms' % (i + 1)] = [
[hyp for synset_id in nouns[words[i]] for hyp in ruwordnet.get_hypernyms_by_id(synset_id)]
for words in tqdm(df['wn_top10'])
]
import os
from collections import Counter
import pymorphy2
from nltk.tokenize import word_tokenize
morph = pymorphy2.MorphAnalyzer()
for df in [df_test, df_train]:
cnt = 0
lens = []
wikt_top1_hypernyms = []
for word, hypernyms_text in zip(df['word'], df['wikt_top1_hypernyms_text_docs']):
res = []
for hypernym in hypernyms_text:
lhypernym = hypernym.lower().replace('ё', 'е')
if lhypernym in nouns:
res.extend(sorted(nouns[lhypernym]))
else:
parsed = morph.parse(lhypernym)
if 'plur' in parsed[0].tag and parsed[0].normal_form in nouns:
res.extend(sorted(nouns[parsed[0].normal_form]))
cnt += 1
lens.append(len(res))
wikt_top1_hypernyms.append(res)
df['wikt_top1_hypernyms_docs'] = wikt_top1_hypernyms
print(cnt)
df_test.head(6)
df_train.head(6)
# +
import codecs
def save_to_file(words_with_hypernyms, output_path, ruwordnet):
with codecs.open(output_path, 'w', encoding='utf-8') as f:
for word, hypernyms in words_with_hypernyms.items():
for hypernym in hypernyms:
f.write(f"{word}\t{hypernym}\t{ruwordnet.get_name_by_id(hypernym)}\n")
# -
def get_top_hypernyms(l, sz=10):
res_set = set()
res = []
for el in sorted(l):
if el[1] not in res_set:
res.append(el[1])
res_set.add(el[1])
return res[:sz]
from itertools import chain
# +
# in df_train some items added to hypernyms multiple times
features = {word: {} for word in chain(df_test['word'], df_train['word'])}
hypernyms = {word: [] for word in chain(df_test['word'], df_train['word'])}
syn_priority_l1 = 4.
syn_priority_l2 = 2.
syn_priority_l3 = 3.
syntail_priority_l1 = 7.
syntail_priority_l2 = 5.
syntail_priority_l3 = 6.
wikhyp_priority2_l1 = 0.
wikhyp_priority2_l2 = 1.
wikhyp_priority3_l1 = 5.
wikhyp_priority3_l2 = 6.
for df in [df_test, df_train]:
for word, hs in zip(df['word'], df['wikt_top1_hypernyms_docs']):
for j, hypernym in enumerate(hs):
features[word].setdefault(hypernym, {})['wikhyp_priority_l1'] = 1
features[word].setdefault(hypernym, {})['wikhyp_priority_l1_pos'] = j
for hyphyp in ruwordnet.get_hypernyms_by_id(hypernym):
features[word].setdefault(hyphyp, {})['wikhyp_priority_l2'] = 1
features[word].setdefault(hyphyp, {})['wikhyp_priority_l2_pos'] = j
for j, hypernym in enumerate(hs[:2]):
hypernyms[word].append((wikhyp_priority2_l1 + j*1e-3, hypernym))
for hyphyp in ruwordnet.get_hypernyms_by_id(hypernym):
hypernyms[word].append((wikhyp_priority2_l2 + j*1e-3, hyphyp))
for j, hypernym in enumerate(hs[2:]):
hypernyms[word].append((wikhyp_priority3_l1 + j*1e-3, hypernym))
for hyphyp in ruwordnet.get_hypernyms_by_id(hypernym):
hypernyms[word].append((wikhyp_priority3_l2 + j*1e-3, hyphyp))
for i in range(2, 11):
for word, hs in zip(df['word'], df['wn_top%d_hypernyms' % i]):
for j, hypernym in enumerate(hs):
features[word].setdefault(hypernym, {})['syn%d_priority_l2'%i] = 1
features[word].setdefault(hypernym, {})['syn%d_priority_l2_pos'%i] = j
hypernyms[word].append((syntail_priority_l2 + (i-2)*1e-3, hypernym))
for hyphyp in ruwordnet.get_hypernyms_by_id(hypernym):
features[word].setdefault(hyphyp, {})['syn%d_priority_l3'%i] = 1
features[word].setdefault(hyphyp, {})['syn%d_priority_l3_pos'%i] = j
hypernyms[word].append((syntail_priority_l3 + (i-2)*1e-3, hyphyp))
for word, hs in zip(df['word'], df['wn_top1_hypernyms']):
for j, hypernym in enumerate(hs):
hypernyms[word].append((syn_priority_l2, hypernym))
features[word].setdefault(hypernym, {})['syn1_priority_l2'] = 1
features[word].setdefault(hypernym, {})['syn1_priority_l2_pos'] = j
for hyphyp in ruwordnet.get_hypernyms_by_id(hypernym):
features[word].setdefault(hyphyp, {})['syn1_priority_l3'] = 1
features[word].setdefault(hyphyp, {})['syn1_priority_l3_pos'] = j
hypernyms[word].append((syn_priority_l3, hyphyp))
for word, words in zip(df['word'], df['wn_top10']):
for synset_id in nouns[words[0]]:
hypernyms[word].append((syn_priority_l1, synset_id))
features[word].setdefault(synset_id, {})['syn1_priority_l1'] = 1
features[word].setdefault(synset_id, {})['syn1_priority_l1_pos'] = 0
for i, word2 in enumerate(words[1:]):
for j, synset_id in enumerate(nouns[word2]):
hypernyms[word].append((syntail_priority_l1 + i*1e-3, synset_id))
features[word].setdefault(synset_id, {})['syn%d_priority_l1'%(i+2)] = 1
features[word].setdefault(synset_id, {})['syn%d_priority_l1_pos'%(i+2)] = j
# -
from nltk.corpus import wordnet as wn
try:
wn.all_synsets
except LookupError as e:
import nltk
nltk.download('wordnet')
def drop_trailing_dot(s):
if s.endswith('.'):
return s[:-1]
return s
ru2en = {}
with open('data/ru.txt', 'r', encoding='utf-8') as f_ru, open('data/en_ya.txt', 'r', encoding='utf-8') as f_en_y:
for i, r, ey in zip(range(100500), f_ru, f_en_y):
r = drop_trailing_dot(r.strip())
ey = drop_trailing_dot(ey.strip())
ru2en[r] = ey
len(ru2en)
en2ru = {}
with open('data/hyp_en.txt', 'r', encoding="utf-8") as f_en, open('data/hyp_ru_ya.txt', 'r', encoding="utf-8") as f_ru_y:
for i, e, ruy, in zip(range(100500), f_en, f_ru_y):
e = drop_trailing_dot(e.strip())
ruy = drop_trailing_dot(ruy.strip())
en2ru[e] = ruy
len(en2ru)
# +
missing = set()
hypernyms_en = {}
hypernyms_en_txt = {}
for df in [df_test, df_train]:
cnt = 0
for word in df["word"]:
hypernyms_en[word] = set()
hypernyms_en_txt[word] = set()
lword = word.lower()
if lword in ru2en:
synsets = wn.synsets(ru2en[lword])
if synsets:
flag = False
for sense in synsets:
for hyp in sense.hypernyms():
for name in hyp.lemma_names():
name = name.replace('_', ' ')
if name in en2ru:
if en2ru[name].lower() in nouns:
flag = True
hypernyms_en_txt[word].add(en2ru[name].lower())
for id_ in nouns[en2ru[name].lower()]:
hypernyms[word].append((0.0, id_))
hypernyms_en[word].add((0.0, id_))
else:
missing.add(name)
if hypernyms_en[word]:
cnt += 1
print(cnt / (len(df["word"])+1e-6))
# -
for word in public_test[:2] + private_test[:2]:
print(word, hypernyms_en_txt[word], hypernyms_en[word])
for word in hypernyms_en:
for score, hypernym in hypernyms_en[word]:
features[word].setdefault(hypernym, {})['wordnet_en_l1'] = 1
for hyphyp in ruwordnet.get_hypernyms_by_id(hypernym):
features[word].setdefault(hyphyp, {})['wordnet_en_l2'] = 1
norm_words = {}
def normalize(s):
res = []
for word in word_tokenize(s.lower()):
if word in norm_words:
res.append(norm_words[word])
else:
mp = morph.parse(word)
if mp:
norm_words[word] = mp[0].normal_form
res.append(norm_words[word])
return " ".join(res)
# +
def innertext(tag):
return (tag.text or '') + ''.join(innertext(e) for e in tag) + (tag.tail or '')
def get_serp_texts(xml_path, k=5):
res = []
if os.path.exists(xml_path):
root = ET.parse(xml_path).getroot()
for e in root.find('response').find('results').find('grouping').findall('group')[:k]:
res.append(innertext(e.find('doc').find('title')))
if e.find('doc').find('passages'):
for passage in e.find('doc').find('passages'):
res.append(innertext(passage))
if e.find('doc').find('headline'):
res.append(innertext(e.find('doc').find('headline')))
return " ".join(res)
# +
synset_norm_serp_ya_cnt = Counter()
synset_norm_serp_g_cnt = Counter()
hypernyms_wserp = {}
serp_priority = -4.
serp_hyp_priority = -4.
meaning_priority = -1.
for df in [
df_test,
df_train
]:
for word in tqdm(df["word"]):
word_file_path = 'data/google_it_all/' + word.lower() + '.tsv'
total_g_serp = ""
if os.path.exists(word_file_path):
with open(word_file_path, 'r', encoding='utf-8') as f:
for line in f:
text = line.split('\t')[1]
total_g_serp += text + " "
norm_total_g_serp = normalize(total_g_serp)
word_file_path = 'data/yandex_it_all/' + word.upper() + '.xml'
total_ya_serp = get_serp_texts(word_file_path, k=10)
norm_total_ya_serp = normalize(total_ya_serp)
total_meaning = ""
if word.lower() in ltitle2doc:
for meaning in parse_wiktionary(ltitle2doc[word.lower()]['text'])['meaning']:
total_meaning += meaning + " "
norm_total_meaning = normalize(total_meaning)
res = []
for score, hypernym in hypernyms[word]:
hypernym_texts = synset2words[hypernym] + [ruwordnet.get_name_by_id(hypernym)]
for hypernym_text in hypernym_texts:
norm_hypernym_text = normalize(hypernym_text)
if norm_hypernym_text in norm_total_g_serp:
score += serp_priority
features[word][hypernym]['serp_g_norm'] = 1
for hyphyp in ruwordnet.get_hypernyms_by_id(hypernym):
if hyphyp in features[word]:
features[word][hyphyp]['serp_g_norm_l2'] = 1
synset_norm_serp_g_cnt[hypernym] += 1
if hypernym_text in total_g_serp:
features[word][hypernym]['serp_g'] = 1
if norm_hypernym_text in norm_total_ya_serp:
features[word][hypernym]['serp_ya_norm'] = 1
synset_norm_serp_ya_cnt[hypernym] += 1
for hyphyp in ruwordnet.get_hypernyms_by_id(hypernym):
if hyphyp in features[word]:
features[word][hyphyp]['serp_ya_norm_l2'] = 1
if hypernym_text in total_ya_serp:
features[word][hypernym]['serp_ya'] = 1
if norm_hypernym_text in norm_total_meaning:
score += meaning_priority
features[word][hypernym]['meaning_norm'] = 1
if hypernym_text in total_meaning:
features[word][hypernym]['meaning'] = 1
res.append((score, hypernym))
hypernyms_wserp[word] = res
# -
# feature_names = set()
# for word in features:
# for synset_id in features[word]:
# for key in features[word][synset_id]:
# feature_names.add(key)
# feature_names = sorted(feature_names)
# len(feature_names)
feature_names = [
"meaning",
"meaning_norm",
# "serp_g",
# "serp_g_norm",
# "serp_g_norm_l2",
# "serp_ya",
# "serp_ya_norm",
# "serp_ya_norm_l2",
"syn1_priority_l1",
"syn1_priority_l2",
"syn1_priority_l3",
"syn2_priority_l1",
"syn2_priority_l2",
"syn2_priority_l3",
"syn3_priority_l1",
"syn3_priority_l2",
"syn3_priority_l3",
"syn4_priority_l1",
"syn4_priority_l2",
"syn4_priority_l3",
"syn5_priority_l1",
"syn5_priority_l2",
"syn5_priority_l3",
"syn6_priority_l1",
"syn6_priority_l2",
"syn6_priority_l3",
"syn7_priority_l1",
"syn7_priority_l2",
"syn7_priority_l3",
"wikhyp_priority_l1",
"wikhyp_priority_l2",
"wordnet_en_l1",
"wordnet_en_l2",
]
total = sum([len(features[x]) for x in df_train["word"]])
X = np.zeros( (total, len(feature_names)) )
y = np.zeros( total )
X.shape, y.shape
pos = 0
for word in df_train["word"]:
lword = word.lower()
true_hypernyms = set()
for synset_id in nouns[lword]:
true_hypernyms.update(ruwordnet.get_hypernyms_by_id(synset_id))
for synset_id in features[word]:
y[pos] = 1 if synset_id in true_hypernyms else 0
X[pos] = [features[word][synset_id].get(fn, 0) for fn in feature_names]
pos += 1
pos
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
y_train.shape, y_test.shape
for C in [0.001, 0.003, 0.01, 0.03, 0.1, 0.3, 1.0, 3, 10, 30, 100, 300, 1000]:
model = LogisticRegression(C=C)
model.fit(X_train, y_train)
print(C, roc_auc_score(y_test, model.predict_proba(X_test)[:, 1]))
model = LogisticRegression(C=0.03)
model.fit(X, y)
weights = {k:v for k,v in zip(feature_names, model.coef_[0])}
# +
pos_weights = {}
for i in range(1, 11):
for j in range(1,4):
pos_weights['syn%d_priority_l%d_pos' % (i, j)] = -0.0001
def calc_score(d):
score = 0.
for feature, weight in weights.items():
score += weight * d.get(feature, 0)
for feature, weight in pos_weights.items():
score += weight * d.get(feature, 0)
return score
# -
prefix = 'subm/subm106_no_gya'
save_to_file({
k: get_top_hypernyms( [(-calc_score(features[k][x]), x) for x in features[k]] )
for k in public_test
}, prefix + '_public_verbs.tsv', ruwordnet)
save_to_file({
k: get_top_hypernyms( [(-calc_score(features[k][x]), x) for x in features[k]] )
for k in private_test
}, prefix + '_private_verbs.tsv', ruwordnet)
weights = {
"meaning": 0.15,
"meaning_norm": 0.15,
"serp_g": 0.20,
"serp_g_norm": 0.50,
"serp_g_norm_l2": 0.20,
"serp_ya": 0.15,
"serp_ya_norm": 0.20,
"serp_ya_norm_l2": 0.20,
"syn1_priority_l1": 0.05,
"syn1_priority_l2": 0.50,
"syn1_priority_l3": 0.30,
"syn2_priority_l1": 0.05,
"syn2_priority_l2": 0.25,
"syn2_priority_l3": 0.15,
"syn3_priority_l1": 0.05,
"syn3_priority_l2": 0.25,
"syn3_priority_l3": 0.15,
"syn4_priority_l1": 0.00,
"syn4_priority_l2": 0.10,
"syn4_priority_l3": 0.15,
"syn5_priority_l1": 0.00,
"syn5_priority_l2": 0.10,
"syn5_priority_l3": 0.00,
"syn6_priority_l1": 0.00,
"syn6_priority_l2": 0.10,
"syn6_priority_l3": 0.00,
"syn7_priority_l1": 0.00,
"syn7_priority_l2": 0.10,
"syn7_priority_l3": 0.00,
"wikhyp_priority_l1": 0.20,
"wikhyp_priority_l2": 0.50,
"wordnet_en_l1": 0.05,
"wordnet_en_l2": 0.20,
}
prefix = 'subm/subm107_no_gya'
save_to_file({
k: get_top_hypernyms( [(-calc_score(features[k][x]), x) for x in features[k]] )
for k in public_test
}, prefix + '_public_verbs.tsv', ruwordnet)
save_to_file({
k: get_top_hypernyms( [(-calc_score(features[k][x]), x) for x in features[k]] )
for k in private_test
}, prefix + '_private_verbs.tsv', ruwordnet)
| code/yuriy/taxonomy_features-ru.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Errors and Exceptions
# Every programmer encounters errors, both those who are just beginning, and those
# who have been programming for years. Encountering errors and exceptions can be
# very frustrating at times, and can make coding feel like a hopeless endeavour.
# However, understanding what the different types of errors are and when you are
# likely to encounter them can help a lot. Once you know *why* you get certain
# types of errors, they become much easier to fix.
#
# Errors in Python have a very specific form, called a traceback. Let's examine
# one:
def favourite_ice_cream():
ice_creams = ["chocolate","vanilla","strawberry"]
print(ice_creams[2])
favourite_ice_cream()
# This particular traceback has two levels.
# You can determine the number of levels by looking for the number of arrows on the left hand side.
# In this case:
# 1. The first shows code from the cell above,
# with an arrow pointing to Line 8 (which is `favorite_ice_cream()`).
#
# 2. The second shows some code in the function `favorite_ice_cream`,
# with an arrow pointing to Line 6 (which is `print(ice_creams[3])`).
#
# The last level is the actual place where the error occurred.
# The other level(s) show what function the program executed to get to the next level down.
# So, in this case, the program first performed a function call to the function `favorite_ice_cream`.
# Inside this function,
# the program encountered an error on Line 6, when it tried to run the code `print(ice_creams[3])`.
#
# <section class="callout panel panel-warning">
# <div class="panel-heading">
# <h2><span class="fa fa-thumb-tack"></span> Long Tracebacks</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p>Sometimes, you might see a traceback that is very long -- sometimes they might even be 20 levels deep!
# This can make it seem like something horrible happened,
# but really it just means that your program called many functions before it ran into the error.
# Most of the time,
# you can just pay attention to the bottom-most level,
# which is the actual place where the error occurred.</p>
#
# </div>
#
# </section>
#
# So what error did the program actually encounter?
#
# In the last line of the traceback, Python helpfully tells us the category or
# type of error (in this case, it is an `IndexError`) and a more detailed error
# message (in this case, it says "list index out of range"). If you encounter an
# error and don't know what it means, it is still important to read the traceback
# closely. That way, if you fix the error, but encounter a new one, you can tell
# that the error changed. Additionally, sometimes just knowing *where* the error
# occurred is enough to fix it, even if you don't entirely understand the message.
#
# If you do encounter an error you don't recognize, try looking at the [official
# documentation on errors](http://docs.python.org/3/library/exceptions.html).
# However, note that you may not always be able to find the error there, as it is
# possible to create custom errors. In that case, hopefully the custom error
# message is informative enough to help you figure out what went wrong.
# ## Syntax Errors
#
# When you forget a colon at the end of a line,
# accidentally add one space too many when indenting under an `if` statement,
# or forget a parenthesis,
# you will encounter a syntax error.
# This means that Python couldn't figure out how to read your program.
# This is similar to forgetting punctuation in English:
# for example,
# this text is difficult to read there is no punctuation there is also no capitalization
# why is this hard because you have to figure out where each sentence ends
# you also have to figure out where each sentence begins
# to some extent it might be ambiguous if there should be a sentence break or not
#
# People can typically figure out what is meant by text with no punctuation,
# but people are much smarter than computers.
# If Python doesn't know how to read the program,
# it will just give up and inform you with an error.
# For example:
def some_function():
print("hello")
# Here, Python tells us that there is a `SyntaxError` on line 1,
# and even puts a little arrow in the place where there is an issue.
# In this case the problem is that the function definition is missing a colon at the end.
#
# Actually, the function above has *two* issues with syntax.
# If we fix the problem with the colon,
# we see that there is *also* an `IndentationError`,
# which means that the lines in the function definition do not all have the same indentation:
def some_function():
a = "hello"
return a
# Both `SyntaxError` and `IndentationError` indicate a problem with the syntax of your program,
# but an `IndentationError` is more specific:
# it *always* means that there is a problem with how your code is indented.
#
# <section class="callout panel panel-warning">
# <div class="panel-heading">
# <h2><span class="fa fa-thumb-tack"></span> Tabs and Spaces</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p>Some indentation errors are harder to spot than others.
# In particular, mixing spaces and tabs can be difficult to spot
# because they are both whitespace.
# In the example below, the first two lines in the body of the function
# <code>some_function</code> are indented with tabs, while the third line — with spaces.
# If you're working in a Jupyter notebook, be sure to copy and paste this example
# rather than trying to type it in manually because Jupyter automatically replaces
# tabs with spaces.</p>
# <div class="codehilite"><pre><span></span><span class="k">def</span> <span class="nf">some_function</span><span class="p">():</span>
# <span class="n">msg</span> <span class="o">=</span> <span class="s2">"hello, world!"</span>
# <span class="k">print</span><span class="p">(</span><span class="n">msg</span><span class="p">)</span>
# <span class="k">return</span> <span class="n">msg</span>
# </pre></div>
#
#
# <p>Visually it is impossible to spot the error.
# Fortunately, Python does not allow you to mix tabs and spaces.</p>
# <div class="codehilite"><pre><span></span> File "<ipython-input-5-653b36fbcd41>", line 4
# return msg
# ^
# TabError: inconsistent use of tabs and spaces in indentation
# </pre></div>
#
# </div>
#
# </section>
#
# ## Variable Name Errors
#
# Another very common type of error is called a `NameError`,
# and occurs when you try to use a variable that does not exist.
# For example:
print(b)
# Variable name errors come with some of the most informative error messages,
# which are usually of the form "name 'the_variable_name' is not defined".
#
# Why does this error message occur?
# That's a harder question to answer,
# because it depends on what your code is supposed to do.
# However,
# there are a few very common reasons why you might have an undefined variable.
# The first is that you meant to use a string, but forgot to put quotes around it:
# The second is that you just forgot to create the variable before using it.
# In the following example,
# `count` should have been defined (e.g., with `count = 0`) before the for loop:
for number in range(10):
count = count + number
print(count)
# Finally, the third possibility is that you made a typo when you were writing your code.
# Let's say we fixed the error above by adding the line `Count = 0` before the for loop.
# Frustratingly, this actually does not fix the error.
# Remember that variables are case-sensitive,
# so the variable `count` is different from `Count`. We still get the same error, because we still have not defined `count`:
del count
count = 0
for number in range(10):
count = count + number
print(count)
# ## Index Errors
#
# Next up are errors having to do with containers (like lists and strings) and the items within them.
# If you try to access an item in a list or a string that does not exist,
# then you will get an error.
# This makes sense:
# if you asked someone what day they would like to get coffee,
# and they answered "caturday",
# you might be a bit annoyed.
# Python gets similarly annoyed if you try to ask it for an item that doesn't exist:
letters = ["a", "b", 3]
letters[1]
letters[4]
# Here, Python is telling us that there is an `IndexError` in our code, meaning we
# tried to access a list index that did not exist.
# ## File Errors
#
# The last type of error we'll cover today
# are those associated with reading and writing files: `FileNotFoundError`.
# If you try to read a file that does not exist,
# you will receive a `FileNotFoundError` telling you so.
# If you attempt to write to a file that was opened read-only, Python 3
# returns an `UnsupportedOperationError`.
# More generally, problems with input and output manifest as
# `IOError`s or `OSError`s, depending on the version of Python you use.
file_handle = open("nonexistent.txt", "r")
file_handle.close()
# One reason for receiving this error is that you specified an incorrect path to the file.
# For example,
# if I am currently in a folder called `myproject`,
# and I have a file in `myproject/writing/myfile.txt`,
# but I try to just open `myfile.txt`,
# this will fail.
# The correct path would be `writing/myfile.txt`.
# It is also possible (like with `NameError`) that you just made a typo.
#
# A related issue can occur if you use the "read" flag instead of the "write" flag.
# Python will not give you an error if you try to open a file for writing when the file does not exist.
# However,
# if you meant to open a file for reading,
# but accidentally opened it for writing,
# and then try to read from it,
# you will get an `UnsupportedOperation` error
# telling you that the file was not opened for reading:
file_handle - open("nonexistent.txt", "w")
file_handle.read()
# These are the most common errors with files,
# though many others exist.
# If you get an error that you've never seen before,
# searching the Internet for that error type
# often reveals common reasons why you might get that error.
#
# <section class="challenge panel panel-success">
# <div class="panel-heading">
# <h2><span class="fa fa-pencil"></span> Reading Error Messages</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p>Read the python code and the resulting traceback below, and answer the following questions:</p>
# <ol>
# <li>How many levels does the traceback have?</li>
# <li>What is the function name where the error occurred?</li>
# <li>On which line number in this function did the error occurr?</li>
# <li>What is the type of error?</li>
# <li>What is the error message?</li>
# </ol>
# <div class="codehilite"><pre><span></span><span class="c1"># This code has an intentional error. Do not type it directly;</span>
# <span class="c1"># use it for reference to understand the error message below.</span>
# <span class="k">def</span> <span class="nf">print_message</span><span class="p">(</span><span class="n">day</span><span class="p">):</span>
# <span class="n">messages</span> <span class="o">=</span> <span class="p">{</span>
# <span class="s2">"monday"</span><span class="p">:</span> <span class="s2">"Hello, world!"</span><span class="p">,</span>
# <span class="s2">"tuesday"</span><span class="p">:</span> <span class="s2">"Today is tuesday!"</span><span class="p">,</span>
# <span class="s2">"wednesday"</span><span class="p">:</span> <span class="s2">"It is the middle of the week."</span><span class="p">,</span>
# <span class="s2">"thursday"</span><span class="p">:</span> <span class="s2">"Today is Donnerstag in German!"</span><span class="p">,</span>
# <span class="s2">"friday"</span><span class="p">:</span> <span class="s2">"Last day of the week!"</span><span class="p">,</span>
# <span class="s2">"saturday"</span><span class="p">:</span> <span class="s2">"Hooray for the weekend!"</span><span class="p">,</span>
# <span class="s2">"sunday"</span><span class="p">:</span> <span class="s2">"Aw, the weekend is almost over."</span>
# <span class="p">}</span>
# <span class="k">print</span><span class="p">(</span><span class="n">messages</span><span class="p">[</span><span class="n">day</span><span class="p">])</span>
#
# <span class="k">def</span> <span class="nf">print_friday_message</span><span class="p">():</span>
# <span class="n">print_message</span><span class="p">(</span><span class="s2">"Friday"</span><span class="p">)</span>
#
# <span class="n">print_friday_message</span><span class="p">()</span>
# </pre></div>
#
#
# <div class="codehilite"><pre><span></span>---------------------------------------------------------------------------
# KeyError Traceback (most recent call last)
# <ipython-input-1-4be1945adbe2> in <module>()
# 14 print_message("Friday")
# 15
# ---> 16 print_friday_message()
#
# <ipython-input-1-4be1945adbe2> in print_friday_message()
# 12
# 13 def print_friday_message():
# ---> 14 print_message("Friday")
# 15
# 16 print_friday_message()
#
# <ipython-input-1-4be1945adbe2> in print_message(day)
# 9 "sunday": "Aw, the weekend is almost over."
# 10 }
# ---> 11 print(messages[day])
# 12
# 13 def print_friday_message():
#
# KeyError: 'Friday'
# </pre></div>
#
# </div>
#
# </section>
#
#
# <section class="solution panel panel-primary">
# <div class="panel-heading">
# <h2><span class="fa fa-eye"></span> Solution</h2>
# </div>
#
#
# <div class="panel-body">
#
# <ol>
# <li>3 levels</li>
# <li><code>print_message</code></li>
# <li>11</li>
# <li><code>KeyError</code></li>
# <li>There isn't really a message; you're supposed to infer that <code>Friday</code> is not a key in <code>messages</code>.</li>
# </ol>
#
# </div>
#
# </section>
#
#
# <section class="challenge panel panel-success">
# <div class="panel-heading">
# <h2><span class="fa fa-pencil"></span> Identifying Syntax Errors</h2>
# </div>
#
#
# <div class="panel-body">
#
# <ol>
# <li>Read the code below, and (without running it) try to identify what the errors are.</li>
# <li>Run the code, and read the error message. Is it a <code>SyntaxError</code> or an <code>IndentationError</code>?</li>
# <li>Fix the error.</li>
# <li>Repeat steps 2 and 3, until you have fixed all the errors.</li>
# </ol>
# <div class="codehilite"><pre><span></span><span class="k">def</span> <span class="nf">another_function</span>
# <span class="k">print</span><span class="p">(</span><span class="s2">"Syntax errors are annoying."</span><span class="p">)</span>
# <span class="k">print</span><span class="p">(</span><span class="s2">"But at least python tells us about them!"</span><span class="p">)</span>
# <span class="k">print</span><span class="p">(</span><span class="s2">"So they are usually not too hard to fix."</span><span class="p">)</span>
# </pre></div>
#
# </div>
#
# </section>
#
#
# <section class="solution panel panel-primary">
# <div class="panel-heading">
# <h2><span class="fa fa-eye"></span> Solution</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p><code>SyntaxError</code> for missing <code>():</code> at end of first line,
# ndentationError` for mismatch between second and third lines.
# A fixed version is:</p>
#
# </div>
#
# </section>
#
def another_function():
print("syntax errors are annoying")
print("but at least python tells us about them")
print("so they are usually not too hard to fix")
#
# <section class="challenge panel panel-success">
# <div class="panel-heading">
# <h2><span class="fa fa-pencil"></span> Identifying Variable Name Errors</h2>
# </div>
#
#
# <div class="panel-body">
#
# <ol>
# <li>Read the code below, and (without running it) try to identify what the errors are.</li>
# <li>Run the code, and read the error message.
# What type of <code>NameError</code> do you think this is?
# In other words, is it a string with no quotes,
# a misspelled variable,
# or a variable that should have been defined but was not?</li>
# <li>Fix the error.</li>
# <li>Repeat steps 2 and 3, until you have fixed all the errors.</li>
# </ol>
# <div class="codehilite"><pre><span></span><span class="k">for</span> <span class="n">number</span> <span class="ow">in</span> <span class="nb">range</span><span class="p">(</span><span class="mi">10</span><span class="p">):</span>
# <span class="c1"># use a if the number is a multiple of 3, otherwise use b</span>
# <span class="k">if</span> <span class="p">(</span><span class="n">Number</span> <span class="o">%</span> <span class="mi">3</span><span class="p">)</span> <span class="o">==</span> <span class="mi">0</span><span class="p">:</span>
# <span class="n">message</span> <span class="o">=</span> <span class="n">message</span> <span class="o">+</span> <span class="n">a</span>
# <span class="k">else</span><span class="p">:</span>
# <span class="n">message</span> <span class="o">=</span> <span class="n">message</span> <span class="o">+</span> <span class="s2">"b"</span>
# <span class="k">print</span><span class="p">(</span><span class="n">message</span><span class="p">)</span>
# </pre></div>
#
# </div>
#
# </section>
#
#
# <section class="solution panel panel-primary">
# <div class="panel-heading">
# <h2><span class="fa fa-eye"></span> Solution</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p>3 <code>NameError</code>s for:</p>
# <ul>
# <li><code>number</code> being misspelled</li>
# <li><code>message</code> not defined</li>
# <li><code>a</code> not being in quotes.</li>
# </ul>
# <p>Fixed version:</p>
#
# </div>
#
# </section>
#
message = ""
for number in range(10):
# use a if the number is a multiple of 3, otherwise use b
if (number % 3) == 0:
message = message + "a"
else:
message = message + "b"
print(message)
#
# <section class="challenge panel panel-success">
# <div class="panel-heading">
# <h2><span class="fa fa-pencil"></span> Identifying Index Errors</h2>
# </div>
#
#
# <div class="panel-body">
#
# <ol>
# <li>Read the code below, and (without running it) try to identify what the errors are.</li>
# <li>Run the code, and read the error message. What type of error is it?</li>
# <li>Fix the error.</li>
# </ol>
# <div class="codehilite"><pre><span></span><span class="n">seasons</span> <span class="o">=</span> <span class="p">[</span><span class="s1">'Spring'</span><span class="p">,</span> <span class="s1">'Summer'</span><span class="p">,</span> <span class="s1">'Fall'</span><span class="p">,</span> <span class="s1">'Winter'</span><span class="p">]</span>
# <span class="k">print</span><span class="p">(</span><span class="s1">'My favorite season is '</span><span class="p">,</span> <span class="n">seasons</span><span class="p">[</span><span class="mi">4</span><span class="p">])</span>
# </pre></div>
#
# </div>
#
# </section>
#
#
# <section class="solution panel panel-primary">
# <div class="panel-heading">
# <h2><span class="fa fa-eye"></span> Solution</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p><code>IndexError</code>; the last entry is <code>seasons[3]</code>, so <code>seasons[4]</code> doesn't make sense.
# A fixed version is:</p>
#
# </div>
#
# </section>
#
| 05-writing-effective-tests/01-errors-and-exceptions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="kox2ddGqIBMZ" outputId="cda131e5-0cb0-4c5c-bbca-92ff23c28756" colab={"base_uri": "https://localhost:8080/"}
# !pip install -U git+https://github.com/d2l-ai/d2l-en.git@master
# + id="SCz0svAwHqDq" outputId="2fce3231-3f98-4164-e314-1aa7eced9f5d" colab={"base_uri": "https://localhost:8080/"}
from d2l import torch as d2l
import torch
from torch import nn
batch_size = 64
train_iter, test_iter, vocab = d2l.load_data_imdb(batch_size)
# + id="tJlrYh_5HsN6"
def corr1d(X, K):
w = K.shape[0]
Y = torch.zeros((X.shape[0] - w + 1))
for i in range(Y.shape[0]):
Y[i] = (X[i: i + w] * K).sum()
return Y
# + id="i96qKjKHHulY" outputId="c7f91440-b730-48da-e8c8-90973a18900c" colab={"base_uri": "https://localhost:8080/"}
X, K = torch.tensor([0, 1, 2, 3, 4, 5, 6]), torch.tensor([1, 2])
corr1d(X, K)
# + id="lrxgfk01HxQ1" outputId="63604f5a-3221-4ea1-b09d-729ff9f3134c" colab={"base_uri": "https://localhost:8080/"}
def corr1d_multi_in(X, K):
# First, we traverse along the 0th dimension (channel dimension) of `X`
# and `K`. Then, we add them together by using * to turn the result list
# into a positional argument of the `add_n` function
return sum(corr1d(x, k) for x, k in zip(X, K))
X = torch.tensor([[0, 1, 2, 3, 4, 5, 6],
[1, 2, 3, 4, 5, 6, 7],
[2, 3, 4, 5, 6, 7, 8]])
K = torch.tensor([[1, 2], [3, 4], [-1, -3]])
corr1d_multi_in(X, K)
# + id="kQ-RhlDVHx2U"
class TextCNN(nn.Module):
def __init__(self, vocab_size, embed_size, kernel_sizes, num_channels,
**kwargs):
super(TextCNN, self).__init__(**kwargs)
self.embedding = nn.Embedding(vocab_size, embed_size)
# The embedding layer does not participate in training
self.constant_embedding = nn.Embedding(vocab_size, embed_size)
self.dropout = nn.Dropout(0.5)
self.decoder = nn.Linear(sum(num_channels), 2)
# The max-over-time pooling layer has no weight, so it can share an
# instance
self.pool = nn.AdaptiveAvgPool1d(1)
self.relu = nn.ReLU()
# Create multiple one-dimensional convolutional layers
self.convs = nn.ModuleList()
for c, k in zip(num_channels, kernel_sizes):
self.convs.append(nn.Conv1d(2 * embed_size, c, k))
def forward(self, inputs):
# Concatenate the output of two embedding layers with shape of
# (batch size, no. of words, word vector dimension) by word vector
embeddings = torch.cat((
self.embedding(inputs), self.constant_embedding(inputs)), dim=2)
# According to the input format required by Conv1d, the word vector
# dimension, that is, the channel dimension of the one-dimensional
# convolutional layer, is transformed into the previous dimension
embeddings = embeddings.permute(0, 2, 1)
# For each one-dimensional convolutional layer, after max-over-time
# pooling, a tensor with the shape of (batch size, channel size, 1)
# can be obtained. Use the flatten function to remove the last
# dimension and then concatenate on the channel dimension
encoding = torch.cat([
torch.squeeze(self.relu(self.pool(conv(embeddings))), dim=-1)
for conv in self.convs], dim=1)
# After applying the dropout method, use a fully connected layer to
# obtain the output
outputs = self.decoder(self.dropout(encoding))
return outputs
# + id="In9xt9XbHz6_"
embed_size, kernel_sizes, nums_channels = 100, [3, 4, 5], [100, 100, 100]
devices = d2l.try_all_gpus()
net = TextCNN(len(vocab), embed_size, kernel_sizes, nums_channels)
def init_weights(m):
if type(m) in (nn.Linear, nn.Conv1d):
nn.init.xavier_uniform_(m.weight)
net.apply(init_weights);
# + id="XTMZcJjhH1cH" outputId="7c2a5b40-b4f9-4fec-b5e0-d9c643c96c72" colab={"base_uri": "https://localhost:8080/"}
glove_embedding = d2l.TokenEmbedding('glove.6b.100d')
embeds = glove_embedding[vocab.idx_to_token]
net.embedding.weight.data.copy_(embeds)
net.constant_embedding.weight.data.copy_(embeds)
net.constant_embedding.weight.requires_grad = False
# + id="OkRHOq-AH3Kw" outputId="cc9cd7dc-ebd9-4e5f-a7cf-3dd3792a89d0" colab={"base_uri": "https://localhost:8080/", "height": 301}
lr, num_epochs = 0.001, 5
trainer = torch.optim.Adam(net.parameters(), lr=lr)
loss = nn.CrossEntropyLoss(reduction="none")
d2l.train_ch13(net, train_iter, test_iter, loss, trainer, num_epochs, devices)
# + id="VKr6U4MhH4Z2" outputId="c60439c0-3796-4982-c055-894aafd5f226" colab={"base_uri": "https://localhost:8080/", "height": 35}
d2l.predict_sentiment(net, vocab, 'this movie is so great')
# + id="7jH-ar_BH6Bs" outputId="04c77daf-85e2-4cfc-f143-c6f34312e643" colab={"base_uri": "https://localhost:8080/", "height": 35}
d2l.predict_sentiment(net, vocab, 'this movie is so bad')
| Ch15_NLP-applications/sentiment-analysis_cnn_pt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression
# 
# ## Importing the data
from sklearn.datasets import load_boston
data=load_boston()
# ## Let's have a look at what the data has in it
print(data.DESCR)
# ## Getting Features and Target values from Data
import pandas as pd
boston=pd.DataFrame(data["data"],columns=data["feature_names"])
boston.head()
prices=pd.DataFrame(data["target"],columns=["price"])
prices.head()
# ## Visualize the features
boston.plot()
# ## Visualize the Target variable
prices.plot()
# ## Splitting the data
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(boston,prices)
print("Train Data Size:",X_train.size)
print("Test Data Size:",X_test.size)
# ## Modelling the data
from sklearn.linear_model import LinearRegression
model=LinearRegression()
model.fit(X_train,y_train)
print(model)
# ## Let's test with our Test Data
predicted=model.predict(X_test)
# ## Let's compare using our metrics in sklearn
# ### Metrics for Regression analysis :
#
# * RMSE ( Root Mean Squared Error )
# * MAE ( Mean Absolute Error )
# * R-Squared
from sklearn.metrics import mean_squared_error, r2_score
print("Mean Squared Error: %.2f"% mean_squared_error(y_test,predicted))
print("R-Squared Error: %.2f"% r2_score(y_test,predicted))
| LinearRegression/LinearRegression_House_Prices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from pandas import Series, DataFrame
np.random.seed(12345)
dframe = DataFrame(np.random.randn(1000,4))
dframe
dframe.describe()
col = dframe[0]
col
col[np.abs(col)>3]
dframe[(np.abs(dframe)>3).any(1)]
dframe[np.abs(dframe)>3] = np.sign(dframe) *3
dframe.describe()
| Working with Data/Outliers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# <h1> Create Keras DNN model </h1>
#
# This notebook illustrates:
# <ol>
# <li> Creating a model using Keras. This requires TensorFlow 2.0
# </ol>
# + deletable=true editable=true
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
# + deletable=true editable=true
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
# + deletable=true editable=true language="bash"
# if ! gsutil ls | grep -q gs://${BUCKET}/; then
# gsutil mb -l ${REGION} gs://${BUCKET}
# fi
# + deletable=true editable=true language="bash"
# ls *.csv
# + [markdown] deletable=true editable=true
# ## Create Keras model
# <p>
# First, write an input_fn to read the data.
# + deletable=true editable=true
import shutil
import numpy as np
import tensorflow as tf
print(tf.__version__)
# + deletable=true editable=true
# Determine CSV, label, and key columns
CSV_COLUMNS = 'weight_pounds,is_male,mother_age,plurality,gestation_weeks,key'.split(',')
LABEL_COLUMN = 'weight_pounds'
KEY_COLUMN = 'key'
# Set default values for each CSV column. Treat is_male and plurality as strings.
DEFAULTS = [[0.0], ['null'], [0.0], ['null'], [0.0], ['nokey']]
# +
def features_and_labels(row_data):
for unwanted_col in ['key']:
row_data.pop(unwanted_col)
label = row_data.pop(LABEL_COLUMN)
return row_data, label # features, label
# load the training data
def load_dataset(pattern, batch_size=1, mode=tf.estimator.ModeKeys.EVAL):
dataset = (tf.data.experimental.make_csv_dataset(pattern, batch_size, CSV_COLUMNS, DEFAULTS)
.map(features_and_labels) # features, label
)
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.shuffle(1000).repeat()
dataset = dataset.prefetch(1) # take advantage of multi-threading; 1=AUTOTUNE
return dataset
# + [markdown] deletable=true editable=true
# Next, define the feature columns. mother_age and gestation_weeks should be numeric.
# The others (is_male, plurality) should be categorical.
# +
## Build a simple Keras DNN using its Functional API
def rmse(y_true, y_pred):
return tf.sqrt(tf.reduce_mean(tf.square(y_pred - y_true)))
# Helper function to handle categorical columns
def categorical_fc(name, values):
return tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_vocabulary_list(name, values))
def build_dnn_model():
# input layer
inputs = {
colname : tf.keras.layers.Input(name=colname, shape=(), dtype='float32')
for colname in ['mother_age', 'gestation_weeks']
}
inputs.update({
colname : tf.keras.layers.Input(name=colname, shape=(), dtype='string')
for colname in ['is_male', 'plurality']
})
# feature columns from inputs
feature_columns = {
colname : tf.feature_column.numeric_column(colname)
for colname in ['mother_age', 'gestation_weeks']
}
feature_columns['is_male'] = categorical_fc('is_male', ['True', 'False', 'Unknown'])
feature_columns['plurality'] = categorical_fc('plurality',
['Single(1)', 'Twins(2)', 'Triplets(3)',
'Quadruplets(4)', 'Quintuplets(5)','Multiple(2+)'])
# the constructor for DenseFeatures takes a list of numeric columns
# The Functional API in Keras requires that you specify: LayerConstructor()(inputs)
dnn_inputs = tf.keras.layers.DenseFeatures(feature_columns.values())(inputs)
# two hidden layers of [64, 32] just in like the BQML DNN
h1 = tf.keras.layers.Dense(64, activation='relu', name='h1')(dnn_inputs)
h2 = tf.keras.layers.Dense(32, activation='relu', name='h2')(h1)
# final output is a linear activation because this is regression
output = tf.keras.layers.Dense(1, activation='linear', name='weight')(h2)
model = tf.keras.models.Model(inputs, output)
model.compile(optimizer='adam', loss='mse', metrics=[rmse, 'mse'])
return model
print("Here is our DNN architecture so far:\n")
model = build_dnn_model()
print(model.summary())
# -
# We can visualize the DNN using the Keras plot_model utility.
tf.keras.utils.plot_model(model, 'dnn_model.png', show_shapes=False, rankdir='LR')
# + [markdown] deletable=true editable=true
# ## Train and evaluate
# + deletable=true editable=true
TRAIN_BATCH_SIZE = 32
NUM_TRAIN_EXAMPLES = 10000 * 5 # training dataset repeats, so it will wrap around
NUM_EVALS = 5 # how many times to evaluate
NUM_EVAL_EXAMPLES = 10000 # enough to get a reasonable sample, but not so much that it slows down
trainds = load_dataset('train*', TRAIN_BATCH_SIZE, tf.estimator.ModeKeys.TRAIN)
evalds = load_dataset('eval*', 1000, tf.estimator.ModeKeys.EVAL).take(NUM_EVAL_EXAMPLES//1000)
steps_per_epoch = NUM_TRAIN_EXAMPLES // (TRAIN_BATCH_SIZE * NUM_EVALS)
history = model.fit(trainds,
validation_data=evalds,
epochs=NUM_EVALS,
steps_per_epoch=steps_per_epoch)
# + [markdown] deletable=true editable=true
# ## Visualize loss curve
# +
# plot
import matplotlib.pyplot as plt
nrows = 1
ncols = 2
fig = plt.figure(figsize=(10, 5))
for idx, key in enumerate(['loss', 'rmse']):
ax = fig.add_subplot(nrows, ncols, idx+1)
plt.plot(history.history[key])
plt.plot(history.history['val_{}'.format(key)])
plt.title('model {}'.format(key))
plt.ylabel(key)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left');
# -
# ## Save the model
# + deletable=true editable=true
import shutil, os, datetime
OUTPUT_DIR = 'babyweight_trained'
shutil.rmtree(OUTPUT_DIR, ignore_errors=True)
EXPORT_PATH = os.path.join(OUTPUT_DIR, datetime.datetime.now().strftime('%Y%m%d%H%M%S'))
tf.saved_model.save(model, EXPORT_PATH) # with default serving function
# + [markdown] deletable=true editable=true
# <h2> Monitor and experiment with training </h2>
# -
# To begin TensorBoard from within AI Platform Notebooks, click the + symbol in the top left corner and select the **Tensorboard** icon to create a new TensorBoard.
# + [markdown] deletable=true editable=true
# In TensorBoard, look at the learned embeddings. Are they getting clustered? How about the weights for the hidden layers? What if you run this longer? What happens if you change the batchsize?
# + [markdown] deletable=true editable=true
# Copyright 2017-2018 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/06_structured/3_keras_dnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
root_directory = 'D:\\'
from multiprocessing.dummy import Pool
from pathlib import Path
import pandas as pd
from dask.utils import format_bytes
import plotly.express as px
# + id="8e7TeCo2QKrP"
def gen_file_row(f):
return {'path':str(f),'filename':f.name,'directory':str(f.parent),'type':f.suffix,'bytes':f.stat().st_size}, f.stat().st_size
def process_dir(d):
paths = []
dir_size = 0
try:
for p in d.iterdir():
try:
if p.is_dir():
res, b = process_dir(p)
paths = paths + res
dir_size += b
elif p.is_file():
res, b = gen_file_row(p)
paths.append(res)
dir_size += b
except (FileNotFoundError, PermissionError):
continue
except (FileNotFoundError, PermissionError):
paths.append({'path':str(d),
'filename':d.name if d.name != '' else str(d),
'directory':str(d.parent) if d.parent != d else "",
'type':'directory',
'bytes':0})
return paths, 0
paths.append({'path':str(d),
'filename':d.name if d.name != '' else str(d),
'directory':str(d.parent) if d.parent != d else "",
'type':'directory',
'bytes':dir_size})
return paths, dir_size
def process_with_Pool(d):
pool = Pool()
glob_list = list(d.glob("**/*"))
dir_list = [{'path':x.name,'directory':str(x.parent),'type':'directory','bytes':0} for x in glob_list if x.is_dir()]
print(dir_list[0:5])
file_list = [x for x in glob_list if not x.is_dir()]
res = pool.map(gen_file_row, file_list)
return dir_list + res
# + colab={"base_uri": "https://localhost:8080/", "height": 477} id="KekjpZuid3OA" outputId="7a13c84d-8d26-471f-d8fe-ea08f352344d"
import pandas as pd
#data=process_dir_Pool(Path(root_directory))
data, __ =process_dir(Path(root_directory))
df = pd.DataFrame(data)
df['filesize'] = df['bytes'].apply(lambda x: format_bytes(x))
# -
with pd.option_context('display.max_colwidth', None):
largest_dir = df.loc[(df['type'] == 'directory') & (df['path'] != root_directory)].sort_values(by=['bytes','directory'],ascending=[False,True]).head(10).reset_index()
largest_dir.index = range(1,11)
print("10 largest directories")
display(largest_dir[['path','directory','filesize']])
with pd.option_context('display.max_colwidth', None):
print("10 largest files")
largest_files = df.loc[df['type'] != 'directory'].nlargest(10,'bytes').reset_index(drop=True)
largest_files.index = range(1,11)
display(largest_files[['filename','directory','type','filesize']])
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="odajmRGSi0en" outputId="337d0bfa-d633-4247-a6b6-5171370f5a74"
fig = px.treemap(df.nlargest(100,'bytes').sort_values(by='directory'),
branchvalues = "total",
names = 'filename',
ids = 'path',
parents = 'directory',
values = 'bytes',
title = '100 largest directories/files',
hover_data = {'filename':True,'path':False,'bytes':False,'filesize':True,'directory':True,'type':True},
color = 'type'
)
#fig.update_traces(root_color="lightgrey")
fig.update_layout(margin = dict(t=50, l=25, r=25, b=25))
fig.show()
# -
| PyDirTreeMap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <NAME>. <NAME>. 2019 г.
#
# ## Газосодержание
# <a id="Rs"></a>
#
# ### Газосодержание, корреляция Стендинга
# <a id="Rs_Standing"></a>
#
# Для расчета газосодержания используется корреляция, обратная корреляции Стендинга для давления насыщения нефти газом.
#
# $$ R_s = \gamma_g \left( \frac{1.92 p}{\ 10^{y_g}}\right)^{1.204} $$
#
# где:
#
# $R_s$ - газосодержание, $м^3/м^3 $
#
# $P$ - давление, $МПа$
#
# $\gamma_g$ - относительная плотность газа, безразмерная величина
#
# $y_g$ - мольная доля газа, $ y_g = 1.225 +0.00164 T - \frac{ 1.769}{\gamma_o}$
#
# $\gamma_o$ - относительная плотность нефти, безразмерная величина
#
# $ T $ - температура, $ ^{\circ}\mathrm{K}$
#
# Газосодержание является одним из ключевых свойств нефти при расчётах производительности скважин и работы скважинного оборудования. Динамика изменения газосодержания во многом определяет количество свободного газа в потоке и должна учитываться при проведении расчётов.
#
# Если известно газосодержание при давлении насыщения, то газосодержание при давлениях ниже давления насыщения может быть получено из пропорции:
#
# $$ R_s = \ R_{sb}\left( \frac{p}{\ P_b}\right)^{1.204} $$
#
# где:
#
# $R_s$ - газосодержание, $м^3/м^3 $
#
# $P$ - давление, $МПа$
#
# $P_b$ - давление насыщения, $МПа$
#
# $R_{sb}$ - газосодержание при давлении насыщения, $м^3/м^3 $
#
# Корреляции Standing базируются на 105 экспериментально определенных давлениях насыщения нефтяных систем Калифорнии. Диапазоны значений основных свойств, использованных для разработки данной корреляции, приведены в таблице ниже.
#
# | <p align="left"> Параметр | Диапазон |
# | :--- | :--- |
# | <p align="left"> давление насыщения,$P_b$ , $ МПа $ | 0.896…48.263 |
# | <p align="left"> температура, $^{\circ}\mathrm{K} $ | 310…400 |
# | <p align="left"> газосодержание при давлении насыщения, $R_{sb}$ , $м^3/м^3 $ | 3.6…254 |
# | <p align="left"> относительная плотность нефти по воде, $\gamma_o$ | 0.725…0.956 |
# | <p align="left"> относительная плотность газа, $\gamma_g$ | 0.59…0.95 |
#
#
# ref "A Pressure-Volume-Temperature Correlation for Mixtures of California Oil and Gases", M.B. Standing, Drill. & Prod. Prac., API, 1947.
import matplotlib.pyplot as plt
import matplotlib as mpl
import numpy as np
from sympy import *
init_printing()
import sys
sys.path.append('../')
import uniflocpy.uPVT.PVT_correlations as PVT
# +
# Корреляция Стендинга для газосодержания реализована
# в виде функции unf_Rs_Standing_m3m3 в модуле PVT_correlations.
# Подробные данные по функции включая исходный код приведены ниже
# PVT.unf_rs_Standing_m3m3??
# -
# параметры определяющие диапазоны значений для построения графиков
p_set=np.arange(1,300,10)
t_set=np.arange(273,380,30)
t_set_def=np.array([313])
gg_set=np.arange(0.6,1,0.1)
gg_set_def=np.array([0.8])
go_set=np.arange(0.8,1,0.05)
go_set_def=np.array([0.86])
# функция для автоматизации построения графиков по давлению насыщения
def prep_plot(func,tset,goset,ggset,plot_title,plot_xlab,plot_ylab):
for t in tset:
for gg in ggset:
for go in goset:
Rs_set=[]
for p in p_set:
Rs_set.append(func(p,t_K = t,gamma_gas = gg,gamma_oil = go))
plt.plot(p_set, Rs_set, label='t = %1.0f $ ^{\circ}\mathrm{K}$'%t +
' $\gamma_g$ = %1.2f'%gg +
' $\gamma_o$ = %1.2f'%go)
plt.title(plot_title)
plt.ylabel(plot_ylab, color = 'black')
plt.xlabel(plot_xlab, color = 'black')
plt.legend()
# +
# код для построения графиков
plt.figure(figsize=(15,8))
f = PVT.unf_rs_Standing_m3m3
# рисуем первый график
plt.subplot(221)
prep_plot(f,t_set,go_set_def,gg_set_def,
'Газосодержание от давления',
'$P, MPa$',
'$R_s, м^3/м^3$')
plt.grid()
# рисуем второй график
plt.subplot(222)
prep_plot(f,t_set_def,go_set,gg_set_def,
'Газосодержание от давления',
'$P, MPa$',
'$R_s, м^3/м^3$')
plt.grid()
# рисуем третий график
plt.subplot(223)
prep_plot(f,t_set_def,go_set_def,gg_set,
'Газосодержание от давления',
'$P, MPa$',
'$R_s, м^3/м^3$')
plt.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.25, wspace=0.35)
# рисуем все
plt.grid()
plt.show()
# -
# ### Газосодержание, корреляция Веларде-Маккейна
# <a id="Rs_McCain"></a>
# Корреляция Веларде-Маккейна (1999) для газосодержания базируется на 718 лабораторных исследованиях разгазирования различных нефтей со всего мира.
#
# $$ R_s = R_{sb}R_{sr} $$
#
# где:
#
# $R_s$ - газосодержание, $м^3/м^3$
#
# $R_{sb}$ - газосодержание при давлении насыщения, $м^3/м^3$
#
#
#
#
# $R_{sr}=a_1p_r^{a_2} + (1-a_1)p_r^{a_3}$ - приведенное газосодержание
#
# $p_r=\frac{(p-14,7)}{(p_b-14,7)}$ - степень превышения давления, $psig/psig$
#
# $a_1=9.73 \cdot 10^{-7}\gamma_{gSP}^{1.672608}API^{0.929870}T^{0.247235}(p_b-14.7)^{1.056052}$
#
# $a_2=0.022339 \gamma_{gSP}^{-1.004750}API^{0.337711}T^{0.132795}(p_b-14.7)^{0.302065}$
#
# $a_3=0.725167 \gamma_{gSP}^{-1.485480}API^{-0.164741}T^{-0.091330}(p_b-14.7)^{0.047094}$
#
# В приведенной таблице представлены диапазоны значений использованных для создания корреляции:
#
# | <p align="left"> Параметр | Мин|Среднее|Макс|
# | :--- | :---: |:---:|:---:|
# | <p align="left"> давление насыщения,$P_b$ , $ МПа $ | 2.861 |15.706|53.434|
# | <p align="left"> температура, $^{\circ}\mathrm{С} $ | 21 |86|160|
# | <p align="left"> относительная плотность газа на сепараторе, $\gamma_g$ | 0.555 |0.793|1.472|
# | <p align="left"> объемный коэффициент нефти при давлении насыщения, $B_{ob}$ , $ м^3/м^3 $ | 1.012 |1.358|2.042|
#
# +
# Корреляция Веларде для газосодержания реализована
# в виде функции unf_Rs_Velarde_m3m3 в модуле PVT_correlations.
# Подробные данные по функции включая исходный код приведены ниже
# PVT.unf_rs_Velarde_m3m3??
# -
# параметры определяющие диапазоны значений для построения графиков
p_set=np.arange(1,11,0.25)
t_set=np.arange(294,400,30)
t_set_def=np.array([313])
gg_set=np.arange(0.6,1,0.1)
gg_set_def=np.array([0.8])
go_set=np.arange(0.8,1,0.05)
go_set_def=np.array([0.86])
# функция для автоматизации построения графиков по давлению насыщения
def prep_plot(func,tset,goset,ggset,plot_title,plot_xlab,plot_ylab):
for t in tset:
for gg in ggset:
for go in goset:
Rs_set=[]
for p in p_set:
Rs_set.append(func(p,t_K = t,gamma_gas = gg,gamma_oil = go))
plt.plot(p_set, Rs_set, label='t = %1.0f $ ^{\circ}\mathrm{K}$'%t +
' $\gamma_g$ = %1.2f'%gg +
' $\gamma_o$ = %1.2f'%go)
plt.title(plot_title)
plt.ylabel(plot_ylab, color = 'black')
plt.xlabel(plot_xlab, color = 'black')
plt.legend()
# +
# код для построения графиков
plt.figure(figsize=(15,8))
f = PVT.unf_rs_Velarde_m3m3
# рисуем первый график
plt.subplot(221)
plt.grid()
prep_plot(f,t_set,go_set_def,gg_set_def,
'Газосодержание от давления',
'$P, MPa$',
'$R_s, м^3/м^3$')
# рисуем второй график
plt.subplot(222)
plt.grid()
prep_plot(f,t_set_def,go_set,gg_set_def,
'Газосодержание от давления',
'$P, MPa$',
'$R_s, м^3/м^3$')
# рисуем третий график
plt.subplot(223)
prep_plot(f,t_set_def,go_set_def,gg_set,
'Газосодержание от давления',
'$P, MPa$',
'$R_s, м^3/м^3$')
plt.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.25, wspace=0.35)
# рисуем все
plt.grid()
plt.show()
# -
# ### Газосодержание при давлении насыщения, Корреляция Маккейна если известны данные в сепараторе и резервуаре
# <a id="Rsb_McCain"></a>
# Во многих корреляциях используется в качестве исходного параметра - газосодержание при давлении насыщения. Значение этого параметра может быть получено с помощью полевых данных как сумма отделяемого газового фактора в сепараторе и резервуаре для товарной нефти.
#
#
# $$ R_{sb} = R_{sp} + R_{st} $$
#
# где:
#
# $R_{sb}$ - газосодержание при давлении насыщения, $м^3/м^3$
#
# $R_{sp}$ - газовый фактор, отделяемый в сепараторе, $м^3/м^3$
#
# $R_{st}$ - газовый фактор в резервуаре для товарной нефти, $м^3/м^3$
#
# Данное уравнение справедливо только если пластовое давление выше давления насыщения
#
# ---
# <img src="pics/McCain_Rsb with stock tank and separator.jpg" width="600" >
#
# Расход газа и дебит нефти в сепараторе обычно измеряются, когда как в резервуаре газ обычно выпускается и не замеряется. Поэтому для более точной оценки газосодержания пластовой нефти при давлении насыщения необходимо оценить газовый фактор в резервуаре.
# Таким образом, был разработана формула на основе GRACE-алгоритма на базе 898 исследований пластового флюида. В таблице приведены набор данных для исследования.
#
# <img src="pics/McCain_Rsb with stock tank and separator_data.jpg" width="600" >
#
# $$ ln R_{st} = 3.955 + 0.83z - 0.024z^2 + 0.075z^3 $$
#
# $$ z =\sum_{n=1}^3 z_n $$
#
# $$ z_n = C0_n + C1_nVAR_n + C2_nVAR_n^2 $$
#
# | <p align="left"> $$n$$ | $$VAR$$|$$CO$$|$$C1$$|$$C2$$|
# | :--- | :---: |:---:|:---:|:---:|
# | <p align="left"> $$1$$ | $$ln p_{sp} $$ |$$-8.005$$|$$2.7$$|$$-0.161$$|
# | <p align="left"> $$2$$ | $$ln T_{sp}$$ |$$1.224$$|$$-0.5$$|$$0$$|
# | <p align="left"> $$3$$ | $$API$$ |$$-1.587$$|$$0.0441$$|$$-2.29 \cdot 10 ^{-5}$$|
#
# $$T_{sp} - °F, p_{sp} - psia $$
#
# Выражение для вычисления газового фактора в резервуаре требует знать температуру и давление в сепараторе, которые обычно не всегда бывают известны. Поэтому в этом случае можно использовать следующее уравнение.
#
# $$ R_{sb} = 1.1618 R_{sp} $$
# +
# Корреляция Маккейна для газосодержания при давлении насыщения реализована
# в виде функции unf_Rsb_Mccain_m3m3 в модуле PVT_correlations.
# Подробные данные по функции включая исходный код приведены ниже
# PVT.unf_rsb_Mccain_m3m3??
# -
# параметры определяющие диапазоны значений для построения графиков
p_set=np.arange(1,11,0.25)
t_set=np.arange(294,400,30)
t_set_def=np.array([313])
go_set=np.arange(0.8,1,0.05)
go_set_def=np.array([0.86])
r_sp = 50
# функция для автоматизации построения графиков по давлению насыщения
def prep_plot(func,tset,goset,plot_title,plot_xlab,plot_ylab):
for t in tset:
for go in goset:
Rs_set=[]
for p in p_set:
Rs_set.append(func(r_sp,go,p,t))
plt.plot(p_set, Rs_set, label='t = %1.0f $ ^{\circ}\mathrm{K}$'%t +
' $\gamma_o$ = %1.2f'%go)
plt.title(plot_title)
plt.ylabel(plot_ylab, color = 'black')
plt.xlabel(plot_xlab, color = 'black')
plt.legend()
# +
# код для построения графиков
plt.figure(figsize=(15,8))
f = PVT.unf_rsb_Mccain_m3m3
# рисуем первый график
plt.subplot(221)
plt.grid()
prep_plot(f,t_set,go_set_def,
'Газосодержание от давления в сепараторе',
'$P, MPa$',
'$R_s, м^3/м^3$')
# рисуем второй график
plt.subplot(222)
plt.grid()
prep_plot(f,t_set_def,go_set,
'Газосодержание от давления в сепараторе',
'$P, MPa$',
'$R_s, м^3/м^3$')
plt.subplots_adjust(top=0.92, bottom=0.08, left=0.10, right=0.95, hspace=0.25, wspace=0.35)
# рисуем все
plt.show()
| notebooks/uPVT. Solution Gas-oil ratio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="8ZtuUVpcCEno" colab_type="code" colab={}
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# + id="l0gBTuTZCPp_" colab_type="code" colab={}
train = pd.read_csv("/content/train_mod.csv")
# + id="gJMuBPLsIBdW" colab_type="code" colab={}
tr = pd.read_csv("/content/train.csv")
y=tr["Survived"]
# + id="XI4MvhrtIBhQ" colab_type="code" colab={}
# + id="kSigtyYyChRw" colab_type="code" colab={}
test = pd.read_csv("/content/test_mod.csv")
# + id="wURpx3hFDF5A" colab_type="code" colab={}
X_test = test
X_test = X_test.drop(columns=["PassengerId"])
# + id="xNilALBuHlVS" colab_type="code" outputId="fd459bb1-f76d-4d8d-d82e-dccc7c9de63c" colab={"base_uri": "https://localhost:8080/", "height": 195}
train.head()
# + id="hcMtnZeXDNNy" colab_type="code" outputId="e61b5d03-b10a-44bd-efe2-5bf8512862fd" colab={"base_uri": "https://localhost:8080/", "height": 403}
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV,StratifiedKFold,cross_val_score,train_test_split
from sklearn.metrics import accuracy_score
rfc = RandomForestClassifier(random_state=100)
param_grid = {
'n_estimators': [200, 500],
'max_features': ['auto', 'sqrt', 'log2'],
'max_depth' : [4,5,6,7,8],
'criterion' :['gini', 'entropy']
}
kf = StratifiedKFold(n_splits=5)
grfc = GridSearchCV(estimator = rfc,param_grid=param_grid,cv=kf)
X_train,X_tes,y_train,y_test = train_test_split(train,y,random_state=69)
grfc.fit(X_train,y_train)
# + id="k2ZbRFVyKsr0" colab_type="code" outputId="3b561e5a-6e16-4ace-acb5-27c8c9f85416" colab={"base_uri": "https://localhost:8080/", "height": 34}
grfc.best_score_
# + id="d6YBB3hiLbns" colab_type="code" outputId="f7774311-0d5e-4664-e926-070c7c5c56c4" colab={"base_uri": "https://localhost:8080/", "height": 84}
grfc.best_params_
# + id="y8155B9zLfBe" colab_type="code" colab={}
y_pred1 = grfc.predict(X_tes)
# + id="hU4F_HEuLkAA" colab_type="code" outputId="a854c134-9e03-4451-9517-b545d524ee45" colab={"base_uri": "https://localhost:8080/", "height": 34}
accuracy_score(y_test,y_pred1)
# + id="691ZsEhwLpQM" colab_type="code" colab={}
y_pred1 = grfc.predict(X_test)
# + id="IF1rxJMtLwfu" colab_type="code" colab={}
rfc_grid= pd.DataFrame({
"PassengerId": test["PassengerId"],
"Survived": y_pred1
})
# + id="l9SAdpu7MBQl" colab_type="code" colab={}
rfc_grid.to_csv("/content/rfc_grid_data12.csv", index=False)
# + id="SHnKXzsiMI6M" colab_type="code" colab={}
| RFC_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model ensembling
#
# This example illustrates how to vectorize model ensembling using vmap.
#
# ## What is model ensembling?
# Model ensembling combines the predictions from multiple models together.
# Traditionally this is done by running each model on some inputs separately
# and then combining the predictions. However, if you're running models with
# the same architecture, then it may be possible to combine them together
# using `vmap`. `vmap` is a function transform that maps functions across
# dimensions of the input tensors. One of its use cases is eliminating
# for-loops and speeding them up through vectorization.
#
# Let's demonstrate how to do this using an ensemble of simple CNNs.
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
from functools import partial
torch.manual_seed(0)
# Here's a simple CNN
class SimpleCNN(nn.Module):
def __init__(self):
super(SimpleCNN, self).__init__()
self.conv1 = nn.Conv2d(1, 32, 3, 1)
self.conv2 = nn.Conv2d(32, 64, 3, 1)
self.fc1 = nn.Linear(9216, 128)
self.fc2 = nn.Linear(128, 10)
def forward(self, x):
x = self.conv1(x)
x = F.relu(x)
x = self.conv2(x)
x = F.relu(x)
x = F.max_pool2d(x, 2)
x = torch.flatten(x, 1)
x = self.fc1(x)
x = F.relu(x)
x = self.fc2(x)
output = F.log_softmax(x, dim=1)
output = x
return output
# -
# Let's generate some dummy data. Pretend that we're working with an MNIST dataset
# where the images are 28 by 28.
# Furthermore, let's say we wish to combine the predictions from 10 different
# models.
device = 'cuda'
num_models = 10
data = torch.randn(100, 64, 1, 28, 28, device=device)
targets = torch.randint(10, (6400,), device=device)
models = [SimpleCNN().to(device) for _ in range(num_models)]
# We have a couple of options for generating predictions. Maybe we want
# to give each model a different randomized minibatch of data, or maybe we
# want to run the same minibatch of data through each model (e.g. if we were
# testing the effect of different model initializations).
# Option 1: different minibatch for each model
minibatches = data[:num_models]
predictions1 = [model(minibatch) for model, minibatch in zip(models, minibatches)]
# Option 2: Same minibatch
minibatch = data[0]
predictions2 = [model(minibatch) for model in models]
# ## Using vmap to vectorize the ensemble
#
# Let's use `vmap` to speed up the for-loop. We must first prepare the models
# for use with `vmap`.
#
# First, let's combine the states of the model together by stacking each parameter.
# For example, `model[i].fc1.weight` has shape `[9216, 128]`; we are going to stack the
# `.fc1.weight` of each of the 10 models to produce a big weight of shape `[10, 9216, 128]`.
#
# functorch offers the following convenience function to do that. It returns a
# stateless version of the model (fmodel) and stacked parameters and buffers.
from functorch import combine_state_for_ensemble
fmodel, params, buffers = combine_state_for_ensemble(models)
[p.requires_grad_() for p in params]
pass
# Option 1: get predictions using a different minibatch for each model.
# By default, vmap maps a function across the first dimension of all inputs to the
# passed-in function. After `combine_state_for_ensemble`, each of of `params`,
# `buffers` have an additional dimension of size `num_models` at the front;
# and `minibatches` has a dimension of size `num_models`.
print([p.size(0) for p in params])
assert minibatches.shape == (num_models, 64, 1, 28, 28)
from functorch import vmap
predictions1_vmap = vmap(fmodel)(params, buffers, minibatches)
assert torch.allclose(predictions1_vmap, torch.stack(predictions1), atol=1e-3, rtol=1e-5)
# Option 2: get predictions using the same minibatch of data
# vmap has an in_dims arg that specify which dimensions to map over.
# Using `None`, we tell vmap we want the same minibatch to apply for all of
# the 10 models.
predictions2_vmap = vmap(fmodel, in_dims=(0, 0, None))(params, buffers, minibatch)
assert torch.allclose(predictions2_vmap, torch.stack(predictions2), atol=1e-3, rtol=1e-5)
# A quick note: there are limitations around what types of functions can be
# transformed by vmap. The best functions to transform are ones that are
# pure functions: a function where the outputs are only determined by the inputs
# that have no side effects (e.g. mutation). vmap is unable to handle mutation of
# arbitrary Python data structures, but it is able to handle many in-place
# PyTorch operations.
# ## Performance
#
# Curious about performance numbers? Here's how the numbers look on my machine with an A100 GPU:
from torch.utils.benchmark import Timer
without_vmap = Timer(
stmt="[model(minibatch) for model, minibatch in zip(models, minibatches)]",
globals=globals())
with_vmap = Timer(
stmt="vmap(fmodel)(params, buffers, minibatches)",
globals=globals())
print(f'Predictions without vmap {without_vmap.timeit(500)}')
print(f'Predictions with vmap {with_vmap.timeit(500)}')
# In general, vectorization with vmap should be faster than running a function in a for-loop. There are some exceptions though, like if we haven't implemented the vmap rule for a particular operation or if the underlying kernels weren't optimized for older hardware. If you see any of these cases, please let us know by opening an issue at our GitHub!
| notebooks/ensembling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import ipymaterialui as m
m.Chip(label=['hoi'])
# +
statusb = m.Chip(label='-')
count = 1
def onclick(e, w, d):
global count
count += 1
statusb.label = f'{count}'
button1 = m.Button(color='primary', variant='contained', children='hoi')
button1.on_event('onClick', onclick)
m.List(children=[button1, statusb])
# -
m.Chip(label=button1)
button1.color='primary'
button1.children='anders'
button1.color='secondary'
switch = m.Switch(checked=True)
switch
switch.checked=False
switch.checked=True
m.Slider(value=10, min=0, max=100)
# +
status = m.Chip(label='-')
tg = m.ToggleButtonGroup(value='bliep', exclusive=True, children=[
m.ToggleButton(value='bla', children='bla'),
m.ToggleButton(value='bliep', children='bliep'),
m.ToggleButton(value='foo', children='foo')
])
def on_t_change(widget, event, data):
status.label=f'{data}'
tg.on_event('onChange', on_t_change)
m.List(children=[tg, status])
# +
status2 = m.Chip(label='-')
display(status2)
def rg_change(widget, event, data):
status2.label=f'{data}'
rg = m.RadioGroup(
name='gender1',
value='male',
children=[
m.FormControlLabel(value='female', control=m.Radio(), label='Female'),
m.FormControlLabel(value='male', control=m.Radio(), label='Male'),
m.FormControlLabel(value='other', control=m.Radio(), label='Other')
]
)
rg.on_event('onChange', rg_change)
m.FormControl(component='fieldset', children=[
m.FormLabel(component='legend'),
rg
])
# +
def do_change(widget, event, data):
status3.label=f'{data}'
sl = m.Slider(value=10)
sl.on_event('onChange', do_change)
status3 = m.Chip(label='-')
display(status3)
m.Grid(container=True,
spacing=2,
children=[
m.Grid(item=True, xs=1, children=m.Icon(children='volume_down')),
m.Grid(item=True, xs=10, children=sl),
m.Grid(item=True, xs=1, children=m.Icon(children='volume_up'))
])
# -
m.Tooltip(
title='Delete',
children=m.Button(children='Hoi'))
m.IconButton( children=m.Icon(children='delete'))
m.Fab(color='primary', children=m.Icon(children='add'))
# +
def toggle_modal(widget, event, data):
modal1.open_ = not modal1.open_
btn3 = m.Button(color='primary', variant='contained', children='Open Modal')
btn3.on_event('onClick', toggle_modal)
modal1 = m.Modal(open_=False, children=m.Button(
style_={'position': 'absolute',
'left': '50%',
'top': '50%'},
color='primary',
variant='contained',
children='button in modal'))
modal1.on_event('onClose', toggle_modal)
m.List(children=[
btn3, modal1
])
# -
btn3.cid
# +
def om_menu_click(widget, event, data):
menu1.open_ = not menu1.open_
btn4 = m.Button(color='primary', variant='contained', children='open menu')
btn4.on_event('onClick', om_menu_click)
def mk_item(i):
item = m.MenuItem(children=i)
item.on_event('onClick', om_menu_click)
return item
items = [mk_item(i) for i in ['Profile', 'My account', 'Logout']]
menu1 = m.Menu(anchor_el=btn4, keep_mounted=True, open_=False, children=items)
menu1.on_event('onClose', om_menu_click)
m.List(children=[btn4, menu1])
# -
m.Checkbox(checked=True)
m.Select(value='20', children=[
m.MenuItem(value=10, children=['Profile']),
m.MenuItem(value=20, children=['My account'])
])
textfield = m.TextField(label='Name', value='', margin='normal')
textfield
datefield = m.TextField(label='Birthday', type='date', value='2017-05-24')
datefield
# +
tabContent = ['Item One', 'Item Two', 'Item Three']
def on_tab_change(widget, even, data):
index = appBar.children[0].value
tabs.children=[appBar, m.Html(tag='div', style_={'padding': '24px'}, children=tabContent[index])]
tab_header = m.Tabs(value=1, children=[
m.Tab(label='Item One'),
m.Tab(label='Item Two'),
m.Tab(label='Item Three'),
])
tab_header.on_event('onChange', on_tab_change)
appBar = m.AppBar(position='static', children=[
tab_header
])
tabs = m.Html(tag='div', children=[
appBar,
m.Html(tag='div', style_={'padding': '24px'}, children='Item Two')
])
tabs
# -
sb = m.Snackbar(open_=True, message=m.Chip(label='hoi'))
sb
sb.open_=False
m.LinearProgress()
| notebooks/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/google-research/football/blob/master/gfootball/colabs/gfootball_example_from_prebuild.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="iCWQ9yUT3W61" colab_type="text"
# # Setup (should take < 100 seconds)
# + id="kB9McZ4THuor" colab_type="code" colab={}
# !apt-get update
# !apt-get install libsdl2-gfx-dev libsdl2-ttf-dev
# 2.8 and 2.9 binary is the same, so we use 2.8 .so file
# !git clone -b v2.9 https://github.com/google-research/football.git
# !mkdir -p football/third_party/gfootball_engine/lib
# !wget https://storage.googleapis.com/gfootball/prebuilt_gameplayfootball_v2.8.so -O football/third_party/gfootball_engine/lib/prebuilt_gameplayfootball.so
# !cd football && GFOOTBALL_USE_PREBUILT_SO=1 python3 -m pip install .
# + [markdown] id="GGCu8kbl21Rx" colab_type="text"
# # Now, you can run it:
# + id="vLg01fIo2lpV" colab_type="code" outputId="13dcac37-3f59-4f1e-<PASSWORD>-fed<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 68}
import gfootball.env as football_env
env = football_env.create_environment(env_name="academy_empty_goal_close", stacked=False, logdir='/tmp/football', write_goal_dumps=False, write_full_episode_dumps=False, render=False)
env.reset()
steps = 0
while True:
obs, rew, done, info = env.step(env.action_space.sample())
steps += 1
if steps % 100 == 0:
print("Step %d Reward: %f" % (steps, rew))
if done:
break
print("Steps: %d Reward: %.2f" % (steps, rew))
| gfootball/colabs/gfootball_example_from_prebuild.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data Preprocessing
# +
#importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import warnings
# ignore all warnings
warnings.filterwarnings('ignore')
# -
#import dataset
dataset = pd.read_csv('iris.data')
# ## Visualizations
sns.set_style("whitegrid");
sns.FacetGrid(dataset, hue="species", size=4) \
.map(plt.scatter, "sepal_length", "sepal_width") \
.add_legend();
#pairplot
sns.set_style("whitegrid");
sns.pairplot(dataset, hue="species", size=3);
# ## Splitting the data into test and train
from sklearn.model_selection import train_test_split
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state =452 )
# ## Applying K Nearest Neighbourhood
# +
#Using KNeighborsClassifier Method of neighbors class to use Nearest Neighbor algorithm
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier(n_neighbors = 5, metric = 'minkowski', p = 2)
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
# Importing metrics for evaluation
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
# Summary of the predictions made by the classifier
print(classification_report(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# Accuracy score
from sklearn.metrics import accuracy_score
print('\nAccuracy is :-',accuracy_score(y_pred,y_test))
| Semester V/Machine Learning (ML) (4659302)/14/ML_PR_14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import rioxarray
import xarray as xr
import numpy as np
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
# +
ds = xr.tutorial.open_dataset('rasm') # use xr.tutorial.load_dataset() for xarray<v0.11.0
ds = ds.rename({'xc': 'lon', 'yc': 'lat'})
ds = ds.rename_vars({'Tair':'temperature'})
spatial_coords = ['lon', 'lat']
for c, coef in zip(spatial_coords, [180, 360]):
ds.coords[c] = (ds.coords[c] + coef) % (coef*2) - coef
ds = ds.transpose(*[x for x in ds.dims if x not in ds.coords.keys()], ...)
ds = ds.sortby(['x', 'y'])
ds['temp2'] = ds['temperature'] + 273.24
ds
# +
def prepare_xdata(ds, spatial_coords=['lon', 'lat']):
for c, coef in zip(spatial_coords, [180, 360]):
ds.coords[c] = (ds.coords[c] + coef) % (coef*2) - coef
dims = list(ds.dims)
ds = ds.sortby(list(ds.dims))
return ds
def get_min_max_coords(coords):
return np.array([coords.min(), coords.max()])
def regrid(xres, yres, data_array, xcoord='lon', ycoord='lat'):
xmin, xmax = get_min_max_coords(data_array.coords[xcoord])
ymin, ymax = get_min_max_coords(data_array.coords[ycoord])
xbins = np.arange(xmin-xres/2, xmax + xres/2, xres)
ybins = np.arange(ymin - ymin/2, ymax + ymin/2, yres)
y = data_array.coords['lat'].values.ravel()
x = data_array.coords['lon'].values.ravel()
# Bin the data onto a 10x10 grid
# Have to reverse x & y due to row-first indexing
zi, yi, xi = np.histogram2d(x, y, bins=[xbins,ybins], weights=data_array.values.flatten(), normed=False)
counts, _, _ = np.histogram2d(x, y, bins=[xbins,ybins])
counts = np.where(np.isclose(counts, np.array([0])), np.nan, counts)
zi = np.where(np.isclose(zi, np.array([0])), np.nan, zi)
zi = zi / counts
#zi = np.ma.masked_invalid(zi)
yshape = yi.size -1
xshape = xi.size -1
return xr.DataArray(zi.reshape( (yshape,
xshape
)),
coords={'lon':yi[:-1],
'lat':xi[:-1]},
dims=['lon', 'lat'],
name=data_array.name)
def interpolate_temporal_dataset(ds, xres, yres , spatial_coords=['lon', 'lat'], time_dim='time'):
ds = prepare_xdata(ds, spatial_coords=spatial_coords)
interp_dataArrays = []
for i in ds.data_vars:
das = []
for idx, group in ds[i].groupby(time_dim):
da = regrid(xres=xres, yres=yres, data_array=group)
da = da.assign_coords({time_dim:idx})
das.append(da)
interp_dataArray = xr.concat(das, dim=time_dim)
interp_dataArrays.append(interp_dataArray)
interp_dataSet = xr.merge(interp_dataArrays)
return interp_dataSet
def interpolate_non_temporal_dataset(ds, xres, yres , spatial_coords=['lon', 'lat']):
ds = prepare_xdata(ds, spatial_coords=spatial_coords)
das = []
for i in ds.data_vars:
da = ds[i]
da = regrid(xres=xres, yres=yres, data_array=da)
das.append(da)
interp_dataSet_non_temporal = xr.merge(das)
return interp_dataSet_non_temporal
if '__main__' == __name__:
interp_dataSet = interpolate_temporal_dataset(ds, xres=5, yres=5 , spatial_coords=['lon', 'lat'], time_dim='time')
# +
fig, ax = plt.subplots(subplot_kw={'projection':ccrs.PlateCarree()})
interp_dataSet['temperature'].isel(time=0).plot(x='lon', y='lat', ax=ax)
ax.coastlines()
ax.gridlines(draw_labels=True)
fig.show()
# +
fig, ax = plt.subplots(subplot_kw={'projection':ccrs.PlateCarree()})
ds['temperature'].isel(time=0).plot(x='lon', y='lat', ax=ax)
ax.coastlines()
ax.gridlines(draw_labels=True)
fig.show()
# -
interp_dataSet.sel({'lon':-30, 'lat':70}, method='nearest')['temperature'].plot()
# +
def dataArray_to_tiff(da, lon_name='lon', lat_name='lat', crs="epsg:4326", pathname='test.tif'):
da = da.rio.set_spatial_dims(lon_name, lat_name, False).rio.write_crs(crs)
if len(interp_dataSet_Tmean.to_array().dims)==3:
transposing_order = [x for x in da.dims if x not in [lon_name, lat_name]]
transposing_order = transposing_order +[lat_name, lon_name]
elif len(interp_dataSet_Tmean.to_array().dims) == 2:
transposing_order = [lat_name, lon_name]
else:
raise('Dimensions must be at most 3, and not below 2')
da = da.transpose(*transposing_order).rio.set_spatial_dims(lon_name, lat_name, False).rio.to_raster(pathname)
# -
dataArray_to_tiff(interp_dataSet.mean('time').to_array())
| scripts/regridding/regridding_by_histogram_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
# # Python for Finance (2nd ed.)
#
# **Mastering Data-Driven Finance**
#
# © Dr. <NAME> | The Python Quants GmbH
#
# <img src="http://hilpisch.com/images/py4fi_2nd_shadow.png" width="300px" align="left">
# # Data Types and Structures
# ## Basic Data Types
# ### Integers
# + uuid="16aa85ac-ab50-480b-bf7c-d49343bbe65a"
a = 10
type(a)
# + uuid="2128044d-a150-4ea6-9aa3-6d6efb93bcf8"
a.bit_length()
# + uuid="19c4eea7-f4b6-48ac-a265-a5b5d61a838f"
a = 100000
a.bit_length()
# + uuid="34c9348e-fc24-4cb3-9f1e-538b517d16b7"
googol = 10 ** 100
googol
# + uuid="55230cf3-5d00-4958-8ae9-2407ae6c968f"
googol.bit_length()
# + uuid="4f0ff5d0-e5aa-4825-8a0c-9db8297d8414"
1 + 4
# + uuid="3b44359c-ea14-4273-b275-f38b946955cb"
1 / 4
# + uuid="1ee18247-5627-481a-8180-ba4ce158ebaf"
type(1 / 4)
# -
# ### Floats
# + uuid="3ba6e56a-a026-4eb7-ad01-0b7593ef5692"
1.6 / 4
# + uuid="37a1e7ff-5949-4f0e-b4de-6ac28ee69e31"
type (1.6 / 4)
# + uuid="82d6e596-d9ec-49d9-85c4-6a8cb33b2a79"
b = 0.35
type(b)
# + uuid="a3f371c1-9a9a-4d19-bfac-3dc119486abb"
b + 0.1
# + uuid="4cf2a3ed-6231-4672-9df1-03c60e3dfe7b"
c = 0.5
c.as_integer_ratio()
# + uuid="af99f61f-afdb-41f8-bbf8-5abd0e7c9890"
b.as_integer_ratio()
# + uuid="5584d3cc-2cdb-4ce9-9ac7-25541dc6e9b8"
import decimal
from decimal import Decimal
# + uuid="40822f81-0da3-4d62-9e31-713d0068de2f"
decimal.getcontext()
# + uuid="775a8fc7-380f-4444-b394-ccdb6d8e6ea9"
d = Decimal(1) / Decimal (11)
d
# + uuid="2a184e95-cc99-4b25-9905-b6d9becb2670"
decimal.getcontext().prec = 4
# + uuid="66cc3ae7-9a70-4612-b64c-a86fd481ef9d"
e = Decimal(1) / Decimal (11)
e
# + uuid="6fa10fc7-4d21-4490-95a1-4a1be00f2c19"
decimal.getcontext().prec = 50
# + uuid="a3515ac1-4029-4645-ade9-094b6e3e87cf"
f = Decimal(1) / Decimal (11)
f
# + uuid="17a5e733-475b-4a49-a320-9b67bd45ce2e"
g = d + e + f
g
# -
# ### Boolean
import keyword
keyword.kwlist
4 > 3
type(4 > 3)
type(False)
4 >= 3
4 < 3
4 <= 3
4 == 3
4 != 3
True and True
True and False
False and False
True or True
True or False
False or False
not True
not False
(4 > 3) and (2 > 3)
(4 == 3) or (2 != 3)
not (4 != 4)
(not (4 != 4)) and (2 == 3)
if 4 > 3:
print('condition true')
i = 0
while i < 4:
print('condition true, i = ', i)
i += 1
int(True)
int(False)
float(True)
float(False)
bool(0)
bool(0.0)
bool(1)
bool(10.5)
bool(-2)
# ### Strings
# + uuid="01840910-c872-4f80-8e26-37d537aae1c5"
t = 'this is a string object'
# + uuid="d961e3c3-59ca-4e85-949d-553727b39477"
t.capitalize()
# + uuid="443fd868-860a-4ea7-a26c-85ad658a2fb7"
t.split()
# + uuid="7ba8fdc2-a567-42a8-b363-847be15d5073"
t.find('string')
# + uuid="8cc71c8c-e447-4272-8b82-74da5f504404"
t.find('Python')
# + uuid="ce57da4c-b95a-417f-a8bd-44f81b28d162"
t.replace(' ', '|')
# + uuid="4a115a34-774e-4544-95eb-95e9ebec36ea"
'http://www.python.org'.strip('htp:/')
# -
# ### Excursion: Printing and String Replacements
print('Python for Finance')
print(t)
i = 0
while i < 4:
print(i)
i += 1
i = 0
while i < 4:
print(i, end='|')
i += 1
'this is an integer %d' % 15
'this is an integer %4d' % 15
'this is an integer %04d' % 15
'this is a float %f' % 15.3456
'this is a float %.2f' % 15.3456
'this is a float %8f' % 15.3456
'this is a float %8.2f' % 15.3456
'this is a float %08.2f' % 15.3456
'this is a string %s' % 'Python'
'this is a string %10s' % 'Python'
'this is an integer {:d}'.format(15)
'this is an integer {:4d}'.format(15)
'this is an integer {:04d}'.format(15)
'this is a float {:f}'.format(15.3456)
'this is a float {:.2f}'.format(15.3456)
'this is a float {:8f}'.format(15.3456)
'this is a float {:8.2f}'.format(15.3456)
'this is a float {:08.2f}'.format(15.3456)
'this is a string {:s}'.format('Python')
'this is a string {:10s}'.format('Python')
i = 0
while i < 4:
print('the number is %d' % i)
i += 1
i = 0
while i < 4:
print('the number is {:d}'.format(i))
i += 1
# ### Excursion: Regular Expressions
# + uuid="8295f915-ee30-44df-896a-539aa6e5287d"
import re
# + uuid="31a001d3-195c-42d3-823b-643eb2be3981"
series = """
'01/18/2014 13:00:00', 100, '1st';
'01/18/2014 13:30:00', 110, '2nd';
'01/18/2014 14:00:00', 120, '3rd'
"""
# + uuid="b62df3db-51da-4bd5-8abe-f536296c756e"
dt = re.compile("'[0-9/:\s]+'") # datetime
# + uuid="02dd3264-3090-4607-a0d9-568d12093158"
result = dt.findall(series)
result
# + uuid="ec54f07e-c507-437e-b5ec-6bd4f6d1e9f8"
from datetime import datetime
pydt = datetime.strptime(result[0].replace("'", ""),
'%m/%d/%Y %H:%M:%S')
pydt
# + uuid="28c72cd0-7efb-43b6-8dae-c48f034c154e"
print(pydt)
# + uuid="d166e305-a9ab-4d28-ac04-ee6016035b88"
print(type(pydt))
# -
# ## Basic Data Structures
# ### Tuples
# + uuid="a84deee3-7f60-459a-bb91-d0cd9c8031aa"
t = (1, 2.5, 'data')
type(t)
# + uuid="9c63a354-9fde-4ded-92f2-bb8ed08ea185"
t = 1, 2.5, 'data'
type(t)
# + uuid="0c6923a3-efab-4cb1-baf7-f31ce725f520"
t[2]
# + uuid="d36fef13-3878-4512-a458-9f5c2dda2cc6"
type(t[2])
# + uuid="d306c24a-5ae0-4395-9fb7-27bf0bb3d1a5"
t.count('data')
# + uuid="10a2d29d-b83d-45bf-aa6b-7d7d028e0fcf"
t.index(1)
# -
# ### Lists
# + uuid="434dd023-faaf-4b7f-953a-69ec45eec4aa"
l = [1, 2.5, 'data']
l[2]
# + uuid="51232410-3f94-4df5-b982-ffcb8438c437"
l = list(t)
l
# + uuid="7fab351f-6bb8-492f-a4be-314dfab9d13b"
type(l)
# + uuid="5f409ddb-67a5-42cc-949c-7b1cc7fb7489"
l.append([4, 3])
l
# + uuid="4e108298-4370-4371-9794-066aff49bfa3"
l.extend([1.0, 1.5, 2.0])
l
# + uuid="9076e4d2-518a-4554-b400-ffc06918a11a"
l.insert(1, 'insert')
l
# + uuid="7049a3a2-3d79-4584-bc57-3f91feb1e941"
l.remove('data')
l
# + uuid="23bf5c19-5dc8-404a-bb5c-bda07c752d89"
p = l.pop(3)
print(l, p)
# + uuid="df15c54e-7500-40d8-9563-ecb305993e6b"
l[2:5]
# -
# ### Excursion: Control Structures
# + uuid="4661533f-3ff0-4699-a17b-cf061e860d8a"
for element in l[2:5]:
print(element ** 2)
# + uuid="c5d97e03-6106-46a9-abb3-0c71cf0b7767"
r = range(0, 8, 1)
r
# + uuid="0ef58c5e-a1d8-438a-bf2f-db598b963187"
type(r)
# + uuid="c1621dce-d184-4331-bd1d-6b6008286c99"
for i in range(2, 5):
print(l[i] ** 2)
# + uuid="474dd7ff-5fdd-429c-a574-48d698919e98"
for i in range(1, 10):
if i % 2 == 0:
print("%d is even" % i)
elif i % 3 == 0:
print("%d is multiple of 3" % i)
else:
print("%d is odd" % i)
# + uuid="d15a40d3-e0c5-444c-83b8-68a08b1066de"
total = 0
while total < 100:
total += 1
print(total)
# + uuid="bcfe427d-ea2c-485a-aace-f2927afefbdd"
m = [i ** 2 for i in range(5)]
m
# -
# ### Excursion: Functional Programming
# + uuid="73fe24f8-e852-4b29-a488-901145b88120"
def f(x):
return x ** 2
f(2)
# + uuid="af1c90bc-8688-4b64-a269-11b81e53b7f2"
def even(x):
return x % 2 == 0
even(3)
# + uuid="6b9746e8-1717-4f33-a9d8-46fbb8d6d114"
list(map(even, range(10)))
# + uuid="b8e6e5a8-704f-4d72-b92c-0c42f15b53e8"
list(map(lambda x: x ** 2, range(10)))
# + uuid="3354de29-f946-4076-a794-ba07969bf3b7"
list(filter(even, range(15)))
# -
# ### Dicts
# + uuid="14981fb1-5636-4c73-93c9-f8e18930cc6b"
d = {
'Name' : '<NAME>',
'Country' : 'Germany',
'Profession' : 'Chancelor',
'Age' : 64
}
type(d)
# + uuid="786249fd-c888-4aa2-9157-973147ff2c2b"
print(d['Name'], d['Age'])
# + uuid="33182240-3b00-4e43-922c-8189296791aa"
d.keys()
# + uuid="98b8e7a4-bac0-4299-9c47-c223269e24d7"
d.values()
# + uuid="48545763-dfe7-4bb2-ade3-d0598498726e"
d.items()
# + uuid="9ed881cc-0eef-4307-b93b-0a728d98d647"
birthday = True
if birthday:
d['Age'] += 1
print(d['Age'])
# + uuid="a2f40c35-348f-4727-87b6-df2e1c5f6f15"
for item in d.items():
print(item)
# + uuid="2dc61967-ece2-48cd-af60-fcf58eb68270"
for value in d.values():
print(type(value))
# -
# ### Sets
# + uuid="7037930a-6fda-493e-8922-2095f50c76e5"
s = set(['u', 'd', 'ud', 'du', 'd', 'du'])
s
# + uuid="c649fb45-81fd-4d46-9ea0-4462203fde81"
t = set(['d', 'dd', 'uu', 'u'])
# + uuid="d592a20e-c8bd-4db4-b0db-f4f30b3ddb9a"
s.union(t)
# + uuid="0c60085c-d1e9-4023-9909-8f8623f1620e"
s.intersection(t)
# + uuid="c2efb482-02dc-41f6-ba00-4bbdccbfa905"
s.difference(t)
# + uuid="ad2a4b31-8cea-41ef-af26-164ba264dd08"
t.difference(s)
# + uuid="1a040fc4-58d0-454d-b21a-59ce4325914d"
s.symmetric_difference(t)
# + uuid="a9173c67-0fff-4313-a1c6-c9f52f911b00"
from random import randint
l = [randint(0, 10) for i in range(1000)]
len(l)
# + uuid="b79db3dd-9264-49c1-8163-ca5ac1dca35d"
l[:20]
# + uuid="918b67ef-4b7e-456c-8133-60de3ae680f6"
s = set(l)
s
# -
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
#
# <a href="http://tpq.io" target="_blank">http://tpq.io</a> | <a href="http://twitter.com/dyjh" target="_blank">@dyjh</a> | <a href="mailto:<EMAIL>"><EMAIL></a>
| code/ch03/03_data_structures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# **Resources**
#
# * [Stanford Dogs Dataset](http://vision.stanford.edu/aditya86/ImageNetDogs/) - subset of ImageNet dataset
# # Imports
import os
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image # show example images
import scipy.io # needed to read .mat files
# # Stanford Dogs Dataset
# Where to download and unpack dataset? Folder will be created if doesn't exist.
dataset_location = './stanford-dogs/'
# **Download and Extract**
def download(url, dest, md5sum):
import os
import urllib
import hashlib
folder, file = os.path.split(dest)
if folder != '':
os.makedirs(folder, exist_ok=True)
if not os.path.isfile(dest):
print('Downloading', file, '...')
urllib.request.urlretrieve(url, dest)
else:
print('Already Exists:', file)
assert hashlib.md5(open(dest, 'rb').read()).hexdigest() == md5sum
download(url='http://vision.stanford.edu/aditya86/ImageNetDogs/images.tar',
dest=os.path.join(dataset_location, 'images.tar'),
md5sum='1bb1f2a596ae7057f99d7d75860002ef')
download(url='http://vision.stanford.edu/aditya86/ImageNetDogs/lists.tar',
dest=os.path.join(dataset_location, 'lists.tar'),
md5sum='edbb9f16854ec66506b5f09b583e0656')
def extract(src, dest):
import os
import tarfile
path, file = os.path.split(src)
extract_path, _ = os.path.splitext(src)
already_extracted = os.path.isdir(dest)
if not already_extracted:
with tarfile.open(src, 'r') as zf:
print('Extracting', file, '...')
zf.extractall(dest)
else:
print('Already Extracted:', file)
assert os.path.isdir(extract_path)
extract(src=os.path.join(dataset_location, 'images.tar'),
dest=os.path.join(dataset_location, 'images'))
extract(src=os.path.join(dataset_location, 'lists.tar'),
dest=os.path.join(dataset_location, 'lists'))
# **Load Train Set**
# Train set is composed of 12000 images split evenly between 120 classes
train_list = scipy.io.loadmat(os.path.join(dataset_location, 'lists/train_list.mat'))
train_files = np.array([item[0][0] for item in train_list['file_list']])
train_labels = train_list['labels'].flatten() - 1 # convert labels [1..120] -> [0..199]
print(train_files.shape)
print(train_files)
print(train_labels.shape)
print(train_labels)
# **Load Test Set**
# Test set is not balanced, each class contains between 48 and 152 images.
test_list = scipy.io.loadmat(os.path.join(dataset_location, 'lists/test_list.mat'))
test_files = [item[0][0] for item in test_list['file_list']]
test_labels = test_list['labels'] -1 # convert labels [1..120] -> [0..199]
plt.hist(test_labels.ravel(), bins=120)
plt.title('Test Set Distribution');
# **Create Human Readable Tags**
idx_to_tags = np.array([x.split('-')[1] for x in sorted(os.listdir('./stanford-dogs/images/Images/'))])
print(idx_to_tags.shape)
print(idx_to_tags[:5])
# **Explore**
# Show image
idx = 1234
img = matplotlib.image.imread(os.path.join(dataset_location, 'images/Images', train_files[idx]))
label = train_labels[idx]
plt.imshow(img)
plt.title(str(label) + ' - ' + idx_to_tags[label]);
# Show raw
idx = 1234
img = matplotlib.image.imread(os.path.join(dataset_location, 'images/Images', train_files[idx]))
print(type(img))
print(img.shape)
print(img.dtype)
print(img[:,:,0]) # red channel [0..255]
# **Save to .npz**
# +
# save_path = os.path.join(dataset_location, 'stanford-dogs-filenames.npz')
# save_path
# +
# np.savez(save_path,
# train_files=train_files,
# train_labels=train_labels,
# test_files=test_files,
# test_labels=test_labels,
# idx_to_tags=idx_to_tags)
# -
# **Load and Save Images**
# Cells below load all images and convert them to (224,224) resolution.
def load_images(folder, files, target_size):
import PIL
images_list = []
for file in files:
img_full_path = os.path.join(folder, file)
img = PIL.Image.open(img_full_path)
img = img.resize(target_size)
if img.mode == 'RGBA':
img = img.convert('RGB')
images_list.append(np.array(img))
return np.array(images_list)
train_images = load_images(folder='./stanford-dogs/images/Images', files=train_files, target_size=(224,224))
test_images = load_images(folder='./stanford-dogs/images/Images', files=test_files, target_size=(224,224))
# Save converted images to .npz file. File will be approx *3.1GB*.
save_path = os.path.join(dataset_location, 'stanford-dogs-images.npz')
save_path
np.savez(save_path,
x_train=train_images,
y_train=train_labels,
x_valid=test_images,
y_valid=test_labels,
idx_to_tags=idx_to_tags)
| Datasets/1430_Stanford_Dogs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
import boto3
import pandas as pd
import numpy as np
from sagemaker import get_execution_role
import sagemaker
import sys
import os
# !pip install xgboost
import data_manipulation as dm
import xgboost as xgb
# ## Setting up session
# +
sm_boto3 = boto3.client('sagemaker')
sess = sagemaker.Session()
region = sess.boto_session.region_name
role = sagemaker.get_execution_role()
bucket = 'lvp-ion-switching'
prefix = 'xgb-lvp-ion-switching'
print('Using bucket ' + bucket)
# -
# ## Loading Data
# +
train_name = "train_clean.csv"
test_name = "test_clean.csv"
train_path = f's3://{bucket}/{train_name}'
test_path = f's3://{bucket}/{test_name}'
all_data = pd.read_csv(train_path)
# test_data = pd.read_csv(test_path)
# +
# dm.plot_signal_and_channels(all_data)
# +
# dm.plot_signal(test_data)
# -
# ## Feature engineering
# Here we are adding the shifted version of the signal variable as feature, with the shift ranging from -9 to +9. As demonstrated in the RandomForest notebook, this results in a much better prediction.
dm.load_shifted_values([all_data], max_shift=10)
all_data.head()
# +
# test_data.head()
# -
# ### Index feature for training data
# This new feature indicates the number of the chunk of 50k samples to which each sample belongs. Since the original data was generated by chunks which were then concatenated, this is helpful for prediction as demonstrated in the RandomForest notebook.
# +
def get_index_mod_int(col, d):
return pd.Series(range(len(col))).floordiv(d)
index_feat = get_index_mod_int(all_data['time'], 50000)
all_data['index_feat'] = index_feat
all_data = all_data.drop(columns=['time'])
# -
# ### Index feature for test data
# For the corresponding feature in the test data, we compute for each 50k chunk of the test data, which 50k chunk of the train data is the closest (by computing the distance induced by the canonical dot product)
all_data.columns
print(sys.getsizeof(all_data))
# ## Format data
# To use the built-in algorithms parameter tuning of sagemaker, the data must:
# * have no missing value
# * be numeric (including categorical variables)
# * be scaled (not always necessary, like here for gradient boosting algorithms)
# * first column should be the target variable
# * CSV File should have no header
previous_columns = [x for x in all_data.columns if x != 'open_channels']
all_data = all_data[['open_channels'] + previous_columns]
all_data.head()
random_permutation = np.random.choice(len(all_data), len(all_data), replace=False)
split_idx = int(0.7 * len(all_data))
train_data = all_data.iloc[random_permutation[:split_idx], :]
val_data = all_data.iloc[random_permutation[split_idx:], :]
train_data.to_csv('train.csv', index=False, header=False)
val_data.to_csv('validation.csv', index=False, header=False)
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train/train.csv')).upload_file('train.csv')
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'validation/validation.csv')).upload_file('validation.csv')
s3_input_train = sagemaker.s3_input(s3_data='s3://{}/{}/train'.format(bucket, prefix), content_type='csv')
s3_input_validation = sagemaker.s3_input(s3_data='s3://{}/{}/validation/'.format(bucket, prefix), content_type='csv')
# # Setup hyperparameter tuning
# +
# %%writefile xgb_model.py
import argparse
import json
import logging
import os
import pandas as pd
import pickle as pkl
from sagemaker_containers import entry_point
from sagemaker_xgboost_container.data_utils import get_dmatrix
from sagemaker_xgboost_container import distributed
import xgboost as xgb
def _xgb_train(params, dtrain, evals, num_boost_round, model_dir, is_master):
"""Run xgb train on arguments given with rabit initialized.
This is our rabit execution function.
:param args_dict: Argument dictionary used to run xgb.train().
:param is_master: True if current node is master host in distributed training, or is running single node training job. Note that rabit_run will include this argument.
"""
booster = xgb.train(params=params, dtrain=dtrain, evals=evals, num_boost_round=num_boost_round)
if is_master:
model_location = model_dir + '/xgboost-model'
pkl.dump(booster, open(model_location, 'wb'))
logging.info("Stored trained model at {}".format(model_location))
if __name__ == '__main__':
parser = argparse.ArgumentParser()
# Hyperparameters are described here. In this simple example we are just including one hyperparameter.
parser.add_argument('--max_depth', type=int,)
parser.add_argument('--eta', type=float)
parser.add_argument('--gamma', type=int)
parser.add_argument('--min_child_weight', type=int)
parser.add_argument('--subsample', type=float)
parser.add_argument('--verbose', type=int)
parser.add_argument('--objective', type=str)
parser.add_argument('--num_round', type=int)
# Sagemaker specific arguments. Defaults are set in the environment variables.
parser.add_argument('--output_data_dir', type=str, default=os.environ['SM_OUTPUT_DATA_DIR'])
parser.add_argument('--model_dir', type=str, default=os.environ['SM_MODEL_DIR'])
parser.add_argument('--train', type=str, default=os.environ['SM_CHANNEL_TRAIN'])
parser.add_argument('--validation', type=str, default=os.environ['SM_CHANNEL_VALIDATION'])
parser.add_argument('--sm_hosts', type=str, default=os.environ['SM_HOSTS'])
parser.add_argument('--sm_current_host', type=str, default=os.environ['SM_CURRENT_HOST'])
args, _ = parser.parse_known_args()
# Get SageMaker host information from runtime environment variables
sm_hosts = json.loads(os.environ['SM_HOSTS'])
sm_current_host = args.sm_current_host
dtrain = get_dmatrix(args.train, 'csv')
dval = get_dmatrix(args.validation, 'csv')
watchlist = [(dtrain, 'train'), (dval, 'validation')] if dval is not None else [(dtrain, 'train')]
train_hp = {
'max_depth': args.max_depth,
'eta': args.eta,
'gamma': args.gamma,
'min_child_weight': args.min_child_weight,
'subsample': args.subsample,
'verbose': args.verbose,
'objective': args.objective}
xgb_train_args = dict(
params=train_hp,
dtrain=dtrain,
evals=watchlist,
num_boost_round=args.num_round,
model_dir=args.model_dir)
if len(sm_hosts) > 1:
# Wait until all hosts are able to find each other
entry_point._wait_hostname_resolution()
# Execute training function after initializing rabit.
distributed.rabit_run(
exec_fun=_xgb_train,
args=xgb_train_args,
include_in_training=(dtrain is not None),
hosts=sm_hosts,
current_host=sm_current_host,
update_rabit_args=True
)
else:
# If single node training, call training method directly.
if dtrain:
xgb_train_args['is_master'] = True
_xgb_train(**xgb_train_args)
else:
raise ValueError("Training channel must have data to train model.")
def model_fn(model_dir):
"""Deserialized and return fitted model.
Note that this should have the same name as the serialized model in the _xgb_train method
"""
model_file = 'xgboost-model'
booster = pkl.load(open(os.path.join(model_dir, model_file), 'rb'))
return booster
# -
nb_classes = len(all_data['open_channels'].value_counts())
print(nb_classes)
# +
hyperparams = {
"max_depth":"5",
"eta":"0.2",
"gamma":"4",
"min_child_weight":"6",
"subsample":"0.7",
"verbose":"1",
"objective":"multi:softmax",
"num_class": str(nb_classes)
"num_round":"100"}
instance_type = "ml.m5.xlarge"
output_path = 's3://{}/{}/{}/output'.format(bucket, prefix, 'liv-ion-xgb')
content_type = "csv"
# +
# Open Source distributed script mode
from sagemaker.session import s3_input, Session
from sagemaker.xgboost.estimator import XGBoost
boto_session = boto3.Session(region_name=region)
session = Session(boto_session=boto_session)
script_path = 'xgb_model.py'
xgb_script_mode_estimator = XGBoost(
entry_point=script_path,
framework_version='0.90-1', # Note: framework_version is mandatory
hyperparameters=hyperparams,
role=role,
train_instance_count=2,
train_instance_type=instance_type,
output_path=output_path)
train_input = s3_input("s3://{}/{}/{}/".format(bucket, prefix, 'train'), content_type=content_type)
validation_input = s3_input("s3://{}/{}/{}/".format(bucket, prefix, 'validation'), content_type=content_type)
# -
xgb_script_mode_estimator.fit({'train': train_input, 'validation': validation_input})
| scripts/XGBoost-Sagemaker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
from pandas.plotting import parallel_coordinates
# +
# teste = pd.read_csv('../data_science/turmas_2015-2017.csv')
# series_teste = (teste['ano'].map(str)+teste['periodo'].map(str))
# teste['periodoano'] = series_teste.astype(int)
# teste.to_csv('../data_science/turmas_new.csv')
# +
dataFrame = pd.read_csv('data_science/turmas_new.csv')
discA = 'CÁLCULO I'
discB = 'CÁLCULO II'
lista_disciplinas = ['CÁLCULO I', 'CÁLCULO II', 'CÁLCULO III']
dataFrame = dataFrame.dropna()
print(dataFrame.columns)
# -
dataFrame[(dataFrame['discente'] == '0120b6290275041d959e36cc17bbf368') &
(dataFrame['nome'] == 'CÁLCULO II')]
# #### Correlação
# correlação entre duas disciplinas
def correlacao(discA, discB):
dataFrame = pd.read_csv('../data_science/turmas_new.csv')
dataFrameA = dataFrame[dataFrame['nome'] == discA]
dataFrameB = dataFrame[dataFrame['nome'] == discB]
# Aprovados no DiscA
dataFrameA = dataFrameA[dataFrameA['descricao'].str.contains('APROVADO')]
series_aprovados = dataFrameA.discente.unique()
df_finalB = pd.DataFrame()
for dis in series_aprovados:
linhas = dataFrameB[dataFrameB['discente'] == dis]
linha = linhas[linhas['periodoano'] == linhas.periodoano.min()]
df_finalB = pd.concat([df_finalB, linha])
# concatenando das tabelas
colunas = ['discente', 'media_final', 'nome']
dataFrameA = dataFrameA[colunas]
df_finalB = df_finalB[colunas]
conc = pd.concat([dataFrameA, df_finalB])
df = pd.crosstab(conc.discente, conc.nome, conc.media_final,aggfunc=np.mean)
df = df.dropna()
df_correlacao = df.corr()
return df_correlacao[discA][discB]
# +
df_retorno = pd.DataFrame(columns=lista_disciplinas)
# matriz de zeros
w, h = len(lista_disciplinas), len(lista_disciplinas)
content = [[0] * w for i in range(h)]
correlacoes = np.array(content, dtype='f')
# calculo das relacões sem repetição
for i in range(0, len(lista_disciplinas)):
for j in range(0, len(lista_disciplinas)):
if i==j:
correlacoes[i][j] = 1
if i<j:
correlacoes[i][j] = correlacao(lista_disciplinas[i], lista_disciplinas[j])
# -
df_retorno = pd.DataFrame(correlacoes,
columns=lista_disciplinas)
df_retorno = df_retorno.set_axis(lista_disciplinas, axis=0, inplace=False)
df_retorno
# df_retorno.set_axis(lista_disciplinas, axis=0, inplace=False)
# #### Coordenadas Paralelas
# + active=""
# dimensions: [{
# range: this.state.range,
# // constraintrange: [3, 6],
# label: this.state.eixos[0],
# values: this.state.valor[0]
# }, {
# range: this.state.range,
# label: this.state.eixos[1],
# values: this.state.valor[1],
# // tickvals: [1.5,3,4.5]
# },...
# -
dataFrame = pd.read_csv('../data_science/turmas_new.csv')
dataFrame.columns
disciplinas = ['CÁLCULO I', 'CÁLCULO II', 'CÁLCULO III']
# +
# Contando reprovações de media_final notnull
df_contagemRep = dataFrame[dataFrame['descricao'].str.contains('REPROVADO')]
df_contagemRep = df_contagemRep[df_contagemRep.media_final.notnull()]
colunas_1 = ['descricao', 'discente', 'media_final', 'id_turma', 'nome']
df_contagemRep = df_contagemRep[colunas_1].drop_duplicates()
# df_contagemRep[df_contagemRep.discente=='f3895ff9fc386821a519db524a8aeaac']
df_contagemRep = df_contagemRep[df_contagemRep['nome'].isin(disciplinas)]
df_contagemRep = df_contagemRep.groupby(['discente']).descricao.count().reset_index()
df_contagemRep.sort_values('descricao', ascending=False).head()
# +
# Aprovados e não foram reprovados
series_Rep = df_contagemRep['discente']
df_NRep = dataFrame[dataFrame['descricao'].str.contains('APROVADO')]
# tirando os reprovados
df_NRep = df_NRep[~df_NRep['discente'].isin(series_Rep)]
df_NRep = df_NRep[df_NRep.media_final.notnull()]
colunas_2 = ['descricao', 'discente', 'media_final', 'id_turma', 'nome']
df_NRep = df_NRep[colunas_2].drop_duplicates()
df_NRep = df_NRep[df_NRep['nome'].isin(disciplinas)]
df_NRep.head()
# +
# junta APROVADOS e REPROVADOS
aprovados = pd.DataFrame();
aprovados['discente'] = df_NRep['discente'];
aprovados['descricao'] = df_NRep['descricao'];
aprovados = aprovados.replace('APROVADO', 0)
aprovados = aprovados.replace('APROVADO POR NOTA', 0)
df_contagem = pd.concat([df_contagemRep, aprovados])
# +
colunas = ['discente', 'nome', 'media_final']
# tirando duplicados e NaN
grafico = dataFrame[colunas].drop_duplicates().dropna()
grafico = grafico[grafico['nome'].isin(disciplinas)]
df_grafico = pd.crosstab(grafico.discente,grafico.nome,grafico.media_final,aggfunc=np.max).reset_index()
df_grafico = pd.merge(df_grafico, df_contagem, on='discente',how='left')
df_grafico= df_grafico.fillna(0)
#eixo = plt.gca()
plt.figure(figsize=(10,10))
df_grafico= df_grafico[['CÁLCULO I', 'CÁLCULO II', 'CÁLCULO III', 'descricao']]
parallel_coordinates(frame=df_grafico.head(300), class_column='descricao',colormap='gist_rainbow')
#eixo.legend_.remove()
# -
np.array(df_grafico['CÁLCULO I'])
# +
import json
response_data = {}
response_data['result'] = 'error'
response_data['message'] = 'Some error message'
# -
# #### Serviços das matérias
codigos = ['ECT2101','ECT2102','ECT2103','ECT2104','ECT2201','ECT2202','ECT2203','ECT2204','ECT2207','ECT2301',
'ECT2303','ECT2304', 'ECT2401', 'ECT2402','ECT2411','ECT2412','ECT2413']
# +
#codigos = dataFrame.codigo.unique()
#codigos
# -
df_disciplinas = pd.read_csv('../data_science/componentes-curriculares-presenciais-ok.csv')
df_disciplinas.columns
df_disciplinas[df_disciplinas['unidade_responsavel'] == 'ESCOLA DE CIÊNCIAS E TECNOLOGIA']
display(df_disciplinas.head())
print()
df_disciplinas.ano_programa.unique()
df_disciplinas[df_disciplinas['unidade_responsavel']=='ESCOLA DE CIÊNCIAS E TECNOLOGIA'].ano_programa.unique()
df_disciplinas[df_disciplinas['unidade_responsavel']=='ESCOLA DE CIÊNCIAS E TECNOLOGIA'].groupby('nome').ano_programa.unique()
# +
df_disciplinas = df_disciplinas[df_disciplinas['unidade_responsavel'] == 'ESCOLA DE CIÊNCIAS E TECNOLOGIA']
df_disciplinas = df_disciplinas[df_disciplinas.codigo.isin(codigos)]
colDisc = ['codigo', 'nome']
df_disciplinas = df_disciplinas[colDisc]
df_disciplinas.shape
# -
df_disciplinas.nome.unique()
periodo1 = ['PRÉ-CÁLCULO','VETORES E GEOMETRIA ANALÍTICA','CÁLCULO I','QUÍMICA GERAL']
periodo2 = ['CÁLCULO II','ÁLGEBRA LINEAR','PROBABILIDADE E ESTATÍSTICA','LÓGICA DE PROGRAMAÇÃO',
'INTRODUÇÃO À FÍSICA CLÁSSICA I']
periodo3 = ['CÁLCULO III','INTRODUÇÃO À FÍSICA CLÁSSICA II','LINGUAGEM DE PROGRAMAÇÃO']
periodo4 = ['COMPUTAÇÃO NUMÉRICA','MECÂNICA DOS SÓLIDOS','MECÂNICA DOS FLUIDOS',
'INTRODUÇÃO À FÍSICA CLÁSSICA III','CIÊNCIA E TECNOLOGIA DOS MATERIAIS']
# +
df_1 = df_disciplinas[df_disciplinas.nome.isin(periodo1)]
df_1['periodo'] = 1
df_2 = df_disciplinas[df_disciplinas.nome.isin(periodo2)]
df_2['periodo'] = 2
df = pd.concat([df_1, df_2])
df_3 = df_disciplinas[df_disciplinas.nome.isin(periodo3)]
df_3['periodo'] = 3
df = pd.concat([df, df_3])
df_4 = df_disciplinas[df_disciplinas.nome.isin(periodo4)]
df_4['periodo'] = 4
df = pd.concat([df, df_4])
# -
df.nome.unique()
df
teste_col = ['-', # PRÉ-CÁLCULO
'-', # CÁLCULO I
'-', # QUÍMICA GERAL
'-', # VETORES E GEOMETRIA ANALÍTICA
'ECT2102',# ÁLGEBRA LINEAR
'ECT2101,ECT2103,ECT2102',# CÁLCULO II
'ECT2103',# PROBABILIDADE E ESTATÍSTICA
'ECT2103',# INTRODUÇÃO À FÍSICA CLÁSSICA I
'-',# LÓGICA DE PROGRAMAÇÃO
'ECT2201',# CÁLCULO III
'ECT2204',# INTRODUÇÃO À FÍSICA CLÁSSICA II
'ECT2203',# LINGUAGEM DE PROGRAMAÇÃO
'ECT2301,ECT2304',# INTRODUÇÃO À FÍSICA CLÁSSICA III
'ECT2303,ECT2103',# COMPUTAÇÃO NUMÉRICA
'ECT2104',# CIÊNCIA E TECNOLOGIA DOS MATERIAIS
'ECT2304,ECT2201',# MECÂNICA DOS FLUIDOS
'ECT2201,ECT2204'# MECÂNICA DOS SÓLIDOS
]
df['pre_requisito'] = teste_col
df = df[['codigo', 'nome', 'periodo', 'pre_requisito']]
df
df.to_csv('data_science/disciplinas_prerequisitos.csv')
# ### Desvio Padrão
def variancia (data):
media = data.mean()
soma = 0
soma += ((data.map(int)-media) ** 2).sum()
variancia = soma / (len(data)-1)
return variancia
# +
df_teste = pd.read_csv('data_science/turmas_2015-2017.csv')
aux = df_teste[df_teste['nome']=='CÁLCULO II'].media_final
print("Desvio de C2:")
print((variancia(aux))**0.5)
# -
# teste
# olhando o ano_programa
ok = pd.read_csv('../data_science/componentes-curriculares-presenciais-ok.csv')
print(ok.columns)
ok[ok['codigo']=='ECT2102']
# olhando a primeira ocorrencia da turma
okTeste = pd.read_csv('../data_science/turmas-2010-2017-ok.csv')
materiaVetores = okTeste[okTeste['id_componente_curricular'] == 2051005]
materiaVetores.ano.unique()
new_table = pd.read_csv('../data_science/disciplinas_prerequisitos.csv')
new_table.columns
# +
vetor = []
for n in new_table.pre_requisito.str.split(','):
x= new_table[new_table['codigo'].isin(n)].nome.values
vetor.append(','.join(tuple(x)))
new_table['nome_prereq'] = vetor
# -
new_table.to_csv('../data_science/disciplinas_prerequisitosnome.csv')
| back/api/back-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import slidexplain as sle
explainer = sle.FunctionExplainer('Raio do circulo')
explainer.display(lambda x, y: x**2 + y**2, boundaries={'x': (-1., 1.), 'y': (-1., 1.)}, target_boundaries=(-2., 2.))
# +
import pandas as pd
df = pd.DataFrame({'a': [1., 5., 7., 2., -1., 9.], 'b': [1., 2., 6., -2., -1., 1.]})
y = 0.2*df['a']**2 + 0.8*df['a'] - 1.2*df['b']
explainer = sle.DataFrameExplainer('Fitting a dataframe.')
explainer.display(df, y, max_degree=2, boundaries={'a': (-1., 1.), 'b': (-1., 1.)}, target_boundaries=(-10., 10.))
# -
df.max()
df.min()['a']
y.min()
| Examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 分析に必要なライブラリをインポート
import pandas as pd
import numpy as np
# 可視化ライブラリのインポート
import matplotlib.pyplot as plt
import seaborn as sns
# データセットを読み込む
df = pd.read_csv("dataset/winequality-red.csv", sep=";")
# ### モデル構築のステップ
# 1. 標準化
# 2. テストデータと訓練データに分割
# 3. 外れ値の除去
# ### モデル概要
# ---
# - 今回は教師あり学習
# -
# 多重共線性があるか分析
#statsmodelsのvifをインポート
from statsmodels.stats.outliers_influence import variance_inflation_factor
# # ↑この続きはあとでやる
# 最適な重回帰モデルをステップワイズ法でモデル選択(AICで評価)
# # ロジスティック回帰モデル
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import cross_validate
# +
# 機械学習用です。
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import train_test_split
# もう一つ、性能の評価用に
from sklearn import metrics
# -
# 教師データの作成
y = df["quality"]
x = df.drop("quality", axis=1)
# データ分割
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.2)
# 機械学習
model = LogisticRegression()
model.fit(x_train, y_train)
# +
# 評価用ライブラリ
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
# 評価
y_pred = model.predict(x_test)
print(classification_report(y_test, y_pred))
print("正解率 = ", accuracy_score(y_test, y_pred))
# -
| analytics/workspace/.ipynb_checkpoints/【現在未使用】04_評価-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CSV to JSON
#
# After finishing to collect data, all files should be converted into JSON format.
# Outputs
OUTPUT_PATHNAME = './world-campus-220212-1.json'
# +
import numpy as np
import pandas as pd
pd.options.display.max_columns = None
import json
import itertools
# Inputs
PRGM_PATHNAME = '../univ-program/program.csv'
UNIV_PATHNAME = '../qs-rankings/univ-with-qs.csv' # With QS Rankings
UNIV_CITY_PATHNAME = '../weather/univ-city.csv' # University - City (for weather)
WTHR_PATHNAME = '../weather/city-weather.json'
CTRY_PATHNAME = '../country/country-stat.csv'
CTRY_CODE_PATHNAME = '../covid19/country-code.csv' # Country - Code (for COVID-19 Graph)
COVD_PATHNAME = '../covid19/covid19-graph.csv'
# +
ctry_mapper = {
'UK': 'United Kingdom',
'United States of America': 'United States'
}
def update_country(df):
# Get columns related to country
cols = [x for x in df.columns if x.endswith('country')]
for col_name, kv in itertools.product(cols, ctry_mapper.items()):
# Convert country
df.loc[df[col_name] == kv[0], col_name] = kv[1]
# Initialize output
output_info = dict()
# -
# ## University (+ City ID + Country Code)
# +
df_univ = pd.read_csv(UNIV_PATHNAME)
print('univ', df_univ.columns)
# Merge University and City ID
df_city = pd.read_csv(UNIV_CITY_PATHNAME)
df_city.rename(columns={
'country': 'univ-country',
'city': 'univ-city',
'CityId': 'city-id'
}, inplace=True)
print('city', df_city.columns)
# Merge University and Country code
df_code = pd.read_csv(CTRY_CODE_PATHNAME)
df_code.rename(columns={'code': 'country-code'}, inplace=True)
print('code', df_code.columns)
# -
# Merge
df_univ = pd.merge(left=df_univ, right=df_city[['sequence', 'univ-city', 'city-id']],
left_on='sequence', right_on='sequence')
df_univ.rename(columns={'univ-city': 'city'}, inplace=True)
# Merge
df_univ = pd.merge(left=df_univ, right=df_code, how='left',
left_on='country', right_on='univ-country')
df_univ.drop(columns=['sequence', 'city', 'univ-country'], inplace=True)
df_univ.tail()
# Set index
df_univ.drop_duplicates(ignore_index=True, inplace=True)
df_univ.set_index('name', inplace=True)
df_univ.head()
update_country(df_univ)
output_info['university'] = json.loads(df_univ.to_json(orient='index'))
# ## Program
df_prgm = pd.read_csv(PRGM_PATHNAME)
update_country(df_prgm)
output_info['program'] = json.loads(df_prgm.to_json(orient='records'))
# ## Weather
df_wthr = pd.read_json(WTHR_PATHNAME)
df_wthr.set_index('cityId', drop=True, inplace=True)
df_wthr.head()
output_info['weather'] = json.loads(df_wthr.to_json(orient='index'))
# ## COVID-19
df_covd = pd.read_csv(COVD_PATHNAME)
df_covd.set_index('label', drop=True, inplace=True)
df_covd.head()
output_info['covid19'] = json.loads(df_covd.to_json(orient='index'))
with open(OUTPUT_PATHNAME, 'w') as f:
json.dump(output_info, f)
| data/json/csv-to-json.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# <table align="center">
# <td align="center"><a target="_blank" href="https://deeplearning.mit.edu">
# <img src="https://deeplearning.mit.edu/files/images/github/icon_mit.png" style="padding-bottom:5px;" />
# Visit MIT Deep Learning</a></td>
# <td align="center"><a target="_blank" href="http://colab.research.google.com/github/lexfridman/mit-deep-learning/blob/master/tutorial_gans/tutorial_gans.ipynb">
# <img src="https://deeplearning.mit.edu/files/images/github/icon_google_colab.png" style="padding-bottom:5px;" />Run in Google Colab</a></td>
# <td align="center"><a target="_blank" href="https://github.com/lexfridman/mit-deep-learning/blob/master/tutorial_gans/tutorial_gans.ipynb">
# <img src="https://deeplearning.mit.edu/files/images/github/icon_github.png" style="padding-bottom:5px;" />View Source on GitHub</a></td>
# <td align="center"><a target="_blank" align="center" href="https://www.youtube.com/watch?v=O5xeyoRL95U&list=PLrAXtmErZgOeiKm4sgNOknGvNjby9efdf">
# <img src="https://deeplearning.mit.edu/files/images/github/icon_youtube.png" style="padding-bottom:5px;" />Watch YouTube Videos</a></td>
# </table>
# # Generative Adversarial Networks (GANs)
#
# This tutorial accompanies lectures of the [MIT Deep Learning](https://deeplearning.mit.edu) series. Acknowledgement to amazing people involved is provided throughout the tutorial and at the end. Introductory lectures on GANs include the following (with more coming soon):
#
# <table>
# <td align="center" style="text-align: center;">
# <a target="_blank" href="https://www.youtube.com/watch?list=PLrAXtmErZgOeiKm4sgNOknGvNjby9efdf&v=O5xeyoRL95U">
# <img src="https://i.imgur.com/FfQVV8q.png" style="padding-bottom:5px;" />
# (Lecture) Deep Learning Basics: Intro and Overview
# </a>
# </td>
# <td align="center" style="text-align: center;">
# <a target="_blank" href="https://www.youtube.com/watch?list=PLrAXtmErZgOeiKm4sgNOknGvNjby9efdf&v=53YvP6gdD7U">
# <img src="https://i.imgur.com/vbNjF3N.png" style="padding-bottom:5px;" />
# (Lecture) Deep Learning State of the Art 2019
# </a>
# </td>
# </table>
#
# Generative Adversarial Networks (GANs) are a framework for training networks optimized for generating new realistic samples from a particular representation. In its simplest form, the training process involves two networks. One network, called the generator, generates new data instances, trying to fool the other network, the discriminator, that classifies images as real or fake. This original form is illustrated as follows (where #6 refers to one of 7 architectures described in the [Deep Learning Basics tutorial](https://github.com/lexfridman/mit-deep-learning/blob/master/tutorial_deep_learning_basics/deep_learning_basics.ipynb)):
#
# <img src="https://i.imgur.com/LweaD1s.png" width="600px">
#
# There are broadly 3 categories of GANs:
#
# 1. **Unsupervised GANs**: The generator network takes random noise as input and produces a photo-realistic image that appears very similar to images that appear in the training dataset. Examples include the [original version of GAN](https://arxiv.org/abs/1406.2661), [DC-GAN](https://arxiv.org/abs/1511.06434), [pg-GAN](https://arxiv.org/abs/1710.10196), etc.
# 3. **Style-Transfer GANs** - Translate images from one domain to another (e.g., from horse to zebra, from sketch to colored images). Examples include [CycleGAN](https://junyanz.github.io/CycleGAN/) and [pix2pix](https://phillipi.github.io/pix2pix/).
# 2. **Conditional GANs** - Jointly learn on features along with images to generate images conditioned on those features (e.g., generating an instance of a particular class). Examples includes [Conditional GAN](https://arxiv.org/abs/1411.1784), [AC-GAN](https://arxiv.org/abs/1610.09585), [Stack-GAN](https://github.com/hanzhanggit/StackGAN), and [BigGAN](https://arxiv.org/abs/1809.11096).
#
# First, we illustrate BigGAN, a state-of-the-art conditional GAN from DeepMind. This illustration is based on the [BigGAN TF Hub Demo](https://colab.research.google.com/github/tensorflow/hub/blob/master/examples/colab/biggan_generation_with_tf_hub.ipynb) and the BigGAN generators on [TF Hub](https://tfhub.dev/deepmind/biggan-256). See the [BigGAN paper on arXiv](https://arxiv.org/abs/1809.11096) [1] for more information about these models.
#
# We'll be adding more parts to this tutorial as additional lectures come out.
# ## Part 1: BigGAN
#
# We recommend that you run this this notebook in the cloud on Google Colab. If you have not done so yet, consider following the setup steps in the [Deep Learning Basics tutorial](https://github.com/lexfridman/mit-deep-learning) and reading the [Deep Learning Basics: Introduction and Overview with TensorFlow](https://medium.com/tensorflow/mit-deep-learning-basics-introduction-and-overview-with-tensorflow-355bcd26baf0) blog post.
# +
# basics
import io
import os
import numpy as np
# deep learning
from scipy.stats import truncnorm
import tensorflow as tf
if tf.__version__.split(".")[0] == '2':
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
import tensorflow_hub as hub
# visualization
from IPython.core.display import HTML
# #!pip install imageio
import imageio
import base64
# check that tensorflow GPU is enabled
tf.test.gpu_device_name() # returns empty string if using CPU
# -
# ### Load BigGAN generator module from TF Hub
# +
# comment out the TF Hub module path you would like to use
# module_path = 'https://tfhub.dev/deepmind/biggan-128/1' # 128x128 BigGAN
# module_path = 'https://tfhub.dev/deepmind/biggan-256/1' # 256x256 BigGAN
module_path = 'https://tfhub.dev/deepmind/biggan-512/1' # 512x512 BigGAN
tf.reset_default_graph()
print('Loading BigGAN module from:', module_path)
module = hub.Module(module_path)
inputs = {k: tf.placeholder(v.dtype, v.get_shape().as_list(), k)
for k, v in module.get_input_info_dict().items()}
output = module(inputs)
# -
# ### Functions for Sampling and Interpolating the Generator
# +
input_z = inputs['z']
input_y = inputs['y']
input_trunc = inputs['truncation']
dim_z = input_z.shape.as_list()[1]
vocab_size = input_y.shape.as_list()[1]
# sample truncated normal distribution based on seed and truncation parameter
def truncated_z_sample(truncation=1., seed=None):
state = None if seed is None else np.random.RandomState(seed)
values = truncnorm.rvs(-2, 2, size=(1, dim_z), random_state=state)
return truncation * values
# convert `index` value to a vector of all zeros except for a 1 at `index`
def one_hot(index, vocab_size=vocab_size):
index = np.asarray(index)
if len(index.shape) == 0: # when it's a scale convert to a vector of size 1
index = np.asarray([index])
assert len(index.shape) == 1
num = index.shape[0]
output = np.zeros((num, vocab_size), dtype=np.float32)
output[np.arange(num), index] = 1
return output
def one_hot_if_needed(label, vocab_size=vocab_size):
label = np.asarray(label)
if len(label.shape) <= 1:
label = one_hot(label, vocab_size)
assert len(label.shape) == 2
return label
# using vectors of noise seeds and category labels, generate images
def sample(sess, noise, label, truncation=1., batch_size=8, vocab_size=vocab_size):
noise = np.asarray(noise)
label = np.asarray(label)
num = noise.shape[0]
if len(label.shape) == 0:
label = np.asarray([label] * num)
if label.shape[0] != num:
raise ValueError('Got # noise samples ({}) != # label samples ({})'
.format(noise.shape[0], label.shape[0]))
label = one_hot_if_needed(label, vocab_size)
ims = []
for batch_start in range(0, num, batch_size):
s = slice(batch_start, min(num, batch_start + batch_size))
feed_dict = {input_z: noise[s], input_y: label[s], input_trunc: truncation}
ims.append(sess.run(output, feed_dict=feed_dict))
ims = np.concatenate(ims, axis=0)
assert ims.shape[0] == num
ims = np.clip(((ims + 1) / 2.0) * 256, 0, 255)
ims = np.uint8(ims)
return ims
def interpolate(a, b, num_interps):
alphas = np.linspace(0, 1, num_interps)
assert a.shape == b.shape, 'A and B must have the same shape to interpolate.'
return np.array([(1-x)*a + x*b for x in alphas])
def interpolate_and_shape(a, b, steps):
interps = interpolate(a, b, steps)
return (interps.transpose(1, 0, *range(2, len(interps.shape))).reshape(steps, -1))
# -
# ### Create a TensorFlow session and initialize variables
initializer = tf.global_variables_initializer()
sess = tf.Session()
sess.run(initializer)
# ### Create video of interpolated BigGAN generator samples
# +
# category options: https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a
category = 947 # mushroom
# important parameter that controls how much variation there is
truncation = 0.2 # reasonable range: [0.02, 1]
seed_count = 10
clip_secs = 36
seed_step = int(100 / seed_count)
interp_frames = int(clip_secs * 30 / seed_count) # interpolation frames
cat1 = category
cat2 = category
all_imgs = []
for i in range(seed_count):
seed1 = i * seed_step # good range for seed is [0, 100]
seed2 = ((i+1) % seed_count) * seed_step
z1, z2 = [truncated_z_sample(truncation, seed) for seed in [seed1, seed2]]
y1, y2 = [one_hot([category]) for category in [cat1, cat2]]
z_interp = interpolate_and_shape(z1, z2, interp_frames)
y_interp = interpolate_and_shape(y1, y2, interp_frames)
imgs = sample(sess, z_interp, y_interp, truncation=truncation)
all_imgs.extend(imgs[:-1])
# save the video for displaying in the next cell, this is way more space efficient than the gif animation
imageio.mimsave('gan.mp4', all_imgs, fps=30)
# -
# %%HTML
<video autoplay loop>
<source src="gan.mp4" type="video/mp4">
</video>
# The above code should generate a 512x512 video version of the following:
#
# 
# # Acknowledgements
#
# The content of this tutorial is based on and inspired by the work of [TensorFlow team](https://www.tensorflow.org) (see their [Colab notebooks](https://www.tensorflow.org/tutorials/)), [Google DeepMind](https://deepmind.com/), our [MIT Human-Centered AI team](https://hcai.mit.edu), and individual pieces referenced in the [MIT Deep Learning](https://deeplearning.mit.edu) course slides.
#
# TF Colab and TF Hub content is copyrighted to The TensorFlow Authors (2018). Licensed under the Apache License, Version 2.0 (the "License"); http://www.apache.org/licenses/LICENSE-2.0
| tutorial_gans/tutorial_gans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas dataframe output from callbacks
import ipywidgets
import pandas as pd
from IPython.display import display, clear_output
data = [['John', 'Mary', 'Bob'], [40, 38, 22], ['London', 'Paris', 'Rome']]
df = pd.DataFrame(data, columns=['name', 'age', 'location'])
# ## IPython.display
#
# Doesn't work in voila. Also displays multiple times if you keep pressing the button, but `clear_output` will remove the widgets too.
# +
def callback1(w):
display(df)
button1 = ipywidgets.Button(description='Run')
button1.on_click(callback1)
button1
# -
# ## HTML widget
#
# Works but without the nice styling of IPython.display.
# +
def callback2(w):
html2.value = df.to_html()
html2 = ipywidgets.HTML()
button2 = ipywidgets.Button(description='Run')
button2.on_click(callback2)
ipywidgets.VBox(children=[button2, html2])
# -
# ## Output widget + Ipython.display
#
# Works nicely ~~but looks like the styling isn't quite the same in voila~~ and voila styling looks the same as notebook.
# +
def callback3(w):
with output3:
clear_output()
display(df)
output3 = ipywidgets.Output()
button3 = ipywidgets.Button(description='Run')
button3.on_click(callback3)
ipywidgets.VBox(children=[button3, output3])
# -
| voila/callback_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Configuring backend
# ### Configuration Files
#
# Like the Jupyter Notebook server, JupyterHub, and other Jupyter interactive
# computing tools, `jupyter-lsp` can be configured via [Python or JSON
# files][notebook-config] in _well-known locations_. You can find out where to put
# them on your system with:
#
# [notebook-config]: https://jupyter-notebook.readthedocs.io/en/stable/config.html
#
# ```bash
# jupyter --paths
# ```
#
# They will be merged from bottom to top, and the directory where you launch your
# `notebook` server wins, making it easy to check in to version control.
# ### Configuration Options
# #### language_servers
#
# `jupyter-lsp` does not come with any Language Servers! However, we will try to
# use [known language servers](./Language%20Servers.ipynb) if they _are_ installed
# and we know about them: you can disable this behavior by configuring
# [autodetect](#autodetect).
#
# If you don't see an implementation for the language server you need, continue
# reading!
#
# > Please consider [contributing your language server spec](./Contributing.ipynb)
# > to `jupyter-lsp`!
# The absolute minimum language server spec requires:
#
# - `argv`, a list of shell tokens to launch the server in `stdio` mode (as
# opposed to `tcp`),
# - the `languages` which the server will respond to, and
# - the schema `version` of the spec (currently only `1`)
#
# ```python
# # ./jupyter_notebook_config.json ---------- unique! -----------
# # | |
# # or e.g. V V
# # $PREFIX/etc/jupyter/jupyter_notebook_config.d/a-language-server-implementation.json
# {
# "LanguageServerManager": {
# "language_servers": {
# "a-language-server-implementation": {
# "version": 1,
# "argv": ["/absolute/path/to/a-language-server", "--stdio"],
# "languages": ["a-language"]
# }
# }
# }
# }
# ```
# A number of other options we hope to use to enrich the user experience are
# available in the [schema][].
#
# [schema]:
# https://github.com/krassowski/jupyterlab-lsp/blob/master/py_src/jupyter_lsp/schema/schema.json
#
# More complex configurations that can't be hard-coded may benefit from the python
# approach:
#
# ```python
# # jupyter_notebook_config.py
# import shutil
#
# # c is a magic, lazy variable
# c.LanguageServerManager.language_servers = {
# "a-language-server-implementation": {
# # if installed as a binary
# "argv": [shutil.which("a-language-server")],
# "languages": ["a-language"],
# "version": 1
# },
# "another-language-implementation": {
# # if run like a script
# "argv": [shutil.which("another-language-interpreter"), "another-language-server"],
# "languages": ["another-language"],
# "version": 1
# }
# }
# ```
# #### nodejs
#
# > default: `None`
#
# An absolute path to your `nodejs` executable. If `None`, `nodejs` will be
# detected in a number of well-known places.
# #### autodetect
#
# > default: `True`
#
# If `True`, `jupyter-lsp` will look for all
# [known language servers](#installing-language-servers). User-configured
# `language_servers` of the same implementation will be preferred over
# `autodetect`ed ones.
# #### node_roots
#
# > default: `[]`
#
# Absolute paths to search for directories named `node_modules`, such as
# `nodejs`-backed language servers. The order is, roughly:
#
# - the folder where `notebook` or `lab` was launched
# - the JupyterLab `staging` folder
# - wherever `conda` puts global node modules
# - wherever some other conventions put it
# #### extra_node_roots
#
# > default: `[]`
#
# Additional places `jupyter-lsp` will look for `node_modules`. These will be
# checked _before_ `node_roots`, and should not contain the trailing
# `node_modules`.
# ### Python `entry_points`
#
# `pip`-installable packages in the same environment as the Jupyter `notebook`
# server can be automatically detected as providing
# [language_servers](#language_servers). These are a little more involved, but
# also more powerful: see more in [Contributing](Contributing.ipynb#Specs).
# Servers configured this way are loaded _before_ those defined in
# [configuration files](#Configuration-Files), so that a user can fine-tune their
# available servers.
| docs/Configuring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from statsmodels.stats.outliers_influence import variance_inflation_factor
plt.rcParams['figure.figsize'] = (8, 8)
df = pd.read_parquet('./data/dataframe.parquet')
df.head()
sns.distplot(df['salary'], color = 'blue')
plt.xlabel('Salary range for players')
plt.ylabel('Count of players')
plt.title('Players\' Salary Distribution')
plt.xticks()
plt.show()
sns.heatmap(df.corr(), annot=True, linewidths=0.5, linecolor='black', cmap='coolwarm')
plt.show()
plt.figure(figsize=(15,8))
plt.title("Salary distribution based on players teams")
x = sns.boxplot(x=df['team'], y=df["salary"])
x.set_xticklabels(x.get_xticklabels(), rotation=90);
plt.figure(figsize=(15,8))
plt.title("Salary distribution based on players positions")
x = sns.boxplot(x=df['position'], y=df["salary"])
x.set_xticklabels(x.get_xticklabels(), rotation=90);
plt.figure(figsize=(15,8))
plt.title("Salary distribution based on players rating")
x = sns.barplot(x=df['rating'], y=df["salary"])
x.set_xticklabels(x.get_xticklabels(), rotation=90);
plt.figure(figsize=(8,8))
plt.title("Salary distribution based on draft round")
x = sns.barplot(x=df['draft_round'], y=df["salary"])
x.set_xticklabels(x.get_xticklabels(), rotation=90);
plt.figure(figsize=(15,8))
plt.title("Salary distribution based on draft position")
x = sns.barplot(x=df['draft_peak'], y=df["salary"])
x.set_xticklabels(x.get_xticklabels(), rotation=90);
plt.figure(figsize=(15,8))
plt.title("Salary distribution based on draft year")
x = sns.barplot(x=df['draft_year'], y=df["salary"])
x.set_xticklabels(x.get_xticklabels(), rotation=90);
plt.figure(figsize=(15,8))
plt.title("Salary distribution based on year born")
x = sns.barplot(x=df['year_born'], y=df["salary"])
x.set_xticklabels(x.get_xticklabels(), rotation=90);
df.info()
plt.figure(figsize=(15,10))
sns.pairplot(df, x_vars=["rating", "height", "weight"], y_vars=["salary"],
height=10, aspect=.8, kind="reg");
plt.xticks()
plt.show()
plt.figure(figsize=(15,10))
sns.pairplot(df, x_vars=["draft_year", "draft_round", "draft_peak"], y_vars=["salary"],
height=10, aspect=.8, kind="reg");
plt.xticks()
plt.show()
plt.figure(figsize=(5,5))
sns.pairplot(df, x_vars=["year_born"], y_vars=["salary"],
height=10, aspect=.8, kind="reg");
plt.xticks()
plt.show()
| 02 - Data Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Claudia
#Fehlermeldung
print("""/Users/info/.virtualenvs/Clava/lib/python3.7/site-packages/requests/__init__.py:91:
RequestsDependencyWarning: urllib3 (1.23) or chardet (2.3.0) doesn’t match a supported version!
RequestsDependencyWarning)""")
# # Michael
# +
#Importe
from bs4 import BeautifulSoup
import requests
import time
import datetime
import pandas as pd
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import Select
import os
# +
#def selenium(startdatum, enddatum):
#definition webdriver (ohne Pfadangabe, da Pfad gespeichert)
driver = webdriver.Chrome()
#definition Website
driver.get("https://www.six-exchange-regulation.com/de/home/publications/management-transactions.html")
#driver.get("https://www.six-exchange-regulation.com/de/home/publications/management-transactions.html#notificationId=T1I2G00075")
#time.sleep(1)
#datum setzen
startdatum = "01012018"
enddatum = "20012018"
datum_von = driver.find_element_by_id('FromTransactionDate')
datum_von.click()
#datum_von.clear() #Code läuft,wenn clear nicht verwendet wird
datum_von.send_keys(startdatum)
datum_bis = driver.find_element_by_name('ToTransactionDate')
datum_bis.click()
datum_bis.clear()
datum_bis.send_keys(enddatum)
#Einfach nur Abschicken
datum_von = driver.find_element_by_id('FromTransactionDate')
datum_von.send_keys(Keys.ENTER)
#time.sleep(5)
#HTML-Inhalt erfassen und übergeben
html = driver.page_source
html
# -
# # Reto
import requests
from bs4 import BeautifulSoup
import pandas as pd
# +
jahre = list(range(2016,2019))
for jahr in jahre:
url = "https://www.atpworldtour.com/en/players/roger-federer/f324/player-activity?year="
url = url+str(jahr)
print(url)
r = requests.get(url)
soup = BeautifulSoup(r.text, 'html.parser')
# -
turniere
# +
federer = []
jahre = list(range(2016,2018))
for jahr in jahre:
url = "https://www.atpworldtour.com/en/players/roger-federer/f324/player-activity?year="
url = url+str(jahr)
print(url)
r = requests.get(url)
soup = BeautifulSoup(r.text, 'html.parser')
turniere = soup.find_all('div', {'class':"activity-tournament-table"}).find_all('a', {'class':'tourney-title'})
daten = soup.find('div', {'class':"activity-tournament-table"}).find_all('span', {'class':'tourney-dates'})
gegner = soup.find('div', {'class':"day-table-name "}).find_all('a', {'class':'mega-player-name'})
for turnier, datum, spieler in zip(turniere, daten, gegner):
turnier = turnier.text
datum = datum.text
gegner = spieler.text
minidict = {'Turnier': turnier,
'Datum': datum,
"Gegner": spieler,
'Saison':str(jahr)}
federer.append(minidict)
# -
federer
| Vertiefungstage5_6/15 Debugging, Text, Server/00 Debugging Examples from Class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# <div class="alert alert-block alert-info">
# <b>Important:</b> This notebook uses <code>ipywidgets</code> that take advantage of the javascript interface in a web browser. The downside is the functionality does not render well on saved notebooks. Run this notebook locally to see the widgets in action.
# </div>
# # Qiskit Jupyter Tools
#
# Qiskit was designed to be used inside of the Jupyter notebook interface. As such it includes many useful routines that take advantage of this platform, and make performing tasks like exploring devices and tracking job progress effortless.
#
# Loading all the qiskit Jupyter goodness is done via:
from qiskit import *
from qiskit_ibm_provider import IBMProvider
import qiskit_ibm_provider.jupyter # This is the where the magic happens (literally).
# ## Table of contents
#
# 1) [IBM Quantum Dashboard](#dashboard)
#
#
# 2) [Backend Details](#details)
#
# To start, load your IBM Quantum account:
provider = IBMProvider()
# ## IBM Quantum Dashboard <a name="dashboard"></a>
#
# Perhaps the most useful Jupyter tool is the `ibm_quantum_dashboard`. This widget consists of a `Devices` tab and a `Jobs` tab. The `Devices` tab provides an overview of all the devices you have access to. The `Jobs` tab automatically tracks and displays information for the jobs submitted in this session.
#
# To start the dashboard you run the Jupyter magic:
# %ibm_quantum_dashboard
# You should now see a small window titled "IBM Quantum Dashboard" in the upper left corner of the notebook. Click on the drop down symbol to see the two tabs. The `Devices` tab may take a few seconds to load as it needs to communicate with the server for device information. The `Jobs` tab should contain no job information as none has been submitted yet.
# ### Getting an Overview of Backends
#
# The `Devices` tab provides an overview of all the backends you have access to. You can use it to compare, for example, the average CNOT error rates. In addition, the number of pending jobs on the devices is continuously being updated along with the operational status.
# ### Automatic Job Tracking
#
# The `Jobs` tab automatically tracks and displays information for the jobs submitted in this session.
#
# Now, let's submit a job to a device to see this in action:
# +
from qiskit_ibm_provider import least_busy
backend = least_busy(
provider.backends(
simulator=False, filters=lambda b: b.configuration().n_qubits >= 5
)
)
qc = QuantumCircuit(2, 2)
qc.h(0)
qc.cx(0, 1)
qc.measure([0, 1], [0, 1])
mapped_circuit = transpile(qc, backend=backend)
job = backend.run(mapped_circuit, shots=1024)
# -
# Click on the `Jobs` tab and you will see that the job has been added to the list of jobs. Its status, queue position (if any), and estimated start time are being automatically tracked and updated. If the job is running, the scheduling mode for the job is also displayed. For example, if the job status is `RUNNING[F]`, that means the job is actively running and was scheduled using a fair-share algorithm. The button to the left of a job ID allows you to cancel the job.
#
# If you want to kill the dashboard you can do so by calling:
# %disable_ibm_quantum_dashboard
# Although the dashboard itself is killed, the underlying framework is still tracking jobs for you and will show this information if loaded once again.
# ## Viewing Backend Details <a name="details"></a>
#
# The IBM Quantum devices contain a large amount of configuration data and properties. This information can be retrieved by calling:
config = backend.configuration()
params = backend.properties()
# However, parsing through this information quickly becomes tedious. Instead, all the information for a single backend can be displayed graphically by just calling the backend instance itself:
backend
# This widget displays all the information about a backend in a single tabbed-window.
# +
from qiskit_ibm_provider.version import __version__
print("qiskit-ibm-provider version: {}".format(__version__))
# +
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
# -
| docs/tutorials/2_jupyter_tools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Batch regression
#
# Carry out regression experiment for a fixed set of predictors and a set of target dates.
# +
import os, sys
from subseasonal_toolkit.utils.notebook_util import isnotebook
if isnotebook():
# Autoreload packages that are modified
# %load_ext autoreload
# %autoreload 2
else:
from argparse import ArgumentParser
# Imports
import numpy as np
import pandas as pd
from sklearn import *
import sys
import json
import subprocess
from datetime import datetime, timedelta
from functools import partial
from multiprocessing import cpu_count
from ttictoc import tic, toc
from subseasonal_data.utils import get_measurement_variable, df_merge, shift_df
from subseasonal_toolkit.utils.general_util import printf
from subseasonal_toolkit.utils.experiments_util import (get_id_name, get_th_name, get_first_year,
get_start_delta, clim_merge)
from subseasonal_toolkit.utils.eval_util import get_target_dates, mean_rmse_to_score, save_metric
from subseasonal_toolkit.utils.fit_and_predict import apply_parallel
from subseasonal_toolkit.utils.models_util import (get_submodel_name, start_logger, log_params, get_forecast_filename,
save_forecasts)
from subseasonal_toolkit.models.perpp.perpp_util import fit_and_predict, years_ago
from subseasonal_data import data_loaders
import re
# +
#
# Specify model parameters
#
model_name = "perpp"
if not isnotebook():
# If notebook run as a script, parse command-line arguments
parser = ArgumentParser()
parser.add_argument("pos_vars",nargs="*") # gt_id and horizon
parser.add_argument('--target_dates', '-t', default="std_contest")
# Number of years to use in training ("all" or integer)
parser.add_argument('--train_years', '-y', default="all")
# Number of month-day combinations on either side of the target combination
# to include when training
# Set to 0 to include only target month-day combo
# Set to "None" to include entire year
parser.add_argument('--margin_in_days', '-m', default="None")
# If True, use cfsv2 ensemble forecast as a feature
parser.add_argument('--use_cfsv2', '-c', default="True")
args, opt = parser.parse_known_args()
# Assign variables
gt_id = get_id_name(args.pos_vars[0]) # "contest_precip" or "contest_tmp2m"
horizon = get_th_name(args.pos_vars[1]) # "34w" or "56w"
target_dates = args.target_dates
train_years = args.train_years
if train_years != "all":
train_years = int(train_years)
if args.margin_in_days == "None":
margin_in_days = None
else:
margin_in_days = int(args.margin_in_days)
use_best_cfsv2 = (args.use_cfsv2 == "True")
else:
# Otherwise, specify arguments interactively
gt_id = "us_precip"
horizon = "56w"
target_dates = "20171114"
train_years = "all"
margin_in_days = None
use_best_cfsv2 = True
#
# Process model parameters
#
if not use_best_cfsv2:
raise ValueError("--use_cfsv2 argument must be True")
# Get list of target date objects
target_date_objs = pd.Series(get_target_dates(date_str=target_dates,horizon=horizon))
# Sort target_date_objs by day of week
target_date_objs = target_date_objs[target_date_objs.dt.weekday.argsort(kind='stable')]
# Identify measurement variable name
measurement_variable = get_measurement_variable(gt_id) # 'tmp2m' or 'precip'
# Column names for gt_col, clim_col and anom_col
gt_col = measurement_variable
clim_col = measurement_variable+"_clim"
anom_col = get_measurement_variable(gt_id)+"_anom" # 'tmp2m_anom' or 'precip_anom'
# Store delta between target date and forecast issuance date
start_delta = timedelta(days=get_start_delta(horizon, gt_id))
# +
#
# Choose regression parameters
#
# Record standard settings of these parameters
base_col = "zeros"
if (gt_id.endswith("tmp2m")) and (horizon == "34w"):
cfsv2_col = 'subx_cfsv2_tmp2m'
x_cols = [
'tmp2m_shift29',
'tmp2m_shift58',
cfsv2_col,
clim_col
]
elif (gt_id.endswith("precip")) and (horizon == "34w"):
cfsv2_col = 'subx_cfsv2_precip'
x_cols = [
'precip_shift29',
'precip_shift58',
cfsv2_col,
clim_col
]
elif (gt_id.endswith("tmp2m")) and (horizon == "56w"):
cfsv2_col = 'subx_cfsv2_tmp2m'
x_cols = [
'tmp2m_shift43',
'tmp2m_shift86',
cfsv2_col,
clim_col
]
elif (gt_id.endswith("precip")) and (horizon == "56w"):
cfsv2_col = 'subx_cfsv2_precip'
x_cols = [
'precip_shift43',
'precip_shift86',
cfsv2_col,
clim_col
]
elif (gt_id.endswith("tmp2m_1.5x1.5")) and (horizon == "34w"):
cfsv2_col = 'iri_cfsv2_tmp2m'
x_cols = [
'tmp2m_shift29',
'tmp2m_shift58',
cfsv2_col,
clim_col
]
elif (gt_id.endswith("precip_1.5x1.5")) and (horizon == "34w"):
cfsv2_col = 'iri_cfsv2_precip'
x_cols = [
'precip_shift29',
'precip_shift58',
cfsv2_col,
clim_col
]
elif (gt_id.endswith("tmp2m_1.5x1.5")) and (horizon == "56w"):
cfsv2_col = 'iri_cfsv2_tmp2m'
x_cols = [
'tmp2m_shift43',
'tmp2m_shift86',
cfsv2_col,
clim_col
]
elif (gt_id.endswith("precip_1.5x1.5")) and (horizon == "56w"):
cfsv2_col = 'iri_cfsv2_precip'
x_cols = [
'precip_shift43',
'precip_shift86',
cfsv2_col,
clim_col
]
group_by_cols = ['lat', 'lon']
# Record submodel names for perpp model
submodel_name = get_submodel_name(
model_name, train_years=train_years, margin_in_days=margin_in_days,
use_cfsv2=use_best_cfsv2)
printf(f"Submodel name {submodel_name}")
if not isnotebook():
# Save output to log file
logger = start_logger(model=model_name,submodel=submodel_name,gt_id=gt_id,
horizon=horizon,target_dates=target_dates)
# Store parameter values in log
params_names = ['gt_id', 'horizon', 'target_dates',
'train_years', 'margin_in_days',
'base_col', 'x_cols', 'group_by_cols', 'use_best_cfsv2'
]
params_values = [eval(param) for param in params_names]
log_params(params_names, params_values)
# +
#
# Load ground truth data
#
printf("Loading ground truth data")
tic()
gt = data_loaders.get_ground_truth(gt_id)[['lat','lon','start_date',gt_col]]
printf(f"elapsed: {toc()}s")
#
# Added shifted ground truth features
#
printf("Adding shifted ground truth features")
lld_data = gt
shifts = [int(re.search(r'\d+$', col).group()) for col in x_cols if col.startswith(gt_col+"_shift")]
tic()
for shift in shifts:
gt_shift = shift_df(gt, shift)
lld_data = df_merge(lld_data, gt_shift, how="right")
printf(f"elapsed: {toc()}s")
#
# Drop rows with empty pred_cols
#
pred_cols = x_cols+[base_col]
exclude_cols = set([clim_col, cfsv2_col, 'zeros'])
lld_data = lld_data.dropna(subset=set(pred_cols) - exclude_cols)
# Add climatology
if clim_col in pred_cols:
printf("Merging in climatology")
tic()
lld_data = clim_merge(lld_data, data_loaders.get_climatology(gt_id))
toc()
# Add zeros
if 'zeros' in pred_cols:
lld_data['zeros'] = 0
# +
#
# Add cfsv2 ensemble forecast as feature
#
printf(f"Forming cfsv2 ensemble forecast...")
shift = 15 if horizon == "34w" else 29
first_lead = 14 if horizon == "34w" else 28
last_lead = 29
suffix = "-us" if gt_id.startswith("us_") else ""
if gt_id.endswith("1.5x1.5"):
prefix = "iri_cfsv2"
suffix += "1_5"
else:
prefix = "subx_cfsv2"
tic(); data = data_loaders.get_forecast(prefix+"-"+gt_id.split("_")[1]+suffix, shift=shift); toc()
printf(f"Aggregating lead {first_lead} with shift {shift}")
tic()
cfsv2 = data[['lat','lon','start_date',f'{prefix}_{gt_col}-{first_lead}.5d_shift{shift}']].set_index(
['lat','lon','start_date']).squeeze().unstack(['lat','lon']).copy()
toc()
for lead in range(first_lead+1,last_lead+1):
printf(f"Aggregating lead {lead} with shift {shift+lead-first_lead}")
tic()
cfsv2 += data[['lat','lon','start_date',f'{prefix}_{gt_col}-{lead}.5d_shift{shift}']].set_index(
['lat','lon','start_date']).squeeze().unstack(['lat','lon']).shift(lead-first_lead)
toc()
del data
num_leads = last_lead - first_lead + 1
cfsv2 /= num_leads
# Drop dates with no forecasts and reshape
cfsv2 = cfsv2.dropna().unstack().rename(cfsv2_col)
# Merge cfsv2 forecast with lld_data
printf(f"Merging {cfsv2_col} with lld_data")
tic()
lld_data = pd.merge(lld_data, cfsv2, left_on=['lat','lon','start_date'],
right_index=True)
toc()
del cfsv2
# +
# specify regression model
fit_intercept = True
model = linear_model.LinearRegression(fit_intercept=fit_intercept)
# Form predictions for each grid point (in parallel) using train / test split
# and the selected model
prediction_func = partial(fit_and_predict, model=model)
num_cores = cpu_count()
# Store rmses
rmses = pd.Series(index=target_date_objs, dtype='float64')
# Restrict data to relevant columns and rows for which predictions can be made
relevant_cols = set(
['start_date','lat','lon',gt_col,base_col]+x_cols).intersection(lld_data.columns)
lld_data = lld_data[relevant_cols].dropna(subset=x_cols+[base_col])
# +
for target_date_obj in target_date_objs:
if not any(lld_data.start_date.isin([target_date_obj])):
printf(f"warning: some features unavailable for target={target_date_obj}; skipping")
continue
target_date_str = datetime.strftime(target_date_obj, '%Y%m%d')
# Skip if forecast already produced for this target
forecast_file = get_forecast_filename(
model=model_name, submodel=submodel_name,
gt_id=gt_id, horizon=horizon,
target_date_str=target_date_str)
if True and os.path.isfile(forecast_file):
printf(f"prior forecast exists for target={target_date_obj}; loading")
tic()
preds = pd.read_hdf(forecast_file)
# Add ground truth for later evaluation
preds = pd.merge(preds, lld_data.loc[lld_data.start_date==target_date_obj,['lat','lon',gt_col]],
on=['lat','lon'])
preds.rename(columns={gt_col:'truth'}, inplace=True)
toc()
else:
printf(f'target={target_date_str}')
# Subset data based on margin
if margin_in_days is not None:
tic()
sub_data = month_day_subset(lld_data, target_date_obj, margin_in_days)
toc()
else:
sub_data = lld_data
# Find the last observable training date for this target
last_train_date = target_date_obj - start_delta
# Only train on train_years worth of data
if train_years != "all":
tic()
sub_data = sub_data.loc[sub_data.start_date >= years_ago(last_train_date, train_years)]
toc()
tic()
preds = apply_parallel(
sub_data.groupby(group_by_cols),
prediction_func,
num_cores=num_cores,
gt_col=gt_col,
x_cols=x_cols,
base_col=base_col,
last_train_date=last_train_date,
test_dates=[target_date_obj])
# Ensure raw precipitation predictions are never less than zero
if gt_id.endswith("precip"):
tic()
preds['pred'] = np.maximum(preds['pred'],0)
toc()
preds = preds.reset_index()
if True:
# Save prediction to file in standard format
save_forecasts(preds.drop(columns=['truth']),
model=model_name, submodel=submodel_name,
gt_id=gt_id, horizon=horizon,
target_date_str=target_date_str)
toc()
# Evaluate and store error
rmse = np.sqrt(np.square(preds.pred - preds.truth).mean())
rmses.loc[target_date_obj] = rmse
print("-rmse: {}, score: {}".format(rmse, mean_rmse_to_score(rmse)))
mean_rmse = rmses.mean()
print("-mean rmse: {}, running score: {}".format(mean_rmse, mean_rmse_to_score(mean_rmse)))
if True:
# Save rmses in standard format
rmses = rmses.reset_index()
rmses.columns = ['start_date','rmse']
save_metric(rmses, model=model_name, submodel=submodel_name, gt_id=gt_id, horizon=horizon, target_dates=target_dates, metric="rmse")
# -
| subseasonal_toolkit/models/perpp/perpp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from database.strategy import Strategy
from database.sec import SEC
from database.market import Market
from transformer.model_transformer import ModelTransformer
from transformer.product_transformer import ProductTransformer
from transformer.predictor_transformer import PredictorTransformer
from preprocessor.model_preprocessor import ModelPreprocessor
from preprocessor.predictor_preprocessor import PredictorPreprocessor
from modeler.modeler import Modeler
from utils.date_utils import DateUtils
import pandas as pd
import matplotlib.pyplot as plt
from datetime import datetime, timedelta, timezone
from tqdm import tqdm
import math
## Loading Constants
start = "2008-01-01"
end = datetime.now().strftime("%Y-%m-%d")
# Loading Databases
strat_db = Strategy("recs")
market = Market()
sec = SEC("sec")
market.connect()
tickers = market.retrieve_data("sp500")
market.close()
reload = False
current_week = int(datetime.now().strftime("%W")) + 1
current_quarter = (datetime.now().month - 1)//3 +1
current_year = datetime.now().year
model_range = range(len(tickers["Symbol"]))
weekly_range = range(current_week - 1,current_week)
weekly_range_final = range(current_week - 15,current_week)
quarterly_range = range(current_quarter-2,current_quarter)
quarterly_fundamental_range = range(current_quarter-1,current_quarter +1)
yearly_fundamental_range = range(current_year-1,current_year)
yearly_range = range(current_year,current_year +1)
ticker_range = range(len(tickers["Symbol"]))
tier_2_yearly_range = range(current_year,current_year+1)
load_quarter = False
refined_regression = []
refined_daily_classification = []
refined_quarterly_classification = []
refined_weekly_classification = []
market.connect()
if reload:
for i in tqdm(ticker_range):
try:
ticker = tickers["Symbol"][i]
if "." in ticker:
ticker = ticker.replace(".","-")
prices = market.retrieve_price_data("prices",ticker)
prices.reset_index(inplace=True)
relev = prices[["date","adjClose"]]
for column in ["date","adjClose"]:
relev.rename(columns={column:"".join(column.split("_")).lower()},inplace=True)
relev["date"] = [datetime.strptime(x.split("T")[0],"%Y-%m-%d") for x in relev["date"]]
relev["ticker"] = ticker
relev.sort_values("date",inplace=True)
## daily transformations
refined_regression.append(relev)
relev_classification = relev.copy()
relev_classification["adjclose"] = [1 if x > 0 else 0 for x in relev_classification["adjclose"].diff()]
refined_daily_classification.append(relev_classification)
## weekly transformations
relev["week"] = [x.week for x in relev["date"]]
relev["year"] = [x.year for x in relev["date"]]
relev["quarter"] = [x.quarter for x in relev["date"]]
relev_weekly_classification = relev.groupby(["year","week"]).mean().reset_index()
relev_weekly_classification["adjclose"] = [1 if x > 0 else 0 for x in relev_weekly_classification["adjclose"].diff()]
relev_weekly_classification["ticker"] = ticker
refined_weekly_classification.append(relev_weekly_classification)
## quarterly transformations
relev_quarterly_classification = relev.groupby(["year","quarter"]).mean().reset_index().drop("week",axis=1)
relev_quarterly_classification["adjclose"] = [1 if x > 0 else 0 for x in relev_quarterly_classification["adjclose"].diff()]
relev_quarterly_classification["ticker"] = ticker
refined_quarterly_classification.append(relev_quarterly_classification)
except Exception as e:
print(str(e),ticker)
market.close()
classification_sets = {"date":refined_daily_classification,
"quarter":refined_quarterly_classification,
"week":refined_weekly_classification}
if reload:
for dataset in classification_sets:
base = pd.concat(classification_sets[dataset])
if dataset == "date":
final = base.pivot_table(index=dataset,values="adjclose",columns="ticker").reset_index()
final = final[final["date"] >= datetime.strptime(start,"%Y-%m-%d")]
else:
final = base.pivot_table(index=["year",dataset],values="adjclose",columns="ticker").reset_index()
final = final[final["year"] >= datetime.strptime(start,"%Y-%m-%d").year]
name = "dataset_{}_classification".format(dataset)
for column in final.columns:
if "_" in column:
final.drop(column,axis=1,inplace=True,errors="ignore")
final.fillna(-99999,inplace=True)
for column in tqdm(final.columns):
if -99999 == final[column].min():
final = final.drop(column,axis=1)
market.drop_table(name)
market.store_data(name,final)
if reload:
base = pd.concat(refined_regression)
final = base.pivot_table(index="date",values="adjclose",columns="ticker").reset_index()
final = final[final["date"] >= datetime.strptime(start,"%Y-%m-%d")]
for column in final.columns:
if "_" in column:
final.drop(column,axis=1,inplace=True,errors="ignore")
final.fillna(-99999,inplace=True)
for column in tqdm(final.columns):
if -99999 == final[column].min():
final.drop(column,axis=1,inplace=True)
market.drop_table("dataset_date_regression")
market.store_data("dataset_date_regression",final)
final["quarter"] = [x.quarter for x in final["date"]]
final["year"] = [x.year for x in final["date"]]
final["week"] = [x.week for x in final["date"]]
for timeframe in ["week","quarter"]:
relev = final.groupby(["year",timeframe]).mean().reset_index()
relev.reset_index(drop=True,inplace=True)
name = "dataset_{}_regression".format(timeframe)
market.drop_table(name)
market.store_data(name,relev)
sims = []
training_days = 100
gap = 90
quarter_gap = int(gap/90)
training_years = 4
training_days = 100
sims = []
sec.connect()
strat_db.connect()
market.connect()
quarterly_classification_data = market.retrieve_data("dataset_quarter_classification")
quarterly_regression_data = market.retrieve_data("dataset_quarter_regression")
price_regression_data = market.retrieve_data("dataset_date_regression")
if load_quarter:
for year in tqdm(yearly_range):
for quarter in tqdm(quarterly_range):
try:
## Setting Up
print(year,quarter)
date_ranges = DateUtils.create_quarterly_training_range_rec(year,quarter,training_years,gap)
training_start,training_end,prediction_start,prediction_end = date_ranges
dates = pd.to_datetime(date_ranges)
quarters = [x.quarter for x in dates]
years = [x.year for x in dates]
print(date_ranges)
### switch to all tickers
for i in model_range:
try:
ticker = tickers.iloc[i]["Symbol"].replace(".","-")
if ticker in quarterly_regression_data.columns:
price = market.retrieve_price_data("prices",ticker)
mt = ModelTransformer(ticker,training_start,training_end,gap)
mr = ModelPreprocessor(ticker)
sp = Modeler(ticker)
prot = ProductTransformer(ticker,prediction_start,prediction_end)
## regression_model
rqpd = mt.quarterly_price_transform(quarterly_regression_data,ticker
,years[0],quarters[0],
years[1],quarters[1],quarter_gap)
qpd = mr.day_trade_preprocess_regression(rqpd.copy(),ticker)
q1c = qpd["X"].columns
rpr = sp.quarterly_model("prices",qpd,str(training_years),str(gap))
## classification_model
cqpd = mt.quarterly_price_transform(quarterly_classification_data,ticker
,years[0],quarters[0],
years[1],quarters[1],quarter_gap)
qpd = mr.day_trade_preprocess_classify(cqpd.copy(),ticker)
q2c = qpd["X"].columns
cpr = sp.classify_wta("prices",qpd,str(training_years),str(gap))
price_results = pd.DataFrame([cpr,rpr])
product_refineds = []
product_qpds = []
for i in range(len(price_results)):
price_result = price_results.iloc[i]
if price_result["classification"] == False:
quarterly_price_data = quarterly_regression_data
else:
quarterly_price_data = quarterly_classification_data
quarterly_price_data = quarterly_price_data[(quarterly_price_data["year"] == years[3]) &
(quarterly_price_data["quarter"] == quarters[3])]
quarterly_price_data["y"] = quarterly_price_data[ticker]
product_qpd = mr.day_trade_preprocess_regression(quarterly_price_data.copy(),ticker)
product_qpds.append(product_qpd)
sim = prot.merge_quarterlies_price(price.copy(),product_qpds,price_results)
strat_db.store_data("quarterly_sim",sim)
except Exception as e:
message = {"status":"quarterly modeling","ticker":ticker,"year":str(year),"quarter":str(quarter),"message":str(e)}
print(message)
except Exception as e:
print(year,quarter,str(e))
sec.close()
market.close()
strat_db.close()
sims = []
training_days = 100
gap = 365
quarter_gap = int(gap/90)
training_years = 4
training_days = 100
sims = []
sec.connect()
strat_db.connect()
market.connect()
quarterly_classification_data = market.retrieve_data("dataset_quarter_classification")
quarterly_regression_data = market.retrieve_data("dataset_quarter_regression")
price_regression_data = market.retrieve_data("dataset_date_regression")
if load_quarter:
for year in tqdm(yearly_fundamental_range):
for quarter in tqdm(quarterly_fundamental_range):
try:
## Setting Up
print(year,quarter)
date_ranges = DateUtils.create_quarterly_training_range_rec(year,quarter,training_years,gap)
training_start,training_end,prediction_start,prediction_end = date_ranges
dates = pd.to_datetime(date_ranges)
quarters = [x.quarter for x in dates]
years = [x.year for x in dates]
### switch to all tickers
for i in model_range:
try:
ticker = tickers.iloc[i]["Symbol"].replace(".","-")
if ticker in quarterly_regression_data.columns:
cik = int(tickers.iloc[i]["CIK"].item())
price = market.retrieve_price_data("prices",ticker)
filing = sec.retrieve_filing_data(cik)
mt = ModelTransformer(ticker,training_start,training_end,gap)
mr = ModelPreprocessor(ticker)
sp = Modeler(ticker)
prot = ProductTransformer(ticker,prediction_start,prediction_end)
## regression_model
rfd = mt.fundamental_merge(price.copy(),filing.copy(),True,classify=False)
refined = mr.fundamental_preprocess(rfd.copy())
rfr = sp.quarterly_model("prices",refined,str(training_years),str(gap))
## classification_model
cfd = mt.fundamental_merge(price.copy(),filing.copy(),True,classify=True)
refined = mr.fundamental_preprocess(cfd.copy())
cfr = sp.classify_wta("prices",refined,str(training_years),str(gap))
fundamental_results = pd.DataFrame([cfr,rfr])
product_refineds = []
product_qpds = []
for i in range(len(fundamental_results)):
fundamental_result = fundamental_results.iloc[i]
if fundamental_result["classification"] == False:
fundamental_data = rfd
else:
fundamental_data = cfd
product_refined = mr.fundamental_preprocess(fundamental_data.copy())
product_refineds.append(product_refined)
sim = prot.merge_quarterlies_fundamental(price.copy(),product_refineds,fundamental_results)
strat_db.store_data("fundamental_quarterly_sim",sim)
except Exception as e:
message = {"status":"quarterly modeling","ticker":ticker,"year":str(year),"quarter":str(quarter),"message":str(e)}
print(message)
except Exception as e:
print(year,quarter,str(e))
sec.close()
market.close()
strat_db.close()
sims = []
test = True
training_days = 100
gap = 5
week_gap = int(gap/5)
training_years = 4
training_days = 100
daily_gap = 2
timeline = DateUtils.create_timeline(start,end)
sims = []
sec.connect()
strat_db.connect()
market.connect()
classification = market.retrieve_data("dataset_week_classification")
regression = market.retrieve_data("dataset_week_regression")
for year in tqdm(yearly_range):
for week in tqdm(weekly_range):
try:
## Setting Up
date_ranges = DateUtils.create_weekly_training_range_rec(timeline,year,week,training_years)
training_start,training_end,prediction_start,prediction_end = date_ranges
dates = pd.to_datetime(date_ranges)
weeks = [x.week for x in dates]
years = [x.year for x in dates]
for i in model_range:
try:
ticker = tickers.iloc[i]["Symbol"].replace(".","-")
if ticker in regression.columns:
price = market.retrieve_price_data("prices",ticker)
mt = ModelTransformer(ticker,training_start,training_end,week_gap)
mr = ModelPreprocessor(ticker)
sp = Modeler(ticker)
prot = ProductTransformer(ticker,prediction_start,prediction_end)
## regression_model
rqpd = mt.weekly_price_transform(regression,ticker
,years[0],weeks[0],
years[1],weeks[1],week_gap)
qpd = mr.day_trade_preprocess_regression(rqpd.copy(),ticker)
q1c = qpd["X"].columns
rpr = sp.quarterly_model("prices",qpd,str(training_years),str(week_gap))
## classification_model
cqpd = mt.weekly_price_transform(classification,ticker
,years[0],weeks[0],
years[1],weeks[1],week_gap)
qpd = mr.day_trade_preprocess_classify(cqpd.copy(),ticker)
q2c = qpd["X"].columns
cpr = sp.classify_wta("prices",qpd,str(training_years),str(week_gap))
price_results = pd.DataFrame([cpr,rpr])
product_qpds = []
for i in range(len(price_results)):
price_result = price_results.iloc[i]
if price_result["classification"] == False:
weekly_price_data = regression
else:
weekly_price_data = classification
weekly_price_data = weekly_price_data[(weekly_price_data["year"] == years[3]) &
(weekly_price_data["week"] == weeks[3])]
weekly_price_data["y"] = weekly_price_data[ticker]
product_qpd = mr.day_trade_preprocess_regression(weekly_price_data.copy(),ticker)
product_qpds.append(product_qpd)
print(week,year,current_week,current_year)
if week == datetime.now().isocalendar()[1] - 1 and year == datetime.now().year:
sim = prot.merge_weeklies_predict(product_qpds,price_results)
strat_db.store_data("prediction_sim",pd.DataFrame([sim]))
else:
sim = prot.merge_weeklies(price.copy(),product_qpds,price_results)
if len(sim) > 0:
strat_db.store_data("weekly_sim",sim)
except Exception as e:
message = {"status":"weekly modeling","ticker":ticker,"year":str(year),"week":str(week),"message":str(e)}
print(message)
except Exception as e:
print(year,week,str(e))
sec.close()
market.close()
strat_db.close()
strat_db.connect()
weekly_sim = strat_db.retrieve_data("weekly_sim")
price_quarterly_sim = strat_db.retrieve_data("quarterly_sim")
fundamental_quarterly_sim = strat_db.retrieve_data("fundamental_quarterly_sim")
strat_db.close()
quarterly_sim = fundamental_quarterly_sim.drop(["_id","adjclose"],axis=1) \
.merge(price_quarterly_sim.drop("_id",axis=1),on=["date","ticker"],how="left")
weekly_sim["passed"] = weekly_sim["wmtd"] > weekly_sim["weekly_price_regression_prediction"]
quarterly_sim_columns = quarterly_sim.columns
weekly_sim_columns = weekly_sim.columns
qsc = [x for x in quarterly_sim_columns if "score" in x]
wsc = [x for x in weekly_sim_columns if "score" in x]
qcc = [x for x in quarterly_sim_columns if "classification_prediction" in x]
wcc = [x for x in weekly_sim_columns if "classification_prediction" in x]
qrc = [x for x in quarterly_sim_columns if "regression_prediction" in x]
wrc = [x for x in weekly_sim_columns if "regression_prediction" in x]
availability = []
sp5 = tickers
end_datetime = datetime.strptime(end,"%Y-%m-%d")
for i in model_range:
ticker = list(sp5["Symbol"])[i]
ticker = ticker.replace(".","-")
td = weekly_sim[(weekly_sim["ticker"]==ticker) & (weekly_sim["date"]<=end_datetime)]
try:
availability.append({"ticker":ticker
,"end_date":end_datetime <= td.iloc[len(td)-1]["date"]
,"days":(td.iloc[len(td)-1]["date"]- td.iloc[0]["date"]).days
,"records":len(td[td["date"] <= end_datetime])
,"pct":math.ceil(len(td[td["date"] <= end_datetime])/(end_datetime - td.iloc[0]["date"]).days * 100)})
except Exception as e:
print(ticker,str(e))
continue
stuff = pd.DataFrame(availability)
sp5_tickers = list(stuff[(stuff["pct"] >= stuff["pct"].mode().item() - 1) | (stuff["pct"] <= stuff["pct"].mode().item() + 1)]["ticker"].values)
sims = []
test = True
gap = 5
week_gap = int(gap/5)
training_years = 1
training_days = 100
timeline = DateUtils.create_timeline(start,end)
sims = []
sec.connect()
strat_db.connect()
market.connect()
for year in tqdm(tier_2_yearly_range):
for week in tqdm(weekly_range_final):
try:
## Setting Up
date_ranges = DateUtils.create_weekly_training_range_rec(timeline,year,week,training_years)
training_start,training_end,prediction_start,prediction_end = date_ranges
print(date_ranges)
for i in range(len(sp5_tickers)):
try:
ticker = sp5_tickers[i].replace(".","-")
quarter_sim = quarterly_sim[(quarterly_sim["ticker"] == ticker) & (quarterly_sim["date"] <= prediction_end) & (quarterly_sim["date"] >= training_start)]
week_sim = weekly_sim[(weekly_sim["ticker"] == ticker) & (weekly_sim["date"] <= prediction_end) & (weekly_sim["date"] >= training_start)]
for col in wrc:
week_sim["{}_delta".format(col)] = (week_sim[col] - week_sim["adjclose"])/week_sim["adjclose"]
for col in qrc:
quarter_sim["{}_delta".format(col)] = (quarter_sim[col] - quarter_sim["adjclose"])/quarter_sim["adjclose"]
sim = week_sim.drop("_id",axis=1).merge(quarter_sim.drop("adjclose",axis=1),on=["ticker","date"],how="left").dropna()
training_set = sim[sim["date"] <= training_end]
prediction_set = sim[sim["date"] >= prediction_start].iloc[:5]
try:
mr = ModelPreprocessor(ticker)
sp = Modeler(ticker)
mt = ModelTransformer(ticker,start,end,week_gap)
## classification_model
ts = mt.trade_signal_transform_classify(training_set.copy())
qpd = mr.trade_signal_preprocess_classify(ts.copy())
price_result = sp.classify_wta("prices",qpd,str(training_years),str(week_gap))
except Exception as e:
message = {"status":"modeling","ticker":ticker,"message":str(e)}
print(message)
try:
if week == datetime.now().isocalendar()[1] - 1 and year == datetime.now().year:
strat_db.connect()
predicts = strat_db.retrieve_data("prediction_sim")
strat_db.close()
current_values = sim.tail(1)
cols = []
for x in current_values.columns:
if "weekly" in x.split("_") or "x" in x.split("_") or "delta" in x.split("_"):
cols.append(x)
current_values.drop(cols,axis=1,inplace=True)
for col in predicts:
current_values[col] = predicts[col]
current_values.drop(["date","ticker","wmtd","adjclose","_id"],axis=1,inplace=True,errors="ignore")
## get adjcloses
adjclose = market.retrieve_price_data("prices",ticker).tail(1)["adjClose"].item()
wmtd = market.retrieve_price_data("prices",ticker).iloc[:5]["adjClose"].max()
current_values["adjclose"] = adjclose
current_values["wmtd"] = wmtd
current_values["passed_x"] = current_values["wmtd"].item() > current_values["weekly_price_regression_prediction"].item()
current_values["quarterly_fundamental_regression_prediction_delta"] = (current_values["quarterly_fundamental_regression_prediction"] - current_values["adjclose"]) / current_values["adjclose"]
current_values["quarterly_price_regression_prediction_delta"] = (current_values["quarterly_price_regression_prediction"] - current_values["adjclose"]) / current_values["adjclose"]
current_values["weekly_price_regression_prediction_delta"] = (current_values["weekly_price_regression_prediction"] - current_values["adjclose"]) / current_values["adjclose"]
classification_predictions = price_result["model"].predict(current_values)
product = current_values
product["trade_signal_prediction"] = classification_predictions[0]
product["trade_signal_prediction_score"] = price_result["score"].item()
product["ticker"] = ticker
product["date"] = datetime.now()
if product.index.size > 0:
strat_db.store_data("recs",product)
else:
weekly_price_data = prediction_set.copy()
weekly_price_data["y"] = weekly_price_data["adjclose"]
product_qpd = mr.trade_signal_preprocess_classify(weekly_price_data.copy())
classification_predictions = price_result["model"].predict(product_qpd["X"])
prediction_set["trade_signal_classification_prediction"] = classification_predictions
prediction_set["trade_signal_classification_score"] = price_result["score"]
historical = prediction_set[["date","ticker"
,"adjclose","weekly_price_regression_prediction"
,"weekly_price_regression_prediction_delta"
,"weekly_price_regression_score","trade_signal_classification_prediction"
,"trade_signal_classification_score"]]
if product.index.size > 0:
strat_db.store_data("sim",historical)
except Exception as e:
message = {"status":"packaging","ticker":ticker,"message":str(e)}
print(message)
except Exception as e:
message = {"status":"weekly modeling","ticker":ticker,"message":str(e)}
print(message)
except Exception as e:
print(year,week,str(e))
sec.close()
market.close()
strat_db.close()
| boiler/unity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `text` plot
#
# This notebook is designed to demonstrate (and so document) how to use the `shap.plots.text` function. It uses a distilled PyTorch BERT model from the transformers package to do sentiment analysis of IMDB movie reviews.
#
# Note that the prediction function we define takes a list of strings and returns a logit value for the positive class.
# +
import shap
import transformers
import nlp
import torch
import numpy as np
import scipy as sp
# load a BERT sentiment analysis model
tokenizer = transformers.DistilBertTokenizerFast.from_pretrained("distilbert-base-uncased")
model = transformers.DistilBertForSequenceClassification.from_pretrained(
"distilbert-base-uncased-finetuned-sst-2-english"
).cuda()
# define a prediction function
def f(x):
tv = torch.tensor([tokenizer.encode(v, padding='max_length', max_length=500, truncation=True) for v in x]).cuda()
outputs = model(tv)[0].detach().cpu().numpy()
scores = (np.exp(outputs).T / np.exp(outputs).sum(-1)).T
val = sp.special.logit(scores[:,1]) # use one vs rest logit units
return val
# build an explainer using a token masker
explainer = shap.Explainer(f, tokenizer)
# explain the model's predictions on IMDB reviews
imdb_train = nlp.load_dataset("imdb")["train"]
shap_values = explainer(imdb_train[:10], fixed_context=1)
# -
# ## Single instance text plot
#
# When we pass a single instance to the text plot we get the importance of each token overlayed on the original text that corresponds to that token. Red regions correspond to parts of the text that increase the output of the model when they are included, while blue regions decrease the output of the model when they are included. In the context of the sentiment analysis model here red corresponds to a more positive review and blue a more negative review.
#
# Note that importance values returned for text models are often hierarchical and follow the structure of the text. Nonlinear interactions between groups of tokens are often saved and can be used during the plotting process. If the Explanation object passed to the text plot has a `.hierarchical_values` attribute, then small groups of tokens with strong non-linear effects among them will be auto-merged together to form coherent chunks. When the `.hierarchical_values` attribute is present it also means that the explainer may not have completely enumerated all possible token perturbations and so has treated chunks of the text as essentially a single unit. This happens since we often want to explain a text model while evaluating it fewer times than the numbers of tokens in the document. Whenever a region of the input text is not split by the explainer, it is show by the text plot as a single unit.
#
# The force plot above the text is designed to provide an overview of how all the parts of the text combine to produce the model's output. See the [force plot]() notebook for more details, but the general structure of the plot is positive red features "pushing" the model output higher while negative blue features "push" the model output lower. The force plot provides much more quantitative information than the text coloring. Hovering over a chuck of text will underline the portion of the force plot that corresponds to that chunk of text, and hovering over a portion of the force plot will underline the corresponding chunk of text.
#
# Note that clicking on any chunk of text will show the sum of the SHAP values attributed to the tokens in that chunk (clicked again will hide the value).
# plot the first sentence's explanation
shap.plots.text(shap_values[3])
# ## Multiple instance text plot
#
# When we pass a multi-row explanation object to the text plot we get the single instance plots for each input instance scaled so they have consistent comparable x-axis and color ranges.
# plot the first sentence's explanation
shap.plots.text(shap_values[:3])
# ## Summarizing text explanations
#
# While plotting several instance-level explanations using the text plot can be very informative, sometime you want global summaries of the impact of tokens over the a large set of instances. See the [Explanation object]() documentation for more details, but you can easily summarize the importance of tokens in a dataset by collapsing a multi-row explanation object over all it's rows (in this case by summing). Doing this treats every text input token type as a feature, so the collapsed Explanation object will have as many columns as there were unique tokens in the orignal multi-row explanation object. If there are hierarchical values present in the Explanation object then any large groups are divided up and each token in the gruop is given an equal share of the overall group importance value.
shap.plots.bar(shap_values.abs.sum(0))
# Note that how you summarize the importance of features can make a big difference. In the plot above the `a` token was very importance both because it had an impact on the model, and because it was very common. Below we instead summize the instances using the `max` function to see the largest impact of a token in any instance.
shap.plots.bar(shap_values.abs.max(0))
# You can also slice out a single token from all the instances by using that token as an input name (note that the gray values to the left of the input names are the original text that the token was generated from).
shap.plots.bar(shap_values[:,"but"])
shap.plots.bar(shap_values[:,"but"])
# ## Text-To-Text Visualization
# +
import numpy as np
from transformers import AutoTokenizer, AutoModelForSeq2SeqLM
import shap
import torch
tokenizer = AutoTokenizer.from_pretrained("Helsinki-NLP/opus-mt-en-es")
model = AutoModelForSeq2SeqLM.from_pretrained("Helsinki-NLP/opus-mt-en-es").cuda()
s=["In this picture, there are four persons: my father, my mother, my brother and my sister."]
explainer = shap.Explainer(model,tokenizer)
shap_values = explainer(s)
# -
# Text-To-Text Visualization contains the input text to the model on the left side and output text on the right side (in the default layout). On hovering over a token on the right (output) side the importance of each input token is overlayed on it, and is signified by the background color of the token. Red regions correspond to parts of the text that increase the output of the model when they are included, while blue regions decrease the output of the model when they are included. The explanation for a particular output token can be anchored by clickling on the output token (it can be un-anchored by clicking again).
#
# Note that similar to the single output plots described above, importance values returned for text models are often hierarchical and follow the structure of the text. Small groups of tokens with strong non-linear effects among them will be auto-merged together to form coherent chunks. Similarly, The explainer may not have completely enumerated all possible token perturbations and so has treated chunks of the text as essentially a single unit. This preprocessing is done for each output token, and the merging behviour can differ for each output token, since the interation effects might be different for each output token. The merged chunks can be viewed by hovering over the input text, once an output token is anchored. All the tokens of a merged chunk are made bold.
#
# Once the ouput text is anchored the input tokens can be clicked on to view the exact shap value (Hovering over input token also brings up a tooltip with the values). Auto merged tokens show the total values divided over the number of tokens in that chunk.
#
# Hovering over the input text shows the SHAP value for that token for each output token. This is again signified by the background color of the output token. This can be anchored by clicking on the input token.
#
# Note: The color scaling for all token (input and output) are consistent and the brightest red is assigned to the maximum SHAP value of input tokens for any output token.
#
# Note: The layout of the two pieces of text can be changed by using the 'Layout' Drop down menu.
shap.plots.text(shap_values)
# <hr>
# Have an idea for more helpful examples? Pull requests that add to this documentation notebook are encouraged!
| notebooks/api_examples/plots/text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using the Sharpe ratio for stock analysis
# <p>An investment may make sense if we expect it to return more money than it costs. But returns are only part of the story because they are risky - there may be a range of possible outcomes. How does one compare different investments that may deliver similar results on average, but exhibit different levels of risks?</p>
# <p>The Sharpe ratio is one method to perform quick portfolio analysis since it compares the expected returns for two investment opportunities and calculates the additional return per unit of risk an investor could obtain by choosing one over the other. In particular, it looks at the difference in returns for two investments and compares the average difference to the standard deviation (as a measure of risk) of this difference. A higher Sharpe ratio means that the reward will be higher for a given amount of risk. It is common to compare a specific opportunity against a benchmark that represents an entire category of investments.</p>
# <p>The Sharpe ratio is usually calculated for a portfolio and uses the risk-free interest rate as benchmark. In this analysis we will analyze stocks and use a stock index as benchmark rather than the risk-free interest rate, e.g, the <a href="https://fred.stlouisfed.org/series/TB3MS">3-month Treasury Bill Rate</a>. </p>
# +
# Importing required modules
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pandas_datareader.data import DataReader #ONLY NEEDED IF YOU'RE PULLING DATA FROM YAHOO
from datetime import date
# %matplotlib inline
# -
# You can read in example data locally by using the cell below, or pull live data from Yahoo Finance if you have installed Pandas datareader
# +
# Reading in example data
#stock_data = pd.read_csv('../dat/stock_data.csv',parse_dates=['Date'],index_col='Date').dropna()
#benchmark_data = pd.read_csv('../dat/benchmark_data.csv',parse_dates=['Date'],index_col='Date').dropna()
# -
# Use the code below if you want to pull data from Yahoo Finance
# +
year = 2018 #can be 2015,2016,2017,2018 because of the 5 year limit from by Yahoo Finance
tickers = ['AAPL','AMZN','FB','GOOG','SSNLF','MCD','TSLA']
start = date(year,1,1)
end = date(year+1,1,1)
stock_data = DataReader(tickers,'yahoo',start,end)['Close'].dropna()
benchmark_data = DataReader('SP500','fred',start,end).dropna()
# -
stock_data.plot(subplots=True,title='Stock Data')
benchmark_data.plot(subplots=True,title='S&P 500')
plt.show()
# The Sharpe Ratio uses the difference in returns between the two investment opportunities under consideration.
#
# However, our data show the historical value of each investment, not the return. To calculate the return, we need to calculate the percentage change in value from one day to the next.
# +
# calculate daily stock_data returns
stock_returns = stock_data.pct_change()
# calculate daily benchmark_data returns
sp_returns = benchmark_data[benchmark_data.columns[0]].pct_change()
# plot the daily returns
sp_returns.plot(title='S&P500 daily pct return')
stock_returns.plot(title='Stocks daily pct return')
plt.show()
# -
# Next, we need to calculate the relative performance of stocks vs. the S&P 500 benchmark. This is calculated as the difference in returns between stock_returns and sp_returns for each day.
# +
# calculate the difference in daily returns
excess_returns = stock_returns.sub(sp_returns,axis=0)
# plot the excess_returns
excess_returns.plot(title='Stock returns relative to S&P500')
plt.show()
# -
# The Sharpe ratio can now be calculated as the ratio of the average of excess_returns and the standard deviation of the excess returns.
#
# <p>The Sharpe Ratio is often <em>annualized</em> by multiplying it by the square root of the number of periods. We have used daily data as input, so we'll use the square root of the number of trading days (5 days, 52 weeks, minus a few holidays): √252</p>
# +
# calculate the mean of excess_returns
avg_excess_return = excess_returns.mean()
avg_excess_return.plot.bar(title = 'Mean Return Difference' )
plt.show()
# calculate the standard deviations
sd_excess_return = excess_returns.std()
sd_excess_return.plot.bar(title = 'StdDev Return Difference: risk measure' )
plt.show()
# +
# calculate the daily sharpe ratio
daily_sharpe_ratio = avg_excess_return.div(sd_excess_return)
# annualize the sharpe ratio
annual_factor = np.sqrt(252)
annual_sharpe_ratio = daily_sharpe_ratio.mul(annual_factor)
# plot the annualized sharpe ratio
annual_sharpe_ratio.plot.bar(title='Annualized Sharpe Ratio: Stocks vs S&P 500')
plt.show()
# -
print('The stock with the highest Sharpe ratio is: '+ str(annual_sharpe_ratio.idxmax()))
# ## Conclusion
#
# The analysis shows which investment you should go for. The Sharpe Ratio provides us the tool to compare different stocks while taking into account the inherent risks when investing in it and helps to make a decision by adjusting the returns by the differences in risk and allows an investor to compare investment opportunities on equal terms, that is, on an 'apples-to-apples' basis.
| src/Sharpe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
a = np.random.choice([0,1], 500, p=[0.3, 0.7])
a
a.mean()
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 8.0
# language: ''
# name: sagemath
# ---
def B(n,f):
var('t')
return sum([binomial(n,p)*f(x=(p/n).n())*(1-t)^(n-p)*t^p for p in srange(n+1)])
f(x)=exp(x);
expand(B(15,f))
plot(f,0,1)+plot(B(100,f),0,1)
g(x)=sin(x*2*pi)
plot(g,0,1)+plot(B(20,g),0,1)
plot(g,0,1)+plot(B(4,g),0,1)
plot(g,0,1)+plot(B(100,g),0,1)
def b(n,p):
var('t')
return binomial(n,p)*(1-t)^(n-p)*t^p
sum([plot(b(5,p),0,1) for p in srange(6)])
| 2_Curso/Laboratorio/SAGE-noteb/IPYNB/APROX/78-APROX-bernstein.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
print("Let´s get started")
# In this notebook, I´m going to explain step by step what I did to get Trump´s twitter data from the first day of his presidency on January 20, 2017, until the 7th of February 2020.
# I´m going to use the tweepy API to extract the twitter data.
#
# - What: Trumps tweets
# - Time: from 07.01.2017 to 20.01.2020
# - How: tweepy AIP
# import the needed libraries
import os
import tweepy as tw
import pandas as pd
# The next step is to get a twitter developer account, this is needed to access the tweets.
# In previous research I created back than an account, therefore I use this access from the past.
#
# - consumer_key = "uYqdnmEfQdo8UCihFdYAKMST0"
# - consumer_secret = "<KEY>"
# - access_token = "<KEY>"
# - access_secret ="<KEY>"
#Twitter credentials for the app
consumer_key= 'uYqdnmEfQdo8UCihFdYAKMST0'
consumer_secret= '<KEY>'
access_token= '<KEY>'
access_token_secret= '<KEY>'
#pass twitter credentials to tweepy
auth = tw.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tw.API(auth, wait_on_rate_limit=True)
# **Search tweet**
# testing the #realDonaldTrump
search_words = "@realDonaldTrump"
date_since = "2020-02-06"
# Collect tweets
tweets = tw.Cursor(api.search,
q=search_words,
lang="en",
since=date_since).items(5)
tweets
type(tweets)
for tweet in tweets:
print("------")
print(tweet.text)
# Collect a list of tweets
[tweet.text for tweet in tweets]
# +
new_search = search_words + " -filter:retweets"
tweets = tw.Cursor(api.search,
q=new_search,
lang="en",
since=date_since).items(5)
users_locs = [[tweet.user.screen_name, tweet.user.location, tweet.text] for tweet in tweets]
users_locs
# -
# create a dataframe so save information
tweet_text = pd.DataFrame(data=users_locs,
columns=['user', "location", "text"])
tweet_text
tweet_text.text[2]
# **Trump example**
#make initial request for most recent tweets (200 is the maximum allowed count)
trumps_tweets = api.user_timeline(screen_name = "@realDonaldTrump",count=10, since_id = "2020-01-06")
trumps_tweets_list = [[tweet.user.screen_name, tweet.created_at, tweet.text] for tweet in trumps_tweets]
trumps_tweets_list
trumps_tweets_df = pd.DataFrame(data=trumps_tweets_list,
columns=['user', "time", "text"])
trumps_tweets_df
trumps_tweets_df.text[0]
# **Problem definition**
#
# - Extract data for a specific **time period**
# - How to access the **full tweeted text** of the sentence
# - Decide if **retweet** should be used or not
# - Need to do **data cleaning**
# - Just **access** to tweets of the **last 7 days**
# ### Problem: Full tweet text
# +
#make initial request
trumps_tweets = api.user_timeline(screen_name = "@realDonaldTrump", count=10,
since_id = "2020-01-06",
tweet_mode="extended")
# filter for specific attributes
trumps_tweets_list = [[tweet.user.screen_name, tweet.created_at, tweet.full_text] for tweet in trumps_tweets]
trumps_tweets_df = pd.DataFrame(data=trumps_tweets_list,
columns=['user', "time", "text"])
trumps_tweets_df
# -
trumps_tweets_df.text[1]
# ### Problem: retweet
#
# Trump tweets a lot (quantify) much, his own created tweets are most interesting.
#
# Other reasons why not retweets:
#
# - smaller amount of tweets
# - own tweets represent more important to him
# - uses twitter havily as communication plattform, retweets a looooottt!!
# - The question is how is emotions influences the US economy, especially his own tweets and retweets shoud have a bigger impact to this question
# **Quantification of his tweet activity**
#
# Therefor, check the frist 10 tweets and count the number of retweets.
nr_of_retweets = 0
for tweet in trumps_tweets_df.text:
word = tweet.split()
if word[0] == "RT":
nr_of_retweets = nr_of_retweets +1
print("Of the first X words are", nr_of_retweets, "retweets.")
end = trumps_tweets_df.time[0]
start = trumps_tweets_df.time[len(trumps_tweets_df.time)-1]
time_tweeted = end - start
print("Days:", time_tweeted.days)
print("Minutes:", round(time_tweeted.seconds/60))
# Therefore I gonna use Tramp´s tweet data without his retweets.
# Create calculation with assumntion of how many tweets he posted in the 3 years.
# 10 tweet per hour * 8 hours a day * 365 for one year * 3 because he is president for 3 years
10 * 8 * 365 * 3
# This amount is less than 100.000 which is the limit of one request per day.
# Therefore I gonna contunie with the retweets and filter later one, maybe I can do some interesting stuff with the retweets...
# ### Problem: access just last 7 days
| Data/tweepy/Get Twitter Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This is the stuff we did on 02/07
astring = "Mike"
whos
number_string = "12.34"
whos
1+number_string
number_string+astring
long_string = """four score and seven years ago
our country"""
print(long_string)
num_list = [1,2,3,4]
whos
num_list[0]
mixed_list = [2, "kids"]
mixed_list[1]
big_list = [num_list , mixed_list]
whos
big_list[1]
del big_list
big_list
whos
num_list
# +
# num_list.clear?
# -
num_list.clear()
whos
num_list = [4,5,6]
whos
whos
num_list
num_list[0] = 100
num_list
my_first_tuple = (1,2,3,4)
whos
my_first_tuple[3]
my_first_tuple[0] = 100
# ### Dictionary
names = {"Dave": "Backus", "Chase": "Colman", "Glenn": "Okun"}
whos
names["Dave"]
print(names)
print(names)
print(names)
integer_dict = {"1": "one", "2": "two", 3: "three"}
integer_dict["2"]
new_names = {"Donald": "DucK", "Mickey": "Mouse", "Donald": "Trump"}
new_names
new_names.add({"Bill": "clinton"})
new_names["bill"] = "clinton"
new_names.update({"ronald": "regan"})
new_names
data = {'Year': [1990, 2000, 2010], 'GDP': [8.95, 12.56, 14.78]}
print(data)
whos
len(number_string)
# +
# len?
# -
type(12.34)
x = 12.34
type(x)
y = str(x)
y
z = float(y)
whos
float("Mike")
x
int(x)
# +
# int?
# -
whos
astring
len(str(list(astring))
| python_fundamentals_one/python_fun_one_020719.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: leukemia
# language: python
# name: leukemia
# ---
import numpy as np
import os
import cv2
import matplotlib.pyplot as plt
from keras.models import load_model
import numpy as np
import pandas as pd
import os
import glob
import cv2
import random
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.optimizers import RMSprop
from keras.layers import Conv2D, MaxPooling2D
from keras.layers import Activation, Dropout, Flatten, Dense
from keras import regularizers
from keras.callbacks import CSVLogger
#from livelossplot import PlotLossesKeras
import os
import numpy as np
#from imgaug import augmenters as iaa
#import cv2
from keras.layers.normalization import BatchNormalization
#import seaborn as sns
import pandas as pd
from keras import initializers
from keras import optimizers
import keras.backend as K
import tensorflow as tf
import operator
val_all_path = r'D:\new_leuk\Leukemia_Work_Revive\dataset\val\all'
val_hem_path = r'D:\new_leuk\Leukemia_Work_Revive\dataset\val\hem'
# +
val_all_list = os.listdir(val_all_path)
#val_all_list.sort()
val_hem_list = os.listdir(val_hem_path)
#val_hem_list.sort()
# -
print('val/all: ', len(val_all_list))
print('val/hem :', len(val_hem_list))
val_all_batch = np.zeros((len(val_all_list), 210, 210, 3), dtype=np.uint8)
val_hem_batch = np.zeros((len(val_hem_list), 210, 210, 3), dtype=np.uint8)
print(val_all_batch.shape, val_hem_batch.shape)
def Read_n_Crop(list_data, batch, path):
i=0
for x in list_data:
image = cv2.imread(os.path.join(path, x))
image = cv2.cvtColor(image,cv2.COLOR_BGR2RGB)
image = crop_center(image, (210,210,3))
batch[i] = image
i+=1
print(type(batch), batch.shape, batch.dtype, batch[0].shape, batch[0].dtype)
return batch
def crop_center(img, bounding):
start = tuple(map(lambda a, da: a//2-da//2, img.shape, bounding))
end = tuple(map(operator.add, start, bounding))
slices = tuple(map(slice, start, end))
return img[slices]
parasite_images=Read_n_Crop(val_all_list, val_all_batch, val_all_path)
uninf_images=Read_n_Crop(val_hem_list, val_hem_batch, val_hem_path)
# +
para_label = np.array([0 for _ in range(len(parasite_images))])
uninf_label = np.array([1 for _ in range(len(uninf_images))])
para_label.shape, uninf_label.shape
# -
x_all = np.concatenate((parasite_images, uninf_images), axis=0)
y_all = np.concatenate((para_label, uninf_label), axis=0)
print(x_all.shape, y_all.shape)
model = None
model = load_model(r"C:\Users\NSU\10x_adam_baseline_vgg.h5", compile = False)
model.summary()
x_all=x_all/255.0
# Make predictions using trained model
y_pred = model.predict(x_all, verbose=1)
print("Predictions: ", y_pred.shape)
y_pred_flat = []
for pred in y_pred:
if pred > 0.5:
y_pred_flat.append(1)
else:
y_pred_flat.append(0)
y_pred_flat = np.array(y_pred_flat)
# +
from sklearn.metrics import confusion_matrix, classification_report
# Classification report
confusion_mtx = confusion_matrix(y_all, y_pred_flat)
print(confusion_mtx)
target_names = ['0', '1']
print(classification_report(y_all, y_pred_flat, target_names=target_names, digits=4))
| scripts/inference/inference.ipynb |