code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Q-learning
#
# This notebook will guide you through implementation of vanilla Q-learning algorithm.
#
# You need to implement QLearningAgent (follow instructions for each method) and use it on a number of tests below.
# +
#XVFB will be launched if you run on a server
import os
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY"))==0:
# !bash ../xvfb start
# %env DISPLAY=:1
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# +
# %%writefile qlearning.py
from collections import defaultdict
import random, math
import numpy as np
class QLearningAgent:
def __init__(self, alpha, epsilon, discount, get_legal_actions):
"""
Q-Learning Agent
based on http://inst.eecs.berkeley.edu/~cs188/sp09/pacman.html
Instance variables you have access to
- self.epsilon (exploration prob)
- self.alpha (learning rate)
- self.discount (discount rate aka gamma)
Functions you should use
- self.get_legal_actions(state) {state, hashable -> list of actions, each is hashable}
which returns legal actions for a state
- self.get_qvalue(state,action)
which returns Q(state,action)
- self.set_qvalue(state,action,value)
which sets Q(state,action) := value
!!!Important!!!
Note: please avoid using self._qValues directly.
There's a special self.get_qvalue/set_qvalue for that.
"""
self.get_legal_actions = get_legal_actions
self._qvalues = defaultdict(lambda: defaultdict(lambda: 0))
self.alpha = alpha
self.epsilon = epsilon
self.discount = discount
def get_qvalue(self, state, action):
""" Returns Q(state,action) """
return self._qvalues[state][action]
def set_qvalue(self,state,action,value):
""" Sets the Qvalue for [state,action] to the given value """
self._qvalues[state][action] = value
#---------------------START OF YOUR CODE---------------------#
def get_value(self, state):
"""
Compute your agent's estimate of V(s) using current q-values
V(s) = max_over_action Q(state,action) over possible actions.
Note: please take into account that q-values can be negative.
"""
possible_actions = self.get_legal_actions(state)
#If there are no legal actions, return 0.0
if len(possible_actions) == 0:
return 0.0
<YOUR CODE HERE>
return value
def update(self, state, action, reward, next_state):
"""
You should do your Q-Value update here:
Q(s,a) := (1 - alpha) * Q(s,a) + alpha * (r + gamma * V(s'))
"""
#agent parameters
gamma = self.discount
learning_rate = self.alpha
<YOUR CODE HERE>
self.set_qvalue(state, action, <YOUR_QVALUE>)
def get_best_action(self, state):
"""
Compute the best action to take in a state (using current q-values).
"""
possible_actions = self.get_legal_actions(state)
#If there are no legal actions, return None
if len(possible_actions) == 0:
return None
<YOUR CODE HERE>
return best_action
def get_action(self, state):
"""
Compute the action to take in the current state, including exploration.
With probability self.epsilon, we should take a random action.
otherwise - the best policy action (self.getPolicy).
Note: To pick randomly from a list, use random.choice(list).
To pick True or False with a given probablity, generate uniform number in [0, 1]
and compare it with your probability
"""
# Pick Action
possible_actions = self.get_legal_actions(state)
action = None
#If there are no legal actions, return None
if len(possible_actions) == 0:
return None
#agent parameters:
epsilon = self.epsilon
<YOUR CODE HERE>
return chosen_action
# -
# ### Try it on taxi
#
# Here we use the qlearning agent on taxi env from openai gym.
# You will need to insert a few agent functions here.
# +
import gym
env = gym.make("Taxi-v2")
n_actions = env.action_space.n
# +
from qlearning import QLearningAgent
agent = QLearningAgent(alpha=0.5, epsilon=0.25, discount=0.99,
get_legal_actions = lambda s: range(n_actions))
# +
def play_and_train(env,agent,t_max=10**4):
"""
This function should
- run a full game, actions given by agent's e-greedy policy
- train agent using agent.update(...) whenever it is possible
- return total reward
"""
total_reward = 0.0
s = env.reset()
for t in range(t_max):
# get agent to pick action given state s.
a = <YOUR CODE>
next_s, r, done, _ = env.step(a)
# train (update) agent for state s
<YOUR CODE HERE>
s = next_s
total_reward +=r
if done: break
return total_reward
# +
from IPython.display import clear_output
rewards = []
for i in range(1000):
rewards.append(play_and_train(env, agent))
agent.epsilon *= 0.99
if i %100 ==0:
clear_output(True)
print('eps =', agent.epsilon, 'mean reward =', np.mean(rewards[-10:]))
plt.plot(rewards)
plt.show()
# -
# ### Submit to Coursera I: Preparation
# from submit import submit_qlearning1
# submit_qlearning1(rewards, <EMAIL>, <TOKEN>)
submit_rewards1 = rewards.copy()
# # Binarized state spaces
#
# Use agent to train efficiently on CartPole-v0.
# This environment has a continuous set of possible states, so you will have to group them into bins somehow.
#
# The simplest way is to use `round(x,n_digits)` (or numpy round) to round real number to a given amount of digits.
#
# The tricky part is to get the n_digits right for each state to train effectively.
#
# Note that you don't need to convert state to integers, but to __tuples__ of any kind of values.
# +
env = gym.make("CartPole-v0")
n_actions = env.action_space.n
print("first state:%s" % (env.reset()))
plt.imshow(env.render('rgb_array'))
# -
# ### Play a few games
#
# We need to estimate observation distributions. To do so, we'll play a few games and record all states.
# +
all_states = []
for _ in range(1000):
all_states.append(env.reset())
done = False
while not done:
s, r, done, _ = env.step(env.action_space.sample())
all_states.append(s)
if done: break
all_states = np.array(all_states)
for obs_i in range(env.observation_space.shape[0]):
plt.hist(all_states[:, obs_i], bins=20)
plt.show()
# -
# ## Binarize environment
from gym.core import ObservationWrapper
class Binarizer(ObservationWrapper):
def _observation(self, state):
#state = <round state to some amount digits.>
#hint: you can do that with round(x,n_digits)
#you will need to pick a different n_digits for each dimension
return tuple(state)
env = Binarizer(gym.make("CartPole-v0"))
# +
all_states = []
for _ in range(1000):
all_states.append(env.reset())
done = False
while not done:
s, r, done, _ = env.step(env.action_space.sample())
all_states.append(s)
if done: break
all_states = np.array(all_states)
for obs_i in range(env.observation_space.shape[0]):
plt.hist(all_states[:,obs_i],bins=20)
plt.show()
# -
# ## Learn binarized policy
#
# Now let's train a policy that uses binarized state space.
#
# __Tips:__
# * If your binarization is too coarse, your agent may fail to find optimal policy. In that case, change binarization.
# * If your binarization is too fine-grained, your agent will take much longer than 1000 steps to converge. You can either increase number of iterations and decrease epsilon decay or change binarization.
# * Having 10^3 ~ 10^4 distinct states is recommended (`len(QLearningAgent._qvalues)`), but not required.
#
agent = QLearningAgent(alpha=0.5, epsilon=0.25, discount=0.99,
getLegalActions = lambda s: range(n_actions))
rewards = []
for i in range(1000):
rewards.append(play_and_train(env,agent))
#OPTIONAL YOUR CODE: adjust epsilon
if i %100 ==0:
clear_output(True)
print('eps =', agent.epsilon, 'mean reward =', np.mean(rewards[-10:]))
plt.plot(rewards)
plt.show()
# ### Submit to Coursera II: Submission
# from submit import submit_qlearning2
# submit_qlearning2(rewards, <EMAIL>, <TOKEN>)
submit_rewards2 = rewards.copy()
from submit import submit_qlearning_all
submit_qlearning_all(submit_rewards1, submit_rewards2, <EMAIL>, <TOKEN>)
| week3_model_free/.ipynb_checkpoints/qlearning-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
from torchtools.core import *
# from torchtools.models import *
# # Torchtools
#
# > Provides customized loss functions, metrics and models for prototyping projects. Mainly geared towards time series data at the moment. Under Construction.
# This file will become your README and also the index of your documentation.
# ## Install
# `pip install torchtools`
# ## How to use
# Provides customized loss functions, metrics and models for prototyping projects. Mainly geared towards time series data at the moment.
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# # MAPSI - TME - Rappels de Proba/stats
# ## I- La planche de Galton (<font color="red"> obligatoire</font>)
# ### I.1- Loi de Bernouilli
# Écrire une fonction `bernouilli: float ->int` qui prend en argument la paramètre $p \in [0,1]$ et qui renvoie aléatoirement $0$ (avec la probabilité $1-p$) ou $1$ (avec la probabilité $p$).
def bernouilli(p):
if(np.random.random(1)>p):
return 1
else:
return 0
# ### I.2- Loi binomiale
# Écrire une fonction `binomiale: int , float -> int` qui prend en argument un entier $n$ et $p \in [0,1]$ et qui renvoie aléatoirement un nimbre tiré selon la distribution ${\cal B}(n,p)$.
def binomiale(n, p):
acc=0
for i in range(n):
acc += bernouilli(p)
return acc
# ### I.3- Histogramme de la loi binomiale
# <img src="tme2_Galton.jpg" title="Planche de Galton" style="float: right;">
#
# Dans cette question, on considère une planche de Galton de hauteur $n$. On rappelle que des bâtons horizontaux (oranges) sont cloués à cette planche comme le montre la figure ci-contre.
#
# Des billes bleues tombent du haut de la planche et, à chaque niveau, se retrouvent à la verticale d'un des bâtons. Elles vont alors tomber soit à gauche, soit à droite du bâton, jusqu'à atteindre le bas de la planche. Ce dernier est constitué de petites boites dont les bords sont symbolisés par les lignes verticales grises.
#
# Chaque boite renferme des billes qui sont passées exactement le même nombre de fois à droite des bâtons oranges. Par exemple, la boite la plus à gauche renferme les billes qui ne sont jamais passées à droite d'un bâton, celle juste à sa droite renferme les billes passées une seule fois à droite d'un bâton et toutes les autres fois à gauche, et ainsi de suite.
#
# La répartition des billes dans les boites suit donc une loi binomiale ${\cal B}(n,0.5)$.
#
# Écrire un script qui crée un tableau de $1000$ cases dont le contenu correspond à $1000$ instanciations de la loi binomiale ${\cal B}(n,0.5)$. Afin de voir la répartition des billes dans la planche de Galton, tracer l'histogramme de ce tableau. Vous pourrez utiliser la fonction hist de matplotlib.pyplot:
# +
import matplotlib.pyplot as plt
nb_billes = 10000
nb_etape = 100
proba = 0.5
tab = np.array([])
for i in range(nb_billes):
tab = np.append(tab, binomiale(nb_etape, proba))
plt.hist(tab, nb_etape, width=1.1)
plt.title("Histogramme répartition des billets dans la planche de Galton")
plt.show()
# -
# Pour le nombre de bins, calculez le nombre de valeurs différentes dans votre tableau.
nb_unique = np.size(np.unique(tab))
print("Nombre de barres de l'histogramme:", nb_unique)
# ## II- Visualisation d'indépendances (<font color="red"> obligatoire</font>)
# ### II.1- Loi normale centrée réduite
# <img src="tme2_normale.jpg" title="<NAME>" style="float: right;">
#
# On souhaite visualiser la fonction de densité de la loi normale. Pour cela, on va créer un ensemble de $k$ points $(x_i,y_i$), pour des $x_i$ équi-espacés variant de $-2σ$ à $2σ$, les $y_i$ correspondant à la valeur de la fonction de densité de la loi normale centrée de variance $σ^2$, autrement dit ${\cal N}(0,σ^2)$.
#
# Écrire une fonction `normale : int , float -> float np.array` qui, étant donné un paramètre entier `k` impair et un paramètre réel `sigma` renvoie l'`array numpy` des $k$ valeurs $y_i$. Afin que l'`array numpy` soit bien symmétrique, on lèvera une exception si $k$ est pair.
# +
import random
import math
def normale(k, sigma):
if(k%2==0):
raise ValueError("le nombre k doit etre impair")
else:
res = np.zeros(shape=(k, 2))
x = np.linspace(-2*sigma, 2*sigma, k)
for i in range(k):
x_val = x[i]
y = (np.exp(-0.5*(x_val/sigma)**2))/(sigma*np.sqrt(2*np.pi))
# print("x:",x)
# print("y:",y)
res[i][0] = x_val
res[i][1] = y
return res
# -
# Vérfier la validité de votre fonction en affichant grâce à la fonction plot les points générés dans une figure.
# +
nb_points = 2001
sigma = 10
points = normale(nb_points, sigma)
plt.plot(points[:,0], points[:,1])
plt.title("Loi normale entre -2 sigma et +2 sigma")
plt.show()
# -
# ### II.2- Distribution de probabilité affine
# <img src="tme2_lineaire.jpg" title="Distribution affine" style="float: right;">
#
# Dans cette question, on considère une généralisation de la distribution uniforme: une distribution affine, c'est-à-dire que la fonction de densité est une droite, mais pas forcément horizontale, comme le montre la figure ci-contre.
#
# Écrire une fonction `proba_affine : int , float -> float np.array` qui, comme dans la question précédente, va générer un ensemble de $k$ points $y_i, i=0,...,k−1$, représentant cette distribution (paramétrée par sa pente `slope`). On vérifiera ici aussi que l'entier $k$ est impair. Si la pente est égale à $0$, c'est-à-dire si la distribution est uniforme, chaque point $y_i$ devrait être égal à $\frac{1}{k}$ (afin que $\sum y_i=1$). Si la pente est différente de $0$, il suffit de choisir, $\forall i=0,...,k−1$,
#
# $$y_i=\frac{1}{k}+(i−\frac{k−1}{2})×slope$$
#
# Vous pourrez aisément vérifier que, ici aussi, $\sum y_i=1$. Afin que la distribution soit toujours positive (c'est quand même un minimum pour une distribution de probabilité), il faut que la pente slope ne soit ni trop grande ni trop petite. Le bout de code ci-dessous lèvera une exception si la pente est trop élevée et indiquera la pente maximale possible.
def proba_affine(k, slope):
if k % 2 == 0:
raise ValueError("le nombre k doit etre impair")
if abs ( slope ) > 2. / ( k * k ):
raise ValueError("la pente est trop raide : pente max = " +
str ( 2. / ( k * k ) ) )
res = np.zeros(shape=(k, 2))
for i in range(k):
res[i][0] = i
res[i][1] = (1/k) + (i - ((k-1)/2)) * slope
return res
nb_points = 101
slope = 1e-4
points = proba_affine(nb_points, slope)
plt.plot(points[:,0], points[:,1])
plt.title("Courbe distribution de probabilité affine")
plt.show()
# ### II.3- Distribution jointe
# Écrire une fonction `Pxy : float np.array , float np.array -> float np.2D-array` qui, étant donné deux tableaux numpy de nombres réels à $1$ dimension générés par les fonctions des questions précédentes et représentant deux distributions de probabilités $P(A)$ et $P(B)$, renvoie la distribution jointe $P(A,B)$ sous forme d'un tableau numpy à $2$ dimensions de nombres réels, en supposant que $A$ et $B$ sont des variables aléatoires indépendantes. Par exemple, si:
PA = np.array ( [0.2, 0.7, 0.1] )
PB = np.array ( [0.4, 0.4, 0.2] )
# alors `Pxy(A,B)` renverra le tableau :
# ```
# np.array([[ 0.08, 0.08, 0.04],
# [ 0.28, 0.28, 0.14],
# [ 0.04, 0.04, 0.02]])
# ```
def Pxy(x,y):
n1 = np.size(x)
n2 = np.size(y)
res = np.zeros(shape=(n1, n2))
for i in range(n1):
for j in range(n2):
res[i][j] = x[i]*y[j]
return res
print("Distribution jointe de PA PB:\n",Pxy(PA, PB))
# ### II.4- Affichage de la distribution jointe
# <img src="tme2_jointe.jpg" title="Distribution jointe" style="float: right;">
#
# Le code ci-dessous permet d'afficher en 3D une probabilité jointe générée par la fonction précédente. Exécutez-le avec une probabilité jointe résultant de la combinaison d'une loi normale et d'une distribution affine.
#
# Si la commande `%matplotlib notebook` fonctione, vous pouvez interagir avec la courbe. Si le contenu de la fenêtre est vide, redimensionnez celle-ci et le contenu devrait apparaître. Cliquez à la souris à l'intérieur de la fenêtre et bougez la souris en gardant le bouton appuyé afin de faire pivoter la courbe. Observez sous différents angles cette courbe. Refaites l'expérience avec une probaiblité jointe résultant de deux lois normales. Essayez de comprendre ce que signifie, visuellement, l'indépendance probabiliste. Vous pouvez également recommencer l'expérience avec le logarithme des lois jointes.
#
# +
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib inline
# essayer `%matplotib notebook` pour interagir avec la visualisation 3D
def dessine ( P_jointe ):
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
x = np.linspace ( -3, 3, P_jointe.shape[0] )
y = np.linspace ( -3, 3, P_jointe.shape[1] )
X, Y = np.meshgrid(x, y)
ax.plot_surface(X, Y, P_jointe, rstride=1, cstride=1 )
ax.set_xlabel('A')
ax.set_ylabel('B')
ax.set_zlabel('P(A) * P(B)')
plt.show ()
# -
dessine(Pxy(PA,PB))
k = 101
sigma = 10
slope = 1e-5
dessine(Pxy(normale(k, sigma)[1], proba_affine(k, slope)[1]))
# ## III- Indépendances conditionnelles (<font color="red"> obligatoire</font>)
#
# Dans cet exercice, on considère quatre variables aléatoires booléennes $X$, $Y$, $Z$ et $T$ ainsi que leur distribution jointe $P(X,Y,Z,T)$ encodée en python de la manière suivante :
# creation de P(X,Y,Z,T)
P_XYZT = np.array([[[[ 0.0192, 0.1728],
[ 0.0384, 0.0096]],
[[ 0.0768, 0.0512],
[ 0.016 , 0.016 ]]],
[[[ 0.0144, 0.1296],
[ 0.0288, 0.0072]],
[[ 0.2016, 0.1344],
[ 0.042 , 0.042 ]]]])
# Ainsi, $\forall (x,y,z,t) \in \{0,1\}^4$, `P_XYZT[x][y][z][t]` correspond à $P(X=x,Y=y,Z=z,T=t)$ ou, en version abrégée, à $P(x,y,z,t)$.
# ### III.1- Indépendance de X et T conditionnellement à (Y,Z)
# On souhaite tester si les variables aléatoires $X$ et $T$ sont indépendantes conditionnellement à $(Y,Z)$. Il s'agit donc de vérifer que dans la loi $P$, $$P(X,T|Y,Z)=P(X|Y,Z)\cdot P(T|Y,Z)$$
#
#
# Pour cela, tout d'abord, calculer à partir de `P_XYZT` le tableau `P_YZ` représentant la distribution $P(Y,Z)$. On rappelle que $$P(Y,Z)=\sum_{X,T} P(X,Y,Z,T)$$
#
# Le tableau `P_YZ` est donc un tableau à deux dimensions, dont la première correspond à $Y$ et la deuxième à $Z$. Si vous ne vous êtes pas trompé(e)s, vous devez obtenir le tableau suivant :
# ```
# np.array([[ 0.336, 0.084],
# [ 0.464, 0.116]])
# ```
#
# Ainsi $P(Y=0,Z=1)=$ `P_YZ[0][1]` $=0.084$
# +
P_YZ = np.zeros((2,2))
for x in range(P_XYZT.shape[0]):
for y in range(P_XYZT.shape[1]):
for z in range(P_XYZT.shape[2]):
for t in range(P_XYZT.shape[3]):
P_YZ[y][z] += P_XYZT[x][y][z][t]
print(P_YZ)
# -
# Ensuite, calculer le tableau `P_XTcondYZ` représentant la distribution $P(X,T|Y,Z)$. Ce tableau a donc 4 dimensions, chacune correspondant à une des variables aléatoires. De plus, les valeurs de `P_XTcondYZ` sont obtenues en utilisant la formule des probabilités conditionnelles: $$P(X,T|Y,Z)=\frac{P(X,Y,Z,T)}{P(Y,Z)}$$
# +
P_XTcondYZ = np.zeros((2,2,2,2))
for x in range(P_XYZT.shape[0]):
for y in range(P_XYZT.shape[1]):
for z in range(P_XYZT.shape[2]):
for t in range(P_XYZT.shape[3]):
P_XTcondYZ[x][y][z][t] += P_XYZT[x][y][z][t] / P_YZ[y][z]
print(P_XTcondYZ)
# -
# Calculer à partir de `P_XTcondYZ` les tableaux à 3 dimensions `P_XcondYZ` et `P_TcondYZ` représentant respectivement les distributions $P(X|Y,Z)$ et $P(T|Y,Z)$. On rappelle que $$P(X|Y,Z)=∑_Y P(X,T|Y,Z)$$
# +
P_XcondYZ = np.zeros((2,2,2))
P_TcondYZ = np.zeros((2,2,2))
for x in range(P_XYZT.shape[0]):
for y in range(P_XYZT.shape[1]):
for z in range(P_XYZT.shape[2]):
for t in range(P_XYZT.shape[3]):
P_XcondYZ[x][y][z] += P_XTcondYZ[x][y][z][t]
P_TcondYZ[y][z][t] += P_XTcondYZ[x][y][z][t]
print(P_XcondYZ)
print(P_TcondYZ)
# -
# Enfin, tester si $X$ et $T$ sont indépendantes conditionnellement à $(Y,Z)$: si c'est bien le cas, on doit avoir $$P(X,T|Y,Z)=P(X|Y,Z)×P(T|Y,Z)$$
# +
# A = np.round(P_XcondYZ * P_TcondYZ, 5) # Erreur !
A = np.zeros((2, 2, 2, 2))
ecart = 0
epsilon = 1e-3
for x in range(P_XYZT.shape[0]):
for y in range(P_XYZT.shape[1]):
for z in range(P_XYZT.shape[2]):
for t in range(P_XYZT.shape[3]):
# A[x][y][z][t] = P_XcondYZ[x][y][z] * P_TcondYZ[y][z][t]
ecart += np.abs((P_XcondYZ[x][y][z] * P_TcondYZ[y][z][t]) - P_XTcondYZ[x][y][z][t])
if ecart < epsilon:
print("indépendant")
else:
print("pas indépendant")
# A = np.round(A, 5)
# B = np.round(P_XTcondYZ, 5)
# print(A.shape)
# print("=======")
# print(B.shape)
# print(A == B)
# -
# ### III.2- Indépendance de X et (Y,Z)
#
# On souhaite maintenant déterminer si $X$ et $(Y,Z)$ sont indépendantes. Pour cela, commencer par calculer à partir de `P_XYZT` le tableau `P_XYZ` représentant la distribution $P(X,Y,Z)$.
#
# Ensuite, calculer à partir de `P_XYZ` les tableaux `P_X` et `P_YZ` représentant respectivement les distributions $P(X)$ et $P(Y,Z)$. On rappelle que $$P(X)=∑_Y∑_Z P(X,Y,Z)$$
#
# Si vous ne vous êtes pas trompé(e), P_X doit être égal au tableau suivant :
# ```
# np.array([ 0.4, 0.6])
# ```
# +
P_XYZ = np.zeros((2,2,2))
for x in range(P_XYZT.shape[0]):
for y in range(P_XYZT.shape[1]):
for z in range(P_XYZT.shape[2]):
for t in range(P_XYZT.shape[3]):
P_XYZ[x][y][z] += P_XYZT[x][y][z][t]
print(P_XYZ)
P_X = np.zeros((2))
for x in range(P_XYZT.shape[0]):
for y in range(P_XYZT.shape[1]):
for z in range(P_XYZT.shape[2]):
P_X[x] += P_XYZ[x][y][z]
print(P_X)
# -
# Enfin, si $X$ et $(Y,Z)$ sont bien indépendantes, on doit avoir $$P(X,Y,Z)=P(X)×P(Y,Z)$$
# +
A = np.zeros((2, 2, 2))
ecart = 0
epsilon = 1e-3
for x in range(P_XYZ.shape[0]):
for y in range(P_XYZ.shape[1]):
for z in range(P_XYZ.shape[2]):
A[x][y][z] = P_X[x] * P_YZ[y][z]
ecart += np.abs(P_X[x] * P_YZ[y][z] - P_XYZ[x][y][z])
if ecart < epsilon:
print("indépendant")
else:
print("pas indépendant")
# A = np.round(A, 5)
# B = np.round(P_XYZ, 5)
# print(A.shape)
# print("=======")
# print(B.shape)
# print(A == B)
# -
# ## IV- Indépendances conditionnelles et consommation mémoire (<font color="red"> obligatoire</font>)
#
# Le but de cet exercice est d'exploiter les probabilités conditionnelles et les indépendances conditionnelles afin de décomposer une probabilité jointe en un produit de "petites probabilités conditionnelles". Cela permet de stocker des probabilités jointes de grandes tailles sur des ordinateurs "standards". Au cours de l'exercice, vous allez donc partir d'une probabilité jointe et, progressivement, construire un programme qui identifie ces indépendances conditionnelles.
#
# Pour simplifier, dans la suite de cet exercice, nous allons considérer un ensemble $X_0,…,X_n$ de variables aléatoires binaires (elles ne peuvent prendre que 2 valeurs : 0 et 1).
#
# ### Simplification du code : utilisation de pyAgrum
#
# Manipuler des probabilités et des opérations sur des probabilités complexes est difficiles avec les outils classiques. La difficulté principale est certainement le problème du mapping entre axe et variable aléatoire. `pyAgrum` propose une gestion de `Potential` qui sont des tableaux multidimensionnels dont les axes sont caractérisés par des variables et sont donc non ambigüs.
#
# Par exemple, après l'initiation du `Potential PABCD` :
# +
import pyAgrum as gum
import pyAgrum.lib.notebook as gnb
X,Y,Z,T=[gum.LabelizedVariable(x,x,2) for x in "XYZT"]
pXYZT=gum.Potential().add(T).add(Z).add(Y).add(X)
pXYZT[:]=[[[[ 0.0192, 0.1728],
[ 0.0384, 0.0096]],
[[ 0.0768, 0.0512],
[ 0.016 , 0.016 ]]],
[[[ 0.0144, 0.1296],
[ 0.0288, 0.0072]],
[[ 0.2016, 0.1344],
[ 0.042 , 0.042 ]]]]
# -
# On peut alors utiliser la méthode `margSumOut` qui supprime les variables par sommations: `p.margSumOut(['X','Y'])` correspond à calculer $\sum_{X,Y} p$
#
# La réponse a question III.1 se calcule donc ainsi :
# +
pXT_YZ=pXYZT/pXYZT.margSumOut(['X','T'])
pX_YZ=pXT_YZ.margSumOut(['T'])
pT_YZ=pXT_YZ.margSumOut(['X'])
if pXT_YZ==pX_YZ*pT_YZ:
print("=> X et T sont indépendants conditionnellemnt à Y et Z")
else:
print("=> pas d'indépendance trouvée")
# -
# La réponse à la question III.2 se calcule ainsi :
pXYZ=pXYZT.margSumOut("T")
pYZ=pXYZ.margSumOut("X")
pX=pXYZ.margSumOut(["Y","Z"])
if pXYZ==pX*pYZ:
print("=> X et YZ sont indépendants")
else:
print("=> pas d'indépendance trouvée")
gnb.sideBySide(pXYZ,pX,pYZ,pX*pYZ,
captions=['$P(X,Y,Z)$','$P(X)$','$P(Y,Z)$','$P(X)\cdot P(Y,Z)$'])
# `asia.txt` contient la description d'une probabilité jointe sur un ensemble de $8$ variables aléatoires binaires (256 paramètres). Le fichier est produit à partir du site web suivant `http://www.bnlearn.com/bnrepository/`.
#
# Le code suivant permet de lire ce fichier et d'en récupérer la probabilité jointe (sous forme d'une `gum.Potential`) qu'il contient :
# +
def read_file ( filename ):
"""
Renvoie les variables aléatoires et la probabilité contenues dans le
fichier dont le nom est passé en argument.
"""
Pres = gum.Potential ()
vars=[]
with open ( filename, 'r' ) as fic:
# on rajoute les variables dans le potentiel
nb_vars = int ( fic.readline () )
for i in range ( nb_vars ):
name, domsize = fic.readline ().split ()
vars.append(name)
variable = gum.LabelizedVariable(name,name,int (domsize))
Pres.add(variable)
# on rajoute les valeurs de proba dans le potentiel
cpt = []
for line in fic:
cpt.append ( float(line) )
Pres.fillWith( cpt )
return vars,Pres
vars,Pjointe=read_file('asia.txt')
# afficher Pjointe est un peu délicat (retire le commentaire de la ligne suivante)
# Pjointe
print('Les variables : '+str(vars))
# -
# Noter qu'il existe une fonction margSumIn qui, à l'inverse de MargSumOut, élimine
# toutes les variables qui ne sont pas dans les arguments
Pjointe.margSumIn(['tuberculosis?','lung_cancer?'])
# ### IV.1- test d'indépendance conditionnelle
#
# En utilisant la méthode `margSumIn` (voir juste au dessus), écrire une fonction `conditional_indep: Potential,str,str,list[str]->bool` qui rend vrai si dans le `Potential`, on peut lire l'indépendance conditionnelle.
#
# Par exemple, l'appel
#
# `conditional_indep(Pjointe,'bronchitis?', 'positive_Xray?',['tuberculosis?','lung_cancer?'])`
#
# vérifie si bronchitis est indépendant de `posititve_Xray` conditionnellement à `tuberculosis?` et `lung_cancer?`
#
# D'un point de vue général, on vérifie que $X$ et $Y$ sont indépendants conditionnellement à $Z_1,\cdots,Z_d$ par l'égalité :
# $$P(X,Y|Z_1,\cdots,Z_d)=P(X|Z_1,\cdot,Z_d)\cdot P(Y|Z_1,\cdots,Z_d)$$
#
# Ces trois probabilités sont calculables à partir de la loi jointe de $P(X,Y,Z_1,\cdots,Z_d)$.
#
# <em>Remarque</em> Vérifier l'égalité `P==Q` de 2 `Potential` peut être problématique si les 2 sont des résultats de calcul : il peut exister une petite variation. Un meilleur test est de vérifier `(P-Q).abs().max()<epsilon` avec `epsilon` assez petit.
# +
epsilon = 1e-3
def conditional_indep(P,X,Y,Zs):
variables = [X, Y]
for var in Zs:
variables.append(var)
if len(Zs) != 0:
pXYZs = P.margSumIn(variables) # Besoin de cette valeur pour calculer PXY_ZS
pXY_Zs = pXYZs/pXYZs.margSumOut([X, Y]) # On enleve X Y pour avoir X et Y conditionnellement à Zs
pX_Zs = pXY_Zs.margSumOut([Y])
pY_Zs = pXY_Zs.margSumOut([X])
A = pX_Zs * pY_Zs
B = pXY_Zs
# print("A:",A)
# print("B:",B)
# print("A-B",A-B)
test = (A-B).abs().max() < epsilon
if(test):
return 1 # Independant
else:
return 0 # Pas independant
else:
pX = P.margSumIn(X)
pY = P.margSumIn(Y)
PXY = P.margSumIn([X, Y])
A = pX * pY
B = PXY
# print("A:",A)
# print("B:",B)
# print("A-B",A-B)
test = (A-B).abs().max() < epsilon
if(test):
return 1
else:
return 0
# -
conditional_indep(Pjointe,
'bronchitis?',
'positive_Xray?',
['tuberculosis?','lung_cancer?'])
conditional_indep(Pjointe,
'bronchitis?',
'visit_to_Asia?',
[])
# ### IV.2- Factorisation compacte de loi jointe
#
# On sait que si un ensemble de variables aléatoires ${\cal S} = \{X_{i_0},\ldots,X_{i_{n-1}}\}$ peut être partitionné en deux sous-ensembles $\cal K$ et $\cal L$ (c'est-à-dire tels que ${\cal K} \cap {\cal L} = \emptyset$ et ${\cal K} \cup {\cal L} = \{X_{i_0},\ldots,X_{i_{n-1}}\}$) tels qu'une variable $X_{i_n}$ est indépendante de ${\cal L}$ conditionnellement à ${\cal K}$, alors:
#
# $$P(X_{i_n}|X_{i_0},\ldots,X_{i_{n-1}}) = P(X_{i_n} | {\cal K},{\cal L}) = P(X_{i_n} | {\cal K})$$
#
# C'est ce que nous avons vu au cours n°2 (cf. définition des probabilités conditionnelles). Cette formule est intéressante car elle permet de réduire la taille mémoire consommée pour stocker $P(X_{i_n}|X_{i_0},\ldots,X_{i_{n-1}})$: il suffit en effet de stocker uniquement $P(X_{i_n} | {\cal K})$ pour obtenir la même information.
# Écrire une fonction `compact_conditional_proba: Potential,str-> Potential` qui, étant donné une probabilité jointe $P(X_{i_0},\ldots,X_{i_n})$, une variable aléatoire $X_{i_n}$, retourne cette probabilité conditionnelle $P(X_{i_n} | {\cal K})$. Pour cela, nous vous proposons l'algorithme itératif suivant:
#
# ```
# K=S
# Pour tout X in K:
# Si X indépendante de Xin conditionnellement à K\{X) alors
# Supprimer X de K
# retourner P(Xin|K)$
# ```
#
# Trois petites aides :
#
# 1- La fonction precédente `conditional_indep` devrait vous servir...
#
# 2- Obtenir la liste des noms des variables dans un `Potential` se fait par l'attribut
# ```
# P.var_names
# ```
#
# 3- Afin que l'affichage soit plus facile à comprendre, il peut être judicieux de placer la variable $X_{i_n}$ en premier dans la liste des variables du Potential, ce que l'on peut faire avec le code suivant :
# ```
# proba = proba.putFirst(Xin)
# ```
#
#
def compact_conditional_proba(P, X):
if P.domainSize() <= 2: # Si tableau vide on retourne le tableau car plus possible de compacter
return P
K = P.var_names # Ensemble des variables dans la Pjointe
K.remove(X)
for k in K:
tmp_K = K.copy()
tmp_K.remove(k)
if conditional_indep(P, k, X, K): # Check si les variables dans K sont independantes
K.remove(k) # Supprime de la liste si independantes
var = []
var.append(X)
for k in K:
var.append(k)
PX_K = P.margSumIn(var) / P.margSumIn(K)
PX_K = PX_K.putFirst(X)
return PX_K
compact_conditional_proba(Pjointe,"visit_to_Asia?")
compact_conditional_proba(Pjointe,"dyspnoea?")
# ### IV.3- Création d'un réseau bayésien
#
# Un réseau bayésien est simplement la décomposition d'une distribution de probabilité jointe en un produit de probabilités conditionnelles: vous avez vu en cours que $P(A,B) = P(A|B)P(B)$, et ce quel que soient les ensembles de variables aléatoires disjoints $A$ et $B$. En posant $A = X_n$ et $B = \{X_0,\ldots,X_{n-1}\}$, on obtient donc:
#
# $$P(X_0,\ldots,X_n) = P(X_n | X_0,\ldots,X_{n-1}) P(X_0,\ldots,X_{n-1})$$
#
# On peut réitérer cette opération pour le terme de droite en posant $A = X_{n-1}$ et $B=\{X_0,\ldots,X_{n-2}\}$, et ainsi de suite. Donc, par récurrence, on a:
#
# $$P(X_0,\ldots,X_n) = P(X_0) \times \prod_{i=1}^n P(X_i | X_0,\ldots,X_{i-1} )$$
#
# Si on applique à chaque terme $P(X_i | X_0,\ldots,X_{i-1} )$ la fonction `compact_conditional_proba`, on obtient une décomposition:
#
# $$P(X_0,\ldots,X_n) = P(X_0) \times \prod_{i=1}^n P(X_i | {\cal K_i})$$
#
# avec $K_i \subseteq \{X_0,\ldots,X_{i-1}\}$}. Cette décomposition est dite ''compacte'' car son stockage nécessite en pratique beaucoup moins de mémoire que celui de la distribution jointe. C'est ce que l'on appelle un réseau bayésien.
#
# Écrire une fonction `create_bayesian_network : Potential -> Potential list` qui, étant donné une probabilité jointe, vous renvoie la liste des $P(X_i | {\cal K_i})$. Pour cela, il vous suffit d'appliquer l'algorithme suivant:
#
# ```
# liste = []
# P = P(X_0,...,X_n)
# Pour i de n à 0 faire:
# calculer Q = compact_conditional_proba(P,X_i)
# afficher la liste des variables de Q
# rajouter Q à liste
# supprimer X_i de P par marginalisation
#
# retourner liste
# ```
#
# Il est intéressant ici de noter les affichages des variables de Q: comme toutes les variables sont binaires, Q nécessite uniquement (2 puissance le nombre de ces variables) nombres réels. Ainsi une probabilité sur 3 variables ne nécessite que {$2^3=8$} nombres réels.
#
def create_bayesian_network(P):
res = []
liste_names = P.var_names
for X_i in reversed(liste_names):
print(X_i)
Q = compact_conditional_proba(P, X_i)
# print(Q)
res.append(Q)
P = P.margSumOut(X_i)
return res
bn = create_bayesian_network(Pjointe)
print(bn)
# ### IV.4- Gain en compression
#
# On souhaite observer le gain en termes de consommation mémoire obtenu par votre décomposition. Si `P` est un `Potential`, alors `P.toarray().size` est égal à la taille (le nombre de paramètres) de la table `P`. Calculez donc le nombre de paramètres nécessaires pour stocker la probabilité jointe lue dans le fichier `asia.txt` ainsi que la somme des nombres de paramètres des tables que vous avez créées grâce à votre fonction `create_bayesian_network`.
# +
Pjointe = read_file('asia.txt')[1]
print("Size before:", Pjointe.toarray().size)
bn = create_bayesian_network(Pjointe)
acc = 0
for i in range(0, len(bn)):
acc += bn[i].domainSize()
print("Size after:", acc)
# -
# ## V- Applications pratiques (optionnelle)
# La technique de décomposition que vous avez vue est effectivement utilisée en pratique. Vous pouvez voir le gain que l'on peut obtenir sur différentes distributions de probabilité du site :
#
# http://www.bnlearn.com/bnrepository/
#
# Cliquez sur le nom du dataset que vous voulez visualiser et téléchargez son .bif ou .dsl. Afin de visualiser le contenu du fichier, vous allez utiliser pyAgrum. Le code suivant vous permettra alors de visualiser votre dataset: la valeur indiquée après "domainSize" est la taille de la probabilité jointe d'origine (en nombre de paramètres) et celle après "dim" est la taille de la probabilité sous forme compacte (somme des tailles des probabilités conditionnelles compactes).
# +
# chargement de pyAgrum
import pyAgrum as gum
import pyAgrum.lib.notebook as gnb
# chargement du fichier bif ou dsl
bn = gum.loadBN ("earthquake.bif")
# affichage de la taille des probabilités jointes compacte et non compacte
print(bn)
# affichage graphique du réseau bayésien
bn
| S1/MAPSI/TME/TME2/enonce/TME2DurandKordon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
data = load_iris().data
targets = load_iris().target
# +
covariance = np.cov(data, rowvar=False)
means = np.mean(data, axis=0)
adjusted = np.subtract(data,means)
# 1/(N-1) * (X - X^)(X - X^)'
covariance = np.subtract(np.matmul(np.transpose(adjusted),adjusted),np.mean(adjusted,axis=0)) / (data.shape[0] -1)
print(covariance)
# -
# For CCA and so on, the correlation matrix can be used instead. Here it's calculated as a diagonalization problem instead of the typical Corr(X,Y) = cov(X,Y)/||X^||
D = np.sqrt(np.diagonal(covariance) * np.identity(4))
D_inv = np.linalg.inv(D)
correlation = np.matmul(np.matmul(D_inv,covariance),D_inv)
print(correlation)
# Get eigenvectors, already sorted on magnitude
val,vec = np.linalg.eig(covariance)
print(val)
# +
num_components = 2
pcadata = np.zeros((num_components,150))
for i in range(num_components):
# Remember to use X - X^ matrix as data, otherwise we don't project it porpperly.
pcadata[i,:] = np.matmul(adjusted,vec[i])
# +
plt.scatter(pcadata[0,:],pcadata[1,:], c = targets)
plt.show()
plt.scatter(data[:,0], data[:,1], c = targets)
plt.show()
# -
| PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Intro to Python
# ----
#
#
# ## Table of Contents
# 1. [Intro to Python](#Intro-to-Python)
# - [Introduction](#Introduction)
# - [Python](#Python)
# - [Python-Setup](#Python-Setup)
# - [Basic-Plotting](#A-simple-example-of-plotting-of-using-numpy-and-plotting)
# - [Getting help/documentation](#Getting-help/documentation)
# - [Data-Basics](#Data-Basics)
# - [Loading Data](#Loading-Data)
# - [Understanding the Data](#Understanding-the-Data)
# - [Value Counts](#value_counts)
# - [Subplots](#Subplots)
# - [Adding-columns](#Adding-columns)
# - [Saving Results](#Saving-Results)
# 4. [Exercises](#Exercises)
# 5. [Resources](#Resources)
#
# ## Introduction
# This course is a practical introduction to the methods and tools that a social scientist can use to make sense of big data, and thus programming resources are also important. We make extensive use of the Python programming language and SQL database management. We recommend that any social scientist who aspires to work with large datasets become proficient in the use of these two systems as well as [Github](http://www.github.com). All three, fortunately, are quite accessible and supported by excellent online resources.
# ## Python
# ---
# - Back to [Table of Contents](#table-of-contents)
#
#
# > Before coming to class, you should have completed the [DataCamp Intro to Python for Data Science](https://www.datacamp.com/courses/intro-to-python-for-data-science) course. It is free and takes about four hours.
#
#
# Python is a high-level interpreted general purpose programming language named after a British Comedy Troupe. Python was created by
# <NAME> (Python's benovolent dictator fore life), and is maintained by an international group of enthusiasts.
#
# As of the time of this writing (10/2016) Python is currently the fifth most popular programming language. It is popular for data science because it is powerful and fast, it "plays well" with other languages, it runs everywhere, it's easy to learn, it's highly readable, open-source and its fast development time compared to other languages. Because of its general-purpose nature and its ability to call compiled languages like FORTRAN or C it can be used in full-stack development. There is a growing and always-improving list of open-source libraries for scientific programming, data manipulation, and data analysis (e.g., Numpy, Scipy, Pandas, Scikit-Learn, Statsmodels, Matplotlib, Seaborn, PyTables, etc.)
#
# [IPython](http://www.ipython.org) is an enhanced, interactive python interpreter that started as a grad school project by <NAME>. The project evolved into the IPython notebook, which allowed users to archive their code, figures, and analysis in a single document, making doing reproducible research and sharing said research much easier. The creators of the IPython notebook quickly realized that the "notebook" aspects were agnostic with respect to programming language, and ported the notebook to other languages including but not limited to Julia, Python and R. This then led to a rebranding known as the Jupyter Project.
#
# This tutorial will go over the basics of Data Analysis in Python using the PyData stack.
#
#
#
# ### Python Setup
# - In Python, we `import` packages. The `import` command allows us to use libraries created by others in our own work by "importing" them. You can think of importing a library as opening up a toolbox and pulling out a specific tool.
# - NumPy is short for numerical python. NumPy is a lynchpin in Python's scientific computing stack. Its strengths include a powerful *N*-dimensional array object, and a large suite of functions for doing numerical computing.
# - Pandas is a library in Python for data analysis that uses the DataFrame object from R which is similiar to a spreedsheet but allows you to do your analysis programaticaly rather than the point-and-click of Excel. It is a lynchpin of the PyData stack.
# - Psycopg2 is a python library for interfacing with a PostGreSQL database.
# - Matplotlib is the standard plotting library in python.
# `%matlplotlib inline` is a so-called "magic" function of Jupyter that enables plots to be displayed inline with the code and text of a notebook.
import matplotlib.pyplot
import numpy as np
import pandas as pd
import psycopg2
import sqlalchemy
from __future__ import print_function
# %matplotlib inline
# In practice we typically load libraries like `numpy` and `pandas` with shortened aliases, e.g, `import numpy as np`. This is like saying, "`import numpy`, and wherever you see `np`, read it as `numpy`." Similarly, you'll often see `import pandas as pd`, or `import matplotlib.pyplot as plt`.
#
# Another shortcut is `%pylab inline`. This command includes both `import numpy as np` and `import matplotlib.pyplot as plt `. This shortcut was invented because it's faster to type `plt.plot()` rather than `matplotlib.pyplot.plot()`, and even programmers don't like to type more than they have to.
#
# In documentation and in examples, you will frequently see `numpy` commands starting with the alias `np` rather than `numpy` (e.g, `np.array()` or `np.argsort`) and `pandas` commands starting with `pd` (e.g., `pd.DataFrame()` or `pd.concat()`). See below for an example of using aliases.
# ```
# # %pylab inline
# import pandas as pd
# import psycopg2
# from __future__ import print_function
# ```
# Now all `numpy` commands will be prefixed with `np` and all plotting commands will be prefixed with `plt`.
# ## A simple example of plotting using numpy and matplotlib
# The main workhorse library for plotting in Python is `matplotlib`. All commands in the `matplotlib` plotting library are stored in the `plt` namespace. A *namespace* contains all of the functions of that library grouped together, denoted by a common prefix. Any command using some function from the `matplotlib` plotting library will start with `plt`; for example, to create a plot we use the `plt.plot()` command. That way, if you've imported multiple libraries that each have a function called `plot()`, you know you're using the *right* one. Below is a very simple example of plotting a sine wave and adding labels to the axes.
x = np.linspace(0,4*np.pi,100) #the linspace command from the numpy library
#creates a set of 100 equally spaced points from 0 to 4pi
y = np.sin(x) #calculates the sin(x) which is stored in the y variable
matplotlib.pyplot.plot(x,y) #plot x vs y
matplotlib.pyplot.ylabel('sin(x)') #set the y-label
matplotlib.pyplot.xlabel('x') # set the x-label
# ## Documentation and getting help
#
# Jupyter has great features for looking up the documentation of a function.
# To get the full documentation of a function, just type a question mark after the function name.
# +
# np.array?
# -
# To get an abbreviated documentation of a function, we can place the cursor in between the parentheses of a function and press `Shift+Tab` to get a shorter version of the documentation.
# try it for yourself
np.array()
# ### Data Basics
# - Information about the data that we're using: where did it come from, what variables are present?
# - How to connect to the database
# In this lesson, we'll be using the [pandas package](http://pandas.pydata.org/) to read in and manipulate data. `pandas` reads data from the PostGreSQL database and stores the data in special table format called a "dataframe," which will be familiar to you if you have used R or STATA for data analysis. Dataframes allow for easy statistical analysis, and can be directly used for machine learning.
#
# `pandas` uses a database engine to connect to databases. In the code cell below, we'll use `sqlalchemy` to connect to the database.
db_name = "appliedda"
db_host = "10.10.2.10"
pgsql_connection = psycopg2.connect( host = db_host, database = db_name )
cur = pgsql_connection.cursor()
pgsql_engine = sqlalchemy.create_engine( "postgresql://10.10.2.10/appliedda" )
# ### Loading Data
# Next, we will use this database connection to have `pandas` retrieve the data. `pandas` has a set of [Input/Output tools](http://pandas.pydata.org/pandas-docs/stable/io.html) that let it read from and write to a large variety of tabular data formats, including CSV and Excel files, databases via SQL, JSON files, and even SAS and Stata data files. In the example below, we'll use the `pandas.read_sql()` function to read the results of an SQL query into a data frame.
#
# - We will change this query to only select the dataset that we want
query = 'SELECT * FROM {table};'.format(table="ildoc_admit")
df_ildoc_admit = pd.read_sql( query, con = pgsql_engine )
# Now, let's see what the data looks like. The `pandas.DataFrame` method `head(number_of_rows)` outputs the first `number_of_rows` rows in a dataframe. Let's look at the first five rows in our data. Since our `pandas.DataFrame` object is called `df_ildoc_admit` and we want to see 5 rows, we'll type `df_ildoc_admit.head(5)`.
#
# In the code cells below, you'll see two ways to output this information. If you just call the method, you'll see an HTML table displayed directly in the IPython notebook. This is a useful way to display the information if you only want to view it in this notebook. If you pass the results of the method to the `print()` function, you'll get text output. This is useful if you want to export the output for use outside the notebook.
# to get a pretty tabular view, just call the method.
df_ildoc_admit.head( 5 )
print(df_ildoc_admit.head(5))
# Now let's look at the *bottom* five rows. As you might be able to guess, `tail(number_of_rows)` does almost the same thing as `head(number_of_rows)`, but gives you the *last* `number_of_rows` rather than the *first* `number_of_rows`.
df_ildoc_admit.tail(5)
# One thing to notice is that we have a pesky column called `unnamed:0`, which is just a duplicate of the index columns. Let's drop this column, since it has no purpose. We'll use the `drop()` method, and specify what we want to drop (`['unnamed: 0']`), which `axis` we're dropping it from (0 for rows, 1 for columns), and whether we want to make a *new* version of `df_ildoc_admit`, or just drop the column `inplace`. If you think you might want to be able to look back at the version of your dataframe before you made the alterations, you can specify `inplace=False`, but you will need to save the altered version under a new name.
df_ildoc_admit.drop(['unnamed: 0'], axis=1, inplace=True)
# To learn more about the `drop` command and the arguments it uses, you can use our handy trick to read the documentation.
# +
# df_ildoc_admit.drop?
# -
# ### Understanding the Data
# In `pandas`, our data is represented by a `DataFrame`. You can think of dataframes as giant spreadsheets which you can *program*, rather than manipulating them with point-and-click tools like Excel. In a dataframe, each column is stored in its own *list*, which `pandas` calls a `Series` (or vector of values), along with a set of **methods** (another name for functions that are tied to objects) that make managing data easy.
#
# A `Series` is a list of values. Each value in a `Series` can have its own *label*, or `index`. If you retrieve a single *row* from your dataframe, it will come along with its `index`, or the names of the columns represented in each cell. If you retrieve a single *column* from your dataframe, the accompanying `index` will tell your the row IDs.
#
# While `DataFrame` and `Series` are separate objects, they share some of the same methods. In general, shared methods are those make sense in both a table and a list context (for example, `head()` and `tail()`, as seen in this notebook, can be used on both `DataFrame` and `Series` objects).
#
# More details on `pandas` data structures:
# - [Data Structures Overview](http://pandas.pydata.org/pandas-docs/stable/dsintro.html)
# - [Series specifics](http://pandas.pydata.org/pandas-docs/stable/dsintro.html#series)
# - [DataFrame specifics](http://pandas.pydata.org/pandas-docs/stable/dsintro.html#dataframe)
# With over 200 columns in our dataframe, it's too hard to look at them all at once. Let's get a list of the column names to see what we're working with.
for i, col in enumerate(df_ildoc_admit.columns):
print(i,col)
# We see that there are 205 columns (although the last number is 204, remember that in Python we start counting from 0). It appears that a few of the columns might be redundant, like `birth_year` and `birth_yr`.
# Let's try to break down this code snippet.
# ```
# for i, col in enumerate(df_ildoc_admit.columns):
# print(i,col)
# ```
# First, `df_ildoc_admit.columns` gives us a list of the column names. We then *pass* that list to the `enumerate()` function, which returns a numerical index (`i`) and the name of the column (`col`) as the for loop goes through the list of columns. We print the values of `i` and `col` at each iteration using `print()`. Note that we can print multiple values (both `i` and `col`) so long as they are separated by a comma.
# The numbers of rows and columns are stored in the `shape` attribute of a `DataFrame`.
df_ildoc_admit.shape
# We can check if there are any duplicate rows in the dataframe by using the `drop_duplicates()` method and checking if that changes the shape.
df_ildoc_admit.drop_duplicates().shape
# Great, no duplicates!
#
# Here we ran the command `df_ildoc_admit.drop_duplicates()`, which returns a `DataFrame` with all duplicates removed, then accessed the `shape` of the *new* dataframe by just adding `.shape` after the `drop_duplicates()`. In many other languages, you would have to take these two steps individually, or nest the commands within each other in a way that makes your code hard to read. In this case, we did it all in one fell swoop, and it's easy to understand what we did, reading from left to right. The chaining of commands is a handy feature of python.
# Now let's start exploring the data!
# As we mentioned before, you can think of a `DataFrame` is an object made up of `Series` objects, which comprise its columns. Let's pull out just the `Series` for `'race'` and examine it.
df_ildoc_admit['race'].head(10) #get first 10 entries of 'race' column
df_ildoc_admit['race'].tail() #output the last five entries
# Note that in the `tail()` example above we didn't *specify* how many rows we wanted, and we got 5, because that's the default for this method. It's good practice to read documentation, so you know the default values methods use for any of the possible arguments.
# ## Getting value counts
#
# Let's look at the distribution of `race` in our dataset.
#
# - **`value_counts()`** - The `value_counts()` "series method and top-level function computes a histogram of a one-dimensional array of values." ( See [documentation](http://pandas.pydata.org/pandas-docs/stable/basics.html#value-counts-histogramming-mode) ). This method returns a `Series` of the counts of the number of times each unique value in the column is present in the column (also known as frequencies), from largest count to least, with the value itself the label for each row.
df_ildoc_admit['race'].value_counts()
# We see that there are six distinct races represented. By far the most represented race is ####, which has more than twice as many entries as the next most common, white. Note that this does not necessarily mean that there are more than twice as many *people* who are ##### as ZZZZZZ represented in this dataset, because there could be multiple rows (i.e. multiple admissions) for an individual.
#
# Let's make a bar plot. `pandas` comes with a handy `plot()` function, which allows us to specify the type of plot we want using the keyword argument `kind`.
#
# When you give a function multiple *arguments*, the function can differentiate between the arguments by their order (**positional arguments**) or by their names (**keyword arguments**).
#
# A method like `head()` *expects* that the first argument you give it will be the number of rows you want, so you can get the first 10 rows of `df_ildoc_admit` by typing `df_ildoc_admit.head(10)`. This is an example of a positional argument. Keyword arguments have to be specified along with their name.
#
# In the case of `pd.Series.plot()`, the keyword argument is `kind`, for the *kind* of plot we want. We pass the string `'bar'` (with single quotes) to indicate we would like to plot a barplot. Some other possible values would be `'line'` or `'box'`. Check out the documentation to learn more.
df_ildoc_admit['race'].value_counts().plot(kind='bar')
# We can even create a horizontal barplot by passing the string `'barh'` into the `plot()` command.
df_ildoc_admit['race'].value_counts().plot(kind='barh')
# We can do a bit more with keyword arguments. When we use the `value_counts()` command, we can normalize the results to get *percentages* (rather than absolute numbers) of prisoners using the keyword argument `normalize=True`, and sort the results from largest to smallest using the keyword argument `ascending=True`. Then we can plot the resulting `Series` as a horizontal bar plot.
df_ildoc_admit['race'].value_counts(normalize=True, ascending=True).plot(kind='barh', xlim=(0,1))
# As we can see, those that identify as $#$#$# make up a larger proportion of the prison population than those that identify as #$#$##.
# Try to do the samething for the `sex` field.
df_ildoc_admit['sex'].value_counts()
df_ildoc_admit['sex'].value_counts(normalize=True, ascending=False).plot(kind='barh')
# Clearly $$$ make up the majority of the prisoners in our dataset.
# # Subplots
# `Matplotlib` also comes with a handy subplot feature where we can plot multiple subplots and tune the display of a plot.
#
# Let's take the last two plots we made and plot them side-by-side, and make the figures bigger. When we invoke the `plt.subplots()` command, the first argument is the number of rows, `1`, and `2` is the number of columns. We then pass a tuple pair `(16,6)` of length and width (in inches) to the keyword argument `figsize`. The `plt.subplots()` function returns a tuple `fig` and `ax` object. The `fig` object controls *figure-level attributes* of the figure, like saving the file, while the `ax` object controls axis-level attributes for each figure respectively. We can create a plots as we did above and specify which axis object to plot them on using the `ax` keyword.
#
fig, ax = matplotlib.pyplot.subplots(1,2, figsize=(16,6))
df_ildoc_admit['race'].value_counts(normalize=True, ascending=True).plot(kind='barh', ax=ax[0])
df_ildoc_admit['sex'].value_counts(normalize=True, ascending=True).plot(kind='barh', ax=ax[1])
fig.savefig('fig1_race_gender_barh.png')
# As a challenge, let's try to make a subplot where the barplots are *horizontally* stacked and the x-axis ranges from 0 to 1 in both plots.
fg, ax = matplotlib.pyplot.subplots(2, figsize=(16,6), sharex=True)
for i,pair in enumerate(df_ildoc_admit[['race','sex']].groupby('sex')):
label = pair[0]
df = pair[1]
df['race'].value_counts(normalize=True, ascending=True).plot(kind='barh', title=label, ax=ax[i], xlim=(0,1))
# Here we can see that *within each sex*, the breakdown of race is generally the same.
# ## Adding columns
#
# - Adding columns to a dataframe, using the map and lambda functions
#
# Now we'll calculate the approximate distribution of ages of people admitted into the corrections system (assuming that they are alive today).
#
# ### Lambda functions
#
# A **lambda function** is an anonymous function, or a function that we can use as a throw-away without explcitly naming it. Typically these are functions we only use once.
#
# In this case, we'll create a a simple function that calculates the difference in years between 2016 and another year.
#
# +
calc_age = lambda x : 2016 - x
print(calc_age(1980))
# -
# Armed with our new lambda function, we can use the **map function**, a method of the `Series` object, to map our function onto the entries in the `Series`. This allows us to use `calc_age()` on all of the entries in the `birth_year` column at once.
df_ildoc_admit['birth_year'].map(calc_age).head()
# Now that we have the ages, we can create a new field called `age` in our dataframe.
df_ildoc_admit['age'] = df_ildoc_admit['birth_year'].map(calc_age)
# We can also use the `describe()` method to get distributions of the ages.
df_ildoc_admit['age'].describe()
# Let's look at a histogram of the ages.
df_ildoc_admit['age'].plot(kind='hist')
# From the above plot, we see that the ages look roughly normally distributed, with the median age around XX, and a standard deviation of about XX years.
# ### Saving Results
# `Pandas` has methods for saving results in a variety of formats, from SQL dumps to Excel, JSON and CSV. Let's save our table as a CSV file using the `to_csv` method. We are going to pass the `index=False` parameter so we don't get those pesky `'Unnamed: 0'` columns when we reload it.
#
# The first argument we pass to `to_csv()` is what we want to call the file. By default, the file will be saved in the directory that you are working in.
df_ildoc_admit.to_csv('ildoc_admit.csv', index=False)
# ## Exercises
# ---
# - Back to [Table of Contents](#table-of-contents)
#
# We have just scratched the surface of what can be done. Let's do a bit more data analysis!
#
#
# # Create a barplot of the escape risk
#
# The data contains a field `escrisk` which stands for escape risk. The categories are:
#
#
# * H: High
# * M: Moderate
# * L: Low
# * P: Pending
#
# Create a barplot of the data. Also include values that are listed as `NaN`. See the documentation on how to include this option.
df_ildoc_admit['escrisk'].unique() #check what all the unique values are.
df_ildoc_admit['escrisk'].value_counts()
df_ildoc_admit['escrisk'].value_counts(normalize=True, ascending=True, dropna=False).plot(kind='barh', xlim=(0,1))
# **Note:** This excludes any row where the escrick is None
# ## Find the number of unique individuals in the data
#
# The IDOC number is a unique five digit number preceded by an alpha character which is assigned to an inmate at first reception to IDOC.
#
# 1. First find how many entries there are in the table by how many ildoc numbers there are.
# 2. Find how many unique ildoc numbers there are.
#
#
df_ildoc_admit['docnbr'].shape
df_ildoc_admit.docnbr.unique().shape
# # What is the maximum number of admits a single prisoner has had in the ildoc system
# Let's group the records by the `docnbr`, then count the number of records in each group.
#
# Sort the values from highest to least, and then get the top value. Which prisoner has the highest number of admits?
df_grpby_docnbr = df_ildoc_admit.groupby('docnbr') #use the groupby commmnd to group the records by ildoc number
s_count_docnbr = df_grpby_docnbr.size() #use the size method to count the number of records in each group
s_count_docnbr.sort_values(ascending=False)[:1] #sort the values and then pull out the first value which is the highest one
# When you get more comfortable with pandas and python this can be done in a single line.
df_ildoc_admit.groupby('docnbr').size().sort_values(ascending=False)[:1]
# # Find the mean, median and minimum values for the admits of a single prisoner in the ildoc system
df_ildoc_admit.groupby('docnbr').size().sort_values(ascending=False).mean()
df_ildoc_admit.groupby('docnbr').size().sort_values(ascending=False).min()
df_ildoc_admit.groupby('docnbr').size().sort_values(ascending=False).median()
# ## Find the distribution of admits by year.
df_ildoc_admit['curadmyr'].value_counts().sort_values(ascending=True).plot(kind='barh')
# ## Resources
# - Back to [Table of Contents](#table-of-contents)
#
# - <NAME>, the creator of pandas, wrote the standard text "Data Analyis in Python: Data wrangling with Pandas, Numpy and IPython"
# - <NAME>'s [Python for Economists](http://www.alexmbell.com/python-tutorial-for-economists/) provides a wonderful 30-page introduction to the use of Python in the social sciences, complete with XKCD cartoons.
# - Economists <NAME> and <NAME> provide a [very useful set of lectures and examples](http://lectures.quantecon.org)
# - For more detail, we recommend <NAME>'s [Python for Informatics: Exploring Information](http://www.pythonlearn.com/book_007.pdf)
# - [Software Carpentry version control tutorial](https://swcarpentry.github.io/git-novice/)
#
| notebooks/session_02-git_and_python_basics/04_pandas_data/introduction-to-python-data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # <font color='blue'>UNINOVE - Ciência de Dados</font>
#
# ## Tópico 10 - Python: Matrizes (NumPy)
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
# ### Criando matrizes em NumPy - https://numpy.org
# Importar a biblioteca NumPy.
import numpy as np
# Exemplo de uma matriz NumPy
matriz = np.array([[1,2,3],[4,5,6],[7,8,9],[10,20,30]])
# Imprimindo dados do vetor
print('Matriz:',matriz)
print('Elemento posicao (1,2):',matriz[1][2])
print('Dimensoes:',matriz.ndim)
# Atribuindo valor a um elemento da matriz
matriz[3][1] = 18
print('Matriz:',matriz)
# ### Operações matemáticas com matrizes
# +
# Importar biblioteca
import numpy as np
# Definindo as matrizes
matriz01 = np.array([[1,2,3],[4,5,6],[7,8,9],[10,20,30]])
matriz02 = np.array([[2,2,2], [6,9,-10], [1,7,9] , [3,6,8]])
matriz03 = np.array([[1,1], [0,4],[2,2]])
# -
# Soma
matrizsoma= matriz01 + matriz02
# Subtração
matrizsub = matriz01 - matriz02
# Função para efetuar multiplicação de matrizes: </b>dot</b> - https://numpy.org/doc/stable/reference/generated/numpy.dot.html
matrizprod = matriz01.dot(matriz03)
# #### Imprimindo as matrizes
print('Matriz 01')
print(matriz01)
print('Matriz 02')
print(matriz02)
print('Matriz 03')
print(matriz03)
print('Soma Matriz 01 + Matriz 02')
print(matrizsoma)
print('Subtracao Matriz 01 - Matriz 02')
print(matrizsub)
print('Produto Matriz 01 x Matriz 03')
print(matrizprod)
# Podemos <b>concatenar</b> e <b>dividir</b> matrizes para isso temos as funções <i>concatenate</i> e <i>split</i>.
#
# https://numpy.org/doc/stable/reference/generated/numpy.concatenate.html
#
# https://numpy.org/doc/stable/reference/generated/numpy.split.html
# ### Concatenação de matrizes
# +
# Importar biblioteca
import numpy as np
# Definir matrizes
matriz01 = np.array([[1,2,3],[4,5,6],[7,8,9],[10,20,30]])
matriz02 = np.array([[2,2,2], [6,9,-10], [1,7,9] , [3,6,8]])
matriz03 = np.zeros((1,3))
# Concatenando matrizes
matrizresultante = np.concatenate((matriz01,matriz02,matriz03))
# -
print('Matriz 01')
print(matriz01)
print('Matriz 02')
print(matriz02)
print('Matriz 03')
print(matriz03)
print('Matriz Concetenada')
print(matrizresultante)
# ### Divisão de matrizes - <i>split</i>
#
# Uma observação importantes a ser feita com a utilização da função <i>split</i>. Ela irá apresentar erro caso a quantidade de linhas na matriz resultante não seja exatamente igual em cada uma das partes em que a matriz foi dividida.
# +
# Importar biblioteca
import numpy as np
# Definir matriz
matriz01 = np.array([[1,2,3],[4,5,6],[7,8,9],[10,20,30]])
# Split
matrizresultante = np.split(matriz01,2)
# -
# Imprimindo matrizes
print('Matriz 01')
print(matriz01)
print('Matriz Resultante parte 1')
print(matrizresultante [0])
print('Matriz Resultante parte 2')
print(matrizresultante [1])
# ### Matriz Transposta
#
# Matriz transposta é aquela que é obtida efetuando-se a troca de linhas por colunas de uma dada matriz.
#
# https://numpy.org/doc/stable/reference/generated/numpy.transpose.html
# +
# Importar biblioteca
import numpy as np
# Definindo matriz
matriz = np.array([[10, 20],[30,40],[50,60],[70,80]])
# Gerando a matriz transposta
matrizt = matriz.transpose()
# -
print('Matriz')
print(matriz)
print('Matriz transposta')
print(matrizt)
# ### Operações matemáticas interessantes com matrizes
# +
# Importar biblioteca
import numpy as np
# Definido matriz
matriz = np.array([[1,2,3],[4,5,6],[7,8,9],[10,20,33]])
# -
# <b>max()</b>: recupera o maior elemento da matriz.
#
# https://numpy.org/doc/stable/reference/generated/numpy.matrix.max.html
print('Max:',matriz.max())
# <b>min()</b>: recupera o menor elemento da matriz.
#
# https://numpy.org/doc/stable/reference/generated/numpy.matrix.min.html
print('Min:',matriz.min())
# <b>ptp()</b>: apresenta a diferença entre o maior elemento da matriz e o menor elemento da matriz.
#
# https://numpy.org/doc/stable/reference/generated/numpy.ptp.html
print('Ptp:',matriz.ptp())
# <b>sum()</b>: soma os elementos de uma matriz.
#
# https://numpy.org/doc/stable/reference/generated/numpy.sum.html
print('Sum:',matriz.sum())
# <b>mean()</b>: média aritmética entre os elementos da matriz.
#
# https://numpy.org/doc/stable/reference/generated/numpy.mean.html
print('Mean:',matriz.mean())
# <b>var()</b>: variância entre os elementos da matriz.
#
# https://numpy.org/doc/stable/reference/generated/numpy.var.html
print('Var:',matriz.var())
# <b>std()</b>: variância entre os elementos da matriz.
#
# https://numpy.org/doc/stable/reference/generated/numpy.std.html
print('Std:',matriz.std())
# Podemos redimensionar nossas matrizes com o uso da função <i>resize</i> ou <i>reshape</i>. A diferença entre as duas abordagens é:
#
# <ul>
# <li><b>reshape</b> não altera a matriz original enquanto que <b>resize</b> altera a matriz original.</li>
# <li><b>reshape</b> tem que manter a mesma quantidade de elementos na matriz enquanto que no <b>resize</b> não existe esta obrigatoriedade.</li>
# </ul>
#
# Por exemplo, uma matriz 3x3 (3 linhas e 3 colunas - 9 elementos) não pode ser feito o <i>reshape</i> para 2x5 pois neste caso teríamos 10 elementos, 1 a mais do que na matriz original. Para <i>resize</i> não há problema algum em se efetuar o resize de uma matriz 3x3 para uma matriz 2x5.
# +
# Importar biblioteca
import numpy as np
# Definindo matriz
matriz = np.array([[1,2,3,4,5],[6,7,8,9,10],[11,12,13,14,15]])
# -
# Imprimindo matrizes
print ('matriz antes do Reshape')
print(matriz)
print('Reshape')
print(matriz.reshape(5,3));
print ('matriz após Reshape')
print(matriz)
print('Efetuar Resize')
matriz.resize(8,2)
print('Matriz após resize')
print(matriz)
| uninove/topico_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import os
import glob
import torch
import torch.nn as nn
import torch.nn.functional as F
from sklearn.preprocessing import MinMaxScaler
import pickle
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
# +
HOME_DIR = '/data'
table_path = f'{HOME_DIR}/cricketers.csv'
wikify_column_name = "cricketers"
final_score_column = "siamese_prediction"
canonical_file_path = f'{HOME_DIR}/temp/canonical.csv'
candidate_file_path = f'{HOME_DIR}/temp/candidates.csv'
aux_field = 'graph_embedding_complex,class_count,property_count'
temp_dir = f'{HOME_DIR}/temp/temp'
aligned_pagerank_candidate_file_path = f'{HOME_DIR}/temp/apr_test.csv'
model_file_path = '/table-linker/notebooks/models/weighted_lr.pkl'
ranking_model_file_path = '/table-linker/notebooks/models/epoch_2_loss_0.09150885790586472_top1_0.9067796610169492.pth'
min_max_scaler_path = '/table-linker/notebooks/models/normalization_factor.pkl'
model_voted_candidate_file_path = f'{HOME_DIR}/temp/mv_test.csv'
graph_embedding_file_path = f'{HOME_DIR}/temp/score_test.csv'
lof_reciprocal_rank_file_path = f'{HOME_DIR}/temp/lof_rr_test.csv'
lof_tfidf_file_path = f'{HOME_DIR}/temp/lof_tfidf_test.csv'
lof_feature_file = f'{HOME_DIR}/temp/lof_feature.csv'
output_model_pred_file = f'{HOME_DIR}/temp/model_prediction.csv'
top5_links = f'{HOME_DIR}/temp/top5_links.csv'
colorized_kg_links = f'{HOME_DIR}/temp/colorized_kg_links.xlsx'
graph_embedding_complex_file = f'{HOME_DIR}/temp/graph_embedding_complex.tsv'
class_count_file = f'{HOME_DIR}/temp/class_count.tsv'
property_count_file = f'{HOME_DIR}/temp/property_count.tsv'
index_url = 'http://ckg07:9200/wikidatadwd-augmented/'
# !mkdir -p $temp_dir
# -
# ### Canonicalize
# !tl canonicalize -c "$wikify_column_name" --add-context "$table_path" \
# > "$canonical_file_path"
pd.read_csv(canonical_file_path, nrows = 5)
# ### Candidate Generation
# !tl clean -c label -o label_clean "$canonical_file_path" \
# / --url http://ckg07:9200 --index wikidatadwd-augmented \
# get-fuzzy-augmented-matches -c label_clean \
# --auxiliary-fields "$aux_field" \
# --auxiliary-folder "$temp_dir" \
# / --url http://ckg07:9200 --index wikidatadwd-augmented \
# get-exact-matches \
# -c label_clean --auxiliary-fields "$aux_field" \
# --auxiliary-folder "$temp_dir" > "$candidate_file_path"
column_rename_dict = {
'graph_embedding_complex': 'embedding',
'class_count': 'class_count',
'property_count': 'property_count'
}
for field in aux_field.split(','):
aux_list = []
for f in glob.glob(f'{temp_dir}/*{field}.tsv'):
aux_list.append(pd.read_csv(f, sep='\t'))
aux_df = pd.concat(aux_list).drop_duplicates(subset=['qnode']).rename(columns={field: column_rename_dict[field]})
aux_df.to_csv(f'{HOME_DIR}/temp/{field}.tsv', sep='\t', index=False)
pd.read_csv(candidate_file_path, nrows=5)
# !ls $temp_dir
# ### Generate lof-related features: lof-graph-embedding-score, lof-reciprocal-rank, lof-tfidf
# ##### Generate required 4 features for voting classifier
# !tl align-page-rank $candidate_file_path \
# / string-similarity -i --method symmetric_monge_elkan:tokenizer=word -o monge_elkan \
# / string-similarity -i --method jaro_winkler -o jaro_winkler \
# / string-similarity -i --method levenshtein -o levenshtein \
# / string-similarity -i --method jaccard:tokenizer=word -c kg_descriptions context -o des_cont_jaccard \
# / normalize-scores -c des_cont_jaccard / smallest-qnode-number \
# / mosaic-features -c kg_labels --num-char --num-tokens \
# / create-singleton-feature -o singleton \
# > $aligned_pagerank_candidate_file_path
features_df = pd.read_csv(aligned_pagerank_candidate_file_path)
features_df.loc[:, ['method', 'pagerank', 'aligned_pagerank', 'smallest_qnode_number', 'monge_elkan', 'des_cont_jaccard_normalized']].head()
# ##### Generate model-voted candidates result
# !tl vote-by-classifier $aligned_pagerank_candidate_file_path \
# --prob-threshold 0.995 \
# --model $model_file_path \
# > $model_voted_candidate_file_path
model_voted_df = pd.read_csv(model_voted_candidate_file_path)
model_voted_df.head()
# ##### Generate graph-embedding-score using centroid-of-lof and lof-strategy
# !tl score-using-embedding $model_voted_candidate_file_path \
# --column-vector-strategy centroid-of-lof \
# --lof-strategy ems-mv \
# -o lof-graph-embedding-score \
# --embedding-file $graph_embedding_complex_file \
# --embedding-url $index_url \
# > $graph_embedding_file_path
score_df = pd.read_csv(graph_embedding_file_path)
score_df.head(5)
score_df.sort_values(by=['lof-graph-embedding-score'], ascending=False).loc[:, [
'kg_id', 'kg_labels', 'kg_descriptions', 'method', 'singleton', 'vote_by_classifier', 'is_lof', 'lof-graph-embedding-score'
]].head(20)
# ##### Generate lof reciprocal rank feature
# !tl generate-reciprocal-rank "$graph_embedding_file_path" \
# -c lof-graph-embedding-score \
# -o lof-reciprocal-rank \
# > "$lof_reciprocal_rank_file_path"
pd.read_csv(lof_reciprocal_rank_file_path, nrows=5)
# ##### Generate lof tfidf feature
# !tl compute-tf-idf "$lof_reciprocal_rank_file_path" \
# --feature-file "$class_count_file" \
# --feature-name class_count \
# --singleton-column is_lof \
# -o lof_class_count_tf_idf_score \
# / compute-tf-idf \
# --feature-file "$property_count_file" \
# --feature-name property_count \
# --singleton-column is_lof \
# -o lof_property_count_tf_idf_score \
# > "$lof_feature_file"
d = pd.read_csv(lof_feature_file, nrows=5)
# ### Model Prediction
ranking_model_file_path
# !tl predict-using-model -o siamese_prediction \
# --ranking_model $ranking_model_file_path \
# --normalization_factor $min_max_scaler_path $lof_feature_file > $output_model_pred_file
# ### Get Top 5 links
# !tl get-kg-links -c $final_score_column -k 5 --k-rows $output_model_pred_file > $top5_links
pd.set_option('display.max_rows', None)
final_output = pd.read_csv(top5_links)
final_output[['column', 'row', 'label', 'context', 'kg_id', 'kg_labels', 'kg_aliases',
'kg_descriptions', 'siamese_prediction']]
# ### Colorized KG Links file
# !tl add-color -c $final_score_column -k 5 $top5_links --output $colorized_kg_links
| notebooks/table-linker-wikifier-pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import networkx as nx
from collections import defaultdict
from itertools import groupby
# +
def data_baker_len(filename="words_alpha.txt"):
"""function that takes in a text file of words and bae them into a python list + dict them according to sizes"""
with open(filename) as f:
words = f.read().split()
words_len = defaultdict(list)
for key, values in groupby(words, key=len):
words_len[key] += list(values)
return words_len
def words_gen(filename="words_alpha.txt"):
with open(filename, 'r') as f:
for i in f.read().split():
yield i
def word_cooker(word):
yield "_" + word
for i in range(len(word)):
yield word[:i] + "_" + word[i+1:]
yield word[:i+1] + "_" + word[i+1:]
def grapher(wordFile):
d = {}
g = nx.Graph()
wfile = open(wordFile,'r')
# create buckets of words that differ by one letter
for line in wfile:
word = line[:-1]
for bucket in word_cooker(word):
if bucket in d:
d[bucket].append(word)
else:
d[bucket] = [word]
# add vertices and edges for words in the same bucket
for bucket in d.keys():
for word1 in d[bucket]:
for word2 in d[bucket]:
if word1 != word2:
g.add_edge(word1,word2)
return g
# -
len_dict = data_baker_len()
graph_eq_words = grapher("words_alpha.txt")
len_dict = data_baker_len()
"->".join(nx.astar_path(nx.subgraph(graph_eq_words, len_dict[4]), "head", "tail"))
nx.write_gpickle(graph_eq_words, "equivalent_words_graph.pickle")
a = nx.read_gpickle("equivalent_words_graph.pickle")
print(nx.info(a))
nx.astar_path(a, "head", "tail")
| misc/graph_edit_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] id="pwLNyamAUw0m"
# # Pokemon
# + [markdown] id="O9-ygBTJUw0p"
# ### Introduction:
#
# This time you will create the data.
#
#
#
# ### Step 1. Import the necessary libraries
# + id="OVI65vO0Uw0r"
import numpy as np
import pandas as pd
# + [markdown] id="g6gc-iN8Uw0s"
# ### Step 2. Create a data dictionary that looks like the DataFrame below
# + id="e3qQ_LLnUw0s"
data_dict = {"name": ['Bulbasaur', 'Charmander','Squirtle','Caterpie'],
"evolution": ['Ivysaur','Charmeleon','Wartortle','Metapod'],
"type": ['grass', 'fire', 'water', 'bug'],
"hp": [45, 39, 44, 45],
"pokedex": ['yes', 'no','yes','no']
}
# + [markdown] id="tM0OHA_1Uw0u"
# ### Step 3. Assign it to a variable called pokemon
# + id="54ZVJ_MaUw0u"
pokemon = pd.DataFrame(data_dict)
# + id="M-3JoIuOVBOl" outputId="717b8c91-48f6-41b3-af61-46ef9c1af707" colab={"base_uri": "https://localhost:8080/", "height": 173}
pokemon.head()
# + [markdown] id="_IFrDBuaUw0x"
# ### Step 4. Ops...it seems the DataFrame columns are in alphabetical order. Place the order of the columns as name, type, hp, evolution, pokedex
# + id="oLuvhNIYUw0y"
pokemon = pokemon[['name', 'type', 'hp', 'evolution', 'pokedex']]
# + id="o2_Oki_UVc2q" outputId="bfffa6df-f0e9-44bb-af9f-d78b8297442c" colab={"base_uri": "https://localhost:8080/", "height": 173}
pokemon.head()
# + [markdown] id="SlpgkgHfUw0y"
# ### Step 5. Add another column called place, and insert what you have in mind.
# + id="9KQ3QDpLUw0z"
pokemon['place'] = ['Greece', 'Bolivia', 'Turkey', 'Spain']
# + id="CmQZyyhLVzNx" outputId="f9ff8edb-fd5b-423d-bcf7-8e831c78e6b0" colab={"base_uri": "https://localhost:8080/", "height": 173}
pokemon
# + [markdown] id="LOyX7V3_Uw0z"
# ### Step 6. Present the type of each column
# + id="qemBJ6X2Uw00" outputId="ed3a2340-ded8-42bd-a276-29b3ecf6ede2" colab={"base_uri": "https://localhost:8080/"}
pokemon.info()
| 08_Creating_Series_and_DataFrames/Pokemon/Exercises[Solved].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/misqualzarabi/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/Copy_of_Copy_of_LS_DS_121_Join_and_Reshape_Data_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="pmU5YUal1eTZ"
# _Lambda School Data Science_
#
# # Join and Reshape datasets
#
# Objectives
# - concatenate data with pandas
# - merge data with pandas
# - understand tidy data formatting
# - melt and pivot data with pandas
#
# Links
# - [Pandas Cheat Sheet](https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf)
# - [Tidy Data](https://en.wikipedia.org/wiki/Tidy_data)
# - Combine Data Sets: Standard Joins
# - Tidy Data
# - Reshaping Data
# - Python Data Science Handbook
# - [Chapter 3.6](https://jakevdp.github.io/PythonDataScienceHandbook/03.06-concat-and-append.html), Combining Datasets: Concat and Append
# - [Chapter 3.7](https://jakevdp.github.io/PythonDataScienceHandbook/03.07-merge-and-join.html), Combining Datasets: Merge and Join
# - [Chapter 3.8](https://jakevdp.github.io/PythonDataScienceHandbook/03.08-aggregation-and-grouping.html), Aggregation and Grouping
# - [Chapter 3.9](https://jakevdp.github.io/PythonDataScienceHandbook/03.09-pivot-tables.html), Pivot Tables
#
# Reference
# - Pandas Documentation: [Reshaping and Pivot Tables](https://pandas.pydata.org/pandas-docs/stable/reshaping.html)
# - Modern Pandas, Part 5: [Tidy Data](https://tomaugspurger.github.io/modern-5-tidy.html)
# + id="5MsWLLW4Xg_i" colab_type="code" outputId="b71cd9da-e448-4165-ac8f-470855720589" colab={"base_uri": "https://localhost:8080/", "height": 202}
# !wget https://s3.amazonaws.com/instacart-datasets/instacart_online_grocery_shopping_2017_05_01.tar.gz
# + id="gfr4_Ya0XkLI" colab_type="code" outputId="60261302-5715-43a4-cb9b-97398b4c3903" colab={"base_uri": "https://localhost:8080/", "height": 235}
# !tar --gunzip --extract --verbose --file=instacart_online_grocery_shopping_2017_05_01.tar.gz
# + id="N4YyGPNdXrT0" colab_type="code" outputId="4a0e1b67-6ad8-4cdd-9e42-1407c8a9471a" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd instacart_2017_05_01
# + id="b26wmLUiXtlM" colab_type="code" outputId="12457990-077c-40a9-f462-18271660df0a" colab={"base_uri": "https://localhost:8080/", "height": 118}
# !ls -lh *.csv
# + [markdown] colab_type="text" id="kAMtvSQWPUcj"
# # Assignment
#
# ## Join Data Practice
#
# These are the top 10 most frequently ordered products. How many times was each ordered?
#
# 1. Banana
# 2. Bag of Organic Bananas
# 3. Organic Strawberries
# 4. Organic Baby Spinach
# 5. Organic Hass Avocado
# 6. Organic Avocado
# 7. Large Lemon
# 8. Strawberries
# 9. Limes
# 10. Organic Whole Milk
#
# First, write down which columns you need and which dataframes have them.
#
# Next, merge these into a single dataframe.
#
# Then, use pandas functions from the previous lesson to get the counts of the top 10 most frequently ordered products.
# + id="vvE0EVHgXMFO" colab_type="code" colab={}
import pandas as pd
# + id="_Eq6WkwBk4Oy" colab_type="code" outputId="df1960fc-2209-4d7b-8cf4-6307696996a7" colab={"base_uri": "https://localhost:8080/", "height": 195}
aisles = pd.read_csv('aisles.csv')
aisles.head()
# + id="6l_xQJAwlCHG" colab_type="code" outputId="9375e479-d648-4647-acc4-3f774802cb7f" colab={"base_uri": "https://localhost:8080/", "height": 34}
aisles.shape
# + id="V9Fn7yKDlFZF" colab_type="code" outputId="5c3c3a38-d67b-4891-ec31-13ec43d56369" colab={"base_uri": "https://localhost:8080/", "height": 195}
departments = pd.read_csv('departments.csv')
departments.head()
# + id="bqmiDD87lfB3" colab_type="code" outputId="9de5af46-bf1c-4e53-a099-a5a111ebff4a" colab={"base_uri": "https://localhost:8080/", "height": 34}
departments.shape
# + id="MLVLEkckloxR" colab_type="code" outputId="925be939-557f-4e0e-bfed-2b6dca0a8897" colab={"base_uri": "https://localhost:8080/", "height": 195}
order_products__prior = pd.read_csv('order_products__prior.csv')
order_products__prior.head()
# + id="1681LK6dmvTj" colab_type="code" outputId="036e39e0-7259-4839-afc4-bc13f713eb9c" colab={"base_uri": "https://localhost:8080/", "height": 34}
order_products__prior.shape # we need product_id , reordered
# + id="-JlL0T-wnIQm" colab_type="code" outputId="524305e8-e243-4f36-bd38-4949b550e696" colab={"base_uri": "https://localhost:8080/", "height": 195}
order_products_train = pd.read_csv('order_products__train.csv') # We need product_id, reordered
order_products_train.head()
# + id="mDXd5F5Knbdq" colab_type="code" outputId="6cb3153b-6eb2-4b06-b7c9-1ec8da0a5b6e" colab={"base_uri": "https://localhost:8080/", "height": 34}
order_products_train.shape
# + id="HmcK_uYCm2-V" colab_type="code" outputId="e2616839-adbf-4f3e-a845-645acdd3c8aa" colab={"base_uri": "https://localhost:8080/", "height": 195}
orders = pd.read_csv('orders.csv')
orders.head()
# + id="mDwz4TZOn5lI" colab_type="code" outputId="789297bc-4db2-4088-f71a-3cbd4adae987" colab={"base_uri": "https://localhost:8080/", "height": 34}
orders.shape
# + id="k-MZAjyln_Bk" colab_type="code" outputId="02e2eb6b-68dc-407e-dd2d-34a2823292a3" colab={"base_uri": "https://localhost:8080/", "height": 195}
products = pd.read_csv('products.csv')
products.head() # We need product_id, product_name
# + id="I_KKjzk0oRXk" colab_type="code" outputId="7a5d234b-521b-49d6-ae3e-31f81a7968b6" colab={"base_uri": "https://localhost:8080/", "height": 34}
products.shape
# + id="YzSwqbuzooqy" colab_type="code" outputId="3bf6adce-9f0a-4436-e889-86ede6804701" colab={"base_uri": "https://localhost:8080/", "height": 195}
df = pd.merge(order_products__prior, order_products_train, how='outer')
df.head()
# + id="G_Czfzhnp6tE" colab_type="code" outputId="a7654fe8-61c2-41da-c825-c91d7f0c73e6" colab={"base_uri": "https://localhost:8080/", "height": 195}
final_df = pd.merge(df, products[['product_id', 'product_name']])
final_df.head()
# + id="FyYEAUUrpEnk" colab_type="code" outputId="835338e7-a123-49de-bf1b-a8a3a10b5a76" colab={"base_uri": "https://localhost:8080/", "height": 1000}
final_df.drop(['order_id','add_to_cart_order'], axis=1)
# + id="NBoUyPUftuUn" colab_type="code" colab={}
columns = ['product_id','product_name','reordered']
final_df = final_df[columns]
# + id="A0gARkRGuKef" colab_type="code" outputId="7909fc74-9312-40ad-d331-cc5f43ab9f75" colab={"base_uri": "https://localhost:8080/", "height": 1000}
final_df
# + id="rO9TDXIyukgN" colab_type="code" outputId="62dd82a3-853e-4609-d600-e1020fac3273" colab={"base_uri": "https://localhost:8080/", "height": 84}
final_df.count()
# + id="Ok_w9Rogx66u" colab_type="code" outputId="d19c6eac-264f-4798-caed-455956939725" colab={"base_uri": "https://localhost:8080/", "height": 1000}
final_df.groupby(["product_name"]).sum().sort_values("reordered", ascending=False)
# + [markdown] id="RsiWi4DuXPLP" colab_type="text"
# ## Reshape Data Section
#
# - Replicate the lesson code
# - Complete the code cells we skipped near the beginning of the notebook
# - Table 2 --> Tidy
# - Tidy --> Table 2
# - Load seaborn's `flights` dataset by running the cell below. Then create a pivot table showing the number of passengers by month and year. Use year for the index and month for the columns. You've done it right if you get 112 passengers for January 1949 and 432 passengers for December 1960.
# + id="MGqR3oHDkn5E" colab_type="code" colab={}
# %matplotlib inline
import pandas as pd
import numpy as np
import seaborn as sns
table1 = pd.DataFrame(
[[np.nan, 2],
[16, 11],
[3, 1]],
index=['<NAME>', '<NAME>', '<NAME>'],
columns=['treatmenta', 'treatmentb'])
table2 = table1.T
# + id="hvmtkuQw2kYN" colab_type="code" outputId="679d8603-f595-403e-bc18-19723f8741ec" colab={"base_uri": "https://localhost:8080/", "height": 136}
table1
# + id="hz6JyxR42zGH" colab_type="code" outputId="87536a89-ca89-409c-8c38-3da712ddf6ff" colab={"base_uri": "https://localhost:8080/", "height": 106}
table2
# + [markdown] id="H5-yR5bb3Ptz" colab_type="text"
# **Table 1 --> Tidy**
# + id="fK0yJIvO2-0g" colab_type="code" outputId="7704e4f7-7cfe-4e0a-ddbf-4b94ec741bf0" colab={"base_uri": "https://localhost:8080/", "height": 136}
table1
# + id="oF_MQ8Hf3t_U" colab_type="code" outputId="7c277f66-71a1-4dde-fdc6-866c49552a73" colab={"base_uri": "https://localhost:8080/", "height": 34}
table1.index
# + id="lEwq3jSx3_fY" colab_type="code" colab={}
table1 = table1.reset_index()
# + id="2d9jVQM34IDy" colab_type="code" outputId="233db80b-b17b-4b22-b851-fae391d13fe3" colab={"base_uri": "https://localhost:8080/", "height": 136}
table1
# + id="HAeB5AvQ4NSJ" colab_type="code" outputId="485ad4a9-768f-474b-e191-4045fe3a626f" colab={"base_uri": "https://localhost:8080/", "height": 225}
tidy = table1.melt(id_vars='index')
tidy
# + id="7MUtZqor5ODA" colab_type="code" outputId="1f71d0a3-32b8-4c35-fabf-7cc159f02813" colab={"base_uri": "https://localhost:8080/", "height": 225}
tidy.columns = ['name','trt','result']
tidy
# + [markdown] id="Y7hjycX_AaMd" colab_type="text"
# **Table 2 --> Tidy**
# + id="KE2OjDD85aj_" colab_type="code" outputId="445b8080-a90e-4d68-9de4-bd9be87c2b6a" colab={"base_uri": "https://localhost:8080/", "height": 106}
table2
# + id="2xfsDcsE5wHQ" colab_type="code" outputId="dc08d8ef-63e1-4d33-8bd3-9af70597fb50" colab={"base_uri": "https://localhost:8080/", "height": 34}
table2.index
# + id="K69NMBor6eI-" colab_type="code" colab={}
table2 = table2.reset_index()
# + id="a8NA2dhP7zol" colab_type="code" outputId="454b83de-9f67-4c1c-d915-b44820a7f8c0" colab={"base_uri": "https://localhost:8080/", "height": 106}
table2
# + id="FLe8VZbs7367" colab_type="code" outputId="d3dde8e9-0960-4a33-c036-acd3020da348" colab={"base_uri": "https://localhost:8080/", "height": 225}
tidy2 = table2.melt(id_vars='index')
tidy2
# + id="CAxVGSMq8vSr" colab_type="code" outputId="58f0502f-5a95-4bee-c49b-587a15834b26" colab={"base_uri": "https://localhost:8080/", "height": 225}
tidy2.columns = ['trt','name','result']
tidy2
# + [markdown] id="7Vas4LmoAvAL" colab_type="text"
# **Tidy --> Table 1**
# + id="OfmSuU_FAWbP" colab_type="code" outputId="c1240b87-951e-443c-9597-1799181fc114" colab={"base_uri": "https://localhost:8080/", "height": 136}
table1
# + id="ae_yq-vBBPTg" colab_type="code" outputId="45716c6d-8646-4354-d27a-5e177ad4a90c" colab={"base_uri": "https://localhost:8080/", "height": 166}
tidy.pivot_table(index='name', columns='trt', values='result')
# + [markdown] id="ZH6N8-DoD1ja" colab_type="text"
# **Tidy --> Table 2**
# + id="FAXaRioSBiG_" colab_type="code" outputId="07684403-3682-4091-ed00-1d055be70c40" colab={"base_uri": "https://localhost:8080/", "height": 106}
table2
# + id="8roFJC19BnBq" colab_type="code" outputId="c8e6578d-fab0-4e53-b2c4-f2204e82ae44" colab={"base_uri": "https://localhost:8080/", "height": 136}
tidy2.pivot_table(index='trt',columns='name',values='result')
# + id="fgxulJQq0uLw" colab_type="code" colab={}
flights = sns.load_dataset('flights')
# + id="adHjeWEQEGH0" colab_type="code" outputId="ead75624-ce4b-449b-d5e1-30fda9098596" colab={"base_uri": "https://localhost:8080/", "height": 1000}
flights
# + id="cbhw3t4BE4y1" colab_type="code" outputId="a596a3b0-2076-4905-d7f7-680f4e586c45" colab={"base_uri": "https://localhost:8080/", "height": 462}
flights.pivot_table(index='year', columns='month')
# + [markdown] id="mnOuqL9K0dqh" colab_type="text"
# ## Join Data Stretch Challenge
#
# The [Instacart blog post](https://tech.instacart.com/3-million-instacart-orders-open-sourced-d40d29ead6f2) has a visualization of "**Popular products** purchased earliest in the day (green) and latest in the day (red)."
#
# The post says,
#
# > "We can also see the time of day that users purchase specific products.
#
# > Healthier snacks and staples tend to be purchased earlier in the day, whereas ice cream (especially Half Baked and The Tonight Dough) are far more popular when customers are ordering in the evening.
#
# > **In fact, of the top 25 latest ordered products, the first 24 are ice cream! The last one, of course, is a frozen pizza.**"
#
# Your challenge is to reproduce the list of the top 25 latest ordered popular products.
#
# We'll define "popular products" as products with more than 2,900 orders.
#
#
# + id="B-QNMrVkYap4" colab_type="code" colab={}
##### YOUR CODE HERE #####
# + [markdown] id="Ij8S60q0YXxo" colab_type="text"
# ## Reshape Data Stretch Challenge
#
# _Try whatever sounds most interesting to you!_
#
# - Replicate more of Instacart's visualization showing "Hour of Day Ordered" vs "Percent of Orders by Product"
# - Replicate parts of the other visualization from [Instacart's blog post](https://tech.instacart.com/3-million-instacart-orders-open-sourced-d40d29ead6f2), showing "Number of Purchases" vs "Percent Reorder Purchases"
# - Get the most recent order for each user in Instacart's dataset. This is a useful baseline when [predicting a user's next order](https://www.kaggle.com/c/instacart-market-basket-analysis)
# - Replicate parts of the blog post linked at the top of this notebook: [Modern Pandas, Part 5: Tidy Data](https://tomaugspurger.github.io/modern-5-tidy.html)
# + id="_d6IA2R0YXFY" colab_type="code" colab={}
##### YOUR CODE HERE #####
| Copy_of_Copy_of_LS_DS_121_Join_and_Reshape_Data_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Node representation learning with Metapath2Vec
#
# + [markdown] nbsphinx="hidden" tags=["CloudRunner"]
# <table><tr><td>Run the latest release of this notebook:</td><td><a href="https://mybinder.org/v2/gh/stellargraph/stellargraph/master?urlpath=lab/tree/demos/embeddings/metapath2vec-embeddings.ipynb" alt="Open In Binder" target="_parent"><img src="https://mybinder.org/badge_logo.svg"/></a></td><td><a href="https://colab.research.google.com/github/stellargraph/stellargraph/blob/master/demos/embeddings/metapath2vec-embeddings.ipynb" alt="Open In Colab" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg"/></a></td></tr></table>
# -
# An example of implementing the Metapath2Vec representation learning algorithm using components from the `stellargraph` and `gensim` libraries.
#
# **References**
#
# **1.** Metapath2Vec: Scalable Representation Learning for Heterogeneous Networks. <NAME>, <NAME>, and <NAME>. ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (KDD), 135–144, 2017. ([link](https://ericdongyx.github.io/papers/KDD17-dong-chawla-swami-metapath2vec.pdf))
#
# **2.** Distributed representations of words and phrases and their compositionality. <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. In Advances in Neural Information Processing Systems (NIPS), pp. 3111-3119, 2013. ([link](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf))
#
# **3.** Gensim: Topic modelling for humans. ([link](https://radimrehurek.com/gensim/))
#
# **4.** Social Computing Data Repository at ASU [http://socialcomputing.asu.edu]. <NAME> and <NAME>, AZ: Arizona State University, School of Computing, Informatics and Decision Systems Engineering. 2009.
# + nbsphinx="hidden" tags=["CloudRunner"]
# install StellarGraph if running on Google Colab
import sys
if 'google.colab' in sys.modules:
# %pip install -q stellargraph[demos]==1.3.0b
# + nbsphinx="hidden" tags=["VersionCheck"]
# verify that we're using the correct version of StellarGraph for this notebook
import stellargraph as sg
try:
sg.utils.validate_notebook_version("1.3.0b")
except AttributeError:
raise ValueError(
f"This notebook requires StellarGraph version 1.3.0b, but a different version {sg.__version__} is installed. Please see <https://github.com/stellargraph/stellargraph/issues/1172>."
) from None
# +
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
from sklearn.decomposition import PCA
import os
import networkx as nx
import numpy as np
import pandas as pd
from stellargraph import datasets
from IPython.display import display, HTML
# %matplotlib inline
# -
# ## Load the dataset
# + [markdown] tags=["DataLoadingLinks"]
# (See [the "Loading from Pandas" demo](../basics/loading-pandas.ipynb) for details on how data can be loaded.)
# + tags=["DataLoading"]
dataset = datasets.BlogCatalog3()
display(HTML(dataset.description))
g = dataset.load()
print(
"Number of nodes {} and number of edges {} in graph.".format(
g.number_of_nodes(), g.number_of_edges()
)
)
# -
# ## The Metapath2Vec algorithm
#
# The Metapath2Vec algorithm introduced in [1] is a 2-step representation learning algorithm. The two steps are:
#
# 1. Use uniform random walks to generate sentences from a graph. A sentence is a list of node IDs. The set of all sentences makes a corpus. The random walk is driven by a metapath that defines the node type order by which the random walker explores the graph.
#
# 2. The corpus is then used to learn an embedding vector for each node in the graph. Each node ID is considered a unique word/token in a dictionary that has size equal to the number of nodes in the graph. The Word2Vec algorithm [2] is used for calculating the embedding vectors.
# ## Corpus generation using random walks
#
# The `stellargraph` library provides an implementation for uniform, first order, random walks as required by Metapath2Vec. The random walks have fixed maximum length and are controlled by the list of metapath schemas specified in parameter `metapaths`.
#
# A metapath schema defines the type of node that the random walker is allowed to transition to from its current location. In the `stellargraph` implementation of metapath-driven random walks, the metapath schemas are given as a list of node types under the assumption that the input graph is not a multi-graph, i.e., two nodes are only connected by one edge type.
#
# See [1] for a detailed description of metapath schemas and metapath-driven random walks.
#
# For the **BlogCatalog3** dataset we use the following 3 metapaths.
#
# - "user", "group", "user"
# - "user", "group", "user", "user"
# - "user", "user"
#
#
# + tags=["parameters"]
walk_length = 100 # maximum length of a random walk to use throughout this notebook
# specify the metapath schemas as a list of lists of node types.
metapaths = [
["user", "group", "user"],
["user", "group", "user", "user"],
["user", "user"],
]
# +
from stellargraph.data import UniformRandomMetaPathWalk
# Create the random walker
rw = UniformRandomMetaPathWalk(g)
walks = rw.run(
nodes=list(g.nodes()), # root nodes
length=walk_length, # maximum length of a random walk
n=1, # number of random walks per root node
metapaths=metapaths, # the metapaths
)
print("Number of random walks: {}".format(len(walks)))
# -
# ## Representation Learning using Word2Vec
#
# We use the Word2Vec [2] implementation in the free Python library gensim [3] to learn representations for each node in the graph.
#
# We set the dimensionality of the learned embedding vectors to 128 as in [1].
# +
from gensim.models import Word2Vec
model = Word2Vec(walks, size=128, window=5, min_count=0, sg=1, workers=2, iter=1)
# -
model.wv.vectors.shape # 128-dimensional vector for each node in the graph
# ## Visualise Node Embeddings
#
# We retrieve the Word2Vec node embeddings that are 128-dimensional vectors and then we project them down to 2 dimensions using the [t-SNE](http://scikit-learn.org/stable/modules/generated/sklearn.manifold.TSNE.html) algorithm.
# Retrieve node embeddings and corresponding subjects
node_ids = model.wv.index2word # list of node IDs
node_embeddings = (
model.wv.vectors
) # numpy.ndarray of size number of nodes times embeddings dimensionality
node_targets = [g.node_type(node_id) for node_id in node_ids]
# Transform the embeddings to 2d space for visualisation
# +
transform = TSNE # PCA
trans = transform(n_components=2)
node_embeddings_2d = trans.fit_transform(node_embeddings)
# +
# draw the points
label_map = {l: i for i, l in enumerate(np.unique(node_targets))}
node_colours = [label_map[target] for target in node_targets]
plt.figure(figsize=(20, 16))
plt.axes().set(aspect="equal")
plt.scatter(node_embeddings_2d[:, 0], node_embeddings_2d[:, 1], c=node_colours, alpha=0.3)
plt.title("{} visualization of node embeddings".format(transform.__name__))
plt.show()
# -
# ## Downstream task
#
# The node embeddings calculated using Metapath2Vec can be used as feature vectors in a downstream task such as node attribute inference (e.g., inferring the gender or age attribute of 'user' nodes), community detection (e.g., clustering of 'user' nodes based on the similarity of their embedding vectors), and link prediction (e.g., prediction of friendship relation between 'user' nodes).
# + [markdown] nbsphinx="hidden" tags=["CloudRunner"]
# <table><tr><td>Run the latest release of this notebook:</td><td><a href="https://mybinder.org/v2/gh/stellargraph/stellargraph/master?urlpath=lab/tree/demos/embeddings/metapath2vec-embeddings.ipynb" alt="Open In Binder" target="_parent"><img src="https://mybinder.org/badge_logo.svg"/></a></td><td><a href="https://colab.research.google.com/github/stellargraph/stellargraph/blob/master/demos/embeddings/metapath2vec-embeddings.ipynb" alt="Open In Colab" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg"/></a></td></tr></table>
| demos/embeddings/metapath2vec-embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dopplerchase/DRpy/blob/master/notebooks/Example_one.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="9MeygYrcTopZ" colab_type="text"
# # Welcome to DRpy (derpy)
#
# Here is a simple notebook to help you get going.
# + [markdown] id="eaOjXXDdTope" colab_type="text"
# ### If you want mapping, please install cartopy
# + id="JW5WeHjUTopg" colab_type="code" outputId="4eb11e63-ab31-4324-c38e-2f75e835e0bc" colab={"base_uri": "https://localhost:8080/", "height": 717}
#first install cartopy
# !apt-get -qq install libproj-dev proj-data proj-bin libgeos-dev
# !pip install Cython
# !pip install cartopy
# + [markdown] id="nNSsCzC-Top0" colab_type="text"
# ### Ok, now go clone the repository
#
# The reason we are cloning the directory is to grab the example file. If you don't need this file, just install this way:
#
# # # !pip install git+https://github.com/dopplerchase/DRpy.git
# + id="92DDacVTTop5" colab_type="code" outputId="3a0f570d-6607-4209-9084-036fb18ea2dc" colab={"base_uri": "https://localhost:8080/", "height": 136}
#clone repository
# !git clone https://github.com/dopplerchase/DRpy.git
# + [markdown] id="qv_45F8uToqB" colab_type="text"
# ### Install it
# + id="e3UKu8rTToqI" colab_type="code" outputId="a53d53a3-7f30-4495-9aed-8d79789e13a6" colab={"base_uri": "https://localhost:8080/", "height": 561}
import os
# #cd into folder
os.chdir('./DRpy/')
#install
# !python setup.py install
# #cd into notebooks
os.chdir('./notebooks/')
# + [markdown] id="sqMGdbJ0ToqP" colab_type="text"
# ## Now we can actually use it!
# + id="gBOdWeQ5ToqS" colab_type="code" outputId="4e77df1d-23ed-41d3-bf60-8a89144a0906" colab={"base_uri": "https://localhost:8080/", "height": 34}
import drpy
import time
# %pylab inline
# + id="MYR7QqdOToqc" colab_type="code" colab={}
filename = '../example_file/2A.GPM.DPR.V820180723.20200117-S181128-E184127.V06A.RT-H5'
# + id="ArLtGDf1Toql" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6913aee4-945f-4735-9c13-3eccecbb6626"
stime = time.time()
dpr = drpy.core.GPMDPR(filename = filename)
dpr.read()
dpr.toxr()
etime = time.time()
print('time elapsed {} seconds'.format(etime-stime))
# + [markdown] id="RKHRtzfLToqw" colab_type="text"
# ### lets take a gander at the xarray datatset (.xrds)
#
# xrds stands for xarray dataset
# We can see that the data have the original shape of the GPM-DPR data
#
#
# along_track,cross_track,range
# + id="gVFhoGmIToqz" colab_type="code" outputId="dd60dbd2-67cf-4121-d81a-b8a26510968a" colab={"base_uri": "https://localhost:8080/", "height": 374}
dpr.xrds
# + [markdown] id="a5kGQpbUtdL8" colab_type="text"
# ### let us plot a map of the data, to see where the swath is in the world
# + id="B2LhkabitcjS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="760e068c-ddac-4d9e-cd1b-d45a3c26df85"
import cartopy
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import matplotlib.ticker as mticker
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
import matplotlib.patheffects as PathEffects
import cartopy.io.shapereader as shpreader
from cartopy.mpl.geoaxes import GeoAxes
from mpl_toolkits.axes_grid1 import AxesGrid
from cartopy.mpl.ticker import LongitudeFormatter, LatitudeFormatter
import matplotlib.colors as colors
import matplotlib.patheffects as PathEffects
#make figure
fig = plt.figure(figsize=(10, 10))
#add the map
ax = fig.add_subplot(1, 1, 1,projection=ccrs.PlateCarree())
ax.add_feature(cfeature.STATES.with_scale('50m'),lw=0.5)
ax.add_feature(cartopy.feature.OCEAN.with_scale('50m'))
ax.add_feature(cartopy.feature.LAND.with_scale('50m'), edgecolor='black',lw=0.5,facecolor=[0.95,0.95,0.95])
ax.add_feature(cartopy.feature.LAKES.with_scale('50m'), edgecolor='black')
ax.add_feature(cartopy.feature.RIVERS.with_scale('50m'))
ax.pcolormesh(dpr.xrds.lons,dpr.xrds.lats,dpr.xrds.nearsurfaceKu,vmin=12,vmax=50,cmap=drpy.graph.cmaps.HomeyerRainbow)
# + [markdown] id="Bfw33Sodwq6B" colab_type="text"
# you can see that the data is quite spread out. Say you are interested in the data near St. Johns Newfoundland, Canada. Lets cut the data to a box around it.
#
# There are two ways to do this, you can reprocess the original file, or you can use the method to cut the box.
# + id="djUlZULIs3Ya" colab_type="code" colab={}
#define box
center_lat = 47.5615
center_lon = -52.7126
corners = [center_lon - 4,center_lon +4, center_lat-1.8,center_lat+1.8]
#
dpr.corners = corners
#method 2: Cut the exisiting datset
dpr.setboxcoords()
#drop dead weight (i.e. blank data)
dpr.xrds = dpr.xrds.dropna(dim='along_track',how='all')
# + id="c3dojbDyxc2o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 514} outputId="c110a661-96e0-440a-8253-ac81056dce70"
import cartopy
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import matplotlib.ticker as mticker
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
import matplotlib.patheffects as PathEffects
import cartopy.io.shapereader as shpreader
from cartopy.mpl.geoaxes import GeoAxes
from mpl_toolkits.axes_grid1 import AxesGrid
from cartopy.mpl.ticker import LongitudeFormatter, LatitudeFormatter
import matplotlib.colors as colors
import matplotlib.patheffects as PathEffects
#make figure
fig = plt.figure(figsize=(10, 10))
#add the map
ax = fig.add_subplot(1, 1, 1,projection=ccrs.PlateCarree())
ax.add_feature(cfeature.STATES.with_scale('50m'),lw=0.5)
ax.add_feature(cartopy.feature.OCEAN.with_scale('50m'))
ax.add_feature(cartopy.feature.LAND.with_scale('50m'), edgecolor='black',lw=0.5,facecolor=[0.95,0.95,0.95])
ax.add_feature(cartopy.feature.LAKES.with_scale('50m'), edgecolor='black')
ax.add_feature(cartopy.feature.RIVERS.with_scale('50m'))
ax.pcolormesh(dpr.xrds.lons,dpr.xrds.lats,dpr.xrds.nearsurfaceKu,vmin=12,vmax=50,cmap=drpy.graph.cmaps.HomeyerRainbow)
# + id="ppTkKroHxfE0" colab_type="code" colab={}
| notebooks/Example_one.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Oversampling using SMOTE
# ## Import and read data
# +
from numpy import where
import matplotlib.pyplot as plt
from collections import Counter
from sklearn.datasets import make_classification
from imblearn.over_sampling import SMOTE
# -
# ## Create an imbalanced dataset
X, y = make_classification(n_samples=1000, n_features=2, n_redundant=0, n_clusters_per_class=1, weights=[0.975], flip_y=0, random_state=1)
counter=Counter(y)
counter
for label, _ in counter.items():
row_ix = where(y == label)[0]
plt.scatter(X[row_ix, 0], X[row_ix, 1], label=str(label))
plt.legend()
plt.show()
# ## Oversample using SMOTE
oversample = SMOTE()
X, y = oversample.fit_resample(X, y)
counter=Counter(y)
counter
for label, _ in counter.items():
row_ix = where(y == label)[0]
plt.scatter(X[row_ix, 0], X[row_ix, 1], label=str(label))
plt.legend()
plt.show()
| M6_Machine_Learning/M6_Machine_Learning_W2_Supervised_Learning_Ensemble_Techniques_Sampling_Techniques/smote.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (conda)
# language: python
# name: python3
# ---
# # Example: noisy identity process on two qubits
#
# This example is featured in the paper.
#
#
# **Contents:**
# * [Python initializations](#Python-initializations)
# * [Data set-up](#Data-set-up)
# * [Bipartite sampling method](#Bipartite-sampling-method)
# * [Bipartite sampling method, optimized](#Bipartite-sampling-method,-optimized)
# * [Fit with our empirical model \#2](#Fit-with-our-empirical-model-#2)
# * [Bipartite sampling method, optimize, entanglement fidelity](#Bipartite-sampling-method,-optimized,-entanglement-fidelity)
# * [Bipartite sampling method, optimize, worst-case entanglement fidelity](#Bipartite-sampling-method,-optimized,-worst-case-entanglement-fidelity)
# * [Prepare data for channel-space methods](#Prepare-data-for-channel-space-methods)
# * [Channel-space method, "$e^{iH}$" variant](#Channel-space-method,-%22$e^{iH}$%22-variant)
# * [Channel-space method, "elementary rotations" variant](#Channel-space-method,-%22elementary-rotation%22-variant)
# * [Channel-space method, "elementary rotations" variant, entanglement fidelity](#Channel-space-method,-%22elementary-rotation%22-variant,-entanglement-fidelity)
# * [Channel-space method, "elementary rotations" variant, worst-case entanglement fidelity](#Channel-space-method,-%22elementary-rotation%22-variant,-worst-case-entanglement-fidelity)
# * [Grand comparison plots](#Grand-comparison-plots)
#
#
# ## Python initializations
# +
from __future__ import print_function
import os.path
import sys
import datetime
import numpy as np
import numpy.linalg as npl
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.font_manager
matplotlib.rc('font', family='Arial')
from IPython.display import display, Markdown, Latex
import QPtomographer.channelspace
import QPtomographer.bistates
import QPtomographer.util
# use this to load pickle files saved with our old python code
sys.modules['pydnorm_util'] = QPtomographer.util
sys.modules['dnormtomo'] = QPtomographer
QPtomographer.util._Ns = QPtomographer.util._Store
import qutip
import tomographer
import tomographer.jpyutil
import tomographer.querrorbars
display(Markdown("Using `QPtomographer` **{}** with `tomographer` **{}**"
.format(QPtomographer.__version__, tomographer.__version__)))
# to save & load computation results
try:
import cPickle as pickle
except:
import pickle
# interact with plots in notebook
# %matplotlib notebook
# -
# utilities for storing & loading calculation results
def save_to_cache(cachesuffix, x):
with open('_CACHE_'+cachesuffix+'.dat', 'wb') as f:
pickle.dump(x, f, 2)
def load_from_cache(cachesuffix):
cachefile = '_CACHE_'+cachesuffix+'.dat'
if not os.path.exists(cachefile):
return None
with open(cachefile, 'rb') as f:
display(Markdown("Loading `{}` from cache".format(cachesuffix)))
return pickle.load(f)
# ## Data set-up
# +
#
# number of outcomes per Pauli pairs
#
NumSamplesPerSetting = 500
# Pauli measurement settings on one system
PauliMeasSettings = [
[
qutip.Qobj(qutip.tensor(QPtomographer.util.projpauli(i1, s1),
QPtomographer.util.projpauli(i2, s2)).data.toarray(),
dims=[[4],[4]])
for s1 in [1, -1]
for s2 in [1, -1]
]
for i1 in [1, 2, 3]
for i2 in [1, 2, 3]
]
#display(PauliMeasSettings)
#
# the "real" states & process from which we simulate outcomes
#
sigmareal_X = qutip.Qobj(np.array([[ 0.35, 0.00, 0.04, 0.1j],
[ 0.00, 0.15, 0.05, 0.00],
[ 0.04, 0.05, 0.32, 0.00],
[-0.1j, 0.00, 0.00, 0.18]]), dims=[[4],[4]])
display(Markdown("Eigenvalues of sigmareal_X = $[" +
",".join("{:.4g}".format(x) for x in npl.eigvalsh(sigmareal_X.data.toarray()))
+ "]$"))
#display(sigmareal_X.eigenstates())
MaxEntgl_XY = qutip.Qobj(np.array([ [ 1. if i==j else 0. ] for i in range(4) for j in range(4)]),
dims=[[4,4],[1,1]])
#display(MaxEntgl_XY.data.toarray())
Ereal_XY = 0.9*MaxEntgl_XY*MaxEntgl_XY.dag() + 0.1*qutip.Qobj(np.identity(16), dims=[[4,4],[4,4]])/4
#display(Ereal_XY.tr())#.data.toarray().diagonal())
#rho_AB = pydnorm_util.process_matrix(sigmareal_X, Ereal_XY)
#display(rho_AB)
#display(rho_AB.data.diagonal())
#display(rho_AB.tr())
def simulate_measurements():
#
# Simulate the outcomes
#
d = QPtomographer.util.simulate_process_measurements(sigmareal_X, Ereal_XY,
PauliMeasSettings,
PauliMeasSettings,
NumSamplesPerSetting)
return d
#
# Only simulate the measurements once. After that, use the same data when comparing methods!!
#
d = load_from_cache('meas_data')
if d is None:
d = simulate_measurements()
save_to_cache('meas_data', d)
display(Markdown("We have **{}** total measurements".format(sum(d.Nm))))
#print(d.__dict__) # prints Emn, Nm ... long outputs!!
# +
#
# Stuff for the analysis, later
#
def print_report(r):
display(Markdown("Calculation ran for **{!s} seconds**".format(datetime.timedelta(seconds=r['elapsed_seconds']))))
display(Markdown("```\n{}\n```".format(r['final_report_runs'])))
def do_analysis(r, name, plots=False, ftox=(0,1), qtyname='diamond distance'):
final_histogram = r['final_histogram']
analysis = tomographer.querrorbars.HistogramAnalysis(final_histogram, ftox=ftox)
fitparams = analysis.printFitParameters()
analysis.printQuantumErrorBars()
p1 = None
p2 = None
if plots:
p1 = analysis.plot(show_plot=False) # linear scale
p1.ax.set_title("Distribution of the %s: %s"%(qtyname, name))
p1.ax.set_xlabel('Diamond Norm distance to the identity channel')
p1.show()
p2 = analysis.plot(log_scale=True, show_plot=False) # log scale
p2.ax.set_title("Distribution of the %s: %s"%(qtyname, name))
p2.ax.set_xlabel('Diamond Norm distance to the identity channel')
p2.show()
return {'r': r, 'name': name, 'analysis': analysis, 'fitparams': fitparams, 'p1': p1, 'p2': p2}
# -
# ## Bipartite sampling method
# +
r_naive = load_from_cache('r_naive')
if r_naive is None:
# perform calculation
with tomographer.jpyutil.RandWalkProgressBar() as prg:
r_naive = QPtomographer.bistates.run(
dimX=4, dimY=4, Emn=d.Emn, Nm=np.array(d.Nm),
hist_params=tomographer.UniformBinsHistogramParams(0.1, 0.3, 100),
mhrw_params=tomographer.MHRWParams(0.001, 1000, 2048, 32768),
progress_fn=prg.progress_fn,
progress_interval_ms=2000,
)
prg.displayFinalInfo(r_naive['final_report_runs'])
save_to_cache('r_naive', r_naive)
print_report(r_naive)
# -
a_naive = do_analysis(r_naive, 'st., std.', plots=True)
# ## Bipartite sampling method, optimized
# +
r_naiveopt = load_from_cache('r_naiveopt')
if r_naiveopt is None:
# perform calculation
with tomographer.jpyutil.RandWalkProgressBar() as prg:
r_naiveopt = QPtomographer.bistates.run(
dimX=4, dimY=4, Emn=d.Emn, Nm=np.array(d.Nm),
hist_params=tomographer.UniformBinsHistogramParams(0.1, 0.3, 100),
mhrw_params=tomographer.MHRWParams(0.001, 4000, 8192, 32768),
progress_fn=prg.progress_fn,
progress_interval_ms=2000,
jumps_method='light',
)
prg.displayFinalInfo(r_naiveopt['final_report_runs'])
save_to_cache('r_naiveopt', r_naiveopt)
print_report(r_naiveopt)
# -
a_naiveopt = do_analysis(r_naiveopt, 'st., opt.', plots=True)
# ## Fit with our empirical model \#2
# +
def fit_fn_test(x, a2, a1, m, p, c):
return -a2* np.square(x) - a1*x - m*np.power(-np.log(x),p) + c
a = tomographer.querrorbars.HistogramAnalysis(
r_naiveopt['final_histogram'], fit_fn=fit_fn_test, bounds=((0,-np.inf,0,0,-np.inf), np.inf)
)
a.printFitParameters()
a.plot(plot_deskewed_gaussian=False, log_scale=True)
# -
# ## Bipartite sampling method, optimized, entanglement fidelity
# +
r_Fe_naiveopt = load_from_cache('r_Fe_naiveopt')
if r_Fe_naiveopt is None:
# perform calculation
with tomographer.jpyutil.RandWalkProgressBar() as prg:
r_Fe_naiveopt = QPtomographer.bistates.run(
dimX=4, dimY=4, Emn=d.Emn, Nm=np.array(d.Nm),
fig_of_merit='entanglement-fidelity',
hist_params=tomographer.UniformBinsHistogramParams(0.8, 0.9, 100),
mhrw_params=tomographer.MHRWParams(0.001, 4000, 8192, 32768),
progress_fn=prg.progress_fn,
progress_interval_ms=2000,
jumps_method='light',
)
prg.displayFinalInfo(r_Fe_naiveopt['final_report_runs'])
save_to_cache('r_Fe_naiveopt', r_Fe_naiveopt)
print_report(r_Fe_naiveopt)
# -
a_Fe_naiveopt = do_analysis(r_Fe_naiveopt, 'st., opt.', plots=True,
qtyname='entanglement fidelity', ftox=(1,-1))
# ## Bipartite sampling method, optimized, worst-case entanglement fidelity
# +
r_wFe_naiveopt = load_from_cache('r_wFe_naiveopt')
if r_wFe_naiveopt is None:
# perform calculation
with tomographer.jpyutil.RandWalkProgressBar() as prg:
r_wFe_naiveopt = QPtomographer.bistates.run(
dimX=4, dimY=4, Emn=d.Emn, Nm=np.array(d.Nm),
fig_of_merit='worst-entanglement-fidelity',
hist_params=tomographer.UniformBinsHistogramParams(0.75, 0.9, 150),
mhrw_params=tomographer.MHRWParams(0.001, 4000, 8192, 32768),
progress_fn=prg.progress_fn,
progress_interval_ms=2000,
jumps_method='light',
)
prg.displayFinalInfo(r_Fe_naiveopt['final_report_runs'])
save_to_cache('r_wFe_naiveopt', r_wFe_naiveopt)
print_report(r_wFe_naiveopt)
# -
a_wFe_naiveopt = do_analysis(r_wFe_naiveopt, 'st., opt., went.', plots=True,
qtyname='worst-case entanglement fidelity', ftox=(1,-1))
# ## Prepare data for channel-space methods
# +
# we need to encode the input state in the POVM effects
sigmareal_X_sqrtm_eyeY = np.kron(sigmareal_X.sqrtm().data.toarray(), np.eye(4))
Emn_for_channelspace = [
np.dot(np.dot(sigmareal_X_sqrtm_eyeY, E), sigmareal_X_sqrtm_eyeY)
for E in d.Emn
]
# -
# ## Channel-space method, "$e^{iH}$" variant
r_eiH = load_from_cache('r_eiH')
if r_eiH is None:
# no stored result, perform computation
with tomographer.jpyutil.RandWalkProgressBar() as prg:
r_eiH = QPtomographer.channelspace.run(
dimX=4, dimY=4, Emn=Emn_for_channelspace, Nm=np.array(d.Nm),
hist_params=tomographer.UniformBinsHistogramParams(0.1, 0.2, 100),
channel_walker_jump_mode=pydnormchannelspace.RandHermExp,
mhrw_params=tomographer.MHRWParams(0.001, 1000, 4096, 32768),
progress_fn=prg.progress_fn,
progress_interval_ms=2000,
ctrl_converged_params={'enabled':False},
)
prg.displayFinalInfo(r_eiH['final_report_runs'])
save_to_cache('r_eiH', r_eiH)
print_report(r_eiH)
a_eiH = do_analysis(r_eiH, 'ch., eiH')
# ## Channel-space method, "elementary-rotation" variant
r_elr = load_from_cache('r_elr')
if r_elr is None:
# no stored result, perform computation
with tomographer.jpyutil.RandWalkProgressBar() as prg:
r_elr = QPtomographer.channelspace.run(
dimX=4, dimY=4, Emn=Emn_for_channelspace, Nm=np.array(d.Nm),
hist_params=tomographer.UniformBinsHistogramParams(0.1, 0.2, 100),
channel_walker_jump_mode=QPtomographer.channelspace.ElemRotations,
mhrw_params=tomographer.MHRWParams(0.005, 500, 4096, 32768),
progress_fn=prg.progress_fn,
progress_interval_ms=2000,
)
prg.displayFinalInfo(r_elr['final_report_runs'])
save_to_cache('r_elr', r_elr)
print_report(r_elr)
a_elr = do_analysis(r_elr, 'ch., elr.')
# ## Channel-space method, "elementary-rotation" variant, entanglement fidelity
r_Fe_elr = load_from_cache('r_Fe_elr')
if r_Fe_elr is None:
# no stored result, perform computation
with tomographer.jpyutil.RandWalkProgressBar() as prg:
r_Fe_elr = QPtomographer.channelspace.run(
dimX=4, dimY=4, Emn=Emn_for_channelspace, Nm=np.array(d.Nm),
fig_of_merit='entanglement-fidelity',
hist_params=tomographer.UniformBinsHistogramParams(0.8, 0.9, 100),
channel_walker_jump_mode=QPtomographer.channelspace.ElemRotations,
mhrw_params=tomographer.MHRWParams(0.005, 500, 4096, 32768),
progress_fn=prg.progress_fn,
progress_interval_ms=2000,
)
prg.displayFinalInfo(r_Fe_elr['final_report_runs'])
save_to_cache('r_Fe_elr', r_Fe_elr)
print_report(r_Fe_elr)
a_Fe_elr = do_analysis(r_Fe_elr, 'ch., elr.', plots=True,
qtyname='entanglement fidelity', ftox=(1,-1))
# ## Channel-space method, "elementary-rotation" variant, worst-case entanglement fidelity
r_wFe_elr = load_from_cache('r_wFe_elr')
if r_wFe_elr is None:
# no stored result, perform computation
with tomographer.jpyutil.RandWalkProgressBar() as prg:
r_wFe_elr = QPtomographer.channelspace.run(
dimX=4, dimY=4, Emn=Emn_for_channelspace, Nm=np.array(d.Nm),
fig_of_merit='worst-entanglement-fidelity',
hist_params=tomographer.UniformBinsHistogramParams(0.75, 0.9, 150),
channel_walker_jump_mode=QPtomographer.channelspace.ElemRotations,
mhrw_params=tomographer.MHRWParams(0.005, 500, 4096, 32768),
progress_fn=prg.progress_fn,
progress_interval_ms=2000,
)
prg.displayFinalInfo(r_wFe_elr['final_report_runs'])
save_to_cache('r_wFe_elr', r_wFe_elr)
print_report(r_wFe_elr)
a_wFe_elr = do_analysis(r_wFe_elr, 'ch., elr., went.', plots=True,
qtyname='worst-case entanglement fidelity', ftox=(1,-1))
# # Grand comparison plots
def do_comparison_plot(alist, fig=None, ax=None, log_scale=False, alt_analysis_args=None, xlabel='', flist=None):
if fig is None:
fig, ax = plt.subplots()
ax.set_xlabel(xlabel, fontsize=12)
ax.set_ylabel('probability density', fontsize=12)
if log_scale:
ax.set_yscale('log')
clist = 'crbgmyk'
class _Ns: pass
thelabels = _Ns()
thelabels.d = {}
def add_label(s):
k = len(thelabels.d)
thelabels.d[k] = s
return str(k)
def last_label():
return thelabels.d.get(len(thelabels.d)-1)
qeb = dict()
for i in range(len(alist)):
a = alist[i]
print("Taking care of plots for {}".format(a['name']))
r = a['r']
h = r['final_histogram'].normalized()
c = clist[i%len(clist)]
f = h.values_center
#analysis = a['analysis']
analysis_dflt = tomographer.querrorbars.HistogramAnalysis(h)
analysis_dflt.printFitParameters()
qeb[a['name']] = analysis_dflt.printQuantumErrorBars()
analysis = None
if alt_analysis_args:
analysis = tomographer.querrorbars.HistogramAnalysis(h, **alt_analysis_args)
analysis.printFitParameters()
if log_scale:
ax.errorbar(x=f, y=h.bins, yerr=h.delta, c=c, fmt='.', label=last_label())
else:
ax.errorbar(x=f, y=h.bins, yerr=h.delta, c=c, fmt='.', label=add_label('{}, his.'.format(a['name'])))
if flist is None:
theflist = np.linspace(np.min(f), np.max(f), 100)
else:
theflist = flist
plist_dflt = np.exp(analysis_dflt.fit_fn(analysis_dflt.ftox(theflist), *analysis_dflt.fit_params))
if log_scale:
ax.plot(theflist, plist_dflt, c=c, ls=':', label=last_label())
else:
ax.plot(theflist, plist_dflt, c=c, ls=':', label=add_label('{}, fit1'.format(a['name'])))
if analysis is not None:
plist = np.exp(analysis.fit_fn(analysis.ftox(theflist), *analysis.fit_params))
if log_scale:
ax.plot(theflist, plist, c=c, label=last_label())
else:
ax.plot(theflist, plist, c=c, label=add_label('{}, fit2'.format(a['name'])))
handles, labels = ax.get_legend_handles_labels()
iorder = sorted(range(len(handles)), key=lambda i: int(labels[i]))
ax.legend([handles[i] for i in iorder], [thelabels.d[int(labels[i])] for i in iorder],fontsize=11)
return dict(qeb=qeb, fig=fig, ax=ax)
# ### Diamond norm analysis
# +
def fit_fn_q_lnxp_dnorm(x, a, xq, m, p, c):
return -a*np.square(x-xq) - m*np.power(-np.log(x)/-np.log(0.16), p) + c
xyz = do_comparison_plot([a_naive, a_naiveopt, a_eiH, a_elr],
alt_analysis_args=dict(fit_fn=fit_fn_q_lnxp_dnorm, maxfev=10000,
bounds=((0,-np.inf,0,0,-np.inf),np.inf),
p0=(12, -13, 0.057, 8.8, 2100),),
xlabel='Diamond norm distance to the identity')
fig = xyz['fig']
fig.tight_layout()
fig.savefig('twoqubits.pdf', format='pdf')
plt.show()
# -
# ### Entanglement fidelity & worst-case entanglement fidelity analysis
# +
def fit_fn_q_lnxp_Fe(x, a, xq, m, p, c):
return -a*np.square(x-xq) - m*np.power(-np.log(x)/-np.log(0.2), p) + c
xyz = do_comparison_plot([a_Fe_naiveopt, a_Fe_elr, a_wFe_naiveopt, a_wFe_elr],
alt_analysis_args=dict(fit_fn=fit_fn_q_lnxp_Fe, maxfev=1000000, ftox=(1,-1),
bounds=((0,-np.inf,0,0,-np.inf),np.inf),
),
xlabel='Worst-case/average entanglement fidelity')
fig, ax = xyz['fig'], xyz['ax']
ax.set_xlim([0.77, 0.91])
ax.set_ylim([5e-4, 2e2])
fig.savefig('comparison_plot_Fe_wFe.pdf', format='pdf')
plt.show()
# -
# ## behavior of $\mu(f)$ for larger values of $f$.
#
# Note: We need to evaluate everything in log space or we get overflows.
# +
# normalized histogram
hn = r_Fe_elr['final_histogram'].normalized()
analysis_dflt_elr = tomographer.querrorbars.HistogramAnalysis(hn)
analysis_dflt_elr.printFitParameters()
analysis_dflt_elr.printQuantumErrorBars()
fig = plt.figure()
ax = fig.add_subplot(111)
ax.set_xlabel('average entanglement fidelity')
ax.set_ylabel('probability density, 10^x')
f = hn.values_center
# careful between log10() and ln()=log()
idx = np.nonzero(hn.bins)
ax.errorbar(x=f[idx], y=np.log10(hn.bins[idx]),
yerr=np.divide(hn.delta[idx], hn.bins[idx])/np.log(10), c='b', fmt='.')
theflist = np.linspace(.5, 1, 256)
y1 = analysis_dflt_elr.fit_fn(analysis_dflt_elr.ftox(theflist),
*analysis_dflt_elr.fit_params)/np.log(10.0)
ax.plot(theflist, y1, c='r', ls=':')
plt.show()
# -
| examples/2qubits-noisy-identity/run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# TNE 1 Linear Regression for Ozone rate prediction
# ===
# +
from pylab import *
import pandas as pd
import seaborn as sns
import numpy as np
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score, mean_absolute_error
from sklearn.linear_model import Ridge
from sklearn.preprocessing import StandardScaler
from copy import deepcopy
# %matplotlib inline
# -
# # Exercice 1
# ## Reading data
data = pd.read_csv('ozone_data.txt', sep='\t')
# ## Data Analysis
#
# First, we'll try to visualize pairwise relationships in a our dataset by using PairPlot function from seaborn.
#
sns.pairplot(data)
# From this visualization, we can see clearly that the pair (temperature, ozone) is the most correlated followed by the pair (wind, ozone).
#
# Let's plot the covariance matrix :
corr = data.corr()
plt.figure(figsize=(9,7))
plt.title('Correlation matrix', size=15)
sns.heatmap(corr, annot=True, cmap='RdBu')
# The covariance matrix confirmed our observations:
# * (temperature and ozone) is the most correlated pair : 0.7
# * (wind and ozone) is the most inversely correlated pair : -0.61
#
# ## Linear Regression
#
# For the linear regression, we decided to use scikitlearn package as we achived the implementation of Linear Regression algorithm before.
# We will perform regression on standarized data (for iterpretation ease).
# +
# Standarize data
N = len(data)
scaler = StandardScaler()
scaler.fit(data[:int(0.75*N)])
# Save a copy of data
data_N = deepcopy(data)
data_N[['ozone', 'radiation', 'temperature', 'wind']] = scaler.transform(data)
# Creating train and test sets
train = data_N[['radiation', 'temperature', 'wind']][:int(0.75*N)] # 75% de la data
train_t = data_N['ozone'][:int(0.75*N)]
test = data_N[['radiation', 'temperature', 'wind']][int(0.75*N):] # 25% de la data
test_t = data_N['ozone'][int(0.75*N):]
# +
# Create linear regression object
regr = linear_model.LinearRegression()
# Train the model using the training sets
regr.fit(train, train_t)
# Make predictions using the testing set
pred_t = regr.predict(test)
# The coefficients
print('Coefficients: \n', regr.coef_)
# The mean squared error
print("Mean squared error: %.5f"
% mean_squared_error(test_t, pred_t))
# -
# **As we standarized the data, the MSE is about 27% of the variance. Our prediction is within a tolerable range around the mean, but still not accurate.**
# ## Feature engineering
# We try to create new features from existing ones in order to have better quality entries.
# +
rad, temp, win = data['radiation'], data['temperature'], data['wind']
# Create new fetures from existing ones (by brainstorming and testing)
rad_temp = pd.DataFrame(data={'rad*temp': rad*temp})
temp_win = pd.DataFrame(data={'temp*win': temp*win})
temp2 = pd.DataFrame(data={'temp^2': temp**3})
win_rad = pd.DataFrame(data={'win*rad': (win*rad)})
win_rad3 = pd.DataFrame(data={'(win*rad)^3': (win*rad)**3})
# Input vectors
I = ['radiation','temperature','wind'] # original input vector
I1 = ['radiation', 'temperature', 'wind', 'rad*temp', 'temp*win', 'temp^2', 'win*rad', '(win*rad)^3']
I2 = ['radiation', 'temperature', 'wind', 'rad*temp', 'temp*win', 'win*rad']
I3 = ['radiation', 'temperature', 'rad*temp', 'temp^2']
I4 = ['temperature','rad*temp','temp^2', '(win*rad)^3']
# +
# New crafted data set
data2 = pd.concat([data, rad_temp, temp_win, temp2, win_rad, win_rad3], axis=1)
# Standarize
scaler = StandardScaler()
scaler.fit(data2[:int(0.75*N)])
data2[['ozone'] + I1] = scaler.transform(data2)
# -
# For each input, we'll predict a linear regression model and compare the corresponding mean square error.
# +
attrib = [I, I1, I2, I3, I4]
err = []
for i in attrib :
# Split data set
train2 = data2[i][: int(0.75*N)]
train2_t = data2['ozone'][:int(0.75*N)]
test2 = data2[i][int(0.75*N):]
test2_t = data2['ozone'][int(0.75*N):]
# Create linear regression object
regr2 = linear_model.LinearRegression()
# Train the model using the training sets
regr2.fit(train2, train2_t)
# Make predictions using the testing set
pred2_t = regr2.predict(test2)
# MSE
err+= [mean_squared_error(test2_t, pred2_t)]
plt.figure(figsize=(8,5))
plt.title('Mean square error as a function of entry sets', size=15)
plt.bar(range(5), err)
# -
# - All crafted features yield a better result than the original one (first column). Beginner chance ?
# - The temperature multipliers and powers improved the results.
# - The MSE has been improved by 50% (between I and I3).
# ## Ridge Regression
# Using ridge regression, we'll try to improve the linear models found earlier.
# +
Lambda = np.linspace(0,99,1000)
err3 = []
for l in Lambda :
# Create linear regression object
regr3 = linear_model.Ridge(l)
# Train the model using the training sets
regr3.fit(train, train_t)
# Make predictions using the testing set
pred3_t = regr3.predict(test)
err3 += [mean_squared_error(test_t, pred3_t)]
plt.figure(figsize=(8,5))
plt.title('Mean square error as a function of $\lambda$', size=15)
plt.plot(Lambda, err3)
# -
Lambda_best = argmin(err3)/10
print(' The best lambda parameter in this case is lambda = ',Lambda_best,'\n The corresponding error is : ', min(err3))
# The MSE was slightly improved, but its performance is still far below the one with modified features.
# Let's try to combine both methods.
# +
# After a first iteration we found that the best lambda is between 0 and 1 (except for original input)
Lambda2 = np.linspace(0,1,1000)
err4 = [min(err3)]
for i in attrib[1:] :
train4 = data2[i][: int(0.75*N)]
train4_t = data2['ozone'][:int(0.75*N)]
test4 = data2[i][int(0.75*N):]
test4_t = data2['ozone'][int(0.75*N):]
errors = []
for l in Lambda2 :
# Create linear regression object
regr4 = linear_model.Ridge(l)
# Train the model using the training sets
regr4.fit(train4, train4_t)
# Make predictions using the testing set
pred4_t = regr4.predict(test4)
errors += [mean_squared_error(test4_t, pred4_t)]
Lambda_best = argmin(errors)/1000
print('\n The best lambda parameter for features : ',i,' is lambda = ',Lambda_best,
'\n The corresponding error is : ', min(errors))
err4 +=[min(errors)]
# -
plt.figure(figsize=(8,5))
plt.title('Mean square error as a function of entry sets before and after ridge regression', size=15)
plt.bar(range(5), err)
plt.bar(range(5), err4)
plt.legend(['before','after'])
# In the third and fourth case, the best lambda is 0, wich means that the best result is obtained without using Ridge Regression.
# +
# Correlation Matrix of the 3rd case's dataset
corr2 = data2[['ozone']+ I3].corr()
plt.figure(figsize=(9,7))
plt.title('Correlation matrix', size=15)
sns.heatmap(corr2, annot=True, cmap='Blues')
# -
# The correlation matrix shows that the features $temperature$, $radiation \times temperature$, and $temperature^2$ are well correlated with targets, which may be the factor that made this combination better than the four other ones.
| TNE1/TNE1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from chesslab.agent_torch import agent
from chesslab.agent_random import agent as agent_r
from chesslab.agent_stockfish import agent as s_agent
from chesslab.tests import versus
from chesslab.agent_mcts import agent_MCTS
import chess
import chess.pgn
import torch.nn as nn
class Model_1(nn.Module):
def __init__(self):
super().__init__()
self.func_1=nn.ELU()
self.func_2=nn.ELU()
self.func_3=nn.ELU()
self.func_4=nn.ELU()
self.cnn_1 = nn.Conv2d(3, 32, kernel_size=7,padding=3)
self.cnn_2 = nn.Conv2d(32, 64, kernel_size=5,padding=2)
self.cnn_3 = nn.Conv2d(64, 128, kernel_size=3,padding=1)
self.linear_1 = nn.Linear(8*8*128,256 )
self.linear_2 = nn.Linear(256, 2)
def forward(self, x ):
out = self.cnn_1(x)
out = self.func_1(out)
out = self.cnn_2(out)
out = self.func_2(out)
out = self.cnn_3(out)
out = self.func_3(out)
out = out.reshape([x.size(0), -1])
out = self.linear_1(out)
out = self.func_4(out)
out = self.linear_2(out)
return out
model = Model_1()
deepbot = agent(model,'../tmp/test_elo.0.5.pt')
randombot = agent_r()
white_wins,black_wins,draws = versus(agent_white=randombot,agent_black=randombot)
print("White wins: {}\nBlack wins: {}\nDraws:{}".format(white_wins,black_wins,draws))
white_wins,black_wins,draws = versus(agent_white=deepbot,agent_black=randombot)
print("White wins: {}\nBlack wins: {}\nDraws:{}".format(white_wins,black_wins,draws))
board=chess.Board()
deepMCTS = agent_MCTS(temperature=2,bot=deepbot,game_state=board,verbose=2,max_iter=1000)
white_wins,black_wins,draws = versus(agent_white=deepMCTS,agent_black=randombot,n_counts=10)
print("White wins: {}\nBlack wins: {}\nDraws:{}".format(white_wins,black_wins,draws))
path = 'E:/database/stockfish_14.exe'
stockfish = s_agent(path,depth=2)
white_wins,black_wins,draws = versus(agent_white=deepMCTS,agent_black=stockfish,n_counts=10)
print("White wins: {}\nBlack wins: {}\nDraws:{}".format(white_wins,black_wins,draws))
deepMCTS.root.game_state
board=chess.Board()
board
# +
game = chess.pgn.Game()
game.headers["White"] = "MCTSBot"
game.headers["Black"] = "Stockfish14"
game.setup(board) # Not required for the standard
# starting position.
blancas=deepMCTS
negras=stockfish
node = game
# -
import chess.svg
from IPython.display import display
# + tags=[]
while not board.is_game_over():
move=blancas.select_move(board)
board.push(move)
node = node.add_variation(move) # Add game node
if not board.is_game_over():
move=negras.select_move(board)
board.push(move)
node = node.add_variation(move) # Add game node
#print(board)
display(chess.svg.board(board,size=350))
print("=="*8)
game.headers["Result"] = board.result()
board.result()
# -
board
print(game)
| examples/versus/VS_random-Copy2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2. KNN
# ### References:
# 1. https://github.com/ypwhs/dogs_vs_cats
# 2. https://www.kaggle.com/yangpeiwen/keras-inception-xception-0-47
# ### Import pkgs
import cv2
import numpy as np
from tqdm import tqdm
import pandas as pd
import os
import random
import matplotlib.pyplot as plt
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
# +
# from keras.preprocessing import image
# from keras.models import Model
# from keras.layers import Dense, GlobalAveragePooling2D
# from keras import backend as K
# from keras.layers import Input
# from keras.layers.core import Lambda
# from keras.applications.vgg16 import VGG16
# from keras.applications.vgg19 import VGG19
# from keras.applications.resnet50 import ResNet50
# from keras.applications.inception_v3 import InceptionV3
# from keras.applications.xception import Xception
# from keras.applications.inception_resnet_v2 import InceptionResNetV2
from keras.layers import *
from keras.models import *
from keras.applications import *
from keras.optimizers import *
from keras.regularizers import *
from keras.applications.inception_v3 import preprocess_input
# -
# ### Load data
cwd = os.getcwd()
df = pd.read_csv(os.path.join(cwd, 'input', 'labels.csv'))
print('lables amount: %d' %len(df))
df.head()
n = len(df)
breed = set(df['breed'])
n_class = len(breed)
class_to_num = dict(zip(breed, range(n_class)))
num_to_class = dict(zip(range(n_class), breed))
width = 299
X = np.zeros((n, width, width, 3), dtype=np.uint8)
y = np.zeros(n, dtype=np.uint8)
for i in tqdm(range(n)):
X[i] = cv2.resize(cv2.imread('.\\input\\train\\%s.jpg' % df['id'][i]), (width, width))
y[i] = class_to_num[df['breed'][i]]
# ### Preview images
plt.figure(figsize=(12, 6))
for i in range(8):
random_index = random.randint(0, n-1)
plt.subplot(2, 4, i+1)
plt.imshow(X[random_index][:,:,::-1])
plt.title(num_to_class[y[random_index].argmax()])
plt.show()
# ## Shulfe and split data
print(X.shape)
print(y.shape)
x_data = X.reshape(X.shape[0],-1)
print(x_data.shape)
print(y[0:10])
from sklearn.utils import shuffle
x_data, y_data = shuffle(x_data, y)
# +
from sklearn.model_selection import train_test_split
x_train, x_val, y_train, y_val = train_test_split(x_data, y_data, test_size=0.05)
print(x_train.shape)
print(y_train.shape)
print(x_val.shape)
print(y_val.shape)
# -
# ### Train model
from sklearn.neighbors import KNeighborsClassifier
clf = KNeighborsClassifier(n_neighbors=80)
clf.fit(x_train, y_train)
# ## Cross validation
from sklearn.metrics import log_loss, accuracy_score
print('Val log_loss: {}'.format(log_loss(log_loss_y_val, val_proba)))
val_proba_limit = np.clip(log_loss_y_val, 0.005, 0.995)
print('Val log_loss: {}'.format(log_loss(log_loss_y_val, val_proba_limit)))
print('Val accuracy_score: {}'.format(accuracy_score(y_val, val_preds)))
# ### Load test data
df2 = pd.read_csv('.\\input\\sample_submission.csv')
n_test = len(df2)
X_test = np.zeros((n_test, width, width, 3), dtype=np.uint8)
for i in tqdm(range(n_test)):
X_test[i] = cv2.resize(cv2.imread('.\\input\\test\\%s.jpg' % df2['id'][i]), (width, width))
# ### Export test data feature
inception_features = get_features(InceptionV3, X_test)
xception_features = get_features(Xception, X_test)
features_test = np.concatenate([inception_features, xception_features], axis=-1)
# ### Get test data prediction and output
y_pred = model.predict(features_test, batch_size=128)
for b in breed:
df2[b] = y_pred[:,class_to_num[b]]
df2.to_csv('.\\output\\pred.csv', index=None)
print('Done !')
| dog-breed-identification/3. KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="D4mfB5aNV4iv" colab_type="text"
# # Data Processing and Analysis
#
# Data Processing is the most important and most time consuming component of the overall lifecycle of any Machine Learning project.
#
# In this notebook, we will analyze a dummy dataset to understand different issues we face with real world datasets and steps to handle the same.
# + [markdown] id="_zbobieHWJtz" colab_type="text"
# ## Utilities
#
# We add in some utility functions here which we will be using across this notebook. We have also packaged it into a `utils.py` file which you can use offline. Since we will be using colab for the tutorials, we add in all the functions in the same notebook to save the hassle of file uploads and drive connects
# + id="bnAJjlj3Wb8c" colab_type="code" colab={}
import datetime
import random
from random import randrange
import numpy as np
import pandas as pd
def _random_date(start,date_count):
"""This function generates a random date based on params
Args:
start (date object): the base date
date_count (int): number of dates to be generated
Returns:
list of random dates
"""
current = start
while date_count > 0:
curr = current + datetime.timedelta(days=randrange(42))
yield curr
date_count-=1
def generate_sample_data(row_count=100):
"""This function generates a random transaction dataset
Args:
row_count (int): number of rows for the dataframe
Returns:
a pandas dataframe
"""
# sentinels
startDate = datetime.datetime(2016, 1, 1, 13)
serial_number_sentinel = 1000
user_id_sentinel = 5001
product_id_sentinel = 101
price_sentinel = 2000
# base list of attributes
data_dict = {
'Serial No':
np.arange(row_count) + serial_number_sentinel,
'Date':
np.random.permutation(
pd.to_datetime([
x.strftime("%d-%m-%Y")
for x in _random_date(startDate, row_count)
]).date),
'User ID':
np.random.permutation(
np.random.randint(0, row_count, size=int(row_count / 10)) +
user_id_sentinel).tolist() * 10,
'Product ID':
np.random.permutation(
np.random.randint(0, row_count, size=int(row_count / 10)) +
product_id_sentinel).tolist() * 10,
'Quantity Purchased':
np.random.permutation(np.random.randint(1, 42, size=row_count)),
'Price':
np.round(
np.abs(np.random.randn(row_count) + 1) * price_sentinel,
decimals=2),
'User Type':
np.random.permutation(
[chr(random.randrange(97, 97 + 3 + 1)) for i in range(row_count)])
}
# introduce missing values
for index in range(int(np.sqrt(row_count))):
data_dict['Price'][np.argmax(
data_dict['Price'] == random.choice(data_dict['Price']))] = np.nan
data_dict['User Type'][np.argmax(
data_dict['User Type'] == random.choice(
data_dict['User Type']))] = np.nan
data_dict['Date'][np.argmax(
data_dict['Date'] == random.choice(data_dict['Date']))] = np.nan
data_dict['Product ID'][np.argmax(data_dict['Product ID'] == random.
choice(data_dict['Product ID']))] = 0
data_dict['Serial No'][np.argmax(data_dict['Serial No'] == random.
choice(data_dict['Serial No']))] = -1
data_dict['User ID'][np.argmax(data_dict['User ID'] == random.choice(
data_dict['User ID']))] = -101
# create data frame
df = pd.DataFrame(data_dict)
return df
# + [markdown] id="6G3lFubNV4ix" colab_type="text"
# ## Import Dependencies
# + id="hj1nhjQhV4iy" colab_type="code" colab={}
# import required libraries
import numpy as np
import pandas as pd
from IPython.display import display
from sklearn import preprocessing
pd.options.mode.chained_assignment = None
# + [markdown] id="ZpSyKkAVV4i0" colab_type="text"
# ## Generate Dataset
#
# # + Question: Generate 1000 sample rows
# + id="EmPoVwXxV4i1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="0c3f6fb7-52b7-45a6-f0d0-34dda0678646"
## Generate a dataset with 1000 rows
df = generate_sample_data(row_count=1000)
df.shape
# + [markdown] id="z9VOI1JLV4i3" colab_type="text"
# ### Analyze generated Dataset
# + id="AbFMdnSTV4i3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="2c9ab654-75d8-4cb0-d7fb-b63104d2e9a5"
df.head()
# + [markdown] id="2TIezuylV4i5" colab_type="text"
# ### Dataframe Stats
#
# Determine the following:
#
# * The number of data points (rows). (*Hint:* check out the dataframe `.shape` attribute.)
# * The column names. (*Hint:* check out the dataframe `.columns` attribute.)
# * The data types for each column. (*Hint:* check out the dataframe `.dtypes` attribute.)
# + id="dUj0WEAeV4i6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="41749dee-2d32-4a2d-caab-b625748fb187"
print("Number of rows::",df.shape[0])
# + [markdown] id="hK32tYiDV4i7" colab_type="text"
# ### Question
# # + Get the number of columns
# + id="VDGc8jGcV4i8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="a67923f8-4adc-4341-c241-4c67ebe5d138"
print("Number of columns::",df.shape[1])
# + id="E3BB40qBV4i9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="9f7e8af3-a7c2-4dbd-ec9f-2c4e4cf7c382"
print("Column Names::",df.columns.values.tolist())
# + id="EgWKounbV4i_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 167} outputId="8ed77b9b-9b83-42a6-ff7d-46ad4f6d3ac4"
print("Column Data Types::\n",df.dtypes)
# + id="_al94gw1V4jB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="222ec914-e183-48f5-b0d0-b4b1d02a858e"
print("Columns with Missing Values::",df.columns[df.isnull().any()].tolist())
# + id="je1JCRB3V4jC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="55e508cb-b04e-4208-fefe-3bdb5bfa0f24"
print("Number of rows with Missing Values::",len(pd.isnull(df).any(1).nonzero()[0].tolist()))
# + [markdown] id="XtqgVRPlV4jE" colab_type="text"
# #### General Stats
# + id="DgsXws4HV4jE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 234} outputId="60e98759-3692-49c2-addd-c9b052c4220c"
print(df.info())
# + id="5PYXAzYUV4jG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 167} outputId="5e724f9e-1b89-40f0-d36b-928711b3afba"
print(df.describe())
# + [markdown] id="PLnxdcjfV4jH" colab_type="text"
# ## Standardize Columns
#
# ### Question
# # + Use ```columns``` attribute and ```tolist()``` method to get the list of all columns
# + id="28lgIrY8V4jI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="c6e6b50d-750a-48b6-89cb-4d2392f8604c"
# list all columns
print("Dataframe columns:\n{}".format(df.columns.tolist()))
# + [markdown] id="-EfxXb1iV4jJ" colab_type="text"
# ### Utility to Standardize Columns
#
# # + Question : We usually use lowercase-snakecased column names in python. Write a utility method to do the same. You may user methods like ```lower, replace```. Setting ```inplace``` = ```True``` avoid creating a copy of your dataframe
#
#
# *Hint:* there are multiple ways to do this, but you could use either the [string processing methods](http://pandas.pydata.org/pandas-docs/stable/text.html) or the [apply method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.apply.html).
# + id="a71tNWVMV4jK" colab_type="code" colab={}
def cleanup_column_names(df,rename_dict={},do_inplace=True):
"""This function renames columns of a pandas dataframe
It converts column names to snake case if rename_dict is not passed.
Args:
rename_dict (dict): keys represent old column names and values point to
newer ones
do_inplace (bool): flag to update existing dataframe or return a new one
Returns:
pandas dataframe if do_inplace is set to False, None otherwise
"""
if not rename_dict:
# lower case and replace <space> with <underscore>
return df.rename(columns={col: col.lower().replace(' ','_')
for col in df.columns.values.tolist()},
inplace=True)
else:
return df.rename(columns=rename_dict,inplace=do_inplace)
# + id="HdKWzjnbV4jL" colab_type="code" colab={}
cleanup_column_names(df)
# + id="CgVeuGwAV4jN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="fb303fa2-16c0-4fb0-ac13-60ab67d986b6"
# Updated column names
print("Dataframe columns:\n{}".format(df.columns.tolist()))
# + [markdown] id="ItCqzQANV4jO" colab_type="text"
# ## Basic Manipulation
# + [markdown] id="sBPpK6IgV4jP" colab_type="text"
# ### Sort basis specific attributes
#
# # + Question: Sort serial_no in ascending and price in descending order.
# + id="d9uTMQsNV4jP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="12bb74af-66c0-4936-f6f9-e3c92922f93b"
# Ascending for Serial No and Descending for Price
display(df.sort_values(['serial_no', 'price'],
ascending=[True, False]).head())
# + [markdown] id="r_Thuwc2V4jR" colab_type="text"
# ### Reorder columns
# + id="BEYFxfuDV4jR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="1abd0f28-dae7-49bc-ecad-87a3ae3cb798"
display(df[['serial_no','date','user_id','user_type',
'product_id','quantity_purchased','price']].head())
# + [markdown] id="SJ8fLBLEV4jU" colab_type="text"
# ### Select Attributes
# + id="du6oDJOMV4jV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="a415626e-04fc-4c77-ca66-1642cb5042f1"
# Using Column Index
# print 10 values from column at index 3
print(df.iloc[:,3].values[0:10])
# + id="pC9WHUkQV4jX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="b3b639a3-5fea-4fef-8fbc-434a991d6b06"
# Using Column Name
# print 10 values of quantity purchased
print(df.quantity_purchased.values[0:10])
# + id="4yGLHTiwV4jZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="2d2b8f1c-d474-4f46-eef2-4072b5f801c6"
# Using Datatype
# print 10 values of columns with data type float
print(df.select_dtypes(include=['float64']).values[:10,0])
# + [markdown] id="XvoVtl2mV4ja" colab_type="text"
# ### Select Rows
# + id="IA4lL0aCV4ja" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 137} outputId="8ebf1dd8-b17a-4685-e685-07fd0b55f917"
# Using Row Index
display(df.iloc[[10,501,20]])
# + id="6h9nAxjeV4jc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="9da40fdd-2b25-45dd-de56-40cc1abb27e4"
# Exclude specific rows
display(df.drop([0,24,51], axis=0).head())
# + [markdown] id="5dFqDng2V4je" colab_type="text"
# ### Question
# # + Show only rows which have quantity purchased greater than 25
# + id="XtugpCFEV4je" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="cce31b39-224f-428a-ec8d-823b9a82faa0"
# Conditional Filtering
# Quantity_Purchased greater than 25
display(df[df.quantity_purchased > 25].head())
# + id="d3z4CHPpV4jf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="55227af1-ed31-4381-9bb0-c15084c79451"
# Offset from Top
display(df[100:].head())
# + id="40lSP0PvV4jh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="c6f17f84-1e65-4faa-f474-df6a7158b6fc"
# Offset from Bottom
display(df[-10:].head())
# + [markdown] id="ONvj9tYjV4ji" colab_type="text"
# ### Type Casting
# + id="cjmATTb9V4jj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 150} outputId="3af7de15-25ef-44c8-aa55-5ffcf4dfb1d1"
# Existing Datatypes
df.dtypes
# + id="HTPtNFc2V4jl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 150} outputId="58ca1b1d-8725-41c0-9b24-155e0c0f9339"
# Set Datatime as dtype for date column
df['date'] = pd.to_datetime(df.date)
print(df.dtypes)
# + [markdown] id="fl1DeQFTV4jn" colab_type="text"
# ### Map/Apply Functionality
# + [markdown] id="7csCsaI5V4jn" colab_type="text"
# ### Question
# # + Write a utility method to create a new column ```user_class``` from ```user_type``` using the following mapping:
# - ```user_type``` __a__ and __b__ map to ```user_class``` __new__
# - ```user_type``` __c__ maps to ```user_class``` __existing__
# - ```user_type``` __d__ maps to ```user_class``` __loyal_existing__
# - map all other ```user_type``` values as __error__
# + id="sn3nEHucV4jo" colab_type="code" colab={}
def expand_user_type(u_type):
"""This function maps user types to user classes
Args:
u_type (str): user type value
Returns:
(str) user_class value
"""
if u_type in ['a','b']:
return 'new'
elif u_type == 'c':
return 'existing'
elif u_type == 'd':
return 'loyal_existing'
else:
return 'error'
# + id="3Lu_p7ukV4jp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="c44493da-6e1f-4698-b652-eca1865e543c"
# Map User Type to User Class
df['user_class'] = df['user_type'].map(expand_user_type)
display(df.tail())
# + [markdown] id="4Jn_JSufV4jr" colab_type="text"
# ### Question
# # + Get range for each numeric attribute, i.e. max-min
# + id="AcCePNNTV4jr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 117} outputId="5e34150b-35e1-45e0-901c-e34854ebdf85"
# Apply: Using apply to get attribute ranges
display(df.select_dtypes(include=[np.number]).apply(lambda x:
x.max()- x.min()))
# + id="HF8RrqaUV4js" colab_type="code" colab={}
# Apply-Map: Extract Week from Date
df['purchase_week'] = df[['date']].applymap(lambda dt:dt.week
if not pd.isnull(dt.week)
else 0)
# + id="QlJLljesV4ju" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="11a7b1f2-12e5-4c57-f1a9-a1241725528a"
display(df.head())
# + [markdown] id="4APDNaLFV4jv" colab_type="text"
# ## Handle Missing Values
# + id="2vjkynTtV4jv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="9e86dc5f-2a12-484f-f220-06ee014f44f3"
# Drop Rows with Missing Dates
df_dropped = df.dropna(subset=['date'])
display(df_dropped.head())
# + id="SyVo_VoHV4jw" colab_type="code" colab={}
# Filling missing price with mean price
df_dropped['price'].fillna(value=np.round(df.price.mean(),decimals=2),
inplace=True)
# + id="FudwcLMOV4jy" colab_type="code" colab={}
# Fill missing user types using values from previous row
df_dropped['user_type'].fillna(method='ffill',inplace=True)
# + [markdown] id="M5N2As03V4j0" colab_type="text"
# ## Handle Duplicates
#
# ### Question
# # + Identify duplicates only for column ```serial_no```
# + id="_zFWhH94V4j0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 213} outputId="0b0a16dd-dfe6-4ea9-cea3-6bdc68524b24"
# sample duplicates. Identify for serial_no
display(df_dropped[df_dropped.duplicated(subset=['serial_no'])].head())
print("Shape of df={}".format(df_dropped.shape))
# + id="msiSpAU0V4j1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 213} outputId="2cff3f28-69fd-440f-9550-dc40571013b6"
# Drop Duplicates
df_dropped.drop_duplicates(subset=['serial_no'],inplace=True)
display(df_dropped.head())
print("Shape of df={}".format(df_dropped.shape))
# + [markdown] id="kxGjq9aNV4j4" colab_type="text"
# ### Question
# # + Remove rows which have less than 3 attributes with non-missing data
# # + Print the shape of dataframe thus prepared
# + id="A52LVNdsV4j4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 213} outputId="cfbdbcef-da5e-4d71-90f2-8fbb718289d0"
# Remove rows which have less than 3 attributes with non-missing data
display(df.dropna(thresh=3).head())
print("Shape of df={}".format(df.dropna(thresh=3).shape))
# + [markdown] id="oTV1r9PoV4j7" colab_type="text"
# ## Handle Categoricals
# + [markdown] id="hlM07JlrV4j8" colab_type="text"
# ### One Hot Encoding
# + id="7aO4ZntMV4j8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 263} outputId="10e0af5e-5086-4c90-c80f-f0791e1cbf08"
display(pd.get_dummies(df,columns=['user_type']).head())
# + [markdown] id="aG6P8bPDV4j-" colab_type="text"
# ### Label Encoding
#
# ### Question
# # + Use a dictionary to encode user_types in sequence of numbers. Replace missing/Nan's with -1
# + id="KkzTAjsrV4j-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="6d4855d0-c5dd-42a8-cbcd-d0021163f99e"
type_map = {'a': 0, 'b': 1, 'c': 2, 'd': 3, np.NAN: -1}
df['encoded_user_type'] = df.user_type.map(type_map)
display((df.tail()))
# + [markdown] id="UxfWk0hyV4j_" colab_type="text"
# ## Handle Numerical Attributes
# + [markdown] id="nMf1AKhMV4kA" colab_type="text"
# ### Min-Max Scalar
# ### Question
# # + Control the range of numerical attribute price by using ```MinMaxScaler``` transformer
# + id="zyaklXmmV4kA" colab_type="code" colab={}
df_normalized = df.dropna().copy()
min_max_scaler = preprocessing.MinMaxScaler()
np_scaled = min_max_scaler.fit_transform(df_normalized['price'].values.reshape(-1,1))
df_normalized['price'] = np_scaled.reshape(-1,1)
# + id="QrOhPZ0uV4kD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="156bc752-5854-4357-cc0d-a1d11854d3ff"
display(df_normalized.head())
# + [markdown] id="wR4h2KiZV4kE" colab_type="text"
# ### Robust Scaler
# + id="HZmiNETLV4kF" colab_type="code" colab={}
df_normalized = df.dropna().copy()
robust_scaler = preprocessing.RobustScaler()
rs_scaled = robust_scaler.fit_transform(df_normalized['quantity_purchased'].values.reshape(-1,1))
df_normalized['quantity_purchased'] = rs_scaled.reshape(-1,1)
# + id="9e4DaFx5V4kG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="07aaf833-d162-430c-cf77-7aa632b2efb0"
display(df_normalized.head())
# + [markdown] id="RA5b4CiDV4kH" colab_type="text"
# ## Group-By
#
# ### Question
# # + Group By attribute ```user_class``` and get sum of quantity_purchased
#
# *Hint:* you may want to use Pandas [`groupby` method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.groupby.html) to group by certain attributes before calculating the statistic.
#
# Try calculating multiple statistics (mean, median, etc) in a single table (i.e. with a single groupby call). See the section of the Pandas documentation on [applying multiple functions at once](http://pandas.pydata.org/pandas-docs/stable/groupby.html#applying-multiple-functions-at-once) for a hint.
# + id="2pl-aq78V4kI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 117} outputId="fc4e93ed-1961-49a3-dece-7fc494804e08"
# Group By attributes user_class and get sum of quantity_purchased
print(df.groupby(['user_class'])['quantity_purchased'].sum())
# + id="Knu9iT1UV4kJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="0f1c6446-b522-4d05-a0c9-75c03f4fc8fd"
# Aggregate Functions. Sum, Mean and Non Zero Row Count
display(
df.groupby(['user_class'])['quantity_purchased'].agg(
[np.sum, np.mean, np.count_nonzero]))
# + id="T3FO3fFRV4kL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 226} outputId="3bf05fa3-1a82-47c5-8c3e-98269982a887"
# Aggregate Functions specific to columns
display(df.groupby(['user_class','user_type']).agg({'price':np.mean,
'quantity_purchased':np.max}))
# + id="vq5dWfxbV4kM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="177144a5-4c4b-4792-ebf4-1325a5481757"
# Multiple Aggregate Functions
display(
df.groupby(['user_class', 'user_type']).agg({
'price': {
'total_price': np.sum,
'mean_price': np.mean,
'variance_price': np.std,
'count': np.count_nonzero
},
'quantity_purchased': np.sum
}))
# + [markdown] id="l_PbgxNNV4kO" colab_type="text"
# ## Pivot Tables
# + id="winLa3toV4kO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="e1f166f6-1f37-4c0d-f519-6dbf285b2447"
display(df.pivot_table(index='date', columns='user_type',
values='price',aggfunc=np.mean))
# + [markdown] id="ZvSyymJ2V4kQ" colab_type="text"
# ## Stacking
# + id="F78Wmf6IV4kQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 217} outputId="790ce799-9acb-4d1d-a067-aa76913843c0"
print(df.stack())
| Chapter 02 - Data Processing and Analysis/data_preprocessing_and_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="w_H_3i8P8eZA" outputId="abe87c69-c879-434d-c4b8-1d42ef8e89de"
# ! pip install -q pyspark==3.1.2 spark-nlp==3.3.2 --upgrade
# + [markdown] id="YUAbznLNGXd3"
# ## 1. Start Spark Session
# + colab={"base_uri": "https://localhost:8080/"} id="ZvBTTIMD9Oiq" outputId="302f0d32-6bd8-4adc-8bf9-dd0337ba3535"
import pandas as pd
import numpy as np
import os
import json
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
import pyspark.sql.functions as F
from sparknlp.annotator import *
from sparknlp.base import *
import sparknlp
from sparknlp.pretrained import PretrainedPipeline
spark = sparknlp.start()
print("Spark NLP version", sparknlp.version())
print("Apache Spark version:", spark.version)
# + [markdown] id="yfJQDKVi7Vfb"
# ## 2. Preparing Input
# + id="j3lBSipoGmA3"
text_list = [
"""Bu sıralar moralim bozuk.""",
"""Sınavımı geçtiğimi öğrenince derin bir nefes aldım.""",
"""Hizmet kalite çok güzel teşekkürler""",
"""Meydana gelen kazada 1 kisi hayatini kaybetti.""",
"""Ocak ayinda deprem bekleniyor""",
"""Gun batimi izlemeyi cok severim.""",
]
files = [f"{i}.txt" for i in (range(1, len(text_list)+1))]
df = spark.createDataFrame(pd.DataFrame({'text': text_list, 'file' : files}))
# + [markdown] id="vxVkYd9e7Zlj"
# ## 3. Define Pipeline
# + colab={"base_uri": "https://localhost:8080/"} id="Z1XUmtpzNv_G" outputId="de2043f8-6687-4820-a972-b7f0616dea13"
document = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
embeddings = UniversalSentenceEncoder.pretrained("tfhub_use_multi", "xx") \
.setInputCols("document") \
.setOutputCol("sentence_embeddings")
sentimentClassifier = ClassifierDLModel.pretrained("classifierdl_use_sentiment", "tr") \
.setInputCols(["document", "sentence_embeddings"]) \
.setOutputCol("class")
sentiment_pipeline = Pipeline(stages=[document, embeddings, sentimentClassifier])
tr_sentiment_pipeline = sentiment_pipeline.fit(spark.createDataFrame([['']]).toDF("text"))
# + [markdown] id="6aVbeJqi7f5T"
# ## 4. Predictions
# + id="G4MCgugsGr4g"
res = tr_sentiment_pipeline.transform(df).toPandas()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="WwxhWHIXHQbL" outputId="dfefb984-e69c-4766-be59-3cfd381e1d02"
pd.set_option('display.max_colwidth', None)
res[['text', 'class']].head()
# + id="4H5bbIOqMDq9" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="d349c331-aa8a-42dd-ba97-501eff4d8726"
res['prediction'], res['score'] = '-', 0
res = res[['text', 'class', 'prediction', 'score']].explode('class')
res['prediction'] = res['class'].apply(lambda row: row[3])
res['score'] = res['class'].apply(lambda row: round(float(row[4][row[3].strip()])*100, 3))
res.head()
| tutorials/streamlit_notebooks/CLASSIFICATION_TR_SENTIMENT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
a = float(input('Enter a degree in Celsius: '))
b = (9 / 5) * a + 32
print('%0.1f Celsius is %0.1f Fahrenheit' %(a,b))
# +
radius,length = eval(input('Enter the radius of a cylinder: '))
area = radius * radius * 3.14159
volume = area * length
print('The area is %0.4f' %(area))
print('The volume is %0.1f'%(volume))
# -
y = float(input('Enter a value for feet: '))
m = y * 0.305
print('%0.1f feet is %0.4f meters' %(y,m))
M = float(input('Enter the amount of water in kilograms: '))
i = float(input('Enter the initial temperature: '))
f = float(input('Enter the final temperature: '))
Q = M * (f - i) *4184
print('The energy needed is %0.1f' %(Q))
c,l = eval(input('Enter balance and interest rate (e.g., 3 for 3%): '))
m = c * (l/1200)
print('The interset is %0.5f' %(m))
v0,v1, t = eval(input('Enter v0,v1, and t:'))
a = (v1-v0)/t
print('The average acceleration is %0.4f' %(a))
# +
a= float(input('Enter the monthly saving amount:'))
account= a * ((1+0.00417)**6+(1+0.00417)**5+(1+0.00417)**4+(1+0.00417)**3+(1+0.00417)**2+(1+0.00417)**1)
print('After the sixth month, the account value is %0.2f' %(account))
| Firstday.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="mq3xfj1yS_hz"
USERNAME = "USERNAME" # replace here
PASSWORD = "PASSWORD" # replace here
ELAB = "ELAB"
# replace above ELAB with : (don't remove the quotes)
# computingskill_c (for c)
# computingskill_cpp (for c++)
# computingskill_java (for java)
# computingskill_ds (for data structure)
# computingskill_ml (for mathslab)
# computingskill_py (for python)
# java1, java2 or ada
# !pip install img2pdf
import requests
import os
import img2pdf
def gen_report(username, password, elabx):
computingskill_c = {'url': 'http://care.srmuniv.ac.in/computingskill/', 'code': 'c/c.code.php', 'key': 'c'}
computingskill_cpp = {'url': 'http://care.srmuniv.ac.in/computingskill/', 'code': 'cpp/cpp.code.php', 'key': 'CPP'}
computingskill_java = {'url': 'http://care.srmuniv.ac.in/computingskill/', 'code': 'java/java.code.php', 'key': 'JAVA'}
computingskill_ds = {'url': 'http://care.srmuniv.ac.in/computingskill/', 'code': 'data-structure/data-structure.code.php', 'key': 'DATA-STRUCTURE'}
computingskill_ml = {'url': 'http://care.srmuniv.ac.in/computingskill/', 'code': 'mathslab/mathslab.code.php', 'key': 'MATHSLAB'}
computingskill_py = {'url': 'http://care.srmuniv.ac.in/computingskill/', 'code': 'python/python.code.php', 'key': 'PYTHON'}
java1 = {'url': 'http://care.srmuniv.ac.in/ktrcsejava1/', 'code': 'java/java.code.php', 'key': 'java'}
java2 = {'url': 'http://care.srmuniv.ac.in/ktrcsejava2/', 'code': 'java/java.code.php', 'key': 'java'}
ada = {'url': 'http://care.srmuniv.ac.in/ktrcseada/', 'code': 'daa/daa.code.php', 'key': 'daa'}
pdd = {'url': 'http://care.srmuniv.ac.in/ktrcsepdd/', 'code': 'c/c.code.php', 'key': 'c'}
it_ada = {'url': 'http://care1.srmuniv.ac.in/ktritada/', 'code': 'daa/daa.code.php', 'key': 'daa'}
it_java = {'url': 'http://care1.srmuniv.ac.in/ktritjava/', 'code': 'java/java.code.php', 'key': 'java'}
if(elabx == 'computingskill_c'):
elab = computingskill_c
elif(elabx == 'computingskill_cpp'):
elab = computingskill_cpp
elif(elabx == 'computingskill_java'):
elab = computingskill_java
elif(elabx == 'computingskill_ds'):
elab = computingskill_ds
elif(elabx == 'computingskill_ml'):
elab = computingskill_ml
elif(elabx == 'computingskill_py'):
elab = computingskill_py
elif(elabx == 'java1'):
elab = java1
elif(elabx == 'java2'):
elab = java2
elif(elabx == 'ada'):
elab = ada
elif(elabx == 'it_java'):
elab = it_java
elif(elabx == 'it_ada'):
elab = it_ada
elif(elabx == 'pdd'):
elab = pdd;
else:
return
login_page = elab['url'] + 'login_check.php'
home_page = elab['url'] + 'login/student/home.php'
question_page = elab['url'] + 'login/student/code/' + elab['code'] + '?id=1&value='
payload = {
'uname': username,
'pass': password
}
print('eLab Report Generator : ' + payload['uname'])
with requests.Session() as s:
# login page
s.post(login_page, data=payload)
# home page
s.get(home_page)
# question page requests & responses
s.get(elab['url'] + 'login/student/question.php')
s.post(elab['url'] + 'login/student/home.helper.php', data={'text': elab['key'].upper()})
s.get(elab['url'] + 'login/student/question.php')
s.get(elab['url'] + 'login/student/question.list.js')
s.post(elab['url'] + 'login/student/course.get.php', data={'q': 'SESSION'})
s.post(elab['url'] + 'login/student/course.get.php', data={'q': 'VALUES'})
# individual question -> code page
s.get(elab['url'] + 'login/student/code/' + elab['code'] + '?id=1&value=0')
s.get(elab['url'] + 'Code-mirror/lib/codemirror.js')
s.get(elab['url'] + 'Code-mirror/mode/clike/clike.js')
s.get(elab['url'] + 'login/student/code/' + elab['key'] + '/code.elab.js')
s.post(elab['url'] + 'login/student/code/code.get.php')
s.post(elab['url'] + 'login/student/code/flag.checker.php')
# get the code, evaluate it and download the report (if 100%)
for i in range(0, 100):
present_question = question_page + str(i)
s.get(present_question)
code = s.get(elab['url'] + 'login/student/code/code.get.php')
if(code.text != ''):
if(elab['key'] == 'daa'):
evaluate_payload_c = s.post(elab['url'] + 'login/student/code/' + elab['key'] + '/code.evaluate.elab.php', data={'code': code.text, 'input': '', 'language': 'c'})
evaluate_payload_cpp = s.post(elab['url'] + 'login/student/code/' + elab['key'] + '/code.evaluate.elab.php', data={'code': code.text, 'input': '', 'language': 'cpp'})
evaluate_payload_java = s.post(elab['url'] + 'login/student/code/' + elab['key'] + '/code.evaluate.elab.php', data={'code': code.text, 'input': '', 'language': 'java'})
evaluate_payload_python = s.post(elab['url'] + 'login/student/code/' + elab['key'] + '/code.evaluate.elab.php', data={'code': code.text, 'input': '', 'language': 'python'})
evaluate_payload_mathslab = s.post(elab['url'] + 'login/student/code/' + elab['key'] + '/code.evaluate.elab.php', data={'code': code.text, 'input': '', 'language': 'mathslab'})
if '100' in [evaluate_payload_c.text[-4:-1], evaluate_payload_cpp.text[-4:-1], evaluate_payload_java.text[-4:-1], evaluate_payload_python.text[-4:-1], evaluate_payload_mathslab[-4:-1]]:
complete_percent = '100'
else:
complete_percent = '0'
else:
evaluate_payload = s.post(elab['url'] + 'login/student/code/' + elab['key'] + '/code.evaluate.elab.php', data={'code': code.text, 'input': ''})
complete_percent = evaluate_payload.text[-4:-1]
if(complete_percent == '100'):
print(str(i + 1) + ' : getting report')
file = s.get(elab['url'] + 'login/student/code/getReport.php')
with open(str(i).zfill(3) + '.png', 'wb') as f:
f.write(file.content)
else:
print(str(i + 1) + ' : evaluation error : Couldn\'t get report')
else:
print(str(i + 1) + ' : No code written')
# put all the images to PDF
global filename
filename = payload['uname'] + '-' + elabx.upper() + '.pdf'
with open(filename, "ab") as f:
f.write(img2pdf.convert([i for i in sorted(os.listdir('.')) if i.endswith('.png')]))
print('PDF file named ' + filename + ' generated')
# remove the image files
for i in range(0, 100):
if(os.path.isfile(str(i).zfill(3) + '.png')):
os.remove(str(i).zfill(3) + '.png')
print('Image files cleared')
filename = 'dummy'
gen_report(USERNAME, PASSWORD, ELAB)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="PnO3RKLXc7JA"
from google.colab import files
uploaded = files.download(filename)
| elabreport.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example 06: How to use synthetic data to enable unsupervised learning
#
# -------------------------------------------
#
# ## Overview
#
#
#
# - AitiaExplorer synthetic data to enable unsupervised learning.
# - This achieved using a BayesianGaussianMixture (BGMM).
# - A BGMM can be used for clustering but it can also be used to model the data distribution that best represents the data.
# - This means that a BGMM can be used to model a data distribution and provide samples from that distribution, allowing the creation of sythentic data.
# - This synthetic data can then be combined with the real data, along with an extra label that separates the synthetic data from the real data.
# - This new dataset can then allow a classifier to be trained to recognise the real data in an unsupervised manner.
# - The code below in the method `get_synthetic_training_data` creates such a dataset.
# - These classifiers are used internally in AitiaExplorer to select the most important features in a dataset.
# ### Imports
# +
import os
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import mixture
from sklearn.linear_model import LinearRegression, SGDClassifier
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
module_path = os.path.abspath(os.path.join('../src'))
if module_path not in sys.path:
sys.path.append(module_path)
from aitia_explorer.app import App
# stop the warning clutter
import warnings
warnings.filterwarnings('ignore')
# -
# ### Define Utility Methods
#
# - `get_gmm_sample_data` creates sample data from a BGMM.
# - `get_synthetic_training_data` creates training data from synthetic and real data.
# - These methods are taken from the internals of AitiaExplorer.
def get_gmm_sample_data(incoming_df, column_list, sample_size):
"""
Unsupervised Learning in the form of BayesianGaussianMixture to create sample data.
"""
gmm = mixture.BayesianGaussianMixture(n_components=2,
covariance_type="full",
n_init=100,
random_state=42).fit(incoming_df)
clustered_data = gmm.sample(sample_size)
clustered_df = pd.DataFrame(clustered_data[0], columns=column_list)
return clustered_df
def get_synthetic_training_data(incoming_df):
"""
Creates synthetic training data by sampling from a BayesianGaussianMixture supplied distribution.
Synthetic data is then labelled differently from the original data.
"""
# number of records in df
number_records = len(incoming_df.index)
# get sample data from the unsupervised BayesianGaussianMixture
df_bgmm = get_gmm_sample_data(incoming_df, list(incoming_df), number_records)
# set the class on the samples
df_bgmm['original_data'] = 0
# add the class to a copy of incoming df, stops weird errors due to changed dataframes
working_df = incoming_df.copy(deep=True)
working_df['original_data'] = 1
# concatinate the two dataframes
df_combined = working_df.append(df_bgmm, ignore_index=True)
# shuffle the data
df_combined = df_combined.sample(frac=1)
# get the X and y
x = df_combined.drop(['original_data'], axis=1).values
y = df_combined['original_data'].values
y = y.ravel()
return x, y
# ### Set up training
#
# - Now we will set up for training the classifiers by creating an AitiaExplorer instance and using it to load the [HEPAR II](https://www.bnlearn.com/bnrepository/#hepar2) dataset.
# - This data will then be divided into training and test datatsets.
aitia = App()
df = aitia.data.hepar2_10k_data()
# get ths synthetic data
X, y = get_synthetic_training_data(df)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
# ### Train the classifiers
#
# - Now we will train some classifiers that normally need labelled data i.e. for supervised learning.
# - However, as we have created a synthetic training set, we can use these classifiers in an unsupervised manner to learn the real data.
# - The scores will then be displayed.
models = [
SGDClassifier,
RandomForestClassifier,
GradientBoostingClassifier,
XGBClassifier]
# +
model_results = dict()
for model in models:
current_model = model()
# fit the model
current_model.fit(X_train, y_train)
# score
model_results[type(current_model).__name__] = [current_model.score(X_test, y_test)]
model_df = pd.DataFrame(model_results)
model_df
# -
# ## Observations
# - Several of the classifiers have an almost perfect score on the synthetic dataset.
# - Even though the SGDClassifier does very poorly, it is still useful for feature selection.
# - LinearRegression has been omitted from this test as the score metric returns meaningless results in regression, however it is also still useful for feature selection.
| notebooks/Example06-Synthetic_Data_and_Unsupervised_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="c9r3yAxdO9px" colab_type="text"
# <center>
# <a href="https://drive.google.com/file/d/1N01IvqI0yxK1CAKi0cfwRTcgvR-_YukL/view?usp=sharing"> <img align="center" src="https://colab.research.google.com/assets/colab-badge.svg"> </a></center>
# + [markdown] id="tapQt1VGqf93" colab_type="text"
# ## Importing Required Libraries
# + id="htIT7EgIaSQ2" colab_type="code" colab={}
from io import open
import matplotlib.pyplot as plt
import numpy as np
import random
import time
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from google.colab import drive
# + id="RlLcUWGRctg2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f7f2b98f-5e82-49db-9162-8f63264e36a5"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("Notebook is running on", device)
# + [markdown] id="WXJE8ELy6kXA" colab_type="text"
# ## Mounting G-Drive
# You can skip this cell if you don't plan to save the model and training plots. For sampling, you can download the pre-trained model.
# + id="VtZOccKHc8nQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="dc525152-8873-4936-9e37-a12ad3481224"
drive.mount('/content/gdrive')
# + [markdown] id="XbD4VA-NqlcA" colab_type="text"
# ## Fetching Data
# Use `!wget -O your_file_name.extension raw_link`
#
# To write contents from *raw_link* into *your_file_name.extension*.
# + id="wTNQjJrFqksJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="cee1056c-4d98-492a-c734-7353b2ebd68c"
# !wget -O Dino_Names.txt https://github.com/IvLabs/Natural-Language-Processing/raw/master/datasets/Dino_Names.txt
# + [markdown] id="v2q0ZV23qo65" colab_type="text"
# ## Data Preprocessing
# + id="GPy0VLxbdZJP" colab_type="code" colab={}
FILE_NAME = "Dino_Names.txt"
data = open(FILE_NAME).read().strip()
data = data.lower()
data = data.replace(" ", "")
data = data.replace("\n", ".")
# + id="CWPQ0665dkna" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7029a527-8c59-4db6-e232-800cc0666e03"
print(data[:50])
# + id="mXqk6wyWeEd2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="7e4a016b-292d-4cc2-ddd6-d71d2564762f"
vocab = list(set(data))
vocab.sort()
print("Vocabulary: ", vocab)
print("Vocabulary Length: ", len(vocab))
# + id="vIKwcnPCeIex" colab_type="code" colab={}
char_idx = {c: i for i, c in enumerate(vocab)}
idx_char = {i: c for i, c in enumerate(vocab)}
# + id="Y0Sk8Yq3ibAO" colab_type="code" colab={}
data = data.replace(".", " ")
data = data.split()
random.shuffle(data)
# + [markdown] id="H_nlHFyNqwXl" colab_type="text"
# ## Helper Functions
# + id="CVXBjK-kzy_6" colab_type="code" colab={}
def Split(corpus, valid_perc = 2, test_perc = 3):
valid_split = int(len(corpus) * valid_perc / 100)
test_split = int(len(corpus) * test_perc / 100)
train_iterator = corpus[: -(valid_split + test_split)]
test_iterator = corpus[-(valid_split + test_split): -valid_split]
valid_iterator = corpus[-valid_split: ]
return train_iterator, valid_iterator, test_iterator
# + id="6TsS7ygGeRTp" colab_type="code" colab={}
def InputTensor(x, vocab_size):
ip_tensor = torch.zeros((len(x), vocab_size)) # [input_tensor] = [seq_len, vocab_size]
for i in range(len(x)):
letter = x[i]
ip_tensor[i][char_idx[letter]] = 1
return ip_tensor
# + id="IZbjlPs0eUL7" colab_type="code" colab={}
def TargetTensor(x):
ids = [char_idx[letter] for letter in x[1:]]
ids.append(char_idx['.'])
target_tensor = torch.LongTensor(ids) # [target_tensor] = [seq_len]
return target_tensor
# + id="TjYx4l7yeXAO" colab_type="code" colab={}
def Example(word, vocab_size):
input_tensor = InputTensor(word, vocab_size)
target_tensor = TargetTensor(word)
return input_tensor, target_tensor
# + [markdown] id="KQvXsQO2q-u_" colab_type="text"
# ## Model Definition
# + id="j7OeAq3LeK5r" colab_type="code" colab={}
class Scratch_RNN(nn.Module):
def __init__(self, input_size, hidden_size, vocab_size):
super(Scratch_RNN, self).__init__()
self.hidden_size = hidden_size
self.vocab_size = vocab_size
self.i2h = nn.Linear(input_size + hidden_size, hidden_size)
self.i2o = nn.Linear(input_size + hidden_size, vocab_size)
self.o2o = nn.Linear(hidden_size + vocab_size, vocab_size)
self.dropout = nn.Dropout(0.25)
# We do not use softmax as nn.CrossEntropyLoss() calculates both LogSoftmax and NLLLoss
def forward(self, input, hidden): # [input] = [1, input_size], [hidden] = [1, hidden_size]
input_combined = torch.cat((input, hidden), 1) # [input_combined] = [1, input_size + hidden_size]
hidden = self.i2h(input_combined) # [hidden] = [1, hidden_size]
output = self.i2o(input_combined) # [output] = [1, vocab_size]
output_combined = torch.cat((hidden, output), 1) # [output_combined] = [1, vocab_size + hidden_size]
output = self.o2o(output_combined) # [output] = [1, vocab_size]
output = self.dropout(output)
return output, hidden
def initHidden(self):
return torch.zeros(1, self.hidden_size) # Since [input] = [1, input_size] and we concat along dim = 1
# + [markdown] id="AHrH4g1mrCAt" colab_type="text"
# ## Training and Evaluation Functions
# + id="QQOHiMTWecv7" colab_type="code" colab={}
def Train(iterator, model, criterion, optimizer):
model.train()
epoch_loss = 0
for word in iterator:
input_tensor, target_tensor = Example(word, model.vocab_size) # [input_tensor] = [seq_len, vocab_size]
seq_len = input_tensor.shape[0]
predictions = torch.zeros(seq_len, model.vocab_size).to(device) # [predictions] = [seq_len, vocab_size]
hidden = model.initHidden()
optimizer.zero_grad()
for i in range(seq_len):
input = input_tensor[i].unsqueeze(0).to(device) # [input] = [1, vocab_size] i.e. One Hot vector
output, hidden = model.forward(input, hidden.to(device)) # [output] = [1, vocab_size]
predictions[i] = output.squeeze(0)
batch_loss = criterion(predictions, target_tensor.to(device))
epoch_loss += batch_loss.item()
batch_loss.backward()
optimizer.step()
return epoch_loss / len(iterator)
# + id="SaWjq6h3rRWJ" colab_type="code" colab={}
def Evaluate(iterator, model, criterion):
model.eval()
epoch_loss = 0
for word in iterator:
input_tensor, target_tensor = Example(word, model.vocab_size)
seq_len = input_tensor.shape[0]
predictions = torch.zeros(seq_len, model.vocab_size).to(device)
hidden = model.initHidden()
with torch.no_grad():
for i in range(seq_len):
input = input_tensor[i].unsqueeze(0).to(device)
output, hidden = model.forward(input, hidden.to(device))
predictions[i] = output.squeeze(0)
batch_loss = criterion(predictions, target_tensor.to(device))
epoch_loss += batch_loss.item()
return epoch_loss / len(iterator)
# + id="LhF4jVZH6T1p" colab_type="code" colab={}
def Epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return (elapsed_mins, elapsed_secs)
# + [markdown] id="OLqo3SqurHNZ" colab_type="text"
# ## Data Iterators, Hyperparameters and Model Initialization
# + id="MpMxn-bv1CbZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3406b8a2-31b4-4dac-c681-c4e95e86d700"
VALID_PERC = 2
TEST_PERC = 3
train_iterator, valid_iterator, test_iterator = Split(data, VALID_PERC, TEST_PERC)
print(f"Number of Training examples: {len(train_iterator)} | Number of Testing examples: {len(test_iterator)} | Number of Validation examples: {len(valid_iterator)}")
# + id="Z3aRcj6WeZiH" colab_type="code" colab={}
NUM_EPOCHS = 40
LR = 4e-4
HIDDEN_DIM = 64
VOCAB_SIZE = len(vocab)
MODEL_STORE_PATH = '/content/gdrive/My Drive/Colab/NLP/Char RNNs/Models/Dino_Names_Scratch.pth'
# + id="m6OPWShuqLOz" colab_type="code" colab={}
rnn = Scratch_RNN(VOCAB_SIZE, HIDDEN_DIM, VOCAB_SIZE).to(device)
optimizer = optim.AdamW(rnn.parameters(), LR)
criterion = nn.CrossEntropyLoss()
# + id="_Xr2iNEFfLDA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e45b18ea-592a-4dd6-c3c6-7eec8d08e614"
total_params = sum(p.numel() for p in rnn.parameters())
total_learn_params = sum(p.numel() for p in rnn.parameters() if p.requires_grad)
print(f"Total Parameters: {total_params} | Total Learnable Parameters: {total_learn_params}")
# + [markdown] id="5U7q0na6F0iu" colab_type="text"
# ## Download Pre-Trained Model
# Run this section instead of the "Training" section if you wish to download the pre-trained model instead of training one yourself.
# + id="9LVqPOYOF28l" colab_type="code" colab={}
# # !wget -O Dino_Names_Scratch.pth https://gitlab.com/rishika2110/weights/-/blob/6e88f0c7acaf5d86d803eca8abb150e0b79f95ef/char_rnns/Dino_Names_Scratch.pth
# + id="DoEc4aEYF8OZ" colab_type="code" colab={}
# MODEL_STORE_PATH = "Dino_Names_Scratch.pth"
# + [markdown] id="0HIYkwwcrM2q" colab_type="text"
# ## Training
# Run this section only if you have mounted G-Drive and want to train the model yourself.
# + id="2htPo5cfej8T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="260bfb20-9e29-4445-8550-95c3ab4f0cd0"
print(f"Learning Rate: {LR}, Hidden Dimensions: {HIDDEN_DIM}")
train_losses = []
valid_losses = []
prev_epoch = 1
min_losses = [float('inf'), float('inf')]
start_time = time.time()
for epoch in range(1, NUM_EPOCHS+1):
train_loss = Train(train_iterator, rnn, criterion, optimizer)
train_losses.append(train_loss)
valid_loss = Evaluate(valid_iterator, rnn, criterion)
valid_losses.append(valid_loss)
if valid_loss < min_losses[0]:
min_losses[0] = valid_loss
min_losses[1] = train_loss
torch.save(rnn.state_dict(), MODEL_STORE_PATH)
if epoch % int(NUM_EPOCHS / 10) == 0:
elapsed_time = Epoch_time(start_time, time.time())
print(f"Time taken for epochs {prev_epoch} to {epoch}: {elapsed_time[0]}m {elapsed_time[1]}s")
start_time = time.time()
prev_epoch = epoch + 1
print(f"Training Loss: {train_losses[epoch - 1]:.4f} | Valid Loss: {valid_losses[epoch - 1]:.4f}")
print(f"Model with Train Loss {min_losses[1]:.4f}, Validation Loss: {min_losses[0]:.4f} was saved.")
# + id="-4R0EyZu3asK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="1a6e9a77-8330-4936-c65c-c4c4c0883b20"
plt.title('RNN from scratch: Learning Curves')
plt.xlabel('Number of Epochs')
plt.ylabel('Cross Entropy Loss')
plt.plot(train_losses, label = 'Training Loss')
plt.plot(valid_losses, label='Validation Loss')
plt.legend()
plt.savefig('/content/gdrive/My Drive/Colab/NLP/Char RNNs/Plots/Dino_Names_Scratch.jpeg')
plt.show()
# + [markdown] id="niooddVPrPXD" colab_type="text"
# ## Sampling
# + id="0Rk5kbUVheoE" colab_type="code" colab={}
max_length = 20
def sample(start_letter, model):
with torch.no_grad():
start_letter = start_letter.lower()
input = InputTensor(start_letter, model.vocab_size)
hidden = model.initHidden()
output_name = str(start_letter)
for i in range(max_length):
output, hidden = model(input.to(device), hidden.to(device))
probs = F.softmax(output, dim = 1)
ind = probs.argmax(1).item()
if ind == char_idx["."]:
break
else:
letter = idx_char[ind]
output_name += letter
input = InputTensor(letter, model.vocab_size)
return output_name
# + id="yrAFAFlvhrRQ" colab_type="code" colab={}
def samples(start_letters, model):
for start_letter in start_letters:
print(f"Starting character: {start_letter} -> Dino name: {sample(start_letter, model)}")
# + id="HlGE2RgenIJm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 476} outputId="8110ca98-7fa9-4a8a-a2c3-25e745cf67a6"
rnn.load_state_dict(torch.load(MODEL_STORE_PATH))
rnn.eval()
print(f"Test Loss of Loaded Model: {Evaluate(test_iterator, rnn, criterion)}")
samples('ABCDEFGHIJKLMNOPQRSTUVWXYZ', rnn)
# + [markdown] id="pxpiwfb9Duv6" colab_type="text"
# ## References:
#
#
# * [Generating Names with a Character-Level RNN (PyTorch)](https://pytorch.org/tutorials/intermediate/char_rnn_generation_tutorial.html)
#
#
| char_rnns/notebooks/Dino_Names_Scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# -
# <img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
#
# # Multi-GPU with MovieLens: ETL and Training
#
# ## Overview
#
# NVIDIA Merlin is a open source framework to accelerate and scale end-to-end recommender system pipelines on GPU. In this notebook, we use NVTabular, Merlin’s ETL component, to scale feature engineering and pre-processing to multiple GPUs and then perform data-parallel distributed training of a neural network on multiple GPUs with PyTorch, [Horovod](https://horovod.readthedocs.io/en/stable/), and [NCCL](https://developer.nvidia.com/nccl).
#
# The pre-requisites for this notebook are to be familiar with NVTabular and its API:
# - You can read more about NVTabular, its API and specialized dataloaders in [Getting Started with Movielens notebooks](../getting-started-movielens).
# - You can read more about scaling NVTabular ETL in [Scaling Criteo notebooks](../scaling-criteo).
#
# **In this notebook, we will focus only on the new information related to multi-GPU training, so please check out the other notebooks first (if you haven’t already.)**
#
# ### Learning objectives
#
# In this notebook, we learn how to scale ETL and deep learning taining to multiple GPUs
# - Learn to use larger than GPU/host memory datasets for ETL and training
# - Use multi-GPU or multi node for ETL with NVTabular
# - Use NVTabular dataloader to accelerate PyTorch pipelines
# - Scale PyTorch training with Horovod
#
# ### Dataset
#
# In this notebook, we use the [MovieLens25M](https://grouplens.org/datasets/movielens/25m/) dataset. It is popular for recommender systems and is used in academic publications. The dataset contains 25M movie ratings for 62,000 movies given by 162,000 users. Many projects use only the user/item/rating information of MovieLens, but the original dataset provides metadata for the movies, as well.
#
# Note: We are using the MovieLens 25M dataset in this example for simplicity, although the dataset is not large enough to require multi-GPU training. However, the functionality demonstrated in this notebook can be easily extended to scale recommender pipelines for larger datasets in the same way.
#
# ### Tools
#
# - [Horovod](https://horovod.readthedocs.io/en/stable/) is a distributed deep learning framework that provides tools for multi-GPU optimization.
# - The [NVIDIA Collective Communication Library (NCCL)](https://developer.nvidia.com/nccl) provides the underlying GPU-based implementations of the [allgather](https://docs.nvidia.com/deeplearning/nccl/user-guide/docs/usage/operations.html#allgather) and [allreduce](https://docs.nvidia.com/deeplearning/nccl/user-guide/docs/usage/operations.html#allreduce) cross-GPU communication operations.
# ## Download and Convert
#
# First, we will download and convert the dataset to Parquet. This section is based on [01-Download-Convert.ipynb](../getting-started-movielens/01-Download-Convert.ipynb).
# #### Download
# +
# External dependencies
import os
import pathlib
import cudf # cuDF is an implementation of Pandas-like Dataframe on GPU
from nvtabular.utils import download_file
INPUT_DATA_DIR = os.environ.get(
"INPUT_DATA_DIR", "~/nvt-examples/multigpu-movielens/data/"
)
BASE_DIR = pathlib.Path(INPUT_DATA_DIR).expanduser()
zip_path = pathlib.Path(BASE_DIR, "ml-25m.zip")
download_file(
"http://files.grouplens.org/datasets/movielens/ml-25m.zip", zip_path, redownload=False
)
# -
# #### Convert
movies = cudf.read_csv(pathlib.Path(BASE_DIR, "ml-25m", "movies.csv"))
movies["genres"] = movies["genres"].str.split("|")
movies = movies.drop("title", axis=1)
movies.to_parquet(pathlib.Path(BASE_DIR, "ml-25m", "movies_converted.parquet"))
# #### Split into train and validation datasets
# +
ratings = cudf.read_csv(pathlib.Path(BASE_DIR, "ml-25m", "ratings.csv"))
ratings = ratings.drop("timestamp", axis=1)
# shuffle the dataset
ratings = ratings.sample(len(ratings), replace=False)
# split the train_df as training and validation data sets.
num_valid = int(len(ratings) * 0.2)
train = ratings[:-num_valid]
valid = ratings[-num_valid:]
train.to_parquet(pathlib.Path(BASE_DIR, "train.parquet"))
valid.to_parquet(pathlib.Path(BASE_DIR, "valid.parquet"))
# -
# ## ETL with NVTabular
#
# We finished downloading and converting the dataset. We will preprocess and engineer features with NVTabular on multiple GPUs. You can read more
# - about NVTabular's features and API in [getting-started-movielens/02-ETL-with-NVTabular.ipynb](../getting-started-movielens/02-ETL-with-NVTabular.ipynb).
# - scaling NVTabular ETL to multiple GPUs [scaling-criteo/02-ETL-with-NVTabular.ipynb](../scaling-criteo/02-ETL-with-NVTabular.ipynb).
# #### Deploy a Distributed-Dask Cluster
#
# This section is based on [scaling-criteo/02-ETL-with-NVTabular.ipynb](../scaling-criteo/02-ETL-with-NVTabular.ipynb) and [multi-gpu-toy-example/multi-gpu_dask.ipynb](../multi-gpu-toy-example/multi-gpu_dask.ipynb)
# +
# Standard Libraries
import shutil
# External Dependencies
import cudf
from dask_cuda import LocalCUDACluster
from dask.distributed import Client
import rmm
# NVTabular
import nvtabular as nvt
from nvtabular.io import Shuffle
from nvtabular.utils import device_mem_size
# +
# define some information about where to get our data
input_path = pathlib.Path(BASE_DIR, "converted", "movielens")
dask_workdir = pathlib.Path(BASE_DIR, "test_dask", "workdir")
output_path = pathlib.Path(BASE_DIR, "test_dask", "output")
stats_path = pathlib.Path(BASE_DIR, "test_dask", "stats")
# Make sure we have a clean worker space for Dask
if pathlib.Path.is_dir(dask_workdir):
shutil.rmtree(dask_workdir)
dask_workdir.mkdir(parents=True)
# Make sure we have a clean stats space for Dask
if pathlib.Path.is_dir(stats_path):
shutil.rmtree(stats_path)
stats_path.mkdir(parents=True)
# Make sure we have a clean output path
if pathlib.Path.is_dir(output_path):
shutil.rmtree(output_path)
output_path.mkdir(parents=True)
# Get device memory capacity
capacity = device_mem_size(kind="total")
# +
# Deploy a Single-Machine Multi-GPU Cluster
protocol = "tcp" # "tcp" or "ucx"
visible_devices = "0,1" # Delect devices to place workers
device_spill_frac = 0.5 # Spill GPU-Worker memory to host at this limit.
# Reduce if spilling fails to prevent
# device memory errors.
cluster = None # (Optional) Specify existing scheduler port
if cluster is None:
cluster = LocalCUDACluster(
protocol=protocol,
CUDA_VISIBLE_DEVICES=visible_devices,
local_directory=dask_workdir,
device_memory_limit=capacity * device_spill_frac,
)
# Create the distributed client
client = Client(cluster)
client
# +
# Initialize RMM pool on ALL workers
def _rmm_pool():
rmm.reinitialize(
pool_allocator=True,
initial_pool_size=None, # Use default size
)
client.run(_rmm_pool)
# -
# #### Defining our Preprocessing Pipeline
#
# This subsection is based on [getting-started-movielens/02-ETL-with-NVTabular.ipynb](../getting-started-movielens/02-ETL-with-NVTabular.ipynb). The only difference is that we initialize the NVTabular workflow using the LocalCUDACluster client with `nvt.Workflow(output, client=client)`.
# +
movies = cudf.read_parquet(pathlib.Path(BASE_DIR, "ml-25m", "movies_converted.parquet"))
joined = ["userId", "movieId"] >> nvt.ops.JoinExternal(movies, on=["movieId"])
cat_features = joined >> nvt.ops.Categorify()
ratings = nvt.ColumnSelector(["rating"]) >> nvt.ops.LambdaOp(lambda col: (col > 3).astype("int8"))
output = cat_features + ratings
# USE client in NVTabular workflow
workflow = nvt.Workflow(output, client=client)
# !rm -rf $BASE_DIR/train
# !rm -rf $BASE_DIR/valid
train_iter = nvt.Dataset([str(pathlib.Path(BASE_DIR, "train.parquet"))], part_size="100MB")
valid_iter = nvt.Dataset([str(pathlib.Path(BASE_DIR, "valid.parquet"))], part_size="100MB")
workflow.fit(train_iter)
workflow.save(pathlib.Path(BASE_DIR, "workflow"))
shuffle = Shuffle.PER_WORKER # Shuffle algorithm
out_files_per_proc = 4 # Number of output files per worker
workflow.transform(train_iter).to_parquet(
output_path=pathlib.Path(BASE_DIR, "train"),
shuffle=shuffle,
out_files_per_proc=out_files_per_proc,
)
workflow.transform(valid_iter).to_parquet(
output_path=pathlib.Path(BASE_DIR, "valid"),
shuffle=shuffle,
out_files_per_proc=out_files_per_proc,
)
client.shutdown()
cluster.close()
# -
# ## Training with PyTorch on multiGPUs
#
# In this section, we will train a PyTorch model with multi-GPU support. In the NVTabular v0.5 release, we added multi-GPU support for NVTabular dataloaders. We will modify the [getting-started-movielens/03-Training-with-PyTorch.ipynb](../getting-started-movielens/03-Training-with-PyTorch.ipynb) to use multiple GPUs. Please review that notebook, if you have questions about the general functionality of the NVTabular dataloaders or the neural network architecture.
#
# #### NVTabular dataloader for PyTorch
#
# We’ve identified that the dataloader is one bottleneck in deep learning recommender systems when training pipelines with PyTorch. The normal PyTorch dataloaders cannot prepare the next training batches fast enough and therefore, the GPU is not fully utilized.
#
# We developed a highly customized tabular dataloader for accelerating existing pipelines in PyTorch. In our experiments, we see a speed-up by 9x of the same training workflow with NVTabular dataloader. NVTabular dataloader’s features are:
# - removing bottleneck of item-by-item dataloading
# - enabling larger than memory dataset by streaming from disk
# - reading data directly into GPU memory and remove CPU-GPU communication
# - preparing batch asynchronously in GPU to avoid CPU-GPU communication
# - supporting commonly used .parquet format
# - easy integration into existing PyTorch pipelines by using similar API
# - **supporting multi-GPU training with Horovod**
#
# You can find more information on the dataloaders in our [blogpost](https://medium.com/nvidia-merlin/training-deep-learning-based-recommender-systems-9x-faster-with-PyTorch-cc5a2572ea49).
# #### Using Horovod with PyTorch and NVTabular
#
# The training script below is based on [getting-started-movielens/03-Training-with-PyTorch.ipynb](../getting-started-movielens/03-Training-with-PyTorch.ipynb), with a few important changes:
#
# - We provide several additional parameters to the `TorchAsyncItr` class, including the total number of workers `hvd.size()`, the current worker's id number `hvd.rank()`, and a function for generating random seeds `seed_fn()`.
#
# ```python
# train_dataset = TorchAsyncItr(
# ...
# global_size=hvd.size(),
# global_rank=hvd.rank(),
# seed_fn=seed_fn,
# )
# ```
# - The seed function uses Horovod to collectively generate a random seed that's shared by all workers so that they can each shuffle the dataset in a consistent way and select partitions to work on without overlap. The seed function is called by the dataloader during the shuffling process at the beginning of each epoch:
#
# ```python
# def seed_fn():
# max_rand = torch.iinfo(torch.int).max // hvd.size()
#
# # Generate a seed fragment
# seed_fragment = cupy.random.randint(0, max_rand)
#
# # Aggregate seed fragments from all Horovod workers
# seed_tensor = torch.tensor(seed_fragment)
# reduced_seed = hvd.allreduce(seed_tensor, name="shuffle_seed", op=hvd.mpi_ops.Sum)
#
# return reduced_seed % max_rand
# ```
#
# - We wrap the PyTorch optimizer with Horovod's `DistributedOptimizer` class and scale the learning rate by the number of workers:
#
# ```python
# optimizer = torch.optim.Adam(model.parameters(), lr=0.01 * lr_scaler)
# optimizer = hvd.DistributedOptimizer(optimizer, named_parameters=model.named_parameters())
# ```
#
# - We broadcast the model and optimizer parameters to all workers with Horovod:
#
# ```python
# hvd.broadcast_parameters(model.state_dict(), root_rank=0)
# hvd.broadcast_optimizer_state(optimizer, root_rank=0)
# ```
#
# The rest of the script is the same as the MovieLens example in [getting-started-movielens/03-Training-with-PyTorch.ipynb](../getting-started-movielens/03-Training-with-PyTorch.ipynb). In order to run it with Horovod, we first need to write it to a file.
# +
# %%writefile './torch_trainer.py'
import argparse
import glob
import os
from time import time
import cupy
import torch
import nvtabular as nvt
from nvtabular.framework_utils.torch.models import Model
from nvtabular.framework_utils.torch.utils import process_epoch
from nvtabular.loader.torch import DLDataLoader, TorchAsyncItr
# Horovod must be the last import to avoid conflicts
import horovod.torch as hvd # noqa: E402, isort:skip
parser = argparse.ArgumentParser(description="Train a multi-gpu model with Torch and Horovod")
parser.add_argument("--dir_in", default=None, help="Input directory")
parser.add_argument("--batch_size", default=None, help="Batch size")
parser.add_argument("--cats", default=None, help="Categorical columns")
parser.add_argument("--cats_mh", default=None, help="Categorical multihot columns")
parser.add_argument("--conts", default=None, help="Continuous columns")
parser.add_argument("--labels", default=None, help="Label columns")
parser.add_argument("--epochs", default=1, help="Training epochs")
args = parser.parse_args()
hvd.init()
gpu_to_use = hvd.local_rank()
if torch.cuda.is_available():
torch.cuda.set_device(gpu_to_use)
BASE_DIR = os.path.expanduser(args.dir_in or "./data/")
BATCH_SIZE = int(args.batch_size or 16384) # Batch Size
CATEGORICAL_COLUMNS = args.cats or ["movieId", "userId"] # Single-hot
CATEGORICAL_MH_COLUMNS = args.cats_mh or ["genres"] # Multi-hot
NUMERIC_COLUMNS = args.conts or []
# Output from ETL-with-NVTabular
TRAIN_PATHS = sorted(glob.glob(os.path.join(BASE_DIR, "train", "*.parquet")))
proc = nvt.Workflow.load(os.path.join(BASE_DIR, "workflow/"))
EMBEDDING_TABLE_SHAPES = nvt.ops.get_embedding_sizes(proc)
# TensorItrDataset returns a single batch of x_cat, x_cont, y.
def collate_fn(x):
return x
# Seed with system randomness (or a static seed)
cupy.random.seed(None)
def seed_fn():
"""
Generate consistent dataloader shuffle seeds across workers
Reseeds each worker's dataloader each epoch to get fresh a shuffle
that's consistent across workers.
"""
max_rand = torch.iinfo(torch.int).max // hvd.size()
# Generate a seed fragment
seed_fragment = cupy.random.randint(0, max_rand)
# Aggregate seed fragments from all Horovod workers
seed_tensor = torch.tensor(seed_fragment)
reduced_seed = hvd.allreduce(seed_tensor, name="shuffle_seed", op=hvd.mpi_ops.Sum)
return reduced_seed % max_rand
train_dataset = TorchAsyncItr(
nvt.Dataset(TRAIN_PATHS),
batch_size=BATCH_SIZE,
cats=CATEGORICAL_COLUMNS + CATEGORICAL_MH_COLUMNS,
conts=NUMERIC_COLUMNS,
labels=["rating"],
device=gpu_to_use,
global_size=hvd.size(),
global_rank=hvd.rank(),
shuffle=True,
seed_fn=seed_fn,
)
train_loader = DLDataLoader(
train_dataset, batch_size=None, collate_fn=collate_fn, pin_memory=False, num_workers=0
)
EMBEDDING_TABLE_SHAPES_TUPLE = (
{
CATEGORICAL_COLUMNS[0]: EMBEDDING_TABLE_SHAPES[CATEGORICAL_COLUMNS[0]],
CATEGORICAL_COLUMNS[1]: EMBEDDING_TABLE_SHAPES[CATEGORICAL_COLUMNS[1]],
},
{CATEGORICAL_MH_COLUMNS[0]: EMBEDDING_TABLE_SHAPES[CATEGORICAL_MH_COLUMNS[0]]},
)
model = Model(
embedding_table_shapes=EMBEDDING_TABLE_SHAPES_TUPLE,
num_continuous=0,
emb_dropout=0.0,
layer_hidden_dims=[128, 128, 128],
layer_dropout_rates=[0.0, 0.0, 0.0],
).cuda()
lr_scaler = hvd.size()
optimizer = torch.optim.Adam(model.parameters(), lr=0.01 * lr_scaler)
hvd.broadcast_parameters(model.state_dict(), root_rank=0)
hvd.broadcast_optimizer_state(optimizer, root_rank=0)
optimizer = hvd.DistributedOptimizer(optimizer, named_parameters=model.named_parameters())
for epoch in range(args.epochs):
start = time()
print(f"Training epoch {epoch}")
train_loss, y_pred, y = process_epoch(train_loader,
model,
train=True,
optimizer=optimizer)
hvd.join(gpu_to_use)
hvd.broadcast_parameters(model.state_dict(), root_rank=0)
print(f"Epoch {epoch:02d}. Train loss: {train_loss:.4f}.")
hvd.join(gpu_to_use)
t_final = time() - start
total_rows = train_dataset.num_rows_processed
print(
f"run_time: {t_final} - rows: {total_rows} - "
f"epochs: {epoch} - dl_thru: {total_rows / t_final}"
)
hvd.join(gpu_to_use)
if hvd.local_rank() == 0:
print("Training complete")
# -
# !horovodrun -np 2 python torch_trainer.py --dir_in $BASE_DIR --batch_size 16384
| examples/multi-gpu-movielens/01-03-MultiGPU-Download-Convert-ETL-with-NVTabular-Training-with-PyTorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Blob Detection
#
# https://www.learnopencv.com/blob-detection-using-opencv-python-c/
#
# cv2.drawKeypoints(input image, keypoints, blank_output_array, color, flags)
#
# flags:
# - cv2.DRAW_MATCHES_FLAGS_DEFAULT
# - cv2.DRAW_MATCHES_FLAGS_DRAW_RICH_KEYPOINTS
# - cv2.DRAW_MATCHES_FLAGS_DRAW_OVER_OUTIMG
# - cv2.DRAW_MATCHES_FLAGS_NOT_DRAW_SINGLE_POINTS
# +
# Standard imports
import cv2
import numpy as np;
# Read image
image = cv2.imread("../images/Sunflowers.jpg",0)
# cv2.imshow("Blobs", image)
# cv2.waitKey(0)
# Set up the detector with default parameters
detector = cv2.SimpleBlobDetector_create()
# Detect blobs
keypoints = detector.detect(image)
# Draw detected blobs
blank = np.zeros((1,1))
blobs = cv2.drawKeypoints(image, keypoints, blank, (0,255,255),cv2.DRAW_MATCHES_FLAGS_DEFAULT)
# Show keypoints
cv2.imshow("Blobs", blobs)
cv2.waitKey(0)
cv2.destroyAllWindows()
# -
#
| 06 - countours extended/02 - Blob Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="wKKzqdLzSSvb"
# In the 20×20 grid below, four numbers along a diagonal line have been marked in red.
#
# `08 02 22 97 38 15 00 40 00 75 04 05 07 78 52 12 50 77 91 08`
#
# `49 49 99 40 17 81 18 57 60 87 17 40 98 43 69 48 04 56 62 00`
#
# `81 49 31 73 55 79 14 29 93 71 40 67 53 88 30 03 49 13 36 65`
#
# `52 70 95 23 04 60 11 42 69 24 68 56 01 32 56 71 37 02 36 91`
#
# `22 31 16 71 51 67 63 89 41 92 36 54 22 40 40 28 66 33 13 80`
#
# `24 47 32 60 99 03 45 02 44 75 33 53 78 36 84 20 35 17 12 50`
#
# `32 98 81 28 64 23 67 10 (26) 38 40 67 59 54 70 66 18 38 64 70`
#
# `67 26 20 68 02 62 12 20 95 (63) 94 39 63 08 40 91 66 49 94 21`
#
# `24 55 58 05 66 73 99 26 97 17 (78) 78 96 83 14 88 34 89 63 72`
#
# `21 36 23 09 75 00 76 44 20 45 35 (14) 00 61 33 97 34 31 33 95`
#
# `78 17 53 28 22 75 31 67 15 94 03 80 04 62 16 14 09 53 56 92`
#
# `16 39 05 42 96 35 31 47 55 58 88 24 00 17 54 24 36 29 85 57`
#
# `86 56 00 48 35 71 89 07 05 44 44 37 44 60 21 58 51 54 17 58`
#
# `19 80 81 68 05 94 47 69 28 73 92 13 86 52 17 77 04 89 55 40`
#
# `04 52 08 83 97 35 99 16 07 97 57 32 16 26 26 79 33 27 98 66`
#
# `88 36 68 87 57 62 20 72 03 46 33 67 46 55 12 32 63 93 53 69`
#
# `04 42 16 73 38 25 39 11 24 94 72 18 08 46 29 32 40 62 76 36`
#
# `20 69 36 41 72 30 23 88 34 62 99 69 82 67 59 85 74 04 36 16`
#
# `20 73 35 29 78 31 90 01 74 31 49 71 48 86 81 16 23 57 05 54`
#
# `01 70 54 71 83 51 54 69 16 92 33 48 61 43 52 01 89 19 67 48`
#
# The product of these numbers is $26 × 63 × 78 × 14 = 1788696$.
#
# What is the greatest product of four adjacent numbers in the same direction (up, down, left, right, or diagonally) in the 20×20 grid?
# + colab={"base_uri": "https://localhost:8080/"} id="OMJ29OvjSTQA" outputId="92c71192-b4e1-4f84-9c34-57896ce7cd2c"
grid = [
"08 02 22 97 38 15 00 40 00 75 04 05 07 78 52 12 50 77 91 08",
"49 49 99 40 17 81 18 57 60 87 17 40 98 43 69 48 04 56 62 00",
"81 49 31 73 55 79 14 29 93 71 40 67 53 88 30 03 49 13 36 65",
"52 70 95 23 04 60 11 42 69 24 68 56 01 32 56 71 37 02 36 91",
"22 31 16 71 51 67 63 89 41 92 36 54 22 40 40 28 66 33 13 80",
"24 47 32 60 99 03 45 02 44 75 33 53 78 36 84 20 35 17 12 50",
"32 98 81 28 64 23 67 10 26 38 40 67 59 54 70 66 18 38 64 70",
"67 26 20 68 02 62 12 20 95 63 94 39 63 08 40 91 66 49 94 21",
"24 55 58 05 66 73 99 26 97 17 78 78 96 83 14 88 34 89 63 72",
"21 36 23 09 75 00 76 44 20 45 35 14 00 61 33 97 34 31 33 95",
"78 17 53 28 22 75 31 67 15 94 03 80 04 62 16 14 09 53 56 92",
"16 39 05 42 96 35 31 47 55 58 88 24 00 17 54 24 36 29 85 57",
"86 56 00 48 35 71 89 07 05 44 44 37 44 60 21 58 51 54 17 58",
"19 80 81 68 05 94 47 69 28 73 92 13 86 52 17 77 04 89 55 40",
"04 52 08 83 97 35 99 16 07 97 57 32 16 26 26 79 33 27 98 66",
"88 36 68 87 57 62 20 72 03 46 33 67 46 55 12 32 63 93 53 69",
"04 42 16 73 38 25 39 11 24 94 72 18 08 46 29 32 40 62 76 36",
"20 69 36 41 72 30 23 88 34 62 99 69 82 67 59 85 74 04 36 16",
"20 73 35 29 78 31 90 01 74 31 49 71 48 86 81 16 23 57 05 54",
"01 70 54 71 83 51 54 69 16 92 33 48 61 43 52 01 89 19 67 48"]
def cell(x, y):
return int(grid[y-1][(x-1)*3 : (x-1)*3+2])
def combos():
for y in range(1, 21):
for x in range(1, 18):
yield [cell(x, y), cell(x+1, y), cell(x+2, y), cell(x+3, y)]
for x in range(1, 21):
for y in range(1, 18):
yield [cell(x, y), cell(x, y+1), cell(x, y+2), cell(x, y+3)]
for x in range(1, 18):
for y in range(1, 18):
yield [cell(x, y), cell(x+1, y+1), cell(x+2, y+2), cell(x+3, y+3)]
for x in range(4, 21):
for y in range(1, 18):
yield [cell(x, y), cell(x-1, y+1), cell(x-2, y+2), cell(x-3, y+3)]
def product(values):
result = 1
for value in values:
result *= value
return result
def all_products():
for nums in combos():
yield product(nums)
print(max(allProducts()))
# + id="jJnuePrXTc1Z"
| 11_Largest_Product_in_a_Grid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # !wget https://github.com/bamps53/convnext-tf/releases/download/v0.1/convnext_small_1k_224_ema.h5
# -
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
import tensorflow as tf
import numpy as np
from convnext import ConvNeXt, model_configs
x = tf.keras.Input(shape=(None, None, 3))
x_placeholder = tf.placeholder(tf.float32, shape = (None, None, None, 3))
tf.keras.layers.Input(tensor=x_placeholder)
def preprocess_input(x):
x = x.astype(np.float32)
x /= 255.
mean = [0.485, 0.456, 0.406]
std = [0.229, 0.224, 0.225]
x[..., 0] -= mean[0]
x[..., 1] -= mean[1]
x[..., 2] -= mean[2]
x[..., 0] /= std[0]
x[..., 1] /= std[1]
x[..., 2] /= std[2]
return x
from skimage.io import imread
from skimage.transform import resize
image = imread('panda.jpg')
image.shape
image_resized = resize(image, (224, 224))
image_resized = preprocess_input(image_resized)
image_resized = np.expand_dims(image_resized, 0)
image_resized.shape
num_classes=1000
include_top=True
cfg = model_configs['convnext_small']
net = ConvNeXt(num_classes, cfg['depths'],
cfg['dims'], include_top)
out = net(x_placeholder)
out
backbone = tf.keras.Model(x_placeholder, out)
backbone
net.load_weights('convnext_small_1k_224_ema.h5')
session = tf.keras.backend.get_session()
from tensorflow.keras.applications.imagenet_utils import decode_predictions
y = session.run(backbone.output, feed_dict = {x_placeholder: image_resized})
decode_predictions(y)
saver = tf.train.Saver()
saver.save(session, "convnext_small_1k_224_ema/model.ckpt")
| pretrained/ign-convnext/convert-small-convnext-224-to-tf1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# +
import numpy as np
from numpy.random import randn
import pandas as pd
from scipy import stats
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
data1 = randn(100)
data2 = randn(100)
sns.boxplot([data1,data2])
sns.boxplot([data1,data2], whis=np.inf)
sns.boxplot(data1, whis=np.inf)
sns.boxplot(data2, whis=np.inf)
sns.boxplot(data = [data1, data2], orient='v', whis=np.inf)
data1 = stats.norm(0,5).rvs(100)
data2 = np.concatenate([stats.gamma(5).rvs(50) - 1,
stats.gamma(5).rvs(50) * - 1])
sns.boxplot(data = [data1, data2], orient='v', whis=np.inf)
sns.violinplot(data=[data1, data2])
sns.violinplot(data=[data1, data2], bw=.01)
sns.violinplot(data=[data1, data2], inner='stick')
| l8/Lecture 51 - Box and Violin Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import matplotlib as mpl
label_size=16
mpl.rcParams['xtick.labelsize']=label_size
mpl.rcParams['ytick.labelsize']=label_size
t=np.arange(-50,50,5)
#t=np.linspace(-50,50,100)
y=np.sin(t)
plt.plot(t,y,'o-')
t=np.linspace(0,50,100)
plt.figure(figsize=(7,4))
plt.plot(t, 0.5*np.cos(0.5*t)+0.1*np.sin(0.5*t),'-',lw=1,ms=4,label='$x(t)$')
plt.plot(t, 0.5*0.1*np.cos(0.5*t)-0.5*0.5*np.sin(0.5*t),'ro-',lw=1,ms=4,label=r'$\dot(x(t))$')
plt.xlabel('$t$',fontsize=20)
plt.legend()
plt.show()
# !pip install seaborn
import seaborn as sns
sns.set(style='ticks',palette='set2')
frecuencias=np.array([0.1,0.2,0.5,0.6])
plt.figure(figsize=(7,4))
for f in frecuencias:
plt.plot(t,0.5*np.cos(f*t)+0.1*np.sin(f*t),'o-',label='$\omega_0=%a$'%f)
plt.xlabel('$t$',fontsize=16)
plt.ylabel('$x(t)$',fontsize=16)
plt.title('Oscilaciones',fontsize=16)
plt.legend(loc='center left',bbox_to_anchor=(1.05,0,5),prop={'size':14})
plt.show()
from ipywidgets import *
def masa_resorte(t=0):
fig=plt.figure()
ax=fig.add_subplot(1,1,1)
ax.plot(.5*np.cos(.5*t)+.1*np.sin(.5*t),[0],'ko',ms=10)
ax.set_xlim(left=-0.6,right=.6)
ax.axvline(x=0,color='r')
ax.axhline(y=0,color='grey',lw=1)
fig.canvas.draw()
interact(masa_resorte,t=(0,50,.01));
| Sistema masa resorte.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: u4-s3-dnn
# kernelspec:
# display_name: U4-S2-NNF-DS10
# language: python
# name: u4-s2-nnf-ds10
# ---
# + [markdown] id="vDJYgMhRRI51" colab_type="text"
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# # Major Neural Network Architectures Challenge
# ## *Data Science Unit 4 Sprint 3 Challenge*
#
# In this sprint challenge, you'll explore some of the cutting edge of Data Science. This week we studied several famous neural network architectures:
# recurrent neural networks (RNNs), long short-term memory (LSTMs), convolutional neural networks (CNNs), and Autoencoders. In this sprint challenge, you will revisit these models. Remember, we are testing your knowledge of these architectures not your ability to fit a model with high accuracy.
#
# __*Caution:*__ these approaches can be pretty heavy computationally. All problems were designed so that you should be able to achieve results within at most 5-10 minutes of runtime locally, on AWS SageMaker, on Colab or on a comparable environment. If something is running longer, double check your approach!
#
# ## Challenge Objectives
# *You should be able to:*
# * <a href="#p1">Part 1</a>: Train a LSTM classification model
# * <a href="#p2">Part 2</a>: Utilize a pre-trained CNN for object detection
# * <a href="#p3">Part 3</a>: Describe a use case for an autoencoder
# * <a href="#p4">Part 4</a>: Describe yourself as a Data Science and elucidate your vision of AI
# + [markdown] colab_type="text" id="-5UwGRnJOmD4"
# <a id="p1"></a>
# ## Part 1 - LSTMSs
#
# Use a LSTM to fit a multi-class classification model on Reuters news articles to distinguish topics of articles. The data is already encoded properly for use in a LSTM model.
#
# Your Tasks:
# - Use Keras to fit a predictive model, classifying news articles into topics.
# - Report your overall score and accuracy
#
# For reference, the [Keras IMDB sentiment classification example](https://github.com/keras-team/keras/blob/master/examples/imdb_lstm.py) will be useful, as well as the LSTM code we used in class.
#
# __*Note:*__ Focus on getting a running model, not on maxing accuracy with extreme data size or epoch numbers. Only revisit and push accuracy if you get everything else done!
# + colab_type="code" id="DS-9ksWjoJit" outputId="1f86f880-20af-4529-c036-741db9998948" colab={"base_uri": "https://localhost:8080/", "height": 51}
from tensorflow.keras.datasets import reuters
(X_train, y_train), (X_test, y_test) = reuters.load_data(num_words=None,
skip_top=0,
maxlen=None,
test_split=0.2,
seed=723812,
start_char=1,
oov_char=2,
index_from=3)
# + colab_type="code" id="fLKqFh8DovaN" outputId="548ae1ec-474d-441d-eab0-2c504aa3c181" colab={"base_uri": "https://localhost:8080/", "height": 102}
# Demo of encoding
word_index = reuters.get_word_index(path="reuters_word_index.json")
print(f"Iran is encoded as {word_index['iran']} in the data")
print(f"London is encoded as {word_index['london']} in the data")
print("Words are encoded as numbers in our dataset.")
# + id="kohfTWBfb3pQ" colab_type="code" colab={}
word_index = {k:(v+3) for k,v in word_index.items()}
word_index["<PAD>"] = 0
word_index["<START>"] = 1
word_index["<UNKNOWN>"] = 2
word_index["<UNUSED>"] = 3
# Perform reverse word lookup and make it callable
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
# + id="r2uDKWT3b-ds" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="cbfbd9c3-f6f8-4d04-e8cb-b04d6aa89bcd"
# Do not change this line. You need the +1 for some reason.
max_features = len(word_index.values()) + 1
import numpy as np
batch_size = 32
class_names = ['']
all_articles = np.concatenate((X_train, X_test), axis=0)
# Review lengths across test and training whole datasets
print("Maximum article length: {}".format(len(max((all_articles), key=len))))
print("Minimum article length: {}".format(len(min((all_articles), key=len))))
result = [len(x) for x in all_articles]
print("Mean article length: {}".format(np.mean(result)))
print("")
print("Machine Readable Article")
print(" Article Text: " + str(X_train[10]))
print(" Article Class: " + str(y_train[10]))
# Print a review and it's class in human readable format. Replace the number
# to select a different review.
print("")
print("Human Readable Article")
print(" Article Text: " + decode_review(X_train[10]))
print(" Article Class: " + str(y_train[10]))
# + id="DDCSLyAbcCY5" colab_type="code" colab={}
# Pad the articles to make them equal length
from tensorflow.keras.preprocessing import sequence
import pandas as pd
import tensorflow as tf
maxlen=225
X_train = sequence.pad_sequences(X_train, maxlen=maxlen)
X_test = sequence.pad_sequences(X_test, maxlen=maxlen)
# + id="uE7fr9dwcHaj" colab_type="code" colab={}
# One hotting our y values
y_train = tf.one_hot(y_train, depth=46, axis=-1)
y_test = tf.one_hot(y_test, depth=46, axis=-1)
# + id="QlQMCX8YcHzW" colab_type="code" colab={}
# create our modle and compile
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Embedding, Dropout, LSTM
lstm = Sequential()
lstm.add(Embedding(max_features +1, 128))
lstm.add(LSTM(32))
lstm.add(Dropout(0.25))
lstm.add(Dense(46, activation='softmax'))
lstm.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# + id="3AKNup5ycM_6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="ff5e2b95-e408-4a9d-e317-962a428b15e9"
# A useful tool is to look at the summary
lstm.summary()
# + id="kD8_2XNLcSj1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="04ecd3ce-bf1e-4ec3-c3ce-8d223d3ad0cb"
# 5 Epochs should show some improvement, but it takes a while!
lstm_history = lstm.fit(X_train,
y_train,
validation_data=(X_test, y_test),
batch_size=batch_size,
epochs=5,
verbose=1)
# + id="KC6D3KhmcUcO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="18ac4a17-e9a0-47c0-fbd2-757f42ba3e98"
# Let's plot it!
import matplotlib.pyplot as plt
plt.plot(lstm_history.history['loss'])
plt.plot(lstm_history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show();
# + [markdown] nteract={"transient": {"deleting": false}} id="kGkJ8BNhRI6A" colab_type="text"
# ## Sequence Data Question
# #### *Describe the `pad_sequences` method used on the training dataset. What does it do? Why do you need it?*
# #####Pad_sequences is a method from Keras Preprocessing that is used to ensure all sequences in a dataset will be the same length. This is done to keep it consistent for proper calculations.
#
#
# ## RNNs versus LSTMs
# #### *What are the primary motivations behind using Long-ShortTerm Memory Cell unit over traditional Recurrent Neural Networks?*
# ##### RNNs have "short-term memory". LSTMs add memorgy gates in order to use earlier weights.
#
#
# ## RNN / LSTM Use Cases
# #### *Name and Describe 3 Use Cases of LSTMs or RNNs and why they are suited to that use case*
# ##### Three cases for LSTMs are Robot control, grammar learning, and time series prediction. These are all cases that do well with LSTMs since there won't be an issue with shirt term memory loss. More data can be input for better predictions.
# + [markdown] colab_type="text" id="yz0LCZd_O4IG"
# <a id="p2"></a>
# ## Part 2- CNNs
#
# ### Find the Frog
#
# Time to play "find the frog!" Use Keras and [ResNet50v2](https://www.tensorflow.org/api_docs/python/tf/keras/applications/resnet_v2) (pre-trained) to detect which of the images with the `frog_images` subdirectory has a frog in it. Note: You will need to upload the images to Colab.
#
# <img align="left" src="https://d3i6fh83elv35t.cloudfront.net/newshour/app/uploads/2017/03/GettyImages-654745934-1024x687.jpg" width=400>
# + id="GauKNxC3fLw2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="9e22a4f5-cd8d-417d-9ef7-861654373ca8"
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + id="y_TdTeX6RZhn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="f0867af0-1de1-46de-892a-0f384f14ed14"
# I am having some serious issues getting any images to load
'''from skimage.io import imread_collection
from skimage.transform import resize
import numpy as np
images = imread_collection('./frog_images/*.jpg')'''
# + [markdown] id="QniSfajgRI6B" colab_type="text"
# The skimage function below will help you read in all the frog images into memory at once. You should use the preprocessing functions that come with ResnetV2 to help resize the images prior to inference.
# + id="0TehPjXFc6r1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 258} outputId="0c0bbb98-13cc-444e-801c-5f6c52f7aa76"
# !pip install google_images_download
# + id="HKbCa7_emgQU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="eb927b71-4642-4ec7-9caf-28b55c7be177"
# Hopefully this works
from google_images_download import google_images_download
response = google_images_download.googleimagesdownload()
arguments = {"keywords": "frog", "limit": 5, "print_urls": True}
absolute_image_paths = response.download(arguments)
# + id="72IgUOC0mgXG" colab_type="code" colab={}
# + [markdown] colab_type="text" id="si5YfNqS50QU"
# Your goal is to validly run ResNet50v2 on the input images - don't worry about tuning or improving the model. Print out the predictions in any way you see fit.
#
# *Hint* - ResNet 50v2 doesn't just return "frog". The three labels it has for frogs are: `bullfrog, tree frog, tailed frog`
#
# *Stretch goals:*
# - Check for other things such as fish.
# - Print out the image with its predicted label
# - Wrap everything nicely in well documented fucntions
# + colab_type="code" id="FaT07ddW3nHz" colab={}
import glob
import numpy as np
from tensorflow.keras.applications.resnet_v2 import ResNet50V2, decode_predictions, preprocess_input
from tensorflow.keras.preprocessing import image
from tensorflow.keras.applications.resnet50 import preprocess_input, decode_predictions
def process_img_path(img_path):
return image.load_img(img_path, target_size=(224, 224))
def img_contains_frog(img):
""" Scans image for Frogs
Should return a boolean (True/False) if a frog is in the image.
Inputs:
---------
img: Precrossed image ready for prediction. The `process_img_path` function should already be applied to the image.
Returns:
---------
frogs (boolean): TRUE or FALSE - There are frogs in the image.
"""
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
model = ResNet50V2(weights='imagenet')
features = model.predict(x)
results = decode_predictions(features)[0]
print(results)
if 'frog' in results[0][1] and results[0][2] > 0.25:
return True
else:
return False
# + id="3mmJ896Fm-bR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 547} outputId="4a225c19-88ca-4b98-8cf3-4faf98d2a941"
# Not one says frog here :/
imagelist = glob.glob('/content/drive/My Drive/frog_images2/*.jpg')
for x in imagelist:
print(img_contains_frog(process_img_path(x)))
# + [markdown] colab_type="text" id="XEuhvSu7O5Rf"
# <a id="p3"></a>
# ## Part 3 - Autoencoders
#
# Describe a use case for an autoencoder given that an autoencoder tries to predict its own input.
#
# __*Your Answer:*__
#
# #####Autoencoders can be used for denoising images, to make the images smoother, or to enhance a certain part of the image beyond what has previously been capable.
#
# + [markdown] colab_type="text" id="626zYgjkO7Vq"
# <a id="p4"></a>
# ## Part 4 - More...
# + [markdown] colab_type="text" id="__lDWfcUO8oo"
# Answer the following questions, with a target audience of a fellow Data Scientist:
#
# - What do you consider your strongest area, as a Data Scientist?
# - What area of Data Science would you most like to learn more about, and why?
# - Where do you think Data Science will be in 5 years?
# - What are the threats posed by AI to our society?
# - How do you think we can counteract those threats?
# - Do you think achieving General Artifical Intelligence is ever possible?
#
# A few sentences per answer is fine - only elaborate if time allows.
#
# #####- I consider Linear Algebra and Applied Modeling my strongest suit.
#
# #####- I would love to dive into Neural Networks some more. We touch on base items, and it's all just overview. But I know it's hard to fit everything into a schedule.
#
# ##### - Data Science will be strong and steady in 5 years. There will be high demand for data scientists!
#
# ##### - Well, any human interaction can soil AI. We are imperfect, and machines are 1s and 0s. We can be seen as a 0 and irridicated.
#
# ##### - We can establish an AI that will help seek out "unlawful" or "unethical" AI systems. Someone would need to keep that in check as well though.
#
# ##### - I do! We just need the processing power for it :)
# + [markdown] colab_type="text" id="_Hoqe3mM_Mtc"
# ## Congratulations!
#
# Thank you for your hard work, and congratulations! You've learned a lot, and you should proudly call yourself a Data Scientist.
#
# + id="WMM0bJnzRI6M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 320} outputId="5a10b7cb-3ad9-4a87-e59b-f5de4546fe64"
from IPython.display import HTML
HTML("""<iframe src="https://giphy.com/embed/26xivLqkv86uJzqWk" width="480" height="270" frameBorder="0" class="giphy-embed" allowFullScreen></iframe><p><a href="https://giphy.com/gifs/mumm-champagne-saber-26xivLqkv86uJzqWk">via GIPHY</a></p>""")
| Sprint Challenge/LS_DS_Unit_4_Sprint_Challenge_3_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#importing libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
#Reading files and storing them in variables
ny=pd.read_csv("yellow_tripdata_2018-01.csv") #database of yellow cabs in the month of January
borough=pd.read_csv("taxi _zone_lookup.csv") #databases with the classification of zones in boroughs
# +
#ny
# -
ny=ny[(ny["PULocationID"]<=263) &(ny["DOLocationID"]<=263)]
#remove taxis outside the boroughs. Seen on taxis lookup zones db
ny
ny.columns #columns of the database
ny=ny[ny["fare_amount"]>2.5]
#removing all amounts lower or equal to 2.5(standard fare)
ny
#ny2
#del ny2
#del ny3
#del ny4
# +
ny=ny[ny["trip_distance"]>=0]#no negative distances
# -
ny=ny[(ny["trip_distance"]>=0) & ((ny["RatecodeID"]==5 )| (ny["PULocationID"]!=ny["DOLocationID"]))]
#ny4[(ny4["trip_distance"]==0) &(ny4["RatecodeID"]==1)]
ny
ny=ny.join(borough.set_index("LocationID"),on="PULocationID")
ny
ny=ny.rename(index = str, columns = {"Borough":"PUBorough"})
ny=ny.drop(["PULocationID","Zone","service_zone","VendorID","store_and_fwd_flag","fare_amount","extra","mta_tax","tip_amount","tolls_amount","improvement_surcharge","total_amount"],axis=1)
#ny6=ny5.join(borough.set_index("LocationID"),on="DOLocationID")
ny=ny.join(borough.set_index("LocationID"),on="DOLocationID")
ny=ny.rename(index = str, columns = {"Borough":"DOBorough"})
ny=ny.drop(["DOLocationID","Zone","service_zone"],axis=1)
ny
ny['tpep_pickup_datetime'] = pd.to_datetime(ny['tpep_pickup_datetime'])
ny['tpep_dropoff_datetime'] = pd.to_datetime(ny['tpep_dropoff_datetime'])
ny["duration"] = ny['tpep_dropoff_datetime'] - ny['tpep_pickup_datetime']
#ny.loc[:, 'duration'] = dif
ny
| NewYork.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sandipniyogi/CenterTrack/blob/main/startfile12.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="3ccOcU5tvgoZ" outputId="376459e0-5711-4c1a-d288-7421b42f68cd"
# !pip install cython; pip install -U 'git+https://github.com/cocodataset/cocoapi.git#subdirectory=PythonAPI'
# !git clone --recursive https://github.com/sandipniyogi/CenterTrack.git
# + colab={"base_uri": "https://localhost:8080/"} id="rldwg_Jvxk3g" outputId="319cafb6-5876-4076-fcf4-c29ea9ac2b41"
# !cp -r /content/drive/MyDrive/ColabNotebooks/ .
# !cd CenterTrack; pip install -r requirements.txt
# !cd CenterTrack/src/lib/model/networks/;git clone https://github.com/lbin/DCNv2.git;
# ! cd CenterTrack/src/lib/model/networks/DCNv2;chmod +x make.sh;./make.sh
# + colab={"base_uri": "https://localhost:8080/"} id="voQKsstoyW46" outputId="bc0bd684-0b19-439b-a467-ac4124551eb6"
# !pip install gdown
#https://drive.google.com/file/d/1tJCEJmdtYIh8VuN8CClGNws3YO7QGd40/view?usp=sharing
#https://drive.google.com/file/d/1sf1bWJ1LutwQ_wp176nd2Y3HII9WeFf0/view?usp=sharing
# !gdown --id 1H0YvFYCOIZ06EzAkC2NxECNQGXxK27hH
# + colab={"base_uri": "https://localhost:8080/"} id="Q9E8hpjh-dK-" outputId="47ec55b3-f14d-407b-f5d7-c7e08a7c4d5e"
# !sudo apt-get install libx264-dev
# + colab={"base_uri": "https://localhost:8080/"} id="DBKdv0iD-gM4" outputId="5945a4fd-6cb5-448b-89e3-737d7386ae2f"
# #!gdown --id 1d82DiCiyp435dyGawaMgr3sbiU8S8rEU
# !gdown --id 1noEHcNaliP2BtvDSuMkkeWNJ6VqRb-Nr
# #!gdown --id 1JXWpV4ivzEDyjwLBuJcFU6A3jwWHLodb
# + colab={"base_uri": "https://localhost:8080/"} id="XL-NhEth_AeX" outputId="5452b0d7-0fdf-4640-8aec-61678172f2af"
# %matplotlib notebook
# %cd /content/CenterTrack/src/
# !mkdir ../results
# !python demo.py tracking --load_model /content/coco_pose_tracking.pth --demo /content/drive/MyDrive/ColabNotebooks/test4.mp4 --save_results
#python demo.py tracking --load_model ../mot17_half.pth --num_class 1 --demo ../videos/nuscenes_mini.mp4
| startfile12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import cv2
img = cv2.imread('../DATA/sammy_face.jpg')
plt.imshow(img)
edges = cv2.Canny(img,threshold1 = 127,threshold2 = 127)
plt.imshow(edges)
edges = cv2.Canny(img,threshold1 = 255,threshold2 = 0)
plt.imshow(edges)
# # Closing Thresholds
med_val = np.median(img)
med_val
lower = int(max(0,0.7 * med_val))
upper = int(max(255,1.3 * med_val))
edges = cv2.Canny(img,lower,upper)
plt.imshow(edges)
blurred_img = cv2.blur(img,(5,5))
plt.imshow(blurred_img)
plt.imshow(img)
edges = cv2.Canny(blurred_img,lower,upper)
plt.imshow(edges)
lower
upper
edges = cv2.Canny(blurred_img,lower+50,upper-50)
plt.imshow(edges)
| 04.Object Detection/02.edge detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.pipeline import make_pipeline, Pipeline
from sklearn.model_selection import cross_val_score
from sklearn.preprocessing import OneHotEncoder
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
# Standard imports
import numpy as np
import pandas as pd
from collections import Counter, OrderedDict
import re
import string
import inspect
import os
import datetime
from sklearn.metrics import f1_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_score, recall_score, precision_recall_curve
from lightgbm import LGBMClassifier
from sklearn.metrics import confusion_matrix
# -
import json
import pickle
import joblib
import pandas as pd
from flask import Flask, jsonify, request
from peewee import (
SqliteDatabase, PostgresqlDatabase, Model, IntegerField,
FloatField, TextField, IntegrityError
)
from playhouse.shortcuts import model_to_dict
from transformers import transf as tf
list(df.head(1).values)
df['Part of a policing operation'] =df['Part of a policing operation'].astype('bool')
# Group feaatures
categorical_features = ['Type','Gender','Age range','Officer-defined ethnicity','Object of search','station']
numeric_features = ['Time (sin)','Time (cos)']
# +
numeric_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='median'))])
categorical_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='constant', fill_value='missing')),
('onehot', OneHotEncoder(handle_unknown='ignore'))])
standard_preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)])
# -
df.columns
target_pipe = Pipeline(
steps=[
('Filter Metropolitain', tf.Filter_ColumnValue(col='station', value='metropolitan')),
('FixNA_OutcomeLinkedSearch', tf.FixNA_OutcomeLinkedSearch()),
('OutcomePositive', tf.Mask_OutcomePositive()),
('SearchSuccess', tf.Mask_SearchSuccess()),
])
initial_pipe = Pipeline(
steps=[
('Filter Metropolitain', tf.Filter_ColumnValue(col='station', value='metropolitan')),
('FixNA_Coordinates', tf.FixNA_Coordinates()),
('DateTransformer', tf.DateTransformer())
])
pipeline = Pipeline([
('Initial Preprocessing Pipeline',initial_pipe),
('preprocessor', standard_preprocessor),
('model', LGBMClassifier(n_jobs=-1, random_state=42))
]
)
columns = ['observation_id', 'Type', 'Date', 'Part of a policing operation',
'Latitude', 'Longitude', 'Gender', 'Age range',
'Officer-defined ethnicity', 'Legislation',
'Object of search', 'station']
outcome_columns = ['Outcome', 'Outcome linked to object of search']
df_target = target_pipe.fit_transform(df)
target = df_target['search_success']
df_target.columns
X_train = df[columns]
y_train = target
pipeline.fit(X_train, y_train)
df[columns].dtypes
# +
##Serializing the columns in the correct order
with open( os.path.join("columns.json"), 'w') as fh:
json.dump(columns, fh)
##Serializing the fitted pipeline
joblib.dump(pipeline,os.path.join( "pipeline.pickle"))
##Serializinf dtypes of columns
with open(os.path.join('dtypes.pickle'), 'wb') as fh:
pickle.dump(df[columns].dtypes, fh)
| 3_create_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import astropy.units as u
import numpy as np
from matplotlib import pyplot as plt
from astroduet.bbmag import bb_abmag_fluence, bb_abmag
import astroduet.config as config
from astroduet.background import background_pixel_rate
from astroduet.utils import duet_abmag_to_fluence
# %load_ext autoreload
# %autoreload 2
from astropy.visualization import quantity_support
import matplotlib
font = {'family' : 'normal',
'weight' : 'bold',
'size' : 22}
matplotlib.rc('font', **font)
# Account for the fact that you're co-adding the two frames here:
duet = config.Telescope(config='minimum_mass_requirement')
# +
# From Marianne:
# Using the median surface brightnesses from Bai:
# Elliptical, D1: 0.56 ph/s/pixel
# Elliptical, D2: 0.64 ph/s/pixel
# Spiral, D1: 2.09 ph/s/pixel
# Spiral, D2: 1.68 ph/s/pixel
pixel_area = duet.pixel * duet.pixel
mean_elliptical_duet1 = 24.94535864867936 * u.ABmag
mean_elliptical_duet2 = 24.730938046220277 * u.ABmag
mean_spiral_duet1 = 23.684754905932877 * u.ABmag
mean_spiral_duet2 = 23.827616937816682 * u.ABmag
rate1 = pixel_area.value * duet.fluence_to_rate(duet_abmag_to_fluence(mean_elliptical_duet1, 1))
#rate1 = pixel_area.value * duet.fluence_to_rate(duet_abmag_to_fluence(mean_spiral_duet1, 1))
print(rate1)
rate2 = pixel_area.value * duet.fluence_to_rate(duet_abmag_to_fluence(mean_elliptical_duet2, 2))
#rate2 = pixel_area.value * duet.fluence_to_rate(duet_abmag_to_fluence(mean_spiral_duet2, 2))
print(rate2)
# +
# Band1
# 5-sigma limiting magnitude in 1 and 5 stacked frames.
# Account for the fact that you're co-adding the two frames here:
duet = config.Telescope(config='minimum_mass_threshold')
bandone = duet.bandpass1
bandtwo = duet.bandpass2
surf_scale = 1.0
exposure = 300*u.s
print()
siglimit=5
dmag = 0.01
print()
[bgd_band1, bgd_band2] = background_pixel_rate(duet, med_zodi=True)
tot_bgd_rate = bgd_band1 + rate1 * surf_scale
for nframes in [8]:
snr = 100
swiftmag = 20.5
while snr > siglimit:
swiftmag += dmag
band1_fluence, band2_fluence = bb_abmag_fluence(duet =duet, swiftmag=swiftmag*u.ABmag, bbtemp=12e3*u.K)
band1_rate = duet.fluence_to_rate(band1_fluence)
band2_rate = duet.fluence_to_rate(band2_fluence)
src_rate = band1_rate
snr = duet.calc_snr(exposure, src_rate, tot_bgd_rate, nint=nframes)
bbmag1, bbmag2 = bb_abmag(swiftmag=swiftmag*u.ABmag, bbtemp=12e3*u.K, bandone = bandone, bandtwo=bandtwo)
print('Band1 {} {}-σ magnitude limit: {}'.format(nframes*exposure, siglimit, bbmag1))
print(snr)
duet1_limit = bbmag1
tot_bgd_rate = bgd_band2 + rate2*surf_scale
for nframes in [8]:
snr = 100
swiftmag = 20.8
while snr > siglimit:
swiftmag += dmag
band1_fluence, band2_fluence = bb_abmag_fluence(duet =duet, swiftmag=swiftmag*u.ABmag, bbtemp=12e3*u.K)
band1_rate = duet.fluence_to_rate(band1_fluence)
band2_rate = duet.fluence_to_rate(band2_fluence)
src_rate = band2_rate
snr = duet.calc_snr(exposure, src_rate, tot_bgd_rate, nint=nframes)
bbmag1, bbmag2 = bb_abmag(swiftmag=swiftmag*u.ABmag, bbtemp=12e3*u.K, bandone = bandone, bandtwo=bandtwo)
print('Band2 {} {}-σ magnitude limit: {}'.format(nframes*exposure, siglimit, bbmag2))
print(snr)
print()
duet2_limit = bbmag2
# -
from astroduet import models
sims = models.Simulations()
sims.emgw_simulations
# +
target = 0.2 *u.day
lc = models.load_model_ABmag('kilonova_0.02.dat', dist = 10*u.pc)
duet1_abmag_at_5hr = np.interp(target.to(u.s).value, lc[0].to(u.s).value, lc[1])
duet2_abmag_at_5hr = np.interp(target.to(u.s).value, lc[0].to(u.s).value, lc[2])
hrs = lc[0].to(u.hr)
plt.figure()
plt.plot(hrs, lc[1])
plt.plot(hrs, lc[2])
plt.ylim([-12, -15])
plt.xlim([0, 10])
plt.xlabel('Hours')
plt.figure()
dist1 = 10 * 10**((duet1_limit.value - duet1_abmag_at_5hr) / 5)*u.pc
print(dist1.to(u.Mpc))
dist2 = 10 * 10**((duet2_limit.value - duet2_abmag_at_5hr) / 5)*u.pc
print(dist2.to(u.Mpc))
lc1 = models.load_model_ABmag('kilonova_0.02.dat', dist = dist1)
plt.plot(hrs, lc1[1])
lc2 = models.load_model_ABmag('kilonova_0.02.dat', dist = dist2)
plt.plot(hrs, lc2[2])
plt.axhline(duet1_limit.value, linestyle = ':')
plt.axhline(duet2_limit.value, linestyle = ':')
plt.axvline(target.to(u.hr).value, linestyle=':')
plt.ylim([24, 17])
plt.xlim([0, 10])
plt.xlabel('Hours')
plt.show()
lc = models.load_model_ABmag('shock_5e10.dat', dist = 10*u.pc)
duet1_abmag_at_5hr = np.interp(target.to(u.s).value, lc[0].to(u.s).value, lc[1])
duet2_abmag_at_5hr = np.interp(target.to(u.s).value, lc[0].to(u.s).value, lc[2])
hrs = lc[0].to(u.hr)
plt.figure()
plt.plot(hrs, lc[1])
plt.plot(hrs, lc[2])
plt.ylim([-12, -18])
plt.xlim([0, 10])
plt.xlabel('Hours')
plt.figure()
dist1 = 10 * 10**((duet1_limit.value - duet1_abmag_at_5hr) / 5)*u.pc
print(dist1.to(u.Mpc))
dist2 = 10 * 10**((duet2_limit.value - duet2_abmag_at_5hr) / 5)*u.pc
print(dist2.to(u.Mpc))
lc1 = models.load_model_ABmag('shock_5e10.dat', dist = dist1)
plt.plot(hrs, lc1[1])
lc2 = models.load_model_ABmag('shock_5e10.dat', dist = dist2)
plt.plot(hrs, lc2[2])
plt.axhline(duet1_limit.value, linestyle = ':')
plt.axhline(duet2_limit.value, linestyle = ':')
plt.axvline(target.to(u.hr).value, linestyle=':')
plt.ylim([24, 17])
plt.xlim([0, 10])
plt.xlabel('Hours')
plt.show()
# -
| notebooks/proposal_versions/EMGW Vs Host Threshold Ellipticals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 使用块的网络(VGG)
# :label:`sec_vgg`
#
# 虽然AlexNet证明深层神经网络卓有成效,但它没有提供一个通用的模板来指导后续的研究人员设计新的网络。
# 在下面的几个章节中,我们将介绍一些常用于设计深层神经网络的启发式概念。
#
# 与芯片设计中工程师从放置晶体管到逻辑元件再到逻辑块的过程类似,神经网络架构的设计也逐渐变得更加抽象。研究人员开始从单个神经元的角度思考问题,发展到整个层,现在又转向块,重复层的模式。
#
# 使用块的想法首先出现在牛津大学的[视觉几何组(visualgeometry group)](http://www.robots.ox.ac.uk/~vgg/)的*VGG网络*中。通过使用循环和子程序,可以很容易地在任何现代深度学习框架的代码中实现这些重复的架构。
#
# ## (**VGG块**)
#
# 经典卷积神经网络的基本组成部分是下面的这个序列:
#
# 1. 带填充以保持分辨率的卷积层;
# 1. 非线性激活函数,如ReLU;
# 1. 汇聚层,如最大汇聚层。
#
# 而一个VGG块与之类似,由一系列卷积层组成,后面再加上用于空间下采样的最大汇聚层。在最初的VGG论文中 :cite:`Simonyan.Zisserman.2014`,作者使用了带有$3\times3$卷积核、填充为1(保持高度和宽度)的卷积层,和带有$2 \times 2$汇聚窗口、步幅为2(每个块后的分辨率减半)的最大汇聚层。在下面的代码中,我们定义了一个名为`vgg_block`的函数来实现一个VGG块。
#
# + [markdown] origin_pos=1 tab=["tensorflow"]
# 该函数有两个参数,分别对应于卷积层的数量`num_convs`和输出通道的数量`num_channels`.
#
# + origin_pos=5 tab=["tensorflow"]
import tensorflow as tf
from d2l import tensorflow as d2l
def vgg_block(num_convs, num_channels):
blk = tf.keras.models.Sequential()
for _ in range(num_convs):
blk.add(tf.keras.layers.Conv2D(num_channels,kernel_size=3,
padding='same',activation='relu'))
blk.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2))
return blk
# + [markdown] origin_pos=6
# ## [**VGG网络**]
#
# 与AlexNet、LeNet一样,VGG网络可以分为两部分:第一部分主要由卷积层和汇聚层组成,第二部分由全连接层组成。如 :numref:`fig_vgg`中所示。
#
# 
# :width:`400px`
# :label:`fig_vgg`
#
# VGG神经网络连接 :numref:`fig_vgg`的几个VGG块(在`vgg_block`函数中定义)。其中有超参数变量`conv_arch`。该变量指定了每个VGG块里卷积层个数和输出通道数。全连接模块则与AlexNet中的相同。
#
# 原始VGG网络有5个卷积块,其中前两个块各有一个卷积层,后三个块各包含两个卷积层。
# 第一个模块有64个输出通道,每个后续模块将输出通道数量翻倍,直到该数字达到512。由于该网络使用8个卷积层和3个全连接层,因此它通常被称为VGG-11。
#
# + origin_pos=7 tab=["tensorflow"]
conv_arch = ((1, 64), (1, 128), (2, 256), (2, 512), (2, 512))
# + [markdown] origin_pos=8
# 下面的代码实现了VGG-11。可以通过在`conv_arch`上执行for循环来简单实现。
#
# + origin_pos=11 tab=["tensorflow"]
def vgg(conv_arch):
net = tf.keras.models.Sequential()
# 卷积层部分
for (num_convs, num_channels) in conv_arch:
net.add(vgg_block(num_convs, num_channels))
# 全连接层部分
net.add(tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(4096, activation='relu'),
tf.keras.layers.Dropout(0.5),
tf.keras.layers.Dense(4096, activation='relu'),
tf.keras.layers.Dropout(0.5),
tf.keras.layers.Dense(10)]))
return net
net = vgg(conv_arch)
# + [markdown] origin_pos=12
# 接下来,我们将构建一个高度和宽度为224的单通道数据样本,以[**观察每个层输出的形状**]。
#
# + origin_pos=15 tab=["tensorflow"]
X = tf.random.uniform((1, 224, 224, 1))
for blk in net.layers:
X = blk(X)
print(blk.__class__.__name__,'output shape:\t', X.shape)
# + [markdown] origin_pos=16
# 正如你所看到的,我们在每个块的高度和宽度减半,最终高度和宽度都为7。最后再展平表示,送入全连接层处理。
#
# ## 训练模型
#
# [**由于VGG-11比AlexNet计算量更大,因此我们构建了一个通道数较少的网络**],足够用于训练Fashion-MNIST数据集。
#
# + origin_pos=18 tab=["tensorflow"]
ratio = 4
small_conv_arch = [(pair[0], pair[1] // ratio) for pair in conv_arch]
# 回想一下,这必须是一个将被放入“d2l.train_ch6()”的函数,为了利用我们现有的CPU/GPU设备,这样模型构建/编译需要在strategy.scope()中
net = lambda: vgg(small_conv_arch)
# + [markdown] origin_pos=19
# 除了使用略高的学习率外,[**模型训练**]过程与 :numref:`sec_alexnet`中的AlexNet类似。
#
# + origin_pos=20 tab=["tensorflow"]
lr, num_epochs, batch_size = 0.05, 10, 128
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size, resize=224)
d2l.train_ch6(net, train_iter, test_iter, num_epochs, lr, d2l.try_gpu())
# + [markdown] origin_pos=21
# ## 小结
#
# * VGG-11使用可复用的卷积块构造网络。不同的VGG模型可通过每个块中卷积层数量和输出通道数量的差异来定义。
# * 块的使用导致网络定义的非常简洁。使用块可以有效地设计复杂的网络。
# * 在VGG论文中,Simonyan和Ziserman尝试了各种架构。特别是他们发现深层且窄的卷积(即$3 \times 3$)比较浅层且宽的卷积更有效。
#
# ## 练习
#
# 1. 打印层的尺寸时,我们只看到8个结果,而不是11个结果。剩余的3层信息去哪了?
# 1. 与AlexNet相比,VGG的计算要慢得多,而且它还需要更多的显存。分析出现这种情况的原因。
# 1. 尝试将Fashion-MNIST数据集图像的高度和宽度从224改为96。这对实验有什么影响?
# 1. 请参考VGG论文 :cite:`Simonyan.Zisserman.2014`中的表1构建其他常见模型,如VGG-16或VGG-19。
#
# + [markdown] origin_pos=24 tab=["tensorflow"]
# [Discussions](https://discuss.d2l.ai/t/1865)
#
| tensorflow/chapter_convolutional-modern/vgg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # K-Nearest Neighbor (KNN)
# ---
# Algoritma k-Nearest Neighbor adalah algoritma supervised learning dimana hasil dari instance yang baru diklasifikasikan berdasarkan mayoritas dari kategori k-tetangga terdekat.
#
# Tujuan dari algoritma ini adalah untuk mengklasifikasikan obyek baru berdasarkan atribut dan sample-sample dari training data.
#
# Algoritma k-Nearest Neighbor menggunakan Neighborhood Classification sebagai nilai prediksi dari nilai instance yang baru.
from luwiji.knn import illustration, demo
# +
demo.knn()
# Simulasi KNN berdasarkan
#parameter jarak dengan tetangga terdekat, banyak hubungan tetangga dan bobot jarak
# -
# Ada beberapa cara untuk menentukan kedekatan jarak dengan tetangga terdekat, salah satunya seperti di bawah ini :
illustration.knn_distance
# algoritma yang berbasis jarak nanti selanjutnya harus dilakukan feature scalling seperti normalisasi atau standarisasi agar skala valuenya tidak terlalu jauh antar kolom
# ### Other Distance Metric
# https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.DistanceMetric.html
| 04 - KNN & Scikit-learn/Part 1 - K Nearest Neighbor Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Mikaykay/DS-Unit-2-Linear-Models/blob/master/DS29_GP_Unit2_Sprint1_Module1_Nivi.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CPzo2e6aFrjx"
# # Lambda School Data Science - Unit 2 Sprint 1 Module 1
# + [markdown] id="2EVx8r5HLa6f"
# # I. Wrangle Data
# + id="xrpXzyGffCOA"
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
pd.options.display.max_rows = 1000
pd.options.display.max_columns = 1000
# + [markdown] id="au_SfudeJyBN"
# ## Write a function!
# + id="kyFh3n_ZzIHg"
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data'
def wrangle(filepath):
df = pd.read_csv(filepath,
na_values=[0.0],
parse_dates=['SALE_DATE'])
# Low cardinality
df.drop(columns=['NEIGHBORHOOD','BUILDING_CLASS_CATEGORY'],inplace=True)
#Dropping due to high cardinality
df.drop(columns=['ADDRESS','APARTMENT_NUMBER'],inplace=True)
return df
df = wrangle(DATA_PATH + '/condos/tribeca.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="JJhYyaULfcqA" outputId="525f6bef-ecbf-492c-dd26-d657ef5109a1"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="3N9ktYNvfqNh" outputId="dcb4e6d7-a463-46eb-8496-9e43557bc66f"
df['GROSS_SQUARE_FEET'].hist(bins=45)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="2tEOwJXEgD0D" outputId="2444d078-9cd1-4fd5-a16e-565cff0a6136"
df['YEAR_BUILT'].hist(bins=90)
# + [markdown] id="GgRcyfQMXhzJ"
# # II. Split Data
#
# Split our dataset into a **feature matrix `X`** and a **target vector `y`**.
# + id="dusfK5DEnHPz"
# Use a double square brackets to create a 2D matrix and a single square bracket when you need a vector.
X = df[['GROSS_SQUARE_FEET']]
#target vector
y = df['SALE_PRICE']
# + colab={"base_uri": "https://localhost:8080/"} id="GmaCkZLsncvL" outputId="88c8e9d5-94c1-4238-dfc0-45cd88d18c7e"
X.shape
# + colab={"base_uri": "https://localhost:8080/"} id="hufmuJmGoPxV" outputId="7418e67e-894d-4dc3-9590-ee15db9ab201"
y.shape
# + [markdown] id="CuwsiXRlYsjG"
# # III. Establish Baseline
#
# If you had a *naïve model* that could only predict one value, how well would you model perform?
# + colab={"base_uri": "https://localhost:8080/"} id="zjDk6v43oamA" outputId="a5d07676-45a9-4e8f-b0f7-cab5799e1726"
y.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="ASaeKSTrpEuR" outputId="b83cfd01-88da-478d-8542-8121dbb40862"
y.mean()
# + id="f1HOUiHrqM0H"
y_baseline = [y.mean()]*len(y)
# + colab={"base_uri": "https://localhost:8080/", "height": 307} id="3E5sMPAIpMrY" outputId="508a43dc-2c7c-4cdd-f051-d4ac7856d6cd"
plt.scatter(X,y)
plt.plot(X,y_baseline,label='Baseline',color='red')
plt.xlabel('Gross sq ft')
plt.ylabel('Sale Price')
plt.legend()
# + [markdown] id="3bFjzSkUdl21"
# # IV. Build Model
# + colab={"base_uri": "https://localhost:8080/"} id="WFDStQ7DsV_h" outputId="4b4a23c2-a984-4641-a0ec-68414fc77232"
#Step1: Import library to use our machine learning algorithm
from sklearn.linear_model import LinearRegression
#Step2: "Instantiate" your model and set parameters.
model_lr = LinearRegression()
#Step3: Fit your model for the given data (Training)
model_lr.fit(X,y)
# + id="TArH0MD0tZ9k"
#dir(model_lr)
# + colab={"base_uri": "https://localhost:8080/"} id="FMMpL1Z2tusD" outputId="d6ff6cc8-af86-4309-d1e0-5d6f7ecc6c9f"
model_slope = model_lr.coef_[0]
print(model_slope)
# + colab={"base_uri": "https://localhost:8080/"} id="7NqgDxg4uET5" outputId="d18380f4-9ff5-46a2-a99c-a1f633a780fd"
model_intercept = model_lr.intercept_
print(model_intercept)
# + [markdown] id="Lo3CypnUfIwc"
# # V. Check Metrics
#
#
# + [markdown] id="YOu_MKTiu0sw"
# 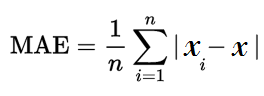
# + colab={"base_uri": "https://localhost:8080/"} id="Ov7F6bSYvUxn" outputId="98a4bce1-3cde-401b-96d2-9ed89b56fd8d"
from sklearn.metrics import mean_absolute_error
print("BASELINE MAE:", mean_absolute_error(y,y_baseline))
# + id="_vUxHC63vsrc"
y_pred_lr = model_lr.predict(X)
# + colab={"base_uri": "https://localhost:8080/"} id="ic3qcI-2v0tw" outputId="84e13b43-5a0a-457d-81d8-ca37b2d6a69d"
y_pred_lr[:20]
# + colab={"base_uri": "https://localhost:8080/"} id="_9X3FNzsv0mI" outputId="8493bb2a-86c1-43ea-e39e-b47ffc1d073b"
from sklearn.metrics import mean_absolute_error
print("BASELINE MAE:", mean_absolute_error(y,y_pred_lr))
# + [markdown] id="w5D3570TZ1oe"
# ### Check the baseline metric
# + [markdown] id="jGYUL1mufdS-"
# # VI. Communicate Results
#
# Plot model
# + colab={"base_uri": "https://localhost:8080/", "height": 307} id="q15rB884wMN_" outputId="07de75b1-a102-4a96-fe87-baef6c385dfa"
plt.scatter(X,y)
plt.plot(X,y_baseline,label='Baseline',color='red')
plt.plot(X,y_pred_lr,label='Linear Regression',color='yellow')
plt.xlabel('GRoss sq ft')
plt.ylabel('SALE price')
plt.legend()
# + [markdown] id="5036hzreQkxd"
# So what is our equation?
# $$\hat{y}=-1505364+3076*SQFT$$
| DS29_GP_Unit2_Sprint1_Module1_Nivi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (walk-env)
# language: python
# name: walk-env
# ---
# # Initial Exploratory Data Analysis
# ## Load necessary packages
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
# %matplotlib inline
# ## Helper functions
def expand_date(dataframe):
"""expands the date to add columns with year, month, day, weekday
Args:
dataframe: a pandas dataframe with column 'date', among others
Returns:
a copy of the dataframe with out the date column, and with the
new year, month, day, weekday columns
"""
df_copy = dataframe.copy()
X = pd.to_datetime(df_copy['date'], format="%Y-%m-%d")
df_copy['year'] = [x.year for x in X]
df_copy['month'] = [x.month for x in X]
df_copy['day'] = [x.day for x in X]
df_copy['weekday'] = [x.weekday() for x in X]
df_copy = df_copy.drop(columns=['date'])
print(df_copy.head())
return df_copy
def plot_w_best_fit_line(x, y):
"""plots two-dimensional data with a dashed best fit line.
Args:
x: x-values
y: y-values
Returns:
none
"""
coef = np.polyfit(x,y,1)
poly1d_fn = np.poly1d(coef)
# poly1d_fn is now a function which takes in x and returns an estimate for y
plt.plot(x,y, 'yo', x, poly1d_fn(x), '--k')
# ## Load data from SQL database
df = pd.read_sql_table('manhattan_loc_d_ar_wea', 'postgresql:///walk')
# ## Data Cleaning
# fill n_arrests with 0 in place of NaN
df['n_arrests'] = df['n_arrests'].fillna(value=0).astype(int)
# expand date features
df_expanded = expand_date(df)
# +
# ozone levels were not recorded for the earlier years of the data set
# they do not seem to be correlated with the arrest count
# decided to drop ozone readings.
# df_expanded = df_expanded.drop(columns=['ozone10'])
# the remaining features all have NaN in place of zero. Filling NaNs with 0s
df_expanded.fillna(0)
# -
from statsmodels.formula.api import ols
# ## Looking at single-feature linear regression for each feature
for col in df_expanded.columns:
outcome = 'n_arrests'
predictor = col
formula = outcome + '~' + predictor
model = ols(formula=formula, data=df_expanded).fit()
print(model.summary())
| notebooks/03_eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
from torchcox import TorchCox
import numpy as np
import torch
import pandas as pd
# -
df_tied = pd.read_csv('../data/ovarian_deduplicated.csv')
df_tied
valdf = pd.DataFrame({'id':['Bob','Sally','James','Ann'], 'time':[1,3,6,10], 'status':[1,1,0,1], 'smoke':[1,0,0,1]})
valdf
# +
tname = 'time'
Xnames = ['smoke']
dname = 'status'
coxmod1 = TorchCox.TorchCox(lr=1)
coxmod1.fit(valdf, Xnames=Xnames, tname=tname, dname=dname)
coxmod1.beta.detach().numpy()[0]
# -
# MLE of Cox model on simple dataset above has closed-form solution, which is ln(2)/2. The result above should equal this, which reassuringly it does.
np.log(2)/2
# Now we will compare our result against the R package 'survival' on another dataset.
# +
coxmod = TorchCox.TorchCox()
tname = 'tyears'
Xnames = ['Karn', 'Ascites']
dname = 'd'
# -
# %load_ext rpy2.ipython
# + language="R"
# library(readr)
# library(survival)
# library(dplyr)
# library(tidyr)
#
# df2 = read_csv("/home/ilan/Desktop/TorchCox/data/ovarian_deduplicated.csv")
#
# #df2 = df2 %>% arrange(tyears)
#
# starttime = Sys.time()
#
# rmod = coxph(Surv(tyears, d) ~ Karn + Ascites, df2, ties="breslow")
# print(coef(rmod))
#
# endtime = Sys.time()
#
# print(endtime-starttime)
# +
# %%time
coxmod.fit(df_tied, Xnames=Xnames, tname=tname, dname=dname, basehaz=False)
# -
# We indeed match that result as well!
# We are about 10x slower than the R package (which runs in C). But simplicity and extensibility of our code compensates for that in my view. Also timings here most likely dominated by overhead of loading libraries, comparison on larger dataset required.
# Fit again, but this time computing the baseline hazard, to ensure that works as well.
# +
# %%time
coxmod.fit(df_tied, Xnames=Xnames, tname=tname, dname=dname, basehaz=True)
# -
coxmod.basehaz
# Predict on the training set to ensure predict_proba() method works as well.
df_tied['pred'] = coxmod.predict_proba(df_tied, Xnames=Xnames, tname=tname)
df_tied
| notebooks/Torch_Cox_package_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sistema de Ecuaciones Lineales
# ## Factorización PALU y Cholesky
import numpy as np
import matplotlib.pyplot as plt
import sympy as sym
from scipy import linalg as spla
from scipy import sparse as scsp
from time import time
# ## Número de Condición
# ### Matriz de Hilbert
#
# Matriz cuadrada simétrica y positiva definida, cuyas entradas se definen como:
# \begin{equation}
# H_{ij} = \frac{1}{i+j-1}
# \end{equation}
#
# Por ejemplo, la matriz $H$ de $5\times 5$ es
# \begin{equation}
# H=
# \begin{bmatrix}
# 1 & \frac {1}{2} & \frac {1}{3} & \frac {1}{4} & \frac {1}{5}\\
# \frac {1}{2} & \frac {1}{3} & \frac {1}{4} & \frac {1}{5} & \frac {1}{6}\\
# \frac {1}{3} & \frac {1}{4} & \frac {1}{5} & \frac {1}{6} & \frac {1}{7}\\
# \frac {1}{4} & \frac {1}{5} & \frac {1}{6} & \frac {1}{7} & \frac {1}{8}\\
# \frac {1}{5} & \frac {1}{6} & \frac {1}{7} & \frac {1}{8} & \frac {1}{9}
# \end{bmatrix}
# \end{equation}
n = 5
H = spla.hilbert(n)
# ¿Simétrica?
np.all(H == H.T)
# ¿Definida positiva?
np.all(np.linalg.eigvals(H) > 0)
# Cuidado ver lo que pasa para valores de $n$ grandes.
# Número de condición $\kappa(A)$
np.linalg.cond(H), np.max(np.abs(np.linalg.eigvals(H)))/np.min(np.abs(np.linalg.eigvals(H)))
# En teoría esta matriz es simétrica y definida positiva, en la práctica para $n$ grandes no se cumple la ultima propiedad. Además es una matriz mal condicionada. ¿Qué pasará si debemos resolver un sistema de ecuaciones lineales con esta matriz?
# # Matriz definida positiva
# Una manera de comprobar si una matriz es definida positiva es calcular los valores propios y comprobar que todos estos sean positivos. Veamos el caso para $n=2$.
np.random.seed(2)
n = 2
A = np.random.rand(n, n)
A = np.dot(A, A.T) + np.eye(n)
eigenvalues = np.linalg.eig(A)[0]
print(eigenvalues)
print(eigenvalues > 0)
# La otra forma es comprobar $\mathbf{x}^{T} A \mathbf{x} > 0$, $\mathbf{x}\neq\mathbf{0}$, para eso podemos hacer uso de simpy:
x, y = sym.symbols('x y')
x = np.array([x,y])
f = sym.factor(sym.expand(np.dot(np.dot(x,A),x)))
fl = sym.lambdify(x, f)
f
# Si analizamos el polinomio anterior tenemos:
#
# \begin{equation}
# \begin{split}
# 0.798294611703302 x^{2} + 0.336458187477782 xy + y^2 & \\
# 0.798294611703302 x^{2} + 0.336458187477782 xy + y^2 + 0.028301027980208573x^2 - 0.028301027980208573x^2 & \\
# 0.7699935837230933 x^{2} + (0.168229093738891x + y)^{2} &
# \end{split}
# \end{equation}
# El polinomio es positivo para cualquier valor de $x,y$, entonces estamos en presencia de una matriz definida positiva
# Otra idea, pero que podría ser más complicado para $\mathbb{R}^{n}$ con $n>3$, es utilizar el criterio de la segunda derivada (en este caso parcial).
fx = sym.diff(f, x[0])
fy = sym.diff(f, x[1])
fxy = sym.diff(fx, x[1])
fyx = sym.diff(fy, x[0])
fxx = sym.diff(fx, x[0])
fyy = sym.diff(fy, x[1])
fxxl = sym.lambdify(x, fxx)
fxyl = sym.lambdify(x, fxy)
fyyl = sym.lambdify(x, fyy)
D = lambda a, b: fxxl(a, b) * fyyl(a, b) - (fxyl(a, b)) ** 2
fx, fy
sym.solve([fx, fy])
D(0, 0), fxxl(0, 0)
fl(0,0)
# La función tienen un mínimo en $(x,y)=(0,0)$ pero por definición no nos interesa ese punto, así que la función efectivamente es positiva. Por lo tanto $A$ es definida positiva.
# # Complejidad temporal y espacial
# Estimación del tamaño del arreglo en [MB]
size = lambda a: a.nbytes * 1e-6
Ne = 10 # Repetición de experimentos
Nf = 10
N = 2 ** np.arange(7, Nf + 1) # N = [2^7, 2^{10}]
Nn = N.shape[-1]
# Para tiempos
times_palu = np.zeros(Nn)
times_chol = np.zeros(Nn)
# Para espacio
storage_palu = np.zeros(Nn)
storage_chol = np.zeros(Nn)
# ## Experimentos
for i in range(Nn):
n = N[i]
A = np.random.rand(n, n)
A = np.dot(A, A.T) + np.eye(n)
# Time PALU
start_time= time()
for j in range(Ne):
P, L, U = spla.lu(A)
end_time = time()
storage_palu[i] = size(P) + size(L) + size(U)
times_palu[i] = (end_time - start_time) / Ne
# Time Cholesky
start_time = time()
for j in range(Ne):
R = np.linalg.cholesky(A) # R corresponde a R^T
end_time = time()
times_chol[i] = (end_time - start_time) / Ne
# Storage
storage_chol[i] = size(R)
# ## Análisis de tiempo computacional
plt.figure(figsize=(12, 6))
plt.plot(N, times_palu, 'bd', label="PALU")
plt.plot(N, times_chol, 'go', label="Cholesky")
# Deben adaptar el coeficiente que acompaña a N**k según los tiempos que obtengan en su computador
plt.plot(N, 1e-8 * N ** 2, 'g--', label=r"$O(n^2)$")
plt.plot(N, 1e-10 * N ** 3, 'r--', label=r"$O(n^3)$")
plt.grid(True)
plt.yscale('log')
plt.xscale('log')
plt.xlabel(r"$n$")
plt.ylabel("Time [s]")
plt.legend()
plt.show()
# Los algoritmos tienen una complejidad que tiende a $O(n^3)$. Dado que hay optimizaciones en las bibliotecas, pueden ver exponentes menores a $3$. En teoría el análisis debería funcionar para $n\to\infty$.
# ## Análisis de memoria
plt.figure(figsize=(12, 6))
plt.plot(N, storage_palu, 'bd', label="PALU")
plt.plot(N, storage_chol, 'go', label="Cholesky")
plt.plot(N, 1.5e-5 * N ** 2, 'g-', label=r"$O(n^2)$")
plt.grid(True)
plt.yscale('log')
plt.xscale('log')
plt.xlabel(r"$n$")
plt.ylabel("Size [GB]")
plt.legend()
plt.show()
# Si guardamos todos los elementos de las matrices, el espacio utlizado debería ser del orden $\sim n^2$. ¿Existirá una manera de ahorrar espacio?
# # Alternativas de almacenamiento
# En el caso de $PALU$ debemos almacenar $P$, $L$ y $U$ $\sim 3n^2$. Con Cholesky solo debemos almacenar $\sim n^2$.
# Dado que hay muchos elementos iguales a $0$, podríamos almacenar de manera eficiente solo los valores distintos de $0$. Las matrices que tienen muchos valores que son $0$ se conocen como **Sparse Matrix**, *Matriz dispersa o rala* en español. $L$, $U$ y $R$ son matrices triangulares, y $P$ solo tiene $n$ elementos distintos de $0$.
#
# Una matriz triangular, tiene $\displaystyle n^2 - \sum_{i=1}^{n-1}i=n^2 - \frac{n(n-1)}{2}=\frac{n^2+ n}{2}$ elementos. Es decir, con una representación *dispersa*, $PALU$ requiere mantener $n^2 + 2n$ elementos mientras que Cholesky solo requiere $\displaystyle \frac{n^2+ n}{2}$.
# ## Almacenamiento *Sparse*
#
# ### Coordinate list (COO)
#
# Se almacena una lista de tuplas con $(fila, columna, valor)$.
#
# ### Compressed sparse row (CSR)
#
# La matriz se almacena por fila. Se mantienen 3 arreglos, $valores$, $indice\_columnas$, $puntero\_fila$. El primer arreglo guarda los valores no nulos, el segundo guarda el índice de la columna donde se encuentra el coeficiente y el último indica qué parte del primer y segundo arreglo corresponde a cada fila.
#
# ### Compressed sparse column (CSC)
#
# Análogo al anterior pero se almacena por columnas. Además el segundo arreglo mantiene el $indice\_filas$ y el tercero $puntero\_columna$.
# ## Ejemplo
M = np.array([[1, 0, 3, 0], [0, 1, 8, 0], [0, 0, 0, 0], [0, 1, 0, 1]])
M
# ### COO
M_coo = scsp.coo_matrix(M)
print("Datos:", M_coo.data)
print("Indice filas:", M_coo.row)
print("Indice columnas:", M_coo.col)
# ### CSR
M_csr = scsp.csr_matrix(M)
print("Datos:", M_csr.data)
print("Indice columnas:", M_csr.indices)
print("Puntero filas:", M_csr.indptr)
for i in range(M_csr.indptr.shape[-1]-1):
print("Los elementos de la fila %d, se encuentran entre las posiciones [%d, %d) del arreglo de indices" % (i, M_csr.indptr[i], M_csr.indptr[i+1]))
# ### CSC
M_csc = scsp.csc_matrix(M)
print("Datos:", M_csc.data)
print("Indice filas:", M_csc.indices)
print("Puntero columnas:", M_csc.indptr)
for i in range(M_csc.indptr.shape[-1]-1):
print("Los elementos de la columna %d, se encuentran entre las posiciones [%d, %d) del arreglo de indices" % (i, M_csc.indptr[i], M_csc.indptr[i+1]))
# Los formatos *CSR* y *CSC* necesitan almacenar $2nz+n+1$ elementos distintos de $0$ ($nz$).
# ## Matrices generadas por los métodos de factorización
# Para la visualización de las matrices.
def plotMatrix(M):
# Solo para ver coeficientes distintos de 0, se muestra la magnitud -> log(|M+eps|), eps para evitar el log(0)
plt.imshow(np.log(np.abs(M+1e-16)))
plt.show()
plotMatrix(A)
# ## PALU
plotMatrix(P); plotMatrix(L); plotMatrix(U)
# ### Matrices densas
palu_dense = size(P) + size(L) + size(U)
palu_dense
# ### Matrices dispersas
# #### Formato Coordenadas
Pc = scsp.coo_matrix(P)
Lc = scsp.coo_matrix(L)
Uc = scsp.coo_matrix(U)
palu_sparse_co = size(Pc.data) + size(Pc.row) + size(Pc.col) + size(Lc.data) + size(Lc.row) + size(Lc.col) + size(Uc.data) + size(Uc.row) + size(Uc.col)
palu_sparse_co
# #### Formato Comprimido
Psr = scsp.csr_matrix(P)
Lsr = scsp.csr_matrix(L)
Usr = scsp.csr_matrix(U)
Psc = scsp.csc_matrix(P)
Lsc = scsp.csc_matrix(L)
Usc = scsp.csc_matrix(U)
palu_sparse_r = size(Psr.data) + size(Psr.indices) + size(Psr.indptr) + size(Lsr.data) + size(Lsr.indices) + size(Lsr.indptr) + size(Usr.data) + size(Usr.indices) + size(Usr.indptr)
palu_sparse_r
len(Psr.data)
palu_sparse_c = size(Psc.data) + size(Psc.indices) + size(Psc.indptr) + size(Lsc.data) + size(Lsc.indices) + size(Lsc.indptr) + size(Usc.data) + size(Usc.indices) + size(Usc.indptr)
palu_sparse_c
# % memoria matrices *densas* vs *dispersas* utilizando coordenadas
palu_sparse_co / palu_dense
# % memoria matrices *densas* vs *dispersas* utilizando formato comprimido
palu_sparse_r / palu_dense
# ## Cholesky
plotMatrix(R)
# ### Matriz densa
cholesky_dense = size(R)
cholesky_dense
# ### Matriz dispersa
# #### Formato Coordenadas
Rc = scsp.coo_matrix(R)
# #### Formato Comprimido
Rsr = scsp.csr_matrix(R)
Rsc = scsp.csc_matrix(R)
# Tamaño utilizando representación *sparse*
cholesky_sparse_co = size(Rc.data) + size(Rc.row) + size(Rc.col)
cholesky_sparse_co
cholesky_sparse_r = size(Rsr.data) + size(Rsr.indices) + size(Rsr.indptr)
cholesky_sparse_r
cholesky_sparse_c = size(Rsc.data) + size(Rsc.indices) + size(Rsc.indptr)
cholesky_sparse_c
# % de memoria utilizando representación *dispersa* formato coordenadas
cholesky_sparse_co / cholesky_dense
# % memoria utilizando representación *dispersa* comprimida.
cholesky_sparse_r / cholesky_dense
# ### Comparación
#
# Si analizamos teóricamente la memoria que podemos ahorrar para ambos tipos de factorizaciones utilizando la representación *sparse* tenemos:
#
# * $PALU$:
# \begin{equation}
# \frac{\text{Representación dispersa}}{\text{Representación densa}}=
# \frac{n^2 + 2n}{3n^2} = \frac{n+2}{3n} = \frac{1}{3} + \frac{2}{3n}
# \implies \lim_{n\to\infty} \left(\frac{1}{3} + \frac{2}{3n}\right) = \dfrac{1}{3}
# \end{equation}
#
# * Cholesky
# \begin{equation}
# \frac{\text{Representación dispersa}}{\text{Representación densa}}=
# \frac{\frac{n^2 + n}{2}}{n^2} = \frac{n+1}{2n} = \frac{1}{2} + \frac{1}{2n}
# \implies \lim_{n\to\infty} \left(\frac{1}{2} + \frac{1}{2n}\right) = \dfrac{1}{2}
# \end{equation}
#
# Notar que esto es válido solo si almacenamos los coeficientes distintos de $0$. En la práctica estas representaciones deben guardar información adicional sobre la posición de los elementos, entre otros.
# Caso $PALU$ (considerando valores no nulos)
(Pc.data.shape[-1] + Lc.data.shape[-1] + Uc.data.shape[-1]) / (3 * N[-1] ** 2)
# Caso Cholesky (considerando valores no nulos)
Rc.data.shape[-1] / N[-1] ** 2
# Se hace énfasis en que este análisis solo compara los valores de coeficientes, pero no se incluye el tamaño de las estructuras adicionales que requieren estas representaciones.
# ## Resolución sistema de ecuaciones
# ### Sistema de ecuaciones lineales
#
# \begin{equation}
# A \mathbf{x} = \mathbf{b}
# \end{equation}
#
# ### Resolución utilizando $PA=LU$
#
# \begin{equation}
# \begin{split}
# A\, \mathbf{x} & = \mathbf{b} \\
# PA\, \mathbf{x} & = P\,\mathbf{b} \\
# LU\, \mathbf{x} & = P\, \mathbf{b} \\
# L\, \mathbf{c} & = P\, \mathbf{b}
# \end{split}
# \end{equation}
#
# Algoritmo:
# 1. Obtener descomposición PALU: $PA=LU$
# 2. Resolver para $\mathbf{c}$: $L\,\mathbf{c} = P\, \mathbf{b}$
# 3. Resolver para $\mathbf{x}$: $U\,\mathbf{x} = \mathbf{c}$
# 4. Retornar $\mathbf{x}$
#
# ### Resolución utilizando Cholesky
#
# \begin{equation}
# \begin{split}
# A \, \mathbf{x} & = \mathbf{b} \\
# R^T\,R \,\mathbf{x} & = \mathbf{b}
# \end{split}
# \end{equation}
#
# Algoritmo:
# 1. Obtener descomposición de Cholesky: $A=R^T\,R$
# 2. Resolver para $\mathbf{c}$: $R^T\mathbf{c} = \mathbf{b}$
# 3. Resolver para $\mathbf{x}$: $R\,\mathbf{x} = \mathbf{c}$
# 4. Retornar $\mathbf{x}$
# ## Ejemplo
# ¿Cómo se resuelven sistemas utilizando las factorizaciones $PA=LU$ y Cholesky?
n = 100 # Probar con distintos valores de n
A = np.random.rand(n, n)
A = np.dot(A, A.T) + np.eye(n)
# Luego de probar con la matriz A simétrica y definida positiva, probar con la matriz de Hilbert...
#A = spla.hilbert(n)
np.linalg.cond(A)
x = np.arange(1, n+1) # Solucion
b = np.dot(A, x)
# +
#x
# -
np.all(A == A.T)
np.all(np.linalg.eigvals(A) > 0)
x_n = np.linalg.solve(A, b)
# ### $PA=LU$
P, L, U = spla.lu(A)
Pb = np.dot(P.T, b) # Permutacion
c = spla.solve_triangular(L, Pb, lower=True) # Lc = Pb
x_p = spla.solve_triangular(U, c) # Ux = c
np.linalg.norm(np.dot(P, np.dot(L, U)) - A) # Verificar que PLU = A
# ### Cholesky
R = np.linalg.cholesky(A) # Entrega R^T de acuerdo a nuestras diapositivas
c = spla.solve_triangular(R, b, lower=True) # R^Tc = b
x_c = spla.solve_triangular(R.T, c) # Rx = c
np.linalg.norm(np.dot(R, R.T) - A) # Veridiar que R^TR = A
# +
# Descomentar para ver soluciones
#print(x_n)
#print(x_p)
#print(x_c)
# -
# ## Backward y Forward Error
# Analizar el error de las soluciones...
backwardError = lambda A, x_c, b: np.linalg.norm(b - np.dot(A, x_c), np.inf)
forwardError = lambda x, x_c: np.linalg.norm(x - x_c, np.inf)
backwardError(A, x_n, b), forwardError(x, x_n) # Solver de NumPy
backwardError(A, x_p, b), forwardError(x, x_p) # PALU
backwardError(A, x_c, b), forwardError(x, x_c) # Cholesky
| material/04_sistemas_ecuaciones/palu_cholesky.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from numpy import linalg as LA
# %matplotlib inline
import scipy.sparse.linalg as SA
import scipy.sparse as S
def V(r, f = 1e-1):
"""
Coulomb potential between two unit charges of same sign in units of Hartree,
if r is in bohr radii. The parameter smoothes the divergence of the potential in 1D."""
return 1 / (f + abs(r))
def build_H2(grid, ions = []):
dx = grid[1] - grid[0]
size = len(grid)
x1, x2 = np.meshgrid(grid, grid)
v = V(x1 - x2)
for Z, k in ions:
v += -Z * (V(x1 - k) + V(x2 - k))
units = 0.5 # Ha = h_bar²/2m_e / r_0²
o = units / dx**2 * np.ones(size ** 2)
H = np.diag(4 * o + v.flat, 0) + \
- np.diag(o[1:], -1) \
- np.diag(o[1:], +1) \
- np.diag(o[n:], -n) \
- np.diag(o[n:], +n)
for i in range(n - 1, n**2 - 1, n):
H[i, i + 1] = 0
H[i + 1, i] = 0
return H
def solve(H):
e, v = LA.eigh(H)
v = v[:, np.argsort(e)]
e = e[np.argsort(e)]
return e, v
def solve2d(H):
e, v = solve(H)
n = int(np.sqrt(len(e)))
return e, v.T.reshape(-1, n, n)
n = 52
x = np.linspace(-5, 5, n)
H0 = build_H2(x)
H1 = build_H2(x, [(2, 0.0)])
H2 = build_H2(x, [(1, 0.5), (1, -0.5)])
plt.matshow(np.diag(H1).reshape(n, n))
plt.matshow(H0)
e1, v1 = solve2d(H1)
plt.plot(e1[:10], "-o")
plt.matshow(v1[3])
plt.colorbar()
for i in range(5):
plt.plot(x, (abs(v1[i])**2).sum(axis = 0),
label = i)
plt.legend()
| Interacting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import random as random
sum([5,7])
def gravity(r,m1,m2):
G=6.67*10**-11
F=G*m1*m2/r**2
return F
gravity(2,4,3)
gravity(10,100000,1000000)
import matplotlib.pyplot as plt
import seaborn as sns
import random as random
# %matplotlib inline
sns.set()
import random as random
x = 0
y = 0
X = [x]
Y = [y]
n=1
isprint = False
while n < 1000000:
r = random.uniform(0,100)
if r < 1.0:
x = 0
y = 0.16*Y[n-1]
X.append(x) ; Y.append(y)
elif r > 1.0 and r < 86.0:
x = 0.85*X[n-1] + 0.04*Y[n-1]
y = -0.04*X[n-1] + 0.85*Y[n-1]+1.6
X.append(x);Y.append(y)
elif r > 86.0 and r < 93.0:
x = 0.2*X[n-1] - 0.26*Y[n-1]
y = 0.23*X[n-1] + 0.22*Y[n-1] + 1.6
X.append(x);Y.append(y)
elif r > 93.0 and r < 100.0:
x = -0.15*X[n-1] + 0.28*Y[n-1]
y = 0.26*X[n-1] + 0.24*Y[n-1] + 0.44
X.append(x);Y.append(y)
if isprint:
print("step: ",n,"random number is: ", r, "coordinate is : ", x,y)
n = n+1
plt.figure(figsize = [5,8])
plt.scatter(X,Y,color = 'g',marker = '.')
plt.show()
| sum and gravity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/neurologic/MotorSystems_BIOL358_SP22/blob/main/BasisSets.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="P7b-VwcyRkbv"
# We have talked a lot about "basis sets" this semester, and it seems like a concept that many are still struggling to build intuition for and visualize. This concept is particularly important for understanding what the cerebellum is doing for adaptive motor control (and whatever other adaptive control functions it is involved in). This notebook providees tools to examine how signals of a basis set can be combined in different proportions to produce a variety of output signals.
# + cellView="form" id="_rdZWOYuJsPd"
#@markdown TASK: run this code cell to import packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from datetime import datetime,timezone,timedelta
import ipywidgets as widgets # interactive display
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
my_layout = widgets.Layout()
# + colab={"base_uri": "https://localhost:8080/", "height": 575} cellView="form" id="Ez4LBxwuKEq9" outputId="d83df27d-09ee-460d-b61e-062f2d45ad1d"
#@markdown TASK: set the frequency for two signals in your basis set
frequency1 = 5 #@param
frequency2 = 10 #@param
#@markdown TASK: Run this code cell to plot the two signals
# sampling rate
sr = 500.0
# sampling interval
ts = 1.0/sr
t = np.arange(0,1,ts)
# frequency of the signal
y1 = np.sin(2*np.pi*frequency1*t)
y2 = np.sin(2*np.pi*frequency2*t)
plt.figure(figsize = (8, 8))
plt.subplot(211)
plt.plot(t, y1, 'purple')
plt.ylabel('Amplitude')
plt.subplot(212)
plt.plot(t, y2, 'green')
plt.ylabel('Amplitude')
plt.xlabel('Time (s)')
plt.show()
# + [markdown] id="VQ5IrXuKLgM2"
# You can think about these two signals as parallel fiber inputs to a post-synaptic cell. The membrane potential response of the post-synaptic cell will change based on the synaptic weight from each input. The set of inputs to the postsynaptic cell is a "basis set" (a set of signals that can be combinned in different proportions to create many different signals).
#
# $$
# response = {baseline\ spike\ rate} + {(w_1 * y1)} + {(w_2 * y2)}
# $$
# + colab={"base_uri": "https://localhost:8080/", "height": 575} cellView="form" id="iXFFqxrPKLRD" outputId="0cc1bedc-7006-4e10-b601-6657664dc2d5"
#@markdown TASK: set the synaptic weight for each input
w1 = 2#@param
w2 = 3#@param
#@markdown You can also change the baseline spike rate of the post-synaptic cell
baseline_rate = 5 #@param
#@markdown TASK: run this code cell to plot the response
#@markdown of the purkinje cell to these inputs
response = baseline_rate + w1*y1 + w2*y2
plt.figure(figsize = (8, 8))
plt.subplot(211)
plt.plot(t, w1*y1, 'purple')
plt.plot(t, w2*y2, 'green')
plt.ylabel('Input Amplitude')
plt.subplot(212)
plt.plot(t, response, 'k')
plt.ylabel('Response Amplitude')
plt.xlabel('Time (s)')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 784, "referenced_widgets": ["f92d162574224368aadc335235824070", "e91192de1177448ea2d099e1206103e9", "4054c598e4414d0492539e6d47dfd88f", "a62222afb8a44aeb85e2987a5ce74b30", "<KEY>", "a47dbb9a71cd4f1bbece9d5268f60989", "eaa5c9dce2f848289ea47be66903784f", "c1adafc7a0284b50860c0c60fbe704d2", "43c1702fc4d545cc9f1f6e1fe6cb0ea2", "510d2531735246558b1bd57bfd58a0ff", "a879afe581d5486494bc3a7656961e73", "7533e7a660cc49428acb8bb1ce8f3d4f", "8458ca8e05874212ba91c5a29bf8e999", "<KEY>", "1e2a21892abd42c1aa651f992841ced0", "<KEY>", "e9e6434aa65a4bd1873d0ac95a9f27f8"]} cellView="form" id="RE79CkzOOdFz" outputId="c248999c-4f12-4230-b115-d32889cab122"
#@markdown TASK: run this code cell and then slide the widgets
#@markdown to set the synaptic weight on each of 5 different basis set signals
#@markdown > The plots of each input and the net response of the postsynaptic cell will update accordingly
my_layout.width = '450px'
@widgets.interact(
w1=widgets.FloatSlider(0., min=0., max=10., step=1.,
layout=my_layout),
w2=widgets.FloatSlider(0., min=0., max=10., step=1.,
layout=my_layout),
w3=widgets.FloatSlider(0., min=0., max=10., step=1.,
layout=my_layout),
w4=widgets.FloatSlider(0., min=0., max=10., step=1.,
layout=my_layout),
w5=widgets.FloatSlider(0., min=0., max=10., step=1.,
layout=my_layout),
baseline_rate=widgets.FloatSlider(0., min=0., max=10., step=1.,
layout=my_layout)
)
def basis_set_combine(w1,w2,w3,w4,w5,baseline_rate):
y1 = np.sin(2*np.pi*2*t)
y2 = np.sin(2*np.pi*4*t)
y3 = np.sin(2*np.pi*6*t)
y4 = np.sin(2*np.pi*8*t)
y5 = np.sin(2*np.pi*10*t)
response = baseline_rate + w1*y1 + w2*y2 + w3*y3 + w4*y4 + w5*y5
plt.figure(figsize = (8, 8))
plt.subplot(211)
plt.plot(t, w1*y1, 'purple')
plt.plot(t, w2*y2, 'green')
plt.plot(t, w3*y3, 'orange')
plt.plot(t, w4*y4, 'brown')
plt.plot(t, w5*y5, 'blue')
plt.ylabel('Input Amplitude')
plt.subplot(212)
plt.plot(t, response, 'k')
plt.ylabel('Response Amplitude')
plt.xlabel('Time (s)')
plt.show()
print('Interactive demo initiated at ' + str(datetime.now(timezone(-timedelta(hours=5)))))
| BasisSets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Creating co-occurence network for visualization with Gephi
# -
import pandas as pd
naver_df = pd.read_csv('/Users/SeoyeonHong/Desktop/text_mining/topic_modeling/naver_kin/naver_q1_topic_representation.tsv', sep = '\t')
aha_df = pd.read_csv('/Users/SeoyeonHong/Desktop/text_mining/topic_modeling/aha/aha_qna_topic_representation.tsv', sep = '\t')
hidoc_df = pd.read_csv('/Users/SeoyeonHong/Desktop/text_mining/topic_modeling/hidoc/hidoc_qna_topic_representation.tsv', sep = '\t')
# +
#topic1_naver = naver_df.loc[naver_df['Topic_Num'] == 2]
topic1_aha_1 = aha_df.loc[aha_df['Topic_Num'] == 23]
#topic1_aha_2 = aha_df.loc[aha_df['Topic_Num'] == 26]
#topic1_aha_3 = aha_df.loc[aha_df['Topic_Num'] == 18]
#topic1_aha = pd.concat([topic1_aha_1, topic1_aha_2])
#topic1_hidoc_1 = hidoc_df.loc[hidoc_df['Topic_Num'] == 9]
#topic1_hidoc_2 = hidoc_df.loc[hidoc_df['Topic_Num'] == 15]
#topic1_hidoc = pd.concat([topic1_hidoc_1, topic1_hidoc_2])
#extra = ["안녕하세요 의사선생님 궁금한게 있어서 질문드립니다 할머니가 지금 폐렴에 걸리셨습니다 근데 기침도 안하시고 하는데 폐렴이랍니다 ㅠㅠ 근데 어머니가 70세이상은 보호자가 꼭 필요하다면서 오늘 할머니 옆에서 주무시기로 했습니다 1인실 특실입니다 창문도 입고요 ㅠㅠ 저는 걱정되는게 폐렴이 전염성이 있어 롬길까봐 걱정입니다 창문도 있고 다 있지만요 폐렴 전염성이 높은건가요? 할머니가 이상하게 기침은 안하신다는데 그리고 할머니는 마스크를 착용 하지 않고 있습니다 어머니는 KF94 마스크 착용하고 계시구요 만약에 오늘 어머니가 그 병실에 자면은 폐렴에 전염 되지 않나요? 어머니는 K94 마스크를 낀 채로 주무신다고 합니다 더 걱정되는건 1인실 창문은 있지만 밀폐된 공간이라 걱정이 많이 되는네요 ㅠㅠ 지금 이 시국에 너무 걱정입니다 폐렴 전염성이 높을까요? 2일뒤면 저도 가야되는데 걱정되어 죽겠습니다 좀 도와주세요", "아파트 인터폰 현관앞에 사람들 많이 지나가는 길에 핸드폰이랑 이어폰을 떨어뜨렸고 그걸 맨손으로 주웠습니다.근데 땅바닥은 사람들이 신발로 밟고 지나가잖아요그럼 침이나 가래같은 걸 밟고 땅에도 묻어있을텐데 거기에 핸드폰을 떨어뜨렸으니 바이러스가 혹시라도 묻어서 그걸 집거나 공기중으로 전파되어 제가 감염될수 있을지 너무 불안합니다..확진자인진 모르지만 사람들이 여러군데 다니고 하잖아요 그럼 그중에 확진자나 잠복기에 있는 사람의 가래나 침이 땅바닥에 묻어있거나 그걸 밟은 신발로 다니니 땅바닥에도 우한폐렴 바이러스가 있을거 같은데요지금은 물티슈로 닦았는데 안심해도 되나요 아님 우한폐렴 걸릴수도 있나요?근데 이미 닦았다고 해도 공기중으로 감염되면 어떡하죠?", "아는사람이 신종플루라는데 타미플루를 처방받고 5일이 지났는데도 잔증상(기침같은)이 남아 계속 병원을 다닙니다2주째 계속되고 있는데 본인은 많이 나아졌다고 하는데 저는 불안합니다이사람이랑 접촉해도 안 옮을까요?", "안녕하세요~요즘 코로나 바이러스 때문에 걱정이 많은데요 카페에서 일을하니 주문을 받기 때문에 여러사람과 대화를 하는데 중국사람들 당골도 많습니다 오늘은 평소에 자주오는 중국 여자 손님과 가까이서 얘기를 했는데 손님은 마스크를 쓰지 않았고 저는 쓰고 몇분얘기했는데 손님이 가고난후에 아차하는 생각과 걱정이 되네요 중국인이라는 생각에 더걱정이 되네요 이런일로 걱정을 해야하는 현실이 너무 싫고 요즘 너무 우울합니다 그리고 사람들과 주문을 받는 과정에서 대화를 해야하는데 어떻게 해야할지요"]
# -
topic1_aha_1.info()
# +
import re
from konlpy.tag import Mecab
from konlpy.tag import Kkma
kkma = Kkma()
mecab = Mecab()
mecab_nouns = []
for sent in topic1_aha['Text']: #21
substring = re.sub(r'[^\w\s]','',str(sent))
substring = ''.join([i for i in str(substring) if not i.isdigit()])
substring = str(substring).replace("가능","").replace("정도","").replace("관련","").replace("지금","").replace("월일","").replace("가요","").replace("동안","").replace("요즘","").replace("평소","").replace("최근","").replace("느낌","").replace("하루","").replace("시간","").replace("오늘","").replace("동안","").replace("새벽","").replace("그때","").replace("예전","").replace("코로나", "").replace("면", "").replace("도", "").replace("은", "").replace("임", "").replace("글", "").replace("감사", "").replace("시", "").replace("때", "").replace("곳", "").replace("문", "").replace("말", "").replace("코로나바이러스", "")
sent_pos = mecab.pos(substring)
nouns = [n for n, tag in sent_pos if tag in ["NNG","NNP"] ]
#print(nouns)
mecab_nouns.append(nouns)
for sent in topic1_hidoc['Text']: #
substring = re.sub(r'[^\w\s]','',str(sent))
substring = ''.join([i for i in str(substring) if not i.isdigit()])
substring = str(substring).replace("가능","").replace("정도","").replace("관련","").replace("지금","").replace("월일","").replace("가요","").replace("동안","").replace("요즘","").replace("평소","").replace("최근","").replace("느낌","").replace("하루","").replace("시간","").replace("오늘","").replace("동안","").replace("새벽","").replace("그때","").replace("예전","").replace("전","").replace("후","").replace("닦","").replace("답변","").replace("안녕","").replace("제목","").replace("도","").replace("나용","").replace("번","").replace("애요","").replace("쌀","").replace("정","").replace("질문","").replace("고","").replace("때","").replace("첨","").replace("칸","").replace("소간","").replace("일","").replace("의","").replace("상","").replace("일","").replace("코로나","").replace("대요","").replace("자","").replace("글","").replace("시", "").replace("코로나바이러스","").replace("문","").replace("달","")
sent_pos = mecab.pos(substring)
nouns = [n for n, tag in sent_pos if tag in ["NNG","NNP"] ]
#print(nouns)
mecab_nouns.append(nouns)
for sent in topic1_naver['Text']: #1701
substring = re.sub(r'[^\w\s]','',str(sent))
substring = ''.join([i for i in str(substring) if not i.isdigit()])
substring = str(substring).replace("가능","").replace("정도","").replace("관련","").replace("지금","").replace("월일","").replace("가요","").replace("동안","").replace("요즘","").replace("평소","").replace("최근","").replace("느낌","").replace("하루","").replace("시간","").replace("오늘","").replace("동안","").replace("새벽","").replace("그때","").replace("예전","").replace("▲","").replace("◇","").replace("-","").replace("코로나", "").replace("코로나바이러스","").replace("내공","").replace("질문","").replace("답변","").replace("안녕하세요","")
sent_pos = mecab.pos(substring)
nouns = [n for n, tag in sent_pos if tag in ["NNG","NNP"] ]
#print(nouns)
mecab_nouns.append(nouns)
#for sent in extra:
# substring = re.sub(r'[^\w\s]','',str(sent))
# substring = ''.join([i for i in str(substring) if not i.isdigit()])
# substring = str(substring).replace("전","").replace("후","").replace("닦","").replace("답변","").replace("안녕","").replace("제목","").replace("도","").replace("나용","").replace("번","").replace("애요","").replace("쌀","").replace("정","").replace("질문","").replace("고","").replace("때","").replace("첨","").replace("칸","").replace("소간","").replace("일","").replace("의","").replace("상","").replace("일","").replace("코로나","").replace("대요","").replace("자","").replace("글","").replace("시", "").replace("코로나바이러스","").replace("문","").replace("달","")
# sent_pos = mecab.pos(substring)
# nouns = [n for n, tag in sent_pos if tag in ["NNG","NNP"] ]
# #print(nouns)
# mecab_nouns.append(nouns)
# -
len(mecab_nouns)
import scipy.sparse as sp
from sklearn.feature_extraction.text import CountVectorizer
class Cooccurrence(CountVectorizer):
"""Co-ocurrence matrix
Convert collection of raw documents to word-word co-ocurrence matrix
Parameters
----------
encoding : string, 'utf-8' by default.
If bytes or files are given to analyze, this encoding is used to
decode.
ngram_range : tuple (min_n, max_n)
The lower and upper boundary of the range of n-values for different
n-grams to be extracted. All values of n such that min_n <= n <= max_n
will be used.
max_df: float in range [0, 1] or int, default=1.0
min_df: float in range [0, 1] or int, default=1
Example
-------
>> import Cooccurrence
>> docs = ['this book is good',
'this cat is good',
'cat is good shit']
>> model = Cooccurrence()
>> Xc = model.fit_transform(docs)
Check vocabulary by printing
>> model.vocabulary_
"""
def __init__(self, encoding='utf-8', ngram_range=(1, 1),
max_df=1.0, min_df=1, max_features=None,
stop_words=None, normalize=True, vocabulary=None):
super(Cooccurrence, self).__init__(
ngram_range=ngram_range,
max_df=max_df,
min_df=min_df,
max_features=max_features,
stop_words=stop_words,
vocabulary=vocabulary
)
self.X = None
self.normalize = normalize
def fit_transform(self, raw_documents, y=None):
"""Fit cooccurrence matrix
Parameters
----------
raw_documents : iterable
an iterable which yields either str, unicode or file objects
Returns
-------
Xc : Cooccurrence matrix
"""
X = super(Cooccurrence, self).fit_transform(raw_documents)
self.X = X
n_samples, n_features = X.shape
Xc = (X.T * X)
if self.normalize:
g = sp.diags(1./Xc.diagonal())
Xc = g * Xc
else:
Xc.setdiag(0)
return Xc
def vocab(self):
tuples = super(Cooccurrence, self).get_feature_names()
vocabulary=[]
for e_tuple in tuples:
tokens = e_tuple.split()
for t in tokens:
if t not in vocabulary:
vocabulary.append(t)
return vocabulary
def word_histgram(self):
word_list = super(Cooccurrence, self).get_feature_names()
count_list = self.X.toarray().sum(axis=0)
return dict(zip(word_list,count_list))
# +
from collections import Counter
from nltk import bigrams
from collections import defaultdict
import operator
import numpy as np
class BaseCooccurrence:
INPUT=[list,str]
OUTPUT=[list,tuple]
class CooccurrenceWorker(BaseCooccurrence):
def __init__(self):
name = 'cooccurrence'
self.inst = Cooccurrence(ngram_range=(2, 2), stop_words='english')
def __call__(self, *args, **kwargs):
# bigram_vectorizer = CountVectorizer(ngram_range=(1, 2), vocabulary={'awesome unicorns': 0, 'batman forever': 1})
co_occurrences = self.inst.fit_transform(args[0])
# print('Printing sparse matrix:', co_occurrences)
# print(co_occurrences.todense())
sum_occ = np.sum(co_occurrences.todense(), axis=0)
# print('Sum of word-word occurrences:', sum_occ)
# Converting itertor to set
result = zip(self.inst.get_feature_names(), np.array(sum_occ)[0].tolist())
result_set = list(result)
return result_set, self.inst.vocab()
class CooccurrenceManager:
def computeCooccurence(self, list):
com = defaultdict(lambda: defaultdict(int))
count_all = Counter()
count_all1 = Counter()
uniqueList = []
for _array in list:
for line in _array:
for word in line:
if word not in uniqueList:
uniqueList.append(word)
terms_bigram = bigrams(line)
# Update the counter
count_all.update(line)
count_all1.update(terms_bigram)
# Build co-occurrence matrix
for i in range(len(line) - 1):
for j in range(i + 1, len(line)):
w1, w2 = sorted([line[i], line[j]])
if w1 != w2:
com[w1][w2] += 1
com_max = []
# For each term, look for the most common co-occurrent terms
for t1 in com:
t1_max_terms = sorted(com[t1].items(), key=operator.itemgetter(1), reverse=True)[:5]
for t2, t2_count in t1_max_terms:
com_max.append(((t1, t2), t2_count))
# Get the most frequent co-occurrences
terms_max = sorted(com_max, key=operator.itemgetter(1), reverse=True)
return terms_max, uniqueList
# -
co = CooccurrenceWorker()
documents = []
for sublist in mecab_nouns:
document = ",".join(sublist)
documents.append(document)
#import itertools
#merged = list(itertools.chain(*mecab_nouns))
co_result, vocab = co.__call__(documents)
# +
import networkx as nx
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.font_manager as fm
import platform
from matplotlib.ft2font import FT2Font
import matplotlib as mpl
class GraphMLCreator:
def __init__(self):
self.G = nx.Graph()
# Hack: offset the most central node to avoid too much overlap
self.rad0 = 0.3
def createGraphML(self, co_occurrence, word_hist, vocabulary, file):
G = nx.Graph()
for obj in vocabulary:
G.add_node(obj)
# convert list to a single dictionary
for pair in co_occurrence:
node1 = ''
node2 = ''
for inner_pair in pair:
if type(inner_pair) is tuple:
node1 = inner_pair[0]
node2 = inner_pair[1]
elif type(inner_pair) is str:
inner_pair=inner_pair.split()
node1 = inner_pair[0]
node2 = inner_pair[1]
elif type(inner_pair) is int:
#print ("X " + node1 + " == " + node2 + " == " + str(inner_pair) + " : " + str(tuple[node1]))
G.add_edge(node1, node2, weight=float(inner_pair))
elif type(inner_pair) is float:
#print ("X " + node1 + " == " + node2 + " == " + str(inner_pair) + " : ")
G.add_edge(node1, node2, weight=float(inner_pair))
for word in word_hist:
G.add_node(word, count=word_hist[word])
self.G = G
print(self.G.number_of_nodes())
nx.write_graphml(G, file)
def createGraphMLWithThreshold(self, co_occurrence, word_hist, vocab, file, threshold=10.0):
G = nx.Graph()
filtered_word_list=[]
for pair in co_occurrence:
node1 = ''
node2 = ''
for inner_pair in pair:
if type(inner_pair) is tuple:
node1 = inner_pair[0]
node2 = inner_pair[1]
elif type(inner_pair) is str:
inner_pair=inner_pair.split()
node1 = inner_pair[0]
node2 = inner_pair[1]
elif type(inner_pair) is int:
if float(inner_pair) >= threshold:
#print ("X " + node1 + " == " + node2 + " == " + str(inner_pair) + " : " + str(tuple[node1]))
G.add_edge(node1, node2, weight=float(inner_pair))
if node1 not in filtered_word_list:
filtered_word_list.append(node1)
if node2 not in filtered_word_list:
filtered_word_list.append(node2)
elif type(inner_pair) is float:
if float(inner_pair) >= threshold:
#print ("X " + node1 + " == " + node2 + " == " + str(inner_pair) + " : ")
G.add_edge(node1, node2, weight=float(inner_pair))
if node1 not in filtered_word_list:
filtered_word_list.append(node1)
if node2 not in filtered_word_list:
filtered_word_list.append(node2)
for word in word_hist:
if word in filtered_word_list:
G.add_node(word, count=word_hist[word])
self.G = G
print(self.G.number_of_nodes())
nx.write_graphml(G, file)
def centrality_layout(self):
centrality = nx.eigenvector_centrality_numpy(self.G)
"""Compute a layout based on centrality.
"""
# Create a list of centralities, sorted by centrality value
cent = sorted(centrality.items(), key=lambda x:float(x[1]), reverse=True)
nodes = [c[0] for c in cent]
cent = np.array([float(c[1]) for c in cent])
rad = (cent - cent[0])/(cent[-1]-cent[0])
rad = self.rescale_arr(rad, self.rad0, 1)
angles = np.linspace(0, 2*np.pi, len(centrality))
layout = {}
for n, node in enumerate(nodes):
r = rad[n]
th = angles[n]
layout[node] = r*np.cos(th), r*np.sin(th)
return layout
def plot_graph(self, title=None, file='graph.png'):
from matplotlib.font_manager import _rebuild
_rebuild()
font_path = '/System/Library/Fonts/Supplemental/AppleGothic.ttf'
font_name = fm.FontProperties(fname=font_path).get_name()
plt.rc('font', family=font_name)
plt.rc('axes', unicode_minus=False)
# 그래프에서 마이너스 폰트 깨지는 문제에 대한 대처
mpl.rcParams['axes.unicode_minus'] = False
#print('버전: ', mpl.__version__)
#print('설치 위치: ', mpl.__file__)
#print('설정 위치: ', mpl.get_configdir())
#print('캐시 위치: ', mpl.get_cachedir())
# size, family
print('# 설정 되어있는 폰트 사이즈')
print(plt.rcParams['font.size'])
print('# 설정 되어있는 폰트 글꼴')
print(plt.rcParams['font.family'])
fig = plt.figure(figsize=(8, 8))
pos = self.centrality_layout()
"""Conveniently summarize graph visually"""
# config parameters
edge_min_width= 3
edge_max_width= 12
label_font = 18
node_font = 22
node_alpha = 0.4
edge_alpha = 0.55
edge_cmap = plt.cm.Spectral
# Create figure
if fig is None:
fig, ax = plt.subplots()
else:
ax = fig.add_subplot(111)
fig.subplots_adjust(0,0,1)
font = FT2Font(font_path)
# Plot nodes with size according to count
sizes = []
degrees = []
for n, d in self.G.nodes(data=True):
sizes.append(d['count'])
degrees.append(self.G.degree(n))
sizes = self.rescale_arr(np.array(sizes, dtype=float), 100, 1000)
# Compute layout and label edges according to weight
pos = nx.spectral_layout(self.G) if pos is None else pos
labels = {}
width = []
for n1, n2, d in self.G.edges(data=True):
w = d['weight']
labels[n1, n2] = w
width.append(w)
width = self.rescale_arr(np.array(width, dtype=float), edge_min_width,
edge_max_width)
# Draw
nx.draw_networkx_nodes(self.G, pos, node_size=sizes, node_color=degrees,
alpha=node_alpha)
nx.draw_networkx_edges(self.G, pos, width=width, edge_color=width,
edge_cmap=edge_cmap, alpha=edge_alpha)
#nx.draw_networkx_edge_labels(self.G, pos, edge_labels=labels,
#font_size=label_font)
nx.draw_networkx_labels(self.G, pos, font_size=node_font, font_family=font_name, font_weight='bold')
if title is not None:
ax.set_title(title, fontsize=label_font)
ax.set_xticks([])
ax.set_yticks([])
# Mark centrality axes
kw = dict(color='k', linestyle='-')
cross = [ax.axhline(0, **kw), ax.axvline(self.rad0, **kw)]
[l.set_zorder(0) for l in cross]
plt.savefig(file)
plt.show()
def rescale_arr(self, arr, amin, amax):
"""Rescale an array to a new range.
Return a new array whose range of values is (amin, amax).
Parameters
----------
arr : array-like
amin : float
new minimum value
amax : float
new maximum value
Examples
--------
>>> a = np.arange(5)
>>> rescale_arr(a,3,6)
array([ 3. , 3.75, 4.5 , 5.25, 6. ])
"""
# old bounds
m = arr.min()
M = arr.max()
# scale/offset
s = float(amax - amin) / (M - m)
d = amin - s * m
# Apply clip before returning to cut off possible overflows outside the
# intended range due to roundoff error, so that we can absolutely guarantee
# that on output, there are no values > amax or < amin.
return np.clip(s * arr + d, amin, amax)
def summarize_centrality(self, limit=10):
centrality = nx.eigenvector_centrality_numpy(self.G)
c = centrality.items()
c = sorted(c, key=lambda x: x[1], reverse=True)
print('\nGraph centrality')
count=0
for node, cent in c:
if count>limit:
break
print ("%15s: %.3g" % (node, float(cent)))
count+=1
def sort_freqs(self, freqs):
"""Sort a word frequency histogram represented as a dictionary.
Parameters
----------
freqs : dict
A dict with string keys and integer values.
Return
------
items : list
A list of (count, word) pairs.
"""
items = freqs.items()
items.sort(key=lambda wc: wc[1])
return items
def plot_word_histogram(self, freqs, show=10, title=None):
"""Plot a histogram of word frequencies, limited to the top `show` ones.
"""
sorted_f = self.sort_freqs(freqs) if isinstance(freqs, dict) else freqs
# Don't show the tail
if isinstance(show, int):
# interpret as number of words to show in histogram
show_f = sorted_f[-show:]
else:
# interpret as a fraction
start = -int(round(show * len(freqs)))
show_f = sorted_f[start:]
# Now, extract words and counts, plot
n_words = len(show_f)
ind = np.arange(n_words)
words = [i[0] for i in show_f]
counts = [i[1] for i in show_f]
fig = plt.figure()
ax = fig.add_subplot(111)
if n_words <= 20:
# Only show bars and x labels for small histograms, they don't make
# sense otherwise
ax.bar(ind, counts)
ax.set_xticks(ind)
ax.set_xticklabels(words, rotation=45)
fig.subplots_adjust(bottom=0.25)
else:
# For larger ones, do a step plot
ax.step(ind, counts)
# If it spans more than two decades, use a log scale
if float(max(counts)) / min(counts) > 100:
ax.set_yscale('log')
if title:
ax.set_title(title)
return ax
# -
cv = CountVectorizer()
cv_fit = cv.fit_transform(documents)
word_list = cv.get_feature_names();
count_list = cv_fit.toarray().sum(axis=0)
word_hist = dict(zip(word_list, count_list))
graph_builder = GraphMLCreator()
graph_builder.createGraphMLWithThreshold(co_result, word_hist, vocab, file="/Users/SeoyeonHong/Desktop/text_mining/qna_topic_preventive.graphml", threshold=30.0)
| topic_depth_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generating animated time series for the crop-mask <img align="right" src="../Supplementary_data/DE_Africa_Logo_Stacked_RGB_small.jpg">
#
# ## Description
# This notebook:
#
# 1. Import a time series of cloud-free satellite imagery
# 2. Calculate annual geomedians
# 3. Open cropmask at the same location
# 4. Vectorize crop-mask for plotting over the animation
# 5. Create an animation
#
# ***
# ## Getting started
#
# > Note: currently set up for running Landsat Collection 2 Provisional so requires setting `aws_unsigned=False` in the dask cluster creation and running it on a C2 sandbox
# ### Load packages
# +
# %matplotlib inline
import sys
import datacube
import skimage.exposure
import geopandas as gpd
import matplotlib.pyplot as plt
from IPython.display import Image
import odc.algo
import xarray as xr
from datacube.utils import masking
from odc.algo import xr_geomedian, int_geomedian
from odc.algo import xr_reproject
sys.path.append('../Scripts')
from deafrica_plotting import xr_animation, rgb, display_map
from deafrica_datahandling import load_ard
from deafrica_dask import create_local_dask_cluster
from deafrica_spatialtools import xr_vectorize, xr_rasterize
# -
create_local_dask_cluster(aws_unsigned=False)
# ### Connect to the datacube
dc = datacube.Datacube(app='Animated_timeseries')
# ## Load satellite data from datacube
#
# +
lat, lon = -8.105, 33.294 #-6.72, 32.214 #tanzania -6.7519, 32.4694
lon_buffer = 0.3
lat_buffer = 0.15
x = (lon-lon_buffer, lon+lon_buffer)
y = (lat-lat_buffer, lat+lat_buffer)
# Create a reusable query
query = {
'x': x,
'y': y,
'time': ('2013','2018'),
'resolution': (-30, 30)
}
# -
display_map(x=x, y=y)
# +
# Load available data
ds = load_ard(dc=dc,
products=['usgs_ls8c_level2_2'],
measurements=['red', 'green', 'blue'], #'nir_1', 'swir_1', 'swir_2'
group_by='solar_day',
output_crs='epsg:6933',
dask_chunks={'x':1000, 'y':1000},
**query)
print(ds)
# -
# ## Annual geomedians
grouped = ds.resample(time='1YS')
gms=grouped.map(xr_geomedian)
gms=gms.compute()
# ## Open crop-mask at same location
#
# reproject to match Landsat
g_id='C-20'
# +
p=xr.open_rasterio('../eastern_cropmask/results/classifications/predicted/20201215/Eastern_tile_'+g_id+'_prediction_pixel_gm_mads_two_seasons_20201215.tif')
p=p.squeeze().sel(x=slice(ds.coords['x'][0].values, ds.coords['x'][-1].values), y=slice(ds.coords['y'][0].values, ds.coords['y'][-1].values))
p=xr_reproject(p, gms.geobox, "nearest")
print(p)
# -
# ## Vectorize for plotting over satellite image
# +
gdf = xr_vectorize(p,
crs=ds.crs,
transform=p.geobox.transform,
mask=p.values==1)
gdf['geometry'] = gdf['geometry'].buffer(0)
gdf['DISS_ID'] = 1
gdf = gdf.dissolve(by='DISS_ID', aggfunc='sum')
# -
# ## Create animation
# +
import numpy as np
import pandas as pd
recent = xr.concat([gms.isel(time=-1), gms.isel(time=-1)], dim='time')
time=pd.to_datetime(np.array(['2018-01-01', '2019-01-01']))
recent['time'] = time
# +
gdf['start_time'] = ['2019-01']
gdf['end_time'] = ['2019-01']
# Get shapefile path
gdf['color'] = '#3266ff90'
# Produce time series animation
xr_animation(ds=recent,
bands=['red', 'green', 'blue'],
output_path='data/animations/tanzania_cropping.gif',
interval=1000,
width_pixels=400,
show_date = None,
show_text = ['2019 True Colour','2019 Crop Mask'],
gdf_kwargs={'edgecolor': 'blue', 'linewidth':0.2},
show_gdf=gdf)
# Plot animated gif
plt.close()
Image(filename='data/animations/tanzania_cropping.gif')
# -
| pre-post_processing/animations_cropmask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from copy import deepcopy
import numpy as np
import torch
from torch.optim import Adam
import pybulletgym
import gym
import time
import spinup.algos.pytorch.lstm_ddpg.core as core
from spinup.utils.logx import EpochLogger
import itertools
import torch
import torch.nn as nn
from torch.nn.utils.rnn import pad_sequence, pack_padded_sequence, pad_packed_sequence
class POMDPWrapper(gym.ObservationWrapper):
def __init__(self, env_name):
super().__init__(gym.make(env_name))
# Remove velocity info
# OpenAIGym
# 1. MuJoCo
if env_name == "HalfCheetah-v3" or env_name == "HalfCheetah-v2":
self.remain_obs_idx = np.arange(0, 8)
elif env_name == "Ant-v3" or env_name == "Ant-v2":
self.remain_obs_idx = list(np.arange(0, 13)) + list(np.arange(27, 111))
elif env_name == 'Walker2d-v3' or env_name == "Walker2d-v2":
self.remain_obs_idx = np.arange(0, 8)
elif env_name == 'Hopper-v3' or env_name == "Hopper-v2":
self.remain_obs_idx = np.arange(0, 5)
elif env_name == "InvertedPendulum-v2":
self.remain_obs_idx = np.arange(0, 2)
elif env_name == "InvertedDoublePendulum-v2":
self.remain_obs_idx = list(np.arange(0, 5)) + list(np.arange(8, 11))
elif env_name == "Swimmer-v3" or env_name == "Swimmer-v2":
self.remain_obs_idx = np.arange(0, 3)
elif env_name == "Thrower-v2":
self.remain_obs_idx = list(np.arange(0, 7)) + list(np.arange(14, 23))
elif env_name == "Striker-v2":
self.remain_obs_idx = list(np.arange(0, 7)) + list(np.arange(14, 23))
elif env_name == "Pusher-v2":
self.remain_obs_idx = list(np.arange(0, 7)) + list(np.arange(14, 23))
elif env_name == "Reacher-v2":
self.remain_obs_idx = list(np.arange(0, 6)) + list(np.arange(8, 11))
elif env_name == 'Humanoid-v3' or env_name == "Humanoid-v2":
self.remain_obs_idx = list(np.arange(0, 22)) + list(np.arange(45, 185)) + list(np.arange(269, 376))
elif env_name == 'HumanoidStandup-v2':
self.remain_obs_idx = list(np.arange(0, 22)) + list(np.arange(45, 185)) + list(np.arange(269, 376))
# PyBulletGym
# 1. MuJoCo
elif env_name == 'HalfCheetahMuJoCoEnv-v0':
self.remain_obs_idx = np.arange(0, 8)
elif env_name == 'AntMuJoCoEnv-v0':
self.remain_obs_idx = list(np.arange(0, 13)) + list(np.arange(27, 111))
elif env_name == 'Walker2DMuJoCoEnv-v0':
self.remain_obs_idx = np.arange(0, 8)
elif env_name == 'HopperMuJoCoEnv-v0':
self.remain_obs_idx = np.arange(0, 7)
elif env_name == 'InvertedPendulumMuJoCoEnv-v0':
self.remain_obs_idx = np.arange(0, 3)
elif env_name == 'InvertedDoublePendulumMuJoCoEnv-v0':
self.remain_obs_idx = list(np.arange(0, 5)) + list(np.arange(8, 11))
# 2. Roboschool
elif env_name == 'HalfCheetahPyBulletEnv-v0':
self.remain_obs_idx = list(set(np.arange(0,26)) - set(np.arange(3,6)))
elif env_name == 'AntPyBulletEnv-v0':
self.remain_obs_idx = list(set(np.arange(0,28)) - set(np.arange(3,6)))
elif env_name == 'Walker2DPyBulletEnv-v0':
self.remain_obs_idx = list(set(np.arange(0,22)) - set(np.arange(3,6)))
elif env_name == 'HopperPyBulletEnv-v0':
self.remain_obs_idx = list(set(np.arange(0,15)) - set(np.arange(3,6)))
elif env_name == 'InvertedPendulumPyBulletEnv-v0':
self.remain_obs_idx = list(set(np.arange(0,5)) - set([1,4]))
elif env_name == 'InvertedDoublePendulumPyBulletEnv-v0':
self.remain_obs_idx = list(set(np.arange(0,9)) - set([1,5,8]))
elif env_name == 'ReacherPyBulletEnv-v0':
self.remain_obs_idx = list(set(np.arange(0,9)) - set([6,8]))
else:
raise ValueError('POMDP for {} is not defined!'.format(env_name))
# Redefine observation_space
obs_low = np.array([-np.inf for i in range(len(self.remain_obs_idx))], dtype="float32")
obs_high = np.array([np.inf for i in range(len(self.remain_obs_idx))], dtype="float32")
self.observation_space = gym.spaces.Box(obs_low, obs_high)
def observation(self, obs):
return obs.flatten()[self.remain_obs_idx]
class ReplayBuffer:
"""
A simple FIFO experience replay buffer for agents.
"""
def __init__(self, obs_dim, act_dim, max_size):
self.obs_dim = obs_dim
self.act_dim = act_dim
self.max_size = max_size
self.obs_buf = np.zeros(core.combined_shape(max_size, obs_dim), dtype=np.float32)
self.obs2_buf = np.zeros(core.combined_shape(max_size, obs_dim), dtype=np.float32)
self.act_buf = np.zeros(core.combined_shape(max_size, act_dim), dtype=np.float32)
self.rew_buf = np.zeros(max_size, dtype=np.float32)
self.done_buf = np.zeros(max_size, dtype=np.float32)
self.ptr, self.size = 0, 0
def store(self, obs, act, rew, next_obs, done):
self.obs_buf[self.ptr] = obs
self.act_buf[self.ptr] = act
self.rew_buf[self.ptr] = rew
self.obs2_buf[self.ptr] = list(next_obs)
self.done_buf[self.ptr] = done
self.ptr = (self.ptr+1) % self.max_size
self.size = min(self.size+1, self.max_size)
def sample_batch(self, batch_size=32):
idxs = np.random.randint(0, self.size, size=batch_size)
batch = dict(obs=self.obs_buf[idxs],
obs2=self.obs2_buf[idxs],
act=self.act_buf[idxs],
rew=self.rew_buf[idxs],
done=self.done_buf[idxs])
return {k: torch.as_tensor(v, dtype=torch.float32) for k,v in batch.items()}
def sample_batch_with_history(self, batch_size=32, max_hist_len=100):
idxs = np.random.randint(max_hist_len, self.size, size=batch_size)
# History
if max_hist_len == 0:
hist_obs = np.zeros([batch_size, 1, self.obs_dim])
hist_act = np.zeros([batch_size, 1, self.act_dim])
hist_obs2 = np.zeros([batch_size, 1, self.obs_dim])
hist_act2 = np.zeros([batch_size, 1, self.act_dim])
hist_rew = np.zeros([batch_size, 1])
hist_done = np.zeros([batch_size, 1])
hist_len = np.zeros(batch_size)
# hist_msk = np.tile((hist_len!=0).astype(float).reshape([-1,1]), [1, 12]).shape
else:
hist_obs = np.zeros([batch_size, max_hist_len, self.obs_dim])
hist_act = np.zeros([batch_size, max_hist_len, self.act_dim])
hist_obs2 = np.zeros([batch_size, max_hist_len, self.obs_dim])
hist_act2 = np.zeros([batch_size, max_hist_len, self.act_dim])
hist_rew = np.zeros([batch_size, max_hist_len])
hist_done = np.zeros([batch_size, max_hist_len])
hist_len = max_hist_len * np.ones(batch_size)
for hist_i in range(max_hist_len):
hist_obs[:, -1-hist_i, :] = self.obs_buf[idxs-hist_i-1, :]
hist_act[:, -1-hist_i, :] = self.act_buf[idxs-hist_i-1, :]
hist_obs2[:, -1-hist_i, :] = self.obs2_buf[idxs-hist_i-1, :]
hist_act2[:, -1-hist_i, :] = self.act_buf[idxs-hist_i, :] # include a_t
hist_rew[:, -1-hist_i] = self.rew_buf[idxs-hist_i-1]
hist_done[:, -1-hist_i] = self.done_buf[idxs-hist_i-1]
# If there is done in the backward experiences, only consider the experiences after the last done.
for batch_i in range(batch_size):
done_idxs_exclude_last_exp = np.where(hist_done[batch_i][:-1] == 1) # Exclude last experience
# If exist done
if done_idxs_exclude_last_exp[0].size != 0:
largest_done_id = done_idxs_exclude_last_exp[0][-1]
hist_len[batch_i] = max_hist_len - (largest_done_id+1)
# Only keep experiences after the last done
obs_keep_part = np.copy(hist_obs[batch_i, largest_done_id+1:, :])
act_keep_part = np.copy(hist_act[batch_i, largest_done_id+1:, :])
obs2_keep_part = np.copy(hist_obs2[batch_i, largest_done_id+1:, :])
act2_keep_part = np.copy(hist_act2[batch_i, largest_done_id+1:, :])
rew_keep_part = np.copy(hist_rew[batch_i, largest_done_id+1:])
done_keep_part = np.copy(hist_done[batch_i, largest_done_id+1:])
# Set to 0 to make sure all experiences are at the beginning
hist_obs[batch_i] = np.zeros([max_hist_len, self.obs_dim])
hist_act[batch_i] = np.zeros([max_hist_len, self.act_dim])
hist_obs2[batch_i] = np.zeros([max_hist_len, self.obs_dim])
hist_act2[batch_i] = np.zeros([max_hist_len, self.act_dim])
hist_rew[batch_i] = np.zeros([max_hist_len])
hist_done[batch_i] = np.zeros([max_hist_len])
# Move kept experiences to the start of the segment
hist_obs[batch_i, :max_hist_len-(largest_done_id+1), :] = obs_keep_part
hist_act[batch_i, :max_hist_len-(largest_done_id+1), :] = act_keep_part
hist_obs2[batch_i, :max_hist_len-(largest_done_id+1), :] = obs2_keep_part
hist_act2[batch_i, :max_hist_len-(largest_done_id+1), :] = act2_keep_part
hist_rew[batch_i, :max_hist_len-(largest_done_id+1)] = rew_keep_part
hist_done[batch_i, :max_hist_len-(largest_done_id+1)] = done_keep_part
#
batch = dict(obs=self.obs_buf[idxs],
obs2=self.obs2_buf[idxs],
act=self.act_buf[idxs],
rew=self.rew_buf[idxs],
done=self.done_buf[idxs],
hist_obs=hist_obs,
hist_act=hist_act,
hist_obs2=hist_obs2,
hist_act2=hist_act2,
hist_rew=hist_rew,
hist_done=hist_done,
hist_len=hist_len)
return {k: torch.as_tensor(v, dtype=torch.float32) for k,v in batch.items()}
# +
class MLPCritic(nn.Module):
def __init__(self, obs_dim, act_dim,
mem_pre_lstm_hid_sizes=(128,),
mem_lstm_hid_sizes=(128,),
cur_feature_hid_sizes=(128,),
post_comb_hid_sizes=(128,), mem_gate=True):
super(MLPCritic, self).__init__()
self.obs_dim = obs_dim
self.act_dim = act_dim
self.mem_gate = mem_gate
#
self.mem_pre_lstm_layers = nn.ModuleList()
self.mem_lstm_layers = nn.ModuleList()
self.mem_gate_layer = nn.ModuleList()
self.cur_feature_layers = nn.ModuleList()
self.post_combined_layers = nn.ModuleList()
# Memory
# Pre-LSTM
mem_pre_lstm_layer_size = [obs_dim+act_dim] + list(mem_pre_lstm_hid_sizes)
for h in range(len(mem_pre_lstm_layer_size)-1):
self.mem_pre_lstm_layers += [nn.Linear(mem_pre_lstm_layer_size[h],
mem_pre_lstm_layer_size[h+1]),
nn.ReLU()]
# LSTM
self.mem_lstm_layer_sizes = [mem_pre_lstm_layer_size[-1]] + list(mem_lstm_hid_sizes)
for h in range(len(self.mem_lstm_layer_sizes)-1):
self.mem_lstm_layers += [nn.LSTM(self.mem_lstm_layer_sizes[h], self.mem_lstm_layer_sizes[h+1], batch_first=True)]
# Memeory Gate
if self.mem_gate:
self.mem_gate_layer += [nn.Linear(self.mem_lstm_layer_sizes[-1]+obs_dim+act_dim, self.mem_lstm_layer_sizes[-1]),
nn.Sigmoid()]
# Current Feature Extraction
cur_feature_layer_size = [obs_dim+act_dim]+list(cur_feature_hid_sizes)
for h in range(len(cur_feature_layer_size)-1):
self.cur_feature_layers += [nn.Linear(cur_feature_layer_size[h], cur_feature_layer_size[h+1]),
nn.ReLU()]
# Post-Combination
post_combined_layer_size = [self.mem_lstm_layer_sizes[-1]+cur_feature_layer_size[-1]]+list(post_comb_hid_sizes) + [1]
for h in range(len(post_combined_layer_size)-2):
self.post_combined_layers += [nn.Linear(post_combined_layer_size[h], post_combined_layer_size[h+1]), nn.ReLU()]
self.post_combined_layers += [nn.Linear(post_combined_layer_size[-2], post_combined_layer_size[-1]), nn.Identity()]
def forward(self, obs, act, hist_obs, hist_act, hist_seg_len):
#
tmp_hist_seg_len = deepcopy(hist_seg_len)
tmp_hist_seg_len[hist_seg_len == 0] = 1
x = torch.cat([hist_obs, hist_act], dim=-1)
# Memory
# Pre-LSTM
for layer in self.mem_pre_lstm_layers:
x = layer(x)
# LSTM
for layer in self.mem_lstm_layers:
x, (lstm_hidden_state, lstm_cell_state) = layer(x)
hist_out = torch.gather(x, 1, (tmp_hist_seg_len-1).view(-1,1).repeat(1, self.mem_lstm_layer_sizes[-1]).unsqueeze(1).long()).squeeze(1)
hist_msk = (hist_seg_len != 0).float().view(-1,1).repeat(1, self.mem_lstm_layer_sizes[-1]).cuda()
# Memory Gate
if self.mem_gate:
memory_gate = torch.cat([hist_out*hist_msk, obs, act], dim=-1)
for layer in self.mem_gate_layer:
memory_gate = layer(memory_gate)
# Current Feature Extraction
x = torch.cat([obs, act], dim=-1)
for layer in self.cur_feature_layers:
x = layer(x)
# Post-Combination
if self.mem_gate:
x = torch.cat([memory_gate*hist_out*hist_msk, x], dim=-1)
else:
x = torch.cat([hist_out*hist_msk, x], dim=-1)
for layer in self.post_combined_layers:
x = layer(x)
return torch.squeeze(x, -1) # Critical to ensure q has right shape.
class MLPActor(nn.Module):
def __init__(self, obs_dim, act_dim, act_limit,
mem_pre_lstm_hid_sizes=(128,),
mem_lstm_hid_sizes=(128,),
cur_feature_hid_sizes=(128,),
post_comb_hid_sizes=(128,), mem_gate=True):
super(MLPActor, self).__init__()
self.obs_dim = obs_dim
self.act_dim = act_dim
self.act_limit = act_limit
self.mem_gate = mem_gate
#
self.mem_pre_lstm_layers = nn.ModuleList()
self.mem_lstm_layers = nn.ModuleList()
self.mem_gate_layer = nn.ModuleList()
self.cur_feature_layers = nn.ModuleList()
self.post_combined_layers = nn.ModuleList()
# Memory
# Pre-LSTM
mem_pre_lstm_layer_size = [obs_dim+act_dim] + list(mem_pre_lstm_hid_sizes)
for h in range(len(mem_pre_lstm_layer_size)-1):
self.mem_pre_lstm_layers += [nn.Linear(mem_pre_lstm_layer_size[h],
mem_pre_lstm_layer_size[h+1]),
nn.ReLU()]
# LSTM
self.mem_lstm_layer_sizes = [mem_pre_lstm_layer_size[-1]] + list(mem_lstm_hid_sizes)
for h in range(len(self.mem_lstm_layer_sizes)-1):
self.mem_lstm_layers += [nn.LSTM(self.mem_lstm_layer_sizes[h], self.mem_lstm_layer_sizes[h+1], batch_first=True)]
# Memeory Gate
if self.mem_gate:
self.mem_gate_layer += [nn.Linear(self.mem_lstm_layer_sizes[-1]+obs_dim, self.mem_lstm_layer_sizes[-1]),
nn.Sigmoid()]
# Current Feature Extraction
cur_feature_layer_size = [obs_dim]+list(cur_feature_hid_sizes)
for h in range(len(cur_feature_layer_size)-1):
self.cur_feature_layers += [nn.Linear(cur_feature_layer_size[h], cur_feature_layer_size[h+1]),
nn.ReLU()]
# Post-Combination
post_combined_layer_size = [self.mem_lstm_layer_sizes[-1]+cur_feature_layer_size[-1]]+list(post_comb_hid_sizes) + [act_dim]
for h in range(len(post_combined_layer_size)-2):
self.post_combined_layers += [nn.Linear(post_combined_layer_size[h], post_combined_layer_size[h+1]), nn.ReLU()]
self.post_combined_layers += [nn.Linear(post_combined_layer_size[-2], post_combined_layer_size[-1]), nn.Tanh()]
def forward(self, obs, hist_obs, hist_act, hist_seg_len):
#
tmp_hist_seg_len = deepcopy(hist_seg_len)
tmp_hist_seg_len[hist_seg_len == 0] = 1
x = torch.cat([hist_obs, hist_act], dim=-1)
# Memory
# Pre-LSTM
for layer in self.mem_pre_lstm_layers:
x = layer(x)
# LSTM
for layer in self.mem_lstm_layers:
x, (lstm_hidden_state, lstm_cell_state) = layer(x)
hist_out = torch.gather(x, 1, (tmp_hist_seg_len-1).view(-1,1).repeat(1, self.mem_lstm_layer_sizes[-1]).unsqueeze(1).long()).squeeze(1)
hist_msk = (hist_seg_len != 0).float().view(-1,1).repeat(1, self.mem_lstm_layer_sizes[-1]).cuda()
# Memory Gate
if self.mem_gate:
memory_gate = torch.cat([hist_out*hist_msk, obs], dim=-1)
for layer in self.mem_gate_layer:
memory_gate = layer(memory_gate)
# Current Feature Extraction
x = obs
for layer in self.cur_feature_layers:
x = layer(x)
# Post-Combination
if self.mem_gate:
x = torch.cat([memory_gate*hist_out*hist_msk, x], dim=-1)
else:
x = torch.cat([hist_out*hist_msk, x], dim=-1)
for layer in self.post_combined_layers:
x = layer(x)
return self.act_limit * x
class MLPActorCritic(nn.Module):
def __init__(self, obs_dim, act_dim, act_limit=1,
critic_mem_pre_lstm_hid_sizes=(128,),
critic_mem_lstm_hid_sizes=(128,),
critic_cur_feature_hid_sizes=(128,),
critic_post_comb_hid_sizes=(128,), critic_mem_gate=True,
actor_mem_pre_lstm_hid_sizes=(128,),
actor_mem_lstm_hid_sizes=(128,),
actor_cur_feature_hid_sizes=(128,),
actor_post_comb_hid_sizes=(128,), actor_mem_gate=True):
super(MLPActorCritic, self).__init__()
self.q1 = MLPCritic(obs_dim, act_dim,
mem_pre_lstm_hid_sizes=critic_mem_pre_lstm_hid_sizes,
mem_lstm_hid_sizes=critic_mem_lstm_hid_sizes,
cur_feature_hid_sizes=critic_cur_feature_hid_sizes,
post_comb_hid_sizes=critic_post_comb_hid_sizes, mem_gate=critic_mem_gate)
self.q2 = MLPCritic(obs_dim, act_dim,
mem_pre_lstm_hid_sizes=critic_mem_pre_lstm_hid_sizes,
mem_lstm_hid_sizes=critic_mem_lstm_hid_sizes,
cur_feature_hid_sizes=critic_cur_feature_hid_sizes,
post_comb_hid_sizes=critic_post_comb_hid_sizes, mem_gate=critic_mem_gate)
self.pi = MLPActor(obs_dim, act_dim, act_limit,
mem_pre_lstm_hid_sizes=actor_mem_pre_lstm_hid_sizes,
mem_lstm_hid_sizes=actor_mem_lstm_hid_sizes,
cur_feature_hid_sizes=actor_cur_feature_hid_sizes,
post_comb_hid_sizes=actor_post_comb_hid_sizes, mem_gate=actor_mem_gate)
def act(self, obs, hist_obs=None, hist_act=None, hist_seg_len=None):
if (hist_obs is None) or (hist_act is None) or (hist_seg_len is None):
hist_obs = torch.zeros(1, 1, self.obs_dim).cuda()
hist_act = torch.zeros(1, 1, self.act_dim).cuda()
hist_seg_len = torch.zeros(1).cuda()
with torch.no_grad():
return self.pi(obs, hist_obs, hist_act, hist_seg_len).cpu().numpy()
# +
def td3(env_name, actor_critic=core.MLPActorCritic, ac_kwargs=dict(), seed=0,
steps_per_epoch=4000, epochs=100, replay_size=int(1e6), gamma=0.99,
polyak=0.995, pi_lr=1e-3, q_lr=1e-3,
batch_size=100, max_hist_len=100,
start_steps=10000,
update_after=1000, update_every=50, act_noise=0.1, target_noise=0.2,
noise_clip=0.5, policy_delay=2, num_test_episodes=10, max_ep_len=1000,
nonstationary_env = True,
gravity_change_pattern = 'gravity_averagely_equal',
partially_observable = False,
freeze_hist_coding = False,
logger_kwargs=dict(), save_freq=1):
"""
Twin Delayed Deep Deterministic Policy Gradient (TD3)
Args:
env_fn : A function which creates a copy of the environment.
The environment must satisfy the OpenAI Gym API.
actor_critic: The constructor method for a PyTorch Module with an ``act``
method, a ``pi`` module, a ``q1`` module, and a ``q2`` module.
The ``act`` method and ``pi`` module should accept batches of
observations as inputs, and ``q1`` and ``q2`` should accept a batch
of observations and a batch of actions as inputs. When called,
these should return:
=========== ================ ======================================
Call Output Shape Description
=========== ================ ======================================
``act`` (batch, act_dim) | Numpy array of actions for each
| observation.
``pi`` (batch, act_dim) | Tensor containing actions from policy
| given observations.
``q1`` (batch,) | Tensor containing one current estimate
| of Q* for the provided observations
| and actions. (Critical: make sure to
| flatten this!)
``q2`` (batch,) | Tensor containing the other current
| estimate of Q* for the provided observations
| and actions. (Critical: make sure to
| flatten this!)
=========== ================ ======================================
ac_kwargs (dict): Any kwargs appropriate for the ActorCritic object
you provided to TD3.
seed (int): Seed for random number generators.
steps_per_epoch (int): Number of steps of interaction (state-action pairs)
for the agent and the environment in each epoch.
epochs (int): Number of epochs to run and train agent.
replay_size (int): Maximum length of replay buffer.
gamma (float): Discount factor. (Always between 0 and 1.)
polyak (float): Interpolation factor in polyak averaging for target
networks. Target networks are updated towards main networks
according to:
.. math:: \\theta_{\\text{targ}} \\leftarrow
\\rho \\theta_{\\text{targ}} + (1-\\rho) \\theta
where :math:`\\rho` is polyak. (Always between 0 and 1, usually
close to 1.)
pi_lr (float): Learning rate for policy.
q_lr (float): Learning rate for Q-networks.
batch_size (int): Minibatch size for SGD.
start_steps (int): Number of steps for uniform-random action selection,
before running real policy. Helps exploration.
update_after (int): Number of env interactions to collect before
starting to do gradient descent updates. Ensures replay buffer
is full enough for useful updates.
update_every (int): Number of env interactions that should elapse
between gradient descent updates. Note: Regardless of how long
you wait between updates, the ratio of env steps to gradient steps
is locked to 1.
act_noise (float): Stddev for Gaussian exploration noise added to
policy at training time. (At test time, no noise is added.)
target_noise (float): Stddev for smoothing noise added to target
policy.
noise_clip (float): Limit for absolute value of target policy
smoothing noise.
policy_delay (int): Policy will only be updated once every
policy_delay times for each update of the Q-networks.
num_test_episodes (int): Number of episodes to test the deterministic
policy at the end of each epoch.
max_ep_len (int): Maximum length of trajectory / episode / rollout.
logger_kwargs (dict): Keyword args for EpochLogger.
save_freq (int): How often (in terms of gap between epochs) to save
the current policy and value function.
"""
logger = EpochLogger(**logger_kwargs)
logger.save_config(locals())
torch.manual_seed(seed)
np.random.seed(seed)
# Wrapper environment if using POMDP
if partially_observable == True:
env, test_env = POMDPWrapper(env_name), POMDPWrapper(env_name)
else:
env, test_env = gym.make(env_name), gym.make(env_name)
obs_dim = env.observation_space.shape[0]
act_dim = env.action_space.shape[0]
# Action limit for clamping: critically, assumes all dimensions share the same bound!
act_limit = env.action_space.high[0]
# Create actor-critic module and target networks
critic_mem_gate = False
actor_mem_gate = False
ac = MLPActorCritic( obs_dim, act_dim, act_limit,
critic_mem_pre_lstm_hid_sizes=(128,),
critic_mem_lstm_hid_sizes=(128,),
critic_cur_feature_hid_sizes=(128,),
critic_post_comb_hid_sizes=(128,), critic_mem_gate=critic_mem_gate,
actor_mem_pre_lstm_hid_sizes=(128,),
actor_mem_lstm_hid_sizes=(128,),
actor_cur_feature_hid_sizes=(128,),
actor_post_comb_hid_sizes=(128,), actor_mem_gate=actor_mem_gate)
ac_targ = deepcopy(ac)
ac.cuda()
ac_targ.cuda()
# # Freeze hist coding
# if freeze_hist_coding:
# ac.q1.layers[0].requires_grad=False
# ac.q2.layers[0].requires_grad=False
# ac.pi.layers[0].requires_grad=False
# Freeze target networks with respect to optimizers (only update via polyak averaging)
for p in ac_targ.parameters():
p.requires_grad = False
# List of parameters for both Q-networks (save this for convenience)
q_params = itertools.chain(ac.q1.parameters(), ac.q2.parameters())
# Experience buffer
replay_buffer = ReplayBuffer(obs_dim=obs_dim, act_dim=act_dim, max_size=replay_size)
# Count variables (protip: try to get a feel for how different size networks behave!)
var_counts = tuple(core.count_vars(module) for module in [ac.pi, ac.q1, ac.q2])
logger.log('\nNumber of parameters: \t pi: %d, \t q1: %d, \t q2: %d\n'%var_counts)
# Set up function for computing TD3 Q-losses
def compute_loss_q(data):
o, a, r, o2, d = data['obs'], data['act'], data['rew'], data['obs2'], data['done']
h_o, h_a, h_o2, h_a2, h_len = data['hist_obs'], data['hist_act'], data['hist_obs2'], data['hist_act2'], data['hist_len']
q1 = ac.q1(o, a, h_o, h_a, h_len)
q2 = ac.q2(o, a, h_o, h_a, h_len)
# Bellman backup for Q functions
with torch.no_grad():
pi_targ = ac_targ.pi(o2, h_o2, h_a2, h_len)
# Target policy smoothing
epsilon = torch.randn_like(pi_targ) * target_noise
epsilon = torch.clamp(epsilon, -noise_clip, noise_clip)
a2 = pi_targ + epsilon
a2 = torch.clamp(a2, -act_limit, act_limit)
# Target Q-values
q1_pi_targ = ac_targ.q1(o2, a2, h_o2, h_a2, h_len)
q2_pi_targ = ac_targ.q2(o2, a2, h_o2, h_a2, h_len)
q_pi_targ = torch.min(q1_pi_targ, q2_pi_targ)
q_pi_targ = q1_pi_targ
backup = r + gamma * (1 - d) * q_pi_targ
# MSE loss against Bellman backup
loss_q1 = ((q1 - backup)**2).mean()
loss_q2 = ((q2 - backup)**2).mean()
loss_q = loss_q1 + loss_q2
loss_q = loss_q1
# Useful info for logging
loss_info = dict(Q1Vals=q1.detach().cpu().numpy(),
Q2Vals=q2.detach().cpu().numpy())
return loss_q, loss_info
# Set up function for computing TD3 pi loss
def compute_loss_pi(data):
o, h_o, h_a, h_len = data['obs'], data['hist_obs'], data['hist_act'], data['hist_len']
q1_pi = ac.q1(o, ac.pi(o, h_o, h_a, h_len), h_o, h_a, h_len)
return -q1_pi.mean()
# Set up optimizers for policy and q-function
pi_optimizer = Adam(ac.pi.parameters(), lr=pi_lr)
q_optimizer = Adam(q_params, lr=q_lr)
# Set up model saving
logger.setup_pytorch_saver(ac)
def update(data, timer):
# First run one gradient descent step for Q1 and Q2
q_optimizer.zero_grad()
loss_q, loss_info = compute_loss_q(data)
loss_q.backward()
q_optimizer.step()
# Record things
logger.store(LossQ=loss_q.item(), **loss_info)
# Possibly update pi and target networks
if timer % policy_delay == 0:
# # Freeze Q-networks so you don't waste computational effort
# # computing gradients for them during the policy learning step.
# for p in q_params:
# p.requires_grad = False
# Next run one gradient descent step for pi.
pi_optimizer.zero_grad()
loss_pi = compute_loss_pi(data)
loss_pi.backward()
pi_optimizer.step()
# # Unfreeze Q-networks so you can optimize it at next DDPG step.
# for p in q_params:
# p.requires_grad = True
# Record things
logger.store(LossPi=loss_pi.item())
# Finally, update target networks by polyak averaging.
with torch.no_grad():
for p, p_targ in zip(ac.parameters(), ac_targ.parameters()):
# NB: We use an in-place operations "mul_", "add_" to update target
# params, as opposed to "mul" and "add", which would make new tensors.
p_targ.data.mul_(polyak)
p_targ.data.add_((1 - polyak) * p.data)
def get_action(o, o_buff, a_buff, o_buff_len, noise_scale):
h_o = torch.tensor(o_buff).view(1, o_buff.shape[0], o_buff.shape[1]).float().cuda()
h_a = torch.tensor(a_buff).view(1, a_buff.shape[0], a_buff.shape[1]).float().cuda()
h_l = torch.tensor([o_buff_len]).float().cuda()
with torch.no_grad():
a = ac.act(torch.as_tensor(o, dtype=torch.float32).view(1,-1).cuda(),
h_o, h_a, h_l).reshape(act_dim)
a += noise_scale * np.random.randn(act_dim)
if a.shape[0]!=act_dim:
import pdb
pdb.set_trace()
return np.clip(a, -act_limit, act_limit)
def test_agent():
for j in range(num_test_episodes):
o, d, ep_ret, ep_len = test_env.reset(), False, 0, 0
if max_hist_len>0:
o_buff = np.zeros([max_hist_len, obs_dim])
a_buff = np.zeros([max_hist_len, act_dim])
o_buff[0,:] = o
o_buff_len = 0
else:
o_buff = np.zeros([1, obs_dim])
a_buff = np.zeros([1, act_dim])
o_buff_len = 0
while not(d or (ep_len == max_ep_len)):
# Take deterministic actions at test time (noise_scale=0)
a = get_action(o, o_buff, a_buff, o_buff_len, 0)
o2, r, d, _ = test_env.step(a)
ep_ret += r
ep_len += 1
# Add short history
if max_hist_len != 0:
if o_buff_len == max_hist_len:
o_buff[:max_hist_len-1] = o_buff[1:]
a_buff[:max_hist_len-1] = a_buff[1:]
o_buff[max_hist_len-1] = list(o)
a_buff[max_hist_len-1] = list(a)
else:
if a.shape[0]!=act_dim:
import pdb
pdb.set_trace()
o_buff[o_buff_len+1-1] = list(o)
a_buff[o_buff_len+1-1] = list(a)
o_buff_len += 1
o = o2
logger.store(TestEpRet=ep_ret, TestEpLen=ep_len)
# Prepare for interaction with environment
total_steps = steps_per_epoch * epochs
start_time = time.time()
o, ep_ret, ep_len = env.reset(), 0, 0
if max_hist_len>0:
o_buff = np.zeros([max_hist_len, obs_dim])
a_buff = np.zeros([max_hist_len, act_dim])
o_buff[0,:] = o
o_buff_len = 0
else:
o_buff = np.zeros([1, obs_dim])
a_buff = np.zeros([1, act_dim])
o_buff_len = 0
# Main loop: collect experience in env and update/log each epoch
start_time = time.time()
for t in range(total_steps):
if t%200 == 0:
end_time = time.time()
print("t={}, {}s".format(t, end_time-start_time))
start_time = end_time
# Until start_steps have elapsed, randomly sample actions
# from a uniform distribution for better exploration. Afterwards,
# use the learned policy (with some noise, via act_noise).
if t > start_steps:
a = get_action(o, o_buff, a_buff, o_buff_len, act_noise)
else:
a = env.action_space.sample()
if nonstationary_env == True:
gravity_cycle = 1000
gravity_base = -9.81
if gravity_change_pattern == 'gravity_averagely_equal':
gravity = gravity_base * 1 / 2 * (np.cos(2 * np.pi / gravity_cycle * t) + 1) + gravity_base / 2
elif gravity_change_pattern == 'gravity_averagely_easier':
gravity = gravity_base * 1 / 2 * (np.cos(2 * np.pi / gravity_cycle * t) + 1)
elif gravity_change_pattern == 'gravity_averagely_harder':
gravity = gravity_base * 1 / 2 * (-np.cos(2 * np.pi / gravity_cycle * t) + 1) + gravity_base
else:
pass
if 'PyBulletEnv' in env_name:
env.env._p.setGravity(0, 0, gravity)
elif 'Roboschool' in env_name:
pass
else:
env.model.opt.gravity[2] = gravity
# Step the env
o2, r, d, _ = env.step(a)
ep_ret += r
ep_len += 1
# Ignore the "done" signal if it comes from hitting the time
# horizon (that is, when it's an artificial terminal signal
# that isn't based on the agent's state)
d = False if ep_len==max_ep_len else d
# Store experience to replay buffer
replay_buffer.store(o, a, r, o2, d)
# Add short history
if max_hist_len != 0:
if o_buff_len == max_hist_len:
o_buff[:max_hist_len-1] = o_buff[1:]
a_buff[:max_hist_len-1] = a_buff[1:]
o_buff[max_hist_len-1] = list(o)
a_buff[max_hist_len-1] = list(a)
else:
if a.shape[0]!=act_dim:
import pdb
pdb.set_trace()
o_buff[o_buff_len+1-1] = list(o)
a_buff[o_buff_len+1-1] = list(a)
o_buff_len += 1
# Super critical, easy to overlook step: make sure to update
# most recent observation!
o = o2
# End of trajectory handling
if d or (ep_len == max_ep_len):
logger.store(EpRet=ep_ret, EpLen=ep_len)
o, ep_ret, ep_len = env.reset(), 0, 0
if max_hist_len>0:
o_buff = np.zeros([max_hist_len, obs_dim])
a_buff = np.zeros([max_hist_len, act_dim])
o_buff[0,:] = o
o_buff_len = 0
else:
o_buff = np.zeros([1, obs_dim])
a_buff = np.zeros([1, act_dim])
o_buff_len = 0
# Update handling
if t >= update_after and t % update_every == 0:
for j in range(update_every):
batch = replay_buffer.sample_batch_with_history(batch_size, max_hist_len)
batch = {k: v.cuda() for k,v in batch.items()}
update(data=batch, timer=j)
# End of epoch handling
if (t+1) % steps_per_epoch == 0:
epoch = (t+1) // steps_per_epoch
# Save model
if (epoch % save_freq == 0) or (epoch == epochs):
logger.save_state({'env': env}, None)
# Test the performance of the deterministic version of the agent.
test_agent()
# Log info about epoch
logger.log_tabular('Epoch', epoch)
logger.log_tabular('EpRet', with_min_and_max=True)
logger.log_tabular('TestEpRet', with_min_and_max=True)
logger.log_tabular('EpLen', average_only=True)
logger.log_tabular('TestEpLen', average_only=True)
logger.log_tabular('TotalEnvInteracts', t)
logger.log_tabular('Q1Vals', with_min_and_max=True)
logger.log_tabular('Q2Vals', with_min_and_max=True)
logger.log_tabular('LossPi', average_only=True)
logger.log_tabular('LossQ', average_only=True)
logger.log_tabular('Time', time.time()-start_time)
logger.dump_tabular()
# +
args = {'env': 'Ant-v2', 'hid': 256, 'l': 2,
'max_hist_len': 5,
'gamma': 0.99, 'seed': 0, 'epochs': 50,
'nonstationary_env':False,
'gravity_change_pattern': 'gravity_averagely_equal',
'partially_observable': True,
'freeze_hist_coding': False,
'exp_name': 'RTD3_MemGate_POMDP_Ant_PreLSTM1L128_HistMemory5Len_LSTM1L128_HidStateLayer0_1L128_NoFreeze'}
from spinup.utils.run_utils import setup_logger_kwargs
logger_kwargs = setup_logger_kwargs(args['exp_name'], args['seed'])
td3(env_name=args['env'], actor_critic=core.MLPActorCritic,
ac_kwargs=dict(hidden_sizes=[args['hid']]*args['l']),
max_hist_len=args['max_hist_len'],
gamma=args['gamma'], seed=args['seed'], epochs=args['epochs'],
nonstationary_env=args['nonstationary_env'],
gravity_change_pattern=args['gravity_change_pattern'],
partially_observable=args['partially_observable'],
freeze_hist_coding=args['freeze_hist_coding'],
logger_kwargs=logger_kwargs)
# +
args = {'env': 'HalfCheetah-v2', 'hid': 256, 'l': 2,
'max_hist_len': 5,
'gamma': 0.99, 'seed': 0, 'epochs': 50,
'nonstationary_env':False,
'gravity_change_pattern': 'gravity_averagely_equal',
'partially_observable': False,
'freeze_hist_coding': False,
'exp_name': 'RTD3_NoMemGate_MDP_HalfCheetah_PreLSTM1L128_HistMemory5Len_LSTM1L128_HidStateLayer0_1L128_NoFreeze'}
from spinup.utils.run_utils import setup_logger_kwargs
logger_kwargs = setup_logger_kwargs(args['exp_name'], args['seed'])
td3(env_name=args['env'], actor_critic=core.MLPActorCritic,
ac_kwargs=dict(hidden_sizes=[args['hid']]*args['l']),
max_hist_len=args['max_hist_len'],
gamma=args['gamma'], seed=args['seed'], epochs=args['epochs'],
nonstationary_env=args['nonstationary_env'],
gravity_change_pattern=args['gravity_change_pattern'],
partially_observable=args['partially_observable'],
freeze_hist_coding=args['freeze_hist_coding'],
logger_kwargs=logger_kwargs)
# +
args = {'env': 'HalfCheetah-v2', 'hid': 256, 'l': 2,
'max_hist_len': 5,
'gamma': 0.99, 'seed': 0, 'epochs': 50,
'nonstationary_env':False,
'gravity_change_pattern': 'gravity_averagely_equal',
'partially_observable': False,
'freeze_hist_coding': False,
'exp_name': 'RTD3_MemGate_MDP_HalfCheetah_PreLSTM1L128_HistMemory5Len_LSTM1L128_HidStateLayer0_1L128_NoFreeze'}
from spinup.utils.run_utils import setup_logger_kwargs
logger_kwargs = setup_logger_kwargs(args['exp_name'], args['seed'])
td3(env_name=args['env'], actor_critic=core.MLPActorCritic,
ac_kwargs=dict(hidden_sizes=[args['hid']]*args['l']),
max_hist_len=args['max_hist_len'],
gamma=args['gamma'], seed=args['seed'], epochs=args['epochs'],
nonstationary_env=args['nonstationary_env'],
gravity_change_pattern=args['gravity_change_pattern'],
partially_observable=args['partially_observable'],
freeze_hist_coding=args['freeze_hist_coding'],
logger_kwargs=logger_kwargs)
# +
args = {'env': 'HalfCheetah-v2', 'hid': 256, 'l': 2,
'max_hist_len': 5,
'gamma': 0.99, 'seed': 0, 'epochs': 50,
'nonstationary_env':False,
'gravity_change_pattern': 'gravity_averagely_equal',
'partially_observable': True,
'freeze_hist_coding': False,
'exp_name': 'RTD3_MemGate_POMDP_HalfCheetah_PreLSTM1L128_HistMemory5Len_LSTM1L128_HidStateLayer0_1L128_NoFreeze'}
from spinup.utils.run_utils import setup_logger_kwargs
logger_kwargs = setup_logger_kwargs(args['exp_name'], args['seed'])
td3(env_name=args['env'], actor_critic=core.MLPActorCritic,
ac_kwargs=dict(hidden_sizes=[args['hid']]*args['l']),
max_hist_len=args['max_hist_len'],
gamma=args['gamma'], seed=args['seed'], epochs=args['epochs'],
nonstationary_env=args['nonstationary_env'],
gravity_change_pattern=args['gravity_change_pattern'],
partially_observable=args['partially_observable'],
freeze_hist_coding=args['freeze_hist_coding'],
logger_kwargs=logger_kwargs)
# +
args = {'env': 'HalfCheetah-v2', 'hid': 256, 'l': 2,
'max_hist_len': 5,
'gamma': 0.99, 'seed': 0, 'epochs': 50,
'nonstationary_env':False,
'gravity_change_pattern': 'gravity_averagely_equal',
'partially_observable': True,
'freeze_hist_coding': False,
'exp_name': 'RTD3_NoMemGate_POMDP_HalfCheetah_PreLSTM1L128_HistMemory5Len_LSTM1L128_HidStateLayer0_1L128_NoFreeze'}
from spinup.utils.run_utils import setup_logger_kwargs
logger_kwargs = setup_logger_kwargs(args['exp_name'], args['seed'])
td3(env_name=args['env'], actor_critic=core.MLPActorCritic,
ac_kwargs=dict(hidden_sizes=[args['hid']]*args['l']),
max_hist_len=args['max_hist_len'],
gamma=args['gamma'], seed=args['seed'], epochs=args['epochs'],
nonstationary_env=args['nonstationary_env'],
gravity_change_pattern=args['gravity_change_pattern'],
partially_observable=args['partially_observable'],
freeze_hist_coding=args['freeze_hist_coding'],
logger_kwargs=logger_kwargs)
# +
args = {'env': 'Ant-v2', 'hid': 256, 'l': 2,
'max_hist_len': 5,
'gamma': 0.99, 'seed': 0, 'epochs': 50,
'nonstationary_env':False,
'gravity_change_pattern': 'gravity_averagely_equal',
'partially_observable': True,
'freeze_hist_coding': False,
'exp_name': 'test_pre_feature_extraction_gated_lstm_DDPG_POMDP_Ant_PreLSTM1L128_HistMemory5Len_LSTM1L128_HidStateLayer0_2L128_NoFreeze'}
from spinup.utils.run_utils import setup_logger_kwargs
logger_kwargs = setup_logger_kwargs(args['exp_name'], args['seed'])
td3(env_name=args['env'], actor_critic=core.MLPActorCritic,
ac_kwargs=dict(hidden_sizes=[args['hid']]*args['l']),
max_hist_len=args['max_hist_len'],
gamma=args['gamma'], seed=args['seed'], epochs=args['epochs'],
nonstationary_env=args['nonstationary_env'],
gravity_change_pattern=args['gravity_change_pattern'],
partially_observable=args['partially_observable'],
freeze_hist_coding=args['freeze_hist_coding'],
logger_kwargs=logger_kwargs)
# -
| spinup/algos/pytorch/lstm_ddpg/Untitled3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="popV-5ukLucj"
# # Murales par arrondissement
# + [markdown] id="7ShroeYKG0aF"
# ## Question
#
# Quel arrondissement (quartier) de Montréal a le plus de murales?
#
# La réponse se trouve dans un des nombreux jeux de données ouvertes de la ville. Ou peut voir ces jeux de données sur le site de la ville:
# https://donnees.montreal.ca/ville-de-montreal
#
# Comme le jeu de données qui nous intéresse est en format CVS, on réutilise la fonction `telecharge_csv` vue dans la partie précédente.
# + [markdown] id="elqpdfPfLfq_"
# ## Fichier des données
# + id="5nkruAbHEA_7"
import csv
import requests
def telecharge_csv(url):
req = requests.get(url)
texte = req.text
lignes = list(csv.reader(texte.splitlines()))
return lignes
MURALES_URL = "https://raw.githubusercontent.com/mtlpy/mp-84-atelier/master/murales_montreal_2021.csv"
data = telecharge_csv(MURALES_URL)
print(data[0])
# + [markdown] id="SzyGcTzsH8J2"
# Regardons à quoi ressemblent les données.
# + id="cuwGtIlqIHKE"
for i in range(5):
print(data[i])
# + [markdown] id="9RxXcCGtIygV"
# La première ligne est le nom des colonnes, les autres sont les données qui nous aident à répondre à la question.
#
# Si on veut savoir quel est l'arrondissement avec le plus de murales, on n'a besoin de consulter une seule de ses colonnes. Laquelle?
# + [markdown] id="S3E2LBQfLiuo"
# ## Question 1
# Quelle colonne doit-on consulter?
#
# 🐍 - nom_arrond
# 🗡️ - no_arrond
# 🐰 - artiste
# 🏰 - annee
# + [markdown] id="whBSlQyzLq6C"
# ## Compte des occurences
#
# On utilise un dictionnaire pour compter le nombre de fois où on rencontre le nom d'un arrondissement. Pour nous aider, on utilise la fonction `len` (pour length) qui nous donne la taille d'une collection.
#
# + id="KJ3wHY4gJGvY"
nb_lignes = len(data)
murales_par_arrondissement = dict()
for i in range(1, nb_lignes):
row = data[i]
arrondissement = row[0]
if arrondissement in murales_par_arrondissement:
murales_par_arrondissement[arrondissement] += 1
else:
murales_par_arrondissement[arrondissement] = 1
print(murales_par_arrondissement)
# + [markdown] id="FrqBga-YQrOg"
# ## Triage des résultats
# + [markdown] id="ckIJ5rYqOImB"
# Il nous reste à trier par nombre d'occurenceces. On ne peut pas changer l'ordre des dictionnaires, mais on peut changer l'ordre d'une liste. On va donc devoir transformer notre dictionnaire en liste puis trier cette dernière. La rebrique d'aide de `dict` et de `list` nous donne les fonctions sur ces types collections que nous n'avons pas encore vu. On peut aussi consulter la rebrique d'aide pour une fonction spécifique pour plus de détails.
# + id="uNAMcjqXN-PV"
help(dict)
# + id="UfgCgC2dSa8F"
help(list)
help(list.sort)
# + id="f5dI8jLINcq3"
top_murales = [[nb_occ, nom] for nom, nb_occ in murales_par_arrondissement.items()]
print(top_murales)
top_murales.sort()
print(top_murales)
# + [markdown] id="vUvFJhi7Su6s"
# ## Exercices
# + [markdown] id="WPJeTK61SyuX"
# 1. Changez l'ordre pour avoir l'arrondissement avec le plus d'occurence en premier
# 1. Affichez les 3 premiers résultats dans un format plus aggréable à lire. Par exemple:
# - Nom 1: 21
# - Nom 2: 10
# 1. Bonnus: Triez la liste en ordre alphabetique sur le nom de l'arondissement plutot que sur le nombre d'occurence.
#
#
# + [markdown] id="-shgXT3vnKxW"
# # License
# + [markdown] id="DQDRMYMKnOVi"
# Copyright 2021 Montréal-Python
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
#
| Murales-1.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 1
# Create a bank acc class that has 2 attributes : 'ownerName' and 'balance'
# and 2 methods : 'deposit' and 'withdraw'
# withdrawals may not exceed the available balance
# instantiate your class, make several deposits and withdrawals and test to make sure that
# this account can't be withdrawn
class Bank():
def __init__(self,owner,balance=0):
self.owner = owner
self.balance = balance
def deposit(self, balance):
self.balance += balance
return self.balance
def withdraw(self,balance):
if self.balance >= balance:
self.balance -= balance
else:
return "Not Possible only " + str(self.balance) + ' is avaliable'
return self.balance
account1 = Bank('account1',0)
account1.deposit(1000)
account1.withdraw(400)
account1.withdraw(700)
# # Assignment 2
# Create a cone class that has 2 attributes
# r = radius and h = height
# and 2 methods :
volume = pi * r * r * (h/3)
surface area = base: pi * r*r , side: pi * r (sqrt of(r**2 + h**2))
# use import math
# +
import math
class Cone():
def __init__(self,r,h):
self.r = r
self.h = h
def volume_func(self):
self.volume = math.pi * self.r * self.r * (self.h/3)
return self.volume
def surface_area_func(self):
base = math.pi * self.r * self.r
side = math.pi * self.r * math.sqrt(self.r**2 + self.h**2)
self.surface_area = base + side
return self.surface_area
# -
cone1 = Cone(3,5)
cone1.volume_func()
cone1.surface_area_func()
| Day 6 - Assignment 1,2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="FseUhwIf_Yfg" colab={"base_uri": "https://localhost:8080/"} outputId="2ff31c2c-bb4c-427e-a1d7-89c3442f8102"
# !pip install keras-tuner
# + id="DxvOsIGq_YSZ"
import tensorflow as tf
from tensorflow import keras
import numpy as np
# + id="FxgoXpboha1e" colab={"base_uri": "https://localhost:8080/"} outputId="5855885c-f96c-4103-84c9-91070e977e8c"
# !wget 'https://s3.amazonaws.com/isic-challenge-data/2019/ISIC_2019_Training_GroundTruth.csv'
# + id="-KeFB_PQhUSG" colab={"base_uri": "https://localhost:8080/"} outputId="6a9ade88-1705-4a85-f7d0-fa9700596746"
# !wget 'https://s3.amazonaws.com/isic-challenge-data/2019/ISIC_2019_Training_Metadata.csv'
# + id="ULKVZWzj56QQ" colab={"base_uri": "https://localhost:8080/"} outputId="5b341ed9-e5a7-464d-f71b-fb1f1a416851"
# !wget 'https://s3.amazonaws.com/isic-challenge-data/2019/ISIC_2019_Training_Input.zip'
# + id="Mx9kopXtf47r" colab={"base_uri": "https://localhost:8080/"} outputId="8f4ff6dd-6fdc-4e75-eb4e-7f219d30ffab"
# !unzip 'ISIC_2019_Training_Input.zip'
# + id="E-CRfWJ0Vrjn"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import keras
# + id="UysT9yEJVroV"
labels = pd.read_csv('ISIC_2019_Training_GroundTruth.csv')
label_df = pd.DataFrame(labels)
# + id="rjeXCMc1Vrrc"
info = pd.read_csv('ISIC_2019_Training_Metadata.csv')
info_df = pd.DataFrame(info)
# + id="xqoWqXhgVruc" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="9c1bac7c-4533-45ef-dc5a-fbdc99f35d76"
label_df.tail()
# + id="OuNLr8rPh3DO" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="be1cbe9c-7beb-4bb3-9893-f47ecc35d13a"
info_df.head()
# + id="A-FY9P28YbvI"
info_df = info_df.drop('image',axis=1)
# + id="K3wIS9vPVr8t" colab={"base_uri": "https://localhost:8080/", "height": 402} outputId="eaf35ac1-effe-411e-ba12-8aa77389d2e9"
info_df
# + id="wK4KHiVxXsem"
data = pd.concat([label_df, info_df], axis=1)
# + id="PHy1xSaXXsgy" colab={"base_uri": "https://localhost:8080/", "height": 343} outputId="c5da8a69-36ec-4f4f-a22a-342bf81ae714"
data.head(10)
# + id="iNgexiXPZGMa"
data['lesion_id'].fillna(method ='bfill', inplace=True)
data["age_approx"].fillna(30.0, inplace = True)
data["sex"].fillna("male", inplace = True)
data["anatom_site_general"].fillna( method ='ffill', inplace = True)
# + id="T9ZSCfoNLS9r" colab={"base_uri": "https://localhost:8080/", "height": 639} outputId="85be4aac-903f-427a-e2a9-5f5c9b886d7a"
data.head(20)
# + id="2Cst5RU5Jo-z" colab={"base_uri": "https://localhost:8080/"} outputId="4e59a394-bddf-4883-ff11-0c343e2cc7d2"
rows_with_nan = []
for index, row in data.iterrows():
is_nan_series = row.isnull()
if is_nan_series.any():
rows_with_nan.append(index)
print(rows_with_nan)
# + id="PYPY2064dd5C" colab={"base_uri": "https://localhost:8080/"} outputId="503930a2-7335-47d2-c2c1-fd7db40bacab"
data['anatom_site_general'].unique()
# + id="19ZwrjLReOya"
anatom_site_general = {'anterior torso': 1,'upper extremity': 2,'posterior torso':3,'lower extremity':4, 'lateral torso':5,'head/neck':6,'palms/soles':7,'oral/genital':8}
data['anatom_site_general'] = [anatom_site_general[item] for item in data['anatom_site_general']]
# + id="6sP8dOKPD4qM"
sex = {'male': 0,'female':1}
data['sex'] = [sex[item] for item in data['sex']]
# + id="8j6kl9A3vxl9" colab={"base_uri": "https://localhost:8080/"} outputId="914b7c1c-998d-4351-d602-7d38d37f08af"
len(data['lesion_id'].unique())
# + id="omH03AXID0cz" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="199b7f58-60c4-4134-8311-2dcde801bf24"
data.head(6)
# + id="_aBWlkGIvw4C"
data = data.drop(['lesion_id'],axis=1)
# + id="rpYN7xLBZIeh"
target = data[['MEL']].values
# + id="__NAZhrsYV_a"
data = data.drop(['image','MEL','NV'],axis=1)
# + id="LqvPEh_HY7-z" colab={"base_uri": "https://localhost:8080/", "height": 402} outputId="342bf9d7-6ab3-4c40-e8d3-a92b3984eb74"
data
# + id="bV667Gp4Y8EQ"
label = target
# + id="yW7RZQdSY8Gw" colab={"base_uri": "https://localhost:8080/"} outputId="741172e4-b48a-4d32-b2a9-b4814cc3339f"
list0 = [data, label]
list1 = ['x_train','y_train']
for i in range(2):
print('The shape of the {} is {}'.format(list1[i],list0[i].shape))
# + id="LvqWm02BY8JX" colab={"base_uri": "https://localhost:8080/"} outputId="7709f31e-c0a0-4067-b550-9a7e67c8df65"
_,D = data.shape
print(D)
# + id="p6zw95gai0um"
from google.colab import files
import cv2
# + [markdown] id="_FymV5RtPtAG"
# **I use this part to upload the downloaded images instead of download them in the colab.**
# + id="uAjeudjQY8ke" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="11db6458-94c8-4e3d-f95a-d50d4354dd6e"
'''uploaded = files.upload()
train_image = []
for i in uploaded.keys():
train_image.append(cv2.resize(cv2.cvtColor(cv2.imread(i), cv2.COLOR_BGR2RGB), (32,32)))'''
# + id="3ayVcghzFwPx"
# Upload the images from folder
import os
def load_images_from_folder(folder):
train_image = []
for filename in os.listdir(folder):
img = cv2.imread(os.path.join(folder,filename))
if img is not None:
train_image.append(cv2.resize(cv2.cvtColor(img, cv2.COLOR_BGR2RGB), (32,32)))
return train_image
# + id="ABlV0xUvnIbC"
images = load_images_from_folder('ISIC_2019_Training_Input')
# + id="fwDcrSq89bpP"
from sklearn.preprocessing import StandardScaler
# + id="vy9NF6QV9bs0"
train_image = images[:20264]
test_image = images[20264:]
x_train = data[:20264]
x_test = data[20264:]
y_train = label[:20264]
y_test = label[20264:]
# + id="4zzRYUtzBrLF"
scaler = StandardScaler()
x_train = scaler.fit_transform(x_train)
x_test = scaler.fit_transform(x_test)
# + id="ccQ04O45BVNe" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="5b6dbce2-a0ce-4512-cbb7-e5e569aeb916"
i = 2
plt.imshow(train_image[i])
plt.xticks([])
plt.yticks([])
plt.show()
print(y_train[i][0])
# + id="sT03PmLl8ck0" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="64f84bec-4476-4ea9-a4cb-41564a295c2b"
i = 0
plt.imshow(test_image[i])
plt.xticks([])
plt.yticks([])
plt.show()
print(y_test[i][0])
# + id="cBmD6_MlkD7a"
train_image = np.asarray(train_image)
test_image = np.asarray(test_image)
# + id="xXMiu6AiY8nV"
train_image = train_image.astype('float32')
test_image = test_image.astype('float32')
# + id="tzHAOr2gY8v-"
mean = np.mean(train_image,axis=(0,1,2,3))
std = np.std(train_image,axis=(0,1,2,3))
train_image = (train_image-mean)/(std+1e-7)
test_image = (test_image-mean)/(std+1e-7)
# + id="dAF_-z0yY80v"
from keras.utils import np_utils
nClasses = 2
y_train = np_utils.to_categorical(y_train, nClasses)
y_test = np_utils.to_categorical(y_test, nClasses)
# + id="dyqmNDSdY8zI" colab={"base_uri": "https://localhost:8080/"} outputId="aea0a33d-d2ed-4037-dc9f-f190abb26bf6"
print(test_image.shape)
print(y_train.shape)
print(y_test.shape)
# + id="5AbBn4b8Y8il"
input_shape = (32,32,3)
# + id="Uw3pmeoWxl6S"
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Activation , GlobalAveragePooling2D
from keras import layers
from keras.models import Model
from keras.models import Sequential
from keras.layers import Dense, Dropout , Input , Flatten , Conv2D , MaxPooling2D
from keras.layers.merge import concatenate
from keras.callbacks import EarlyStopping , ModelCheckpoint
from keras.optimizers import Adam, SGD, RMSprop
from keras import regularizers
from kerastuner.tuners import RandomSearch
# + [markdown] id="SQIwrkCF10wV"
# # **Transfer Learning**
# + [markdown] id="_DQopSrjJHEm"
# Transfer Learning with Keras-Tuner to find the best hyper parameters.
# + id="omX51bV-lx7k" colab={"base_uri": "https://localhost:8080/"} outputId="a5a1e820-7b8e-4d57-9b61-9ac24531e49e"
def build_model(hp):
base_model = keras.applications.MobileNetV2(input_shape=input_shape, alpha=1.0, include_top=False, weights="imagenet", input_tensor=None, pooling=None, classes=1000, classifier_activation="softmax")
base_model.trainable = False
# model_1
model1_in = keras.Input(shape=input_shape)
x = base_model(model1_in, training=False)
x = layers.Conv2D(64,(1,1),padding='same', activation='relu')(x)
x = layers.Conv2D(64,(1,1), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(1, 1))(x)
x = layers.Conv2D(128,(1,1),padding='same', activation='relu')(x)
x = layers.Conv2D(128,(1,1), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(1,1))(x)
x = layers.Conv2D(256,(1,1),padding='same', activation='relu')(x)
x = layers.Conv2D(256,(1,1), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(1,1))(x)
x = layers.Conv2D(512,(1,1),padding='same', activation='relu')(x)
x = layers.Conv2D(512,(1,1), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(1,1))(x)
x = layers.Flatten()(x)
x = layers.Dense(512, activation='relu')(x)
x = layers.Dense(256, activation='relu')(x)
x = layers.Dense(512, activation='relu')(x)
x = layers.Dense(1024, activation='relu')(x)
model1_out = layers.Dense(nClasses, activation='sigmoid',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
model1 = keras.Model(model1_in, model1_out)
# model_2
model2_in = keras.Input(shape=(D,))
x = layers.Dense(16384, kernel_initializer='normal')(model2_in)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(8192, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(4096, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(2048, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(1024, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(512, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(128, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(64, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(32, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(16, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(4, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
model2_out = layers.Dense(nClasses, kernel_initializer='normal',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
model2 = keras.Model(model2_in, model2_out)
# concatanation
concatenated = concatenate([model1_out, model2_out])
x = layers.Dense(units=hp.Int('units', min_value=32, max_value=512, step=32), activation='relu')(concatenated)
out = Dense(2, activation='sigmoid', name='output_layer',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
merged_model = Model([model1_in, model2_in], out)
merged_model.compile(optimizer=keras.optimizers.Adam(hp.Choice('learning_rate',values=[1e-2, 1e-3, 1e-4])),loss='binary_crossentropy',metrics=['accuracy'])
return merged_model
tuner = RandomSearch(build_model,objective='val_accuracy',max_trials=5,executions_per_trial=3,directory='my_dir',project_name='helloworld')
# + id="eeG__svxD8eK" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="57663906-c80c-42c9-9eb1-4d676486af69"
tuner.search_space_summary()
# + id="Xn4ITy2AEADR" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c554236e-0521-4e8c-dfb1-5eae4242a0e7"
tuner.search([train_image,x_train], y_train,
epochs=5,
validation_data=([test_image,x_test],y_test))
# + [markdown] id="cRiL6vFTJaXV"
# Transfer Learning with Augmentation
# + id="Ow4xiCqvsmQ4"
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Activation , GlobalAveragePooling2D
from keras import layers
from keras.models import Model
from keras.models import Sequential
from keras.layers import Dense, Dropout , Input , Flatten , Conv2D , MaxPooling2D
from keras.layers.merge import concatenate
from keras.callbacks import EarlyStopping , ModelCheckpoint
from keras.optimizers import Adam, SGD, RMSprop
from keras import regularizers
# + id="oM4V0SlxX1n2"
input_shape = (32,32,3)
# + id="5GKsFPHztJ-L"
def pure_cnn_model():
base_model = keras.applications.MobileNetV2(input_shape=input_shape, alpha=1.0, include_top=False, weights="imagenet", input_tensor=None, pooling=None, classes=1000, classifier_activation="softmax")
base_model.trainable = False
# model_1
model1_in = keras.Input(shape=input_shape)
x = base_model(model1_in, training=False)
x = layers.Conv2D(32,(1,1),padding='same', activation='sigmoid')(x)
x = layers.MaxPooling2D(pool_size=(1, 1))(x)
x = layers.Conv2D(128,(1,1),padding='same', activation='relu')(x)
x = layers.Conv2D(128,(1,1), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(1,1))(x)
x = layers.Conv2D(256,(1,1),padding='same', activation='relu')(x)
x = layers.Conv2D(256,(1,1), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(1,1))(x)
x = layers.Conv2D(512,(1,1),padding='same', activation='relu')(x)
x = layers.Conv2D(512,(1,1), activation='relu')(x)
x = layers.MaxPooling2D(pool_size=(1,1))(x)
x = layers.Flatten()(x)
x = layers.Dense(256, activation='relu')(x)
x = layers.Dense(512, activation='relu')(x)
x = layers.Dense(1024, activation='relu')(x)
model1_out = layers.Dense(2, activation='sigmoid',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
model1 = keras.Model(model1_in, model1_out)
# model_2
model2_in = keras.Input(shape=(D,))
x = layers.Dense(16384, kernel_initializer='normal')(model2_in)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(8192, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(4096, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(2048, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(1024, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(512, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(128, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(64, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(32, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(16, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
x = layers.Dense(4, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
#x = layers.Dropout(0.1)(x)
model2_out = layers.Dense(2, kernel_initializer='normal',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
model2 = keras.Model(model2_in, model2_out)
# concatanation
concatenated = concatenate([model1_out, model2_out])
x = Dense(16384, activation='relu')(concatenated)
x = layers.Dense(8192, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(4096, kernel_initializer='normal')(x)
x = layers.BatchNormalization()(x)
x = layers.Activation('relu')(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(2048, activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(1024, activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(512, activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(256, activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(128, activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Dense(64, activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(32, activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Dense(16, activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Dropout(0.1)(x)
x = layers.Dense(8, activation='relu',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
x = layers.BatchNormalization()(x)
x = layers.Dense(4, activation='relu',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
out = Dense(2, activation='sigmoid', name='output_layer',kernel_regularizer=regularizers.l1_l2(l1=1e-5, l2=1e-4))(x)
merged_model = Model([model1_in, model2_in], out)
return merged_model
# + id="fiWDD39Lpon6" colab={"base_uri": "https://localhost:8080/"} outputId="17fc6852-5082-432e-8bdf-fbc60863963c"
datagen = ImageDataGenerator(
featurewise_center=True,
samplewise_center=True,
featurewise_std_normalization=True,
samplewise_std_normalization=True,
zca_whitening=True,
rotation_range=45,
width_shift_range=0.2,
height_shift_range=0.2,
horizontal_flip=True,
vertical_flip=True)
datagen.fit(train_image)
# + id="Jk9iR6JYpom3" colab={"base_uri": "https://localhost:8080/"} outputId="ed2e54f1-b031-4748-b4d0-05eaf8f7c624"
augmented_model = pure_cnn_model()
# + id="HaNt13Rgpol4"
augmented_checkpoint = ModelCheckpoint('augmented_best_model.h5',
monitor='val_loss',
verbose=0,
save_best_only= True,
mode='auto')
# + id="VMvG74_HCniW"
augmented_model.compile(loss='binary_crossentropy',
optimizer=Adam(lr=1.0e-4),
metrics = ['accuracy'])
# + id="5nN3GWmcpok6" colab={"base_uri": "https://localhost:8080/"} outputId="5f7150d1-1490-4e19-fa12-a94788b20880"
augmented_model_details = augmented_model.fit_generator(datagen.flow([train_image, x_train], y_train, batch_size = 32),
steps_per_epoch = len(train_image) / 32,
validation_data=([test_image, x_test], y_test),
epochs= 10,
callbacks=[augmented_checkpoint],
verbose=1)
| project_melanomia.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.1
# language: julia
# name: julia-1.3
# ---
# Import needed libs
using CSV
using Optim
using Statistics
using Plots
# Read data
X_train = convert(Matrix, CSV.read("X_train.csv"));
X_test = convert(Matrix, CSV.read("X_test.csv"));
y_train = CSV.read("y_train.csv")[:, 1];
y_test = CSV.read("y_test.csv")[:, 1];
# +
# Normalise the training design matrix
function scale_features(X)
μ = mean(X, dims=1)
σ = std(X, dims=1)
X_norm = (X .- μ) ./ σ
return (X_norm, μ, σ);
end
# Normalise the testing design matrix
function transform_features(X, μ, σ)
X_norm = (X .- μ) ./ σ
return X_norm;
end
# Scale training features and get artificats for future use
X_train_scaled, μ, σ = scale_features(X_train);
# Transform the testing features by using the learned artifacts
X_test_scaled = transform_features(X_test, μ, σ);
# -
function sigmoid(z)
return 1 ./ (1 .+ exp.(.-z))
end
function regularised_cost(X, y, θ, λ)
m = length(y)
# Sigmoid predictions at current batch
h = sigmoid(X * θ)
# left side of the cost function
positive_class_cost = ((-y)' * log.(h))
# right side of the cost function
negative_class_cost = ((1 .- y)' * log.(1 .- h))
# lambda effect
lambda_regularization = (λ/(2*m) * sum(θ[2 : end] .^ 2))
# Current batch cost. Basically mean of the batch cost plus regularization penalty
𝐉 = (1/m) * (positive_class_cost - negative_class_cost) + lambda_regularization
# Gradients for all the theta members with regularization except the constant
∇𝐉 = (1/m) * (X') * (h-y) + ((1/m) * (λ * θ)) # Penalise all members
∇𝐉[1] = (1/m) * (X[:, 1])' * (h-y) # Exclude the constant
return (𝐉, ∇𝐉)
end
function logistic_regression_sgd(X, y, λ, fit_intercept=true, η=0.01, max_iter=1000)
# Initialize some useful values
m = length(y); # number of training examples
if fit_intercept
# Add a constant of 1s if fit_intercept is specified
constant = ones(m, 1)
X = hcat(constant, X)
else
X # Assume user added constants
end
# Use the number of features to initialise the theta θ vector
n = size(X)[2]
θ = zeros(n)
# Initialise the cost vector based on the number of iterations
𝐉 = zeros(max_iter)
for iter in range(1, stop=max_iter)
# Calcaluate the cost and gradient (∇𝐉) for each iter
𝐉[iter], ∇𝐉 = regularised_cost(X, y, θ, λ)
# Update θ using gradients (∇𝐉) for direction and (η) for the magnitude of steps in that direction
θ = θ - (η * ∇𝐉)
end
return (θ, 𝐉)
end
θ, 𝐉 = logistic_regression_sgd(X_train_scaled, y_train, 0.0001, true, 0.3, 3000);
plot(𝐉, color="blue", title="Cost Per Iteration", legend=false,
xlabel="Num of iterations", ylabel="Cost")
function predict_proba(X, θ, fit_intercept=true)
m = size(X)[1]
if fit_intercept
# Add a constant of 1s if fit_intercept is specified
constant = ones(m, 1)
X = hcat(constant, X)
else
X
end
h = sigmoid(X * θ)
return h
end
function predict_class(proba, threshold=0.5)
return proba .>= threshold
end
train_score = mean(y_train .== predict_class(predict_proba(X_train_scaled, θ)));
println("Training score: ", round(train_score, sigdigits=4))
test_score = mean(y_test .== predict_class(predict_proba(X_test_scaled, θ)));
println("Testing score: ", round(test_score, sigdigits=4))
| Tutorials/HTRU2_julia_project/updated_log_reg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="UU22Gz_V2S5T"
# # ***Hierarchial Clustering***
# + [markdown] id="ocTYyRnW2dEr"
# ### Importing Libraries
# + id="nSZbooWR2LkS"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] id="BtuE_-5I2lS5"
# ### Importing Dataset
# + id="A03wI6l22n_E"
dataset = pd.read_csv('Mall_Customers.csv') #loading/reading dataset using pandas
X = dataset.iloc[:, [3, 4]].values #selecting all values of indexes 3 and 4 which are annual income and spending score
#y = dataset.iloc[:, 3].values
# + [markdown] id="iwdY7Mih2s0S"
# ### Splitting the dataset into the Training set and Test set
# + id="RhxVaiJ22tp0"
'''from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)'''
# + [markdown] id="0UKSddZ72zvD"
# ### Feature Scaling
# + id="YQTHnBVw2zMX"
"""from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
X_train = sc_X.fit_transform(X_train)
X_test = sc_X.transform(X_test)
sc_y = StandardScaler()
y_train = sc_y.fit_transform(y_train)"""
# + [markdown] id="56z1sbFK275A"
# ### Using the dendrogram to find the optimal number of clusters
# + id="V-CWFBfm259g"
import scipy.cluster.hierarchy as sch #importing cluster.hierarchy from scipy as sch
dendrogram = sch.dendrogram(sch.linkage(X, method = 'ward')) #using dendogram function and inputting linkage function as its argument. Method ward is used for minimizing variance inside clusters
plt.title('Dendrogram') #Naming Title of graph
plt.xlabel('Customers') #Naming entity being represented on x axis
plt.ylabel('Euclidean distances') #Naming entity being represented on y axis
plt.show()
# + [markdown] id="3UGk5jkw2_6Y"
# ### Fitting Hierarchical Clustering to the datase
# + id="ir4KKSvB3Crn"
from sklearn.cluster import AgglomerativeClustering
hc = AgglomerativeClustering(n_clusters = 5, affinity = 'euclidean', linkage = 'ward') #hc is an object of AgglomerativeClustering. We specify Number of clusters as 5 based on dendograms, affinity is the distance method you want to use.
y_hc = hc.fit_predict(X)
# + [markdown] id="39V2-09J3FKo"
# ### Visualising the clusters
# + id="sOXNyysQ3IL3"
plt.scatter(X[y_hc == 0, 0], X[y_hc == 0, 1], s = 100, c = 'red', label = 'Cluster 1')
plt.scatter(X[y_hc == 1, 0], X[y_hc == 1, 1], s = 100, c = 'blue', label = 'Cluster 2')
plt.scatter(X[y_hc == 2, 0], X[y_hc == 2, 1], s = 100, c = 'green', label = 'Cluster 3')
plt.scatter(X[y_hc == 3, 0], X[y_hc == 3, 1], s = 100, c = 'cyan', label = 'Cluster 4')
plt.scatter(X[y_hc == 4, 0], X[y_hc == 4, 1], s = 100, c = 'magenta', label = 'Cluster 5')
plt.title('Clusters of customers')
plt.xlabel('Annual Income (k$)')
plt.ylabel('Spending Score (1-100)')
plt.legend()
plt.show()
| 5_Clustering/Hierarchical_Clustering/hc_writeup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
pew = pd.read_csv('../data/pew.csv')
pew
pd.melt(pew, id_vars='religion')
pd.melt(pew, id_vars='religion',
var_name='income', value_name='count')
billboard = pd.read_csv('../data/billboard.csv')
billboard.head()
pd.melt(billboard,
id_vars=['year', 'artist', 'track', 'time', 'date.entered'],
var_name='week',
value_name='rank')
tb = pd.read_csv('../data/tb.csv')
tb
ebola = pd.read_csv('../data/ebola_country_timeseries.csv')
ebola.head()
ebola_melt = pd.melt(ebola,
id_vars=['Date', 'Day'],
var_name='cd_country',
value_name='count')
ebola_melt
var_split = ebola_melt['cd_country'].str.split('_')
status_values = var_split.str.get(0)
country_values = var_split.str.get(1)
ebola_melt['status'] = status_values
ebola_melt['country'] = country_values
ebola_melt.head()
variable_split = ebola_melt['cd_country'].str.split('_', expand=True)
variable_split
variable_split.columns = ['status1', 'country1']
variable_split.head()
ebola_clean = pd.concat([ebola_melt, variable_split], axis=1)
ebola_clean.head()
weather = pd.read_csv('../data/weather.csv')
weather.head()
weather_melt = pd.melt(weather,
id_vars=['id', 'year', 'month', 'element'],
var_name='day',
value_name='temp')
weather_melt.head()
weather_tidy = weather_melt.pivot_table(
index=['id', 'year', 'month', 'day'],
columns='element',
values='temp'
).reset_index()
weather_tidy
'd10'[1:]
# +
# pd.pivot_table?
# -
| 02-lesson/04-tidy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><h1> Data for Visualization </center></h1>
#
# This notebook shows the code for preparing some data for visualization.
# ## Trending Topics
# +
# Import necessary libraries
import pandas as pd
from wordcloud import WordCloud, STOPWORDS
stopwords = set(STOPWORDS) #Set of English Stopwords
import numpy as npy
from PIL import Image
maskArray = npy.array(Image.open("mask.png")) # Twitter Logo as a mask for wordcloud
# +
# Import relevant files
sent_df = pd.read_csv("Processed Data.csv",sep='\t')
topic_df = pd.read_csv("Topic Modelled Data.csv",sep='\t')
# -
relevant_df = sent_df.loc[:,['date','time','id']]
df = pd.merge(left=relevant_df, right=topic_df, left_on='id', right_on='id') #Merge the dataframe with respect to ID
df.sample(3) #Sample Data
# +
#Drop irrevelant data
df.drop(['id','topic_perc_contrib','time','processed_text'],axis=1,inplace=True)
# -
adf=df.groupby('date')['topic_data'].value_counts()
# +
#Group the data with respect to date and count the number of topics
df=df.groupby('date')['topic_data'].value_counts()
df = pd.DataFrame(df)
df.rename(columns={'topic_data': 'topic_count'},inplace=True)
df.reset_index(inplace=True)
df
# +
#Split the data into different lockdown phases and save the data
lockdown1 = df[(df['date'] >= '2020-03-25') & (df['date'] <= '2020-04-14')]
lockdown1 = lockdown1.groupby('topic_data').sum()
lockdown1.reset_index(inplace=True)
lockdown1.sort_values(by='topic_count',inplace=True,ascending=False)
lockdown1.insert(0, "Phase", "LD1")
lockdown2 = df[(df['date'] >= '2020-04-15') & (df['date'] <= '2020-05-03')]
lockdown2 = lockdown2.groupby('topic_data').sum()
lockdown2.reset_index(inplace=True)
lockdown2.sort_values(by='topic_count',inplace=True,ascending=False)
lockdown2.insert(0, "Phase", "LD2")
lockdown3 = df[(df['date'] >= '2020-05-04') & (df['date'] <= '2020-05-17')]
lockdown3 = lockdown3.groupby('topic_data').sum()
lockdown3.reset_index(inplace=True)
lockdown3.sort_values(by='topic_count',inplace=True,ascending=False)
lockdown3.insert(0, "Phase", "LD3")
lockdown4 = df[(df['date'] >= '2020-05-18') & (df['date'] <= '2020-05-31')]
lockdown4 = lockdown4.groupby('topic_data').sum()
lockdown4.reset_index(inplace=True)
lockdown4.sort_values(by='topic_count',inplace=True,ascending=False)
lockdown4.insert(0, "Phase", "LD4")
unlock1 = df[(df['date'] >= '2020-06-01') & (df['date'] <= '2020-06-14')]
unlock1 = unlock1.groupby('topic_data').sum()
unlock1.reset_index(inplace=True)
unlock1.sort_values(by='topic_count',inplace=True,ascending=False)
unlock1.insert(0, "Phase", "Unlock1")
combined_df = pd.concat([lockdown1,lockdown2,lockdown3,lockdown4,unlock1],ignore_index=True)
combined_df.to_csv("Topic data.csv",index=False)
# -
# ## Trending Hashtags
# Read the data
hash_df = sent_df.loc[:,['date','hashtags']]
hash_df.sample(3)
# +
#Split the data into different lockdown phases, find the top 10 hashtags and save the data
lockdown1 = hash_df[(hash_df['date'] >= '2020-03-25') & (hash_df['date'] <= '2020-04-14')]
lockdown1 = pd.DataFrame(lockdown1['hashtags'].str.split(expand=True).stack().value_counts()).head(10)
lockdown1.reset_index(inplace=True)
lockdown1.insert(0, "Phase", "LD1")
lockdown1.rename({0:"value","index":"hashtag"},axis=1,inplace=True)
lockdown2 = hash_df[(hash_df['date'] >= '2020-04-15') & (hash_df['date'] <= '2020-05-03')]
lockdown2 = pd.DataFrame(lockdown2['hashtags'].str.split(expand=True).stack().value_counts()).head(10)
lockdown2.reset_index(inplace=True)
lockdown2.insert(0, "Phase", "LD2")
lockdown2.rename({0:"value","index":"hashtag"},axis=1,inplace=True)
lockdown3 = hash_df[(hash_df['date'] >= '2020-05-04') & (hash_df['date'] <= '2020-05-17')]
lockdown3 = pd.DataFrame(lockdown3['hashtags'].str.split(expand=True).stack().value_counts()).head(10)
lockdown3.reset_index(inplace=True)
lockdown3.insert(0, "Phase", "LD3")
lockdown3.rename({0:"value","index":"hashtag"},axis=1,inplace=True)
lockdown4 = hash_df[(hash_df['date'] >= '2020-05-18') & (hash_df['date'] <= '2020-05-31')]
lockdown4 = pd.DataFrame(lockdown4['hashtags'].str.split(expand=True).stack().value_counts()).head(10)
lockdown4.reset_index(inplace=True)
lockdown4.insert(0, "Phase", "LD4")
lockdown4.rename({0:"value","index":"hashtag"},axis=1,inplace=True)
unlock1 = hash_df[(hash_df['date'] >= '2020-06-01') & (hash_df['date'] <= '2020-06-14')]
unlock1 = pd.DataFrame(unlock1['hashtags'].str.split(expand=True).stack().value_counts()).head(10)
unlock1.reset_index(inplace=True)
unlock1.insert(0, "Phase", "Unlock1")
unlock1.rename({0:"value","index":"hashtag"},axis=1,inplace=True)
combined_df = pd.concat([lockdown1,lockdown2,lockdown3,lockdown4,unlock1],ignore_index=True)
combined_df.to_csv("Hashtag data.csv",index=False)
# -
# ## Tone Analyser
# +
# Read the data and remove unnecessary columns
tone_df = pd.read_csv("Sentiment Data.csv",sep='\t')
tone_df = tone_df.loc[:,['date','sadness','confident','joy','analytical','anger','tentative','fear']]
tone_df.sample(3)
# +
#Split the data into different lockdown phases and save the data
lockdown1 = tone_df[(tone_df['date'] >= '2020-03-25') & (tone_df['date'] <= '2020-04-14')]
lockdown1 = pd.DataFrame(lockdown1.sum())
lockdown1 = lockdown1.T
lockdown1.drop(['date'],axis=1,inplace=True)
lockdown1.insert(0,"Phase","LD1")
lockdown2 = tone_df[(tone_df['date'] >= '2020-04-15') & (tone_df['date'] <= '2020-05-03')]
lockdown2 = pd.DataFrame(lockdown2.sum())
lockdown2 = lockdown2.T
lockdown2.drop(['date'],axis=1,inplace=True)
lockdown2.insert(0,"Phase","LD2")
lockdown3 = tone_df[(tone_df['date'] >= '2020-05-04') & (tone_df['date'] <= '2020-05-17')]
lockdown3 = pd.DataFrame(lockdown3.sum())
lockdown3 = lockdown3.T
lockdown3.drop(['date'],axis=1,inplace=True)
lockdown3.insert(0,"Phase","LD3")
lockdown4 = tone_df[(tone_df['date'] >= '2020-05-18') & (tone_df['date'] <= '2020-05-31')]
lockdown4 = pd.DataFrame(lockdown4.sum())
lockdown4 = lockdown4.T
lockdown4.drop(['date'],axis=1,inplace=True)
lockdown4.insert(0,"Phase","LD4")
unlock1 = tone_df[(tone_df['date'] >= '2020-06-01') & (tone_df['date'] <= '2020-06-14')]
unlock1 = pd.DataFrame(unlock1.sum())
unlock1 = unlock1.T
unlock1.drop(['date'],axis=1,inplace=True)
unlock1.insert(0,"Phase","Unlock1")
combined_df = pd.concat([lockdown1,lockdown2,lockdown3,lockdown4,unlock1],ignore_index=True)
combined_df = combined_df.melt("Phase")
combined_df.rename({"variable":"Tone","value":"Value"},axis=1,inplace=True)
combined_df = combined_df.sort_values(by="Phase")
combined_df.to_csv("Tone data.csv",index=False)
# -
# ## Wordcloud
# +
# Read the data
df = pd.read_csv("Processed Data.csv",sep='\t')
df['processed_text']=df['processed_text'].astype(str)
df.sample(3)
# +
# Remove unnecessary data
cloud_df = df.loc[:,['date','processed_text']]
cloud_df
# +
#Split the data into different lockdown phases, generate wordcloud and save the data
lockdown1 = cloud_df[(cloud_df['date'] >= '2020-03-25') & (cloud_df['date'] <= '2020-04-14')]
text = []
for item in lockdown1['processed_text']:
text.append(str(data) for data in item)
string = ["".join(data) for data in text]
lockdown1text = " ".join(string)
lockdown1cloud = WordCloud(background_color = "white",stopwords = stopwords,mask = maskArray)
lockdown1cloud.generate(lockdown1text)
lockdown1cloud.to_file("Lockdown1 cloud.png")
lockdown2 = cloud_df[(cloud_df['date'] >= '2020-04-15') & (cloud_df['date'] <= '2020-05-03')]
text = []
for item in lockdown2['processed_text']:
text.append(str(data) for data in item)
string = ["".join(data) for data in text]
lockdown2text = " ".join(string)
lockdown2cloud = WordCloud(background_color = "white",stopwords = stopwords,mask = maskArray)
lockdown2cloud.generate(lockdown2text)
lockdown2cloud.to_file("Lockdown2 cloud.png")
lockdown3 = cloud_df[(cloud_df['date'] >= '2020-05-04') & (cloud_df['date'] <= '2020-05-17')]
text = []
for item in lockdown3['processed_text']:
text.append(str(data) for data in item)
string = ["".join(data) for data in text]
lockdown3text = " ".join(string)
lockdown3cloud = WordCloud(background_color = "white",stopwords = stopwords,mask = maskArray)
lockdown3cloud.generate(lockdown3text)
lockdown3cloud.to_file("Lockdown3 cloud.png")
lockdown4 = cloud_df[(cloud_df['date'] >= '2020-05-18') & (cloud_df['date'] <= '2020-05-31')]
text = []
for item in lockdown4['processed_text']:
text.append(str(data) for data in item)
string = ["".join(data) for data in text]
lockdown4text = " ".join(string)
lockdown4cloud = WordCloud(background_color = "white",stopwords = stopwords,mask = maskArray)
lockdown4cloud.generate(lockdown4text)
lockdown4cloud.to_file("Lockdown4 cloud.png")
unlock1 = cloud_df[(cloud_df['date'] >= '2020-06-01') & (cloud_df['date'] <= '2020-06-14')]
text = []
for item in unlock1['processed_text']:
text.append(str(data) for data in item)
string = ["".join(data) for data in text]
unlock1text = " ".join(string)
unlock1cloud = WordCloud(background_color = "white",stopwords = stopwords,mask = maskArray)
unlock1cloud.generate(unlock1text)
unlock1cloud.to_file("Unlock1 cloud.png")
#General Data
text = []
for item in cloud_df['processed_text']:
text.append(str(data) for data in item)
string = ["".join(data) for data in text]
generaltext = " ".join(string)
generalcloud = WordCloud(background_color = "white",stopwords = stopwords,mask = maskArray)
generalcloud.generate(generaltext)
generalcloud.to_file("General cloud.png")
| Jupyter Notebooks/Visualisation Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Write a Pandas program to join the two given dataframes along rows and assign all data
import pandas as pd
import numpy as np
data1=pd.DataFrame({
'student_id': ['S1', 'S2', 'S3', 'S4', 'S5'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [200, 210, 190, 222, 199]})
data2=pd.DataFrame({
'student_id': ['S4', 'S5', 'S6', 'S7', 'S8'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [201, 200, 198, 219, 201]})
print("printing a original data of first:")
print(data1)
print("printing a original data of second:")
print(data2)
final=pd.concat([data1,data2])
print(final)
#Write a Pandas program to join the two given dataframes along columns and assign all data.
data1=pd.DataFrame({
'student_id': ['S1', 'S2', 'S3', 'S4', 'S5'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [200, 210, 190, 222, 199]})
data2=pd.DataFrame({
'student_id': ['S4', 'S5', 'S6', 'S7', 'S8'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [201, 200, 198, 219, 201]})
print("original data:")
print(data1)
print("-------------------")
print(data2)
print("joinign with column:")
final=pd.concat([data1,data2],axis=1)
print(final)
# # ADDING A ROW IN A DATAFRAME
#Write a Pandas program to append rows to an existing DataFrame and display the combined data.
import numpy as np
import pandas as pd
data1=pd.DataFrame({
'student_id': ['S1', 'S2', 'S3', 'S4', 'S5'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [200, 210, 190, 222, 199]})
print("original data:")
print(data1)
x=pd.Series(['S6','carlos','295'],index=['student_id','name','marks'])
print(x)
main=data1.append(x,ignore_index=True)
print(main)
#Write a Pandas program to append a list of dictioneries or series to a existing DataFrame and display the combined data.
import pandas as pd
import numpy as np
data1=pd.DataFrame({
'student_id': ['S1', 'S2', 'S3', 'S4', 'S5'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [200, 210, 190, 222, 199]})
print(data1)
x=pd.Series(['S6','Bibek','295'],index=['student_id','name','marks'])
print(x)
y=pd.DataFrame({'student_id':'s6','name':'Bibek','marks':260},index=[5])
#print(y)
# yy=pd.DataFrame(y,index=[1])
# print(yy)
final=data1.append(y)
print(final)
import pandas as pd
import numpy as np
data1=pd.DataFrame({
'student_id': ['S1', 'S2', 'S3', 'S4', 'S5'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [200, 210, 190, 222, 199]})
print(data1)
x=[{'student_id':'s6','name':'Bibek','marks':260}]
print(x)
final=data1.append(x,ignore_index=True)
print(final)
import pandas as pd
y={'student_id':'s6','name':'Bibek','marks':260}
yy=pd.DataFrame(y,index=['a'])
print(yy)
#Write a Pandas program to join the two given dataframes along rows and merge with another dataframe along the common column id.
import numpy as np
import pandas as pd
data1=pd.DataFrame({
'student_id': ['S1', 'S2', 'S3', 'S4', 'S5'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [200, 210, 190, 222, 199]})
data2=pd.DataFrame({
'student_id': ['S4', 'S5', 'S6', 'S7', 'S8'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [201, 200, 198, 219, 201]})
exam_data = pd.DataFrame({
'student_id': ['S1', 'S2', 'S3', 'S4', 'S5', 'S7', 'S8', 'S9', 'S10', 'S11', 'S12', 'S13'],
'exam_id': [23, 45, 12, 67, 21, 55, 33, 14, 56, 83, 88, 12]})
print(data1)
print("-----------------------------------------")
print(data2)
print("-----------------------------------------")
print(exam_data)
print("------------------------------------------")
final=data1.append(data2,ignore_index=True)
print("merging data1 and data2:")
print(final)
print("-------------------------------------------")
final_merge=pd.merge(final,exam_data,on='student_id')
print(final_merge)
#Write a Pandas program to join the two dataframes with matching records from both sides where available.
import numpy as np
import pandas as pd
data1=pd.DataFrame({
'student_id': ['S1', 'S2', 'S3', 'S4', 'S5'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [200, 210, 190, 222, 199]})
data2=pd.DataFrame({
'student_id': ['S4', 'S5', 'S6', 'S7', 'S8'],
'name': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'marks': [201, 200, 198, 219, 201]})
print(data1)
print("-----------------------------------")
print(data2)
print("after merging a data:")
final=pd.merge(data1,data2,on='student_id',how='outer')
print(final)
#Write a Pandas program to join (left join) the two dataframes using keys from left dataframe only.
import pandas as pd
data1=pd.DataFrame({'key1': ['<KEY>'],
'key2': ['K0', 'K1', 'K0', 'K1'],
'P': ['P0', 'P1', 'P2', 'P3'],
'Q': ['Q0', 'Q1', 'Q2', 'Q3']})
data2=pd.DataFrame({'key1': ['K0', '<KEY>'],
'key2': ['K0', 'K0', '<KEY>'],
'R': ['R0', 'R1', 'R2', 'R3'],
'S': ['S0', 'S1', 'S2', 'S3']})
print(data1)
print("-----------------------------")
print(data2)
print("print final merge data:")
final=pd.merge(data1,data2,how='left',on=['key1','key2'])
print(final)
#Write a Pandas program to join two dataframes using keys from right dataframe only.
#Write a Pandas program to join (left join) the two dataframes using keys from left dataframe only.
import pandas as pd
data1=pd.DataFrame({'key1': ['K0', 'K0', 'K1', 'K2'],
'key2': ['K0', '<KEY>'],
'P': ['P0', 'P1', 'P2', 'P3'],
'Q': ['Q0', 'Q1', 'Q2', 'Q3']})
data2=pd.DataFrame({'key1': ['K0', '<KEY> 'K2'],
'key2': ['K0', '<KEY>'],
'R': ['R0', 'R1', 'R2', 'R3'],
'S': ['S0', 'S1', 'S2', 'S3']})
print(data1)
print("-----------------------------")
print(data2)
print("print final merge data:")
final=pd.merge(data1,data2,how='right',on=['key1','key2'])
print(final)
#Write a Pandas program to merge two given datasets using multiple join keys
import pandas as pd
data1=pd.DataFrame({'key1': ['K0', 'K0', 'K1', 'K2'],
'key2': ['K0', 'K1', 'K0', 'K1'],
'P': ['P0', 'P1', 'P2', 'P3'],
'Q': ['Q0', 'Q1', 'Q2', 'Q3']})
data2=pd.DataFrame({'key1': ['K0', 'K1', 'K1', 'K2'],
'key2': ['K0', 'K0', 'K0', 'K0'],
'R': ['R0', 'R1', 'R2', 'R3'],
'S': ['S0', 'S1', 'S2', 'S3']})
print(data1)
print("-----------------------------")
print(data2)
print("print final merge data:")
final=pd.merge(data1,data2,on=['key1','key2'])
print(final)
#Write a Pandas program to create a combination from two dataframes where a column id combination
#appears more than once in both dataframes
import pandas as pd
data1=pd.DataFrame({'key1': ['K0', 'K0', 'K1', 'K2'],
'key2': ['K0', 'K1', 'K0', 'K1'],
'P': ['P0', 'P1', 'P2', 'P3'],
'Q': ['Q0', 'Q1', 'Q2', 'Q3']})
data2=pd.DataFrame({'key1': ['K0', 'K1', 'K1', 'K2'],
'key2': ['K0', 'K0', 'K0', 'K0'],
'R': ['R0', 'R1', 'R2', 'R3'],
'S': ['S0', 'S1', 'S2', 'S3']})
print(data1)
print("-----------------------------")
print(data2)
print("print final merge data(many to many join):")
final=pd.merge(data1,data2,on='key1')
print(final)
#Write a Pandas program to combine the columns of two potentially differently-indexed DataFrames into a single result DataFrame
import pandas as pd
import numpy as np
data1=pd.DataFrame({'A': ['A0', 'A1', 'A2'],
'B': ['B0', 'B1', 'B2']},
index=['K0', 'K1', 'K2'])
data2=pd.DataFrame({'C': ['C0', 'C2', 'C3'],
'D': ['D0', 'D2', 'D3']},
index=['K0', 'K2', 'K3'])
print(data1)
print(data2)
print("final after merging data:")
final=data1.join(data2)
print(final)
#Write a Pandas program to merge two given dataframes with different columns.
import pandas as pd
data1 = pd.DataFrame({'key1': ['K0', 'K0', 'K1', 'K2'],
'key2': ['K0', 'K1', 'K0', 'K1'],
'P': ['P0', 'P1', 'P2', 'P3'],
'Q': ['Q0', 'Q1', 'Q2', 'Q3']})
data2 = pd.DataFrame({'key1': ['K0', 'K1', 'K1', 'K2'],
'key2': ['K0', 'K0', 'K0', 'K0'],
'R': ['R0', 'R1', 'R2', 'R3'],
'S': ['S0', 'S1', 'S2', 'S3']})
print(data1)
print(data2)
final=pd.concat([data1,data2],ignore_index=True)
print(final)
| Pandas Joining and merging DataFrame.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Auto Accidents(1975-2017)
# ## Import cuxfilter
import cuxfilter
from cuxfilter import charts
from cuxfilter.layouts import feature_and_base
#update data_dir if you have downloaded datasets elsewhere
DATA_DIR = './data'
# ## Download required datasets
from cuxfilter.sampledata import datasets_check
datasets_check('auto_accidents', base_dir=DATA_DIR)
cux_df = cuxfilter.DataFrame.from_arrow('./data/auto_accidents.arrow')
cux_df.data['ST_CASE'] = cux_df.data['ST_CASE'].astype('float64')
# ## Define Charts
# +
# add mappings for day of week
label_map = {
1: 'Sunday',
2: 'Monday',
3: 'Tuesday',
4: 'Wednesday',
5: 'Thursday',
6: 'Friday',
7: 'Saturday',
9: 'Unknown'
}
gtc_demo_red_blue_palette = [ "#3182bd", "#6baed6", "#7b8ed8", "#e26798", "#ff0068" , "#323232" ]
# -
from bokeh.tile_providers import get_provider as gp
tile_provider = gp('CARTODBPOSITRON')
# Uncomment the below lines and replace MAPBOX_TOKEN with mapbox token string if you want to use mapbox map-tiles. Can be created for free here -https://www.mapbox.com/help/define-access-token/
# +
#from cuxfilter.assets.custom_tiles import get_provider, Vendors
#tile_provider = get_provider(Vendors.MAPBOX_LIGHT, access_token=MAPBOX_TOKEN)
# +
chart1 = charts.scatter(x='dropoff_x', y='dropoff_y', aggregate_col='DAY_WEEK', aggregate_fn='mean',
tile_provider=tile_provider,
color_palette=gtc_demo_red_blue_palette,pixel_shade_type='linear')
chart2 = charts.bar('YEAR')
chart3 = charts.multi_select('DAY_WEEK', label_map=label_map)
# -
# ## Create a dashboard object
d = cux_df.dashboard([chart1, chart2], sidebar=[chart3], layout=cuxfilter.layouts.feature_and_base,theme = cuxfilter.themes.light, title='Auto Accident Dataset')
#dashboard object
d
# ## Starting the dashboard
# 1. d.show('current_notebook_url:current_notebook_port') remote dashboard
#
# 2. d.app('current_notebook_url:current_notebook_port') inline in notebook (layout is ignored, and charts are displayed one below another)
#
# Incase you need to stop the server:
#
# - d.stop()
await d.preview()
# ## Export the queried data into a dataframe
queried_df = d.export()
| docs/source/examples/auto_accidents_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import des librairies ##
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import re as re
import matplotlib.pyplot as plt
from sklearn.model_selection import StratifiedShuffleSplit
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import regularizers
from tensorflow.keras.callbacks import History,LearningRateScheduler
from tensorflow.keras.layers import Dropout,Dense,Activation
print('TensorFlow %s, Keras %s, numpy %s, pandas %s'%(tf.__version__,keras.__version__, np.__version__,pd.__version__))
__DEBUG__=False
# -
# ## Paramètres ##
# +
## Structure du réseau et nombre d'epochs (nombre de fois où on passe sur le DataSet)
num_hidden_layers=4
first_layer_size = 128
other_layer_size = 512
epochs=200
###Valeurs A tester dans la cross validation
lst_init_learning_rate = [0.01,0.003, 0.1]
lst_dropout_prob=[0.15,0.05]
n_splits=10
# -
# ## Fonctions ##
# +
#Calcule les valeurs min/max et moyennes de chaque colonne dans lst_cols du dataframe pandas df
def get_columns_metadata(df, lst_cols):
header_df = pd.DataFrame( data = lst_cols, columns=['var_name'])
header_df['mean']=df[lst_cols].mean().values
header_df['min']= df[lst_cols].min().values
header_df['max']= df[lst_cols].max().values
header_df.set_index('var_name',inplace=True)
return header_df
#Normalisation de chaque colonne du dataframe pandas df en utilisant les valeurs de header_df
def normalize(df,header_df):
for col in df.columns:
if col in header_df.index :
valmin = header_df["min"][col]
valmax = header_df["max"][col]
valmoy = header_df["mean"][col]
df[col] = 2*((df[col] - valmoy)/(valmax - valmin))
# + [markdown] _cell_guid="25b1e1db-8bc5-7029-f719-91da523bd121" _uuid="5c867fcbb300bcf3c9b8986bba9949da2a2df931"
# ## lecture des données ##
# + _cell_guid="2ce68358-02ec-556d-ba88-e773a50bc18b" _uuid="5ec0878acc5c7ab3903410e671c2a2c6cfeafeea"
# La fonction pandas pd.read_csv permet de créer un objet Dataframe à partir d'un csv
# Données avec labels
train = pd.read_csv('Data/passagers.csv', header = 0, dtype={'Age': np.float64})
# Données de tests sans label. Les prédictions de survie seront envoyées à kaggle
test = pd.read_csv('Data/test.csv' , header = 0, dtype={'Age': np.float64})
# On réunit les données dans une liste (pour pouvoir boucler sur les 2 dataframes)
full_data = [train, test]
#On garde les passagers ID des données test, car on en aura besoin pour le fichiers résultats de kaggle (voir l'exemple gender_submission.csv)
finalfile_index=test.PassengerId #Index des données de test pour le résultat final
#La fonction info() permet de répérer les colonnes avec des valeurs nulles
print('\nTrain data:')
train.info()
print('\nTest data:')
test.info()
# + [markdown] _cell_guid="f9595646-65c9-6fc4-395f-0befc4d122ce" _uuid="66273d64a2548d7a88464ab2a73dbdedfbdc488b"
# # Analyse des données #
# + [markdown] _cell_guid="9b4c278b-aaca-e92c-ba77-b9b48379d1f1" _uuid="c2b62e14d493c270ec8df9f3af1938c479361ef3"
# ## 1. Pclass ##
# Impact de la classe sur la Survie.
# + _cell_guid="4680d950-cf7d-a6ae-e813-535e2247d88e" _uuid="f02533e7b85bba0cca7fcf2cc598c8da92d7646d"
print (train[['Pclass', 'Survived']].groupby(['Pclass'], as_index=False).mean())
# + [markdown] _cell_guid="5e70f81c-d4e2-1823-f0ba-a7c9b46984ff" _uuid="3ca2394409e52b8d6c40d13b6ce557c85e4cd4fd"
# ## 2. Sex ##
# Impact du genre sur la Survie.
# + _cell_guid="6729681d-7915-1631-78d2-ddf3c35a424c" _uuid="2b50b53008fa018127b9d9ee2fb519347b22edcc"
print (train[["Sex", "Survived"]].groupby(['Sex'], as_index=False).mean())
# + [markdown] _cell_guid="7c58b7ee-d6a1-0cc9-2346-81c47846a54a" _uuid="88185e9222c26d5d23caaeb209c18710a231b5f9"
# ## 3. SibSp and Parch ##
# Impacte de la taille de la famille.
# + _cell_guid="1a537f10-7cec-d0b7-8a34-fa9975655190" _uuid="ccc4a4cf7624dd4be450fd62ca3ad478d4e75696"
for dataset in full_data:
dataset['FamilySize'] = dataset['SibSp'] + dataset['Parch'] + 1
print (train[['FamilySize', 'Survived']].groupby(['FamilySize'], as_index=False).mean())
# + [markdown] _cell_guid="e4861d3e-10db-1a23-8728-44e4d5251844" _uuid="f8d7354e5c9160a7da108726a752f7dc366cb0aa"
# Introduction d'une distinction sur les personnes seules
# + _cell_guid="8c35e945-c928-e3bc-bd9c-d6ddb287e4c9" _uuid="87f79dc0711c29f39c0db1a4f7a2e8a84c0c7edb"
for dataset in full_data:
dataset['IsAlone'] = 0
dataset.loc[dataset['FamilySize'] == 1, 'IsAlone'] = 1
print (train[['IsAlone', 'Survived']].groupby(['IsAlone'], as_index=False).mean())
# + [markdown] _cell_guid="8aa419c0-6614-7efc-7797-97f4a5158b19" _uuid="dd18a31086cfeca6330b05f83caf3cc02f687253"
# ## 4. Embarked ##
# Impact du Port d'embarquement sur la Survie.
# + _cell_guid="0e70e9af-d7cc-8c40-b7d4-2643889c376d" _uuid="b4f7ccb3df98da6915bda1c7c225b905fc37845b"
for dataset in full_data:
dataset['Embarked'] = dataset['Embarked'].fillna('S')
print (train[['Embarked', 'Survived']].groupby(['Embarked'], as_index=False).mean())
# + [markdown] _cell_guid="e08c9ee8-d6d1-99b7-38bd-f0042c18a5d9" _uuid="bbeb369d4bb1b086fcc3257218fdeeb6bcdb53c0"
# ## 5. Fare ##
# On remplace les valeurs manquantes par la moyenne. Puis on regarde l'impact du prix du ticket
# -
for dataset in full_data:
dataset.loc[dataset.Fare.isnull(), 'Fare'] = train['Fare'].mean()
train['CategoricalFare'] = pd.qcut(train['Fare'], 4)
print (train[['CategoricalFare', 'Survived']].groupby(['CategoricalFare'], as_index=False).mean())
# + [markdown] _cell_guid="ec8d1b22-a95f-9f16-77ab-7b60d2103852" _uuid="7d96d8817432fa25d8acbcb229df0bd0633b75fa"
# ## 6. Age ##
# Pour les valeurs vides, on gnère des ages aléatoires entre (mean - std) and (mean + std).
# Ensuite on analyse l'impact
# + _cell_guid="b90c2870-ce5d-ae0e-a33d-59e35445500e" _uuid="2af2b56d51752be08b84dbb2684466976758faa7"
for dataset in full_data:
age_avg = dataset['Age'].mean() # Calcul de la valeur moyenne
age_std = dataset['Age'].std() # Calcul de l'écart type
age_null_count = dataset['Age'].isnull().sum() # nombre de valuer nulle
#On génère une valeur aléatoire pour chaque valeur nulle, puis on l'arrondit à l'entier
age_null_random_list = np.random.randint(age_avg - age_std, age_avg + age_std, size=age_null_count)
dataset.loc[np.isnan(dataset['Age']),'Age'] = age_null_random_list
dataset['Age'] = dataset['Age'].astype(int)
#Impact de l'age sur le taux de survie
train['CategoricalAge'] = pd.qcut(train['Age'],5)
print (train[['CategoricalAge', 'Survived']].groupby(['CategoricalAge'], as_index=False).mean())
# + [markdown] _cell_guid="68fa2057-e27a-e252-0d1b-869c00a303ba" _uuid="52bcf7b36b8edb12d40f2a1f9e80060b55d91ad3"
# # Mise en Forme des données #
# ### > Remplacement des données textuelles par des données numériques
# ### > Suppressions des colonnes inutiles (sans impact sur la survie ou créées ci-dessus)###
#
# ## ATTENTION : Il faut lancer "Run All Above Selected Cell" dans le menu Run pour pouvoir relancer ce bloc
# + _cell_guid="2502bb70-ce6f-2497-7331-7d1f80521470" _uuid="1aa110c1043f1f43c091a771abc64054a211f784"
for dataset in full_data:
# Traitement variable 'Sex'
dataset['Sex'].replace('female',0,inplace=True )
dataset['Sex'].replace('male',1,inplace=True)
# Traitement variable 'Embarked'
dataset['Embarked'].replace('S',0,inplace=True)
dataset['Embarked'].replace('C',1,inplace=True)
dataset['Embarked'].replace('Q',2,inplace=True)
# Suppression des colonnes inutiles (Traitements différents sur Train et Test => on ne peut pas mettre ces instruction dans la boucle)
drop_elements = ['PassengerId', 'Name', 'Ticket', 'Cabin', 'SibSp','Parch', 'FamilySize']
train = train.drop(drop_elements, axis = 1)
train = train.drop(['CategoricalAge', 'CategoricalFare'], axis = 1)
### N'oubliez pas de mettre à jour la fonction normalize !
header_df=get_columns_metadata(train,list(train.columns.values))
print(header_df)
normalize(train,header_df)
test = test.drop(drop_elements, axis = 1)
normalize(test,header_df)
print('\nTrain data:')
print (train.head(10))
print('\nTest data:')
print (test.head(10))
# -
# ## Création du modèle et initialisation Training ##
# +
def set_model(init_learning_rate,dropout_prob):
#Architecture du réseau
model = keras.Sequential()
model.add(keras.layers.Dense(first_layer_size, activation='relu'))
### Ajouter ici une ligne pour gérer le sur-apprentissage (fait)
model.add(Dense(64, input_dim=64,kernel_regularizer=regularizers.l2(0.01),activity_regularizer=regularizers.l1(0.01)))
#Couches cachées (Hidden Layers)
for i in range(num_hidden_layers):
# Adds a densely-connected layer to the model:
model.add(keras.layers.Dense(other_layer_size, activation='relu'))
### Ajouter ici une ligne pour gérer le sur-apprentissage
model.add(keras.layers.Dropout(dropout_prob))
# Couche de Sortie (avec fonction Softmax):
model.add(keras.layers.Dense(2, activation='softmax'))
global_step = tf.Variable(0, trainable=False)
learning_rate = tf.train.exponential_decay(init_learning_rate, global_step,1000, 0.96, staircase=True)
### Ici vous pouvez essayer différents algos de descentes de gradients
#Définiton de l'optimizer en charge de la Gradient Descent, de la fonction de coût et de la métrique.
model.compile(optimizer=tf.train.GradientDescentOptimizer(learning_rate),#RMSPropOptimizer(learning_rate), #GradientDescentOptimizer(learning_rate),AdamOptimizer
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
# -
# ## Vérification du Sur-Apprentissage ##
# +
###Essayez Différents jeu de paramètre pour réduire le sur-appentissage
init_learning_rate=0.003
dropout_prob= 0.15
check_epochs=200
pourcentage_validation= 0.2
#A partir des données Train, on sépare features (X) et labels "Survived"
lst_col=list(train.columns.values)
lst_col.remove('Survived')
X=train[lst_col]
y=train['Survived']
# On calcule la position de la séparation pour une répartition 80/20
position_validation_data=int(train.shape[0] * (1-pourcentage_validation))
print('position_validation_data=',position_validation_data)
# Construction des Features pour l'apprentissage et la validation. Transformation du Dataframe Pandas en Numpy Array (attendu par Keras)
X_train, X_val = X[lst_col][:position_validation_data].values, X[lst_col][position_validation_data:].values
# Construction des Labels pour l'apprentissage et la validation. Hot Encoding
y_train, y_val = np.transpose([1-y[:position_validation_data], y[:position_validation_data]]), \
np.transpose([1-y[position_validation_data:], y[position_validation_data:]])
#Construction du modèle en appelant la fonction set_model
model = set_model(init_learning_rate,dropout_prob)
#définition d'une fonction History pour récupérer la fonction de coût et la métrique à chaque epoch.
hist = History()
model.fit(X_train, y_train, epochs=check_epochs, batch_size=512,validation_data=(X_val, y_val),verbose=False, callbacks=[hist])
print(hist.history.keys())
plt.rcParams["figure.figsize"] = (40,20)
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax1.plot(hist.history['val_loss'], color= 'g')
ax2.plot(hist.history['loss'], color= 'b')
ax1.set_xlabel('epochs')
ax1.set_ylabel('Validation data Error', color='g')
ax2.set_ylabel('Training Data Error', color='b')
plt.show()
# -
# ## Fonction de cross validation ##
# +
#Pour un modèle donné, on exécute la cross validation en utilisant un objet sss sklearn StratifiedShuffleSplit
def cv_run(model, name, sss):
loop=1
for train_index, test_index in sss.split(X, y):
### A vous de completer les 2 lignes ci-dessous.
### Il faut extraire les données d'apprentissage et de test des données du dataframe train en utilisant les index renvoyé par la fonction split
### Vous pouvez vous inspirer du code du bloc "Vérification du Sur-Apprentissage"
X_train, X_val = X[train_index], X[test_index]
y_train, y_val = np.transpose([1-y[train_index], y[train_index]]), \
np.transpose([1-y[test_index], y[test_index]])
# Apprentissage et évaluation
hist = History()
model.fit(X_train, y_train, epochs=epochs, batch_size=32,validation_data=(X_val, y_val),verbose=False, callbacks=[hist])
[loss, acc] = model.evaluate(X_val, y_val, batch_size=32,verbose=False)
#Ajout de la performance dans les dictionnaires "loss_dict" et "acc_dict"
if name in acc_dict:
acc_dict[name] += acc
loss_dict[name] += loss
else:
acc_dict[name] = acc
loss_dict[name] = loss
#Affichage de l'avancement
print(loop,':',[loss, acc])
loop+=1
# + [markdown] _cell_guid="23b55b45-572b-7276-32e7-8f7a0dcfd25e" _uuid="4caf4fa8b262c029a4f220883b4c95ed3f25c88f"
# ## Hyperparametrage ##
# ### Ce traitement va être long. Commencer par une faible valeur du paramètre epochs
# + _cell_guid="31ded30a-8de4-6507-e7f7-5805a0f1eaf1" _uuid="b745532338e187d58ff5ee6d961d384b2a5f7bf9"
#Données utilisées pour la méthode split de l'objet StratifiedShuffleSplit
X = train.values[0::, 1::]
y = train.values[0::, 0]
#Créatio d'un dictionnaire pour stocker les modèles
model_dict={}
sss = StratifiedShuffleSplit(n_splits=n_splits, test_size=0.1, random_state=0)
#Créatio d'un dataframe pour logger les résultatsc
log_cols = ["Classifier", "Accuracy"]
log = pd.DataFrame(columns=log_cols)
#Boucle sur des valeurs de init_learning_rate et de dropout_prob
for init_learning_rate in lst_init_learning_rate:
for dropout_prob in lst_dropout_prob :
#Initialisation des dictionnaires utilisés dans la cross validation
acc_dict = {}
loss_dict = {}
#Construction du nom du modèle, en fonction des paramètres
name="lr_%s_do_%s"%(init_learning_rate,dropout_prob)
#Création de l'objet modèle
model = set_model(init_learning_rate,dropout_prob)
#Ajout du modèle au dico pour sélectionner le meilleur dans le suivant
model_dict[name]=model
cv_run(model, name, sss)
# Calcul de la performance du modèle comme moyenne pour chaque itération dans cross-validation
for clf in acc_dict:
acc_dict[clf] = acc_dict[clf] / n_splits
log_entry = pd.DataFrame([[clf, acc_dict[clf]]], columns=log_cols)
log = log.append(log_entry)
print (log.values)
# + [markdown] _cell_guid="438585cf-b7ad-73ba-49aa-87688ff21233" _uuid="f9e6b51b6b3c4cf3098bbdf90f984f827c2f7fd1"
# # Prediction #
# Maintenant on utilise le meilleur jeu de paramètre pour faire la prédiction
# +
###A vous de completer les 3 lignes ci-dessous, sans oublier la normalisation !
### Analyser les résultats du bloc précédent pour choisir le meilleur paramètre
best_model = model_dict["lr_0.003_do_0.15"]
X = train.values[0::, 1::]
y = train.values[0::, 0]
y_hot = np.transpose([1-y, y])
#Apprentissage sur toutes les données, avec le modèle sélectionné
best_model.fit(X,y_hot, epochs=epochs, batch_size=32,verbose=False)
print(pd.DataFrame(best_model.evaluate(X, y_hot, batch_size=32,verbose=False),index=model.metrics_names))
#Inférence des données du fichier test et Construction du fichier à envoyer à Kaggle
prediction=best_model.predict(test.values, batch_size=32)
results=pd.DataFrame(np.argmax(prediction,axis=1), index = finalfile_index, columns=['Survived'])
results.to_csv('resultats.csv')
print(results.sum())
results.describe()
| notebooks/tp3_ANN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Generative Adversarial Networks
#
# Generative adversarial networks (GANs) are a powerful approach for
# probabilistic modeling (I. Goodfellow et al., 2014; I. Goodfellow, 2016).
# They posit a deep generative model and they enable fast and accurate
# inferences.
#
# We demonstrate with an example in Edward. A webpage version is available at
# http://edwardlib.org/tutorials/gan.
# +
from __future__ import absolute_import
from __future__ import print_function
from __future__ import division
import edward as ed
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import numpy as np
import os
import tensorflow as tf
from edward.models import Uniform
from observations import mnist
# -
def plot(samples):
fig = plt.figure(figsize=(4, 4))
gs = gridspec.GridSpec(4, 4)
gs.update(wspace=0.05, hspace=0.05)
for i, sample in enumerate(samples):
ax = plt.subplot(gs[i])
plt.axis('off')
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.set_aspect('equal')
plt.imshow(sample.reshape(28, 28), cmap='Greys_r')
return fig
# +
ed.set_seed(42)
data_dir = "/tmp/data"
out_dir = "/tmp/out"
if not os.path.exists(out_dir):
os.makedirs(out_dir)
M = 128 # batch size during training
d = 100 # latent dimension
# -
# ## Data
#
# We use training data from MNIST, which consists of 55,000 $28\times
# 28$ pixel images (LeCun, Bottou, Bengio, & Haffner, 1998). Each image is represented
# as a flattened vector of 784 elements, and each element is a pixel
# intensity between 0 and 1.
#
# 
#
#
# The goal is to build and infer a model that can generate high quality
# images of handwritten digits.
#
# During training we will feed batches of MNIST digits. We instantiate a
# TensorFlow placeholder with a fixed batch size of $M$ images.
#
# We also define a helper function to select the next batch of data
# points from the full set of examples. It keeps track of the current
# batch index and returns the next batch using the function ``next()``.
# We will generate batches from `x_train_generator` during inference.
def generator(array, batch_size):
"""Generate batch with respect to array's first axis."""
start = 0 # pointer to where we are in iteration
while True:
stop = start + batch_size
diff = stop - array.shape[0]
if diff <= 0:
batch = array[start:stop]
start += batch_size
else:
batch = np.concatenate((array[start:], array[:diff]))
start = diff
batch = batch.astype(np.float32) / 255.0 # normalize pixel intensities
batch = np.random.binomial(1, batch) # binarize images
yield batch
(x_train, _), (x_test, _) = mnist(data_dir)
x_train_generator = generator(x_train, M)
x_ph = tf.placeholder(tf.float32, [M, 784])
# ## Model
#
# GANs posit generative models using an implicit mechanism. Given some
# random noise, the data is assumed to be generated by a deterministic
# function of that noise.
#
# Formally, the generative process is
#
# \begin{align*}
# \mathbf{\epsilon} &\sim p(\mathbf{\epsilon}), \\
# \mathbf{x} &= G(\mathbf{\epsilon}; \theta),
# \end{align*}
#
# where $G(\cdot; \theta)$ is a neural network that takes the samples
# $\mathbf{\epsilon}$ as input. The distribution
# $p(\mathbf{\epsilon})$ is interpreted as random noise injected to
# produce stochasticity in a physical system; it is typically a fixed
# uniform or normal distribution with some latent dimensionality.
#
# In Edward, we build the model as follows, using `tf.layers` to
# specify the neural network. It defines a 2-layer fully connected neural
# network and outputs a vector of length $28\times28$ with values in
# $[0,1]$.
# +
def generative_network(eps):
net = tf.layers.dense(eps, 128, activation=tf.nn.relu)
net = tf.layers.dense(net, 784, activation=tf.sigmoid)
return net
with tf.variable_scope("Gen"):
eps = Uniform(tf.zeros([M, d]) - 1.0, tf.ones([M, d]))
x = generative_network(eps)
# -
# We aim to estimate parameters of the generative network such
# that the model best captures the data. (Note in GANs, we are
# interested only in parameter estimation and not inference about any
# latent variables.)
#
# Unfortunately, probability models described above do not admit a tractable
# likelihood. This poses a problem for most inference algorithms, as
# they usually require taking the model's density. Thus we are
# motivated to use "likelihood-free" algorithms
# (<NAME>, Robert, & Ryder, 2012), a class of methods which assume one
# can only sample from the model.
# ## Inference
#
# A key idea in likelihood-free methods is to learn by
# comparison (e.g., Rubin (1984; <NAME>, Rasch, Schölkopf, & Smola, 2012)): by
# analyzing the discrepancy between samples from the model and samples
# from the true data distribution, we have information on where the
# model can be improved in order to generate better samples.
#
# In GANs, a neural network $D(\cdot;\phi)$ makes this comparison,
# known as the discriminator.
# $D(\cdot;\phi)$ takes data $\mathbf{x}$ as input (either
# generations from the model or data points from the data set), and it
# calculates the probability that $\mathbf{x}$ came from the true data.
#
# In Edward, we use the following discriminative network. It is simply a
# feedforward network with one ReLU hidden layer. It returns the
# probability in the logit (unconstrained) scale.
def discriminative_network(x):
"""Outputs probability in logits."""
net = tf.layers.dense(x, 128, activation=tf.nn.relu)
net = tf.layers.dense(net, 1, activation=None)
return net
# Let $p^*(\mathbf{x})$ represent the true data distribution.
# The optimization problem used in GANs is
#
# \begin{equation*}
# \min_\theta \max_\phi~
# \mathbb{E}_{p^*(\mathbf{x})} [ \log D(\mathbf{x}; \phi) ]
# + \mathbb{E}_{p(\mathbf{x}; \theta)} [ \log (1 - D(\mathbf{x}; \phi)) ].
# \end{equation*}
#
# This optimization problem is bilevel: it requires a minima solution
# with respect to generative parameters and a maxima solution with
# respect to discriminative parameters.
# In practice, the algorithm proceeds by iterating gradient updates on
# each. An
# additional heuristic also modifies the objective function for the
# generative model in order to avoid saturation of gradients
# (<NAME>, 2014).
#
# Many sources of intuition exist behind GAN-style training. One, which
# is the original motivation, is based on idea that the two neural
# networks are playing a game. The discriminator tries to best
# distinguish samples away from the generator. The generator tries
# to produce samples that are indistinguishable by the discriminator.
# The goal of training is to reach a Nash equilibrium.
#
# Another source is the idea of casting unsupervised learning as
# supervised learning
# (<NAME>, <NAME>, & Corander, 2014; <NAME> & Hyvärinen, 2010).
# This allows one to leverage the power of classification—a problem that
# in recent years is (relatively speaking) very easy.
#
# A third comes from classical statistics, where the discriminator is
# interpreted as a proxy of the density ratio between the true data
# distribution and the model
# (Mohamed & Lakshminarayanan, 2016; Sugiyama, Suzuki, & Kanamori, 2012). By augmenting an
# original problem that may require the model's density with a
# discriminator (such as maximum likelihood), one can recover the
# original problem when the discriminator is optimal. Furthermore, this
# approximation is very fast, and it justifies GANs from the perspective
# of approximate inference.
#
# In Edward, the GAN algorithm (`GANInference`) simply takes the
# implicit density model on `x` as input, binded to its
# realizations `x_ph`. In addition, a parameterized function
# `discriminator` is provided to distinguish their
# samples.
inference = ed.GANInference(
data={x: x_ph}, discriminator=discriminative_network)
# We'll use ADAM as optimizers for both the generator and discriminator.
# We'll run the algorithm for 15,000 iterations and print progress every
# 1,000 iterations.
# +
optimizer = tf.train.AdamOptimizer()
optimizer_d = tf.train.AdamOptimizer()
inference = ed.GANInference(
data={x: x_ph}, discriminator=discriminative_network)
inference.initialize(
optimizer=optimizer, optimizer_d=optimizer_d,
n_iter=15000, n_print=1000)
# -
# We now form the main loop which trains the GAN. At each iteration, it
# takes a minibatch and updates the parameters according to the
# algorithm. At every 1000 iterations, it will print progress and also
# saves a figure of generated samples from the model.
# +
sess = ed.get_session()
tf.global_variables_initializer().run()
idx = np.random.randint(M, size=16)
i = 0
for t in range(inference.n_iter):
if t % inference.n_print == 0:
samples = sess.run(x)
samples = samples[idx, ]
fig = plot(samples)
plt.savefig(os.path.join(out_dir, '{}.png').format(
str(i).zfill(3)), bbox_inches='tight')
plt.close(fig)
i += 1
x_batch = next(x_train_generator)
info_dict = inference.update(feed_dict={x_ph: x_batch})
inference.print_progress(info_dict)
# -
# Examining convergence of the GAN objective can be meaningless in
# practice. The algorithm is usually run until some other criterion is
# satisfied, such as if the samples look visually okay, or if the GAN
# can capture meaningful parts of the data.
# ## Criticism
#
# Evaluation of GANs remains an open problem---both in criticizing their
# fit to data and in assessing convergence.
# Recent advances have considered alternative objectives and
# heuristics to stabilize training (see also Soumith Chintala's
# [GAN hacks repo](https://github.com/soumith/ganhacks)).
#
# As one approach to criticize the model, we simply look at generated
# images during training. Below we show generations after 14,000
# iterations (that is, 14,000 gradient updates of both the generator and
# the discriminator).
#
# 
#
# The images are meaningful albeit a little blurry. Suggestions for
# further improvements would be to tune the hyperparameters in the
# optimization, to improve the capacity of the discriminative and
# generative networks, and to leverage more prior information (such as
# convolutional architectures).
| notebooks/gan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="65aRJNe5mMKk" colab_type="code" outputId="33acd327-2177-4986-ccf8-2716426c9538" colab={"base_uri": "https://localhost:8080/", "height": 74}
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
plt.style.use('fivethirtyeight')
import warnings
warnings.filterwarnings('ignore')
import missingno as msno
from sklearn.model_selection import train_test_split
from sklearn import metrics
# + id="c_Z_qYY2mMKx" colab_type="code" colab={}
df = pd.read_csv('/content/loan_sample.csv')
# + id="JPqqZ-qamMK3" colab_type="code" outputId="7a575d9f-6c25-4172-d318-90673baca0db" colab={"base_uri": "https://localhost:8080/", "height": 334}
df.head()
# + id="-WxWmXmPmMLB" colab_type="code" outputId="3089c49f-56c9-4956-b9e0-50ee837efeac" colab={"base_uri": "https://localhost:8080/", "height": 108}
df.info()
# + id="yHBx_6MxmMLL" colab_type="code" outputId="6093df21-112f-4dfe-e904-1694a082c54b" colab={"base_uri": "https://localhost:8080/", "height": 235}
df.dtypes
# + id="BUu5kexkmMLR" colab_type="code" outputId="8e443fee-8063-4352-df15-5d170b519c46" colab={"base_uri": "https://localhost:8080/", "height": 35}
df.shape
# + id="vF2k1TI1mMLX" colab_type="code" outputId="44c2cc3e-03f3-43e2-9ce5-45dfbbcd5d30" colab={"base_uri": "https://localhost:8080/", "height": 235}
df.isna().mean()
# + [markdown] id="SAf1rBmdmMLe" colab_type="text"
# ### Dropping columns with 80% NaN
# + id="3rx_kDpomMLg" colab_type="code" colab={}
#axis=1 specifies column, inplace=True overwrites df
df.dropna(thresh=(df.shape[0] * 0.8), axis=1, inplace=True)
# + id="tmP5RnVYmMLl" colab_type="code" outputId="cca89402-f7bc-4ae5-f6f3-89be428d27eb" colab={"base_uri": "https://localhost:8080/", "height": 35}
# we can now see how many columns have been thrashed
df.shape
# + id="LtUqEBYsmMLr" colab_type="code" outputId="2a8621b6-b546-4ef3-ffad-2224d5840650" colab={"base_uri": "https://localhost:8080/", "height": 446}
# visualizing the missingness partern
msno.matrix(df[0:100000])
# + id="TgQ_8UuymMLy" colab_type="code" outputId="d9b496f9-57c6-491d-99b7-fff47f910eb3" colab={"base_uri": "https://localhost:8080/", "height": 35}
df.dtypes.unique()
# + id="bnMNxRYGmML6" colab_type="code" colab={}
int_df = df.select_dtypes(include=['float64','int64'])
# + id="fTcsCVa3mMMC" colab_type="code" outputId="43081aff-1696-4d37-c4a4-a44d4d7a9fc1" colab={"base_uri": "https://localhost:8080/", "height": 218}
int_df.head()
# + id="9ZUdsRygmMMT" colab_type="code" outputId="79a529f6-f407-47b7-bdea-fa8cbc4ad336" colab={"base_uri": "https://localhost:8080/", "height": 978}
np.absolute(int_df.corr()[['loan_status']]).sort_values(by='loan_status')[35:]
# + id="-kzZISRMmMMf" colab_type="code" outputId="fd1d7be4-9113-46f0-b43e-df6530e4da34" colab={"base_uri": "https://localhost:8080/", "height": 199}
np.absolute(int_df.corr()[['loan_status']]).sort_values(by='loan_status')[35:].index
# + id="qBsTdhrDmMMk" colab_type="code" colab={}
cat_df = df.select_dtypes(include=['O'])
# + id="6uyjXz5DmMMo" colab_type="code" outputId="43dcd8f6-a0ee-4625-b4b4-24ec21cd6271" colab={"base_uri": "https://localhost:8080/", "height": 305}
cat_df.head()
# + id="LSbmbzBymMMt" colab_type="code" outputId="c1b8aa18-e28d-4ff5-d4b3-57e6a74cab68" colab={"base_uri": "https://localhost:8080/", "height": 35}
cat_df.shape
# + id="dKnelV_vmMMz" colab_type="code" outputId="98c42a25-43cd-418b-e4bc-e81a6a100295" colab={"base_uri": "https://localhost:8080/", "height": 35}
cat_df.grade.unique()
# + id="xhGXSbxCmMM9" colab_type="code" outputId="e0056e65-54a2-4d22-dd35-d3c9ac2f2d5e" colab={"base_uri": "https://localhost:8080/", "height": 145}
cat_df.columns
# + id="CtlmlipJ3sOy" colab_type="code" colab={}
# Encoding grade as a predictor variable
encode = {'A': 1, 'B': 2, 'C': 3, 'D': 4, 'E': 5, 'F': 6,'G': 7}
cat_df['grade'].replace(encode, inplace=True)
# adding encoded grade feature to int_df dataframe for modeling
int_df['grade'] = cat_df['grade'].copy()
# + [markdown] id="9gqhQP1emMNF" colab_type="text"
# ## Modeling with continuous predictor variables
# + id="xIK4OLCumMNG" colab_type="code" outputId="7fa8a12e-6fea-4b5f-f492-0213fb79f1ef" colab={"base_uri": "https://localhost:8080/", "height": 322}
# plotting the distribution of the target variable
sns.countplot(df['loan_status'])
# + id="RYV41USsFY5y" colab_type="code" colab={}
# filling missing values with IterativeImputer
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
imp = IterativeImputer(max_iter=1, verbose=0)
imputed_df = imp.fit_transform(int_df)
int_df = pd.DataFrame(imputed_df, columns=int_df.columns)
# + id="bQkp3dA0KX9v" colab_type="code" colab={}
int_df['loan_status'] = df['loan_status'].copy()
# + id="Ilh7dM77KEmF" colab_type="code" outputId="5965953d-b45a-4c63-b328-dc5424aa075b" colab={"base_uri": "https://localhost:8080/", "height": 235}
# finding the sum of null values in the df after Iterative imputation
int_df.isna().sum()
# + id="FjUh5RceIjgu" colab_type="code" colab={}
from google.colab import files
int_df.to_csv('int_df.csv')
files.download('int_df.csv')
# + [markdown] id="Mmx16ZeyMq8U" colab_type="text"
# ## Model Feature Selection
# + id="vNxQ5ZMnznQN" colab_type="code" colab={}
X_test_df = int_df[['total_rec_late_fee', 'int_rate', 'total_pymnt_inv', 'total_pymnt', 'grade', 'last_pymnt_amnt', 'total_rec_prncp',
'collection_recovery_fee', 'recoveries', 'loan_status']].copy()
# + id="8g8oU7qGxihW" colab_type="code" outputId="ae4ae2a4-0ce7-4eb2-ba2e-ff8ca49e0f18" colab={"base_uri": "https://localhost:8080/", "height": 456}
# checking for little or no multicollinearity between good predictor variables
sns.heatmap(X_test_df.corr(), annot=True)
# + id="sc8fS1M9Melj" colab_type="code" colab={}
# feature selection while dealing with multicollinearity
X = int_df[['total_rec_int', 'mths_since_recent_inq', 'total_rev_hi_lim','revol_util', 'inq_last_6mths','funded_amnt_inv','loan_amnt','funded_amnt',
'bc_util', 'percent_bc_gt_75', 'num_rev_tl_bal_gt_0','num_actv_rev_tl', 'tot_cur_bal', 'total_bc_limit', 'mort_acc',
'tot_hi_cred_lim', 'avg_cur_bal', 'bc_open_to_buy','num_tl_op_past_12m', 'dti', 'acc_open_past_24mths', 'total_rec_late_fee',
'int_rate','total_rec_prncp','recoveries']]
# Dropped columns as a result of multicollinearity: 'collection_recovery_fee','total_pymnt_inv','total_pymnt','grade','last_pymnt_amnt'
y = int_df['loan_status']
# + id="2rsZ8X-NmMNY" colab_type="code" outputId="0d8145d3-5c37-4d18-c9ca-95bbd77b1c54" colab={"base_uri": "https://localhost:8080/", "height": 308}
# values to be scaled later
X.describe()
# + id="haU3oQN0mMNe" colab_type="code" colab={}
# splitting data into train and tests sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=123, stratify=y)
# + [markdown] id="pFUG4Ictc05H" colab_type="text"
# ### Logistic Regression
# + id="1KZKMs71mMNk" colab_type="code" colab={}
logreg_model_1 = LogisticRegression()
logreg_model_1.fit(X_train,y_train)
#y_pred_train = logreg_model_1.predict(X_train)
y_pred_test = logreg_model_1.predict(X_test)
# + id="1H_-EXbEmMNt" colab_type="code" outputId="dac058bd-8dfa-4e95-bb31-c7947634fb12" colab={"base_uri": "https://localhost:8080/", "height": 35}
metrics.accuracy_score(y_test, y_pred_test)
# + id="HFcmr7vTmMNy" colab_type="code" outputId="18fe986f-49b2-49cf-ac63-17c040a30d2d" colab={"base_uri": "https://localhost:8080/", "height": 54}
metrics.confusion_matrix(y_test, y_pred_test)
# + id="SoJAjcdlmMN7" colab_type="code" outputId="7a9d628f-cade-4061-9d24-334ba4178c56" colab={"base_uri": "https://localhost:8080/", "height": 181}
print(metrics.classification_report(y_test, y_pred_test))
# + id="FPY4xKo4YtOB" colab_type="code" colab={}
y_pred_prob = logreg_model_1.predict_proba(X_test)[:,1]
# + id="DWcMJ2JCWrqi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a172098e-25dc-4418-96fd-ae2fc31c964c"
metrics.roc_auc_score(y_test, y_pred_prob)
# + [markdown] id="Bm75oE9wfHuR" colab_type="text"
# ### Support Vector Machine Classifier
# + colab_type="code" id="uYd0cVzpE2sI" colab={}
#svm_model_1 = SVC(kernel='linear')
#svm_model_1.fit(X_train,y_train)
#y_pred_train = svm_model_1.predict(X_train)
#y_pred_test = svm_model_1.predict(X_test)
# + id="Pn5UzJqGTpx0" colab_type="code" colab={}
#metrics.accuracy_score(y_test, y_pred_test)
# + id="SS5Qdxp9mMPO" colab_type="code" colab={}
#metrics.confusion_matrix(y_test, y_pred_test)
# + colab_type="code" id="g8oYUQMRE1PF" colab={}
#print(metrics.classification_report(y_test, y_pred_test))
# + [markdown] id="mcWBAssDT2bi" colab_type="text"
# ### RandomForestClassifier
# + id="Uqhc6Y3BUAcU" colab_type="code" colab={}
rf_model_1 = RandomForestClassifier()
rf_model_1.fit(X_train,y_train)
#y_pred_train = rf_model_1.predict(X_train)
y_pred_test = rf_model_1.predict(X_test)
# + id="pNkkx5oJU2Ub" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="db55cec0-7520-43cc-bf41-f74af6141423"
metrics.accuracy_score(y_test, y_pred_test)
# + id="xycpVzZBVYFX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="3818ba76-ebf5-457d-9592-d73c02eb1bf5"
metrics.confusion_matrix(y_test, y_pred_test)
# + id="1IDV-WoDVdvd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 181} outputId="dad9e868-bd5b-4f79-cb30-43524b0c2cbc"
print(metrics.classification_report(y_test, y_pred_test))
# + id="00w5u6msVkCW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="aff107d9-14ba-4d50-d674-dc44b9bdb77f"
y_pred_prob = rf_model_1.predict_proba(X_test)[:,1]
metrics.roc_auc_score(y_test, y_pred_prob)
# + [markdown] id="G2VTiNJXbwuZ" colab_type="text"
# ## MinMaxScaler
# * Scaling features and remodeling
# + id="CZhEm5Qzb7oE" colab_type="code" colab={}
from sklearn.preprocessing import MinMaxScaler, StandardScaler
# + id="dA0OklnxdUPZ" colab_type="code" colab={}
mm = MinMaxScaler()
feature_names = X_train.columns
X_train_mm = mm.fit_transform(X_train)
X_train_mm = pd.DataFrame(X_train_mm, columns=feature_names)
X_test_mm = mm.transform(X_test)
X_test_mm = pd.DataFrame(X_test_mm, columns=feature_names)
# + id="ZY2kUgSvi2Rk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 308} outputId="68ba83fe-5639-4d09-9374-f73928423c58"
# showing a DESC of the normalized X_train features
X_train_mm.describe()
# + [markdown] colab_type="text" id="9ybzawEvsUGX"
# ### Logistic Regression
# + colab_type="code" id="BIV0kTF0sUGh" colab={}
logreg_model_2 = LogisticRegression()
logreg_model_2.fit(X_train_mm,y_train)
#y_pred_train = logreg_model_2.predict(X_train_mm)
y_pred_test = logreg_model_2.predict(X_test_mm)
# + colab_type="code" outputId="46edd277-6329-4e1e-fb31-f4307a3720b0" id="MX4NCR6fsUGw" colab={"base_uri": "https://localhost:8080/", "height": 35}
metrics.accuracy_score(y_test, y_pred_test)
# + colab_type="code" outputId="446a4792-80e4-400c-dd00-d8dc43e0913c" id="lv7L6o4fsUHF" colab={"base_uri": "https://localhost:8080/", "height": 54}
metrics.confusion_matrix(y_test, y_pred_test)
# + colab_type="code" outputId="c6147e06-28de-406b-d4be-7e27df50a935" id="OaL90KIHsUHQ" colab={"base_uri": "https://localhost:8080/", "height": 181}
print(metrics.classification_report(y_test, y_pred_test))
# + colab_type="code" id="vFUQL001sUHc" colab={}
y_pred_prob = logreg_model_2.predict_proba(X_test_mm)[:,1]
# + colab_type="code" outputId="19f794e0-4ecd-468a-f4a2-09eed0e017b0" id="MT1BONaSsUHn" colab={"base_uri": "https://localhost:8080/", "height": 35}
metrics.roc_auc_score(y_test, y_pred_prob)
# + [markdown] colab_type="text" id="kAWLcKaBuN7x"
# ### Support Vector Machine Classifier
# + colab_type="code" id="hvGCOzYFuN72" colab={}
#svm_model_2 = SVC(kernel='linear')
#svm_model_2.fit(X_train_mm,y_train)
#y_pred_train = svm_model_2.predict(X_train_mm)
#y_pred_test = svm_model_2.predict(X_test_mm)
# + colab_type="code" id="LJL0rGLNuN8F" colab={}
#metrics.accuracy_score(y_test, y_pred_test)
# + colab_type="code" id="BkrqjBwGuN8S" colab={}
#metrics.confusion_matrix(y_test, y_pred_test)
# + colab_type="code" id="twYOr-mLuN88" colab={}
#print(metrics.classification_report(y_test, y_pred_test))
# + [markdown] colab_type="text" id="r8xFNqL_v8i0"
# ### RandomForestClassifier
# + colab_type="code" id="Nwvx2Oftv8i-" colab={}
rf_model_2 = RandomForestClassifier()
rf_model_2.fit(X_train_mm,y_train)
#y_pred_train = rf_model_2.predict(X_train_mm)
y_pred_test = rf_model_2.predict(X_test_mm)
# + colab_type="code" outputId="7df06c6f-1999-486e-8196-2994e235f7de" id="ZC7p6f1Lv8jN" colab={"base_uri": "https://localhost:8080/", "height": 35}
metrics.accuracy_score(y_test, y_pred_test)
# + colab_type="code" outputId="40b470e5-962a-4386-e587-07c780a7ba54" id="r_JfTKlTv8jh" colab={"base_uri": "https://localhost:8080/", "height": 54}
metrics.confusion_matrix(y_test, y_pred_test)
# + colab_type="code" outputId="c4775091-a9a9-4c04-edc3-f1f1b68f9eda" id="nc87yp0Hv8ju" colab={"base_uri": "https://localhost:8080/", "height": 181}
print(metrics.classification_report(y_test, y_pred_test))
# + colab_type="code" outputId="f01a2a2c-1ca0-465c-9707-7c2f55c1caf9" id="fWwUr8Psv8kB" colab={"base_uri": "https://localhost:8080/", "height": 35}
y_pred_prob = rf_model_2.predict_proba(X_test_mm)[:,1]
metrics.roc_auc_score(y_test, y_pred_prob)
# + [markdown] id="XNZLb2cGx4uK" colab_type="text"
# ### StandardScaler
# + id="JfrvtY7CxwgY" colab_type="code" colab={}
sc = StandardScaler()
X_train_sc = sc.fit_transform(X_train)
X_train_sc = pd.DataFrame(X_train_sc, columns=feature_names)
X_test_sc = sc.transform(X_test)
X_test_sc = pd.DataFrame(X_test_sc, columns=feature_names)
# + id="g0hbvOgiyyk4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 395} outputId="e01d8f2d-b880-4897-d468-39648ef7223f"
X_train_sc.describe()
# + [markdown] colab_type="text" id="-itbjivVy_vd"
# ### Logistic Regression
# + colab_type="code" id="_rt2ZHdAy_vp" colab={}
logreg_model_3 = LogisticRegression()
logreg_model_3.fit(X_train_sc,y_train)
#y_pred_train = logreg_model_3.predict(X_train_sc)
y_pred_test = logreg_model_3.predict(X_test_sc)
# + colab_type="code" outputId="5f964127-35ad-4599-d420-20dbd09f047a" id="_PAGJxVay_v4" colab={"base_uri": "https://localhost:8080/", "height": 35}
metrics.accuracy_score(y_test, y_pred_test)
# + colab_type="code" outputId="17dacb90-919c-4813-f9e0-65327d70f3a3" id="MK-R6Ugsy_wE" colab={"base_uri": "https://localhost:8080/", "height": 54}
metrics.confusion_matrix(y_test, y_pred_test)
# + colab_type="code" outputId="a1be50d2-90bd-4a24-f52c-d8ca557e5a7c" id="-FERpZt0y_wS" colab={"base_uri": "https://localhost:8080/", "height": 181}
print(metrics.classification_report(y_test, y_pred_test))
# + colab_type="code" id="d4dDQd2ry_wc" colab={}
y_pred_prob = logreg_model_3.predict_proba(X_test_sc)[:,1]
# + colab_type="code" outputId="e5b904e3-0fa7-47ae-96cd-66a533659c17" id="kCRQmoiUy_wv" colab={"base_uri": "https://localhost:8080/", "height": 35}
metrics.roc_auc_score(y_test, y_pred_prob)
# + [markdown] colab_type="text" id="MbpNzmU-zjVH"
# ### Support Vector Machine Classifier
# + colab_type="code" id="T0YnIxUSzjVP" colab={}
#svm_model_3 = SVC(kernel='linear')
#svm_model_3.fit(X_train_mm,y_train)
#y_pred_train = svm_model_3.predict(X_train_sc)
#y_pred_test = svm_model_3.predict(X_test_sc)
# + colab_type="code" id="uwFTDRC1zjVe" colab={}
#metrics.accuracy_score(y_test, y_pred_test)
# + colab_type="code" id="XpPHTAuDzjVo" colab={}
#metrics.confusion_matrix(y_test, y_pred_test)
# + colab_type="code" id="4XUPtmBvzjV4" colab={}
#print(metrics.classification_report(y_test, y_pred_test))
# + [markdown] colab_type="text" id="FvCTIeZNzvL8"
# ### RandomForestClassifier
# + colab_type="code" id="RDtofIrRzvL_" colab={}
rf_model_3 = RandomForestClassifier()
rf_model_3.fit(X_train_sc,y_train)
#y_pred_train = rf_model_3.predict(X_train_sc)
y_pred_test = rf_model_3.predict(X_test_sc)
# + colab_type="code" outputId="b67df572-93c5-42c0-d002-fae6d19dd997" id="l8FhHdqHzvMM" colab={"base_uri": "https://localhost:8080/", "height": 35}
metrics.accuracy_score(y_test, y_pred_test)
# + colab_type="code" outputId="f7a01c95-7cdf-4e17-9fd2-b0c4de9585ca" id="wMBv_grEzvMa" colab={"base_uri": "https://localhost:8080/", "height": 54}
metrics.confusion_matrix(y_test, y_pred_test)
# + colab_type="code" outputId="3a80fe89-fccb-4544-8eae-727826107809" id="LNaP54gNzvMl" colab={"base_uri": "https://localhost:8080/", "height": 181}
print(metrics.classification_report(y_test, y_pred_test))
# + colab_type="code" outputId="2ac6b76c-09db-4cb1-b02c-932acb30a837" id="p3NyQE2azvMw" colab={"base_uri": "https://localhost:8080/", "height": 35}
y_pred_prob = rf_model_3.predict_proba(X_test_sc)[:,1]
metrics.roc_auc_score(y_test, y_pred_prob)
# + id="BWR_qJto1btI" colab_type="code" colab={}
| Project_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Naive Bayes Modeling
# This notebook will focus on using Gaussian Naive Bayes to model customer churn.
# +
# import libraries
from warnings import filterwarnings
filterwarnings("ignore")
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.naive_bayes import GaussianNB
from sklearn.model_selection import train_test_split, GridSearchCV, cross_val_score
from sklearn.metrics import plot_precision_recall_curve, plot_roc_curve
from imblearn.over_sampling import SMOTENC
from sklearn.feature_selection import SelectFpr, SelectPercentile, SelectKBest
from src.seed import SEED
from src.helper import confmat, praf1
# %matplotlib inline
sns.set(font_scale=1.2)
# +
# load data, split data
train = pd.read_csv("../data/processed/train.csv")
X = train.iloc[:, :-1]
y = train.iloc[:, -1]
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=SEED, stratify=y)
# -
# ## Vanilla Modeling
# +
# create instance, fit data, and make predictions
nb = GaussianNB()
nb.fit(X_train, y_train)
train_pred = nb.predict(X_train)
test_pred = nb.predict(X_test)
# -
# output results
a = praf1(y_train, train_pred, "Training")
b = praf1(y_test, test_pred, "Testing")
pd.concat([a, b])
# confusion matrix
confmat([y_train, y_test], [train_pred, test_pred], ["Training", "Testing"])
plt.show()
# +
# output roc/auc curve
fig, ax = plt.subplots(figsize=(10, 8))
plot_roc_curve(nb, X_train, y_train, name="Training", ax=ax)
plot_roc_curve(nb, X_test, y_test, name="Testing", ax=ax)
line = np.linspace(0, 1)
plt.plot(line, line, "--")
plt.show()
# -
# Our basic model with class imbalance not accounted for, has accuracy below random guessing (50/50 chance) for both our training and testing sets. Although our recall is high, it is at the cost of precision.
#
# Future models should look into removing unnecessary features, and accounting for class imbalance.
# ## Class Imbalance
# To tune our model, we'll first start off by balancing out our classes.
# use SMOTENC to account for class imbalance
sm = SMOTENC(np.arange(19, 69), random_state=SEED, n_jobs=-1)
X_train, y_train = sm.fit_resample(X_train, y_train)
# +
# create instance, fit data, and make predictions
nb1 = GaussianNB()
nb1.fit(X_train, y_train)
train_pred = nb1.predict(X_train)
test_pred = nb1.predict(X_test)
# -
# output results
a = praf1(y_train, train_pred, "Training")
b = praf1(y_test, test_pred, "Testing")
pd.concat([a, b])
# confusion matrix
confmat([y_train, y_test], [train_pred, test_pred], ["Training", "Testing"])
plt.show()
# +
# output roc/auc curve
fig, ax = plt.subplots(figsize=(10, 8))
plot_roc_curve(nb1, X_train, y_train, name="Training", ax=ax)
plot_roc_curve(nb1, X_test, y_test, name="Testing", ax=ax)
line = np.linspace(0, 1)
plt.plot(line, line, "--")
plt.show()
# -
# Adjusting for class imbalance has lead to a model which performs well on training data, but testing data is still lackluster. Our accuracy in both training and testing has adjusted to above 50%, making it better than random chance, however we see low precision and low recall.
# ## Feature Selection
# To expand our model and see if we can make it perform better, we'll look to reducing our feature set. Since we don't have l1 or l2 regularization like in logistic regression it'll take a little more effort to do feature selection.
# The feature selection classes that will be used, use the f_classif function, this function computes the ANOVA F-value for the provided sample between classes. For a classification task such as ours, that means it'll compute the value between the churn classes. It examines each feature individually to determine the strength of the relationship of the feature with the response variable.
# iteratively train models with different feature sets and find train/test scores
scores = []
for feats in np.arange(1, X_train.shape[1] + 1):
selector = SelectKBest(k=feats)
selection = selector.fit_transform(X_train, y_train)
train_score = cross_val_score(GaussianNB(), selection, y_train, scoring="f1", n_jobs=-1).mean()
test_score = cross_val_score(GaussianNB(), selector.transform(X_test), y_test, scoring="f1", n_jobs=-1).mean()
scores.append((feats, train_score, test_score))
# find the optimal feature size with a good testing score
feat_df = pd.DataFrame(scores, columns=["feats", "train_score", "test_score"])
feat_df.sort_values("test_score", ascending=False).head()
# +
# what are the best 6 features
selector = SelectKBest(k=6)
selector.fit(X_train, y_train)
X_train.columns[selector.get_support()]
# -
# adjust our X_train, and X_test
X_train_fs = selector.fit_transform(X_train, y_train)
X_test_fs = selector.transform(X_test)
# +
# fit and predict with our new model
nb_fs = GaussianNB()
nb_fs.fit(X_train_fs, y_train)
train_pred = nb_fs.predict(X_train_fs)
test_pred = nb_fs.predict(X_test_fs)
# -
# output results
a = praf1(y_train, train_pred, "Training")
b = praf1(y_test, test_pred, "Testing")
pd.concat([a, b])
confmat([y_train, y_test], [train_pred, test_pred], ["Training", "Testing"])
# +
# output roc/auc curve
fig, ax = plt.subplots(figsize=(10, 8))
plot_roc_curve(nb_fs, X_train_fs, y_train, name="Training", ax=ax)
plot_roc_curve(nb_fs, X_test_fs, y_test, name="Testing", ax=ax)
line = np.linspace(0, 1)
plt.plot(line, line, "--")
plt.show()
# -
# With just 6 features, we see an immense improvement in our model, our f1 score is now almost .5, and our accuracy in both training and testing shows promising results. Our precision is still lackluster, hitting almost 40% in tests, but our recall is well in the 70% range in tests.
#
# Based on tests our model is still misclassifying about 28% of customers who will soon about to leave (recall), and approximately 60% of the positive predictions are actually incorrect (precision).
| notebooks/05-Skellet0r-naive-bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dev
# language: python
# name: dev
# ---
import os
import pandas as pd
# %matplotlib inline
from nltk.sentiment.vader import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
# # News Headlines Sentiment
#
# Use the news api to pull the latest news articles for bitcoin and ethereum and create a DataFrame of sentiment scores for each coin.
#
# Use descriptive statistics to answer the following questions:
# 1. Which coin had the highest mean positive score?
# 2. Which coin had the highest negative score?
# 3. Which coin had the highest positive score?
# +
# Read your api key environment variable
# +
# Create a newsapi client
# +
# Fetch the Bitcoin news articles
# +
# Fetch the Ethereum news articles
# +
# Create the Bitcoin sentiment scores DataFrame
# +
# Create the ethereum sentiment scores DataFrame
# +
# Describe the Bitcoin Sentiment
# +
# Describe the Ethereum Sentiment
# -
# ### Questions:
#
# Q: Which coin had the highest mean positive score?
#
# A:
#
# Q: Which coin had the highest compound score?
#
# A:
#
# Q. Which coin had the highest positive score?
#
# A:
# ---
# # Tokenizer
#
# In this section, you will use NLTK and Python to tokenize the text for each coin. Be sure to:
# 1. Lowercase each word
# 2. Remove Punctuation
# 3. Remove Stopwords
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer, PorterStemmer
from string import punctuation
import re
# +
# Expand the default stop words list if necessary
# -
# Complete the tokenizer function
def tokenizer(text):
"""Tokenizes text."""
# Create a list of the words
# Convert the words to lowercase
# Remove the punctuation
# Remove the stop words
# Lemmatize Words into root words
return tokens
# +
# Create a new tokens column for bitcoin
# +
# Create a new tokens column for ethereum
# -
# ---
# # NGrams and Frequency Analysis
#
# In this section you will look at the ngrams and word frequency for each coin.
#
# 1. Use NLTK to produce the n-grams for N = 2.
# 2. List the top 10 words for each coin.
from collections import Counter
from nltk import ngrams
# +
# Generate the Bitcoin N-grams where N=2
# +
# Generate the Ethereum N-grams where N=2
# -
# Use the token_count function to generate the top 10 words from each coin
def token_count(tokens, N=10):
"""Returns the top N tokens from the frequency count"""
return Counter(tokens).most_common(N)
# +
# Get the top 10 words for Bitcoin
# +
# Get the top 10 words for Ethereum
# -
# # Word Clouds
#
# In this section, you will generate word clouds for each coin to summarize the news for each coin
from wordcloud import WordCloud
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = [20.0, 10.0]
# +
# Generate the Bitcoin word cloud
# +
# Generate the Ethereum word cloud
# -
# # Named Entity Recognition
#
# In this section, you will build a named entity recognition model for both coins and visualize the tags using SpaCy.
import spacy
from spacy import displacy
# +
# Optional - download a language model for SpaCy
# # !python -m spacy download en_core_web_sm
# -
# Load the spaCy model
nlp = spacy.load('en_core_web_sm')
# ## Bitcoin NER
# +
# Concatenate all of the bitcoin text together
# +
# Run the NER processor on all of the text
# Add a title to the document
# +
# Render the visualization
# +
# List all Entities
# -
# ---
# ## Ethereum NER
# +
# Concatenate all of the bitcoin text together
# +
# Run the NER processor on all of the text
# Add a title to the document
# +
# Render the visualization
# +
# List all Entities
| Starter_Code/crypto_sentiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stellargraph Ensembles for node attribute inference
#
# This notebook demonstrates the use of `stellargraph`'s `Ensemble` class for node attribute inference using the Cora and Pubmed-Diabetes citation datasets.
#
# The `Ensemble` class brings ensemble learning to `stellargraph`'s graph neural network models, e.g., `GraphSAGE` and `GCN`, quantifying prediction variance and potentially improving prediction accuracy.
#
# **References**
#
# 1. Inductive Representation Learning on Large Graphs. <NAME>, <NAME>, and <NAME> arXiv:1706.02216
# [cs.SI], 2017.
#
#
# 2. Semi-Supervised Classification with Graph Convolutional Networks. <NAME>, <NAME>. ICLR 2017. arXiv:1609.02907
#
#
# 3. Graph Attention Networks. <NAME> al. ICLR 2018
# +
import networkx as nx
import pandas as pd
import numpy as np
import itertools
import os
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
import stellargraph as sg
from stellargraph.mapper import GraphSAGENodeGenerator, FullBatchNodeGenerator
from stellargraph.layer import GraphSAGE, GCN, GAT
from stellargraph import globalvar
from stellargraph import Ensemble, BaggingEnsemble
from tensorflow.keras import layers, optimizers, losses, metrics, Model, models
from sklearn import preprocessing, feature_extraction, model_selection
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
def plot_history(history):
def remove_prefix(text, prefix):
return text[text.startswith(prefix) and len(prefix):]
figsize=(6, 4)
c_train = 'b'
c_test = 'g'
metrics = sorted(set([remove_prefix(m, "val_") for m in list(history[0].history.keys())]))
for m in metrics:
# summarize history for metric m
plt.figure(figsize=figsize)
for h in history:
plt.plot(h.history[m], c=c_train)
plt.plot(h.history['val_' + m], c=c_test)
plt.title(m)
plt.ylabel(m)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='best')
# ### Loading the network data
# **Downloading the CORA dataset:**
#
# The dataset used in this demo can be downloaded from https://linqs-data.soe.ucsc.edu/public/lbc/cora.tgz
#
# The following is the description of the dataset:
# > The Cora dataset consists of 2708 scientific publications classified into one of seven classes.
# > The citation network consists of 5429 links. Each publication in the dataset is described by a
# > 0/1-valued word vector indicating the absence/presence of the corresponding word from the dictionary.
# > The dictionary consists of 1433 unique words. The README file in the dataset provides more details.
#
# Download and unzip the cora.tgz file to a location on your computer and set the `data_dir` variable to
# point to the location of the dataset (the directory containing "cora.cites" and "cora.content").
#
# **Downloading the PubMed-Diabetes dataset:**
#
# The dataset used in this demo can be downloaded from https://linqs-data.soe.ucsc.edu/public/Pubmed-Diabetes.tgz
#
# The following is the description of the dataset:
#
# >The Pubmed Diabetes dataset consists of 19717 scientific publications from PubMed database pertaining to diabetes classified into one of three classes. The citation network consists of 44338 links. Each publication in the dataset is described by a TF/IDF weighted word vector from a dictionary which consists of 500 unique words.
#
# Download and unzip the Pubmed-Diabetes.tgz file to a location on your computer.
#
# Set the data_dir variable to point to the location of the processed dataset.
# First, we select the dataset to use, either Cora or Pubmed-Diabetes
use_cora = True # Select the dataset; if False, then Pubmed-Diabetes dataset is used.
if use_cora:
data_dir = os.path.expanduser("~/data/cora")
else:
data_dir = os.path.expanduser("~/data/pubmed/Pubmed-Diabetes/data")
def load_cora(data_dir, largest_cc=False):
g_nx = nx.read_edgelist(path=os.path.expanduser(os.path.join(data_dir, "cora.cites")))
for edge in g_nx.edges(data=True):
edge[2]['label'] = 'cites'
# load the node attribute data
cora_data_location = os.path.expanduser(os.path.join(data_dir, "cora.content"))
node_attr = pd.read_csv(cora_data_location, sep='\t', header=None)
values = { str(row.tolist()[0]): row.tolist()[-1] for _, row in node_attr.iterrows()}
nx.set_node_attributes(g_nx, values, 'subject')
if largest_cc:
# Select the largest connected component. For clarity we ignore isolated
# nodes and subgraphs; having these in the data does not prevent the
# algorithm from running and producing valid results.
g_nx_ccs = (g_nx.subgraph(c).copy() for c in nx.connected_components(g_nx))
g_nx = max(g_nx_ccs, key=len)
print("Largest subgraph statistics: {} nodes, {} edges".format(
g_nx.number_of_nodes(), g_nx.number_of_edges()))
feature_names = ["w_{}".format(ii) for ii in range(1433)]
column_names = feature_names + ["subject"]
node_data = pd.read_csv(os.path.join(data_dir, "cora.content"),
sep="\t", header=None,
names=column_names)
node_data.index = node_data.index.map(str)
node_data = node_data[node_data.index.isin(list(g_nx.nodes()))]
for nid in node_data.index:
g_nx.node[nid][globalvar.TYPE_ATTR_NAME] = "paper" # specify node type
return g_nx, node_data, feature_names
def load_pubmed(data_dir):
edgelist = pd.read_csv(os.path.join(data_dir, 'Pubmed-Diabetes.DIRECTED.cites.tab'),
sep="\t", skiprows=2,
header=None )
edgelist.drop(columns=[0,2], inplace=True)
edgelist.columns = ['source', 'target']
# delete unneccessary prefix
edgelist['source'] = edgelist['source'].map(lambda x: x.lstrip('paper:'))
edgelist['target'] = edgelist['target'].map(lambda x: x.lstrip('paper:'))
edgelist["label"] = "cites" # set the edge type
# Load the graph from the edgelist
g_nx = nx.from_pandas_edgelist(edgelist, edge_attr="label")
# Load the features and subject for each node in the graph
nodes_as_dict = []
with open(os.path.join(os.path.expanduser(data_dir),
"Pubmed-Diabetes.NODE.paper.tab")) as fp:
for line in itertools.islice(fp, 2, None):
line_res = line.split("\t")
pid = line_res[0]
feat_name = ['pid'] + [l.split("=")[0] for l in line_res[1:]][:-1] # delete summary
feat_value = [l.split("=")[1] for l in line_res[1:]][:-1] # delete summary
feat_value = [pid] + [ float(x) for x in feat_value ] # change to numeric from str
row = dict(zip(feat_name, feat_value))
nodes_as_dict.append(row)
# Create a Pandas dataframe holding the node data
node_data = pd.DataFrame(nodes_as_dict)
node_data.fillna(0, inplace=True)
node_data['label'] = node_data['label'].astype(int)
node_data['label'] = node_data['label'].astype(str)
node_data.index = node_data['pid']
node_data.drop(columns=['pid'], inplace=True)
node_data.head()
for nid in node_data.index:
g_nx.node[nid][globalvar.TYPE_ATTR_NAME] = "paper" # specify node type
feature_names = list(node_data.columns)
feature_names.remove("label")
return g_nx, node_data, feature_names
# Load the graph data.
if use_cora:
Gnx, node_data, feature_names = load_cora(data_dir)
else:
Gnx, node_data, feature_names = load_pubmed(data_dir)
# We aim to train a graph-ML model that will predict the "subject" or "label" attribute on the nodes depending on the selected dataset. These subjects are one of 7 or 3 categories for Cora and PubMed-Diabetes respectively:
# Print the class names for the selected dataset
if use_cora:
print(set(node_data["subject"]))
else:
print(set(node_data["label"]))
# ### Splitting the data
# For machine learning we want to take a subset of the nodes for training, and use the rest for validation and testing. We'll use scikit-learn again to do this
# +
node_label = "label"
if use_cora:
node_label = "subject"
train_data, test_data = model_selection.train_test_split(node_data,
train_size=0.2, #140
test_size=None,
stratify=node_data[node_label],
random_state=42)
val_data, test_data = model_selection.train_test_split(test_data,
train_size=0.2, #500,
test_size=None,
stratify=test_data[node_label],
random_state=100)
# -
# ### Converting to numeric arrays
# For our categorical target, we will use one-hot vectors that will be fed into a soft-max Keras layer during training.
# +
target_encoding = feature_extraction.DictVectorizer(sparse=False)
train_targets = target_encoding.fit_transform(train_data[[node_label]].to_dict('records'))
val_targets = target_encoding.transform(val_data[[node_label]].to_dict('records'))
test_targets = target_encoding.transform(test_data[[node_label]].to_dict('records'))
# -
# We now do the same for the node attributes we want to use to predict the subject. These are the feature vectors that the Keras model will use as input.
node_features = node_data[feature_names]
# ### Specify global parameters
#
# Here we specify some parameters that control the type of model we are going to use. For example, we specify the base model type, e.g., GCN, GraphSAGE, etc, and the number of estimators in the ensemble as well as model-specific parameters.
# +
model_type = 'graphsage' # Can be either gcn, gat, or graphsage
use_bagging=True # If True, each model in the ensemble is trained on a bootstrapped sample
# of the given training data; otherwise, the same training data are used
# for training each model.
if model_type == "graphsage":
# For GraphSAGE model
batch_size = 50;
num_samples = [10, 10]
n_estimators = 5 # The number of estimators in the ensemble
n_predictions = 10 # The number of predictions per estimator per query point
epochs = 50 # The number of training epochs
elif model_type == "gcn":
# For GCN model
n_estimators = 5 # The number of estimators in the ensemble
n_predictions = 10 # The number of predictions per estimator per query point
epochs = 50 # The number of training epochs
elif model_type == "gat":
# For GAT model
layer_sizes = [8, train_targets.shape[1]]
attention_heads = 8
n_estimators = 5 # The number of estimators in the ensemble
n_predictions = 10 # The number of predictions per estimator per query point
epochs = 200 # The number of training epochs
# -
# ## Creating the base graph machine learning model in Keras
# Now create a `StellarGraph` object from the `NetworkX` graph and the node features and targets. It is `StellarGraph` objects that we use in this library to perform machine learning tasks on.
G = sg.StellarGraph(Gnx, node_features=node_features)
print(G.info())
# To feed data from the graph to the Keras model we need a generator that feeds data from the graph into the model. The generators are specialized to the model and the learning task.
# For training we use only the training nodes returned from our splitter and the target values. The `shuffle=True` argument is given to the `flow` method to improve training for those generators that support shuffling.
if model_type == 'graphsage':
generator = GraphSAGENodeGenerator(G, batch_size, num_samples)
train_gen = generator.flow(train_data.index, train_targets, shuffle=True)
elif model_type == 'gcn':
generator = FullBatchNodeGenerator(G, method="gcn")
train_gen = generator.flow(train_data.index, train_targets) # does not support shuffle
elif model_type == 'gat':
generator = FullBatchNodeGenerator(G, method="gat")
train_gen = generator.flow(train_data.index, train_targets) # does not support shuffle
len(train_data.index)
# Now we can specify our machine learning model, we need a few more parameters for this but the parameters are model-specific.
if model_type == 'graphsage':
base_model = GraphSAGE(
layer_sizes=[16, 16],
generator=train_gen,
bias=True,
dropout=0.5,
normalize="l2"
)
x_inp, x_out = base_model.node_model(flatten_output=True)
prediction = layers.Dense(units=train_targets.shape[1], activation="softmax")(x_out)
elif model_type == 'gcn':
base_model = GCN(
layer_sizes=[32, train_targets.shape[1]],
generator = generator,
bias=True,
dropout=0.5,
activations=["elu", "softmax"]
)
x_inp, x_out = base_model.node_model()
prediction = x_out
elif model_type == 'gat':
base_model = GAT(
layer_sizes=layer_sizes,
attn_heads=attention_heads,
generator=generator,
bias=True,
in_dropout=0.5,
attn_dropout=0.5,
activations=["elu", "softmax"],
)
x_inp, prediction = base_model.node_model()
# Let's have a look at the shape of the output tensor.
prediction.shape
# ### Create a Keras model and then an Ensemble
# Now let's create the actual Keras model with the graph inputs `x_inp` provided by the `base_model` and outputs being the predictions from the softmax layer.
model = Model(inputs=x_inp, outputs=prediction)
# Next, we create the ensemble model consisting of `n_estimators` models.
#
# We are also going to specify that we want to make `n_predictions` per query point per model. These predictions will differ because of the application of `dropout` and, in the case of ensembling GraphSAGE models, the sampling of node neighbourhoods.
# +
if use_bagging:
model = BaggingEnsemble(model, n_estimators=n_estimators, n_predictions=n_predictions)
else:
model = Ensemble(model, n_estimators=n_estimators, n_predictions=n_predictions)
model.compile(
optimizer=optimizers.Adam(lr=0.005),
loss=losses.categorical_crossentropy,
metrics=["acc"],
)
# -
# The model is of type stellargraph.utils.ensemble.Ensemble but has
# a very similar interface to a Keras model
model
# The ensemble has `n_estimators` models. Let's have a look at the first model's layers.
model.layers(0)
# Train the model, keeping track of its loss and accuracy on the training set, and its performance on the validation set during the training (e.g., for early stopping), and generalization performance of the final model on a held-out test set (we need to create another generator over the test data for this)
val_gen = generator.flow(val_data.index, val_targets)
test_gen = generator.flow(test_data.index, test_targets)
# Note that the amount of time to train the ensemble is linear to `n_estimators`.
#
# Also, we are going to use early stopping by monitoring the accuracy on the validation data and stopping if the accuracy does not increase after 10 training epochs (this is the default grace value specified by the `Ensemble` class but we can set it to a different value by using `model.early_stopping_patience=20` for example.)
if use_bagging:
# When using bootstrap samples to train each model in the ensemble, we must specify
# the IDs of the training nodes (train_data) and their corresponding target values
# (train_targets)
history = model.fit_generator(
generator,
train_data = train_data.index,
train_targets = train_targets,
epochs=epochs,
validation_data=val_gen,
verbose=0,
shuffle=False,
bag_size=None,
use_early_stopping=True, # Enable early stopping
early_stopping_monitor="val_acc",
)
else:
history = model.fit_generator(
train_gen,
epochs=epochs,
validation_data=val_gen,
verbose=0,
shuffle=False,
use_early_stopping=True, # Enable early stopping
early_stopping_monitor="val_acc",
)
plot_history(history)
# Now we have trained the model, let's evaluate it on the test set. Note that the `.evaluate_generator()` method of the `Ensemble` class returns mean and standard deviation of each evaluation metric.
# +
test_metrics_mean, test_metrics_std = model.evaluate_generator(test_gen)
print("\nTest Set Metrics of the trained models:")
for name, m, s in zip(model.metrics_names, test_metrics_mean, test_metrics_std):
print("\t{}: {:0.4f}±{:0.4f}".format(name, m, s))
# -
# ### Making predictions with the model
# Now let's get the predictions for all nodes, using a new generator for all nodes:
all_nodes = node_data.index
all_gen = generator.flow(all_nodes)
all_predictions = model.predict_generator(generator=all_gen)
all_predictions.shape
# For full-batch methods, the batch dimension is 1 so we will remove any singleton dimensions
all_predictions = np.squeeze(all_predictions)
all_predictions.shape
# These predictions will be the output of the softmax layer, so to get final categories we'll use the `inverse_transform` method of our target attribute specifcation to turn these values back to the original categories
# For demonstration, we are going to select one of the nodes in the graph, and plot the ensemble's predictions for that node.
selected_query_point = -1
# The array `all_predictions` has dimensionality $MxKxNxF$ where $M$ is the number of estimators in the ensemble (`n_estimators`); $K$ is the number of predictions per query point per estimator (`n_predictions`); $N$ is the number of query points (`len(all_predictions)`); and $F$ is the output dimensionality of the specified layer determined by the shape of the output layer.
#
# Since we are only interested in the predictions for a single query node, e.g., `selected_query_point`, we are going to slice the array to extract them.
# Select the predictions for the point specified by selected_query_point
qp_predictions = all_predictions[:, :, selected_query_point, :]
# The shape should be n_estimators x n_predictions x size_output_layer
qp_predictions.shape
# Next, to facilitate plotting the predictions using either a density plot or a box plot, we are going to reshape `qp_predictions` to $R\times F$ where $R$ is equal to $M\times K$ as above and $F$ is the output dimensionality of the output layer.
qp_predictions = qp_predictions.reshape(np.product(qp_predictions.shape[0:-1]), qp_predictions.shape[-1])
qp_predictions.shape
inv_subject_mapper = {k: v for k, v in enumerate(target_encoding.feature_names_)}
inv_subject_mapper
# We'd like to assess the ensemble's confidence in its predictions in order to decide if we can trust them or not. Utilising density plots, we can visually inspect the ensemble's distribution of prediction probabilities for a node's label.
#
# This is better demonstrated if the ensemble's base mode is `GraphSAGE` because the predictions of the base model vary most (when compared to GCN and GAT) due to the random sampling of node neighbours during prediction in addition to the inherent stocasticity of the ensemble itself.
#
# If the density plot for the predicted node label is well separated from those of the other labels with little overlap then we can be confident trusting the model's prediction.
if model_type not in ['gcn', 'gat']:
fig, ax = plt.subplots(figsize=(12,6))
for i in range(qp_predictions.shape[1]):
sns.kdeplot(data=qp_predictions[:, i].reshape((-1,)), label=inv_subject_mapper[i])
plt.xlabel("Predicted Probability")
plt.title("Density plots of predicted probabilities for each subject")
# An alternative and possibly more informative view of the distribution of node predictions is a box plot.
fig, ax = plt.subplots(figsize=(12,6))
ax.boxplot(x=qp_predictions)
ax.set_xticklabels(target_encoding.feature_names_)
ax.tick_params(axis='x', rotation=45)
if model_type == "graphsage":
y = np.argmax(target_encoding.transform(node_data[[node_label]].to_dict('records')), axis=1)
elif model_type == "gcn" or model_type == "gat":
y = np.argmax(target_encoding.transform(node_data.reindex(G.nodes())[[node_label]].to_dict('records')), axis=1)
plt.title("Correct "+target_encoding.feature_names_[y[selected_query_point]])
plt.ylabel("Predicted Probability")
plt.xlabel("Subject")
# The above example shows that the ensemble predicts the correct node label with high confidence so we can trust its prediction.
#
# (Note that due to the stochastic nature of training neural network algorithms, the above conclusion may not be valid if you re-run the notebook; however, the general conclusion that the use of ensemble learning can be used to quantify the model's uncertainty about its predictions still holds.)
# ## Node embeddings
#
# Evaluate node embeddings as activations of the output of one of the graph convolutional or aggregation layers in the ensemble model, and visualise them, coloring nodes by their subject label.
#
# You can find the index of the layer of interest by calling the `Ensemble` class's method `layers`, e.g., `model.layers()`.
if model_type == 'graphsage':
# For GraphSAGE, we are going to use the output activations of the second GraphSAGE layer
# as the node embeddings
emb = model.predict_generator(generator=generator,
predict_data=node_data.index,
output_layer=-4) # this selects the output layer
elif model_type == 'gcn' or model_type == 'gat':
# For GCN and GAT, we are going to use the output activations of the first GCN or Graph
# Attention layer as the node embeddings
emb = model.predict_generator(generator=generator,
predict_data=node_data.index,
output_layer=6) # this selects the output layer
# The array `emb` has dimensionality $MxKxNxF$ (or $MxKx1xNxF$for full batch methods) where $M$ is the number of estimators in the ensemble (`n_estimators`); $K$ is the number of predictions per query point per estimator (`n_predictions`); $N$ is the number of query points (`len(node_data.index)`); and $F$ is the output dimensionality of the specified layer determined by the shape of the readout layer as specified above.
emb.shape
emb = np.squeeze(emb)
emb.shape
# Next we are going to average the predictions over the number of models and the number of predictions per query point.
#
# The dimensionality of the array will then be **NxF** where N is the number of points to predict (equal to the number of nodes in the graph for this example) and F is the dimensionality of the embeddings that depends on the output shape of the readout layer as specified above.
#
# Note that we could have achieved the same by specifying `summarise=True` in the call to the method `predict_generator` above.
emb = np.mean(emb, axis=(0,1))
emb.shape
# Project the embeddings to 2d using either TSNE or PCA transform, and visualise, coloring nodes by their subject label
X = emb
if model_type == 'graphsage':
y = np.argmax(target_encoding.transform(node_data[[node_label]].to_dict('records')),
axis=1)
elif model_type == 'gcn' or model_type =='gat':
y = np.argmax(target_encoding.transform(node_data.reindex(G.nodes())[[node_label]].to_dict('records')),
axis=1)
if X.shape[1] > 2:
transform = TSNE # PCA
trans = transform(n_components=2)
emb_transformed = pd.DataFrame(trans.fit_transform(X), index=node_data.index)
emb_transformed['label'] = y
else:
emb_transformed = pd.DataFrame(X, index=node_data.index)
emb_transformed = emb_transformed.rename(columns = {'0':0, '1':1})
emb_transformed['label'] = y
# +
alpha = 0.7
fig, ax = plt.subplots(figsize=(7,7))
ax.scatter(emb_transformed[0], emb_transformed[1], c=emb_transformed['label'].astype("category"),
cmap="jet", alpha=alpha)
ax.set(aspect="equal", xlabel="$X_1$", ylabel="$X_2$")
plt.title('{} visualization of {} embeddings for cora dataset'.format(model_type, transform.__name__))
plt.show()
| demos/ensembles/ensemble-node-classification-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing Data from Multiple Files
# We now have almost everything we need to process all our data files.
# The only thing that's missing is a library with a rather unpleasant name:
import glob
# The `glob` library contains a function, also called `glob`,
# that finds files and directories whose names match a pattern.
# We provide those patterns as strings:
# the character `*` matches zero or more characters,
# while `?` matches any one character.
# We can use this to get the names of all the CSV files in the current directory:
files = glob.glob("inflammation-*.csv")
files.sort()
files
# As these examples show,
# `glob.glob`'s result is a list of file and directory paths in arbitrary order.
# This means we can loop over it
# to do something with each filename in turn.
# In our case,
# the "something" we want to do is generate a set of plots for each file in our inflammation dataset.
# If we want to start by analyzing just the first three files in alphabetical order, we can use the
# `sorted` built-in function to generate a new sorted list from the `glob.glob` output:
# %matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
# +
for f in files:
data=np.loadtxt(f,delimiter=",")
fig= plt.figure(figsize=(9,3))
ax1 = fig.add_subplot(1,3,1)
ax2 = fig.add_subplot(1,3,2)
ax3 = fig.add_subplot(1,3,3)
axes[i,0].set_ylabel()
fig,axes =fig.subplot()
ax1.set_ylabel("average")
ax1.plot(np.mean(data,axis=0))
ax2.set_ylabel("max")
ax2.plot(np.max(data,axis=0))
ax3.set_ylabel("min")
ax3.plot(np.min(data,axis=0))
fig.tight_layout()
plt.show()
# -
# Sure enough,
# the maxima of the first two data sets show exactly the same ramp as the first,
# and their minima show the same staircase structure;
# a different situation has been revealed in the third dataset,
# where the maxima are a bit less regular, but the minima are consistently zero.
#
# <section class="challenge panel panel-success">
# <div class="panel-heading">
# <h2><span class="fa fa-pencil"></span> Challenge: Plotting Differences</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p>Plot the difference between the average of the first dataset
# and the average of the second dataset,
# i.e., the difference between the leftmost plot of the first two figures.</p>
#
# </div>
#
# </section>
#
#
# <section class="solution panel panel-primary">
# <div class="panel-heading">
# <h2><span class="fa fa-eye"></span> Solution</h2>
# </div>
#
# </section>
#
#
# <section class="challenge panel panel-success">
# <div class="panel-heading">
# <h2><span class="fa fa-pencil"></span> Challenge: Generate Composite Statistics</h2>
# </div>
#
#
# <div class="panel-body">
#
# <p>Use each of the files once to generate a dataset containing values averaged over all patients:</p>
#
# </div>
#
# </section>
#
# Then use pyplot to generate average, max, and min for all patients.
#
# <section class="solution panel panel-primary">
# <div class="panel-heading">
# <h2><span class="fa fa-eye"></span> Solution</h2>
# </div>
#
# </section>
#
# ---
# The material in this notebook is derived from the Software Carpentry lessons
# © [Software Carpentry](http://software-carpentry.org/) under the terms
# of the [CC-BY 4.0](https://creativecommons.org/licenses/by/4.0/) license.
| 03-fundamentals-of-python/04-processing-files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import pandas
import numpy
recs = pandas.read_csv('./Project-2/data/recs2015_public_v4.csv')
recs['NGXBTU'] = recs['NGXBTU'].fillna(0)
recs['btu_log'] = numpy.log1p(recs['TOTALBTU'])
# features_categorical = ['CELLAR', # 1 (cellar), 0 (no cellar), -2 (not applicable)
# 'WINDOWS' # 10 (1-2), 20 (3-5), 30 (6-9), 41 (10-15), 42 (16-19), 50 (20-29), 60 (30+)
# ]
# features_numeric = ['BEDROOMS', # Bedroom count
# 'TOTROOMS', # Total room count
# 'TOTSQFT_EN', # Total Square Feet
# 'TOTHSQFT' # Total Heated Square Feet
# ]
# features = features_numeric + features_categorical
potential_targets = ['btu_log',
'TOTALBTU',
'TOTALDOL',
'KWH',
'BTUEL',
'DOLLAREL',
'CUFEETNG',
'BTUNG',
'DOLLARNG',
'GALLONLP',
'BTULP',
'DOLLARLP',
'GALLONFO',
'BTUFO',
'DOLLARFO',
'WOODAMT',
'WOODBTU',
'PELLETAMT',
'PELLETBTU']
not_features = ["btu_log",
"KWH", # Numeric 8 Total site electricity usage, in kilowatthours, 2015
"KWHSPH", # Numeric 8 Electricity usage for space heating, main and secondary, in kilowatthours, 2015
"KWHCOL", # Numeric 8 Electricity usage for air conditioning (central systems and individual units), in kilowatthours, 2015
"KWHWTH", # Numeric 8 Electricity usage for water heating, main and secondary, in kilowatthours, 2015
"KWHRFG", # Numeric 8 Electricity usage for all refrigerators, in kilowatthours, 2015
"KWHRFG1", # Numeric 8 Electricity usage for first refrigerators, in kilowatthours, 2015
"KWHRFG2", # Numeric 8 Electricity usage for second refrigerators, in kilowatthours, 2015
"KWHFRZ", # Numeric 8 Electricity usage for freezers, in kilowatthours, 2015
"KWHCOK", # Numeric 8 Electricity usage for cooking (stoves, cooktops, and ovens), in kilowatthours, 2015
"KWHMICRO", # Numeric 8 Electricity usage for microwaves, in kilowatthours, 2015
"KWHCW", # Numeric 8 Electricity usage for clothes washers, in kilowatthours, 2015
"KWHCDR", # Numeric 8 Electricity usage for clothes dryers, in kilowatthours, 2015
"KWHDWH", # Numeric 8 Electricity usage for dishwashers, in kilowatthours, 2015
"KWHLGT", # Numeric 8 Electricity usage for indoor and outdoor lighting, in kilowatthours, 2015
"KWHTVREL", # Numeric 8 Electricity usage for all televisions and related peripherals, in kilowatthours, 2015
"KWHTV1", # Numeric 8 Electricity usage for first televisions, in kilowatthours, 2015
"KWHTV2", # Numeric 8 Electricity usage for second televisions, in kilowatthours, 2015
"KWHAHUHEAT", # Numeric 8 Electricity usage for air handlers and boiler pumps used for heating, in kilowatthours, 2015
"KWHAHUCOL", # Numeric 8 Electricity usage for air handlers used for cooling, in kilowatthours, 2015
"KWHEVAPCOL", # Numeric 8 Electricity usage for evaporative coolers, in kilowatthours, 2015
"KWHCFAN", # Numeric 8 Electricity usage for ceiling fans, in kilowatthours, 2015
"KWHDHUM", # Numeric 8 Electricity usage for dehumidifiers, in kilowatthours, 2015
"KWHHUM", # Numeric 8 Electricity usage for humidifiers, in kilowatthours, 2015
"KWHPLPMP", # Numeric 8 Electricity usage for swimming pool pumps, in kilowatthours, 2015
"KWHHTBPMP", # Numeric 8 Electricity usage for hot tub pumps, in kilowatthours, 2015
"KWHHTBHEAT", # Numeric 8 Electricity usage for hot tub heaters, in kilowatthours, 2015
"KWHNEC", # Numeric 8 Electricity usage for other devices and purposes not elsewhere classified, in kilowatthours, 2015
"BTUEL", # Numeric 8 Total site electricity usage, in thousand Btu, 2015
"BTUELSPH", # Numeric 8 Electricity usage for space heating, main and secondary, in thousand Btu, 2015
"BTUELCOL", # Numeric 8 Electricity usage for air conditioning (central systems and individual units), in thousand Btu, 2015
"BTUELWTH", # Numeric 8 Electricity usage for water heating, main and secondary, in thousand Btu, 2015
"BTUELRFG", # Numeric 8 Electricity usage for all refrigerators, in thousand Btu, 2015
"BTUELRFG1", # Numeric 8 Electricity usage for first refrigerators, in thousand Btu, 2015
"BTUELRFG2", # Numeric 8 Electricity usage for second refrigerators, in thousand Btu, 2015
"BTUELFRZ", # Numeric 8 Electricity usage for freezers, in thousand Btu, 2015
"BTUELCOK", # Numeric 8 Electricity usage for cooking (stoves, cooktops, and ovens), in thousand Btu, 2015
"BTUELMICRO", # Numeric 8 Electricity usage for microwaves, in thousand Btu, 2015
"BTUELCW", # Numeric 8 Electricity usage for clothes washers, in thousand Btu, 2015
"BTUELCDR", # Numeric 8 Electricity usage for clothes dryers, in thousand Btu, 2015
"BTUELDWH", # Numeric 8 Electricity usage for dishwashers, in thousand Btu, 2015
"BTUELLGT", # Numeric 8 Electricity usage for indoor and outdoor lighting, in thousand Btu, 2015
"BTUELTVREL", # Numeric 8 Electricity usage for all televisions and related peripherals, in thousand Btu, 2015
"BTUELTV1", # Numeric 8 Electricity usage for first televisions, in thousand Btu, 2015
"BTUELTV2", # Numeric 8 Electricity usage for second televisions, in thousand Btu, 2015
"BTUELAHUHEAT", # Numeric 8 Electricity usage for air handlers and boiler pumps used for heating, in thousand Btu, 2015
"BTUELAHUCOL", # Numeric 8 Electricity usage for air handlers used for cooling, in thousand Btu, 2015
"BTUELEVAPCOL", # Numeric 8 Electricity usage for evaporative coolers, in thousand Btu, 2015
"BTUELCFAN", # Numeric 8 Electricity usage for ceiling fans, in thousand Btu, 2015
"BTUELDHUM", # Numeric 8 Electricity usage for dehumidifiers, in thousand Btu, 2015
"BTUELHUM", # Numeric 8 Electricity usage for humidifiers, in thousand Btu, 2015
"BTUELPLPMP", # Numeric 8 Electricity usage for swimming pool pumps, in thousand Btu, 2015
"BTUELHTBPMP", # Numeric 8 Electricity usage for hot tub pumps, in thousand Btu, 2015
"BTUELHTBHEAT", # Numeric 8 Electricity usage for hot tub heaters, in thousand Btu, 2015
"BTUELNEC", # Numeric 8 Electricity usage for other devices and purposes not elsewhere classified, in thousand Btu, 2015
"DOLLAREL", # Numeric 8 Total electricity cost, in dollars, 2015
"DOLELSPH", # Numeric 8 Electricity cost for space heating, main and secondary, in dollars, 2015
"DOLELCOL", # Numeric 8 Electricity cost for air conditioning (central systems and individual units), in dollars, 2015
"DOLELWTH", # Numeric 8 Electricity cost for water heating, main and secondary, in dollars, 2015
"DOLELRFG", # Numeric 8 Electricity cost for all refrigerators, in dollars, 2015
"DOLELRFG1", # Numeric 8 Electricity cost for first refrigerators, in dollars, 2015
"DOLELRFG2", # Numeric 8 Electricity cost for second refrigerators, in dollars, 2015
"DOLELFRZ", # Numeric 8 Electricity cost for freezers, in dollars, 2015
"DOLELCOK", # Numeric 8 Electricity cost for cooking (stoves, cooktops, and ovens), in dollars, 2015
"DOLELMICRO", # Numeric 8 Electricity cost for microwaves, in dollars, 2015
"DOLELCW", # Numeric 8 Electricity cost for clothes washers, in dollars, 2015
"DOLELCDR", # Numeric 8 Electricity cost for clothes dryers, in dollars, 2015
"DOLELDWH", # Numeric 8 Electricity cost for dishwashers, in dollars, 2015
"DOLELLGT", # Numeric 8 Electricity cost for indoor and outdoor lighting, in dollars, 2015
"DOLELTVREL", # Numeric 8 Electricity cost for all televisions and related peripherals, in dollars, 2015
"DOLELTV1", # Numeric 8 Electricity cost for first televisions, in dollars, 2015
"DOLELTV2", # Numeric 8 Electricity cost for second televisions, in dollars, 2015
"DOLELAHUHEAT", # Numeric 8 Electricity cost for air handlers and boiler pumps used for heating, in dollars, 2015
"DOLELAHUCOL", # Numeric 8 Electricity cost for air handlers used for cooling, in dollars, 2015
"DOLELEVAPCOL", # Numeric 8 Electricity cost for evaporative coolers, in dollars, 2015
"DOLELCFAN", # Numeric 8 Electricity cost for ceiling fans, in dollars, 2015
"DOLELDHUM", # Numeric 8 Electricity cost for dehumidifiers, in dollars, 2015
"DOLELHUM", # Numeric 8 Electricity cost for humidifiers, in dollars, 2015
"DOLELPLPMP", # Numeric 8 Electricity cost for swimming pool pumps, in dollars, 2015
"DOLELHTBPMP", # Numeric 8 Electricity cost for hot tub pumps, in dollars, 2015
"DOLELHTBHEAT", # Numeric 8 Electricity cost for hot tub heaters, in dollars, 2015
"DOLELNEC", # Numeric 8 Electricity cost for other devices and purposes not elsewhere classified, in dollars, 2015
"CUFEETNG", # Numeric 8 Total natural gas usage, in hundred cubic feet, 2015
"CUFEETNGSPH", # Numeric 8 Natural gas usage for space heating, main and secondary, in hundred cubic feet, 2015
"CUFEETNGWTH", # Numeric 8 Natural gas usage for water heating , main and secondary, in hundred cubic feet, 2015
"CUFEETNGCOK", # Numeric 8 Natural gas usage for cooking (stoves, cooktops, and ovens), in hundred cubic feet, 2015
"CUFEETNGCDR", # Numeric 8 Natural gas usage for clothes dryers, in hundred cubic feet, 2015
"CUFEETNGPLHEAT", # Numeric 8 Natural gas usage for swimming pool heaters, in hundred cubic feet, 2015
"CUFEETNGHTBHEAT", # Numeric 8 Natural gas usage for hot tub heaters, in hundred cubic feet, 2015
"CUFEETNGNEC", # Numeric 8 Natural gas usage for other devices and purposes not elsewhere classified, in hundred cubic feet, 2015
"BTUNG", # Numeric 8 Total natural gas usage, in thousand Btu, 2015
"BTUNGSPH", # Numeric 8 Natural gas usage for space heating, main and secondary, in thousand Btu, 2015
"BTUNGWTH", # Numeric 8 Natural gas usage for water heating, main and secondary, in thousand Btu, 2015
"BTUNGCOK", # Numeric 8 Natural gas usage for cooking (stoves, cooktops, and ovens), in thousand Btu, 2015
"BTUNGCDR", # Numeric 8 Natural gas usage for clothes dryers, in thousand Btu, 2015
"BTUNGPLHEAT", # Numeric 8 Natural gas usage for swimming pool heaters, in thousand Btu, 2015
"BTUNGHTBHEAT", # Numeric 8 Natural gas usage for hot tub heaters, in thousand Btu, 2015
"BTUNGNEC", # Numeric 8 Natural gas usage for other devices and purposes not elsewhere classified, in thousand Btu, 2015
"DOLLARNG", # Numeric 8 Total natural gas cost, in dollars, 2015
"DOLNGSPH", # Numeric 8 Natural Gas cost for space heating, main and secondary, in dollars, 2015
"DOLNGWTH", # Numeric 8 Natural gas cost for water heating, main and secondary, in dollars, 2015
"DOLNGCOK", # Numeric 8 Natural gas cost for cooking (stoves, cooktops, and ovens), in dollars, 2015
"DOLNGCDR", # Numeric 8 Natural gas cost for clothes dryers, in dollars, 2015
"DOLNGPLHEAT", # Numeric 8 Natural gas cost for swimming pool heaters, in dollars, 2015
"DOLNGHTBHEAT", # Numeric 8 Natural gas cost for hot tub heaters, in dollars, 2015
"DOLNGNEC", # Numeric 8 Natural gas cost for other devices and purposes not elsewhere classified, in dollars, 2015
"GALLONLP", # Numeric 8 Total propane usage, in gallons, 2015
"GALLONLPSPH", # Numeric 8 Propane usage for space heating, main and secondary, in gallons, 2015
"GALLONLPWTH", # Numeric 8 Propane usage for water heating, main and secondary, in gallons, 2015
"GALLONLPCOK", # Numeric 8 Propane usage for cooking (stoves, cooktops, and ovens), in gallons, 2015
"GALLONLPCDR", # Numeric 8 Propane usage for clothes dryers, in gallons, 2015
"GALLONLPNEC", # Numeric 8 Propane usage for other devices and purposes not elsewhere classified, in gallons, 2015
"BTULP", # Numeric 8 Total propane usage, in thousand Btu, 2015
"BTULPSPH", # Numeric 8 Propane usage for space heating, main and secondary, in thousand Btu, 2015
"BTULPWTH", # Numeric 8 Propane usage for water heating, main and secondary, in thousand Btu, 2015
"BTULPCOK", # Numeric 8 Propane usage for cooking (stoves, cooktops, and ovens), in thousand Btu, 2015
"BTULPCDR", # Numeric 8 Propane usage for clothes dryers, in thousand Btu, 2015
"BTULPNEC", # Numeric 8 Propane usage for other devices and purposes not elsewhere classified, in thousand Btu, 2015
"DOLLARLP", # Numeric 8 Total cost of propane, in dollars, 2015
"DOLLPSPH", # Numeric 8 Propane cost for space heating, main and secondary, in dollars, 2015
"DOLLPWTH", # Numeric 8 Propane cost for water heating, main and secondary, in dollars, 2015
"DOLLPCOK", # Numeric 8 Propane cost for cooking (stoves, cooktops, and ovens), in dollars, 2015
"DOLLPCDR", # Numeric 8 Propane cost for clothes dryers, in dollars, 2015
"DOLLPNEC", # Numeric 8 Propane cost for other devices and purposes not elsewhere classified, in dollars, 2015
"GALLONFO", # Numeric 8 Total fuel oil/kerosene usage, in gallons, 2015
"GALLONFOSPH", # Numeric 8 Fuel oil/kerosene usage for space heating, main and secondary, in gallons, 2015
"GALLONFOWTH", # Numeric 8 Fuel oil/kerosene usage for water heating, main and secondary, in gallons, 2015
"GALLONFONEC", # Numeric 8 Fuel oil/kerosene usage for other devices and purposes not elsewhere classified, in gallons, 2015
"BTUFO", # Numeric 8 Total fuel oil/kerosene usage, in thousand Btu, 2015
"BTUFOSPH", # Numeric 8 Fuel oil/kerosene usage for space heating, main and secondary, in thousand Btu, 2015
"BTUFOWTH", # Numeric 8 Fuel oil/kerosene usage for water heating, main and secondary, in thousand Btu, 2015
"BTUFONEC", # Numeric 8 Fuel oil/kerosene usage for other devices and purposes not elsewhere classified, in thousand Btu, 2015
"DOLLARFO", # Numeric 8 Total cost of fuel oil/kerosene, in dollars, 2015
"DOLFOSPH", # Numeric 8 Fuel oil/kerosene cost for space heating, main and secondary, in dollars, 2015
"DOLFOWTH", # Numeric 8 Fuel oil/kerosene cost for water heating, main and secondary, in dollars, 2015
"DOLFONEC", # Numeric 8 Fuel oil/kerosene cost for other devices and purposes not elsewhere classified, in dollars, 2015
"TOTALBTU", # Numeric 8 Total usage, in thousand Btu, 2015
"TOTALDOL", # Numeric 8 Total cost, in dollars, 2015
"TOTALBTUSPH", # Numeric 8 Total usage for space heating, main and secondary, in thousand Btu, 2015
"TOTALDOLSPH", # Numeric 8 Total cost for space heating, main and secondary, in dollars, 2015
"TOTALBTUWTH", # Numeric 8 Total usage for water heating, main and secondary, in thousand Btu, 2015
"TOTALDOLWTH", # Numeric 8 Total cost for water heating, main and secondary, in dollars, 2015
"TOTALBTUCOK", # Numeric 8 Total usage for cooking (stoves, cooktops, and ovens), in thousand Btu, 2015
"TOTALDOLCOK", # Numeric 8 Total cost for cooking (stoves, cooktops, and ovens), in dollars, 2015
"TOTALBTUCDR", # Numeric 8 Total usage for clothes dryers, in thousand Btu, 2015
"TOTALDOLCDR", # Numeric 8 Total cost for clothes dryers, in dollars, 2015
"TOTALBTUPL", # Numeric 8 Total usage for swimming pool pumps and heaters, in thousand Btu, 2015
"TOTALDOLPL", # Numeric 8 Total cost for swimming pool pumps and heaters, in dollars, 2015
"TOTALBTUHTB", # Numeric 8 Total usage for hot tub pumps and heaters, in thousand Btu, 2015
"TOTALDOLHTB", # Numeric 8 Total cost for hot tub pumps and heaters, in dollars, 2015
"TOTALBTUNEC", # Numeric 8 Total usage for other devices and purposes not elsewhere classified, in thousand Btu, 2015
"TOTALDOLNEC", # Numeric 8 Total cost for other devices and purposes not elsewhere classified, in thousand Btu, 2015
"WOODAMT", # Numeric 8 Cords of wood used in the last year
"ZWOODAMT", # Numeric 8 Imputation flag for WOODAMT
"WOODBTU", # Numeric 8 Total cordwood usage, in thousand Btu, 2015 (Wood consumption is not included in TOTALBTU or TOTALDOL)
"PELLETAMT", # Numeric 8 Number of 40-pound wood pellet bags used in the last year
"ZPELLETAMT", # Numeric 8 Imputation flag for PELLETAMT
"PELLETBTU" # Numeric 8 Total wood pellet usage, in thousand Btu, 2015 (Wood consumption is not included in TOTALBTU or TOTALDOL)
]
features = recs.columns.drop(not_features)
weights = ['NWEIGHT']
targets = ['btu_log']
features
# -
uniques = recs[features].nunique()
encode = list(uniques[uniques<=11].index)
# +
from category_encoders import OneHotEncoder
from sklearn.pipeline import Pipeline
oneHot = OneHotEncoder( use_cat_names=True, cols=encode)
features_transformed = oneHot.fit_transform(recs[features])
# -
import numpy
try:
features_transformed.describe(exclude=numpy.number)
except ValueError:
print('No non-numeric cols!')
features_transformed.iloc[0]
# +
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val, w_train, w_val = train_test_split(features_transformed, recs[potential_targets].values, recs[weights], random_state=3)
# +
from sklearn.linear_model import LinearRegression
lr_model = LinearRegression()
lr_model.fit(X_train, y_train)
lr_model.score(X_val, y_val)
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.pipeline import Pipeline
from sklearn.model_selection import RandomizedSearchCV
rfr_model = RandomForestRegressor(n_estimators=170, min_samples_leaf=12, n_jobs=-1, verbose=1)
pipeline = Pipeline([('RandomForestRegressor', rfr_model)])
searchCV = RandomizedSearchCV( pipeline,
param_distributions={},
n_iter=1,
cv=9,
scoring='r2',
verbose=10,
return_train_score=True,
n_jobs=-1)
searchCV.fit(X_train, y_train)
print('best_score_: ', searchCV.best_score_)#.score(X_val, y_val)
print('best_estimator_.score: ', searchCV.best_estimator_.score(X_val, y_val))
# -
searchCV.best_estimator_.predict(X_val.iloc[0:1,:])
# +
y_val[0:1,:]
# -
# y_train[0]
recs[recs['TOTALBTU']==84251.317][potential_targets]
# +
from eli5.sklearn import PermutationImportance
models = {}
feature_importances = {}
for i in range(len(potential_targets)):
target = potential_targets[i]
print('Training on', target)
y_train_t = y_train[:,i]
y_val_t = y_val[:,i]
#n_estimators=120
rfr_model_t = RandomForestRegressor(n_estimators=40, min_samples_leaf=12, n_jobs=-2, verbose=0)
pipeline_t = Pipeline([('RandomForestRegressor', rfr_model)])
searchCV = RandomizedSearchCV( pipeline,
param_distributions={},
n_iter=1,
cv=3,
scoring='r2',
verbose=0,
return_train_score=True,
n_jobs=-2)
searchCV.fit(X_train, y_train_t)
searchCV.best_estimator_.verbose=0
models[target] = searchCV.best_estimator_
print(target, 'score:', models[target].score(X_val, y_val_t))
permuter = PermutationImportance(searchCV.best_estimator_, n_iter=2)
permuter.fit(X_val, y_val_t)
importances = [feature for _,feature in sorted(zip(permuter.feature_importances_,list(X_val.columns)), reverse=True)]
print(target, 'feature importances top 15:', importances[:15])
feature_importances[target] = importances
# +
scores_t = {}
for i in range(len(potential_targets)):
target = potential_targets[i]
y_val_t = y_val[:,i]
scores_t[target] = models[target].score(X_val, y_val_t)
# -
for target in potential_targets:
print(target, 'score:', scores_t[target])
print(target, 'feature importances top 10:', feature_importances[target][:10])
print('TOTALBTU', models['TOTALBTU'].score(X_val, y_val[:,1]))
# +
# from xgboost import XGBRegressor
# from sklearn.multioutput import MultiOutputRegressor
# xgbr_model = XGBRegressor( random_state=1,
# n_jobs=-1,
# n_estimators=4000,
# min_samples_leaf=35,
# # max_depth=9,
# learning_rate=.05,
# verbosity=3)
# eval_set = [(X_train, y_train),
# (X_val, y_val)]
# multioutputregressor = MultiOutputRegressor(xgbr_model)
# # multioutputregressor.set_params(eval_set=eval_set, early_stopping_rounds=50)
# multioutputregressor.fit(X_train, y_train)#, eval_set=eval_set, early_stopping_rounds=50)
# # xgbr_model.fit(X_train, y_train, eval_set=eval_set, early_stopping_rounds=50)
# # xgbr_model.score(X_val, y_val)
# multioutputregressor.score(X_val, y_val)
# -
corr = recs.corr()
for target in potential_targets:
print(corr[target].sort_values().iloc[0:5])
# +
# import seaborn
# import matplotlib.pyplot as pyplot
# pyplot.rcParams['figure.facecolor'] = '#002B36'
# pyplot.rcParams['axes.facecolor'] = 'black'
# pyplot.rcParams['figure.figsize'] = (10,400)
# seaborn.heatmap(corr[potential_targets])
# pyplot.show()
# -
# +
# corr_s = corr.stack()
# corr_s
| Project-2/c.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="e4b1iEVqjUEn"
# # 排序算法
# + [markdown] colab_type="text" id="KoJf4BYLjreT"
# ## 问题定义
#
# **Input** 一个包含$n$个数字的序列 $s=<a_1, a_2, a_3, ..., a_n>$.
#
# **Output**原序列$s$的其中一个排列(permutation)$s'=<a_1', a_2', a_3', ..., a_n'>$, 使得$a_1' \leq a_2' \leq ... \leq a_n'$.
#
# ---
# + [markdown] colab_type="text" id="1VT63sL0tD63"
# ## Heap Sort 堆排序
#
# 时间复杂度: $O(n log n)$
#
# 借助了数据结构 "堆 (Heap)", 所以在将堆排序之前我们需要先了解**堆**的一些基本性质.
# + [markdown] colab_type="text" id="gPg7NOcduRGo"
# ### 堆
#
# 堆(二叉堆 binary heap) 是一种用**近似**完全二叉树来表示的数据结构 (*完全二叉树的意思为除了最底层, 其余层所有节点都有左右两个子节点*).
#
# 而堆也可以用数组来存储, 其中堆的根节点为Heap[1].
#
# 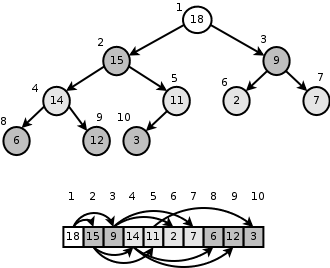
#
# 对于堆中的某个节点$i$, 计算它的父节点和左右节点可以使用以下方法:
#
# $parent(i)=floor({i \over 2})$
#
# $left(i)=2i$
#
# $right(i)=2i+1$
#
# 堆分为两种: **最大堆**和**最小堆**. 对于最大堆来说, 它的特性为:
#
# 任意节点$i \neq 1$, 有$Heap[parent(i)] \geq Heap[i]$. 所以最大堆中最大元素存放在根节点.
#
# 最小堆与最大堆相反, 它的特性为:
#
# 任意节点$i \neq 1$, 有$Heap[parent(i)] \leq Heap[i]$. 所以最小堆中最小元素存放在根节点.
#
#
# +
def left(i):
return i * 2
def right(i):
return i * 2 + 1
def parent(i):
return int(i / 2)
# -
# 对于堆排序, 我们使用最大堆. 由二叉树的定义易知堆的$height=\Theta(log n)$
#
# 下面定义堆的几种操作:
# ### Max-heapify (Sink)
#
# 该操作用于维护最大堆的性质, 它的输入是一个数组$A$和一个下标$i$. 当它被调用时, 假设以$i$节点的两个子节点为根的两棵子树已经满足最大堆的性质, 而节点$i$的值A\[i\]有可能小于它子节点的值, 该情况不满足最大堆的定义. 所以在max-heapify操作中, A\[i\]会往下**下沉**, 完成操作后以$i$为根节点的子树将满足最大堆的性质. 我们将这个操作起名为下沉 (sink).
# **举例** sink(a, 2)的过程模拟
# 
# +
def sink(array, i, heap_size):
l, r = left(i), right(i)
largest = i
if l <= heap_size and array[l] > array[largest]:
largest = l
if r <= heap_size and array[r] > array[largest]:
largest = r
if largest == i:
return
array[i], array[largest] = array[largest], array[i]
sink(array, largest, heap_size)
array = [0, 16, 4, 10, 14, 7, 9, 3, 2, 8, 1]
sink(array, 2, len(array) - 1)
print(array)
# -
# #### Sink操作的时间复杂度分析
#
# 1. 在判断节点$i$与其两个子节点的关系并且交换位置的步骤复杂度为$\Theta(1)$
# 2. 在递归调用某一个子节点时, 假设以$i$为根节点的树共有$n$个节点, 那么以其子节点为根的子树至多有${2n}\over 3$个节点
# 3. 只有一次递归调用
#
# 综上, 我们可以写出Sink的$T(n)$如下:
#
# $T(n) \leq T({ {2n} \over 3}) + O(1)$
#
# 运用Master Theorem, 得出a=1, b=${3 \over 2}=1.5$, d=0.
#
# $$
# T(n)=
# \begin{cases}
# O(n^dlogn)& ,\text{if } a=b^d\\
# O(n^d)& ,\text{if } a < b^d\\
# O(n^{log_ba})& ,\text{if } a > b^d
# \end{cases}
# $$
#
# $b^d=1 = a$, 属于case 1, 所以复杂度为$O(logn)$
# ### Build heap
#
# 建堆的过程非常容易, 只需反复调用Sink操作即可实现. 通过分析我们知道假设数组$A$长度为$n$, 那么$A[n/2+1...n]$都是叶节点, 即它们都是长度为1的堆, 我们只需要关心前半部分也同样满足堆的性质即可.
# +
def build_heap(array):
heap_size = len(array) - 1
for i in range(int(heap_size / 2), 0, -1):
sink(array, i, heap_size)
array = [0, 6, 4, 10, 14, 7, 9, 3, 2, 8, 1]
build_heap(array)
print(array)
# -
# ***思考*** 如何证明该操作的正确性?
# #### Build heap的时间复杂度分析 \[重点\]
#
# 由前文可得到每一次Sink操作消耗$O(log n)$的时间, 并且build-heap进行了${n \over 2}=O(n)$次调用, 所以复杂度为$O(logn) \times O(n) = O(nlogn)$.
#
# 然而这是一个很松弛的渐进上界, 因为当对很底层的节点作sink操作时, 虽然堆中的节点数并没有发生改变, 但sink操作的时间会比对根节点进行sink操作短很多. 因此我们可以得出一个更好更紧密的上界.
#
# 通过观察我们知道$n$个节点的堆的高度为$log n$, 且高度正好为$h$的节点数至多有${n \over {2^{h+1}}}$个. 而高度为$h$的树, 对其根节点进行sink操作的复杂度为$O(h)$, 所以我们可以将build-heap的复杂度表示为
# $$
# O\Big(\sum_{h=0}^{logn} {n \over {2^{h+1}}} O(h) \Big) = O\Big( n \sum_{h=0}^{logn} {h \over {2^{h+1}}} \Big)
# $$
#
# 令$k=logn, S=\sum_{h=0}^{k} {h \over {2^{h+1}}}$, 则$2S=\sum_{h=0}^{k} {h \over {2^{h}}}$.
#
# $2S - S=({1\over2}+{2\over4}+{3\over8}+...+{k\over{2^k}}) - ({1\over4}+{2\over8}+...+{{k-1}\over{2^k}} + {k\over{2^{k+1}}})$
#
# $S={1\over2} + {1\over4} + {1\over8} + ... + {1\over{2^k}} - {k\over{2^{k+1}}}$
#
# 显而易见,
# $$S \lt 1 - {k\over{2^{k+1}}} \lt 1$$
#
# 所以$O\Big( n \sum_{h=0}^{logn} {h \over {2 ^ {h + 1}}} \Big)=O(n \times S)=O(n)$, 即build-heap的复杂度其实为$ O(n)$, 牛逼得抠脚!
# ### The MIGHTY heapsort algorithm
#
# 接下来就是堆排序的主体部分了. 用自然语言描述堆排序的过程为:
#
# 1. 对输入序列$A[1...n]$建堆 (build-heap操作).
# 2. 由于最大堆的定义是最大的值在根节点, 所以我们将$A[1]$与$A[n]$交换位置, 并将堆的大小-1.
# 3. 下沉$A[1]$, 因为此时$A[1]$是从最后直接上来的, 并不一定满足堆的性质.
# 4. 重复2, 3步骤, 直到堆中剩余元素为2.
#
# 经过上述过程之后的原序列$A[1...n]$即为排序后的序列, 也就是说, 堆排序是不额外消耗空间的.
#
# 图示过程:
#
# 
#
# 上代码:
# +
def heapsort(array):
# 在数组前添加一个占位符, 使得真正的数组从1开始
array.insert(0, 0)
# 建堆
build_heap(array)
heap_size = len(array) - 1
# 从当前堆的最后一个叶节点开始与array[1]交换, 并将交换后的节点排除在堆之外
# 最后下沉array[1]
for i in range(len(array)-1, 1, -1):
array[1], array[i] = array[i], array[1]
heap_size -= 1
sink(array, 1, heap_size)
# 将最开始的占位符删掉
array.pop(0)
a = [16, 1, 10, 8, 7, 9, 3, 2, 4, 14]
heapsort(a)
print(a)
# -
# #### 时间复杂度分析
#
# 建堆需要$O(n)$的时间, 每次循环内的下沉需要$O(log n)$的时间, 循环n-1次, 所以复杂度为$O(n + n log n)=O(nlogn)$
#
# ---
# ## 快速排序
#
# 快速排序使用了分治的手段, 以下为使用分治排序序列$A[p...r]$的三个典型步骤:
# ### Divide
#
# 将$A[p...r]$分成两个子序列$A_1=A[p...q-1]$和$A_2=A[q+1...r]$, 使得$A_1$中所有元素都≤$A[q]$, 且$A_2$中所有元素都≥$A[q]$. 这一步骤的关键即为计算索引$q$, 该索引对应的值我们称为pivot.
#
# 图示过程:
#
# 
def partition(array, p, r):
pivot = array[r]
i = p - 1
for j in range(p, r):
if array[j] <= pivot:
i += 1
array[i], array[j] = array[j], array[i]
array[i + 1], array[r] = array[r], array[i + 1]
return i + 1
# ### Conquer
#
# 使用递归调用分别对$A_1, A_2$进行排序.
# ### Combine
#
# 将排序后的$A_1, A[q], A_2$拼接起来.
#
#
# 上述步骤的代码实现:
# +
def quick_sort(array, p, r):
if p < r:
q = partition(array, p, r)
quick_sort(array, p, q - 1)
quick_sort(array, q + 1, r)
import random
a = [1, 4, 2, 9, 0, 6, 5, 7, 3, 10, 8, 1, 0]
quick_sort(a, 0, len(a) - 1)
print(a)
# -
# ### 效率
#
# 快速排序的运行效率取决于partition步骤是否能尽量平均地将原序列砍成两部分, 也就是说问题的关键落在了寻找合适的pivot上. 当partition很平均时, 快速排序的效率会非常高, 因为每一个子问题的规模都缩小了一半, 通过Master Theorem得知运行效率为$O(nlogn)$; 当partition不平均时, 其中一个子问题size为0, 另一个缩小为n-1, 这是非常糟糕的运行效率, 一开始我们就讲过.
#
# #### 最坏情况直观分析
#
# 假设每次递归调用中我们都只能将序列分为n-1和0, 并且partition消耗$\Theta(n)$时间, 有:
#
# $T(n)=T(n-1)+T(0)+\Theta(n) = T(n-1) + \Theta(n)=O(n^2)$ (如何证明?)
#
# #### 最佳情况直观分析
#
# 若partition能做到最大限度的平均分, 那么得到的两个子问题规模都为原来的$1\over 2$, 这时快速排序会运行得非常快:
#
# $T(n)=2T({n\over2})+\Theta(n)=O(nlogn)$ (如何证明?)
#
# #### 一般情况直观分析
#
# 假设在每一次partition中我们分别得到两个子问题的规模为${n \over 10}$和${{9n} \over 10}$, 则:
#
# $T(n)=T({9\over10}n)+T({n \over 10}) + cn$
#
# 根据定义, 在这里用cn来代替$\Theta(n)$是成立的. 接下来我们观察下图:
#
# 
#
# 易知总是被分为$1\over10$的那一支的深度为$log_{10}n$, 因为问题规模每次都缩小为原来的$1 \over 10$. 类似地, 总是被分为$9\over10$的那一支深度为$log_{10\over9}n$. 而其他所有分支的深度都在这个范围之间. 且在当深度≤$log_{10}n$时, 这一层所有节点中的序列长度和为n, 即这一层中每个节点处理partition的复杂度之和为$\Theta(n)=cn$. 而再往下走, 每一层的partition复杂度之和一定会≤$cn$. 那么我们有如下关系:
# $$
# T(n)=O\big({log_{10}n \times cn} + (log_{10\over9}n - log_{10}n) \times cn \big)
# =O\big( cnlogn + cn \times logn \big)=O(nlogn)
# $$
#
# 通过以上分析我们得出结论, 就算是partition做得非常不好, 如9-1分, 甚至是99-1分, 只要能满足每次都是按照常数比例, 在渐进意义(asymptotically)上快速排序的时间复杂度总是为$O(nlogn)$.
# ### 随机化快速排序
#
# 很多算法都会采用随机化来获取更好的期望效率, 随机化同样可以用在快速排序中以获取更好的效率, 但做法并不是将原序列随机排序, 而是在partition过程中取代$pivot=A[r]$的做法, 让$pivot=A[i], i=random(p, r)$. 这种做法叫做**random sampling**.
def randomized_partition(array, p, r):
i = random.randint(p, r)
array[i], array[r] = array[r], array[i]
return partition(array, p, r)
# 使用随机化的partition来实现快速排序也非常容易, 用`randomized_partition`来取代原来的`partition`即可:
# +
def randomized_quicksort(array, p, r):
if p < r:
q = randomized_partition(array, p, r)
randomized_quicksort(array, p, q - 1)
randomized_quicksort(array, q + 1, r)
a = [1, 4, 2, 9, 0, 6, 5, 7, 3, 10, 8, 1, 0]
randomized_quicksort(a, 0, len(a) - 1)
print(a)
# -
# #### 复杂度分析
#
# 通过观察和分析算法, 不难知道快速排序的时间消耗主要集中在partition上. 每一次partition被调用时, 一个pivot会被选中, 并且在之后的所有递归调用中这个pivot将不再会被包含. 因此, 在整个快速排序的过程中partition至多被调用n次. 接下来分析partition中的时间消耗:
# ```python
# 1. def partition(array, p, r):
# 2. pivot = array[r]
# 3. i = p - 1
# 4. for j in range(p, r):
# 5. if array[j] <= pivot:
# 6. i += 1
# 7. array[i], array[j] = array[j], array[i]
# 8. array[i + 1], array[r] = array[r], array[i + 1]
# 9. return i + 1
# ```
# 在一次partition中, 2-3行消耗常数时间, 4-7行消耗时间取决于for循环的次数, 具体等于(循环次数)$\times$(5-7行消耗时间) (某个常数). 那么如果我们能算出在整个算法执行中有多少次第5行的比较, 就能得出有多少时间被消耗在了partition中的for循环上.
#
# 设X为比较次数, 那么我们可以将快速排序的时间复杂度表示为$O(n+X)$. (想想如何证明这个表示是正确的)
#
# **计算X**
#
# 要计算X, 我们必须理解数组中的两个元素在何时需要比较, 何时不需要. 我们假设有一个新的序列$Z=<z_1, z_2, ..., z_n>, z_i$表示数组A中第i小的元素. $Z_{ij}=<z_i, z_{i+1}, ..., z_j>$
#
# 在算法中, $z_i, z_j$至多会被比较一次. 因为partition中每个元素都只跟pivot作比较, 而pivot不会被第二次使用.
#
# 定义$X_{ij}=I\{z_i\text{是否与}z_j\text{比较}\}$
#
# 那么我们就有
# $$
# X=\sum_{i=1}^{n-1} \sum_{j=i+1}^{n} X_{ij}
# $$
# 同时对等号两边求数学期望:
# $$
# \begin{align*}
# E[X]&=E\Big[ \sum_{i=1}^{n-1} \sum_{j=i+1}^{n} X_{ij} \Big] \\
# &=\sum_{i=1}^{n-1} \sum_{j=i+1}^{n} E[X_{ij}] \\
# &=\sum_{i=1}^{n-1} \sum_{j=i+1}^{n} p(z_i, z_j)
# \end{align*}
# $$
# 其中$p(a, b)$为a与b需要比较的概率. 算这个概率我们首先需要分析两个数不会比较的情况.
#
# 假设有一列数1到10, 我们选取了7作为pivot, 那么partition之后我们得到了两个序列, 分别用**集合**表示为{1, 2, 3, 4, 5, 6}与{8, 9, 10}. 在这个过程中, 7与所有数字都比较过一次, 但任意取这两个集合中的各一个元素, 二者都永远不会互相比较了.
#
# 更客观地来描述, 假设我们从$Z_{ij}$中取出pivot$=x$, 且$z_i < x < z_j$, 那么$z_i$与$ z_j$将永不比较; 反之若$x \leq z_i$, 那么$z_i$将与$Z_{ij}$中的所有元素比较 (除了自己), 当$x \geq z_j$时也是类似的情况. 也就是说在$Z_{ij}$中, $z_i$和$z_j$需要比较的话, pivot就必须刚好选到它们二者之一. 由于我们是采取的随机选pivot的做法, 那么每个元素被选中的概率是均匀的, 在$Z_{ij}$中一共有$j-i+1$个元素, 所以每个元素被选中的概率是$1 \over {j-i+1}$, 即:
# $$
# \begin{align*}
# p(z_i, z_j)&=p(z_i\text{被选中} | z_j\text{被选中})\\
# &=p(z_i\text{被选中}) + p(z_j\text{被选中}) \\
# &={1 \over {j-i+1}} \times 2 \\
# &={2 \over {j-i+1}}
# \end{align*}
# $$
#
# 代入原式
#
# $$
# E[X]=\sum_{i=1}^{n-1} \sum_{j=i+1}^{n} {2 \over {j-i+1}}
# $$
#
# 设$k=j-i$
#
# $$
# \begin{align*}
# E[X]&=\sum_{i=1}^{n-1} \sum_{j=i+1}^{n} {2 \over {j-i+1}} \\
# &=\sum_{i=1}^{n-1} \sum_{k=1}^{n-i} {2 \over {k + 1}} \\
# &<\sum_{i=1}^{n-1} \sum_{k=1}^{n} {2 \over k} \\
# \end{align*}
# $$
#
# 求$\sum_{k=1}^n {2 \over k}$这个式子叫做**调和级数**, 很容易证明这个求和是发散的, 并没有通项公式, 那怎么办? 其实也很简单, 当k趋近于$+\infty$时, 它的和为:
#
# $$
# \lim_{n\to+\infty} \sum_{k=1}^n {2 \over k}=2\int_{1}^{+\infty} {1 \over x} dx = 2ln(n)
# $$
#
# 代入后有
#
# $$
# E[X]<\sum_{i=1}^{n-1} 2ln(n)=2n \times ln(n)
# $$
#
# 所以$O(nlogn)$
| lectures/lecture1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] editable=true
# # Part I. ETL Pipeline for Pre-Processing the Files
# + [markdown] editable=true
# ## PLEASE RUN THE FOLLOWING CODE FOR PRE-PROCESSING THE FILES
# + [markdown] editable=true
# #### Import Python packages
# + editable=true
# Import Python packages
import pandas as pd
import cassandra
import re
import os
import glob
import numpy as np
import json
import csv
# + [markdown] editable=true
# #### Creating list of filepaths to process original event csv data files
# + editable=true
# checking your current working directory
print(os.getcwd())
# Get your current folder and subfolder event data
filepath = os.getcwd() + '/event_data'
# Create a for loop to create a list of files and collect each filepath
for root, dirs, files in os.walk(filepath):
# join the file path and roots with the subdirectories using glob
file_path_list = glob.glob(os.path.join(root,'*'))
#print(file_path_list)
# + [markdown] editable=true
# #### Processing the files to create the data file csv that will be used for Apache Casssandra tables
# + editable=true
# initiating an empty list of rows that will be generated from each file
full_data_rows_list = []
# for every filepath in the file path list
for f in file_path_list:
# reading csv file
with open(f, 'r', encoding = 'utf8', newline='') as csvfile:
# creating a csv reader object
csvreader = csv.reader(csvfile)
next(csvreader)
# extracting each data row one by one and append it
for line in csvreader:
#print(line)
full_data_rows_list.append(line)
#print(len(full_data_rows_list))
#print(full_data_rows_list)
# creating a smaller event data csv file called event_data_file_new csv that will be used to insert data into the \
# Apache Cassandra tables
csv.register_dialect('myDialect', quoting=csv.QUOTE_ALL, skipinitialspace=True)
with open('event_datafile_new.csv', 'w', encoding = 'utf8', newline='') as f:
writer = csv.writer(f, dialect='myDialect')
writer.writerow(['artist','firstName','gender','itemInSession','lastName','length',\
'level','location','sessionId','song','userId'])
for row in full_data_rows_list:
if (row[0] == ''):
continue
writer.writerow((row[0], row[2], row[3], row[4], row[5], row[6], row[7], row[8], row[12], row[13], row[16]))
# + editable=true
# check the number of rows in your csv file
with open('event_datafile_new.csv', 'r', encoding = 'utf8') as f:
print(sum(1 for line in f))
# + [markdown] editable=true
# # Part II. Complete the Apache Cassandra coding portion of your project.
#
# ## Now you are ready to work with the CSV file titled <font color=red>event_datafile_new.csv</font>, located within the Workspace directory. The event_datafile_new.csv contains the following columns:
# - artist
# - firstName of user
# - gender of user
# - item number in session
# - last name of user
# - length of the song
# - level (paid or free song)
# - location of the user
# - sessionId
# - song title
# - userId
#
# The image below is a screenshot of what the denormalized data should appear like in the <font color=red>**event_datafile_new.csv**</font> after the code above is run:<br>
#
# <img src="images/image_event_datafile_new.jpg">
# + [markdown] editable=true
# ## Begin writing your Apache Cassandra code in the cells below
# + [markdown] editable=true
# #### Creating a Cluster
# + editable=true
# This should make a connection to a Cassandra instance your local machine
# (127.0.0.1)
from cassandra.cluster import Cluster
cluster = Cluster()
# To establish connection and begin executing queries, need a session
session = cluster.connect()
# + [markdown] editable=true
# #### Create Keyspace
# + editable=true
try:
session.execute("""
CREATE KEYSPACE IF NOT EXISTS sparkifydb
WITH REPLICATION =
{ 'class' : 'SimpleStrategy', 'replication_factor' : 1 }"""
)
except Exception as e:
print(e)
# + [markdown] editable=true
# #### Set Keyspace
# + editable=true
try:
session.set_keyspace('sparkifydb')
except Exception as e:
print(e)
# + [markdown] editable=true
# ### With Apache Cassandra you model the database tables on the queries you want to run. So, we are modeling tables for three queries below.
# + editable=true
# Mapping column location from csv
# 0 --> artist_name
# 1 --> first_name
# 2 --> gender
# 3 --> item_in_session
# 4 --> last_name
# 5 --> length
# 6 --> level
# 7 --> location
# 8 --> session_id
# 9 --> song_title
# 10 --> user_id
# + [markdown] editable=true
# ### 1. Give me the artist, song title and song's length in the music app history that was heard during sessionId = 338, and itemInSession = 4
#
#
# + [markdown] editable=true
# - Query 1 asks us to give artist, song title and song's length for particular session id and item in session. So we will use session id as partition key and item in session as clustering key to model our table and name it as song_details.
# + editable=true
# Query to create the table
query = "CREATE TABLE IF NOT EXISTS song_details"
query = query + "(session_id int, item_in_session int, artist_name text, song_title text, song_length FLOAT, \
PRIMARY KEY (session_id, item_in_session))"
try:
session.execute(query)
except Exception as e:
print(e)
# + editable=true
# Query to populated the data in the table from the csv file
file = 'event_datafile_new.csv'
with open(file, encoding = 'utf8') as f:
csvreader = csv.reader(f)
next(csvreader)
for line in csvreader:
query = "INSERT INTO song_details(session_id, item_in_session, artist_name, song_title, song_length)"
query = query + "VALUES(%s, %s, %s, %s, %s)"
session.execute(query, (int(line[8]), int(line[3]), line[0], line[9], float(line[5])))
# + editable=true
# Query to get the results from the table
song_details_query = "SELECT artist_name, song_title, song_length FROM song_details WHERE session_id = %s AND item_in_session = %s"
try:
rows = session.execute(song_details_query, (338, 4))
except Exception as e:
print(e)
for row in rows:
print("The artist_name is: {}".format(row.artist_name))
print("The song_title is: {}".format(row.song_title))
print("The song_length is: {}".format(row.song_length))
print("******************************************************************************************************************************")
# + [markdown] editable=true
# ### 2. Give me only the following: name of artist, song (sorted by itemInSession) and user (first and last name) for userid = 10, sessionid = 182
#
# + [markdown] editable=true
# - Query 2 asks us to give artist/s, song title, user's first and last name for particular user id and session id. So we will use user id and session id as composite partition key and item in session as clustering key since we need data sorted by item in session to model our table and name it as user_details.
# + editable=true
# Query to create the table
query = "CREATE TABLE IF NOT EXISTS user_details"
query = query + "(user_id int, session_id int, item_in_session int, artist_name text, song_title text, first_name text, last_name text, \
PRIMARY KEY ((user_id, session_id), item_in_session))"
try:
session.execute(query)
except Exception as e:
print(e)
# + editable=true
# Query to populated the data in the table from the csv file
file = 'event_datafile_new.csv'
with open(file, encoding = 'utf8') as f:
csvreader = csv.reader(f)
next(csvreader) # skip header
for line in csvreader:
query = "INSERT INTO user_details(user_id, session_id, item_in_session, artist_name, song_title, first_name, last_name)"
query = query + "VALUES(%s, %s, %s, %s, %s, %s, %s)"
session.execute(query, (int(line[10]), int(line[8]), int(line[3]), line[0], line[9], line[1], line[4]))
# + editable=true
# Query to get the results from the table
user_details_query = "SELECT artist_name, song_title, first_name, last_name FROM user_details WHERE user_id = %s AND session_id = %s"
try:
rows = session.execute(user_details_query, (10, 182))
except Exception as e:
print(e)
for row in rows:
print("The artist_name is: {}".format(row.artist_name))
print("The song_title is: {}".format(row.song_title))
print("The first_name is: {}".format(row.first_name))
print("The last_name is: {}".format(row.last_name))
print("******************************************************************************************************************************")
# + [markdown] editable=true
# ### 3. Give me every user name (first and last) in my music app history who listened to the song 'All Hands Against His Own'
# + [markdown] editable=true
# - Query 3 asks us to give user's first and last name is music app history who listened to the song All Hands Against His Own. Here we use song_title as partiton key and user_id as clustering key and create a table named event_details.
# + editable=true
# Query to create the table
query = "CREATE TABLE IF NOT EXISTS event_details"
query = query + "(song_title text, user_id int, first_name text, last_name text, PRIMARY KEY (song_title, user_id))"
try:
session.execute(query)
except Exception as e:
print(e)
# + editable=true
# Query to populated the data in the table from the csv file
file = 'event_datafile_new.csv'
with open(file, encoding = 'utf8') as f:
csvreader = csv.reader(f)
next(csvreader) # skip header
for line in csvreader:
query = "INSERT INTO event_details(song_title, user_id, first_name, last_name)"
query = query + "VALUES(%s, %s, %s, %s)"
session.execute(query, (line[9], int(line[10]), line[1], line[4]))
# + editable=true
# Query to get the results from the table
event_details_query = "SELECT first_name, last_name FROM event_details WHERE song_title = %s"
try:
rows = session.execute(event_details_query, ('All Hands Against His Own', ))
except Exception as e:
print(e)
for row in rows:
print("All Hands Against His Own is played by")
print("User first_name: {}".format(row.first_name))
print("User last_name: {}".format(row.last_name))
print("******************************************************************************************************************************")
# + [markdown] editable=true
# ### Drop the tables before closing out the sessions
# + editable=true
session.execute("DROP TABLE IF EXISTS song_details")
session.execute("DROP TABLE IF EXISTS user_details")
session.execute("DROP TABLE IF EXISTS event_details")
# + editable=true
# + [markdown] editable=true
# ### Close the session and cluster connection¶
# + editable=true
session.shutdown()
cluster.shutdown()
# + editable=true
| 1. Data Modeling with Postgres and Apache Cassandra/Data Modeling with Apache Cassandra/Data_Modeling_With_Apache_Cassandra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pytesseract
import matplotlib.pyplot as plt
import cv2
image = cv2.imread('opportunity-road.jpg')
image_gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
plt.imshow(image_gray,cmap='gray')
plt.show()
pytesseract.image_to_string(image_gray)
| deprecated/OCR/ocr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Uncommon operation with dataframes
#
# Cheat sheet on uncommand operation with pandas such as reading a big file.
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# ## Pointer on notebooks
#
# * [Rappel de ce que vous savez déjà mais avez peut-être oublié](http://www.xavierdupre.fr/app/ensae_teaching_cs/helpsphinx/notebooks/td2_eco_rappels_1a.html)
# * [Python pour un Data Scientist / Economiste](http://www.xavierdupre.fr/app/ensae_teaching_cs/helpsphinx/td_2a.html)
# * [Exercices Pratiques](http://www.xavierdupre.fr/app/actuariat_python/helpsphinx/i_seances_base.html)
# ## List of strings into binaries features
import pandas
df = pandas.DataFrame([{"target":0, "features":["a", "b", "c"]},
{"target":1, "features":["a", "b"]},
{"target":2, "features":["c", "b"]}])
df
df.features.str.join("*").str.get_dummies("*")
# ## Big files
#
# Let's save some data first.
from sklearn.datasets import load_breast_cancer
data = load_breast_cancer()
import pandas
df = pandas.DataFrame(data.data, columns=data.feature_names)
df.to_csv("cancer.txt", sep="\t", encoding="utf-8", index=False)
# ### first lines : nrows
df = pandas.read_csv("cancer.txt", nrows=3)
df
df = pandas.read_csv("cancer.txt", nrows=3, sep="\t")
df
# ### middle lines : nrows + skiprows
df = pandas.read_csv("cancer.txt", nrows=3, skiprows=100, sep="\t", header=None)
df
# ### big files : iterator
for piece, df in enumerate(pandas.read_csv("cancer.txt", iterator=True, sep="\t", chunksize=3)):
print(piece, df.shape)
if piece > 2:
break
# ### sample on big files : iterator + concat
samples = []
for df in pandas.read_csv("cancer.txt", iterator=True, sep="\t", chunksize=30):
sample = df.sample(3)
samples.append(sample)
dfsample = pandas.concat(samples)
dfsample.shape
| _doc/notebooks/cheat_sheets/chsh_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="-nJ7YEw_vyjG" colab_type="text"
# # **Not an ideal network**
#
# We follow a What+Why framework for each cell.
# + [markdown] id="kO4QF3CokXs7" colab_type="text"
#
#
# - What: Installing keras via pip package manager, Keras is a wrapper over Tensorflow. Then we import it for use
# - Why: This package will help us define our DNN layer by layer
# + id="kGWv5hBhv2jf" colab_type="code" outputId="532ff27c-1456-46f9-d8c8-02ced0edf5d1" colab={"base_uri": "https://localhost:8080/", "height": 51}
# https://keras.io/
# !pip install -q keras
import keras
import tensorflow as tf
tf.test.gpu_device_name()
# + [markdown] id="bTvASHdGk9XT" colab_type="text"
# - What: Import each Library, Classes from Keras.layers will be used to define our network, also import mnist dataset and numpy.
# - Why: We will be doing Mnist classification
# + id="wnMlDJQKv4VG" colab_type="code" colab={}
import numpy as np
from keras.models import Sequential
from keras.layers import Flatten
from keras.layers import Convolution2D
from keras.utils import np_utils
from keras.datasets import mnist
# + [markdown] id="BK0TQUyMlRM1" colab_type="text"
# We first use `help` to check the signature of the function. Using `help` command is going to be useful it seems.
#
# - What: So It gets the mnist data partitioned into Train-Test set
# - Why: Train-test split will be used to train and then test the model on test set
# + id="bx0-PdCZlWg1" colab_type="code" outputId="ae0a501d-0d48-44c3-922f-ecb87bd5fc55" colab={"base_uri": "https://localhost:8080/", "height": 221}
help(mnist.load_data)
# + id="8CdSu2lMwB9s" colab_type="code" outputId="6d8fb6a6-da32-4cc8-ceb5-da53ab423a5c" colab={"base_uri": "https://localhost:8080/", "height": 51}
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# + [markdown] id="mYJuaglEmVh6" colab_type="text"
# - What:
# - We explore our data a little.
# - We print number of training and test examples
# - We also show a example to see what our data looks like
# - `%matplotlib inline` commands makes the plots appear within our notebook
#
# - Why
# - Knowing train and test size and data dimensions will help us design/debug our ConvNet
#
# **Note**
# - the image looks color but it isn't, its just the way `plt.imshow` gives color to different cell values in a 2D matrix
# - See that we have a single channel in the initial image, its a grayscale not RGB image.
# + id="tLaDf0-rwCmj" colab_type="code" outputId="923e79b6-8cfe-4d8a-e43e-cf3b830f863f" colab={"base_uri": "https://localhost:8080/", "height": 320}
print (X_train.shape)
# add a line to print test set shape as well
print (X_test.shape)
from matplotlib import pyplot as plt
# %matplotlib inline
plt.imshow(X_train[0])
# + [markdown] id="GMKfvkffm52n" colab_type="text"
# Once you change CMAP parameter we can see how it is a grayscale image
# + id="ladotsMYucuv" colab_type="code" outputId="edffc261-467d-4afd-dbed-9059ecdc09db" colab={"base_uri": "https://localhost:8080/", "height": 286}
plt.imshow(X_train[0],cmap="binary")
# + [markdown] id="JjR8E1pewFhJ" colab_type="text"
# What: We reshape to get single channel as dimension, the last 1 in each reshape is for the channel. Since the image is single channel so this is done.
#
# If image was 3 channel then
#
# ```python
# X_train = X_train.reshape(X_train.shape[0], 28, 28,3)
# ```
#
# + id="erb11jNwwFwl" colab_type="code" colab={}
X_train = X_train.reshape(X_train.shape[0], 28, 28,1)
X_test = X_test.reshape(X_test.shape[0], 28, 28,1)
# + [markdown] id="dBrpFWY4xTAH" colab_type="text"
# What
# - Convert datatype to float32, and normalize to 0-1 range
#
# Why
# - NNs can use only 0-1 data for our ConvNets, otherwise gradients will go to zero.
# - float32 is used since gpu has 32 bit calculation units
#
# Reference
#
# https://devtalk.nvidia.com/default/topic/994172/how-to-tell-if-gpu-cores-are-actually-32-64-bit-processors/?offset=2
#
# ```
#
# "Most GPUs have a native 32 bit integer multiply machine language instruction"
#
# "The most commonly used criterion for an N-bit processor is the width of integer registers. Based on that, all "recent" NVIDIA GPUs (i.e. Tesla through Pascal architectures, 2007 through today), are 32-bit processors."
#
# ```
# + id="RLK4YDoRwHet" colab_type="code" colab={}
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
# + id="JNKLOmhlwJQl" colab_type="code" outputId="449f1bdb-0688-4617-fc7c-a06c414ef271" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_train[:10]
# + [markdown] id="ICv8LQpuzVzG" colab_type="text"
# - Lets the spread of data to see if it is evenly spread. or some classes are more in number
# + id="CsNvC8wQy91_" colab_type="code" outputId="602652fe-a5ee-43e3-a2c0-f57ae8c0bdbd" colab={"base_uri": "https://localhost:8080/", "height": 286}
import seaborn as sns
sns.countplot(y_train)
# + id="36DUTwGazgfj" colab_type="code" outputId="4b31a514-be67-4bcc-b3f7-5ada3d5bd788" colab={"base_uri": "https://localhost:8080/", "height": 323}
help(np_utils.to_categorical)
# + id="xjU3-DMZznTd" colab_type="code" outputId="b2867434-86be-4327-9443-074dfb028456" colab={"base_uri": "https://localhost:8080/", "height": 187}
np_utils.to_categorical(range(0,10), 10)
# + [markdown] id="1n2CmFKAztX2" colab_type="text"
# What
# - We do one hot encoding of the target labels for using categorical cross entropy loss.
#
# Why
# - Cross entropy loss function for multi-class problems requires your target labels to be one hot encoded.
# + id="YusMJguiwKsM" colab_type="code" colab={}
# Convert 1-dimensional class arrays to 10-dimensional class matrices
Y_train = np_utils.to_categorical(y_train, 10)
Y_test = np_utils.to_categorical(y_test, 10)
# + id="upxc99AswMW0" colab_type="code" outputId="a1f20e04-94c0-45d8-9e50-0a0385b549e3" colab={"base_uri": "https://localhost:8080/", "height": 187}
Y_train[:10]
# + [markdown] id="CEapHPh00Fa8" colab_type="text"
# ## Da Model!!
#
# Possible ideas before we start
#
# - Check receptive fields
# - Check if Maxpool is used after atleast 2/3 layers.
# - Check if channels,kernels,channel dimensions are ok
# - Check if the number of channels are correct for each layer and if we are possibly losing info
# - Count the total number of parameters in the network and see if network is overly complex.
# + id="irTVUE47wNwr" colab_type="code" outputId="24866d20-ea0e-4079-932f-1ddfa6280a9f" colab={"base_uri": "https://localhost:8080/", "height": 853}
from keras.layers import Activation, MaxPooling2D
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
# 28 x 28 x 1 | 1x1 | 32x(3x3x1) = 288
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
# 26x26x32| 3x3 | 64x(3x3x32) = 18432
model.add(Convolution2D(64, 3, 3, activation='relu'))
# 24x24x64 | 5x5 | 128x(3x3x64) = 73728
model.add(Convolution2D(128, 3, 3, activation='relu'))
# 22x22x128| 7x7 | Maxpool layer has no params
model.add(MaxPooling2D(pool_size=(2, 2)))
# Maxpool halves the image in each dimension, so it increases receptive field by 2 times
# 11x11x128 | 14x14 | 256x(3x3x128) = 294912
model.add(Convolution2D(256, 3, 3, activation='relu'))
# 9x9x256 | 16x16 | 512x(3x3x256) = 1179648
model.add(Convolution2D(512, 3, 3, activation='relu'))
# 7x7x512 | 18x18 | 1024x(3x3x512) = 4718592
model.add(Convolution2D(1024, 3, 3, activation='relu'))
# 5x5x1024 | 20x20 | 2048x(3x3x1024) ~ 18x10^6 = 18 Million
model.add(Convolution2D(2048, 3, 3, activation='relu'))
# 3x3x2048 | 22x22 | 10x(3x3x2048) = 184320
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
# I now know how without using Fully Connected layers we can still do classification.
# Had asked in 1st class. :)
model.add(Flatten())
# Use softmax activation for giving probabilities.
model.add(Activation('softmax'))
model.summary()
# + [markdown] id="q1ZQiSBs3MgF" colab_type="text"
# What
# - Compile the network with loss function and optimiser
# - `metrics=['accuracy']` tells what to print as score after each epoch and as score when `model.evaluate` is called. We are concerned with accuracy.
# + id="VYZOpRb6yG7_" colab_type="code" colab={}
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# + id="5O248wVQyMft" colab_type="code" outputId="fa83463e-45d1-4cc0-c7b9-6ea15f939b82" colab={"base_uri": "https://localhost:8080/", "height": 479}
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
# + [markdown] id="Aa6lZwaB2_ZQ" colab_type="text"
# **Observations**
#
# - The accuracy is not increasing.
# - The network is slow as shit, seems the network is overly complex
# + id="Sst4KneiyOL5" colab_type="code" colab={}
score = model.evaluate(X_test, Y_test, verbose=0)
# + id="CfJiXOKsyj4y" colab_type="code" outputId="2c5afe60-086b-4f33-9952-d9c4cb567091" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(score)
# + id="hwLSXt7nyn_0" colab_type="code" colab={}
y_pred = model.predict(X_test)
# + id="WWKKoOKwyppN" colab_type="code" outputId="e7a1e3ff-cdc7-49b6-8f6b-aed71ee47c87" colab={"base_uri": "https://localhost:8080/", "height": 187}
print(y_pred[:9])
print(y_test[:9])
# + [markdown] id="iHHwVQg3bBY4" colab_type="text"
# ### Remove last Relu - Session 3 suggestion
# + id="aRRt3MhV0WYV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 768} outputId="2afc8979-2ed3-44f3-8f86-8603d6a8e167"
from keras.layers import Activation, MaxPooling2D
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
# 28 x 28 x 1 | 1x1 | 32x(3x3x1) = 288
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
# 26x26x32| 3x3 | 64x(3x3x32) = 18432
model.add(Convolution2D(64, 3, 3, activation='relu'))
# 24x24x64 | 5x5 | 128x(3x3x64) = 73728
model.add(Convolution2D(128, 3, 3, activation='relu'))
# 22x22x128| 7x7 | Maxpool layer has no params
model.add(MaxPooling2D(pool_size=(2, 2)))
# Maxpool halves the image in each dimension, so it increases receptive field by 2 times
# 11x11x128 | 14x14 | 256x(3x3x128) = 294912
model.add(Convolution2D(256, 3, 3, activation='relu'))
# 9x9x256 | 16x16 | 512x(3x3x256) = 1179648
model.add(Convolution2D(512, 3, 3, activation='relu'))
# 7x7x512 | 18x18 | 1024x(3x3x512) = 4718592
model.add(Convolution2D(1024, 3, 3, activation='relu'))
# 5x5x1024 | 20x20 | 2048x(3x3x1024) ~ 18x10^6 = 18 Million
model.add(Convolution2D(2048, 3, 3, activation='relu'))
# 3x3x2048 | 22x22 | 10x(3x3x2048) = 184320
model.add(Convolution2D(10, 3, 3))
# Output from conv layers
# 1x1x10
# I now know how without using Fully Connected layers we can still do classification.
# Had asked in 1st class. :)
model.add(Flatten())
# Use softmax activation for giving probabilities.
model.add(Activation('softmax'))
model.summary()
# + id="toWPpLWAboVH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 479} outputId="2c3ab8dd-2ad0-4fa2-8d35-76d4ef47e993"
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + id="gTsUVXTEg93x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 649} outputId="0cee0f3f-0080-45f7-ebc5-0c18b6b02aff"
from keras.layers import Activation, MaxPooling2D
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
# 28 x 28 x 1 | 1x1 | 32x(3x3x1) = 288
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
# 26x26x32| 3x3 | 64x(3x3x32) = 18432
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
# 3x3x2048 | 22x22 | 10x(3x3x2048) = 184320
model.add(Convolution2D(10, 3, 3))
# Output from conv layers
# 1x1x10
# I now know how without using Fully Connected layers we can still do classification.
# Had asked in 1st class. :)
model.add(Flatten())
# Use softmax activation for giving probabilities.
model.add(Activation('softmax'))
model.summary()
# + id="CEXm-Epog907" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="fceabb06-86f5-4204-ef12-6277c1347b74"
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + id="Ygk453jmkiJG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 649} outputId="885587ad-7205-4d77-f35f-cc9444804b3d"
from keras.layers import Activation, MaxPooling2D
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
# 28 x 28 x 1 | 1x1 | 32x(3x3x1) = 288
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
# 26x26x32| 3x3 | 64x(3x3x32) = 18432
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
# 3x3x2048 | 22x22 | 10x(3x3x2048) = 184320
model.add(Convolution2D(10, 3, 3))
# Output from conv layers
# 1x1x10
# I now know how without using Fully Connected layers we can still do classification.
# Had asked in 1st class. :)
model.add(Flatten())
# Use softmax activation for giving probabilities.
model.add(Activation('softmax'))
model.summary()
# + id="romUk-2tkn3i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="e22ba2a9-5be9-480f-de81-24541488eb05"
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + [markdown] id="go2beia31Cd3" colab_type="text"
# # What's wrong
#
# - Receptive field at last layer was 22x22, we are not seeing the full image since image is 28x28
# - Number of parameters is too huge.
# - We use maxpool only once, which is why we have too many conv layers, we can use maxpool 2 times easily
# - Too many conv layers lead to last layer having 2048 kernels and over 18 Million parameters for a single layer
# - No image augmentation, Neither did we do any rotate or any other augmentation like changing brightness levels or invert colors (black to white n vice versa)
#
# - The 1st Maxpool happens at 7x7 receptive field, it should be done at either 9x9 or 11x11, since at 7x7 receptive field the CNN may not be able identify any edges/gradients.
#
# - We are not using blocks (conv-conv-conv-maxpool) style.
#
# - Our overall arch is 32->64->128->MP->256->512->1024->2048->Output layer.
#
# - A better way would be
# - 1st block detect edges/grads as 32->64->128->MP (11x11x128 output)
# - 2nd block detect overall patterns 32->64->128->256->Output Layer.
# - The first block detects edges/gradients of 128 different types, the 2nd block we again start from 32 and proceed, 1st block gives 128 different types of edges, but since our overall labels are few, the number of patterns that can be created from these edges and gradients is also low. Hence we don't start our 2nd block with 256 but rather 32.
#
# Please tell me if the `better way` I am saying is applicable or not?
# + [markdown] id="Uiy_GAmd3iJp" colab_type="text"
# # Make a correct network
# + [markdown] id="zIPIjk_rTQ0X" colab_type="text"
# ## Double Max Pool
#
# - Double max pool may hurt performance
# + id="g2fp9zxE3lLo" colab_type="code" outputId="1b069e74-6e66-4416-ad04-8f011ae374d6" colab={"base_uri": "https://localhost:8080/", "height": 564}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + [markdown] id="nxFEp0BMTaRW" colab_type="text"
# ## Block style architecture
#
#
# + [markdown] id="H7vDCEO2T0NQ" colab_type="text"
# ### Style 1
#
# - 1st block detect edges/grads as 32->64->128->MP (11x11x128 output)
# - 2nd block detect overall patterns 32->64->128->256->Output Layer.
# + id="HokISj71Qk_e" colab_type="code" outputId="30aec03a-b61b-4639-8d01-a170ff55367c" colab={"base_uri": "https://localhost:8080/", "height": 666}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(32, 3, 3, activation='relu'))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + [markdown] id="r1IbEE8xT4rO" colab_type="text"
# ### Style 2
#
# - 1st block detect edges/grads as 32->64->128->256->256->MP (9x9x256 output)
# - 2nd block detect overall patterns 32->64->128->Output Layer.
# + id="wdeaJAWVT3zQ" colab_type="code" outputId="90d00edf-1c54-4b89-b586-67f9d16ef2fc" colab={"base_uri": "https://localhost:8080/", "height": 700}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(32, 3, 3, activation='relu'))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + [markdown] id="bhOp_B1dU0is" colab_type="text"
# ### Style 3
#
# - 1st block detect edges/grads as 32->64->MP (12x12)
# - 2nd Block 64->128->256->MP->Output
# + id="bPOaEAI4U00-" colab_type="code" outputId="eec8bb82-bdca-4b6c-a2f1-f7cd45868018" colab={"base_uri": "https://localhost:8080/", "height": 615}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + id="bWXogMviXCn5" colab_type="code" colab={}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(Convolution2D(512, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + [markdown] id="FkIYmcKgWJ7E" colab_type="text"
# ### Style 4 (Funnel shape of kernels)
#
#
# + id="_4kIvCdhWZjD" colab_type="code" outputId="16bf94c4-9f40-4dae-9c3d-a4294f31e524" colab={"base_uri": "https://localhost:8080/", "height": 700}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(Convolution2D(512, 3, 3, activation='relu'))
model.add(Convolution2D(512, 3, 3, activation='relu'))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + [markdown] id="R0vkT4BZWspr" colab_type="text"
# ### Style 5 (Another funnel shape)
# + id="7B8jaixoWwWB" colab_type="code" outputId="ff326718-0e63-43fc-bab2-4ef4dfd63deb" colab={"base_uri": "https://localhost:8080/", "height": 666}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(Convolution2D(256, 3, 3, activation='relu'))
model.add(Convolution2D(512, 3, 3, activation='relu'))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + [markdown] id="QSMvPAYdXK0e" colab_type="text"
# ### Drastically reduced params model
# + id="hqOE6KW_XOCx" colab_type="code" outputId="1297bca0-a1f9-4a76-a614-b621d13cca38" colab={"base_uri": "https://localhost:8080/", "height": 683}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(32, 3, 3, activation='relu'))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2))) # 11x11
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + id="SANV3lcZXwq1" colab_type="code" outputId="927bc4c0-7214-405b-9c27-91841861dd61" colab={"base_uri": "https://localhost:8080/", "height": 581}
model = Sequential()
# Input: width x height x num channels (depth) | Inputs receptive field width x height | parameters this layer
model.add(Convolution2D(32, 3, 3, activation='relu', input_shape=(28,28,1)))
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2))) # 12x12
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(128, 3, 3, activation='relu'))
model.add(Convolution2D(10, 3, 3, activation='relu'))
# Output from conv layers
# 1x1x10
model.add(Flatten())
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.fit(X_train, Y_train, batch_size=32, nb_epoch=10, verbose=1)
score = model.evaluate(X_test, Y_test, verbose=0)
print(score)
# + [markdown] id="8Ua9x2Bd3l8i" colab_type="text"
# # Pytorch
#
# work in progress
# + id="UVLPjE1i3ncJ" colab_type="code" colab={}
# + [markdown] id="k67cBpavbJZ8" colab_type="text"
# # What's wrong (Repeated cell for convinience)
#
# - Receptive field at last layer was 22x22, we are not seeing the full image since image is 28x28
# - Number of parameters is too huge.
# - We use maxpool only once, which is why we have too many conv layers, we can use maxpool 2 times easily
# - Too many conv layers lead to last layer having 2048 kernels and over 18 Million parameters for a single layer
# - No image augmentation, Neither did we do any rotate or any other augmentation like changing brightness levels or invert colors (black to white n vice versa)
#
# - The 1st Maxpool happens at 7x7 receptive field, it should be done at either 9x9 or 11x11, since at 7x7 receptive field the CNN may not be able identify any edges/gradients.
#
# - We are not using blocks (conv-conv-conv-maxpool) style.
#
# - Our overall arch is 32->64->128->MP->256->512->1024->2048->Output layer.
#
# - A better way would be
# - 1st block detect edges/grads as 32->64->128->MP (11x11x128 output)
# - 2nd block detect overall patterns 32->64->128->256->Output Layer.
# - The first block detects edges/gradients of 128 different types, the 2nd block we again start from 32 and proceed, 1st block gives 128 different types of edges, but since our overall labels are few, the number of patterns that can be created from these edges and gradients is also low. Hence we don't start our 2nd block with 256 but rather 32.
#
# Please tell me if the `better way` I am saying is applicable or not?
# + id="gWVPkPlZbM3v" colab_type="code" colab={}
| assignment-2/Session2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/DannielM/Ola--Mundo/blob/master/Aula01.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="z9XYg7h2FcNq" outputId="c11a384b-0045-4b17-f1da-8cdaa37143a1" colab={"base_uri": "https://localhost:8080/", "height": 244}
import pandas as pd
fonte = "https://github.com/alura-cursos/imersao-dados-2-2020/blob/master/MICRODADOS_ENEM_2019_SAMPLE_43278.csv?raw=true"
dados = pd.read_csv(fonte)
dados.head()
# + id="roHaxkDKHfDt" outputId="9b43b159-dacb-4011-e9bf-e1fcfba6217c" colab={"base_uri": "https://localhost:8080/", "height": 34}
dados.shape
# + id="nE1TlKlrHr1y" outputId="c189ff27-78ac-47fe-f665-431cc89d9eaa" colab={"base_uri": "https://localhost:8080/", "height": 218}
dados["SG_UF_RESIDENCIA"]
# + id="iKsMyeSjJvCa" outputId="8a9262f3-2c45-499f-e1a3-525f48f7e973" colab={"base_uri": "https://localhost:8080/", "height": 672}
dados.columns.values
# + id="4RinKz2RKws6" outputId="2d38bfd2-b4e9-404d-fbaa-967e36bf6095" colab={"base_uri": "https://localhost:8080/", "height": 402}
dados[["SG_UF_RESIDENCIA", "Q025"]]
# + id="9XyBWm9DL3hI" outputId="6b36ebf8-d04a-4ba0-88dc-a4890c591648" colab={"base_uri": "https://localhost:8080/", "height": 218}
dados["SG_UF_RESIDENCIA"]
# + id="WEH6_lABMT2T" outputId="77732434-1354-4861-93b7-097e5ef8fb93" colab={"base_uri": "https://localhost:8080/", "height": 67}
dados["SG_UF_RESIDENCIA"].unique()
# + id="kDXEe1rxMY0D" outputId="5daa12ce-1713-41e8-d8de-a2b5451829ce" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(dados["SG_UF_RESIDENCIA"].unique())
# + id="yyQaMKoyMeaK" outputId="0fbf3170-ff28-42da-8064-5a4498ae4152" colab={"base_uri": "https://localhost:8080/", "height": 487}
dados["SG_UF_RESIDENCIA"].value_counts()
# + id="ML7wBPY1MrZL" outputId="46f5a12f-2a03-48a8-bfca-ab1ff55b60b7" colab={"base_uri": "https://localhost:8080/", "height": 218}
dados["NU_IDADE"].value_counts()
# + id="xj2Lmq2_NNPk" outputId="22ee15c5-c1c1-4d74-a9a3-a95c8131c989" colab={"base_uri": "https://localhost:8080/", "height": 218}
dados["NU_IDADE"].value_counts().sort_index()
# + id="ioQ7i7Q1Oln8" outputId="e9ba28dd-d98e-41ea-b667-2b678f2bd0da" colab={"base_uri": "https://localhost:8080/", "height": 285}
dados["NU_IDADE"].hist()
# + id="TYd8dPfNO1-n" outputId="1c6c0355-1c6c-4d50-f081-ce8ca42d1fe2" colab={"base_uri": "https://localhost:8080/", "height": 500}
dados["NU_IDADE"].hist(bins = 20, figsize = (10,8))
# + id="Al4Yeg7wSFN6"
# + id="VrQVzO4vNyTP" outputId="299bd33d-af75-476b-8116-d1db32d9a8ea" colab={"base_uri": "https://localhost:8080/", "height": 655}
dados.query("IN_TREINEIRO == 1")["NU_IDADE"].value_counts().sort_index()
# + id="NCPdzdkeNioz" outputId="1c8f71c9-e16b-43a6-e397-e4da971a0fc3" colab={"base_uri": "https://localhost:8080/", "height": 391}
dados["NU_NOTA_REDACAO"].hist(bins = 20, figsize=(8, 6))
# + id="2tJt5q4dSolR" outputId="cc019a85-32b0-47af-f44b-ff2091e661bc" colab={"base_uri": "https://localhost:8080/", "height": 391}
dados["NU_NOTA_LC"].hist(bins = 20, figsize=(8, 6))
# + id="BDLcSca0S8nd" outputId="c2b4c76f-c929-4d62-eaf5-a0a00fd9f071" colab={"base_uri": "https://localhost:8080/", "height": 34}
dados["NU_NOTA_REDACAO"].mean()
# + id="Vz4VGeogTHRw" outputId="da7be8bc-e3f9-4e48-b91f-730143ca5f39" colab={"base_uri": "https://localhost:8080/", "height": 34}
dados["NU_NOTA_REDACAO"].std()
# + id="YeKjZt4oTS5o" outputId="2d55e6ef-e649-4ad9-ecea-a1866a5c58ec" colab={"base_uri": "https://localhost:8080/", "height": 284}
provas = ["NU_NOTA_CN","NU_NOTA_CH","NU_NOTA_MT","NU_NOTA_LC","NU_NOTA_REDACAO"]
dados[provas].describe()
# + id="FMaMoSUyUD_8" outputId="ba489f7f-17b3-44dc-a2a8-ea03dac518f1" colab={"base_uri": "https://localhost:8080/", "height": 34}
dados["NU_NOTA_LC"].quantile(0.1)
# + id="03RSXG1vVQOi" outputId="b43dd5f9-8ad9-4790-f0f6-5f2160529285" colab={"base_uri": "https://localhost:8080/", "height": 392}
dados["NU_NOTA_LC"].plot.box(grid = True, figsize=(8,6))
# + id="y_62npBVWvW5" outputId="f6280615-431b-4c1f-b0a0-8e732845a343" colab={"base_uri": "https://localhost:8080/", "height": 501}
dados[provas].boxplot(grid=True, figsize= (10,8))
# + [markdown] id="2iyYYnhjNzRr"
# Desafio01: Proporção dos inscritos por idade.
#
# Desafio02: Descobrir de quais estados são os inscritos com 13 anos.
#
# Desafio03: Adicionar título no gráfico
#
# Desafio04: Plotar os Histogramas das idades dos do treineiro e não treineiros.
#
# Desafio05: Comparar as distribuições das provas em inglês espanhol
#
# Desafio06: Explorar a documentações e visualizações com matplotlib ou pandas e gerar novas visualizações.
# + id="X-vN4RHqZrak"
| Aula01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# +
# Libs initialization
import pandas as pd
import nltk
nltk.download('stopwords')
nltk.download('wordnet')
nltk.download('omw-1.4')
import re
# import scipy
# from scipy.signal import fftconvolve
# -
# # 1. Data file preparation
# </br> Retrieving Data from a dataset collected using the Twitter API, renaming classes and randomizing.
df = pd.read_csv('Data.csv', sep= ';', index_col=0)
df = df.dropna()
df.shape
df.drop_duplicates(subset={'Text'}, inplace=True)
df.head(7)
# Converting Class names to codes/numbers
j = df['True class'].unique().tolist()
z = df['True class'].unique().tolist()
for i in range (0, len(j)):
df.loc[df['True class'] == j[i], 'True class'] = i
df.loc[df['Class'] == j[i], 'Class'] = i
df = df.reset_index(drop=True)
# A List of English contractions from https://en.wikipedia.org/wiki/Wikipedia%3aList_of_English_contractions
c_dict = {
"ain't": "am not",
"aren't": "are not",
"can't": "cannot",
"can't've": "cannot have",
"'cause": "because",
"could've": "could have",
"couldn't": "could not",
"couldn't've": "could not have",
"didn't": "did not",
"doesn't": "does not",
"don't": "do not",
"hadn't": "had not",
"hadn't've": "had not have",
"hasn't": "has not",
"haven't": "have not",
"he'd": "he would",
"he'd've": "he would have",
"he'll": "he will",
"he's": "he is",
"how'd": "how did",
"how'll": "how will",
"how's": "how is",
"i'd": "i would",
"i'll": "i will",
"i'm": "i am",
"i've": "i have",
"isn't": "is not",
"it'd": "it would",
"it'll": "it will",
"it's": "it is",
"let's": "let us",
"ma'am": "madam",
"mayn't": "may not",
"might've": "might have",
"mightn't": "might not",
"must've": "must have",
"mustn't": "must not",
"needn't": "need not",
"oughtn't": "ought not",
"shan't": "shall not",
"sha'n't": "shall not",
"she'd": "she would",
"she'll": "she will",
"she's": "she is",
"should've": "should have",
"shouldn't": "should not",
"that'd": "that would",
"that's": "that is",
"there'd": "there had",
"there's": "there is",
"they'd": "they would",
"they'll": "they will",
"they're": "they are",
"they've": "they have",
"wasn't": "was not",
"we'd": "we would",
"we'll": "we will",
"we're": "we are",
"we've": "we have",
"weren't": "were not",
"what'll": "what will",
"what're": "what are",
"what's": "what is",
"what've": "what have",
"where'd": "where did",
"where's": "where is",
"who'll": "who will",
"who's": "who is",
"won't": "will not",
"wouldn't": "would not",
"you'd": "you would",
"you'll": "you will",
"you're": "you are",
"1st": "first",
"2nd": "second",
"3rd": "third",
"4th": "forth",
"5th": "fifth",
"6th": "sixth",
"7th": "seventh",
"8th": "eighth",
"9th": "ninth"
}
# # 2. Text Filtering
# </br> Four-step raw tex filter, using [nltk](https://www.nltk.org) libraries. Filter includes:
# * Constructions filter (special thanks for [arturomp](https://stackoverflow.com/users/583834/arturomp) for [converting](https://stackoverflow.com/posts/19794953/revisions) wikipedia contraction-to-expansion page into a python dictionary)
# * Stopwords filter
# * Unwanted characters filter
# +
def text_filter(text):
# Convert words to lower case
text = text.lower()
# Remove constructions
text = text.split()
new_text = []
for word in text:
if word in c_dict:
new_text.append(c_dict[word])
else:
new_text.append(word)
text = " ".join(new_text)
# Remove unwanted characters
text = re.sub(r'https?:\/\/.*[\r\n]*', '', text, flags=re.MULTILINE)
text = re.sub(r'[_"\-;%()|+&=*%.,!?:#$@\[\]/]', ' ', text)
text = re.sub(r'\<a href', ' ', text)
text = re.sub(r'&', '', text)
text = re.sub(r'<br />', ' ', text)
text = re.sub(r'\'', ' ', text)
# Remove SW
text = text.split()
sw = set(nltk.corpus.stopwords.words("english"))
text = [w for w in text if not w in sw]
text = " ".join(text)
# Split numbers and words
text = text.split()
new_text2 = []
for word in text:
if (word.isalpha() or word.isdigit()):
new_text2.append(word)
else:
for i in range(0,len(word)-1):
if ((word[i].isdigit() and word[i+1].isalpha()) or (word[i+1].isdigit() and word[i].isalpha())):
word1 = word[0:(i+1)]
word2 = word[(i+1):len(word)]
new_text2.append(word1)
new_text2.append(word2)
text = " ".join(new_text2)
return text
def text_tokenizer(text):
text = nltk.WordPunctTokenizer().tokenize(text)
return text
def joinclean(text):
text = str(' '.join(text))
return text
# -
df['CleanedText'] = list(map(text_filter, df['Text']))
df['TokenizedText'] = list(map(text_tokenizer, df['CleanedText']))
lemm = nltk.stem.WordNetLemmatizer()
df['TokenizedText'] = list(map(lambda word:
list(map(lemm.lemmatize, word)),
df['TokenizedText']))
df['CleanedText'] = list(map(joinclean, df['TokenizedText']))
df.head(7)
# +
# Creating full corpus of words
corpus = []
for i in range(0, len(df)):
corpus.append(df.loc[i, 'TokenizedText'])
# w_list = df.loc[i, 'Tokenized Text']
# for j in range (0, len(w_list)):
# corpus.append(w_list[j])
# corpus = set(corpus)
df['Words quantity'] = 0
for i in range(0, len(df)):
word_list = df.TokenizedText[i]
df.loc[i, ['Words quantity']] = len(word_list)
df.head(3)
# -
# # 3. Filtered Data output
# </br> Data is output to files by categories for Logistic Regression and to one file for other types of algorithms.
# Creating the final dataset
df = df[['CleanedText', 'TokenizedText', 'Words quantity', 'Class', 'True class']]
df.to_csv('FilteredData.csv', sep=';', encoding='utf-8', index=False)
df.head(7)
| Data/Filtering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style="color:blue">Pivot basics</h1>
import pandas as pd
import numpy as np
df = pd.read_csv("weather.csv")
df
df.pivot(index='city', columns='date')
df.pivot(index='city', columns='date', values="humidity")
df.pivot(index='date', columns='city')
df.pivot(index='humidity', columns='city')
# <h1 style="color:blue">Pivot Table</h1>
df = pd.read_csv("weather2.csv")
df
# df.pivot_table(index="city")
pd.pivot_table(df, index="city")
pd.pivot_table(df, index="city", values='temperature')
pd.pivot_table(df, index="city", values='temperature', columns="humidity")
df.pivot_table(index="city", columns="date")
# <h2 style="color:brown">Margins</h2>
df.pivot_table(index="city", columns="date", margins=True)
df.pivot_table(index="city", columns="date", margins=True, aggfunc=np.sum)
df.pivot_table(index="city", columns="date", margins=True, aggfunc='median')
# <h2 style="color:brown">Grouper</h2>
df = pd.read_csv("weather3.csv")
df
df['date'] = pd.to_datetime(df['date'])
df.pivot_table(index=pd.Grouper(freq='M',key='date'),columns='city')
| 8_Grouping and Aggregating/3_Pivot_table/pandas_pivot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp numpy
# -
# # 00_Numpy
#
# > Building an example `Dataset` and `DataLoader` with `NumPy`
#hide
from nbdev.showdoc import *
#export
from fastai2.tabular.all import *
# For our data we'll first utilize `TabularPandas` for pre-processing. One potential is to use `TabularPandas` for pre-processing only, or to integrate `NumPy` directly into it
path = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(path/'adult.csv')
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race']
cont_names = ['age', 'fnlwgt', 'education-num']
procs = [Categorify, FillMissing, Normalize]
y_names = 'salary'
splits = RandomSplitter()(range_of(df))
# We'll still build our regular `TabularPandas`, as we haven't done any `NumPy` modifications yet
to = TabularPandas(df, procs=procs, cat_names=cat_names, cont_names=cont_names,
y_names=y_names, splits=splits)
#export
class NumpyDataset():
"A `Numpy` dataset object from `TabularPandas`"
def __init__(self, to:TabularPandas):
self.cats = to.cats.to_numpy().astype(np.long)
self.conts = to.conts.to_numpy().astype(np.float32)
self.ys = to.ys.to_numpy()
def __getitem__(self, idx):
idx = idx[0]
return self.cats[idx:idx+self.bs], self.conts[idx:idx+self.bs], self.ys[idx:idx+self.bs]
def __len__(self): return len(self.cats)
ds = NumpyDataset(to)
ds.bs = 3
a,b,c = ds[[0]]
test_eq(len(a), 3)
#export
class NumpyDataLoader(DataLoader):
def __init__(self, dataset, bs=1, **kwargs):
"A `DataLoader` for a `NumpyDataset`"
super().__init__(dataset, bs=bs, **kwargs)
self.dataset.bs = bs
def create_item(self, s): return s
def create_batch(self, b):
cat, cont, y = self.dataset[b]
return tensor(cat).to(self.device), tensor(cont).to(self.device), tensor(y).to(self.device)
dl = NumpyDataLoader(ds, bs=3)
batch = next(iter(dl))
test_eq(len(dl), len(ds)//3+1)
#export
@patch
def shuffle_fn(x:NumpyDataLoader):
"Shuffle the interior dataset"
rng = np.random.permutation(len(x.dataset))
x.dataset.cats = x.dataset.cats[rng]
x.dataset.conts = x.dataset.conts[rng]
x.dataset.ys = x.dataset.ys[rng]
#export
@patch
def get_idxs(x:NumpyDataLoader):
"Get index's to select"
idxs = Inf.count if x.indexed else Inf.nones
if x.n is not None: idxs = list(range(len(x.dataset)))
if x.shuffle: x.shuffle_fn()
return idxs
# To ensure that we still see an improvement, we'll compare timings
train_ds = NumpyDataset(to.train)
valid_ds = NumpyDataset(to.valid)
train_dl = NumpyDataLoader(train_ds, bs=64, shuffle=True, drop_last=True)
valid_dl = NumpyDataLoader(valid_ds, bs=64)
dls = to.dataloaders(bs=64)
# %%timeit
# Numpy
for _ in train_dl: pass
# %%timeit
# fastai
for _ in dls[0]: pass
# %%timeit
# Numpy
for _ in valid_dl: pass
# %%timeit
# fastai
for _ in dls[1]: pass
# export
class NumpyDataLoaders(DataLoaders):
def __init__(self, to, bs=64, val_bs=None, shuffle_train=True, device='cpu', **kwargs):
train_ds = NumpyDataset(to.train)
valid_ds = NumpyDataset(to.valid)
val_bs = bs if val_bs is None else val_bs
train = NumpyDataLoader(train_ds, bs=bs, shuffle=shuffle_train, device=device, drop_last=True, **kwargs)
valid = NumpyDataLoader(valid_ds, bs=val_bs, shuffle=False, device=device, **kwargs)
super().__init__(train, valid, device=device, **kwargs)
df_np = df.to_numpy()
col_names = df.columns
idx_2_col = dict(zip(range(len(col_names)), col_names)); idx_2_col
col_2_idx = {v: k for k, v in idx_2_col.items()}
cat_idxs = [col_2_idx[name] for name in cat_names]
cont_idxs = [col_2_idx[name] for name in cont_names]
y_idxs = [col_2_idx[name] for name in [y_names]]
class NumpyFillMissing(TabularProc):
def __init__(self, fill_strategy=FillStrategy.median, add_col=True, fill_vals=None):
if fill_vals is None: fill_vals = defaultdict(int)
store_attr(self, 'fill_strategy,add_col,fill_vals')
def setups(self, dsets):
missing = [np.isnan(np.sum(df_np[:,idx])) for idx in cont_idxs]
self.na_dict = {n:self.fill_strategy(dsets[n])}
fs = FillStrategy.median
missing = [np.isnan(np.sum(df_np[:,idx])) for idx in cont_idxs]; missing
missing = {}
for idx in cont_idxs:
missing[idx] = np.isnan(np.sum(df_np[:,idx]))
missing
missing
fs
missing.keys()
{n:fs(df_np[:,n], defaultdict(int))
for n in missing.keys()}
np.all(df_np[:,cont_idxs], axis=1).isnan()
df_np[:,cont_idxs]
class TabularNumpy(CollBase, GetAttr, FilteredBase):
_default, with_cont='procs',True
def __init__(self, df):
super().__init__(df)
t_np = TabularNumpy(df_np)
len(t_np)
class TabularNumpy(Tabular):
_default, with_cont='procs',True
def __init__(self, df, procs=None, cat_names=None, cont_names=None, y_names=None, y_block=None, splits=None,
do_setup=True, device=None, inplace=False, reduce_memory=True):
self.df = df[cat_names+cont_names+y_names].to_numpy()
self.idx_2_col = dict(zip(range(len(df.columns)), df.columns))
self.col_2_idx = {v: k for k, v in self.idx_2_col.items()}
class Tabular(CollBase, GetAttr, FilteredBase):
"A `DataFrame` wrapper that knows which cols are cont/cat/y, and returns rows in `__getitem__`"
_default,with_cont='procs',True
def __init__(self, df, procs=None, cat_names=None, cont_names=None, y_names=None, y_block=None, splits=None,
do_setup=True, device=None, inplace=False, reduce_memory=True):
if inplace and splits is not None and pd.options.mode.chained_assignment is not None:
warn("Using inplace with splits will trigger a pandas error. Set `pd.options.mode.chained_assignment=None` to avoid it.")
if not inplace: df = df.copy()
if splits is not None: df = df.iloc[sum(splits, [])]
self.dataloaders = delegates(self._dl_type.__init__)(self.dataloaders)
super().__init__(df)
self.y_names,self.device = L(y_names),device
if y_block is None and self.y_names:
# Make ys categorical if they're not numeric
ys = df[self.y_names]
if len(ys.select_dtypes(include='number').columns)!=len(ys.columns): y_block = CategoryBlock()
else: y_block = RegressionBlock()
if y_block is not None and do_setup:
if callable(y_block): y_block = y_block()
procs = L(procs) + y_block.type_tfms
self.cat_names,self.cont_names,self.procs = L(cat_names),L(cont_names),Pipeline(procs)
self.split = len(df) if splits is None else len(splits[0])
if reduce_memory:
if len(self.cat_names) > 0: self.reduce_cats()
if len(self.cont_names) > 0: self.reduce_conts()
if do_setup: self.setup()
def new(self, df):
return type(self)(df, do_setup=False, reduce_memory=False, y_block=TransformBlock(),
**attrdict(self, 'procs','cat_names','cont_names','y_names', 'device'))
def subset(self, i): return self.new(self.items[slice(0,self.split) if i==0 else slice(self.split,len(self))])
def copy(self): self.items = self.items.copy(); return self
def decode(self): return self.procs.decode(self)
def decode_row(self, row): return self.new(pd.DataFrame(row).T).decode().items.iloc[0]
def reduce_cats(self): self.train[self.cat_names] = self.train[self.cat_names].astype('category')
def reduce_conts(self): self[self.cont_names] = self[self.cont_names].astype(np.float32)
def show(self, max_n=10, **kwargs): display_df(self.new(self.all_cols[:max_n]).decode().items)
def setup(self): self.procs.setup(self)
def process(self): self.procs(self)
def loc(self): return self.items.loc
def iloc(self): return _TabIloc(self)
def targ(self): return self.items[self.y_names]
def x_names (self): return self.cat_names + self.cont_names
def n_subsets(self): return 2
def y(self): return self[self.y_names[0]]
def new_empty(self): return self.new(pd.DataFrame({}, columns=self.items.columns))
def to_device(self, d=None):
self.device = d
return self
def all_col_names (self):
ys = [n for n in self.y_names if n in self.items.columns]
return self.x_names + self.y_names if len(ys) == len(self.y_names) else self.x_names
| 00_Numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon reviews : Sentiment analysis
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Introduction" data-toc-modified-id="Introduction-1"><span class="toc-item-num">1 </span>Introduction</a></span></li><li><span><a href="#Data-Analysis" data-toc-modified-id="Data-Analysis-2"><span class="toc-item-num">2 </span>Data Analysis</a></span><ul class="toc-item"><li><span><a href="#Top-10-most-reviewed-products" data-toc-modified-id="Top-10-most-reviewed-products-2.1"><span class="toc-item-num">2.1 </span>Top 10 most reviewed products</a></span></li><li><span><a href="#Top-10-best-brand" data-toc-modified-id="Top-10-best-brand-2.2"><span class="toc-item-num">2.2 </span>Top 10 best brand</a></span></li><li><span><a href="#Top-10-worst-products" data-toc-modified-id="Top-10-worst-products-2.3"><span class="toc-item-num">2.3 </span>Top 10 worst products</a></span></li><li><span><a href="#Best-budget-product" data-toc-modified-id="Best-budget-product-2.4"><span class="toc-item-num">2.4 </span>Best budget product</a></span></li><li><span><a href="#Best-high-end-product" data-toc-modified-id="Best-high-end-product-2.5"><span class="toc-item-num">2.5 </span>Best high end product</a></span></li><li><span><a href="#Price-vs-Rating-distribution" data-toc-modified-id="Price-vs-Rating-distribution-2.6"><span class="toc-item-num">2.6 </span>Price vs Rating distribution</a></span></li></ul></li><li><span><a href="#Model" data-toc-modified-id="Model-3"><span class="toc-item-num">3 </span>Model</a></span><ul class="toc-item"><li><span><a href="#Feature-extraction-from-text" data-toc-modified-id="Feature-extraction-from-text-3.1"><span class="toc-item-num">3.1 </span>Feature extraction from text</a></span><ul class="toc-item"><li><span><a href="#Tfidf" data-toc-modified-id="Tfidf-3.1.1"><span class="toc-item-num">3.1.1 </span>Tfidf</a></span></li></ul></li><li><span><a href="#Logistic-Regression" data-toc-modified-id="Logistic-Regression-3.2"><span class="toc-item-num">3.2 </span>Logistic Regression</a></span></li><li><span><a href="#n-grams" data-toc-modified-id="n-grams-3.3"><span class="toc-item-num">3.3 </span>n-grams</a></span></li></ul></li></ul></div>
# ## Introduction
# About this Dataset
# Context
#
# PromptCloud extracted 400 thousand reviews of unlocked mobile phones sold on Amazon.com to find out insights with respect to reviews, ratings, price and their relationships.
# Content
#
# Given below are the fields:
#
# Product Title
# Brand
# Price
# Rating
# Review text
# Number of people who found the review helpful
#
# Data was acquired in December, 2016 by the crawlers build to deliver our data extraction services.
# Initial Analysis
#
# It can be accessed here: http://www.kdnuggets.com/2017/01/data-mining-amazon-mobile-phone-reviews-interesting-insights.html
# The goal of this notebook is to predict wheter a review is positive or negative using Logistic Regression, MLP and NN
# **Data preparation**
#
# First read the dataset in panda dataframe. We are interseted in positive and negative reviews only. No such column exist in dataframe.
#
# To create the column we will remove neutral rating i.e. 3. After that values those are below 3 will be treated negative review and above 3 will be treated as positive review.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import pandas as pd
plt.style.use('fivethirtyeight')
# Read in the data
df = pd.read_csv('../input/amazon-reviews-unlocked-mobile-phones/Amazon_Unlocked_Mobile.csv')
df.head()
# -
# ## Data Analysis
# ### Top 10 most reviewed products
# +
mostreviewd = (df.set_index('Product Name').groupby(level=0)['Reviews']
.agg(['count'])).sort_values(['count'], ascending=False)[:10]
plt.figure(figsize=(12, 8))
sns.barplot(mostreviewd.reset_index().index, y=mostreviewd['count'], hue=mostreviewd.index.str[:50] + '...', dodge=False)
plt.ylim(1000,)
plt.xticks([]);
plt.ylabel('Reviews count')
plt.title('Top 10 most reviewed products');
# -
# ### Top 10 best brand
# +
bestbrand = (df[df['Rating'] > 3].set_index('Brand Name').groupby(level=0)['Reviews'].
agg(['count'])).sort_values(['count'], ascending=False)[:10]
plt.figure(figsize=(12, 8))
sns.barplot(bestbrand.index, y=bestbrand['count'], hue=bestbrand.index, dodge=False)
plt.legend([])
plt.ylabel('Positive reviews count')
plt.title('Top 10 best brand');
# -
# ### Top 10 worst products
# +
# Filter out rating above 3 and get the review count
worstproduct = (df[df['Rating'] < 3].set_index('Product Name').groupby(level=0)['Reviews'].
agg(['count'])).sort_values(['count'], ascending=False)[:10]
plt.figure(figsize=(12, 8))
sns.barplot(worstproduct.reset_index().index, y=worstproduct['count'], hue=worstproduct.index.str[:50] + '...', dodge=False)
plt.ylim(250,)
plt.xticks([]);
plt.ylabel('Negative reviews count')
plt.title('Top 10 worst products');
# -
# ### Best budget product
# +
## Best budget product
budget = (df[(df['Rating'] > 3) & (df['Price'] < 500)].set_index('Product Name').groupby(level=0)['Price'].
agg(['count'])).sort_values(['count'], ascending=False)[:10]
grouped = df.set_index('Product Name').loc[budget.index].groupby(level=0)
price = pd.Series(index = budget.index)
for name, group in grouped:
price.loc[name] = group.Price.iloc[0]
budget['Price'] = price
budget.reset_index(inplace=True)
plt.figure(figsize=(12, 8))
sns.barplot(x='Price', y='count', dodge=False, hue='Product Name', data=budget, palette=sns.color_palette("cubehelix", 12))
plt.ylim(750,)
plt.ylabel('Positive reviews count')
plt.title('Best budget products under $500');
# Put the legend out of the figure
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.);
# -
# ### Best high end product
# +
highend = (df[(df['Rating'] > 3) & (df['Price'] > 900)].set_index('Product Name').groupby(level=0)['Price'].
agg(['count'])).sort_values(['count'], ascending=False)[:10]
grouped = df.set_index('Product Name').loc[highend.index].groupby(level=0)
price = pd.Series(index = budget.index)
for name, group in grouped:
price.loc[name] = group.Price.iloc[0]
highend['Price'] = price
highend.reset_index(inplace=True)
plt.figure(figsize=(8, 8))
sns.barplot(x='Price', y='count', dodge=False, hue='Product Name', data=highend, palette=sns.color_palette("cubehelix", 12))
plt.ylabel('Positive reviews count')
plt.title('Best high end products under $2000');
# Put the legend out of the figure
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.);
# -
# ### Price vs Rating distribution
plt.figure(figsize=(10,8))
sns.violinplot(x="Rating", y="Price", data=df)
plt.title('Price vs Rating distribution');
# ## Model
# +
# Drop missing values
df.dropna(inplace=True)
# Remove any 'neutral' ratings equal to 3
df = df[df['Rating'] != 3]
# Encode 4s and 5s as 1 (rated positively)
# Encode 1s and 2s as 0 (rated poorly)
df['Positively Rated'] = np.where(df['Rating'] > 3, 1, 0)
df.head(10)
# -
df.describe()
# As you can see most of the reviews are positive.
# +
# Get training and test data from dataset.
from sklearn.model_selection import train_test_split
# Split data into training and test sets
X_train, X_test, y_train, y_test = train_test_split(df['Reviews'],
df['Positively Rated'],
random_state=0)
# -
# ### Feature extraction from text
# The main and only feature for this model is **Review**. We will be parsing review and train the model. Finally, model should be able to predict whether review is positive or negative.
#
# * Feature : Reviews
# * Target : Positively Rated
#
# #### Tfidf
#
# Convert a collection of raw documents to a matrix of TF-IDF features. https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html
#
#
# +
from sklearn.feature_extraction.text import TfidfVectorizer
# Fit the TfidfVectorizer to the training data specifiying a minimum document frequency of 5
vect = TfidfVectorizer(min_df=5).fit(X_train)
len(vect.get_feature_names())
# -
# TfidfVectorizer created 17951 features from review text. Now we can feed this features to out model. Lets see top features extracted by TfidfVectorizer
X_train_vectorized = vect.transform(X_train)
# +
feature_names = np.array(vect.get_feature_names())
sorted_tfidf_index = X_train_vectorized.max(0).toarray()[0].argsort()
print('Smallest tfidf:\n{}\n'.format(feature_names[sorted_tfidf_index[:10]]))
print('Largest tfidf: \n{}'.format(feature_names[sorted_tfidf_index[:-11:-1]]))
# -
# ### Logistic Regression
# +
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import roc_auc_score
X_train_vectorized = vect.transform(X_train)
model = LogisticRegression(solver='saga')
model.fit(X_train_vectorized, y_train)
predictions = model.predict(vect.transform(X_test))
print('AUC: ', roc_auc_score(y_test, model.decision_function(vect.transform(X_test))))
# -
# Our model's roc score is very good. Below are the lists of words from Logistic Regression model coefficiants
# +
sorted_coef_index = model.coef_[0].argsort()
print('Smallest Coefs:\n{}\n'.format(feature_names[sorted_coef_index[:10]]))
print('Largest Coefs: \n{}'.format(feature_names[sorted_coef_index[:-11:-1]]))
# +
from wordcloud import WordCloud, STOPWORDS
import matplotlib.pyplot as plt
def PlotWordCloud(words, title):
wordcloud = WordCloud(width = 800, height = 800,
background_color ='white'
).generate(words)
# plot the WordCloud image
plt.figure(figsize = (10, 10), facecolor = None)
plt.imshow(wordcloud)
plt.axis("off")
plt.tight_layout(pad = 0)
plt.title(title, fontsize=50)
plt.show()
# -
negative = ''
for word in feature_names[sorted_coef_index[:100]]:
negative += word + ' '
PlotWordCloud(negative, 'Most negative words')
positive = ''
for word in feature_names[sorted_coef_index[:-101:-1]]:
positive += word + ' '
PlotWordCloud(positive, 'Most positive words')
# Model is wroking as expected. Lets try to give it some difficult reviews
print(model.predict(vect.transform(['not an issue, phone is working',
'an issue, phone is not working'])))
# If you can notice that our model is predicting both the reviews as negative. It is only considering single word. Now lets make it understand two word combination.
# ### n-grams
# +
# extracting 1-grams and 2-grams
vect = TfidfVectorizer(min_df=5, ngram_range=(1,2)).fit(X_train)
X_train_vectorized = vect.transform(X_train)
len(vect.get_feature_names())
# -
# The features count reached to whopping 198917 from 17951. Lets train the model again with new features
# +
model = LogisticRegression(solver='saga')
model.fit(X_train_vectorized, y_train)
predictions = model.predict(vect.transform(X_test))
print('AUC: ', roc_auc_score(y_test, model.decision_function(vect.transform(X_test))))
# +
feature_names = np.array(vect.get_feature_names())
sorted_coef_index = model.coef_[0].argsort()
print('Smallest Coefs:\n{}\n'.format(feature_names[sorted_coef_index[:10]]))
print('Largest Coefs: \n{}'.format(feature_names[sorted_coef_index[:-11:-1]]))
# -
print(model.predict(vect.transform(['not an issue, phone is working',
'an issue, phone is not working'])))
# Thats exactly we want our model should predict. It is now able to diffrentiate reviews based on tow words combination.
| Sentiment analysis with Logistic Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# +
import random
import pandas as pd
class Main():
def __init__(self):
self.c1 = "Ecuador"
self.c2 = "Singapore"
self.c3 = "Portugal"
self.People = []
self.Stocks = []
self.WealthC1 = []
self.WealthC2 = []
self.WealthC3 = []
self.Avoid = False
self.x = []
def create_population(self, size, nationality, chance):
for i in range(0, size):
person = People()
person.chance = chance
person.nationality = nationality
self.People.append(person)
def create_stocks(self, size):
for i in range(0, size):
stock = Stocks()
self.Stocks.append(stock)
def FilterSort(self, c):
lst = list(filter(lambda person: person.nationality == c, self.People))
lst.sort(key=lambda person: person.money, reverse=True)
return lst
def AvoidIt(self, chance, risk):
if (chance <= risk):
self.Avoid = False
else:
self.Avoid = True
def Round(self, rounds):
for i in range(0, rounds):
self.x.append(i)
self.WealthC1.append(self.addwealth(self.c1))
self.WealthC2.append(self.addwealth(self.c2))
self.WealthC3.append(self.addwealth(self.c3))
for person in self.People:
if (person.broke == False):
if (person.encounterchance <= 0.7):
person.encounterchance = random.random()
stock = random.choice(self.Stocks)
self.AvoidIt(person.chance, stock.risk)
if (self.Avoid):
stock.encounter += 1
person.Trade(stock.risk)
person.Broke()
def addwealth(self, nationality):
Wealth = 0
for person in self.People:
# if(person.broke == False):
if (person.nationality == nationality):
Wealth += person.money
return round(Wealth)
def addwealthC(self, lst):
Wealth = 0
for person in lst:
Wealth += person.money
return round(Wealth)
def results(self):
c1p = self.FilterSort(self.c1)
c2p = self.FilterSort(self.c2)
c3p = self.FilterSort(self.c3)
topbottom = int(len(c1p)*(10/100))
richestc1 = 0
richestc2 = 0
richestc3 = 0
for person in self.People:
if (person.nationality == self.c1):
if (person.money > richestc1):
richestc1 = round(person.money)
if (person.nationality == self.c2):
if (person.money > richestc2):
richestc2 = round(person.money)
if (person.nationality == self.c3):
if (person.money > richestc3):
richestc3 = round(person.money)
print('amount of money left {0} : {1}'.format(self.c1, self.addwealth(self.c1)))
print('Richest 10% {0} : {1}'.format(self.c1, self.addwealthC(c1p[:topbottom])))
print('Poorest 10% {0} : {1}'.format(self.c1, self.addwealthC(c1p[-topbottom:])))
print('Richest {0} : {1}'.format(self.c1, richestc1))
print('amount of money left {0}: {1}'.format(self.c2, self.addwealth(self.c2)))
print('Richest 10% {0} : {1}'.format(self.c2, self.addwealthC(c2p[:topbottom])))
print('Poorest 10% {0} : {1}'.format(self.c2, self.addwealthC(c2p[-topbottom:])))
print('Richest {0} : {1}'.format(self.c2, richestc2))
print('amount of money left {0} : {1}'.format(self.c3, self.addwealth(self.c3)))
print('Richest 10% {0} : {1}'.format(self.c3, self.addwealthC(c3p[:topbottom])))
print('Poorest 10% {0} : {1}'.format(self.c3, self.addwealthC(c3p[-topbottom:])))
print('Richest {0} : {1}'.format(self.c3, richestc3))
print('amount of People who are broke in {0} : {1}'.format(self.c1, len(list(filter(lambda person: person.broke == True and person.nationality == self.c1, self.People)))))
print('amount of People who are broke in {0} : {1}'.format(self.c2, len(list(filter(lambda person: person.broke == True and person.nationality == self.c2, self.People)))))
print('amount of People who are broke in {0} : {1}'.format(self.c3, len(list(filter(lambda person: person.broke == True and person.nationality == self.c3, self.People)))))
self.y = {self.c1: self.WealthC1, self.c2: self.WealthC2, self.c3: self.WealthC3}
self.y2 = {self.c1: self.WealthC1[:10], self.c2: self.WealthC2[:10], self.c3: self.WealthC3[:10]}
self.y3 = {self.c1: self.WealthC1[:20], self.c2: self.WealthC2[:20], self.c3: self.WealthC3[:20]}
self.y4 = {self.c1: self.WealthC1[:50], self.c2: self.WealthC2[:50], self.c3: self.WealthC3[:50]}
self.y5 = {self.c1: self.WealthC1[:100], self.c2: self.WealthC2[:100], self.c3: self.WealthC3[:100]}
graph = pd.DataFrame(self.y, self.x)
graph.plot(kind='line', grid=True, title="Total wealth per round", ylabel="wealth", xlabel="rounds")
graph2 = pd.DataFrame(self.y2, self.x[:10])
graph2.plot(kind='line', grid=True, title="Total wealth per round", ylabel="wealth", xlabel="rounds")
graph3 = pd.DataFrame(self.y3, self.x[:20])
graph3.plot(kind='line', grid=True, title="Total wealth per round", ylabel="wealth", xlabel="rounds")
graph4 = pd.DataFrame(self.y4, self.x[:50])
graph4.plot(kind='line', grid=True, title="Total wealth per round", ylabel="wealth", xlabel="rounds")
graph5 = pd.DataFrame(self.y5, self.x[:100])
graph5.plot(kind='line', grid=True, title="Total wealth per round", ylabel="wealth", xlabel="rounds")
c1wealth = []
c2wealth = []
c3wealth = []
for person in self.People:
if (person.nationality == self.c1):
c1wealth.append(person.money)
if (person.nationality == self.c2):
c2wealth.append(person.money)
if (person.nationality == self.c3):
c3wealth.append(person.money)
data={self.c1:c1wealth}
grapfscat1 = pd.DataFrame(data = data)
grapfscat1.plot.scatter(x = self.c1, y = self.c1, s = 20, title=self.c1)
grapfscat1.hist(bins=40)
data={self.c2:c2wealth}
grapfscat2 = pd.DataFrame(data = data)
grapfscat2.plot.scatter(x = self.c2, y = self.c2, s = 20, title=self.c2)
grapfscat2.hist(bins=40)
data={self.c3:c3wealth}
grapfscat3 = pd.DataFrame(data = data)
grapfscat3.plot.scatter(x = self.c3, y = self.c3, s = 20, title=self.c3)
grapfscat3.hist(bins=40)
def simulate(self):
self.create_population(1000000, self.c1, 0.33)
self.create_population(1000000, self.c2, 0.92)
self.create_population(1000000, self.c3, 0.01)
random.shuffle(self.People)
self.create_stocks(100)
self.Round(1000)
self.results()
class People():
def __init__(self):
self.chance = random.uniform(0.3,0.6)
self.money = 1000
self.broke = False
self.encounterchance = random.random()
self.nationality = "nothing"
def Broke(self):
if (self.money <= 0):
self.broke = True
def Trade(self, risk):
rng = random.random()
if (rng < risk):
self.money = self.money - ((self.money*1.5) * risk)
else:
self.money = self.money + ((self.money*1.5) * risk)
class Stocks():
def __init__(self):
self.risk = random.random()
self.encounter = 0
if __name__ == "__main__":
main = Main()
main.simulate()
| main-graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/Gregory-Eales/NeuralNet-Digit-Recognition/blob/master/Digit_Recognition.ipynb)
# + id="ii2wESXNnl-S" colab_type="code" colab={}
# import dependencies
import numpy as np
from sklearn import datasets
from matplotlib import pyplot as plt
# + id="iaZ1bHtRrkIe" colab_type="code" colab={}
# load digit data set
x, uncleaned_y = datasets.load_digits(return_X_y=True)
x = x/10
# create a y for each classification: numbers 0-9 and stores it in 'answers'
answers = []
for i in range(10):
zero = [0, 0, 0, 0, 0, 0, 0, 0, 0, 0]
zero[i] = 1
answers.append(zero)
# iterate through 'uncleaned_y' and add the correct classification for each y
y = []
for i in uncleaned_y:
for j in range(10):
if i == j:
y.append(answers[j])
# convert y to an array
y = np.array(y)
# + id="BBZvWKSyQDuV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="2694a425-a43e-4a60-f2eb-4d2437ffcc48"
print()
# + id="HVLUZ-IxoKjL" colab_type="code" colab={}
# define neural network model
class NeuralNetwork(object):
def __init__(self, x, y, alpha=0.1, iterations=1000, num_layers=3, hidden_addition=1):
# initiate class properties
self.x = x
self.y = y
self.alpha = alpha
self.iterations = iterations
self.num_layers = num_layers
self.w = {}
self.b = {}
self.z = {}
self.a = {}
self.e = {}
self.historical_cost = []
# create layer weights and layer variables
for i in range(1, self.num_layers+hidden_addition):
# if not the last weight initiate with eacher layer one bigger than the input size of x
if i != self.num_layers:
if i == 1:
self.w["w"+str(i)] = np.random.randn(self.x.shape[1], self.x.shape[1]+hidden_addition)*0.5
self.b["b"+str(i)] = np.random.randn(1, self.x.shape[1]+hidden_addition)*0.5
else:
self.w["w"+str(i)] = np.random.randn(self.x.shape[1]+hidden_addition, self.x.shape[1]+hidden_addition)*0.5
self.b["b"+str(i)] = np.random.randn(1, self.x.shape[1]+hidden_addition)*0.5
# if the last weight initiate with output = dimensions of y
else:
self.w["w"+str(i)] = np.random.randn(self.x.shape[1]+hidden_addition, self.y.shape[1])*0.1
self.b["b"+str(i)] = np.random.randn(1, self.y.shape[1])*0.1
# doesnt matter, will be changed later
self.z["z" + str(i)] = np.zeros([self.x.shape[0], self.x.shape[1]])
self.a["a" + str(i)] = np.zeros([self.x.shape[0], self.x.shape[1]])
# calculate and make predictions
def forward_propagation(self):
# initiate forward propigation with dotting x an w1
self.a['a0'] = self.x
self.z["z1"] = np.dot(self.x, self.w["w1"]) + self.b['b1']
self.a['a1'] = np.tanh(self.z['z1'])
# iterate through all dots and activations
for i in range(2, self.num_layers):
self.z["z" + str(i)] = np.dot(self.a['a'+str(i-1)], self.w['w' + str(i)]) + self.b['b' + str(i)]
self.a['a'+ str(i)] = np.tanh(self.z["z" + str(i)])
# on the last iteration use sigmoid instead of tanh for classification
self.z["z" + str(self.num_layers)] = np.dot(self.a['a'+str(self.num_layers-1)], self.w['w' + str(self.num_layers)]) + self.b['b' + str(self.num_layers)]
self.a['a'+ str(self.num_layers)] = self.sigmoid(self.z["z" + str(self.num_layers)])
return self.a['a'+str(self.num_layers)]
# adjust weights based on cost function
def backward_propagation(self):
self.create_updates()
#self.w['w3'] = self.w['w3'] - (self.alpha/self.x.shape[0])*np.dot((self.sigmoid_prime(self.z['z3'])*self.j_prime()).T, self.a['a2']).T
# iterate throught weights
for i in reversed(range(1, self.num_layers+1)):
self.w["w"+str(i)] = self.w["w"+str(i)] - (self.alpha/self.x.shape[0])*np.dot(self.a['a'+str(i-1)].T, self.e['e'+str(i)])
self.b["b"+str(i)] = self.b["b"+str(i)] - (self.alpha/self.x.shape[0])*np.sum(self.e['e'+str(i)], axis=0)
# returns the derivative of the cost
def j_prime(self):
return (self.y/self.a['a' + str(self.num_layers)] - (1-self.y)/(1-self.a['a' + str(self.num_layers)]))
# creates update cache
def create_updates(self):
self.e['e'+str(self.num_layers)] = self.j_prime()*self.sigmoid_prime(self.z['z'+str(self.num_layers)])
for i in reversed(range(1, self.num_layers)):
self.e['e'+str(i)] = np.dot(self.e['e' + str(i+1)], self.w['w' + str(i+1)].T)*self.tanh_prime(self.z['z'+str(i)])
# optimize model based on inputes
def optimize(self):
for i in range(self.iterations):
if i%100 == 0:
print(str(i/10) + "%")
self.forward_propagation()
self.backward_propagation()
self.historical_cost.append(self.cost_function())
print("Complete")
# calculate the cost of prections
def cost_function(self):
j = -(np.sum(self.y*np.log(self.a['a'+str(self.num_layers)]) + (1-self.y)*np.log(1 - self.a['a'+ str(self.num_layers)]))/self.x.shape[0])
return j
# sigmoid activation function
def sigmoid(self, z):
return 1/(1+np.exp(-z))
# derivative of sigmoid activation function
def sigmoid_prime(self, z):
return -np.exp(-z)/np.square(1 + np.exp(-z))
# derivative of tanh activation function
def tanh_prime(self, z):
return 1 - np.square(np.tanh(z))
# + id="jHXVy8LzoKtD" colab_type="code" colab={}
NN = NeuralNetwork(x[0:1700], y[0:1700], alpha=0.12, iterations=1000, num_layers=5, hidden_addition=1)
# + id="cPBk0gwO_qC8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="d13835ca-3535-49c4-9be9-02203f052e3b"
NN.optimize()
# + id="Ci8LLlJxL0Pi" colab_type="code" colab={}
x_axis = []
for i in range(NN.iterations):
x_axis.append(i)
# + id="BuVnjCcEoK0j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 724} outputId="32cc5b8d-8b5b-45db-cfac-c6755638896a"
plt.plot(x_axis, NN.historical_cost)
plt.show()
print(NN.historical_cost)
NN.x = x[1700:x.shape[0]]
NN.y = y[1700:x.shape[0]]
predictions = (NN.forward_propagation())
p = np.zeros_like(predictions)
p[np.arange(len(predictions)), predictions.argmax(1)] = 1
print(p[30:40])
print(y[30:40])
# + id="CIBXyZO68hRr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="32e4d7fa-4409-46cf-e095-8f1231506b85"
correct = 0
for i in range(len(p)):
if p.tolist()[i] == y[1700:y.shape[0]].tolist()[i]:
correct += 1
print("The model is " + str((correct/len(p))*100) + "% accurate")
| Digit_Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
#
#
# # Attenuation
#
# Attenuation is the phenomenon of light's intensity being gradually dampened as
# it propagates through a medium. In PyVista positional lights can show attenuation.
# The quadratic attenuation model uses three parameters to describe attenuation:
# a constant, a linear and a quadratic parameter. These parameters
# describe the decrease of the beam intensity as a function of the distance, `I(r)`.
# In a broad sense the constant, linear and quadratic components correspond to
# `I(r) = 1`, `I(r) = 1/r` and `I(r) = 1/r^2` decay of the intensity with distance
# from the point source. In all cases a larger attenuation value (of a given kind)
# means stronger dampening (weaker light at a given distance).
#
# So the constant attenuation parameter corresponds roughly to a constant intensity
# component. The linear and the quadratic attenuation parameters correspond to intensity
# components that decay with distance from the source. For the same parameter value the
# quadratic attenuation produces a beam that is shorter in range than that produced
# by linear attenuation.
#
# Three spotlights with three different attenuation profiles each:
#
#
# +
# sphinx_gallery_thumbnail_number = 3
import pyvista as pv
plotter = pv.Plotter(lighting='none')
billboard = pv.Plane(direction=(1, 0, 0), i_size=6, j_size=6)
plotter.add_mesh(billboard, color='white')
all_attenuation_values = [(1, 0, 0), (0, 2, 0), (0, 0, 2)]
offsets = [-2, 0, 2]
for attenuation_values, offset in zip(all_attenuation_values, offsets):
light = pv.Light(position=(0.1, offset, 2), focal_point=(0.1, offset, 1), color='cyan')
light.positional = True
light.cone_angle = 20
light.intensity = 15
light.attenuation_values = attenuation_values
plotter.add_light(light)
plotter.view_yz()
plotter.show()
# -
# It's not too obvious but it's visible that the rightmost light with quadratic
# attenuation has a shorter range than the middle one with linear attenuation.
# Although it seems that even the leftmost light with constant attenuation loses
# its brightness gradually, this partly has to do with the fact that we sliced
# the light beams very close to their respective axes, meaning that light hits
# the surface in a very small angle. Altering the scene such that the lights
# are further away from the plane changes this:
#
#
# +
plotter = pv.Plotter(lighting='none')
billboard = pv.Plane(direction=(1, 0, 0), i_size=6, j_size=6)
plotter.add_mesh(billboard, color='white')
all_attenuation_values = [(1, 0, 0), (0, 2, 0), (0, 0, 2)]
offsets = [-2, 0, 2]
for attenuation_values, offset in zip(all_attenuation_values, offsets):
light = pv.Light(position=(0.5, offset, 3), focal_point=(0.5, offset, 1), color='cyan')
light.positional = True
light.cone_angle = 20
light.intensity = 15
light.attenuation_values = attenuation_values
plotter.add_light(light)
plotter.view_yz()
plotter.show()
# -
# Now the relationship of the three kinds of attenuation seems clearer.
#
# For a more practical comparison, let's look at planes that are perpendicular
# to the axis of each light (making use of the fact that shadowing between
# objects is not handled by default):
#
#
# +
plotter = pv.Plotter(lighting='none')
# loop over three lights with three kinds of attenuation
all_attenuation_values = [(2, 0, 0), (0, 2, 0), (0, 0, 2)]
light_offsets = [-6, 0, 6]
for attenuation_values, light_x in zip(all_attenuation_values, light_offsets):
# loop over three perpendicular planes for each light
for plane_y in [2, 5, 10]:
screen = pv.Plane(center=(light_x, plane_y, 0), direction=(0, 1, 0),
i_size=5, j_size=5)
plotter.add_mesh(screen, color='white')
light = pv.Light(position=(light_x, 0, 0), focal_point=(light_x, 1, 0),
color='cyan')
light.positional = True
light.cone_angle = 15
light.intensity = 5
light.attenuation_values = attenuation_values
light.show_actor()
plotter.add_light(light)
plotter.view_vector((1, -2, 2))
plotter.show()
| locale/examples/04-lights/attenuation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Feature Engineering
# ## Textual Data
#
# This notebook show cases some of the common methods for feature extraction and engineering on textual data.
# ## Important Imports
# +
import numpy as np
import pandas as pd
from collections import Counter
# pandas display data frames as tables
from IPython.display import display, HTML
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
# -
# ---
#
# ## Prepare a Sample Corpus
# +
corpus = ['pack my box with five dozen liquor jugs.',
'pack my box',
'the quick brown fox jumps over the lazy dog.',
'the brown fox is quick and the blue dog is lazy',
'pack my box with five dozen liquor jugs and biscuits',
'the dog is lazy but the brown fox is quick']
labels = ['picnic', 'picnic', 'animals', 'animals', 'picnic', 'animals']
corpus = np.array(corpus)
corpus_df = pd.DataFrame({'document': corpus,
'category': labels})
corpus_df = corpus_df[['document', 'category']]
corpus_df
# -
# ## Bag of Words
# +
cv = CountVectorizer(min_df=0., max_df=1.)
cv_matrix = cv.fit_transform(corpus_df.document)
cv_matrix = cv_matrix.toarray()
vocab = cv.get_feature_names()
pd.DataFrame(cv_matrix, columns=vocab)
# -
# ## TF-IDF
# +
tv = TfidfVectorizer(min_df=0., max_df=1., use_idf=True)
tv_matrix = tv.fit_transform(corpus_df.document)
tv_matrix = tv_matrix.toarray()
vocab = tv.get_feature_names()
pd.DataFrame(np.round(tv_matrix, 2), columns=vocab)
# -
# ## N-Gram Vectorizer
bv = CountVectorizer(ngram_range=(2,2))
bv_matrix = bv.fit_transform(corpus_df.document)
bv_matrix = bv_matrix.toarray()
vocab = bv.get_feature_names()
pd.DataFrame(bv_matrix, columns=vocab)
| notebooks/Ch01 - Machine Learning Fundamentals/feature_engineering_text_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="wJ4LiB_So76e"
# # All
# + id="EGawA05dBnAA" executionInfo={"status": "ok", "timestamp": 1628780168791, "user_tz": 240, "elapsed": 426, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}}
from google.colab import auth
auth.authenticate_user()
# + colab={"base_uri": "https://localhost:8080/"} id="D-bm3dCglIX5" executionInfo={"status": "ok", "timestamp": 1628779089785, "user_tz": 240, "elapsed": 124, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="dc4887f7-28c8-495f-f0bf-662b74d9525d"
from google.colab import drive
drive.mount('/content/drive')
# + id="3NQieOHYlJrb"
# %%capture
# !pip install datasets
# + id="2WELNTlBBrQe" colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"status": "ok", "timestamp": 1628780249499, "user_tz": 240, "elapsed": 34099, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="fc57f227-208d-42d7-ac2c-505b3822efe5"
# # %%capture
# !pip install --force-reinstall git+https://github.com/CaitlinJCorbin/transformers
# + id="0W7MvbTjB2fp" executionInfo={"status": "ok", "timestamp": 1628780263131, "user_tz": 240, "elapsed": 3922, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}}
from transformers import EncoderDecoderModel, DistilBertTokenizer, DistilBertConfig, DistilBertModel
import torch
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="ypPLSYBqCTMU" executionInfo={"status": "ok", "timestamp": 1628780264503, "user_tz": 240, "elapsed": 136, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="8d777e4d-f404-40c7-f1e4-d75ef6a5ed98"
if torch.cuda.is_available():
device = torch.device('cuda')
print(torch.cuda.get_device_name())
else:
device = torch.device('cpu')
# + [markdown] id="qN4eZVPqPYAl"
# ## Data #1
#
# + colab={"base_uri": "https://localhost:8080/"} id="dWic3hohnmhD" executionInfo={"status": "ok", "timestamp": 1628780271783, "user_tz": 240, "elapsed": 1643, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="e2887ebd-17e1-49bb-9e26-7f155ad999b7"
# !gsutil -m cp /content/drive/MyDrive/immunolinguistics/frames_converted/*otsu.*.txt.gz .
# + colab={"base_uri": "https://localhost:8080/"} id="qo0RnK7WnpX-" executionInfo={"status": "ok", "timestamp": 1628780271915, "user_tz": 240, "elapsed": 135, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="74e6563a-b885-48d5-e5f0-561463309ee2"
# !gzip -d *.gz
# + colab={"base_uri": "https://localhost:8080/"} id="srgR52x6nrAM" executionInfo={"status": "ok", "timestamp": 1628780273361, "user_tz": 240, "elapsed": 1447, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="c171835f-898c-4c3e-f5cb-263d1677393b"
# !mkdir otsu
# !gsutil -m cp *txt otsu/
# + id="NGciYRaKnsCf" colab={"base_uri": "https://localhost:8080/", "height": 253} executionInfo={"status": "error", "timestamp": 1628780288011, "user_tz": 240, "elapsed": 246, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="3807e851-8374-4480-d16e-041b5833ef3b"
import os
# !mkdir otsu_cleaned
for file_name in os.listdir('/content/otsu'):
with open(f'otsu/{file_name}') as file:
with open(f'otsu_cleaned/{file_name}', 'w') as out_file:
for line in file:
line = ' '.join(line.strip().split()[1:-1])
out_file.write(f'{line}\n')
# + id="3v7fOt65ntY1" executionInfo={"status": "aborted", "timestamp": 1628780273797, "user_tz": 240, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}}
# !gsutil -m cp otsu_cleaned/* gs://cytereader/otsu/
# + [markdown] id="7wurB7oMpAPh"
# ## Train
# + id="3iJekoKc7P7h" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1628780299214, "user_tz": 240, "elapsed": 2843, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="c06def8a-9a34-4732-ce8d-b825e6f5ab90"
# !pip install tokenizers
# #!mkdir cellAttention
# + id="qs1jCahOaotj" executionInfo={"status": "ok", "timestamp": 1628780300734, "user_tz": 240, "elapsed": 115, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}}
#from google.colab import files
#uploaded = files.upload()
# + id="zNEiywff0DT_" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1628780304488, "user_tz": 240, "elapsed": 1438, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="2806ffc0-fb8d-459f-fce7-1c801f2a936e"
# !gsutil cp gs://cytereader/preprocessed_cell_corpus_0.txt .
# #!gsutil cp gs://cytereader/preprocessed_cell_corpus_1.txt .
# + id="-9XXbbzWpCRi"
#from tokenizers import BertWordPieceTokenizer
# wb_tokenizer = BertWordPieceTokenizer(clean_text=True,
# strip_accents=True, lowercase=True)
# wb_tokenizer.train(['preprocessed_cell_corpus_0.txt', 'preprocessed_cell_corpus_1.txt'],
# vocab_size=10000, min_frequency=2,
# special_tokens=["[PAD]", "[UNK]", "[CLS]", "[SEP]", "[MASK]"])
# wb_tokenizer.save_model("./cellAttention")
# + id="EsROp3Se1QeA" executionInfo={"status": "ok", "timestamp": 1628780307838, "user_tz": 240, "elapsed": 1147, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}}
from transformers import BertConfig, AutoTokenizer, BertTokenizer, BertLMHeadModel, DistilBertForMaskedLM
from transformers.models.bert.tokenization_bert import CellBertTokenizer
configuration = DistilBertConfig(vocab_size=1000)
model = DistilBertForMaskedLM(configuration)
#tokenizer = BertTokenizer.from_pretrained("./cellAttention", max_len=64)
#tokenizer = CellBertTokenizer.from_pretrained('./cellAttention/',vocab_file="vocab.txt")
# + id="Zo006Ohb7Odq" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1628780310924, "user_tz": 240, "elapsed": 1826, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="8f8b64fb-4ffa-42dd-c619-bc2ec10eb6b1"
# !mkdir cellAttention
# !gsutil cp gs://cytereader/vocab.txt cellAttention/
tokenizer = CellBertTokenizer.from_pretrained('./cellAttention',vocab_file="./cellAttention/vocab.txt")
#tokenizer = CellBertTokenizer.from_pretrained('./cellAttention/',vocab_file="vocab.txt")
#model.num_parameters()
# + colab={"base_uri": "https://localhost:8080/"} id="P7KQwANfmxCn" executionInfo={"status": "ok", "timestamp": 1628780314578, "user_tz": 240, "elapsed": 141, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="171999a3-77d5-4b84-84c3-7720035143b9"
tokenizer.vocab
tokenizer.encode('CD45+ CD196_CCR6+'.lower())
# + id="834hY8Pam2z6"
#cnt = 0
#with open('small.txt','w') as file:
# for line in open("preprocessed_cell_corpus_0.txt"):
# file.write(line.lower())
# cnt += 1
# if cnt == 128:
# break
# + id="9raHGglZm4qj" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1628780437131, "user_tz": 240, "elapsed": 120646, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="a3a8bdf7-44d4-48e9-e048-109d0a6fb719"
# !mkdir otsu
# !gsutil -m cp -r gs://cytereader/otsu/ otsu/
# + id="27tNvSbD1i0Z" colab={"base_uri": "https://localhost:8080/", "height": 156, "referenced_widgets": ["e965a389ff8d441f9ad9d394b7096122", "ddcbf65603b240d59cdfcef04fd1511c", "ebbae0b2e4634f92b6e59fff42072755", "335cd04efa9a49eda05c905f60560d6c", "195c5447bfac4e5fb26fa2898d8b0dbd", "883ff2ec60a042839e581deb44081161", "b0aa3ff564e94a4d83ede532f8f25c4a", "06ec719188524e70ba301cfde5007c71", "43fd1060acdd4e99826760f78a6935b5", "<KEY>", "7432bb2201e44b98b67406003c6ebaca", "b4be1c5481f44aa3a6e1c513e530aecc", "56c7d3fdef6842558a808184e665e289", "2aecd932546942fa822cd578eb854fec", "43d1613df4b44ee4837a901b77900dc9", "<KEY>", "f3da79286df04ae099912bd8db913c68", "<KEY>", "f08e561a0d804656b7896e2126398af3", "650957cdfe6442239e4a43058f5d7de0", "d9bb294224c3402a99fa550e4936b5dc", "1fa460dfbfca4ffaa4e8ebe113bae932"]} executionInfo={"status": "ok", "timestamp": 1628780593627, "user_tz": 240, "elapsed": 154284, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}} outputId="d163d010-af61-4ba7-b44a-e60241694f57"
# %%time
from transformers import LineByLineTextDataset
from datasets import load_dataset
import os
files_name = ['otsu/otsu/' + x for x in os.listdir('/content/otsu/otsu/')][0:2]
dataset = load_dataset("text", data_files=files_name, split='train')
max_length = 128
batch_size=64
def tokenize_function(examples):
tokenizer_ = tokenizer([x.lower() for x in examples['text']], max_length=max_length, truncation=True, padding='max_length')
return tokenizer_
train_data_batch = dataset.map(
tokenize_function,
batched=True,
batch_size=batch_size,
)
# + id="sDvACIyx2l73" executionInfo={"status": "ok", "timestamp": 1628780602858, "user_tz": 240, "elapsed": 132, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}}
from transformers.data.data_collator import DataCollatorForLanguageModeling, DataCollatorForSOP, DataCollatorForNetutralCellModeling
data_collator = DataCollatorForNetutralCellModeling(
tokenizer=tokenizer, ncm=True, mlm_probability=0.15)
# + id="Fn_oz7tu9Q13" executionInfo={"status": "ok", "timestamp": 1628780618214, "user_tz": 240, "elapsed": 12044, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjQwMhgU2MXlA7_i_w6BbygBqnT39n-ka-rltoS=s64", "userId": "16277796292436344578"}}
from transformers import Trainer, TrainingArguments
training_args = TrainingArguments(
output_dir="./cellAttention",
overwrite_output_dir=True,
num_train_epochs=1,
per_device_train_batch_size=32,
save_steps=10_000,
learning_rate=1e-4,
save_total_limit=2,
prediction_loss_only=True,
)
trainer = Trainer(
model=model,
args=training_args,
data_collator=data_collator,
train_dataset=train_data_batch,
)
# + id="-_mKjdOV9UF8" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b04180be-65eb-4ce9-d622-56fa6ebbbd87"
# # %%time
trainer.train()
# + id="YNfsthQO9WYk"
trainer.save_model("./cellAttention")
# + id="KaQacVClEQP8"
from transformers import pipeline
fill_mask = pipeline(
"fill-mask",
model="./cellAttention",
tokenizer="./cellAttention"
)
# + id="b-Cg3tatEZuT"
fill_mask("CD45+ CD196_CCR6+ CD181_CXCR1- HLA_DR- CD15- CD31_PECAM1- CD8a- CD182_CXCR2[MASK] CD66ace- CD63- CD14- CD66b- CD62L_Lselectin- CD3+ CD27- CD86+ CD10- CD197_CCR7+ CD28- CD11c- CD33- CD161- CD45RO- CD24- CD38+ CD278_ICOS- CD32- CD152_CTLA4+ IgM+ CD184_CXCR4+ CD279_PD1- CD56+ CD16-")
# + id="GU-Uqz46FfmS"
input = 'CD45+ CD196_CCR6+ CD181_CXCR1- HLA_DR- CD15-'
masked_input = 'CD45+ CD196_CCR6+ CD181_CXCR1[MASKED] HLA_DR- CD15-'
output = 'CD45+ CD196_CCR6+ CD181_CXCR1-/+ HLA_DR- CD15-'
# + id="F07vRaaZnMEH"
encode = tokenizer.encode('CD45+')
encode
# + id="eV9u_NqNnNe7"
tokenizer.decode(encode)
# + id="Ilxb54vNnPL8"
tokenizer.convert_ids_to_tokens([56, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37,
37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37,
37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37,
37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37,
37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37, 37,
37, 61])
# + id="XsLaINy7nQhB"
tokenizer.convert_tokens_to_ids('CD45+ CD196_CCR6+ CD181_CXCR1- HLA_DR- CD15- CD31_PECAM1- CD8a- CD182_CXCR2- IgA- CD66ace- CD63- CD14- CD66b- CD62L_Lselectin- CD3+ CD27- CD86+ CD10- CD197_CCR7+ CD28- CD11c- CD33- CD161- CD45RO- CD24- CD38+ CD278_ICOS- CD32- CD152_CTLA4+ IgM+ CD184_CXCR4+ CD279_PD1- CD56+ CD16-'.split())
# + id="fdR3nopynSR4"
tokenizer.convert_tokens_to_ids(['CD45+', 'CD196_CCR6+', 'CD181_CXCR1-', 'HLA_DR-', 'CD15-', 'CD31_PECAM1-', 'CD8a-', 'CD182_CXCR2-', 'IgA-', 'CD66ace-', 'CD63-', 'CD14-', 'CD66b-', 'CD62L_Lselectin-', 'CD3+', 'CD27-', 'CD86+', 'CD10-', 'CD197_CCR7+', 'CD28-', 'CD11c-', 'CD33-', 'CD161-', 'CD45RO-', 'CD24-', 'CD38+', 'CD278_ICOS-', 'CD32-', 'CD152_CTLA4+', 'IgM+', 'CD184_CXCR4+', 'CD279_PD1-', 'CD56+', 'CD16-']
)
# + id="jcsu8F95rJTo"
# !gsutil cp -r /content/cellAttention/*.bin gs://cytereader/longphan/test_bert_0/
# !gsutil cp -r /content/cellAttention/config.json gs://cytereader/longphan/test_bert_0/
# !gsutil cp -r /content/cellAttention/vocab.txt gs://cytereader/longphan/test_bert_0/
| training/CellAttention_DistilBERT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 4
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim library
from modsim import *
# -
# ## Returning values
# Here's a simple function that returns a value:
def add_five(x):
return x + 5
# And here's how we call it.
y = add_five(3)
# If you run a function on the last line of a cell, Jupyter displays the result:
add_five(5)
#
# But that can be a bad habit, because usually if you call a function and don't assign the result in a variable, the result gets discarded.
#
# In the following example, Jupyter shows the second result, but the first result just disappears.
add_five(3)
add_five(5)
# When you call a function that returns a variable, it is generally a good idea to assign the result to a variable.
# +
y1 = add_five(3)
y2 = add_five(5)
print(y1, y2)
# -
# **Exercise:** Write a function called `make_state` that creates a `State` object with the state variables `olin=10` and `wellesley=2`, and then returns the new `State` object.
#
# Write a line of code that calls `make_state` and assigns the result to a variable named `init`.
def make_state():
x =State(olin=10, wellesley=2)
return x
init = make_state()
# ## Running simulations
# Here's the code from the previous notebook.
# +
def step(state, p1, p2):
"""Simulate one minute of time.
state: bikeshare State object
p1: probability of an Olin->Wellesley customer arrival
p2: probability of a Wellesley->Olin customer arrival
"""
if flip(p1):
bike_to_wellesley(state)
if flip(p2):
bike_to_olin(state)
def bike_to_wellesley(state):
"""Move one bike from Olin to Wellesley.
state: bikeshare State object
"""
if state.olin == 0:
state.olin_empty += 1
return
state.olin -= 1
state.wellesley += 1
def bike_to_olin(state):
"""Move one bike from Wellesley to Olin.
state: bikeshare State object
"""
if state.wellesley == 0:
state.wellesley_empty += 1
return
state.wellesley -= 1
state.olin += 1
def decorate_bikeshare():
"""Add a title and label the axes."""
decorate(title='Olin-Wellesley Bikeshare',
xlabel='Time step (min)',
ylabel='Number of bikes')
# -
# Here's a modified version of `run_simulation` that creates a `State` object, runs the simulation, and returns the `State` object.
def run_simulation(p1, p2, num_steps):
"""Simulate the given number of time steps.
p1: probability of an Olin->Wellesley customer arrival
p2: probability of a Wellesley->Olin customer arrival
num_steps: number of time steps
"""
state = State(olin=10, wellesley=2,
olin_empty=0, wellesley_empty=0)
for i in range(num_steps):
step(state, p1, p2)
return state
# Now `run_simulation` doesn't plot anything:
state = run_simulation(0.4, 0.2, 60)
# But after the simulation, we can read the metrics from the `State` object.
state.olin_empty
# Now we can run simulations with different values for the parameters. When `p1` is small, we probably don't run out of bikes at Olin.
state = run_simulation(0.2, 0.2, 60)
state.olin_empty
# When `p1` is large, we probably do.
state = run_simulation(0.6, 0.2, 60)
state.olin_empty
# ## More for loops
# `linspace` creates a NumPy array of equally spaced numbers.
p1_array = linspace(0, 1, 5)
# We can use an array in a `for` loop, like this:
for p1 in p1_array:
print(p1)
# This will come in handy in the next section.
#
# `linspace` is defined in `modsim.py`. You can get the documentation using `help`.
help(linspace)
# `linspace` is based on a NumPy function with the same name. [Click here](https://docs.scipy.org/doc/numpy/reference/generated/numpy.linspace.html) to read more about how to use it.
# **Exercise:**
# Use `linspace` to make an array of 10 equally spaced numbers from 1 to 10 (including both).
array = linspace (1,10,10)
# **Exercise:** The `modsim` library provides a related function called `linrange`. You can view the documentation by running the following cell:
help(linrange)
# Use `linrange` to make an array of numbers from 1 to 11 with a step size of 2.
array = linrange (1,11,2)
# ## Sweeping parameters
# `p1_array` contains a range of values for `p1`.
p2 = 0.2
num_steps = 60
p1_array = linspace(0, 1, 11)
# The following loop runs a simulation for each value of `p1` in `p1_array`; after each simulation, it prints the number of unhappy customers at the Olin station:
for p1 in p1_array:
state = run_simulation(p1, p2, num_steps)
print(p1, state.olin_empty)
# Now we can do the same thing, but storing the results in a `SweepSeries` instead of printing them.
#
#
# +
sweep = SweepSeries()
for p1 in p1_array:
state = run_simulation(p1, p2, num_steps)
sweep[p1] = state.olin_empty
# -
# And then we can plot the results.
# +
plot(sweep, label='Olin')
decorate(title='Olin-<NAME>',
xlabel='Arrival rate at Olin (p1 in customers/min)',
ylabel='Number of unhappy customers')
savefig('figs/chap02-fig02.pdf')
# -
# ## Exercises
#
# **Exercise:** Wrap this code in a function named `sweep_p1` that takes an array called `p1_array` as a parameter. It should create a new `SweepSeries`, run a simulation for each value of `p1` in `p1_array`, store the results in the `SweepSeries`, and return the `SweepSeries`.
#
# Use your function to plot the number of unhappy customers at Olin as a function of `p1`. Label the axes.
def sweep_p1(p1_array):
sweep = SweepSeries()
for p1 in p1_array:
state = run_simulation(p1, p2, num_steps)
sweep[p1] = state.olin_empty
return sweep
plot(sweep, 'bo', label= 'Olin')
decorate(title='<NAME>',
xlabel='Arrival rate at Olin (p1 in customers/min)',
ylabel='Number of unhappy customers')
# **Exercise:** Write a function called `sweep_p2` that runs simulations with `p1=0.5` and a range of values for `p2`. It should store the results in a `SweepSeries` and return the `SweepSeries`.
#
# +
p1 = 0.5
num_steps = 60
p2_array = linspace(0, 1, 11)
def sweep_p2 (p2_array):
sweep = SweepSeries ()
for p2 in p2_array:
state = run_simulation(p1, p2, num_steps)
sweep[p2] = state.olin_empty
return sweep
sweep_p2(p2_array)
# +
plot(sweep, 'bo', label='Olin')
decorate (title='<NAME>',
xlabel='Arrival Rate at Wellesley (p2 in Customers per Minute)',
ylabel='Number of Unhappy Customers')
# -
# ## Optional exercises
#
# The following two exercises are a little more challenging. If you are comfortable with what you have learned so far, you should give them a try. If you feel like you have your hands full, you might want to skip them for now.
#
# **Exercise:** Because our simulations are random, the results vary from one run to another, and the results of a parameter sweep tend to be noisy. We can get a clearer picture of the relationship between a parameter and a metric by running multiple simulations with the same parameter and taking the average of the results.
#
# Write a function called `run_multiple_simulations` that takes as parameters `p1`, `p2`, `num_steps`, and `num_runs`.
#
# `num_runs` specifies how many times it should call `run_simulation`.
#
# After each run, it should store the total number of unhappy customers (at Olin or Wellesley) in a `TimeSeries`. At the end, it should return the `TimeSeries`.
#
# Test your function with parameters
#
# ```
# p1 = 0.3
# p2 = 0.3
# num_steps = 60
# num_runs = 10
# ```
#
# Display the resulting `TimeSeries` and use the `mean` function provided by the `TimeSeries` object to compute the average number of unhappy customers.
# +
# Solution goes here
# +
# Solution goes here
# -
# **Exercise:** Continuting the previous exercise, use `run_multiple_simulations` to run simulations with a range of values for `p1` and
#
# ```
# p2 = 0.3
# num_steps = 60
# num_runs = 20
# ```
#
# Store the results in a `SweepSeries`, then plot the average number of unhappy customers as a function of `p1`. Label the axes.
#
# What value of `p1` minimizes the average number of unhappy customers?
# +
# Solution goes here
# +
# Solution goes here
| code/chap04mine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Handling Volume with Apache Spark
# Use Apache Spark to perform word count on product names after tokenization.
# ## License
# MIT License
#
# Copyright (c) 2018 PT Bukal<EMAIL>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all
# copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
# SOFTWARE.
# ## Software Versions
import sys, os
print("Python %s" % sys.version)
import time
import pyspark
print("PySpark %s" % pyspark.__version__)
from pyspark.sql import SparkSession
import platform
print("platform %s" % platform.__version__)
print("OS", platform.platform())
import tensorflow as tf
print("TensorFlow %s" % tf.__version__)
from tensorflow.keras.preprocessing.text import text_to_word_sequence
# + language="bash"
# /usr/local/spark/bin/spark-submit --version
# -
# ## Perform Word Count using Notebook (NB)
# Setup spark.
APP_NAME = "bukalapak-core-ai.big-data-3v.volume-spark"
spark = SparkSession \
.builder \
.appName(APP_NAME) \
.getOrCreate()
sc = spark.sparkContext
sc
# Input and output URLs.
product_names_text_filename = \
"file:/home/jovyan/work/" + \
"data/product_names_sample/" + \
"product_names.rdd"
product_names_text_filename
product_names_word_count_nb_orc_filename = \
"file:/home/jovyan/work/" + \
"data/product_names_sample/" + \
"product_names_word_count_nb.orc"
product_names_word_count_nb_orc_filename
# Read input file.
product_names_df = spark.read.text(product_names_text_filename)
product_names_df
product_names_df.head(10)
product_names_rdd = product_names_df.rdd
product_names_rdd
product_names_rdd.getNumPartitions()
product_names_rdd.top(10)
# Perform tokenization.
def tokenize(words):
return text_to_word_sequence(words['value'])
tokenized_product_names_rdd = \
product_names_rdd.flatMap(lambda product_name: tokenize(product_name))
tokenized_product_names_rdd
tokenized_product_names_rdd.top(20)
# Perform word count.
word_count_product_names_rdd = \
tokenized_product_names_rdd.map(lambda word: (word, 1)) \
.reduceByKey(lambda a, b: a + b)
word_count_product_names_rdd
word_count_product_names_rdd.top(20)
# Save the output. __Note:__ Don't forget to delete existing `product_names_word_count_nb.orc` directory in `data/product_names_sample`. Following Spark implementation does not overwrite existing data but it will throw error.
word_count_product_names_df = spark.createDataFrame(word_count_product_names_rdd)
word_count_product_names_df.write.save(product_names_word_count_nb_orc_filename, \
format="orc")
# Read back the word count.
new_word_count_product_names_df = spark.read.orc(product_names_word_count_nb_orc_filename)
new_word_count_product_names_df
new_word_count_product_names_df.head(20)
# Stop Spark.
sc.stop()
spark.stop()
# ## Perform Word Count using Spark Submit (SS)
# +
# %%writefile bukalapak-core-ai.big-data-3v.volume-spark.py
# Copyright (c) 2018 PT Bukalapak.com
#
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all
# copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
# SOFTWARE.
from pyspark.sql import SparkSession
APP_NAME = "bukalapak-core-ai.big-data-3v.volume-spark"
def tokenize(words):
from tensorflow.keras.preprocessing.text import text_to_word_sequence
return text_to_word_sequence(words['value'])
def main(spark):
# Input
product_names_text_filename = \
"file:/home/jovyan/work/" + \
"data/product_names_sample/" + \
"product_names.rdd"
# Output
product_names_word_count_ss_orc_filename = \
"file:/home/jovyan/work/" + \
"data/product_names_sample/" + \
"product_names_word_count_ss.orc"
# Read input
product_names_df = spark.read.text(product_names_text_filename)
product_names_rdd = product_names_df.rdd
# Perform tokenization and word count
tokenized_product_names_rdd = \
product_names_rdd.flatMap(lambda product_name: tokenize(product_name))
word_count_product_names_rdd = \
tokenized_product_names_rdd.map(lambda word: (word, 1)) \
.reduceByKey(lambda a, b: a + b)
# Write output
word_count_product_names_df = spark.createDataFrame(word_count_product_names_rdd)
word_count_product_names_df.write.save(product_names_word_count_ss_orc_filename, \
format="orc")
if __name__ == "__main__":
# Configure Spark
spark = SparkSession \
.builder \
.appName(APP_NAME) \
.getOrCreate()
main(spark)
spark.stop()
# -
# __Note:__ Don't forget to delete existing `product_names_word_count_ss.orc` directory in `data/product_names_sample`. Following Spark implementation does not overwrite existing data but it will throw error.
# + language="bash"
# /usr/local/spark/bin/spark-submit \
# --executor-memory 1g --executor-cores 1 --num-executors 2 \
# bukalapak-core-ai.big-data-3v.volume-spark.py
# -
# Read back the word count.
APP_NAME = "bukalapak-core-ai.big-data-3v.volume-spark"
spark = SparkSession \
.builder \
.appName(APP_NAME) \
.getOrCreate()
product_names_word_count_ss_orc_filename = \
"file:/home/jovyan/work/" + \
"data/product_names_sample/" + \
"product_names_word_count_ss.orc"
product_names_word_count_ss_orc_filename
new_word_count_product_names_df = spark.read.orc(product_names_word_count_ss_orc_filename)
new_word_count_product_names_df
new_word_count_product_names_df.head(20)
# Stop Spark.
sc.stop()
spark.stop()
# ## Software Versions
# + language="bash"
# cat /etc/os-release
# + language="bash"
# pip freeze
# + language="bash"
# conda list
| volume/notebook/spark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import ticker
import pycountry_convert as pc
import folium
from datetime import datetime,date
from scipy.interpolate import make_interp_spline, BSpline
# %matplotlib inline
import os
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
import cufflinks as cf
# -
def Reverse(lst):
return [ele for ele in reversed(lst)]
path='data/COVID19/JHU/COVID-19-master/csse_covid_19_data/csse_covid_19_time_series'
confirmed_fn=os.path.join(path,'time_series_covid19_confirmed_global.csv')
deaths_fn=os.path.join(path,'time_series_covid19_deaths_global.csv')
recovered_fn=os.path.join(path,'time_series_covid19_recovered_global.csv')
confirmed_fn
df_confirmed = pd.read_csv(confirmed_fn)
df_deaths = pd.read_csv(deaths_fn)
df_recovered = pd.read_csv(recovered_fn)
df_confirmed.head()
df_confirmed = df_confirmed.rename(columns={"Province/State":"state","Country/Region": "country"})
df_deaths = df_deaths.rename(columns={"Province/State":"state","Country/Region": "country"})
df_recovered = df_recovered.rename(columns={"Province/State":"state","Country/Region": "country"})
# +
# Changing the conuntry names as required by pycountry_convert Lib
df_confirmed.loc[df_confirmed['country'] == "US", "country"] = "USA"
df_deaths.loc[df_deaths['country'] == "US", "country"] = "USA"
df_recovered.loc[df_recovered['country'] == "US", "country"] = "USA"
df_confirmed.loc[df_confirmed['country'] == 'Korea, South', "country"] = 'South Korea'
df_deaths.loc[df_deaths['country'] == 'Korea, South', "country"] = 'South Korea'
df_recovered.loc[df_recovered['country'] == 'Korea, South', "country"] = 'South Korea'
df_confirmed.loc[df_confirmed['country'] == 'Taiwan*', "country"] = 'Taiwan'
df_deaths.loc[df_deaths['country'] == 'Taiwan*', "country"] = 'Taiwan'
df_recovered.loc[df_recovered['country'] == 'Taiwan*', "country"] = 'Taiwan'
df_confirmed.loc[df_confirmed['country'] == 'Congo (Kinshasa)', "country"] = 'Democratic Republic of the Congo'
df_deaths.loc[df_deaths['country'] == 'Congo (Kinshasa)', "country"] = 'Democratic Republic of the Congo'
df_recovered.loc[df_recovered['country'] == 'Congo (Kinshasa)', "country"] = 'Democratic Republic of the Congo'
df_confirmed.loc[df_confirmed['country'] == "Cote d'Ivoire", "country"] = "Côte d'Ivoire"
df_deaths.loc[df_deaths['country'] == "Cote d'Ivoire", "country"] = "Côte d'Ivoire"
df_recovered.loc[df_recovered['country'] == "Cote d'Ivoire", "country"] = "Côte d'Ivoire"
df_confirmed.loc[df_confirmed['country'] == "Reunion", "country"] = "Réunion"
df_deaths.loc[df_deaths['country'] == "Reunion", "country"] = "Réunion"
df_recovered.loc[df_recovered['country'] == "Reunion", "country"] = "Réunion"
df_confirmed.loc[df_confirmed['country'] == 'Congo (Brazzaville)', "country"] = 'Republic of the Congo'
df_deaths.loc[df_deaths['country'] == 'Congo (Brazzaville)', "country"] = 'Republic of the Congo'
df_recovered.loc[df_recovered['country'] == 'Congo (Brazzaville)', "country"] = 'Republic of the Congo'
df_confirmed.loc[df_confirmed['country'] == 'Bahamas, The', "country"] = 'Bahamas'
df_deaths.loc[df_deaths['country'] == 'Bahamas, The', "country"] = 'Bahamas'
df_recovered.loc[df_recovered['country'] == 'Bahamas, The', "country"] = 'Bahamas'
df_confirmed.loc[df_confirmed['country'] == 'Gambia, The', "country"] = 'Gambia'
df_deaths.loc[df_deaths['country'] == 'Gambia, The', "country"] = 'Gambia'
df_recovered.loc[df_recovered['country'] == 'Gambia, The', "country"] = 'Gambia'
# getting all countries
countries = np.asarray(df_confirmed["country"])
# -
df_active = df_confirmed.copy()
df_active.iloc[:,5:] = df_active.iloc[:,5:] - df_recovered.iloc[:,5:] - df_deaths.iloc[:,5:]
df_active.head(5)
df_confirmed = df_confirmed.replace(np.nan, '', regex=True)
df_deaths = df_deaths.replace(np.nan, '', regex=True)
df_recovered = df_recovered.replace(np.nan, '', regex=True)
df_active = df_active.replace(np.nan, '', regex=True)
# +
def get_total_cases(cases, country = "All"):
if(country == "All") :
return np.sum(np.asarray(cases.iloc[:,5:]),axis = 0)[-1]
else :
return np.sum(np.asarray(cases[cases["country"] == country].iloc[:,5:]),axis = 0)[-1]
def get_mortality_rate(confirmed,deaths, continent = None, country = None):
if continent != None:
params = ["continent",continent]
elif country != None:
params = ["country",country]
else :
params = ["All", "All"]
if params[1] == "All" :
Total_confirmed = np.sum(np.asarray(confirmed.iloc[:,5:]),axis = 0)
Total_deaths = np.sum(np.asarray(deaths.iloc[:,5:]),axis = 0)
mortality_rate = np.round((Total_deaths/Total_confirmed)*100,2)
else :
Total_confirmed = np.sum(np.asarray(confirmed[confirmed[params[0]] == params[1]].iloc[:,5:]),axis = 0)
Total_deaths = np.sum(np.asarray(deaths[deaths[params[0]] == params[1]].iloc[:,5:]),axis = 0)
mortality_rate = np.round((Total_deaths/Total_confirmed)*100,2)
return np.nan_to_num(mortality_rate)
def dd(date1,date2):
return (datetime.strptime(date1,'%m/%d/%y') - datetime.strptime(date2,'%m/%d/%y')).days
# +
confirmed_cases = df_confirmed.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).iloc[:,-1]
recovered_cases = df_recovered.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).iloc[:,-1]
deaths = df_deaths.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).iloc[:,-1]
active_cases = df_active.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).iloc[:,-1]
confirmed_cases.name = "Confirmed Cases"
recovered_cases.name = "Recovered Cases"
deaths.name = "Deaths Reported"
active_cases.name = "Active Cases"
df_countries_cases = pd.DataFrame([confirmed_cases,recovered_cases,deaths,active_cases]).transpose()
# -
df_countries_cases.head()
# +
f = plt.figure(figsize=(10,5))
f.add_subplot(111)
plt.barh(df_countries_cases.sort_values('Confirmed Cases')["Confirmed Cases"].index[-10:],df_countries_cases.sort_values('Confirmed Cases')["Confirmed Cases"].values[-10:],)
plt.tick_params(size=5,labelsize = 13)
plt.xlabel("Confirmed Cases",fontsize=18)
plt.title("Top 10 Countries (Confirmed Cases)",fontsize=20)
# plt.savefig('Top 10 Countries (Confirmed Cases).png')
plt.grid(alpha=0.3)
# +
f = plt.figure(figsize=(10,5))
f.add_subplot(111)
plt.barh(df_countries_cases.sort_values('Deaths Reported')["Deaths Reported"].index[-10:],df_countries_cases.sort_values('Deaths Reported')["Deaths Reported"].values[-10:])
plt.tick_params(size=5,labelsize = 13)
plt.xlabel("Confirmed Cases",fontsize=18)
plt.title("Top 10 Countries (Deaths Cases)",fontsize=20)
# plt.savefig('Top 10 Countries (Deaths Cases).png')
plt.grid(alpha=0.3)
# -
Top10_Deaths=list(df_countries_cases.sort_values('Deaths Reported')["Deaths Reported"].index[-10:])
Top10_Confirmed=list(df_countries_cases.sort_values('Confirmed Cases')["Confirmed Cases"].index[-10:])
Top10_Recovered=list(df_countries_cases.sort_values('Recovered Cases')["Recovered Cases"].index[-10:])
Top10_Active=list(df_countries_cases.sort_values('Active Cases')["Active Cases"].index[-10:])
Top10_Confirmed=Reverse(list(df_countries_cases.sort_values('Confirmed Cases')["Confirmed Cases"].index[-10:]))
confirmed_series = df_confirmed.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).loc[Top10_Confirmed].T
recovered_series = df_recovered.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).loc[Top10_Confirmed].T
deaths_series = df_deaths.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).loc[Top10_Recovered].T
active_series = df_active.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).loc[Top10_Active].T
#https://plotly.com/~ziwang/69/
@interact
def top10_confirmed_country():
confirmed_series.iplot(xTitle='Date', yTitle='Convirmed Cases',title='Top 10 country: Confirmed COVID-19 Cases',theme='solar')
# +
Top21_Confirmed=Reverse(list(df_countries_cases.sort_values('Confirmed Cases')["Confirmed Cases"].index[-51:]))
print(Top21_Confirmed)
# -
confirmed_series_21 = df_confirmed.groupby(["country"]).sum().drop(['Lat','Long'],axis =1).loc[Top21_Confirmed].T
#https://plotly.com/~ziwang/136/
@interact
def plot_top20country_confirmed_cases():
confirmed_series_21.iplot(subplots=True, shape=(17,3), shared_xaxes=True, fill=False,
subplot_titles=True,legend=False, xTitle="",yTitle="",
title='Top 51 countries: Confirmed COVID-19 Cases',theme='pearl')
| codes/A01_globalCasesAnalysis_run01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: csc790
# language: python
# name: csc790
# ---
# +
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import keras
from tensorflow.random import set_seed
from tqdm import tqdm
from contextlib import redirect_stdout
import os
import utils
# -
# Define a model name for repeated use
modelName = "SimpleResNet"
n_folds = 10 # 1 = normal experiment
batch_size = 128 # 128 is default across the models
#Seeding random state to 13 always, for reproducibility
np.random.seed(utils.seed)
set_seed(utils.seed)
# +
### Used to select GPU 0=first device, 1=second device, etc...
os.environ["CUDA_VISIBLE_DEVICES"]="0"
gpus = tf.config.experimental.list_physical_devices('GPU')
print('gpus:',gpus)
if gpus:
try:
tf.config.experimental.set_visible_devices(gpus[0], 'GPU')
tf.config.experimental.set_memory_growth(gpus[0], True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPU,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Virtual devices must be set before GPUs have been initialized
print(e)
# -
dataset = utils.makeDataset(pathFromCwdToDataRoot="Data")
kFCV_sets, weights_dict = utils.makeFolds(dataset=dataset, n_folds=n_folds, batch_size=batch_size)
print(weights_dict)
# +
k_results=pd.DataFrame(columns = ['Fold', 'Loss', 'Accuracy'])
for i, k in enumerate(kFCV_sets):
print("Fold", i+1, "of", len(kFCV_sets))
train_generator, test_generator, val_generator = k
#Build Model
model = utils.makeModel(inputShape=(150, 150, 3), modelName=modelName)
opt = tf.optimizers.Adam()
model.compile(
optimizer = opt,
loss = keras.losses.BinaryCrossentropy(from_logits = True),
metrics = ["accuracy"]
)
#Fit data
model.fit(
train_generator,
validation_data=val_generator,
callbacks = utils.callbacks,
epochs = 100,
class_weight = weights_dict,
max_queue_size = 10,
workers = os.cpu_count(),
)
#Test accuracy
results = model.evaluate(
test_generator,
max_queue_size = 10,
workers = os.cpu_count(),
)
k_results = k_results.append({'Fold':i+1, 'Loss':results[0], 'Accuracy':results[1]}, ignore_index=True)
if i == 0:
# Write the summary to a file
with open(f'Results/{modelName}Summary.txt', 'w') as f:
with redirect_stdout(f):
print(model.summary())
#Save the model
model.save(f'Models/{modelName}')
keras.backend.clear_session()
k_results = k_results.append({'Fold':"Avg", 'Loss':np.average(k_results['Loss']), 'Accuracy':np.average(k_results['Accuracy'])}, ignore_index=True)
# -
if n_folds != 1:
k_results.to_csv(f'Results/k-fcv_{modelName}.csv')
| SimpleResNet_EXT_k-fold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Справка
#
# API Cross Web осуществляет расчет данных проекта Cross Web (измерение аудитории в интернете).
#
# Обращаться к API Cross Web будем с помощью Jupyter Notebook, для этого необходимо владеть некоторыми терминами.
#
# Ниже описаны основные из них.
#
# ### usetype - тип пользования интернетом
#
# Возможные варианты:
#
# - 1 - Web Desktop
# - 2 - Web Mobile
# - 3 - App Mobile
#
# ## Типы расчетов
#
#
# ### Audience - расчет объема аудитории интернет-проектов
#
# На текущий момент доступны следующие статистики:
#
# - adr - Average Daily Reach
# - adrPer - Average Daily Reach %
# - affinity
# - affinityInternet
# - affinityAdr
# - affinityAdrInternet
# - averageAge - Average Age
# - dr - Days Reached
# - drfd - Days Reached Frequency Distribution
# - frequency
# - ots
# - reach
# - reachN - Reach N+ Distribution
# - reachPer - Reach %
# - uni - Universe
# - att - Attention
# - adatt - Average Daily Attention
# - addPerU - Average Daily Duration на пользователей
# - addPerP - Average Daily Duration на население
# # Библиотека Mediascope API
# Библиотека Mediascope содержит набор классов и методов, которые позволяют упростить работу с API.
#
# Импортируйте библиотеку Mediascope API
#
#
# +
# %reload_ext autoreload
# %autoreload 2
import sys
import os
import re
import json
import datetime
import time
import pandas as pd
from pathlib import Path
from IPython.display import JSON
from mediascope_api.core import net as msnet
from mediascope_api.crossweb import catalogs as cwc
from mediascope_api.crossweb import tasks as cwt
# -
# Создайте объекты для работы с API
mtask = cwt.CrossWebTask()
# # Работа с Mediascope Jupyter Notebook
#
# В работе с Mediascope Jupyter Notebook можно выделить два основных действия:
#
# - расчет заданий (создание задания, расчет, получение результата);
# - работа со справочниками.
# ## Расчет заданий
#
# Стандартный сценарий работы с заданиями:
#
# - инициализация - импорт библиотеки и создание объектов для работы с API Cross Web;
# - формирование задания;
# - отправление задания на расчет и ожидание результата;
# - получение результата и его преобразование в pandas.DataFrame;
# - при необходимости сохранение результата в Excel.
#
# При формировании задания можно указать параметры, приведенные ниже.
#
# - **filters** - фильтры нескольких типов:
# - **date_filter** - фильтр по периодам
# - **usetype_filter** - фильтр по типам пользования интернетом
# - **geo_filter** - фильтр по географии
# - **demo_filter** - фильтр по соц.-дему
# - **mart_filter** - фильтр по медиа-объектам
# - **slices** - срезы
# - **statistics** - статистики
# - **scales** - шкалы, если заданы статистики со шкалами (drfd, reachN)
#
# Из перечисленных параметров обязательными являются:
#
# - **date_filter** - фильтр по периоду
# - **statistics** - статистики
# - **scales** - шкалы, если заданы статистики со шкалами (drfd, reachN)
#
# Без их указания рассчитать задание невозможно.
#
# Для расчета заданий существуют методы в модуле mediascope_api.crossweb.tasks:
#
# - **build_audience_task** - формирует задание: принимает указанные параметры, проверяет их и создает JSON для API Cross Web;
# - **send_audience_task** - отправляет задание на расчет;
# - **wait_task** - ожидает расчет;
# - **get_result** - получает результат;
# - **result2table** - преобразует результат из JSON в pandas.DataFrame.
#
# **При формировании задания необходимо корректно задавать атрибуты для параметров.**
#
# Для получения списка допустимых атрибутов воспользуйтесь методом `get_media_unit()` модуля `mediascope_api.crossweb.catalogs`.
#
# Создавать выражения для фильтров **geo_filter**, **demo_filter** и **mart_filter** можно с помощью следующих операторов:
#
# = (равно)
# != (не равно)
# > (больше)
# < (меньше)
# >= (больше или равно)
# <= (меньше или равно)
# AND (И)
# OR (ИЛИ)
# IN() (вхождение в список значений)
# NIN() (невхождение в список значений)
# ## Вывод справки
#
# Все методы библиотеки Mediascope имеют справку.
#
# Для вывода справки нужно нажать комбинацию клавиш __Shift+Tab__
# ```
# mtask.build_task(
# ```
# или воспользоваться командой __help__
help(mtask.build_task)
# ## Работа со справочниками
#
# Для работы со справочниками в библиотеке Mediascope содержатся методы в модуле `mediascope_api.crossweb.catalogs`.
#
# В ноутбуке [catalogs](catalogs.ipynb) приведены примеры получения данных из всех доступных справочников.
| crossweb/help.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# name: ir
# ---
# + id="IlOR1i1fc7SJ"
variable<- 'happy'
install.packages('ghql')
library(ghql)
library(tidyverse)
con <- GraphqlClient$new(
url = "https://flowing-crow-42.hasura.app/v1/graphql")
qry <- Query$new()
qry$query('mydata', paste0('{
gss_gss_data {
Education_gap
year
',variable, '_num
wtssall
}
}'))
data<- qry$queries$mydata
# returns json
x <- con$exec(qry$queries$mydata)
data<- jsonlite::fromJSON(x)
d<- data$data[[1]]
timetrend<- d %>% group_by(year, Education_gap) %>%
summarise(avg = weighted.mean(!!sym(paste0(variable, '_num')), w = wtssall, na.rm=TRUE))
timetrend
| colab/notebooks/happy-timetrend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# %load_ext rpy2.ipython
# +
import os
import numpy as np
import pandas as pd
import Scorer
# -
# Global parameters
BASE = os.path.realpath('../')
TEST_SET = os.path.join(BASE, 'data', 'test_data','input_data', 'TestSet8')
# # Test Set
# +
abundance_file = os.path.join(TEST_SET, "TestSet8_abundance_thresholds.txt")
counts_file = os.path.join(TEST_SET, "TestSet8_timepoint_counts.txt")
times = np.array([21,28])
# initialize screen object
screen = Scorer.Scorer(counts_file, times, abundance_file)
# set some parameters
# number of iterations to perform
screen.niter = 2
screen.testing = True
screen.null_target_id = '0'
screen.verbose = True
# perform construct fitness calculation
screen.run_construct_fitting()
# run bootstraps
screen.run_sampling()
# summarize results
screen.summarize()
# view results
screen.results.head()
# -
# ### Compare to CTG benchmark pi scores
# +
benchmark = os.path.join(BASE, 'data', 'test_data', 'output_data', 'Notebook8Test_pi.txt')
benchmark = pd.read_csv(benchmark, sep="\t", header=0)
benchmark_pi_mean = benchmark[['geneA','geneB','pi']]
benchmark_pi_mean = benchmark_pi_mean.sort_values(['geneA','geneB']).reset_index(drop=True)
#pi_scores_mean = screen.results.mean(axis=1).reset_index()
pi_scores_mean = screen.results[['geneA','geneB','pi_mean']]
comb = pd.merge(pi_scores_mean, benchmark_pi_mean, on=['geneA','geneB'])
print('All pi-scores close:')
np.allclose(comb['pi_mean'],comb['pi'], atol=1e-6, equal_nan=True)
# + magic_args="-i comb" language="R"
# jointPlot <- function(df, alpha=0.1, title=NULL, correlation=FALSE){
#
# # make a joint plot from the data frame of two columns first one is on x axis
# require(ggplot2)
# require(cowplot)
# # get info
# x_l = colnames(df)[1]
# y_l = colnames(df)[2]
# coord_min = min(na.omit(df))
# coord_max = max(na.omit(df))
# # make scatter plot
# scatter = ggplot(df, aes_string(x=x_l,y=y_l))+ # init grob
# geom_point(shape=1, alpha=alpha, size=4)+ # add points
# geom_abline(slope=1,intercept=0,color="red",linetype="dotted")+ # add perfect 1:1 line
# theme_bw(base_size=18)+ # use preset theme
# xlim(coord_min,coord_max)+ylim(coord_min,coord_max)+ # make square plot
# labs(x=x_l,y=y_l) # add axis labels
# # add correlation info to scatter plot
# if (correlation){
# # get correaltion
# res = cor.test(df[,x_l], df[,y_l])
# p_value = res$p.value
# corr = res$estimate
# # add annotation
# scatter = scatter + annotate("text",
# hjust=0,
# label=paste("Corr: ", format(round(corr, 4), nsmall=3),
# sep=""),
# x=coord_min,y=coord_max)
# scatter = scatter + geom_smooth(se=FALSE, method="lm")
# }
#
# # add title to scatter plot
# if (!is.null(title)){
# scatter = scatter + labs(title=title)
# }
#
# xdens = axis_canvas(scatter, axis="x")+
# geom_histogram(data=df, aes_string(x=x_l), color="black",fill="white",bins=50)
#
# ydens = axis_canvas(scatter, axis="y", coord_flip=TRUE)+
# geom_histogram(data=df, aes_string(x=y_l), color="black",fill="white",bins=50)+
# coord_flip()
#
# p1 <- insert_xaxis_grob(scatter, xdens, grid::unit(.2,'null'),position="top")
# p2 <- insert_yaxis_grob(p1, ydens, grid::unit(.2,'null'),position="right")
# ggdraw(p2)
#
# }
# df = comb[,c('pi_mean','pi')]
# colnames(df) = c('CrAPPY', 'CTG')
# jointPlot(df, title="Test Set 8 Pi Scores", correlation=TRUE)
#
# -
# # Testing abundance thresholds
# +
abundance_file = os.path.join(TEST_SET, "TestSet8_abundance_thresholds.txt")
counts_file = os.path.join(TEST_SET, "TestSet8_timepoint_counts.txt")
times = np.array([21,28])
# initialize screen object
screen = Scorer.Scorer(counts_file, times, min_counts_threshold=8,verbose=True)
# set some parameters
# number of iterations to perform
screen.niter = 2
screen.testing = True
screen.null_target_id = '0'
screen.verbose = True
# perform construct fitness calculation
screen.run_construct_fitting()
# run bootstraps
screen.run_sampling()
# summarize results
screen.summarize()
# view results
screen.results.head()
# +
benchmark = os.path.join(BASE, 'data', 'test_data', 'output_data', 'Notebook8Test_pi.txt')
benchmark = pd.read_csv(benchmark, sep="\t", header=0)
benchmark_pi_mean = benchmark[['geneA','geneB','pi']]
benchmark_pi_mean = benchmark_pi_mean.sort_values(['geneA','geneB']).reset_index(drop=True)
#pi_scores_mean = screen.results.mean(axis=1).reset_index()
pi_scores_mean = screen.results[['geneA','geneB','pi_mean']]
comb = pd.merge(pi_scores_mean, benchmark_pi_mean, on=['geneA','geneB'])
print('All pi-scores close:')
np.allclose(comb['pi_mean'],comb['pi'], atol=1e-6, equal_nan=True)
# -
def rmse(x,y):
return np.power(np.sum(np.power(x - y,2))/len(x), 0.5)
rmse(comb['pi_mean'], comb['pi'])
np.corrcoef(comb[['pi_mean', 'pi']].T)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.scatter(comb['pi_mean'], comb['pi'], s=5)
# # A549
# +
abundance_file = "/cellar/users/bpmunson/crappy/data/test_data/input_data/A549/A549_abundance_thresholds.txt"
counts_file = "/cellar/users/bpmunson/crappy/data/test_data/input_data/A549/A549_timepoint_counts.txt"
times = np.array([3,14,21,28])
# initialize screen object
screen = Scorer.Scorer(counts_file, times)
# set some parameters
# number of iterations to perform
screen.niter = 2
screen.null_target_id = '0'
screen.verbose = True
# perform construct fitness calculation
screen.run_construct_fitting()
# run bootstraps
screen.run_sampling()
# summarize results
screen.summarize()
# view results
screen.results.head()
# +
abundance_file = "/cellar/users/bpmunson/crappy/data/test_data/input_data/A549/A549_abundance_thresholds.txt"
counts_file = "/cellar/users/bpmunson/crappy/data/test_data/input_data/A549/A549_timepoint_counts.txt"
times = np.array([3,14,21,28])
# initialize screen object
screen = Scorer.Scorer(counts_file, times, abundance_file)
# set some parameters
# number of iterations to perform
screen.niter = 2
screen.null_target_id = '0'
screen.verbose = True
# perform construct fitness calculation
screen.run_construct_fitting()
# run bootstraps
screen.run_sampling()
# summarize results
screen.summarize()
# view results
screen.results.head()
# +
sampleName
testing_T3_1 3.025
testing_T3_2 3.525
testing_T14_1 5.525
testing_T14_2 3.025
testing_T21_1 4.525
testing_T21_2 3.025
testing_T28_1 3.025
testing_T28_2 3.525
sampleName
testing_T3_1 3.335947
testing_T3_2 3.475000
testing_T14_1 3.338280
testing_T14_2 4.089135
testing_T21_1 4.525000
testing_T21_2 4.325000
testing_T28_1 4.225000
testing_T28_2 3.575000
# -
2**3.025
| examples/Example_Execution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # multiple Linear Regression (mLR) with scikit-learn (Example for lesson ML05)
#
# Powered by: Dr. <NAME>, DHBW Stuttgart(Germany); July 2020
#
# Following ideas from:
# "Linear Regression in Python" by <NAME>, 28.4.2020
# (see details: https://realpython.com/linear-regression-in-python/#what-is-regression)
#
# You can obtain the properties of the model the same way as in the case of simple linear regression:
# The example is from Lecture: "ML_Concept&Algorithm" (WS2020); Homework 5.4 part c:"mLR (k=2) manual calculations of Adj.R²
# & Jupyter Notebook (Python) to check results"
#
# So let’s start with the next level of Linear Regression, which is multipe Linear Regression (mLR).
# There are five basic steps when you’re implementing linear regression:
#
# 1. Import the packages and classes you need.
# 2. Provide data to work with and eventually do appropriate transformations.
# 3. Create a regression model and fit it with existing data.
# 4. Check the results of model fitting to know whether the model is satisfactory.
# 5. Apply the model for predictions.
# These steps are more or less general for most of the regression approaches and implementations.
# ## Steps 1 and 2: Import packages and classes, and provide data
#
# First, you import numpy and sklearn.linear_model.LinearRegression and provide known inputs and output:
# +
# First, you import numpy and sklearn.linear_model.LinearRegression and
# provide known inputs and output.
import numpy as np
from sklearn.linear_model import LinearRegression
# That’s a simple way to define the input x and output y.
x = [[1, 2], [3, 3], [2, 2], [4, 3]]
y = [3, 4, 4, 6]
x, y = np.array(x), np.array(y)
# +
# You can print x and y to see how they look now:
print('x looks like: ', x)
print('y looks like: ', y)
# -
# ## Step 3: Create a model and fit it
#
# The next step is to create the regression model as an instance of LinearRegression and fit it with .fit():
# +
# The result of this statement is the variable model referring to the object of type LinearRegression.
# It represents the regression model fitted with existing data.
model = LinearRegression().fit(x, y)
# -
# ## Step 4: Get results
#
# You can obtain the properties of the model the same way as in the case of simple linear regression:
r_sq = model.score(x, y)
print('coefficient of determination:', r_sq)
print('intercept:', model.intercept_)
print('coefficients:', model.coef_)
# You obtain the value of 𝑅² using .score() and the values of the estimators of regression coefficients with .intercept_ and .coef_.
# Again, .intercept_ holds the bias 𝑏₀, while now .coef_ is an array containing 𝑏₁ and 𝑏₂ respectively.
#
# In this example, the intercept is approximately 4.25, and this is the value of the predicted response when 𝑥₁ = 𝑥₂ = 0.
# The increase of 𝑥₁ by 1 yields the rise of the predicted response by 1.5. Similarly, when 𝑥₂ grows by 1, the response
# declined by -1.5.
# ## Step 5: Predict response
#
# Predictions also work the same way as in the case of simple linear regression:
# The predicted response is obtained with .predict()
y_pred = model.predict(x)
print('predicted response:', y_pred, sep='\n')
# You can predict the output values also (which is similar to run ".predict()"), by multiplying each column of the input with the appropriate weight, summing the results and adding the intercept to the sum.
y_pred = model.intercept_ + np.sum(model.coef_ * x, axis=1)
print('predicted response:', y_pred, sep='\n')
# print current date and time
import time
print("date",time.strftime("%d.%m.%Y %H:%M:%S"))
print ("end")
| ML5-Homework-H5_4c.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gensim
import config as cfg
LOGS = False
if LOGS:
import logging
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
# + pycharm={"name": "#%%\n"}
model = gensim.models.KeyedVectors.load_word2vec_format(cfg.RESOURCE.w2v_google, binary=True)
# + pycharm={"name": "#%%\n"}
# model.most_similar(positive=['good', 'shoot'], negative=['bad'])
model.most_similar(positive=['arrivals'])
# + pycharm={"name": "#%%\n"}
from nltk.corpus import wordnet as wn
for syn in wn.synsets("good"):
for l in syn.lemmas():
if l.antonyms():
for ant in l.antonyms():
print(ant.name())
| toxic_language_mining/similarity_search_word_embedding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > **Note:** In most sessions you will be solving exercises posed in a Jupyter notebook that looks like this one. Because you are cloning a Github repository that only we can push to, you should **NEVER EDIT** any of the files you pull from Github. Instead, what you should do, is either make a new notebook and write your solutions in there, or **make a copy of this notebook and save it somewhere else** on your computer, not inside the `sds` folder that you cloned, so you can write your answers in there. If you edit the notebook you pulled from Github, those edits (possible your solutions to the exercises) may be overwritten and lost the next time you pull from Github. This is important, so don't hesitate to ask if it is unclear.
# # Exercise set 1: Introduction to Python
#
# *Morning, August 12, 2019*
#
# In this session you will be working with core Python. We will go over some of the basic operations and functions in Python. You will learn to set up a `for` loop and write your own function.
# ### Exercise Section 1.0.: Check modules are working
#
# Run the cell below to check you have all the relevant packages installed. You should get a scatter plot of `total_bill` and `tip` in the dataset `tips`.
# +
# This part import the relevant packages
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import requests
import seaborn as sns
# We load the dataset 'tips' from the seaborn package, and call the dataset tips
tips = sns.load_dataset('tips')
# We use seaborn to make a jointplot, with total_bill on the x-axis and tip on the y-axis, form the dataset tips.
sns.jointplot(x='total_bill', y='tip', data=tips)
# The plot is displayed
plt.show()
# -
# ## Exercises
# ### Exercise Section 1.1: Basic operations and functions
# We start with some simple procedures for core Python.
# > **Ex. 1.1.1**: Division in Python has two ways of usage. What is the output of `5/2`? What is the output of `5//2`? What is the fundamental data type for each of the two operations? Explain your results.
#
# > Note: Python 2 and 3 are different on what `/` does to integers!
# +
# [Answer to Ex. 1.1.1]
# -
# > **Ex. 1.1.2
# **: What happens if we use the modulus operator instead, i.e. `5%2`? What is the output of `5**2`? Explain your answers.
# +
# [Answer to Ex. 1.1.2]
# -
# > **Ex. 1.1.3**: use expressions `<`, `>`, `==`, `!=` to compare 4 to 6 (e.g. run `4<6`)- what are the input data types? What are the output data types?
# +
# [Answer to Ex. 1.1.3]
# -
# > **Ex. 1.1.4**: Use the functions `abs`, `round` to return the absolute value of -3.14159265 with 3 decimals.
# +
# [Answer to Ex. 1.1.4]
# -
# ### Exercise Section 1.2: Logic and loops
# > **Ex. 1.2.1**: `if` & `else`
# Imagine that we have a car that can fit 5 passengers, and we want to know, if the car can fit more passengers.
# Using 'if' and 'else', make a piece of code that prints out "The car is full" if *passengers* is 5 or more, and "The car is not full yet" if *passengers* are less than 5.
#
# > *Note*: Multiple conditions can be inserted with **elif** statement(s):
#
# +
# [Answer to Ex. 1.2.1]
# -
# > **Ex. 1.2.2**: compute and print each element in the range 1 to 5 cubic (i.e. lifted to power 3) using a `for` loop
#
# > *Hint*: Python's `range` may be useful
# +
# [Answer to Ex. 1.2.2]
# -
# > **Ex. 1.2.3**: Now make a `while` loops that prints the square (i.e. lifted to power 2) of all the values from 1 to 12.
#
# > *Hint*: Add 1 to your iterator variable at the end of each loop. (You can use the += operator this)
# +
# [Answer to Ex. 1.2.3]
# -
# ### Exercise Section 1.3: Container and arrays
# > **Ex. 1.3.1**: make two lists, `A` and `B` with respectively integers 2,3,1 and 3,7,4. What is the index of 3 in list Β?
# +
# [Answer to Ex. 1.3.1]
# -
# > **Ex. 1.3.2**: make a list `C` which consist of first `A`, then `B`. How might we describe this list? Convert `C` into a `numpy array` named `C_a`. What are the dimensions `C` after being converted to an array.
#
# > *Hint 1*: you must import `numpy` before you make the array
#
# > *Hint 2*: numpy arrays have the attribute `shape`.
# +
# [Answer to Ex. 1.3.2]
# -
# It is often useful to access lists through its indices. Let `L=[0,2,1,3]`. Slicing of a list works by returning a subsequence: `B[2:]` returns `[1,3]` and `L[1:3]` returns `[2,1]`.
#
#
# > **Ex. 1.3.3**:
# - Use `in` operator to check if which values are in `B`
# - Slice `B` to print only its second and third element
# +
# [Answer to Ex. 1.3.3]
# -
# We can change the elements in a list. Substitution of an element in the list can be done by slicing.
#
# > **Ex. 1.3.4**: replace middle element of `B` with 9
# +
# [Answer to Ex. 1.3.4]
# -
# > **Ex. 1.3.5**: Use functions for containers: Use `max` to find the maximal value of B, and `sorted` to print the elements in B from low to high
# +
# [Answer to Ex. 1.3.5]
# -
# > **Ex. 1.3.6**: try to add `B` to `A`. What is the new length of `A`? Are lists mutable? (e.i. can you change them?)
#
# > *Hint*: one way of doing this is with `append`. Adding elements to a list can be done as `L.append(O)` where `L` is our list and `O` is the object we add.
#
# > *Note*: More methods for lists can be seen [here](https://docs.python.org/3/tutorial/datastructures.html#more-on-lists), including removal of elements with **pop** and **remove**.
# +
# [Answer to Ex. 1.3.6]
# -
#
# > **Ex. 1.3.7 - BONUS**: Convert `A`, `B` into sets named respectively `A_s`, `B_s`. Combine the set `A_s` and `B_s` into one set `C_s`. How many elements are there in `C_s`?
#
# > Note: read about the `set` container in Python [here](https://docs.python.org/3.6/tutorial/datastructures.html#sets).
# +
# [Answer to Ex. 1.3.7]
# -
# ### Exercise Section 1.4: Reusable code; functions and objects
# > **Ex. 1.4.1**: make a function named `my_subtract` that takes two arguments and subtracts the second argument from the first. Try altering the function where the second argument is equal to 3 as default; can you execute the function with only a single argument?
# +
# [Answer to Ex. 1.4.1]
# -
# Numpy arrays is a class like lists, sets, dictonaries ect. Numpy arrays are both fast and easy to work with, when you are doing linear algebra.
#
# Like other classes Numpy arrays have certain functions, that are specific to Numpy arrays. These functions are called by first specifing the Numpy array, and then the a punctuation follow by the function (e.g. `myarray.sort()`)
#
# > **Ex. 1.4.2**: compute the sum over all elements in the matrix (i.e. 2d-array), `C_a`. Transpose the matrix `C_a`.
#
# > *Hint*: numpy has the function `sum` which may be of use
# +
# [Answer to Ex. 1.4.2]
# -
# > **Ex. 1.4.3 - BONUS**: Make a class called employee, which take the arguments first_name, last_name and pay. Use it to make an object `employee_1`, and then print the first name, last name, and the pay of the employee.
#
#
# > *Hint:* Within the class you need first to define the "\_\_init\_\_" function, which take the argument *self*
# +
# [Answer to Ex. 1.4.3]
| material/session_1/exercise_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import sklearn
import matplotlib.pyplot as plt
from pathlib import Path
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.cluster import KMeans
# +
#1 Read crypto_data.csv into Pandas. The dataset was obtained from CryptoCompare.
crypto_raw_df = pd.read_csv(Path('crypto_data.csv'))
crypto_raw_df.head(5)
# +
#2 Discard all cryptocurrencies that are not being traded. In other words, filter for currencies that are currently being traded.
# Once you have done this, drop the IsTrading column from the dataframe.
traded_cryptos_df = crypto_raw_df.loc[crypto_raw_df["IsTrading"]==True]
traded_cryptos_df = traded_cryptos_df.drop(columns=['Unnamed: 0','IsTrading'],axis=1)
traded_cryptos_df.head(5)
# +
#3 Remove all rows that have at least one null value.
traded_cryptos_df = traded_cryptos_df.dropna(axis=0)
# +
#4 Filter for cryptocurrencies that have been mined. That is, the total coins mined should be greater than zero.
mined_cryptos_df = traded_cryptos_df.loc[traded_cryptos_df.TotalCoinsMined > 0]
# +
# 5. In order for your dataset to be comprehensible to a machine learning algorithm, its data should be numeric.
# Since the coin names do not contribute to the analysis of the data, delete the CoinName from the original dataframe.
mined_cryptos_df = mined_cryptos_df.drop(columns='CoinName',axis=1)
# +
#6 Your next step in data preparation is to convert the remaining features with text values, Algorithm and ProofType,
# into numerical data. To accomplish this task, use Pandas to create dummy variables. Examine the number of rows and
# columns of your dataset now. How did they change?
dummy_df = pd.get_dummies(mined_cryptos_df, columns=['Algorithm', 'ProofType'])
len(dummy_df.columns)
#Data changed in that Algorithm and ProofType values rows became columns
# +
#7 Standardize your dataset so that columns that contain larger values do not unduly influence the outcome.
scaler = StandardScaler()
scaled_dummy_df = scaler.fit_transform(dummy_df)
# +
# Creating dummy variables above dramatically increased the number of features in your dataset.
# Perform dimensionality reduction with PCA. Rather than specify the number of principal components
# when you instantiate the PCA model, it is possible to state the desired explained variance.
pca = PCA(n_components = 5)
pca_fit_df = pca.fit_transform(scaled_dummy_df)
crypto_pca_df = pd.DataFrame(data=pca_fit_df, columns=["PCA1", "PCA2", "PCA3", "PCA4", "PCA5"])
crypto_pca_df.head()
# -
pca.explained_variance_ratio_
# +
# For example, say that a dataset has 100 features. Using PCA(n_components=0.99) creates a model that
# will preserve approximately 99% of the explained variance, whether that means reducing the dataset to
# 80 principal components or 3. For this project, preserve 90% of the explained variance in dimensionality
# reduction. How did the number of the features change?
# +
# Preserve 90% of the explained variance in dimensionality reduction using PCA
pca2 = PCA(n_components=.90)
crypto_pca = pca2.fit_transform(scaled_dummy_df)
crypto_pca
# -
# Transform PCA data to a DataFrame
transformed_crypto_pca = pd.DataFrame(data=crypto_pca)
transformed_crypto_pca.head()
pca.explained_variance_ratio_
features = range(pca.n_components_)
plt.bar(features, pca.explained_variance_ratio_, color='skyblue')
plt.xlabel('PCA Features')
plt.ylabel('% Var')
plt.xticks(features)
# Number of features
len(transformed_crypto_pca.columns)
# +
# Next, further reduce the dataset dimensions with t-SNE and visually inspect the results.
# In order to accomplish this task, run t-SNE on the principal components: the output of the PCA transformation.
# Then create a scatter plot of the t-SNE output. Observe whether there are distinct clusters or not.
tsne = TSNE(learning_rate=35)
# -
tsne_features = tsne.fit_transform(transformed_crypto_pca)
tsne_features.shape
# +
x = tsne_features[:,0]
y = tsne_features[:,1]
plt.scatter(x, y)
plt.show()
# +
#Create an elbow plot to identify the best number of clusters. Use a for-loop to determine the inertia for each k between 1 through 10. Determine, if possible, where the elbow of the plot is,
#and at which value of k it appears.
inertia_container = []
l_range = list(range(1, 11))
for i in l_range:
km = KMeans(n_clusters=i, random_state=42)
km.fit(transformed_crypto_pca)
inertia_container.append(km.inertia_)
# Elbow Curve
elbow_data = {"K": l_range, "Inertia": inertia_container}
df_elbow = pd.DataFrame(elbow_data)
plt.plot(df_elbow['K'], df_elbow['Inertia'])
plt.xticks(range(1,11))
plt.xlabel('# Clusters')
plt.ylabel('Inertia')
plt.show()
# +
# There isn't a discernable elbow in the graph. THe K-value beyond changes, albiet a very small amount.
| .ipynb_checkpoints/crypto_compare-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import numpy as np
import pandas as pd
import collections
from sklearn import metrics
from sklearn.preprocessing import LabelEncoder
import tensorflow as tf
from sklearn.model_selection import train_test_split
from unidecode import unidecode
from tqdm import tqdm
import time
rules_normalizer = {
'experience': 'pengalaman',
'bagasi': 'bagasi',
'kg': 'kampung',
'kilo': 'kilogram',
'g': 'gram',
'grm': 'gram',
'k': 'okay',
'abgkat': 'abang dekat',
'abis': 'habis',
'ade': 'ada',
'adoi': 'aduh',
'adoii': 'aduhh',
'aerodarat': 'kapal darat',
'agkt': 'angkat',
'ahh': 'ah',
'ailior': 'air liur',
'airasia': 'air asia x',
'airasiax': 'penerbangan',
'airline': 'penerbangan',
'airlines': 'penerbangan',
'airport': 'lapangan terbang',
'airpot': 'lapangan terbang',
'aje': 'sahaja',
'ajelah': 'sahajalah',
'ajer': 'sahaja',
'ak': 'aku',
'aq': 'aku',
'all': 'semua',
'ambik': 'ambil',
'amek': 'ambil',
'amer': 'amir',
'amik': 'ambil',
'ana': 'saya',
'angkt': 'angkat',
'anual': 'tahunan',
'apapun': 'apa pun',
'ape': 'apa',
'arab': 'arab',
'area': 'kawasan',
'aritu': 'hari itu',
'ask': 'tanya',
'astro': 'astro',
'at': 'pada',
'attitude': 'sikap',
'babi': 'khinzir',
'back': 'belakang',
'bag': 'beg',
'bang': 'abang',
'bangla': 'bangladesh',
'banyk': 'banyak',
'bard': 'pujangga',
'bargasi': 'bagasi',
'bawak': 'bawa',
'bawanges': 'bawang',
'be': 'jadi',
'behave': 'berkelakuan baik',
'belagak': 'berlagak',
'berdisiplin': 'berdisplin',
'berenti': 'berhenti',
'beskal': 'basikal',
'bff': 'rakan karib',
'bg': 'bagi',
'bgi': 'bagi',
'biase': 'biasa',
'big': 'besar',
'bike': 'basikal',
'bile': 'bila',
'binawe': 'binatang',
'bini': 'isteri',
'bkn': 'bukan',
'bla': 'bila',
'blom': 'belum',
'bnyak': 'banyak',
'body': 'tubuh',
'bole': 'boleh',
'boss': 'bos',
'bowling': 'boling',
'bpe': 'berapa',
'brand': 'jenama',
'brg': 'barang',
'briefing': 'taklimat',
'brng': 'barang',
'bro': 'abang',
'bru': 'baru',
'bruntung': 'beruntung',
'bsikal': 'basikal',
'btnggjwb': 'bertanggungjawab',
'btul': 'betul',
'buatlh': 'buatlah',
'buh': 'letak',
'buka': 'buka',
'but': 'tetapi',
'bwk': 'bawa',
'by': 'dengan',
'byr': 'bayar',
'bz': 'sibuk',
'camera': 'kamera',
'camni': 'macam ini',
'cane': 'macam mana',
'cant': 'tak boleh',
'carakerja': 'cara kerja',
'care': 'jaga',
'cargo': 'kargo',
'cctv': 'kamera litar tertutup',
'celako': 'celaka',
'cer': 'cerita',
'cheap': 'murah',
'check': 'semak',
'ciput': 'sedikit',
'cite': 'cerita',
'citer': 'cerita',
'ckit': 'sikit',
'ckp': 'cakap',
'class': 'kelas',
'cm': 'macam',
'cmni': 'macam ini',
'cmpak': 'campak',
'committed': 'komited',
'company': 'syarikat',
'complain': 'aduan',
'corn': 'jagung',
'couldnt': 'tak boleh',
'cr': 'cari',
'crew': 'krew',
'cube': 'cuba',
'cuma': 'cuma',
'curinyaa': 'curinya',
'cust': 'pelanggan',
'customer': 'pelanggan',
'd': 'di',
'da': 'dah',
'dn': 'dan',
'dahh': 'dah',
'damaged': 'rosak',
'dapek': 'dapat',
'day': 'hari',
'dazrin': 'dazrin',
'dbalingnya': 'dibalingnya',
'de': 'ada',
'deep': 'dalam',
'deliberately': 'sengaja',
'depa': 'mereka',
'dessa': 'desa',
'dgn': 'dengan',
'dh': 'dah',
'didunia': 'di dunia',
'diorang': 'mereka',
'diorng': 'mereka',
'direct': 'secara terus',
'diving': 'junam',
'dkt': 'dekat',
'dlempar': 'dilempar',
'dlm': 'dalam',
'dlt': 'padam',
'dlu': 'dulu',
'done': 'siap',
'dont': 'jangan',
'dorg': 'mereka',
'dpermudhkn': 'dipermudahkan',
'dpt': 'dapat',
'dr': 'dari',
'dri': 'dari',
'dsb': 'dan sebagainya',
'dy': 'dia',
'educate': 'mendidik',
'ensure': 'memastikan',
'everything': 'semua',
'ewahh': 'wah',
'expect': 'sangka',
'fb': 'facebook',
'fired': 'pecat',
'first': 'pertama',
'fkr': 'fikir',
'flight': 'kapal terbang',
'for': 'untuk',
'free': 'percuma',
'friend': 'kawan',
'fyi': 'untuk pengetahuan anda',
'gantila': 'gantilah',
'gantirugi': 'ganti rugi',
'gentlemen': 'lelaki budiman',
'gerenti': 'jaminan',
'gile': 'gila',
'gk': 'juga',
'gnti': 'ganti',
'go': 'pergi',
'gomen': 'kerajaan',
'goment': 'kerajaan',
'good': 'baik',
'ground': 'tanah',
'guarno': 'macam mana',
'hampa': 'mereka',
'hampeh': 'teruk',
'hanat': 'jahanam',
'handle': 'kawal',
'handling': 'kawalan',
'hanta': 'hantar',
'haritu': 'hari itu',
'hate': 'benci',
'have': 'ada',
'hawau': 'celaka',
'henpon': 'telefon',
'heran': 'hairan',
'him': 'dia',
'his': 'dia',
'hmpa': 'mereka',
'hntr': 'hantar',
'hotak': 'otak',
'hr': 'hari',
'i': 'saya',
'hrga': 'harga',
'hrp': 'harap',
'hu': 'sedih',
'humble': 'merendah diri',
'ibon': 'ikon',
'ichi': 'inci',
'idung': 'hidung',
'if': 'jika',
'ig': 'instagram',
'iklas': 'ikhlas',
'improve': 'menambah baik',
'in': 'masuk',
'isn t': 'tidak',
'isyaallah': 'insyallah',
'ja': 'sahaja',
'japan': 'jepun',
'jd': 'jadi',
'je': 'saja',
'jee': 'saja',
'jek': 'saja',
'jepun': 'jepun',
'jer': 'saja',
'jerr': 'saja',
'jez': 'saja',
'jg': 'juga',
'jgk': 'juga',
'jgn': 'jangan',
'jgnla': 'janganlah',
'jibake': 'celaka',
'jjur': 'jujur',
'job': 'kerja',
'jobscope': 'skop kerja',
'jogja': 'jogjakarta',
'jpam': 'jpam',
'jth': 'jatuh',
'jugak': 'juga',
'ka': 'ke',
'kalo': 'kalau',
'kalu': 'kalau',
'kang': 'nanti',
'kantoi': 'temberang',
'kasi': 'beri',
'kat': 'dekat',
'kbye': 'ok bye',
'kearah': 'ke arah',
'kecik': 'kecil',
'keja': 'kerja',
'keje': 'kerja',
'kejo': 'kerja',
'keksongan': 'kekosongan',
'kemana': 'ke mana',
'kene': 'kena',
'kenekan': 'kenakan',
'kesah': 'kisah',
'ketempat': 'ke tempat',
'kije': 'kerja',
'kijo': 'kerja',
'kiss': 'cium',
'kite': 'kita',
'kito': 'kita',
'kje': 'kerja',
'kjr': 'kerja',
'kk': 'okay',
'kmi': 'kami',
'kt': 'kat',
'tlg': 'tolong',
'kl': 'kuala lumpur',
'klai': 'kalau',
'klau': 'kalau',
'klia': 'klia',
'klo': 'kalau',
'klu': 'kalau',
'kn': 'kan',
'knapa': 'kenapa',
'kne': 'kena',
'ko': 'kau',
'kompom': 'sah',
'korang': 'kamu semua',
'korea': 'korea',
'korg': 'kamu semua',
'kot': 'mungkin',
'krja': 'kerja',
'ksalahan': 'kesalahan',
'kta': 'kita',
'kuar': 'keluar',
'kut': 'mungkin',
'la': 'lah',
'laa': 'lah',
'lahabau': 'celaka',
'lahanat': 'celaka',
'lainda': 'lain dah',
'lak': 'pula',
'last': 'akhir',
'le': 'lah',
'leader': 'ketua',
'leave': 'pergi',
'ler': 'lah',
'less': 'kurang',
'letter': 'surat',
'lg': 'lagi',
'lgi': 'lagi',
'lngsong': 'langsung',
'lol': 'hehe',
'lorr': 'lah',
'low': 'rendah',
'lps': 'lepas',
'luggage': 'bagasi',
'lumbe': 'lumba',
'lyak': 'layak',
'maap': 'maaf',
'maapkan': 'maafkan',
'mahai': 'mahal',
'mampos': 'mampus',
'mart': 'kedai',
'mau': 'mahu',
'mcm': 'macam',
'mcmtu': 'macam itu',
'memerlukn': 'memerlukan',
'mengembirakan': 'menggembirakan',
'mengmbilnyer': 'mengambilnya',
'mengtasi': 'mengatasi',
'mg': 'memang',
'mihak': 'memihak',
'min': 'admin',
'mingu': 'minggu',
'mintak': 'minta',
'mjtuhkn': 'menjatuhkan',
'mkyong': 'mak yong',
'mlibatkn': 'melibatkan',
'mmg': 'memang',
'mmnjang': 'memanjang',
'mmpos': 'mampus',
'mn': 'mana',
'mna': 'mana',
'mntak': 'minta',
'mntk': 'minta',
'mnyusun': 'menyusun',
'mood': 'suasana',
'most': 'paling',
'mr': 'tuan',
'msa': 'masa',
'msia': 'malaysia',
'mst': 'mesti',
'mu': 'awak',
'much': 'banyak',
'muko': 'muka',
'mum': 'emak',
'n': 'dan',
'nah': 'nah',
'nanny': 'nenek',
'napo': 'kenapa',
'nati': 'nanti',
'ngan': 'dengan',
'ngn': 'dengan',
'ni': 'ini',
'nie': 'ini',
'nii': 'ini',
'nk': 'nak',
'nmpk': 'nampak',
'nye': 'nya',
'ofis': 'pejabat',
'ohh': 'oh',
'oii': 'hoi',
'one': 'satu',
'online': 'dalam talian',
'or': 'atau',
'org': 'orang',
'orng': 'orang',
'otek': 'otak',
'p': 'pergi',
'paid': 'dah bayar',
'palabana': 'kepala otak',
'pasni': 'lepas ini',
'passengers': 'penumpang',
'passengger': 'penumpang',
'pastu': 'lepas itu',
'pd': 'pada',
'pegi': 'pergi',
'pekerje': 'pekerja',
'pekrja': 'pekerja',
'perabih': 'perabis',
'perkerja': 'pekerja',
'pg': 'pergi',
'phuii': 'puih',
'pikir': 'fikir',
'pilot': 'juruterbang',
'pk': 'fikir',
'pkerja': 'pekerja',
'pkerjaan': 'pekerjaan',
'pki': 'pakai',
'please': 'tolong',
'pls': 'tolong',
'pn': 'pun',
'pnh': 'pernah',
'pnt': 'penat',
'pnya': 'punya',
'pon': 'pun',
'priority': 'keutamaan',
'properties': 'harta benda',
'ptugas': 'petugas',
'pub': 'kelab malam',
'pulak': 'pula',
'puye': 'punya',
'pwrcuma': 'percuma',
'pyahnya': 'payahnya',
'quality': 'kualiti',
'quit': 'keluar',
'ramly': 'ramly',
'rege': 'harga',
'reger': 'harga',
'report': 'laporan',
'resigned': 'meletakkan jawatan',
'respect': 'hormat',
'rizal': 'rizal',
'rosak': 'rosak',
'rosok': 'rosak',
'rse': 'rasa',
'sacked': 'buang',
'sado': 'tegap',
'salute': 'sanjung',
'sam': 'sama',
'same': 'sama',
'samp': 'sampah',
'sbb': 'sebab',
'sbgai': 'sebagai',
'sblm': 'sebelum',
'sblum': 'sebelum',
'sbnarnya': 'sebenarnya',
'sbum': 'sebelum',
'sdg': 'sedang',
'sebb': 'sebab',
'sebijik': 'sebiji',
'see': 'lihat',
'seen': 'dilihat',
'selangor': 'selangor',
'selfie': 'swafoto',
'sempoi': 'cantik',
'senaraihitam': 'senarai hitam',
'seorg': 'seorang',
'service': 'perkhidmatan',
'sgt': 'sangat',
'shared': 'kongsi',
'shirt': 'kemeja',
'shut': 'tutup',
'sib': 'nasib',
'skali': 'sekali',
'sket': 'sikit',
'sma': 'sama',
'smoga': 'semoga',
'smpoi': 'cantik',
'sndiri': 'sendiri',
'sndr': 'sendiri',
'sndri': 'sendiri',
'sne': 'sana',
'so': 'jadi',
'sop': 'tatacara pengendalian piawai',
'sorang': 'seorang',
'spoting': 'pembintikan',
'sronok': 'seronok',
'ssh': 'susah',
'staff': 'staf',
'standing': 'berdiri',
'start': 'mula',
'steady': 'mantap',
'stiap': 'setiap',
'stress': 'stres',
'student': 'pelajar',
'study': 'belajar',
'studycase': 'kajian kes',
'sure': 'pasti',
'sykt': 'syarikat',
'tah': 'entah',
'taik': 'tahi',
'takan': 'tak akan',
'takat': 'setakat',
'takde': 'tak ada',
'takkan': 'tak akan',
'taknak': 'tak nak',
'tang': 'tentang',
'tanggungjawab': 'bertanggungjawab',
'taraa': 'sementara',
'tau': 'tahu',
'tbabit': 'terbabit',
'team': 'pasukan',
'terbaekk': 'terbaik',
'teruknye': 'teruknya',
'tgk': 'tengok',
'that': 'itu',
'thinking': 'fikir',
'those': 'itu',
'time': 'masa',
'tk': 'tak',
'tnggongjwb': 'tanggungjawab',
'tngok': 'tengok',
'tngu': 'tunggu',
'to': 'kepada',
'tosak': 'rosak',
'tp': 'tapi',
'tpi': 'tapi',
'tpon': 'telefon',
'transfer': 'pindah',
'trgelak': 'tergelak',
'ts': 'tan sri',
'tstony': 'tan sri tony',
'tu': 'itu',
'tuh': 'itu',
'tula': 'itulah',
'umeno': 'umno',
'unfortunately': 'malangnya',
'unhappy': 'tidak gembira',
'up': 'naik',
'upkan': 'naikkan',
'ur': 'awak',
'utk': 'untuk',
'very': 'sangat',
'viral': 'tular',
'vote': 'undi',
'warning': 'amaran',
'warranty': 'waranti',
'wassap': 'whatsapp',
'wat': 'apa',
'weii': 'wei',
'well': 'maklumlah',
'win': 'menang',
'with': 'dengan',
'wt': 'buat',
'x': 'tak',
'tw': 'tahu',
'ye': 'ya',
'yee': 'ya',
'yg': 'yang',
'yng': 'yang',
'you': 'awak',
'your': 'awak',
'sakai': 'selekeh',
'rmb': 'billion ringgit',
'rmj': 'juta ringgit',
'rmk': 'ribu ringgit',
'rm': 'ringgit',
}
# +
permulaan = [
'bel',
'se',
'ter',
'men',
'meng',
'mem',
'memper',
'di',
'pe',
'me',
'ke',
'ber',
'pen',
'per',
]
hujung = ['kan', 'kah', 'lah', 'tah', 'nya', 'an', 'wan', 'wati', 'ita']
def naive_stemmer(word):
assert isinstance(word, str), 'input must be a string'
hujung_result = [e for e in hujung if word.endswith(e)]
if len(hujung_result):
hujung_result = max(hujung_result, key = len)
if len(hujung_result):
word = word[: -len(hujung_result)]
permulaan_result = [e for e in permulaan if word.startswith(e)]
if len(permulaan_result):
permulaan_result = max(permulaan_result, key = len)
if len(permulaan_result):
word = word[len(permulaan_result) :]
return word
def build_dataset(words, n_words):
count = [['GO', 0], ['PAD', 1], ['EOS', 2], ['UNK', 3]]
counter = collections.Counter(words).most_common(n_words)
count.extend(counter)
dictionary = dict()
for word, _ in count:
dictionary[word] = len(dictionary)
data = list()
unk_count = 0
for word in words:
index = dictionary.get(word, 3)
if index == 0:
unk_count += 1
data.append(index)
count[0][1] = unk_count
reversed_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
return data, count, dictionary, reversed_dictionary
def classification_textcleaning(string):
string = re.sub(
'http\S+|www.\S+',
'',
' '.join(
[i for i in string.split() if i.find('#') < 0 and i.find('@') < 0]
),
)
string = unidecode(string).replace('.', ' . ').replace(',', ' , ')
string = re.sub('[^A-Za-z ]+', ' ', string)
string = re.sub(r'[ ]+', ' ', string.lower()).strip()
string = [rules_normalizer.get(w, w) for w in string.split()]
string = [naive_stemmer(word) for word in string]
return ' '.join([word for word in string if len(word) > 1])
def str_idx(corpus, dic, maxlen, UNK = 3):
X = np.zeros((len(corpus), maxlen))
for i in range(len(corpus)):
for no, k in enumerate(corpus[i].split()[:maxlen][::-1]):
X[i, -1 - no] = dic.get(k, UNK)
return X
# -
classification_textcleaning('kerajaan sebenarnya sangat bencikan rakyatnya, minyak naik dan segalanya')
# +
df = pd.read_csv('sentiment-data-v2.csv')
Y = LabelEncoder().fit_transform(df.label)
with open('polarity-negative-translated.txt','r') as fopen:
texts = fopen.read().split('\n')
labels = [0] * len(texts)
with open('polarity-positive-translated.txt','r') as fopen:
positive_texts = fopen.read().split('\n')
labels += [1] * len(positive_texts)
texts += positive_texts
texts += df.iloc[:,1].tolist()
labels += Y.tolist()
assert len(labels) == len(texts)
# +
import json
with open('bm-amazon.json') as fopen:
amazon = json.load(fopen)
with open('bm-imdb.json') as fopen:
imdb = json.load(fopen)
with open('bm-yelp.json') as fopen:
yelp = json.load(fopen)
texts += amazon['negative']
labels += [0] * len(amazon['negative'])
texts += amazon['positive']
labels += [1] * len(amazon['positive'])
texts += imdb['negative']
labels += [0] * len(imdb['negative'])
texts += imdb['positive']
labels += [1] * len(imdb['positive'])
texts += yelp['negative']
labels += [0] * len(yelp['negative'])
texts += yelp['positive']
labels += [1] * len(yelp['positive'])
# +
import os
for i in [i for i in os.listdir('negative') if 'Store' not in i]:
with open('negative/'+i) as fopen:
a = json.load(fopen)
texts += a
labels += [0] * len(a)
import os
for i in [i for i in os.listdir('positive') if 'Store' not in i]:
with open('positive/'+i) as fopen:
a = json.load(fopen)
texts += a
labels += [1] * len(a)
# -
for i in range(len(texts)):
texts[i] = classification_textcleaning(texts[i])
concat = ' '.join(texts).split()
vocabulary_size = len(list(set(concat)))
data, count, dictionary, rev_dictionary = build_dataset(concat, vocabulary_size)
print('vocab from size: %d'%(vocabulary_size))
print('Most common words', count[4:10])
print('Sample data', data[:10], [rev_dictionary[i] for i in data[:10]])
def str_idx(corpus, dic, maxlen, UNK = 3):
X = np.zeros((len(corpus), maxlen))
for i in range(len(corpus)):
for no, k in enumerate(corpus[i].split()[:maxlen][::-1]):
val = dic[k] if k in dic else UNK
X[i, -1 - no] = val
return X
# +
def attention(inputs, attention_size):
hidden_size = inputs.shape[2].value
w_omega = tf.Variable(
tf.random_normal([hidden_size, attention_size], stddev = 0.1)
)
b_omega = tf.Variable(tf.random_normal([attention_size], stddev = 0.1))
u_omega = tf.Variable(tf.random_normal([attention_size], stddev = 0.1))
with tf.name_scope('v'):
v = tf.tanh(tf.tensordot(inputs, w_omega, axes = 1) + b_omega)
vu = tf.tensordot(v, u_omega, axes = 1, name = 'vu')
alphas = tf.nn.softmax(vu, name = 'alphas')
output = tf.reduce_sum(inputs * tf.expand_dims(alphas, -1), 1)
return output, alphas
class Model:
def __init__(
self,
size_layer,
num_layers,
dimension_output,
learning_rate,
dropout,
dict_size,
):
def cells(size, reuse = False):
return tf.contrib.rnn.DropoutWrapper(
tf.nn.rnn_cell.LSTMCell(
size,
initializer = tf.orthogonal_initializer(),
reuse = reuse,
),
state_keep_prob = dropout,
output_keep_prob = dropout,
)
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None])
encoder_embeddings = tf.Variable(
tf.random_uniform([dict_size, size_layer], -1, 1)
)
encoder_embedded = tf.nn.embedding_lookup(encoder_embeddings, self.X)
for n in range(num_layers):
(out_fw, out_bw), (
state_fw,
state_bw,
) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = cells(size_layer),
cell_bw = cells(size_layer),
inputs = encoder_embedded,
dtype = tf.float32,
scope = 'bidirectional_rnn_%d' % (n),
)
encoder_embedded = tf.concat((out_fw, out_bw), 2)
self.outputs, self.attention = attention(encoder_embedded, size_layer)
W = tf.get_variable(
'w',
shape = (size_layer * 2, 2),
initializer = tf.orthogonal_initializer(),
)
b = tf.get_variable(
'b', shape = (2), initializer = tf.zeros_initializer()
)
self.logits = tf.add(tf.matmul(self.outputs, W), b, name = 'logits')
self.cost = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits = self.logits, labels = self.Y
)
)
self.optimizer = tf.train.AdamOptimizer(
learning_rate = learning_rate
).minimize(self.cost)
self.accuracy = tf.reduce_mean(
tf.cast(tf.nn.in_top_k(self.logits, self.Y, 1), tf.float32)
)
# +
size_layer = 256
num_layers = 2
dropout = 0.8
dimension_output = 2
learning_rate = 1e-4
batch_size = 32
maxlen = 100
dropout = 0.8
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(
size_layer,
num_layers,
dimension_output,
learning_rate,
dropout,
len(dictionary),
)
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'hierarchical/model.ckpt')
# -
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 'alphas' in n.name)
and 'Adam' not in n.name
and 'beta' not in n.name
]
)
strings.split(',')
tf.trainable_variables()
train_X, test_X, train_Y, test_Y = train_test_split(texts,
labels,
test_size = 0.2)
# +
from tqdm import tqdm
import time
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 3, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:%d\n' % (EPOCH))
break
train_acc, train_loss, test_acc, test_loss = 0, 0, 0, 0
pbar = tqdm(
range(0, len(train_X), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
batch_x = str_idx(train_X[i : min(i + batch_size, len(train_X))], dictionary, maxlen)
batch_y = train_Y[i : min(i + batch_size, len(train_X))]
batch_x_expand = np.expand_dims(batch_x,axis = 1)
acc, cost, _ = sess.run(
[model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.Y: batch_y,
model.X: batch_x
},
)
assert not np.isnan(cost)
train_loss += cost
train_acc += acc
pbar.set_postfix(cost = cost, accuracy = acc)
pbar = tqdm(range(0, len(test_X), batch_size), desc = 'test minibatch loop')
for i in pbar:
batch_x = str_idx(test_X[i : min(i + batch_size, len(test_X))], dictionary, maxlen)
batch_y = test_Y[i : min(i + batch_size, len(test_X))]
batch_x_expand = np.expand_dims(batch_x,axis = 1)
acc, cost = sess.run(
[model.accuracy, model.cost],
feed_dict = {
model.Y: batch_y,
model.X: batch_x
},
)
test_loss += cost
test_acc += acc
pbar.set_postfix(cost = cost, accuracy = acc)
train_loss /= len(train_X) / batch_size
train_acc /= len(train_X) / batch_size
test_loss /= len(test_X) / batch_size
test_acc /= len(test_X) / batch_size
if test_acc > CURRENT_ACC:
print(
'epoch: %d, pass acc: %f, current acc: %f'
% (EPOCH, CURRENT_ACC, test_acc)
)
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
else:
CURRENT_CHECKPOINT += 1
print('time taken:', time.time() - lasttime)
print(
'epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'
% (EPOCH, train_loss, train_acc, test_loss, test_acc)
)
EPOCH += 1
# +
real_Y, predict_Y = [], []
pbar = tqdm(
range(0, len(test_X), batch_size), desc = 'validation minibatch loop'
)
for i in pbar:
batch_x = str_idx(test_X[i : min(i + batch_size, len(test_X))], dictionary, maxlen)
batch_y = test_Y[i : min(i + batch_size, len(test_X))]
predict_Y += np.argmax(
sess.run(
model.logits, feed_dict = {model.X: batch_x, model.Y: batch_y}
),
1,
).tolist()
real_Y += batch_y
# -
print(
metrics.classification_report(
real_Y, predict_Y, target_names = ['negative', 'positive']
)
)
text = classification_textcleaning('kerajaan sebenarnya sangat bencikan rakyatnya, minyak naik dan segalanya')
new_vector = str_idx([text], dictionary, len(text.split()))
sess.run(tf.nn.softmax(model.logits), feed_dict={model.X:new_vector})
import json
with open('hierarchical-sentiment.json','w') as fopen:
fopen.write(json.dumps({'dictionary':dictionary,'reverse_dictionary':rev_dictionary}))
saver.save(sess, 'hierarchical/model.ckpt')
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('hierarchical', strings)
def load_graph(frozen_graph_filename):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
g = load_graph('hierarchical/frozen_model.pb')
x = g.get_tensor_by_name('import/Placeholder:0')
logits = g.get_tensor_by_name('import/logits:0')
alphas = g.get_tensor_by_name('import/alphas:0')
test_sess = tf.InteractiveSession(graph = g)
result = test_sess.run([logits, alphas], feed_dict = {x: new_vector})
| session/sentiment/hierarchical.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating your own GAN III: LR-GAN
#
# In the last notebook we implemented the `Pix2Pix` GAN together, this time we're tackling the Latent-Regressor GAN. It is described in one of our favorite papers describing a really amazing algorithm: The [BicycleGAN](https://arxiv.org/pdf/1711.11586.pdf). This will soon be implemented as well in `vegan`. As the time of writing this notebook (2021-04-08 20:22, only one hour passed since we started implementing the Pix2Pix :) ) there are only 4 (9) GAN architectures implemented in `vegans`: `VanillaGAN`, `WasssersteinGAN`, `WassersteinGANGP`, `LSGAN` and all there conditional variants plus the Pix2Pix which only is a conditional algorithm. The LR-GAN finally tackles the problem of mode collapse which means that all random inputs into the generator are mapped to a single output which might fool the discriminator perfectly but does not look like a real image to humans. Even if i does it is not desirable as we want a high variety of output images. This is one of the most pressing problems so it is good that we finally deal with it (somewhat).
#
# We suppose you have read the previous notebooks on the creation of the `LSGAN` and `Pix2Pix`. If not, please go over it so you have a understanding of the abstract base classes. We will not present it here and jump basically right into the implementation.
#
# First import the usual libraries:
# +
import os
import torch
os.chdir("/home/thomas/Backup/Algorithmen/GAN-pytorch")
from vegans.GAN import ConditionalWassersteinGAN, ConditionalWassersteinGANGP
from vegans.utils.utils import plot_losses, plot_images, get_input_dim
# -
# ## Latent-Regressor GAN
#
# Remember that so far we only looked at two networks learning against each other. A discriminator which is trained to differentiate between real and fake images and a generator which is trained to fool the discriminator. As mentioned above this quite often leads to mode collapse where the generator produces a single "perfect" image regardless of input. In the well known example of generating handwritten digits, the generator might learn to produce the perfect image of a zero but produces nothing else. The discriminator can never tell if the generator images are real or fake so the generator is content and will stop learning. The latent regressor GAN approaches this problem elegantly by introducing another helper network (Note that another technique to deal with this is minibatch discrimination which is not yet supported in `vegan`, but hopefully is in the future (it might even be implemented by the time you read this)).
#
# The job of this helper network is to take the output of the generator (often a generated image) and compress it back into the latent space. So if we started from a random image (say with shape [1, 4, 4]) the generator produces an image from it and the helper network maps at back to a space of the initial dimension (again [1, 4, 4]). For this reason this network is called an **Encoder**.
# The output of the encoder is then compared to the initial latent input to the generator and a L1 (L2) norm is computed between the two. The goal of the generator (and encoder) is to minimise this L1 (L2) norm. That's why it is called a Latent-Space Regressor.
#
# This helps against mode collapse because if every latent input to the generator is mapped to basically the same output image the encoder will transform all those images back to one single latent vector. This vector will most of the times be quite different from the original input so the L1 norm increases which the generator has to minimize. Therefore it needs to produce a reproducible output for every input. The schema looks like this (again taken from the paper for BiCycleGAN):
#
# 
#
# Let's now start with the implementation of the LRGAN. First note that unlike before we can't use the parent class `AbstractGAN1v1` because we now have three networks working against each other. There is no base class for this case in `vegan` (yet), so we can only inherit from `AbstractGenerativeModel` and do a lot of the footwork on our own. This means we need to implement all abstract methods:
#
# - __init__(self, x_dim, z_dim, optim, optim_kwargs, fixed_noise_size, device, folder, ngpu):
# This takes care of the initializaton and the method
#
# super().__init__(
# x_dim=x_dim, z_dim=z_dim, optim=optim, optim_kwargs=optim_kwargs,
# fixed_noise_size=fixed_noise_size, device=device, folder=folder, ngpu=ngpu
# )
#
# must be called at the end of the `__init__` method.
#
# - _default_optimizer(self): returns an optimizer from torch.optim that is used if the user doesn't specify any optimizers in the `optim` keyword when constructing a class.
#
# - _define_loss(self): Not strictly necessary but it is still kept as an abstract method so that the user has to think about what he wants to implement here. You can also implement it with a single `pass` statement. However, we will show you it's intended use here.
#
# - calculate_losses(self, X_batch, Z_batch, who): The core function that needs to be implemented. For every batch it must populate an already existing (but empty) dictionary `self._losses`. The keys of the dictionary must include at least the keys used in `self.neural_nets` (explained below), but can also contain other losses. We will discuss this further in later implementations.
#
# We start with the `__init__` method which **MUST** create the self.neural_nets dictionary. No worries, if you forget to specify it the `AbstractGenerativeModel` class will kindly remind you to populate the dictionary. I will copy some of the code from [AbstractGAN1v1](https://github.com/tneuer/GAN-pytorch/blob/main/vegans/models/unconditional/AbstractGAN1v1.py) just to get a feeling for how to start. I will reuse some of the code of course because that's what past-me would have wanted.
from vegans.models.unconditional.AbstractGenerativeModel import AbstractGenerativeModel
from torch.nn import BCELoss, L1Loss
from torch.nn import MSELoss as L2Loss
from vegans.utils.networks import Generator, Adversariat, Encoder
class LRGAN(AbstractGenerativeModel):
#########################################################################
# Actions before training
#########################################################################
def __init__(
self,
generator,
adversariat,
encoder,
x_dim,
z_dim,
optim=None,
optim_kwargs=None,
lambda_L1=10,
fixed_noise_size=32,
device=None,
folder="./AbstractGAN1v1",
ngpu=0):
if device is None:
device = "cuda" if torch.cuda.is_available() else "cpu"
self.generator = Generator(generator, input_size=z_dim, device=device, ngpu=ngpu)
self.adversariat = Adversariat(adversariat, input_size=x_dim, adv_type="Discriminator", device=device, ngpu=ngpu)
self.encoder = Encoder(encoder, input_size=x_dim, device=device, ngpu=ngpu)
self.neural_nets = {
"Generator": self.generator, "Adversariat": self.adversariat, "Encoder": self.encoder
}
AbstractGenerativeModel.__init__(
self, x_dim=x_dim, z_dim=z_dim, optim=optim, optim_kwargs=optim_kwargs,
fixed_noise_size=fixed_noise_size, device=device, folder=folder, ngpu=ngpu
)
self.lambda_L1 = lambda_L1
self.hyperparameters["lambda_L1"] = lambda_L1
def _default_optimizer(self):
return torch.optim.Adam
def _define_loss(self):
self.loss_functions = {"Generator": BCELoss(), "Adversariat": BCELoss(), "L1": L1Loss()}
# I included the construction of three `vegan` classes in the vegans.utils.networks module: Generator, Adversariat, Encoder. If you are sneaky and look at the [code](https://github.com/tneuer/GAN-pytorch/blob/main/vegans/utils/networks.py) you will notice that there is absolutely no difference between the implementations of Generator and Encoder (apart from the `name` attribute which only shows up when printing the network). On an abstract level both do the same thing: take an input (image or vector), applying weights and biases and finally producing an output (image or vector). Everything else is implementation detail. We also already implemented the `_default_optimizer` method (which is always the easiest part, mostly `torch.optim.Adam`) and the `_define_loss` method which includes the L1 loss for the encoder and generator. We also included the parameter `lambda_L1` which will be later used to weight the L1 metric in the generator loss.
#
# If you have worked through the previous notebooks you might note that the names in the self.neural_nets dictionary are extremely important! They will be from now in used in every relevant dictionary (`self.optimizers`, `self.steps`, `self._losses` to refer to this network. Give it a descriptive but concise name).
#
# Next we will implement the last abstract method: calculate_losses(...)! I again copy and modify existing code from the [AbstractGAN1v1](https://github.com/tneuer/GAN-pytorch/blob/main/vegans/models/unconditional/AbstractGAN1v1.py) class.
class LRGAN(AbstractGenerativeModel):
#########################################################################
# Actions before training
#########################################################################
def __init__(
self,
generator,
adversariat,
encoder,
x_dim,
z_dim,
optim=None,
optim_kwargs=None,
lambda_L1=10,
fixed_noise_size=32,
device=None,
folder="./AbstractGAN1v1",
ngpu=0):
if device is None:
device = "cuda" if torch.cuda.is_available() else "cpu"
self.generator = Generator(generator, input_size=z_dim, device=device, ngpu=ngpu)
self.adversariat = Adversariat(adversariat, input_size=x_dim, adv_type="Discriminator", device=device, ngpu=ngpu)
self.encoder = Encoder(encoder, input_size=x_dim, device=device, ngpu=ngpu)
self.neural_nets = {
"Generator": self.generator, "Adversariat": self.adversariat, "Encoder": self.encoder
}
AbstractGenerativeModel.__init__(
self, x_dim=x_dim, z_dim=z_dim, optim=optim, optim_kwargs=optim_kwargs,
fixed_noise_size=fixed_noise_size, device=device, folder=folder, ngpu=ngpu
)
self.lambda_L1 = lambda_L1
self.hyperparameters["lambda_L1"] = lambda_L1
def _default_optimizer(self):
return torch.optim.Adam
def _define_loss(self):
self.loss_functions = {"Generator": BCELoss(), "Adversariat": BCELoss(), "L1": L1Loss()}
#########################################################################
# Actions during training
#########################################################################
def encode(self, x):
return self.encoder(x)
def calculate_losses(self, X_batch, Z_batch, who=None):
if who == "Generator":
self._calculate_generator_loss(X_batch=X_batch, Z_batch=Z_batch)
elif who == "Adversariat":
self._calculate_adversariat_loss(X_batch=X_batch, Z_batch=Z_batch)
elif who == "Encoder":
self._calculate_encoder_loss(X_batch=X_batch, Z_batch=Z_batch)
else:
self._calculate_generator_loss(X_batch=X_batch, Z_batch=Z_batch)
self._calculate_adversariat_loss(X_batch=X_batch, Z_batch=Z_batch)
self._calculate_encoder_loss(X_batch=X_batch, Z_batch=Z_batch)
def _calculate_generator_loss(self, X_batch, Z_batch):
fake_images = self.generate(z=Z_batch)
fake_predictions = self.predict(x=fake_images)
encoded_space = self.encode(x=fake_images)
gen_loss_original = self.loss_functions["Generator"](
fake_predictions, torch.ones_like(fake_predictions, requires_grad=False)
)
latent_space_regression = self.loss_functions["L1"](
encoded_space, Z_batch
)
gen_loss = gen_loss_original + self.lambda_L1*latent_space_regression
self._losses.update({
"Generator": gen_loss,
"Generator_Original": gen_loss_original,
"Generator_L1": latent_space_regression
})
def _calculate_adversariat_loss(self, X_batch, Z_batch):
fake_images = self.generate(z=Z_batch).detach()
fake_predictions = self.predict(x=fake_images)
real_predictions = self.predict(x=X_batch.float())
adv_loss_fake = self.loss_functions["Adversariat"](
fake_predictions, torch.zeros_like(fake_predictions, requires_grad=False)
)
adv_loss_real = self.loss_functions["Adversariat"](
real_predictions, torch.ones_like(real_predictions, requires_grad=False)
)
adv_loss = 0.5*(adv_loss_fake + adv_loss_real)
self._losses.update({
"Adversariat": adv_loss,
"Adversariat_fake": adv_loss_fake,
"Adversariat_real": adv_loss_real,
"RealFakeRatio": adv_loss_real / adv_loss_fake
})
def _calculate_encoder_loss(self, X_batch, Z_batch):
fake_images = self.generate(z=Z_batch).detach()
encoded_space = self.encoder(fake_images)
latent_space_regression = self.loss_functions["L1"](
encoded_space, Z_batch
)
self._losses.update({
"Encoder": latent_space_regression
})
# This should do the trick and wasn't to complicated. To implement the conditional version allow for `y_dim`, `y_batch` and concatenate at the correct positions. Look at the source code to see that it is almost the same :)
# Please again note that this is a prelimanary tutorial implementation which might or might not change in future releases of `vegan`. So this implementation might not be completely up-to-date, but still is a viable implementation nonetheless.
| notebooks/07_create-your-own-LR-GAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
a = {'a':1,'b':2,'c':3}
if 'a' in a:
print("yes")
if 'a' in a.keys():
print('yes2')
print(a.keys())
print(a.items())
# -
print(a)
a.update({'d':123,'e':456,'f':789})
print(a)
for x in a.items():
print(type(x))
print(x[0],":",x[1])
print(a.get('c'))
print(a.get('c',234567))
print(a.get('g'))
print(a.get('g',9876))
import pandas as pd
# +
df = pd.read_csv('../data/surnames/surnames_with_splits.csv')
print(df.head())
print(df.iloc[1])
# -
class Test(object):
def __init__(self):
print("init")
self.a = [1,3,34,4,5,6,76,7,8,9]
def __str__(self):
return "%d" % len(self)
def __len__(self):
return len(self.a)
a = Test()
a.__str__
print(a)
# +
import numpy as np
import torch
hiddens = []
for x in range(5):
hiddens.append(torch.ones(5) * x)
print(hiddens)
#hiddens = torch.from_numpy(hiddens)
hiddens = torch.stack(hiddens)
print(hiddens)
# +
print(hiddens.size())
hiddens = torch.unsqueeze(hiddens,0)
print(hiddens.size())
print(hiddens)
hiddens = hiddens.permute(1, 0, 2)
print(hiddens.size())
print(hiddens)
# -
| chapters/ttttttest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="TBFXQGKYUc4X"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="1z4xy2gTUc4a"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="FE7KNzPPVrVV"
# # Dogs vs Cats Image Classification With Image Augmentation
# + [markdown] colab_type="text" id="KwQtSOz0VrVX"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l05c02_dogs_vs_cats_with_augmentation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l05c02_dogs_vs_cats_with_augmentation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="gN7G9GFmVrVY"
# In this tutorial, we will discuss how to classify images into pictures of cats or pictures of dogs. We'll build an image classifier using `tf.keras.Sequential` model and load data using `tf.keras.preprocessing.image.ImageDataGenerator`.
#
# ## Specific concepts that will be covered:
# In the process, we will build practical experience and develop intuition around the following concepts
#
# * Building _data input pipelines_ using the `tf.keras.preprocessing.image.ImageDataGenerator` class — How can we efficiently work with data on disk to interface with our model?
# * _Overfitting_ - what is it, how to identify it, and how can we prevent it?
# * _Data Augmentation_ and _Dropout_ - Key techniques to fight overfitting in computer vision tasks that we will incorporate into our data pipeline and image classifier model.
#
# ## We will follow the general machine learning workflow:
#
# 1. Examine and understand data
# 2. Build an input pipeline
# 3. Build our model
# 4. Train our model
# 5. Test our model
# 6. Improve our model/Repeat the process
#
# <hr>
#
# **Before you begin**
#
# Before running the code in this notebook, reset the runtime by going to **Runtime -> Reset all runtimes** in the menu above. If you have been working through several notebooks, this will help you avoid reaching Colab's memory limits.
#
# + [markdown] colab_type="text" id="zF9uvbXNVrVY"
# # Importing packages
# + [markdown] colab_type="text" id="VddxeYBEVrVZ"
# Let's start by importing required packages:
#
# * os — to read files and directory structure
# * numpy — for some matrix math outside of TensorFlow
# * matplotlib.pyplot — to plot the graph and display images in our training and validation data
# + colab={} colab_type="code" id="rtPGh2MAVrVa"
from __future__ import absolute_import, division, print_function, unicode_literals
# + colab={} colab_type="code" id="in3OdvpUG_9_"
try:
# Use the %tensorflow_version magic if in colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
# + colab={} colab_type="code" id="L1WtoaOHVrVh"
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# + colab={} colab_type="code" id="ede3_kbeHOjR"
import os
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] colab_type="text" id="UZZI6lNkVrVm"
# # Data Loading
# + [markdown] colab_type="text" id="DPHx8-t-VrVo"
# To build our image classifier, we begin by downloading the dataset. The dataset we are using is a filtered version of <a href="https://www.kaggle.com/c/dogs-vs-cats/data" target="_blank">Dogs vs. Cats</a> dataset from Kaggle (ultimately, this dataset is provided by Microsoft Research).
#
# In previous Colabs, we've used <a href="https://www.tensorflow.org/datasets" target="_blank">TensorFlow Datasets</a>, which is a very easy and convenient way to use datasets. In this Colab however, we will make use of the class `tf.keras.preprocessing.image.ImageDataGenerator` which will read data from disk. We therefore need to directly download *Dogs vs. Cats* from a URL and unzip it to the Colab filesystem.
# + colab={} colab_type="code" id="OYmOylPlVrVt"
_URL = 'https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip'
zip_dir = tf.keras.utils.get_file('cats_and_dogs_filterted.zip', origin=_URL, extract=True)
# + [markdown] colab_type="text" id="Giv0wMQzVrVw"
# The dataset we have downloaded has following directory structure.
#
# <pre style="font-size: 10.0pt; font-family: Arial; line-height: 2; letter-spacing: 1.0pt;" >
# <b>cats_and_dogs_filtered</b>
# |__ <b>train</b>
# |______ <b>cats</b>: [cat.0.jpg, cat.1.jpg, cat.2.jpg ....]
# |______ <b>dogs</b>: [dog.0.jpg, dog.1.jpg, dog.2.jpg ...]
# |__ <b>validation</b>
# |______ <b>cats</b>: [cat.2000.jpg, cat.2001.jpg, cat.2002.jpg ....]
# |______ <b>dogs</b>: [dog.2000.jpg, dog.2001.jpg, dog.2002.jpg ...]
# </pre>
# + [markdown] colab_type="text" id="VpmywIlsVrVx"
# We'll now assign variables with the proper file path for the training and validation sets.
# + colab={} colab_type="code" id="sRucI3QqVrVy"
base_dir = os.path.join(os.path.dirname(zip_dir), 'cats_and_dogs_filtered')
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
# + colab={} colab_type="code" id="Utv3nryxVrV0"
train_cats_dir = os.path.join(train_dir, 'cats') # directory with our training cat pictures
train_dogs_dir = os.path.join(train_dir, 'dogs') # directory with our training dog pictures
validation_cats_dir = os.path.join(validation_dir, 'cats') # directory with our validation cat pictures
validation_dogs_dir = os.path.join(validation_dir, 'dogs') # directory with our validation dog pictures
# + [markdown] colab_type="text" id="ZdrHHTy2VrV3"
# ### Understanding our data
# + [markdown] colab_type="text" id="LblUYjl-VrV3"
# Let's look at how many cats and dogs images we have in our training and validation directory
# + colab={} colab_type="code" id="vc4u8e9hVrV4"
num_cats_tr = len(os.listdir(train_cats_dir))
num_dogs_tr = len(os.listdir(train_dogs_dir))
num_cats_val = len(os.listdir(validation_cats_dir))
num_dogs_val = len(os.listdir(validation_dogs_dir))
total_train = num_cats_tr + num_dogs_tr
total_val = num_cats_val + num_dogs_val
# + colab={} colab_type="code" id="g4GGzGt0VrV7"
print('total training cat images:', num_cats_tr)
print('total training dog images:', num_dogs_tr)
print('total validation cat images:', num_cats_val)
print('total validation dog images:', num_dogs_val)
print("--")
print("Total training images:", total_train)
print("Total validation images:", total_val)
# + [markdown] colab_type="text" id="tdsI_L-NVrV_"
# # Setting Model Parameters
# + [markdown] colab_type="text" id="8Lp-0ejxOtP1"
# For convenience, let us set up variables that will be used later while pre-processing our dataset and training our network.
# + colab={} colab_type="code" id="3NqNselLVrWA"
BATCH_SIZE = 100
IMG_SHAPE = 150 # Our training data consists of images with width of 150 pixels and height of 150 pixels
# + [markdown] colab_type="text" id="RLciCR_FVrWH"
# After defining our generators for training and validation images, **flow_from_directory** method will load images from the disk and will apply rescaling and will resize them into required dimensions using single line of code.
# + [markdown] colab_type="text" id="UOoVpxFwVrWy"
# # Data Augmentation
# + [markdown] colab_type="text" id="Wn_QLciWVrWy"
# Overfitting often occurs when we have a small number of training examples. One way to fix this problem is to augment our dataset so that it has sufficient number and variety of training examples. Data augmentation takes the approach of generating more training data from existing training samples, by augmenting the samples through random transformations that yield believable-looking images. The goal is that at training time, your model will never see the exact same picture twice. This exposes the model to more aspects of the data, allowing it to generalize better.
#
# In **tf.keras** we can implement this using the same **ImageDataGenerator** class we used before. We can simply pass different transformations we would want to our dataset as a form of arguments and it will take care of applying it to the dataset during our training process.
#
# To start off, let's define a function that can display an image, so we can see the type of augmentation that has been performed. Then, we'll look at specific augmentations that we'll use during training.
# + colab={} colab_type="code" id="GBYLOFgOXPJ9"
# This function will plot images in the form of a grid with 1 row and 5 columns where images are placed in each column.
def plotImages(images_arr):
fig, axes = plt.subplots(1, 5, figsize=(20,20))
axes = axes.flatten()
for img, ax in zip(images_arr, axes):
ax.imshow(img)
plt.tight_layout()
plt.show()
# + [markdown] colab_type="text" id="rlVj6VqaVrW0"
# ### Flipping the image horizontally
# + [markdown] colab_type="text" id="xcdvx4TVVrW1"
# We can begin by randomly applying horizontal flip augmentation to our dataset and seeing how individual images will look after the transformation. This is achieved by passing `horizontal_flip=True` as an argument to the `ImageDataGenerator` class.
# + colab={} colab_type="code" id="Bi1_vHyBVrW2"
image_gen = ImageDataGenerator(rescale=1./255, horizontal_flip=True)
train_data_gen = image_gen.flow_from_directory(batch_size=BATCH_SIZE,
directory=train_dir,
shuffle=True,
target_size=(IMG_SHAPE,IMG_SHAPE))
# + [markdown] colab_type="text" id="zJpRSxJ-VrW7"
# To see the transformation in action, let's take one sample image from our training set and repeat it five times. The augmentation will be randomly applied (or not) to each repetition.
# + colab={} colab_type="code" id="RrKGd_jjVrW7"
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
# + [markdown] colab_type="text" id="i7n9xcqCVrXB"
# ### Rotating the image
# + [markdown] colab_type="text" id="qXnwkzFuVrXB"
# The rotation augmentation will randomly rotate the image up to a specified number of degrees. Here, we'll set it to 45.
# + colab={} colab_type="code" id="1zip35pDVrXB"
image_gen = ImageDataGenerator(rescale=1./255, rotation_range=45)
train_data_gen = image_gen.flow_from_directory(batch_size=BATCH_SIZE,
directory=train_dir,
shuffle=True,
target_size=(IMG_SHAPE, IMG_SHAPE))
# + [markdown] colab_type="text" id="deaqZLsfcZ15"
# To see the transformation in action, let's once again take a sample image from our training set and repeat it. The augmentation will be randomly applied (or not) to each repetition.
# + colab={} colab_type="code" id="kVoWh4OIVrXD"
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
# + [markdown] colab_type="text" id="FOqGPL76VrXM"
# ### Applying Zoom
# + [markdown] colab_type="text" id="NvqXaD8BVrXN"
# We can also apply Zoom augmentation to our dataset, zooming images up to 50% randomly.
# + colab={} colab_type="code" id="tGNKLa_YVrXR"
image_gen = ImageDataGenerator(rescale=1./255, zoom_range=0.5)
train_data_gen = image_gen.flow_from_directory(batch_size=BATCH_SIZE,
directory=train_dir,
shuffle=True,
target_size=(IMG_SHAPE, IMG_SHAPE))
# + [markdown] colab_type="text" id="WgPWieSZcctO"
# One more time, take a sample image from our training set and repeat it. The augmentation will be randomly applied (or not) to each repetition.
# + colab={} colab_type="code" id="VOvTs32FVrXU"
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
# + [markdown] colab_type="text" id="usS13KCNVrXd"
# ### Putting it all together
# + [markdown] colab_type="text" id="OC8fIsalVrXd"
# We can apply all these augmentations, and even others, with just one line of code, by passing the augmentations as arguments with proper values.
#
# Here, we have applied rescale, rotation of 45 degrees, width shift, height shift, horizontal flip, and zoom augmentation to our training images.
# + colab={} colab_type="code" id="gnr2xujaVrXe"
image_gen_train = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
train_data_gen = image_gen_train.flow_from_directory(batch_size=BATCH_SIZE,
directory=train_dir,
shuffle=True,
target_size=(IMG_SHAPE,IMG_SHAPE),
class_mode='binary')
# + [markdown] colab_type="text" id="AW-pV5awVrXl"
# Let's visualize how a single image would look like five different times, when we pass these augmentations randomly to our dataset.
# + colab={} colab_type="code" id="z2m68eMhVrXm"
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
# + [markdown] colab_type="text" id="J8cUd7FXVrXq"
# ### Creating Validation Data generator
# + [markdown] colab_type="text" id="a99fDBt7VrXr"
# Generally, we only apply data augmentation to our training examples, since the original images should be representative of what our model needs to manage. So, in this case we are only rescaling our validation images and converting them into batches using ImageDataGenerator.
# + colab={} colab_type="code" id="54x0aNbKVrXr"
image_gen_val = ImageDataGenerator(rescale=1./255)
val_data_gen = image_gen_val.flow_from_directory(batch_size=BATCH_SIZE,
directory=validation_dir,
target_size=(IMG_SHAPE, IMG_SHAPE),
class_mode='binary')
# + [markdown] colab_type="text" id="b5Ej-HLGVrWZ"
# # Model Creation
# + [markdown] colab_type="text" id="wEgW4i18VrWZ"
# ## Define the model
#
# The model consists of four convolution blocks with a max pool layer in each of them.
#
# Before the final Dense layers, we're also applying a Dropout probability of 0.5. It means that 50% of the values coming into the Dropout layer will be set to zero. This helps to prevent overfitting.
#
# Then we have a fully connected layer with 512 units, with a `relu` activation function. The model will output class probabilities for two classes — dogs and cats — using `softmax`.
# + cellView="both" colab={} colab_type="code" id="Evjf8jZk2zi-"
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(150, 150, 3)),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Dropout(0.5),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dense(2)
])
# + [markdown] colab_type="text" id="DADWLqMSJcH3"
# ### Compiling the model
#
# As usual, we will use the `adam` optimizer. Since we output a softmax categorization, we'll use `sparse_categorical_crossentropy` as the loss function. We would also like to look at training and validation accuracy on each epoch as we train our network, so we are passing in the metrics argument.
# + colab={} colab_type="code" id="08rRJ0sn3Tb1"
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# + [markdown] colab_type="text" id="uurnCp_H4Hj9"
# ### Model Summary
#
# Let's look at all the layers of our network using **summary** method.
# + colab={} colab_type="code" id="b66qAJF_4Jnw"
model.summary()
# + [markdown] colab_type="text" id="N06iqE8VVrWj"
# ### Train the model
# + [markdown] colab_type="text" id="oub9RtoFVrWk"
# It's time we train our network.
#
# Since our batches are coming from a generator (`ImageDataGenerator`), we'll use `fit_generator` instead of `fit`.
# + colab={} colab_type="code" id="tk5NT1PW3j_P"
epochs=100
history = model.fit_generator(
train_data_gen,
steps_per_epoch=int(np.ceil(total_train / float(BATCH_SIZE))),
epochs=epochs,
validation_data=val_data_gen,
validation_steps=int(np.ceil(total_val / float(BATCH_SIZE)))
)
# + [markdown] colab_type="text" id="ojJNteAGVrWo"
# ### Visualizing results of the training
# + [markdown] colab_type="text" id="LZPYT-EmVrWo"
# We'll now visualize the results we get after training our network.
# + colab={} colab_type="code" id="8CfngybnFHQR"
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(epochs)
plt.figure(figsize=(8, 8))
plt.subplot(1, 2, 1)
plt.plot(epochs_range, acc, label='Training Accuracy')
plt.plot(epochs_range, val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(1, 2, 2)
plt.plot(epochs_range, loss, label='Training Loss')
plt.plot(epochs_range, val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.show()
| courses/udacity_intro_to_tensorflow_for_deep_learning/l05c02_dogs_vs_cats_with_augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.3
# language: julia
# name: julia-1.0
# ---
# # Homework 3: Due March 22 11:00PM
# ## Link to generate the homework repository: https://classroom.github.com/g/JHO5iO_C
# Make sure your code is well-commented.
# # Problem 1: Pseudo-spectral approximation
#
# Code up a function `project_cheb(f, n, lb, ub)` that takes in some **two-dimensional** function `f`, vector of degrees of approximation `n`, and vectors of lower bounds `lb` and upper bounds `ub`, and constructs a Chebyshev approximation on an grid of Chebyshev nodes.
#
# 1. Using collocation methods, approximate `sin(x + y)` on $[0,2\pi]\times[0,2\pi]$ using n=3, 5, and 10 degree approximations.
# 2. Plot your approximation for n=10.
# 3. Plot the relative error for n=5 in log10 units on an evenly spaced 100x100 grid in $[0,2\pi]\times[0,2\pi]$ where relative error is given by
# $$\text{relative error} = \log_{10}\left(\left|\frac{\hat{\sin}(x+y) - \sin(x+y)}{\sin(x+y)}\right|\right)$$
# where the hat indicates your approximation.
#
# log10 units often makes the plots more readable since errors can change size rapidly.
# # Problem 2: Complete Chebyshev basis
#
# Code up a function `project_lin_spline(f, knots)` that takes in some **two-dimensional** function `f`, and an $n\times2$ grid of knots `knots`, and constructs a linear spline approximation on the grid.
#
# 1. Using finite-element methods, approximate `sin(x + y)` on $[0,2\pi]\times[0,2\pi]$ using n=3, 5, and 10 knots on each dimension (9, 25, 100 total).
# 2. Plot your approximation for n=10.
# 3. Plot the relative error for n=5 in log10 units on an evenly spaced 100x100 grid in $[0,2\pi]\times[0,2\pi]$.
| problem-sets/3_ps/3_ps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 ('xaod-usage-w68Kx7k0-py3.9')
# language: python
# name: python3
# ---
# # Advanced, Common, Features
#
# The ATLAS data model's primary objects, like the `Jet` or `Electron` (or any others) have some features in common. This chapter talks about access an objet's decorations (and please let us know if there are others that should be described).
from config import ds_bphys as ds
import matplotlib.pyplot as plt
import awkward as ak
from func_adl_servicex_xaodr21 import cpp_float, cpp_string
# ## Object Decorations
#
# The XAOD data model is designed to be extensible. Another object member can be added at anytime, even if the object doesn't have an accessor method. In fact, almost all data in the data model is stored as a named column: when you access `jet.pt()` the `pt` method actually does a lookup on a decorator with the name `pt` associated with the `jet` object.
#
# This feature is used extensively in reconstruction and calibration processing. Since decorators can be almost any type of access, in `func_adl` you have to used parameterized access to get at them. Here is an example from a b-physics derived dataset. We'll demo this by translating a snippet of *b*-physics code provided by <NAME>.
#
# If you've not worked with derived BPHYS datasets, they are unique-ish in the ATLAS world. Since statistics are the name of the game, almost everything is stripped out. Here we want to go after the `QUAD_mass` decoration on the `BPHY4Quads` collection in the sample. First job is to figure out what the `BPHY4Quads` collection is. The ATLAS `checkxAOD.py` tool is very useful here. If you do that, you'll find the following:
#
# ```bash
# [bash][atlas AnalysisBase-21.2.62]:data > checkxAOD.py DAOD_BPHY4.999031._000001.pool.root.1
# ======================================================================================================================== File: DAOD_BPHY4.999031._000001.pool.root.1
# ------------------------------------------------------------------------------------------------------------------------ Memory size Disk Size Size/Event Compression Entries Name (Type)
# ------------------------------------------------------------------------------------------------------------------------ 333.24 kB 50.19 kB 0.00 kB/event 6.64 20788 TrigConfKeys (xAOD::TrigConfKeys_v1) [Trig]
# 2544.13 kB 516.87 kB 0.02 kB/event 4.92 20788 GSFConversionVertices (DataVector<xAOD::Vertex_v1>) [egamma]
# 4089.23 kB 1132.31 kB 0.05 kB/event 3.61 20788 LVL1MuonRoIs (DataVector<xAOD::MuonRoI_v1>) [Trig]
# 6630.11 kB 1189.60 kB 0.06 kB/event 5.57 20788 InDetForwardTrackParticles (DataVector<xAOD::TrackParticle_v1>) [InDet]
# 12605.41 kB 1757.67 kB 0.08 kB/event 7.17 20788 EventInfo (xAOD::EventInfo_v1) [EvtId]
# 7545.86 kB 2787.27 kB 0.13 kB/event 2.71 20788 HLT_xAOD__MuonContainer_MuonEFInfo_FullScan (DataVector<xAOD::Muon_v1>) [Trig]
# 23009.07 kB 2875.53 kB 0.14 kB/event 8.00 20788 McEventInfo (PileUpEventInfo_p5) [EvtId]
# 14812.24 kB 3296.27 kB 0.16 kB/event 4.49 20788 HLT_xAOD__TrigBphysContainer_EFTrackMass (DataVector<xAOD::TrigBphys_v1>) [Trig]
# 21498.52 kB 4903.93 kB 0.24 kB/event 4.38 20788 HLT_xAOD__MuonContainer_MuonEFInfo (DataVector<xAOD::Muon_v1>) [Trig]
# 61415.48 kB 4976.95 kB 0.24 kB/event 12.34 20788 HLT_xAOD__TrigBphysContainer_EFBMuMuXFex (DataVector<xAOD::TrigBphys_v1>) [Trig]
# 217772.43 kB 5590.42 kB 0.27 kB/event 38.95 20788 xTrigDecision (xAOD::TrigDecision_v1) [Trig]
# 107825.61 kB 7299.25 kB 0.35 kB/event 14.77 20788 HLT_xAOD__TrigBphysContainer_EFMultiMuFex (DataVector<xAOD::TrigBphys_v1>) [Trig]
# 26836.23 kB 8364.40 kB 0.40 kB/event 3.21 20788 Electrons (DataVector<xAOD::Electron_v1>) [egamma]
# 104940.43 kB 8684.58 kB 0.42 kB/event 12.08 20788 HLT_xAOD__TrigBphysContainer_L2BMuMuFex (DataVector<xAOD::TrigBphys_v1>) [Trig]
# 14616.80 kB 8693.32 kB 0.42 kB/event 1.68 20788 MuonSpectrometerTrackParticles (DataVector<xAOD::TrackParticle_v1>) [Muon]
# 20589.57 kB 11065.87 kB 0.53 kB/event 1.86 20788 GSFTrackParticles (DataVector<xAOD::TrackParticle_v1>) [egamma]
# 38546.04 kB 11397.73 kB 0.55 kB/event 3.38 20788 Photons (DataVector<xAOD::Photon_v1>) [egamma]
# 23701.12 kB 13832.01 kB 0.67 kB/event 1.71 20788 HLT_xAOD__TrackParticleContainer_InDetTrigTrackingxAODCnv_Muon_IDTrig (DataVector<xAOD::TrackParticle_v1>) [Trig]
# 453682.19 kB 22308.57 kB 1.07 kB/event 20.34 20788 HLT_xAOD__TrigBphysContainer_EFBMuMuFex (DataVector<xAOD::TrigBphys_v1>) [Trig]
# 521885.12 kB 24422.69 kB 1.17 kB/event 21.37 20788 BPHY4RefittedPrimaryVertices (DataVector<xAOD::Vertex_v1>) [*Unknown*]
# 71110.69 kB 28750.34 kB 1.38 kB/event 2.47 20788 ExtrapolatedMuonTrackParticles (DataVector<xAOD::TrackParticle_v1>) [Muon]
# 72210.54 kB 31226.66 kB 1.50 kB/event 2.31 20788 CombinedMuonTrackParticles (DataVector<xAOD::TrackParticle_v1>) [Muon]
# 66508.79 kB 35456.61 kB 1.71 kB/event 1.88 20788 egammaClusters (DataVector<xAOD::CaloCluster_v1>) [egamma]
# 142288.23 kB 45957.38 kB 2.21 kB/event 3.10 20788 Muons (DataVector<xAOD::Muon_v1>) [Muon]
# 322969.71 kB 56110.05 kB 2.70 kB/event 5.76 20788 PrimaryVertices (DataVector<xAOD::Vertex_v1>) [InDet]
# 99516.93 kB 59326.76 kB 2.85 kB/event 1.68 20788 BPHY4Quads (DataVector<xAOD::Vertex_v1>) [*Unknown*]
# 110682.30 kB 62286.51 kB 3.00 kB/event 1.78 20788 BPHY4Pairs (DataVector<xAOD::Vertex_v1>) [*Unknown*]
# 421894.21 kB 117419.71 kB 5.65 kB/event 3.59 20788 InDetTrackParticles (DataVector<xAOD::TrackParticle_v1>) [InDet]
# 1064386.64 kB 292559.83 kB 14.07 kB/event 3.64 20788 TrigNavigation (xAOD::TrigNavigation_v1) [Trig]
# ------------------------------------------------------------------------------------------------------------------------ 4056446.90 kB 874239.27 kB 42.05 kB/event Total
# ```
#
# So we know we can access it as a vertex!
quad_mass = (
ds.Select(lambda e: e.Vertices('BPHY4Quads'))
.Select(lambda quads: {
'mass': [q.auxdataConst[cpp_float]('QUAD_mass') for q in quads],
'chi2': [q.chiSquared() for q in quads],
'charge_code': [q.auxdataConst[cpp_string]('ChargeCode') for q in quads],
})
.AsAwkwardArray()
.value()
)
# And a two panel plot showing the difference in mass and fit $\chi^2$ for the two types of reconstructed vertices:
# +
fig, [ax1, ax2] = plt.subplots(1, 2, sharey=True, figsize=(10, 5))
two_plus_charges = ak.sum(ak.without_parameters(quad_mass.charge_code) == ord('+'), axis=1) == 2
ax1.hist(ak.flatten(quad_mass['mass']), bins=100, range=(0, 2e6), label='All')
ax1.hist(ak.flatten(quad_mass[two_plus_charges]['mass']), bins=100, range=(0, 2e6), label='++')
ax1.set_yscale('log')
ax1.set_xlabel('Mass [eV]')
ax1.legend()
ax1.set_title('QUAD Mass')
ax2.hist(ak.flatten(quad_mass['chi2']), bins=100, range=(0,100), label='All')
ax2.hist(ak.flatten(quad_mass[two_plus_charges]['chi2']), bins=100, range=(0, 100), label='++')
ax2.set_xlabel('$\chi^2$')
ax2.legend()
_ = ax2.set_title('Fit $\chi^2$')
| book/chapters/xaod_objects.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
score = np.random.randint(40, 100, (10, 5))
score_df = pd.DataFrame(score)
score_df
subjects = ["语文", "数学", "英语", "政治", "体育"]
stu = ['同学' + str(i) for i in range(score_df.shape[0])]
data = pd.DataFrame(score, columns=subjects, index=stu)
data
data.shape
data.index
data.columns
data.values
data.T
data.head(5)
data.tail(5)
stu = ["学生_" + str(i) for i in range(score_df.shape[0])]
data.index = stu
data
data.reset_index(drop=True)
| deeplearningDemo/hello.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
Utility to fetch mu, sigma and proba
__author__ : <NAME>
"""
import numpy as np
VERBOSE = True
class gaussian_params:
def __init__ (self, mu, sigma):
self.mu = np.array(mu).reshape((1,-1))
self.sigma = np.array(sigma)
if VERBOSE:
print ("mu = ", self.mu)
print ("sigma = ", self.sigma)
def proba(self, x):
t1 = (1./np.sqrt(np.linalg.det(self.sigma)))
t2 = np.exp(-(x - self.mu).dot(np.linalg.pinv(self.sigma)).dot((x-self.mu).T)/2.)
prob = t1 * t2
return prob.diagonal()
def _var(self, sigma):
delta = abs(sigma - self.sigma)
self.sigma = sigma
return np.sqrt(e.mean())/2
def _mu(self, mu):
delta = abs(self, mu)
self.mu = mu
return delta.mean()/2
# -
| _Templates/Utility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img style="float: right;" src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAAOIAAAAjCAYAAACJpNbGAAAABHNCSVQ<KEY>zAAALEgAACxIB0t1+/AAAABR0RVh0Q3JlYXRpb24gVGltZQAzLzcvMTNND4u/AAAAHHRFWHRTb2Z0d2FyZQBBZG9iZSBGaXJld29ya3MgQ1M26LyyjAAACMFJREFUeJztnD1y20gWgD+6nJtzAsPhRqKL3AwqwQdYDpXDZfoEppNNTaWbmD7BUEXmI3EPMFCR2YI1UDQpdAPqBNzgvRZA/BGUZEnk9FeFIgj0z2ugX7/XP+jGer2mLv/8b6d+4Efgf/8KG0+Zn8XyXLx+bgEslqegcfzxSY3Irrx6bgEsFssBWsRGowGufwHAYtq7u+H6fUCOxTTWax4wBAbr+SRqNDKesOv3gN/133sW0yh927j1mucIaFWINl7PJ+OcvMcfW8Bol3iN44+mLIOsTCp3UJFfAETr+WRQcG8EOJpunEnTyDlYzycbeWr5xxq3jOF6PglK8ix9buv5xCsrAzBkMV1l5OwD/aJ4BXzV3+8F9z4gz/hTSbz8cxc84FuNvDc4VIsYA7+qohmGwAnycA194G22YqUYlZxv4vpN4AuwBv4oON5m8k3TVLnK4sYFcRyN86dWvCwnlCvFCeUVvwX8CkSZZ5eWs5mLJWE/VZThBMgpfirPk5J4f1SU4QsQ6LNP4+j9OkSUKdRiGlD87CWe3PcyR5PFdAhc1cz/joOziMoIeVF95GX1EGVY6bWhvsAeZQrm+kON80PDneD6PRbTi4LQpmJfsZieFaR1qXlXURh3y2BaBPyG63sspv0t6e+CKJTrf2YxHe8Qr6z8AXBdGbMoHgCTshgr4AiItfxljenPJGv5roCi+rGVw1TExTTWl99ThRsglfYHUnF7SMv+Bhjn4idxbhFLGiAu6gjXD3LuUBF5VzWi3CoAfMP1kxe7mNYZMT5DLFgf13eAXi3ZtvMOsUb3V3J5/mmqy+/66RbnTC1LFdfIu/kd8Qx2bTQeg2GBTPfiUF1TgHNE0QaIq/JDX9RKr/WBy/V8EhfEHWncWMO2EKV8S7UypYnYdE2r+o8gyj5MHXVYsZh+JnG7A+3LPQxR5g9II/UJ148ockmrybqm2+Qapo6gppwB8J7EM6jqaz8u0lhfkXgB58BKPam6rvEdh2kRARbTMa7/HXEfVqnW8hxxWwE+5+JJRTYd9CM90gxw/XFuMKMo/yTNDzUkLnbr6rCYnuH6N8igQ3CvNPJproDPuH6MKMd4Z5kMUjnrh98tn1if72/Ie729Vzq708L0YV3/HGmgB4iHsjOProhhd1lrEr4zaz/FvM4lolTnqWum/6jKmeuDmFb1jHylNg96hPQbhcU0wPVBXESvQI4W5aNshsK4jeOPhSOcOaThMVb48dhU8m2UlR+29ZHzrqyhLL0EaTROteGt67EYIsT6F1HXC/ikcvS00dl51PRwLaIwQtzCxGWRFnRMkT8v/SyAy8I+iliHJtDUsHHq7imipE42GtJanxdcB6mgQcm9MmKNs1m5F9MI13+n+cXZSEpAeV8mQgZqNkmU/HsuT7kf4PrGhXcK0h1SXv7iPKsJKCrDYvoV17+meMqhiDFlll7GEb4U3iseAf+k7mqksmU9qUoaj73E7TEtol3iZnks7Moai8WylUN3TS0WANbzyYv2rqxFtFheANYi7iGNRoPOrO2QGTQIu8vhU8vSmbWNDAHQD7vLYWfWbgFx2F3ee3FBZ9ZuIgMpTWAQdpeRXm9pPoPOrD3UMCtkQM4BRmF3ubG6ZZdxkOfCWsT9pU96CuX56KfOjeIFVC8Ar8NI0xuyOQJsVkWl8xzptQGPNY/6xFiLuL+0gIu0FVTrNESmbK7C7tLrzNpmPW0EeGF32UyFN19UnCAT4ZHGWWnYqDNrB4jViZBK/kbD9sLuMiBZSD8AVp1Z+0LD/NmZta+BIzOS3pm1xwBhd9kvkeEGUbQeqSmIdHhkXnGs5fIQRUxPV1x0Zm2zMuoq7C69rU/yBWAt4v7iAd86s/ZaDweZP+wBvwBOZ9b2SCrrmPzk+AWizA09j1QxMK4gZumcWKUWMvkdA56mfxN2l7GmHWk6V2F32Qi7yxaIsmnYHvkJ9zEQqAwBotQXwK2m0c+EN/Kk8zPTZiOkIWrp/xNTnpeOtYh7iFauN+k5W+0vXab6UsbyecAw229SxWiG3aVZ7NBCKrGHuneazy2iyBeIuxkjk9UDE1bzOtJ4IzbdwysNN0D6dnf9Rk3/iKSBWOnhUbASSWW+DbvLWM+HKreZ3O/r77gza5u842w6LxFrEfcTj+Jv3mK4q7Co63hE+fI6E94hUaT0cry+XushSuvoNZO2CdsCrlXJHDYVMUIUJso2BmhfL+wuV6rMvVR6AXnS1428XupaE7Hwnrqkg4cMGD0lr3NfpVegrUw1m2sN0+crNirEX1uTqiPbPoyI/QSKKmqA9I9aer+fcR2zxIj7GiMV+EYVIkZc3r5eH2rYI+0vnpBYIE/vGwUCdYM7s3agbqXJu58VIOwug86sfd2ZtSPNKwi7S9PHy4UnscCmXKuUZQRdsqbPwCHp2754pKYnW0akcZBO/x2df29XnvA//6iV8T3TSluBmOQlR+v5JNvaHixlDZRalRZifbZaAg3vIIrkmP6YVu6owI1M9x2r0vVIFCBGXNLS96Ph45IGY2ey6e1DY20UMaLGItUXoIhVvCv5tvDg2MWLqYNaoKBKWe6Z7gBR8OwAzZOyD4poBmtidlwt/gIxw/QHz0+oWKIoj19fRz8p3YOjoV8195F5l31ltZ5PfnluISyW+/IK6SPstRIiH/FaLHvLa2R+6F6f978AVsD7v0vf0HK4vNK9VfbVojSBceP4o/PcglgsD8GMmjaRbRCc1PEQIrbv45nlIfleIrs778XkrcWSZXMcXPZyqbvfxy7ckuyqHJPslJzH9c3We2ZRbx1O/07ziJbDI1FE2Qwp4n4DNzHJhkZF16+3bnwrCmi40U2eWoj7KZvobn7+YtKO1vPJVyyWPSZrER1kNU0TqfienpvlaWZR7oX+3tba6lxcX7MK3tNfo2RlpNc8tthsIFbAKYtpsA+TtRbLNp5/H4/EFXX0MOfbOGUxvbCKaDkEnl8Rq0jc1ayFjhFFjKwiWg6B/wNk+JCXXNBIXQAAAABJRU5ErkJggg==">
#
#
# # Running PCSE/WOFOST with custom input data (1)
#
# This Jupyter notebook will show you how to read inputs from files for running PCSE/WOFOST.
#
# <NAME>, March 2018
#
# **Prerequisites for running this notebook**
#
# Several packages need to be installed for running PCSE/WOFOST:
#
# 1. `PCSE` and its dependencies. See the [PCSE user guide](http://pcse.readthedocs.io/en/stable/installing.html) for more information;
# 2. The `pandas` module for processing and storing WOFOST output;
# 3. The `matplotlib` module for generating charts
#
#
# ## Introduction
#
# For running PCSE/WOFOST (and PCSE models in general) with your own data sources you need three different types of inputs:
#
# 1. Model parameters that parameterize the different model components. These parameters usually consist of a set of crop parameters (or multiple sets in case of crop rotations), a set of soil parameters and a set of site parameters. The latter provide ancillary parameters that are specific for a location, for example the initial amount of moisture in the soil.
# 2. Driving variables represented by weather data which can be derived from various sources.
# 3. Agromanagement actions which specify the farm activities that will take place on the field that is simulated by PCSE.
#
# For this example we will run a simulation for sugar beet in Wageningen (Netherlands) and we will read the input data step by step from several different sources instead of using the pre-configured start_wofost() script. For the example we will assume that data files are in the `data` directory within the directory where this notebook is located. This will be the case if you downloaded the notebooks from github.
#
# ## Importing the relevant modules
#
# +
# %matplotlib inline
import sys, os
import matplotlib
matplotlib.style.use("ggplot")
import matplotlib.pyplot as plt
import pandas as pd
data_dir = os.path.join(os.getcwd(), "data")
import pcse
print("This notebook was built with:")
print("python version: %s " % sys.version)
print("PCSE version: %s" % pcse.__version__)
# -
# ## Reading model parameters
# ### Crop parameters
#
# The crop parameters consist of parameter names and the corresponding parameter values that are needed to parameterize the components of the crop simulation model. These are crop-specific values regarding phenology, assimilation, respiration, biomass partitioning, etc. The parameter file for sugar beet is taken from the crop files in the WOFOST Control Centre.
#
# As many crop models in Wageningen were written in FORTRAN, the crop parameters for many models in Wageningen are often provided in the CABO format that could be read with the TTUTIL FORTRAN library. This CABO format will be gradually phased out and PCSE will move to a new format based on YAML, see [here for an example](https://github.com/ajwdewit/WOFOST_crop_parameters/blob/master/wheat.yaml). However, PCSE tries to be backward compatible as much as possible and provides the `CABOFileReader` for reading parameter files in CABO format. The `CABOFileReader` returns a dictionary with the parameter name/value pairs:
from pcse.fileinput import CABOFileReader
cropfile = os.path.join(data_dir, 'crop', 'sug0601.crop')
cropdata = CABOFileReader(cropfile)
# ### Soil parameters
# The soildata dictionary provides the parameter name/value pairs related to the soil type and soil physical properties. The number of parameters is variable depending on the soil water balance type that is used for the simulation. For this example, we will use the water balance for freely draining soils and use the soil file for medium fine sand: `ec3.soil`. This file is also taken from the soil files in the [WOFOST Control Centre](http://www.wageningenur.nl/wofost).
soilfile = os.path.join(data_dir, 'soil', 'ec3.soil')
soildata = CABOFileReader(soilfile)
# ### Site parameters
#
# The site parameters provide ancillary parameters that are not related to the crop or the soil. Examples are the initial conditions of the water balance such as the initial soil moisture content (WAV) and the initial and maximum surface storage (SSI, SSMAX). Also the atmospheric $CO_{2}$
# concentration is a typical site parameter. For the moment, we can define these parameters directly on the Python commandline as a simple python dictionary. However, it is more convenient to use the `WOFOST71SiteDataProvider` that documents the site parameters and provides sensible defaults:
from pcse.util import WOFOST71SiteDataProvider
sitedata = WOFOST71SiteDataProvider(WAV=100, CO2=360)
print(sitedata)
# ### Packaging all parameters
# Finally, we need to pack the different sets of parameters into one variable using the `ParameterProvider`. This is needed because PCSE expects one variable that contains all parameter values. Using this approach has the additional advantage that parameter value can be easily overridden in case of running multiple simulations with slightly different parameter values:
from pcse.base import ParameterProvider
parameters = ParameterProvider(cropdata=cropdata, soildata=soildata, sitedata=sitedata)
# ## Agromanagement
# The agromanagement inputs provide the start date of the agricultural campaign, the start_date/start_type of the crop simulation, the end_date/end_type of the crop simulation and the maximum duration of the crop simulation. The latter is included to avoid unrealistically long simulations for example as a results of a too high temperature sum requirement.
#
# The agromanagement inputs are defined with a special syntax called [YAML](http://yaml.org/) which allows to easily create more complex structures which is needed for defining the agromanagement. The agromanagement file for sugar beet in Wageningen `sugarbeet_calendar.agro` can be read with the `YAMLAgroManagementReader`:
from pcse.fileinput import YAMLAgroManagementReader
agromanagement_file = os.path.join(data_dir, 'agro', 'sugarbeet_calendar.agro')
agromanagement = YAMLAgroManagementReader(agromanagement_file)
print(agromanagement)
# ## Daily weather observations
# Daily weather variables are needed for running the simulation. There are several data providers in PCSE for reading weather data, see the section on [weather data providers](http://pcse.readthedocs.io/en/stable/reference_guide.html#weather-data-providers) to get an overview.
#
# For this example we will use weather data from an excel file which provides daily weather data for Wageningen for the period 2004 to 2008. We will read the data from the file using the ExcelWeatherDataProvider:
from pcse.fileinput import ExcelWeatherDataProvider
weatherfile = os.path.join(data_dir, 'meteo', 'nl1.xlsx')
wdp = ExcelWeatherDataProvider(weatherfile)
print(wdp)
# ## Importing, initializing and running a PCSE model
#
# Internally, PCSE uses a simulation engine to run a crop simulation. This engine takes a configuration file that specifies the components for the crop, the soil and the agromanagement that need to be used for the simulation. So any PCSE model can be started by importing the engine and initializing it with a given configuration file and the corresponding parameters, weather data and agromanagement.
#
# However, as many users of PCSE only need a particular configuration (for example the WOFOST model for potential production), preconfigured Engines are provided in `pcse.models`. For the sugarbeet example we will import the WOFOST model for water-limited simulation under freely draining soil conditions:
from pcse.models import Wofost71_WLP_FD
wofsim = Wofost71_WLP_FD(parameters, wdp, agromanagement)
# We can then run the simulation and retrieve the time series of daily simulation output using the get_output() method on the WOFOST object. Finally, we convert the simulation reults to a pandas dataframe:
wofsim.run_till_terminate()
df_results = pd.DataFrame(wofsim.get_output())
df_results = df_results.set_index("day")
df_results.tail()
# ## Visualizing simulation results
# Finally, we can generate some figures of WOFOST variables such as the development (DVS), total biomass (TAGP), leaf area index (LAI) and root-zone soil moisture (SM) using the MatPlotLib plotting package:
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(12,10))
for var, ax in zip(["DVS", "TAGP", "LAI", "SM"], axes.flatten()):
ax.plot_date(df_results.index, df_results[var], 'b-')
ax.set_title(var)
fig.autofmt_xdate()
| 02 Running with custom input data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SKLearn Spacy Reddit Text Classification Example
#
# In this example we will be buiding a text classifier using the reddit content moderation dataset.
#
# For this, we will be using SpaCy for the word tokenization and lemmatization.
#
# The classification will be done with a Logistic Regression binary classifier.
#
# The steps in this tutorial include:
#
# 1) Train and build your NLP model
#
# 2) Build your containerized model
#
# 3) Test your model as a docker container
#
# 4) Run Seldon in your kubernetes cluster
#
# 5) Deploy your model with Seldon
#
# 6) Interact with your model through API
#
# 7) Clean your environment
#
#
# ### Before you start
# Make sure you install the following dependencies, as they are critical for this example to work:
#
# * Helm v3.0.0+
# * A Kubernetes cluster running v1.13 or above (minkube / docker-for-windows work well if enough RAM)
# * kubectl v1.14+
# * Python 3.6+
# * Python DEV requirements (we'll install them below)
#
# Let's get started! 🚀🔥
#
# ## 1) Train and build your NLP model
# Let's first install any dependencies
# !pip install -r requirements.txt
# +
import pandas as pd
from sklearn.model_selection import train_test_split
import numpy as np
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import LogisticRegression
from seldon_core.seldon_client import SeldonClient
import dill
import sys, os
# This import may take a while as it will download the Spacy ENGLISH model
from ml_utils import CleanTextTransformer, SpacyTokenTransformer
# +
df_cols = ["prev_idx", "parent_idx", "body", "removed"]
TEXT_COLUMN = "body"
CLEAN_COLUMN = "clean_body"
TOKEN_COLUMN = "token_body"
# Downloading the 50k reddit dataset of moderated comments
df = pd.read_csv("https://raw.githubusercontent.com/axsauze/reddit-classification-exploration/master/data/reddit_train.csv",
names=df_cols, skiprows=1, encoding="ISO-8859-1")
df.head()
# -
# Let's see how many examples we have of each class
df["removed"].value_counts().plot.bar()
x = df["body"].values
y = df["removed"].values
x_train, x_test, y_train, y_test = train_test_split(
x, y,
stratify=y,
random_state=42,
test_size=0.1, shuffle=True)
# Clean the text
clean_text_transformer = CleanTextTransformer()
x_train_clean = clean_text_transformer.transform(x_train)
# Tokenize the text and get the lemmas
spacy_tokenizer = SpacyTokenTransformer()
x_train_tokenized = spacy_tokenizer.transform(x_train_clean)
# +
# Build tfidf vectorizer
tfidf_vectorizer = TfidfVectorizer(
max_features=10000,
preprocessor=lambda x: x,
tokenizer=lambda x: x,
token_pattern=None,
ngram_range=(1, 3))
tfidf_vectorizer.fit(x_train_tokenized)
# -
# Transform our tokens to tfidf vectors
x_train_tfidf = tfidf_vectorizer.transform(
x_train_tokenized)
# Train logistic regression classifier
lr = LogisticRegression(C=0.1, solver='sag')
lr.fit(x_train_tfidf, y_train)
# These are the models we'll deploy
with open('tfidf_vectorizer.model', 'wb') as model_file:
dill.dump(tfidf_vectorizer, model_file)
with open('lr.model', 'wb') as model_file:
dill.dump(lr, model_file)
# ## 2) Build your containerized model
# This is the class we will use to deploy
# !cat RedditClassifier.py
# test that our model works
from RedditClassifier import RedditClassifier
# With one sample
sample = x_test[0:1]
print(sample)
print(RedditClassifier().predict(sample, ["feature_name"]))
# ### Create Docker Image with the S2i utility
# Using the S2I command line interface we wrap our current model to seve it through the Seldon interface
# To create a docker image we need to create the .s2i folder configuration as below:
# !cat .s2i/environment
# As well as a requirements.txt file with all the relevant dependencies
# !cat requirements.txt
# !s2i build . seldonio/seldon-core-s2i-python3:0.13 reddit-classifier:0.1
# ## 3) Test your model as a docker container
# Remove previously deployed containers for this model
# !docker rm -f reddit_predictor
# !docker run --name "reddit_predictor" -d --rm -p 5001:5000 reddit-classifier:0.1
# ### Make sure you wait for language model
# SpaCy will download the English language model, so you have to make sure the container finishes downloading it before it can be used. You can view this by running the logs until you see "Linking successful".
# Here we need to wait until we see "Linking successful", as it's downloading the Spacy English model
# You can hit stop when this happens
# !docker logs -t -f reddit_predictor
# +
# We now test the REST endpoint expecting the same result
endpoint = "0.0.0.0:5001"
batch = sample
payload_type = "ndarray"
sc = SeldonClient(microservice_endpoint=endpoint)
response = sc.microservice(
data=batch,
method="predict",
payload_type=payload_type,
names=["tfidf"])
print(response)
# -
# We now stop it to run it in docker
# !docker stop reddit_predictor
# ## 4) Run Seldon in your kubernetes cluster
#
# ## Setup Seldon Core
#
# Use the setup notebook to [Setup Cluster](../../seldon_core_setup.ipynb#Setup-Cluster) with [Ambassador Ingress](../../seldon_core_setup.ipynb#Ambassador) and [Install Seldon Core](../../seldon_core_setup.ipynb#Install-Seldon-Core). Instructions [also online](./seldon_core_setup.html).
# ## 5) Deploy your model with Seldon
# We can now deploy our model by using the Seldon graph definition:
# We'll use our seldon deployment file
# !cat reddit_clf.json
# !kubectl apply -f reddit_clf.json
# !kubectl get pods
# ## 6) Interact with your model through API
# Now that our Seldon Deployment is live, we are able to interact with it through its API.
#
# There are two options in which we can interact with our new model. These are:
#
# a) Using CURL from the CLI (or another rest client like Postman)
#
# b) Using the Python SeldonClient
#
# #### a) Using CURL from the CLI
# + language="bash"
# curl -X POST -H 'Content-Type: application/json' \
# -d "{'data': {'names': ['text'], 'ndarray': ['Hello world this is a test']}}" \
# http://127.0.0.1/seldon/default/reddit-classifier/api/v0.1/predictions
# -
# #### b) Using the Python SeldonClient
# +
from seldon_core.seldon_client import SeldonClient
import numpy as np
host = "localhost"
port = "80" # Make sure you use the port above
batch = np.array(["Hello world this is a test"])
payload_type = "ndarray"
deployment_name="reddit-classifier"
transport="rest"
namespace="default"
sc = SeldonClient(
gateway="ambassador",
ambassador_endpoint=host + ":" + port,
namespace=namespace)
client_prediction = sc.predict(
data=batch,
deployment_name=deployment_name,
names=["text"],
payload_type=payload_type,
transport="rest")
print(client_prediction)
# -
# ## 7) Clean your environment
# !kubectl delete -f reddit_clf.json
# !helm del ambassador
# !helm del seldon-core-operator
| examples/models/sklearn_spacy_text/sklearn_spacy_text_classifier_example.ipynb |