code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="JDfSG9_9dZ9M"
# # Programa principal
# + [markdown] id="VZdlIZ4-d64x"
# ## Imports y variables globales
# + id="iV0Xev8caIOJ"
import pandas as pd
import numpy as np
import tensorflow as tf
import sklearn
from sklearn import preprocessing
from sklearn import compose
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LinearRegression, Ridge, Lasso, ElasticNet
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import mean_squared_error, mean_absolute_error, r2_score
from matplotlib import pyplot as plt
data_url = "https://raw.githubusercontent.com/BoscoDomingo/ML-P1-Regression/main/data/"
csv_filenames = ["audi.csv","bmw.csv", "cclass.csv", "focus.csv", "ford.csv" , "hyundi.csv", "merc.csv", "skoda.csv", "toyota.csv", "vauxhall.csv", "vw.csv"]
# There is no elegant solution to this yet. Google and Microsoft, pls.
random_state = 1337
# + [markdown] id="8-1ZpKr82Ar0"
# ## Lectura de datos
# + id="XPzPqyGSeBPe"
df = pd.DataFrame(columns=["brand"])
for file in csv_filenames:
partial_df = pd.read_csv(f"{data_url}{file}")
# Fix "tax" column
if "tax(£)" in partial_df:
partial_df.rename(columns={"tax(£)":"tax"}, inplace=True)
partial_df["brand"] = file.split(".")[0]
df = pd.concat([df, partial_df])
# Ensure correct data type
df.infer_objects()
df.year = pd.to_numeric(df.year, downcast='unsigned')
# + [markdown] id="H80JXiVo17g2"
# ## Análisis de Datos y Preprocesamiento
# Analizamos el DataFrame resultante de unificar los datos
# + id="xjeDQLz0IKnp"
df_backup = df.copy() # Backup so as not to have to re-read CSVs all the time
# Price goes first for easier data treatment later on
df = df[['price', 'brand', 'model', 'year', 'transmission', 'mileage',
'fuelType', 'tax', 'mpg', 'engineSize']]
# + id="guyw2TVkizTF" outputId="e3eabd0c-cc06-4215-e6e1-a5660b9a34d5" colab={"base_uri": "https://localhost:8080/", "height": 300}
df_backup.describe()
# + id="fOv5bHVEmFow" outputId="05919b05-d533-48f7-a782-4258023ff850" colab={"base_uri": "https://localhost:8080/", "height": 363}
df.head(10)
# + id="whV-cZ05UoSP" outputId="84c2e395-939a-4251-90ce-275cfd84fa38" colab={"base_uri": "https://localhost:8080/", "height": 1000}
for column in df_backup.columns:
plt.figure()
plt.xlabel(column)
plt.hist(df_backup[column])
# + id="3fzCfi3eUt0r" outputId="9755b897-48aa-430b-842e-f2b11de1f98b" colab={"base_uri": "https://localhost:8080/", "height": 677}
index = 1
plt.figure(figsize=(14,10))
plt.suptitle('Price scatter plots v1', size = 28);
for column in df_backup.columns:
if column == "price":
continue
plt.subplot(3,3,index)
plt.scatter(df_backup[column], df_backup['price'])
plt.xlabel(column)
index += 1
# Source: https://datascience.stackexchange.com/a/31755
# + [markdown] id="Ne_2sJFXpywG"
# #### Anomalías observadas
#
# * Las columnas `tax` y `tax(£)` (que unificamos en el proceso de lectura)
# * `year` superiores a 2020 (imposible) y un par de valores muy bajos, cuando la gran mayoría son del 2000 en adelante
# * `engineSize` con valor 0 (imposible)
# * `mpg` con valores imposibles (menores a 11 MPG o superiores a 94 MPG)
# * `mileage` con valores extremadamente altos y bajos
#
#
# + [markdown] id="_m0jP5LiiXVp"
# #### Filtrado inicial
#
#
# 1. Eliminar filas con valores en `year` superiores a 2020 (que son imposibles) y eliminar los valores menores a 2000 (que son un par de unidades y pueden causar problemas)
# 2. Eliminar filas con valores en `engineSize` menores que 1 (no existen coches con tamaño menor a 1L)
# 3. Eliminar anomalías en `mpg` y `mileage` tomando sólo las filas que caigan dentro de µ ± 2σ (asumiendo una distribución normal esto sería el 95% de los casos en torno a la media)
#
#
#
#
# + id="7le8CAoV8Zbo"
# Filtering year
df = df[(df["year"] <= 2020) & (df["year"] >= 2000) ]
# Filtering engineSize
df = df[df["engineSize"] >= 0.5]
# Reducing sample size by mpg and mileage
df = df[(df.mpg > df.mpg.quantile(.025)) & (df.mpg < df.mpg.quantile(.975)) & (df.mileage > df.mileage.quantile(.025)) & (df.mileage < df.mileage.quantile(.975))]
# + [markdown] id="Bcli2t9L8SqH"
# ##### Resultados tras las primeras operaciones
# + id="6FcEbMF18cnh" outputId="dc71b650-29d1-498c-b8af-adfdf829de98" colab={"base_uri": "https://localhost:8080/", "height": 300}
df.describe()
# + id="5UhrElBPPWTM" outputId="e3739aa8-0928-4a38-b42b-44055508fb97" colab={"base_uri": "https://localhost:8080/", "height": 1000}
for column in df.columns[1:]:
plt.figure()
plt.xlabel(column)
plt.hist(df[column])
# + id="rMBIQxGvSoc7" outputId="6d733247-48f7-44d8-aa47-8df08813bae5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
index = 1
plt.figure(figsize=(14,20))
plt.suptitle('Price scatter plots v2', size = 28);
for column in df.columns[:6]:
if column == "price":
continue
plt.subplot(6,1,index)
plt.scatter(df[column], df['price'])
plt.xlabel(column)
index += 1
# Source: https://datascience.stackexchange.com/a/31755
# + [markdown] id="xyIOjDMtQTfq"
# ### Procesado de datos
# Tras ver los histogramas, decidimos procesar features a fin de obtener escalas similares y normalizar aquellos valores que consideramos nos pueden aportar utilidad
# + [markdown] id="ygdZVRDINVmd"
# ### Reescalado
# + id="2h5uyFI5NUnK"
min_max_scaler = sklearn.preprocessing.MinMaxScaler()
df['year'] = min_max_scaler.fit_transform(df[['year']])
# The ones below didn't bring any performance increases
#df['tax'] = min_max_scaler.fit_transform(df[['tax']])
#df['engineSize'] = min_max_scaler.fit_transform(df[['engineSize']])
#df['mpg'] = min_max_scaler.fit_transform(df[['mpg']])
#df['mileage'] = min_max_scaler.fit_transform(df[['mileage']])
# + id="o5iLXxz30uSl" outputId="7854e3b8-74c7-4f24-cd7a-7e462b85226d" colab={"base_uri": "https://localhost:8080/", "height": 300}
df.describe()
# + [markdown] id="m2lcK4ktOXUz"
# ### One-Hot Encoding
# *One-Hot Encoder* a las clases `brand`, `model`, `transmission`, `fuelType`
# + id="suRnRsuy0Uy4"
# One-Hot Encoder
for column_name, prefix in {"brand":"", "model":"model", "transmission": "gearbox", "fuelType":"fuel"}.items():
if prefix:
df = pd.concat([df, pd.get_dummies(df[column_name], prefix=prefix)], axis=1)
else:
df = pd.concat([df, pd.get_dummies(df[column_name])], axis=1)
df.drop([column_name], axis=1, inplace=True)
# + id="P7z-iDJ5jkky" outputId="864b751b-35e8-4fee-b5ee-204beca714d1" colab={"base_uri": "https://localhost:8080/", "height": 290}
df.head()
# + [markdown] id="T9xRbhJg7PhR"
# ### Resultados finales
# + id="vgrpfEM17gYk" outputId="fc1a32ae-56c3-43df-eb9c-69268a869d6b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
for column in df.columns[1:6]:
plt.figure()
plt.xlabel(column)
plt.hist(df[column])
# + id="5ZWyHBmU7hCR" outputId="46a2e2b8-692e-45e8-d5aa-ae7c386a969e" colab={"base_uri": "https://localhost:8080/", "height": 1000}
index = 1
plt.figure(figsize=(10,30))
plt.suptitle('Price scatter plots v3', size = 28);
for column in df.columns[:6]:
if column == "price":
continue
plt.subplot(6,1,index)
plt.scatter(df[column], df['price'])
plt.xlabel(column)
index += 1
# Source: https://datascience.stackexchange.com/a/31755
# + [markdown] id="QIPKWUltl4nR"
# ### Obtener X e y (X_train, X_test, y_train, y_test)
# + id="3hLBWf3NlLrH" outputId="c57dcb6a-c67c-47cb-efbb-1213eaa8b367" colab={"base_uri": "https://localhost:8080/"}
X = np.array(df.iloc[:, 1:])
X
# + id="ISNVM1RylyR7" outputId="f5b62516-010f-48e1-ffdc-47aeecadcd3b" colab={"base_uri": "https://localhost:8080/"}
y = np.array(df['price'])
y
# + id="vo_SbnVa9CIg"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=random_state)
# + [markdown] id="6-b7hn8Wsa5m"
# ## Optimización de hiperparámetros mediante GridSearch con Cross Validation
#
#
# + id="6W3ESTazlXgn"
# Parameter cv is 5 by default, adding more scoring methods would make it take too long to train
ridge_grid = GridSearchCV(
Ridge(),
{
'alpha': (0.001, 0.005, 0.01), # Started with (0.001, 0.01, 0.1, 0.5, 0.9, 0.99)
'max_iter': (10,50,100), # Started with (500, 1000, 1500)
},
scoring= ('neg_mean_absolute_error', "neg_root_mean_squared_error"),
refit = 'neg_mean_absolute_error',
n_jobs=-1,
)
lasso_grid = GridSearchCV(
Lasso(),
{
'alpha': (0.01, 0.1), # Started with (0.01, 0.1, 1)
'max_iter': (1500, 2000), # Started with (500, 1000, 1500)
},
scoring= ('neg_mean_absolute_error', "neg_root_mean_squared_error"),
refit = 'neg_mean_absolute_error',
n_jobs=-1
)
elastic_net_grid = GridSearchCV(
ElasticNet(),
{
'alpha': (0.01, 0.02), # Started with (0.01, 0.1, 1)
'l1_ratio': (0.01, 0.5, 0.99), # Started with np.linspace(0.15, 0.85, 3),
'max_iter': (1500, 2000), # Started with (500, 1000)
},
scoring= ('neg_mean_absolute_error', "neg_root_mean_squared_error"),
refit = 'neg_mean_absolute_error',
n_jobs=-1
)
# + [markdown] id="QQR2M-rRgytK"
# ## Modelos Machine Learning
#
# + id="lWkExxtIhEUl"
linear = LinearRegression().fit(X_train, y_train)
# + id="l4UHARFjzocS" outputId="50350f8c-4fd8-4b9a-f955-98c6127b7f23" colab={"base_uri": "https://localhost:8080/"}
with tf.device('/device:GPU:0'):
print("Ridge - Training")
ridge_grid.fit(X_train, y_train)
print("Ridge - DONE")
# + id="QTiFzbwOzNxG" outputId="d978d576-2839-4c6b-c735-3bc322d963c6" colab={"base_uri": "https://localhost:8080/"}
with tf.device('/device:GPU:0'):
print("Lasso - Training")
lasso_grid.fit(X_train, y_train)
print("Lasso - DONE")
# + id="lHRTMthJ3KlZ" outputId="cedf32ba-58c2-4bbd-881f-708c64700a13" colab={"base_uri": "https://localhost:8080/"}
with tf.device('/device:GPU:0'):
print("Elastic Net - Training")
elastic_net_grid.fit(X_train, y_train)
print("Elastic Net - DONE")
# It simply became a Lasso function when we last tried
# + [markdown] id="loNoX115mRlC"
# ### Scoring
# + id="8-8wEY3QpUjS" outputId="95744b00-642b-4208-bd2b-2d717f2647a4" colab={"base_uri": "https://localhost:8080/"}
print("Lineal:\n\tMAE")
print(f"\t\ttrain:\t{mean_absolute_error(y_train, linear.predict(X_train))}")
print(f"\t\ttest:\t{mean_absolute_error(y_test, linear.predict(X_test))}")
print("\tRMSE")
print(f"\t\ttrain:\t{mean_squared_error(y_train, linear.predict(X_train))**0.5}")
print(f"\t\ttest:\t{mean_squared_error(y_test, linear.predict(X_test))**0.5}")
print("\tR2")
print(f"\t\ttrain:\t{r2_score(y_train, linear.predict(X_train))}")
print(f"\t\ttest:\t{r2_score(y_test, linear.predict(X_test))}")
print('\n\t------\n')
print("Ridge:\n\tMAE")
print(f"\t\ttrain:\t{mean_absolute_error(y_train, ridge_grid.predict(X_train))}")
print(f"\t\ttest:\t{mean_absolute_error(y_test, ridge_grid.predict(X_test))}")
print("\tRMSE")
print(f"\t\ttrain:\t{mean_squared_error(y_train, ridge_grid.predict(X_train))**0.5}")
print(f"\t\ttest:\t{mean_squared_error(y_test, ridge_grid.predict(X_test))**0.5}")
print("\tR2")
print(f"\t\ttrain:\t{r2_score(y_train, ridge_grid.predict(X_train))}")
print(f"\t\ttest:\t{r2_score(y_test, ridge_grid.predict(X_test))}")
print('\n\t------\n')
print("Lasso:\n\tMAE")
print(f"\t\ttrain:\t{mean_absolute_error(y_train, lasso_grid.predict(X_train))}")
print(f"\t\ttest:\t{mean_absolute_error(y_test, lasso_grid.predict(X_test))}")
print("\tRMSE")
print(f"\t\ttrain:\t{mean_squared_error(y_train, lasso_grid.predict(X_train))**0.5}")
print(f"\t\ttest:\t{mean_squared_error(y_test, lasso_grid.predict(X_test))**0.5}")
print("\tR2")
print(f"\t\ttrain:\t{r2_score(y_train, lasso_grid.predict(X_train))}")
print(f"\t\ttest:\t{r2_score(y_test, lasso_grid.predict(X_test))}")
print('\n\t------\n')
print("Elastic Net:\n\tMAE")
print(f"\t\ttrain:\t{mean_absolute_error(y_train, elastic_net_grid.predict(X_train))}")
print(f"\t\ttest:\t{mean_absolute_error(y_test, elastic_net_grid.predict(X_test))}")
print("\tRMSE")
print(f"\t\ttrain:\t{mean_squared_error(y_train, elastic_net_grid.predict(X_train))**0.5}")
print(f"\t\ttest:\t{mean_squared_error(y_test, elastic_net_grid.predict(X_test))**0.5}")
print("\tR2")
print(f"\t\ttrain:\t{r2_score(y_train, elastic_net_grid.predict(X_train))}")
print(f"\t\ttest:\t{r2_score(y_test, elastic_net_grid.predict(X_test))}")
# + [markdown] id="4bdosuvLzh8P"
# # Conclusión:
# Tras haber realizado el procesamiento de datos, haber hecho optimización de hiperparámetros con GridSearchCV, y haber entrenado los modelos con los datos, comparando los scores (MAE, RMSE y R²) se puede observar que los modelos Linear, Ridge y Lasso son los que mejores resultados ofrecen, descartando Elastic Net debido a su alto error en comparación con estos modelos (cuando no se convierte directamente en una Lasso, en cuyo caso es más útil emplear dicho modelo directamente).
#
# A partir de aquí, ya obtendríamos el modelo que más se ajustase a nuestros requisitos, con los parámetros más óptimos, y podríamos emplear dicho modelo para que sea capaz de predecir los precios de los coches.
# + [markdown] id="8-8V3LB4diiR"
# # Testing
# + [markdown] id="Wegw__Ukd7sy"
# ## Análisis de datos
# + id="ySCdI2XkgX7q"
df_backup.brand.nunique() # should be 11
# + id="k_CpxaXvrnLT"
df_backup.fuelType.describe()
# + id="HM9q42Sjr67r"
df_backup.transmission.describe()
# + id="_G2OzyQYr-DL"
df_backup.transmission.unique()
# + id="zaXO5qbesHR-"
df_backup.fuelType.unique()
# + id="k0Hnq75bsQ3h"
print(sorted(df_backup.year.unique())) # To detect the anomaly
# + id="l6g9CROlm_0e"
len(X)
# + id="wG11dBmZjzSM"
pd.plotting.scatter_matrix(df_backup, alpha=0.2, figsize=(20,20))
# + id="nd71xAglpQ2s"
# We found a more efficient way to do it
column_transformer = sklearn.compose.ColumnTransformer(transformers=[
("scale", sklearn.preprocessing.StandardScaler(), [0]),
("passthrough", "passthrough", [1]),
("scale", sklearn.preprocessing.StandardScaler(), [2]),
("min-max", sklearn.preprocessing.MinMaxScaler(), [3]),
("one-hot", sklearn.preprocessing.OneHotEncoder(), [4])
]);
# + id="cbiNV4aeB1ov"
# Doesn't work
sigma = df_backup.describe().loc['std'][0]
mean = df_backup.describe().loc['mean'][0]
df_backup[(df_backup["mpg"] > mean - 2*sigma) & (df_backup["mpg"] < mean + 2*sigma)].describe()
# + id="VA7F17HCDpGt"
df_backup[(df_backup.mpg > df_backup.mpg.quantile(.025)) & (df_backup.mpg < df_backup.mpg.quantile(.975))].describe()
# + id="Oofi-MB8HSaC"
df_backup[df_backup["mpg"] == 11].head()
# + id="-_6aXDyF_oIJ"
df_backup[(df_backup["mpg"] > 11) & (df_backup["mpg"] <= 94)].describe()
# + id="DZG0V1BEIyeb"
df_backup[df_backup["mileage"] > 250000].head()
# + id="zQyFPxN9KA4f"
df_backup[(df_backup.tax >= df_backup.tax.quantile(.003)) & (df_backup.tax <= df.tax.quantile(.997))].describe()
# + id="z01wHoWJMh_8"
print(sorted(df_backup.engineSize.unique()))
# + id="VwRHqs5cRhim"
# Plotting experiment
import seaborn as sns
sns.pairplot(data=df_backup,y_vars=['price'],x_vars=df_backup.columns)
# Source: https://stackoverflow.com/a/43322569/9022642
# + id="ixxiq_MvsZJa" outputId="d154c55b-1853-42b5-85dd-9132636a4b5e" colab={"base_uri": "https://localhost:8080/", "height": 71}
# K-FOLDS CV to train Linear Reg. resulted in worse performance
print(f"\tMAE: {np.mean(cross_val_score(LinearRegression(), X_train, y_train, cv=5, scoring='neg_mean_absolute_error'))}")
print(f"\tMSE: {np.mean(cross_val_score(LinearRegression(), X_train, y_train, cv=5, scoring='neg_mean_squared_error'))}")
print(f"\tRMSE: {np.mean(cross_val_score(LinearRegression(), X_train, y_train, cv=5, scoring='neg_root_mean_squared_error'))}")
# + [markdown] id="cEkQJ94o0Ewh"
# ## Normalización
# + id="m8czwsKa0PO6" outputId="348e4c51-df22-45d5-be51-a8982ca00084" colab={"base_uri": "https://localhost:8080/", "height": 677}
# Normalisation (l1, l2, max) for this dataset doesn't give any advantages and in fact makes the data somewhat weird
# So we didn't apply it despite Linear Regression assuming Normal distributions
norm = 'l2'
df['engineSize'] = sklearn.preprocessing.normalize(np.array(df['engineSize']).reshape(-1, 1), norm=norm)
df['tax'] = sklearn.preprocessing.normalize(np.array(df['tax']).reshape(-1, 1), norm=norm)
df['mpg'] = sklearn.preprocessing.normalize(np.array(df['mpg']).reshape(-1, 1), norm=norm)
df['mileage'] = sklearn.preprocessing.normalize(np.array(df['mileage']).reshape(-1, 1), norm=norm)
# + [markdown] id="mfJUhLw1aumY"
# ## Estandarización
# + id="b1adRi7qaxe2"
# Tried even though it's not a normal distribution
df['mpg'] = sklearn.preprocessing.scale(df['mpg'])
# + [markdown] id="ieYxLMbX4Qjl"
# ## Iteraciones de grid search
#
# + id="qShV0VMJ2Xa2" outputId="ebef7871-7009-459b-f5cf-8ef8a6afae83" colab={"base_uri": "https://localhost:8080/"}
ridge_grid.best_estimator_
# + id="I76GIEBG2cfG" outputId="23fcb4c3-c309-42af-fd7d-9140c132d470" colab={"base_uri": "https://localhost:8080/"}
ridge_grid.best_score_
# + id="xNY6AFAA2iWJ" outputId="872d6997-d0fe-4dc7-bc4f-7587c395244e" colab={"base_uri": "https://localhost:8080/"}
lasso_grid.best_estimator_
# + id="lmigwuaF2kkh" outputId="07177834-4e5d-411d-d93b-643fbbf82fb5" colab={"base_uri": "https://localhost:8080/"}
lasso_grid.best_score_
# + id="9mk97eXl28UP" outputId="435e1ec1-8fb7-4d5c-d07f-ac6c3f1fe26e" colab={"base_uri": "https://localhost:8080/"}
elastic_net_grid.best_estimator_ # It went full Lasso
# + id="TvXt6ygA28aP" outputId="268689d0-34e5-425c-f307-7b9c2e40859e" colab={"base_uri": "https://localhost:8080/"}
elastic_net_grid.best_score_
# + id="CvC0_8Yvtk2i" outputId="ccadcd5f-fe90-4c22-e834-538fc69a45fc" colab={"base_uri": "https://localhost:8080/"}
ridge_grid_train = GridSearchCV(
Ridge(),
{
'alpha': (0.001, 0.01, 0.1, 0.5, 0.9, 0.99),
'max_iter': (500, 1000, 1500),
},
scoring= ('neg_mean_absolute_error', "neg_root_mean_squared_error"),
refit = 'neg_mean_absolute_error',
n_jobs=-1,
)
ridge_grid_train.fit(X_train, y_train)
# + id="62dEygvvQv9V" outputId="9fad64e5-4028-418c-8c85-9c1b019f5642" colab={"base_uri": "https://localhost:8080/"}
print(ridge_grid_train.best_score_*-1)
print(mean_absolute_error(y_test, ridge_grid_train.predict(X_test)))
# + id="agZuWhqrQp4k" outputId="5471f36b-a76b-407f-c0e0-480df438ee3a" colab={"base_uri": "https://localhost:8080/", "height": 34}
mean_absolute_error(y_test, ridge_grid.predict(X_test))
# + id="z0n242MpT6No" outputId="0e4aa151-9bcd-4590-9661-4d40094f1efa" colab={"base_uri": "https://localhost:8080/", "height": 51}
ridge = Ridge(alpha=0.01, max_iter=1000)
ridge.fit(X,y)
# + id="k9qjQzkzUHBH" outputId="528c3552-ac9a-4922-a1c3-ca33669d331d" colab={"base_uri": "https://localhost:8080/", "height": 34}
mean_absolute_error(y_test, ridge.predict(X_test))
# + [markdown] id="v5EPp-qDDeZc"
# # Referencias
#
# 1. https://www.inchcape.co.uk/blog/guides/what-size-engine-do-i-need/
# 2. https://motorway.co.uk/guides/best-cars-for-mpg
# 3. https://www.caranddriver.com/features/g30256394/worst-fuel-economy-cars/?slide=1
#
#
| Bosco Domingo, Luis de Marcos - P1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import multiprocessing
def cube(x):
return x**3
# %%timeit
for x in [1, 2, 3]:
cube(x)
# +
from multiprocessing import Process
def f(name):
print('hello', name)
if __name__ == '__main__':
p = Process(target=f, args=('bob',))
p.start()
p.join()
# -
multiprocessing.cpu_count()
# +
from multiprocessing import Pool, TimeoutError
import time
import os
def f(x):
return x*x
if __name__ == '__main__':
# start 4 worker processes
with Pool(processes=4) as pool:
# print "[0, 1, 4,..., 81]"
print(pool.map(f, range(10)))
# print same numbers in arbitrary order
for i in pool.imap_unordered(f, range(10)):
print(i)
# evaluate "f(20)" asynchronously
res = pool.apply_async(f, (20,)) # runs in *only* one process
print(res.get(timeout=1)) # prints "400"
# evaluate "os.getpid()" asynchronously
res = pool.apply_async(os.getpid, ()) # runs in *only* one process
print(res.get(timeout=1)) # prints the PID of that process
# launching multiple evaluations asynchronously *may* use more processes
multiple_results = [pool.apply_async(os.getpid, ()) for i in range(4)]
print([res.get(timeout=1) for res in multiple_results])
# make a single worker sleep for 10 secs
res = pool.apply_async(time.sleep, (10,))
try:
print(res.get(timeout=1))
except TimeoutError:
print("We lacked patience and got a multiprocessing.TimeoutError")
print("For the moment, the pool remains available for more work")
# exiting the 'with'-block has stopped the pool
print("Now the pool is closed and no longer available")
# -
| parallel processing play.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# 
Task1 = pd.read_excel("header.xlsx")
Task1.head()
# -----------------
# header=1 ---> Column names preesent in second row.
df = pd.read_excel("header.xlsx", header=1)
df.head()
# -------------------------------
| 1_Data_Frame/9_Header.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.version
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import sklearn
import json
import os
import joblib
from operator import itemgetter
import networkx as nx
from networkx.readwrite import json_graph
import matplotlib.colors as mcolors
import collections
DATA_PATH = 'data/keras-example/graph/train/'
PREFIX = 'keras'
# -
flatten = lambda l: [y for x in l for y in x]
var_map = json.load(open(DATA_PATH+PREFIX+"-var_map.json"))
vocab =list(set(var_map.values()))
flat_vocab = [x.split('_') for x in vocab]
flat_vocab = flatten(flat_vocab)
len(flat_vocab)
import keras
# +
from _ast import *
import sys
MOD_SYMBOLS = [Module, Interactive, Expression, Suite]
STMT_SYMBOLS = [FunctionDef, AsyncFunctionDef, ClassDef, Return, Delete, \
Assign, AugAssign, For, AsyncFor, While, If, With, AsyncWith, \
Raise, Try, Assert, Import, ImportFrom, Global, Nonlocal, \
Expr, Pass, Break, Continue]
EXPR_SYMBOLS = [BoolOp, BinOp, UnaryOp, Lambda, IfExp, Dict, Set, ListComp, \
SetComp, DictComp, GeneratorExp, Await, Yield, YieldFrom, \
Compare, Call, Num, Str, FormattedValue, JoinedStr, Bytes, \
NameConstant, Ellipsis, Constant, Attribute, Subscript, \
Starred, Name, List, Tuple]
EXPR_CONTENT_SYMBOLS = [Load, Store, Del, AugLoad, AugStore, Param]
SLICE_SYMBOLS = [Slice, ExtSlice, Index]
BOOLOP_SYMBOLS = [And, Or]
OPERATOR_SYMBOLS = [Add, Sub, Mult, MatMult, Div, Mod, Pow, LShift, RShift, \
BitOr, BitXor, BitAnd, FloorDiv]
UNARYOP_SYMBOLS = [Invert, Not, UAdd, USub]
CMPOP_SYMBOLS = [Eq, NotEq, Lt, LtE, Gt, GtE, Is, IsNot, In, NotIn]
COMPREHENSION_SYMBOLS = [comprehension]
EXCEPT_SYMBOLS = [excepthandler, ExceptHandler]
ARG_SYMBOLS = [arguments, arg, keyword]
IMPORT_SYMBOLS = [alias, withitem]
PYTHON_SYMBOLS = MOD_SYMBOLS + STMT_SYMBOLS + EXPR_SYMBOLS \
+ EXPR_CONTENT_SYMBOLS + SLICE_SYMBOLS \
+ BOOLOP_SYMBOLS + OPERATOR_SYMBOLS \
+ UNARYOP_SYMBOLS + CMPOP_SYMBOLS \
+ EXCEPT_SYMBOLS + ARG_SYMBOLS + IMPORT_SYMBOLS \
+ COMPREHENSION_SYMBOLS
AST_SYMBOL_DICT = dict((v, k) for (k, v) in enumerate(PYTHON_SYMBOLS))
# -
inv_ast_symbol_dict = joblib.load(filename='inv_ast_symbol_dict')
import csv
lst = [k.__name__ for k in AST_SYMBOL_DICT.keys()]
with open(..., 'wb') as myfile:
wr = csv.writer(myfile, quoting=csv.QUOTE_ALL)
wr.writerow(mylist)
| notebook/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="8Xdmm79IChoP" colab_type="code" colab={}
import pandas as pd
import scipy.stats as stats
# + id="6o9H0nwZCkJk" colab_type="code" colab={}
df_collection = pd.read_csv('https://raw.githubusercontent.com/niravjdn/Software-Measurement-Project/master/data/jacoc-by-version/collection/collections-v4.2.csv', error_bad_lines=False)
# + id="F4Zm04R7Dir8" colab_type="code" colab={}
import matplotlib.pyplot as plt
# + id="U1nY7nhdFF4e" colab_type="code" colab={}
df_collection['Statement_Percentage'] = (df_collection['LINE_COVERED'] / (df_collection['LINE_COVERED'] + df_collection['LINE_MISSED'])) * 100
# + id="0sGHQ9qLFw1u" colab_type="code" colab={}
df_collection['Branch_Percentage'] = (df_collection['BRANCH_COVERED'] / (df_collection['BRANCH_COVERED'] + df_collection['BRANCH_MISSED'])) * 100
# + id="G1uiGZmiGBe8" colab_type="code" colab={}
df_collection['CC'] = df_collection['COMPLEXITY_COVERED'] + df_collection['COMPLEXITY_MISSED'];
# + id="cbxZwOnTDpfv" colab_type="code" outputId="2831c861-efbc-4c97-8e12-3a0f52fbcec1" executionInfo={"status": "ok", "timestamp": 1554515897023, "user_tz": 240, "elapsed": 1317, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 660}
df_collection.head()
# + id="Ztm1rZvLEeyS" colab_type="code" outputId="6ccd3ca0-767e-4c95-cc8a-7919b405f084" executionInfo={"status": "ok", "timestamp": 1554515897617, "user_tz": 240, "elapsed": 1872, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 378}
df_collection.plot(x='CC', y='Statement_Percentage', style='o')
# + id="RyiTWuCqo9DT" colab_type="code" outputId="ed77b24a-77e1-44d9-ab4b-7d591dca0564" executionInfo={"status": "ok", "timestamp": 1554515898845, "user_tz": 240, "elapsed": 3076, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 378}
df_collection.plot(x='CC', y='Branch_Percentage', style='o')
# + id="ufAGflaPGfZD" colab_type="code" outputId="3294db8c-e6fb-472f-cd69-1a88aace7d45" executionInfo={"status": "ok", "timestamp": 1554515899279, "user_tz": 240, "elapsed": 3490, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 347}
plt.scatter(df_collection['CC'], df_collection['Statement_Percentage'])
plt.show() # Depending on whether you use IPython or interactive mode, etc.
# + id="yXwX1zpHHhOU" colab_type="code" outputId="6016c2c4-9272-4c98-d2b7-fa3aedc4ed43" executionInfo={"status": "ok", "timestamp": 1554515899286, "user_tz": 240, "elapsed": 3477, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 477}
df_collection.corr(method ='spearman')
# + id="LlZdN8Q8Ig0l" colab_type="code" outputId="b8e36918-549c-45ff-a714-e842d0d54054" executionInfo={"status": "ok", "timestamp": 1554515899290, "user_tz": 240, "elapsed": 3461, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 112}
df_collection[['CC','Statement_Percentage']].corr(method ='spearman')
# + id="T86BizaEQ9Mb" colab_type="code" outputId="a725ff67-67f0-4e80-9385-4ddfaaaafa2e" executionInfo={"status": "ok", "timestamp": 1554516409171, "user_tz": 240, "elapsed": 476, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df_clean = df_collection.dropna()
stats.spearmanr(df_clean['Statement_Percentage'], df_clean['CC'])
# + id="1DAV9QEjIxpy" colab_type="code" outputId="1e4aeb0d-7f1f-498f-e8e6-ef7f9f0be94a" executionInfo={"status": "ok", "timestamp": 1554515899296, "user_tz": 240, "elapsed": 3415, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 112}
df_collection[['CC','Branch_Percentage']].corr(method ='spearman')
# + id="LdbShMPAQ-R9" colab_type="code" outputId="7c5663d7-ecca-43a0-a029-4a730c34d9c9" executionInfo={"status": "ok", "timestamp": 1554516074047, "user_tz": 240, "elapsed": 396, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df_clean = df_collection.dropna()
stats.spearmanr(df_clean['Branch_Percentage'], df_clean['CC'])
# + id="3FBbpGRwlP5l" colab_type="code" outputId="2d49f1e5-13b1-4e82-8172-6df4f4c97989" executionInfo={"status": "ok", "timestamp": 1554515899305, "user_tz": 240, "elapsed": 3376, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
print('Total Statment Coverage '+str((df_collection.LINE_COVERED.sum()/(df_collection.LINE_MISSED.sum() + df_collection.LINE_COVERED.sum()))*100))
# + id="PWuIdk61pENk" colab_type="code" outputId="dca88330-4ba6-4d18-ac66-36487cab50a7" executionInfo={"status": "ok", "timestamp": 1554515899309, "user_tz": 240, "elapsed": 3353, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MXavU57lU4k/AAAAAAAAAAI/AAAAAAAAMFw/tzJ-F4ETDCM/s64/photo.jpg", "userId": "16537180351831762327"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
print('Total Branch Coverage '+str((df_collection.BRANCH_COVERED.sum()/(df_collection.BRANCH_MISSED.sum() + df_collection.BRANCH_COVERED.sum()))*100))
# + id="MVF4iS4X6VoF" colab_type="code" colab={}
| Jupyter Notebook/Jupyter Notebok/Final Metrics Correlation/Metric 1,2 to 3,4/Jacoco/Collection/Collection 4.2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Checkbox
# +
from PyQt5 import QtCore, QtGui, QtWidgets
class Ui_Dialog(object):
def setupUi(self, Dialog):
Dialog.setObjectName("Dialog")
Dialog.resize(413, 306)
self.verticalLayout_3 = QtWidgets.QVBoxLayout(Dialog)
self.verticalLayout_3.setObjectName("verticalLayout_3")
self.verticalLayout = QtWidgets.QVBoxLayout()
self.verticalLayout.setObjectName("verticalLayout")
self.label = QtWidgets.QLabel(Dialog)
font = QtGui.QFont()
font.setPointSize(14)
font.setBold(True)
font.setWeight(75)
self.label.setFont(font)
self.label.setObjectName("label")
self.verticalLayout.addWidget(self.label)
self.horizontalLayout = QtWidgets.QHBoxLayout()
self.horizontalLayout.setObjectName("horizontalLayout")
self.label_2 = QtWidgets.QLabel(Dialog)
self.label_2.setObjectName("label_2")
self.horizontalLayout.addWidget(self.label_2)
self.verticalLayout_2 = QtWidgets.QVBoxLayout()
self.verticalLayout_2.setObjectName("verticalLayout_2")
self.checkBoxPizza = QtWidgets.QCheckBox(Dialog)
self.checkBoxPizza.setObjectName("checkBoxPizza")
self.checkBoxPizza.stateChanged.connect(self.checked_item)
self.verticalLayout_2.addWidget(self.checkBoxPizza)
self.checkBoxSalad = QtWidgets.QCheckBox(Dialog)
self.checkBoxSalad.setObjectName("checkBoxSalad")
self.verticalLayout_2.addWidget(self.checkBoxSalad)
self.checkBoxSalad.stateChanged.connect(self.checked_item)
self.checkBoxSausage = QtWidgets.QCheckBox(Dialog)
self.checkBoxSausage.setObjectName("checkBoxSausage")
self.verticalLayout_2.addWidget(self.checkBoxSausage)
self.horizontalLayout.addLayout(self.verticalLayout_2)
#added newly after conversion for connecting signal to slot
self.checkBoxSausage.stateChanged.connect(self.checked_item)
self.verticalLayout.addLayout(self.horizontalLayout)
self.labelResult = QtWidgets.QLabel(Dialog)
font = QtGui.QFont()
font.setPointSize(12)
font.setBold(True)
font.setWeight(75)
self.labelResult.setFont(font)
self.labelResult.setText("")
self.labelResult.setObjectName("labelResult")
self.verticalLayout.addWidget(self.labelResult)
self.verticalLayout_3.addLayout(self.verticalLayout)
self.retranslateUi(Dialog)
QtCore.QMetaObject.connectSlotsByName(Dialog)
def retranslateUi(self, Dialog):
_translate = QtCore.QCoreApplication.translate
Dialog.setWindowTitle(_translate("Dialog", "PyQt5 QCheckBoxes"))
self.label.setText(_translate("Dialog", "Regular Tuna Price : 20"))
self.label_2.setText(_translate("Dialog", "Select Extra"))
self.checkBoxPizza.setText(_translate("Dialog", "Pizza : 3"))
self.checkBoxSalad.setText(_translate("Dialog", "Salad : 4"))
self.checkBoxSausage.setText(_translate("Dialog", "Sausage : 5"))
#this is method is added after conversion
def checked_item(self):
price = 20
if self.checkBoxPizza.isChecked():
price = price + 3
if self.checkBoxSalad.isChecked():
price = price + 4
if self.checkBoxSausage.isChecked():
price = price + 5
self.labelResult.setText("Total Price Is : {} ".format(price))
if __name__ == "__main__":
import sys
app = QtWidgets.QApplication(sys.argv)
Dialog = QtWidgets.QDialog()
ui = Ui_Dialog()
ui.setupUi(Dialog)
Dialog.show()
sys.exit(app.exec_())
# +
from PyQt5.QtWidgets import QApplication,QDialog,QHBoxLayout, QVBoxLayout, QLabel, QCheckBox
import sys
from PyQt5.QtGui import QIcon, QIcon, QFont
from PyQt5.QtCore import QSize
class Window(QDialog):
def __init__(self):
super().__init__()
#window title, icon and geometry
self.setGeometry(200,200, 400,200)
self.setWindowTitle("PyQt5 CheckBox")
self.setWindowIcon(QIcon('python.png'))
self.create_checkbox()
def create_checkbox(self):
#this is our hboxlayout
hbox = QHBoxLayout()
#these are checkboxes
self.check1 = QCheckBox("Football")
self.check1.setIcon(QIcon('football.png'))
self.check1.setIconSize(QSize(40,40))
self.check1.setFont(QFont("Sanserif", 13))
self.check1.toggled.connect(self.item_selected)
hbox.addWidget(self.check1)
self.check2 = QCheckBox("Cricket")
self.check2.setIcon(QIcon('cricket.png'))
self.check2.setIconSize(QSize(40, 40))
self.check2.setFont(QFont("Sanserif", 13))
self.check2.toggled.connect(self.item_selected)
hbox.addWidget(self.check2)
self.check3 = QCheckBox("Tennis")
self.check3.setIcon(QIcon('tennis.png'))
self.check3.setIconSize(QSize(40, 40))
self.check3.setFont(QFont("Sanserif", 13))
self.check3.toggled.connect(self.item_selected)
hbox.addWidget(self.check3)
#this is the vboxlayout
vbox = QVBoxLayout()
#we have created a label in here
self.label = QLabel("Hello")
self.label.setFont(QFont("Sanserif", 15))
#add the label in the vbox layout
vbox.addWidget(self.label)
#add the hbox layout in the vbox layout
vbox.addLayout(hbox)
#set the layout for the main window
self.setLayout(vbox)
def item_selected(self):
value = ""
#check for the check box value
if self.check1.isChecked():
value = self.check1.text()
if self.check2.isChecked():
value = self.check2.text()
if self.check3.isChecked():
value = self.check3.text()
self.label.setText("You have selected : " + value)
app = QApplication(sys.argv)
window = Window()
window.show()
sys.exit(app.exec_())
# -
# ### Spinbox
# +
from PyQt5.QtWidgets import QApplication, QDialog, QHBoxLayout, QSpinBox, QLabel, QLineEdit
import sys
from PyQt5.QtGui import QIcon
class Window(QDialog):
def __init__(self):
super().__init__()
#our window requirements like icon,title
self.setGeometry(200,200,400,300)
self.setWindowTitle("PyQt5 SpinBox")
self.setWindowIcon(QIcon("python.png"))
self.create_spinbox()
def create_spinbox(self):
#cretae hboxlayout
hbox = QHBoxLayout()
#in here we have created our label
label = QLabel("Laptop Price : ")
#this is our linedit
self.lineEdit = QLineEdit()
#we need to create the object of QSpinBox class
self.spinbox = QSpinBox()
#we have connected valueChanged signal
self.spinbox.valueChanged.connect(self.spin_selected)
self.totalResult = QLineEdit()
#add widgets to your hbox layout
hbox.addWidget(label)
hbox.addWidget(self.lineEdit)
hbox.addWidget(self.spinbox)
hbox.addWidget(self.totalResult)
self.setLayout(hbox)
def spin_selected(self):
if self.lineEdit.text() != 0:
price = int(self.lineEdit.text())
totalPrice = self.spinbox.value() * price
self.totalResult.setText(str(totalPrice))
else:
print("Wrong value")
App = QApplication(sys.argv)
window = Window()
window.show()
sys.exit(App.exec())
# +
from PyQt5.QtWidgets import QApplication, QDialog, QLineEdit,\
QDoubleSpinBox
from PyQt5 import uic
class UI(QDialog):
def __init__(self):
super().__init__()
#loading the ui file with uic module
uic.loadUi("doublespin.ui", self)
#finding child in the ui file
self.linePrice = self.findChild(QLineEdit, "lineEditPrice")
self.doublespin = self.findChild(QDoubleSpinBox, "doubleSpinBox")
self.doublespin.valueChanged.connect(self.spin_selected)
self.lineResult = self.findChild(QLineEdit, "lineEditTotal")
def spin_selected(self):
if self.linePrice.text() != 0:
price = int(self.linePrice.text())
totalPrice = self.doublespin.value() * price
self.lineResult.setText(str(totalPrice))
app = QApplication([])
window = UI()
window.show()
app.exec_()
# -
| Day 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 推測統計の基本
# ## 母集団と標本
# ### 標本の抽出方法
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %precision 3
# %matplotlib inline
# -
df = pd.read_csv('../data/ch4_scores400.csv')
scores = np.array(df['点数'])
scores[:10]
np.random.choice([1, 2, 3], 3)
np.random.choice([1, 2, 3], 3, replace=False)
np.random.seed(0)
np.random.choice([1, 2, 3], 3)
# +
np.random.seed(0)
sample = np.random.choice(scores, 20)
sample.mean()
# -
scores.mean()
for i in range(5):
sample = np.random.choice(scores, 20)
print(f'{i+1}回目の無作為抽出で得た標本平均', sample.mean())
# ## 確率モデル
# ### 確率の基本
# ### 確率分布
dice = [1, 2, 3, 4, 5, 6]
prob = [1/21, 2/21, 3/21, 4/21, 5/21, 6/21]
np.random.choice(dice, p=prob)
num_trial = 100
sample = np.random.choice(dice, num_trial, p=prob)
sample
freq, _ = np.histogram(sample, bins=6, range=(1, 7))
pd.DataFrame({'度数':freq,
'相対度数':freq / num_trial},
index = pd.Index(np.arange(1, 7), name='出目'))
fig = plt.figure(figsize=(10, 6))
ax = fig.add_subplot(111)
ax.hist(sample, bins=6, range=(1, 7), density=True, rwidth=0.8)
# 真の確率分布を横線で表示
ax.hlines(prob, np.arange(1, 7), np.arange(2, 8), colors='gray')
# 棒グラフの[1.5, 2.5, ..., 6.5]の場所に目盛りをつける
ax.set_xticks(np.linspace(1.5, 6.5, 6))
# 目盛りの値は[1, 2, 3, 4, 5, 6]
ax.set_xticklabels(np.arange(1, 7))
ax.set_xlabel('出目')
ax.set_ylabel('相対度数')
plt.show()
# +
num_trial = 10000
sample = np.random.choice(dice, size=num_trial, p=prob)
fig = plt.figure(figsize=(10, 6))
ax = fig.add_subplot(111)
ax.hist(sample, bins=6, range=(1, 7), density=True, rwidth=0.8)
ax.hlines(prob, np.arange(1, 7), np.arange(2, 8), colors='gray')
ax.set_xticks(np.linspace(1.5, 6.5, 6))
ax.set_xticklabels(np.arange(1, 7))
ax.set_xlabel('出目')
ax.set_ylabel('相対度数')
plt.show()
# -
# ## 推測統計における確率
fig = plt.figure(figsize=(10, 6))
ax = fig.add_subplot(111)
ax.hist(scores, bins=100, range=(0, 100), density=True)
ax.set_xlim(20, 100)
ax.set_ylim(0, 0.042)
ax.set_xlabel('点数')
ax.set_ylabel('相対度数')
plt.show()
np.random.choice(scores)
# +
sample = np.random.choice(scores, 10000)
fig = plt.figure(figsize=(10, 6))
ax = fig.add_subplot(111)
ax.hist(sample, bins=100, range=(0, 100), density=True)
ax.set_xlim(20, 100)
ax.set_ylim(0, 0.042)
ax.set_xlabel('点数')
ax.set_ylabel('相対度数')
plt.show()
# +
sample_means = [np.random.choice(scores, 20).mean()
for _ in range(10000)]
fig = plt.figure(figsize=(10, 6))
ax = fig.add_subplot(111)
ax.hist(sample_means, bins=100, range=(0, 100), density=True)
# 母平均を縦線で表示
ax.vlines(np.mean(scores), 0, 1, 'gray')
ax.set_xlim(50, 90)
ax.set_ylim(0, 0.13)
ax.set_xlabel('点数')
ax.set_ylabel('相対度数')
plt.show()
# -
# ## これから学ぶこと
| notebook/chap04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/noiroiseauun/MLOverflow/blob/main/NB_tests.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="k_angBtGWmRL"
# Import statements
import csv
import string
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import confusion_matrix
from sklearn.linear_model import SGDRegressor
from sklearn.naive_bayes import CategoricalNB
from datetime import datetime, timedelta
import gc
import pandas as pd
import random
import re
from google.colab import drive
# + colab={"base_uri": "https://localhost:8080/"} id="AH5Al09iXIC5" outputId="415949e0-c0f9-4c5b-fd21-d5f393984f76"
drive.mount('/content/drive')
datapath = '/content/drive/Shared drives/CMPUT 466 - Project/data/'
questionsPath = f"{datapath}noHTMLQuestions.csv"
answersPath = f"{datapath}Answers-Final.csv"
tagsPath = f"{datapath}Tags-Final.csv"
# + id="L3frLVbvXQsW"
# answerdf = pd.read_csv("/content/drive/Shared drives/CMPUT 466 - Project/data/Answers-Final.csv", encoding="ISO-8859-1")
answerdf = pd.read_csv(answersPath, encoding="ISO-8859-1")
questiondf = pd.read_csv(questionsPath, encoding="ISO-8859-1", header=None)
tagdf = pd.read_csv(tagsPath)
# + id="HBU8QboB_5YI"
questionCols = {0:"Id", 1:"OwnerUserId", 2:"CreationDate", 3:"ClosedDate", 4:"Score", 5:"Title", 6:"Body"}
questiondf.rename(mapper=questionCols, axis='columns', inplace=True)
# + id="ofYszUeIZIKM"
tagCounts = tagdf["Tag"].value_counts(normalize=False)
tmpDict = tagCounts.to_dict()
tagdf["counts"] = tagdf["Tag"].map(tmpDict)
tagdf = tagdf[tagdf["counts"] > 101180]
# + id="bLoa0Vp7b4sM"
jointdf = pd.merge(tagdf, questiondf, how="left", left_on="Id", right_on="Id")
# + id="J7h3JbxIjK4O"
SEED = 19
random.seed(SEED)
SAMPLE = 10000
# + id="vOoRkm6lcLdR"
tagSample = jointdf.sample(SAMPLE)
vectorizer = CountVectorizer(
lowercase=True, stop_words='english',
max_df=1.0, min_df=1, max_features=None, binary=True,
encoding="ISO-8859-1"
)
def removeWhitespace(val):
return re.sub('\s', ' ', val)
# tagSample = jointdf[:10000] # For consistency, don't sample yet
# tagSample["Body"] = tagSample["Body"].apply(removeWhitespace)
body = tagSample["Body"].apply(removeWhitespace).to_numpy()
tag = tagSample["Tag"].to_numpy()
X = vectorizer.fit_transform(body).toarray()
classifier = CategoricalNB()
# + colab={"base_uri": "https://localhost:8080/"} id="nCRJLj3uPZUF" outputId="d48a25a0-223d-4c91-92c7-7320d804f53a"
spliter = 9000
x_train = X[0:spliter]
x_test = X[spliter:]
y_train = tag[0:spliter]
y_test = tag[spliter:]
print("x_train shape: {}".format(x_train.shape))
print("y_train shape: {}".format(y_train.shape))
print("x_test shape: {}".format(x_test.shape))
print("y_test shape: {}".format(y_test.shape))
# + colab={"base_uri": "https://localhost:8080/"} id="CQXWs0YVeLNv" outputId="4f86cef2-009c-4a8d-eb8f-e9bff6d0c295"
classifier.fit(x_train, y_train)
# + id="ry03gQe7JoTq"
# classifier.score(x_test, y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="cZdtGpReJwxU" outputId="cf06adc4-0d60-41f1-ae53-ab0008f3bf2a"
failList = list()
missList = list()
count = 0
tmpResultlist = list()
length = len(x_test)
for index in range(length):
try:
result = classifier.predict(x_test[index:index+1])
tmpResultlist.append(result[0])
if result[0] == y_test[index]: count += 1
else:
missList.append(index)
# print("result was {} and true was {}".format(result[0], y_test[index]))
except Exception as e:
# print("index {} failed: {}".format(index, e))
failList.append(index)
failed = len(failList)
missed = len(missList)
print("FAILED: {}".format(failed))
print("Correct: {}".format(count/(length-failed)))
print("missed: {}".format(missed/(length-failed)))
# print("count: {}".format(count))
# print(tmpResultlist)
# + id="njneR1-Jk7SF"
| NB_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table> <tr>
# <td style="background-color:#ffffff;">
# <a href="http://qworld.lu.lv" target="_blank"><img src="../images/qworld.jpg" width="25%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <a href="http://abu.lu.lv" target="_blank"><NAME></a> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# <h2>Quantum Teleportation</h2>
#
# [Watch Lecture](https://youtu.be/4PYeoqALKHk)
#
# <hr>
#
# _**Prepare a few blank sheets of paper**_
# - _**to draw the circuit of the following protocol step by step and**_
# - _**to solve some of tasks requiring certain calculations.**_
#
# <hr>
# Asja wants to send a qubit to Balvis by using only classical communication.
#
# Let $ \ket{v} = \myvector{a\\b} \in \mathbb{R}^2 $ be the quantum state.
#
# _Discussion:_ If Asja has many copies of this qubit, then she can collect the statistics based on these qubits and obtain an approximation of $ a $ and $ b $, say $ \tilde{a} $ and $\tilde{b}$, respectively. After this, Asja can send $ \tilde{a} $ and $\tilde{b}$ by using many classical bits, the number of which depends on the precision of the amplitudes.
# On the other hand, If Asja and Balvis share the entangaled qubits in state $ \sqrttwo\ket{00} + \sqrttwo\ket{11} $ in advance, then it is possible for Balvis to create $ \ket{v} $ in his qubit after receiving two bits of information from Asja.
# <h3> Protocol </h3>
#
# The protocol uses three qubits as specified below:
#
# <img src='../images/quantum_teleportation_qubits.png' width="25%" align="left">
# Asja has two qubits and Balvis has one qubit.
#
# Asja's quantum message (key) is $ \ket{v} = \myvector{a\\b} = a\ket{0} + b\ket{1} $.
#
# The entanglement between Asja's second qubit and Balvis' qubit is $ \sqrttwo\ket{00} + \sqrttwo\ket{11} $.
#
# So, the quantum state of the three qubits is
#
# $$ \mypar{a\ket{0} + b\ket{1}}\mypar{\sqrttwo\ket{00} + \sqrttwo\ket{11}}
# = \sqrttwo \big( a\ket{000} + a \ket{011} + b\ket{100} + b \ket{111} \big). $$
# <h4> CNOT operator by Asja </h4>
#
# Asja applies CNOT gate to her qubits where $q[2]$ is the control qubit and $q[1]$ is the target qubit.
# <h3>Task 1</h3>
#
# Calculate the new quantum state after this CNOT operator.
# <a href="B56_Quantum_Teleportation_Solutions.ipynb#task1">click for our solution</a>
# <h3>Hadamard operator by Asja</h3>
#
# Asja applies Hadamard gate to $q[2]$.
# <h3>Task 2</h3>
#
# Calculate the new quantum state after this Hadamard operator.
#
# Verify that the resulting quantum state can be written as follows:
#
# $$
# \frac{1}{2} \ket{00} \big( a\ket{0}+b\ket{1} \big) +
# \frac{1}{2} \ket{01} \big( a\ket{1}+b\ket{0} \big) +
# \frac{1}{2} \ket{10} \big( a\ket{0}-b\ket{1} \big) +
# \frac{1}{2} \ket{11} \big( a\ket{1}-b\ket{0} \big) .
# $$
# <a href="B56_Quantum_Teleportation_Solutions.ipynb#task2">click for our solution</a>
# <h3> Measurement by Asja </h3>
#
# Asja measures her qubits. With probability $ \frac{1}{4} $, she can observe one of the basis states.
#
# Depeding on the measurement outcomes, Balvis' qubit is in the following states:
# <ol>
# <li> "00": $ \ket{v_{00}} = a\ket{0} + b \ket{1} $ </li>
# <li> "01": $ \ket{v_{01}} = a\ket{1} + b \ket{0} $ </li>
# <li> "10": $ \ket{v_{10}} = a\ket{0} - b \ket{1} $ </li>
# <li> "11": $ \ket{v_{11}} = a\ket{1} - b \ket{0} $ </li>
# </ol>
# As can be observed, the amplitudes $ a $ and $ b $ are "transferred" to Balvis' qubit in each case.
#
# If Asja sends the measurement outcomes, then Balvis can construct $ \ket{v} $ exactly.
# <h3>Task 3</h3>
#
# Asja sends the measurement outcomes to Balvis by using two classical bits: $ x $ and $ y $.
#
# For each $ (x,y) $ pair, determine the quantum operator(s) that Balvis can apply to obtain $ \ket{v} = a\ket{0}+b\ket{1} $ exactly.
# <a href="B56_Quantum_Teleportation_Solutions.ipynb#task3">click for our solution</a>
# <h3> Task 4 </h3>
#
# Create a quantum circuit with three qubits as described at the beginning of this notebook and two classical bits.
#
# Implement the protocol given above until Asja makes the measurements (included).
# - The state of $q[2]$ can be set by the rotation with a randomly picked angle.
# - Remark that Balvis does not make the measurement.
#
# At this point, read the state vector of the circuit by using "statevector_simulator".
#
# _When a circuit having measurement is simulated by "statevector_simulator", the simulator picks one of the outcomes, and so we see one of the states after the measurement._
#
# Verify that the state of Balvis' qubit is in one of these: $ \ket{v_{00}}$, $ \ket{v_{01}}$, $ \ket{v_{10}}$, and $ \ket{v_{11}}$.
#
# Guess the measurement outcome obtained by "statevector_simulator".
#
# your code is here
#
# <a href="B56_Quantum_Teleportation_Solutions.ipynb#task4">click for our solution</a>
# <h3> Task 5 </h3>
#
# Implement the protocol above by including the post-processing part done by Balvis, i.e., the measurement results by Asja are sent to Balvis and then he may apply $ X $ or $ Z $ gates depending on the measurement results.
#
# We use the classically controlled quantum operators.
#
# Since we do not make measurement on $ q[0] $, we define only 2 classical bits, each of which can also be defined separated.
#
# q = QuantumRegister(3)
# c2 = ClassicalRegister(1,'c2')
# c1 = ClassicalRegister(1,'c1')
# qc = QuantumCircuit(q,c1,c2)
# ...
# qc.measure(q[1],c1)
# ...
# qc.x(q[0]).c_if(c1,1) # x-gate is applied to q[0] if the classical bit c1 is equal to 1
#
# Read the state vector and verify that Balvis' state is $ \myvector{a \\ b} $ after the post-processing.
#
# your code is here
#
# <a href="B56_Quantum_Teleportation_Solutions.ipynb#task5">click for our solution</a>
# <!--
# <h3> Task 6 (optional) </h3>
#
# Observe that Balvis can also t
#
# Create a quantum circuit with four qubits and four classical bits.
#
# Assume that Asja has the first two qubits (number 3 and 2) and Balvis has the last two qubits (number 1 and 0).
#
# Create an entanglement between qubits 2 and 1.
#
# Implement the protocol (the state of the qubit can be set by a rotation with randomly picked angle):
# - If Asja teleports a qubit, then set the state of qubit 3.
# - If Balvis teleports a qubit, then set the state of qubit 0.
# -->
| bronze/B56_Quantum_Teleportation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''deeplearning'': conda)'
# language: python
# name: python37564bitdeeplearningconda2f5dcc693383402099797ed40bd3951d
# ---
# %load_ext autoreload
# %autoreload 2
import timm
import sys
sys.path.append('../src')
from models import SimpleClassificationModel
from dataset import WindDataset
from transforms import get_valid_transforms
import neptune
from os.path import join as join_path
import pandas as pd
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
import torch
import numpy as np
from tqdm.auto import tqdm
from sklearn.metrics import mean_squared_error
import seaborn as sns
# %matplotlib inline
data_dir = '../data/'
image_dir = '../data/all_images/'
sample_sub = pd.read_csv(join_path(data_dir, 'submission_format.csv'))
train_labels = pd.read_csv(join_path(data_dir, 'training_set_labels.csv'))
test_images = [x+'.jpg' for x in sample_sub['image_id']]
# Get test loader
test_dataset = WindDataset(test_images, image_dir, get_valid_transforms(256), 1)
test_loader = DataLoader(test_dataset, batch_size=16, shuffle=False)
model = SimpleClassificationModel('resnest200e', pretrained=False)
weights = torch.load('../logs/21.14_resnest200e_adamw_heavy_fold_0/21.14_resnest200e_adamw_heavy_fold_0_best_val_loss=67.265.ckpt')['state_dict']
weights = {k.replace('net.', ''): v for k, v in weights.items() if k in weights}
model.load_state_dict(weights)
model.cuda()
model.eval()
preds = []
with torch.no_grad():
for sample in tqdm(test_loader, total=len(test_loader)):
preds.append(model(sample['features'].cuda()).cpu().numpy().squeeze())
preds = np.concatenate(preds)
sample_sub['wind_speed'] = np.round(preds).astype(int)
sample_sub.to_csv('../data/21.14_resnest200e_adamw_heavy_fold_0_best_val_loss=67.265_sub.csv', index=False)
# Get validation loader and plot results
val_df = pd.read_csv(join_path(data_dir, 'fold0_val.csv'))
val_df['image_order'] = val_df['image_id'].apply(lambda x: int(str(x).split('_')[-1]))
val_df = val_df.sort_values(['image_order', 'storm_id'])
val_images = [ x+'.jpg' for x in val_df['image_id']]
val_dataset = WindDataset(val_images, image_dir, get_valid_transforms(256), 1)
val_loader = DataLoader(val_dataset, batch_size=16, shuffle=False)
preds_val = []
with torch.no_grad():
for sample in tqdm(val_loader, total=len(val_loader)):
preds_val.append(model(sample['features'].cuda()).cpu().numpy().squeeze())
val_df['wind_speed_predicted'] = np.concatenate(preds_val)
val_df['type'] = 'val'
val_df['wind_speed_diff'] = val_df['wind_speed_predicted'] - val_df['wind_speed']
rmse = mean_squared_error(val_df['wind_speed_predicted'], val_df['wind_speed'], squared=False)
print(f'rmse: {rmse}')
val_df.head()
sample_sub['storm_id'] = sample_sub['image_id'].apply(lambda x: x.split('_')[0])
sample_sub['image_order'] = sample_sub['image_id'].apply(lambda x: int(str(x).split('_')[-1]))
sample_sub['type'] = 'test'
sample_sub.head()
# # Let's make some diagnostic plots
merged_preds = pd.concat([
sample_sub[['image_id', 'wind_speed', 'storm_id', 'image_order', 'type']],
val_df[['image_id', 'wind_speed', 'storm_id', 'image_order', 'type']]]
)
merged_preds.head()
storms_val = set(val_df['storm_id'].value_counts().index.tolist())
storms_test = set(sample_sub['storm_id'].value_counts().index.tolist())
common_storms = list(storms_test.intersection(storms_val))
print(f'common storms val-test: {len(common_storms)}')
f, ax = plt.subplots(5, 5, figsize=(25, 25))
ax = ax.flatten()
for idx_storm in range(min(len(ax), len(common_storms))):
selected_storm = common_storms[idx_storm]
sns.lineplot(
y='wind_speed',
x='image_order',
style='type',
hue='storm_id',
ax=ax[idx_storm],
data=merged_preds.loc[merged_preds['storm_id']==selected_storm, :])
| notebooks/Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
iris = load_iris()
X, y = iris.data[:, :2], iris.target
print(X.shape)
print(y.shape)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=33)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
| codecheatsheet/split_training_and_test_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# NOTE: To get files for this assignment # 2, run inverted index 5.0 and get files from there.
# -
# # Overview
# In this assignment, you will use the index you created in Assignment 1 to rank documents
# and create a search engine. You will implement two different scoring functions and compare
# their results against a baseline ranking produced by expert analysts.
# # Running Queries
#
# For this assignment, you will need the following two files:
#
# <font color=red> </font> <font color=blue> topics.xml (\\sandata\xeon\Maryam Bashir\Information Retrieval\topics.xml) </font> contains the queries you will be testing.
#
# You should run the queries using the text stored in the <font color=green> query </font> elements. The <font color=green> description </font> elements are only there to clarify the information need which the query is trying to express</font> .
#
#
# <font color=red> </font> <font color=blue> corpus.qrel (\\sandata\xeon\Maryam Bashir\Information Retrieval\corpus.qrel)</font> contains the relevance grades from expert assessors. While these grades are not necessarily entirely correct (and defining correctness unambiguously is quite difficult), they are fairly reliable and we will treat them as being correct here.
#
# The format here is:
# <font color=green> topic </font> <font color=green> 0 </font> <font color=green> docid </font> <font color=green> grade </font>
#
# <font color=red> o </font> <font color=green> topic </font> is the ID of the query for which the document was assessed.
#
# <font color=red> o </font> <font color=green> 0 </font> is part of the format and can be ignored.
#
# <font color=red> o </font> <font color=green> docid </font> is the name of one of the documents which you have indexed.
#
# <font color=red> o </font> <font color=green> grade </font> is a value in the set <font color=blue> {-2, 0, 1, 2, 3, 4} </font>, where a higher value means that the document is more relevant to the query.
# The value -2 indicates a spam document, and 0 indicates a non-spam document which is completely non- relevant.
# Most queries do not have any document with a grade of 4, and many queries do not have any document with a grade of 3.
# This is a consequence of the specific meaning assigned to these grades here and the manner in which the documents were collected.
#
# This <font color=green> QREL </font> does not have assessments for every <font color=blue>(query, document) </font> pair. If an assessment is missing, we assume the correct grade for the pair is 0 (non-relevant).
#
# You will write a program which takes the name of a scoring function as a command line argument and which prints a ranked list of documents for all queries found in topics.xml using that scoring function.
#
# For example:
#
# <font color=red> $ </font> <font color=green> ./query.py --score TF-IDF </font>
#
# <font color=blue> 202 clueweb12-0000tw-13-04988 1 0.73 run1 </font>
#
# <font color=blue> 202 clueweb12-0000tw-13-04901 2 0.33 run1 </font>
#
# <font color=blue> 202 clueweb12-0000tw-13-04932 3 0.32 run1 </font> ...
#
# <font color=blue> 214 clueweb12-0000tw-13-05088 1 0.73 run1 </font>
#
# <font color=blue> 214 clueweb12-0000tw-13-05001 2 0.33 run1 </font>
#
# <font color=blue> 214 clueweb12-0000tw-13-05032 3 0.32 run1 </font> ...
#
# <font color=blue> 250 clueweb12-0000tw-13-05032 500 0.002 run1 </font>
#
#
# The output should have one row for each document which your program ranks for each query it runs.
# These lines should have the format:
#
# <font color=green> topic </font> <font color=green> docid </font> <font color=green> rank </font> <font color=green> score </font> <font color=green> run </font>
#
# <font color=red> </font> <font color=green> topic </font> is the ID of the query for which the document was ranked.
#
# <font color=red> </font> <font color=green> docid </font> is the document identifier.
#
# <font color=red> </font> <font color=green> rank </font> is the order in which to present the document to the user. The document with the highest score will be assigned a rank of 1, the second highest a rank of 2, and so on.
#
# <font color=red> </font> <font color=green> score </font> is the actual score the document obtained for that query.
#
# <font color=red> </font> <font color=green> run </font> is the name of the run. You can use any value here. It is meant to allow research teams to submit multiple runs for evaluation in competitions such as TREC.
from bs4 import BeautifulSoup
import re
from nltk.stem import PorterStemmer
from nltk.tokenize import word_tokenize
import random
import os
import operator
import xml.dom.minidom
import numpy as np
import math
import sys
#from sets import Set
#from html.parser import HTMLParser
def get_directory_path(mode):
"""
It takes only path of folder, no file name.
It only returns the folder which contain all the text file.
Argument:
mode -- string specifying input or output for directory
Returns:
dp -- directory path which contains all the txt files.
"""
if (mode == "input"):
dp = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/"
elif (mode == "output"):
dp = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/out/"
else:
raise ValueError('Unspecified mode for I/O.')
dp = None
return dp
# Function : read_stop_list
def read_text_in_list_form(file_path):
"""
This function takes the path of stop words file and reads it and returns a list of words.
Argument:
stop_file_path -- path should be like: "(dir) + file_Name.extension"
"/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/stoplist.txt".
Returns:
lst -- list of words containg all the stop_words.
"""
lst = [line.rstrip('\n') for line in open(file_path)]
return lst
# +
# stop_word_file = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/stoplist.txt"
# stop_words = read_text_in_list_form(stop_word_file)
# -
## Version 2.0 using dictionary
def xml_parser(file_name):
"""
It takes file name of xml file, which will contain the
queries with their unique topics-ids.
It returns the dictionary which contain topic-ids as keys and query of words values.
Argument:
file_name -- this will be "(dir)+topics.xml"
Returns:
queries -- directory with topic-ids as keys and query of words values.
"""
doc = xml.dom.minidom.parse(file_name)
qrys = doc.getElementsByTagName('query')
tpcs = doc.getElementsByTagName('topic')
queries = dict()
i = 0
for elem in qrys:
queries[tpcs[i].attributes['number'].value] = elem.firstChild.data
i = i + 1
return queries
# +
# file_name = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/topics.xml"
# qrys = xml_parser(file_name)
# #print(qrys[str(202)])
# print(qrys)
# -
# # Query Processing
# Before running any scoring function, you should process the text of the query in exactly the same way that you processed the text of a document. That is:
# 1. Split the query into tokens (it is most correct to use the regular expression, but for these queries it suffices to split on whitespace)
# 2. Convert all tokens to lowercase
# 3. Apply stop-wording to the query using the same list you used in assignment 1
# 4. Apply the same stemming algorithm to the query which you used in your indexer
def stem_words(tokenized_words_without_stop_words):
"""
This function takes in list of words which do not contain stop_words.
It uses the PorterStemmer() to reduce the words to their root word.
Argument:
removed_all_stop_words -- list of all words which do not have stop_words.
Returns:
stemmed_words -- list of words which are reduced to their origin word.
"""
ps = PorterStemmer()
stemmed_words = list()
for w in tokenized_words_without_stop_words:
stemmed_words.append(ps.stem(w))
stemmed_words.sort()
return stemmed_words
def query_processing(query_string):
"""
This function takes in a query string and does the pre-processing on it.
It will first load stop words from directory, then split the query into
single-single terms. Then remove stop words from it.
Argument:
query_string -- a string of query.
Returns:
stemmed_tokens -- tokens of query after being stemmed.
"""
path_to_stop_words = get_directory_path("input") +"stoplist.txt"
stop_words = read_text_in_list_form(path_to_stop_words)
#splited_query = list(re.split(query))
splited_query = list(query_string.split())
#splited_query.lower()
cleaned_tokens_from_stop_words = list(set(splited_query) - set(stop_words))
stemmed_tokens = stem_words(cleaned_tokens_from_stop_words)
return stemmed_tokens
# +
# #print(type(qrys[str(202)]))
# x = query_processing(qrys[str(202)])
# print((x))
# print(len(x))
# -
# # Scoring Function 1: Okapi BM25
# Implement BM25 scores. This should use the following scoring function for document d and query q:
#
# Where k1,k2, and b are constants. For start, you can use the values suggested in the lecture on BM25 (k1 = 1.2, k2 varies from 0 to 1000, b = 0.75). Feel free to experiment with different values for these
# constants to learn their effect and try to improve performance.
def get_all_documents_length(docid_hashed_file):
"""
It takes file name of a file which will contain doc-ids and
adjacent to it will be document name and length of each document.
It only returns the dictionary which will have unique DOC_IDS as KEYS
and documents length as values.
Argument:
docid_hashed_file -- this will be "(dir)+docid_hashed.txt"
Returns:
doc_lengths -- is a dictionary as {doc-id : length_of_doc}.
"""
doc_lengths = dict()
file = open(docid_hashed_file, 'r' , encoding = "utf-8")
for each_line in file:
x = each_line.split()
doc_lengths[x[0]] = x[2]
return doc_lengths
# +
# docid_hashed_file = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/docid_hashed.txt"
# all_doc_lengths = get_all_documents_length(docid_hashed_file)
# #c = int(all_doc_lengths[str(3058)])
# #print(int(all_doc_lengths[str(3058)]))
# print((all_doc_lengths))
# -
def get_all_documents_name(docid_hashed_file):
"""
It takes file name of a file which will contain doc-ids and
adjacent to it will be document name and length of each document.
It only returns the dictionary which will have unique DOC_IDS as KEYS
and document names as values.
Argument:
docid_hashed_file -- this will be "(dir)+docid_hashed.txt"
Returns:
doc_name-- it is a dictionary as {doc-id : doc-name}.
"""
doc_names = dict()
i = 0
file = open(docid_hashed_file,'r',encoding = "utf-8")
for each_line in file:
x = each_line.split()
doc_names[x[0]] = x[1]
return doc_names
# +
# docid_hashed_file = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/docid_hashed.txt"
# all_doc_names = get_all_documents_name(docid_hashed_file)
# print((all_doc_names))
# -
def get_all_vocablury(file_name):
"""
It takes file name of term-ids which will contain termids and
adjacent to termids will be terms.
It only returns the dictionary which will have unique TERMS as KEYS
and TERM_IDS will be the VALUES..
Argument:
file_name -- this will be "(dir)+termid_hashed.txt"
Returns:
vocablury -- it is a dictionary as {terms: term-ids}.
"""
vocablury = dict()
file = open(file_name,'r',encoding = "utf-8")
for each_line in file:
x = each_line.split()
#Interesting: making term as key and term_id as value (;
vocablury[x[1]] = x[0]
return vocablury
# +
# voc_file = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/termid_hashed.txt"
# vocablury = get_all_vocablury(voc_file)
# #print(vocablury)
# -
#### version 2.0
def get_document_postings(document_postings_file):
"""
It takes file name of a file which will contain termids and
adjacent to it will be list of documents in which this termid appears.
It only returns the dictionary which will have unique TERMS_IDS as KEYS
and document list as values.
Argument:
document_postings_file -- this will be "(dir)+document_postings.txt"
Returns:
termid_with_doc_postings -- it is a dictionary as {terms-ids: list(documents)}.
"""
termid_with_doc_postings = dict()
file = open(document_postings_file,'r',encoding = "utf-8")
for each_line in file:
x = (re.split("\n",each_line))
#print("x = " , (x), " len of x = " ,len(x))
y = (re.split("\t", x[0]))
#print("y = " , (y), " len of y = " ,len(y))
temp_str = y[(len(y)-1)]
temp_str = temp_str.strip("[]")
temp_str = temp_str.replace(" ", "")
lst = list()
lst = temp_str.split(",")
current_term_id = y[0]
# create a key in dict_with_docid_and_its_positions
termid_with_doc_postings[current_term_id] = lst
return termid_with_doc_postings
# +
# doc_postings_file = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/document_postings.txt"
# dict_termid_with_docs_postings = get_document_postings(doc_postings_file)
# #print(dict_termid_with_docs_postings)
# -
# version 2.0
def get_inverted_index(term_index_hashed_file):
"""
It takes file name of a inverted index file which will contain
term-ids and all of it's documents with all of it's positions.
It returns the Nested dictionary which will have unique term-id
as key to outer dictionary and on a single term-id there may
have multiple documents, these documents will be used a keys to
inner dictionary, and on a single-document-id there may have
multiple positions on which that term appeared.
Argument:
term_index_hashed_file -- this will be file name as "(dir)+docid_hashed.txt"
Returns:
nested_dict_with_termid_and_its_docs_and_occurance -- it is a dictionary as {term-id : { doc-id : positions }}.
"""
nested_dict_with_termid_and_its_docs_and_occurance = dict()
doc_id_with_positions = dict()
file = open(term_index_hashed_file, 'r' , encoding = "utf-8")
length = 0
current_term_id = 0
for each_line in file:
x = (re.split("\n",each_line))
y = (re.split("\t", x[0]))
if y[0] == '':
y = y[1:]
# It means a New term_id aya hai, tou dict ki new key bnani
if (len(y) == 4):
# pick up last element, which will be list of positions in one document.
# and filter it with strip(), replace() methods.
temp_str = y[(len(y)-1)]
temp_str = temp_str.strip("[]")
temp_str = temp_str.replace(" ", "")
lst = list()
lst = temp_str.split(",")
# reset length variable
doc_id_with_positions = dict()
length = 0
# new term_id now found
current_term_id = y[0]
current_document_id = y[1]
# create a key in dict_with_docid_and_its_positions
doc_id_with_positions[current_document_id] = lst
#length = len(lst) #len((y[(len(y)-1)]))
nested_dict_with_termid_and_its_docs_and_occurance[current_term_id] = doc_id_with_positions
elif (len(y) == 3):
# pick up last element, which will be list of positions in one document.
# and filter it with strip(), replace() methods.
temp_str = y[(len(y)-1)]
temp_str = temp_str.strip("[]")
temp_str = temp_str.replace(" ", "")
lst = list()
lst = temp_str.split(",")
current_document_id = y[0]
doc_id_with_positions[current_document_id] = lst
nested_dict_with_termid_and_its_docs_and_occurance.update({current_term_id:doc_id_with_positions})
else:
print("\nI Don't know what to do.\n")
return nested_dict_with_termid_and_its_docs_and_occurance
# +
# hashed_term_id = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/term_index_hashed.txt"
# dict_term_id_with_frequencies = get_term_frequency(hashed_term_id)
# print((dict_term_id_with_frequencies[str(583007)]))
# #print((dict_term_id_with_frequencies[str(583007)][str(5256)]))
# #print(len(dict_term_id_with_frequencies[str(583007)][str(5256)]))
# #print(dict_term_id_with_frequencies)
# -
### build version 2.3 : Final version
# Changes Made:
# finally removed unnecessary comments & print statements, and added proper comments &
# replaced doc_id with doc_names as key of docs_score_for_each_query()
def calculate_okapi_bm25(parameters):
k_1 = parameters['k1']
k_2 = parameters['k2']
b = parameters['b']
D = parameters['D']
input_directory = get_directory_path("input")
output_directory = get_directory_path("output")
txt_extension = ".txt"
xml_extension = ".xml"
queries_file_name = input_directory + "topics" + xml_extension
queries_dict = xml_parser(queries_file_name)
doc_info_file = input_directory+ "docid_hashed" + txt_extension
all_doc_lengths = get_all_documents_length(doc_info_file)
docid_hashed_file = input_directory + "docid_hashed" + txt_extension
all_doc_names = get_all_documents_name(docid_hashed_file)
voc_file = input_directory + "termid_hashed" + txt_extension
vocablury = get_all_vocablury(voc_file)
doc_postings_file = input_directory + "document_postings" + txt_extension
dict_termid_with_docs_postings = get_document_postings(doc_postings_file)
inverted_index_file = input_directory + "term_index_hashed" + txt_extension
hashed_ii = get_inverted_index(inverted_index_file)
#hashed_ii
avg_length = 0
for doc_id, length in all_doc_lengths.items():
avg_length += int(length)
avg_length /= len(all_doc_lengths)
scores_dictionary = dict()
# Will run for number of times of queries in topics.xml
for query_id, query in queries_dict.items(): # run for 10 times
# Split one query in terms
splitted_query = query_processing(query)
score_of_each_term_for_single_doc = 0
# Reset : docs_score_for_each_query.
docs_score_for_each_query = dict()
#scores_dictionary[query_id] = dict()
# For loop for each term in single queries,
# i.e. if there is 3 word query, it will run for 3 times.
for i in range(0,len(splitted_query)):
# score_of_each_term_for_single_doc = 0
# Check If this splitted term (from query) exists in my vocablury.
if splitted_query[i] in vocablury: # if that term exists
# If YES, then get term_id of this splitted term (from query).
# Now, get value = term_id by passing term as key.
term_id = vocablury[splitted_query[i]]
# Get list of documents in which this term exists
list_of_all_docs_in_which_term_exists = dict_termid_with_docs_postings[term_id]
# Get it's document_frequency, i.e. In how many docs it is present.
df_i = len(list_of_all_docs_in_which_term_exists)
# Now, run this loop for all docs in which it is present.
# i.e if it is present in 3 docs, it will run for 3
for j in range(0,len(list_of_all_docs_in_which_term_exists)):
# Now, pick one by one doc_id, and compute score.
doc_id = list_of_all_docs_in_which_term_exists[j]
doc_name = all_doc_names[doc_id]
# Check IF that doc_id is present in my doc_postings file
if doc_id in all_doc_lengths:
# Get this term's frquency in this document
tf_d_i = len(hashed_ii[str(term_id)][doc_id])
tf_q_i = 1
length_of_doc_id = int(all_doc_lengths[doc_id])
capital_K = k_1 * ((1-b) + (b * (length_of_doc_id/avg_length)))
a = float(D + 0.5)
b = float(df_i + 0.5)
c = float(math.log(a/b))
d = float((1+k_1) * tf_d_i)
e = float(capital_K + tf_d_i)
f = float((1+k_2) * tf_q_i)
g = float(k_2+tf_q_i)
score_of_each_term_for_single_doc = c * (d/e) * (f/g)
# Check IF already a term might have calculated score for this document (for same query).
# Or we can say, multiple term words of single query might present in same document.
# If YES: Else NO
if doc_name in docs_score_for_each_query:
prev_score = docs_score_for_each_query[doc_name]
new_score = prev_score + score_of_each_term_for_single_doc
docs_score_for_each_query[doc_name] = new_score
sorted_docs_score_for_each_query = sorted(docs_score_for_each_query.items(), key=operator.itemgetter(1), reverse=True)
scores_dictionary[query_id] = sorted_docs_score_for_each_query
# Or maybe we found new document. Let's create a new key of doc_id on same query_id
else:
docs_score_for_each_query[doc_name] = dict()
docs_score_for_each_query[doc_name] = score_of_each_term_for_single_doc
sorted_docs_score_for_each_query = sorted(docs_score_for_each_query.items(), key=operator.itemgetter(1),reverse=True)
scores_dictionary[query_id] = sorted_docs_score_for_each_query
#scores_dictionary[query_id] = dict()
scores_dictionary[query_id] = sorted_docs_score_for_each_query
# Or maybe there is a document which is not in my possession.
else:
docs_score_for_each_query[doc_name] = 0
scores_dictionary[query_id] = docs_score_for_each_query
# If this term is not in my Vocablury
else:
print("Terms of Queries which are not in my collection = ", splitted_query[i])
return scores_dictionary
# +
parameters = dict()
parameters['k1'] = 1.2
parameters['k2'] = 500
parameters['b'] = 0.75
parameters['D'] = 17
okapi_bmi25_score = calculate_okapi_bm25(parameters)
#print(okapi_bmi25_score[str(214)])
#a = (okapi_bmi25_score[str(214)])[0]
#print(a[1])
#print((okapi_bmi25_score[str(214)])[1])
# -
## version v3.0
def dirichlet_smoothing():
input_directory = get_directory_path("input")
output_directory = get_directory_path("output")
txt_extension = ".txt"
xml_extension = ".xml"
queries_file_name = input_directory + "topics" + xml_extension
queries_dict = xml_parser(queries_file_name)
doc_info_file = input_directory+ "docid_hashed" + txt_extension
all_doc_lengths = get_all_documents_length(doc_info_file)
docid_hashed_file = input_directory + "docid_hashed" + txt_extension
all_doc_names = get_all_documents_name(docid_hashed_file)
voc_file = input_directory + "termid_hashed" + txt_extension
vocablury = get_all_vocablury(voc_file)
doc_postings_file = input_directory + "document_postings" + txt_extension
dict_termid_with_docs_postings = get_document_postings(doc_postings_file)
inverted_index_file = input_directory + "term_index_hashed" + txt_extension
hashed_ii = get_inverted_index(inverted_index_file)
mu = 0
total_length_of_collection = 0
for doc_id, length in all_doc_lengths.items():
total_length_of_collection += int(length)
mu = total_length_of_collection/len(all_doc_lengths)
scores_dictionary = dict()
# Will run for number of times of queries in topics.xml
for query_id, query in queries_dict.items(): # run for 10 times
# Split one query in terms
splitted_query = query_processing(query)
score_of_each_term_for_single_doc = 0
# Reset : docs_score_for_each_query.
docs_score_for_each_query = dict()
# For loop for each term in single queries,
# i.e. if there is 3 word query, it will run for 3 times.
for i in range(0,len(splitted_query)):
# Check If this splitted term (from query) exists in my vocablury.
if splitted_query[i] in vocablury: # if that term exists
# If YES, then get term_id of this splitted term (from query).
# Now, get value = term_id by passing term as key.
term_id = vocablury[splitted_query[i]]
# Get list of documents in which this term exists.
list_of_all_docs_in_which_term_exists = dict_termid_with_docs_postings[(term_id)]
# Count the number of times each word occurs in Corpora, divide by total length.
# Basically add-up all lengths of documents.
sum_of_term_in_whole_corpora = 0
for d_idx, positinos in hashed_ii[term_id].items():
sum_of_term_in_whole_corpora += len(positinos) # len(hashed_ii[term_id][positinos]) #
prob_of_term_occuring_in_whole_corpora = sum_of_term_in_whole_corpora/total_length_of_collection
# Now, run this loop for all docs in which it is present.
# i.e if it is present in 3 docs, it will run for 3.
for j in range(0,len(list_of_all_docs_in_which_term_exists)):
# Now, pick one by one doc_id, and compute score.
doc_id = list_of_all_docs_in_which_term_exists[j]
doc_name = all_doc_names[doc_id]
# Check IF that doc_id is present in my doc_postings file.
if doc_id in all_doc_lengths:
N = int(all_doc_lengths[(doc_id)]) # doc length
lamdba = N/(N+mu)
one_minus_lamdba = 1 - lamdba
# Count the number of times word occurs in document, divide by document length.
count_of_term_in_single_doc = len(hashed_ii[term_id][doc_id])
prob_occuring_in_single_doc = count_of_term_in_single_doc / N
score_of_each_term_for_single_doc = (lamdba * prob_occuring_in_single_doc) + (one_minus_lamdba * prob_of_term_occuring_in_whole_corpora)
# Check IF already a term might have calculated score for this document (for same query).
# Or we can say, multiple term words of single query might present in same document.
# If YES: Else NO
if doc_name in docs_score_for_each_query:
prev_score = docs_score_for_each_query[doc_name]
new_score = prev_score + score_of_each_term_for_single_doc
docs_score_for_each_query.update({doc_name : new_score})
sorted_docs_score_for_each_query = sorted(docs_score_for_each_query.items(), key=operator.itemgetter(1), reverse=True)
scores_dictionary[query_id] = sorted_docs_score_for_each_query
# Or maybe we found new document. Let's create a new key of doc_id on same query_id.
else:
docs_score_for_each_query[doc_name] = dict()
docs_score_for_each_query[doc_name] = score_of_each_term_for_single_doc
sorted_docs_score_for_each_query = sorted(docs_score_for_each_query.items(), key=operator.itemgetter(1),reverse=True)
scores_dictionary[query_id] = sorted_docs_score_for_each_query
#scores_dictionary[query_id] = dict()
scores_dictionary[query_id] = sorted_docs_score_for_each_query
# Or maybe there is a document which is not in my possession.
else:
docs_score_for_each_query[doc_name] = 0
scores_dictionary[query_id] = docs_score_for_each_query
# If this term is not in my Vocablury.
else:
print("Terms of Queries which are not in my collection = ", splitted_query[i])
return scores_dictionary
dirichlet_score = dirichlet_smoothing()
#print(dirichlet_score)
def save_scores(output_file_name, scores_dictionary):
fh = open(output_file_name, "a+")
for query_id , doc_score_list in scores_dictionary.items():
# term_id
q_id = str(query_id)
i = 1
for j in range(0, len(doc_score_list)):
doc_name = doc_score_list[j][0]
score = doc_score_list[j][1]
lne = q_id + "\t"+ doc_name + "\t" + str(i)+"\t"+ str(score) + (" run 1")
fh.write(lne)
fh.write('\n')
i+=1
fh.close()
# +
okapi_output_file = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/out/okapi_scores.txt"
### okapi_bmi25_score -> getting from okapi_bmi25_score = calculate_okapi_bm25(parameters)
save_scores(output_file, okapi_bmi25_score)
#dirichlet_output_file = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/out/dirichlet_scores.txt"
#### dirichlet_score -> getting from dirichlet_score = dirichlet_smoothing()
#save_scores(output_file, dirichlet_score)
# -
def print_socre(scores_dictionary):
for query_id , doc_score_list in scores_dictionary.items():
# term_id
q_id = str(query_id)
i = 1
for j in range(0, len(doc_score_list)):
doc_name = doc_score_list[j][0]
score = doc_score_list[j][1]
lne = q_id + "\t"+ doc_name + "\t" + str(i)+"\t"+ str(score) + (" run 1")
print(lne)
i+=1
# +
print("\nOkapi BM-25 Score :\n")
print_socre(okapi_bmi25_score)
#print("\nDirichlet Score :\n")
#print_socre(dirichlet_score)
# -
# # Evaluation
#
# To evaluate your results, we will write a program that computes mean average precision of the
# rank list of documents for different queries. The input to program will be the <font color=blue> qrel file
# (relevance judgments) </font> and scoring file that has rank list of documents.
#
# The output should be following measures:
#
# <font color=red> </font> <font color=green> P@5 </font>
#
# <font color=red> </font> <font color=green> P@10 </font>
#
# <font color=red> </font> <font color=green> P@20 </font>
#
# <font color=red> </font> <font color=green> P@30 </font>
#
# <font color=red> </font> <font color=green> MAP </font>
#
# These measures should be computed for each query. Average for all queries should also be computed.
#
def qrel_reader_and_parser(qrel_file):
nested_dict_with_topics_and_its_docs_and_grades = dict()
doc_id_with_grades = dict()
file = open(qrel_file, 'r' , encoding = "utf-8")
length = 0
current_topic_id = 0
for each_line in file:
x = (re.split("\n",each_line))
y = (re.split(" ", x[0]))
#print(y)
current_topic_id = y[0]
# If current_topic already is already in my dict
if current_topic_id in nested_dict_with_topics_and_its_docs_and_grades:
current_doc_name = y[2]
doc_id_with_grades[current_doc_name] = int()
val = 0
if int(y[3]) > 0:
val = 1
# name_of_doc # grade
doc_id_with_grades[current_doc_name] = val
nested_dict_with_topics_and_its_docs_and_grades.update({current_topic_id : doc_id_with_grades})
# Current_topic pehle se mujood naaii hai
else:
# Now new topic found, thus reset.
nested_dict_with_topics_and_its_docs_and_grades[current_topic_id] = dict()
doc_id_with_grades = dict()
current_doc_name = y[2]
doc_id_with_grades[current_doc_name] = int()
val = 0
if int(y[3]) > 0:
val = 1
# name_of_doc # grade
doc_id_with_grades[current_doc_name] = val
nested_dict_with_topics_and_its_docs_and_grades.update({current_topic_id : doc_id_with_grades})
return nested_dict_with_topics_and_its_docs_and_grades
qrel_file_path = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/relevance judgements.qrel"
dict_for_qrel = qrel_reader_and_parser(qrel_file_path)
#print(dict_for_qrel)
def calculate_precison(qrel_file_path, precision_mode, score_algorithm):
dict_for_qrel = qrel_reader_and_parser(qrel_file_path)
if (score_algorithm == "dirichlet"):
dirichlet_score = dirichlet_smoothing()
elif (score_algorithm == "bm25"):
parameters = dict()
parameters['k1'] = 1.2
parameters['k2'] = 500
parameters['b'] = 0.75
parameters['D'] = 17
# okapi_bmi25_score = calculate_okapi_bm25(parameters)
# Naming it as dirichlet_score, because then I would have to change
# dictionary name dirichley_socre everywhere in code.
dirichlet_score = calculate_okapi_bm25(parameters)
else:
raise ValueError('Unspecified Scoring algorithm.')
sys.exit(0)
if precision_mode == "p5":
dict_for_precision = dict()
precision = 0
relevant = 0
#retrieved2 = 5
for query_id, zipi in dirichlet_score.items():
count = 0
retrieved = len(dirichlet_score[query_id])
for i in range(0,len(zipi)):
#print(zipi[i][0]) = Doc_name
#print(zipi[i][0]) = calculated_score
doc_name = zipi[i][0]
if(count == 5):
break
# IF THAT FOLDER EXISTS IN QREL AND HAS SCORE 1
if doc_name in dict_for_qrel[query_id] and dict_for_qrel[query_id][doc_name] > 0:
relevant+=1
count+=1
#retrieved2+=1
precision = float(relevant/retrieved)
dict_for_precision[query_id] = precision
elif precision_mode == "p10":
dict_for_precision = dict()
precision = 0
relevant = 0
#retrieved2 = 10
for query_id, zipi in dirichlet_score.items():
count = 0
retrieved = len(dirichlet_score[query_id])
for i in range(0,len(zipi)):
#print(zipi[i][0]) = Doc_name
#print(zipi[i][0]) = calculated_score
doc_name = zipi[i][0]
if(count == 10):
break
# IF THAT FOLDER EXISTS IN QREL AND HAS SCORE 1
if doc_name in dict_for_qrel[query_id] and dict_for_qrel[query_id][doc_name] > 0:
relevant+=1
count+=1
#retrieved2+=1
precision = float(relevant/retrieved)
dict_for_precision[query_id] = precision
elif precision_mode == "p20":
dict_for_precision = dict()
precision = 0
relevant = 0
#retrieved2 = 20
for query_id, zipi in dirichlet_score.items():
count = 0
retrieved = len(dirichlet_score[query_id])
for i in range(0,len(zipi)):
#print(zipi[i][0]) = Doc_name
#print(zipi[i][0]) = calculated_score
doc_name = zipi[i][0]
if(count == 20):
break
# IF THAT FOLDER EXISTS IN QREL AND HAS SCORE 1
if doc_name in dict_for_qrel[query_id] and dict_for_qrel[query_id][doc_name] > 0:
relevant+=1
count+=1
#retrieved2+=1
precision = float(relevant/retrieved)
dict_for_precision[query_id] = precision
elif precision_mode == "p30":
dict_for_precision = dict()
precision = 0
relevant = 0
#retrieved2 = 30
for query_id, zipi in dirichlet_score.items():
count = 0
retrieved = len(dirichlet_score[query_id])
for i in range(0,len(zipi)):
#print(zipi[i][0]) = Doc_name
#print(zipi[i][0]) = calculated_score
doc_name = zipi[i][0]
if(count == 30):
break
# IF THAT FOLDER EXISTS IN QREL AND HAS SCORE 1
if doc_name in dict_for_qrel[query_id] and dict_for_qrel[query_id][doc_name] > 0:
relevant+=1
count+=1
#retrieved2+=1
precision = float(relevant/retrieved)
dict_for_precision[query_id] = precision
elif precision_mode == "map":
print("TODO")
else:
raise ValueError('Unspecified mode for Precision.')
sys.exit(0)
print(dict_for_precision)
# +
qrel_file_path = "/Users/imbilalbutt/Documents/Semesters/Semester 9/Information Retrieval/Assignment/hw2/input/relevance judgements.qrel"
i_mode = "p30"
#score_algorithm = "dirichlet"
score_algorithm = "bm25"
calculate_precison(qrel_file_path, i_mode, score_algorithm)
# -
D = dict()
ls1 = ['1','2','3']
ls2 = ['1','2','3']
s = list(zip(ls1,ls2))
len(s)
print(s[0][0])
| IR Assignment 2 v3.1 .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import modcell as mods
# +
import sys
sys.path.append('../module_notebook')
# -
# if source_info is True, modcell export source code information to the output file, which is useful for debug
source_info = True
# ### Compile hello and fizzbuzz
mod_useless = mods.ModCell()
mod_useless._import('hello')
mod_useless._import('fizzbuzz')
with open('../module/useless.py', mode='w') as f:
mod_useless.compile(out=f, source_info = source_info)
# ### Compile game and animmal (with sellective import)
mod_funny = mods.ModCell()
mod_funny._import('game')
mod_funny._import('animal', tag='Mammals')
mod_funny._import('animal', tag='Birds')
with open('../module/funny.py', mode='w') as f:
mod_funny.compile(out=f, source_info = source_info)
| notebooks/compile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Name
# Submitting a Cloud Machine Learning Engine training job as a pipeline step
#
# # Label
# GCP, Cloud ML Engine, Machine Learning, pipeline, component, Kubeflow, Kubeflow Pipeline
#
# # Summary
# A Kubeflow Pipeline component to submit a Cloud ML Engine training job as a step in a pipeline.
#
# # Details
# ## Intended use
# Use this component to submit a training job to Cloud ML Engine from a Kubeflow Pipeline.
#
# ## Runtime arguments
# | Argument | Description | Optional | Data type | Accepted values | Default |
# |:------------------|:------------------|:----------|:--------------|:-----------------|:-------------|
# | project_id | The ID of the Google Cloud Platform (GCP) project of the job. | No | GCPProjectID | | |
# | python_module | The name of the Python module to run after installing the training program. | Yes | String | | None |
# | package_uris | The Cloud Storage location of the packages that contain the training program and any additional dependencies. The maximum number of package URIs is 100. | Yes | List | | None |
# | region | The Compute Engine region in which the training job is run. | Yes | GCPRegion | | us-central1 |
# | args | The command line arguments to pass to the training program. | Yes | List | | None |
# | job_dir | A Cloud Storage path in which to store the training outputs and other data needed for training. This path is passed to your TensorFlow program as the `job-dir` command-line argument. The benefit of specifying this field is that Cloud ML validates the path for use in training. | Yes | GCSPath | | None |
# | python_version | The version of Python used in training. If it is not set, the default version is 2.7. Python 3.5 is available when the runtime version is set to 1.4 and above. | Yes | String | | None |
# | runtime_version | The runtime version of Cloud ML Engine to use for training. If it is not set, Cloud ML Engine uses the default. | Yes | String | | 1 |
# | master_image_uri | The Docker image to run on the master replica. This image must be in Container Registry. | Yes | GCRPath | | None |
# | worker_image_uri | The Docker image to run on the worker replica. This image must be in Container Registry. | Yes | GCRPath | | None |
# | training_input | The input parameters to create a training job. | Yes | Dict | [TrainingInput](https://cloud.google.com/ml-engine/reference/rest/v1/projects.jobs#TrainingInput) | None |
# | job_id_prefix | The prefix of the job ID that is generated. | Yes | String | | None |
# | job_id | The ID of the job to create, takes precedence over generated job id if set. | Yes | String | - | None |
# | wait_interval | The number of seconds to wait between API calls to get the status of the job. | Yes | Integer | | 30 |
#
#
#
# ## Input data schema
#
# The component accepts two types of inputs:
# * A list of Python packages from Cloud Storage.
# * You can manually build a Python package and upload it to Cloud Storage by following this [guide](https://cloud.google.com/ml-engine/docs/tensorflow/packaging-trainer#manual-build).
# * A Docker container from Container Registry.
# * Follow this [guide](https://cloud.google.com/ml-engine/docs/using-containers) to publish and use a Docker container with this component.
#
# ## Output
# | Name | Description | Type |
# |:------- |:---- | :--- |
# | job_id | The ID of the created job. | String |
# | job_dir | The Cloud Storage path that contains the trained model output files. | GCSPath |
#
#
# ## Cautions & requirements
#
# To use the component, you must:
#
# * Set up a cloud environment by following this [guide](https://cloud.google.com/ml-engine/docs/tensorflow/getting-started-training-prediction#setup).
# * The component can authenticate to GCP. Refer to [Authenticating Pipelines to GCP](https://www.kubeflow.org/docs/gke/authentication-pipelines/) for details.
# * Grant the following access to the Kubeflow user service account:
# * Read access to the Cloud Storage buckets which contain the input data, packages, or Docker images.
# * Write access to the Cloud Storage bucket of the output directory.
#
# ## Detailed description
#
# The component builds the [TrainingInput](https://cloud.google.com/ml-engine/reference/rest/v1/projects.jobs#TrainingInput) payload and submits a job via the [Cloud ML Engine REST API](https://cloud.google.com/ml-engine/reference/rest/v1/projects.jobs).
#
# The steps to use the component in a pipeline are:
#
#
# 1. Install the Kubeflow Pipeline SDK:
#
# +
# %%capture --no-stderr
# !pip3 install kfp --upgrade
# -
# 2. Load the component using KFP SDK
# +
import kfp.components as comp
mlengine_train_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/pipelines/1.1.2/components/gcp/ml_engine/train/component.yaml')
help(mlengine_train_op)
# -
# ### Sample
# Note: The following sample code works in an IPython notebook or directly in Python code.
#
# In this sample, you use the code from the [census estimator sample](https://github.com/GoogleCloudPlatform/cloudml-samples/tree/master/census/estimator) to train a model in Cloud ML Engine. To upload the code to Cloud ML Engine, package the Python code and upload it to a Cloud Storage bucket.
#
# Note: You must have read and write permissions on the bucket that you use as the working directory.
# #### Set sample parameters
# + tags=["parameters"]
# Required Parameters
PROJECT_ID = '<Please put your project ID here>'
GCS_WORKING_DIR = 'gs://<Please put your GCS path here>' # No ending slash
# -
# Optional Parameters
EXPERIMENT_NAME = 'CLOUDML - Train'
TRAINER_GCS_PATH = GCS_WORKING_DIR + '/train/trainer.tar.gz'
OUTPUT_GCS_PATH = GCS_WORKING_DIR + '/train/output/'
# #### Clean up the working directory
# %%capture --no-stderr
# !gsutil rm -r $GCS_WORKING_DIR
# #### Download the sample trainer code to local
# %%capture --no-stderr
# !wget https://github.com/GoogleCloudPlatform/cloudml-samples/archive/master.zip
# !unzip master.zip
# #### Package code and upload the package to Cloud Storage
# %%capture --no-stderr
# %%bash -s "$TRAINER_GCS_PATH"
pushd ./cloudml-samples-master/census/estimator/
python setup.py sdist
gsutil cp dist/preprocessing-1.0.tar.gz $1
popd
# rm -fr ./cloudml-samples-master/ ./master.zip ./dist
# #### Example pipeline that uses the component
import kfp.dsl as dsl
import json
@dsl.pipeline(
name='CloudML training pipeline',
description='CloudML training pipeline'
)
def pipeline(
project_id = PROJECT_ID,
python_module = 'trainer.task',
package_uris = json.dumps([TRAINER_GCS_PATH]),
region = 'us-central1',
args = json.dumps([
'--train-files', 'gs://cloud-samples-data/ml-engine/census/data/adult.data.csv',
'--eval-files', 'gs://cloud-samples-data/ml-engine/census/data/adult.test.csv',
'--train-steps', '1000',
'--eval-steps', '100',
'--verbosity', 'DEBUG'
]),
job_dir = OUTPUT_GCS_PATH,
python_version = '',
runtime_version = '1.10',
master_image_uri = '',
worker_image_uri = '',
training_input = '',
job_id_prefix = '',
job_id = '',
wait_interval = '30'):
task = mlengine_train_op(
project_id=project_id,
python_module=python_module,
package_uris=package_uris,
region=region,
args=args,
job_dir=job_dir,
python_version=python_version,
runtime_version=runtime_version,
master_image_uri=master_image_uri,
worker_image_uri=worker_image_uri,
training_input=training_input,
job_id_prefix=job_id_prefix,
job_id=job_id,
wait_interval=wait_interval)
# #### Compile the pipeline
pipeline_func = pipeline
pipeline_filename = pipeline_func.__name__ + '.zip'
import kfp.compiler as compiler
compiler.Compiler().compile(pipeline_func, pipeline_filename)
# #### Submit the pipeline for execution
# +
#Specify pipeline argument values
arguments = {}
#Get or create an experiment and submit a pipeline run
import kfp
client = kfp.Client()
experiment = client.create_experiment(EXPERIMENT_NAME)
#Submit a pipeline run
run_name = pipeline_func.__name__ + ' run'
run_result = client.run_pipeline(experiment.id, run_name, pipeline_filename, arguments)
# -
# #### Inspect the results
#
# Use the following command to inspect the contents in the output directory:
# !gsutil ls $OUTPUT_GCS_PATH
# ## References
# * [Component python code](https://github.com/kubeflow/pipelines/blob/master/components/gcp/container/component_sdk/python/kfp_component/google/ml_engine/_train.py)
# * [Component docker file](https://github.com/kubeflow/pipelines/blob/master/components/gcp/container/Dockerfile)
# * [Sample notebook](https://github.com/kubeflow/pipelines/blob/master/components/gcp/ml_engine/train/sample.ipynb)
# * [Cloud Machine Learning Engine job REST API](https://cloud.google.com/ml-engine/reference/rest/v1/projects.jobs)
#
# ## License
# By deploying or using this software you agree to comply with the [AI Hub Terms of Service](https://aihub.cloud.google.com/u/0/aihub-tos) and the [Google APIs Terms of Service](https://developers.google.com/terms/). To the extent of a direct conflict of terms, the AI Hub Terms of Service will control.
| components/gcp/ml_engine/train/sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # 3.13 Self Check
# **2. _(IPython Session)_** What happens if you try to print the items in `range(10,` `0,` `2)`?
for number in range(10, 0, 2):
print(number, end=' ')
# **3. _(IPython Session)_** Use a `for` statement, `range` and `print` to display on one line the sequence of values `99` `88` `77` `66` `55` `44` `33` `22` `11` `0`, each separated by one space.
for number in range(99, -1, -11):
print(number, end=' ')
# **4. _(IPython Session)_** Use `for` and `range` to sum the even integers from 2 through 100, then display the sum.
total = 0
for number in range(2, 101, 2):
total += number
total
##########################################################################
# (C) Copyright 2019 by Deitel & Associates, Inc. and #
# Pearson Education, Inc. All Rights Reserved. #
# #
# DISCLAIMER: The authors and publisher of this book have used their #
# best efforts in preparing the book. These efforts include the #
# development, research, and testing of the theories and programs #
# to determine their effectiveness. The authors and publisher make #
# no warranty of any kind, expressed or implied, with regard to these #
# programs or to the documentation contained in these books. The authors #
# and publisher shall not be liable in any event for incidental or #
# consequential damages in connection with, or arising out of, the #
# furnishing, performance, or use of these programs. #
##########################################################################
| examples/ch03/snippets_ipynb/03_13selfcheck.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="MlOO1FiPSeGC"
# # Import Libraries
# + id="lF2vmbIpFa1L"
# Lib & Dependencies
import pandas as pd
import numpy as np
from sklearn import model_selection
from sklearn.model_selection._split import _BaseKFold, indexable, _num_samples
from sklearn import metrics
from scipy.stats import spearmanr
import xgboost as xgb
# + [markdown] id="kR5PXochSiGa"
# # Load data
# + id="7uNfyX3AFjCI"
train_datalink_X = 'https://tournament.datacrunch.com/data/X_train.csv'
train_datalink_y = 'https://tournament.datacrunch.com/data/y_train.csv'
hackathon_data_link = 'https://tournament.datacrunch.com/data/X_test.csv'
# + id="CrHm3uYAFat4"
train_data = pd.read_csv(train_datalink_X)
test_data = pd.read_csv(hackathon_data_link)
train_targets = pd.read_csv(train_datalink_y)
# + id="HFUCQugROCBg"
train = train_data.merge(train_targets, left_index=True, right_index=True, how='inner')
# + colab={"base_uri": "https://localhost:8080/", "height": 215} id="8aprBdSuFakX" outputId="0a3b2d28-ed8d-428e-f3f4-a56e152d90e1"
train.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="ruCBxNygF4hd" outputId="4ff90b3e-40dc-4609-fe47-5ac364d2a9a2"
test_data.head()
# + [markdown] id="SiXkj9LMSniZ"
# # Set features
# + id="X8lhzXniMIKV" colab={"base_uri": "https://localhost:8080/"} outputId="67abef0a-917e-4931-bc2f-d3018f854d9b"
# Feature columns
features = train.columns[train.columns.str.startswith('Feature')]
features
# + [markdown] id="OXIw4yr2SqQD"
# # Set Targets
# + colab={"base_uri": "https://localhost:8080/"} id="p3HOqPyvOITY" outputId="0d6162ea-7ce0-4b0b-8b76-75e024c529e4"
# Targets columns
targets = train.columns[train.columns.str.startswith('target')]
targets
# + id="7hztXOF3tB45"
# Pick target_r
target = 'target_r'
# + [markdown] id="0lIVHkPYquZA"
# # Set Time group (Moons)
# + colab={"base_uri": "https://localhost:8080/"} id="bZxYqV0Fqut4" outputId="c5b2dd17-8c42-449a-bf05-acdc008e5539"
moons = train.Moons
moons
# + [markdown] id="HexJqSmrqK7L"
# # Standard Cross Validations
# + id="f2kLGdnOqJs5"
crossvalidators = [
model_selection.KFold(3),
model_selection.KFold(3, shuffle=True),
model_selection.GroupKFold(3),
model_selection.TimeSeriesSplit(3)
]
# + [markdown] id="WZ9KMRaZre35"
# # Metric Spearman Rank Correlation
# + id="VRzm5mR6rfMs"
def spearman(y_true, y_pred):
return spearmanr(y_pred, y_true).correlation
# + [markdown] id="3AXHEmcEr6zD"
# # Set Model
# + id="2hYUCjg8r7Hm"
model = xgb.XGBRegressor(objective="reg:squarederror", max_depth=5, learning_rate=0.01, n_estimators=200, n_jobs=-1, colsample_bytree=0.5)
# + [markdown] id="wEk0UC8fuHoo"
# # Calculate Cross Validations Scores
# + colab={"base_uri": "https://localhost:8080/"} id="iKZpmejjqJ1S" outputId="e0cbd3d8-6e0e-4547-f005-42b308a7a5ec"
for cv in crossvalidators:
print(cv)
print(np.mean(
model_selection.cross_val_score(
model,
train[features],
train[target],
cv=cv,
n_jobs=1,
groups=moons,
scoring=metrics.make_scorer(spearman, greater_is_better=True)
)))
print()
# + [markdown] id="bZ8hQBQGtYp_"
# # Here is a more elaborated Time-Series CV
# + id="SG-yflGrqJ6X"
class TimeSeriesSplitGroups(_BaseKFold):
def __init__(self, n_splits=5):
super().__init__(n_splits, shuffle=False, random_state=None)
def split(self, X, y=None, groups=None):
X, y, groups = indexable(X, y, groups)
n_samples = _num_samples(X)
n_splits = self.n_splits
n_folds = n_splits + 1
group_list = np.unique(groups)
n_groups = len(group_list)
if n_folds > n_groups:
raise ValueError(
("Cannot have number of folds ={0} greater"
" than the number of samples: {1}.").format(n_folds,
n_groups))
indices = np.arange(n_samples)
test_size = (n_groups // n_folds)
test_starts = range(test_size + n_groups % n_folds,
n_groups, test_size)
test_starts = list(test_starts)[::-1]
for test_start in test_starts:
yield (indices[groups.isin(group_list[:test_start])],
indices[groups.isin(group_list[test_start:test_start + test_size])])
# + colab={"base_uri": "https://localhost:8080/"} id="031hpfi-qJ9n" outputId="dc807379-a255-42a5-a339-54f08829f8b9"
print(np.mean(
model_selection.cross_val_score(
model,
train[features],
train[target],
cv=TimeSeriesSplitGroups(3),
n_jobs=1,
groups=moons,
scoring=metrics.make_scorer(spearman, greater_is_better=True)
)))
print(cv)
# + [markdown] id="UlIxOTqJv6rQ"
# # About
# Last updated: 2021-02-25
#
# Created by: [<NAME>](https://github.com/jberros)
#
# Greatly inspired by the works from: [<NAME>](https://github.com/jonrtaylor) and [<NAME>](https://github.com/the-moliver)
| quickstart/Datacrunch_Cross_Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ###### Import libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
# ###### Import Data
dirty_data = pd.read_csv("datasets/train.csv")
data = dirty_data.dropna()
# ###### Plotting Data
plt.scatter(data.x,data.y)
plt.xlabel("x value")
plt.ylabel("y value")
plt.rcParams["figure.figsize"] = [25,15]
plt.show()
# ###### Linear Regression
linear_reg = LinearRegression()
x = data.x.values.reshape(-1,1)
y = data.y.values.reshape(-1,1)
linear_reg.fit(x,y)
# ###### Prediction
b0 = linear_reg.predict([[0]])
print("b0 =",b0)
b0_ = linear_reg.intercept_
print("b0_ =",b0_)
b1 = linear_reg.coef_
print("b1 =",b1)
linear_reg.predict([[25]])
# ###### Line Visulation
array = np.array(range(0,101)).reshape(-1,1)
y_head = linear_reg.predict(array)
plt.scatter(x,y)
plt.plot(array,y_head, color = "red")
plt.show()
| Machine Learning Algorithms/a. Linear Regression/other works/Linear Regression Datasets(test&train)/Study Assignment-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "notes"} toc-hr-collapsed=false
# # Parsing and Recombining Inputs
#
# In the chapter on [Grammars](Grammars.ipynb), we discussed how grammars can be
# used to represent various languages. We also saw how grammars can be used to
# generate strings of the corresponding language. Grammars can also perform the
# reverse. That is, given a string, one can decompose the string into its
# constituent parts that correspond to the parts of grammar used to generate it
# – the _derivation tree_ of that string. These parts (and parts from other similar
# strings) can later be recombined using the same grammar to produce new strings.
#
# In this chapter, we use grammars to parse and decompose a given set of valid seed inputs into their corresponding derivation trees. This structural representation allows us to mutate, crossover, and recombine their parts in order to generate new valid, slightly changed inputs (i.e., fuzz)
# + [markdown] slideshow={"slide_type": "skip"}
# **Prerequisites**
#
# * You should have read the [chapter on grammars](Grammars.ipynb).
# * An understanding of derivation trees from the [chapter on grammar fuzzer](GrammarFuzzer.ipynb)
# is also required.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Fuzzing a Simple Proram
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is a simple program that accepts a CSV file of vehicle details and processes this information.
# + slideshow={"slide_type": "subslide"}
def process_vehicle(vehicle):
year, kind, company, model, *_ = vehicle.split(',')
if kind == 'van':
print("We have a %s %s van from %s vintage." % (company, model, year))
iyear = int(year)
if year > 2010:
print("It is a recent model!")
else:
print("It is an old but reliable model!")
elif kind == 'car':
print("We have a %s %s car from %s vintage." % (company, model, year))
iyear = int(year)
if year > 2016:
print("It is a recent model!")
else:
print("It is an old but reliable model!")
else:
raise Exception('Invalid entry')
# + [markdown] slideshow={"slide_type": "subslide"}
# Here is a sample of inputs that the `process_vehicle()` accepts.
# + slideshow={"slide_type": "fragment"}
mystring = """\
1997,van,Ford,E350
2000,car,Mercury,Cougar\
"""
print(mystring)
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us try to fuzz this program. Given that it takes a CSV file, we can write a simple grammar for generating comma separated values, and generate the required inputs.
# + slideshow={"slide_type": "skip"}
import string
# + slideshow={"slide_type": "subslide"}
CSV_GRAMMAR = {
'<start>' : ['<csvline>'],
'<csvline>': ['<items>'],
'<items>' : ['<item>,<items>', '<item>'],
'<item>' : ['<letters>'],
'<letters>': ['<letter><letters>', '<letter>'],
'<letter>' : list(string.ascii_letters + string.digits + string.punctuation + ' \t\n')
}
# + [markdown] slideshow={"slide_type": "fragment"}
# We need some infrastructure first for viewing the grammar.
# + slideshow={"slide_type": "skip"}
import fuzzingbook_utils
# + slideshow={"slide_type": "skip"}
from Grammars import EXPR_GRAMMAR, START_SYMBOL, RE_NONTERMINAL, is_valid_grammar, syntax_diagram
from Fuzzer import Fuzzer
from GrammarFuzzer import GrammarFuzzer, FasterGrammarFuzzer, display_tree, tree_to_string, dot_escape
from ExpectError import ExpectError
from Coverage import Coverage
from Timer import Timer
# + slideshow={"slide_type": "subslide"}
syntax_diagram(CSV_GRAMMAR)
# + [markdown] slideshow={"slide_type": "subslide"}
# We generate `1000` values, and evaluate the `process_vehicle()` with each.
# + slideshow={"slide_type": "subslide"}
gf = GrammarFuzzer(CSV_GRAMMAR, min_nonterminals=4)
trials = 1000
valid = []
time = 0
for i in range(trials):
with Timer() as t:
vehicle_info = gf.fuzz()
try:
process_vehicle(vehicle_info)
valid.append(vehicle_info)
except:
pass
time += t.elapsed_time()
print("%d valid strings, that is GrammarFuzzer generated %f%% valid entries from %d inputs" % (len(valid), len(valid)*100.0/trials , trials))
print("Total time of %f seconds" % time)
# + [markdown] slideshow={"slide_type": "subslide"}
# This is obviously not working. But why?
# + slideshow={"slide_type": "subslide"}
gf = GrammarFuzzer(CSV_GRAMMAR, min_nonterminals=4)
trials = 10
valid = []
time = 0
for i in range(trials):
vehicle_info = gf.fuzz()
try:
print(repr(vehicle_info), end="")
process_vehicle(vehicle_info)
except Exception as e:
print("\t", e)
else:
print()
# + [markdown] slideshow={"slide_type": "subslide"}
# None of the entries will get through unless the fuzzer can produce either `van` or `car`.
# Indeed, the reason is that the grammar itself does not capture the complete information about the format. So here is another idea. We modify the `GrammarFuzzer` to know a bit about our format.
# + slideshow={"slide_type": "skip"}
from copy import deepcopy
import random
class PooledGrammarFuzzer(GrammarFuzzer):
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self._node_cache = {}
def update_cache(self, key, values):
self._node_cache[key] = values
def expand_node_randomly(self, node):
(symbol, children) = node
assert children is None
if symbol in self._node_cache:
if random.randint(0, 1) == 1:
return super().expand_node_randomly(node)
return deepcopy(random.choice(self._node_cache[symbol]))
return super().expand_node_randomly(node)
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us try again!
# + slideshow={"slide_type": "subslide"}
gf = PooledGrammarFuzzer(CSV_GRAMMAR, min_nonterminals=4)
gf.update_cache('<item>', [
('<item>', [('car', [])]),
('<item>', [('van', [])]),
])
trials = 10
valid = []
time = 0
for i in range(trials):
vehicle_info = gf.fuzz()
try:
print(repr(vehicle_info), end="")
process_vehicle(vehicle_info)
except Exception as e:
print("\t", e)
else:
print()
# + [markdown] slideshow={"slide_type": "subslide"}
# At least we are getting somewhere! It would be really nice if we could incorporate what we know about the sample data in our fuzzer. Infact, it would be nice if we could extract the template and valid values from samples, and use them in our fuzzing. How do we do that?
# + [markdown] slideshow={"slide_type": "slide"}
# ## An Ad Hoc Parser
# + [markdown] slideshow={"slide_type": "fragment"}
# As we saw in the previous section, programmers often have to extract parts of data that obeys certain rules. For example, for *CSV* files, each element in a raw is separated by *commas*, and multiple raws are used to store the data.
# + [markdown] slideshow={"slide_type": "fragment"}
# To extract the information, we write an ad hoc parser `parse_csv()`.
# + slideshow={"slide_type": "fragment"}
def parse_csv(mystring):
children = []
tree = (START_SYMBOL, children)
for i,line in enumerate(mystring.split('\n')):
children.append(("record %d" %i, [(cell,[]) for cell in line.split(',')]))
return tree
# + [markdown] slideshow={"slide_type": "fragment"}
# We also change the default orientation of the graph to *left to right* rather than *top to bottom* for easier viewing using `lr_graph()`.
# + slideshow={"slide_type": "subslide"}
def lr_graph(dot):
dot.attr('node', shape='plain')
dot.graph_attr['rankdir'] = 'LR'
# + [markdown] slideshow={"slide_type": "fragment"}
# The `display_tree()` shows the structure of our CSV file after parsing.
# + slideshow={"slide_type": "fragment"}
tree = parse_csv(mystring)
display_tree(tree, graph_attr=lr_graph)
# + [markdown] slideshow={"slide_type": "fragment"}
# This is of course simple. What if we encounter slightly more complexity? Again, another example from the Wikipedia.
# + slideshow={"slide_type": "subslide"}
mystring = '''\
1997,Ford,E350,"ac, abs, moon",3000.00\
'''
print(mystring)
# + [markdown] slideshow={"slide_type": "fragment"}
# We define a new annotation method `highlight_node()` to mark the nodes that are interesting.
# + slideshow={"slide_type": "fragment"}
def highlight_node(predicate):
def hl_node(dot, nid, symbol, ann):
if predicate(dot, nid, symbol, ann):
dot.node(repr(nid), dot_escape(symbol), fontcolor='red')
else:
dot.node(repr(nid), dot_escape(symbol))
return hl_node
# + [markdown] slideshow={"slide_type": "subslide"}
# Using `highlight_node()` we can highlight particular nodes that we were wrongly parsed.
# + slideshow={"slide_type": "fragment"}
tree = parse_csv(mystring)
bad_nodes = {5,6,7,12,13,20,22,23,24,25}
hl_predicate = lambda _d,nid,_s,_a: nid in bad_nodes
highlight_err_node = highlight_node(hl_predicate)
display_tree(tree, log=False, node_attr=highlight_err_node, graph_attr=lr_graph)
# + [markdown] slideshow={"slide_type": "fragment"}
# The marked nodes indicate where our parsing went wrong. We can of course extend our parser to understand quotes. First we define some of the helper functions `parse_quote()`, `find_comma()` and `comma_split()`
# + slideshow={"slide_type": "subslide"}
def parse_quote(string, i):
v = string[i+1:].find('"')
return v+i+1 if v >= 0 else -1
def find_comma(string, i):
slen = len(string)
while i < slen:
if string[i] == '"':
i = parse_quote(string, i)
if i == -1:
return -1
if string[i] == ',':
return i
i+=1
return -1
def comma_split(string):
slen = len(string)
i = 0
while i < slen:
c = find_comma(string, i)
if c == -1:
yield string[i:]
return
else:
yield string[i:c]
i = c+1
# + [markdown] slideshow={"slide_type": "subslide"}
# We can update our `parse_csv()` procedure to use our advanced quote parser.
# + slideshow={"slide_type": "fragment"}
def parse_csv(mystring):
children = []
tree = (START_SYMBOL, children)
for i,line in enumerate(mystring.split('\n')):
children.append(("record %d" %i, [(cell,[]) for cell in comma_split(line)]))
return tree
# + [markdown] slideshow={"slide_type": "fragment"}
# Our new `parse_csv()` can now handle quotes correctly.
# + slideshow={"slide_type": "fragment"}
tree = parse_csv(mystring)
display_tree(tree, graph_attr=lr_graph)
# + [markdown] slideshow={"slide_type": "fragment"}
# That of course does not survive long:
# + slideshow={"slide_type": "subslide"}
mystring = '''\
1999,Chevy,"Venture \\"Extended Edition, Very Large\\"",,5000.00\
'''
print(mystring)
# + [markdown] slideshow={"slide_type": "fragment"}
# A few embedded quotes are sufficient to confuse our parser again.
# + slideshow={"slide_type": "fragment"}
tree = parse_csv(mystring)
bad_nodes = {4,5}
display_tree(tree, node_attr=highlight_err_node, graph_attr=lr_graph)
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is another record from that CSV file:
# + slideshow={"slide_type": "subslide"}
mystring = '''\
1996,Jeep,Grand Cherokee,"MUST SELL!
air, moon roof, loaded",4799.00
'''
print(mystring)
# + slideshow={"slide_type": "fragment"}
tree = parse_csv(mystring)
bad_nodes = {5, 6,7,8,9,10}
display_tree(tree, node_attr=highlight_err_node, graph_attr=lr_graph)
# + [markdown] slideshow={"slide_type": "fragment"}
# Fixing this would require modifying both inner `parse_quote()` and the outer `parse_csv()` procedures.
# + [markdown] slideshow={"slide_type": "subslide"}
# Indeed, each additional improvement falls apart even with a little extra complexity. The problem becomes severe when one encounters recursive expressions. For example, JSON is a common alternative to CSV files for saving data. Similarly, one may have to parse data from an HTML table instead of a CSV file if one is getting the data from the web.
#
# One might be tempted to fix it with a little more ad hoc parsing, with a bit of *regular expressions* thrown in. However, that is the [path to insanity](https://stackoverflow.com/a/1732454).
#
# It is here that the formal parsers shine. The main idea is that, any given set of strings belong to a language, and these languages can be specified by their grammars (as we saw in the [chapter on grammars](Grammars.ipynb)). The great thing about grammars is that they can be *composed*. That is, one can introduce finer and finer details into an internal structure without affecting the external structure, and similarly, one can change the external structure without much impact on the internal structure. We briefly describe grammars in the next section.
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true
# ## Grammars
#
# A grammar, as you have read from the [chapter on grammars](Grammars.ipynb) is a set of _rules_ that explain how the start symbol can be expanded. Each rule has a name, also called a _nonterminal_, and a set of _alternative choices_ in how the nonterminal can be expanded.
# + slideshow={"slide_type": "subslide"}
A1_GRAMMAR = {
"<start>": ["<expr>"],
"<expr>": ["<expr>+<expr>", "<expr>-<expr>", "<integer>"],
"<integer>": ["<digit><integer>", "<digit>"],
"<digit>": ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"]
}
# + slideshow={"slide_type": "subslide"}
syntax_diagram(A1_GRAMMAR)
# + [markdown] slideshow={"slide_type": "notes"}
# In the above expression, the rule `<expr> : [<expr>+<expr>,<expr>-<expr>,<integer>]` corresponds to how the nonterminal `<expr>` might be expanded. The expression `<expr>+<expr>` corresponds to one of the alternative choices. We call this an _alternative_ expansion for the nonterminal `<expr>`. Finally, in an expression `<expr>+<expr>`, each of `<expr>`, `+`, and `<expr>` are _symbols_ in that expansion. A symbol could be either a nonterminal or a terminal symbol based on whether its expansion is available in the grammar.
# + [markdown] slideshow={"slide_type": "slide"}
# Here is a string that represents an arithmetic expression that we would like to parse, which is specified by the grammar above:
# + slideshow={"slide_type": "fragment"}
mystring = '1+2'
# + [markdown] slideshow={"slide_type": "subslide"}
# The _derivation tree_ for our expression from this grammar is given by:
# + slideshow={"slide_type": "fragment"}
tree = ('<start>', [('<expr>',
[('<expr>', [('<integer>', [('<digit>', [('1', [])])])]),
('+', []),
('<expr>', [('<integer>', [('<digit>', [('2',
[])])])])])])
assert mystring == tree_to_string(tree)
display_tree(tree)
# + [markdown] slideshow={"slide_type": "subslide"}
# While a grammar can be used to specify a given language, there could be multiple
# grammars that correspond to the same language. For example, here is another
# grammar to describe the same addition expression.
# + slideshow={"slide_type": "subslide"}
A2_GRAMMAR = {
"<start>": ["<expr>"],
"<expr>": ["<integer><expr_>"],
"<expr_>": ["+<expr>", "-<expr>", ""],
"<integer>": ["<digit><integer_>"],
"<integer_>": ["<integer>", ""],
"<digit>": ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"]
}
# + slideshow={"slide_type": "subslide"}
syntax_diagram(A2_GRAMMAR)
# + [markdown] slideshow={"slide_type": "subslide"}
# The corresponding derivation tree is given by:
# + slideshow={"slide_type": "subslide"}
tree = ('<start>', [('<expr>', [('<integer>', [('<digit>', [('1', [])]),
('<integer_>', [])]),
('<expr_>', [('+', []),
('<expr>',
[('<integer>',
[('<digit>', [('2', [])]),
('<integer_>', [])]),
('<expr_>', [])])])])])
assert mystring == tree_to_string(tree)
display_tree(tree)
# + [markdown] slideshow={"slide_type": "notes"}
# Indeed, there could be different classes of grammars that
# describe the same language. For example, the first grammar `A1_GRAMMAR`
# is a grammar that sports both _right_ and _left_ recursion, while the
# second grammar `A2_GRAMMAR` does not have left recursion in the
# nonterminals in any of its productions, but contains _epsilon_ productions.
# (An epsilon production is a production that has empty string in its right
# hand side.)
#
# You would have noticed that we reuse the term `<expr>` in its own definition. Using the same nonterminal in its own definition is called *recursion*. There are two specific kinds of recursion one should be aware of in parsing, as we see in the next section.
# + [markdown] slideshow={"slide_type": "notes"}
# #### Recursion
#
# A grammar is _left recursive_ if any of its nonterminals are left recursive,
# and a nonterminal is directly left-recursive if the left-most symbol of
# any of its productions is itself.
# + slideshow={"slide_type": "fragment"}
LR_GRAMMAR = {
'<start>': ['<A>'],
'<A>': ['<A>a', ''],
}
# + slideshow={"slide_type": "subslide"}
syntax_diagram(LR_GRAMMAR)
# + slideshow={"slide_type": "fragment"}
mystring = 'aaaaaa'
display_tree(('<start>', (('<A>', (('<A>', (('<A>', []), ('a', []))), ('a', []))), ('a', []))))
# + [markdown] slideshow={"slide_type": "subslide"}
# A grammar is indirectly left-recursive if any
# of the left-most symbols can be expanded using their definitions to
# produce the nonterminal as the left-most symbol of the expansion. The left
# recursion is called a _hidden-left-recursion_ if during the series of
# expansions of a nonterminal, one reaches a rule where the rule contains
# the same nonterminal after a prefix of other symbols, and these symbols can
# dervive the empty string. For example, in `A1_GRAMMAR`, `<integer>` will be
# considered hidden-left recursive if `<digit>` could derive an empty string.
#
# Right recursive grammars are defined similarly.
# Below is the derivation tree for the right recursive grammar that represents the same
# language as that of `LR_GRAMMAR`.
# + slideshow={"slide_type": "subslide"}
RR_GRAMMAR = {
'<start>': ['<A>'],
'<A>': ['a<A>', ''],
}
# + slideshow={"slide_type": "fragment"}
syntax_diagram(RR_GRAMMAR)
# + slideshow={"slide_type": "fragment"}
display_tree(('<start>', (('<A>', (('a', []), ('<A>', (('a', []), ('<A>', (('a', []), ('<A>', []))))))),)))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Ambiguity
#
# To complicate matters further, there could be
# multiple derivation trees – also called _parses_ – corresponding to the
# same string from the same grammar. For example, a string `1+2+3` can be parsed
# in two ways as we see below using the `A1_GRAMMAR`
# + slideshow={"slide_type": "subslide"}
mystring = '1+2+3'
tree = ('<start>',
[('<expr>',
[('<expr>', [('<expr>', [('<integer>', [('<digit>', [('1', [])])])]),
('+', []),
('<expr>', [('<integer>',
[('<digit>', [('2', [])])])])]), ('+', []),
('<expr>', [('<integer>', [('<digit>', [('3', [])])])])])])
assert mystring == tree_to_string(tree)
display_tree(tree)
# + slideshow={"slide_type": "subslide"}
tree = ('<start>',
[('<expr>', [('<expr>', [('<integer>', [('<digit>', [('1', [])])])]),
('+', []),
('<expr>',
[('<expr>', [('<integer>', [('<digit>', [('2', [])])])]),
('+', []),
('<expr>', [('<integer>', [('<digit>', [('3',
[])])])])])])])
assert tree_to_string(tree) == mystring
display_tree(tree)
# + [markdown] slideshow={"slide_type": "subslide"}
# There are many ways to resolve ambiguities. One approach taken by *Parsing Expression Grammars* explained in next section is to specify a particular order of resolution, and choose the first one. Another approach is to simply return all possible derivation trees, which is the approach taken by *Earley parser* we develop later.
# + [markdown] slideshow={"slide_type": "subslide"}
# Next, we develop different parsers. To do that, we define a minimal interface for parsing that is obeyed by all parsers. There are two approaches to parsing a string using a grammar.
#
# 1. The traditional approach is to use a *lexer* (also called a *tokenizer* or a *scanner*) to first tokenize the incoming string, and feed the grammar one token at a time. The lexer is typically a smaller parser that accepts a *regular language*. The advantage of this approach is that the grammar used by the parser can eschew the details of tokenization. Further, one gets a shallow derivation tree at the end of the parsing which can be directly used for generating the *Abstract Syntax Tree*.
# 2. The second approach is to use a tree pruner after the complete parse. With this approach, one uses a grammar that incorporates complete details of the syntax. Next, the nodes corresponding to tokens are pruned and replaced with their corresponding strings as leaf nodes. The utility of this approach is that the parser is more powerful, and further there is no artificial distinction between *lexing* and *parsing*.
#
# In this chapter, we use the second approach.
# + [markdown] slideshow={"slide_type": "subslide"}
# The *Parser* class we define below provides the minimal interface. The main methods that need to be implemented by the classes implementing this interface are `parse_prefix` and `parse`. The `parse_prefix` returns a tuple, which contains the index until which parsing was completed successfully, and the parse forest until that index. The method `parse` returns a list of derivation trees if the parse was successful.
# + slideshow={"slide_type": "subslide"}
class Parser(object):
def __init__(self, grammar, **kwargs):
self._grammar = grammar
self.start_symbol = kwargs.get('start_symbol') or START_SYMBOL
self.log = kwargs.get('log') or False
self.tokens = kwargs.get('tokens') or set()
def grammar(self):
return self._grammar
def parse_prefix(self, text):
"""Return pair (cursor, forest) for longest prefix of text"""
raise NotImplemented()
def parse(self, text):
cursor, forest = self.parse_prefix(text)
if cursor < len(text):
raise SyntaxError("at " + repr(text[cursor:]))
return [self.prune_tree(tree) for tree in forest]
def prune_tree(self, tree):
name, children = tree
if name in self.tokens:
return (name, [(tree_to_string(tree), [])])
else:
return (name, [self.prune_tree(c) for c in children])
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true
# ## Parsing Expression Grammars
# + [markdown] slideshow={"slide_type": "subslide"}
# A _[Parsing Expression Grammar](http://bford.info/pub/lang/peg)_ (*PEG*) \cite{Ford2004} is a type of _recognition based formal grammar_ that specifies the sequence of steps to take to parse a given string.
# A _parsing expression grammar_ is very similar to a _context-free grammar_ (*CFG*) such as the ones we saw in the [chapter on grammars](Grammars.ipynb). As in a CFG, a parsing expression grammar is represented by a set of nonterminals and corresponding alternatives representing how to match each. For example, here is a PEG that matches `a` or `b`.
# + slideshow={"slide_type": "fragment"}
PEG1 = {
'<start>': ['a', 'b']
}
# + [markdown] slideshow={"slide_type": "notes"}
# However, unlike the _CFG_, the alternatives represent *ordered choice*. That is, rather than choosing all rules that can potentially match, we stop at the first match that succeed. For example, the below _PEG_ can match `ab` but not `abc` unlike a _CFG_ which will match both. (We call the sequence of ordered choice expressions *choice expressions* rather than alternatives to make the distinction from _CFG_ clear.)
# + slideshow={"slide_type": "subslide"}
PEG2 = {
'<start>': ['ab', 'abc']
}
# + [markdown] slideshow={"slide_type": "notes"}
# Each choice in a _choice expression_ represents a rule on how to satisfy that particular choice. The choice is a sequence of symbols (terminals and nonterminals) that are matched against a given text as in a _CFG_.
# + [markdown] slideshow={"slide_type": "skip"}
# Beyond the syntax of grammar definitions we have seen so far, a _PEG_ can also contain a few additional elements. See the exercises at the end of the chapter for additional information.
#
# The PEGs model the typical practice in handwritten recursive descent parsers, and hence it may be considered more intuitive to understand. We look at parsers for PEGs next.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### The Packrat Parser for Predicate Expression Grammars
#
# Short of hand rolling a parser, _Packrat_ parsing is one of the simplest parsing techniques, and is one of the techniques for parsing PEGs.
# The _Packrat_ parser is so named because it tries to cache all results from simpler problems in the hope that these solutions can be used to avoid re-computation later. We develop a minimal _Packrat_ parser next.
#
# But before that, we need to implement a few supporting tools.
# + [markdown] slideshow={"slide_type": "skip"}
# The `EXPR_GRAMMAR` we import from the [chapter on grammars](Grammars.ipynb) is oriented towards generation. In particular, the production rules are stored as strings. We need to massage this representation a little to conform to a canonical representation where each token in a rule is represented separately. The `canonical` format uses separate tokens to represent each symbol in an expansion.
# + slideshow={"slide_type": "skip"}
import re
def canonical(grammar, letters=False):
def split(rule):
return [token for token in re.split(RE_NONTERMINAL, rule) if token]
def tokenize(word):
return list(word) if letters else [word]
def canonical_expr(expression):
return [
token for word in split(expression)
for token in ([word] if word in grammar else tokenize(word))
]
return {
k: [canonical_expr(expression) for expression in alternatives]
for k, alternatives in grammar.items()
}
# + slideshow={"slide_type": "skip"}
canonical(EXPR_GRAMMAR)
# + [markdown] slideshow={"slide_type": "skip"}
# It is easier to work with the `canonical` representation during parsing. Hence, we update our parser class to store the `canonical` representation also.
# + slideshow={"slide_type": "subslide"}
class Parser(Parser):
def __init__(self, grammar, **kwargs):
self._grammar = grammar
self.start_symbol = kwargs.get('start_symbol') or START_SYMBOL
self.log = kwargs.get('log') or False
self.tokens = kwargs.get('tokens') or set()
self.cgrammar = canonical(grammar)
# + [markdown] slideshow={"slide_type": "subslide"} toc-hr-collapsed=false
# ### The Parser
#
# We derive from the `Parser` base class first, and we accept the text to be parsed in the `parse()` method, which in turn calls `unify_key()` with the `start_symbol`.
#
# __Note.__ While our PEG parser can produce only a single unambiguous parse tree, other parsers can produce multiple parses for ambiguous grammars. Hence, we return a list of trees (in this case with a single element).
# + slideshow={"slide_type": "subslide"}
class PEGParser(Parser):
def parse_prefix(self, text):
cursor, tree = self.unify_key(self.start_symbol, text, 0)
return cursor, [tree]
# + [markdown] slideshow={"slide_type": "notes"}
# #### Unify Key
# The `unify_key()` algorithm is simple. If given a terminal symbol, it tries to match the symbol with the current position in the text. If the symbol and text match, it returns successfully with the new parse index `at`.
#
# If on the other hand, it was given a nonterminal, it retrieves the choice expression corresponding to the key, and tries to match each choice *in order* using `unify_rule()`. If **any** of the rules succeed in being unified with the given text, the parse is considered a success, and we return with the new parse index returned by `unify_rule()`.
# + button=false code_folding=[] new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
class PEGParser(PEGParser):
def unify_key(self, key, text, at=0):
if self.log:
print("unify_key: %s with %s" % (repr(key), repr(text[at:])))
if key not in self.cgrammar:
if text[at:].startswith(key):
return at + len(key), (key, [])
else:
return at, None
for rule in self.cgrammar[key]:
to, res = self.unify_rule(rule, text, at)
if res:
return (to, (key, res))
return 0, None
# + slideshow={"slide_type": "subslide"}
mystring = "1"
peg = PEGParser(EXPR_GRAMMAR, log=True)
peg.unify_key('1', mystring)
# + slideshow={"slide_type": "fragment"}
mystring = "2"
peg.unify_key('1', mystring)
# + [markdown] slideshow={"slide_type": "notes"}
# #### Unify Rule
#
# The `unify_rule()` method is similar. It retrieves the tokens corresponding to the rule that it needs to unify with the text, and calls `unify_key()` on them in sequence. If **all** tokens are successfully unified with the text, the parse is a success.
# + slideshow={"slide_type": "subslide"}
class PEGParser(PEGParser):
def unify_rule(self, rule, text, at):
if self.log:
print('unify_rule: %s with %s' % (repr(rule), repr(text[at:])))
results = []
for token in rule:
at, res = self.unify_key(token, text, at)
if res is None:
return at, None
results.append(res)
return at, results
# + slideshow={"slide_type": "subslide"}
mystring = "0"
peg = PEGParser(EXPR_GRAMMAR, log=True)
peg.unify_rule(peg.cgrammar['<digit>'][0], mystring, 0)
# + slideshow={"slide_type": "subslide"}
mystring = "12"
peg.unify_rule(peg.cgrammar['<integer>'][0], mystring, 0)
# + slideshow={"slide_type": "subslide"}
mystring = "1 + 2"
peg = PEGParser(EXPR_GRAMMAR, log=False)
peg.parse(mystring)
# + [markdown] slideshow={"slide_type": "notes"}
# The two methods are mutually recursive, and given that `unify_key()` tries each alternative until it succeeds, `unify_key` can be called multiple times with the same arguments. Hence, it is important to memoize the results of `unify_key`. Python provides a simple decorator `lru_cache` for memoizing any function call that has hashable arguments. We add that to our implementation so that repeated calls to `unify_key()` with the same argument get cached results.
#
# This memoization gives the algorithm its name – _Packrat_.
# + slideshow={"slide_type": "skip"}
from functools import lru_cache
# + slideshow={"slide_type": "subslide"}
class PEGParser(PEGParser):
@lru_cache(maxsize=None)
def unify_key(self, key, text, at=0):
if key not in self.cgrammar:
if text[at:].startswith(key):
return at + len(key), (key, [])
else:
return at, None
for rule in self.cgrammar[key]:
to, res = self.unify_rule(rule, text, at)
if res:
return (to, (key, res))
return 0, None
# + [markdown] slideshow={"slide_type": "notes"}
# We wrap initialization and calling of `PEGParser` in a method `parse()` already implemented in the `Parser` base class that accepts the text to be parsed along with the grammar.
# + [markdown] slideshow={"slide_type": "notes"}
# Here are a few examples of our parser in action.
# + slideshow={"slide_type": "subslide"}
mystring = "1 + (2 * 3)"
peg = PEGParser(EXPR_GRAMMAR)
for tree in peg.parse(mystring):
assert tree_to_string(tree) == mystring
display_tree(tree)
# + slideshow={"slide_type": "subslide"}
mystring = "1 * (2 + 3.35)"
for tree in peg.parse(mystring):
assert tree_to_string(tree) == mystring
display_tree(tree)
# + [markdown] slideshow={"slide_type": "notes"}
# One should be aware that while the grammar looks like a *CFG*, the language described by a *PEG* may be different. Indeed, only *LL(1)* grammars are guaranteed to represent the same language for both PEGs and other parsers. Behavior of PEGs for other classes of grammars could be surprising \cite{redziejowski2008}.
# + [markdown] slideshow={"slide_type": "fragment"}
# We previously showed how using fragments of existing data can help quite a bit with fuzzing. We now explore this idea in more detail.
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true
# ## Recombining Parsed Inputs
# + [markdown] slideshow={"slide_type": "notes"}
# Recombining parsed inputs was pioneered by _Langfuzz_ \cite{Holler2012}. The main challenge is that program inputs often carry additional constraints beyond what is described by the syntax. For example, in Java, one needs to declare a variable (using a specific format for declaration) before it can be used in an expression. This restriction is not captured in the _Java CFG_. Checking for type correctness is another example for additional restrictions carried by program definitions.
# + [markdown] slideshow={"slide_type": "notes"}
# When fuzzing compilers and interpreters, naive generation of programs using the language *CFG* often fails to achieve significant deeper coverage due to these kinds of checks external to the grammar. Holler et al. suggests using pre-existing valid code fragments to get around these restrictions. The idea is that the pre-existing valid code fragments already conform to the restrictions external to the grammar, and can often provide a means to evade validity checks.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### A Grammar-based Mutational Fuzzer
#
# The idea is that one can treat the derivation tree of a preexisting program as the scaffolding, poke holes in it, and patch it with generated inputs from our grammar. Given below is a grammar for a language that allows assignment of variables.
# + slideshow={"slide_type": "subslide"}
import string
# + slideshow={"slide_type": "skip"}
from Grammars import crange
# + slideshow={"slide_type": "subslide"}
VAR_GRAMMAR = {
'<start>': ['<statements>'],
'<statements>': ['<statement>;<statements>', '<statement>'],
'<statement>': ['<assignment>'],
'<assignment>': ['<identifier>=<expr>'],
'<identifier>': ['<word>'],
'<word>': ['<alpha><word>', '<alpha>'],
'<alpha>': list(string.ascii_letters),
'<expr>': ['<term>+<expr>', '<term>-<expr>', '<term>'],
'<term>': ['<factor>*<term>', '<factor>/<term>', '<factor>'],
'<factor>':
['+<factor>', '-<factor>', '(<expr>)', '<identifier>', '<number>'],
'<number>': ['<integer>.<integer>', '<integer>'],
'<integer>': ['<digit><integer>', '<digit>'],
'<digit>': crange('0', '9')
}
# + slideshow={"slide_type": "subslide"}
syntax_diagram(VAR_GRAMMAR)
# + [markdown] slideshow={"slide_type": "notes"}
# Let us use our new grammar to parse a program.
# + slideshow={"slide_type": "fragment"}
mystring = 'va=10;vb=20'
hl_predicate = lambda _d,_n,symbol,_a: symbol in {'<number>', '<identifier>'}
parser = PEGParser(VAR_GRAMMAR)
for tree in parser.parse(mystring):
display_tree(tree, node_attr=highlight_node(hl_predicate))
# + [markdown] slideshow={"slide_type": "fragment"}
# As can be seen from the above example, our grammar is rather detailed. So we need to define our token nodes, which correspond to the *red* nodes above.
# + slideshow={"slide_type": "fragment"}
VAR_TOKENS = {'<number>', '<identifier>'}
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is a slightly more complex program to parse, but with the tree pruned using tokens:
# + slideshow={"slide_type": "subslide"}
mystring = 'avar=1.3;bvar=avar-3*(4+300)'
parser = PEGParser(VAR_GRAMMAR, tokens=VAR_TOKENS)
for tree in parser.parse(mystring):
display_tree(tree, node_attr=highlight_node(hl_predicate))
# + [markdown] slideshow={"slide_type": "notes"}
# We develop a `LangFuzzer` class that generates recombined inputs. To apply the _Langfuzz_ approach, we need a few parsed strings.
# + slideshow={"slide_type": "fragment"}
mystrings = [
'abc=12+(3+3.3)',
'a=1;b=2;c=a+b',
'avar=1.3;bvar=avar-3*(4+300)',
'a=1.3;b=a-1*(4+3+(2/a))',
'a=10;b=20;c=34;d=-b+(b*b-4*a*c)/(2*a)',
'x=10;y=20;z=(x+y)*(x-y)',
'x=23;y=51;z=x*x-y*y',
]
# + [markdown] slideshow={"slide_type": "notes"}
# We recurse through any given tree, collecting parsed fragments corresponding to each nonterminal. Further, we also name each node so that we can address each node separately.
# + slideshow={"slide_type": "subslide"}
class LangFuzzer(Fuzzer):
def __init__(self, parser):
self.parser = parser
self.fragments = {k: [] for k in self.parser.cgrammar}
def traverse_tree(self, node):
counter = 1
nodes = {}
def helper(node, id):
nonlocal counter
name, children = node
new_children = []
nodes[id] = node
for child in children:
counter += 1
new_children.append(helper(child, counter))
return name, new_children, id
return helper(node, counter), nodes
def fragment(self, strings):
self.trees = []
for string in strings:
for tree in self.parser.parse(string):
tree, nodes = self.traverse_tree(tree)
self.trees.append((tree, nodes))
for node in nodes:
symbol = nodes[node][0]
if symbol in self.fragments:
self.fragments[symbol].append(nodes[node])
return self.fragments
# + [markdown] slideshow={"slide_type": "skip"}
# We thus obtain all valid fragments from our parsed strings.
# + slideshow={"slide_type": "subslide"}
lf = LangFuzzer(PEGParser(VAR_GRAMMAR, tokens=VAR_TOKENS))
fragments = lf.fragment(mystrings)
for key in fragments:
print("%s: %d" % (key, len(fragments[key])))
# + [markdown] slideshow={"slide_type": "notes"}
# All that remains is to actually find a place to poke a hole using `candidate()`, and patch that hole using `generate_new_tree()`. We will explain how to do this next.
#
# But before that, we update our `initialization` method with a call to `fragment().
# + slideshow={"slide_type": "subslide"}
import random
# + slideshow={"slide_type": "fragment"}
class LangFuzzer(LangFuzzer):
def __init__(self, parser, strings):
self.parser = parser
self.fragments = {k: [] for k in self.parser.cgrammar}
self.fragment(strings)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Candidate
# `LangFuzzer` accepts a list of strings, which are stored as derivation trees in the object.
#
# The method `candidate()` chooses one of the derivation trees randomly as the template, and identifies a node such that it can be replaced by another node that is different from itself. That is, it chooses a node such that, if the non-terminal name of the node is `node_type`, there is at least one other entry in `fragment[node_type])`
# + slideshow={"slide_type": "subslide"}
class LangFuzzer(LangFuzzer):
def candidate(self):
tree, nodes = random.choice(self.trees)
interesting_nodes = [
n for n in nodes if nodes[n][0] in self.fragments
and len(self.fragments[nodes[n][0]]) > 1
]
node = random.choice(interesting_nodes)
return tree, node
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is how it is used -- the *red* node is the node chosen.
# + slideshow={"slide_type": "subslide"}
random.seed(1)
lf = LangFuzzer(PEGParser(VAR_GRAMMAR, tokens=VAR_TOKENS), mystrings)
tree, node = lf.candidate()
hl_predicate = lambda _d,nid,_s,_a: nid in {node}
display_tree(tree, node_attr=highlight_node(hl_predicate))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Generate New Tree
# Once we have identified the node, one can generate a new tree by replacing that node with another node of similar type from our fragment pool.
# + slideshow={"slide_type": "fragment"}
class LangFuzzer(LangFuzzer):
def generate_new_tree(self, node, choice):
name, children, id = node
if id == choice:
return random.choice(self.fragments[name])
else:
return (name, [self.generate_new_tree(c, choice)
for c in children])
# + [markdown] slideshow={"slide_type": "fragment"}
# Again, the red node indicates where the replacement has occurred.
# + slideshow={"slide_type": "subslide"}
random.seed(1)
lf = LangFuzzer(PEGParser(VAR_GRAMMAR, tokens=VAR_TOKENS), mystrings)
tree, node = lf.candidate()
hl_predicate = lambda _d,nid,_s,_a: nid in {node}
new_tree = lf.generate_new_tree(tree, node)
for s in [tree_to_string(i) for i in [tree, new_tree]]:
print(s)
display_tree(new_tree, node_attr=highlight_node(hl_predicate))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Fuzz
#
# The `fuzz()` method simply calls the procedures defined before in order.
# + slideshow={"slide_type": "fragment"}
class LangFuzzer(LangFuzzer):
def fuzz(self):
tree, node = self.candidate()
modified = self.generate_new_tree(tree, node)
return tree_to_string(modified)
# + [markdown] slideshow={"slide_type": "notes"}
# Here is our fuzzer in action.
# + slideshow={"slide_type": "subslide"}
lf = LangFuzzer(PEGParser(VAR_GRAMMAR, tokens=VAR_TOKENS), mystrings)
for i in range(10):
print(lf.fuzz())
# + [markdown] slideshow={"slide_type": "fragment"}
# How effective was our fuzzing? Let us findout!
# + slideshow={"slide_type": "subslide"}
trials = 100
lf = LangFuzzer(PEGParser(VAR_GRAMMAR, tokens=VAR_TOKENS), mystrings)
valid = []
time = 0
for i in range(trials):
with Timer() as t:
s = lf.fuzz()
try:
exec(s,{},{})
valid.append((s, t.elapsed_time()))
except:
pass
time += t.elapsed_time()
print("%d valid strings, that is LangFuzzer generated %f%% valid entries" % (len(valid), len(valid)*100.0/trials ))
print("Total time of %f seconds" % time)
# + slideshow={"slide_type": "subslide"}
gf = GrammarFuzzer(VAR_GRAMMAR)
valid = []
time = 0
for i in range(trials):
with Timer() as t:
s = gf.fuzz()
try:
exec(s,{},{})
valid.append(s)
except:
pass
time += t.elapsed_time()
print("%d valid strings, that is GrammarFuzzer generated %f%% valid entries" % (len(valid), len(valid)*100.0/trials ))
print("Total time of %f seconds" % time)
# + [markdown] slideshow={"slide_type": "subslide"}
# That is, our `LangFuzzer` is rather effective on generating valid entries when compared to the `GrammarFuzzer`.
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true
# ## Parsing Context-Free Grammars
# + [markdown] slideshow={"slide_type": "slide"}
# ### Problems with PEG
# While _PEGs_ are simple at first sight, their behavior in some cases might be a bit unintuitive. For example, here is an example \cite{redziejowski}:
# + slideshow={"slide_type": "subslide"}
PEG_SURPRISE = {
"<A>": ["a<A>a", "aa"]
}
# + [markdown] slideshow={"slide_type": "notes"}
# When interpreted as a *CFG* and used as a string generator, it will produce strings of the form `aa, aaaa, aaaaaa` that is, it produces strings where the number of `a` is $ 2*n $ where $ n > 0 $.
# + slideshow={"slide_type": "subslide"}
strings = []
for e in range(4):
f = GrammarFuzzer(PEG_SURPRISE, '<A>')
tree = ('<A>', None)
for _ in range(e):
tree = f.expand_tree_once(tree)
tree = f.expand_tree_with_strategy(tree, f.expand_node_min_cost)
strings.append(tree_to_string(tree))
display_tree(tree)
strings
# + [markdown] slideshow={"slide_type": "subslide"}
# However, the _PEG_ parser can only recognize strings of the form $2^n$
# + slideshow={"slide_type": "subslide"}
peg = PEGParser(PEG_SURPRISE, start_symbol='<A>')
for s in strings:
with ExpectError():
for tree in peg.parse(s):
display_tree(tree)
print(s)
# + [markdown] slideshow={"slide_type": "notes"}
# This is not the only problem with _Parsing Expression Grammars_. While *PEGs* are expressive and the *packrat* parser for parsing them is simple and intuitive, *PEGs* suffer from a major deficiency for our purposes. *PEGs* are oriented towards language recognition, and it is not clear how to translate an arbitrary *PEG* to a *CFG*. As we mentioned earlier, a naive re-interpretation of a *PEG* as a *CFG* does not work very well. Further, it is not clear what is the exact relation between the class of languages represented by *PEG* and the the class of languages represented by *CFG*. Since our primary focus is *fuzzing* – that is _generation_ of strings – , we next look at _parsers that can accept context-free grammars_.
# + [markdown] slideshow={"slide_type": "fragment"}
# The general idea of *CFG* parser is the following: Peek at the input text for the allowed number of characters, and use these, and our parser state to determine which rules can be applied to complete parsing. We next look at a typical *CFG* parsing algorithm, the Earley Parser.
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=false
# ### The Earley Parser
# + [markdown] slideshow={"slide_type": "notes"}
# The Earley parser is a general parser that is able to parse any arbitrary *CFG*. It was invented by <NAME> \cite{Earley1970} for use in computational linguistics. While its computational complexity is $O(n^3)$ for parsing strings with arbitrary grammars, it can parse strings with unambiguous grammars in $O(n^2)$ time, and all *[LR(k)](https://en.wikipedia.org/wiki/LR_parser)* grammars in linear time ($O(n)$ \cite{Leo1991}). Further improvements – notably handling epsilon rules – were invented by Aycock et al. \cite{Aycock2002}.
# + [markdown] slideshow={"slide_type": "notes"}
# Note that one restriction of our implementation is that the start symbol can have only one alternative in its alternative expressions. This is not a restriction in practice because any grammar with multiple alternatives for its start symbol can be extended with a new start symbol that has the original start symbol as its only choice. That is, given a grammar as below,
#
# ```
# grammar = {
# '<start>': ['<A>', '<B>'],
# ...
# }
# ```
# one may rewrite it as below to conform to the *single-alternative* rule.
# ```
# grammar = {
# '<start>': ['<start_>'],
# '<start_>': ['<A>', '<B>'],
# ...
# }
# ```
# + [markdown] slideshow={"slide_type": "notes"}
# We first implement a simpler parser that is a parser for nearly all *CFGs*, but not quite. In particular, our parser does not understand _epsilon rules_ – rules that derive empty string. We show later how the parser can be extended to handle these.
# + [markdown] slideshow={"slide_type": "fragment"}
# We use the following grammar in our examples below.
# + slideshow={"slide_type": "fragment"}
SAMPLE_GRAMMAR = {
'<start>': ['<A><B>'],
'<A>': ['a<B>c', 'a<A>'],
'<B>': ['b<C>', '<D>'],
'<C>': ['c'],
'<D>': ['d']
}
C_SAMPLE_GRAMMAR = canonical(SAMPLE_GRAMMAR)
# + slideshow={"slide_type": "subslide"}
syntax_diagram(SAMPLE_GRAMMAR)
# + [markdown] slideshow={"slide_type": "notes"}
# The basic idea of Earley parsing is the following
# * Start with the alternative expressions corresponding to the START_SYMBOL. These represent the possible ways to parse the string from a high level. Essentially each expression represents a parsing path. Queue each expression in our set of possible parses of the string. The parsed index of an expression is the part of expression that has already been recognized. In the beginning of parse, the parsed index of all expressions is at the beginning. Further, each letter gets a queue of expressions that recognizes that letter at that point in our parse.
# * Examine our queue of possible parses and check if any of them start with a nonterminal. If it does, then that nonterminal needs to be recognized from the input before the given rule can be parsed. Hence, add the alternative expressions corresponding to the nonterminal to the queue. Do this recursively.
# * At this point, we are ready to advance. Examine the current letter in the input, and select all expressions that have that particular letter at the parsed index. These expressions can now advance one step. Advance these selected expressions by incrementing their parsed index and add them to the queue of expressions in line for recognizing the next input letter.
# * If while doing these things, we find that any of the expressions have finished parsing, we fetch its corresponding nonterminal, and advance all expressions that have that nonterminal at their parsed index.
# * Continue this procedure recursively until all expressions that we have queued for the current letter have been processed. Then start processing the queue for the next letter.
#
# We explain each step in detail with examples in the coming sections.
#
# The parser uses dynamic programming to generate a table containing a _forest of possible parses_ at each letter index – the table contains as many columns as there are letters in the input, and each column contains different parsing rules at various stages of the parse.
#
# For example, given an input `adcd`, the Column 0 would contain the following:
# ```
# <start> : ● <A> <B>
# ```
# which is the starting rule that indicates that we are currently parsing the rule `<start>`, and the parsing state is just before identifying the symbol `<A>`. It would also contain the following which are two alternative paths it could take to complete the parsing.
#
# ```
# <A> : ● a <B> c
# <A> : ● a <A>
# ```
#
# Column 1 would contain the following, which represents the possible completion after reading `a`.
# ```
# <A> : a ● <B> c
# <A> : a ● <A>
# <B> : ● b <C>
# <B> : ● <D>
# <A> : ● a <B> c
# <A> : ● a <A>
# <D> : ● d
# ```
# Column 2 would contain the following after reading `d`
# ```
# <D> : d ●
# <B> : <D> ●
# <A> : a <B> ● c
# ```
# Similarly, Column 3 would contain the following after reading `c`
# ```
# <A> : a <B> c ●
# <start> : <A> ● <B>
# <B> : ● b <C>
# <B> : ● <D>
# <D> : ● d
# ```
# Finally, Column 4 would contain the following after reading `d`, with the `●` at the end of the `<start>` rule indicating that the parse was successful.
# ```
# <D> : d ●
# <B> : <D> ●
# <start> : <A> <B> ●
# ```
#
# As you can see from above, we are essentially filling a table (a table is also called a **chart**) of entries based on each letter we read, and the grammar rules that can be applied. This chart gives the parser its other name -- Chart parsing.
# + [markdown] slideshow={"slide_type": "notes"}
# ### Columns
#
# We define the `Column` first. The `Column` is initialized by its own `index` in the input string, and the `letter` at that index. Internally, we also keep track of the states that are added to the column as the parsing progresses.
# + slideshow={"slide_type": "subslide"}
class Column(object):
def __init__(self, index, letter):
self.index, self.letter = index, letter
self.states, self._unique = [], {}
def __str__(self):
return "%s chart[%d]\n%s" % (self.letter, self.index, "\n".join(
str(state) for state in self.states if state.finished()))
# + [markdown] slideshow={"slide_type": "notes"}
# The `Column` only stores unique `states`. Hence, when a new `state` is `added` to our `Column`, we check whether it is already known.
# + slideshow={"slide_type": "subslide"}
class Column(Column):
def add(self, state):
if state in self._unique:
return self._unique[state]
self._unique[state] = state
self.states.append(state)
state.e_col = self
return self._unique[state]
# + [markdown] slideshow={"slide_type": "notes"}
# ### Items
#
# An item represents a _parse in progress for a specific rule._ Hence the item contains the name of the nonterminal, and the corresponding alternative expression (`expr`) which together form the rule, and the current position of parsing in this expression -- `dot`.
#
#
# **Note.** If you are familiar with [LR parsing](https://en.wikipedia.org/wiki/LR_parser), you will notice that an item is simply an `LR0` item.
# + slideshow={"slide_type": "subslide"}
class Item(object):
def __init__(self, name, expr, dot):
self.name, self.expr, self.dot = name, expr, dot
# + [markdown] slideshow={"slide_type": "notes"}
# We also provide a few convenience methods. The method `finished()` checks if the `dot` has moved beyond the last element in `expr`. The method `advance()` produces a new `Item` with the `dot` advanced one token, and represents an advance of the parsing. The method `at_dot()` returns the current symbol being parsed.
# + slideshow={"slide_type": "subslide"}
class Item(Item):
def finished(self):
return self.dot >= len(self.expr)
def advance(self):
return Item(self.name, self.expr, self.dot + 1)
def at_dot(self):
return self.expr[self.dot] if self.dot < len(self.expr) else None
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is how an item could be used. We first define our item
# + slideshow={"slide_type": "subslide"}
item_name = '<B>'
item_expr = C_SAMPLE_GRAMMAR[item_name][1]
an_item = Item(item_name,tuple(item_expr), 0)
# + [markdown] slideshow={"slide_type": "fragment"}
# To determine where the status of parsing, we use `at_dot()`
# + slideshow={"slide_type": "fragment"}
an_item.at_dot()
# + [markdown] slideshow={"slide_type": "fragment"}
# That is, the next symbol to be parsed is `<D>`
# + [markdown] slideshow={"slide_type": "fragment"}
# If we advance the item, we get another item that represents the finished parsing rule `<B>`.
# + slideshow={"slide_type": "fragment"}
another_item = an_item.advance()
# + slideshow={"slide_type": "fragment"}
another_item.finished()
# + [markdown] slideshow={"slide_type": "notes"}
# ### States
#
# For `Earley` parsing, the state of the parsing is simply one `Item` along with some meta information such as the starting `s_col` and ending column `e_col` for each state. Hence we inherit from `Item` to create a `State`.
# We also store a reference to all the child states (that is, all states that originates from nonterminals in the `expr`). Since we are interested in comparing states, we define `hash()` and `eq()` with the corresponding methods.
# + slideshow={"slide_type": "subslide"}
class State(Item):
def __init__(self, name, expr, dot, s_col):
super().__init__(name, expr, dot)
self.s_col, self.e_col = s_col, None
def __str__(self):
return self.name + ':= ' + ' '.join([
str(p)
for p in [*self.expr[:self.dot], '|', *self.expr[self.dot:]]
]) + "(%d,%d)" % (self.s_col.index, self.e_col.index)
def _t(self):
return (self.name, self.expr, self.dot, self.s_col.index)
def __hash__(self):
return hash(self._t())
def __eq__(self, other):
return self._t() == other._t()
def advance(self):
return State(self.name, self.expr, self.dot + 1, self.s_col)
# + [markdown] slideshow={"slide_type": "subslide"}
# The `State` is used similar to `Item`. The only difference is that it is used along with the `Column` to track the parsing state. For example, we initialize the first column as follows:
# + slideshow={"slide_type": "fragment"}
col_0 = Column(0, None)
item_expr = tuple(*C_SAMPLE_GRAMMAR[START_SYMBOL])
start_state = State(START_SYMBOL, item_expr, 0, col_0)
col_0.add(start_state)
start_state.at_dot()
# + [markdown] slideshow={"slide_type": "fragment"}
# The first column is then updated by using `add()` method of `Column`
# + slideshow={"slide_type": "subslide"}
sym = start_state.at_dot()
for alt in C_SAMPLE_GRAMMAR[sym]:
col_0.add(State(sym, tuple(alt), 0, col_0))
for s in col_0.states:
print(s)
# + [markdown] slideshow={"slide_type": "subslide"} toc-hr-collapsed=false
# ### The Parsing Algorithm
# + [markdown] slideshow={"slide_type": "notes"}
# The _Earley_ algorithm starts by initializing the chart with columns (as many as there are letters in the input). We also seed the first column with a state representing the expression corresponding to the start symbol. In our case, the state corresponds to the start symbol with the `dot` at `0` is represented as below. The `●` symbol represents the parsing status. In this case, we have not parsed anything.
#
# ```
# <start>: ● <A> <B>
# ```
# We pass this partial chart to a method for filling the rest of the parse chart.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(Parser):
def __init__(self, grammar, **kwargs):
super().__init__(grammar, **kwargs)
self.cgrammar = canonical(grammar, letters=True)
# + [markdown] slideshow={"slide_type": "notes"}
# Before starting to parse, we seed the chart with the state representing the ongoing parse of the start symbol.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def chart_parse(self, words, start):
alt = tuple(*self.cgrammar[start])
chart = [Column(i, tok) for i, tok in enumerate([None, *words])]
chart[0].add(State(start, alt, 0, chart[0]))
return self.fill_chart(chart)
# + [markdown] slideshow={"slide_type": "notes"}
# The main parsing loop in `fill_chart()` has three fundamental operations. `predict()`, `scan()`, and `complete()`. We discuss `predict` next.
# + [markdown] slideshow={"slide_type": "notes"}
# ### Predicting States
#
# We have already seeded `chart[0]` with a state `[<A>,<B>]` with `dot` at `0`. Next, given that `<A>` is a nonterminal, we `predict` the possible parse continuations of this state. That is, it could be either `a <B> c` or `A <A>`.
#
# The general idea of `predict()` is as follows: Say you have a state with name `<A>` from the above grammar, and expression containing `[a,<B>,c]`. Imagine that you have seen `a` already, which means that the `dot` will be on `<B>`. Below, is a representation of our parse status. The left hand side of ● represents the portion already parsed (`a`), and the right hand side represents the portion yet to be parsed (`<B> c`).
#
# ```
# <A>: a ● <B> c
# ```
# + [markdown] slideshow={"slide_type": "notes"}
# To recognize `<B>`, we look at the definition of `<B>`, which has different alternative expressions. The `predict()` step adds each of these alternatives to the set of states, with `dot` at `0`.
#
# ```
# <A>: a ● <B> c
# <B>: ● b c
# <B>: ● <D>
# ```
#
# In essence, the `predict()` method, when called with the current nonterminal, fetches the alternative expressions corresponding to this nonterminal, and adds these as predicted _child_ states to the _current_ column.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def predict(self, col, sym, state):
for alt in self.cgrammar[sym]:
col.add(State(sym, tuple(alt), 0, col))
# + [markdown] slideshow={"slide_type": "fragment"}
# To see how to use `predict`, we first construct the 0th column as before, and we assign the constructed column to an instance of the EarleyParser.
# + slideshow={"slide_type": "fragment"}
col_0 = Column(0, None)
col_0.add(start_state)
ep = EarleyParser(SAMPLE_GRAMMAR)
ep.chart = [col_0]
# + [markdown] slideshow={"slide_type": "fragment"}
# It should contain a single state -- `<start> at 0`
# + slideshow={"slide_type": "subslide"}
for s in ep.chart[0].states:
print(s)
# + [markdown] slideshow={"slide_type": "fragment"}
# We apply predict to fill out the 0th column, and the column should contain the possible parse paths.
# + slideshow={"slide_type": "fragment"}
ep.predict(col_0, '<A>', s)
for s in ep.chart[0].states:
print(s)
# + [markdown] slideshow={"slide_type": "notes"}
# ### Scanning Tokens
#
# What if rather than a nonterminal, the state contained a terminal symbol such as a letter? In that case, we are ready to make some progress. For example, consider the second state:
# ```
# <B>: ● b c
# ```
# We `scan` the next column's letter. Say the next token is `b`.
# If the letter matches what we have, then create a new state by advancing the current state by one letter.
#
# ```
# <B>: b ● c
# ```
# This new state is added to the next column (i.e the column that has the matched letter).
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def scan(self, col, state, letter):
if letter == col.letter:
col.add(state.advance())
# + [markdown] slideshow={"slide_type": "fragment"}
# As before, we construct the partial parse first, this time adding a new column so that we can observe the effects of `scan()`
# + slideshow={"slide_type": "fragment"}
ep = EarleyParser(SAMPLE_GRAMMAR)
col_1 = Column(1, 'a')
ep.chart = [col_0, col_1]
ep.predict(col_0, '<A>', s)
# + slideshow={"slide_type": "fragment"}
new_state = ep.chart[0].states[1]
print(new_state)
# + slideshow={"slide_type": "subslide"}
ep.scan(col_1, new_state, 'a')
for s in ep.chart[1].states:
print(s)
# + [markdown] slideshow={"slide_type": "notes"}
# ### Completing Processing
#
# When we advance, what if we actually `complete()` the processing of the current rule? If so, we want to update not just this state, but also all the _parent_ states from which this state was derived.
# For example, say we have states as below.
# ```
# <A>: a ● <B> c
# <B>: b c ●
# ```
# The state `<B>: b c ●` is now complete. So, we need to advance `<A>: a ● <B> c` one step forward.
#
# How do we determine the parent states? Note from `predict` that we added the predicted child states to the _same_ column as that of the inspected state. Hence, we look at the starting column of the current state, with the same symbol `at_dot` as that of the name of the completed state.
#
# For each such parent found, we advance that parent (because we have just finished parsing that non terminal for their `at_dot`) and add the new states to the current column.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def complete(self, col, state):
return self.earley_complete(col, state)
def earley_complete(self, col, state):
parent_states = [
st for st in state.s_col.states if st.at_dot() == state.name
]
for st in parent_states:
col.add(st.advance())
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is an example of completed processing. First we complete the Column 0
# + slideshow={"slide_type": "subslide"}
ep = EarleyParser(SAMPLE_GRAMMAR)
col_1 = Column(1, 'a')
col_2 = Column(2, 'd')
ep.chart = [col_0, col_1,col_2]
ep.predict(col_0, '<A>', s)
for s in ep.chart[0].states:
print(s)
# + [markdown] slideshow={"slide_type": "fragment"}
# Then we use `scan()` to populate Column 1
# + slideshow={"slide_type": "subslide"}
for state in ep.chart[0].states:
if state.at_dot() not in SAMPLE_GRAMMAR:
ep.scan(col_1, state, 'a')
for s in ep.chart[1].states:
print(s)
# + slideshow={"slide_type": "subslide"}
for state in ep.chart[1].states:
if state.at_dot() in SAMPLE_GRAMMAR:
ep.predict(col_1, state.at_dot(), state)
for s in ep.chart[1].states:
print(s)
# + [markdown] slideshow={"slide_type": "fragment"}
# Then we use `scan()` again to populate Column 2
# + slideshow={"slide_type": "subslide"}
for state in ep.chart[1].states:
if state.at_dot() not in SAMPLE_GRAMMAR:
ep.scan(col_2, state, state.at_dot())
for s in ep.chart[2].states:
print(s)
# + [markdown] slideshow={"slide_type": "fragment"}
# Now, we can use `complete()`:
# + slideshow={"slide_type": "subslide"}
for state in ep.chart[2].states:
if state.finished():
ep.complete(col_2, state)
for s in ep.chart[2].states:
print(s)
# + [markdown] slideshow={"slide_type": "notes"}
# ### Filling the Chart
#
# The main driving loop in `fill_chart()` essentially calls these operations in order. We loop over each column in order.
# * For each column, fetch one state in the column at a time, and check if the state is `finished`.
# * If it is, then we `complete()` all the parent states depending on this state.
# * If the state was not finished, we check to see if the state's current symbol `at_dot` is a nonterminal.
# * If it is a nonterminal, we `predict()` possible continuations, and update the current column with these states.
# * If it was not, we `scan()` the next column and advance the current state if it matches the next letter.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def fill_chart(self, chart):
for i, col in enumerate(chart):
for state in col.states:
if state.finished():
self.complete(col, state)
else:
sym = state.at_dot()
if sym in self.cgrammar:
self.predict(col, sym, state)
else:
if i + 1 >= len(chart):
continue
self.scan(chart[i + 1], state, sym)
if self.log:
print(col)
return chart
# + [markdown] slideshow={"slide_type": "subslide"}
# We now can recognize a given string as belonging to a language represented by a grammar.
# + slideshow={"slide_type": "subslide"}
ep = EarleyParser(SAMPLE_GRAMMAR, log=True)
columns = ep.chart_parse('adcd', START_SYMBOL)
# + [markdown] slideshow={"slide_type": "subslide"}
# The chart we printed above only shows completed entries at each index. The parenthesized expression indicates the column just before the first character was recognized, and the ending column.
#
# Notice how the `<start>` nonterminal shows fully parsed status.
# + slideshow={"slide_type": "fragment"}
last_col = columns[-1]
for s in last_col.states:
if s.name == '<start>':
print(s)
# + [markdown] slideshow={"slide_type": "notes"}
# Since `chart_parse()` returns the completed table, we now need to extract the derivation trees.
# + [markdown] slideshow={"slide_type": "notes"}
# ### The Parse Method
#
# For determining how far we have managed to parse, we simply look for the last index from `chart_parse()` where the `start_symbol` was found.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def parse_prefix(self, text):
table = self.chart_parse(text, self.start_symbol)
for col in reversed(table):
states = [st for st in col.states if st.name == self.start_symbol]
if states:
return col.index, states
return -1, []
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is the `parse_prefix()` in action.
# + slideshow={"slide_type": "subslide"}
ep = EarleyParser(SAMPLE_GRAMMAR)
cursor, last_states = ep.parse_prefix('adcd')
print(cursor, [str(s) for s in last_states])
# + [markdown] slideshow={"slide_type": "subslide"}
# The following is adapted from the excellent reference on Earley parsing by [Loup Vaillant](http://loup-vaillant.fr/tutorials/earley-parsing/).
#
# The `parse()` method has to construct the parsed forest. Our chart is a table of states that contains production rules that end at that index. For easier parsing, we switch to a table that contains rules that begin at that index.
# + [markdown] slideshow={"slide_type": "fragment"}
# We define a `reverse()` method to parse and extract the parse forest. We need to extract the parse forest from the chart as we show in the next section.
# + code_folding=[] slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def reverse(self, table):
f_table = [Column(c.index, c.letter) for c in table]
for col in table:
finished = [s for s in col.states if s.finished()]
for s in finished:
f_table[s.s_col.index].states.append(s)
return f_table
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is the reversed table
# + slideshow={"slide_type": "subslide"}
ep = EarleyParser(SAMPLE_GRAMMAR)
reversed_table = ep.reverse(columns)
for col in reversed_table:
print(col)
# + [markdown] slideshow={"slide_type": "subslide"}
# Our `parse()` method is as follows. It depends on two methods `parse_forest()` and `extract_trees()` that will be defined next.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def parse(self, text):
cursor, states = self.parse_prefix(text)
if cursor != len(text):
return []
table = self.chart_parse(text, self.start_symbol)
f_table = self.reverse(table)
start = next(s for s in states if s.finished())
return self.extract_trees(self.parse_forest(f_table, start))
# + [markdown] slideshow={"slide_type": "notes"}
# ### Parsing Paths
#
# The `parse_paths()` method tries to unify the given expression in `named_expr` with the parsed string. For that, it extracts the first symbol in `named_expr` and checks if it is a terminal symbol. If it is, then it checks the chart to see if the letter corresponding to the position matches the terminal symbol. If it does, advance our start index by the length of the symbol.
#
# If the symbol was a nonterminal symbol, then we retrieve the parsed states at the current start index that correspond to the nonterminal symbol, and collect the end index. These are the start indexes for the remaining expression.
#
# Given our list of start indexes, we obtain the parse paths from the remaining expression. If we can obtain any, then we return the parse paths. If not, we return an empty list.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def parse_paths(self, named_expr, chart, frm, til):
var, *expr = named_expr
starts = None
if var not in self.cgrammar:
starts = ([(var, frm + len(var))]
if frm < til and chart[frm + 1].letter == var else [])
else:
starts = [(s, s.e_col.index) for s in chart[frm].states
if s.name == var]
paths = []
for state, start in starts:
if not expr:
paths.extend([[state]] if start == til else [])
else:
res = self.parse_paths(expr, chart, start, til)
paths.extend([[state] + r for r in res])
return paths
# + [markdown] slideshow={"slide_type": "subslide"}
# Here is the `parse_paths()` in action
# + slideshow={"slide_type": "fragment"}
print(SAMPLE_GRAMMAR['<start>'])
ep = EarleyParser(SAMPLE_GRAMMAR)
completed_start = last_states[0]
reversed_table = ep.reverse(columns)
paths = ep.parse_paths(completed_start.expr, reversed_table, 0, 4)
for path in paths:
print([str(s) for s in path])
# + [markdown] slideshow={"slide_type": "subslide"}
# That is, the parse path for `<start>` given the input `adcd` included recognizing the expression `<A><B>`. This was recognized by the two states: `<A>` from input(0) to input(2) which further involved recognizing the rule `a<B>c`, and the next state `<B>` from input(3) which involved recognizing the rule `<D>`.
# + [markdown] slideshow={"slide_type": "notes"}
# ### Parsing Forests
#
# The `parse_forest()` method takes the state which represents the completed parse, and determines the possible ways that its expressions corresponded to the parsed expression. For example, say we are parsing `1+2+3`, and the state has `[<expr>,+,<expr>]` in `expr`. It could have been parsed as either `[{<expr>:1+2},+,{<expr>:3}]` or `[{<expr>:1},+,{<expr>:2+3}]`.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def parse_forest(self, chart, state):
if not state.expr:
return (state.name, [])
pathexprs = self.parse_paths(state.expr, chart, state.s_col.index,
state.e_col.index)
paths_ = []
for pathexpr in pathexprs:
pathexpr_ = []
for varexpr in pathexpr:
completion = (self.parse_forest(chart, varexpr) if isinstance(
varexpr, State) else (varexpr, []))
pathexpr_.append(completion)
paths_.append(pathexpr_)
return (state.name, paths_)
# + slideshow={"slide_type": "subslide"}
ep = EarleyParser(SAMPLE_GRAMMAR)
reversed_table = ep.reverse(columns)
result = ep.parse_forest(reversed_table, last_states[0])
result
# + [markdown] slideshow={"slide_type": "notes"}
# ### Extracting Trees
# + [markdown] slideshow={"slide_type": "notes"}
# What we have from `parse_forest()` is a forest of trees. We need to extract a single tree from that forest. That is accomplished as follows.
# + [markdown] slideshow={"slide_type": "skip"}
# (For now, we return the first available derivation tree. To do that, we need to extract the parse forest from the state corresponding to `start`.)
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def extract_a_tree(self, forest_node):
name, paths = forest_node
if not paths:
return (name, [])
return (name, [self.extract_a_tree(p) for p in paths[0]])
def extract_trees(self, forest):
return [self.extract_a_tree(forest)]
# + [markdown] slideshow={"slide_type": "skip"}
# We now verify that our parser can parse a given expression.
# + slideshow={"slide_type": "subslide"}
A3_GRAMMAR = {
"<start>": ["<expr>"],
"<expr>": ["<expr>+<expr>", "<expr>-<expr>", "(<expr>)", "<integer>"],
"<integer>": ["<digit><integer>", "<digit>"],
"<digit>": ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"]
}
# + slideshow={"slide_type": "subslide"}
syntax_diagram(A3_GRAMMAR)
# + slideshow={"slide_type": "subslide"}
mystring = '(1+24)-33'
parser = EarleyParser(A3_GRAMMAR)
for tree in parser.parse(mystring):
assert tree_to_string(tree) == mystring
display_tree(tree)
# + [markdown] slideshow={"slide_type": "fragment"}
# We now have a complete parser that can parse almost arbitrary *CFG*. There remains a small corner to fix -- the case of epsilon rules as we will see later.
# + [markdown] slideshow={"slide_type": "skip"}
# ### Ambiguous Parsing
# + [markdown] slideshow={"slide_type": "skip"}
# Ambiguous grammars are grammars that can produce multiple derivation trees for some given string. For example, the `A3_GRAMMAR` can parse `1+2+3` in two different ways – `[1+2]+3` and `1+[2+3]`.
#
# Extracting a single tree might be reasonable for unambiguous parses. However, what if the given grammar produces ambiguity when given a string? We need to extract all derivation trees in that case. We enhance our `extract_trees()` method to extract multiple derivation trees.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def extract_trees(self, forest_node):
name, paths = forest_node
if not paths:
return [(name, [])]
results = []
for path in paths:
ptrees = zip(*[self.extract_trees(p) for p in path])
results.extend([(name, p) for p in ptrees])
return results
# + [markdown] slideshow={"slide_type": "skip"}
# As before, we verify that everything works.
# + slideshow={"slide_type": "skip"}
mystring = '12+23-34'
parser = EarleyParser(A1_GRAMMAR)
for tree in parser.parse(mystring):
assert mystring == tree_to_string(tree)
display_tree(tree)
# + [markdown] slideshow={"slide_type": "fragment"}
# One can also use a `GrammarFuzzer` to verify that everything works.
# + slideshow={"slide_type": "subslide"}
gf = GrammarFuzzer(A1_GRAMMAR)
for i in range(5):
s = gf.fuzz()
print(i, s)
for tree in parser.parse(s):
assert tree_to_string(tree) == s
# + [markdown] slideshow={"slide_type": "skip"} toc-hr-collapsed=true
# ### The Aycock Epsilon Fix
#
# While parsing, one often requires to know whether a given nonterminal can derive an empty string. For example, in the following grammar A can derive an empty string, while B can't. The nonterminals that can derive an empty string are called _nullable_ nonterminals. For example, in the below grammar `E_GRAMMAR_1`, `<A>` is _nullable_, and since `<A>` is one of the alternatives of `<start>`, `<start>` is also _nullable_. But `<B>` is not _nullable_.
# + slideshow={"slide_type": "skip"}
E_GRAMMAR_1 = {
'<start>': ['<A>', '<B>'],
'<A>': ['a', ''],
'<B>': ['b']
}
# + [markdown] slideshow={"slide_type": "notes"}
# One of the problems with the original Earley implementation is that it does not handle rules that can derive empty strings very well. For example, the given grammar should match `a`
# + slideshow={"slide_type": "subslide"}
EPSILON = ''
E_GRAMMAR = {
'<start>': ['<S>'],
'<S>': ['<A><A><A><A>'],
'<A>': ['a', '<E>'],
'<E>': [EPSILON]
}
# + slideshow={"slide_type": "subslide"}
syntax_diagram(E_GRAMMAR)
# + slideshow={"slide_type": "subslide"}
mystring = 'a'
parser = EarleyParser(E_GRAMMAR)
trees = parser.parse(mystring)
print(trees)
# + [markdown] slideshow={"slide_type": "skip"}
# Aycock et al.\cite{Aycock2002} suggests a simple fix. Their idea is to pre-compute the `nullable` set and use it to advance the `nullable` states. However, before we do that, we need to compute the `nullable` set. The `nullable` set consists of all nonterminals that can derive an empty string.
# + [markdown] slideshow={"slide_type": "skip"}
# Computing the `nullable` set requires expanding each production rule in the grammar iteratively and inspecting whether a given rule can derive the empty string. Each iteration needs to take into account new terminals that have been found to be `nullable`. The procedure stops when we obtain a stable result. This procedure can be abstracted into a more general method `fixpoint`.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Fixpoint
#
# A `fixpoint` of a function is an element in the function's domain such that it is mapped to itself. For example, 1 is a `fixpoint` of square root because `squareroot(1) == 1`.
#
# (We use `str` rather than `hash` to check for equality in `fixpoint` because the data structure `set`, which we would like to use as an argument has a good string representation but is not hashable).
# + slideshow={"slide_type": "subslide"}
def fixpoint(f):
def helper(arg):
while True:
sarg = str(arg)
arg_ = f(arg)
if str(arg_) == sarg:
return arg
arg = arg_
return helper
# + [markdown] slideshow={"slide_type": "skip"}
# Remember `my_sqrt()` from [the first chapter](Intro_Testing.ipynb)? We can define `my_sqrt()` using fixpoint.
# + slideshow={"slide_type": "subslide"}
def my_sqrt(x):
@fixpoint
def _my_sqrt(approx):
return (approx + x / approx) / 2
return _my_sqrt(1)
# + slideshow={"slide_type": "fragment"}
my_sqrt(2)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Nullable
#
# Similarly, we can define `nullable` using `fixpoint`. We essentially provide the definition of a single intermediate step. That is, assuming that `nullables` contain the current `nullable` nonterminals, we iterate over the grammar looking for productions which are `nullable` -- that is, productions where the entire sequence can yield an empty string on some expansion.
# + [markdown] slideshow={"slide_type": "skip"}
# We need to iterate over the different alternative expressions and their corresponding nonterminals. Hence we define a `rules()` method converts our dictionary representation to this pair format.
# + slideshow={"slide_type": "skip"}
def rules(grammar):
return [(key, choice)
for key, choices in grammar.items()
for choice in choices]
# + [markdown] slideshow={"slide_type": "skip"}
# The `terminals()` method extracts all terminal symbols from a `canonical` grammar representation.
# + slideshow={"slide_type": "skip"}
def terminals(grammar):
return set(token
for key, choice in rules(grammar)
for token in choice if token not in grammar)
# + slideshow={"slide_type": "subslide"}
def nullable_expr(expr, nullables):
return all(token in nullables for token in expr)
# + slideshow={"slide_type": "subslide"}
def nullable(grammar):
productions = rules(grammar)
@fixpoint
def nullable_(nullables):
for A, expr in productions:
if nullable_expr(expr, nullables):
nullables |= {A}
return (nullables)
return nullable_({EPSILON})
# + slideshow={"slide_type": "subslide"}
for key, grammar in {
'E_GRAMMAR': E_GRAMMAR,
'E_GRAMMAR_1': E_GRAMMAR_1
}.items():
print(key, nullable(canonical(grammar)))
# + [markdown] slideshow={"slide_type": "notes"}
# So, once we have the `nullable` set, all that we need to do is, after we have called `predict` on a state corresponding to a nonterminal, check if it is `nullable` and if it is, advance and add the state to the current column.
# + slideshow={"slide_type": "subslide"}
class EarleyParser(EarleyParser):
def __init__(self, grammar, **kwargs):
super().__init__(grammar, **kwargs)
self.cgrammar = canonical(grammar, letters=True)
self.epsilon = nullable(self.cgrammar)
def predict(self, col, sym, state):
for alt in self.cgrammar[sym]:
col.add(State(sym, tuple(alt), 0, col))
if sym in self.epsilon:
col.add(state.advance())
# + slideshow={"slide_type": "subslide"}
mystring = 'a'
parser = EarleyParser(E_GRAMMAR)
for tree in parser.parse(mystring):
display_tree(tree)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### More Earley Parsing
#
# A number of other optimizations exist for Earley parsers. A fast industrial strength Earley parser implementation is the [Marpa parser](https://jeffreykegler.github.io/Marpa-web-site/). Further, Earley parsing need not be restricted to character data. One may also parse streams (audio and video streams) \cite{qi2018generalized} using a generalized Earley parser.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Testing the Parsers
#
# While we have defined two parser variants, it would be nice to have some confirmation that our parses work well. While it is possible to formally prove that they work, it is much more satisfying to generate random grammars, their corresponding strings, and parse them using the same grammar.
# + slideshow={"slide_type": "subslide"}
def prod_line_grammar(nonterminals, terminals):
g = {
'<start>': ['<symbols>'],
'<symbols>': ['<symbol><symbols>', '<symbol>'],
'<symbol>': ['<nonterminals>', '<terminals>'],
'<nonterminals>': ['<lt><alpha><gt>'],
'<lt>': ['<'],
'<gt>': ['>'],
'<alpha>': nonterminals,
'<terminals>': terminals
}
if not nonterminals:
g['<nonterminals>'] = ['']
return g
# + slideshow={"slide_type": "subslide"}
syntax_diagram(prod_line_grammar(["A", "B", "C"], ["1", "2", "3"]))
# + slideshow={"slide_type": "subslide"}
def make_rule(nonterminals, terminals, num_alts):
prod_grammar = prod_line_grammar(nonterminals, terminals)
gf = GrammarFuzzer(prod_grammar, min_nonterminals=3, max_nonterminals=5)
name = "<%s>" % ''.join(random.choices(string.ascii_uppercase, k=3))
return (name, [gf.fuzz() for _ in range(num_alts)])
# + slideshow={"slide_type": "fragment"}
make_rule(["A", "B", "C"], ["1", "2", "3"], 3)
# + slideshow={"slide_type": "subslide"}
def make_grammar(num_symbols=3, num_alts=3):
a = list(string.ascii_lowercase)
grammar = {}
name = None
for _ in range(num_symbols):
name, expansions = make_rule([k[1:-1]
for k in grammar.keys()], a, num_alts)
grammar[name] = expansions
grammar[START_SYMBOL] = [name]
# assert is_valid_grammar(grammar)
return grammar
# + slideshow={"slide_type": "subslide"}
make_grammar()
# + [markdown] slideshow={"slide_type": "fragment"}
# Now we verify if our arbitrary grammars can be used by the Earley parser.
# + slideshow={"slide_type": "subslide"}
for i in range(5):
my_grammar = make_grammar()
print(my_grammar)
parser = EarleyParser(my_grammar)
mygf = GrammarFuzzer(my_grammar)
s = mygf.fuzz()
print(s)
for tree in parser.parse(s):
assert tree_to_string(tree) == s
display_tree(tree)
# + [markdown] slideshow={"slide_type": "subslide"}
# With this, we have completed both implementation and testing of *arbitrary* CFG, which can now be used along with `LangFuzzer` to generate better fuzzing inputs.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Background
#
#
# Numerous parsing techniques exist that can parse a given string using a
# given grammar, and produce corresponding derivation tree or trees. However,
# some of these techniques work only on specific classes of grammars.
# These classes of grammars are named after the specific kind of parser
# that can accept grammars of that category. That is, the upper bound for
# the capabilities of the parser defines the grammar class named after that
# parser.
#
# The *LL* and *LR* parsing are the main traditions in parsing. Here, *LL* means left-to-right, leftmost derivation, and it represents a top-down approach. On the other hand, and LR (left-to-right, rightmost derivation) represents a bottom up approach. Another way to look at it is that LL parsers compute the derivation tree incrementally in *pre-order* while LR parsers compute the derivation tree in *post-order* \cite{pingali2015graphical}).
#
# Different classes of grammars differ in the features that are available to
# the user for writing a grammar of that class. That is, the corresponding
# kind of parser will be unable to parse a grammar that makes use of more
# features than allowed. For example, the `A2_GRAMMAR` is an *LL*
# grammar because it lacks left recursion, while `A1_GRAMMAR` is not an
# *LL* grammar. This is because an *LL* parser parses
# its input from left to right, and constructs the leftmost derivation of its
# input by expanding the nonterminals it encounters. If there is a left
# recursion in one of these rules, an *LL* parser will enter an infinite loop.
#
# Similarly, a grammar is LL(k) if it can be parsed by an LL parser with k lookahead token, and LR(k) grammar can only be parsed with LR parser with at least k lookahead tokens. These grammars are interesting because both LL(k) and LR(k) grammars have $O(n)$ parsers, and can be used with relatively restricted computational budget compared to other grammars.
#
# The languages for which one can provide an *LL(k)* grammar is called *LL(k)* languages (where k is the minimum lookahead required). Similarly, *LR(k)* is defined as the set of languages that have an *LR(k)* grammar. In terms of languages, LL(k) $\subset$ LL(k+1) and LL(k) $\subset$ LR(k), and *LR(k)* $=$ *LR(1)*. All deterministic *CFLs* have an *LR(1)* grammar. However, there exist *CFLs* that are inherently ambiguous \cite{ogden1968helpful}, and for these, one can't provide an *LR(1)* grammar.
#
# The other main parsing algorithms for *CFGs* are GLL \cite{scott2010gll}, GLR \cite{tomita1987efficient,tomita2012generalized}, and CYK \cite{grune1990parsing}.
# The ALL(\*) (used by ANTLR) on the other hand is a grammar representation that uses *Regular Expression* like predicates (similar to advanced PEGs – see [Exercise](#Exercise-3:-PEG-Predicates)) rather than a fixed lookahead. Hence, ALL(\*) can accept a larger class of grammars than CFGs.
#
# In terms of computational limits of parsing, the main CFG parsers have a complexity of $O(n^3)$ for arbitrary grammars. However, parsing with arbitrary *CFG* is reducible to boolean matrix multiplication \cite{Valiant1975} (and the reverse \cite{Lee2002}). This is at present bounded by $O(2^{23728639}$) \cite{LeGall2014}. Hence, worse case complexity for parsing arbitrary CFG is likely to remain close to cubic.
#
# Regarding PEGs, the actual class of languages that is expressible in *PEG* is currently unknown. In particular, we know that *PEGs* can express certain languages such as $a^n b^n c^n$. However, we do not know if there exist *CFLs* that are not expressible with *PEGs*. In Section 2.3, we provided an instance of a counter-intuitive PEG grammar. While important for our purposes (we use grammars for generation of inputs) this is not a criticism of parsing with PEGs. PEG focuses on writing grammars for recognizing a given language, and not necessarily in interpreting what language an arbitrary PEG might yield. Given a Context-Free Language to parse, it is almost always possible to write a grammar for it in PEG, and given that 1) a PEG can parse any string in $O(n)$ time, and 2) at present we know of no CFL that can't be expressed as a PEG, and 3) compared with *LR* grammars, a PEG is often more intuitive because it allows top-down interpretation, when writing a parser for a language, PEGs should be under serious consideration.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ## Lessons Learned
#
# * Grammars can be used to generate derivation trees for a given string.
# * Parsing Expression Grammars are intuitive, and easy to implement, but require care to write.
# * Earley Parsers can parse arbitrary Context Free Grammars.
# * How to generate a pool of fragments using the Langfuzz approach, and use it to generate nearly valid strings.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ## Next Steps
#
# * [Use the automatic grammar miner for obtaining a grammar](GrammarMiner.ipynb)
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "skip"} toc-hr-collapsed=false
# ## Exercises
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# ### Exercise 1: An Alternative Packrat
#
# In the _Packrat_ parser, we showed how one could implement a simple _PEG_ parser. That parser kept track of the current location in the text using an index. Can you modify the parser so that it simply uses the current substring rather than tracking the index? That is, it should no longer have the `at` parameter.
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# **Solution.** Here is a possible solution:
# + slideshow={"slide_type": "subslide"} solution2="hidden"
class PackratParser(Parser):
def parse_prefix(self, text):
txt, res = self.unify_key(self.start_symbol, text)
return len(txt), [res]
def parse(self, text):
remain, res = self.parse_prefix(text)
if remain:
raise SyntaxError("at " + res)
return res
def unify_rule(self, rule, text):
results = []
for token in rule:
text, res = self.unify_key(token, text)
if res is None:
return text, None
results.append(res)
return text, results
def unify_key(self, key, text):
if key not in self.cgrammar:
if text.startswith(key):
return text[len(key):], (key, [])
else:
return text, None
for rule in self.cgrammar[key]:
text_, res = self.unify_rule(rule, text)
if res:
return (text_, (key, res))
return text, None
# + button=false code_folding=[] new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution2="hidden"
mystring = "1 + (2 * 3)"
for tree in PackratParser(EXPR_GRAMMAR).parse(mystring):
assert tree_to_string(tree) == mystring
display_tree(tree)
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
# ### Exercise 2: More PEG Syntax
#
# The _PEG_ syntax provides a few notational conveniences reminiscent of regular expressions. For example, it supports the following operators (letters `T` and `A` represents tokens that can be either terminal or nonterminal. `ε` is an empty string, and `/` is the ordered choice operator similar to the non-ordered choice operator `|`):
#
# * `T?` represents an optional greedy match of T and `A := T?` is equivalent to `A := T/ε`.
# * `T*` represents zero or more greedy matches of `T` and `A := T*` is equivalent to `A := T A/ε`.
# * `T+` represents one or more greedy matches – equivalent to `TT*`
#
# If you look at the three notations above, each can be represented in the grammar in terms of basic syntax.
# Remember the exercise from [the chapter on grammars](Grammars.ipynb) that developed `define_ex_grammar()` that can represent grammars as Python code? extend `define_ex_grammar()` to `define_peg()` to support the above notational conveniences. The decorator should rewrite a given grammar that contain these notations to an equivalent grammar in basic syntax.
# + [markdown] button=false heading_collapsed=true new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution_first=true
# ### Exercise 3: PEG Predicates
#
# Beyond these notational conveniences, it also supports two predicates that can provide a powerful lookahead facility that does not consume any input.
#
# * `T&A` represents an _And-predicate_ that matches `T` if `T` is matched, and it is immediately followed by `A`
# * `T!A` represents a _Not-predicate_ that matches `T` if `T` is matched, and it is *not* immediately followed by `A`
#
# Implement these predicates in our _PEG_ parser.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# ### Exercise 4: Earley Fill Chart
#
# In the `Earley Parser`, `Column` class, we keep the states both as a `list` and also as a `dict` even though `dict` is ordered. Can you explain why?
#
# **Hint**: see the `fill_chart` method.
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# **Solution.** Python allows us to append to a list in flight, while a dict, eventhough it is ordered does not allow that facility.
#
# That is, the following will work
#
# ```python
# values = [1]
# for v in values:
# values.append(v*2)
# ```
#
# However, the following will result in an error
# ```python
# values = {1:1}
# for v in values:
# values[v*2] = v*2
# ```
#
# In the `fill_chart`, we make use of this facility to modify the set of states we are iterating on, on the fly.
# + [markdown] slideshow={"slide_type": "skip"}
# ### Exercise 5: Leo Parser
#
# One of the problems with the original Earley parser is that while it can parse strings using arbitrary _Context Free Gramamrs_, its performance on right-recursive grammars is quadratic. That is, it takes $O(n^2)$ runtime and space for parsing with right-recursive grammars. For example, consider the parsing of the following string by two different grammars `LR_GRAMMAR` and `RR_GRAMMAR`.
# + slideshow={"slide_type": "fragment"}
mystring = 'aaaaaa'
# + [markdown] slideshow={"slide_type": "skip"}
# We enable logging to see the parse progress
# + slideshow={"slide_type": "skip"}
result = EarleyParser(LR_GRAMMAR, log=True).parse(mystring)
# + slideshow={"slide_type": "skip"}
result = EarleyParser(RR_GRAMMAR, log=True).parse(mystring)
# + [markdown] slideshow={"slide_type": "skip"}
# As can be seen from the parsing log for each letter, the number of states with representation `<A>:= a <A> | (i, j)` increases at each stage, and these are simply a left over from the previous letter. They do not contribute anything more to the parse other than to simply complete these entries. However, they take up space, and require resources for inspection, contributing a factor of `n` in analysis.
#
# Jo<NAME> \cite{Leo1991} found that this inefficiency can be avoided by detecting right recursion. The idea is that before starting the `completion` step, check whether the current item has a _deterministic reduction path_. If such a path exists, add a copy of the topmost element of the _deteministic reduction path_ to the current column, and return. If not, perform the original `completion` step.
#
# Finding a _deterministic reduction path_ is as follows:
#
# Given a complete state, represented by `<A> := ... | (i)` where `(i)` is the starting column, there is a _deterministic reduction path_ above it if two criteria are satisfied.
# * There exist an item `<B> := ... <A> | (k)` in the current column.
# * It should have a _single_ predecessor of the form `<B> := ... | <A> (k)` in the current column.
# Following this chain (looking for the link above `<B> := ... <A> | (k)`), the topmost item is the item that does not have a parent.
#
# See [Loup Vaillant](http://loup-vaillant.fr/tutorials/earley-parsing/right-recursion) for further information.
# + slideshow={"slide_type": "skip"}
class State(State):
def __init__(self, name, expr, dot, s_col, tag=None):
super().__init__(name, expr, dot, s_col)
self.tag = tag
def copy(self, tag=None):
return State(self.name, self.expr, self.dot, self.s_col, tag)
def __str__(self):
init = "%s %s" % (self.tag or " ", self.name)
return init + ':= ' + ' '.join([
str(p)
for p in [*self.expr[:self.dot], '|', *self.expr[self.dot:]]
]) + "(%d,%d)" % (self.s_col.index, self.e_col.index)
# + slideshow={"slide_type": "skip"}
class LeoParser(EarleyParser):
def complete(self, col, state):
return self.leo_complete(col, state)
def leo_complete(self, col, state):
detred = self.deterministic_reduction(state)
if detred:
col.add(detred.copy())
else:
self.earley_complete(col, state)
def deterministic_reduction(self, state):
...
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# Can you implement the `deterministic_reduction` method to obtain the top most element?
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# **Solution.** Here is a possible solution:
# + slideshow={"slide_type": "skip"} solution2="hidden"
from functools import reduce
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden"
# The `splitlst` takes a predicate and an iterable, and splits the iterable into two lists. The first list contains all items where the predicate is `True`, and the second list contains all items where the predicate was `False`.
# + slideshow={"slide_type": "fragment"} solution2="hidden"
def splitlst(predicate, iterable):
return reduce(lambda res, e: res[predicate(e)].append(e)
or res, iterable, ([], []))
# + slideshow={"slide_type": "subslide"} solution2="hidden"
class LeoParser(LeoParser):
def check_single_item(self, st, remain):
res = [
s for s in remain if s.name == st.name and s.expr == st.expr
and s.s_col.index == st.s_col.index and s.dot == (st.dot - 1)
]
return len(res) == 1
@lru_cache(maxsize=None)
def get_above(self, state):
remain, finished = splitlst(lambda s: s.finished(), state.s_col.states)
res = [
st for st in finished
if len(st.expr) > 1 and state.name == st.expr[-1]
]
vals = [st for st in res if self.check_single_item(st, remain)]
if vals:
assert len(vals) == 1
return vals[0]
return None
def deterministic_reduction(self, state):
st = state
while True:
_st = self.get_above(st)
if not _st:
break
st = _st
return st if st != state else None
def complete(self, col, state):
return self.leo_complete(col, state)
def leo_complete(self, col, state):
detred = self.deterministic_reduction(state)
if detred:
col.add(detred.copy(state.name))
else:
self.earley_complete(col, state)
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# Now, both LR and RR grammars should work within $O(n)$ bounds.
# + slideshow={"slide_type": "subslide"} solution2="hidden"
result = LeoParser(RR_GRAMMAR, log=True).parse(mystring)
# + slideshow={"slide_type": "subslide"} solution2="hidden"
result = LeoParser(LR_GRAMMAR, log=True).parse(mystring)
for tree in result:
print(tree_to_string(tree))
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# __Advanced:__ We have fixed the complexity bounds. However, because we are saving only the top most item of a right recursion, we need to fix our `parse_forest` and `extract_a_tree` to be aware of our fix. Can you fix both?
#
# __Hint:__ When you start extracting the parse trees, any time you see a tagged state, look at its end point `e_col.index`. Until you reach this end point, you can carry along this state for each token you see, with the starting column suitably adjusted. Does your solution work for the following grammar?
# + slideshow={"slide_type": "fragment"} solution2="hidden"
RR_GRAMMAR2 = {
'<start>': ['<A>'],
'<A>': ['ab<A>', ''],
}
mystring2 = 'abababab'
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# How about this one? (Does this obey the requirements?)
# + slideshow={"slide_type": "fragment"} solution2="hidden" solution2_first=true
RR_GRAMMAR3 = {
'<start>': ['<A>'],
'<A>': ['ab<B>', ''],
'<B>': ['<A>'],
}
mystring3 = 'abababab'
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# How about this?
# + slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
RR_GRAMMAR3 = {
'<start>': ['<A>'],
'<A>': ['a<B>', ''],
'<B>': ['b<A>'],
}
mystring3 = 'abababab'
# + slideshow={"slide_type": "subslide"} solution2="hidden"
class Tagged:
def __init__(self, tbl):
self.col, self.tbl = [{} for _ in tbl], tbl
@lru_cache(maxsize=None)
def get(self, frm, var):
if frm == -1:
return []
def filter_states(ends):
lst = []
for (state, e) in ends:
if e > frm and state.name == var:
sc = state.copy()
sc.s_col, sc.e_col = self.tbl[frm], self.tbl[e]
lst.append((sc, e))
return lst
lst = filter_states(self.get(frm - 1, var))
st_dict = self.col[frm]
ends = [(state, s) for state in st_dict for s in st_dict[state]
if s > frm]
return lst + filter_states(ends)
# + slideshow={"slide_type": "subslide"} solution2="hidden"
class LeoParser(LeoParser):
def parse(self, text):
cursor, states = self.parse_prefix(text)
if cursor != len(text):
return []
table = self.chart_parse(text, self.start_symbol)
f_table = self.reverse(table)
self.tagged_array = Tagged(f_table)
for i, col in enumerate(f_table):
tcol = self.tagged_array.col[i]
for state in col.states:
if state.tag:
if state not in tcol:
tcol[state] = set()
tcol[state].add(state.e_col.index)
start = next(s for s in states if s.finished())
return self.extract_trees(self.parse_forest(f_table, start))
def parse_paths(self, expr_, chart, frm, til):
var, *expr = expr_
ends = None
if var not in self.cgrammar:
ends = ([(var, frm + len(var))]
if frm < til and chart[frm + 1].letter == var else [])
else:
tagged_ends = self.tagged_array.get(frm, var)
ends = [(s, s.e_col.index) for s in chart[frm].states
if s.name == var and not s.tag] + tagged_ends
paths = []
for state, end in ends:
if not expr:
paths.extend([[state]] if end == til else [])
else:
res = self.parse_paths(expr, chart, end, til)
paths.extend([[state] + r for r in res])
return paths
# + slideshow={"slide_type": "subslide"} solution2="hidden"
p = LeoParser(RR_GRAMMAR)
for tree in p.parse(mystring):
assert tree_to_string(tree) == mystring
display_tree(tree)
# + slideshow={"slide_type": "fragment"} solution2="hidden"
p = LeoParser(RR_GRAMMAR2)
for tree in p.parse(mystring2):
assert tree_to_string(tree) == mystring2
display_tree(tree)
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# ### Exercise 6: First Set of a Nonterminal
#
# We previously gave a way to extract a the `nullable` (epsilon) set, which is often used for parsing.
# Along with `nullable`, parsing algorithms often use two other sets [`first` and `follow`](https://en.wikipedia.org/wiki/Canonical_LR_parser#FIRST_and_FOLLOW_sets).
# The first set of a terminal symbol is itself, and the first set of a nonterminal is composed of terminal symbols that can come at the beginning of any derivation
# of that nonterminal. The first set of any nonterminal that can derive the empty string should contain `EPSILON`. For example, using our `A1_GRAMMAR`, the first set of both `<expr>` and `<start>` is `{0,1,2,3,4,5,6,7,8,9}`. The extraction first set for any self-recursive nonterminal is simple enough. One simply has to recursively compute the first set of the first element of its choice expressions. The computation of `first` set for a self-recursive nonterminal is tricky. One has to recursively compute the first set until one is sure that no more terminals can be added to the first set.
#
# Can you implement the `first` set using our `fixpoint()` decorator?
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# **Solution.** The first set of all terminals is the set containing just themselves. So we initialize that first. Then we update the first set with rules that derive empty strings.
# + slideshow={"slide_type": "fragment"} solution2="hidden"
def firstset(grammar, nullable):
first = {i: {i} for i in terminals(grammar)}
for k in grammar:
first[k] = {EPSILON} if k in nullable else set()
return firstset_((rules(grammar), first, nullable))[1]
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden"
# Finally, we rely on the `fixpoint` to update the first set with the contents of the current first set until the first set stops changing.
# + slideshow={"slide_type": "subslide"} solution2="hidden"
def first_expr(expr, first, nullable):
tokens = set()
for token in expr:
tokens |= first[token]
if token not in nullable:
break
return tokens
# + slideshow={"slide_type": "fragment"} solution2="hidden"
@fixpoint
def firstset_(arg):
(rules, first, epsilon) = arg
for A, expression in rules:
first[A] |= first_expr(expression, first, epsilon)
return (rules, first, epsilon)
# + slideshow={"slide_type": "subslide"} solution2="hidden"
firstset(canonical(A1_GRAMMAR), EPSILON)
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# ### Exercise 7: Follow Set of a Nonterminal
#
# The follow set definition is similar to the first set. The follow set of a nonterminal is the set of terminals that can occur just after that nonterminal is used in any derivation. The follow set of the start symbol is `EOF`, and the follow set of any nonterminal is the super set of first sets of all symbols that come after it in any choice expression.
#
# For example, the follow set of `<expr>` in `A1_GRAMMAR` is the set `{EOF, +, -}`.
#
# As in the previous exercise, implement the `followset()` using the `fixpoint()` decorator.
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# **Solution.** The implementation of `followset()` is similar to `firstset()`. We first initialize the follow set with `EOF`, get the epsilon and first sets, and use the `fixpoint()` decorator to iteratively compute the follow set until nothing changes.
# + slideshow={"slide_type": "fragment"} solution2="hidden"
EOF = '\0'
# + slideshow={"slide_type": "fragment"} solution2="hidden"
def followset(grammar, start):
follow = {i: set() for i in grammar}
follow[start] = {EOF}
epsilon = nullable(grammar)
first = firstset(grammar, epsilon)
return followset_((grammar, epsilon, first, follow))[-1]
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden"
# Given the current follow set, one can update the follow set as follows:
# + slideshow={"slide_type": "subslide"} solution2="hidden"
@fixpoint
def followset_(arg):
grammar, epsilon, first, follow = arg
for A, expression in rules(grammar):
f_B = follow[A]
for t in reversed(expression):
if t in grammar:
follow[t] |= f_B
f_B = f_B | first[t] if t in epsilon else (first[t] - {EPSILON})
return (grammar, epsilon, first, follow)
# + slideshow={"slide_type": "subslide"} solution2="hidden"
followset(canonical(A1_GRAMMAR), START_SYMBOL)
# + [markdown] slideshow={"slide_type": "skip"} toc-hr-collapsed=true
# ### Exercise 8: A LL(1) Parser
#
# As we mentioned previously, there exist other kinds of parsers that operate left-to-right with right most derivation (*LR(k)*) or left-to-right with left most derivation (*LL(k)*) with _k_ signifying the amount of lookahead the parser is permitted to use.
#
# What should one do with the lookahead? That lookahead can be used to determine which rule to apply. In the case of an *LL(1)* parser, the rule to apply is determined by looking at the _first_ set of the different rules. We previously implemented `first_expr()` that takes a an expression, the set of `nullables`, and computes the first set of that rule.
#
# If a rule can derive an empty set, then that rule may also be applicable if of sees the `follow()` set of the corresponding nonterminal.
# + [markdown] slideshow={"slide_type": "skip"}
# #### Part 1: A LL(1) Parsing Table
#
# The first part of this exercise is to implement the _parse table_ that describes what action to take for an *LL(1)* parser on seeing a terminal symbol on lookahead. The table should be in the form of a _dictionary_ such that the keys represent the nonterminal symbol, and the value should contain another dictionary with keys as terminal symbols and the particular rule to continue parsing as the value.
#
# Let us illustrate this table with an example. The `parse_table()` method populates a `self.table` data structure that should conform to the following requirements:
# + slideshow={"slide_type": "skip"}
class LL1Parser(Parser):
def parse_table(self):
self.my_rules = rules(self.cgrammar)
self.table = ... # fill in here to produce
def rules(self):
for i, rule in enumerate(self.my_rules):
print(i, rule)
def show_table(self):
ts = list(sorted(terminals(self.cgrammar)))
print('Rule Name\t| %s' % ' | '.join(t for t in ts))
for k in self.table:
pr = self.table[k]
actions = list(str(pr[t]) if t in pr else ' ' for t in ts)
print('%s \t| %s' % (k, ' | '.join(actions)))
# + [markdown] slideshow={"slide_type": "skip"}
# On invocation of `LL1Parser(A2_GRAMMAR).show_table()`
# It should result in the following table:
# + slideshow={"slide_type": "skip"}
for i, r in enumerate(rules(canonical(A2_GRAMMAR))):
print("%d\t %s := %s" % (i, r[0], r[1]))
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# |Rule Name || + | - | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9|
# |-----------||---|---|---|---|---|---|---|---|---|---|---|--|
# |start || | | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0|
# |expr || | | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1|
# |expr_ || 2 | 3 | | | | | | | | | | |
# |integer || | | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5|
# |integer_ || 7 | 7 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6|
# |digit || | | 8 | 9 |10 |11 |12 |13 |14 |15 |16 |17|
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# **Solution.** We define `predict()` as we explained before. Then we use the predicted rules to populate the parse table.
# + slideshow={"slide_type": "subslide"} solution2="hidden"
class LL1Parser(LL1Parser):
def predict(self, rulepair, first, follow, epsilon):
A, rule = rulepair
rf = first_expr(rule, first, epsilon)
if nullable_expr(rule, epsilon):
rf |= follow[A]
return rf
def parse_table(self):
self.my_rules = rules(self.cgrammar)
epsilon = nullable(self.cgrammar)
first = firstset(self.cgrammar, epsilon)
# inefficient, can combine the three.
follow = followset(self.cgrammar, self.start_symbol)
ptable = [(i, self.predict(rule, first, follow, epsilon))
for i, rule in enumerate(self.my_rules)]
parse_tbl = {k: {} for k in self.cgrammar}
for i, pvals in ptable:
(k, expr) = self.my_rules[i]
parse_tbl[k].update({v: i for v in pvals})
self.table = parse_tbl
# + slideshow={"slide_type": "subslide"} solution2="hidden"
ll1parser = LL1Parser(A2_GRAMMAR)
ll1parser.parse_table()
ll1parser.show_table()
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# #### Part 2: The Parser
#
# Once we have the parse table, implementing the parser is as follows: Consider the first item from the sequence of tokens to parse, and seed the stack with the start symbol.
#
# While the stack is not empty, extract the first symbol from the stack, and if the symbol is a terminal, verify that the symbol matches the item from the input stream. If the symbol is a nonterminal, use the symbol and input item to lookup the next rule from the parse table. Insert the rule thus found to the top of the stack. Keep track of the expressions being parsed to build up the parse table.
#
# Use the parse table defined previously to implement the complete LL(1) parser.
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# **Solution.** Here is the complete parser:
# + slideshow={"slide_type": "subslide"} solution2="hidden"
class LL1Parser(LL1Parser):
def parse_helper(self, stack, inplst):
inp, *inplst = inplst
exprs = []
while stack:
val, *stack = stack
if isinstance(val, tuple):
exprs.append(val)
elif val not in self.cgrammar: # terminal
assert val == inp
exprs.append(val)
inp, *inplst = inplst or [None]
else:
if inp is not None:
i = self.table[val][inp]
_, rhs = self.my_rules[i]
stack = rhs + [(val, len(rhs))] + stack
return self.linear_to_tree(exprs)
def parse(self, inp):
self.parse_table()
k, _ = self.my_rules[0]
stack = [k]
return self.parse_helper(stack, inp)
def linear_to_tree(self, arr):
stack = []
while arr:
elt = arr.pop(0)
if not isinstance(elt, tuple):
stack.append((elt, []))
else:
# get the last n
sym, n = elt
elts = stack[-n:] if n > 0 else []
stack = stack[0:len(stack) - n]
stack.append((sym, elts))
assert len(stack) == 1
return stack[0]
# + slideshow={"slide_type": "subslide"} solution2="hidden"
ll1parser = LL1Parser(A2_GRAMMAR)
tree = ll1parser.parse('1+2')
display_tree(tree)
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# ### Exercise 9: A Different LangFuzzer
#
# Sometimes we do not want to use our pool of strings for various reasons – the number of items in the pool may be inadequate, or not varied enough. Extend the `LangFuzzer` to use a separate function to check if the number of items in the pool corresponding to the selected non-terminal is large enough (say greater than 10), and if not, use the tree expansion technique from `GrammarFuzzer` to patch the hole.
#
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# **Solution.** Here is a possible solution. Before we can make use of `GrammarFuzzer`, we need to change it a little bit. GrammarFuzzer relies on the grammar being in the `fuzzing` format with the expansions represented as strings. Our `LangFuzz` expects the expansions to be a list of tokens. So we fix the output of `GrammarFuzzer`.
# + slideshow={"slide_type": "subslide"} solution2="hidden"
class LangFuzzer2(LangFuzzer):
def __init__(self, parser, strings):
super().__init__(parser, strings)
self.gfuzz = GrammarFuzzer(parser.grammar())
def check_diversity(self, pool):
return len(pool) > 10
def candidate(self):
tree, nodes = random.choice(self.trees)
interesting_nodes = [
n for n in nodes if nodes[n][0] in self.fragments
and nodes[n][0] is not self.parser.start_symbol
and len(self.fragments[nodes[n][0]]) > 0
]
node = random.choice(interesting_nodes)
return tree, node
def generate_new_tree(self, node, choice):
name, children, id = node
if id == choice:
pool = self.fragments[name]
if self.check_diversity(pool):
return random.choice(self.fragments[name])
else:
return None
else:
return (name,
[self.generate_new_tree(c, choice) for c in children])
def fuzz(self):
tree, node = self.candidate()
tree_with_a_hole = self.generate_new_tree(tree, node)
modified = self.gfuzz.expand_tree(tree_with_a_hole)
return tree_to_string(modified)
# + slideshow={"slide_type": "subslide"} solution2="hidden"
parser = EarleyParser(VAR_GRAMMAR, tokens=VAR_TOKENS)
lf = LangFuzzer2(parser, mystrings)
for i in range(100):
print(lf.fuzz())
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# With these changes, our `LangFuzzer2` is able to use the pool of fragments when necessary, but can rely on the grammar when the pool is not sufficient.
| docs/beta/notebooks/Parser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.2 64-bit (''bilateral-gp'': conda)'
# name: python392jvsc74a57bd0ffdb161abba9b4c6ed90ea70f14a998c2b70ba8faeee50224e9918737cdc1ef1
# ---
# +
import torch
import numpy as np
import pandas as pd
import gpytorch as gp
import seaborn as sns
import matplotlib.pyplot as plt
from tqdm.auto import tqdm
device = 'cuda:0' if torch.cuda.is_available() else None
sns.set(font_scale=2.0, style='whitegrid')
# -
class ExactGPModel(gp.models.ExactGP):
def __init__(self, train_x, train_y):
likelihood = gp.likelihoods.GaussianLikelihood()
super().__init__(train_x, train_y, likelihood)
self.mean_module = gp.means.ZeroMean()
self.covar_module = gp.kernels.ScaleKernel(gp.kernels.RBFKernel())
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gp.distributions.MultivariateNormal(mean_x, covar_x)
# +
df = pd.read_csv('snelson.csv')
train_x = torch.from_numpy(df.x.values[:, np.newaxis]).float().to(device)[:10]
train_y = torch.from_numpy(df.y.values).float().to(device)[:10]
train_x = (train_x - train_x.mean(dim=0, keepdim=True)) / (train_x.std(dim=0, keepdim=True) + 1e-6)
train_y = (train_y - train_y.mean(dim=0, keepdim=True)) / (train_y.std(dim=0, keepdim=True) + 1e-6)
train_x.shape, train_y.shape
# -
model = ExactGPModel(train_x, train_y).to(device)
with torch.no_grad():
all_x = torch.linspace(-3., 3., 200).to(device).unsqueeze(-1)
prior = model.forward(all_x)
# +
def train(x, y, model, mll, optim):
model.train()
optim.zero_grad()
output = model(x)
loss = -mll(output, y)
loss.backward()
optim.step()
return { 'train/mll': -loss.detach().item() }
optim = torch.optim.Adam(model.parameters(), lr=.1)
mll = gp.mlls.ExactMarginalLogLikelihood(model.likelihood, model)
for i in tqdm(range(50)):
print(train(train_x, train_y, model, mll, optim))
model.eval()
with torch.no_grad():
posterior = model(all_x)
# +
def plot_fns(x, f):
y = torch.cat([x.expand(-1, 3).permute(1, 0).unsqueeze(-1), f.unsqueeze(-1)], axis=-1)
viz_data = []
for i in range(3):
for idx in range(200):
viz_data.append({ 'id': i, 'x': y[i][idx][0].item(), 'y': y[i][idx][1].item() })
viz_data = pd.DataFrame(viz_data)
fig, ax = plt.subplots(figsize=(11,7))
sns.lineplot(ax=ax, data=viz_data, x='x', y='y', hue='id', legend=False,
palette=sns.color_palette('Set1', 3), alpha=.7)
return fig, ax
fig_prior, ax_prior = plot_fns(all_x, prior.sample(torch.Size([3])))
ax_prior.plot(all_x.cpu().numpy().flatten(), prior.mean.cpu().numpy(), linestyle=(0, (10,5)),
color='black', alpha=.6, linewidth=3)
with torch.no_grad():
ax_prior.fill_between(all_x.cpu().numpy().flatten(),
prior.mean.cpu().numpy() - 2. * prior.variance.sqrt().cpu().numpy(),
prior.mean.cpu().numpy() + 2. * prior.variance.sqrt().cpu().numpy(),
color='grey', alpha=.15)
ax_prior.set_title('Prior')
ax_prior.set_yticks(np.arange(-2, 2.1))
ax_prior.set_ylim([-2.5,2.5])
fig_post, ax_post = plot_fns(all_x, posterior.sample(torch.Size([3])))
ax_post.plot(all_x.cpu().numpy().flatten(), posterior.mean.cpu().numpy(), linestyle=(0, (10,5)),
color='black', alpha=.6, linewidth=3)
with torch.no_grad():
ax_post.fill_between(all_x.cpu().numpy().flatten(),
posterior.mean.cpu().numpy() - 2. * posterior.variance.sqrt().cpu().numpy(),
posterior.mean.cpu().numpy() + 2. * posterior.variance.sqrt().cpu().numpy(),
color='grey', alpha=.15)
sns.scatterplot(ax=ax_post, x=train_x.squeeze(-1).cpu().numpy(), y=train_y.cpu().numpy(),
color='red', s=100, edgecolor='black', linewidth=1)
ax_post.set_title('Posterior')
ax_post.set_yticks(np.arange(-2, 2.1))
ax_post.set_ylim([-2.5,2.5]);
fig_prior.savefig('prior.pdf', bbox_inches='tight')
fig_post.savefig('post.pdf', bbox_inches='tight')
# -
| notebooks/viz_exact.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
# -
# ## load train data
pd_df_train = pd.read_csv("data/preprocessing_data/task1_preprocess_training_data.csv")
pd_df_train.head()
# ## Grouping and creating one hot encoding for intersection_id and time_window for train data
intersection_id_index = pd_df_train.groupby('intersection_id').ngroup()
intersection_id_class_vec = pd.get_dummies(intersection_id_index, prefix= "intesection_id_index")
time_window_index = pd_df_train.groupby('time_window').ngroup()
time_window_class_vec = pd.get_dummies(time_window_index)
pd_df_train = pd.concat([pd_df_train, intersection_id_index, intersection_id_class_vec,
time_window_index, time_window_class_vec], axis=1)
pd_df_train.head()
train_set_X = pd_df_train.drop(["intersection_id", "time_window"], axis=1)
train_set_X = train_set_X.rename(columns={"average_travl_time":"label"})
train_set_X.head()
train_set_X = train_set_X.loc[:,~train_set_X.columns.duplicated()]
train_set_y = train_set_X.label
pca = PCA(n_components=10)
features_pca=pca.fit_transform(train_set_X.drop("label", axis=1))
scaler=StandardScaler()
features_scled=scaler.fit_transform(features_pca)
features_scled.shape
# ## load test data
pd_df_test = pd.read_csv("data/preprocessing_data/task1_preprocess_test_data.csv")
pd_df_test.head()
# ## Grouping and creating one hot encoding for intersection_id and time_window for test data
intersection_id_index_test = pd_df_test.groupby('intersection_id').ngroup()
intersection_id_class_vec_test = pd.get_dummies(intersection_id_index_test, prefix= "intesection_id_index")
time_window_index_test = pd_df_test.groupby('time_window').ngroup()
time_window_class_vec_test = pd.get_dummies(time_window_index_test)
pd_df_test = pd.concat([pd_df_test, intersection_id_index_test, intersection_id_class_vec_test,
time_window_index_test, time_window_class_vec_test], axis=1)
pd_df_test.tail()
test_set = pd_df_test.drop(["intersection_id", "time_window"], axis=1)
pca = PCA(n_components=10)
test_features_pca=pca.fit_transform(test_set.drop("average_travl_time", axis=1))
scaler=StandardScaler()
test_features_scled=scaler.fit_transform(test_features_pca)
test_features_scled.shape
# ## Split data train set for train and cross validation
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(features_scled,
train_set_y,
test_size=0.30)
# ## Implementing Random Forest Regressor
from sklearn.ensemble import RandomForestRegressor
r_model = RandomForestRegressor(n_estimators=10)
r_model.fit(X_train, y_train)
y_pred = r_model.predict(X_test)
# ## Evaluation metrics
# +
from sklearn.metrics import mean_squared_error
from math import sqrt
# MAE,R^2:
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
# -
# ## MAPE Calculation method
def Mean_Absolute_Percentage_Error(labl,predction):
labl, predction = np.array(labl), np.array(predction)
return np.mean(np.abs((labl - predction) / labl))
# ## MAPE for Random Forest Regressor
print("MAPE for RF", Mean_Absolute_Percentage_Error(y_test, y_pred))
# ## Mean Abosute error for Random Forest Regressor
mae_rf=mean_absolute_error(y_test, y_pred)
print("MAE for RF", mae_rf)
# ## r2_score for Random Forest Regressor
r2_rf=r2_score(y_test,y_pred)
print("r2_score for RF", r2_rf)
# ## Root Mean Squared Error For Random Forest Regressor
# +
#root mean square error metric:
rms = np.sqrt(mean_squared_error(y_test, y_pred))
print("RMSE for RF", rms)
# +
#Not needed, unusually large value to compare the error performance metric.
mse = mean_squared_error(y_test, y_pred)
print("MSE for RF", mse)
# -
# ## Implementing Gradient Boosting Regressor
from sklearn.ensemble import GradientBoostingRegressor
gb_reg = GradientBoostingRegressor()
gb_reg.fit(X_train, y_train)
gb_pred = gb_reg.predict(X_test)
# ## Evaluation metrics
# +
# mean squared error
mse = mean_squared_error(y_test, gb_pred)
print("MSE for GBR", mse)
# +
# root mean squared error
rms_gbt=np.sqrt(mean_squared_error(y_test, gb_pred))
print("RMSE for GBR", rms_gbt)
# -
# ## MAPE for Gradient Boosting Regressor
print("MAPE for GBR", Mean_Absolute_Percentage_Error(y_test, gb_pred))
# ## MAE for Gradient Boosting Regressor
mae_gbr=mean_absolute_error(y_test, gb_pred)
print("MAE for GBR", mae_gbr)
# ## r_2 score for Gradient Boosting Regressor
r2_gb=r2_score(y_test,gb_pred)
print("r2_score for GBR", r2_gb)
# ## Implementing XGBOOST
# +
import xgboost as xgb
xgb_model = xgb.XGBRegressor()
xgb_model.fit(X_train, y_train)
y_pred_xgb = xgb_model.predict(X_test)
# -
# ## Evaluation of XGBoost performance
print("MAPE for XGB", Mean_Absolute_Percentage_Error(y_test, y_pred_xgb))
print("MAE for XBG", mean_absolute_error(y_test, y_pred_xgb))
print("RMSE for XGB", np.sqrt(mean_squared_error(y_test, y_pred_xgb)))
print("Mean Squared Error XGB", mean_squared_error(y_test, y_pred_xgb))
print("r_2 score XGB", r2_score(y_test,y_pred_xgb))
# ## Conclusion
# Based on the above experiment we choose Gradient Boosting Regressor as the best model as it has the lowest MAPE score
# ## Predicting on test data
pred = gb_reg.predict(test_features_scled)
pd_df_test["avg_travel_time"] = pred
f_data = pd_df_test[["intersection_id","tollgate_id","time_window","avg_travel_time"]]
f_data.head()
f_data.to_csv(path_or_buf="data/task1_submission.csv", index=False)
| task1_modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# a simple cnn model to separate noisy images from clean images
# +
import numpy as np
import pandas as pd
import os
import matplotlib.image as mpimg
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision
from torch.utils.data import DataLoader, Dataset
import torch.utils.data as utils
from torchvision import transforms
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
# -
# dataloader
import glob
import cv2
class ImageData(Dataset):
def __init__(self, pos_path_list, neg_path_list, transform): # pos = clean, neg = noisy
super().__init__()
self.imgs = []
self.labels = []
for p_path in pos_path_list:
for f in glob.glob(p_path):
self.imgs.append(f)
self.labels.append([0., 1.])
for n_path in neg_path_list:
for f in glob.glob(n_path):
self.imgs.append(f)
self.labels.append([1., 0.])
self.transform = transform
def __len__(self):
return len(self.imgs)
def __getitem__(self, index):
img_name = self.imgs[index]
label = self.labels[index]
image = mpimg.imread(img_path)
image = self.transform(image)
return image, label
data_transf = transforms.Compose([transforms.ToPILImage(), transforms.ToTensor()])
train_data = ImageData(["../dump/"] transform = data_transf)
train_loader = DataLoader(dataset = train_data, batch_size = 64)
from efficientnet_pytorch import EfficientNet
model = EfficientNet.from_pretrained('efficientnet-b0')
print(model)
model._fc =
| .ipynb_checkpoints/simple_cnn_noisy_clean_lp_classification-checkpoint.ipynb |
# <font size="10"><b><center>Heterogeneity and Macro Modeling</center></b></font>
# <font size="10"><b><center>In Policymaking Institutions</center></b></font>
# <br><br>
# <font size="6"><b><center><NAME></center></b></font>
# <font size="6"><b><center>Johns Hopkins University</center></b></font>
# <br><br>
# <font size="6"><b><center>"Heads of Research of the European Central Banks"</center></font>
# <font size="6"><b><center>London, June 24, 2019</center></b></font><br>
#
# # Bottom lines
#
# 1. Rep Agent ('RA') models are badly wrong
# * Inconsistent with basic (micro) theory _and_ evidence
# * $\Rightarrow$ least reliable when most urgently needed
# <!-- Not hard to match reality in Great Moderation: Nothing ever happened, many models can reproduce that! -->
# 1. New tools can solve these problems
# * Now: Hand-crafted, bespoke "works of art"
# * "Irreproducible": a compliment for artworks
# * For models, not so much
# <!-- spoke to someone from Very Important CB who said had spent weeks trying to understand zip file of code for one well known model and finally gave up -->
# * Need: "Open source" tools for HA models
# * Like DYNARE for RA
# * Creatable, maintainable by staff economists
#
# <!-- 1:30 -->
# # Teaser
#
# Bayer and Luetticke (2019): What Made Great Recession Great?
#
# Two models:
#
# * RANK: Sudden, massive, highly persistent neg pty shock
# * Why? Where did it come from?
# * HANK: Sudden, massive neg aggregate demand shock
# * Interpretation: Uncertainty _a la_ Nick Bloom
# * Dynamics are endogenous instead of assumed
# * Recovery is _autonomously_ slow <!-- Fed forecasts overpredicted recovery for 3 years running -->
# <font size="5"><b><center>Representative Agent New Keyensian ('RANK')</center></b></font>
#
# <center><img src="./BayerLuetticke/RANK_DS_Smoother_of_logY.png" width="600">
#
# <font size="5"><b><center>Heterogeneous Agent New Keyensian ('HANK')</center></b></font>
#
#
# <center><img src="./BayerLuetticke/HANK_DS_Smoother_of_logY.png" width="600">
# ### Diametrically opposite policy prescriptions:
#
# * RANK: Austerity
# * HANK: Stimulus
#
# IMF: "Stimulus worked; austerity didn't"
# ## Framing: It's All About the MPC ('$\kappa$')
#
# ### "Old Keynesian ('OK')" Cross ($\approx$ ZLB)
#
# \begin{eqnarray}
# \texttt{Period 0: } \hat{Y} & = & 1 \\
# \texttt{Period 1: } \phantom{\hat{Y}} & = & 1 + \kappa \\
# \texttt{Period 2: } \phantom{\hat{Y}} & = & 1 + \kappa + \kappa^2
# \end{eqnarray}
#
# <!--
# | $\phantom{.}$ | $\phantom{Periods:}$ | Date of Unit Shock <br> 0 | <br> 1 | <br> 2 | <br> ... |
# | :---: | :---: | :---: | --- | --- | --- |
# | $\Delta Y$ | = | 1 |
# | . | = | 1 | + $\kappa$ | |
# | . | = | 1 | + $\kappa$ | +$\kappa^{2}$ |
# | $\vdots$ |
# | . | = | $\frac{1}{(1-\kappa)}$ | |
# | . | Periods: | 0 (date of shock) | 1 | 2 | 3 | ... |
# | . | = | 1 | + $\kappa$ | +$\kappa^{2}$ | +$\kappa^{3}$ | -->
#
# Suppose "period" is a year and $\kappa = 0.75$
#
# 2-year "Multipliers"
# * $G = 1.75$
# * $T \approx 1.3 (= 0.75+0.75^2)$
# #### Anti-Old-Keynesians ('New Classicals'):
#
# $\kappa=0.75$ has "no microfoundations"
# * Perf Foresight unconstrained model $\Rightarrow \kappa \approx 0.04$
# <!-- More like 0.01 in models with habit formation -->
# #### "RA New Keynesian" (RANK) Models
#
# Fiscal Policy ('FP')?
#
# Doesn't work because $\kappa \approx 0.04$ and
# 1. Ricardian Equivalence for taxes
# 1. $G$ mostly changes _split_ of $Y$ between $C$, $I$, $G$
# #### "RA New Keynesian" (RANK) Models
#
# Monetary Policy ('MP')?
#
# Only works via Intertemporal Elasticity of Substitution (IES)
# * Problem: All micro and macro estimates are IES $\approx$ 0
# * $\Rightarrow$ MP doesn't work either
# * Kludge: Assume high IES despite evidence
# <!-- Excuse: It's macroeconomics so we can do this if we want
# * Blanchard, Eichenbaum
# * One of the earliest criticisms of RA models
# * No satisfactory excuse ever given -->
# * No "redistribution" channel (everyone identical)
# #### "Two Agent New Keynesian" (TANK) Models <!-- Will discuss role below -->
#
# * "Savers" like RA in RANK model
# * "Spenders" like in OK model
# ### Recent Update (Last 10 Years)
#
# #### Theory and Data Finally Agree
#
# ##### Theory (with uncertainty, liq constraints, illiquid assets, heterogeneity):
#
# * Easy to get $\bar{\kappa} = 0.5$ or more
# * Lots of heterogeneity in $\kappa$
#
# ##### Data (e.g., National registries)
# * Estimates are robustly $\bar{\kappa} = 0.5$ or more
# * Lots of heterogeneity in $\kappa$
# ### Auclert (2017)
#
# _In a model where everyone is optimizing_:
#
# FP that changes income:
# * $\bar{\kappa}$: is a 'sufficient statistic'
#
# MP:
# <!--* Channels: -->
# 1. IES channel still exists
# 1. _Also_ an "Old Keynesian"-like channel:
# * Effects of redist _between_ people with *different* $\kappa$'s
# ### Crawley (2019): Measure Auclert Stuff<!-- Kohn:"Monetary policy is like war -- conducted in a fog." But at least engineers know how the guns work! -->
#
# Relative size of MP channels?
# * Danish registry data for hetero in $\kappa$
# * Mechanism: $i \uparrow$
# * Reduces $Y$ for debtors ($\kappa = 0.75$)
# * Increases $Y$ for creditors ($\kappa = 0.25$)
# * $\kappa$ hetero channel is $\approx$ 5 times size of IES channel
# ### Crawley and Moon (2019)
# #### Does Auclert Decomposition Work in Theoretical Models?
#
# Can theory reproduce Crawley (2019) empirical findings?
#
# * Yes -- if you include heavily indebted consumers
# <font size="5"><b><center>Heterogeneous Agent New Keyensian ('HANK')</center></b></font>
#
#
# <center><img src="./Figures/KeynesianDebt_sigma3.png" width="600">
#
# #### Policy Transmission Channels
#
# Hetero $\kappa$:
# * crucial for _both_ MP _and_ FP
#
# If: we know how a policy:
# 1. Redistributes $Y$ betw groups w different $\kappa$'s
# 1. Changes constraints (which can have $\kappa >> 1$)
# Then: Micro evidence can _predict_:
# 1. How previously untried _macro_ policies might work
# 1. How effects of given policy should change over time
# * "Operation Twist" could have v diff effects:
# * US vs UK
# * 1960s vs 2020s
# #### TANK: Combines Defects of Both Progenitors
#
# 1. No credible microfoundations
# * For either class <!-- $\Rightarrow$ little confidence in predictions OoS -->
# * Can't use micro data to think about macro questions
# 1. Dynamics are wrong
# * Fails to produce observed sluggish dynamics
# * Half life of $C$ shocks is maybe a year
# 1. Says nothing about dynamic roles of:
# * Uncertainty
# * Constraints
# * Finance
#
# <!-- 25 mins to get here -->
# <!-- So, why hasn't everybody already adopted HANK models? Too hard! -->
# ## Challenge: HA Models Are Too Hard?
#
# ### That Can Change (Endogenously)
#
# * DSGE RA models once viewed as 'too hard'
# * Where there's a will there's a way: DYNARE
# <!-- Like me, senior policymakers in this room are probably old enough to
# remember when the same thing was said about RA RE macro models; we need to keep
# our Old Keynesian econometric models because RE models are too complex for
# government work -->
# ## Feasibility of 'DYNARE for HA models'?
#
# A New Insight is Very Good News: <!-- Kaplan et al, Krusell, Bayer and Luetticke, cstwMPC -->
# * Macro and Micro Largely Separable
# * Solve micro model for steady-state ('StSt') ONCE
# * Using powerful micro tools
# * Macro fluctuations?
# * Deviations from StSt
# * Not too hard to handle
#
# [Econ-ARK/HARK](https://github.com/econ-ark.org) open source toolkit is a start
# * So far mostly focused on micro problem
# * Have begun to integrate with macro
# ## Short Run
#
# * CB's should build 'Toy' Models Now
# * Norway has a head start
# * FRB has a number of people
# * Crawley starts there in Sep
# * Crucial:
# * Need to be able to swap models
# * Learn from each other
# * $\Rightarrow$ common (open source) toolkit
# * Do not use TROLL!
# ## Long Run
#
# #### 'Main model' key features:
# 1. Optimizing consumers subject to:
# 1. Constraints
# 1. Uncertainty
# 1. Tracks Income (and perceptions thereof), wealth distributions
# 1. 'seriously': resembles micro evidence
#
# #### 'Auxiliary models'
# 1. Fiscal policy (track distributional consequences)
# 1. MacroPru
# 1. Detailed monetary policy
# ## Digression on MacroPru
#
# The _whole_ point is to understand how many _micro_ consumers will stop paying <!-- Not, of course, that MacroPru authorities care about their financial distress -- they (rightly) are concerned about the distress they may inflict on fin -->
#
# Impossible to learn reliable answers using AR(1) models of aggregated data
#
# U.S. micro data: "double trigger"
#
# People don't default unless _both_:
# 1. Deeply underwater
# 1. Unemployment or other big neg income shock
#
# Conclusion?
#
# Optimal MacroPru rules:
# -- Depend on relative sizes of $Y$ vs $P$ shocks
#
# Micro modeling $\Rightarrow$ better macro policy
# ## Getting There
#
# * A long way to go yet
# * DYNARE wasn't built in a day
#
#
# * Institutional support of infrastructure development
# * Like DYNARE has had
# * As is done in other scientific/technical fields
# * [Astronomy, Artificial Intelligence, Bayesian Statistics, Biology, ...](https://www.scipy.org/topical-software.html)
#
# Feasible with modern collaborative software development tools:
# * Modular
# * Open-source
# * Platform-Independent
# * Automatic testing/debugging tools
# * Robust reproducibility
| Hetero-And-Macro/Hetero-And-Macro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="rqad6cNyIsR8" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593269814985, "user_tz": -120, "elapsed": 753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}}
import pandas as pd
import matplotlib.pyplot as plt
from collections import Counter
# + id="9qZkIlCdJNKX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593267745632, "user_tz": -120, "elapsed": 868, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="3bf4fa40-eddc-4918-bea6-88ee53d1de1d"
# cd 'drive/My Drive/Colab Notebooks/matrix/dw_matrix_three'
# + id="XWSgj0LYKG_E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593268109227, "user_tz": -120, "elapsed": 808, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="32940874-ee4b-4362-ca26-b89bd7a53779"
# cd data
# + id="61zuYa9SK7wa" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593268115155, "user_tz": -120, "elapsed": 4026, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}}
train = pd.read_pickle('train.p')
# + id="7fGivGJfLhaO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593268127038, "user_tz": -120, "elapsed": 949, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="15bc1d09-c2c6-4fb4-d2a5-0fcf9258104d"
train.keys()
# + id="HVC1oAkXLlbv" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593268227110, "user_tz": -120, "elapsed": 697, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}}
X_train = train['features']
y_train = train['labels']
# + id="Z39gVsQML89L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593268250268, "user_tz": -120, "elapsed": 951, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="e86acbda-bd6c-464c-f4a3-656643fc945d"
X_train.shape, y_train.shape
# + id="SFufucFIMTKT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} executionInfo={"status": "ok", "timestamp": 1593268376024, "user_tz": -120, "elapsed": 700, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="e28f9491-9365-4818-e1ac-98b83c8977af"
plt.imshow(X_train[10000])
# + id="G1xwbF4WM1-O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"status": "ok", "timestamp": 1593268600081, "user_tz": -120, "elapsed": 802, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="3ded9b65-d9fa-4fde-b851-39fe26aa9d37"
signs = pd.read_csv('signnames.csv')
signs.
# + id="Rz2KtfV4NfnK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1593268694832, "user_tz": -120, "elapsed": 714, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="4b78ffb1-1184-479f-baeb-fdbde5d54382"
dict_signs = signs.to_dict()['b']
dict_signs[30]
# + id="MitK4a0cN2WJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593268761455, "user_tz": -120, "elapsed": 941, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="6d635ac8-d502-453e-dfd2-a52f31445c44"
X_train[ y_train == 10].shape
# + id="j9vKHOC6OPpN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000, "output_embedded_package_id": "1-JUxvhaz6hwv2v4iAuTj731liRzo2mu4"} executionInfo={"status": "ok", "timestamp": 1593271456135, "user_tz": -120, "elapsed": 18927, "user": {"displayName": "J\u0<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="9e34fe49-d343-4d8c-96fe-7a0933769113"
for id_sign in dict_signs.keys():
given_signs = X_train[ y_train == id_sign ]
plt.figure(figsize = (15, 5))
for i in range(9):
plt.subplot('19{}'.format(i+1))
plt.imshow( given_signs[i] )
plt.axis('off')
plt.tight_layout()
plt.show()
# + id="kORyST_wSdbf" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593269970657, "user_tz": -120, "elapsed": 772, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}}
cnt = Counter(y_train).most_common()
# + id="wo6FgzlsSm4I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 578} executionInfo={"status": "ok", "timestamp": 1593270536347, "user_tz": -120, "elapsed": 1427, "user": {"displayName": "J\u01<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="60162e75-cff1-4d2f-afec-0dcb30f889b0"
id_labels, cnt_labels = zip(*cnt)
ids = range(len(id_labels))
plt.figure(figsize = (15, 5))
plt.bar(ids, cnt_labels)
plt.xlabel = 'Znaki'
labels = [dict_signs[ id_labels [id_ ] ] for id_ in id_labels]
plt.xticks(ids, labels, rotation = 'vertical')
plt.title('Znaki Drogowe - wystąpienia wg znaku')
plt.show()
# + id="TD2ImEaHVlxV" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593271121379, "user_tz": -120, "elapsed": 18422, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}}
# !git add data
# !git add dw_matrix_three/
# + id="v5P6Bn8MW7Uj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"status": "ok", "timestamp": 1593271311651, "user_tz": -120, "elapsed": 7062, "user": {"displayName": "J\u0119<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="0ca0c0a5-c0f8-4d8d-f19c-a78393fbb8bb"
# !git commit -m "add data and visualisation"
# + id="DxbPomikXlZ_" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1593271302971, "user_tz": -120, "elapsed": 6517, "user": {"displayName": "J\u01<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}}
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "Jędrzej"
# + id="DlYe-0DaXsfA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} executionInfo={"status": "ok", "timestamp": 1593271327028, "user_tz": -120, "elapsed": 12390, "user": {"displayName": "J\u0119<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gig_U8mFugPG7q5TrmLKDfkMjCZ8-cv3YvMUVSVxQ=s64", "userId": "16499995068208684705"}} outputId="400a0673-dad4-4277-c775-91445c26a3d6"
# !git push
| dw_matrix_three/day2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ChelseyGuasis/CPEN-21A-CPE-1-1/blob/main/Loop_Statement.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Jj_xx_NbrTF_"
# ##For Loop Statement
# + colab={"base_uri": "https://localhost:8080/"} id="xQSefsGArMf_" outputId="12d8feda-1b5e-42df-d501-990850d45fc3"
week = ["Sunday","Monday","Tuesday","Wednesday","Thursday","Friday","Saturday"]
for x in week:
print(x)
# + [markdown] id="2Bie16ootWJk"
# The break statement
# + colab={"base_uri": "https://localhost:8080/"} id="dKRcdWGys011" outputId="43a49000-c694-4fb8-ced3-5ffca3388629"
week = ["Sunday","Monday","Tuesday","Wednesday","Thursday","Friday","Saturday"]
for x in week:
print(x)
if x=="Thursday":
break
# + colab={"base_uri": "https://localhost:8080/"} id="8HMy8hP0t4yf" outputId="07b2e5a9-b4e1-4172-a401-85ad6ce18ff2"
week = ["Sunday","Monday","Tuesday","Wednesday","Thursday","Friday","Saturday"]
for x in week:
if x=="Thursday":
break
print(x)
# + [markdown] id="PNJQaO4fvEkT"
# Looping through a string
# + colab={"base_uri": "https://localhost:8080/"} id="hlrU6iFRvJIP" outputId="7add4583-0488-41e5-d07a-b117701ac2de"
for x in "week":
print(x)
# + [markdown] id="jww_-akhvqQl"
# The range() function
# + colab={"base_uri": "https://localhost:8080/"} id="Ai015pwMvth9" outputId="a0225270-9a4b-4c4f-87e1-abed0e0fdf05"
for x in range(6):
print("Example 1",x)
for x in range(2,6):
print("Example 2",x)
# + [markdown] id="YLscbeSBw9Bn"
# Nested Loops
# + colab={"base_uri": "https://localhost:8080/"} id="vMXJJrYOxXIL" outputId="a2d4dd96-091e-4fb0-a165-33e63ed402dc"
adjective = ["red","big","tasty"]
fruits = ["apple","banana","cherry"]
for x in adjective:
for y in fruits:
print(x,y)
# + [markdown] id="_qPYr5sIzLLQ"
# ##While loop
# + colab={"base_uri": "https://localhost:8080/"} id="1p34lQ6DzOwE" outputId="417daa71-b23f-4208-fc4a-62e8fd738a79"
i = 1
while i<6:
print(i)
i+=1 #Assignment operator for addition
# + [markdown] id="PJzTh6pw0doU"
# The break statement
# + colab={"base_uri": "https://localhost:8080/"} id="yxwdLXLF0HsT" outputId="d5a25fe5-b8ca-4c41-baee-8ab97aa80406"
i = 1
while i<6:
print(i)
if i==3:
break
i+=1 #Assignment operator for addition
# + colab={"base_uri": "https://localhost:8080/"} id="cU0ww6u02oyl" outputId="facdf12d-5613-4eac-e7a0-98bb50723429"
i= 1
while i<6:
i+=1 #Assignment operator for addition
if i==3:
continue
else:
print(i)
# + [markdown] id="QgNBschd3XsG"
# The else statement
# + colab={"base_uri": "https://localhost:8080/"} id="wYEiDY8P3Zxu" outputId="0750f11a-a5b7-4bcf-b17b-cf84378cc313"
i= 1
while i<6:
print(i)
i+=1
else:
print("i is no longer less than 6")
# + [markdown] id="v5rAZ-Eb33Z8"
# ###Application 1
# + colab={"base_uri": "https://localhost:8080/"} id="_txSAmd_5Agm" outputId="fa36da75-1b04-4b5f-f785-7bfc898fd976"
hello = ["Hello 0","Hello 1","Hello 2","Hello 3","Hello 4","Hello 5", "Hello 6","Hello 7","Hello 8","Hello 9","Hello 10"]
for x in hello:
print(x)
i=0
while i<=10:
print("Hello",i)
i+=1
# + colab={"base_uri": "https://localhost:8080/"} id="sqcVcMqK362c" outputId="fe10eccc-219e-4cc7-92f3-45e313143735"
for x in range(11):
print("Hello",x)
# + [markdown] id="1d6peJVm5j-O"
# ##Application 2
# + colab={"base_uri": "https://localhost:8080/"} id="q9qO2jov5pOJ" outputId="7723a0da-3a17-4c14-9731-6d67f7c83ed8"
for x in range(3,10):
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="ovh5cidn85pI" outputId="3cc8e878-819a-4d72-86b2-82c3d32687c4"
i=3
while i<10:
print(i)
if i==10:
break
i+=1
| Loop_Statement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from sklearn.base import BaseEstimator, RegressorMixin, clone
from sklearn.metrics.pairwise import rbf_kernel
from sklearn.utils.validation import check_X_y, check_array, check_is_fitted
import sys
sys.path.insert(0,'/Users/eman/Documents/code_projects/kernellib/')
import matplotlib.pyplot as plt
# %matplotlib inline
# + code_folding=[]
def sample_data():
"""Gets some sample data."""
d_dimensions = 1
n_samples = 20
noise_std = 0.1
seed = 123
rng = np.random.RandomState(seed)
n_train = 20
n_test = 5000
xtrain = np.linspace(-4, 5, n_train).reshape(n_train, 1)
xtest = np.linspace(-4, 5, n_test).reshape(n_test, 1)
f = lambda x: np.sin(x) * np.exp(0.2 * x)
ytrain = f(xtrain) + noise_std * rng.randn(n_train, 1)
ytest = f(xtest)
return xtrain, xtest, ytrain, ytest
# +
random_state = 0
rng = np.random.RandomState(random_state)
x_train, x_test, y_train, y_test = sample_data()
# # Training data is 11 points in [0,1] inclusive regularly spaced# Traini
# x_train = np.linspace(0, 1, 25).reshape(-1, 1)
# # True function is sin(2*pi*x) with Gaussian noise
# y_train = np.sin(x_train * (4 * np.pi)) + rng.randn(x_train.shape[0], 1) * 0.3
# y_train = np.squeeze(y_train)
# x_test = np.linspace(0, 1, 1000).reshape(-1, 1)
# +
fig, ax = plt.subplots()
ax.scatter(x_train, y_train)
plt.show()
# -
# ### RBF Kernel
from scipy.spatial.distance import pdist, cdist, squareform
from sklearn.metrics.pairwise import euclidean_distances
from sklearn.metrics.pairwise import rbf_kernel
# ## KRR with Cross Validation
from sklearn.base import BaseEstimator, RegressorMixin
from sklearn.metrics.pairwise import check_pairwise_arrays
from sklearn.linear_model.ridge import _solve_cholesky_kernel
from sklearn.utils import check_array, check_X_y
from sklearn.utils.validation import check_is_fitted
from sklearn.kernel_ridge import KernelRidge
from sklearn.model_selection import GridSearchCV
print(np.logspace(0, 10, 10))
# +
# %%time
krr_grid = GridSearchCV(KernelRidge(kernel='rbf'), cv=2, n_jobs=2,
param_grid={"alpha": [1e0, 0.1, 1e-2, 1e-3],
"gamma": np.logspace(-3, 3, 20)})
krr_grid.fit(x_train, y_train)
krr_model = krr_grid.best_estimator_
print(krr_model.gamma, krr_model.alpha)
weights = krr_model.dual_coef_
gamma = krr_model.gamma
alpha = krr_model.alpha
K = rbf_kernel(x_test, x_train, gamma=gamma)
y_pred = krr_model.predict(x_test)
# -
# # Full Derivative Map
# [Nx dx] = size(x);
# [Ns ds] = size(sv);
# Kreg = kernel2(x,sv,sigma);
# % for p=1:Nx
# % for q=1:Ns
# % for xxx = 1:dx
# % map0(xxx,p,q) = alpha(q) * (x(p,xxx)-sv(q,xxx)) * Kreg(p,q);
# % end
# % end
# % end
# map2 = zeros(dx,Nx,Ns);
# for p=1:Nx
# map2(:,p,:) = (repmat(alpha,1,size(sv,2)) .* (repmat(x(p,:),size(sv,1),1)-sv) .* repmat(Kreg(p,:)',1,size(sv,2)))';
# # %p
# end
# +
import sys
sys.path.insert(0, '/home/emmanuel/Drives/erc/code/kernellib')
from kernellib.derivatives import ard_derivative
# -
print(np.ndim(x_test))
# + [markdown] heading_collapsed=true
# ## Pure Python
# + hidden=true
def rbf_full_py(x_train, x_function, K, weights, gamma):
n_test, d_dims = x_function.shape
n_train, d_dims = x_train.shape
derivative = np.zeros(shape=(n_test, n_train, d_dims))
constant = - 2* gamma
for itest in range(n_test):
for itrain in range(n_train):
for idim in range(d_dims):
derivative[itest, itrain, idim] = \
weights[itrain] \
* (x_function[itest, idim] - x_train[itrain, idim]) \
* K[itest, itrain]
derivative *= constant
return derivative
# + hidden=true
d1_map_py = rbf_full_py(x_train, x_test, K, weights, gamma)
d1_der_py = d1_map_py.sum(axis=1)
print(d1_der_py.shape)
# + hidden=true
# pyed = %timeit -o rbf_full_py(x_train, x_test, K, weights, gamma)
# + hidden=true
fig, ax = plt.subplots()
ax.scatter(x_train, y_train, color='r', label='Training Data')
ax.plot(x_test, y_pred, color='k', label='Predictions')
ax.plot(x_test, d1_der_py, color='b', label='Derivatives')
ax.legend()
plt.show()
# + [markdown] heading_collapsed=true hidden=true
# #### Check Gradient
# + hidden=true
from scipy.optimize import check_grad, approx_fprime
# + code_folding=[] hidden=true
def my_grad(X):
full_d = rbf_full_py(x_train, X.reshape(-1,1), K, weights, gamma)
d1 = full_d.sum(axis=1).flatten()
return d1
def my_pred(X):
pred = krr_model.predict(X.reshape(-1, 1)).flatten()
return pred.flatten()
d1_numerical = list()
for i in x_test:
d1_numerical.append(approx_fprime(i, my_pred, [eps]))
d1_numerical = np.array(d1_numerical)
eps = np.sqrt(np.finfo(float).eps)
x = np.linspace(0, 1, 100)
errs = list()
for i in x_test:
errs.append(check_grad(my_pred, my_grad, i))
fig, ax = plt.subplots()
ax.plot(x_test, d1_numerical)
ax.plot(x_test, d1_der_py)
plt.show()
fig, ax = plt.subplots()
ax.plot(x_test, np.array(errs))
plt.show()
# -
# ## Numpy
# + code_folding=[8]
def rbf_full_numpy(x_train, x_function, K, weights, gamma):
n_test, d_dims = x_function.shape
n_train, d_dims = x_train.shape
derivative = np.zeros(shape=(n_test, n_train, d_dims))
constant = -2*gamma
for itest in range(n_test):
term1 = (np.tile(x_function[itest, :], (n_train, 1)) - x_train)
term2 = np.tile(weights, (1, d_dims))
term3 = np.tile(K[itest, :].T, (1, d_dims)).T
derivative[itest, :, :] = term1 * term2 * term3\
derivative *= constant
return derivative
# -
d1_map_np = rbf_full_numpy(x_train, x_test, K, weights, gamma)
# npyed = %timeit -o rbf_full_numpy(x_train, x_test, K, weights, gamma)
print('Numpy Speed up: ', pyed.best / npyed.best)
np.testing.assert_array_equal(d1_map_py, d1_map_np)
# ## Numba
import numba
from numba import njit, jit, prange, double
K = rbf_kernel(x_test, x_train, gamma=gamma)
weights = krr_model.dual_coef_
# +
import numba
@njit(fastmath=True)
def rbf_full_numba(x_train, x_function, K, weights, gamma):
n_test, d_dims = x_function.shape
n_train, d_dims = x_train.shape
derivative = np.zeros(shape=(n_test, n_train, d_dims))
for idim in prange(d_dims):
for itrain in range(n_train):
w = weights[itrain]
for itest in range(n_test):
# print(weights.shape)
derivative[itest, itrain, idim] = \
w * (x_function[itest, idim] - x_train[itrain, idim]) * K[itest, itrain]
derivative *= - 2 * gamma
return derivative
# -
d1_map_numba = rbf_full_numba(x_train, x_test, K, weights.flatten(), gamma)
# jitted = %timeit -o rbf_full_numba(x_train, x_test, K, weights.flatten(), gamma)
print('Jitted Speed up (Numpy): ', npyed.best / jitted.best)
print('Numba Speed up (Python): ', pyed.best / jitted.best)
np.testing.assert_array_equal(d1_map_py, d1_map_numba)
# # Derivative
# + [markdown] heading_collapsed=true
# ## Pure Python
# + hidden=true
def rbf_derivative_py(x_train, x_function, weights, gamma):
if np.ndim(x_function) == 1:
x_function = x_function[np.newaxis, :]
if np.ndim(weights) == 1:
weights = weights[:, np.newaxis]
n_test, d_dims = x_function.shape
n_train, d_dims = x_train.shape
derivative = np.zeros(shape=(n_test, n_train, d_dims))
K = rbf_kernel(x_function, x_train, gamma=gamma)
constant = -2*gamma
for itest in range(n_test):
term1 = (np.tile(x_function[itest, :], (n_train, 1)) - x_train)
term2 = np.tile(weights, (1, d_dims))
term3 = np.tile(K[itest, :].T, (1, d_dims)).T
derivative[itest, :, :] = term1 * term2 * term3\
derivative *= constant
derivative = np.sum(derivative, axis=1)
return derivative
# + hidden=true
d1_der_py = rbf_derivative_py(x_train, x_test, weights, gamma)
# + hidden=true
# pyedd = %timeit -o rbf_derivative_py(x_train, x_test, weights, gamma)
# -
# ## Numpy
# +
def rbf_derivative_np(x_train, x_function, K, weights, gamma):
# # check the sizes of x_train and x_test
# err_msg = "xtrain and xtest d dimensions are not equivalent."
# np.testing.assert_equal(x_function.shape[1], x_train.shape[1], err_msg=err_msg)
# # check the n_samples for x_train and weights are equal
# err_msg = "Number of training samples for xtrain and weights are not equal."
# np.testing.assert_equal(x_train.shape[0], weights.shape[0], err_msg=err_msg)
n_test, n_dims = x_function.shape
derivative = np.zeros(shape=x_function.shape)
for itest in range(n_test):
derivative[itest, :] = np.dot((x_function[itest, :] - x_train).T,
(K[itest, :].reshape(-1, 1) * weights))
derivative *= - 2 * gamma
return derivative
# +
# K = pairwise_kernels(x_function, x_train, gamma=gamma, metric='rbf')
K = rbf_kernel(x_test, x_train, gamma=gamma)
weights = krr_model.dual_coef_.reshape(-1, 1)
d1_der_np = rbf_derivative_np(x_train, x_test, K, weights, gamma)
# -
np.testing.assert_array_almost_equal(d1_der_py, d1_der_np, decimal=12)
# npyedd = %timeit -o rbf_derivative_np(x_train, x_train, K, weights, gamma)
print('Numpy Speed up: ', pyedd.best / npyedd.best)
# + [markdown] heading_collapsed=true
# #### Version II
# + hidden=true
# def rbf_derivative_np2(x_train, x_function, K, weights, gamma):
# # # check the sizes of x_train and x_test
# # err_msg = "xtrain and xtest d dimensions are not equivalent."
# # np.testing.assert_equal(x_function.shape[1], x_train.shape[1], err_msg=err_msg)
# # # check the n_samples for x_train and weights are equal
# # err_msg = "Number of training samples for xtrain and weights are not equal."
# # np.testing.assert_equal(x_train.shape[0], weights.shape[0], err_msg=err_msg)
# n_test, n_dims = x_function.shape
# derivative = np.zeros(shape=x_function.shape)
# constant = - 2 * gamma
# # print(x_train.T.shape)
# # print(np.diag(weights.flatten()).shape, K.T.shape)
# # print(weights.T.shape, K.T.shape)
# derivative = x_train.T.dot(np.diag(weights.flatten()).dot(K.T) - np.diag(weights.T.dot(K.T))).T
# derivative *= constant
# return derivative
# + hidden=true
d1_der_np2 = rbf_derivative_np2(x_train, x_train, K, weights, gamma)
# + hidden=true
# npyedd2 = %timeit -o rbf_derivative_np2(x_train, x_train, K, weights, gamma)
# + hidden=true
print('Numpy2 Speed up: ', pyedd.best / npyedd2.best)
# + hidden=true
np.testing.assert_almost_equal(d1_der_py, d1_der_np2, decimal=3)
# -
# ## Numba
# +
@njit(fastmath=True)
def rbf_derivative_numba(x_train, x_function, K, weights, gamma):
# # check the sizes of x_train and x_test
# err_msg = "xtrain and xtest d dimensions are not equivalent."
# np.testing.assert_equal(x_function.shape[1], x_train.shape[1], err_msg=err_msg)
# # check the n_samples for x_train and weights are equal
# err_msg = "Number of training samples for xtrain and weights are not equal."
# np.testing.assert_equal(x_train.shape[0], weights.shape[0], err_msg=err_msg)
n_test, n_dims = x_function.shape
derivative = np.zeros(shape=x_function.shape)
constant = - 2 * gamma
for itest in prange(n_test):
derivative[itest, :] = np.dot((x_function[itest, :] - x_train).T,
(K[itest, :].reshape(-1, 1) * weights))
derivative *= constant
return derivative
# -
d1_der_nmb = rbf_derivative_numba(x_train, x_test, K, weights, gamma)
np.testing.assert_array_almost_equal(d1_der_py, d1_der_nmb, decimal=10)
# jittedd = %timeit -o rbf_derivative_numba(x_train, x_train, K, weights, gamma)
print('Numpy Speed up: ', npyedd.best / jittedd.best)
print('Python Speed up: ', pyedd.best / jittedd.best)
# # 2nd Map Derivative
# $$\frac{\partial^2f}{x^2}=2\gamma \left[2 \gamma( x^j - y^j )^2 - 1 \right] K(x,y)$$
# ## Pure Python
def rbf_full2_py(x_train, x_function, K, weights, gamma):
n_test, d_dims = x_function.shape
n_train, d_dims = x_train.shape
derivative = np.zeros(shape=(n_test, n_train, d_dims))
constant = 2* gamma
for itest in range(n_test):
for itrain in range(n_train):
for idim in range(d_dims):
derivative[itest, itrain, idim] = \
weights[itrain, 0] \
* (constant * (x_function[itest, idim] - x_train[itrain, idim])**2 - 1) \
* K[itest, itrain]
derivative *= constant
return derivative
d2_map_py = rbf_full2_py(x_train, x_test, K, weights, gamma)
d2_der_py = d2_map_py.sum(axis=1)
print(d2_der_py.shape)
# pyed2 = %timeit -o rbf_full2_py(x_train, x_test, K, weights, gamma)
# +
fig, ax = plt.subplots()
ax.scatter(x_train, y_train, color='r', label='Training Data')
ax.plot(x_test, y_pred, color='k', label='Predictions')
ax.plot(x_test, d1_der_py, color='b', label='1st Derivative')
ax.plot(x_test, d2_der_py, color='g', label='2nd Derivative')
ax.legend()
plt.show()
# -
# ## Numpy
# +
def rbf_full2_numpy(x_train, x_function, K, weights, gamma):
n_test, d_dims = x_function.shape
n_train, d_dims = x_train.shape
derivative = np.zeros(shape=(n_test, n_train, d_dims))
constant = 2*gamma
term2 = np.tile(weights, (1, d_dims))
for itest in range(n_test):
term1 = constant * (np.tile(x_function[itest, :], (n_train, 1)) - x_train)**2 - 1
# print(term1.shape)
# print(term2.shape)
term3 = np.tile(K[itest, :].T, (1, d_dims)).T
# print(term3.shape)
derivative[itest, :, :] = term1 * term2 * term3
# break
derivative *= constant
return derivative
# -
d2_map_np = rbf_full2_numpy(x_train, x_test, K, weights, gamma)
# npyed2 = %timeit -o rbf_full2_numpy(x_train, x_test, K, weights, gamma)
print('Numpy Speed up: ', pyed2.best / npyed2.best)
np.testing.assert_array_almost_equal(d2_map_py, d2_map_np)
# ## Numba
@njit(fastmath=True)
def rbf_full2_numba(x_train, x_function, K, weights, gamma):
n_test, d_dims = x_function.shape
n_train, d_dims = x_train.shape
derivative = np.zeros(shape=(n_test, n_train, d_dims))
constant = 2* gamma
for idim in range(d_dims):
for itrain in range(n_train):
for itest in prange(n_test):
derivative[itest, itrain, idim] = \
weights[itrain]\
* (constant * (x_function[itest, idim] - x_train[itrain, idim])**2 - 1) \
* K[itest, itrain]
derivative *= constant
return derivative
d2_map_numba = rbf_full2_numba(x_train, x_test, K, weights.flatten(), gamma)
# jitted2 = %timeit -o rbf_full2_numba(x_train, x_test, K, weights.flatten(), gamma)
print('Python Speed up: ', pyed2.best / jitted2.best)
print('Numpy Speed up: ', npyed2.best / jitted2.best)
| notebooks/uncategorized/derivative/rbf_derivative_speedup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Handling NaN values
# When analysing tabular data, sometimes table cells are present that does not contain data. In Python this typically means the value is _Not a Number_ ([NaN](https://en.wikipedia.org/wiki/NaN)). We cannot assume these values are `0` or `-1` or any other value because that would distort descriptive statistics, for example. We need to deal with these NaN entries differently and this notebook will introduce how.
#
# To get a first view where NaNs play a role, we load again an example table and sort it.
import pandas as pd
data = pd.read_csv('../../data/Results.csv', index_col=0, delimiter=';')
data.sort_values(by = "Area", ascending=False)
# As you can see, there are rows at the bottom containing NaNs. These are at the bottom of the table because pandas cannot sort them.
# A quick check if there are NaNs anywhere in a DataFrame is an important quality check and good scientific practice.
data.isnull().values.any()
# We can also get some deeper insights in which columns these NaN values are located.
data.isnull().sum()
# For getting a glimpse about if we can further process that tabel, we may want to know the percentage of NaNs for each column?
data.isnull().mean().sort_values(ascending=False) *100
# # Dropping rows that containt NaNs
# Depending on what kind of data analysis should be performed, it might make sense to just ignore columns that contain NaN values. Alternatively, it is possible to delete rows that contain NaNs.
#
# It depends on your project and what is important or not for the analysis. Its not an easy answer.
data_no_nan = data.dropna(how="any")
data_no_nan
# On the bottom of that table, you can see that it still contains 374 of the original 391 columns. If you remove rows, you should document in your later scientific publication, home many out of how many datasets were analysed.
#
# We can now also check again if NaNs are present.
data_no_nan.isnull().values.any()
| docs/40_tabular_data_wrangling/handling_NaNs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Helaena-Cambel/OOP-58001/blob/main/Operations%20and%20Expressions%20in%20Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YXIzc2ydkjjg"
# **Boolean Operators**
#
# + id="nQ3FGy_okyO2" outputId="99bb1e58-4b05-49a5-b692-94c3cc5fefbf" colab={"base_uri": "https://localhost:8080/"}
print(10>9)
print(10==9)
print(10!=9)
print(10<9)
# + [markdown] id="Nuj9Il6XmHl2"
# **bool() Fnction**
# + id="3e_3ddeUmM17" outputId="f66bc104-4a99-4292-cb70-d8956c05a791" colab={"base_uri": "https://localhost:8080/"}
print(bool("Maria"))
print(bool(19))
print(bool([]))
print(bool(0))
print(bool(1))
print(bool(None))
print(bool(False))
# + [markdown] id="wQ73RYbHm6WJ"
# **Function can return a Boolean**
# + id="jmAf0ZSLm9Vt" outputId="a9acaf4b-9638-4642-90a5-9b442b96f7f7" colab={"base_uri": "https://localhost:8080/"}
def myFunction():
return False
print(myFunction())
# + id="b9JAC7uxn3kS" outputId="9b900cc1-50b6-4cea-8cb5-3e3b3740ef78" colab={"base_uri": "https://localhost:8080/"}
def myFunction():
return True
if myFunction():
print("Yes!")
else:
print("No!")
# + [markdown] id="Ve60svD4ozSJ"
# **Application 1**
# + id="reESfd-Qo1yj" outputId="d8173756-df37-462b-8823-d301aa7100e0" colab={"base_uri": "https://localhost:8080/"}
print(10>9)
a=6
b=7
print(a==b)
print(a!=a)
# + id="5ZZEIvvcqz3H" outputId="72f1d1e1-85e3-4c9e-9eb1-e91578aa8b34" colab={"base_uri": "https://localhost:8080/"}
a=60
b=13
print(a>b)
if a<b:
print("a is less than b")
else:
print("a is greater than b")
# + [markdown] id="zjaMMI3LrfMH"
# **Python Operators**
# + id="dXgqD_2frinl" outputId="f1ec40c2-3c8e-4385-8f39-2f6219d45291" colab={"base_uri": "https://localhost:8080/"}
print(10+3)
print(10-3)
print(10*3)
print(10%3)
print(10/3)
# + [markdown] id="ZLjczaXNr8S0"
# **Bitwise Operators**
# + id="apqnsufysAYc" outputId="9613a1b6-ea9d-496d-d695-a5eaa4a0151d" colab={"base_uri": "https://localhost:8080/"}
# a = 60, binary 0011 1100
# b = 13, binary 0000 1101
print(a&b)
print(a|b)
print(a^b)
print(a<<1)
print(a<<2)
# + [markdown] id="Xrxwhh5rvoHo"
# **Application 2**
# + id="MLT3AdLfvrdK" outputId="492154e1-1345-4f79-bafb-6642284b5821" colab={"base_uri": "https://localhost:8080/"}
#Assignment Operators
x=2
x+=3 #same as x=x+3
print(x)
x-=3 #same as x=x-3
print(x)
x*=3 #same as x=x*3
print(x)
x/=3 #same as x=x/3
print(x)
x%=3 #same as x=x%3
print(x)
# + [markdown] id="ckfe_q_5xNRj"
# **Logical Operators**
# + id="CuFjQIsoyLQ5" outputId="4c83633a-5ff6-43ec-f982-d40ffb2bd03d" colab={"base_uri": "https://localhost:8080/"}
k = True
l = False
print(k and l)
print(k or l)
print(not(k or l))
# + [markdown] id="0uemzPsmy6Mg"
# **Identity Operators**
# + id="9ttBpTNzy8dc" outputId="3994d92e-a1e8-4353-a07e-a0d0d56e4918" colab={"base_uri": "https://localhost:8080/"}
k is l
k is not l
# + [markdown] id="xEkf1iMuz0WH"
# #Control Structure
# + [markdown] id="_tpLUzldz3Ze"
# **If Statement**
# + id="JjdMjL3ez_Is" outputId="a0b1b6c2-8200-433c-d8be-e98d53e43903" colab={"base_uri": "https://localhost:8080/"}
v = 2
z = 1
if 1<2:
print("1 is less than 2")
# + [markdown] id="jPCs0Fej0N3G"
# **Elif Statement**
# + id="t8ITviOP0MeF" outputId="7b5e59e3-3de3-4721-8d0e-9c7b12f21ed9" colab={"base_uri": "https://localhost:8080/"}
if v<z:
print("v is less than z")
elif v>z:
print("v is greater than z")
# + [markdown] id="SqZRkY4z0uSc"
# **Else Statement**
# + id="Z5Slms1D06rt" outputId="97358aa7-a5d5-40f1-e63e-1c13ee3edd79" colab={"base_uri": "https://localhost:8080/"}
number = int(input()) #to know if the number is positive or negative
if number>0:
print("Positive")
elif number<0:
print("Negative")
else:
print("number is equal to zero")
# + [markdown] id="ay41uur-24L9"
# **Application 3**
# + id="Rha7W94k28nk" outputId="eddd346a-ce82-4b56-e6fc-4877c2654e43" colab={"base_uri": "https://localhost:8080/"}
#Develop a Python program that will accept if the person is entitled to vote or not
age = int(input())
if age>=18:
print("You are qualified to vote")
else:
print("You are not qualified to vote")
# + [markdown] id="5LLCnUzG3lAG"
# **Nested If...Else**
# + id="tA19Q6wp3nqO" outputId="0a69adaa-523a-423d-8134-df4a77439e8d" colab={"base_uri": "https://localhost:8080/"}
u = int(input())
if u>10:
print("u is above 10")
if u>20:
print("u is above 20")
if u>30:
print("u is above 30")
if u>40:
print("u is above 40")
else:
print("u is below 40")
if u>50:
print("u is above 50")
else:
print("u is below 50")
# + [markdown] id="brHpYgov7IzO"
# **Loop Structure**
# + id="jvYk6ojS7Ley" outputId="bc08b7ae-1dfa-4088-fce2-8904595bc61f" colab={"base_uri": "https://localhost:8080/"}
week = ['Sunday', 'Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday']
season = ["Rainy", "Sunny"]
for x in week:
for y in season:
print(y,x)
# + [markdown] id="YX3tgr0b8ywj"
# **Break Statement**
# + id="HhNdr7XI81yU" outputId="467168b9-7e4a-4b41-f84a-e114916bd626" colab={"base_uri": "https://localhost:8080/"}
for x in week:
print(x)
if x == "Thursday":
break
# + [markdown] id="8v6tm_3zAMy5"
# **While Loop**
# + id="t4_M9cJ0APJa" outputId="554fd1e0-9faa-4cce-e7ed-1f70881c268b" colab={"base_uri": "https://localhost:8080/"}
i=1
while i<6:
print(i)
i+=1
# + [markdown] id="v8yhsr7nAiW3"
# **Application 4**
#
#
# + id="tdVowPndAm-c" outputId="4e754f51-c2aa-49ed-dc66-981bf73d1038" colab={"base_uri": "https://localhost:8080/"}
#Create a Python program that displays numbers from 1 to 4 using while loop statement
n=1
while n<5:
print(n)
n+=1
# + [markdown] id="K-63xsqPBQTt"
# **Application 5**
# + id="O_EUaegoBLz-" outputId="972d7cd7-2b11-4751-a762-127d37b88923" colab={"base_uri": "https://localhost:8080/"}
#Create a Python program that displays 4 numbers using while loop and break statement
r=1
while r<=4:
if r==4:
print(r)
r+=1
| Operations and Expressions in Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyze Performance
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import json
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
# +
repetitions = 10
duration = 20000
start_time = 2000
end_time = start_time + duration
scale_factor = 1 / 100
# +
"""Load data"""
with open('./num-insets.json') as f:
num_insets = json.load(f)
num_insets = np.array(num_insets)
# Substract the start time
num_insets[:,2] -= num_insets[:,2][0]
fps_inner = []
for i in range(repetitions):
with open('inner-fps-{}.json'.format(i)) as f:
tmp = json.load(f)
fps_inner.append(np.array(tmp))
fps_outer = []
for i in range(repetitions):
with open('outer-fps-{}.json'.format(i)) as f:
tmp = json.load(f)
fps_outer.append(np.array(tmp))
# -
# ## Align profiles
#
# Since we have to [manually start the performance profiling in Chrome](https://stackoverflow.com/questions/56675684/programmatically-start-the-performance-profiling-in-chrome), we need to first align the frames before it makes sense to aggregate the results.
#
# We know that the animation runs for 20 seconds so we'll have to find the last frame. The initial frames have already been aligned in the data by removing all frames until and inclusive a frame with FPS ~1 or below. Such low FPS are related to a static view before the animation started.
# +
"""Get FPS by milliseconds"""
fps_inner_msec = np.zeros((repetitions, duration))
for r in range(repetitions):
last = 0
frames_duration = 1000 / fps_inner[r]
frames_duration = np.cumsum(frames_duration)
for i in range(frames_duration.size):
if frames_duration[i] < duration:
curr = np.int(frames_duration[i])
fps_inner_msec[r, last:curr] = fps_inner[r][i]
last = curr
else:
fps_inner_msec[r, last:] = fps_inner[r][i]
break
fps_outer_msec = np.zeros((repetitions, duration))
for r in range(repetitions):
last = 0
frames_duration = 1000 / fps_outer[r]
frames_duration = np.cumsum(frames_duration)
for i in range(frames_duration.size):
if frames_duration[i] < duration:
curr = np.int(frames_duration[i])
fps_outer_msec[r, last:curr] = fps_outer[r][i]
last = curr
else:
fps_outer_msec[r, last:] = fps_outer[r][i]
break
# +
"""Sanity check the FPS per repetition"""
from scipy import ndimage
for i in range(10):
plt.figure(figsize=(16,2))
plt.plot(ndimage.zoom(fps_msec[i], scale_factor))
plt.margins(0, 0.05)
plt.show()
# +
"""Get number of drawn insets and annotations per milliseconds"""
num_insets_msec = np.zeros(duration)
num_annos_msec = np.zeros(duration)
last = 0
for i in range(num_insets.shape[0]):
if num_insets[i, 2] > start_time:
if num_insets[i, 2] <= end_time:
num_insets_msec[last:num_insets[i, 2]] = num_insets[i, 0]
num_annos_msec[last:num_insets[i, 2]] = num_insets[i, 1]
last = num_insets[i, 2]
else:
num_insets_msec[last:] = num_insets[i, 0]
num_annos_msec[last:] = num_insets[i, 1]
break
# +
from scipy import stats
x = range(int(duration * scale_factor))
x_ticks = range(0, 220, 20)
x_tick_labels = range(0, 22, 2)
# FPS Inner
mean_fps_inner = ndimage.zoom(np.mean(fps_inner_msec, axis=0), scale_factor, mode='nearest')
se_fps_inner = ndimage.zoom(stats.sem(fps_inner_msec, axis=0), scale_factor, mode='nearest')
plt.figure(figsize=(16,2))
plt.axhline(y=60, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.axhline(y=40, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.axhline(y=20, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.plot(x, mean_fps_inner, color='#000000', linewidth=1)
plt.fill_between(x, mean_fps_inner - se_fps_inner, mean_fps_inner + se_fps_inner, color='#999999')
plt.axvspan(65, 68, facecolor='#FF2B00', alpha=0.5)
plt.axvspan(132.5, 135, facecolor='#FF2B00', alpha=0.5)
# To align the plots by pixels...
plt.yticks([10, 20, 40, 60], ['10', '20', '40', '060'])
plt.xticks(x_ticks, x_tick_labels)
plt.ylim(8, 70)
plt.margins(0)
plt.show()
# FPS Outer
mean_fps_outer = ndimage.zoom(np.mean(fps_outer_msec, axis=0), scale_factor, mode='nearest')
se_fps_outer = ndimage.zoom(stats.sem(fps_outer_msec, axis=0), scale_factor, mode='nearest')
plt.figure(figsize=(16,2))
plt.axhline(y=60, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.axhline(y=40, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.axhline(y=20, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.plot(x, mean_fps_outer, color='#000000', linewidth=1)
plt.fill_between(x, mean_fps_outer - se_fps_outer, mean_fps_outer + se_fps_outer, color='#999999')
plt.axvspan(65, 68, facecolor='#FF2B00', alpha=0.5)
plt.axvspan(132.5, 135, facecolor='#FF2B00', alpha=0.5)
# To align the plots by pixels...
plt.yticks([10, 20, 40, 60], ['10', '20', '40', '060'])
plt.xticks(x_ticks, x_tick_labels)
plt.ylim(8, 70)
plt.margins(0)
plt.show()
# Drawn insets
plt.figure(figsize=(16,1.5))
plt.axhline(y=40, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.axhline(y=30, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.axhline(y=20, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.plot(ndimage.zoom(num_insets_msec, scale_factor), color='#000000')
plt.margins(0, 0.05)
plt.axvspan(65, 68, facecolor='#FF2B00', alpha=0.5)
plt.axvspan(132.5, 135, facecolor='#FF2B00', alpha=0.5)
# To align the plots by pixels...
plt.yticks([10, 20, 30, 40], ['10', '20', '30', '040'])
plt.xticks(x_ticks, x_tick_labels)
plt.show()
# Drawn annotations
plt.figure(figsize=(16,1.5))
plt.axhline(y=750, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.axhline(y=500, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.axhline(y=250, color='#bbbbbb', linestyle='dashed', linewidth=1)
plt.plot(ndimage.zoom(num_annos_msec, scale_factor), color='#000000')
plt.margins(0, 0.05)
plt.axvspan(65, 68, facecolor='#FF2B00', alpha=0.5)
plt.axvspan(132.5, 135, facecolor='#FF2B00', alpha=0.5)
plt.xticks(x_ticks, x_tick_labels)
plt.show()
# +
"""For the Table 2 in the manuscript"""
print(np.round(np.mean(np.mean(fps_inner_msec, axis=0).reshape((10, -1)), axis=1)), np.round(np.mean(fps_inner_msec)))
print(np.round(np.mean(np.mean(fps_outer_msec, axis=0).reshape((10, -1)), axis=1)), np.round(np.mean(fps_outer_msec)))
print(np.round(np.mean(num_insets_msec.reshape((10, -1)), axis=1)))
print(np.round(np.mean(num_annos_msec.reshape((10, -1)), axis=1)))
# -
| experiment/analyze-performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/microprediction/timeseries-notebooks/blob/main/pycaret_ts_univariate.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="9wb8F5n1v8qT"
# !pip uninstall numpy
# !pip install sktime
# + id="qw0V5cxYkbXV"
# !pip install pycaret-ts-alpha
# !pip install statsmodels
# !pip install git+https://github.com/microprediction/timemachines.git
# + [markdown] id="gaQ8l9Pu1D7w"
#
#
#
#
# ## Using Pycaret for univariate prediction
# Predict five steps ahead using median of the best three models pycaret finds
#
#
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 870, "referenced_widgets": ["5d258a66744048ed8d93238bdd59b26a", "7dd211c587f54f7793b3de3e10e957ed", "a684b0478d1a4c1b8b0bcff77a8e500e", "d26400c0aadb43839a7817bf5faea18c", "2891f78dc5054300b1bab60f7808009d", "15dd6eae0e4f4d83868e617c2be3b46b"]} id="DfOacBJ_iuU-" outputId="3850da7d-6970-441b-b16f-f2b43d2b5432"
from timemachines.skaters.pycrt.pycaretwrapper import pycrt_iskater
import numpy as np
y = np.random.randn(500)
x,x_std = pycrt_iskater(y=y,k=5)
print(x)
| pycaret_ts_univariate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 机器学习纳米学位
# ## 非监督学习
# ## 项目 3: 创建用户分类
# 欢迎来到机器学习工程师纳米学位的第三个项目!在这个 notebook 文件中,有些模板代码已经提供给你,但你还需要实现更多的功能来完成这个项目。除非有明确要求,你无须修改任何已给出的代码。以**'练习'**开始的标题表示接下来的代码部分中有你必须要实现的功能。每一部分都会有详细的指导,需要实现的部分也会在注释中以 **'TODO'** 标出。请仔细阅读所有的提示!
#
# 除了实现代码外,你还**必须**回答一些与项目和你的实现有关的问题。每一个需要你回答的问题都会以**'问题 X'**为标题。请仔细阅读每个问题,并且在问题后的**'回答'**文字框中写出完整的答案。我们将根据你对问题的回答和撰写代码所实现的功能来对你提交的项目进行评分。
#
# >**提示:**Code 和 Markdown 区域可通过 **Shift + Enter** 快捷键运行。此外,Markdown 可以通过双击进入编辑模式。
# ## 开始
#
# 在这个项目中,你将分析一个数据集的内在结构,这个数据集包含很多客户真对不同类型产品的年度采购额(用**金额**表示)。这个项目的任务之一是如何最好地描述一个批发商不同种类顾客之间的差异。这样做将能够使得批发商能够更好的组织他们的物流服务以满足每个客户的需求。
#
# 这个项目的数据集能够在[UCI机器学习信息库](https://archive.ics.uci.edu/ml/datasets/Wholesale+customers)中找到.因为这个项目的目的,分析将不会包括 'Channel' 和 'Region' 这两个特征——重点集中在6个记录的客户购买的产品类别上。
#
# 运行下面的的代码单元以载入整个客户数据集和一些这个项目需要的 Python 库。如果你的数据集载入成功,你将看到后面输出数据集的大小。
# 检查你的Python版本
from sys import version_info
if version_info.major != 3:
raise Exception('请使用Python 3.x 来完成此项目')
# +
# 引入这个项目需要的库
import numpy as np
import pandas as pd
import visuals as vs
from IPython.display import display # 使得我们可以对DataFrame使用display()函数
# 设置以内联的形式显示matplotlib绘制的图片(在notebook中显示更美观)
# %matplotlib inline
# 高分辨率显示
# # %config InlineBackend.figure_format='retina'
# 载入整个客户数据集
try:
data = pd.read_csv("customers.csv")
data.drop(['Region', 'Channel'], axis = 1, inplace = True)
print("Wholesale customers dataset has {} samples with {} features each.".format(*data.shape))
except:
print("Dataset could not be loaded. Is the dataset missing?")
# -
# ## 分析数据
# 在这部分,你将开始分析数据,通过可视化和代码来理解每一个特征和其他特征的联系。你会看到关于数据集的统计描述,考虑每一个属性的相关性,然后从数据集中选择若干个样本数据点,你将在整个项目中一直跟踪研究这几个数据点。
#
# 运行下面的代码单元给出数据集的一个统计描述。注意这个数据集包含了6个重要的产品类型:**'Fresh'**, **'Milk'**, **'Grocery'**, **'Frozen'**, **'Detergents_Paper'**和 **'Delicatessen'**。想一下这里每一个类型代表你会购买什么样的产品。
# 显示数据集的一个描述
display(data.describe())
# ### 练习: 选择样本
# 为了对客户有一个更好的了解,并且了解代表他们的数据将会在这个分析过程中如何变换。最好是选择几个样本数据点,并且更为详细地分析它们。在下面的代码单元中,选择**三个**索引加入到索引列表`indices`中,这三个索引代表你要追踪的客户。我们建议你不断尝试,直到找到三个明显不同的客户。
# +
# TODO:从数据集中选择三个你希望抽样的数据点的索引
indices = []
# 为选择的样本建立一个DataFrame
samples = pd.DataFrame(data.loc[indices], columns = data.keys()).reset_index(drop = True)
print("Chosen samples of wholesale customers dataset:")
display(samples)
# -
# ### 问题 1
# 在你看来你选择的这三个样本点分别代表什么类型的企业(客户)?对每一个你选择的样本客户,通过它在每一种产品类型上的花费与数据集的统计描述进行比较,给出你做上述判断的理由。
#
#
# **提示:** 企业的类型包括 markets (市场)、cafes (咖啡馆、餐厅)、delis (熟食店、快餐店),wholesale retailers (大宗零售商/超级市场) 及其他。注意不要使用具体企业的名字,比如说在描述一个餐饮业客户时,你不能使用麦当劳。
# **回答:**
# ### 练习: 特征相关性
# 一个有趣的想法是,考虑这六个类别中的一个(或者多个)产品类别,是否对于理解客户的购买行为具有实际的相关性。也就是说,当用户购买了一定数量的某一类产品,我们是否能够确定他们必然会成比例地购买另一种类的产品。有一个简单的方法可以检测相关性:我们用移除了某一个特征之后的数据集来构建一个监督学习(回归)模型,然后用这个模型去预测那个被移除的特征,再对这个预测结果进行评分,看看预测结果如何。
#
# 在下面的代码单元中,你需要实现以下的功能:
# - 使用 `DataFrame.drop` 函数移除数据集中你选择的不需要的特征,并将移除后的结果赋值给 `new_data` 。
# - 使用 `sklearn.model_selection.train_test_split` 将数据集分割成训练集和测试集。
# - 使用移除的特征作为你的目标标签。设置 `test_size` 为 `0.25` 并设置一个 `random_state` 。
#
#
# - 导入一个 DecisionTreeRegressor (决策树回归器),设置一个 `random_state`,然后用训练集训练它。
# - 使用回归器的 `score` 函数输出模型在测试集上的预测得分。
# +
# TODO:为DataFrame创建一个副本,用'drop'函数丢弃一个特征# TODO:
new_data = None
# TODO:使用给定的特征作为目标,将数据分割成训练集和测试集
X_train, X_test, y_train, y_test = (None, None, None, None)
# TODO:创建一个DecisionTreeRegressor(决策树回归器)并在训练集上训练它
regressor = None
# TODO:输出在测试集上的预测得分
score = None
# -
# ### 问题 2
# 你尝试预测哪一个特征?预测的得分是多少?这个特征对于区分用户的消费习惯来说必要吗?为什么?
# **提示:** 决定系数(coefficient of determination),$R^2$ 结果在0到1之间,1表示完美拟合,一个负的 $R^2$ 表示模型不能够拟合数据。
# **回答:**
# ### 可视化特征分布
# 为了能够对这个数据集有一个更好的理解,我们可以对数据集中的每一个产品特征构建一个散布矩阵(scatter matrix)。如果你发现你在上面尝试预测的特征对于区分一个特定的用户来说是必须的,那么这个特征和其它的特征可能不会在下面的散射矩阵中显示任何关系。相反的,如果你认为这个特征对于识别一个特定的客户是没有作用的,那么通过散布矩阵可以看出在这个数据特征和其它特征中有关联性。运行下面的代码以创建一个散布矩阵。
# 对于数据中的每一对特征构造一个散布矩阵
pd.plotting.scatter_matrix(data, alpha = 0.3, figsize = (14,8), diagonal = 'kde');
# ### 问题 3
# 这里是否存在一些特征他们彼此之间存在一定程度相关性?如果有请列出。这个结果是验证了还是否认了你尝试预测的那个特征的相关性?这些特征的数据是怎么分布的?
#
# **提示:** 这些数据是正态分布(normally distributed)的吗?大多数的数据点分布在哪?
# **回答:**
# ## 数据预处理
# 在这个部分,你将通过在数据上做一个合适的缩放,并检测异常点(你可以选择性移除)将数据预处理成一个更好的代表客户的形式。预处理数据是保证你在分析中能够得到显著且有意义的结果的重要环节。
# ### 练习: 特征缩放
# 如果数据不是正态分布的,尤其是数据的平均数和中位数相差很大的时候(表示数据非常歪斜)。这时候通常用一个[非线性的缩放](https://github.com/czcbangkai/translations/blob/master/use_of_logarithms_in_economics/use_of_logarithms_in_economics.pdf)是很合适的,[(英文原文)](http://econbrowser.com/archives/2014/02/use-of-logarithms-in-economics) — 尤其是对于金融数据。一种实现这个缩放的方法是使用 [Box-Cox 变换](http://scipy.github.io/devdocs/generated/scipy.stats.boxcox.html),这个方法能够计算出能够最佳减小数据倾斜的指数变换方法。一个比较简单的并且在大多数情况下都适用的方法是使用自然对数。
#
# 在下面的代码单元中,你将需要实现以下功能:
# - 使用 `np.log` 函数在数据 `data` 上做一个对数缩放,然后将它的副本(不改变原始data的值)赋值给 `log_data`。
# - 使用 `np.log` 函数在样本数据 `samples` 上做一个对数缩放,然后将它的副本赋值给 `log_samples`。
# +
# TODO:使用自然对数缩放数据
log_data = None
# TODO:使用自然对数缩放样本数据
log_samples = None
# 为每一对新产生的特征制作一个散射矩阵
pd.plotting.scatter_matrix(log_data, alpha = 0.3, figsize = (14,8), diagonal = 'kde');
# -
# ### 观察
# 在使用了一个自然对数的缩放之后,数据的各个特征会显得更加的正态分布。对于任意的你以前发现有相关关系的特征对,观察他们的相关关系是否还是存在的(并且尝试观察,他们的相关关系相比原来是变强了还是变弱了)。
#
# 运行下面的代码以观察样本数据在进行了自然对数转换之后如何改变了。
# 展示经过对数变换后的样本数据
display(log_samples)
# ### 练习: 异常值检测
# 对于任何的分析,在数据预处理的过程中检测数据中的异常值都是非常重要的一步。异常值的出现会使得把这些值考虑进去后结果出现倾斜。这里有很多关于怎样定义什么是数据集中的异常值的经验法则。这里我们将使用[ Tukey 的定义异常值的方法](http://datapigtechnologies.com/blog/index.php/highlighting-outliers-in-your-data-with-the-tukey-method/):一个异常阶(outlier step)被定义成1.5倍的四分位距(interquartile range,IQR)。一个数据点如果某个特征包含在该特征的 IQR 之外的特征,那么该数据点被认定为异常点。
#
# 在下面的代码单元中,你需要完成下面的功能:
# - 将指定特征的 25th 分位点的值分配给 `Q1` 。使用 `np.percentile` 来完成这个功能。
# - 将指定特征的 75th 分位点的值分配给 `Q3` 。同样的,使用 `np.percentile` 来完成这个功能。
# - 将指定特征的异常阶的计算结果赋值给 `step`。
# - 选择性地通过将索引添加到 `outliers` 列表中,以移除异常值。
#
# **注意:** 如果你选择移除异常值,请保证你选择的样本点不在这些移除的点当中!
# 一旦你完成了这些功能,数据集将存储在 `good_data` 中。
# +
# 对于每一个特征,找到值异常高或者是异常低的数据点
for feature in log_data.keys():
# TODO: 计算给定特征的Q1(数据的25th分位点)
Q1 = None
# TODO: 计算给定特征的Q3(数据的75th分位点)
Q3 = None
# TODO: 使用四分位范围计算异常阶(1.5倍的四分位距)
step = None
# 显示异常点
print("Data points considered outliers for the feature '{}':".format(feature))
display(log_data[~((log_data[feature] >= Q1 - step) & (log_data[feature] <= Q3 + step))])
# TODO(可选): 选择你希望移除的数据点的索引
outliers = []
# 以下代码会移除outliers中索引的数据点, 并储存在good_data中
good_data = log_data.drop(log_data.index[outliers]).reset_index(drop = True)
# -
# ### 问题 4
# 请列出所有在多于一个特征下被看作是异常的数据点。这些点应该被从数据集中移除吗?为什么?把你认为需要移除的数据点全部加入到到 `outliers` 变量中。
# **回答:**
# ## 特征转换
# 在这个部分中你将使用主成分分析(PCA)来分析批发商客户数据的内在结构。由于使用PCA在一个数据集上会计算出最大化方差的维度,我们将找出哪一个特征组合能够最好的描绘客户。
# ### 练习: 主成分分析(PCA)
#
# 既然数据被缩放到一个更加正态分布的范围中并且我们也移除了需要移除的异常点,我们现在就能够在 `good_data` 上使用PCA算法以发现数据的哪一个维度能够最大化特征的方差。除了找到这些维度,PCA 也将报告每一个维度的解释方差比(explained variance ratio)--这个数据有多少方差能够用这个单独的维度来解释。注意 PCA 的一个组成部分(维度)能够被看做这个空间中的一个新的“特征”,但是它是原来数据中的特征构成的。
#
# 在下面的代码单元中,你将要实现下面的功能:
# - 导入 `sklearn.decomposition.PCA` 并且将 `good_data` 用 PCA 并且使用6个维度进行拟合后的结果保存到 `pca` 中。
# - 使用 `pca.transform` 将 `log_samples` 进行转换,并将结果存储到 `pca_samples` 中。
# +
# TODO:通过在good data上进行PCA,将其转换成6个维度
pca = None
# TODO:使用上面的PCA拟合将变换施加在log_samples上
pca_samples = None
# 生成PCA的结果图
pca_results = vs.pca_results(good_data, pca)
# -
# ### 问题 5
# 数据的第一个和第二个主成分**总共**表示了多少的方差? 前四个主成分呢?使用上面提供的可视化图像,从用户花费的角度来讨论前四个主要成分中每个主成分代表的消费行为并给出你做出判断的理由。
#
# **提示:**
# * 对每个主成分中的特征分析权重的正负和大小。
# * 结合每个主成分权重的正负讨论消费行为。
# * 某一特定维度上的正向增长对应正权特征的增长和负权特征的减少。增长和减少的速率和每个特征的权重相关。[参考资料:Interpretation of the Principal Components](https://onlinecourses.science.psu.edu/stat505/node/54)
# **回答:**
# ### 观察
# 运行下面的代码,查看经过对数转换的样本数据在进行一个6个维度的主成分分析(PCA)之后会如何改变。观察样本数据的前四个维度的数值。考虑这和你初始对样本点的解释是否一致。
# 展示经过PCA转换的sample log-data
display(pd.DataFrame(np.round(pca_samples, 4), columns = pca_results.index.values))
# ### 练习:降维
# 当使用主成分分析的时候,一个主要的目的是减少数据的维度,这实际上降低了问题的复杂度。当然降维也是需要一定代价的:更少的维度能够表示的数据中的总方差更少。因为这个,**累计解释方差比(cumulative explained variance ratio)**对于我们确定这个问题需要多少维度非常重要。另外,如果大部分的方差都能够通过两个或者是三个维度进行表示的话,降维之后的数据能够被可视化。
#
# 在下面的代码单元中,你将实现下面的功能:
# - 将 `good_data` 用两个维度的PCA进行拟合,并将结果存储到 `pca` 中去。
# - 使用 `pca.transform` 将 `good_data` 进行转换,并将结果存储在 `reduced_data` 中。
# - 使用 `pca.transform` 将 `log_samples` 进行转换,并将结果存储在 `pca_samples` 中。
# +
# TODO:通过在good data上进行PCA,将其转换成两个维度
pca = None
# TODO:使用上面训练的PCA将good data进行转换
reduced_data = None
# TODO:使用上面训练的PCA将log_samples进行转换
pca_samples = None
# 为降维后的数据创建一个DataFrame
reduced_data = pd.DataFrame(reduced_data, columns = ['Dimension 1', 'Dimension 2'])
# -
# ### 观察
# 运行以下代码观察当仅仅使用两个维度进行 PCA 转换后,这个对数样本数据将怎样变化。观察这里的结果与一个使用六个维度的 PCA 转换相比较时,前两维的数值是保持不变的。
# 展示经过两个维度的PCA转换之后的样本log-data
display(pd.DataFrame(np.round(pca_samples, 4), columns = ['Dimension 1', 'Dimension 2']))
# ## 可视化一个双标图(Biplot)
# 双标图是一个散点图,每个数据点的位置由它所在主成分的分数确定。坐标系是主成分(这里是 `Dimension 1` 和 `Dimension 2`)。此外,双标图还展示出初始特征在主成分上的投影。一个双标图可以帮助我们理解降维后的数据,发现主成分和初始特征之间的关系。
#
# 运行下面的代码来创建一个降维后数据的双标图。
# 可视化双标图
vs.biplot(good_data, reduced_data, pca)
# ### 观察
#
# 一旦我们有了原始特征的投影(红色箭头),就能更加容易的理解散点图每个数据点的相对位置。
#
# 在这个双标图中,哪些初始特征与第一个主成分有强关联?哪些初始特征与第二个主成分相关联?你观察到的是否与之前得到的 pca_results 图相符?
# ## 聚类
#
# 在这个部分,你讲选择使用 K-Means 聚类算法或者是高斯混合模型聚类算法以发现数据中隐藏的客户分类。然后,你将从簇中恢复一些特定的关键数据点,通过将它们转换回原始的维度和规模,从而理解他们的含义。
# ### 问题 6
# 使用 K-Means 聚类算法的优点是什么?使用高斯混合模型聚类算法的优点是什么?基于你现在对客户数据的观察结果,你选用了这两个算法中的哪一个,为什么?
# **回答:**
# ### 练习: 创建聚类
#
# 针对不同情况,有些问题你需要的聚类数目可能是已知的。但是在聚类数目不作为一个**先验**知道的情况下,我们并不能够保证某个聚类的数目对这个数据是最优的,因为我们对于数据的结构(如果存在的话)是不清楚的。但是,我们可以通过计算每一个簇中点的**轮廓系数**来衡量聚类的质量。数据点的[轮廓系数](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.silhouette_score.html)衡量了它与分配给他的簇的相似度,这个值范围在-1(不相似)到1(相似)。**平均**轮廓系数为我们提供了一种简单地度量聚类质量的方法。
#
# 在接下来的代码单元中,你将实现下列功能:
# - 在 `reduced_data` 上使用一个聚类算法,并将结果赋值到 `clusterer`,需要设置 `random_state` 使得结果可以复现。
# - 使用 `clusterer.predict` 预测 `reduced_data` 中的每一个点的簇,并将结果赋值到 `preds`。
# - 使用算法的某个属性值找到聚类中心,并将它们赋值到 `centers`。
# - 预测 `pca_samples` 中的每一个样本点的类别并将结果赋值到 `sample_preds`。
# - 导入 `sklearn.metrics.silhouette_score` 包并计算 `reduced_data` 相对于 `preds` 的轮廓系数。
# - 将轮廓系数赋值给 `score` 并输出结果。
# +
# TODO:在降维后的数据上使用你选择的聚类算法
clusterer = None
# TODO:预测每一个点的簇
preds = None
# TODO:找到聚类中心
centers = None
# TODO:预测在每一个转换后的样本点的类
sample_preds = None
# TODO:计算选择的类别的平均轮廓系数(mean silhouette coefficient)
score = None
# -
# ### 问题 7
#
# 汇报你尝试的不同的聚类数对应的轮廓系数。在这些当中哪一个聚类的数目能够得到最佳的轮廓系数?
# **回答:**
# ### 聚类可视化
# 一旦你选好了通过上面的评价函数得到的算法的最佳聚类数目,你就能够通过使用下面的代码块可视化来得到的结果。作为实验,你可以试着调整你的聚类算法的聚类的数量来看一下不同的可视化结果。但是你提供的最终的可视化图像必须和你选择的最优聚类数目一致。
# 从已有的实现中展示聚类的结果
vs.cluster_results(reduced_data, preds, centers, pca_samples)
# ### 练习: 数据恢复
# 上面的可视化图像中提供的每一个聚类都有一个中心点。这些中心(或者叫平均点)并不是数据中真实存在的点,但是是所有预测在这个簇中的数据点的平均。对于创建客户分类的问题,一个簇的中心对应于那个分类的平均用户。因为这个数据现在进行了降维并缩放到一定的范围,我们可以通过施加一个反向的转换恢复这个点所代表的用户的花费。
#
# 在下面的代码单元中,你将实现下列的功能:
# - 使用 `pca.inverse_transform` 将 `centers` 反向转换,并将结果存储在 `log_centers` 中。
# - 使用 `np.log` 的反函数 `np.exp` 反向转换 `log_centers` 并将结果存储到 `true_centers` 中。
#
# +
# TODO:反向转换中心点
log_centers = None
# TODO:对中心点做指数转换
true_centers = None
# 显示真实的中心点
segments = ['Segment {}'.format(i) for i in range(0,len(centers))]
true_centers = pd.DataFrame(np.round(true_centers), columns = data.keys())
true_centers.index = segments
display(true_centers)
# -
# ### 问题 8
# 考虑上面的代表性数据点在每一个产品类型的花费总数,你认为这些客户分类代表了哪类客户?为什么?需要参考在项目最开始得到的统计值来给出理由。
#
# **提示:** 一个被分到`'Cluster X'`的客户最好被用 `'Segment X'`中的特征集来标识的企业类型表示。
# **回答:**
# ### 问题 9
# 对于每一个样本点**问题 8 **中的哪一个分类能够最好的表示它?你之前对样本的预测和现在的结果相符吗?
#
# 运行下面的代码单元以找到每一个样本点被预测到哪一个簇中去。
# 显示预测结果
for i, pred in enumerate(sample_preds):
print("Sample point", i, "predicted to be in Cluster", pred)
# **回答:**
# ## 结论
#
# 在最后一部分中,你要学习如何使用已经被分类的数据。首先,你要考虑不同组的客户**客户分类**,针对不同的派送策略受到的影响会有什么不同。其次,你要考虑到,每一个客户都被打上了标签(客户属于哪一个分类)可以给客户数据提供一个多一个特征。最后,你会把客户分类与一个数据中的隐藏变量做比较,看一下这个分类是否辨识了特定的关系。
# ### 问题 10
# 在对他们的服务或者是产品做细微的改变的时候,公司经常会使用 [A/B tests ](https://en.wikipedia.org/wiki/A/B_testing)以确定这些改变会对客户产生积极作用还是消极作用。这个批发商希望考虑将他的派送服务从每周5天变为每周3天,但是他只会对他客户当中对此有积极反馈的客户采用。这个批发商应该如何利用客户分类来知道哪些客户对它的这个派送策略的改变有积极的反馈,如果有的话?你需要给出在这个情形下A/B 测试具体的实现方法,以及最终得出结论的依据是什么?
#
# **提示:** 我们能假设这个改变对所有的客户影响都一致吗?我们怎样才能够确定它对于哪个类型的客户影响最大?
# **回答:**
# ### 问题 11
# 通过聚类技术,我们能够将原有的没有标记的数据集中的附加结构分析出来。因为每一个客户都有一个最佳的划分(取决于你选择使用的聚类算法),我们可以把用户分类作为数据的一个[工程特征](https://en.wikipedia.org/wiki/Feature_learning#Unsupervised_feature_learning)。假设批发商最近迎来十位新顾客,并且他已经为每位顾客每个产品类别年度采购额进行了预估。进行了这些估算之后,批发商该如何运用它的预估和非监督学习的结果来对这十个新的客户进行更好的预测?
#
# **提示**:在下面的代码单元中,我们提供了一个已经做好聚类的数据(聚类结果为数据中的cluster属性),我们将在这个数据集上做一个小实验。尝试运行下面的代码看看我们尝试预测‘Region’的时候,如果存在聚类特征'cluster'与不存在相比对最终的得分会有什么影响?这对你有什么启发?
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
# 读取包含聚类结果的数据
cluster_data = pd.read_csv("cluster.csv")
y = cluster_data['Region']
X = cluster_data.drop(['Region'], axis = 1)
# 划分训练集测试集
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=24)
clf = RandomForestClassifier(random_state=24)
clf.fit(X_train, y_train)
score_with_cluster = clf.score(X_test, y_test)
# 移除cluster特征
X_train = X_train.copy()
X_train.drop(['cluster'], axis=1, inplace=True)
X_test = X_test.copy()
X_test.drop(['cluster'], axis=1, inplace=True)
clf.fit(X_train, y_train)
score_no_cluster = clf.score(X_test, y_test)
print("不使用cluster特征的得分: %.4f"%score_no_cluster)
print("使用cluster特征的得分: %.4f"%score_with_cluster)
# -
# **回答:**
# ### 可视化内在的分布
#
# 在这个项目的开始,我们讨论了从数据集中移除 `'Channel'` 和 `'Region'` 特征,这样在分析过程中我们就会着重分析用户产品类别。通过重新引入 `Channel` 这个特征到数据集中,并施加和原来数据集同样的 PCA 变换的时候我们将能够发现数据集产生一个有趣的结构。
#
# 运行下面的代码单元以查看哪一个数据点在降维的空间中被标记为 `'HoReCa'` (旅馆/餐馆/咖啡厅)或者 `'Retail'`。另外,你将发现样本点在图中被圈了出来,用以显示他们的标签。
# 根据‘Channel‘数据显示聚类的结果
vs.channel_results(reduced_data, outliers, pca_samples)
# ### 问题 12
#
# 你选择的聚类算法和聚类点的数目,与内在的旅馆/餐馆/咖啡店和零售商的分布相比,有足够好吗?根据这个分布有没有哪个簇能够刚好划分成'零售商'或者是'旅馆/饭店/咖啡馆'?你觉得这个分类和前面你对于用户分类的定义是一致的吗?
# **回答:**
# > **注意**: 当你写完了所有的代码,并且回答了所有的问题。你就可以把你的 iPython Notebook 导出成 HTML 文件。你可以在菜单栏,这样导出**File -> Download as -> HTML (.html)**把这个 HTML 和这个 iPython notebook 一起做为你的作业提交。
| Project4_Unsupervised_learning/project/customer_segments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os
from copy import deepcopy
from pommerman.agents import SimpleAgent
from pommerman.configs import ffa_competition_env
from pommerman.constants import BOARD_SIZE
from pommerman.envs.v0 import Pomme
from tqdm import tqdm_notebook as tqdm
from matplotlib import pyplot as plt
# %matplotlib inline
# +
initial_rollouts = 600
min_episode_length = 35
train_data_path = './dataset/'
train_data_obs = os.path.join(train_data_path, 'obs.npy')
train_data_labels = os.path.join(train_data_path, 'labels.npy')
train_data_reward = os.path.join(train_data_path, 'reward.npy')
train_data_obs_map = os.path.join(train_data_path, 'obs_map.npy')
if not os.path.isdir(train_data_path):
os.makedirs(train_data_path)
# -
class Episode:
def __init__(self, agent_id, episode_id):
self.agent_id = agent_id
self.episode_id = episode_id
self.observations = []
self.actions = []
self.reward = []
self.done = False
def record(self, obs, action, reward):
self.observations.append(deepcopy(obs))
self.actions.append(deepcopy(action))
# Agent not win or not die
if reward != 0:
self.done = True
self.reward = [reward] * self.get_num_steps()
def get_num_steps(self):
return len(self.observations)
# Environment wrapper
class Stimulator:
def __init__(self, env):
self.env = env
self.episodes = []
def record(self, agents, obs, actions, rewards):
for agent_id in range(len(agents)):
agent = agents[agent_id]
# If an agent is dead (or win) we should not record a history
if not agent.done:
agent.record(obs[agent_id], actions[agent_id], rewards[agent_id])
def stimulate(self, num_rollouts):
for cur_episode in tqdm(range(num_rollouts)):
# Create a history for each agent
agents = []
for agent_id in range(4):
agents.append(Episode(agent_id, cur_episode))
done = False
# Obtain initial observations
obs = self.env.reset()
try:
while not done:
# FUCK self.env.act change "POSITION"!!!!
obs_to_save = deepcopy(obs)
# Produce actions
actions = self.env.act(obs)
# Make an episode step. Save an observations as new_obs, because we want to record previous one
obs, rewards, done, _ = self.env.step(actions)
# Record observations and actions
self.record(agents, obs_to_save, actions, rewards)
except:
print("Error occurs")
continue
self.episodes.extend(agents)
def get_episodes(self):
return deepcopy(self.episodes)
# +
# Instantiate the environment
config = ffa_competition_env()
env = Pomme(**config["env_kwargs"])
# Random seed
env.seed(0)
# Add agents
agents = []
for agent_id in range(4):
agents.append(SimpleAgent(config["agent"](agent_id, config["game_type"])))
env.set_agents(agents)
env.set_init_game_state(None)
# -
# Generate training data
stimulator = Stimulator(env)
stimulator.stimulate(initial_rollouts)
episodes = stimulator.get_episodes()
len(episodes)
for episode in tqdm(episodes):
for observation in episode.observations:
board = observation['board']
pos = observation['position']
if board[pos[0], pos[1]] != (episode.agent_id + 10):
print("ERROR")
# ## Win/Lose in corners
# +
wins = []
loses = []
for episode in episodes:
if episode.reward[-1] == -1:
loses.append(episode.agent_id)
elif episode.reward[-1] == 1:
wins.append(episode.agent_id)
else:
print("FUCK")
plt.hist(loses)
plt.hist(wins)
plt.show()
# -
np.bincount(loses)
# ## Episode length
# +
episode_length = []
for episode in episodes:
episode_length.append(episode.get_num_steps())
plt.figure(figsize=(17,4))
plt.bar(range(0, len(episode_length)), episode_length)
plt.plot([0, len(episode_length)], [np.mean(episode_length), np.mean(episode_length)], 'r')
plt.legend(['mean', 'episode length'])
plt.show()
# -
np.min(episode_length), np.max(episode_length), np.sum(np.array(episode_length) < 30)
# ## Rewards
# +
nr_wins = [0] * 4
nr_loose = [0] * 4
nr_tie = [0] * 4
average_score = [[], [], [], []]
for episode in episodes:
if episode.reward[-1] == 1:
nr_wins[episode.agent_id] += 1
else:
nr_loose[episode.agent_id] += 1
average_score[episode.agent_id].append(episode.reward[-1])
average_score[0] = np.mean(average_score[0])
average_score[1] = np.mean(average_score[1])
average_score[2] = np.mean(average_score[2])
average_score[3] = np.mean(average_score[3])
plt.figure(figsize=(17,4))
plt.subplot(141)
plt.bar(range(0,4), nr_wins)
plt.title('nr_wins')
plt.subplot(142)
plt.bar(range(0,4), nr_loose)
plt.title('nr_loose')
plt.subplot(143)
plt.bar(range(0,4), average_score)
plt.title('average_score')
plt.show()
# -
# ## Consequtive actions
actions_length = []
for episode in episodes:
current_action = episode.actions[0]
action_length = 0
for action in episode.actions:
if current_action != action:
actions_length.append(action_length)
current_action = action
action_length = 0
action_length += 1
np.mean(actions_length), np.median(actions_length), np.max(actions_length)
plt.figure(figsize=(17,10))
plt.plot(actions_length)
plt.plot([np.mean(actions_length)] * len(actions_length))
plt.title('action length')
plt.show()
# ## Actions frequency
# +
actions_count = np.zeros(6)
for episode in episodes:
for action in episode.actions:
actions_count[action] += 1
plt.figure(figsize=(17,10))
plt.bar(range(0,6), actions_count)
plt.title('actions count')
plt.show()
# -
# # Some data preprocessing
# Remove episodes with too small length. Maybe it is suicide.
episodes = [episode for episode in episodes if episode.get_num_steps() > 35 or episode.reward[-1] == 1]
len(episodes)
# Let's remove consequtive actions. I will use threshold 12.
# +
threshold = 12
for episode in tqdm(episodes):
# New observations
obs = []
act = []
rew = []
# Initial values
cur_action = episode.actions[0]
seq_length = 0
for ind in range(len(episode.actions)):
# Current action
action = episode.actions[ind]
# If an action the same
if action == cur_action:
seq_length += 1
else:
seq_length = 0
cur_action = action
# If a sequence is less than threashold
if seq_length <= threshold:
obs.append(episode.observations[ind])
act.append(episode.actions[ind])
rew.append(episode.reward[ind])
# Save preprocessed observations
episode.actions = act
episode.observations = obs
episode.reward = rew
# -
# Let's rerun plots.
# ## Save stuff
# +
observations = []
actions = []
rewards = []
for episode in episodes:
observations.extend(episode.observations)
actions.extend(episode.actions)
rewards.extend(episode.reward)
observations = np.array(observations)
actions = np.array(actions)
rewards = np.array(rewards)
# -
np.save(train_data_obs, observations)
np.save(train_data_labels, actions)
np.save(train_data_reward, rewards)
observations = np.load(train_data_obs)
actions = np.load(train_data_labels)
rewards = np.load(train_data_reward)
observations.shape, actions.shape, rewards.shape
# ## Featurize
def featurize(obs):
shape = (BOARD_SIZE, BOARD_SIZE, 1)
def get_matrix(board, key):
res = board[key]
return res.reshape(shape).astype(np.float32)
def get_map(board, item):
map = np.zeros(shape)
map[board == item] = 1
return map
board = get_matrix(obs, 'board')
path_map = get_map(board, 0) # Empty space
rigid_map = get_map(board, 1) # Rigid = 1
wood_map = get_map(board, 2) # Wood = 2
bomb_map = get_map(board, 3) # Bomb = 3
flames_map = get_map(board, 4) # Flames = 4
fog_map = get_map(board, 5) # TODO: not used for first two stages Fog = 5
extra_bomb_map = get_map(board, 6) # ExtraBomb = 6
incr_range_map = get_map(board, 7) # IncrRange = 7
kick_map = get_map(board, 8) # Kick = 8
skull_map = get_map(board, 9) # Skull = 9
position = obs["position"]
my_position = np.zeros(shape)
my_position[position[0], position[1], 0] = 1
team_mates = get_map(board, obs["teammate"].value) # TODO during documentation it should be an array
enemies = np.zeros(shape)
for enemy in obs["enemies"]:
enemies[board == enemy.value] = 1
bomb_blast_strength = get_matrix(obs, 'bomb_blast_strength')
bomb_life = get_matrix(obs, 'bomb_life')
ammo = np.full((BOARD_SIZE, BOARD_SIZE, 1), obs["ammo"])
blast_strength = np.full((BOARD_SIZE, BOARD_SIZE, 1), obs["blast_strength"])
can_kick = np.full((BOARD_SIZE, BOARD_SIZE, 1), int(obs["can_kick"]))
obs = np.concatenate([my_position, enemies, team_mates, path_map, rigid_map,
wood_map, bomb_map, flames_map, fog_map, extra_bomb_map,
incr_range_map, kick_map, skull_map, bomb_blast_strength,
bomb_life, ammo, blast_strength, can_kick], axis=2)
return obs.astype(np.int16)
observations_map = []
for obs in tqdm(observations):
observations_map.append(featurize(obs))
np.save(train_data_obs_map, observations_map)
| rl_agent/generate_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_pytorch_p27)
# language: python
# name: conda_pytorch_p27
# ---
# %matplotlib inline
#
# Neural Networks
# ===============
#
# Neural networks can be constructed using the ``torch.nn`` package.
#
# Now that you had a glimpse of ``autograd``, ``nn`` depends on
# ``autograd`` to define models and differentiate them.
# An ``nn.Module`` contains layers, and a method ``forward(input)``\ that
# returns the ``output``.
#
# For example, look at this network that classfies digit images:
#
# .. figure:: /_static/img/mnist.png
# :alt: convnet
#
# convnet
#
# It is a simple feed-forward network. It takes the input, feeds it
# through several layers one after the other, and then finally gives the
# output.
#
# A typical training procedure for a neural network is as follows:
#
# - Define the neural network that has some learnable parameters (or
# weights)
# - Iterate over a dataset of inputs
# - Process input through the network
# - Compute the loss (how far is the output from being correct)
# - Propagate gradients back into the network’s parameters
# - Update the weights of the network, typically using a simple update rule:
# ``weight = weight - learning_rate * gradient``
#
# Define the network
# ------------------
#
# Let’s define this network:
#
#
# +
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# 1 input image channel, 6 output channels, 5x5 square convolution
# kernel
self.conv1 = nn.Conv2d(1, 6, 5)
self.conv2 = nn.Conv2d(6, 16, 5)
# an affine operation: y = Wx + b
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
# Max pooling over a (2, 2) window
x = F.max_pool2d(F.relu(self.conv1(x)), (2, 2))
# If the size is a square you can only specify a single number
x = F.max_pool2d(F.relu(self.conv2(x)), 2)
x = x.view(-1, self.num_flat_features(x))
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
def num_flat_features(self, x):
size = x.size()[1:] # all dimensions except the batch dimension
num_features = 1
for s in size:
num_features *= s
return num_features
net = Net()
print(net)
# -
# You just have to define the ``forward`` function, and the ``backward``
# function (where gradients are computed) is automatically defined for you
# using ``autograd``.
# You can use any of the Tensor operations in the ``forward`` function.
#
# The learnable parameters of a model are returned by ``net.parameters()``
#
#
params = list(net.parameters())
print(len(params))
print(params[0].size()) # conv1's .weight
# The input to the forward is an ``autograd.Variable``, and so is the output.
# Note: Expected input size to this net(LeNet) is 32x32. To use this net on
# MNIST dataset,please resize the images from the dataset to 32x32.
#
#
input = Variable(torch.randn(1, 1, 32, 32))
out = net(input)
print(out)
# Zero the gradient buffers of all parameters and backprops with random
# gradients:
#
#
net.zero_grad()
out.backward(torch.randn(1, 10))
# <div class="alert alert-info"><h4>Note</h4><p>``torch.nn`` only supports mini-batches The entire ``torch.nn``
# package only supports inputs that are a mini-batch of samples, and not
# a single sample.
#
# For example, ``nn.Conv2d`` will take in a 4D Tensor of
# ``nSamples x nChannels x Height x Width``.
#
# If you have a single sample, just use ``input.unsqueeze(0)`` to add
# a fake batch dimension.</p></div>
#
# Before proceeding further, let's recap all the classes you’ve seen so far.
#
# **Recap:**
# - ``torch.Tensor`` - A *multi-dimensional array*.
# - ``autograd.Variable`` - *Wraps a Tensor and records the history of
# operations* applied to it. Has the same API as a ``Tensor``, with
# some additions like ``backward()``. Also *holds the gradient*
# w.r.t. the tensor.
# - ``nn.Module`` - Neural network module. *Convenient way of
# encapsulating parameters*, with helpers for moving them to GPU,
# exporting, loading, etc.
# - ``nn.Parameter`` - A kind of Variable, that is *automatically
# registered as a parameter when assigned as an attribute to a*
# ``Module``.
# - ``autograd.Function`` - Implements *forward and backward definitions
# of an autograd operation*. Every ``Variable`` operation, creates at
# least a single ``Function`` node, that connects to functions that
# created a ``Variable`` and *encodes its history*.
#
# **At this point, we covered:**
# - Defining a neural network
# - Processing inputs and calling backward.
#
# **Still Left:**
# - Computing the loss
# - Updating the weights of the network
#
# Loss Function
# -------------
# A loss function takes the (output, target) pair of inputs, and computes a
# value that estimates how far away the output is from the target.
#
# There are several different
# `loss functions <http://pytorch.org/docs/nn.html#loss-functions>`_ under the
# nn package .
# A simple loss is: ``nn.MSELoss`` which computes the mean-squared error
# between the input and the target.
#
# For example:
#
#
# +
output = net(input)
target = Variable(torch.arange(1, 11)) # a dummy target, for example
criterion = nn.MSELoss()
loss = criterion(output, target)
print(loss)
# -
# Now, if you follow ``loss`` in the backward direction, using it’s
# ``.grad_fn`` attribute, you will see a graph of computations that looks
# like this:
#
# ::
#
# input -> conv2d -> relu -> maxpool2d -> conv2d -> relu -> maxpool2d
# -> view -> linear -> relu -> linear -> relu -> linear
# -> MSELoss
# -> loss
#
# So, when we call ``loss.backward()``, the whole graph is differentiated
# w.r.t. the loss, and all Variables in the graph will have their
# ``.grad`` Variable accumulated with the gradient.
#
# For illustration, let us follow a few steps backward:
#
#
print(loss.grad_fn) # MSELoss
print(loss.grad_fn.next_functions[0][0]) # Linear
print(loss.grad_fn.next_functions[0][0].next_functions[0][0]) # ReLU
# Backprop
# --------
# To backpropagate the error all we have to do is to ``loss.backward()``.
# You need to clear the existing gradients though, else gradients will be
# accumulated to existing gradients
#
#
# Now we shall call ``loss.backward()``, and have a look at conv1's bias
# gradients before and after the backward.
#
#
# +
net.zero_grad() # zeroes the gradient buffers of all parameters
print('conv1.bias.grad before backward')
print(net.conv1.bias.grad)
loss.backward()
print('conv1.bias.grad after backward')
print(net.conv1.bias.grad)
# -
# Now, we have seen how to use loss functions.
#
# **Read Later:**
#
# The neural network package contains various modules and loss functions
# that form the building blocks of deep neural networks. A full list with
# documentation is `here <http://pytorch.org/docs/nn>`_
#
# **The only thing left to learn is:**
#
# - updating the weights of the network
#
# Update the weights
# ------------------
# The simplest update rule used in practice is the Stochastic Gradient
# Descent (SGD):
#
# ``weight = weight - learning_rate * gradient``
#
# We can implement this using simple python code:
#
# .. code:: python
#
# learning_rate = 0.01
# for f in net.parameters():
# f.data.sub_(f.grad.data * learning_rate)
#
# However, as you use neural networks, you want to use various different
# update rules such as SGD, Nesterov-SGD, Adam, RMSProp, etc.
# To enable this, we built a small package: ``torch.optim`` that
# implements all these methods. Using it is very simple:
#
#
# +
import torch.optim as optim
# create your optimizer
optimizer = optim.SGD(net.parameters(), lr=0.01)
# in your training loop:
optimizer.zero_grad() # zero the gradient buffers
output = net(input)
loss = criterion(output, target)
loss.backward()
optimizer.step() # Does the update
| neural_networks_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sankalpachowdhury/Galaxy-Classification-using-CNN/blob/master/Model1_Galaxy_classification_sankalpa_v3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="t254vRoD7cyj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="23261e10-0baf-4778-d4e1-02dfa3c1e16d"
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
import tensorflow as tf
import keras
from keras.models import Sequential
import matplotlib.pyplot as plt
# + id="K9BQa8C07zyj" colab_type="code" colab={}
# %matplotlib inline
import os, random, shutil
from keras_preprocessing import image
# + id="CJC996oP7r33" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 128} outputId="246c9d55-8882-4822-8f65-e65d3eef055a"
from google.colab import drive
drive.mount('/content/drive')
# + id="Ro87h7kL7tNT" colab_type="code" colab={}
# #!unzip "/content/drive/My Drive/Astronomical /Test.zip"
# + id="RZbr7GaW8L1k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="19146d5e-f808-4c89-aa8a-107e61eb26f6"
# !unzip "/content/drive/My Drive/Galaxy_classification_project/images_training_rev1.zip"
# + id="4PCnUgLA8WKk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 272} outputId="429bf51d-e4bf-4fe0-9dfe-27bbaa77b5e7"
df = pd.read_csv('/content/drive/My Drive/Galaxy_classification_project/training_solutions_rev1.csv')
#df_t = pd.read_csv('/content/drive/My Drive/Astronomical /Test.csv')
print(df.head)
# + id="7c_K62av8bKM" colab_type="code" colab={}
cols = df.columns
new = list(map(lambda s: s.replace('Class','Q'), cols))
df.columns = new
# + id="qi7kqsdd-WjQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1ecfbb3b-3428-45cd-a41f-5eed44bcdf25"
df.shape
# + id="-qRCpKpI-aZT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="14a7e1eb-8417-46e5-e191-8a146ee7cf5d"
df.head(5)
# + id="WkWHsUBu-iij" colab_type="code" colab={}
ellipticals = df[(df['Q1.1']>0.8) & (df['Q7.1']>0.4)]['GalaxyID'].tolist()
#ellipticals
# + id="Zd8lzfvk-jbk" colab_type="code" colab={}
lenticulars = df[(df['Q1.1']>0.8) & (df['Q7.2']>0.4)]['GalaxyID'].tolist()
#lenticulars
# + id="2LZGJZzq-r21" colab_type="code" colab={}
spirals = df[(df['Q1.2']>0.8) & (df['Q2.1']>0.4)]['GalaxyID'].tolist()
#spirals
# + id="v-MHLYFC-30N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="955e358d-c632-4f56-8d2d-baa802bd22b4"
print('Total number of elliptical examples: ', len(ellipticals))
print('Total number of lenticular examples: ', len(lenticulars))
print('Total number of spiral examples: ', len(spirals))
# + id="Gtocen_ND1s9" colab_type="code" colab={}
source_path = '/content/images_training_rev1'
dest_path = '/content/data'
# + id="AurD3A0cHZCj" colab_type="code" colab={}
def _proc_images(src, dst, label, arr, percent):
train_dir = os.path.join(dst, 'train')
val_dir = os.path.join(dst, 'validation')
train_dest = os.path.join(train_dir, label)
val_dest = os.path.join(val_dir, label)
if not os.path.exists(train_dest):
os.makedirs(train_dest)
if not os.path.exists(val_dest):
os.makedirs(val_dest)
random.shuffle(arr)
idx = int(len(arr)*percent)
for i in arr[0:idx]:
shutil.copyfile(os.path.join(src, str(i)+'.jpg'), os.path.join(train_dest, str(i)+'.jpg'))
for i in arr[idx:]:
shutil.copyfile(os.path.join(src, str(i)+'.jpg'), os.path.join(val_dest, str(i)+'.jpg'))
print(label, 'done!')
# + id="kk-lOaJjHy_m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="0ecff43f-9dad-4896-ef55-cd926068ae2e"
_proc_images(source_path, dest_path, 'elliptical', ellipticals, 0.9)
_proc_images(source_path, dest_path, 'lenticular', lenticulars, 0.9)
_proc_images(source_path, dest_path, 'spiral', spirals, 0.9)
# + [markdown] id="nsDF2sEfIq7E" colab_type="text"
# Data Processing
# + id="rUsadeQEIt1s" colab_type="code" colab={}
import os
import PIL
import PIL.Image
# + id="1iNz4OXnISWF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 120} outputId="1d363f83-8229-4a2d-f8ff-4fdf305be3bd"
'''
import matplotlib.pyplot as plt
from tqdm import tqdm
from skimage.transform import resize
ORIG_SHAPE = (424,424)
CROP_SIZE = (256,256)
IMG_SHAPE = (64,64)
def get_image(path, x1,y1, shape, crop_size):
x = plt.imread(path)
x = x[x1:x1+crop_size[0], y1:y1+crop_size[1]]
# x = resize(x, shape)
x = x/255.
return x
def get_all_images(path,df,shape=IMG_SHAPE, crop_size=CROP_SIZE, orig_size = ORIG_SHAPE, r=600):
x1 = (ORIG_SHAPE[0]-CROP_SIZE[0])//2
y1 = (ORIG_SHAPE[1]-CROP_SIZE[1])//2
sel = df.values
ids = sel[:r,0].astype(int).astype(str)
y_batch = sel[:r,1:]
x_batch = []
for i in tqdm(ids):
x = get_image(path+i+'.jpg', x1,y1, shape=shape, crop_size=crop_size)
x_batch.append(x)
print(i)
x_batch = np.array(x_batch)
return x_batch, y_batch
x_train, y_train = get_all_images('/content/Train/',df)
x_test, y_test = get_all_images('/content/Test/',df_t)
print(x_train.shape)
print(y_train.shape)
print(type(y_train))
print(x_test.shape)
print(y_test.shape)
print(type(y_test))
'''
# + id="GIWtRTXtJv9M" colab_type="code" colab={}
train_dir = 'data/train'
validation_dir = 'data/validation'
# make a directory in colabs "data" and copy the path.
# + id="_eD_EfeBJyVo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="df45a97a-eb9a-4c37-c310-423406e5951a"
total_train = 0
for c in ['elliptical', 'lenticular', 'spiral']:
total_train += len(os.listdir(os.path.join(train_dir, c)))
print('Total train:', total_train)
total_validation = 0
for c in ['elliptical', 'lenticular', 'spiral']:
total_validation += len(os.listdir(os.path.join(validation_dir, c)))
print('Total validation:', total_validation)
# + id="ltqDEaLKKPBg" colab_type="code" colab={}
# parameters
target = (150, 150)
batch_size = 32
# + id="Qloo1BtqKJbY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="ff6c80ba-4d57-47f0-80af-226f384a2184"
train_datagen = tf.keras.preprocessing.image.ImageDataGenerator(
rescale=1.0/255,
rotation_range=25,
width_shift_range=.15,
height_shift_range=.15,
horizontal_flip=True,
zoom_range=0.2)
validation_datagen = tf.keras.preprocessing.image.ImageDataGenerator(
rescale=1.0/255.)
train_generator = train_datagen.flow_from_directory(train_dir,
target_size=target,
batch_size=batch_size,
shuffle=True,
class_mode='categorical')
validation_generator = train_datagen.flow_from_directory(validation_dir,
target_size=target,
batch_size=batch_size,
shuffle=True,
class_mode='categorical')
# + id="hSlBtPOmMN5I" colab_type="code" colab={}
r_coeff = 0.001
# + id="usuXy04WL3s8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 120} outputId="ad8c2091-a32b-4053-92c3-ac6e77d6d02c"
'''
from keras import regularizers
model = keras.models.Sequential([keras.layers.Conv2D(32,(3,3),activation='relu',input_shape=(256,256,3)),
keras.layers.Conv2D(64,(3,3),activation='relu', kernel_regularizer = regularizers.l2(r_coeff)),
keras.layers.MaxPooling2D((20,20),strides=(2,2)),
keras.layers.Conv2D(128,(3,3),activation='relu',kernel_regularizer = regularizers.l2(r_coeff)),
keras.layers.Conv2D(128,(3,3),activation='relu',kernel_regularizer = regularizers.l2(r_coeff)),
keras.layers.MaxPooling2D((2,2)),# Deafault takes stride = (2,2)
keras.layers.Flatten(),
keras.layers.Dense(512,activation='relu',kernel_regularizer = regularizers.l2(r_coeff)),
keras.layers.Dense(512,activation='relu',kernel_regularizer = regularizers.l2(r_coeff)),
keras.layers.Dense(3,activation = 'softmax',kernel_regularizer = regularizers.l2(r_coeff))
])
#model1.add()
model.summary()
'''
# + id="dmwiARwXz-k-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 672} outputId="bce5360e-e6c4-4c46-af2d-490412fcca84"
from keras import regularizers
model2 = tf.keras.models.Sequential([
# first convolution layer, input is an 150x150 image x3 colors
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(150, 150, 3)),
tf.keras.layers.MaxPooling2D(2, 2),
# second convolution layer
tf.keras.layers.Conv2D(32, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# third convolution layer
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# fourth convolution layer
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
#tf.keras.layers.Conv2D(256, (3,3), activation='relu'),
#tf.keras.layers.MaxPooling2D(2,2),
# flatten the image pixels
tf.keras.layers.Flatten(),
# 512 neuron fully connected hidden layer
#tf.keras.layers.Dense(1024, activation='relu'),
tf.keras.layers.Dense(512, activation='relu'),
#tf.keras.layers.Dense(256, activation='relu'),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(3, activation='softmax')
])
#model1.add()
#tf.keras.layers.Dropout(0.5),
model2.summary()
# + id="0R7NnmynUVeT" colab_type="code" colab={}
from keras import regularizers
from keras.layers import BatchNormalization
model3 = tf.keras.models.Sequential([
# first convolution layer, input is an 150x150 image x3 colors
tf.keras.layers.Conv2D(64, (3,3), activation='relu', input_shape=(180, 180, 3)),
tf.keras.layers.MaxPooling2D(2, 2),
# second convolution layer
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# third convolution layer
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# fourth convolution layer
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(256, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(256, (3,3), activation='relu'),
#tf.keras.layers.MaxPooling2D(2,2),
# flatten the image pixels
tf.keras.layers.Flatten(),
tf.keras.layers.BatchNormalization(),
# 512 neuron fully connected hidden layer
tf.keras.layers.Dense(256, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.BatchNormalization(),
#tf.keras.layers.Dense(256, activation='relu'),
#tf.keras.layers.BatchNormalization(),
#tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(3, activation='softmax')
])
#model1.add()
#tf.keras.layers.Dropout(0.5),
model3.summary()
# + id="rojiuduiMTOK" colab_type="code" colab={}
from tensorflow.keras.optimizers import Adam
model2.compile(optimizer='Adam',
loss = 'categorical_crossentropy',
metrics = ['acc'])
#(learning_rate=0.001)
# + id="Sws0F4VkM2Wq" colab_type="code" colab={}
EPOCHS = 60
# + id="n4iX_jyEMhB0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="64be9272-a269-407c-a5f1-7471d669bf00"
from keras.callbacks import TensorBoard
from time import time
# Create a TensorBoard instance with the path to the logs directory
tensorboard = TensorBoard(log_dir='logs/{}'.format(time()))
history = model2.fit_generator(train_generator,
epochs=EPOCHS,
validation_data=validation_generator,
#callbacks=[plot_lossesd],
verbose=1)
# + id="Py4P74vyn4I4" colab_type="code" colab={}
# + id="uzYArx-mT7bl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 499} outputId="34aec92a-2c27-41b9-c743-93f6330d9520"
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(len(acc)) # range for the number of epochs
plt.figure(figsize=(16, 8))
plt.subplot(1, 2, 1)
plt.plot(epochs_range, acc, label='Training Accuracy')
plt.plot(epochs_range, val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(1, 2, 2)
plt.plot(epochs_range, loss, label='Training Loss')
plt.plot(epochs_range, val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.savefig('./plots-v2.png')
plt.show()
# + id="j2eJO0-TMcds" colab_type="code" colab={}
'''
model.fit(x_train, y_train, batch_size = 64, epochs= EPOCHS, validation_data=(x_test, y_test))
history = model.fit(train_generator,
epochs=EPOCHS,
validation_data=validation_generator,
verbose=1)
'''
# + id="iK8pAY0NWPwH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="483f4d88-59d5-4c38-8d6b-edd261254edd"
model.save('galaxy-convnet-v2.h5')
# + id="N78dssyOYPWB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 454} outputId="85ab4858-6a51-42a5-f5f3-a9cc050a60e1"
# !pip install Tensorboard
# + [markdown] id="0Sni_Xft4X-D" colab_type="text"
# filter size = (fh=3, fw = 3, fc = 3)
# nc[l+1] = 32
# w.shape = 32*(3*3*3) = 864
# b.shape = 32
# total = 896
# + [markdown] id="NBiXZH6hfJjg" colab_type="text"
# TUMI OFF KORE DIYO LAPTOP< OKK
# + id="FX0-vpNzfX08" colab_type="code" colab={}
tensorboard --logdir=logs/
# + id="mXopkgLnB3ht" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 178} outputId="36f606c7-1548-4f12-95b1-805105f40693"
# launch TensorBoard (data won't show up until after the first epoch)
tensorboard("logs/run_a")
# + id="vea2aSm8CT7U" colab_type="code" colab={}
# a minimal example (sort of)
class PlotLosses(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.i = 0
self.x = []
self.losses = []
self.val_losses = []
self.fig = plt.figure()
self.logs = []
def on_epoch_end(self, epoch, logs={}):
self.logs.append(logs)
self.x.append(self.i)
self.losses.append(logs.get('loss'))
self.val_losses.append(logs.get('val_loss'))
self.i += 1
clear_output(wait=True)
plt.plot(self.x, self.losses, label="loss")
plt.plot(self.x, self.val_losses, label="val_loss")
plt.legend()
plt.show();
plot_lossesd = PlotLosses()
# + id="VmFlLF2vCmn7" colab_type="code" colab={}
| Model testing/Model1_Galaxy_classification_sankalpa_v3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
from SmartAnno.utils.ReviewRBInit import ReviewRBInit
from SmartAnno.utils.ReviewRBLoop import ReviewRBLoop
from SmartAnno.utils.ReviewMLInit import ReviewMLInit
from SmartAnno.utils.ReviewMLLoop import ReviewMLLoop
from SmartAnno.gui.Workflow import Workflow
from sqlalchemy_dao import Dao
from SmartAnno.db.ORMs import Document
from SmartAnno.utils.IntroStep import IntroStep
from SmartAnno.gui.PreviousNextWidgets import PreviousNextHTML
import sqlalchemy_dao
import os
from conf.ConfigReader import ConfigReader
from SmartAnno.models.logistic.LogisticBOWClassifier import LogisticBOWClassifier
import logging
logging.getLogger().setLevel(logging.DEBUG)
ConfigReader('../conf/smartanno_conf.json')
wf=Workflow()
rb=ReviewRBInit(name="rb_review_init")
wf.append(rb)
rv = ReviewRBLoop(name='rb_review', rush_rule='../conf/rush_rules.tsv')
wf.append(rv)
wf.append(PreviousNextHTML('<h2>Congratuations!</h2><h4>You have finished the initial review on the rule-base preannotations. </h4>',
name='intro'))
wf.append(ReviewMLInit(name='ml_review_init'))
wf.append(ReviewMLLoop(name='ml_review',ml_classifier_cls=LogisticBOWClassifier))
wf.filters={'Eng':['heart'],'NotEng':['exam']}
wf.types=['TypeA','TypeB']
wf.task_id=1
wf.umls_extended={}
wf.we_extended={}
wf.dao=Dao('sqlite+pysqlite:///../data/demo.sqlite', sqlalchemy_dao.POOL_DISABLED)
# -
wf.start()
| SmartAnno/test/TestReviewRB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
from __future__ import print_function
# %matplotlib inline
import tensorflow as tf
import numpy
import matplotlib.pyplot as plt
import pandas as pd
# -
rng = numpy.random
# Different parameters for learning
learning_rate = 0.01
training_epochs = 1000
display_step = 100
# +
#import data
df= pd.read_csv("../data/mon.csv")
#divisione train & test set modo 1
#train=df.sample(frac=0.8,random_state=200)
#test=df.drop(train.index)
#divisione train & test set modo 2
from sklearn.model_selection import train_test_split
train, test = train_test_split(df, test_size = 0.2)
# -
train.head(5)
#creo train & test set
train_X = train['sys']
train_Y = train['hr']
n_samples = train_X.shape[0]
test_X = test['sys']
test_Y = test['hr']
type(train_X)
# +
# Training Data
#train_X = numpy.asarray([3.3,4.4,5.5,6.71,6.93,4.168,9.779,6.182,7.59,2.167,7.042,10.791,5.313,7.997,5.654,9.27,3.1])
#train_Y = numpy.asarray([1.7,2.76,2.09,3.19,1.694,1.573,3.366,2.596,2.53,1.221,2.827,3.465,1.65,2.904,2.42,2.94,1.3])
#n_samples = train_X.shape[0]
# -
# Create placeholder for providing inputs
X = tf.placeholder("float")
Y = tf.placeholder("float")
# create weights and bias and initialize with random number
W = tf.Variable(rng.randn(), name="weight")
b = tf.Variable(rng.randn(), name="bias")
# Construct a linear model using Y=WX+b
pred = tf.add(tf.mul(X, W), b)
# Calculate Mean squared error
cost = tf.reduce_sum(tf.pow(pred-Y, 2))/(2*n_samples)
# Gradient descent to minimize mean sequare error
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(cost)
# Initializing the variables
# init = tf.initialize_all_variables
init = tf.global_variables_initializer()
# Launch the graph
with tf.Session() as sess:
sess.run(init)
print("Training started")
# Fit all training data
for epoch in range(training_epochs):
for (x, y) in zip(train_X, train_Y):
#create small batch of trining and testing data and feed it to model
sess.run(optimizer, feed_dict={X: x, Y: y})
# Display training information after each N step
if (epoch+1) % display_step == 0:
c = sess.run(cost, feed_dict={X: train_X, Y:train_Y})
print("Epoch:", '%04d' % (epoch+1), "cost=", "{:.9f}".format(c), \
"W=", sess.run(W), "b=", sess.run(b))
print("Training completed")
training_cost = sess.run(cost, feed_dict={X: train_X, Y: train_Y})
print("Training cost=", training_cost, "W=", sess.run(W), "b=", sess.run(b), '\n')
# Testing
print("Testing started")
#test_X = numpy.asarray([6.83, 4.668, 8.9, 7.91, 5.7, 8.7, 3.1, 2.1])
#test_Y = numpy.asarray([1.84, 2.273, 3.2, 2.831, 2.92, 3.24, 1.35, 1.03])
#Calculate Mean square error
print("Calculate Mean square error")
testing_cost = sess.run(tf.reduce_sum(tf.pow(pred-Y, 2)) / (2 * test_X.shape[0]),feed_dict={X: test_X, Y: test_Y}) # same function as cost above
print("Testing cost=", testing_cost)
print("Absolute mean square loss difference:", abs(training_cost - testing_cost))
plt.plot(test_X, test_Y, 'bo', label='Testing data')
plt.plot(train_X, sess.run(W) * train_X + sess.run(b), label='Fitted line')
plt.legend()
plt.show()
# +
# -
| develop/2017-03-12-SG-TestRegressioneLineareTF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="fig/cover-small.jpg">
#
# *Este notebook es una adaptación realizada por <NAME> del material "[Whirlwind Tour of Python](http://www.oreilly.com/programming/free/a-whirlwind-tour-of-python.csp)" de Jake VanderPlas; tanto el [contenido original](https://github.com/jakevdp/WhirlwindTourOfPython) como la [adpatación actual](https://github.com/rrgalvan/PythonIntroMasterMatemat)] están disponibles en Github.*
#
#
# *The text and code are released under the [CC0](https://github.com/jakevdp/WhirlwindTourOfPython/blob/master/LICENSE) license; see also the companion project, the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# <!--NAVIGATION-->
# < [Built-In Data Structures](06-Built-in-Data-Structures.ipynb) | [Contents](Index.ipynb) | [Defining and Using Functions](08-Defining-Functions.ipynb) >
# # Control Flow
# *Control flow* is where the rubber really meets the road in programming.
# Without it, a program is simply a list of statements that are sequentially executed.
# With control flow, you can execute certain code blocks conditionally and/or repeatedly: these basic building blocks can be combined to create surprisingly sophisticated programs!
#
# Here we'll cover *conditional statements* (including "``if``", "``elif``", and "``else``"), *loop statements* (including "``for``" and "``while``" and the accompanying "``break``", "``continue``", and "``pass``").
# ## Conditional Statements: ``if``-``elif``-``else``:
# Conditional statements, often referred to as *if-then* statements, allow the programmer to execute certain pieces of code depending on some Boolean condition.
# A basic example of a Python conditional statement is this:
# +
x = -15
if x == 0:
print(x, "is zero")
elif x > 0:
print(x, "is positive")
elif x < 0:
print(x, "is negative")
else:
print(x, "is unlike anything I've ever seen...")
# -
# Note especially the use of colons (``:``) and whitespace to denote separate blocks of code.
#
# Python adopts the ``if`` and ``else`` often used in other languages; its more unique keyword is ``elif``, a contraction of "else if".
# In these conditional clauses, ``elif`` and ``else`` blocks are optional; additionally, you can optinally include as few or as many ``elif`` statements as you would like.
# ## ``for`` loops
# Loops in Python are a way to repeatedly execute some code statement.
# So, for example, if we'd like to print each of the items in a list, we can use a ``for`` loop:
for N in [2, 3, 5, 7]:
print(N, end=' ') # print all on same line
# Notice the simplicity of the ``for`` loop: we specify the variable we want to use, the sequence we want to loop over, and use the "``in``" operator to link them together in an intuitive and readable way.
# More precisely, the object to the right of the "``in``" can be any Python *iterator*.
# An iterator can be thought of as a generalized sequence, and we'll discuss them in [Iterators](10-Iterators.ipynb).
#
# For example, one of the most commonly-used iterators in Python is the ``range`` object, which generates a sequence of numbers:
for i in range(10):
print(i, end=' ')
# Note that the range starts at zero by default, and that by convention the top of the range is not included in the output.
#
# ---
# - **Nota**: obsérvese que ``range(n)``$=[0,n)\cap\mathbb{Z}$ y en general ``range(m,n)``$=[m,n)\cap\mathbb{Z}$
# ---
# Range objects can also have more complicated values:
# range from 5 to 10
list(range(5, 10))
# range from 0 to 10 by 2
list(range(0, 10, 2))
# You might notice that the meaning of ``range`` arguments is very similar to the slicing syntax that we covered in [Lists](06-Built-in-Data-Structures.ipynb#Lists).
#
# Note that the behavior of ``range()`` is one of the differences between Python 2 and Python 3: in Python 2, ``range()`` produces a list, while in Python 3, ``range()`` produces an iterable object.
# ## ``while`` loops
# The other type of loop in Python is a ``while`` loop, which iterates until some condition is met:
i = 0
while i < 10:
print(i, end=' ')
i += 1
# The argument of the ``while`` loop is evaluated as a boolean statement, and the loop is executed until the statement evaluates to False.
# ## ``break`` and ``continue``: Fine-Tuning Your Loops
# There are two useful statements that can be used within loops to fine-tune how they are executed:
#
# - The ``break`` statement breaks-out of the loop entirely
# - The ``continue`` statement skips the remainder of the current loop, and goes to the next iteration
#
# These can be used in both ``for`` and ``while`` loops.
#
# Here is an example of using ``continue`` to print a string of odd numbers.
# In this case, the result could be accomplished just as well with an ``if-else`` statement, but sometimes the ``continue`` statement can be a more convenient way to express the idea you have in mind:
for n in range(20):
# if the remainder of n / 2 is 0, skip the rest of the loop
if n % 2 == 0:
continue
print(n, end=' ')
# Here is an example of a ``break`` statement used for a less trivial task.
# This loop will fill a list with all Fibonacci numbers up to a certain value:
# +
a, b = 0, 1
amax = 100
L = []
while True:
(a, b) = (b, a + b)
if a > amax:
break
L.append(a)
print(L)
# -
# Notice that we use a ``while True`` loop, which will loop forever unless we have a break statement!
# ## Loops with an ``else`` Block
# One rarely used pattern available in Python is the ``else`` statement as part of a ``for`` or ``while`` loop.
# We discussed the ``else`` block earlier: it executes if all the ``if`` and ``elif`` statements evaluate to ``False``.
# The loop-``else`` is perhaps one of the more confusingly-named statements in Python; I prefer to think of it as a ``nobreak`` statement: that is, the ``else`` block is executed only if the loop ends naturally, without encountering a ``break`` statement.
#
# As an example of where this might be useful, consider the following (non-optimized) implementation of the *Sieve of Eratosthenes*, a well-known algorithm for finding prime numbers:
# +
L = []
nmax = 30
for n in range(2, nmax):
for factor in L:
if n % factor == 0:
break
else: # no break
L.append(n)
print(L)
# -
# The ``else`` statement only executes if none of the factors divide the given number.
# The ``else`` statement works similarly with the ``while`` loop.
# <!--NAVIGATION-->
# < [Built-In Data Structures](06-Built-in-Data-Structures.ipynb) | [Contents](Index.ipynb) | [Defining and Using Functions](08-Defining-Functions.ipynb) >
| 07-Control-Flow-Statements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fenotebook
# language: python
# name: fenotebook
# ---
# # Missing value imputation: DropMissingData
#
# Deletes rows with missing values.
#
# DropMissingData works both with numerical and categorical variables. When no variable list is passed, it will default to all variables in the dataset. In addition, in the parameter missing_only, we can indicate if we want to drop observations in all variables, or only for those that showed missing data during fit, that is, in the train set.
#
# **For this demonstration, we use the Ames House Prices dataset produced by Professor <NAME>:**
#
# <NAME> (2011) Ames, Iowa: Alternative to the Boston Housing
# Data as an End of Semester Regression Project, Journal of Statistics Education, Vol.19, No. 3
#
# http://jse.amstat.org/v19n3/decock.pdf
#
# https://www.tandfonline.com/doi/abs/10.1080/10691898.2011.11889627
#
# The version of the dataset used in this notebook can be obtained from [Kaggle](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data)
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from feature_engine.imputation import DropMissingData
# +
# load data
data = pd.read_csv('houseprice.csv')
# show first 5 rows
data.head()
# +
# let's separate into training and testing set
X_train, X_test, y_train, y_test = train_test_split(
data.drop(['Id', 'SalePrice'], axis=1), data['SalePrice'], test_size=0.3, random_state=0)
X_train.shape, X_test.shape
# +
# let's create an instance of the imputer
# we want to drop na only in 4 variables from the dataset
na_imputer = DropMissingData(
missing_only = True,
variables=['Alley', 'MasVnrType', 'LotFrontage', 'MasVnrArea'])
# imputer checks that the indicated variables have NA
# in the train set
na_imputer.fit(X_train)
# +
# variables from which observations with NA will be deleted
na_imputer.variables_
# +
# Number of observations with NA before the transformation
X_train[na_imputer.variables].isna().sum()
# +
# After the transformation the rows with NA values are
# deleted form the dataframe
train_t = na_imputer.transform(X_train)
test_t = na_imputer.transform(X_test)
# +
# Number of observations with NA after transformation
train_t[na_imputer.variables].isna().sum()
# +
# shape of dataframe before transformation
X_train.shape
# +
# shape of dataframe after transformation
train_t.shape
# +
# The "return_na_data" method, returns a dataframe that contains
# the observations with NA, that would be dropped if we applied
# the transform method
tmp = na_imputer.return_na_data(X_train)
tmp.shape
# +
# total obs - obs with NA = final dataframe shape
1022-963
# -
# ## Automatically select all variables
# +
# let's create an instance of the imputer
na_imputer = DropMissingData(missing_only=True)
# the transformer will find the variables with NA
na_imputer.fit(X_train)
# +
# variables with NA in the train set
na_imputer.variables_
# +
# Number of observations with NA
X_train[na_imputer.variables_].isna().sum()
# +
# After the transformation the rows with NA are deleted form the dataframe
train_t = na_imputer.transform(X_train)
test_t = na_imputer.transform(X_test)
# -
# Number of observations with NA after the transformation
train_t[na_imputer.variables_].isna().sum()
# +
# in this case, all observations will be dropped
# because all of them show NA at least in 1 variable
train_t.shape
# -
| examples/imputation/DropMissingData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: widgets-tutorial
# language: python
# name: widgets-tutorial
# ---
# <!--NAVIGATION-->
# < [Widgets Events](05.00-Widget_Events.ipynb) | [Contents](00.00-index.ipynb) | [*OPTIONAL* Password generator: `observe`](05.02-OPTIONAL-Widget_Events_2_--_bad_password_generator,_version_1.ipynb) >
# # *OPTIONAL* Three approaches to events
# The next series of notebooks presents three ways of handling widget events. In each of the notebooks we'll construct the same simple password generator.
#
# The first uses a mix of functions and global variables, an approach like that in the notebook introducing widget events.
#
# The second separates the logic of the password generation from the password generation user interface.
#
# The third takes that separation a step further by using a class for the user interface, a class for the logic, and a class to bring them together.
# <!--NAVIGATION-->
# < [Widgets Events](05.00-Widget_Events.ipynb) | [Contents](00.00-index.ipynb) | [*OPTIONAL* Password generator: `observe`](05.02-OPTIONAL-Widget_Events_2_--_bad_password_generator,_version_1.ipynb) >
| notebooks/05.01-OPTIONAL-Widget_Events_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# language: python
# name: python3
# ---
# # Starting Working on Pandas and Data Analysis
# +
import pandas
# reading the files
data = pandas.read_csv("salaries_by_college_major.csv")
# -
data.head(5)
data.columns
data.shape
data.tail().isna()
#remove na values
cleaned_data = data.dropna()
cleaned_data.tail()
# find the row with highest starting median salary
cleaned_data["Starting Median Salary"].idxmax()
# get that entire row
cleaned_data.loc[43]
# What college major has the highest mid-career salary? How much do
# graduates with this major earn? (Mid-career is defined as having 10+ years of experience).
cleaned_data.loc[cleaned_data["Mid-Career Median Salary"].idxmax()]
# +
# Which college major has the lowest starting salary
# and how much do graduates earn after university?
cleaned_data.loc[cleaned_data["Starting Median Salary"].idxmin()]
# +
# Which college major has the lowest mid-career
# salary and how much can people expect to earn with this degree?
cleaned_data.loc[cleaned_data["Mid-Career Median Salary"].idxmin()]
# +
# How would we calculate the
# difference between the earnings of the
# 10th and 90th percentile?
#Calculate
spread_col = cleaned_data['Mid-Career 90th Percentile Salary'] - cleaned_data['Mid-Career 10th Percentile Salary']
#insert new column
cleaned_data.insert(1, 'Spread', spread_col)
#display the data
# -
cleaned_data.head()
# +
# Sort by lowest risk
low_risk = cleaned_data.sort_values('Spread')
low_risk[['Undergraduate Major', 'Spread']].head()
# +
# Using the .sort_values() method, can you find the degrees with
# the highest potential? Find the top 5 degrees
# with the highest values in the 90th percentile.
high_value =cleaned_data.sort_values("Mid-Career 90th Percentile Salary", ascending=False)
high_value.head(5)
# -
# ## Grouping and Pivoting Data with Pandas
cleaned_data.groupby("Group").count()
pandas.options.display.float_format = '{:,.2f}'.format
cleaned_data.groupby("Group").mean()
| Day_25/college_salaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
students = {}
class Ranking_System:
def __init__(self, name, surname):
self.name = name
self.surname = surname
self.quiz_score = 0
self.homework_score = 0
self.project_score = 0
self.rating = 0
self.id = ''
def set_rating(self, new_rating):
self.rating = new_rating * 1
def generate_id(self):
self.id = '12' + str(random.randint(1,1000))
def add(self):
students[self.id] = [self.name, self.surname]
print(self.name, self.surname + " is added")
def update(user_id):
pass
def menu():
while True:
item = input("Press: \n 0 to add new student \n 1 to show the list of students \n 2 and ID to print all information \n 3 and ID to update the rating \n 4 and ID to delete student ").split()
start_menu = int(item[0])
if start_menu in [2,3,4]:
id_new = int(item[1])
if item[0] == '0':
#add user
user_info = input("Input: Name, Surname, 3 RATINGs ").split(" ")
name = user_info[0]
surname = user_info[1]
quiz_score = user_info[2]
homework_score = user_info[3]
project_score = user_info[4]
rating = int(quiz_score) + int(homework_score) + int(project_score)
new_user = Ranking_System(name, surname)
new_user.set_rating(rating)
new_user.generate_id()
new_user.add()
elif item[0] == '1':
#print list of students
for key, value in sorted(students.items(), key=lambda kv: kv[1]):
print (key, ':', str(value), sep=' ')
elif item[0] == '2':
for every_stud in dir(students[key]):
for key in students:
#print information about student, how to do 2_id to input?
print ("\n Username: %s %s, \n quiz_score: %s, \n homework_score: %s, \n project_score %s, \n rating: %s" %(name, surname, quiz_score, homework_score, project_score, rating))
menu()
# -
| Raiting System Neeeeew.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from keras.layers import (LSTM, Activation, Dense, Dropout,
Embedding, GlobalMaxPool1D, Input)
from keras.models import Model
from keras.preprocessing.sequence import pad_sequences
from keras.preprocessing.text import Tokenizer
from run_binary_classifier import _load_comments, run
# -
def create_model(max_features):
inp = Input(shape=(200, ))
embed_size = 128
x = Embedding(max_features, embed_size)(inp)
x = LSTM(60, return_sequences=True,name='lstm_layer')(x)
x = GlobalMaxPool1D()(x)
x = Dropout(0.1)(x)
x = Dense(50, activation="relu")(x)
x = Dropout(0.1)(x)
x = Dense(1, activation="sigmoid")(x)
model = Model(inputs=inp, outputs=x)
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
return model
comments_X, comments_y = _load_comments('../../../data/train_binary_labels.csv')
max_features = 20000
tokenizer = Tokenizer(num_words=max_features)
tokenizer.fit_on_texts(list(comments_X))
comments_X = tokenizer.texts_to_sequences(comments_X)
comments_X = pad_sequences(comments_X, maxlen=200)
comments_X_train, comments_X_test, comments_y_train, comments_y_test = train_test_split(comments_X, comments_y, train_size=0.7, random_state=1)
model = create_model(comments_X_train.shape[0])
hist = model.fit(comments_X_train, comments_y_train, batch_size=64, epochs=1, validation_split=0.2)
# +
print('\n================= Classification report =================')
predictions = model.predict(comments_X_test, batch_size=64)
predictions[predictions > 0.5] = 1
predictions[predictions <= 0.5] = 0
print(classification_report(comments_y_test, predictions))
# -
| binary_classifiers/DemoLSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# Adding needed libraries and reading data
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import ElasticNet, Lasso, BayesianRidge, LassoLarsIC
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
from sklearn.kernel_ridge import KernelRidge
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import RobustScaler
from sklearn.base import BaseEstimator, TransformerMixin, RegressorMixin, clone
from sklearn.model_selection import KFold, cross_val_score, train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import r2_score, mean_squared_error
from sklearn.utils import shuffle
from xgboost.sklearn import XGBRegressor
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# +
train = pd.read_csv("../input/sales_train_v2.csv")
test = pd.read_csv("../input/test.csv")
items = pd.read_csv("../input/items.csv")
categories = pd.read_csv("../input/item_categories.csv")
print (train.shape, test.shape, items.shape, categories.shape)
# -
train.head()
test.head()
items.head()
categories.head()
# ## Aggregating sales by month
train.date_block_num.describe()
last_month_sales = train.loc[train.date_block_num == 33]
last_month_sales.reset_index(inplace=True)
print last_month_sales.shape
last_month_sales.head()
last_month_sales.pop('date')
last_month_sales.pop('date_block_num')
last_month_sales.pop('item_price')
last_month_sales.pop('index')
last_month_sales.head()
last_month_sales.item_cnt_day.describe()
last_month_sales = last_month_sales.groupby(['shop_id', 'item_id'], as_index=False).agg({'item_cnt_day':sum})
# +
#last_month_sales['total'] = last_month_sales.groupby(['shop_id', 'item_id'])['item_cnt_day'].transform('sum')
# -
print last_month_sales.shape
last_month_sales.head()
last_month_sales.item_cnt_day.describe()
# +
#last_month_sales.drop_duplicates(subset=['shop_id', 'item_id'], inplace=True)
# -
print last_month_sales.shape
last_month_sales.head()
test.head()
last_month_sales['total'] = last_month_sales['item_cnt_day']
last_month_sales.head()
t = last_month_sales.loc[(last_month_sales.shop_id == 45) & (last_month_sales.item_id==13881)]
t.total.values[0]
test_y = np.zeros(shape=test.shape[0])
print test_y.shape
print test.shape[0]
for i in range(test.shape[0]):
id0 = test['ID'][i]
shop_id0 = test['shop_id'][i]
item_id0 = test['item_id'][i]
#print id0, shop_id0, item_id0
if shop_id0 in last_month_sales.shop_id.values:
possibles = last_month_sales.loc[last_month_sales.shop_id == shop_id0]
if item_id0 in possibles.item_id.values:
t = possibles.loc[possibles.item_id==item_id0]
#if t.shape[0] > 0:
test_y[id0] = t.total.values[0]
#else:
# test_y[id0] = 0.0
else:
test_y[id0] = 0.0
else:
test_y[id0] = 0.0
test_id = test.ID
test_submit = pd.DataFrame({'ID': test_id, 'item_cnt_month': test_y})
test_submit.describe()
test_submit.item_cnt_month[test_submit.item_cnt_month > 20] = 20
test_submit.item_cnt_month[test_submit.item_cnt_month < 0] = 0
test_submit.describe()
print test_submit.shape
test_submit.to_csv('last_month_sales.csv', index=False)
test_submit.head()
| script/EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Rivaldop/metodologidatascience/blob/main/Pertemuan_10.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="x88FvpokX-dc"
# 1. DATASET
# + id="_JPZG7peYCzZ"
from sklearn import datasets
iris = datasets.load_iris()
x = iris.data
y = iris.target
# + id="e_1kzKF-Y4ZZ" colab={"base_uri": "https://localhost:8080/"} outputId="8ae8e267-0110-4714-db32-d1106a980dbe"
print("Banyaknya data: ",len(x))
# + id="WNgzzw0MZLcp" colab={"base_uri": "https://localhost:8080/"} outputId="fd500cf0-cf34-4a0a-88f8-39c1e91a1907"
print("15 data pertama: ")
print(x[:15])
# + [markdown] id="c0r-zSRTZZmd"
# 2. SPLIT DATA
# + id="f-S9moH1ZbP8"
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x, y, train_size = 0.7)
# + [markdown] id="Za0OGFj9aq6c"
# K-fold
# + id="HdxmFkBcatFC" colab={"base_uri": "https://localhost:8080/"} outputId="cab5a62a-bc84-405e-8432-d264966d742d"
from sklearn.model_selection import cross_val_score
from sklearn.svm import SVC
model = SVC(kernel='linear', C=1)
scores = cross_val_score(model, x, y, cv = 5)
print("Akurasi SVN untuk setiap fold: ", scores)
print("Akurasi SVN dengan 5-fold: ", scores.mean())
# + [markdown] id="ffNwJm46baX7"
# UJI COBA MODEL KLASIFIKASI
# + [markdown] id="cg5zRxsYbfBr"
# 1. KNN
# + id="V5EzricGbhBo" colab={"base_uri": "https://localhost:8080/"} outputId="c6d7fd6b-4246-40c5-b3ec-c66b52fd56db"
from sklearn.neighbors import KNeighborsClassifier
from sklearn import metrics
knn = KNeighborsClassifier()
knn.fit(x_train, y_train)
y_pred = knn.predict(x_test)
score = metrics.accuracy_score(y_test, y_pred)
print("Akurasi model KNN sebesar: ", score)
# + [markdown] id="pF4Q1zD7dWTX"
# 2. D-tree
# + id="gD2Trf2hdYBu" colab={"base_uri": "https://localhost:8080/"} outputId="cf3d9fd5-d9cf-4243-ce9c-226abfd39d22"
from sklearn.tree import DecisionTreeClassifier
from sklearn import metrics
dt = DecisionTreeClassifier(max_depth= None, min_samples_split = 2)
dt.fit(x_train, y_train)
y_pred = dt.predict(x_test)
score = metrics.accuracy_score(y_test, y_pred)
print("Akurasi model D-Tree sebesar: ", score)
# + [markdown] id="yG7O4ENxeqiH"
# 3. SVM
# + id="AzwYHz_pfF0T" colab={"base_uri": "https://localhost:8080/"} outputId="40870904-18fe-4388-a832-1b57a8778718"
from sklearn.svm import SVC
from sklearn import metrics
svm = SVC(kernel='rbf', C = 1, gamma = 0.01)
svm.fit(x_train, y_train)
y_pred = svm.predict(x_test)
score = metrics.accuracy_score(y_test, y_pred)
print("Akurasi model SVM sebesar: ", score)
# + id="4fmwkWwbgfS1" colab={"base_uri": "https://localhost:8080/"} outputId="1e225b59-ae54-4226-ac3e-c5ed61b0d6cf"
svmLinear = SVC(kernel='linear', C = 1)
svmLinear.fit(x_train, y_train)
y_pred = svmLinear.predict(x_test)
score = metrics.accuracy_score(y_test, y_pred)
print("Akurasi model SVM-Linear sebesar: ", score)
# + [markdown] id="jb-vESzrhOVy"
# 4. Logistic Regresion
# + id="ky4gEGBmhTul" colab={"base_uri": "https://localhost:8080/"} outputId="5ea8453f-f239-42c7-adb4-884b07209ea7"
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
lr = LogisticRegression()
lr.fit(x_train, y_train)
y_pred = lr.predict(x_test)
score = metrics.accuracy_score(y_test, y_pred)
print("Akurasi model LogisticRegression sebesar: ", score)
# + [markdown] id="YBfJW_pBiSP_"
# 5. Naive Bayes
# + id="rnXWG17TiaPv" colab={"base_uri": "https://localhost:8080/"} outputId="8944cf4b-4338-43c6-9c1a-d7658dc56ea3"
from sklearn.naive_bayes import BernoulliNB
from sklearn import metrics
nb = BernoulliNB()
nb.fit(x_train, y_train)
y_pred = nb.predict(x_test)
score = metrics.accuracy_score(y_test, y_pred)
print("Akurasi model Naive Bayes sebesar: ", score)
| Pertemuan_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook reproduces the computational examples in Figures 3-5 from the manuscript.
# add path to code
import sys
sys.path.insert(0, '../code')
import numpy as np
# First, we specify that we are inverting for the basal vertical velocity $w_b$:
inv_w = 1 # turn basal velocity inversion 'on'
inv_beta = 0 # turn basal drag inversion 'off'
# We are going to make some synthetic data for the example inversion.
# This is done by prescribing an oscillating Gaussian anomaly of the form
# $$ w_b^\mathrm{true}(x,y,t) = 5\exp\left(-\frac{x^2+y^2 }{2\sigma^2}\right)\sin(2\pi t\,/\,T) $$
# where $T=10$ yr is the final time and $\sigma = 20/3$ km determines the width of the anomaly.
# For later comparison, we will want this "true" inverse solution defined above, so we obtain that via:
from synthetic_data import make_fields
sol_true = make_fields(inv_w,inv_beta)
# The "true" elevation is computed by application of the forward operator $\mathcal{H}_{w_b}$:
# $$h^\mathrm{true} = \mathcal{H}_{w_b}(w_b^\mathrm{true}) $$
# and the synthetic data is constructed via
# $$h^\mathrm{obs} = h^\mathrm{true} + \text{noise}.$$
# The magnitude of the noise is set by the $\texttt{noise}\_\texttt{level}$ parameter, which determines the deviation from
# the smooth elevation by the relative "error"
# $$\|h^\mathrm{obs}-h^\mathrm{true} \|/\|h^\mathrm{true}\| = \texttt{noise}\_\texttt{level}.$$
# Here the norm over space and time is defined via
# $$\|f\|^2 = \int_0^T\int_{-\infty}^{+\infty}\int_{-\infty}^{+\infty} |f(x,y,t)|^2\;\mathrm{d}x\,\mathrm{d}y\,\mathrm{d}t,$$
# where obviously the infinite spatial domain is replaced by a "large enough" box.
# +
from synthetic_data import make_data
noise_level = 0.01 # noise level (scaled relative to elevation anomaly norm)
data = make_data(inv_w,inv_beta,noise_level) # make the synthetic data
# -
# The least-squares inverse solution is obtained by solving the normal equation
# $$ \mathcal{H}_{w_b}^\dagger(\mathcal{H}_{w_b}(w_b)) + \mathcal{R}'(w_b) = \mathcal{H}_{w_b}^\dagger (h^\mathrm{obs}) $$
# with the conjugate gradient method, where $\mathcal{R}'$ is a regularization term. An analogous equation is used for the basal drag coefficient ($\beta$) inversion. In these examples, we choose an $H^1$-type regularization of the form
# $$ \mathcal{R}'(w_b) = -\varepsilon\nabla^2 w_b$$
# where $\varepsilon$ is the regularization parameter.
# The goal now is to determine the optimal regularization parameter $\varepsilon$ that minimizes the misfit without overfitting the data.
# We are not using surface velocity data for these examples, so we set the velocity "locations" all to zero:
vel_locs = np.zeros(np.shape(data[0]),dtype=int)
# To find the optimal regularization parameter ($\varepsilon$), we will test a range of values, then
# pick the one that minimizes the misfit without overfitting the data:
eps_w = np.array([1e-2,1e-1,1e0,1e1,1e2]) # array of regularization parameters
mis_w = np.zeros(np.shape(eps_w)) # array of misfits
# The $\texttt{main}$ function returns the inverse solution $\texttt{sol}$ ($w_b$ in this case), as well as the associated forward solution $\texttt{fwd}$ ($h$ in this case), and the relative misfit $\texttt{mis}=\|h^\mathrm{obs}-h \|/\|h^\mathrm{obs}\|$.
# Convergence information is printed during the conjugate gradient iterations.
from main import main
for i in range(np.size(eps_w)):
print('------------- testing eps = '+str(eps_w[i])+' -------------')
sol,fwd,mis_w[i] = main(data,vel_locs,inv_w,inv_beta,eps_w=eps_w[i],eps_beta=0);
print('||h-h_obs||/||h_obs|| = '+str(mis_w[i])+' (target = '+str(noise_level)+') \n')
# We now determine the optimal paramter via interpolation and root finding:
from scipy.interpolate import interp1d
mis_w_int = interp1d(eps_w,mis_w,kind='linear')
# +
from scipy.optimize import root_scalar
eps_w_opt = root_scalar(lambda x: mis_w_int(x)-noise_level,x0=eps_w[0],x1=eps_w[-1]).root
# -
# We will plot the "L-curve" later, but first let's see what the optimal inverse solution looks like:
sol,fwd,mis = main(data,vel_locs,inv_w,inv_beta,eps_w=eps_w_opt,eps_beta=0);
from plotting import snapshots,plot_movie
snapshots(data,fwd,sol,sol_true,inv_w,inv_beta)
#plot_movie(data,fwd,sol,sol_true,inv_w,inv_beta) # uncomment to plot a png at every time step
# Next, we will repeat the same example for the basal drag coefficient ($\beta$) inversion. Here, he assume that a slippery spot emerges and disappeares over the observation time. The "true" field is given by
# $$ \beta^\mathrm{true}(x,y,t) = -8\times 10^{-2}\exp\left(-\frac{x^2+y^2 }{2\sigma^2}\right)B(t) $$
# where $B$ is a continuous box-type function that controls the emergence and disappearance of the anomaly (see synthetic_data.py).
# Omitting the same level of detail as above, we repeat the test for this input below:
# +
inv_w = 0 # turn basal velocity inversion 'off'
inv_beta = 1 # turn basal drag inversion 'on'
sol_true = make_fields(inv_w,inv_beta) # get the "true" inverse solution
data = make_data(inv_w,inv_beta,noise_level) # create the data
eps_b = np.array([1e2,1e3,1e4,1e5,1e6]) # array of regularization parameters
mis_b = np.zeros(np.shape(eps_b)) # array of misfits
for i in range(np.size(eps_b)):
print('------------- testing eps = '+str(eps_b[i])+' -------------')
sol,fwd,mis_b[i] = main(data,vel_locs,inv_w,inv_beta,eps_beta=eps_b[i],eps_w=0);
print('||h-h_obs||/||h_obs|| = '+str(mis_b[i])+' (target = '+str(noise_level)+') \n')
mis_b_int = interp1d(eps_b,mis_b,kind='linear') # interpolate misfits and find the optimal reg. parameter
eps_b_opt = root_scalar(lambda x: mis_b_int(x)-noise_level,x0=eps_b[0],x1=eps_b[-1]).root
print('--------------------------------------------------------------------')
print('Getting inverse solution at optimal regularization parameter value\n')
sol,fwd,mis = main(data,vel_locs,inv_w,inv_beta,eps_beta=eps_b_opt,eps_w=0);
snapshots(data,fwd,sol,sol_true,inv_w,inv_beta)
#plot_movie(data,fwd,sol,sol_true,inv_w,inv_beta) # uncomment to plot a png at every time step
# -
# Clearly the reconstructed basal drag field has a smaller amplitude than the "true" solution. In the next notebooks, we show how incorporation of velocity data can remedy this issue.
# Finaly, we can plot the "L-curve" for both inversion examples:
import matplotlib.pyplot as plt
plt.figure(figsize=(8,4))
plt.axhline(y=noise_level,color='k',linestyle='--',linewidth=2)
plt.plot(eps_w,mis_w,'o-',color='C3',linewidth=2,markersize=8,mec='k',label=r'$w_b$')
plt.plot([eps_w_opt],[mis_w_int(eps_w_opt)],'*',color='C3',markersize=20,mec='k')
plt.plot(eps_b,mis_b,'^-',color='C0',linewidth=2,markersize=8,mec='k',label=r'$\beta$')
plt.plot([eps_b_opt],[mis_b_int(eps_b_opt)],'*',color='C0',markersize=20,mec='k')
plt.annotate(r'noise level',xy=(3e-1,1.1e-2),fontsize=18,color='k')
plt.gca().set_yscale('log')
plt.gca().set_xscale('log')
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.gca().invert_xaxis()
plt.xlabel(r'$\varepsilon$',fontsize=20)
plt.ylabel(r'$\Vert h^\mathrm{obs}-h^\varepsilon \Vert\,/\,\Vert h^\mathrm{obs}\Vert$',fontsize=20)
plt.legend(fontsize=18,loc='upper right')
plt.tight_layout()
plt.savefig('fig3',bbox_inches='tight')
plt.show()
plt.close()
| notebooks/1_Figs3-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="2d74572580e24b09f3bbd0ec9d74bde002234be0"
# Existing model is forked From [https://github.com/ashishpatel26/Facial-Expression-Recognization-using-JAFFE](http://) . But I have removed the labeling error and changed the CNN Model . I have also updated the existing model that gives better accuracy than previous.
# + id="ODyVs_urTCSH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 94} outputId="3954b8fb-c4b7-4f38-a59f-8c3be26b6217" _uuid="be0ca550e4cd455834b3177a5ee43c33801cfe11"
import os,cv2
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from pylab import rcParams
rcParams['figure.figsize'] = 20, 10
from sklearn.utils import shuffle
from sklearn.cross_validation import train_test_split
import keras
from keras.utils import np_utils
from keras import backend as K
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation, Flatten
from keras.layers.convolutional import Convolution2D, MaxPooling2D
from keras.optimizers import SGD,RMSprop,adam
from keras.preprocessing.image import ImageDataGenerator
# + id="uQI820JhUYj9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="0c96a197-6b10-454a-bc23-c23b3fbf6056" _uuid="b2ccc8d12aa2c567760421bd87d536144f00ae90"
data_path = '../input/jaffefacialexpression/jaffe/jaffe'
data_dir_list = os.listdir(data_path)
img_rows=256
img_cols=256
num_channel=1
num_epoch=10
img_data_list=[]
for dataset in data_dir_list:
img_list=os.listdir(data_path+'/'+ dataset)
print ('Loaded the images of dataset-'+'{}\n'.format(dataset))
for img in img_list:
input_img=cv2.imread(data_path + '/'+ dataset + '/'+ img )
#input_img=cv2.cvtColor(input_img, cv2.COLOR_BGR2GRAY)
input_img_resize=cv2.resize(input_img,(128,128))
img_data_list.append(input_img_resize)
img_data = np.array(img_data_list)
img_data = img_data.astype('float32')
img_data = img_data/255
img_data.shape
# + id="hbOl7E-8UcX9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="00424152-50c9-422f-ed15-117c33ebe6dc" _uuid="1079555608d0b0036a349af00f76481caa79eebd"
num_classes = 7
num_of_samples = img_data.shape[0]
labels = np.ones((num_of_samples,),dtype='int64')
labels[0:29]=0 #30
labels[30:58]=1 #29
labels[59:90]=2 #32
labels[91:121]=3 #31
labels[122:151]=4 #30
labels[152:182]=5 #31
labels[183:]=6 #30
names = ['ANGRY','DISGUST','FEAR','HAPPY','NEUTRAL','SAD','SURPRISE']
def getLabel(id):
return ['ANGRY','DISGUST','FEAR','HAPPY','NEUTRAL','SAD','SURPRISE'][id]
# + id="xUmcqKiMUgZk" colab_type="code" colab={} _uuid="29f2c1a4c161db04c594667fc55023a6e5da0be8"
Y = np_utils.to_categorical(labels, num_classes)
#Shuffle the dataset
x,y = shuffle(img_data,Y, random_state=2)
# Split the dataset
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.15, random_state=2)
x_test=X_test
#X_train=X_train.reshape(X_train.shape[0],128,128,1)
#X_test=X_test.reshape(X_test.shape[0],128,128,1)
# + _uuid="89517c9ec41b09ed60e4d005a08fb7fe78ac2eb0"
x_test.shape
# + id="UEcO-PMyUkhO" colab_type="code" colab={} _uuid="4425acad3bc8aa0019f50e0b29c7ce4e1eeb7d57"
from keras.models import Sequential
from keras.layers import Dense , Activation , Dropout ,Flatten
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
from keras.metrics import categorical_accuracy
from keras.models import model_from_json
from keras.callbacks import ModelCheckpoint
from keras.optimizers import *
from keras.layers.normalization import BatchNormalization
# + id="YpCL9im7VHpC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 150} outputId="1ad4d180-1c8b-43c9-9399-a25e79194520" _uuid="b59b01ab135eca80576a5915c4a1fae0721328f2"
input_shape=(128,128,3)
model = Sequential()
model.add(Conv2D(6, (5, 5), input_shape=input_shape, padding='same', activation = 'relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(16, (5, 5), padding='same', activation = 'relu'))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3), activation = 'relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(128, activation = 'relu'))
model.add(Dropout(0.5))
model.add(Dense(7, activation = 'softmax'))
# Classification
# model.add(Flatten())
# model.add(Dense(64))
# model.add(Activation('relu'))
# model.add(Dropout(0.5))
# model.add(Dense(num_classes))
# model.add(Activation('softmax'))
#Compile Model
model.compile(loss='categorical_crossentropy', optimizer='adam',metrics=["accuracy"])
# + id="Gjmj9Cz3VT1b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 734} outputId="2ffb75fb-f6eb-4f49-8afe-68e86641b5f4" _uuid="888f7ccffbe9b2a0e7f5eea9886509c4c2da53b7"
model.summary()
model.get_config()
model.layers[0].get_config()
model.layers[0].input_shape
model.layers[0].output_shape
model.layers[0].get_weights()
np.shape(model.layers[0].get_weights()[0])
model.layers[0].trainable
# + id="8CNczAIPVhDy" colab_type="code" colab={} _uuid="36548e567c32aa7fe143611962071bb306b0e0fb"
from keras import callbacks
filename='model_train_new.csv'
filepath="Best-weights-my_model-{epoch:03d}-{loss:.4f}-{acc:.4f}.hdf5"
csv_log=callbacks.CSVLogger(filename, separator=',', append=False)
checkpoint = callbacks.ModelCheckpoint(filepath, monitor='val_loss', verbose=1, save_best_only=True, mode='min')
callbacks_list = [csv_log,checkpoint]
callbacks_list = [csv_log]
# + id="7s8Uy0G8WnTa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1980} outputId="a5ce06fa-576f-47b0-b6d8-da606e32fd7f" _uuid="b44a3fe424841532a630635f4702c1f980f2b3ac"
hist = model.fit(X_train, y_train, batch_size=7, epochs=50, verbose=1, validation_data=(X_test, y_test),callbacks=callbacks_list)
# + id="tYAXlAm7WzSw" colab_type="code" colab={} _uuid="5dca58ec635d18a8baf11fc70ec826da6e9f3c1e"
#Model Save
model.save_weights('model_weights.h5')
model.save('model_keras.h5')
# + id="Enxigu7LYX5e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 581} outputId="05f9582a-efd7-41ee-a2ae-0443a9b13440" _uuid="7e0d02f0f04af3345997c0236194c2ff7937c1dc"
# visualizing losses and accuracy
# %matplotlib inline
train_loss=hist.history['loss']
val_loss=hist.history['val_loss']
train_acc=hist.history['acc']
val_acc=hist.history['val_acc']
epochs = range(len(train_acc))
plt.plot(epochs,train_loss,'r', label='train_loss')
plt.plot(epochs,val_loss,'b', label='val_loss')
plt.title('train_loss vs val_loss')
plt.legend()
plt.figure()
plt.plot(epochs,train_acc,'r', label='train_acc')
plt.plot(epochs,val_acc,'b', label='val_acc')
plt.title('train_acc vs val_acc')
plt.legend()
plt.figure()
# + id="nvqxVr7qYbxX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 707} outputId="3554ef0e-d6b6-4956-ef8a-6ff78f88246c" _uuid="c5de70c87dc739e401ddc27262eea4433fa34d9f"
# Evaluating the model
score = model.evaluate(X_test, y_test, verbose=0)
print('Test Loss:', score[0])
print('Test accuracy:', score[1])
test_image = X_test[0:1]
print (test_image.shape)
print(model.predict(test_image))
print(model.predict_classes(test_image))
print(y_test[0:1])
res = model.predict_classes(X_test[9:18])
plt.figure(figsize=(10, 10))
for i in range(0, 9):
plt.subplot(330 + 1 + i)
plt.imshow(x_test[i],cmap=plt.get_cmap('gray'))
plt.gca().get_xaxis().set_ticks([])
plt.gca().get_yaxis().set_ticks([])
plt.ylabel('prediction = %s' % getLabel(res[i]), fontsize=14)
# show the plot
plt.show()
# + id="1Y1ASlPjYgZY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="fd0767e8-5066-4bfc-dc84-4b758dfc2157" _uuid="dac2263fcba5f2d1bbd0d3eceb3df32843f51f93"
from sklearn.metrics import confusion_matrix
results = model.predict_classes(X_test)
cm = confusion_matrix(np.where(y_test == 1)[1], results)
plt.matshow(cm)
plt.title('Confusion Matrix')
plt.colorbar()
plt.ylabel('True Label')
plt.xlabel('Predicted Label')
plt.show()
# + id="L-3b02owi6b5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 333} outputId="79fb1279-f850-400c-84ce-c03dd2988a65" _uuid="864e957558ded986a03307bbbb129b9bf26623c3"
testimg_data_list=[]
test_img=cv2.imread('../input/facial-expression/facial/facial/Shawon.jpg',True)
test_img_resize=cv2.resize(test_img,(128,128))
testimg_data_list.append(test_img_resize)
testimg_data = np.array(testimg_data_list)
testimg_data = testimg_data.astype('float32')
testimg_data = testimg_data/255
testimg_data.shape
print("test image original shape",testimg_data[0].shape)
print("image original shape",img_data[0].shape)
results = model.predict_classes(testimg_data)
plt.imshow(test_img,cmap=plt.get_cmap('Set2'))
plt.gca().get_xaxis().set_ticks([])
plt.gca().get_yaxis().set_ticks([])
plt.xlabel('prediction = %s' % getLabel(results[0]), fontsize=25)
# + id="w0Pdii74jBVn" colab_type="code" colab={} _uuid="aa6a25862ae18ba49caddc6e11a8bca667ccb249"
| jaffe-with-resnet50.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# author: <NAME>, Ed.D., February 2019
# # *Pulling and Processing PDFs *
#
# This is a KEY piece of code. This code
# 1. pulls a .pdf from a directory
# 1. rejects the .pdf if it has "._"_ in the title
# 1. pulls the text from the .pdf
# 1. cleans the text
# 1. tokenizes the text by sentence
# 1. searches each sentence for our designated list of keywords
# 1. saves the sentences that contain one of our keywords in a dataframe
# - saves the name of the file for reference
# - saves the keyword in the same row for reference
# - saves the sentence in the same row
#
#
# # Load Libraries
#
# `__nlp_init_nsv__.py` is a local library that I produced. It is necessary to run this program. It will take a moment to run as it is rather long. It is loading a lot of things.
# %run lib/__nlp_init_nsv__.py
# # The Real Work of the Program
# +
# Pulling only those files with a .pdf extention
import os
df = pd.DataFrame(columns=['1'])
i = 0
# directory is where you point Python to where the .pdfs are stored in a blob
directory = 'test_pdfs/'
#keywords are those words or parts of words for which we are searching
keywords = ['correlat',
'male',
'race',
'ethn',
'will be used',
'female',
'youth',
'weapon',
'adolescent',
'gun',
'knife',
' caus',
]
for root, dirs, files in os.walk(directory):
for file in files:
if file.endswith(".pdf"):
# print(os.path.join(root, file)) ## This line is only used to check our work
if '._' in file:
# This line tells us which files do not make the criteria for being processed
# This printed line is NOT saved.
print('\n\n',(os.path.join(root, file)),'ignored.', '\n\n')
pass
else:
#if there is a unicodedecode error because of the .pdf, this will skip the bad file.
try:
s = str(directory+file)
# This line tells us which files are being processed. It is not saved.
print('\n\n',s, 'being processed.','\n\n')
# import the text from the .pdfs, convert it to a string, and clean it
text = import_pdf(s)
text = str(text)
text = clean_the_text(text) #uses my internal library from __nlp_init_nsv__.py
#Where the real work is done
for term in keywords:
for sentence in sent_tokenize(text):
if term in sentence:
sentence = clean_the_text(sentence)
print(term, '\n', sentence)
df.loc[i, 'Article'] = s
df.loc[i, 'Keyword'] = term
df.loc[i, 'Sentence'] = sentence
i += 1
except UnicodeDecodeError:
print('\n\n',(os.path.join(root, file)),'ignored.', '\n\n')
continue
df
# -
# # Save to File
#
# I included a copy of the output to the file just to give you an idea of what it should come out looking like.
# +
# Save the file
df.to_csv('KeywordExtract.csv')
# -
| Pulling and Processing PDFs .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import Perceptron
import pandas
import numpy
import matplotlib as plt
from sklearn.pipeline import make_pipeline
from sklearn import linear_model, feature_extraction
def categorical_features(row):
d = {}
d["STATE"] = row[1]["STATE"]
return d
def last_poll(full_data):
"""
Create feature from last poll in each state
"""
# Only care about republicans
repub = full_data[full_data["PARTY"] == "Rep"]
# Sort by date
chron = repub.sort_values(by="DATE", ascending=True)
# Only keep the last one
dedupe = chron.drop_duplicates(subset="STATE", keep="last")
# Remove national polls
return dedupe[dedupe["STATE"] != "US"]
if __name__ == "__main__":
# Read in the X data
all_data = pandas.read_csv("data.csv")
# Remove non-states
all_data = all_data[pandas.notnull(all_data["STATE"])]
# split between testing and training
train_x = last_poll(all_data[all_data["TOPIC"] == '2012-president'])
train_x.set_index("STATE")
test_x = last_poll(all_data[all_data["TOPIC"] == '2016-president'])
test_x.set_index("STATE")
# Read in the Y data
y_data = pandas.read_csv("../data/2012_pres.csv", sep=';')
y_data = y_data[y_data["PARTY"] == "R"]
y_data = y_data[pandas.notnull(y_data["GENERAL %"])]
y_data["GENERAL %"] = [float(x.replace(",", ".").replace("%", ""))
for x in y_data["GENERAL %"]]
y_data["STATE"] = y_data["STATE ABBREVIATION"]
y_data.set_index("STATE")
backup = train_x
train_x = y_data.merge(train_x, on="STATE",how='left')
# make sure we have all states in the test data
for ii in set(y_data.STATE) - set(test_x.STATE):
new_row = pandas.DataFrame([{"STATE": ii}])
test_x = test_x.append(new_row)
# format the data for regression
train_x = pandas.concat([train_x.STATE.astype(str).str.get_dummies(),
train_x], axis=1)
test_x = pandas.concat([test_x.STATE.astype(str).str.get_dummies(),
test_x], axis=1)
# handle missing data
for dd in train_x, test_x:
dd["NOPOLL"] = pandas.isnull(dd["VALUE"])
dd["VALUE"] = dd["VALUE"].fillna(0.0)
dd["NOMOE"] = pandas.isnull(dd["MOE"])
dd["MOE"] = dd["MOE"].fillna(0.0)
# create feature list
features = list(y_data.STATE)
features.append("VALUE")
features.append("NOPOLL")
features.append("MOE")
features.append("NOMOE")
features_rep = list(y_data.PARTY)# from y data
features_rep = [ord(i) for i in features_rep]# changing R into ascii value
features_obs = list(all_data.OBS)#from csv all_data since its not in y data
features_val = list(all_data.VALUE)# adding value as features
features_moe = list(all_data.MOE)
feature_matrix=[] #creating empty matrix
for i in range(len(features_rep)):
feature_matrix.append([features_rep[i],features_obs[i],features_val[i],features_moe[i]])# appending to list R and OBS
features_matrix=numpy.array(feature_matrix)#returning an array with value of R and OBs
feature_matrix1=[] #creating empty matrix
for i in range(len(features_rep)):
feature_matrix1.append([features_rep[i],features_obs[i],features_val[i],features_moe[i]])# appending to list R and OBS
features_matrix1=numpy.array(feature_matrix1)#returning an array with value of R and OBs
# fit the regression
#mod = linear_model.LinearRegression()
#mod = linear_model.Ridge(alpha=.667)
#mod = linear_model.BayesianRidge()
#mod.fit(train_x[features], train_x["GENERAL %"])
#pipelining
mod=make_pipeline(PolynomialFeatures(degree=6),linear_model.LinearRegression())# making a model for polynomial regression
mod.fit(feature_matrix, train_x["GENERAL %"])# passing matris with feature
#modd=make_pipeline(PolynomialFeatures(degree=4),linear_model.BayesianRidge())# making a model for polynomial regression
#modd.fit(feature_matrix1, train_x["GENERAL %"])# passing matris with feature
modd=make_pipeline(PolynomialFeatures(degree=4),linear_model.Lasso())# making a model for polynomial regression
modd.fit(feature_matrix1, train_x["GENERAL %"])# passing matris with feature
# Write out the model
#with open("model.txt", 'w') as out:
#out.write("BIAS\t%f\n" % mod.intercept_)
#for jj, kk in zip(features, mod.coef_):
#out.write("%s\t%f\n" % (jj, kk))
# Write the predictions
pred_test = mod.predict(features_matrix)
with open("pred.txt", 'w') as out:
for ss, vv in sorted(zip(list(test_x.STATE), pred_test)):
out.write("%s\t%f\n" % (ss, vv))
# Write the predictions
pred_test = modd.predict(feature_matrix1)
with open("pred.txt", 'w') as out:
for ss, vv in sorted(zip(list(test_x.STATE), pred_test)):
out.write("%s\t%f\n" % (ss, vv))
print("Mean squared error: %.2f"
% numpy.mean((mod.predict(features_matrix) - train_x["GENERAL %"]) ** 2))
#test data - train data
#(prediction - observed)^2
# Explained variance score: 1 is perfect prediction
print('Variance score: %.2f' % mod.score(features_matrix,train_x["GENERAL %"]))
print("Mean squared error2: %.2f"
% numpy.mean((modd.predict(features_matrix1) - train_x["GENERAL %"]) ** 2))
#test data - train data
#(prediction - observed)^2
# Explained variance score: 1 is perfect prediction
print('Variance score2: %.2f' % modd.score(features_matrix1,train_x["GENERAL %"]))
# -
| regression/new code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
from your_lib.core import *
# # Parameter Identifiability
#
# This project contains the code and the results of the experiment in progress to evaluate the identifiability of the distributed parameters in a Watershed.
# ## Install
# `git clone https://github.com/nicolas998/Parameter_identifiability.git`
#
# `pip3 setup.py install --user`
# ## Questions
# Following a syncronic cascade, which is the required level of complexity to explain the hydrograps at different scales.
#
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_ner1upo"
# # Counting Inversions
#
# The number of *inversions* in a disordered list is the number of pairs of elements that are inverted (out of order) in the list.
#
# Here are some examples:
# - [0,1] has 0 inversions
# - [2,1] has 1 inversion (2,1)
# - [3, 1, 2, 4] has 2 inversions (3, 2), (3, 1)
# - [7, 5, 3, 1] has 6 inversions (7, 5), (3, 1), (5, 1), (7, 1), (5, 3), (7, 3)
#
# The number of inversions can also be thought of in the following manner.
#
# >Given an array `arr[0 ... n-1]` of `n` distinct positive integers, for indices `i and j`, if `i < j` and `arr[i] > arr[j]` then the pair `(i, j)` is called an inversion of `arr`.
#
# ## Problem statement
#
# Write a function, `count_inversions`, that takes an array (or Python list) as input, and returns a count of the total number of inversions present in the input.
#
# Mergesort provides an efficient way to solve this problem.
# + graffitiCellId="id_jf6a5ys"
def count_inversions(arr):
start_index = 0
end_index = len(arr) - 1
output = inversion_count_func(arr, start_index, end_index)
return output
def inversion_count_func(arr, start_index, end_index):
if start_index >= end_index:
return 0
mid_index = start_index + (end_index - start_index) // 2
# find number of inversions in left-half
left_answer = inversion_count_func(arr, start_index, mid_index)
# find number of inversions in right-half
right_answer = inversion_count_func(arr, mid_index + 1, end_index)
output = left_answer + right_answer
# merge two sorted halves and count inversions while merging
output += merge_two_sorted_halves(arr, start_index, mid_index, mid_index + 1, end_index)
return output
def merge_two_sorted_halves(arr, start_one, end_one, start_two, end_two):
count = 0
left_index = start_one
right_index = start_two
output_length = (end_two - start_two + 1) + (end_one - start_one + 1)
output_list = [0 for _ in range(output_length)]
index = 0
while index < output_length:
# if left <= right, it's not an inversion
if arr[left_index] <= arr[right_index]:
output_list[index] = arr[left_index]
left_index += 1
else:
count = count + (end_one - left_index + 1) # left > right hence it's an inversion
output_list[index] = arr[right_index]
right_index += 1
index = index + 1
if left_index > end_one:
for i in range(right_index, end_two + 1):
output_list[index] = arr[i]
index += 1
break
elif right_index > end_two:
for i in range(left_index, end_one + 1):
output_list[index] = arr[i]
index += 1
break
index = start_one
for i in range(output_length):
arr[index] = output_list[i]
index += 1
return count
# + [markdown] graffitiCellId="id_8809fp2"
# <span class="graffiti-highlight graffiti-id_8809fp2-id_8br31oi"><i></i><button>Hide Solution</button></span>
# + graffitiCellId="id_8br31oi"
def count_inversions(arr):
start_index = 0
end_index = len(arr) - 1
output = inversion_count_func(arr, start_index, end_index)
return output
def inversion_count_func(arr, start_index, end_index):
if start_index >= end_index:
return 0
mid_index = start_index + (end_index - start_index) // 2
# find number of inversions in left-half
left_answer = inversion_count_func(arr, start_index, mid_index)
# find number of inversions in right-half
right_answer = inversion_count_func(arr, mid_index + 1, end_index)
output = left_answer + right_answer
# merge two sorted halves and count inversions while merging
output += merge_two_sorted_halves(arr, start_index, mid_index, mid_index + 1, end_index)
return output
def merge_two_sorted_halves(arr, start_one, end_one, start_two, end_two):
count = 0
left_index = start_one
right_index = start_two
output_length = (end_two - start_two + 1) + (end_one - start_one + 1)
output_list = [0 for _ in range(output_length)]
index = 0
while index < output_length:
# if left <= right, it's not an inversion
if arr[left_index] <= arr[right_index]:
output_list[index] = arr[left_index]
left_index += 1
else:
count = count + (end_one - left_index + 1) # left > right hence it's an inversion
output_list[index] = arr[right_index]
right_index += 1
index = index + 1
if left_index > end_one:
for i in range(right_index, end_two + 1):
output_list[index] = arr[i]
index += 1
break
elif right_index > end_two:
for i in range(left_index, end_one + 1):
output_list[index] = arr[i]
index += 1
break
index = start_one
for i in range(output_length):
arr[index] = output_list[i]
index += 1
return count
# + graffitiCellId="id_l6xh0rg"
def test_function(test_case):
arr = test_case[0]
solution = test_case[1]
if count_inversions(arr) == solution:
print("Pass")
else:
print("Fail")
# + graffitiCellId="id_4aqr272"
arr = [2, 5, 1, 3, 4]
solution = 4
test_case = [arr, solution]
test_function(test_case)
# + graffitiCellId="id_0sqy9z2"
arr = [54, 99, 49, 22, 37, 18, 22, 90, 86, 33]
solution = 26
test_case = [arr, solution]
test_function(test_case)
# + graffitiCellId="id_s5l67ma"
arr = [1, 2, 4, 2, 3, 11, 22, 99, 108, 389]
solution = 2
test_case = [arr, solution]
test_function(test_case)
# + graffitiCellId="id_yueoudx"
| concepts/Basic Algorithms/09 Counting Inversions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
# %matplotlib inline
from matplotlib import pyplot as plt
import seaborn as sns
os.chdir('D:/Practical Time Series/')
zero_mean_series = np.random.normal(loc=0.0, scale=1., size=100)
plt.figure(figsize=(5.5, 5.5))
g = sns.tsplot(zero_mean_series)
g.set_title('Zero mean model')
g.set_xlabel('Time index')
plt.savefig('plots/ch1/B07887_01_12.png', format='png', dpi=300)
random_walk = np.cumsum(zero_mean_series)
plt.figure(figsize=(5.5, 5.5))
g = sns.tsplot(random_walk)
g.set_title('Random Walk')
g.set_xlabel('Time index')
plt.savefig('plots/ch1/B07887_01_13.png', format='png', dpi=300)
| Chapter01/Chapter_1_Models_for_Time_Series_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="TWSHyGLvFHfl"
# ## Low level PyTorch interface
# + colab={} colab_type="code" id="_QstD-ySFHfo"
import numpy as np # results verification during grading
import math # Pi
import matplotlib.pyplot as plt # 2D plot in task 1
# %matplotlib inline
import torch
assert torch.__version__ >= '1.3.0'
# + colab={} colab_type="code" id="DBMRmAyOFHf-"
from notmnist import load_notmnist
# + [markdown] colab_type="text" id="763SfU_dFHgF"
# While learning PyTorch, you will have lots of questions, e.g.
#
# * how to choose between `.sqrt()` and `.sqrt_()`,
# * when to use `.view()` and how is it different from `.reshape()`,
# * which `dtype` to use etc.
#
# To find the answers, you are expected to study [documentation](https://pytorch.org/docs/stable/index.html) and google a lot.
#
# If this is your first PyTorch experience, you may want to [start here](https://pytorch.org/tutorials/).
#
# Quick documentation on functions and modules is available with `?` and `help()`, like so:
# + [markdown] colab_type="text" id="lhgyOraEFHg9"
# ## Task
#
# You have to solve yet another character recognition problem: *notMNIST* dataset of 10 letters and ~14 000 train samples.
#
# For this, we ask you to build a multilayer perceptron (*i.e. a neural network of linear layers*) from scratch using **low-level** PyTorch interface.
#
# Requirements:
# 1. at least 82% validation accuracy,
# 2. at least 2 linear layers,
# 3. no convolutions,
# 3. use [softmax followed by categorical cross-entropy](https://gombru.github.io/2018/05/23/cross_entropy_loss/).
#
# Tips:
#
# * Don't use the structures from homework 1 (`Sequential`, `.forward()` etc.). I suggest that your `NeuralNet.predict()` consists of 5-7 lines.
# * Pick random batches (either shuffle data before each epoch or sample each batch randomly).
# * Do not initialize weights with zeros ([learn why](https://stats.stackexchange.com/questions/27112/danger-of-setting-all-initial-weights-to-zero-in-backpropagation)). Gaussian noise with small variance will do.
# * 50 hidden neurons and a sigmoid nonlinearity will do for a start. Many ways to improve.
# * To improve accuracy, consider changing layers' sizes, nonlinearities, optimization methods, weights initialization.
# * Don't use GPU yet.
#
# Happy googling!
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="IBQ8Zz9rFHg9" outputId="7fd30075-ee9e-42eb-f576-e9b7edf6c83f"
letters = 'ABCDEFGHIJ'
X_train, y_train, X_val, y_val = map(torch.tensor, load_notmnist(letters=letters))
X_train.squeeze_()
X_val.squeeze_();
# + colab={"base_uri": "https://localhost:8080/", "height": 154} colab_type="code" id="UGgRCZtDFHhD" outputId="7107b6f9-7ebf-43af-d7b6-83f3efa127d8"
fig, axarr = plt.subplots(2, 10, figsize=(10, 2))
for idx, ax in enumerate(axarr.ravel()):
ax.imshow(X_train[idx], cmap='gray')
ax.axis('off')
ax.set_title(letters[y_train[idx]])
fig.tight_layout()
# + colab={} colab_type="code" id="7yUp8x9MFHhH"
np.random.seed(666)
torch.manual_seed(666)
# + [markdown] colab_type="text" id="QosI2zTDFHhK"
# The cell below has an example layout for encapsulating your neural network. Feel free to modify the interface if you need to (add arguments, add return values, add methods etc.). For example, you may want to add a method `do_gradient_step()` that executes one optimization algorithm (SGD / Adadelta / Adam / ...) step.
# + colab={} colab_type="code" id="Xo9_lwWvFHhL"
class NeuralNet:
EPS = 0.01
LR = 0.1
def __init__(self):
self.linear1 = torch.randn((28*28, 28*28//2)) * self.EPS
self.bias1 = torch.randn(28*28//2) * self.EPS
self.linear2 = torch.randn((28*28//2, 10)) * self.EPS
self.bias2 = torch.randn(10) * self.EPS
self.linear1.requires_grad_()
self.bias1.requires_grad_()
self.linear2.requires_grad_()
self.bias2.requires_grad_()
def predict(self, images):
"""
images:
`torch.tensor`, shape == `batch_size x height x width`, dtype == `torch.float32`
A minibatch of images -- the input to the neural net.
return:
prediction:
`torch.tensor`, shape == `batch_size x 10`, dtype == `torch.float32`
The probabilities of each input image to belong to each of the dataset classes.
Namely, `output[i, j]` is the probability of `i`-th
minibatch sample to belong to `j`-th class.
"""
batch_size = images.shape[0]
out1 = images.reshape(batch_size, -1)
out2 = out1 @ self.linear1 + self.bias1
out3 = out2.sigmoid()
out4 = out3 @ self.linear2 + self.bias2
out5 = out4.softmax(dim=1)
return out5
def do_gradient_step(self):
self.linear1.data = self.linear1.data - self.LR * self.linear1.grad.detach()
self.bias1.data = self.bias1.data - self.LR * self.bias1.grad.detach()
self.linear2.data = self.linear2.data - self.LR * self.linear2.grad.detach()
self.bias2.data = self.bias2.data - self.LR * self.bias2.grad.detach()
self.linear1.grad.zero_()
self.bias1.grad.zero_()
self.linear2.grad.zero_()
self.bias2.grad.zero_()
# + colab={} colab_type="code" id="A49vVGCOFHhP"
def accuracy(model, images, labels):
"""
model:
`NeuralNet`
images:
`torch.tensor`, shape == `N x height x width`, dtype == `torch.float32`
labels:
`torch.tensor`, shape == `N`, dtype == `torch.int64`
Class indices for each sample in `images`.
return:
value:
`float`
The fraction of samples from `images` correctly classified by `model`.
`0 <= value <= 1`.
"""
with torch.no_grad():
probs = model.predict(images)
preds = torch.argmax(probs, dim=1)
return torch.sum(preds == labels).item() / labels.shape[0]
# + colab={} colab_type="code" id="dwSr04TTFHhT"
model = NeuralNet()
batch_size = 128
epochs = 6
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="6_JWl8BoFHhY" outputId="6e8f6d92-af5a-4f94-f887-fb2e7e80546a"
# %%time
torch.set_num_threads(1)
for k in range(epochs):
print('{} epoch has started'.format(k + 1))
idxs = torch.randperm(X_train.shape[0])
X_train = X_train[idxs]
y_train = y_train[idxs]
for i in range(0, X_train.shape[0], batch_size):
X_train_batch, y_train_batch = X_train[i: i + batch_size], y_train[i: i + batch_size]
preds = model.predict(X_train_batch)
loss = -torch.mean(torch.log(torch.gather(preds, 1, y_train_batch.unsqueeze(1)).squeeze()))
loss.backward()
model.do_gradient_step()
# +
train_accuracy = accuracy(model, X_train, y_train) * 100
val_accuracy = accuracy(model, X_val, y_val) * 100
print("Training accuracy: %.2f, validation accuracy: %.2f" % (train_accuracy, val_accuracy))
assert val_accuracy >= 82.0
print("Well done!")
# + colab={} colab_type="code" id="74RxN0umvFIG"
| lecture_8/cnn_in_torch/low_level_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/zronyj/matematica/blob/master/FactorizacionDiagonalizacion.ipynb)
# + [markdown] id="yynf_kq3VWk_" colab_type="text"
# # ¡Te volvemos a dar la bienvenida a un cuaderno interactivo de Colaboratory!
#
# En esta ocasión vamos a ir directamente al grano y comenzar a trabajar con matrices de Hückel. Según la práctica de laboratorio, una matriz de Hückel se construye para moléculas con enlaces $\pi$ en resonancia. Para ello vamos a comenzar con un ejemplo sencillo: una molécula de **benceno**.
#
# La matriz para el benceno va a tener 6 columnas y 6 filas, pues son solo 6 los átomos que cuentan con enlaces $\pi$ en resonancia. Siguiendo con lo propuesto, la diagonal principal se debe llenar con valores $\alpha$ que representan la energía de ionización del orbital **p**, y luego se deben de colocar $\beta$ en aquella combinación de fila-columna en donde se encuentre un enlace.
#
# Por ejemplo, la fila 1, columna 2 representa al átomo de carbono 1 y el átomo de carbono 2 en la molécula de benceno. De estos podemos esperar que haya un enlace, por lo que colocamos un $\beta$ en esa entrada de la matriz. Siguiendo así terminaríamos con la siguiente matriz:
#
# $$\mathbf{H} = \left [ \begin{matrix}
# \alpha & \beta & 0 & 0 & 0 & \beta\\
# \beta & \alpha & \beta & 0 & 0 & 0\\
# 0 & \beta & \alpha & \beta & 0 & 0\\
# 0 & 0 & \beta & \alpha & \beta & 0\\
# 0 & 0 & 0 & \beta & \alpha & \beta\\
# \beta & 0 & 0 & 0 & \beta & \alpha
# \end{matrix} \right ]$$
#
# + [markdown] id="tSC10Btfb7qC" colab_type="text"
# Sin embargo, al colocar esto dentro de la ecuación de valores propios, resulta que se puede manipular más este problema para facilitar su resolución.
#
# $$\left [ \begin{matrix}
# \alpha & \beta & 0 & 0 & 0 & \beta\\
# \beta & \alpha & \beta & 0 & 0 & 0\\
# 0 & \beta & \alpha & \beta & 0 & 0\\
# 0 & 0 & \beta & \alpha & \beta & 0\\
# 0 & 0 & 0 & \beta & \alpha & \beta\\
# \beta & 0 & 0 & 0 & \beta & \alpha
# \end{matrix} \right ] \cdot
# \left [ \begin{matrix}
# c_1\\
# c_2\\
# c_3\\
# c_4\\
# c_5\\
# c_6
# \end{matrix} \right ] =
# \left [ \begin{matrix}
# E_1 & 0 & 0 & 0 & 0 & 0\\
# 0 & E_2 & 0 & 0 & 0 & 0\\
# 0 & 0 & E_3 & 0 & 0 & 0\\
# 0 & 0 & 0 & E_4 & 0 & 0\\
# 0 & 0 & 0 & 0 & E_5 & 0\\
# 0 & 0 & 0 & 0 & 0 & E_6
# \end{matrix} \right ] \cdot
# \left [ \begin{matrix}
# c_1\\
# c_2\\
# c_3\\
# c_4\\
# c_5\\
# c_6
# \end{matrix} \right ]$$
# + [markdown] id="uwKnGRHjcA7N" colab_type="text"
# Entonces, factorizando un poco y dividiendo entre $\beta$, se puede tener la siguiente situación:
#
# $$\left [ \begin{matrix}
# \frac{\left ( \alpha - E \right )}{\beta}& 1 & 0 & 0 & 0 & 1\\
# 1 & \frac{\left ( \alpha - E \right )}{\beta} & 1 & 0 & 0 & 0\\
# 0 & 1 & \frac{\left ( \alpha - E \right )}{\beta} & 1 & 0 & 0\\
# 0 & 0 & 1 & \frac{\left ( \alpha - E \right )}{\beta} & 1 & 0\\
# 0 & 0 & 0 & 1 & \frac{\left ( \alpha - E \right )}{\beta} & 1\\
# 1 & 0 & 0 & 0 & 1 & \frac{\left ( \alpha - E \right )}{\beta}
# \end{matrix} \right ] \cdot
# \left [ \begin{matrix}
# c_1\\
# c_2\\
# c_3\\
# c_4\\
# c_5\\
# c_6
# \end{matrix} \right ] = 0$$
# + [markdown] id="N0PRMmt9cEwc" colab_type="text"
# Si definimos $x_i = \frac{E_i - \alpha}{\beta}$, entonces es posible reescribir la anterior ecuación como:
#
# $$\left [ \begin{matrix}
# -x_1& 1 & 0 & 0 & 0 & 1\\
# 1 & -x_2 & 1 & 0 & 0 & 0\\
# 0 & 1 & -x_3 & 1 & 0 & 0\\
# 0 & 0 & 1 & -x_4 & 1 & 0\\
# 0 & 0 & 0 & 1 & -x_5 & 1\\
# 1 & 0 & 0 & 0 & 1 & -x_6
# \end{matrix} \right ] \cdot
# \left [ \begin{matrix}
# c_1\\
# c_2\\
# c_3\\
# c_4\\
# c_5\\
# c_6
# \end{matrix} \right ] = 0$$
# + [markdown] id="NC5OZOHWcLph" colab_type="text"
# Lo cual es simplemente el problema de valores propios del que partimos, pero con $E_i = \alpha + x_i \beta$.
#
# $$\left [ \begin{matrix}
# 0 & 1 & 0 & 0 & 0 & 1\\
# 1 & 0 & 1 & 0 & 0 & 0\\
# 0 & 1 & 0 & 1 & 0 & 0\\
# 0 & 0 & 1 & 0 & 1 & 0\\
# 0 & 0 & 0 & 1 & 0 & 1\\
# 1 & 0 & 0 & 0 & 1 & 0
# \end{matrix} \right ] \cdot
# \left [ \begin{matrix}
# c_1\\
# c_2\\
# c_3\\
# c_4\\
# c_5\\
# c_6
# \end{matrix} \right ] =
# x_i \cdot
# \left [ \begin{matrix}
# c_1\\
# c_2\\
# c_3\\
# c_4\\
# c_5\\
# c_6
# \end{matrix} \right ]$$
# + [markdown] id="gAu2_Sy8dCb4" colab_type="text"
# Ahora vamos a proceder a resolver el problema de valores propios, y así encontrar los valores de energía del sistema. Lo primero que vamos a intentar será descomponer la matriz $\mathbf{H}$ en matrices $Q$ y $R$.
# + id="2R9UdVNHVLFA" colab_type="code" colab={}
# Ejecuta esta celda sin realizar cambios en ella
import numpy as np
import numpy.linalg as LA
import matplotlib.pyplot as plt
import matplotlib.cm as cm
np.set_printoptions(suppress=True)
# + id="uxciXMqadrMY" colab_type="code" colab={}
# Creamos la matriz H de la molecula de benceno
benceno = np.array([[0,1,0,0,0,1],
[1,0,1,0,0,0],
[0,1,0,1,0,0],
[0,0,1,0,1,0],
[0,0,0,1,0,1],
[1,0,0,0,1,0]])
# + id="MbvS4sageXLl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="b74a26c6-670f-45b4-d63f-491e0c6675fd"
# Calculamos las matrices Q y R a partir de la matriz de Huckel del benceno
q, r = LA.qr(benceno)
np.around(q, decimals=4)
# + id="WyBjJdJUetyS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="d37f02d6-4566-4813-da14-e84988ac8ff7"
np.around(r, decimals=4)
# + [markdown] id="XEN-IeIaiJ70" colab_type="text"
# En este punto es probable que surja la pregunta: ¿Y para qué me sirve poder realizar la descomposición QR? Pues esta es la parte más interesante de todo lo que se hará hoy. Hasta ahora, solo sabíamos calcular los valores y vectores propios de una matriz mediante su polinomio característico. Sin embargo, hace unos momentos vimos cómo es que diagonalizar una matriz lleva al mismo resultado. Entonces, recordando en principio cómo se diagonaliza una matriz $A$ (se busca una matriz $P$ tal que $P^{-1} A P = D$ ) y recordando cómo se halla la matriz $R$ en una descomposición QR ($R = Q^{-1} A$), vamos a realizar el siguiente análisis:
#
# $$A = QR$$
#
# $$R Q = Q^{-1} A Q = A'$$
#
# Entonces, ¡multiplicar $R$ por $Q$ es equivalente a diagonalizar $A$! El problema es que esto no sucede inmediatamente. $A'$ se aproxima a la forma diagonalizada de $A$, pero no es la matriz diagonal. ¿Entonces qué? Pues repetimos este procedimiento. Esto se repite tantas veces como sea necesario hasta que los valores de $Q$ y de $R$ ya no cambien.
# + id="ptkKwuzhfJM3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="1cd38f76-9546-4801-91bd-83b42346cab6"
# Primero vamos a volver a la matriz H del benceno; a la larga es Q por R
b = np.dot(q,r)
np.around(b, decimals=4)
# + id="WKQa9R2egNOI" colab_type="code" colab={}
# Ahora vamos a repetir el proceso de factorizar la matriz
# y generar una nueva matriz 200 veces
for i in xrange(200):
q, r = LA.qr(b)
b = np.dot(np.dot(np.transpose(q),b), q)
# + id="5t_KCzIggN9T" colab_type="code" colab={}
# Finalmente vamos a volver a hallar las matrices Q y R
q, r = LA.qr(b)
# + id="K3CisMeshpdI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="71e62b54-6788-456f-ec99-e326e69fb632"
# Y es la matriz R la contiene los valores propios en la diagonal principal
np.around(r, decimals=4)
# + [markdown] id="z4z3Lg26oUF_" colab_type="text"
# A este punto vale la pena decir que los valores propios de $\mathbf{H}$ son `[-2, -1, -1, 1, 1, 2]`. Entonces, ¿existe algún error? Lo que sucede es que nuestros ordenadores están trabajando con pocos decimales relativamente y este problema requiere mucho mayor precisión. Por la misma razón, se han tenido que buscar alternativas para este tipo de problemas. Sin embargo, vale la pena resaltar que nuestros ordenadores **sí** utilizan el algoritmo QR para este tipo de problemas, solo que este cuenta con otro par de modificaciones para evitar estos errores. A continuación se presenta la manera en que lo hace NumPy.
# + id="ePnEDGxNnrUD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3e604bcb-d613-4820-b8a1-e94a76ebe45e"
# Calculamos los valores y vectores propios de la matriz H en una linea
w, v = LA.eigh(benceno)
# Luego mostramos los valores propios
np.around(w, decimals=4)
# + id="N6ZFQO1wooDW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="d6cc1596-b22a-4b9f-cfd8-9eb26ba2a2c5"
# Y finalmente, observamos los vectores propios
np.around(v, decimals=4)
# + id="vLi5DAHuqo3Y" colab_type="code" colab={}
# Ejecuta esta celda sin hacer cambios en ella
# Funcion de onda para orbital 2p de un atomo de carbono
def Psi(x, y, z, x0, y0, z0):
r = ((x - x0)**2 + (y - y0)**2 + (z - z0)**2)**0.5
A = (3.25**5 / (32*np.pi))**0.5
ex = -3.25 * r / 2
return A * (z - z0) * np.e**ex
# + id="2_eJZB-uwtIN" colab_type="code" colab={}
# Ejecuta esta celda sin hacer cambios en ella
# Funcion para graficar los orbitales
def orbital2D(coords, constantes, z=1, orbital=0, delta=0.05):
if len(coords) != len(constantes):
raise ValueError, """El numero de coordenadas no coincide
con el numero de constantes"""
n = len(coords)
lx = min([c[0] for c in coords])
ux = max([c[0] for c in coords])
ly = min([c[1] for c in coords])
uy = max([c[1] for c in coords])
domx = np.arange(lx - 2, ux + 2 + delta, delta)
domy = np.arange(ly - 2, uy + 2 + delta, delta)
densmap = [[0]*len(domx)]*len(domy)
for iy in range(len(domy)):
for ix in range(len(domx)):
densmap[iy][ix] = 0
for i in range(n):
p = Psi(domx[ix], domy[iy], z, coords[i][0], coords[i][1], coords[i][2])
densmap[iy][ix] += ( constantes[i][n-orbital-1] * p )
densmap[iy] = tuple(densmap[iy])
dm = np.array(densmap, dtype=np.float32)
plt.imshow(dm, cmap=cm.viridis)
plt.grid(False)
plt.axis('equal')
plt.axis('off')
plt.show()
# + id="CCDVk-8jxFh0" colab_type="code" colab={}
# Ahora vamos a necesitar las coordenadas de los atomos de carbono en el benceno
benceno_coords = [[2.6,0,0], [1.3,2.2517,0],[-1.3,2.2517,0],
[-2.6,0,0],[-1.3,-2.2517,0],[1.3,-2.2517,0]]
# + id="qDpP0ddIzuUd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 350} outputId="d9bef2e6-047e-41af-c29c-68accae6923d"
# Finalmente, graficamos los orbitales pi del benceno
# Para cambiar el orbital, cambia el numero aqui abajo y corre otra vez la celda
# En el benceno hallamos 6 niveles de energia
# Entonces hay orbitales del 0 al 5
orbital2D(benceno_coords, v, orbital=0)
# + [markdown] id="ZADHF0R3_Jte" colab_type="text"
# Este momento es para detenerse y reflexionar: ¿Qué acabamos de hacer? Con una simple **función** y un poco de **álgebra de matrices** acabamos de encontrar la ubicación de los electrones que se hallan en resonancia en el benceno. No solo eso, ¡hallamos cómo se comportan a diferentes niveles de energía! Es importante recordar que solo estamos hablando de los electrones en los orbitales $\pi$. Sin embargo, esto ya complementa la intuición química sobre el comportamiento de los electrones en moléculas con resonancia.
#
# Otras observaciones que quizá vale la pena mencionar, a pesar de que no son tema de esta práctica directamente son las siguientes:
# 1. Aquellos orbitales que tienen la misma energía (i.e. el mismo valor propio) son llamados *degenerados*
# 2. Los orbitales negativos de mayor energía son llamados **orbitales moleculares ocupados de más energía**, u **HOMO** por sus siglas en inglés.
# 3. Los orbitales positivos de menor energía son llamados **orbitales moleculares desocupados de menor energía**, u **LUMO** por sus siglas en inglés.
# 4. Es conveniente poner atención a las simetrías que se presentan entre los orbitales degenerados. Los planos de simetría suelen ser peculiares cuando se trata de este tipo de orbital.
#
# ---
#
# Ahora que ya hemos visto cómo hacer esto para el benceno, vamos a repetir el mismo ejercicio con el naftaleno, el estireno, el azuleno o el coroneno. **¡Solo es necesario hacer uno!**
#
# El procedimiento se desarrollará de la siguiente manera:
# * Crear la matriz de Hückel de 0s y 1s
# * Factorizar la matriz en Q y R
# * Encontrar los valores propios utilizando el algoritmo QR
# * Encontrar los valores y vectores propios utilizando la función de NumPy: `LA.eigh()`
# * Graficar los orbitales utilizando la función `orbital2D()`
# * Guardar el cuaderno interactivo e imprimir un PDF del mismo
# + [markdown] id="nc4qTTA7JRBQ" colab_type="text"
# |Azuleno|Coroneno|Estireno|Naftaleno
# |---|---|---|---|
# |||||
# + id="hCLBo5e40AQy" colab_type="code" colab={}
# Coordenadas para las moleculas
azuleno_coords = [[-1.21800105,-3.02633781,0],[-0.0819981,-3.97084248,0],
[1.21800105,-3.02633781,0],[0.72144557,-1.49809715,0],
[-0.72144557,-1.49809715,0],[-1.62107347,-0.37,0],
[-1.3,1.0367154,0],[0,1.6627624,0],
[1.3,1.0367154,0],[1.62107347,-0.37,0]]
coroneno_coords = [[0,2.6,0],[-2.2517,1.3,0],[-2.2517,-1.3,0],
[0,-2.6,0],[2.2517,-1.3,0],[2.2517,1.3,0],
[-4.5033,2.6,0],[-6.7551,1.3,0],[-6.7551,-1.3,0],
[-4.5033,-2.6,0],[4.5033,2.6,0],[4.5033,-2.6,0],
[6.7551,-1.3,0],[6.7551,1.3,0],[2.2517,6.5,0],
[0,5.2,0],[4.5033,5.2,0],[-2.2517,6.5,0],
[-4.5033,5.2,0],[-4.5033,-5.2,0],[-2.2517,-6.5,0],
[0,-5.2,0],[2.2517,-6.5,0],[4.5033,-5.2,0]]
estireno_coords = [[-2.2517,2.6,0],[-4.5033,1.3,0],[-4.5033,-1.3,0],
[-2.2517,-2.6,0],[0,-1.3,0],[0,1.3,0],
[2.2517,2.6,0],[4.5033,1.3,0]]
naftaleno_coords = [[-2.2517,2.6,0],[-4.5033,1.3,0],[-4.5033,-1.3,0],
[-2.2517,-2.6,0],[0,-1.3,0],[0,1.3,0],
[2.2517,2.6,0],[2.2517,-2.6,0],[4.5033,-1.3,0],
[4.5033,1.3,0]]
# + id="wV5VMzpddLpH" colab_type="code" colab={}
| FactorizacionDiagonalizacion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Extrai informações da base de dados do EPM
import pyodbc
import pandas as pd
from tqdm import tqdm
import os
# > Parâmetros de conexão na base de dados.
server = '172.16.17.32'
database = 'EPM_Database1'
username = 'epasa'
password = '<PASSWORD>'
# > Busca da base de dados a tags que foram cadastradas no sistema.
cnxn = pyodbc.connect('DRIVER={ODBC Driver 17 for SQL Server};SERVER='+server+';DATABASE='+database+';UID='+username+';PWD='+ password)
df_tags = pd.read_sql_query("select * from Epm_Tags", cnxn)
tags_list = df_tags['Name'].unique()
cnxn.close()
# > Lista de tags para serem extraidas da base de dados
tags_generating_unit = ['1ST1000','1TE6575','1TE6180A','1TE6180B','2PT6180A','2PT6180B','1ST1004A','1ST1004B','2GT1022','1PT5070','1TE5070','ANALOG_164']
tags_factory = ['FQ003']
# > Monta um dicionário com todas as tags que tem de extrair para cada unidade geradora
# +
# tag_per_machine = {}
# for i in range(1,41):
# machine_number = 'UGD{}'.format(i)
# filtered_tags = []
# for tag in tags_generating_unit:
# filtered_tags = filtered_tags + list(filter(lambda x: machine_number+"_" in x and tag in x, tags_list))
# tag_per_machine[machine_number] = filtered_tags
# -
# > Busca as tags e escreve os arquivos, um arquivo para cada tag
# +
# for key, item in tqdm(tag_per_machine.items()):
# if key not in ['UGD1','UGD2','UGD3','UGD4','UGD5']:
# os.mkdir(+key)
# for i in range(len(item)):
# cnxn = pyodbc.connect('DRIVER={ODBC Driver 17 for SQL Server};SERVER='+server+';DATABASE='+database+';UID='+username+';PWD='+ password)
# query = "SELECT CAST(A.Timestamp as smalldatetime) as DataHora, CAST(A.Value as numeric) AS {} FROM dbo.EpmQueryRawFunction(-3,'01/01/2018 00:00:00','05/30/2020 00:00:00',0,0,'{}') AS A".format(item[i], item[i])
# unidade_geradora = pd.read_sql_query(query, cnxn)
# unidade_geradora.to_csv('./../../dataset/01.real/epm/{}/{}_{}.tar.xz'.format(key,key,item[i]), chunksize=10000, compression='xz')
# cnxn.close()
# -
# > Escreve um arquivo exlusivo para uma tag.
# +
# # pd.read_csv('UGD1_Main_1ST1000.tar.xz', compression='xz')
# tag = "UGD1_Main_1ST1000"
# cnxn = pyodbc.connect('DRIVER={ODBC Driver 17 for SQL Server};SERVER='+server+';DATABASE='+database+';UID='+username+';PWD='+ password)
# query = "SELECT CAST(A.Timestamp as smalldatetime) as DataHora, CAST(A.Value as numeric) AS {} FROM dbo.EpmQueryRawFunction(-3,'01/01/2018 00:00:00','02/02/2018 00:00:00',0,0,'{}') AS A".format(tag, tag)
# unidade_geradora = pd.read_sql_query(query, cnxn)
# unidade_geradora.to_csv('{}.tar.xz'.format('UGD1_Main_1ST1000'), chunksize=10000, compression='xz')
# cnxn.close()
# -
def get_tag_from_database(tag):
print(tag)
# > Capturar todas as tags do banco de dados
| util-code/01.notebook/1_get_data_sql.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CIFAR-10 Differential Attack
#
# ## Configuration
# +
import os
import numpy as np
import tensorflow as tf
# -
FILE_PATH = os.getcwd()
MODEL_PATH = os.path.join(FILE_PATH, "../models/my_vgg.h5")
os.environ["TF_FORCE_GPU_ALLOW_GROWTH"] = "true"
# ## Dataset preparation
#
# We work with categorical (binary class matrix) instead of class vectors (integers).
# +
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.utils import to_categorical
def __prepare_datasets():
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
return (x_train, y_train), (x_test[:500], y_test[:500])
# -
data_train, data_test = __prepare_datasets()
x_train, y_train = data_train
x_test, y_test = data_test
print(f"x_train.shape = {x_train.shape} y_train.shape = {y_train.shape}")
print(f"x_test.shape = {x_test.shape} y_test.shape = {y_test.shape}")
# ## Model preparation
#
# We use our own VGG model.
# +
from integration_tests.models.my_vgg import my_vgg
def __prepare_model(data_train, data_test):
if os.path.exists(MODEL_PATH):
print("---Using Existing Model---")
model: tf.keras.Model = tf.keras.models.load_model(MODEL_PATH)
else:
print("---Training Model---")
print(f"GPU IS AVAILABLE: {tf.config.list_physical_devices('GPU')}")
model: tf.keras.Model = my_vgg()
model.fit(
*data_train,
epochs=100,
batch_size=64,
validation_data=data_test,
)
model.save(MODEL_PATH)
model.summary()
return model
# -
model = __prepare_model(data_train, data_test)
# ## Prior Prediction
y_preds = model.predict(x_test)
y_preds.shape
# ## Main Algorithm
#
# Original algorithm:
#
# ```txt
# CornerSearch:
# - One pixel attack : Choose N pixels with differential evolution
# - For each pixel (x, y, r, g, b), create 8 images (x,y,0,0,0), (x,y,255,0,0), (x,y,0,255,0)... (x,y,255,255,255)
# - Test the 8 images and save the pertubations where M(x_fake) != y_true
#
# Assuming Y_fake, a group of prediction
# For each pair (image, label) x_test, y_true in X (group of images) and Y (group of true prediction):
# - Create a fragile pixel pool P of x based on the algorithm above
# - For each pixel p=(x, y, r, g, b) in the fragile pixel pool P:
# - x_fake = InjectFault(p, x_test) (or InjectFaultByBit(p.x, p.y, msb_index, channel, bit_flip/set/reset))
# - M(x_fake) = y_fake, and we store y_fake in a temporary group called Y_temp
# - If one of Y_temp has y_fake != y_true:
# - Y_fake append y_fake
# Else:
# - Y_fake append y_pred (normal prediction W/O FI)
# - Measure the accuracy of Y_fake using mean
# ```
#
# We do a little modification :
#
# ```txt
# CornerSearch:
# - One pixel attack : Choose N pixels with differential evolution
# - For each pixel (x, y, r, g, b), create 8 images (x,y,0,0,0), (x,y,255,0,0), (x,y,0,255,0)... (x,y,255,255,255)
# - Test the 8 images and save the pertubations where M(x_fake) != y_true
#
# Assuming Y_fake, a group of prediction
# For each pair (image, label) x_test, y_true in X (group of images) and Y (group of true prediction):
# - If already missclassified:
# - Y_fake append y_pred (normal prediction W/O FI)
# - Create a fragile pixel pool P of x based on the algorithm above
# - For each pixel p=(x, y, r, g, b) in the fragile pixel pool P:
# - x_fake = InjectFault(p, x_test) (or InjectFaultByBit(p.x, p.y, msb_index, channel, bit_flip/set/reset))
# - M(x_fake) = y_fake, and we store y_fake in a temporary group called Y_temp
# - If one of Y_temp has y_fake != y_true:
# - Y_fake append y_fake
# Else:
# - Y_fake append y_pred (normal prediction W/O FI)
# - Measure the accuracy of Y_fake using mean
# ```
#
# ```python
# CORNERS = (
# (0, 0, 0),
# (255, 255, 255),
# (0, 0, 255),
# (0, 255, 0),
# (0, 255, 255),
# (255, 0, 0),
# (255, 0, 255),
# (255, 255, 0),
# )
#
#
# def corner_search(
# image_id: int,
# pixels: np.ndarray,
# data_test: np.ndarray,
# model: tf.keras.Model,
# ) -> Iterable[Tuple[np.ndarray, np.ndarray, PixelFault]]:
# x_test, y_test = data_test
#
# y_true = y_test[image_id]
# y_true_index = np.argmax(y_true)
#
# for pixel in pixels:
# corner_pixels = [PixelFault(pixel.x, pixel.y, r, g, b) for r, g, b in CORNERS]
#
# x_fakes = np.array(
# [
# build_perturb_image([corner_pixel])(x_test[image_id])
# for corner_pixel in corner_pixels
# ]
# )
# y_preds = model.predict(x_fakes)
#
# for x_fake, y_pred, corner_pixel in zip(
# x_fakes,
# y_preds,
# corner_pixels,
# ):
# y_pred_index = np.argmax(y_pred)
# if y_true_index != y_pred_index:
# yield x_fake, y_pred, corner_pixel
#
# ```
# +
from inputtensorfi.attacks.utils import attack
from inputtensorfi.manipulation.img.faults import PixelFault
def _look_for_pixels(
image_id: int,
data_test: np.ndarray,
model: tf.keras.Model,
pixel_count=1,
):
x_test, y_test = data_test
x = x_test[image_id]
y_true = y_test[image_id]
y_true_index = np.argmax(y_true)
pixels = attack(
x,
y_true_index,
model,
pixel_count=pixel_count,
maxiter=10,
verbose=False,
).astype(np.uint8)
# Convert [x_0, y_0, r_0, g_0, b_0, x_1, ...]
# to [pixel_fault_0, pixel_fault_1, ...]
return np.array([PixelFault(*pixels[0:5]) for i in range(len(pixels) // 5)])
# +
from typing import Dict
from inputtensorfi.attacks.corner_search import corner_search
length = len(y_preds)
y_fake = y_preds.copy()
total_faults: Dict[int, PixelFault] = dict()
for image_id, _ in enumerate(y_test):
if np.argmax(y_preds[image_id]) != np.argmax(y_test[image_id]):
print(f"MISPREDICTED {image_id}/{length}")
continue
pixels = _look_for_pixels(image_id, data_test, model, pixel_count=10)
try:
first_pred = next(corner_search(image_id, pixels, data_test, model))
_, y_pred, pixel = first_pred
y_fake[image_id] = y_pred
total_faults[image_id] = pixel
print(
f"FAULT {image_id}/{length}, {pixel}, original={data_test[0][image_id, pixel.x, pixel.y]}"
)
except StopIteration:
# print(f"NO FAULT image_id={image_id}")
pass
# +
import json
dict_data = {key: fault.to_dict() for key, fault in total_faults.items()}
print(f"total_faults={json.dumps(dict_data, indent=2)}")
# -
# ## Accuracies
y_true_acc = np.array([np.max(y) for y in y_test])
y_preds_acc = np.array([y[np.argmax(y_true)] for y, y_true in zip(y_preds, y_test)])
y_fake_acc = np.array([y[np.argmax(y_true)] for y, y_true in zip(y_fake, y_test)])
print(f"y_true_acc={np.mean(y_true_acc)}")
print(f"y_prior_acc={np.mean(y_preds_acc)}")
print(f"y_fake_acc={np.mean(y_fake_acc)}")
| integration_tests/attacks/cifar10_differential_attack_with_corner_search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:splicing]
# language: python
# name: conda-env-splicing-py
# ---
# %load_ext autoreload
# %autoreload 2
from dataloader import SplicingVCFDataloader
fasta = "example_files/hg19.nochr.chr17.fa"
gtf = "example_files/test.gtf"
vcf = "example_files/test.vcf.gz"
dl = SplicingVCFDataloader(gtf,
fasta,
vcf,
# out_file="/tmp/test.pkl",
overhang=(100,100),
split_seq=False)
dliter = dl.batch_iter()
test = next(dliter)
test['inputs'].shape
from model import MMSpliceModel
model = MMSpliceModel()
model.predict_on_batch(test['inputs'])
| MMSplice/splicingEfficiency/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set()
pima_df = pd.read_csv('../data/pima.csv', dtype = {'diabetes': int})
pima_df.head(3)
drinks_df = pd.read_csv('../data/drinks-by-country.csv')
drinks_df.head(3)
flight_df = sns.load_dataset('flights')
flight_df.head(3)
ufo_df = pd.read_csv('../data/ufo.csv')
ufo_df.columns = ufo_df.columns.str.lower().str.replace(' ', '_')
ufo_df.time = pd.to_datetime(ufo_df.time)
ufo_df['year'] = ufo_df.time.dt.year
ufo_df.head(3)
imdb_df = pd.read_csv('../data/imdb.csv')
imdb_df.head(3)
# # Bar Chart
# +
sns.set_context('talk')
plt.figure(figsize = (15, 5))
drinks_df.groupby('continent').mean().plot(kind = 'bar', figsize = (15, 5), cmap = 'viridis', rot = 0)
plt.show()
# +
sns.set_context('talk')
pd.crosstab(imdb_df.genre, imdb_df.content_rating).plot(kind = 'bar', rot = 45, figsize = (16, 6),
stacked = True, cmap = 'icefire')
plt.show()
# -
# # Pie Chart
# +
sns.set_context('talk')
with sns.color_palette("Spectral", 12):
flight_df.groupby('year').mean().plot(kind = 'pie', subplots = True, figsize = (7, 7), legend = False, autopct = '%1.1f%%')
plt.ylabel('')
plt.title('Passengers\nYear by Year')
plt.show()
# -
# # Line Chart
# +
sns.set_context('talk')
imdb_df.star_rating.plot(figsize = (10, 4))
plt.show()
# +
sns.set_context('talk')
pd.crosstab(imdb_df.genre, imdb_df.content_rating).plot(figsize = (16, 6), rot = 45)
plt.show()
# -
# # Histogram
# +
sns.set_context('talk')
ufo_df.plot(kind = 'hist', figsize = (10, 4), bins = 20)
plt.show()
# -
# # Box Plot
# Pandas Box Plot Looks Ugly
# # Scatter Plot
# +
sns.set_context('talk')
pima_df.plot(kind = 'scatter', x = 'num_preg', y = 'diastolic_bp', figsize = (15, 5), s = pima_df['age'] * 2)
plt.show()
# -
# # Hexbin
# Pandas Hexbin is difficult to setup
# # KDE Plot
# +
sns.set_context('talk')
ufo_df.plot(kind = 'kde', figsize = (10, 4))
plt.show()
# -
# # Andrew Plot
from pandas.plotting import andrews_curves
iris_df = pd.read_csv('../data/iris.csv')
iris_df.head(3)
# +
sns.set_context('talk')
plt.figure(figsize = (10, 6))
andrews_curves(iris_df, 'species', colormap = 'viridis_r')
plt.show()
| visualization/pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Analysis based on wards
# +
from random import shuffle, randint
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.collections import PatchCollection
from mpl_toolkits.basemap import Basemap
from shapely.geometry import Polygon, Point, MultiPoint, MultiPolygon, LineString, mapping, shape
from descartes import PolygonPatch
import random
import fiona
import numpy as np
import csv
from fiona import collection
import geopandas as gpd
from geopandas.tools import sjoin # rtree index in-build, used with inner, intersection
import pandas as pd
from collections import defaultdict
# +
def find_intersections(o):
from collections import defaultdict
paired_ind = [o.index_PP, o.index_WARD]
d_over_ind = defaultdict(list)
# creating a dictionary that has prescints as keys and associated small areas as values
for i in range(len(paired_ind[0].values)):
if not paired_ind[0].values[i]==paired_ind[1].values[i]: # it shows itself as intersection
d_over_ind[paired_ind[0].values[i]].append(paired_ind[1].values[i])
# get rid of the pol precincts with no small areas associated to them- not the most efficient way
d_temp = {}
for l in d_over_ind:
if len(d_over_ind[l]):
d_temp[l] = d_over_ind[l]
return d_temp
def calculate_join_indices(g1_reind, g2_reind):
# the indexing:
out = sjoin(g1_reind, g2_reind, how ="inner", op = "intersects")
out.drop('index_right', axis=1, inplace=True) # one index doubles, so we drop one
# output retains only 1 area (left or right join), and gives no intersection area.
# so we create an array with paired indices: police precincts with a ward
# we use it in a loop in a function below
dict_over_ind = find_intersections(out)
return dict_over_ind
# -
def calculate_join(dict_over_ind, g1_reind, g2_reind):
data_aggreg = []
for index1, crim in g1_reind.iterrows():
try:
index1 = crim.index_PP
wards_found = dict_over_ind[index1]
for ward in range(len(wards_found)):
pom = g2_reind[g2_reind.index_WARD == wards_found[ward]]['geometry']
area_int = pom.intersection(crim['geometry']).area.values[0]
area_crim = crim['geometry'].area
area_ward = pom.values[0].area
popu_ward = g2_reind[g2_reind.index_WARD == wards_found[ward]]['WARD_POP'].values[0]
murd_count = crim['murd_cnt']
pp_province = crim['province']
compnt_nm = crim['compnt_nm']
popu_frac = (area_int / area_ward) * popu_ward# fraction of the pop area contained inside the crim
extra_info_col_names = ['MUNICNAME', 'PROVINCE', 'WARD_ID']
extra_cols = g2_reind[g2_reind.index_WARD == wards_found[ward]][extra_info_col_names]#.filter(regex=("NAME"))
data_aggreg.append({'geometry': pom.intersection(crim['geometry']).values[0],\
'index_PP': index1,'index_WARD': wards_found[ward] ,'area_pp': area_crim,\
'area_ward': area_ward,'area_inter': area_int,\
'popu_inter' : popu_frac, 'popu_ward': popu_ward,\
'murd_cnt': murd_count,'province': pp_province,\
'MUNICNAME': extra_cols.MUNICNAME.values[0],\
'PROVINCE': extra_cols.PROVINCE.values[0],\
'WARD_ID': extra_cols.WARD_ID.values[0],\
'PoliceStation': compnt_nm})
except:
pass
df_t = gpd.GeoDataFrame(data_aggreg,columns=['geometry', 'index_PP','index_WARD','area_pp',\
'area_ward','area_inter', 'popu_inter',\
'popu_ward', 'murd_cnt','province',\
'MUNICNAME','PROVINCE','WARD_ID', 'PoliceStation'])
#df_t.to_file(out_name)
return df_t
# this function adds the remaining columns, calculates fractions etc
def compute_final_col(df_temp):
# add population data per PP, ratio, etc to the main table
temp = df_temp.groupby(by=['index_PP'])['popu_inter'].sum().reset_index()
data_with_population = pd.merge(df_temp, temp, on='index_PP', how='outer')\
.rename(columns={'popu_inter_y':'popu_frac_per_pp', 'popu_inter_x':'popu_inter'})
data_with_population['murd_per_int'] = data_with_population['popu_inter']/data_with_population['popu_frac_per_pp']\
* data_with_population['murd_cnt']
data_mur_per_int = data_with_population.groupby(by=['index_WARD'])['murd_per_int'].sum().reset_index()
data_mur_per_sal = data_mur_per_int.rename(columns={'murd_per_int':'murd_per_ward'})
data_with_population['scal_fac_per_int'] = data_with_population['popu_inter']/data_with_population['popu_frac_per_pp']\
data_complete = pd.merge(data_with_population, data_mur_per_sal, on='index_WARD', how='outer')
return data_complete
# LOAD the data
# +
ppSHP = 'shapefiles/updated/polPrec_murd2015_prov_aea.shp'
warSHP = '../maps/data/Wards2011_aea.shp'
geo_ward = gpd.GeoDataFrame.from_file(warSHP)
geo_pp = gpd.GeoDataFrame.from_file(ppSHP)
geo_pp_reind = geo_pp.reset_index().rename(columns={'index':'index_PP'})
geo_ward_reind = geo_ward.iloc[:,[2,3,7,8,9]].reset_index().rename(columns={'index':'index_WARD'})
dict_int = calculate_join_indices(geo_pp_reind,geo_ward_reind)
# -
# number of wards
len(geo_ward)
geo_pp_reind.head(n=2)
# +
from timeit import default_timer as timer
start = timer()
df_int = calculate_join(dict_int ,geo_pp_reind, geo_ward_reind)
end = timer()
print("time: ", end - start)
# -
# There are 101,546 intersections
df_int_aea = compute_final_col(df_int) # add final calculations
df_int_aea.head(n=2)
# add column to indicate type of unit
df_int_aea['unit'] = 'ward'
df_int_aea.head(n=2)
df_int_aea_waz = df_int_aea.iloc[:,[18,12,11,10,13,6,16,15]].sort('WARD_ID').reset_index().drop('index', axis=1)
#df_int_aea_waz.to_csv('data/wards_pp_intersections_basic.csv')
df_int_aea.to_csv('data/wards_pp_intersections_with_geo.csv')
df_int_aea_waz.head(n=2)
# Check whether the sums over provinces add up to the official/initial numbers:
df_int_aea.head()
data_prov = df_int_aea[['PROVINCE','province','murd_per_int']]
data_prov.groupby('province')['murd_per_int'].sum()
# Adding the remaining crimes based on the PP ID:
def csv_dict_reader(file_obj):
"""
Read a CSV file using csv.DictReader
"""
reader = csv.DictReader(file_obj, delimiter=',')
data_list=[]
for line in reader:
data_list.append(line)
return data_list
# read in the additional crime data
pp_data=pd.read_csv("data/pp_crime_2015.csv")
pp_data.head(n=2)
# pivot for indexing per police station and crime types as columnd
df_crimes = pp_data.pivot('PoliceStation' ,'Crime','Incidents').reset_index()
# replace by lower case to merge with the existing table
df_crimes['PoliceStation'] = df_crimes['PoliceStation'].map(lambda x: x if type(x)!=str else x.lower())
df_crimes.head()
temp = pd.merge(df_int_aea_waz, df_crimes, on = 'PoliceStation', how ='outer')
temp.columns # 55 column
temp.iloc[:,8:55] = temp.iloc[:,8:55].multiply(temp["scal_fac_per_int"], axis="index")
temp.head()
len(df_int_aea_waz_full)
df_int_aea_waz_full = temp
df_int_aea_waz_full.to_csv('data/wards2011_pp_intersections_full.csv')
df_int_aea_waz_full.head()
| crime_by_wards.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (cv_fastai)
# language: python
# name: cv_fastai
# ---
import sys
sys.path.append("..")
from utils.download_dataset import download_dataset, extract_zip
# +
import numpy as np
import PIL
from fastai.vision.all import *
from pathlib import Path
from tqdm.auto import tqdm
from shutil import copyfile, rmtree
# +
DATA_PATH = Path.cwd()/'data'
if not DATA_PATH.exists():
DATA_PATH.mkdir(exist_ok=True)
fpath = download_dataset(dataset_name='MAGNETIC_TILE_SURFACE_DEFECTS', dest_dir=DATA_PATH)
fpath
# -
dir_path = extract_zip(fpath)
dir_path
dir_path.ls()
if (dir_path/'MT_Free').exists():
rmtree(dir_path/'MT_Free')
classes = []
for ii in (dir_path).ls():
if ii.is_dir() and ii.stem.startswith('MT_'):
classes.append(ii.stem)
classes
classes_dict = {c:i+1 for i, c in enumerate(classes)}
classes_dict
img_fpaths = [get_files(dir_path/c, extensions='.jpg') for c in classes]
img_fpaths = [ii for sublist in img_fpaths for ii in sublist]
len(img_fpaths)
msk_fpaths = [get_files(dir_path/c, extensions='.png') for c in classes]
msk_fpaths = [ii for sublist in msk_fpaths for ii in sublist]
len(msk_fpaths)
assert len(img_fpaths) == len(msk_fpaths)
# +
path_images = dir_path/'images'
if not path_images.exists():
path_images.mkdir(exist_ok=True)
path_labels = dir_path/'labels'
if not path_labels.exists():
path_labels.mkdir(exist_ok=True)
# -
for msk_path in tqdm(msk_fpaths):
c = msk_path.parent.parent.stem
i = classes_dict[c]
msk = np.array(Image.open(msk_path))
msk[msk>0] = i
Image.fromarray(msk).save(path_labels/msk_path.name)
for img_fpath in tqdm(img_fpaths):
copyfile(img_fpath, path_images/img_fpath.name)
new_img_fpaths = get_files(path_images)
assert len(new_img_fpaths) == len(img_fpaths)
| segmentation/ProcessData_MAGNETIC_TILE_SURFACE_DEFECTS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 作業 : (Kaggle)鐵達尼生存預測
# ***
# +
# 做完特徵工程前的所有準備
import pandas as pd
import numpy as np
import copy
from sklearn.preprocessing import LabelEncoder, MinMaxScaler
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
# 因為擬合(fit)與編碼(transform)需要分開, 因此不使用.get_dummy, 而採用 sklearn 的 OneHotEncoder
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_curve
data_path = 'data/data2/'
df = pd.read_csv(data_path + 'titanic_train.csv')
train_Y = df['Survived']
df = df.drop(['PassengerId', 'Survived'] , axis=1)
df.head()
# -
# 因為需要把類別型與數值型特徵都加入, 故使用最簡版的特徵工程
LEncoder = LabelEncoder()
MMEncoder = MinMaxScaler()
for c in df.columns:
df[c] = df[c].fillna(-1)
if df[c].dtype == 'object':
df[c] = LEncoder.fit_transform(list(df[c].values))
df[c] = MMEncoder.fit_transform(df[c].values.reshape(-1, 1))
df.head()
train_X = df.values
# 因為訓練邏輯斯迴歸時也要資料, 因此將訓練及切成三部分 train / val / test, 採用 test 驗證而非 k-fold 交叉驗證
# train 用來訓練梯度提升樹, val 用來訓練邏輯斯迴歸, test 驗證效果
train_X, test_X, train_Y, test_Y = train_test_split(train_X, train_Y, test_size=0.5)
train_X, val_X, train_Y, val_Y = train_test_split(train_X, train_Y, test_size=0.5)
print('train_X shape: {}'.format(train_X.shape))
print('val_X shape: {}'.format(val_X.shape))
print('test_X shape: {}'.format(test_X.shape))
# # 作業1
# * 請對照範例,完成隨機森林的鐵達尼生存率預測,以及對應的葉編碼+邏輯斯迴歸
# +
# 隨機森林擬合後, 再將葉編碼 (*.apply) 結果做獨熱 / 邏輯斯迴歸
rf = RandomForestClassifier(n_estimators=20, min_samples_split=10, min_samples_leaf=5,
max_features=4, max_depth=3, bootstrap=True)
onehot = OneHotEncoder()
lr = LogisticRegression(solver='lbfgs', max_iter=1000)
"""
Your Code Here (Hint : 隨機森林的葉編碼(.apply)不需要加上[:, :, 0], 直接用rf.apply()調用即可, 本作業其餘寫法相同)
"""
rf.fit(train_X, train_Y)
onehot.fit(rf.apply(train_X))
onehot_val_X = onehot.transform(rf.apply(val_X))
lr.fit(onehot_val_X, val_Y)
# +
# 將隨機森林+葉編碼+邏輯斯迴歸結果輸出
"""
Your Code Here
"""
pred_rf_lr = lr.predict(onehot.transform(rf.apply(test_X)))
fpr_rf_lr, tpr_rf_lr, _ = roc_curve(test_Y, pred_rf_lr)
# 將隨機森林結果輸出
"""
Your Code Here
"""
pred_rf = rf.predict_proba(test_X)[:, 1]
fpr_rf, tpr_rf, _ = roc_curve(test_Y, pred_rf)
# -
label_rf_lr = np.where(pred_rf_lr > 0.5, 1, 0)
label_rf = np.where(pred_rf > 0.5, 1, 0)
print("Gradient Boosting Accuracy: {:.3f}".format(sum(label_rf == test_Y)/len(label_rf)))
print("GBT + LR Accuracy: {:.3f}".format(sum(pred_rf_lr == test_Y)/len(label_rf_lr)))
# # 作業2
# * 上述的結果,葉編碼是否有提高預測的正確性呢?
import matplotlib.pyplot as plt
# %matplotlib inline
# 將結果繪圖
"""
Your Code Here
"""
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_rf, tpr_rf, label='RF')
plt.plot(fpr_rf_lr, tpr_rf_lr, label='RF + LR')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve')
plt.legend(loc='best')
plt.show()
| Day_030_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import pickle
import math
# # Time Consumption
# ## Number of Supports
tree_time_dic = pickle.load(open("exp/mnist_time_support_FastPSD.pk", "rb"))
naive_tree_time_dic = pickle.load(open("exp/mnist_time_support_PSD.pk", "rb"))
ibp_time_dic = pickle.load(open("exp/mnist_time_support_IBP.pk", "rb"))
n_pixels = [28*math.sqrt(1/2), 28, 28*math.sqrt(2), 28*math.sqrt(3), 28*math.sqrt(4), 28*math.sqrt(5), 28*math.sqrt(6)]
x = [int(n_pixel)**2 for n_pixel in n_pixels]
# +
tree_time_list = []
naive_tree_time_list = []
ibp_time_list = []
for key in tree_time_dic.keys():
tree_time_list.append(sum(tree_time_dic[key]) / len(tree_time_dic[key]))
naive_tree_time_list.append(sum(naive_tree_time_dic[key]) / len(naive_tree_time_dic[key]))
ibp_time_list.append(sum(ibp_time_dic[key]) / len(ibp_time_dic[key]))
# -
fig = plt.figure(figsize=(5, 6))
plt.plot(x, tree_time_list, color='blue', label="FS-TSWB (FastPSD)", marker='o')
#plt.plot([size**2 for size in n_pixels], fast_quadtree_time_list, marker='o', color='blue')
plt.plot(x, naive_tree_time_list, color='green', label="FS-TSWB (PSD)", marker='s')
#plt.plot([size**2 for size in n_pixels], quadtree_time_list, marker='o', color='green')
plt.plot(x, ibp_time_list, color='r', label="FS-WB (IBP)", marker='v')
#plt.plot([size**2 for size in n_pixels], ibp_time_list, marker='o', color='r')
plt.xlabel('number of supports', fontsize=20)
plt.ylabel('time consumption [seconds]', fontsize=20)
plt.xticks(x)
plt.xticks(rotation=20)
plt.legend(fontsize=14)
plt.tick_params(labelsize=14)
#plt.yscale("log")
#plt.xscale("log"
# ## Number of Samples
tree_time_dic = pickle.load(open("exp/mnist_time_sample_FastPSD.pk", "rb"))
naive_tree_time_dic = pickle.load(open("exp/mnist_time_sample_PSD.pk", "rb"))
ibp_time_dic = pickle.load(open("exp/mnist_time_sample_IBP.pk", "rb"))
n_samples = [100, 1000, 2000, 3000, 4000, 5000]
# +
tree_time_list = []
naive_tree_time_list = []
ibp_time_list = []
for key in tree_time_dic.keys():
tree_time_list.append(sum(tree_time_dic[key]) / len(tree_time_dic[key]))
naive_tree_time_list.append(sum(naive_tree_time_dic[key]) / len(naive_tree_time_dic[key]))
ibp_time_list.append(sum(ibp_time_dic[key]) / len(ibp_time_dic[key]))
# +
fig = plt.figure(figsize=(5, 6))
plt.plot(n_samples, tree_time_list, color='blue', label="FS-TSWB (FastPSD)", marker="o")
#plt.plot(n_samples, tree_time_list, 'o', color='blue', markersize=4)
plt.plot(n_samples, naive_tree_time_list, color='green', label="FS-TSWB (PSD)", marker="s")
#plt.plot(n_samples, naive_tree_time_list, 'o', color='green', markersize=4)
plt.plot(n_samples, ibp_time_list, color='r', label="FS-WB (IBP)", marker="v")
#plt.plot(n_samples, ibp_time_list, 'o', color='r', markersize=4)
plt.xlabel('number of samples', fontsize=20)
plt.ylabel('time consumption [seconds]', fontsize=20)
plt.xticks(n_samples)
plt.xticks(rotation=20)
plt.legend(fontsize=14)
plt.tick_params(labelsize=14)
# -
# # Memory Consumption
# ## Number of Supports
tree_mem = pickle.load(open("exp/mnist_memory_support_FastPSD.pk", "rb"))
naive_tree_mem = pickle.load(open("exp/mnist_memory_support_PSD.pk", "rb"))
ibp_mem = pickle.load(open("exp/mnist_memory_support_IBP.pk", "rb"))
# +
mean_tree_mem = []
mean_naive_tree_mem = []
mean_ibp_mem = []
for key in tree_mem.keys():
mean_tree_mem.append(sum(tree_mem[key]) / len(tree_mem[key]))
mean_naive_tree_mem.append(sum(naive_tree_mem[key]) / len(naive_tree_mem[key]))
mean_ibp_mem.append(sum(ibp_mem[key]) / len(ibp_mem[key]))
# -
n_pixels = [28*math.sqrt(1/2), 28, 28*math.sqrt(2), 28*math.sqrt(3), 28*math.sqrt(4), 28*math.sqrt(5), 28*math.sqrt(6)]
x = [int(n_pixel)**2 for n_pixel in n_pixels]
# +
fig = plt.figure(figsize=(5, 6))
plt.plot(x, mean_tree_mem, marker='o', label="FS-TSWB (FastPSD)", color="blue")
plt.plot(x, mean_naive_tree_mem, marker='s', label="FS-TSWB (PSD)", color="green")
plt.plot(x, mean_ibp_mem, marker='v', label="FS-WB (IBP)", color="red")
plt.xlabel("number of supports", fontsize=20)
plt.ylabel("memory consumption [MB]", fontsize=20)
plt.xticks(x, fontsize=14)
plt.xticks(rotation=20)
plt.yticks(fontsize=14)
plt.legend(fontsize=14)#, bbox_to_anchor=(1.00, 0.7), loc="upper right")
# -
# # Number of Samples
tree_mem = pickle.load(open("exp/mnist_memory_sample_FastPSD.pk", "rb"))
naive_tree_mem = pickle.load(open("exp/mnist_memory_sample_PSD.pk", "rb"))
ibp_mem = pickle.load(open("exp/mnist_memory_sample_IBP.pk", "rb"))
# +
mean_tree_mem = []
mean_naive_tree_mem = []
mean_ibp_mem = []
for key in tree_mem.keys():
mean_tree_mem.append(sum(tree_mem[key]) / len(tree_mem[key]))
mean_naive_tree_mem.append(sum(naive_tree_mem[key]) / len(naive_tree_mem[key]))
mean_ibp_mem.append(sum(ibp_mem[key]) / len(ibp_mem[key]))
# -
x = [500, 1000, 2000, 3000, 4000, 5000]
# +
fig = plt.figure(figsize=(5, 6))
plt.plot(x, mean_tree_mem, marker='o', label="FS-TSWB (FastPSD)", color="blue")
plt.plot(x, mean_naive_tree_mem, marker='s', label="FS-TSWB (PSD)", color="green")
plt.plot(x, mean_ibp_mem, marker='v', label="FS-WB (IBP)", color="red")
plt.xlabel("number of samples", fontsize=20)
plt.ylabel("memory consumption [MB]", fontsize=20)
plt.xticks(x, fontsize=14)
plt.xticks(rotation=20)
plt.yticks(fontsize=14)
plt.legend(fontsize=14)#, bbox_to_anchor=(1.00, 0.7), loc="upper right")
| figures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import requests
import json
import random, time
### get_matches returns a dataframe from the opendota API with given parameters
### don't run this too often, as there is a request limit of one per second and 50000 pr. month
def get_matches(matches=10000, start_time_less_than=1594771200, mmr_lower=2000, mmr_upper=2500):
# we build the base url for the opendota api
base = 'https://api.opendota.com/api/explorer'
# we create the SQL percent encoded query. The query is as follows
#"""
#SELECT *
#FROM public_matches
#LEFT JOIN public_player_matches
#ON public_matches.match_id = public_player_matches.match_id
#WHERE lobby_type=7 AND game_mode=22 AND avg_mmr!=0 AND start_time<1594771200
#ORDER BY start_time DESC
#LIMIT 10000
#"""
matches_request = f'?sql=SELECT%20*%0AFROM%20public_matches%0ALEFT%20JOIN%20public_player_matches%0AON%20public_matches.match_id%20%3D%20public_player_matches.match_id%0AWHERE%20lobby_type%3D7%20AND%20game_mode%3D22%20AND%20avg_mmr!%3D0%20AND%20start_time<{start_time_less_than}%20AND%20avg_mmr>{mmr_lower}%20AND%20avg_mmr<{mmr_upper}%20AND%20hero_id!%3D0%0AORDER%20BY%20start_time%20DESC%0ALIMIT%20{str(matches*10)}'
url = base + matches_request
result = requests.get(url)
print(result)
j_matches = result.json()
return pd.DataFrame(j_matches["rows"])
# +
def practice_data(matches, sleep):
mmr_steps = [i for i in range(1000, 8001) if i%1000 == 0]
mmr_brackets = [[mmr_steps[i], mmr_steps[i+1]] for i in range(len(mmr_steps)) if mmr_steps[i] < 8000]
dataframes = [get_matches(matches, start_time_less_than=1594771200, mmr_lower=lower, mmr_upper=upper) for lower,upper in mmr_brackets if time.sleep(sleep) is None]
return dataframes
#for lower,upper in mmr_brackets:
# practice_data_f"{lower}"_f"{upper}" = get_matches(matches=100, start_time_less_than=1594771200, mmr_lower=lower, mmr_upper=upper)
# +
#practice_data(100, 2)
# -
practice_data_2000_3000 = get_matches(matches=100000, start_time_less_than=1594771200, mmr_lower=2000, mmr_upper=3000)
practice_data_3000_4000 = get_matches(matches=100000, start_time_less_than=1594771200, mmr_lower=3000, mmr_upper=4000)
practice_data_no_limit = get_matches(matches=100000, start_time_less_than=1594771200, mmr_lower=0, mmr_upper=10000)
practice_data_2000_3000.to_pickle("../data/practice_data_2000_3000.pkl")
practice_data_3000_4000.to_pickle("../data/practice_data_3000_4000.pkl")
practice_data_no_limit.to_pickle("../data/practice_data_no_limit.pkl")
# **We now get some test data in those brackets, which we will take from the day after, to avoid duplicates**
test_data_2000_3000 = get_matches(matches=10000, start_time_less_than=1594857600, mmr_lower=2000, mmr_upper=3000)
test_data_3000_4000 = get_matches(matches=10000, start_time_less_than=1594857600, mmr_lower=3000, mmr_upper=4000)
test_data_no_limit = get_matches(matches=10000, start_time_less_than=1594857600, mmr_lower=0, mmr_upper=10000)
test_data_2000_3000.to_pickle("../data/test_data_2000_3000.pkl")
test_data_3000_4000.to_pickle("../data/test_data_3000_4000.pkl")
test_data_no_limit.to_pickle("../data/test_data_no_limit.pkl")
| notebooks/data_mining_test_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Manipulação e tratamento das bases
import pandas as pd
import numpy as np
#Pré-Processamento das bases
# !pip install imblearn
from imblearn.over_sampling import SMOTE
from sklearn.model_selection import train_test_split
#Modelagem de Dados
from sklearn.ensemble import BaggingClassifier
from sklearn.metrics import accuracy_score
# -
Durante= pd.read_csv('Base_Tratada.csv', sep= ',')
Durante= Durante.loc[Durante['CO_MUN_NOT'].isin([351880])]
Durante=Durante[(Durante['Periodo']==2.0)]
Durante= Durante.drop(columns=["CO_MUN_NOT", "Periodo"])
Durante.head()
# # PRÉ-PROCESSAMENTO
Durante['CS_GESTANT'].replace({1.0: 1, 2.0: 1, 3.0 :1, 4.0 : 1}, inplace= True)
Durante['CS_GESTANT'].replace({5.0: 0, 6.0:0, 9.0:0}, inplace= True)
Durante['CS_RACA'].fillna(9,inplace= True)
Durante['CS_ESCOL_N'].fillna(9,inplace= True)
Durante['SURTO_SG'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['SURTO_SG'].fillna(0,inplace= True)
Durante['NOSOCOMIAL'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['NOSOCOMIAL'].fillna(0,inplace= True)
Durante['FEBRE'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['FEBRE'].fillna(0,inplace= True)
Durante['TOSSE'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['TOSSE'].fillna(0,inplace= True)
Durante['GARGANTA'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['GARGANTA'].fillna(0,inplace= True)
Durante['DISPNEIA'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['DISPNEIA'].fillna(0,inplace= True)
Durante['DESC_RESP'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['DESC_RESP'].fillna(0,inplace= True)
Durante['SATURACAO'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['SATURACAO'].fillna(0,inplace= True)
Durante['DIARREIA'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['DIARREIA'].fillna(0,inplace= True)
Durante['VOMITO'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['VOMITO'].fillna(0,inplace= True)
Durante['PUERPERA'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['PUERPERA'].fillna(0,inplace= True)
Durante['CARDIOPATI'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['CARDIOPATI'].fillna(0,inplace= True)
Durante['HEMATOLOGI'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['HEMATOLOGI'].fillna(0,inplace= True)
Durante['SIND_DOWN'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['SIND_DOWN'].fillna(0,inplace= True)
Durante['HEPATICA'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['HEPATICA'].fillna(0,inplace= True)
Durante['ASMA'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['ASMA'].fillna(0,inplace= True)
Durante['DIABETES'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['DIABETES'].fillna(0,inplace= True)
Durante['NEUROLOGIC'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['NEUROLOGIC'].fillna(0,inplace= True)
Durante['PNEUMOPATI'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['PNEUMOPATI'].fillna(0,inplace= True)
Durante['IMUNODEPRE'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['IMUNODEPRE'].fillna(0,inplace= True)
Durante['RENAL'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['RENAL'].fillna(0,inplace= True)
Durante['OBESIDADE'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['OBESIDADE'].fillna(0,inplace= True)
Durante['ASMA'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['ASMA'].fillna(0,inplace= True)
Durante['ANTIVIRAL'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['ANTIVIRAL'].fillna(0,inplace= True)
Durante['UTI'].replace({2.0: 0, 9.0: 0}, inplace= True)
Durante['UTI'].fillna(0,inplace= True)
Durante['SUPORT_VEN'].replace({3.0: 0, 9.0: 0}, inplace= True)
Durante['SUPORT_VEN'].fillna(0,inplace= True)
Durante['PCR_RESUL'].fillna(4,inplace= True)
Durante['HISTO_VGM'].replace({0: 2}, inplace= True)
Durante['DOR_ABD'].replace({9.0: 0, 2.0 :0}, inplace= True)
Durante['DOR_ABD'].fillna(0,inplace= True)
Durante['FADIGA'].replace({9.0: 0, 2.0 :0}, inplace= True)
Durante['FADIGA'].fillna(0,inplace= True)
Durante['PERD_OLFT'].replace({9.0: 0, 2.0 :0}, inplace= True)
Durante['PERD_OLFT'].fillna(0,inplace= True)
Durante['PERD_PALA'].replace({9.0: 0, 2.0 :0}, inplace= True)
Durante['PERD_PALA'].fillna(0,inplace= True)
Durante['VACINA'].fillna(0,inplace= True)
Durante['FATOR_RISC'].replace({'S': 1, 'N':2, '1':1, '2':2}, inplace= True)
Durante['FATOR_RISC'].fillna(0,inplace= True)
# - Resetando o Index novamente.
Durante=Durante.reset_index(drop=True)
Durante.head()
# - Aplicação da Dummy nas Features Categóricas
Durante=pd.get_dummies(Durante, columns=['CS_SEXO', 'CS_GESTANT', 'CS_RACA', 'CS_ESCOL_N',
'SURTO_SG', 'NOSOCOMIAL', 'FEBRE', 'TOSSE', 'GARGANTA', 'DISPNEIA',
'DESC_RESP', 'SATURACAO', 'DIARREIA', 'VOMITO', 'PUERPERA',
'FATOR_RISC', 'CARDIOPATI', 'HEMATOLOGI', 'SIND_DOWN', 'HEPATICA',
'ASMA', 'DIABETES', 'NEUROLOGIC', 'PNEUMOPATI', 'IMUNODEPRE', 'RENAL',
'OBESIDADE', 'VACINA', 'ANTIVIRAL', 'UTI', 'SUPORT_VEN', 'PCR_RESUL',
'HISTO_VGM', 'DOR_ABD', 'FADIGA', 'PERD_OLFT', 'PERD_PALA'], drop_first=True)
Durante.head()
# # Verificando o Balanceamento
Durante["EVOLUCAO"].value_counts(normalize=True)
# +
X = Durante[['IDADE_ANOS','CS_SEXO_M','CS_RACA_4.0','FEBRE_1.0','DISPNEIA_1.0','SATURACAO_1.0','UTI_1.0',
'SUPORT_VEN_1.0', 'SUPORT_VEN_2.0', 'PCR_RESUL_2.0','TOSSE_1.0','DESC_RESP_1.0', 'FATOR_RISC_2']]
y = Durante['EVOLUCAO']
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, test_size=0.3, random_state=42)
# -
Xtrain.shape, Xtest.shape, ytrain.shape, ytest.shape
Xtrain.columns
# +
smote = SMOTE(sampling_strategy = 'minority', random_state = 42)
Xtrain_over, ytrain_over = smote.fit_resample(Xtrain,ytrain)
Xtest_over, ytest_over = smote.fit_resample(Xtest,ytest)
Xtrain_over.shape, ytrain_over.shape, Xtest_over.shape, ytest_over.shape
# -
# # Aplicação do Modelo Escolhido
random_state=42
BCG=BaggingClassifier()
BCG.fit(Xtrain_over, ytrain_over)
previsoes = BCG.predict(Xtest_over)
previsoes
accuracy_score(ytest_over, previsoes)
# +
# Testar Modelo
idade = 43.0
sexo = 1
raca = 0
febre = 1
dispneia = 1
saturacao = 0
uti = 1
suport1 = 1
suport2 = 0
pcr = 1
tosse = 1
descresp = 0
frisc = 0
prediction = BCG.predict(np.array([idade, sexo, raca, febre, dispneia, saturacao, uti, suport1, suport2, pcr, tosse, descresp, frisc]).reshape(1, -1))
print(prediction)
| Notebooks cidades/Guarulhos_Durante.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="xTCDJCXJgjL8" colab_type="text"
# #RUN NN
# + id="Suu0L7jqUwFp" colab_type="code" outputId="8a4c128a-f029-404c-fdd0-7677c59aad66" colab={"base_uri": "https://localhost:8080/", "height": 367}
# -*- coding: utf-8 -*-
"""
Created on Sat Dec 8 22:05:38 2018
@author: Chandar_S
"""
from cnn import cnn
from fnn import fnn
from rnn import rnn
from nn_utilities_py import nn_utilities
import tensorflow as tf
from scipy.misc import imread
import os
import numpy as np
import pylab
import matplotlib.pyplot as plt
import sys
data_path = './'
nn_utilities_obj = nn_utilities(data_path)
def run_test():
nn_utilities_obj.load_PneumothoraxDataset()
# nn_utilities_obj.load_fashion_data()
# nn_utilities_obj.load_mnist_digit_data()
# nn_utilities_obj.prepare_digits_image_inputs()
def run_fnn():
fnn_obj = fnn(data_path)
# Flag makes it run with new simplified code and does not run validation accuracy for quicker response
legacy_run = False
## GET INPUT DATA
# input_data = nn_utilities_obj.prepare_digits_image_inputs()
input_data = nn_utilities_obj.load_mnist_digit_data()
# input_data = nn_utilities_obj.load_fashion_data()
## 2 LAYER FNN INPUTS
hiddenlayer_1_width = 256
hiddenlayer_2_width = 256
## Override the default learning rate
fnn_obj.learning_rate_var = 0.001
if legacy_run == True:
## CREATE FNN MODEL
optimizer, cost, accuracy, fnn_model = fnn_obj.create_model(fnn_obj.x, input_data["x_train"].shape[1], hiddenlayer_1_width, hiddenlayer_2_width, input_data["y_train"].shape[1])
else:
## CREATE FNN MODEL
optimizer, cost, accuracy, fnn_model = fnn_obj.create_simplified_model(fnn_obj.x, input_data["x_train"].shape[1], hiddenlayer_1_width, hiddenlayer_2_width, input_data["y_train"].shape[1] )
## TRAIN THE MODEL AND TEST PREDICTION
run_nn(fnn_obj, input_data, optimizer, cost, accuracy, fnn_model, "fnn_"+input_data["name"])
def run_cnn():
cnn_obj = cnn(data_path)
# Flag makes it run with new simplified code and does not run validation accuracy for quicker response
legacy_run = False
''' WE NEED THIS FOR LOOKING AT HEAT MAP OVER IMAGE'''
single_layer_fnn = True
## Override the default learning rate
cnn_obj.learning_rate_var = 0.0001
## GET INPUT DATA
# input_data = nn_utilities_obj.prepare_digits_image_inputs()
# input_data = nn_utilities_obj.load_mnist_digit_data()
# input_data = nn_utilities_obj.load_fashion_data()
input_data = nn_utilities_obj.load_PneumothoraxDataset()
## 2 LAYER FNN INPUTS
hiddenlayer_1_width = 500
hiddenlayer_2_width = 500
## Assuming it's a SQUARE IMAGE
image_height = int(np.sqrt(input_data["x_train"].shape[1]))
image_width = image_height
if legacy_run == True:
## CREATE CNN & DNN MODEL
optimizer, cost, accuracy, cnn_fnn_model = cnn_obj.create_model([image_height, image_width], hiddenlayer_1_width, hiddenlayer_2_width, input_data["y_train"].shape[1], single_layer_fnn)
else:
## CREATE CNN & DNN MODEL
optimizer, cost, accuracy, cnn_fnn_model = cnn_obj.create_simplified_model([image_height, image_width], hiddenlayer_1_width, hiddenlayer_2_width, input_data["y_train"].shape[1], single_layer_fnn)
## TRAIN THE MODEL AND TEST PREDICTION
run_nn(cnn_obj, input_data, optimizer, cost, accuracy, cnn_fnn_model, "cnn_"+input_data["name"], False)
def run_rnn():
rnn_obj = rnn(data_path)
## GET INPUT DATA
# input_data = nn_utilities_obj.prepare_digits_image_inputs()
input_data = nn_utilities_obj.load_fashion_data()
## Override the default learning rate
rnn_obj.learning_rate_var = 0.05
## Assuming it's a SQUARE IMAGE
image_height = int(np.sqrt(input_data["x_train"].shape[1]))
image_width = image_height
# Network Parameters
num_input = image_height # MNIST data input (img shape: 28*28)
timesteps = image_width # timesteps
num_hidden = 128 # hidden layer num of features
num_classes = 10 # MNIST total classes (0-9 digits)
## CREATE RNN MODEL
optimizer, cost, accuracy, rnn_model = rnn_obj.create_model(num_input, timesteps, num_hidden, num_classes)
input_data["x_train"] = np.reshape(input_data["x_train"],[input_data["x_train"].shape[0], timesteps,num_input])
input_data["x_validation"] = np.reshape(input_data["x_validation"],[input_data["x_validation"].shape[0], timesteps,num_input])
## TRAIN THE MODEL AND TEST PREDICTION
run_nn(rnn_obj, input_data, optimizer, cost, accuracy, rnn_model, "rnn_"+input_data["name"])
def run_nn(obj, input_data, optimizer, cost, accuracy, model, model_name=None, run_validation_accuracy=True):
# Python optimisation variables
training_epochs = 20
display_step = 100
batch_size = 100
quick_training = False
print ("Starting session")
#### TRAIN AND TEST NN
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
# TRAIN
trained_model = obj.train_model(sess, model, training_epochs, display_step, batch_size, optimizer, cost, accuracy, input_data["x_train"], input_data["x_train_4D"], input_data["y_train"], input_data["x_validation"], input_data["y_validation"], quick_training, model_name, run_validation_accuracy)
## TEST
test = input_data["test"]
if (test is not None):
data_dir = input_data["data_dir"]
img_name = obj.rng.choice(test.filename)
filepath = os.path.join(data_dir, 'Numbers', 'Images', 'test', img_name)
img = imread(filepath, flatten=True)
# convert list to ndarray and PREP AS PER INPUT FORMAT
x_test = np.stack(img)
if len(input_data["x_train"].shape) == 2:
x_test = x_test.reshape(-1, input_data["x_train"].shape[1])
else:
x_test = x_test.reshape(-1, input_data["x_train"].shape[1], input_data["x_train"].shape[2])
## PREDICT AND VALIDATE
predicted_test = obj.predictvalue(trained_model, x_test)
print("Prediction is: ", predicted_test[0])
pylab.imshow(img, cmap='gray')
pylab.axis('off')
pylab.show()
print ("Ending session")
## DO MIT CAM Analysis to print the Heatmap
CAM_analysis = True
if (CAM_analysis == True):
load_saved_model(model_name, obj, input_data)
def load_saved_model(model_name, obj, input_data):
with tf.Session() as sess:
saver = tf.train.Saver()
print ("Restoring Model")
saver.restore(sess, data_path + ""+model_name+".ckpt")
print ("Starting with CAM Analysis")
"""DOING CAM Heatmaps Analysis"""
'''extract the features and weights using the function defined directly above '''
(feature_maps, dense_weights) = extract_features_weights(sess, obj) #TODO
# print("Feature Maps: "+str(feature_maps))
# print("Dense Weights: "+str(dense_weights))
'''TODO: compute the CAM for a pneumothorax detection using the function above'''
WHICH_OPTION_INDEX = 1
cam = compute_cam(WHICH_OPTION_INDEX, feature_maps, dense_weights)
## Assuming it's a SQUARE IMAGE
image_height = int(np.sqrt(input_data["x_train"].shape[1]))
image_width = image_height
''' upsample the CAM Tensor to a 28\times 28 image '''
cam_upsampled = tf.image.resize_bilinear(cam, [image_height,image_width])
inds = []
for check_index in range (1,20):
if np.argmax(input_data["y_validation"][check_index]) == WHICH_OPTION_INDEX:
inds.extend([check_index])
print (inds)
# inds= [79, 31]
input_data["y_validation"] = np.stack(input_data["y_validation"])
# print (type(input_data["x_validation"][1]))
# print (input_data["y_validation"][1])
for im, cl in zip(input_data["x_validation"][inds], input_data["y_validation"][inds]):
heatmap = sess.run(
cam_upsampled,
feed_dict={
obj.x: im[np.newaxis,:],
})
vis_cam(im, np.squeeze(heatmap), input_data)
"""DOING CAM Heatmaps Analysis"""
''' Extract the last Layer weights of CNN and FNN for CAM manipulation'''
def extract_features_weights(sess, cnn_obj):
#access feature map activations directly from the model declaration
feature_maps = cnn_obj.cnn_output
# graph = tf.get_default_graph()
# for op in graph.get_operations():
# print(op.name)
# we have implemented 2 different methods, so handling both scenarios
try:
#access the weights by searching by name
dense_weights = sess.graph.get_tensor_by_name('fnn/FNN_Output_Weight:0')
except:
#access the weights by searching by name
dense_weights = sess.graph.get_tensor_by_name('dense_layer/kernel:0')
return (feature_maps, dense_weights)
''' Forms a CAM operation given a class name, feature maps, and weights
Params:
- class_index: index of the class to measure
- fmap: (1 x h x w x d) tf.Tensor of activations from the final convolutional layer
- weights: (features x #ofoutputclasses) tf.Tensor with the learned weights of the final FC layer
Returns:
- (16 x 16) tf.Tensor of downscaled CAMs
'''
def compute_cam(class_index, fmap, weights):
w_vec = tf.expand_dims(weights[:, class_index], 1)
_, h, w, c = fmap.shape.as_list()
fmap = tf.squeeze(fmap) # remove batch dim
fmap = tf.reshape(fmap, [h * w, c])
# compute the CAM! Remeber to look at the equation defining CAMs above to do this
CAM = tf.matmul(fmap, w_vec) # TODO
CAM = tf.reshape(CAM, [1, h, w, 1])
return CAM
""" Visualize class activation heatmap, overlaying on image."""
def vis_cam(image, cam, input_data, save_file=None):
# print (cam)
if (cam.min() != cam.max()):
cam = (cam - cam.min()) / (cam.max() - cam.min()) # TODO: check
## Assuming it's a SQUARE IMAGE
image_height = int(np.sqrt(input_data["x_train"].shape[1]))
image_width = image_height
image = image.reshape(image_height, image_width, 1 )
plt.imshow(255-image.squeeze(), cmap=plt.cm.gray)
plt.imshow(1-cam, cmap=plt.cm.jet, alpha=0.5, interpolation='nearest', vmin=0, vmax=1)
if save_file:
plt.savefig(save_file)
plt.show()
plt.close()
if __name__ == "__main__":
code = 'cnn'
if (code == "cnn"):
print ("Running CNN model")
run_cnn()
elif (code == "fnn"):
print ("Running FNN model")
run_fnn()
elif (code == "rnn"):
print ("Running RNN model")
run_rnn()
elif (code == "test"):
print ("Running Test")
run_test()
# + [markdown] id="YlqxYJtKXXDz" colab_type="text"
#
| DeepLearning/DeepNetworks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from os import path
from wordcloud import WordCloud
import matplotlib.pyplot as plt
#
# Read in job descriptions 1-5
#
f = open('data/job1.txt','r')
job1 = f.read().lower()
f = open('data/job2.txt','r')
job2 = f.read().lower()
f = open('data/job3.txt','r')
job3 = f.read().lower()
f = open('data/job4.txt','r')
job4 = f.read().lower()
f = open('data/job5.txt','r')
job5 = f.read().lower()
#
# Concatenate the data.
#
descriptions = job1 + job2 + job3 + job4 + job5
# +
#
# Generate a category word cloud.
#
wordcloud = WordCloud(max_words=75, width=1024, height=512, background_color="white", colormap="Blues").generate_from_text(descriptions)
#
# Display the generated image
#
plt.figure(figsize=(15,10))
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.margins(x=0, y=0)
plt.show()
| jobs/job-desc-wordcloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import codecs
file = codecs.open("SastaSundar.html", "r", "utf-8")
#print(file.read())
with open("filename.html","w+") as f:
f.write("SastaSundar.html")
infile = open('SastaSundar.html', 'r')
img_list = []
img_count = 0
for line in infile:
if "img" in line:
img_list.append(line+"\n")
img_count = img_count + 1
#print(line+"\n")
print(img_list)
print(img_count)
# +
import requests
import sys
from bs4 import BeautifulSoup
from urllib.parse import urlparse
from urllib.parse import urljoin
searched_links = []
broken_links = []
infile = open('SastaSundar.html', 'r')
URL = "https://www.sastasundar.com/"
def getLinksFromHTML(infile):
def getLink(el):
return el["href"]
return list(map(getLink, BeautifulSoup(infile, features="html.parser").select("a[href]")))
def find_broken_links(domainToSearch, URL, parentURL):
if (not (URL in searched_links)) and (not URL.startswith("mailto:")) and (not ("javascript:" in URL)) and (not URL.endswith(".png")) and (not URL.endswith(".jpg")) and (not URL.endswith(".jpeg")):
try:
requestObj = requests.get(URL);
searched_links.append(URL)
if(requestObj.status_code == 404):
broken_links.append("BROKEN: link " + URL + " from " + parentURL)
print(broken_links[-1])
else:
print("NOT BROKEN: link " + URL + " from " + parentURL)
if urlparse(URL).netloc == domainToSearch:
for link in getLinksFromHTML(requestObj.text):
find_broken_links(domainToSearch, urljoin(URL, link), URL)
except Exception as e:
print("ERROR: " + str(e));
searched_links.append(domainToSearch)
find_broken_links(urlparse(sys.argv[1]).netloc, sys.argv[1], "")
print("\n--- DONE! ---\n")
print("The following links were broken:")
for link in broken_links:
print ("\t" + link)
# -
from bs4 import BeautifulSoup
#import urllib2
import re
count1 = 0
#html_page = urllib2.urlopen("http://imgur.com")
soup = BeautifulSoup(infile)
images = []
for img in soup.findAll('img'):
images.append(img.get('src'))
count1 = count1 + 1
print(images)
print(count1)
| Python Web Scrapping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 07: Creating Custom Environments
#
# This tutorial walks you through the process of creating custom environments in Flow. Custom environments contain specific methods that define the problem space of a task, such as the state and action spaces of the RL agent and the signal (or reward) that the RL algorithm will optimize over. By specifying a few methods within a custom environment, individuals can use Flow to design traffic control tasks of various types, such as optimal traffic light signal timing and flow regulation via mixed autonomy traffic (see the figures below). Finally, these environments are compatible with OpenAI Gym.
#
# The rest of the tutorial is organized as follows: in section 1 walks through the process of creating an environment for mixed autonomy vehicle control where the autonomous vehicles perceive all vehicles in the network, and section two implements the environment in simulation.
#
# <img src="img/sample_envs.png">
#
#
# ## 1. Creating an Environment Class
#
# In this exercise we will create an environment in which the accelerations of a handful of vehicles in the network are specified by a single centralized agent, with the objective of the agent being to improve the average speed of all vehicle in the network. In order to create this environment, we begin by inheriting the base environment class located in *flow.envs*:
# +
# import the base environment class
from flow.envs import Env
# define the environment class, and inherit properties from the base environment class
class myEnv(Env):
pass
# -
# `Env` provides the interface for running and modifying a SUMO simulation. Using this class, we are able to start sumo, provide a scenario to specify a configuration and controllers, perform simulation steps, and reset the simulation to an initial configuration.
#
# By inheriting Flow's base environment, a custom environment for varying control tasks can be created by adding the following functions to the child class:
# * **action_space**
# * **observation_space**
# * **apply_rl_actions**
# * **get_state**
# * **compute_reward**
#
# Each of these components are covered in the next few subsections.
#
# ### 1.1 ADDITIONAL_ENV_PARAMS
#
# The features used to parametrize components of the state/action space as well as the reward function are specified within the `EnvParams` input, as discussed in tutorial 1. Specifically, for the sake of our environment, the `additional_params` attribute within `EnvParams` will be responsible for storing information on the maximum possible accelerations and decelerations by the autonomous vehicles in the network. Accordingly, for this problem, we define an `ADDITIONAL_ENV_PARAMS` variable of the form:
ADDITIONAL_ENV_PARAMS = {
"max_accel": 1,
"max_decel": 1,
}
# All environments presented in Flow provide a unique `ADDITIONAL_ENV_PARAMS` component containing the information needed to properly define some environment-specific parameters. We assume that these values are always provided by the user, and accordingly can be called from `env_params`. For example, if we would like to call the "max_accel" parameter, we simply type:
#
# max_accel = env_params.additional_params["max_accel"]
#
# ### 1.2 action_space
#
# The `action_space` method defines the number and bounds of the actions provided by the RL agent. In order to define these bounds with an OpenAI gym setting, we use several objects located within *gym.spaces*. For instance, the `Box` object is used to define a bounded array of values in $\mathbb{R}^n$.
from gym.spaces.box import Box
# In addition, `Tuple` objects (not used by this exercise) allow users to combine multiple `Box` elements together.
from gym.spaces.tuple_space import Tuple
# Once we have imported the above objects, we are ready to define the bounds of our action space. Given that our actions consist of a list of n real numbers (where n is the number of autonomous vehicles) bounded from above and below by "max_accel" and "max_decel" respectively (see section 1.1), we can define our action space as follows:
class myEnv(myEnv):
@property
def action_space(self):
num_actions = self.scenario.vehicles.num_rl_vehicles
accel_ub = self.env_params.additional_params["max_accel"]
accel_lb = - abs(self.env_params.additional_params["max_decel"])
return Box(low=accel_lb,
high=accel_ub,
shape=(num_actions,))
# ### 1.3 observation_space
# The observation space of an environment represents the number and types of observations that are provided to the reinforcement learning agent. For this example, we will be observe two values for each vehicle: its position and speed. Accordingly, we need a observation space that is twice the size of the number of vehicles in the network.
class myEnv(myEnv): # update my environment class
@property
def observation_space(self):
return Box(
low=0,
high=float("inf"),
shape=(2*self.scenario.vehicles.num_vehicles,),
)
# ### 1.4 apply_rl_actions
# The function `apply_rl_actions` is responsible for transforming commands specified by the RL agent into actual actions performed within the simulator. The vehicle kernel within the environment class contains several helper methods that may be of used to facilitate this process. These functions include:
# * **apply_acceleration** (list of str, list of float) -> None: converts an action, or a list of actions, into accelerations to the specified vehicles (in simulation)
# * **apply_lane_change** (list of str, list of {-1, 0, 1}) -> None: converts an action, or a list of actions, into lane change directions for the specified vehicles (in simulation)
# * **choose_route** (list of str, list of list of str) -> None: converts an action, or a list of actions, into rerouting commands for the specified vehicles (in simulation)
#
# For our example we consider a situation where the RL agent can only specify accelerations for the RL vehicles; accordingly, the actuation method for the RL agent is defined as follows:
class myEnv(myEnv): # update my environment class
def _apply_rl_actions(self, rl_actions):
# the names of all autonomous (RL) vehicles in the network
rl_ids = self.k.vehicle.get_rl_ids()
# use the base environment method to convert actions into accelerations for the rl vehicles
self.k.vehicle.apply_acceleration(rl_ids, rl_actions)
# ### 1.5 get_state
#
# The `get_state` method extracts features from within the environments and provides then as inputs to the policy provided by the RL agent. Several helper methods exist within flow to help facilitate this process. Some useful helper method can be accessed from the following objects:
# * **self.k.vehicle**: provides current state information for all vehicles within the network
# * **self.k.traffic_light**: provides state information on the traffic lights
# * **self.k.scenario**: information on the scenario, which unlike the vehicles and traffic lights is static
# * More accessor objects and methods can be found within the Flow documentation at: http://berkeleyflow.readthedocs.io/en/latest/
#
# In order to model global observability within the network, our state space consists of the speeds and positions of all vehicles (as mentioned in section 1.3). This is implemented as follows:
# +
import numpy as np
class myEnv(myEnv): # update my environment class
def get_state(self, **kwargs):
# the get_ids() method is used to get the names of all vehicles in the network
ids = self.k.vehicle.get_ids()
# we use the get_absolute_position method to get the positions of all vehicles
pos = [self.k.vehicle.get_x_by_id(veh_id) for veh_id in ids]
# we use the get_speed method to get the velocities of all vehicles
vel = [self.k.vehicle.get_speed(veh_id) for veh_id in ids]
# the speeds and positions are concatenated to produce the state
return np.concatenate((pos, vel))
# -
# ### 1.6 compute_reward
#
# The `compute_reward` method returns the reward associated with any given state. These value may encompass returns from values within the state space (defined in section 1.5) or may contain information provided by the environment but not immediately available within the state, as is the case in partially observable tasks (or POMDPs).
#
# For this exercise, we choose the reward function to be the average speed of all vehicles currently in the network. In order to extract this information from the environment, we use the `get_speed` method within the Vehicle kernel class to collect the current speed of all vehicles in the network, and return the average of these speeds as the reward. This is done as follows:
# +
import numpy as np
class myEnv(myEnv): # update my environment class
def compute_reward(self, rl_actions, **kwargs):
# the get_ids() method is used to get the names of all vehicles in the network
ids = self.k.vehicle.get_ids()
# we next get a list of the speeds of all vehicles in the network
speeds = self.k.vehicle.get_speed(ids)
# finally, we return the average of all these speeds as the reward
return np.mean(speeds)
# -
# ## 2. Testing the New Environment
#
#
# ### 2.1 Testing in Simulation
# Now that we have successfully created our new environment, we are ready to test this environment in simulation. We begin by running this environment in a non-RL based simulation. The return provided at the end of the simulation is indicative of the cumulative expected reward when jam-like behavior exists within the netowrk.
# +
from flow.controllers import IDMController, ContinuousRouter
from flow.core.experiment import Experiment
from flow.core.params import SumoParams, EnvParams, \
InitialConfig, NetParams
from flow.core.params import VehicleParams
from flow.scenarios.loop import LoopScenario, ADDITIONAL_NET_PARAMS
sumo_params = SumoParams(sim_step=0.1, render=True)
vehicles = VehicleParams()
vehicles.add(veh_id="idm",
acceleration_controller=(IDMController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=22)
env_params = EnvParams(additional_params=ADDITIONAL_ENV_PARAMS)
additional_net_params = ADDITIONAL_NET_PARAMS.copy()
net_params = NetParams(additional_params=additional_net_params)
initial_config = InitialConfig(bunching=20)
scenario = LoopScenario(name="sugiyama",
vehicles=vehicles,
net_params=net_params,
initial_config=initial_config)
#############################################################
######## using my new environment for the simulation ########
#############################################################
env = myEnv(env_params, sumo_params, scenario)
#############################################################
exp = Experiment(env)
_ = exp.run(1, 1500)
# -
# ### 2.2 Training the New Environment
#
# Next, we wish to train this environment in the presence of the autonomous vehicle agent to reduce the formation of waves in the network, thereby pushing the performance of vehicles in the network past the above expected return.
#
# In order for an environment to be trainable in either RLLib for rllab (as we have shown in tutorials 2 and 3), the environment must be acccessable via import from *flow.envs*. In order to do so, copy the above envrionment onto a .py and import the environment in `flow.envs.__init__.py`. You can ensure that the process was successful by running the following command:
# NOTE: only runs if the above procedure have been performed
from flow.envs import myEnv
# Once this is done, the below code block may be used to train the above environment using the Trust Region Policy Optimization (TRPO) algorithm provided by rllab. We do not recommend training this environment to completion within a jupyter notebook setting; however, once training is complete, visualization of the resulting policy should show that the autonomous vehicle learns to dissipate the formation and propagation of waves in the network.
# +
from rllab.envs.normalized_env import normalize
from rllab.misc.instrument import run_experiment_lite
from rllab.algos.trpo import TRPO
from rllab.baselines.linear_feature_baseline import LinearFeatureBaseline
from rllab.policies.gaussian_gru_policy import GaussianGRUPolicy
from flow.scenarios.loop import LoopScenario
from flow.controllers import RLController, IDMController, ContinuousRouter
from flow.core.params import VehicleParams
from flow.core.params import SumoParams, EnvParams, NetParams, InitialConfig
from rllab.envs.gym_env import GymEnv
HORIZON = 1500
def run_task(*_):
sumo_params = SumoParams(sim_step=0.1, render=False)
vehicles = VehicleParams()
vehicles.add(veh_id="rl",
acceleration_controller=(RLController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=1)
vehicles.add(veh_id="idm",
acceleration_controller=(IDMController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=21)
env_params = EnvParams(horizon=HORIZON,
additional_params=ADDITIONAL_ENV_PARAMS)
additional_net_params = ADDITIONAL_NET_PARAMS.copy()
net_params = NetParams(additional_params=additional_net_params)
initial_config = InitialConfig(bunching=20)
scenario = LoopScenario(name="sugiyama-training",
vehicles=vehicles,
net_params=net_params,
initial_config=initial_config)
#######################################################
######## using my new environment for training ########
#######################################################
env_name = "myEnv"
#######################################################
pass_params = (env_name, sumo_params, vehicles, env_params, net_params,
initial_config, scenario)
env = GymEnv(env_name, record_video=False, register_params=pass_params)
horizon = env.horizon
env = normalize(env)
policy = GaussianGRUPolicy(
env_spec=env.spec,
hidden_sizes=(5,),
)
baseline = LinearFeatureBaseline(env_spec=env.spec)
algo = TRPO(
env=env,
policy=policy,
baseline=baseline,
batch_size=30000,
max_path_length=horizon,
n_itr=500,
discount=0.999,
)
algo.train(),
exp_tag = "stabilizing-the-ring"
for seed in [5]: # , 20, 68]:
run_experiment_lite(
run_task,
n_parallel=1,
snapshot_mode="all",
seed=seed,
mode="local",
exp_prefix=exp_tag,
)
# -
| tutorials/tutorial07_environments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Hc0XEe7S6Ft4" colab_type="text"
# # Seq2Seq with Attention for German-English Neural Machine Translation
# - Network architecture based on this [paper](https://arxiv.org/abs/1409.0473)
# - Fit to run on Google Colaboratory
# + id="jCfquy219jiJ" colab_type="code" colab={}
import os
import io
import math
import time
import random
import spacy
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torchtext.data import Field, BucketIterator
from torchtext.datasets import TranslationDataset, Multi30k
# + [markdown] id="txr1BWK6dxLW" colab_type="text"
# ## Set random seed
# + id="uAy9-XbGdzp4" colab_type="code" colab={}
SEED = 2015010720
random.seed(SEED)
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
# + [markdown] id="Rk3DQJRdkqmc" colab_type="text"
# ## Check Spacy (이미 설치되어 있음)
# + id="ub_8K6GxkU_V" colab_type="code" outputId="e90a7d5f-358b-431a-ed58-e35d56462247" executionInfo={"status": "ok", "timestamp": 1564688612003, "user_tz": -540, "elapsed": 5555, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 199}
# 설치가 되어있는지 확인
# !pip show spacy
# + id="KIB9ZBpWln-G" colab_type="code" outputId="85a13822-3086-4092-def4-f4736454db39" executionInfo={"status": "ok", "timestamp": 1564688622914, "user_tz": -540, "elapsed": 15056, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 456}
# 설치가 되어있는지 확인 (없다면 자동설치됨)
# !python -m spacy download en
# !python -m spacy download de
# + id="18MJjo21kyG1" colab_type="code" colab={}
import spacy
spacy_en = spacy.load('en')
spacy_de = spacy.load('de')
# + [markdown] id="skY4owhASmK5" colab_type="text"
# # 1. Define Tokenizing Functions
# 문장을 받아 그보다 작은 어절 혹은 형태소 단위의 리스트로 반환해주는 함수를 각 언어에 대해 작성
# + [markdown] id="0R17vYJMT8mq" colab_type="text"
# ## German Tokenizer
# + id="Nh5klj0N0VQz" colab_type="code" colab={}
def tokenize_de(text):
return [t.text for t in spacy_de.tokenizer(text)]
# + id="xH9qI4OH1x0t" colab_type="code" outputId="d3f34177-0477-4e70-dee0-8f22513afd6e" executionInfo={"status": "ok", "timestamp": 1564688628162, "user_tz": -540, "elapsed": 505, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
# Usage example
print(tokenize_de("Was ich nicht erschaffen kann, verstehe ich nicht."))
# + [markdown] id="-HF7H_h_UAXb" colab_type="text"
# ## English Tokenizer
# + id="XoJWSFgLTTjG" colab_type="code" colab={}
def tokenize_en(text):
return [t.text for t in spacy_en.tokenizer(text)]
# + id="CW3hvo5Lf1G3" colab_type="code" outputId="59a7fb55-5f73-4253-b3be-efecb59762f2" executionInfo={"status": "ok", "timestamp": 1564688630539, "user_tz": -540, "elapsed": 621, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
# Usage example
print(tokenize_en("What I cannot create, I don't understand."))
# + [markdown] id="tkprvaRwR2HB" colab_type="text"
# # 2. Data Preprocessing
# + [markdown] id="31Rm1iulT4Wb" colab_type="text"
# ##Define Fields
# + id="XbFJiDz7hGub" colab_type="code" colab={}
GERMAN = Field(
tokenize = tokenize_de,
init_token='<sos>',
eos_token='<eos>',
lower=True,
include_lengths=True,
batch_first=False # time first
)
ENGLISH = Field(
tokenize=tokenize_en,
init_token='<sos>',
eos_token='<eos>',
lower=True,
include_lengths=True,
batch_first=False # time first
)
# + [markdown] id="cxDjYnUZhbHh" colab_type="text"
# ## Load Data
# + id="xusPXahxhee4" colab_type="code" outputId="363c2670-0bf8-47f1-e937-25e637a19dd5" executionInfo={"status": "ok", "timestamp": 1564688683302, "user_tz": -540, "elapsed": 35072, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 126}
train_set, valid_set, test_set = Multi30k.splits(
exts=('.de', '.en'),
fields=(GERMAN, ENGLISH)
)
# + id="XqOQMiMZiiN1" colab_type="code" outputId="7e5b54b1-9dfc-4c01-bb3a-f5ba0fb5ed43" executionInfo={"status": "ok", "timestamp": 1564688694302, "user_tz": -540, "elapsed": 605, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 72}
print('#. train examples:', len(train_set.examples))
print('#. valid examples:', len(valid_set.examples))
print('#. test examples:', len(test_set.examples))
# + id="q8frH5waRAfa" colab_type="code" outputId="794d2289-c4e9-4c40-bfcc-3aec6916f10d" executionInfo={"status": "ok", "timestamp": 1564688699271, "user_tz": -540, "elapsed": 603, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 345}
# Training example (GERMAN, source language)
train_set.examples[50].src
# + id="1OukZ7MqRCff" colab_type="code" outputId="75f743ac-2795-430d-f9ed-e44d00aa98a7" executionInfo={"status": "ok", "timestamp": 1564688701241, "user_tz": -540, "elapsed": 613, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 290}
# Training example (ENGLISH, target language)
train_set.examples[50].trg
# + [markdown] id="g9mfm9nyzA8f" colab_type="text"
# ## Build Vocabulary
# - 각 언어별 생성: `Field`의 인스턴스를 활용
# - 최소 빈도수(`MIN_FREQ`) 값을 작게 하면 vocabulary의 크기가 커짐.
# - 최소 빈도수(`MIN_FREQ`) 값을 크게 하면 vocabulary의 크기가 작아짐.
#
# + id="tqTH_3EelkFC" colab_type="code" colab={}
MIN_FREQ = 2 # TODO: try different values
# + [markdown] id="kmovdbPipDHh" colab_type="text"
# ### German vocab
# + id="gB3ssFibRLOi" colab_type="code" outputId="e2434ece-632f-4b14-f0c9-0d6306042b84" executionInfo={"status": "ok", "timestamp": 1564688711101, "user_tz": -540, "elapsed": 613, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
# Build vocab for German
GERMAN.build_vocab(train_set, min_freq=MIN_FREQ) # de
print('Size of source vocab (de):', len(GERMAN.vocab))
# + id="vJC9V0C9WbY0" colab_type="code" outputId="7f01440e-aca9-4ee6-e385-4b4fdab4a443" executionInfo={"status": "ok", "timestamp": 1564688712867, "user_tz": -540, "elapsed": 594, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 199}
GERMAN.vocab.freqs.most_common(10)
# + id="myIIOwR29spe" colab_type="code" outputId="39fc6dba-3b36-491e-e08c-505fdf811f6e" executionInfo={"status": "ok", "timestamp": 1564688715159, "user_tz": -540, "elapsed": 656, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 90}
# Check indices of some important tokens
tokens = ['<unk>', '<pad>', '<sos>', '<eos>']
for token in tokens:
print(f"{token} -> {GERMAN.vocab.stoi[token]}")
# + [markdown] id="V8gBAHlqpFws" colab_type="text"
# ### English vocab
# + id="YYV-xvYnzL-k" colab_type="code" outputId="c4750496-8f64-4172-e26e-e44ee634e3b9" executionInfo={"status": "ok", "timestamp": 1564688717421, "user_tz": -540, "elapsed": 720, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
# Build vocab for English
ENGLISH.build_vocab(train_set, min_freq=MIN_FREQ) # en
print('Size of target vocab (en):', len(ENGLISH.vocab))
# + id="r5zgO9adWmzR" colab_type="code" outputId="78a45592-646e-47d0-d113-6781f8167e9d" executionInfo={"status": "ok", "timestamp": 1564688717421, "user_tz": -540, "elapsed": 534, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 199}
ENGLISH.vocab.freqs.most_common(10)
# + id="ygH06w7p-XFS" colab_type="code" outputId="08b7bf60-d401-4ebe-f22f-455f9cf5cac4" executionInfo={"status": "ok", "timestamp": 1564688717673, "user_tz": -540, "elapsed": 557, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 90}
# Check indices of some important tokens
tokens = ['<unk>', '<pad>', '<sos>', '<eos>']
for token in tokens:
print(f"{token} -> {ENGLISH.vocab.stoi[token]}")
# + [markdown] id="rGZS2-xBz8I-" colab_type="text"
# ## Configure Device
# - *'런타임' -> '런타임 유형변경'* 에서 하드웨어 가속기로 **GPU** 선택
# + id="39RfXwWszTi8" colab_type="code" outputId="f3f264f1-55e5-4b32-bd64-1949832cc2af" executionInfo={"status": "ok", "timestamp": 1564688721452, "user_tz": -540, "elapsed": 742, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print('Device to use:', device)
# + [markdown] id="VGVUyRY_0f7s" colab_type="text"
# ## Create Data Iterators
# - 데이터를 미니배치(mini-batch) 단위로 반환해주는 역할
# - `train_set`, `dev_set`, `test_set`에 대해 개별적으로 정의해야 함
# - `BATCH_SIZE`를 정의해주어야 함
# - `torchtext.data.BucketIterator`는 하나의 미니배치를 서로 비슷한 길이의 관측치들로 구성함
# - [Bucketing](https://medium.com/@rashmi.margani/how-to-speed-up-the-training-of-the-sequence-model-using-bucketing-techniques-9e302b0fd976)의 효과: 하나의 미니배치 내 padding을 최소화하여 연산의 낭비를 줄여줌
#
# + id="ZmBkViElnEAj" colab_type="code" colab={}
BATCH_SIZE = 128
# + id="PoX4TwE5m6si" colab_type="code" outputId="296aaacc-e68a-4978-fad7-abbccf56c261" executionInfo={"status": "ok", "timestamp": 1564688733880, "user_tz": -540, "elapsed": 601, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
#from torchtext.data import BucketIterator
# Train iterator
train_iterator = BucketIterator(
train_set,
batch_size=BATCH_SIZE,
train=True,
sort_within_batch=True,
sort_key=lambda x: len(x.src),
device=device,
)
print(f'Number of minibatches per epoch: {len(train_iterator)}')
# + id="Pc0B3HS7nBkl" colab_type="code" outputId="a4e3ff69-9909-4893-8e73-46c79a7e4541" executionInfo={"status": "ok", "timestamp": 1564688761412, "user_tz": -540, "elapsed": 591, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
#from torchtext.data import BucketIterator
# Dev iterator
valid_iterator = BucketIterator(
valid_set,
batch_size=BATCH_SIZE,
train=False,
sort_within_batch=True,
sort_key=lambda x: len(x.src),
device=device
)
print(f'Number of minibatches per epoch: {len(valid_iterator)}')
# + id="3YNd4heenVGn" colab_type="code" outputId="a2741088-9d8e-474c-ee78-be96b35683b1" executionInfo={"status": "ok", "timestamp": 1564688767105, "user_tz": -540, "elapsed": 595, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
#from torchtext.data import BucketIterator
# Test iterator
test_iterator = BucketIterator(
test_set,
batch_size=BATCH_SIZE,
train=False,
sort_within_batch=True,
sort_key=lambda x: len(x.src),
device=device
)
print(f'Number of minibatches per epoch: {len(test_iterator)}')
# + id="Ivq4PK8O1qG2" colab_type="code" outputId="96567933-6f67-485e-fff1-bcc2db16d5cf" executionInfo={"status": "ok", "timestamp": 1564688806725, "user_tz": -540, "elapsed": 6538, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 53}
train_batch = next(iter(train_iterator))
src, src_len = train_batch.src
trg, trg_len = train_batch.trg
print('a batch of source examples has shape:', src.size()) # (source_seq_len, batch_size)
print('a batch of target examples has shape:', trg.size()) # (target_seq_len, batch_size)
# + id="2CIJ8fdp8zqw" colab_type="code" outputId="8df9e13a-9bbe-49fd-d229-b83cdf674a74" executionInfo={"status": "ok", "timestamp": 1564688809112, "user_tz": -540, "elapsed": 581, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 144}
src
# + id="mGfoj3T_83dh" colab_type="code" outputId="f7a0e570-7f75-4a37-8597-b7dbf790c96b" executionInfo={"status": "ok", "timestamp": 1564688813019, "user_tz": -540, "elapsed": 663, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 163}
src_len # sorted in descending order
# + id="x3nSjgcq84KS" colab_type="code" outputId="b8b3bac4-42dc-4160-ac78-82dfb285096f" executionInfo={"status": "ok", "timestamp": 1564688814236, "user_tz": -540, "elapsed": 524, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 144}
trg
# + id="LxU0H8X19zdm" colab_type="code" outputId="94183aec-ad43-4057-dfc6-8ca432ff66d4" executionInfo={"status": "ok", "timestamp": 1564688824974, "user_tz": -540, "elapsed": 568, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 163}
trg_len
# + id="otfd7srHBec9" colab_type="code" outputId="7e618536-5c3e-4b81-89ce-7cab2c7d8467" executionInfo={"status": "ok", "timestamp": 1564688934470, "user_tz": -540, "elapsed": 637, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 235}
# Checking last sample in mini-batch (GERMAN, source lang)
src, src_len = train_batch.src
de_indices = src[:, -1]
de_tokens = [GERMAN.vocab.itos[i] for i in de_indices]
for t, i in zip(de_tokens, de_indices):
print(f"{t} ({i})")
del de_indices, de_tokens
# + id="cizBVHogBhbY" colab_type="code" outputId="1baf3399-72b2-4814-a19a-7dd76857bdd3" executionInfo={"status": "ok", "timestamp": 1564688957407, "user_tz": -540, "elapsed": 628, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 381}
# Checking last sample in mini-batch (EN, target lang)
trg, trg_len = train_batch.trg
en_indices = trg[:, -1]
en_tokens = [ENGLISH.vocab.itos[i] for i in en_indices]
for t, i in zip(en_tokens, en_indices):
print(f"{t} ({i})")
del en_indices, en_tokens
# + [markdown] id="E4h6GTTJ1_vs" colab_type="text"
# # 3. Building Seq2Seq Model
# + [markdown] id="k_Eq8AyXMmpH" colab_type="text"
# ## Hyperparameters
# + id="MWed4n21MkGN" colab_type="code" outputId="f81959ed-55f4-431d-c741-80d2c6bffd14" executionInfo={"status": "ok", "timestamp": 1564688968135, "user_tz": -540, "elapsed": 623, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 144}
# Hyperparameters
INPUT_DIM = len(GERMAN.vocab)
OUTPUT_DIM = len(ENGLISH.vocab)
ENC_EMB_DIM = DEC_EMB_DIM = 256
ENC_HID_DIM = DEC_HID_DIM = 512
USE_BIDIRECTIONAL = False
print('source vocabulary size:', INPUT_DIM)
print('source word embedding size:', ENC_EMB_DIM)
print(f'encoder RNN hidden size: {ENC_HID_DIM} ({ENC_HID_DIM * 2} if bidirectional)')
print('-' * 50)
print('target vocabulary size:', OUTPUT_DIM)
print('target word embedding size:', ENC_EMB_DIM)
print('decoder RNN hidden size:', ENC_HID_DIM)
# + [markdown] id="hWK3vdrW2Io3" colab_type="text"
# ## Encoder
# + id="iPIPAhi01ruC" colab_type="code" colab={}
class Encoder(nn.Module):
"""
Learns an embedding for the source text.
Arguments:
input_dim: int, size of input language vocabulary.
emb_dim: int, size of embedding layer output.
enc_hid_dim: int, size of encoder hidden state.
dec_hid_dim: int, size of decoder hidden state.
bidirectional: uses bidirectional RNNs if True. default is False.
"""
def __init__(self, input_dim, emb_dim, enc_hid_dim, dec_hid_dim, bidirectional=False):
super(Encoder, self).__init__()
self.input_dim = input_dim
self.emb_dim = emb_dim
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.bidirectional = bidirectional
self.embedding = nn.Embedding(
num_embeddings=self.input_dim,
embedding_dim=self.emb_dim
)
self.rnn = nn.GRU(
input_size=self.emb_dim,
hidden_size=self.enc_hid_dim,
bidirectional=self.bidirectional,
batch_first=False
)
self.rnn_output_dim = self.enc_hid_dim
if self.bidirectional:
self.rnn_output_dim *= 2
self.fc = nn.Linear(self.rnn_output_dim, self.dec_hid_dim)
self.dropout = nn.Dropout(.2)
def forward(self, src, src_len):
"""
Arguments:
src: 2d tensor of shape (S, B)
src_len: 1d tensor of shape (B).
Returns:
outputs: 3d tensor of shape (input_seq_len, batch_size, num_directions * enc_h)
hidden: 2d tensor of shape (b, dec_h). This tensor will be used as the initial
hidden state value of the decoder (h0 of decoder).
"""
assert len(src.size()) == 2, 'Input requires dimension (input_seq_len, batch_size).'
# Shape: (b, s, h)
embedded = self.embedding(src)
embedded = self.dropout(embedded)
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded, src_len)
packed_outputs, hidden = self.rnn(packed_embedded)
outputs, _ = nn.utils.rnn.pad_packed_sequence(packed_outputs)
if self.bidirectional:
# (2, b, enc_h) -> (b, 2 * enc_h)
hidden = torch.cat((hidden[-2, :, :], hidden[-1, :, :]), dim=1)
else:
# (1, b, enc_h) -> (b, enc_h)
hidden = hidden.squeeze(0)
# (b, num_directions * enc_h) -> (b, dec_h)
hidden = self.fc(hidden)
hidden = torch.tanh(hidden)
return outputs, hidden # (S, B, enc_h * num_directions), (B, dec_h)
# + [markdown] id="iJtFNEYp-ett" colab_type="text"
# ## Attention
# + id="EdjDNWD1gnS_" colab_type="code" colab={}
class Attention(nn.Module):
def __init__(self, enc_hid_dim, dec_hid_dim, encoder_is_bidirectional=False):
super(Attention, self).__init__()
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.encoder_is_bidirectional = encoder_is_bidirectional
self.attention_input_dim = enc_hid_dim + dec_hid_dim
if self.encoder_is_bidirectional:
self.attention_input_dim += enc_hid_dim # 2 * h_enc + h_dec
self.linear = nn.Linear(self.attention_input_dim, dec_hid_dim)
self.v = nn.Parameter(torch.rand(dec_hid_dim))
def forward(self, hidden, encoder_outputs, mask):
"""
Arguments:
hidden: 2d tensor with shape (batch_size, dec_hid_dim).
encoder_outputs: 3d tensor with shape (input_seq_len, batch_size, enc_hid_dim * num_directions).
mask: 2d tensor with shape(batch_size, input_seq_len)
"""
# Shape check
assert hidden.dim() == 2
assert encoder_outputs.dim() == 3
seq_len, batch_size, _ = encoder_outputs.size()
# (b, dec_h) -> (b, s, dec_h)
hidden = hidden.unsqueeze(1).expand(-1, seq_len, -1)
# (s, b, enc_h * num_directions) -> (b, s, enc_h * num_directions)
encoder_outputs = encoder_outputs.permute(1, 0, 2)
# concat; shape results in (b, s, enc_h + dec_h).
# if encoder is bidirectional, (b, s, 2 * h_enc + h_dec).
concat = torch.cat((hidden, encoder_outputs), dim=2)
# energy; shape is (b, s, dec_h)
energy = torch.tanh(self.linear(concat))
# tile v; (dec_h, ) -> (b, dec_h) -> (b, dec_h, 1)
v = self.v.unsqueeze(0).expand(batch_size, -1).unsqueeze(2)
# attn; (b, s, dec_h) @ (b, dec_h, 1) -> (b, s, 1) -> (b, s)
attn_scores = torch.bmm(energy, v).squeeze(-1)
# mask padding indices
attn_scores = attn_scores.masked_fill(mask == 0, -1e10)
assert attn_scores.dim() == 2 # Final shape check: (b, s)
return F.softmax(attn_scores, dim=1)
# + [markdown] id="6-d9zt0SF9Ff" colab_type="text"
# ## Decoder
# + id="IcG1tbhDC8-1" colab_type="code" colab={}
class Decoder(nn.Module):
"""
Unlike the encoder, a single forward pass of
a `Decoder` instance is defined for only a single timestep.
Arguments:
output_dim: int,
emb_dim: int,
enc_hid_dim: int,
dec_hid_dim: int,
attention_module: torch.nn.Module,
encoder_is_bidirectional: False
"""
def __init__(self, output_dim, emb_dim, enc_hid_dim, dec_hid_dim, attention_module, encoder_is_bidirectional=False):
super(Decoder, self).__init__()
self.emb_dim = emb_dim
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.output_dim = output_dim
self.encoder_is_bidirectional = encoder_is_bidirectional
if isinstance(attention_module, nn.Module):
self.attention_module = attention_module
else:
raise ValueError
self.rnn_input_size = enc_hid_dim + emb_dim # enc_h + dec_emb_dim
if self.encoder_is_bidirectional:
self.rnn_input_size += enc_hid_dim # 2 * enc_h + dec_emb_dim
self.embedding = nn.Embedding(output_dim, emb_dim)
self.rnn = nn.GRU(
input_size=self.rnn_input_size,
hidden_size=dec_hid_dim,
bidirectional=False,
batch_first=False,
)
self.out_input_size = emb_dim + dec_hid_dim + enc_hid_dim
if self.encoder_is_bidirectional:
self.out_input_size += enc_hid_dim
self.out = nn.Linear(self.out_input_size, output_dim)
self.dropout = nn.Dropout(.2)
def forward(self, inp, hidden, encoder_outputs, mask, temperature=1.):
"""
Arguments:
inp: 1d tensor with shape (batch_size, )
hidden: 2d tensor with shape (batch_size, dec_hid_dim).
This `hidden` tensor is the hidden state vector from the previous timestep.
encoder_outputs: 3d tensor with shape (input_seq_len, batch_size, enc_hid_dim * num_directions).
mask: 2d tensor of shape (batch_size, input_seq_len).
"""
assert inp.dim() == 1
assert hidden.dim() == 2
assert encoder_outputs.dim() == 3
# (b, ) -> (1, b)
inp = inp.unsqueeze(0)
# (1, b) -> (1, b, emb)
embedded = self.embedding(inp)
embedded = self.dropout(embedded)
# attention probabilities; (b, s)
attn_probs = self.attention_module(hidden, encoder_outputs, mask)
# (b, 1, s)
attn_probs = attn_probs.unsqueeze(1)
# (s, b, ~) -> (b, s, ~)
encoder_outputs = encoder_outputs.permute(1, 0, 2)
# (b, 1, s) @ (b, s, ~) -> (b, 1, enc_h * num_directions)
weighted = torch.bmm(attn_probs, encoder_outputs)
# (1, b, ~)
weighted = weighted.permute(1, 0, 2)
# (b, 1, emb + enc_h * num_directions)
rnn_input = torch.cat((embedded, weighted), dim=2)
# output; (b, 1, dec_h)
# new_hidden; (1, b, dec_h)
output, new_hidden = self.rnn(rnn_input, hidden.unsqueeze(0))
assert (output == new_hidden).all()
embedded = embedded.squeeze(0) # (1, b, emb) -> (b, emb)
output = output.squeeze(0) # (1, b, dec_h) -> (b, dec_h)
weighted = weighted.squeeze(0) # (1, b, enc_h * num_d) -> (b, enc_h * num_d)
# output; (b, emb + enc_h + dec_h) -> (b, output_dim)
# if encoder is bidirectional, (b, emb + 2 * enc_h + dec_h) -> (b, output_dim)
output = self.out(torch.cat((output, weighted, embedded), dim=1))
output = output / temperature
return output, new_hidden.squeeze(0), attn_probs.squeeze(1)
# + [markdown] id="1FfbPnfr3IgQ" colab_type="text"
# ## Seq2Seq
# + id="in1zWsUvQZdv" colab_type="code" colab={}
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder, pad_idx, sos_idx, eos_idx, device):
super(Seq2Seq, self).__init__()
self.encoder = encoder
self.decoder = decoder
self.pad_idx = pad_idx # 1
self.sos_idx = sos_idx # 2
self.eos_idx = eos_idx # 3
self.device = device
def create_mask(self, src):
mask = (src != self.pad_idx).permute(1, 0) # (b, s)
return mask
def forward(self, src, src_len, trg=None, teacher_forcing_ratio=.5):
batch_size = src.size(1)
max_seq_len = trg.size(0) if trg is not None else 100
trg_vocab_size = self.decoder.output_dim
if trg is None:
assert teacher_forcing_ratio == 0., "Must be zero during inference."
inference = True
trg = torch.zeros(max_seq_len, batch_size).long().fill_(self.sos_idx).to(self.device)
else:
inference = False
# An empty tensor to store decoder outputs (time index first for faster indexing)
outputs_shape = (max_seq_len, batch_size, trg_vocab_size)
outputs = torch.zeros(outputs_shape).to(self.device)
# empty tensor to store attention probs
attns_shape = (max_seq_len, batch_size, src.size(0))
attns = torch.zeros(attns_shape).to(self.device)
encoder_outputs, hidden = self.encoder(src, src_len)
mask = self.create_mask(src)
# first input to the decoder is '<sos>'
# trg; shape (batch_size, seq_len)
initial_dec_input = output = trg[0, :] # get first timestep token
for t in range(1, max_seq_len):
output, hidden, attn = self.decoder(output, hidden, encoder_outputs, mask)
outputs[t] = output # Save output for timestep t, for 1 <= t <= max_len
attns[t] = attn
_, idx = output.max(dim=1)
teacher_force = torch.rand(1).item() < teacher_forcing_ratio
new_dec_input = output = trg[t] if teacher_force else idx
if inference and output.item() == self.eos_idx:
return outputs[:t], attns[:t]
return outputs, attns
# + [markdown] id="AmnGbjjJ3GrO" colab_type="text"
# ## Build Model
# + id="v1lU0L8UOBB7" colab_type="code" outputId="cfbfacc2-4797-4166-b412-0b4c81540e51" executionInfo={"status": "ok", "timestamp": 1564690077641, "user_tz": -540, "elapsed": 802, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 126}
# Define encoder
enc = Encoder(
input_dim=INPUT_DIM,
emb_dim=ENC_EMB_DIM,
enc_hid_dim=ENC_HID_DIM,
dec_hid_dim=DEC_HID_DIM,
bidirectional=USE_BIDIRECTIONAL
)
print(enc)
# + id="ut6eTqZnORIX" colab_type="code" outputId="438f5977-6608-492a-a70f-6eda0056eb96" executionInfo={"status": "ok", "timestamp": 1564690079517, "user_tz": -540, "elapsed": 733, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 72}
# Define attention layer
attn = Attention(
enc_hid_dim=ENC_HID_DIM,
dec_hid_dim=DEC_HID_DIM,
encoder_is_bidirectional=USE_BIDIRECTIONAL
)
print(attn)
# + id="RHmtjt6sOMp6" colab_type="code" outputId="d49d396e-615d-424f-874c-dccc1f810ee2" executionInfo={"status": "ok", "timestamp": 1564690079763, "user_tz": -540, "elapsed": 772, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 181}
# Define decoder
dec = Decoder(
output_dim=OUTPUT_DIM,
emb_dim=DEC_EMB_DIM,
enc_hid_dim=ENC_HID_DIM,
dec_hid_dim=DEC_HID_DIM,
attention_module=attn,
encoder_is_bidirectional=USE_BIDIRECTIONAL
)
print(dec)
# + id="qCFci4L5FLFO" colab_type="code" outputId="f1b01de3-aa38-4b56-f3dd-075cb4b8be79" executionInfo={"status": "ok", "timestamp": 1564690081394, "user_tz": -540, "elapsed": 754, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 72}
PAD_IDX = GERMAN.vocab.stoi['<pad>']
SOS_IDX = ENGLISH.vocab.stoi['<sos>']
EOS_IDX = ENGLISH.vocab.stoi['<eos>']
print('PAD INDEX:', PAD_IDX)
print('SOS INDEX:', SOS_IDX)
print('EOS INDEX:', EOS_IDX)
# + id="pvUGmhhZOP2d" colab_type="code" outputId="a330b765-525f-4cf3-bf55-fa44bd9d8b6b" executionInfo={"status": "ok", "timestamp": 1564690081396, "user_tz": -540, "elapsed": 569, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 326}
model = Seq2Seq(enc, dec, PAD_IDX, SOS_IDX, EOS_IDX, device).to(device)
print(model)
# + [markdown] id="qRQAQp8uX_61" colab_type="text"
# ## Count trainable parameters
# + id="VKaAzVGAOwjw" colab_type="code" outputId="33a196b6-bba7-4855-c157-c0892b640c29" executionInfo={"status": "ok", "timestamp": 1564690083562, "user_tz": -540, "elapsed": 606, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters.')
# + [markdown] id="u8MlgN7eYD00" colab_type="text"
# ## Initialize trainable parameters
# + id="gMWPa_PBYGFa" colab_type="code" outputId="c307955d-f1ce-4668-8a48-fb69da10dfd7" executionInfo={"status": "ok", "timestamp": 1564690085636, "user_tz": -540, "elapsed": 584, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 326}
def init_parameters(model):
for name, param in model.named_parameters():
if 'weight' in name:
nn.init.normal_(param.data, mean=0., std=0.01)
else:
nn.init.constant_(param.data, 0.)
model.apply(init_parameters)
# + [markdown] id="MxqW8sfvPV2z" colab_type="text"
# # 4. Train
# + [markdown] id="Sg3cOfxAPYzV" colab_type="text"
# ## Optimizer
# - Use `optim.Adam` or `optim.RMSprop`.
# + id="J0cJYGriPXc8" colab_type="code" colab={}
optimizer = optim.Adam(model.parameters(), lr=0.001)
#optimizer = optim.RMSprop(model.parameters(), lr=0.01)
# + [markdown] id="wo2OgmdYPjs5" colab_type="text"
# ## Loss function
# + id="zN84sIssPklu" colab_type="code" outputId="6b0de651-5b52-405e-b212-99aff058adde" executionInfo={"status": "ok", "timestamp": 1564690106080, "user_tz": -540, "elapsed": 596, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
criterion = nn.CrossEntropyLoss(ignore_index=PAD_IDX)
print(f"<pad> index in target vocab (en): '{PAD_IDX}' will be ignored when loss is calculated.")
# + [markdown] id="-sdvuh9VP1nC" colab_type="text"
# ## Train function
# + id="N7jbrmD6P2cc" colab_type="code" colab={}
def train(seq2seq_model, iterator, optimizer, criterion, grad_clip=1.0):
seq2seq_model.train()
epoch_loss = .0
for i, batch in enumerate(iterator):
print('.', end='')
src, src_len = batch.src
trg, _ = batch.trg
optimizer.zero_grad()
decoder_outputs, _ = seq2seq_model(src, src_len, trg, teacher_forcing_ratio=.5)
trg_seq_len, batch_size, trg_vocab_size = decoder_outputs.size() # (s, b, trg_vocab)
# (s-1, b, trg_vocab)
decoder_outputs = decoder_outputs[1:]
# (s-1 * b, trg_vocab)
decoder_outputs = decoder_outputs.view(-1, trg_vocab_size)
# (s, b) -> (s-1 * b, )
trg = trg[1:].view(-1)
loss = criterion(decoder_outputs, trg)
loss.backward()
# Gradient clipping; remedy for exploding gradients
torch.nn.utils.clip_grad_norm_(seq2seq_model.parameters(), grad_clip)
optimizer.step()
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# + [markdown] id="3lSnoKXuWS20" colab_type="text"
# ## Evaluate function
# + id="UOlqFlRTSjR1" colab_type="code" colab={}
def evaluate(seq2seq_model, iterator, criterion):
seq2seq_model.eval()
epoch_loss = 0.
with torch.no_grad():
for i, batch in enumerate(iterator):
print('.', end='')
src, src_len = batch.src
trg, _ = batch.trg
decoder_outputs, _ = seq2seq_model(src, src_len, trg, teacher_forcing_ratio=0.)
trg_seq_len, batch_size, trg_vocab_size = decoder_outputs.size() # (s, b, trg_vocab)
# (s-1, b, trg_vocab)
decoder_outputs = decoder_outputs[1:]
# (s-1 * b, trg_vocab)
decoder_outputs = decoder_outputs.view(-1, trg_vocab_size)
# (s, b) -> (s-1 * b, )
trg = trg[1:].view(-1)
loss = criterion(decoder_outputs, trg)
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# + [markdown] id="K4tw3PJUWVUw" colab_type="text"
# ## Epoch time measure function
# + id="2gI5DjKwPQhZ" colab_type="code" colab={}
def epoch_time(start_time, end_time):
"""Returns elapsed time in mins & secs."""
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# + [markdown] id="UtMu_ZQREe1v" colab_type="text"
# ## Train for multiple epochs
# + id="xx4zXJ4JR5lv" colab_type="code" colab={}
NUM_EPOCHS = 10
# + id="kOdqo0ieR9zs" colab_type="code" outputId="804e8ba5-4d02-4b4b-e009-08b0b8bd2ca6" executionInfo={"status": "ok", "timestamp": 1564691052938, "user_tz": -540, "elapsed": 785182, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 947}
best_valid_loss = float('inf')
for epoch in range(NUM_EPOCHS):
start_time = time.time()
train_loss = train(model, train_iterator, optimizer, criterion)
valid_loss = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), './best_model_de2en.pt')
print("\n")
print(f"Epoch: {epoch + 1:>02d} | Time: {epoch_mins}m {epoch_secs}s")
print(f"Train Loss: {train_loss:>.4f} | Train Perplexity: {math.exp(train_loss):7.3f}")
print(f"Valid Loss: {valid_loss:>.4f} | Valid Perplexity: {math.exp(valid_loss):7.3f}")
# + [markdown] id="UcQ-BE01W-c-" colab_type="text"
# ## Save last model (overfitted)
# + id="ig5cGvhwXC-b" colab_type="code" colab={}
torch.save(model.state_dict(), './last_model_de2en.pt')
# + [markdown] id="F6xofPzNVckv" colab_type="text"
# # 5. Test
# + [markdown] id="xgmzL9n34IEd" colab_type="text"
# ## Evaluate on test data
# + id="vYSg7hpx38hF" colab_type="code" outputId="29eeff1a-63b6-4d62-a799-f6a4eb84575d" executionInfo={"status": "ok", "timestamp": 1564691419364, "user_tz": -540, "elapsed": 1635, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
model.load_state_dict(torch.load('best_model_de2en.pt'))
test_loss = evaluate(model, test_iterator, criterion)
print(f'| Test Loss: {test_loss:.3f} | Test PPL: {math.exp(test_loss):7.3f} |')
# + [markdown] id="r-NR8RlmRQGt" colab_type="text"
# ## Function to convert indices to original text strings (translate)
# + id="ezBTAU3CmcI4" colab_type="code" colab={}
def translate_sentence(seq2seq_model, sentence):
seq2seq_model.eval()
# Tokenize sentence
tokenized = tokenize_de(sentence)
# lower tokens
tokenized = [t.lower() for t in tokenized]
# Add <sos> & <eos> tokens to the front and back of the sentence
tokenized = ['<sos>'] + tokenized + ['<eos>']
# tokens -> indices
numericalized = [GERMAN.vocab.stoi[s] for s in tokenized]
sent_length = torch.tensor([len(numericalized)]).long().to(device)
tensor = torch.LongTensor(numericalized).unsqueeze(1).to(device)
translation_logits, attention = seq2seq_model(tensor, sent_length, None, 0.)
translation_tensor = torch.argmax(translation_logits.squeeze(1), dim=1)
translation = [ENGLISH.vocab.itos[i] for i in translation_tensor]
translation, attention = translation[1:], attention[1:]
#assert translation.__len__() == 1 # (sequence_length, )
#assert attention.dim() == 2
return translation, attention
# + id="92FWzU8E3kLo" colab_type="code" colab={}
def display_attention(candidate, translation, attention):
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111)
attention = attention.squeeze(1).cpu().detach().numpy()
cax = ax.matshow(attention, cmap='bone')
ax.tick_params(labelsize=15)
ax.set_xticklabels([''] + ['<sos>'] + [t.lower() for t in tokenize_de(candidate)] + ['<eos>'],
rotation=45)
ax.set_yticklabels([''] + translation)
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
plt.close()
# + id="kGcZZpGCsHEc" colab_type="code" outputId="bedcc89d-c5cf-40e9-c956-263ec969d3dc" executionInfo={"status": "ok", "timestamp": 1564691261734, "user_tz": -540, "elapsed": 624, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 53}
example_idx = 4
src = ' '.join(train_set.examples[example_idx].src)
trg = ' '.join(train_set.examples[example_idx].trg)
print(f'src = {src}')
print(f'trg = {trg}')
# + id="v9QO3FDksmQn" colab_type="code" outputId="d44aacac-3f39-4fe6-a5c9-85b53822bb3d" executionInfo={"status": "ok", "timestamp": 1564691262979, "user_tz": -540, "elapsed": 791, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
translation, attention = translate_sentence(model, src)
print(f'predicted = {translation}')
# + [markdown] id="hqzkjE6g3k9R" colab_type="text"
# ## Alignment
# + id="ZRaSpRma4PAe" colab_type="code" outputId="ca327d51-b92a-49f0-986c-73bbde8b202b" executionInfo={"status": "ok", "timestamp": 1564691481233, "user_tz": -540, "elapsed": 798, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 515}
display_attention(src, translation, attention)
# + id="_wAzQI8e4T7b" colab_type="code" outputId="29a87ffa-5877-4ca9-9364-5342879361bb" executionInfo={"status": "ok", "timestamp": 1564691564246, "user_tz": -540, "elapsed": 983, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 652}
example_idx = 35
src = ' '.join(valid_set.examples[example_idx].src)
trg = ' '.join(valid_set.examples[example_idx].trg)
print(f'src = {src}')
print(f'trg = {trg}')
translation, attention = translate_sentence(model, src)
print(f'predicted trg = {translation}')
display_attention(src, translation, attention)
# + id="hos6p0s64gP2" colab_type="code" outputId="414530ab-0f24-4f1c-bcc9-aa83820896b1" executionInfo={"status": "ok", "timestamp": 1564691593956, "user_tz": -540, "elapsed": 1055, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 576}
example_idx = 7
src = ' '.join(test_set.examples[example_idx].src)
trg = ' '.join(test_set.examples[example_idx].trg)
print(f'src = {src}')
print(f'trg = {trg}')
translation, attention = translate_sentence(model, src)
print(f'predicted trg = {translation}')
display_attention(src, translation, attention)
# + [markdown] id="de77guAiCPgZ" colab_type="text"
# # 6. Download Model
# + id="5yYiqOcuY530" colab_type="code" outputId="a35a1c12-3b02-41df-e741-f6dd0ad3770d" executionInfo={"status": "ok", "timestamp": 1564691681357, "user_tz": -540, "elapsed": 2536, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 163}
# !ls -al
# + id="DptY_D9qCVTN" colab_type="code" outputId="5647dcad-97ea-4bdd-bbd3-47c29f44c4f7" executionInfo={"status": "ok", "timestamp": 1564691883128, "user_tz": -540, "elapsed": 66715, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
from google.colab import files
print('Downloading best model...') # Known bug; if using Firefox, a print statement in the same cell is necessary.
files.download('./best_model_de2en.pt')
# + id="AymW4-HXf8qO" colab_type="code" outputId="fb5331ae-f6cf-46b7-f09f-fdfa46b11caf" executionInfo={"status": "ok", "timestamp": 1564691955648, "user_tz": -540, "elapsed": 66699, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
print('Downloading last model...') # Known bug; if using Firefox, a print statement in the same cell is necessary.
files.download('./last_model_de2en.pt')
# + id="fTiU0LlA53qr" colab_type="code" colab={}
| colab/NMT-Seq2SeqWithAttention-de2en.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nama
# language: python
# name: nama
# ---
# %load_ext autoreload
# %autoreload 2
# # Clean preferred names and split into separate given and surname datasets
# +
from os.path import join
from mpire import WorkerPool
import pandas as pd
from pathlib import Path
from src.data.filesystem import glob
from src.data.normalize import normalize
# -
in_path = "s3://familysearch-names/raw/tree-preferred/"
given_out_path = "s3://familysearch-names/interim/tree-preferred-given/"
surname_out_path = "s3://familysearch-names/interim/tree-preferred-surname/"
# + pycharm={"name": "#%%\n"}
def normalize_given_and_join(name):
return " ".join(normalize(name, False))
def normalize_surname_and_join(name):
return " ".join(normalize(name, True))
# + pycharm={"name": "#%%\n"}
def process_file(shared, filename):
given_out_path, surname_out_path = shared
basename = Path(filename).stem
df = pd.read_csv(
filename,
sep="|",
compression="gzip",
names=["name"],
dtype={"name": str},
na_filter=False,
encoding="utf-8",
)
# create separate given and surname dataframes
given_df = df[["name"]].copy()
surname_df = df[["name"]].copy()
del df
# split names into given and surname
given_df["name"] = given_df["name"].str.replace("\^.*$", "", regex=True)
surname_df["name"] = surname_df["name"].str.replace("^.*\^", "", regex=True)
# filter out non-latin names
given_df = given_df[
given_df["name"].str.endswith("~Latn")
]
surname_df = surname_df[
surname_df["name"].str.endswith("~Latn")
]
# remove ~Latn suffix and lowercase
given_df["name"] = given_df["name"].str.replace(
"~Latn$", "", regex=True
).str.lower()
surname_df["name"] = surname_df["name"].str.replace(
"~Latn$", "", regex=True
).str.lower()
# normalize names and join the pieces back into a single space-separated string
given_df["name"] = given_df["name"].map(normalize_given_and_join)
surname_df["name"] = surname_df["name"].map(normalize_surname_and_join)
# write files
given_df.to_parquet(
join(given_out_path, basename) + ".parquet", engine="pyarrow", compression="snappy"
)
surname_df.to_parquet(
join(surname_out_path, basename) + ".parquet", engine="pyarrow", compression="snappy"
)
# -
# process files
filenames = glob(join(in_path,"*.gz"))
with WorkerPool(shared_objects=(given_out_path, surname_out_path)) as pool:
pool.map(process_file, filenames, progress_bar=True)
# + pycharm={"name": "#%%\n"}
| notebooks/11_clean_preferred.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# pandas and numpy for data manipulation
import pandas as pd
import numpy as np
# matplotlib and seaborn for plotting
import matplotlib.pyplot as plt
import seaborn as sns
# Suppress warnings from pandas
import warnings
warnings.filterwarnings('ignore')
plt.style.use('fivethirtyeight')
# -
# Read in bureau
bureau = pd.read_csv('./input/home-credit-default-risk/bureau.csv')
bureau.head()
# Groupby the client id (SK_ID_CURR), count the number of previous loans, and rename the column
previous_loan_counts = bureau.groupby('SK_ID_CURR', as_index=False)['SK_ID_BUREAU'].count().rename(columns = {'SK_ID_BUREAU': 'previous_loan_counts'})
previous_loan_counts.head()
# +
# Join to the training dataframe
train = pd.read_csv('./input/home-credit-default-risk/application_train.csv')
train = train.merge(previous_loan_counts, on = 'SK_ID_CURR', how = 'left')
# Fill the missing values with 0
train['previous_loan_counts'] = train['previous_loan_counts'].fillna(0)
train.head()
# -
# Plots the disribution of a variable colored by value of the target
def kde_target(var_name, df):
# Calculate the correlation coefficient between the new variable and the target
corr = df['TARGET'].corr(df[var_name])
# Calculate medians for repaid vs not repaid
avg_repaid = df.ix[df['TARGET'] == 0, var_name].median()
avg_not_repaid = df.ix[df['TARGET'] == 1, var_name].median()
plt.figure(figsize = (12, 6))
# Plot the distribution for target == 0 and target == 1
sns.kdeplot(df.ix[df['TARGET'] == 0, var_name], label = 'TARGET == 0')
sns.kdeplot(df.ix[df['TARGET'] == 1, var_name], label = 'TARGET == 1')
# label the plot
plt.xlabel(var_name); plt.ylabel('Density'); plt.title('%s Distribution' % var_name)
plt.legend();
# print out the correlation
print('The correlation between %s and the TARGET is %0.4f' % (var_name, corr))
# Print out average values
print('Median value for loan that was not repaid = %0.4f' % avg_not_repaid)
print('Median value for loan that was repaid = %0.4f' % avg_repaid)
kde_target('EXT_SOURCE_3', train)
kde_target('previous_loan_counts', train)
# Group by the client id, calculate aggregation statistics
bureau_agg = bureau.drop(columns = ['SK_ID_BUREAU']).groupby('SK_ID_CURR', as_index = False).agg(['count', 'mean', 'max', 'min', 'sum']).reset_index()
bureau_agg.head()
# +
# List of column names
columns = ['SK_ID_CURR']
# Iterate through the variables names
for var in bureau_agg.columns.levels[0]:
# Skip the id name
if var != 'SK_ID_CURR':
# Iterate through the stat names
for stat in bureau_agg.columns.levels[1][:-1]:
# Make a new column name for the variable and stat
columns.append('bureau_%s_%s' % (var, stat))
# -
# Assign the list of columns names as the dataframe column names
bureau_agg.columns = columns
bureau_agg.head()
# Merge with the training data
train = train.merge(bureau_agg, on = 'SK_ID_CURR', how = 'left')
train.head()
# +
# List of new correlations
new_corrs = []
# Iterate through the columns
for col in columns:
# Calculate correlation with the target
corr = train['TARGET'].corr(train[col])
# Append the list as a tuple
new_corrs.append((col, corr))
# -
# Sort the correlations by the absolute value
# Make sure to reverse to put the largest values at the front of list
new_corrs = sorted(new_corrs, key = lambda x: abs(x[1]), reverse = True)
new_corrs[:15]
kde_target('bureau_DAYS_CREDIT_mean', train)
def agg_numeric(df, group_var, df_name):
"""Aggregates the numeric values in a dataframe. This can
be used to create features for each instance of the grouping variable.
Parameters
--------
df (dataframe):
the dataframe to calculate the statistics on
group_var (string):
the variable by which to group df
df_name (string):
the variable used to rename the columns
Return
--------
agg (dataframe):
a dataframe with the statistics aggregated for
all numeric columns. Each instance of the grouping variable will have
the statistics (mean, min, max, sum; currently supported) calculated.
The columns are also renamed to keep track of features created.
"""
# Remove id variables other than grouping variable
for col in df:
if col != group_var and 'SK_ID' in col:
df = df.drop(columns = col)
group_ids = df[group_var]
numeric_df = df.select_dtypes('number')
numeric_df[group_var] = group_ids
# Group by the specified variable and calculate the statistics
agg = numeric_df.groupby(group_var).agg(['count', 'mean', 'max', 'min', 'sum']).reset_index()
# Need to create new column names
columns = [group_var]
# Iterate through the variables names
for var in agg.columns.levels[0]:
# Skip the grouping variable
if var != group_var:
# Iterate through the stat names
for stat in agg.columns.levels[1][:-1]:
# Make a new column name for the variable and stat
columns.append('%s_%s_%s' % (df_name, var, stat))
agg.columns = columns
return agg
bureau_agg_new = agg_numeric(bureau.drop(columns = ['SK_ID_BUREAU']), group_var = 'SK_ID_CURR', df_name = 'bureau')
bureau_agg_new.head()
# Function to calculate correlations with the target for a dataframe
def target_corrs(df):
# List of correlations
corrs = []
# Iterate through the columns
for col in df.columns:
print(col)
# Skip the target column
if col != 'TARGET':
# Calculate correlation with the target
corr = df['TARGET'].corr(df[col])
# Append the list as a tuple
corrs.append((col, corr))
# Sort by absolute magnitude of correlations
corrs = sorted(corrs, key = lambda x: abs(x[1]), reverse = True)
return corrs
categorical = pd.get_dummies(bureau.select_dtypes('object'))
categorical['SK_ID_CURR'] = bureau['SK_ID_CURR']
categorical.head()
categorical_grouped = categorical.groupby('SK_ID_CURR').agg(['sum', 'mean'])
categorical_grouped.head()
categorical_grouped.columns.levels[0][:10]
categorical_grouped.columns.levels[1]
# +
group_var = 'SK_ID_CURR'
# Need to create new column names
columns = []
# Iterate through the variables names
for var in categorical_grouped.columns.levels[0]:
# Skip the grouping variable
if var != group_var:
# Iterate through the stat names
for stat in ['count', 'count_norm']:
# Make a new column name for the variable and stat
columns.append('%s_%s' % (var, stat))
# Rename the columns
categorical_grouped.columns = columns
categorical_grouped.head()
# -
train = train.merge(categorical_grouped, left_on = 'SK_ID_CURR', right_index = True, how = 'left')
train.head()
train.shape
train.iloc[:10, 123:]
def count_categorical(df, group_var, df_name):
"""Computes counts and normalized counts for each observation
of `group_var` of each unique category in every categorical variable
Parameters
--------
df : dataframe
The dataframe to calculate the value counts for.
group_var : string
The variable by which to group the dataframe. For each unique
value of this variable, the final dataframe will have one row
df_name : string
Variable added to the front of column names to keep track of columns
Return
--------
categorical : dataframe
A dataframe with counts and normalized counts of each unique category in every categorical variable
with one row for every unique value of the `group_var`.
"""
# Select the categorical columns
categorical = pd.get_dummies(df.select_dtypes('object'))
# Make sure to put the identifying id on the column
categorical[group_var] = df[group_var]
# Groupby the group var and calculate the sum and mean
categorical = categorical.groupby(group_var).agg(['sum', 'mean'])
column_names = []
# Iterate through the columns in level 0
for var in categorical.columns.levels[0]:
# Iterate through the stats in level 1
for stat in ['count', 'count_norm']:
# Make a new column name
column_names.append('%s_%s_%s' % (df_name, var, stat))
categorical.columns = column_names
return categorical
bureau_counts = count_categorical(bureau, group_var = 'SK_ID_CURR', df_name = 'bureau')
bureau_counts.head()
# Read in bureau balance
bureau_balance = pd.read_csv('./input/home-credit-default-risk/bureau_balance.csv')
bureau_balance.head()
# Counts of each type of status for each previous loan
bureau_balance_counts = count_categorical(bureau_balance, group_var = 'SK_ID_BUREAU', df_name = 'bureau_balance')
bureau_balance_counts.head()
# Calculate value count statistics for each `SK_ID_CURR`
bureau_balance_agg = agg_numeric(bureau_balance, group_var = 'SK_ID_BUREAU', df_name = 'bureau_balance')
bureau_balance_agg.head()
# +
# Dataframe grouped by the loan
bureau_by_loan = bureau_balance_agg.merge(bureau_balance_counts, right_index = True, left_on = 'SK_ID_BUREAU', how = 'outer')
# Merge to include the SK_ID_CURR
bureau_by_loan = bureau_by_loan.merge(bureau[['SK_ID_BUREAU', 'SK_ID_CURR']], on = 'SK_ID_BUREAU', how = 'left')
bureau_by_loan.head()
# -
bureau_balance_by_client = agg_numeric(bureau_by_loan.drop(columns = ['SK_ID_BUREAU']), group_var = 'SK_ID_CURR', df_name = 'client')
bureau_balance_by_client.head()
# Free up memory by deleting old objects
import gc
gc.enable()
del train, bureau, bureau_balance, bureau_agg, bureau_agg_new, bureau_balance_agg, bureau_balance_counts, bureau_by_loan, bureau_balance_by_client, bureau_counts
gc.collect()
# Read in new copies of all the dataframes
train = pd.read_csv('./input/home-credit-default-risk/application_train.csv')
bureau = pd.read_csv('./input/home-credit-default-risk/bureau.csv')
bureau_balance = pd.read_csv('./input/home-credit-default-risk/bureau_balance.csv')
bureau_counts = count_categorical(bureau, group_var = 'SK_ID_CURR', df_name = 'bureau')
bureau_counts.head()
bureau_agg = agg_numeric(bureau.drop(columns = ['SK_ID_BUREAU']), group_var = 'SK_ID_CURR', df_name = 'bureau')
bureau_agg.head()
bureau_balance_counts = count_categorical(bureau_balance, group_var = 'SK_ID_BUREAU', df_name = 'bureau_balance')
bureau_balance_counts.head()
bureau_balance_agg = agg_numeric(bureau_balance, group_var = 'SK_ID_BUREAU', df_name = 'bureau_balance')
bureau_balance_agg.head()
# +
# Dataframe grouped by the loan
bureau_by_loan = bureau_balance_agg.merge(bureau_balance_counts, right_index = True, left_on = 'SK_ID_BUREAU', how = 'outer')
# Merge to include the SK_ID_CURR
bureau_by_loan = bureau[['SK_ID_BUREAU', 'SK_ID_CURR']].merge(bureau_by_loan, on = 'SK_ID_BUREAU', how = 'left')
# Aggregate the stats for each client
bureau_balance_by_client = agg_numeric(bureau_by_loan.drop(columns = ['SK_ID_BUREAU']), group_var = 'SK_ID_CURR', df_name = 'client')
# -
original_features = list(train.columns)
print('Original Number of Features: ', len(original_features))
# +
# Merge with the value counts of bureau
train = train.merge(bureau_counts, on = 'SK_ID_CURR', how = 'left')
# Merge with the stats of bureau
train = train.merge(bureau_agg, on = 'SK_ID_CURR', how = 'left')
# Merge with the monthly information grouped by client
train = train.merge(bureau_balance_by_client, on = 'SK_ID_CURR', how = 'left')
# -
new_features = list(train.columns)
print('Number of features using previous loans from other institutions data: ', len(new_features))
def missing_values_table(df):
# Total missing values
mis_val = df.isnull().sum()
# Percentage of missing values
mis_val_percent = 100 * df.isnull().sum() / len(df)
# Make a table with the results
mis_val_table = pd.concat([mis_val, mis_val_percent], axis=1)
# Rename the columns
mis_val_table_ren_columns = mis_val_table.rename(
columns = {0 : 'Missing Values', 1 : '% of Total Values'})
# Sort the table by percentage of missing descending
mis_val_table_ren_columns = mis_val_table_ren_columns[
mis_val_table_ren_columns.iloc[:,1] != 0].sort_values(
'% of Total Values', ascending=False).round(1)
# Print some summary information
print ("Your selected dataframe has " + str(df.shape[1]) + " columns.\n"
"There are " + str(mis_val_table_ren_columns.shape[0]) +
" columns that have missing values.")
# Return the dataframe with missing information
return mis_val_table_ren_columns
missing_train = missing_values_table(train)
missing_train.head(10)
missing_train_vars = list(missing_train.index[missing_train['% of Total Values'] > 90])
len(missing_train_vars)
# +
# Read in the test dataframe
test = pd.read_csv('./input/home-credit-default-risk/application_test.csv')
# Merge with the value counts of bureau
test = test.merge(bureau_counts, on = 'SK_ID_CURR', how = 'left')
# Merge with the stats of bureau
test = test.merge(bureau_agg, on = 'SK_ID_CURR', how = 'left')
# Merge with the value counts of bureau balance
test = test.merge(bureau_balance_by_client, on = 'SK_ID_CURR', how = 'left')
# -
print('Shape of Testing Data: ', test.shape)
# +
train_labels = train['TARGET']
# Align the dataframes, this will remove the 'TARGET' column
train, test = train.align(test, join = 'inner', axis = 1)
train['TARGET'] = train_labels
# -
print('Training Data Shape: ', train.shape)
print('Testing Data Shape: ', test.shape)
missing_test = missing_values_table(test)
missing_test.head(10)
missing_test_vars = list(missing_test.index[missing_test['% of Total Values'] > 90])
len(missing_test_vars)
missing_columns = list(set(missing_test_vars + missing_train_vars))
print('There are %d columns with more than 90%% missing in either the training or testing data.' % len(missing_columns))
# Drop the missing columns
train = train.drop(columns = missing_columns)
test = test.drop(columns = missing_columns)
train.to_csv('./input/train_bureau_raw.csv', index = False)
test.to_csv('./input/test_bureau_raw.csv', index = False)
# Calculate all correlations in dataframe
corrs = train.corr()
# +
corrs = corrs.sort_values('TARGET', ascending = False)
# Ten most positive correlations
pd.DataFrame(corrs['TARGET'].head(10))
# -
# Ten most negative correlations
pd.DataFrame(corrs['TARGET'].dropna().tail(10))
kde_target(var_name='client_bureau_balance_MONTHS_BALANCE_count_mean', df=train)
kde_target(var_name='bureau_CREDIT_ACTIVE_Active_count_norm', df=train)
# +
# Set the threshold
threshold = 0.8
# Empty dictionary to hold correlated variables
above_threshold_vars = {}
# For each column, record the variables that are above the threshold
for col in corrs:
above_threshold_vars[col] = list(corrs.index[corrs[col] > threshold])
# +
# columns to remove and columns already examined
cols_to_remove = []
cols_seen = []
cols_to_remove_pair = []
# Iterate through columns and correlated columns
for key, value in above_threshold_vars.items():
# Keep track of columns already examined
cols_seen.append(key)
for x in value:
if x == key:
next
else:
# Only want to remove one in a pair
if x not in cols_seen:
cols_to_remove.append(x)
cols_to_remove_pair.append(key)
cols_to_remove = list(set(cols_to_remove))
print('Number of columns to remove: ', len(cols_to_remove))
# +
train_corrs_removed = train.drop(columns = cols_to_remove)
test_corrs_removed = test.drop(columns = cols_to_remove)
print('Training Corrs Removed Shape: ', train_corrs_removed.shape)
print('Testing Corrs Removed Shape: ', test_corrs_removed.shape)
# +
import lightgbm as lgb
from sklearn.model_selection import KFold
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import LabelEncoder
import gc
import matplotlib.pyplot as plt
# -
def plot_feature_importances(df):
"""
Plot importances returned by a model. This can work with any measure of
feature importance provided that higher importance is better.
Args:
df (dataframe): feature importances. Must have the features in a column
called `features` and the importances in a column called `importance
Returns:
shows a plot of the 15 most importance features
df (dataframe): feature importances sorted by importance (highest to lowest)
with a column for normalized importance
"""
# Sort features according to importance
df = df.sort_values('importance', ascending = False).reset_index()
# Normalize the feature importances to add up to one
df['importance_normalized'] = df['importance'] / df['importance'].sum()
# Make a horizontal bar chart of feature importances
plt.figure(figsize = (10, 6))
ax = plt.subplot()
# Need to reverse the index to plot most important on top
ax.barh(list(reversed(list(df.index[:15]))),
df['importance_normalized'].head(15),
align = 'center', edgecolor = 'k')
# Set the yticks and labels
ax.set_yticks(list(reversed(list(df.index[:15]))))
ax.set_yticklabels(df['feature'].head(15))
# Plot labeling
plt.xlabel('Normalized Importance'); plt.title('Feature Importances')
plt.show()
return df
train_control = pd.read_csv('./input/home-credit-default-risk/application_train.csv')
test_control = pd.read_csv('./input/home-credit-default-risk/application_test.csv')
submission, fi, metrics = model(train_control, test_control)
metrics
fi_sorted = plot_feature_importances(fi)
submission.to_csv('./submission/control.csv', index = False)
submission_raw, fi_raw, metrics_raw = model(train, test)
metrics_raw
fi_raw_sorted = plot_feature_importances(fi_raw)
# +
top_100 = list(fi_raw_sorted['feature'])[:100]
new_features = [x for x in top_100 if x not in list(fi['feature'])]
print('%% of Top 100 Features created from the bureau data = %d.00' % len(new_features))
# -
submission_raw.to_csv('./submission/test_one.csv', index = False)
submission_corrs, fi_corrs, metrics_corr = model(train_corrs_removed, test_corrs_removed)
metrics_corr
fi_corrs_sorted = plot_feature_importances(fi_corrs)
submission_corrs.to_csv('./submission/test_two.csv', index = False)
| mykernel2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="DjUA6S30k52h"
# ##### Copyright 2021 The TensorFlow Authors.
# + cellView="form" id="SpNWyqewk8fE"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="6x1ypzczQCwy"
# # Feature Engineering using TFX Pipeline and TensorFlow Transform
#
# ***Transform input data and traing a model with a TFX pipeline.***
# + [markdown] id="HU9YYythm0dx"
# Note: We recommend running this tutorial in a Colab notebook, with no setup required! Just click "Run in Google Colab".
#
# <div class="devsite-table-wrapper"><table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/tfx/tutorials/tfx/penguin_tft">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png"/>View on TensorFlow.org</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/tfx/blob/master/docs/tutorials/tfx/penguin_tft.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png">Run in Google Colab</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/tfx/tree/master/docs/tutorials/tfx/penguin_tft.ipynb">
# <img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">View source on GitHub</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/tfx/docs/tutorials/tfx/penguin_tft.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a></td>
# </table></div>
# + [markdown] id="_VuwrlnvQJ5k"
# In this notebook-based tutorial, we will create and run a TFX pipeline
# to ingest raw input data and preprocess it appropriately for ML training.
# This notebook is based on the TFX pipeline we built in
# [Data validation using TFX Pipeline and TensorFlow Data Validation Tutorial](https://www.tensorflow.org/tfx/tutorials/tfx/penguin_tfdv).
# If you have not read that one yet, you should read it before proceeding with
# this notebook.
#
# You can increase the predictive quality of your data and/or reduce
# dimensionality with feature engineering. One of the benefits of using TFX is
# that you will write your transformation code once, and the resulting transforms
# will be consistent between training and serving in
# order to avoid training/serving skew.
#
# We will add a `Transform` component to the pipeline. The Transform component is
# implemented using the
# [tf.transform](https://www.tensorflow.org/tfx/transform/get_started) library.
#
# Please see
# [Understanding TFX Pipelines](https://www.tensorflow.org/tfx/guide/understanding_tfx_pipelines)
# to learn more about various concepts in TFX.
# + [markdown] id="Fmgi8ZvQkScg"
# ## Set Up
# We first need to install the TFX Python package and download
# the dataset which we will use for our model.
#
# ### Upgrade Pip
#
# To avoid upgrading Pip in a system when running locally,
# check to make sure that we are running in Colab.
# Local systems can of course be upgraded separately.
# + id="as4OTe2ukSqm"
try:
import colab
# !pip install --upgrade pip
except:
pass
# + [markdown] id="MZOYTt1RW4TK"
# ### Install TFX
#
# + id="iyQtljP-qPHY"
# !pip install -U tfx
# + [markdown] id="EwT0nov5QO1M"
# ### Did you restart the runtime?
#
# If you are using Google Colab, the first time that you run
# the cell above, you must restart the runtime by clicking
# above "RESTART RUNTIME" button or using "Runtime > Restart
# runtime ..." menu. This is because of the way that Colab
# loads packages.
# + [markdown] id="BDnPgN8UJtzN"
# Check the TensorFlow and TFX versions.
# + id="6jh7vKSRqPHb"
import tensorflow as tf
print('TensorFlow version: {}'.format(tf.__version__))
from tfx import v1 as tfx
print('TFX version: {}'.format(tfx.__version__))
# + [markdown] id="aDtLdSkvqPHe"
# ### Set up variables
#
# There are some variables used to define a pipeline. You can customize these
# variables as you want. By default all output from the pipeline will be
# generated under the current directory.
# + id="EcUseqJaE2XN"
import os
PIPELINE_NAME = "penguin-transform"
# Output directory to store artifacts generated from the pipeline.
PIPELINE_ROOT = os.path.join('pipelines', PIPELINE_NAME)
# Path to a SQLite DB file to use as an MLMD storage.
METADATA_PATH = os.path.join('metadata', PIPELINE_NAME, 'metadata.db')
# Output directory where created models from the pipeline will be exported.
SERVING_MODEL_DIR = os.path.join('serving_model', PIPELINE_NAME)
from absl import logging
logging.set_verbosity(logging.INFO) # Set default logging level.
# + [markdown] id="qsO0l5F3dzOr"
# ### Prepare example data
# We will download the example dataset for use in our TFX pipeline. The dataset
# we are using is
# [Palmer Penguins dataset](https://allisonhorst.github.io/palmerpenguins/articles/intro.html).
#
# However, unlike previous tutorials which used an already preprocessed dataset,
# we will use the **raw** Palmer Penguins dataset.
#
# + [markdown] id="11J7XiCq6AFP"
# Because the TFX ExampleGen component reads inputs from a directory, we need
# to create a directory and copy the dataset to it.
# + id="4fxMs6u86acP"
import urllib.request
import tempfile
DATA_ROOT = tempfile.mkdtemp(prefix='tfx-data') # Create a temporary directory.
_data_path = 'https://storage.googleapis.com/download.tensorflow.org/data/palmer_penguins/penguins_size.csv'
_data_filepath = os.path.join(DATA_ROOT, "data.csv")
urllib.request.urlretrieve(_data_path, _data_filepath)
# + [markdown] id="ASpoNmxKSQjI"
# Take a quick look at what the raw data looks like.
# + id="-eSz28UDSnlG"
# !head {_data_filepath}
# + [markdown] id="OTtQNq1DdVvG"
# There are some entries with missing values which are represented as `NA`.
# We will just delete those entries in this tutorial.
# + id="fQhpoaqff9ca"
# !sed -i '/\bNA\b/d' {_data_filepath}
# !head {_data_filepath}
# + [markdown] id="z8EOfCy1dzO2"
# You should be able to see seven features which describe penguins. We will use
# the same set of features as the previous tutorials - 'culmen_length_mm',
# 'culmen_depth_mm', 'flipper_length_mm', 'body_mass_g' - and will predict
# the 'species' of a penguin.
#
# **The only difference will be that the input data is not preprocessed.** Note
# that we will not use other features like 'island' or 'sex' in this tutorial.
# + [markdown] id="Jtbrkjjc-IKA"
# ### Prepare a schema file
#
# As described in
# [Data validation using TFX Pipeline and TensorFlow Data Validation Tutorial](https://www.tensorflow.org/tfx/tutorials/tfx/penguin_tfdv),
# we need a schema file for the dataset. Because the dataset is different from the previous tutorial we need to generate it again. In this tutorial, we will skip those steps and just use a prepared schema file.
#
# + id="EDoB97m8B9nG"
import shutil
SCHEMA_PATH = 'schema'
_schema_uri = 'https://raw.githubusercontent.com/tensorflow/tfx/master/tfx/examples/penguin/schema/raw/schema.pbtxt'
_schema_filename = 'schema.pbtxt'
_schema_filepath = os.path.join(SCHEMA_PATH, _schema_filename)
os.makedirs(SCHEMA_PATH, exist_ok=True)
urllib.request.urlretrieve(_schema_uri, _schema_filepath)
# + [markdown] id="gKJ_HDJQB94b"
# This schema file was created with the same pipeline as in the previous tutorial
# without any manual changes.
# + [markdown] id="nH6gizcpSwWV"
# ## Create a pipeline
#
# TFX pipelines are defined using Python APIs. We will add `Transform`
# component to the pipeline we created in the
# [Data Validation tutorial](https://www.tensorflow.org/tfx/tutorials/tfx/penguin_tfdv).
#
# A Transform component requires input data from an `ExampleGen` component and
# a schema from a `SchemaGen` component, and produces a "transform graph". The
# output will be used in a `Trainer` component. Transform can optionally
# produce "transformed data" in addition, which is the materialized data after
# transformation.
# However, we will transform data during training in this tutorial without
# materialization of the intermediate transformed data.
#
# One thing to note is that we need to define a Python function,
# `preprocessing_fn` to describe how input data should be transformed. This is
# similar to a Trainer component which also requires user code for model
# definition.
#
# + [markdown] id="lOjDv93eS5xV"
# ### Write preprocessing and training code
#
# We need to define two Python functions. One for Transform and one for Trainer.
#
# #### preprocessing_fn
# The Transform component will find a function named `preprocessing_fn` in the
# given module file as we did for `Trainer` component. You can also specify a
# specific function using the
# [`preprocessing_fn` parameter](https://github.com/tensorflow/tfx/blob/142de6e887f26f4101ded7925f60d7d4fe9d42ed/tfx/components/transform/component.py#L113)
# of the Transform component.
#
# In this example, we will do two kinds of transformation. For continuous numeric
# features like `culmen_length_mm` and `body_mass_g`, we will normalize these
# values using the
# [tft.scale_to_z_score](https://www.tensorflow.org/tfx/transform/api_docs/python/tft/scale_to_z_score)
# function. For the label feature, we need to convert string labels into numeric
# index values. We will use
# [`tf.lookup.StaticHashTable`](https://www.tensorflow.org/api_docs/python/tf/lookup/StaticHashTable)
# for conversion.
#
# To identify transformed fields easily, we append a `_xf` suffix to the
# transformed feature names.
#
# #### run_fn
#
# The model itself is almost the same as in the previous tutorials, but this time
# we will transform the input data using the transform graph from the Transform
# component.
#
# One more important difference compared to the previous tutorial is that now we
# export a model for serving which includes not only the computation graph of the
# model, but also the transform graph for preprocessing, which is generated in
# Transform component. We need to define a separate function which will be used
# for serving incoming requests. You can see that the same function
# `_apply_preprocessing` was used for both of the training data and the
# serving request.
#
# + id="aES7Hv5QTDK3"
_module_file = 'penguin_utils.py'
# + id="Gnc67uQNTDfW"
# %%writefile {_module_file}
from typing import List, Text
from absl import logging
import tensorflow as tf
from tensorflow import keras
from tensorflow_metadata.proto.v0 import schema_pb2
import tensorflow_transform as tft
from tensorflow_transform.tf_metadata import schema_utils
from tfx import v1 as tfx
from tfx_bsl.public import tfxio
# Specify features that we will use.
_FEATURE_KEYS = [
'culmen_length_mm', 'culmen_depth_mm', 'flipper_length_mm', 'body_mass_g'
]
_LABEL_KEY = 'species'
_TRAIN_BATCH_SIZE = 20
_EVAL_BATCH_SIZE = 10
# NEW: Transformed features will have '_xf' suffix.
def _transformed_name(key):
return key + '_xf'
# NEW: TFX Transform will call this function.
def preprocessing_fn(inputs):
"""tf.transform's callback function for preprocessing inputs.
Args:
inputs: map from feature keys to raw not-yet-transformed features.
Returns:
Map from string feature key to transformed feature.
"""
outputs = {}
# Uses features defined in _FEATURE_KEYS only.
for key in _FEATURE_KEYS:
# tft.scale_to_z_score computes the mean and variance of the given feature
# and scales the output based on the result.
outputs[_transformed_name(key)] = tft.scale_to_z_score(inputs[key])
# For the label column we provide the mapping from string to index.
# We could instead use `tft.compute_and_apply_vocabulary()` in order to
# compute the vocabulary dynamically and perform a lookup.
# Since in this example there are only 3 possible values, we use a hard-coded
# table for simplicity.
table_keys = ['Adelie', 'Chinstrap', 'Gentoo']
initializer = tf.lookup.KeyValueTensorInitializer(
keys=table_keys,
values=tf.cast(tf.range(len(table_keys)), tf.int64),
key_dtype=tf.string,
value_dtype=tf.int64)
table = tf.lookup.StaticHashTable(initializer, default_value=-1)
outputs[_transformed_name(_LABEL_KEY)] = table.lookup(inputs[_LABEL_KEY])
return outputs
# NEW: This function will apply the same transform operation to training data
# and serving requests.
def _apply_preprocessing(raw_features, tft_layer):
transformed_features = tft_layer(raw_features)
if _LABEL_KEY in raw_features:
transformed_label = transformed_features.pop(_transformed_name(_LABEL_KEY))
return transformed_features, transformed_label
else:
return transformed_features, None
# NEW: This function will create a handler function which gets a serialized
# tf.example, preprocess and run an inference with it.
def _get_serve_tf_examples_fn(model, tf_transform_output):
# We must save the tft_layer to the model to ensure its assets are kept and
# tracked.
model.tft_layer = tf_transform_output.transform_features_layer()
@tf.function(input_signature=[
tf.TensorSpec(shape=[None], dtype=tf.string, name='examples')
])
def serve_tf_examples_fn(serialized_tf_examples):
# Expected input is a string which is serialized tf.Example format.
feature_spec = tf_transform_output.raw_feature_spec()
# Because input schema includes unnecessary fields like 'species' and
# 'island', we filter feature_spec to include required keys only.
required_feature_spec = {
k: v for k, v in feature_spec.items() if k in _FEATURE_KEYS
}
parsed_features = tf.io.parse_example(serialized_tf_examples,
required_feature_spec)
# Preprocess parsed input with transform operation defined in
# preprocessing_fn().
transformed_features, _ = _apply_preprocessing(parsed_features,
model.tft_layer)
# Run inference with ML model.
return model(transformed_features)
return serve_tf_examples_fn
def _input_fn(file_pattern: List[Text],
data_accessor: tfx.components.DataAccessor,
tf_transform_output: tft.TFTransformOutput,
batch_size: int = 200) -> tf.data.Dataset:
"""Generates features and label for tuning/training.
Args:
file_pattern: List of paths or patterns of input tfrecord files.
data_accessor: DataAccessor for converting input to RecordBatch.
tf_transform_output: A TFTransformOutput.
batch_size: representing the number of consecutive elements of returned
dataset to combine in a single batch
Returns:
A dataset that contains (features, indices) tuple where features is a
dictionary of Tensors, and indices is a single Tensor of label indices.
"""
dataset = data_accessor.tf_dataset_factory(
file_pattern,
tfxio.TensorFlowDatasetOptions(batch_size=batch_size),
schema=tf_transform_output.raw_metadata.schema)
transform_layer = tf_transform_output.transform_features_layer()
def apply_transform(raw_features):
return _apply_preprocessing(raw_features, transform_layer)
return dataset.map(apply_transform).repeat()
def _build_keras_model() -> tf.keras.Model:
"""Creates a DNN Keras model for classifying penguin data.
Returns:
A Keras Model.
"""
# The model below is built with Functional API, please refer to
# https://www.tensorflow.org/guide/keras/overview for all API options.
inputs = [
keras.layers.Input(shape=(1,), name=_transformed_name(f))
for f in _FEATURE_KEYS
]
d = keras.layers.concatenate(inputs)
for _ in range(2):
d = keras.layers.Dense(8, activation='relu')(d)
outputs = keras.layers.Dense(3)(d)
model = keras.Model(inputs=inputs, outputs=outputs)
model.compile(
optimizer=keras.optimizers.Adam(1e-2),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=[keras.metrics.SparseCategoricalAccuracy()])
model.summary(print_fn=logging.info)
return model
# TFX Trainer will call this function.
def run_fn(fn_args: tfx.components.FnArgs):
"""Train the model based on given args.
Args:
fn_args: Holds args used to train the model as name/value pairs.
"""
tf_transform_output = tft.TFTransformOutput(fn_args.transform_output)
train_dataset = _input_fn(
fn_args.train_files,
fn_args.data_accessor,
tf_transform_output,
batch_size=_TRAIN_BATCH_SIZE)
eval_dataset = _input_fn(
fn_args.eval_files,
fn_args.data_accessor,
tf_transform_output,
batch_size=_EVAL_BATCH_SIZE)
model = _build_keras_model()
model.fit(
train_dataset,
steps_per_epoch=fn_args.train_steps,
validation_data=eval_dataset,
validation_steps=fn_args.eval_steps)
# NEW: Save a computation graph including transform layer.
signatures = {
'serving_default': _get_serve_tf_examples_fn(model, tf_transform_output),
}
model.save(fn_args.serving_model_dir, save_format='tf', signatures=signatures)
# + [markdown] id="blaw0rs-emEf"
# Now you have completed all of the preparation steps to build a TFX pipeline.
# + [markdown] id="w3OkNz3gTLwM"
# ### Write a pipeline definition
#
# We define a function to create a TFX pipeline. A `Pipeline` object
# represents a TFX pipeline, which can be run using one of the pipeline
# orchestration systems that TFX supports.
#
# + id="M49yYVNBTPd4"
def _create_pipeline(pipeline_name: str, pipeline_root: str, data_root: str,
schema_path: str, module_file: str, serving_model_dir: str,
metadata_path: str) -> tfx.dsl.Pipeline:
"""Implements the penguin pipeline with TFX."""
# Brings data into the pipeline or otherwise joins/converts training data.
example_gen = tfx.components.CsvExampleGen(input_base=data_root)
# Computes statistics over data for visualization and example validation.
statistics_gen = tfx.components.StatisticsGen(
examples=example_gen.outputs['examples'])
# Import the schema.
schema_importer = tfx.dsl.Importer(
source_uri=schema_path,
artifact_type=tfx.types.standard_artifacts.Schema).with_id(
'schema_importer')
# Performs anomaly detection based on statistics and data schema.
example_validator = tfx.components.ExampleValidator(
statistics=statistics_gen.outputs['statistics'],
schema=schema_importer.outputs['result'])
# NEW: Transforms input data using preprocessing_fn in the 'module_file'.
transform = tfx.components.Transform(
examples=example_gen.outputs['examples'],
schema=schema_importer.outputs['result'],
materialize=False,
module_file=module_file)
# Uses user-provided Python function that trains a model.
trainer = tfx.components.Trainer(
module_file=module_file,
examples=example_gen.outputs['examples'],
# NEW: Pass transform_graph to the trainer.
transform_graph=transform.outputs['transform_graph'],
train_args=tfx.proto.TrainArgs(num_steps=100),
eval_args=tfx.proto.EvalArgs(num_steps=5))
# Pushes the model to a filesystem destination.
pusher = tfx.components.Pusher(
model=trainer.outputs['model'],
push_destination=tfx.proto.PushDestination(
filesystem=tfx.proto.PushDestination.Filesystem(
base_directory=serving_model_dir)))
components = [
example_gen,
statistics_gen,
schema_importer,
example_validator,
transform, # NEW: Transform component was added to the pipeline.
trainer,
pusher,
]
return tfx.dsl.Pipeline(
pipeline_name=pipeline_name,
pipeline_root=pipeline_root,
metadata_connection_config=tfx.orchestration.metadata
.sqlite_metadata_connection_config(metadata_path),
components=components)
# + [markdown] id="mJbq07THU2GV"
# ## Run the pipeline
#
# We will use `LocalDagRunner` as in the previous tutorial.
# + id="fAtfOZTYWJu-"
tfx.orchestration.LocalDagRunner().run(
_create_pipeline(
pipeline_name=PIPELINE_NAME,
pipeline_root=PIPELINE_ROOT,
data_root=DATA_ROOT,
schema_path=SCHEMA_PATH,
module_file=_module_file,
serving_model_dir=SERVING_MODEL_DIR,
metadata_path=METADATA_PATH))
# + [markdown] id="ppERq0Mj6xvW"
# You should see "INFO:absl:Component Pusher is finished." if the pipeline
# finished successfully.
#
# The pusher component pushes the trained model to the `SERVING_MODEL_DIR` which
# is the `serving_model/penguin-transform` directory if you did not change
# the variables in the previous steps. You can see the result from the file
# browser in the left-side panel in Colab, or using the following command:
# + id="NTHROkqX6yHx"
# List files in created model directory.
# !find {SERVING_MODEL_DIR}
# + [markdown] id="VTqM-WiZkPbt"
# You can also check the signature of the generated model using the
# [`saved_model_cli` tool](https://www.tensorflow.org/guide/saved_model#show_command).
# + id="YBfUzD_OkOq_"
# !saved_model_cli show --dir {SERVING_MODEL_DIR}/$(ls -1 {SERVING_MODEL_DIR} | sort -nr | head -1) --tag_set serve --signature_def serving_default
# + [markdown] id="DkAxFs_QszoZ"
# Because we defined `serving_default` with our own `serve_tf_examples_fn`
# function, the signature shows that it takes a single string.
# This string is a serialized string of tf.Examples and will be parsed with the
# [tf.io.parse_example()](https://www.tensorflow.org/api_docs/python/tf/io/parse_example)
# function as we defined earlier.
#
# We can load the exported model and try some inferences with a few examples.
# + id="Z1Yw5yYdvqKf"
# Find a model with the latest timestamp.
model_dirs = (item for item in os.scandir(SERVING_MODEL_DIR) if item.is_dir())
model_path = max(model_dirs, key=lambda i: int(i.name)).path
loaded_model = tf.keras.models.load_model(model_path)
inference_fn = loaded_model.signatures['serving_default']
# + id="xrOHIvnIv0-4"
# Prepare an example and run inference.
features = {
'culmen_length_mm': tf.train.Feature(float_list=tf.train.FloatList(value=[49.9])),
'culmen_depth_mm': tf.train.Feature(float_list=tf.train.FloatList(value=[16.1])),
'flipper_length_mm': tf.train.Feature(int64_list=tf.train.Int64List(value=[213])),
'body_mass_g': tf.train.Feature(int64_list=tf.train.Int64List(value=[5400])),
}
example_proto = tf.train.Example(features=tf.train.Features(feature=features))
examples = example_proto.SerializeToString()
result = inference_fn(examples=tf.constant([examples]))
print(result['output_0'].numpy())
# + [markdown] id="cri3mTgZ0SQ2"
# The third element, which corresponds to 'Gentoo' species, is expected to be the
# largest among three.
# + [markdown] id="08R8qvweThRf"
# ## Next steps
#
# If you want to learn more about Transform component, see
# [Transform Component guide](https://www.tensorflow.org/tfx/guide/transform).
# You can find more resources on https://www.tensorflow.org/tfx/tutorials.
#
# Please see
# [Understanding TFX Pipelines](https://www.tensorflow.org/tfx/guide/understanding_tfx_pipelines)
# to learn more about various concepts in TFX.
#
| site/en-snapshot/tfx/tutorials/tfx/penguin_tft.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import re
import pandas as pd
from bs4 import BeautifulSoup
response = requests.get('https://www.billboard.com/charts/hot-100')
doc = BeautifulSoup(response.text)
doc
chart = doc.find_all(class_ = "chart-list-item")
# +
rows = []
for info in chart:
row = {}
row['song'] = info.find(class_ = 'chart-list-item__title-text').text.strip()
row['artist'] = info.find(class_ = 'chart-list-item__artist').text.strip()
row['rank'] = info.find(class_ = 'chart-list-item__rank').text.strip()
rows.append(row)
rows
# -
df = pd.DataFrame(rows)
df
df.to_csv('Billboard100.csv', index = False)
| 09-homework/Billboard Hot 100 Scraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # The Rastrigin function: plotting results
#
# We are going to make the following plots for the results of the evolution stored in the database:
# * Error bars graph (raw scores).
# * Error bars graph (fitness scores).
# * Max/min/avg/std. dev. graph (raw scores).
# * Max/min/avg/std. dev. graph (fitness scores).
# * Raw and Fitness min/max difference graph.
# * Heat map of population raw score distribution.
# %matplotlib inline
from pyevolve_plot import plot_errorbars_raw, plot_errorbars_fitness, \
plot_maxmin_raw, plot_maxmin_fitness, \
plot_diff_raw, plot_pop_heatmap_raw
plot_errorbars_raw('rastrigin.db','ex1')
plot_errorbars_fitness('rastrigin.db','ex1')
plot_maxmin_raw('rastrigin.db','ex1')
plot_maxmin_raw('rastrigin.db','ex1')
plot_diff_raw('rastrigin.db','ex1')
plot_pop_heatmap_raw('rastrigin.db','ex1')
| Rastrigin_check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Comparing Regression Models
# In this notebook, we'll look at methods for comparing regression models. In this notebook, we'll use results from the paper [Validation of AMBER/GAFF for Relative Free Energy Calculations](https://chemrxiv.org/articles/Validation_of_AMBER_GAFF_for_Relative_Free_Energy_Calculations/7653434) to compare two different approaches to free energy calculations. More information can be found in this [blog post](http://practicalcheminformatics.blogspot.com/2019/02/some-thoughts-on-evaluating-predictive.html).
# Install the necessary Python libraries
# !pip install pandas matplotlib seaborn numpy scipy sklearn
# Import the necessary Python libraries
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import math
import numpy as np
from scipy.stats import norm, pearsonr
from sklearn.metrics import mean_squared_error
from scipy.stats import pearsonr
from collections import namedtuple
from math import sqrt
import warnings
# #### Read and Parse the Input Data
# As a first step, we want to grab the data from a set of Excel worksheets in the supporting material. The Excel workbook has multiple sheets, we need to get the names of those sheets. Fortunately Pandas has all sorts of facilities for manipulating Excel files.
xls_name = "https://raw.githubusercontent.com/PatWalters/practical_cheminformatics_tutorials/main/data/FEP_vs_GTI-dG-SI.xlsx"
df_dict = pd.read_excel(xls_name,sheet_name=None,header=None)
# Now we can define a function that will grab the data from a particular sheet. In this case the data we want is in columns 15-17. We'll grab those columns and create a new dataframe with the columns labeled as Experimental Δ G, FEP Δ G, and TI Δ G. We will also include columns with the target and with the target concatenated with a sequential index. Ok, in truth, we're going to label the columns as Experimental_dG, etc. for convenience. The final row in columns 15-17 is the sum of the values in the column. We don't want this, so we'll use df = df.query("Experimental_dG > -20") to get rid of the sum row.
def get_data_from_sheet(df_dict, sheet_name, col_names):
df = df_dict[sheet_name]
df = df[df.columns[15:18]]
col_names = ["Experimental_dG","FEP_dG","TI_dG"]
df.columns = col_names
df = df.dropna()
df = df.query("Experimental_dG > -20")
rows,_ = df.shape
df.insert(0,"Target",[sheet_name]*rows)
df.insert(1,"ID",[f"{sheet_name}_{x}" for x in range(0,rows)])
return df
# Define the number of decimal places displayed in the Pandas table.
pd.options.display.float_format = '{:,.2f}'.format
# Read the Excel sheets and put the data into a Pandas dataframe.
sheet_names = df_dict.keys()
col_names = ["Experimental_dG","FEP_dG","TI_dG"]
big_df = pd.DataFrame(columns=["Target","ID"]+col_names)
for sheet in sheet_names:
tmp_df = get_data_from_sheet(df_dict,sheet,col_names)
big_df = big_df.append(tmp_df)
big_df.sort_values("Target",inplace=True)
big_df.head()
# #### Reformat the Data for Plotting
# In order to make some of our plots we need to reshape the data so that "FEP_dG" and "TI_dG" are in one column. We can use the Pandas "melt" function to do this.
mdf = big_df.melt(id_vars=["Target","ID","Experimental_dG"])
mdf["DataSet"] = mdf.Target + "_" + mdf.variable.str.replace("_dG","")
mdf.sort_values("DataSet",inplace=True)
cols = list(mdf.columns)
cols[4] = "Predicted_dG"
mdf.columns = cols
mdf.head()
# In order to make the plots look better we'll add another column called "Method" to label FEP and TI
mdf["Method"] = [x.replace("_dG","") for x in mdf.variable.values]
mdf.head()
# Ok, now we're ready to make some plots. I want to use the Seaborn FacetGrid class to put plots side by side. In order to do this, I'm going to define a custom plotting function that plots Experimental $\Delta$G vs Predicted $\Delta$G. We also want to put in lines at 2 kcal above and below the unity line. I could make this function more customizable and pass a bunch of kwargs, but that will also make the function a lot more complicated. For things like this, I often find it easier just to hard code the parameters I want. The only argument being passed is "skip_text" which tells the function whether to display the $R^2$ for the correlation.
def draw_dG_facet(*args,**kwargs):
skip_text = False
if kwargs.get("skip_text"):
kwargs.pop("skip_text")
skip_text = True
xlim, ylim = np.array([[-15,-4],[-15,-4]])
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=FutureWarning)
ax = sns.scatterplot(*args,**kwargs)
r2 = pearsonr(args[0],args[1])[0]**2
pad = 2
if not skip_text:
ax.text(-14,-6,f"$R^2$={r2:.2f}")
ax.plot(xlim,ylim,linewidth=2, color='blue')
ax.plot(xlim,ylim+pad,color="black",linestyle="--")
ax.plot(xlim,ylim-pad,color="black",linestyle="--")
ax.set(xlim=xlim,ylim=ylim)
# #### Putting Everything on One Plot (Please Don't Do This)
# Make a plot like the one in Wang et al. https://pubs.acs.org/doi/10.1021/ja512751q I find this plot very confusing, but I wanted to prove to myself that I could reproduce it.
sns.set(font_scale=2)
sns.set_style("white")
g = sns.FacetGrid(mdf,col="Method",hue="Target",col_wrap=2,height=8,legend_out=True)
g.map(draw_dG_facet,"Experimental_dG","Predicted_dG",skip_text=True,s=100)
_ = g.set_titles(col_template="{col_name}")
g.set_xlabels("Experimental $\Delta$G (kcal/mol)")
g.set_ylabels("Predicted $\Delta$G (kcal/mol)")
_ = g.add_legend()
# #### Trellising the Data
# I think a better way to plot this data is to trellis by target/method and to put the plots side by side. I also think it's important to display the $R^2$ for the correlations.
sns.set(font_scale=1)
sns.set_style("white")
g = sns.FacetGrid(mdf,col="DataSet",col_wrap=4)
g.map(draw_dG_facet,"Experimental_dG","Predicted_dG")
_ = g.set_titles(col_template="{col_name}")
g.set_xlabels("Experimental $\Delta$G (kcal/mol)")
_= g.set_ylabels("Predicted $\Delta$G (kcal/mol)")
# #### Calculating Confidence Intervals for Correlation Coefficients
# Another way to look at this data is to plot the correlations as bar plots. Of course the bar plots should have error bars. We can use this function to calculate the 95% confidence interval for the correlations.
def pearson_confidence(r, num, interval=0.95):
"""
Calculate upper and lower 95% CI for a Pearson r (not R**2)
Inspired by https://stats.stackexchange.com/questions/18887
:param r: Pearson's R
:param num: number of data points
:param interval: confidence interval (0-1.0)
:return: lower bound, upper bound
"""
stderr = 1.0 / math.sqrt(num - 3)
interval = interval + (1-interval)/2
z_score = norm.ppf(interval)
delta = z_score * stderr
lower = math.tanh(math.atanh(r) - delta)
upper = math.tanh(math.atanh(r) + delta)
return lower, upper
# This code just calculates Pearson r for each of the target/method combinations. I used the Pearson r rather than $R^2$ because I felt it made the differences more apparent in the plots.
gb = big_df.groupby("Target")
res = []
for k,v in gb:
num = len(v.Experimental_dG)
r_FEP = pearsonr(v.Experimental_dG,v.FEP_dG)[0]
r_TI = pearsonr(v.Experimental_dG,v.TI_dG)[0]
lb_FEP, ub_FEP = pearson_confidence(r_FEP,num)
lb_TI, ub_TI = pearson_confidence(r_TI,num)
# Currently calculate Pearson r, use the line below to get R**2
#res.append([k,len(v)]+[x**2 for x in [lb_FEP,r_FEP,ub_FEP,lb_TI,r_TI,ub_TI]])
res.append([k,len(v)]+[x for x in [lb_FEP,r_FEP,ub_FEP,lb_TI,r_TI,ub_TI]])
res_df = pd.DataFrame(res,columns=["Target","Num","FEP_LB","FEP Correlation","FEP_UB","TI_LB","TI Correlation","TI_UB"])
res_df
# #### Plotting Correlations With Error Bars
# In order to make the plots, we need to reshape the data.
melt_res_df = res_df[["Target","FEP Correlation","TI Correlation"]].melt(id_vars="Target")
melt_res_df.sort_values("Target",inplace=True)
melt_res_df
sns.set(rc={'figure.figsize':(12,12)},font_scale=1.5)
ax = sns.barplot(x="Target",y="value",hue="variable",data=melt_res_df)
x_pos = np.arange(8)-0.2
y_val = res_df["FEP Correlation"]
lb = y_val - res_df.FEP_LB
ub = res_df.FEP_UB - y_val
ax.errorbar(x_pos,y_val,yerr=[lb,ub],fmt="none",capsize=0,color='black')
x_pos = np.arange(8)+0.2
y_val = res_df["TI Correlation"]
lb = y_val - res_df.TI_LB
ub = res_df.TI_UB - y_val
a = ax.errorbar(x_pos,y_val,yerr=[lb,ub],fmt="none",capsize=0,color='black')
handles, labels = ax.get_legend_handles_labels()
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0., handles=handles[0:2], labels=labels[0:2])
_ = ax.set(xlabel='', ylabel="Pearson r")
# #### Calculating the Effect Size
# As [<NAME>](https://link.springer.com/article/10.1007/s10822-016-9904-5) points out, when we have a data set like this with dependent errors, the fact that error bars overlap does not necessarily imply that the methods are equivalent. However, we can use the mean of differences / std deviation of the differences to calculate [Cohen's d](https://machinelearningmastery.com/effect-size-measures-in-python/), which measures the effect size. Cohen’s d measures the difference between the mean from two Gaussian-distributed variables. It is a standard score that summarizes the difference in terms of the number of standard deviations. Because the score is standardized, there is a table for the interpretation of the result, summarized as:
#
# - Small Effect Size: d=0.20
# - Medium Effect Size: d=0.50
# - Large Effect Size: d=0.80
#
delta = res_df["FEP Correlation"].values-res_df["TI Correlation"].values
np.mean(delta)/np.std(delta)
# We get a Cohen's d of 1.2, which is a very large effect size. We can define a function to convert d to an approximate probability between 0 and 1.
def d_to_probability(d):
return 0.25 * abs(d) + 0.5
d_to_probability(1.2)
# So for the 8 datasets we examined, FEP should outperform TI in 8 of 10 cases. To look at this visually, let's plot a kernel density estimate for the Pearson r distributions.
sns.set(font_scale=1.5)
sns.set_style("white")
sns.kdeplot(res_df["FEP Correlation"],color="r",shade=True)
ax = sns.kdeplot(res_df["TI Correlation"],color="b",shade=True)
_ = ax.set(xlabel="Pearson r",ylabel="Density")
# We can also calculate Cohen's d for the RMSE.
gb = big_df.groupby("Target")
res = []
for k,v in gb:
res.append([k,mean_squared_error(v.Experimental_dG,v.FEP_dG),mean_squared_error(v.Experimental_dG,v.TI_dG)])
rmse_df = pd.DataFrame(res,columns=["Target","FEP RMSE","TI RMSE"])
delta = rmse_df["FEP RMSE"].values - rmse_df["TI RMSE"].values
np.mean(delta)/np.std(delta)
# Again we have a very large effect size. As above, we can covert this to a probability.
d_to_probability(-1.08)
| ml_models/comparing_regression_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # S3 equal temperament
#
# 3C6 Section 3: equal temperament and the major third
#
# ## imports and definitions
# +
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.animation as animation
matplotlib.rcParams.update({'font.size': 12,'font.family':'serif'})
from ipywidgets import interact
import time
import sounddevice as sd
# -
# %matplotlib notebook
# ## Setup parameters
# +
# Parameters
fs = 44.1e3
T = 6
Tc = 6
t = np.arange(0,T,1/fs)
z = np.zeros(np.int(T*fs/10))
# A and major third: equal temperament versus 5/4
f1 = 440
y1 = np.exp(-t/Tc) * np.sin(2*np.pi*f1*t)
y2 = np.exp(-t/Tc) * np.sin(2*np.pi*f1*5/4*t)
y3 = np.exp(-t/Tc) * np.sin(2*np.pi*f1*2**(4/12)*t)
y1 = np.concatenate((y1,z))/2
y2 = np.concatenate((z,y2))/2
y3 = np.concatenate((z,y3))/2
# -
# by exact ratio 5/4
sd.play(y1+y2,fs)
# by equal temperament:
sd.play(y1+y3,fs)
| S3_equal_temperament.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Predicting Credit Default
# +
# sudo pip install imblearn
# sudo pip install xgboost
# +
import pandas as pd
# ignore all future warnings
from warnings import simplefilter
simplefilter(action='ignore', category=FutureWarning)
from imblearn.over_sampling import SMOTE
from sklearn.preprocessing import StandardScaler #OneHotEncoder,
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
#LabelEncoder, label_binarize, StandardScaler, MinMaxScaler
from collections import defaultdict
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
import xgboost as xgb
import matplotlib.pyplot as plt
# -
#seed = 1234 # credit, age, duration
seed = 1234
# +
# https://archive.ics.uci.edu/ml/datasets/statlog+(german+credit+data)
col_names = names = ["checking_account", "duration", "credit_history", "purpose", "credit_amount",
"savings", "employment_since", "installment_rate", "status", "debtors_guarantors",
"residence", "property", "age", "other_installments", "housing",
"credits", "job", "dependents", "telephone", "foreign_worker", "credit"]
data_df = pd.read_csv("german.data",names = col_names, delimiter=' ')
data_df.head()
# -
# ## Feature engineering
# +
# Remap the target attribute: 1 - good credit, 0 - bad credit
data_df["credit"].replace([1,2], [1,0], inplace=True)
num_attr_names = ["duration", "credit_amount", "installment_rate", "residence",
"age", "credits", "dependents"]
cat_attr_names = ["checking_account", "credit_history", "purpose", "savings", "employment_since",
"status", "debtors_guarantors", "property", "other_installments", "housing",
"job", "telephone", "foreign_worker"]
num_attr_norm = pd.DataFrame(StandardScaler().fit_transform(data_df[num_attr_names]),
columns=num_attr_names)
num_attr_norm.head()
# +
dd = defaultdict(LabelEncoder)
cat_attr = data_df[cat_attr_names].apply(lambda col: dd[col.name].fit_transform(col))
cat_attr_dummy = pd.get_dummies(data_df[cat_attr_names])
cat_attr_dummy.head()
# -
clean_df = pd.concat([cat_attr_dummy, num_attr_norm, data_df["credit"]], axis = 1)
clean_df.head()
X = clean_df.loc[:, clean_df.columns != "credit"]
y = clean_df["credit"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=seed)
y_train.value_counts()
oversample = SMOTE(random_state=seed)
X_train, y_train = oversample.fit_resample(X_train, y_train)
y_train.value_counts()
# ## Model training
# change max_depth 15 -> 5 and look at feature importance
xgc = xgb.XGBClassifier(n_estimators=500, max_depth=15, base_score=0.5,
objective="binary:logistic", random_state=seed, use_label_encoder=False, eval_metric="logloss")
xgc.fit(X_train, y_train)
y_pred = xgc.predict(X_test)
print("Accuracy: %.2f" % accuracy_score(y_pred, y_test))
print("F1 score: %.2f" % f1_score(y_pred, y_test))
# ## Feature importance
xgb.plot_importance(xgc, importance_type="cover", max_num_features=10, show_values=False);
| MLCONF2021/CreditDefault.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Compare SQL phenotype to R phenotype
#
# <div class="alert alert-block alert-success">
# <b>For AoU there are some logic changes that will affect <i>which of a person's measurements</i> is used.</b> And <i>which measurement</i> is used will affect the <b>age</b>, since its age at time of measurement, and the <b>statin use indicator</b>, since the measurment must occur with in the statin use interval to be true.
# <ol>
# <li>AoU: We now retain only measurements where <kbd>value_as_number IS NOT NULL AND value_as_number > 0</kbd>.</li>
# <li>AoU: Previously the R code was modifying LDL during the lipids adjustment. Now LDL is the original value from the measurements table. Adjustments only occur within LDL_adjusted.
# <li>AoU: A single age and statin use indicator was previously chosen per person, even though those values could vary between a person's different lipid measurements. Now each measurement is retaining the age and statin use flag associated with the datetime of the measurment.</li>
# <li>AoU: When choosing the "most recent" measurement, the SQL code goes to greater lengths to make the result reproducible by sorting not only by measurement date, but also by measurement time, and measurement id in the case of ties.</li>
# <li>AoU: The SQL JOIN logic for measurements and statin use intervals uses the datetime instead of the date.</li>
# <li>UKB: 148 UKB samples were getting dropped incorrectly. I narrowed it down to the na.omit command being used to keep only people with all four lipids. Since na.omit is run on the entire dataframe, it checks other columns for NAs too such as the european ancestry column.</li>
# <li>UKB: the lipids adjustment is not the same formula, specifically the rule If TG > 400, then LDL = NA` was not applied to to ldladj in the natarajan dataframe provided.</li>
# </ol>
# </div>
# # Setup
lapply(c('hexbin', 'hrbrthemes', 'skimr', 'viridis'),
function(pkg) { if(! pkg %in% installed.packages()) { install.packages(pkg)} } )
library(hexbin)
library(hrbrthemes)
library(skimr)
library(tidyverse)
ORIG_R_PHENO <- c(
HDL = 'gs://fc-secure-fd6786bf-6c28-4f33-ac30-3860fbeee5bb/data/MergedData_HDL_Iteration2_ForGWAS.csv',
LDL = 'gs://fc-secure-fd6786bf-6c28-4f33-ac30-3860fbeee5bb/data/MergedData_LDL_Iteration2_ForGWAS.csv',
TC = 'gs://fc-secure-fd6786bf-6c28-4f33-ac30-3860fbeee5bb/data/MergedData_TC_Iteration2_ForGWAS.csv',
TG = 'gs://fc-secure-fd6786bf-6c28-4f33-ac30-3860fbeee5bb/data/MergedData_TG_Iteration2_ForGWAS.csv'
)
NEW_SQL_PHENO <- 'gs://fc-secure-fd6786bf-6c28-4f33-ac30-3860fbeee5bb/data/pooled/phenotypes/20211224/aou_alpha2_ukb_pooled_lipids_phenotype.tsv'
# +
# Set some visualiation defaults.
theme_set(theme_ipsum(base_size = 16)) # Default theme for plots.
#' Returns a data frame with a y position and a label, for use annotating ggplot boxplots.
#'
#' @param d A data frame.
#' @return A data frame with column y as max and column label as length.
get_boxplot_fun_data <- function(df) {
return(data.frame(y = max(df), label = stringr::str_c('N = ', length(df))))
}
# -
# # Load data
orig_hdl <- read_csv(pipe(str_glue('gsutil cat {ORIG_R_PHENO[["HDL"]]}')))
orig_ldl <- read_csv(pipe(str_glue('gsutil cat {ORIG_R_PHENO[["LDL"]]}')))
orig_tc <- read_csv(pipe(str_glue('gsutil cat {ORIG_R_PHENO[["TC"]]}')))
orig_tg <- read_csv(pipe(str_glue('gsutil cat {ORIG_R_PHENO[["TG"]]}')))
orig_pheno_wide <- orig_hdl %>%
full_join(orig_ldl) %>%
full_join(orig_tc) %>%
full_join(orig_tg) %>%
mutate(
FID = paste0(sampleid, '_', cohort),
IID = FID
)
# +
nrow(orig_pheno_wide)
length(unique(orig_pheno_wide$IID))
stopifnot(nrow(orig_pheno_wide) == length(unique(orig_pheno_wide$IID)))
# -
colnames(orig_pheno_wide)
new_pheno_wide = read_tsv(pipe(str_glue('gsutil cat {NEW_SQL_PHENO}')))
colnames(new_pheno_wide)
# # Compare data
dim(orig_pheno_wide)
dim(new_pheno_wide)
# <div class="alert alert-block alert-success">
# We've retained more non-zero and non-null measurements.
# </div>
# +
length(unique(orig_pheno_wide$IID))
length(unique(new_pheno_wide$IID))
nrow(new_pheno_wide) - nrow(orig_pheno_wide)
# -
new_pheno_wide %>%
filter(!IID %in% orig_pheno_wide$IID) %>%
group_by(cohort) %>%
summarize(count = n())
# <div class="alert alert-block alert-success">
# We've also included more genomes.
# </div>
# +
pheno_versions <- inner_join(
new_pheno_wide,
orig_pheno_wide,
suffix = c('_sql_phenotypes', '_r_phenotypes'),
by = c('FID', 'IID')
)
dim(pheno_versions)
# -
stopifnot(nrow(orig_pheno_wide) == nrow(pheno_versions))
colnames(pheno_versions)
# ## Check age
sum(abs(pheno_versions$age_sql_phenotypes - pheno_versions$age_r_phenotypes) > 2)
pheno_versions %>%
select(IID, age_r_phenotypes, age_sql_phenotypes) %>%
filter(age_sql_phenotypes - age_r_phenotypes > 2)
# ## Check cohort
table(pheno_versions$cohort_r_phenotypes, pheno_versions$cohort_sql_phenotypes)
# <div class="alert alert-block alert-success">
# The results are identical.
# </div>
# ## Check sex_at_birth
table(pheno_versions$sex, pheno_versions$sex_at_birth)
# <div class="alert alert-block alert-success">
# The results are identical.
# </div>
# ## Check PCs
skim(pheno_versions %>%
select(pc1, PC1, pc2, PC2, pc3, PC3, pc4, PC4, pc5, PC5, pc6, PC6, pc7, PC7, pc8, PC8, pc9, PC9, pc10, PC10))
# <div class="alert alert-block alert-success">
# The results are identical.
# </div>
# ## Check raw lipids
skim(pheno_versions %>%
filter(cohort_r_phenotypes == 'AOU') %>%
select(HDLraw, HDL, LDLraw, LDL,
TCraw, TC, TGraw, TG))
skim(pheno_versions %>%
filter(cohort_r_phenotypes == 'UKB') %>%
select(HDLraw, HDL, LDLraw, LDL,
TCraw, TC, TGraw, TG))
# <div class="alert alert-block alert-success">
# The results have minor differences, but no major differences.
# </div>
# ## Check adjusted lipids
skim(pheno_versions %>%
filter(cohort_r_phenotypes == 'AOU') %>%
select(HDLadj, HDL, LDLadj, LDL_adjusted,
TCadj, TC_adjusted, TGadj, TG_adjusted))
skim(pheno_versions %>%
filter(cohort_r_phenotypes == 'UKB') %>%
select(HDLadj, HDL, LDLadj, LDL_adjusted,
TCadj, TC_adjusted, TGadj, TG_adjusted))
# <div class="alert alert-block alert-success">
# The results have minor differences, but no unexpected major differences. (It is expected that we have more NA values for LDL_adjusted.)
# </div>
# ## Check normalized lipids
skim(pheno_versions %>%
filter(cohort_r_phenotypes == 'AOU') %>%
select(HDLnorm, HDL_norm, LDLnorm, LDL_adjusted_norm,
TCnorm, TC_adjusted_norm, TGnorm, TG_adjusted_norm))
skim(pheno_versions %>%
filter(cohort_r_phenotypes == 'UKB') %>%
select(HDLnorm, HDL_norm, LDLnorm, LDL_adjusted_norm,
TCnorm, TC_adjusted_norm, TGnorm, TG_adjusted_norm))
# <div class="alert alert-block alert-success">
# The results have minor differences, but no major differences.
# </div>
# # Provenance
devtools::session_info()
| aou_workbench_pooled_analyses/phenotype_exploration/compare_sql_phenotype_to_r_phenotype.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Target Connectivity
# # Configurable logging system
# All LISA modules have been updated to use a more consistent logging which can be configured using a single configuraton file:
# !head -n12 $LISA_HOME/logging.conf
# Each module has a unique name which can be used to assign a priority level for messages generated by that module.
# !head -n30 $LISA_HOME/logging.conf | tail -n5
# The default logging level for a notebook can also be easily configured using this few lines
# + run_control={"marked": false}
import logging
from conf import LisaLogging
LisaLogging.setup(level=logging.INFO)
# -
# ## Removed Juno/Juno2 distinction
# Juno R0 and Juno R2 boards are now accessible by specifying "juno" in the target configuration.
# The previous distinction was required because of a different way for the two boards to report HWMON channels.
# This distinction is not there anymore and thus Juno boards can now be connected using the same platform data.
# +
from env import TestEnv
te = TestEnv({
'platform' : 'linux',
'board' : 'juno',
'host' : '10.1.210.45',
'username' : 'root'
})
target = te.target
# -
# # Executor Module
#
# ## Simplified tests definition using in-code configurations
# Automated LISA tests previously configured the Executor using JSON files. This is still possible, but the existing tests now use Python dictionaries directly in the code. In the short term, this allows de-duplicating configuration elements that are shared between multiple tests. It will later allow more flexible test configuration.
#
# See `tests/eas/acceptance.py` for an example of how this is currently used.
# ## Support to write files from Executor configuration
# https://github.com/ARM-software/lisa/pull/209
#
# A new "files" attribute can be added to Executor configurations which allows
# to specify a list files (e.g. sysfs and procfs) and values to be written to that files.
#
# For example, the following test configuration:
tests_conf = {
"confs" : [
{
"tag" : "base",
"flags" : "ftrace",
"sched_features" : "NO_ENERGY_AWARE",
"cpufreq" : {
"governor" : "performance",
},
"files" : {
'/proc/sys/kernel/sched_is_big_little' : '0',
'!/proc/sys/kernel/sched_migration_cost_ns' : '500000'
},
}
]
}
# can be used to run a test where the platform is configured to
# - disable the "sched_is_big_little" flag (if present)
# - set to 50ms the "sched_migration_cost_ns"
#
# Nortice that a value written in a file is verified only if the file path is
# **prefixed** by a '/'. Otherwise, the write never fails, e.g. if the file does not exists.
# ## Support to freeze user-space across a test
# https://github.com/ARM-software/lisa/pull/227
#
# Executor learned the `"freeze_userspace"` conf flag. When this flag is present, LISA uses the devlib freezer to freeze as much of userspace as possible while the experiment workload is executing, in order to reduce system noise.
#
# The `Executor` example notebook:
#
# https://github.com/ARM-software/lisa/blob/master/ipynb/examples/utils/executor_example.ipynb
#
# gives an example of using this feature.
# # Trace module
# ## Tasks name pre-loading
# When the Trace module is initialized, by default all the tasks in that trace are identified and exposed via the usual getTask() method:
# +
from trace import Trace
import json
with open('/home/patbel01/Code/lisa/results/LisaInANutshell_Backup/platform.json', 'r') as fh:
platform = json.load(fh)
trace = Trace('/home/patbel01/Code/lisa/results/LisaInANutshell_Backup/trace.dat',
['sched_switch'], platform
))
# -
logging.info("%d tasks loaded from trace", len(trace.getTasks()))
logging.info("The rt-app task in this trace has these PIDs:")
logging.info(" %s", trace.getTasks()['rt-app'])
# # Android Support
# ## Added support for Pixel Phones
# A new platform definition file has been added which allows to easily setup
# a connection with an Pixel device:
# !cat $LISA_HOME/libs/utils/platforms/pixel.json
# +
from env import TestEnv
te = TestEnv({
'platform' : 'android',
'board' : 'pixel',
'ANDROID_HOME' : '/home/patbel01/Code/lisa/tools/android-sdk-linux/'
}, force_new=True)
target = te.target
# -
# ## Added UiBench workload
# A new Android benchmark has been added to run UiBench provided tests.
# Here is a notebook which provides an example of how to run this test on your
# android target:
#
# https://github.com/ARM-software/lisa/blob/master/ipynb/examples/android/benchmarks/Android_UiBench.ipynb
# # Tests
# ## Intial version of the preliminary tests
# Preliminary tests aim at verifying some basic support required for a
# complete functional EAS solution.
#
# A initial version of these preliminary tests is now available:
#
# https://github.com/ARM-software/lisa/blob/master/tests/eas/preliminary.py
#
# and it will be extended in the future to include more and more tests.
# ## Capacity capping test
# A new test has been added to verify that capacity capping is working
# as expected:
#
# https://github.com/ARM-software/lisa/blob/master/tests/eas/capacity_capping.py
#
# ## Acceptance tests reworked
# The EAS acceptace test collects a set of platform independent tests to verify
# basic EAS beahviours.
#
# This test has been cleaned up and it's now avaiable with a detailed documentation:
#
# https://github.com/ARM-software/lisa/blob/master/tests/eas/acceptance.py
#
# # Notebooks
# ## Added scratchpad notebooks
# A new **scratchpad** folder has been added under the ipynb folder which collects the available notebooks:
# !tree -L 1 ~/Code/lisa/ipynb
# This folder is configured to be ignored by git, thus it's the best place to place your work-in-progress notebooks.
# ## Example notebook restructoring
# Example notebooks has been consolidated and better organized by topic:
# !tree -L 1 ~/Code/lisa/ipynb/examples
# This is the folder to look into when it comes to undedrstand how a specific
# LISA API works.
#
# Here is where we will provide a dedicated folder and set of notebooks for each of the main LISA modules.
| ipynb/deprecated/releases/ReleaseNotes_v16.12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="MjG6VZL35yNF"
# # OPTION A
# + [markdown] id="t2xlS3sR6Oy3"
# ### Present the code and methods for acquiring the data. Loading the data into appropieate format for analysis. Explain the process and results
# + [markdown] id="zTgB7pwc6es6"
# #### At the beginning we must import so numpy as panda libraries
# + id="hievc64p4CbB"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="2n7K1MWr6z24"
# #### We are importing csv file from my drive on google
#
# + id="bqctHK0w4VYT"
df = pd.read_csv("AB_NYC_2019.csv")
# + [markdown] id="oPyDeBb38gv-"
# #### To be sure dataset has been created successfully we can check the first terms of it. To do that let us use next attribute of dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="3qQmEx6C9CHh" outputId="54d117b2-6d86-46e0-c5dd-20b614237117"
df.head()
# + [markdown] id="UbZDEBcS9n7O"
# #### We are checking the name of the categories of tha dataset
# + colab={"base_uri": "https://localhost:8080/"} id="2ikXSHdL4411" outputId="86b1d804-9b99-4e4e-a1ee-6dd856ba8867"
columns = df.columns
print(columns)
categories_number = len(columns)
print("Number of categories: ", categories_number)
# + [markdown] id="S_bs71LhFge9"
# #### Let us see how many rows dataset has right now
# + colab={"base_uri": "https://localhost:8080/"} id="HTm9ScHWFda9" outputId="1f83315c-7c21-4ce0-de31-a6c815f5c18f"
df.shape
# + [markdown] id="gZR9i4wLCV9a"
# #### Let us set up how many rows will be shown when we use methods like head(), tail(), describe()...
# + id="uogTMVG-C9oT"
pd.options.display.max_rows = 20
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="AEBN8oIHDDq8" outputId="292fdbb9-4ea3-41e9-fda6-615e245f66d0"
df.head()
# + [markdown] id="2b7BHAhyOX7g"
# #### Let us check how many NotANumber are present in dataset
# + colab={"base_uri": "https://localhost:8080/"} id="KLvHcagdPvHT" outputId="450d26f1-bd92-440b-cc15-5c27ea729c8c"
not_a_number = df.isna().sum()
print(not_a_number)
# + [markdown] id="Yov9NatNQKRF"
# #### We can verify only name, host_name, last_review and reviews_per_month contain NAN elements. To solve that situation we may replace the previous terms that are equal to NAN by 0
# + [markdown] id="mXShkroRiJuz"
# #### It does not matter whether either name or host_name have some terms equal to NAN, what is to say, we do not need to fix them up. On the other hand we might try to replace NAN terms presented in last_review, the best way to do this would be purge all NAN terms because this category is a date category and we do not have any way to guess or interpolate those NAN terms
# + colab={"base_uri": "https://localhost:8080/", "height": 479} id="yYOLdo1-jqeK" outputId="2b73666e-2b93-4d0d-9f94-7e9d6f18d638"
df_cleaned = df.dropna()
df_cleaned.head()
# + [markdown] id="1Jris7axuHWK"
# #### Let us check whether there is any NAN term in dataframe
# + colab={"base_uri": "https://localhost:8080/"} id="XeqrAyEbuOU2" outputId="87d6f968-6c7c-4738-bac8-3052838dfade"
df_cleaned.isna().sum()
# + [markdown] id="ylyo_eZivbu1"
# #### Let us start with latitude category
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="qZViGFHYvi29" outputId="75b76686-8829-4f79-dcc7-4ff185b16653"
new_len = df_cleaned.shape[0]
x_axis = np.array(range(1,new_len+1))
y_axis = df_cleaned["latitude"]
y_max = df_cleaned["latitude"].max()
y_min = df_cleaned["latitude"].min()
print(f"Type of y-axis: {y_axis.dtype}")
print(f"Latitude: y_max: {y_max} || y_min: {y_min}")
plt.plot(x_axis, y_axis)
# + [markdown] id="xUYqX5BCyMWG"
# #### We have just checked latitude values are within [40.9306, 40.50605] interval. There is not any singular value in latitude category in order to type of y_axis is float64. If there were any non float64 type term, the type of y_axis would be object.
# + [markdown] id="LGNLxQBKy9bA"
# #### We are proceeding in the same way as before for longitude category
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="zRQ1Q-H0zdYw" outputId="b4b359eb-7928-4012-f616-aa2f65e1b40f"
y_axis = df_cleaned["longitude"]
y_max = df_cleaned["longitude"].max()
y_min = df_cleaned["longitude"].min()
print(f"Type of y_axis: {y_axis.dtype}")
print(f"Longitude: y_max: {y_max} || y_min: {y_min}")
plt.plot(x_axis, y_axis)
# + [markdown] id="T9gXovYN9yK5"
# #### In this case, we can affirm type of y_axis is float64 too, therefore any value of longitude category belongs to [-73.713, -74.25] interval
# + [markdown] id="RF21Ltld-Ll-"
# #### Let us proceed with price category. We can confirm any singular value is present in this category
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="7lk-Ak4S_0xf" outputId="bba34c41-a226-4d1d-b1f1-7ad0479629d9"
y_axis = df_cleaned["price"]
y_max = df_cleaned["price"].max()
y_min = df_cleaned["price"].min()
print(f"Type of y_axis is: {y_axis.dtype}")
print(f"Price: y-max: {y_max} || y_min: {y_min}")
plt.plot(x_axis, y_axis)
# + colab={"base_uri": "https://localhost:8080/"} id="8-zMBxxRH1Gt" outputId="1c78fdc8-77c7-45bc-95ed-bd50b31460b5"
df_cleaned.loc[df_cleaned.price > 1500,:].value_counts().sum()
# + [markdown] id="lD_FTen7Ib81"
# #### We have just checked that there are only 58 items whose values are greater then 1500$. They are going to su deleted because they can be considered as outliers.
# + id="YdXgquuTJBAX"
df_cleaned = df_cleaned.loc[df_cleaned.price < 1500,:]
# + [markdown] id="s0lYh52CBvT_"
# #### Let us proceed with minimum_nights
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="pc_Xca8MCAZC" outputId="3c2b05ec-2d22-4921-be66-8ad2bea6243e"
x_axis = np.array(range(df_cleaned.shape[0]))
y_axis = df_cleaned["minimum_nights"]
y_max = df_cleaned["minimum_nights"].max()
y_min = df_cleaned["minimum_nights"].min()
print(f"Type of df_cleaned.minimum_nights is: ",y_axis.dtype)
print(f"Minimum_nights: y-max: {y_max}|| y-min: {y_min}")
plt.plot(x_axis, y_axis)
# + [markdown] id="NmgAxKPjC3sq"
# #### In this case we can affirm there are three singular values at least. Every value is lower than 600. We are removing these singular rows from dataframe
# + colab={"base_uri": "https://localhost:8080/"} id="Z1IkFfR5Dds2" outputId="bb654360-fd3c-483a-dd6e-b41b5a115b7e"
df_minimum_nights_high = df_cleaned.loc[df_cleaned.minimum_nights >= 800,:]
print(df_minimum_nights_high.minimum_nights)
df_cleaned_2 = df_cleaned.loc[df_cleaned.minimum_nights < 800,:]
print(df_cleaned_2)
# + [markdown] id="FRFXi4vEFzJY"
# #### We do affirm there were only three unusual values of minimum_nights because we had 38752 rows in df_cleaned, we have determined there are only three values in minimum_nights category over 800.
# + [markdown] id="-da4-_tMa-c2"
# #### Let us proceed with number_of_reviews category. In this case we must create another x_axis with whose length must be equals to 38818
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="w-MgjcqXbLWb" outputId="632b2f2e-0ec9-4fe0-8f94-83507bf7bf67"
new_len = df_cleaned_2.shape[0]
x_axis = np.array(range(1, new_len+1))
y_axis = df_cleaned_2["number_of_reviews"]
y_max = df_cleaned_2["number_of_reviews"].max()
y_min = df_cleaned_2["number_of_reviews"].min()
print(f"Type of y-axis: ", y_axis.dtype)
print(f"Number of reviews: y-max: {y_max} || y-min: {y_min}")
plt.plot(x_axis, y_axis)
# + [markdown] id="nfwwat0IfGtg"
# #### In this case, we can affirm there is not any singular value in number_of_reviewes category, because its dtype is int64 what means every value in this category is int64 dtype. Moreover every value of this category belongs to [1, 629] interval. We also do affirm values of this category are in decline
# + [markdown] id="XnceRRtWjDKW"
# #### Let us start with reviews_per_month
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="rDd-Zb7hkDPA" outputId="999ea661-6e1b-43ea-931e-522cc0979c1c"
y_axis = df_cleaned_2["reviews_per_month"]
y_max = df_cleaned_2["reviews_per_month"].max()
y_min = df_cleaned_2["reviews_per_month"].min()
print(f"Reviews-per-month: y-max: {y_max} || y-min: {y_min}")
print(f"dtype of y_axis: {y_axis.dtype}")
plt.plot(x_axis, y_axis)
# + [markdown] id="Vy5EWf6AlnZV"
# #### We can check it seems to show a singular value what might alters the results.One way to remove this singular value may be
# + colab={"base_uri": "https://localhost:8080/"} id="lwVGMxDPnY9K" outputId="084d84c6-441e-4c88-d257-fa4eb061dab2"
df_cleaned_2_dropped = df_cleaned_2.loc[df_cleaned_2.reviews_per_month > 50,:]
print(len(df_cleaned_2_dropped))
# + [markdown] id="b6Gm49-hoSBb"
# #### There is only one singular value in this category.
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="ncH2o2Qhow6w" outputId="d79fb9d5-19af-4489-c0e6-6cf94d24ae5a"
df_cleaned_3 = df_cleaned_2.loc[df_cleaned_2.reviews_per_month <= 50,:]
new_len = df_cleaned_3.shape[0]
x_axis = np.array(range(1,new_len+1))
y_axis = df_cleaned_3["reviews_per_month"]
y_max = df_cleaned_3["reviews_per_month"].max()
y_min = df_cleaned_3["reviews_per_month"].min()
print(f"dtype of reviews_per_month_dropeed: ", y_axis.dtype)
print(f"reviews_per_month_dropped: y-max: {y_max} || y-min: {y_min}")
plt.plot(x_axis, y_axis)
# + [markdown] id="gSe3bBwUrA5B"
# #### Values in this category are in increase
# + [markdown] id="Z-oR60MLrRh8"
# #### Let us start with calculated_host_listings_count
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="X6epaQdwsKQO" outputId="c138fb74-e911-4c04-cc59-b046f6d326e0"
y_axis = df_cleaned_3["calculated_host_listings_count"]
y_max = df_cleaned_3["calculated_host_listings_count"].max()
y_min = df_cleaned_3["calculated_host_listings_count"].min()
print(f"dtype of y_axis: ", y_axis.dtype)
print(f"calculated_host_listings_count: y-max: {y_max} || y-min: {y_min}")
plt.plot(x_axis, y_axis)
# + [markdown] id="3cdyHzG9t15y"
# #### This picture does not show any singular value
# + [markdown] id="xyeDvYPEvNUf"
# #### Let us start with availability_365 category
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="vRcOOd6Qv5bx" outputId="cc0d03c2-2f3b-434d-ddb6-65dc6da0734b"
y_axis = df_cleaned_3["availability_365"]
y_max = df_cleaned_3["availability_365"].max()
y_min = df_cleaned_3["availability_365"].min()
print(f"dtype of df_cleaned_3.availability_365: ", y_axis.dtype)
print(f"df_cleaned_3.availability_365: y-max: {y_max} || y-min: {y_min}")
plt.plot(x_axis, y_axis)
# + [markdown] id="2FVJUL-CxaaR"
# #### This category does not seem to show any singular data either
# + [markdown] id="HQ4AblRZzozC"
# ### In this point we have cleaned the initial dataset. We are checking the correlation matrix of df_cleaned_3
# + colab={"base_uri": "https://localhost:8080/"} id="lGVFNBF20cOS" outputId="aa23b192-0b4d-4668-9303-73ffc9ebc467"
corr_df = df_cleaned_3.corr(method="pearson")
print(corr_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 275} id="naZ0MI7L1cAq" outputId="949da196-26f6-46a4-e92d-def4c0004057"
plt.matshow(corr_df)
plt.show()
# + [markdown] id="Mm09B9vX4LUJ"
# #### We can check the correlations coefficients betweent categories. We could use heatmap from seaborn library to have a similar graphic with all correlation coefficients as legends
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="fccO3IZs3vZC" outputId="5b3cd7b0-5794-4e5e-e37f-f6730a8f82c7"
import seaborn as sns
sns.heatmap(corr_df, annot=True)
# + [markdown] id="dfhWAaygkbXO"
# #### The next grapich does take over 1 minuto to be shown, it consists in every correlation scatter plot for avery couple of cartegories that are present in df_cleaned_3. Thus we can appreciate which category has more influence over price values
# + id="uLwSb7urjYyu"
#sns.pairplot(df_cleaned_3, size=2.5)
# + [markdown] id="WN5770RiApdp"
# #### We also can have a similar graphic using attributes of df_Corr object
# + colab={"base_uri": "https://localhost:8080/", "height": 250} id="KA-HKoU5AkDb" outputId="db6a7680-dc3c-40f7-9ef4-108d1d4d96b0"
corr_df.style.background_gradient(cmap='coolwarm')
# + [markdown] id="6lCI8Fo7B-0B"
# #### There are some categories having correlations with each other. These caterories are:
# - id | host_id
# - number_of_reviews | reviews_per_month
# + [markdown] id="uOP0dsnLEpJT"
# #### The only correlation that might me interesting is the second one. To see more in details we can try to get the positive correlation existing between number_of_reviews and reviews_per_month categories
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="xF1Y2sHiGAjz" outputId="7baa183a-1d01-460d-de74-971554215cb9"
x_axis = df_cleaned_3["reviews_per_month"]
y_axis = df_cleaned_3["number_of_reviews"]
plt.scatter(x_axis, y_axis)
# + [markdown] id="E3cCeislKJ-_"
# #### As picture above shows, all values of number_of_reviews are distributed within two wrapping lines
# + [markdown] id="NAxzKo8-u5VA"
# #### To get both wrapping lines we must manipulate last values of x_axis and y_axis. We are getting a subsample of y_axis and x_axis whose values are whithin the interval defining by df_cleaned_3.loc[df_cleaned_3.number_of_reviews <= 20]...
# + colab={"base_uri": "https://localhost:8080/", "height": 316} id="O2Xpkj3dvc9s" outputId="a640af2d-a301-4a26-9578-b1d776f56c5c"
from sklearn import linear_model
x_subsample = df_cleaned_3.loc[df_cleaned_3.number_of_reviews <= 20, "reviews_per_month"]
y_subsample = df_cleaned_3.loc[df_cleaned_3.number_of_reviews <= 20, "number_of_reviews"]
print("len subsample: ", len(x_subsample))
plt.scatter(x_subsample, y_subsample)
regr = linear_model.LinearRegression()
x_train = np.array(x_subsample).reshape(-1,1)
y_train = np.array(y_subsample).reshape(-1,1)
regr.fit(x_train, y_train)
y_pred = regr.predict(x_train)
plt.plot(x_train, y_pred)
print("Coeficient: ", regr.coef_)
print("Intercept: ", regr.intercept_)
# + [markdown] id="5WBiVUwyRVvp"
# #### We already know the expression or the lower wrapping liner:
# *y(x) = 1.456 x + 4.9949*
# + [markdown] id="o9NbhnCSSnyq"
# #### However we must keep in mind we have got the expression of lower wrapping line based on a subsample of df_3_cleaned, what is to say, the mathematical expression is based on an x_axis whose length is 25581. On the contrary df_cleaned_3 has 38817 rows.
#
# + [markdown] id="0Z_jZA1-UFhz"
# In short, the expression of lower wrapping line might not be strictly correct but it seems to fit properly to df_cleaned_3
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="zDU7S2n2Uo8c" outputId="a138e316-cfbc-46b8-d5fd-2d8a4881f018"
x_axis = df_cleaned_3["reviews_per_month"]
y_axis = df_cleaned_3["number_of_reviews"]
plt.scatter(x_axis, y_axis)
y_lwl = regr.coef_[0][0] * x_axis + regr.intercept_[0]
plt.plot(x_axis, y_lwl)
# + [markdown] id="4uo143-vXlHM"
# #### Let us analyze price category, we may consider to use either a *technique of machine learning* or even some *kind of neural network*, but first of all we should know the behaviour of values of this category. To summarize, we must know whether values of price category are randomly distributed, to know that we must apply a goodness of fit test to price category.
# + [markdown] id="uLUm_uiMTD9N"
# #### We must determine CFD function of this category
# + [markdown] id="rXHdvWKTUJCG"
# #### We are taking values of the price category in order to try to make some predicictions
# + [markdown] id="Q5Y-WbIuXcOz"
# #### Let us analyze price category, we may consider to use either a *technique of machine learning* or even some *kind of neural network*, but first of all we should know the behaviour of values of this category. To summarize, we must know whether values of price category are randomly distributed, to know that we must apply a goodness of fit test to price category.
# + [markdown] id="xdoj5WJuXeHc"
# #### We must determine Cumulative Frecuency Distribution (CFD) of price values
# + colab={"base_uri": "https://localhost:8080/", "height": 707} id="y0FQh83wVGeZ" outputId="96074375-b90c-4eb9-91eb-72825b89251c"
#cfd = pd.DataFrame(x_train).value_counts().cumsum() / 12400
cfd = df_cleaned_3.loc[:,"price"].value_counts().cumsum() / 38751
print(cfd)
print(cfd)
x_axis = np.array(range(0, len(cfd)))
print(len(cfd))
plt.plot(x_axis, cfd)
# + colab={"base_uri": "https://localhost:8080/"} id="Hq8sK1ixWVNk" outputId="cac60f65-eab7-4f52-a4a3-f6313e91e365"
from scipy.stats import anderson, kstest
ad_test = anderson(cfd, dist="norm")
ks_test = kstest(cfd, "norm", N=len(cfd))
print("Test: ", ad_test)
print("ks_test: ", ks_test)
# + [markdown] id="-B007Bn1X83q"
# #### We must take in consideration critical statistics values of Kolmogorov-Smirnov (KS) test. If n is the size of the sample and is greater than 35:
# - alpha = 0.2 ---> s* = 1.07 / n^0.5 --> 4.85e-2
# - alpha = 0.15 --> s* = 1.14 / n^0.5 --> 5.17e-2
# - alpha = 0.1 ---> s* = 1.22 / n^0.5 --> 5.53e-2
# - alpha = 0.05 --> s* = 1.36 / n^0.5 --> 6.17e-2
# - alpha = 0.01 --> s* = 1.63 / n^0.5 --> 7.39e-2
#
# alpha would be the significance_level of results of Anderson-Darling test
# + [markdown] id="nbz9VYqkbaEo"
# #### In this point we can consider two hypothesis:
# - Ho : price values follow a Normal distribution
# - Hi : price values do not follow a Normal distribution
#
# In Anderson-Darling test we have got a statistic value which is greater than every of its *critical_values*, moreover in case of Kolmogorov-Smirnov test we also have got a statistic value which is greater than every of its s* (critical_values) with p-value = 2.49e-249
#
# As a consecuence we do affirm we have not enough information to reject Hi, in addition we also do affirm *price category does not follow a Normal distribution*
# + [markdown] id="LPCEySEveoFb"
# #### As a previous result we can consider to use some kinf of predictor, for instance a predictor based on cross-validation methods
# + [markdown] id="M8uDLV_VpG_2"
# #### In this point we should know what categories seems to have more influence over price values. To get these categories we have already used a correlogram-table, so in this case
# + [markdown] id="onOnpmSl2ZZy"
# #### Checking neighbourhood category in dataframe we can see there are three principal groups: Manhattan, Brooklyn and Queens. Instead of trying to make an estimation of rental price in NY we can split up the problem into three parts.
# #### In short, we are doing three models
# + [markdown] id="VyV26l2k3rq1"
# ##Manhattan
# + colab={"base_uri": "https://localhost:8080/", "height": 250} id="zXu41rVZmpCZ" outputId="7409f36a-9a96-4f37-d7f3-f02d207294e9"
df_manhattan = df_cleaned_3.loc[df_cleaned_3.neighbourhood_group == "Manhattan", :]
corr_manhattan = df_manhattan.corr(method="pearson")
corr_manhattan.style.background_gradient(cmap='coolwarm')
# + [markdown] id="EwL5CmyiqAtq"
# #### The categories have more influence on price are: longitude, latitude and availability_365.
# #### Both of them have a negative correlation coefficient respect to price
# + colab={"base_uri": "https://localhost:8080/"} id="XVtZLmjIrFCM" outputId="b491d70d-ec92-454c-9b23-540b82819252"
from sklearn.model_selection import train_test_split
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
features = df_manhattan.loc[:,["latitude", "longitude", "availability_365"]]
target = df_manhattan.loc[:,"price"]
scaler = MinMaxScaler(feature_range=(0,1))
prices_man = np.array(df_manhattan.loc[:,"price"]).reshape(-1,1)
latitude_man = np.array(df_manhattan.loc[:,"latitude"]).reshape(-1,1)
longitude_man = np.array(df_manhattan.loc[:,"longitude"]).reshape(-1,1)
availability_man = np.array(df_manhattan.loc[:,"availability_365"]).reshape(-1,1)
prices_min = prices_man.min()
prices_max = prices_man.max()
prices_n_man = scaler.fit_transform(prices_man)
latitude_n_man = scaler.fit_transform(latitude_man)
longitude_n_man = scaler.fit_transform(longitude_man)
availability_n_man = scaler.fit_transform(availability_man)
new_len = len(prices_man)
features_matrix = np.zeros((new_len, 4))
features_matrix[:,0] = latitude_n_man[:,0]
features_matrix[:,1] = longitude_n_man[:,0]
features_matrix[:,2] = availability_n_man[:,0]
features_matrix[:,3] = prices_n_man[:,0]
features_df_man = pd.DataFrame(features_matrix)
features_df_man.columns = ["latitude", "longitude", "availability_365", "price"]
features_n_man = features_df_man.loc[:,["latitude", "longitude", "availability_365"]]
target_n_man = features_df_man.loc[:,"price"]
print(features_df_man)
# + id="qpULbMrByew_"
X_train, X_test, y_train, y_test = train_test_split(features_n_man, target_n_man, test_size=0.25, random_state=42)
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="jSaK6akRM3Ow" outputId="6ac87b31-ceb8-4403-ce53-abb457c1882f"
X_train
# + colab={"base_uri": "https://localhost:8080/"} id="eOXewMoIVWPX" outputId="b89271b7-e889-478b-8e82-b6cd06bb5aaf"
from keras.models import Sequential
from keras.layers import Dense, Dropout
from tensorflow.keras.optimizers import RMSprop, Adam
model = Sequential()
model.add(Dense(128, activation="sigmoid"))
model.add(Dropout(0.3))
model.add(Dense(64, activation="sigmoid"))
model.add(Dropout(0.2))
model.add(Dense(32, activation="sigmoid"))
model.add(Dropout(0.25))
model.add(Dense(1, activation="sigmoid"))
my_optimizer = Adam(learning_rate=0.001)
model.compile(loss="mean_squared_error", optimizer= my_optimizer ,metrics=["mean_absolute_error", "mean_squared_error"])
model.fit(X_train, y_train, epochs=100, batch_size=50)
# + id="hGfqA9iOMfEd"
y_pred = model.predict(X_test)
# + id="bOd4hs8PAf4j"
y_test_np = np.array(y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="dKpNZaNPCEEB" outputId="a015ca1c-4475-4af2-be76-b5f5b857c817"
x_axis = np.array(range(len(y_test_np)))
x_axis.size
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="J-qHpfbkCQvk" outputId="a029c5d7-7bba-4187-e23a-ad48a72c1520"
plt.scatter(x_axis, y_test, label="real")
plt.scatter(x_axis, y_pred[:,0], label="estimado")
plt.title("Indicador de precio MinMaxScaled", fontsize=10, color="SteelBlue", fontweight="bold")
plt.ylabel("Indicador de precio MinMaxScaled estimado y real", fontsize=7, color="SteelBlue", fontweight="bold")
plt.legend()
# + colab={"base_uri": "https://localhost:8080/"} id="DdXd9aRTzBcE" outputId="50ab5842-3d5a-4872-d38f-dfa7063981f8"
y_pred
# + colab={"base_uri": "https://localhost:8080/", "height": 766} id="z0Uf1hvONRLK" outputId="12946191-5b32-4a11-bf6a-e4292300e6fe"
price_test = y_test * (prices_max - prices_min) + prices_min
price_pred = y_pred * (prices_max - prices_min) + prices_min
print(type(price_test))
print(type(price_pred))
price_test_np = np.array(price_test)
print(type(price_test_np))
print(price_test_np)
print(np.array(price_pred))
error = abs(price_pred[:,0] - price_test)
x_axis = np.array(range(len(price_pred)))
plt.scatter(x_axis, error)
plt.title("abs. Error Precio estimado - Precio real sobre x_test",fontsize=10,color="SteelBlue",fontweight="bold")
plt.ylabel("Error en valor absoluto de precio estimado y precio real en $", color="SteelBlue", fontweight="bold")
error
# + colab={"base_uri": "https://localhost:8080/"} id="8dA3-q4Y9Orr" outputId="10379499-eced-42fb-edaa-a382e3d56e42"
y_test
# + colab={"base_uri": "https://localhost:8080/"} id="ottgbAKv9Qhw" outputId="a10a65b2-d82e-4bda-940d-9397e385e882"
y_pred
# + colab={"base_uri": "https://localhost:8080/"} id="y7Aw-kCBCYP3" outputId="1c478b88-7f9c-43ed-9403-07bdce888ef5"
error.describe()
# + [markdown] id="ZRNHf-Nd3wA6"
# ##Brooklyn
# + colab={"base_uri": "https://localhost:8080/", "height": 250} id="gR_GeSvy377i" outputId="a5f0de2f-a0d4-4166-8204-17ead94672ca"
df_brooklyn = df_cleaned_3.loc[df_cleaned_3.neighbourhood_group == "Brooklyn", :]
corr_brooklyn = df_brooklyn.corr(method="pearson")
corr_brooklyn.style.background_gradient(cmap='coolwarm')
# + [markdown] id="Lv2Nkb5I4FgS"
# #### We can see in the correlation matrix above the categories that has more influence in price values are: latitude, longitude and availability_365
# + colab={"base_uri": "https://localhost:8080/"} id="X4SHHsfB44Vm" outputId="2ab8ccda-f170-4608-b0d1-6ae71124a76d"
from sklearn.model_selection import train_test_split
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
features_bro = df_brooklyn.loc[:,["latitude", "longitude", "availability_365"]]
target_bro = df_brooklyn.loc[:,"price"]
scaler_bro = MinMaxScaler(feature_range=(0,1))
prices_bro = np.array(df_brooklyn.loc[:,"price"]).reshape(-1,1)
latitude_bro = np.array(df_brooklyn.loc[:,"latitude"]).reshape(-1,1)
longitude_bro = np.array(df_brooklyn.loc[:,"longitude"]).reshape(-1,1)
availability_bro = np.array(df_brooklyn.loc[:,"availability_365"]).reshape(-1,1)
prices_min_bro = prices.min()
prices_max_bro = prices.max()
prices_n_bro = scaler_bro.fit_transform(prices_bro)
latitude_n_bro = scaler_bro.fit_transform(latitude_bro)
longitude_n_bro = scaler_bro.fit_transform(longitude_bro)
availability_n_bro = scaler_bro.fit_transform(availability_bro)
new_len = len(prices_bro)
features_matrix_bro = np.zeros((new_len, 4))
features_matrix_bro[:,0] = latitude_n_bro[:,0]
features_matrix_bro[:,1] = longitude_n_bro[:,0]
features_matrix_bro[:,2] = availability_n_bro[:,0]
features_matrix_bro[:,3] = prices_n_bro[:,0]
features_df_bro = pd.DataFrame(features_matrix_bro)
features_df_bro.columns = ["latitude", "longitude", "availability_365", "price"]
features_n_bro = features_df_bro.loc[:,["latitude", "longitude", "availability_365"]]
target_n_bro = features_df_bro.loc[:,"price"]
print(features_df_bro)
# + id="3NoIXvRn5YZe"
X_train_bro, X_test_bro, y_train_bro, y_test_bro = train_test_split(features_n_bro, target_n_bro, test_size=0.25, random_state=27)
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="p3A7O7K_MaRO" outputId="6e64cc3d-02d4-4561-d4d7-3ee9438c2bf1"
X_train_bro
# + colab={"base_uri": "https://localhost:8080/"} id="EDBFbBpB5Zh2" outputId="becc41c5-24d5-4759-e444-572026df0ba5"
from keras.models import Sequential
from keras.layers import Dense, Dropout
from tensorflow.keras.optimizers import RMSprop, Adam
model = Sequential()
model.add(Dense(128, activation="sigmoid"))
model.add(Dropout(0.2))
model.add(Dense(64, activation="sigmoid"))
model.add(Dropout(0.2))
model.add(Dense(32, activation="sigmoid"))
model.add(Dense(1, activation="sigmoid"))
my_optimizer = Adam(learning_rate=0.001)
model.compile(loss="mean_squared_error", optimizer= my_optimizer ,metrics=["mean_absolute_error", "mean_squared_error"])
model.fit(X_train_bro, y_train_bro, epochs=100, batch_size=60)
# + id="ZRXU8k6I5qRx"
y_pred_bro = model.predict(X_test_bro)
# + id="HfluJ0mJ5-9D"
y_test_np_bro = np.array(y_test_bro)
# + colab={"base_uri": "https://localhost:8080/"} id="5jVDqu276GAi" outputId="a5548558-0afe-435e-d022-1cd288465efe"
x_axis = np.array(range(len(y_test_np_bro)))
x_axis.size
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="Q4-kcfNS6Kq6" outputId="19ad8a1d-03eb-49ab-e8ca-0197150d5bb5"
plt.scatter(x_axis, y_test_bro, label="real")
plt.scatter(x_axis, y_pred_bro[:,0], label="estimado")
plt.title("Indicador de precio MinMaxScaled", fontsize=10, color="SteelBlue", fontweight="bold")
plt.ylabel("Indicador de precio MinMaxScaled estimado y real", fontsize=7, color="SteelBlue", fontweight="bold")
plt.legend()
# + colab={"base_uri": "https://localhost:8080/", "height": 783} id="02o_r23b6SSP" outputId="66f7ae53-cb27-426b-c433-1c902488404c"
price_test_bro = y_test_bro * (prices_max_bro - prices_min_bro) + prices_min_bro
price_pred_bro = y_pred_bro * (prices_max_bro - prices_min_bro) + prices_min_bro
print(type(price_test_bro))
print(type(price_pred_bro))
price_test_np_bro = np.array(price_test_bro)
print(type(price_test_np_bro))
print(price_test_np_bro)
print(np.array(price_pred_bro))
error_bro = abs(price_pred_bro[:,0] - price_test_bro)
x_axis_bro = np.array(range(len(price_pred_bro)))
plt.scatter(x_axis_bro, error_bro)
plt.title("abs. Error Precio estimado - Precio real sobre x_test",fontsize=10,color="SteelBlue",fontweight="bold")
plt.ylabel("Error en valor absoluto de precio estimado y precio real en $", color="SteelBlue", fontweight="bold")
error_bro
# + colab={"base_uri": "https://localhost:8080/"} id="gdTaG30o9BlC" outputId="14e68eaf-0ae2-41d4-fbb8-158a0d03f297"
y_test_bro
# + colab={"base_uri": "https://localhost:8080/"} id="HwwBCdgO9EKD" outputId="ac025491-a3bb-4baa-e4f6-af746099e3f8"
y_pred_bro
# + colab={"base_uri": "https://localhost:8080/"} id="gO3IbZeqCpXR" outputId="793a8001-166d-44b4-bfd9-8993cd730d28"
error_bro.describe()
# + [markdown] id="qBZ2NAeb6aB0"
# ## Queens
# + colab={"base_uri": "https://localhost:8080/", "height": 250} id="dN0WHCam6eK4" outputId="7aeee455-5037-4a83-bb76-bd4cb16cd6f0"
df_queens = df_cleaned_3.loc[df_cleaned_3.neighbourhood_group == "Queens", :]
corr_queens = df_queens.corr(method="pearson")
corr_queens.style.background_gradient(cmap='coolwarm')
# + [markdown] id="Xv63WEn561Qp"
# #### We can see in the correlation matrix above the categories that has more influence in price values are: reviews_per_month, calculated_host_listings_count and availability_365
# + colab={"base_uri": "https://localhost:8080/"} id="qtdjYOrr69Uf" outputId="aeb61d19-a632-4490-e605-61b37fc6ccbf"
from sklearn.model_selection import train_test_split
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
features_que = df_queens.loc[:,["reviews_per_month", "calculated_host_listings_count", "availability_365"]]
target_que = df_queens.loc[:,"price"]
scaler_que = MinMaxScaler(feature_range=(0,1))
prices_que = np.array(df_queens.loc[:,"price"]).reshape(-1,1)
reviews_que = np.array(df_queens.loc[:,"reviews_per_month"]).reshape(-1,1)
calculated_que = np.array(df_queens.loc[:,"calculated_host_listings_count"]).reshape(-1,1)
availability_que = np.array(df_queens.loc[:,"availability_365"]).reshape(-1,1)
prices_min_que = prices_que.min()
prices_max_que = prices_que.max()
prices_n_que = scaler_que.fit_transform(prices_que)
reviews_n_que = scaler_que.fit_transform(reviews_que)
calculated_n_que = scaler_que.fit_transform(calculated_que)
availability_n_que = scaler_que.fit_transform(availability_que)
new_len_que = len(prices_que)
features_matrix_que = np.zeros((new_len_que, 4))
features_matrix_que[:,0] = reviews_n_que[:,0]
features_matrix_que[:,1] = calculated_n_que[:,0]
features_matrix_que[:,2] = availability_n_que[:,0]
features_matrix_que[:,3] = prices_n_que[:,0]
features_df_que = pd.DataFrame(features_matrix_que)
features_df_que.columns = ["reviews_per_month", "calculated_host_listings_count", "availability_365", "price"]
features_n_que = features_df_que.loc[:,["reviews_per_month", "calculated_host_listings_count", "availability_365"]]
target_n_que = features_df_que.loc[:,"price"]
print(features_df_que)
# + id="su0qxWMA7XeZ"
X_train_que, X_test_que, y_train_que, y_test_que = train_test_split(features_n_que, target_n_que, test_size=0.25, random_state=73)
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="tbmkkkOHMIqc" outputId="d25bac08-2835-4bcd-928e-a1e663d60f0d"
X_train_que
# + colab={"base_uri": "https://localhost:8080/"} id="lTH3-RlQ7b0J" outputId="ff2f169c-f39a-4fe4-e4da-4b6100a327f1"
from keras.models import Sequential
from keras.layers import Dense, Dropout
from tensorflow.keras.optimizers import RMSprop, Adam
model = Sequential()
model.add(Dense(128, activation="tanh"))
model.add(Dropout(0.2))
model.add(Dense(64, activation="tanh"))
model.add(Dropout(0.2))
model.add(Dense(32, activation="tanh"))
model.add(Dense(1, activation="tanh"))
my_optimizer = Adam(learning_rate=0.001)
model.compile(loss="mean_squared_error", optimizer= my_optimizer ,metrics=["mean_absolute_error", "mean_squared_error"])
model.fit(X_train_que, y_train_que, epochs=100, batch_size=50)
# + id="gIKi0gp_7kj3"
y_pred_que = model.predict(X_test_que)
# + id="zhgbs7f97pbj"
y_test_np_que = np.array(y_test_que)
# + colab={"base_uri": "https://localhost:8080/"} id="h3L5o85x7tib" outputId="0ced415f-c33a-489e-b65b-3f18a4d4bb4b"
x_axis_que = np.array(range(len(y_test_np_que)))
x_axis_que.size
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="m6KJjG237xV9" outputId="b71fde9a-9f96-42cd-df34-ee71ef97121e"
plt.scatter(x_axis_que, y_test_que, label="real")
plt.scatter(x_axis_que, y_pred_que[:,0], label="estimado")
plt.title("Indicador de precio MinMaxScaled", fontsize=10, color="SteelBlue", fontweight="bold")
plt.ylabel("Indicador de precio MinMaxScaled estimado y real", fontsize=7, color="SteelBlue", fontweight="bold")
plt.legend()
# + colab={"base_uri": "https://localhost:8080/", "height": 766} id="UVnN3jCF74P5" outputId="671a1b26-4cb2-4a5b-c0fa-3d6b912853f6"
price_test_que = y_test_que * (prices_max_que - prices_min_que) + prices_min_que
price_pred_que = y_pred_que * (prices_max_que - prices_min_que) + prices_min_que
print(type(price_test_que))
print(type(price_pred_que))
price_test_np_que = np.array(price_test_que)
print(type(price_test_np_que))
print(price_test_np_que)
print(np.array(price_pred_que))
error_que = abs(price_pred_que[:,0] - price_test_que)
x_axis_que = np.array(range(len(price_pred_que)))
plt.scatter(x_axis_que, error_que)
plt.title("abs. Error Precio estimado - Precio real sobre x_test",fontsize=10,color="SteelBlue",fontweight="bold")
plt.ylabel("Error en valor absoluto de precio estimado y precio real en $", color="SteelBlue", fontweight="bold")
error_que
# + colab={"base_uri": "https://localhost:8080/"} id="GHBf13Kv8JuA" outputId="c058ef23-8540-46b5-9da0-620f39214d6c"
price_test_que
# + colab={"base_uri": "https://localhost:8080/"} id="B_0eYZ3R8QE3" outputId="61772679-5d30-4dcb-dea6-d0cf374c0bd5"
price_pred_que
# + colab={"base_uri": "https://localhost:8080/"} id="dpOed6SJDUTe" outputId="93a08dd6-9d52-4984-ba8b-5cc815dcc2f3"
error_que.describe()
| Prueba_5_Raul_Vergel_Romero_Opcion_A.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotly Fun
# Jupyter Notebooks look quite interesting, particularly the charting from [Plotly]. Let's give the it a go! 🚀
#
# First up: let's import all the libraries we need for this notebook now (this was adapted from the [offline] docs:
#
# [Plotly]: https://plot.ly/python/v3/ipython-notebook-tutorial/
# [offline]: https://plot.ly/python/v3/offline/
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
import pandas
import plotly.figure_factory as ff
import plotly.graph_objs as go
# Now for any other prior setup:
init_notebook_mode(connected=True)
# Now let's grab a simple data set and plot it:
data = pandas.read_csv("https://raw.githubusercontent.com/plotly/datasets/master/school_earnings.csv")
table = ff.create_table(data)
iplot(table, filename="jupyter-table1")
# We can look at this data however we want! Let's try some simple inspection of the data:
data.School[0]
# How about an interactive chart, though?
data = [go.Bar(x=data.School, y=data.Gap)]
iplot(data, filename='jupyter-basic_bar')
# Not too much effort, and this is looking really interesting!
| plotly-tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="NA5FvNG-KdqJ" colab_type="text"
# ## finance.yahoo.co.jp
# + id="Cfl503QHH1Fm" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1600280229896, "user_tz": -540, "elapsed": 518, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNeNR8zYe2bcZus4tq_IiWYynmssgG5VWC92Lo=s64", "userId": "14098338404840441752"}}
import pandas as pd
# + id="G7eUHp3tJtc9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 131} executionInfo={"status": "ok", "timestamp": 1600280585632, "user_tz": -540, "elapsed": 1613, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNeNR8zYe2bcZus4tq_IiWYynmssgG5VWC92Lo=s64", "userId": "14098338404840441752"}} outputId="77f3d7e5-972c-4b0c-ccf3-441105997482"
url = 'https://finance.yahoo.co.jp/'
table = pd.read_html(url)
print("No. of Table: ", len(table))
df = table[0]
df
# + id="-UCotdxNHi_B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} executionInfo={"status": "ok", "timestamp": 1600280466313, "user_tz": -540, "elapsed": 1941, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNeNR8zYe2bcZus4tq_IiWYynmssgG5VWC92Lo=s64", "userId": "14098338404840441752"}} outputId="3a614ba6-4438-4a19-ee14-177a3dcdf62a"
url = 'https://info.finance.yahoo.co.jp/ranking/?kd=4'
table = pd.read_html(url)
print("No. of Table: ", len(table))
df = table[0]
df.head(5)
# + id="JxNWbnSTJKx4" colab_type="code" colab={}
| notebooks/web_scraping_yahoo_fin_ranking_table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
# %matplotlib inline
cell_file = '../cells/poolosyn.cell.nml'
cell_id = 'poolosyncell'
from pyneuroml.analysis import generate_current_vs_frequency_curve
curve = generate_current_vs_frequency_curve(cell_file,
cell_id,
custom_amps_nA = [-0.4,-0.35,-0.3,-0.25,-0.2,-0.15,-0.1,-0.05,0,0.3],
analysis_duration = 500,
pre_zero_pulse = 50,
post_zero_pulse = 50,
analysis_delay = 0,
dt = 0.025,
simulator = 'jNeuroML_NEURON',
plot_voltage_traces = True,
plot_if = False,
plot_iv = False,
temperature = '34degC',
title_above_plot = True)
# +
# Longer duration, more points
curve = generate_current_vs_frequency_curve(cell_file,
cell_id,
start_amp_nA = -0.1,
end_amp_nA = 0.4,
step_nA = 0.02,
analysis_duration = 1000, ######### 2000
pre_zero_pulse = 0,
post_zero_pulse = 0,
analysis_delay = 100,
dt = 0.025,
simulator = 'jNeuroML_NEURON',
plot_voltage_traces = False,
plot_if = True,
plot_iv = True,
temperature = '34degC',
title_above_plot = True)
# -
from pyneuroml import pynml
pynml.run_jneuroml("", cell_file, '-png')
from IPython.display import Image
Image(filename=cell_file.replace('.nml','.png'),width=500)
| NeuroML2/notebooks/Test_poolosyn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Challenge Notebook
# ## Problem: Given a knapsack with a total weight capacity and a list of items with weight w(i) and value v(i), determine which items to select to maximize total value.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Can we replace the items once they are placed in the knapsack?
# * No, this is the 0/1 knapsack problem
# * Can we split an item?
# * No
# * Can we get an input item with weight of 0 or value of 0?
# * No
# * Can we assume the inputs are valid?
# * No
# * Are the inputs in sorted order by val/weight?
# * Yes, if not we'd need to sort them first
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * items or total weight is None -> Exception
# * items or total weight is 0 -> 0
# * General case
#
# <pre>
# total_weight = 8
# items
# v | w
# 0 | 0
# a 2 | 2
# b 4 | 2
# c 6 | 4
# d 9 | 5
#
# max value = 13
# items
# v | w
# b 4 | 2
# d 9 | 5
# </pre>
# ## Algorithm
#
# Refer to the [Solution Notebook](). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
class Item(object):
def __init__(self, label, value, weight):
self.label = label
self.value = value
self.weight = weight
def __repr__(self):
return self.label + ' v:' + str(self.value) + ' w:' + str(self.weight)
class Knapsack(object):
def fill_knapsack(self, input_items, total_weight):
# TODO: Implement me
pass
# ## Unit Test
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_knapsack.py
from nose.tools import assert_equal, assert_raises
class TestKnapsack(object):
def test_knapsack_bottom_up(self):
knapsack = Knapsack()
assert_raises(TypeError, knapsack.fill_knapsack, None, None)
assert_equal(knapsack.fill_knapsack(0, 0), 0)
items = []
items.append(Item(label='a', value=2, weight=2))
items.append(Item(label='b', value=4, weight=2))
items.append(Item(label='c', value=6, weight=4))
items.append(Item(label='d', value=9, weight=5))
total_weight = 8
expected_value = 13
results = knapsack.fill_knapsack(items, total_weight)
assert_equal(results[0].label, 'd')
assert_equal(results[1].label, 'b')
total_value = 0
for item in results:
total_value += item.value
assert_equal(total_value, expected_value)
print('Success: test_knapsack_bottom_up')
def test_knapsack_top_down(self):
knapsack = KnapsackTopDown()
assert_raises(TypeError, knapsack.fill_knapsack, None, None)
assert_equal(knapsack.fill_knapsack(0, 0), 0)
items = []
items.append(Item(label='a', value=2, weight=2))
items.append(Item(label='b', value=4, weight=2))
items.append(Item(label='c', value=6, weight=4))
items.append(Item(label='d', value=9, weight=5))
total_weight = 8
expected_value = 13
assert_equal(knapsack.fill_knapsack(items, total_weight), expected_value)
print('Success: test_knapsack_top_down')
def main():
test = TestKnapsack()
test.test_knapsack_bottom_up()
test.test_knapsack_top_down()
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook]() for a discussion on algorithms and code solutions.
| recursion_dynamic/knapsack_01/knapsack_challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
# +
file_name="Resources/clean_2019.csv"
clean_2019=pd.read_csv(file_name)
# clean_2019.drop("Unnamed:0")
clean_2019.tail()
# They have no impact on the total score reported for each country, but they do explain why some countries rank higher than others.
# -
clean_2019.drop(["Unnamed: 0"], axis=1)
clean_2019.describe()
# +
# Assign the data to X and y
X = clean_2019[["gdp_per_capita", "social_support","healthy_life_expectancy","freedom_choice","generosity","perceptions_of_corruption"]]
y = clean_2019["happiness_score"].values.reshape(-1, 1)
print(X.shape, y.shape)
X
# -
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
model = LinearRegression()
# +
model.fit(X_train, y_train)
training_score = model.score(X_train, y_train)
testing_score = model.score(X_test, y_test)
### END SOLUTION
print(f"Training Score: {training_score}")
print(f"Testing Score: {testing_score}")
# -
### BEGIN SOLUTION
plt.scatter(model.predict(X_train), model.predict(X_train) - y_train, c="blue", label="Training Data")
plt.scatter(model.predict(X_test), model.predict(X_test) - y_test, c="orange", label="Testing Data")
plt.legend()
plt.hlines(y=0, xmin=y.min(), xmax=y.max())
plt.title("Residual Plot")
### END SOLUTION
# from mpl_toolkits.mplot3d import Axes3D
# fig = plt.figure(1, figsize=(5, 5))
# axes = Axes3D(fig, elev=20, azim=45)
# axes.scatter(X[:,0], X[:,1], X[:,2], X[:,3], X[:,4], X[:,5], c=y, cmap=plt.cm.get_cmap("Spectral"))
# plt.show()
print('Weight coefficients: ', model.coef_)
print('y-axis intercept: ', model.intercept_)
# +
# LASSO model
# Note: Use an alpha of .01 when creating the model for this activity
from sklearn.linear_model import Lasso
from sklearn.metrics import mean_squared_error
### BEGIN SOLUTION
lasso = Lasso(alpha=.01).fit(X_train, y_train)
predictions = lasso.predict(X_test)
MSE = mean_squared_error(y_test, predictions)
r2 = lasso.score(X_test, y_test)
### END SOLUTION
print(f"MSE: {MSE}, R2: {r2}")
# +
# Ridge model
# Note: Use an alpha of .01 when creating the model for this activity
from sklearn.linear_model import Ridge
### BEGIN SOLUTION
ridge = Ridge(alpha=.01).fit(X_train, y_train)
predictions = ridge.predict(X_test)
MSE = mean_squared_error(y_test, predictions)
r2 = ridge.score(X_test, y_test)
### END SOLUTION
print(f"MSE: {MSE}, R2: {r2}")
# +
# ElasticNet model
# Note: Use an alpha of .01 when creating the model for this activity
from sklearn.linear_model import ElasticNet
### BEGIN SOLUTION
elasticnet = ElasticNet(alpha=.01).fit(X_train, y_train)
predictions = elasticnet.predict(X_test)
MSE = mean_squared_error(y_test, predictions)
r2 = elasticnet.score(X_test, y_test)
### END SOLUTION
print(f"MSE: {MSE}, R2: {r2}")
# -
| .ipynb_checkpoints/Country_Happiness_Predictor-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
import warnings
import imageio
from skimage.transform import resize
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
warnings.filterwarnings("ignore")
# -
label_df = pd.read_csv('label.csv')
len(label_df)
# +
fake_train_dir = '//nas5.pmi.rwth-aachen.de/mri-scratch/DeepLearning/Fake_Stanford_256_final/'
fake_img_list = os.listdir(fake_train_dir)
len(fake_img_list)
#### super nice sort algorithm! ############
import re
digits = re.compile(r'(\d+)')
def tokenize(filename):
return tuple(int(token) if match else token
for token, match in
((fragment, digits.search(fragment))
for fragment in digits.split(filename)))
# Now you can sort your PDF file names like so:
fake_img_list.sort(key=tokenize)
# -
label_df.insert(loc=0, column='Fake Image Index', value=fake_img_list)
Data_Dir = '/media/tianyu.han/mri-scratch/DeepLearning/Fake_Stanford_256_2/'
label_df['Image Path'] = label_df['Fake Image Index'].map(lambda x: Data_Dir + x)
nih_df = pd.read_csv('fake_nih_78468.csv')
nih_train_df, nih_validtest_df = [x for _, x in nih_df.groupby(nih_df['fold'] != 'train')]
nih_train_df = nih_train_df.reset_index(drop=True)
len(nih_train_df)
len(nih_validtest_df)
nih_train_df.head()
nih_validtest_df.head()
label_df = label_df.drop(columns=['Fake Image Index', 'Path', 'No Finding', 'Enlarged Cardiomediastinum',
'Lung Lesion', 'Pleural Other', 'Fracture', 'Image Index',
'Support Devices'])
label_df.insert(loc=0, column='Image Index', value=fake_img_list)
label_df.insert(loc=8, column='fold', value='train')
label_df.rename({'Pleural Effusion': 'Effusion'}, axis=1, inplace=True)
label_df
nih_column = nih_df.columns.tolist()
label_df = label_df[nih_column]
label_df.head()
validtest_img_dir = '/media/tianyu.han/mri-scratch/DeepLearning/NIH_Chest/images/'
train_img_dir = '/media/tianyu.han/mri-scratch/DeepLearning/Fake_78468/'
nih_train_df = nih_train_df.drop(columns='Image Path')
nih_validtest_df = nih_validtest_df.drop(columns='Image Path')
nih_train_df['Image Path'] = nih_train_df['Image Index'].map(lambda x: train_img_dir + x)
nih_validtest_df['Image Path'] = nih_validtest_df['Image Index'].map(lambda x: validtest_img_dir + x)
nih_train_df.head()
nih_validtest_df.head()
frames = [label_df, nih_train_df, nih_validtest_df]
nih_stanford_df = pd.concat(frames)
nih_stanford_df = nih_stanford_df.reset_index(drop=True)
len(nih_stanford_df)
nih_stanford_df.tail()
saveDir = '/media/tianyu.han/mri-scratch/DeepLearning/CheXpert_Dataset/DL_DENSENET/Densenet_Stanford_lr/'
file = 'nih_stanford.csv'
nih_stanford_df.to_csv(os.path.join(saveDir, file), index=False)
| Preprocess/Stanford_CSV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FloPy
#
# ### Lake Example
#
# First set the path and import the required packages. The flopy path doesn't have to be set if you install flopy from a binary installer. If you want to run this notebook, you have to set the path to your own flopy path.
# +
import os
import sys
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
workspace = os.path.join('data')
# make sure workspace directory exists
if not os.path.exists(workspace):
os.makedirs(workspace)
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
# -
# We are creating a square model with a specified head equal to `h1` along all boundaries. The head at the cell in the center in the top layer is fixed to `h2`. First, set the name of the model and the parameters of the model: the number of layers `Nlay`, the number of rows and columns `N`, lengths of the sides of the model `L`, aquifer thickness `H`, hydraulic conductivity `k`
name = 'lake_example'
h1 = 100
h2 = 90
Nlay = 10
N = 101
L = 400.0
H = 50.0
k = 1.0
# Create a MODFLOW model and store it (in this case in the variable `ml`, but you can call it whatever you want). The modelname will be the name given to all MODFLOW files (input and output). The exe_name should be the full path to your MODFLOW executable. The version is either 'mf2k' for MODFLOW2000 or 'mf2005'for MODFLOW2005.
ml = flopy.modflow.Modflow(modelname=name, exe_name='mf2005', version='mf2005', model_ws=workspace)
# Define the discretization of the model. All layers are given equal thickness. The `bot` array is build from the `Hlay` values to indicate top and bottom of each layer, and `delrow` and `delcol` are computed from model size `L` and number of cells `N`. Once these are all computed, the Discretization file is built.
bot = np.linspace(-H/Nlay,-H,Nlay)
delrow = delcol = L/(N-1)
dis = flopy.modflow.ModflowDis(ml,nlay=Nlay,nrow=N,ncol=N,delr=delrow,delc=delcol,top=0.0,botm=bot,laycbd=0)
# Next we specify the boundary conditions and starting heads with the Basic package. The `ibound` array will be `1` in all cells in all layers, except for along the boundary and in the cell at the center in the top layer where it is set to `-1` to indicate fixed heads. The starting heads are used to define the heads in the fixed head cells (this is a steady simulation, so none of the other starting values matter). So we set the starting heads to `h1` everywhere, except for the head at the center of the model in the top layer.
Nhalf = int((N-1) / 2)
ibound = np.ones((Nlay, N, N), dtype=np.int)
ibound[:,0,:] = -1; ibound[:,-1,:] = -1; ibound[:,:,0] = -1; ibound[:,:,-1] = -1
ibound[0,Nhalf,Nhalf] = -1
start = h1 * np.ones((N,N))
start[Nhalf,Nhalf] = h2
bas = flopy.modflow.ModflowBas(ml,ibound=ibound,strt=start)
# The aquifer properties (really only the hydraulic conductivity) are defined with the LPF package.
lpf = flopy.modflow.ModflowLpf(ml, hk=k)
# Finally, we need to specify the solver we want to use (PCG with default values), and the output control (using the default values). Then we are ready to write all MODFLOW input files and run MODFLOW.
pcg = flopy.modflow.ModflowPcg(ml)
oc = flopy.modflow.ModflowOc(ml)
ml.write_input()
ml.run_model()
# Once the model has terminated normally, we can read the heads file. First, a link to the heads file is created with `HeadFile`. The link can then be accessed with the `get_data` function, by specifying, in this case, the step number and period number for which we want to retrieve data. A three-dimensional array is returned of size `nlay, nrow, ncol`. Matplotlib contouring functions are used to make contours of the layers or a cross-section.
hds = flopy.utils.HeadFile(os.path.join(workspace, name+'.hds'))
h = hds.get_data(kstpkper=(0, 0))
x = y = np.linspace(0, L, N)
c = plt.contour(x, y, h[0], np.arange(90,100.1,0.2))
plt.clabel(c, fmt='%2.1f')
plt.axis('scaled');
x = y = np.linspace(0, L, N)
c = plt.contour(x, y, h[-1], np.arange(90,100.1,0.2))
plt.clabel(c, fmt='%1.1f')
plt.axis('scaled');
z = np.linspace(-H/Nlay/2, -H+H/Nlay/2, Nlay)
c = plt.contour(x, z, h[:,50,:], np.arange(90,100.1,.2))
plt.axis('scaled');
| examples/Notebooks/flopy3_lake_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="GDZTwH3tS9BM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="13802475-a2cc-47f5-fdda-788c5a5243a0" executionInfo={"status": "ok", "timestamp": 1583270933814, "user_tz": -60, "elapsed": 7675, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
# !pip install --upgrade tables
# + id="rhvTDrujTMEi" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# + id="2m_fwWGTTej7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="affe6c47-3e7b-4fe9-e5cf-8c6e6e24b4c1" executionInfo={"status": "ok", "timestamp": 1583270989937, "user_tz": -60, "elapsed": 3325, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
# ls
# + id="bHZnvOhyTgJj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dc00d3a7-7311-428b-ee5f-d61631dac57f" executionInfo={"status": "ok", "timestamp": 1583271013992, "user_tz": -60, "elapsed": 3065, "user": {"displayName": "Micha\u01<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
# ls
# + id="iGZEcu6nTmHP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="13e4c055-529f-48ad-b65e-2932a8ba6049" executionInfo={"status": "ok", "timestamp": 1583271050147, "user_tz": -60, "elapsed": 755, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
# cd '/content/drive/My Drive/Colab Notebooks/matrix/matrixtwo/dw_matrix_car'
# + id="q60NcJFwTozC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="261a0b80-0386-455b-df6d-da59e6d2e41a" executionInfo={"status": "ok", "timestamp": 1583271055341, "user_tz": -60, "elapsed": 2840, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
# ls
# + id="H1swm9dRTwP5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d173c1bf-333b-4af1-9ce2-7b01eaafa790" executionInfo={"status": "ok", "timestamp": 1583271094598, "user_tz": -60, "elapsed": 4314, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="1ErXT9-1T5gk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4104a61e-a461-4150-894a-952204b7fd61" executionInfo={"status": "ok", "timestamp": 1583271116141, "user_tz": -60, "elapsed": 873, "user": {"displayName": "Micha\u01<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
df.columns.values
# + id="B_hZAY6-T_jo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="2ed0336c-5ff4-4084-cf73-c0174df55f18" executionInfo={"status": "ok", "timestamp": 1583271186709, "user_tz": -60, "elapsed": 1127, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
df['price_value'].hist(bins=100)
# + id="9PpNDR1QUQxN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="ab0d9e5d-5f94-43f4-aa94-cce1530ea9c4" executionInfo={"status": "ok", "timestamp": 1583271259711, "user_tz": -60, "elapsed": 907, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
df['price_value'].describe()
# + id="LwVx21qpUapr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="3a9ca5d7-c8ee-4a68-bb70-7a7093a27c10" executionInfo={"status": "ok", "timestamp": 1583271351388, "user_tz": -60, "elapsed": 585, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
df['param_marka-pojazdu'].unique()
# + id="myUfhuXnU5Gv" colab_type="code" colab={}
def group_barplot(feat_groupby, feat_agg='price_value', agg_funcs= [np.mean, np.median, np.size], feat_sort='mean', top=50, subplots=True):
return(
df
.groupby(feat_groupby)[feat_agg]
.agg(agg_funcs)
.sort_values(by=feat_sort, ascending = False)
.head(top)
).plot(kind='bar', figsize = (30,8), subplots = subplots)
# + id="XfFoFUO6VQ6j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 580} outputId="81be0b18-50e3-4b95-91a0-af5d245b02f2" executionInfo={"status": "ok", "timestamp": 1583272667606, "user_tz": -60, "elapsed": 3003, "user": {"displayName": "Micha\u0142 Klimczak", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
group_barplot('param_marka-pojazdu');
# + id="HzsHZWXjZV2v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 599} outputId="0b063286-06d0-4905-84de-db9d5a71a1dc" executionInfo={"status": "ok", "timestamp": 1583272777751, "user_tz": -60, "elapsed": 2417, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
group_barplot('param_kraj-pochodzenia');
# + id="_ef0QujlaLch" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 553} outputId="fe6de33a-8a83-4e97-e4dc-a661dff11e34" executionInfo={"status": "ok", "timestamp": 1583272895121, "user_tz": -60, "elapsed": 1990, "user": {"displayName": "Micha\u0142 Klimczak", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjOwMFoju4G1lg6oWLZRVB_jrFchbit4ekIqIEEsg=s64", "userId": "14327755442440161367"}}
group_barplot('param_kolor', feat_sort='mean');
# + id="D1CrE2Auasnl" colab_type="code" colab={}
| m2_day2_visual.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Griff-Kaiga/facial-detection-model/blob/main/Facial_detection_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="eVs2tZQ_sMWe"
# Moraan, mostly we run models using jupyter notebook or colab(like this one). To run this, click on connect, clickon the runtime tab, seleect restart and run all option and you good to go.
# + id="uk3b_0fVZ4vr"
# importing the neccesary libraries
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from keras.utils import to_categorical
from tensorflow.keras.models import Sequential
from keras.models import Sequential
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from keras.layers import Flatten
from keras.layers import Dense
from keras.layers import Activation, Dense
from keras.layers import Dropout
# + colab={"base_uri": "https://localhost:8080/"} id="iwY1hrZi_k9L" outputId="bbf6a8a2-4d43-4862-9ef8-8fb709a394aa"
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + id="I4XF5fp2KP2B"
import os
data_dir = "/content/drive/MyDrive/humans"
data_paths = [os.path.join(data_dir, f) for f in os.listdir(data_dir)]
data_paths = [i for i in data_paths if os.path.isfile(i)]
# + id="yjjZErK8m97y"
# loading the images off disk and defining some parameters for it
batch_size = 32
img_height = 180
img_width = 180
# + colab={"base_uri": "https://localhost:8080/"} id="NtDh5DnanDkF" outputId="b6992a24-f4ff-4982-b256-14960e2ae974"
# creating a training and validation test set
train_ds = tf.keras.preprocessing.image_dataset_from_directory(
data_dir,
validation_split= 0.2,
subset= 'training',
seed= 123,
image_size = (img_height, img_width),
batch_size = batch_size,
labels = 'inferred'
)
print(train_ds.class_names)
# + colab={"base_uri": "https://localhost:8080/"} id="vWSyDxmLtRJF" outputId="1d46e23c-5893-4a98-ae1c-8db835995fea"
# validation dataset
valid_ds = tf.keras.preprocessing.image_dataset_from_directory(
data_dir,
validation_split = 0.2,
subset = 'validation',
seed= 123,
image_size = (img_height,img_width),
batch_size = batch_size
)
# + id="BZx1JdsuvEtC"
# configuring the dataset for performace
AUTOTUNE = tf.data.AUTOTUNE
train_ds = train_ds.cache().shuffle(1000).prefetch(buffer_size=AUTOTUNE)
valid_ds = valid_ds.cache().prefetch(buffer_size=AUTOTUNE)
# + id="y-qumBDdvz9Z"
# standardizing my data
normalization_layer = layers.experimental.preprocessing.Rescaling(1./255)
# + colab={"background_save": true} id="e7aS45PlwFul" outputId="06afae0e-2a7b-44d7-dfb8-0d5defcee8c0"
normalized_ds = train_ds.map(lambda x, y: (normalization_layer(x), y))
image_batch, labels_batch = next(iter(normalized_ds))
first_image = image_batch[0]
# Notice the pixels values are now in `[0,1]`.
print(np.min(first_image), np.max(first_image))
# + id="1F3jfM0qz1aY"
# model creation
num_classes = 3
model = Sequential([
layers.experimental.preprocessing.Rescaling(1./255, input_shape=(img_height, img_width, 3)),
layers.Conv2D(16, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(32, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(64, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dense(num_classes, activation = 'softmax')
])
# + id="5mY_L0SP0FrL"
# compile it
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="u1ZXdAul0P1R" outputId="69cd46a9-6366-4477-a4c8-146ef61314ab"
# get the summary
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="Y8YQ5T_c0S47" outputId="5b69ecb0-1162-4267-eeeb-f82dfe4f5edc"
# fitting in the training data
epochs=10
history = model.fit(
train_ds,
validation_data=valid_ds,
epochs=epochs
)
# + id="AA5-N2X-7p8E" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="d37b2521-c110-4534-8092-aea48ffc75d8"
# performing data augmentation to allow for generalization
data_augmentation = keras.Sequential(
[
layers.experimental.preprocessing.RandomFlip("horizontal",
input_shape=(img_height,
img_width,
3)),
layers.experimental.preprocessing.RandomRotation(0.1),
layers.experimental.preprocessing.RandomZoom(0.1),
]
)
# + colab={"base_uri": "https://localhost:8080/", "height": 575} id="s86NyfD37tLj" outputId="d2326fc7-e4f1-4e9d-ab81-e79043592326"
# visualizing a few augmentated examples
index = 1
plt.figure(figsize=(10, 10))
for images, _ in train_ds.take(index):
for i in range(9):
augmented_images = data_augmentation(images)
ax = plt.subplot(3, 3, i + 1)
plt.imshow(augmented_images[0].numpy().astype("uint8"))
plt.axis("off")
# + id="GBAupUsO74Pa"
# introducing dropout layer to prevent overfitting
model = Sequential([
data_augmentation,
layers.experimental.preprocessing.Rescaling(1./255),
layers.Conv2D(16, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(32, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(64, 3, padding='same', activation='relu'),
layers.MaxPooling2D(),
layers.Dropout(0.2),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dense(num_classes)
])
# + id="5H9WlhdZ8EZy"
# compiling the model
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="0Rlx7HR_8SpG" outputId="b248534d-46d9-414b-e5c5-e48579eadf25"
# get the summary
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="zcyQAeKI8VUf" outputId="25c02f01-4621-4b1a-d29f-e72d49fc0a70"
# training the model
epochs = 10
history = model.fit(
train_ds,
validation_data=valid_ds,
epochs=epochs
)
# + colab={"base_uri": "https://localhost:8080/"} id="ZxdxXLAMzMzA" outputId="65f97404-07e5-4599-fc46-5b738995704b"
# evaluate the model using the test dataset
model.evaluate(valid_ds)[1]
# + [markdown] id="lcWY0b0Gtwo1"
# This is the point where you test your model using a photo. run the first cell below and click on the file to upload, assuming you have the photo on your machine.
#
# + id="oeIoklUOz1ON"
# test the model with an example
from google.colab import files
uploaded_file = files.upload()
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="lEE4F8Q08Mqk" outputId="57480018-ee69-47ea-8a47-c4102219ef5a"
# preview the image
new_image = plt.imread('632316-gp.jpg')
test_image = plt.imshow(new_image)
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="cY9rcp1V7XIu" outputId="85b44ae2-fcb7-445c-d34d-cc0085e6f5df"
# resize the image
from skimage.transform import resize
resized_image = resize(new_image, (180,180,3))
# show the resized image
image = plt.imshow(resized_image)
# + colab={"base_uri": "https://localhost:8080/"} id="jubIJAS5-Me3" outputId="c70e9e59-d496-4adb-edc4-39b06849fd2e"
resized_image.shape
# models prediction
predictions =model.predict(np.array([resized_image]))
# showing the predictions
predictions
# + colab={"base_uri": "https://localhost:8080/"} id="Xx8hKFcDE2pL" outputId="89d70326-9117-4ba2-8972-cfbf5f4bb418"
# sort the predictions from the least to greatest
list_index = [0,1,2]
x = predictions
for i in range(2):
for j in range(2):
if x[0][list_index[i]] > x[0][list_index[j]]:
temp = list_index[i]
list_index[i] = list_index[j]
list_index[j] = temp
# show sorted labels in order
print(list_index)
# + id="-S4ZR3f5NyWG"
| Facial_detection_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import plotly
import plotly.plotly as py
plotly.tools.set_credentials_file(username='falrashidi', api_key='XaO64TRYU0N3Sdup8Z3H')
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
# ### Plotly & Cufflinks
# At this point you will need to isntall `cufflinks`. `Cufflinks` binds `Plotly` directly to pandas dataframes.
# ```python
# # ! pip install cufflinks --upgrade
# ```
# +
import cufflinks as cf
print(cf.__version__)
import pandas as pd
import numpy as np
import gzip
# Configure cufflings
cf.set_config_file(offline=False, world_readable=True, theme='pearl')
# -
# ## Loading the Data
# The below functions are provided directly from the [Amazon Review Data link](http://jmcauley.ucsd.edu/data/amazon/) by the author and it is used to load the [5-cores](https://en.wikipedia.org/wiki/Degeneracy_(graph_theory)) book reviews as a `panda dataframe`.
def parse(path):
g = gzip.open(path, 'rb')
for l in g:
yield eval(l)
def getDF(path):
i = 0
df = {}
for d in parse(path):
df[i] = d
i += 1
return pd.DataFrame.from_dict(df, orient='index')
df = getDF('/Users/falehalrashidi/Downloads/reviews_Books_5.json.gz')
# I used the below snippet to monitor the memory requirements for the loading.
# %load_ext memory_profiler
# %memit
# Below you can see the fields loaded and a count of the values per field;
df.count()
# A sample of the overal data appears next:
df[0:10]
# ## Column Fields of Interest
# In general, the loaded dataframe, include 7 fields:
# * `reviewerID: A `String` (probably a hashText) that uniquely identifies the user that submitted the review.
# * `asin`: ASIN stands for **Amazon Standard Identification Number**. Almost every product on **Amazon** has its own **ASIN**, a unique code used to identify it. For books, the **ASIN** is the same as the book's **ISBN** number.
# * `reviewerName`: The name of the reviewer.
# * `helpful`: Amazon has implemented an interface that allows customers to vote on whether a particular review
# has been helpful or unhelpful. This is captured by this field, which represents a rating of the review, e.g. if `[2,3] --> 2/3`.
# * `reviewText`: The actual review provided by the reviewer.
# * `overall`: The product's rating attributed by the same reviewer.
# * `summary`: A summary of the review.
# * `unixReviewTime`: Time of the review (unix time).
# * `reviewTime`: Time of the review (raw).
#
# Of these fields, for the purposes of this project we care to keep the `reviewerID`, `asin`, `reviewText`, `overall` and `helpful`. Specifically, we keep `reviewerID` only to merge it with `asin` and create unique identifier (`key`) per review, e.g.:
#
# ```python
# key = reviewerID:"<KEY>" + asin:"000100039X"
# ```
#
# `asin` is obviously necessary to identify the distinct books in the dataset, while the rest are necessary for the analysis (`overall`, `reviewText`) and for evaluation (`helpful`) purposes.
# ## Data Inspection
# +
# Number of reviews:
number_of_reviews=len(df)
my_number_string = '{:0,.0f}'.format(number_of_reviews)
print('Number of Reviews: ' + my_number_string + '.')
# +
# Unique number of items:
unique_books=len(df['asin'].unique())
my_number_string = '{:0,.0f}'.format(unique_books)
print('Number of Books: ' + my_number_string + '.')
# -
# ### Distribution of ratings amongst all reviews
# Distribution of Ratings (too many to plot with plotly)
fig = df['overall'].plot.hist(alpha=0.5, title='Ratings Distribution', figsize=(15,7), grid=True)
fig.set_xlabel("Ratings")
fig.set_ylabel("Number of Review")
df10 = df[['overall','asin']]
df11 = pd.DataFrame(df10.groupby(['asin'])['overall'].mean())
# ### Distribution of Average Book Ratings
len(df11)
df11 = df11.reset_index()
df11.head()
#df11['overall'].iplot(kind='histogram', bins=100, xTitle='Rating (0-5)',yTitle='Number of Books', title='Average Book Ratings')
df11.plot.hist(alpha=0.5,bins=100)
# ### Books per Year
df20 = df[['asin','reviewTime']]
def get_year(reviewTime):
day_month_year_list = reviewTime.split(',')
if(len(day_month_year_list)==2):
return day_month_year_list[1]
else:
return fillna(0)
df20['reviewYear'] = pd.DataFrame(df20['reviewTime'].apply(lambda time: get_year(time)))
df20.head()
books_per_year = pd.DataFrame(df20.groupby(['reviewYear']).size())
books_per_year.columns = ['counts']
books_per_year.iplot(kind='bar', xTitle='Years', yTitle='Number of Reviews', title='Number of Reviews per Year')
df30 = df[['asin','reviewTime', 'overall']]
df30['reviewYear'] = pd.DataFrame(df30['reviewTime'].apply(lambda time: get_year(time)))
df30.head()
books_per_rating_per_year = df30.groupby(['reviewYear','overall']).size().reset_index(name='counts')
books_per_rating_per_year[0:10]
pivot_df = books_per_rating_per_year.pivot(index='reviewYear', columns='overall', values='counts')
pivot_df.iplot(kind='bar', barmode='stack', xTitle='Years', yTitle='Number of Reviews', title='Number of Reviews per Rating per Year')
# ### Helpfulness
df40 = df[['asin', 'helpful']]
# Create new Column for the enumerator
df40 = df40.assign(enum = df40['helpful'].apply(lambda enum_denom:enum_denom[0]))
# Create new Column for the denominator
df40 = df40.assign(denom = df40['helpful'].apply(lambda enum_denom:enum_denom[1]))
# Filter on the denom
df40 = df40.loc[df40['denom'] != 0]
df40[0:15]
len(df40)
bin_values = np.arange(start=0,stop=100,step=1)
df40['denom'].plot.hist(alpha=0.5, bins=bin_values, figsize=(15,7), grid=True, title='Distribution of Binary Helpfulness Ratings Counts per Review')
# Focus on [10,100] range of rating per review
df40 = df40.loc[df40['denom'] > 15]
df40 = df40.loc[df40['denom'] < 100]
len(df40)
df50 = df40.assign(percentage = df40['enum']/df40['denom'])
df50['percentage'].iplot(kind='histogram', title='Distribution of Helpfulness Percentage')
df50.head()
threshold = 0.7
df60 = df50.loc[df50['percentage'] > threshold]
len(df60)
# +
# END OF FILE
| notebooks/000_data_inspection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="tAbV8ljjT_fC" colab_type="text"
# # Introducition to Python 2
#
# ## Str - Lists - Tuples - Dicts
#
# ## Herramientas Computacionales
# ## Uniandes
#
# Based on [SoloLearn Python](https://www.sololearn.com/Play/Python)
# + [markdown] id="-FzvZO1uUmNC" colab_type="text"
# ## Strings
# + id="psqA04lLUnYV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277289620, "user_tz": 300, "elapsed": 474, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="52a256db-d252-4f76-a7ef-82d0e0559f31"
# Concatenation
print("Hello" +" " +"World")
# + id="QxSTmrZGVEEc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277292793, "user_tz": 300, "elapsed": 810, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="ba01d325-ca13-4eb4-d884-dd2a42de7234"
# Muliplication of strings
print("Hello"*3)
# + id="HFBUQMl-VN74" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277295192, "user_tz": 300, "elapsed": 929, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="a4dad89b-ef26-4bf1-f45e-968c40d19714"
# Join Function
name = " ".join(("Maria", "Judith", "Andrea"))
print(name)
# + id="Juu9mcwPYWwj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277297514, "user_tz": 300, "elapsed": 914, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="748334d5-79cb-4ec6-9587-7d723aa06d9b"
# Strings are unmutable
name.upper()
print(name)
# + id="pVSre8AxYdNi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277299701, "user_tz": 300, "elapsed": 1004, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="f0d1aab5-ab95-40b9-f87b-e8169bbe65f0"
# Upper or Lower Functions
name_upper = name.upper()
print(name_upper)
# + id="6rtj2emWVsCe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277301859, "user_tz": 300, "elapsed": 754, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="45ea1380-1afc-4db1-a4be-3f2d54994843"
# Split Function
name.split("a")
# + id="0nYYlaGqV-Gx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277304356, "user_tz": 300, "elapsed": 952, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="6950c46c-bbc2-4020-8021-493ab01e0e4e"
# String Formating
age = 56
print("Maria is {} years old".format(age))
# + id="gTRC1FvwWW64" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"status": "ok", "timestamp": 1600277307707, "user_tz": 300, "elapsed": 790, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="65b343bf-3edd-4bce-e853-102fa104f4e7"
# Access to substrings as a list
name[3:7]
# + [markdown] id="bgUZzwHQSG9K" colab_type="text"
#
# + id="9MThyP2KSMxK" colab_type="code" colab={}
age = 56
# + id="aKN3gzCbSIIa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600282797890, "user_tz": 300, "elapsed": 455, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="c1f2f61b-ec1d-4701-ea46-5e7ae10fcf89"
### format
print("La edad de Maria es {}".format(age))
# + [markdown] id="V437kox8Wttv" colab_type="text"
# ## Lists
# + id="u0WVYnhBWkQS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} executionInfo={"status": "ok", "timestamp": 1600277311434, "user_tz": 300, "elapsed": 722, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="a755e86d-221c-4b29-8eb3-db1f1ad5131d"
# Taken from https://www.sololearn.com/Play/Python
squares = [0, 1, 4, 9, 16, 25, 36, 49, 64, 81]
print(squares[2:6])
print(squares[3:8])
print(squares[0:1])
# + id="8bSE3OS1YEA-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277314167, "user_tz": 300, "elapsed": 799, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="d3cf5a2e-8186-4639-8c4e-78efa66680b2"
# Find the lenght of a list
len(squares)
# + id="NhXuNeclW5Av" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"status": "ok", "timestamp": 1600277316954, "user_tz": 300, "elapsed": 672, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="3a217849-7ebb-4740-d0d7-6758f14faee1"
# Taken from https://www.sololearn.com/Play/Python
print(squares[::2])
print(squares[2:8:3])
# + id="ivzIxyqvXIcy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277319974, "user_tz": 300, "elapsed": 957, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="f8dd6dbf-4404-4b85-fb47-8271a5d76e18"
# List Comprehension
print([i**2 for i in range(10)])
# + id="ZZyvlbDaXawc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277322171, "user_tz": 300, "elapsed": 782, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="c1923fcd-f88f-41dc-b4a6-9223e7d84fae"
# List inside list
matrix = [[1, 2],[3, 4]]
print(matrix)
# + id="N_08Y_xFZra1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277329847, "user_tz": 300, "elapsed": 640, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="97876ecc-9a7f-4825-c74f-1eef3be522a1"
matrix[1][1] = 7
print(matrix)
# + id="fMjkhyJRXyvY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} executionInfo={"status": "ok", "timestamp": 1600277332120, "user_tz": 300, "elapsed": 991, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="f7f1c664-3c73-45ea-b6cb-1e7f6410c2c3"
print(matrix[0])
print(matrix[1])
print(matrix[0][1])
# + [markdown] id="WUjNB3-oZR5t" colab_type="text"
# ## Tuples
# + id="4YaoKF7lZDYf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} executionInfo={"status": "error", "timestamp": 1600277336290, "user_tz": 300, "elapsed": 867, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="8f86203e-c854-4f2d-961f-141a0cbf62fa"
# Tuples are inmutable
# Taken from https://www.sololearn.com/Play/Python
words = ("spam", "eggs", "sausages")
words[1] = "cheese"
# + id="pw0Hf-wWZkVV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277339628, "user_tz": 300, "elapsed": 728, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="adecda8b-3a1e-4f40-a1a5-ef27c5bf6c65"
# List of words
words = ["spam", "eggs", "sausages", "eggs"]
print(words)
# + id="56pqQJSpaLMA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277341887, "user_tz": 300, "elapsed": 735, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="69799e26-3093-4060-87d7-675fafd99bbb"
words_tpl = tuple(words)
print(words_tpl)
# + [markdown] id="z3vDZ178cxSQ" colab_type="text"
# ## Dictionaries
# + id="1u4_8vpra5G5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277346041, "user_tz": 300, "elapsed": 855, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="98ba0e0a-5ac7-456c-da44-096ca3c433a4"
# Dictionaries map keys to value, keys must be inmutable
ages = {"David":34, "Felipe":50, "Ana": 25}
type(ages)
# + id="2LVgpByrdFfi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277347929, "user_tz": 300, "elapsed": 630, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="411cb6cb-a05b-4500-c5e4-26b0ca42a3fc"
ages["Felipe"]
# + id="cXaiq-fGdSs0" colab_type="code" colab={}
# Change the value of the dictionary
ages["Felipe"] = 80
# + id="QVfFphnZdnYo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277350856, "user_tz": 300, "elapsed": 914, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="5cf9587d-0ea4-4ee6-e58f-20ba59fbad1c"
print(ages)
# + id="YVQKSugwdogo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277424767, "user_tz": 300, "elapsed": 657, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="609d82db-9e9f-418e-f374-3d5cc5f81ac6"
# Check wheter an element is in the dictinary
print("Camila" in ages)
# + id="vXi74GfjeKMx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277430639, "user_tz": 300, "elapsed": 675, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="a6280dde-7078-4d50-cd38-622cd5098ecd"
# Use one dictionary function
ages.keys()
# + id="egAJ5l0T99-K" colab_type="code" colab={}
ages["Camila"] = 18
# + id="OszZIkH2-HZc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600277482060, "user_tz": 300, "elapsed": 511, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="eda0a0e1-5129-4724-92e6-b6bffbb7b977"
ages
# + id="6TIwNnTQfMCV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600204755817, "user_tz": 300, "elapsed": 593, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="34f298c7-230b-43dc-e28b-ae1eb083b430"
integers = []
i = 1
while 54*i < 1000:
integers.append(54*i)
i += 1
print(integers)
# + id="TApHuQnVoi3-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1600204776795, "user_tz": 300, "elapsed": 478, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="00fd8584-f22b-4ecd-eb3b-775b1619b678"
sum(integers)
# + [markdown] id="EQ95z7C8uaFs" colab_type="text"
# ## Exercise
# + id="HovPi-gvowNH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"status": "ok", "timestamp": 1600205378777, "user_tz": 300, "elapsed": 2924, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg6lrk9agmplTKK54rGVqavR8_tM4odcJqDil2B2w=s64", "userId": "02270832771774592167"}} outputId="f720f311-fd75-4a5e-9a6d-4c1a56048ece"
number = input("Por favor ingrese su numero magico:")
integers = []
i = 1
number = int(number)
while number*i < 1000:
integers.append(54*i)
i += 1
print("Los múltiplos de {} entre 1 y 1000 son:".format(number))
print(integers)
print("Su suma es: {}".format(sum(integers)))
| Notebooks/python 2 - lists tuples dicts strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''base'': conda)'
# name: python3710jvsc74a57bd0b3ba2566441a7c06988d0923437866b63cedc61552a5af99d1f4fb67d367b25f
# ---
# # Player Logs
# +
# %config IPCompleter.greedy=True
# %matplotlib inline
# Import the dependencies.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# Path
from pathlib import Path
# NBA
import nba_api
from nba_api.stats.static import players
from nba_api.stats.static import teams
from nba_api.stats.endpoints import leaguegamefinder,boxscoretraditionalv2, boxscoreadvancedv2, playergamelog
import requests
# -
# ## Look Up Tables
#
# > We want to create a couple of look up tables so when we pull the previous day's games we could use team id or player id to filter and display the correct information.
# +
# This gets all the teams put in a dictionary
nba_teams = teams.get_teams()
print(nba_teams)
# This gets all the teams put in a dictionary
nba_players = players.get_players()
print(nba_players)
# Need to get team ids to a csv
nba_team_ids = pd.DataFrame(nba_teams)
nba_team_ids.to_csv('nba_team_ids.csv', index=False)
# Need to get team ids to a csv
nba_player_ids = pd.DataFrame(nba_players)
nba_player_ids.to_csv('nba_player_ids.csv', index=False)
# +
# From our Team ID look up table we want to go through all teams and grab all played games
# Create list of Team Ids from the nba_team_ids dataframe
# list_of_team_ids = nba_team_ids['id'].unique().tolist()
games_list = []
for value in nba_team_ids['id']:
gamefinder = leaguegamefinder.LeagueGameFinder(team_id_nullable=value, season_nullable="2020-21")
games = gamefinder.get_data_frames()[-1]
games_list.append(games)
games_df = pd.concat(df)
games_df
# -
print(games)
| exploratory/PlayerLogsRebounds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # More on python
# Python has many high-level builtin features, time to learn some more!
# ## 3.02 Functions
# Functions can be defined using a lambda expression or via `def`. Python provides for functions both positional and keyword-based arguments.
square = lambda x: x * x
square(10)
# roots of ax^2 + bx + c
quadratic_root = lambda a, b, c: ((-b - (b * b - 4 * a * c) ** .5) / (2 * a), (-b + (b * b - 4 * a * c) ** .5) / (2 * a))
quadratic_root(1, 5.5, -10.5)
# a clearer function using def
def quadratic_root(a, b, c):
d = (b * b - 4 * a * c) ** .5
coeff = .5 / a
return (coeff * (-b - d), coeff * (-b + d))
quadratic_root(1, 5.5, -10.5)
# Functions can have positional arguments and keyword based arguments. Positional arguments have to be declared before keyword args
# name is a positional argument, message a keyword argument
def greet(name, message='Hello {}, how are you today?'):
print(message.format(name))
greet('Tux')
greet('Tux', 'Hi {}!')
greet('Tux', message='What\'s up {}?')
# this doesn't work
greet(message="Hi {} !", 'Tux')
# keyword arguments can be used to define default values
# +
import math
def log(num, base=math.e):
return math.log(num) / math.log(base)
# -
log(math.e)
log(10)
log(1000, 10)
# ## 3.03 builtin functions, attributes
#
# Python provides a rich standard library with many builtin functions. Also, bools/ints/floats/strings have many builtin methods allowing for concise code.
#
# One of the most useful builtin function is `help`. Call it on any object to get more information, what methods it supports.
# +
s = 'This is a test string!'
print(s.lower())
print(s.upper())
print(s.startswith('This'))
print('string' in s)
print(s.isalnum())
# -
# For casting objects, python provides several functions closely related to the constructors
# `bool, int, float, str, list, tuple, dict, ...`
tuple([1, 2, 3, 4])
str((1, 2, 3))
str([1, 4.5])
# ## 4.01 Dictionaries
#
# Dictionaries (or associate arrays) provide a structure to lookup values based on keys. I.e. they're a collection of k->v pairs.
list(zip(['brand', 'model', 'year'], ['Ford', 'Mustang', 1964])) # creates a list of tuples by "zipping" two list
# convert a list of tuples to a dictionary
D = dict(zip(['brand', 'model', 'year'], ['Ford', 'Mustang', 1964]))
D
D['brand']
D = dict([('brand', 'Ford'), ('model', 'Mustang')])
D['model']
# Dictionaries can be also directly defined using `{ ... : ..., ...}` syntax
D = {'brand' : 'Ford', 'model' : 'Mustang', 'year' : 1964}
D
# +
# dictionaries have serval useful functions implemented
# help(dict)
# -
# adding a new key
D['price'] = '48k'
D
# removing a key
del D['year']
D
# checking whether a key exists
'brand' in D
# returning a list of keys
D.keys()
# casting to a list
list(D.keys())
D
# iterating over a dictionary
for k in D.keys():
print(k)
for v in D.values():
print(v)
for k, v in D.items():
print('{}: {}'.format(k, v))
# ## 4.02 Calling functions with tuples/dicts
#
# Python provides two special operators `*` and `**` to call functions with arguments specified through a tuple or dictionary. I.e. `*` unpacks a tuple into positional args, whereas `**` unpacks a dictionary into keyword arguments.
quadratic_root(1, 5.5, -10.5)
args=(1, 5.5, -10.5)
quadratic_root(*args)
args=('Tux',) # to create a tuple with one element, need to append , !
kwargs={'message' : 'Hi {}!'}
greet(*args, **kwargs)
# ## 4.03 Sets
#
# python has builtin support for sets (i.e. an unordered list without duplicates). Sets can be defined using `{...}`.\
#
# **Note:** `x={}` defines an empty dictionary! To define an empty set, use
S = set()
type(S)
S = {1, 2, 3, 1, 4}
S
2 in S
# +
# casting can be used to get unique elements from a list!
L = [1, 2, 3, 4, 3, 2, 5, 3, 65, 19]
list(set(L))
# -
# set difference via `-` or `difference`
{1, 2, 3} - {2, 3}, {1, 2, 3}.difference({2, 3})
# set union via `+` or `union`
{1, 2, 3} | {4, 5}, {1, 2, 3}.union({4, 5})
# set intersection via `&` or `intersection`
{1, 5, 3, 4} & {2, 3}
{1, 5, 3, 4}.intersection({2, 3})
# ## 4.04 Comprehensions
#
# Instead of creating list, dictionaries or sets via explicit extensional declaration, you can use a comprehension expression. This is especially useful for conversions.
# +
# list comprehension
L = ['apple', 'pear', 'banana', 'cherry']
[(1, x) for x in L]
# -
# special case: use if in comprehension for additional condition
[(len(x), x) for x in L if len(x) > 5]
# if else must come before for
# ==> here ... if ... else ... is an expression!
[(len(x), x) if len(x) % 2 == 0 else None for x in L]
# The same works also for sets AND dictionaries. The collection to iterate over doesn't need to be of the same type.
# +
L = ['apple', 'pear', 'banana', 'cherry']
length_dict = {k : len(k) for k in L}
length_dict
# +
import random
[random.randint(0, 10) for i in range(10)]
# -
{random.randint(0, 10) for _ in range(20)}
[(k, v) for k, v in length_dict.items()]
# filter out elements from dict based on condition
{k : v for k,v in length_dict.items() if k[0] < 'c'}
# ## 5.01 More on functions
# Nested functions + decorators
#
# ==> Functions are first-class citizens in python, i.e. we can return them
def make_plus_one(f):
def inner(x):
return f(x) + 1
return inner
# +
fun = make_plus_one(lambda x: x)
fun(2), fun(3), fun(4)
# -
# A more complicated function can be created to create functions to evaluate a polynomial defined through a vectpr $p = (p_1, ..., p_n)^T$
#
#
# $$ f(x) = \sum_{i=1}^n p_i x^i$$
def make_polynomial(p):
def f(x):
if 0 == len(p):
return 0.
y = 0
xq = 1
for a in p:
y += a * xq
xq *= x
return y
return f
poly = make_polynomial([1])
poly(2)
quad_poly = make_polynomial([1, 2, 1])
quad_poly(1)
# Basic idea is that when declaring nested functions, the inner ones have access to the enclosing functions scope. When returning them, a closure is created.
#
#
# We can use this to change the behavior of functions by wrapping them with another!
#
# ==> we basically decorate the function with another, thus the name decorator
def greet(name):
return 'Hello {}!'.format(name)
greet('Tux')
# Let's say we want to shout the string, we could do:
greet('Tux').upper()
# ==> however, we would need to change this everywhere
# However, what if we want to apply uppercase to another function?
def state_an_important_fact():
return 'The one and only answer to ... is 42!'
state_an_important_fact().upper()
# with a wrapper we could create an upper version
def make_upper(f):
def inner(*args, **kwargs):
return f(*args, **kwargs).upper()
return inner
GREET = make_upper(greet)
STATE_AN_IMPORTANT_FACT = make_upper(state_an_important_fact)
GREET('tux')
STATE_AN_IMPORTANT_FACT()
# Instead of explicitly having to create the decoration via make_upper, we can also use python's builtin support for this via the @ statement. I.e.
@make_upper
def say_hi(name):
return 'Hi ' + name + '.'
say_hi('sealion')
# It's also possible to use multiple decorators
def split(function):
def wrapper():
res = function()
return res.split()
return wrapper
@split
@make_upper
def state_an_important_fact():
return 'The one and only answer to ... is 42!'
state_an_important_fact()
# More on decorators here: <https://www.datacamp.com/community/tutorials/decorators-python>.
#
#
# ==> Flask (the framework we'll learn next week) uses decorators extensively, therefore they're included here.
# **Summary**: What is a decorator?
#
# A decorator is a design pattern to add/change behavior to an individual object. In python decorators are typically used for functions (later: also for classes)
# ## 6.01 Generators
#
# Assume we want to generate all square numbers. We could do so using a list comprehension:
[x * x for x in range(10)]
# However, this will create a list of all numbers. Sometimes, we just want to consume the number. I.e. we could do this via a function
square = lambda x: x * x
print(square(1))
print(square(2))
print(square(3))
# However, what about a more complicated sequence? I.e. fibonnaci numbers?
def fib(n):
if n <= 0:
return 0
if n <= 1:
return 1
a, b = 0, 1
for i in range(n):
a, b = b, a + b
return a
n = 10
for i in range(n):
print(fib(i))
# Complexity is n^2! However, with generators we can stop execution.
#
# The pattern is to call basically generator.next()
def fib():
a, b = 0, 1
yield a
while True:
a, b = b, a + b
yield a
fib()
g = fib()
for i in range(5):
print(next(g))
# `enumerate` and `zip` are both generator objects!
L = ['a', 'b', 'c', 'd']
g = enumerate(L)
print(next(g))
print(next(g))
print(next(g))
print(next(g))
# stop iteration exception will be done
print(next(g))
g = range(3)
g
# +
L = ['a', 'b', 'c', 'd']
i = list(range(len(L)))[::-1]
g = zip(L, i)
for el in g:
print(el)
# -
# **Note**: There is no hasNext in python. Use a loop with `in` to iterate over the full generator.
for i, n in enumerate(fib()):
if i > 10:
break
print(n)
# ## 7.01 Higher order functions
# python provides two builitn higher order functions: `map` and `filter`. A higher order function is a function which takes another function as argument or returns a function (=> decorators).
#
# In python3, `map` and `filter` yield a generator object.
map(lambda x: x * x, range(7))
for x in map(lambda x: x * x, range(7)):
print(x)
# +
# display squares which end with 1
list(filter(lambda x: x % 10 == 1, map(lambda x: x * x, range(25))))
# -
# ## 8.01 Basic I/O
# Python has builtin support to handle files
# +
f = open('file.txt', 'w')
f.write('Hello world')
f.close()
# -
# Because a file needs to be closed (i.e. the file object destructed), python has a handy statement to deal with auto-closing/destruction: The `with` statement.
with open('file.txt', 'r') as f:
lines = f.readlines()
print(lines)
# Again, `help` is useful to understand what methods a file object has
# +
# uncomment here to get the full help
# help(f)
# -
# # 7.01 classes
# In python you can define compound types using `class`
class Animal:
def __init__(self, name, weight):
self.name = name
self.weight = weight
def print(self):
print('{} ({} kg)'.format(self.name, self.weight))
def __str__(self):
return '{} ({} kg)'.format(self.name, self.weight)
dog = Animal('dog', 20)
dog
print(dog)
dog.print()
# Basic inheritance is supported in python
class Elephant(Animal):
def __init__(self):
Animal.__init__(self, 'elephant', 1500)
#alternative:
# super().__init__(...)
e = Elephant()
print(e)
# # 8.01 Modules and packages
# More on this at <https://docs.python.org/3.7/tutorial/modules.html>. A good explanation of relative imports can be found here <https://chrisyeh96.github.io/2017/08/08/definitive-guide-python-imports.html>.
#
#
# ==> Each file represents a module in python. One or more modules make up a package.
#
# Let's say we want to package our `quad_root` function into a separate module `solver`
# !rm -r solver*
# !ls
# +
# %%file solver.py
# a clearer function using def
def quadratic_root(a, b, c):
d = (b * b - 4 * a * c) ** .5
coeff = .5 / a
return (coeff * (-b - d), coeff * (-b + d))
# -
# !cat solver.py
import solver
solver.quadratic_root(1, 1, -2)
# Alternative is to import the name quadratic_root directly into the current scope
from solver import quadratic_root
quadratic_root(1, 1, -2)
# To import everything, you can use `from ... import *`. To import multiple specific functions, use `from ... import a, b`.
#
# E.g. `from flask import render_template, request, abort, jsonify, make_response`.
#
# To organize modules in submodules, subsubmodules, ... you can use folders.
# I.e. to import a function from a submodule, use `from solver.algebraic import quadratic_root`.
#
# There's a special file `__init__.py` that is added at each level, which gets executed when `import folder` is run.
# !rm *.py
# !mkdir -p solver/algebraic
# %%file solver/__init__.py
# this file we run when import solver is executed
print('import solver executed!')
# %%file solver/algebraic/__init__.py
# run when import solver.algebraic is used
print('import solver.algebraic executed!')
# +
# %%file solver/algebraic/quadratic.py
print('solver.algebraic.quadratic executed!')
# a clearer function using def
def quadratic_root(a, b, c):
d = (b * b - 4 * a * c) ** .5
coeff = .5 / a
return (coeff * (-b - d), coeff * (-b + d))
# +
# %%file test.py
import solver
# -
# !python3 test.py
# +
# %%file test.py
import solver.algebraic
# -
# !python3 test.py
# +
# %%file test.py
import solver.algebraic.quadratic
# +
# %%file test.py
import solver.algebraic.quadratic
# -
# !python3 test.py
# +
# %%file test.py
from solver.algebraic.quadratic import *
print(quadratic_root(1, 1, -2))
# -
# !python3 test.py
# One can also use relative imports to import from other files via `.` or `..`!
# %%file solver/version.py
__version__ = "1.0"
# !tree solver
# +
# %%file solver/algebraic/quadratic.py
from ..version import __version__
print('solver.algebraic.quadratic executed!')
print('package version is {}'.format(__version__))
# a clearer function using def
def quadratic_root(a, b, c):
d = (b * b - 4 * a * c) ** .5
coeff = .5 / a
return (coeff * (-b - d), coeff * (-b + d))
# -
# !python3 test.py
# This can be also used to bring certain functions into scope!
# +
# %%file solver/algebraic/__init__.py
from .quadratic import *
# use this to restrict what functions to "export"
__all__ = [quadratic_root.__name__]
# +
# %%file test.py
from solver.algebraic import *
print(quadratic_root(1, 1, -2))
# -
# !python3 test.py
# Of course there's a lot more on how to design packages in python! However, these are the essentials you need to know.
# *End of lecture*
| lecture15/02_More_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from joblib import load, dump
import seaborn as sns
colors = sns.color_palette(palette = 'rainbow',n_colors=5)
sns.palplot(colors)
# +
task_name = 'ESOL'
batch_sizes, res = load('./%s.x1.res' % task_name)
sns.set(style = 'white', font_scale = 1.6)
fig, axes = plt.subplots(ncols=2,figsize=(16,6))
for item, ax in zip(['val_loss', 'val_r2'], axes):
# l_16 = sum(pd.DataFrame(res[0])['process'])/3
# l_64 = sum(pd.DataFrame(res[1])['process'])/3
# l_128 = sum(pd.DataFrame(res[2])['process'])/3
l_16 = pd.DataFrame(res[0])['process'].iloc[1]
l_64 = pd.DataFrame(res[1])['process'].iloc[1]
l_128 = pd.DataFrame(res[2])['process'].iloc[1]
l16 = l_16[item].to_frame(name = '8')
l64 = l_64[item].to_frame(name = '64')
l128 = l_128[item].to_frame(name = '128')
df = pd.concat([ l16, l64, l128], axis=1)
df.columns.name = 'batch size'
df.iloc[:500].rolling(3).apply(np.mean).plot(colors = colors[-3:], lw = 2, ax = ax)
if item == 'val_loss':
item = 'MSE loss of validation set'
else:
item = 'R squared of validation set'
ax.set_ylabel(item, fontsize = 20)
ax.set_xlabel('Epoch', fontsize = 16)
fig.tight_layout()
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.3, hspace=None)
fig.savefig( 'val_%s.svg' % task_name, dpi=300, format='svg')
# -
dfr = pd.concat([pd.DataFrame(a) for a in res])
dfr = dfr[['batch_size', 'valid_rmse', 'valid_r2', 'test_rmse','test_r2']]
dfr = dfr.set_index('batch_size')
app = []
for i in dfr.columns:
df1 = dfr[i].to_frame(name = 'value')
df1['metric'] = i
app.append(df1)
dfapp = pd.concat(app).reset_index()
dfapp.batch_size = dfapp.batch_size.astype(str)
df1 = dfapp[dfapp.metric.isin(['valid_rmse', 'test_rmse'])]
df2 = dfapp[dfapp.metric.isin(['valid_r2', 'test_r2'])]
df1.metric = df1.metric.map({'valid_rmse': 'valid_best_rmse', 'test_rmse': 'test_rmse'})
df2.metric = df2.metric.map({'valid_r2': 'valid_best_r2', 'test_r2': 'test_r2'})
# +
sns.set(style = 'white', font_scale = 2)
fig, axes = plt.subplots(nrows=1, ncols=1, figsize=(9,6), sharex=False, sharey=False, )
ax1 = axes
g1 = sns.barplot(x = 'metric',y = 'value',data = df1, hue ='batch_size', ax=ax1,
hue_order=['8', '64', '128'], palette=colors.as_hex()[-3:], capsize = 0.1,
)
ax1.set_ylabel('RMSE')
ax1.set_xlabel('')
ax1.set_ylim(0.0, 0.8)
fontdict = {'fontsize':18, "color": "#663300"}
#[0.475, 0.494, 0.507, 0.547, 0.569, 0.56]
g1.text(-0.36, 0.38, '0.475', fontdict = fontdict)
g1.text(-0.10, 0.38, '0.494', fontdict = fontdict)
g1.text(0.16, 0.38, '0.507', fontdict = fontdict)
g1.text(1-0.36, 0.38, '0.547', fontdict = fontdict)
g1.text(1-0.10, 0.38, '0.569', fontdict = fontdict)
g1.text(1+0.16, 0.38, '0.560', fontdict = fontdict)
fig.tight_layout()
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.3, hspace=None)
fig.savefig( '1_val_test_bar_%s.svg' % task_name, format='svg')
# +
sns.set(style = 'white', font_scale = 2)
fig, axes = plt.subplots(nrows=1, ncols=1, figsize=(9,6), sharex=False, sharey=False, )
ax2 = axes
g2 = sns.barplot(x = 'metric',y = 'value',data = df2, hue ='batch_size', ax=ax2,
hue_order=['8', '64', '128'], palette=colors.as_hex()[-3:], capsize = 0.1,
)
ax2.set_ylabel('R2')
ax2.set_xlabel('')
ax2.set_ylim(0.0, 1.05)
#[0.946, 0.942, 0.938, 0.924, 0.92, 0.921]
g2.text(-0.36, 0.68, '0.946', fontdict = fontdict)
g2.text(-0.10, 0.68, '0.942', fontdict = fontdict)
g2.text(0.16, 0.68, '0.938', fontdict = fontdict)
g2.text(1-0.36, 0.68, '0.924', fontdict = fontdict)
g2.text(1-0.10, 0.68, '0.920', fontdict = fontdict)
g2.text(1+0.16, 0.68, '0.921', fontdict = fontdict)
fig.tight_layout()
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=0.3, hspace=None)
fig.savefig( '2_val_test_bar_%s.svg' % task_name, format='svg')
# -
df1.groupby(['metric','batch_size'])[['value']].apply(np.mean).sort_index(ascending = False).round(3)['value'].tolist()
df2.groupby(['metric','batch_size'])[['value']].apply(np.mean).sort_index(ascending = False).round(3)['value'].tolist()
pd.DataFrame(res[0])
pd.DataFrame(res[1])
pd.DataFrame(res[2])
| paper/09_batchsize_effect/04_analysis_plot_ESOL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear regression performance benchmark: scikit-learn vs. statsmodels
#
# <NAME>
#
# 2019-05-03
#
# This notebook tests the performance of running ordinary least square (OLS) linear regression on samples of an intermediate size (*n* ~ 10<sup>2</sup>–10<sup>4</sup>) using [Scikit-learn](https://scikit-learn.org/stable/) vs. using [Statsmodels](https://www.statsmodels.org/stable/index.html).
import numpy as np
import pandas as pd
import statsmodels.api as sm
from scipy.stats import pearsonr
from sklearn import datasets
from sklearn.linear_model import LinearRegression
# ## Version information
# +
import pkg_resources
for pkg in ["numpy", "scipy", "pandas", "statsmodels", "scikit-learn"]:
version = pkg_resources.get_distribution(pkg).version
print(f"{pkg} version = {version}")
# -
# ## Load the dataset
#
# Load the Boston house prices dataset from Scikit-learn.
dataset = datasets.load_boston()
X = dataset.data
y = dataset.target
# add constants
X_with_const = np.hstack([np.ones((X.shape[0], 1)), X])
# ## Benchmark regressions
#
# ### Baseline: normal equation
# +
coefs = np.linalg.solve(X_with_const.T @ X_with_const, X_with_const.T @ y)
r2 = pearsonr(X_with_const @ coefs, y)[0] ** 2
print(f"""Linear regression results from the normal equation
* coefs = {coefs}
* R^2 = {r2}""")
# -
# %%timeit -n 1000
coefs = np.linalg.solve(X_with_const.T @ X_with_const, X_with_const.T @ y)
r2 = pearsonr(X_with_const @ coefs, y)[0] ** 2
# ### Statsmodels OLS
lm_sm = sm.OLS(y, X_with_const).fit()
print(lm_sm.summary())
# %%timeit -n 1000
lm_sm = sm.OLS(y, X_with_const).fit()
# ## Scikit-learn linear model
#
sk_ols = LinearRegression(fit_intercept=False)
lm_sk = sk_ols.fit(X_with_const, y)
score_sk = lm_sk.score(X_with_const, y) # this calculates the R^2
print(f"""Scikit-learn linear regression results
* coefs = {lm_sk.coef_}
* R^2 = {score_sk}""")
# %%timeit -n 500
lm_sk = sk_ols.fit(X_with_const, y)
score_sk = lm_sk.score(X_with_const, y)
# ## Summary
#
# 1. Speed ranking: normal equation > Scikit-learn `LinearRegression` ≈ Statsmodels `OLS`.
# 2. The Statsmodels `OLS` class provides a rich set of statistics for diagnosing the goodness of the fit, which does not exist in the results from other methods.
# 3. Given that there is not a significant difference between the performance of Scikit-learn `LinearRegression` and that of Statsmodels `OLS`, **Statsmodels should be the preferred package for performing linear regression in Python**.
# 4. For bootstrapping when the performance of a single iteration is critical, the normal equation may be preferred.
| statsmodels/benchmark_linreg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# <center><h1> Assignment I </h1><br> <h2>Linear Regression</h2></center>
#
# + [markdown] colab_type="text" id="ylRMAvt4VYPM"
# # Generating Synthetic Data
# + [markdown] colab_type="text" id="siDqkSD4OvA8"
# This assignment shows how we can extend ordinary least squares regression, which uses the hypothesis class of linear regression functions, to non-linear regression functions modeled using polynomial basis functions and radial basis functions. The function we want to fit is $y_\mathsf{true} \, = \, f_\mathsf{true}(x) \, = \, 6 (\sin(x + 2) + \sin(2x + 4)) $. This is a **univariate function** as it has only one input variable. First, we generate synthetic input (data) $x_i$ by sampling $n=750$ points from a uniform distribution on the interval $[-7.5, \, 7.5]$.
# + colab={} colab_type="code" id="Zss-4fRm1lhx"
# The true function
def f_true(x):
y = 6.0 * (np.sin(x + 2) + np.sin(2*x + 4))
return y
# + [markdown] colab_type="text" id="KvCvfnxhB5zY"
# We can generate a synthetic data set, with Gaussian noise.
# + colab={} colab_type="code" id="NP0yQliVP_ST"
import numpy as np # For all our math needs
n = 750 # Number of data points
X = np.random.uniform(-7.5, 7.5, n) # Training examples, in one dimension
e = np.random.normal(0.0, 5.0, n) # Random Gaussian noise
y = f_true(X) + e # True labels with noise
# + [markdown] colab_type="text" id="__4x3YpHQ7f0"
# Now, we plot the raw data as well as the true function (without noise).
# + colab={"base_uri": "https://localhost:8080/", "height": 364} colab_type="code" id="frdKT7-lRF_c" outputId="1fdc1039-7781-4d0a-c4ae-2cbcd002d121"
import matplotlib.pyplot as plt # For all our plotting needs
plt.figure()
# Plot the data
plt.scatter(X, y, 12, marker='o')
# Plot the true function, which is really "unknown"
x_true = np.arange(-7.5, 7.5, 0.05)
y_true = f_true(x_true)
plt.plot(x_true, y_true, marker='None', color='r')
# + [markdown] colab_type="text" id="jvz-0RjMqtT9"
# Recall that we want to build a model to **generalize well on future data**, and in order to generalize well on future data, we need to pick a model that trade-off well between fit and complexity (that is, bias and variance). We randomly split the overall data set ($\mathcal{D}$) into three subsets:
# * **Training set**: $\mathcal{D}_\mathsf{trn}$ consists of the actual training examples that will be used to **train the model**;
# * **Validation set**: $\mathcal{D}_\mathsf{val}$ consists of validation examples that will be used to **tune model hyperparameters** (such as $\lambda > 0$ in ridge regression) in order to find the best trade-off between fit and complexity (that is, the value of $\lambda$ that produces the best model);
# * **Test set**: $\mathcal{D}_\mathsf{tst}$ consists of test examples to **estimate how the model will perform on future data**.
# 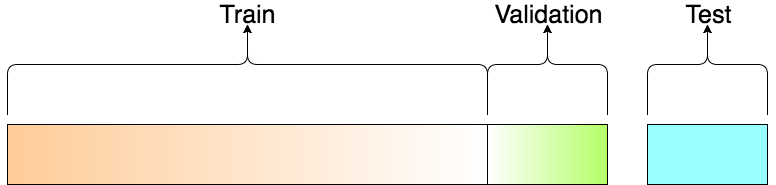
#
# For this example, let us randomly partition the data into three non-intersecting sets: $\mathcal{D}_\mathsf{trn} = 60\%$ of $\mathcal{D}$, $\mathcal{D}_\mathsf{val} = 10\%$ of $\mathcal{D}$ and $\mathcal{D}_\mathsf{tst} = 30\%$ of $\mathcal{D}$.
# + colab={"base_uri": "https://localhost:8080/", "height": 364} colab_type="code" id="ONF5BvRIj0b5" outputId="baa43a32-dedc-4b33-a283-27c23b47778e"
# scikit-learn has many tools and utilities for model selection
from sklearn.model_selection import train_test_split
tst_frac = 0.3 # Fraction of examples to sample for the test set
val_frac = 0.1 # Fraction of examples to sample for the validation set
# First, we use train_test_split to partition (X, y) into training and test sets
X_trn, X_tst, y_trn, y_tst = train_test_split(X, y, test_size=tst_frac, random_state=42)
# Next, we use train_test_split to further partition (X_trn, y_trn) into training and validation sets
X_trn, X_val, y_trn, y_val = train_test_split(X_trn, y_trn, test_size=val_frac, random_state=42)
# Plot the three subsets
plt.figure()
plt.scatter(X_trn, y_trn, 12, marker='o', color='orange')
plt.scatter(X_val, y_val, 12, marker='o', color='green')
plt.scatter(X_tst, y_tst, 12, marker='o', color='blue')
# + [markdown] colab_type="text" id="OqcB075kVcvr"
# # 1. <font color='#556b2f'> **Regression with Polynomial Basis Functions**</font>, 30 points.
# + [markdown] colab_type="text" id="4aL3ual-l0zb"
# This problem extends **ordinary least squares regression**, which uses the hypothesis class of _linear regression functions_, to _non-linear regression functions_ modeled using **polynomial basis functions**. In order to learn nonlinear models using linear regression, we have to explicitly **transform the data** into a higher-dimensional space. The nonlinear hypothesis class we will consider is the set of $d$-degree polynomials of the form $f(x) \, = \, w_0 + w_1 x + w_2 x^2 + ... + w_d x^d$ or **a linear combination of polynomial basis function**:
#
# <div align="center">
# $
# f(x) = [w_0, \, w_1,\, w_2 \, ..., w_d]^T \left[ \begin{array}{c} 1 \\ x \\ x^2 \\ \vdots \\ x^d\end{array} \right]
# $.
# </div>
#
# The monomials $\{ 1, \, x, \, x^2, \, ..., \, x^d\}$ are called **basis functions**, and each basis function $x^k$ has a corresponding weight $w_k$ associated with it, for all $k \, = \, 1, ..., d$. We transform each univariate data point $x_i$ into into a multivariate ($d$-dimensional) data point via $\phi(x_i) \rightarrow [1, \, x_i, \, x_i^2, \, \, ..., \, x^d_i]$. When this transformation is applied to every data point, it produces the **Vandermonde matrix**:
#
# <div align="center">
# $
# \Phi \, = \,
# \left[
# \begin{array}{ccccc}
# 1 & x_1 & x_1^2 & ... & x_1^d\\
# 1 & x_2 & x_2^2 & ... & x_2^d\\
# \vdots & \vdots & \vdots & \ddots & \vdots\\
# 1 & x_n & x_n^2 & \cdots & x_n^d\\
# \end{array}
# \right]
# $.
# </div>
# + [markdown] colab_type="text" id="mYeBBOe0K-NG"
# ---
# ### **a**. (10 points)
# Complete the Python function below that takes univariate data as input and computes a Vandermonde matrix of dimension $d$. This transforms one-dimensional data into $d$-dimensional data in terms of the polynomial basis and allows us to model regression using a $d$-degree polynomial.
# + colab={} colab_type="code" id="KwoVO3Cal3Iz"
# X float(n, ): univariate data
# d int: degree of polynomial
def polynomial_transform(X, d):
# convert data to np.array
X = np.array(X)
# power univariate data to d-th and transpose
return np.transpose(np.array([np.power(X, i) for i in range(d + 1)]))
# + [markdown] colab_type="text" id="2UvaLwVPL8tD"
# ---
# ### **b**. (10 points)
# Complete the Python function below that takes a Vandermonde matrix $\Phi$ and the labels $\mathbf{y}$ as input and learns weights via **ordinary least squares regression**. Specifically, given a Vandermonde matrix $\Phi$, implement the computation of $\mathbf{w} \, = \, (\Phi^T \Phi)^{-1}\Phi^T\mathbf{y}$. _Remember that in Python, @ performs matrix multiplication, while * performs element-wise multiplication. Alternately, [numpy.dot](https://docs.scipy.org/doc/numpy-1.15.0/reference/generated/numpy.dot.html) also performs matrix multiplication._
# + colab={} colab_type="code" id="7Ks6HU01LHB0"
# Phi float(n, d): transformed data
# y float(n, ): labels
def train_model(Phi, y):
Phi_t = np.transpose(Phi) # transpose phi
# comptue weight
return np.dot(np.dot(np.linalg.inv(np.dot(Phi_t, Phi)),Phi_t), y)
# + [markdown] colab_type="text" id="qrLFAGW4OSBo"
# ---
# ### **c**. (5 points)
# Complete the Python function below that takes a Vandermonde matrix $\Phi$, corresponding labels $\mathbf{y}$, and a linear regression model $\mathbf{w}$ as input and evaluates the model using **mean squared error**. That is, $\epsilon_\mathsf{MSE} \, = \, \frac{1}{n} \sum_{i=1}^n \, (y_i \, - \, \mathbf{w}^T \Phi_i)^2$.
# + colab={} colab_type="code" id="QQoZE89VOQT3"
# Phi float(n, d): transformed data
# y float(n, ): labels
# w float(d, ): linear regression model
def evaluate_model(Phi, y, w):
# wT@phi -> converted to phi@w
# this is due to the piece-wise subtraction
return np.sum(np.power((y - np.dot(Phi, w)), 2)) / len(y)
# + [markdown] colab_type="text" id="p5BwgXfLQEih"
# ---
# ### **d**. (5 points, **Discussion**)
# We can explore the **effect of complexity** by varying $d = 3, 6, 9,\, \cdots, 24$ to steadily increase the non-linearity of the models. For each model, we train using the transformed training data ($\Phi$, whose dimension increases) and evaluate its performance on the transformed validation data and estimate what our future accuracy will be using the test data.
#
# From plot of $d$ vs. validation error below, which choice of $d$ do you expect will generalize best?
# + colab={} colab_type="code" id="28aQeQ5xRFxD"
w = {} # Dictionary to store all the trained models
validationErr = {} # Validation error of the models
testErr = {} # Test error of all the models
for d in range(3, 25, 3): # Iterate over polynomial degree
Phi_trn = polynomial_transform(X_trn, d) # Transform training data into d dimensions
w[d] = train_model(Phi_trn, y_trn) # Learn model on training data
Phi_val = polynomial_transform(X_val, d) # Transform validation data into d dimensions
validationErr[d] = evaluate_model(Phi_val, y_val, w[d]) # Evaluate model on validation data
Phi_tst = polynomial_transform(X_tst, d) # Transform test data into d dimensions
testErr[d] = evaluate_model(Phi_tst, y_tst, w[d]) # Evaluate model on test data
# Plot all the models
plt.figure()
plt.plot(validationErr.keys(), validationErr.values(), marker='o', linewidth=3, markersize=12)
plt.plot(testErr.keys(), testErr.values(), marker='s', linewidth=3, markersize=12)
plt.xlabel('Polynomial degree', fontsize=16)
plt.ylabel('Validation/Test error', fontsize=16)
plt.xticks(list(validationErr.keys()), fontsize=12)
plt.legend(['Validation Error', 'Test Error'], fontsize=16)
plt.axis([2, 25, 15, 60])
# + [markdown] colab_type="text" id="1VZZKMgFdAd1"
# ---
# Finally, let's visualize each learned model.
# + colab={} colab_type="code" id="XZCJZdiedPMA"
plt.figure()
plt.plot(x_true, y_true, marker='None', linewidth=5, color='k')
for d in range(9, 25, 3):
X_d = polynomial_transform(x_true, d)
y_d = X_d @ w[d]
plt.plot(x_true, y_d, marker='None', linewidth=2)
plt.legend(['true'] + list(range(9, 25, 3)))
plt.axis([-8, 8, -15, 15])
# + [markdown] colab_type="text" id="Qooi6kMBVfJN"
# ---
# # 2. <font color='#556b2f'> **Regression with Radial Basis Functions**</font>, 70 points
#
# + [markdown] colab_type="text" id="Pj4oYRGBl6xz"
# In the previous case, we considered a nonlinear extension to linear regression using a linear combination of polynomial basis functions, where each basis function was introduced as a feature $\phi(x) = x^k$. Now, we consider Gaussian radial basis functions of the form:
# <div align="center">
# $\phi(\mathbf{x}) = e^{-\gamma \, (x - \mu)^2}$,
# </div>
# whose shape is defined by its center $\mu$ and its width $\gamma > 0$. In the case of polynomial basis regression, the user's choice of the dimension $d$ determined the transformation and the model. For radial basis regression, we have to contend with deciding how many radial basis functions we should have, and what their center and width parameters should be. For simplicity, let's assume that $\gamma = 0.1$ is fixed. Instead of trying to identify the number of radial basis functions or their centers, we can treat **each data point as the center of a radial basis function**, which means that the model will be:
#
# <div align="center">
# $
# f(x) = [w_1, \, w_2,\, w_3 \, ..., w_n]^T \left[ \begin{array}{c} e^{-\gamma \, (x - x_1)^2} \\ e^{-\gamma \, (x - x_2)^2} \\ e^{-\gamma \, (x - x_2)^2} \\ ... \\ e^{-\gamma \, (x - x_n)^2} \end{array} \right]
# $.
# </div>
#
# This transformation uses radial basis functions centered around data points $e^{-\gamma \, (x - x_i)^2}$ and each basis function has a corresponding weight $w_i$ associated with it, for all $i \, = \, 1, ..., n$. We transform each univariate data point $x_j$ into into a multivariate ($n$-dimensional) data point via $\phi(x_j) \rightarrow [..., e^{-\gamma \, (x_j - x_i)^2}, \, ...]$. When this transformation is applied to every data point, it produces the **radial-basis kernel**:
#
# <div align="center">
# $
# \Phi \, = \,
# \left[
# \begin{array}{ccccc}
# 1 & e^{-\gamma \, (x_1 - x_2)^2} & e^{-\gamma \, (x_1 - x_3)^2} & ... & e^{-\gamma \, (x_1 - x_n)^2}\\
# e^{-\gamma \, (x_2 - x_1)^2} & 1 & e^{-\gamma \, (x_2 - x_3)^2} & ... & e^{-\gamma \, (x_2 - x_n)^2}\\
# \vdots & \vdots & \vdots & \ddots & \vdots\\
# e^{-\gamma \, (x_n - x_1)^2} & e^{-\gamma \, (x_n - x_2)^2} & e^{-\gamma \, (x_n - x_3)^2} & \cdots & 1\\
# \end{array}
# \right]
# $.
# </div>
# + [markdown] colab_type="text" id="a6XGeuTbjjku"
# ---
# ### **a**. (15 points)
# Complete the Python function below that takes univariate data as input and computes a radial-basis kernel. This transforms one-dimensional data into $n$-dimensional data in terms of Gaussian radial-basis functions centered at each data point and allows us to model nonlinear (kernel) regression.
# + colab={} colab_type="code" id="DpqR-II-jjky"
# X float(n, ): univariate data
# B float(n, ): basis functions
# gamma float : standard deviation / scaling of radial basis kernel
def radial_basis_transform(X, B, gamma=0.1):
B = np.array(B)
# compute radial basis
return np.exp(-gamma * np.square(np.array([B - x for x in X])))
# + [markdown] colab_type="text" id="ZBMj1VpWm5lq"
# ---
# ### **b**. (15 points)
# Complete the Python function below that takes a radial-basis kernel matrix $\Phi$, the labels $\mathbf{y}$, and a regularization parameter $\lambda > 0$ as input and learns weights via **ridge regression**. Specifically, given a radial-basis kernel matrix $\Phi$, implement the computation of $\mathbf{w} \, = \, \left( \Phi^T \Phi + \lambda I_n \right)^{-1} \, \Phi^T\mathbf{y}$.
# + colab={} colab_type="code" id="QienR3VVm5lu"
# Phi float(n, d): transformed data
# y float(n, ): labels
# lam float : regularization parameter
def train_ridge_model(Phi, y, lam):
# transpose phi
Phi_t = np.transpose(Phi)
return np.dot(np.dot(
np.linalg.inv(np.dot(Phi_t, Phi) + np.dot(lam, np.identity(Phi.shape[-1]))), Phi_t), y)
# + [markdown] colab_type="text" id="FQ1_cQmzpYM5"
# ---
# ### **c**. (30 points)
# As before, we can explore the tradeoff between **fit and complexity** by varying $\lambda \in [10^{-3}, 10^{-2}\, \cdots, 1, \, \cdots\, 10^3]$. For each model, train using the transformed training data ($\Phi$) and evaluate its performance on the transformed validation and test data. Plot two curves: (i) $\lambda$ vs. validation error and (ii) $\lambda$ vs. test error, as above.
#
# What are some ideal values of $\lambda$?
# + colab={} colab_type="code" id="Y3hsN6OkpYM8"
w = {} # Dictionary to store all the trained models
validationErr = {} # Validation error of the models
testErr = {} # Test error of all the models
Phi_trn = radial_basis_transform(X_trn, X_trn, gamma=0.1)
Phi_val = radial_basis_transform(X_val, X_trn, gamma=0.1)
Phi_tst = radial_basis_transform(X_tst, X_trn, gamma=0.1)
for lam in range(-3, 4): # lambdas
w[lam] = train_ridge_model(Phi_trn, y_trn, 10 ** lam)
validationErr[lam] = evaluate_model(Phi_val, y_val, w[lam]) # Evaluate model on validation data
testErr[lam] = evaluate_model(Phi_tst, y_tst, w[lam]) # Evaluate model on test data
# Plot lambda vs validation error
plt.figure()
plt.plot(validationErr.keys(), validationErr.values(), marker='o', linewidth=3, markersize=12)
plt.plot(testErr.keys(), testErr.values(), marker='s', linewidth=3, markersize=12)
plt.xlabel('Lambda values', fontsize=16)
plt.ylabel('Validation/Test error', fontsize=16)
plt.xticks(list(validationErr.keys()), fontsize=12)
plt.legend(['Validation Error', 'Test Error'], fontsize=16)
# + [markdown] colab_type="text" id="n426nYfCFVIL"
# ### **d**. (10 points, **Discussion**)
# Plot the learned models as well as the true model similar to the polynomial basis case above. How does the linearity of the model change with $\lambda$?
# + colab={} colab_type="code" id="VgJrAuKbyBHt"
plt.figure()
plt.plot(x_true, y_true, marker='None', linewidth=5, color='k')
for lam in range(-3, 4):
X_d = radial_basis_transform(x_true, X_trn, gamma = 0.1)
y_d = X_d @ w[lam]
plt.plot(x_true, y_d, marker='None', linewidth=2)
plt.legend(['true'] + list(range(-3, 4)), loc = 'lower right')
plt.axis([-8, 8, -15, 15])
# -
# Look at the plot, the lambda at -3 is most close to the true value. This is due to the larger lambda value that leads to larger oscilation. Hence, it is predicted that the smaller value may lead to the higher accuracy prediction.
| ml-course/assignment_1/dqn170000_assignment_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 5GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
train_df = pd.read_csv('/kaggle/input/train.csv')
test_df = pd.read_csv('/kaggle/input/test.csv')
submit = pd.read_csv('/kaggle/input/Submission.csv')
# -
train_df
train_df.isnull().sum()
test_df.isnull().sum()
train_df.describe()
train_df.hist(figsize=(8,8))
h = plt.hist((np.power(train_df.Item_Outlet_Sales, 1/3)), bins=70)
# +
# train_df.Item_Outlet_Sales = np.power(train_df.Item_Outlet_Sales, 1/3)
# -
train_df.info()
# +
num_cols = []
cat_cols = []
for col in train_df.columns:
if train_df[col].dtype=='int64' or train_df[col].dtype=='float64':
num_cols.append(col)
else:
cat_cols.append(col)
cat_cols
# -
for col in cat_cols:
d = dict()
d[col] = train_df[col].unique()
if len(d[col]) < 20:
print(d)
print(train_df.Item_Fat_Content.unique())
train_df.replace({'reg':'Regular','LF':'Low Fat','low fat':'Low Fat'},inplace = True)
test_df.replace({'reg':'Regular','LF':'Low Fat','low fat':'Low Fat'},inplace = True)
train_df['Item_Category'] = train_df['Item_Identifier'].apply(lambda x: x[0:2]).map({'FD':'Food',
'NC':'Non-Consumable',
'DR':'Drinks'})
test_df['Item_Category'] = test_df['Item_Identifier'].apply(lambda x: x[0:2]).map({'FD':'Food',
'NC':'Non-Consumable',
'DR':'Drinks'})
cat_cols.append('Item_Category')
for col in cat_cols[1:]:
sns.barplot(data=train_df, x=col, y='Item_Outlet_Sales')
t = plt.xticks(rotation=90)
plt.show()
sns.barplot(data=train_df, x='Outlet_Establishment_Year', y='Item_Outlet_Sales')
#x = 70, 138, 202
sns.scatterplot(data=train_df, x='Item_MRP', y='Item_Outlet_Sales')
plt.plot([70, 70], [0,12000])
plt.plot([137, 137], [0,12000])
plt.plot([203, 203], [0,12000])
# * 0-70
# * 71-138
# * 139-202
# * 203-..
# +
# def func(x):
# if x<71:
# return 1
# elif x<139:
# return 2
# elif x<203:
# return 3
# else:
# return 4
# train_df['Item_MRP_Classes'] = train_df.Item_MRP.apply(lambda x:func(x))
# test_df['Item_MRP_Classes'] = test_df.Item_MRP.apply(lambda x:func(x))
# -
train_df.columns
corr = train_df.corr()
corr['Item_Outlet_Sales']
_ = train_df.groupby(['Item_Category', 'Item_Fat_Content', 'Item_Type']).describe()['Item_Weight']['mean'].reindex()
_
# +
for i in range(len(_.index)):
row = _.index[i]
train_df.loc[(train_df['Item_Weight'].isnull()) & (train_df['Item_Category']==row[0]) & (train_df['Item_Fat_Content']==row[1]) & (train_df['Item_Type']==row[2]), 'Item_Weight'] = _.values[i]
train_df['Item_Weight'] = train_df['Item_Weight'].fillna((train_df['Item_Weight']).mean())
for i in range(len(_.index)):
row = _.index[i]
test_df.loc[(test_df['Item_Weight'].isnull()) & (test_df['Item_Category']==row[0]) & (test_df['Item_Fat_Content']==row[1]) & (test_df['Item_Type']==row[2]), 'Item_Weight'] = _.values[i]
test_df['Item_Weight'] = test_df['Item_Weight'].fillna((test_df['Item_Weight']).mean())
# -
train_df.groupby(['Outlet_Location_Type','Outlet_Identifier'])['Outlet_Type'].value_counts()
train_df.groupby(['Outlet_Location_Type','Outlet_Identifier', 'Outlet_Type'])['Outlet_Size'].value_counts()
# So Outlets OUT017, OUT045, OUT010 have outlet_size as missing values.
# Now
# 1. Since OUT010 is a Grocery Store of tier 3 we can fill with medium in outlet_size
# 2. Since OUT017 and OUT045 are of supermarkettype 1 of tier 2, it's outlet_size should be small
# +
train_df[train_df['Outlet_Identifier']=='OUT010'] = \
train_df[train_df['Outlet_Identifier']=='OUT010'].fillna('Medium', axis=0)
train_df[train_df['Outlet_Identifier']=='OUT045'] = \
train_df[train_df['Outlet_Identifier']=='OUT045'].fillna('Small', axis=0)
train_df[train_df['Outlet_Identifier']=='OUT017'] = \
train_df[train_df['Outlet_Identifier']=='OUT017'].fillna('Small', axis=0)
test_df[test_df['Outlet_Identifier']=='OUT010'] = \
test_df[test_df['Outlet_Identifier']=='OUT010'].fillna('Medium', axis=0)
test_df[test_df['Outlet_Identifier']=='OUT045'] = \
test_df[test_df['Outlet_Identifier']=='OUT045'].fillna('Small', axis=0)
test_df[test_df['Outlet_Identifier']=='OUT017'] = \
test_df[test_df['Outlet_Identifier']=='OUT017'].fillna('Small', axis=0)
# -
cat_cols
# +
dum_cols = ['Item_Fat_Content', 'Outlet_Size',
'Outlet_Location_Type',
'Outlet_Type',
'Item_Category']
categ_cols = ['Item_Type','Outlet_Identifier']
# +
from sklearn.preprocessing import LabelEncoder
for col in cat_cols:
le = LabelEncoder()
train_df[col] = le.fit_transform(train_df[col])
test_df[col] = le.transform(test_df[col])
# train_df = pd.get_dummies(data=train_df, columns=dum_cols, drop_first=True)
# test_df = pd.get_dummies(data=test_df, columns=dum_cols, drop_first=True)
# -
train_df.head(2)
train_df['fat_per_weight'] = train_df['Item_Fat_Content']/train_df['Item_Weight']
test_df['fat_per_weight'] = test_df['Item_Fat_Content']/test_df['Item_Weight']
train_df.columns
# +
# features = ['Item_Weight', 'Item_Visibility', 'Item_Type',
# 'Item_MRP', 'Outlet_Identifier', 'Outlet_Establishment_Year',
# 'Item_Fat_Content_Regular', 'Outlet_Size_Medium',
# 'Outlet_Size_Small', 'Outlet_Location_Type_Tier 2',
# 'Outlet_Location_Type_Tier 3', 'Outlet_Type_Supermarket Type1',
# 'Outlet_Type_Supermarket Type2', 'Outlet_Type_Supermarket Type3',
# 'Item_Category_Food', 'Item_Category_Non-Consumable', 'Item_Outlet_Sales']
# corr = train_df[features].corr()
# features = features[:-1]
# corr['Item_Outlet_Sales']
# -
features = ['Item_Fat_Content', 'Item_Visibility', 'fat_per_weight',
'Item_Type', 'Item_MRP', 'Outlet_Identifier',
'Outlet_Size', 'Outlet_Location_Type', 'Outlet_Establishment_Year',
'Outlet_Type', 'Item_Category', 'Item_Outlet_Sales']
corr = train_df[features].corr()
features = features[:-1]
corr['Item_Outlet_Sales']
# +
from sklearn.model_selection import train_test_split
train_X, val_X, train_y, val_y = train_test_split(train_df[features], train_df['Item_Outlet_Sales'], test_size=0.2, random_state=42)
# +
from sklearn.metrics import mean_squared_error as mse
def get_e(model, df, target):
pred = model.predict(df)
print(len(pred[pred<=0]))
pred[pred<=0] = 2200
e = mse(target, pred)
print(np.sqrt(e))
def preds_to_csv(model, name):
pred = model.predict(test_df[features])
pred[pred<=0] = 2200
submit['Item_Outlet_Sales'] = pred
submit.to_csv('big_mart_sales_'+name+'.csv', index=False)
# pd.DataFrame({'Item_Identifier':test_df.Item_Identifier, 'Outlet_Identifier':test_df.Outlet_Identifier, 'Item_Outlet_Sales':pred}).to_csv(name+'_big_mart_sales.csv', index=False)
# -
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor, ExtraTreesRegressor
from sklearn.linear_model import LinearRegression
import xgboost as xgb
import lightgbm as lgb
# rm -r *.csv
# +
# rfr = RandomForestRegressor()
# rfr.fit(train_df[features], train_df.Item_Outlet_Sales)
# get_e(rfr, val_X, val_y)
# preds_to_csv(rfr, 'rfr')
# +
# from sklearn.model_selection import KFold
# gbr = GradientBoostingRegressor()
# X = train_df[features].values
# y = train_df['Item_Outlet_Sales'].values
# kf = KFold(n_splits=3, random_state=42)
# for train_index, test_index in kf.split(X):
# X_train, X_val = X[train_index], X[test_index]
# y_train, y_val = y[train_index], y[test_index]
# gbr.fit(X_train, y_train)
# get_e(gbr, X_val, y_val)
# get_e(gbr, val_X, val_y)
# preds_to_csv(gbr, 'k_fold_gbr')
# -
gbr = GradientBoostingRegressor()
gbr.fit(train_df[features], train_df.Item_Outlet_Sales)
get_e(gbr, val_X, val_y)
preds_to_csv(gbr, 'gbr')
gbr.get_params()
parameters = {
"learning_rate": [0.01, 0.05, 0.1, 0.15, 0.2],
"min_samples_split": np.linspace(0.1, 1, 6),
"min_samples_leaf": np.linspace(0.1, 0.5, 5),
"max_depth":[2,3,4,5,8],
"max_features":["auto", "log2","sqrt"],
"criterion": ["friedman_mse"],
"n_estimators":[50, 75, 100, 125, 150]
}
# +
from sklearn.model_selection import GridSearchCV
grid_search = GridSearchCV(gbr, parameters, cv=2, n_jobs=-1, verbose=1)
grid_search.fit(train_df[features], train_df.Item_Outlet_Sales)
get_e(grid_search, val_X, val_y)
preds_to_csv(grid_search, 'grid_search')
# -
grid_search.best_estimator_
grid_search.best_params_
# +
# lgbr = lgb.LGBMRegressor()
# lgbr.fit(train_df[features], train_df.Item_Outlet_Sales)
# get_e(lgbr, val_X, val_y)
# preds_to_csv(lgbr, 'lgbr')
# +
# lr = LinearRegression()
# lr.fit(train_X, train_y)
# get_e(lr, val_X, val_y)
# preds_to_csv(lr, 'lr')
# +
# xgbr = xgb.XGBRegressor()
# xgbr.fit(train_X, train_y)
# get_e(xgbr, val_X, val_y)
# preds_to_csv(xgbr, 'xgbr')
# +
# etr = ExtraTreesRegressor()
# etr.fit(train_X, train_y)
# get_e(etr, val_X, val_y)
# preds_to_csv(etr, 'etr')
# -
# GBR with whole train data gives a good score
# +
from tpot import TPOTRegressor
tpr = TPOTRegressor(generations=100, population_size=10, n_jobs=-1, early_stop=20, verbosity=3, random_state=42)
tpr.fit(train_df[features], train_df.Item_Outlet_Sales)
get_e(tpr, val_X, val_y)
preds_to_csv(tpr, 'tpr')
# -
tpr.fitted_pipeline_
| big-mart-sales-prediction/big-mart-sales-prediction (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Running database reconstruction attacks on the Iris dataset
# In this tutorial we will show how to run a database reconstruction attack on the Iris dataset and evaluate its effectiveness against models trained non-privately (i.e., naively with scikit-learn) and models trained with differential privacy guarantees.
# ## Preliminaries
# The database reconstruction attack takes a trained machine learning model `model`, which has been trained by a training dataset of `n` examples. Then, using `n-1` examples of the training dataset (i.e., with the target row removed), we seek to reconstruct the `n`th example of the dataset by using `model`.
#
# In this example, we train a Gaussian Naive Bayes classifier (`model`) with the training dataset, then remove a single row from that dataset, and seek to reconstruct that row using `model`. For typical examples, this attack is successful up to machine precision.
#
# We then show that launching the same attack on a ML model trained with differential privacy guarantees provides protection for the traning dataset, and prevents learning the target row with precision.
# ## Example usage
# ## Load data
# First, we load the data of interest and split into train/test subsets.
# +
from sklearn import datasets
from sklearn.model_selection import train_test_split
import numpy as np
dataset = datasets.load_iris()
# -
x_train, x_test, y_train, y_test = train_test_split(dataset.data, dataset.target, test_size=0.2)
# ## Train model
# We can now train a Gaussian naive Bayes classifier using the full training dataset. This is the model that will be used to attack the training dataset later.
# +
import sklearn.naive_bayes as naive_bayes
from art.estimators.classification.scikitlearn import ScikitlearnGaussianNB
model1 = naive_bayes.GaussianNB().fit(x_train, y_train)
non_private_art = ScikitlearnGaussianNB(model1)
# -
print("Model accuracy (on the test dataset): {}".format(model1.score(x_test, y_test)))
# ## Launch and evaluate attack
# We now select a row from the training dataset that we will remove. This is the **target row** which the attack will seek to reconstruct. The attacker will have access to `x_public` and `y_public`.
# +
target_row = int(np.random.random() * x_train.shape[0])
x_public = np.delete(x_train, target_row, axis=0)
y_public = np.delete(y_train, target_row, axis=0)
# -
# We can now launch the attack, and seek to infer the value of the target row. This is typically completed in less than a second.
# +
from art.attacks.inference.reconstruction import DatabaseReconstruction
dbrecon = DatabaseReconstruction(non_private_art)
x, y = dbrecon.reconstruct(x_public, y_public)
# -
# We can evaluate the accuracy of the attack using root-mean-square error (RMSE), showing a high level of accuracy in the inferred value.
print("Inference RMSE: {}".format(
np.sqrt(((x_train[target_row] - x) ** 2).sum() / x_train.shape[1])))
# We can confirm that the attack also inferred the correct label `y`.
np.argmax(y) == y_train[target_row]
# # Attacking a model trained with differential privacy
# We can mitigate against this attack by training the public ML model with differential privacy. We will use [diffprivlib](https://github.com/Trusted-AI/differential-privacy-library) to train a differentially private Guassian naive Bayes classifier. We can mitigate against any loss in accuracy of the model by choosing an `epsilon` value appropriate to our needs.
# ## Train the model
# +
from diffprivlib import models
model2 = models.GaussianNB(bounds=([4.3, 2.0, 1.1, 0.1], [7.9, 4.4, 6.9, 2.5]), epsilon=3).fit(x_train, y_train)
private_art = ScikitlearnGaussianNB(model2)
model2.score(x_test, y_test)
# -
# ## Launch and evaluate attack
# We then launch the same attack as before. In this case, the attack may take a number of seconds to return a result.
# +
dbrecon = DatabaseReconstruction(private_art)
x_dp, y_dp = dbrecon.reconstruct(x_public, y_public)
# -
# In this case, the RMSE shows our attack has not been as successful
print("Inference RMSE (with differential privacy): {}".format(
np.sqrt(((x_train[target_row] - x_dp) ** 2).sum() / x_train.shape[1])))
# This is confirmed by inspecting the inferred value and the true value.
x_dp, x_train[target_row]
# In fact, the attack may not even be able to correctly infer the target label.
np.argmax(y_dp), y_train[target_row]
| notebooks/attack_database_reconstruction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext watermark
# %watermark -d -v -a '<NAME>' -p scikit-learn,nltk,numpy
# <font size="1.5em">[More information](https://github.com/rasbt/watermark) about the `watermark` magic command extension.</font>
# <br>
# <br>
# # Lyrics Mood Classification - Training
# <br>
# <br>
# ### Sections
# - [Preprocessing](#Preprocessing)
# - [Reading the dataset](#Reading-the-dataset)
# - [Label Encoder](#Label-Encoder)
# - [Feature extraction: Word counts and Vectorizers](#Feature-extraction:-Word-counts-and-Vectorizers)
# - [Porter Stemmer](#Porter-Stemmer)
# - [Stop word list](#Stop-word-list)
# - [Count Vectorizer](#Count-Vectorizer)
# - [Tfidf Vectorizer](#Tfidf-Vectorizer)
# - [Model Selection](#Model-Selection)
# - [Grid search and hyperparameter tuning](#Grid-search-and-hyperparameter-tuning)
# - [Grid Search](#Grid-Search)
# - [Grid Search ROC Curves](#Grid-Search-ROC-Curves)
# - [Hyperparameter tuning 1 - max_features](#Hyperparameter-tuning-1---max_features)
# - [Hyperparameter tuning 2 - min_df](#Hyperparameter-tuning-2---min_df)
# - [Hyperparameter tuning 3 - alpha](#Hyperparameter-tuning-3---alpha)
# - [Validation](#Validation)
# - [Confusion matrix](#Confusion-matrix)
# - [ROC AUC, Accuracy, Precision, Recall, and F1-score](#ROC-AUC,-Accuracy,-Precision,-Recall,-and-F1-score)
# - [Save classifier](#Save-classifier)
# <br>
# <br>
# # Preprocessing
# [[back to top](#Sections)]
# <br>
# <br>
# ### Reading the dataset
# [[back to top](#Sections)]
# +
import pandas as pd
df = pd.read_csv('../../dataset/training/train_lyrics_1000.csv')
df.head()
# -
# <br>
# <br>
# ### Label Encoder
# [[back to top](#Sections)]
# +
from sklearn.preprocessing import LabelEncoder
import pickle
import numpy as np
X_train = df['lyrics'].values
y_train = df['mood'].values
print('before: %s ...' %y_train[:5])
le = LabelEncoder()
le.fit(y_train)
y_train = le.transform(y_train)
print('after: %s ...' %y_train[:5])
# +
# Save object to disk
import pickle
pickle_out = open('./lyrics_label_encoder_py.pkl', 'wb')
pickle.dump(le, pickle_out)
pickle_out.close()
# -
# <br>
# <br>
# ## Feature extraction: Word counts and Vectorizers
# [[back to top](#Sections)]
# <br>
# <br>
# ### Porter Stemmer
# [[back to top](#Sections)]
# +
# Porter Stemmer
import nltk
import string
import re
porter_stemmer = nltk.stem.porter.PorterStemmer()
def porter_tokenizer(text, stemmer=porter_stemmer):
"""
A Porter-Stemmer-Tokenizer hybrid to splits sentences into words (tokens)
and applies the porter stemming algorithm to each of the obtained token.
Tokens that are only consisting of punctuation characters are removed as well.
Only tokens that consist of more than one letter are being kept.
Parameters
----------
text : `str`.
A sentence that is to split into words.
Returns
----------
no_punct : `str`.
A list of tokens after stemming and removing Sentence punctuation patterns.
"""
lower_txt = text.lower()
tokens = nltk.wordpunct_tokenize(lower_txt)
stems = [porter_stemmer.stem(t) for t in tokens]
no_punct = [s for s in stems if re.match('^[a-zA-Z]+$', s) is not None]
return no_punct
# -
# #### Test
porter_tokenizer("Don't !!! --- want swimming. ")
# <br>
# <br>
# ### Stop word list
# [[back to top](#Sections)]
# +
# Commented out to prevent overwriting files:
#
# stp = nltk.corpus.stopwords.words('english')
# with open('./stopwords_eng.txt', 'w') as outfile:
# outfile.write('\n'.join(stp))
with open('./stopwords_eng.txt', 'r') as infile:
stop_words = infile.read().splitlines()
print('stop words %s ...' %stop_words[:5])
# -
# <br>
# <br>
# ### Count Vectorizer
# [[back to top](#Sections)]
# +
# Count Vectorizer
from sklearn.feature_extraction.text import CountVectorizer
vec = CountVectorizer(
encoding='utf-8',
decode_error='replace',
strip_accents='unicode',
analyzer='word',
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1)
)
# -
# #### Test 1
# +
vocab = ["123 1 The\n swimmer likes swimming so he swims. Don't didn`t"]
vec = vec.fit(vocab)
sentence1 = vec.transform([u'The swimmer likes swimming.'])
sentence2 = vec.transform(['The\nswimmer \nswims.'])
print('TEST:')
print('Vocabulary: %s' %vec.get_feature_names())
print('Sentence 1: %s' %sentence1.toarray())
print('Sentence 2: %s' %sentence2.toarray())
# -
# #### Fitting the lyrics
vec = vec.fit(X_train.ravel())
print('Vocabulary size: %s' %len(vec.get_feature_names()))
# #### Test 2 (N-grams = 2)
# +
vec = CountVectorizer(
encoding='utf-8',
decode_error='replace',
strip_accents='unicode',
analyzer='word',
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(2,2)
)
vocab = ["123 1 The\n swimmer likes swimming so he swims. Don't didn`t"]
vec = vec.fit(vocab)
sentence1 = vec.transform([u'The swimmer likes swimming.'])
sentence2 = vec.transform(['The\nswimmer \nswims.'])
print('TEST:')
print('Vocabulary: %s' %vec.get_feature_names())
print('Sentence 1: %s' %sentence1.toarray())
print('Sentence 2: %s' %sentence2.toarray())
# -
# <br>
# <br>
# ### Tfidf Vectorizer
# [[back to top](#Sections)]
# +
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer(
encoding='utf-8',
decode_error='replace',
strip_accents='unicode',
analyzer='word',
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer
)
# -
# #### Test
# +
vocab = ["123 1 The\n swimmer likes swimming so he swims. Don't didn`t"]
tfidf = tfidf.fit(vocab)
sentence1 = tfidf.transform([u'The swimmer likes swimming.'])
sentence2 = tfidf.transform(['The\nswimmer \nswims.'])
print('TEST:')
print('Vocabulary: %s' %tfidf.get_feature_names())
print('Sentence 1: %s' %sentence1.toarray())
print('Sentence 2: %s' %sentence2.toarray())
# -
# #### Fitting the lyrics
# +
tfidf = tfidf.fit(X_train.ravel())
print('Vocabulary size: %s' %len(tfidf.get_feature_names()))
# -
# <br>
# <br>
# # Model Selection
# [[back to top](#Sections)]
# <br>
# <br>
# ### Grid search and hyperparameter tuning
# [[back to top](#Sections)]
# <br>
# <br>
# #### Models: Multivariate Bernoulli and Multinomial naive Bayes
from sklearn.naive_bayes import MultinomialNB
from sklearn.naive_bayes import BernoulliNB
from sklearn.pipeline import Pipeline
# #### Performance metric: F1-score
# +
# Custom scorer methods to account for positive-negative class labels
from sklearn import metrics
# `pos_label` for positive class, since we have sad=1, happy=0
f1_scorer = metrics.make_scorer(metrics.f1_score, greater_is_better=True, pos_label=0)
# -
# <br>
# <br>
# #### Grid Search
# [[back to top](#Sections)]
# +
from sklearn.grid_search import GridSearchCV
from pprint import pprint
pipeline_1 = Pipeline([
('vect', CountVectorizer()),
('clf', BernoulliNB())
])
parameters_1 = dict(
vect__binary=[True],
vect__stop_words=[stop_words, None],
vect__tokenizer=[porter_tokenizer, None],
vect__ngram_range=[(1,1), (2,2), (3,3)],
)
grid_search_1 = GridSearchCV(pipeline_1,
parameters_1,
n_jobs=1,
verbose=1,
scoring=f1_scorer,
cv=10
)
print("Performing grid search...")
print("pipeline:", [name for name, _ in pipeline_1.steps])
print("parameters:")
pprint(parameters_1, depth=2)
grid_search_1.fit(X_train, y_train)
print("Best score: %0.3f" % grid_search_1.best_score_)
print("Best parameters set:")
best_parameters_1 = grid_search_1.best_estimator_.get_params()
for param_name in sorted(parameters_1.keys()):
print("\t%s: %r" % (param_name, best_parameters_1[param_name]))
# +
from sklearn.grid_search import GridSearchCV
pipeline_3 = Pipeline([
('vect', CountVectorizer()),
('clf', MultinomialNB())
])
parameters_3 = dict(
vect__binary=[False],
vect__stop_words=[stop_words, None],
vect__tokenizer=[porter_tokenizer, None],
vect__ngram_range=[(1,1), (2,2), (3,3)],
)
grid_search_3 = GridSearchCV(pipeline_3,
parameters_3,
n_jobs=1,
verbose=1,
scoring=f1_scorer,
cv=10
)
print("Performing grid search...")
print("pipeline:", [name for name, _ in pipeline_3.steps])
print("parameters:")
pprint(parameters_3, depth=2)
grid_search_3.fit(X_train, y_train)
print("Best score: %0.3f" % grid_search_3.best_score_)
print("Best parameters set:")
best_parameters_3 = grid_search_3.best_estimator_.get_params()
for param_name in sorted(parameters_3.keys()):
print("\t%s: %r" % (param_name, best_parameters_3[param_name]))
# +
from sklearn.grid_search import GridSearchCV
pipeline_4 = Pipeline([
('vect', TfidfVectorizer()),
('clf', MultinomialNB())
])
parameters_4 = dict(
vect__binary=[False],
vect__stop_words=[stop_words, None],
vect__tokenizer=[porter_tokenizer, None],
vect__ngram_range=[(1,1), (2,2), (3,3)],
)
grid_search_4 = GridSearchCV(pipeline_4,
parameters_4,
n_jobs=1,
verbose=1,
scoring=f1_scorer,
cv=10
)
print("Performing grid search...")
print("pipeline:", [name for name, _ in pipeline_4.steps])
print("parameters:")
pprint(parameters_4, depth=2)
grid_search_4.fit(X_train, y_train)
print("Best score: %0.3f" % grid_search_4.best_score_)
print("Best parameters set:")
best_parameters_4 = grid_search_4.best_estimator_.get_params()
for param_name in sorted(parameters_4.keys()):
print("\t%s: %r" % (param_name, best_parameters_4[param_name]))
# -
# <br>
# <br>
# #### Grid Search ROC Curves
# [[back to top](#Sections)]
# +
from sklearn.metrics import roc_curve, auc
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.cross_validation import StratifiedKFold
from scipy import interp
sns.set()
sns.set_style("whitegrid")
clf_1 = Pipeline([
('vect', CountVectorizer(
binary=True,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1),
)
),
('clf', BernoulliNB()),
])
clf_2 = Pipeline([
('vect', CountVectorizer(
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1),
)
),
('clf', MultinomialNB()),
])
clf_3 = Pipeline([
('vect', TfidfVectorizer(
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1),
)
),
('clf', MultinomialNB()),
])
colors = ['#1947D1', '#CC3300', 'k']
linestyles = ['-', '--', '-.']
classifiers = [clf_1, clf_2, clf_3]
labels = ['1: MV Bernoulli NB, stop words, porter stemmer, \nuni-gram, df',
'2: Multinomial NB, stop words, porter stemmer, \nuni-gram, tf',
'3: Multinomial NB, stop words, porter stemmer, \nuni-gram, tf-idf',
]
for clf,col,ls,lab in zip(classifiers, colors, linestyles, labels):
mean_tpr = 0.0
mean_fpr = np.linspace(0, 1, 100)
all_tpr = []
cv = StratifiedKFold(y_train, n_folds=10, random_state=123)
for i, (train, test) in enumerate(cv):
probas_ = clf.fit(X_train[train], y_train[train]).predict_proba(X_train[test])
# Compute ROC curve and area the curve
fpr, tpr, thresholds = roc_curve(y_train[test], probas_[:, 1])
mean_tpr += interp(mean_fpr, fpr, tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr, tpr)
mean_tpr /= len(cv)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
plt.plot(mean_fpr,
mean_tpr,
color=col,
linestyle=ls,
label='%s (ROC AUC = %0.2f)' % (lab, mean_auc),
lw=2
)
plt.plot([0, 1], [0, 1], '--', color=(0.6, 0.6, 0.6), label='Random Guessing')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc="lower right")
plt.savefig('./images/roc_gridsearch_1.eps', dpi=300)
plt.show()
# -
# <br>
# <br>
# #### Hyperparameter tuning 1 - max_features
# [[back to top](#Sections)]
# +
from sklearn.metrics import roc_curve, auc
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.cross_validation import StratifiedKFold
from scipy import interp
sns.set()
sns.set_style("whitegrid")
colors = ['#1947D1', '#CC3300', 'k', '#339933']
linestyles = ['-', '--', '-.', ':']
params = [1000,3000,5000,None]
labels = ['max features = 1000',
'max features = 3000',
'max features = 5000',
'max features = all (=8550)',
]
for param,col,ls,lab in zip(params, colors, linestyles, labels):
clf = Pipeline([
('vect', TfidfVectorizer(
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1),
max_features=param,
)
),
('clf', MultinomialNB()),
])
mean_tpr = 0.0
mean_fpr = np.linspace(0, 1, 100)
all_tpr = []
cv = StratifiedKFold(y_train, n_folds=10, random_state=123)
for i, (train, test) in enumerate(cv):
probas_ = clf.fit(X_train[train], y_train[train]).predict_proba(X_train[test])
# Compute ROC curve and area the curve
fpr, tpr, thresholds = roc_curve(y_train[test], probas_[:, 1])
mean_tpr += interp(mean_fpr, fpr, tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr, tpr)
mean_tpr /= len(cv)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
plt.plot(mean_fpr,
mean_tpr,
color=col,
linestyle=ls,
label='%s (ROC AUC = %0.2f)' % (lab, mean_auc),
lw=2
)
plt.plot([0, 1], [0, 1], '--', color=(0.6, 0.6, 0.6), label='Random Guessing')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Multinomial NB, stop words, porter stemmer, uni-gram, tf-idf')
plt.legend(loc="lower right")
plt.savefig('./images/roc_maxfeatures.eps', dpi=300)
plt.show()
# -
# <br>
# <br>
# #### Hyperparameter tuning 2 - min_df
# [[back to top](#Sections)]
# +
from sklearn.metrics import roc_curve, auc
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.cross_validation import StratifiedKFold
from scipy import interp
sns.set()
sns.set_style("whitegrid")
colors = ['#1947D1', '#CC3300', 'k', ]
linestyles = ['-', '--', '-.']
params = [1, 0.1, 0.01]
labels = ['no cutoff',
'min. df = 0.1',
'min. df = 0.01',
]
for param,col,ls,lab in zip(params, colors, linestyles, labels):
clf = Pipeline([
('vect', TfidfVectorizer(
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1),
min_df=param,
)
),
('clf', MultinomialNB()),
])
mean_tpr = 0.0
mean_fpr = np.linspace(0, 1, 100)
all_tpr = []
cv = StratifiedKFold(y_train, n_folds=10, random_state=123)
for i, (train, test) in enumerate(cv):
probas_ = clf.fit(X_train[train], y_train[train]).predict_proba(X_train[test])
# Compute ROC curve and area the curve
fpr, tpr, thresholds = roc_curve(y_train[test], probas_[:, 1])
mean_tpr += interp(mean_fpr, fpr, tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr, tpr)
mean_tpr /= len(cv)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
plt.plot(mean_fpr,
mean_tpr,
color=col,
linestyle=ls,
label='%s (ROC AUC = %0.2f)' % (lab, mean_auc),
lw=2
)
plt.plot([0, 1], [0, 1], '--', color=(0.6, 0.6, 0.6), label='Random Guessing')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc="lower right")
plt.title('Multinomial NB, stop words, porter stemmer, uni-gram, tf-idf')
plt.savefig('./images/roc_mindf.eps', dpi=300)
plt.show()
# -
# <br>
# <br>
# #### Hyperparameter tuning 3 - alpha
# [[back to top](#Sections)]
# +
from sklearn.metrics import roc_curve, auc
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.cross_validation import StratifiedKFold
from scipy import interp
sns.set()
sns.set_style("whitegrid")
colors = ['#1947D1', '#CC3300', 'k', '#339933']
linestyles = ['-', '--', '-.', ':']
params = [0.05, 0.1, 1.0, 2.0]
labels = ['alpha = 0.05',
'alpha = 0.1',
'alpha = 1.0',
'alpha = 2.0',
]
for param,col,ls,lab in zip(params, colors, linestyles, labels):
clf = Pipeline([
('vect', TfidfVectorizer(
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1),
)
),
('clf', MultinomialNB(alpha=param)),
])
mean_tpr = 0.0
mean_fpr = np.linspace(0, 1, 100)
all_tpr = []
cv = StratifiedKFold(y_train, n_folds=10, random_state=123)
for i, (train, test) in enumerate(cv):
probas_ = clf.fit(X_train[train], y_train[train]).predict_proba(X_train[test])
# Compute ROC curve and area the curve
fpr, tpr, thresholds = roc_curve(y_train[test], probas_[:, 1])
mean_tpr += interp(mean_fpr, fpr, tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr, tpr)
mean_tpr /= len(cv)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
plt.plot(mean_fpr,
mean_tpr,
color=col,
linestyle=ls,
label='%s (ROC AUC = %0.2f)' % (lab, mean_auc),
lw=2
)
plt.plot([0, 1], [0, 1], '--', color=(0.6, 0.6, 0.6), label='Random Guessing')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc="lower right")
plt.title('Multinomial NB, stop words, porter stemmer, uni-gram, tf-idf')
plt.savefig('./images/roc_alpha.eps', dpi=300)
plt.show()
# -
# <br>
# <br>
# #### n-gram comparison
# [[back to top](#Sections)]
# +
from sklearn.metrics import roc_curve, auc
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.cross_validation import StratifiedKFold
from scipy import interp
sns.set()
sns.set_style("whitegrid")
colors = ['#1947D1', '#CC3300', 'k', ]
linestyles = ['-', '--', '-.',]
params = [(1,1), (2,2), (3,3),]
labels = ['1-gram',
'2-gram',
'3-gram',
]
for param,col,ls,lab in zip(params, colors, linestyles, labels):
clf = Pipeline([
('vect', TfidfVectorizer(
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=param,
)
),
('clf', MultinomialNB(alpha=1.0)),
])
mean_tpr = 0.0
mean_fpr = np.linspace(0, 1, 100)
all_tpr = []
cv = StratifiedKFold(y_train, n_folds=10, random_state=123)
for i, (train, test) in enumerate(cv):
probas_ = clf.fit(X_train[train], y_train[train]).predict_proba(X_train[test])
# Compute ROC curve and area the curve
fpr, tpr, thresholds = roc_curve(y_train[test], probas_[:, 1])
mean_tpr += interp(mean_fpr, fpr, tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr, tpr)
mean_tpr /= len(cv)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
plt.plot(mean_fpr,
mean_tpr,
color=col,
linestyle=ls,
label='%s (ROC AUC = %0.2f)' % (lab, mean_auc),
lw=2
)
plt.plot([0, 1], [0, 1], '--', color=(0.6, 0.6, 0.6), label='Random Guessing')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend(loc="lower right")
plt.title('Multinomial NB, stop words, porter stemmer, uni-gram, tf-idf')
plt.savefig('./images/roc_ngrams.eps', dpi=300)
plt.show()
# -
# <br>
# <br>
# +
from sklearn.metrics import roc_curve, auc
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.cross_validation import StratifiedKFold
from scipy import interp
sns.set()
sns.set_style("whitegrid")
classifier = Pipeline([
('vect', TfidfVectorizer(
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1),
)
),
('clf', MultinomialNB()),
])
cv = StratifiedKFold(y_train, n_folds=10, random_state=123)
mean_tpr = 0.0
mean_fpr = np.linspace(0, 1, 100)
all_tpr = []
for i, (train, test) in enumerate(cv):
probas_ = classifier.fit(X_train[train], y_train[train]).predict_proba(X_train[test])
# Compute ROC curve and area the curve
fpr, tpr, thresholds = roc_curve(y_train[test], probas_[:, 1])
mean_tpr += interp(mean_fpr, fpr, tpr)
mean_tpr[0] = 0.0
roc_auc = auc(fpr, tpr)
plt.plot(fpr, tpr, lw=1, label='ROC fold %d (area = %0.2f)' % (i+1, roc_auc))
plt.plot([0, 1], [0, 1], '--', color=(0.6, 0.6, 0.6), label='Random Guessing')
mean_tpr /= len(cv)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
plt.plot(mean_fpr, mean_tpr, 'k--',
label='Mean ROC (area = %0.2f)' % mean_auc, lw=2)
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver Operator Characteristic of the Lyrics Classifier')
plt.legend(loc="lower right")
plt.savefig('./images/roc_final.eps', dpi=300)
plt.show()
# -
# <br>
# <br>
# # Validation
# [[back to top](#Sections)]
final_clf = Pipeline([
('vect', TfidfVectorizer(
binary=False,
stop_words=stop_words,
tokenizer=porter_tokenizer,
ngram_range=(1,1),
)
),
('clf', MultinomialNB(alpha=1.0)),
])
final_clf.fit(X_train, y_train)
# <br>
# <br>
# ### Confusion matrix
# [[back to top](#Sections)]
# +
import matplotlib as mpl
import numpy as np
cm = metrics.confusion_matrix(y_train, final_clf.predict(X_train))
np.set_printoptions(suppress=True)
mpl.rc("figure", figsize=(4, 2))
hm = sns.heatmap(cm,
cbar=False,
annot=True,
square=True,
fmt='d',
yticklabels=['happy','sad'],
xticklabels=['happy','sad'],
cmap='Blues'
)
plt.title('Confusion matrix - Training dataset')
plt.ylabel('actual class')
plt.xlabel('predicted class')
plt.tight_layout()
plt.savefig('./images/confmat_training.eps', dpi=300)
plt.show()
# +
df = pd.read_csv('../../dataset/validation/valid_lyrics_200.csv')
X_valid = df['lyrics'].values
y_valid = df['mood'].values
y_valid = le.transform(y_valid)
# +
cm = metrics.confusion_matrix(y_valid, final_clf.predict(X_valid))
np.set_printoptions(suppress=True)
mpl.rc("figure", figsize=(4, 2))
hm = sns.heatmap(cm,
cbar=False,
annot=True,
square=True,
fmt='d',
yticklabels=['happy','sad'],
xticklabels=['happy','sad'],
cmap='Blues'
)
plt.title('Confusion matrix - Validation dataset')
plt.ylabel('actual class')
plt.xlabel('predicted class')
plt.tight_layout()
plt.savefig('./images/confmat_valid.eps', dpi=300)
plt.show()
# -
# <br>
# <br>
# ### ROC AUC, Accuracy, Precision, Recall, and F1-score
# [[back to top](#Sections)]
# +
# Custom scorer methods to account for positive-negative class labels
from sklearn import metrics
# `pos_label` for positive class, since we have sad=1, happy=0
acc_scorer = metrics.make_scorer(metrics.accuracy_score, greater_is_better=True)
pre_scorer = metrics.make_scorer(metrics.precision_score, greater_is_better=True, pos_label=0)
rec_scorer = metrics.make_scorer(metrics.recall_score, greater_is_better=True, pos_label=0)
f1_scorer = metrics.make_scorer(metrics.f1_score, greater_is_better=True, pos_label=0)
auc_scorer = metrics.make_scorer(metrics.roc_auc_score, greater_is_better=True)
# -
d = {'Data':['Training', 'Validation'],
'ACC (%)':[],
'PRE (%)':[],
'REC (%)':[],
'F1 (%)':[],
'ROC AUC (%)':[],
}
# +
d['ACC (%)'].append(acc_scorer(estimator=final_clf, X=X_train, y_true=y_train))
d['PRE (%)'].append(pre_scorer(estimator=final_clf, X=X_train, y_true=y_train))
d['REC (%)'].append(rec_scorer(estimator=final_clf, X=X_train, y_true=y_train))
d['F1 (%)'].append(f1_scorer(estimator=final_clf, X=X_train, y_true=y_train))
d['ROC AUC (%)'].append(auc_scorer(estimator=final_clf, X=X_train, y_true=y_train))
d['ACC (%)'].append(acc_scorer(estimator=final_clf, X=X_valid, y_true=y_valid))
d['PRE (%)'].append(pre_scorer(estimator=final_clf, X=X_valid, y_true=y_valid))
d['REC (%)'].append(rec_scorer(estimator=final_clf, X=X_valid, y_true=y_valid))
d['F1 (%)'].append(f1_scorer(estimator=final_clf, X=X_valid, y_true=y_valid))
d['ROC AUC (%)'].append(auc_scorer(estimator=final_clf, X=X_valid, y_true=y_valid))
# -
df_perform = pd.DataFrame(d)
df_perform = df_perform[['ACC (%)', 'PRE (%)', 'REC (%)', 'F1 (%)', 'ROC AUC (%)']]
df_perform.index=(['Training', 'Validation'])
df_perform = df_perform*100
df_perform = np.round(df_perform, decimals=2)
df_perform
df_perform.to_csv('./clf_performance.csv', index_label=False)
# <br>
# <br>
# ## Save classifier
# [[back to top](#Sections)]
# +
lyrics_clf_1000 = final_clf
pickle_out = open('./lyrics_clf_1000_py27.pkl', 'wb')
pickle.dump(lyrics_clf_1000, pickle_out)
pickle_out.close()
# -
# # New pickle objects for webapp
# +
import pickle
pickle_out = open('./lyrics_label_encoder.pkl', 'rb')
le = pickle.load(pickle_out)
pickle_out.close()
# +
from sklearn.naive_bayes import MultinomialNB
with open('./stopwords_eng.txt', 'r') as infile:
stop_words = infile.read().splitlines()
# +
# Porter Stemmer
import nltk
import string
import re
porter_stemmer = nltk.stem.porter.PorterStemmer()
def porter_tokenizer(text, stemmer=porter_stemmer):
"""
A Porter-Stemmer-Tokenizer hybrid to splits sentences into words (tokens)
and applies the porter stemming algorithm to each of the obtained token.
Tokens that are only consisting of punctuation characters are removed as well.
Only tokens that consist of more than one letter are being kept.
Parameters
----------
text : `str`.
A sentence that is to split into words.
Returns
----------
no_punct : `str`.
A list of tokens after stemming and removing Sentence punctuation patterns.
"""
lower_txt = text.lower()
tokens = nltk.wordpunct_tokenize(lower_txt)
stems = [porter_stemmer.stem(t) for t in tokens]
no_punct = [s for s in stems if re.match('^[a-zA-Z]+$', s) is not None]
return no_punct
# +
import pandas as pd
df = pd.read_csv('../../dataset/training/train_lyrics_1000.csv')
X_train = df['lyrics'].values
y_train = df['mood'].values
# +
from sklearn.preprocessing import LabelEncoder
from sklearn.pipeline import Pipeline
le = LabelEncoder()
le.fit(y_train)
y_train = le.transform(y_train)
# +
from sklearn.feature_extraction.text import TfidfVectorizer
final_clf = Pipeline([
('vect', TfidfVectorizer(
binary=False,
stop_words=stop_words,
ngram_range=(1,1),
)
),
('clf', MultinomialNB(alpha=1.0)),
])
final_clf.fit(X_train, y_train)
# +
from sklearn.externals import joblib
from sklearn.pipeline import Pipeline
import pickle
pickle_out = open('./lyrics_label_encoder_np.pkl', 'wb')
pickle.dump(le, pickle_out)
pickle_out.close()
joblib.dump(final_clf, 'lyrics_clf_1000_np.pkl')
# +
from sklearn.externals import joblib
final_clf = joblib.load('lyrics_clf_1000_jb.pkl')
# -
final_clf.predict(X_train)[:3]
joblib.dump(le, 'lyrics_label_encoder_jb.pkl')
from sklearn.externals import joblib
lyrics_label_encoder = joblib.load('lyrics_label_encoder_jb.pkl')
| .ipynb_checkpoints/nb_init_model-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="BdP8BRszLRvq" outputId="b4776e1f-fc07-4921-ff5a-46d4d683c830"
import nltk
from sklearn.model_selection import train_test_split
import torch
from xtagger import xtagger_dataset_to_df
from xtagger import df_to_torchtext_data
from transformers import AutoTokenizer
nltk_data = list(nltk.corpus.treebank.tagged_sents(tagset='universal'))
train_set,test_set =train_test_split(nltk_data,train_size=0.8,test_size=0.2,random_state = 2112)
df_train = xtagger_dataset_to_df(train_set)
df_test = xtagger_dataset_to_df(test_set)
device = torch.device("cuda")
tokenizer = AutoTokenizer.from_pretrained("bert-base-cased")
train_iterator, valid_iterator, test_iterator, TEXT, TAGS = df_to_torchtext_data(
df_train,
df_test,
device,
transformers = True,
tokenizer = tokenizer,
batch_size=32
)
# + colab={"base_uri": "https://localhost:8080/"} id="orAhtoENM0N3" outputId="6bedad4c-94d6-4e6f-f854-aeed30659aab"
from xtagger import BERTForTagging
model = BERTForTagging(
output_dim = 13,
TEXT = TEXT,
TAGS = TAGS,
dropout = 0.2,
device = device,
cuda = True
)
# + colab={"base_uri": "https://localhost:8080/", "height": 749, "referenced_widgets": ["798e515b1b3a4d6da750ddaf656dcf87", "169e607e47d64258abe6bec7bae40c15", "37f2d6ce6840446e86fcf1a7cdd60256", "<KEY>", "3684d5174a6a47b39031cad53afad33d", "<KEY>", "7a987bfefe664116936e9dae898a34e0", "2cb40d4558904b8e81155a420321188c", "142d3f938d1e40f4b4d3f4394637e7f6", "fe93ecdb9977410aaa17788d68ceea99", "c26be8088c8b4d219126d76732a55de0", "304c3fc253744f2fb24a528e10a40168", "22af1e6f785c47d4b5437c054e53fdde", "<KEY>", "bdf2d144ff4f431f810c89df751fa92e", "<KEY>", "1e0f8067ea7c42ea9a0d2ccc4b29879c", "<KEY>", "<KEY>", "45e2c0dbeffa4a4f9e9543142d308a58", "308ad9fb706643978fe3da26009bd6a3", "<KEY>", "3499b4a1cbd04f569a8fbe4d7f0be384", "ac42c1ce963447979066e0c58c451cef", "<KEY>", "<KEY>", "2da5c0a61bb94798963d517e8f3c812d", "e73acbe542ed4eebb802377d162f5428", "4385e234504c48be9bc01c6e16af3c02", "2993dc7cf4e94825a9d177d7094f33aa", "c708d18b6d5a46918cf824406be5e21d", "3480d66eb85d4389a86e9a12d6c81bd1", "<KEY>", "6cc662eef23c4e99899d8fc0b67dbd10", "bd47e686385f4e9ea5b604f9f276864b", "<KEY>", "<KEY>", "bf922c8d0d434b77a2e2370ff2ffec3e", "<KEY>", "66287c0d9a0f4797814473d07a73df76", "f08f40444f5743fe825f8475f554f19d", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "1d10a30db91644e5af4a41cf5719e1ce", "21e3a711539246fdb8e5ff9562a6d29c", "<KEY>", "7334f6ab6d45434abd984e8b05220b10", "<KEY>", "<KEY>", "fb933df2616f4216a7595b9e4004b998", "<KEY>", "<KEY>", "54e8eb48b3f14159a5da1175dac31e7c", "<KEY>", "233433da05174d19b41e2ceb91870154", "<KEY>", "701145ceb48044c98e153e2f48b88337", "<KEY>", "<KEY>", "<KEY>", "d6cc607538ce4e07892e4e24a5875280", "<KEY>", "11c211036a604b8399028b4555be408d", "8e111ff9843146a085a60c5a68d676e4", "297a458418c84621b171b0a88b1e508e", "<KEY>", "<KEY>", "748d1ff67b8048dbae17616d6e061e92", "3936451a655f458da532da97b98c9e0a", "cad3625430cd4e0e8fd68021bdda14b5", "<KEY>", "5392eec0c19545209d6ab1be031fa49b", "<KEY>", "<KEY>", "<KEY>", "e38dc7539c314defbe308908ace15873", "<KEY>", "b2cee6e8699746e4a62299c3ebbd5f65", "<KEY>", "<KEY>", "<KEY>", "60571183ece64c1e938264d7ab0f8714", "<KEY>", "d139ac2fc1e94c3483ef38e84baa7a7a", "<KEY>", "<KEY>", "1d3f7796aeab4aab9d5fd5697b14a92d", "<KEY>", "<KEY>", "beefe5f07bfa41fda56a12f3906105e0", "6dabfd6e6532440585195202d854fc64", "ed13a5dfba0147c682a9d7e1253b4a89", "<KEY>", "<KEY>", "96fbafb1ab914389a5ab32e88ff4ed4d", "e30cc4e38827479face901cd41f07d96", "0a507ceb7fdf4fa3904fede7d0a0323a", "<KEY>", "<KEY>", "d9a2428eacfb42be81a7d78e86bcd00e", "<KEY>", "487744df4b7348c2aca465e906a57355", "8faf99a700ad4de09cc5bb91248b3839", "e9e154151639453f905c49ea5761f3b1", "ad021a62fadf4ea2a3724a56ec4dd545", "<KEY>", "dc9c2d8a3ff749e1a19325797f25b281", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "94de78f8e2aa4da28571def99284d222", "fd4c2d1d84864455a0ade9837cf13d9d", "<KEY>", "ded0d0732d0744129fe58f4f40297d48", "3178d40e3a9a49f7be519f7977fa6473", "ae0aca3199314cbeaa0ff473ccea2376", "29603d2b4b1e4ef28ced0977f0f166d2"]} id="SoycocBNNEP7" outputId="6399581b-916c-4952-abad-fcdc6d743337"
model.fit(
train_iterator,
valid_iterator,
eval_metrics = ["acc", "avg_f1"],
epochs = 10
)
# + colab={"base_uri": "https://localhost:8080/", "height": 121, "referenced_widgets": ["9671ed62e6a54dc6a4f9f56c11d60170", "d26ea2f8974d466a92d25c777dd9b48e", "fc278189bb18476ea2de288137755f29", "a381fd48ccd04c0d92cffff0553aa86c", "3cf46f5adab54501a112c7667b48eddd", "e6c9e178a32a4ec98ff0d4dc3a0ceaa5", "ed8ca5114e9d46e4aaba9070acf01e73", "0515428d455b4a51a82b6769b4951240", "<KEY>", "11dcc0b4dd6b4133af8ea392586b4e3d", "13556840c1824e3c883bb97f4a659a4b"]} id="74eSO0x4NMQ2" outputId="8d15d303-d425-4701-b1e3-e92afee780b4"
model.evaluate(test_iterator, eval_metrics = ["acc", "avg_f1"])
# + colab={"base_uri": "https://localhost:8080/"} id="1ihW749vN4gR" outputId="6ec7f58a-878b-4e26-99b0-2eeb0ce9ea5f"
s = ["there", "are", "no", "two", "words", "in", "the", "english",
"language", "more", "harmful", "than", "good", "job"]
model.predict(s, tokenizer)
| examples/Sequence Labeling: Pretrained BERT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating a GRU model using Trax: Ungraded Lecture Notebook
# For this lecture notebook you will be using Trax's layers. These are the building blocks for creating neural networks with Trax.
import trax
from trax import layers as tl
# Trax allows to define neural network architectures by stacking layers (similarly to other libraries such as Keras). For this the `Serial()` is often used as it is a combinator that allows to stack layers serially using function composition.
#
# Next you can see a simple vanilla NN architecture containing 1 hidden(dense) layer with 128 cells and output (dense) layer with 10 cells on which we apply the final layer of logsoftmax.
mlp = tl.Serial(
tl.Dense(128),
tl.Relu(),
tl.Dense(10),
tl.LogSoftmax()
)
# Each of the layers within the `Serial` combinator layer is considered a sublayer. Notice that unlike similar libraries, **in Trax the activation functions are considered layers.** To know more about the `Serial` layer check the docs [here](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Serial).
#
# You can try printing this object:
print(mlp)
# Printing the model gives you the exact same information as the model's definition itself.
#
# By just looking at the definition you can clearly see what is going on inside the neural network. Trax is very straightforward in the way a network is defined, that is one of the things that makes it awesome!
# ## GRU MODEL
# To create a `GRU` model you will need to be familiar with the following layers (Documentation link attached with each layer name):
# - [`ShiftRight`](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.attention.ShiftRight) Shifts the tensor to the right by padding with zeros on axis 1. The `mode` should be specified and it refers to the context in which the model is being used. Possible values are: 'train', 'eval' or 'predict', predict mode is for fast inference. Defaults to "train". With this layer, the input sequence is shifted to the right so, at every time step, the GRU cell doesn't get as input the same element that needs to be predicted. Note that this layer isn't always neccessary, its inclusion depends on the NLP task, but for this week's assignment you'll need to use it.
# - [`Embedding`](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.Embedding) Maps discrete tokens to vectors. It will have shape `(vocabulary length X dimension of output vectors)`. The dimension of output vectors (also called `d_feature`) is the number of elements in the word embedding.
# - [`GRU`](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.rnn.GRU) The GRU layer. It leverages another Trax layer called [`GRUCell`](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.rnn.GRUCell). The hidden state dimension should be specified as `n_units` and should match the number of elements in the word embedding --by design in Trax. If you want to stack two consecutive GRU layers, it can be done by using python's list comprehension.
# - [`Dense`](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.Dense) Vanilla Dense layer.
# - [`LogSoftMax`](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.LogSoftmax) Log Softmax function.
#
# Putting everything together the GRU model will look like this:
# +
mode = 'train'
vocab_size = 256
model_dimension = 512
n_layers = 2
GRU = tl.Serial(
tl.ShiftRight(mode=mode), # Do remember to pass the mode parameter if you are using it for interence/test as default is train
tl.Embedding(vocab_size=vocab_size, d_feature=model_dimension),
[tl.GRU(n_units=model_dimension) for _ in range(n_layers)], # You can play around n_layers if you want to stack more GRU layers together
tl.Dense(n_units=vocab_size),
tl.LogSoftmax()
)
# -
# Next is a helper function that prints information for every layer (sublayer within `Serial`):
#
# _Try changing the parameters defined before the GRU model and see how it changes!_
#
# +
def show_layers(model, layer_prefix="Serial.sublayers"):
print(f"Total layers: {len(model.sublayers)}\n")
for i in range(len(model.sublayers)):
print('========')
print(f'{layer_prefix}_{i}: {model.sublayers[i]}\n')
show_layers(GRU)
# -
# Hope you are now more familiarized with creating GRU models using Trax.
#
# You will train this model in this week's assignment and see it in action.
#
#
# **GRU and the trax minions will return, in this week's endgame.**
| Part3_Sequence_Models/C3_W2_lecture_nb_4_GRU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yashpatel5400/crypto-prediction/blob/main/stats_601_project.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="mNp93N3fBzaM"
# ## Setup
# + id="ccADqYamxwt0" colab={"base_uri": "https://localhost:8080/"} outputId="fdfa40fb-6ffb-46d1-d921-b25927436962"
import os
import datetime
import time
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import lightgbm as lgb
from sklearn.linear_model import RidgeCV
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import RidgeClassifierCV
from sklearn.linear_model import HuberRegressor
from sklearn.linear_model import ElasticNet
from sklearn.tree import DecisionTreeRegressor
from sklearn.preprocessing import PolynomialFeatures
from sklearn.ensemble import AdaBoostRegressor
from sklearn.linear_model import BayesianRidge
from sklearn.linear_model import ARDRegression
import urllib.request
from joblib import dump, load
url = "https://media.githubusercontent.com/media/yashpatel5400/crypto-prediction/main/log_pr.csv"
urllib.request.urlretrieve(url, "log_pr.csv")
url = "https://media.githubusercontent.com/media/yashpatel5400/crypto-prediction/main/volu.csv"
urllib.request.urlretrieve(url, "volu.csv")
# + id="GzB9_4WAMHo2" colab={"base_uri": "https://localhost:8080/"} outputId="2e527214-74c9-41c3-a140-82e685277cf3"
log_pr = pd.read_csv("log_pr.csv", index_col= 0)
volu = pd.read_csv("volu.csv", index_col= 0)
print(len(log_pr))
log_pr.index = pd.to_datetime(log_pr.index)
volu.index = pd.to_datetime(volu.index)
in_sample_size = 50000
out_sample_size = 25000
set_size = in_sample_size + out_sample_size
start_index = 0
log_pr_kfolds = []
volu_kfolds = []
while start_index + set_size <= len(log_pr):
end_index = start_index + set_size
log_prfold = log_pr.iloc[start_index:end_index, :]
volufold = volu.iloc[start_index:end_index, :]
log_pr_kfolds.append((log_prfold.iloc[:-out_sample_size, :], log_prfold.iloc[-out_sample_size:, :])) #(train, test)
volu_kfolds.append((volufold.iloc[:-out_sample_size, :], volufold.iloc[-out_sample_size:, :]))
start_index += out_sample_size
print(len(log_pr_kfolds))
train_size = 44160
out_sample_size = 44160
log_pr_anchoredfolds = []
volu_anchoredfolds = []
while train_size + out_sample_size <= len(log_pr):
log_pr_anchoredfolds.append((log_pr.iloc[0:train_size, :] , log_pr.iloc[train_size:train_size+out_sample_size, :])) #(train, test)
print(len(log_pr.iloc[0:train_size, :]), len(log_pr.iloc[train_size:train_size+out_sample_size, :]))
volu_anchoredfolds.append((volu.iloc[0:train_size, :] , volu.iloc[train_size:train_size+out_sample_size, :])) #(train, test)
train_size += out_sample_size
print(len(log_pr_anchoredfolds))
# + id="Ps0EegrBDUHr"
vol_train_mean = volu.mean()
vol_train_sd = volu.std()
np.save("vol_train_mean.npy", vol_train_mean)
np.save("vol_train_sd.npy", vol_train_sd)
vol_train_mean = np.load("vol_train_mean.npy")
vol_train_sd = np.load("vol_train_sd.npy")
# + [markdown] id="jDjRe7xoJdZc"
# # Construct features and dataset
# + id="eRn3ihxa2ENe"
def construct_features(log_pr_df, vol_df):
df = log_pr_df.copy()
vol_standardized = (vol_df - vol_train_mean)/(vol_train_sd)
df = pd.concat([df, vol_standardized], axis=1)
return df
# + id="yrHfzn9Zxwt2"
def construct_dataset(window_size, features, log_prices, classification= False):
"""
window: look-back window size for constructing X (in minutes)
"""
window_dt = datetime.timedelta(minutes=window_size)
predict_dt = datetime.timedelta(minutes=30)
window_X = []
window_y = []
for t in features.index[window_size:-30:10]: # compute the predictions every 10 minutes
window_X.append(features.loc[(t - window_dt):t])
if classification:
window_y.append(np.sign(log_prices.loc[t + predict_dt] - log_prices.loc[t])) #changed to classification
else:
window_y.append(log_prices.loc[t + predict_dt] - log_prices.loc[t])
return np.array(window_X), np.array(window_y)
# + id="ohmDUTY2S7cr"
def split_dataset_by_asset(dataset_train):
NUM_ASSETS = 10
return [dataset_train[..., asset::NUM_ASSETS] for asset in range(NUM_ASSETS)]
# + id="XDKaL6zYCYqg"
def construct_local_features(asset_window, asset_idx):
"""
construct features for *single windowed asset*
NOTE: for adding new features, see asset_window[...,0] for np.std
Here, 0 is the feature that is being pulled out (corresponds to the index from global_feature construction)
from which you can do whatever transforms you want
"""
if 0 <= asset_idx and asset_idx <= 9:
poly = PolynomialFeatures(4, include_bias= False)
neg_back_30_for = -(asset_window[...,0][:, -1] - asset_window[...,0][:, -30]).reshape(-1, 1)
median_vol = volt_pr = np.median(asset_window[...,1], axis=-1, keepdims=True)
volt_vol = np.std(asset_window[...,1], axis=-1, keepdims=True)
volt_pr = np.std(asset_window[...,0], axis=-1, keepdims=True)
poly_pr = poly.fit_transform(neg_back_30_for)
test = [poly_pr, volt_pr, median_vol, volt_vol]
test = np.hstack(test)
return test
# + id="RD0WrSmOVznw"
def train_models_split(dataset_train_by_asset, y_train, train_idxs, model_types):
NUM_ASSETS = 10
return [(model_types[asset].fit(construct_local_features(dataset_train_by_asset[asset], asset),
y_train[:, asset]), asset)
for asset in range(NUM_ASSETS) if asset in train_idxs]
# + id="YKsnMlpodBgc"
def get_r_hat_clean(A,B):
window_size = 31
input_features = (construct_features(A, B)).iloc[-window_size:] # only retain tail window
predictions = -(input_features.iloc[-1, :10] - input_features.iloc[-30,:10]).values # init baseline
input_features = input_features.values
split_features_by_asset = split_dataset_by_asset(input_features)
for model, asset_idx in split_models:
# print(asset_idx)
# if model is not None:
# expand_dims is used to align dimensions from "batching" used in training
predictions[asset_idx] = model.predict(construct_local_features(np.expand_dims(split_features_by_asset[asset_idx], axis=0), asset_idx))
return predictions
# + id="8Qu_XQxiGoz8"
models = []
#alphas=[1e-10,1e-8, 1e-6, 1e-4, 1e-2,1e-1,1,1e1,1e2]
for _ in range(10):
models.append(BayesianRidge(n_iter = 1000))
# models[2] = RidgeCV(alphas=[1e-10,1e-8, 1e-6, 1e-4, 1e-2,1e-1,1,1e1,1e2])
# models[5] = RidgeCV(alphas=[1e-10,1e-8, 1e-6, 1e-4, 1e-2,1e-1,1,1e1,1e2])
# models[7] = RidgeCV(alphas=[1e-10,1e-8, 1e-6, 1e-4, 1e-2,1e-1,1,1e1,1e2])
# models[6] = RidgeCV(alphas=[1e-10,1e-8, 1e-6, 1e-4, 1e-2,1e-1,1,1e1,1e2])
# models[9] = RidgeCV(alphas=[1e-10,1e-8, 1e-6, 1e-4, 1e-2,1e-1,1,1e1,1e2])
# + colab={"base_uri": "https://localhost:8080/"} id="UdOLoczdlLOJ" outputId="b11a0b38-8937-45c5-c484-aab19f633658"
tracker = []
split_models_tracker = []
for k in range(len(log_pr_anchoredfolds)):
print(f"Performing {k} fold...")
log_pr_train, log_pr_test = log_pr_anchoredfolds[k]
volu_train, volu_test = volu_anchoredfolds[k]
features = construct_features(log_pr_train, volu_train)
print(log_pr_train.shape)
print(log_pr_test.shape)
window_size = 30 # in minutes
X_train, y_train = construct_dataset(window_size, features, log_pr_train, classification= False)
X_train_by_asset = split_dataset_by_asset(X_train)
# 0, 3, 5 this gave 0.022
print(X_train.shape)
# print(X_train_by_asset.shape)
split_models = train_models_split(X_train_by_asset, y_train, [0, 1, 2, 3, 4, 5, 6, 7, 8, 9], models)
split_models_tracker.append(split_models)
def get_model_corr(test_log_pr, test_volu):
w = 30
t0 = time.time()
dt = datetime.timedelta(days=1)
r_hat = pd.DataFrame(index=test_log_pr.index[w::10], columns=np.arange(10), dtype=np.float64)
print(r_hat.shape)
for t in test_log_pr.index[w::10]: # compute the predictions every 10 minutes
r_hat.loc[t, :] = get_r_hat_clean(test_log_pr.loc[(t - dt):t], test_volu.loc[(t - dt):t])
t_used = time.time() - t0
r_fwd = (test_log_pr.shift(-30) - test_log_pr).iloc[w::10].rename(columns={f"input_df_{i}": i for i in range(10)})
kuku = r_fwd.corrwith(r_hat)
r_fwd_all = r_fwd.iloc[:-3].values.ravel() # the final "ignore_rows" rows are NaNs.
r_hat_all = r_hat.iloc[:-3].values.ravel()
individual = np.zeros((5, 10))
for j in range(10):
individual[k, j] = np.corrcoef(r_fwd.iloc[:-3, j], r_hat.iloc[:-3, j])[0, 1]
print(individual[k])
return np.corrcoef(r_fwd_all, r_hat_all)[0, 1], kuku
t0 = time.time()
ans, kuku = get_model_corr(log_pr_test, volu_test)
# ans = get_model_corr(log_pr, volu)
tracker.append(ans)
t_used = time.time() - t0
# + colab={"base_uri": "https://localhost:8080/"} id="drxRkjz7wMvR" outputId="c43f9bd1-e6c1-4839-b402-fadcb11faba6"
print(tracker)
print(np.average(tracker))
print(np.median(tracker))
print(np.max(tracker))
print(np.min(tracker))
print(np.average(np.abs(tracker)))
# + colab={"base_uri": "https://localhost:8080/"} id="zkjwTCY9WZUj" outputId="f3bd4e22-e879-4eb1-da44-77822b4c6df6"
for model, idx in split_models:
print(idx, model.coef_)
# + [markdown] id="biH35k64IjnW"
# #Train model on the full dataset
# + colab={"base_uri": "https://localhost:8080/"} id="ok-HEnzPIjLQ" outputId="4dc69ce4-f5b3-4718-ad53-c8742ab87c68"
log_pr_train, _ = log_pr_anchoredfolds[-1]
volu_train, _ = volu_anchoredfolds[-1]
print(log_pr_train.shape)
print(volu_train.shape)
features = construct_features(log_pr_train, volu_train)
print(features.shape)
window_size = 30 # in minutes
X_train, y_train = construct_dataset(window_size, features, log_pr_train)
X_train_by_asset = split_dataset_by_asset(X_train)
# 0, 3, 5 this gave 0.022
print(X_train.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="-NPaS9nlJ8a6" outputId="0b88d850-12ad-4a42-acc1-454980e45952"
split_models = train_models_split(X_train_by_asset, y_train, [0, 1, 2, 3, 4, 5, 6, 7, 8, 9], models)
dump(split_models, 'split_models.joblib')
# + colab={"base_uri": "https://localhost:8080/"} id="J3FXaV0QKEZW" outputId="2b9a0b4d-bf8a-4052-a12b-cb3f246abe7e"
split_models = load('split_models.joblib')
for model, asset_idx in split_models:
print(asset_idx)
print(model)
print(model.coef_)
# + [markdown] id="p_HVo5MMSLWB"
# # main.py to submit
# + colab={"base_uri": "https://localhost:8080/"} id="W3xgQwrASNrw" outputId="09c0b177-0fe0-49d3-c4f7-14f6c4bd5c0f"
split_models = load('split_models.joblib')
for model, asset_idx in split_models:
print(asset_idx, model)
def get_r_hat(A, B):
"""
A: 1440-by-10 dataframe of log prices with columns log_pr_0, ... , log_pr_9
B: 1440-by-10 dataframe of trading volumes with columns volu_0, ... , volu_9
return: a numpy array of length 10, corresponding to the predictions for the forward 30-minutes returns of assets 0, 1, 2, ..., 9
"""
def split_dataset_by_asset(dataset_train):
NUM_ASSETS = 10
return [dataset_train[..., asset::NUM_ASSETS] for asset in range(NUM_ASSETS)]
def construct_local_features(asset_window):
"""
construct features for *single windowed asset*
NOTE: for adding new features, see asset_window[...,0] for np.std
Here, 0 is the feature that is being pulled out (corresponds to the index from global_feature construction)
from which you can do whatever transforms you want
"""
poly = PolynomialFeatures(4, include_bias= False)
neg_back_30_for = -(asset_window[...,0][:, -1] - asset_window[...,0][:, -30]).reshape(-1, 1)
test = poly.fit_transform(neg_back_30_for)
test = [test]
test = np.hstack(test)
return test
def construct_features(log_pr_df, vol_df):
df = log_pr_df.copy()
df = pd.concat([df], axis=1)
return df
window_size = 31
input_features = (construct_features(A, B)).iloc[-window_size:] # only retain tail window
predictions = -(input_features.iloc[-1, :10] - input_features.iloc[-30,:10]).values # init baseline
input_features = input_features.values
split_features_by_asset = split_dataset_by_asset(input_features)
for model, asset_idx in split_models:
predictions[asset_idx] = model.predict(construct_local_features(np.expand_dims(split_features_by_asset[asset_idx], axis=0)))
return predictions
# + [markdown] id="9FN2_cehSYAC"
# ## test for speed
# + colab={"base_uri": "https://localhost:8080/"} id="cVlmYrOASZlo" outputId="5b41cb68-5014-40f3-b391-292a193a9325"
def get_model_corr(log_pr, volu_pr, get_r_hat):
t0 = time.time()
dt = datetime.timedelta(days=1)
r_hat = pd.DataFrame(index=log_pr.index[30::10], columns=np.arange(10), dtype=np.float64)
print(r_hat.shape)
for t in log_pr.index[30::10]: # compute the predictions every 10 minutes
r_hat.loc[t, :] = get_r_hat(log_pr.loc[(t - dt):t], volu_pr.loc[(t - dt):t])
t_used = time.time() - t0
print(t_used)
r_fwd = (log_pr.shift(-30) - log_pr).iloc[30::10].rename(columns={f"input_df_{i}": i for i in range(10)})
# kuku = r_fwd.corrwith(r_hat)
r_fwd_all = r_fwd.iloc[:-3].values.ravel() # the final "ignore_rows" rows are NaNs.
r_hat_all = r_hat.iloc[:-3].values.ravel()
return np.corrcoef(r_fwd_all, r_hat_all)[0, 1]
ans = get_model_corr(log_pr, volu, get_r_hat)
print(ans)
# + [markdown] id="mKUmT18dGBqn"
# # Clustering Analysis
# + id="m-nYo3qSGBPl"
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
std_pr = log_pr.std(axis=0).values.reshape(-1, 1)
r_fwd = (log_pr.shift(-30) - log_pr).iloc[30::10]
mean_pr = r_fwd.mean(axis=0).values.reshape(-1, 1)
std_fwd_pr =r_fwd.std(axis=0).values.reshape(-1, 1)
volu_log = np.log(volu + 1)
v_fwd = (volu_log.shift(-30) - volu_log).iloc[30::10]
std_vol = volu_log.std(axis=0).values.reshape(-1, 1)
mean_vol = v_fwd.mean(axis=0).values.reshape(-1, 1)
std_fwd_vol = v_fwd.std(axis=0).values.reshape(-1, 1)
features = np.concatenate((mean_pr, std_pr, std_vol, mean_vol, std_fwd_vol, std_fwd_pr), axis = 1)
print(features.shape)
distortions = []
K = range(1,10)
for k in K:
kmeanModel = KMeans(n_clusters=k)
kmeanModel.fit(features)
distortions.append(kmeanModel.inertia_)
# + id="6MGAJ1OALqkX"
plt.plot(K, distortions, 'bx-')
plt.xlabel('k')
plt.ylabel('Distortion')
plt.title('The Elbow')
plt.show()
# + id="P1t9qN83M1_Y"
kmeanModel = KMeans(n_clusters=3)
kmeanModel.fit(features)
print(kmeanModel.labels_)
# + id="ev7qIyZ6M8hP"
from sklearn.cluster import AgglomerativeClustering
clustering = AgglomerativeClustering().fit(features)
clustering.labels_
# + [markdown] id="hTSIJGH7JqHt"
# # Data Visualization
# + id="hA3imcLD9o2S"
log_pr.plot(figsize=(12, 8))
# + id="TZaE-FpL8USC"
volu.plot(figsize=(12, 8))
# + [markdown] id="6dR1OlnV0jLX"
# # Appendix
# + [markdown] id="4_v9yGy_Jiec"
# ## Models
# + [markdown] id="ZS5aTovmSQTc"
# ##LSTMs
# + id="izfnP2X2ZrLz"
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Variable
if torch.cuda.is_available():
device = torch.device("cuda")
else:
device = torch.device("cpu")
print(device)
# + id="Kt55VF6CYXx1"
X_train_tensors = Variable(torch.Tensor(X_train).to(device))
y_train_tensors = Variable(torch.Tensor(y_train).to(device))
print("Training Shape", X_train_tensors.shape, y_train_tensors.shape)
# + id="CQHjSqpJSPxG"
class LSTM1(nn.Module):
def __init__(self, output_size, input_size, hidden_size, num_layers):
super(LSTM1, self).__init__()
self.output_size = output_size #number of classes
self.num_layers = num_layers #number of layers
self.input_size = input_size #input size
self.hidden_size = hidden_size #hidden state
self.lstm = nn.LSTM(input_size=input_size, hidden_size=hidden_size,
num_layers=num_layers, batch_first=True) #lstm
self.fc1 = nn.Linear(hidden_size, 64)
self.relu1 = nn.ReLU()
self.fc2 = nn.Linear(64, 32)
self.relu2 = nn.ReLU()
self.fc = nn.Linear(32, output_size) #fully connected last layer
def forward(self,x):
output, (hn, cn) = self.lstm(x) #lstm with input, hidden, and internal state
hn = hn.view(-1, self.hidden_size) #reshaping the data for Dense layer next
out = self.fc1(hn)
out = self.relu1(out)
out = self.fc2(out)
out = self.relu2(out)
out = self.fc(out) #Final Output
return out
# + id="9ndy3ZdgV94S"
num_epochs = 100 #1000 epochs
learning_rate = 0.001 #0.001 lr
input_size = X_train.shape[-1] #number of features
print(input_size)
hidden_size = 128 #number of features in hidden state
num_layers = 1 #number of stacked lstm layers
output_size = 10 #number of output classes
lstm1 = LSTM1(output_size, input_size, hidden_size, num_layers)
lstm1 = lstm1.to(device)
criterion = torch.nn.MSELoss() # mean-squared error for regression
optimizer = torch.optim.Adam(lstm1.parameters(), lr=learning_rate)
print(X_train_tensors.shape)
# + id="08j9WqBBWsSm"
for epoch in range(num_epochs):
outputs = lstm1.forward(X_train_tensors)
optimizer.zero_grad()
# obtain the loss function
loss = criterion(outputs, y_train_tensors)
loss.backward()
optimizer.step() #improve from loss, i.e backprop
if epoch % 2 == 0:
print("Epoch: %d, loss: %1.5f" % (epoch, loss.item()))
# + id="yzZ-1cPQc436"
lstm1 = lstm1.to(torch.device("cpu"))
torch.save(lstm1.state_dict(), "/content/lstm_model.pth")
# + id="uZvySfrKfYEQ"
print( np.expand_dims(log_pr.iloc[-31:, :].to_numpy(), 0).shape)
# + id="blDR3M_w4Kiq"
lstm1.load_state_dict(torch.load("/content/lstm_model.pth"))
lstm1.eval()
lstm1 = lstm1.to(torch.device("cpu"))
# + [markdown] id="vcDi4DmPggul"
# ##GBoost
# + id="q9hoezbohP2r"
model_0 = lgb.LGBMRegressor()
print(X_train.shape)
X_train_asset0 = np.array(X_train[:, :, 0])
X_train_asset2 = np.array(X_train[:, :, 2])
X_train_asset3 = np.array(X_train[:, :, 3])
X_train_asset1 = np.array(X_train[:, :, 1])
y_train_asset0 = np.array(y_train[:, 0])
y_train_asset2 = np.array(y_train[:, 2])
y_train_asset3 = np.array(y_train[:, 3])
y_train_asset1 = np.array(y_train[:, 1])
model_2 = lgb.LGBMRegressor()
model_3 = lgb.LGBMRegressor()
model_1 = lgb.LGBMRegressor()
model_0.fit(X_train_asset0, y_train_asset0)
model_2.fit(X_train_asset2, y_train_asset2)
model_3.fit(X_train_asset3, y_train_asset3)
model_1.fit(X_train_asset1, y_train_asset1)
model_0.booster_.save_model('model_0.txt')
model_2.booster_.save_model('model_2.txt')
model_3.booster_.save_model('model_3.txt')
model_1.booster_.save_model('model_1.txt')
# + id="MMX0vKg-a-va"
model_0 = lgb.Booster(model_file='model_0.txt')
model_2 = lgb.Booster(model_file='model_2.txt')
model_3 = lgb.Booster(model_file='model_3.txt')
model_1 = lgb.Booster(model_file='model_1.txt')
# + [markdown] id="Ms0aBYa1_wnb"
# ##GBoost + Ridge
# + id="HeF86fu3_wZE"
model_2 = lgb.LGBMRegressor()
model_01 = Ridge()
model_35 = Ridge()
#X_train_asset2 = np.array(X_train[:, :, 2])
X_train_asset2 = np.concatenate([X_train[:, :, 2], X_train[:, :, 12], np.array([np.std(X_train[:, :, 2], axis = 1)]).T], axis =1)
X_train_asset01 = np.concatenate((X_train[:, :, 0], X_train[:, :, 1]), axis=1)
X_train_asset35 = np.concatenate((X_train[:, :, 3], X_train[:, :, 5]), axis=1)
y_train_asset2 = np.array(y_train[:, 2])
y_train_asset01 = np.concatenate((y_train[:, 0:1], y_train[:, 1:2]), axis=1)
y_train_asset35 = np.concatenate((y_train[:, 3:4], y_train[:, 5:6]), axis=1)
model_2.fit(X_train_asset2, y_train_asset2)
model_01.fit(X_train_asset01, y_train_asset01)
model_35.fit(X_train_asset35, y_train_asset35)
# + [markdown] id="U6eo1J0-lmpT"
# ##Ridge+Huber
#
#
#
#
# + id="_57x3AyjSK6r"
X_train
# + id="ndINNU0wlwvJ"
def train_models(X_train, y_train):
X_train_asset0 = np.concatenate([X_train[:, :, 0], X_train[:, :, 10], np.array([np.std(X_train[:, :, 0], axis = 1)]).T], axis =1)
X_train_asset1 = np.concatenate([X_train[:, :, 1], X_train[:, :, 11], np.array([np.std(X_train[:, :, 1], axis = 1)]).T], axis =1)
X_train_asset2 = np.concatenate([X_train[:, :, 2], X_train[:, :, 12], np.array([np.std(X_train[:, :, 2], axis = 1)]).T], axis =1)
X_train_asset3 = np.concatenate([X_train[:, :, 3], X_train[:, :, 13], np.array([np.std(X_train[:, :, 3], axis = 1)]).T], axis =1)
X_train_asset4 = np.concatenate([X_train[:, :, 4], X_train[:, :, 14], np.array([np.std(X_train[:, :, 4], axis = 1)]).T], axis =1)
X_train_asset5 = np.concatenate([X_train[:, :, 5], X_train[:, :, 15], np.array([np.std(X_train[:, :, 5], axis = 1)]).T], axis =1)
X_train_asset6 = np.array(X_train[:, :, 6])
X_train_asset7 = np.array(X_train[:, :, 7])
X_train_asset8 = np.array(X_train[:, :, 8])
X_train_asset9 = np.array(X_train[:, :, 9])
y_train_asset0 = np.array(y_train[:, 0])
y_train_asset1 = np.array(y_train[:, 1])
y_train_asset2 = np.array(y_train[:, 2])
y_train_asset3 = np.array(y_train[:, 3])
y_train_asset4 = np.array(y_train[:, 4])
y_train_asset5 = np.array(y_train[:, 5])
y_train_asset6 = np.array(y_train[:, 6])
y_train_asset7 = np.array(y_train[:, 7])
y_train_asset8 = np.array(y_train[:, 8])
y_train_asset9 = np.array(y_train[:, 9])
model_0 = Ridge()
model_1 = Ridge()
model_2 = Ridge()
model_3 = Ridge()
model_4 = Ridge()
model_5 = Ridge()
print(X_train_asset0.shape)
model_0.fit(X_train_asset0, y_train_asset0)
model_1.fit(X_train_asset1, y_train_asset1)
model_2.fit(X_train_asset2, y_train_asset2)
model_3.fit(X_train_asset3, y_train_asset3)
#model_4.fit(X_train_asset4, y_train_asset4)
model_5.fit(X_train_asset5, y_train_asset5)
#model_6.fit(X_train_asset6, y_train_asset6)
##model_7.fit(X_train_asset7, y_train_asset7)
#model_8.fit(X_train_asset8, y_train_asset8)
#model_9.fit(X_train_asset9, y_train_asset9)
return model_0, model_1, model_2, model_3, model_5
# + [markdown] id="PqnE4kzzhSsY"
# ## Evaluation
# + id="9mprkHj3xwt6"
# Use the negative 30-minutes backward log-returns to predict the 30-minutes forward log-returns
#predict the log price, and then do correlation
def get_r_hat_baseline(A, B):
return -(A.iloc[-1] - A.iloc[-30]).values
# + id="u41btF83fFM8"
def get_r_hat_lstm(A, B):
input = np.expand_dims(construct_features(A, B).values, axis=0)
input = Variable(torch.Tensor(input))
pred = lstm1(input).detach().cpu().numpy()
return pred.squeeze()
# + id="1tEScM3UotCy"
def get_r_hat_complex(A, B):
w = 31
input = construct_features(A, B)
tmp = -(input.iloc[-1] - input.iloc[-30]).values
asset_0_pred = model_0.predict(np.expand_dims(input.iloc[-w:, 0].values, axis=0))
#asset_2_pred = model_2.predict(np.expand_dims(input.iloc[-w:, 2].values, axis=0))
asset_2_pred = model_2.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 2].values, input.iloc[-w:, 12].values, [np.std(input.iloc[-w:, 2].values)]]), axis=0))
asset_3_pred = model_3.predict(np.expand_dims(input.iloc[-w:, 3].values, axis=0))
asset_1_pred = model_1.predict(np.expand_dims(input.iloc[-w:, 1].values, axis=0))
tmp[0] = asset_0_pred[0]
tmp[2] = asset_2_pred[0]
tmp[3] = asset_3_pred[0]
tmp[1] = asset_1_pred[0]
return tmp
# + id="Hzduuz7BlZw3"
def get_r_hat_more_features(A,B):
w = 61
input = construct_features(A, B)
tmp = -(input.iloc[-1, :10] - input.iloc[-30,:10]).values
asset_0_pred = model_0.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 0].values, input.iloc[-w:, 10].values, [np.std(input.iloc[-w:, 0].values)]]), axis=0))
asset_2_pred = model_2.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 2].values, input.iloc[-w:, 12].values, [np.std(input.iloc[-w:, 2].values)]]), axis=0))
asset_3_pred = model_3.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 3].values, input.iloc[-w:, 13].values, [np.std(input.iloc[-w:, 3].values)]]), axis=0))
asset_1_pred = model_1.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 1].values, input.iloc[-w:, 11].values, [np.std(input.iloc[-w:, 1].values)]]), axis=0))
#asset_4_pred = model_4.predict(np.expand_dims(np.concatenate([input.iloc[-61:, 4].values, input.iloc[-61:, 14].values, [np.std(input.iloc[-61:, 4].values)]]), axis=0))
asset_5_pred = model_5.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 5].values, input.iloc[-w:, 15].values, [np.std(input.iloc[-w:, 5].values)]]), axis=0))
#asset_5_pred = model_5.predict(np.expand_dims(input.iloc[-61:, 5].values, axis=0))
#asset_6_pred = model_6.predict(np.expand_dims(input.iloc[-61:, 6].values, axis=0))
#asset_7_pred = model_7.predict(np.expand_dims(input.iloc[-61:, 7].values, axis=0))
#asset_8_pred = model_8.predict(np.expand_dims(input.iloc[-61:, 8].values, axis=0))
#asset_9_pred = model_9.predict(np.expand_dims(input.iloc[-61:, 9].values, axis=0))
tmp[0] = asset_0_pred[0]
tmp[2] = asset_2_pred[0]
tmp[3] = asset_3_pred[0]
tmp[1] = asset_1_pred[0]
#tmp[4] = asset_4_pred[0]
tmp[5] = asset_5_pred[0]
#tmp[6] = asset_6_pred[0]
#tmp[7] = asset_7_pred[0]
#tmp[8] = asset_8_pred[0]
#tmp[9] = asset_9_pred[0]
return tmp
# + id="_vGoGX5jot55"
def get_r_hat_simple(A, B):
input = construct_features(A, B)
return -(input.iloc[-1] - input.iloc[-30]).values
# + id="8Vc2T3S8Be-q"
def get_r_hat_ridge_boost(A, B):
input = construct_features(A, B)
tmp = -(input.iloc[-1] - input.iloc[-30]).values
#asset_2_pred = model_2.predict(np.expand_dims(input.iloc[-61:, 2].values, axis=0))
asset_2_pred = model_2.predict(np.expand_dims(np.concatenate([input.iloc[-61:, 2].values, input.iloc[-61:, 12].values, [np.std(input.iloc[-61:, 2].values)]]), axis=0))
input_01 = np.concatenate((input.iloc[-61:, 0:1].values, input.iloc[-61:, 1:2].values), axis=1).reshape(1, -1)
asset_01_pred = np.squeeze(model_01.predict(input_01))
input_35 = np.concatenate((input.iloc[-61:, 3:4].values, input.iloc[-61:, 5:6].values), axis=1).reshape(1, -1)
asset_35_pred = np.squeeze(model_35.predict(input_35))
tmp[2] = asset_2_pred[0]
tmp[0] = asset_01_pred[0]
tmp[1] = asset_01_pred[1]
tmp[3] = asset_35_pred[0]
tmp[5] = asset_35_pred[1]
return tmp
# + id="qMc8AuzExwt6"
# An example of get_r_hat
ACTIVE_R_HAT = "more_features"
r_hat_implementations = {
"baseline": get_r_hat_baseline,
"lstm": get_r_hat_lstm,
"simple":get_r_hat_simple,
"complex": get_r_hat_complex,
"ridge_boost": get_r_hat_ridge_boost,
"more_features": get_r_hat_more_features
}
def get_r_hat(A, B):
"""
A: 1440-by-10 dataframe of log prices with columns log_pr_0, ... , log_pr_9
B: 1440-by-10 dataframe of trading volumes with columns volu_0, ... , volu_9
return: a numpy array of length 10, corresponding to the predictions for the forward 30-minutes returns of assets 0, 1, 2, ..., 9
"""
return r_hat_implementations[ACTIVE_R_HAT](A, B)
# + id="RN6LhZA_xwt7"
def get_model_corr(test_log_pr, test_volu):
t0 = time.time()
dt = datetime.timedelta(days=1)
r_hat = pd.DataFrame(index=test_log_pr.index[1440::10], columns=np.arange(10), dtype=np.float64)
for t in test_log_pr.index[1440::10]: # compute the predictions every 10 minutes
r_hat.loc[t, :] = get_r_hat(test_log_pr.loc[(t - dt):t], test_volu.loc[(t - dt):t])
t_used = time.time() - t0
r_fwd = (test_log_pr.shift(-30) - test_log_pr).iloc[1440::10].rename(columns={f"input_df_{i}": i for i in range(10)})
r_fwd.corrwith(r_hat)
r_fwd_all = r_fwd.iloc[:-3].values.ravel() # the final "ignore_rows" rows are NaNs.
r_hat_all = r_hat.iloc[:-3].values.ravel()
return np.corrcoef(r_fwd_all, r_hat_all)[0, 1]
# + id="LXKZAmnjRUap"
tracker = []
for k in range(len(log_pr_kfolds)):
print(k)
log_pr_train, log_pr_test = log_pr_kfolds[k]
volu_train, volu_test = log_pr_kfolds[k]
features = construct_features(log_pr_train, volu_train)
window_size = 60 # in minutes
X_train, y_train = construct_dataset(window_size, features, log_pr_train)
print(X_train.shape)
model_0, model_1, model_2, model_3, model_5 = train_models(X_train, y_train)
def get_r_hat(A,B):
w = 61
input = construct_features(A, B)
tmp = -(input.iloc[-1, :10] - input.iloc[-30,:10]).values
asset_0_pred = model_0.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 0].values, input.iloc[-w:, 10].values, [np.std(input.iloc[-w:, 0].values)]]), axis=0))
asset_2_pred = model_2.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 2].values, input.iloc[-w:, 12].values, [np.std(input.iloc[-w:, 2].values)]]), axis=0))
asset_3_pred = model_3.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 3].values, input.iloc[-w:, 13].values, [np.std(input.iloc[-w:, 3].values)]]), axis=0))
asset_1_pred = model_1.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 1].values, input.iloc[-w:, 11].values, [np.std(input.iloc[-w:, 1].values)]]), axis=0))
asset_5_pred = model_5.predict(np.expand_dims(np.concatenate([input.iloc[-w:, 5].values, input.iloc[-w:, 15].values, [np.std(input.iloc[-w:, 5].values)]]), axis=0))
tmp[0] = asset_0_pred[0]
tmp[2] = asset_2_pred[0]
tmp[3] = asset_3_pred[0]
tmp[1] = asset_1_pred[0]
tmp[5] = asset_5_pred[0]
return tmp
def get_model_corr(test_log_pr, test_volu):
w = 60
t0 = time.time()
dt = datetime.timedelta(days=1)
r_hat = pd.DataFrame(index=test_log_pr.index[w::10], columns=np.arange(10), dtype=np.float64)
print(r_hat.shape)
for t in test_log_pr.index[w::10]: # compute the predictions every 10 minutes
r_hat.loc[t, :] = get_r_hat_clean(test_log_pr.loc[(t - dt):t], test_volu.loc[(t - dt):t])
t_used = time.time() - t0
r_fwd = (test_log_pr.shift(-30) - test_log_pr).iloc[w::10].rename(columns={f"input_df_{i}": i for i in range(10)})
r_fwd.corrwith(r_hat)
r_fwd_all = r_fwd.iloc[:-3].values.ravel() # the final "ignore_rows" rows are NaNs.
r_hat_all = r_hat.iloc[:-3].values.ravel()
return np.corrcoef(r_fwd_all, r_hat_all)[0, 1]
t0 = time.time()
ans = get_model_corr(log_pr_test, volu_test)
tracker.append(ans)
t_used = time.time() - t0
# + id="xC69U6aoW8sg"
print(tracker)
print(np.average(tracker))
print(np.median(tracker))
print(np.max(tracker))
print(np.min(tracker))
print(np.average(np.abs(tracker)))
| stats_601_project.ipynb |