code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizations
# +
# Observe the rug plot of the first 1000 rows of Sales that you created in the previous question.
# Which of these ranges has not a single data point present between it?
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# the commonly used alias for seaborn is sns
import seaborn as sns
# set a seaborn style of your taste
sns.set_style("whitegrid")
# data
df = pd.read_csv("../Data/global_sales_data/market_fact.csv")
# rug = True
# plotting only a few points since rug takes a long while
sns.distplot(df['Sales'][:1000], rug=True)
plt.show()
# -
df = df[(df.Profit > 0)]
sns.jointplot('Sales', 'Profit', df)
plt.show()
# +
# Now, say you want to do this the other way round - different sub-plots for each categories,
# and divisions for customer segments inside each sub-plot.
market_df = pd.read_csv("../Data/global_sales_data/market_fact.csv")
customer_df = pd.read_csv("../Data/global_sales_data/cust_dimen.csv")
product_df = pd.read_csv("../Data/global_sales_data/prod_dimen.csv")
shipping_df = pd.read_csv("../Data/global_sales_data/shipping_dimen.csv")
orders_df = pd.read_csv("../Data/global_sales_data/orders_dimen.csv")
df = pd.merge(market_df, product_df, how='inner', on='Prod_id')
df = pd.merge(df, customer_df, how='inner', on='Cust_id')
# set figure size for larger figure
plt.figure(num=None, figsize=(12, 8), dpi=80, facecolor='w', edgecolor='k')
# specify hue="categorical_variable"
sns.boxplot(x='Product_Category', y='Profit', hue="Customer_Segment", data=df)
plt.show()
| Practice/Visualizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Local homology: tutorial
#
# <iframe src="./implemented_variants.pdf" width=200 height=200></iframe>
# %load_ext autoreload
# %autoreload 2
# +
import numpy as np
from gudhi import plot_persistence_diagram
from gudhi.datasets.generators import points
from sklearn import datasets
import matplotlib.pyplot as plt
from local_homology import compute_local_homology_alpha, compute_local_homology_r
from local_homology.alpha_filtration import distance_to_point_outside_ball, distance_to_expanding_boundary, distance_to_boundary
from local_homology.dataset import intersecting_lines
from local_homology.r_filtration import plot_one_skeleton
from local_homology.vis import plot_disc, plot_point_cloud, plot_rectangle
# -
np.random.seed(0)
# ## Local homology at the intersection of two segments
# +
point_cloud = intersecting_lines(100, 0.01)
x0 = np.array([[0., 0.]])
epsilon = 0.2
plot_point_cloud(point_cloud)
# -
# ### $\alpha$-filtration
# We consider the points in the disc and we perform a Vietoris-Rips filtration by growing balls around those points. There are three variants on how to compute the distance of a point to the boundary.
plot_point_cloud(point_cloud)
plot_disc(x0[0], epsilon, color="r", label=r"Neighborhood for $\alpha$-filtr")
alpha_dgm = compute_local_homology_alpha(point_cloud, x0, epsilon, 2,
distances=distance_to_point_outside_ball)
_ = plot_persistence_diagram(alpha_dgm)
alpha_dgm_1 = [(b, d)for dim, (b, d) in alpha_dgm if dim==1.]
alpha_dgm_1
# We see indeed 3 prominent points in $H_1$, what corresponds to the 4 branches coming out from the center.
# **Surprisingly, we also see some $H_2$ if we call the fnuction with `max_dimension=2`.**
# +
alpha_dgm_boundary = compute_local_homology_alpha(
point_cloud, x0, epsilon, 2, distances=distance_to_boundary
)
alpha_dgm_expanding = compute_local_homology_alpha(
point_cloud, x0, epsilon, 2, distances=distance_to_expanding_boundary)
fig, axes = plt.subplots(1, 3, figsize=(10, 3))
_ = plot_persistence_diagram(alpha_dgm, axes=axes[0])
_ = plot_persistence_diagram(alpha_dgm_boundary, axes=axes[1])
_ = plot_persistence_diagram(alpha_dgm_expanding, axes=axes[2])
plt.suptitle("Distance to")
axes[0].set_title("point outside")
axes[1].set_title("boundary")
axes[2].set_title("expanding boundary")
plt.tight_layout()
# -
# ### $r$ filtration
#
# We fix a Rips-scale $\alpha$ and we build the filtration by considering a smaller and smaller neighborhood around $x_0$. **Since the homology is computed via a duality theorem, we do not get $H_0$.**
# +
alpha = 0.1
plot_point_cloud(point_cloud)
for x in point_cloud:
plot_disc(x, alpha, alpha=alpha, color="r", label="_nolegend_")
# the keyword parameter alpha is the opacity of the circles, no the scale.
plot_one_skeleton(point_cloud, x0, alpha)
# -
r_dgm = compute_local_homology_r(point_cloud, x0, alpha, 2)
_ = plot_persistence_diagram(r_dgm)
r_dgm_1 = [(b, d)for dim, (b, d) in r_dgm if dim==1.]
r_dgm_1
# ### Different center
# For the chosen parameters, both methods show 3 persistent points, recovering the correct local homology. Let's pick a point where local homology is different.
#
# At $x_1 = (-0.25, -0.25)$, the local homology group in dimension 1 has one generator and groups in all other dimensions are trivial.
# +
x1 = np.array([[-0.25, -0.25]])
plot_point_cloud(point_cloud)
for x in point_cloud:
plot_disc(x, alpha, alpha=0.03, color="r", label="_nolegend_")
plot_disc(x1[0], epsilon, alpha=0.4, color="blue")
# +
x = x1
alpha_dgm1 = compute_local_homology_alpha(point_cloud, x, epsilon, 2)
_ = plot_persistence_diagram(alpha_dgm1)
# -
# There is one (persistent) point in $H_1$, as expected.
r_dgm1 = compute_local_homology_r(point_cloud, x, alpha, 2)
plot_persistence_diagram(r_dgm1)
r_dgm1
# This diagram is more interesting. Remember that we are quotienting by everything but a smaller and smaller neighborhood.
#
# There are three points in total. Two appear, and die, for high parameter values, namely $[0.38, 075]$ is an interval where they both persist. In this particular example, we should be able to see them for $r= 0.6$. Visual inspection confirms that the $r$-ball contains the intersection of lines and so that the points correspond to things happening "far" from the localization point.
#
# The point closer to the origin persists through $(0.03, 0.33)$. Let's inspect $r=0.2$ and $r=0.5$. We see that we recover the correct homology in that interval. The homology can be read-off as the number of points in the rectangle $\rbrack-\infty,r\rbrack\times\rbrack r, \infty\lbrack$ in the diagram.
plot_point_cloud(point_cloud)
plot_disc(x[0], 0.5, color="red", label="r=0.5")
plot_disc(x[0], 0.2, color="green", label="r=0.2")
plt.legend()
r_dgm1 = compute_local_homology_r(point_cloud, x, alpha, 2)
plot_persistence_diagram(r_dgm1)
plot_rectangle(0.2, color="green")
plot_rectangle(0.5, color="red")
r_dgm1
# An interesting question is what happens when we place the point closer to the intersection point, so that the bottom-left endpoint of the segment is outside of the ball while the intersection is still inside. Let us examine that situation for both filtrations.
x = np.array([[-0.21, -0.21]])
r_dgm2 = compute_local_homology_r(point_cloud, x, alpha, 2)
r_dgm2
# +
plot_persistence_diagram(r_dgm2)
plot_rectangle(0.38, color="red")
plt.figure()
plot_point_cloud(point_cloud)
for p in point_cloud:
plot_disc(p, alpha, alpha=0.03, color="r", label="_nolegend_")
plot_disc(x[0], 0.38, color="green", label="r=0.5")
# -
# For the $r-$filtration, it does not change much. We have the same number of points, but now, all bars live through (an interval that includes) $[0.33, 0.39]$.
# +
alpha_dgm2 = compute_local_homology_alpha(point_cloud, x, epsilon, 2)
_ = plot_persistence_diagram(alpha_dgm2)
plt.figure()
plot_point_cloud(point_cloud)
plot_disc(x[0], epsilon, color="red", label=r"$\epsilon$=" + str(epsilon))
# -
alpha_dgm2_h1, alpha_dgm1_h1 = [[p for p in dgm if p[0]==1]
for dgm in [alpha_dgm2, alpha_dgm1]]
alpha_dgm2_h1, alpha_dgm1_h1
# For the $\alpha$-filtration, it does not change anything, since the intersection point is not in the $\epsilon$-ball of the center $(0,0)\notin B(x, \epsilon)$.
# ## Densely sampled square
np.random.seed(2)
# +
point_cloud = np.random.rand(400, 2)
plot_point_cloud(point_cloud)
# +
x0 = np.array([[0.25, 0.5]])
epsilon = 0.2
plot_point_cloud(point_cloud)
plot_disc(x0[0], epsilon)
# -
alpha_square = compute_local_homology_alpha(point_cloud, x0, epsilon, 2)
_ = plot_persistence_diagram(alpha_square)
alpha = 0.2
r_square = compute_local_homology_r(point_cloud, x0, alpha, 2)
_ = plot_persistence_diagram(r_square)
r_square
# Both diagrams show some $H_2$, but it's not very persistent. it dies at 0.24 and it might be because the ball centered at $(0.25, 0.25)$ approaches the boundary $y=0$ of the support of the distribution.
# ## Sphere
# +
point_cloud = points.sphere(n_samples=3000, ambient_dim=3, radius=1, sample="random")
point_cloud = point_cloud[point_cloud[:,2]>=0.0]
ax = plt.axes(projection='3d')
_ = ax.scatter3D(point_cloud[:,0], point_cloud[:, 1], point_cloud[:, 2],)
# +
x0 = np.array([[0., 0., 1.]])
epsilon = 0.35
from local_homology.alpha_filtration import is_point_in_ball
in_ball, _ = is_point_in_ball(point_cloud, x0, epsilon, return_distances=True)
pts_in_ball = point_cloud[in_ball]
ax = plt.axes(projection='3d')
_ = ax.scatter3D(pts_in_ball[:,0], pts_in_ball[:, 1], pts_in_ball[:, 2],)
# -
alpha_sphere = compute_local_homology_alpha(point_cloud, x0, epsilon, 2)
_ = plot_persistence_diagram(alpha_sphere)
_ = plt.title(r"Pseudo $\alpha$-filtration")
# *Observation: The proposed $\alpha$-filtration with `distances=distance_to_boundary` is close to ~~equivalent~~ to covering the boundary of the disc with points and performing a Vietoris-Rips filtration. It is not equivalent, because a 2-simplex is added as soon as there is an edge between points, which are connected with the boundary. They do not need to connect to the same point.*
alpha_sphere = compute_local_homology_alpha(point_cloud, x0, epsilon, 2, distance_to_boundary)
_ = plot_persistence_diagram(alpha_sphere)
_ = plt.title(r"Pseudo $\alpha$-filtration, distance to boundary")
alpha_sphere = compute_local_homology_alpha(point_cloud, x0, epsilon, 2, distance_to_expanding_boundary)
_ = plot_persistence_diagram(alpha_sphere)
_ = plt.title(r"Pseudo $\alpha$-filtration, distance to expanding boundary")
# +
alpha = 0.2
r_sphere = compute_local_homology_r(point_cloud, x0, alpha, 2)
_ = plot_persistence_diagram(r_sphere)
_ = plt.title(r"$r$-filtration")
# -
# For the $\alpha$-filtration, the expanding boundary seems to fill out the interior too quickly for $H_2$ to appear. *I* suspect that the points connect to the boundary quicker than they connect to each other, so that no 2-simplices have time to form. Indeed, if we disable this behavior, we observe a non-trivial $2$-dim hole.
#
# Still, we can get an interesting observation from here. By comparing the persistence diagram from `distance_to_boundary` and `distance_to_expanding_boundary`, we can speculate which points in the diagram correspond to cycles that were created by joining the boundary, and which were created purely inside the $r$-ball. For the first category, their birth value will be smaller, while the other ones should see their death values affected.
#
# For the $r-$filtration, this appears immediately quickly and persists. Of course, this is due to a good scale choice for Rips.
# ## Swiss hole
# +
### from https://github.com/scikit-learn/scikit-learn/blob/main/examples/manifold/plot_swissroll.py
sh_points, sh_color = datasets.make_swiss_roll(
n_samples=11000, hole=True, random_state=0
)
t = np.array([7.853, 12.5, 12.5])
y = np.array([8., 20., 8.])
pts = np.stack([t*np.cos(t), y, t*np.sin(t)], axis=1)
# -
fig = plt.figure(figsize=(8, 6))
ax = fig.add_subplot(111, projection="3d")
fig.add_axes(ax)
ax.scatter(
sh_points[:, 0], sh_points[:, 1], sh_points[:, 2],
c=sh_color, s=5, alpha=0.15
)
ax.scatter(pts[:, 0], pts[:, 1], pts[:, 2], c='r', s=20, alpha=0.8)
ax.set_title("Swiss-Hole in Ambient Space")
ax.view_init(azim=-66, elev=12)
_ = ax.text2D(0.8, 0.05, s="n_samples=1500", transform=ax.transAxes)
pt_hole, pt_boundary, pt_inside = pts
# +
alpha = 0.9
r_sh = [compute_local_homology_r(sh_points, x0, alpha, 2, max_r=9.) for x0 in pts]
# +
fig, axes = plt.subplots(1, 3, figsize=(10, 3))
for (ind, dgm), name in zip(enumerate(r_sh), ["hole", "boundary", "inside"]):
_ = plot_persistence_diagram(dgm, axes=axes[ind])
axes[ind].set_title(name)
_ = plt.suptitle(r"$r$-filtration")
# -
epsilon = 2.
dgms = [compute_local_homology_alpha(sh_points, x0[None,:], epsilon, 2,
distances=distance_to_point_outside_ball, approximate=True) for x0 in pts]
min_, max_ = 0., epsilon
for dgm, name in zip(dgms, ["hole", "boundary", "inside"]):
ax = plot_persistence_diagram(dgm, )
ax.set_xlim([min_, max_]),
ax.set_ylim([min_, max_])
ax.set_title(name)
| notebooks/Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 视频动作识别
# 视频动作识别是指对一小段视频中的内容进行分析,判断视频中的人物做了哪种动作。视频动作识别与图像领域的图像识别,既有联系又有区别,图像识别是对一张静态图片进行识别,而视频动作识别不仅要考察每张图片的静态内容,还要考察不同图片静态内容之间的时空关系。比如一个人扶着一扇半开的门,仅凭这一张图片无法判断该动作是开门动作还是关门动作。
#
# 视频分析领域的研究相比较图像分析领域的研究,发展时间更短,也更有难度。视频分析模型完成的难点首先在于,需要强大的计算资源来完成视频的分析。视频要拆解成为图像进行分析,导致模型的数据量十分庞大。视频内容有很重要的考虑因素是动作的时间顺序,需要将视频转换成的图像通过时间关系联系起来,做出判断,所以模型需要考虑时序因素,加入时间维度之后参数也会大量增加。
#
# 得益于PASCAL VOC、ImageNet、MS COCO等数据集的公开,图像领域产生了很多的经典模型,那么在视频分析领域有没有什么经典的模型呢?答案是有的,本案例将为大家介绍视频动作识别领域的经典模型并进行代码实践。
#
# 由于本案例的代码是在华为云ModelArts Notebook上运行,所以需要先按照如下步骤来进行Notebook环境的准备。
#
# ### 进入ModelArts
#
# 点击如下链接:https://www.huaweicloud.com/product/modelarts.html , 进入ModelArts主页。点击“立即使用”按钮,输入用户名和密码登录,进入ModelArts使用页面。
#
# ### 进入ModelArts
#
# 点击如下链接:https://www.huaweicloud.com/product/modelarts.html , 进入ModelArts主页。点击“立即使用”按钮,输入用户名和密码登录,进入ModelArts使用页面。
#
# ### 创建ModelArts notebook
#
# 下面,我们在ModelArts中创建一个notebook开发环境,ModelArts notebook提供网页版的Python开发环境,可以方便的编写、运行代码,并查看运行结果。
#
# 第一步:在ModelArts服务主界面依次点击“开发环境”、“创建”
#
# 
#
# 第二步:填写notebook所需的参数:
#
# 
#
# 第三步:配置好notebook参数后,点击下一步,进入notebook信息预览。确认无误后,点击“立即创建”
# 
#
# 第四步:创建完成后,返回开发环境主界面,等待Notebook创建完毕后,打开Notebook,进行下一步操作。
# 
#
# ### 在ModelArts中创建开发环境
#
# 接下来,我们创建一个实际的开发环境,用于后续的实验步骤。
#
# 第一步:点击下图所示的“启动”按钮,加载后“打开”按钮变从灰色变为蓝色后点击“打开”进入刚刚创建的Notebook
# 
# 
#
#
# 第二步:创建一个Python3环境的的Notebook。点击右上角的"New",然后选择TensorFlow 1.13.1开发环境。
#
# 第三步:点击左上方的文件名"Untitled",并输入一个与本实验相关的名称,如"action_recognition"
# 
# 
#
#
# ### 在Notebook中编写并执行代码
#
# 在Notebook中,我们输入一个简单的打印语句,然后点击上方的运行按钮,可以查看语句执行的结果:
# 
#
#
# 开发环境准备好啦,接下来可以愉快地写代码啦!
# ### 准备源代码和数据
#
# 这一步准备案例所需的源代码和数据,相关资源已经保存在OBS中,我们通过[ModelArts SDK](https://support.huaweicloud.com/sdkreference-modelarts/modelarts_04_0002.html)将资源下载到本地,并解压到当前目录下。解压后,当前目录包含data、dataset_subset和其他目录文件,分别是预训练参数文件、数据集和代码文件等。
import os
if not os.path.exists('videos'):
from modelarts.session import Session
session = Session()
session.download_data(bucket_path="ai-course-common-26-bj4/video/video.tar.gz", path="./video.tar.gz")
# 使用tar命令解压资源包
os.system("tar xf ./video.tar.gz")
# 使用rm命令删除压缩包
os.system("rm ./video.tar.gz")
# 上一节课我们已经介绍了视频动作识别有HMDB51、UCF-101和Kinetics三个常用的数据集,本案例选用了UCF-101数据集的部分子集作为演示用数据集,接下来,我们播放一段UCF-101中的视频:
video_name = "./data/v_TaiChi_g01_c01.avi"
# +
from IPython.display import clear_output, Image, display, HTML
import time
import cv2
import base64
import numpy as np
def arrayShow(img):
_,ret = cv2.imencode('.jpg', img)
return Image(data=ret)
cap = cv2.VideoCapture(video_name)
while True:
try:
clear_output(wait=True)
ret, frame = cap.read()
if ret:
tmp = cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)
img = arrayShow(frame)
display(img)
time.sleep(0.05)
else:
break
except KeyboardInterrupt:
cap.release()
cap.release()
# -
# ## 视频动作识别模型介绍
#
# 在图像领域中,ImageNet作为一个大型图像识别数据集,自2010年开始,使用此数据集训练出的图像算法层出不穷,深度学习模型经历了从AlexNet到VGG-16再到更加复杂的结构,模型的表现也越来越好。在识别千种类别的图片时,错误率表现如下:
#
# <img src="./img/ImageNet.png" width="500" height="500" align=center>
#
# 在图像识别中表现很好的模型,可以在图像领域的其他任务中继续使用,通过复用模型中部分层的参数,就可以提升模型的训练效果。有了基于ImageNet模型的图像模型,很多模型和任务都有了更好的训练基础,比如说物体检测、实例分割、人脸检测、人脸识别等。
#
# 那么训练效果显著的图像模型是否可以用于视频模型的训练呢?答案是yes,有研究证明,在视频领域,如果能够复用图像模型结构,甚至参数,将对视频模型的训练有很大帮助。但是怎样才能复用上图像模型的结构呢?首先需要知道视频分类与图像分类的不同,如果将视频视作是图像的集合,每一个帧将作为一个图像,视频分类任务除了要考虑到图像中的表现,也要考虑图像间的时空关系,才可以对视频动作进行分类。
#
# 为了捕获图像间的时空关系,论文[I3D](https://arxiv.org/pdf/1705.07750.pdf)介绍了三种旧的视频分类模型,并提出了一种更有效的Two-Stream Inflated 3D ConvNets(简称I3D)的模型,下面将逐一简介这四种模型,更多细节信息请查看原论文。
#
# ### 旧模型一:卷积网络+LSTM
#
# 模型使用了训练成熟的图像模型,通过卷积网络,对每一帧图像进行特征提取、池化和预测,最后在模型的末端加一个LSTM层(长短期记忆网络),如下图所示,这样就可以使模型能够考虑时间性结构,将上下文特征联系起来,做出动作判断。这种模型的缺点是只能捕获较大的工作,对小动作的识别效果较差,而且由于视频中的每一帧图像都要经过网络的计算,所以训练时间很长。
#
# <img src="./img/video_model_0.png" width="200" height="200" align=center>
#
# ### 旧模型二:3D卷积网络
#
# 3D卷积类似于2D卷积,将时序信息加入卷积操作。虽然这是一种看起来更加自然的视频处理方式,但是由于卷积核维度增加,参数的数量也增加了,模型的训练变得更加困难。这种模型没有对图像模型进行复用,而是直接将视频数据传入3D卷积网络进行训练。
#
# <img src="./img/video_model_1.png" width="150" height="150" align=center>
#
# ### 旧模型三:Two-Stream 网络
#
# Two-Stream 网络的两个流分别为**1张RGB快照**和**10张计算之后的光流帧画面组成的栈**。两个流都通过ImageNet预训练好的图像卷积网络,光流部分可以分为竖直和水平两个通道,所以是普通图片输入的2倍,模型在训练和测试中表现都十分出色。
#
# <img src="./img/video_model_2.png" width="400" height="400" align=center>
#
# #### 光流视频 optical flow video
#
# 上面讲到了光流,在此对光流做一下介绍。光流是什么呢?名字很专业,感觉很陌生,但实际上这种视觉现象我们每天都在经历,我们坐高铁的时候,可以看到窗外的景物都在快速往后退,开得越快,就感受到外面的景物就是“刷”地一个残影,这种视觉上目标的运动方向和速度就是光流。光流从概念上讲,是对物体运动的观察,通过找到相邻帧之间的相关性来判断帧之间的对应关系,计算出相邻帧画面中物体的运动信息,获取像素运动的瞬时速度。在原始视频中,有运动部分和静止的背景部分,我们通常需要判断的只是视频中运动部分的状态,而光流就是通过计算得到了视频中运动部分的运动信息。
#
# 下面是一个经过计算后的原视频及光流视频。
#
# 原视频
# 
# 光流视频
# 
#
# ### 新模型:Two-Stream Inflated 3D ConvNets
#
# 新模型采取了以下几点结构改进:
# - 拓展2D卷积为3D。直接利用成熟的图像分类模型,只不过将网络中二维$ N × N $的 filters 和 pooling kernels 直接变成$ N × N × N $;
# - 用 2D filter 的预训练参数来初始化 3D filter 的参数。上一步已经利用了图像分类模型的网络,这一步的目的是能利用上网络的预训练参数,直接将 2D filter 的参数直接沿着第三个时间维度进行复制N次,最后将所有参数值再除以N;
# - 调整感受野的形状和大小。新模型改造了图像分类模型Inception-v1的结构,前两个max-pooling层改成使用$ 1 × 3 × 3 $kernels and stride 1 in time,其他所有max-pooling层都仍然使用对此的kernel和stride,最后一个average pooling层使用$ 2 × 7 × 7 $的kernel。
# - 延续了Two-Stream的基本方法。用双流结构来捕获图片之间的时空关系仍然是有效的。
#
# 最后新模型的整体结构如下图所示:
#
# <img src="./img/video_model_3.png" width="200" height="200" align=center>
#
# 好,到目前为止,我们已经讲解了视频动作识别的经典数据集和经典模型,下面我们通过代码来实践地跑一跑其中的两个模型:**C3D模型**( 3D卷积网络)以及**I3D模型**(Two-Stream Inflated 3D ConvNets)。
#
# ### C3D模型结构
#
#
# 我们已经在前面的“旧模型二:3D卷积网络”中讲解到3D卷积网络是一种看起来比较自然的处理视频的网络,虽然它有效果不够好,计算量也大的特点,但它的结构很简单,可以构造一个很简单的网络就可以实现视频动作识别,如下图所示是3D卷积的示意图:
#
# 
#
# a)中,一张图片进行了2D卷积, b)中,对视频进行2D卷积,将多个帧视作多个通道, c)中,对视频进行3D卷积,将时序信息加入输入信号中。
#
# ab中,output都是一张二维特征图,所以无论是输入是否有时间信息,输出都是一张二维的特征图,2D卷积失去了时序信息。只有3D卷积在输出时,保留了时序信息。2D和3D池化操作同样有这样的问题。
#
# 如下图所示是一种[C3D](https://www.cv-foundation.org/openaccess/content_iccv_2015/papers/Tran_Learning_Spatiotemporal_Features_ICCV_2015_paper.pdf)网络的变种:(如需阅读原文描述,请查看I3D论文 2.2 节)
#
# 
#
# C3D结构,包括8个卷积层,5个最大池化层以及2个全连接层,最后是softmax输出层。
#
# 所有的3D卷积核为$ 3 × 3 × 3$ 步长为1,使用SGD,初始学习率为0.003,每150k个迭代,除以2。优化在1.9M个迭代的时候结束,大约13epoch。
#
# 数据处理时,视频抽帧定义大小为:$ c × l × h × w$,$c$为通道数量,$l$为帧的数量,$h$为帧画面的高度,$w$为帧画面的宽度。3D卷积核和池化核的大小为$ d × k × k$,$d$是核的时间深度,$k$是核的空间大小。网络的输入为视频的抽帧,预测出的是类别标签。所有的视频帧画面都调整大小为$ 128 × 171 $,几乎将UCF-101数据集中的帧调整为一半大小。视频被分为不重复的16帧画面,这些画面将作为模型网络的输入。最后对帧画面的大小进行裁剪,输入的数据为$16 × 112 × 112 $
# ### C3D模型训练
# 接下来,我们将对C3D模型进行训练,训练过程分为:数据预处理以及模型训练。在此次训练中,我们使用的数据集为UCF-101,由于C3D模型的输入是视频的每帧图片,因此我们需要对数据集的视频进行抽帧,也就是将视频转换为图片,然后将图片数据传入模型之中,进行训练。
#
# 在本案例中,我们随机抽取了UCF-101数据集的一部分进行训练的演示,感兴趣的同学可以下载完整的UCF-101数据集进行训练。
#
# [UCF-101下载](https://www.crcv.ucf.edu/data/UCF101.php)
#
# 数据集存储在目录` dataset_subset`下
#
# 如下代码是使用cv2库进行视频文件到图片文件的转换
import cv2
import os
# 视频数据集存储位置
video_path = './dataset_subset/'
# 生成的图像数据集存储位置
save_path = './dataset/'
# 如果文件路径不存在则创建路径
if not os.path.exists(save_path):
os.mkdir(save_path)
# 获取动作列表
action_list = os.listdir(video_path)
# 遍历所有动作
for action in action_list:
if action.startswith(".")==False:
if not os.path.exists(save_path+action):
os.mkdir(save_path+action)
video_list = os.listdir(video_path+action)
# 遍历所有视频
for video in video_list:
prefix = video.split('.')[0]
if not os.path.exists(os.path.join(save_path, action, prefix)):
os.mkdir(os.path.join(save_path, action, prefix))
save_name = os.path.join(save_path, action, prefix) + '/'
video_name = video_path+action+'/'+video
# 读取视频文件
# cap为视频的帧
cap = cv2.VideoCapture(video_name)
# fps为帧率
fps = int(cap.get(cv2.CAP_PROP_FRAME_COUNT))
fps_count = 0
for i in range(fps):
ret, frame = cap.read()
if ret:
# 将帧画面写入图片文件中
cv2.imwrite(save_name+str(10000+fps_count)+'.jpg',frame)
fps_count += 1
# 此时,视频逐帧转换成的图片数据已经存储起来,为模型训练做准备。
#
# ### 模型训练
# 首先,我们构建模型结构。
#
# C3D模型结构我们之前已经介绍过,这里我们通过`keras`提供的Conv3D,MaxPool3D,ZeroPadding3D等函数进行模型的搭建。
# +
from keras.layers import Dense,Dropout,Conv3D,Input,MaxPool3D,Flatten,Activation, ZeroPadding3D
from keras.regularizers import l2
from keras.models import Model, Sequential
# 输入数据为 112×112 的图片,16帧, 3通道
input_shape = (112,112,16,3)
# 权重衰减率
weight_decay = 0.005
# 类型数量,我们使用UCF-101 为数据集,所以为101
nb_classes = 101
# 构建模型结构
inputs = Input(input_shape)
x = Conv3D(64,(3,3,3),strides=(1,1,1),padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(inputs)
x = MaxPool3D((2,2,1),strides=(2,2,1),padding='same')(x)
x = Conv3D(128,(3,3,3),strides=(1,1,1),padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = MaxPool3D((2,2,2),strides=(2,2,2),padding='same')(x)
x = Conv3D(128,(3,3,3),strides=(1,1,1),padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = MaxPool3D((2,2,2),strides=(2,2,2),padding='same')(x)
x = Conv3D(256,(3,3,3),strides=(1,1,1),padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = MaxPool3D((2,2,2),strides=(2,2,2),padding='same')(x)
x = Conv3D(256, (3, 3, 3), strides=(1, 1, 1), padding='same',
activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = MaxPool3D((2, 2, 2), strides=(2, 2, 2), padding='same')(x)
x = Flatten()(x)
x = Dense(2048,activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = Dropout(0.5)(x)
x = Dense(2048,activation='relu',kernel_regularizer=l2(weight_decay))(x)
x = Dropout(0.5)(x)
x = Dense(nb_classes,kernel_regularizer=l2(weight_decay))(x)
x = Activation('softmax')(x)
model = Model(inputs, x)
# -
# 通过keras提供的`summary()`方法,打印模型结构。可以看到模型的层构建以及各层的输入输出情况。
model.summary()
# 通过keras的`input`方法可以查看模型的输入形状,shape分别为`( batch size, width, height, frames, channels) ` 。
model.input
# 可以看到模型的数据处理的维度与图像处理模型有一些差别,多了frames维度,体现出时序关系在视频分析中的影响。
#
# 接下来,我们开始将图片文件转为训练需要的数据形式。
# +
# 引用必要的库
from keras.optimizers import SGD,Adam
from keras.utils import np_utils
import numpy as np
import random
import cv2
import matplotlib.pyplot as plt
# 自定义callbacks
from schedules import onetenth_4_8_12
# -
# 参数定义
img_path = save_path # 图片文件存储位置
results_path = './results' # 训练结果保存位置
if not os.path.exists(results_path):
os.mkdir(results_path)
# 数据集划分,随机抽取4/5 作为训练集,其余为验证集。将文件信息分别存储在`train_list`和`test_list`中,为训练做准备。
cates = os.listdir(img_path)
train_list = []
test_list = []
# 遍历所有的动作类型
for cate in cates:
videos = os.listdir(os.path.join(img_path, cate))
length = len(videos)//5
# 训练集大小,随机取视频文件加入训练集
train= random.sample(videos, length*4)
train_list.extend(train)
# 将余下的视频加入测试集
for video in videos:
if video not in train:
test_list.append(video)
print("训练集为:")
print( train_list)
print("共%d 个视频\n"%(len(train_list)))
print("验证集为:")
print(test_list)
print("共%d 个视频"%(len(test_list)))
# 接下来开始进行模型的训练。
#
# 首先定义数据读取方法。方法`process_data`中读取一个batch的数据,包含16帧的图片信息的数据,以及数据的标注信息。在读取图片数据时,对图片进行随机裁剪和翻转操作以完成数据增广。
def process_data(img_path, file_list,batch_size=16,train=True):
batch = np.zeros((batch_size,16,112,112,3),dtype='float32')
labels = np.zeros(batch_size,dtype='int')
cate_list = os.listdir(img_path)
def read_classes():
path = "./classInd.txt"
with open(path, "r+") as f:
lines = f.readlines()
classes = {}
for line in lines:
c_id = line.split()[0]
c_name = line.split()[1]
classes[c_name] =c_id
return classes
classes_dict = read_classes()
for file in file_list:
cate = file.split("_")[1]
img_list = os.listdir(os.path.join(img_path, cate, file))
img_list.sort()
batch_img = []
for i in range(batch_size):
path = os.path.join(img_path, cate, file)
label = int(classes_dict[cate])-1
symbol = len(img_list)//16
if train:
# 随机进行裁剪
crop_x = random.randint(0, 15)
crop_y = random.randint(0, 58)
# 随机进行翻转
is_flip = random.randint(0, 1)
# 以16 帧为单位
for j in range(16):
img = img_list[symbol + j]
image = cv2.imread( path + '/' + img)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = cv2.resize(image, (171, 128))
if is_flip == 1:
image = cv2.flip(image, 1)
batch[i][j][:][:][:] = image[crop_x:crop_x + 112, crop_y:crop_y + 112, :]
symbol-=1
if symbol<0:
break
labels[i] = label
else:
for j in range(16):
img = img_list[symbol + j]
image = cv2.imread( path + '/' + img)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = cv2.resize(image, (171, 128))
batch[i][j][:][:][:] = image[8:120, 30:142, :]
symbol-=1
if symbol<0:
break
labels[i] = label
return batch, labels
# +
batch, labels = process_data(img_path, train_list)
print("每个batch的形状为:%s"%(str(batch.shape)))
print("每个label的形状为:%s"%(str(labels.shape)))
# -
# 定义data generator, 将数据批次传入训练函数中。
def generator_train_batch(train_list, batch_size, num_classes, img_path):
while True:
# 读取一个batch的数据
x_train, x_labels = process_data(img_path, train_list, batch_size=16,train=True)
x = preprocess(x_train)
# 形成input要求的数据格式
y = np_utils.to_categorical(np.array(x_labels), num_classes)
x = np.transpose(x, (0,2,3,1,4))
yield x, y
def generator_val_batch(test_list, batch_size, num_classes, img_path):
while True:
# 读取一个batch的数据
y_test,y_labels = process_data(img_path, train_list, batch_size=16,train=False)
x = preprocess(y_test)
# 形成input要求的数据格式
x = np.transpose(x,(0,2,3,1,4))
y = np_utils.to_categorical(np.array(y_labels), num_classes)
yield x, y
# 定义方法`preprocess`, 对函数的输入数据进行图像的标准化处理。
def preprocess(inputs):
inputs[..., 0] -= 99.9
inputs[..., 1] -= 92.1
inputs[..., 2] -= 82.6
inputs[..., 0] /= 65.8
inputs[..., 1] /= 62.3
inputs[..., 2] /= 60.3
return inputs
# 训练一个epoch大约需4分钟
# 类别数量
num_classes = 101
# batch大小
batch_size = 4
# epoch数量
epochs = 1
# 学习率大小
lr = 0.005
# 优化器定义
sgd = SGD(lr=lr, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['accuracy'])
# 开始训练
history = model.fit_generator(generator_train_batch(train_list, batch_size, num_classes,img_path),
steps_per_epoch= len(train_list) // batch_size,
epochs=epochs,
callbacks=[onetenth_4_8_12(lr)],
validation_data=generator_val_batch(test_list, batch_size,num_classes,img_path),
validation_steps= len(test_list) // batch_size,
verbose=1)
# 对训练结果进行保存
model.save_weights(os.path.join(results_path, 'weights_c3d.h5'))
# ## 模型测试
# 接下来我们将训练之后得到的模型进行测试。随机在UCF-101中选择一个视频文件作为测试数据,然后对视频进行取帧,每16帧画面传入模型进行一次动作预测,并且将动作预测以及预测百分比打印在画面中并进行视频播放。
# 首先,引入相关的库。
from IPython.display import clear_output, Image, display, HTML
import time
import cv2
import base64
import numpy as np
# 构建模型结构并且加载权重。
from models import c3d_model
model = c3d_model()
model.load_weights(os.path.join(results_path, 'weights_c3d.h5'), by_name=True) # 加载刚训练的模型
# 定义函数arrayshow,进行图片变量的编码格式转换。
def arrayShow(img):
_,ret = cv2.imencode('.jpg', img)
return Image(data=ret)
# 进行视频的预处理以及预测,将预测结果打印到画面中,最后进行播放。
# 加载所有的类别和编号
with open('./ucfTrainTestlist/classInd.txt', 'r') as f:
class_names = f.readlines()
f.close()
# 读取视频文件
video = './videos/v_Punch_g03_c01.avi'
cap = cv2.VideoCapture(video)
clip = []
# 将视频画面传入模型
while True:
try:
clear_output(wait=True)
ret, frame = cap.read()
if ret:
tmp = cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)
clip.append(cv2.resize(tmp, (171, 128)))
# 每16帧进行一次预测
if len(clip) == 16:
inputs = np.array(clip).astype(np.float32)
inputs = np.expand_dims(inputs, axis=0)
inputs[..., 0] -= 99.9
inputs[..., 1] -= 92.1
inputs[..., 2] -= 82.6
inputs[..., 0] /= 65.8
inputs[..., 1] /= 62.3
inputs[..., 2] /= 60.3
inputs = inputs[:,:,8:120,30:142,:]
inputs = np.transpose(inputs, (0, 2, 3, 1, 4))
# 获得预测结果
pred = model.predict(inputs)
label = np.argmax(pred[0])
# 将预测结果绘制到画面中
cv2.putText(frame, class_names[label].split(' ')[-1].strip(), (20, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.6,
(0, 0, 255), 1)
cv2.putText(frame, "prob: %.4f" % pred[0][label], (20, 40),
cv2.FONT_HERSHEY_SIMPLEX, 0.6,
(0, 0, 255), 1)
clip.pop(0)
# 播放预测后的视频
lines, columns, _ = frame.shape
frame = cv2.resize(frame, (int(columns), int(lines)))
img = arrayShow(frame)
display(img)
time.sleep(0.02)
else:
break
except:
print(0)
cap.release()
# ## I3D 模型
# 在之前我们简单介绍了I3D模型,[I3D官方github库](https://github.com/deepmind/kinetics-i3d)提供了在Kinetics上预训练的模型和预测代码,接下来我们将体验I3D模型如何对视频进行预测。
# 首先,引入相关的包
# +
import numpy as np
import tensorflow as tf
import i3d
# -
# 进行参数的定义
# +
# 输入图片大小
_IMAGE_SIZE = 224
# 视频的帧数
_SAMPLE_VIDEO_FRAMES = 79
# 输入数据包括两部分:RGB和光流
# RGB和光流数据已经经过提前计算
_SAMPLE_PATHS = {
'rgb': 'data/v_CricketShot_g04_c01_rgb.npy',
'flow': 'data/v_CricketShot_g04_c01_flow.npy',
}
# 提供了多种可以选择的预训练权重
# 其中,imagenet系列模型从ImageNet的2D权重中拓展而来,其余为视频数据下的预训练权重
_CHECKPOINT_PATHS = {
'rgb': 'data/checkpoints/rgb_scratch/model.ckpt',
'flow': 'data/checkpoints/flow_scratch/model.ckpt',
'rgb_imagenet': 'data/checkpoints/rgb_imagenet/model.ckpt',
'flow_imagenet': 'data/checkpoints/flow_imagenet/model.ckpt',
}
# 记录类别文件
_LABEL_MAP_PATH = 'data/label_map.txt'
# 类别数量为400
NUM_CLASSES = 400
# -
# 定义参数:
# - imagenet_pretrained :如果为`True`,则调用预训练权重,如果为`False`,则调用ImageNet转成的权重
imagenet_pretrained = True
# 加载动作类型
kinetics_classes = [x.strip() for x in open(_LABEL_MAP_PATH)]
tf.logging.set_verbosity(tf.logging.INFO)
# 构建RGB部分模型
# +
rgb_input = tf.placeholder(tf.float32, shape=(1, _SAMPLE_VIDEO_FRAMES, _IMAGE_SIZE, _IMAGE_SIZE, 3))
with tf.variable_scope('RGB', reuse=tf.AUTO_REUSE):
rgb_model = i3d.InceptionI3d(NUM_CLASSES, spatial_squeeze=True, final_endpoint='Logits')
rgb_logits, _ = rgb_model(rgb_input, is_training=False, dropout_keep_prob=1.0)
rgb_variable_map = {}
for variable in tf.global_variables():
if variable.name.split('/')[0] == 'RGB':
rgb_variable_map[variable.name.replace(':0', '')] = variable
rgb_saver = tf.train.Saver(var_list=rgb_variable_map, reshape=True)
# -
# 构建光流部分模型
# +
flow_input = tf.placeholder(tf.float32,shape=(1, _SAMPLE_VIDEO_FRAMES, _IMAGE_SIZE, _IMAGE_SIZE, 2))
with tf.variable_scope('Flow', reuse=tf.AUTO_REUSE):
flow_model = i3d.InceptionI3d(NUM_CLASSES, spatial_squeeze=True, final_endpoint='Logits')
flow_logits, _ = flow_model(flow_input, is_training=False, dropout_keep_prob=1.0)
flow_variable_map = {}
for variable in tf.global_variables():
if variable.name.split('/')[0] == 'Flow':
flow_variable_map[variable.name.replace(':0', '')] = variable
flow_saver = tf.train.Saver(var_list=flow_variable_map, reshape=True)
# -
# 将模型联合,成为完整的I3D模型
model_logits = rgb_logits + flow_logits
model_predictions = tf.nn.softmax(model_logits)
# 开始模型预测,获得视频动作预测结果。
# 预测数据为开篇提供的RGB和光流数据:
#
# 
#
# 
with tf.Session() as sess:
feed_dict = {}
if imagenet_pretrained:
rgb_saver.restore(sess, _CHECKPOINT_PATHS['rgb_imagenet']) # 加载rgb流的模型
else:
rgb_saver.restore(sess, _CHECKPOINT_PATHS['rgb'])
tf.logging.info('RGB checkpoint restored')
if imagenet_pretrained:
flow_saver.restore(sess, _CHECKPOINT_PATHS['flow_imagenet']) # 加载flow流的模型
else:
flow_saver.restore(sess, _CHECKPOINT_PATHS['flow'])
tf.logging.info('Flow checkpoint restored')
start_time = time.time()
rgb_sample = np.load(_SAMPLE_PATHS['rgb']) # 加载rgb流的输入数据
tf.logging.info('RGB data loaded, shape=%s', str(rgb_sample.shape))
feed_dict[rgb_input] = rgb_sample
flow_sample = np.load(_SAMPLE_PATHS['flow']) # 加载flow流的输入数据
tf.logging.info('Flow data loaded, shape=%s', str(flow_sample.shape))
feed_dict[flow_input] = flow_sample
out_logits, out_predictions = sess.run(
[model_logits, model_predictions],
feed_dict=feed_dict)
out_logits = out_logits[0]
out_predictions = out_predictions[0]
sorted_indices = np.argsort(out_predictions)[::-1]
print('Inference time in sec: %.3f' % float(time.time() - start_time))
print('Norm of logits: %f' % np.linalg.norm(out_logits))
print('\nTop classes and probabilities')
for index in sorted_indices[:20]:
print(out_predictions[index], out_logits[index], kinetics_classes[index])
| notebook/DL_video_action_recognition/action_recognition.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.3
# language: julia
# name: julia-0.6
# ---
using Revise
using JuMIT
using Plots
model = JuMIT.Gallery.Seismic(:acou_homo1);
acqgeom = JuMIT.Gallery.Geom(model.mgrid,:xwell);
tgrid = JuMIT.Gallery.M1D(:acou_homo1);
wav = Signals.Wavelets.ricker(10.0, tgrid, tpeak=0.25, );
# source wavelet for modelling
acqsrc = JuMIT.Acquisition.Src_fixed(acqgeom.nss,1,[:P],wav,tgrid);
vp0=mean(JuMIT.Models.χ(model.χvp,model.vp0,-1))
ρ0=mean(JuMIT.Models.χ(model.χρ,model.ρ0,-1))
rec1 = JuMIT.Analytic.mod(vp0=vp0, model_pert=model, ρ0=ρ0, acqgeom=acqgeom, acqsrc=acqsrc, tgridmod=tgrid, src_flag=2)
pa=JuMIT.Fdtd.Param(npw=1,model=model, acqgeom=[acqgeom], acqsrc=[acqsrc], sflags=[2], rflags=[1], tgridmod=tgrid, verbose=true);
pab=JuMIT.Fdtd.Param(npw=2,model=model, acqgeom=[acqgeom, acqgeom], acqsrc=[acqsrc, acqsrc], sflags=[2, 0], rflags=[1, 1], tgridmod=tgrid, verbose=true, born_flag=true);
@time JuMIT.Fdtd.mod!(pab);
# +
# least-squares misfit
paerr=JuMIT.Data.P_misfit(rec1, pa.c.data[1])
err = JuMIT.Data.func_grad!(paerr)
# normalization
error = err[1]
# desired accuracy?
@test error<1e-2
println(error)
# -
plot(rec1.d[1,1])
plot!(pab.c.data[2].d[1,1])
| Modeling/Fdtd_accuracy.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// + dotnet_interactive={"language": "fsharp"}
#load "../include/MathDev.fsx"
open System
open System.Linq
open Sylvester
// + dotnet_interactive={"language": "fsharp"}
let dice = seq {1..6} |> Seq
dice
// + dotnet_interactive={"language": "csharp"}
let outcomes = (dice * dice)
outcomes.
// + dotnet_interactive={"language": "csharp"}
let s = dice.Prod
s |> Util.Table
// + dotnet_interactive={"language": "csharp"}
dice.AsSigmaAlgebra
// + dotnet_interactive={"language": "csharp"}
dice.AsSigmaAlgebra.Count()
// + dotnet_interactive={"language": "csharp"}
| examples/math/Probability.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# <div id="body">
# <center>
# <a href="06 Remotes in GitHub.ipynb"> <font size="6"> < </font></a>
# <a href="index.ipynb"> <font size="6"> Version Control with Git </font> </a>
# <a href="08 Conflicts.ipynb"> <font size="6"> > </font></a>
# </center>
# </div>
# # Collaborating
# ## Questions
#
# * How can I use version control to collaborate with other people?
#
#
# ## Objectives
#
# * Clone a remote repository.
# * Collaborate by pushing to a common repository.
# * Describe the basic collaborative workflow.
#
#
# For the next step, get into pairs. One person will be the “Owner” and the other will be the “Collaborator”. The goal is that the Collaborator add changes into the Owner’s repository. We will switch roles at the end, so both persons will play Owner and Collaborator.
# <blockquote class="note">
# <h2>Practicing By Yourself</h2>
#
# If you’re working through this lesson on your own, you can carry on by opening a second terminal window. This window will represent your partner, working on another computer. You won’t need to give anyone access on GitHub, because both ‘partners’ are you.
# </blockquote>
# The Owner needs to give the Collaborator access. On GitHub, click the settings button on the right, then select Collaborators, and enter your partner’s username.
#
# <img src="static/images/github-add-collaborators.png">
#
# o accept access to the Owner’s repo, the Collaborator needs to go to https://github.com/notifications. Once there she can accept access to the Owner’s repo.
#
# Next, the Collaborator needs to download a copy of the Owner’s repository to her machine. This is called “cloning a repo”. To clone the Owner’s repo into her Desktop folder, the Collaborator enters:
#
# ```
# git clone https://github.com/epinux/git_tutorial.git ~/Desktop/epinux-git_tutorial
# ```
#
# Replace ‘epinux’ with the Owner’s username.
#
# <img src="static/images/github-collaboration.svg">
git clone https://github.com/epinux/git_tutorial.git ~/Desktop/epinux-git_tutorial
# The Collaborator can now make a change in her clone of the Owner’s repository, exactly the same way as we’ve been doing before:
#
#
cd ~/Desktop/epinux-git_tutorial
touch newfile.txt
echo "# this is going to be my first contribution" > newfile.txt
cat newfile.txt
git add newfile.txt
git commit -m "Add notes about my first contribution"
# Then push the change to the Owner’s repository on GitHub:
# +
# git push origin master
# -
# **Note:** To run the command above you may need to issue username and password. To do so in a jupyter environment execute the command from a terminal.
#
# ```bash
# Enumerating objects: 4, done.
# Counting objects: 4, done.
# Delta compression using up to 4 threads.
# Compressing objects: 100% (2/2), done.
# Writing objects: 100% (3/3), 306 bytes, done.
# Total 3 (delta 0), reused 0 (delta 0)
# To https://github.com/epinux/git_tutorial.git
# b324f10..28773e7 master -> master
# ```
#
# Note that we didn’t have to create a remote called origin: Git uses this name by default when we clone a repository. (This is why origin was a sensible choice earlier when we were setting up remotes by hand.)
# <blockquote class="note">
# <h2>Some more about remotes</h2>
# In this episode and the previous one, our local repository has had a single “remote”, called `origin`. A remote is a copy of the repository that is hosted somewhere else, that we can push to and pull from, and there’s no reason that you have to work with only one. For example, on some large projects you might have your own copy in your own GitHub account (you’d probably call this `origin`) and also the main “upstream” project repository (let’s call this `upstream` for the sake of examples). You would pull from `upstream` from time to time to get the latest updates that other people have committed.
#
# Remember that the name you give to a remote only exists locally. It’s an alias that you choose - whether `origin`, or `upstream`, or `fred` - and not something intrinstic to the remote repository.
#
# The `git remote` family of commands is used to set up and alter the remotes associated with a repository. Here are some of the most useful ones:
#
# * `git remote -v` lists all the remotes that are configured (we already used this in the last episode)
# * `git remote add [name] [url]` is used to add a new remote
# * `git remote remove [name]` removes a remote. Note that it doesn’t affect the remote repository at all - it just removes the link to it from the local repo.
# * `git remote set-url [name] [newurl]` changes the URL that is associated with the remote. This is useful if it has moved, e.g. to a different GitHub account, or from GitHub to a different hosting service. Or, if we made a typo when adding it!
# * `git remote rename [oldname] [newname]` changes the local alias by which a remote is known - its name. For example, one could use this to change `upstream` to fred.
#
# </blockquote>
#
# Take a look to the Owner’s repository on its GitHub website now (maybe you need to refresh your browser.) You should be able to see the new commit made by the Collaborator.
#
# To download the Collaborator’s changes from GitHub, the Owner now enters:
#
git pull origin master
# Now the three repositories (Owner’s local, Collaborator’s local, and Owner’s on GitHub) are back in sync.
# <blockquote class="note">
# <h2>A Basic Collaborative Workflow</h2>
#
#
# In practice, it is good to be sure that you have an updated version of the repository you are collaborating on, so you should git pull before making our changes. The basic collaborative workflow would be:
#
# * update your local repo with `git pull origin master`,
# * make your changes and stage them with `git add`,
# * commit your changes with `git commit -m`, and
# * upload the changes to GitHub with `git push origin master`
#
# It is better to make many commits with smaller changes rather than of one commit with massive changes: small commits are easier to read and review.
#
# </blockquote>
# <blockquote class="note">
# <h2>Review Changes</h2>
# The Owner pushed commits to the repository without giving any information to the Collaborator. How can the Collaborator find out what has changed with command line? And on GitHub?
#
# * On the command line, the Collaborator can use `git fetch origin master` to get the remote changes into the local repository, but without merging them. Then by running `git diff master origin/master` the Collaborator will see the changes output in the terminal.
#
# * On GitHub, the Collaborator can go to their own fork of the repository and look right above the light blue latest commit bar for a gray bar saying “This branch is 1 commit behind Our-Repository:master.” On the far right of that gray bar is a Compare icon and link. On the Compare page the Collaborator should change the base fork to their own repository, then click the link in the paragraph above to “compare across forks”, and finally change the head fork to the main repository. This will show all the commits that are different.
# </blockquote>
# <blockquote class="note">
# <h2>Comment Changes in GitHub</h2>
#
# The Collaborator has some questions about one line change made by the Owner and has some suggestions to propose.
#
# With GitHub, it is possible to comment the diff of a commit. Over the line of code to comment, a blue comment icon appears to open a comment window.
#
# The Collaborator posts its comments and suggestions using GitHub interface.
# </blockquote>
# <blockquote class="keypoints">
# <h2>Key Points</h2>
#
# * `git clone` copies a remote repository to create a local repository with a remote called `origin` automatically set up.
# </blockquote>
#
# <div id="body">
# <center>
# <a href="06 Remotes in GitHub.ipynb"> <font size="4"> < </font></a>
# <a href="index.ipynb"> <font size="4"> Version Control with Git </font> </a>
# <a href="08 Conflicts.ipynb"> <font size="4"> > </font></a>
# </center>
# </div>
| 07 Collaborating.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kernel CCA (KCCA)
# This algorithm runs KCCA on two views of data. The kernel implementations, parameter 'ktype', are linear, polynomial and gaussian. Polynomial kernel has two parameters: 'constant', 'degree'. Gaussian kernel has one parameter: 'sigma'.
#
# Useful information, like canonical correlations between transformed data and statistical tests for significance of these correlations can be computed using the get_stats() function of the KCCA object.
#
# When initializing KCCA, you can also initialize the following parameters: the number of canonical components 'n_components', the regularization parameter 'reg', the decomposition type 'decomposition', and the decomposition method 'method'. There are two decomposition types: 'full' and 'icd'. In some cases, ICD will run faster than the full decomposition at the cost of performance. The only method as of now is 'kettenring-like'.
#
# +
import numpy as np
import sys
sys.path.append("../../..")
from mvlearn.embed.kcca import KCCA
from mvlearn.plotting.plot import crossviews_plot
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import stats
import warnings
import matplotlib.cbook
warnings.filterwarnings("ignore",category=matplotlib.cbook.mplDeprecation)
# -
# Function creates Xs, a list of two views of data with a linear relationship, polynomial relationship (2nd degree) and a gaussian (sinusoidal) relationship.
def make_data(kernel, N):
# # # Define two latent variables (number of samples x 1)
latvar1 = np.random.randn(N,)
latvar2 = np.random.randn(N,)
# # # Define independent components for each dataset (number of observations x dataset dimensions)
indep1 = np.random.randn(N, 4)
indep2 = np.random.randn(N, 5)
if kernel == "linear":
x = 0.25*indep1 + 0.75*np.vstack((latvar1, latvar2, latvar1, latvar2)).T
y = 0.25*indep2 + 0.75*np.vstack((latvar1, latvar2, latvar1, latvar2, latvar1)).T
return [x,y]
elif kernel == "poly":
x = 0.25*indep1 + 0.75*np.vstack((latvar1**2, latvar2**2, latvar1**2, latvar2**2)).T
y = 0.25*indep2 + 0.75*np.vstack((latvar1, latvar2, latvar1, latvar2, latvar1)).T
return [x,y]
elif kernel == "gaussian":
t = np.random.uniform(-np.pi, np.pi, N)
e1 = np.random.normal(0, 0.05, (N,2))
e2 = np.random.normal(0, 0.05, (N,2))
x = np.zeros((N,2))
x[:,0] = t
x[:,1] = np.sin(3*t)
x += e1
y = np.zeros((N,2))
y[:,0] = np.exp(t/4)*np.cos(2*t)
y[:,1] = np.exp(t/4)*np.sin(2*t)
y += e2
return [x,y]
# ## Linear kernel implementation
# Here we show how KCCA with a linear kernel can uncover the highly correlated latent distribution of the 2 views which are related with a linear relationship, and then transform the data into that latent space. We use an 80-20, train-test data split to develop the embedding.
#
# Also, we use statistical tests (Wilk's Lambda) to check the significance of the canonical correlations.
# +
np.random.seed(1)
Xs = make_data('linear', 100)
Xs_train = [Xs[0][:80],Xs[1][:80]]
Xs_test = [Xs[0][80:],Xs[1][80:]]
kcca_l = KCCA(n_components = 4, reg = 0.01)
kcca_l.fit(Xs_train)
linearkcca = kcca_l.transform(Xs_test)
# -
# ### Original Data Plotted
crossviews_plot(Xs, ax_ticks=False, ax_labels=True, equal_axes=True)
# ### Transformed Data Plotted
crossviews_plot(linearkcca, ax_ticks=False, ax_labels=True, equal_axes=True)
# Now, we assess the canonical correlations achieved on the testing data, and the p-values for significance using a Wilk's Lambda test
# +
stats = kcca_l.get_stats()
print("Below are the canonical correlations and the p-values of a Wilk's Lambda test for each components:")
print(stats['r'])
print(stats['pF'])
# -
# ## Polynomial kernel implementation
# Here we show how KCCA with a polynomial kernel can uncover the highly correlated latent distribution of the 2 views which are related with a polynomial relationship, and then transform the data into that latent space.
Xsp = make_data("poly", 150)
kcca_p = KCCA(ktype ="poly", degree = 2.0, n_components = 4, reg=0.001)
polykcca = kcca_p.fit_transform(Xsp)
# ### Original Data Plotted
crossviews_plot(Xsp, ax_ticks=False, ax_labels=True, equal_axes=True)
# ### Transformed Data Plotted
crossviews_plot(polykcca, ax_ticks=False, ax_labels=True, equal_axes=True)
# Now, we assess the canonical correlations achieved on the testing data
# +
stats = kcca_p.get_stats()
print("Below are the canonical correlations for each components:")
print(stats['r'])
# -
# ## Gaussian Kernel Implementation
# Here we show how KCCA with a gaussian kernel can uncover the highly correlated latent distribution of the 2 views which are related with a sinusoidal relationship, and then transform the data into that latent space.
Xsg = make_data("gaussian", 100)
Xsg_train = [Xsg[0][:20],Xsg[1][:20]]
Xsg_test = [Xsg[0][20:],Xsg[1][20:]]
kcca_g = KCCA(ktype ="gaussian", sigma = 1.0, n_components = 2, reg = 0.01)
kcca_g.fit(Xsg)
gausskcca = kcca_g.transform(Xsg)
# ### Original Data Plotted
crossviews_plot(Xsg, ax_ticks=False, ax_labels=True, equal_axes=True)
# ### Transformed Data Plotted
crossviews_plot(gausskcca, ax_ticks=False, ax_labels=True, equal_axes=True)
# Now, we assess the canonical correlations achieved on the testing data
# +
stats = kcca_g.get_stats()
print("Below are the canonical correlations for each components:")
print(stats['r'])
| docs/tutorials/embed/kcca_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Reinforcement Learning <em> in Action </em>
# ## Ch. 4 - Policy Gradients
import gym
import numpy as np
import torch
from matplotlib import pylab as plt
# #### Helper functions
def running_mean(x, N=50):
cumsum = np.cumsum(np.insert(x, 0, 0))
return (cumsum[N:] - cumsum[:-N]) / float(N)
# #### Defining Network
# +
l1 = 4
l2 = 150
l3 = 2
model = torch.nn.Sequential(
torch.nn.Linear(l1, l2),
torch.nn.LeakyReLU(),
torch.nn.Linear(l2, l3),
torch.nn.Softmax()
)
learning_rate = 0.0009
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# -
# #### Objective Function
def loss_fn(preds, r):
# pred is output from neural network, a is action index
# r is return (sum of rewards to end of episode), d is discount factor
return -torch.sum(r * torch.log(preds)) # element-wise multipliy, then sum
# #### Training Loop
# +
env = gym.make('CartPole-v0')
MAX_DUR = 200
MAX_EPISODES = 500
gamma_ = 0.99
time_steps = []
for episode in range(MAX_EPISODES):
curr_state = env.reset()
done = False
transitions = [] # list of state, action, rewards
for t in range(MAX_DUR): #while in episode
act_prob = model(torch.from_numpy(curr_state).float())
action = np.random.choice(np.array([0,1]), p=act_prob.data.numpy())
prev_state = curr_state
curr_state, reward, done, info = env.step(action)
transitions.append((prev_state, action, reward))
if done:
break
# Optimize policy network with full episode
ep_len = len(transitions) # episode length
time_steps.append(ep_len)
preds = torch.zeros(ep_len)
discounted_rewards = torch.zeros(ep_len)
for i in range(ep_len): #for each step in episode
discount = 1
future_reward = 0
# discount rewards
for i2 in range(i, ep_len):
future_reward += transitions[i2][2] * discount
discount = discount * gamma_
discounted_rewards[i] = future_reward
state, action, _ = transitions[i]
pred = model(torch.from_numpy(state).float())
preds[i] = pred[action]
loss = loss_fn(preds, discounted_rewards)
optimizer.zero_grad()
loss.backward()
optimizer.step()
env.close()
# -
plt.figure(figsize=(10,7))
plt.ylabel("Duration")
plt.xlabel("Episode")
plt.plot(running_mean(time_steps, 50), color='green')
| old_but_more_detailed/Ch4_PolicyGradients.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 2 Required Coding Activity
# Introduction to Python Unit 1
#
# This is an activity based on code similar to the Jupyter Notebook **`Practice_MOD02_1-3_IntroPy.ipynb`** which you may have completed.
#
# | Some Assignment Requirements |
# |:-------------------------------|
# | **NOTE:** This program requires a **function** be defined, created and called. The call will send values based on user input. The function call must capture a `return` value that is used in print output. The function will have parameters and `return` a string and should otherwise use code syntax covered in module 2. |
#
# ## Program: fishstore()
# create and test fishstore()
# - **fishstore() takes 2 string arguments: fish & price**
# - **fishstore returns a string in sentence form**
# - **gather input for fish_entry and price_entry to use in calling fishstore()**
# - **print the return value of fishstore()**
# >example of output: **`Fish Type: Guppy costs $1`**
# +
# [ ] create, call and test fishstore() function
# then PASTE THIS CODE into edX
# -
# ### Need assignment tips and clarification?
# See the video on the "End of Module coding assignment > Module 2 Required Code Description" course page on [edX](https://courses.edx.org/courses/course-v1:Microsoft+DEV236x+4T2017/course)
#
# # Important: [How to submit the code in edX by pasting](https://courses.edx.org/courses/course-v1:Microsoft+DEV236x+1T2017/wiki/Microsoft.DEV236x.3T2018/paste-code-end-module-coding-assignments/)
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Required_Code_MOD2_IntroPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from bokeh.plotting import output_file, show, output_notebook
from bokeh.models import GeoJSONDataSource
from bokeh.plotting import figure
from bokeh.sampledata.sample_geojson import geojson
geo_source = GeoJSONDataSource(geojson=geojson)
p = figure()
p.circle(x='x', y='y', alpha=0.9, source=geo_source)
output_notebook()
show(p)
# -
| notebooks/bokeh-visualizations/geojson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/seel-channel/AMPD_Mask_RCNN/blob/master/Train_Mask_RCNN_(AMPD).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="rGWTPWBIx370"
# ## **1. Installation**
#
# Load your dataset
# + id="Md87Hxgtn6zi"
# %tensorflow_version 1.x
# !pip install --upgrade h5py==2.10.0
# !git clone https://github.com/seel-channel/AMPD_Mask_RCNN
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/"} id="O0LSXkUGsYMS" outputId="dace138c-e28e-4951-dcc5-65a988e28277"
import sys
sys.path.append("/content/AMPD_Mask_RCNN")
from train_mask_rcnn import *
# + id="JvM9sdw3Vou_" colab={"base_uri": "https://localhost:8080/"} outputId="e63c2751-b7be-47ef-e693-00cf042a6857"
# !nvidia-smi
# + [markdown] id="Omb3Yl6ABqiJ"
# ## **2. Image Dataset**
#
# Load your annotated dataset
#
# + id="MnRU9zVkRktW" colab={"base_uri": "https://localhost:8080/"} outputId="203e1fc2-f046-4d26-c8e8-ca2a7977bcef"
from google.colab import drive
drive.mount('/content/gdrive')
# + id="IlNYqGhvqb_p" colab={"base_uri": "https://localhost:8080/"} outputId="b6377426-2156-41bc-b575-a48e787a34c8"
# Extract Images
drive_folder = "/content/gdrive/MyDrive/ampd/"
cache_folder = "/content/dataset/"
test_folder = cache_folder + "test/"
train_folder = cache_folder + "train/"
images_folder = "images/"
generated_folder = "generated/"
annotations_filename = "coco_annotations.json"
train_annotations_path = train_folder + annotations_filename
test_annotations_path = test_folder + annotations_filename
print("Extracting: train")
extract_images(drive_folder + "dataset_train.zip", train_folder)
print("Extracting: test")
extract_images(drive_folder + "dataset_test.zip", test_folder)
# + colab={"base_uri": "https://localhost:8080/"} id="J520fnwgUMTI" outputId="5411eccf-f570-4115-e76c-3e29f9774e3a"
import json
def open_json(path: str):
with open(path) as f:
return json.loads(f.read())
def save_json(dict: dict, path: str):
with open(path, 'w+') as f:
json.dump(dict, f)
def remove_unused_images(annotations_path):
coco_json = open_json(annotations_path)
used_images = []
images = []
for annotation in coco_json['annotations']:
image_id = annotation['image_id']
if not image_id in used_images:
used_images.append(image_id)
for image in coco_json['images']:
image_id = image['id']
if image_id in used_images:
images.append(image)
coco_json['images'] = images
save_json(coco_json, annotations_path)
# Ignore images without annotations
print("Removing train unused images...")
remove_unused_images(train_annotations_path)
print("Removing test unused images...")
remove_unused_images(test_annotations_path)
# + id="MnW8ETPKzqFT" colab={"base_uri": "https://localhost:8080/"} outputId="a06d3893-48ad-4637-cba3-5da04801a65e"
dataset_train = load_image_dataset(train_annotations_path, train_folder, "train")
dataset_val = load_image_dataset(test_annotations_path, test_folder, "val")
class_number = dataset_train.count_classes()
print('Train: %d' % len(dataset_train.image_ids))
print('Validation: %d' % len(dataset_val.image_ids))
print("Classes: {}".format(class_number))
# + id="umeaqvVeBqiL" colab={"base_uri": "https://localhost:8080/", "height": 617} outputId="bbb4ab64-5fba-44fe-990f-6214aa93ea1f"
# Load image samples
display_image_samples(dataset_train)
# + [markdown] id="Z9k3Wm0_BqiN"
# ##**3. Training**
#
# Train Mask RCNN on your custom Dataset.
# + id="axkqWaZ7z_4p" colab={"base_uri": "https://localhost:8080/"} outputId="df9e88be-4ce6-4409-8eeb-887c0142c833"
# Load Configuration
import os
def make_folders(folders: list, remove=False):
for path in folders:
if not os.path.exists(path):
os.makedirs(path)
elif(remove):
os.remove(path)
model_dir = drive_folder + "pretrained/"
make_folders([model_dir])
config = CustomConfig(class_number)
model = load_training_model(config, model_dir)
# + id="SyzLXzF5BqiN" colab={"base_uri": "https://localhost:8080/"} outputId="09bf5c5d-0ec7-4f0e-9194-32de525b79e7"
# Start Training. This operation might take a long time.
train_head(model, dataset_train, dataset_train, config, epochs=10)
# + [markdown] id="6npLKIL3BqiO"
# ## **4. Detection (test your model on a random image)**
# + id="lUwXQ6h7BqiO" colab={"base_uri": "https://localhost:8080/"} outputId="98572e62-6b09-40e3-da9e-2583f38da4f1"
# Load the latest trained model will be loaded
test_model, inference_config = load_test_model(class_number, model_dir)
# + id="H8uzE5U3BqiP" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="9d890a36-8541-48b2-a6b1-c7ec84ab1c14"
# Test on a random image
test_random_image(test_model, dataset_val, inference_config)
| Train_Mask_RCNN_(AMPD).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from eppy import modeleditor
from eppy.modeleditor import IDF
from matplotlib import pyplot as plt
import os
import pandas as pd
import numpy as np
from besos import eppy_funcs as ef
from besos import sampling
from besos.evaluator import EvaluatorEP
from besos.parameters import RangeParameter, FieldSelector, FilterSelector, Parameter, expand_plist, wwr, CategoryParameter, GenericSelector
from besos.problem import EPProblem
import matplotlib.pyplot as plt
output_path="./output/simulation_monthly_2017/"
iddfile='./energyplus/EnergyPlus-9-0-1/Energy+.idd'
fname = './input/on_double.idf'
weather='./input/ITA_Torino_160590_IWEC.epw'
IDF.setiddname(iddfile)
idf = IDF(fname,weather)
# +
zones = idf.idfobjects["ZONE"]
total_area=0
for i in range(len(zones)):
zone = zones[i]
area = modeleditor.zonearea(idf, zone.Name)
total_area=total_area+area
print("total one area = %s" % (total_area, ))
# -
samples_temp =[]
samples_temp.append({'Orientation': 270,
'Insulation Thickness': 0.35,
'Window to Wall Ratio': 0.15,})
samples = pd.DataFrame.from_dict(samples_temp)
samples
# +
building = ef.get_building(fname)
insulation = FieldSelector(class_name='Material', object_name='MW Glass Wool (rolls)_O.1445',
field_name='Thickness')
insulationPR=Parameter(selector=insulation,name='Insulation Thickness')
window_to_wall = wwr(CategoryParameter([0.15]))
orientation = FieldSelector(class_name='Building', field_name='North Axis')
orientationPR = Parameter(selector=orientation, value_descriptor=CategoryParameter(options=[270]),
name='Orientation')
parameters = [orientationPR , window_to_wall, insulationPR]
objectives = ['Electricity:Facility','DistrictHeating:Facility','DistrictCooling:Facility'] # these get made into `MeterReader` or `VariableReader`
problem=EPProblem(parameters, objectives) # problem = parameters + objectives
# -
evaluator = EvaluatorEP(problem, building, out_dir=output_path, err_dir=output_path,
epw=weather) # evaluator = problem + building
#simulation 2019
run_period = idf.idfobjects['RunPeriod'][0]
run_period.Begin_Year = 2017
run_period.End_Year = 2017
run_period
idf.idfobjects['RunPeriod'][0]
#monthly simulation
for i in range(len(idf.idfobjects['OUTPUT:VARIABLE'])):
idf.idfobjects['OUTPUT:VARIABLE'][i].Reporting_Frequency='monthly'
for i in range(len(idf.idfobjects['OUTPUT:METER'])):
idf.idfobjects['OUTPUT:METER'][i].Reporting_Frequency='monthly'
idf.idfobjects['OUTPUT:ENVIRONMENTALIMPACTFACTORS'][0].Reporting_Frequency='monthly'
#Now we run the evaluator with the given parameters
result = evaluator([270,0.15,0.35])
values = dict(zip(objectives, result))
for key, value in values.items():
print(key, " :: ", "{0:.2f}".format(value/3.6e6), "kWh")
idf.run(readvars=True,output_directory=output_path,annual= True)
idf_data =pd.read_csv(output_path + 'eplusout.csv')
for i in idf_data.columns:
if 'Temperature' in i and 'Zone' in i:
print(i)
#getting required columns
columns=['Date/Time']
for i in idf_data.columns:
if ('Zone Operative Temperature' in i or 'District' in i or 'Electricity:Facility' in i or 'Drybulb' in i) and ('Monthly' in i):
columns=columns+[i]
columns
df_columns = {'Date/Time':'Date_Time',
'Environment:Site Outdoor Air Drybulb Temperature [C](Monthly)':'t_out',
'BLOCK1:ZONE3:Zone Operative Temperature [C](Monthly)':'t_in_ZONE3',
'BLOCK1:ZONE1:Zone Operative Temperature [C](Monthly)':'t_in_ZONE1',
'BLOCK1:ZONE4:Zone Operative Temperature [C](Monthly)':'t_in_ZONE4',
'BLOCK1:ZONE2:Zone Operative Temperature [C](Monthly)':'t_in_ZONE2',
'DistrictHeating:Facility [J](Monthly)':'power_heating',
'DistrictCooling:Facility [J](Monthly)':'power_cooling' ,
'Electricity:Facility [J](Monthly)':'power_electricity'}
idf_data=idf_data[columns]
data = idf_data.rename(columns =df_columns)
data
data['t_in'] = data[['t_in_ZONE3','t_in_ZONE1','t_in_ZONE4', 't_in_ZONE2']].mean(axis=1)
data=data.drop(['t_in_ZONE3', 't_in_ZONE1', 't_in_ZONE4','t_in_ZONE2'], axis = 1)
data
data['temp_diff'] =data['t_in'] - data['t_out']
#data['Date_Time'] = '2019/' + data['Date_Time'].str.strip()
#data['Date_Time'] = data['Date_Time'].str.replace('24:00:00','23:59:59')
data
#converting from J to kWh
data['power_heating'] /= 3.6e6
data['power_cooling'] /= 3.6e6
data['power_electricity'] /= 3.6e6
data
data['total_power'] = data['power_heating']+ data['power_cooling']
data = data[['Date_Time','t_in','t_out','temp_diff','power_heating','power_cooling','power_electricity','total_power']]
data.to_csv(path_or_buf=output_path+'simulation_data_monthly.csv',index=False)
data
y=data['power_electricity'].values
x=data['Date_Time'].values
y
y/total_area
fig, ax = plt.subplots(figsize=(20,10))
labels= data['Date_Time'].values
plt.xticks(np.arange(1, len(x)+1, 1.0))
ax.set_xticklabels(labels)
plt.xticks(rotation='vertical')
plt.xlabel('Months')
plt.ylabel('Electricity Consumption per m2 (kWh/m2)')
plt.title(' Annual Electricity Consumption per m2 in 2019')
plt.plot(y)
plt.show()
fig, ax = plt.subplots(figsize=(10,5))
labels= data['Date_Time'].values
plt.xticks(np.arange(1, len(x)+1, 1.0))
ax.set_xticklabels(labels)
plt.xticks(rotation='vertical')
plt.plot(data['power_cooling'].values/total_area)
plt.xlabel('Months')
plt.ylabel('Cooling Power per m2 (kWh/m2)')
plt.title('Monthly Cooling Power Consumption per m2 from in 2019')
plt.show()
fig, ax = plt.subplots(figsize=(10,5))
labels= data['Date_Time'].values
plt.xticks(np.arange(1, len(x)+1, 1.0))
ax.set_xticklabels(labels)
plt.xticks(rotation='vertical')
plt.plot(data['power_heating'].values/total_area)
plt.xlabel('Months')
plt.ylabel('Heating Power per m2(kWh/m2)')
plt.title('Monthly Heating Power Consumption per m2 from in 2019')
plt.show()
| 2-Simulations_monthly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # About the environment - "vector_cv_project"
# !which python
# !echo $PYTHONPATH
# !echo $LD_LIBRARY_PATH
# !echo $PATH
# +
# imports
import argparse
import logging
import time
from tqdm import tqdm
import numpy as np
import torch
from vector_cv_tools import datasets
from vector_cv_tools import transforms as T
from vector_cv_tools import utils
import albumentations
import torchvision
from torch.utils.data import DataLoader
torch.cuda.is_available()
# -
kinetics_annotation_path = "./datasets/kinetics/kinetics700/train.json"
kinetics_data_path = "./datasets/kinetics/train"
# # A basic, un-transformed kinetics dataset
#
# +
# define basic spatial and temporal transforms
base_spatial_transforms = T.ComposeVideoSpatialTransform([albumentations.ToFloat(max_value=255)])
base_temporal_transforms = T.ComposeVideoTemporalTransform([T.video_transforms.ToTensor()])
# create raw dataset
data_raw = datasets.KineticsDataset(
fps=10,
max_frames=128,
round_source_fps=False,
annotation_path = kinetics_annotation_path,
data_path = kinetics_data_path,
class_filter = ["push_up", "pull_ups"],
spatial_transforms=base_spatial_transforms,
temporal_transforms=base_temporal_transforms)
# -
labels = data_raw.metadata.labels
print("Looping through the dataset, {} labels, {} data points in total".
format(data_raw.num_classes, len(data_raw)))
for label, info in labels.items():
print("{:<40} ID: {} size: {} {}".
format(label, info["id"], len(info["indexes"]), len(info["indexes"])//20 * "|"))
data_point, label = data_raw[0]
print(data_point.shape)
print(label)
vid = (data_point.numpy() * 255).astype(np.uint8)
utils.create_GIF("raw_img.gif", vid)
# # A dataset with video transformations
# +
###############################################
##### NOW PRESENT TO YOU: VideoTransforms!!####
###############################################
# compatibility with others
transform1 = T.from_torchvision(
torchvision.transforms.ColorJitter())
transform2 = T.from_torchvision(
torchvision.transforms.functional.hflip)
transform3 = T.from_albumentation(
albumentations.VerticalFlip(p=1))
# Spatial: in-house
transform4 = T.RandomResizedSpatialCrop((280, 280), scale=(0, 1))
transform5 = T.RandomSpatialCrop((480, 480))
transform6 = T.RandomTemporalCrop(size=50, pad_if_needed=True, padding_mode="wrap")
transform7 = T.SampleEveryNthFrame(2)
transform8 = T.ToTensor()
spatial_transforms = base_spatial_transforms
# define temporal transforms
temporal_transforms = [transform1, transform2, transform3, transform4,
transform6, transform7, transform8]
temporal_transforms = T.ComposeVideoTemporalTransform(temporal_transforms)
print("Spatial transforms: \n{}".format(spatial_transforms))
print("Temporal transforms: \n{}".format(temporal_transforms))
# +
# create a dataset with transformations
data_transformed = datasets.KineticsDataset(
fps=10,
max_frames=128,
round_source_fps=False,
annotation_path = kinetics_annotation_path,
data_path = kinetics_data_path,
class_filter = ["push_up", "pull_ups"],
spatial_transforms=spatial_transforms,
temporal_transforms=temporal_transforms,)
# -
data_point, label = data_transformed[0]
print(data_point.shape)
print(label)
vid = (data_point.numpy() * 255).astype(np.uint8)
utils.create_GIF("transformed_img.gif", vid)
| examples/video/VideoTransformationDemo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="VBECACYMYNny" outputId="687227ce-54c9-4efb-9a61-a73acfdbff15"
# !pip install lightkurve
# + colab={"base_uri": "https://localhost:8080/"} id="8UZm2_6xXNt7" outputId="a301d552-e133-483d-951d-906834ebb0c0"
# !pip install exoplanet
# + colab={"base_uri": "https://localhost:8080/"} id="riIVV1fJXfF_" outputId="0a95d3f8-8ac2-4b26-cb77-2f440971a5f0"
import exoplanet
exoplanet.utils.docs_setup()
print(f"exoplanet.__version__ = '{exoplanet.__version__}'")
# + colab={"base_uri": "https://localhost:8080/", "height": 386} id="9EmVYe7LfHVk" outputId="f0b9910e-db51-4ebd-8efe-6a3e11a37714"
lc = lk.search_lightcurve('TIC 375506058', mission="TESS", sector=15).download()
lc.plot();
# + colab={"base_uri": "https://localhost:8080/", "height": 520} id="kT-rdAakXqee" outputId="0f2b3f6d-036e-43fe-f77c-7b046c198a7d"
import numpy as np
import lightkurve as lk
import matplotlib.pyplot as plt
from astropy.io import fits
lc_file = lk.search_lightcurve(
"TIC 375506058", sector=15, mission="TESS"
).download(quality_bitmask="hardest", flux_column="pdcsap_flux")
lc = lc_file.remove_nans().normalize().remove_outliers()
time = lc.time.value
flux = lc.flux
# For the purposes of this example, we'll discard some of the data
m = (lc.quality == 0) & (
np.random.default_rng(375506058).uniform(size=len(time)) < 0.3
)
with fits.open(lc_file.filename) as hdu:
hdr = hdu[1].header
texp = hdr["FRAMETIM"] * hdr["NUM_FRM"]
texp /= 60.0 * 60.0 * 24.0
ref_time = 0.5 * (np.min(time) + np.max(time))
x = np.ascontiguousarray(time[m] - ref_time, dtype=np.float64)
y = np.ascontiguousarray(1e3 * (flux[m] - 1.0), dtype=np.float64)
plt.plot(x, y, ".k")
plt.xlabel("time [days]")
plt.ylabel("relative flux [ppt]")
_ = plt.xlim(x.min(), x.max())
# + colab={"base_uri": "https://localhost:8080/"} id="xPO_oxdwfWAu" outputId="a8cf54c6-5cef-4f7b-a17e-c18e89cbc8c4"
pg = lc.normalize(unit='ppm').to_periodogram()
pg
# + [markdown] id="wSzNvRVBfiZm"
# **Box Least Square**
# + colab={"base_uri": "https://localhost:8080/", "height": 643} id="a5J9lWYiYaRE" outputId="388b3b1a-0938-4d21-c80a-16444948fedb"
from astropy.timeseries import BoxLeastSquares
period_grid = np.exp(np.linspace(np.log(1), np.log(15), 50000))
bls = BoxLeastSquares(x, y)
bls_power = bls.power(period_grid, 0.1, oversample=20)
# Save the highest peak as the planet candidate
index = np.argmax(bls_power.power)
bls_period = bls_power.period[index]
bls_t0 = bls_power.transit_time[index]
bls_depth = bls_power.depth[index]
transit_mask = bls.transit_mask(x, bls_period, 0.2, bls_t0)
fig, axes = plt.subplots(2, 1, figsize=(10, 10))
# Plot the periodogram
ax = axes[0]
ax.axvline(np.log10(bls_period), color="C1", lw=5, alpha=0.8)
ax.plot(np.log10(bls_power.period), bls_power.power, "k")
ax.annotate(
"period = {0:.4f} d".format(bls_period),
(0, 1),
xycoords="axes fraction",
xytext=(5, -5),
textcoords="offset points",
va="top",
ha="left",
fontsize=12,
)
ax.set_ylabel("bls power")
ax.set_yticks([])
ax.set_xlim(np.log10(period_grid.min()), np.log10(period_grid.max()))
ax.set_xlabel("log10(period)")
# Plot the folded transit
ax = axes[1]
x_fold = (x - bls_t0 + 0.5 * bls_period) % bls_period - 0.5 * bls_period
m = np.abs(x_fold) < 0.4
ax.plot(x_fold[m], y[m], ".k")
# Overplot the phase binned light curve
bins = np.linspace(-0.41, 0.41, 32)
denom, _ = np.histogram(x_fold, bins)
num, _ = np.histogram(x_fold, bins, weights=y)
denom[num == 0] = 1.0
ax.plot(0.5 * (bins[1:] + bins[:-1]), num / denom, color="C1")
ax.set_xlim(-0.3, 0.3)
ax.set_ylabel("de-trended flux [ppt]")
_ = ax.set_xlabel("time since transit")
# + [markdown] id="lDF_Wg1PfcKn"
# **Lomb-Scargle**
# + colab={"base_uri": "https://localhost:8080/", "height": 352} id="AjKlBzVFYMo8" outputId="8396aaae-1684-4e78-80ba-94ad9ebbdc34"
pg.plot();
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="ZLcYuLU_fsJs" outputId="d77fadb6-e5a1-41cc-e519-72aeaf287137"
pg.plot(scale='log');
| Tugas_Exoplanet_BLS_dan_LombScargle_Adoni.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/As-12/Temple-Image-Classification/blob/master/3_Temple_Classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Ob65Dl6mwENs" colab_type="text"
# # Temple Classification
#
# ### Author
# <NAME>
#
# <EMAIL>
#
# https://thanaphon.dev
#
# ### MIT 3.0 License
#
# Copyright 2020 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# + [markdown] id="MbZYBaOEwKKR" colab_type="text"
# # Introduction
#
# Wat Phra Kaew and Wat Pho are one of the most popular temples in Thailand. Thousands of visitors across the world travel to Thailand every year to appreciate the architecture wonders the temples have to offer.
#
# Unfortunately, many visitors of the temples often get confused about whether their vacation pictures are taken at Wat Phra Kaew or Wat Pho. In order to salvage what's remained of their trip, a machine learning specialist is contacted and tasked with creating a Machine Learning model that can distinguish the photos taken from Wat Phra Kaew and Wat Pho.
#
#
#
# 
#
#
# A sample image of Wat Phra Kaew
#
#
# 
#
# A sample image of Wat Pho
#
# # Objective
#
# The goal is to develop a model that predicts whether a photo is taken at Wat Phra Kaew or Wat Pho - a binary classification problem. There is a possibility of pictures containing neither temples, but an assumption will be made that no such pictures exist in the samples.
#
# # Approach
#
# This machine learning project consists of two primary tasks. Data mining and model development. As this model will not be deployed to production yet, model deployment and operationalization will not be covered.
#
# First, the training dataset will be mined from the internet using Google and Bing image search API. Roughly around 1,000 samples for each class should be sufficient.
#
# Second, because there is an abundant sample of the training dataset, a deep convolutional neural network will be deployed to handle this task. Convolutional Neural Network is generally better suited to handle high dimensionality of image data but it requires specialized hardware such as a Graphical Processing Unit (GPU) to process the information. Without specialized hardware and big training dataset, traditional computer vision approaches such as SIFT feature extraction and bag of SIFTs classification may be more suitable.
#
# Complicated Deep Neural Network usually requires extensive training with millions of training samples. With limited resources and time available, the transfer learning approach will be used in combination with state of the art CNN model, specifically, Google's inception v3 model. The Inception model is user-friendly and very robust against many types of variations such as scale, translation, and rotations. Image preprocessing steps such as color standardization and data augmentation are generally not necessary.
#
# # Training Dataset
#
# The training dataset will be collected by mining images from Google Image Search and Bing Image Search API. See Data Mining notebook for the documentation regarding this process. You can access it [here](https://colab.research.google.com/drive/1YVZQXqzqbGA1kbSkhwtQ9S_vMl7j6qe-?usp=sharing).
#
#
# # Performance Measure
#
# The data provided by the reviewing committee will be used to validate the performance of the model. The model will be optimized and evaluated for precision as there is no repercussion for False Positive and False Negative. Nevertheless, ROC Curve and F1 score will also be calculated for reference purposes.
#
# + [markdown] id="IFVLv7CJ1K5n" colab_type="text"
# # 1. Configurations
#
# ## 1.1 Imports
#
# All the imports go here.
#
# + id="v_bPtODrwJIp" colab_type="code" outputId="c847fd42-b3f0-46cc-a0ff-63716c7e63d2" colab={"base_uri": "https://localhost:8080/", "height": 68}
# set the matplotlib backend so figures can be saved in the background
import os
os.environ['KERAS_BACKEND'] = 'tensorflow'
import matplotlib
import keras
# import the necessary packages
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ModelCheckpoint
from keras.optimizers import Adam
from keras.optimizers import SGD
from sklearn.model_selection import train_test_split
from keras.preprocessing.image import img_to_array
from keras.utils import to_categorical
from imutils import paths
import matplotlib.pyplot as plt
import numpy as np
import argparse
import random
import cv2
import shutil
import fnmatch
import pickle
from keras import Model
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import Flatten
from keras.layers import BatchNormalization
from keras.layers import Activation
from keras.layers import Input
from keras.constraints import max_norm
from keras import regularizers
from keras.layers.advanced_activations import LeakyReLU
import keras.initializers
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
import keras.optimizers
from keras.utils import np_utils
from keras import backend as K
from keras.applications.inception_v3 import InceptionV3
from sklearn.metrics import confusion_matrix
import seaborn as sns
import pandas as pd
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import f1_score, roc_curve, auc
from sklearn.metrics import confusion_matrix, log_loss, precision_recall_fscore_support
from statistics import stdev, mean
# %matplotlib inline
# Ignore warnings for final delivery
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="_mXox0AN5UKu" colab_type="text"
# ## 1.2 Global Settings
#
# These variables will be used across the project.
# + id="kFIGucWDutpR" colab_type="code" colab={}
#Random seeds
SEED = 2082018
np.random.seed(SEED)
#Image Settings
IMAGE_RESIZE = (299,299) # Image size. Process on the by data generator.
IMAGE_INPUT_SIZE = (299,299,3) #Image Input size to the neural network
#Training Settings
BATCH_SIZE = 32
EPOCH = 10
#Directories
# NOTE: The labels are determined by subfolders. PNG or JPEG images only.
TRAIN_DIR = '/content/drive/My Drive/Colab Notebooks/dataset/training/'
VAL_DIR = '/content/drive/My Drive/Colab Notebooks/dataset/validation/'
TEST_DIR = '/content/drive/My Drive/Colab Notebooks/dataset/test/'
#Index of the class label represents numerical representation
CLASS_LABELS = ['0', '1']
CLASS_MEANING = ['Wat Pho', 'Wat Phra Kaew']
NUM_CLASSES = 2
#Checkpoints and save files
#Saving every epochs that improve val accuracy
MODEL_WEIGHT_FILE="temple-classification-inception-v3.hdf5"
# + [markdown] id="-8Bn8bBiAT6H" colab_type="text"
# ## 1.3 Mount Google Drive
# + id="OxWmtX9HATbx" colab_type="code" outputId="f8b4efcf-9090-4887-ba30-6b01be44d4c5" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="9wtOh1EHMTnM" colab_type="text"
# ## 1.4 Execution Environment Information
#
# Obtaining available system memory and graphic card. This can influence training decisions. A low amount of available RAM will require the use of data streaming such as data generators and it will also affect image batch size.
#
# + id="rUFoG17-MSzj" colab_type="code" outputId="a4fe86b8-634f-453e-ec97-6cef86feafd8" colab={"base_uri": "https://localhost:8080/", "height": 34}
from psutil import virtual_memory
ram_gb = virtual_memory().total / 1e9
print("There are {:.1f} GB of RAM available".format(ram_gb))
# + id="wOWg19X3MuCN" colab_type="code" outputId="1da41499-e5b6-45ef-fee1-d78a24921e6b" colab={"base_uri": "https://localhost:8080/", "height": 306}
# !nvidia-smi
# + [markdown] id="Qk54UtMc7gLD" colab_type="text"
# # 2.0 Data Processing
#
#
# ## 2.1 Training / Valiation Split
#
# Because it may not be feasible to load every image into one dataset and split them into several copies for training and testing. This task will copy the image from the original source and split them into an appropriate directory.
# + [markdown] id="SV-MdAYIAnnO" colab_type="text"
# ### 2.1.1 Wat Pho (Label 0)
#
# There are 1000 images of Wat Pho. Split 200 for training validation.
# + id="eKLnEGi-7cBs" colab_type="code" outputId="836031ad-6644-4f7b-f8fa-c89d80e4f088" colab={"base_uri": "https://localhost:8080/", "height": 1000}
run_this = False # Preventing accidental execution
if run_this == True:
split_size = 200
dataset_directory = '/content/drive/My Drive/Colab Notebooks/dataset/bing/wat pho/'
train_directory = '/content/drive/My Drive/Colab Notebooks/dataset/training/0/'
valid_directory = '/content/drive/My Drive/Colab Notebooks/dataset/validation/0/'
# Split Validation Set
count = 0
for file in os.listdir(os.fsencode(dataset_directory)):
filename = os.fsdecode(file)
if count < split_size:
src = os.path.join(dataset_directory, filename)
dst = os.path.join(valid_directory, filename)
print("Copying Validation: {} to {}".format(src, dst))
else:
src = os.path.join(dataset_directory, filename)
dst = os.path.join(train_directory, filename)
print("Copying Training: {} to {}".format(src, dst))
shutil.copyfile(src, dst)
count += 1
# + [markdown] id="myHnp3O6DyA3" colab_type="text"
# ### 2.1.2 Wat Phra Kaew (Label 1)
#
# There are 1000 images of Wat Phra Kaew. Split 200 for training validation.
# + id="ii9hPhI2DyMW" colab_type="code" outputId="e81a7a3f-b8c9-4154-b1fb-25e7ee200145" colab={"base_uri": "https://localhost:8080/", "height": 1000}
run_this = False # Preventing accidental execution
if run_this == True:
split_size = 200
dataset_directory = '/content/drive/My Drive/Colab Notebooks/dataset/bing/wat phra kaew/'
train_directory = '/content/drive/My Drive/Colab Notebooks/dataset/training/1/'
valid_directory = '/content/drive/My Drive/Colab Notebooks/dataset/validation/1/'
# Split Validation Set
count = 0
for file in os.listdir(os.fsencode(dataset_directory)):
filename = os.fsdecode(file)
if count < split_size:
src = os.path.join(dataset_directory, filename)
dst = os.path.join(valid_directory, filename)
print("Copying Validation: {} to {}".format(src, dst))
else:
src = os.path.join(dataset_directory, filename)
dst = os.path.join(train_directory, filename)
print("Copying Training: {} to {}".format(src, dst))
shutil.copyfile(src, dst)
count += 1
# + [markdown] id="gS4v3r2S7nYA" colab_type="text"
# ## 2.2 Data Preprocessing
#
# Minimal data preprocessing is required because Google's Inception v3 model is user-friendly and is generally robust against all kinds of variations.
#
# ### 2.2.1 Resizing
# Resize the image to fit the input layer.
# + id="fPQ40ZLE79qk" colab_type="code" colab={}
def buildImageDataset(path, imageResize=None, shuffle=False, seed=0):
"""
Load dataset into an array. Labels are defined by folder name.
"""
filenames = []
data = []
labels = []
imagePaths = sorted(list(paths.list_images(path)))
if shuffle == True:
random.seed(seed)
random.shuffle(imagePaths)
for imagePath in imagePaths:
image = cv2.imread(imagePath)
if imageResize != None:
image = cv2.resize(image, imageResize)
image = img_to_array(image)
data.append(image)
filenames.append(imagePath)
label = imagePath.split(os.path.sep)[-2]
labels.append(CLASS_LABELS.index(label))
return (np.array(data), np.array(labels), np.array(filenames))
# + id="RBVDxKEyOYZu" colab_type="code" colab={}
X_train, y_train, train_files = buildImageDataset(TRAIN_DIR,imageResize=IMAGE_RESIZE,seed=SEED)
X_valid, y_valid, valid_files = buildImageDataset(VAL_DIR,imageResize=IMAGE_RESIZE,seed=SEED)
# + [markdown] id="4QB2aVZKTPL5" colab_type="text"
# Sanity Check - Review the dimensions of the data
# + id="ZFcb7iLSOmht" colab_type="code" outputId="6c49473b-8f8c-491a-9ba4-5ec972943faf" colab={"base_uri": "https://localhost:8080/", "height": 85}
print ("Shape of the Training Input: {}".format(X_train.shape))
print ("Shape of the Training Target: {}".format(y_train.shape))
print ("Shape of the Validation Input: {}".format(X_valid.shape))
print ("Shape of the Validation Target: {}".format(y_valid.shape))
# + [markdown] id="je76iP8w8wN6" colab_type="text"
# ### 2.2.2 Normalization and Category Encoding
# The Inception v3 Network expects the image pixel data to have a value between 0 and 1. Unfortunately, the typical image pixel is represented as a byte with a value between 0 and 255. Therefore, the image input needs to be normalized by diving it with 255.0.
#
# The classification layer will be 2 class softmax. This requires the target output will be one-hot encoded.
# + id="LBMJUJD0jErF" colab_type="code" colab={}
def preprocess_data(input, output):
return input/255.0 , keras.utils.to_categorical(output)
# + id="hyK9mKzI8wZ3" colab_type="code" colab={}
X_train, y_train = preprocess_data(X_train, y_train)
X_valid, y_valid = preprocess_data(X_valid, y_valid)
# + id="bh5MWhHSnWvt" colab_type="code" outputId="f2fc4789-f7ba-4c4d-cf59-640873fb137d" colab={"base_uri": "https://localhost:8080/", "height": 85}
# Sanity Check
print ("Shape of the Training Input: {}".format(X_train.shape))
print ("Shape of the Training Target: {}".format(y_train.shape))
print ("Shape of the Validation Input: {}".format(X_valid.shape))
print ("Shape of the Validation Target: {}".format(y_valid.shape))
# + [markdown] id="NLQESciHvQPk" colab_type="text"
# # 3. Model Creation
#
# ## 3.1 Inception V3
#
# The Inception v3 network with pre-trained weights from ImageNet is used to initialize the model. The classification layer is replaced by two softmax functions for binary classification.
# + id="Be3DtSetYvCO" colab_type="code" colab={}
# Building Models
def InceptionNet():
#https://keras.io/applications/#inceptionv3
#Use Inception 3 without the last layer.
#Replace last layer with two class softmax for binary classification
sgd = SGD(lr=0.01, momentum=0.9,nesterov=False)
model = keras.applications.inception_v3.InceptionV3(include_top=False,
weights='imagenet', #Use pre-train weight from ImageNet for transfer learning
input_tensor=Input(shape=IMAGE_INPUT_SIZE),
input_shape=None,
pooling='avg',
classes=NUM_CLASSES)
final = Model(input=model.input,output=Dense(NUM_CLASSES, activation='softmax')(model.output))
final.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['accuracy'])
return final
# + id="-WxI528SY4ad" colab_type="code" colab={}
def generateTrainingPlots(history):
plt.style.use('ggplot')
accuracy_plot = plt.figure(figsize=(15,10))
for k in ['val_accuracy', 'accuracy']:
data = np.array(history[k])
plt.plot(data)
plt.title('Model Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch Number')
plt.ylim((0,1.2))
plt.legend(['acc(valid)', 'acc(train)'], loc='upper left')
plt.grid(True)
plt.show()
loss_plot = plt.figure(figsize=(15,10))
for k in ['loss', 'val_loss']:
data = np.array(history[k])
plt.plot(data)
plt.title('Model Loss')
plt.ylabel('Error (Log Loss)')
plt.xlabel('Epoch Number')
plt.grid(True)
plt.legend(['error(train)', 'error(valid)'], loc='upper left')
plt.show()
# + [markdown] id="ETJAdARG09IV" colab_type="text"
# ## 3.1 Overview of the machine learning model
# + id="uXmcxtbpyR_H" colab_type="code" outputId="200090cd-4b92-4275-e28b-2ddeaeab2866" colab={"base_uri": "https://localhost:8080/", "height": 1000}
InceptionNet().summary()
# + [markdown] id="iviUe2V-vYj4" colab_type="text"
# # 4. Training
#
# Due to long training time, only 10 epochs will be evaluated.
# + id="zXAtv5EiZC3d" colab_type="code" outputId="254880c3-9f98-4d82-c47b-3e2c885ab604" colab={"base_uri": "https://localhost:8080/", "height": 1000}
run_this = False #Prevent accidental execution
if run_this:
model = InceptionNet()
# Early stopping by only saving the weight with the best validation accuracy
checkpoint = ModelCheckpoint(MODEL_WEIGHT_FILE, monitor='val_acc', verbose=1, save_best_only=True, mode='max')
callbacks_list = [checkpoint]
#Training
history = model.fit(x=X_train,y=y_train,
validation_data=(X_valid,y_valid),
batch_size=BATCH_SIZE,
epochs = EPOCH,
callbacks = callbacks_list)
#Generate training graphs
generateTrainingPlots(history.history)
# + [markdown] id="a31c-QNWmrls" colab_type="text"
# # 5. Evaluation
#
# The model is evaluated against the test samples. Unfortunately, there are only 8 test samples available. A high variation of scores will be expected.
# + id="zsFqPSOvdVFq" colab_type="code" colab={}
X_test, y_test, _ = buildImageDataset(TEST_DIR, imageResize=IMAGE_RESIZE, seed=SEED)
X_test /= 255.0
# + id="XvU-xX6Am1VW" colab_type="code" outputId="f1a7769c-8574-4892-b403-be5b42db56a4" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Sanity Check
print ("Shape of the Evaluation Input: {}".format(X_test.shape))
print ("Shape of the Evaluation Target: {}".format(y_test.shape))
# + id="czpc4hE8Z_RU" colab_type="code" colab={}
y_pred = model.predict(X_test)
# + id="BaPp2LoadraD" colab_type="code" colab={}
# Translate the sigmoid output into 0s and 1s
predict_result = []
for a,b in y_pred:
if a > b:
predict_result.append(0)
else:
predict_result.append(1)
# + id="G1FzIXuSdsca" colab_type="code" colab={}
def plot_confusion_matrix(cm, classes=None, title='Confusion matrix'):
"""Plots a confusion matrix."""
if classes is not None:
sns.heatmap(cm, xticklabels=classes, yticklabels=classes, vmin=0., vmax=1., annot=True)
else:
sns.heatmap(cm, vmin=0., vmax=1., annot=True)
plt.title(title)
plt.ylabel('True label')
plt.xlabel('Predicted label')
def plot_ROC(y, y_pred):
fpr, tpr, threshold = roc_curve(y, y_pred)
roc_auc = auc(fpr, tpr)
plt.title('Receiver Operating Characteristic')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# + id="3bQ44MJdd4Nj" colab_type="code" outputId="2b6c2078-e75f-4e9a-d2f0-31395a78fa21" colab={"base_uri": "https://localhost:8080/", "height": 598}
# Display CM from one of tests
cm = confusion_matrix(y_test, predict_result)
cm = cm / cm.sum(axis=1)[:, np.newaxis]
plot_confusion_matrix(cm, title="CM for XGBoost (Ground Truth only)")
plt.show()
# Plot ROC
plot_ROC(y_test, predict_result)
# Summarize Scores
scores = precision_recall_fscore_support(y_test, predict_result, average="macro")
print("Precision {} Recall {} F1 {}".format(scores[0], scores[1], scores[2]))
# + [markdown] id="Mdqjoe3zv3Xp" colab_type="text"
# # 5. Analysis
#
# The model performs quite well with 1 missed label on the given 8 images.
# Based on the 99% validation performance over hundreds of images in the validation set, I believe this model is suitable for production deployment.
#
# # 6. Conclusion
#
# In this task, I developed a machine learning model capable of classifying pictures of Wat Pho and Wat Phra Kaew for images in the wild. The training data is mined using Bing and Google Image Search API. Due to limited resources, I leveraged transfer learning using Inception-v3 Convolutional Neural Network initialized with pre-trained weights from ImageNet competition. The model has over 99% accuracy in the validation set, and it can correctly classify 7 out of 8 test images given by the committee.
# + id="-vpS9o1qxlaY" colab_type="code" colab={}
| 3_Temple_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Data Processing: Missing Data and Class Imbalance
#
# _<NAME>, <NAME>_
#
# 12nd February 2022
# +
from copy import deepcopy
from datetime import datetime
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OrdinalEncoder, StandardScaler
# -
# You can download the dataset here: [https://www.kaggle.com/jackdaoud/marketing-data](https://www.kaggle.com/jackdaoud/marketing-data)
# +
data = pd.read_csv('marketing_data.csv')
y = np.array(data['Response']).astype(np.float16)
x = data.drop(['Response'], axis=1)
x = x.drop(['ID'], axis=1)
# -
x.shape
# Let's view the number of missing values.
x.isna().sum()
def remove_value(x):
# Randomly remove values from the data
a, b = x.shape
n_missing = int(a * b * 0.1)
print(f'Removing {n_missing} values randomly.')
for i in range(n_missing):
coor = (np.random.rand(2) * [*x.shape]).astype(np.int16)
coor = np.clip(coor, a_min=0, a_max=max(x.shape) - 1)
x.iloc[coor[0], coor[1]] = np.nan
return x
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.2)
x_train_orig = deepcopy(x_train)
x_test_orig = deepcopy(x_test)
# We remove some of the values randomly.
x_train = remove_value(x_train)
x_test = remove_value(x_test)
# Now we should have more missing values.
x_train.isna().sum()
x_test.isna().sum()
# We summarize the dataset.
pos_res = y_train[y_train == 1].shape[0]
neg_res = y_train[y_train == 0].shape[0]
print(f'Number of positive responses: {pos_res}')
print(f'Number of negative responses: {neg_res}')
print(f'Accuracy if the model always predicts positive: {100 * pos_res/(pos_res+neg_res)}%')
print(f'Accuracy if the model always predicts negative: {100 * neg_res/(pos_res+neg_res)}%')
x_train.isna().sum()
x_train.dtypes
x_train.columns[x_train.dtypes == object]
x_train[x_train.columns[x_train.dtypes == object]].nunique()
# We further structure non-number features.
def ordinal_encode(X, cats):
X = deepcopy(X)
for cat in cats:
X[cat][X[cat].isna()] = 'nan'
enc=OrdinalEncoder()
enc.fit(X[cats])
transformed = enc.transform(X[cats])
for i in range(len(cats)):
nan_pos = np.where(enc.categories_[i] == 'nan')
transformed[[np.where(transformed[:, i] == nan_pos)], i] = np.nan
return transformed, enc
cats = ['Education', 'Marital_Status', 'Country']
a, b = ordinal_encode(x_train, cats)
for i, cat in enumerate(cats):
x_train[cat] = a[:, i]
# +
for cat in cats:
x_test[cat][x_test[cat].isna()] = 'nan'
c = b.transform(x_test[cats])
for i in range(len(cats)):
nan_pos = np.where(b.categories_[i] == 'nan')
c[[np.where(c[:, i] == nan_pos)], i] = np.nan
for i, cat in enumerate(cats):
x_test[cat] = c[:, i]
# +
for cat in cats:
x_train_orig[cat][x_train_orig[cat].isna()] = 'nan'
d = b.transform(x_train_orig[cats])
for i in range(len(cats)):
nan_pos = np.where(b.categories_[i] == 'nan')
d[[np.where(d[:, i] == nan_pos)], i] = np.nan
for i, cat in enumerate(cats):
x_train_orig[cat] = d[:, i]
# -
x_train
x_test
# +
# x_train = pd.get_dummies(x_train, columns=['Education', 'Marital_Status', 'Country'], dummy_na=True)
# x_test = pd.get_dummies(x_test, columns=['Education', 'Marital_Status', 'Country'], dummy_na=True)
# +
# x_train_orig = pd.get_dummies(x_train_orig, columns=['Education', 'Marital_Status', 'Country'], dummy_na=True)
# -
incomes = [float(inc.lstrip('$').replace(',', '')) for inc in x_train[' Income '] if isinstance(inc, str)]
x_train[' Income '][x_train[' Income '].apply(lambda x: isinstance(x, str))] = incomes
x_train[' Income '] = x_train[' Income '].astype(np.float64)
incomes = [float(inc.lstrip('$').replace(',', '')) for inc in x_train_orig[' Income '] if isinstance(inc, str)]
x_train_orig[' Income '][x_train_orig[' Income '].apply(lambda x: isinstance(x, str))] = incomes
x_train_orig[' Income '] = x_train_orig[' Income '].astype(np.float64)
x_train
incomes = [float(inc.lstrip('$').replace(',', '')) for inc in x_test[' Income '] if isinstance(inc, str)]
x_test[' Income '][x_test[' Income '].apply(lambda x: isinstance(x, str))] = incomes
x_test[' Income '] = x_test[' Income '].astype(np.float64)
x_train['Dt_Customer'] = (datetime.today() - pd.to_datetime(x_train['Dt_Customer'])).dt.days
x_test['Dt_Customer'] = (datetime.today() - pd.to_datetime(x_test['Dt_Customer'])).dt.days
x_train_orig['Dt_Customer'] = (datetime.today() - pd.to_datetime(x_train_orig['Dt_Customer'])).dt.days
x_train.insert(2, 'Age', [2021 - birth for birth in x_train['Year_Birth']])
x_test.insert(2, 'Age', [2021 - birth for birth in x_test['Year_Birth']])
x_train_orig.insert(2, 'Age', [2021 - birth for birth in x_train_orig['Year_Birth']])
x_train.dtypes
# + [markdown] tags=[]
# ## Correlation
# + tags=[]
corr_mat = x_train.corr()
# + tags=[]
x_train.corrwith(pd.DataFrame(y_train)[0])
# + tags=[]
corr_mat.to_csv('corr_mat.csv')
# + tags=[]
corr_mat
# + tags=[]
corr_mat['Age'][np.abs(corr_mat['Age']) > 0.5]
# + [markdown] tags=[]
# ## Scaling
# -
# In general, models behave heavily biased on unscaled datasets. Therefore, it is better to scale the data.
sc = StandardScaler()
x_train_orig = pd.DataFrame(sc.fit_transform(x_train_orig), columns=x_train.columns)
x_train = pd.DataFrame(sc.fit_transform(x_train), columns=x_train.columns)
x_test = pd.DataFrame(sc.transform(x_test), columns=x_test.columns)
# + [markdown] tags=[]
# ## Filling Missing Data
#
# Mainly, there are two types of techniques:
#
# - Univariate methods: By considering only that specific feature.
# - Mean
# - Median
# - Mod
# - Fix value
# - Multivariate methods: By considering not only one feature but also a mix of other features
# - Closest Neighbor
# - Train a model for the missing data
#
# We will first demonstrate univariate methods.
# -
# ### Univariate Methods: Mean, Median, Mod, Fixed Value
from sklearn.impute import SimpleImputer
# +
imputers = []
imp_mean = SimpleImputer(missing_values=np.nan, strategy='mean')
imputers.append(imp_mean)
imp_median = SimpleImputer(missing_values=np.nan, strategy='median')
imputers.append(imp_median)
imp_mode = SimpleImputer(missing_values=np.nan, strategy='most_frequent')
imputers.append(imp_mode)
imp_constant = SimpleImputer(missing_values=np.nan, strategy='constant', fill_value=0)
imputers.append(imp_constant)
# -
for imp in imputers:
imp.fit(x_train)
x_train.mode().iloc[0]
print(x_train.mean())
print(x_train.median())
x_mean_train = pd.DataFrame(imp_mean.transform(x_train), columns=x_train.columns)
x_median_train = pd.DataFrame(imp_median.transform(x_train), columns=x_train.columns)
x_mode_train = pd.DataFrame(imp_mode.transform(x_train), columns=x_train.columns)
x_const_train = pd.DataFrame(imp_mode.transform(x_train), columns=x_train.columns)
# + [markdown] tags=[]
# ### Multivariate Methods
#
# We will demonstrate K-Nearest Neighbor.
# + [markdown] tags=[]
# #### K-Nearest Neighbor
# -
from sklearn.impute import KNNImputer
imp_knn = KNNImputer(missing_values=np.nan, n_neighbors=5, weights='distance')
imp_knn.fit(x_train)
x_knn_train = pd.DataFrame(imp_knn.transform(x_train), columns=x_train.columns)
x_knn_train
# #### Iterative Methods
#
# We can generate missing data using another classifier.
from sklearn.ensemble import RandomForestRegressor
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
from sklearn.linear_model import BayesianRidge
from sklearn.tree import DecisionTreeRegressor
imp_linear = IterativeImputer(max_iter=100, random_state=0, estimator=BayesianRidge())
imp_tree = IterativeImputer(max_iter=100, random_state=0, estimator=DecisionTreeRegressor(max_features='sqrt', random_state=0))
# imp_forest = IterativeImputer(max_iter = 25, random_state=0, estimator=RandomForestRegressor(n_estimators=20, random_state=0))
imp_linear = IterativeImputer(max_iter=100, random_state=0, estimator=BayesianRidge())
imp_linear.fit(x_train)
imp_tree.fit(x_train)
# +
# imp_forest.fit(x_train)
# -
x_linear_train = pd.DataFrame(imp_linear.transform(x_train), columns=x_train.columns)
x_tree_train = pd.DataFrame(imp_tree.transform(x_train), columns=x_train.columns)
# x_forest_train = pd.DataFrame(imp_forest.transform(x_train), columns=x_train.columns)
# ### Comparison
print('Metric: Mean Absolute Error (MAE)\n')
print(f'Imputing by Mean: {((np.abs(x_train_orig - x_mean_train))).mean().mean()}')
print(f'Imputing by Median: {((np.abs(x_train_orig - x_median_train))).mean().mean()}')
print(f'Imputing by Mode: {((np.abs(x_train_orig - x_mode_train))).mean().mean()}')
print(f'Imputing by Fixed Value: {((np.abs(x_train_orig - x_const_train))).mean().mean()}')
print(f'Imputing by K-Nearest Neighbour: {((np.abs(x_train_orig - x_knn_train))).mean().mean()}')
print(f'Imputing by Linear Reggression Regresyona Göre: {((np.abs(x_train_orig - x_linear_train))).mean().mean()}')
print(f'Imputing by Decision TreeKarar Ağacına Göre: {((np.abs(x_train_orig - x_tree_train))).mean().mean()}')
# ### Transforming Test Set
x_test = pd.DataFrame(imp_knn.transform(x_test), columns=x_test.columns)
x_test
# Inverse transform to get the originals
pd.DataFrame(sc.inverse_transform(x_test), columns=x_test.columns)
# + [markdown] tags=[]
# ## Handling Class Imbalance
#
# Class imbalance is having unequal number of samples in each class, especially when most of the data belong to only one class. This can be problematic since our model usually learns to choose the most popular class, since it maximizes accuracy.
#
# Here are some methods to mitigate class imbalance problem:
#
# - Shifting decision boundary: For example, we can shift decision boundary of 0.5 of logistic regression and set it to a higher of lower value according to our distribution.
# - Oversampling: Sampling new data from the given dataset. There are a variety of methods for this having their advantages and disadvantages.
# - Undersampling: Why not just throw some of the data?
# - Generating new data: These are more complex.
# - Generative Adversarial Networks (GANs)
# - Normalizing Flows
# - Variational Autoencoders
#
# ### SMOTE
#
# We add a new data point between two data points of the least represented class.
#
# <figure>
# <img src="img/smote.png">
# <figcaption>Visualisation of SMOTE algorithm. <br> (Figure from: <i>https://www.kaggle.com/rafjaa/resampling-strategies-for-imbalanced-datasets</i>)</figcaption>
# </figure>
#
# ### Tomek
#
# We remove close and different data point pairs in order to make decision boundary clearer.
#
# <figure>
# <img src="img/tomek.png">
# <figcaption>Visualisation of Tomek algorithm. <br> (Figure from: <i>https://www.kaggle.com/rafjaa/resampling-strategies-for-imbalanced-datasets</i>)</figcaption>
# </figure>
# -
from imblearn.combine import SMOTETomek, SMOTEENN
from imblearn.over_sampling import RandomOverSampler
from lightgbm import LGBMClassifier
from sklearn.metrics import classification_report
from sklearn.linear_model import LogisticRegression
smote_tomek = SMOTETomek(random_state=0)
smote_enn = SMOTEENN(random_state=0)
ros = RandomOverSampler(random_state=0)
x_resampled_tomek, y_resampled_tomek = smote_tomek.fit_resample(x_linear_train, y_train)
x_resampled_enn, y_resampled_enn = smote_enn.fit_resample(x_linear_train, y_train)
x_oversampled, y_oversampled = ros.fit_resample(x_linear_train, y_train)
# +
clf = [LGBMClassifier(), LGBMClassifier(), LGBMClassifier(), LGBMClassifier()]
clf[0].fit(x_knn_train, y_train)
clf[1].fit(x_oversampled, y_oversampled)
clf[2].fit(x_resampled_tomek, y_resampled_tomek)
clf[3].fit(x_resampled_enn, y_resampled_enn)
print('Base')
print(classification_report(y_test, clf[0].predict(x_test)))
print('Random Oversampling')
print(classification_report(y_test, clf[1].predict(x_test)))
print('SMOTE Tomek')
print(classification_report(y_test, clf[2].predict(x_test)))
print('SMOTE ENN')
print(classification_report(y_test, clf[0].predict(x_test)))
# +
clf = [LogisticRegression(), LogisticRegression(), LogisticRegression(), LogisticRegression()]
clf[0].fit(x_knn_train, y_train)
clf[1].fit(x_oversampled, y_oversampled)
clf[2].fit(x_resampled_tomek, y_resampled_tomek)
clf[3].fit(x_resampled_enn, y_resampled_enn)
print('Base')
print(classification_report(y_test, clf[0].predict(x_test)))
print('Random Oversampling')
print(classification_report(y_test, clf[1].predict(x_test)))
print('SMOTE Tomek')
print(classification_report(y_test, clf[2].predict(x_test)))
print('SMOTE ENN')
print(classification_report(y_test, clf[0].predict(x_test)))
# + tags=[]
print('Shifting Decision Boundary')
print(classification_report(y_test,
np.array(
pd.DataFrame(clf[0].predict_proba(x_test)).applymap(
lambda x: 1 if x>0.4 else 0)[1]
)))
| Batch_6_January2022/Week5/notebooks/2. Data Processing, Missing Data and Class Imbalance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from __future__ import print_function
import sklearn
from sklearn.ensemble import RandomForestClassifier
from sklearn import preprocessing
from datetime import datetime
import os
# %matplotlib inline
# %config InlineBackend.figure_format = 'png'
pd.set_option("max_columns",50)
# -
# %%time
train = pd.read_csv("../data/train.csv", index_col=0)
train = train.reset_index(drop=True)
train = train[train["is_booking"] == 1]
np.random.seed(402)
train = train.ix[np.random.choice(train.index, 100000, replace=False)]
train = train.reset_index(drop=True)
# +
# dtype_set = {'site_name':np.int32, 'posa_continent':np.int32, 'user_location_country':np.int32, 'orig_destination_distance':np.int32, \
# 'user_id':np.int32, 'is_mobile':bool, 'is_package':bool, 'channel':np.int32, 'srch_adults_cnt':np.int32, 'srch_children_cnt':np.int32, \
# 'srch_rm_cnt':np.int32, 'srch_destination_id':np.int32, 'srch_destination_type_id':np.int32, 'is_booking':bool, \
# 'cnt':np.int32, 'hotel_country':np.int32, 'hotel_cluster':np.int32}
# usecols_set = ['user_id', 'site_name', 'posa_continent', 'user_location_country', 'orig_destination_distance', 'is_mobile', 'is_package', 'channel', \
# 'srch_adults_cnt', 'srch_children_cnt', 'srch_rm_cnt', 'srch_destination_id', 'srch_destination_type_id', 'is_booking', 'cnt', 'hotel_country', 'hotel_cluster']
# # %%time
# train = pd.read_csv('../data/train.csv',
# dtype=dtype_set,
# usecols=usecols_set,
# chunksize=1000000)
# -
train["num"] = 1
train.head()
table = pd.pivot_table(train, values='num', index=['hotel_cluster', 'user_id'], columns=['is_booking'], aggfunc=np.sum)
train[train["user_id"] == 12053]
train = train.drop_duplicates()
train.groupby(['srch_destination_id',
'hotel_cluster'])['is_booking'].agg(['sum','count']).head()
train[train.srch_destination_id == 4][train.hotel_cluster == 78]
# +
train[["is_booking","cnt"]].head()
train["ratio"] = train["is_booking"]/train["cnt"]
# -
train["cnt"].value_counts() # 대부분 한번에 보고 결제해버림..! => 다른 대체물을 많이 보는편이 아님.
train["ratio"].value_counts()
train[train.cnt == 8]
train[train.user_id == 1195399]
train.groupby("hotel_cluster").agg(sum)
# +
train1 = pd.read_csv('../data/train_2013.csv',
dtype={'is_booking':bool,'srch_destination_id':np.int32, 'hotel_cluster':np.int32},
usecols=['srch_destination_id','is_booking','hotel_cluster'],
chunksize=1000000)
# dtype을 설정하고(bool, np.int32) chunksize로 끊어서 하면 더 빠르게 데이터를 처리할 수 있음!!
aggs = []
print('-'*38)
for chunk in train1:
agg = chunk.groupby(['srch_destination_id',
'hotel_cluster'])['is_booking'].agg(['sum','count'])
agg.reset_index(inplace=True)
aggs.append(agg)
print('.',end='')
print('')
aggs = pd.concat(aggs, axis=0)
# -
CLICK_WEIGHT = 0.05
agg = aggs.groupby(['srch_destination_id','hotel_cluster']).sum().reset_index()
agg.head()
agg['count'] -= agg['sum']
# sum은 실제 booking과 count를 합친 것..!
agg = agg.rename(columns={'sum':'bookings','count':'clicks'})
agg['relevance'] = agg['bookings'] + CLICK_WEIGHT * agg['clicks']
agg.head()
# +
def most_popular(group, n_max=5):
relevance = group['relevance'].values
hotel_cluster = group['hotel_cluster'].values
most_popular = hotel_cluster[np.argsort(relevance)[::-1]][:n_max]
return np.array_str(most_popular)[1:-1] # remove square brackets
most_pop = agg.groupby(['srch_destination_id']).apply(most_popular)
most_pop = pd.DataFrame(most_pop).rename(columns={0:'hotel_cluster'})
most_pop.head()
# -
most_pop.tail()
train
train["hotel_cluster"]
delete_list = ["posa_continent", "user_location_country", "user_location_region", "user_location_city", "is_mobile", "channel"]
train = train.drop(delete_list, axis=1)
train.sort_values("hotel_cluster")
train.sort_values("user_id")
train.groupby(['srch_destination_id',
'hotel_cluster'])['is_booking'].agg(['sum','count']).sort_values("count", ascending=False)
train[train["hotel_cluster"] == 3].sort_values("is_booking")
train[train["hotel_cluster"] == 44]
train[train["srch_destination_id"] == 8250].sort_values("is_booking", ascending=False)
agg.head()
| notebook/09. Find unique People.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# # image and station data cleaning
stations_by_name=pd.read_json("raw_data/stations_by_name_with_alias.json",orient='index')
stations_by_name["img"]=stations_by_name.ST_NAME_EN+".png"
stations_by_name=stations_by_name[stations_by_name.index!="严御路"]
stations_by_name=stations_by_name[stations_by_name.index!="外高桥保税区北"]
stations_by_name=stations_by_name[stations_by_name.index!="李子园路"]
stations_by_name=stations_by_name[stations_by_name.index!="淞浜路"]
stations_by_name=stations_by_name[stations_by_name.index!="外高桥保税区南"]
stations_by_name=stations_by_name[stations_by_name.index!="上海大学站"]
stations_by_name=stations_by_name[stations_by_name.index!="上海野生动物园"]
stations_by_name=stations_by_name.drop_duplicates(subset=None, keep='first', inplace=False)
stations_by_name
stations_by_name.to_json(r"/Users/HYT/Desktop/CS573 Data Visualization/git/gradfinal/data/stations_by_name.json",orient='index',force_ascii=False)
# # exit and entry data cleaning
stations_by_name[stations_by_name.duplicated(keep=False)]
stations_by_name[stations_by_name.ST_NAME=="淞浜路"]
stations_by_name[stations_by_name.ST_NAME=="淞浜路"]
stations_by_name[stations_by_name.index=="李子园"]
| data/SHMTR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Two Rectangles // Intersection Area
#
# Given two rectangles on a 2D graph, return the area of their intersection. If the rectangles don't intersect, return 0.
#
# For example, given the following rectangles:
#
# {
# "top_left": (1, 4),
# "dimensions": (3, 3) # width, height
# }
# and
#
# {
# "top_left": (0, 5),
# "dimensions": (4, 3) # width, height
# }
# return 6.
#
#
# ### Answer
# The trick here is that you need to find a few points without knowing how they're placed. The right-most most left border, the bottom-most top border, the left-most right border, and the top-most bottom border. If these pairs don't cross, then you know they don't intersect.
def rectangles(rec1, rec2):
left_x = max(rec1["top_left"][0], rec2["top_left"][0])
right_x = min(rec1["top_left"][0] + rec1["dimensions"][0],
rec2["top_left"][0] + rec2["dimensions"][0])
top_y = min(rec1["top_left"][1], rec2["top_left"][1])
bottom_y = max(rec1["top_left"][1] - rec1["dimensions"][1],
rec2["top_left"][1] - rec2["dimensions"][1])
if left_x > right_x or bottom_y > top_y:
return 0
return (right_x - left_x) * (top_y - bottom_y)
| Daily/Overlapping Rectangles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:qiskit]
# language: python
# name: conda-env-qiskit-py
# ---
# +
# default_exp overlap.swap
# +
#export
import numpy as np
from fisherman.utils import sym_from_triu
from qiskit import BasicAer
from qiskit.utils import QuantumInstance
from qiskit.circuit import ParameterExpression
from qiskit.providers import Backend, BaseBackend
from qiskit import QuantumCircuit, ClassicalRegister
from qiskit.opflow import CircuitStateFn, CircuitOp
from collections.abc import Iterable
from typing import Optional, Union, Dict, List, Iterable
# -
# # Swap test
# > Functionalities to compute the overlap between states through a swap test.
# +
#export
def swap_test_overlap(
state0: Union[QuantumCircuit, CircuitStateFn],
state1: Optional[Union[QuantumCircuit, CircuitStateFn, Iterable[Union[QuantumCircuit, CircuitStateFn]]]] = None,
param_dict: Optional[Dict[ParameterExpression, List[float]]] = None,
backend: Optional[Union[Backend, QuantumInstance]] = None
) -> np.ndarray:
"""Returns overlap between states using swap test."""
if state1 is not None and param_dict is not None:
raise ValueError(
"swap_test_overlap only accepts one optional input "
"either `state1` or `param_dict`."
)
backend = BasicAer.get_backend('qasm_simulator') if backend is None else backend
qi = QuantumInstance(backend) if isinstance(backend, Backend) else backend
if qi.is_statevector:
raise ValueError("swap_test_overlap does not suport statevector simulator yet.")
if isinstance(state0, QuantumCircuit):
state0 = CircuitStateFn(state0)
if state1 is not None:
if not isinstance(state1, Iterable):
state1 = [state1]
state1 = [CircuitStateFn(s) if isinstance(s, QuantumCircuit) else s for s in state1]
n_qubits = state0.num_qubits
swap_op = _swap_test_operator(n_qubits)
ancilla = CircuitStateFn(QuantumCircuit(1))
def _swap_circuit(s0, s1):
circuit = (swap_op @ (s1 ^ s0 ^ ancilla)).primitive
circuit.add_register(ClassicalRegister(1))
circuit.measure(0, 0)
return circuit
if state1 is not None:
circuits = [_swap_circuit(state0, s) for s in state1]
elif param_dict is not None:
states = state0.bind_parameters(param_dict)
circuits = [_swap_circuit(s_i, s_j)
for i, s_i in enumerate(states) for s_j in states[i:]]
else:
circuits = [_swap_circuit(state0, state0)]
counts = qi.execute(circuits).get_counts()
if not isinstance(counts, list):
counts = [counts]
p0 = [c.get('0', 0)/sum(c.values()) for c in counts]
overlaps = 2*np.array(p0) - 1
return overlaps.squeeze() if param_dict is None else sym_from_triu(overlaps, len(states))
def _swap_test_operator(n_qubits):
swap_qc = QuantumCircuit(2*n_qubits+1)
swap_qc.h(0)
for q in range(n_qubits):
swap_qc.cswap(0, q + 1, q + 1 + n_qubits)
swap_qc.h(0)
return CircuitOp(swap_qc)
# -
# The swap test is one of the most common ways of computing the overlap between quantum states. It relies on the construction of a circuit of $2n+1$ qubits, where $n$ is the number of qubits of the states that are to be compared. The additional qubit is used as control qubit for a set of control-swap gates performed between the qubits of the states.
#
# We have taken the convention of having the additional qubit as the first one. This way, the operator applied to the tensored states looks as follows for two 3-qubit states.
_swap_test_operator(3).primitive.draw('mpl')
# The first state $|\psi\rangle$ would, for instance, take qubits $q_1, q_2, q_3$ and the second state $|\phi\rangle$ would take $q_4, q_5, q_6$. The ancilla qubit, $q0$ is then measured and the probability of measuring zero $P_0$ is proportional to the overlap between the states, such that $$\text{Tr}\left[\rho_\psi\rho_\phi\right] = 2P_0 - 1.$$
#
# The behaviour is analogous to the other overlap computation functions with the exception that `swap_test_overlap` does accept `QuantumCircuit` inputs, provided that it heavily relies on quantum circuit construction.
#
# See some function call examples below and refer to [the basic usage](https://borjarequena.github.io/Quantum-Fisherman/#Basic-usage) for a joint explanation of the overlap computation functions.
#hide
from qiskit.circuit import Parameter
qc0 = QuantumCircuit(2)
qc0.x(0)
qc0.draw('mpl')
theta0 = Parameter('θ')
qc1 = QuantumCircuit(2)
qc1.x(0)
qc1.rx(theta0, 1)
qc1.draw('mpl')
state0 = CircuitStateFn(qc0)
purity = swap_test_overlap(state0)
purity
theta_values = [0, np.pi/2, np.pi, 3*np.pi/2, 2*np.pi]
param_dict = {theta0: theta_values}
state0 = CircuitStateFn(qc0)
state1 = CircuitStateFn(qc1).bind_parameters(param_dict)
overlaps = swap_test_overlap(state0, state1)
overlaps
param_state = CircuitStateFn(qc1)
overlaps = swap_test_overlap(param_state, param_dict=param_dict)
overlaps
overlaps.diagonal()
| nbs/01_overlap.swap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pydeck Earth Engine Introduction
#
# This is an introduction to using [Pydeck](https://pydeck.gl) and [Deck.gl](https://deck.gl) with [Google Earth Engine](https://earthengine.google.com/) in Jupyter Notebooks.
# If you wish to run this locally, you'll need to install some dependencies. Installing into a new Conda environment is recommended. To create and enter the environment, run:
# ```
# conda create -n pydeck-ee -c conda-forge python jupyter notebook pydeck earthengine-api requests -y
# source activate pydeck-ee
# jupyter nbextension install --sys-prefix --symlink --overwrite --py pydeck
# jupyter nbextension enable --sys-prefix --py pydeck
# ```
# then open Jupyter Notebook with `jupyter notebook`.
# Now in a Python Jupyter Notebook, let's first import required packages:
from pydeck_earthengine_layers import EarthEngineLayer
import pydeck as pdk
import requests
import ee
# ## Authentication
#
# Using Earth Engine requires authentication. If you don't have a Google account approved for use with Earth Engine, you'll need to request access. For more information and to sign up, go to https://signup.earthengine.google.com/.
# If you haven't used Earth Engine in Python before, you'll need to run the following authentication command. If you've previously authenticated in Python or the command line, you can skip the next line.
#
# Note that this creates a prompt which waits for user input. If you don't see a prompt, you may need to authenticate on the command line with `earthengine authenticate` and then return here, skipping the Python authentication.
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
# ## Create Map
#
# Next it's time to create a map. Here we create an `ee.Image` object
# Initialize objects
ee_layers = []
view_state = pdk.ViewState(latitude=37.7749295, longitude=-122.4194155, zoom=10, bearing=0, pitch=45)
# +
# %%
# Add Earth Engine dataset
# Load SRTM Digital Elevation Model data.
image = ee.Image('CGIAR/SRTM90_V4');
# Define an SLD style of discrete intervals to apply to the image.
sld_intervals = \
'<RasterSymbolizer>' + \
'<ColorMap type="intervals" extended="false" >' + \
'<ColorMapEntry color="#0000ff" quantity="0" label="0"/>' + \
'<ColorMapEntry color="#00ff00" quantity="100" label="1-100" />' + \
'<ColorMapEntry color="#007f30" quantity="200" label="110-200" />' + \
'<ColorMapEntry color="#30b855" quantity="300" label="210-300" />' + \
'<ColorMapEntry color="#ff0000" quantity="400" label="310-400" />' + \
'<ColorMapEntry color="#ffff00" quantity="1000" label="410-1000" />' + \
'</ColorMap>' + \
'</RasterSymbolizer>';
# Define an sld style color ramp to apply to the image.
sld_ramp = \
'<RasterSymbolizer>' + \
'<ColorMap type="ramp" extended="false" >' + \
'<ColorMapEntry color="#0000ff" quantity="0" label="0"/>' + \
'<ColorMapEntry color="#00ff00" quantity="100" label="100" />' + \
'<ColorMapEntry color="#007f30" quantity="200" label="200" />' + \
'<ColorMapEntry color="#30b855" quantity="300" label="300" />' + \
'<ColorMapEntry color="#ff0000" quantity="400" label="400" />' + \
'<ColorMapEntry color="#ffff00" quantity="500" label="500" />' + \
'</ColorMap>' + \
'</RasterSymbolizer>';
# Add the image to the map using both the color ramp and interval schemes.
view_state = pdk.ViewState(longitude=-76.8054, latitude=42.0289, zoom=8)
ee_layers.append(EarthEngineLayer(ee_object=image.sldStyle(sld_intervals), vis_params={}))
ee_layers.append(EarthEngineLayer(ee_object=image.sldStyle(sld_ramp), vis_params={}))
# -
# Then just pass these layers to a `pydeck.Deck` instance, and call `.show()` to create a map:
r = pdk.Deck(layers=ee_layers, initial_view_state=view_state)
r.show()
| Visualization/image_color_ramp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with libraries
# ##3 Most of the power of a programming language is in its libraries.
#
# * A *library* is a collection of files (called *modules*) that contains
# functions for use by other programs.
# * May also contain data values (e.g., numerical constants) and other things.
# * Library's contents are supposed to be related, but there's no way to enforce that.
# * The Python [standard library][stdlib] is an extensive suite of modules that comes
# with Python itself.
# * Many additional libraries are available from [PyPI][pypi] (the Python Package Index).
# * We will see later how to write new libraries.
#
# ```{Note}
# ## Most of the power of a programming language is in its libraries.
#
# * A *library* is a collection of files (called *modules*) that contains
# functions for use by other programs.
# * May also contain data values (e.g., numerical constants) and other things.
# * Library's contents are supposed to be related, but there's no way to enforce that.
# * The Python [standard library][stdlib] is an extensive suite of modules that comes
# with Python itself.
# * Many additional libraries are available from [PyPI][pypi] (the Python Package Index).
# * We will see later how to write new libraries.
# ```
# +
import math
print('pi is', math.pi)
print('cos(pi) is', math.cos(math.pi))
# -
# * Have to refer to each item with the module's name.
# * `math.cos(pi)` won't work: the reference to `pi`
# doesn't somehow "inherit" the function's reference to `math`.
#
# ## Use `help` to learn about the contents of a library module.
#
# * Works just like help for a function.
help(math)
# ## Import specific items from a library module to shorten programs.
#
# * Use `from ... import ...` to load only specific items from a library module.
# * Then refer to them directly without library name as prefix.
# +
from math import cos, pi
print('cos(pi) is', cos(pi))
# -
# ## Create an alias for a library module when importing it to shorten programs.
#
# * Use `import ... as ...` to give a library a short *alias* while importing it.
# * Then refer to items in the library using that shortened name.
# +
import math as m
print('cos(pi) is', m.cos(m.pi))
# -
# * Commonly used for libraries that are frequently used or have long names.
# * E.g., the `matplotlib` plotting library is often aliased as `mpl`.
# * But can make programs harder to understand,
# since readers must learn your program's aliases.
# ```{admonition} Exercise: Exploring the Math Module
# 1. What function from the `math` module can you use to calculate a square root
# *without* using `sqrt`?
# 2. Since the library contains this function, why does `sqrt` exist?
#
# ```
#
# :::{admonition} See Solution
# :class: tip, dropdown
# 1. Using `help(math)` we see that we've got `pow(x,y)` in addition to `sqrt(x)`, so we could use `pow(x, 0.5)` to find a square root.
# 2. The `sqrt(x)` function is arguably more readable than `pow(x, 0.5)` when implementing equations. Readability is a cornerstone of good programming, so it makes sense to provide a special function for this specific common case.
# Also, the design of Python's `math` library has its origin in the C standard, which includes both `sqrt(x)` and `pow(x,y)`, so a little bit of the history of programming is showing in Python's function names.
# :::
#
# ```{admonition} Exercise: Locating the right Module
# You want to select a random character from a string:
#
# ~~~
# bases = 'ACTTGCTTGAC'
# ~~~~
#
# 1. Which [standard library][stdlib] module could help you?
# 2. Which function would you select from that module? Are there alternatives?
# 3. Try to write a program that uses the function.
#
# ```
# :::{admonition} See Solution
# :class: tip, dropdown
# The [random module][randommod] seems like it could help you.
#
# The string has 11 characters, each having a positional index from 0 to 10.
# You could use `random.randrange` function (or the alias `random.randint` if you find that easier to remember) to get a random integer between 0 and 10, and then pick out the character at that position:
#
# ~~~
# from random import randrange
#
# random_index = randrange(len(bases))
# print(bases[random_index])
# ~~~
#
# ~~~
# from random import randrange
#
# print(bases[randrange(len(bases))])
# ~~~
# Perhaps you found the `random.sample` function? It allows for slightly less typing:
# ~~~
# from random import sample
# print(sample(bases, 1)[0])
# ~~~
# Note that this function returns a list of values. There's also other functions you could use, but with more convoluted
# code as a result.
# :::
#
| 02_Day_2/exercise/B0_Exercise_libs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (tensorflow)
# language: python
# name: tensorflow
# ---
# <a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_08_2_keras_ensembles.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # T81-558: Applications of Deep Neural Networks
# **Module 8: Kaggle Data Sets**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Module 8 Material
#
# * Part 8.1: Introduction to Kaggle [[Video]](https://www.youtube.com/watch?v=v4lJBhdCuCU&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_1_kaggle_intro.ipynb)
# * **Part 8.2: Building Ensembles with Scikit-Learn and Keras** [[Video]](https://www.youtube.com/watch?v=LQ-9ZRBLasw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_2_keras_ensembles.ipynb)
# * Part 8.3: How Should you Architect Your Keras Neural Network: Hyperparameters [[Video]](https://www.youtube.com/watch?v=1q9klwSoUQw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_3_keras_hyperparameters.ipynb)
# * Part 8.4: Bayesian Hyperparameter Optimization for Keras [[Video]](https://www.youtube.com/watch?v=sXdxyUCCm8s&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_4_bayesian_hyperparameter_opt.ipynb)
# * Part 8.5: Current Semester's Kaggle [[Video]](https://www.youtube.com/watch?v=PHQt0aUasRg&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_5_kaggle_project.ipynb)
#
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow.
# Running the following code will map your GDrive to ```/content/drive```.
# +
try:
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
COLAB = True
print("Note: using Google CoLab")
# %tensorflow_version 2.x
except:
print("Note: not using Google CoLab")
COLAB = False
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
# -
# # Part 8.2: Building Ensembles with Scikit-Learn and Keras
#
# ### Evaluating Feature Importance
#
# Feature importance tells us how important each of the features (from the feature/import vector is to the prediction of a neural network or another model. There are many different ways to evaluate the feature importance for neural networks. The following paper presents an excellent (and readable) overview of the various means of assessing the significance of neural network inputs/features.
#
# * An accurate comparison of methods for quantifying variable importance in artificial neural networks using simulated data [[Cite:olden2004accurate]](http://depts.washington.edu/oldenlab/wordpress/wp-content/uploads/2013/03/EcologicalModelling_2004.pdf). *Ecological Modelling*, 178(3), 389-397.
#
# In summary, the following methods are available to neural networks:
#
# * Connection Weights Algorithm
# * Partial Derivatives
# * Input Perturbation
# * Sensitivity Analysis
# * Forward Stepwise Addition
# * Improved Stepwise Selection 1
# * Backward Stepwise Elimination
# * Improved Stepwise Selection
#
# For this class, we will use the **Input Perturbation** feature ranking algorithm. This algorithm will work with any regression or classification network. I provide an implementation of the input perturbation algorithm for scikit-learn in the next section. This code implements a function below that will work with any scikit-learn model.
#
# [<NAME>](https://en.wikipedia.org/wiki/Leo_Breiman) provided this algorithm in his seminal paper on random forests. [[Citebreiman2001random:]](https://www.stat.berkeley.edu/~breiman/randomforest2001.pdf) Although he presented this algorithm in conjunction with random forests, it is model-independent and appropriate for any supervised learning model. This algorithm, known as the input perturbation algorithm, works by evaluating a trained model’s accuracy with each of the inputs individually shuffled from a data set. Shuffling an input causes it to become useless—effectively removing it from the model. More important inputs will produce a less accurate score when they are removed by shuffling them. This process makes sense because important features will contribute to the accuracy of the model. I first presented the TensorFlow implementation of this algorithm in the following paper.
#
# * Early stabilizing feature importance for TensorFlow deep neural networks[[Cite:heaton2017early]](https://www.heatonresearch.com/dload/phd/IJCNN%202017-v2-final.pdf)
#
# This algorithm will use log loss to evaluate a classification problem and RMSE for regression.
# +
from sklearn import metrics
import scipy as sp
import numpy as np
import math
from sklearn import metrics
def perturbation_rank(model, x, y, names, regression):
errors = []
for i in range(x.shape[1]):
hold = np.array(x[:, i])
np.random.shuffle(x[:, i])
if regression:
pred = model.predict(x)
error = metrics.mean_squared_error(y, pred)
else:
pred = model.predict_proba(x)
error = metrics.log_loss(y, pred)
errors.append(error)
x[:, i] = hold
max_error = np.max(errors)
importance = [e/max_error for e in errors]
data = {'name':names,'error':errors,'importance':importance}
result = pd.DataFrame(data, columns = ['name','error','importance'])
result.sort_values(by=['importance'], ascending=[0], inplace=True)
result.reset_index(inplace=True, drop=True)
return result
# -
# ### Classification and Input Perturbation Ranking
#
# We now look at the code to perform perturbation ranking for a classification neural network. The implementation technique is slightly different for classification vs regression, so I must provide two different implementations. The primary difference between classification and regression is how we evaluate the accuracy of the neural network in each of these two network types. For regression neural networks, we will use the Root Mean Square (RMSE) error calculation; whereas, we will use log loss for classification.
#
# The code presented below creates a classification neural network that will predict for the classic iris dataset.
# +
import pandas as pd
import io
import requests
import numpy as np
from sklearn import metrics
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
from tensorflow.keras.callbacks import EarlyStopping
from sklearn.model_selection import train_test_split
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/iris.csv",
na_values=['NA', '?'])
# Convert to numpy - Classification
x = df[['sepal_l', 'sepal_w', 'petal_l', 'petal_w']].values
dummies = pd.get_dummies(df['species']) # Classification
species = dummies.columns
y = dummies.values
# Split into train/test
x_train, x_test, y_train, y_test = train_test_split(
x, y, test_size=0.25, random_state=42)
# Build neural network
model = Sequential()
model.add(Dense(50, input_dim=x.shape[1], activation='relu')) # Hidden 1
model.add(Dense(25, activation='relu')) # Hidden 2
model.add(Dense(y.shape[1],activation='softmax')) # Output
model.compile(loss='categorical_crossentropy', optimizer='adam')
model.fit(x_train,y_train,verbose=2,epochs=100)
# -
# Next, we evaluate the accuracy of the trained model. Here we see that the neural network is performing great, with an accuracy of 1.0. For a more complex dataset, we might fear overfitting with such high accuracy. However, for this example, we are more interested in determining the importance of each column.
# +
from sklearn.metrics import accuracy_score
pred = model.predict(x_test)
predict_classes = np.argmax(pred,axis=1)
expected_classes = np.argmax(y_test,axis=1)
correct = accuracy_score(expected_classes,predict_classes)
print(f"Accuracy: {correct}")
# -
# We are now ready to call the input perturbation algorithm. First, we extract the column names and remove the target column. The target column does not have importance, as it is the objective, not one of the inputs. In supervised learning, the target is of the utmost importance.
#
# We can see the importance displayed in the following table. The most important column is always 1.0, and lessor columns will continue in a downward trend. The least important column will have the lowest rank.
# +
# Rank the features
from IPython.display import display, HTML
names = list(df.columns) # x+y column names
names.remove("species") # remove the target(y)
rank = perturbation_rank(model, x_test, y_test, names, False)
display(rank)
# -
# ### Regression and Input Perturbation Ranking
#
# We now see how to use input perturbation ranking for a regression neural network. We will use the MPG dataset as a demonstration. The code below loads the MPG dataset and creates a regression neural network for this dataset. The code trains the neural network and calculates an RMSE evaluation.
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
from sklearn.model_selection import train_test_split
import pandas as pd
import io
import os
import requests
import numpy as np
from sklearn import metrics
save_path = "."
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA', '?'])
cars = df['name']
# Handle missing value
df['horsepower'] = df['horsepower'].fillna(df['horsepower'].median())
# Pandas to Numpy
x = df[['cylinders', 'displacement', 'horsepower', 'weight',
'acceleration', 'year', 'origin']].values
y = df['mpg'].values # regression
# Split into train/test
x_train, x_test, y_train, y_test = train_test_split(
x, y, test_size=0.25, random_state=42)
# Build the neural network
model = Sequential()
model.add(Dense(25, input_dim=x.shape[1], activation='relu')) # Hidden 1
model.add(Dense(10, activation='relu')) # Hidden 2
model.add(Dense(1)) # Output
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(x_train,y_train,verbose=2,epochs=100)
# Predict
pred = model.predict(x)
# -
# Just as before, we extract the column names and discard the target. We can now create a ranking of the importance of each of the input features. The feature with a ranking of 1.0 is the most important.
# +
# Rank the features
from IPython.display import display, HTML
names = list(df.columns) # x+y column names
names.remove("name")
names.remove("mpg") # remove the target(y)
rank = perturbation_rank(model, x_test, y_test, names, True)
display(rank)
# -
# ### Biological Response with Neural Network
#
# The following sections will demonstrate how to use feature importance ranking and ensembling with a more complex dataset. Ensembling is the process where you combine multiple models for greater accuracy. Kaggle competition winners frequently make use of ensembling for high ranking solutions.
#
# We will use the biological response dataset, a Kaggle dataset, where there is an unusually high number of columns. Because of the large number of columns, it is essential to use feature ranking to determine the importance of these columns. We begin by loading the dataset and preprocessing. This Kaggle dataset is a binary classification problem. You must predict if certain conditions will cause a biological response.
#
# * [Predicting a Biological Response](https://www.kaggle.com/c/bioresponse)
# +
import pandas as pd
import os
import numpy as np
from sklearn import metrics
from scipy.stats import zscore
from sklearn.model_selection import KFold
from IPython.display import HTML, display
if COLAB:
path = "/content/drive/My Drive/data/"
else:
path = "./data/"
filename_train = os.path.join(path,"bio_train.csv")
filename_test = os.path.join(path,"bio_test.csv")
filename_submit = os.path.join(path,"bio_submit.csv")
df_train = pd.read_csv(filename_train,na_values=['NA','?'])
df_test = pd.read_csv(filename_test,na_values=['NA','?'])
activity_classes = df_train['Activity']
# -
# A large number of columns is evident when we display the shape of the dataset.
print(df_train.shape)
# The following code constructs a classification neural network and trains it for the biological response dataset. Once trained, the accuracy is measured.
# +
import os
import pandas as pd
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
from sklearn.model_selection import train_test_split
from tensorflow.keras.callbacks import EarlyStopping
import numpy as np
import sklearn
# Encode feature vector
# Convert to numpy - Classification
x_columns = df_train.columns.drop('Activity')
x = df_train[x_columns].values
y = df_train['Activity'].values # Classification
x_submit = df_test[x_columns].values.astype(np.float32)
# Split into train/test
x_train, x_test, y_train, y_test = train_test_split(
x, y, test_size=0.25, random_state=42)
print("Fitting/Training...")
model = Sequential()
model.add(Dense(25, input_dim=x.shape[1], activation='relu'))
model.add(Dense(10))
model.add(Dense(1,activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam')
monitor = EarlyStopping(monitor='val_loss', min_delta=1e-3,
patience=5, verbose=1, mode='auto')
model.fit(x_train,y_train,validation_data=(x_test,y_test),
callbacks=[monitor],verbose=0,epochs=1000)
print("Fitting done...")
# Predict
pred = model.predict(x_test).flatten()
# Clip so that min is never exactly 0, max never 1
pred = np.clip(pred,a_min=1e-6,a_max=(1-1e-6))
print("Validation logloss: {}".format(
sklearn.metrics.log_loss(y_test,pred)))
# Evaluate success using accuracy
pred = pred>0.5 # If greater than 0.5 probability, then true
score = metrics.accuracy_score(y_test, pred)
print("Validation accuracy score: {}".format(score))
# Build real submit file
pred_submit = model.predict(x_submit)
# Clip so that min is never exactly 0, max never 1 (would be a NaN score)
pred = np.clip(pred,a_min=1e-6,a_max=(1-1e-6))
submit_df = pd.DataFrame({'MoleculeId':[x+1 for x \
in range(len(pred_submit))],'PredictedProbability':\
pred_submit.flatten()})
submit_df.to_csv(filename_submit, index=False)
# -
# ### What Features/Columns are Important
# The following uses perturbation ranking to evaluate the neural network.
# +
# Rank the features
from IPython.display import display, HTML
names = list(df_train.columns) # x+y column names
names.remove("Activity") # remove the target(y)
rank = perturbation_rank(model, x_test, y_test, names, False)
display(rank)
# -
# ### Neural Network Ensemble
#
# A neural network ensemble combines neural network predictions with other models. The program determines the exact blend of all of these models by logistic regression. The following code performs this blend for a classification. If you present the final predictions from the ensemble to Kaggle, you will see that the result is very accurate.
# +
import numpy as np
import os
import pandas as pd
import math
from tensorflow.keras.wrappers.scikit_learn import KerasClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import StratifiedKFold
from sklearn.ensemble import RandomForestClassifier,
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.linear_model import LogisticRegression
PATH = "./data/"
SHUFFLE = False
FOLDS = 10
def build_ann(input_size,classes,neurons):
model = Sequential()
model.add(Dense(neurons, input_dim=input_size, activation='relu'))
model.add(Dense(1))
model.add(Dense(classes,activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam')
return model
def mlogloss(y_test, preds):
epsilon = 1e-15
sum = 0
for row in zip(preds,y_test):
x = row[0][row[1]]
x = max(epsilon,x)
x = min(1-epsilon,x)
sum+=math.log(x)
return( (-1/len(preds))*sum)
def stretch(y):
return (y - y.min()) / (y.max() - y.min())
def blend_ensemble(x, y, x_submit):
kf = StratifiedKFold(FOLDS)
folds = list(kf.split(x,y))
models = [
KerasClassifier(build_fn=build_ann,neurons=20,
input_size=x.shape[1],classes=2),
KNeighborsClassifier(n_neighbors=3),
RandomForestClassifier(n_estimators=100, n_jobs=-1,
criterion='gini'),
RandomForestClassifier(n_estimators=100, n_jobs=-1,
criterion='entropy'),
ExtraTreesClassifier(n_estimators=100, n_jobs=-1,
criterion='gini'),
ExtraTreesClassifier(n_estimators=100, n_jobs=-1,
criterion='entropy'),
GradientBoostingClassifier(learning_rate=0.05,
subsample=0.5, max_depth=6, n_estimators=50)]
dataset_blend_train = np.zeros((x.shape[0], len(models)))
dataset_blend_test = np.zeros((x_submit.shape[0], len(models)))
for j, model in enumerate(models):
print("Model: {} : {}".format(j, model) )
fold_sums = np.zeros((x_submit.shape[0], len(folds)))
total_loss = 0
for i, (train, test) in enumerate(folds):
x_train = x[train]
y_train = y[train]
x_test = x[test]
y_test = y[test]
model.fit(x_train, y_train)
pred = np.array(model.predict_proba(x_test))
# pred = model.predict_proba(x_test)
dataset_blend_train[test, j] = pred[:, 1]
pred2 = np.array(model.predict_proba(x_submit))
#fold_sums[:, i] = model.predict_proba(x_submit)[:, 1]
fold_sums[:, i] = pred2[:, 1]
loss = mlogloss(y_test, pred)
total_loss+=loss
print("Fold #{}: loss={}".format(i,loss))
print("{}: Mean loss={}".format(model.__class__.__name__,
total_loss/len(folds)))
dataset_blend_test[:, j] = fold_sums.mean(1)
print()
print("Blending models.")
blend = LogisticRegression(solver='lbfgs')
blend.fit(dataset_blend_train, y)
return blend.predict_proba(dataset_blend_test)
if __name__ == '__main__':
np.random.seed(42) # seed to shuffle the train set
print("Loading data...")
filename_train = os.path.join(PATH, "bio_train.csv")
df_train = pd.read_csv(filename_train, na_values=['NA', '?'])
filename_submit = os.path.join(PATH, "bio_test.csv")
df_submit = pd.read_csv(filename_submit, na_values=['NA', '?'])
predictors = list(df_train.columns.values)
predictors.remove('Activity')
x = df_train[predictors].values
y = df_train['Activity']
x_submit = df_submit.values
if SHUFFLE:
idx = np.random.permutation(y.size)
x = x[idx]
y = y[idx]
submit_data = blend_ensemble(x, y, x_submit)
submit_data = stretch(submit_data)
####################
# Build submit file
####################
ids = [id+1 for id in range(submit_data.shape[0])]
submit_filename = os.path.join(PATH, "bio_submit.csv")
submit_df = pd.DataFrame({'MoleculeId': ids,
'PredictedProbability':
submit_data[:, 1]},
columns=['MoleculeId',
'PredictedProbability'])
submit_df.to_csv(submit_filename, index=False)
# -
| t81_558_class_08_2_keras_ensembles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
time = np.array([3.4365310668945312,
1.8143439292907715,
1.2134535312652588,
1.011009931564331,
0.7168467044830322,
0.669912576675415,
0.5913188457489014,
0.5408780574798584,
0.48679542541503906,
0.44424986839294434,
0.415529727935791,
0.3681631088256836,
0.35811352729797363,
0.2830197811126709,
0.24335241317749023,
0.2270965576171875,
0.2328789234161377,
0.251446008682251,
0.2357335090637207,
0.23085975646972656])
n = [i for i in range(1, 21)]
plt.plot(n, time)
plt.xlabel('Number of Cores')
plt.ylabel('Computation Time')
plt.title('Computation Time VS Number of Cores')
plt.savefig("health_simulation_mpi.png")
| Assignment 1/Assignment1_Q1(a).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Za8-Nr5k11fh"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="Eq10uEbw0E4l" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="nuRx7K-sirJr"
# # Forecasting with an RNN
# + [markdown] colab_type="text" id="97jsq1rHh2Ds"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c06_forecasting_with_rnn.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c06_forecasting_with_rnn.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="vidayERjaO5q"
# ## Setup
# + colab_type="code" id="m_rX1u_nv0Nn" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
# + colab_type="code" id="m-I80IS32DBl" colab={}
try:
# Use the %tensorflow_version magic if in colab.
# %tensorflow_version 2.x
except Exception:
pass
# + colab_type="code" id="gqWabzlJ63nL" colab={}
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
keras = tf.keras
# + colab_type="code" id="cg1hfKCPldZG" colab={}
def plot_series(time, series, format="-", start=0, end=None, label=None):
plt.plot(time[start:end], series[start:end], format, label=label)
plt.xlabel("Time")
plt.ylabel("Value")
if label:
plt.legend(fontsize=14)
plt.grid(True)
def trend(time, slope=0):
return slope * time
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def white_noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
def window_dataset(series, window_size, batch_size=32,
shuffle_buffer=1000):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(window_size + 1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_size + 1))
dataset = dataset.shuffle(shuffle_buffer)
dataset = dataset.map(lambda window: (window[:-1], window[-1]))
dataset = dataset.batch(batch_size).prefetch(1)
return dataset
def model_forecast(model, series, window_size):
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size))
ds = ds.batch(32).prefetch(1)
forecast = model.predict(ds)
return forecast
# + colab_type="code" id="iL2DDjV3lel6" colab={}
time = np.arange(4 * 365 + 1)
slope = 0.05
baseline = 10
amplitude = 40
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
noise_level = 5
noise = white_noise(time, noise_level, seed=42)
series += noise
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
# + colab_type="code" id="Zmp1JXKxk9Vb" colab={}
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
# + [markdown] colab_type="text" id="vDs_w3kZ8OIw"
# ## Simple RNN Forecasting
# + colab_type="code" id="YU4xRp9G8OIx" colab={}
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = window_dataset(x_train, window_size, batch_size=128)
model = keras.models.Sequential([
keras.layers.Lambda(lambda x: tf.expand_dims(x, axis=-1),
input_shape=[None]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.SimpleRNN(100),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
lr_schedule = keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-7 * 10**(epoch / 20))
optimizer = keras.optimizers.SGD(lr=1e-7, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
# + colab_type="code" id="YJTlFAXF8OIy" colab={}
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-7, 1e-4, 0, 30])
# + colab_type="code" id="T3yNjxWE8OIz" colab={}
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = window_dataset(x_train, window_size, batch_size=128)
valid_set = window_dataset(x_valid, window_size, batch_size=128)
model = keras.models.Sequential([
keras.layers.Lambda(lambda x: tf.expand_dims(x, axis=-1),
input_shape=[None]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.SimpleRNN(100),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
optimizer = keras.optimizers.SGD(lr=1.5e-6, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
early_stopping = keras.callbacks.EarlyStopping(patience=50)
model_checkpoint = keras.callbacks.ModelCheckpoint(
"my_checkpoint", save_best_only=True)
model.fit(train_set, epochs=500,
validation_data=valid_set,
callbacks=[early_stopping, model_checkpoint])
# + colab_type="code" id="4KuPtKFe8OI0" colab={}
model = keras.models.load_model("my_checkpoint")
# + colab_type="code" id="cxq09qOg8OI1" colab={}
rnn_forecast = model_forecast(
model,
series[split_time - window_size:-1],
window_size)[:, 0]
# + colab_type="code" id="PkC_JssS8OI2" colab={}
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
# + colab_type="code" id="1mwfgEK08OI3" colab={}
keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
# + [markdown] colab_type="text" id="KNG7s8jt8OI4"
# ## Sequence-to-Sequence Forecasting
# + colab_type="code" id="bsKGxfiE8OI4" colab={}
def seq2seq_window_dataset(series, window_size, batch_size=32,
shuffle_buffer=1000):
series = tf.expand_dims(series, axis=-1)
ds = tf.data.Dataset.from_tensor_slices(series)
ds = ds.window(window_size + 1, shift=1, drop_remainder=True)
ds = ds.flat_map(lambda w: w.batch(window_size + 1))
ds = ds.shuffle(shuffle_buffer)
ds = ds.map(lambda w: (w[:-1], w[1:]))
return ds.batch(batch_size).prefetch(1)
# + colab_type="code" id="5Nk2C7WP8OI5" colab={}
for X_batch, Y_batch in seq2seq_window_dataset(tf.range(10), 3,
batch_size=1):
print("X:", X_batch.numpy())
print("Y:", Y_batch.numpy())
# + colab_type="code" id="4JSc-Btk8OI7" colab={}
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = seq2seq_window_dataset(x_train, window_size,
batch_size=128)
model = keras.models.Sequential([
keras.layers.SimpleRNN(100, return_sequences=True,
input_shape=[None, 1]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200)
])
lr_schedule = keras.callbacks.LearningRateScheduler(
lambda epoch: 1e-7 * 10**(epoch / 30))
optimizer = keras.optimizers.SGD(lr=1e-7, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
history = model.fit(train_set, epochs=100, callbacks=[lr_schedule])
# + colab_type="code" id="YGNsWceq8OI8" colab={}
plt.semilogx(history.history["lr"], history.history["loss"])
plt.axis([1e-7, 1e-4, 0, 30])
# + colab_type="code" id="G9lDnb0X8OI9" colab={}
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
window_size = 30
train_set = seq2seq_window_dataset(x_train, window_size,
batch_size=128)
valid_set = seq2seq_window_dataset(x_valid, window_size,
batch_size=128)
model = keras.models.Sequential([
keras.layers.SimpleRNN(100, return_sequences=True,
input_shape=[None, 1]),
keras.layers.SimpleRNN(100, return_sequences=True),
keras.layers.Dense(1),
keras.layers.Lambda(lambda x: x * 200.0)
])
optimizer = keras.optimizers.SGD(lr=1e-6, momentum=0.9)
model.compile(loss=keras.losses.Huber(),
optimizer=optimizer,
metrics=["mae"])
early_stopping = keras.callbacks.EarlyStopping(patience=10)
model.fit(train_set, epochs=500,
validation_data=valid_set,
callbacks=[early_stopping])
# + colab_type="code" id="4mglBRex8OI_" colab={}
rnn_forecast = model_forecast(model, series[..., np.newaxis], window_size)
rnn_forecast = rnn_forecast[split_time - window_size:-1, -1, 0]
# + colab_type="code" id="Zl_FkcdI8OJA" colab={}
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, rnn_forecast)
# + colab_type="code" id="cznEtSVK8OJB" colab={}
keras.metrics.mean_absolute_error(x_valid, rnn_forecast).numpy()
| Intro-to-TensorFlow/Time-Series-Forecasting/forecasting_with_rnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## What is Python? Why are we going to use it?
# Python is an interpreted high-level language; other high-level languages you might have heard of are C, C++, Perl, and Java.
# You can go deep and far with any programming language, but we wanted to teach Python because...
# * We want you to focus on programming, not learning a programming language
# * Struggle less, Learn more
# * Have more fun!
# * Python is general purpose and allows you to work on all kinds of interesting projects
# * Data science
# * Scientific and mathematical computing
# * Web development
# * Finance and trading
# * System automation and administration
# * Computer graphics
# * Basic game development
# * Security and penetration testing
# * General and application-specific scripting
# * Mapping and geography (GIS software)
# * Python is widely used in data analysis/ data science
# ## Python Expressions
# **Value**: what a program has to work with.
#
# There are different types of values
# String, Int, Float
type("Hello, World!")
type(3)
type(3.14)
# **Variable**, a name that points to a value
#
# Variable names in Python have some restrictions...
# Python is wonderful
a = "Today"
a
7coolthings = "sevencoolthings"
awesome@ = 45
class = "my bootcamp"
# Example:
# If you run a 10 kilometer race in 43 minutes 30 seconds, what is your average time per mile? What is your average speed in miles per hour? (Hint: there are 1.61 kilometers in a mile).
kilometers = 10
kilom_per_mile = 1.61
distance_in_miles = kilometers / kilom_per_mile
distance_in_miles
# Average time per mile
43.5 / distance_in_miles
time_in_hours = 43.5 / 60
distance_in_miles / time_in_hours
# ## Python Data types (what you'll be working with during this bootcamp)
# From the [great talk](https://youtu.be/j6VSAsKAj98?list=LLnOPU_YQJoz4VlqjCDgsBAQ) by <NAME>ley
# ## *Tuple*
# AKA:
# >`Record`
#
# KNOWN FOR:
# > A row in a database. Packing and unpacking things.
#
# BASIC USAGE:
# > `record = (val1, val2, val3)`
# > `a, b, c = record`
# > `val = record[n]`
#
# ACCOMPLICE:
# > `collections.namedtuple`
row = ("Bob", "Davis", "2391 695 UT", 84121)
row[3]
from collections import namedtuple
Person = namedtuple("Person", ["first", "last", "address", "zip"])
row = Person("Sam", "Douglas", "2391 E 695 S SLC UT", 84121)
row.last
# ## *List*
# AKA:
# >`Mutable Sequence, Array`
#
# KNOWN FOR:
# > Enforcing order.
#
# BASIC USAGE:
# > `items = [val1, val2, val3,..., valn]`
# > `x = items[n]`
# > `del items[n]`
# > `items.append(value)`
# > `items.sort()`
my_list = [7, "john", "hello world", 2.4]
my_list
my_list.append("add me too!")
my_list
my_list.insert(2, "eh?")
my_list
for x in my_list:
print("The number " + str(x))
lower_names = ["judy", "charlie", "sam", "chuck", "jim", "sandy"]
for name in lower_names:
print(name.upper())
# ## *Set*
# AKA:
# >`Set`
#
# KNOWN FOR:
# > Uniqueness, membership tests.
#
# BASIC USAGE:
# > `s = {val1, val2, ..., valn}`
# > `s.add(val)`
# > `s.remove(val)`
# > `val in s`
#
names = ["Dave", "Nate", "Carol", "John", "Dave"]
names
set(names)
names_two = {"Dave", "Nate", "carol", "John", "Amber", "Dave"}
names_two
# ### *Dict*
# AKA:
# >`Mapping, Associative Array`
#
# KNOWN FOR:
# > Lookup tables, indices.
#
# BASIC USAGE:
# > `d = {key1: val1, key2: val2, key3: val3}`
# > `val = d[key]`
# > `d[key] = val`
# > `del d[key]`
# > `key in d`
#
prices = {
"ACME": 45.23,
"YOW": {"key1": "val1", "key2": "val2", "key3": "val3"},
"APPL": ["Dave", "Nate", "Carol", "John", "Dave"]
}
prices['APPL']
prices["YOW"]["key1"]
# ### *Counter*
# AKA:
# >`collections.Counter`
#
# KNOWN FOR:
# > Counting, Histograms, Tabluation.
#
# BASIC USAGE:
# > `c = Counter(sequence)`
# > `c[key] += n`
# > `c.most_common(n)`
from collections import Counter
c = Counter("xxyxxzzt")
c
c['a'] += 10
c
# ### *Defaultdict*
# AKA:
# >`collections.defaultdict`
#
# KNOWN FOR:
# > Multidicts, One-to-many relationships, Grouping.
#
# BASIC USAGE:
# > `d = defaultdict(list)`
# > `d[key].append(val)`
# > `values = d[key]`
#
from collections import defaultdict
d = defaultdict(list)
d['stuff'].append(19)
d['stuff'].append(34)
d['spam'].append("hi!")
d
# ### OrderderDict
# AKA
# `collections.OrderedDict`
#
# KNOWN FOR:
# `An dictionary which keeps the order of the items`
#
# BASIC USAGE:
# `d = OrderedDict.fromkeys('abcde')
# d.move_to_end('b')`
from collections import OrderedDict
d = OrderedDict.fromkeys('ABCDE')
d
d.move_to_end('B')
d
d['C'] = "Hello World"
d
| day1/2. Lab/.ipynb_checkpoints/introduction_to_python-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
H = int(input())
# ピラミッドの深さを求める
count = 1
while H//2 != 0:
count += 1
H = H//2
# 深さに応じた戦闘回数を求める
ret = 0
for c in range(count):
ret += 2**c
print(ret)
# -
| 2020/20201217_ABC153_D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''py3.9'': conda)'
# name: python3
# ---
# for good measure
# %config Completer.use_jedi = False
# +
from ml_algorithms.linear_regression import LinearRegression
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn import datasets
# -
# some handy functions we need
def mse(y_true, y_pred):
return np.mean((y_true-y_pred)**2)
# +
X, y = datasets.make_regression(
n_samples=1000, n_features=1, noise=10, random_state=42
)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=42
)
# -
linear_regressor = LinearRegression(lr=0.01, n_iters=10000)
linear_regressor.fit(X_train, y_train)
predictions = linear_regressor.predict(X_test)
mse = mse(y_test, predictions)
print('MSE: %f' % mse)
corr_matrix = np.corrcoef(y_test, predictions)
corr = corr_matrix[0, 1]**2
print('Corr: ', corr)
y_pred_line = linear_regressor.predict(X)
# cmap = plt.get_cmap("viridis")
fig = plt.figure(figsize=(8, 6))
m1 = plt.scatter(X_train, y_train, s=10)
m2 = plt.scatter(X_test, y_test, s=10)
plt.plot(X, y_pred_line, color="black", linewidth=2, label="Prediction")
plt.show()
| notebooks/linear_regression_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import ipywidgets
from ipywidgets import FloatProgress
import mne
from mne.datasets.sleep_physionet.age import fetch_data
from mne.time_frequency import psd_welch
result = []
for i in range(0,100): #2 to test
if i not in [36, 39, 52, 68, 69, 78, 79]:
try:
subject = fetch_data(subjects = [i], recording = [1])
raw_train = mne.io.read_raw_edf(subject[0][0])
annot_train = mne.read_annotations(subject[0][1])
raw_train.set_annotations(annot_train, emit_warning=False)
annotation_desc_2_event_id = {'Sleep stage W': 1,
'Sleep stage 1': 2,
'Sleep stage 2': 3,
'Sleep stage 3': 4,
'Sleep stage 4': 4,
'Sleep stage R': 5}
# keep last 30-min wake events before sleep and first 30-min wake events after
# sleep and redefine annotations on raw data
annot_train.crop(annot_train[1]['onset'] - 30 * 60,
annot_train[-2]['onset'] + 30 * 60)
raw_train.set_annotations(annot_train, emit_warning=False)
events_train, _ = mne.events_from_annotations(
raw_train, event_id=annotation_desc_2_event_id, chunk_duration=30.)
# create a new event_id that unifies stages 3 and 4
event_id = {'Sleep stage W': 1,
'Sleep stage 1': 2,
'Sleep stage 2': 3,
'Sleep stage 3/4': 4,
'Sleep stage R': 5}
tmax = 30. - 1. / raw_train.info['sfreq'] # tmax in included
epochs_train = mne.Epochs(raw=raw_train, events=events_train,
event_id=event_id, tmin=0., tmax=tmax, baseline=None)
index, scalings = ['epoch'], dict(eeg=1e6, mag=1e15, grad=1e13)
df = epochs_train.to_data_frame(picks=None, scalings=scalings)
#take first condition value per epoch (is there 1 epoch to condition? not sure)
df = pd.DataFrame(df.groupby('epoch').first()['condition']).merge(df.groupby('epoch').mean().drop('time', axis = 1), left_index = True, right_index = True, how = 'inner')
df['patient_id'] = i + 1
result.append(df)
except:
pass
result = pd.concat(result)
result[['patient_id','condition', 'EEG Fpz-Cz', 'EEG Pz-Oz', 'EOG horizontal',
'Resp oro-nasal', 'EMG submental', 'Temp rectal', 'Event marker']].to_csv('sleep_edf_raw.csv')
| notebooks/physio_edf data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# default_exp predefined_problems.ner_data
# %load_ext autoreload
# %autoreload 2
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
# # Pre-defined Problems
#
# Preprocessing functions of pre-defined problems.
# +
# export
from glob import glob
import re
import random
from sklearn.model_selection import train_test_split
from m3tl.utils import cluster_alphnum
from m3tl.preproc_decorator import preprocessing_fn
NER_TYPE = ['LOC', # location
'GPE',
'PER', # person
'ORG', # organization
'PRD', # Product
]
def gold_horse_ent_type_process_fn(d):
"""golden horse ent type process fn
Source: https://github.com/hltcoe/golden-ho rse
Entity type:
B, I, O: Begining \ In middle \ Outside of entity
GPE: Country, City, District...
LOC: Location, zoo, school...
PER: Person
ORG: Organiazation
NAM: Entity
NOM: More general, 女生, 男的...
Example:
B-PER.NAM
Only keep NAM here
So after process:
B-PER
Arguments:
ent_type {str} -- ent type from gold_horse data
Returns:
str -- processed enttype
"""
ent_type = d.split('\t')[1].replace('\n', '')
# keep nam only
ent_type = ent_type if 'NAM' in ent_type else 'O'
ent_type = ent_type.replace('.NAM', '')
return ent_type
def chinese_literature_ent_type_process_fn(d):
"""Not match my need
Arguments:
d {[type]} -- [description]
Returns:
[type] -- [description]
"""
ent_type = d.split(' ')[1].replace('\n', '')
return ent_type
def read_ner_data(file_pattern='data/ner/weiboNER*', proc_fn=None):
"""Read data from golden horse data
Arguments:
file_pattern {str} -- file patterns
Returns:
dict -- dict, key: 'train', 'eval', value: dict {'inputs', 'target'}
"""
result_dict = {
'train': {
'inputs': [],
'target': []
},
'eval': {
'inputs': [],
'target': []
}
}
file_list = glob(file_pattern)
for file_path in file_list:
with open(file_path, 'r', encoding='utf8') as f:
raw_data = f.readlines()
inputs_list = [[]]
target_list = [[]]
for d in raw_data:
if d != '\n':
# put first char to input
inputs_list[-1].append(d[0])
ent_type = proc_fn(d)
target_list[-1].append(ent_type)
else:
inputs_list.append([])
target_list.append([])
# remove trailing empty str/list
if not inputs_list[-1]:
del inputs_list[-1]
if not target_list[-1]:
del target_list[-1]
inputs_with_ent = []
target_with_ent = []
for inputs, target in zip(inputs_list, target_list):
# if len(set(target)) > 1:
inputs_with_ent.append(inputs)
target_with_ent.append(target)
if 'train' in file_path or 'dev' in file_path:
result_dict['train']['inputs'] = inputs_with_ent
result_dict['train']['target'] = target_with_ent
else:
result_dict['eval']['inputs'] = inputs_with_ent
result_dict['eval']['target'] = target_with_ent
return result_dict
def get_weibo_ner_fn(file_path):
@preprocessing_fn
def weibo_ner(params, mode):
data = read_ner_data(file_pattern=file_path,
proc_fn=gold_horse_ent_type_process_fn)
if mode == 'train':
data = data['train']
else:
data = data['eval']
inputs_list = data['inputs']
target_list = data['target']
return inputs_list, target_list
return weibo_ner
def gold_horse_segment_process_fn(d):
ent_type = d.split('\t')[0][-1]
if ent_type not in ['0', '1', '2']:
ent_type = '0'
return ent_type
def get_weibo_cws_fn(file_path):
@preprocessing_fn
def weibo_cws(params, mode):
data = read_ner_data(file_pattern=file_path,
proc_fn=gold_horse_segment_process_fn)
if mode == 'train':
data = data['train']
else:
data = data['eval']
inputs_list = data['inputs']
target_list = data['target']
return inputs_list, target_list
return weibo_cws
def read_bosonnlp_data(file_pattern, eval_size=0.2):
file_list = glob(file_pattern)
sentence_split = r'[!?。?!]'
project_table = {
'person_name': 'PER',
'company_name': 'ORG',
'location': 'LOC',
'product_name': 'PRD',
'time': 'TME',
'org_name': 'ORG2'
}
input_list = []
target_list = []
if not file_list:
raise FileNotFoundError('Please make sure you have downloaded BosonNLP\
data and put it in the path you specified. \
Download: https://bosonnlp.com/resources/BosonNLP_NER_6C.zip')
for file_path in file_list:
with open(file_path, 'r', encoding='utf8') as f:
data_list = f.readlines()
for doc in data_list:
if '}}}}' in doc:
continue
splited_doc = re.split(sentence_split, doc)
for sentence in splited_doc:
# split doc into sentences
input_list.append([])
target_list.append([])
# split by {{
doc_chunk_list = sentence.split('{{')
for chunk in doc_chunk_list:
if '}}' not in chunk or ':' not in chunk:
target_list[-1] += ['O']*len(chunk)
input_list[-1] += list(chunk)
else:
ent_chunk, text_chunk = chunk.split('}}')
punc_ind = ent_chunk.index(':')
ent_type = ent_chunk[:punc_ind]
ent = ent_chunk[punc_ind+1:]
if ent_type in project_table:
ent = cluster_alphnum(ent)
for char_ind, ent_char in enumerate(ent):
if char_ind == 0:
loc_char = 'B'
else:
loc_char = 'I'
target_list[-1].append(loc_char +
'-'+project_table[ent_type])
input_list[-1].append(ent_char)
else:
target_list[-1] += ['O']*len(ent)
input_list[-1] += list(ent)
target_list[-1] += ['O']*len(text_chunk)
input_list[-1] += list(text_chunk)
return_input, return_target = [], []
for inp, tar in zip(input_list, target_list):
if inp and tar:
return_input.append(inp)
return_target.append(tar)
assert len(inp) == len(tar)
train_input, eval_input, train_target, eval_target = train_test_split(
return_input, return_target, test_size=eval_size, random_state=1024)
result_dict = {
'train': {},
'eval': {}
}
result_dict['train']['inputs'] = train_input
result_dict['train']['target'] = train_target
result_dict['eval']['inputs'] = eval_input
result_dict['eval']['target'] = eval_target
return result_dict
def read_msra(file_pattern, eval_size):
file_list = glob(file_pattern)
project_table = {
'nr': 'PER',
'nt': 'ORG',
'ns': 'LOC'
}
input_list = []
target_list = []
for file_path in file_list:
with open(file_path, 'r', encoding='utf8') as f:
data_list = f.readlines()
for sentence in data_list:
sentence = sentence.replace('\n', '')
input_list.append([])
target_list.append([])
sentence_word_list = sentence.split(' ')
for word in sentence_word_list:
if word:
ent, ent_type = word.split('/')
ent = cluster_alphnum(ent)
if ent_type not in project_table:
input_list[-1] += list(ent)
target_list[-1] += ['O'] * len(ent)
else:
for char_ind, ent_char in enumerate(ent):
if char_ind == 0:
loc_char = 'B'
else:
loc_char = 'I'
target_list[-1].append(loc_char +
'-'+project_table[ent_type])
input_list[-1].append(ent_char)
return_input, return_target = [], []
for inp, tar in zip(input_list, target_list):
if inp and tar:
return_input.append(inp)
return_target.append(tar)
assert len(inp) == len(tar)
train_input, eval_input, train_target, eval_target = train_test_split(
return_input, return_target, test_size=eval_size, random_state=1024)
result_dict = {
'train': {},
'eval': {}
}
result_dict['train']['inputs'] = train_input
result_dict['train']['target'] = train_target
result_dict['eval']['inputs'] = eval_input
result_dict['eval']['target'] = eval_target
return result_dict
def get_msra_ner_fn(file_path):
@preprocessing_fn
def msra_ner(params, mode):
msra_data = read_msra(
file_pattern=file_path, eval_size=0.2)
inputs_list = []
target_list = []
for data in [msra_data]:
if mode == 'train':
inputs_list += data['train']['inputs']
target_list += data['train']['target']
else:
inputs_list += data['eval']['inputs']
target_list += data['eval']['target']
return inputs_list, target_list
return msra_ner
def get_boson_ner_fn(file_path):
@preprocessing_fn
def boson_ner(params, mode):
boson_data = read_bosonnlp_data(
file_pattern=file_path, eval_size=0.2)
inputs_list = []
target_list = []
for data in [boson_data]:
if mode == 'train':
inputs_list += data['train']['inputs']
target_list += data['train']['target']
else:
inputs_list += data['eval']['inputs']
target_list += data['eval']['target']
return inputs_list, target_list
return boson_ner
# -
| source_nbs/09_predefined_problems_ner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # SELECT Tutorial
#
#
# MLDB comes with a powerful implementation of [SQL's `SELECT` Queries](../../../../doc/#builtin/sql/Sql.md.html). This tutorial will walk you through the basics of `SELECT`, and some MLDB-specific features.
#
# The notebook cells below use `pymldb`'s `.query()` method; you can check out the [Using `pymldb` Tutorial](../../../../doc/nblink.html#_tutorials/Using pymldb Tutorial) for more details.
from pymldb import Connection
mldb = Connection()
# # `SELECT`
#
# All queries start with the keyword `SELECT`. Here is the simplest possible query: we ask for 1 and we get a very short result set consisting of one row with one column named 1 and the single cell in it also contains 1.
mldb.query("""
select 1
""")
# Of course we can ask for more: the query below does a little math and shows how you can rename your columns with the `as` keyword. Note that single-quotes (`'`) are used to denote strings and double-quotes (`"`) denote column names, both of which can contain any Unicode character.
mldb.query("""
select 1+1, 3+4 as var, 'UTF8 striñg' as "hello, François"
""")
# We can use a variety of operators in a `SELECT` expression, like this:
mldb.query("""
select
1 between 0 and 2,
2 in (1,2,3),
3 is integer,
(case when 4<5 then 'yes' else 'no' end)
""")
# # `FROM` and `LIMIT`
#
# Queries are mostly useful when run on actual datasets, so let's import part of the passenger manifest from the Titanic.
print mldb.put('/v1/procedures/import_titanic', {
"type": "import.text",
"params": {
"dataFileUrl": "http://public.mldb.ai/titanic_train.csv",
"outputDataset": "titanic",
"runOnCreation": True
}
})
# Now let's query all columns with the star (`*`) operator `FROM` our `titanic` dataset, using the `LIMIT` keyword to avoid getting too much output.
mldb.query("""
select *
from titanic
limit 10
""")
# We can also ask for just certain columns by name.
mldb.query("""
select Name, Age
from titanic
limit 10
""")
# # `ORDER BY`
#
# When we've used the `LIMIT` keyword above, we were just getting an arbitrary set of 10 rows. Using the `ORDER BY` keyword we can ask for the 'top 10' according to some criterion, for example `Age`.
mldb.query("""
select Name, Age
from titanic
order by Age desc
limit 10
""")
# # `WHERE`
#
# Beyond limiting the number of records, sometimes we want to look at records which match certain criteria, which we can do with the `WHERE` keyword. You can use the same operators in the `WHERE` clause as in the `SELECT` clause.
mldb.query("""
select Name, Age, Pclass, Sex, SibSp, Parch, label
from titanic
where Pclass in (1,3) and Sex='female' and (SibSp>3 or Parch=2) and label=1 and Age is null
""")
# In the query above we used the special operator `is` to retrieve only rows where `Age is null`. This is worth pointing out because `null` is a special value in SQL: it means "unknown". `null` has some strange properties, as you can see below: any comparison between `Age` and 1 returns `null`. This makes sense because if, say, `Age` is unknown, then we don't know if `Age` is less than, equal to or greater than anything else. SQL works according to [3-valued logic](https://en.wikipedia.org/wiki/Null_(SQL)).
#
# The only reliable way to check if a value is null is with the `is null` operator.
mldb.query("""
select Age, Age = 1, Age < 1, Age > 1, Age + 1, Age / 1, Age is null, Age is not null
from titanic
where Age is null
limit 1
""")
# # Functions and Aggregate Functions
#
# MLDB comes with a number of builtin functions to operate on your data. Here's an example where we convert a string to uppercase and lowercase.
mldb.query("""
select Name, upper(Name), lower(Name)
from titanic
order by Age desc limit 10
""")
# The functions below are special: they're aggregate functions, so they operate on multiple rows and give you a single output. They operate only on non-`null` values of their input.
mldb.query("""
select count(Age), sum(Age), sum(Age)/count(Age), avg(Age), min(Age), max(Age)
from titanic
""")
# The `count` aggregate function is special in that it accepts `*` as an input, in which case it will return the count of all rows:
mldb.query("""
select count(*)
from titanic
""")
# # GROUP BY & HAVING
#
# You can get aggregate functions to return multiple rows by grouping the input according to some criteria with the `GROUP BY` keyword. If you use an aggregate function in your `SELECT` clause, then you cannot use any non-aggregate expressions unless they appear in a `GROUP BY` clause.
mldb.query("""
select Pclass, avg(Age)
from titanic
group by Pclass
""")
# You cannot use aggregate functions in a `WHERE` clause. The `HAVING` clause is a little bit like a `WHERE` clause which is applied after `GROUP BY`, and in which you can use aggregate functions.
mldb.query("""
select Pclass, avg(Age)
from titanic
group by Pclass
having avg(Age) > 30
""")
# # Advanced `FROM` with subqueries
#
# SQL allows you to use the output of one query as the input to another by putting queries in the `FROM` clause, at which point they become "subqueries". The following example shows how to emulate the `HAVING` example above with a subquery, although it should be noted that the `HAVING` form will be faster in this case.
mldb.query("""
select *
from (
select Pclass, avg(Age) as mean_age
from titanic
group by Pclass
)
where mean_age > 30
""")
# # `INTO`: supported via `transform` Procedures
#
# Standard SQL defines an `INTO` clause to create new datasets from the output of queries. MLDB `SELECT` queries are idempotent (they do not modify anything) so `INTO` is not supported directly. You can accomplish the same task with a `transform` procedure, however:
# +
not_supported = """
select Pclass, avg(Age) as mean_age
into class_stats
from titanic
group by Pclass
"""
supported = mldb.post('/v1/procedures', {
"type": "transform",
"params": {
"inputData": """
select Pclass, avg(Age) as mean_age
from titanic
group by Pclass
""",
"outputDataset": "class_stats",
"runOnCreation": True
}
})
print supported
# -
# We can now query our new dataset!
mldb.query("""
select *
from class_stats
""")
# # Advanced `FROM` with `JOIN`
#
# You can run queries across multiple datasets with the `JOIN` keyword, using the `ON` keyword to define how to combine the datasets.
mldb.query("""
select titanic.Name, titanic.Pclass, class_stats.*
from titanic JOIN class_stats ON titanic.Pclass = class_stats.Pclass
order by Age desc limit 10
""")
# ----------
#
# # MLDB extensions to conventional SQL
# MLDB has some notable differences with more conventional SQL databases like PostgreSQL, MySQL, Oracle or SQLServer. For example, MLDB datasets are not SQL tables:
#
# * datasets have no fixed schema
# * datasets can have a variable number of columns, numbering into the millions
# * columns can contain mixed types (i.e. both numbers and strings in the same column)
# * both rows and columns have names
#
# In order to accomodate this, MLDB provides a number of extensions to standard SQL. Examples are provided below.
# Selecting columns based on a prefix:
mldb.query("""
select P*
from titanic
limit 1
""")
# Renaming columns based on a prefix pattern:
mldb.query("""
select P* as x*
from titanic
limit 1
""")
# Excluding columns from a selection:
mldb.query("""
select * excluding(P*)
from titanic
limit 1
""")
# NOTE: Selecting a column which is not in the dataset will not cause an error, instead it will return `NULL`.
mldb.query("""
select nothing
from titanic
limit 1
""")
# MLDB supports JSON-like objects in queries.
mldb.query("""
select {a: 1, b:2, c: 'hello'}
""")
mldb.query("""
select {a: 1, b:2, c: 'hello'} as obj
""")
mldb.query("""
select {a: 1, b:{x:2}, c: 'hello'} as *
""")
# This is especially useful for tokenizing text into bags of words, or importing semi-structured JSON data.
mldb.query("""
select tokenize('Hello world, Hello!', {splitChars: ' ,!'}) as *
""")
mldb.query("""
select parse_json('{"hello":"world","list":[1,2,3,4]}') as *
""")
# MLDB's object notation also allows you to run aggregate functions on multiple columns at once, with the special `{*}` notation, which refers to all fields in the current row as an object.
mldb.query("""
select Pclass, count(*), count({*})
from titanic
group by Pclass
""")
# MLDB's flexible output model also enables powerful aggregate functions like `pivot()` to operate.
mldb.query("""
select Pclass, pivot(Sex, "count(*)") as *
from (
select Pclass, Sex, count(*) from titanic group by Pclass, Sex
)
group by Pclass
""")
# MLDB supports multi-dimensional arrays called embeddings, also known as tensors.
mldb.query("""
select [1,2,3] as embedding
""")
mldb.query("""
select normalize([1,2,3], 1) as n, [1,2,3] / norm([1,2,3] ,1) as d
""")
# MLDB datasets have named rows as well as columns, and the `NAMED` keyword allows you to control the names of your output rows.
mldb.query("""
select *
named Name + ' aged ' + cast(Age as string)
from titanic
order by Age desc limit 10
""")
# Having named rows as well as columns allows us to easily operate on the transpose of a dataset
mldb.query("""
select * from transpose(
(select * from titanic order by Age desc limit 5)
)
""")
# MLDB supports inline Javascript application via the `jseval()` function.
mldb.query("""
select Name,
jseval(
'return Name.replace(/([A-Z])/g, function(m, p) { return " "+p; });',
'Name', Name
) as processed_name
from titanic
order by Age desc limit 10
""")
# MLDB datasets handle millions of columns, and deal very well with sparse datasets, making them ideal for operating on bags of words.
mldb.query("""
select tokenize(
jseval('
return Name.replace(/([A-Z])/g, function(m, p) { return " "+p; });
', 'Name', Name),
{splitChars: ' .()"', quoteChar:''}) as *
from titanic
order by Age desc
limit 10
""")
# Putting it all together, here are the top 20 tokens present in the names of Titanic passengers.
mldb.query("""
select * from transpose((
select sum(
tokenize(
jseval(
'return Name.replace(/([A-Z])/g, function(m, p) { return " "+p; });',
'Name', Name
),
{splitChars: ' .()"', quoteChar:''}
)
) as *
named 'counts'
from titanic
))
order by counts desc limit 20
""")
# ## Where to next?
#
# Check out the other [Tutorials and Demos](../../../../doc/#builtin/Demos.md.html).
| container_files/tutorials/SELECT Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # matplotlib api入门
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Figure和Subplot
# mapplotlib的图像都是位于Figure对象中
fig = plt.figure()
ax1 = fig.add_subplot(2, 2, 1) # 图像是2x2的,现在创建第一个
ax2 = fig.add_subplot(2, 2, 2)
ax3 = fig.add_subplot(2, 2, 3)
ax4 = fig.add_subplot(2, 2, 4)
plt.plot([1.5, 3.5, -2, 1.6])
# +
import numpy as np
plt.plot(np.random.randn(50).cumsum(), 'k--') # 'k'表示是黑色
# -
ax1.hist(np.random.randn(100), bins=20, color='k', alpha=0.3) # ???
ax2.scatter(np.arange(30), np.arange(30) + 3 * np.random.randn(30)) # ???
# 直接创建一个新的Figure,并返回含有已经创建好的subplot对象的numpy数组
fig, axes = plt.subplots(2, 3)
axes
# ## 调整subplot周围的间距
fig, axes = plt.subplots(2, 2, sharex=True, sharey=True)
for i in range(2):
for j in range(2):
axes[i, j].hist(np.random.randn(500), bins=50, color='k', alpha=0.5)
plt.subplots_adjust(wspace=1, hspace=0)
# ## 颜色、标记和线形
plt.plot(np.random.randn(30).cumsum(), 'ko--')
plt.plot(np.random.randn(30).cumsum(), color='k', linestyle='dashed', marker='o')
data = np.random.randn(30).cumsum()
plt.plot(data, 'k--', label='Default')
plt.plot(data, 'k-', drawstyle='steps-post', label='steps-post')
plt.legend(loc='best')
# ## 刻度、标签和图例
plt.xlim() # 不带参数则表示当前x轴绘图范围
plt.xlim([0, 10])
# ## 设置标题、轴标签、刻度以及刻度标签
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
ax.plot(np.random.randn(1000).cumsum())
ticks = ax.set_xticks([0, 250, 500, 750, 1000])
labels = ax.set_xticklabels(['one', 'two', 'three', 'four', 'five'],
rotation=30, fontsize='small')
ax.set_title('My first matplotlib plot')
ax.set_xlabel('Stages')
ax.plot(np.random.randn(1000).cumsum())
# ## 添加图例
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
ax.plot(np.random.randn(100).cumsum(), 'k', label='one')
ax.plot(np.random.randn(100).cumsum(), 'k--', label='two')
ax.plot(np.random.randn(100).cumsum(), 'k.', label='three')
ax.legend(loc='best')
# ## 注解以及在subplot上绘图
# ## 将图标保存到文件
# +
# plt.savefig('some-path')
# 并非一定要写入磁盘,可以写入stringio,方便web上传递图片
# from io import StringIO
# buffer = StringIO()
# plt.savefig(buffer)
# plot_data = buffer.getvalue()
# -
# ## matplotlib配置
# # Pandas中的绘图函数
# ## 线性图
import pandas as pd
| books/Python-for-Data-Analysis/08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ytv3LlPdlb7R" colab_type="text"
# # Visualizing Chipotle's Data
# + [markdown] id="gNDbBe_5lb7W" colab_type="text"
# This time we are going to pull data directly from the internet.
# Special thanks to: https://github.com/justmarkham for sharing the dataset and materials.
#
# ### Step 1. Import the necessary libraries
# + id="A-czVtMDlb7d" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as plt
from collections import Counter
# set this so the graphs open internally
# %matplotlib inline
# + [markdown] id="HnQtxyfUlb7x" colab_type="text"
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv).
# + [markdown] id="P5enpNOclb70" colab_type="text"
# ### Step 3. Assign it to a variable called chipo.
# + id="tRIQ4Yedlb72" colab_type="code" colab={}
url = "https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv"
chipo = pd.read_csv(url, sep = "\t")
# + [markdown] id="CkRJln-vlb8I" colab_type="text"
# ### Step 4. See the first 10 entries
# + id="4Seyie4Dlb8N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="7caf39b8-04bd-4fff-a1ec-2fc24d979c27"
chipo.head()
# + [markdown] id="KcHQswOBlb8x" colab_type="text"
# ### Step 5. Create a histogram of the top 5 items bought
# + id="ZtWZHmG5lb8z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 412} outputId="760d1eb0-4309-460d-c3d9-79d459cc6be2"
top_items = chipo.groupby("item_name")["quantity"].sum().sort_values()[-5:]
#top_items.plot(kind="bar")
top_items.plot.bar()
plt.xlabel("Items")
plt.ylabel("Orders")
plt.title("Most ordered items")
# + [markdown] id="tw9RwU_nlb9L" colab_type="text"
# ### Step 6. Create a scatterplot with the number of items orderered per order price
# #### Hint: Price should be in the X-axis and Items ordered in the Y-axis
# + id="09X8rk8ilb9P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 296} outputId="b329474a-7a7d-493e-b07c-c256e014ff76"
chipo["item_price"] = [float(p[1:]) for p in chipo.item_price]
orders = chipo.groupby("order_id").sum()
orders.plot.scatter(x="item_price", y="quantity", s=15)
plt.xlabel("sum per order")
plt.ylabel("num of orders")
| 07_Visualization/Chipotle/my_Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# http://blog.yhat.com/tutorials/5-Feature-Engineering.html
# +
# Import libraries
import math
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.ensemble import RandomForestClassifier
# -
# Download and read cvs example from AWS.
df = pd.read_csv("https://rodeo-tutorials.s3.amazonaws.com/data/credit-data-trainingset.csv")
df.head(5)
df['serious_dlqin2yrs'].hist()
# Set features.
features = np.array(['revolving_utilization_of_unsecured_lines',
'age', 'number_of_time30-59_days_past_due_not_worse',
'debt_ratio', 'monthly_income','number_of_open_credit_lines_and_loans',
'number_of_times90_days_late', 'number_real_estate_loans_or_lines',
'number_of_time60-89_days_past_due_not_worse', 'number_of_dependents'])
# Build RF classifier.
clf = RandomForestClassifier()
clf.fit(df[features], df['serious_dlqin2yrs'])
# from the calculated importances, order them from most to least important
# and make a barplot so we can visualize what is/isn't important
importances = clf.feature_importances_
sorted_idx = np.argsort(importances)
padding = np.arange(len(features)) + 0.5
plt.barh(padding, importances[sorted_idx], align='center')
plt.yticks(padding, features[sorted_idx])
plt.xlabel("Relative Importance")
plt.title("Variable Importance")
plt.show()
# Bucketing Continuous Values
df['income_bins'] = pd.cut(df.monthly_income, bins=15)
pd.value_counts(df['income_bins'])
# not very usefull
mi_max = df['monthly_income'].max()
mi_max
df['monthly_income'].hist(bins=10,range=[50000,mi_max])
# ### Bin monthly income into buckets
front = np.linspace(0, 10000, 9)
mid = np.delete(np.linspace(10000, 50000, 3), 0, 0)
tail = np.delete(np.linspace(50000, math.ceil(df['monthly_income'].max()), 3),0, 0)
bins = np.append(front, [mid, tail])
bins
df['income_bins'] = pd.cut(df.monthly_income, bins=bins, labels=False)
pd.value_counts(df.income_bins)
# Remove nulls :]
df['income_bins'].isnull().sum()
df['income_bins'] = df['income_bins'].fillna(-1)
# ### Inspecting the predictive value of your bins
# A quick and dirty way to see the effectiveness of derived bins is to inspect their distribution
# as it relates to the variable you're predicting. For us that means cross-tablulating it against serious_dlqin2yrs.
# +
# In this case, it's very easy to see in the plot that serious_dlqin2yrs tends to be higher
# where the income_bins are lower.
df[["income_bins", "serious_dlqin2yrs"]].groupby("income_bins").mean()
# -
cols = ["income_bins", "serious_dlqin2yrs"]
df[cols].groupby("income_bins").mean().plot()
# continuous distributions
cols = ['age', 'serious_dlqin2yrs']
age_means = df[cols].groupby("age").mean()
age_means.plot()
# Bin age into buckets
mybins = [0] + list(range(20, 80, 5)) + [120]
df['age_bucket'] = pd.cut(df.age, bins=mybins)
df['age_bucket'].value_counts()
# Using the age bins, calculate the percent of customers that were delinquent for each bucket
df[["age_bucket", "serious_dlqin2yrs"]].groupby("age_bucket").mean().fillna(0)
df[["age_bucket", "serious_dlqin2yrs"]].groupby("age_bucket").mean().plot()
# ### Factorize - turning categoricals into numericals
# Use factorize to encode categorical data ("A", "B", "C", "A") into numerical data (0, 1, 2, 0).
#
#
labels, levels = pd.factorize(df.age_bucket)
df.age_bucket = labels
df.age_bucket.head()
# ### Write something that buckets debt_ratio into 4 (nearly) equally sized groups.
# Hint: use the quantile method for Series
# +
bins = []
for q in [0.2, 0.4, 0.6, 0.8, 1.0]:
bins.append(df.debt_ratio.quantile(q))
debt_ratio_binned = pd.cut(df.debt_ratio, bins=bins)
debt_ratio_binned
print(pd.value_counts(debt_ratio_binned))
# -
# ### Scaling Features
# Some algorithms will work better if your data is centered around 0. The StandardScaler module in scikit-learn makes it very easy to quickly scale columns in your data frame.
#
# +
from sklearn.preprocessing import StandardScaler
df['monthly_income_scaled'] = StandardScaler().fit_transform(df.monthly_income.values.reshape(-1,1))
print(df.monthly_income_scaled.describe())
print("Mean at 0?", round(df.monthly_income_scaled.mean(), 10)==0)
# -
plt.hist(df.monthly_income_scaled, bins=np.linspace(-0.5,2.5,50))
# ### feature importance (again)
# Let's redo our feature importance calculations to include the engineered features we just created.
# +
features = np.array(['revolving_utilization_of_unsecured_lines',
'age', 'number_of_time30-59_days_past_due_not_worse',
'debt_ratio', 'monthly_income','number_of_open_credit_lines_and_loans',
'number_of_times90_days_late', 'number_real_estate_loans_or_lines',
'number_of_time60-89_days_past_due_not_worse', 'number_of_dependents',
'age_bucket', 'monthly_income_scaled', 'income_bins'])#,
clf = RandomForestClassifier()
clf.fit(df[features], df['serious_dlqin2yrs'])
importances = clf.feature_importances_
sorted_idx = np.argsort(importances)
padding = np.arange(len(features)) + 0.5
plt.barh(padding, importances[sorted_idx], align='center')
plt.yticks(padding, features[sorted_idx])
plt.xlabel("Relative Importance")
plt.title("Variable Importance")
plt.show()
# -
best_features = features[sorted_idx][::-1]
pd.DataFrame({ "name": best_features })
| sklearn/feature-importance-random-forrest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="omJl3NRs12zw" colab_type="text"
# _Lambda School Data Science_
#
# # Ordinary Least Squares Regression - notes
#
# ## What is Linear Regression?
#
# Linear Regression is a statistical model that seeks to describe the relationship between some y variable and one or more x variables.
#
# 
#
# In the simplest case, linear regression seeks to fit a straight line through a cloud of points. This line is referred to as the "regression line" or "line of best fit." This line tries to summarize the relationship between our X and Y in a way that enables us to use the equation for that line to make predictions.
#
# ### Synonyms for "y variable"
# - Dependent Variable
# - Response Variable
# - Outcome Variable
# - Predicted Variable
# - Measured Variable
# - Explained Variable
# - Label
# - Target
#
# ### Synonyms for "x variable"
# - Independent Variable
# - Explanatory Variable
# - Regressor
# - Covariate
# - Feature
# + [markdown] id="GLfiyLr-12z0" colab_type="text"
# # Simple Linear Regresion (bivariate)
#
# ## Making Predictions
#
# Say that we were trying to create a model that captured the relationship between temperature outside and ice cream sales. In Machine Learning our goal is often different that of other flavors of Linear Regression Analysis, because we're trying to fit a model to this data with the intention of making **predictions** on new data (in the future) that we don't have yet.
#
# ## What are we trying to predict?
#
# So if we had measured ice cream sales and the temprature outside on 11 different days, at the end of our modeling **what would be the thing that we would want to predict? - Ice Cream Sales or Temperature?**
#
# We would probably want to be measuring temperature with the intention of using that to **forecast** ice cream sales. If we were able to successfully forecast ice cream sales from temperature, this might help us know beforehand how much ice cream to make or how many cones to buy or on which days to open our store, etc. Being able to make predictions accurately has a lot of business implications. This is why making accurate predictions is so valuable (And in large part is why data scientists are paid so well).
#
# ### Y Variable Intuition
#
# We want the thing that we're trying to predict to serve as our **y** variable. This is why it's sometimes called the "predicted variable." We call it the "dependent" variable because our prediction for how much ice cream we're going to sell "depends" on the temperature outside.
#
# ### X Variable Intuition
#
# All other variables that we use to predict our y variable (we're going to start off just using one) we call our **x** variables. These are called our "independent" variables because they don't *depend* on y, they "explain" y. Hence they are also referred to as our "explanatory" variables.
# + id="p0X_but-12z5" colab_type="code" colab={}
# %matplotlib inline
from ipywidgets import interact
from matplotlib.patches import Rectangle
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
import statsmodels.api as sm
# + id="i17Ht2zf120L" colab_type="code" colab={}
columns = ['Year','Incumbent Party Candidate','Other Candidate','Incumbent Party Vote Share']
data = [[1952,"Stevenson","Eisenhower",44.6],
[1956,"Eisenhower","Stevenson",57.76],
[1960,"Nixon","Kennedy",49.91],
[1964,"Johnson","Goldwater",61.34],
[1968,"Humphrey","Nixon",49.60],
[1972,"Nixon","McGovern",61.79],
[1976,"Ford","Carter",48.95],
[1980,"Carter","Reagan",44.70],
[1984,"Reagan","Mondale",59.17],
[1988,"<NAME>r.","Dukakis",53.94],
[1992,"<NAME>r.","Clinton",46.55],
[1996,"Clinton","Dole",54.74],
[2000,"Gore","Bush, Jr.",50.27],
[2004,"Bush, Jr.","Kerry",51.24],
[2008,"McCain","Obama",46.32],
[2012,"Obama","Romney",52.00]]
df = pd.DataFrame(data=data, columns=columns)
# + id="TmAj9NrL120X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="6851ba3d-9ecc-46f2-daf6-8884ade0a64a"
df
# + id="pT0AAnic120j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="330a08e4-145a-4cf9-b245-3ee08d8f52a1"
df.plot(x='Year', y='Incumbent Party Vote Share');
# + id="Y8MKji6k120z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="3cae751c-3d9d-4e0e-efec-a1c799ef4745"
df['Incumbent Party Vote Share'].describe()
# + id="IuMStnkH1203" colab_type="code" colab={}
target = 'Incumbent Party Vote Share'
df['Prediction'] = df[target].mean()
df['Error'] = df['Prediction'] - df[target]
# + id="dBJlc_v3120_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="a7e2c6c8-e451-4ca2-bdda-4c5330ffce27"
df
# + id="AhqQBZsV121F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="85704f76-bb97-43b4-eeb7-e69115b9680c"
df['Error'].sum()
# + id="GgbrVyC2121N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0387cacd-8081-4ddb-b8da-f6a5215af385"
df['Absolute Error'] = df['Error'].abs()
df['Absolute Error'].sum()
# + id="wPmTNGtk121S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a7516d03-61d2-4454-cd9b-b07e2765f3e8"
df['Absolute Error'].mean()
# + id="00PO9ett121W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4ccc2774-1327-4720-9eb0-898eb9e8b42a"
mean_absolute_error(y_true=df[target], y_pred=df['Prediction'])
# + id="YaSlOKCZ121b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dde2d83f-9cfb-4595-943e-a9dd15a66b91"
r2_score(y_true=df[target], y_pred=df['Prediction'])
# + id="hcQLXMEF121h" colab_type="code" colab={}
columns = ['Year','Average Recent Growth in Personal Incomes']
data = [[1952,2.40],
[1956,2.89],
[1960, .85],
[1964,4.21],
[1968,3.02],
[1972,3.62],
[1976,1.08],
[1980,-.39],
[1984,3.86],
[1988,2.27],
[1992, .38],
[1996,1.04],
[2000,2.36],
[2004,1.72],
[2008, .10],
[2012, .95]]
growth = pd.DataFrame(data=data, columns=columns)
# + id="MWe0Nj9A121l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 562} outputId="8bb4f5d9-ca2b-43b0-dbc9-e0a044bbda17"
df = df.merge(growth)
df
# + id="h-Cyego5121p" colab_type="code" colab={}
feature = 'Average Recent Growth in Personal Incomes'
# + id="L4jA1Grm121u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="560a6e82-fe0b-4957-ba11-e1191981482d"
df.plot(x=feature, y=target, kind='scatter');
# + [markdown] id="u7OoSyAI121y" colab_type="text"
# We can see from the scatterplot that these data points seem to follow a somewhat linear relationship. This means that we could probably summarize their relationship well by fitting a line of best fit to these points. Lets do it.
#
#
# ## The Equation for a Line
#
# As we know a common equation for a line is:
#
# \begin{align}
# y = mx + b
# \end{align}
#
# Where $m$ is the slope of our line and $b$ is the y-intercept.
#
# If we want to plot a line through our cloud of points we figure out what these two values should be. Linear Regression seeks to **estimate** the slope and intercept values that describe a line that best fits the data points.
# + id="OuoILQfr1210" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e17acaa4-fff5-45a8-f52d-b78cc402693d"
m = 4
b = 44
df['Prediction'] = m * df[feature] + b
df['Error'] = df['Prediction'] - df[target]
df['Absolute Error'] = df['Error'].abs()
df['Absolute Error'].sum()
# + id="0_k2PYLD1215" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="924b038f-914a-4691-fdcb-e0a1e741856c"
df['Absolute Error'].mean()
# + id="J73RTrlJ1218" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0345544b-2090-4041-8f9c-71416f9d8a8b"
r2_score(y_true=df[target], y_pred=df['Prediction'])
# + id="-LVNdAdL122A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="3c5bf43f-5f8d-47a5-cc29-28dc8ce951b1"
ax = df.plot(x=feature, y=target, kind='scatter')
df.plot(x=feature, y='Prediction', kind='line', ax=ax);
# + id="RkyVu4qE122H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 458} outputId="1a5f0003-5571-4a8f-8287-3b12d27efe1c"
def regression(m, b):
df['Prediction'] = m * df[feature] + b
df['Error'] = df['Prediction'] - df[target]
df['Absolute Error'] = df['Error'].abs()
sum_absolute_error = df['Absolute Error'].sum()
title = f'Sum of absolute errors: {sum_absolute_error}'
ax = df.plot(x=feature, y=target, kind='scatter', title=title, figsize=(7, 7))
df.plot(x=feature, y='Prediction', kind='line', ax=ax)
regression(m=3, b=46)
# + [markdown] id="fl7lVFED122L" colab_type="text"
# ## Residual Error
#
# The residual error is the distance between points in our dataset and our regression line.
# + id="x3F3aMHc122M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 458} outputId="dc601b31-98c6-481e-9c3f-4cba4319415d"
def regression(m, b):
df['Prediction'] = m * df[feature] + b
df['Error'] = df['Prediction'] - df[target]
df['Absolute Error'] = df['Error'].abs()
sum_absolute_error = df['Absolute Error'].sum()
title = f'Sum of absolute errors: {sum_absolute_error}'
ax = df.plot(x=feature, y=target, kind='scatter', title=title, figsize=(7, 7))
df.plot(x=feature, y='Prediction', kind='line', ax=ax)
for x, y1, y2 in zip(df[feature], df[target], df['Prediction']):
ax.plot((x, x), (y1, y2), color='grey')
regression(m=3.5, b=46)
# + id="XoYM68Zu122P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="5a421d96-2c14-4120-c6df-e3fbfa3b0515"
interact(regression, m=(-10,10,0.5), b=(40,60,0.5));
# + id="4QPg_F2U122R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 562} outputId="2a6c51e0-73ac-4417-908d-acc99fc4101e"
df
# + id="yLcIg0Df122T" colab_type="code" colab={}
df['Square Error'] = df['Error'] **2
# + id="vyc477ez122X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 562} outputId="5649e29e-e014-46b0-d1f1-a27949b6fa65"
df
# + id="h5sDSbXI122a" colab_type="code" colab={}
def regression(m, b):
df['Prediction'] = m * df[feature] + b
df['Error'] = df['Prediction'] - df[target]
df['Absolute Error'] = df['Error'].abs()
df['Square Error'] = df['Error'] **2
sum_square_error = df['Square Error'].sum()
title = f'Sum of square errors: {sum_square_error}'
ax = df.plot(x=feature, y=target, kind='scatter', title=title, figsize=(7, 7))
df.plot(x=feature, y='Prediction', kind='line', ax=ax)
xmin, xmax = ax.get_xlim()
ymin, ymax = ax.get_ylim()
scale = (xmax-xmin)/(ymax-ymin)
for x, y1, y2 in zip(df[feature], df[target], df['Prediction']):
bottom_left = (x, min(y1, y2))
height = abs(y1 - y2)
width = height * scale
ax.add_patch(Rectangle(xy=bottom_left, width=width, height=height, alpha=0.1))
# + id="KiCmd3hP122d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="9322cc3b-d3a8-403f-edd4-72429e0d7c1a"
interact(regression, m=(-10,10,0.5), b=(40,60,0.5));
# + id="-WZCy-mkAV7V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 458} outputId="e608b88b-f721-4274-c357-4137d4bf491b"
regression(3, 46.5)
# + id="kyR-P28w122g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1289} outputId="62f84c38-ebbc-4e8c-c9c2-148c9550df4c"
b = 46
ms = np.arange(-10,10,0.5)
sses = []
for m in ms:
predictions = m * df[feature] + b
errors = predictions - df[target]
square_errors = errors ** 2
sse = square_errors.sum()
sses.append(sse)
hypotheses = pd.DataFrame({'Slope': ms})
hypotheses['Intercept'] = b
hypotheses['Sum of Square Errors'] = sses
hypotheses
# + id="2AnrMiRP122j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="392e6926-d65a-439e-873b-01c8fc06312d"
hypotheses.plot(x='Slope', y='Sum of Square Errors',
title=f'Intercept={b}');
# + id="0MXhGpft122o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="168fc54f-a239-47cf-c5a4-b5258ca8bde4"
X = df[[feature]]
y = df[target]
X.shape, y.shape
# + id="nVPrfuRt122r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="b577bf80-94f1-4f5d-bca2-4be41a0e2970"
model = LinearRegression()
model.fit(X, y)
# + id="8oo33tpb122v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="79ba6bd3-f09b-42f1-eb4d-baf43c348053"
model.coef_, model.intercept_
# + id="k-tOWY_L122y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="823cdbff-cc93-4ec2-e7bc-5c7db9b69222"
model.predict([[1]])
# + id="xTmPs7BO1222" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6a0cab91-1153-4fcf-e25c-8d35fa0cd353"
model.predict([[2]])
# + id="tsAlmUAK1224" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6841f462-ef3f-4d8f-c453-edca1cd5f68d"
model.predict([[3]])
# + id="GYGftGc81226" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="4d4222bc-d70a-4b8c-9a24-3fc1add57b9a"
model.predict(X)
# + id="nlGPNhKU1227" colab_type="code" colab={}
df['Prediction'] = model.predict(X)
# + id="LkMzv9SN122-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="19aaabc2-22c8-4755-c3e5-187c654620ec"
ax = df.plot(x=feature, y=target, kind='scatter', title='sklearn LinearRegression')
df.plot(x=feature, y='Prediction', kind='line', ax=ax);
# + id="ApuX9mlU123C" colab_type="code" colab={}
df['Error'] = df['Prediction'] - y
# + id="Xa-zVZl_123E" colab_type="code" colab={}
df['Absolute Error'] = df['Error'].abs()
df['Square Error'] = df['Error'] ** 2
# + id="9X1kA7Ri123G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4b37b51b-92dd-45c5-e222-c5c0cad961c4"
df['Square Error'].mean()
# + id="QGIFHfZj123I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1fb992bf-d6b6-4835-a655-edea5eae2756"
mean_squared_error(y_true=y, y_pred=model.predict(X))
# + id="5nzJ8wf-123L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a87c6043-43e2-4d08-97c4-5294136206ed"
np.sqrt(mean_squared_error(y, model.predict(X)))
# + [markdown] id="W9MRIt4c123O" colab_type="text"
# # R Squared: $R^2$
#
# One final attribute of linear regressions that we're going to talk about today is a measure of goodness of fit known as $R^2$ or R-squared. $R^2$ is a statistical measure of how close the data are fitted to our regression line. A helpful interpretation for the $R^2$ is the percentage of the dependent variable that is explained by the model.
#
# In other words, the $R^2$ is the percentage of y that is explained by the x variables included in the model. For this reason the $R^2$ is also known as the "coefficient of determination," because it explains how much of y is explained (or determined) by our x varaibles. We won't go into the calculation of $R^2$ today, just know that a higher $R^2$ percentage is nearly always better and indicates a model that fits the data more closely.
# + id="dSDVxwF3123P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f17fd758-bc00-4f2f-a208-4b22b6b1b785"
model.score(X, y)
# + id="osXOE4Ew123R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4b145b79-485e-4296-e359-5ae87244db73"
r2_score(y, model.predict(X))
# + [markdown] id="Rsrdnk1d123V" colab_type="text"
# ### Statsmodels
# + [markdown] id="0zt4dl8O123V" colab_type="text"
# https://www.statsmodels.org/dev/examples/notebooks/generated/ols.html
# + id="_Jov649z123W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 550} outputId="1fb1986b-778b-41cf-a571-2ea14b2d7e80"
model = sm.OLS(y, sm.add_constant(X))
results = model.fit()
print(results.summary())
# + [markdown] id="8_vwwnCg123X" colab_type="text"
# # The Anatomy of Linear Regression
#
# - Intercept: The $b$ value in our line equation $y=mx+b$
# - Slope: The $m$ value in our line equation $y=mx+b$. These two values together define our regression line.
#
# 
#
# - $\hat{y}$ : A prediction
# - Line of Best Fit (Regression Line)
# - Predicted (fitted) Values: Points on our regression line
# - Observed Values: Points from our dataset
# - Error: The distance between predicted and observed values.
#
# 
#
# + [markdown] id="5aYmOfe_123Z" colab_type="text"
# # More Formal Notation
#
# 
#
# We have talked about a line of regression being represented like a regular line $y=mx+b$ but as we get to more complicated versions we're going to need to extend this equation. So lets establish the proper terminology.
#
# **X** - Independent Variable, predictor variable, explanatory variable, regressor, covariate
#
# **Y** - Response variable, predicted variable, measured vairable, explained variable, outcome variable
#
# $\beta_0$ - "Beta Naught" or "Beta Zero", the intercept value. This is how much of y would exist if X were zero. This is sometimes represented by the letter "a" but I hate that. So it's "Beta 0" during my lecture.
#
# $\beta_1$ - "Beta One" The primary coefficient of interest. This values is the slope of the line that is estimated by "minimizing the sum of the squared errors/residuals" - We'll get to that.
#
# $\epsilon$ - "Epsilon" The "error term", random noise, things outside of our model that affect y.
# + [markdown] id="tNz-iXN0123a" colab_type="text"
# # How Does it do it?
#
# ## Minimizing the Sum of the Squared Error
#
# The most common method of estimating our $\beta$ parameters is what's known as "Ordinary Least Squares" (OLS). (There are different methods of arriving at a line of best fit). OLS estimates the parameters that minimize the squared distance between each point in our dataset and our line of best fit.
#
# \begin{align}
# SSE = \sum(y_i - \hat{y})^2
# \end{align}
#
#
# + [markdown] id="lGzEoymn123a" colab_type="text"
# ## Linear Algebra!
#
# The same result that is found by minimizing the sum of the squared errors can be also found through a linear algebra process known as the "Least Squares Solution:"
#
# 
#
# Before we can work with this equation in its linear algebra form we have to understand how to set up the matrices that are involved in this equation.
# + [markdown] id="ayDaqc0D123b" colab_type="text"
# ### The $\beta$ vector
#
# The $\beta$ vector represents all the parameters that we are trying to estimate, our $y$ vector and $X$ matrix values are full of data from our dataset. The $\beta$ vector holds the variables that we are solving for: $\beta_0$ and $\beta_1$
#
# Now that we have all of the necessary parts we can set them up in the following equation:
#
# \begin{align}
# y = X \beta + \epsilon
# \end{align}
#
# Since our $\epsilon$ value represents **random** error we can assume that it will equal zero on average.
#
# \begin{align}
# y = X \beta
# \end{align}
#
# The objective now is to isolate the $\beta$ matrix. We can do this by pre-multiplying both sides by "X transpose" $X^{T}$.
#
# \begin{align}
# X^{T}y = X^{T}X \beta
# \end{align}
#
# Since anything times its transpose will result in a square matrix, if that matrix is then an invertible matrix, then we should be able to multiply both sides by its inverse to remove it from the right hand side. (We'll talk tomorrow about situations that could lead to $X^{T}X$ not being invertible.)
#
# \begin{align}
# (X^{T}X)^{-1}X^{T}y = (X^{T}X)^{-1}X^{T}X \beta
# \end{align}
#
# Since any matrix multiplied by its inverse results in the identity matrix, and anything multiplied by the identity matrix is itself, we are left with only $\beta$ on the right hand side:
#
# \begin{align}
# (X^{T}X)^{-1}X^{T}y = \hat{\beta}
# \end{align}
#
# We will now call it "beta hat" $\hat{\beta}$ because it now represents our estimated values for $\beta_0$ and $\beta_1$
#
# ### Lets calculate our $\beta$ coefficients with numpy!
# + id="6lS8o1Tx123b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 958} outputId="661db205-ded2-4287-cf78-654a5d3fc58b"
X = sm.add_constant(df[feature]).values
print('X')
print(X)
y = df[target].values[:, np.newaxis]
print('y')
print(y)
X_transpose = X.T
print('X Transpose')
print(X_transpose)
X_transpose_X = X_transpose @ X
print('X Transpose X')
print(X_transpose_X)
X_transpose_X_inverse = np.linalg.inv(X_transpose_X)
print('X Transpose X Inverse')
print(X_transpose_X_inverse)
X_transpose_y = X_transpose @ y
print('X Transpose y')
print(X_transpose_y)
beta_hat = X_transpose_X_inverse @ X_transpose_y
print('Beta Hat')
print(beta_hat)
# + [markdown] id="eXlj3Juu123d" colab_type="text"
# # Multiple Regression
#
# Simple or bivariate linear regression involves a single $x$ variable and a single $y$ variable. However, we can have many $x$ variables. A linear regression model that involves multiple x variables is known as **Multiple** Regression - NOT MULTIVARIATE!
#
# 
# + id="yLBSiVgM123e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 579} outputId="7e8d4079-296a-4392-b4ba-7f391e2a0790"
df.sort_values(by='Square Error', ascending=False)
# + id="YG6lPV8b123f" colab_type="code" colab={}
"""
Fatalities denotes the cumulative number of American military
fatalities per millions of US population the in Korea, Vietnam,
Iraq and Afghanistan wars during the presidential terms
preceding the 1952, 1964, 1968, 1976 and 2004, 2008 and
2012 elections.
http://www.douglas-hibbs.com/HibbsArticles/HIBBS-PRESVOTE-SLIDES-MELBOURNE-Part1-2014-02-26.pdf
"""
columns = ['Year','US Military Fatalities per Million']
data = [[1952,190],
[1956, 0],
[1960, 0],
[1964, 1],
[1968,146],
[1972, 0],
[1976, 2],
[1980, 0],
[1984, 0],
[1988, 0],
[1992, 0],
[1996, 0],
[2000, 0],
[2004, 4],
[2008, 14],
[2012, 5]]
deaths = pd.DataFrame(data=data, columns=columns)
# + id="5N9t0IHG123h" colab_type="code" colab={}
df = df.merge(deaths)
# + id="h1DdAls_123l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 567} outputId="f2d07109-05d2-4ab0-ff14-f9afb14042f2"
features = ['Average Recent Growth in Personal Incomes',
'US Military Fatalities per Million']
target = 'Incumbent Party Vote Share'
X = df[features]
y = df[target]
model = sm.OLS(y, sm.add_constant(X))
results = model.fit()
print(results.summary())
# + id="U5JPDLhp123o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="fac677a9-a5f6-458b-a6fc-8ee0cb08fdc4"
model = LinearRegression()
model.fit(X, y)
print('Intercept:', model.intercept_)
pd.Series(model.coef_, features)
# + id="wf8q2cEY123p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="91a57527-256a-42b9-bab8-18b9068c508f"
np.sqrt(mean_squared_error(y, model.predict(X)))
# + [markdown] id="TovMOsNi123r" colab_type="text"
# # Train / Test Split
# + id="ku5UBe3Z123s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="93724778-3151-4e5e-8d1a-974a3e5ba3ce"
train = df.query('Year < 2008')
test = df.query('Year >= 2008')
X_train = train[features]
y_train = train[target]
X_test = test[features]
y_test = test[target]
X_train.shape, y_train.shape, X_test.shape, y_test.shape
# + id="mXWF1eVn123w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1cb61154-2692-4524-be1f-98dcdf8e8689"
model.fit(X_train, y_train)
model.predict(X_test)
# + id="44k2j2sv123z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 145} outputId="74b6faa0-e140-4626-d61e-e5a279c47988"
test
# + [markdown] id="b-8E5BX81232" colab_type="text"
# ### More about the "Bread & Peace" model
# - https://fivethirtyeight.com/features/what-do-economic-models-really-tell-us-about-elections/
# - https://statmodeling.stat.columbia.edu/2007/12/15/bread_and_peace/
# - https://avehtari.github.io/RAOS-Examples/ElectionsEconomy/hibbs.html
# - https://douglas-hibbs.com/
# - http://www.douglas-hibbs.com/HibbsArticles/HIBBS-PRESVOTE-SLIDES-MELBOURNE-Part1-2014-02-26.pdf
# + [markdown] id="aDWksVbf1233" colab_type="text"
# # Dimensionality in Linear Regression!
#
# Muliple Regression is simply an extension of the bivariate case. The reason why we see the bivariate case demonstrated so often is simply because it's easier to graph and all of the intuition from the bivariate case is the same as we keep on adding explanatory variables.
#
# As we increase the number of $x$ values in our model we are simply fitting a n-1-dimensional plane to an n-dimensional cloud of points within an n-dimensional hypercube.
# + [markdown] id="eSErkf1J1234" colab_type="text"
# # Interpreting Coefficients
#
# One of Linear Regression's strengths is that the parameters of the model (coefficients) are readily interpretable and useful. Not only do they describe the relationship between x and y but they put a number on just how much x is associated with y. We should be careful to not speak about this relationshiop in terms of causality because these coefficients are in fact correlative measures. We would need a host of additional techniques in order to estimate a causal effect using linear regression (econometrics).
#
# \begin{align}
# \hat{\beta} = \frac{Cov(x,y)}{Var(y)}
# \end{align}
#
# Going back to the two equations for the two models that we have estimated so far, lets replace their beta values with their actual values to see if we can make sense of how to interpret these beta coefficients.
# + [markdown] id="Mrcl_weP1235" colab_type="text"
# ## Bivariate Model
#
# $y_i = \beta_0 + \beta_1temperature + \epsilon$
#
# $sales_i = -596.2 + 24.69temperature + \epsilon$
#
# What might $\beta_0$ in this model represent? It represents the level of sales that we would have if temperature were 0. Since this is negative one way of interpreting it is that it's so cold outside that you would have to pay people to eat ice cream. A more appropriate interpretation is probably that the ice cream store owner should close his store down long before the temperature reaches 0 degrees farenheit (-17.7 celsius). The owner can compare his predicted sales with his costs of doing business to know how warm the weather has to get before he should open his store.
#
# What might the $beta_1$ in this model reprsent? it represents the increase in sales for each degree of temperature increase. For every degree that the temperature goes up outside he has $25 more in sales.
# + [markdown] id="1BCj64eR1236" colab_type="text"
# ## Multiple Regression Model
#
# $y_i = \beta_0 + \beta_1age_i + \beta_2weight_i + \epsilon$
#
# $BloodPressure_i = 30.99+ .86age_i + .33weight_i + \epsilon$
#
# The interpretation of coefficients in this example are similar. The intercept value repesents the blood pressure a person would have if they were 0 years old and weighed 0 pounds. This not a super useful interpretation. If we look at our data it is unlikely that we have any measurements like these in the dataset. This means that our interpretation of our intercept likely comes from extrapolating the regression line (plane). Coefficients having straightforward interpretations is a strength of linear regression if we're careful about extrapolation and only interpreting our data within the context that it was gathered.
#
# The interpretation of our other coefficients can be a useful indicator for how much a person similar to those in our dataset's blood pressure will go up on average with each additional year of age and pound of weight.
# + [markdown] id="3X7NGUij1237" colab_type="text"
# # Basic Model Validation
#
# One of the downsides of relying on $R^2$ too much is that although it tells you when you're fitting the data well, it doesn't tell you when you're *overfitting* the data. The best way to tell if you're overfitting the data is to get some data that your model hasn't seen yet, and evaluate how your predictions do. This is essentially what "model validation" is.
# + [markdown] id="NVrAKh3S1238" colab_type="text"
# # Why is Linear Regression so Important?
#
# ## Popularity
#
# Linear Regression is an extremely popular technique that every data scientist **needs** to understand. It's not the most advanced technique and there are supervised learning techniques that will obtain a higher accuracy, but where it lacks in accuracy it makes up for it in interpretability and simplicity.
#
# ## Interpretability
#
# Few other models possess coefficients that are so directly linked to their variables with a such a clear interpretation. Tomorrow we're going to learn about ways to make them even easier to interpret.
#
# ## Simplicity
#
# A linear regression model can be communicated just by writing out its equation. It's kind of incredible that such high dimensional relationships can be described from just a linear combination of variables and coefficients.
| module1-ols-regression/ols-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DrivenData: Pump it Up Starter Code
#
# This notebook provides some starter code for the water table project from DrivenData. First, we'll import the libraries will be using. Feel free to import others if you need.
import pandas as pd
import seaborn as sns
import requests
# Then, we'll get the list of url's to our dataset.
# +
train_features_url = "https://drivendata-prod.s3.amazonaws.com/data/7/public/4910797b-ee55-40a7-8668-10efd5c1b960.csv?X-Amz-Algorithm=AWS4-HMAC-SHA256&X-Amz-Credential=<KEY>LMPSY%2F20201002%2Fus-east-1%2Fs3%2Faws4_request&X-Amz-Date=20201002T044617Z&X-Amz-Expires=86400&X-Amz-SignedHeaders=host&X-Amz-Signature=d30aed3efa8ffd5d8e00373b85f3313f41e3a55a09730d8ebfa442258f371a6b"
train_labels_url = "https://drivendata-prod.s3.amazonaws.com/data/7/public/0bf8bc6e-30d0-4c50-956a-603fc693d966.csv?X-Amz-Algorithm=AWS4-HMAC-SHA256&X-Amz-Credential=AKIARVBOBDCYVI2LMPSY%2F20201002%2Fus-east-1%2Fs3%2Faws4_request&X-Amz-Date=20201002T044617Z&X-Amz-Expires=86400&X-Amz-SignedHeaders=host&X-Amz-Signature=bc5f308eba4daa23639d22ddb053462d04b76bffe90ddec0b4af4d166bdad9b3"
test_features_url = "https://drivendata-prod.s3.amazonaws.com/data/7/public/702ddfc5-68cd-4d1d-a0de-f5f566f76d91.csv?X-Amz-Algorithm=AWS4-HMAC-SHA256&X-Amz-Credential=<PASSWORD>SY%2F20201002%2Fus-east-1%2Fs3%2Faws4_request&X-Amz-Date=20201002T044617Z&X-Amz-Expires=86400&X-Amz-SignedHeaders=host&X-Amz-Signature=9f1572b0d8cf8568acc5cf4f785b5ebe00fd59b1e28148e84323163b6c7055a6"
sample_sub_url = "https://drivendata-prod.s3.amazonaws.com/data/7/public/SubmissionFormat.csv?X-Amz-Algorithm=AWS4-HMAC-SHA256&X-Amz-Credential=AKI<PASSWORD>%2F20201002%2Fus-east-1%2Fs3%2Faws4_request&X-Amz-Date=20201002T044617Z&X-Amz-Expires=86400&X-Amz-SignedHeaders=host&X-Amz-Signature=0d8750e142897c8447d7810cefde94d064b43d3430f40838404051df934e7807"
data_dict = {'train_features.csv': train_features_url,
'train_labels.csv': train_labels_url,
'test_features.csv': test_features_url,
'sample_submission.csv': sample_sub_url}
# -
# And use the `requests` library to download the data.
for dataset in data_dict:
r = requests.get(data_dict[dataset])
with open(dataset, 'wb') as f:
f.write(r.content)
#
| starter_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import onnxruntime
import torch
import numpy as np
import torch.nn.functional as F
import matplotlib.pyplot as plt
input_path = 'checkpoint'
tags_path = os.path.join(input_path, 'tags.txt')
model_path = os.path.join(input_path, 'model.onnx')
generator_path = os.path.join(input_path, 'Gs.pth')
device = device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
batch_size = 4
seed = 0
# let's run one image to checkout if it works
C = onnxruntime.InferenceSession(model_path)
with open(tags_path, 'r') as tags_stream:
tags = np.array([tag for tag in (tag.strip() for tag in tags_stream) if tag])
# +
import stylegan2
from stylegan2 import utils
G = stylegan2.models.load(generator_path, map_location=device)
G.to(device)
# -
def to_image_tensor(image_tensor, pixel_min=-1, pixel_max=1):
if pixel_min != 0 or pixel_max != 1:
image_tensor = (image_tensor - pixel_min) / (pixel_max - pixel_min)
return image_tensor.clamp(min=0, max=1)
# +
torch.manual_seed(seed)
qlatents = torch.randn(1, G.latent_size).to(device=device, dtype=torch.float32)
generated = G(qlatents)
images = to_image_tensor(generated)
# 299 is the input size of the model
images = F.interpolate(images, size=(299, 299), mode='bilinear')
ort_inputs = {C.get_inputs()[0].name: images.detach().cpu().numpy()}
[predicted_labels] = C.run(None, ort_inputs)
# print out some tags
plt.imshow(images[0].detach().cpu().permute(1, 2, 0))
labels = [tags[i] for i, score in enumerate(predicted_labels[0]) if score > 0.5]
print(labels)
# +
# reset seed
torch.manual_seed(seed)
iteration = 5000
progress = utils.ProgressWriter(iteration)
progress.write('Generating images...', step=False)
qlatents_data = torch.Tensor(0, G.latent_size).to(device=device, dtype=torch.float32)
dlatents_data = torch.Tensor(0, 16, G.latent_size).to(device=device, dtype=torch.float32)
labels_data = torch.Tensor(0, len(tags)).to(device=device, dtype=torch.float32)
for i in range(iteration):
qlatents = torch.randn(batch_size, G.latent_size).to(device=device, dtype=torch.float32)
with torch.no_grad():
generated, dlatents = G(latents=qlatents, return_dlatents=True)
# inplace to save memory
generated = to_image_tensor(generated)
# 299 is the input size of the model
# resize the image to 299 * 299
images = F.interpolate(generated, size=(299, 299), mode='bilinear')
labels = []
## tagger does not take input as batch, need to feed one by one
for image in images:
ort_inputs = {C.get_inputs()[0].name: image.reshape(1, 3, 299, 299).detach().cpu().numpy()}
[[predicted_labels]] = C.run(None, ort_inputs)
labels.append(predicted_labels)
# store the result
labels_tensor = torch.Tensor(labels).to(device=device, dtype=torch.float32)
qlatents_data = torch.cat((qlatents_data, qlatents))
dlatents_data = torch.cat((dlatents_data, dlatents))
labels_data = torch.cat((labels_data, labels_tensor))
progress.step()
progress.write('Done!', step=False)
progress.close()
# -
torch.save({
'qlatents_data': qlatents_data.cpu(),
'dlatents_data': dlatents_data.cpu(),
'labels_data': labels_data.cpu(),
'tags': tags
}, 'latents.pth')
| Generate labeled anime data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.056945, "end_time": "2020-11-16T15:32:12.292309", "exception": false, "start_time": "2020-11-16T15:32:12.235364", "status": "completed"} tags=[]
# - a notebook to save preprocessing model and train/save NN models
# - all necessary ouputs are stored in MODEL_DIR = output/kaggle/working/model
# - put those into dataset, and load it from inference notebook
# + papermill={"duration": 6.296766, "end_time": "2020-11-16T15:32:18.644686", "exception": false, "start_time": "2020-11-16T15:32:12.347920", "status": "completed"} tags=[]
import sys
sys.path.append('../input/iterative-stratification/iterative-stratification-master')
sys.path.append('../input/umaplearn/umap')
# %mkdir model
# %mkdir interim
from scipy.sparse.csgraph import connected_components
from umap import UMAP
from iterstrat.ml_stratifiers import MultilabelStratifiedKFold
import numpy as np
import random
import pandas as pd
import matplotlib.pyplot as plt
import os
import copy
import seaborn as sns
import time
from sklearn import preprocessing
from sklearn.metrics import log_loss
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA,FactorAnalysis
from sklearn.manifold import TSNE
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
print(torch.cuda.is_available())
import warnings
# warnings.filterwarnings('ignore')
# + papermill={"duration": 0.069706, "end_time": "2020-11-16T15:32:18.771815", "exception": false, "start_time": "2020-11-16T15:32:18.702109", "status": "completed"} tags=[]
torch.__version__
# + papermill={"duration": 0.069099, "end_time": "2020-11-16T15:32:18.897467", "exception": false, "start_time": "2020-11-16T15:32:18.828368", "status": "completed"} tags=[]
NB = '25'
IS_TRAIN = True
MODEL_DIR = "model" # "../model"
INT_DIR = "interim" # "../interim"
NSEEDS = 5 # 5
DEVICE = ('cuda' if torch.cuda.is_available() else 'cpu')
EPOCHS = 15
BATCH_SIZE = 256
LEARNING_RATE = 5e-3
WEIGHT_DECAY = 1e-5
EARLY_STOPPING_STEPS = 10
EARLY_STOP = False
NFOLDS = 5 # 5
PMIN = 0.0005
PMAX = 0.9995
SMIN = 0.0
SMAX = 1.0
# + papermill={"duration": 6.072238, "end_time": "2020-11-16T15:32:25.030415", "exception": false, "start_time": "2020-11-16T15:32:18.958177", "status": "completed"} tags=[]
train_features = pd.read_csv('../input/lish-moa/train_features.csv')
train_targets_scored = pd.read_csv('../input/lish-moa/train_targets_scored.csv')
train_targets_nonscored = pd.read_csv('../input/lish-moa/train_targets_nonscored.csv')
test_features = pd.read_csv('../input/lish-moa/test_features.csv')
sample_submission = pd.read_csv('../input/lish-moa/sample_submission.csv')
# + papermill={"duration": 0.262629, "end_time": "2020-11-16T15:32:25.351370", "exception": false, "start_time": "2020-11-16T15:32:25.088741", "status": "completed"} tags=[]
train_targets_nonscored = train_targets_nonscored.loc[:, train_targets_nonscored.sum() != 0]
print(train_targets_nonscored.shape)
# + papermill={"duration": 2.920741, "end_time": "2020-11-16T15:32:28.329561", "exception": false, "start_time": "2020-11-16T15:32:25.408820", "status": "completed"} tags=[]
# for c in train_targets_scored.columns:
# if c != "sig_id":
# train_targets_scored[c] = np.maximum(PMIN, np.minimum(PMAX, train_targets_scored[c]))
for c in train_targets_nonscored.columns:
if c != "sig_id":
train_targets_nonscored[c] = np.maximum(PMIN, np.minimum(PMAX, train_targets_nonscored[c]))
# + papermill={"duration": 0.095352, "end_time": "2020-11-16T15:32:28.528120", "exception": false, "start_time": "2020-11-16T15:32:28.432768", "status": "completed"} tags=[]
print("(nsamples, nfeatures)")
print(train_features.shape)
print(train_targets_scored.shape)
print(train_targets_nonscored.shape)
print(test_features.shape)
print(sample_submission.shape)
# + papermill={"duration": 0.094918, "end_time": "2020-11-16T15:32:28.705597", "exception": false, "start_time": "2020-11-16T15:32:28.610679", "status": "completed"} tags=[]
GENES = [col for col in train_features.columns if col.startswith('g-')]
CELLS = [col for col in train_features.columns if col.startswith('c-')]
# + papermill={"duration": 0.104299, "end_time": "2020-11-16T15:32:28.894146", "exception": false, "start_time": "2020-11-16T15:32:28.789847", "status": "completed"} tags=[]
def seed_everything(seed=1903):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.deterministic = True
seed_everything(seed=1903)
# + papermill={"duration": 0.058739, "end_time": "2020-11-16T15:32:29.026640", "exception": false, "start_time": "2020-11-16T15:32:28.967901", "status": "completed"} tags=[]
# + papermill={"duration": 271.569729, "end_time": "2020-11-16T15:37:00.654972", "exception": false, "start_time": "2020-11-16T15:32:29.085243", "status": "completed"} tags=[]
# GENES
n_comp = 90
n_dim = 45
data = pd.concat([pd.DataFrame(train_features[GENES]), pd.DataFrame(test_features[GENES])])
if IS_TRAIN:
fa = FactorAnalysis(n_components=n_comp, random_state=1903).fit(data[GENES])
pd.to_pickle(fa, f'{MODEL_DIR}/{NB}_factor_analysis_g.pkl')
umap = UMAP(n_components=n_dim, random_state=1903).fit(data[GENES])
pd.to_pickle(umap, f'{MODEL_DIR}/{NB}_umap_g.pkl')
else:
fa = pd.read_pickle(f'{MODEL_DIR}/{NB}_factor_analysis_g.pkl')
umap = pd.read_pickle(f'{MODEL_DIR}/{NB}_umap_g.pkl')
data2 = (fa.transform(data[GENES]))
data3 = (umap.transform(data[GENES]))
train2 = data2[:train_features.shape[0]]
test2 = data2[-test_features.shape[0]:]
train3 = data3[:train_features.shape[0]]
test3 = data3[-test_features.shape[0]:]
train2 = pd.DataFrame(train2, columns=[f'fa_G-{i}' for i in range(n_comp)])
train3 = pd.DataFrame(train3, columns=[f'umap_G-{i}' for i in range(n_dim)])
test2 = pd.DataFrame(test2, columns=[f'fa_G-{i}' for i in range(n_comp)])
test3 = pd.DataFrame(test3, columns=[f'umap_G-{i}' for i in range(n_dim)])
train_features = pd.concat((train_features, train2, train3), axis=1)
test_features = pd.concat((test_features, test2, test3), axis=1)
#CELLS
n_comp = 50
n_dim = 25
data = pd.concat([pd.DataFrame(train_features[CELLS]), pd.DataFrame(test_features[CELLS])])
if IS_TRAIN:
fa = FactorAnalysis(n_components=n_comp, random_state=1903).fit(data[CELLS])
pd.to_pickle(fa, f'{MODEL_DIR}/{NB}_factor_analysis_c.pkl')
umap = UMAP(n_components=n_dim, random_state=1903).fit(data[CELLS])
pd.to_pickle(umap, f'{MODEL_DIR}/{NB}_umap_c.pkl')
else:
fa = pd.read_pickle(f'{MODEL_DIR}/{NB}_factor_analysis_c.pkl')
umap = pd.read_pickle(f'{MODEL_DIR}/{NB}_umap_c.pkl')
data2 = (fa.transform(data[CELLS]))
data3 = (umap.fit_transform(data[CELLS]))
train2 = data2[:train_features.shape[0]]
test2 = data2[-test_features.shape[0]:]
train3 = data3[:train_features.shape[0]]
test3 = data3[-test_features.shape[0]:]
train2 = pd.DataFrame(train2, columns=[f'fa_C-{i}' for i in range(n_comp)])
train3 = pd.DataFrame(train3, columns=[f'umap_C-{i}' for i in range(n_dim)])
test2 = pd.DataFrame(test2, columns=[f'fa_C-{i}' for i in range(n_comp)])
test3 = pd.DataFrame(test3, columns=[f'umap_C-{i}' for i in range(n_dim)])
train_features = pd.concat((train_features, train2, train3), axis=1)
test_features = pd.concat((test_features, test2, test3), axis=1)
# drop_cols = [f'c-{i}' for i in range(n_comp,len(CELLS))]
# + papermill={"duration": 0.059235, "end_time": "2020-11-16T15:37:00.774014", "exception": false, "start_time": "2020-11-16T15:37:00.714779", "status": "completed"} tags=[]
# + papermill={"duration": 120.954735, "end_time": "2020-11-16T15:39:01.787539", "exception": false, "start_time": "2020-11-16T15:37:00.832804", "status": "completed"} tags=[]
from sklearn.preprocessing import QuantileTransformer
for col in (GENES + CELLS):
vec_len = len(train_features[col].values)
vec_len_test = len(test_features[col].values)
raw_vec = pd.concat([train_features, test_features])[col].values.reshape(vec_len+vec_len_test, 1)
if IS_TRAIN:
transformer = QuantileTransformer(n_quantiles=100, random_state=123, output_distribution="normal")
transformer.fit(raw_vec)
pd.to_pickle(transformer, f'{MODEL_DIR}/{NB}_{col}_quantile_transformer.pkl')
else:
transformer = pd.read_pickle(f'{MODEL_DIR}/{NB}_{col}_quantile_transformer.pkl')
train_features[col] = transformer.transform(train_features[col].values.reshape(vec_len, 1)).reshape(1, vec_len)[0]
test_features[col] = transformer.transform(test_features[col].values.reshape(vec_len_test, 1)).reshape(1, vec_len_test)[0]
# + papermill={"duration": 0.069192, "end_time": "2020-11-16T15:39:01.917099", "exception": false, "start_time": "2020-11-16T15:39:01.847907", "status": "completed"} tags=[]
# PCAS = [col for col in train_features.columns if col.startswith('pca_')]
# UMAPS = [col for col in train_features.columns if col.startswith('umap_')]
# + papermill={"duration": 0.078788, "end_time": "2020-11-16T15:39:02.058710", "exception": false, "start_time": "2020-11-16T15:39:01.979922", "status": "completed"} tags=[]
# from sklearn.preprocessing import PolynomialFeatures
# n_deg = 2
# data = pd.concat([pd.DataFrame(train_features[PCAS]), pd.DataFrame(test_features[PCAS])])
# data2 = (PolynomialFeatures(degree=n_deg, include_bias=False).fit_transform(data[PCAS]))
# # print(data2)
# # data4 = (UMAP(n_components=n_dim, n_neighbors=5, random_state=1903).fit_transform(data[GENES]))
# # data5 = (UMAP(n_components=n_dim, min_dist=0.01, random_state=1903).fit_transform(data[GENES]))
# train2 = data2[:train_features.shape[0]]
# test2 = data2[-test_features.shape[0]:]
# # print(train2.shape)
# train2 = pd.DataFrame(train2, columns=[f'poly_C-{i}' for i in range(train2.shape[1])])
# test2 = pd.DataFrame(test2, columns=[f'poly_C-{i}' for i in range(train2.shape[1])])
# # drop_cols = [f'c-{i}' for i in range(n_comp,len(GENES))]
# # train_features = pd.concat((train_features, train2, train3, train4, train5), axis=1)
# # test_features = pd.concat((test_features, test2, test3, test4, test5), axis=1)
# train_features = pd.concat((train_features, train2), axis=1)
# test_features = pd.concat((test_features, test2), axis=1)
# data = pd.concat([pd.DataFrame(train_features[UMAPS]), pd.DataFrame(test_features[UMAPS])])
# data2 = (PolynomialFeatures(degree=n_deg, include_bias=False).fit_transform(data[UMAPS]))
# # print(data2)
# # data4 = (UMAP(n_components=n_dim, n_neighbors=5, random_state=1903).fit_transform(data[GENES]))
# # data5 = (UMAP(n_components=n_dim, min_dist=0.01, random_state=1903).fit_transform(data[GENES]))
# train2 = data2[:train_features.shape[0]]
# test2 = data2[-test_features.shape[0]:]
# # print(train2.shape)
# train2 = pd.DataFrame(train2, columns=[f'poly_C-{i}' for i in range(train2.shape[1])])
# test2 = pd.DataFrame(test2, columns=[f'poly_C-{i}' for i in range(train2.shape[1])])
# # drop_cols = [f'c-{i}' for i in range(n_comp,len(GENES))]
# # train_features = pd.concat((train_features, train2, train3, train4, train5), axis=1)
# # test_features = pd.concat((test_features, test2, test3, test4, test5), axis=1)
# train_features = pd.concat((train_features, train2), axis=1)
# test_features = pd.concat((test_features, test2), axis=1)
# + papermill={"duration": 0.069787, "end_time": "2020-11-16T15:39:02.198165", "exception": false, "start_time": "2020-11-16T15:39:02.128378", "status": "completed"} tags=[]
print(train_features.shape)
print(test_features.shape)
# + papermill={"duration": 0.058859, "end_time": "2020-11-16T15:39:02.316948", "exception": false, "start_time": "2020-11-16T15:39:02.258089", "status": "completed"} tags=[]
# + papermill={"duration": 0.865346, "end_time": "2020-11-16T15:39:03.241502", "exception": false, "start_time": "2020-11-16T15:39:02.376156", "status": "completed"} tags=[]
# train = train_features.merge(train_targets_scored, on='sig_id')
train = train_features.merge(train_targets_nonscored, on='sig_id')
train = train[train['cp_type']!='ctl_vehicle'].reset_index(drop=True)
test = test_features[test_features['cp_type']!='ctl_vehicle'].reset_index(drop=True)
# target = train[train_targets_scored.columns]
target = train[train_targets_nonscored.columns]
# + papermill={"duration": 0.148851, "end_time": "2020-11-16T15:39:03.473560", "exception": false, "start_time": "2020-11-16T15:39:03.324709", "status": "completed"} tags=[]
train = train.drop('cp_type', axis=1)
test = test.drop('cp_type', axis=1)
# + papermill={"duration": 0.07072, "end_time": "2020-11-16T15:39:03.628960", "exception": false, "start_time": "2020-11-16T15:39:03.558240", "status": "completed"} tags=[]
print(target.shape)
print(train_features.shape)
print(test_features.shape)
print(train.shape)
print(test.shape)
# + papermill={"duration": 0.081147, "end_time": "2020-11-16T15:39:03.774324", "exception": false, "start_time": "2020-11-16T15:39:03.693177", "status": "completed"} tags=[]
target_cols = target.drop('sig_id', axis=1).columns.values.tolist()
# + papermill={"duration": 2.963638, "end_time": "2020-11-16T15:39:06.798361", "exception": false, "start_time": "2020-11-16T15:39:03.834723", "status": "completed"} tags=[]
folds = train.copy()
mskf = MultilabelStratifiedKFold(n_splits=NFOLDS)
for f, (t_idx, v_idx) in enumerate(mskf.split(X=train, y=target)):
folds.loc[v_idx, 'kfold'] = int(f)
folds['kfold'] = folds['kfold'].astype(int)
folds
# + papermill={"duration": 0.078361, "end_time": "2020-11-16T15:39:06.940337", "exception": false, "start_time": "2020-11-16T15:39:06.861976", "status": "completed"} tags=[]
print(train.shape)
print(folds.shape)
print(test.shape)
print(target.shape)
print(sample_submission.shape)
# + papermill={"duration": 0.078343, "end_time": "2020-11-16T15:39:07.088435", "exception": false, "start_time": "2020-11-16T15:39:07.010092", "status": "completed"} tags=[]
class MoADataset:
def __init__(self, features, targets):
self.features = features
self.targets = targets
def __len__(self):
return (self.features.shape[0])
def __getitem__(self, idx):
dct = {
'x' : torch.tensor(self.features[idx, :], dtype=torch.float),
'y' : torch.tensor(self.targets[idx, :], dtype=torch.float)
}
return dct
class TestDataset:
def __init__(self, features):
self.features = features
def __len__(self):
return (self.features.shape[0])
def __getitem__(self, idx):
dct = {
'x' : torch.tensor(self.features[idx, :], dtype=torch.float)
}
return dct
# + papermill={"duration": 0.082009, "end_time": "2020-11-16T15:39:07.236374", "exception": false, "start_time": "2020-11-16T15:39:07.154365", "status": "completed"} tags=[]
def train_fn(model, optimizer, scheduler, loss_fn, dataloader, device):
model.train()
final_loss = 0
for data in dataloader:
optimizer.zero_grad()
inputs, targets = data['x'].to(device), data['y'].to(device)
# print(inputs.shape)
outputs = model(inputs)
loss = loss_fn(outputs, targets)
loss.backward()
optimizer.step()
scheduler.step()
final_loss += loss.item()
final_loss /= len(dataloader)
return final_loss
def valid_fn(model, loss_fn, dataloader, device):
model.eval()
final_loss = 0
valid_preds = []
for data in dataloader:
inputs, targets = data['x'].to(device), data['y'].to(device)
outputs = model(inputs)
loss = loss_fn(outputs, targets)
final_loss += loss.item()
valid_preds.append(outputs.sigmoid().detach().cpu().numpy())
final_loss /= len(dataloader)
valid_preds = np.concatenate(valid_preds)
return final_loss, valid_preds
def inference_fn(model, dataloader, device):
model.eval()
preds = []
for data in dataloader:
inputs = data['x'].to(device)
with torch.no_grad():
outputs = model(inputs)
preds.append(outputs.sigmoid().detach().cpu().numpy())
preds = np.concatenate(preds)
return preds
# + papermill={"duration": 0.077352, "end_time": "2020-11-16T15:39:07.376366", "exception": false, "start_time": "2020-11-16T15:39:07.299014", "status": "completed"} tags=[]
class Model(nn.Module):
def __init__(self, num_features, num_targets, hidden_size):
super(Model, self).__init__()
self.batch_norm1 = nn.BatchNorm1d(num_features)
self.dropout1 = nn.Dropout(0.15)
self.dense1 = nn.utils.weight_norm(nn.Linear(num_features, hidden_size))
self.batch_norm2 = nn.BatchNorm1d(hidden_size)
self.dropout2 = nn.Dropout(0.3)
self.dense2 = nn.Linear(hidden_size, hidden_size)
self.batch_norm3 = nn.BatchNorm1d(hidden_size)
self.dropout3 = nn.Dropout(0.25)
self.dense3 = nn.utils.weight_norm(nn.Linear(hidden_size, num_targets))
def forward(self, x):
x = self.batch_norm1(x)
x = self.dropout1(x)
x = F.leaky_relu(self.dense1(x))
x = self.batch_norm2(x)
x = self.dropout2(x)
x = F.leaky_relu(self.dense2(x))
x = self.batch_norm3(x)
x = self.dropout3(x)
x = self.dense3(x)
return x
# + papermill={"duration": 0.071597, "end_time": "2020-11-16T15:39:07.510415", "exception": false, "start_time": "2020-11-16T15:39:07.438818", "status": "completed"} tags=[]
def process_data(data):
data = pd.get_dummies(data, columns=['cp_time','cp_dose'])
# data.loc[:, 'cp_time'] = data.loc[:, 'cp_time'].map({24: 0, 48: 1, 72: 2})
# data.loc[:, 'cp_dose'] = data.loc[:, 'cp_dose'].map({'D1': 0, 'D2': 1})
# --------------------- Normalize ---------------------
# for col in GENES:
# data[col] = (data[col]-np.mean(data[col])) / (np.std(data[col]))
# for col in CELLS:
# data[col] = (data[col]-np.mean(data[col])) / (np.std(data[col]))
#--------------------- Removing Skewness ---------------------
# for col in GENES + CELLS:
# if(abs(data[col].skew()) > 0.75):
# if(data[col].skew() < 0): # neg-skewness
# data[col] = data[col].max() - data[col] + 1
# data[col] = np.sqrt(data[col])
# else:
# data[col] = np.sqrt(data[col])
return data
# + papermill={"duration": 0.215368, "end_time": "2020-11-16T15:39:07.817895", "exception": false, "start_time": "2020-11-16T15:39:07.602527", "status": "completed"} tags=[]
feature_cols = [c for c in process_data(folds).columns if c not in target_cols]
feature_cols = [c for c in feature_cols if c not in ['kfold','sig_id']]
len(feature_cols)
# + papermill={"duration": 0.072169, "end_time": "2020-11-16T15:39:07.953285", "exception": false, "start_time": "2020-11-16T15:39:07.881116", "status": "completed"} tags=[]
num_features=len(feature_cols)
num_targets=len(target_cols)
hidden_size=2048
# hidden_size=4096
# hidden_size=9192
# + papermill={"duration": 0.094675, "end_time": "2020-11-16T15:39:08.111587", "exception": false, "start_time": "2020-11-16T15:39:08.016912", "status": "completed"} tags=[]
def run_training(fold, seed):
seed_everything(seed)
train = process_data(folds)
test_ = process_data(test)
trn_idx = train[train['kfold'] != fold].index
val_idx = train[train['kfold'] == fold].index
train_df = train[train['kfold'] != fold].reset_index(drop=True)
valid_df = train[train['kfold'] == fold].reset_index(drop=True)
x_train, y_train = train_df[feature_cols].values, train_df[target_cols].values
x_valid, y_valid = valid_df[feature_cols].values, valid_df[target_cols].values
train_dataset = MoADataset(x_train, y_train)
valid_dataset = MoADataset(x_valid, y_valid)
trainloader = torch.utils.data.DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True)
validloader = torch.utils.data.DataLoader(valid_dataset, batch_size=BATCH_SIZE, shuffle=False)
model = Model(
num_features=num_features,
num_targets=num_targets,
hidden_size=hidden_size,
)
model.to(DEVICE)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE, weight_decay=WEIGHT_DECAY)
# scheduler = optim.lr_scheduler.OneCycleLR(optimizer=optimizer, pct_start=0.3, div_factor=1000,
# max_lr=1e-2, epochs=EPOCHS, steps_per_epoch=len(trainloader))
scheduler = optim.lr_scheduler.OneCycleLR(optimizer=optimizer, pct_start=0.2, div_factor=1e3,
max_lr=1e-2, epochs=EPOCHS, steps_per_epoch=len(trainloader))
loss_fn = nn.BCEWithLogitsLoss()
early_stopping_steps = EARLY_STOPPING_STEPS
early_step = 0
oof = np.zeros((len(train), target.iloc[:, 1:].shape[1]))
best_loss = np.inf
best_loss_epoch = -1
if IS_TRAIN:
for epoch in range(EPOCHS):
train_loss = train_fn(model, optimizer, scheduler, loss_fn, trainloader, DEVICE)
valid_loss, valid_preds = valid_fn(model, loss_fn, validloader, DEVICE)
if valid_loss < best_loss:
best_loss = valid_loss
best_loss_epoch = epoch
oof[val_idx] = valid_preds
torch.save(model.state_dict(), f"{MODEL_DIR}/{NB}-nonscored-SEED{seed}-FOLD{fold}_.pth")
elif(EARLY_STOP == True):
early_step += 1
if (early_step >= early_stopping_steps):
break
if epoch % 10 == 0 or epoch == EPOCHS-1:
print(f"seed: {seed}, FOLD: {fold}, EPOCH: {epoch}, train_loss: {train_loss:.6f}, valid_loss: {valid_loss:.6f}, best_loss: {best_loss:.6f}, best_loss_epoch: {best_loss_epoch}")
#--------------------- PREDICTION---------------------
x_test = test_[feature_cols].values
testdataset = TestDataset(x_test)
testloader = torch.utils.data.DataLoader(testdataset, batch_size=BATCH_SIZE, shuffle=False)
model = Model(
num_features=num_features,
num_targets=num_targets,
hidden_size=hidden_size,
)
model.load_state_dict(torch.load(f"{MODEL_DIR}/{NB}-nonscored-SEED{seed}-FOLD{fold}_.pth"))
model.to(DEVICE)
if not IS_TRAIN:
valid_loss, valid_preds = valid_fn(model, loss_fn, validloader, DEVICE)
oof[val_idx] = valid_preds
predictions = np.zeros((len(test_), target.iloc[:, 1:].shape[1]))
predictions = inference_fn(model, testloader, DEVICE)
return oof, predictions
# + papermill={"duration": 0.073356, "end_time": "2020-11-16T15:39:08.253237", "exception": false, "start_time": "2020-11-16T15:39:08.179881", "status": "completed"} tags=[]
def run_k_fold(NFOLDS, seed):
oof = np.zeros((len(train), len(target_cols)))
predictions = np.zeros((len(test), len(target_cols)))
for fold in range(NFOLDS):
oof_, pred_ = run_training(fold, seed)
predictions += pred_ / NFOLDS
oof += oof_
return oof, predictions
# + papermill={"duration": 522.044473, "end_time": "2020-11-16T15:47:50.364869", "exception": false, "start_time": "2020-11-16T15:39:08.320396", "status": "completed"} tags=[]
SEED = range(NSEEDS) #[0, 1, 2, 3 ,4]#, 5, 6, 7, 8, 9, 10]
oof = np.zeros((len(train), len(target_cols)))
predictions = np.zeros((len(test), len(target_cols)))
time_start = time.time()
for seed in SEED:
oof_, predictions_ = run_k_fold(NFOLDS, seed)
oof += oof_ / len(SEED)
predictions += predictions_ / len(SEED)
print(f"elapsed time: {time.time() - time_start}")
train[target_cols] = oof
test[target_cols] = predictions
print(oof.shape)
print(predictions.shape)
# + papermill={"duration": 0.7287, "end_time": "2020-11-16T15:47:51.194273", "exception": false, "start_time": "2020-11-16T15:47:50.465573", "status": "completed"} tags=[]
train.to_pickle(f"{INT_DIR}/{NB}-train_nonscore_pred.pkl")
test.to_pickle(f"{INT_DIR}/{NB}-test_nonscore_pred.pkl")
# + papermill={"duration": 0.105139, "end_time": "2020-11-16T15:47:51.400195", "exception": false, "start_time": "2020-11-16T15:47:51.295056", "status": "completed"} tags=[]
len(target_cols)
# + papermill={"duration": 2.221951, "end_time": "2020-11-16T15:47:53.717415", "exception": false, "start_time": "2020-11-16T15:47:51.495464", "status": "completed"} tags=[]
train[target_cols] = np.maximum(PMIN, np.minimum(PMAX, train[target_cols]))
valid_results = train_targets_nonscored.drop(columns=target_cols).merge(train[['sig_id']+target_cols], on='sig_id', how='left').fillna(0)
y_true = train_targets_nonscored[target_cols].values
y_true = y_true > 0.5
y_pred = valid_results[target_cols].values
score = 0
for i in range(len(target_cols)):
score_ = log_loss(y_true[:, i], y_pred[:, i])
score += score_ / target.shape[1]
print("CV log_loss: ", score)
# + [markdown] papermill={"duration": 0.094811, "end_time": "2020-11-16T15:47:53.910977", "exception": false, "start_time": "2020-11-16T15:47:53.816166", "status": "completed"} tags=[]
# CV log_loss: 0.014761779358699672
# CV log_loss: 0.014519859174255039
# CV log_loss: 0.014525173864593479
# CV log_loss: 0.014354930596928602 # 3 umap features
# CV log_loss: 0.014353604854355429 # more umap features
# CV log_loss: 0.01436484670778641 # more hidden nodes
# + papermill={"duration": 0.104858, "end_time": "2020-11-16T15:47:54.110885", "exception": false, "start_time": "2020-11-16T15:47:54.006027", "status": "completed"} tags=[]
EPOCHS = 25
# NFOLDS = 5
# + papermill={"duration": 0.103088, "end_time": "2020-11-16T15:47:54.310590", "exception": false, "start_time": "2020-11-16T15:47:54.207502", "status": "completed"} tags=[]
# sub = sample_submission.drop(columns=target_cols).merge(test[['sig_id']+target_cols], on='sig_id', how='left').fillna(0)
# sub.to_csv('submission.csv', index=False)
# + papermill={"duration": 0.125327, "end_time": "2020-11-16T15:47:54.532597", "exception": false, "start_time": "2020-11-16T15:47:54.407270", "status": "completed"} tags=[]
nonscored_target = [c for c in train[train_targets_nonscored.columns] if c != "sig_id"]
# + papermill={"duration": 0.126509, "end_time": "2020-11-16T15:47:54.768823", "exception": false, "start_time": "2020-11-16T15:47:54.642314", "status": "completed"} tags=[]
nonscored_target
# + papermill={"duration": 0.476692, "end_time": "2020-11-16T15:47:55.348157", "exception": false, "start_time": "2020-11-16T15:47:54.871465", "status": "completed"} tags=[]
train = pd.read_pickle(f"{INT_DIR}/{NB}-train_nonscore_pred.pkl")
test = pd.read_pickle(f"{INT_DIR}/{NB}-test_nonscore_pred.pkl")
# + papermill={"duration": 0.683948, "end_time": "2020-11-16T15:47:56.130247", "exception": false, "start_time": "2020-11-16T15:47:55.446299", "status": "completed"} tags=[]
# use nonscored target in the given file as feature
# if comment out below, use predicted nonscored target
# train = train.drop(nonscored_target, axis=1)
# train = train.merge(train_targets_nonscored, on="sig_id")
# train = train_features.merge(train_targets_scored, on='sig_id')
train = train.merge(train_targets_scored, on='sig_id')
# train = train[train['cp_type']!='ctl_vehicle'].reset_index(drop=True)
# test = test[test['cp_type']!='ctl_vehicle'].reset_index(drop=True)
# target = train[train_targets_scored.columns]
target = train[train_targets_scored.columns]
# + papermill={"duration": 3.539532, "end_time": "2020-11-16T15:47:59.773101", "exception": false, "start_time": "2020-11-16T15:47:56.233569", "status": "completed"} tags=[]
# from sklearn.preprocessing import QuantileTransformer
for col in (nonscored_target):
vec_len = len(train[col].values)
vec_len_test = len(test[col].values)
raw_vec = train[col].values.reshape(vec_len, 1)
if IS_TRAIN:
transformer = QuantileTransformer(n_quantiles=100, random_state=0, output_distribution="normal")
transformer.fit(raw_vec)
pd.to_pickle(transformer, f"{MODEL_DIR}/{NB}_{col}_quantile_nonscored.pkl")
else:
transformer = pd.read_pickle(f"{MODEL_DIR}/{NB}_{col}_quantile_nonscored.pkl")
train[col] = transformer.transform(raw_vec).reshape(1, vec_len)[0]
test[col] = transformer.transform(test[col].values.reshape(vec_len_test, 1)).reshape(1, vec_len_test)[0]
# + papermill={"duration": 0.145033, "end_time": "2020-11-16T15:48:00.029050", "exception": false, "start_time": "2020-11-16T15:47:59.884017", "status": "completed"} tags=[]
target_cols = target.drop('sig_id', axis=1).columns.values.tolist()
# + papermill={"duration": 0.144341, "end_time": "2020-11-16T15:48:00.274053", "exception": false, "start_time": "2020-11-16T15:48:00.129712", "status": "completed"} tags=[]
train
# + papermill={"duration": 4.259285, "end_time": "2020-11-16T15:48:04.644957", "exception": false, "start_time": "2020-11-16T15:48:00.385672", "status": "completed"} tags=[]
folds = train.copy()
mskf = MultilabelStratifiedKFold(n_splits=NFOLDS)
for f, (t_idx, v_idx) in enumerate(mskf.split(X=train, y=target)):
folds.loc[v_idx, 'kfold'] = int(f)
folds['kfold'] = folds['kfold'].astype(int)
folds
# + papermill={"duration": 0.125759, "end_time": "2020-11-16T15:48:04.880143", "exception": false, "start_time": "2020-11-16T15:48:04.754384", "status": "completed"} tags=[]
print(train.shape)
print(folds.shape)
print(test.shape)
print(target.shape)
print(sample_submission.shape)
# + papermill={"duration": 0.123118, "end_time": "2020-11-16T15:48:05.130247", "exception": false, "start_time": "2020-11-16T15:48:05.007129", "status": "completed"} tags=[]
def process_data(data):
data = pd.get_dummies(data, columns=['cp_time','cp_dose'])
# data.loc[:, 'cp_time'] = data.loc[:, 'cp_time'].map({24: 0, 48: 1, 72: 2})
# data.loc[:, 'cp_dose'] = data.loc[:, 'cp_dose'].map({'D1': 0, 'D2': 1})
# --------------------- Normalize ---------------------
# for col in GENES:
# data[col] = (data[col]-np.mean(data[col])) / (np.std(data[col]))
# for col in CELLS:
# data[col] = (data[col]-np.mean(data[col])) / (np.std(data[col]))
#--------------------- Removing Skewness ---------------------
# for col in GENES + CELLS:
# if(abs(data[col].skew()) > 0.75):
# if(data[col].skew() < 0): # neg-skewness
# data[col] = data[col].max() - data[col] + 1
# data[col] = np.sqrt(data[col])
# else:
# data[col] = np.sqrt(data[col])
return data
# + papermill={"duration": 0.359577, "end_time": "2020-11-16T15:48:05.594014", "exception": false, "start_time": "2020-11-16T15:48:05.234437", "status": "completed"} tags=[]
feature_cols = [c for c in process_data(folds).columns if c not in target_cols]
feature_cols = [c for c in feature_cols if c not in ['kfold','sig_id']]
len(feature_cols)
# + papermill={"duration": 0.160731, "end_time": "2020-11-16T15:48:05.861292", "exception": false, "start_time": "2020-11-16T15:48:05.700561", "status": "completed"} tags=[]
num_features=len(feature_cols)
num_targets=len(target_cols)
hidden_size=2048
# hidden_size=4096
# hidden_size=9192
# + papermill={"duration": 0.18558, "end_time": "2020-11-16T15:48:06.198381", "exception": false, "start_time": "2020-11-16T15:48:06.012801", "status": "completed"} tags=[]
def run_training(fold, seed):
seed_everything(seed)
train = process_data(folds)
test_ = process_data(test)
trn_idx = train[train['kfold'] != fold].index
val_idx = train[train['kfold'] == fold].index
train_df = train[train['kfold'] != fold].reset_index(drop=True)
valid_df = train[train['kfold'] == fold].reset_index(drop=True)
x_train, y_train = train_df[feature_cols].values, train_df[target_cols].values
x_valid, y_valid = valid_df[feature_cols].values, valid_df[target_cols].values
train_dataset = MoADataset(x_train, y_train)
valid_dataset = MoADataset(x_valid, y_valid)
trainloader = torch.utils.data.DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True)
validloader = torch.utils.data.DataLoader(valid_dataset, batch_size=BATCH_SIZE, shuffle=False)
model = Model(
num_features=num_features,
num_targets=num_targets,
hidden_size=hidden_size,
)
model.to(DEVICE)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE, weight_decay=WEIGHT_DECAY)
# scheduler = optim.lr_scheduler.OneCycleLR(optimizer=optimizer, pct_start=0.3, div_factor=1000,
# max_lr=1e-2, epochs=EPOCHS, steps_per_epoch=len(trainloader))
scheduler = optim.lr_scheduler.OneCycleLR(optimizer=optimizer, pct_start=0.2, div_factor=1e3,
max_lr=1e-2, epochs=EPOCHS, steps_per_epoch=len(trainloader))
loss_fn = nn.BCEWithLogitsLoss()
early_stopping_steps = EARLY_STOPPING_STEPS
early_step = 0
oof = np.zeros((len(train), target.iloc[:, 1:].shape[1]))
best_loss = np.inf
best_loss_epoch = -1
if IS_TRAIN:
for epoch in range(EPOCHS):
train_loss = train_fn(model, optimizer, scheduler, loss_fn, trainloader, DEVICE)
valid_loss, valid_preds = valid_fn(model, loss_fn, validloader, DEVICE)
if valid_loss < best_loss:
best_loss = valid_loss
best_loss_epoch = epoch
oof[val_idx] = valid_preds
torch.save(model.state_dict(), f"{MODEL_DIR}/{NB}-scored-SEED{seed}-FOLD{fold}_.pth")
elif(EARLY_STOP == True):
early_step += 1
if (early_step >= early_stopping_steps):
break
if epoch % 10 == 0 or epoch == EPOCHS-1:
print(f"seed: {seed}, FOLD: {fold}, EPOCH: {epoch}, train_loss: {train_loss:.6f}, valid_loss: {valid_loss:.6f}, best_loss: {best_loss:.6f}, best_loss_epoch: {best_loss_epoch}")
#--------------------- PREDICTION---------------------
x_test = test_[feature_cols].values
testdataset = TestDataset(x_test)
testloader = torch.utils.data.DataLoader(testdataset, batch_size=BATCH_SIZE, shuffle=False)
model = Model(
num_features=num_features,
num_targets=num_targets,
hidden_size=hidden_size,
)
model.load_state_dict(torch.load(f"{MODEL_DIR}/{NB}-scored-SEED{seed}-FOLD{fold}_.pth"))
model.to(DEVICE)
if not IS_TRAIN:
valid_loss, valid_preds = valid_fn(model, loss_fn, validloader, DEVICE)
oof[val_idx] = valid_preds
predictions = np.zeros((len(test_), target.iloc[:, 1:].shape[1]))
predictions = inference_fn(model, testloader, DEVICE)
return oof, predictions
# + papermill={"duration": 0.163112, "end_time": "2020-11-16T15:48:06.522900", "exception": false, "start_time": "2020-11-16T15:48:06.359788", "status": "completed"} tags=[]
def run_k_fold(NFOLDS, seed):
oof = np.zeros((len(train), len(target_cols)))
predictions = np.zeros((len(test), len(target_cols)))
for fold in range(NFOLDS):
oof_, pred_ = run_training(fold, seed)
predictions += pred_ / NFOLDS
oof += oof_
return oof, predictions
# + papermill={"duration": 924.623178, "end_time": "2020-11-16T16:03:31.245873", "exception": false, "start_time": "2020-11-16T15:48:06.622695", "status": "completed"} tags=[]
SEED = range(NSEEDS) #[0, 1, 2, 3 ,4]#, 5, 6, 7, 8, 9, 10]
oof = np.zeros((len(train), len(target_cols)))
predictions = np.zeros((len(test), len(target_cols)))
time_start = time.time()
for seed in SEED:
oof_, predictions_ = run_k_fold(NFOLDS, seed)
oof += oof_ / len(SEED)
predictions += predictions_ / len(SEED)
print(f"elapsed time: {time.time() - time_start}")
train[target_cols] = oof
test[target_cols] = predictions
# + papermill={"duration": 0.879941, "end_time": "2020-11-16T16:03:32.272105", "exception": false, "start_time": "2020-11-16T16:03:31.392164", "status": "completed"} tags=[]
train.to_pickle(f"{INT_DIR}/{NB}-train-score-pred.pkl")
test.to_pickle(f"{INT_DIR}/{NB}-test-score-pred.pkl")
# + papermill={"duration": 0.149996, "end_time": "2020-11-16T16:03:32.570961", "exception": false, "start_time": "2020-11-16T16:03:32.420965", "status": "completed"} tags=[]
len(target_cols)
# + papermill={"duration": 1.489532, "end_time": "2020-11-16T16:03:34.202115", "exception": false, "start_time": "2020-11-16T16:03:32.712583", "status": "completed"} tags=[]
train[target_cols] = np.maximum(PMIN, np.minimum(PMAX, train[target_cols]))
valid_results = train_targets_scored.drop(columns=target_cols).merge(train[['sig_id']+target_cols], on='sig_id', how='left').fillna(0)
y_true = train_targets_scored[target_cols].values
y_true = y_true > 0.5
y_pred = valid_results[target_cols].values
score = 0
for i in range(len(target_cols)):
score_ = log_loss(y_true[:, i], y_pred[:, i])
score += score_ / target.shape[1]
print("CV log_loss: ", score)
# + [markdown] papermill={"duration": 0.140342, "end_time": "2020-11-16T16:03:34.483167", "exception": false, "start_time": "2020-11-16T16:03:34.342825", "status": "completed"} tags=[]
# - CV log_loss: 0.014761779358699672
# - CV log_loss: 0.014519859174255039
# - CV log_loss: 0.014525173864593479
# - CV log_loss: 0.014354930596928602 # 3 umap features
# - CV log_loss: 0.014353604854355429 # more umap features
# - CV log_loss: 0.01436484670778641 # more hidden nodes
# - CV log_loss: 0.014344688083211073
# - using predicted unscored targets as feature
# - CV log_loss: 0.013368097791623873
# - using given unscored targets as feature
# - bad in public lb
# - CV log_loss: 0.01434373547175235
# - rankgauss predicted unscored targets
# - CV log_loss: 0.014346100008158216
# - unscored targets pca/umap
# - CV log_loss: 0.014328486629791769
# - NFOLDS=10, Epoch=20
# - CV log_loss: 0.014299741080816082
# - NFOLDS=10, Epoch=20, 25
# - CV log_loss: 0.014311301224480969
# - NFOLDS=10, Epoch=25
# - CV log_loss: 0.01429269446076626
# - NFOLDS=10, Epoch=15, 25
# + papermill={"duration": 0.149808, "end_time": "2020-11-16T16:03:34.777462", "exception": false, "start_time": "2020-11-16T16:03:34.627654", "status": "completed"} tags=[]
# train = pd.read_pickle(f"../interim/23-train-score-pred.pkl")
# test = pd.read_pickle(f"../interim/23-test-score-pred.pkl")
# + papermill={"duration": 0.601558, "end_time": "2020-11-16T16:03:35.519666", "exception": false, "start_time": "2020-11-16T16:03:34.918108", "status": "completed"} tags=[]
train = pd.read_pickle(f"{INT_DIR}/{NB}-train-score-pred.pkl")
test = pd.read_pickle(f"{INT_DIR}/{NB}-test-score-pred.pkl")
# + papermill={"duration": 0.151029, "end_time": "2020-11-16T16:03:35.814627", "exception": false, "start_time": "2020-11-16T16:03:35.663598", "status": "completed"} tags=[]
EPOCHS = 25
# NFOLDS = 5
# + papermill={"duration": 0.966342, "end_time": "2020-11-16T16:03:36.920644", "exception": false, "start_time": "2020-11-16T16:03:35.954302", "status": "completed"} tags=[]
PMIN = 0.0005
PMAX = 0.9995
for c in train_targets_scored.columns:
if c != "sig_id":
train_targets_scored[c] = np.maximum(PMIN, np.minimum(PMAX, train_targets_scored[c]))
# + papermill={"duration": 0.160927, "end_time": "2020-11-16T16:03:37.230306", "exception": false, "start_time": "2020-11-16T16:03:37.069379", "status": "completed"} tags=[]
train_targets_scored.columns
# + papermill={"duration": 0.594604, "end_time": "2020-11-16T16:03:37.971540", "exception": false, "start_time": "2020-11-16T16:03:37.376936", "status": "completed"} tags=[]
train = train[train_targets_scored.columns]
train.columns = [c + "_pred" if (c != 'sig_id' and c in train_targets_scored.columns) else c for c in train.columns]
# + papermill={"duration": 0.468599, "end_time": "2020-11-16T16:03:38.595101", "exception": false, "start_time": "2020-11-16T16:03:38.126502", "status": "completed"} tags=[]
test = test[train_targets_scored.columns]
test.columns = [c + "_pred" if (c != 'sig_id' and c in train_targets_scored.columns) else c for c in test.columns]
# + papermill={"duration": 0.215285, "end_time": "2020-11-16T16:03:39.196380", "exception": false, "start_time": "2020-11-16T16:03:38.981095", "status": "completed"} tags=[]
train
# + papermill={"duration": 0.3471, "end_time": "2020-11-16T16:03:39.714810", "exception": false, "start_time": "2020-11-16T16:03:39.367710", "status": "completed"} tags=[]
# use nonscored target in the given file as feature
# if comment out below, use predicted nonscored target
# train = train.drop(nonscored_target, axis=1)
# train = train.merge(train_targets_nonscored, on="sig_id")
# train = train_features.merge(train_targets_scored, on='sig_id')
train = train.merge(train_targets_scored, on='sig_id')
# train = train[train['cp_type']!='ctl_vehicle'].reset_index(drop=True)
# test = test[test['cp_type']!='ctl_vehicle'].reset_index(drop=True)
# target = train[train_targets_scored.columns]
target = train[train_targets_scored.columns]
# + papermill={"duration": 0.15914, "end_time": "2020-11-16T16:03:40.023029", "exception": false, "start_time": "2020-11-16T16:03:39.863889", "status": "completed"} tags=[]
# train["cp_time"] = train_features[train_features["cp_type"]=="trt_cp"].reset_index(drop=True)["cp_time"]
# train["cp_dose"] = train_features[train_features["cp_type"]=="trt_cp"].reset_index(drop=True)["cp_dose"]
# test["cp_time"] = test_features[test_features["cp_type"]=="trt_cp"].reset_index(drop=True)["cp_time"]
# test["cp_dose"] = test_features[test_features["cp_type"]=="trt_cp"].reset_index(drop=True)["cp_dose"]
# + papermill={"duration": 2.357458, "end_time": "2020-11-16T16:03:42.542418", "exception": false, "start_time": "2020-11-16T16:03:40.184960", "status": "completed"} tags=[]
from sklearn.preprocessing import QuantileTransformer
scored_target_pred = [c + "_pred" for c in train_targets_scored.columns if c != 'sig_id']
for col in (scored_target_pred):
# transformer = QuantileTransformer(n_quantiles=100, random_state=0, output_distribution="normal")
vec_len = len(train[col].values)
vec_len_test = len(test[col].values)
raw_vec = train[col].values.reshape(vec_len, 1)
# transformer.fit(raw_vec)
if IS_TRAIN:
transformer = QuantileTransformer(n_quantiles=100, random_state=0, output_distribution="normal")
transformer.fit(raw_vec)
pd.to_pickle(transformer, f"{MODEL_DIR}/{NB}_{col}_quantile_scored.pkl")
else:
transformer = pd.read_pickle(f"{MODEL_DIR}/{NB}_{col}_quantile_scored.pkl")
train[col] = transformer.transform(raw_vec).reshape(1, vec_len)[0]
test[col] = transformer.transform(test[col].values.reshape(vec_len_test, 1)).reshape(1, vec_len_test)[0]
# + papermill={"duration": 0.161805, "end_time": "2020-11-16T16:03:42.855775", "exception": false, "start_time": "2020-11-16T16:03:42.693970", "status": "completed"} tags=[]
# train = train.drop('cp_type', axis=1)
# test = test.drop('cp_type', axis=1)
# + papermill={"duration": 0.182302, "end_time": "2020-11-16T16:03:43.192663", "exception": false, "start_time": "2020-11-16T16:03:43.010361", "status": "completed"} tags=[]
target_cols = target.drop('sig_id', axis=1).columns.values.tolist()
# + papermill={"duration": 0.200993, "end_time": "2020-11-16T16:03:43.795699", "exception": false, "start_time": "2020-11-16T16:03:43.594706", "status": "completed"} tags=[]
train
# + papermill={"duration": 2.132589, "end_time": "2020-11-16T16:03:46.088915", "exception": false, "start_time": "2020-11-16T16:03:43.956326", "status": "completed"} tags=[]
folds = train.copy()
mskf = MultilabelStratifiedKFold(n_splits=NFOLDS)
for f, (t_idx, v_idx) in enumerate(mskf.split(X=train, y=target)):
folds.loc[v_idx, 'kfold'] = int(f)
folds['kfold'] = folds['kfold'].astype(int)
folds
# + papermill={"duration": 0.180432, "end_time": "2020-11-16T16:03:46.435453", "exception": false, "start_time": "2020-11-16T16:03:46.255021", "status": "completed"} tags=[]
print(train.shape)
print(folds.shape)
print(test.shape)
print(target.shape)
print(sample_submission.shape)
# + papermill={"duration": 0.194062, "end_time": "2020-11-16T16:03:46.778434", "exception": false, "start_time": "2020-11-16T16:03:46.584372", "status": "completed"} tags=[]
folds
# + papermill={"duration": 0.167835, "end_time": "2020-11-16T16:03:47.108622", "exception": false, "start_time": "2020-11-16T16:03:46.940787", "status": "completed"} tags=[]
def process_data(data):
# data = pd.get_dummies(data, columns=['cp_time','cp_dose'])
# data.loc[:, 'cp_time'] = data.loc[:, 'cp_time'].map({24: 0, 48: 1, 72: 2, 0:0, 1:1, 2:2})
# data.loc[:, 'cp_dose'] = data.loc[:, 'cp_dose'].map({'D1': 0, 'D2': 1, 0:0, 1:1})
# --------------------- Normalize ---------------------
# for col in GENES:
# data[col] = (data[col]-np.mean(data[col])) / (np.std(data[col]))
# for col in CELLS:
# data[col] = (data[col]-np.mean(data[col])) / (np.std(data[col]))
#--------------------- Removing Skewness ---------------------
# for col in GENES + CELLS:
# if(abs(data[col].skew()) > 0.75):
# if(data[col].skew() < 0): # neg-skewness
# data[col] = data[col].max() - data[col] + 1
# data[col] = np.sqrt(data[col])
# else:
# data[col] = np.sqrt(data[col])
return data
# + papermill={"duration": 0.16363, "end_time": "2020-11-16T16:03:47.425632", "exception": false, "start_time": "2020-11-16T16:03:47.262002", "status": "completed"} tags=[]
feature_cols = [c for c in folds.columns if c not in target_cols]
feature_cols = [c for c in feature_cols if c not in ['kfold','sig_id']]
len(feature_cols)
# + papermill={"duration": 0.163302, "end_time": "2020-11-16T16:03:47.737838", "exception": false, "start_time": "2020-11-16T16:03:47.574536", "status": "completed"} tags=[]
feature_cols
# + papermill={"duration": 0.206348, "end_time": "2020-11-16T16:03:48.096266", "exception": false, "start_time": "2020-11-16T16:03:47.889918", "status": "completed"} tags=[]
folds
# + papermill={"duration": 0.183773, "end_time": "2020-11-16T16:03:48.450156", "exception": false, "start_time": "2020-11-16T16:03:48.266383", "status": "completed"} tags=[]
EPOCHS = 25
num_features=len(feature_cols)
num_targets=len(target_cols)
hidden_size=1024
# hidden_size=4096
# hidden_size=9192
# + papermill={"duration": 0.202728, "end_time": "2020-11-16T16:03:48.820303", "exception": false, "start_time": "2020-11-16T16:03:48.617575", "status": "completed"} tags=[]
def run_training(fold, seed):
seed_everything(seed)
train = process_data(folds)
test_ = process_data(test)
trn_idx = train[train['kfold'] != fold].index
val_idx = train[train['kfold'] == fold].index
train_df = train[train['kfold'] != fold].reset_index(drop=True)
valid_df = train[train['kfold'] == fold].reset_index(drop=True)
x_train, y_train = train_df[feature_cols].values, train_df[target_cols].values
x_valid, y_valid = valid_df[feature_cols].values, valid_df[target_cols].values
train_dataset = MoADataset(x_train, y_train)
valid_dataset = MoADataset(x_valid, y_valid)
trainloader = torch.utils.data.DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True)
validloader = torch.utils.data.DataLoader(valid_dataset, batch_size=BATCH_SIZE, shuffle=False)
model = Model(
num_features=num_features,
num_targets=num_targets,
hidden_size=hidden_size,
)
model.to(DEVICE)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE, weight_decay=WEIGHT_DECAY)
# scheduler = optim.lr_scheduler.OneCycleLR(optimizer=optimizer, pct_start=0.3, div_factor=1000,
# max_lr=1e-2, epochs=EPOCHS, steps_per_epoch=len(trainloader))
scheduler = optim.lr_scheduler.OneCycleLR(optimizer=optimizer, pct_start=0.2, div_factor=1e3,
max_lr=1e-2, epochs=EPOCHS, steps_per_epoch=len(trainloader))
loss_fn = nn.BCEWithLogitsLoss()
early_stopping_steps = EARLY_STOPPING_STEPS
early_step = 0
oof = np.zeros((len(train), target.iloc[:, 1:].shape[1]))
best_loss = np.inf
best_loss_epoch = -1
if IS_TRAIN:
for epoch in range(EPOCHS):
train_loss = train_fn(model, optimizer, scheduler, loss_fn, trainloader, DEVICE)
valid_loss, valid_preds = valid_fn(model, loss_fn, validloader, DEVICE)
if valid_loss < best_loss:
best_loss = valid_loss
best_loss_epoch = epoch
oof[val_idx] = valid_preds
torch.save(model.state_dict(), f"{MODEL_DIR}/{NB}-scored2-SEED{seed}-FOLD{fold}_.pth")
elif(EARLY_STOP == True):
early_step += 1
if (early_step >= early_stopping_steps):
break
if epoch % 10 == 0 or epoch == EPOCHS-1:
print(f"seed: {seed}, FOLD: {fold}, EPOCH: {epoch}, train_loss: {train_loss:.6f}, valid_loss: {valid_loss:.6f}, best_loss: {best_loss:.6f}, best_loss_epoch: {best_loss_epoch}")
#--------------------- PREDICTION---------------------
x_test = test_[feature_cols].values
testdataset = TestDataset(x_test)
testloader = torch.utils.data.DataLoader(testdataset, batch_size=BATCH_SIZE, shuffle=False)
model = Model(
num_features=num_features,
num_targets=num_targets,
hidden_size=hidden_size,
)
model.load_state_dict(torch.load(f"{MODEL_DIR}/{NB}-scored2-SEED{seed}-FOLD{fold}_.pth"))
model.to(DEVICE)
if not IS_TRAIN:
valid_loss, valid_preds = valid_fn(model, loss_fn, validloader, DEVICE)
oof[val_idx] = valid_preds
predictions = np.zeros((len(test_), target.iloc[:, 1:].shape[1]))
predictions = inference_fn(model, testloader, DEVICE)
return oof, predictions
# + papermill={"duration": 0.18549, "end_time": "2020-11-16T16:03:49.171044", "exception": false, "start_time": "2020-11-16T16:03:48.985554", "status": "completed"} tags=[]
def run_k_fold(NFOLDS, seed):
oof = np.zeros((len(train), len(target_cols)))
predictions = np.zeros((len(test), len(target_cols)))
for fold in range(NFOLDS):
oof_, pred_ = run_training(fold, seed)
predictions += pred_ / NFOLDS
oof += oof_
return oof, predictions
# + papermill={"duration": 548.772004, "end_time": "2020-11-16T16:12:58.140578", "exception": false, "start_time": "2020-11-16T16:03:49.368574", "status": "completed"} tags=[]
SEED = range(NSEEDS) # [0, 1, 2, 3 ,4]#, 5, 6, 7, 8, 9, 10]
oof = np.zeros((len(train), len(target_cols)))
predictions = np.zeros((len(test), len(target_cols)))
time_start = time.time()
for seed in SEED:
oof_, predictions_ = run_k_fold(NFOLDS, seed)
oof += oof_ / len(SEED)
predictions += predictions_ / len(SEED)
print(f"elapsed time: {time.time() - time_start}")
train[target_cols] = oof
test[target_cols] = predictions
# + papermill={"duration": 0.398196, "end_time": "2020-11-16T16:12:58.722619", "exception": false, "start_time": "2020-11-16T16:12:58.324423", "status": "completed"} tags=[]
train.to_pickle(f"{INT_DIR}/{NB}-train-score-stack-pred.pkl")
test.to_pickle(f"{INT_DIR}/{NB}-test-score-stack-pred.pkl")
# + papermill={"duration": 1.563964, "end_time": "2020-11-16T16:13:00.465440", "exception": false, "start_time": "2020-11-16T16:12:58.901476", "status": "completed"} tags=[]
train[target_cols] = np.maximum(PMIN, np.minimum(PMAX, train[target_cols]))
valid_results = train_targets_scored.drop(columns=target_cols).merge(train[['sig_id']+target_cols], on='sig_id', how='left').fillna(0)
y_true = train_targets_scored[target_cols].values
y_true = y_true > 0.5
y_pred = valid_results[target_cols].values
y_pred = np.minimum(SMAX, np.maximum(SMIN, y_pred))
score = 0
for i in range(len(target_cols)):
score_ = log_loss(y_true[:, i], y_pred[:, i])
score += score_ / target.shape[1]
print("CV log_loss: ", score)
# + papermill={"duration": 2.416383, "end_time": "2020-11-16T16:13:03.065859", "exception": false, "start_time": "2020-11-16T16:13:00.649476", "status": "completed"} tags=[]
# for c in test.columns:
# if c != "sig_id":
# test[c] = np.maximum(PMIN, np.minimum(PMAX, test[c]))
sub = sample_submission.drop(columns=target_cols).merge(test[['sig_id']+target_cols], on='sig_id', how='left').fillna(0)
sub.to_csv('submission_kibuna_nn.csv', index=False)
# + papermill={"duration": 0.251153, "end_time": "2020-11-16T16:13:03.594013", "exception": false, "start_time": "2020-11-16T16:13:03.342860", "status": "completed"} tags=[]
sub
# + papermill={"duration": 0.197781, "end_time": "2020-11-16T16:13:03.999143", "exception": false, "start_time": "2020-11-16T16:13:03.801362", "status": "completed"} tags=[]
# + papermill={"duration": 0.185191, "end_time": "2020-11-16T16:13:04.374176", "exception": false, "start_time": "2020-11-16T16:13:04.188985", "status": "completed"} tags=[]
# + papermill={"duration": 0.185531, "end_time": "2020-11-16T16:13:04.746246", "exception": false, "start_time": "2020-11-16T16:13:04.560715", "status": "completed"} tags=[]
# + papermill={"duration": 0.186445, "end_time": "2020-11-16T16:13:05.115683", "exception": false, "start_time": "2020-11-16T16:13:04.929238", "status": "completed"} tags=[]
| models/nn model with non scored[old cv].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# +
# #!/usr/bin/env python
# coding: utf-8
# In[1]:
import os
import pytz
from datetime import datetime
import boto3
import sagemaker
import sagemaker.session
from sagemaker.transformer import Transformer
from sagemaker.estimator import Estimator
from sagemaker.inputs import TrainingInput,TransformInput,CreateModelInput
from sagemaker.processing import (
ProcessingInput,
ProcessingOutput,
ScriptProcessor,
)
from sagemaker.network import NetworkConfig
from sagemaker.sklearn.processing import SKLearnProcessor
from sagemaker.workflow.conditions import (
ConditionGreaterThanOrEqualTo,
)
from sagemaker.workflow.condition_step import (
ConditionStep,
JsonGet,
)
from sagemaker.model_metrics import (
MetricsSource,
ModelMetrics,
)
from sagemaker.workflow.parameters import (
ParameterInteger,
ParameterString,
)
from sagemaker.workflow.pipeline import Pipeline
from sagemaker.workflow.properties import PropertyFile
from sagemaker.workflow.steps import (
ProcessingStep,
TrainingStep,
TransformStep,
CreateModelStep
)
from sagemaker.workflow.step_collections import RegisterModel
from sagemaker.model import Model
sess = sagemaker.Session()
# -
def get_session(region, default_bucket):
"""Gets the sagemaker session based on the region.
Args:
region: the aws region to start the session
default_bucket: the bucket to use for storing the artifacts
Returns:
`sagemaker.session.Session instance
"""
boto_session = boto3.Session(region_name=region)
sagemaker_client = boto_session.client("sagemaker")
runtime_client = boto_session.client("sagemaker-runtime")
return sagemaker.session.Session(
boto_session=boto_session,
sagemaker_client=sagemaker_client,
sagemaker_runtime_client=runtime_client,
default_bucket=default_bucket,
)
default_bucket = sess.default_bucket()
account_id = boto3.client("sts").get_caller_identity().get("Account")
region = boto3.session.Session().region_name
input_path = f"s3://{default_bucket}/mobile_price_pred/input/"
# +
import pandas as pd
df = pd.read_csv("data/data.csv")
df.to_csv(input_path)
# -
# ## Pipeline for Preprocess Job + Training Job
def get_pipeline(
region,
#subnets, #if your orginization concern about security
#security_group_ids,
role=None,
default_bucket=None,
model_package_group_name="MobilePricePred-ModelPackageGroup", # Choose any name
pipeline_name="MobilePricePred-pipeline-", # You can find your pipeline name in the Studio UI (project -> Pipelines -> name)
base_job_prefix="MobilePricePred", # Choose any name
env='dev' #Choose the enviornment
):
"""Gets a SageMaker ML Pipeline instance working with on CustomerChurn data.
Args:
region: AWS region to create and run the pipeline.
role: IAM role to create and run steps and pipeline.
default_bucket: the bucket to use for storing the artifacts
Returns:
an instance of a pipeline
"""
srilanka_tz = pytz.timezone('Asia/Colombo')
s3 = boto3.client('s3')
#working input data path
input_data = "s3://{default_bucket}/mobile_price_pred/input/"
directory_name = "mobile-price-pred"
date_folder = datetime.now(srilanka_tz).strftime("%Y-%m-%d")
sagemaker_session = get_session(region, default_bucket)
if role is None:
role = sagemaker.session.get_execution_role(sagemaker_session)
account_id = boto3.client("sts").get_caller_identity().get("Account")
region = boto3.session.Session().region_name
# Parameters for pipeline execution
processing_instance_count = ParameterInteger(
name="ProcessingInstanceCount", default_value=1
)
processing_instance_type = ParameterString(
name="ProcessingInstanceType", default_value="ml.m5.2xlarge"
)
preprocess_output_data = f"s3://{default_bucket}/mobile_price_pred/preprocess_output_data/{date_folder}/X_train/"
preprocess_output_data1 = f"s3://{default_bucket}/mobile_price_pred/preprocess_output_data/{date_folder}/X_test/"
preprocess_output_data2 = f"s3://{default_bucket}/mobile_price_pred/preprocess_output_data/{date_folder}/y_train/"
preprocess_output_data3 = f"s3://{default_bucket}/mobile_price_pred/preprocess_output_data/{date_folder}/y_test/"
training_instance_type = ParameterString(
name="TrainingInstanceType", default_value="ml.m5.xlarge"
)
model_path = ParameterString(
name="ModelPath",
default_value=f"s3://{default_bucket}/mobile_price_pred/preprocess_output_data/{date_folder}/model/xgboost/",
)
# --------------------- PREPROCESSING --------------------------------------------------------------------
ecr_repository = "mobile-price-pred-preprocessing-image"
tag = ":latest"
uri_suffix = "amazonaws.com"
preprocessing_repository_uri = "{}.dkr.ecr.{}.{}/{}".format(
account_id, region, uri_suffix, ecr_repository + tag
)
script_processor = ScriptProcessor(
command = ["python3"],
image_uri = preprocessing_repository_uri,
role = role,
instance_count = 1,
instance_type = processing_instance_type,
#network_config = NetworkConfig(subnets=subnets.split(':'), security_group_ids=security_group_ids.split(':'))
)
step_preprocess = ProcessingStep(
name="mobile-price-pred-preprocessing",
processor=script_processor,
code="mobile_price_preprocessing/preprocessing.py",
inputs=[ProcessingInput(source=input_data, destination="/opt/ml/processing/input")],
outputs=[
ProcessingOutput(output_name="trainX", destination=preprocess_output_data, source="/opt/ml/processing/train-x"),
ProcessingOutput(output_name="testX", destination=preprocess_output_data1, source="/opt/ml/processing/test-x"),
ProcessingOutput(output_name="trainy", destination=preprocess_output_data2, source="/opt/ml/processing/train-y"),
ProcessingOutput(output_name="testy", destination=preprocess_output_data3, source="/opt/ml/processing/test-y"),
]
#job_arguments=["--env", env],
)
# --------------------- TRAINING --------------------------------------------------------------------
# Training step for generating model artifacts
ecr_repository = "mobile-price-pred-training-image"
tag = ":latest"
uri_suffix = "amazonaws.com"
recommender_image_uri = "{}.dkr.ecr.{}.{}/{}".format(
account_id, region, uri_suffix, ecr_repository + tag
)
estimator = Estimator(image_uri=recommender_image_uri,
role=role,
sagemaker_session=sess, # Technical object
output_path=model_path,
base_job_name='Mobile-price-pred-training-job',
input_mode='File', # Copy the dataset and then train
train_instance_count=1,
train_instance_type=training_instance_type,
debugger_hook_config=False,
disable_profiler = True,
metric_definitions=[
# Only 40 Metrics can be accomodated
{'Name': 'Training Accuracy:' , 'Regex': 'Training Accuracy: ([-+]?[0-9]*\.?[0-9]+)'},
{'Name': 'MAE' , 'Regex': 'MAE: ([-+]?[0-9]*\.?[0-9]+)'},
{'Name': 'MSE' , 'Regex': 'MSE: ([-+]?[0-9]*\.?[0-9]+)'},
{'Name': 'RMSE' , 'Regex': 'RMSE: ([-+]?[0-9]*\.?[0-9]+)'},
#{'Name': 'F1 score:' , 'Regex': 'F1 score:([-+]?[0-9]*\.?[0-9]+)'}
],
#subnets = subnets.split(':'),
#security_group_ids = security_group_ids.split(':')
)
# start training
step_train = TrainingStep(
name="Mobile-price-pred-training",
estimator=estimator,
inputs = {
"train-x": TrainingInput(
s3_data=step_preprocess.properties.ProcessingOutputConfig.Outputs["trainX"].S3Output.S3Uri,
content_type="text/csv",
),
"train-y": TrainingInput(
s3_data=step_preprocess.properties.ProcessingOutputConfig.Outputs["trainy"].S3Output.S3Uri,
content_type="text/csv",
),
"test-x": TrainingInput(
s3_data=step_preprocess.properties.ProcessingOutputConfig.Outputs["testX"].S3Output.S3Uri,
content_type="text/csv",
),
"test-y": TrainingInput(
s3_data=step_preprocess.properties.ProcessingOutputConfig.Outputs["testy"].S3Output.S3Uri,
content_type="text/csv",
),
}
)
# ========================================= PIPELINE ORCHESTRATION ================================================
# Pipeline instance
pipeline = Pipeline(
name=pipeline_name+env,
parameters=[
processing_instance_type,
processing_instance_count,
training_instance_type,
model_path,
],
steps=[
step_preprocess,
step_train,
],
sagemaker_session=sagemaker_session,
)
return pipeline
# ## Execute the Pipeline
# +
region = region
role = sagemaker.get_execution_role()
#dev_subnets = dev_subnets
#dev_sg = dev_sg
model_package_group_name="MobilePricePred-ModelPackageGroup"
pipeline_name="MobilePricePred-pipeline-"
default_bucket = sagemaker.session.Session().default_bucket()
default_bucket='pipeline-sagemaker-test'
pipeline_def = get_pipeline(
region=region,
#subnets = dev_subnets,
#security_group_ids = dev_sg,
role=role,
default_bucket=default_bucket,
model_package_group_name=model_package_group_name,
pipeline_name=pipeline_name,
env='dev'
)
# -
pipeline_def.upsert(role_arn=role)
execution = pipeline_def.start()
# ## Clean Up
# +
import boto3
client = boto3.client('sagemaker')
response = client.delete_pipeline(
PipelineName='mobilepricepred-pipeline-dev',
ClientRequestToken='<PASSWORD>'
)
| pipeline_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import scipy
print('scipy: %s' % scipy.__version__)
# +
# %matplotlib inline
import os
#load the data
import s3fs
import numpy as np
import pandas as pd
from pandas import set_option
from matplotlib import pyplot as plt
from pandas import read_csv
from pandas.plotting import scatter_matrix
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.svm import SVR
from sklearn.tree import DecisionTreeRegressor
from sklearn.neighbors import KNeighborsRegressor
from sklearn.linear_model import Lasso
from sklearn.linear_model import ElasticNet
from sklearn.ensemble import BaggingClassifier
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn import preprocessing
import xgboost as xgb
from yellowbrick.classifier import ROCAUC
import numpy as np
from sklearn import model_selection
from sklearn.linear_model import LogisticRegression
from sklearn import datasets
from sklearn.metrics import accuracy_score
# -
model_df_encode = pd.read_csv('model_df_encoded')
from pandas import set_option
set_option('display.max_columns', None)
model_df_encode.head(10)
# +
from sklearn.model_selection import train_test_split
features = model_df_encode.drop(columns = ['exclusion_flag']).columns
target = ['exclusion_flag']
X = model_df_encode[features]
y = model_df_encode[target]
#test size
test_size = 0.33
seed = 7
#cross-validation settings
kfold = model_selection.KFold(n_splits=10, random_state=seed)
#Model instance
model = SVC()
# -
#Evaluate model performance
scoring = 'accuracy'
results = model_selection.cross_val_score(model, X, y, cv=kfold, scoring=scoring)
print('Accuracy -val set: %.2f%% (%.2f)' % (results.mean()*100, results.std()))
#split data
X_train, X_test, y_train, y_test = model_selection.train_test_split(X, y, test_size=test_size, random_state=seed)
#fit model
model.fit(X_train, y_train)
#accuracy on test set
result = model.score(X_test, y_test)
print("Accuracy - test set: %.2f%%" % (result*100.0))
| Analysis_and_Modeling/Classification accuracy SVC model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Non-Associative Commutative Algebras for Dynamical Systems
#
# We will consider quadratic dynamical systems, and go to general quasipolynomial differential equations of the form $$\dot{x_i} = x_i\sum_{j=1}^mA_{ij}\prod_{k=1}^nx_k^{B_{jk}}$$
# In one dimension we have $$\dot{x}=ax^2+bx+c$$ or in the notation above $$\dot{x}=x(ax+b+c/x)=x\sum_{j=1}^3A_{j}x^{B_j}.$$ With $A_1=a$, $A_2=b$, $A_3=c$ and $B_1=1$, $B_2=0$, $B_3=-1$.
import numpy as np
import sympy as sp
a1,a2,a3=sp.Symbol("a1"),sp.Symbol("a2"),sp.Symbol("a3")
b1,b2,b3=sp.Symbol("b1"),sp.Symbol("b2"),sp.Symbol("b3")
A=np.array([a1,a2,a3])
B=np.array([b1,b2,b3])
M=np.outer(B,A)
print M
| NACA/.ipynb_checkpoints/NACA-quadratic-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import ujson as json
import matplotlib.pyplot as plt
import os
from sklearn.model_selection import train_test_split
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dense, Conv2D , MaxPool2D , Flatten , Dropout, BatchNormalization
import tensorflow as tf
from sklearn.metrics import classification_report
#DECLARE CONSTANTS
IMG_HEIGHT = 28
IMG_WIDTH = 28
NUM_FILES = 25000
N_EPOCHS = 315
RANDOM_SEED = 42
#GETTING ALL FILE NAMES
data_files = os.listdir('./data/')
data_files = [f'./data/{file}' for file in data_files]
# data_files.remove('./data/.DS_Store')
N_CATEGORIES = len(data_files)
print(data_files)
print(len(data_files))
# +
#POPULATING TOTAL X AND Y
all_drawings = []
all_categories = []
for idx, file in enumerate(data_files):
data = np.load(file)[:NUM_FILES]
for d in data:
reshaped_img = np.array(np.reshape(d, (-1, 28))).astype(np.float32)
all_drawings.append(reshaped_img)
all_categories.append(idx)
all_drawings = np.array(all_drawings)
all_categories = np.array(all_categories)
# +
from sklearn.model_selection import train_test_split
X_train_full, X_test, y_train_full, y_test = train_test_split(all_drawings, all_categories, test_size=0.2, random_state=RANDOM_SEED)
# -
# ## Estabelecendo uma base
# Antes de iniciar a montagem da rede convulacional, vamos utilizar um classificador simples (Random Forrest) para estabelecer um minimo de performance para nosso modelo oficial ja que caso a rede neural montada possua desempenho similar a um classificado aleatorio, nao eh possivel dizer que obtivemos qualquer sucesso.
#
# Como o classificador Random forrest recebe dados de treinamento de uma forma diferente que uma rede convulacional precisamos dividir a base de treinamento de teste.
# +
# from sklearn.ensemble import RandomForestClassifier
# from sklearn.model_selection import cross_val_score
# random_forest_X_train = np.array([x.flatten() for x in X_train_full])
# random_forrest_clf = RandomForestClassifier(random_state=RANDOM_SEED, n_jobs=-1)
# cross_val_score(random_forrest_clf, random_forest_X_train, y_train_full, cv=5, n_jobs=-1)
# +
slice_index = int(len(X_train_full)*0.8)
X_train = X_train_full[:slice_index][..., np.newaxis]
X_valid = X_train_full[slice_index:][..., np.newaxis]
y_train = y_train_full[:slice_index][..., np.newaxis]
y_valid = y_train_full[slice_index:][..., np.newaxis]
X_test = X_test[..., np.newaxis]
# -
model = keras.models.Sequential([
Conv2D(filters=8, kernel_size=3,padding="same", activation="relu", input_shape=(IMG_HEIGHT,IMG_WIDTH,1)),
MaxPool2D(pool_size=2),
Conv2D(filters=16, kernel_size=3, padding="same", activation="relu"),
MaxPool2D(pool_size=2),
Conv2D(filters=32, kernel_size=3, padding="same", activation="relu"),
MaxPool2D(pool_size=2),
Flatten(),
BatchNormalization(),
Dropout(0.5, seed=RANDOM_SEED),
Dense(units=30,activation="relu"),
BatchNormalization(),
Dropout(0.5, seed=RANDOM_SEED),
Dense(units=N_CATEGORIES, activation="softmax"),
])
model.compile(optimizer=keras.optimizers.Adam(learning_rate=1e-4), loss = keras.losses.SparseCategoricalCrossentropy(from_logits=False) , metrics = ['accuracy'])
history = model.fit(X_train,y_train,epochs = N_EPOCHS, validation_data=(X_valid, y_valid), batch_size=512)
# +
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(N_EPOCHS)
plt.figure(figsize=(15, 15))
plt.subplot(2, 2, 1)
plt.plot(epochs_range, acc, label='Training Accuracy')
plt.plot(epochs_range, val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(2, 2, 2)
plt.plot(epochs_range, loss, label='Training Loss')
plt.plot(epochs_range, val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.show()
# +
predictions = model.predict_classes(X_test)
# categories = [x.split('/')[-1].split('.')[0] for x in data_files]
print(classification_report(y_test, predictions, target_names = categories))
# -
| model/Projeto_GabrielZezze_PedroLuizV2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
data = open('day13.txt', 'r').read().split('\n')
start, buses = data[0],data[1]
start = int(start)
buses = [int(b) if b != "x" else None for b in buses.split(",")]
import math
tot = 0
minV = math.inf
for i in buses:
if i == None:
continue
else:
val = start//i
inc = abs(start - (i*val+ i))
if inc < minV:
tot = inc* i
minV = inc
print("Part 1:",tot)
# +
M = math.prod(b for b in buses if b is not None)
result = 0
M
for i, b in enumerate(buses):
if b is None:
continue
Mi = M // b
mi = pow(Mi, -1, b)
result += (-i) * Mi * mi
print("Part 2:", result % M)
| day_13/day-13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Uncomment and run this cell if you're on Colab or Kaggle
# # !git clone https://github.com/nlp-with-transformers/notebooks.git
# # %cd notebooks
# from install import *
# install_requirements(is_chapter7=True)
# -
#hide
from utils import *
setup_chapter()
#hide
# %env TOKENIZERS_PARALLELISM=false
#hide
# Suppress Haystack logging
import logging
for module in ["farm.utils", "farm.infer", "haystack.reader.farm.FARMReader",
"farm.modeling.prediction_head", "elasticsearch", "haystack.eval",
"haystack.document_store.base", "haystack.retriever.base",
"farm.data_handler.dataset"]:
module_logger = logging.getLogger(module)
module_logger.setLevel(logging.ERROR)
# # Question Answering
# <img alt="<NAME>" width="500" caption="A Google search query and corresponding answer snippet" src="images/chapter07_marie-curie.png" id="marie-curie"/>
# ## Building a Review-Based QA System
# ### The Dataset
# <img alt="Phone with Query" width="400" caption="A question about a product and the corresponding review (the answer span is underlined)" src="images/chapter07_phone.png" id="phone"/>
# +
from datasets import get_dataset_config_names
domains = get_dataset_config_names("subjqa")
domains
# +
#hide_output
from datasets import load_dataset
subjqa = load_dataset("subjqa", name="electronics")
# -
print(subjqa["train"]["answers"][1])
# +
import pandas as pd
dfs = {split: dset.to_pandas() for split, dset in subjqa.flatten().items()}
for split, df in dfs.items():
print(f"Number of questions in {split}: {df['id'].nunique()}")
# -
#hide_output
qa_cols = ["title", "question", "answers.text",
"answers.answer_start", "context"]
sample_df = dfs["train"][qa_cols].sample(2, random_state=7)
sample_df
start_idx = sample_df["answers.answer_start"].iloc[0][0]
end_idx = start_idx + len(sample_df["answers.text"].iloc[0][0])
sample_df["context"].iloc[0][start_idx:end_idx]
# +
counts = {}
question_types = ["What", "How", "Is", "Does", "Do", "Was", "Where", "Why"]
for q in question_types:
counts[q] = dfs["train"]["question"].str.startswith(q).value_counts()[True]
pd.Series(counts).sort_values().plot.barh()
plt.title("Frequency of Question Types")
plt.show()
# -
for question_type in ["How", "What", "Is"]:
for question in (
dfs["train"][dfs["train"].question.str.startswith(question_type)]
.sample(n=3, random_state=42)['question']):
print(question)
# ### Sidebar: The Stanford Question Answering Dataset
# <img alt="SQuAD SotA" width="600" caption="Progress on the SQuAD 2.0 benchmark (image from Papers with Code)" src="images/chapter07_squad-sota.png" id="squad-sota"/>
# ### End sidebar
# ### Extracting Answers from Text
# #### Span classification
# <img alt="QA Head" caption="The span classification head for QA tasks" src="images/chapter07_qa-head.png" id="qa-head"/>
# <img alt="SQuAD models" width="600" caption="A selection of extractive QA models on the Hugging Face Hub" src="images/chapter07_squad-models.png" id="squad-models"/>
# #### Tokenizing text for QA
# +
#hide_output
from transformers import AutoTokenizer
model_ckpt = "deepset/minilm-uncased-squad2"
tokenizer = AutoTokenizer.from_pretrained(model_ckpt)
# -
question = "How much music can this hold?"
context = """An MP3 is about 1 MB/minute, so about 6000 hours depending on \
file size."""
inputs = tokenizer(question, context, return_tensors="pt")
#hide_input
input_df = pd.DataFrame.from_dict(tokenizer(question, context), orient="index")
input_df
print(tokenizer.decode(inputs["input_ids"][0]))
# +
import torch
from transformers import AutoModelForQuestionAnswering
model = AutoModelForQuestionAnswering.from_pretrained(model_ckpt)
with torch.no_grad():
outputs = model(**inputs)
print(outputs)
# -
start_logits = outputs.start_logits
end_logits = outputs.end_logits
print(f"Input IDs shape: {inputs.input_ids.size()}")
print(f"Start logits shape: {start_logits.size()}")
print(f"End logits shape: {end_logits.size()}")
# +
#hide_input
#id qa-scores
#caption Predicted logits for the start and end tokens; the token with the highest score is colored in orange
# The idea for this visualisation comes from https://mccormickml.com/2020/03/10/question-answering-with-a-fine-tuned-BERT
import numpy as np
import matplotlib.pyplot as plt
s_scores = start_logits.detach().numpy().flatten()
e_scores = end_logits.detach().numpy().flatten()
tokens = tokenizer.convert_ids_to_tokens(inputs["input_ids"][0])
fig, (ax1, ax2) = plt.subplots(nrows=2, sharex=True)
colors = ["C0" if s != np.max(s_scores) else "C1" for s in s_scores]
ax1.bar(x=tokens, height=s_scores, color=colors)
ax1.set_ylabel("Start Scores")
colors = ["C0" if s != np.max(e_scores) else "C1" for s in e_scores]
ax2.bar(x=tokens, height=e_scores, color=colors)
ax2.set_ylabel("End Scores")
plt.xticks(rotation="vertical")
plt.show()
# +
import torch
start_idx = torch.argmax(start_logits)
end_idx = torch.argmax(end_logits) + 1
answer_span = inputs["input_ids"][0][start_idx:end_idx]
answer = tokenizer.decode(answer_span)
print(f"Question: {question}")
print(f"Answer: {answer}")
# +
from transformers import pipeline
pipe = pipeline("question-answering", model=model, tokenizer=tokenizer)
pipe(question=question, context=context, topk=3)
# -
pipe(question="Why is there no data?", context=context,
handle_impossible_answer=True)
# #### Dealing with long passages
# +
#hide_input
#id subjqa-dist
#caption Distribution of tokens for each question-context pair in the SubjQA training set
def compute_input_length(row):
inputs = tokenizer(row["question"], row["context"])
return len(inputs["input_ids"])
dfs["train"]["n_tokens"] = dfs["train"].apply(compute_input_length, axis=1)
fig, ax = plt.subplots()
dfs["train"]["n_tokens"].hist(bins=100, grid=False, ec="C0", ax=ax)
plt.xlabel("Number of tokens in question-context pair")
ax.axvline(x=512, ymin=0, ymax=1, linestyle="--", color="C1",
label="Maximum sequence length")
plt.legend()
plt.ylabel("Count")
plt.show()
# -
# <img alt="Sliding window" caption="How the sliding window creates multiple question-context pairs for long documents—the first bar corresponds to the question, while the second bar is the context captured in each window" src="images/chapter07_sliding-window.png" id="sliding-window"/>
example = dfs["train"].iloc[0][["question", "context"]]
tokenized_example = tokenizer(example["question"], example["context"],
return_overflowing_tokens=True, max_length=100,
stride=25)
for idx, window in enumerate(tokenized_example["input_ids"]):
print(f"Window #{idx} has {len(window)} tokens")
for window in tokenized_example["input_ids"]:
print(f"{tokenizer.decode(window)} \n")
# ### Using Haystack to Build a QA Pipeline
# <img alt="QA Architecture" caption="The retriever-reader architecture for modern QA systems" src="images/chapter07_retriever-reader.png" id="retriever-reader"/>
# #### Initializing a document store
url = """https://artifacts.elastic.co/downloads/elasticsearch/\
elasticsearch-7.9.2-linux-x86_64.tar.gz"""
# !wget -nc -q {url}
# !tar -xzf elasticsearch-7.9.2-linux-x86_64.tar.gz
# +
import os
from subprocess import Popen, PIPE, STDOUT
# Run Elasticsearch as a background process
# !chown -R daemon:daemon elasticsearch-7.9.2
es_server = Popen(args=['elasticsearch-7.9.2/bin/elasticsearch'],
stdout=PIPE, stderr=STDOUT, preexec_fn=lambda: os.setuid(1))
# Wait until Elasticsearch has started
# !sleep 30
# +
#hide
# Alternative if Docker is installed
from haystack.utils import launch_es
launch_es()
# -
# !curl -X GET "localhost:9200/?pretty"
# +
from haystack.document_store.elasticsearch import ElasticsearchDocumentStore
# Return the document embedding for later use with dense retriever
document_store = ElasticsearchDocumentStore(return_embedding=True)
# -
#hide
# It's a good idea to flush Elasticsearch with each notebook restart
if len(document_store.get_all_documents()) or len(document_store.get_all_labels()) > 0:
document_store.delete_documents("document")
document_store.delete_documents("label")
# +
for split, df in dfs.items():
# Exclude duplicate reviews
docs = [{"text": row["context"],
"meta":{"item_id": row["title"], "question_id": row["id"],
"split": split}}
for _,row in df.drop_duplicates(subset="context").iterrows()]
document_store.write_documents(docs, index="document")
print(f"Loaded {document_store.get_document_count()} documents")
# -
# #### Initializing a retriever
# +
from haystack.retriever.sparse import ElasticsearchRetriever
es_retriever = ElasticsearchRetriever(document_store=document_store)
# -
item_id = "B0074BW614"
query = "Is it good for reading?"
retrieved_docs = es_retriever.retrieve(
query=query, top_k=3, filters={"item_id":[item_id], "split":["train"]})
print(retrieved_docs[0])
# #### Initializing a reader
# +
#hide_output
from haystack.reader.farm import FARMReader
model_ckpt = "deepset/minilm-uncased-squad2"
max_seq_length, doc_stride = 384, 128
reader = FARMReader(model_name_or_path=model_ckpt, progress_bar=False,
max_seq_len=max_seq_length, doc_stride=doc_stride,
return_no_answer=True)
# -
print(reader.predict_on_texts(question=question, texts=[context], top_k=1))
# #### Putting it all together
# +
from haystack.pipeline import ExtractiveQAPipeline
pipe = ExtractiveQAPipeline(reader, es_retriever)
# +
n_answers = 3
preds = pipe.run(query=query, top_k_retriever=3, top_k_reader=n_answers,
filters={"item_id": [item_id], "split":["train"]})
print(f"Question: {preds['query']} \n")
for idx in range(n_answers):
print(f"Answer {idx+1}: {preds['answers'][idx]['answer']}")
print(f"Review snippet: ...{preds['answers'][idx]['context']}...")
print("\n\n")
# -
# ## Improving Our QA Pipeline
# ### Evaluating the Retriever
# +
from haystack.pipeline import Pipeline
from haystack.eval import EvalDocuments
class EvalRetrieverPipeline:
def __init__(self, retriever):
self.retriever = retriever
self.eval_retriever = EvalDocuments()
pipe = Pipeline()
pipe.add_node(component=self.retriever, name="ESRetriever",
inputs=["Query"])
pipe.add_node(component=self.eval_retriever, name="EvalRetriever",
inputs=["ESRetriever"])
self.pipeline = pipe
pipe = EvalRetrieverPipeline(es_retriever)
# +
from haystack import Label
labels = []
for i, row in dfs["test"].iterrows():
# Metadata used for filtering in the Retriever
meta = {"item_id": row["title"], "question_id": row["id"]}
# Populate labels for questions with answers
if len(row["answers.text"]):
for answer in row["answers.text"]:
label = Label(
question=row["question"], answer=answer, id=i, origin=row["id"],
meta=meta, is_correct_answer=True, is_correct_document=True,
no_answer=False)
labels.append(label)
# Populate labels for questions without answers
else:
label = Label(
question=row["question"], answer="", id=i, origin=row["id"],
meta=meta, is_correct_answer=True, is_correct_document=True,
no_answer=True)
labels.append(label)
# -
print(labels[0])
document_store.write_labels(labels, index="label")
print(f"""Loaded {document_store.get_label_count(index="label")} \
question-answer pairs""")
labels_agg = document_store.get_all_labels_aggregated(
index="label",
open_domain=True,
aggregate_by_meta=["item_id"]
)
print(len(labels_agg))
print(labels_agg[109])
def run_pipeline(pipeline, top_k_retriever=10, top_k_reader=4):
for l in labels_agg:
_ = pipeline.pipeline.run(
query=l.question,
top_k_retriever=top_k_retriever,
top_k_reader=top_k_reader,
top_k_eval_documents=top_k_retriever,
labels=l,
filters={"item_id": [l.meta["item_id"]], "split": ["test"]})
run_pipeline(pipe, top_k_retriever=3)
print(f"Recall@3: {pipe.eval_retriever.recall:.2f}")
# +
#hide_output
def evaluate_retriever(retriever, topk_values = [1,3,5,10,20]):
topk_results = {}
for topk in topk_values:
# Create Pipeline
p = EvalRetrieverPipeline(retriever)
# Loop over each question-answers pair in test set
run_pipeline(p, top_k_retriever=topk)
# Get metrics
topk_results[topk] = {"recall": p.eval_retriever.recall}
return pd.DataFrame.from_dict(topk_results, orient="index")
es_topk_df = evaluate_retriever(es_retriever)
# +
def plot_retriever_eval(dfs, retriever_names):
fig, ax = plt.subplots()
for df, retriever_name in zip(dfs, retriever_names):
df.plot(y="recall", ax=ax, label=retriever_name)
plt.xticks(df.index)
plt.ylabel("Top-k Recall")
plt.xlabel("k")
plt.show()
plot_retriever_eval([es_topk_df], ["BM25"])
# -
# #### Dense Passage Retrieval
# <img alt="DPR Architecture" caption="DPR's bi-encoder architecture for computing the relevance of a document and query" src="images/chapter07_dpr.png" id="dpr"/>
# +
#hide_output
from haystack.retriever.dense import DensePassageRetriever
dpr_retriever = DensePassageRetriever(document_store=document_store,
query_embedding_model="facebook/dpr-question_encoder-single-nq-base",
passage_embedding_model="facebook/dpr-ctx_encoder-single-nq-base",
embed_title=False)
# -
#hide_output
document_store.update_embeddings(retriever=dpr_retriever)
dpr_topk_df = evaluate_retriever(dpr_retriever)
plot_retriever_eval([es_topk_df, dpr_topk_df], ["BM25", "DPR"])
# ### Evaluating the Reader
# +
from farm.evaluation.squad_evaluation import compute_f1, compute_exact
pred = "about 6000 hours"
label = "6000 hours"
print(f"EM: {compute_exact(label, pred)}")
print(f"F1: {compute_f1(label, pred)}")
# -
pred = "about 6000 dollars"
print(f"EM: {compute_exact(label, pred)}")
print(f"F1: {compute_f1(label, pred)}")
# +
#hide_output
from haystack.eval import EvalAnswers
def evaluate_reader(reader):
score_keys = ['top_1_em', 'top_1_f1']
eval_reader = EvalAnswers(skip_incorrect_retrieval=False)
pipe = Pipeline()
pipe.add_node(component=reader, name="QAReader", inputs=["Query"])
pipe.add_node(component=eval_reader, name="EvalReader", inputs=["QAReader"])
for l in labels_agg:
doc = document_store.query(l.question,
filters={"question_id":[l.origin]})
_ = pipe.run(query=l.question, documents=doc, labels=l)
return {k:v for k,v in eval_reader.__dict__.items() if k in score_keys}
reader_eval = {}
reader_eval["Fine-tune on SQuAD"] = evaluate_reader(reader)
# +
def plot_reader_eval(reader_eval):
fig, ax = plt.subplots()
df = pd.DataFrame.from_dict(reader_eval)
df.plot(kind="bar", ylabel="Score", rot=0, ax=ax)
ax.set_xticklabels(["EM", "F1"])
plt.legend(loc='upper left')
plt.show()
plot_reader_eval(reader_eval)
# -
# ### Domain Adaptation
# <img alt="SQuAD Schema" caption="Visualization of the SQuAD JSON format" src="images/chapter07_squad-schema.png" id="squad-schema"/>
def create_paragraphs(df):
paragraphs = []
id2context = dict(zip(df["review_id"], df["context"]))
for review_id, review in id2context.items():
qas = []
# Filter for all question-answer pairs about a specific context
review_df = df.query(f"review_id == '{review_id}'")
id2question = dict(zip(review_df["id"], review_df["question"]))
# Build up the qas array
for qid, question in id2question.items():
# Filter for a single question ID
question_df = df.query(f"id == '{qid}'").to_dict(orient="list")
ans_start_idxs = question_df["answers.answer_start"][0].tolist()
ans_text = question_df["answers.text"][0].tolist()
# Fill answerable questions
if len(ans_start_idxs):
answers = [
{"text": text, "answer_start": answer_start}
for text, answer_start in zip(ans_text, ans_start_idxs)]
is_impossible = False
else:
answers = []
is_impossible = True
# Add question-answer pairs to qas
qas.append({"question": question, "id": qid,
"is_impossible": is_impossible, "answers": answers})
# Add context and question-answer pairs to paragraphs
paragraphs.append({"qas": qas, "context": review})
return paragraphs
#hide_output
product = dfs["train"].query("title == 'B00001P4ZH'")
create_paragraphs(product)
# ```python
# [{'qas': [{'question': 'How is the bass?',
# 'id': '2543d296da9766d8d17d040ecc781699',
# 'is_impossible': True,
# 'answers': []}],
# 'context': 'I have had Koss headphones ...',
# 'id': 'd476830bf9282e2b9033e2bb44bbb995',
# 'is_impossible': False,
# 'answers': [{'text': 'Bass is weak as expected', 'answer_start': 1302},
# {'text': 'Bass is weak as expected, even with EQ adjusted up',
# 'answer_start': 1302}]}],
# 'context': 'To anyone who hasn\'t tried all ...'},
# {'qas': [{'question': 'How is the bass?',
# 'id': '455575557886d6dfeea5aa19577e5de4',
# 'is_impossible': False,
# 'answers': [{'text': 'The only fault in the sound is the bass',
# 'answer_start': 650}]}],
# 'context': "I have had many sub-$100 headphones ..."}]
# ```
# +
import json
def convert_to_squad(dfs):
for split, df in dfs.items():
subjqa_data = {}
# Create `paragraphs` for each product ID
groups = (df.groupby("title").apply(create_paragraphs)
.to_frame(name="paragraphs").reset_index())
subjqa_data["data"] = groups.to_dict(orient="records")
# Save the result to disk
with open(f"electronics-{split}.json", "w+", encoding="utf-8") as f:
json.dump(subjqa_data, f)
convert_to_squad(dfs)
# +
#hide_output
train_filename = "electronics-train.json"
dev_filename = "electronics-validation.json"
reader.train(data_dir=".", use_gpu=True, n_epochs=1, batch_size=16,
train_filename=train_filename, dev_filename=dev_filename)
# -
reader_eval["Fine-tune on SQuAD + SubjQA"] = evaluate_reader(reader)
plot_reader_eval(reader_eval)
#hide_output
minilm_ckpt = "microsoft/MiniLM-L12-H384-uncased"
minilm_reader = FARMReader(model_name_or_path=minilm_ckpt, progress_bar=False,
max_seq_len=max_seq_length, doc_stride=doc_stride,
return_no_answer=True)
#hide_output
minilm_reader.train(data_dir=".", use_gpu=True, n_epochs=1, batch_size=16,
train_filename=train_filename, dev_filename=dev_filename)
reader_eval["Fine-tune on SubjQA"] = evaluate_reader(minilm_reader)
plot_reader_eval(reader_eval)
# ### Evaluating the Whole QA Pipeline
#hide_output
# Initialize retriever pipeline
pipe = EvalRetrieverPipeline(es_retriever)
# Add nodes for reader
eval_reader = EvalAnswers()
pipe.pipeline.add_node(component=reader, name="QAReader",
inputs=["EvalRetriever"])
pipe.pipeline.add_node(component=eval_reader, name="EvalReader",
inputs=["QAReader"])
# Evaluate!
run_pipeline(pipe)
# Extract metrics from reader
reader_eval["QA Pipeline (top-1)"] = {
k:v for k,v in eval_reader.__dict__.items()
if k in ["top_1_em", "top_1_f1"]}
#hide_input
#id reader-vs-pipeline
#caption Comparison of EM and _F_~1~ scores for the reader against the whole QA pipeline
plot_reader_eval({"Reader": reader_eval["Fine-tune on SQuAD + SubjQA"],
"QA pipeline (top-1)": reader_eval["QA Pipeline (top-1)"]})
# ## Going Beyond Extractive QA
# <img alt="RAG Architecture" width="600" caption="The RAG architecture for fine-tuning a retriever and generator end-to-end (courtesy of <NAME>)" src="images/chapter07_rag-architecture.png" id="rag"/>
# +
#hide_output
from haystack.generator.transformers import RAGenerator
generator = RAGenerator(model_name_or_path="facebook/rag-token-nq",
embed_title=False, num_beams=5)
# +
#hide_output
from haystack.pipeline import GenerativeQAPipeline
pipe = GenerativeQAPipeline(generator=generator, retriever=dpr_retriever)
# -
def generate_answers(query, top_k_generator=3):
preds = pipe.run(query=query, top_k_generator=top_k_generator,
top_k_retriever=5, filters={"item_id":["B0074BW614"]})
print(f"Question: {preds['query']} \n")
for idx in range(top_k_generator):
print(f"Answer {idx+1}: {preds['answers'][idx]['answer']}")
generate_answers(query)
generate_answers("What is the main drawback?")
# ## Conclusion
# <img alt="QA Pyramid" caption="The QA hierarchy of needs" src="images/chapter07_qa-pyramid.png" id="qa-pyramid"/>
#
| 07_question-answering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import re
import pandas as pd
import numpy as np
import unicodedata
import spacy
from nltk.corpus import stopwords
sw = set(stopwords.words('portuguese'))
# -
# ## Pre processing functions
# +
def remove_tt_username(text):
text = str(text)
no_tt_username = re.sub(r'\@\S+', '', text)
return no_tt_username
def identify_emoticons(text):
text = str(text)
text = re.sub(r'\:\-?\)+', ' cara feliz ', text)
text = re.sub(r'\:\-?[dDpP]+', ' cara feliz ', text)
text = re.sub(r'\:\-?\'?\(+', ' cara triste ', text)
text = re.sub(r'\>\:\-?\(+', ' cara brava ', text)
return text
def remove_hashtags(text):
text = str(text)
no_hashtags = re.sub(r'\#\S+', '', text)
return no_hashtags
def remove_phone(text):
text = str(text)
text = re.sub(r'(\(?(\d{2,3})\)?)?\ ?\d{4,5}\-?\ ?\d{4}', ' ', text)
return text
def remove_url(text):
text = str(text)
text = re.sub(r'https?\:\/\/\S+', ' ', text)
text = re.sub(r'www\.\S+', '', text)
text = re.sub(r'[a-zA-Z|.]+\.com(\.br)?', ' link ', text)
return text
def remove_date(text):
text = str(text)
text = re.sub(r'((\d{1,2}\/)(\d{1,2}\/?)(\d{2,4})?)', ' ', text)
text = re.sub(r'((\d{1,2}\-)(\d{1,2}\-?)(\d{2,4})?)', ' ', text)
text = re.sub(r'((\d+(\s+[deDE]+\s+)[aA-zZ|ç|Ç]+((\s+[deDE]+\s+)\d+)?))', ' ', text)
return text
def remove_hour(text):
text = str(text)
text = re.sub(r'(\d+)\:(\d+)[hH]?(\:\d+)?[hH]?[rsRS]\w?', ' ', text)
text = re.sub(r'(\d+)[hH](\d+)', ' < hora > ', text)
return text
def remove_number(text):
text = str(text)
text = re.sub(r'[0-9]', '', text)
return text
def lowercase(text):
text = str(text)
text = text.lower()
return text
def remove_oneword(text):
text = str(text)
if len(text.split()) > 1:
return text
return
def remove_stopword(text):
text = str(text)
text = [word for word in text.split() if word not in sw]
text = ' '.join(text)
return text
def remove_accent(text):
text = str(text)
text = unicodedata.normalize('NFKD', text).encode('ASCII', 'ignore').decode("utf-8")
return text
def remove_emoji(text):
text = str(text)
emoji_pattern = re.compile("["
u"\U0001F600-\U000E007F"
u"\U0001F300-\U0001F5FF"
u"\U0001F680-\U0001F6FF"
u"\U0001F1E0-\U0001F1FF"
u"\U00002702-\U000027B0"
u"\U000024C2-\U0001F251"
"]+", flags=re.UNICODE)
text = emoji_pattern.sub(r'', text)
return text
def remove_punction(text):
text = str(text)
text = re.sub(r'[!"#$%&\'()*+,-.º<>/:;=?@[/\/\]^_`{|}~]', ' ', text)
return text
def preprocessing(data):
data = pd.Series(data)
data = data.apply(remove_tt_username)
data = data.apply(remove_hashtags)
data = data.apply(identify_emoticons)
data = data.apply(remove_url)
data = data.apply(remove_phone)
data = data.apply(remove_hour)
data = data.apply(remove_date)
data = data.apply(remove_number)
data = data.apply(remove_emoji)
data = data.apply(lowercase)
data = data.apply(remove_stopword)
data = data.apply(remove_accent)
data = data.apply(remove_punction)
data = data.apply(remove_oneword)
return data
| Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 다변수 정규분포
#
# - D차원 확률밀도함수는 평균벡터 $\mu$와 공분산행렬 $\sum$ 두 개의 모수를 가짐.
#
# $N(x;\mu,\sum) = \frac{1}{(2\pi)^{D/2}|\sum|^{1/2}}exp(-\frac{1}{2}(x-\mu)^T\sum^{-1}(x-\mu))$
#
#
# - $x \in R^D$ : 확률변수벡터
# - $\mu \in R^D$ : 평균벡터
# - $\sum \in R^{D \times D}$ : 공분산행렬
#
# * 공분산행렬은 양의 정부호인 대칭행렬이기 때문에 역행렬이 항상 존재한다.
# * 공분산의 역행렬은 $\sum^{-1}$ : 정밀도 행렬이라고 한다.
#
# ----
# ### 다변수 정규분포와 고유값 분해
#
# $\sum^{-1} = V\Lambda^{-1}V^T$
#
# ---
# ### 다변수정규분포의 조건부확률분포
#
# - 다변수정규분포인 확률변수벡터 중 어떤 원소의 값이 주어지면 다른 확률변수의 조건부 확률분포는 다변수정규분포다.
#
# ---
# ### 다변수 정규분포의 주변확률분포
#
# - 다변수 정규분포의 주변확률분포는 다변수정규분포다.
#
# $p(x_1) = \int p(x_1, x_2)dx_2 = N(x_1;\mu_1,\sum_{11})$
# ### 베타분포
#
# - 표본공간은 0과 1사이의 실수 $Beta(x;a,b) 0 \leq x \leq 1$
# - 확률밀도함수
#
# $Beta(x;a,b) = \frac{\Gamma(a+b)}{\Gamma(a)\Gamma(b)}x^{a-1}(1-x)^{(b-1)}$
#
# $\Gamma(a) = \int^\infty_0 x^{a-1}e^{-x}dx$
#
#
# 1) 기대값
#
# $E[X] = \frac{a}{a+b}$
#
# 2) 최빈값
#
# $mode = \frac {a-1}{a+b-2}$
#
# 3) 분산
#
# $Var[X] = \frac{ab}{(a+b)^2(a+b+1)}$
#
# ---
# ### 감마분포
#
# - 0에서 무한대의 값을 가지는 양수 값 추정
#
# $Gam(x;a,b) = \frac{1}{\Gamma(a,b)} b^ax^{a-1}e^{-bx}$
#
# 1) 기대값
#
# $E[X] = \frac{a}{b}$
#
# 2) 최빈값
#
# $mode = \frac {a-1}{b}$
#
# 3) 분산
#
# $Var[X] = \frac{a}{b^2}$
#
#
# ---
# ### 디리클레분포
#
# - Beta 분포의 확장
# - 카테고리 분포의 모수 $\mu$
#
# - 제한조건
# 1) $0 \leq x_i \leq 1$
#
# 2) $\sum^k_{i=1}x_i = 1$
#
#
# $B(\alpha_1,\alpha_2,\cdots,\alpha_K) = \frac{\prod^K_{i=1} \Gamma(\alpha_i)}{\Gamma(\sum^K_{i=1}\alpha_i)}$
#
#
# - 베타분포와 디리클레분포의 관계
#
# < 베타분포는 K=2인 디리클레분포라고 볼 수 있다. >
#
# - $Beta(x;a,b) = \frac{1}{B(\alpha_1,\alpha_2)}\prod^2_{i=1}x_i^{\alpha_i-1}$
#
# 1) 기대값
#
# $E[X_k] = \frac{a_k}{\sum \alpha}$
#
# 2) 최빈값
#
# $mode = \frac {\alpha_k-1}{\sum\alpha - K}$
#
# 3) 분산
#
# $Var[X_k] = \frac{\alpha_k(\sum\alpha- \alpha_k)}{(\sum\alpha)^2(\sum\alpha + 1)}$
#
# ---
#
# ### 확률분포의 추정
#
# #### 확률 분포의 결정
#
# 1) 어떤 확률분포를 따르는가?
#
# 2) 확률분포의 모수 값
#
# ---
# #### 모수추정 방법론
#
# 1) 모멘트방법
#
# 2) 최대가능도 추정법
#
# 3) 베이즈 추정법
#
# ---
# 1) 모멘트 방법
#
# $\mu = E[X] \triangleq \bar x = \frac {1}{N}\sum^N_{i=1}x_i$
#
# $\sigma^2 = E[(X-\mu)^2]\triangleq\bar s^2 = \frac {1}{N-1}\sum^N_{i=1}(x_i - \bar x)^2$
# ### 베이즈 추정법
xx = np.linspace(0,1,100)
a0 , b0 = 3,2
plt.plot(xx,sp.stats.beta(a0,b0).pdf(xx), c = 'r' , ls = '--', label = 'A')
a1, b1 = 60+1, 40+1
plt.plot(xx, sp.stats.beta(a1,b1).pdf(xx), c = 'g', ls = '-', label = 'B')
plt.legend()
plt.title('bayesian estimation')
plt.grid()
plt.show()
| MATH/29_MVN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] cell_id="f42807bf-593a-4f72-874e-359af9cf4906" deepnote_cell_type="markdown" tags=[]
# # NumPy and pandas
#
# Review of NumPy and examples using pandas.
#
# [Recording of lecture from 1/12/2022](https://uci.zoom.us/rec/share/ZX9cbbE7zeJR-MRWu9Rmj1_r7IMlliQOe01P27fln1RgxddHgchdl8x6HYmFKnvU.DVx5i65Sb_b3JrC_)
# + [markdown] cell_id="178277ce-6df8-49e0-a113-f1b13535cce6" deepnote_cell_type="markdown" tags=[]
# ## Warm-up exercise
#
# 1. Define an 8x4 NumPy array A of random integers between 1 and 10 (inclusive).
#
# 2. Each row of A has four columns. Let [x,y,z,w] denote one of these rows. What is the probability that x > y?
# + cell_id="cc72be75-252f-4d36-97ed-773a02b193e9" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1642003501548 source_hash="c2602aa8" tags=[]
import numpy as np
# + cell_id="e5b412ac-d51d-449d-bbf2-e017fa209f66" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1642003544053 source_hash="b6da422c" tags=[]
rng = np.random.default_rng()
# + cell_id="850e0692-32a6-47b0-86e9-fb7fa81acee7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1642003580458 output_cleared=true source_hash="ce27750d" tags=[]
help(rng.integers)
# + cell_id="2d37c613-9e7a-461f-9e27-b6da97fd2eea" deepnote_cell_type="code" deepnote_output_heights=[155.5] deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1642006440072 source_hash="fdd9d46f" tags=[]
A = rng.integers(1,11,size=(8,4))
A
# -
# In mathematics, it doesn't make sense to ask if a vector is strictly greater than another vector. In NumPy, this comparison is done *elementwise*.
# + cell_id="d653988d-bb2e-469e-bc0f-e9c281fc83d6" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1642006444778 source_hash="8c463a8c" tags=[]
A[:,0] > A[:,1]
# -
# It's the same with equality: they are compared elementwise.
# + cell_id="09bf87b3-3e6d-4b09-90dd-3e5a3e1bb80a" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=7 execution_start=1642006446096 source_hash="138912a0" tags=[]
A[:,0] == A[:,1]
# -
# With a `list` instead of an `np.array`, then equality means "are the lists exactly the same, with the same elements in the same positions?"
# + cell_id="3b72907a-71d2-4c94-be6c-31984bb73838" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1642004515392 source_hash="1d9e7361" tags=[]
[4,2,3] == [4,2,3]
# + cell_id="bc7882ab-027d-45aa-aac4-c5d6c70e0ff5" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=16 execution_start=1642004525151 source_hash="c326d95b" tags=[]
[4,2,3] == [4,3,2]
# + cell_id="70864497-9269-40fa-aa20-785d74c61245" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1642004472353 source_hash="926cc0f3" tags=[]
np.array([1,2,3]) == np.array([4,2,3])
# + cell_id="318530c1-bdae-4724-85c9-8555fd27e8e9" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1642004606788 source_hash="61e327e" tags=[]
set([4,2,3]) == set([4,3,2,2,4,2,2])
# + cell_id="402e35f4-0e81-47e0-9660-4a7c3fff089e" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1642004451301 source_hash="13986264" tags=[]
[1,2,3] == [4,2,3]
# -
# This next cell produces an example of a Boolean array.
# + cell_id="d394be65-93a1-41d0-8860-b48ad650e9c5" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1642003903267 source_hash="8c463a8c" tags=[]
A[:,0] > A[:,1]
# -
# Counting how often `True` appears.
# + cell_id="495a7fcd-481e-4377-8de1-5130c071bb89" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1642003965109 source_hash="24323b43" tags=[]
np.count_nonzero(A[:,0] > A[:,1])
# -
# We think of each row as being one "experiment". We can find the number of rows using `len`.
# + cell_id="5410bf82-041b-4de4-b287-16f695dfaf9a" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1642004001081 source_hash="aa33360a" tags=[]
# number of experiments = number of rows
len(A)
# -
# We estimate the probability using "number of successes"/"number of experiments". It won't be accurate yet, because we are using so few experiments.
# + cell_id="4e985af4-1019-4fd9-aca4-a1c5ed621429" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1642004066064 source_hash="74181c50" tags=[]
# prob estimate using len(A) experiments
np.count_nonzero(A[:,0] > A[:,1])/len(A)
# -
# Using ten million experiments.
# + cell_id="239d747f-d300-4358-9935-9ff2ddc53c08" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=222 execution_start=1642004262270 source_hash="1b2dbd10" tags=[]
A = rng.integers(1,11,size=(10**7,4))
np.count_nonzero(A[:,0] > A[:,1])/len(A)
# -
# If we do the same thing, we should get a very similar answer, but it won't be exactly the same, since these are estimates using random experiments.
A = rng.integers(1,11,size=(10**7,4))
np.count_nonzero(A[:,0] > A[:,1])/len(A)
# + [markdown] cell_id="dd9a91bb-9dc9-4d29-81e8-d710bd1e92bc" deepnote_cell_type="markdown" tags=[]
# ## pandas
#
# Probably the most important Python library for Math 10. Essentially everything we did earlier in this notebook, we can also do in pandas. The library pandas also has a lot of extra functionality that will help us work with datasets.
# + cell_id="77361c88-ceaf-4f44-b693-f18d8cbbe5ff" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1642004725311 source_hash="9b82ee11" tags=[]
import pandas as pd
# + cell_id="6638e5e5-aa6f-4c2c-a68b-326a4dccf1fa" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=13 execution_start=1642004737111 source_hash="3b396f85" tags=[]
A = rng.integers(1,11,size=(8,4))
type(A)
# + cell_id="d10318c6-d9f2-42f7-b53f-0834b3359efb" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=14 execution_start=1642004771343 source_hash="992de289" tags=[]
A.shape
# -
# We convert this NumPy array to a pandas DataFrame. (Make sure you capitalize DataFrame correctly.)
# + cell_id="2e894679-7bde-4401-beca-d77df15b8615" deepnote_cell_type="code" deepnote_output_heights=[264] deepnote_to_be_reexecuted=false execution_millis=22 execution_start=1642004956555 source_hash="2846b409" tags=[]
df = pd.DataFrame(A)
df
# -
# The syntax for getting the zeroth column of a pandas DataFrame is a little longer than the NumPy syntax.
# + cell_id="b4237e71-4af3-4dfd-8b51-982ff5e814e1" deepnote_cell_type="code" deepnote_output_heights=[174.6875] deepnote_to_be_reexecuted=false execution_millis=906 execution_start=1642005106609 source_hash="c47ecc93" tags=[]
# zeroth column of df
df.iloc[:,0]
# -
# This column is a pandas Series.
type(df.iloc[:,0])
# We can compare the entries in these columns elementwise, just like we did using NumPy.
df.iloc[:,0] > df.iloc[:,1]
# Here is the most efficient way I know to count `True`s in a pandas Boolean Series.
# + cell_id="dbc438f9-1837-412e-9fd6-8717fa1d689f" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=14 execution_start=1642005149834 source_hash="e9cd51c" tags=[]
(df.iloc[:,0] > df.iloc[:,1]).sum()
# -
# We can again get the number of rows using `len`.
# + cell_id="b76b5b7a-6a24-4d30-9f9d-43e85476b58f" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1642005192610 source_hash="2f88c77a" tags=[]
len(df)
# -
df.shape
# Here is the probability estimate.
# Not using enough experiments
((df.iloc[:,0] > df.iloc[:,1]).sum())/len(df)
# Here we increase the number of experiments, but we forget to change `df`.
# + cell_id="6e37ad6f-62a6-439c-9003-ffb95220436c" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=236 execution_start=1642005416267 source_hash="e7411d7" tags=[]
# forgot to update df
A = rng.integers(1,11,size=(10**7,4))
((df.iloc[:,0] > df.iloc[:,1]).sum())/len(df)
# -
# Here is the correct version.
# + cell_id="25bd496a-2bc5-4253-b651-e6642c3eaee1" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1642005511172 source_hash="23f614b9" tags=[]
A = rng.integers(1,11,size=(10**7,4))
df = pd.DataFrame(A)
((df.iloc[:,0] > df.iloc[:,1]).sum())/len(df)
# + cell_id="6848806a-2619-45b5-8663-7d63ac57c436" deepnote_cell_type="code" deepnote_output_heights=[155.5] deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1642005521035 source_hash="d12c9a80" tags=[]
A = rng.integers(1,11,size=(8,4))
df = pd.DataFrame(A)
# -
A
# + cell_id="3a257743-346f-4389-a02d-07a1ce96f403" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=20 execution_start=1642005527244 source_hash="f804c160" tags=[]
df
# -
# Changing column names.
# + cell_id="2afedd95-0268-4bd6-924d-2cb19263c764" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1642005682093 source_hash="14faa496" tags=[]
df.columns = ["a","b","m","chris"]
# -
df
# There are two similar operations, `df.loc` and `df.iloc`. The operation `df.loc` refers to rows and columns by their names, whereas `df.iloc` refers to rows and columns by their index.
# + cell_id="bab08e69-b25c-4a47-a4e4-f5b1faa58f67" deepnote_cell_type="code" deepnote_output_heights=[174.6875] deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1642005733063 source_hash="43964642" tags=[]
df.loc[:,"b"]
# -
df.iloc[:,1]
# There is a common shortcut for referring to a column by its name.
# + cell_id="c4169e5d-9d52-4cad-a22d-dfc5a8919542" deepnote_cell_type="code" deepnote_output_heights=[174.6875] deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1642005832575 source_hash="1fc5282b" tags=[]
# abbreviation
df["b"]
# -
# This next command says, give me the 1st-4th rows (not including the right endpoint) in the 2nd column.
# + cell_id="e5a0987e-3a05-49a7-9cd0-d2a2f13dc0d4" deepnote_cell_type="code" deepnote_output_heights=[78.75] deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1642005926593 source_hash="2dc5eaa7" tags=[]
df.iloc[1:4,2]
# -
# Somewhat confusingly, right endpoints are included when using `loc`.
# + cell_id="aaaf2fda-6578-4313-997a-4a555703bd62" deepnote_cell_type="code" deepnote_output_heights=[97.9375] deepnote_to_be_reexecuted=false execution_millis=9 execution_start=1642005965729 source_hash="b818fd71" tags=[]
df.loc[1:4,"m"]
# -
# You can use this same sort of notation to set values.
df
# + cell_id="f6bc73b5-ce05-4ee9-8985-e0de6f8146bd" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1642006046274 source_hash="6742961c" tags=[]
df.iloc[1:4,2] = -1000
# + cell_id="b14d13c2-0fe0-41ed-bdda-6bd15dc59550" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=34 execution_start=1642006054735 source_hash="f804c160" tags=[]
df
# -
# That same sort of notation also works for NumPy arrays.
# + cell_id="7fc6abd4-031a-40ec-9a50-d5cb838c0fd8" deepnote_cell_type="code" deepnote_output_heights=[155.5] deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1642006090153 source_hash="450d0f5c" tags=[]
B = np.array(df)
B
# + cell_id="86619754-0282-47a6-b8d6-06f3ea455b6d" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1642006104306 source_hash="c69bf8b9" tags=[]
B[1:4,0] = 3
# + cell_id="7d5c4e8a-ef99-4b9c-84aa-6490f9260740" deepnote_cell_type="code" deepnote_output_heights=[155.5] deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1642006108156 source_hash="ddbccdce" tags=[]
B
# -
# You can also set multiple different values. The following says, in the 1st column (remember that we start counting at 0), set the elements from the 5th, 6th, 7th rows to be 100, 200, 300, respectively.
# + cell_id="8dede0b4-29c3-4dc8-a2f9-75c0c85c923b" deepnote_cell_type="code" deepnote_output_heights=[21.1875] deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1642006146418 source_hash="13df3535" tags=[]
B[5:,1] = [100,200,300]
# + cell_id="e8ca6f04-d9ac-49f6-9734-0c8f7b197e9d" deepnote_cell_type="code" deepnote_output_heights=[155.5] deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1642006149671 source_hash="ddbccdce" tags=[]
B
| _build/jupyter_execute/Week2/Week2-Wednesday.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Model Serving Function
# -
import mlrun
import os
# +
import numpy as np
from cloudpickle import load
class LGBMModel(mlrun.serving.V2ModelServer):
def load(self):
model_file, extra_data = self.get_model('.pkl')
self.model = load(open(model_file, 'rb'))
def predict(self, body):
try:
feats = np.asarray(body['inputs'])
result = self.model.predict(feats)
return result.tolist()
except Exception as e:
raise Exception("Failed to predict %s" % e)
# +
# nuclio: end-code
# -
# ## Deploy and Test The Function
# This demo uses a Model file from MLRUn demo data repository(by default stored in Wasabi object-store service).
models_path = mlrun.get_sample_path('models/lightgbm/SampleModel.pkl')
fn = mlrun.code_to_function('lightgbm-serving',
description="LightGBM Serving",
categories=['serving', 'ml'],
labels={'author': 'edmondg', 'framework': 'lightgbm'},
code_output='.',
image='mlrun/mlrun',
kind='serving')
fn.spec.build.commands = ['pip install lightgbm']
fn.spec.default_class = 'LGBMModel'
fn.add_model('nyc-taxi-server', model_path=models_path)
# deploy the function
fn.apply(mlrun.platforms.auto_mount())
address = fn.deploy()
# test the function
my_data = '''{"inputs":[[5.1, 3.5, 1.4, 3, 5.1, 3.5, 1.4, 0.2, 5.1, 3.5, 1.4, 0.2, 5.1, 3.5, 1.4, 0.2]]}'''
fn.invoke('/v2/models/nyc-taxi-server/predict', my_data)
| howto/converting-to-mlrun/model-serving.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from pandas import DataFrame, Series
df = pd.read_csv('ex1.csv')
df
pd.read_table('ex1.csv', sep=',')
pd.read_csv('ex2.csv', header=None)
names = ['a', 'b', 'c', 'd', 'message']
pd.read_csv('ex2.csv', names = names, index_col = 'message')
parsed = pd.read_csv('csv_minindex.csv', index_col = ['key1', 'key2'])
parsed
list(open('ex3.txt'))
result = pd.read_table('ex3.txt', sep='\s+')
result
pd.read_csv('ex4.csv', skiprows=[0, 2, 3])
result = pd.read_csv('ex5.csv')
result
pd.isnull(result)
result = pd.read_csv('ex5.csv', na_values=['NULL'])
result
sentinels = {'message' : ['foo', 'NA'],
'something' : ['two']}
pd.read_csv('ex5.csv', na_values=sentinels)
result = pd.read_csv('ex6.csv')
result
pd.read_csv('ex6.csv', nrows=5)
chunker = pd.read_csv('ex6.csv', chunksize=1000)
chunker
tot = Series([])
for piece in pd.read_csv('ex6.csv', chunksize=1000):
tot = tot.add(piece['key'].value_counts(), fill_value=0)
tot = tot.sort_values(ascending = False)
tot
data = pd.read_csv('ex5.csv')
data
data.to_csv('out.csv')
# !cat out.csv
import sys
data.to_csv(sys.stdout, sep='|')
data.to_csv(sys.stdout, na_rep='NULL')
data.to_csv(sys.stdout, index = False, header = False)
data.to_csv(sys.stdout, index = False, columns = ['a', 'b', 'c'])
dates = pd.date_range('1/1/2000', periods = 7)
dates
ts = Series(np.arange(7), index=dates)
ts
ts.to_csv('tseries.csv')
# !cat tseries.csv
Series.from_csv('tseries.csv', parse_dates=True)
import csv
f = open('ex7.csv')
reader = csv.reader(f)
for line in reader:
print(line)
lines = list(csv.reader(open('ex7.csv')))
lines
header, values = lines[0], lines[1:]
data_dict = {h : v for h, v in zip(header, zip(*values))}
data_dict
obj = """
{"name": "Wes",
"places_lived": ["United States", "Spain", "Germany"],
"pet": null,
"siblings": [{"name": "Scott", "age": 25, "pet": "Zuko"},
{"name": "Katie", "age": 33, "pet": "Cisco"}]
}
"""
import json
result = json.loads(obj)
result
asjson = json.dumps(result)
asjson
siblings = DataFrame(result['siblings'], columns = ['name', 'age'])
siblings
# +
from lxml.html import parse
from urllib2 import urlopen
parsed = parse(urlopen('http://finance.yahoo.com/q/op?s=AAPL+Options'))
doc = parsed.getroot()
doc
# -
frame = pd.read_csv('ex1.csv')
| Python_for_data_analysis/Chapter_06/Chapter_06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
#
# In this notebook we'll look at how we can interact with APIs over HTTP using the `requests` package.
#
# We will cover the following:
#
# * How to make requests (including HTTP method)
# * How to work with the response
# * What kinds of HTTP errors we might encounter
# * How to raise appropriate HTTP errors
import requests
# ## `requests`
#
# [requests](https://requests.readthedocs.io/en/master/) is a library for making HTTP requests and handling their responses. It is a de facto standard in Python as it provides a neater interface that the built-in libraries.
#
# We'll do a whirlwind tour of HTTP now, but I do recommend reading the [Mozilla Developer Network article on HTTP](https://developer.mozilla.org/en-US/docs/Web/HTTP/Overview)
#
# ### What is an HTTP request?
#
# An HTTP request is a message sent to an HTTP Server (aka web server) for a given resource (expressed as a URL).
#
# Most commonly these are the requests that your browser makes when you visit a website.
#
# For example, consider the requests made when you visit https://news.ycombinator.com
#
# 
#
# Requests **must** also specify an HTTP Method. Typically this is either `GET` or `POST`.
#
# Requests may optionally specify headers that can be useful for providing authentication or specifying expect content types.
#
# In the simplest (and original configuration) almost all requests corresponded files on a webserver that would be returned. Somebody would write an HTML file and then when the URL for that file was requested, the webserver would return the file contents.
#
# However this has evolved and now a URL (universal *resource* locator) frequently corresponds to some API method or logical action that the webserver will invoke and return the results of.
#
# In either case the webserver will send back an HTTP response.
#
# ### What is an HTTP response?
#
# A message sent in response to an HTTP request. This contains:
#
# * An HTTP status: the result of the request (did it succeed, if so how, did it fail, if so how)
# * Payload/response content/body: the actual data. In the case of a file request this returns the content of the requested file.
# * Response headers: metadata pertaining to the payload and the request
#
# 
#
# ### HTTP Status Codes
#
# Status codes are numbers that correspond to messages. We will use the notation 1xx to refer to 100 to 199.
#
# **1xx**
# Informational messages (I have never received one of these)
#
# **2xx**
# Success messages. While there are different types of success, you almost always looking for:
# 200 - OK
#
# **3xx**
# Redirect messages. These are common but are frequently handled by your client library.
# Essentially the resource you have asked for has moved and you should make a new request with the redirect data.
#
# **4xx**
# User error messages.
#
# These happen when the webserver is working as expected but something is wrong with the request. If you resend the same request you will receive the same error because the issue is with the request, not the webserver.
#
# Most frequently you might see:
# 403 - Forbidden: you are not authorized to access the URL
# 404 - File Not Found
#
# **5xx**
# Server error messages.
#
# These happen when something has gone wrong on the webserver. Typically this either due to the server encountering an error while trying to respond (programming error in the API) or a timeout or the server being unreachable.
#
# Depending on the status code, this family of errors can be handled by retrying. Sometimes the server timed out because it was too busy when you last made the request and it may succeed at a later point with the same request.
#
# [More details on MDN](https://developer.mozilla.org/en-US/docs/Web/HTTP/Status)
# ### Making some requests in Python
#
# Let's take a look at what we get when we request https://news.ycombinator.com:
hacker_news_homepage_response = requests.get('https://news.ycombinator.com')
type(hacker_news_homepage_response)
# We can inspect the status code:
hacker_news_homepage_response.status_code
hacker_news_homepage_response.reason
# We can inspect the response headers:
hacker_news_homepage_response.headers
# The headers tell us some metadata about the response: when it was received, the server we spoke to (nginx), the type of content (text/html) and a bunch of other things that you don't always have to be aware of.
#
# Let's examine the actual content.
hacker_news_homepage_response.content
# This is the raw HTML that the webserver will display.
#
# Let's try a request for a non-existent URL:
doomed_response = requests.get('https://news.ycombinator.com/foobar.html')
doomed_response.status_code
doomed_response.reason
if doomed_response:
print('doomed_response is truthy')
else:
print('doomed response is not truthy')
# An idiomatic way of raising errors:
def do_something(url):
response = requests.get(url)
if response:
# use response.content or response.json()
print('That worked!')
else:
response.raise_for_status()
do_something('https://news.ycombinator.com/foobar.html')
do_something('https://news.ycombinator.com/')
# ### Working with JSON responses
#
# Here we'll be using a public API for medicine prices in South Africa: https://medicineprices.org.za/
myprodol_search_result = requests.get('https://medicineprices.org.za/api/v2/search-lite?q=myprodol')
print(myprodol_search_result.status_code)
myprodol_search_result.headers
# So far, so good.
#
# Note that the `Content-Type` header is telling us that the data returned is `application/json` (this value is a MIME type, meaning there are well-defined constants)
#
# If we take a look at the raw content it won't be immediately useful to us:
myprodol_search_result.content
# This is a binary representation of the JSON data as a string. To work with we need to parse the string.
#
# Mercifully this is such a common task that `requests` will do this for us:
myprodol_json = myprodol_search_result.json()
myprodol_json
# Now we have a dict of values and we can apply our regular Python tricks.
myprodol_tabs = [medicine for medicine in myprodol_json if medicine['dosage_form'] == 'tablet']
myprodol_tabs
# ### Query Parameters
#
# You may have noticed that our previous example had our search string embedded in our URL in the [Query String portion](https://en.wikipedia.org/wiki/Query_string).
#
# This appeared as `?q=myprodol`
#
# `?` signals the start of the query string. It is then followed by `name=value`
#
# We could have multiple parameters (this only has any effect if the server looks for those parameters)
#
# This would look like: `?q=myprodol&max_results=5`
#
# Let's see how we can include parameters in requests *without doing any String manipulation*
improved_myprodol_results = requests.get('https://medicineprices.org.za/api/v2/search-lite',
params={'q': 'myprodol tablets'})
improved_myprodol_results.json()
# Let us examine some real life API docs and how we might use them with our new knowledge:
#
# https://developer.apple.com/documentation/appstoreconnectapi/download_sales_and_trends_reports
| notebooks/04 - Requests and HTTP APIs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 1: Explore the BBC News archive
#
# Welcome! In this assignment you will be working with a variation of the [BBC News Classification Dataset](https://www.kaggle.com/c/learn-ai-bbc/overview), which contains 2225 examples of news articles with their respective categories (labels).
#
# Let's get started!
# + colab={"base_uri": "https://localhost:8080/"} id="zrZevCPJ92HG" outputId="4ba10d10-1433-42fc-dc8e-83b297705cfe"
import csv
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
# -
# Begin by looking at the structure of the csv that contains the data:
with open("./bbc-text.csv", 'r') as csvfile:
print(f"First line (header) looks like this:\n\n{csvfile.readline()}")
print(f"Each data point looks like this:\n\n{csvfile.readline()}")
# As you can see, each data point is composed of the category of the news article followed by a comma and then the actual text of the article.
# ## Removing Stopwords
#
# One important step when working with text data is to remove the **stopwords** from it. These are the most common words in the language and they rarely provide useful information for the classification process.
#
# Complete the `remove_stopwords` below. This function should receive a string and return another string that excludes all of the stopwords provided.
# GRADED FUNCTION: remove_stopwords
def remove_stopwords(sentence):
# List of stopwords
stopwords = ["a", "about", "above", "after", "again", "against", "all", "am", "an", "and", "any", "are", "as", "at", "be", "because", "been", "before", "being", "below", "between", "both", "but", "by", "could", "did", "do", "does", "doing", "down", "during", "each", "few", "for", "from", "further", "had", "has", "have", "having", "he", "he'd", "he'll", "he's", "her", "here", "here's", "hers", "herself", "him", "himself", "his", "how", "how's", "i", "i'd", "i'll", "i'm", "i've", "if", "in", "into", "is", "it", "it's", "its", "itself", "let's", "me", "more", "most", "my", "myself", "nor", "of", "on", "once", "only", "or", "other", "ought", "our", "ours", "ourselves", "out", "over", "own", "same", "she", "she'd", "she'll", "she's", "should", "so", "some", "such", "than", "that", "that's", "the", "their", "theirs", "them", "themselves", "then", "there", "there's", "these", "they", "they'd", "they'll", "they're", "they've", "this", "those", "through", "to", "too", "under", "until", "up", "very", "was", "we", "we'd", "we'll", "we're", "we've", "were", "what", "what's", "when", "when's", "where", "where's", "which", "while", "who", "who's", "whom", "why", "why's", "with", "would", "you", "you'd", "you'll", "you're", "you've", "your", "yours", "yourself", "yourselves" ]
# Sentence converted to lowercase-only
sentence = sentence.lower()
### START CODE HERE
list_sentence = sentence.split()
copy_list_sentence = sentence.split()
for word in copy_list_sentence:
if word in stopwords:
list_sentence.remove(word)
sentence = " ".join(list_sentence)
### END CODE HERE
return sentence
# Test your function
remove_stopwords("I am about to go to the store and get any snack")
# ***Expected Output:***
# ```
# 'go store get snack'
#
# ```
# ## Reading the raw data
#
# Now you need to read the data from the csv file. To do so, complete the `parse_data_from_file` function.
#
# A couple of things to note:
# - You should omit the first line as it contains the headers and not data points.
# - There is no need to save the data points as numpy arrays, regular lists is fine.
# - To read from csv files use [`csv.reader`](https://docs.python.org/3/library/csv.html#csv.reader) by passing the appropriate arguments.
# - `csv.reader` returns an iterable that returns each row in every iteration. So the label can be accessed via row[0] and the text via row[1].
# - Use the `remove_stopwords` function in each sentence.
def parse_data_from_file(filename):
sentences = []
labels = []
with open(filename, 'r') as csvfile:
### START CODE HERE
reader = csv.reader(csvfile, delimiter=",")
for i,row in enumerate(reader):
if i == 0:
continue
labels.append(row[0])
sentences.append(remove_stopwords(row[1]))
### END CODE HERE
return sentences, labels
# +
# Test your function
sentences, labels = parse_data_from_file("./bbc-text.csv")
print(f"There are {len(sentences)} sentences in the dataset.\n")
print(f"First sentence has {len(sentences[0].split())} words (after removing stopwords).\n")
print(f"There are {len(labels)} labels in the dataset.\n")
print(f"The first 5 labels are {labels[:5]}")
# -
# ***Expected Output:***
# ```
# There are 2225 sentences in the dataset.
#
# First sentence has 436 words (after removing stopwords).
#
# There are 2225 labels in the dataset.
#
# The first 5 labels are ['tech', 'business', 'sport', 'sport', 'entertainment']
#
# ```
# ## Using the Tokenizer
#
# Now it is time to tokenize the sentences of the dataset.
#
# Complete the `fit_tokenizer` below.
#
# This function should receive the list of sentences as input and return a [Tokenizer](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/text/Tokenizer) that has been fitted to those sentences. You should also define the "Out of Vocabulary" token as `<OOV>`.
def fit_tokenizer(sentences):
### START CODE HERE
# Instantiate the Tokenizer class by passing in the oov_token argument
tokenizer = Tokenizer(oov_token="<OOV>")
# Fit on the sentences
tokenizer.fit_on_texts(sentences)
### END CODE HERE
return tokenizer
# +
tokenizer = fit_tokenizer(sentences)
word_index = tokenizer.word_index
print(f"Vocabulary contains {len(word_index)} words\n")
print("<OOV> token included in vocabulary" if "<OOV>" in word_index else "<OOV> token NOT included in vocabulary")
# -
# ***Expected Output:***
# ```
# Vocabulary contains 29714 words
#
# <OOV> token included in vocabulary
#
# ```
def get_padded_sequences(tokenizer, sentences):
### START CODE HERE
# Convert sentences to sequences
sequences = tokenizer.texts_to_sequences(sentences)
# Pad the sequences using the post padding strategy
padded_sequences = pad_sequences(sequences, padding="post")
### END CODE HERE
return padded_sequences
padded_sequences = get_padded_sequences(tokenizer, sentences)
print(f"First padded sequence looks like this: \n\n{padded_sequences[0]}\n")
print(f"Numpy array of all sequences has shape: {padded_sequences.shape}\n")
print(f"This means there are {padded_sequences.shape[0]} sequences in total and each one has a size of {padded_sequences.shape[1]}")
# ***Expected Output:***
# ```
# First padded sequence looks like this:
#
# [ 96 176 1157 ... 0 0 0]
#
# Numpy array of all sequences has shape: (2225, 2438)
#
# This means there are 2225 sequences in total and each one has a size of 2438
#
# ```
def tokenize_labels(labels):
### START CODE HERE
# Instantiate the Tokenizer class
# No need to pass additional arguments since you will be tokenizing the labels
label_tokenizer = Tokenizer()
# Fit the tokenizer to the labels
label_tokenizer.fit_on_texts(labels)
# Save the word index
label_word_index = label_tokenizer.word_index
# Save the sequences
label_sequences = label_tokenizer.texts_to_sequences(labels)
### END CODE HERE
return label_sequences, label_word_index
label_sequences, label_word_index = tokenize_labels(labels)
print(f"Vocabulary of labels looks like this {label_word_index}\n")
print(f"First ten sequences {label_sequences[:10]}\n")
# ***Expected Output:***
# ```
# Vocabulary of labels looks like this {'sport': 1, 'business': 2, 'politics': 3, 'tech': 4, 'entertainment': 5}
#
# First ten sequences [[4], [2], [1], [1], [5], [3], [3], [1], [1], [5]]
#
# ```
# + [markdown] id="6rITvNKqT-51"
# **Congratulations on finishing this week's assignment!**
#
# You have successfully implemented functions to process various text data processing ranging from pre-processing, reading from raw files and tokenizing text.
#
# **Keep it up!**
| Natural Language Processing Tensorflow/C3W1_Assignment.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.2
# language: julia
# name: julia-1.7
# ---
using Distributions
using Plots
using BSON
function transition(system::Array{Int64,1}, parameters::Dict{String,Any})::Array{Float64,2}
# initialize -
TM = zeros(2,2)
A = system[1]
B = system[2]
# get parameters -
K1 = parameters["K1"]
n1 = parameters["n1"]
K2 = parameters["K2"]
n2 = parameters["n2"]
Z1 = parameters["Z1"]
Z2 = parameters["Z2"]
# compute state factors -
p1 = (A^n1/(K1^n1+A^n1))
p2 = (B^n2/(K2^n2+B^n2))
# package -
TM[1,1] = 1 - p1
TM[1,2] = p1
TM[2,1] = p2
TM[2,2] = 1 - p2
@show (A, B, p1,p2)
# return -
return TM
end
function machine(state::Int64, system::Array{Int64,1}, parameters::Dict{String,Any})::Int64
# compute the probabilities -
T = transition(system, parameters)
# get possible transitions that can occur -
choices = round.(abs.(T[state,:]),digits=2)
# create a new categorical distribution -
d = Categorical(choices)
# what is the new state?
new_state = rand(d)
@show (choices, state, new_state)
# sample -
return new_state
end
function simulation(parameters::Dict{String,Any}, initial::Array{Int64,1}; T::Int64 = 100, 𝒫::Int64 = 100)
# build the Poisson -
λ = 13.7
d = Poisson(λ)
# S -
S = [-1 0 ; 0 1]
# set the system dimensions -
number_of_chemical_species = length(initial)
number_of_time_steps = T
number_of_sample_paths = 𝒫
# initialize: machine state -
machine_state = Array{Int64,2}(undef, number_of_time_steps, number_of_sample_paths)
# initialize: chemical state -
chemical_state = Array{Int64,3}(undef, number_of_time_steps, number_of_chemical_species, number_of_sample_paths)
# main simulation loop -
for s ∈ 1:number_of_sample_paths
# setup the chemical state -
chemical_state[1,1,s] = initial[1]
chemical_state[1,2,s] = initial[2]
# setup the machine state -
machine_state[1,s] = rand(1:2)
for t ∈ 2:number_of_time_steps
# run the machine -
machine_state[t,s] = machine(machine_state[t-1,s], chemical_state[t-1,:,s], parameters)
# the enzyme has activity if machine in state 2 -
Δ = ones(2)*rand(d)
rV = S*Δ
# if machine state == 2, then the enzyme has activity
if (machine_state[t,s] == 2 && (chemical_state[t-1,1,s] + rV[1]) >= 0)
# update the chemical state -
chemical_state[t,1,s] = chemical_state[t-1,1,s] + rV[1]
chemical_state[t,2,s] = chemical_state[t-1,2,s] + rV[2]
else
# no reaction, the chemical state stays the same -
chemical_state[t,1,s] = chemical_state[t-1,1,s]
chemical_state[t,2,s] = chemical_state[t-1,2,s]
end
end
end
# return the chemical state -
return (chemical_state, machine_state)
end
# +
# Setup parameters -
K1 = 500
n1 = 6
K2 = 500
n2 = 1
T = 137
# compute normalizing constant -
MA = range(0,stop=1000,step=1) |> collect;
Z1 = sum((MA.^n1)./(K1^n1 .+ MA.^n1));
Z2 = sum((MA.^n2)./(K2^n2 .+ MA.^n2));
parameters = Dict{String,Any}()
parameters["K1"] = K1
parameters["n1"] = n1
parameters["K2"] = K2
parameters["n2"] = n2
parameters["Z1"] = Z1
parameters["Z2"] = Z2
# how many sample paths?
N = 1000
initial = [1000,0]
(chemical_state, machine_state) = simulation(parameters,initial; T = T, 𝒫 = N);
# -
s=1000
plot(chemical_state[:, 1,s], legend=false, c=:red)
plot!(chemical_state[:,2,s], c=:black)
xlabel!("Time step index", fontsize=18)
ylabel!("Molecule number", fontsize=18)
plot(machine_state[:,88])
# +
MA = range(0,stop=1000,step=1) |> collect;
# grab the parameters -
K1 = parameters["K1"]
n1 = parameters["n1"]
K2 = parameters["K2"]
n2 = parameters["n2"]
P1 = (MA.^n1)./(K1^n1 .+ MA.^n1);
P2 = (MA.^n2)./(K2^n2 .+ MA.^n2);
plot(MA, P1, label="p₁", legend=:topleft, lw=2)
plot!(MA, P2, label="p₂", lw=2)
# -
pwd()
# +
# write simulation results to disk -
# package the simulation state -
simulation_results = Dict{String,Any}()
simulation_results["parameters"] = parameters
simulation_results["initial"] = initial
simulation_results["machine_state"] = machine_state
simulation_results["chemical_state"] = chemical_state
simulation_results["T"] = T
simulation_results["𝒫"] = N
# write simulation state to disk -
_path_to_simulation_state = joinpath(pwd(),"SIMULATION-CASE-2.bson")
bson(_path_to_simulation_state, simulation_results)
# -
| prelim_2/P2-Q1-Soln-S2022.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py37_dowhy]
# language: python
# name: conda-env-py37_dowhy-py
# ---
# +
#default_exp causal_model
# -
# # Causal Model
# > Implementing instrumental variable and backdoor criteria plus propensity and regression methods
# **Notes**
#
# * Causal Bayesian Networkx: [here](http://conference.scipy.org/proceedings/scipy2015/pdfs/mike_pacer.pdf)
#
# dowhy > causal_model.py CausalModel > causal_graph.py CausalGraph
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
#export
import dowhy as dw
from typing import List, Union
import networkx as nx
import itertools
import sympy as sp
import matplotlib.pyplot as plt
import sklearn
from sklearn import linear_model, neighbors
import numpy as np
from scipy import stats
from bcg.basics import CommonCauses, Instruments, EffectModifiers, Treatments, Outcomes, get_Xy
import pandas as pd
plt.style.use('bmh')
treatments = ['V0', 'V1']
outcome = 'Y'
common_causes = ['W0']
effect_modifiers = ['X0']
instruments = []
observed_nodes = treatments + [outcome] + instruments + effect_modifiers + common_causes
add_unobserved_confounder = True
missing_nodes_as_confounders = True
cg_ref = dw.causal_graph.CausalGraph(treatments,
[outcome], graph=None,
common_cause_names=common_causes,
instrument_names=instruments,
effect_modifier_names=effect_modifiers,
observed_node_names=observed_nodes)
cg_ref._graph.nodes(data=True)
#export
class CausalGraph:
def __init__(self, treatments:List[str], outcome:str='Y',
common_causes:List[str]=None, effect_modifiers:List[str]=None,
instruments:List[str]=None, observed_nodes:List[str]=None,
missing_nodes_as_confounders:bool=False,
add_unobserved_confounder:bool=True):
if common_causes is None: common_causes = []
if effect_modifiers is None: effect_modifiers = []
if instruments is None: instruments = []
if missing_nodes_as_confounders:
all_passed_nodes = treatments + [outcome] + \
common_causes + effect_modifiers + instruments
missing_nodes = [node for node in all_passed_nodes if node not in observed_nodes]
common_causes = list(common_causes) + missing_nodes
self.g = self.create_nx_digraph(treatments, outcome,
common_causes, instruments,
effect_modifiers, add_unobserved_confounder)
@staticmethod
def create_nx_digraph(treatments:List[str], outcome:str, common_causes:List[str],
instruments:List[str], effect_modifiers:List[str],
add_unobserved_confounder:bool=False):
g = nx.DiGraph()
g.add_edges_from([(treatment, outcome) for treatment in treatments])
g.add_edges_from([(common_cause, treatment)
for common_cause, treatment in itertools.product(common_causes, treatments)])
g.add_edges_from([(common_cause, outcome)
for common_cause in common_causes])
g.add_edges_from([(effect_modifier, outcome) for effect_modifier in effect_modifiers])
g.add_edges_from([(instrument, treatment) for instrument, treatment in itertools.product(instruments, treatments)])
nx.set_node_attributes(g, True, 'observed')
if add_unobserved_confounder:
g.add_node('U', observed=False)
g.add_edges_from([('U', treatment) for treatment in treatments])
g.add_edge('U', outcome)
return g
cg = CausalGraph(treatments=treatments, outcome=outcome, common_causes=common_causes,
effect_modifiers=effect_modifiers, observed_nodes=observed_nodes,
missing_nodes_as_confounders=missing_nodes_as_confounders,
add_unobserved_confounder=add_unobserved_confounder)
cg.g.nodes['U']['observed']
# +
#export
def show_graph(g:nx.Graph, kind:str='spectral'):
try:
layout = getattr(nx, f'{kind}_layout')(g)
except AttributeError as ae:
raise AttributeError(f'No nx.{kind}_layout found')
nx.draw(g, layout=layout, with_labels=True)
def view_graph(self, kind:str='spectral'):
show_graph(self.g, kind=kind)
CausalGraph.view_graph = view_graph
# -
cg.view_graph()
# +
#export
def get_ancestors(self, node:str, g:nx.DiGraph=None,
parents_only:bool=False):
if parents_only:
f = self.g if g is None else g
return f.predecessors(node)
return nx.ancestors(self.g if g is None else g, node)
CausalGraph.get_ancestors = get_ancestors
# -
cg.get_ancestors('V0')
#hide
assert cg_ref.get_ancestors('V0') == cg.get_ancestors('V0')
# +
#export
def cut_edges(self, edges_to_cut:List[tuple]=None):
if edges_to_cut is None: return None
g_cut = self.g.copy()
g_cut.remove_edges_from(edges_to_cut)
return g_cut
CausalGraph.cut_edges = cut_edges
# -
g_cut = cg.cut_edges([('U','Y'), ('W0', 'V1')])
show_graph(g_cut)
#hide
assert cg.cut_edges() is None
# +
#export
def get_causes(self, nodes:List[str],
edges_to_cut:List[tuple]=None):
g_cut = self.cut_edges(edges_to_cut)
causes = set()
for node in nodes:
causes.update(self.get_ancestors(node, g_cut))
return causes
CausalGraph.get_causes = get_causes
# -
#hide
assert {'X0', 'U', 'W0', 'V0', 'V1'} == cg.get_causes(['Y'])
assert {'U', 'W0'} == cg.get_causes(['V0'])
assert cg_ref.get_causes(['Y']) == cg.get_causes(['Y'])
assert cg_ref.get_causes(['V0']) == cg.get_causes(['V0'])
cg.get_causes(['V0'])
# +
#export
def get_instruments(self, treatments:List[str], outcome:str):
treatment_parents_edges = set()
treatment_parents = set()
for treatment in treatments:
parents = self.get_ancestors(treatment, parents_only=True)
treatment_parents.update(parents)
treatment_parents_edges.update([(parent, treatment)
for parent in parents])
g_cut = self.cut_edges(treatment_parents_edges)
outcome_ancestors = self.get_ancestors(outcome, g_cut)
instruments_candidates = treatment_parents.difference(outcome_ancestors)
descendants = set()
for parent in outcome_ancestors:
descendants.update(nx.descendants(g_cut, parent))
instruments = instruments_candidates.difference(descendants)
return instruments
CausalGraph.get_instruments = get_instruments
# -
cg.get_instruments(treatments, outcome)
#hide
assert set(cg_ref.get_instruments(treatments, [outcome])) == set(cg.get_instruments(treatments, outcome))
# +
#export
def get_effect_modifiers(self, treatments:List[str], outcomes:List[str]):
modifiers = set()
for outcome in outcomes:
modifiers.update(self.get_ancestors(outcome))
modifiers = modifiers.difference(treatments)
for treatment in treatments:
modifiers = modifiers.difference(self.get_ancestors(treatment))
return list(modifiers)
CausalGraph.get_effect_modifiers = get_effect_modifiers
# -
cg.get_effect_modifiers(treatments, [outcome])
#hide
assert set(cg_ref.get_effect_modifiers(treatments, [outcome])) == set(cg.get_effect_modifiers(treatments, [outcome])), f'{effect_modifiers} != {cg.get_effect_modifiers(treatments, [outcome])}'
#export
class CausalModel:
def __init__(self, treatments:List[str], outcome:str='Y',
common_causes:List[str]=None, effect_modifiers:List[str]=None,
instruments:List[str]=None, causal_graph_kwargs=None):
if not causal_graph_kwargs: causal_graph_kwargs = dict()
self.cg = CausalGraph(treatments, outcome,
common_causes=common_causes,
effect_modifiers=effect_modifiers,
instruments=instruments,
**causal_graph_kwargs)
self.treatments = treatments
self.outcome = outcome
self.common_causes = common_causes
self.effect_modifiers = effect_modifiers
self.instruments = instruments
def identify_effect(self):
pass
def estimate_effect(self):
pass
def refute_estimate(self):
pass
# +
treatments = ['V0',] # 'V1']
outcome = 'Y'
common_causes = ['W0']
effect_modifiers = ['X0']
instruments = []
observed_nodes = treatments + [outcome] + instruments + effect_modifiers
add_unobserved_confounder = True
missing_nodes_as_confounders = True
cg_kwargs = dict(
missing_nodes_as_confounders=missing_nodes_as_confounders,
add_unobserved_confounder=add_unobserved_confounder,
observed_nodes=observed_nodes
)
# -
cm = CausalModel(treatments=treatments, outcome=outcome, common_causes=common_causes,
effect_modifiers=effect_modifiers,
causal_graph_kwargs=cg_kwargs)
# +
#export
def identify_effect(self, estimand_type:str='nonparametric-ate'):
causes = {
'treatments': self.cg.get_causes(self.treatments),
'effects': self.cg.get_causes([self.outcome],
edges_to_cut=[(t, self.outcome) for t in self.treatments])
}
print(f'causes: {causes}')
common_causes = causes['treatments'].intersection(causes['effects'])
print(f'common causes: {common_causes}')
instruments = self.cg.get_instruments(self.treatments, self.outcome)
# constructing backdoor estimand
backdoor = self.construct_backdoor(self.treatments, self.outcome,
common_causes, estimand_type=estimand_type)
print('Backdoor:', backdoor)
# constructing instrumental variable estimand
instrumental_variable = None
if len(instruments) > 0:
instrumental_variable = self.construct_instrumental_variable(treatments, outcome, instruments,
estimand_type=estimand_type)
print('Instrumental variable:', instrumental_variable)
return {
'observed_common_causes': common_causes,
'backdoor': backdoor,
'instrumental_variable': instrumental_variable
}
def construct_backdoor(self, treatments:List[str], outcome:str,
common_causes:List[str],
estimand_type:str='nonparametric-ate'):
if estimand_type != 'nonparametric-ate': raise NotImplementedError
# treatment variables
sym_treatments = sp.Array([sp.Symbol(treatment) for treatment in treatments])
# outcome given common causes
expr = f'{outcome} | {",".join(common_causes)}' \
if len(common_causes) > 0 else outcome
# assigning a normal distribution to the outcome given common causes
sym_mu = sp.Symbol("mu")
sym_sigma = sp.Symbol("sigma", positive=True)
sym_outcome = sp.stats.Normal(expr, sym_mu, sym_sigma)
# expected outcome given common causes
sym_conditional_outcome = sp.stats.Expectation(sym_outcome)
# effect of treatment on outcome given common causes
sym_effect = sp.Derivative(sym_conditional_outcome, sym_treatments)
return sym_effect
def construct_instrumental_variable(self, treatments:List[str], outcome:str,
instruments:List[str], estimand_type:str='nonparametric-ate'):
if estimand_type != 'nonparametric-ate': raise NotImplementedError
sym_mu, sym_sigma = 0, 1
sym_outcome = sp.stats.Normal(outcome, sym_mu, sym_sigma)
sym_treatments = sp.Array([sp.Symbol(sp.stats.Normal(treatment, sym_mu, sym_sigma))
for treatment in treatments])
sym_instruments = sp.Array([sp.Symbol(instrument)
for instrument in instruments])
sym_effect = sp.stats.Expectation(
sp.Derivative(sym_outcome, sym_instruments)
/
sp.Derivative(sym_treatments, sym_instruments)
)
return sym_effect
CausalModel.construct_backdoor = construct_backdoor
CausalModel.construct_instrumental_variable = construct_instrumental_variable
CausalModel.identify_effect = identify_effect
# -
estimands = cm.identify_effect(); estimands
# Regression estimators based on sklearn regression classes
isinstance(linear_model.LinearRegression(), sklearn.base.RegressorMixin)
#export
class RegressionEstimator:
def __init__(self, model:sklearn.base.RegressorMixin):
assert isinstance(model, sklearn.base.RegressorMixin)
self.m = model
def fit(self, X:np.ndarray, y:np.ndarray, ix:int, ix_confounders:List[int], reset:bool=True):
if not isinstance(ix_confounders, list):
ix_confounders = list(ix_confounders)
self.ix = ix
self.ix_confounders = ix_confounders
_ix = [ix] + ix_confounders
self._ix = _ix
if reset:
self.m.fit(X[:,self._ix],y)
def estimate_effect(self, X:np.ndarray, treatment:Union[int, float], control:Union[int, float],
y:np.ndarray=None):
n, _ = X.shape
_X = X.copy()
_X[:, self.ix] = treatment
treatment_outcomes = self.m.predict(_X[:, self._ix])
_X[:, self.ix] = control
control_outcomes = self.m.predict(_X[:, self._ix])
treatment_mean = treatment_outcomes.mean()
control_mean = control_outcomes.mean()
ate = treatment_mean - control_mean
return ate
# Sanity checking on a quadratic polynomial toy dataset
# +
X = np.linspace(-1, 1, 200)
X = np.array([X**2, X, np.ones(len(X))*.5]).T
w = np.array([2, 0, .5])
y = X @ w
fig, ax = plt.subplots(figsize=(8,4), constrained_layout=True)
ax.scatter(X[:,-2], y)
ax.set(xlabel='x', ylabel='y', title='dataset')
plt.show()
# -
regression_model = linear_model.LinearRegression()
estimator = RegressionEstimator(regression_model)
estimator.fit(X, y, ix=0, ix_confounders=[1])
ate = estimator.estimate_effect(X=X, treatment=1, control=0)
print(f'ate = {ate:.3f} coefs {estimator.m.coef_}')
#hide
assert np.isclose(ate, w[0], atol=.1)
# Classification estimator
#
# propensity score: common causes -> prediction of treatment (class) -> grouping by score to select pairs of most similar treatment and control group samples to compute the difference in outcome
#
# grouping is done using some nearest neighbour search:
# - ATC if nearest neighbor is set up with the **treated** group and for each **control** group sample a match is looked up and then the difference of the outcome is computed
# - ATT if nearest neighbor is set up with the **control** group and for each **treated** group sample a match is looked up and then the difference of the outcome is computed
# **TODO**: test `PropensityScoreMatcher` on data generated using `bcg.basics` classes
# +
n = 200
x_treatment = np.random.choice([True, False], p=[.5, .5], size=n)
x_common_causes = np.array([
[np.random.normal(loc=v, scale=.1) for v in x_treatment],
[np.random.normal(loc=10-v, scale=.1) for v in x_treatment],
])
y_outcome = np.array([np.random.normal(loc=v, scale=.1) for v in x_treatment])
fig, axs = plt.subplots(figsize=(8,6), nrows=4, constrained_layout=True)
axs[0].hist(x_treatment.astype(float))
axs[0].set(xlabel='treatment')
axs[1].hist(x_common_causes[0])
axs[1].set(xlabel='cc0')
axs[2].hist(x_common_causes[1])
axs[2].set(xlabel='cc1')
axs[3].hist(y_outcome)
axs[3].set(xlabel='outcome')
plt.show()
fig, ax = plt.subplots(figsize=(8,4), constrained_layout=True)
ax.scatter(x_treatment, y_outcome)
ax.set(xlabel='treatment', ylabel='outcome', title='dataset')
plt.show()
X, y = np.concatenate((x_treatment[:,None], x_common_causes.T), axis=1), y_outcome
X.shape, y.shape
# -
class PropensityScoreMatcher:
def __init__(self, propensity_model:sklearn.base.ClassifierMixin):
assert isinstance(propensity_model, sklearn.base.ClassifierMixin)
self.pm = propensity_model
def fit(self, X:np.ndarray, y:np.ndarray, ix:int, ix_confounders:List[int], reset:bool=True):
'''building the classifier model & nearest neigbhor search thingy
ix: needs to point to a binary variable
'''
if not isinstance(ix_confounders, list):
ix_confounders = list(ix_confounders)
self.ix = ix
self.ix_confounders = ix_confounders
_ix = [ix] + ix_confounders
self._ix = _ix
if reset:
self.pm.fit(X[:, self.ix_confounders], X[:,self.ix])
def estimate_effect(self, X:np.ndarray, treatment:Union[int, bool], control:Union[int, bool],
y:np.ndarray=None, kind:str='ate'):
assert y is not None, 'Cannot be None. That\'s just the default to have consistent method parameters.'
assert kind in ['ate', 'att', 'atc']
propensity_score = self.pm.predict(X[:, self.ix_confounders])
ix_treat, ix_control = X[:,self.ix] == treatment, X[:,self.ix] == control
X_treat, X_cont = X[ix_treat,:], X[ix_control,:]
y_treat, y_cont = y[ix_treat], y[ix_control]
searcher = neighbors.NearestNeighbors(n_neighbors=1)
def get_att():
searcher.fit(propensity_score[ix_control][:,None])
distances, indices = searcher.kneighbors(propensity_score[ix_treat][:,None])
att = 0
n_treat = ix_treat.sum()
for i in range(n_treat):
out_treat = y_treat[i]
out_cont = y_cont[indices[i][0]]
att += out_treat - out_cont
return att / n_treat
def get_atc():
searcher.fit(propensity_score[ix_treat][:,None])
distances, indices = searcher.kneighbors(propensity_score[ix_control][:,None])
atc = 0
n_cont = ix_control.sum()
for i in range(n_cont):
out_treat = y_treat[indices[i][0]]
out_cont = y_cont[i]
atc += out_treat - out_cont
return atc / n_cont
def get_ate():
n_treat = ix_treat.sum()
n_cont = ix_control.sum()
att = get_att()
atc = get_atc()
return (att*n_treat + atc*n_cont) / (n_treat + n_cont)
if kind == 'ate':
return get_ate()
elif kind == 'att':
return get_att()
elif kind == 'atc':
return get_atc()
else:
raise NotImplementedError
propensity_model = linear_model.LogisticRegression(solver='lbfgs')
estimator = PropensityScoreMatcher(propensity_model)
estimator.fit(X, y, ix=0, ix_confounders=[1, 2])
ate = estimator.estimate_effect(X=X, treatment=True, control=False, y=y)
print(f'ate = {ate:.3f}')
# Generating data for the graphical model using `bcg.basics` functions
# +
outcome_is_binary = True
treatment_is_binary = True
n = 333
n_common_causes = len(common_causes)
n_instruments = len(instruments)
n_eff_mods = len(effect_modifiers)
n_treatments = len(treatments)
beta = 1 # max random value
cc = CommonCauses.get_obs(n, n_common_causes)
ins = Instruments.get_obs(n, n_instruments)
em = EffectModifiers.get_obs(n, n_eff_mods)
treat = Treatments.get_obs(n, n_treatments, cc, ins, beta, treatment_is_binary=treatment_is_binary)
out = Outcomes.get_obs(treat, cc, em, outcome_is_binary=outcome_is_binary)
obs = pd.concat((treat.obs, cc.obs, em.obs, ins.obs, out.obs), axis=1)
X, y, not_target = get_Xy(obs, target=outcome)
# -
obs.head(), obs.tail()
not_target.index('V0')
# Adding effect estimate functionality to `CausalModel`
# Changing the implementation of `get_Xy`, incorporating products with effect modifiers, based on lns 59-71 in `causal_estimators/linear_regression_estimator.py` with the new argument`feature_product_groups`. The variable is supposed to consist of two lists, each containing features in `obs`, of which all products will be computed.
#export
def get_Xy_with_products(obs:pd.DataFrame, target:str='Y', feature_product_groups:List[list]=None):
'feaure_product_groups (e.g. [["V0", "V1", "W0"], ["X0", "X1"]]) to compute products between each var in the first and second list (not within each list)'
not_target = [c for c in obs.columns if c != target and c not in feature_product_groups[1]]
# out_cols = [col for col in obs.columns if col != target ]
X, y = obs.loc[:, not_target].values, \
obs.loc[:, target].values.ravel()
if feature_product_groups:
assert isinstance(feature_product_groups, list)
assert len(feature_product_groups) == 2
assert all([isinstance(f, list) for f in feature_product_groups])
product_cols = [(t,e) for t,e in itertools.product(*feature_product_groups)]
cols = list(obs.columns.values)
for t, e in product_cols:
ix_t = cols.index(t)
ix_e = cols.index(e)
x = (obs[t] * obs[e]).values
X = np.concatenate((X, x[:,None]), axis=1)
not_target.append(f'{t}_{e}')
return X, y, not_target
get_Xy_with_products(obs, target=outcome, feature_product_groups=[treatments, effect_modifiers])
#hide
X, y, not_target = get_Xy_with_products(obs, target=outcome, feature_product_groups=[treatments, effect_modifiers])
assert y.shape[0] == X.shape[0]
assert X.shape[1] == len(not_target)
n_not_target = len(not_target)
rest_cols = [v for v in obs.columns.values if v not in effect_modifiers and v != outcome]
n_not_target_ideal = len(treatments) * len(effect_modifiers) + len(rest_cols)
assert len(not_target) == n_not_target_ideal, f'{n_not_target} != {n_not_target_ideal}: not_target = {not_target}'
# +
#export
def estimate_effect(self, estimands:dict, control_value:float,
treatment_name:str, treatment_value:float,
obs:pd.DataFrame, outcome:str='Y',
causal_method:str='backdoor',
model:Union[sklearn.base.RegressorMixin,sklearn.base.ClassifierMixin]=None, target_unit:str='ate',
effect_modifiers:List[str]=None,
supervised_type_is_regression:bool=True):
assert causal_method in {'backdoor', 'instrumental_variable'}
assert target_unit == 'ate'
print('model', model)
if model is None:
if supervised_type_is_regression:
model = linear_model.LinearRegression()
else:
model = linear_model.LogisticRegression(solver='lbfgs')
if effect_modifiers is None:
effect_modifiers = self.effect_modifiers
# decide on approach given causal_method and model_type
# estimate the effect using the arrived on approach
X, y, not_outcome = get_Xy_with_products(obs, target=outcome, feature_product_groups=[treatments, effect_modifiers])
if supervised_type_is_regression:
estimator = RegressionEstimator(model)
else:
estimator = PropensityScoreMatcher(model)
ix = [v.lower() for v in not_outcome].index(treatment_name)
confounders = self.treatments + list(estimands['observed_common_causes']) + effect_modifiers
print('confounders', confounders)
ix_confounders = [_i for _i,_v in enumerate(obs.columns.values) if _v in confounders]
estimator.fit(X, y, ix, ix_confounders)
effect = estimator.estimate_effect(X=X, treatment=treatment_value, control=control_value, y=y)
return effect
CausalModel.estimate_effect = estimate_effect
# propensity_model = linear_model.LogisticRegression(solver='lbfgs')
# estimator = PropensityScoreMatcher(propensity_model)
# estimator.fit(X, y, ix=0, ix_confounders=[1, 2])
# ate = estimator.estimate_effect(X=X, y=y, treatment=True, control=False)
# print(f'ate = {ate:.3f}')
# +
causal_method = 'backdoor'
control_value = 0
treatment_name = 'v0'
treatment_value = 2
effect_modifiers = effect_modifiers
target_unit = 'ate'
# model = linear_model.LinearRegression()
# model = linear_model.LogisticRegression()
model = None
supervised_type_is_regression = False
cm.estimate_effect(estimands, control_value, treatment_name, treatment_value,
obs, outcome=outcome, causal_method=causal_method, model=model,
target_unit=target_unit, effect_modifiers=effect_modifiers,
supervised_type_is_regression=supervised_type_is_regression)
# -
a = np.linspace(1, 4, 5)
b = a[:, np.newaxis]; b
| notebooks/02_causal_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="e8cc38f081fa109d3b8477d3beb06c809d000563"
# If you want see another interesting Kernels please check here https://www.kaggle.com/kabure/kernels
#
# ### *Please, don't forget to upvote this Kernel to keep me motivated ! *
# + [markdown] _uuid="8d2102167ab6bfebd6143032ef5e074cb94e487e"
# English is not my native language, so sorry for any error.
# + [markdown] _uuid="47748c0985b935b4e46d2a6c83af0eecf1516f65"
# # Google Analytics Customer Revenue Prediction
# + [markdown] _uuid="7fbc08fd2ec0ec81825c6d794576fbb7a06ff9c1"
# ## Presenting the initial data:
#
# <b>Data Fields: </b>
# <b>fullVisitorIdv</b> - A unique identifier for each user of the Google Merchandise Store. <br>
# <b>channelGrouping</b> - The channel via which the user came to the Store.<br>
# <b>date</b> - The date on which the user visited the Store.<br>
# <b>device </b>- The specifications for the device used to access the Store.<br>
# <b>geoNetwork</b> - This section contains information about the geography of the user.<br>
# <b>sessionId</b> - A unique identifier for this visit to the store.<br>
# <b>socialEngagementType</b> - Engagement type, either "Socially Engaged" or "Not Socially Engaged".<br>
# <b>totals</b> - This section contains aggregate values across the session.<br>
# <b>trafficSource</b> - This section contains information about the Traffic Source from which the session originated.<br>
# <b>visitId</b> - An identifier for this session. This is part of the value usually stored as the _utmb cookie. This is only unique to the user. For a completely unique ID, you should use a combination of fullVisitorId and visitId.<br>
# <b>visitNumber</b> - The session number for this user. If this is the first session, then this is set to 1.<br>
# <b>visitStartTime</b> - The timestamp (expressed as POSIX time).<br>
# + [markdown] _uuid="f8f57d502c6739ff78f74aa89519987d05641cae"
# First of all, the data are becoming in Json format, so we will need to handle with it and I will use a chunk that I saw in almost all kernel of this competition.
# + [markdown] _uuid="d5db3cb7d849d9fad6163f3f8b1e671cd6f17b93"
# # Objectives:
#
# - I will explore if we have some difference between the browser and if browser is significant to predict sells. <br>
# - Which countrys and continents have more acesses and sales ? How it's distributed?! <br>
# - Which type of device are most normal in our dataset?<br>
# - What's the mobile % of accesses? <br>
# - Which is the most frequent Operational System? <br>
# - What's the most frequent channelGrouping ?<br>
# - Whats the most frequent Weekdays, months, days, year with highest accesses and revenue?
#
# And another bunch of ideas that I will have when start exploring.
#
#
#
# + [markdown] _uuid="6ea6f1fa6570a74176e88faf94c3da6782824043"
# ## Importing necessary librarys
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# Necessary librarys
import os # it's a operational system library, to set some informations
import random # random is to generate random values
import pandas as pd # to manipulate data frames
import numpy as np # to work with matrix
from scipy.stats import kurtosis, skew # it's to explore some statistics of numerical values
import matplotlib.pyplot as plt # to graphics plot
import seaborn as sns # a good library to graphic plots
import squarify # to better understand proportion of categorys - it's a treemap layout algorithm
# Importing librarys to use on interactive graphs
from plotly.offline import init_notebook_mode, iplot, plot
import plotly.graph_objs as go
import json # to convert json in df
from pandas.io.json import json_normalize # to normalize the json file
# to set a style to all graphs
plt.style.use('fivethirtyeight')
init_notebook_mode(connected=True)
# + [markdown] _uuid="59dd8b9cf783a9a649fd59eddf9108f6341551f7"
# ## Some columns are in Json format so it will be necessary to handle with this problem.
#
# I will use a chunk code inspiration that almost all kernels are using I dont know who did first, but I got on SRK kernel and I did some modifications
# + _uuid="17c123bd94775fa116a13113e729dc5113f01aad"
columns = ['device', 'geoNetwork', 'totals', 'trafficSource'] # Columns that have json format
dir_path = "../input/" # you can change to your local
# p is a fractional number to skiprows and read just a random sample of the our dataset.
p = 0.07 # *** In this case we will use 50% of data set *** #
#Code to transform the json format columns in table
def json_read(df):
#joining the [ path + df received]
data_frame = dir_path + df
#Importing the dataset
df = pd.read_csv(data_frame,
converters={column: json.loads for column in columns}, # loading the json columns properly
dtype={'fullVisitorId': 'str'}, # transforming this column to string
skiprows=lambda i: i>0 and random.random() > p)# Number of rows that will be imported randomly
for column in columns: #loop to finally transform the columns in data frame
#It will normalize and set the json to a table
column_as_df = json_normalize(df[column])
# here will be set the name using the category and subcategory of json columns
column_as_df.columns = [f"{column}.{subcolumn}" for subcolumn in column_as_df.columns]
# after extracting the values, let drop the original columns
df = df.drop(column, axis=1).merge(column_as_df, right_index=True, left_index=True)
# Printing the shape of dataframes that was imported
print(f"Loaded {os.path.basename(data_frame)}. Shape: {df.shape}")
return df # returning the df after importing and transforming
# + [markdown] _uuid="86ee07b708307e5bfb907e41672393d374216bfe"
# # Importing the datasets
# + _uuid="5ea0bc7871409e13de182189ebc0f30ddf7f3573"
# %%time
# # %%time is used to calculate the timing of code chunk execution #
# We will import the data using the name and extension that will be concatenated with dir_path
df_train = json_read("train.csv")
# The same to test dataset
#df_test = json_read("test.csv")
# + [markdown] _uuid="4b5cd86c5d9a59ed86a7be4f3fa19b079ab5dfbb"
# Nice. <br>
# After the import and transformation, we have 54 columns. <br>
# Now, let's see our data and handle with problemns that we will find
# + _uuid="fb4b16cc25fe51887b9b4ebb3a161a0a6d683866"
# This command shows the first 5 rows of our dataset
df_train.head()
# + [markdown] _uuid="8feee60f1daee4adba7db4836a25348977a65ba9"
# It's interesting because we can see that <b>SessionId</b> has the <b>fullVisitorId</b> and <b>VisitStartTime</b> and <b>visitId</b>
#
# Also, the date column we need to transform in datetime format and extract another datetime informations contained in the columns that I quoted above
# + [markdown] _uuid="473c414cf04168f496cd66a053029b39d5d19973"
# ## Knowing the missing values
# + _uuid="fec2d2803b7bd55c5867242cb88af572b600290d"
# code chunk that I saw in Gabriel Preda kernel
def missing_values(data):
total = data.isnull().sum().sort_values(ascending = False) # getting the sum of null values and ordering
percent = (data.isnull().sum() / data.isnull().count() * 100 ).sort_values(ascending = False) #getting the percent and order of null
df = pd.concat([total, percent], axis=1, keys=['Total', 'Percent']) # Concatenating the total and percent
print("Total columns at least one Values: ")
print (df[~(df['Total'] == 0)]) # Returning values of nulls different of 0
print("\n Total of Sales % of Total: ", round((df_train[df_train['totals.transactionRevenue'] != np.nan]['totals.transactionRevenue'].count() / len(df_train['totals.transactionRevenue']) * 100),4))
return
# + _uuid="8e82a6e0f0f5e949c8142c7f7ecdc464441a0256"
# calling the missing values function
missing_values(df_train)
# + [markdown] _uuid="51ec5f59a8ae95afffd418e95f7f878f269e7c36"
# ### Nice. We can see that we have: <br>
# Our target have just 1.3% of non-null values <br>
# 6 columns with 97%+ of missing values <br>
# 4 columns with 50%+ of missing values <br>
# 1 column with 22.22% <br>
# 1 column with 0.004% <br>
#
# We will explore to understand what it looks like
#
# + [markdown] _uuid="e8d964f6e05fca00667ec2a3c2b0e62e7e7266d0"
# ## Let's take a look on datatypes of all columns
# + [markdown] _uuid="13f65c013634d3793ec70c2143c8e9cee926eb2b"
# - If you want see the code click in "code"
# - If you want see the ouput click in "output"
# + _kg_hide-output=true _uuid="fbe23d09172475042da33a2185600b7ffc5a7653"
print(df_train.info())
# + [markdown] _uuid="17e0aa2c0b6f99008990785e8f360ee49be48a85"
# ## Nice!
#
# Data Types contained in our dataframe: <br>
# - bool(1) <br>
# - int64(4) <br>
# - object(49) <br>
# + [markdown] _uuid="dc1ed41a1c9a92a3f552b42207c116adf50e21bf"
# ## Creating the function to handle with date
# + _uuid="4284e6cb9d9930d26460c4653c5935a2deeed422"
# library of datetime
from datetime import datetime
# This function is to extract date features
def date_process(df):
df["date"] = pd.to_datetime(df["date"], format="%Y%m%d") # seting the column as pandas datetime
df["_weekday"] = df['date'].dt.weekday #extracting week day
df["_day"] = df['date'].dt.day # extracting day
df["_month"] = df['date'].dt.month # extracting day
df["_year"] = df['date'].dt.year # extracting day
df['_visitHour'] = (df['visitStartTime'].apply(lambda x: str(datetime.fromtimestamp(x).hour))).astype(int)
return df #returning the df after the transformations
# + _uuid="80bc44e6661561bdb69f06eab88215cb31311abb"
### Calling the function
# + _uuid="7619fa329dcb6c0b5bea70b032145fe59a6ec70d"
df_train = date_process(df_train) #calling the function that we created above
df_train.head(n=2) #printing the first 2 rows of our dataset
# + [markdown] _uuid="6f3bb45b16bf649d463276cb50eb6e49da5b8a43"
# ## Before look the unique values in each column, I will drop the constant values that is not useful and will make the df lighter
# for it, I will need to give some attention to numerical values
# + [markdown] _uuid="82d578ffa3e569ff729f973868a51ad99812bc74"
# Defining some functions that I will use to call clean the data
# - If you want see, click in "code"
# + _uuid="ad6d4dd89351fda4bcf87f45a0236ec6c1168b29"
def FillingNaValues(df): # fillna numeric feature
df['totals.pageviews'].fillna(1, inplace=True).astype(int) #filling NA's with 1
df['totals.newVisits'].fillna(0, inplace=True).astype(int) #filling NA's with 0
df['totals.bounces'].fillna(0, inplace=True).astype(int) #filling NA's with 0
df["totals.transactionRevenue"] = df["totals.transactionRevenue"].fillna(0.0).astype(float) #filling NA with zero
df['trafficSource.isTrueDirect'].fillna(False, inplace=True) # filling boolean with False
df['trafficSource.adwordsClickInfo.isVideoAd'].fillna(True, inplace=True) # filling boolean with True
df_train.loc[df_train['geoNetwork.city'] == "(not set)", 'geoNetwork.city'] = np.nan
df_train['geoNetwork.city'].fillna("NaN", inplace=True)
return df #return the transformed dataframe
# + _uuid="18f9f89cbe2074b38e687d33ad06c63499df7d69" _kg_hide-input=true
def NumericalColumns(df): # fillna numeric feature
df['totals.pageviews'].fillna(1, inplace=True) #filling NA's with 1
df['totals.newVisits'].fillna(0, inplace=True) #filling NA's with 0
df['totals.bounces'].fillna(0, inplace=True) #filling NA's with 0
df['trafficSource.isTrueDirect'].fillna(False, inplace=True) # filling boolean with False
df['trafficSource.adwordsClickInfo.isVideoAd'].fillna(True, inplace=True) # filling boolean with True
df["totals.transactionRevenue"] = df["totals.transactionRevenue"].fillna(0.0).astype(float) #filling NA with zero
df['totals.pageviews'] = df['totals.pageviews'].astype(int) # setting numerical column as integer
df['totals.newVisits'] = df['totals.newVisits'].astype(int) # setting numerical column as integer
df['totals.bounces'] = df['totals.bounces'].astype(int) # setting numerical column as integer
df["totals.hits"] = df["totals.hits"].astype(float) # setting numerical to float
df['totals.visits'] = df['totals.visits'].astype(int) # seting as int
return df #return the transformed dataframe
# + [markdown] _uuid="f94022c26f4a61c2d3c3693254b51e7a1e57cb98"
# Normalize
# + _uuid="9ae19cdafba7f9a37c64b4d398a9f8be342a56fc" _kg_hide-input=true
from sklearn import preprocessing
def Normalizing(df):
# Use MinMaxScaler to normalize the column
df["totals.hits"] = (df['totals.hits'] - min(df['totals.hits'])) / (max(df['totals.hits']) - min(df['totals.hits']))
# normalizing the transaction Revenue
df['totals.transactionRevenue'] = df_train['totals.transactionRevenue'].apply(lambda x: np.log1p(x))
# return the modified df
return df
# + [markdown] _uuid="751aca8f7276d7ba4968664b9349d063f0b4009b"
# ### Let's investigate some constant columns
# + _uuid="42321bff6c5bdf02b01bdc6f4fec332e0696cc13"
# We will takeoff all columns where we have a unique value (constants)
# It is useful because this columns don't give us none information
discovering_consts = [col for col in df_train.columns if df_train[col].nunique() == 1]
# printing the total of columns dropped and the name of columns
print("Columns with just one value: ", len(discovering_consts), "columns")
print("Name of constant columns: \n", discovering_consts)
# + [markdown] _uuid="f58c88c804aeeca68d597f6cf0ebf61709467d84"
#
# + _uuid="622fda5f6a0b8ffb3a008e755f19b1bee77c4597"
#Here are all columns that the unique value is 'not available in demo dataset'
not_aval_cols = ['socialEngagementType','device.browserSize','device.browserVersion', 'device.flashVersion',
'device.language' ,'device.mobileDeviceBranding', 'device.mobileDeviceInfo','device.mobileDeviceMarketingName',
'device.mobileDeviceModel', 'device.mobileInputSelector' , 'device.operatingSystemVersion','device.screenColors',
'device.screenResolution', 'geoNetwork.cityId', 'geoNetwork.latitude' ,'geoNetwork.longitude',
'geoNetwork.networkLocation','trafficSource.adwordsClickInfo.criteriaParameters']
# + [markdown] _uuid="c8171bdb0370f576f7962687f81aef8cae73d386"
# # It's useul to we have notion that might we have 23 constant columns
#
# - Below I will set a function to better investigate our data and correctly categorize them
#
# + _uuid="356bfb3c7fae18c60e524d069aea3fd7ff3a974d"
# seting the function to show
def knowningData(df, data_type=object, limit=3): #seting the function with df,
n = df.select_dtypes(include=data_type) #selecting the desired data type
for column in n.columns: #initializing the loop
print("##############################################")
print("Name of column ", column, ': \n', "Uniques: ", df[column].unique()[:limit], "\n",
" | ## Total nulls: ", (round(df[column].isnull().sum() / len(df[column]) * 100,2)),
" | ## Total unique values: ", df_train.nunique()[column]) #print the data and % of nulls)
# print("Percentual of top 3 of: ", column)
# print(round(df[column].value_counts()[:3] / df[column].value_counts().sum() * 100,2))
print("#############################################")
# + [markdown] _uuid="08e06b3957406abc12ec9d781297de6c5d175852"
#
# ### I will by object data Type. <br>
# Click on "Output" to see the result
# + _kg_hide-output=true _uuid="070121f44dbf08c698aae911696f612c94aa07f8"
# calling our function: object is default
knowningData(df_train)
# + [markdown] _uuid="721eb4c604e2e7f8c6811d9af128a13ee0d1db9d"
# Printing Integers
# + _kg_hide-output=true _uuid="212795fc1d2d738a8365d1139d5a0205a8e790a1"
knowningData(df_train, data_type=int)
# + [markdown] _uuid="82938d22dcc4c7b69b58c1ac847e039a811990c7"
# Printing Float
# + _kg_hide-output=true _uuid="bbebcb86dafb074cc93cace4f2e19777617add30"
knowningData(df_train, data_type=float)
# + [markdown] _uuid="c6dc9005bec4736a6f2a08d6c47020e5f4a7a2d4"
# We haven't float datatype yet. <br>
# + [markdown] _uuid="5799654350f222aacbe552535cda00b21d583a84"
# ## I will drop some of this features and fillna or missing in some of them
# + _uuid="b8561c320415f62f4cd245091a47de9c62dd7659"
to_drop = ["socialEngagementType",'device.browserVersion', 'device.browserSize', 'device.flashVersion', 'device.language',
'device.mobileDeviceBranding', 'device.mobileDeviceInfo', 'device.mobileDeviceMarketingName', 'device.mobileDeviceModel',
'device.mobileInputSelector', 'device.operatingSystemVersion', 'device.screenColors', 'device.screenResolution',
'geoNetwork.cityId', 'geoNetwork.latitude', 'geoNetwork.longitude','geoNetwork.networkLocation',
'trafficSource.adwordsClickInfo.criteriaParameters', 'trafficSource.adwordsClickInfo.gclId', 'trafficSource.campaign',
'trafficSource.adwordsClickInfo.page', 'trafficSource.referralPath', 'trafficSource.adwordsClickInfo.slot',
'trafficSource.adContent', 'trafficSource.keyword']
# + _uuid="1a4b221505ff10c5e4c51e842365b6fe70c87d59"
df_train.drop(to_drop, axis=1, inplace=True)
# + _uuid="5ec423223d17d45ab7c5cf8e2678a852abc1c863"
print("Total features dropped: ", len(to_drop))
print("Shape after dropping: ", df_train.shape)
# + _uuid="ccf9c8f37da21f67b244b80506654ecd51d58fed"
# call the function to transform the numerical columns
df_train = NumericalColumns(df_train)
# Call the function that will normalize some features
df_train = Normalizing(df_train)
# + [markdown] _uuid="de06e72aa764f9a9ac4575e124358bfc209ddd60"
# # looking if we have any mistake on
#
# for c in dummy_feaures:
# if c in to_drop: print(c)
# + _uuid="4c933bff1466c8ba84d45e5aa691f4ca12ee307c"
# + [markdown] _uuid="83fdb2b3c0572195dc601187643a5622a2ce2587"
# ### Let's see the unique values in our dataset. <br>
#
# - if you want see click in "output"
# + _uuid="df45d83eba81c1688403c0219685818e1df43a8e"
# We will takeoff all columns where we have a unique value
# It is useful because this columns don't give us none information
clean_consts = [col for col in df_train.columns if df_train[col].nunique() == 1]
# this function drop all constant columns, inplacing the data
df_train.drop('trafficSource.adwordsClickInfo.adNetworkType', axis=1, inplace=True)
# printing the total of columns dropped and the name of columns
print("This useful action will drop: ", len(clean_consts), "columns")
print("All dropped columns: \n", clean_consts)
# + [markdown] _uuid="3965cebcc69089630b042f7a1df40040b7eb0309"
# The output show us totals.visits and trafficSource.adwordsClickInfo.adNetworkType, but totals,visits can be useful, so I will drop just trafficSource feature
# + _uuid="cf78dc9e490c5ca03ec89893de56d6b2636e95ea" _kg_hide-output=true
df_train.nunique()
# + [markdown] _uuid="f3e67ac43a41f8976098caddc7b2c5f215aae8cd"
# Excellent. Now we don't have more constant values
# + [markdown] _uuid="609ecb02fbeb7b45e0b434592d2e2d4c837f5062"
# ### Based on this output I will select and set a variable with all features by category
# + _uuid="463656bcaa564dc8530e96c3b645209927d989b8"
'trafficSource.adwordsClickInfo.adNetworkType'
# + _kg_hide-output=true _kg_hide-input=true _uuid="7a67f1a41573aada51a45e0bef3055210c96ff9e"
dummy_feaures =['channelGrouping', 'device.browser', 'device.deviceCategory', 'geoNetwork.city', 'device.operatingSystem',
'trafficSource.medium', 'trafficSource.source',
'geoNetwork.continent', 'geoNetwork.country', 'geoNetwork.metro', 'geoNetwork.networkDomain', 'geoNetwork.region',
'geoNetwork.subContinent']
numericals = ['totals.visits', '_visitHour', '_day', '_month', '_weekday']
# + [markdown] _uuid="646b37c2ea75c5da1fe2bc88b92b22424ab96b3f"
# # First, let see the distribuition of transactions Revenues
#
# I will start exploring the quantile
# + _uuid="8fddcf823310f3ac3b47a280817ac10361be34f9"
# Printing some statistics of our data
print("Transaction Revenue Min Value: ",
df_train[df_train['totals.transactionRevenue'] > 0]["totals.transactionRevenue"].min()) # printing the min value
print("Transaction Revenue Mean Value: ",
df_train[df_train['totals.transactionRevenue'] > 0]["totals.transactionRevenue"].mean()) # mean value
print("Transaction Revenue Median Value: ",
df_train[df_train['totals.transactionRevenue'] > 0]["totals.transactionRevenue"].median()) # median value
print("Transaction Revenue Max Value: ",
df_train[df_train['totals.transactionRevenue'] > 0]["totals.transactionRevenue"].max()) # the max value
# It I did to plot the quantiles but are not working
#print(round(df_train['totals.transactionRevenue'].quantile([.025,.25,.5,.75,.975]),2))
# seting the figure size of our plots
plt.figure(figsize=(14,5))
# Subplot allow us to plot more than one
# in this case, will be create a subplot grid of 2 x 1
plt.subplot(1,2,1)
# seting the distribuition of our data and normalizing using np.log on values highest than 0 and +
# also, we will set the number of bins and if we want or not kde on our histogram
ax = sns.distplot(np.log(df_train[df_train['totals.transactionRevenue'] > 0]["totals.transactionRevenue"] + 0.01), bins=40, kde=True)
ax.set_xlabel('Transaction RevenueLog', fontsize=15) #seting the xlabel and size of font
ax.set_ylabel('Distribuition', fontsize=15) #seting the ylabel and size of font
ax.set_title("Distribuition of Revenue Log", fontsize=20) #seting the title and size of font
# setting the second plot of our grid of graphs
plt.subplot(1,2,2)
# ordering the total of users and seting the values of transactions to understanding
plt.scatter(range(df_train.shape[0]), np.sort(df_train['totals.transactionRevenue'].values))
plt.xlabel('Index', fontsize=15) # xlabel and size of words
plt.ylabel('Revenue value', fontsize=15) # ylabel and size of words
plt.title("Revenue Value Distribution", fontsize=20) # Setting Title and fontsize
plt.show()
# + [markdown] _uuid="c8527ffb1923d433cd68c7cdcb06cc98687bf34f"
# Nice distribuition... We have very high values on the Transactions Revenue.
#
# + [markdown] _uuid="c71ed04299a99b634d251443a76cc8b2b960285a"
# ## I will see the kurtosis and Skewness of Transaction Revenue
# Skew and Kurtosis: 2 Important Statistics terms you need to know
#
#
# ## Skewness
# It is the degree of distortion from the symmetrical bell curve or the normal distribution. It measures the lack of symmetry in data distribution.<br>
# It differentiates extreme values in one versus the other tail. A symmetrical distribution will have a skewness of 0.
#
# <b>Positive Skewness </b>means when the tail on the right side of the distribution is longer or fatter. The mean and median will be greater than the mode.<br>
# <b>Negative Skewness </b> is when the tail of the left side of the distribution is longer or fatter than the tail on the right side. The mean and median will be less than the mode.
#
# #### So, when is the skewness too much?
# The rule of thumb seems to be:<br>
# If the skewness is between -0.5 and 0.5, the data are fairly symmetrical. <br>
# If the skewness is between -1 and -0.5(negatively skewed) or between 0.5 and 1(positively skewed), the data are moderately skewed.<br>
# If the skewness is less than -1(negatively skewed) or greater than 1(positively skewed), the data are highly skewed.<br>
#
#
# ## Kurtosis
# Kurtosis is all about the tails of the distribution — not the peakedness or flatness. It is used to describe the extreme values in one versus the other tail. <br>It is actually the measure of outliers present in the distribution.
#
# <b>High kurtosis</b> in a data set is an indicator that data has heavy tails or outliers. If there is a high kurtosis, then, we need to investigate why do we have so many outliers. It indicates a lot of things, maybe wrong data entry or other things. Investigate! <br>
# <b>Low kurtosis</b> in a data set is an indicator that data has light tails or lack of outliers. If we get low kurtosis(too good to be true), then also we need to investigate and trim the dataset of unwanted results
# + _uuid="1a471f060bdea1f011dbdd9dead1838f56a75ef8"
print('Excess kurtosis of normal distribution (should be 0): {}'.format(
kurtosis(df_train[df_train['totals.transactionRevenue'] > 0]["totals.transactionRevenue"])))
print( 'Skewness of normal distribution (should be 0): {}'.format(
skew((df_train[df_train['totals.transactionRevenue'] > 0]["totals.transactionRevenue"]))))
# + [markdown] _uuid="ec787d718c72215b52417e7bf1f6d2b3272e5065"
# Our data are fairly symmetrical skewed and have a High Kurtosis.
#
# I will see how many outliers we have on this dataset.
#
# + [markdown] _uuid="711c63585ac0f1fecad8ef51f0fb3ed2da1c9fda"
# ### Function that I created to find the map outlier values
# - Click on "code" to see the function
# + _uuid="d12790b819bd9bce414203233c2c1578fe59f2e7"
def CalcOutliers(df_num):
'''
<NAME> 20/10/2018
Set a numerical value and it will calculate the upper, lower and total number of outliers
It will print a lot of statistics of the numerical feature that you set on input
'''
# calculating mean and std of the array
data_mean, data_std = np.mean(df_num), np.std(df_num)
# seting the cut line to both higher and lower values
# You can change this value
cut = data_std * 3
#Calculating the higher and lower cut values
lower, upper = data_mean - cut, data_mean + cut
# creating an array of lower, higher and total outlier values
outliers_lower = [x for x in df_num if x < lower]
outliers_higher = [x for x in df_num if x > upper]
outliers_total = [x for x in df_num if x < lower or x > upper]
# array without outlier values
outliers_removed = [x for x in df_num if x > lower and x < upper]
print('Identified lowest outliers: %d' % len(outliers_lower)) # printing total number of values in lower cut of outliers
print('Identified upper outliers: %d' % len(outliers_higher)) # printing total number of values in higher cut of outliers
print('Identified outliers: %d' % len(outliers_total)) # printing total number of values outliers of both sides
print('Non-outlier observations: %d' % len(outliers_removed)) # printing total number of non outlier values
print("Total percentual of Outliers: ", round((len(outliers_total) / len(outliers_removed) )*100, 4)) # Percentual of outliers in points
return
# + _uuid="31a1e992143d98e30aabd5569d97f70bd92fcc72"
CalcOutliers(df_train['totals.transactionRevenue']) # Call
# + _uuid="5d132e66fde158c6a1d8daede257b93282a2fa5e"
CalcOutliers(df_train['totals.pageviews']) # Call
# + [markdown] _uuid="ed7eb5dda27ed49dc8208f258f1d5377750da132"
# # Device Browsers
# + _uuid="fd2613fb0a19aa87a0ec016378a9bca180fb3887"
# the top 10 of browsers represent % of total
print("Percentual of Browser usage: ")
print(df_train['device.browser'].value_counts()[:7] ) # printing the top 7 percentage of browsers
# seting the graph size
plt.figure(figsize=(14,6))
# Let explore the browser used by users
sns.countplot(df_train[df_train['device.browser']\
.isin(df_train['device.browser']\
.value_counts()[:10].index.values)]['device.browser'], palette="hls") # It's a module to count the category's
plt.title("TOP 10 Most Frequent Browsers", fontsize=20) # Adding Title and seting the size
plt.xlabel("Browser Names", fontsize=16) # Adding x label and seting the size
plt.ylabel("Count", fontsize=16) # Adding y label and seting the size
plt.xticks(rotation=45) # Adjust the xticks, rotating the labels
plt.show() #use plt.show to render the graph that we did above
# + [markdown] _uuid="a9993b94cbee5276e05776fd137a196e32b847ee"
# ### In our top 5 browsers we have more than 94% of total
# - TOP 1 - CHROME - 69,08%
# - TOP 2 - SAFARI - 20,04%
# - TOP 3 - FIREFOX - 3,77%
#
# Nothing new under the sun... Chrome is the most used followed by Safari and firefox.
#
# + [markdown] _uuid="ffb50b603e096223e98bc11d90711156998e8489"
# ## What if we cross the Revenue and Browser?
#
# + _uuid="c3ab87bfb285973429b671b966a60dc2c28881d6"
plt.figure(figsize=(13,6)) #figure size
#It's another way to plot our data. using a variable that contains the plot parameters
g1 = sns.boxenplot(x='device.browser', y='totals.transactionRevenue',
data=df_train[(df_train['device.browser'].isin((df_train['device.browser'].value_counts()[:10].index.values))) &
df_train['totals.transactionRevenue'] > 0])
g1.set_title('Browsers Name by Transactions Revenue', fontsize=20) # title and fontsize
g1.set_xticklabels(g1.get_xticklabels(),rotation=45) # It's the way to rotate the xticks when we use variable to our graphs
g1.set_xlabel('Device Names', fontsize=18) # Xlabel
g1.set_ylabel('Trans Revenue(log) Dist', fontsize=18) #Ylabel
plt.show()
# + [markdown] _uuid="7a1c76caa6384c3d353c87e051990dccab49534c"
# I think that it's very insightful information.
#
# Chrome have highest values in general but the highest value of transactions was did on Firefox.<br>
# We can see a "small" but consistent sells in Safari. Also IE and Edge give some results to Google;
# + [markdown] _uuid="7e8b0a9780d18ad0db49a53d7c996a85d16c632f"
# ## Let's see the Channel Grouping
# - The channel via which the user came to the Store.
# + _uuid="1019d4bf94a102d9ca256cd222fbc716afc94a58"
# the top 10 of browsers represent % of total
print("Percentual of Channel Grouping used: ")
print((df_train['channelGrouping'].value_counts()[:5])) # printing the top 7 percentage of browsers
# seting the graph size
plt.figure(figsize=(14,7))
# let explore the browser used by users
sns.countplot(df_train["channelGrouping"], palette="hls") # It's a module to count the category's
plt.title("Channel Grouping Count", fontsize=20) # seting the title size
plt.xlabel("Channel Grouping Name", fontsize=18) # seting the x label size
plt.ylabel("Count", fontsize=18) # seting the y label size
plt.show() #use plt.show to render the graph that we did above
# + [markdown] _uuid="56670cf50c09a55ee50200fec29a6fac3e1c4d7a"
# The TOP 5 Grouping Channels represents 97% of total values.
# Respectivelly:
# - TOP 1 => Organic Search - 42.99%
# - TOP 2 => Social - 24.39%
# - TOP 3 => Direct - 15.42%
# - TOP 4 => Referral - 11.89%
# - TOP 5 => Paid Search - 2.55%
#
# I have a new insight that I will explore furthuer. How wich channel are distributed by browsers?
#
# + [markdown] _uuid="c9672347d2dc14c59050af7e86499cc509b822ec"
# ## Crossing Channel Grouping x Browsers
# + _uuid="3f461736ae52ed6a36c892e77a265399ad3b0484"
## I will use the crosstab to explore two categorical values
# At index I will use set my variable that I want analyse and cross by another
crosstab_eda = pd.crosstab(index=df_train['channelGrouping'], normalize=True,
# at this line, I am using the isin to select just the top 5 of browsers
columns=df_train[df_train['device.browser'].isin(df_train['device.browser']\
.value_counts()[:5].index.values)]['device.browser'])
# Ploting the crosstab that we did above
crosstab_eda.plot(kind="bar", # select the bar to plot the count of categoricals
figsize=(14,7), # adjusting the size of graphs
stacked=True) # code to unstack
plt.title("Channel Grouping % for which Browser", fontsize=20) # seting the title size
plt.xlabel("The Channel Grouping Name", fontsize=18) # seting the x label size
plt.ylabel("Count", fontsize=18) # seting the y label size
plt.xticks(rotation=0)
plt.show() # rendering
# + [markdown] _uuid="270e9319019ef6f1fd0d25260d33e5455886eef2"
# Very cool! Interesting patterns
# + [markdown] _uuid="9dca0278af87c758626cefe003e9fb428b0542cc"
# # Operational System
# + _uuid="74d59c258d5102197b8c4e0ec15097bfed8aaafd"
# the top 5 of browsers represent % of total
print("Percentual of Operational System: ")
print(df_train['device.operatingSystem'].value_counts()[:5]) # printing the top 7 percentage of browsers
# seting the graph size
plt.figure(figsize=(14,7))
# let explore the browser used by users
sns.countplot(df_train["device.operatingSystem"], palette="hls") # It's a module to count the category's
plt.title("Operational System used Count", fontsize=20) # seting the title size
plt.xlabel("Operational System Name", fontsize=16) # seting the x label size
plt.ylabel("OS Count", fontsize=16) # seting the y label size
plt.xticks(rotation=45) # Adjust the xticks, rotating the labels
plt.show() #use plt.show to render the graph that we did above
# + [markdown] _uuid="b55e6e5f6d48e7fa519294f466ff287d0b77cfa7"
# The TOP 5 of Operational System corresponds to 96%.
#
# TOP 1 => Windows - 38.75% <br>
# TOP 2 => Macintosh - 28.04% <br>
# TOP 3 => Android - 14.15% <br>
# TOP 4 => iOS - 11.75% <br>
# TOP 5 => Linux - 3.91% <br>
#
# It's very interestign to me. In my country macbook isn't the most common SO. I will investigate further the SO by Country's
# + [markdown] _uuid="93e7fe418401f38b0dd67cf211194f9f089ee797"
# ## Now let's investigate the most used brower by Operational System
# + _uuid="13d40e390372c97905e30d0ca10ebad16719db6e"
# At index I will use isin to substitute the loop and get just the values with more than 1%
crosstab_eda = pd.crosstab(index=df_train[df_train['device.operatingSystem']\
.isin(df_train['device.operatingSystem']\
.value_counts()[:6].index.values)]['device.operatingSystem'],
# at this line, I am using the isin to select just the top 5 of browsers
columns=df_train[df_train['device.browser'].isin(df_train['device.browser']\
.value_counts()[:5].index.values)]['device.browser'])
# Ploting the crosstab that we did above
crosstab_eda.plot(kind="bar", # select the bar to plot the count of categoricals
figsize=(14,7), # adjusting the size of graphs
stacked=True) # code to unstack
plt.title("Most frequent OS's by Browsers of users", fontsize=22) # adjusting title and fontsize
plt.xlabel("Operational System Name", fontsize=19) # adjusting x label and fontsize
plt.ylabel("Count OS", fontsize=19) # adjusting y label and fontsize
plt.xticks(rotation=0) # Adjust the xticks, rotating the labels
plt.show() # rendering
# + [markdown] _uuid="f94c946788533ecedd3d397fc773c2f77317155a"
# ### Cool! <br>
#
# It's visually clear to see that chrome is the most used in all OS, less in iOS, that is a mobile OS.
#
# I will see if we can see a diference between the Revenues of transactions are different
# + [markdown] _uuid="5dabdf87282a0408a54388c08540f1a575edef7b"
# ## I will explore the distribuition of transaction Revenue by each OS
# + _uuid="d34317b1ba6d6e9f1f66c58a23c487a65e32ab11"
(sns.FacetGrid(df_train[(df_train['device.operatingSystem']\
.isin(df_train['device.operatingSystem']\
.value_counts()[:6].index.values)) & df_train['totals.transactionRevenue'] > 0],
hue='device.operatingSystem', height=5, aspect=2)
.map(sns.kdeplot, 'totals.transactionRevenue', shade=True)
.add_legend()
)
plt.show()
# + [markdown] _uuid="2fa6a06048c63a484b21d6295e9d5e56beea7bfd"
# Cool, we can have a better understanding of the distribution of Revenue by OS
# + [markdown] _uuid="47aec47461b0fe1b8dce656e1f1e5ce678c98a9a"
# ## Let's investigate the Device Category
# + _uuid="c27df69fb49cb019460d6d611e28970922dbfcd0"
# the top 5 of browsers represent % of total
print("Percentual of Operational System: ")
print(round(df_train['device.deviceCategory'].value_counts() / len(df_train['device.deviceCategory']) * 100, 2)) # printing the top 7 percentage of browsers
# seting the graph size
plt.figure(figsize=(14,5))
plt.subplot(1,2,1)
# let explore the browser used by users
sns.countplot(df_train["device.deviceCategory"], palette="hls") # It's a module to count the category's
plt.title("Device Category Count", fontsize=20) # seting the title size
plt.xlabel("Device Category", fontsize=18) # seting the x label size
plt.ylabel("Count", fontsize=16) # seting the y label size
plt.xticks(fontsize=18) # Adjust the xticks, rotating the labels
plt.subplot(1,2,2)
sns.boxenplot(x="device.deviceCategory", y = 'totals.transactionRevenue',
data=df_train[df_train['totals.transactionRevenue'] > 0], palette="hls") # It's a module to count the category's
plt.title("Device Category Revenue Distribuition", fontsize=20) # seting the title size
plt.xlabel("Device Category", fontsize=18) # seting the x label size
plt.ylabel("Revenue(Log)", fontsize=16) # seting the y label size
plt.xticks(fontsize=18) # Adjust the xticks, rotating the labels
plt.subplots_adjust(hspace = 0.9, wspace = 0.5)
plt.show() #use plt.show to render the graph that we did above
# + [markdown] _uuid="61c436b3b60cfa1e02720aae8c8a26dc2c2267fe"
# In percentual, we can see that :
# - desktop represents 73.5%
# - mobile represents 23.12%
# - tablet represents 3.38%
#
# I thought that the Revenue is almost all did by desktops. Let's explore it further.
# + [markdown] _uuid="8607632f5c289d212d4192bc9ba5ee3d910d8bec"
# ## Let's see the difference distribution between Devices
# + _uuid="e6fe29c132df19a74cc391c13d74b778a70c8d6f"
(sns.FacetGrid(df_train[df_train['totals.transactionRevenue'] > 0],
hue='device.deviceCategory', height=5, aspect=2)
.map(sns.kdeplot, 'totals.transactionRevenue', shade=True)
.add_legend()
)
plt.show()
# + [markdown] _uuid="af8ec25d7455e41340fe78fa43be71c4aa12db14"
# We have We can see the distribuition of
# + [markdown] _uuid="c1aa33b590e5f129f6b737e6cfe8888938aed000"
# ## Now, lets investigate the Device Category by Browsers
# + _uuid="815651b3895bf595d523d82c3b0d619edcbdf295"
# At index I will use isin to substitute the loop and get just the values with more than 1%
crosstab_eda = pd.crosstab(index=df_train['device.deviceCategory'], # at this line, I am using the isin to select just the top 5 of browsers
columns=df_train[df_train['device.operatingSystem']\
.isin(df_train['device.operatingSystem']\
.value_counts()[:6].index.values)]['device.operatingSystem'])
# Ploting the crosstab that we did above
crosstab_eda.plot(kind="bar", # select the bar to plot the count of categoricals
figsize=(14,7), # adjusting the size of graphs
stacked=True) # code to unstack
plt.title("Most frequent OS's by Device Categorys of users", fontsize=22) # adjusting title and fontsize
plt.xlabel("Device Name", fontsize=19) # adjusting x label and fontsize
plt.ylabel("Count Device x OS", fontsize=19) # adjusting y label and fontsize
plt.xticks(rotation=0) # Adjust the xticks, rotating the labels
plt.show() # rendering
# + [markdown] _uuid="fb3fb2e737a1aaf4bf2ea56d6ac906c66ce93432"
# Very interesting values.
# + [markdown] _uuid="0854b097c081212058908ef9c6ae156f1ff6ec44"
# # SubContinent
# + _uuid="2affc90a0388834255a5bb4114ee6ea773cff3d1"
# the top 8 of browsers represent % of total
print("Description of SubContinent count: ")
print(df_train['geoNetwork.subContinent'].value_counts()[:8]) # printing the top 7 percentage of browsers
# seting the graph size
plt.figure(figsize=(16,7))
# let explore the browser used by users
sns.countplot(df_train[df_train['geoNetwork.subContinent']\
.isin(df_train['geoNetwork.subContinent']\
.value_counts()[:15].index.values)]['geoNetwork.subContinent'], palette="hls") # It's a module to count the category's
plt.title("TOP 15 most frequent SubContinents", fontsize=20) # seting the title size
plt.xlabel("subContinent Names", fontsize=18) # seting the x label size
plt.ylabel("SubContinent Count", fontsize=18) # seting the y label size
plt.xticks(rotation=45) # Adjust the xticks, rotating the labels
plt.show() #use plt.show to render the graph that we did above
# + [markdown] _uuid="636413869fa5b17720448f9824bccebf1530d030"
# WoW, We have a very high number of users from North America.
#
# TOP 5 regions are equivalent of almost 70% +- of total
#
# TOP 1 => Northern America - 44.18% <br>
# TOP 2 => Southeast Asia - 8.29% <br>
# TOP 3 => Northern Europe - 6.73% <br>
# TOP 4 => Southern Asia - 6.33% <br>
# TOP 5 => Western Europe - 6.23% <br>
# + [markdown] _uuid="5bd94c1e9c0408a7faa215775f6df1d2915d8354"
# ## Let's cross the SubContinent by Browser
# + _uuid="27b2fd721d60cd80f9a5e3dc26e40fb91a8463a0"
## I will use the crosstab to explore two categorical values
# At index I will use isin to substitute the loop and get just the values with more than 1%
crosstab_eda = pd.crosstab(index=df_train[df_train['geoNetwork.subContinent']\
.isin(df_train['geoNetwork.subContinent']\
.value_counts()[:10].index.values)]['geoNetwork.subContinent'],
# at this line, I am using the isin to select just the top 5 of browsers
columns=df_train[df_train['device.browser'].isin(df_train['device.browser']\
.value_counts()[:5].index.values)]['device.browser'])
# Ploting the crosstab that we did above
crosstab_eda.plot(kind="bar", # select the bar to plot the count of categoricals
figsize=(16,7), # adjusting the size of graphs
stacked=True) # code to unstack
plt.title("TOP 10 Most frequent Subcontinents by Browsers used", fontsize=22) # adjusting title and fontsize
plt.xlabel("Subcontinent Name", fontsize=19) # adjusting x label and fontsize
plt.ylabel("Count Subcontinent", fontsize=19) # adjusting y label and fontsize
plt.xticks(rotation=45) # Adjust the xticks, rotating the labels
plt.legend(loc=1, prop={'size': 12}) # to
plt.show() # rendering
# + [markdown] _uuid="87d3c4aed0e28994d300405382507983aff5afe5"
# Nice, this graph is very insightful.
# The North America have a low ratio of Safari x Chrome... I thought that it was the contrary
#
# Firefox have a relative high presence in North America too.
# + _uuid="02e16f6e9bb5fa0d3195bb079012cabb08cbccea"
print('train date:', min(df_train['date']), 'to', max(df_train['date']))
# + _uuid="bc77dd4a9a1aed2aafe58484a4e8e51db06ed276"
year = df_train['_year'].value_counts() # counting the Year with value counts
month = df_train['_month'].value_counts() # coutning months
weeday = df_train['_weekday'].value_counts() # Couting weekday
day = df_train['_day'].value_counts() # counting Day
date = df_train['date'].value_counts() # Counting date
# + [markdown] _uuid="4e851acd6ae4f98f5e63c8a7befdbc3ec0e64016"
# ## INTERACTIVE DATE FEATURES
# + [markdown] _uuid="45e293c2c406fdb42409cd56c76597bbdbfdc010"
# ## First I will explore revenue and number of visits by day
# + _uuid="2cb8cdf11837f745da37f1faf9eb7d7215115d4b" _kg_hide-input=true
# I saw and take a lot of inspiration to this interactive plots in kernel:
# https://www.kaggle.com/jsaguiar/complete-exploratory-analysis-all-columns
# I learned a lot in this kernel and I will implement and adapted some ideas
#seting some static color options
color_op = ['#5527A0', '#BB93D7', '#834CF7', '#6C941E', '#93EAEA', '#7425FF', '#F2098A', '#7E87AC',
'#EBE36F', '#7FD394', '#49C35D', '#3058EE', '#44FDCF', '#A38F85', '#C4CEE0', '#B63A05',
'#4856BF', '#F0DB1B', '#9FDBD9', '#B123AC']
# Visits by time train
# couting all entries by date to get number of visits by each date
dates_temp = df_train['date'].value_counts().to_frame().reset_index().sort_values('index')
# renaming the columns to apropriate names
dates_temp = dates_temp.rename(columns = {"date" : "visits"}).rename(columns = {"index" : "date"})
# creating the first trace with the necessary parameters
trace = go.Scatter(x=dates_temp.date.astype(str), y=dates_temp.visits,
opacity = 0.8, line = dict(color = color_op[3]), name= 'Visits by day')
# Below we will get the total values by Transaction Revenue Log by date
dates_temp_sum = df_train.groupby('date')['totals.transactionRevenue'].sum().to_frame().reset_index()
# using the new dates_temp_sum we will create the second trace
trace1 = go.Scatter(x=dates_temp_sum.date.astype(str), line = dict(color = color_op[1]), name="RevenueLog by day",
y=dates_temp_sum['totals.transactionRevenue'], opacity = 0.8)
# Getting the total values by Transactions by each date
dates_temp_count = df_train[df_train['totals.transactionRevenue'] > 0].groupby('date')['totals.transactionRevenue'].count().to_frame().reset_index()
# using the new dates_temp_count we will create the third trace
trace2 = go.Scatter(x=dates_temp_count.date.astype(str), line = dict(color = color_op[5]), name="Sellings by day",
y=dates_temp_count['totals.transactionRevenue'], opacity = 0.8)
#creating the layout the will allow us to give an title and
# give us some interesting options to handle with the outputs of graphs
layout = dict(
title= "Informations by Date",
xaxis=dict(
rangeselector=dict(
buttons=list([
dict(count=1, label='1m', step='month', stepmode='backward'),
dict(count=3, label='3m', step='month', stepmode='backward'),
dict(count=6, label='6m', step='month', stepmode='backward'),
dict(step='all')
])
),
rangeslider=dict(visible = True),
type='date'
)
)
# creating figure with the both traces and layout
fig = dict(data= [trace, trace1, trace2], layout=layout)
#rendering the graphs
iplot(fig) #it's an equivalent to plt.show()
# + [markdown] _uuid="58dbc1d65e92ba700c741839c9a4222bf35c11c7"
# ### Creating an Sofistcated interactive graphics to better understanding of date features
#
# To see the code click in "code".
# ## SELECT THE OPTION:
# + _uuid="864f5577ef8b0ab3ac4f3a14537196b8b40eb2e3" _kg_hide-output=false _kg_hide-input=true
# Setting the first trace
trace1 = go.Histogram(x=df_train["_year"],
name='Year Count')
# Setting the second trace
trace2 = go.Histogram(x=df_train["_month"],
name='Month Count')
# Setting the third trace
trace3 = go.Bar(y=day.values,
x=day.index.values,
name='Day Count')
# Setting the fourth trace
trace4 = go.Bar(y=weeday.values,
x=weeday.index.values,
name='Weekday Count')
# puting all traces in the same "array of graphics" to we render it below
data = [trace1, trace2, trace4, trace3]
#Creating the options to be posible we use in our
updatemenus = list([
dict(active=-1,
x=-0.15,
buttons=list([
dict(
label = 'Years Count',
method = 'update',
args = [{'visible': [True, False, False, False,False]},
{'title': 'Count of Year'}]),
dict(
label = 'Months Count',
method = 'update',
args = [{'visible': [False, True, False, False,False]},
{'title': 'Count of Months'}]),
dict(
label = 'WeekDays Count',
method = 'update',
args = [{'visible': [False, False, True, False, False]},
{'title': 'Count of WeekDays'}]),
dict(
label = 'Days Count ',
method = 'update',
args = [{'visible': [False, False, False, True,False]},
{'title': 'Count of Day'}]) ])
)
])
layout = dict(title='The percentual Distribuitions of Date Features (Select from Dropdown)',
showlegend=False,
updatemenus=updatemenus,
# xaxis = dict(
# type="category"
# ),
barmode="group"
)
fig = dict(data=data, layout=layout)
print("SELECT BELOW: ")
iplot(fig)
# + [markdown] _uuid="0a9bcb4e07e0416ac16ab3a7d31ae21e717270b8"
# ******* *How can I set order to my year, months and days?* *******
# + [markdown] _uuid="b144feec1b14923c1237d7eaa8666906ebf1c101"
# ### Very Cool graphs.
#
# WE can see that the number of access are clearly downing by the through the time.
#
# - The months with highest accesses are October and November.
# - On the Weekend the trafic is lower than other days.
# - The 5 days with highest number of accesses is 1 and 5
# - Considering the full count of dates, we can see that the days with highest accesses are almost all in november/2016
#
# + [markdown] _uuid="27d53dc8b53550a8a937f7b30b96f6da9b9ef08c"
# #### Let's investigate the VisitHour and weekday to see if we can find some interesting patterns
# + _uuid="e1975529c7fbc0866acd700cba2fd6b6e234070e"
date_sales = ['_visitHour', '_weekday'] #seting the desired
cm = sns.light_palette("green", as_cmap=True)
pd.crosstab(df_train[date_sales[0]], df_train[date_sales[1]],
values=df_train["totals.transactionRevenue"], aggfunc=[np.sum]).style.background_gradient(cmap = cm)
# tab.columns.levels[1] = ["Sun", "Mon", "Thu", "wed", "Thi","Fri","Sat"]
# + [markdown] _uuid="b22a4329690db833383cde3d3aedb351ddb1666b"
# Very interesting, we can see that from 17 to 20 hour we have the highest numbers of
# + [markdown] _uuid="32dc0d6b64c8d5bb05139b472abc47d17176f00e"
# ## I will use a interesting graphic called Squarify
# - I will apply it in feature Country to discovery where the user access the store
# + _uuid="963ce4b784c466f9aeb7490df57935f440c61096"
number_of_colors = 20 # total number of different collors that we will use
# Here I will generate a bunch of hexadecimal colors
color = ["#"+''.join([random.choice('0123456789ABCDEF') for j in range(6)])
for i in range(number_of_colors)]
# + [markdown] _uuid="66ede41d597a59aa2c44955b29858855b8f37123"
# ## Exploring Countrys
# + _uuid="3b4f42be8780ba5abb86491b95b6c05384d43309"
country_tree = df_train["geoNetwork.country"].value_counts() #counting the values of Country
print("Description most frequent countrys: ")
print(country_tree[:15]) #printing the 15 top most
country_tree = round((df_train["geoNetwork.country"].value_counts()[:30] \
/ len(df_train['geoNetwork.country']) * 100),2)
plt.figure(figsize=(14,5))
g = squarify.plot(sizes=country_tree.values, label=country_tree.index,
value=country_tree.values,
alpha=.4, color=color)
g.set_title("'TOP 30 Countrys - % size of total",fontsize=20)
g.set_axis_off()
plt.show()
# + [markdown] _uuid="a4b96e6e0e11be37f004dd6e1a1d49a3e416292d"
# USA have a very highest value than another countrys.
#
# Below I will take a look on cities and find for the highest revenues from them
#
# + [markdown] _uuid="a39f2bd479c36e09ad97fc5cb7233b625b700b81"
# ## Now, I will look on City feature and see the principal cities in the dataset
# + _uuid="5142be3a58f77f0de9b8d8b03cbfb8230155179f"
df_train.loc[df_train["geoNetwork.city"] == "not available in demo dataset", 'geoNetwork.city'] = np.nan
# + _uuid="744e3f288212cfcd87fb077569b04314bbeb4fef"
city_tree = df_train["geoNetwork.city"].value_counts() #counting
print("Description most frequent Citys: " )
print(city_tree[:15])
city_tree = round((city_tree[:30] / len(df_train['geoNetwork.city']) * 100),2)
plt.figure(figsize=(14,5))
g = squarify.plot(sizes=city_tree.values, label=city_tree.index,
value=city_tree.values,
alpha=.4, color=color)
g.set_title("'TOP 30 Citys - % size of total",fontsize=20)
g.set_axis_off()
plt.show()
# + [markdown] _uuid="8cb4ca4f8fca49baeab424aa9fd9140f0567effe"
# Nicelly distributed clients that accessed the store.
# (non set) have 3.81% of total, so I dont will consider in top five, but it was the top 2 most frequent.
#
# The top 5 are:
# - Montain View
# - New York
# - San Francisco
# - Sunnyvale
# - London
#
# And in terms of money, how the Countrys and Cities are ?
#
# + [markdown] _uuid="9344c85fbc4351d8e35912e1bce2c388e421a486"
# ____________________
# + [markdown] _uuid="a4b4b66d07c41db917702540e6758d959a6a6c19"
# ### Creating a function with plotly to better investigate the dataset
#
# - Click in "code" to see the commented code
# + _kg_hide-input=true _uuid="ba3cacf3d0cc127d8b35b41133604e6e26cbd571"
def PieChart(df_colum, title, limit=15):
"""
This function helps to investigate the proportion of visits and total of transction revenue
by each category
"""
count_trace = df_train[df_colum].value_counts()[:limit].to_frame().reset_index()
rev_trace = df_train.groupby(df_colum)["totals.transactionRevenue"].sum().nlargest(10).to_frame().reset_index()
trace1 = go.Pie(labels=count_trace['index'], values=count_trace[df_colum], name= "% Acesses", hole= .5,
hoverinfo="label+percent+name", showlegend=True,domain= {'x': [0, .48]},
marker=dict(colors=color))
trace2 = go.Pie(labels=rev_trace[df_colum],
values=rev_trace['totals.transactionRevenue'], name="% Revenue", hole= .5,
hoverinfo="label+percent+name", showlegend=False, domain= {'x': [.52, 1]})
layout = dict(title= title, height=450, font=dict(size=15),
annotations = [
dict(
x=.25, y=.5,
text='Visits',
showarrow=False,
font=dict(size=20)
),
dict(
x=.80, y=.5,
text='Revenue',
showarrow=False,
font=dict(size=20)
)
])
fig = dict(data=[trace1, trace2], layout=layout)
iplot(fig)
# + [markdown] _uuid="23e1f787c9d000554066aea293f8808a8de424af"
# ## Device Category feature
# + _uuid="104d01069fe11a66ff3738f206b791d18150af3b"
PieChart("device.deviceCategory", "Device Category")
# + [markdown] _uuid="38b54ca57e57bf4ddade4cd7889485c7c88911e7"
# ## I will apply the Prie Chart in Country's again
# + _uuid="1648ff7cdb2548efe8d357f0a9e88b6549451ce9"
# call the function
PieChart("geoNetwork.city", "Top Cities by Accesses and Revenue", limit=12)
# + [markdown] _uuid="6e77cfdac5dea3046ae40f35713b61d1272a2232"
# - New York is responsible by 14% of visits and 31% of revenues.
# - Montain view have 19% in visists but just 16% of revenues
# - Chicago have just 3.5% of visits but have a high significance in revenues
# + [markdown] _uuid="3b95a7ba1b2337466a8e708019a668b8a4ed49e2"
# ## Seeing again Channel Grouping more specified
# + _uuid="b3296caa545f703647e2e8e46acb0fb422370715"
PieChart("channelGrouping", "Channel Grouping Visits and Revenues")
# + [markdown] _uuid="73c48a5cbeb35949101fc3f909246563b457482a"
# It's interesting to note that Referral have a less number of Visits but is responsible for almost 40% of revenues****
# + [markdown] _uuid="3706ce71368c3957579e38bdb40e032d5e22d18d"
# ### Months in pizza graph
# + [markdown] _uuid="036612794be2040bac93d4918d73bb6d9f9b8515"
# ## Let's see the NetWork Domain
# - I will plot visits and revenues by each category, including the non-set and unknown accesses and revenues
#
# + _uuid="202155e1496d32e68b2e611bbfa9674b4e0ebc7f"
PieChart('geoNetwork.networkDomain', "Network Domain")
# + [markdown] _uuid="b3dd496495bed44e84d481d2a8c2241ca138a212"
# Wow, another very cool information.
#
# - (not set) domain have almost 50% of total visits and 62% of Revenues.
# - Unknown is responsible by 28% of visits but just 2.70% of Revenues
# - comcast.net have 5.5% of visits and 7.4% Revenues.
# + [markdown] _uuid="81305283f850873c66a777edb7315611fd257680"
# Let's take a look on Mobile and Browser proportions
# + _uuid="e1081175860b562099b45c115c751d3487f9c24c"
PieChart("device.deviceCategory", "Device Category")
# + [markdown] _uuid="f11af04271ed24a89c23fde30403cff01020d129"
# The absolutelly high part of revenues are from Desktop Devices
# + [markdown] _uuid="620cc879e9f23ebf410b2f37ccf2bdedcca9fcbf"
# ## Trafic Source Medium
# + _uuid="04fb4fe637312415b073ecd3528bc1198881844a"
PieChart("trafficSource.medium", "Trafic Source - Medium")
# + [markdown] _uuid="e8755bf3ed97eca7561b5d27f1e3cd1311223c22"
# - Organic have highest number of visits but is the third in revenues
# - Referral have almost 40% in both Visits and Revenues
# - The none category have almost 16% of visits but almost 40% of revenues
# + [markdown] _uuid="dc86a9a774f05532d6c45ec817cfb7d2451db84f"
# Now I will take a look on trafficSource section, the Source to access the store
# + _uuid="d86ecf58c561858347391fe8e16bf518dfcb4bd2"
PieChart('trafficSource.source', "Visits and Revenue by TOP Sources", limit=8)
# + [markdown] _uuid="4cabc13925d8cc56bc28591806ac9975ce4cd11d"
# We have a high number of visits from youtube but 0 sales. <br>
# the mall.googleplex is have a low number of access but have the highest value in revenues
# + [markdown] _uuid="6be8ccdae823f3ccaad68ec01ac87a7c88862407"
# # I will continue this notebook! Votes up the kernel and stay tuned to next updates
# + _uuid="6b63bc0f0ba6b6c9f7350878d11759576b370a08"
df_train.corr()['totals.transactionRevenue']
# + [markdown] _uuid="3259db00fe1f717acb96ae78a6f068b518c81cee"
# Seeing the crosstab with heatmap
# + _uuid="fe02aae49c1f4962bebdd02013d8a89f63e66380"
country_repayment = ['channelGrouping', '_weekday'] #seting the desired
cm = sns.light_palette("green", as_cmap=True)
pd.crosstab(df_train[country_repayment[0]], df_train[country_repayment[1]],
values=df_train["totals.transactionRevenue"], aggfunc=[np.sum]).style.background_gradient(cmap = cm)
# tab.columns.levels[1] = ["Sun", "Mon", "Thu", "wed", "Thi","Fri","Sat"]
# + [markdown] _uuid="4fd8dfe3a2014e8eadfdb3b8190ea866de926f23"
# ## Geolocation plot to visually understand the data
# + _uuid="f35db04e9e4dbceeb77029df6af763b1700d55cc" _kg_hide-input=true
# Counting total visits by countrys
countMaps = pd.DataFrame(df_train['geoNetwork.country'].value_counts()).reset_index()
countMaps.columns=['country', 'counts'] #renaming columns
countMaps = countMaps.reset_index().drop('index', axis=1) #reseting index and droping the column
data = [ dict(
type = 'choropleth',
locations = countMaps['country'],
locationmode = 'country names',
z = countMaps['counts'],
text = countMaps['country'],
autocolorscale = False,
marker = dict(
line = dict (
color = 'rgb(180,180,180)',
width = 0.5
) ),
colorbar = dict(
autotick = False,
tickprefix = '',
title = 'Number of Visits'),
) ]
layout = dict(
title = 'Couting Visits Per Country',
geo = dict(
showframe = False,
showcoastlines = True,
projection = dict(
type = 'Mercator'
)
)
)
figure = dict( data=data, layout=layout )
iplot(figure, validate=False, filename='map-countrys-count')
# + [markdown] _uuid="932679723c6b45dd99e3d10709650571545b8361"
# ## Total Revenues by Country
# + _uuid="349766424b33710198240cc3f068b0a0926a250d" _kg_hide-input=true
# I will crete a variable of Revenues by country sum
sumRevMaps = df_train[df_train['totals.transactionRevenue'] > 0].groupby("geoNetwork.country")["totals.transactionRevenue"].count().to_frame().reset_index()
sumRevMaps.columns = ["country", "count_sales"] # renaming columns
sumRevMaps = sumRevMaps.reset_index().drop('index', axis=1) #reseting index and drop index column
data = [ dict(
type = 'choropleth',
locations = sumRevMaps['country'],
locationmode = 'country names',
z = sumRevMaps['count_sales'],
text = sumRevMaps['country'],
autocolorscale = False,
marker = dict(
line = dict (
color = 'rgb(180,180,180)',
width = 0.5
) ),
colorbar = dict(
autotick = False,
tickprefix = '',
title = 'Count of Sales'),
) ]
layout = dict(
title = 'Total Sales by Country',
geo = dict(
showframe = False,
showcoastlines = True,
projection = dict(
type = 'Mercator'
)
)
)
figure = dict( data=data, layout=layout )
iplot(figure, validate=False, filename='map-countrys-total')
# + [markdown] _uuid="d0bce740990e19317f2495dbe741eda30a2b1aea"
# ### Some tests that I am doing to try find interesting feature engineering approaches
# + _uuid="0cb4f9106082d8d96af3c1236550dc33d0cd4175" _kg_hide-output=true _kg_hide-input=true
df_train['month_unique_user_count'] = df_train.groupby('_month')['fullVisitorId'].transform('nunique')
df_train['day_unique_user_count'] = df_train.groupby('_day')['fullVisitorId'].transform('nunique')
df_train['weekday_unique_user_count'] = df_train.groupby('_weekday')['fullVisitorId'].transform('nunique')
df_train['traf_sourc_browser_count'] = df_train.groupby(['trafficSource.medium', 'device.browser'])['totals.pageviews'].transform('nunique')
df_train['Id_browser_pageviews_sumprod'] = df_train.groupby(['fullVisitorId', 'device.browser'])['totals.pageviews'].transform('cumprod')
df_train['Id_browser_hits_sumprod'] = df_train.groupby(['fullVisitorId', 'device.browser'])['totals.hits'].transform('cumprod')
df_train['Id_browser_hits_sumprod'] = df_train.groupby(['fullVisitorId', 'device.browser'])['totals.hits'].transform('cumprod')
df_train['Id_browser_hits_sumprod_mob'] = df_train.groupby(['fullVisitorId', 'device.browser', 'device.isMobile'])['totals.hits'].transform('sum')
df_train['Id_networkDomain_hits'] = df_train.groupby(['fullVisitorId', 'geoNetwork.networkDomain'])['totals.hits'].transform('var')
df_train['Id_networkDomain_country_hits'] = df_train.groupby(['fullVisitorId', 'geoNetwork.networkDomain', 'geoNetwork.country'])['totals.hits'].transform('unique')
# + _uuid="6437a9d5ff5bbb8b0eb6791e832cc2401926fc2c" _kg_hide-output=true _kg_hide-input=true
df_train[["totals.transactionRevenue", 'Id_browser_hits_sumprod', 'Id_networkDomain_hits','Id_networkDomain_country_hits', 'Id_browser_hits_sumprod_mob']].corr()
# + [markdown] _uuid="da64b44e601cf75d0d75673f90f0641a4c0950d6"
# ## Preprocessing the fulldataset and creating new features
# + _uuid="133ea13b036829110319da4f4e69eee0b6334185"
aggs = {
'date': ['min', 'max'],
'totals.hits': ['sum', 'min', 'max', 'mean', 'median'],
'totals.pageviews': ['sum', 'min', 'max', 'mean', 'median'],
'totals.bounces': ['sum', 'mean', 'median'],
'totals.newVisits': ['sum', 'mean', 'median']
}
# Previous applications categorical features
cat_aggregations = {}
for cat in dummy_feaures:
cat_aggregations[cat] = ['min', 'max', 'mean']
prev_agg = df_train.groupby('fullVisitorId').agg({**aggs})
prev_agg.columns = pd.Index(['Agg_' + e[0] + "_" + e[1].upper() for e in prev_agg.columns.tolist()])
# + _uuid="52dc6b6e78b98fb547f4a6012095c3fbb6b7b870"
prev_agg
# + _uuid="a7ef38820e9fb49ad5f1d71f253b2fa803e80029"
new_columns = [
k + '_' + agg for k in aggs.keys() for agg in aggs[k]
]
new_columns
# + _uuid="d4b67b55347c758ce33b2e725c399cbb666e7e84"
dummy_feaures
# + _uuid="b5bfd985138a4abbe288f12a8838a97e763b1bcf"
### Testing some grouping approaches
# + _uuid="55e6693014ca3cb1bc2938d64b6b3371b8a42ddd"
df_train['cumcount'] = df_train.groupby('fullVisitorId').cumcount() + 1
# + [markdown] _uuid="3139a6d399c80ee47c84ee0d05d5f9d9066bdbd6"
# Some tests to feature engineering
# + _uuid="29532ab5a4ec9c636d6d9937f678d708dc2b25e6"
aggs = {
'date': ['min', 'max'],
'totals.transactionRevenue': ['sum', 'size'],
'totals.hits': ['sum', 'min', 'max', 'count', 'median'],
'totals.pageviews': ['sum', 'min', 'max', 'mean', 'median'],
'totals.bounces': ['sum', 'mean', 'median'],
'totals.newVisits': ['sum', 'mean', 'median']
}
# Previous applications categorical features
cat_aggregations = {}
for cat in dummy_feaures:
cat_aggregations[cat] = ['min', 'max', 'mean']
prev_agg = df_train.groupby('fullVisitorId').agg({**aggs})
prev_agg.head()
# + [markdown] _uuid="23b75275bdf104dc889476245e3e9bbc43723b27"
# # I will continue working on this kernel, stay tuned
#
# ******** Please, if you liked this kernel don't forget to votes up and give your feedback ********
# + _uuid="8c8ee2fd6974dacf00d5590a9b701f7c10d7639c"
prev_agg.columns = ["_".join(x) for x in prev_agg.columns.ravel()]
# + _uuid="6fb8904cdf0549f7011328aba6a18f5baac97f11"
prev_agg.head()
# + _uuid="8f46e405dd6662edc927dd42aa30c97d57ab1e2b"
| 9 google customer revenue prediction/exploring-the-consumer-patterns-ml-pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.12 ('two-hearts')
# language: python
# name: python3
# ---
# + [markdown] id="qIERnt9pC-yz"
# # Questioning the Effect of Physiological Heartbeat Synchrony in Romantic Dyads. A Preregistered Deep Learning Analysis.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 26463, "status": "ok", "timestamp": 1653394797559, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="W4yMu0MJkcft" outputId="c4d410c0-0b5c-4d22-afef-06f91837346a"
# For Google Colab / local machine
import tensorflow.keras
ver = tensorflow.version.VERSION
if float(ver[:3]) > 2.7:
print("Latest TensorFlow version detected -> Prepare Google Colab usage\n")
google_colab = 1
from google.colab import drive
drive.mount('/content/drive/')
colab_path = "/content/drive/MyDrive/Masterarbeit/Code/two-hearts/"
import sys
sys.path.append(colab_path)
else:
colab_path = ""
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1606, "status": "ok", "timestamp": 1653394799152, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="JXC8BydxC-zA" outputId="6c1a58ca-4831-4478-dfe3-2514c347131e"
# Import libraries
from lists import dyads, num_dyads, participants, num_participants
import os
import datetime
import random
import pickle
import IPython
import IPython.display
import numpy as np
from numpy import array, hstack
import pandas as pd
from copy import deepcopy
import matplotlib.pyplot as plt
from tensorflow.python.keras.layers import deserialize, serialize
from tensorflow.python.keras.saving import saving_utils
from tensorflow.keras.models import Sequential, Model, load_model
from tensorflow.keras.layers import LSTM, Dense, RepeatVector, TimeDistributed, Input, BatchNormalization, multiply, concatenate, Flatten, Activation, dot
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.utils import plot_model
from tensorflow.keras.callbacks import EarlyStopping
import pydot as pyd
from tensorflow.keras.utils import plot_model, model_to_dot
tensorflow.keras.utils.pydot = pyd
print("TensorFlow version:", tensorflow.version.VERSION)
# + executionInfo={"elapsed": 24, "status": "ok", "timestamp": 1653394799155, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="SL0-lMWtAS3r"
# Function for controlling random number generator
def seed_value(seed):
os.environ['PYTHONHASHSEED'] = str(seed)
random.seed(seed)
np.random.seed(seed)
tensorflow.random.set_seed(seed)
# + [markdown] id="iYhFodT5C-y_"
# ## Deep Learning
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 26, "status": "ok", "timestamp": 1653394799159, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="YT9ANStQiIMB" outputId="8b18909c-6411-48c0-bd86-e12f25c2e417"
# Set sampling rate
sampling_rate = 50
print("Sampling rate:", sampling_rate)
# Set number of time steps
n_steps_in, n_steps_out = 5*sampling_rate, 2*sampling_rate
print("Time steps:", n_steps_in, n_steps_out)
# Set conditions
condition = ["sit", "gaze", "gaze_swap"]
print("Conditions:", condition)
# Show number of participants
print("Participants:", num_participants)
# Show number of dyads
print("Dyads:", num_dyads)
# Set number of trials
trial = ["01", "02", "03"]
# trial = list(range(0,3))
print("Number of training trials:", len(trial))
# + executionInfo={"elapsed": 25, "status": "ok", "timestamp": 1653394799161, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="gCmGEuUyKyFc"
# Prepare sample data
def sample_preperation(condition):
# Load data
data = np.load(f"{colab_path}data/data_{condition}.npy")
print(f"Loaded data with shape {data.shape} and type {data.dtype}")
# Create empty samples
X_input_train = np.empty((0, n_steps_in, 2))
X_input_vali = np.empty((0, n_steps_in, 2))
X_input_test = np.empty((0, n_steps_in, 2))
y_output_train = np.empty((0, n_steps_out, 2))
y_output_vali = np.empty((0, n_steps_out, 2))
y_output_test = np.empty((0, n_steps_out, 2))
# Split a multivariate sequence into samples (modified from Brownlee 2018, p.156)
def split_sequences(sequences, n_steps_in, n_steps_out):
X, y = list(), list()
for i in range(len(sequences)):
if i % (sampling_rate) == 0: # to reduce redundancy in samples
# find the end of this pattern
end_ix = i + n_steps_in
out_end_ix = end_ix + n_steps_out
# check if we are beyond the dataset
if out_end_ix > len(sequences):
break
# gather input and output parts of the pattern
seq_x, seq_y = sequences[i:end_ix, :], sequences[end_ix:out_end_ix, :]
X.append(seq_x)
y.append(seq_y)
return array(X), array(y)
for i, idx in enumerate(list(range(num_participants))[::2]): # index for dyads
# define input sequence
in_seq1 = data[idx]
in_seq2 = data[idx+1]
# convert to [rows, columns] structure
in_seq1 = in_seq1.reshape((len(in_seq1), 1))
in_seq2 = in_seq2.reshape((len(in_seq2), 1))
# horizontally stack columns
dataset = hstack((in_seq1, in_seq2))
# covert into input/output
X, y = split_sequences(dataset, n_steps_in, n_steps_out)
# data split
num_train_samples = int(0.6 * len(X))
num_val_samples = int(0.2 * len(X))
num_test_samples = len(X) - num_train_samples - num_val_samples
# append data for multiple dyads
X_input_train = np.append(
X_input_train,
X[:num_train_samples],
axis=0)
y_output_train = np.append(
y_output_train,
y[:num_train_samples],
axis=0)
X_input_vali = np.append(
X_input_vali,
X[num_train_samples:(num_train_samples+num_val_samples)],
axis=0)
y_output_vali = np.append(
y_output_vali,
y[num_train_samples:(num_train_samples+num_val_samples)],
axis=0)
X_input_test = np.append(
X_input_test,
X[(num_train_samples+num_val_samples):],
axis=0)
y_output_test = np.append(
y_output_test,
y[(num_train_samples+num_val_samples):],
axis=0)
# Create dictionary
samples = {
"X_input_train": X_input_train,
"y_output_train": y_output_train,
"X_input_vali": X_input_vali,
"y_output_vali": y_output_vali,
"X_input_test": X_input_test,
"y_output_test": y_output_test
}
print("Length of samples for each set:",
len(X_input_train),
len(X_input_vali),
len(X_input_test))
return samples
# + executionInfo={"elapsed": 25, "status": "ok", "timestamp": 1653394799162, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="jpbTCapSdNK7"
# Define simple seq2seq model
# Modified from Wieniawska 2020
# (https://levelup.gitconnected.com/building-seq2seq-lstm-with-luong-attention-in-keras-for-time-series-forecasting-1ee00958decb)
def lstm_decoder_encoder(samples, units=100):
# Input layer
input_train = Input(
shape=(samples["X_input_train"].shape[1],
samples["X_input_train"].shape[2]))
output_train = Input(
shape=(samples["y_output_train"].shape[1],
samples["y_output_train"].shape[2]))
# Encoder LSTM with last state_h and state_c
encoder_last_h1, encoder_last_h2, encoder_last_c = LSTM(
units,
activation='elu',
dropout=0.2,
recurrent_dropout=0.2,
return_sequences=False,
return_state=True)(input_train)
# Batch normalisation to avoid gradient explosion
encoder_last_h1 = BatchNormalization(momentum=0.6)(encoder_last_h1)
encoder_last_c = BatchNormalization(momentum=0.6)(encoder_last_c)
# Decoder LSTM
decoder = RepeatVector(output_train.shape[1])(encoder_last_h1)
decoder = LSTM(
units,
activation='elu',
dropout=0.2,
recurrent_dropout=0.2,
return_state=False,
return_sequences=True)(decoder, initial_state=[encoder_last_h1, encoder_last_c])
# Dense layer with repeated weights
out = TimeDistributed(Dense(output_train.shape[2]))(decoder)
# Compile model
model = Model(inputs=input_train, outputs=out)
opt = Adam(learning_rate=0.001, clipnorm=1)
model.compile(optimizer=opt, loss='mean_squared_error', metrics=['mae'])
print(model)
return model
# + executionInfo={"elapsed": 27, "status": "ok", "timestamp": 1653394799165, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="IVg46Gb1iIMI"
# Define seq2seq model with Luong attention
# Modified from Wieniawska 2020
# (https://levelup.gitconnected.com/building-seq2seq-lstm-with-luong-attention-in-keras-for-time-series-forecasting-1ee00958decb)
def lstm_decoder_encoder_luong_attention(samples, units=100):
# Input layer
input_train = Input(
shape=(samples["X_input_train"].shape[1],
samples["X_input_train"].shape[2]))
output_train = Input(
shape=(samples["y_output_train"].shape[1],
samples["y_output_train"].shape[2]))
# Encoder LSTM
encoder_stack_h, encoder_last_h, encoder_last_c = LSTM(
units,
activation='elu',
dropout=0.2,
recurrent_dropout=0.2,
return_state=True,
return_sequences=True)(input_train)
# Batch normalisation to avoid gradient explosion
encoder_last_h = BatchNormalization(momentum=0.6)(encoder_last_h)
encoder_last_c = BatchNormalization(momentum=0.6)(encoder_last_c)
# Decoder LSTM
decoder_input = RepeatVector(output_train.shape[1])(encoder_last_h)
decoder_stack_h = LSTM(
units,
activation='elu',
dropout=0.2,
recurrent_dropout=0.2,
return_state=False,
return_sequences=True)(decoder_input, initial_state=[encoder_last_h, encoder_last_c])
# Attention layer
attention = dot([decoder_stack_h, encoder_stack_h], axes=[2, 2])
attention = Activation('softmax')(attention)
# Calculate context vector with batch normalisation
context = dot([attention, encoder_stack_h], axes=[2, 1])
context = BatchNormalization(momentum=0.6)(context)
# Combine context vector with stacked hidden states of decoder for input to the last dense layer
decoder_combined_context = concatenate([context, decoder_stack_h])
# Dense layer with repeated weights
out = TimeDistributed(Dense(output_train.shape[2]))(decoder_combined_context)
# Compile model
model = Model(
inputs=input_train,
outputs=out)
opt = Adam(
learning_rate=0.001,
clipnorm=1)
model.compile(loss='mean_squared_error', optimizer=opt, metrics=['mae'])
print(model)
return model
# + executionInfo={"elapsed": 28, "status": "ok", "timestamp": 1653394799167, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="LIgKaIn-dNK8"
# Fit model
def fit_model(model, samples):
epc = 500
es = EarlyStopping(
monitor='val_loss',
mode='min',
patience=50,
restore_best_weights=True)
history = model.fit(
samples["X_input_train"],
samples["y_output_train"],
validation_data=(
samples["X_input_vali"],
samples["y_output_vali"]),
epochs=epc,
verbose=1,
callbacks=[es],
batch_size=64,
shuffle=True)
return model, history
# + colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"elapsed": 29, "status": "ok", "timestamp": 1653394799168, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="pEB0YsnAKyFo" outputId="8e4bde8d-cf44-48d9-e065-12e2b77e7908"
''' TODO: Try RMSE und MAE ; trial = 1'''
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3677953, "status": "ok", "timestamp": 1653415279900, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="8CpFzMHZs_vx" outputId="1c07e4c2-0567-41e1-cd43-52fbe55de85e"
# Execute over all conditions for n trials
samples_all = {}
model_all = {}
history_all = {}
results_all = {}
predictions_all = {}
data_all = {}
for x in trial: data_all[x] = {} # prepare nested dictionary
# iterate over trials
for j in range(len(trial)):
seed_value(j) # control random number generator for each trial
# iterate over conditions
for i in range(len(condition)):
print(f"Trial: {trial[j]}, Condition: {condition[i]}")
samples = sample_preperation(condition[i])
samples_all[condition[i]] = samples
# use lstm_decoder_encoder_luong_attention() for model with attention
model = lstm_decoder_encoder(samples)
model, history = fit_model(model, samples)
model_path = f"model/model_{trial[j]}_{condition[i]}.h5"
# extra path because models aren't pickable in tensorflow < 2.7 and can't be imported via pickle
model_all[condition[i]] = model_path
model.save(model_path) # -> using model.save() instead
history_all[condition[i]] = history.history
results = model.evaluate(
samples["X_input_test"],
samples["y_output_test"],
batch_size=64)
results_all[condition[i]] = results
predictions = model.predict(
samples["X_input_test"],
batch_size=64)
predictions_all[condition[i]] = predictions
# deepcopy to prevent reusing the same dictionary
samples_all_copy = deepcopy(samples_all) # (samples are the same in each trial)
model_all_copy = deepcopy(model_all)
history_all_copy = deepcopy(history_all)
results_all_copy = deepcopy(results_all)
predictions_all_copy = deepcopy(predictions_all)
# create nested dictionary for everything
data_all[trial[j]]["samples_all"] = samples_all_copy
data_all[trial[j]]["model_all"] = model_all_copy
data_all[trial[j]]["history_all"] = history_all_copy
data_all[trial[j]]["results_all"] = results_all_copy
data_all[trial[j]]["predictions_all"] = predictions_all_copy
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 9, "status": "ok", "timestamp": 1653415279913, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="7GU9Yq_0s_vz" outputId="ae35bb8b-3500-4321-90c9-bfe7ee60d35d"
for x in trial: print(x,data_all[x]["results_all"])
# + executionInfo={"elapsed": 222, "status": "ok", "timestamp": 1653415280134, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="0S8nvji_KyFy"
# save dictionary as pickle file
with open('model/data_all.pickle', 'wb') as handle:
pickle.dump(data_all, handle, protocol=pickle.HIGHEST_PROTOCOL)
# + colab={"base_uri": "https://localhost:8080/", "height": 17} executionInfo={"elapsed": 5, "status": "ok", "timestamp": 1653415280135, "user": {"displayName": "<NAME>", "userId": "08013796147118011125"}, "user_tz": -120} id="dM2voI-SlBhC" outputId="c0f918c4-13a4-4829-acab-bc565c1ffd2f"
if google_colab == 1:
from google.colab import files
files.download('model/data_all.pickle')
for x in list(data_all.keys()):
for y in list(data_all[x]["model_all"].keys()):
files.download((data_all[x]["model_all"][y]))
| 02_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# # Wavefield Separation FWI sensitivity kernel
# We compare sensitivity kernels in a half-space model with wavefield separation imaging conditions. Wavefield separation in the imaging condition can both enhance long wavelength FWI gradients (for FWI), and remove long wavelength backscattered energy (for RTM) and is implemented as a wavenumber filter in Kz following Faqi et. al. (2011).
#
# ### Workflow for the standard FWI imaging condition
# The standard imaging condition will have short wavelengths in the FWI image, and also long wavelengths in the RTM image.
# 1. Generate forward modeled data in true model
# 1. Generate forward modeled data in perturbed model
# 1. Backproject the residual
# 1. At every time step
# * Zero lag correlation of forward source wavefield `S` with backward receiver wavefield `R` at every point in the model: <br>
# `Image += S * R`
#
# ### Workflow for the wavefield separation FWI imaging condition
# This will enhance long wavelengths in the image.
# 1. Generate forward modeled data in true model
# 1. Generate forward modeled data in perturbed model
# 1. Backproject the residual
# 1. At every time step
# * Spatial Fourier transform the source and receiver wavefields in depth (2 FFTs) --> `ZS`, `ZR`
# * Apply diagonal operators to zero positive and negative frequencies to both `ZS` and `ZR`, generating `ZSup`, `ZSdn`, `ZRup`, and `ZRdn`
# * Inverse spatial Fourier transform in depth (4 FFTs) to generate `Sup`, `Sdn`, `Rup`, `Rdn`
# * Zero lag correlation at every point in the model: <br>
# `Image += Sup * Rup + Sdn * Rdn`
#
# ### Workflow for the wavefield separation RTM imaging condition
# This will remove long wavelength backscattered energy from the image.
# 1. Generate forward modeled data in true model
# 1. Generate forward modeled data in water velocity model
# 1. Subtracting water velocity data from true data to remove direct arrival
# 1. Backproject the direct arrival removed true data
# 1. At every time step
# * Spatial Fourier transform the source and receiver wavefields in depth (2 FFTs) --> `ZS`, `ZR`
# * Apply diagonal operators to zero positive and negative frequencies to both `ZS` and `ZR`, generating `ZSup`, `ZSdn`, `ZRup`, and `ZRdn`
# * Inverse spatial Fourier transform in depth (4 FFTs) to generate `Sup`, `Sdn`, `Rup`, `Rdn`
# * Zero lag correlation at every point in the model: <br>
# `Image += Sup * Rdn + Sdn * Rup`
#
# ### Note on conjugate symmetry
# We can simplify further and only require two inverse Fourier transforms for each time step by following Faqi et. al. (2011). Please see the discussion between equation 9 and 10, this is what is implmented in `JetPackWaveFD`.
#
# ### Reference
# * *An effective imaging condition for reverse-time migration using wavefield decomposition* 2011<br>
# <NAME>, <NAME>, <NAME>, and <NAME>
# https://library.seg.org/doi/full/10.1190/1.3533914
# + tags=[]
using PyPlot, Jets, WaveFD, JetPackWaveFD, Random, LinearAlgebra
# -
# ## Make three models
# * true halfspace model with velocities 1500 m/s over 2000 m/s
# * perturbed halfspace model with velocities 1505 m/s over 2000 m/s
# * water velocity model at 1500 m/s
#
# We make the perturbed model slightly faster in the upper half-space, which indicates that the gradients for FWI should be all positive.
# +
nz,nx = 150,600
dz,dx = 20.0,20.0
z2 = 101
vtrue = 1500 .* ones(Float32,nz,nx);
vtrue[z2:end,:] .= 2000;
vpert = 1510 .* ones(Float32,nz,nx);
vpert[z2:end,:] .= 2000;
vwater = 1500 .* ones(Float32,nz,nx);
# -
figure(figsize=(10,10))
vmin,vmax = extrema(vtrue)
subplot(2,2,1);imshow(vtrue,aspect="auto"); title("True model"); clim(vmin,vmax);
colorbar(orientation="horizontal")
subplot(2,2,2);imshow(vpert,aspect="auto"); title("Perturbed model"); clim(vmin,vmax)
colorbar(orientation="horizontal");
subplot(2,2,3);imshow(vwater,aspect="auto"); title("Water model"); clim(vmin,vmax)
colorbar(orientation="horizontal");
# ## Note on scratch space for temporary files
# When dealing with serialized nonlinear wavefields as in this example, we need to specify the location where scratch files will be written.
#
# You may need to change this to point to a temporary directory available on your system.
scratch = "/mnt/scratch"
@assert isdir(scratch)
# ## Make three operators
# * Standard imaging condition
# * FWI imaging condition
# * RTM imaging condition
# +
ntrec = 2001
dtrec = 0.004
dtmod = 0.002
sz = dz * 1;
sx = dx * div(nx,2);
rz = dz .* ones(Float32, nx);
rx = dx .* Float32[0:(nx-1);];
compscale = 1e-1
Fstd = JopNlProp2DAcoIsoDenQ_DEO2_FDTD(; b = ones(Float32,size(vtrue)),
comptype = UInt32, compscale = compscale,
nthreads = Sys.CPU_THREADS, isinterior = true, nsponge = 80,
ntrec = ntrec, dtrec = dtrec, dtmod = dtmod, dz = dz, dx = dx,
wavelet = WaveletCausalRicker(f=5), sx = sx, sz = sz, rx = rx, rz = rz,
srcfieldfile = joinpath(scratch, "field-$(randstring()).bin"),
reportinterval=0, imgcondition = "standard");
Ffwi = JopNlProp2DAcoIsoDenQ_DEO2_FDTD(; b = ones(Float32,size(vtrue)),
comptype = UInt32, compscale = compscale,
nthreads = Sys.CPU_THREADS, isinterior = true, nsponge = 80,
ntrec = ntrec, dtrec = dtrec, dtmod = dtmod, dz = dz, dx = dx,
wavelet = WaveletCausalRicker(f=5), sx = sx, sz = sz, rx = rx, rz = rz,
srcfieldfile = joinpath(scratch, "field-$(randstring()).bin"),
reportinterval=0, imgcondition = "FWI");
Frtm = JopNlProp2DAcoIsoDenQ_DEO2_FDTD(; b = ones(Float32,size(vtrue)),
comptype = UInt32, compscale = compscale,
nthreads = Sys.CPU_THREADS, isinterior = true, nsponge = 80,
ntrec = ntrec, dtrec = dtrec, dtmod = dtmod, dz = dz, dx = dx,
wavelet = WaveletCausalRicker(f=5), sx = sx, sz = sz, rx = rx, rz = rz,
srcfieldfile = joinpath(scratch, "field-$(randstring()).bin"),
reportinterval=0, imgcondition = "RTM");
# -
# ## Forward modeling
# Generate data to be backprojected:
# * FWI: backproject (d_pert - d_true)
# * RTM: backproject (d_true - d_water)
# + tags=[]
dtrue = Fstd*vtrue;
dpert = Fstd*vpert;
dwater = Fstd*vwater;
rfwi = dpert .- dtrue
rrtm = dtrue .- dwater
@show extrema(dtrue)
@show extrema(dpert)
@show extrema(dwater)
@show extrema(rfwi)
@show extrema(rrtm)
nothing
# -
# ## Plot the generated data
# +
scale = 2.5 / sqrt(norm(dtrue)^2 / length(dtrue))
@show scale
figure(figsize=(6,7)); clf()
subplot(3,2,1); imshow(scale .* dtrue,cmap="gray",aspect="auto",clim=[-1,+1]);
title("True data")
subplot(3,2,2); imshow(scale .* rfwi,cmap="gray",aspect="auto",clim=[-1,+1]);
title("FWI residual data")
subplot(3,2,3); imshow(scale .* dpert,cmap="gray",aspect="auto",clim=[-1,+1]);
title("Perturbed data")
subplot(3,2,4); imshow(scale .* rrtm,cmap="gray",aspect="auto",clim=[-1,+1]);
title("RTM residual data")
subplot(3,2,5); imshow(scale .* dwater,cmap="gray",aspect="auto",clim=[-1,+1]);
title("Water data")
tight_layout()
nothing
# -
# ## Apply the three imaging conditions
# Note we run the three forward modeling operators on the perturbed velocity in order to accurately time the application of the Jacobian.
# + tags=[]
Jstd = jacobian!(Fstd, vpert)
Jfwi = jacobian!(Ffwi, vpert)
Jrtm = jacobian!(Frtm, vpert)
dfwi1 = Fstd * vpert;
dfwi2 = Ffwi * vpert;
drtm2 = Frtm * vpert;
tfwi1 = @elapsed fwi1 = Jstd' * rfwi;
tfwi2 = @elapsed fwi2 = Jfwi' * rfwi;
trtm1 = @elapsed rtm1 = Jstd' * rrtm;
trtm2 = @elapsed rtm2 = Jrtm' * rrtm;
@show extrema(fwi1)
@show extrema(fwi2)
@show extrema(rtm1)
@show extrema(rtm2)
@info "FWI timing std=$(tfwi1) fwi=$(tfwi2) ratio=$(tfwi2/tfwi1)"
@info "RTM timing std=$(trtm1) fwi=$(trtm2) ratio=$(trtm2/trtm1)"
nothing
# -
# ## Plot the results
# * Because in the perturbed model the upper half-space is faster, we expect the gradient to be positive (red).
# * Note how the FWI wavefield separation enhances long wavelengths
# * Note how the RTM wavefield separation removes long wavelength backscattered energy
# + tags=[]
figure(figsize=(10,10))
scale_fwi1 = 0.5 / maximum(abs, fwi1[z2-10:z2+10,:])
scale_fwi2 = 0.5 / maximum(abs, fwi2[z2-10:z2+10,:])
scale_rtm1 = 0.5 / maximum(abs, rtm1[z2-10:z2+10,:])
scale_rtm2 = 0.5 / maximum(abs, rtm2[z2-10:z2+10,:])
@show scale_fwi1,scale_fwi2
@show scale_rtm1,scale_rtm2
subplot(2,2,1); imshow(fwi1.*scale_fwi1,aspect="auto",cmap="seismic",clim=[-1,1]);
title("Standard FWI IC");colorbar(orientation="horizontal")
subplot(2,2,2); imshow(fwi2.*scale_fwi2,aspect="auto",cmap="seismic",clim=[-1,1]);
title("Wavefield Separation FWI IC");colorbar(orientation="horizontal")
subplot(2,2,3); imshow(rtm1.*scale_rtm1,aspect="auto",cmap="seismic",clim=[-1,1]);
title("Standard RTM IC");colorbar(orientation="horizontal")
subplot(2,2,4); imshow(rtm2.*scale_rtm2,aspect="auto",cmap="seismic",clim=[-1,1]);
title("Wavefield Separation RTM IC");colorbar(orientation="horizontal")
nothing
# -
| 40_sensitivity/11_WavefieldSeparationSensitivity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hello bot!
# Open in [Colaboratory](https://colab.research.google.com/github/deepmipt/DeepPavlov/blob/master/docs/intro/hello_bot.ipynb)
# !pip install -q deeppavlov
# Import key components to build HelloBot.
from deeppavlov.skills.pattern_matching_skill import PatternMatchingSkill
from deeppavlov.agents.default_agent.default_agent import DefaultAgent
from deeppavlov.agents.processors.highest_confidence_selector import HighestConfidenceSelector
# Create skills as pre-defined responses for a user's input containing specific keywords and regular expressions. Every skill returns response and confidence.
hello = PatternMatchingSkill(responses=['Hello world!'], patterns=["hi", "hello", "good day"])
bye = PatternMatchingSkill(['Goodbye world!', 'See you around'],
patterns=["bye", "chao", "see you"])
fallback = PatternMatchingSkill(["I don't understand, sorry", 'I can say "Hello world!"'])
# Agent executes skills and then takes response from the skill with the highest confidence.
agent = DefaultAgent([hello, bye, fallback], skills_selector=HighestConfidenceSelector())
# Give the floor to the HelloBot!
agent(['Hello', 'Bye', 'Or not'])
| docs/intro/hello_bot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# .. _tut_stats_cluster_sensor_1samp_tfr:
#
# # Non-parametric 1 sample cluster statistic on single trial power
#
#
# This script shows how to estimate significant clusters
# in time-frequency power estimates. It uses a non-parametric
# statistical procedure based on permutations and cluster
# level statistics.
#
# The procedure consists in:
#
# - extracting epochs
# - compute single trial power estimates
# - baseline line correct the power estimates (power ratios)
# - compute stats to see if ratio deviates from 1.
#
#
#
# +
# Authors: <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne import io
from mne.time_frequency import single_trial_power
from mne.stats import permutation_cluster_1samp_test
from mne.datasets import sample
print(__doc__)
# -
# Set parameters
# --------------
#
#
# +
data_path = sample.data_path()
raw_fname = data_path + '/MEG/sample/sample_audvis_raw.fif'
event_id = 1
tmin = -0.3
tmax = 0.6
# Setup for reading the raw data
raw = io.read_raw_fif(raw_fname)
events = mne.find_events(raw, stim_channel='STI 014')
include = []
raw.info['bads'] += ['MEG 2443', 'EEG 053'] # bads + 2 more
# picks MEG gradiometers
picks = mne.pick_types(raw.info, meg='grad', eeg=False, eog=True,
stim=False, include=include, exclude='bads')
# Load condition 1
event_id = 1
epochs = mne.Epochs(raw, events, event_id, tmin, tmax, picks=picks,
baseline=(None, 0), reject=dict(grad=4000e-13, eog=150e-6))
data = epochs.get_data() # as 3D matrix
data *= 1e13 # change unit to fT / cm
# Time vector
times = 1e3 * epochs.times # change unit to ms
# Take only one channel
ch_name = raw.info['ch_names'][97]
data = data[:, 97:98, :]
evoked_data = np.mean(data, 0)
# data -= evoked_data[None,:,:] # remove evoked component
# evoked_data = np.mean(data, 0)
# Factor to down-sample the temporal dimension of the PSD computed by
# single_trial_power. Decimation occurs after frequency decomposition and can
# be used to reduce memory usage (and possibly computational time of downstream
# operations such as nonparametric statistics) if you don't need high
# spectrotemporal resolution.
decim = 5
frequencies = np.arange(8, 40, 2) # define frequencies of interest
sfreq = raw.info['sfreq'] # sampling in Hz
epochs_power = single_trial_power(data, sfreq=sfreq, frequencies=frequencies,
n_cycles=4, n_jobs=1,
baseline=(-100, 0), times=times,
baseline_mode='ratio', decim=decim)
# Crop in time to keep only what is between 0 and 400 ms
time_mask = (times > 0) & (times < 400)
evoked_data = evoked_data[:, time_mask]
times = times[time_mask]
# The time vector reflects the original time points, not the decimated time
# points returned by single trial power. Be sure to decimate the time mask
# appropriately.
epochs_power = epochs_power[..., time_mask[::decim]]
epochs_power = epochs_power[:, 0, :, :]
epochs_power = np.log10(epochs_power) # take log of ratio
# under the null hypothesis epochs_power should be now be 0
# -
# Compute statistic
# -----------------
#
#
threshold = 2.5
T_obs, clusters, cluster_p_values, H0 = \
permutation_cluster_1samp_test(epochs_power, n_permutations=100,
threshold=threshold, tail=0)
# View time-frequency plots
# -------------------------
#
#
# +
plt.clf()
plt.subplots_adjust(0.12, 0.08, 0.96, 0.94, 0.2, 0.43)
plt.subplot(2, 1, 1)
plt.plot(times, evoked_data.T)
plt.title('Evoked response (%s)' % ch_name)
plt.xlabel('time (ms)')
plt.ylabel('Magnetic Field (fT/cm)')
plt.xlim(times[0], times[-1])
plt.ylim(-100, 250)
plt.subplot(2, 1, 2)
# Create new stats image with only significant clusters
T_obs_plot = np.nan * np.ones_like(T_obs)
for c, p_val in zip(clusters, cluster_p_values):
if p_val <= 0.05:
T_obs_plot[c] = T_obs[c]
vmax = np.max(np.abs(T_obs))
vmin = -vmax
plt.imshow(T_obs, cmap=plt.cm.gray,
extent=[times[0], times[-1], frequencies[0], frequencies[-1]],
aspect='auto', origin='lower', vmin=vmin, vmax=vmax)
plt.imshow(T_obs_plot, cmap=plt.cm.RdBu_r,
extent=[times[0], times[-1], frequencies[0], frequencies[-1]],
aspect='auto', origin='lower', vmin=vmin, vmax=vmax)
plt.colorbar()
plt.xlabel('time (ms)')
plt.ylabel('Frequency (Hz)')
plt.title('Induced power (%s)' % ch_name)
plt.show()
| 0.12/_downloads/plot_stats_cluster_1samp_test_time_frequency.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAZ Diplomarbeit - Biodiversität in der Schweiz#
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
df = pd.read_csv("01_source/xls_ch/eintags_stein_koecherfliegenarten_ch.csv")
df.head()
len (df)
df.shape
df["CAT"].value_counts()
df.head(1)
df.rename(columns={"GROUP":"group","ORDER":"order","FAMILY":"family","GENUS":"genus","SPECIES":"species","Scientific name":"sc name","Deutscher Name":"dt name","CAT":"cat", "Criteria": "criteria", "Bemerkungen":"bemerkungen"},inplace=True)
df
df.pop("criteria")
df.pop("genus")
df.pop("species")
df
df['einteilung'] = 'Insekten'
df['untergruppe'] = 'Eintags,-Stein-,Köcherfliegen'
df.head(5)
df["cat"].value_counts()
# +
values = [0, 27, 51, 68, 81, 71, 186, 15, 8]
colors = ['r', 'r', 'r', 'r', 'r', 'g', "g", "grey", "grey"]
labels = [
'EX: ausgestorben', 'RE: CH ausgestorben',
'CR: v. aussterben bedroht', 'EN: stark gefährdet',
'VU: verletzlich',"NT: potenziell gef.",
"LC: nicht gef.","DD: ungen. Daten","NE: nicht beurteilt"]
fig1, ax1 = plt.subplots(figsize=(7, 6))
fig1.subplots_adjust(0.3,0,1,1)
plt.pie(values, colors=colors, labels= values, counterclock=False,startangle=90)
plt.title('Gefährdung Eintags-, Stein- u. Köcherfliegen, 507 Arten')
total = sum(values)
plt.legend(
loc='upper left',
labels=['%s, %1.1f%%' % (
l, (float(s) / total) * 100) for l, s in zip(labels, values)],
bbox_to_anchor=(1.0, 0.5),
bbox_transform=fig1.transFigure
)
plt.show()
# -
| Eigene Projekte/Diplomarbeit_Biodiversitaet/001_einlesen_csv_eintags_stein_koechefliegen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LeNet Lab Solution
# 
# Source: <NAME>
# ## Load Data
#
# Load the MNIST data, which comes pre-loaded with TensorFlow.
#
# You do not need to modify this section.
# +
# Load pickled data# Load
import pickle
# TODO: Fill this in based on where you saved the training and testing data
training_file = 'train.p'
validation_file= 'valid.p'
testing_file = 'test.p'
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# -
# The MNIST data that TensorFlow pre-loads comes as 28x28x1 images.
#
# However, the LeNet architecture only accepts 32x32xC images, where C is the number of color channels.
#
# In order to reformat the MNIST data into a shape that LeNet will accept, we pad the data with two rows of zeros on the top and bottom, and two columns of zeros on the left and right (28+2+2 = 32).
#
# You do not need to modify this section.
# ## Visualize Data
#
# View a sample from the dataset.
#
# You do not need to modify this section.
# +
import random
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
index = random.randint(0, len(X_train))
image = X_train[index].squeeze()
plt.figure(figsize=(1,1))
plt.imshow(image)
print(y_train[index])
# +
from sklearn.utils import shuffle
# -
# ## Setup TensorFlow
# The `EPOCH` and `BATCH_SIZE` values affect the training speed and model accuracy.
#
# You do not need to modify this section.
# +
import tensorflow as tf
EPOCHS = 10
BATCH_SIZE = 256
# -
# ## SOLUTION: Implement LeNet-5
# Implement the [LeNet-5](http://yann.lecun.com/exdb/lenet/) neural network architecture.
#
# This is the only cell you need to edit.
# ### Input
# The LeNet architecture accepts a 32x32xC image as input, where C is the number of color channels. Since MNIST images are grayscale, C is 1 in this case.
#
# ### Architecture
# **Layer 1: Convolutional.** The output shape should be 28x28x6.
#
# **Activation.** Your choice of activation function.
#
# **Pooling.** The output shape should be 14x14x6.
#
# **Layer 2: Convolutional.** The output shape should be 10x10x16.
#
# **Activation.** Your choice of activation function.
#
# **Pooling.** The output shape should be 5x5x16.
#
# **Flatten.** Flatten the output shape of the final pooling layer such that it's 1D instead of 3D. The easiest way to do is by using `tf.contrib.layers.flatten`, which is already imported for you.
#
# **Layer 3: Fully Connected.** This should have 120 outputs.
#
# **Activation.** Your choice of activation function.
#
# **Layer 4: Fully Connected.** This should have 84 outputs.
#
# **Activation.** Your choice of activation function.
#
# **Layer 5: Fully Connected (Logits).** This should have 10 outputs.
#
# ### Output
# Return the result of the 2nd fully connected layer.
# +
from tensorflow.contrib.layers import flatten
def LeNet(x):
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
# SOLUTION: Layer 1: Convolutional. Input = 32x32x1. Output = 28x28x6.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 3, 6), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
# SOLUTION: Activation.
conv1 = tf.nn.relu(conv1)
# SOLUTION: Pooling. Input = 28x28x6. Output = 14x14x6.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# SOLUTION: Layer 2: Convolutional. Output = 10x10x16.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
# SOLUTION: Activation.
conv2 = tf.nn.relu(conv2)
# SOLUTION: Pooling. Input = 10x10x16. Output = 5x5x16.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# SOLUTION: Flatten. Input = 5x5x16. Output = 400.
fc0 = flatten(conv2)
# SOLUTION: Layer 3: Fully Connected. Input = 400. Output = 120.
fc1_W = tf.Variable(tf.truncated_normal(shape=(400, 120), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(120))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# SOLUTION: Activation.
fc1 = tf.nn.relu(fc1)
# SOLUTION: Layer 4: Fully Connected. Input = 120. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape=(120, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# SOLUTION: Activation.
fc2 = tf.nn.relu(fc2)
# SOLUTION: Layer 5: Fully Connected. Input = 84. Output = 10.
fc3_W = tf.Variable(tf.truncated_normal(shape=(84, 43), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(43))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
# -
# ## Features and Labels
# Train LeNet to classify [MNIST](http://yann.lecun.com/exdb/mnist/) data.
#
# `x` is a placeholder for a batch of input images.
# `y` is a placeholder for a batch of output labels.
#
# You do not need to modify this section.
x = tf.placeholder(tf.float32, (None, 32, 32, 3))
y = tf.placeholder(tf.int32, (None))
one_hot_y = tf.one_hot(y, 43)
# ## Training Pipeline
# Create a training pipeline that uses the model to classify MNIST data.
#
# You do not need to modify this section.
# +
rate = 0.001
logits = LeNet(x)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y, logits=logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation)
# -
# ## Model Evaluation
# Evaluate how well the loss and accuracy of the model for a given dataset.
#
# You do not need to modify this section.
# +
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
# -
# ## Train the Model
# Run the training data through the training pipeline to train the model.
#
# Before each epoch, shuffle the training set.
#
# After each epoch, measure the loss and accuracy of the validation set.
#
# Save the model after training.
#
# You do not need to modify this section.
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y})
validation_accuracy = evaluate(X_validation, y_validation)
print("EPOCH {} ...".format(i+1))
print("Validation Accuracy = {:.3f}".format(validation_accuracy))
print()
saver.save(sess, './lenet')
print("Model saved")
# ## Evaluate the Model
# Once you are completely satisfied with your model, evaluate the performance of the model on the test set.
#
# Be sure to only do this once!
#
# If you were to measure the performance of your trained model on the test set, then improve your model, and then measure the performance of your model on the test set again, that would invalidate your test results. You wouldn't get a true measure of how well your model would perform against real data.
#
# You do not need to modify this section.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
| LeNet-Lab-Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import SQL Alchemy
from sqlalchemy import create_engine
# Import and establish Base for which classes will be constructed
from sqlalchemy.ext.declarative import declarative_base
Base = declarative_base()
# Import modules to declare columns and column data types
from sqlalchemy import Column, Integer, String, Float
# +
# Create Surfer and Board classes
# ----------------------------------
class Surfer(Base):
__tablename__ = 'surfers'
id = Column(Integer, primary_key=True)
name = Column(String(255))
hometown = Column(String(255))
wipeouts = Column(Integer)
rank = Column(Integer)
class Board(Base):
__tablename__ = 'surfboards'
id = Column(Integer, primary_key=True)
surfer_id = Column(Integer)
board_name = Column(String(255))
color = Column(String(255))
length = Column(Integer)
# -
# Create specific instances of the Surfer and Board classes
surfer=Surfer(1,"<NAME>","S"
# ----------------------------------
# Create a new surfer named "Bruno"
# Create a new board and associate it with a surfer's ID
# Create Database Connection
# ----------------------------------
# Establish Connection to a sqlite database
# Create both the Surfer and Board tables within the database
# To push the objects made and query the server we use a Session object
# Add "Bruno" to the current session
# Add "Awwwyeah" to the current session
# Commit both objects to the database
# Query the database and collect all of the surfers in the Surfer table
| 10-Advanced-Data-Storage-and-Retrieval/1/Activities/11-Stu_Surfer_SQL/Unsolved/.ipynb_checkpoints/Surfer_SQL-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# Fill in any place that says `# YOUR CODE HERE` or YOUR ANSWER HERE, as well as your name and collaborators below.
# Grading for pre-lecture assignments is all or nothing. Partial credit is available for in-class assignments and checkpoints, but **only when code is commented**.
# -
NAME = ""
COLLABORATORS = ""
# ---
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "c78e6643854ee561a78f24f5f9af1078", "grade": false, "grade_id": "cell-ec0c8f83ffb0d9c7", "locked": true, "schema_version": 3, "solution": false}
# # Learning Objectives
#
# This lecture will show you how to:
# 1. Apply Gaussian elimination to solve simultaneous equations
# 2. Implement partial pivoting to extend Gaussian elimination
# 3. Use `scipy.linalg.solve`
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "d98d39327803ae41271792ef226900dd", "grade": false, "grade_id": "cell-abd1b2cca923116d", "locked": true, "schema_version": 3, "solution": false}
# imports
import numpy as np
import matplotlib.pyplot as plt
from scipy import linalg # linear algebra
import grading_helper as _test
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "17d45a532e5db426da0bcd8a980838a8", "grade": false, "grade_id": "cell-12b43295a0f000dd", "locked": true, "schema_version": 3, "solution": false}
# # Gaussian Elimination
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "edf55d5e830a2407baa66d30ba73acaa", "grade": false, "grade_id": "cell-e9692b2c91e495dc", "locked": true, "schema_version": 3, "solution": false}
# %video WIU55nqfKZo
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "4f607ab6ca19ee090d607add3a7af4ba", "grade": false, "grade_id": "cell-ef2de58511545032", "locked": true, "schema_version": 3, "solution": false}
# Summary:
#
# - Consider the matrix equation $\mathbf{A}\mathbf{x} = \mathbf{v}$. The rows of matrix $\mathbf{A}$ can be modified as long as the rows of $\mathbf{v}$ are modified in the same manner. We repeated apply this idea to our matrix equation to simplify it in a process called **Gaussian elimination**.
# - Our main tools are dividing a row by a constant, and subtracting multiples of a row from other rows. Eventually, we make $\mathbf{A}$ **upper triangular**. Solving is then trivial.
#
# -
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "18ac6bb8cad905f5ded0136b62f8a443", "grade": false, "grade_id": "cell-e97e11861c0dcf51", "locked": true, "schema_version": 3, "solution": false}
# ## Your Turn
#
# Complete the function `solve3x3` that solves three equations with three unknowns using Gaussian elimination. The inputs to the function are $\mathbf{A}$ and $\mathbf{v}$ (both implemented as arrays).
# + deletable=false nbgrader={"cell_type": "code", "checksum": "56d96b42e67e9fc2bc39a2b57f6a26d7", "grade": false, "grade_id": "cell-35ad44336baecb36", "locked": false, "schema_version": 3, "solution": true}
# %%graded # 3 points
def solve3x3(A, v):
# make copies of A and v
A, v = A.copy(), v.copy()
# divide 1st row by A[0,0]
A[0], v[0] = A[0]/A[0,0], v[0]/A[0,0]
# subtract A[1,0]*A[0] from A[1]
A[1], v[1] = A[1] - A[1,0]*A[0], v[1] - A[1,0]*v[0]
# subtract A[2,0]*A[0] from A[2]
A[2], v[2] = A[2] - A[2,0]*A[0], v[2] - A[2,0]*v[0]
# YOUR CODE HERE
return x, y, z
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "3b920fb912fb23439a068cf960305d30", "grade": true, "grade_id": "cell-da8b5d94fab34508", "locked": true, "points": 3, "schema_version": 3, "solution": false}
# %%tests
A = np.array([[4., 1., 2.],
[2., 2., 1.],
[1., 4., 1.]])
v = np.array([1.,
0.,
2.])
x, y, z = solve3x3(A,v)
_test.similar(x, -8/3)
_test.similar(y, -1/3)
_test.similar(z, 6)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "19b88d548f980a3a33f7a3fe9dc0a82d", "grade": false, "grade_id": "cell-c3979e64ecb0f78e", "locked": true, "schema_version": 3, "solution": false}
# # Partial Pivoting
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "2f73a7c92f4853a92804c90cc2b4ce6f", "grade": false, "grade_id": "cell-e55d933eef813232", "locked": true, "schema_version": 3, "solution": false}
# %video gU50Zp1zonw
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "41166fea83319a98e6a49bb0a5dcc3b0", "grade": false, "grade_id": "cell-3fc428fa4e7fa976", "locked": true, "schema_version": 3, "solution": false}
# Summary:
#
# - There are two issues we may encounter:
# 1. Some sets of equations don't have solutions (although that really isn't an issue in physics).
# 2. Our simple implementation of Gaussian elimination may have us divide by zero.
# - The solution to the second case is **partial pivoting**: before we do any division step, we look for the value with the greatest absolute value in the current column. We then swap the current row with that row.
# -
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "7209cffa50cafdd8acc8f281da732b92", "grade": false, "grade_id": "cell-816be27c799e9a5e", "locked": true, "schema_version": 3, "solution": false}
# # Using `scipy.linalg.solve`
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "4691bd7a7282342475ff6f6506362ba6", "grade": false, "grade_id": "cell-938a3475f0b78917", "locked": true, "schema_version": 3, "solution": false}
# %video clvL26_M_TU
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "423c0a15eecbd1e3074acb7d6b57419f", "grade": false, "grade_id": "cell-3b9c0f7fca8fa09f", "locked": true, "schema_version": 3, "solution": false}
# Summary:
#
# - If you look at our simple function for Gaussian elimination, there are some redundancies. In particular, our modifications to the matrices can be combined into fewer steps.
# - In brief, we follow a particular sequence of simplifications that lead to $\mathbf{LU}=\mathbf{A}$, called the **LU decomposition** of $\mathbf{A}$.
# - This technique, along with partial pivoting, is employed by `scipy.linalg.solve(A, v)`
# -
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "83b0f8a618983999c7d7fa6ea8743dae", "grade": false, "grade_id": "cell-ee75c910cc1dccc4", "locked": true, "schema_version": 3, "solution": false}
# ## Your Turn
#
# Use `linalg.solve` to solve this system of equations:
# $$u + v - 2w + x + 3y - z = 4$$
# $$2u - v + w + 2x + y - 3z = 20$$
# $$u + 3v - 3w - x + 2y + z = -15$$
# $$5u + 2v - w - x + 2y + z = - 3$$
# $$-3u - v + 2w + 3x + y + 3z = 16$$
# $$4u + 3v + w - 6x - 3y - 2z = -27$$
#
# Store your answers in the variables `u`, `v`, `w`, `x`, `y`, and `z`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "26f02434add65933f44fb71dc563c003", "grade": false, "grade_id": "cell-c85222addd195fd0", "locked": false, "schema_version": 3, "solution": true}
# %%graded # 2 points
# YOUR CODE HERE
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "69b5f4f8a40061c19742d542854c6292", "grade": true, "grade_id": "cell-66bb15c80b0dc98c", "locked": true, "points": 2, "schema_version": 3, "solution": false}
# %%tests
_test.code_contains("solve")
_test.similar(u, 1)
_test.similar(v, -2)
_test.similar(w, 3)
_test.similar(x, 4)
_test.similar(y, 2)
_test.similar(z, -1)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "56ed9a7b06f793ad47f204af025e8080", "grade": false, "grade_id": "cell-1ae60099cab3bd92", "locked": true, "schema_version": 3, "solution": false}
# # Additional Resources
#
# - Textbook section 6.1
| Assignments/07.1 Lecture - Linear Equations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/brunoalvoliv/analise-de-empresa/blob/main/carteira_invest3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-_N1qNHecAkY"
# ##Bibliotecas
# + colab={"base_uri": "https://localhost:8080/"} id="SWixDC_Yc26t" outputId="c0018a61-651c-453b-ef3a-5b60d33961a6"
# !pip install yfinance
# + id="bDE7cvQub414"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import yfinance as yf
# + id="pPHMPpfwc73i"
plt.style.use('ggplot')
# + [markdown] id="CkTxXImZcUx1"
# ##Retorno do ibovespa versus carteira
# + id="W7ymbgRH2yZE" colab={"base_uri": "https://localhost:8080/", "height": 463} outputId="6b113303-0f58-4411-95e5-3831ceabe541"
#Buscando os dados
tickers = ['PETR3.SA', 'MGLU3.SA', 'ITSA4.SA', 'RDOR3.SA', 'LCAM3.SA', 'BPAC11.SA',
'CSAN3.SA', 'SMFT3.SA', 'ARZZ3.SA', 'TRPL3.SA', 'B3SA3.SA', 'WEGE3.SA', '^BVSP']
carteira1 = pd.DataFrame()
for i in tickers:
carteira1[i] = yf.download(i, start='2021-11-01', end='2021-11-30')['Adj Close']
carteira1.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="84uiIeFsdVvF" outputId="54339d81-ce47-4002-b1a9-00a2a3faaced"
#Obtendo retorno simples
ret_simples = carteira1.pct_change()
ret_simples.fillna(0, inplace=True)
ret_simples.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="uKTbWmgnzv1t" outputId="aeb0fa24-bd29-44b3-be56-0bf32526e8c6"
#Obtendo o retorno acumulado e normalizando a base de dados
ret_acum = (1 + ret_simples).cumprod()
ret_acum.head()
# + colab={"base_uri": "https://localhost:8080/"} id="PcREyEIzgI-S" outputId="a4aa6e1a-5a2f-4c13-9da3-b01b8403ff62"
ret_simples.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="si8pqPWkoMkM" outputId="669acb0d-0416-4d77-c73c-462800f0da4b"
#Calculando a média diária do retorno simples
mean_retsimples = ret_simples.mean(axis=1)
mean_retsimples = pd.DataFrame(mean_retsimples.head())
mean_retsimples.rename(columns={0:'Carteira'}, inplace=True)
mean_retsimples['Mercado'] = ret_simples['^BVSP']
mean_retsimples.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="rl0x2LsHEVpV" outputId="f2c64427-8b5d-448b-862e-fddafe5bac3d"
mean_retsimples.mean()
# + colab={"base_uri": "https://localhost:8080/", "height": 580} id="f9r3hRuQrf0r" outputId="ef1267f6-3c92-47b3-aa7e-e90013e93f8d"
#Visualização da média diária do retorno simples
plt.figure(figsize=(18, 9));
plt.title('Desempenho da carteira (NOV/2021)', fontsize=20);
x, = plt.plot(mean_retsimples.Carteira, marker='^', color='blue', linewidth=2.5, label='Carteira');
y, = plt.plot(mean_retsimples.Mercado, marker='o', color='orange', linewidth=2.5, label='IBOVESPA');
plt.xlabel('Data', fontsize=17);
plt.ylabel('Retorno simples', fontsize=17);
plt.text(np.datetime64('2021-11-05'), -0.017, 'By: Invest Jr. - Yahoo Finance', fontsize=14);
plt.legend(fontsize=15, loc=2);
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="pu4M178U0UoB" outputId="5b0214f4-c0a7-41c3-ebeb-492655e7a0b5"
#Calculando a média diária do retorno acumulado
mean_retacum = ret_acum.mean(axis=1)
mean_retacum = pd.DataFrame(mean_retacum.head())
mean_retacum.rename(columns={0:'Carteira'}, inplace=True)
mean_retacum['Mercado'] = ret_acum['^BVSP']
mean_retacum.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="Xi-jdCbaEjh0" outputId="37f3dc96-6739-49d1-9945-3bcd98cad239"
mean_retacum.mean()
# + colab={"base_uri": "https://localhost:8080/", "height": 580} id="M6Tcuz-N0QgD" outputId="278ae935-dca1-4c96-dcde-5d7507fe0699"
#Visualização da média diária do retorno acumulado
plt.figure(figsize=(18, 9));
plt.title('Desempenho acumulado da carteira (NOV/2021)', fontsize=20);
x, = plt.plot(mean_retacum.Carteira, marker='^', color='blue', linewidth=2.5, label='Carteira');
y, = plt.plot(mean_retacum.Mercado, marker='o', color='orange', linewidth=2.5, label='IBOVESPA');
plt.xlabel('Data', fontsize=17);
plt.ylabel('Retorno acumulado', fontsize=17);
plt.text(np.datetime64('2021-11-05'), 0.983, 'By: Invest Jr. - Yahoo Finance', fontsize=14);
plt.legend(fontsize=15, loc=2);
# + [markdown] id="ZhuMriJLlwrF"
# Criando a tabela de desempenho
# + colab={"base_uri": "https://localhost:8080/"} id="VIDwoJlijWfa" outputId="6f955a40-8b6e-4917-e76f-623b5053e23c"
preco_inicial = np.round(carteira1.iloc[:1, :12].values.T, 2)
preco_inicio = []
for i in preco_inicial:
preco_inicio.append(f'R$ {str(i[0])}')
preco_inicio
# + colab={"base_uri": "https://localhost:8080/"} id="2xD7UGE7k88Z" outputId="197f57ab-6305-433b-999a-447244e42e32"
preco_final = np.round(carteira1.iloc[-1:, :12].values.T, 2)
preco_fim = []
for i in preco_final:
preco_fim.append(f'R$ {i[0]}')
preco_fim
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="WNzssShXvE4n" outputId="be005017-005d-499e-a29e-a1756122e3a6"
#1º passo: obter o retorno simples da carteira
ret_simples.head().iloc[1:, :12]
# + colab={"base_uri": "https://localhost:8080/"} id="uo0EVN76vt2Z" outputId="c9b296d1-80eb-498b-fd74-26b3f0891b37"
#2º passo: calcular a média dos retornos esperados
#mean_carteira = np.round(np.mean(retorno_carteira.iloc[1:, :12]) * 100, 2)
mean_carteira = ret_simples.iloc[1:, :12].mean()
mean_carteira
# + colab={"base_uri": "https://localhost:8080/", "height": 426} id="N-7uYR4bwQPG" outputId="85d068dd-791c-4ceb-9478-38f3411a8a53"
#3º passo: calcular a covariância da carteira
cov_carteira = ret_simples.head().iloc[1:, :12].cov()
cov_carteira
# + colab={"base_uri": "https://localhost:8080/"} id="mPZdmb5pzQn0" outputId="675266cc-6b23-47f4-9ab0-857e88c06aaf"
#4º passo: definir os pesos
pesos_carteira = np.array([0.091, 0.054, 0.073, 0.058, 0.077, 0.042, 0.098, 0.056,
0.053, 0.205, 0.036, 0.156])
pesos_carteira = np.round(pesos_carteira * 100, 2)
composicao = []
for i in pesos_carteira:
composicao.append(f'{i}%')
composicao
# + colab={"base_uri": "https://localhost:8080/"} id="Dtqv1Ic511_u" outputId="50c620fa-f2bf-4ef9-a146-76d0dfb982d0"
#5º passo: retornos individuais
retornos_individuais = (pesos_carteira * mean_carteira) * 100
retornos_individuais = np.round(retornos_individuais, 2)
retornos_individual = []
for i in retornos_individuais:
retornos_individual.append(f'{i}%')
retornos_individual
# + colab={"base_uri": "https://localhost:8080/"} id="NuAUD9480bq7" outputId="b5cb4c9c-f57e-49d8-8bf6-6b87e8149682"
#6º passo: retorno da carteira
retorno_carteira = np.sum(pesos_carteira * mean_carteira)
retorno_carteira
# + colab={"base_uri": "https://localhost:8080/"} id="P8RstLZZ1p9t" outputId="2dd3c3bc-4404-48a4-8410-bb80ca661333"
#7º risco da carteira
risco_carteira = np.sqrt(np.dot(pesos_carteira.T, np.dot(cov_carteira, pesos_carteira)))
risco_carteira
# + colab={"base_uri": "https://localhost:8080/"} id="bDLs3uxL0_MK" outputId="fa8f3729-6c8b-4e56-a19b-0dd6d89e4ad5"
ret_simples_mean = np.round(ret_simples.iloc[:, :12].head().mean() * 100, 2)
ret_ativo = []
for i in ret_simples_mean:
ret_ativo.append(f'{i}%')
ret_ativo
# + colab={"base_uri": "https://localhost:8080/", "height": 426} id="5VKS79Uwda9L" outputId="43b07763-279e-4fca-dd84-d1a1bb49d1f8"
#Tabela de desempenho
desempenho = pd.DataFrame(index=tickers[:12])
desempenho['Composição'] = composicao
desempenho['Preço Inicial'] = preco_inicio
desempenho['Preço Final'] = preco_fim
desempenho['Retorno do Ativo'] = ret_ativo
desempenho['Retorno na Carteira'] = retornos_individual
desempenho
# + id="wwDWHuScRTl6"
#desempenho.to_csv('desempenho_invest.csv', sep=',');
| carteira_invest3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Imports
import sp
from astropy.io import ascii
import time
import ccdproc as ccdp
from astropy.nddata import CCDData
import numpy as np
import glob
import itertools
from astropy.table import Table
from astropy import table
import Logs
from astropy.convolution import convolve, Box1DKernel
import matplotlib.pyplot as plt
# Photometry
# +
science=ccdp.ImageFileCollection('Final_Science/median')
# sciencelist=glob.glob('Corrected_Science/r*fit')
#ascii.write(sc,'sc_summary.csv',format='csv',overwrite=True)
#found=Table(names=('fname','x','y','sh','ro'),dtype=('str','float32','float32','float32','float32'))
trackerf=[]
trackera=[]
start=time.time()
for imsd,imsn in science.data(return_fname=True):
print('\nassessing',imsn)
try:
x,y,flux,sh,ro=sp.find(imsd,hmin=3000,fwhm=17)
if len(x) == 2:
for i in range(len(x)):
trackerf.append([imsn,x[i],y[i],sh[i],ro[i]])
flux,eflux,sky,skerr=sp.aper(imsd,x,y,phpadu=2.9,apr=[5],skyrad=[15,20],flux=True)
trackera.append([imsn,flux[0][0],flux[1][0],sky[0],sky[1],eflux[0][0],eflux[1][0]])
print(imsn,'flux calculated')
elif len(x) != 2:
print('skipping',imsn,'since it did not return exactly 2 star coordinates')
except Exception as e:
print(imsn,'did not meet sharpness or roundness criteria')
print(e)
tf=np.reshape(trackerf,(-1,5))
ta=np.reshape(trackera,(-1,7))
print(time.time()-start)
foundf=Table(tf,names=('fname','x','y','sh','ro'),dtype=('str','float64','float64','float64','float64'))
fluxt=Table(ta,names=('fname','fluxcalib','fluxtarget','skycalib','skytarget','fluxerrcalib','fluxerrtarget')
,dtype=('str','float64','float64','float64','float64','float64','float64'))
ascii.write(foundf,'stars.csv',format='csv',overwrite=True)
ascii.write(fluxt,'fluxes.csv',format='csv',overwrite=True)
# -
# Get logs
# +
log1=Logs.parse_logfile('HD115709/log_files/run_log_20180428.int')
log2=Logs.parse_logfile('HD115709/log_files/run_log_20180430.int')
log1t=Table(list(log1['Observations'].values()),names=tuple(log1['Observations'].keys()))
log2t=Table(list(log2['Observations'].values()),names=tuple(log2['Observations'].keys()))
# logc=table.vstack([log1t,log2t])
# -
# Data manipulation
# +
# fluxt=ascii.read('fluxes.csv')
c=[]
for i, val in enumerate(fluxt['fname']):
c.append(int(val[1:8]))
fluxt['Run']=c
final1=table.join(fluxt,log1t,keys='Run')
final2=table.join(fluxt,log2t,keys='Run')
final1['truefluxcalib']=final1['fluxcalib']/final1['Exptime']
final2['truefluxcalib']=final2['fluxcalib']/final2['Exptime']
final1['truefluxtarget']=final1['fluxtarget']/final1['Exptime']
final2['truefluxtarget']=final2['fluxtarget']/final2['Exptime']
final1['diff']=final1['truefluxcalib']-final1['truefluxtarget']
final2['diff']=final2['truefluxcalib']-final2['truefluxtarget']
final1['relflux']=final1['diff']/np.nanmedian(final1['diff'])
final2['relflux']=final2['diff']/np.nanmedian(final2['diff'])
# -
# Convolve
# +
final1['smadiff']=convolve(list(final1['relflux']),kernel=Box1DKernel(10),boundary='extend')
final2['smadiff']=convolve(list(final2['relflux']),kernel=Box1DKernel(10),boundary='extend')
ascii.write(final1,'final1.csv',overwrite=True,format='csv')
ascii.write(final2,'final2.csv',overwrite=True,format='csv')
# -
# Plot
fig, axs=plt.subplots(1,2,figsize=(40,10))
axs[0].scatter(final1['MJD']+2400000.5,final1['smadiff'])
axs[0].ticklabel_format(axis='x',useOffset=True,useLocale=True)
axs[1].scatter(final2['MJD']+2400000.5,final2['smadiff'])
axs[1].ticklabel_format(axis='x',useOffset=True,useLocale=True)
plt.show()
| .ipynb_checkpoints/PLot interm run-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/misbahsy/APMonitor-do/blob/master/NonlinearControl.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="JYQajzOVzjkl" colab_type="text"
# #Nonlinear Model Predictive Control
#
# <p class='vspace'>Dynamic control is also known as Nonlinear Model Predictive Control (NMPC) or simply as Nonlinear Control (NLC). NLC with predictive models is a dynamic optimization approach that seeks to follow a trajectory or drive certain values to maximum or minimum levels.
# </p>
# <div class='vspace'></div>
#
# + [markdown] id="i_CzzSVVX-Kf" colab_type="text"
#
# ##Exercise
#
# <p><strong>Objective:</strong> Design a controller to maintain temperature of a chemical reactor. Develop 3 separate controllers (PID, Linear MPC, Nonlinear MPC) in Python, MATLAB, or Simulink. Demonstrate controller performance with steps in the set point and disturbance changes. <em>Estimated time: 3 hours.</em>
# </p>
# <div class='vspace'></div><div><img src='http://apmonitor.com/do/uploads/Main/download.png' alt='' title='' /> <a class='urllink' href='http://apmonitor.com/do/uploads/Main/cstr_control.zip' rel='nofollow'>CSTR Source Files</a> | <a class='urllink' href='http://apmonitor.com/do/uploads/Main/cstr_control.pdf' rel='nofollow'>Problem Information</a></div>
# <div class='vspace'></div><div><img width='300px' src='http://apmonitor.com/do/uploads/Main/cstr.png' alt='' title='' /></div>
# <p class='vspace'>A reactor is used to convert a hazardous chemical <strong>A</strong> to an acceptable chemical <strong>B</strong> in waste stream before entering a nearby lake. This particular reactor is dynamically modeled as a Continuously Stirred Tank Reactor (CSTR) with a simplified kinetic mechanism that describes the conversion of reactant <strong>A</strong> to product <strong>B</strong> with an irreversible and exothermic reaction. It is desired to maintain the temperature at a constant setpoint that maximizes the destruction of A (highest possible temperature). Adjust the jacket temperature (<em>T<sub>c</sub></em>) to maintain a desired reactor temperature and minimize the concentration of <strong>A</strong>. The reactor temperature should never exceed 400 K. The cooling jacket temperature can be adjusted between 250 K and 350 K.
# </p>
#
# + [markdown] id="Hbit8iXpYJOF" colab_type="text"
#
# ##Step Testing
# <p>Step testing is required to obtain a process model for the PID controller and the linear model predictive controller. It is a first step in developing a controller. The following code implements either a doublet test or multiple steps to different levels. A doublet test starts with the system at steady state. Three moves of Manipulated Variable (MV) are made with sufficient time to nearly reach steady state conditions at two other operating points. The steps are above and below the nominal operating conditions. In this case, the cooling jacket temperature is raised, lowered, and brought back to 300 K (nominal operating condition.
# </p>
# <div class='vspace'></div><div><img width='550px' src='http://apmonitor.com/do/uploads/Main/cstr_doublet.png' alt='' title='' /></div>
# + id="WiYfb7NgzGO9" colab_type="code" cellView="form" colab={}
#@markdown Python Simulation Code
try:
from pip import main as pipmain
except:
from pip._internal import main as pipmain
pipmain(['install','gekko'])
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
# define CSTR model
def cstr(x,t,u,Tf,Caf):
# Inputs (3):
# Temperature of cooling jacket (K)
Tc = u
# Tf = Feed Temperature (K)
# Caf = Feed Concentration (mol/m^3)
# States (2):
# Concentration of A in CSTR (mol/m^3)
Ca = x[0]
# Temperature in CSTR (K)
T = x[1]
# Parameters:
# Volumetric Flowrate (m^3/sec)
q = 100
# Volume of CSTR (m^3)
V = 100
# Density of A-B Mixture (kg/m^3)
rho = 1000
# Heat capacity of A-B Mixture (J/kg-K)
Cp = 0.239
# Heat of reaction for A->B (J/mol)
mdelH = 5e4
# E - Activation energy in the Arrhenius Equation (J/mol)
# R - Universal Gas Constant = 8.31451 J/mol-K
EoverR = 8750
# Pre-exponential factor (1/sec)
k0 = 7.2e10
# U - Overall Heat Transfer Coefficient (W/m^2-K)
# A - Area - this value is specific for the U calculation (m^2)
UA = 5e4
# reaction rate
rA = k0*np.exp(-EoverR/T)*Ca
# Calculate concentration derivative
dCadt = q/V*(Caf - Ca) - rA
# Calculate temperature derivative
dTdt = q/V*(Tf - T) \
+ mdelH/(rho*Cp)*rA \
+ UA/V/rho/Cp*(Tc-T)
# Return xdot:
xdot = np.zeros(2)
xdot[0] = dCadt
xdot[1] = dTdt
return xdot
# Steady State Initial Conditions for the States
Ca_ss = 0.87725294608097
T_ss = 324.475443431599
x0 = np.empty(2)
x0[0] = Ca_ss
x0[1] = T_ss
# Steady State Initial Condition
u_ss = 300.0
# Feed Temperature (K)
Tf = 350
# Feed Concentration (mol/m^3)
Caf = 1
# Time Interval (min)
t = np.linspace(0,25,251)
# Store results for plotting
Ca = np.ones(len(t)) * Ca_ss
T = np.ones(len(t)) * T_ss
u = np.ones(len(t)) * u_ss
# Step cooling temperature to 295
u[10:100] = 303.0
u[100:190] = 297.0
u[190:] = 300.0
# Simulate CSTR
for i in range(len(t)-1):
ts = [t[i],t[i+1]]
y = odeint(cstr,x0,ts,args=(u[i+1],Tf,Caf))
Ca[i+1] = y[-1][0]
T[i+1] = y[-1][1]
x0[0] = Ca[i+1]
x0[1] = T[i+1]
# Construct results and save data file
# Column 1 = time
# Column 2 = cooling temperature
# Column 3 = reactor temperature
data = np.vstack((t,u,T)) # vertical stack
data = data.T # transpose data
np.savetxt('data_doublet.txt',data,delimiter=',',\
header='Time,Tc,T',comments='')
# Plot the results
plt.figure()
plt.subplot(3,1,1)
plt.plot(t,u,'b--',linewidth=3)
plt.ylabel('Cooling T (K)')
plt.legend(['Jacket Temperature'],loc='best')
plt.subplot(3,1,2)
plt.plot(t,Ca,'r-',linewidth=3)
plt.ylabel('Ca (mol/L)')
plt.legend(['Reactor Concentration'],loc='best')
plt.subplot(3,1,3)
plt.plot(t,T,'k.-',linewidth=3)
plt.ylabel('T (K)')
plt.xlabel('Time (min)')
plt.legend(['Reactor Temperature'],loc='best')
plt.show()
# + [markdown] id="oaQsjvhB1Mtm" colab_type="text"
# <p class='vspace'>Additional steps are preferred for systems that show a high degree a nonlinearity or when there is little additional expense to obtain the data. The following code generates data at multiple input levels and with varying different step time intervals. The cooling jacket temperature is not raised above 305 K to avoid reactor instability in open loop.
# </p>
# <div class='vspace'></div><div><img width='550px' src='http://apmonitor.com/do/uploads/Main/cstr_step_tests.png' alt='' title='' /></div>
# + id="SgCH6Rbj0WaQ" colab_type="code" cellView="form" colab={}
#@markdown Python Step Test Generator
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
# define CSTR model
def cstr(x,t,Tc):
Ca = x[0]
T = x[1]
Tf = 350
Caf = 1.0
q = 100
V = 100
rho = 1000
Cp = 0.239
mdelH = 5e4
EoverR = 8750
k0 = 7.2e10
UA = 5e4
rA = k0*np.exp(-EoverR/T)*Ca
dCadt = q/V*(Caf - Ca) - rA
dTdt = q/V*(Tf - T) \
+ mdelH/(rho*Cp)*rA \
+ UA/V/rho/Cp*(Tc-T)
xdot = np.zeros(2)
xdot[0] = dCadt
xdot[1] = dTdt
return xdot
# Steady State Initial Conditions for the States
Ca_ss = 0.87725294608097
T_ss = 324.475443431599
x0 = np.empty(2)
x0[0] = Ca_ss
x0[1] = T_ss
# Steady State Initial Condition
Tc_ss = 300.0
# Time Interval (min)
t = np.linspace(0,50,501)
# Store results for plotting
Ca = np.ones(len(t)) * Ca_ss
T = np.ones(len(t)) * T_ss
Tc = np.ones(len(t)) * Tc_ss
# Step cooling temperature
Tc[10:100] = 303.0
Tc[100:200] = 297.0
Tc[200:300] = 300.0
Tc[300:350] = 290.0
Tc[350:400] = 302.0
Tc[400:450] = 302.0
Tc[450:] = 299.0
# Simulate CSTR
for i in range(len(t)-1):
ts = [t[i],t[i+1]]
y = odeint(cstr,x0,ts,args=(Tc[i+1],))
Ca[i+1] = y[-1][0]
T[i+1] = y[-1][1]
x0[0] = Ca[i+1]
x0[1] = T[i+1]
# Construct results and save data file
# Column 1 = time
# Column 2 = cooling temperature
# Column 3 = reactor temperature
data = np.vstack((t,Tc,T)) # vertical stack
data = data.T # transpose data
np.savetxt('cstr_step_tests.txt',data,delimiter=',',\
header='Time,Tc,T',comments='')
# Plot the results
plt.figure()
plt.subplot(3,1,1)
plt.plot(t,Tc,'b--',linewidth=3)
plt.ylabel('Cooling T (K)')
plt.legend(['Jacket Temperature'],loc='best')
plt.subplot(3,1,2)
plt.plot(t,Ca,'r-',linewidth=3)
plt.ylabel('Ca (mol/L)')
plt.legend(['Reactor Concentration'],loc='best')
plt.subplot(3,1,3)
plt.plot(t,T,'k.-',linewidth=3)
plt.ylabel('T (K)')
plt.xlabel('Time (min)')
plt.legend(['Reactor Temperature'],loc='best')
plt.show()
# + [markdown] id="Hp_Wep1l1tD6" colab_type="text"
# ##Model Identification
#
# <p>There are many methods to develop a controller model. For a PID controller, an <a class='urllink' href='https://apmonitor.com/pdc/index.php/Main/FirstOrderOptimization' rel='nofollow'>FOPDT model</a> is one method to obtain <a class='urllink' href='https://apmonitor.com/pdc/index.php/Main/ProportionalIntegralDerivative' rel='nofollow'>IMC tuning parameters</a>. For linear MPC, there are many options to obtain a controller model through <a class='wikilink' href='http://apmonitor.com/do/index.php/Main/ModelIdentification'>identification methods</a>. For nonlinear MPC, the nonlinear simulator equations can be used to develop the controller. This section demonstrates how to obtain a linear model for the MPC application using the step test data generated in the prior section.
# </p>
# <div class='vspace'></div><div><img width='550px' src='http://apmonitor.com/do/uploads/Main/cstr_arx_fit.png' alt='' title='' /></div>
# + id="v28xWSe21jwZ" colab_type="code" cellView="form" colab={}
#@markdown Python ARX System ID
try:
from pip import main as pipmain
except:
from pip._internal import main as pipmain
pipmain(['install','gekko'])
from gekko import GEKKO
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# load data and parse into columns
url = 'http://apmonitor.com/do/uploads/Main/cstr_step_tests.txt'
data = pd.read_csv(url)
print(data.head())
# generate time-series model
t = data['Time']
u = data['Tc']
y = data['T']
m = GEKKO(remote=True) # remote=True for MacOS
# system identification
na = 2 # output coefficients
nb = 2 # input coefficients
yp,p,K = m.sysid(t,u,y,na,nb,shift='init',scale=True,objf=100,diaglevel=1)
# plot results of fitting
plt.figure()
plt.subplot(2,1,1)
plt.plot(t,u)
plt.legend([r'$T_c$'])
plt.ylabel('MV')
plt.subplot(2,1,2)
plt.plot(t,y)
plt.plot(t,yp)
plt.legend([r'$T_{meas}$',r'$T_{pred}$'])
plt.ylabel('CV')
plt.xlabel('Time')
plt.savefig('sysid.png')
# step test model
yc,uc = m.arx(p)
# rename MV and CV
Tc = uc[0]
T = yc[0]
# steady state initialization
m.options.IMODE = 1
Tc.value = 300
m.solve(disp=False)
# dynamic simulation (step test validation)
m.time = np.linspace(0,2,21)
m.options.IMODE = 4
Tc.value = np.ones(21)*300
Tc.value[5:] = 305
m.solve(disp=False)
plt.figure()
plt.subplot(2,1,1)
plt.title('Step Test')
plt.plot(m.time,Tc.value,'b-',label='Cooling Jacket')
plt.ylabel(r'$T_c (K)$')
plt.legend()
plt.subplot(2,1,2)
plt.plot(m.time,T.value,'r-',label='Reactor')
plt.ylabel('T (K)')
plt.xlabel('Time (min)')
plt.legend()
plt.show()
# + id="Eh4VBqRJ19_M" colab_type="code" cellView="form" outputId="6bcdc135-bb56-4d1c-d6f9-b71b46496546" colab={"base_uri": "https://localhost:8080/", "height": 336}
#@title ##Predictive Control
# %%html
<iframe width="560" height="315" src="https://www.youtube.com/embed/ZvvQ0_PdMPk" frameborder="0" allow="autoplay; encrypted-media" allowfullscreen></iframe>
# + id="0YQrmIh02OMW" colab_type="code" cellView="form" colab={}
#@markdown GEKKO Linear First-Order MPC Code
try:
from pip import main as pipmain
except:
from pip._internal import main as pipmain
pipmain(['install','gekko'])
from IPython.display import clear_output
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
from gekko import GEKKO
# Steady State Initial Condition
u_ss = 280.0
# Feed Temperature (K)
Tf = 350
# Feed Concentration (mol/m^3)
Caf = 1
# Steady State Initial Conditions for the States
Ca_ss = 1
T_ss = 304
x0 = np.empty(2)
x0[0] = Ca_ss
x0[1] = T_ss
#%% GEKKO linear MPC
m = GEKKO(remote=True)
m.time = [0,0.02,0.04,0.06,0.08,0.1,0.15,0.2,0.3,0.4,0.5]
# initial conditions
Tc0 = 280
T0 = 304
Ca0 = 1.0
tau = m.Const(value = 0.5)
Kp = m.Const(value = 1)
m.Tc = m.MV(value = Tc0,lb=250,ub=350)
m.T = m.CV(value = T_ss)
m.Equation(tau * m.T.dt() == -(m.T - T0) + Kp * (m.Tc - Tc0))
#MV tuning
m.Tc.STATUS = 1
m.Tc.FSTATUS = 0
m.Tc.DMAX = 100
m.Tc.DMAXHI = 5 # constrain movement up
m.Tc.DMAXLO = -100 # quick action down
#CV tuning
m.T.STATUS = 1
m.T.FSTATUS = 1
m.T.SP = 330
m.T.TR_INIT = 2
m.T.TAU = 1.0
m.options.CV_TYPE = 2
m.options.IMODE = 6
m.options.SOLVER = 3
#%% define CSTR model
def cstr(x,t,u,Tf,Caf):
# Inputs (3):
# Temperature of cooling jacket (K)
Tc = u
# Tf = Feed Temperature (K)
# Caf = Feed Concentration (mol/m^3)
# States (2):
# Concentration of A in CSTR (mol/m^3)
Ca = x[0]
# Temperature in CSTR (K)
T = x[1]
# Parameters:
# Volumetric Flowrate (m^3/sec)
q = 100
# Volume of CSTR (m^3)
V = 100
# Density of A-B Mixture (kg/m^3)
rho = 1000
# Heat capacity of A-B Mixture (J/kg-K)
Cp = 0.239
# Heat of reaction for A->B (J/mol)
mdelH = 5e4
# E - Activation energy in the Arrhenius Equation (J/mol)
# R - Universal Gas Constant = 8.31451 J/mol-K
EoverR = 8750
# Pre-exponential factor (1/sec)
k0 = 7.2e10
# U - Overall Heat Transfer Coefficient (W/m^2-K)
# A - Area - this value is specific for the U calculation (m^2)
UA = 5e4
# reaction rate
rA = k0*np.exp(-EoverR/T)*Ca
# Calculate concentration derivative
dCadt = q/V*(Caf - Ca) - rA
# Calculate temperature derivative
dTdt = q/V*(Tf - T) \
+ mdelH/(rho*Cp)*rA \
+ UA/V/rho/Cp*(Tc-T)
# Return xdot:
xdot = np.zeros(2)
xdot[0] = dCadt
xdot[1] = dTdt
return xdot
# Time Interval (min)
t = np.linspace(0,10,501)
# Store results for plotting
Ca = np.ones(len(t)) * Ca_ss
T = np.ones(len(t)) * T_ss
Tsp = np.ones(len(t)) * T_ss
u = np.ones(len(t)) * u_ss
# Set point steps
Tsp[0:100] = 330.0
Tsp[100:200] = 350.0
Tsp[200:300] = 370.0
Tsp[300:] = 390.0
# Create plot
plt.figure(figsize=(10,7))
plt.ion()
plt.show()
# Simulate CSTR
for i in range(len(t)-1):
# simulate one time period (0.05 sec each loop)
ts = [t[i],t[i+1]]
y = odeint(cstr,x0,ts,args=(u[i],Tf,Caf))
# retrieve measurements
Ca[i+1] = y[-1][0]
T[i+1] = y[-1][1]
# insert measurement
m.T.MEAS = T[i+1]
# update setpoint
m.T.SP = Tsp[i+1]
# solve MPC
m.solve(disp=True)
# change to a fixed starting point for trajectory
m.T.TR_INIT = 2
# retrieve new Tc value
u[i+1] = m.Tc.NEWVAL
# update initial conditions
x0[0] = Ca[i+1]
x0[1] = T[i+1]
#%% Plot the results
clear_output()
plt.clf()
plt.subplot(3,1,1)
plt.plot(t[0:i],u[0:i],'b--',linewidth=3)
plt.ylabel('Cooling T (K)')
plt.legend(['Jacket Temperature'],loc='best')
plt.subplot(3,1,2)
plt.plot(t[0:i],Ca[0:i],'r-',linewidth=3)
plt.ylabel('Ca (mol/L)')
plt.legend(['Reactor Concentration'],loc='best')
plt.subplot(3,1,3)
plt.plot(t[0:i],Tsp[0:i],'k-',linewidth=3,label=r'$T_{sp}$')
plt.plot(t[0:i],T[0:i],'b.-',linewidth=3,label=r'$T_{meas}$')
plt.ylabel('T (K)')
plt.xlabel('Time (min)')
plt.legend(['Reactor Temperature'],loc='best')
plt.draw()
plt.pause(0.01)
# + id="EeFminjP2i5x" colab_type="code" cellView="form" colab={}
#@markdown GEKKO ARX MPC Code
try:
from pip import main as pipmain
except:
from pip._internal import main as pipmain
pipmain(['install','gekko'])
from IPython.display import clear_output
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
import pandas as pd
from gekko import GEKKO
# load data and parse into columns
url = 'http://apmonitor.com/do/uploads/Main/cstr_step_tests.txt'
data = pd.read_csv(url)
print(data.head())
# generate time-series model
t = data['Time']
u = data['Tc']
y = data['T']
m = GEKKO(remote=True)
# system identification
na = 2 # output coefficients
nb = 2 # input coefficients
yp,p,K = m.sysid(t,u,y,na,nb,shift='init',scale=True,objf=100,diaglevel=1)
# plot results of fitting
plt.figure()
plt.subplot(2,1,1)
plt.plot(t,u)
plt.legend([r'$T_c$'])
plt.ylabel('MV')
plt.subplot(2,1,2)
plt.plot(t,y)
plt.plot(t,yp)
plt.legend([r'$T_{meas}$',r'$T_{pred}$'])
plt.ylabel('CV')
plt.xlabel('Time')
plt.savefig('sysid.png')
plt.show()
# step test model
yc,uc = m.arx(p)
# rename MV and CV
m.Tc = uc[0]
m.T = yc[0]
# steady state initialization
m.options.IMODE = 1
m.Tc.value = 280
m.solve(disp=True)
# GEKKO linear MPC
m.time = np.linspace(0,2,21)
# MV tuning
m.Tc.STATUS = 1
m.Tc.FSTATUS = 0
m.Tc.DMAX = 100
m.Tc.DCOST = 0.1
m.Tc.DMAXHI = 5 # constrain movement up
m.Tc.DMAXLO = -100 # quick action down
m.Tc.UPPER = 350
m.Tc.LOWER = 250
# CV tuning
m.T.STATUS = 1
m.T.FSTATUS = 1
m.T.SP = 330
m.T.TR_INIT = 1
m.T.TAU = 1.2
m.options.CV_TYPE = 2
m.options.IMODE = 6
m.options.SOLVER = 3
# define CSTR (plant)
def cstr(x,t,Tc):
Ca,T = x
Tf = 350; Caf = 1.0; q = 100; V = 100
rho = 1000; Cp = 0.239; mdelH = 5e4
EoverR = 8750; k0 = 7.2e10; UA = 5e4
rA = k0*np.exp(-EoverR/T)*Ca
dCadt = q/V*(Caf - Ca) - rA
dTdt = q/V*(Tf - T) + mdelH/(rho*Cp)*rA + UA/V/rho/Cp*(Tc-T)
return [dCadt,dTdt]
# Time Interval (min)
t = np.linspace(0,20,201)
# Store results for plotting
Ca_ss = 1; T_ss = 304; Tc_ss = 280
Ca = np.ones(len(t)) * Ca_ss
T = np.ones(len(t)) * T_ss
Tsp = np.ones(len(t)) * T_ss
Tc = np.ones(len(t)) * Tc_ss
# Set point steps
Tsp[0:40] = 330.0
Tsp[40:80] = 350.0
Tsp[80:120] = 370.0
Tsp[120:] = 390.0
# Create plot
plt.figure(figsize=(10,7))
plt.ion()
plt.show()
# Simulate CSTR
x0 = [Ca_ss,T_ss]
for i in range(len(t)-1):
y = odeint(cstr,x0,[0,0.05],args=(Tc[i],))
# retrieve measurements
Ca[i+1] = y[-1][0]
T[i+1] = y[-1][1]
# insert measurement
m.T.MEAS = T[i+1]
# update setpoint
m.T.SP = Tsp[i+1]
# solve MPC
m.solve(disp=True)
# retrieve new Tc value
Tc[i+1] = m.Tc.NEWVAL
# update initial conditions
x0[0] = Ca[i+1]
x0[1] = T[i+1]
#%% Plot the results
clear_output()
plt.clf()
plt.subplot(3,1,1)
plt.plot(t[0:i],Tc[0:i],'b--',linewidth=3)
plt.ylabel('Cooling T (K)')
plt.legend(['Jacket Temperature'],loc='best')
plt.subplot(3,1,2)
plt.plot(t[0:i],Ca[0:i],'r-',linewidth=3)
plt.ylabel('Ca (mol/L)')
plt.legend(['Reactor Concentration'],loc='best')
plt.subplot(3,1,3)
plt.plot(t[0:i],Tsp[0:i],'k-',linewidth=3,label=r'$T_{sp}$')
plt.plot(t[0:i],T[0:i],'b.-',linewidth=3,label=r'$T_{meas}$')
plt.ylabel('T (K)')
plt.xlabel('Time (min)')
plt.legend(['Temperature SP','Reactor Temperature'],loc='best')
plt.draw()
plt.pause(0.01)
# + id="ceO8_L--2zOr" colab_type="code" cellView="form" colab={}
#@markdown GEKKO Non-Linear MPC Code
try:
from pip import main as pipmain
except:
from pip._internal import main as pipmain
pipmain(['install','gekko'])
from IPython.display import clear_output
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import odeint
from gekko import GEKKO
# Steady State Initial Condition
u_ss = 280.0
# Feed Temperature (K)
Tf = 350
# Feed Concentration (mol/m^3)
Caf = 1
# Steady State Initial Conditions for the States
Ca_ss = 1
T_ss = 304
x0 = np.empty(2)
x0[0] = Ca_ss
x0[1] = T_ss
#%% GEKKO nonlinear MPC
m = GEKKO(remote=True)
m.time = [0,0.02,0.04,0.06,0.08,0.1,0.12,0.15,0.2]
# Volumetric Flowrate (m^3/sec)
q = 100
# Volume of CSTR (m^3)
V = 100
# Density of A-B Mixture (kg/m^3)
rho = 1000
# Heat capacity of A-B Mixture (J/kg-K)
Cp = 0.239
# Heat of reaction for A->B (J/mol)
mdelH = 5e4
# E - Activation energy in the Arrhenius Equation (J/mol)
# R - Universal Gas Constant = 8.31451 J/mol-K
EoverR = 8750
# Pre-exponential factor (1/sec)
k0 = 7.2e10
# U - Overall Heat Transfer Coefficient (W/m^2-K)
# A - Area - this value is specific for the U calculation (m^2)
UA = 5e4
# initial conditions
Tc0 = 280
T0 = 304
Ca0 = 1.0
tau = m.Const(value=0.5)
Kp = m.Const(value=1)
m.Tc = m.MV(value=Tc0,lb=250,ub=350)
m.T = m.CV(value=T_ss)
m.rA = m.Var(value=0)
m.Ca = m.CV(value=Ca_ss)
m.Equation(m.rA == k0*m.exp(-EoverR/m.T)*m.Ca)
m.Equation(m.T.dt() == q/V*(Tf - m.T) \
+ mdelH/(rho*Cp)*m.rA \
+ UA/V/rho/Cp*(m.Tc-m.T))
m.Equation(m.Ca.dt() == q/V*(Caf - m.Ca) - m.rA)
#MV tuning
m.Tc.STATUS = 1
m.Tc.FSTATUS = 0
m.Tc.DMAX = 100
m.Tc.DMAXHI = 20 # constrain movement up
m.Tc.DMAXLO = -100 # quick action down
#CV tuning
m.T.STATUS = 1
m.T.FSTATUS = 1
m.T.TR_INIT = 1
m.T.TAU = 1.0
DT = 0.5 # deadband
m.Ca.STATUS = 0
m.Ca.FSTATUS = 0 # no measurement
m.Ca.TR_INIT = 0
m.options.CV_TYPE = 1
m.options.IMODE = 6
m.options.SOLVER = 3
#%% define CSTR model
def cstr(x,t,u,Tf,Caf):
# Inputs (3):
# Temperature of cooling jacket (K)
Tc = u
# Tf = Feed Temperature (K)
# Caf = Feed Concentration (mol/m^3)
# States (2):
# Concentration of A in CSTR (mol/m^3)
Ca = x[0]
# Temperature in CSTR (K)
T = x[1]
# Parameters:
# Volumetric Flowrate (m^3/sec)
q = 100
# Volume of CSTR (m^3)
V = 100
# Density of A-B Mixture (kg/m^3)
rho = 1000
# Heat capacity of A-B Mixture (J/kg-K)
Cp = 0.239
# Heat of reaction for A->B (J/mol)
mdelH = 5e4
# E - Activation energy in the Arrhenius Equation (J/mol)
# R - Universal Gas Constant = 8.31451 J/mol-K
EoverR = 8750
# Pre-exponential factor (1/sec)
k0 = 7.2e10
# U - Overall Heat Transfer Coefficient (W/m^2-K)
# A - Area - this value is specific for the U calculation (m^2)
UA = 5e4
# reaction rate
rA = k0*np.exp(-EoverR/T)*Ca
# Calculate concentration derivative
dCadt = q/V*(Caf - Ca) - rA
# Calculate temperature derivative
dTdt = q/V*(Tf - T) \
+ mdelH/(rho*Cp)*rA \
+ UA/V/rho/Cp*(Tc-T)
# Return xdot:
xdot = np.zeros(2)
xdot[0] = dCadt
xdot[1] = dTdt
return xdot
# Time Interval (min)
t = np.linspace(0,8,401)
# Store results for plotting
Ca = np.ones(len(t)) * Ca_ss
T = np.ones(len(t)) * T_ss
Tsp = np.ones(len(t)) * T_ss
u = np.ones(len(t)) * u_ss
# Set point steps
Tsp[0:100] = 330.0
Tsp[100:200] = 350.0
Tsp[200:300] = 370.0
Tsp[300:] = 390.0
# Create plot
plt.figure(figsize=(10,7))
plt.ion()
plt.show()
# Simulate CSTR
for i in range(len(t)-1):
# simulate one time period (0.05 sec each loop)
ts = [t[i],t[i+1]]
y = odeint(cstr,x0,ts,args=(u[i],Tf,Caf))
# retrieve measurements
Ca[i+1] = y[-1][0]
T[i+1] = y[-1][1]
# insert measurement
m.T.MEAS = T[i+1]
# solve MPC
m.solve(disp=True)
m.T.SPHI = Tsp[i+1] + DT
m.T.SPLO = Tsp[i+1] - DT
# retrieve new Tc value
u[i+1] = m.Tc.NEWVAL
# update initial conditions
x0[0] = Ca[i+1]
x0[1] = T[i+1]
#%% Plot the results
clear_output()
plt.clf()
plt.subplot(3,1,1)
plt.plot(t[0:i],u[0:i],'b--',linewidth=3)
plt.ylabel('Cooling T (K)')
plt.legend(['Jacket Temperature'],loc='best')
plt.subplot(3,1,2)
plt.plot(t[0:i],Ca[0:i],'b.-',linewidth=3,label=r'$C_A$')
plt.plot([0,t[i-1]],[0.2,0.2],'r--',linewidth=2,label='limit')
plt.ylabel(r'$C_A$ (mol/L)')
plt.legend(loc='best')
plt.subplot(3,1,3)
plt.plot(t[0:i],Tsp[0:i],'k-',linewidth=3,label=r'$T_{sp}$')
plt.plot(t[0:i],T[0:i],'b.-',linewidth=3,label=r'$T_{meas}$')
plt.plot([0,t[i-1]],[400,400],'r--',linewidth=2,label='limit')
plt.ylabel('T (K)')
plt.xlabel('Time (min)')
plt.legend(loc='best')
plt.draw()
plt.pause(0.01)
# + [markdown] id="mpKwN6s53e74" colab_type="text"
# ##Solution in Python
#
# <div><img src='http://apmonitor.com/do/uploads/Main/download.png' alt='' title='' /> <a class='urllink' href='http://apmonitor.com/do/uploads/Main/cstr_pid_solution_Python.zip' rel='nofollow'>PID for CSTR Control (Python)</a> - <a class='urllink' href='https://youtu.be/tSOMSxGLzQo' rel='nofollow'>Solution Video</a></div>
# <div class='vspace'></div><div><img src='http://apmonitor.com/do/uploads/Main/download.png' alt='' title='' /> <a class='urllink' href='http://apmonitor.com/do/uploads/Main/cstr_mpc2_solution_Python.zip' rel='nofollow'>Linear MPC for CSTR Control (APM Python)</a> - <a class='urllink' href='https://youtu.be/nqv6jFeVUYA' rel='nofollow'>Solution Video</a></div>
# <div class='vspace'></div><div><img src='http://apmonitor.com/do/uploads/Main/download.png' alt='' title='' /> <a class='urllink' href='http://apmonitor.com/do/uploads/Main/cstr_nmpc_solution_Python.zip' rel='nofollow'>Nonlinear MPC for CSTR Control (APM Python)</a> - <a class='urllink' href='https://youtu.be/Jxpk4-daDLI' rel='nofollow'>Solution Video</a></div>
# <div class='vspace'></div><h4>Solution in Simulink</h4>
# <div><img src='http://apmonitor.com/do/uploads/Main/download.png' alt='' title='' /> <a class='urllink' href='http://apmonitor.com/do/uploads/Main/cstr_control_solution_PID.zip' rel='nofollow'>PID for CSTR Control (Simulink)</a> - <a class='urllink' href='https://youtu.be/sfhHcSF2i90' rel='nofollow'>Solution Video</a></div>
# <div class='vspace'></div><div><img src='http://apmonitor.com/do/uploads/Main/download.png' alt='' title='' /> <a class='urllink' href='http://apmonitor.com/do/uploads/Main/cstr_control_solution_Linear_MPC.zip' rel='nofollow'>Linear MPC for CSTR Control (Simulink)</a> - <a class='urllink' href='https://youtu.be/lBx10LvT8uA' rel='nofollow'>Solution Video</a></div>
# <div class='vspace'></div><div><img src='http://apmonitor.com/do/uploads/Main/download.png' alt='' title='' /> <a class='urllink' href='http://apmonitor.com/do/uploads/Main/cstr_control_solution_Nonlinear_MPC.zip' rel='nofollow'>Nonlinear MPC for CSTR Control (Simulink)</a> - <a class='urllink' href='https://youtu.be/PyrLMlht-PU' rel='nofollow'>Solution Video</a></div>
| notebooks/NonlinearControl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import statsmodels.formula.api as smf
import numpy as np
import matplotlib.pyplot as plt
# # <font face="gotham" color="purple"> What Is Panel Data? </font>
# **Panel data** is a hybrid data type that has feature of both _cross section_ and _time series_. Actually panel data are the most common data type in industry, for instance a car manufacturer has record of its suppliers' price level over time, a bank has full history of its clients' monthly balance for many years. Needless to say, to carry out serious researches, you must use panel data.
#
#
# Here we will use the data from "Why has Productivity Declined? Productivity and Public Investment" written by <NAME>.
#
# Variable names defined as below:
# ```
# STATE = state name
# ST_ABB = state abbreviation
# YR = 1970,...,1986
# P_CAP = public capital
# HWY = highway capital
# WATER = water utility capital
# UTIL = utility capital
# PC = private capital
# GSP = gross state product
# EMP = employment
# UNEMP = unemployment rate
# ```
df = pd.read_excel('Basic_Econometrics_practice_data.xlsx', sheet_name = 'Prod_PubInvestment')
df.head(5)
df.tail(5)
# Each state is recorded over time in several aspects, such as public capitals, highway capital, water facility capital and etc. If each state is recorded in equal length of time period, we call it **balanced panel**, otherwise **unbalanced panel**.
# Estimation methods includes four approaches
# 1. Pooled OLS model
# 2. Fixed effects least square dummy variable (LSDV) model
# 3. Fixed effects within-in group model
# 4. Random effects model
# # <font face="gotham" color="purple">Pooled OLS Regression </font>
# \begin{aligned}
# ln{GSP}_{i t} &=\beta_{1}+\beta_{2} \ln{PCAP}_{i t}+\beta_{3} \ln{HWY}_{i t}+\beta_{4} \ln{WATER}_{i t}+\beta_{5} \ln{UTIL}_{i t}+\beta_{6} \ln{EMP}_{i t}+u_{i t}
# \end{aligned}
# where $i$ means the $i$the state, $t$ means time period.
model = smf.ols(formula='np.log(GSP) ~ np.log(P_CAP) + np.log(PC) + np.log(HWY) + np.log(WATER) + np.log(UTIL) + np.log(EMP)', data=df)
results = model.fit()
print(results.summary())
# The common symptoms of pooled regression on panel data is that all most of coefficients will be highly significant and also $R^2$ is exceedingly high. However, we can still spot some problems, the conditional number is high, meaning multicollinearity and Durbin-Watson test is close to $0$ meaning autocorrelation or specification error.
# But the most prominent issue of this model is that it camouflages the heterogeneity that may exist among states. The heterogeneity of each state is subsumed by the disturbance term, which causes correlation between independent variables and disturbance terms, therefore OLS estimates are bound to be biased and inconsistent.
# # <font face="gotham" color="purple">The Fixed Effect LSDV Model</font>
# LSDV model allows heterogeneity to take part in by adding different intercept value
# \begin{aligned}
# ln{GSP}_{i t} &=\beta_{1i}+\beta_{2} \ln{PCAP}_{i t}+\beta_{3} \ln{HWY}_{i t}+\beta_{4} \ln{WATER}_{i t}+\beta_{5} \ln{UTIL}_{i t}+\beta_{6} \ln{EMP}_{i t}+u_{i t}
# \end{aligned}
# $\beta_{1i}$ represents the intercept for each state $i$. There are various possible reasons contributing to heterogeneity among states, such as population, average education level and urbanization rate, etc.
#
# _Fixed effect_ means that though each state has its own intercept, but it is **time-invariant**, i.e. constant over the time. If we assume **time-variant** intercept, the notation would be $\beta_{1it}$
df
# +
fig, ax = plt.subplots(nrows = 2, ncols = 3, figsize = (18, 12))
ax[0, 0].scatter(df['GSP'], df['P_CAP'], c ='r', s = 5)
ax[0, 0].grid()
ax[0, 0].set_xlabel('Public Capital')
ax[0, 0].set_ylabel('Gross Regional Produce')
ax[0, 1].scatter(df['GSP'], df['HWY'], c ='r', s = 5)
ax[0, 1].grid()
ax[0, 1].set_xlabel('High Way Capital')
ax[0, 1].set_ylabel('Gross Regional Produce')
ax[0, 2].scatter(df['GSP'], df['WATER'], c ='r', s = 5)
ax[0, 2].grid()
ax[0, 2].set_xlabel('Water Facility')
ax[0, 2].set_ylabel('Gross Regional Produce')
ax[1, 0].scatter(df['GSP'], df['UTIL'], c ='r', s = 5)
ax[1, 0].grid()
ax[1, 0].set_xlabel('Utiltiy Capital')
ax[1, 0].set_ylabel('Gross Regional Produce')
ax[1, 1].scatter(df['GSP'], df['PC'], c ='r', s = 5)
ax[1, 1].grid()
ax[1, 1].set_xlabel('Private Capital')
ax[1, 1].set_ylabel('Gross Regional Produce')
ax[1, 2].scatter(df['GSP'], df['EMP'], c ='r', s = 5)
ax[1, 2].grid()
ax[1, 2].set_xlabel('Employement')
ax[1, 2].set_ylabel('Gross Regional Produce')
plt.show()
# -
# Check how many states are there in the panel data
print(df['STATE'].unique())
print(len(df['STATE'].unique()))
# To avoid dummy variable trap, we can define $47$ dummy intercepts.
# Add dummies onto the intercept
#
# \begin{aligned}
# ln{GSP}_{i t} &=\alpha_{1}+ \sum_{j=2}^{48}\alpha_{j} D_{j i}+\beta_{2} \ln{PCAP}_{i t}+\beta_{3} \ln{HWY}_{i t}+\beta_{4} \ln{WATER}_{i t}+\beta_{5} \ln{UTIL}_{i t}+\beta_{6} \ln{EMP}_{i t}+u_{i t}
# \end{aligned}
#
# Use ```STATE``` as the dummy column and add ```drop_fist``` to avoid dummy trap.
# + tags=[]
df_dum = pd.get_dummies(data = df, columns = ['STATE'], drop_first=True)
# -
df_dum
| 9. Panel Data Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# ## Introduction
#
# ### Data
#
# This project aims to generate a model to predict the presence of a heart disease. [The UCI heart disease database](https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/) contains 76 attributes, but all published experiments refer to using a subset of 14. The target attribute is an integer valued from 0 (no presence) to 4. However, for sake of simplicity it will be reduced to binary classification, i.e, `0` vs `0 <`.
#
# *The authors of the databases: Hungarian Institute of Cardiology. Budapest: <NAME>, M.D. University Hospital, Zurich, Switzerland: <NAME>, M.D. University Hospital, Basel, Switzerland: <NAME>, M.D. V.A. Medical Center, Long Beach and Cleveland Clinic Foundation: <NAME>, M.D., Ph.D.*
#
# ### Attributes
#
# | | Description | Variable | Type |
# |:---------|:---------------------------------------------------------------------------------------------|:-----------|:--------|
# | age | age in years | continuous | `int` |
# | sex | 1 = male, 0 = female | categorial | `int` |
# | cp | chest pain type: 1: typical angina, 2: atypical angina, 3: non-anginal pain, 4: asymptomatic | categorial | `int` |
# | trestbps | resting blood pressure in mm Hg | continuous | `float` |
# | chol | serum cholestoral in mg/dl | continuous | `float` |
# | fbs | fasting blood sugar > 120 mg/dl: 1 = true, 0 = false | categorial | `int` |
# | restecg | 0: normal, 1: having ST-T wave abnormality, 2: left ventricular hypertrophy | categorial | `int` |
# | thalach | maximum heart rate achieved | continuous | `float` |
# | exang | exercise induced angina (1 = yes; 0 = no) | categorial | `int` |
# | oldpeak | ST depression induced by exercise relative to rest | continuous | `float` |
# | slope | the slope of the peak exercise ST segment: 1: upsloping, 2: flat, 3: downsloping | categorial | `int` |
# | ca | number of major vessels: (0-3) colored by flourosopy | continuous | `int` |
# | thal | 3: normal, 6: fixed defect, 7: reversable defect | categorial | `int` |
# | target | diagnosis of heart disease: (0 = false, 1 = true | categorial | `int` |
#
#
# ### Flow
#
# [Data fetching](#data-fetching) --> [Wrangling](#wrangling) --> [Data analysis]() --> [Modeling]() --> [evaluation]()
# ## Imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.api as sm
from sklearn.feature_selection import RFE
from sklearn import model_selection
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split as split
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn import svm
# ## Configuration
# %matplotlib inline
sns.set(style="white")
sns.set(style="whitegrid", color_codes=True)
# +
### Data fetching
# +
path_cleveland = "https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.cleveland.data"
path_hungary = "https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.hungarian.data"
path_swiss = "https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.switzerland.data"
path_veniceb = "https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/processed.va.data"
paths = [path_cleveland, path_hungary, path_swiss, path_veniceb]
names = ["age", "sex", "cp", "trestbps", "chol", "fbs", "restecg", "thalach", "exang",
"oldpeak", "slope", "ca", "thal", "target"]
df = pd.concat(pd.read_csv(path, names=names) for path in paths)
df.head()
# -
# ## Wrangling
# ### Handling missing values
df.replace("?", np.nan, inplace=True)
df.isnull().sum()
df.dropna(axis=0, inplace=True)
df.reset_index(drop = True, inplace = True)
# ### Correcting data types
df['ca'] = pd.to_numeric(df['ca'], errors='coerce')
df['thal'] = pd.to_numeric(df['thal'], errors='coerce')
df[['age', 'sex', 'cp', 'fbs', 'restecg', 'exang', 'ca', 'slope', 'thal']] = df[['age', 'sex', 'cp', 'fbs', 'restecg', 'exang', 'ca', 'slope', 'thal']].astype(int)
df[['trestbps', 'chol', 'thalach', 'oldpeak']] = df[['trestbps', 'chol', 'thalach', 'oldpeak']].astype(float)
df['target'].replace(to_replace=[1, 2, 3, 4], value=1, inplace=True)
# ## Exploritory data analysis
# ### Target
# +
fig_target, ax = plt.subplots(nrows=1, ncols=1, figsize=(4, 4))
sns.countplot(x='target', data=df, ax=ax)
for i, p in enumerate(ax.patches):
count = df['target'].value_counts().values[i]
ax.text(p.get_x() + p.get_width() / 2.,
p.get_height() + 3,
)
# -
| heart_disease_prediction_python3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # How to define a machine
#
# This tutorial shows the different ways to define electrical machine. To do so, it presents the definition of the **Toyota Prius 2004** interior magnet with distributed winding \[1\].
#
# The notebook related to this tutorial is available on [GitHub](https://github.com/Eomys/pyleecan/tree/master/Tutorials/tuto_Machine.ipynb).
#
# ## Type of machines Pyleecan can model
# Pyleecan handles the geometrical modelling of main 2D radial flux machines such as:
# - surface or interior permanent magnet machines (SPMSM, IPMSM)
# - synchronous reluctance machines (SynRM)
# - squirrel-cage induction machines and doubly-fed induction machines (SCIM, DFIM)
# - would rotor synchronous machines and salient pole synchronous machines (WSRM)
# - switched reluctance machines (SRM)
#
# The architecture of Pyleecan also enables to define other kinds of machines (with more than two laminations for instance). More information in our ICEM 2020 pyblication \[2\]
#
# Every machine can be defined by using the **Graphical User Interface** or directly in **Python script**.
#
# ## Defining machine with Pyleecan GUI
# The GUI is the easiest way to define machine in Pyleecan. Its purpose is to create or load a machine and save it in JSON format to be loaded in a python script. The interface enables to define step by step in a user-friendly way every characteristics of the machine such as:
# - topology
# - dimensions
# - materials
# - winding
#
# Each parameter is explained by a tooltip and the machine can be previewed at each stage of the design.
#
# ## Start the GUI
# The GUI can be started by running the following command in the notebook:
#
# To use it on Anaconda you may need to create the system variable:
#
# QT_QPA_PLATFORM_PLUGIN_PATH : path\to\anaconda3\Lib\site-packages\PySide2\plugins\platforms
#
# ```python
# # Start Pyleecan GUI from the Jupyter Notebook
# # %run -m pyleecan
# ```
#
# The GUI can also be launched in a terminal by calling the following command in a terminal:
# ```
# Path/to/python.exe -m pyleecan
# ```
# ## load a machine
# Once the machine defined in the GUI it can be loaded with the following commands:
# +
# %matplotlib notebook
# Load the machine
from os.path import join
from pyleecan.Functions.load import load
from pyleecan.definitions import DATA_DIR
IPMSM_A = load(join(DATA_DIR, "Machine", "IPMSM_A.json"))
IPMSM_A.plot()
# -
# ## Defining Machine in scripting mode
# Pyleecan also enables to define the machine in scripting mode, using different classes. Each class is defined from a csv file in the folder _pyleecan/Generator/ClasseRef_ and the documentation of every class is available on the dedicated [webpage](https://www.pyleecan.org/pyleecan.Classes.html).
# The following image shows the machine classes organization :
#
# 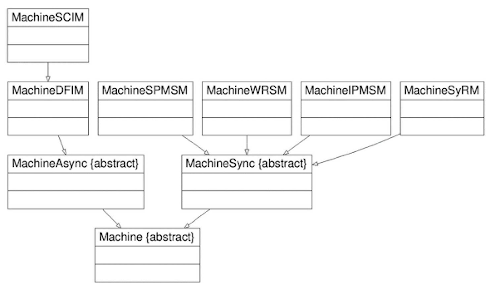
#
# Every rotor and stator can be created with the **Lamination** class or one of its daughters.
#
# 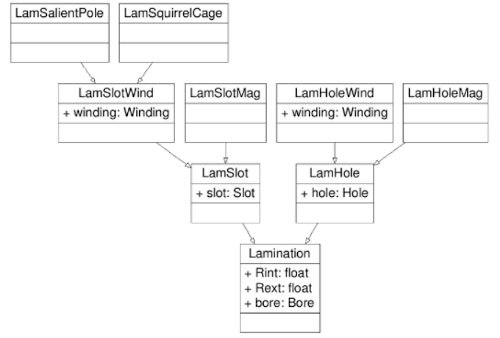
#
# The scripting enables to define some complex and exotic machine that can't be defined in the GUI such as this one:
# +
from pyleecan.Classes.MachineUD import MachineUD
from pyleecan.Classes.LamSlotWind import LamSlotWind
from pyleecan.Classes.LamSlot import LamSlot
from pyleecan.Classes.WindingCW2LT import WindingCW2LT
from pyleecan.Classes.SlotW10 import SlotW10
from pyleecan.Classes.SlotW22 import SlotW22
from numpy import pi
machine = MachineUD()
# Main geometry parameter
Rext = 170e-3 # Exterior radius of outter lamination
W1 = 30e-3 # Width of first lamination
A1 = 2.5e-3 # Width of the first airgap
W2 = 20e-3
A2 = 10e-3
W3 = 20e-3
A3 = 2.5e-3
W4 = 60e-3
# Outer stator
lam1 = LamSlotWind(Rext=Rext, Rint=Rext - W1, is_internal=False, is_stator=True)
lam1.slot = SlotW22(
Zs=12, W0=2 * pi / 12 * 0.75, W2=2 * pi / 12 * 0.75, H0=0, H2=W1 * 0.65
)
lam1.winding = WindingCW2LT(qs=3, p=3)
# Outer rotor
lam2 = LamSlot(
Rext=lam1.Rint - A1, Rint=lam1.Rint - A1 - W2, is_internal=True, is_stator=False
)
lam2.slot = SlotW10(Zs=22, W0=25e-3, W1=25e-3, W2=15e-3, H0=0, H1=0, H2=W2 * 0.75)
# Inner rotor
lam3 = LamSlot(
Rext=lam2.Rint - A2,
Rint=lam2.Rint - A2 - W3,
is_internal=False,
is_stator=False,
)
lam3.slot = SlotW10(
Zs=22, W0=17.5e-3, W1=17.5e-3, W2=12.5e-3, H0=0, H1=0, H2=W3 * 0.75
)
# Inner stator
lam4 = LamSlotWind(
Rext=lam3.Rint - A3, Rint=lam3.Rint - A3 - W4, is_internal=True, is_stator=True
)
lam4.slot = SlotW10(Zs=12, W0=25e-3, W1=25e-3, W2=1e-3, H0=0, H1=0, H2=W4 * 0.75)
lam4.winding = WindingCW2LT(qs=3, p=3)
# Machine definition
machine.lam_list = [lam1, lam2, lam3, lam4]
# Plot, check and save
machine.plot()
# -
# ## Stator definition
# To define the stator, we initialize a [**LamSlotWind**](http://pyleecan.org/pyleecan.Classes.LamSlotWind.html) object with the different parameters. In pyleecan, all the parameters must be set in SI units.
# +
from pyleecan.Classes.LamSlotWind import LamSlotWind
mm = 1e-3 # Millimeter
# Lamination setup
stator = LamSlotWind(
Rint=80.95 * mm, # internal radius [m]
Rext=134.62 * mm, # external radius [m]
L1=83.82 * mm, # Lamination stack active length [m] without radial ventilation airducts
# but including insulation layers between lamination sheets
Nrvd=0, # Number of radial air ventilation duct
Kf1=0.95, # Lamination stacking / packing factor
is_internal=False,
is_stator=True,
)
# -
# Then we add 48 slots using [**SlotW11**](http://pyleecan.org/pyleecan.Classes.SlotW11.html) which is one of the 25 Slot classes:
# +
from pyleecan.Classes.SlotW11 import SlotW11
# Slot setup
stator.slot = SlotW11(
Zs=48, # Slot number
H0=1.0 * mm, # Slot isthmus height
H1=0, # Height
H2=33.3 * mm, # Slot height below wedge
W0=1.93 * mm, # Slot isthmus width
W1=5 * mm, # Slot top width
W2=8 * mm, # Slot bottom width
R1=4 * mm # Slot bottom radius
)
# -
# As for the slot, we can define the winding and its conductor with [**WindingDW1L**](http://pyleecan.org/pyleecan.Classes.WindingDW1L.html) and [**CondType11**](http://pyleecan.org/pyleecan.Classes.CondType11.html). The conventions for winding are further explained on [pyleecan website](https://pyleecan.org/winding.convention.html)
# +
from pyleecan.Classes.WindingDW1L import WindingDW1L
from pyleecan.Classes.CondType11 import CondType11
# Winding setup
stator.winding = WindingDW1L(
qs=3, # number of phases
Lewout=0, # staight length of conductor outside lamination before EW-bend
p=4, # number of pole pairs
Ntcoil=9, # number of turns per coil
Npcpp=1, # number of parallel circuits per phase
Nslot_shift_wind=0, # 0 not to change the stator winding connection matrix built by pyleecan number
# of slots to shift the coils obtained with pyleecan winding algorithm
# (a, b, c becomes b, c, a with Nslot_shift_wind1=1)
is_reverse_wind=False # True to reverse the default winding algorithm along the airgap
# (c, b, a instead of a, b, c along the trigonometric direction)
)
# Conductor setup
stator.winding.conductor = CondType11(
Nwppc_tan=1, # stator winding number of preformed wires (strands)
# in parallel per coil along tangential (horizontal) direction
Nwppc_rad=1, # stator winding number of preformed wires (strands)
# in parallel per coil along radial (vertical) direction
Wwire=0.000912, # single wire width without insulation [m]
Hwire=2e-3, # single wire height without insulation [m]
Wins_wire=1e-6, # winding strand insulation thickness [m]
type_winding_shape=0, # type of winding shape for end winding length calculation
# 0 for hairpin windings
# 1 for normal windings
)
# -
# ## Rotor definition
#
# For this example, we use the [**LamHole**](http://www.pyleecan.org/pyleecan.Classes.LamHole.html) class to define the rotor as a lamination with holes to contain magnet.
#
# In the same way as for the stator, we start by defining the lamination:
# +
from pyleecan.Classes.LamHole import LamHole
# Rotor setup
rotor = LamHole(
Rint=55.32 * mm, # Internal radius
Rext=80.2 * mm, # external radius
is_internal=True,
is_stator=False,
L1=stator.L1 # Lamination stack active length [m]
# without radial ventilation airducts but including insulation layers between lamination sheets
)
# -
# After that, we can add holes with magnets to the rotor using the class [**HoleM50**](http://www.pyleecan.org/pyleecan.Classes.HoleM50.html):
from pyleecan.Classes.HoleM50 import HoleM50
rotor.hole = list()
rotor.hole.append(
HoleM50(
Zh=8, # Number of Hole around the circumference
W0=42.0 * mm, # Slot opening
W1=0, # Tooth width (at V bottom)
W2=0, # Distance Magnet to bottom of the V
W3=14.0 * mm, # Tooth width (at V top)
W4=18.9 * mm, # Magnet Width
H0=10.96 * mm, # Slot Depth
H1=1.5 * mm, # Distance from the lamination Bore
H2=1 * mm, # Additional depth for the magnet
H3=6.5 * mm, # Magnet Height
H4=0, # Slot top height
)
)
# The holes are defined as a list to enable to create several layers of holes and/or to combine different kinds of holes
#
# ## Create a shaft and a frame
#
# The classes [**Shaft**](http://www.pyleecan.org/pyleecan.Classes.Shaft.html) and [**Frame**](http://www.pyleecan.org/pyleecan.Classes.Frame.html) enable to add a shaft and a frame to the machine. For this example there is no shaft:
# +
from pyleecan.Classes.Shaft import Shaft
from pyleecan.Classes.Frame import Frame
# Set shaft
shaft = Shaft(Drsh=rotor.Rint * 2, # Diamater of the rotor shaft [m]
# used to estimate bearing diameter for friction losses
Lshaft=1.2 # length of the rotor shaft [m]
)
# -
# ## Set materials and magnets
#
# Every Pyleecan object can be saved in JSON using the method `save` and can be loaded with the `load` function.
# In this example, the materials *M400_50A* and *Copper1* are loaded while the material *Magnet_prius* is created with the classes [**Material**](http://www.pyleecan.org/pyleecan.Classes.Material.html) and [**MatMagnetics**](http://www.pyleecan.org/pyleecan.Classes.MatMagnetics.html).
# +
from pyleecan.Classes.Material import Material
from pyleecan.Classes.MatMagnetics import MatMagnetics
# Loading Materials
M400_50A = load(join(DATA_DIR, "Material", "M400-50A.json"))
Copper1 = load(join(DATA_DIR, "Material", "Copper1.json"))
# Defining magnets
Magnet_prius = Material(name="Magnet_prius")
# Definition of the magnetic properties of the material
Magnet_prius.mag = MatMagnetics(
mur_lin = 1.05, # Relative magnetic permeability
Hc = 902181.163126629, # Coercitivity field [A/m]
alpha_Br = -0.001, # temperature coefficient for remanent flux density /°C compared to 20°C
Brm20 = 1.24, # magnet remanence induction at 20°C [T]
Wlam = 0, # lamination sheet width without insulation [m] (0 == not laminated)
)
# Definition of the electric properties of the material
Magnet_prius.elec.rho = 1.6e-06 # Resistivity at 20°C
# Definition of the structural properties of the material
Magnet_prius.struct.rho = 7500.0 # mass per unit volume [kg/m3]
# Set Materials
stator.mat_type = M400_50A
rotor.mat_type = M400_50A
stator.winding.conductor.cond_mat = Copper1
# Set magnets in the rotor hole
rotor.hole[0].magnet_0.mat_type = Magnet_prius
rotor.hole[0].magnet_1.mat_type = Magnet_prius
rotor.hole[0].magnet_0.type_magnetization = 1
rotor.hole[0].magnet_1.type_magnetization = 1
# -
# ## Create, save and plot the machine
# Finally, the Machine object can be created with [**MachineIPMSM**](http://www.pyleecan.org/pyleecan.Classes.MachineIPMSM.html) and saved using the `save` method.
# +
from pyleecan.Classes.MachineIPMSM import MachineIPMSM
# %matplotlib notebook
IPMSM_Prius_2004 = MachineIPMSM(
name="Toyota Prius 2004",
stator=stator,
rotor=rotor,
shaft=shaft,
frame=None
)
IPMSM_Prius_2004.save('IPMSM_Toyota_Prius_2004.json')
im=IPMSM_Prius_2004.plot()
# -
# Note that Pyleecan also handles ventilation duct thanks to the classes :
# - [**VentilationCirc**](http://www.pyleecan.org/pyleecan.Classes.VentilationCirc.html)
# - [**VentilationPolar**](http://www.pyleecan.org/pyleecan.Classes.VentilationPolar.html)
# - [**VentilationTrap**](http://www.pyleecan.org/pyleecan.Classes.VentilationTrap.html)
# [1] <NAME>, <NAME> and <NAME>, "Electromagnetic and vibrational characteristic of IPM over full torque-speed range", *2013 International Electric Machines & Drives Conference*, Chicago, IL, 2013, pp. 295-302.
#
# [2] <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>, “Design Optimization of Innovative Electrical Machines Topologies Based on Pyleecan Opensource Object-Oriented Software,” in 24th International Conference on Electrical Machines (ICEM), 2020.
| Tutorials/tuto_Machine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="CCQY7jpBfMur"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="z6X9omPnfO_h"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="F1xIRPtY0E1w"
# # Keras: A quick overview
# + [markdown] colab_type="text" id="VyOjQZHhZxaA"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/keras/overview"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/keras/overview.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="VUJTep_x5-R8"
# This guide gives you the basics to get started with Keras. It's a 10-minute read.
# + [markdown] colab_type="text" id="IsK5aF2xZ-40"
# ## Import tf.keras
#
# `tf.keras` is TensorFlow's implementation of the
# [Keras API specification](https://keras.io). This is a high-level
# API to build and train models that includes first-class support for
# TensorFlow-specific functionality, such as [eager execution](#eager_execution),
# `tf.data` pipelines, and [Estimators](./estimators.md).
# `tf.keras` makes TensorFlow easier to use without sacrificing flexibility and
# performance.
#
# To get started, import `tf.keras` as part of your TensorFlow program setup:
# + colab={} colab_type="code" id="TgPcBFru0E1z"
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# # %tensorflow_version only exists in Colab.
import tensorflow.compat.v2 as tf
except Exception:
pass
tf.enable_v2_behavior()
from tensorflow.compat.v2 import keras
# + [markdown] colab_type="text" id="lj03RamP0E13"
# `tf.keras` can run any Keras-compatible code, but keep in mind:
#
# * The `tf.keras` version in the latest TensorFlow release might not be the same
# as the latest `keras` version from PyPI. Check `tf.keras.__version__`.
# * When [saving a model's weights](#weights_only), `tf.keras` defaults to the
# [checkpoint format](./checkpoints.md). Pass `save_format='h5'` to
# use HDF5 (or pass a filename that ends in `.h5`).
# + [markdown] colab_type="text" id="7e1LPcXx0gR6"
# ## Build a simple model
#
# ### Sequential model
#
# In Keras, you assemble *layers* to build *models*. A model is (usually) a graph
# of layers. The most common type of model is a stack of layers: the
# `tf.keras.Sequential` model.
#
# To build a simple, fully-connected network (i.e. multi-layer perceptron):
# + colab={} colab_type="code" id="WM-DUVQB0E14"
from tensorflow.compat.v2.keras import layers
model = tf.keras.Sequential()
# Adds a densely-connected layer with 64 units to the model:
model.add(layers.Dense(64, activation='relu'))
# Add another:
model.add(layers.Dense(64, activation='relu'))
# Add a softmax layer with 10 output units:
model.add(layers.Dense(10, activation='softmax'))
# + [markdown] colab_type="text" id="I2oH0-cxH7YA"
# You can find a complete, short example of how to use Sequential models [here](https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/quickstart/beginner.ipynb).
#
# To learn about building more advanced models than Sequential models, see:
# - [Guide to the Keras Functional API](./functional.ipynb)
# - [Guide to writing layers and models from scratch with subclassing](./custom_layers_and_models.ipynb)
# + [markdown] colab_type="text" id="-ztyTipu0E18"
# ### Configure the layers
#
# There are many `tf.keras.layers` available. Most of them share some common constructor
# arguments:
#
# * `activation`: Set the activation function for the layer. This parameter is
# specified by the name of a built-in function or as a callable object. By
# default, no activation is applied.
# * `kernel_initializer` and `bias_initializer`: The initialization schemes
# that create the layer's weights (kernel and bias). This parameter is a name or
# a callable object. This defaults to the `"Glorot uniform"` initializer.
# * `kernel_regularizer` and `bias_regularizer`: The regularization schemes
# that apply the layer's weights (kernel and bias), such as L1 or L2
# regularization. By default, no regularization is applied.
#
# The following instantiates `tf.keras.layers.Dense` layers using constructor
# arguments:
# + colab={} colab_type="code" id="MlL7PBtp0E19"
# Create a sigmoid layer:
layers.Dense(64, activation='sigmoid')
# Or:
layers.Dense(64, activation=tf.keras.activations.sigmoid)
# A linear layer with L1 regularization of factor 0.01 applied to the kernel matrix:
layers.Dense(64, kernel_regularizer=tf.keras.regularizers.l1(0.01))
# A linear layer with L2 regularization of factor 0.01 applied to the bias vector:
layers.Dense(64, bias_regularizer=tf.keras.regularizers.l2(0.01))
# A linear layer with a kernel initialized to a random orthogonal matrix:
layers.Dense(64, kernel_initializer='orthogonal')
# A linear layer with a bias vector initialized to 2.0s:
layers.Dense(64, bias_initializer=tf.keras.initializers.Constant(2.0))
# + [markdown] colab_type="text" id="9NR6reyk0E2A"
# ## Train and evaluate
#
# ### Set up training
#
# After the model is constructed, configure its learning process by calling the
# `compile` method:
# + colab={} colab_type="code" id="sJ4AOn090E2A"
model = tf.keras.Sequential([
# Adds a densely-connected layer with 64 units to the model:
layers.Dense(64, activation='relu', input_shape=(32,)),
# Add another:
layers.Dense(64, activation='relu'),
# Add a softmax layer with 10 output units:
layers.Dense(10, activation='softmax')])
model.compile(optimizer=tf.keras.optimizers.Adam(0.01),
loss='categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] colab_type="text" id="HG-RAa9F0E2D"
# `tf.keras.Model.compile` takes three important arguments:
#
# * `optimizer`: This object specifies the training procedure. Pass it optimizer
# instances from the `tf.keras.optimizers` module, such as
# `tf.keras.optimizers.Adam` or
# `tf.keras.optimizers.SGD`. If you just want to use the default parameters, you can also specify optimizers via strings, such as `'adam'` or `'sgd'`.
# * `loss`: The function to minimize during optimization. Common choices include
# mean square error (`mse`), `categorical_crossentropy`, and
# `binary_crossentropy`. Loss functions are specified by name or by
# passing a callable object from the `tf.keras.losses` module.
# * `metrics`: Used to monitor training. These are string names or callables from
# the `tf.keras.metrics` module.
# * Additionally, to make sure the model trains and evaluates eagerly, you can make sure to pass `run_eagerly=True` as a parameter to compile.
#
#
# The following shows a few examples of configuring a model for training:
# + colab={} colab_type="code" id="St4Mgdar0E2E"
# Configure a model for mean-squared error regression.
model.compile(optimizer=tf.keras.optimizers.Adam(0.01),
loss='mse', # mean squared error
metrics=['mae']) # mean absolute error
# Configure a model for categorical classification.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.01),
loss=tf.keras.losses.CategoricalCrossentropy(),
metrics=[tf.keras.metrics.CategoricalAccuracy()])
# + [markdown] colab_type="text" id="yjI5rbi80E2G"
# ### Train from NumPy data
#
# For small datasets, use in-memory [NumPy](https://www.numpy.org/)
# arrays to train and evaluate a model. The model is "fit" to the training data
# using the `fit` method:
# + colab={} colab_type="code" id="3CvP6L-m0E2I"
import numpy as np
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
model.fit(data, labels, epochs=10, batch_size=32)
# + [markdown] colab_type="text" id="N-pnVaFe0E2N"
# `tf.keras.Model.fit` takes three important arguments:
#
# * `epochs`: Training is structured into *epochs*. An epoch is one iteration over
# the entire input data (this is done in smaller batches).
# * `batch_size`: When passed NumPy data, the model slices the data into smaller
# batches and iterates over these batches during training. This integer
# specifies the size of each batch. Be aware that the last batch may be smaller
# if the total number of samples is not divisible by the batch size.
# * `validation_data`: When prototyping a model, you want to easily monitor its
# performance on some validation data. Passing this argument—a tuple of inputs
# and labels—allows the model to display the loss and metrics in inference mode
# for the passed data, at the end of each epoch.
#
# Here's an example using `validation_data`:
# + colab={} colab_type="code" id="gFcXzVQa0E2N"
import numpy as np
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
val_data = np.random.random((100, 32))
val_labels = np.random.random((100, 10))
model.fit(data, labels, epochs=10, batch_size=32,
validation_data=(val_data, val_labels))
# + [markdown] colab_type="text" id="-6ImyXzz0E2Q"
# ### Train from tf.data datasets
#
# Use the [Datasets API](../data.md) to scale to large datasets
# or multi-device training. Pass a `tf.data.Dataset` instance to the `fit`
# method:
# + colab={} colab_type="code" id="OziqhpIj0E2R"
# Instantiates a toy dataset instance:
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
model.fit(dataset, epochs=10)
# + [markdown] colab_type="text" id="I7BcMHkB0E2U"
# Since the `Dataset` yields batches of data, this snippet does not require a `batch_size`.
#
# Datasets can also be used for validation:
# + colab={} colab_type="code" id="YPMb3A0N0E2V"
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
val_dataset = tf.data.Dataset.from_tensor_slices((val_data, val_labels))
val_dataset = val_dataset.batch(32)
model.fit(dataset, epochs=10,
validation_data=val_dataset)
# + [markdown] colab_type="text" id="IgGdlXso0E2X"
# ### Evaluate and predict
#
# The `tf.keras.Model.evaluate` and `tf.keras.Model.predict` methods can use NumPy
# data and a `tf.data.Dataset`.
#
# Here's how to *evaluate* the inference-mode loss and metrics for the data provided:
# + colab={} colab_type="code" id="mhDbOHEK0E2Y"
# With Numpy arrays
data = np.random.random((1000, 32))
labels = np.random.random((1000, 10))
model.evaluate(data, labels, batch_size=32)
# With a Dataset
dataset = tf.data.Dataset.from_tensor_slices((data, labels))
dataset = dataset.batch(32)
model.evaluate(dataset)
# + [markdown] colab_type="text" id="UXUTmDfb0E2b"
# And here's how to *predict* the output of the last layer in inference for the data provided,
# as a NumPy array:
# + colab={} colab_type="code" id="9e3JsSoQ0E2c"
result = model.predict(data, batch_size=32)
print(result.shape)
# + [markdown] colab_type="text" id="GuTb71gYILLG"
# For a complete guide on training and evaluation, including how to write custom training loops from scratch, see the [Guide to Training & Evaluation](./training_and_evaluation.ipynb).
# + [markdown] colab_type="text" id="fzEOW4Cn0E2h"
# ## Build complex models
#
# ### The Functional API
#
# The `tf.keras.Sequential` model is a simple stack of layers that cannot
# represent arbitrary models. Use the
# [Keras functional API](./functional.ipynb)
# to build complex model topologies such as:
#
# * Multi-input models,
# * Multi-output models,
# * Models with shared layers (the same layer called several times),
# * Models with non-sequential data flows (e.g. residual connections).
#
# Building a model with the functional API works like this:
#
# 1. A layer instance is callable and returns a tensor.
# 2. Input tensors and output tensors are used to define a `tf.keras.Model`
# instance.
# 3. This model is trained just like the `Sequential` model.
#
# The following example uses the functional API to build a simple, fully-connected
# network:
# + colab={} colab_type="code" id="mROj832r0E2i"
inputs = tf.keras.Input(shape=(32,)) # Returns an input placeholder
# A layer instance is callable on a tensor, and returns a tensor.
x = layers.Dense(64, activation='relu')(inputs)
x = layers.Dense(64, activation='relu')(x)
predictions = layers.Dense(10, activation='softmax')(x)
# + [markdown] colab_type="text" id="AFmspHeG1_W7"
# Instantiate the model given inputs and outputs.
# + colab={} colab_type="code" id="5k5uzlyu16HM"
model = tf.keras.Model(inputs=inputs, outputs=predictions)
# The compile step specifies the training configuration.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Trains for 5 epochs
model.fit(data, labels, batch_size=32, epochs=5)
# + [markdown] colab_type="text" id="EcKSLH3i0E2k"
# ### Model subclassing
#
# Build a fully-customizable model by subclassing `tf.keras.Model` and defining
# your own forward pass. Create layers in the `__init__` method and set them as
# attributes of the class instance. Define the forward pass in the `call` method.
#
# Model subclassing is particularly useful when
# [eager execution](./eager.md) is enabled, because it allows the forward pass
# to be written imperatively.
#
# Note: if you need your model to *always* run imperatively, you can set `dynamic=True` when calling the `super` constructor.
#
# > Key Point: Use the right API for the job. While model subclassing offers
# flexibility, it comes at a cost of greater complexity and more opportunities for
# user errors. If possible, prefer the functional API.
#
# The following example shows a subclassed `tf.keras.Model` using a custom forward
# pass that does not have to be run imperatively:
# + colab={} colab_type="code" id="KLiHWzcn2Fzk"
class MyModel(tf.keras.Model):
def __init__(self, num_classes=10):
super(MyModel, self).__init__(name='my_model')
self.num_classes = num_classes
# Define your layers here.
self.dense_1 = layers.Dense(32, activation='relu')
self.dense_2 = layers.Dense(num_classes, activation='sigmoid')
def call(self, inputs):
# Define your forward pass here,
# using layers you previously defined (in `__init__`).
x = self.dense_1(inputs)
return self.dense_2(x)
# + [markdown] colab_type="text" id="ShDD4fv72KGc"
# Instantiate the new model class:
# + colab={} colab_type="code" id="42C-qQHm0E2l"
model = MyModel(num_classes=10)
# The compile step specifies the training configuration.
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Trains for 5 epochs.
model.fit(data, labels, batch_size=32, epochs=5)
# + [markdown] colab_type="text" id="yqRQiKj20E2o"
# ### Custom layers
#
# Create a custom layer by subclassing `tf.keras.layers.Layer` and implementing
# the following methods:
#
# * `__init__`: Optionally define sublayers to be used by this layer.
# * `build`: Create the weights of the layer. Add weights with the `add_weight`
# method.
# * `call`: Define the forward pass.
# * Optionally, a layer can be serialized by implementing the `get_config` method
# and the `from_config` class method.
#
# Here's an example of a custom layer that implements a `matmul` of an input with
# a kernel matrix:
# + colab={} colab_type="code" id="l7BFnIHr2WNc"
class MyLayer(layers.Layer):
def __init__(self, output_dim, **kwargs):
self.output_dim = output_dim
super(MyLayer, self).__init__(**kwargs)
def build(self, input_shape):
# Create a trainable weight variable for this layer.
self.kernel = self.add_weight(name='kernel',
shape=(input_shape[1], self.output_dim),
initializer='uniform',
trainable=True)
def call(self, inputs):
return tf.matmul(inputs, self.kernel)
def get_config(self):
base_config = super(MyLayer, self).get_config()
base_config['output_dim'] = self.output_dim
return base_config
@classmethod
def from_config(cls, config):
return cls(**config)
# + [markdown] colab_type="text" id="8wXDRgXV2ZrF"
# Create a model using your custom layer:
# + colab={} colab_type="code" id="uqH-cY0h0E2p"
model = tf.keras.Sequential([
MyLayer(10),
layers.Activation('softmax')])
# The compile step specifies the training configuration
model.compile(optimizer=tf.keras.optimizers.RMSprop(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# Trains for 5 epochs.
model.fit(data, labels, batch_size=32, epochs=5)
# + [markdown] colab_type="text" id="llipvR5wIl_t"
# Learn more about creating new layers and models from scratch with subclassing in the [Guide to writing layers and models from scratch](./custom_layers_and_models.ipynb).
# + [markdown] colab_type="text" id="Lu8cc3AJ0E2v"
# ## Callbacks
#
# A callback is an object passed to a model to customize and extend its behavior
# during training. You can write your own custom callback, or use the built-in
# `tf.keras.callbacks` that include:
#
# * `tf.keras.callbacks.ModelCheckpoint`: Save checkpoints of your model at
# regular intervals.
# * `tf.keras.callbacks.LearningRateScheduler`: Dynamically change the learning
# rate.
# * `tf.keras.callbacks.EarlyStopping`: Interrupt training when validation
# performance has stopped improving.
# * `tf.keras.callbacks.TensorBoard`: Monitor the model's behavior using
# [TensorBoard](https://tensorflow.org/tensorboard).
#
# To use a `tf.keras.callbacks.Callback`, pass it to the model's `fit` method:
# + colab={} colab_type="code" id="rdYwzSYV0E2v"
callbacks = [
# Interrupt training if `val_loss` stops improving for over 2 epochs
tf.keras.callbacks.EarlyStopping(patience=2, monitor='val_loss'),
# Write TensorBoard logs to `./logs` directory
tf.keras.callbacks.TensorBoard(log_dir='./logs')
]
model.fit(data, labels, batch_size=32, epochs=5, callbacks=callbacks,
validation_data=(val_data, val_labels))
# + [markdown] colab_type="text" id="ghhaGfX62abv"
#
# ## Save and restore
# + [markdown] colab_type="text" id="qnl7K-aI0E2z"
# ### Save just the weights values
#
# Save and load the weights of a model using `tf.keras.Model.save_weights`:
# + colab={} colab_type="code" id="uQIANjB94fLB"
model = tf.keras.Sequential([
layers.Dense(64, activation='relu', input_shape=(32,)),
layers.Dense(10, activation='softmax')])
model.compile(optimizer=tf.keras.optimizers.Adam(0.001),
loss='categorical_crossentropy',
metrics=['accuracy'])
# + colab={} colab_type="code" id="4eoHJ-ny0E21"
# Save weights to a TensorFlow Checkpoint file
model.save_weights('./weights/my_model')
# Restore the model's state,
# this requires a model with the same architecture.
model.load_weights('./weights/my_model')
# + [markdown] colab_type="text" id="u25Id3xe0E25"
# By default, this saves the model's weights in the
# [TensorFlow checkpoint](../checkpoints.md) file format. Weights can
# also be saved to the Keras HDF5 format (the default for the multi-backend
# implementation of Keras):
# + colab={} colab_type="code" id="JSAYoFEd0E26"
# Save weights to a HDF5 file
model.save_weights('my_model.h5', save_format='h5')
# Restore the model's state
model.load_weights('my_model.h5')
# + [markdown] colab_type="text" id="mje_yKL10E29"
# ### Save just the model configuration
#
# A model's configuration can be saved—this serializes the model architecture
# without any weights. A saved configuration can recreate and initialize the same
# model, even without the code that defined the original model. Keras supports
# JSON and YAML serialization formats:
# + colab={} colab_type="code" id="EbET0oJTzGkq"
# Serialize a model to JSON format
json_string = model.to_json()
json_string
# + colab={} colab_type="code" id="pX_badhH3yWV"
import json
import pprint
pprint.pprint(json.loads(json_string))
# + [markdown] colab_type="text" id="Q7CIa05r4yTb"
# Recreate the model (newly initialized) from the JSON:
# + colab={} colab_type="code" id="J9UFv9k00E2_"
fresh_model = tf.keras.models.model_from_json(json_string)
# + [markdown] colab_type="text" id="t5NHtICh4uHK"
# Serializing a model to YAML format requires that you install `pyyaml` *before you import TensorFlow*:
# + colab={} colab_type="code" id="aj24KB3Z36S4"
yaml_string = model.to_yaml()
print(yaml_string)
# + [markdown] colab_type="text" id="O53Kerfl43v7"
# Recreate the model from the YAML:
# + colab={} colab_type="code" id="77yRuwg03_MG"
fresh_model = tf.keras.models.model_from_yaml(yaml_string)
# + [markdown] colab_type="text" id="xPvOSSzM0E3B"
# Caution: Subclassed models are not serializable because their architecture is
# defined by the Python code in the body of the `call` method.
# + [markdown] colab_type="text" id="iu8qMwld4-71"
#
# ### Save the entire model in one file
#
# The entire model can be saved to a file that contains the weight values, the
# model's configuration, and even the optimizer's configuration. This allows you
# to checkpoint a model and resume training later—from the exact same
# state—without access to the original code.
# + colab={} colab_type="code" id="45oNY34Z0E3C"
# Create a simple model
model = tf.keras.Sequential([
layers.Dense(10, activation='softmax', input_shape=(32,)),
layers.Dense(10, activation='softmax')
])
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(data, labels, batch_size=32, epochs=5)
# Save entire model to a HDF5 file
model.save('my_model.h5')
# Recreate the exact same model, including weights and optimizer.
model = tf.keras.models.load_model('my_model.h5')
# + [markdown] colab_type="text" id="wGVBURDtI_I6"
# Learn more about saving and serialization for Keras models in the guide to [save and serialize models](./saving_and_serializing.ipynb).
# + [markdown] colab_type="text" id="PMOWhDOB0E3E"
# ## Eager execution
#
# [Eager execution](./eager.md) is an imperative programming
# environment that evaluates operations immediately. This is not required for
# Keras, but is supported by `tf.keras` and useful for inspecting your program and
# debugging.
#
# All of the `tf.keras` model-building APIs are compatible with eager execution.
# And while the `Sequential` and functional APIs can be used, eager execution
# especially benefits *model subclassing* and building *custom layers*—the APIs
# that require you to write the forward pass as code (instead of the APIs that
# create models by assembling existing layers).
#
# See the [eager execution guide](./eager.ipynb#build_a_model) for
# examples of using Keras models with custom training loops and `tf.GradientTape`.
# You can also find a complete, short example [here](https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/quickstart/advanced.ipynb).
# + [markdown] colab_type="text" id="2wG3NVco5B5V"
# ## Distribution
#
# + [markdown] colab_type="text" id="6PJZ6e9J5JHF"
# ### Multiple GPUs
#
# `tf.keras` models can run on multiple GPUs using
# `tf.distribute.Strategy`. This API provides distributed
# training on multiple GPUs with almost no changes to existing code.
#
# Currently, `tf.distribute.MirroredStrategy` is the only supported
# distribution strategy. `MirroredStrategy` does in-graph replication with
# synchronous training using all-reduce on a single machine. To use
# `distribute.Strategy`s , nest the optimizer instantiation and model construction and compilation in a `Strategy`'s `.scope()`, then
# train the model.
#
# The following example distributes a `tf.keras.Model` across multiple GPUs on a
# single machine.
#
# First, define a model inside the distributed strategy scope:
# + colab={} colab_type="code" id="sbaRr7g-0E3I"
strategy = tf.distribute.MirroredStrategy()
with strategy.scope():
model = tf.keras.Sequential()
model.add(layers.Dense(16, activation='relu', input_shape=(10,)))
model.add(layers.Dense(1, activation='sigmoid'))
optimizer = tf.keras.optimizers.SGD(0.2)
model.compile(loss='binary_crossentropy', optimizer=optimizer)
model.summary()
# + [markdown] colab_type="text" id="rO9MiL6X0E3O"
# Next, train the model on data as usual:
# + colab={} colab_type="code" id="BEwFq4PM0E3P"
x = np.random.random((1024, 10))
y = np.random.randint(2, size=(1024, 1))
x = tf.cast(x, tf.float32)
dataset = tf.data.Dataset.from_tensor_slices((x, y))
dataset = dataset.shuffle(buffer_size=1024).batch(32)
model.fit(dataset, epochs=1)
# + [markdown] colab_type="text" id="N6BXU5F90E3U"
# For more information, see the [full guide on Distributed Training in TensorFlow](../distribute_strategy.ipynb).
| site/en/guide/keras/overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''pytorch_env'': conda)'
# name: python3
# ---
# +
import numpy as np
import cv2 as cv
import os
from pathlib import Path
from imutils.paths import list_images
from sklearn.preprocessing import LabelEncoder
from tqdm import tqdm
import joblib
import pv_vision.defective_cell_detection.model.rf_train_val as rf_tool
# -
# # Load the data
# Folder structure is: \
# . \
# |-- rf_train_inference.ipynb \
# |-- segmented_cells \
# ....|-- train \
# ....|...|-- class 1 \
# ....|...|-- class 2 \
# ....|...|-- class ... \
# ....|...\`-- class n \
# ....|-- val \
# ....|...|-- class 1 \
# ....|...|-- class 2 \
# ....|...|-- class ... \
# ....|...\`-- class n \
# ....\`-- test \
# ........|-- class 1 \
# ........|-- class 2 \
# ........|-- class ... \
# ........`-- class n
#
#
# +
# load training data
images_train = []
names_train = []
labels_train = []
for im_path in list(list_images('segmented_cells/train')):
images_train.append(cv.imread(im_path, cv.IMREAD_UNCHANGED))
labels_train.append(im_path.split('/')[-2])
names_train.append(os.path.splitext(os.path.split(im_path)[-1])[0])
# load val + test data
images_val = []
names_val = []
labels_val = []
images_test = []
names_test = []
labels_test = []
for im_path in list(list_images('segmented_cells/val')):
images_val.append(cv.imread(im_path, cv.IMREAD_UNCHANGED))
labels_val.append(im_path.split('/')[-2])
names_val.append(os.path.splitext(os.path.split(im_path)[-1])[0])
for im_path in list(list_images('segmented_cells/test')):
images_test.append(cv.imread(im_path, cv.IMREAD_UNCHANGED))
labels_test.append(im_path.split('/')[-2])
names_test.append(os.path.splitext(os.path.split(im_path)[-1])[0])
images_train = np.array(images_train)
images_val = np.array(images_val)
images_test = np.array(images_test)
# transform labels into integers
le = LabelEncoder()
le.fit(labels_train)
y_train = le.transform(labels_train)
y_val = le.transform(labels_val)
y_test = le.transform(labels_test)
# -
# # Model training
# +
# convert to grayscale if original image is 3 channel grayscale
#images_train_g = np.array([image[:, :, 0] for image in images_train])
#images_val_g = np.array([image[:, :, 0] for image in images_val])
#images_test_g = np.array([image[:, :, 0] for image in images_test])
# augment the training set
images_train_aug, y_train_aug = rf_tool.im_aug(images_train, y_train)
# +
# hyperparameters for tuning
rf_para_grid = {
'n_estimators': [10, 20, 30, 40, 50, 80, 100, 200, 400, 1000],
'criterion': ['gini', 'entropy'],
'max_depth': [10, 20, 40, 60, 80, 100, None],
'bootstrap': [True, False]
}
# hyperparameters tuning in 50 iterations. Return the model with best macro F1 score on val set
rf_best, para_best, score_best = rf_tool.random_search_rf(images_train_aug, y_train_aug, images_val, y_val, 50, rf_para_grid)
# -
# save the model and optimal parameters
os.makedirs('RF', exist_ok=True)
joblib.dump(rf_best, 'RF/rf_aug_model.pkl')
joblib.dump(para_best, 'RF/rf_best_para.pkl')
# # Evaluation & Inference
import pv_vision.defective_cell_detection.result_analysis as analysis
import pickle
from sklearn import preprocessing
# +
# load model
rf_fit = joblib.load('RF/rf_aug_model.pkl')
# define save path
save_path = Path('RF')/'results'
os.makedirs(save_path, exist_ok=True)
# Do prediction on testing set
X_test = rf_tool.im_flatten(images_test)
pred_test = rf_fit.predict(X_test)
# This can output the probability of each class
#prob_test = model_fit.predict_proba(X_test)
# -
# save wrongly predicted
failed = analysis.predict_failed(y_test, pred_test, images_test)
with open(save_path/'rf_failed.pkl', 'wb') as f:
pickle.dump(failed, f)
# +
# Mapping the value of y into label names.
defect_name = {
0: 'crack',
1: 'intact',
2: 'intra',
3: 'oxygen',
4: 'solder'
}
# confusion matrix
analysis.draw_cm(defect_name, y_true=y_test, y_pred=pred_test)
# +
# metrics report
test_report = analysis.metrics_report(y_test, pred_test,
label_names=['crack', 'intact', 'intra', 'oxygen', 'solder'])
test_report.to_pickle(save_path/'rf_test_report.pkl')
# +
# save the prediction
# no need to save ground truth when you do prediction on unlabelled images
le = preprocessing.LabelEncoder()
le.fit(['crack', 'intact', 'intra', 'oxygen', 'solder'])
with open(save_path/'rf_predicted.pkl', 'wb') as f:
pickle.dump({'name': np.array(names_test),
'defects_pred': le.inverse_transform(pred_test),
'defects_true': le.inverse_transform(y_test),
'y_pred': np.array(pred_test),
'y_true': np.array(y_test)}, f)
| tutorials/old_rf_train_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# [](https://githubtocolab.com/giswqs/leafmap/blob/master/examples/notebooks/40_plotly_gui.ipynb)
# [](https://gishub.org/leafmap-binder)
#
# Uncomment the following line to install [leafmap](https://leafmap.org) if needed.
# +
# # !pip install leafmap
# -
import leafmap.plotlymap as leafmap
# **Note:** For best experience, please use Jupyter notebook. The toolbar GUI is not working very well with JupyterLab at the moment.
m = leafmap.Map()
m.add_basemap("Stamen.Terrain")
m.add_heatmap_demo()
m.add_scatter_plot_demo()
m.show()
# 
| examples/notebooks/40_plotly_gui.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %pylab inline
import pandas as pd
from pykalman import KalmanFilter
df = pd.read_csv("../data/ChungCheonDC/CompositeETCdata.csv")
df_DC = pd.read_csv("../data/ChungCheonDC/CompositeDCdata.csv")
df_DCprc = pd.read_csv("../data/ChungCheonDC/CompositeDCdata_processed.csv")
df_DCstd = pd.read_csv("../data/ChungCheonDC/CompositeDCstddata.csv")
# missininds = np.arange(df_DC[electrodeID[elecind]].values.size)[np.isnan(df_DC[electrodeID[elecind]].values)]
electrodeID = df_DC.keys()[1:-1]
# +
from scipy import interpolate
sys.path.append("../codes/")
from DCdata import readReservoirDC_all
directory = "../data/ChungCheonDC/"
dat_temp,height_temp, ID = readReservoirDC_all(directory+"20151231180000.apr")
locs = dat_temp[:,:4]
mida = locs[:,:2].sum(axis=1)
midb = locs[:,2:].sum(axis=1)
mid = (mida + midb)*0.5
dz = mida-midb
x = np.linspace(mid.min(), mid.max(), 100)
z = np.linspace(dz.min(), dz.max(), 100)
grid_x, grid_z = np.meshgrid(x,z)
def vizDCtimeSeries(idatum, itime, itime_ref, colors, flag, df_DC):
fig = plt.figure(figsize = (12, 12))
ax1 = plt.subplot(411)
ax2 = plt.subplot(412)
valsratio = df_DC[electrodeID].values[itime,:].flatten() / df_DC[electrodeID].values[itime_ref,:].flatten()
valsDC = np.log10(df_DC[electrodeID].values[itime,:].flatten())
valsDCstd = df_DCstd[electrodeID].values[itime,:].flatten()
grid_rho_ratio = griddata(mid, dz, valsratio, grid_x, grid_z, interp='linear')
grid_rho_ratio = grid_rho_ratio.reshape(grid_x.shape)
if flag =="std":
vmin, vmax = 0, 10
grid_rho = griddata(mid, dz, valsDCstd, grid_x, grid_z, interp='linear')
elif flag =="rho":
vmin, vmax = np.log10(20), np.log10(200)
grid_rho = griddata(mid, dz, valsDC, grid_x, grid_z, interp='linear')
grid_rho = grid_rho.reshape(grid_x.shape)
ax1.contourf(grid_x, grid_z, grid_rho, 200, vmin =vmin, vmax = vmax, clim=(vmin, vmax), cmap="jet")
vmin, vmax = 0.9, 1.1
ax2.contourf(grid_x, grid_z, grid_rho_ratio, 200, vmin =vmin, vmax = vmax, clim=(vmin, vmax), cmap="jet")
ax1.scatter(mid, dz, s=20, c = valsDC, edgecolor="None", vmin =vmin, vmax = vmax, clim=(vmin, vmax))
ax1.plot(mid, dz, 'k.')
ax2.scatter(mid, dz, s=20, c = valsratio, edgecolor="None", vmin =vmin, vmax = vmax, clim=(vmin, vmax))
ax2.plot(mid, dz, 'k.')
for i in range(len(colors)):
ax1.plot(mid[idatum[i]], dz[idatum[i]], 'o', color=colors[i])
ax2.plot(mid[idatum[i]], dz[idatum[i]], 'o', color=colors[i])
ax3 = plt.subplot(413)
ax3_1 = ax3.twinx()
df.plot(x='date', y='reservoirH', ax=ax3_1, color='k', linestyle='-', lw=2)
df.plot(x='date', y='upperH_med', ax=ax3_1, color='b', linestyle='-', lw=2)
df.plot(x='date', y='Temp (degree)', ax=ax3, color='r', linestyle='-', lw=2)
df.plot(x='date', y='Rainfall (mm)', ax=ax3, color='b', linestyle='-', marker="o", ms=4)
ax3.legend(loc=3, bbox_to_anchor=(1.05, 0.7))
ax3_1.legend(loc=3, bbox_to_anchor=(1.05, 0.4))
itime_ref0 = itime_ref
itime_ref1 = itime
ax3.plot(np.r_[itime_ref0, itime_ref0], np.r_[-5, 40], 'k--', lw=2)
ax3.plot(np.r_[itime_ref1, itime_ref1], np.r_[-5, 40], 'k--', lw=2)
ax4 = plt.subplot(414)
df_DC.plot(x='date', y=electrodeID[idatum], ax=ax4)
ax4.legend(loc=3, bbox_to_anchor=(1.05, 0.7))
ax4.set_yscale('log')
temp = df_DC[electrodeID[elecind]].values
vmax = np.median(temp[~np.isnan(temp)]) + np.std(temp[~np.isnan(temp)])*3
vmin = np.median(temp[~np.isnan(temp)]) - np.std(temp[~np.isnan(temp)])*3
ax4.plot(np.r_[itime_ref1, itime_ref1], np.r_[vmin, vmax], 'k--', lw=2)
ax4.plot(np.r_[itime_ref0, itime_ref0], np.r_[vmin, vmax], 'k--', lw=2)
ax4.set_ylim(vmin, vmax)
# -
ax1 = plt.subplot(111)
ax1_1 = ax1.twinx()
df.plot(figsize=(12,3), x='date', y='reservoirH', ax=ax1_1, color='k', linestyle='-', lw=2)
df.plot(figsize=(12,3), x='date', y='upperH_med', ax=ax1_1, color='b', linestyle='-', lw=2)
df.plot(figsize=(12,3), x='date', y='Temp (degree)', ax=ax1, color='r', linestyle='-', lw=2)
ax1.legend(loc=3, bbox_to_anchor=(1.05, 0.7))
ax1_1.legend(loc=3, bbox_to_anchor=(1.05, 0.4))
itime_ref0 = 255
itime_ref1 = 115
ax1.plot(np.r_[itime_ref0, itime_ref0], np.r_[-5, 35], 'k-')
ax1.plot(np.r_[itime_ref1, itime_ref1], np.r_[-5, 35], 'k-')
# print df['date'].values[itime_ref]
# +
# ax1 = plt.subplot(111)
# ax1_1 = ax1.twinx()
# df_DC.plot(figsize=(12,3), x='date', y=electrodeID[elecind], ax=ax1, colors=['k', 'b', 'r'])
# df.plot(figsize=(12,3), x='date', y='reservoirH', ax=ax1_1, color='k', linestyle='-', lw=2)
# ax1.legend(loc=3, bbox_to_anchor=(1.05, 0.7))
# ax1_1.legend(loc=3, bbox_to_anchor=(1.05, 0.4))
# ax1.set_yscale('linear')
# +
# ax1 = plt.subplot(111)
# df_DCstd.plot(figsize=(12,3), x='date', y=electrodeID[elecind], ax=ax1, colors=['k', 'b', 'r'], linestyle="-", marker='.', lw=1)
# ax1.set_yscale('log')
# ax1.legend(loc=3, bbox_to_anchor=(1.05, 0.7))
# -
txrxID = df_DC.keys()[1:-1]
xmasking = lambda x: np.ma.masked_where(np.isnan(x.values), x.values)
#x= electrodeID[elecind]
x= df_DC[txrxID]
max3 = pd.rolling_max(x, 3)
# +
# #pd.rolling_max??
# +
# plt.plot(x)
# plt.plot(max3)
# -
from ipywidgets import interact
# making matrix like max3 (but with zeros)
newdata = np.zeros_like(max3)
newdata.shape
ndata = newdata.shape[1]
for i in range(ndata):
x= df_DC[txrxID[i]]
#median10 = pd.rolling_median(x, 6)
mean10 = pd.rolling_max(x, 3)
# Masking array having NaN
xm = xmasking(mean10)
kf = KalmanFilter(transition_matrices = [1],
observation_matrices = [1],
initial_state_mean = x[0],
initial_state_covariance = 1,
observation_covariance=1,
transition_covariance=1)
# Use the observed values of the price to get a rolling mean
state_means, _ = kf.filter(xm)
newdata[:,i] = state_means.flatten()
df_DC_new = df_DC.copy()
for i,index in enumerate(txrxID):
df_DC_new.loc[:,index] = newdata[:,i].flatten()
# df_DC_new.to_csv("../data/ChungCheonDC/CompositeDCdata_processed.csv")
# +
from ipywidgets import interact, IntSlider, ToggleButtons
itime = 93
itime_ref = 86
print df['date'].values[itime]
elecind = [5, 150,200]
# vizDCtimeSeries(elecind, itime, itime_ref, ['k','b','r'])
viz = lambda idatum, itime, flag: vizDCtimeSeries([idatum], itime, itime_ref, ['r'], flag, df_DC_new)
interact(viz, idatum=IntSlider(min=0, max=379, step=1, value=294)\
,itime=IntSlider(min=0, max=360, step=1, value=200)\
,flag=ToggleButtons(options=["std", "rho"]))
# -
for i in range(0,379,100):
x= df_DC[txrxID[i]]
x1 = df_DC_new[txrxID[i]]
plt.plot(newdata[:,i], 'k')
plt.plot(x1, 'ro')
plt.plot(x, 'k.', ms=2)
plt.plot(newdata[:,i], 'k')
x1 = df_DC_new[txrxID[i]]
# +
# for index in txrxID:
# df_DC_new.loc[:,index] = newdata[:,i].flatten()
# +
i = 112
def viz(i):
x= df_DC[txrxID[i]]
#median10 = pd.rolling_median(x, 6)
mean10 = pd.rolling_max(x, 3)
#x1 = median10
#x2 = mean10
# Masking array having NaN
xm = xmasking(mean10)
# Construct a Kalman filter
# kf = KalmanFilter(transition_matrices = [1],
# observation_matrices = [1],
# initial_state_mean = x[0],
# initial_state_covariance = 1,
# observation_covariance=1,
# transition_covariance=1)
# # Use the observed values of the price to get a rolling mean
# state_means, _ = kf.filter(xm)
state_means= df_DC_new[txrxID[i]]
plt.plot(x)
plt.plot(mean10, 'k.')
#plt.plot(x1)
#plt.plot(x2)
plt.plot(state_means)
# plt.legend([ i, 'Kalman Estimate'])
# print df_DC[txrxID[i]]
interact(viz, i=(0,389,10))
# +
i = 105
x= df_DC[txrxID[i]]
#median10 = pd.rolling_median(x, 6)
mean10 = pd.rolling_max(x, 3)
#x1 = median10
#x2 = mean10
# Masking array having NaN
xm = xmasking(mean10)
# Construct a Kalman filter
kf = KalmanFilter(transition_matrices = [1],
observation_matrices = [1],
initial_state_mean = 67.6,
initial_state_covariance = 1,
observation_covariance=1,
transition_covariance=1)
# Use the observed values of the price to get a rolling mean
state_means, _ = kf.filter(xm)
#plt.plot(x1)
plt.plot(x)
#plt.plot(x1)
#plt.plot(x2)
plt.plot(state_means)
plt.legend([ 'origin x','Kalman Estimate'])
# +
i = 300
x= df_DC[txrxID[i]]
#median10 = pd.rolling_median(x, 6)
mean10 = pd.rolling_max(x, 3)
#x1 = median10
#x2 = mean10
# Masking array having NaN
xm = xmasking(mean10)
# Construct a Kalman filter
kf = KalmanFilter(transition_matrices = [1],
observation_matrices = [1],
initial_state_mean = 67.6,
initial_state_covariance = 1,
observation_covariance=1,
transition_covariance=1)
# Use the observed values of the price to get a rolling mean
state_means, _ = kf.filter(xm)
#plt.plot(x1)
plt.plot(x)
#plt.plot(x1)
#plt.plot(x2)
plt.plot(state_means)
plt.legend([ 'origin x','Kalman Estimate'])
# -
| notebook/ChengCheonData_MaxKf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (duet)
# language: python
# name: duet
# ---
from astroduet.config import Telescope
from astropy import units as u
from astroduet.utils import get_neff
duet = Telescope()
duet.neff
for tel_type in duet.config_list:
duet = Telescope(config=tel_type)
print(tel_type, duet.psf_fwhm, duet.calc_psf_fwhm())
duet.psf_fwhm
duet.pointing_rms
req_rms = 6.2 / 1.65
req_rms
duet.pointing_rms = 3.75*u.arcsec
duet.calc_psf_fwhm()
duet.psf_fwhm = 13.2 * u.arcsec
get_neff(duet.psf_fwhm, duet.pixel)
# On axis values:
duet = Telescope()
duet.psf_params['sig'] = [2.36 * u.arcsec]
duet.calc_psf_fwhm()
| notebooks/PSF Sanity Check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from mpl_toolkits.mplot3d import Axes3D
#svd 奇异值分解
A = np.random.randint(-10,100,size=(16,8))
U,s,Vt = np.linalg.svd(A)
U.shape,Vt.shape
print(s,len(s))
plt.imshow(A)
plt.subplot(1,3,1)
plt.imshow(U)
plt.subplot(1,3,2)
plt.imshow(np.diag(s))
plt.subplot(1,3,3)
plt.imshow(Vt)
#svd压缩还原 选择前1,3,5维
selt = [1,3,5]
out = []
for dim in selt:
sigma = np.diag(s[:dim])
Ud = U[:,:dim]
Vtd = Vt[:dim]
As = np.dot(np.dot(Ud,sigma),Vtd)
out.append(As)
plt.subplot(1,4,1)
plt.title('Original')
plt.imshow(A)
plt.subplot(1,4,2)
plt.title("1-ndim")
plt.imshow(out[0])
plt.subplot(1,4,3)
plt.title("3-ndim")
plt.imshow(out[1])
plt.subplot(1,4,4)
plt.title("5-ndim")
plt.imshow(out[2])
# +
#完整还原
sigma = np.zeros((U.shape[0],Vt.shape[0]))
for i in range(len(s)):
sigma[i,i] = s[i]
As = np.dot(np.dot(U,sigma),Vt)
plt.subplot(1,2,1)
plt.title('Original')
plt.imshow(A)
plt.subplot(1,2,2)
plt.title("Recovery")
plt.imshow(As)
# +
#PCA主成分分析-降维算法
A = np.random.randn(1900,2)
G1 = np.random.rand(400,1)
G2 = np.subtract(np.array([[2]*400]).T,G1)
G3 = np.random.rand(900,1)
G4 = np.subtract(np.array([[5]*900]).T,G3)
G5 = np.random.rand(600,1)
G6 = np.subtract(np.array([[1]*600]).T,G5)
G = np.concatenate((G1,G3,G5),axis=0)
Gn = np.concatenate((G2,G4,G6),axis=0)
A = np.concatenate((A,G),axis=1)
A = np.concatenate((A,Gn),axis=1)
label = [236]*400+[36]*900+[178]*600
trans = np.random.randint(-9,8,size=(4,4))
h = np.dot(A,trans)
#h = A
#h = np.concatenate((h,np.array([label]).T),axis=1)
fig = plt.figure()
ax = Axes3D(fig)
ax.scatter(h[:,0],h[:,1],h[:,2],c=label)
plt.show()
# -
for i,(x,y) in enumerate([(0,1),(1,2),(0,2)]):#,(1,3),(0,3),(2,3)]):
plt.subplot(1,3,i+1)
plt.scatter(h[:,x],h[:,y],c=label)
plt.show()
pca = PCA()
s = pca.fit_transform(h)
s.shape
for i,(x,y) in enumerate([(0,1),(1,2),(0,2)]):
plt.subplot(1,3,i+1)
plt.scatter(s[:,x],s[:,y],c=label)
plt.show()
fig = plt.figure()
ax = Axes3D(fig)
ax.scatter(s[:,0],s[:,1],s[:,2],c=label)
plt.show()
plt.scatter(s[:,0],s[:,2],c=label)
pca = PCA(n_components=2)
s = pca.fit_transform(h)
plt.scatter(s[:,0],s[:,1],c=label)
pca = PCA(n_components=1)
s = pca.fit_transform(h)
plt.scatter(s[:,0],[1]*1900,c=label)
plt.scatter(A[:,3],[1]*1900,c=label)
| Svd and PCA demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://qworld.net" target="_blank" align="left"><img src="../qworld/images/header.jpg" align="left"></a>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# $ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
# $ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
# $ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
# $ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
# $ \newcommand{\blackbit}[1] {\mathbf{{\color{black}#1}}} $
# <font style="font-size:28px;" align="left"><b> Visualization of a (Real-Valued) Qubit </b></font>
# <br>
# _prepared by <NAME>_
# <br><br>
# [<img src="../qworld/images/watch_lecture.jpg" align="left">](https://youtu.be/p4HjmbAmUh8)
# <br><br><br>
# _We use certain tools from python library "<b>matplotlib.pyplot</b>" for drawing. Check the notebook [Python: Drawing](../python/Python06_Drawing.ipynb) for the list of these tools._
# Suppose that we have a single qubit.
#
# Each possible (real-valued) quantum state of this qubit is a point on 2-dimensional space.
#
# It can also be represented as a vector from origin to that point.
#
# We start with the visual representation of the following quantum states:
#
# $$ \ket{0} = \myvector{1\\0}, ~~ \ket{1} = \myvector{0\\1} , ~~ -\ket{0} = \myrvector{-1\\0}, ~~\mbox{and}~~ -\ket{1} = \myrvector{0\\-1}. $$
# We draw these quantum states as points.
#
# We use one of our predefined functions for drawing axes: "draw_axes()". We include our predefined functions with the following line of code:
#
# # %run quantum.py
# +
# import the drawing methods
from matplotlib.pyplot import plot, figure, show
# draw a figure
figure(figsize=(6,6), dpi=80)
# include our predefined functions
# %run quantum.py
# draw the axes
draw_axes()
# draw the origin
plot(0,0,'ro') # a point in red color
# draw these quantum states as points (in blue, green, yellow, and cyan colors)
plot(1,0,'bo')
plot(0,1,'go')
plot(-1,0,'yo')
plot(0,-1,'co')
show()
# -
# Now, we draw the quantum states as arrows (vectors):
# +
# import the drawing methods
from matplotlib.pyplot import figure, arrow, show
# draw a figure
figure(figsize=(6,6), dpi=80)
# include our predefined functions
# %run quantum.py
# draw the axes
draw_axes()
# draw the quantum states as vectors (in red, blue, green, and yellow colors)
arrow(0,0,0.92,0,head_width=0.04, head_length=0.08, color="r")
arrow(0,0,0,0.92,head_width=0.04, head_length=0.08, color="b")
arrow(0,0,-0.92,0,head_width=0.04, head_length=0.08, color="g")
arrow(0,0,0,-0.92,head_width=0.04, head_length=0.08, color="y")
show()
# -
# <h3> Task 1 </h3>
#
# Write a function that returns a randomly created 2-dimensional (real-valued) quantum state.
#
# _You can use your code written for [a task given in notebook "Quantum State](B28_Quantum_State.ipynb#task2)._
#
# Create 100 random quantum states by using your function and then draw all of them as points.
#
# Create 1000 random quantum states by using your function and then draw all of them as points.
#
# The different colors can be used when drawing the points ([matplotlib.colors](https://matplotlib.org/2.0.2/api/colors_api.html)).
# +
# import the drawing methods
from matplotlib.pyplot import plot, figure
# draw a figure
figure(figsize=(6,6), dpi=60)
# draw the origin
plot(0,0,'ro')
from random import randrange
colors = ['ro','bo','go','yo','co','mo','ko']
#
# your solution is here
#
# -
# <a href="Q32_Visualization_of_a_Qubit_Solutions.ipynb#task1">click for our solution</a>
# <h3> Task 2 </h3>
#
# Repeat the previous task by drawing the quantum states as vectors (arrows) instead of points.
#
# The different colors can be used when drawing the points ([matplotlib.colors](https://matplotlib.org/2.0.2/api/colors_api.html)).
#
# _Please keep the codes below for drawing axes for getting a better visual focus._
# +
# import the drawing methods
from matplotlib.pyplot import plot, figure, arrow
# draw a figure
figure(figsize=(6,6), dpi=60)
# include our predefined functions
# %run quantum.py
# draw the axes
draw_axes()
# draw the origin
plot(0,0,'ro')
from random import randrange
colors = ['r','b','g','y','b','c','m']
#
# your solution is here
#
# -
# <a href="Q32_Visualization_of_a_Qubit_Solutions.ipynb#task2">click for our solution</a>
# <h3> Unit circle </h3>
#
# All (real-valued) quantum states of a qubit form the unit circle.
#
# The length of each quantum state is 1.
#
# All points that are 1 unit away from the origin form the circle with radius 1 unit.
#
# We can draw the unit circle with python.
#
# We have a predefined function for drawing the unit circle: "draw_unit_circle()".
# +
# define a figure
from matplotlib.pyplot import figure
figure(figsize=(6,6), dpi=80) # size of the figure
# include our predefined functions
# %run quantum.py
# draw axes
draw_axes()
# draw the unit circle
draw_unit_circle()
# show the diagram
show_plt()
# -
# <h3>Quantum state of a qubit</h3>
# Suppose that we have a single qubit.
#
# Each possible (real-valued) quantum state of this qubit is a point on 2-dimensional space.
#
# It can also be represented as a vector from origin to that point.
#
# We draw the quantum state $ \myvector{3/5 \\ 4/5} $ and its elements.
# <i style="font-size:10pt;">
# Our predefined function "draw_qubit()" draws a figure, the origin, the axes, the unit circle, and base quantum states.
# <br>
# Our predefined function "draw_quantum_state(x,y,name)" draws an arrow from (0,0) to (x,y) and associates it with <u>name</u>.
# <br>
# We include our predefined functions with the following line of code:
#
# # %run quantum.py
# </i>
# +
# %run quantum.py
draw_qubit()
draw_quantum_state(3/5,4/5,"|v>")
show_plt()
# -
# Now, we draw its angle with $ \ket{0} $-axis and its projections on both axes.
#
# <i> For drawing the angle, we use the method "Arc" from library "matplotlib.patches". </i>
# +
# %run quantum.py
draw_qubit()
draw_quantum_state(3/5,4/5,"|v>")
from matplotlib.pyplot import arrow, text, gca
# the projection on |0>-axis
arrow(0,0,3/5,0,color="blue",linewidth=1.5)
arrow(0,4/5,3/5,0,color="blue",linestyle='dotted')
text(0.1,-0.1,"cos(a)=3/5")
# the projection on |1>-axis
arrow(0,0,0,4/5,color="blue",linewidth=1.5)
arrow(3/5,0,0,4/5,color="blue",linestyle='dotted')
text(-0.1,0.55,"sin(a)=4/5",rotation="90")
# drawing the angle with |0>-axis
from matplotlib.patches import Arc
gca().add_patch( Arc((0,0),0.4,0.4,angle=0,theta1=0,theta2=53) )
text(0.08,0.05,'.',fontsize=30)
text(0.21,0.09,'a')
# -
# <b> Observations: </b>
# <ul>
# <li> The angle of quantum state with state $ \ket{0} $ is $a$.</li>
# <li> The amplitude of state $ \ket{0} $ is $ \cos(a) = \frac{3}{5} $.</li>
# <li> The probability of observing state $ \ket{0} $ is $ \cos^2(a) = \frac{9}{25} $.</li>
# <li> The amplitude of state $ \ket{1} $ is $ \sin(a) = \frac{4}{5} $.</li>
# <li> The probability of observing state $ \ket{1} $ is $ \sin^2(a) = \frac{16}{25} $.</li>
# </ul>
# <h3> The angle of a quantum state </h3>
#
# The angle of a vector (in radians) on the unit circle is the length of arc in counter-clockwise direction that starts from $ (1,0) $ and with the points representing the vector.
#
# We execute the following code a couple of times to see different examples, where the angle is picked randomly in each run.
#
# You can also set the value of "myangle" manually for seeing a specific angle.
# +
# set the angle
from random import randrange
myangle = randrange(361)
################################################
from matplotlib.pyplot import figure,gca
from matplotlib.patches import Arc
from math import sin,cos,pi
# draw a figure
figure(figsize=(6,6), dpi=60)
# %run quantum.py
draw_axes()
print("the selected angle is",myangle,"degrees")
ratio_of_arc = ((1000*myangle/360)//1)/1000
print("it is",ratio_of_arc,"of a full circle")
print("its length is",ratio_of_arc,"x 2\u03C0","=",ratio_of_arc*2*pi)
myangle_in_radian = 2*pi*(myangle/360)
print("its radian value is",myangle_in_radian)
gca().add_patch( Arc((0,0),0.2,0.2,angle=0,theta1=0,theta2=myangle,color="red",linewidth=2) )
gca().add_patch( Arc((0,0),2,2,angle=0,theta1=0,theta2=myangle,color="brown",linewidth=2) )
x = cos(myangle_in_radian)
y = sin(myangle_in_radian)
draw_quantum_state(x,y,"|v>")
# the projection on |0>-axis
arrow(0,0,x,0,color="blue",linewidth=1)
arrow(0,y,x,0,color="blue",linestyle='dashed')
# the projection on |1>-axis
arrow(0,0,0,y,color="blue",linewidth=1)
arrow(x,0,0,y,color="blue",linestyle='dashed')
print()
print("the amplitude of state |0> is",x)
print("the amplitude of state |1> is",y)
print()
print("the probability of observing state |0> is",x*x)
print("the probability of observing state |1> is",y*y)
print("the total probability is",round(x*x+y*y,6))
# -
# <h3> Random quantum states </h3>
#
# Any quantum state of a (real-valued) qubit is a point on the unit circle.
#
# We use this fact to create random quantum states by picking a random point on the unit circle.
#
# For this purpose, we randomly pick an angle between zero and 360 degrees and then find the amplitudes of the quantum state by using the basic trigonometric functions.
# <a id="task3"></a>
# <h3> Task 3 </h3>
#
# Define a function randomly creating a quantum state based on this idea.
#
# Randomly create a quantum state by using this function.
#
# Draw the quantum state on the unit circle.
#
# Repeat the task for a few times.
#
# Randomly create 100 quantum states and draw all of them.
# <i> You can save your function for using later: comment out the first command, give an appropriate file name, and then run the cell.</i>
# # %%writefile FILENAME.py
# your function is here
from math import cos, sin, pi
from random import randrange
def random_qstate_by_angle():
#
# your codes are here
#
# <i style="font-size:10pt;">
# Our predefined function "draw_qubit()" draws a figure, the origin, the axes, the unit circle, and base quantum states.
# <br>
# Our predefined function "draw_quantum_state(x,y,name)" draws an arrow from (0,0) to (x,y) and associates it with <u>name</u>.
# <br>
# We include our predefined functions with the following line of code:
#
# # %run quantum.py
# </i>
# +
# visually test your function
# %run quantum.py
draw_qubit()
#
# your solution is here
#
# draw_quantum_state(x,y,"")
# -
# <a href="Q32_Visualization_of_a_Qubit_Solutions.ipynb#task3">click for our solution</a>
| quantum-with-qiskit/Q32_Visualization_of_a_Qubit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Tutorial Setup
# ### Check your install
import numpy
import matplotlib
import sklearn
import pandas
# Finding the location of an installed package and its version:
numpy.__path__
numpy.__version__
# Or check it all at once: pip install version_information and check versions with a magic command.
# !pip install version_information
# %load_ext version_information
# %version_information numpy, scipy, matplotlib, pandas, tensorflow, sklearn, skflow
# ## A NumPy primer
# ### NumPy array dtypes and shapes
import numpy as np
a = np.array([1, 2, 3])
a
b = np.array([[0, 2, 4], [1, 3, 5]])
b
b.shape
b.dtype
a.shape
a.dtype
np.zeros(5)
np.ones(shape=(3, 4), dtype=np.int32)
# ### Common array operations
c = b * 0.5
c
c.shape
c.dtype
a
d = a + c
d
d[0]
d[0, 0]
d[:, 0]
d.sum()
d.mean()
d.sum(axis=0)
d.mean(axis=1)
# ### Reshaping and inplace update
e = np.arange(12)
e
f = e.reshape(3, 4)
f
e
e[5:] = 0
e
f
# ### Combining arrays
a
b
d
np.concatenate([a, a, a])
np.vstack([a, b, d])
np.hstack([b, d])
# Also see this fun "100 numpy exercises" [site](https://github.com/rougier/numpy-100)
# ## A Matplotlib primer
# %matplotlib inline
import matplotlib.pyplot as plt
x = np.linspace(0, 2, 10)
x
plt.plot(x, 'o-');
# +
plt.plot(x, x, 'o-', label='linear')
plt.plot(x, x ** 2, 'x-', label='quadratic')
plt.legend(loc='best')
plt.title('Linear vs Quadratic progression')
plt.xlabel('Input')
plt.ylabel('Output');
# -
samples = np.random.normal(loc=1.0, scale=0.5, size=1000)
samples.shape
samples.dtype
samples[:30]
plt.hist(samples, bins=50);
samples_1 = np.random.normal(loc=1, scale=.5, size=10000)
samples_2 = np.random.standard_t(df=10, size=10000)
bins = np.linspace(-3, 3, 50)
_ = plt.hist(samples_1, bins=bins, alpha=0.5, label='samples 1')
_ = plt.hist(samples_2, bins=bins, alpha=0.5, label='samples 2')
plt.legend(loc='upper left');
plt.scatter(samples_1, samples_2, alpha=0.1)
samples_3 = np.random.normal(loc=2, scale=.5, size=10000)
fig = plt.figure()
ax1 = fig.add_subplot(111)
ax1.scatter(samples_1, samples_2, alpha=0.1, c='b', marker="s", label='first')
ax1.scatter(samples_3, samples_2, alpha=0.1, c='r', marker="o", label='second')
plt.show()
# Credits
# =======
#
# Most of this material is adapted from the Olivier Grisel's 2015 tutorial:
#
# [https://github.com/ogrisel/parallel_ml_tutorial](https://github.com/ogrisel/parallel_ml_tutorial)
#
# Original author:
#
# - <NAME> [@ogrisel](https://twitter.com/ogrisel) | http://ogrisel.com
#
| 00.Setup and Primers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### pandas索引
import pandas as pd
df = pd.read_csv('titanic_train.csv')
df.head()
df['Age'][:5]
df[['Age', 'Fare']][:5]
# * loc 用label来去定位
# * iloc 用position进行定位
df.iloc[0:5, 1:3]
df = df.set_index('Name')
df.loc['Heikkinen, Miss. Laina']
df.loc['Heikkinen, Miss. Laina', 'Fare']
df.loc['Heikkinen, Miss. Laina':'Allen, Mr. <NAME>',:]
df.loc['Heikkinen, Miss. Laina', 'Fare'] = 1000
df.head()
# deprecationWarning
df.ix['Heikkinen, Miss. Laina', 'Fare']
# bool类型的索引
df['Fare'] > 40
df[df['Fare'] > 40]
df[df['Sex'] == 'male'][:5]
df.loc[df['Sex'] =='male', 'Age'].mean()
sum(df['Age'] > 70)
| pandas_demo/pandas_index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prevendo Turnover
# 
# ## Uma empresa está sofrendo com muitos gastos relacionados a Turnover, sua missão é ajudar a organização a prever possíveis 'saintes' e também encontrar fatores que possam estar levando essas pessoas a sairem da empresa
# ### É esperado três entregas:
#
# * Um algoritmo capaz de prever se um funcionário vai sair da empresa com pelo menos 99% de acurácia
# * (Considerando que você separe pelo menos 30% dos dados para teste)
# * Uma Análise exploratória dos dados
# * (escolha no máximo 3 gráficos que lhe ajudem a 'contar uma história' sobre a situação atual)
# * Um modelo de previsão mais simples mas que seja de fácil entendimento pelos gestores
# # Good Job!
#
# 
| curso/4 - Machine Learning/Trees/Projeto HR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <font size=6>
#
# <b>Curso de Programación en Python</b>
# </font>
#
# <font size=4>
#
# Curso de formación interna, CIEMAT. <br/>
# Madrid, Octubre de 2021
#
# <NAME>
# </font>
#
# https://github.com/andelpe/curso-intro-python/
#
# <br/>
# -
# # Tema 6 - Funciones y módulos
# ## Objetivos
#
# - Aprender a definir funciones y utilizarlas
#
# - Entender las diferentes maneras de pasar argumentos a una función
#
#
# - Conocer la creación y uso de módulos y paquetes
#
# - Introducir el _scope_ (alcance) y _namespaces_ de los objetos Python, en particular para funciones y módulos
#
# - Conocer el uso de los _docstrings_ para documentar código Python
#
# ## Funciones en Python
#
# Una función es un bloque de instrucciones que se ejecutan cuando la función es llamada.
#
# - Permiten reutilizar código, sin tener que reescribirlo.
# - Son esenciales para cualquier programa no trivial.
#
# Una función se define con la sentencia `def`:
#
# def mi_funcion(arg1, arg2, ...):
# instruccion
# instruccion
#
# - La ejecución de la sentencia `def` crear un nuevo _objeto función_ ligado al nombre `mi_funcion`.
# - El _cuerpo_ de la función no se interpreta hasta que la función es usada: `mi_funcion(args)`
#
# En el cuerpo de la función:
#
# - Los identificadores de argumentos se pueden usar como variables locales
# - La sentencia `return` especifica el valor devuelto por la función (por defecto es `None`)
# +
def suma(x, y):
res = x + y
return res
s = suma(3, 4)
print(s)
# -
# <div style="background-color:powderblue;">
#
# **EJERCICIO e6_1:**
#
# - Crear una función que acepte un argumento numérico y que devuelva el doble del valor pasado.
# - Probarla con las siguientes entradas: `2`, `-10.0`, `'abcd'`
# ### Argumentos de funciones
#
# Los argumentos de una función de Python se _pasan por asignación_ (equivalente a _por referencia_).
#
# - El valor pasado _se asigna_ a una variable local (no se hace una copia)
# - Si el valor es modificable, y se modifica, la variable externa verá el mismo cambio
# - Como las variables no tienen tipo, tampoco lo tienen los argumentos de una función
# +
def addElem(var, val):
print("\n-- 'var' interna original:", var)
var.append(val)
print("-- 'var' interna modificada:", var)
nums = [0, 1]
print("Variable externa 'nums' antes:", nums)
addElem(nums, 2)
print("\nVariable externa 'nums' después:", nums)
# +
def autoSuma(var, amount):
print("\n-- 'var' interna original:", var)
var += amount
print("-- 'var' interna modificada:", var)
x = 3
print("Variable externa 'x' antes:", x)
autoSuma(x, 5)
print("\nVariable externa 'x' después:", x)
# -
# #### Formas de pasar argumentos
#
# - Por posición: `f(3, 4)`
# - Nombrados: `f(x=3, y=4)`
# - Expansión:
# - `f(*(3, 4))` equivale a `f(3, 4)`
# - `f(**{x:3, y:4}` equivale a `f(x=3, y=4)`
#
# Si se combinan, el orden siempre debe respetar:
# - Nombrados después de posición
# - Expandidos después de no expandidos.
#
# P.ej.:
#
# f(1, z=3, y=3)
# f(1 *[2,3])
# f(1, *mytuple, w=10, **mydict)
# +
def f(a1, a2, a3):
print(f'a1: {a1} a2: {a2} a3: {a3}\n')
f(3, *(4,5))
f(3, 4, a3=5)
f(*(3,4), a3=5)
f(3, **{'a2':4, 'a3':5})
# -
# #### Formas de recoger argumentos
#
# - Argumentos con valores por defecto (si no son especificados por el llamante):
# +
def f1(a1, a2=0):
print(f'a1: {a1} a2: {a2}')
f1(3, 4)
f1(3)
# -
# - Resto de argumentos recogidos en una tupla:
# +
def f2(a1, *rest):
print(f'a1: {a1} rest: {rest}')
f2(3, 4, 5)
f2(3)
# -
# - Resto de argumentos _nombrados_ recogidos en un diccionario:
# +
def f3(a1, **rest):
print(f'a1: {a1} rest: {rest}')
f3(a1=3, y=5, x=4)
f3(3, x=4, y=5)
f3(3)
# +
def f3(a1, *vrest, **drest):
print(f'a1: {a1} vrest: {vrest} drest: {drest}')
f3(3, 4, 5, pepe=8)
# -
# ### Polimorfismo
#
# En Python, los argumentos de una función no tienen tipo, por lo que no tiene sentido tener diferentes definiciones de la función para diferentes tipos de argumentos (como sucede en otros lenguajes: _sobrecarga_).
#
# Para que nuestra función soporte diferentes argumentos solo se requiere... usarlo.
#
# - _Duck type: If it walks like a duck and quacks like a duck..._
#
# Esta filosofía gusta a unos más y a otros menos, pero es una herramienta muy potente
#
# - ¡Es importante documentar bien las funciones!
# +
def muestra(iterable):
print(' -- EN muestra --')
for i, x in enumerate(iterable):
if i > 3: break
print(str(x).strip())
muestra( ['a', 2, (3,3), 4 ])
muestra( 'astring' )
muestra( open('README.md') )
# -
# <div style="background-color:powderblue;">
#
# **EJERCICIO e6_2:**
#
# - Utilizar el código de los ejercicios e5_3 y e5_4, para crear dos funciones `readRows(lines)` y `readCols(lines)`, que aceptan un iterable (p.ej., lista de líneas, o objeto fichero), y devuelven un diccionario con los datos.
# <div style="background-color:powderblue;">
#
# **EJERCICIO e6_3:**
#
# - Implementar una función `readData(fname, cols=False)`, que acepta un nombre de fichero, y, opcionalmente, un _boolean_, que indica si se lee por filas (por defecto), o columans (`cols=True`). La función `readData` hará uso de las funciones del ejercicio e6_2, para devolver un diccionario con los datos.
# <div style="background-color:powderblue;">
#
# **EJERCICIO e6_4:**
#
# Crear una función `showTotals(fname, cols=False, **kwd)`, que llama a `readData` del ejercicio e6_3, y muestra la suma para cada clave presente. Además, si `mult=X` aparece en `kwd`, multiplica las sumas por `X`.
# ### Funciones como objetos
#
# La definición de una función crea un objeto función.
#
# - No confundir la función, con el resultado de su invocación
#
# Un objeto función (como cualquier otro objeto) puede copiarse, pasarse como argumento, devolverse con `return`, etc.
#
# - Paradigma de programación funcional con Python
# +
# Function that creates and returns a new function
def funcFactory(x):
print(f"--Fijamos x={x} y creamos una f({x}, y)")
def f(y): print(f'{x} * {y} = {x*y}')
return f
# Function that receives a function as argument, and calls it
def funcCaller(func):
print(f"--Llamamos a 'func(y)' con y=4")
func(4)
# Produce a function with fixed x=3
myfunc = funcFactory(3)
# Assign the function
a = myfunc
# Call the function with y=5
a(5)
# Pass the function as argument (it will be called with y=4)
funcCaller(a)
# -
# <div style="background-color:powderblue;">
#
# **EJERCICIO e6_5:**
#
# Definir una función `func`, tal que dado el iterable `v`, la expresión `sorted(v, key=func)` devuelva `v` ordenado por la longitud de sus elementos.
#
# - Nota: `key` espera una función que se aplica a cada elemento de `v` antes de ordenar.
#
# Por ejemplo:
#
# ```python
# v = ['a', 'bbb', 'xx']
# sorted(v, key=func) ---> ['a','xx','bbb']
# ```
#
# <div style="background-color:powderblue;">
#
# A continuación, buscar otra `func` que ordene un iterable por la longitud del segundo elemento de cada miembro. P. ej.:
#
# ```python
# v = [(0, 'a'), (1, 'bbb'), (2, 'xx')]
# sorted(v, key=func) ---> [(0, 'a'), (2, 'xx'), (1, 'bbb')]
# ```
v = 'a', 'xx', 'bbb'
sorted(v)
v = ['a', 'bbb', 'xx']
# fill this
v = [(0, 'a'), (1, 'bbb'), (2, 'xx')]
# fill this
# ### Recursividad
#
# - Una función puede llamarse a sí misma
# - Funciona igual que en cualquier otro lenguaje
# - Se necesita una condición de salida que siempre se alcance
# +
def factorial(x):
if x < 2:
return 1
else:
return x * factorial(x-1)
print(factorial(5))
# -
# <div style="background-color:powderblue;">
#
# **EJERCICIO e6_6:** Recursividad.
#
# Definir una función que busca un path entre 2 nodos de un grafo (definido según el ejercicio e4_3. Su firma será:
# ```python
# find_path(grafo, start, end, path = [])
# ```
#
# - Donde `path` es el camino ya recorrido (en una llamada inicial, simplemente se omite).
#
# Se puede comprobar el resultado (buscar un camino posible entre dos nodos) con:
#
# ```python
# import modulos.graph_plot as gplt
# gplt.plotAll(grafo, path)
# ```
#
# - Donde `path` es un iterable con dos elementos, el nodo de inicio y el de final: `(start, end)`
# ## Namespace y scope
#
# Los _namespaces_ dividen el conjunto de identificadores de objetos, de manera que sea posible repetir el mismo nombre en dos espacios independientes, sin que haya colisión.
#
# - Es análogo a como uno puede tener dos ficheros con el mismo nombre si están en directorios diferentes.
#
# Python define muchos espacios de nombres diferentes.
#
# - P.ej. existe un espacio de nombres para los objetos _built-in_, así como uno para cada módulo.
# - Cada función Python define su propio espacio de nombres para sus variables (por tanto, son locales).
#
# Una variable siempre puede identificarse como: `namespace.identificador`, p.ej:
#
# math.log
# __builtins__.print
#
# Un concepto relacionado es el de _scope_ (alcance). El _scope_ de un identificador (variable) es en qué partes del programa es accesible, sin usar un prefijo (indicando su namespace).
#
# - Las variables _built-in_ están siempre accesibles
# - Las variables del espacio de nombres global de un módulo son accesibles dentro de ese módulo
# - Las variables locales a una función (incluidos los argumentos) solo son accesible desde el propio cuerpo de la función
#
# Si no se especifica el namespace, una variable se busca primero en el local, luego en el módulo (global), y luego en _built-in_.
#
# - La sentencia `global <variable>` permite indicar que nos referimos a la variable global, y no a la local
# +
a = 0
b = 1
def func1(x): # x local (param)
a = b + x # a local, b global (lectura)
print("func1: a =", a)
def func2(x):
global b # b global
b = a # a global (lectura)
print("func2: b =", b)
func1(2)
func2(2)
print("out: a =", a)
print("out: b =", b)
# -
# ## Módulos y paquetes
#
# ### Módulos
#
# Un módulo es un fichero que agrupa código Python, principalmente definiciones de objetos, para su reutilización.
#
# - El caso más habitual es que el módulo `foo` corresponda al fichero `foo.py`
#
# - Nota: también existen módulos de código compilados de _C_: `foo.so`
#
# Un módulo crea su propio espacio de nombres, que se hace accesible al importar el módulo.
#
# - Y accedemos a sus objetos con la notación `modulo.objeto`
# +
import math # Ligamos el identificador 'math' al namespace del módulo
print(math.pi) # Accedemos al objeto `pi` en ese namespace
import math as mod # Ligamos el identificador 'mod' al namespace del módulo 'math'
print(mod.pi)
from math import pi as PI # Ligamos el id 'PI' al objeto 'pi' del módulo 'math'
print(PI)
# -
# Los módulos también pueden ejecutar otro tipo de instrucciones, además de las de asignación (creación de objetos).
#
# - Cualquier instrucción contenida en el módulo se ejecuta cuando se llama a `import` (pero solo la primera vez)
# - Esto permite incluir código de inicialización, o utilizar un `.py` a la vez como módulo o como script
# Ejemplo de módulo (contenidos de `modulos/samplemod.py`)
#
# ```python
# print('Loading...')
#
# def double(x):
# return 2*x
#
# if __name__ == "__main__":
# import sys
# print('Tu valor doblado:', float(sys.argv[1]))
# ```
#
# Si lo importamos, veremos el resultado del `print`, y podremos usar `double`.
import modulos.samplemod
modulos.samplemod.double(34)
# A continuación, lanzamos el `.py` como un script:
# !python modulos/samplemod.py 34
# También se puede ver ejecutando `python modulos/samplemod.py 34` en una terminal.
# #### Módulos y bytecodes
#
# Cuando un módulo (o un script) se usa por primera vez, Python lo compila a _bytecodes_, y genera un fichero `.pyc`.
#
# Para acelerar las ejecuciones, si el fichero no se modifica, las siguientes veces que se utilice el módulo, Python ejecutará directamente el código pre-compilado, en lugar de generarlo de nuevo.
# ### Paquetes
#
# Los paquetes son agrupaciones de módulos.
#
# - Físicamente, se corresponden con directorios que albergan ficheros `.py`
#
# mypack/__init__.py
# mypack/mymod1.py
# mypack/mymod2.py
# mypack/subpack/__init__.py
# mypack/subpack/mymod1.py
#
#
# - Desde el punto de vista lógico, organizan jerárquicamente los namespaces:
#
# ```python
# # Import a module preserving namespace path
# import mypack.mymod1
# mypack.mymod1.some_function()
#
# # Import a module into our namespace
# from mypack.subpack import mymod1
# mymod1.other_func()
#
# # Import a function from within a module
# from mypack.subpack.mymod1 import other_func
# other_func()
# ```
# Para que un directorio se considere un paquete (puedan importarse módulos de él), debe albergar el fichero `__init__.py` aunque sea vacío.
#
# - `__init__.py` puede contener código de inicialización
#
# - También pueden configurar los efectos de `import mypack`/`from mypack import *`
# - Por defecto, no importarán nada (supondría un riesgo, y es mala práctica, por contaminar el namespace)
# - Se puede ver un ejemplo en: `modulos/__init__.py` y `modulos/pack/__init__.py`
import modulos
modulos.submod.f(3)
# + [markdown] tags=[]
# ### Búsqueda de módulos
#
# Cuando se usa la instrucción `import`, el módulo se busca primero en los _built-in_.
#
# Si no se encuentra, se busca en los directorios contenidos en la variable `sys.path`. Esta variable contiene:
#
# - El directorio del script en ejecución
# - Los dirs de la variable de entorno `PYTHONPATH`
# - Los dirs por defecto de la instalación (librerías del sistema)
#
# Notas:
#
# - Si definimos módulos con el mismo nombre que los del sistema (en el dir actual o en `PYTHONPATH`), ocultaremos los del sistema.
# - La variable `sys.path` se puede variar en ejecución.
# -
import sys
for dir in sys.path: print(dir)
# ### Recargar módulos
#
# Por defecto, los módulos solo se cargan una vez (_it's a feature!_).
#
# Pero si estamos desarrollando código, y probándolo, quizás queramos recargarlos cuando realicemos cambios. Se puede hacer con:
#
# ```python
# import importlib
# importlib.reload(module)
# ```
#
# De hecho, con Jupyter (o Ipython), el cambio puede ser automático cuando haya cambios, con:
#
# ```python
# # %load_ext autoreload
# # %autoreload 2
# ```
# +
import modulos.samplemod
print('Nothing happened.\n\nNow, see:')
import importlib
mod = importlib.reload(modulos.samplemod)
# -
# ### Sobre Jupyter y los módulos
#
# Aunque Jupyter es un entorno muy potente, cuando uno desarrolla código _en serio_, es conveniente (al menos, es mi opinión) ir moviendo el código a módulos Python (`.py`, no `.ipynb`), de manera que sea fácilmente usable en otros programas, u otras personas, o incluso ejecutable como script.
#
# Igualmente, es muy recomendable usar control de versiones, como Git (incluso con los notebooks: extensión _@jupyterlab/git_)
# ## Docstrings
#
# - Sirven para documentar python
# - Cualquier string comenzando módulos, clases, funciones, se considera documentación
# - Es lo que vemos con `help()` (también hay herramientas para generar html...)
#
# ¡Es muy importante documentar el código!
# +
def funcFactory(x):
"""
Creates and returns a new function that multiplies its argument by 'x'.
"""
def f(y): print(f'{x} * {y} = {x*y}')
return f
help(funcFactory)
# -
help(modulos.samplemod)
# <div style="background-color:powderblue;">
#
# **EJERCICIO e6_7:**
#
# Crear un módulo en nuevo fichero `mydata.py`, que agrupe las funciones del ejercicio e6_2, e6_3 y e6_4.
#
# - El módulo solo debe mostrar como interfaz las funciones `readData` y `showTotals`. Cualquier otra función se considera auxiliar, y se debe marcar como privada nombrándola con un `_` inicial.
#
# - Incluir un _docstring_ al principio del módulo.
#
# Colocar el nuevo módulo en la carpeta `modulos`. Comprobar los ejemplos de e6_4 en el notebook, importando el módulo creado. Ver la documentación del módulo con `help`.
| tema_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nmdc
# language: python
# name: nmdc
# ---
# +
import os
from dotenv import load_dotenv
load_dotenv(os.path.expanduser("~/.nmdc_mongo.env"))
from nmdc_mongo import get_db, add_to_db
db_share = get_db("dwinston_share")
# -
biosamples = [
{
"name":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States - ",
"description":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States",
"lat_lon":{
"has_raw_value":"38.9206 -106.9489",
"latitude":38.9206,
"longitude":-106.9489
},
"geo_loc_name":"USA: Colorado",
"collection_date":"2017-05-09",
"env_broad_scale":{
"has_raw_value":"ENVO_00000446",
"type":"ControlledTermValue"
},
"env_local_scale":{
"has_raw_value":"ENVO_00000292",
"type":"ControlledTermValue"
},
"env_medium":{
"has_raw_value":"ENVO_00001998",
"type":"ControlledTermValue"
},
"ecosystem":"Environmental",
"ecosystem_category":"Terrestrial",
"ecosystem_type":"Soil",
"ecosystem_subtype":"Unclassified",
"specific_ecosystem":"Unclassified",
"depth":15,
"ncbi_taxonomy_name":"soil metagenome",
"community":"microbial communities",
"location":"The East River watershed near Crested Butte, Colorado, USA",
"habitat":"soil",
"sample_collection_site":"soil",
"add_date":"22-Jun-18 04.28.47.015000 PM",
"mod_date":"01-Oct-19 09.41.01.459000 AM",
"id":"igsn:IEWFS000I",
"identifier":"igsn:IEWFS000I"
},
{
"name":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States - ",
"description":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States",
"lat_lon":{
"has_raw_value":"38.9206 -106.9489",
"latitude":38.9206,
"longitude":-106.9489
},
"geo_loc_name":"USA: Colorado",
"collection_date":"2017-05-09",
"env_broad_scale":{
"has_raw_value":"ENVO_00000446",
"type":"ControlledTermValue"
},
"env_local_scale":{
"has_raw_value":"ENVO_00000292",
"type":"ControlledTermValue"
},
"env_medium":{
"has_raw_value":"ENVO_00001998",
"type":"ControlledTermValue"
},
"ecosystem":"Environmental",
"ecosystem_category":"Terrestrial",
"ecosystem_type":"Soil",
"ecosystem_subtype":"Unclassified",
"specific_ecosystem":"Unclassified",
"depth":15,
"ncbi_taxonomy_name":"soil metagenome",
"community":"microbial communities",
"location":"The East River watershed near Crested Butte, Colorado, USA",
"habitat":"soil",
"sample_collection_site":"soil",
"add_date":"22-Jun-18 04.28.47.015000 PM",
"mod_date":"01-Oct-19 09.41.01.459000 AM",
"id":"igsn:IEWFS000K",
"identifier":"igsn:IEWFS000K"
},
{
"name":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States - ",
"description":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States",
"lat_lon":{
"has_raw_value":"38.9206 -106.9489",
"latitude":38.9206,
"longitude":-106.9489
},
"geo_loc_name":"USA: Colorado",
"collection_date":"2017-05-09",
"env_broad_scale":{
"has_raw_value":"ENVO_00000446",
"type":"ControlledTermValue"
},
"env_local_scale":{
"has_raw_value":"ENVO_00000292",
"type":"ControlledTermValue"
},
"env_medium":{
"has_raw_value":"ENVO_00001998",
"type":"ControlledTermValue"
},
"ecosystem":"Environmental",
"ecosystem_category":"Terrestrial",
"ecosystem_type":"Soil",
"ecosystem_subtype":"Unclassified",
"specific_ecosystem":"Unclassified",
"depth":15,
"ncbi_taxonomy_name":"soil metagenome",
"community":"microbial communities",
"location":"The East River watershed near Crested Butte, Colorado, USA",
"habitat":"soil",
"sample_collection_site":"soil",
"add_date":"22-Jun-18 04.28.47.015000 PM",
"mod_date":"01-Oct-19 09.41.01.459000 AM",
"id":"igsn:IEWFS000B",
"identifier":"igsn:IEWFS000B"
},
{
"name":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States - ",
"description":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States",
"lat_lon":{
"has_raw_value":"38.9206 -106.9489",
"latitude":38.9206,
"longitude":-106.9489
},
"geo_loc_name":"USA: Colorado",
"collection_date":"2017-05-09",
"env_broad_scale":{
"has_raw_value":"ENVO_00000446",
"type":"ControlledTermValue"
},
"env_local_scale":{
"has_raw_value":"ENVO_00000292",
"type":"ControlledTermValue"
},
"env_medium":{
"has_raw_value":"ENVO_00001998",
"type":"ControlledTermValue"
},
"ecosystem":"Environmental",
"ecosystem_category":"Terrestrial",
"ecosystem_type":"Soil",
"ecosystem_subtype":"Unclassified",
"specific_ecosystem":"Unclassified",
"depth":15,
"ncbi_taxonomy_name":"soil metagenome",
"community":"microbial communities",
"location":"The East River watershed near Crested Butte, Colorado, USA",
"habitat":"soil",
"sample_collection_site":"soil",
"add_date":"22-Jun-18 04.28.47.015000 PM",
"mod_date":"01-Oct-19 09.41.01.459000 AM",
"id":"igsn:IEWFS000A",
"identifier":"igsn:IEWFS000A"
},
{
"name":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States - ",
"description":"Soil microbial communities from the East River watershed near Crested Butte, Colorado, United States",
"lat_lon":{
"has_raw_value":"38.9206 -106.9489",
"latitude":38.9206,
"longitude":-106.9489
},
"geo_loc_name":"USA: Colorado",
"collection_date":"2017-05-09",
"env_broad_scale":{
"has_raw_value":"ENVO_00000446",
"type":"ControlledTermValue"
},
"env_local_scale":{
"has_raw_value":"ENVO_00000292",
"type":"ControlledTermValue"
},
"env_medium":{
"has_raw_value":"ENVO_00001998",
"type":"ControlledTermValue"
},
"ecosystem":"Environmental",
"ecosystem_category":"Terrestrial",
"ecosystem_type":"Soil",
"ecosystem_subtype":"Unclassified",
"specific_ecosystem":"Unclassified",
"depth":15,
"ncbi_taxonomy_name":"soil metagenome",
"community":"microbial communities",
"location":"The East River watershed near Crested Butte, Colorado, USA",
"habitat":"soil",
"sample_collection_site":"soil",
"add_date":"22-Jun-18 04.28.47.015000 PM",
"mod_date":"01-Oct-19 09.41.01.459000 AM",
"id":"igsn:IEWFS000J",
"identifier":"igsn:IEWFS000J"
}
]
add_to_db(biosamples, db_share, collection_name="biosample_set")
assert db_share.biosample_set.count_documents({"id": {"$in": [d["id"] for d in biosamples]}}) == len(biosamples)
| metadata-translation/notebooks/ghissue_272.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("books_new_N2.csv",delimiter=',')
df[['Last name','First name']] = df['Author'].str.split(',',expand=True)
df.fillna("",inplace=True)
df['Full name of Author'] = df['First name'] + ' ' + df['Last name']
df = df.drop(['Author', 'First name', 'Last name'], 1)
second_column = df.pop('Full name of Author')
df.insert(1, 'Author', second_column)
df[['Book name','A/The']] = df['Title'].str.split(',',expand=True)
df.fillna("",inplace=True)
df['Full name of book'] = df['A/The'] + ' ' + df['Book name']
df = df.drop(['Book name', 'A/The','Title'], 1)
second_column = df.pop('Full name of book')
df.insert(0, 'Title', second_column)
df.to_csv('dulieuchuanhoa.csv')
df.head(50)
| non-function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 프로젝트 - Movielens 영화 추천 실습
# 이전 스텝에서 배운 MF 모델 학습 방법을 토대로, 내가 좋아할 만한 영화 추천 시스템을 제작해 보겠습니다.
#
# cloud jupyter 를 열고 aiffel/recommendata_iu/ 디렉토리에 python3로 ipynb파일을 생성 해 주세요 jupyter notebook 환경에서 프로젝트 코드를 작성하고 제출하시면 됩니다.
#
# 이번에 활용할 데이터셋은 추천시스템의 MNIST라고 부를만한 Movielens 데이터입니다.
#
# 유저가 영화에 대해 평점을 매긴 데이터가 데이터 크기 별로 있습니다. MovieLens 1M Dataset 사용을 권장합니다.
# 별점 데이터는 대표적인 explicit 데이터입니다. 하지만 implicit 데이터로 간주하고 테스트해볼 수 있습니다.
# 별점을 시청횟수로 해석해서 생각하겠습니다.
# 또한 유저가 3점 미만으로 준 데이터는 선호하지 않는다고 가정하고 제외하겠습니다.
# Cloud Storage에 미리 업로드 된 ml-1m폴더 내 파일을 심볼릭 링크로 개인 storage에 연결 해 줍니다.
#
# Cloud shell에서 아래 명령어를 입력해주세요.
# +
import os
import pandas as pd
rating_file_path=os.getenv('HOME') + '/aiffel/EXP_09_recommendata_iu/data/ml-1m/ratings.dat'
ratings_cols = ['user_id', 'movie_id', 'ratings', 'timestamp']
ratings = pd.read_csv(rating_file_path, sep='::', names=ratings_cols, engine='python', encoding = "ISO-8859-1")
orginal_data_size = len(ratings)
ratings.head()
# +
# 3점 이상만 남깁니다.
ratings = ratings[ratings['ratings']>=3]
filtered_data_size = len(ratings)
print(f'orginal_data_size: {orginal_data_size}, filtered_data_size: {filtered_data_size}')
print(f'Ratio of Remaining Data is {filtered_data_size / orginal_data_size:.2%}')
# -
# ratings 컬럼의 이름을 counts로 바꿉니다.
ratings.rename(columns={'ratings':'counts'}, inplace=True)
ratings['counts']
# 영화 제목을 보기 위해 메타 데이터를 읽어옵니다.
movie_file_path=os.getenv('HOME') + '/aiffel/EXP_09_recommendata_iu/data/ml-1m/movies.dat'
cols = ['movie_id', 'title', 'genre']
movies = pd.read_csv(movie_file_path, sep='::', names=cols, engine='python', encoding='ISO-8859-1')
movies.head()
# movie_id는 3952까지 존재
movies.tail()
# 여기까지가 전처리입니다. 이후에는 이전 스텝에 소개했던 것과 동일한 방식으로 MF model을 구성하여 내가 좋아할 만한 영화를 추천해 볼 수 있습니다.
#
#
# ### 2) 분석해 봅시다.
#
# - ratings에 있는 유니크한 영화 개수
# - ratings에 있는 유니크한 사용자 수
# - 가장 인기 있는 영화 30개(인기순)
#
# +
# 실습 위에 설명보고 이해해서 만들어보기
from scipy.sparse import csr_matrix
# ratings에 있는 유니크한 영화 개수
ratings['movie_id'].nunique()
# ratings에 있는 유니크한 사용자 수
ratings['user_id'].nunique()
# ratings 와 movies 데이터 프레임 합치기
# movie_id를 key로 ratings와 movies 데이터프레임을 조인
ratings = ratings.merge(movies, how='left', on='movie_id')
ratings.groupby('title')['user_id'].count().sort_values(ascending=False)[:30]
# -
# ratings의 데이터타입 체크
ratings.info()
# #### 결측치
# 결측값 존재 여부를 체크해보자
ratings.isnull().sum()
# #### 3.2 Unique Genre
# genre는 pipe-separated(|)로 구분 되어있으므로 unique한 genre가 무엇이 있는지 확인해보자
genres = list(movies['genre'].value_counts().keys())
genres[:20]
# genre는 pipe-separated(|)로 구분 되어있으므로 unique한 genre가 무엇이 있는지 확인해보자
genres = list(movies['genre'].value_counts().keys())
genres[:20]
# +
from collections import Counter
result = [] # 중복을 포함해서 genre 출현 빈도를 count
for genre in ratings['genre']:
result.extend(genre.split('|'))
# -
genre_counter = Counter(result)
genre_counter
sorted_genre_counter = genre_counter.most_common()
sorted_genre_counter
# #### 데이터 시각화 by using plotly
# +
# !pip install plotly
# Horizontal bar
import pandas as pd
import plotly.express as px
import plotly.io as pio
pio.renderers.default = 'notebook_connected'
sorted_genre_counter_df = pd.DataFrame(sorted_genre_counter, columns=['genre', 'count'])
fig = px.bar(sorted_genre_counter_df, x='count', y='genre', orientation='h')
fig.show()
# -
# Pie chart
fig = px.pie(sorted_genre_counter_df, values='count', names='genre', title='Pie chart for Genre')
fig.show()
# helper function
def get_unique_genre(genres):
result = [] # unique 장르를 담을 list
for genre in genres:
result.extend(genre.split('|'))
return sorted(set(result))
# unique한 genre는 다음과 같다.
unique_genres = get_unique_genre(genres)
unique_genres.sort
print(unique_genres)
# #### Timestamp
ratings.head()
# +
from datetime import datetime
import time
s = int('978300760')
datetime.fromtimestamp(s).strftime('%Y-%m-%d')
# timestamp를 %Y-%m-%d 형식으로 변경해봅니다.
# ratings.rename(columns={'timestamp' : 'date'}, inplace=True)
ratings['timestamp'] = ratings['timestamp'].apply(lambda s : datetime.fromtimestamp(int(s)).strftime('%Y-%m-%d'))
ratings.head()
# -
# #### year
# +
ratings['year'] = ratings['title'].str.extract('.*\((.*)\).*',expand = False).astype(int)
ratings.head()
# +
# 가장 인기있는 영화 top 10을 시각화 해봅니다.
import plotly.graph_objects as go
movies_top10 = ratings.groupby('title')['user_id'].count().sort_values(ascending=False)[:10]
fig = go.Figure(data=go.Scatter(
x=movies_top10.index,
y=movies_top10.values,
mode='markers',
marker=dict(size=[100,90,80,70,60,50,40,30,20,10],
color=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
))
fig.show()
# +
# 90년대 인기 영화 top 10을 시각화
cond1 = 'year >= 1990 and year <= 2000'
movies_90s = ratings.query(cond1)
movies_90s_top10 = movies_90s.groupby('title')['user_id'].count().sort_values(ascending=False)[:10]
fig1 = go.Figure(data=go.Scatter(
x=movies_90s_top10.index,
y=movies_90s_top10.values,
mode='markers',
marker=dict(size=[100,90,80,70,60,50,40,30,20,10],
color=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
))
fig1.show()
# +
# 80년대 인기 영화 top 10
cond2 = 'year >= 1980 and year < 1990'
movies_80s = ratings.query(cond2)
movies_80s_top10 = movies_80s.groupby('title')['user_id'].count().sort_values(ascending=False)[:10]
fig2 = go.Figure(data=go.Scatter(
x=movies_80s_top10.index,
y=movies_80s_top10.values,
mode='markers',
marker=dict(size=[100,90,80,70,60,50,40,30,20,10],
color=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
))
fig2.show()
# +
# 70년대 인기 영화 top 10
cond3 = 'year >= 1970 and year < 1980'
movies_70s = ratings.query(cond3)
movies_70s_top10 = movies_70s.groupby('title')['user_id'].count().sort_values(ascending=False)[:10]
fig3 = go.Figure(data=go.Scatter(
x=movies_70s_top10.index,
y=movies_70s_top10.values,
mode='markers',
marker=dict(size=[100,90,80,70,60,50,40,30,20,10],
color=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
))
fig3.show()
# -
# #### rating
# +
avg_ratings = ratings[['movie_id', 'counts', 'year']].groupby('movie_id').mean()
fig = px.scatter(avg_ratings, x="year", y="counts", color='counts')
fig.show()
# -
# ### 3) 내가 선호하는 영화를 5가지 골라서 ratings에 추가해 줍시다.
#
# user_id : 기존 id가 6040까지 있으므로 내 id는 6041으로 설정한다.
# movie_id : 내가 선택할 영화의 id가 리스트에 존재한다면 해당 id로 설정하고 그렇지 않다면 3952 이후로 설정
# count : 내가 좋아하는 영화이므로 5점을 부여
# title : title 형식에 맞게 '제목 (연도)'로 전처리
# genre : 장르가 여러개 존재할 경우 |로 연결
# +
# 2000년 이전 영화 중에서 내가 좋아하는 영화 5가지를 선택한다.
# 라이언 일병 구하기 Saving Private Ryan, 1998 War, Action, Drama
# 인생은 아름다워 Life Is Beautiful, 1997, Drama, Comedy
# 쇼생크 탈출 The Shaw<NAME>, 1994, Drama
# 타이타닉 Titanic, 1997, Romance, Drama
# 매트릭스 The Matrix, 1999 Sci-Fi, Action
# 영화 하나 하나 검색하며 title이 일치하는지 여부를 확인한다.
favorite_movies = ['Saving Private Ryan (1998)',
'Life Is Beautiful (1997)',
'Star Wars: Episode IV - A New Hope (1977)',
'Titanic (1997)',
'The Matrix (1999)']
# -
# 라이언 일병 구하기
movie_name = 'Saving Private Ryan (1998)'
ratings[ratings['title'].apply(lambda x : x == movie_name)]
# +
# 인생은 아름다워
# Life Is Beautiful (1997)으로는 인덱싱이 되지 않는다'
# 제목을 'Life Is Beautiful (La Vita è bella) (1997)'로 수정한다.
movie_name = 'Life Is Beautiful (1997)'
favorite_movies[1] = 'Life Is Beautiful (La Vita è bella) (1997)'
ratings[ratings['title'].apply(lambda x : x[:4] == 'Life')]
# +
# 스타워즈
movie_name = 'Star Wars: Episode IV - A New Hope (1977)'
movies[2] = 'Star Wars: Episode IV - A New Hope (1977)'
movies[movies['title'].apply(lambda x : (x[:7] == 'Star Wa'))]
# -
# 타이타닉
movie_name = 'Titanic (1997)'
ratings[ratings['title'].apply(lambda x : x == movie_name)]
# 매트릭스
# 매트릭스는 Matrix, The (1999)로 수정한다.
movie_name = 'The Matrix (1999)'
favorite_movies[4] = 'Matrix, The (1999)'
ratings[ratings['title'].apply(lambda x : x[:6] == 'Matrix')]
favorite_movies
my_movies = {
'user_id' : [6041] * 5,
'movie_id' : [2028, 2324, 318, 1721, 2571],
'count' : [5]*5,
'title' : favorite_movies,
'genre': ['Action|Drama|War',
'Comedy|Drama',
'Drama',
'Drama|Romance',
'Action|Sci-Fi|Thriller']
}
my_movies_df = pd.DataFrame(my_movies)
ratings = ratings.append(my_movies_df, ignore_index=True)
ratings.tail(10)
# 이제 필요한 칼럼만 남기고 제거한다.
ratings.drop(['timestamp', 'genre', 'year'], axis=1, inplace=True)
ratings
# +
# user_id와 title의 unique 값을 저장
user_unique = ratings['user_id'].unique()
movie_unique = ratings['title'].unique()
# unique 값의 개수만큼 user_id와 title을 indexing
user_to_idx = {v:k for k,v in enumerate(user_unique)}
movie_to_idx = {v:k for k,v in enumerate(movie_unique)}
# +
# 데이터 컬럼 내 값을 indexing된 값으로 교체
# user_to_idx.get을 통해 user_id 컬럼의 모든 값을 인덱싱한 Series를 구해 봅시다.
# 혹시 정상적으로 인덱싱되지 않은 row가 있다면 인덱스가 NaN이 될 테니 dropna()로 제거합니다.
temp_user_data = ratings['user_id'].map(user_to_idx.get).dropna()
if len(temp_user_data) == len(ratings): # 모든 row가 정상적으로 인덱싱되었다면
print('user_id column indexing OK!!')
ratings['user_id'] = temp_user_data # data['user_id']을 인덱싱된 Series로 교체해 줍니다.
else:
print('user_id column indexing Fail!!')
# movie_to_idx을 통해 title 컬럼도 동일한 방식으로 인덱싱해 줍니다.
temp_movie_data = ratings['title'].map(movie_to_idx.get).dropna()
if len(temp_movie_data) == len(ratings):
print('title column indexing OK!!')
ratings['title'] = temp_movie_data
else:
print('title column indexing Fail!!')
ratings
# +
from scipy.sparse import csr_matrix
num_user = ratings['user_id'].nunique()
num_movie = ratings['title'].nunique()
csr_data = csr_matrix((ratings['count'], (ratings['user_id'], ratings['title'])), shape=(num_user, num_movie))
csr_data
# -
# !pip install implicit
# +
from implicit.als import AlternatingLeastSquares
import os
import numpy as np
# implicit 라이브러리에서 권장하고 있는 부분
os.environ['OPENBLAS_NUM_THREADS']='1'
os.environ['KMP_DUPLICATE_LIB_OK']='True'
os.environ['MKL_NUM_THREADS']='1'
# -
# Implicit AlternatingLeastSquares 모델의 선언
als_model = AlternatingLeastSquares(factors=100, regularization=0.01, use_gpu=False, iterations=30, dtype=np.float32)
# als 모델은 input으로 item X user 꼴의 matrix를 받기 때문에 Transpose 해준다.
csr_data_transpose = csr_data.T
csr_data_transpose
# 모델 훈련
als_model.fit(csr_data_transpose)
# +
my_id = user_to_idx[6041] # 내가 새로 추가한 유저 아이디
saving_ryan = title_to_idx['Saving Private Ryan (1998)']
my_vector, saving_ryan_vector = als_model.user_factors[my_id], als_model.item_factors[saving_ryan]
# -
my_vector
matrix_vector
# 라이언 일병 구하기를 좋아한다!
favorite_movie = 'Saving Private Ryan (1998)'
movie_id = movie_to_idx[favorite_movie]
similar_movie = als_model.similar_items(movie_id, N=15)
similar_movie
# +
# 라이언 일병 구하기와 유사한 추천 영화 15개
# 쇼생크 탈출이 두개 보이고.. 쉰들러 리스트는 좋아하는 영화다!
idx_to_movie = {v:k for k,v in movie_to_idx.items()}
[idx_to_movie[i[0]] for i in similar_movie]
# -
# 비슷한 영화를 알려주는 함수
def get_similar_movie(movie_title):
movie_id = movie_to_idx[movie_title]
similar_movie = als_model.similar_items(movie_id)
similar_movie = [idx_to_movie[i[0]] for i in similar_movie]
return similar_movie
# 쉰들러리스트와 유사한 영화
get_similar_movie("Schindler's List (1993)")
# 레옹도 좋아한다. 이중에서 SEVEN도 좋아하는 영화이다.
get_similar_movie('Professional, The (a.k.a. Leon: The Professional) (1994)')
# +
# 나를 위한 영화 추천
for_me = user_to_idx[6041]
recommended_movie = als_model.recommend(for_me, csr_data, N=20, filter_already_liked_items=True)
recommended_movie
# -
# 인덱스를 영화 제목으로 변환
[idx_to_movie[i[0]] for i in recommended_movie]
# +
# 사용자가 기록을 남긴 데이터가 특정 영화 추천에 기여한 정도를 확인
recommended = movie_to_idx['Star Wars: Episode IV - A New Hope (1977)']
explain = als_model.explain(for_me, csr_data, itemid=recommended)
[(idx_to_movie[i[0]], i[1]) for i in explain[1]]
# -
# ### 4) CSR matrix를 직접 만들어 봅시다
# +
csr_data = csr_matrix((data.play, (data.user_id, data.artist)), shape= (num_user, num_artist))
csr_data
# -
# ### 5) als_model = AlternatingLeastSquares 모델을 직접 구성하여 훈련시켜 봅시다.
#
# ### 6) 내가 선호하는 5가지 영화 중 하나와 그 외의 영화 하나를 골라 훈련된 모델이 예측한 나의 선호도를 파악해 보세요.
# ### 7) 내가 좋아하는 영화와 비슷한 영화를 추천받아 봅시다.
#
# ### 8) 내가 가장 좋아할 만한 영화들을 추천받아 봅시다.
| EXP_09_movie_rec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import utils
import os
import numpy as np
from collections import OrderedDict
from vgg_face_dag import vgg_face_dag
from ami_model import AmIModel
device = torch.device('cuda')
# vgg_weight = './vgg_face_caffe.pth'
# vgg_weight = './keras_vgg_face.pth'
vgg_weight = './vgg_face_dag.pth'
# ami model
SKIP_LAYERS = utils.SKIP_LAYERS
attribute_model = AmIModel(vgg_face_dag(vgg_weight), 60., 2.15, 50.)
attribute_model.to(device)
attribute_model.eval()
attribute_model.register_my_hook(skip_layers=SKIP_LAYERS, ami_data=utils.load_neuron_set_lists())
# original model
vgg_net = vgg_face_dag(vgg_weight)
vgg_net.to(device)
vgg_net.eval()
# -
vgg_root = '../../data/vgg_face_caffe/'
vgg_names = utils.read_list(vgg_root + 'names.txt')
# +
img_dir = '/tmp/sized_full_images/'
img_list = utils.read_list('./benign_list.txt')
img_count = 0
adv_count = 0
with torch.no_grad():
for img_name in img_list:
img_name = img_name.strip()
img_path = os.path.join(img_dir, img_name)
prob_original = vgg_net(utils.get_data(img_path).to(device))
prob_attribute = attribute_model(utils.get_data(img_path).to(device))
id_original = torch.max(prob_original, 1)[1].item()
id_attribute = torch.max(prob_attribute, 1)[1].item()
id_gold = utils.get_identity(img_name, vgg_names)
if id_gold == id_original:
img_count += 1
### consistency observer for detecting adversarial samples ###
if id_original != id_attribute:
adv_count += 1
# print('{:3} Gold-Original-Attribute: {:4}-{:4}-{:4} | Adversary: {:.4f} [{:3}/{:3}]'
# .format(img_count, id_gold, id_original, id_attribute,
# 1.0*adv_count/img_count, adv_count, img_count))
print(f'False positive rate: {adv_count}/{img_count}={adv_count/img_count}')
| src/pytorch/benign_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT License.
# 
# # Using Azure Machine Learning Pipelines for batch prediction
#
# In this notebook we will demonstrate how to run a batch scoring job using Azure Machine Learning pipelines. Our example job will be to take an already-trained image classification model, and run that model on some unlabeled images. The image classification model that we'll use is the __[Inception-V3 model](https://arxiv.org/abs/1512.00567)__ and we'll run this model on unlabeled images from the __[ImageNet](http://image-net.org/)__ dataset.
#
# The outline of this notebook is as follows:
#
# - Register the pretrained inception model into the model registry.
# - Store the dataset images in a blob container.
# - Use the registered model to do batch scoring on the images in the data blob container.
# ## Prerequisites
# If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc.
from azureml.core import Experiment
from azureml.core.compute import AmlCompute, ComputeTarget
from azureml.core.datastore import Datastore
from azureml.core.runconfig import CondaDependencies, RunConfiguration
from azureml.data.data_reference import DataReference
from azureml.pipeline.core import Pipeline, PipelineData
from azureml.pipeline.steps import PythonScriptStep
# +
import os
from azureml.core import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# -
# ## Set up machine learning resources
# ### Set up datastores
# First, let’s access the datastore that has the model, labels, and images.
#
# ### Create a datastore that points to a blob container containing sample images
#
# We have created a public blob container `sampledata` on an account named `pipelinedata`, containing images from the ImageNet evaluation set. In the next step, we create a datastore with the name `images_datastore`, which points to this container. In the call to `register_azure_blob_container` below, setting the `overwrite` flag to `True` overwrites any datastore that was created previously with that name.
#
# This step can be changed to point to your blob container by providing your own `datastore_name`, `container_name`, and `account_name`.
# +
account_name = "pipelinedata"
datastore_name="images_datastore"
container_name="sampledata"
batchscore_blob = Datastore.register_azure_blob_container(ws,
datastore_name=datastore_name,
container_name= container_name,
account_name=account_name,
overwrite=True)
# -
# Next, let’s specify the default datastore for the outputs.
def_data_store = ws.get_default_datastore()
# ### Configure data references
# Now you need to add references to the data, as inputs to the appropriate pipeline steps in your pipeline. A data source in a pipeline is represented by a DataReference object. The DataReference object points to data that lives in, or is accessible from, a datastore. We need DataReference objects corresponding to the following: the directory containing the input images, the directory in which the pretrained model is stored, the directory containing the labels, and the output directory.
input_images = DataReference(datastore=batchscore_blob,
data_reference_name="input_images",
path_on_datastore="batchscoring/images",
mode="download"
)
model_dir = DataReference(datastore=batchscore_blob,
data_reference_name="input_model",
path_on_datastore="batchscoring/models",
mode="download"
)
label_dir = DataReference(datastore=batchscore_blob,
data_reference_name="input_labels",
path_on_datastore="batchscoring/labels",
mode="download"
)
output_dir = PipelineData(name="scores",
datastore=def_data_store,
output_path_on_compute="batchscoring/results")
# ### Create and attach Compute targets
# Use the below code to create and attach Compute targets.
# +
# choose a name for your cluster
aml_compute_name = os.environ.get("AML_COMPUTE_NAME", "gpu-cluster")
cluster_min_nodes = os.environ.get("AML_COMPUTE_MIN_NODES", 0)
cluster_max_nodes = os.environ.get("AML_COMPUTE_MAX_NODES", 1)
vm_size = os.environ.get("AML_COMPUTE_SKU", "STANDARD_NC6")
if aml_compute_name in ws.compute_targets:
compute_target = ws.compute_targets[aml_compute_name]
if compute_target and type(compute_target) is AmlCompute:
print('found compute target. just use it. ' + aml_compute_name)
else:
print('creating a new compute target...')
provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size, # NC6 is GPU-enabled
vm_priority = 'lowpriority', # optional
min_nodes = cluster_min_nodes,
max_nodes = cluster_max_nodes)
# create the cluster
compute_target = ComputeTarget.create(ws, aml_compute_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it will use the scale settings for the cluster
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# For a more detailed view of current Azure Machine Learning Compute status, use get_status()
print(compute_target.get_status().serialize())
# -
# ## Prepare the Model
# ### Download the Model
#
# Download and extract the model from http://download.tensorflow.org/models/inception_v3_2016_08_28.tar.gz to `"models"`
# create directory for model
model_dir = 'models'
if not os.path.isdir(model_dir):
os.mkdir(model_dir)
# +
import tarfile
import urllib.request
url="http://download.tensorflow.org/models/inception_v3_2016_08_28.tar.gz"
response = urllib.request.urlretrieve(url, "model.tar.gz")
tar = tarfile.open("model.tar.gz", "r:gz")
tar.extractall(model_dir)
# -
# ### Register the model with Workspace
# +
import shutil
from azureml.core.model import Model
# register downloaded model
model = Model.register(model_path = "models/inception_v3.ckpt",
model_name = "inception", # this is the name the model is registered as
tags = {'pretrained': "inception"},
description = "Imagenet trained tensorflow inception",
workspace = ws)
# remove the downloaded dir after registration if you wish
shutil.rmtree("models")
# -
# ## Write your scoring script
# To do the scoring, we use a batch scoring script `batch_scoring.py`, which is located in the same directory that this notebook is in. You can take a look at this script to see how you might modify it for your custom batch scoring task.
#
# The python script `batch_scoring.py` takes input images, applies the image classification model to these images, and outputs a classification result to a results file.
#
# The script `batch_scoring.py` takes the following parameters:
#
# - `--model_name`: the name of the model being used, which is expected to be in the `model_dir` directory
# - `--label_dir` : the directory holding the `labels.txt` file
# - `--dataset_path`: the directory containing the input images
# - `--output_dir` : the script will run the model on the data and output a `results-label.txt` to this directory
# - `--batch_size` : the batch size used in running the model.
#
# ## Build and run the batch scoring pipeline
# You have everything you need to build the pipeline. Let’s put all these together.
# ### Specify the environment to run the script
# Specify the conda dependencies for your script. You will need this object when you create the pipeline step later on.
# +
from azureml.core.runconfig import DEFAULT_GPU_IMAGE
cd = CondaDependencies.create(pip_packages=["tensorflow-gpu==1.13.1", "azureml-defaults"])
# Runconfig
amlcompute_run_config = RunConfiguration(conda_dependencies=cd)
amlcompute_run_config.environment.docker.enabled = True
amlcompute_run_config.environment.docker.gpu_support = True
amlcompute_run_config.environment.docker.base_image = DEFAULT_GPU_IMAGE
amlcompute_run_config.environment.spark.precache_packages = False
# -
# ### Specify the parameters for your pipeline
# A subset of the parameters to the python script can be given as input when we re-run a `PublishedPipeline`. In the current example, we define `batch_size` taken by the script as such parameter.
from azureml.pipeline.core.graph import PipelineParameter
batch_size_param = PipelineParameter(name="param_batch_size", default_value=20)
# ### Create the pipeline step
# Create the pipeline step using the script, environment configuration, and parameters. Specify the compute target you already attached to your workspace as the target of execution of the script. We will use PythonScriptStep to create the pipeline step.
# +
inception_model_name = "inception_v3.ckpt"
batch_score_step = PythonScriptStep(
name="batch_scoring",
script_name="batch_scoring.py",
arguments=["--dataset_path", input_images,
"--model_name", "inception",
"--label_dir", label_dir,
"--output_dir", output_dir,
"--batch_size", batch_size_param],
compute_target=compute_target,
inputs=[input_images, label_dir],
outputs=[output_dir],
runconfig=amlcompute_run_config
)
# -
# ### Run the pipeline
# At this point you can run the pipeline and examine the output it produced.
# + tags=["pipelineparameterssample"]
pipeline = Pipeline(workspace=ws, steps=[batch_score_step])
pipeline_run = Experiment(ws, 'batch_scoring').submit(pipeline, pipeline_parameters={"param_batch_size": 20})
# -
# ### Monitor the run
from azureml.widgets import RunDetails
RunDetails(pipeline_run).show()
pipeline_run.wait_for_completion(show_output=True)
# ### Download and review output
step_run = list(pipeline_run.get_children())[0]
step_run.download_file("./outputs/result-labels.txt")
import pandas as pd
df = pd.read_csv("result-labels.txt", delimiter=":", header=None)
df.columns = ["Filename", "Prediction"]
df.head()
# ## Publish a pipeline and rerun using a REST call
# ### Create a published pipeline
# Once you are satisfied with the outcome of the run, you can publish the pipeline to run it with different input values later. When you publish a pipeline, you will get a REST endpoint that accepts invoking of the pipeline with the set of parameters you have already incorporated above using PipelineParameter.
# +
published_pipeline = pipeline_run.publish_pipeline(
name="Inception_v3_scoring", description="Batch scoring using Inception v3 model", version="1.0")
published_pipeline
# -
# ### Get published pipeline
#
# You can get the published pipeline using **pipeline id**.
#
# To get all the published pipelines for a given workspace(ws):
# ```css
# all_pub_pipelines = PublishedPipeline.get_all(ws)
# ```
# +
from azureml.pipeline.core import PublishedPipeline
pipeline_id = published_pipeline.id # use your published pipeline id
published_pipeline = PublishedPipeline.get(ws, pipeline_id)
published_pipeline
# -
# ## Rerun the pipeline using the REST endpoint
# ### Get AAD token
# [This notebook](https://aka.ms/pl-restep-auth) shows how to authenticate to AML workspace.
# +
from azureml.core.authentication import InteractiveLoginAuthentication
import requests
auth = InteractiveLoginAuthentication()
aad_token = auth.get_authentication_header()
# -
# ### Run published pipeline
rest_endpoint = published_pipeline.endpoint
# specify batch size when running the pipeline
response = requests.post(rest_endpoint,
headers=aad_token,
json={"ExperimentName": "batch_scoring",
"ParameterAssignments": {"param_batch_size": 50}})
run_id = response.json()["Id"]
# ### Monitor the new run
# +
from azureml.pipeline.core.run import PipelineRun
published_pipeline_run = PipelineRun(ws.experiments["batch_scoring"], run_id)
RunDetails(published_pipeline_run).show()
| how-to-use-azureml/machine-learning-pipelines/pipeline-batch-scoring/pipeline-batch-scoring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from skimage.io import imread, imsave, imshow
from skimage.data import cells3d
import napari_mahotas_image_processing as nmh
import numpy as np
blobs = imread('https://samples.fiji.sc/blobs.png')
imshow(blobs)
# ## Gaussian blur
blurred = nmh.gaussian_blur(blobs, sigma=5)
imshow(blurred)
# ## Thresholding
binary = nmh.threshold_otsu(blobs)
imshow(binary * 1)
# ## Labeling
labeled = nmh.connected_component_labeling(binary)
imshow(labeled)
# ## Sobel operator
edges = nmh.sobel_edge_detector(blobs)
imshow(edges)
# ## Fill holes
# +
cells = cells3d()
nuclei = cells[30, 1]
imshow(nuclei)
# -
binary_cells = nmh.threshold_otsu(nmh.gaussian_blur(nuclei))
imshow(binary_cells * 1)
filled = nmh.binary_fill_holes(binary_cells)
imshow(filled * 1)
# ## Seeded watershed
seeds = nmh.connected_component_labeling(filled)
imshow(seeds)
membranes = cells[30, 0]
imshow(membranes)
labels = nmh.seeded_watershed(membranes, seeds)
imshow(labels)
# ## Distance Map
distance_map = nmh.euclidean_distance_map(binary)
imshow(distance_map)
# ## Split touching objects
split = nmh.split_touching_objects(binary)
imshow(split * 1)
# ## Voronoi-Otsu-Labeling
# image = blobs
# spot_sigma = 2
# outline_sigma = 2
#
# blurred_spots = mh.gaussian_filter(image, spot_sigma)
# spot_centroids = mh.locmax(blurred_spots)
#
# blurred_outline = mh.gaussian_filter(image, outline_sigma)
# binary_otsu = threshold_otsu(blurred_outline)
#
# remaining_spots = spot_centroids * binary_otsu
#
# labeled_spots, num_labes = mh.label(remaining_spots)
# labels = mh.cwatershed(binary_otsu, labeled_spots) # * binary_otsu
#
#
# imshow(labels)
#
# ### ... doesn't work :-( yet :-)
| docs/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://raw.githubusercontent.com/Qiskit/qiskit-tutorials/master/images/qiskit-heading.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# ## Iterative Phase Estimation Algorithm
#
#
# The latest version of this notebook is available on https://github.com/qiskit/qiskit-tutorial.
#
# For more information about how to use the IBM Q Experience (QX), consult the [tutorials](https://quantumexperience.ng.bluemix.net/qstage/#/tutorial?sectionId=c59b3710b928891a1420190148a72cce&pageIndex=0), or check out the [community](https://quantumexperience.ng.bluemix.net/qstage/#/community).
#
# ***
# ### Contributors
# <NAME>, <NAME>, <NAME>
#
# ### Qiskit Package Versions
import qiskit
qiskit.__qiskit_version__
# ## Quantum Phase Estimation (QPE)
#
# The Quantum Phase Estimation (QPE) algorithm solves the problem of finding unknown eigenvalues of a unitary operator. The attractiveness of the QPE algorithm is due to the fact that it is a key ingredient of some other very powerful algorithms, like order-finding and Shor's.
#
# In a standard textbook, such as Nielsen & Chuang <i>Quantum Computation and Quantum Information</i>, in the QPE, each bit of the phase is encoded in a different qubit on a register using the phase kickback property of controlled-unitary operations. This is followed by an inverse Quantum Fourier Transform operation, which yields an n-bit approximation to the phase by reading the n-qubit register.
# ## Iterative Phase Estimation Algorithm (IPEA)
# The QPE algorithm can, however, be realized in a much smaller qubit system, by iterating the steps on a system of just two qubits. This is called the Iterative Phase Estimation Algorithm (IPEA).
#
# Consider the problem of finding $\varphi$ given $|\Psi\rangle$ and $U$ in $U |\Psi\rangle = e^{i \phi} | \Psi \rangle$, with $\phi = 2 \pi \varphi$. Let's assume for now that $\varphi$ can be written as $\varphi = \varphi_1/2 + \varphi_2/4 + ... + \varphi_m/2^m = 0.\varphi_1 \varphi_2 ... \varphi_m$, where we have defined the notation $0.\varphi_1 \varphi_2 ... \varphi_m$. Now, if we have two qubits, $q_0$ and $q_1$, and we initialize them as $q_0 \rightarrow |+\rangle$ and $q_1 \rightarrow |\Psi \rangle$, then, after applying a control-U between $q_0$ and $q_1$ $2^t$ times, the state of $q_0$ can be written as $|0\rangle + e^{i 2 \pi 2^{t} \varphi} | 1 \rangle$. That is, the phase of $U$ has been kicked back into $q_0$ as many times as the control operation has been performed.
#
# For $t=0$, we have a total phase in $q_0$ of $e^{i 2 \pi 2^{0} \varphi} = e^{i 2 \pi \varphi} = e^{i 2 \pi 0.\varphi_1 \varphi_2 ... \varphi_m}$
#
# For $t=1$, the phase would be $e^{i 2 \pi 2^{1} \varphi} = e^{i 2 \pi \varphi_1} e^{i 2 \pi 0.\varphi_2 \varphi_3 ... \varphi_m}$
#
# For $t=2$, $e^{i 2 \pi 2^{2} \varphi} = e^{i 2 \pi 2 \varphi_1} e^{i 2 \pi \varphi_2} e^{i 2 \pi 0.\varphi_3 \varphi_4 ... \varphi_m}$
#
# And for $t=m-1$, $e^{i 2 \pi 2^{m-1} \varphi} = e^{i 2 \pi 2^{m-2} \varphi_1} e^{i 2 \pi 2^{m-3} \varphi_2} ... e^{i 2 \pi 2^{-1} \varphi_m} = e^{i 2 \pi 0.\varphi_m}$. Note that if we perform a Hadamard operation on the state $|0\rangle + e^{i 2 \pi 0.\varphi_m}|1\rangle$ and perform a measurement in the standard basis, we obtain $|0\rangle$ if $\varphi_m = 0$ and $|1\rangle$ if $\varphi_m = 1$.
#
# In the first step of the IPEA, we directly measure the least significant bit of the phase $\varphi$, $\varphi_m$, by initializing the 2-qubit register as described above, performing $2^{m-1}$ control-$U$ operations between the qubits, and measuring $q_0$ in the diagonal basis.
#
# For the second step, we initialize the register in the same way and apply $2^{m-2}$ control-$U$ operations. The phase in $q_0$ after these operations is now $e^{i 2 \pi 0.\varphi_{m-1}\varphi_{m}}= e^{i 2 \pi 0.\varphi_{m-1}} e^{i 2 \pi \varphi_m/4}$. We see that prior to extracting the phase bit $\varphi_{m-1}$, we must perform a phase correction of $\varphi_m /2$. This is equivalent to a rotation around the $Z-$axis of angle $-\varphi_m /4$.
#
# Therefore, the $k$th step of the IPEA, giving $\varphi_{m-k+1}$, consists of the register initialization ($q_0$ in $|+\rangle$, $q_1$ in $|\Psi\rangle$), the application of control-$U$ $2^{m-k}$ times, a rotation around $Z$ of angle $\omega_k = -2 \pi 0.0\varphi_{k+1} ... \varphi_m$, a Hadamard transform to $q_0$, and a measurement of $q_0$ in the standard basis. Note that $q_1$ remains in the state $|\Psi\rangle$ throughout the algorithm.
# ## IPEA circuit
#
# Let's first initialize the API and import the necessary packages
# +
from math import pi
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
# %matplotlib inline
# importing Qiskit
from qiskit import BasicAer, IBMQ
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister
from qiskit import execute
from qiskit.tools.visualization import plot_histogram
from qiskit.tools.monitor import job_monitor
# -
# Load saved IBMQ accounts
IBMQ.load_account()
# Now you can try the following circuit in the quantum simulator for a phase of $-5\pi/8 = 2 \pi \varphi$ and $m=4$. Note that the IPEA cannot be run in the real device in this form, due to the current lack of feedback capability.
# +
# We first define controlled gates used in the IPEA
def cu1fixed(qProg, c, t, a):
qProg.u1(-a, t)
qProg.cx(c, t)
qProg.u1(a, t)
qProg.cx(c, t)
def cu5pi8(qProg, c, t):
cu1fixed(qProg, c, t, -5.0*pi/8.0)
# We then prepare quantum and classical registers and the circuit
qr = QuantumRegister(2)
cr = ClassicalRegister(4)
circuitName="IPEAonSimulator"
ipeaCircuit = QuantumCircuit(qr, cr)
# Apply IPEA
ipeaCircuit.h(qr[0])
for i in range(8):
cu5pi8(ipeaCircuit, qr[0], qr[1])
ipeaCircuit.h(qr[0])
ipeaCircuit.measure(qr[0], cr[0])
ipeaCircuit.reset(qr[0])
ipeaCircuit.h(qr[0])
for i in range(4):
cu5pi8(ipeaCircuit, qr[0], qr[1])
ipeaCircuit.u1(-pi/2, qr[0]).c_if(cr, 1)
ipeaCircuit.h(qr[0])
ipeaCircuit.measure(qr[0], cr[1])
ipeaCircuit.reset(qr[0])
ipeaCircuit.h(qr[0])
for i in range(2):
cu5pi8(ipeaCircuit, qr[0], qr[1])
ipeaCircuit.u1(-pi/4, qr[0]).c_if(cr, 1)
ipeaCircuit.u1(-pi/2, qr[0]).c_if(cr, 2)
ipeaCircuit.u1(-3*pi/4, qr[0]).c_if(cr, 3)
ipeaCircuit.h(qr[0])
ipeaCircuit.measure(qr[0], cr[2])
ipeaCircuit.reset(qr[0])
ipeaCircuit.h(qr[0])
cu5pi8(ipeaCircuit, qr[0], qr[1])
ipeaCircuit.u1(-pi/8, qr[0]).c_if(cr, 1)
ipeaCircuit.u1(-2*pi/8, qr[0]).c_if(cr, 2)
ipeaCircuit.u1(-3*pi/8, qr[0]).c_if(cr, 3)
ipeaCircuit.u1(-4*pi/8, qr[0]).c_if(cr, 4)
ipeaCircuit.u1(-5*pi/8, qr[0]).c_if(cr, 5)
ipeaCircuit.u1(-6*pi/8, qr[0]).c_if(cr, 6)
ipeaCircuit.u1(-7*pi/8, qr[0]).c_if(cr, 7)
ipeaCircuit.h(qr[0])
ipeaCircuit.measure(qr[0], cr[3])
backend = BasicAer.get_backend('qasm_simulator')
shots = 1000
results = execute(ipeaCircuit, backend=backend, shots=shots).result()
plot_histogram(results.get_counts())
# -
# The results are given in terms of $\varphi = 0.\varphi_1 \varphi_2 \varphi_3 \varphi_4$, with the least significant digit ($\varphi_4$) as the leftmost bit in the classical register. The result is $\varphi = 11/16$, from which $\phi = 2\pi \varphi = 11 \pi/8 = 2 \pi - 5\pi/8$, as encoded in the circuit.
# ## IPEA in the real device
#
# As we have mentioned before, we currently lack the ability to use measurement feedback or feedforward, along with qubit resetting, on the real device in the Quantum Experience. However, we still can implement a segmentized version of the IPEA by extracting the information about the phase one bit at a time.
#
# Try the following four circuits in the real device. They estimate the same phase as in the previous example (-5$\pi/8$), one bit at a time, from least ($\varphi_4$) to most ($\varphi_1$) significant bit.
# +
# We then prepare quantum and classical registers and the circuit
qr = QuantumRegister(5)
cr = ClassicalRegister(5)
realStep1Circuit = QuantumCircuit(qr, cr)
# Apply IPEA
realStep1Circuit.h(qr[0])
for i in range(8):
cu5pi8(realStep1Circuit, qr[0], qr[1])
realStep1Circuit.h(qr[0])
realStep1Circuit.measure(qr[0], cr[0])
#connect to remote API to be able to use remote simulators and real devices
print("Available backends:", [BasicAer.backends(), IBMQ.backends()])
backend = IBMQ.get_backend("ibmq_5_yorktown")
shots = 1000
job_exp1 = execute(realStep1Circuit, backend=backend, shots=shots)
job_monitor(job_exp1)
# -
results1 = job_exp1.result()
plot_histogram(results1.get_counts())
# In the first step of IPEA as above, we obtain the bit "1" with probability close to one. We then proceed to the second step of IPEA, assuming that we have identified the result of the first step correctly, as below.
# +
realStep2Circuit = QuantumCircuit(qr, cr)
# Apply IPEA
realStep2Circuit.h(qr[0])
for i in range(4):
cu5pi8(realStep2Circuit, qr[0], qr[1])
realStep2Circuit.u1(-pi/2, qr[0]) # Assuming the value of the measurement on Step 1
realStep2Circuit.h(qr[0])
realStep2Circuit.measure(qr[0], cr[0])
job_exp2 = execute(realStep2Circuit, backend=backend, shots=shots)
job_monitor(job_exp1)
# -
results2 = job_exp2.result()
plot_histogram(results2.get_counts())
# In the second step of IPEA as above, we obtain the bit "1" with probability close to one. We then proceed to the third step of IPEA, assuming that we have identified the result of the first and second steps correctly, as below.
# +
realStep3Circuit = QuantumCircuit(qr, cr)
# Apply IPEA
realStep3Circuit.h(qr[0])
for i in range(2):
cu5pi8(realStep3Circuit, qr[0], qr[1])
realStep3Circuit.u1(-3*pi/4, qr[0]) # Assuming the value of the measurement on Step 1 and Step 2
realStep3Circuit.h(qr[0])
realStep3Circuit.measure(qr[0], cr[0])
job_exp3 = execute(realStep3Circuit, backend=backend, shots=shots)
job_monitor(job_exp3)
# -
results3 = job_exp3.result()
plot_histogram(results3.get_counts())
# In the third step of IPEA as above, we obtain the bit "0" with probability close to one. We then proceed to the fourth step of IPEA, assuming that we have identified the result of the first, second, and third steps correctly, as below.
# +
realStep4Circuit = QuantumCircuit(qr, cr)
# Apply IPEA
realStep4Circuit.h(qr[0])
cu5pi8(realStep4Circuit, qr[0], qr[1])
realStep4Circuit.u1(-3*pi/8, qr[0]) # Assuming the value of the measurement on Step 1, 2, and 3
realStep4Circuit.h(qr[0])
realStep4Circuit.measure(qr[0], cr[0])
job_exp4 = execute(realStep4Circuit, backend=backend, shots=shots)
job_monitor(job_exp4)
# -
results4 = job_exp4.result()
plot_histogram(results4.get_counts())
# In the fourth step of the IPEA, we identify the bit "1" with high probability. In summary, we can conclude with high probability that the binary string of the phase is "1011"; that is, eleven in the decimal.
#
# We have left aside the case when $\varphi$ does not accept a decomposition of the form $\varphi = \varphi_1/2 + \varphi_2/4 + ... + \varphi_m/2^m$. In that case, it can be shown that we can still use the IPEA to obtain $\varphi$ to an accuracy of $2^{-m}$ with greater than a constant probability independent of $m$ (around $81\%$ [1]).
# ### References
#
# [1] <NAME> *et al. Phys. Rev. A* **76**, 030306 (2007)
| algorithms/iterative_phase_estimation_algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings('ignore')
import scipy.io.wavfile as file
import IPython.display as ipd
from utils import *
from ola import OLA
import numpy as np
import librosa
import matplotlib.pyplot as plt
# -
out_file_path = '_output/test.wav'
in_file_path = '_input/test.wav'
base_file_name = os.path.basename(in_file_path).split('.')[0]
sr, w = read_audio('_input/', 'test.wav')
# +
tester = OLA()
# using some response
response = tester.equalizer(db1=10, db2 = 10, db12 = 20)
# response = 1- tester.lowpass_resp(tester.coef_no)
# computing frequencies corresponding to coefficient
freqs = [i * tester.sample_rate / (2 * (tester.coef_no - 1)) for i in range(tester.coef_no)]
# printing response function
plt.plot(freqs, response)
plt.show()
# -
w_out, proc_time = tester.test(w, response)
w_out = w_out.astype('int16')
# w_out = w_out / 2**15
file.write(out_file_path, sr, w_out)
print('file {} is processed, time: {}'.format(base_file_name, proc_time))
ipd.Audio(w, rate=sr)
plot_waveform(w, 0, len(w))
ipd.Audio(w_out, rate=sr)
plot_waveform(w_out, 0, len(w_out))
| Homeworks/11_Equalizer/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Daily Coding Problem #1 [Easy]
# Good morning! Here's your coding interview problem for today.
#
# This problem was recently asked by Google.
#
# Given a list of numbers and a number k, return whether any two numbers from the list add up to k.
#
# For example, given `[10, 15, 3, 7]` and `k` of `17`, return true since `10 + 7` is `17`.
#
# Bonus: Can you do this in one pass?
# +
def problem1(arr, target):
temp = set(arr)
for x in temp:
if target-x in temp:
return x, target-x
arr = [10, 15, 3, 7]
problem1(arr, 17)
# -
# This runs in O(N) since we have to iterate over the whole list/array. The lookup of target-x does only cost O(1) because a set is implemented with a hash table. Hash tables do have an average lookup time of O(1).
| problem_#1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook compares the size of serialized sklearn `NearestNeighbors` models depending of the neighboring algorithm.
# +
import itertools
import numpy as np
import os
import pandas as pd
import pickle
from sklearn.neighbors import NearestNeighbors
# +
# %%time
n_samples_range = np.logspace(3, 5, num=3, base=10, dtype=int)
n_features_range = np.logspace(1, 3, num=3, base=10, dtype=int)
algorithms = ['brute', 'kd_tree', 'ball_tree']
temp_path = 'temp_model.pickle'
results = []
for n_samples, n_features in itertools.product(n_samples_range, n_features_range):
experiment_results = {'n_samples': n_samples, 'n_features': n_features}
X = np.random.randn(n_samples, n_features)
for algo in algorithms:
model = NearestNeighbors(algorithm=algo)
model.fit(X)
pickle.dump(model, open(temp_path, 'wb'))
pickle_size = os.path.getsize(temp_path)
experiment_results[f'{algo}'] = round(pickle_size / 1e6, 2)
results.append(experiment_results)
os.remove(temp_path)
# -
df = pd.DataFrame(results)
print(df.to_markdown())
| sklearn_nn_pickled_size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
skempi_data = pd.read_csv("../data/skempi_ABlike_singleMut.Final.csv",index_col=0)
abbind_data = pd.read_csv('../data/ab_bind_filtered_2020.csv',index_col=0)
skempi_data.head()
abbind_data.head()
set(abbind_data['#PDB']).intersection(set(skempi_data['#PDB']))
abbind_data.loc[abbind_data['#PDB']=='2JEL']
skempi_data.loc[skempi_data['#PDB'] == ('1DQJ')]
# +
1BJ1 mutantes en distinta cadena, mantiene mismo nombre. 1CZ8 mutante en distinta cadena, mantiene mismo nombre.
1DQJ cadena AB == LH, 1MHP partners esta dada vuelta, 1MLC cadena AB == LH, 1N8Z cadena AB == LH,1VFB cadena AB==LH
1YY9 cadena CD == LH, 2NYY cadena DC == HL, 2NZ9 DC==HL, 3BN9 CD == LH,
# -
AB_LH_chain = ['1DQJ','1MLC','1N8Z','1VFB']
CD_LH_chain = ['1YY9','2NYY','2NZ9','3BN9']
for pdb in AB_LH_chain:
chain_map = {'L':'A','H':'B'}
abbind_data.loc[abbind_data['#PDB']==pdb,'chain.2'] = abbind_data.loc[abbind_data['#PDB']==pdb]['chain'].map(chain_map)
abbind_data.loc[:,'chain.2'].fillna(abbind_data['chain'],inplace=True)
for pdb in CD_LH_chain:
chain_map = {'L':'C','H':'D'}
abbind_data.loc[abbind_data['#PDB']==pdb,'chain.2'] = abbind_data.loc[abbind_data['#PDB']==pdb]['chain'].map(chain_map)
abbind_data.loc[:,'chain.2'].fillna(abbind_data['chain'],inplace=True)
abbind_data.iloc[65:,:]
abbind_data['pdb_mutation_chain'] = abbind_data[['pdb_mutation','chain.2']].sum(axis=1)
skempi_data['pdb_mutation_chain'] = skempi_data[['pdb_mutation','chain']].sum(axis=1)
mutants_union = set(skempi_data['pdb_mutation_chain']).union(set(abbind_data['pdb_mutation_chain']))
len(mutants_union)
len(set(abbind_data['pdb_mutation_chain']))
from matplotlib_venn import venn2, venn2_circles, venn2_unweighted
from matplotlib_venn import venn3, venn3_circles
import matplotlib.pyplot as plt
# +
plt.subplots(figsize=(7, 5))
v = venn3([mutants_union,set(skempi_data['pdb_mutation_chain']),set(abbind_data['pdb_mutation_chain'])],
set_labels=["ABPRED","SKEMPI2","AB-BIND"],set_colors=('r', 'g','b'))
v.get_label_by_id('10').set_text("")
# c = venn3_circles([mutants_union,set(skempi_data['pdb_mutation_chain']),set(abbind_data['pdb_mutation_chain'])],
# linestyle='dashed')
plt.savefig("Abpred_database2020.png",dpi=300,bbox_inches="tight")
# -
len(mutants_union)
# +
total_mutants_series = pd.Series(list(mutants_union))
skempi_mutants_toselect = total_mutants_series[total_mutants_series.isin(skempi_data['pdb_mutation_chain'])]
abbind_mutants_toselect = total_mutants_series[-total_mutants_series.isin(skempi_data['pdb_mutation_chain'])]
len(skempi_mutants_toselect),len(abbind_mutants_toselect)
# -
# data selected for each database,
abbind_data_selected = abbind_data[['#PDB','mutant','chain','partners','ddG']].loc[abbind_data['pdb_mutation_chain'].isin(abbind_mutants_toselect)]
abbind_data_selected['database'] = 'ab-bind'
abbind_data_selected
# +
skempi_data_selected = skempi_data[['#PDB','Mutation(s)_cleaned','chain','partners','ddG']].loc[skempi_data['pdb_mutation_chain'].isin(skempi_mutants_toselect)]
skempi_data_selected['database'] = 'skempi'
skempi_data_selected.rename(columns={'Mutation(s)_cleaned':'mutant'},inplace=True)
skempi_data_selected['mutant'] = skempi_data_selected['mutant'].str[0]+skempi_data_selected['mutant'].str[2:]
# -
with open('../data/mutant_list_ab-bind.txt','w') as f:
pdbs = abbind_data_selected['#PDB']
mutants = abbind_data_selected['mutant']
chains = abbind_data_selected['chain']
partners = abbind_data_selected['partners']
raw_text = (pdbs+'\t'+mutants+'\t'+chains+'\t'+partners).str.cat(sep='\n')
f.write(raw_text)
with open('../data/mutant_list_skempi.txt','w') as f:
pdbs = skempi_data_selected['#PDB']
mutants = skempi_data_selected['mutant']
chains = skempi_data_selected['chain']
partners = skempi_data_selected['partners']
raw_text = (pdbs+'\t'+mutants+'\t'+chains+'\t'+partners).str.cat(sep='\n')
f.write(raw_text)
final_database = pd.concat([skempi_data_selected,abbind_data_selected])
final_database.drop_duplicates(subset=["#PDB","mutant","chain"],inplace=True)
final_database.head()
print(final_database[["#PDB"]].drop_duplicates().to_string(index=False))
# +
# final_database.to_string?
# +
#final_database.to_csv('../data/Data_skempi_ab-bind_Final-merge.csv')
# -
for n,data in final_database.groupby("database"):
with open('mutant_list_{}.txt'.format(n),'w') as f:
pdbs = data['#PDB']
mutants = data['mutant']
chains = data['chain']
partners = data['partners']
raw_text = (pdbs+'\t'+mutants+'\t'+chains+'\t'+partners).str.cat(sep='\n')
f.write(raw_text)
| notebooks/abpred_ab-bind_skempi_database.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CoxPH tests
# nuclio: ignore
import nuclio
import warnings
warnings.simplefilter(action="ignore", category=FutureWarning)
# +
import os
import pandas as pd
from mlrun.datastore import DataItem
from mlrun.artifacts import get_model
from cloudpickle import load
from mlrun.mlutils.models import eval_class_model
def cox_test(
context,
models_path: DataItem,
test_set: DataItem,
label_column: str,
plots_dest: str = "plots",
model_evaluator = None
) -> None:
"""Test one or more classifier models against held-out dataset
Using held-out test features, evaluates the peformance of the estimated model
Can be part of a kubeflow pipeline as a test step that is run post EDA and
training/validation cycles
:param context: the function context
:param model_file: model artifact to be tested
:param test_set: test features and labels
:param label_column: column name for ground truth labels
:param score_method: for multiclass classification
:param plots_dest: dir for test plots
:param model_evaluator: WIP: specific method to generate eval, passed in as string
or available in this folder
"""
xtest = test_set.as_df()
ytest = xtest.pop(label_column)
model_file, model_obj, _ = get_model(models_path.url, suffix='.pkl')
model_obj = load(open(str(model_file), "rb"))
try:
# there could be different eval_models, type of model (xgboost, tfv1, tfv2...)
if not model_evaluator:
# binary and multiclass
eval_metrics = eval_class_model(context, xtest, ytest, model_obj)
# just do this inside log_model?
model_plots = eval_metrics.pop("plots")
model_tables = eval_metrics.pop("tables")
for plot in model_plots:
context.log_artifact(plot, local_path=f"{plots_dest}/{plot.key}.html")
for tbl in model_tables:
context.log_artifact(tbl, local_path=f"{plots_dest}/{plot.key}.csv")
context.log_results(eval_metrics)
except:
#dummy log:
context.log_dataset("cox-test-summary", df=model_obj.summary, index=True, format="csv")
context.logger.info("cox tester not implemented")
# +
# nuclio: end-code
# -
task_params = {
"name" : "tasks cox test",
"params": {
"label_column" : "labels",
"plots_dest" : "churn/test/plots"}}
DATA_URL = "https://raw.githubusercontent.com/yjb-ds/testdata/master/demos/churn/churn-tests.csv"
# ## Run Locally
# +
from mlrun import run_local, NewTask, mlconf
run = run_local(NewTask(**task_params),
handler=cox_test,
inputs={"test_set": DATA_URL,
"models_path" : "models/cox"},
workdir=mlconf.artifact_path+"/churn")
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Run Remotely
# + pycharm={"name": "#%%\n"}
from mlrun import import_function
from mlrun.platforms.other import auto_mount
GPU = False
fn = import_function("hub://coxph_test")
if GPU:
fn.image = "mlrun/ml-models-gpu"
fn.apply(auto_mount())
run = fn.run(
NewTask(**task_params),
inputs={
"test_set" : DATA_URL,
"models_path" : "models/cox"},
workdir=os.path.join(mlconf.artifact_path, "churn"))
# -
| coxph_test/coxph_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: basement
# language: python
# name: basement
# ---
# # Figure S6: Distribution of misfits
# This notebook produces figure S6 of Tankersley et al. 2022. The figure shows the distribution of misfits between OIB, ANTOSTRAT, and ROSETTA-Ice basement models.
# import necessary python packages
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from matplotlib import gridspec
import pygmt
import os
import geopandas as gpd
# all file paths assume this notebook is in /RIS_basement_sediment/figures
pwd = os.getcwd()
pwd
# ## Load data
# +
# OIB to ANTOSTRAT comparison
oib_to_anto = pd.read_csv("../data/results/oib_basement.csv", header=0, index_col=None,sep=',', compression='gzip')
oib_to_anto = oib_to_anto[['Line', 'ANTOSTRAT_basement', 'filt_clip']]
oib_to_anto.rename(columns={'filt_clip':'OIB_basement'}, inplace=True)
oib_to_anto['dif'] = oib_to_anto.ANTOSTRAT_basement - oib_to_anto.OIB_basement
oib_to_anto['abs_dif'] = np.abs(oib_to_anto.dif)
# ROSETTTA to OIB comparison
rosetta_to_oib = pd.read_csv("../data/results/rs_oib_basement.csv", header=0, index_col=None,sep=',', compression='gzip')
rosetta_to_oib = rosetta_to_oib[['Line', 'filt_clip','filt_clip_oib']]
rosetta_to_oib.rename(columns={'filt_clip':'ROSETTA_basement', 'filt_clip_oib':'OIB_basement'}, inplace=True)
rosetta_to_oib['dif'] = rosetta_to_oib.ROSETTA_basement - rosetta_to_oib.OIB_basement
rosetta_to_oib['abs_dif'] = np.abs(rosetta_to_oib.dif)
# prep OIB & ROSETTA magnetics data
# these csv are the outputs of the Werner deconvolution process, which was performed in Geosoft Oasis Montaj.
ROSETTA='../data/input/Geosoft_mag_data.csv.gz'
OIB='../data/input/Geosoft_Werner_output.csv.gz'
# load as pandas dataframes
df_rs = pd.read_csv(ROSETTA, index_col=None, compression='gzip')
df_oib = pd.read_csv(OIB, index_col=None, compression='gzip')
# drop unnecessary columns
df_oib.drop(df_oib.columns.difference(['x','y','Line','Mag_anom_draped_1000']),axis=1, inplace=True)
# drop unneccessary lines, and NaNs
df_oib_403=df_oib[df_oib.Line.isin(['L403.1', 'L403.3'])].copy()
df_oib_404=df_oib[df_oib.Line.isin(['L404.590', 'L404.650', 'L590', 'L650'])].copy()
df_rs.dropna(inplace=True)
df_oib_403.dropna(inplace=True)
df_oib_404.dropna(inplace=True)
df_oib_404_590=df_oib_404[df_oib_404.Line.isin(['L404.590', 'L590'])].copy()
df_oib_404_590.dropna(how='any', inplace=True)
# -
# ## Set figure parameters
# inset map details
e= -760_000
n= -2_200_000
w= 600_000
s= -350_000
#set figure height, width and map scale
fig_height = 50 # in mm
fig_width = fig_height*(w-e)/(s-n)
fig_ratio = (s-n)/(fig_height/1000)
# Make a GMT region string and projection strings in both ESPG3031 and Lat/Lon
fig_reg = f'{e}/{w}/{n}/{s}'
fig_proj = f"x1:{fig_ratio}"
# ## Plot figure
# +
fig = pygmt.Figure()
data = oib_to_anto.abs_dif
upper_lim = 14
with pygmt.config(FONT_LABEL= '18p,black', FONT_ANNOT='14p,black'):
fig.histogram(data=data, projection='X14c',
region=[data.min(), data.max(), 0, upper_lim],
fill='grey',
frame=['SW','xa500f100+l"OIB to ANTOSTRAT misfit (m)"', 'ya2f.5+l"Frequency"+u" %"'],
series=100, # bar width in meters
pen='.1p',
histtype=1, # % of total data
)
# add lines for mean / median valus
fig.plot(x=[data.mean(),data.mean()], y=[0,100], pen='3p,blue,-')
fig.plot(x=[data.median(),data.median()], y=[0,100], pen='3p,darkorange,-')
# add mean values as text
fig.text(x=data.mean(), y=upper_lim-upper_lim*.2, text=f' mean: {int(data.mean())} m',
justify='ML', font='16p,Helvetica-Bold,blue', fill='white')
# add mean values as text
fig.text(x=data.median(), y=upper_lim-upper_lim*.1, text=f' median: {int(data.median())} m',
justify='ML', font='16p,Helvetica-Bold,darkorange', fill='white')
# Add inset map to show location
with fig.inset(position=f'JTR+w{fig_width}c+o-6c/-6'):
fig.plot(region=fig_reg, projection=fig_proj,
data = gpd.read_file('../data/shapefiles/Coastline_Antarctica_v02.shp'), color = 'lightblue')
fig.plot(region=fig_reg, projection=fig_proj,
data = gpd.read_file('../data/shapefiles/GroundingLine_Antarctica_v02.shp'), color = 'grey')
fig.plot(region=fig_reg, projection=fig_proj,
data = gpd.read_file('../data/shapefiles/Coastline_Antarctica_v02.shp'), frame=["nsew"], pen='0.5p,black')
# plot flight paths of OIB 403
fig.plot(region=fig_reg, projection=fig_proj,
x=df_oib_403.x[df_oib_403.Line=='L403.1'], y=df_oib_403.y[df_oib_403.Line=='L403.1'],
pen='1p,black,4_2:2p')
fig.plot(region=fig_reg, projection=fig_proj,
x=df_oib_403.x[df_oib_403.Line=='L403.3'], y=df_oib_403.y[df_oib_403.Line=='L403.3'],
pen='1p,black,4_2:2p')
"""
Shift origin and add sublot to right
"""
fig.shift_origin(xshift='15c')
data = rosetta_to_oib.abs_dif
upper_lim = 8
with pygmt.config(FONT_LABEL= '18p,black', FONT_ANNOT='14p,black'):
fig.histogram(data=data, projection='X14c',
region=[data.min(), data.max(), 0, upper_lim],
fill='grey',
frame=['SW','xa500f100+l"ROSETTA to OIB misfit (m)"', 'ya2f.5+l"Frequency"+u" %"'],
series=50, # bar width in meters
pen='.1p',
histtype=1, # % of total data
)
# add lines for mean / median valus
fig.plot(x=[data.mean(),data.mean()], y=[0,100], pen='3p,blue,-')
fig.plot(x=[data.median(),data.median()], y=[0,100], pen='3p,darkorange,-')
# add mean values as text
fig.text(x=data.mean(), y=upper_lim-upper_lim*.2, text=f' mean: {int(data.mean())} m',
justify='ML', font='16p,Helvetica-Bold,blue', fill='white')
# add mean values as text
fig.text(x=data.median(), y=upper_lim-upper_lim*.1, text=f' median: {int(data.median())} m',
justify='ML', font='16p,Helvetica-Bold,darkorange', fill='white')
# Add inset map to show location
with fig.inset(position=f'JTR+w{fig_width}c+o-6c/-6'):
fig.plot(region=fig_reg, projection=fig_proj,
data = gpd.read_file('../data/shapefiles/Coastline_Antarctica_v02.shp'), color = 'lightblue')
fig.plot(region=fig_reg, projection=fig_proj,
data = gpd.read_file('../data/shapefiles/GroundingLine_Antarctica_v02.shp'), color = 'grey')
fig.plot(region=fig_reg, projection=fig_proj,
data = gpd.read_file('../data/shapefiles/Coastline_Antarctica_v02.shp'), frame=["nsew"], pen='0.5p,black')
# plot flight paths of OIB 404
fig.plot(region=fig_reg, projection=fig_proj,
x=df_oib_404_590.x[df_oib_404_590.Line=='L590'], y=df_oib_404_590.y[df_oib_404_590.Line=='L590'],
pen='1.2p,black,4_2:2p')
fig.plot(region=fig_reg, projection=fig_proj,
x=df_oib_404.x[df_oib_404.Line=='L404.650'], y=df_oib_404.y[df_oib_404.Line=='L404.650'],
pen='1.2p,black,4_2:2p')
fig.show(width=400)
# -
# ## Save figure
out='outputs/FigS6_misfits.jpg'
fig.savefig(out, dpi=400)
| Figures/FigS6_misfits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## This is an end to end example of using ArcGIS with python
# this has been mostly followed as a simplefied version from here https://github.com/orhuna/RF-Demo/blob/master/RF_Demo_Fl.ipynb
#
# For better visualization, maps have been published into the GIS server with a public read access to display it in the notebook cell
#
#
from sklearn.ensemble import RandomForestClassifier
import numpy as NUM
import arcpy as ARCPY
import arcpy.da as DA
import pandas as PD
import seaborn as SEA
import matplotlib.pyplot as PLOT
import pandas as PD
import arcgisscripting as ARC
import arcpy as ARCPY
import SSUtilities as UTILS
from arcgis.gis import *
import os as OS
mygis = GIS()
from IPython.display import IFrame
def show_web_map_by_url(URL):
url = 'http://www.arcgis.com/home/webmap/viewer.html?url=' + URL
return IFrame(url, width='100%', height=500)
analysis_map_URL = 'https://services3.arcgis.com/oZfKvdlWHN1MwS48/ArcGIS/rest/services/MachineLearningSeagrass/FeatureServer/1&source=sd'
# This is the initial map which has florida data to start the data analysis
# +
#Where the seagrasses are in florida
show_web_map_by_url(analysis_map_URL)
# -
#Load the features from the shapefile stored in GIS Desktop
inputDir = 'C:/GISDemo/SeaGrass/SeaGrass.gdb/FloridaSeaGrass'
#Names of Prediction Variables
predictVars = ['salinity', 'temp', 'phosphate','nitrate',
'silicate', 'dissO2', 'NameEMU']
#Name of Classification Variable
classVar = ['Present']
#List of all Variables
allVars = predictVars + classVar
# +
#GIS data to Pandas dataframe
#Get Data from ArcGIS Desktop
FlLayer = DA.FeatureClassToNumPyArray(inputDir, ["SHAPE@XY"] + allVars)
#Obtain Spatial Reference
spatRef = ARCPY.Describe(inputDir).spatialReference
#Define Main Dataframe
data = PD.DataFrame(FlLayer, columns = allVars)
#Display Portion of the Data Frame
data.head()
# -
#Process Categorical Data for Analysis flatten the categorical data
#Create Numeric Fields for One-Hot Encoding of the Categorical Variable
catVars = PD.get_dummies(data[predictVars[-1]])
#Remove raw Categories from Dataset
data = data.drop(predictVars[-1], axis = 1)
#Add Processed Categories Back into the Data Frame
data = data.join(catVars)
#Abbreviate Long Categorical Variable Names
newNames = ['c1','c2','c3']
for ind, name in enumerate(newNames):
data.rename(columns={data.columns[len(predictVars)+ind]:name}, inplace=True)
#Update Predict Variable Names
predictVarsNew = predictVars[:-1] + newNames
#Display Portion of the Data Frame
data.head()
# +
##EVALUATE CORRELATION BETWEEN PREDICTORS- EXCLUDE CATEGORICAL
#Calculate Correlation Coefficient between Prediction Variables
corr = data.drop(data.columns[-3:], axis = 1).astype('float64').corr()
#Plot Correlation Matrix Between Prediction Variables
ax = SEA.heatmap(corr, cmap=SEA.diverging_palette(220, 10, as_cmap=True),
square=True, annot = True, linecolor = 'k', linewidths = 1)
PLOT.show()
# -
##PERFORM RANDOM FOREST CLASSIFICATION
#Fraction of data to be used in Training
fracNum = 0.3
#Seperate the Data into Training and Test Datasets
train_c1 = data[data[classVar[0]] == 1].sample(frac = fracNum)
train_c0 = data[data[classVar[0]] == 0].sample(frac = fracNum)
#Create Training Dataset
train_set = PD.concat([train_c0, train_c1])
#Create Testing Dataset
test_set = data.drop(train_set.index)
#Encode Seagrass Presence as Classes
indicator, _ = PD.factorize(train_set[classVar[0]])
#Print Test and Train Data Set Sizes
print('Training Data Size = ' + str(train_set.size))
print('Test Data Size = ' + str(test_set.size))
##Create Random Forest Classification Object
rfco = RandomForestClassifier(n_estimators = 500, oob_score = True)
#Perform Classification Using Training Set
rfco.fit(train_set[predictVars[:-1]], indicator)
#Plot Variable Importance
#Dummy Variables for Y-Ticks
#Predict Seagrass Occurance for the Test Dataset
seagrassPred = rfco.predict(test_set[predictVars[:-1]])
#Calculate Prediction Accuracy
test_seagrass = test_set['Present'].values.flatten()
#Calculate Estimation Error
error = NUM.sum(test_seagrass - seagrassPred)/len(seagrassPred) * 100
#Print Accuracy Metrics
print('Accuracy = ' + str(100 - NUM.abs(error)) + ' % ')
print('Locations with Seagrass = ' + str(len(NUM.where(test_seagrass==1)[0])) )
print('Predicted Locations with Seagrass = ' + str(len(NUM.where(seagrassPred==1)[0])))
#Delete the feature as ARCPY.da.NumPyArrayToFeatureClass does not overwrite
#ARCPY.DeleteFeatures_management("C:/GISDemo/SeaGrass/SeaGrass.gdb\Florida_Seagrass_Prediction")
##BRING OUTPUT BACK INTO ArcGIS Desktop
#Allow overwriting Feature Classes
ARCPY.env.overwriteOutput = True
#Get Indexes for the Test Dataset
outputDir = r'C:/GISDemo/SeaGrass/SeaGrass.gdb'
nameFC = 'Florida_Seagrass_Prediction_Py'
#Locations with Seagrass
grassExists = FlLayer[["SHAPE@XY"]][test_set.index[NUM.where(seagrassPred==1)]]
# Write Locations with Seagrass to Feature Class
ARCPY.da.NumPyArrayToFeatureClass(grassExists, OS.path.join(outputDir, nameFC), ['SHAPE@XY'], spatRef)
predicted_fl_map_url ='https://www.arcgis.com/home/webmap/viewer.html?webmap=2e65c37b14764e7a875efb5b5bad4633'
IFrame(predicted_fl_map_url, width='100%', height=500)
# Now create a model from the USA data set and we will use this to predict the sea grass locations worldwide
#Names of Prediction Variables#Import USA Seagrass Data
inputDir = 'C:/GISDemo/SeaGrass/SeaGrass.gdb/USASeaGrass_training'
predictVars = ['salinity', 'temp', 'phosphate','nitrate',
'silicate', 'dissO2']
#Name of Classification Variable
classVar = ['Present']
#List of all Variables
allVars = predictVars + classVar
#Create a Data Object
USAlayer = DA.FeatureClassToNumPyArray(inputDir, ["SHAPE@XY"] + allVars)
#Obtain Spatial Reference
spatRefGlobal = ARCPY.Describe(inputDir).spatialReference
#Define Main Dataframe
USA_Train = PD.DataFrame(USAlayer, columns = allVars)
#Display Portion of the Data Frame
USA_Train.head()
##Train Random Forest Using USA Data
#Encode Seagrass Presence as Classes
indicatorUSA, _ = PD.factorize(USA_Train['Present'])
#Create Random Forest Classification Object
rfco = RandomForestClassifier(n_estimators = 500)
#Perform Classification Using Training Set
rfco.fit(USA_Train[predictVars[:-1]], indicatorUSA)
predictVars[:-1]
## Import Global Data for Prediction ##
##globalUrl = r'https://services3.arcgis.com/oZfKvdlWHN1MwS48/arcgis/rest/services/MachineLearningSeagrass/FeatureServer/2?token=<KEY>.'
globalUrl = 'C:/GISDemo/SeaGrass/SeaGrass.gdb/Global_Seagrass_Prediction'
#Create a SS Data Object
globalData = DA.FeatureClassToNumPyArray(globalUrl, ["SHAPE@XY"] + predictVars[:-1])
#Obtain Spatial Reference
spatRefGlobal = ARCPY.Describe(globalUrl).spatialReference
#Define Dataframe
globalTrain = PD.DataFrame(globalData, columns = predictVars[:-1])
#Predict Global Seagrass Occurance
seagrassPredGlobal = rfco.predict(globalTrain)
#Bring Output Back to ArcGIS
nameFC = 'GlobSeagrass'
outputDir = r'C:/GISDemo/SeaGrass/SeaGrass.gdb'
nameFC = 'Global_Seagrass_Prediction_Done'
grassExists = globalData[["SHAPE@XY"]][globalTrain.index[NUM.where(seagrassPredGlobal==1)]]
# Write Locations with Seagrass to Feature Class
ARCPY.da.NumPyArrayToFeatureClass(grassExists, OS.path.join(outputDir, nameFC), ['SHAPE@XY'], spatRefGlobal)
predicted_gl_map_url='https://www.arcgis.com/home/webmap/viewer.html?webmap=503efd8acb7b48e3b92b4f4ac4d6e82e'
# Global prediction data as shown on the map
IFrame(predicted_gl_map_url, width='100%', height=500)
| Notebooks/ArcGIS/Python walkthrough ArcGIS Data analysis and ML.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala 2.11 with Spark 2.0
// language: scala
// name: scala-spark20
// ---
// <table style="border: none" align="left">
// <tr style="border: none">
// <th style="border: none"><font face="verdana" size="5" color="black"><b>Predict outdoor equipment purchase with IBM Watson Machine Learning</b></th>
// <th style="border: none"><img src="https://github.com/pmservice/customer-satisfaction-prediction/blob/master/app/static/images/ml_icon_gray.png?raw=true" alt="Watson Machine Learning icon" height="40" width="40"></th>
// </tr>
// <tr style="border: none">
// <th style="border: none"><img src="https://github.com/pmservice/wml-sample-models/blob/master/spark/product-line-prediction/images/products_graphics.png?raw=true" alt="Icon" width="800"> </th>
// </tr>
// </table>
// This notebook contains steps and code to get data from the IBM Data Science Experience Community, create a predictive model, and start scoring new data. This notebook introduces commands for getting data and for basic data cleaning and exploration, pipeline creation, model training, model persistance to Watson Machine Learning repository, model deployment, and scoring.
//
// Some familiarity with Scala is helpful. This notebook uses Scala 2.11 and Apache® Spark 2.0.
//
// You will use a publicly available data set, **GoSales Transactions for Naive Bayes Model**, which details anonymous outdoor equipment purchases. Use the details of this data set to predict clients' interests in terms of product line, such as golf accessories, camping equipment, and others.
//
// ## Learning goals
//
// The learning goals of this notebook are:
//
// - Load a CSV file into an Apache® Spark DataFrame.
// - Explore data.
// - Prepare data for training and evaluation.
// - Create an Apache® Spark machine learning pipeline.
// - Train and evaluate a model.
// - Persist a pipeline and model in Watson Machine Learning repository.
// - Deploy a model for online scoring using Wastson Machine Learning API.
// - Score sample scoring data using the Watson Machine Learning API.
//
//
// ## Contents
//
// This notebook contains the following parts:
//
// 1. [Setup](#setup)
// 2. [Load and explore data](#load)
// 3. [Create spark ml model](#model)
// 4. [Persist model](#persistence)
// 5. [Predict locally and visualize](#visualization)
// 6. [Deploy and score in a Cloud](#scoring)
// 7. [Summary and next steps](#summary)
// <a id="setup"></a>
// ## 1. Setup
//
// Before you use the sample code in this notebook, you must perform the following setup tasks:
//
// - Create a [Watson Machine Learning Service](https://console.ng.bluemix.net/catalog/services/ibm-watson-machine-learning/) instance (a free plan is offered).
// - Upload **GoSales Transactions** data as a data asset in IBM Data Science Experience.
// - Make sure that you are using a Spark 2.0 kernel.
//
// ### Create the GoSales Transactions Data Asset
//
// The GOSales data is a freely available data set on the Data Science Experience home page.
//
// 1. Go to the [GoSales Transactions for Naive Bayes Model](https://apsportal.ibm.com/exchange-api/v1/entries/8044492073eb964f46597b4be06ff5ea/data?accessKey=9561295fa407698694b1e254d0099600) data card on the Data Science Experience **Community** page and open the card by double-clicking it.
// 2. Click the link icon.
// 4. Select the link, copy it by pressing Ctrl+C, and then, click **Close**.
// 5. In the following cell, replace the **link_to_data** variable value with the link.
// <a id="load"></a>
// ## 2. Load and explore data
//
// In this section you will load the data as an Apache® Spark DataFrame and perform a basic exploration.
//
// Load the data to the Spark DataFrame by using *wget* to upload the data to gpfs and then *read* method.
// +
import scala.sys.process._
"wget https://apsportal.ibm.com/exchange-api/v1/entries/8044492073eb964f46597b4be06ff5ea/data?accessKey=9561295fa407698694b1e254d0099600".!
// -
val filename = "data?accessKey=9561295fa407698694b1e254d0099600"
val df_data = spark.
read.format("org.apache.spark.sql.execution.datasources.csv.CSVFileFormat").
option("header", "true").
option("inferSchema", "true").
load(filename)
df_data.take(5)
// Explore the loaded data by using the following Apache® Spark DataFrame methods:
// - print schema
// - print top ten records
// - count all records
df_data.printSchema()
// As you can see, the data contains five fields. PRODUCT_LINE field is the one we would like to predict (label).
df_data.show()
df_data.count()
// As you can see, the data set contains 60252 records.
// <a id="model"></a>
// ## 3. Create an Apache® Spark machine learning model
//
// In this section you will learn how to prepare data, create an Apache® Spark machine learning pipeline, and train a model.
// ### 3.1: Prepare data
//
// In this subsection you will split your data into: train, test and predict datasets.
// +
val splits = df_data.randomSplit(Array(0.8, 0.18, 0.02), seed = 24L)
val training_data = splits(0).cache()
val test_data = splits(1)
val prediction_data = splits(2)
println("Number of training records: " + training_data.count())
println("Number of testing records: " + test_data.count())
println("Number of prediction records: " + prediction_data.count())
// -
// As you can see our data has been successfully split into three datasets:
//
// - The train data set, which is the largest group, is used for training.
// - The test data set will be used for model evaluation and is used to test the assumptions of the model.
// - The predict data set will be used for prediction.
// ### 3.2: Create pipeline and train a model
// In this section you will create an Apache® Spark machine learning pipeline and then train the model.
//
// In the first step you need to import the Apache® Spark machine learning packages that will be needed in the subsequent steps.
// +
// Spark ML libraries
import org.apache.spark.ml.classification.RandomForestClassifier
import org.apache.spark.ml.feature.{OneHotEncoder, StringIndexer, IndexToString, VectorAssembler}
import org.apache.spark.ml.evaluation.MulticlassClassificationEvaluator
import org.apache.spark.ml.{Model, Pipeline, PipelineStage, PipelineModel}
import org.apache.spark.sql.SparkSession
// -
// In the following step, convert all the string fields to numeric ones by using the StringIndexer transformer.
val stringIndexer_label = new StringIndexer().setInputCol("PRODUCT_LINE").setOutputCol("label").fit(df_data)
val stringIndexer_prof = new StringIndexer().setInputCol("PROFESSION").setOutputCol("PROFESSION_IX")
val stringIndexer_gend = new StringIndexer().setInputCol("GENDER").setOutputCol("GENDER_IX")
val stringIndexer_mar = new StringIndexer().setInputCol("MARITAL_STATUS").setOutputCol("MARITAL_STATUS_IX")
// In the following step, create a feature vector by combining all features together.
val vectorAssembler_features = new VectorAssembler().setInputCols(Array("GENDER_IX", "AGE", "MARITAL_STATUS_IX", "PROFESSION_IX")).setOutputCol("features")
// Next, define estimators you want to use for classification. Random Forest is used in the following example.
val rf = new RandomForestClassifier().setLabelCol("label").setFeaturesCol("features").setNumTrees(10)
// Finally, indexed labels back to original labels.
val labelConverter = new IndexToString().setInputCol("prediction").setOutputCol("predictedLabel").setLabels(stringIndexer_label.labels)
// Let's build the pipeline now. A pipeline consists of transformers and an estimator.
val pipeline_rf = new Pipeline().setStages(Array(stringIndexer_label, stringIndexer_prof, stringIndexer_gend, stringIndexer_mar, vectorAssembler_features, rf, labelConverter))
// Now, you can train your Random Forest model by using the previously defined **pipeline** and **training data**.
training_data.printSchema()
val model_rf = pipeline_rf.fit(training_data)
// You can check your **model accuracy** now. To evaluate the model, use **test data**.
val predictions = model_rf.transform(test_data)
val evaluatorRF = new MulticlassClassificationEvaluator().setLabelCol("label").setPredictionCol("prediction").setMetricName("accuracy")
val accuracy = evaluatorRF.evaluate(predictions)
println("Accuracy = " + accuracy)
println("Test Error = " + (1.0 - accuracy))
// You can tune your model now to achieve better accuracy. For simplicity of this example tuning section is omitted.
// <a id="persistence"></a>
// ## 4. Persist model
//
// In this section you will learn how to store your pipeline and model in Watson Machine Learning repository by using Scala client libraries.
//
// First, you must import client libraries.
//
// **Note**: Apache® Spark 2.0 or higher is required.
// +
// WML client library
import com.ibm.analytics.ngp.repository._
// Helper libraries
import scalaj.http.{Http, HttpOptions}
import scala.util.{Success, Failure}
import java.util.Base64
import java.nio.charset.StandardCharsets
import play.api.libs.json._
// -
// Authenticate to Watson Machine Learning service on Bluemix.
//
// **Action**: Put authentication information from your instance of Watson Machine Learning service here.</div>
val service_path = "https://ibm-watson-ml.mybluemix.net"
val instance_id = "***"
val username = "***"
val password = "***"
// **Tip**: service_path, user and password can be found on **Service Credentials** tab of service instance created in Bluemix. If you cannot see **instance_id** field in **Serice Credentials** generate new credentials by pressing **New credential (+)** button.
val client = MLRepositoryClient(service_path)
client.authorize(username, password)
// Create model artifact (abstraction layer).
val model_artifact = MLRepositoryArtifact(model_rf, training_data, "WML Product Line Prediction Model")
// **Tip**: The MLRepositoryArtifact method expects a trained model object, training data, and a model name. (It is this model name that is displayed by the Watson Machine Learning service).
// ### 4.1: Save pipeline and model
//
// In this subsection you will learn how to save pipeline and model artifacts to your Watson Machine Learning instance.
val saved_model = client.models.save(model_artifact).get
// Get saved model metadata from Watson Machine Learning.
//
// **Tip**: Use *meta.availableProps* to get the list of available props.
saved_model.meta.availableProps
println("modelType: " + saved_model.meta.prop("modelType"))
println("trainingDataSchema: " + saved_model.meta.prop("trainingDataSchema"))
println("creationTime: " + saved_model.meta.prop("creationTime"))
println("modelVersionHref: " + saved_model.meta.prop("modelVersionHref"))
println("label: " + saved_model.meta.prop("label"))
// **Tip**: **modelVersionHref** is our model unique indentifier in the Watson Machine Learning repository.
// ### 4.2: Load model and make predictions
//
// In this subsection you will learn how to load back saved model from specified instance of Watson Machine Learning.
val model_version_href = saved_model.meta.prop("modelVersionHref").get
val loaded_model_artifact = client.models.version(model_version_href).get
// You can print for example model name to make sure that model artifact has been loaded correctly.
loaded_model_artifact.name.mkString
// As you can see the name is correct.
loaded_model_artifact match {
case SparkPipelineModelLoader(Success(model)) => {
val predictions = model.transform(prediction_data)
}
case SparkPipelineModelLoader(Failure(e)) => "Loading failed."
case _ => println(s"Unexpected artifact class: ${loaded_model_artifact.getClass}")
}
predictions.select("GENDER", "AGE", "MARITAL_STATUS", "PROFESSION","predictedLabel").show()
// By tabulating a count, you can see which product line is the most popular.
predictions.select("predictedLabel").groupBy("predictedLabel").count().show()
// You have already learned how save and load the model from Watson Machine Learning repository.
// <a id="scoring"></a>
// ## 5. Deploy and score in a Cloud
//
// In this section you will learn how to create online scoring and to score a new data record by using the Watson Machine Learning REST API.
// For more information about REST APIs, see the [Swagger Documentation](http://watson-ml-api.mybluemix.net/).
//
// To work with the Watson Machine Leraning REST API you must generate an access token. To do that you can use the following sample code:
// +
// Get WML service instance token
val wml_auth_header = "Basic " + Base64.getEncoder.encodeToString((username + ":" + password).getBytes(StandardCharsets.UTF_8))
val wml_url = service_path + "/v3/identity/token"
val wml_response = Http(wml_url).header("Authorization", wml_auth_header).asString
val wmltoken_json: JsValue = Json.parse(wml_response.body)
val wmltoken = (wmltoken_json \ "token").asOpt[String] match {
case Some(x) => x
case None => ""
}
// -
wmltoken
// ### 5.1: Create online scoring endpoint
//
// Now you can create an online scoring endpoint. Execute the following sample code that uses the publishedModelId value to create the scoring endpoint to the Bluemix repository.
// #### Get published_models url from instance details
val endpoint_instance = service_path + "/v3/wml_instances/" + instance_id
val wml_response_instance = Http(endpoint_instance).header("Content-Type", "application/json").header("Authorization", "Bearer " + wmltoken).option(HttpOptions.connTimeout(10000)).option(HttpOptions.readTimeout(50000)).asString
wml_response_instance
val published_models_json: JsValue = Json.parse(wml_response_instance.body)
val published_models_url = (((published_models_json \ "entity") \\ "published_models")(0) \ "url").as[JsString].value
published_models_url
// #### Get list of published models
val wml_models = Http(published_models_url).header("Content-Type", "application/json").header("Authorization", "Bearer " + wmltoken).option(HttpOptions.connTimeout(10000)).option(HttpOptions.readTimeout(50000)).asString
wml_models
var deployment_endpoint: String = _
wml_models.body.split("\"").map{ s => {if ((s contains "deployments") & (s contains saved_model.uid.mkString)) {deployment_endpoint = s}}}
deployment_endpoint
// #### Create online deployment for published model
val payload_name = "Online scoring"
val payload_data_online = Json.stringify(Json.toJson(Map("type" -> "online", "name" -> payload_name)))
val response_online = Http(deployment_endpoint).postData(payload_data_online).header("Content-Type", "application/json").header("Authorization", "Bearer " + wmltoken).option(HttpOptions.connTimeout(50000)).option(HttpOptions.readTimeout(50000)).asString
val scoring_url_json: JsValue = Json.parse(response_online.body)
val scoring_url = (scoring_url_json \ "entity" \ "scoring_url").asOpt[String] match {
case Some(x) => x
case None => ""
}
print(scoring_url)
val payload_scoring = Json.stringify(Json.toJson(Map("fields" -> Json.toJson(List(Json.toJson("GENDER"), Json.toJson("AGE"), Json.toJson("MARITAL_STATUS"), Json.toJson("PROFESSION"))),
"values" -> Json.toJson(List(List(Json.toJson("M"), Json.toJson(55), Json.toJson("Single"), Json.toJson("Executive")))))))
payload_scoring
// Now, you can send (POST) new scoring records (new data) for which you would like to get predictions. To do that, execute the following sample code:
val response_scoring = Http(scoring_url).postData(payload_scoring).header("Content-Type", "application/json").header("Authorization", "Bearer " + wmltoken).option(HttpOptions.method("POST")).option(HttpOptions.connTimeout(10000)).option(HttpOptions.readTimeout(50000)).asString
print(response_scoring)
// As we can see we predict that a 55-year-old single male executive is interested in Mountaineering Equipment (prediction: 2.0).
// <a id="summary"></a>
// ## 6. Summary and next steps
//
// You successfully completed this notebook! You learned how to use Apache Spark machine learning as well as Watson Machine Learning for model creation and deployment. Check out our _[Online Documentation](https://console.ng.bluemix.net/docs/services/PredictiveModeling/pm_service_api_spark.html)_ for more samples, tutorials, documentation, how-tos, and blog posts.
//
// ### Authors
//
// **<NAME>** is Data Scientist in IBM with a track record of developing enterprise-level applications that substantially increases clients' ability to turn data into actionable insights.
//
// Copyright © 2017 IBM. This notebook and its source code are released under the terms of the MIT License.
| JupyterNotebooks/Training/DSX/DSX-From+spark+ml+model+to+online+scoring+with+scala.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Academic Integrity Statement
#
# As a matter of Departmental policy, **we are required to give you a 0** unless you **type your name** after the following statement:
#
# > *I certify on my honor that I have neither given nor received any help, or used any non-permitted resources, while completing this evaluation.*
#
# \[TYPE YOUR NAME HERE\]
#
# # Problem 1 (50 points)
#
# Rampant disinformation---often called "fake news"---has emerged as one of the fundamental crises over our time.
#
# <figure class="image" style="width:30%">
# <img src="https://s3.amazonaws.com/libapps/accounts/63707/images/21392935.jpg" alt="A portrait of <NAME>, wearing a purple suit and brown top hat. His face is condescending. The caption reads: 'Oh, so you read it on the internet? Well then I guess it must be true.'">
# <figcaption><i></i></figcaption>
# </figure>
#
# There is a growing movement for online platforms to regulate fake news. Doing so at scale requires combing through millions of news items every day, making it very expensive to do by hand. Can an algorithm do it instead?
#
# The following two URLs each contain part of a data set.
#
# - **Fake news items**: `https://raw.githubusercontent.com/PhilChodrow/PIC16A/master/datasets/fake_news/Fake.csv`
# - **Real news items**: `https://raw.githubusercontent.com/PhilChodrow/PIC16A/master/datasets/fake_news/true.csv`
#
# Use the data at these urls to **construct a fake news classifier.**
#
# 1. Your model must be able to **make predictions** about whether or not an unseen news item is fake or real.
# 2. Because fake news models must be able to make millions of predictions per day, it must be able to make predictions very quickly. More columns mean more computation time. **Your final model should use no more than 50 columns.**
#
# You are free to create any columns that you need, and to use any functions that we have or have not covered in the course. You may also use any machine learning model.
#
# Please use Markdown headers with \#\# signs to clearly distinguish the different stages of your solution.
#
# ### Requirements
#
# 1. Any operations that you perform multiple times (such processing that you perform on both the training and test sets) must be contained in function with informative docstrings. Comments and explanations are expected throughout. It is especially important to explain how you chose the columns to use in your final model.
# 2. You should not use for-loops to iterate over the rows of data frames or arrays.
# 3. You must fit your model on the training data, and not use the test data for fitting.
#
# ### Hints
#
# - `pd.concat()` is a good way to combine data frames.
# - Try fitting a model with as many columns as you want first. See if you can get a representation of which columns are important, and then select your final columns from this list.
# - In class, we talked about greedy stagewise feature selection and exhaustive enumeration for determining a good set of columns. Neither of these methods are recommended for this problem.
# - If you want to be creative about your model choice, then please go for it. If you want a safe option, try logistic regression.
# - If a model takes too long to fit on the full data set, try fitting it on, say, 10% of the data.
# - You might find the some of the [cheatsheets](https://philchodrow.github.io/PIC16A/resources/) to be helpful.
#
# ### Rubric
#
# - (**15 points**): clearly written code that makes economical use of skills from the course to manipulate data.
# - (**15 points**): comments, explanatory surrouding text, and docstrings for any functions and classes.
# - (**20 points**): computed according to the formula `20 x score`, where `score` is your model's prediction performance on unseen data. Models that use more than 50 columns can receive up to 15 of these points. Scores will be rounded up. For example, if you obtain an 84% predictive performance with 50 columns, then the score of `20 x 0.84 = 16.8` will be rounded up to 17 points.
| prior_exams/F20/final/final-P1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# conda environment to use on nightingale: /scratch/conda_envs/elmo-embeddings
# %load_ext autoreload
# +
# %autoreload
from allennlp.commands.elmo import ElmoEmbedder
import os
from sys import path
path.append('..')
from relation_extraction.data import utils
import h5py
import numpy as np
data_path = '/data/medg/misc/semeval_2010'
def res(path): return os.path.join(data_path, path)
# -
elmo = ElmoEmbedder()
tokens = ["I", "ate", "an", "apple", "for", "breakfast"]
vectors = elmo.embed_sentence(tokens)
assert(len(vectors) == 3) # one for each layer in the ELMo output
assert(len(vectors[0]) == len(tokens)) # the vector elements correspond with the input tokens
# +
# # the dimensionality of this vector is layer in ELMO LSTM X num_of_words_in_sentence X the size of the ELMO embedding
# vectors
# -
# In order to incorporate ELMO into my model (well the very first thing is just to use the first layer) and then beyond that use the other layers. But in order to map a sentence to the ELMO representation (which is the larger goal), I should re-generate my pickle files to include the embedding of the sentence with it so I can finally use a linearly weighted combinations of the embeddings from the different layers of the LSTM. One important question is though, that they claim that the very first layer includes context insensitive embeddings. Does this mean that in two sentences, if I have the word "claims," they would both have the same vector? Let's check
new_sentence = ["The", "apple", "that", "I", "ate", "was", "not", "delicious"]
vector2 = elmo.embed_sentence(new_sentence)
assert(len(vector2) == 3)
assert(len(vector2[0] == len(new_sentence)))
vector2[0][2]
vectors[0][3]
# Nope so the embeddings are different, so this is not the best way to generate individual word embeddings via elmo
# # Simplest possible way of generating embedding file similar to Glove/Senna etc
# Read the train and test file of Semeval data, get the tokenized version of the sentence and then feed in individual word to generate an embedding for
res('train.txt')
def get_duplicate_words_from_file(filename):
words = []
with open(res(filename)) as f:
for line in f:
line = line.strip().lower().split()
sentence = line[5:]
words.extend(sentence)
return words
dup_words = get_duplicate_words_from_file('train.txt') + get_duplicate_words_from_file('test.txt')
words = list(set(dup_words))
vectorlist = []
# +
# consider doing some pre processing of the data where you either turn it into a number or some range of numbers
# -
for word in words:
vectors = elmo.embed_sentence([word])
assert(len(vectors) == 3) # one for each layer in the ELMo output
assert(len(vectors[0]) == 1) # the vector elements correspond with the input tokens
# grab the first layer and then write that into a file
vector = vectors[0][0]
list_a = [word]
list_a.extend(vector)
vectorlist.append(list_a)
# +
# word = words[0]
# vectors = elmo.embed_sentence([word])
# assert(len(vectors) == 3) # one for each layer in the ELMo output
# assert(len(vectors[0]) == 1) # the vector elements correspond with the input tokens
# # grab the first layer and then write that into a file
# vector = vectors[0][0]
# list_a = [word]
# list_a.extend(vector)
# vectorlist.append(list_a)
# -
vectorlist
with open(res('elmo/elmo.layer1.txt'), 'w') as f:
for vector in vectorlist:
vectorstring = ""
for val in vector:
vectorstring = vectorstring + " " + str(val)
vectorstring = vectorstring.strip()
f.write(vectorstring + "\n")
f.close()
# ## Next, work on getting all the layers of the elmo embeddings
def get_sentences(filename):
data = open(res(filename))
data = utils.split_data_cut_sentence(data, 50)
sentences = data[0]
return sentences
train_sentences = get_sentences('train.txt')
test_sentences = get_sentences('test.txt')
len(train_sentences)
len(test_sentences)
def write_sentences_to_txt(sentences, filename):
with open(res('elmo/input-sentences/'+filename), 'w') as f:
for sentence in sentences:
sentence_string = " ".join(sentence)
f.write(sentence_string + "\n")
write_sentences_to_txt(train_sentences, 'train-sentences.txt')
write_sentences_to_txt(test_sentences, 'test-sentences.txt')
# Now, follow the instructions on https://github.com/allenai/allennlp/blob/master/tutorials/how_to/elmo.md to generate the h5py file
# Nope, that does not work
with open(res('elmo/input-sentences/train-sentences.txt')) as train_file:
elmo.embed_file(input_file=train_file,
output_file_path = res('elmo/train-elmo-full.hdf5'))
with open(res('elmo/input-sentences/test-sentences.txt')) as test_file:
elmo.embed_file(input_file=test_file,
output_file_path = res('elmo/test-elmo-full.hdf5'))
# ## Read the h5py file and write it as a np array on disk
def get_elmo_embeddings(filename):
h5py_file = h5py.File(filename, 'r')
elmo_embeddings = []
for i in range(0, len(h5py_file) - 1):
embedding = h5py_file.get(str(i))
elmo_embeddings.append(np.array(embedding))
return elmo_embeddings
elmo_embedding_train = get_elmo_embeddings(res('elmo/train-elmo-full.hdf5'))
elmo_embedding_test = get_elmo_embeddings(res('elmo/test-elmo-full.hdf5'))
np.save(res('elmo/train-elmo-full.npy'), elmo_embedding_train)
h5py_file = h5py.File("/data/medg/misc/semeval_2010/elmo/train-elmo-full.hdf5", 'r')
embedding = h5py_file.get("0")
embedding
np.array(embedding).shape
assert(len(embedding) == 3)
len(h5py_file)
# 8000 + (1 is extra) for all the sentences in the training set
# dimensions is layers, words_in_sentences, dimensions_per_layer
elmo_embed_train = []
for i in range(0, len(h5py_file) - 1):
embedding = h5py_file.get(str(i))
elmo_embed_train.append(np.array(embedding))
len(elmo_embed_train)
elmo_embed_train[0].shape
first_sentence = elmo_embed_train[0]
first_sentence.shape
max_len = 0
# because of the new line character, elmo generated embeddings for the blank line at the end.
for i in range(0, len(elmo_embed_train)):
sentence = elmo_embed_train[i]
if sentence.shape[1] > max_len:
max_len = sentence.shape[1]
# ## create a new embedding with the padding
new_elmo_embed_train=[]
for i in range(0, len(elmo_embed_train)):
sentence = elmo_embed_train[i]
num_of_words_to_pad = max_len - sentence.shape[1] # gives the length of the sentence
array_to_pad = np.zeros(shape=(sentence.shape[0], num_of_words_to_pad, sentence.shape[2]), dtype='float32')
appended_array = np.append(sentence, array_to_pad, axis=1) # appending along the sentence axis
new_elmo_embed_train.append(appended_array)
# try to pad the elmo_embed_train with 0's in the second dimension acc to max sentence length 18
a = np.array([1,2,3], dtype='float32')
np.append(a, 0)
toappend = np.zeros(shape=(3,1,1024), dtype='float32')
appended = np.append(first_sentence, toappend, axis=1)
appended.shape
# Let's make sure to see with a simple example
a = np.zeros(shape=(3,4,2), dtype='float32')
b = np.ones(shape=(3,1,2), dtype='float32')
np.append(a, b, axis=1)
elmo_embed_train[0].shape
# next will need to see how to append to the original train array
sentences = np.zeros((3,5), dtype=int)
e1 = np.ones((3,1), dtype=int)
batch = sentences, e1
batch
batch = list(zip(*batch))
batch = (x for x in zip(*batch))
batch
(sentences, e1) = batch
sentences
np.vstack(sentences)
e1
| notebooks/junk/Elmo-Embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import torch
torch.set_printoptions(edgeitems=2, threshold=50, linewidth=75)
bikes_numpy = np.loadtxt(
"../data/p1ch4/bike-sharing-dataset/hour-fixed.csv",
dtype=np.float32,
delimiter=",",
skiprows=1,
converters={1: lambda x: float(x[8:10])}) # <1>
bikes = torch.from_numpy(bikes_numpy)
bikes
bikes.shape, bikes.stride()
daily_bikes = bikes.view(-1, 24, bikes.shape[1])
daily_bikes.shape, daily_bikes.stride()
daily_bikes = daily_bikes.transpose(1, 2)
daily_bikes.shape, daily_bikes.stride()
first_day = bikes[:24].long()
weather_onehot = torch.zeros(first_day.shape[0], 4)
first_day[:,9]
weather_onehot.scatter_(
dim=1,
index=first_day[:,9].unsqueeze(1).long() - 1, # <1>
value=1.0)
torch.cat((bikes[:24], weather_onehot), 1)[:1]
daily_weather_onehot = torch.zeros(daily_bikes.shape[0], 4,
daily_bikes.shape[2])
daily_weather_onehot.shape
daily_weather_onehot.scatter_(
1, daily_bikes[:,9,:].long().unsqueeze(1) - 1, 1.0)
daily_weather_onehot.shape
daily_bikes = torch.cat((daily_bikes, daily_weather_onehot), dim=1)
daily_bikes[:, 9, :] = (daily_bikes[:, 9, :] - 1.0) / 3.0
temp = daily_bikes[:, 10, :]
temp_min = torch.min(temp)
temp_max = torch.max(temp)
daily_bikes[:, 10, :] = ((daily_bikes[:, 10, :] - temp_min)
/ (temp_max - temp_min))
temp = daily_bikes[:, 10, :]
daily_bikes[:, 10, :] = ((daily_bikes[:, 10, :] - torch.mean(temp))
/ torch.std(temp))
| p1ch4/4_time_series_bikes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sergiocabrales/python/blob/main/DoubleExponentialMovingAverage.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ncx40TlXSpit"
#Description: This program uses the Double Exponential Moving Average (DEMA) to determine when to buy and sell stock
#Import the libraries
import pandas as pd
import numpy as np
import pandas_datareader as web
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="Fy0JFqPPTAm-" outputId="b405f168-5cfa-4896-fa74-b4ac30ed3cdc"
# Get the stock quote
df = web.DataReader('AAPL', data_source='yahoo',start='2021-01-01', end='2021-04-28')
df
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="iGS_P4b3TE8r" outputId="ff5f06b7-cd3b-4621-cd68-909879ecb906"
#Visually show the close price
df['Close'].plot(figsize=(12.2,6.4)) #Plot the data
plt.title('Close Price for AAPL ')
plt.ylabel('USD Price ($)')
plt.xlabel('Date')
plt.show();
# + id="T4LRGELETL4L"
#Create a function to calculate the Double Exponential Moving Average (DEMA)
def DEMA(data, time_period, column):
#Calculate the Exponential Moving Average for some time_period (in days)
EMA = data[column].ewm(span=time_period, adjust=False).mean()
#Calculate the DEMA
DEMA = 2*EMA - EMA.ewm(span=time_period, adjust=False).mean()
return DEMA
# + id="IxmB_-0QTPOW"
df['DEMA_short'] = DEMA(df, 20, 'Close') #Store the short term DEMA
df['DEMA_long'] = DEMA(df, 50, 'Close') #Store the long term DEMA
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="ayKjkTkiTUCR" outputId="d2245254-4980-4ff8-89f6-93a2d410a18f"
#Plot the chart
#Create a list of columns to keep
column_list = ['DEMA_short', 'DEMA_long', 'Close']
df[column_list].plot(figsize=(12.2,6.4)) #Plot the data
plt.title('DEMA for AMZN')
plt.ylabel('USD Price ($)')
plt.xlabel('Date')
plt.show()
# + id="zTzcQq-YTczn"
def DEMA_Strategy(data):
buy_list = [] #Create a list to store the price at which to buy
sell_list = [] #Create a list to store the price at which to sell
flag = False #Create a flag to determine when the indicators cross
#Loop through the data
for i in range(0,len(data)):
#Check if the Short Term DEMA crosses above the Long Term DEMA
if data['DEMA_short'][i] > data['DEMA_long'][i] and flag == False:
buy_list.append(data['Close'][i])
sell_list.append(np.nan)
flag = True
#Check if the Short Term DEMA crosses below the Long Term DEMA
elif data['DEMA_short'][i] < data['DEMA_long'][i] and flag == True:
buy_list.append(np.nan)
sell_list.append(data['Close'][i])
flag = False
else:#Else they didn't cross
buy_list.append(np.nan)
sell_list.append(np.nan)
#Store the Buy and Sell signals in the data set
data['Buy'] = buy_list
data['Sell'] = sell_list
# + id="BGf1zEKwTeRR"
#Run the Strategy to get the buy and sell signals
DEMA_Strategy(df)
# + id="ra1aSsL5TiKZ" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="707530d3-c26d-4424-f89c-6479a822b289"
# Visually Show The Stock Buy and Sell Signals
#Create and plot the graph
plt.figure(figsize=(12.2,5.5)) #width = 12.2in, height = 4.5
plt.scatter(df.index, df['Buy'], color = 'green', label='Buy Signal', marker = '^', alpha = 1) #Plot the buy signal
plt.scatter(df.index, df['Sell'], color = 'red', label='Sell Signal', marker = 'v', alpha = 1) #Plot the sell signal
plt.plot( df['Close'], label='Close Price', alpha = 0.35)#plt.plot( X-Axis , Y-Axis, line_width, alpha_for_blending, label)
plt.plot( df['DEMA_short'], label='DEMA_short', alpha = 0.35) #plot the Short Term DEMA
plt.plot( df['DEMA_long'], label='DEMA_long', alpha = 0.35) #plot the Long Term DEMA
plt.xticks(rotation=45)#Rotate the dates 45 degrees
plt.title('Close Price History Buy / Sell Signals')
plt.xlabel('Date',fontsize=18)
plt.ylabel('Close Price USD ($)',fontsize=18)
plt.legend( loc='upper left')
plt.show()
| DoubleExponentialMovingAverage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/desabuh/elliptic_curves_cryptography_plots/blob/master/plot_for_eliptic_curves.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="5KshdH61sego"
import matplotlib.pyplot as plt
import math
import numpy as np
# %matplotlib inline
from ipywidgets import interact
from sympy import nsolve
# + [markdown] id="H6vBdrkvzbYZ"
# TOTIENT FUNCTION
#
# + id="SbcX1uXIsnOI"
def gcd(a, b):
if b==0:
return a
return gcd(b, a % b)
# + id="2Y_7nOE2spEA" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="8ecd851a-ab31-44b3-ec96-4c08fd092356"
plt.gca().spines['top'].set_visible(False)
plt.gca().spines['right'].set_visible(False)
plt.xlabel("N")
plt.ylabel("phi(N)")
plt.scatter([*range(1,2500)], [sum(gcd(n, i) == 1 for i in range(1,n)) for n in range(1, 2500)], s = 1, c='green');
plt.show()
# + [markdown] id="w-xduxm4zfJg"
# GENERAL TO WEISTRASS FORM
#
# + colab={"base_uri": "https://localhost:8080/", "height": 425, "referenced_widgets": ["6bcb9d38763642e3b480d547a434192d", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "9e4d794eb82e4aa382c9fab2612fa9ed", "<KEY>", "0ab00e52009942d88675b7de3f3a25f9", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "fc6d5d171628405da738581f83a8ecab", "385530ab40074c949e46f1d31a767639", "c37a997de1ac47d4a66a198b668394cb", "e594957bc004448c99f4bad10bc5c34f", "2b9efaa274ef4a49828b7b5ad77a2ce8", "a10260f6243546e4a5e5efc7466c3ef6"]} id="V9EDDhjcB8FO" outputId="ee24d046-f1a6-456f-eb62-06cfafb25722"
@interact(a = (-10,10,0.1), b=(-10,10,0.1), c=(-10,10,0.1), d=(-10,10,0.1), e=(-10,10,0.1))
def ell_curve(a, b, c, d, e):
mx2, mx1 = np.ogrid[-10:10:0.1,-15:15:0.1]
def evaluate_general(x,y):
return np.power(y,2) + a*x*y + b*y - np.power(x, 3) - c * np.power(x,2) - d*x - e
def transform_coord(x,y):
return x - ((a**2 + 4*c) / 12), y - (a / 2)*x + ((a**3 + 4*a*c - 12 * b) / 24)
def evaluate_normal(x,y):
x, y = transform_coord(x,y)
return np.power(y,2) - np.power(x,3) - d*x - e
plt.contour(mx1.ravel(), mx2.ravel(), evaluate_general(mx1, mx2), [0], colors="blue")
plt.contour(mx1.ravel(), mx2.ravel(), evaluate_normal(mx1, mx2), [0], colors="red")
plt.show()
# + [markdown] id="Sx_x8EzfoA7Y"
# FINITE CURVE
# + id="SIsyW0TlZq8Y"
def display_finite_curve(a, b, N):
def is_point(x, y, a, b, N):
return (y**2) % N == (x**3+ a*x + b) % N
points = [(x,y) for x in range(N) for y in range(N) if is_point(x,y,a,b,N)]
plt.text(-5,-5,s = "p = {}\n a = {}\n b= {}".format(N,a,b),c = "black",bbox={'facecolor': 'green', 'alpha': 0.5})
plt.scatter(list(zip(*points))[0], list(zip(*points))[1], s=10)
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="PaDfw_3oaAx-" outputId="b4b3d8cd-ce95-40c0-e5bc-1fd38d0df4be"
display_finite_curve(1, -1, 39)
| plot_for_eliptic_curves.ipynb |
# ---
# layout: post
# title: "영상입력 이진분류 모델 레시피"
# author: 김태영
# date: 2017-08-18 02:00:00
# categories: Lecture
# comments: true
# image: http://tykimos.github.io/warehouse/2017-8-18-Image_Input_Binary_Classification_Model_Recipe_title_m.png
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# 영상을 입력해서 이진분류할 수 있는 모델들에 대해서 알아보겠습니다. 숫자 손글씨 데이터셋인 MNIST을 이용하여 홀수/짝수를 구분을 위한 데이터셋을 생성해보고, 다층퍼셉트론 및 컨볼루션 신경망 모델을 구성하고 학습 시켜보겠습니다. 이 모델은 임의의 영상으로부터 A와 B를 구분하는 문제나 양성과 음성을 구분하는 문제를 풀 수 있습니다. 아래 문제들에 활용 용 기대해봅니다.
# * 입력된 얼굴 사진에 남자인지 여자인지 구분
# * 촬영된 부품 사진이 정상인지 불량인지 구분
# * 의료영상을 보고 질병유무 판독
# ---
# ### 데이터셋 준비
#
# 케라스 함수에서 제공하는 숫자 손글씨 데이터셋인 MNIST을 이용하겠습니다. 초기 라벨값은 0에서 9까지 정수로 지정되어 있습니다. 데이터 정규화를 위해서 255.0으로 나누었습니다. 아래는 다층퍼셉트론 신경망 모델에 입력하기 위해 데이터셋 생성하는 코드입니다.
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(60000, width*height).astype('float32') / 255.0
x_test = x_test.reshape(10000, width*height).astype('float32') / 255.0
# 아래는 컨볼루션 신경망 모델에 입력하기 위해 데이터셋 생성하는 코드입니다. 샘플수, 너비, 높이, 채널수로 총 4차원 배열로 구성합니다.
x_train = x_train.reshape(60000, width, height, 1).astype('float32') / 255.0
x_test = x_test.reshape(10000, width, height, 1).astype('float32') / 255.0
# 불러온 훈련셋을 다시 훈련셋 50,000개와 검증셋 10,000개로 나누었습니다.
x_val = x_train[50000:]
y_val = y_train[50000:]
x_train = x_train[:50000]
y_train = y_train[:50000]
# 라벨값은 다중클래스 분류로 0에서 9까지 지정되어 있으나 이것을 홀수/짝수로 바꾸어서 이진분류 라벨로 지정하겠습니다. '1'은 홀수를 의미하고, '0'은 짝수를 의미합니다.
y_train = y_train % 2
y_val = y_val % 2
y_test = y_test % 2
# 만든 데이터셋 일부를 가시화 해보겠습니다.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
plt_row = 5
plt_col = 5
plt.rcParams["figure.figsize"] = (10,10)
f, axarr = plt.subplots(plt_row, plt_col)
for i in range(plt_row*plt_col):
sub_plt = axarr[i/plt_row, i%plt_col]
sub_plt.axis('off')
sub_plt.imshow(x_test[i].reshape(width, height))
sub_plt_title = 'R: '
if y_test[i] :
sub_plt_title += 'odd '
else:
sub_plt_title += 'even '
sub_plt.set_title(sub_plt_title)
plt.show()
# -
# 
# ---
# ### 레이어 준비
#
# 본 장에서 새롭게 소개되는 블록은 'Dropout'입니다.
#
# |블록|이름|설명|
# |:-:|:-:|:-|
# ||Dropout|과적합을 방지하기 위해서 학습 시에 지정된 비율만큼 임의의 입력 뉴런(1차원)을 제외시킵니다.|
# ||Dropout|과적합을 방지하기 위해서 학습 시에 지정된 비율만큼 임의의 입력 뉴런(2차원)을 제외시킵니다.|
# ---
# ### 모델 준비
#
# 영상을 입력하여 이진분류를 하기 위해 `다층퍼셉트론 신경망 모델`, `컨볼루션 신경망 모델`, `깊은 컨볼루션 신경망 모델`을 준비했습니다.
#
# #### 다층퍼셉트론 신경망 모델
#
# model = Sequential()
# model.add(Dense(256, input_dim=width*height, activation='relu'))
# model.add(Dense(256, activation='relu'))
# model.add(Dense(256, activation='relu'))
# model.add(Dense(1, activation='sigmoid'))
#
#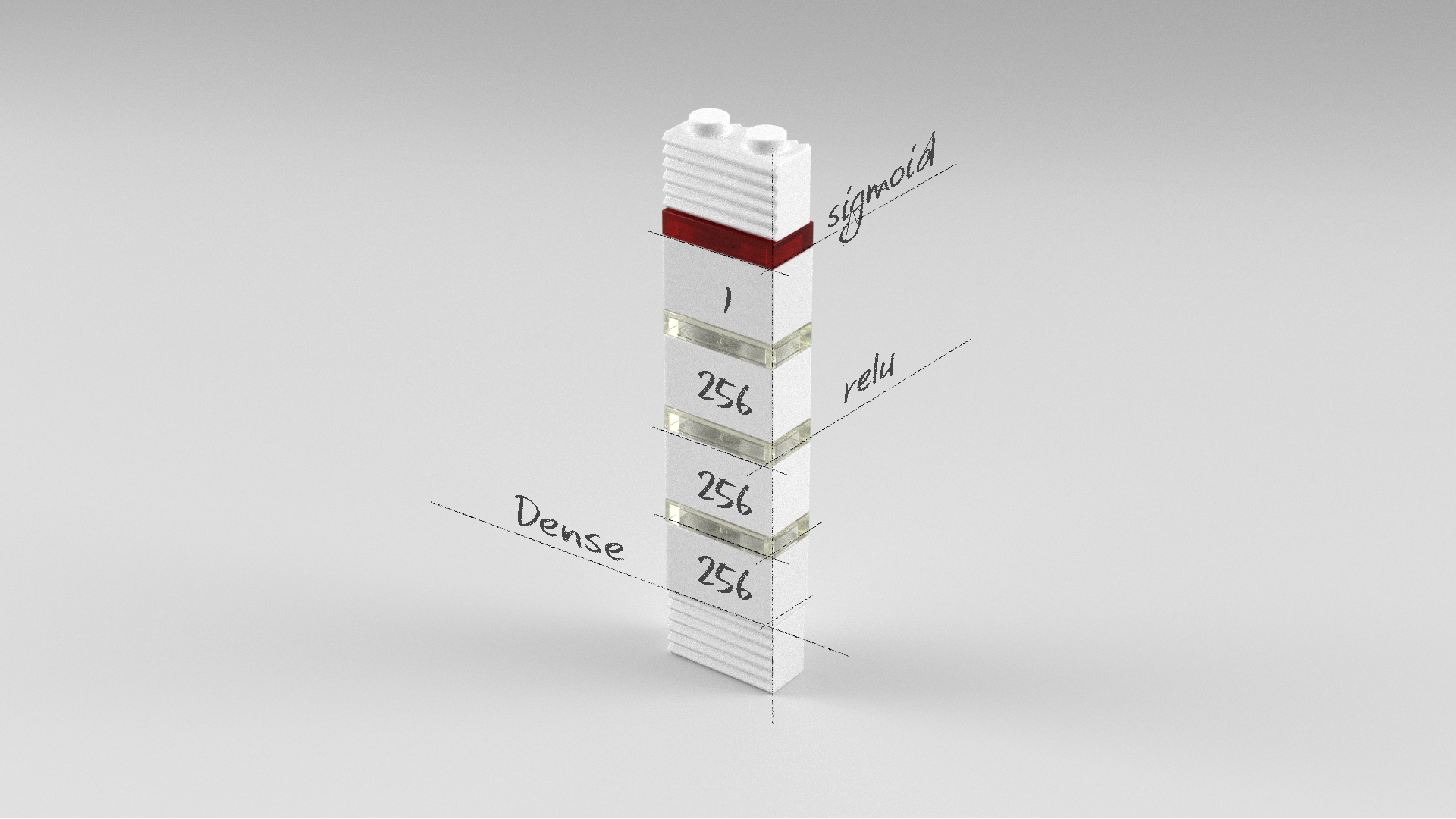 
#
# #### 컨볼루션 신경망 모델
#
# model = Sequential()
# model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(width, height, 1)))
# model.add(MaxPooling2D(pool_size=(2, 2)))
# model.add(Conv2D(32, (3, 3), activation='relu'))
# model.add(MaxPooling2D(pool_size=(2, 2)))
# model.add(Flatten())
# model.add(Dense(256, activation='relu'))
# model.add(Dense(1, activation='sigmoid'))
#
#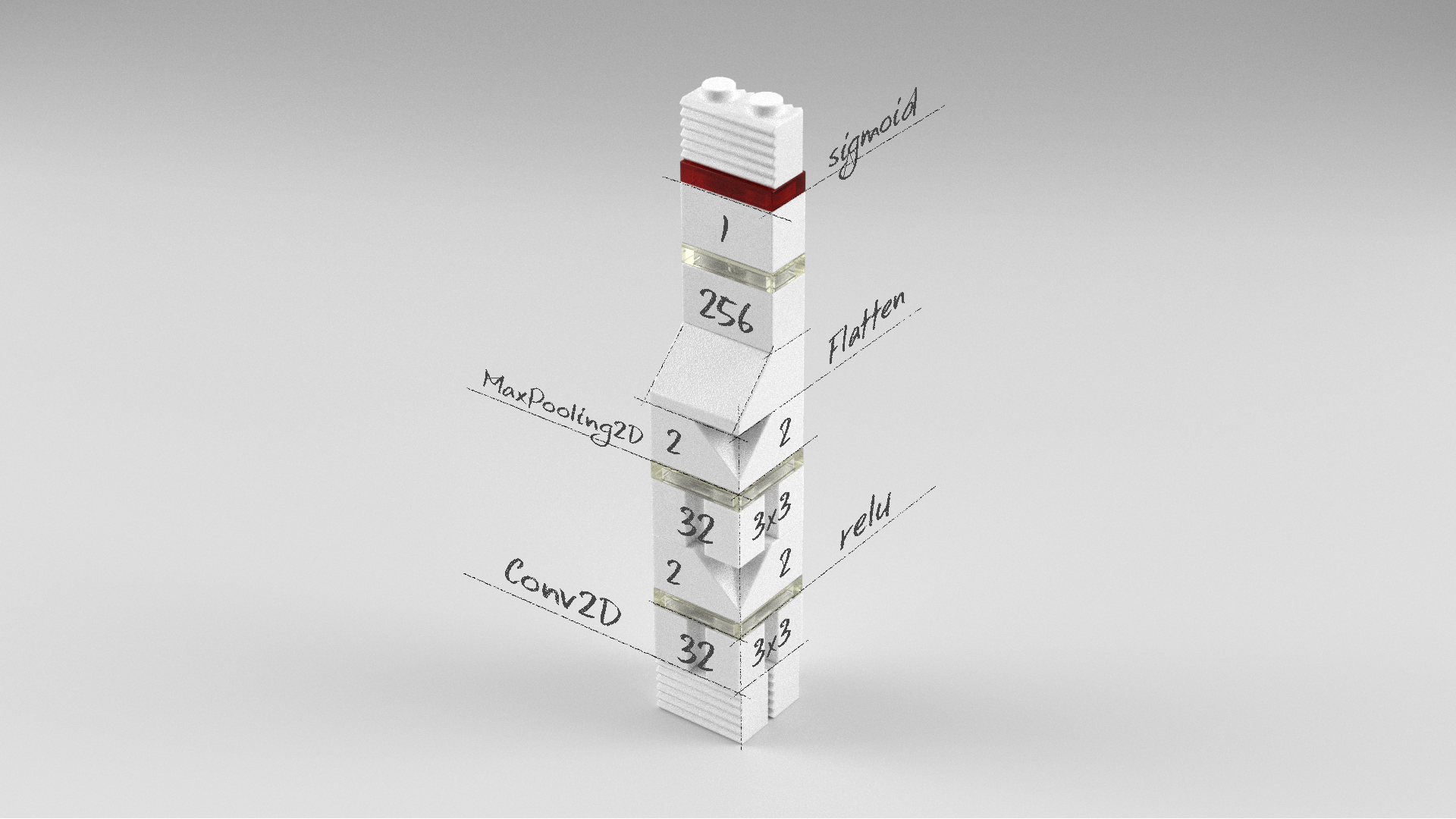 
#
# #### 깊은 컨볼루션 신경망 모델
#
# model = Sequential()
# model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(width, height, 1)))
# model.add(Conv2D(32, (3, 3), activation='relu'))
# model.add(MaxPooling2D(pool_size=(2, 2)))
# model.add(Dropout(0.25))
# model.add(Conv2D(64, (3, 3), activation='relu'))
# model.add(Conv2D(64, (3, 3), activation='relu'))
# model.add(MaxPooling2D(pool_size=(2, 2)))
# model.add(Dropout(0.25))
# model.add(Flatten())
# model.add(Dense(256, activation='relu'))
# model.add(Dropout(0.5))
# model.add(Dense(1, activation='sigmoid'))
#
#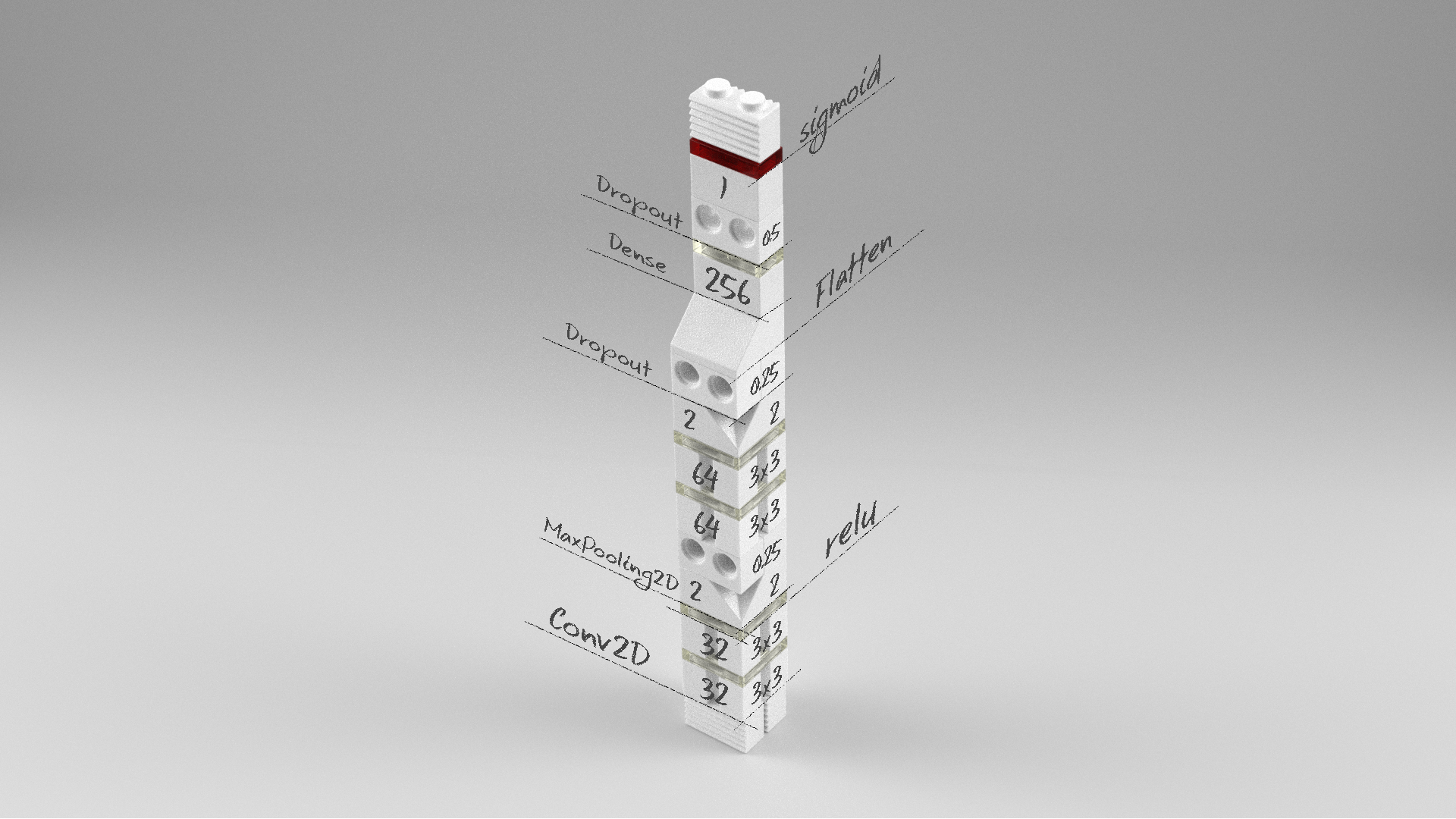 
# ---
# ### 전체 소스
#
# 앞서 살펴본 `다층퍼셉트론 신경망 모델`, `컨볼루션 신경망 모델`, `깊은 컨볼루션 신경망 모델`의 전체 소스는 다음과 같습니다.
# #### 다중퍼셉트론 신경망 모델
# +
# 0. 사용할 패키지 불러오기
from keras.utils import np_utils
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Activation
width = 28
height = 28
# 1. 데이터셋 생성하기
# 훈련셋과 시험셋 불러오기
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(60000, width*height).astype('float32') / 255.0
x_test = x_test.reshape(10000, width*height).astype('float32') / 255.0
# 훈련셋과 검증셋 분리
x_val = x_train[50000:]
y_val = y_train[50000:]
x_train = x_train[:50000]
y_train = y_train[:50000]
# 데이터셋 전처리 : 홀수는 1, 짝수는 0으로 변환
y_train = y_train % 2
y_val = y_val % 2
y_test = y_test % 2
# 2. 모델 구성하기
model = Sequential()
model.add(Dense(256, input_dim=width*height, activation='relu'))
model.add(Dense(256, activation='relu'))
model.add(Dense(256, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
# 3. 모델 학습과정 설정하기
model.compile(loss='binary_crossentropy', optimizer='sgd', metrics=['accuracy'])
# 4. 모델 학습시키기
hist = model.fit(x_train, y_train, epochs=30, batch_size=32, validation_data=(x_val, y_val))
# 5. 학습과정 살펴보기
# %matplotlib inline
import matplotlib.pyplot as plt
fig, loss_ax = plt.subplots()
acc_ax = loss_ax.twinx()
loss_ax.plot(hist.history['loss'], 'y', label='train loss')
loss_ax.plot(hist.history['val_loss'], 'r', label='val loss')
loss_ax.set_ylim([0.0, 0.5])
acc_ax.plot(hist.history['acc'], 'b', label='train acc')
acc_ax.plot(hist.history['val_acc'], 'g', label='val acc')
acc_ax.set_ylim([0.8, 1.0])
loss_ax.set_xlabel('epoch')
loss_ax.set_ylabel('loss')
acc_ax.set_ylabel('accuray')
loss_ax.legend(loc='upper left')
acc_ax.legend(loc='lower left')
plt.show()
# 6. 모델 평가하기
loss_and_metrics = model.evaluate(x_test, y_test, batch_size=32)
print('## evaluation loss and_metrics ##')
print(loss_and_metrics)
# 7. 모델 사용하기
yhat_test = model.predict(x_test, batch_size=32)
# %matplotlib inline
import matplotlib.pyplot as plt
plt_row = 5
plt_col = 5
plt.rcParams["figure.figsize"] = (10,10)
f, axarr = plt.subplots(plt_row, plt_col)
for i in range(plt_row*plt_col):
sub_plt = axarr[i/plt_row, i%plt_col]
sub_plt.axis('off')
sub_plt.imshow(x_test[i].reshape(width, height))
sub_plt_title = 'R: '
if y_test[i] :
sub_plt_title += 'odd '
else:
sub_plt_title += 'even '
sub_plt_title += 'P: '
if yhat_test[i] >= 0.5 :
sub_plt_title += 'odd '
else:
sub_plt_title += 'even '
sub_plt.set_title(sub_plt_title)
plt.show()
# -
# 시험셋으로 예측한 결과 일부를 비교해봤습니다. 25개 샘플 중 9번째 샘플을 제외하고는 모두 맞췄습니다.
#
# 
# #### 컨볼루션 신경망 모델
# +
# 0. 사용할 패키지 불러오기
from keras.utils import np_utils
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.layers import Conv2D, MaxPooling2D, Flatten
width = 28
height = 28
# 1. 데이터셋 생성하기
# 훈련셋과 시험셋 불러오기
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(60000, width, height, 1).astype('float32') / 255.0
x_test = x_test.reshape(10000, width, height, 1).astype('float32') / 255.0
# 훈련셋과 검증셋 분리
x_val = x_train[50000:]
y_val = y_train[50000:]
x_train = x_train[:50000]
y_train = y_train[:50000]
# 데이터셋 전처리 : 홀수는 1, 짝수는 0으로 변환
y_train = y_train % 2
y_val = y_val % 2
y_test = y_test % 2
# 2. 모델 구성하기
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(width, height, 1)))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(32, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
# 3. 모델 학습과정 설정하기
model.compile(loss='binary_crossentropy', optimizer='sgd', metrics=['accuracy'])
# 4. 모델 학습시키기
hist = model.fit(x_train, y_train, epochs=30, batch_size=32, validation_data=(x_val, y_val))
# 5. 학습과정 살펴보기
# %matplotlib inline
import matplotlib.pyplot as plt
fig, loss_ax = plt.subplots()
acc_ax = loss_ax.twinx()
loss_ax.plot(hist.history['loss'], 'y', label='train loss')
loss_ax.plot(hist.history['val_loss'], 'r', label='val loss')
loss_ax.set_ylim([0.0, 0.5])
acc_ax.plot(hist.history['acc'], 'b', label='train acc')
acc_ax.plot(hist.history['val_acc'], 'g', label='val acc')
acc_ax.set_ylim([0.8, 1.0])
loss_ax.set_xlabel('epoch')
loss_ax.set_ylabel('loss')
acc_ax.set_ylabel('accuray')
loss_ax.legend(loc='upper left')
acc_ax.legend(loc='lower left')
plt.show()
# 6. 모델 평가하기
loss_and_metrics = model.evaluate(x_test, y_test, batch_size=32)
print('## evaluation loss and_metrics ##')
print(loss_and_metrics)
# 7. 모델 사용하기
yhat_test = model.predict(x_test, batch_size=32)
# %matplotlib inline
import matplotlib.pyplot as plt
plt_row = 5
plt_col = 5
plt.rcParams["figure.figsize"] = (10,10)
f, axarr = plt.subplots(plt_row, plt_col)
for i in range(plt_row*plt_col):
sub_plt = axarr[i/plt_row, i%plt_col]
sub_plt.axis('off')
sub_plt.imshow(x_test[i].reshape(width, height))
sub_plt_title = 'R: '
if y_test[i] :
sub_plt_title += 'odd '
else:
sub_plt_title += 'even '
sub_plt_title += 'P: '
if yhat_test[i] >= 0.5 :
sub_plt_title += 'odd '
else:
sub_plt_title += 'even '
sub_plt.set_title(sub_plt_title)
plt.show()
# -
# 시험셋으로 예측한 결과 일부를 비교해봤습니다. '다층퍼셉트론 신경망 모델'과 마찬가지로 25개 샘플 중 9번째 샘플을 제외하고는 모두 맞췄습니다.
#
# 
# #### 깊은 컨볼루션 신경망 모델
# +
# 0. 사용할 패키지 불러오기
from keras.utils import np_utils
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.layers import Conv2D, MaxPooling2D, Flatten
from keras.layers import Dropout
width = 28
height = 28
# 1. 데이터셋 생성하기
# 훈련셋과 시험셋 불러오기
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(60000, width, height, 1).astype('float32') / 255.0
x_test = x_test.reshape(10000, width, height, 1).astype('float32') / 255.0
# 훈련셋과 검증셋 분리
x_val = x_train[50000:]
y_val = y_train[50000:]
x_train = x_train[:50000]
y_train = y_train[:50000]
# 데이터셋 전처리 : 홀수는 1, 짝수는 0으로 변환
y_train = y_train % 2
y_val = y_val % 2
y_test = y_test % 2
# 2. 모델 구성하기
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(width, height, 1)))
model.add(Conv2D(32, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(1, activation='sigmoid'))
# 3. 모델 학습과정 설정하기
model.compile(loss='binary_crossentropy', optimizer='sgd', metrics=['accuracy'])
# 4. 모델 학습시키기
hist = model.fit(x_train, y_train, epochs=30, batch_size=32, validation_data=(x_val, y_val))
# 5. 학습과정 살펴보기
# %matplotlib inline
import matplotlib.pyplot as plt
fig, loss_ax = plt.subplots()
acc_ax = loss_ax.twinx()
loss_ax.plot(hist.history['loss'], 'y', label='train loss')
loss_ax.plot(hist.history['val_loss'], 'r', label='val loss')
loss_ax.set_ylim([0.0, 0.5])
acc_ax.plot(hist.history['acc'], 'b', label='train acc')
acc_ax.plot(hist.history['val_acc'], 'g', label='val acc')
acc_ax.set_ylim([0.8, 1.0])
loss_ax.set_xlabel('epoch')
loss_ax.set_ylabel('loss')
acc_ax.set_ylabel('accuray')
loss_ax.legend(loc='upper left')
acc_ax.legend(loc='lower left')
plt.show()
# 6. 모델 평가하기
loss_and_metrics = model.evaluate(x_test, y_test, batch_size=32)
print('## evaluation loss and_metrics ##')
print(loss_and_metrics)
# 7. 모델 사용하기
yhat_test = model.predict(x_test, batch_size=32)
# %matplotlib inline
import matplotlib.pyplot as plt
plt_row = 5
plt_col = 5
plt.rcParams["figure.figsize"] = (10,10)
f, axarr = plt.subplots(plt_row, plt_col)
for i in range(plt_row*plt_col):
sub_plt = axarr[i/plt_row, i%plt_col]
sub_plt.axis('off')
sub_plt.imshow(x_test[i].reshape(width, height))
sub_plt_title = 'R: '
if y_test[i] :
sub_plt_title += 'odd '
else:
sub_plt_title += 'even '
sub_plt_title += 'P: '
if yhat_test[i] >= 0.5 :
sub_plt_title += 'odd '
else:
sub_plt_title += 'even '
sub_plt.set_title(sub_plt_title)
plt.show()
# -
# 시험셋으로 예측한 결과 일부를 비교해봤습니다. 이전 모델에선 틀렸던 9번째 샘플도 맞췄습니다.
#
# 
# ---
#
# ### 학습결과 비교
#
# 다층퍼셉트론 신경망 모델은 훈련정확도는 검증 손실값은 높아지고 있어 과적합이 발생하였습니다. 컨볼루션 신경망 모델은 다층퍼셉트론 신경망 모델에 비해 높은 성능을 보이고 있습니다. 깊은 컨볼루션 신경망 모델은 드랍아웃(Dropout) 레이어 덕분에 과적합이 발생하지 않고 검증 손실값이 지속적으로 떨어지고 있음을 확인할 수 있습니다.
#
# |다층퍼셉트론 신경망 모델|컨볼루션 신경망 모델|깊은 컨볼루션 신경망 모델|
# |:-:|:-:|:-:|
# |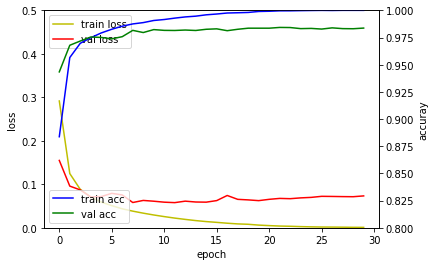|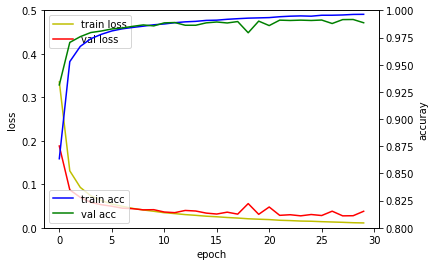|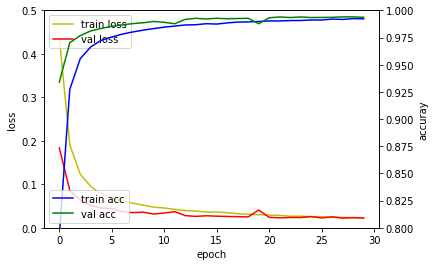
# ---
#
# ### 요약
#
# 영상를 입력하여 이진분류하는 다층퍼셉트론 신경망 모델, 컨볼루션 신경망 모델, 깊은 컨볼루션 신경망 모델을 살펴보고 그 성능을 확인 해봤습니다.
#
#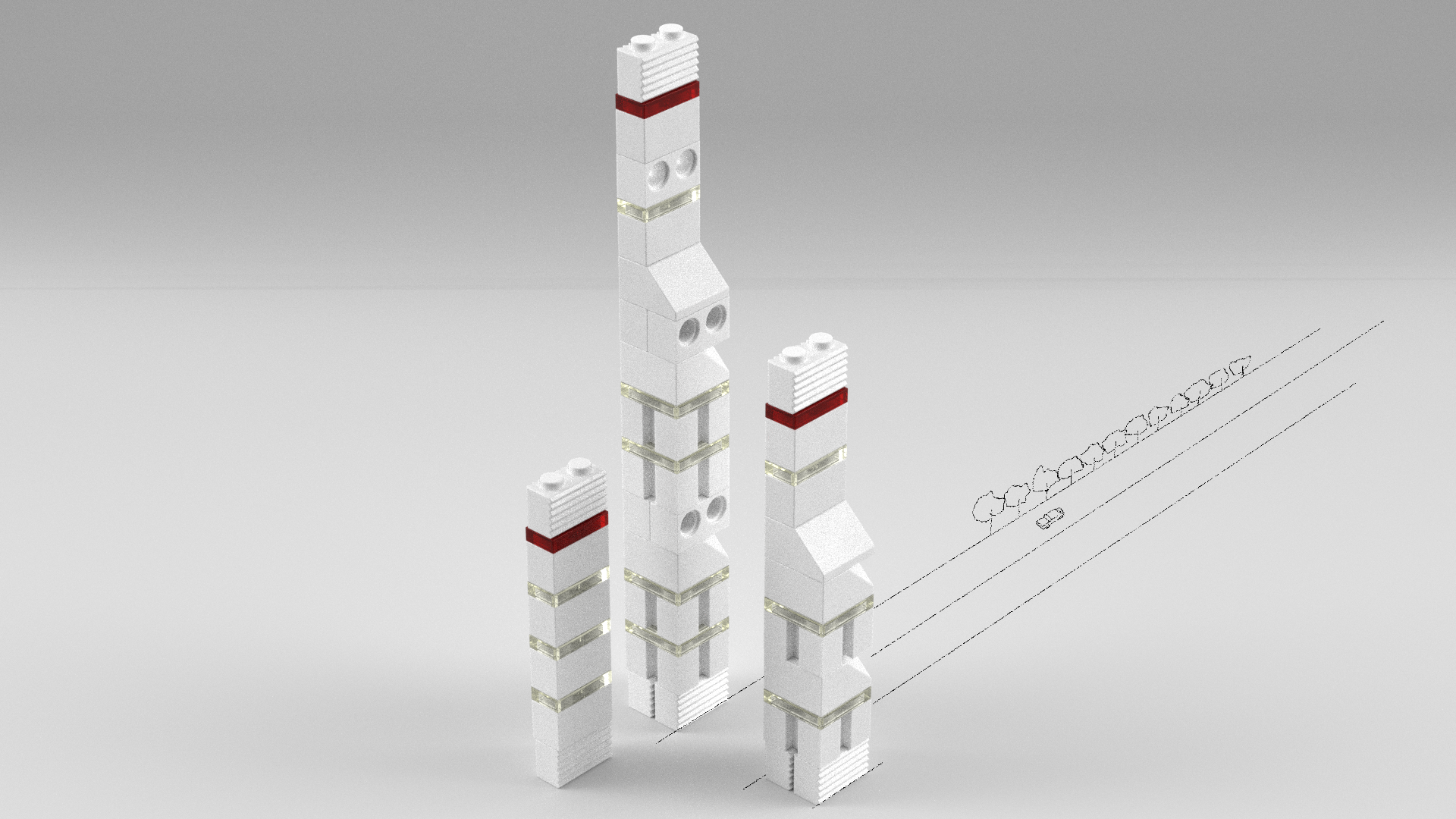 
# ---
#
# ### 같이 보기
#
# * [강좌 목차](https://tykimos.github.io/lecture/)
# * 이전 : [영상입력 수치예측 모델 레시피](https://tykimos.github.io/2017/08/20/Image_Input_Numerical_Prediction_Model_Recipe/)
# * 다음 : [영상입력 다중클래스분류 모델 레시피](https://tykimos.github.io/2017/08/18/Image_Input_Multiclass_Classification_Model_Recipe/)
| _writing/2017-8-18-Image_Input_Binary_Classification_Model_Recipe.ipynb |