code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="9aSTkPJcoCGf"
# 
# + [markdown] id="Ns7qmq7moEno"
# # **PySpark Tutorial-8 Custom Annotators UDF and Light Pipelines**
# + [markdown] id="A4XVzhsZU_Yd"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/PySpark/8.PySpark_CustomAnnotators_UDF_and_Lightpipelines.ipynb)
# + [markdown] id="mo2CP3QgoNdS"
# In this notebook, some special Spark NLP annotators have been performed.
#
#
#
# + [markdown] id="n8dM5YYFoLrg"
# ### Install PySpark
# + id="YiQLvv4lnuZy"
# install PySpark
# ! pip install -q pyspark==3.2.0 spark-nlp
# + [markdown] id="ymaOkjF5oUgz"
# ### Initializing Spark
# + colab={"base_uri": "https://localhost:8080/", "height": 254} id="4pNShfO7oPDo" executionInfo={"status": "ok", "timestamp": 1645430762418, "user_tz": -180, "elapsed": 27227, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="82d0bd86-0729-472c-8dd1-88310fd38bbc"
import sparknlp
spark = sparknlp.start(spark32=True)
# params =>> gpu=False, spark23=False (start with spark 2.3)
print("Spark NLP version", sparknlp.version())
print("Apache Spark version:", spark.version)
spark
# + id="UqWgoDS0oY7k"
# DO NOT FORGET WHEN YOU'RE DONE => spark.stop()
# + id="-cdjTAnKrUdI"
from sparknlp.base import *
import pandas as pd
from sparknlp.functions import *
from pyspark.sql.functions import col
from pyspark.sql.types import ArrayType, StringType
from pyspark.ml.param.shared import HasInputCol, HasOutputCol
from pyspark.ml.util import DefaultParamsReadable, DefaultParamsWritable
import pyspark.sql.functions as F
import pyspark.sql.types as T
from pyspark.sql import Row
# + [markdown] id="1u8dhq8C0paw"
# # Annotators and Transformer Concepts
# + [markdown] id="YSTTBzLux43n"
# In Spark NLP, all Annotators are either Estimators or Transformers as we see in Spark ML. An Estimator in Spark ML is an algorithm which can be fit on a DataFrame to produce a Transformer. E.g., a learning algorithm is an Estimator which trains on a DataFrame and produces a model. A Transformer is an algorithm which can transform one DataFrame into another DataFrame. E.g., an ML model is a Transformer that transforms a DataFrame with features into a DataFrame with predictions. In Spark NLP, there are two types of annotators: AnnotatorApproach and AnnotatorModel AnnotatorApproach extends Estimators from Spark ML, which are meant to be trained through fit(), and AnnotatorModel extends Transformers which are meant to transform data frames through transform(). Some of Spark NLP annotators have a Model suffix and some do not. The model suffix is explicitly stated when the annotator is the result of a training process. Some annotators, such as Tokenizer are transformers but do not contain the suffix Model since they are not trained, annotators. Model annotators have a pre-trained() on its static object, to retrieve the public pre-trained version of a model. Long story short, if it trains on a DataFrame and produces a model, it’s an AnnotatorApproach; and if it transforms one DataFrame into another DataFrame through some models, it’s an AnnotatorModel (e.g. WordEmbeddingsModel) and it doesn’t take Model suffix if it doesn’t rely on a pre-trained annotator while transforming a DataFrame (e.g. Tokenizer).
# + id="b6Txfllfx5Oo"
# !wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jupyter/annotation/english/spark-nlp-basics/sample-sentences-en.txt
# + colab={"base_uri": "https://localhost:8080/"} id="PJsfwrxbyRGN" executionInfo={"status": "ok", "timestamp": 1645430784761, "user_tz": -180, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="d08a6ce3-2bbb-445c-a29a-ad5b84c7be73"
with open('./sample-sentences-en.txt') as f:
print (f.read())
# + colab={"base_uri": "https://localhost:8080/"} id="ftQtm_2SySi2" executionInfo={"status": "ok", "timestamp": 1645430792844, "user_tz": -180, "elapsed": 6333, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="7a3d311e-2870-46f3-95d1-da66f974f662"
spark_df = spark.read.text('./sample-sentences-en.txt').toDF('text')
spark_df.show(truncate=False)
# + [markdown] id="VlSKSOHai53C"
# ## Spark NLP Annotators
# + [markdown] id="GstOz1mpyrvk"
# ### Document Assembler
#
# To get through the process in Spark NLP, we need to get raw data transformed into Document type at first.
#
# DocumentAssembler() is a special transformer that does this for us; it creates the first annotation of type Document which may be used by annotators down the road.
# + colab={"base_uri": "https://localhost:8080/"} id="EAW6EBHlybXV" executionInfo={"status": "ok", "timestamp": 1645430798645, "user_tz": -180, "elapsed": 1830, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="fb410a2a-1835-4c17-8269-6e5effaf77dc"
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")\
.setCleanupMode("shrink")
doc_df = documentAssembler.transform(spark_df)
doc_df.show(truncate=30)
# + colab={"base_uri": "https://localhost:8080/"} id="o2N_Ay0gyt0H" executionInfo={"status": "ok", "timestamp": 1645430798922, "user_tz": -180, "elapsed": 285, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="b346e59e-c7b8-4c1c-a18d-69fa45edab66"
doc_df.printSchema()
# + colab={"base_uri": "https://localhost:8080/"} id="_SG41Iery73v" executionInfo={"status": "ok", "timestamp": 1645430801370, "user_tz": -180, "elapsed": 689, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="51580356-e15e-414a-b37d-33f08b5fdce0"
doc_df.select('document.result','document.begin','document.end').show(truncate=False)
# + colab={"base_uri": "https://localhost:8080/"} id="CvUVfZE3y_fP" executionInfo={"status": "ok", "timestamp": 1645430802732, "user_tz": -180, "elapsed": 394, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="08c59f9a-f863-4d66-c3f5-8196573b4c80"
doc_df.select("document.result").take(1)
# + [markdown] id="Jric5ouozk-C"
# ### Sentence Detector
# Finds sentence bounds in raw text.
# `setCustomBounds(string)`: Custom sentence separator text e.g. `["\n"]`
# + id="WhKAIDWlzl_J"
from sparknlp.annotator import *
# we feed the document column coming from Document Assembler
sentenceDetector = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentences')
# + colab={"base_uri": "https://localhost:8080/"} id="gMb0j2NyzvCX" executionInfo={"status": "ok", "timestamp": 1645430806069, "user_tz": -180, "elapsed": 1029, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="03b876d0-fefa-450e-dd75-31cc1e839a2d"
sent_df = sentenceDetector.transform(doc_df)
sent_df.show(truncate=False)
# + colab={"base_uri": "https://localhost:8080/"} id="3Adh-Mt70JYf" executionInfo={"status": "ok", "timestamp": 1645430806784, "user_tz": -180, "elapsed": 362, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="c0a5ce87-b014-4adc-803a-dfed354c1045"
sent_df.select('sentences.result').take(5)
# + [markdown] id="qgkepVjctCse"
# ### Tokenizer
#
# Identifies tokens with tokenization open standards. It is an **Annotator Approach, so it requires .fit()**.
#
# A few rules will help customizing it if defaults do not fit user needs.
#
# setExceptions(StringArray): List of tokens to not alter at all. Allows composite tokens like two worded tokens that the user may not want to split.
# + id="cNeLmh5EtPfD"
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
# + id="j68Vzej2tdsZ"
text = '<NAME> (Spiderman) is a nice guy and lives in New York but has no e-mail!'
spark_df = spark.createDataFrame([[text]]).toDF("text")
# + colab={"base_uri": "https://localhost:8080/"} id="YnUF3LCHtdpu" executionInfo={"status": "ok", "timestamp": 1645430813366, "user_tz": -180, "elapsed": 1545, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="d27ce3a9-4a3c-4567-85a0-fd943f56a31e"
doc_df = documentAssembler.transform(spark_df)
token_df = tokenizer.fit(doc_df).transform(doc_df)
token_df.show(truncate=100)
# + colab={"base_uri": "https://localhost:8080/"} id="72BG3XpAtdll" executionInfo={"status": "ok", "timestamp": 1645430813755, "user_tz": -180, "elapsed": 392, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="c8a7102f-cdc9-<PASSWORD>"
token_df.select('token.result').take(1)
# + [markdown] id="A9RNPnggt2me"
# ### Perceptron Model
#
# POS - Part of speech tags
#
# Averaged Perceptron model to tag words part-of-speech. Sets a POS tag to each word within a sentence.
#
# This is the instantiated model of the PerceptronApproach. For training your own model, please see the documentation of that class.
# + colab={"base_uri": "https://localhost:8080/"} id="y4mP6WcWtdi-" executionInfo={"status": "ok", "timestamp": 1645430834642, "user_tz": -180, "elapsed": 19454, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="54cd1587-1d07-4b30-d496-83e7fd6b2c14"
pos = PerceptronModel.pretrained()\
.setInputCols(['document', 'token'])\
.setOutputCol('pos')
# + [markdown] id="bzhQm1kyq33D"
# ## Custom Annotator
# + [markdown] id="C7jz3ViSrFyo"
# ### SentenceChecking
# + id="XEsjBGUfrIaP"
class SentenceChecking(
Transformer, HasInputCol, HasOutputCol,
DefaultParamsReadable, DefaultParamsWritable):
output_annotation_type = "document"
def __init__(self,f,output_annotation_type="document"):
super(SentenceChecking, self).__init__()
self.f = f
def setInputCol(self, value):
"""
Sets the value of :py:attr:`inputCol`.
"""
return self._set(inputCol=value)
def setOutputCol(self, value):
"""
Sets the value of :py:attr:`outputCol`.
"""
return self._set(outputCol=value)
def _transform(self, dataset):
t = Annotation.arrayType()
out_col = self.getOutputCol()
in_col = dataset[self.getInputCol()]
return dataset.withColumn(out_col, map_annotations(self.f, t)(in_col).alias(out_col, metadata={
'annotatorType': self.output_annotation_type}))
# + id="lcg6td14rIXs"
def checking_sentences(annotations):
anns = []
for a in annotations:
result = a.result + " - CHECKED SENTENCE"
anns.append(sparknlp.annotation.Annotation(a.annotator_type, a.begin, a.begin + len(result), result, a.metadata, a.embeddings))
return anns
# + [markdown] id="JnKkOz1ju-JJ"
# ## Creating Pipeline with Custom Annotator
# + id="nzvTSZldrIU8"
document_assembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentence_detector = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentences')
sentence_checker = SentenceChecking(f=checking_sentences, output_annotation_type="document")\
.setInputCol("sentences")\
.setOutputCol("checked_sentences")
tokenizer = Tokenizer()\
.setInputCols(["checked_sentences"])\
.setOutputCol("tokens")
pipeline = Pipeline(stages=[document_assembler,
sentence_detector,
sentence_checker,
tokenizer
])
# + id="kemBz_wMrISa"
test_string = "This is a sample text with multiple sentences. It aims to show our custom annotator problem."
test_data = spark.createDataFrame([[test_string]]).toDF("text")
# + id="IpQd6ncasP9H" executionInfo={"status": "ok", "timestamp": 1645430915397, "user_tz": -180, "elapsed": 393, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} colab={"base_uri": "https://localhost:8080/"} outputId="c143b105-0b8a-4a7f-cd4f-6d923608e38d"
# %%time
fitted_pipeline = pipeline.fit(test_data)
spark_results = fitted_pipeline.transform(test_data)
# + colab={"base_uri": "https://localhost:8080/"} id="A9OxPLwnU9HC" executionInfo={"status": "ok", "timestamp": 1645430984215, "user_tz": -180, "elapsed": 702, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="74608d58-c574-4c0c-91b4-7004e40b9540"
# %%time
spark_results.show(truncate=False)
# + colab={"base_uri": "https://localhost:8080/"} id="uX8_5WTHsP6q" executionInfo={"status": "ok", "timestamp": 1645430929364, "user_tz": -180, "elapsed": 1998, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="380e9c93-fd59-4be1-daab-6870dd8f28fb"
# %%time
spark_results.select("checked_sentences").show(truncate=False)
# + colab={"base_uri": "https://localhost:8080/"} id="4YIExvdWsP3_" executionInfo={"status": "ok", "timestamp": 1645431006646, "user_tz": -180, "elapsed": 687, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="36a81271-2f68-4c58-873d-25c78a6df3bf"
spark_results.select("tokens.result").show(truncate=False)
# + [markdown] id="7AiJebnhWRkG"
# ## LightPipeline
# + [markdown] id="T9jIjMWAXVv0"
# Spark NLP LightPipelines are Spark ML pipelines converted into a single machine but the multi-threaded task, becoming more than **10x times faster** for smaller amounts of data (small is relative, but 50k sentences are roughly a good maximum). To use them, we simply plug in a trained (fitted) pipeline and then annotate a plain text. We don't even need to convert the input text to DataFrame in order to feed it into a pipeline that's accepting DataFrame as an input in the first place. This feature would be quite useful when it comes to getting a prediction for a few lines of text from a trained ML model. Here is the medium post [Spark NLP 101: LightPipeline](https://medium.com/spark-nlp/spark-nlp-101-lightpipeline-a544e93f20f1)
# + colab={"base_uri": "https://localhost:8080/"} id="9DaKaytMWb7F" executionInfo={"status": "ok", "timestamp": 1645431977456, "user_tz": -180, "elapsed": 4041, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="c30325f4-aad3-483e-9d8d-48fd310df8f2"
document_assembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentence_detector = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentences')
tokenizer = Tokenizer()\
.setInputCols(["sentences"])\
.setOutputCol("token")
pos = PerceptronModel.pretrained()\
.setInputCols(['document', 'token'])\
.setOutputCol('pos')
pipeline = Pipeline(stages=[document_assembler,
sentence_detector,
tokenizer,
pos
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = pipeline.fit(empty_data)
# + [markdown] id="27K8XSl-m_qO"
# **IMPORTANT!**
# In Lightpipelines, you can not use Custom annotators
# + id="hyPiXfT7W3MI"
from sparknlp.base import LightPipeline
light_model = LightPipeline(model)
# + id="366eQ_9XX5bf"
light_result = light_model.annotate("John and Peter are brothers. However they don't support each other that much.")
# + colab={"base_uri": "https://localhost:8080/"} id="MVPieouOX5Uh" executionInfo={"status": "ok", "timestamp": 1645432025858, "user_tz": -180, "elapsed": 3, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="a130b909-7933-41a8-af9f-70e5f<PASSWORD>a7"
list(zip(light_result['token'], light_result['pos']))
# + [markdown] id="fn5AVab0wJxN"
# -------------
# # Spark NLP Annotation UDFs
# + colab={"base_uri": "https://localhost:8080/"} id="GMwLediPtdg2" executionInfo={"status": "ok", "timestamp": 1645431024087, "user_tz": -180, "elapsed": 4466, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="dafe2f6e-a9df-43fa-a7bd-a4c2a82fec11"
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")\
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
pos = PerceptronModel.pretrained()\
.setInputCols(['document', 'token'])\
.setOutputCol('pos')
pipeline = Pipeline().setStages([
documentAssembler,
tokenizer,
pos])
# + colab={"base_uri": "https://localhost:8080/"} id="BfIRbDFNtdbg" executionInfo={"status": "ok", "timestamp": 1645431025674, "user_tz": -180, "elapsed": 270, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="69e53ce8-2c74-4812-ff0f-2a415c01e309"
data = spark.read.text('./sample-sentences-en.txt').toDF('text')
data.show(truncate = False)
# + id="L5rYM95qv9sE"
model = pipeline.fit(data)
# + id="TKH9HamlwALt"
result = model.transform(data)
# + colab={"base_uri": "https://localhost:8080/"} id="eFvVor1OwAJa" executionInfo={"status": "ok", "timestamp": 1645431033195, "user_tz": -180, "elapsed": 400, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="e37e7d14-e4ae-4403-b063-fe39c1db1511"
result.show(5)
# + colab={"base_uri": "https://localhost:8080/"} id="9cZDB3TrwAEI" executionInfo={"status": "ok", "timestamp": 1645431036538, "user_tz": -180, "elapsed": 349, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="4ad2a3c7-5c23-451f-b1f6-146083d966c2"
result.select('pos').show(1, truncate=False)
# + id="VM8F8D5Dw5Z0"
@udf( StringType())
def nn_annotation(res,meta):
nn = []
for i,j in zip(res,meta):
if i == "NN" or i == "NNP":
nn.append(j["word"])
return nn
# + colab={"base_uri": "https://localhost:8080/"} id="MRzmPohYw5XR" executionInfo={"status": "ok", "timestamp": 1645431041130, "user_tz": -180, "elapsed": 406, "user": {"displayName": "Monster C", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08787989274818793476"}} outputId="8d140c18-1b88-4c11-b607-9e9e0451641b"
result.withColumn("nn & NNp tokens", nn_annotation(col("pos.result"), col("pos.metadata")))\
.select("nn & NNp tokens")\
.show(truncate=False)
|
tutorials/PySpark/8.PySpark_CustomAnnotators_UDF_and_Lightpipelines.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # End-To-End Example: Get Calories For Popular Beers
#
# This example uses a data file `"ETEE-beer-calories.txt"` which contains calorie information for 254 popular beers. The calories are per 12 fluid ounces.
#
# The data file looks like this:
# ```
# ...
# Abita Purple Haze,128
# Abita Restoration,167
# Abita Turbodog,168
# Amstel Light,99
# Anchor Porter,209
# ...
# ```
#
# Let's write a program to search for a beer by name and retrieve the number of calories in 12 ounces.
#
# Example Run:
#
# ```
# Enter a beer name: Stella
# Searching for Stella...
# Stella Artois has 154 calories per 12oz.
# ```
filename = "ETEE-beer-calories.txt"
try:
with open (filename,"r") as f:
beer = input("Enter a beer name: ").title()
print("Searching for %s..." % (beer))
for line in f.readlines():
if beer in line:
beer_name = line.split(',')[0]
calories = int(line.split(',')[1])
print("%s has %d calories per 12oz." % (beer_name, calories))
break
else:
print("I could not find %s" % (beer))
except FileNotFoundError:
print("Could not find data file '%s'" % (filename))
# +
|
content/lessons/08/End-To-End-Example/ETEE-Beer-Calories.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
from nltk.corpus import reuters
nltk.download('reuters')
reuters.fileids()
reuters.categories()
reuters.fileids(['wheat','rice'])
nltk.download('punkt')
for fileid in reuters.fileids(['wheat','rice']):
num_chars = len(reuters.raw(fileid))
num_words = len(reuters.words(fileid))
num_sents = len(reuters.sents(fileid))
num_vocab = len(set(w.lower() for w in reuters.words(fileid)))
print(fileid, " : ", num_chars, num_words, num_sents, num_vocab)
fileid = 'test/15618'
reuters.raw(fileid)
reuters.words(fileid)
reuters.sents(fileid)
set (w.lower() for w in reuters.words(fileid))
nltk.download('stopwords')
from nltk.corpus import stopwords
wordList = [w for w in reuters.words(fileid) if w.lower() not in stopwords.words('english')]
wordList
from nltk import word_tokenize
tokens = word_tokenize (reuters.raw(fileid))
wordList = reuters.words(fileid)
tokens
wordList[12:20]
reuters.raw(fileid).find('MARKET')
nltk.download('wordnet')
from nltk.corpus import wordnet as wn
wn.synsets('trade')
wn.synset('trade.n.02').lemma_names()
wn.synset('trade.n.01').lemma_names()
text = nltk.Text(word.lower() for fileid in reuters.fileids(['wheat','rice']) for word in reuters.words(fileid))
text.similar('wheat')
nltk.download('averaged_perceptron_tagger')
nltk.pos_tag(tokens)
pattern = """NP: {<DT>?<RB.?>?<VBG>?<JJ.?>*(<NN.?>|<PRP.?>)+}
VP: {<VB.?>+<.*>*}
"""
mySentence = "A fastly running beautiful deer skidded off the road"
myParser = nltk.RegexpParser(pattern)
myParsedSentence = myParser.parse(nltk.pos_tag(nltk.word_tokenize(mySentence)))
print(myParsedSentence)
from IPython.display import display
display(myParsedSentence)
mySentence = "I left to do my homework"
myParser = nltk.RegexpParser(pattern)
myParsedSentence = myParser.parse(nltk.pos_tag(nltk.word_tokenize(mySentence)))
print(myParsedSentence)
from IPython.display import display
display(myParsedSentence)
# +
pattern = """NP: {<DT>?<RB.?>?<VBG>?<JJ.?>*(<NN.?>|<PRP.?>)+}
VP: {<VB.?>+<.*>*}
}(<VBG>|(<TO><.*>*)){
"""
mySentence = "I left to do my homework"
myParser = nltk.RegexpParser(pattern)
myParsedSentence = myParser.parse(nltk.pos_tag(nltk.word_tokenize(mySentence)))
print(myParsedSentence)
# -
from IPython.display import display
display(myParsedSentence)
# +
from nltk import CFG
myGrammar = nltk.CFG.fromstring("""
S -> NP VP
VP -> VB NP
VP -> VB
VP -> VB PRP
NP -> DET NN
VB -> "chased"|"ate"
DET -> "another"|"the"
NN -> "cat"|"rat"|"snake"
PRP -> "it"
""")
from nltk.parse.generate import generate
for sent in generate(myGrammar):
print(' '.join(sent))
# -
fdist = nltk.FreqDist(wordList)
fdist
# +
from nltk.util import ngrams
bigrams = ngrams(tokens,2)
for b in bigrams:
print(b)
# -
trigrams = ngrams(tokens, 3)
for t in trigrams:
print(t)
# +
nltk.download('brown')
from nltk.corpus import brown
fileid = 'ck23'
from nltk.corpus import stopwords
wordList = [w for w in brown.words(fileid) if w.lower() not in stopwords.words('english')]
# +
import string
wordList = [w for w in wordList if w not in string.punctuation]
# -
from nltk.stem import PorterStemmer
from nltk.stem import LancasterStemmer
porter = PorterStemmer()
lancaster = LancasterStemmer()
stemmersCompared = [word + ' : ' + porter.stem(word) + ' : ' + lancaster.stem(word) for word in wordList]
stemmersCompared
# +
from nltk.stem import WordNetLemmatizer
wordNet = WordNetLemmatizer()
stemmersCompared = [word + ' : ' + porter.stem(word) + ' : ' + lancaster.stem(word) + " : " + wordNet.lemmatize(word) for word in wordList]
stemmersCompared
# -
|
NLP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <hr style="border-width:2px;border-color:#84C7F7">
# <center><h1> Meta-learning competition </h1></center>
# <center><h2> Few-shot learning </h2></center>
# <hr style="border-width:2px;border-color:#84C7F7">
#
# Make sure you have the **meta_dataset** and **metadl** packages installed in your kernel environment. If you ran the <code>quick_start.sh</code> script, make sure you activated the **metadl** conda environment before launching the jupyter notebook. Here is the link of the [CodaLab competition](https://competitions.codalab.org/competitions/26212?secret_key=<KEY>) where you can submit your code and check the leaderboard.
#
#
# <u>**Outline**</u> :
# * **I - Data exploration** : We define the few-shot learning setup and explore how the data is formatted
# * **II - Submission details** : We present how a submission should be organized
# * **III - Test and submission** : We present how to test a potential submission and also how to zip your scripts to submit your code on CodaLab.
# +
import os
from collections import Counter
import gin
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from meta_dataset.data import config
from meta_dataset.data import pipeline
from meta_dataset.data import learning_spec
from meta_dataset.data import dataset_spec as dataset_spec_lib
tf.get_logger().setLevel('INFO')
def plot_episode(support_images, support_class_ids, query_images,
query_class_ids, size_multiplier=1, max_imgs_per_col=10,
max_imgs_per_row=10):
"""Plots the content of an episode. Episodes are composed of a support set
(training set) and a query set (test set). The different numbers of examples in
each set will be detailled in the starting kit.
Args:
- support_images : tuple, (Batch_size_support, Height, Width, Channels)
- support_class_ids : tuple, (Batch_size_support, N_class)
- query_images : tuple, (Batch_size_query, Height, Width, Channels)
- size_multiplier : dilate or shrink the size of displayed images
- max_imgs_per_col : Integer, Number of images in a column
- max_imgs_per_row : Integer, Number of images in a row
"""
for name, images, class_ids in zip(('Support', 'Query'),
(support_images, query_images),
(support_class_ids, query_class_ids)):
n_samples_per_class = Counter(class_ids)
n_samples_per_class = {k: min(v, max_imgs_per_col)
for k, v in n_samples_per_class.items()}
id_plot_index_map = {k: i for i, k
in enumerate(n_samples_per_class.keys())}
num_classes = min(max_imgs_per_row, len(n_samples_per_class.keys()))
max_n_sample = max(n_samples_per_class.values())
figwidth = max_n_sample
figheight = num_classes
if name == 'Support':
print('#Classes: %d' % len(n_samples_per_class.keys()))
figsize = (figheight * size_multiplier, figwidth * size_multiplier)
fig, axarr = plt.subplots(
figwidth, figheight, figsize=figsize)
fig.suptitle('%s Set' % name, size='15')
fig.tight_layout(pad=3, w_pad=0.1, h_pad=0.1)
reverse_id_map = {v: k for k, v in id_plot_index_map.items()}
for i, ax in enumerate(axarr.flat):
ax.patch.set_alpha(0)
# Print the class ids, this is needed since, we want to set the x axis
# even there is no picture.
ax.set(xlabel=reverse_id_map[i % figheight], xticks=[], yticks=[])
ax.label_outer()
for image, class_id in zip(images, class_ids):
# First decrement by one to find last spot for the class id.
n_samples_per_class[class_id] -= 1
# If class column is filled or not represented: pass.
if (n_samples_per_class[class_id] < 0 or
id_plot_index_map[class_id] >= max_imgs_per_row):
continue
# If width or height is 1, then axarr is a vector.
if axarr.ndim == 1:
ax = axarr[n_samples_per_class[class_id]
if figheight == 1 else id_plot_index_map[class_id]]
else:
ax = axarr[n_samples_per_class[class_id], id_plot_index_map[class_id]]
ax.imshow(image / 2 + 0.5)
plt.show()
def iterate_dataset(dataset, n):
""" Iterates over an episode generator represented by dataset.
It yields n episodes. An episode is a tuple containing images from
the support (train set) and query set (test set). A full episode description
is available in the starting kit.
"""
if not tf.executing_eagerly():
iterator = dataset.make_one_shot_iterator()
next_element = iterator.get_next()
with tf.Session() as sess:
for idx in range(n):
yield idx, sess.run(next_element)
else:
for idx, episode in enumerate(dataset):
if idx == n:
break
yield idx, episode
# -
# # I - Data exploration
# The goal of this section is to familiarize participants with the data format used in the challenge.
#
# Few-shot learning procedures aim to produce a Learner that is able to quickly adapt to unseen tasks with a few examples.
# In the standard Machine Learning setting, we usually split the data in train/test sets, these datasets then contain **examples** assumed to be generated from the same distribution. In few-shot learning, we have the same idea but with one additional level of abstraction : we have a meta-train and meta-test split (optionnally a meta-validation split as well). Indeed, meta-train and meta-test dataset are assumed to have **classes** generated from the same **task distribution**. For instance, we consider the Omniglot dataset during the public phase of the challenge. Omniglot is composed of 1623 classes that makes it interesting for meta-learning problems. We seperate these classes in 3 splits, a meta-train, meta-validation and meta-test sets.
# * **Meta-training** : with data sampled from the meta-train pool, we could meta-train a MetaLearner, i.e. try to learn the best approach to tackle different tasks.
# * **Meta-validation** : with data sampled from the meta-validation pool, we could adjust the meta-learner's hyper-parameters without worrying about any data leakage.
# * **Meta-testing** : with data sampled from the meta-test pool, we evaluate the Learner produced by the meta-learning procedure to quickly adapt to new unseen tasks. In order to measure the performance of such behavior, we define what we call **episodes**. These are small tasks, i.e. with a few training examples of unseen classes.
#
# Let's formalize some of the ideas exposed above.
#
# ## Definitions
# Previously we mentionned the possibility of generating data from a specific split pool. There are 2 different ways to generate data in this challenge. We can either generate data in the form of **episodes** or **batches**. Let's first describe these 2 methods :
#
# An **episode**, which represents a **task**, is described as follows :
# $$ \mathcal{T} = \{ \mathcal{D}_{train}, \mathcal{D}_{test}\}$$
# where $\mathcal{D}_{train} = \{x_{i}, y_{i}\}_{i \in \mathcal{I}_{train}}$ is the training set of the task, often called **support set**. $\mathcal{D}_{test} = \{x_{i}, y_{i}\}_{i \in \mathcal{I}_{test}}$ is the test set of the task, often called **query set**. Note that $\mathcal{I}_{train}$ and $\mathcal{I}_{test}$ are indices of the train and test set examples respectively.
#
# A **batch** is a collection of examples sampled from a split pool but **without enforcing a configuration**. For instance, let's say we want to generate data in batch mode from the meta-train pool. We can specify the batch size which is the number of examples to be sampled from the pool. We would directly sample examples from the pool without sampling **classes** as it is the case for episodes. More importantly, there would be no aforementionned $\mathcal{D}_{test}$ unlike the episodic setting. In order to better visualize the difference with the episode setting, we provide a figure below to illustrate these 2 methods.
#
# 
#
#
# ## The few-shot learning problem
#
# The few-shot learning problems are often referred as N-way K-shots problem. This name refers to episodes configuration at **meta-test time**. The number of **ways** N denotes the number of classes in an episode that represents an image classification problem. The number of **shots** K denotes the number of examples per class in the **support set**. In our case, we focus on the **5-way 1-shot** setting. In other words, episodes at meta-test time represent image classification problems with exactly 5 classes, and the **support set** contains 1 labelled example per class. More formally, $|\mathcal{I}_{train} | = 5$.
# Let's summarize the different parts of the meta-learning procedure.
#
# * At **meta-train** time : This is the part you have control on. You can choose to generate data from the meta-train split in the form of **episodes** or **batches**.
# * At **meta-test** time : We always evaluate your few-shot learning algorithm using the same setting, we generate new unseen tasks from the meta-test pool in the form of episodes. Actually these episodes have a fixed configuration, the 5-way 1-shot setting. It essentially means that when you receive a new unseen task, the support set (i.e. train set) will be composed of 5 examples, 1 for each class represented. The query set (i.e. test set) is composed of multiple unlabelled examples corresponding to theses classes. We control the number of examples in the query set, and it depends on the challenge phase. For instance, for the Omniglot dataset, the episodes' query sets at meta-test time are composed of 19 examples per class (95 examples in total).
#
# In this challenge, the episodes are generated **on the fly** from our datasets. Also, it is worth mentioning that the episodes and batches are coming from **generators**, meaning that there are virtually infinite.
#
# The number of examples in the **query set** usually depends on the number of examples for each class available in a dataset. In the public dataset (Omniglot), each class has 20 examples and we thus decide to set $|\mathcal{I}_{test}| = |N| * 19$ = 95. A visual example using the Omniglot dataset using this setting is displayed in the figure below. Note that this setting is an example, one could change the way the data is received at meta-train time. For example we could change the number of classes in an episode at meta-train time as in the prototypical networks algorithm. That is, you creates episodes containing 60 classes at meta-train time and evaluate the meta-learning algorithm performance with episodes containing 5 classes at meta-test time.
#
#
#
#
# **Note** : Make sure you have downloaded the public data under <code>omniglot/</code> directory in the root directory of this project, i.e. **../metadl**.
# Let's see how it looks like in practice :
# +
from metadl.data.dataset import DataGenerator
config_episode = [28, 5, 1, 19] # [img_size, N_ways, K_shot, nbr_query_ex]
meta_train_dir = '../../omniglot/meta_train' # Path to Public data
# The DataGenerator initialization creates 2 generators as attributes :
# Meta-train data generator : meta_train_pipeline
# Meta-valid data generator : meta_valid_pipeline
data_generator = DataGenerator(path_to_records=meta_train_dir,
batch_config=None,
episode_config=config_episode,
valid_episode_config=config_episode,
pool='train',
mode='episode')
meta_train_generator = data_generator.meta_train_pipeline
meta_valid_generator = data_generator.meta_valid_pipeline
# -
# In the previous cell, we created a <code>DataGenerator</code> object. You receive data during meta-training through this object. Notice that you can specify the configuration of meta-train and meta-valid episodes, but you could switch to <code>mode='batch'</code> if you think it would improve your meta-algorithm performance. We are going to visualize data generated as **episodes** and **batches** in the next code cells.
# +
N_EPISODES=2
dataset_spec = dataset_spec_lib.load_dataset_spec(meta_train_dir)
all_dataset_specs = [dataset_spec]
for idx, (episode, source_id) in iterate_dataset(meta_train_generator, N_EPISODES):
print('Episode id: %d from source %s' % (idx, all_dataset_specs[source_id].name))
episode = [a.numpy() for a in episode]
plot_episode(support_images=episode[0], support_class_ids=episode[2],
query_images=episode[3], query_class_ids=episode[5])
# -
# In the figures above, you can observe the composition of an episode : A **support set** (train) and a **query set** (test). In the next cell, we present some useful caracteristics of an episode.
# +
print('Length of the tuple describing an episode : {} \n'\
.format(len(episode)))
print('#'*70)
print('\nThe episode tuple is organized the following way : \n \n ' +
'[Support_images, Support_labels, Support_original_labels,' +
'Query_images, Query_labels, Query_original_labels] \n')
print('#'*70)
print('\nThe support set images are of the following shape : {} \n'\
.format(episode[0].shape))
print('The support set labels are : {} and their shape : {} \n'\
.format(episode[1], episode[1].shape))
print('The support set original labels in the dataset from which' +
' they are sampled : {} and shape : {}\n'\
.format(episode[2], episode[2].shape))
print('#'*70)
print('\nThe query set images are of the following shape : {} \n'\
.format(episode[3].shape))
print('The query set labels shape is : {} \n'\
.format(episode[4].shape))
print('The query set original labels shape in the dataset from which' +
' they are sampled is : {} \n'\
.format(episode[5].shape))
# -
# Now let's take a look at the **batch** mode.
# Let's define the batch configuration as \[28, 30\] indicating that we'd like to receive data from the meta-train split in batches of 30 images of shape \[28,28,3\].
# +
# The DataGenerator initialization creates 2 generators as attributes :
# Meta-train data generator : meta_train_pipeline
# Meta-valid data generator : meta_valid_pipeline
batch_data_generator = DataGenerator(path_to_records=meta_train_dir,
batch_config=[28, 30],
episode_config=None,
valid_episode_config=[28,5,1,19],
pool='train',
mode='batch')
meta_train_generator = batch_data_generator.meta_train_pipeline
meta_valid_generator = batch_data_generator.meta_valid_pipeline
meta_train_iterator = meta_train_generator.__iter__()
((images, labels), _) = next(meta_train_iterator)
print(f'Batch images shape : {images.shape}')
print(f'Batch labels shape : {labels.shape}')
def plot_batch(images, labels, size_multiplier=1):
""" Plot the images in a batch. Notice that labels,
corresponds to images original class id_s.
Args:
images: tf.Tensor, shape
(batch_size, image_size, image_size, 3)
labels: tf.Tensor, shape (batch_size,)
size_multiplier: Float, defines how big images will
be displayed.
"""
num_examples = len(labels)
figwidth = np.ceil(np.sqrt(num_examples)).astype('int32')
figheight = num_examples // figwidth
figsize = (figwidth * size_multiplier, (figheight + 2.5) * size_multiplier)
_, axarr = plt.subplots(figwidth, figheight, dpi=300, figsize=figsize)
for i, ax in enumerate(axarr.transpose().ravel()):
# Images are between -1 and 1.
ax.imshow(images[i] / 2 + 0.5)
ax.set(xlabel=str(labels[i].numpy()), xticks=[], yticks=[])
plt.show()
plot_batch(images, labels)
# -
# For the challenge, you don't need to create your generators, you will receive a DataGenerator object (thus already initialized). The way you receive the DataGenerator object will be described in the next section. The default setting is the episodic setting 5-way 1-shot for every meta-split. However, if you do think you could achieve better performance with your own meta-training setting, you can specify it. In order to specify your own setting, you need to write down your settings in a gin file named **config.gin** and put it in your submission folder before zipping it. We will go over the structure of submission folder in the next sections. Here is an example of a config file for the prototypical networks algorithms :
#
# **Content of a <code>config.gin</code> file**:
# ```bash
# DataGenerator.batch_config = None
# DataGenerator.episode_config = [ 28, 60, 1, 5 ]
# DataGenerator.valid_episode_config = [ 28, 5, 1, 19 ]
# DataGenerator.pool = 'train'
# DataGenerator.mode = 'episode'
# ```
# First, notice the configuration of episodes coming from the meta-train split, described by **episode_config**. The first value denotes the image size of the received images, here we kept the original value 28. Then you can specify the number of classes in your episodes, here it is set to 60! Then you specify the number of shots **K**, here 1. And finally you can specify the number of query examples per class, here 5. In this example, the meta-validation episodes description, **valid_episode_config**, is set to <code>[28, 5, 1, 19]</code> to match the episode configuration at meta-test time.
#
#
# For clarity here are the configuration descriptions :
#
# <code>episode_config = [img_size, num_ways, num_shots_per_class, num_query_per_class]</code>
#
# <code>batch_config = [img_size, batch_size]</code>
#
# ---
#
# **Section summary** :
#
# * You can choose to generate data from the meta-train split in the form of episodes or batches. Default configurations are episodic but you can change it via a **config.gin** file that you put in your folder submission.
# * You can choose to have access to episodes coming from the meta-validation split to match the evaluation at meta-test time. However, we do not allow you to generate data from the meta-validation split in batch mode.
# # II - Submission details
# In this section, we will review the structure of a valid submission. We will see that the data we receive for the few-shot learning algorithm follows the aforementioned structure.
#
#
#
# The participants would have to submit a zip file containing one or several files. The crucial file to add is <code>model.py</code>. It contains the meta-learning algorithm logic. This file **has** to follow the specific API that we defined for the challenge described in the following figure :
#
# 
#
# The 3 classes with their associated methods that need to be overriden are the following :
# * **MetaLearner** : The meta-learner contains the meta-algorithm logic. The <code>meta_fit(data_generator)</code> method has to be overriden with your own meta-learning algorithm. It receives a DataGenerator object initialized with default setting or your **config.gin** file.
#
# * **Learner** : It encapsulates the logic to learn from a new unseen task. Several methods need to be overriden :
# * <code>fit(D_train)</code>: Takes a support (train) set as an argument and fit the learner according to this dataset.
# * <code>save()</code> : You need to implement a way to save your model in a pre-defined directory.
# * <code>load()</code> : You need to implement a way to load your model from the file you created in <code>save()</code>.
# * **Predictor** : The predictor contains the logic of your model to make predictions once the learner is fitted. The <code>predict(D_test)</code> encapsulates this step and takes a query (test) set as an argument, i.e. unlabelled examples.
# ## Walkthrough a submission example
#
# In this sub-section, we present how your code submission folder should look like before zipping it.
#
# **Example of a submission directory**
# ```
# proto
# | metadata (Mandatory)
# │ model.py (Mandatory)
# │ model.gin (Optional but has to have this name)
# | config.gin (Optional but has to have this name)
# │ helper.py (Optional)
# │ utils.py (Optional)
# │ ...
# ```
# <code>model.py</code> and <code>metadata</code> are the crucial files to be added. The former contains your few-shot learning algorithm and the latter is just a file for the competition server to work properly, you simply add it to your folder without worrying about it (you can find this file in any given baseline's folder). Other files could be added and it us up to you to organize your code as you'd like.
#
# ## Defining the classes
# We go through a dummy example to understand how to create a model. In the code cell below, you can find the **zero** baseline. There are 2 important remarks :
# * First, it is mandatory to **write a file** in the <code>model_dir</code> given as an argument in the <code>save()</code> method. It could be a any file, some metadata that you gathered and/or your serialized neural network, but you need to include one.
# * Then, one can notice that the shape of the tensor returned by the <code>predict</code> method is (95,5). Indeed, The number of query examples is set to 95 for the episodes generated from the **meta-test** dataset, in the **Omniglot** dataset (i.e. the public dataset). Make sure your own predictions match the shape of the corresponding challenge phase. You can check the outputs shape in the CodaLab competition website.
#
# **Note** : You can always test your algorithm with <code>run.py</code> to verify everything is working properly. We explain how to run the script in the next section.
# +
from metadl.api.api import MetaLearner, Learner, Predictor
class MyMetaLearner(MetaLearner):
def __init__(self):
super().__init__()
def meta_fit(self, meta_dataset_generator) -> Learner:
"""
Args:
meta_dataset_generator : a DataGenerator object. We can access
the meta-train and meta-validation data via its attributes.
Refer to the metadl/data/dataset.py for more details.
Returns:
MyLearner object : a Learner that stores the meta-learner's
learning object. (e.g. a neural network trained on meta-train
episodes)
"""
return MyLearner()
class MyLearner(Learner):
def __init__(self):
super().__init__()
def fit(self, dataset_train) -> Predictor:
"""
Args:
dataset_train : a tf.data.Dataset object. It is an iterator over
the support examples.
Returns:
ModelPredictor : a Predictor.
"""
return MyPredictor()
def save(self, model_dir):
""" Saves the learning object associated to the Learner. It could be
a neural network for example.
Note : It is mandatory to write a file in model_dir. Otherwise, your
code won't be available in the scoring process (and thus it won't be
a valid submission).
"""
if(os.path.isdir(model_dir) != True):
raise ValueError(('The model directory provided is invalid. Please'
+ ' check that its path is valid.'))
# Save a file for the code submission to work correctly.
with open(os.path.join(model_dir,'dummy_sample.csv'), 'w', newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=' ',
quotechar='|', quoting=csv.QUOTE_MINIMAL)
writer.writerow(['Dummy example'])
def load(self, model_dir):
""" Loads the learning object associated to the Learner. It should
match the way you saved this object in save().
"""
if(os.path.isdir(model_dir) != True):
raise ValueError(('The model directory provided is invalid. Please'
+ ' check that its path is valid.'))
class MyPredictor(Predictor):
def __init__(self):
super().__init__()
def predict(self, dataset_test):
""" Predicts the label of the examples in the query set which is the
dataset_test in this case. The prototypes are already computed by
the Learner.
Args:
dataset_test : a tf.data.Dataset object. An iterator over the
unlabelled query examples.
Returns:
preds : tensors, shape (num_examples, N_ways). We are using the
Sparse Categorical Accuracy to evaluate the predictions. Valid
tensors can take 2 different forms described below.
Case 1 : The i-th prediction row contains the i-th example logits.
Case 2 : The i-th prediction row contains the i-th example
probabilities.
Since in both cases the SparseCategoricalAccuracy behaves the same way,
i.e. taking the argmax of the row inputs, both forms are valid.
Note : In the challenge N_ways = 5 at meta-test time.
"""
# mimick the softmax outputs
dummy_pred = tf.constant([[1.0, 0, 0, 0 ,0]], dtype=tf.float32)
dummy_pred = tf.broadcast_to(dummy_pred, (95, 5))
return dummy_pred
# -
# You can refer to the <code>metadl/baselines/</code> folder if you want to see submission examples. Here are the algorithms provided :
# * The **dummy zero** baseline
# * The **Prototypical Networks** based on [J. Snell et al. - Prototypical Networks for Few-shot Learning (2017)](https://arxiv.org/pdf/1703.05175)
# * The **fo-MAML** algorithm based on [<NAME> et al. - Model-Agnostic Meta-Learning for Fast Adaptation of Deep Networks (2017)](https://arxiv.org/pdf/1703.03400)
# # III - Test and Submission
#
# Here we present the <code>run.py</code> script. It is meant to mimick what is happenning on the CodaLab platform, i.e. the competition server. Let's say you worked on an algorithm and you are ready to test it before submitting it. More specifically, it will create your MetaLearner object, run the meta-fit method and evaluate your meta-algorithm on test episodes generated from the meta-test split. You can run the script command with the following arguments :
# * <code>meta_dataset_dir</code> : The path which contains the **2 meta-datasets**, the meta-train dataset and the meta-test dataset. The <code>quick_start.sh</code> script that you executed or the Docker image, downloaded the public dataset : Omniglot.
# * <code>code_dir</code> : The path which contains your **algorithm's code** following the format we previously defined.
#
# !python -m metadl.core.run --meta_dataset_dir=../../omniglot --code_dir=../baselines/zero
# ## Prepare a ZIP file ready for submission
# Here we present how to zip your code to submit it on the CodaLab platform. As an example, we zip the folder <code>metadl/baselines/zero/</code> which corresponds to the dummy baseline which was introduced in the previous section.
# +
from zip_utils import zipdir
model_dir = '../baselines/zero/'
submission_filename = 'mysubmission.zip'
zipdir(submission_filename, model_dir)
print('Submit this file :' + submission_filename)
# -
# ## Summary
# For clarity, we summarize the steps that you should be aware of while making a submission :
# * Follow the **MetaLearner**/**Learner**/**Predictor** API to encapsulate your few-shot learning algorithm. Please make sure you name your subclasses as **MyMetaLearner**, **MyLearner** and **MyPredictor** respectively.
# * Make sure you <u>save</u> at least a file in the given <code>model_dir</code> path. If this is a trained neural network, you need to serialize it in the <code>save()</code> method, and provide code to deserialize it in the <code>load()</code> method. Examples are provided in <code>metadl/baselines/</code>.
# * In your algorithm folder, make sure you have <code>model.py</code> and <code>metadata</code> with these **exact** names. If you do use gin files, be sure to use the corresponding names as in the baselines, i.e. **model.gin** for your own model parameters and **config.gin** for the data generation configuration (batch vs episodes).
#
# ---
#
# ## Next steps
# Now you know all the steps required to create a valid code submission.
#
# Good luck !
|
starting_kit/tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tweepy
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import time
from datetime import datetime, timezone
from matplotlib.legend_handler import HandlerLine2D
from config import (consumer_key,
consumer_secret,
access_token,
access_token_secret)
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
# -
analyzer = SentimentIntensityAnalyzer()
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth, parser=tweepy.parsers.JSONParser())
def perform_analysis(target_user):
compound_list = []
positive_list = []
negative_list = []
neutral_list = []
tweet_list=[]
account_list=[]
for x in range(0, 25):
public_tweets=api.user_timeline(target_user, page=x)
for tweet in public_tweets:
result=analyzer.polarity_scores(tweet["text"])
compound_list.append(result["compound"])
positive_list.append(result["pos"])
negative_list.append(result["neg"])
neutral_list.append(result["neu"])
tweet_list.append(tweet["text"])
account_list.append(target_user)
analyze_tweet_df=pd.DataFrame({"User":account_list,
"Compound":compound_list,
"Positive":positive_list,
"Negative":negative_list,
"Neutral":neutral_list,
"Tweet":tweet_list
})
####Plotting. Do not want to send DataFrame as an arughment so plotting it same function
try:
with plt.style.context('seaborn-dark'):
plt.figure(figsize=(12,9))
plt.xlim([len(analyze_tweet_df)*-1,0])
plt.ylim(-1,1)
plt.ylabel("Tweet Polarity")
plt.xlabel("Tweets Ago")
plt.grid(color='white', linestyle='-', linewidth=1.25)
plt.title(f"Sentiment Analysis {datetime.now().strftime('%m/%d/%Y')}")
g,=plt.plot(range(len(analyze_tweet_df)*-1,0), compound_list, marker="o", linewidth=0.75, alpha=0.8, color="steelblue",label=f"Tweets\n{target_user}")
#plt.legend([g],[f"Tweet {target_user}"])
#plt.legend([g],loc='upper right',bbox_to_anchor=(1.20, 1))
plt.legend( handler_map={g: HandlerLine2D(numpoints=1)},loc='upper right',bbox_to_anchor=(1.12, 1))
plt.savefig(f"{target_user}{datetime.now().strftime('%m-%d-%Y')}.png")
except:
print("There is an issue creating plot")
return "ERROR_IN_CREATING_IMAGE"
return f"{target_user}{datetime.now().strftime('%m-%d-%Y')}.png"
def AnalyazeString(tweet_text):
#If analyze word missing then no need to analyze
#print(tweet_text)
if("analyze" in tweet_text.replace(':','').lower()):
split_text=tweet_text.split(" ")
else:
return {"exist":False}
# If users don't exist then No need to analyze
try:
my_user=api.me()["screen_name"]
mention_user=api.get_user(split_text[2])["screen_name"]
except:
return {"exist":False}
if(split_text[0].replace('@','').lower()!=my_user.lower()):
return {"exist":False}
#Second argument
if(split_text[1].replace(':','').lower()!="analyze"):
return {"exist":False}
if not mention_user:
return {"exist":False}
#If everything looks good then send true
return {"exist":True,"mention_user":split_text[2]}
# +
##starting main program
## Search analysis tweet.
mybotname="@redhotmarket"
### This variable will be used for search tweet id greater than last one.
search_tweetid=0
while(True):
#Keeping aside first find for analyze
first_found=True
#This will check if it's been published already within a day
Already_published=False
tweets=api.search(mybotname,rpp=100,since_id=search_tweetid)
for tweet in tweets["statuses"]:
### Filtering tweets. It will parse tweet to check if it's called for analyze
analyzeString_result=AnalyazeString(tweet["text"])
if(analyzeString_result["exist"]):
#print(analyzeString_result)
if(first_found):
first_found=False
#print(analyzeString_result["mention_user"])
first_analyze=analyzeString_result["mention_user"]
first_create=tweet["created_at"]
first_tweetid=tweet["id"]
first_tweetedby=tweet["user"]["screen_name"]
else:
if((first_analyze==analyzeString_result["mention_user"]) and ((datetime.strptime(first_create, "%a %b %d %H:%M:%S %z %Y")-datetime.strptime(tweet["created_at"], "%a %b %d %H:%M:%S %z %Y")).seconds <=86400)):
Already_published=True
break
##If Analaysis is completed for same user then reply to user.Else publish the result
if(Already_published):
print(f"{first_analyze} is already published")
api.update_status(f"Hi @{first_tweetedby}, analysis for @{first_analyze} completed within 24hrs. Try later",first_tweetid)
elif((datetime.strptime(str(datetime.now(timezone.utc).strftime("%a %b %d %H:%M:%S %z %Y")),"%a %b %d %H:%M:%S %z %Y")-datetime.strptime(first_create, "%a %b %d %H:%M:%S %z %Y")).seconds <=120):
#print(first_analyze)
imgname=perform_analysis(first_analyze)
#print(imgname)
if(imgname !="ERROR_IN_CREATING_IMAGE"):
api.update_with_media(imgname,f"Thank you @{first_tweetedby} for using my plot !!")
print(f"Thank you @{first_tweetedby} for using my plot !!")
else:
print("Nothing to print")
#perform_analysis(first_analyze)
#Initializing variables. THis will ensure on second run variables don't have old values
first_analyze=''
first_create=''
first_tweetid=''
first_tweetedby=''
analyzeString_result={}
search_tweetid=first_tweetid
time.sleep(120)
# -
|
SentimentAnalyzerBot.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// [](https://mybinder.org/v2/gh/oddrationale/AdventOfCode2020CSharp/main?urlpath=lab%2Ftree%2FDay21.ipynb)
// # --- Day 21: Allergen Assessment ---
using System.IO;
record Food
{
public List<string> Ingredients { get; init; }
public List<string> Allergens { get; init; }
public Food(string input)
{
Ingredients = input
.Split(" (contains ")
.First()
.Split(" ")
.ToList();
Allergens = input
.Split(" (contains ")
.Last()
.Replace(")", "")
.Split(", ")
.ToList();
}
}
var foods = File.ReadAllLines(@"input/21.txt").Select(line => new Food(line));
var allergens = foods
.Select(food => food.Allergens)
.SelectMany(allergen => allergen)
.Distinct()
.ToDictionary(
k => k,
v => foods
.Where(food => food.Allergens.Contains(v))
.Select(food => food.Ingredients)
.Aggregate<IEnumerable<string>>((a, b) => a.Intersect(b))
.ToList()
);
allergens
while (allergens.Where(kv => kv.Value.Count() > 1).Count() > 1)
{
foreach (var allergen in allergens.Where(kv => kv.Value.Count() > 1))
{
allergen.Value.RemoveAll(ingredient =>
allergens
.Where(kv => kv.Value.Count() == 1)
.Select(kv => kv.Value.First())
.Contains(ingredient)
);
}
}
allergens
var allergenIngredients = allergens.Select(kv => kv.Value.First());
var nonAllergenIngredients = Enumerable.Except(
foods.Select(food => food.Ingredients).SelectMany(f => f).Distinct(),
allergenIngredients
);
nonAllergenIngredients
.Select(i => foods
.Select(food => food.Ingredients)
.SelectMany(f => f)
.Where(f => f == i).Count()
)
.Sum()
// # --- Part Two ---
string.Join(",", allergens.OrderBy(kv => kv.Key).Select(kv => kv.Value.First()))
|
Day21.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### I import pyplot and image from matplotlib. I also import numpy for operating on the image.
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
# Read in the image and print out some stats
image = mpimg.imread('imagedata/test.jpg')
print('This image is: ',type(image),
'with dimensions:', image.shape)
# Grab the x and y size and make a copy of the image
ysize = image.shape[0]
xsize = image.shape[1]
# Note: always make a copy rather than simply using "="
color_select = np.copy(image)
print('This image is: ',type(color_select),
'with dimensions:', color_select.shape)
# Define our color selection criteria
# Note: if you run this code, you'll find these are not sensible values!!
# But you'll get a chance to play with them soon in a quiz
red_threshold = 200
green_threshold = 200
blue_threshold = 200
rgb_threshold = [red_threshold, green_threshold, blue_threshold]
color_select.size
print(color_select[0])
# +
# Identify pixels below the threshold
thresholds = (image[:,:,0] < rgb_threshold[0]) \
| (image[:,:,1] < rgb_threshold[1]) \
| (image[:,:,2] < rgb_threshold[2])
color_select[thresholds] = [0,0,0]
# Display the image
plt.imshow(color_select)
plt.show()
|
Term1/Coding_up_a_Color_Selection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %env GOOGLE_APPLICATION_CREDENTIALS = /Users/blevine/goonalytics/resources/gcloud-cred.json
from goonalytics.io.gcloudio import BigQueryer, get_thread_posts, random_thread_id
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.cluster import KMeans
from time import time
from sklearn import metrics
tposts = get_thread_posts(random_thread_id(post_count_min=4000))
labels = list(tposts.keys())
values = [tposts[key] for key in labels]
clf = TfidfVectorizer(input='content', stop_words='english', analyzer='word', norm='l2')
X = clf.fit_transform(values, labels)
true_k = len(labels)
km = KMeans(n_clusters=true_k, init='k-means++', max_iter=100, n_init=1,
verbose=True)
t0 = time()
km.fit(X)
print("done in %0.3fs" % (time() - t0))
print()
# +
km = KMeans(n_clusters=true_k, init='k-means++', max_iter=100, n_init=1,
verbose=True)
print("Clustering sparse data with %s" % km)
t0 = time()
km.fit(X)
print("done in %0.3fs" % (time() - t0))
print()
print("Homogeneity: %0.3f" % metrics.homogeneity_score(labels, km.labels_))
print("Completeness: %0.3f" % metrics.completeness_score(labels, km.labels_))
print("V-measure: %0.3f" % metrics.v_measure_score(labels, km.labels_))
print("Adjusted Rand-Index: %.3f"
% metrics.adjusted_rand_score(labels, km.labels_))
print("Silhouette Coefficient: %0.3f"
% metrics.silhouette_score(X, km.labels_, sample_size=1000))
print()
# -
|
python/goonalytics/ipy/.ipynb_checkpoints/clustering-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 02 Using Nipype to load fMRI data
# =====================
# #### Date: Feb 7 2018; Author: Farahana
# +
from nipype import SelectFiles, Node, DataSink
from nipype.interfaces.base import Bunch
import pandas as pd
# -
# We will try to do "Preprocessing" based on figure below;
#
# 
#
# We are using OpenfMRI.org dataset according to BIDS structure.
# +
# The template string
templates = { 'anat' : 'sub*/anatomy/highres001.nii*',
'func' : 'sub*/BOLD/task001_run{ses_no}/bold.nii*'}
# How to address and import using SelectFiles node
sf = Node(SelectFiles(templates),
name='selectfiles')
sf.inputs.base_directory = '/home/farahana/Documents/dataset/Multi_Subject/ds117'
# -
# We will feed the {}-based placeholder strings with values
sf.inputs.ses_no = "001"
#sf.inputs.task_name = 'reversalweatherprediction'
sf.run().outputs
# Let us look at the TSV file of the dataset
trialinfo = pd.read_table('/home/farahana/Documents/dataset/ds052/sub-01/func/sub-01_task-reversalweatherprediction_run-1_events.tsv')
trialinfo.head()
# We will split based on two conditions:
for group in trialinfo.groupby('trial_type'):
print(group)
# +
conditions = []
onsets = []
durations = []
for group in trialinfo.groupby('trial_type'):
conditions.append(group[0])
onsets.append(group[1].onset.tolist())
durations.append(group[1].duration.tolist())
subject_info = Bunch(conditions=conditions,
onsets=onsets,
durations=durations)
#subject_info.items()
# -
sink = DataSink()
experiment_dir = '/experiment_folder'
sink.inputs.base_directory = experiment_dir + '/output_folder'
import nipype.interfaces.afni as afni
realign =afni.Retroicor()
import nipype.interfaces.freesurfer as fs
coreg = fs.BBRegister()
import nipype.interfaces.ants as ants
normalize = ants.WarpTimeSeriesImageMultiTransform()
from nipype.pipeline.engine import Workflow
# Create a preprocessing workflow
preproc = Workflow(name='preproc')
preproc.connect([(sf -> realign),
(realign -> coreg),
(coreg -> normalize),
(normalize -> sink)])
preproc.connect(sf, realign, [("out_file", "in_file")])
# !bet /home/farahana/Documents/dataset/ds052/sub-01/anat/sub-01_run-1_T1w.nii.gz output/T1_bet.nii.gz
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import nibabel as nib
# +
from nilearn.plotting import plot_img, plot_anat, plot_stat_map
# Import BET from the FSL interface
from nipype.interfaces.fsl import BET
# -
# ### Plotting Before and after Brain Extraction of Anatomical maps
skullstrip = BET(in_file="/home/farahana/Documents/dataset/Multi_Subject/ds117/sub003/anatomy/highres001.nii.gz",
out_file = "output/ds117_run001_bet.nii.gz", mask = True)
skullstrip.run()
plot_anat('/home/farahana/Documents/dataset/Multi_Subject/ds117/sub003/anatomy/highres001.nii.gz',
cut_coords=(36, -27, 30))
plot_anat('output/ds117_run001_bet.nii.gz', cut_coords=(36, -27, 30))
# ### Plotting Before and after Brain Extraction of Functional maps
skullstrip_func = BET(in_file="/home/farahana/Documents/dataset/Multi_Subject/ds117/sub003/BOLD/task001_run001/bold.nii",
out_file = "output/func_ds117_run001_bet.nii.gz")
skullstrip_func.run()
from nilearn import image
func_ds117_mean = image.mean_img('/home/farahana/Documents/dataset/Multi_Subject/ds117/sub003/BOLD/task001_run001/bold.nii')
func_ds117_bet_mean = image.mean_img('output/func_ds117_run001_bet.nii.gz')
plot_anat(np.asarray (data_1.dataobj[:,:,:,0]),
cut_coords=(36, -27, 30))
aff = data_1.get_affine()
ctr = np.dot(np.linalg.inv(aff), [0, 0, 0, 1])[:3]
vmin, vmax = (0, 1) if data_1.get_fdata().dtype == np.int16 else (30, 150)
data = data_1.get_fdata()
# %matplotlib inline
import matplotlib.pyplot as plt
plt.imshow(np.rot90(data_1[:, :, ctr[2] + 5]),
cmap="gray", vmin=vmin, vmax=vmax)
plot_anat(func_ds117_bet_mean,
cut_coords=(36, -27, 30))
data_1 = nib.load('/home/farahana/Documents/dataset/Multi_Subject/ds117/sub003/BOLD/task001_run001/bold.nii')
a = np.array(data_1)
a = data_1.get_data()
a.shape
b = a[:,:,:,0]
b.shape
plot_anat(b,cut_coords=(36, -27, 30) )
a = data_1
|
02-Nipype.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import numpy as np
# get all the keys
annotation = np.load("DATA/annotation_list.npy")[()]
image_list = annotation.keys()
image_list
# +
# get a viusalization
imgfile = "2010_000948"
img = plt.imread("DATA/{}.jpg".format(imgfile))
box = annotation[imgfile]
print box
# Create figure and axes
fig,ax = plt.subplots(1)
ax.imshow(img)
for eachbox in box:
cx, cy, w, h = eachbox
# Create a Rectangle patch
rect = patches.Rectangle((cx-w/2,cy-h/2),w,h,linewidth=1,edgecolor='r',facecolor='none')
# Add the patch to the Axes
ax.add_patch(rect)
plt.show()
print "The box is {}".format(box)
print "image size is {}".format(img.shape)
# -
|
week6/mxnet-week4n5-final-project/demo_raw_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Прореживание нейронных сетей
#
# В этой лабораторной мы попробуем уменьшить размер нейронной сети за счет удаления из нее части весов.
# + pycharm={"name": "#%%\n"}
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data
from torch.autograd import Variable
from sklearn.model_selection import train_test_split
# + pycharm={"name": "#%%\n"}
SEED=9876
torch.manual_seed(SEED)
# -
# В качестве данных будем использовать стандартный mnist
# + pycharm={"name": "#%%\n"}
df = pd.read_csv('/data/mnist_784.csv')
df.head()
# + pycharm={"name": "#%%\n"}
y = df['class'].values
X = df.drop(['class'],axis=1).values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=100)
# + pycharm={"name": "#%%\n"}
plt.imshow(X_train[0].reshape(28, 28))
# -
# Первое, что мы попробуем сделать - это собрать какую-то несложную архитектуру нейронной сети и просто обучить ее на данных.
#
# После этого мы замерим ее размер, а также качество, которое она выдает. Все дальнейшие полученные модели будем сравнивать с этими результатами, как с базовыми и понимать - получилось лучше или хуже.
#
# Вначале просто подготовим данные для обучения.
# + pycharm={"name": "#%%\n"}
BATCH_SIZE = 32
torch_X_train = torch.from_numpy(X_train).type(torch.LongTensor)
torch_y_train = torch.from_numpy(y_train).type(torch.LongTensor)
torch_X_test = torch.from_numpy(X_test).type(torch.LongTensor)
torch_y_test = torch.from_numpy(y_test).type(torch.LongTensor)
train = torch.utils.data.TensorDataset(torch_X_train,torch_y_train)
test = torch.utils.data.TensorDataset(torch_X_test,torch_y_test)
train_loader = torch.utils.data.DataLoader(train, batch_size = BATCH_SIZE, shuffle = False)
test_loader = torch.utils.data.DataLoader(test, batch_size = BATCH_SIZE, shuffle = False)
# -
# В реальной жизни для задачи распознавания числа на картинке мы бы скорее всего использовали более продвинутую архитектуру сети, однако для наглядности мы возьмем простую сеть, которая при этом имеет много параметров. В ней будут просто три полносвязных слоя: 784 - 250 - 100 - 10
# + pycharm={"name": "#%%\n"}
class MLP(nn.Module):
def __init__(self):
super(MLP, self).__init__()
self.linear1 = nn.Linear(784,250)
self.linear2 = nn.Linear(250,100)
self.linear3 = nn.Linear(100,10)
def forward(self,X):
X = F.relu(self.linear1(X))
X = F.relu(self.linear2(X))
X = self.linear3(X)
return F.log_softmax(X, dim=1)
mlp = MLP()
print(mlp)
# -
# Обучаем самым обычным способом, используя кросс-энтропию в качестве меры ошибки и используя 5 эпох
# + pycharm={"name": "#%%\n"}
def fit(model, train_loader, epoch_number=5):
optimizer = torch.optim.Adam(model.parameters())
error = nn.CrossEntropyLoss()
model.train()
for epoch in range(epoch_number):
correct = 0
for batch_idx, (X_batch, y_batch) in enumerate(train_loader):
var_X_batch = Variable(X_batch).float()
var_y_batch = Variable(y_batch)
optimizer.zero_grad()
output = model(var_X_batch)
loss = error(output, var_y_batch)
loss.backward()
optimizer.step()
predicted = torch.max(output.data, 1)[1]
correct += (predicted == var_y_batch).sum()
if batch_idx % 50 == 0:
print('Epoch : {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}\t Accuracy:{:.3f}%'.format(
epoch, batch_idx*len(X_batch), len(train_loader.dataset), 100.*batch_idx / len(train_loader), loss.data, float(correct*100) / float(BATCH_SIZE*(batch_idx+1))))
# + pycharm={"name": "#%%\n"}
torch.manual_seed(SEED)
fit(mlp, train_loader)
# -
# В качестве метрики качества возьмем обычный accuracy
# + pycharm={"name": "#%%\n"}
def evaluate(model):
correct = 0
for test_imgs, test_labels in test_loader:
test_imgs = Variable(test_imgs).float()
output = model(test_imgs)
predicted = torch.max(output,1)[1]
correct += (predicted == test_labels).sum()
print("Test accuracy:{:.3f}% ".format( float(correct) / (len(test_loader)*BATCH_SIZE)))
evaluate(mlp)
# -
# Весьма неплохое качество, учитывая, что мы почти ничего не придумывали с сетью.
#
# Посмотрим, сколько параметров нам потребовалось, чтобы получить это качество.
# + pycharm={"name": "#%%\n"}
def calc_weights(model):
result = 0
for layer in model.children():
result += len(layer.weight.reshape(-1))
return result
# + pycharm={"name": "#%%\n"}
calc_weights(mlp)
# -
# Видно, что полносвязные слои достаточно тяжелые и всего три слоя дали нам больше чем 200 000 параметров. Попробуем ужать это число, не сильно уменьшим при этом качество.
# # Удаляем связи внутри сети
#
# Для того, чтобы начать оптимизировать размер сети, нам нужен инструментарий для удаления связей внутри нашей модели.
#
# Нам потребуется особый полносвязный слой, в котором мы можем отключать конкретные веса. Используя такие слои, соберем такую же архитектруру с тремя полносвязными.
#
# Отключать сами веса мы будем исходя из их абсолютного значения - задавая пороговое значение, мы будем занулять только те веса, которые меньше этого значения.
# + pycharm={"name": "#%%\n"}
class MaskedLinear(nn.Module):
def __init__(self, in_size, out_size):
super(MaskedLinear, self).__init__()
# Обычный полносвязный слой
self._linear = nn.Linear(in_size, out_size)
# Маска для слоя. Для связи из оригинального слоя, здесь будут хранится 0 и 1.
# 1 - связь действует, 0 - связь не действует.
self._mask = nn.Linear(in_size, out_size)
# Изначально все числа в маске - 1. То есть изначально мы не выключаем вообще никакие веса
self._mask.weight.data = torch.ones(self._mask.weight.size())
def forward(self, x):
# Чтобы применить этот слой, нужно вначале умножить веса на маску.
# Тогда те веса, которые мы выключили, занулятся, что и будет означать, что мы их просто выкинули
self._linear.weight.data = torch.mul(self._linear.weight, self._mask.weight)
return self._linear(x)
def prune(self, threshold):
# Для того, чтобы выключить часть связей задается threshold
# Если значение веса по модулю в сети меньше, чем threshold, то мы его выключаем, а значит выставляем 0 в маске.
self._mask.weight.data = torch.mul(torch.gt(torch.abs(self._linear.weight), threshold).float(), self._mask.weight)
# -
# Составляем точно такую же архитектуру, но используя наши особенные полносвязные слои, в которых мы можем отключать веса
# + pycharm={"name": "#%%\n"}
class AutoCompressMLP(nn.Module):
def __init__(self):
super(AutoCompressMLP, self).__init__()
self.linear1 = MaskedLinear(784,250)
self.linear2 = MaskedLinear(250,100)
self.linear3 = MaskedLinear(100,10)
def forward(self,X):
X = F.relu(self.linear1(X))
X = F.relu(self.linear2(X))
X = self.linear3(X)
return F.log_softmax(X, dim=1)
def prune(self, threshold):
self.linear1.prune(threshold)
self.linear2.prune(threshold)
self.linear3.prune(threshold)
# -
# Чтобы удалить какую-то долю связей из сети, необходимо вначале подсчитать необходимое пороговое значение.
#
# Так, чтобы удалить N% связей по этой схеме, необходимо найти такое число, чтобы ровно N% связей имело вес меньше этого числа по модулю. Другими словами найти N-перцентиль.
#
# Напишем функцию, которая будет искать такое пороговое значение.
# + pycharm={"name": "#%%\n"}
def calc_threshhold(model, rate):
all_weights = torch.Tensor()
for layer in model.children():
all_weights = torch.cat( (layer._linear.weight.view(-1), all_weights.view(-1)) )
abs_weight = torch.abs(all_weights)
return np.percentile(abs_weight.detach().cpu().numpy(), rate)
# + pycharm={"name": "#%%\n"}
acmlp = AutoCompressMLP()
t = calc_threshhold(acmlp, 50.0)
t
# -
# Чтобы следить за тем, сколько параметров осталось внури нашей сети, нам потребуется немного другая функция подсчета активных весов, учитываящая маску.
# + pycharm={"name": "#%%\n"}
def calc_pruned_weights(model):
result = 0
for layer in model.children():
result += torch.sum(layer._mask.weight.reshape(-1))
return int(result.item())
# + pycharm={"name": "#%%\n"}
acmlp.prune(t)
calc_pruned_weights(acmlp)
# -
# # Итеративное прореживание
#
# Идин из способов сжатия нейронных сетей - итеративное прореживание (Incremental Magnitude Pruning). Он достаточно ресурсоемкий, однако позволяет достаточно несложными методами добиться неплохого результата.
# + pycharm={"name": "#%%\n"}
acmlp = AutoCompressMLP()
# -
# Вначале просто обучим нашу модель, никаким образом ее не модифицируя.
# + pycharm={"name": "#%%\n"}
torch.manual_seed(SEED)
fit(acmlp, train_loader)
# + pycharm={"name": "#%%\n"}
evaluate(acmlp)
# -
# Отлично, получили примерно такую же модель, как и в самом начале.
#
# Сейчас модель уже имеет хорошие веса для предсказаний. Теперь попробуем убрать из нее 50% связей и посмотрим, насколько ей удастся сохранить качество.
#
# Как уже отмечалось, отключим 50% наиболее слабых связей в сети.
# + pycharm={"name": "#%%\n"}
import copy
acmlp_test1 = copy.deepcopy(acmlp)
# + pycharm={"name": "#%%\n"}
t_50 = calc_threshhold(acmlp_test1, 50.0)
acmlp_test1.prune(t_50)
# + pycharm={"name": "#%%\n"}
evaluate(acmlp_test1)
# -
# Можно заметить, что таким образом выкинутые веса почти не повлияли на качество сети. При этом мы выкинули половину всех коэффициентов! Весьма неплохой результат.
#
# Давайте посмотрим, можем ли мы с таким же успехом выкинуть 90% сети?
# + pycharm={"name": "#%%\n"}
acmlp_test2 = copy.deepcopy(acmlp)
t_90 = calc_threshhold(acmlp_test2, 90.0)
acmlp_test2.prune(t_90)
# + pycharm={"name": "#%%\n"}
evaluate(acmlp_test2)
# -
# Увы, так просто выкинуть 90% и оставить качество не получается. Будем использовать более хитрый подход.
#
# Будет идти с шагом в 10%. Каждый раз будет отключать внутри сети 10% связей. После отключения, оставшиеся веса дообучим на всех данных используя всего одну эпоху. Ожидается, что так как мы выкинули за один раз не очень много, то оставшиеся связи "перехватят" ответственность тех слабых, которые мы только что отключили.
#
# Таким образом за P таких итераций мы выкинем 10P% всей сети и не должны при этом потерять сильно в качестве.
# + pycharm={"name": "#%%\n"}
def smart_prune(model, train_loader, compress_rate):
# Создаем именно новую модель, старую не трогаем
model = copy.deepcopy(model)
optimizer = torch.optim.Adam(model.parameters())
error = nn.CrossEntropyLoss()
model.train()
for rate in range(0, compress_rate+1, 10): # Идем с шагом в 10%
t = calc_threshhold(model, float(rate)) # Считаем очередное пороговое значение
model.prune(t) # Отключаем слабые связи
correct = 0
for batch_idx, (X_batch, y_batch) in enumerate(train_loader): # Далее дообучаем модель как обычно в течение одной эпохи
var_X_batch = Variable(X_batch).float()
var_y_batch = Variable(y_batch)
optimizer.zero_grad()
output = model(var_X_batch)
loss = error(output, var_y_batch)
loss.backward()
optimizer.step()
predicted = torch.max(output.data, 1)[1]
correct += (predicted == var_y_batch).sum()
if batch_idx % 20 == 0:
print('Rate : {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}\t Accuracy:{:.3f}%'.format(
rate, batch_idx*len(X_batch), len(train_loader.dataset), 100.*batch_idx / len(train_loader), loss.data, float(correct*100) / float(BATCH_SIZE*(batch_idx+1))))
return model
# -
# Попробуем для начала выкинуть 70% таким образом
# + pycharm={"name": "#%%\n"}
torch.manual_seed(SEED)
pruned_model = smart_prune(acmlp, train_loader, 70)
# + pycharm={"name": "#%%\n"}
evaluate(pruned_model)
# -
# На моем компьютере получилось качество около 0.97. Формально это даже чуточку лучше, чем оригинальная модель! Получается, что лишние веса в оригинальной модели могли мешали выявить зависимость в данных.
#
# Давайте посчитаем количество ненулевых весов в модели
# + pycharm={"name": "#%%\n"}
calc_pruned_weights(acmlp)
# + pycharm={"name": "#%%\n"}
calc_pruned_weights(pruned_model)
# -
# Оставили около 60 000 весов, мы получили почти такое же качество для модели!
#
# Можем ли мы таким же образом выкинуть 90%?
# + pycharm={"name": "#%%\n"}
torch.manual_seed(SEED)
pruned_model_90 = smart_prune(acmlp, train_loader, 90)
# + pycharm={"name": "#%%\n"}
evaluate(pruned_model_90)
# + pycharm={"name": "#%%\n"}
calc_pruned_weights(pruned_model_90)
# -
# Выкинув большую часть сети, мы все еще имеем относительно неплохое качество, хоть и меньше, чем изначально.
#
# Вполне возможно проблема в том, что мы слишком агрессивно удаляем связи, когда их остается совсем мало. Давайте попробуем более аккуратные шаги.
#
# + pycharm={"name": "#%%\n"}
def smart_prune_shed(model, train_loader, schedule):
# Создаем именно новую модель, старую не трогаем
model = copy.deepcopy(model)
optimizer = torch.optim.Adam(model.parameters())
error = nn.CrossEntropyLoss()
model.train()
for rate, epochs in schedule: # Идем шагами, согласно тому расписанию, которое передали в функцию
t = calc_threshhold(model, float(rate)) # Считаем очередное пороговое значение
model.prune(t) # Отключаем слабые связи
for i in range(epochs):
correct = 0
for batch_idx, (X_batch, y_batch) in enumerate(train_loader): # Далее дообучаем модель как обычно в течение указанного количества эпох
var_X_batch = Variable(X_batch).float()
var_y_batch = Variable(y_batch)
optimizer.zero_grad()
output = model(var_X_batch)
loss = error(output, var_y_batch)
loss.backward()
optimizer.step()
predicted = torch.max(output.data, 1)[1]
correct += (predicted == var_y_batch).sum()
if batch_idx % 20 == 0:
print('Rate : {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}\t Accuracy:{:.3f}%'.format(
rate, batch_idx*len(X_batch), len(train_loader.dataset), 100.*batch_idx / len(train_loader), loss.data, float(correct*100) / float(BATCH_SIZE*(batch_idx+1))))
return model
# + pycharm={"name": "#%%\n"}
torch.manual_seed(SEED)
pruned_model_90 = smart_prune_shed(acmlp, train_loader, [
(0, 1),
(20, 1),
(40, 1),
(50, 1),
(60, 1),
(70, 1),
(75, 1),
(80, 2),
(83, 2),
(85, 2),
(86, 2),
(87, 2),
(88, 2),
(89, 2),
(90, 2)
])
# + pycharm={"name": "#%%\n"}
evaluate(pruned_model_90)
# + pycharm={"name": "#%%\n"}
calc_pruned_weights(pruned_model_90)
# -
# Чтож, это оригинальное качество за всего 10% сети.
#
# Ради интереса попробуем "дожать до победы" и удалим 99%.
# + pycharm={"name": "#%%\n"}
torch.manual_seed(SEED)
pruned_model_99 = smart_prune_shed(pruned_model_90, train_loader, [
(90, 2),
(92, 2),
(94, 2),
(95, 2),
(96, 2),
(97, 2),
(98, 2),
(99, 2)
])
# + pycharm={"name": "#%%\n"}
evaluate(pruned_model_99)
# + pycharm={"name": "#%%\n"}
calc_pruned_weights(pruned_model_99)
# -
# Видно, что метод все таки имеет свои ограничения. По логу видно, что где-то в районе 94% мы видимо задели какой-то очень важный участок сети, после удаления которого она уже не смогла восстановится.
#
# Однако результат в 90% - это тоже вполне неплохо!
# + pycharm={"name": "#%%\n"}
# -
# # Готовые реализации
#
# Сама техника достаточно популярна и часто имеет уже готовые реализации. В Pytorch имеется отдельный модуль для проведения прореживания сети.
# + pycharm={"name": "#%%\n"}
import torch.nn.utils.prune as prune
# + pycharm={"name": "#%%\n"}
class PytorchPrunedMLP(nn.Module):
def __init__(self):
super(PytorchPrunedMLP, self).__init__()
self.linear1 = nn.Linear(784,250)
self.linear2 = nn.Linear(250,100)
self.linear3 = nn.Linear(100,10)
def forward(self,X):
X = F.relu(self.linear1(X))
X = F.relu(self.linear2(X))
X = self.linear3(X)
return F.log_softmax(X, dim=1)
def prune(self, rate):
# Используем l1_unstructured вместо нашего подхода
# unstructured говорит о том, что нет ограничений на удаляемые веса
# l1 говорит о том, что нужно смотреть на модуль веса
prune.l1_unstructured(self.linear1, 'weight', amount=rate)
prune.l1_unstructured(self.linear2, 'weight', amount=rate)
prune.l1_unstructured(self.linear3, 'weight', amount=rate)
# + pycharm={"name": "#%%\n"}
torch.manual_seed(SEED)
ppmlp = PytorchPrunedMLP()
fit(ppmlp, train_loader)
# + pycharm={"name": "#%%\n"}
evaluate(ppmlp)
# + pycharm={"name": "#%%\n"}
ppmlp.prune(0.5)
# + pycharm={"name": "#%%\n"}
evaluate(ppmlp)
# + pycharm={"name": "#%%\n"}
def calc_pytorch_weights(model):
result = 0
for layer in model.children():
if hasattr(layer, 'weight_mask'):
result += int(torch.sum(layer.weight_mask.reshape(-1)).item())
else:
result += len(layer.weight.reshape(-1))
return result
# + pycharm={"name": "#%%\n"}
calc_pytorch_weights(ppmlp)
# -
# Точно таким же образом мы только что выкинули 50% самых слабый весов из сети.
#
# # Групповое (структурированное) прореживание
#
# В библиотеке реализованы также и более продвинутые версии этого алгоритма. Например мы можем делать более структурированное прореживание, удаляя не единичные связи, а целиком нейроны из сети.
#
# Для того, чтобы понять, насколько тот или иной нейрон важен для работы сети, будем смотреть на все веса, связанные с ним. Если веса значительно отличаются от нуля, значит нейрон важный, если близки к нулю - значит скорее всего его можно удалить.
#
# Понимать, насколько группа нейронов близка к нулю можно разными способами. Наиболее популярный - L-нормы. Так например при L1 мы будем смотреть на сумму по модулю все весов для нейрона, а при L2 - на корень из суммы квадратов весов.
# + pycharm={"name": "#%%\n"}
class StructuredPrunedMLP(nn.Module):
def __init__(self):
super(StructuredPrunedMLP, self).__init__()
self.linear1 = nn.Linear(784,250)
self.linear2 = nn.Linear(250,100)
self.linear3 = nn.Linear(100,10)
def forward(self,X):
X = F.relu(self.linear1(X))
X = F.relu(self.linear2(X))
X = self.linear3(X)
return F.log_softmax(X, dim=1)
def prune(self, rate):
# Используем ln_structured для удаления нейронов целиком
# Для оценивания значимости нейрона будем использовать L2, поэтому указываем n=2
# Указываем dim=1 - это укажет, как именно нужно групировать веса. Для dim=1 - это группировка по нейронам
prune.ln_structured(self.linear1, 'weight', amount=rate, n=2, dim=1)
prune.ln_structured(self.linear2, 'weight', amount=rate, n=2, dim=1)
# В последнем слое удалять нейроны нельзя, потому как они отвечают за ответ сети
# + pycharm={"name": "#%%\n"}
torch.manual_seed(SEED)
spmlp = StructuredPrunedMLP()
fit(spmlp, train_loader)
# + pycharm={"name": "#%%\n"}
evaluate(spmlp)
# + pycharm={"name": "#%%\n"}
spmlp.prune(0.5)
# + pycharm={"name": "#%%\n"}
evaluate(spmlp)
# + pycharm={"name": "#%%\n"}
calc_pytorch_weights(spmlp)
# -
# Можно посмотреть на устройство весов в наших последних двух моделях
#
# У модели, которую прореживали по весам, у каждого нейрона отключены какие-то элементы
# + pycharm={"name": "#%%\n"}
ppmlp.linear1.weight_mask.T[0]
# -
# У модели, которую прореживали по нейронам, нейрон или отключен совсем
# + pycharm={"name": "#%%\n"}
spmlp.linear1.weight_mask.T[0]
# -
# Или работает целиком
# + pycharm={"name": "#%%\n"}
spmlp.linear1.weight_mask.T[100]
|
pruning/.ipynb_checkpoints/1-compress--seminar-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from datascience import *
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
# -
# ## Example: Benford's Law
digits = np.arange(1, 10)
benford_model = np.log10(1 + 1/digits)
benford = Table().with_columns(
'First digit', digits,
'Benford model prob', benford_model)
benford.barh('First digit')
# You don't have to understand how this function works, since it uses Python features from beyond Data 8.
def first_digit(num):
return int(str(num)[0])
first_digit(32)
first_digit(17719087)
# County populations from the census data
counties = Table.read_table('counties.csv')
counties = counties.where('SUMLEV', 50).select(5,6,9).relabeled(0,'State').relabeled(1,'County').relabeled(2,'Population')
counties.show(3)
first_digits = counties.apply(first_digit, 'Population')
counties = counties.with_column('First digit', first_digits)
counties.show(3)
num_counties = counties.num_rows
by_digit = counties.group('First digit')
proportions = by_digit.column('count')/num_counties
by_digit = by_digit.with_columns(
'Proportion', proportions,
'Benford proportion', benford_model
)
by_digit.drop('count').barh('First digit')
# Null hypothesis:
# Alternative hypothesis:
# Test statistic: ___
#
# Fill in the blank with "Bigger" or "Smaller":
#
# ___ values of the test statistic favor the alternative
observed_tvd = sum(abs(proportions - benford_model))/2
observed_tvd
sample_proportions(num_counties, benford_model)
simulated_frequencies = sample_proportions(num_counties, benford_model)
tvd = sum(abs(simulated_frequencies - benford_model))/2
tvd
def simulate_county_first_digits():
simulated_frequencies = sample_proportions(num_counties, benford_model)
tvd = sum(abs(simulated_frequencies - benford_model))/2
return tvd
# +
simulated_tvds = make_array()
for i in np.arange(10000):
simulated_tvds = np.append(simulated_tvds, simulate_county_first_digits())
# -
Table().with_column('Simulated TVD', simulated_tvds).hist(0)
np.count_nonzero(simulated_tvds >= observed_tvd) / 10000
# Are the data consistent with the null hypothesis?
# ## Example: sleep survey
survey = Table.read_table('welcome_survey_v4.csv')
survey
# +
def simplify(sleep_position):
if sleep_position == 'On your left side' or sleep_position == 'On your right side':
return 'side'
else:
return 'back or stomach'
survey = survey.with_column(
'position',
survey.apply(simplify, 'Sleep position')
).select('position', 'Hours of sleep')
survey
# -
survey.group('position', np.average)
# Null hypothesis:
# Alternative hypothesis:
# Test statistic: ___
#
# Fill in the blank with "Bigger" or "Smaller":
#
# ___ values of the test statistic favor the alternative
def compute_test_statistic(tbl):
grouped = tbl.group('position', np.average)
avgs = grouped.column('Hours of sleep average')
return avgs.item(1) - avgs.item(0)
obs_test_stat = compute_test_statistic(survey)
obs_test_stat
random_labels = survey.sample(with_replacement=False).column('position')
def simulate_under_null():
random_labels = survey.sample(with_replacement=False).column('position')
relabeled_tbl = survey.with_column('position', random_labels)
return compute_test_statistic(relabeled_tbl)
simulated_diffs = make_array()
for i in np.arange(1000):
null_stat = simulate_under_null()
simulated_diffs = np.append(simulated_diffs, null_stat)
Table().with_column('Simulated difference', simulated_diffs).hist(0)
obs_test_stat
np.mean(simulated_diffs <= obs_test_stat)
# Are the data consistent with the null hypothesis?
|
lec/lec21.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# In this part(final part), the free energy is computed for a "2 part" run (e.g going backward then forward)
# The free energy is computed using EMUS PMF
# The github repo for EMUS is https://github.com/ehthiede/EMUS
# Their AlaDipeptide_1D example demonstrates many EMUS features
import sys, os, os.path
import glob
import scipy as sp
import numpy as np
from emus import usutils as uu
from emus import emus, avar
import matplotlib
import matplotlib.pyplot as pp
from mpl_toolkits.mplot3d import Axes3D
import yt
from yt.frontends.boxlib.data_structures import AMReXDataset
from tempfile import TemporaryFile
# %pylab inline
# Additonal EMUS parameters should be set here
period=None
dim=1
T=0.01
k_B=1
# -
#prepare collective varrible trajectories (samples) and umbrella biasing functions (psi) for emus
meta_file_1 = 'ONE_TO_074_META.txt' # Path to Meta File
psis_1, cv_trajs_1, neighbors_1 = uu.data_from_meta(
meta_file_1, dim, T=T, k_B=k_B, period=period)
meta_file_2 = '074_TO_ONE_META.txt' # Path to Meta File
psis_2, cv_trajs_2, neighbors_2 = uu.data_from_meta(
meta_file_2, dim, T=T, k_B=k_B, period=period)
# compute one iteration of emus, typically MANY iterations are needed
z, F = emus.calculate_zs(psis=psis_1, neighbors=neighbors_1)
# Calculate the PMF from EMUS
cv_trajs=cv_trajs_1
psis=psis_1
nbins = 60 # Number of Histogram Bins.
kT=k_B*T
domain = ((0.74, 0.97))
pmf, edges = emus.calculate_pmf(
cv_trajs, psis, domain, z, nbins=nbins, kT=kT, use_iter=False) # Calculate the pmf
pmf_centers = (edges[0][1:]+edges[0][:-1])/2.0
pp.figure()
pp.plot(pmf_centers, pmf, label='EMUS PMF')
pp.legend(['KT=0.01, $\gamma=1$'])
pp.xlabel('$\phi_x$')
pp.ylabel('$\hat{X}$')
pp.title('1.0 to 0.74')
# +
# Calculate z using the MBAR type iteration.
# error messages sometimes appear when data overlaps "too much" or not "enough" in some regions such that
# the overlap matrix F is poorly conditioned
# This is difficult to avoid, so it is important to make sure iterations converge
#z_iter_25, F_iter_25 = emus.calculate_zs(psis, n_iter=25)
z_iter_50_1, F_iter_50_1 = emus.calculate_zs(psis_1, n_iter=50)
#z_iter_100_1, F_iter_100_1 = emus.calculate_zs(psis=psis_1, n_iter=100)
#z_iter_100_2, F_iter_100_2 = emus.calculate_zs(psis=psis_2, n_iter=100)
z_iter_350_1, F_iter_350_1 = emus.calculate_zs(psis_1, n_iter=350)
z_iter_350_2, F_iter_350_2 = emus.calculate_zs(psis_1, n_iter=350)
#z_iter_1k, F_iter_1k = emus.calculate_zs(psis, n_iter=1000)
# -
nbins = 40 # Number of Histogram Bins.
kT=k_B*T
domain = ((0.74, 0.99))
iterpmf, edges = emus.calculate_pmf(
cv_trajs_1, psis_1, domain, nbins=nbins, z=z_iter_100_1, kT=kT)
pmf_centers_iter = (edges[0][1:]+edges[0][:-1])/2.
pp.plot(pmf_centers_iter, iterpmf, label='Iter EMUS PMF')
pp.legend(['KT=0.01, $\gamma=1$'])
pp.xlabel('$\phi_x$')
pp.ylabel('$\hat{X}$')
pp.title('1.0 to 0.74')
nbins = 40 # Number of Histogram Bins.
kT=k_B*T
domain = ((0.74, 0.99))
iterpmf, edges = emus.calculate_pmf(
cv_trajs_2, psis_2, domain, nbins=nbins, z=z_iter_100_2, kT=kT)
pmf_centers_iter = (edges[0][1:]+edges[0][:-1])/2.
pp.plot(pmf_centers_iter, iterpmf, label='Iter EMUS PMF')
pp.legend(['KT=0.01, $\gamma=1$'])
pp.xlabel('$\phi_x$')
pp.ylabel('$\hat{X}$')
pp.title('0.74 to 1.0')
# bellow is an example of checking the convergence of a run
a,=pp.plot(-np.log(z_iter_1),label="Iteration 1")
b,=pp.plot(-np.log(z_iter_2),label="Iteration 2")
c,=pp.plot(-np.log(z_iter_5),label="Iteration 5")
d,=pp.plot(-np.log(z_iter_10),label="Iteration 10")
e,=pp.plot(-np.log(z_iter_15),label="Iteration 15")
f,=pp.plot(-np.log(z_iter_55),label="Iteration 55")
h,=pp.plot(-np.log(z_iter_150),label="Iteration 150")
j,=pp.plot(-np.log(z_iter_350),label="Iteration 350")
k,=pp.plot(-np.log(z_iter_1000),label="Iteration 1000")
pp.legend(handles=[a, b, c,d,e,f,h,j,k])
pp.show()
|
unmaintained/_GL_alt/Python_notebooks_KL_new/EMUS_scripts/Comp_EMU_PMF(step3).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
# Data for plotting
t = np.arange(0.0, 2.0, 0.01)
s = 1 + np.sin(2 * np.pi * t)
fig, ax = plt.subplots()
ax.plot(t, s)
ax.set(xlabel='time (s)', ylabel='voltage (mV)',
title='About as simple as it gets, folks')
ax.grid()
fig.savefig("test.png")
plt.show()
# -
|
Notebookmatplotlib02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from collections import Counter
from sklearn.feature_extraction.text import TfidfVectorizer
import nltk
from sklearn.naive_bayes import MultinomialNB
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
datas = pd.read_csv('datasets/ISEAR.csv')
datas.head()
datas.columns
datas.drop('0', axis=1, inplace=True)
datas.size
datas.shape
column_name = datas.columns
datas = datas.rename(columns={column_name[0]: "Emotion",
column_name[1]: "Sentence"})
datas.head()
# Adding $joy$ back to the dataset
missing_data = {"Emotion": column_name[0],
"Sentence": column_name[1]}
missing_data
datas = datas.append(missing_data, ignore_index=True)
datas.isna().sum()
datas.tail()
y = datas['Emotion']
y.head()
X = datas['Sentence']
X.head()
Counter(y)
tfidf = TfidfVectorizer(tokenizer=nltk.word_tokenize, stop_words='english', min_df=3, ngram_range=(1, 3))
X = tfidf.fit_transform(X)
tfidf.vocabulary_
bayes_classification = MultinomialNB()
X_train, X_test, y_train, y_test = train_test_split(X, y)
bayes_classification.fit(X_train, y_train)
bayes_pred = bayes_classification.predict(X_test)
accuracy_score(y_test, bayes_pred)
|
notebooks/01.01_PL_sentiment_analysis_2020_05_13.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Atelier 1 : Apprentissage et classification Automatique
#
# ## A : Apprenstissage non-supervisé
# ### Jeu de données :
# Le jeu de données Iris a été utilisé à l''article classique de Fisher, publié en 1936, intitulé "L'utilisation de plusieurs mesures dans des problèmes taxonomiques", est également disponible dans le référentiel UCI Machine Learning.
#
# Il comprend trois espèces d’iris de 50 échantillons chacune, ainsi que des propriétés propres à chaque fleur. Une espèce de fleur est séparable linéairement des deux autres, mais les deux autres ne sont pas séparables linéairement l'une de l'autre.
#
# Les colonnes de cet ensemble de données sont:
#
# Id
# Longueur du Sépale Cm
# Largeur du Sépale Cm
# Longueur du Pétale cm
# Largeur du Pétale Cm
# Espèce : classe : Iris Setosa, Iris Versicolor ou Iris Virginica.
#
# Un échantillon : (4.9,3.6,1.4,0.1, “Iris-setosa”)
# ## Regroupement (Clustering) et visualisation de données
# #### Importation des données
# Avec la fonction read_csv de Pandas: on peut mettre dans notre dataframe le contenu du fichier csv, en indiquant comme paramètre (1: le chemin ou la source où se trouve le fichier csv, 2: les séparateurs entre les valeurs dans notre cas ces des vergules) en troisième position, un paramètre facultatif pour spécifier le type d'encodage de notre fichier exemple encoding ="UTF8".
# +
# importation des lib
import pandas as pd
import numpy as np
df = pd.read_csv('datasets/Iris.csv')
df.head()
# -
df.columns
df.Species.unique()
# +
df_features = df[['SepalLengthCm', 'SepalWidthCm',
'PetalLengthCm', 'PetalWidthCm']]
# -
# Visualisation des types de fleurs, selon un découpage de variables par paires, en utilisant l'outil pairplot de seaborn.
#
# Vous pouvez consulter la documentation sur : https://seaborn.pydata.org/generated/seaborn.pairplot.html
# Main flowers species visualization
import seaborn as sns
from matplotlib import pyplot as plt
sns.pairplot(df.drop("Id", axis=1), hue="Species") #, diag_kind=False
plt.show()
# +
# Visualisation des valeurs de distribution des principales dimensions de plantes
df.drop("Id", axis=1).boxplot(by="Species", figsize=(12, 6))
plt.show()
# -
# ### Clustering par l'algorithme K-means
#
#
# On remarque que la classe (Species) est une chaine de caractères. Pour pouvoir représenter cette information en un tableau ou shéma, il faut transformer ces valeurs en des valeurs entiers.
# Pour voir qulles sont les types de fleurs
df['Species'].unique()
# On peut réaliser cette opération en utilisant le __Label Encoder__ comme suit :
#targetss = df['Species'].ravel()
# +
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
df['Species'] = le.fit_transform(df['Species'])
df_labels = df['Species']
df_labels
# -
from sklearn.cluster import KMeans
km = KMeans(n_clusters=3, random_state=10)
km.fit(df_features)
km.labels_
# +
#Visualisation des clusters
plt.scatter(df_features.PetalLengthCm, df_features.PetalWidthCm)
# -
colormap=np.array(['Red','green','blue'])
#Visualisation des clusters réels
plt.scatter(df_features.PetalLengthCm, df_features.PetalWidthCm,
c=colormap[df_labels],s=40)
plt.title('Clustering réel')
#Visualisation des clusters prédits
plt.scatter(df_features.PetalLengthCm, df_features.PetalWidthCm,
c=colormap[km.labels_],s=40)
plt.title('Clustering prédit')
# Si on veut visualiser le clustering avec les centroids :
# +
centroids = km.cluster_centers_
plt.scatter(centroids[:, 2], centroids[:, 3],
marker='x', s=169, linewidths=3,
color='orange', zorder=10)
plt.scatter(df_features.PetalLengthCm, df_features.PetalWidthCm,
c=colormap[km.labels_],s=40)
plt.title('Clustering prédit')
# -
# #### Evaluation du Clustering
# +
# Run the Kmeans algorithm and get the index of data points clusters
sse = []
list_k = list(range(1, 10))
for k in list_k:
km = KMeans(n_clusters=k)
km.fit(df_features)
sse.append(km.inertia_)
# Plot sse against k
plt.figure(figsize=(6, 6))
plt.plot(list_k, sse, '-o')
plt.xlabel(r'Number of clusters *k*')
plt.ylabel('Sum of squared distance');
# -
# +
from sklearn.metrics import silhouette_samples, silhouette_score
range_n_clusters = [2, 3, 4, 5, 6]
for n_clusters in range_n_clusters:
#for n_clusters in range_n_clusters:
clusterer = KMeans (n_clusters=n_clusters)
preds = clusterer.fit_predict(df_features)
centers = clusterer.cluster_centers_
score = silhouette_score (df_features, preds, metric='euclidean')
print ("For n_clusters = {}, silhouette score is {})".format(n_clusters, score))
# -
# #### Visualisation 3D des clusters
# Si on veut visaliser en 3D les clusters
# +
from mpl_toolkits.mplot3d import Axes3D
X = df.drop("Id", axis=1).drop("Species", axis=1).values
y = df_labels
#centers = [[1, 1], [-1, -1], [1, -1]]
centers = [[0, 0], [0, 0], [0, 0]]
# Plot the ground truth
fig = plt.figure(1, figsize=(5, 4))
plt.clf()
ax = Axes3D(fig, rect=[0, 0, .95, 1], elev=48, azim=134)
plt.cla()
for name, label in [('Iris-setosa', 0),
('Iris-versicolour', 1),
('Iris-virginica', 2)]:
ax.text3D(X[y == label, 3].mean(),
X[y == label, 0].mean() + 1.5,
X[y == label, 2].mean(), name,
horizontalalignment='center',
bbox=dict(alpha=.5, edgecolor='w', facecolor='w'))
# Reorder the labels to have colors matching the cluster results
y = np.choose(y, [1, 2, 0]).astype(np.float)
ax.scatter(X[:, 3], X[:, 0], X[:, 2], c=y)
ax.set_xlabel('Petal width')
ax.set_ylabel('Sepal length')
ax.set_zlabel('Petal length')
plt.show()
# -
# ## B. La classification supervisée :
# C’est l’opération qui permet de placer chaque individu de la population dans une classe parmi l’ensemble des classes préétablies, en suivant un processus d’apprentissage supervisé.
# le choix de la classe d’un individu dépend de ses caractéristiques.
# - Algorithme KNN (K Nearest Neighbors)
# - Arbre de décision
# - Machine à vecteurs de support (SVM)
# - Réseau de neurones
# - ...
# ### Problématique:
# #### Données :
# - Une liste d’exemples X {1..n} caractérisés par un ensemble d’attributs P.
# - Un ensemble C de classes préétablies.
# - Les caractéristiques d'un nouvel exemple «newX».
# #### Question :
# - Quelle est la classe appropriée à «newX» ?
#
# ### Jeu de données :
# Le jeu de données Iris a été utilisé à l''article classique de Fisher, publié en 1936, intitulé "L'utilisation de plusieurs mesures dans des problèmes taxonomiques", est également disponible dans le référentiel UCI Machine Learning.
#
# Il comprend trois espèces d’iris de 50 échantillons chacune, ainsi que des propriétés propres à chaque fleur. Une espèce de fleur est séparable linéairement des deux autres, mais les deux autres ne sont pas séparables linéairement l'une de l'autre.
#
# Les colonnes de cet ensemble de données sont:
#
# Id
# Longueur du Sépale Cm
# Largeur du Sépale Cm
# Longueur du Pétale cm
# Largeur du Pétale Cm
# Espèce : classe : Iris Setosa, Iris Versicolor ou Iris Virginica.
#
# Un échantillon : (4.9,3.6,1.4,0.1, “Iris-setosa”)
# ### A. importation des librairies
# Avec Pandas on peut manipuler lire (et/ou écrire) nos jeux de données, généralement avec une extension .csv
# +
# importation des lib
import pandas as pd
# -
# ### B. Importation des données
# Avec la fonction read_csv de Pandas: on peut mettre dans notre dataframe le contenu du fichier csv, en indiquant comme paramètre (1: le chemin ou la source où se trouve le fichier csv, 2: les séparateurs entre les valeurs dans notre cas ces des vergules) en troisième position, un paramètre facultatif pour spécifier le type d'encodage de notre fichier exemple encoding ="UTF8".
df = pd.read_csv('datasets/Iris.csv')
df.head()
# ### QUESTION 1
# Quelle est la moyenne de la longueure des petales de la setosa ?
# ### Reponse 1
# +
# Il y a plein de manière d'écrire cette commande
rep = df[df["Species"]=='Iris-setosa'].PetalLengthCm.mean()
#df[df.Outcome==1].SkinThickness.mean()
#df[df["Outcome"]==1]["SkinThickness"].mean()
rep
# -
# ## QUESTION 2
# Quelle est la longueure maximale des sepales de la setosa ?
# +
rep = df[df["Species"]=='Iris-setosa'].SepalLengthCm.max()
rep
# -
# ### C. Statistiques descriptives élémentaires
# Lire les informations sur nos données (Types d'attributs, valeurs manquantes...) Pandas nous permet de voir les informations sur notre benchmark exemple: avec dataframe.info() il nous affiche tout les attributs de notre fichier avec le type de donnée et le nombre de valeurs de chaque colonne
# dataframe.columns permet de citer les noms de toutes les colonnes
df.info() #donner les infos de notre data frame
#
# On peut supprimer la colonne ID :
#
# df.drop('Id',axis=1,inplace=True)
#
# #dropping the Id column as it is unecessary, axis=1 specifies that it should be column wise, inplace =1 means the changes should be reflected into the dataframe
#
#
# ### D. préparation des données
# Dans cette étape nous déterminons les attributs choisis pour l'entrainement et nous définissons l'attribut "classe" de notre benchmark
# définir les attraibuts qui nous intéréssent
df_features = df[['SepalLengthCm', 'SepalWidthCm', 'PetalLengthCm', 'PetalWidthCm' ]]
# définir l'attribut classe
df_labels = df[['Species']]
df['Species'].unique()
# Si on veut schématiser la distribution des classes, il suffit de faire appel à la libraire seaborn, en suite définir l'attribut concerné
# +
import seaborn as sns
# schématiser la distribution des classes
sns.countplot(df['Species'])
# -
# ### E. Transformer la colonne des classes en labels numériques
df_labelss=df['Species']#.ravel()
df_labelss
# +
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
df['Species'] = le.fit_transform(df['Species'])
df_labels = df['Species']
df_labels
# -
# ## F. Diviser le dataset en données d'entrainement et données de teste
# Ceci est réalisable avec sklearn qui permet de prendre aléatoirement des données de teste à partir du benchmark et laisser le reste pour l'apprentissage.
# La fonction train_test_split(param1,param2,param3,param4) prends 4 paramétres:
# le premier dédié à l'ensemble d'entrainement, le deuxième à l'ensemble de teste, le troisième c'est le paramètre du % de l'ensemble de test (généralement entre 15 et 40%),
#
# le 4 ème paramétre (facultatif) pour spécifier quel type de fonction random utiliser:
# si vous utilisez random_state = some_number, vous pouvez garantir que la sortie de Run 1 sera égale à la sortie de Run 2, c'est-à-dire que votre split sera toujours le même. Peu importe ce que le nombre réel random_state est 42, 0, 21, ... L'important est que chaque fois que vous utilisez 42, vous obtiendrez toujours la même sortie la première fois que vous faites la division. Ceci est utile si vous voulez des résultats reproductibles, par exemple dans la documentation, afin que tout le monde puisse toujours voir les mêmes nombres lors de l'exécution des exemples.
#
# Cette fonction retourne 4 sorties:
# La 1ere est le sous-ensembles aléatoires d'entrainement
# La 2éme est le vecteur de leurs labels (leurs classes).
# La 3ème est le sous-ensemble aléatoire pour le teste.
# La 4ème est le vecteur de leurs labels (leurs classes).
#
#
from sklearn.model_selection import train_test_split
#decouper le data set en 30% pour test et 70% pour train
X_train, X_test, y_train, y_test = train_test_split(df_features,
df_labels, test_size=0.4,
random_state=42)
# .shape permet de savoir la dimension d'un ensemble.
print('x_train shape:', X_train.shape) # .shape permet de voir la
print('x_test shape:', X_test.shape)
print('y_train shape:', y_train.shape)
print('y_test shape:', y_test.shape)
X_train.shape[0]
# ### Méthode des K plus proches Voisins ( K nearest neigbors)
from sklearn.neighbors import KNeighborsClassifier # le classifieur
# +
# Definir l'algorithme que je veux utiliser (KNN) avec le paramètre k=3
mon_knn = KNeighborsClassifier(n_neighbors=3)
#fitting : Lancer l'apprentissage ( données,labels)
mon_knn.fit(X_train, y_train)#.values.ravel())
# Evaluer l'entrainement de mon modèle
train_score = mon_knn.score(X_train, y_train)
print('train score = ',train_score )
# +
print('---- L Ensemble de test ----- \n',X_test)
#ypred : contient les prédictions de l'ensemble de teste
ypred = mon_knn.predict(X_test)
print('---- Les classes prédites par mon Algo ----- \n',ypred)
print('---- Les classes réelles ----- \n',y_test)
# -
# ### Evaluation du modèle
# #### A. Accuracy :
# Documentation sur accuracy_score de sk-learn ici : https://scikit-learn.org/stable/modules/generated/sklearn.metrics.accuracy_score.html?highlight=accuracy%20score#sklearn.metrics.accuracy_score
from sklearn.metrics import accuracy_score # Evaluation
print ('KNN accuracy score')
print (accuracy_score(y_test, ypred))
# #### B. Par Validation Croisée :
# Documentation sur la validation croisée de sk-learn ici : https://scikit-learn.org/stable/modules/cross_validation.html
# +
from sklearn.model_selection import cross_val_score
scores = cross_val_score(mon_knn, X_train, y_train, cv=5)
#scores = cross_val_score(mon_knn, df_features, df_labels, cv=5)
scores
# -
print("par validation croisée: " , scores.mean())
# #### C. Recall, Precision et F-score:
# documentation sur recall/ precision
# https://scikit-learn.org/stable/modules/generated/sklearn.metrics.recall_score.html?highlight=recall%20score#sklearn.metrics.recall_score
from sklearn.metrics import f1_score, precision_score, recall_score # Evaluation
print ('KNN recall score')
print (recall_score(y_test, ypred, average=None))
print ('KNN precision score')
print (precision_score(y_test, ypred,average=None))
print ('f1 score')
print (f1_score(y_test, ypred,average=None))
# Documentation sur la F-score : https://scikit-learn.org/stable/modules/generated/sklearn.metrics.f1_score.html?highlight=f1%20score#sklearn.metrics.f1_score
# #### D. Par matrice de confusion
from sklearn.metrics import confusion_matrix
print(confusion_matrix(y_test, ypred))
# +
# Function to plot confusion matrix
import matplotlib.pyplot as plt
import itertools
import numpy as np
def plot_confusion_matrix(cm, classes, normalize=False, title=' confusion matrix ', cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
"""
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Predict the values from the validation dataset
Y_pred = mon_knn.predict(X_test)
# Convert predictions classes to one hot vectors
#Y_pred_classes = np.argmax(Y_pred , axis = 1)
# Convert validation observations to one hot vectors
Y_true = y_test#np.argmax(y_test,axis = 1)
# compute the confusion matrix
confusion_mtx = confusion_matrix(Y_true, Y_pred)
# plot the confusion matrix
plot_confusion_matrix(confusion_mtx, classes = range(3))
# -
# ## Méthode des arbres de décision
# Je vous invite à consulter la documentation détaillée de cette méthode sur le site de sklearn : https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html?highlight=treeclassifier#sklearn.tree.DecisionTreeClassifier
# +
# De la meme manière que pour le KNN
#importer l'algorithme tree
from sklearn import tree
clf = tree.DecisionTreeClassifier(max_depth=3)
#fitting : Lancer l'apprentissage ( données,labels)
clf.fit(X_train, y_train)
# Evaluer l'entrainement de mon modèle
train_score = clf.score(X_train, y_train)
print('train score = ',train_score )
# +
#ypred : contient les prédictions de l'ensemble de teste
ypred = clf.predict(X_test)
print ('Decision tree accuracy score')
print (accuracy_score(y_test, ypred))
# -
# #### Ploter mon arbre de decision
#tree.DecisionTreeClassifier(max_depth=3)
tree.plot_tree(clf.fit(df_features, df_labels),max_depth=5)
# #### Une autre manière de schématiser un arbre de decision:
from sklearn.tree.export import export_text
from sklearn import tree
algo_tree = tree.DecisionTreeClassifier(max_depth=3)
algo_tree = algo_tree.fit(df_features, df_labels)
r = export_text(clf , feature_names = ['SepalLengthCm', 'SepalWidthCm', 'PetalLengthCm', 'PetalWidthCm' ])
print(r)
# +
ypred = clf.predict(X_test)
print ('Tree accuracy score')
print (accuracy_score(y_test, ypred))
# +
confusion_mtx = confusion_matrix(Y_true, ypred)
# plot the confusion matrix
plot_confusion_matrix(confusion_mtx, classes = range(3))
# -
confusion_mtx
# ### Exercice :
# En se basant sur ce notebook :
# - Ajouter un code qui cherche les meilleurs paramètres pour chaque méthode. ( vous pouvez utiliser gridsearch)
# - Ajouter d'autre méthodes de classification à ce notebook ( exmple: Naive Bayes, SVM, Random Forest, Réseaux de neurones multi-couches ... etc)
# - Evaluer toutes vos méthodes par validation croisée (nbr de paquet = 5).
|
1-Apprentissage_supervise_et_non_supervise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit (system)
# name: python3
# ---
# # Working with Text Data
# This notebook is about string operations in Pandas Series.
# +
# # !pip install numpy
# # !pip install pandas
# -
import pandas as pd
import numpy as np
s = pd.Series(['Tommy ', '<NAME>', 'John\n',
'ALBER@T', np.nan, '1234','SteveSmith', 34])
s
# **lower()**: Converts strings in the Series/Index to lower case.
s.str.lower()
# **upper()**: Converts strings in the Series/Index to upper case.
s.str.upper()
# **swapcase()**: swaps the case lower/upper.
s.str.swapcase()
# **islower()**: checks whether all characters in each string in the Series/Index in lower case or not. Returns Boolean
s.str.islower()
s.str.lower().str.islower()
s
# **isupper()**: checks whether all characters in each string in the Series/Index in upper case or not. Returns Boolean.
s.str.isupper()
# **isnumeric()**: checks whether all characters in each string in the Series/Index are numeric. Returns Boolean.
s.str.isnumeric()
# **len()**: Computes String length().
s.str.len()
# **strip()**: Helps strip whitespace (including newline) from each string in the Series/index from both the sides.
#
# Observe 'John\n' was changed o 'John'
s.str.strip()
# **split()**: Splits each string with the given pattern. The result is a list for each row
s.str.split(' ')
# + tags=[]
for r in s.str.split(' '):
if type(r) == list:
print('list with ',len(r),'elements',r)
else: print(r)
# -
# **cat(sep='')**: concatenates the series/index elements with given separator
s = pd.Series(['Tom ',' John','<NAME>','123'])
s.str.cat(sep='_')
# **contains(pattern)**: returns a Boolean value True for each element if the substring contains in the element, else False
s.str.contains(' ')
# **replace(a,b)**: replaces the value a with the value b.
s.str.replace(' ','_')
# **repeat(value)**: repeats each element with specified number of times.
s.str.repeat(2)
s.str.repeat(5)
# Observe that the lenght of the Serie is the same:
# + tags=[]
print(len(s))
print(len(s.str.repeat(5)))
# -
# What changes is the lenght of the elements:
# + tags=[]
print(len(s[0]))
print(len(s.str.repeat(5)[0]))
# -
# **count(pattern)**: returns count of appearance of pattern in each element.
s.str.count('o')
# **startswith(pattern)**: returns true if the element in the Series/Index starts with the pattern.
s.str.startswith(' ')
s.str.startswith('w')
s.str.startswith('W')
s.str.lower().str.startswith('w')
# **endswith(pattern)**: returns true if the element in the Series/Index ends with the pattern.
s.str.endswith(' ')
# **find(pattern)**: returns the first position of the first occurrence of the pattern. It returns -1 if the string is not found.
s.str.find('2')
s.str.find('ll')
# **findall(pattern)**: returns a list of all occurrence of the pattern.
s.str.findall('ll')
s = pd.Series(['red','orange','yellow','green','blue'])
s.str.find('e')
s.str.findall('e')
s.str.endswith('e')
# **get_dummies()**: returns the DataFrame with One-Hot Encoded values.
country = pd.Series(['USA','Colombia','Ecuador',
'Rep. Dominicana','Puerto Rico'])
country.str.get_dummies()
sex = pd.Series(['Male','Female'])
sex.str.get_dummies()
|
04-Introduction to Pandas/05-Working with Text Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Conditional Probability Activity & Exercise
# + [markdown] deletable=true editable=true
# Below is some code to create some fake data on how much stuff people purchase given their age range.
#
# It generates 100,000 random "people" and randomly assigns them as being in their 20's, 30's, 40's, 50's, 60's, or 70's.
#
# It then assigns a lower probability for young people to buy stuff.
#
# In the end, we have two Python dictionaries:
#
# "totals" contains the total number of people in each age group.
# "purchases" contains the total number of things purchased by people in each age group.
# The grand total of purchases is in totalPurchases, and we know the total number of people is 100,000.
#
# Let's run it and have a look:
# + deletable=true editable=true
from numpy import random
import numpy as np
random.seed(0)
totals = {20:0, 30:0, 40:0, 50:0, 60:0, 70:0}
purchases = {20:0, 30:0, 40:0, 50:0, 60:0, 70:0}
totalPurchases = 0
for _ in range(100000):
ageDecade = random.choice([20, 30, 40, 50, 60, 70])
purchaseProbability = 0.5
totals[ageDecade] += 1
if (random.random() < purchaseProbability):
totalPurchases += 1
purchases[ageDecade] += 1
# + deletable=true editable=true
totals
# + deletable=true editable=true
purchases
# + deletable=true editable=true
totalPurchases
# + [markdown] deletable=true editable=true
# Let's play with conditional probability.
#
# First let's compute P(E|F), where E is "purchase" and F is "you're in your 30's". The probability of someone in their 30's buying something is just the percentage of how many 30-year-olds bought something:
# + deletable=true editable=true
PEF = float(purchases[30]) / float(totals[30])
print('P(purchase | 30s): ' + str(PEF))
# + [markdown] deletable=true editable=true
# P(F) is just the probability of being 30 in this data set:
# + deletable=true editable=true
PF = float(totals[30]) / 100000.0
print("P(30's): " + str(PF))
# + [markdown] deletable=true editable=true
# And P(E) is the overall probability of buying something, regardless of your age:
# + deletable=true editable=true
PE = float(totalPurchases) / 100000.0
print("P(Purchase):" + str(PE))
# + [markdown] deletable=true editable=true
# If E and F were independent, then we would expect P(E | F) to be about the same as P(E). But they're not; PE is 0.45, and P(E|F) is 0.3. So, that tells us that E and F are dependent (which we know they are in this example.)
#
# What is P(E)P(F)?
# + deletable=true editable=true
print("P(30's)P(Purchase)" + str(PE * PF))
# + [markdown] deletable=true editable=true
# P(E,F) is different from P(E|F). P(E,F) would be the probability of both being in your 30's and buying something, out of the total population - not just the population of people in their 30's:
# + deletable=true editable=true
print("P(30's, Purchase)" + str(float(purchases[30]) / 100000.0))
# + [markdown] deletable=true editable=true
# P(E,F) = P(E)P(F), and they are pretty close in this example. But because E and F are actually dependent on each other, and the randomness of the data we're working with, it's not quite the same.
#
# We can also check that P(E|F) = P(E,F)/P(F) and sure enough, it is:
# + deletable=true editable=true
print((purchases[30] / 100000.0) / PF)
# + [markdown] deletable=true editable=true
# ## Your Assignment
# + [markdown] deletable=true editable=true
# Modify the code above such that the purchase probability does NOT vary with age, making E and F actually independent.
#
# Then, confirm that P(E|F) is about the same as P(E), showing that the conditional probability of purchase for a given age is not any different than the a-priori probability of purchase regardless of age.
#
# + deletable=true editable=true
|
ConditionalProbabilityExercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:analysis27]
# language: python
# name: conda-env-analysis27-py
# ---
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ### Import CMIP5 from the module and start a session
# + [markdown] deletable=true editable=true slideshow={"slide_type": "subslide"}
# The latest ARCCSSive stable version is available from the conda **analysis27** environment
# Anyone can load them both from raijin and the remote desktop.
#
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
# ! module use /g/data3/hh5/public/modules
# ! module load conda/analysis27
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# The database location is saved in the **$CMIP5_DB** environment variable. This is defined automatically if you have loaded ARCCSSive from conda/analysis27.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
# ! export CMIP5_DB=sqlite:////g/data1/ua6/unofficial-ESG-replica/tmp/tree/cmip5_raijin_latest.db
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# Import **CMIP5** from the module and use the *method* **connect()** to open a connection to the database.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
from ARCCSSive import CMIP5
db=CMIP5.connect()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "notes"}
# Opening a connection creates a **session object** (in this case *db*). A *session* manages all the comunication with the database and contains all the objects which you’ve loaded or associated with it during its lifespan. Every query to the database is run through the *session*.
# There are a number of helper functions for common operations:
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
db.models()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# models() return all the models recorded in the database,
# experiments(), variables(), mips() produce similar lists for each respective field
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ### Perform a simple search
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# To perform a search you can use the **outputs( )** function.
# **outputs( )** is a 'shortcut' to perform a session.query on the Instances table.
# The following example shows all the input arguments you can use, the order doesn't matter and you can omit any of them.
#
# > db.outputs( column-name='value', ... )
#
# will return all the rows for the Instances table in the database.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results=db.outputs(variable='tas',experiment='historical',mip='Amon',model='MIROC-ESM-CHEM',ensemble='r1i1p1')
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# You can check how many *instances* your search returned by using the *query* method **count()**
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results.count()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "notes"}
# In this case we defined every possible constraint for the table and hence we get just one instance.
# This should always be the case, if you use all the five attributes, because every *instance* is fully defined by these and each *instance* is unique.
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# We can loop through the instances returned by the search and access their attributes and their *children* ( i.e. related versions and files) attributes.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
for o in results:
print(o.model,o.variable,o.ensemble)
print()
print("drstree path is " + str(o.drstree_path()))
for v in o.versions:
print()
print('version', v.version)
print('dataset-id', v.dataset_id)
print('is_latest', v.is_latest, 'checked on', v.checked_on)
print()
print(v.path)
for f in v.files:
print(f.filename, f.tracking_id)
print(f.md5, f.sha256)
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ### Navigate through search results
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# Let's have a better look at **results**
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results=db.outputs(variable='tas',experiment='historical',mip='Amon',model='MIROC-ESM-CHEM',ensemble='r1i1p1')
type(results)
# + [markdown] deletable=true editable=true slideshow={"slide_type": "notes"}
# **results** is a *query object* but as we saw before we can loop through it as we do with a list.
# In this particular case we have only one *instance* returned in results, but we still need to use an index to access it.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
type(results[0])
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# A useful attribute of an *instance* is **versions**, this is a list of all the versions associated to that particular instance.
# From a database point of view these are all the rows in the **Versions table** which are related to that particular instance.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results[0].versions
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# We have two versions available for this *instance*, we can loop through them and retrieve their attributes:
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
for o in results:
for v in o.versions:
print()
print(v.version)
print()
print(v.path)
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# If we want to get only the *latest version*, we can use the **latest( )** method of the Instance class.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results[0].latest()[0].version
# + [markdown] deletable=true editable=true slideshow={"slide_type": "subslide"}
# As you might have noticed **latest( )** returns a list of Version objects rather than only one.
# This is because there might be different copies of the same version, downloaded from different servers.
# Currently the database lists all of them so that if you used one rather than the other in the past you can still find it.
# There are plans though to keep just one copy per version to facilitate the collection management and save storage resources.
#
# Other methods available for the Instances table (*objects*) are:
#
# # + **filenames( )**
# # + **drstree_path( )**
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results[0].filenames()
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results[0].drstree_path()
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
% ls -l /g/data1/ua6/DRSv2/CMIP5/CCSM4/rcp45/day/atmos/r1i1p1/tas/latest
# + [markdown] deletable=true editable=true slideshow={"slide_type": "notes"}
# #### !!Warning!!
# In most cases you can use directly the drstee_path() method to get to the files, but it can be useful to find all the available versions.
# For example if you want to make sure that a new version hasn't been added recently, DRSv2 it is updated only once a week.
# Or if you find that the version linked by the DRSv2 is incomplete, there might be another copy of the same version.
# We hope eventually to be able to have just one copy for each version and all of them clearly defined.
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ### Filter search results
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# We can refine our results by using the SQLalchemy **filter( )** function.
#
# We will use the attributes ( or columns ) of the database tables as *constraints*.
# So, first we need to import the tables definitions from ARCCSSive.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
from ARCCSSive.CMIP5.Model import Instance, Version, VersionFile
#print(type(Instance))
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# We can also import the **unique( )** function. This function will give us all the possible values we can use to filter over a particular attribute.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
from ARCCSSive.CMIP5.other_functions import unique
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# Let's do a new query
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results=db.outputs(variable='tas',experiment='rcp45',mip='day')
results.count()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# We would like to filter the results by ensemble, so we will use **unique( )** to get all the possible ensemble values.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
ensembles=unique(results,'ensemble')
print(ensembles)
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# **unique( results, 'attribute' )** takes two inputs:
# * results is a *query object* on the Instances table, for example what is returned by the db.outputs( ) function
# * 'attribute' is a string defining a particular attribute or column of the Instances table, for example 'model'
#
# **unique( )** lists all the distinct values returned by the query for that particular attribute.
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# Now that we know all the ensembles values, let's choose one to filter our results.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
r6i1p1_ens=results.filter(Instance.ensemble == 'r6i1p1')
print( r6i1p1_ens.count() )
unique(r6i1p1_ens,'ensemble')
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# We used the **==** equals operator to select all the r61i1p1 ensembles.
# If we wanted all the "r6i1p#" ensembles regardless of their physics (p) value we could have used the **like** operator.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
r6i1_ens=results.filter(Instance.ensemble.like('r6i1p%'))
print( r6i1_ens.count() )
unique(r6i1_ens,'ensemble')
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# If we want to search two variables at the same time we can leave the variable constraints out of the query inputs,
# and then use **filter** with the **in_** operator to select them.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results=db.outputs(ensemble='r1i1p1',experiment='rcp45',mip='day')\
.filter(Instance.variable.in_(['tasmin','tasmax']))
results.count()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# As you can see filter can follow directly the query, i.e. the **outputs( )** function.
# In fact, you can refine a query with how many successive filters as you want.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results=db.outputs(ensemble='r1i1p1',experiment='rcp45',mip='day')\
.filter(Instance.variable.in_(['tasmin','tasmax']))\
.filter(Instance.model.like('%ESM%'))
results.count()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ### Using the search results to open the files
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# Once we have found the instances and versions we want to use we can use their path to find the files and work with them.
# First we load numpy and the netcdf module.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
import numpy as np
from netCDF4 import MFDataset
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# All you need to open a file is the location, this is stored in the Versions table in the database as *path*.
# Alternatively you can use the *drstree* path, that is returned by the Instance *drstree_path( )* method.
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# Let's define a simple function that reads a variable from a file and calculate its maximum value.
# We will use MFDataset( ) from the netcDF4 module to open all the netcdf files in the input path as one aggregated file.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
def var_max(var,path):
''' calculate max value for variable '''
# MFDataset will open all netcdf files in path as one aggregated file
print(path+"/*.nc")
# open the file
nc=MFDataset(path+"/*.nc",'r')
# read the variable from file into a numpy array
data = nc.variables[var][:]
# close the file
nc.close()
# return the maximum
return np.max(data)
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# Now we perform a search, loop through the results and pass the Version path attribute to the var_max( ) function
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results=db.outputs(ensemble='r1i1p1',experiment='rcp45',mip='day').filter(Instance.model.like('MIROC%'))\
.filter(Instance.variable.in_(['tas','pr']))
print(results.count())
for o in results[:2]:
var = o.variable
for v in o.versions:
path=str(v.path)
varmax=var_max(var,path)
print()
print('Maximum value for variable %s, version %s is %d' % (var, v.version, varmax))
# + [markdown] deletable=true editable=true slideshow={"slide_type": "notes"}
# NB if you pass directly *v.path* value you get an error because the databse return unicode string, so you need to use the str( ) function to convert to a normal string.
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# ### How to integrate ARCCSSive in your python script
# + [markdown] deletable=true editable=true slideshow={"slide_type": "slide"}
# In the previous example we simply looped through the results returned by the search as they were and passed them to a function that opened the files.
# But what if we want to do something more complex?
# Let's say that we want to pass two variables to a function and do it for every model/ensemble that has both of them for a fixed experiment and mip
# Mostly users would somehow loop over the drstree path, doing something like:
# > cd /g/data1/ua6/DRSv2/CMIP5
# > list all models and save in model_list
# > for model in model_list:
# > list all eavailable ensembles and save in ensemble_list
# > for ensemble in ensemble_list:
# > call_function(var1_path, var2_path)
#
# Using ARCCSSIve we can do the same using the **unique( )** function to return the list of all available models/ensembles.
# Let's start from defining a simple function that calculates the difference bewteen the values of two variables.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
def vars_difference(var1,path1,var2,path2):
''' calculate difference between the mean of two variables '''
# open the files and read both variables
nc1=MFDataset(path1+"/*.nc",'r')
data1 = nc1.variables[var1][:]
nc1.close()
nc2=MFDataset(path2+"/*.nc",'r')
data2 = nc2.variables[var2][:]
nc2.close()
# return the difference between the two means
return np.mean(data2) - np.mean(data1)
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# Now let's do the another search and get tasmin and tasmax
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
results=db.outputs(ensemble='r1i1p1',experiment='rcp45',mip='Amon').filter(Instance.model.like('MIROC%'))\
.filter(Instance.variable.in_(['tasmin','tasmax']))
results.count()
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# Get the list of distinct models and ensembles using unique
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
models=unique(results,'model')
ensembles=unique(results,'ensemble')
# + [markdown] deletable=true editable=true slideshow={"slide_type": "fragment"}
# Now we loop over the models and the ensembles, for each model-ensemble combination we call the function if we have an instance for both variables.
# + deletable=true editable=true slideshow={"slide_type": "fragment"}
for mod in models:
for ens in ensembles:
# we filter twice the reuslts, using the model and ensemble values plus one the variable at the time
tasmin_inst=results.filter(Instance.model==mod, Instance.ensemble==ens, Instance.variable=='tasmin').first()
tasmax_inst=results.filter(Instance.model==mod, Instance.ensemble==ens, Instance.variable=='tasmax').first()
# we check that both filters returned something and call the function if they did
if tasmax_inst and tasmin_inst:
tasmin_path=tasmin_inst.latest()[0].path
tasmax_path=tasmax_inst.latest()[0].path
diff=vars_difference('tasmin',str(tasmin_path),'tasmax',str(tasmax_path))
print('Difference for model %s and ensemble %s is %d' % (mod, ens, diff))
# + [markdown] deletable=true editable=true slideshow={"slide_type": "notes"}
# **NB** we used **first( )** after the filter because we know we should be getting back either 1 instance or None. We cannot use **one( )** because that would return an error if it can't find anything.
# Also we should have checked that we are using the same versions for both variables rather than just getting the latest!
#
# This is just an attempt to replicate the way we use drstree but when you get more familiar with the module and with SQLalchemy you can set up more sophysticated searches.
|
examples/arccssive_training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
def plot_bar_charts(res_tab, ordered=False):
def preproc_name(method, reg, score):
name = method[0]
if score not in ['MD', 'MP']:
name += f'+{score[0]}'
if reg != '-':
name += f'+{reg[0]}'
return name
datasets = np.unique([col[0] for col in list(res_tab.columns[3:])])
metrics = np.unique([col[1] for col in list(res_tab.columns[3:])])
for dataset in datasets:
n_methods = len(res_tab['Method'].values)
fig = plt.figure(figsize = (n_methods, 5))
gs = fig.add_gridspec(1, len(metrics))
for i, metric in enumerate(metrics):
means = np.array([float(value.split('±')[0]) for value in res_tab[(dataset, metric)].values])
stds = np.array([float(value.split('±')[1]) for value in res_tab[(dataset, metric)].values])
methods = res_tab['Method'].values
regs = res_tab['Reg. Type'].values
scores = res_tab['UE Score'].values
names = np.array([preproc_name(m,r,s) for m,r,s in zip(methods, regs, scores)])
ax = fig.add_subplot(gs[0, i])
cmap = matplotlib.cm.get_cmap('Spectral')
colors = []
for i in range(cmap.N):
rgb = cmap(i)[:3]
colors.append(matplotlib.colors.rgb2hex(rgb))
colors = colors[::len(colors) // len(names)]
x_pos = np.array(list(range(len(names))))
if ordered:
order = np.argsort(means)[::-1]
else:
order = x_pos
ax.bar(x_pos, means[order], yerr=stds[order], width=0.8, align='center', alpha=1, color=colors, edgecolor='black', ecolor='black', capsize=6)
ax.set_ylabel(f'{metric.upper()}')
ax.set_xticks(x_pos)
ax.set_xticklabels(x_pos[order])
ax.tick_params(labelsize=8)
ax.set_title(f'{metric.upper()} for {dataset}')
ax.yaxis.grid(True)
patches = [matplotlib.patches.Patch(color=v, label=f'{x}. {k}') for x, k, v in zip(x_pos, names, colors)]
plt.tight_layout()
plt.legend(handles=patches, loc='center', fontsize=10, bbox_to_anchor=(-0.1, -0.16), ncol=4, edgecolor='black')
plt.savefig(f'../../new_{dataset}.pdf', bbox_inches='tight')
plt.savefig(f'../../new_{dataset}.png', bbox_inches='tight')
plt.show()
res_tab = pd.read_csv('../../new_conll2003.csv', header=[0, 1])
res_tab = res_tab.iloc[[1,10,19,22,28,31,32,39,41,43]]
res_tab = res_tab[res_tab.columns[[0,1,2,4,5,7,8]]]
plot_bar_charts(res_tab, ordered=False)
res_tab = pd.read_csv('../../deberta_all_glue.csv', header=[0, 1])
mrpc_tab = res_tab[res_tab.columns[[0,1,2,4,5]]]
mrpc_tab = mrpc_tab.iloc[[4,23,27,28,29,33,54,58,63,62]]
plot_bar_charts(mrpc_tab, ordered=False)
cola_tab = res_tab[res_tab.columns[[0,1,2,7,8]]]
cola_tab = cola_tab.iloc[[10,23,27,28,29,39,54,58,63,62]]
plot_bar_charts(cola_tab, ordered=False)
sst2_tab = res_tab[res_tab.columns[[0,1,2,10,11]]]
sst2_tab = sst2_tab.iloc[[13,23,27,28,29,39,58,63,62]]
plot_bar_charts(sst2_tab, ordered=False)
res_tab = pd.read_csv('../../deberta_all_conll2003.csv', header=[0, 1])
table_final = res_tab[res_tab.columns[[0,1,2,4,5,7,8]]]
table_final = table_final.iloc[[19,22,1,10,58,28,63,61]].reset_index(drop=True)
plot_bar_charts(table_final, ordered=False)
sst2_tab = sst2_tab.reset_index(drop=True)
# +
sst2_tab.loc[0, 'Reg. Type'] = '-'
sst2_tab.loc[0, 'UE Score'] = 'PV'
sst2_tab.loc[0, ('SST-2', 'rcc-auc')] = '17.04±2.72'
sst2_tab.loc[0, ('SST-2', 'rpp')] = '1.14±0.21'
sst2_tab.loc[1, 'Reg. Type'] = '-'
sst2_tab.loc[1, 'UE Score'] = 'SMP'
sst2_tab.loc[1, ('SST-2', 'rcc-auc')] = '13.12±3.27'
sst2_tab.loc[1, ('SST-2', 'rpp')] = '0.88±0.17'
sst2_tab.loc[2, ('SST-2', 'rcc-auc')] = '12.16±1.93'
sst2_tab.loc[2, ('SST-2', 'rpp')] = '0.83±0.11'
sst2_tab.loc[3, 'Reg. Type'] = 'CER'
sst2_tab.loc[3, ('SST-2', 'rcc-auc')] = '12.90±3.55'
sst2_tab.loc[3, ('SST-2', 'rpp')] = '0.87±0.23'
sst2_tab.loc[4, ('SST-2', 'rcc-auc')] = '10.89±1.25'
sst2_tab.loc[4, ('SST-2', 'rpp')] = '0.75±0.06'
sst2_tab.loc[6, ('SST-2', 'rcc-auc')] = '13.43±1.84'
sst2_tab.loc[6, ('SST-2', 'rpp')] = '0.87±0.08'
sst2_tab.loc[7, 'Method'] = 'SR'
sst2_tab.loc[7, ('SST-2', 'rcc-auc')] = '16.68±2.92'
sst2_tab.loc[7, ('SST-2', 'rpp')] = '1.11±0.24'
sst2_tab.loc[8, ('SST-2', 'rcc-auc')] = '18.07±6.11'
sst2_tab.loc[8, ('SST-2', 'rpp')] = '1.23±0.41'
# -
sst2_tab = sst2_tab.iloc[[0,1,2,3,4,6,7,8]]
plot_bar_charts(sst2_tab, ordered=False)
df = pd.DataFrame({'Method':['baseline', 'MC dropout', 'Deep Ensemble'],
'UE Score':['MP', 'SMP', 'SMP'],
'0%':['43.3±0.2', '43.8±0.2', '44.8±0.2'],
'5%':['44.7±0.3', '45.4±0.3', '46.5±0.2'],
'10%':['46.4±0.3', '47.0±0.3', '48.2±0.3'],
'20%':['49.4±0.1', '50.2±0.4', '51.4±0.3'],
'30%':['52.9±0.3', '53.6±0.4', '54.8±0.3'],
'40%':['56.6±0.3', '57.5±0.5', '58.8±0.3'],})
df
print(str(df.to_latex(index=False)).replace('±', '$\pm$'))
|
src/exps_notebooks/bar_chart.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# default_exp callback.experimental
# -
# # Experimental Callbacks
#
# > Miscellaneous experimental callbacks for timeseriesAI.
#export
import torch.multiprocessing
torch.multiprocessing.set_sharing_strategy('file_system')
#export
from fastai.callback.all import *
from tsai.imports import *
from tsai.utils import *
from tsai.data.preprocessing import *
from tsai.data.transforms import *
from tsai.models.layers import *
from tsai.callback.MVP import *
# ## Gambler's loss: noisy labels
# +
#export
class GamblersCallback(Callback):
"A callback to use metrics with gambler's loss"
def after_loss(self): self.learn.pred = self.learn.pred[..., :-1]
def gambler_loss(reward=2):
def _gambler_loss(model_output, targets):
outputs = torch.nn.functional.softmax(model_output, dim=1)
outputs, reservation = outputs[:, :-1], outputs[:, -1]
gain = torch.gather(outputs, dim=1, index=targets.unsqueeze(1)).squeeze()
doubling_rate = (gain + reservation / reward).log()
return - doubling_rate.mean()
return
# +
from tsai.data.external import *
from tsai.data.core import *
from tsai.models.InceptionTime import *
from tsai.models.layers import *
from tsai.learner import *
from fastai.metrics import *
from tsai.metrics import *
X, y, splits = get_UCR_data('NATOPS', return_split=False)
tfms = [None, TSCategorize()]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits)
dls = TSDataLoaders.from_dsets(dsets.train, dsets.valid, bs=[64, 128])
loss_func = gambler_loss()
learn = ts_learner(dls, InceptionTime(dls.vars, dls.c + 1), loss_func=loss_func, cbs=GamblersCallback, metrics=[accuracy])
learn.fit_one_cycle(1)
# -
# ## Uncertainty-based data augmentation
#export
class UBDAug(Callback):
r"""A callback to implement the uncertainty-based data augmentation."""
def __init__(self, batch_tfms:list, N:int=2, C:int=4, S:int=1):
r'''
Args:
batch_tfms: list of available transforms applied to the combined batch. They will be applied in addition to the dl tfms.
N: # composition steps (# transforms randomly applied to each sample)
C: # augmented data per input data (# times N transforms are applied)
S: # selected data points used for training (# augmented samples in the final batch from each original sample)
'''
self.C, self.S = C, min(S, C)
self.batch_tfms = L(batch_tfms)
self.n_tfms = len(self.batch_tfms)
self.N = min(N, self.n_tfms)
def before_fit(self):
assert hasattr(self.loss_func, 'reduction'), "You need to pass a loss_function with a 'reduction' attribute"
self.red = self.loss_func.reduction
def before_batch(self):
if self.training:
with torch.no_grad():
setattr(self.loss_func, 'reduction', 'none')
for i in range(self.C):
idxs = np.random.choice(self.n_tfms, self.N, False)
x_tfm = compose_tfms(self.x, self.batch_tfms[idxs], split_idx=0)
loss = self.loss_func(self.learn.model(x_tfm), self.y).reshape(-1,1)
if i == 0:
x2 = x_tfm.unsqueeze(1)
max_loss = loss
else:
losses = torch.cat((max_loss, loss), dim=1)
x2 = torch.cat((x2, x_tfm.unsqueeze(1)), dim=1)
x2 = x2[np.arange(x2.shape[0]).reshape(-1,1), losses.argsort(1)[:, -self.S:]]
max_loss = losses.max(1)[0].reshape(-1,1)
setattr(self.loss_func, 'reduction', self.red)
x2 = x2.reshape(-1, self.x.shape[-2], self.x.shape[-1])
if self.S > 1: self.learn.yb = (torch_tile(self.y, 2),)
self.learn.xb = (x2,)
def __repr__(self): return f'UBDAug({[get_tfm_name(t) for t in self.batch_tfms]})'
# +
from tsai.models.utils import *
X, y, splits = get_UCR_data('NATOPS', return_split=False)
tfms = [None, TSCategorize()]
dsets = TSDatasets(X, y, tfms=tfms, splits=splits)
dls = TSDataLoaders.from_dsets(dsets.train, dsets.valid, batch_tfms=[TSStandardize()])
model = build_ts_model(InceptionTime, dls=dls)
TS_tfms = [TSMagScale(.75, p=.5), TSMagWarp(.1, p=0.5), TSWindowWarp(.25, p=.5),
TSSmooth(p=0.5), TSRandomResizedCrop(.1, p=.5),
TSRandomCropPad(.3, p=0.5),
TSMagAddNoise(.5, p=.5)]
ubda_cb = UBDAug(TS_tfms, N=2, C=4, S=2)
learn = ts_learner(dls, model, cbs=ubda_cb, metrics=accuracy)
learn.fit_one_cycle(1)
# -
# # BatchLossFilter
#export
class BatchLossFilter(Callback):
""" Callback that selects the hardest samples in every batch representing a percentage of the total loss"""
def __init__(self, loss_perc=1., schedule_func:Optional[callable]=None):
store_attr()
def before_fit(self):
self.run = not hasattr(self, "gather_preds")
if not(self.run): return
self.crit = self.learn.loss_func
if hasattr(self.crit, 'reduction'): self.red = self.crit.reduction
def before_batch(self):
if not self.training: return
if self.schedule_func is None: loss_perc = self.loss_perc
else: loss_perc = self.loss_perc * self.schedule_func(self.pct_train)
if loss_perc == 1.: return
with torch.no_grad():
if hasattr(self.crit, 'reduction'): setattr(self.crit, 'reduction', 'none')
losses = self.crit(self.learn.model(self.x), self.y)
if losses.ndim == 2: losses = losses.mean(-1)
if hasattr(self.crit, 'reduction'): setattr(self.crit, 'reduction', self.red)
losses /= losses.sum()
idxs = torch.argsort(losses, descending=True)
cut_idx = max(1, torch.argmax((losses[idxs].cumsum(0) > loss_perc).float()))
idxs = idxs[:cut_idx]
self.learn.xb = tuple(xbi[idxs] for xbi in self.learn.xb)
self.learn.yb = tuple(ybi[idxs] for ybi in self.learn.yb)
def after_fit(self):
if hasattr(self.learn.loss_func, 'reduction'): setattr(self.learn.loss_func, 'reduction', self.red)
# # RandomWeightLossWrapper
# +
# export
class RandomWeightLossWrapper(Callback):
def before_fit(self):
self.run = not hasattr(self, "gather_preds")
if not(self.run): return
self.crit = self.learn.loss_func
if hasattr(self.crit, 'reduction'): self.red = self.crit.reduction
self.learn.loss_func = self._random_weight_loss
def _random_weight_loss(self, input: Tensor, target: Tensor) -> Tensor:
if self.training:
setattr(self.crit, 'reduction', 'none')
loss = self.crit(input, target)
setattr(self.crit, 'reduction', self.red)
rw = torch.rand(input.shape[0], device=input.device)
rw /= rw.sum()
non_red_loss = loss * rw
return non_red_loss.sum()
else:
return self.crit(input, target)
def after_fit(self):
if hasattr(self.crit, 'reduction'): setattr(self.crit, 'reduction', self.red)
self.learn.loss_func = self.crit
# -
# # BatchMasker
# +
# export
class BatchMasker(Callback):
""" Callback that applies a random mask to each sample in a training batch
Args:
====
r: probability of masking.
subsequence_mask: apply a mask to random subsequences.
lm: average mask len when using stateful (geometric) masking.
stateful: geometric distribution is applied so that average mask length is lm.
sync: all variables have the same masking.
variable_mask: apply a mask to random variables. Only applicable to multivariate time series.
future_mask: used to train a forecasting model.
schedule_func: if a scheduler is passed, it will modify the probability of masking during training.
"""
def __init__(self, r:float=.15, lm:int=3, stateful:bool=True, sync:bool=False, subsequence_mask:bool=True,
variable_mask:bool=False, future_mask:bool=False, schedule_func:Optional[callable]=None):
store_attr()
def before_fit(self):
self.run = not hasattr(self, "gather_preds")
if not(self.run): return
def before_batch(self):
if not self.training: return
r = self.r * self.schedule_func(self.pct_train) if self.schedule_func is not None else self.r
mask = create_mask(self.x, r=r, lm=self.lm, stateful=self.stateful, sync=self.sync,
subsequence_mask=self.subsequence_mask, variable_mask=self.variable_mask, future_mask=self.future_mask)
self.learn.xb = (self.xb[0].masked_fill(mask, 0),)
# In my tests, mask-based compensation doesn't seem to be important. ??
# mean_per_seq = (torch.max(torch.ones(1, device=mask.device), torch.sum(mask, dim=-1).unsqueeze(-1)) / mask.shape[-1])
# self.learn.xb = (self.xb[0].masked_fill(mask, 0) / (1 - mean_per_seq), )
# -
# # SamplerWithReplacement
# +
# export
class SamplerWithReplacement(Callback):
""" Callback that modify the sampler to select a percentage of samples and/ or sequence steps with replacement from each training batch"""
def before_fit(self):
self.run = not hasattr(self, "gather_preds")
if not(self.run): return
self.old_get_idxs = self.learn.dls.train.get_idxs
self.learn.dls.train.get_idxs = self._get_idxs
def _get_idxs(self):
dl = self.learn.dls.train
if dl.n==0: return []
if dl.weights is not None:
return np.random.choice(dl.n, dl.n, p=dl.weights)
idxs = Inf.count if dl.indexed else Inf.nones
if dl.n is not None: idxs = np.random.choice(dl.n,dl.n,True)
if dl.shuffle: idxs = dl.shuffle_fn(idxs)
return idxs
def after_fit(self):
self.learn.dls.train.get_idxs = self.old_get_idxs
# -
#hide
from tsai.imports import create_scripts
from tsai.export import get_nb_name
nb_name = get_nb_name()
# nb_name = "060_callback.experimental.ipynb"
create_scripts(nb_name);
|
nbs/060_callback.experimental.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (MAE6286)
# language: python
# name: py36-mae6286
# ---
# ###### Content under Creative Commons Attribution license CC-BY 4.0, code under MIT license © 2015 <NAME>, <NAME>, <NAME>
# # Relax and hold steady
# This is the fourth and last notebook of **Module 5** (*"Relax and hold steady"*), dedicated to elliptic PDEs. In the [previous notebook](https://nbviewer.jupyter.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/05_relax/05_03_Iterate.This.ipynb), we examined how different algebraic formulations can speed up the iterative solution of the Laplace equation, compared to the simplest (but slowest) Jacobi method. The Gauss-Seidel and successive-over relaxation methods both provide faster algebraic convergence than Jacobi. But there is still room for improvement.
#
# In this lesson, we'll take a look at the very popular [conjugate gradient](https://en.wikipedia.org/wiki/Conjugate_gradient_method) (CG) method.
# The CG method solves linear systems with coefficient matrices that are symmetric and positive-definite. It is either used on its own, or in conjunction with multigrid—a technique that we'll explore later on its own (optional) course module.
#
# For a real understanding of the CG method, there is no better option than studying the now-classic monograph by <NAME>: *"An introduction to the conjugate gradient method without the agonizing pain"* (1994). Here, we try to give you a brief summary to explain the implementation in Python.
# ### Test problem
# Let's return to the Poisson equation example from [Lesson 2](https://nbviewer.jupyter.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/05_relax/05_02_2D.Poisson.Equation.ipynb).
#
# $$
# \begin{equation}
# \nabla^2 p = -2\left(\frac{\pi}{2}\right)^2\sin\left( \frac{\pi x}{L_x} \right) \cos\left(\frac{\pi y}{L_y}\right)
# \end{equation}
# $$
#
# in the domain
#
# $$
# \left\lbrace \begin{align*}
# 0 &\leq x\leq 1 \\
# -0.5 &\leq y \leq 0.5
# \end{align*} \right.
# $$
#
# where $L_x = L_y = 1$ and with boundary conditions
#
# $$
# p=0 \text{ at } \left\lbrace
# \begin{align*}
# x&=0\\
# y&=0\\
# y&=-0.5\\
# y&=0.5
# \end{align*} \right.
# $$
#
# We will solve this equation by assuming an initial state of $p=0$ everywhere, and applying boundary conditions to relax via the Laplacian operator.
# ## Head in the right direction!
# Recall that in its discretized form, the Poisson equation reads,
#
# $$
# \frac{p_{i+1,j}^{k}-2p_{i,j}^{k}+p_{i-1,j}^{k}}{\Delta x^2}+\frac{p_{i,j+1}^{k}-2 p_{i,j}^{k}+p_{i,j-1}^{k}}{\Delta y^2}=b_{i,j}^{k}
# $$
#
# The left hand side represents a linear combination of the values of $p$ at several grid points and this linear combination has to be equal to the value of the source term, $b$, on the right hand side.
#
# Now imagine you gather the values $p_{i,j}$ of $p$ at all grid points into a big vector ${\bf p}$ and you do the same for $b$ using the same ordering. Both vectors ${\bf p}$ and ${\bf b}$ contain $N=nx*ny$ values and thus belong to $\mathbb{R}^N$. The discretized Poisson equation corresponds to the following linear system:
#
# $$
# \begin{equation}
# A{\bf p}={\bf b},
# \end{equation}
# $$
#
# where $A$ is an $N\times N$ matrix. Although we will not directly use the matrix form of the system in the CG algorithm, it is useful to examine the problem this way to understand how the method works.
#
# All iterative methods start with an initial guess, $\mathbf{p}^0$, and modify it in a way such that we approach the solution. This can be viewed as modifying the vector of discrete $p$ values on the grid by adding another vector, i.e., taking a step of magnitude $\alpha$ in a direction $\mathbf{d}$, as follows:
#
# $$
# \begin{equation}
# {\bf p}^{k+1}={\bf p}^k + \alpha {\bf d}^k
# \end{equation}
# $$
#
# The iterations march towards the solution by taking steps along the direction vectors ${\bf d}^k$, with the scalar $\alpha$ dictating how big a step to take at each iteration. We *could* converge faster to the solution if we just knew how to carefully choose the direction vectors and the size of the steps. But how to do that?
# ## The residual
# One of the tools we use to find the right direction to step to is called the *residual*. What is the residual? We're glad you asked!
#
# We know that, as the iterations proceed, there will be some error between the calculated value, $p^k_i$, and the exact solution $p^{exact}_i$. We may not know what the exact solution is, but we know it's out there. The error is:
#
# $$
# \begin{equation}
# e^k_i = p^k_i - p^{exact}_i
# \end{equation}
# $$
#
# **Note:** We are talking about error at a specific point $i$, not a measure of error across the entire domain.
#
# What if we recast the Poisson equation in terms of a not-perfectly-relaxed $\bf p^k$?
#
# $$
# \begin{equation}
# A \bf p^k \approx b
# \end{equation}
# $$
#
# We write this as an approximation because $\bf p^k \neq p$. To "fix" the equation, we need to add an extra term to account for the difference in the Poisson equation $-$ that extra term is called the residual. We can write out the modified Poisson equation like this:
#
# $$
# \begin{equation}
# {\bf r^k} + A \bf p^k = b
# \end{equation}
# $$
# ## The method of steepest descent
# Before considering the more-complex CG algorithm, it is helpful to introduce a simpler approach called the *method of steepest descent*. At iteration $0$, we choose an initial guess. Unless we are immensely lucky, it will not satisfy the Poisson equation and we will have,
#
# $$
# \begin{equation}
# {\bf b}-A{\bf p}^0={\bf r}^0\ne {\bf 0}
# \end{equation}
# $$
#
# The vector ${\bf r}^0$ is the initial residual and measures how far we are from satisfying the linear system. We can monitor the residual vector at each iteration, as it gets (hopefully) smaller and smaller:
#
# $$
# \begin{equation}
# {\bf r}^k={\bf b}-A{\bf p}^k
# \end{equation}
# $$
#
# We make two choices in the method of steepest descent:
#
# 1. the direction vectors are the residuals ${\bf d}^k = {\bf r}^k$, and
# 2. the length of the step makes the $k+1^{th}$ residual orthogonal to the $k^{th}$ residual.
#
# There are good (not very complicated) reasons to justify these choices and you should read one of the references to understand them. But since we want you to converge to the end of the notebook in a shorter time, please accept them for now.
#
# Choice 2 requires that,
#
# $$
# \begin{align}
# {\bf r}^{k+1}\cdot {\bf r}^{k} = 0 \nonumber \\
# \Leftrightarrow ({\bf b}-A{\bf p}^{k+1}) \cdot {\bf r}^{k} = 0 \nonumber \\
# \Leftrightarrow ({\bf b}-A({\bf p}^{k}+\alpha {\bf r}^k)) \cdot {\bf r}^{k} = 0 \nonumber \\
# \Leftrightarrow ({\bf r}^k-\alpha A{\bf r}^k) \cdot {\bf r}^{k} = 0 \nonumber \\
# \alpha = \frac{{\bf r}^k \cdot {\bf r}^k}{A{\bf r}^k \cdot {\bf r}^k}.
# \end{align}
# $$
#
# We are now ready to test this algorithm.
#
# To begin, let's import libraries and some helper functions and set up our mesh.
import numpy
from helper import l2_norm, poisson_2d_jacobi, poisson_solution
# +
# Set parameters.
nx = 101 # number of points in the x direction
ny = 101 # number of points in the y direction
xmin, xmax = 0.0, 1.0 # limits in the x direction
ymin, ymax = -0.5, 0.5 # limits in the y direction
Lx = xmax - xmin # domain length in the x direction
Ly = ymax - ymin # domain length in the y direction
dx = Lx / (nx - 1) # grid spacing in the x direction
dy = Ly / (ny - 1) # grid spacing in the y direction
# Create the gridline locations and the mesh grid.
x = numpy.linspace(xmin, xmax, num=nx)
y = numpy.linspace(ymin, ymax, num=ny)
X, Y = numpy.meshgrid(x, y)
# Create the source term.
b = (-2.0 * (numpy.pi / Lx) * (numpy.pi / Ly) *
numpy.sin(numpy.pi * X / Lx) *
numpy.cos(numpy.pi * Y / Ly))
# Set the initial conditions.
p0 = numpy.zeros((ny, nx))
# Compute the analytical solution.
p_exact = poisson_solution(x, y, Lx, Ly)
# -
# ### Time to code steepest descent!
#
# Let's quickly review the solution process:
#
# 1. Calculate the residual, $\bf r^k$, which also serves as the direction vector, $\bf d^k$
# 2. Calculate the step size $\alpha$
# 3. Update ${\bf p}^{k+1}={\bf p}^k + \alpha {\bf d}^k$
# ##### How do we calculate the residual?
#
# We have an equation for the residual above:
#
# $$
# \begin{equation}
# {\bf r}^k={\bf b}-A{\bf p}^k
# \end{equation}
# $$
#
# Remember that $A$ is just a stand-in for the discrete Laplacian, which taking $\Delta x=\Delta y$ is:
#
# $$
# \begin{equation}
# \nabla^2 p^k = \frac{-4p^k_{i,j} + \left(p^{k}_{i,j-1} + p^k_{i,j+1} + p^{k}_{i-1,j} + p^k_{i+1,j} \right)}{\Delta x^2}
# \end{equation}
# $$
# ##### What about calculating $\alpha$?
#
# The calculation of $\alpha$ is relatively straightforward, but does require evaluating the term $A{\bf r^k}$, but we just wrote the discrete $A$ operator above. You just need to apply that same formula to $\mathbf{r}^k$.
def poisson_2d_steepest_descent(p0, b, dx, dy,
maxiter=20000, rtol=1e-6):
"""
Solves the 2D Poisson equation on a uniform grid,
with the same grid spacing in both directions,
for a given forcing term
using the method of steepest descent.
The function assumes Dirichlet boundary conditions with value zero.
The exit criterion of the solver is based on the relative L2-norm
of the solution difference between two consecutive iterations.
Parameters
----------
p0 : numpy.ndarray
The initial solution as a 2D array of floats.
b : numpy.ndarray
The forcing term as a 2D array of floats.
dx : float
Grid spacing in the x direction.
dy : float
Grid spacing in the y direction.
maxiter : integer, optional
Maximum number of iterations to perform;
default: 20000.
rtol : float, optional
Relative tolerance for convergence;
default: 1e-6.
Returns
-------
p : numpy.ndarray
The solution after relaxation as a 2D array of floats.
ite : integer
The number of iterations performed.
conv : list
The convergence history as a list of floats.
"""
def A(p):
# Apply the Laplacian operator to p.
return (-4.0 * p[1:-1, 1:-1] +
p[1:-1, :-2] + p[1:-1, 2:] +
p[:-2, 1:-1] + p[2:, 1:-1]) / dx**2
p = p0.copy()
r = numpy.zeros_like(p) # initial residual
Ar = numpy.zeros_like(p) # to store the mat-vec multiplication
conv = [] # convergence history
diff = rtol + 1 # initial difference
ite = 0 # iteration index
while diff > rtol and ite < maxiter:
pk = p.copy()
# Compute the residual.
r[1:-1, 1:-1] = b[1:-1, 1:-1] - A(p)
# Compute the Laplacian of the residual.
Ar[1:-1, 1:-1] = A(r)
# Compute the step size.
alpha = numpy.sum(r * r) / numpy.sum(r * Ar)
# Update the solution.
p = pk + alpha * r
# Dirichlet boundary conditions are automatically enforced.
# Compute the relative L2-norm of the difference.
diff = l2_norm(p, pk)
conv.append(diff)
ite += 1
return p, ite, conv
# Let's see how it performs on our example problem.
# Compute the solution using the method of steepest descent.
p, ites, conv_sd = poisson_2d_steepest_descent(p0, b, dx, dy,
maxiter=20000,
rtol=1e-10)
print('Method of steepest descent: {} iterations '.format(ites) +
'to reach a relative difference of {}'.format(conv_sd[-1]))
# Compute the relative L2-norm of the error.
l2_norm(p, p_exact)
# Not bad! it took only *two* iterations to reach a solution that meets our exit criterion. Although this seems great, the steepest descent algorithm is not too good when used with large systems or more complicated right-hand sides in the Poisson equation (we'll examine this below!). We can get better performance if we take a little more care in selecting the direction vectors, $\bf d^k$.
# ## The method of conjugate gradients
# With steepest descent, we know that two **successive** jumps are orthogonal, but that's about it. There is nothing to prevent the algorithm from making several jumps in the same (or a similar) direction. Imagine you wanted to go from the intersection of 5th Avenue and 23rd Street to the intersection of 9th Avenue and 30th Street. Knowing that each segment has the same computational cost (one iteration), would you follow the red path or the green path?
# <img src="./figures/jumps.png" width=350>
# #### Figure 1. Do you take the red path or the green path?
# + [markdown] variables={"\\bf r}^{k+1} \\cdot {\\bf r}^{k+1": {}}
# The method of conjugate gradients reduces the number of jumps by making sure the algorithm never selects the same direction twice. The size of the jumps is now given by:
#
# $$
# \begin{equation}
# \alpha = \frac{{\bf r}^k \cdot {\bf r}^k}{A{\bf d}^k \cdot {\bf d}^k}
# \end{equation}
# $$
#
# and the direction vectors by:
#
# $$
# \begin{equation}
# {\bf d}^{k+1}={\bf r}^{k+1}+\beta{\bf d}^{k}
# \end{equation}
# $$
#
# where $\beta = \frac{{\bf r}^{k+1} \cdot {\bf r}^{k+1}}{{\bf r}^k \cdot {\bf r}^k}$.
#
# The search directions are no longer equal to the residuals but are instead a linear combination of the residual and the previous search direction. It turns out that CG converges to the exact solution (up to machine accuracy) in a maximum of $N$ iterations! When one is satisfied with an approximate solution, many fewer steps are needed than with any other method. Again, the derivation of the algorithm is not immensely difficult and can be found in Shewchuk (1994).
# -
# ### Implementing Conjugate Gradients
# + [markdown] variables={"\\bf r}^{k+1} \\cdot {\\bf r}^{k+1": {}}
# We will again update $\bf p$ according to
#
# $$
# \begin{equation}
# {\bf p}^{k+1}={\bf p}^k + \alpha {\bf d}^k
# \end{equation}
# $$
#
# but use the modified equations above to calculate $\alpha$ and ${\bf d}^k$.
#
# You may have noticed that $\beta$ depends on both ${\bf r}^{k+1}$ and ${\bf r}^k$ and that makes the calculation of ${\bf d}^0$ a little bit tricky. Or impossible (using the formula above). Instead we set ${\bf d}^0 = {\bf r}^0$ for the first step and then switch for all subsequent iterations.
#
# Thus, the full set of steps for the method of conjugate gradients is:
#
# Calculate ${\bf d}^0 = {\bf r}^0$ (just once), then
#
# 1. Calculate $\alpha = \frac{{\bf r}^k \cdot {\bf r}^k}{A{\bf d}^k \cdot {\bf d}^k}$
# 2. Update ${\bf p}^{k+1}$
# 3. Calculate ${\bf r}^{k+1} = {\bf r}^k - \alpha A {\bf d}^k$ $\ \ \ \ $(see <a href='#references'>Shewchuk (1994)</a>)
# 4. Calculate $\beta = \frac{{\bf r}^{k+1} \cdot {\bf r}^{k+1}}{{\bf r}^k \cdot {\bf r}^k}$
# 5. Calculate ${\bf d}^{k+1}={\bf r}^{k+1}+\beta{\bf d}^{k}$
# 6. Repeat!
# -
def poisson_2d_conjugate_gradient(p0, b, dx, dy,
maxiter=20000, rtol=1e-6):
"""
Solves the 2D Poisson equation on a uniform grid,
with the same grid spacing in both directions,
for a given forcing term
using the method of conjugate gradients.
The function assumes Dirichlet boundary conditions with value zero.
The exit criterion of the solver is based on the relative L2-norm
of the solution difference between two consecutive iterations.
Parameters
----------
p0 : numpy.ndarray
The initial solution as a 2D array of floats.
b : numpy.ndarray
The forcing term as a 2D array of floats.
dx : float
Grid spacing in the x direction.
dy : float
Grid spacing in the y direction.
maxiter : integer, optional
Maximum number of iterations to perform;
default: 20000.
rtol : float, optional
Relative tolerance for convergence;
default: 1e-6.
Returns
-------
p : numpy.ndarray
The solution after relaxation as a 2D array of floats.
ite : integer
The number of iterations performed.
conv : list
The convergence history as a list of floats.
"""
def A(p):
# Apply the Laplacian operator to p.
return (-4.0 * p[1:-1, 1:-1] +
p[1:-1, :-2] + p[1:-1, 2:] +
p[:-2, 1:-1] + p[2:, 1:-1]) / dx**2
p = p0.copy()
r = numpy.zeros_like(p) # initial residual
Ad = numpy.zeros_like(p) # to store the mat-vec multiplication
conv = [] # convergence history
diff = rtol + 1 # initial difference
ite = 0 # iteration index
# Compute the initial residual.
r[1:-1, 1:-1] = b[1:-1, 1:-1] - A(p)
# Set the initial search direction to be the residual.
d = r.copy()
while diff > rtol and ite < maxiter:
pk = p.copy()
rk = r.copy()
# Compute the Laplacian of the search direction.
Ad[1:-1, 1:-1] = A(d)
# Compute the step size.
alpha = numpy.sum(r * r) / numpy.sum(d * Ad)
# Update the solution.
p = pk + alpha * d
# Update the residual.
r = rk - alpha * Ad
# Update the search direction.
beta = numpy.sum(r * r) / numpy.sum(rk * rk)
d = r + beta * d
# Dirichlet boundary conditions are automatically enforced.
# Compute the relative L2-norm of the difference.
diff = l2_norm(p, pk)
conv.append(diff)
ite += 1
return p, ite, conv
# Compute the solution using the method of conjugate gradients.
p, ites, conv_cg = poisson_2d_conjugate_gradient(p0, b, dx, dy,
maxiter=20000,
rtol=1e-10)
print('Method of conjugate gradients: {} iterations '.format(ites) +
'to reach a relative difference of {}'.format(conv_cg[-1]))
# Compute the relative L2-norm of the error.
l2_norm(p, p_exact)
# The method of conjugate gradients also took two iterations to reach a solution that meets our exit criterion. But let's compare this to the number of iterations needed for the Jacobi iteration:
# Compute the solution using Jacobi relaxation.
p, ites, conv_jacobi = poisson_2d_jacobi(p0, b, dx, dy,
maxiter=40000,
rtol=1e-10)
print('Jacobi relaxation: {} iterations '.format(ites) +
'to reach a relative difference of {}'.format(conv_jacobi[-1]))
# For our test problem, we get substantial gains in terms of computational cost using the method of steepest descent or the conjugate gradient method.
# ## More difficult Poisson problems
# The conjugate gradient method really shines when one needs to solve more difficult Poisson problems. To get an insight into this, let's solve the Poisson problem using the same boundary conditions as the previous problem but with the following right-hand side,
#
# $$
# \begin{equation}
# b = \sin\left(\frac{\pi x}{L_x}\right) \cos\left(\frac{\pi y}{L_y}\right) + \sin\left(\frac{6\pi x}{L_x}\right) \cos\left(\frac{6\pi y}{L_y}\right)
# \end{equation}
# $$
# Modify the source term of the Poisson system.
b = (numpy.sin(numpy.pi * X / Lx) *
numpy.cos(numpy.pi * Y / Ly) +
numpy.sin(6.0 * numpy.pi * X / Lx) *
numpy.cos(6.0 * numpy.pi * Y / Ly))
maxiter, rtol = 40000, 1e-10
p, ites, conv = poisson_2d_jacobi(p0, b, dx, dy,
maxiter=maxiter, rtol=rtol)
print('Jacobi relaxation: {} iterations'.format(ites))
p, ites, conv = poisson_2d_steepest_descent(p0, b, dx, dy,
maxiter=maxiter,
rtol=rtol)
print('Method of steepest descent: {} iterations'.format(ites))
p, ites, conv = poisson_2d_conjugate_gradient(p0, b, dx, dy,
maxiter=maxiter,
rtol=rtol)
print('Method of conjugate gradients: {} iterations'.format(ites))
# Now we can really appreciate the marvel of the CG method!
# ## References
# <a id='references'></a>
# <NAME>. (1994). [An Introduction to the Conjugate Gradient Method Without the Agonizing Pain (PDF)](http://www.cs.cmu.edu/~quake-papers/painless-conjugate-gradient.pdf)
#
# <NAME>, [The Concept of Conjugate Gradient Descent in Python](http://ikuz.eu/2015/04/15/the-concept-of-conjugate-gradient-descent-in-python/)
# ---
# ###### The cell below loads the style of this notebook.
from IPython.core.display import HTML
css_file = '../../styles/numericalmoocstyle.css'
HTML(open(css_file, 'r').read())
|
lessons/05_relax/05_04_Conjugate.Gradient.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
# default_exp encoder
# -
# # Imports
#
# > API details.
#hide
from nbdev.showdoc import *
from fastcore.test import *
# +
#export
import pandas as pd # package for high-performance, easy-to-use data structures and data analysis
import re
import string
from fastai2.basics import *
from fastai2.text.all import *
from fastai2.callback.all import *
# -
#export
import os
if os.path.basename(os.path.normpath(os.getcwd())) == "projetos":
os.chdir(Path(os.getcwd())/"gquest_nbdev")
print(os.listdir("../data/gquest_data/"))
data_path = Path("../data/gquest_data/")
import pdb
# # Reading data
# +
#export
print('Reading data...')
train_data = pd.read_csv(data_path/'train/train.csv')
test_data = pd.read_csv(data_path/'test/test.csv')
sample_submission = pd.read_csv(str(data_path/'sample_submission.csv'))
print('Reading data completed')
# -
#export
print('Size of train_data', train_data.shape)
print('Size of test_data', test_data.shape)
print('Size of sample_submission', sample_submission.shape)
test_eq(train_data.shape,(6079,41))
test_eq(test_data.shape,(476,11))
test_eq(sample_submission.shape,(476,31))
train_data.head()
train_data.columns
sample_submission.columns
targets = list(sample_submission.columns[1:])
targets
text_columns=['question_title', 'question_body', 'question_user_name',
'question_user_page', 'answer', 'answer_user_name', 'answer_user_page',
'url', 'category', 'host']
train_data[targets].describe()
train_data['question_body'][0]
text_columns
df_tokenized,token_count=tokenize_df(train_data,text_columns)
df_tokenized.head()
vocab = make_vocab(token_count)
vocab
import pickle
with open(data_path/'vocab.pkl', 'wb') as vocab_file:
pickle.dump(vocab, vocab_file)
tfm = Numericalize(make_vocab(token_count))
splits = RandomSplitter()(df_tokenized)
splits
dsrc = DataSource(df_tokenized, [[attrgetter("text"), tfm]], splits=splits, dl_type=LMDataLoader)
bs,sl = 32,72
dbch = dsrc.databunch(bs=bs, seq_len=sl)
dbch.show_batch()
config = awd_lstm_lm_config.copy()
config.update({'input_p': 0.6, 'output_p': 0.4, 'weight_p': 0.5, 'embed_p': 0.1, 'hidden_p': 0.2})
model = get_language_model(AWD_LSTM, len(vocab), config=config)
opt_func = partial(Adam, wd=0.1, eps=1e-7)
cb_funcs = [partial(MixedPrecision, clip=0.1), partial(RNNTrainer, alpha=2, beta=1)]
learn = language_model_learner(dbch, AWD_LSTM, metrics=[accuracy, Perplexity()], path=data_path, opt_func = partial(Adam, wd=0.1)).to_fp16()
import torch
torch.cuda.is_available()
learn.freeze()
learn.fit_one_cycle(1, 1e-2, moms=(0.8,0.7,0.8))
learn.unfreeze()
learn.fit_one_cycle(4, 1e-3, moms=(0.8,0.7,0.8))
learn.fit_one_cycle(4, 1e-5)
learn.show_results()
learn.save_encoder('enc1')
from nbdev.export import notebook2script
notebook2script()
|
00_encoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:databases19]
# language: python
# name: conda-env-databases19-py
# ---
# # B+ Trees
class BPTree(object):
def __init__(self, internal_capacity, leaf_capacity):
self.internal_capacity = internal_capacity
self.leaf_capacity = leaf_capacity
self.root = InternalNode(internal_capacity, leaf_capacity)
def __repr__(self):
return "Root: " + str(self.root)
def split_root(self):
"""
Creates a new root node and places the split current rood node
as its children
"""
promote, new_right = self.root.split()
new_root = InternalNode(self.internal_capacity, self.leaf_capacity)
new_root.items.append(promote)
new_root.children.append(self.root)
new_root.children.append(new_right)
self.root = new_root
def insert(self, tup):
"""
Inserts tuple into B+Tree, handles splitting if root node
is over capacity
"""
self.root.insert(tup)
if self.root.over_capacity():
self.split_root()
def in_order_traversal(self):
""" Returns the tuples from the smallest to largest """
node = self.root
while type(node) is not LeafNode:
node = node.children[0]
while node is not None:
for tup in node.items:
yield tup
node = node.rsibling
def depth(self):
""" Computes the depth of the B+Tree """
d = 0
node = self.root
while type(node) is not LeafNode:
node = node.children[0]
d += 1
return d
class InternalNode(object):
def __init__(self, internal_capacity, leaf_capacity):
self.capacity = internal_capacity
self.leaf_capacity = leaf_capacity
self.items = []
self.children = []
def __repr__(self):
s = ''
s += 'Internal Node\n'
s += f'Capacity: {self.capacity}\n'
s += f'Items: {self.items}\n'
s += f'Children: ['
for child in self.children:
c = ''.join([' ' + x + '\n' for x in str(child).splitlines()])
s += f'\n{c}'
s = s.rstrip(',')
s += ']'
return s
def over_capacity(self):
""" Returns True if node is over capacity """
return len(self.items) > self.capacity
def insert(self, tup):
"""
insert handles insertion into both internal and leaf nodes by
calling the appropriate functions
"""
if len(self.children) == 0:
self.children.append(LeafNode(self.leaf_capacity))
if isinstance(self.children[0], LeafNode):
self.insert_into_leaf(tup)
elif isinstance(self.children[0], InternalNode):
self.insert_into_internal(tup)
else:
raise Exception(f"Children of type: {type(self.children[0])}")
def split(self):
"""
copes half of this internal node's items and children into a new internal
node - new_right
"""
new_right = InternalNode(self.capacity, self.leaf_capacity)
split_idx = self.capacity//2
promote = self.items[split_idx]
new_right.items = self.items[split_idx+1:]
new_right.children = self.children[split_idx+1:]
self.items = self.items[:split_idx]
self.children = self.children[:split_idx+1]
return promote, new_right
def insert_into_internal(self, tup):
"""
insert_into_internal inserts a tuple into a child internal node
it will also split that child node if it is over capacity
"""
# index of child internal node
child_idx = len(self.items)
for i, item in enumerate(self.items):
if tup < item:
child_idx = i
break
node = self.children[child_idx]
# insert into node
node.insert(tup)
if node.over_capacity():
promote, new_right = node.split()
self.items.insert(child_idx, promote)
self.children.insert(child_idx + 1, new_right)
def split_leaf(self, leaf, child_idx):
"""
splits a leaf into two nodes. Also inserts new split node as a child of
this current internal node
"""
new_right = leaf.split()
self.items.insert(child_idx, new_right.items[0])
self.children.insert(child_idx + 1, new_right)
def insert_into_leaf(self, tup):
""" inserts a tuple into appropriate child leaf node """
# index of child leaf node
child_idx = len(self.items)
for i, item in enumerate(self.items):
if tup < item:
child_idx = i
break
leaf = self.children[child_idx]
# insert tuple into leaf node
leaf.insert(tup)
# fix leaf node if it is over capacity
if leaf.over_capacity():
self.split_leaf(leaf, child_idx)
class LeafNode(object):
def __init__(self, capacity):
self.capacity = capacity
self.items = []
self.rsibling = None
def __repr__(self):
s = ''
s += 'Leaf Node\n'
s += f'Capacity: {self.capacity}\n'
s += f'Items: {self.items}\n'
return s
def over_capacity(self):
""" Returns True if node is over capacity """
return len(self.items) > self.capacity
def insert(self, tup):
"""
Checks if tup can be inserted. If it can, insert leaf node into items
and sort items. (easier than locating proper index to insert)
O(n log(n)) instead of O(n) so this method is fine for small n
"""
if tup in self.items:
raise Exception(f"Can not insert tuple, {tup}, already exists")
self.items.append(tup)
self.items.sort()
def split(self):
""" splits a leaf node in place """
# split items
new_right = LeafNode(self.capacity)
new_right.items = self.items[self.capacity//2 + 1:]
self.items = self.items[:self.capacity//2 + 1]
# change siblings
new_right.rsibling = self.rsibling
self.rsibling = new_right
return new_right
from random import shuffle
intn_cap = 5
leaf_cap = 3
BPT = BPTree(intn_cap, leaf_cap)
numbers = list(range(30))
shuffle(numbers)
for num in numbers:
BPT.insert(num)
BPT
BPT.depth()
in_ord_trav = list(BPT.in_order_traversal())
for i in range(len(in_ord_trav) - 1):
assert(in_ord_trav[i] < in_ord_trav[i+1])
|
B+ Tree.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.common.exceptions import TimeoutException
from selenium.common.exceptions import NoSuchElementException
from selenium.webdriver.common.keys import Keys
import time
import pandas as pd
from bs4 import BeautifulSoup
import re
def get_info(url):
driver = webdriver.Firefox()
driver.set_page_load_timeout(30)
driver.get(url)
bbsq = driver.find_elements_by_class_name('zsg-photo-card-info')
status =
price =
_bbsq = [x.text for x in bbsq]
# +
url = 'https://www.zillow.com/homes/for_sale/Milpitas-CA-95035/house,condo,apartment_duplex,townhouse_type/1-_baths/20000-_price/82-_mp/globalrelevanceex_sort/0_mmm/'
# -
info
|
src/.ipynb_checkpoints/Untitled3-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualize Neural Network
# +
import warnings
warnings.filterwarnings('ignore')
import os
import numpy as np
import pandas as pd
from csrank.callbacks import DebugOutput
from csrank import FATEObjectRanker, ObjectRankingDatasetGenerator, FETAObjectRanker
from csrank.tensorflow_util import configure_numpy_keras
from csrank.util import setup_logging
from keras.utils import plot_model
from keras import backend as K
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
import pandas as pd
from csrank.losses import *
from csrank.metrics import *
from sklearn.utils import check_random_state
from collections import OrderedDict
import logging
# -
# ## Defining constants for the experiments
# Initializing the variables for the experiment. Configuring the keras and tensorflow. Defining the parameters for dataset reader.
SUB_FOLDER = "gr_vis"
log_path = os.path.join(os.getcwd(), SUB_FOLDER, "gr.log")
configure_numpy_keras(seed=42)
setup_logging(log_path)
logger = logging.getLogger('Experiment')
# Generate the medoid dataset for evaluating the model
n_objects = 3
n_features = 2
n_train_instances = 100000
n_test_instances = 6
random_state = check_random_state(42)
params = {'n_train_instances': n_train_instances,
'n_test_instances': n_test_instances,
'n_features': n_features,
'n_objects': n_objects,
'random_state': random_state}
or_generator = ObjectRankingDatasetGenerator(**params)
X_train, Y_train, X_test, Y_test = or_generator.get_single_train_test_split()
n_instances, n_objects, n_features = X_train.shape
# Define the parameters for the FATEObjectRanker
epochs = 5
n_hidden_joint_units = 5
n_hidden_set_units = 7
n_hidden_joint_layers = 1
n_hidden_set_layers = 1
ranker_params = {"n_objects": n_objects,
"n_object_features": n_features,
"n_hidden_joint_layers" : n_hidden_joint_layers,
"n_hidden_set_layers" : n_hidden_set_layers,
"n_hidden_set_units" : n_hidden_set_units,
"n_hidden_joint_units" : n_hidden_joint_units,
"use_early_stopping": True}
logger.info("n_hidden_joint_units {} and n_hidden_set_units {}".format(n_hidden_joint_units, n_hidden_set_units))
logger.info("############################# With set layers ##############################")
# Create the model and fit the ranker on it to to check the visualization
gor = FETAObjectRanker(**ranker_params)
gor.fit(X_train, Y_train, epochs=epochs)
# Create the complete model and visualize it
SVG(model_to_dot(gor.model, show_shapes=True,
rankdir='LR').create(prog='dot', format='svg'))
# To store the model into a .png file
model_path = os.path.join(os.getcwd(), SUB_FOLDER, "completeModel.png")
plot_model(plot_model(gor.model,to_file=model_path,show_shapes=True, rankdir='LR'))
|
docs/notebooks/Visualize-NeuralNetwork.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear regression tutorial
#
#
# ##### improved from https://towardsdatascience.com/introduction-to-linear-regression-in-python-c12a072bedf0
#
#
# #### The basic idea
# The basic idea is that if we can fit a linear regression model to observed data, we can then use the model to predict any future values. For example, let’s assume that we have found from historical data that the price (P) of a house is linearly dependent upon its size (S) — in fact, we found that a house’s price is exactly 90 times its size. The equation will look like this:
#
# P = 90 * S
#
# [ Note that the unit of x, meters, cancels out
#
# P = 90 dollars/meters * S meters = 90 * S dollars
# ]
#
#
# With this model, we can then predict the cost of any house. If we have a house that is 1,500 square feet, we can calculate its price to be:
#
# P = 90*1500 = $135,000
#
# This concept is commonly taught in Algebra in the form of:
#
# $y = mx + b$, where m is the slope and it is equal to $ \Delta y $ / $ \Delta x $, and b is the y-intersect (or bias), the value y at x = 0.
#
# $y$ is the dependent variable, because it depends on x, meaning it varies with it. $x$ is the independent variable since its value is, in theory, independent from that of other variables.
#
# A value of $\beta > 0$ or $\beta<0$ implies that x and y are correlated.
#
# In statistical learning, the notation changes to $y = \beta_1 x + \beta_0 $, absorbing both parameters into the vector $ \beta = [m, b] = [ \beta_1, \beta_0] $, which is a cleaner notation to do math and to generalize the problem beyond a single independent variable, x. We will use the statistical learning notation for the tutorial for these reasons.
#
# However, perhaps the best way to think of a linear model is to think of m or $\beta_1$ as a weight as $\beta_0$ as a bias.
#
# $y = weight * x + bias$
#
#
#
# #### The model
#
# There are two kinds of variables in a linear regression model:
#
# The input or predictor variable is the variable(s) that help predict the value of the output variable. It is commonly referred to as X.
# The output variable is the variable that we want to predict. It is commonly referred to as Y.
# To estimate Y using linear regression, we assume the equation:
#
# $y_e = \beta_1 x + \beta_0 $
# where $y_e$ is the estimated or predicted value of Y based on our linear equation.
#
# Our goal is to find statistically significant values of the parameters $\beta_1$ and $ \beta_0 $ that minimise the difference between the true Y and our estimate $y_e$.
#
#
# This practical tutorial will show how to acomplish this using the library sci-kit learn. A basic machine learning primer follows the Tutorial. These sections introduce other important concepts such as model evaluation and crossvalidation.
#
# A description of a theoretical solution to to this problem can be found in the (Appendix), as well as a numerical solution coded from scratch. These sections set the stage for machine learning, and can help build useful intuitions on the subject since ML combines similar theoretical intuitions with efficient algorithms for numerical computations to solve these types of problems. A small machine learning appendix is also include. The appendix is absolutely not obligatory to understand and apply Linear regression and/or machine learning, which is why we begin with the Practical Tutorial straight away.
#
# # Practical Tutorial
# # Load Libraries
# +
# Load standard libraries
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
# Load simple Linear regression library
from sklearn.linear_model import LinearRegression
# -
# # Load and visualize data
# Import and display first five rows of advertising dataset
data = pd.read_csv('advertising.csv')
data.head()
# Plot Sales data against TV advertising spending
plt.scatter(data.TV, data.sales)
plt.title('Sales vs TV Ads Spending')
plt.xlabel('TV Ads Spending')
plt.ylabel('Sales')
# # Linear Regression of Sales onto TV Ads
# We apply the linear model
#
# $y = \beta_1 x + \beta_0 $
#
# to model sales as a function of TV ads spending, that is:
#
# $sales = \beta_1 TV + \beta_0 $
#
# Where $y = \beta_1 x$ represents the linear relationship between x and y, where parameter $\beta_1$ is the weight that needs to be applied to x to transform it into y, and $\beta_0$ $ is the baseline (e.g. the y-intersect or mean bias of the model).
#
# Ignoring the effects of any other variables, $ b_0 $ tells us how many sales occur when TV Ads spending is 0.
#
# Parameters $\beta_0 $ and $\beta_1$ are chosen such that the difference between the model output $y_{e} = \beta_1 x + \beta_0$ and the true value of $y_{true}$ is at its minimum value. This is done automatically by sci-kit learn using the Ordinary Least Squares algorithm (see Apendix).
# +
# Build linear regression model using TV as predictor
# Split data into predictors X and output Y
predictors = ['TV']
X = data[predictors]
y = data['sales']
# Initialise and fit model
lm = LinearRegression()
model = lm.fit(X, y)
# Print Coefficients
print(f'beta_0 = {model.intercept_}')
print(f'beta = {model.coef_}')
# -
# ### Sales = 0.0475*TV + 7.03
# Overlay the linear fit to the plot of sales vs tv spending
linear_prediction = model.predict(X)
plt.plot(data.TV, linear_prediction, 'r')
plt.scatter(data.TV, data.sales)
plt.title('Sales vs TV Ads Spending')
plt.xlabel('TV Ads Spending')
plt.ylabel('Sales')
# # Exercise
#
# ### Do a linear regression of sales onto Radio Ads Spending (radio), then plot the data and overlay the linear fit.
#
# ### Please rename the variable linear_prediction (e.g. linear_prediction_radio), all other variables can be overwritten.
#
# ##### (The variable linear_prediction is used later in the code for comparing the accuracy of different models)
#
# +
# Exercise Code
# -
# # Can we do better with a quadratic term in the model?
# ## $y = \beta_2 x^2 + \beta_1 x^1 + \beta_0 $
# +
# Build linear regression model using TV and TV^2 as predictors
# First we have to create variable TV^2, we simply add it to the dataframe data
data['TV2'] = data['TV']*data['TV']
predictors = ['TV', 'TV2']
X = data[predictors]
y = data['sales']
# Initialise and fit model
lm2 = LinearRegression()
model_2 = lm2.fit(X, y)
# Print Coefficients
print(f'beta_0 = {model_2.intercept_}')
print(f'betas = {model_2.coef_}')
# -
# ### Sales = -6.84693373e-05 * $TV^2$ + 6.72659270e-02*TV + 6.114
# Overlay the quadratic fit to the plot of sales vs tv spending
quadratic_prediction = model_2.predict(X)
plt.plot(data.TV, quadratic_prediction, '.r')
plt.scatter(data.TV, data.sales)
plt.title('Sales vs TV Ads Spending')
plt.xlabel('TV Ads Spending')
plt.ylabel('Sales')
# +
# We can visualize this in 3d
# This import registers the 3D projection
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(data.TV, data.TV2, quadratic_prediction)
ax.scatter(data.TV, data.TV2, data.sales)
ax.view_init(elev=60., azim=-30)
ax.set_xlabel('TV')
ax.set_ylabel('$TV^2$')
ax.set_zlabel('Sales')
# -
# The solution $\beta = [\beta_0, \beta_1, \beta_2] $ is a vector of dimension 3, which defines plane that can be visualized with a contour plot.
#
#
# Note: 2 non-intersecting points define a line in 2d space, 3 non-intersecting points define a plane in 3d space.
#
# Notice the model allows us to predict sales even for values of $TV$ and $TV^2$ not sampled in the available data.
# +
def f(x, x2):
return model.intercept_ + model_2.coef_[0]*x + model_2.coef_[1]*x2
x, x2 = np.meshgrid(data.TV, data.TV2)
Z = f(x, x2)
fig = plt.figure(figsize=(10,10))
ax = plt.axes(projection='3d')
ax.contour3D(x, x2, Z, 50, cmap='binary')
ax.scatter(data.TV, data.TV2, data.sales)
ax.view_init(elev=60., azim=-30)
ax.set_xlabel('TV')
ax.set_ylabel('$TV^2$')
ax.set_zlabel('Sales')
# -
# What are these betas? The $\beta$etas are often called weights, since they determine the weight of the relationship of the independent variable to each dependent variable. Mathematically, they are the slope of the dependent variable (sales) along the dimension of each variable. In 3 dimensions, as in the figure above, the higher the slope, the more tilted the plane is towards that axis.
#
# The $\beta_1$ for the model $y = \beta_1 TV + \beta_0$
#
# would be the same as $\beta_1$ in the model $y = beta_1TV+ \beta_2 radio + \beta_0$
#
# if and only if TV and radio are linearly independent (i.e. are not correlated).
#
# Since TV and TV^2 are highly correlated, the $\beta_1$ for the model $y = \beta_1 TV + \beta_0$
#
# is guaranteed to be different than the $\beta_1$ in the model $y = beta_1TV+ \beta_2 TV^2 + \beta_0$
np.corrcoef(data.TV, data.TV2)
# # Model Evaluation
#
# In order to choose a model, we need to compare their performance, in science and statistics it is common to use the 'goodness-of-fit' R2. Business users prefer other evaluation metrics such as the Mean Absolute Percentage Error, which is expressed in units of $y$.
#
# The coefficient of determination, denoted R2 or r2 and pronounced "R squared", is the proportion of the variance in the dependent variable that is predictable from the independent variable(s).
#
#
#
# $ R^{2}\equiv 1-{SS_{\rm {res}} \over SS_{\rm {tot}}} $,
#
# where
#
# $ SS_{\text{res}}=\sum _{i}(y_{i}-f_{i})^{2}=\sum _{i}e_{i}^{2} $
#
# is the sum of the residual squared errors, $e_i$.
#
# and
#
# $ SS_{\text{tot}}=\sum _{i}(y_{i}-{\bar {y}})^{2} $
#
# is proportional to the variance of the data
#
#
# $ \sigma^2 =\sum _{i}^{n}(y_{i}-{\bar {y}})^{2} / n$
#
# ### Bottom Line
#
# #### R2 = 1 - Sum of the Errors / Variability in the data
#
# #### The higher R2 the better the model fit. Best possible R2 score is 1.0 and it can be negative (because the model can be arbitrarily worse).
#
# #### A constant model that always predicts the expected value of y, disregarding the input features, would get a R^2 score of 0.0.
#
#
# +
from sklearn.metrics import r2_score
#r^2 (coefficient of determination) regression score function.
print(f'linear model = {r2_score(y, linear_prediction)}')
print(f'quadratic model = {r2_score(y, quadratic_prediction)}')
# -
# #### In business, it is best practice to report things in business units, so metrics like the mean absolute percentage error (MAPE) can be more useful than R2. When evaluating a model using the MAPE metric, the winnig model is the one with the lowest value.
# +
# Define MAPE evaluation metric
def mean_absolute_percentage_error(y_true, y_pred):
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
# Here the winning model is the one with the lower MAPE
print(f'linear model = {mean_absolute_percentage_error(y, linear_prediction)}')
print(f'quadratic model = {mean_absolute_percentage_error(y, quadratic_prediction)}')
# -
# # Multiple linear regression
#
# The problem naturally expands to the case of more independent variables , for example with 2 variables, $x_1$ and $ x_2$ , $y = \beta_0 + \beta_1*x_1 + \beta_1*x_2 $
#
# Luckily we can use the exact same code from the simple linear regression to perform a multiple linear regression.
#
# +
from scipy import stats
# Build linear regression model using TV as predictor
# Split data into predictors X and output Y
predictors = ['TV', 'radio']
# It is good practice to z-score predictors with different units
# such that they are all varying from approximately from -2 to 2 (z-distributed)
X = stats.zscore(data[predictors])
y = data['sales']
# Initialise and fit model
lm = LinearRegression()
model = lm.fit(X, y)
# Print Coefficients
print(f'beta_0 = {model.intercept_}')
print(f'betas = {model.coef_}')
# Produce a prediction with these 2 variables
mlr_prediction = model.predict(X)
# -
# Again we visualize the model fit in 3d.
#
# This type of visualization is useless when we have more than 2 independent variables.
#
# However in this case the 3d plot allows us to see that the model can predict sales even for values of $TV$ and $radio$ not sampled in the available data.
#
# +
def f(X, Y):
return model.intercept_ + model.coef_[0]*X + model.coef_[1]*Y
X, Y = np.meshgrid(stats.zscore(data.TV), stats.zscore(data.radio))
Z = f(X,Y)
fig = plt.figure(figsize=(10,10))
ax = plt.axes(projection='3d')
ax.contour3D(X, Y, Z, 50, cmap='binary')
ax.scatter(stats.zscore(data.TV),stats.zscore(data.radio),data.sales)
ax.view_init(elev=60., azim=-30)
ax.set_xlabel('TV')
ax.set_ylabel('$radio$')
ax.set_zlabel('Sales')
# +
# Lets compare the 3 models first with R2
print(f'linear model = {r2_score(y, linear_prediction)}')
print(f'quadratic model = {r2_score(y, quadratic_prediction)}')
print(f'multiple linear regression = {r2_score(y, mlr_prediction)}')
# +
# and now with MAPE
print(f'linear model = {mean_absolute_percentage_error(y, linear_prediction)}')
print(f'quadratic model = {mean_absolute_percentage_error(y, quadratic_prediction)}')
print(f'multiple linear regression = {mean_absolute_percentage_error(y, mlr_prediction)}')
# -
# # Exercise
# As an exercise, use TV, Radio, and Newspaper to predict sales using the linear model
# Code exercise
# # Machine Learning Primer
#
# ### Crossvalidation (Train/Test Split)
# In statistical learning and machine learning it is very common to split the data set into a training set that is used to fit the model, and a testing set that is used to evaluate the performance of the model on previously unseen data. This is called cross-validation, and it is one of the main tools we use to assure ourselves that the model is actually learning a relationship, rather than overfitting (i.e. memorizing what it has seen).
#
#
# It is best practice to choose the model with the best test set performance.
#
#
# +
from sklearn.model_selection import train_test_split
predictors = ['TV', 'radio', 'newspaper']
# It is good practice to z-score predictors with different units
X = data[predictors]
X_c = stats.zscore(X)
y = data['sales']
# Test set (test_size) is typically between 0.1 to 0.3 of the data
X_train, X_test, y_train, y_test = train_test_split(
X_c, y, test_size=0.2)
model = LinearRegression().fit(X_train, y_train)
print(f'Betas = {model.coef_}')
print(f'R2 Score = {model.score(X_test, y_test)}')
print(f'MAPE = {mean_absolute_percentage_error(y_test,model.predict(X_test))}')
# -
# ### Random Forest Regression
# ##### Default settings
#
# RandomForest and other tree-based methods such as XGBoost often perform very well right out of the box, without the need for any tuning by the user.
# +
from sklearn.ensemble import RandomForestRegressor
regr = RandomForestRegressor()
regr.fit(X_train, y_train)
print(f'Feat.Importance = {regr.feature_importances_}')
print(f'r2 = {r2_score(y_test, regr.predict(X_test) )}')
print(f'MAPE = {mean_absolute_percentage_error(y_test, regr.predict(X_test) )}')
# -
features = regr.fit(X_train, y_train).feature_importances_
feature_imp_df = pd.DataFrame({'Importance': features},
index=X.columns.ravel()).sort_values('Importance', ascending=False)
n=20
plt.figure(figsize=(15, 7))
feature_imp_df.head(n).plot(kind='bar')
plt.grid(True, axis='y')
plt.title('RF Feature Importance')
plt.hlines(y=0, xmin=0, xmax=n, linestyles='dashed');
# ### RandomForestRegressor often performs better than LinearRegression.
#
# ### Importantly, we did not clean up the data (i.e. remove outliers) before modeling. Outliers typically introduce strong biases on the weights ($\beta$), which can lead to bad predictions on test data. Thus, data cleaning is an important pre-processing step that should be considered when fitting linear models.
#
# ### In contrast, RandomForest is a lot more robust to outliers or missing data, and these do not need to be removed before modeling. This is because RandomForest isolates outliers in separate leafs covering small regions of the feature space, meaning they will not impact the mean of other leafs. This is one of the reasons that RandomForest is considered an "off-the-shelf" ready to use algorithm.
#
#
# ## Food for thought
#
#
# # The End
# # Linear regression Appendix
#
# Let's try to solve the simple linear model without using sci-kit learn!
# Build linear regression model using TV as predictor
# Split data into predictors X and output Y
X = data['TV']
y = data['sales']
# ### Ordinary Least Squares
#
# We want to find an approximation or estimate $y_e$ for variable $y$ which has the smallest error possible.
# This can be accomplished by minimize the residual sum of squared errors (RSSE). The squared term is so that positive and negative deviations from $y$ are penalized equally.
#
# residual sum of squared errors = $\sum _{i=1}^{n}(y_{i}-x_{i}^{\mathrm {T} }\beta)^{2}=(y-X\beta)^{\mathrm {T} }(y-X\beta)$
#
# We next show how numerical (trial and error) methods can be used in order to find the value of vector $\beta$ that produces the smallest sum of the squared values. However for the linear model it is actually posible to find an analytial solution (a formula) with a bit of calculus.
#
# We will skip the calculus, but highlight that since our goal is to minimize the RSSE, and this quantity depends on $\beta^2$, we know that we can always find a solution since RSSE is a quadratic equation (i.e. it is a parabolla, see numerical solution for a visual).
#
# ## Analytical Solution
#
# The bit of calculus involves taking the derivative of RSSE and setting it equal to zero in order to solve for $\beta$, but we skip the proof and jump to the result.
#
# It can be shown that $\beta_1 = Cov(X, Y) / Var(X) $.
#
# Using the computed value of $\beta_1$, we can then find $\beta_0 = \mu_y - \beta_1 * \mu_x $
#
# , where $\mu_x $ and $\mu_y$ are the means of x and y.
# +
# We skip the calculus, and just show that the analytical formulas give the same result as sci-kit learn
# Calculate the mean of X and y
xmean = np.mean(X)
ymean = np.mean(y)
# Calculate the terms needed for the numator and denominator of beta
xycov = (X - xmean) * (y - ymean)
xvar = (X - xmean)**2
# Calculate beta and alpha
beta = xycov.sum() / xvar.sum()
alpha = ymean - (beta * xmean)
print(f'alpha = {alpha}')
print(f'beta = {beta}')
# -
# # Sci-kit learn
# +
# Compare this to what we got using sci-kit learn
predictors = ['TV']
X = data[predictors]
y = data['sales']
# Initialise and fit model
lm = LinearRegression()
model = lm.fit(X, y)
# Print Coefficients
print(f'beta_0 = {model.intercept_}')
print(f'beta = {model.coef_}')
# -
# # Numerical solution
# Let's iteratively take some brute-force guesses of the values of $\beta_1$ and $\beta_0$ and record their performance in terms of the sum of residual squared errors.
# +
# This can take a couple minutes, it's the worst algorithm one could write, but it gets the job done.
rss = []
beta1s = []
beta0s = []
# We cheat a bit narrowing our search for the optimal Betas on range 1 and range 2
range1 = np.linspace(-1,1,200)
range0 = np.linspace(0,10,200)
# Compute the RSS over all values of range 1 and range 0
for beta1 in range1:
for beta0 in range0:
rss.append(np.sum((y-(X['TV']*beta1+beta0))**2))
beta1s.append(beta1)
beta0s.append(beta0)
# -
# Print the solution
print(f'beta_0 = {beta0s[np.argmin(rss)]}')
print(f'beta = {beta1s[np.argmin(rss)]}')
# +
# We visualize the Betas and their corresponding RSS
from mpl_toolkits.mplot3d import Axes3D # noqa: F401 unused import
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(beta0s, beta1s, rss)
ax.scatter(beta0s[np.argmin(rss)], beta1s[np.argmin(rss)], np.min(rss), c = 'r', s = 50)
ax.set_xlabel('beta_0')
ax.set_ylabel('beta_1')
ax.set_zlabel('RSS')
ax.set_title(' Cost Function (RSS vs Betas)')
# -
# The Residual Sum of Squares has a concave shape by construction, since we defined it as a parabolla. This makes the RSS a good Cost Function to evaluate different model parameters in order to choose the ones that minimize mistakes because:
#
# 1) it is guaranteed to have a minimum
#
# 2) It is possible to compute this minimum using numerical methods that are much more efficient than the brute force approach we wrote. For example, say we start with completely Random weights, and compute the RSS. Then we change the weights a little (say, make one weight a little bigger) and produce a new prediction. If this prediction has a lower RSS than our previous guess, we can make the weight yet a little bigger and again see if RSS improves again. If we produce a worse prediction (higher RSS), we can modify the weights in the opposite direction (e.g. make the weight a little smaller). Applying this iterative process to all the weights until we cannot longer improve our RSS by more than some threshold value would eventually find us the minimum RSS.
#
# ##### This is the basic idea underlying Gradient decent, which is how machine learning alrorithms minimize their cost function (also known as an objective function).
#
# Gradient descent is an optimization algorithm used to minimize some function by iteratively moving in the direction of steepest descent as defined by the negative of the gradient. A gradient is a multidimensional derivative (i.e. a fancy name for a slope). A lot of research has been done to guarantee the learning algorithms go down the sleepest slope, 99% of data scientists, including myself, just take it for granted and use Adam because this is the default setting. Really, I just read a blog like this one to learn a bit more about it, not even a paper.
#
# https://machinelearningmastery.com/adam-optimization-algorithm-for-deep-learning/
#
# ##### There are other useful Cost Functions, but RSS (also known as mean squared error) is the one most commonly used for regression.
# 
# # Machine Learning Primer Appendix
# ### K-Fold Crossvalidation
#
#
# The training set can actually be sub-split into a train/test and evaluate the model fit more precisely (prevents bias).
#
# This process can be done automatically for you many times using cross_val_score.
# +
# ideally, k-fold crossvalidation is used during model parameter fitting on training data, and a test data for evaluation
from sklearn.model_selection import cross_val_score
scores = cross_val_score(model, X_train, y_train, scoring="r2", cv=5)
print(scores)
# -
# # Random Forest Regression
# ## (with k-fold crossvalidation and custom hyperparameters)
#
#
# These options will become more important as we have more input features.
#
# There are 2 crucial parts to machine learning:
#
# 1) Engineering good features for the model, and eliminating bad ones
#
# 2) Hyper-parameter tuning.
#
# 1 is a lot more important than 2 in my experience.
# +
from sklearn.model_selection import GridSearchCV
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(start = 10, stop = 100, num = 2)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(10, 110, num = 2)]
# Method of selecting samples for training each tree
bootstrap = [True, False]
# Create the random grid
param_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'bootstrap': bootstrap}
regr = RandomForestRegressor()
grid_regr = GridSearchCV(regr, param_grid, cv=5)
grid_regr.fit(X_train, y_train)
print(f'r2 = {r2_score(y_test, grid_regr.predict(X_test) )}')
print(f'MAPE = {mean_absolute_percentage_error(y_test, grid_regr.predict(X_test) )}')
|
LinearRegression/LinearRegression_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Plot Loop-Closure-Detection
#
# Plots statistics on loop closure detection as well as optimized trajectory RPE, APE and trajectory against ground truth.
# +
import yaml
import os
import copy
import pandas as pd
import numpy as np
import logging
log = logging.getLogger(__name__)
log.setLevel(logging.INFO)
if not log.handlers:
ch = logging.StreamHandler()
ch.setLevel(logging.INFO)
ch.setFormatter(logging.Formatter('%(levelname)s - %(message)s'))
log.addHandler(ch)
from evo.tools import file_interface
from evo.tools import plot
from evo.tools import pandas_bridge
from evo.core import sync
from evo.core import trajectory
from evo.core import metrics
from evo.core import transformations
from evo.core import lie_algebra as lie
# %matplotlib inline
# # %matplotlib notebook
import matplotlib.pyplot as plt
# -
# ## Data Locations
#
# Make sure to set the following paths.
#
# `vio_output_dir` is the path to the directory containing `output_*.csv` files obtained from logging a run of SparkVio.
#
# `gt_data_file` is the absolute path to the `csv` file containing ground truth data for the absolute pose at each timestamp of the dataset.
# Define directory to VIO output csv files as well as ground truth absolute poses.
vio_output_dir = "/home/sparklab/code/SparkVIO/output_logs/"
gt_data_file = "/home/sparklab/datasets/EuRoC/mh_04_difficult/mav0/state_groundtruth_estimate0/data.csv"
# +
def get_ape(data, metric):
""" Gets APE and APE statistics for two trajectories and a given pose_relation.
Args:
data: tuple of trajectories, the first being the reference trajectory
and the second being the estimated trajectory.
metric: a metrics.PoseRelation instance representing the pose relation
to use when computing APE.
Returns:
A metrics.APE instance containing the APE for both trajectories according
to the given metric.
"""
ape = metrics.APE(metric)
ape.process_data(data)
return ape
def plot_ape(x_axis, ape, size=(18,10), title=None):
""" Plots APE error against time for a given metrics.APE instance.
Args:
x_axis: An array-type of values for all the x-axis values (time).
rpe: A metrics.APE instance with pre-processed data.
size: A tuple optionally containing the size of the figure to be plotted.
"""
if title is None:
title = "APE w.r.t. " + ape.pose_relation.value
fig = plt.figure(figsize=size)
plot.error_array(fig, ape.error, x_array=x_axis, statistics=ape.get_all_statistics(),
name="APE", title=title, xlabel="$t$ (s)")
plt.show()
def get_rpe(data, metric):
""" Gets RPE and RPE statistics for two trajectories and a given pose_relation.
Args:
data: tuple of trajectories, the first being the reference trajectory
and the second being the estimated trajectory.
metric: a metrics.PoseRelation instance representing the pose relation
to use when computing RPE.
Returns:
A metrics.RPE instance containing the RPE for both trajectories according
to the given metric.
"""
# normal mode
delta = 1
delta_unit = metrics.Unit.frames
all_pairs = False
rpe = metrics.RPE(metric, delta, delta_unit, all_pairs)
rpe.process_data(data)
return rpe
def plot_rpe(x_axis, rpe, size=(18,10), title=None):
""" Plots RPE error against time for a given metrics.RPE instance.
Args:
x_axis: An array-type of values for all the x-axis values (time).
rpe: A metrics.RPE instance with pre-processed data.
size: A tuple optionally containing the size of the figure to be plotted.
"""
if title == None:
title = "RPE w.r.t. " + rpe.pose_relation.value
fig = plt.figure(figsize=size)
plot.error_array(fig, rpe.error, x_array=x_axis, statistics=rpe.get_all_statistics(),
name="RPE", title=title, xlabel="$t$ (s)")
plt.show()
def downsize_lc_df(df):
""" Remove all entries from a pandas DataFrame object that have '0' for the timestamp, which
includes all entries that do not have loop closures. Returns this cleaned DataFrame.
Args:
df: A pandas.DataFrame object representing loop-closure detections, indexed by timestamp.
Returns:
A pandas.DataFrame object with only loop closure entries.
"""
df = df[~df.index.duplicated()]
ts = np.array(df.index.tolist())
good_ts = ts[np.where(ts>0)]
res = df.reindex(index=good_ts)
return res
def convert_abs_traj_to_rel_traj_lcd(df, lcd_df, to_scale=True):
""" Converts an absolute-pose trajectory to a relative-pose trajectory.
The incoming DataFrame df is processed element-wise. At each kf timestamp (which is the
index of the DataFrame row) starting from the second (index 1), the relative pose
from the match timestamp to the query stamp is calculated (in the match-
timestamp's coordinate frame). This relative pose is then appended to the
resulting DataFrame.
The resulting DataFrame has timestamp indices corresponding to poses that represent
the relative transformation between the match timestamp and the query one.
Args:
df: A pandas.DataFrame object with timestamps as indices containing, at a minimum,
columns representing the xyz position and wxyz quaternion-rotation at each
timestamp, corresponding to the absolute pose at that time.
lcd_df: A pandas.DataFrame object with timestamps as indices containing, at a minimum,
columns representing the timestamp of query frames and the timestamps of the
match frames.
to_scale: A boolean. If set to False, relative poses will have their translation
part normalized.
Returns:
A pandas.DataFrame object with xyz position and wxyz quaternion fields for the
relative pose trajectory corresponding to the absolute one given in 'df', and
relative by the given match and query timestamps.
"""
rows_list = []
index_list = []
for i in range(len(lcd_df.index)):
match_ts = lcd_df.timestamp_match[lcd_df.index[i]]
query_ts = lcd_df.timestamp_query[lcd_df.index[i]]
try:
w_t_bi = np.array([df.at[match_ts, idx] for idx in ['x', 'y', 'z']])
w_q_bi = np.array([df.at[match_ts, idx] for idx in ['qw', 'qx', 'qy', 'qz']])
w_T_bi = transformations.quaternion_matrix(w_q_bi)
w_T_bi[:3,3] = w_t_bi
except:
print "Failed to convert an abs pose to a rel pose. Timestamp ", \
match_ts, " is not available in ground truth df."
continue
try:
w_t_bidelta = np.array([df.at[query_ts, idx] for idx in ['x', 'y', 'z']])
w_q_bidelta = np.array([df.at[query_ts, idx] for idx in ['qw', 'qx', 'qy', 'qz']])
w_T_bidelta = transformations.quaternion_matrix(w_q_bidelta)
w_T_bidelta[:3,3] = w_t_bidelta
except:
print "Failed to convert an abs pose to a rel pose. Timestamp ", \
query_ts, " is not available in ground truth df."
continue
index_list.append(lcd_df.index[i])
bi_T_bidelta = lie.relative_se3(w_T_bi, w_T_bidelta)
bi_R_bidelta = copy.deepcopy(bi_T_bidelta)
bi_R_bidelta[:,3] = np.array([0, 0, 0, 1])
bi_q_bidelta = transformations.quaternion_from_matrix(bi_R_bidelta)
bi_t_bidelta = bi_T_bidelta[:3,3]
if not to_scale:
norm = np.linalg.norm(bi_t_bidelta)
if norm > 1e-6:
bi_t_bidelta = bi_t_bidelta / np.linalg.norm(bi_t_bidelta)
new_row = {'x': bi_t_bidelta[0], 'y': bi_t_bidelta[1], 'z': bi_t_bidelta[2],
'qw': bi_q_bidelta[0], 'qx': bi_q_bidelta[1], 'qy': bi_q_bidelta[2],
'qz': bi_q_bidelta[3],}
rows_list.append(new_row)
return pd.DataFrame(data=rows_list, index=index_list)
def rename_euroc_gt_df(df):
""" Renames a DataFrame built from a EuRoC ground-truth data csv file to be easier to read.
Column labels are changed to be more readable and to be identical to the generic pose
trajectory format used with other csv files. Note that '#timestamp' will not actually
be renamed if it is the index of the DataFrame (which it should be). It will be
appropriately renamed if it is the index name.
This operation is 'inplace': It does not return a new DataFrame but simply changes
the existing one.
Args:
df: A pandas.DataFrame object.
"""
df.index.names = ["timestamp"]
df.rename(columns={" p_RS_R_x [m]": "x",
" p_RS_R_y [m]": "y",
" p_RS_R_z [m]": "z",
" q_RS_w []": "qw",
" q_RS_x []": "qx",
" q_RS_y []": "qy",
" q_RS_z []": "qz",
" v_RS_R_x [m s^-1]": "vx",
" v_RS_R_y [m s^-1]": "vy",
" v_RS_R_z [m s^-1]": "vz",
" b_w_RS_S_x [rad s^-1]": "bgx",
" b_w_RS_S_y [rad s^-1]": "bgy",
" b_w_RS_S_z [rad s^-1]": "bgz",
" b_a_RS_S_x [m s^-2]": "bax",
" b_a_RS_S_y [m s^-2]": "bay",
" b_a_RS_S_z [m s^-2]": "baz"}, inplace=True)
def rename_lcd_result_df(df):
""" Renames a DataFrame built from an LCD results measurements csv file to be converted to a trajectory.
This is an 'inplace' argument and returns nothing.
Args:
df: A pandas.DataFrame object.
"""
df.index.names = ["timestamp"]
df.rename(columns={"px": "x",
"py": "y",
"pz": "z"
}, inplace=True)
# -
# ## LoopClosureDetector Statistics Plotting
#
# Gather and plot various statistics on LCD module performance, including RANSAC information, keyframe status (w.r.t. loop closure detection), and loop closure events and the quality of their relative poses.
# ### LCD Status Frequency Chart
#
# Each keyframe is processed for potential loop closures. During this process, the loop-closure detector can either identify a loop closure or not. There are several reasons why a loop closure would not be detected. This plot helps to identify why loop closures are not detected between keyframes.
# +
output_lcd_status_filename = os.path.join(os.path.expandvars(vio_output_dir), "output_lcd_status.csv")
lcd_debuginfo_df = pd.read_csv(output_lcd_status_filename, sep=',', index_col=0)
status_freq_map = {}
for status in lcd_debuginfo_df.lcd_status:
if status not in status_freq_map:
status_freq_map[status] = 1
else:
status_freq_map[status] += 1
print("Full Size of PGO: ", lcd_debuginfo_df.pgo_size.tolist()[-1])
# Print the overall number of loop closures detected over all time.
if "LOOP_DETECTED" in status_freq_map:
print("Loop Closures Detected: ", status_freq_map["LOOP_DETECTED"])
else:
print("Loop Closures Detected: 0")
print("Loop Closures Registered by PGO by End: ", lcd_debuginfo_df.pgo_lc_count.tolist()[-1])
print("Loop Closures Accepted by PGO at End: ", lcd_debuginfo_df.pgo_lc_inliers.tolist()[-1])
# Plot failure modes as a histogram.
fig = plt.figure(figsize=(18,10))
plt.bar(status_freq_map.keys(), status_freq_map.values(), width=1.0)
plt.xticks(status_freq_map.keys(), list(status_freq_map.keys()))
plt.ylabel('Status Frequency')
plt.title('LoopClosureDetector Status Histogram')
plt.show()
# -
# ### LCD RANSAC Performance Charts
#
# Plot the performance of the geometric-verification and pose-recovery steps. These are handled by Nister (5pt) RANSAC and Arun (3pt) RANSAC respectively.
#
# inlier percentages and iterations are plotted for both methods.
# +
lcd_debuginfo_small_df = downsize_lc_df(lcd_debuginfo_df)
#Helper functions for processing data summary.
def get_mean(attrib):
ls = lcd_debuginfo_small_df[attrib].tolist()
return float(sum(ls)) / len(ls)
def get_min(attrib):
return min(lcd_debuginfo_small_df[attrib])
def get_max(attrib):
return max(lcd_debuginfo_small_df[attrib])
# Construct and visualize summary. TODO(marcus): use a LaTeX table.
summary_stats = [
("Average number of mono ransac inliers", get_mean("mono_inliers")),
("Average size of mono ransac input", get_mean("mono_input_size")),
("Average number of stereo ransac inliers", get_mean("stereo_inliers")),
("Average size of stereo ransac input", get_mean("stereo_input_size")),
("Maximum mono ransac iterations", get_max("mono_iters")),
("Maximum stereo ransac iterations", get_max("stereo_iters")),
]
attrib_len = [len(attrib[0]) for attrib in summary_stats]
max_attrib_len = max(attrib_len)
print "\nRANSAC Statistic Summary for Loop Closures ONLY:\n"
for entry in summary_stats:
attrib = entry[0]
value = entry[1]
spacing = max_attrib_len - len(attrib)
print attrib + " "*spacing + ": " + str(value)
# Plot ransac inlier and iteration statistics.
fig1, axes1 = plt.subplots(nrows=1, ncols=2, figsize=(18,10), squeeze=False)
lcd_debuginfo_small_df.plot(kind="hist", y="mono_inliers", ax=axes1[0,0])
lcd_debuginfo_small_df.plot(kind="hist", y="stereo_inliers", ax=axes1[0,0])
lcd_debuginfo_small_df.plot(kind="hist", y="mono_iters", ax=axes1[0,1])
lcd_debuginfo_small_df.plot(kind="hist", y="stereo_iters", ax=axes1[0,1])
plt.show()
# -
# ### LCD Relative Pose Error Plotting
#
# Calculate error statistics for all individual loop closures and plot their error as compared to ground truth. These plots give insight into how reliable the pose determination between two frames is for each loop closure. This pose determination is done via a combination of 5-pt and 3-pt RANSAC matching of the stereo images from the camera.
# +
gt_df = pd.read_csv(gt_data_file, sep=',', index_col=0)
rename_euroc_gt_df(gt_df)
output_loop_closures_filename = os.path.join(os.path.expandvars(vio_output_dir), "output_lcd_result.csv")
output_loop_closures_df = pd.read_csv(output_loop_closures_filename, sep=',', index_col=0)
# -
small_lc_df = downsize_lc_df(output_loop_closures_df)
rename_lcd_result_df(small_lc_df)
gt_rel_df = convert_abs_traj_to_rel_traj_lcd(gt_df, small_lc_df, to_scale=True)
# +
# Convert the gt relative-pose DataFrame to a trajectory object.
traj_ref = pandas_bridge.df_to_trajectory(gt_rel_df)
# Use the mono ransac file as estimated trajectory.
traj_est = pandas_bridge.df_to_trajectory(small_lc_df)
traj_ref, traj_est = sync.associate_trajectories(traj_ref, traj_est)
print "traj_ref: ", traj_ref
print "traj_est: ", traj_est
# +
# Get RPE for entire relative trajectory.
rpe_rot = get_rpe((traj_ref, traj_est), metrics.PoseRelation.rotation_angle_deg)
rpe_tran = get_rpe((traj_ref, traj_est), metrics.PoseRelation.translation_part)
# Print rotation RPE statistics:
rot_summary_stats = [
("mean", rpe_rot.get_statistic(metrics.StatisticsType.mean)),
("median", rpe_rot.get_all_statistics()["median"]),
("rmse", rpe_rot.get_statistic(metrics.StatisticsType.rmse)),
("std", rpe_rot.get_statistic(metrics.StatisticsType.std)),
("min", rpe_rot.get_statistic(metrics.StatisticsType.min)),
("max", rpe_rot.get_statistic(metrics.StatisticsType.max))
]
attrib_len = [len(attrib[0]) for attrib in rot_summary_stats]
max_attrib_len = max(attrib_len)
print "\nRotation RPE Statistics Summary:\n"
for entry in rot_summary_stats:
attrib = entry[0]
value = entry[1]
spacing = max_attrib_len - len(attrib)
print attrib + " "*spacing + ": " + str(value)
# Print translation RPE statistics:
tram_summary_stats = [
("mean", rpe_tran.get_statistic(metrics.StatisticsType.mean)),
("median", rpe_tran.get_all_statistics()["median"]),
("rmse", rpe_tran.get_statistic(metrics.StatisticsType.rmse)),
("std", rpe_tran.get_statistic(metrics.StatisticsType.std)),
("min", rpe_tran.get_statistic(metrics.StatisticsType.min)),
("max", rpe_tran.get_statistic(metrics.StatisticsType.max))
]
attrib_len = [len(attrib[0]) for attrib in tram_summary_stats]
max_attrib_len = max(attrib_len)
print "\nTranslation RPE Statistics Summary:\n"
for entry in tram_summary_stats:
attrib = entry[0]
value = entry[1]
spacing = max_attrib_len - len(attrib)
print attrib + " "*spacing + ": " + str(value)
# -
# ## LoopClosureDetector PGO-Optimized Trajectory Plotting
#
# Plot the APE, RPE, and trajectory of the Pose-graph-optimized trajectory, including loop closures on top of regular odometry updates.
#
# The results are visualized against both ground truth and the odometry-estimate alone to show the performance gain from loop closure detection.
# +
# Load ground truth and estimated data as csv DataFrames.
gt_df = pd.read_csv(gt_data_file, sep=',', index_col=0)
output_poses_filename = os.path.join(os.path.expandvars(vio_output_dir), "output_posesVIO.csv")
output_poses_df = pd.read_csv(output_poses_filename, sep=',', index_col=0)
output_pgo_poses_filename = os.path.join(os.path.expandvars(vio_output_dir), "output_lcd_optimized_traj.csv")
output_pgo_poses_df = pd.read_csv(output_pgo_poses_filename, sep=',', index_col=0)
# +
gt_df = gt_df[~gt_df.index.duplicated()]
rename_euroc_gt_df(gt_df)
# +
# Convert the gt relative-pose DataFrame to a trajectory object.
traj_ref = pandas_bridge.df_to_trajectory(gt_df)
# Compare against the VIO without PGO.
traj_ref_cp = copy.deepcopy(traj_ref)
traj_vio = pandas_bridge.df_to_trajectory(output_poses_df)
traj_ref_cp, traj_vio = sync.associate_trajectories(traj_ref_cp, traj_vio)
traj_vio = trajectory.align_trajectory(traj_vio, traj_ref_cp, correct_scale=False,
discard_n_start_poses = int(discard_n_start_poses),
discard_n_end_poses = int(discard_n_end_poses))
# Use the PGO output as estimated trajectory.
traj_est = pandas_bridge.df_to_trajectory(output_pgo_poses_df)
# Associate the data.
traj_ref, traj_est = sync.associate_trajectories(traj_ref, traj_est)
traj_est = trajectory.align_trajectory(traj_est, traj_ref, correct_scale=False,
discard_n_start_poses = int(discard_n_start_poses),
discard_n_end_poses = int(discard_n_end_poses))
print "traj_ref: ", traj_ref
print "traj_vio: ", traj_vio
print "traj_est: ", traj_est
# -
# ## Absolute-Pose-Error Plotting
#
# Plot absolute-pose-error along the entire trajectory. APE gives a good sense of overall VIO performance across the entire trajectory.
# +
# Plot APE of trajectory rotation and translation parts.
num_of_poses = traj_est.num_poses
traj_est.reduce_to_ids(range(int(discard_n_start_poses), int(num_of_poses - discard_n_end_poses), 1))
traj_ref.reduce_to_ids(range(int(discard_n_start_poses), int(num_of_poses - discard_n_end_poses), 1))
traj_vio.reduce_to_ids(range(int(discard_n_start_poses), int(num_of_poses - discard_n_end_poses), 1))
seconds_from_start = [t - traj_est.timestamps[0] for t in traj_est.timestamps]
ape_tran = get_ape((traj_ref, traj_est), metrics.PoseRelation.translation_part)
plot_ape(seconds_from_start, ape_tran, title="VIO+PGO ATE in Meters")
# +
# Plot the ground truth and estimated trajectories against each other with APE overlaid.
plot_mode = plot.PlotMode.xy
fig = plt.figure(figsize=(18,10))
ax = plot.prepare_axis(fig, plot_mode)
plot.traj(ax, plot_mode, traj_ref, '--', "gray", "reference")
plot.traj(ax, plot_mode, traj_vio, '.', "gray", "vio without pgo")
plot.traj_colormap(ax, traj_est, ape_tran.error, plot_mode,
min_map=ape_tran.get_all_statistics()["min"],
max_map=ape_tran.get_all_statistics()["max"],
title="VIO+PGO Trajectory Tracking - Color Coded by ATE")
ax.legend()
plt.show()
# -
# ## Relative-Pose-Error Plotting
#
# Plot relative-pose-error along the entire trajectory. RPE gives a good sense of overall VIO performance from one frame to the next.
# Get RPE for entire relative trajectory.
rpe_rot = get_rpe((traj_ref, traj_est), metrics.PoseRelation.rotation_angle_deg)
rpe_tran = get_rpe((traj_ref, traj_est), metrics.PoseRelation.translation_part)
# +
# Plot RPE of trajectory rotation and translation parts.
seconds_from_start = [t - traj_est.timestamps[0] for t in traj_est.timestamps[1:]]
plot_rpe(seconds_from_start, rpe_rot, title="VIO+PGO RRE in Degrees")
plot_rpe(seconds_from_start, rpe_tran, title="VIO+PGO RTE in Meters")
# +
# important: restrict data to delta ids for plot.
traj_ref_plot = copy.deepcopy(traj_ref)
traj_est_plot = copy.deepcopy(traj_est)
traj_ref_plot.reduce_to_ids(rpe_rot.delta_ids)
traj_est_plot.reduce_to_ids(rpe_rot.delta_ids)
# Plot the ground truth and estimated trajectories against each other with RPE overlaid.
plot_mode = plot.PlotMode.xy
fig = plt.figure(figsize=(18,10))
ax = plot.prepare_axis(fig, plot_mode)
plot.traj(ax, plot_mode, traj_ref_plot, '--', "gray", "reference")
plot.traj_colormap(ax, traj_est_plot, rpe_rot.error, plot_mode,
min_map=rpe_rot.get_all_statistics()["min"],
max_map=rpe_rot.get_all_statistics()["max"],
title="VIO+PGO Trajectory Tracking - Color Coded by RRE")
ax.legend()
plt.show()
# +
traj_vio = pandas_bridge.df_to_trajectory(output_poses_df)
traj_ref, traj_vio = sync.associate_trajectories(traj_ref, traj_est)
traj_vio = trajectory.align_trajectory(traj_vio, traj_ref, correct_scale=False)
# Plot the trajectories for quick error visualization.
fig = plt.figure(figsize=(18,10))
traj_by_label = {
"VIO only": traj_vio,
"VIO + PGO": traj_est,
"reference": traj_ref
}
plot.trajectories(fig, traj_by_label, plot.PlotMode.xyz, title="PIM Trajectory Tracking in 3D")
plt.show()
|
scripts/plotting/jupyter/plot_lcd.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: DESI master
# language: python
# name: desi-master
# ---
# # This notebook is for BGS specific results
import numpy as np
import fitsio
from matplotlib import pyplot as plt
import os
ff = fitsio.read('/global/cfs/cdirs/desi/survey/catalogs/SV3/LSS/LSScats/test/BGS_ANYAlltiles_full.dat.fits')
print('total number of unique reachable BGS targets is '+str(len(ff)))
wo = ff['LOCATION_ASSIGNED'] == 1
print('total number of unique observed BGS targets is '+str(len(ff[wo])))
wz = ff['ZWARN'] == 0
print('total number of unique BGS targets with good redshifts is '+str(len(ff[wz])))
print('targeting completeness is '+str(len(ff[wo])/len(ff)))
print('redshift success rate is '+str(len(ff[wz])/len(ff[wo])))
ngl = [len(ff[wz])]
ntm = [1]
for nt in range(1,7):
wt = ff['NTILE'] > nt
ntm.append(nt+1)
ngl.append(len(ff[wz&wt]))
plt.plot(ntm,np.array(ngl)/len(ff[wz]),color='purple')
plt.xlabel('minimum number of tiles')
plt.ylabel('fraction of good BGS redshifts kept')
plt.show()
#plot n(z)
nz = np.loadtxt('/global/cfs/cdirs/desi/survey/catalogs/SV3/LSS/LSScats/test/BGS_ANY_N_nz.dat').transpose()
plt.plot(nz[0],nz[3],':',color='darkorchid',label='BASS/MzLS')
nz = np.loadtxt('/global/cfs/cdirs/desi/survey/catalogs/SV3/LSS/LSScats/test/BGS_ANY_S_nz.dat').transpose()
plt.plot(nz[0],nz[3],'--',color='orchid',label='DECaLS')
plt.legend()
plt.xlim(0.01,0.7)
plt.ylim(0,0.1)
xl = [0.1,0.1]
yl = [0,0.1]
plt.plot(xl,yl,'k-')
xl = [0.3,0.3]
yl = [0,0.1]
plt.plot(xl,yl,'k-')
xl = [0.5,0.5]
yl = [0,0.1]
plt.plot(xl,yl,'k-')
plt.plot(xl,yl,'k-')
plt.xlabel('redshift')
plt.ylabel(r'comoving number density ($h$/Mpc)$^3$')
plt.show()
zl = [0.1,0.3,0.5]
for i in range(0,len(zl)):
if i == len(zl)-1:
zmin=zl[0]
zmax=zl[-1]
else:
zmin = zl[i]
zmax = zl[i+1]
xils = np.loadtxt('/global/cscratch1/sd/ajross/SV3xi/xi024SV3_testBGS_ANY_S'+str(zmin)+str(zmax)+'5st0.dat').transpose()
xil = np.loadtxt('/global/cscratch1/sd/ajross/SV3xi/xi024SV3_testBGS_ANY'+str(zmin)+str(zmax)+'5st0.dat').transpose()
xiln = np.loadtxt('/global/cscratch1/sd/ajross/SV3xi/xi024SV3_testBGS_ANY_N'+str(zmin)+str(zmax)+'5st0.dat').transpose()
plt.plot(xil[0],xil[0]**2.*xiln[1],'^:',color='darkorchid',label='BASS/MzLS')
plt.plot(xil[0],xil[0]**2.*xils[1],'v--',color='orchid',label='DECaLS')
plt.plot(xil[0],xil[0]**2.*xil[1],'s-',color='purple',label='combined')
xilin = np.loadtxt(os.environ['HOME']+'/BAOtemplates/xi0Challenge_matterpower0.44.04.08.015.00.dat').transpose()
plt.plot(xilin[0],xilin[0]**2.*xilin[1]*1.,'k-.',label=r'$\xi_{\rm 0}(z=0),b/D(z)=\sqrt{1.},\beta=0.4$')
plt.title('BGS_ANY SV3, '+str(zmin)+' < z < '+str(zmax))
plt.xlim(0,50)
plt.ylim(-30,80)
plt.xlabel(r'$s$ (Mpc/h)')
plt.ylabel(r'$s^2\xi_0$')
plt.legend()
plt.show()
# ## wacky things clearly going on for s > 40 mpc/h or so...
|
Sandbox/SV3BGS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit
# name: python3
# ---
# # MNIST Character Detection
# ## Impleneted by CNN using tensorflow
!#pip install tensorflow numpy pandas matplotlib
# +
import tensorflow as tf
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
np.set_printoptions(linewidth=200)
# -
chart = pd.read_csv("./data/train.csv")
# +
x_train = chart.to_numpy()
y_train, x_train = x_train[:, 0], x_train[:,1:]
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1)
x_train = x_train/255.0
# +
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if (logs.get('accuracy') >= 0.95): # Experiment with changing this value
print("\nReached 95% accuracy so cancelling training!")
self.model.stop_training = True
callbacks = myCallback()
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(64, (3,3), activation='relu', input_shape=(28,28,1)),
tf.keras.layers.MaxPooling2D(2,2),
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
model.compile(
optimizer = tf.optimizers.Adam(),
loss = 'sparse_categorical_crossentropy',
metrics=['accuracy'])
# -
model.fit(x_train, y_train, epochs=10, callbacks=[callbacks])
test_variables = pd.read_csv("./data/test.csv")
x_test = test_variables.to_numpy()
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
predictions = model.predict(x_test)
predicted_value = np.argmax(predictions, axis=1)
print(predicted_value.shape)
# +
predicted_value = predicted_value.reshape(predicted_value.shape[0], 1)
img_id = np.arange(start=1,
stop=(predicted_value.shape[0] + 1),
step=1,
dtype=int).reshape(predicted_value.shape[0], 1)
predicted_value = np.hstack((img_id, predicted_value))
print(predicted_value)
# -
results = pd.DataFrame(predicted_value, columns=['ImageId', 'Label'])
results.to_csv('./my_submission.csv', index=False)
|
MNIST_kaggle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
list_of_dataframe = []
rows_in_a_chunk = 10
num_chunks = 5
import pandas as pd
df_dummy = pd.read_csv("Boston_housing.csv",nrows=2)
colnames = df_dummy.columns
for i in range(0,num_chunks*rows_in_a_chunk,rows_in_a_chunk):
df = pd.read_csv("Boston_housing.csv",header=0,skiprows=i,nrows=rows_in_a_chunk,names=colnames)
list_of_dataframe.append(df)
|
Chapter05/.ipynb_checkpoints/Exercise 5.05-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pprint
print([100] > [-100])
print([1, 2, 100] > [1, 2, -100])
print([1, 2, 100] > [1, 100])
# +
l_2d = [[2, 30, 100], [1, 20, 300], [3, 10, 200]]
pprint.pprint(l_2d, width=40)
# +
l_2d.sort()
pprint.pprint(l_2d, width=40)
# +
l_2d.sort(key=lambda x: x[1])
pprint.pprint(l_2d, width=40)
# +
l_2d.sort(key=lambda x: x[2], reverse=True)
pprint.pprint(l_2d, width=40)
# +
l_sorted = sorted(l_2d, key=lambda x: x[0], reverse=True)
pprint.pprint(l_sorted, width=40)
# +
l_3d = [[[0, 1, 2], [2, 30, 100]], [[3, 4, 5], [1, 20, 300]], [[6, 7, 8], [3, 10, 200]]]
pprint.pprint(l_3d, width=40)
# +
l_sorted = sorted(l_3d, key=lambda x: x[1][0])
pprint.pprint(l_sorted, width=40)
|
notebook/list_2d_sort.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ProtoNN in Tensorflow
#
# This is a simple notebook that illustrates the usage of Tensorflow implementation of ProtoNN. We are using the USPS dataset. Please refer to `fetch_usps.py` and `process_usps.py`for more details on downloading the dataset.
# +
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT license.
from __future__ import print_function
import sys
import os
import numpy as np
import tensorflow as tf
from edgeml_tf.trainer.protoNNTrainer import ProtoNNTrainer
from edgeml_tf.graph.protoNN import ProtoNN
import edgeml_tf.utils as utils
import helpermethods as helper
# -
# # USPS Data
#
# It is assumed that the USPS data has already been downloaded and set up with the help of [fetch_usps.py](fetch_usps.py) and is placed in the `./usps10` subdirectory.
# +
# Load data
DATA_DIR = './usps10'
train, test = np.load(DATA_DIR + '/train.npy'), np.load(DATA_DIR + '/test.npy')
x_train, y_train = train[:, 1:], train[:, 0]
x_test, y_test = test[:, 1:], test[:, 0]
numClasses = max(y_train) - min(y_train) + 1
numClasses = max(numClasses, max(y_test) - min(y_test) + 1)
numClasses = int(numClasses)
y_train = helper.to_onehot(y_train, numClasses)
y_test = helper.to_onehot(y_test, numClasses)
dataDimension = x_train.shape[1]
numClasses = y_train.shape[1]
# -
# # Model Parameters
#
# Note that ProtoNN is very sensitive to the value of the hyperparameter $\gamma$, here stored in valiable `GAMMA`. If `GAMMA` is set to `None`, median heuristic will be used to estimate a good value of $\gamma$ through the `helper.getGamma()` method. This method also returns the corresponding `W` and `B` matrices which should be used to initialize ProtoNN (as is done here).
PROJECTION_DIM = 60
NUM_PROTOTYPES = 60
REG_W = 0.000005
REG_B = 0.0
REG_Z = 0.00005
SPAR_W = 0.8
SPAR_B = 1.0
SPAR_Z = 1.0
LEARNING_RATE = 0.05
NUM_EPOCHS = 200
BATCH_SIZE = 32
GAMMA = 0.0015
W, B, gamma = helper.getGamma(GAMMA, PROJECTION_DIM, dataDimension,
NUM_PROTOTYPES, x_train)
# +
# Setup input and train protoNN
X = tf.placeholder(tf.float32, [None, dataDimension], name='X')
Y = tf.placeholder(tf.float32, [None, numClasses], name='Y')
protoNN = ProtoNN(dataDimension, PROJECTION_DIM,
NUM_PROTOTYPES, numClasses,
gamma, W=W, B=B)
trainer = ProtoNNTrainer(protoNN, REG_W, REG_B, REG_Z,
SPAR_W, SPAR_B, SPAR_Z,
LEARNING_RATE, X, Y, lossType='xentropy')
sess = tf.Session()
trainer.train(BATCH_SIZE, NUM_EPOCHS, sess, x_train, x_test, y_train, y_test,
printStep=600, valStep=10)
# -
# # Model Evaluation
acc = sess.run(protoNN.accuracy, feed_dict={X: x_test, Y: y_test})
# W, B, Z are tensorflow graph nodes
W, B, Z, _ = protoNN.getModelMatrices()
matrixList = sess.run([W, B, Z])
sparcityList = [SPAR_W, SPAR_B, SPAR_Z]
nnz, size, sparse = helper.getModelSize(matrixList, sparcityList)
print("Final test accuracy", acc)
print("Model size constraint (Bytes): ", size)
print("Number of non-zeros: ", nnz)
nnz, size, sparse = helper.getModelSize(matrixList, sparcityList, expected=False)
print("Actual model size: ", size)
print("Actual non-zeros: ", nnz)
|
examples/tf/ProtoNN/protoNN_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Watch Me Code 2: The Need For Exception Handling
# This demonstrates the need for exception handling.
# +
# this generates a run-time error when you enter a non-number
# for example enter "heavy" and you get a ValueError
weight = float(input("Enter product weight in Kg: "))
# -
# This example uses try..except to catch the ValueError
try:
weight = float(input("Enter product weight in Kg: "))
print ("Weight is:", weight)
except ValueError:
print("You did not enter a number! ")
|
lessons/03-Conditionals/WMC2-The-Need-For-Exception-Handling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import Statement
from pyspark.sql import SQLContext
from pyspark.sql import functions as sf
from matplotlib import pyplot as plt
from pyspark.sql.functions import col, avg
import pandas as pd
import pyspark
from datetime import datetime, timedelta, date
from pyspark.sql.functions import broadcast
from pyspark.sql.types import DateType
from pyspark.sql.functions import col, avg, date_format,month,hour,lag, date_sub,lit
log4jLogger = sc._jvm.org.apache.log4j
LOGGER = log4jLogger.LogManager.getLogger(__name__)
LOGGER.error("pyspark script logger initialized")
sc.stop()
sc = pyspark.SparkContext(master="spark://172.16.27.208:7077",appName="spark")
sc
base_path = "/home/test5/Desktop/smart-meters-in-london/"
sqlcontext = SQLContext(sc)
#sqlcontext
household_info = sqlcontext.read.csv(base_path+"informations_households.csv",header=True,inferSchema=True)
#household_mini = sc.parallelize(household_info.take(1)).toDF()
household_mini = household_info
household_mini.printSchema()
column_list = []
for i in range(48):
column_list.append("hh_"+str(i))
column_list
new_column_list = []
for i in range(1,49):
if i<20:
new_column_list.append("0"+str(i*0.5))
else:
new_column_list.append(str(i*0.5))
flag = 0
df_full = []
df_file = household_mini.select("file").distinct()
# exprs = {x: "avg" for x in new_column_list}
exprs1 = [avg(x) for x in column_list[0:40]]
exprs2 = [avg(x) for x in column_list[40:48]] #due to the fact large number of column giving error so divide
count = 0
for row in df_file.rdd.collect():
file = row.file
print(file,count)
count += 1
file_path = base_path + "hhblock_dataset/"+ file+".csv"
half_hourly_consumption_data = sqlcontext.read.csv(file_path,header=True,inferSchema=True).cache()
half_hourly_consumption_data.dropna(how='any')
#half_hourly_consumption_data2 = half_hourly_consumption_data.groupBy('LCLid').agg(*exprs2)
#half_hourly_consumption_data = half_hourly_consumption_data.groupBy('LCLid').agg(*exprs1)
#half_hourly_consumption_data = half_hourly_consumption_data.join(half_hourly_consumption_data2,["LCLid"])
#half_hourly_consumption_data.dropna(how='any')
#half_hourly_consumption_data.printSchema()
if flag == 0:
df_full = sqlcontext.createDataFrame([],half_hourly_consumption_data.schema)
flag = 1
df_full = df_full.union(half_hourly_consumption_data)
df_full = df_full.cache()
#avg_house_data.take(1)
df_full = df_full.repartition(480,"LCLid")
df_full.printSchema()
# ## filtering data as per requirement
# Total user in 2013 5528
#
# Total user in 2013 with full evidence 3961
#
df_full = df_full.withColumn("day",df_full["day"].cast(DateType()))
df_full.printSchema()
#df_full = df_full.withColum("date",date_format(df_full["date"],"yyyy-MM-dd").cast(Datetype()))
# df_full = df_full.filter((df_full.day >= date(2013,1,1)) & (df_full.day <= date(2013,10,31)))
df_full = df_full.na.drop()
# print ("Total user in 2013 ", df_full.select("LCLid").distinct().count())
# year_df = df_full.groupBy("LCLid").count()
# year_df = year_df.filter(year_df["count"] >= 365 )
# print("Total user in 2013 with full evidence ", year_df.select("LCLid").distinct().count())
LCLid_under_Consideration = sqlcontext.read.csv(base_path+"Feature_File/Cleaned_2013_Features_mth_5.csv",header=True)
LCLid_under_Consideration = LCLid_under_Consideration.select("LCLid").distinct()
df_full = df_full.join(broadcast(LCLid_under_Consideration),["LCLid"])
df_full = df_full.filter((df_full.day >= date(2013,1,1)) & (df_full.day <= date(2013,10,31)))
# LCLid_under_Consideration.count() #3930
# year_df = df_full.groupBy("LCLid").count()
# year_df.take(1)
# year_df = year_df.filter(year_df["count"] >= 365 )
half_hourly_consumption_df = df_full#.join(broadcast(year_df),["LCLid"])
half_hourly_consumption_df.take(1)
# +
#flag = 0
#avg_house_data = []
#block_read = set([])
#for row in household_mini.rdd.collect():
# house_id = row.LCLid
# file = row.file
# print(house_id,file)
# file_path = base_path + "hhblock_dataset/"+ file+".csv"
# if file not in block_read:
## print("hi")
# block_read.add(file)
# half_hourly_consumption_data = sqlcontext.read.csv(file_path,header=True,inferSchema=True)
# half_hourly_consumption_data.dropna(how='any')
# for c,n in zip(column_list,new_column_list):
# half_hourly_consumption_data=half_hourly_consumption_data.withColumnRenamed(c,n)
# indiv_house_data = half_hourly_consumption_data.where(col("LCLid") == house_id)
# indiv_house_data = indiv_house_data.toHandy()
# if indiv_house_data.rdd.isEmpty():
# print("Missing Id = {} in file = {}".format(house_id,file))
# continue
# indiv_house_data = sqlcontext.createDataFrame(indiv_house_data.stratify(['LCLid']).cols[new_column_list].mean().reset_index())
# indiv_house_data.printSchema()
# if flag == 0:
# avg_house_data = sqlcontext.createDataFrame([],indiv_house_data.schema)
# flag = 1
# avg_house_data = avg_house_data.union(indiv_house_data)
# -
half_hourly_consumption_data2 = half_hourly_consumption_df.groupBy('LCLid').agg(*exprs2)
half_hourly_consumption_data = half_hourly_consumption_df.groupBy('LCLid').agg(*exprs1)
avg_house_data = half_hourly_consumption_data.join(half_hourly_consumption_data2,["LCLid"])
avg_house_data=avg_house_data.dropna(how='any')
avg_house_data.printSchema()
for c,n in zip(avg_house_data.columns[1:],new_column_list):
avg_house_data=avg_house_data.withColumnRenamed(c,n)
avg_house_data.printSchema()
avg_house_data = avg_house_data.toPandas()
# pd.options.display.max_columns = None
avg_house_data.shape
plot = avg_house_data.set_index("LCLid").T.plot(figsize=(13,8), legend=False, color='blue',alpha=0.5)
# +
#avg_house_data1 = sqlcontext.createDataFrame(avg_house_data)
#avg_house_data1.write.format("csv").save(base_path+"avg.csv")
# -
plot.get_figure().savefig(base_path+"/plot/Avg_LP.png")
avg_house_data.to_csv(base_path+"avg.csv", header=True)
|
Average_load_profile.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mukulsa/tensorflow_ai/blob/main/TensorFlow%20In%20Practice/Course%203%20-%20NLP/Course%203%20-%20Week%202%20-%20Exercise%20-%20Answer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="zX4Kg8DUTKWO"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="view-in-github"
# <a href="https://colab.research.google.com/github/lmoroney/dlaicourse/blob/master/TensorFlow%20In%20Practice/Course%203%20-%20NLP/Course%203%20-%20Week%202%20-%20Exercise%20-%20Answer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="rX8mhOLljYeM"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" id="BZSlp3DAjdYf"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + id="gnwiOnGyW5JK"
import csv
import tensorflow as tf
import numpy as np
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/bbc-text.csv \
# -O /tmp/bbc-text.csv
# + id="EYo6A4v5ZABQ"
vocab_size = 1000
embedding_dim = 16
max_length = 120
trunc_type='post'
padding_type='post'
oov_tok = "<OOV>"
training_portion = .8
# + id="iU1qq3_SZBx_"
sentences = []
labels = []
stopwords = [ "a", "about", "above", "after", "again", "against", "all", "am", "an", "and", "any", "are", "as", "at", "be", "because", "been", "before", "being", "below", "between", "both", "but", "by", "could", "did", "do", "does", "doing", "down", "during", "each", "few", "for", "from", "further", "had", "has", "have", "having", "he", "he'd", "he'll", "he's", "her", "here", "here's", "hers", "herself", "him", "himself", "his", "how", "how's", "i", "i'd", "i'll", "i'm", "i've", "if", "in", "into", "is", "it", "it's", "its", "itself", "let's", "me", "more", "most", "my", "myself", "nor", "of", "on", "once", "only", "or", "other", "ought", "our", "ours", "ourselves", "out", "over", "own", "same", "she", "she'd", "she'll", "she's", "should", "so", "some", "such", "than", "that", "that's", "the", "their", "theirs", "them", "themselves", "then", "there", "there's", "these", "they", "they'd", "they'll", "they're", "they've", "this", "those", "through", "to", "too", "under", "until", "up", "very", "was", "we", "we'd", "we'll", "we're", "we've", "were", "what", "what's", "when", "when's", "where", "where's", "which", "while", "who", "who's", "whom", "why", "why's", "with", "would", "you", "you'd", "you'll", "you're", "you've", "your", "yours", "yourself", "yourselves" ]
print(len(stopwords))
# Expected Output
# 153
# + id="eutB2xMiZD0e"
with open("/tmp/bbc-text.csv", 'r') as csvfile:
reader = csv.reader(csvfile, delimiter=',')
next(reader)
for row in reader:
labels.append(row[0])
sentence = row[1]
for word in stopwords:
token = " " + word + " "
sentence = sentence.replace(token, " ")
sentences.append(sentence)
print(len(labels))
print(len(sentences))
print(sentences[0])
# Expected Output
# 2225
# 2225
# tv future hands viewers home theatre systems plasma high-definition tvs digital video recorders moving living room way people watch tv will radically different five years time. according expert panel gathered annual consumer electronics show las vegas discuss new technologies will impact one favourite pastimes. us leading trend programmes content will delivered viewers via home networks cable satellite telecoms companies broadband service providers front rooms portable devices. one talked-about technologies ces digital personal video recorders (dvr pvr). set-top boxes like us s tivo uk s sky+ system allow people record store play pause forward wind tv programmes want. essentially technology allows much personalised tv. also built-in high-definition tv sets big business japan us slower take off europe lack high-definition programming. not can people forward wind adverts can also forget abiding network channel schedules putting together a-la-carte entertainment. us networks cable satellite companies worried means terms advertising revenues well brand identity viewer loyalty channels. although us leads technology moment also concern raised europe particularly growing uptake services like sky+. happens today will see nine months years time uk adam hume bbc broadcast s futurologist told bbc news website. likes bbc no issues lost advertising revenue yet. pressing issue moment commercial uk broadcasters brand loyalty important everyone. will talking content brands rather network brands said tim hanlon brand communications firm starcom mediavest. reality broadband connections anybody can producer content. added: challenge now hard promote programme much choice. means said stacey jolna senior vice president tv guide tv group way people find content want watch simplified tv viewers. means networks us terms channels take leaf google s book search engine future instead scheduler help people find want watch. kind channel model might work younger ipod generation used taking control gadgets play them. might not suit everyone panel recognised. older generations comfortable familiar schedules channel brands know getting. perhaps not want much choice put hands mr hanlon suggested. end kids just diapers pushing buttons already - everything possible available said mr hanlon. ultimately consumer will tell market want. 50 000 new gadgets technologies showcased ces many enhancing tv-watching experience. high-definition tv sets everywhere many new models lcd (liquid crystal display) tvs launched dvr capability built instead external boxes. one example launched show humax s 26-inch lcd tv 80-hour tivo dvr dvd recorder. one us s biggest satellite tv companies directtv even launched branded dvr show 100-hours recording capability instant replay search function. set can pause rewind tv 90 hours. microsoft chief bill gates announced pre-show keynote speech partnership tivo called tivotogo means people can play recorded programmes windows pcs mobile devices. reflect increasing trend freeing multimedia people can watch want want.
# + id="XfdaWh06ZGe3"
train_size = int(len(sentences) * training_portion)
train_sentences = sentences[:train_size]
train_labels = labels[:train_size]
validation_sentences = sentences[train_size:]
validation_labels = labels[train_size:]
print(train_size)
print(len(train_sentences))
print(len(train_labels))
print(len(validation_sentences))
print(len(validation_labels))
# Expected output (if training_portion=.8)
# 1780
# 1780
# 1780
# 445
# 445
# + id="ULzA8xhwZI22"
tokenizer = Tokenizer(num_words = vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(train_sentences)
word_index = tokenizer.word_index
train_sequences = tokenizer.texts_to_sequences(train_sentences)
train_padded = pad_sequences(train_sequences, padding=padding_type, maxlen=max_length)
print(len(train_sequences[0]))
print(len(train_padded[0]))
print(len(train_sequences[1]))
print(len(train_padded[1]))
print(len(train_sequences[10]))
print(len(train_padded[10]))
# Expected Ouput
# 449
# 120
# 200
# 120
# 192
# 120
# + id="c8PeFWzPZLW_"
validation_sequences = tokenizer.texts_to_sequences(validation_sentences)
validation_padded = pad_sequences(validation_sequences, padding=padding_type, maxlen=max_length)
print(len(validation_sequences))
print(validation_padded.shape)
# Expected output
# 445
# (445, 120)
# + id="XkWiQ_FKZNp2"
label_tokenizer = Tokenizer()
label_tokenizer.fit_on_texts(labels)
training_label_seq = np.array(label_tokenizer.texts_to_sequences(train_labels))
validation_label_seq = np.array(label_tokenizer.texts_to_sequences(validation_labels))
print(training_label_seq[0])
print(training_label_seq[1])
print(training_label_seq[2])
print(training_label_seq.shape)
print(validation_label_seq[0])
print(validation_label_seq[1])
print(validation_label_seq[2])
print(validation_label_seq.shape)
# Expected output
# [4]
# [2]
# [1]
# (1780, 1)
# [5]
# [4]
# [3]
# (445, 1)
# + id="HZ5um4MWZP-W"
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.GlobalAveragePooling1D(),
tf.keras.layers.Dense(24, activation='relu'),
tf.keras.layers.Dense(6, activation='softmax')
])
model.compile(loss='sparse_categorical_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
# Expected Output
# Layer (type) Output Shape Param #
# =================================================================
# embedding (Embedding) (None, 120, 16) 16000
# _________________________________________________________________
# global_average_pooling1d (Gl (None, 16) 0
# _________________________________________________________________
# dense (Dense) (None, 24) 408
# _________________________________________________________________
# dense_1 (Dense) (None, 6) 150
# =================================================================
# Total params: 16,558
# Trainable params: 16,558
# Non-trainable params: 0
# + id="XsfdxySKZSXu"
num_epochs = 30
history = model.fit(train_padded, training_label_seq, epochs=num_epochs, validation_data=(validation_padded, validation_label_seq), verbose=2)
# + id="dQ0BX2apXS9u"
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
plot_graphs(history, "accuracy")
plot_graphs(history, "loss")
# + id="w7Xc-uWxXhML"
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_sentence(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
# + id="OhnFA_TDXrih"
e = model.layers[0]
weights = e.get_weights()[0]
print(weights.shape) # shape: (vocab_size, embedding_dim)
# Expected output
# (1000, 16)
# + id="_POzcWWAXudL"
import io
out_v = io.open('vecs.tsv', 'w', encoding='utf-8')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for word_num in range(1, vocab_size):
word = reverse_word_index[word_num]
embeddings = weights[word_num]
out_m.write(word + "\n")
out_v.write('\t'.join([str(x) for x in embeddings]) + "\n")
out_v.close()
out_m.close()
# + id="VmqpQMZ_XyOa"
try:
from google.colab import files
except ImportError:
pass
else:
files.download('vecs.tsv')
files.download('meta.tsv')
|
TensorFlow In Practice/Course 3 - NLP/Course 3 - Week 2 - Exercise - Answer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# fds.datax
# --------
#
# fds.datax offers instant access to define and retrieve a time-series universe of securities that is cached into a centralized data store. Having access to a cached data store shifts the research period focused on building a universe to immediately analyzing FactSet’s core & 3rd Party data using a consistent process.
#
#
# Main Features
# --------------------
#
# - Generate a daily universe from FDS Ownership based on an ETF-Identifier _(SPY-US)_
# - Generate files for a universe that will contain FDS Symbology, security reference, pricing and corporate actions data
# - Maintain and view a list of data stores available locally
# - Functions to support the generation of FDS symbology, security reference, pricing, and corporate actions Dataframes.
#
#
#
# Usage
# ---------
# The FactSet team recommends users leverage the `fds.datax` library to easily create and manage caches of pre-linked FactSet content for a universe of securities defined by the user.
#
# #### Doc Strings
# Throughout the `fds.datax Overview` notebook, you can leverage python's native doc strings by hitting `Shift + Tab`.
#
#
# Module Functionality
# ------------------------------
# The `fds.datax` library has 2 core modules. Each module has a series of underlying functionality that is described in the details section of the user guide.
#
# 1. **Universe**
#
# - locate - view or delete a data caches within a given data store
# - create – create, load or rebuild a data cache
# - rebuild - refresh an existing data store based on previous criteria
# - delete - remove an existing data cache
# - read - read in data files for a specific data cache
#
#
# 2. **GetSDFdata**
#
# This method contains the underlying elements that can be used to retrieve symbology, prices, corporate actions, and fx rates. Please refer to the User Guide & API details for more information.
#
#
# Interactive Code Sections
# -------------------------
# - [1. Setting the Working Directory](#step1)
# - [2. Setting the Working Directory](#step2)
# - [3. Locating Existing Data Caches](#step3)
# - [4. Building Your First Data Cache](#step4)
# - [5. Reading in Elements of a Data Cache](#step5)
# - [6. Rebuilding a Data Cache](#step6)
# - [7. Deleting a Data Cache](#step7)
#
# _____________________
# <a id='step1'></a>
#
# # Getting Started
#
# ## 1. Import Library
# Import the `fds.datax` Library in the first code block.
import fds.datax as dx
import os
# <a id='step2'></a>
# ## 2. Setting the Working Directory
#
# This package can be instantiated with a default directory path. This path will define the location of your data store.
#
# Let's save an instance with a new default directory.
# **Note**: If left blank, the data store defaults to the current working directory.
#
# We will use the namespace, `ds`, to represent "data store".
# +
fds_path = os.getcwd()
univ = dx.Universe(dir_path=fds_path)
# Display the working directory
univ.dir_path
# -
# <a id='step3'></a>
#
# ____________
#
# ## 3. Locating Existing Data Caches
#
# Next, let's use the `ds.locate` function to examine the existing data store. If the current working directory does not have any cached data files, the message below will be displayed.
#
# No existing Data Cache Universes exist in this Data Store. Use the fds.datax.Universe.create() function to generate a new data cache universe.
univ.locate()
# <a id='step4'></a>
#
# _________________
# ## 4. Building Your First Data Cache
#
# The `ds.create` function creates a series of data files that will be added to the data store within our working directory defined above. Let's use this function to build a cache for the inputs specified below.
# +
# Specify a cache name
cn = "my_first_data_cache"
# DSN name for a connection to a MSSQL Server DB containing FDS Standard DataFeeds content.
# `SDF` is the default as it connects to the FactSet Standard DataFeed data
mssql_dsn = "SDF"
# ETF ticker in a ticker-region format.
etf_ticker = "IWV-US"
# Currency for pricing data to be return, if local currency is desired set to "LOCAL"
currency = "USD"
# Earliest available report date in YYYY-MM-DD format
start_date = "2019-10-31"
# Most recent available report date to be returned in YYYY-MM-DD format
end_date = "2019-12-31"
# -
# Call the `create()` function to generate a time-series universe.
univ.create(
"generate",
cache_name=cn,
mssql_dsn=mssql_dsn,
etf_ticker=etf_ticker,
currency=currency,
start_date=start_date,
end_date=end_date,
)
# ### Checking the Available Files
#
# Once a data cache is created, use the `ds.locate` function to see the full catalog of caches in the data store.
univ.locate()
# <a id='step5'></a>
# _________________
# ## 5. Reading in Elements of a Data Cache
#
# The `ds.read` function will load data files specific to the universe and cache name that is specified. Let's read in the data that was created above.
#
# The `option` parameter accepts one of the following values:
# - `universe` – load time series symbology information for a data cache universe
# - `sec ref` – load security reference data for a data cache universe
# - `prices` – load pricing data for a data cache universe. Choose unadjusted, split, or split & spin-off data
# - `corp actions` – load corporate action factors for a data cache universe
# ### Load Symbology (Time Series)
univ.read(option="universe", cache_name="my_first_data_cache").head()
# ### Load Sec Reference Data
univ.read(option="sec ref", cache_name="my_first_data_cache").head()
# ### Load Pricing Data
#
# Pricing is unique as it has an additional option besides the **cache_name.** The user can specific the type of adjustments to be applied to the data using the `adj` parameter:
#
# * 0 = Unadjusted Data
# * 1 = Split Adjusted Data
# * 2 = Split and Spin-off Adjusted Data
univ.read(option="Prices", cache_name="my_first_data_cache", adj=2).head()
# ### Corporate Action Adjustment Factors
univ.read(option="Corp Actions", cache_name="my_first_data_cache").head()
# <a id='step6'></a>
# ## 6. Rebuilding a Data Cache
#
# To rebuild an existing data cache `ds.rebuild` can be used. This will trigger a refresh of the content based on the original inputs found within `ds.locate` accounting for corporate actions since the last rund date.
#
#
univ.rebuild(cache_name="my_first_data_cache")
# <a id='step7'></a>
# ## 7. Deleting a Data Cache
#
# Using `ds.delete` will remove the files associated with the data cache from the given data store. Let's first run the `ds.locate()` function to view an available cache. Then call the `ds.delete()` function to remove the cache from the Data Store.
univ.locate()
univ.delete(cache_name="my_first_data_cache")
univ.locate()
|
examples/fds.datax Overview.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: chineseocr
# language: python
# name: chineseocr
# ---
# ## 加载模型
#
# +
import os
import json
import time
import web
import numpy as np
from PIL import Image
from config import *
from apphelper.image import union_rbox,adjust_box_to_origin,base64_to_PIL
from application import trainTicket,idcard
if yoloTextFlag =='keras' or AngleModelFlag=='tf' or ocrFlag=='keras':
if GPU:
os.environ["CUDA_VISIBLE_DEVICES"] = str(GPUID)
import tensorflow as tf
from keras import backend as K
config = tf.ConfigProto()
config.gpu_options.allocator_type = 'BFC'
config.gpu_options.per_process_gpu_memory_fraction = 0.3## GPU最大占用量
config.gpu_options.allow_growth = True##GPU是否可动态增加
K.set_session(tf.Session(config=config))
K.get_session().run(tf.global_variables_initializer())
else:
##CPU启动
os.environ["CUDA_VISIBLE_DEVICES"] = ''
if yoloTextFlag=='opencv':
scale,maxScale = IMGSIZE
from text.opencv_dnn_detect import text_detect
elif yoloTextFlag=='darknet':
scale,maxScale = IMGSIZE
from text.darknet_detect import text_detect
elif yoloTextFlag=='keras':
scale,maxScale = IMGSIZE[0],2048
from text.keras_detect import text_detect
else:
print( "err,text engine in keras\opencv\darknet")
from text.opencv_dnn_detect import angle_detect
if ocr_redis:
##多任务并发识别
from helper.redisbase import redisDataBase
ocr = redisDataBase().put_values
else:
from crnn.keys import alphabetChinese,alphabetEnglish
if ocrFlag=='keras':
from crnn.network_keras import CRNN
if chineseModel:
alphabet = alphabetChinese
if LSTMFLAG:
ocrModel = ocrModelKerasLstm
else:
ocrModel = ocrModelKerasDense
else:
ocrModel = ocrModelKerasEng
alphabet = alphabetEnglish
LSTMFLAG = True
elif ocrFlag=='torch':
from crnn.network_torch import CRNN
if chineseModel:
alphabet = alphabetChinese
if LSTMFLAG:
ocrModel = ocrModelTorchLstm
else:
ocrModel = ocrModelTorchDense
else:
ocrModel = ocrModelTorchEng
alphabet = alphabetEnglish
LSTMFLAG = True
elif ocrFlag=='opencv':
from crnn.network_dnn import CRNN
ocrModel = ocrModelOpencv
alphabet = alphabetChinese
else:
print( "err,ocr engine in keras\opencv\darknet")
nclass = len(alphabet)+1
if ocrFlag=='opencv':
crnn = CRNN(alphabet=alphabet)
else:
crnn = CRNN( 32, 1, nclass, 256, leakyRelu=False,lstmFlag=LSTMFLAG,GPU=GPU,alphabet=alphabet)
if os.path.exists(ocrModel):
crnn.load_weights(ocrModel)
else:
print("download model or tranform model with tools!")
ocr = crnn.predict_job
from main import TextOcrModel
model = TextOcrModel(ocr,text_detect,angle_detect)
from apphelper.image import xy_rotate_box,box_rotate,solve
# +
import cv2
import numpy as np
def plot_box(img,boxes):
blue = (0, 0, 0) #18
tmp = np.copy(img)
for box in boxes:
cv2.rectangle(tmp, (int(box[0]),int(box[1])), (int(box[2]), int(box[3])), blue, 1) #19
return Image.fromarray(tmp)
def plot_boxes(img,angle, result,color=(0,0,0)):
tmp = np.array(img)
c = color
h,w = img.shape[:2]
thick = int((h + w) / 300)
i = 0
if angle in [90,270]:
imgW,imgH = img.shape[:2]
else:
imgH,imgW= img.shape[:2]
for line in result:
cx =line['cx']
cy = line['cy']
degree =line['degree']
w = line['w']
h = line['h']
x1,y1,x2,y2,x3,y3,x4,y4 = xy_rotate_box(cx, cy, w, h, degree/180*np.pi)
x1,y1,x2,y2,x3,y3,x4,y4 = box_rotate([x1,y1,x2,y2,x3,y3,x4,y4],angle=(360-angle)%360,imgH=imgH,imgW=imgW)
cx =np.mean([x1,x2,x3,x4])
cy = np.mean([y1,y2,y3,y4])
cv2.line(tmp,(int(x1),int(y1)),(int(x2),int(y2)),c,1)
cv2.line(tmp,(int(x2),int(y2)),(int(x3),int(y3)),c,1)
cv2.line(tmp,(int(x3),int(y3)),(int(x4),int(y4)),c,1)
cv2.line(tmp,(int(x4),int(y4)),(int(x1),int(y1)),c,1)
mess=str(i)
cv2.putText(tmp, mess, (int(cx), int(cy)),0, 1e-3 * h, c, thick // 2)
i+=1
return Image.fromarray(tmp).convert('RGB')
def plot_rboxes(img,boxes,color=(0,0,0)):
tmp = np.array(img)
c = color
h,w = img.shape[:2]
thick = int((h + w) / 300)
i = 0
for box in boxes:
x1,y1,x2,y2,x3,y3,x4,y4 = box
cx =np.mean([x1,x2,x3,x4])
cy = np.mean([y1,y2,y3,y4])
cv2.line(tmp,(int(x1),int(y1)),(int(x2),int(y2)),c,1)
cv2.line(tmp,(int(x2),int(y2)),(int(x3),int(y3)),c,1)
cv2.line(tmp,(int(x3),int(y3)),(int(x4),int(y4)),c,1)
cv2.line(tmp,(int(x4),int(y4)),(int(x1),int(y1)),c,1)
mess=str(i)
cv2.putText(tmp, mess, (int(cx), int(cy)),0, 1e-3 * h, c, thick // 2)
i+=1
return Image.fromarray(tmp).convert('RGB')
# +
import time
from PIL import Image
p = './test/idcard-demo.jpeg'
img = cv2.imread(p)
h,w = img.shape[:2]
timeTake = time.time()
scale=608
maxScale=2048
result,angle= model.model(img,
detectAngle=True,##是否进行文字方向检测
scale=scale,
maxScale=maxScale,
MAX_HORIZONTAL_GAP=80,##字符之间的最大间隔,用于文本行的合并
MIN_V_OVERLAPS=0.6,
MIN_SIZE_SIM=0.6,
TEXT_PROPOSALS_MIN_SCORE=0.1,
TEXT_PROPOSALS_NMS_THRESH=0.7,
TEXT_LINE_NMS_THRESH = 0.9,##文本行之间测iou值
LINE_MIN_SCORE=0.1,
leftAdjustAlph=0,##对检测的文本行进行向左延伸
rightAdjustAlph=0.1,##对检测的文本行进行向右延伸
)
timeTake = time.time()-timeTake
print('It take:{}s'.format(timeTake))
for line in result:
print(line['text'])
plot_boxes(img,angle, result,color=(0,0,0))
# -
boxes,scores = model.detect_box(img,608,2048)
plot_box(img,boxes)
|
test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create a logistic regression model to predict several mutations from covariates
# +
import os
import itertools
import warnings
import collections
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import preprocessing, grid_search
from sklearn.linear_model import SGDClassifier
from sklearn.cross_validation import train_test_split
from sklearn.metrics import roc_auc_score, roc_curve
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from scipy.special import logit
# -
# %matplotlib inline
plt.style.use('seaborn-notebook')
# ## Load Data
path = os.path.join('..', '..', 'download', 'mutation-matrix.tsv.bz2')
Y = pd.read_table(path, index_col=0)
# Read sample information and create a covariate TSV
url = 'https://github.com/cognoma/cancer-data/raw/54140cf6addc48260c9723213c40b628d7c861da/data/covariates.tsv'
covariate_df = pd.read_table(url, index_col=0)
covariate_df.head(2)
# ## Specify the type of classifier
# +
param_grid = {
'alpha': [10 ** x for x in range(-4, 2)],
'l1_ratio': [0, 0.05, 0.1, 0.2, 0.5, 0.8, 0.9, 0.95, 1],
}
clf = SGDClassifier(
random_state=0,
class_weight='balanced',
loss='log',
penalty='elasticnet'
)
# joblib is used to cross-validate in parallel by setting `n_jobs=-1` in GridSearchCV
# Supress joblib warning. See https://github.com/scikit-learn/scikit-learn/issues/6370
warnings.filterwarnings('ignore', message='Changing the shape of non-C contiguous array')
clf_grid = grid_search.GridSearchCV(estimator=clf, param_grid=param_grid, n_jobs=-1, scoring='roc_auc')
pipeline = make_pipeline(
StandardScaler(),
clf_grid
)
# -
# ## Specify covariates and outcomes
# +
def expand_grid(data_dict):
"""Create a dataframe from every combination of given values."""
rows = itertools.product(*data_dict.values())
return pd.DataFrame.from_records(rows, columns=data_dict.keys())
mutations = {
'7157': 'TP53', # tumor protein p53
'7428': 'VHL', # von Hippel-Lindau tumor suppressor
'29126': 'CD274', # CD274 molecule
'672': 'BRCA1', # BRCA1, DNA repair associated
'675': 'BRCA2', # BRCA2, DNA repair associated
'238': 'ALK', # anaplastic lymphoma receptor tyrosine kinase
'4221': 'MEN1', # menin 1
'5979': 'RET', # ret proto-oncogene
}
options = collections.OrderedDict()
options['mutation'] = list(mutations)
binary_options = [
'disease_covariate',
'organ_covariate',
'gender_covariate',
'mutation_covariate',
'survival_covariate'
]
for opt in binary_options:
options[opt] = [0, 1]
option_df = expand_grid(options)
option_df['symbol'] = option_df.mutation.map(mutations)
option_df.head(2)
# -
covariate_to_columns = {
'gender': covariate_df.columns[covariate_df.columns.str.startswith('gender')].tolist(),
'disease': covariate_df.columns[covariate_df.columns.str.startswith('disease')].tolist(),
'organ': covariate_df.columns[covariate_df.columns.str.contains('organ')].tolist(),
'mutation': covariate_df.columns[covariate_df.columns.str.contains('n_mutations')].tolist(),
'survival': ['alive', 'dead'],
}
# ## Compute performance
# +
def get_aurocs(X, y, series):
"""
Fit the classifier specified by series and add the cv, training, and testing AUROCs.
series is a row of option_df, which specificies the which covariates and mutation
status to use in the classifier.
"""
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=0)
series['positive_prevalence'] = np.mean(y)
pipeline.fit(X=X_train, y=y_train)
y_pred_train = pipeline.decision_function(X_train)
y_pred_test = pipeline.decision_function(X_test)
cv_score_df = grid_scores_to_df(clf_grid.grid_scores_)
series['mean_cv_auroc'] = cv_score_df.score.max()
series['training_auroc'] = roc_auc_score(y_train, y_pred_train)
series['testing_auroc'] = roc_auc_score(y_test, y_pred_test)
return series
def grid_scores_to_df(grid_scores):
"""
Convert a sklearn.grid_search.GridSearchCV.grid_scores_ attribute to
a tidy pandas DataFrame where each row is a hyperparameter-fold combinatination.
"""
rows = list()
for grid_score in grid_scores:
for fold, score in enumerate(grid_score.cv_validation_scores):
row = grid_score.parameters.copy()
row['fold'] = fold
row['score'] = score
rows.append(row)
df = pd.DataFrame(rows)
return df
# -
rows = list()
for i, series in option_df.iterrows():
columns = list()
for name, add_columns in covariate_to_columns.items():
if series[name + '_covariate']:
columns.extend(add_columns)
if not columns:
continue
X = covariate_df[columns]
y = Y[series.mutation]
rows.append(get_aurocs(X, y, series))
auroc_df = pd.DataFrame(rows)
auroc_df.sort_values(['symbol', 'testing_auroc'], ascending=[True, False], inplace=True)
auroc_df.head()
auroc_df.to_csv('auroc.tsv', index=False, sep='\t', float_format='%.5g')
# ## Covariate performance by mutation
# Filter for models which include all covariates
plot_df = auroc_df[auroc_df[binary_options].all(axis='columns')]
plot_df = pd.melt(plot_df, id_vars='symbol', value_vars=['mean_cv_auroc', 'training_auroc', 'testing_auroc'], var_name='kind', value_name='auroc')
grid = sns.factorplot(y='symbol', x='auroc', hue='kind', data=plot_df, kind="bar")
xlimits = grid.ax.set_xlim(0.5, 1)
|
explore/confounding/confounding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Temp1, Temp2 Temp3 e Temp4 são temperaturas medidas em diferentes partes da planta
# Target representa o estado da qualidade da amostra (temp1, temp2, temp3 e temp4)
# +
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
#p1_data_test_df = pd.read_csv('p1_data_test.csv',header=0)
df = pd.read_csv('../p1_data_train.csv',header=0)
print len(df)
pct = int(len(df)*0.5)
print pct
new_df = df[df.index > pct]
new_df
# -
def get_outliers_index(df, columns, gama = 1.5):
index_to_drop = []
for column in columns:
q2 = df[column].median()
q3 = df[df[column] > q2][column].median()
q1 = df[df[column] < q2][column].median()
IQR = q3 - q1
index_to_drop += list(df[(df[column] > q3 + gama*IQR) | (df[column] < q1 - gama*IQR)][column].index.values)
return list(np.unique(index_to_drop))
# +
df.head()
index_to_drop = get_outliers_index(df,['Temp1','Temp2','Temp3','Temp4'])
print df.shape
print len(index_to_drop)
print index_to_drop
df = df.drop(df.index[index_to_drop])
print df.shape
# -
data = {'name': ['Jason', 'Molly', 'Tina', 'Jake', 'Amy'],
'year': [2012, 2012, 2013, 2014, 2014],
'reports': [4, 24, 31, 2, 3]}
df = pd.DataFrame(data, index = ['Cochice', 'Pima', '<NAME>', 'Maricopa', 'Yuma'])
df
df.drop(df.index[[0,1,2]])
|
notebook/Desafio Radix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Condensed Matter Physics
#
# University of Sydney
# April 2020
#
# ## Lecture 17
# N and A separate
|
Lecture 18/Lecture 18.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ''
# language: python
# name: ''
# ---
# # Welding Example #02: TCP movements and Weaving
# In this example we will focus on more complex tcp movements along the workpiece and how to combine different motion shapes like weaving.
# ## Imports
# + nbsphinx="hidden"
# enable interactive plots on Jupyterlab with ipympl and jupyterlab-matplotlib installed
# # %matplotlib widget
# +
import numpy as np
import pandas as pd
from weldx import (
Q_,
CoordinateSystemManager,
Geometry,
LinearHorizontalTraceSegment,
LocalCoordinateSystem,
TimeSeries,
Trace,
WXRotation,
get_groove,
util,
welding,
)
from weldx.welding.util import sine
# -
# ## General setup
# We will use the same workpiece geometry as defined in the previous example.
# ### groove shape
groove = get_groove(
groove_type="VGroove",
workpiece_thickness=Q_(0.5, "cm"),
groove_angle=Q_(50, "deg"),
root_face=Q_(1, "mm"),
root_gap=Q_(1, "mm"),
)
# ### workpiece geometry
# +
# define the weld seam length in mm
seam_length = Q_(150, "mm")
# create a linear trace segment a the complete weld seam trace
trace_segment = LinearHorizontalTraceSegment(seam_length)
trace = Trace(trace_segment)
# create 3d workpiece geometry from the groove profile and trace objects
geometry = Geometry(groove.to_profile(width_default=Q_(5, "mm")), trace)
# rasterize geometry
profile_raster_width = "2mm" # resolution of each profile in mm
trace_raster_width = "30mm" # space between profiles in mm
geometry_data_sp = geometry.rasterize(
profile_raster_width=profile_raster_width, trace_raster_width=trace_raster_width
)
# -
# ### Coordinate system manager
# +
# crete a new coordinate system manager with default base coordinate system
csm = CoordinateSystemManager("base")
# add the workpiece coordinate system
csm.add_cs(
coordinate_system_name="workpiece",
reference_system_name="base",
lcs=trace.coordinate_system,
)
# add the geometry data of the specimen
csm.assign_data(
geometry.spatial_data(profile_raster_width, trace_raster_width),
"specimen",
"workpiece",
)
# -
# ## Movement definitions
# Like in the previous example we start by defining the general linear movement along the weld seam with a constant welding speed.
# +
tcp_start_point = Q_([5.0, 0.0, 2.0], "mm")
tcp_end_point = Q_([seam_length.m - 5.0, 0.0, 2.0], "mm")
v_weld = Q_(10, "mm/s")
s_weld = (tcp_end_point - tcp_start_point)[0] # length of the weld
t_weld = s_weld / v_weld
t_start = pd.Timedelta("0s")
t_end = pd.Timedelta(str(t_weld))
rot = WXRotation.from_euler("x", 180, degrees=True)
coords = [tcp_start_point.magnitude, tcp_end_point.magnitude]
tcp_wire = LocalCoordinateSystem(
coordinates=coords, orientation=rot, time=[t_start, t_end]
)
# -
# Let's add the linear movement to the coordinate system manager and see a simple plot:
csm.add_cs(
coordinate_system_name="tcp_wire", reference_system_name="workpiece", lcs=tcp_wire
)
csm
# +
def ax_setup(ax, rotate=170):
ax.legend()
ax.set_xlabel("x / mm")
ax.set_ylabel("y / mm")
ax.set_zlabel("z / mm")
ax.view_init(30, -10)
ax.set_ylim([-5.5, 5.5])
ax.view_init(30, rotate)
ax.legend()
color_dict = {
"tcp_sine": (255, 0, 0),
"tcp_wire_sine": (255, 0, 0),
"tcp_wire_sine2": (255, 0, 0),
"tcp_wire": (0, 150, 0),
"specimen": (0, 0, 255),
}
# -
ax = csm.plot(
coordinate_systems=["tcp_wire"],
colors=color_dict,
limits=[(0, 140), (-5, 5), (0, 12)],
show_vectors=False,
show_wireframe=True,
)
ax_setup(ax)
# ## add a sine wave to the TCP movement
# We now want to add a weaving motion along the y-axis (horizontal plane) of our TCP motion. We can define a general weaving motion using the `weldx.utility.sine` function that creates `TimeSeries` class.
ts_sine = sine(f=Q_(0.5 * 2 * np.pi, "Hz"), amp=Q_([0, 0.75, 0], "mm"))
# We now define a simple coordinate system that contains only the weaving motion.
tcp_sine = LocalCoordinateSystem(coordinates=ts_sine)
# One approach to combine the weaving motion with the existing linear `tcp_wire` movement is to use the coordinate system manager. We can add the `tcp_sine` coordinate system relative to the `tcp_wire` system:
csm.add_cs(
coordinate_system_name="tcp_sine", reference_system_name="tcp_wire", lcs=tcp_sine
)
csm
# Lets see the result:
t = pd.timedelta_range(start=t_start, end=t_end, freq="10ms")
ax = csm.plot(
coordinate_systems=["tcp_wire", "tcp_sine"],
colors=color_dict,
limits=[(0, 140), (-5, 5), (0, 12)],
show_origins=False,
show_vectors=False,
show_wireframe=True,
time=t,
)
ax_setup(ax)
# Here a little bit closer to see the actual sine wave:
ax = csm.plot(
coordinate_systems=["tcp_wire", "tcp_sine"],
colors=color_dict,
limits=[(0, 5), (-2, 2), (0, 12)],
show_origins=False,
show_vectors=False,
show_wireframe=False,
time=t,
)
ax_setup(ax)
# Another approach would be to combine both systems before adding them to the coordinate system manager. We can combine both coordinate systems using the __+__ operator to generate the superimposed weaving coordinate system.
tcp_wire_sine = tcp_sine.interp_time(t) + tcp_wire
# Note the difference in reference coordinate system compared to the first example.
csm.add_cs("tcp_wire_sine", "workpiece", tcp_wire_sine)
csm
# We get the same result:
ax = csm.plot(
coordinate_systems=["tcp_wire", "tcp_wire_sine"],
colors=color_dict,
limits=[(0, 140), (-5, 5), (0, 12)],
show_origins=False,
show_vectors=False,
show_wireframe=True,
)
ax_setup(ax)
ax = csm.plot(
coordinate_systems=["tcp_wire", "tcp_sine"],
colors=color_dict,
limits=[(0, 5), (-2, 2), (0, 12)],
show_origins=False,
show_vectors=False,
show_wireframe=False,
)
ax_setup(ax)
# Adding every single superposition step in the coordinate system manager can be more flexible and explicit, but will clutter the CSM instance for complex movements.
# ## plot with time interpolation
# Sometimes we might only be interested in a specific time range of the experiment or we want to change the time resolution. For this we can use the time interpolation methods of the CSM (or the coordinate systems).
#
# Let's say we want to weave only 8 seconds of our experiment (starting from 2020-04-20 10:03:00) but interpolate steps of 1 ms.
t_interp = pd.timedelta_range(start="3s", end="11s", freq="1ms")
ax = csm.interp_time(t_interp).plot(
coordinate_systems=["tcp_wire", "tcp_wire_sine"],
colors=color_dict,
limits=[(0, 140), (-5, 5), (0, 12)],
show_origins=False,
show_vectors=False,
show_wireframe=True,
)
ax_setup(ax)
# ## Adding a second weaving motion
# We now want to add a second weaving motion along the z-axis that only exists for a limited time. Lets generate the motion first:
ts_sine = sine(f=Q_(1 / 8 * 2 * np.pi, "Hz"), amp=Q_([0, 0, 1], "mm"))
# We define a new `LocalCoordinateSystem` and interpolate it our specified timestamps.
t = pd.timedelta_range(start="0s", end="8s", freq="25ms")
tcp_sine2 = LocalCoordinateSystem(coordinates=ts_sine).interp_time(t)
tcp_sine2
# adding all the movements together. We have to be careful with the time-axis in this case !
t_interp = pd.timedelta_range(
start=tcp_wire.time.index[0], end=tcp_wire.time.index[-1], freq="20ms"
)
tcp_wire_sine2 = (
tcp_sine2.interp_time(t_interp) + tcp_sine.interp_time(t_interp)
) + tcp_wire
csm.add_cs("tcp_wire_sine2", "workpiece", tcp_wire_sine2)
csm
ax = csm.plot(
coordinate_systems=["tcp_wire", "tcp_wire_sine2"],
colors=color_dict,
limits=[(0, 140), (-5, 5), (0, 12)],
show_origins=False,
show_vectors=False,
)
ax_setup(ax, rotate=110)
ax = csm.plot(
coordinate_systems=["tcp_wire", "tcp_wire_sine2"],
colors=color_dict,
limits=[(60, 100), (-2, 2), (0, 12)],
show_origins=False,
show_vectors=False,
)
ax_setup(ax, rotate=110)
# + [markdown] nbsphinx="hidden"
# ## K3D Visualization
# + nbsphinx="hidden"
csm.plot(
backend="k3d",
coordinate_systems=["tcp_wire_sine2"],
colors=color_dict,
show_vectors=False,
show_traces=True,
show_data_labels=False,
show_labels=False,
show_origins=True,
)
|
tutorials/welding_example_02_weaving.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy import signal, io
from matplotlib import pyplot as plt
from scipy import signal
from copy import deepcopy
num = 128
filename = './n0' + str(num) + '_LGS_trs.sav'
telemetry = io.readsav(filename)['a']
print(telemetry.dtype)
telemetry = io.readsav(filename)['a']
commands = deepcopy(telemetry['TTCOMMANDS'][0])
commands = commands - np.mean(commands, axis=0)
residuals = telemetry['RESIDUALWAVEFRONT'][0][:,349:351]
pol = residuals[1:] + commands[:-1]
plt.loglog(*signal.periodogram(pol[:,0], fs=1000))
plt.ylim(1e-10)
# +
s = 1000
P = np.zeros(s // 2 + 1,)
for i in range(10):
f, P_t = signal.periodogram(pol[s * i:s * (i + 1),0], fs=1000)
P += P_t
plt.figure(figsize=(10,10))
plt.plot(*signal.periodogram(pol[:s * 10,0], fs=1000))
plt.plot(f, P / 10)
plt.ylim(1e-9, 1e-3)
# -
plt.loglog(*signal.periodogram(commands[:,0], fs=1000))
plt.ylim(1e-10)
|
telemetry/quickviews.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: analysis
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
np.random.seed(42)
# ### A) Expected data format for SSAML code.
# Make sure you have a .csv file with the following columns and row formats. The format differs for non-survival-analysis and survival-analysis tasks. Within runner_power.sh you will find a boolean parameter survivalTF to be set (True for survival analysis, False for non-survival analysis), and a boolean parameter peopleTF (True for patient-level analysis and False for event-level analysis). This notebook is therefore a preprocessing guide/tutorial to re-format existing data to make it ready for SSAML algorithm and the runner_power.sh code. The analysis method is not determined here but with the aforementioned parameters in runner_power.sh.
#
# 1. 'regular', non-survival analysis model.
# columns:
# -- ID: unique patient identifier (integers)
# -- event: ground truth / label (integers)
# -- p: model output, event probability
#
# rows are data observations (i.e. one row per event/patient)
#
# 2. survival analysis model.
# columns:
# -- ID: unique patient identifier (integers)
# -- C: censhorhip information (i.e. 1 for censored, 0 for not censored)
# -- z is the z-score value, a covariate for Cox proportional hazard.
# -- T: time to event
#
# rows are data observations (i.e. one row per event/patient)
# ### B) sample datasets, as presented in the paper
#
# here, we present the format for three distinct tasks, as presented in the paper.
# #### B.1) seizure risk prediction ('seizure tracker (ST) data')
# +
# c = pd.read_csv(big_file,sep=',',names=['ID','szTF','AI','RMR'])
# uids = pd.unique(c.ID)
# c.rename(columns={'szTF':'event'},inplace=True)
# c.rename(columns={'AI':'p'},inplace=True)
# peopleTF=True
# survivalTF=False
# -
data = pd.DataFrame(columns=['ID', 'event', 'p'])
data['ID'] = np.arange(100)
data['event'] = np.random.randint(0, high=2, size=data.shape[0]) # binary outcome, high excluded
data['p'] = np.random.rand(data.shape[0]) # model output, probability values between 0 and 1
print(f'data shape: {data.shape}')
print(f'events contained: \n{data.event.value_counts()}')
data.head()
data.to_csv('sample_data_st.csv', index=False)
# #### B.2) covid hospitalization risk prediciton ('COVA dataset')
data['ID'] = np.arange(100)
data['event'] = np.random.randint(0, high=2, size=data.shape[0]) # binary outcome, high excluded
data['p'] = np.random.rand(data.shape[0]) # model output, probability values between 0 and 1
# +
data_raw = pd.read_csv('COVA-FAKE.csv', sep=',')
data = pd.DataFrame()
data['ID'] = np.array(range(data_raw.shape[0]))
event_categories = ['Prob-dead','Prob-ICU-MV','Prob-Hosp']
data['p'] = (data_raw[event_categories[0]] + data_raw[event_categories[1]] + data_raw[event_categories[2]])/100
data['event'] = 0.0 + (data_raw['actual']>0)
# -
print(f'data shape: {data.shape}')
print(f'events contained: \n{data.event.value_counts()}')
data.head()
data.to_csv('sample_data_cova.csv', index=False)
# #### B.3) Brain age - mortality risk prediction (survival analysis)
# This database file has the following columns: 'z','T','C', reflecting a z score (output of ML), T=time, and C=censored yes=1, no=0
# The ID numbers were not supplied, so row number can be used to produce a sequential ID number here in preprocessing.
# +
# c = pd.read_csv(big_file,sep=',')
# uids = uids = np.array(range(c.shape[0]))
# c['ID'] = uids
# peopleTF=True
# survivalTF=True
# -
data = pd.DataFrame(columns=['ID','z', 'T', 'C'])
data['ID'] = np.arange(100)
data['T'] = np.random.randint(0, high=21, size=data.shape[0]) # random integer values for time to event
data['C'] = np.random.randint(0, 2, size=data.shape[0]) # random binary censorship information Yes/No
data['z'] = np.random.normal(loc=0, scale=1, size=data.shape[0]) # random z-scored confounding variable.
print(f'data shape: {data.shape}')
print(f'events contained: \n{data.C.value_counts()}')
data.head()
data.to_csv('sample_data_bai_mortality.csv', index=False)
# +
## After you rrun runner_power with your modified parameters, you will get output files.
## if you had enabled "doEXTRA=True" in power.py, then you can plot the zing files as
## as follows or from modifying the make-power-pix.py
import matplotlib
#matplotlib.use("Agg")
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from scipy import stats
import os
import sys
import time
# CONSTANTS
# put whatever your local directory is that has your files from SSAML
mydir = '/Users/danisized/Documents/GitHub/SSAML/OUTcovaFAKE/'
# Note, the numLIST numbers here are hard coded for the number of patients/events we tested.
# change to whatever you like here
numLIST = [100,500,1000]
# FUNCTION DEFINITIONS
def getZING(prefixN,middleOne,numLIST):
# load up the ZING files and compose a pandas dataframe from it
print('Loading %s...' % prefixN)
for howmany in numLIST:
fn = prefixN + str(howmany).zfill(4) + '.csv'
dat = pd.read_csv(fn,sep=',',header=None)
dat.columns =['Slope',middleOne,'CIL']
dat['N'] = dat.Slope*0 + howmany
if howmany == numLIST[0]:
bigD = dat
else:
bigD = bigD.append(dat,ignore_index=True)
return bigD
def plotC(dat,numLIST,fig,ax,tName):
#plot the column number colNum
C=(.7,.7,.7)
colNum=1
plt.subplot(3,3,colNum)
ax[0,colNum-1] = sns.boxplot(x="N", y="Slope",fliersize=0,color=C, data=dat)
plt.grid(True,axis='y')
plt.ylim(0,2.1)
plt.xlabel('')
plt.title(tName)
plt.subplot(3,3,3+colNum)
ax[1,colNum-1] = sns.boxplot(x="N", y="C-index",fliersize=0,color=C, data=dat)
plt.grid(True,axis='y')
plt.ylim(0,1.1)
plt.xlabel('')
plt.subplot(3,3,6+colNum)
ax[2,colNum-1] = sns.boxplot(x="N", y="CIL",fliersize=0,color=C,data=dat)
plt.grid(True,axis='y')
plt.ylim(0,1.5)
ax[0,colNum-1].axes.xaxis.set_ticklabels([])
ax[1,colNum-1].axes.xaxis.set_ticklabels([])
ax[2,colNum-1].axes.xaxis.set_ticklabels(numLIST)
if colNum>1:
ax[0,colNum-1].axes.yaxis.set_ticklabels([])
ax[1,colNum-1].axes.yaxis.set_ticklabels([])
ax[2,colNum-1].axes.yaxis.set_ticklabels([])
ax[0,colNum-1].set_ylabel('')
ax[1,colNum-1].set_ylabel('')
ax[2,colNum-1].set_ylabel('')
return
# MAIN
os.chdir(mydir)
bigD = getZING('smallZ','C-index',numLIST)
print(bigD)
print('plotting...')
fig, ax = plt.subplots(3,2,sharex='col',sharey='row',figsize=(8,8))
plotC(bigD,numLIST,fig,ax,'BAI')
# make a little extra space between the subplots
fig.subplots_adjust(hspace=0.2)
plt.show()
print('saving...')
# jpeg in 300 dpi
fig.savefig('ZplotFull-v2.jpg',dpi=300)
|
data_format_preprocess.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import datetime
import seaborn as sns
import matplotlib.pyplot as plt
file=r'C:\Users\yashn\Desktop\Technocolabs\Project\MSFT.csv'
df=pd.read_csv(file,index_col="Date", parse_dates=True)
df.head()
df.shape
df.info()
df.isnull().sum()
df[df.duplicated()]
df['Volume'].value_counts()
df.describe()
plt.figure(figsize=(14,6))
sns.lineplot(data=df)
sns.scatterplot(x=df['High'], y=df['Low'])
sns.scatterplot(x=df['Open'], y=df['Close'])
sns.distplot(a=df['Volume'], kde=False)
|
Data cleaning of MSFT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # English Premier League VAR Analysis
# ## Part 3 - Analysis of Teams' VAR Incident Statistics
# ___
# **Questions**
# - Do the big 6 teams have more decisions in (or against) their favour? **[DONE]**
# - Which team is involved in the most VAR incidents? Which team had the most FOR decisions, and which team had the most AGAINST decisions **[DONE]**
# - What is the impact of VAR for decisions on the team's final league position and points tally? **[DONE]**
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
import re
import difflib
import seaborn as sns
from collections import Counter
from datetime import datetime as dt
from scipy.stats import ttest_ind
pd.options.display.max_rows = 500
# -
# ### Data Processing
file_date = '20210524'
# Import team stats dataset
teamstats_df = pd.read_csv(f'data/EPL_VAR_Team_Stats_Raw_{file_date}.csv')
# +
# NOT using this method because of misspellings in incidents column (from original data). To use
# VAR decisions tally directly from the website
# # Add column of decisions FOR and decisions AGAINST (from incidents data)
# incidents_df_raw = pd.read_csv('data/EPL_VAR_Incidents_Processed_20210510.csv')
# incidents_df_1 = incidents_df_raw[['team', 'year', 'team_decision', 'incident']]
# incidents_df_2 = incidents_df_raw[['opposition', 'year', 'opposition_decision', 'incident']]
# incidents_df_2.columns = incidents_df_1.columns
# incidents_df = pd.concat([incidents_df_1, incidents_df_2])
# incidents_df.reset_index(drop=True, inplace=True)
# incidents_df
# # Create pivot table and reset index
# decision_count = incidents_df.pivot_table(index=['team', 'year'], columns='team_decision', aggfunc='size', fill_value=0)
# decision_count = decision_count.rename_axis(None, axis=1).reset_index(drop=False)
# decision_count.columns = ['team_name', 'year', 'decisions_against',
# 'decisions_for', 'decisions_neutral']
# Clean up decisions and incidents columns
# for i, row in incidents_df.iterrows():
# if row['team_decision'] not in ['For', 'Against', 'Neutral']:
# incidents_df.loc[i, 'team_decision'] = 'Neutral'
# text = row['incident']
# text_clean = text.split(" - ")[0] # Remove the - FOR and - AGAINST strings
# incidents_df.loc[i, 'incident'] = text_clean
# -
# Create helper function to rename team names so that they can be merged
def rename_team_names(df, col_name):
for i, row in df.iterrows():
if row[col_name] == 'Brighton':
df.loc[i, col_name] = 'Brighton & Hove Albion'
if row[col_name] == 'Leicester':
df.loc[i, col_name] = 'Leicester City'
if row[col_name] == 'Man City':
df.loc[i, col_name] = 'Manchester City'
if row[col_name] == 'Man United':
df.loc[i, col_name] = 'Manchester United'
if row[col_name] == 'Newcastle':
df.loc[i, col_name] = 'Newcastle United'
if row[col_name] == 'Tottenham':
df.loc[i, col_name] = 'Tottenham Hotspur'
if row[col_name] == 'WBA':
df.loc[i, col_name] = 'West Bromwich Albion'
if row[col_name] == 'West Brom':
df.loc[i, col_name] = 'West Bromwich Albion'
if row[col_name] == 'Norwich':
df.loc[i, col_name] = 'Norwich City'
if row[col_name] == 'AFC Bournemouth':
df.loc[i, col_name] = 'Bournemouth'
if row[col_name] == 'Wolves':
df.loc[i, col_name] = 'Wolverhampton Wanderers'
if row[col_name] == 'Leeds':
df.loc[i, col_name] = 'Leeds United'
if row[col_name] == 'West Ham':
df.loc[i, col_name] = 'West Ham United'
return df
decision_count = pd.read_csv(f'./data/EPL_VAR_Decisions_{file_date}.csv')
decision_count = rename_team_names(decision_count, 'team')
decision_count = decision_count.rename(columns={'team': 'team_name'})
decision_count
# +
# Combine decisions count to team stats dataframe
teamstats_df = rename_team_names(teamstats_df, 'team_name')
teamstats_df = pd.merge(teamstats_df, decision_count, how = 'left',
on = ['team_name', 'year'])
# Rearrange columns
cols_to_shift = ['team_name', 'year', 'net_score', 'decisions_for', 'decisions_against']
teamstats_df = teamstats_df[cols_to_shift + [c for c in teamstats_df if c not in cols_to_shift]]
teamstats_df.sort_values(by=['team_name'])
# -
# The `net_score` column serves as an additional sanity check to ensure that the dataframe is processed properly. This is done through asserting `net_score` = `decisions_for` - `decisions_against`
# ### Feature Description
# **Specifics on each of the features:**
#
# `team_name`: Name of the EPL team (categorical: 'Arsenal', 'Burnley', 'Chelsea', ...)
#
# `net_score`: Net score to assess benefit of VAR overturns to the team (Calculated by **decisions_for** - **decisions_against**) (numeric)
#
# `overturns_total`: Total number of overturned decisions by VAR (numeric)
#
# `overturns_rejected`:Number of decisions rejected by the referee at the review screen (numeric)
#
# `leading_to_goals_for`: Number of VAR decisions leading to goals for the team (numeric)
#
# `leading_to_goals_against`: Number of VAR decisions leading to goals against the team (numeric)
#
# `disallowed_goals_for`: Number of VAR decisions resulting in disallowed goals for the team (Detriment) (numeric)
#
# `disallowed_goals_against`: Number of VAR decisions resulting in disallowed goals for the team's opposition (Benefit) (numeric)
#
# `net_goal_score`: Net goal score (Calculated by **leading_to_goals_for - leading_to_goals_against + disallowed_goals_against - disallowed_goals_for**) (numeric)
#
# `subj_decisions_for`: Number of subjective VAR decisions (i.e. referee decision) for the team (numeric)
#
# `subj_decisions_against`: Number of subjective VAR decisions (i.e. referee decision) against the team (numeric)
#
# `net_subjective_score`: Net subjective score (`subj_decisions_for` minus `subj_decisions_against` ) (numeric)
#
# `penalties_for`: Number of VAR decisions resulting in penalties for the team (numeric)
#
# `penalties_against`: Number of VAR decisions resulting in penalties against the team (numeric)
#
# `year`: EPL Season (categorical: '2019/2020', '2020/2021')
#
# `decisions_against`: Number of VAR decisions against the team (numeric)
#
# `decisions_for`: Number of VAR decisions for the team (numeric)
#
# `decisions_neutral`: Number of VAR decisions neutral to the team (numeric)
# ___
# ## Analysis
#
# ### (1) Do VAR overturn decisions favour the big six teams?
#
# ##### Net score
# Categorize big six teams (based on ESL incident in 2021)
big_six_teams = ['Arsenal', 'Chelsea', 'Liverpool', 'Manchester City', 'Manchester United', 'Tottenham Hotspur']
teamstats_df['big_six'] = np.where(teamstats_df['team_name'].isin(big_six_teams), 'Yes', 'No')
teamstats_df
df_net_score = pd.DataFrame(teamstats_df.groupby(['team_name', 'big_six'])['net_score'].agg('sum')).reset_index(drop=False)
df_net_score
big_6_net_score = df_net_score[df_net_score['big_six'] == 'Yes']
big_6_net_score['net_score'].describe()
non_big_6_net_score = df_net_score[df_net_score['big_six'] == 'No']
non_big_6_net_score['net_score'].describe()
ax = sns.boxplot(x="big_six", y="net_score", data=df_net_score)
# If you have two independent samples but you do not know that they have equal variance, you can use Welch's t-test.
#
# Reference: https://stackoverflow.com/questions/13404468/t-test-in-pandas/13413842
# Welch's t-test for net score
ttest_ind(big_6_net_score['net_score'], non_big_6_net_score['net_score'], equal_var=False)
# p-value returned from Welch's t-test = 0.749 (i.e. no statistically significant difference in net score between big six and non big six teams
# ##### Net Goal score
df_net_goal_score = pd.DataFrame(teamstats_df.groupby(['team_name', 'big_six'])['net_goal_score'].agg('sum')).reset_index(drop=False)
df_net_goal_score
ax = sns.boxplot(x="big_six", y="net_goal_score", data=df_net_goal_score)
big_6_net_goal_score = df_net_goal_score[df_net_goal_score['big_six'] == 'Yes']
big_6_net_goal_score['net_goal_score'].describe()
non_big_6_net_goal_score = df_net_goal_score[df_net_goal_score['big_six'] == 'No']
non_big_6_net_goal_score['net_goal_score'].describe()
# Welch t-test for net goal score
from scipy.stats import ttest_ind
ttest_ind(big_6_net_goal_score['net_goal_score'], non_big_6_net_goal_score['net_goal_score'], equal_var=False)
# ##### Net Subjective score
df_net_subj_score = pd.DataFrame(teamstats_df.groupby(['team_name', 'big_six'])['net_subjective_score'].agg('sum')).reset_index(drop=False)
df_net_subj_score
ax = sns.boxplot(x="big_six", y="net_subjective_score", data=df_net_subj_score)
big_6_net_subj_score = df_net_subj_score[df_net_subj_score['big_six'] == 'Yes']
big_6_net_subj_score['net_subjective_score'].describe()
non_big_6_net_subj_score = df_net_subj_score[df_net_subj_score['big_six'] == 'No']
non_big_6_net_subj_score['net_subjective_score'].describe()
# Welch t-test for net subjective score
ttest_ind(big_6_net_subj_score['net_subjective_score'], non_big_6_net_subj_score['net_subjective_score'], equal_var=False)
# **Summary**
# p-values returned from Welch's t-test AND Student's t-test for all 3 scores (net score, net goal score, net subjective score) are all >0.05 (i.e. no statistically significant difference in net score between big six and non big six teams)
# ___
# ### (2) Which EPL teams were involved in most VAR overturn incidents, had the most FOR decisions, and the most AGAINST decisions?
# ##### Count of VAR incident overturn involvement
df_decisions_total = pd.DataFrame(teamstats_df.groupby(['team_name'])['overturns_total'].agg('sum')).reset_index(drop=False)
df_decisions_total.sort_values(by='overturns_total', ascending=False, inplace=True)
df_decisions_total.reset_index(drop=True, inplace=True)
df_decisions_total
# ##### Count of VAR decision for
df_decisions_for = pd.DataFrame(teamstats_df.groupby(['team_name'])['decisions_for'].agg('sum')).reset_index(drop=False)
df_decisions_for.sort_values(by='decisions_for', ascending=False, inplace=True)
df_decisions_for.reset_index(drop=True, inplace=True)
df_decisions_for
# ##### Count of VAR decision against
df_decisions_against = pd.DataFrame(teamstats_df.groupby(['team_name'])['decisions_against'].agg('sum')).reset_index(drop=False)
df_decisions_against.sort_values(by='decisions_against', ascending=False, inplace=True)
df_decisions_against.reset_index(drop=True, inplace=True)
df_decisions_against
# ##### Percentage For (based on total VAR overturn incidents)
df_decisions_for_percent = pd.DataFrame(teamstats_df.groupby(['team_name'])['decisions_for', 'overturns_total'].agg('sum')).reset_index(drop=False)
df_decisions_for_percent['decisions_for_percent'] = round((df_decisions_for_percent['decisions_for']/df_decisions_for_percent['overturns_total']) * 100, 1)
df_decisions_for_percent.sort_values(by='decisions_for_percent', inplace=True, ascending=False)
df_decisions_for_percent.reset_index(drop=True, inplace=True)
df_decisions_for_percent
# This leads to the next question, do these VAR decisions have a correlation with the team's league position and points tally?
# ___
# ### (3) Do VAR decisions correlate with EPL league positions and points tally?
epl_table_df = pd.read_csv(f'./data/EPL_Table_{file_date}.csv')
epl_table_df = rename_team_names(epl_table_df, 'team')
epl_table_df.columns = ['team_name', 'year', 'position', 'points']
epl_table_df.head()
df_decisions_for_percent_yearly = pd.DataFrame(teamstats_df.groupby(['team_name', 'year'])['decisions_for', 'overturns_total'].agg('sum')).reset_index(drop=False)
df_decisions_for_percent_yearly['decisions_for_percent'] = round((df_decisions_for_percent_yearly['decisions_for']/df_decisions_for_percent_yearly['overturns_total']) * 100, 1)
df_decisions_for_percent_yearly.sort_values(by='team_name', inplace=True, ascending=True)
df_decisions_for_percent_yearly.reset_index(drop=True, inplace=True)
df_decisions_for_percent_yearly
table_decision_for_df = pd.merge(epl_table_df, df_decisions_for_percent_yearly, how='left',
on=['team_name', 'year']).reset_index(drop=True)
table_decision_for_df
sns.scatterplot(data=table_decision_for_df, x="decisions_for_percent", y="points");
ax = sns.scatterplot(data=table_decision_for_df, x="decisions_for_percent", y="position");
ax.invert_yaxis()
# Conclusion: No correlation between VAR decision-for ratio and the final EPL points tally (or league position)
|
03_Team_Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Further Python Basics
names = ['alice', 'jonathan', 'bobby']
ages = [24, 32, 45]
ranks = ['kinda cool', 'really cool', 'insanely cool']
for (name, age, rank) in zip(names, ages, ranks):
print name, age, rank
for index, (name, age, rank) in enumerate(zip(names, ages, ranks)):
print index, name, age, rank
# +
# return, esc, shift+enter, ctrl+enter
# text keyboard shortcuts -- cmd > (right), < left,
# option delete (deletes words)
# type "h" for help
# tab
# shift-tab
# keyboard shortcuts
# - a, b, y, m, dd, h, ctrl+shift+-
# +
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
import matplotlib.pyplot as plt
# no pylab
import seaborn as sns
sns.set_context('talk')
sns.set_style('darkgrid')
plt.rcParams['figure.figsize'] = 12, 8 # plotsize
import numpy as np
# don't do `from numpy import *`
import pandas as pd
# -
# If you have a specific function that you'd like to import
from numpy.random import randn
x = np.arange(100)
y = np.sin(x)
plt.plot(x, y);
# %matplotlib notebook
x = np.arange(10)
y = np.sin(x)
plt.plot(x, y)#;
# ## Magics!
#
# - % and %% magics
# - interact
# - embed image
# - embed links, youtube
# - link notebooks
# Check out http://matplotlib.org/gallery.html select your favorite.
# + language="bash"
# for num in {1..5}
# do
# for infile in *;
# do
# echo $num $infile
# done
# wc $infile
# done
# -
print "hi"
# !pwd
# !ping google.com
this_is_magic = "Can you believe you can pass variables and strings like this?"
# !echo $this_is_magic
hey
# # Numpy
#
# If you have arrays of numbers, use `numpy` or `pandas` (built on `numpy`) to represent the data. Tons of very fast underlying code.
# +
x = np.arange(10000)
print x # smart printing
# -
print x[0] # first element
print x[-1] # last element
print x[0:5] # first 5 elements (also x[:5])
print x[:] # "Everything"
print x[-5:] # last five elements
print x[-5:-2]
print x[-5:-1] # not final value -- not inclusive on right
x = np.random.randint(5, 5000, (3, 5))
x
np.sum(x)
x.sum()
np.sum(x)
np.sum(x, axis=0)
np.sum(x, axis=1)
x.sum(axis=1)
# Multi dimension array slice with a comma
x[:, 2]
y = np.linspace(10, 20, 11)
y
# +
# np.linspace?
# -
np.linspace()
# shift-tab; shift-tab-tab
np.
def does_it(first=x, second=y):
"""This is my doc"""
pass
y[[3, 5, 7]]
does_it()
num = 3000
x = np.linspace(1.0, 300.0, num)
y = np.random.rand(num)
z = np.sin(x)
np.savetxt("example.txt", np.transpose((x, y, z)))
# %less example.txt
# !wc example.txt
# !head example.txt
# +
#Not a good idea
a = []
b = []
for line in open("example.txt", 'r'):
a.append(line[0])
b.append(line[2])
a[:10] # Whoops!
# +
a = []
b = []
for line in open("example.txt", 'r'):
line = line.split()
a.append(line[0])
b.append(line[2])
a[:10] # Strings!
# +
a = []
b = []
for line in open("example.txt", 'r'):
line = line.split()
a.append(float(line[0]))
b.append(float(line[2]))
a[:10] # Lists!
# -
# Do this!
a, b = np.loadtxt("example.txt", unpack=True, usecols=(0,2))
a
# ## Matplotlib and Numpy
#
from numpy.random import randn
num = 50
x = np.linspace(2.5, 300, num)
y = randn(num)
plt.scatter(x, y)
y > 1
y[y > 1]
y[(y < 1) & (y > -1)]
plt.scatter(x, y, c='b', s=50)
plt.scatter(x[(y < 1) & (y > -1)], y[(y < 1) & (y > -1)], c='r', s=50)
y[~((y < 1) & (y > -1))] = 1.0
plt.scatter(x, y, c='b')
plt.scatter(x, np.clip(y, -0.5, 0.5), color='red')
num = 350
slope = 0.3
x = randn(num) * 50. + 150.0
y = randn(num) * 5 + x * slope
plt.scatter(x, y, c='b')
# plt.scatter(x[(y < 1) & (y > -1)], y[(y < 1) & (y > -1)], c='r')
# np.argsort, np.sort, complicated index slicing
dframe = pd.DataFrame({'x': x, 'y': y})
g = sns.jointplot('x', 'y', data=dframe, kind="reg")
# ## Grab Python version of ggplot http://ggplot.yhathq.com/
from ggplot import ggplot, aes, geom_line, stat_smooth, geom_dotplot, geom_point
ggplot(aes(x='x', y='y'), data=dframe) + geom_point() + stat_smooth(colour='blue', span=0.2)
|
deliver/08-More_basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Basic training functionality
# + hide_input=true
from fastai.basic_train import *
from fastai.gen_doc.nbdoc import *
from fastai.vision import *
from fastai.distributed import *
# -
# [`basic_train`](/basic_train.html#basic_train) wraps together the data (in a [`DataBunch`](/basic_data.html#DataBunch) object) with a PyTorch model to define a [`Learner`](/basic_train.html#Learner) object. Here the basic training loop is defined for the [`fit`](/basic_train.html#fit) method. The [`Learner`](/basic_train.html#Learner) object is the entry point of most of the [`Callback`](/callback.html#Callback) objects that will customize this training loop in different ways. Some of the most commonly used customizations are available through the [`train`](/train.html#train) module, notably:
#
# - [`Learner.lr_find`](/train.html#lr_find) will launch an LR range test that will help you select a good learning rate.
# - [`Learner.fit_one_cycle`](/train.html#fit_one_cycle) will launch a training using the 1cycle policy to help you train your model faster.
# - [`Learner.to_fp16`](/train.html#to_fp16) will convert your model to half precision and help you launch a training in mixed precision.
# + hide_input=true
show_doc(Learner, title_level=2)
# -
# The main purpose of [`Learner`](/basic_train.html#Learner) is to train `model` using [`Learner.fit`](/basic_train.html#Learner.fit). After every epoch, all *metrics* will be printed and also made available to callbacks.
#
# The default weight decay will be `wd`, which will be handled using the method from [Fixing Weight Decay Regularization in Adam](https://arxiv.org/abs/1711.05101) if `true_wd` is set (otherwise it's L2 regularization). If `true_wd` is set it will affect all optimizers, not only Adam. If `bn_wd` is `False`, then weight decay will be removed from batchnorm layers, as recommended in [Accurate, Large Minibatch SGD: Training ImageNet in 1 Hour](https://arxiv.org/abs/1706.02677). If `train_bn`, batchnorm layer learnable params are trained even for frozen layer groups.
#
# To use [discriminative layer training](#Discriminative-layer-training), pass a list of [`nn.Module`](https://pytorch.org/docs/stable/nn.html#torch.nn.Module) as `layer_groups`; each [`nn.Module`](https://pytorch.org/docs/stable/nn.html#torch.nn.Module) will be used to customize the optimization of the corresponding layer group.
#
# If `path` is provided, all the model files created will be saved in `path`/`model_dir`; if not, then they will be saved in `data.path`/`model_dir`.
#
# You can pass a list of [`callback`](/callback.html#callback)s that you have already created, or (more commonly) simply pass a list of callback functions to `callback_fns` and each function will be called (passing `self`) on object initialization, with the results stored as callback objects. For a walk-through, see the [training overview](/training.html) page. You may also want to use an [application](applications.html) specific model. For example, if you are dealing with a vision dataset, here the MNIST, you might want to use the [`cnn_learner`](/vision.learner.html#cnn_learner) method:
# + hide_input=false
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
learn = cnn_learner(data, models.resnet18, metrics=accuracy)
# -
# ### Model fitting methods
# + hide_input=true
show_doc(Learner.lr_find)
# -
# Runs the learning rate finder defined in [`LRFinder`](/callbacks.lr_finder.html#LRFinder), as discussed in [Cyclical Learning Rates for Training Neural Networks](https://arxiv.org/abs/1506.01186).
learn.lr_find()
learn.recorder.plot()
# + hide_input=true
show_doc(Learner.fit)
# -
# Uses [discriminative layer training](#Discriminative-layer-training) if multiple learning rates or weight decay values are passed. To control training behaviour, use the [`callback`](/callback.html#callback) system or one or more of the pre-defined [`callbacks`](/callbacks.html#callbacks).
learn.fit(1)
# + hide_input=true
show_doc(Learner.fit_one_cycle)
# -
# Use cycle length `cyc_len`, a per cycle maximal learning rate `max_lr`, momentum `moms`, division factor `div_factor`, weight decay `wd`, and optional callbacks [`callbacks`](/callbacks.html#callbacks). Uses the [`OneCycleScheduler`](/callbacks.one_cycle.html#OneCycleScheduler) callback. Please refer to [What is 1-cycle](/callbacks.one_cycle.html#What-is-1cycle?) for a conceptual background of 1-cycle training policy and more technical details on what do the method's arguments do.
learn.fit_one_cycle(1)
# ### See results
# + hide_input=true
show_doc(Learner.predict)
# -
# `predict` can be used to get a single prediction from the trained learner on one specific piece of data you are interested in.
learn.data.train_ds[0]
# Each element of the dataset is a tuple, where the first element is the data itself, while the second element is the target label. So to get the data, we need to index one more time.
data = learn.data.train_ds[0][0]
data
pred = learn.predict(data)
pred
# The first two elements of the tuple are, respectively, the predicted class and label. Label here is essentially an internal representation of each class, since class name is a string and cannot be used in computation. To check what each label corresponds to, run:
learn.data.classes
# So category 0 is 3 while category 1 is 7.
probs = pred[2]
# The last element in the tuple is the predicted probabilities. For a categorization dataset, the number of probabilities returned is the same as the number of classes; `probs[i]` is the probability that the `item` belongs to `learn.data.classes[i]`.
learn.data.valid_ds[0][0]
# You could always check yourself if the probabilities given make sense.
# + hide_input=true
show_doc(Learner.get_preds)
# -
# It will run inference using the learner on all the data in the `ds_type` dataset and return the predictions; if `n_batch` is not specified, it will run the predictions on the default batch size. If `with_loss`, it will also return the loss on each prediction.
# Here is how you check the default batch size.
learn.data.batch_size
preds = learn.get_preds()
preds
# The first element of the tuple is a tensor that contains all the predictions.
preds[0]
# While the second element of the tuple is a tensor that contains all the target labels.
preds[1]
preds[1][0]
# For more details about what each number mean, refer to the documentation of [`predict`](/basic_train.html#predict).
#
# Since [`get_preds`](/basic_train.html#get_preds) gets predictions on all the data in the `ds_type` dataset, here the number of predictions will be equal to the number of data in the validation dataset.
len(learn.data.valid_ds)
len(preds[0]), len(preds[1])
# To get predictions on the entire training dataset, simply set the `ds_type` argument accordingly.
learn.get_preds(ds_type=DatasetType.Train)
# To also get prediction loss along with the predictions and the targets, set `with_loss=True` in the arguments.
learn.get_preds(with_loss=True)
# Note that the third tensor in the output tuple contains the losses.
# + hide_input=true
show_doc(Learner.validate)
# -
# Return the calculated loss and the metrics of the current model on the given data loader `dl`. The default data loader `dl` is the validation dataloader.
# You can check the default metrics of the learner using:
str(learn.metrics)
learn.validate()
learn.validate(learn.data.valid_dl)
learn.validate(learn.data.train_dl)
# + hide_input=true
show_doc(Learner.show_results)
# -
# Note that the text number on the top is the ground truth, or the target label, the one in the middle is the prediction, while the image number on the bottom is the image data itself.
learn.show_results()
learn.show_results(ds_type=DatasetType.Train)
# + hide_input=true
show_doc(Learner.pred_batch)
# -
# Note that the number of predictions given equals to the batch size.
learn.data.batch_size
preds = learn.pred_batch()
len(preds)
# Since the total number of predictions is too large, we will only look at a part of them.
preds[:10]
item = learn.data.train_ds[0][0]
item
batch = learn.data.one_item(item)
batch
learn.pred_batch(batch=batch)
# + hide_input=true
show_doc(Learner.interpret, full_name='interpret')
# + hide_input=true
jekyll_note('This function only works in the vision application.')
# -
# For more details, refer to [ClassificationInterpretation](/vision.learner.html#ClassificationInterpretation)
# ### Model summary
# + hide_input=true
show_doc(Learner.summary)
# -
# ### Test time augmentation
# + hide_input=true
show_doc(Learner.TTA, full_name = 'TTA')
# -
# Applies Test Time Augmentation to `learn` on the dataset `ds_type`. We take the average of our regular predictions (with a weight `beta`) with the average of predictions obtained through augmented versions of the training set (with a weight `1-beta`). The transforms decided for the training set are applied with a few changes `scale` controls the scale for zoom (which isn't random), the cropping isn't random but we make sure to get the four corners of the image. Flipping isn't random but applied once on each of those corner images (so that makes 8 augmented versions total).
# ### Gradient clipping
# + hide_input=true
show_doc(Learner.clip_grad)
# -
# ### Mixed precision training
# + hide_input=true
show_doc(Learner.to_fp16)
# -
# Uses the [`MixedPrecision`](/callbacks.fp16.html#MixedPrecision) callback to train in mixed precision (i.e. forward and backward passes using fp16, with weight updates using fp32), using all [NVIDIA recommendations](https://docs.nvidia.com/deeplearning/sdk/mixed-precision-training/index.html) for ensuring speed and accuracy.
# + hide_input=true
show_doc(Learner.to_fp32)
# -
# ### Distributed training
# If you want to use ditributed training or [`torch.nn.DataParallel`](https://pytorch.org/docs/stable/nn.html#torch.nn.DataParallel) these will directly wrap the model for you.
# + hide_input=true
show_doc(Learner.to_distributed, full_name='to_distributed')
# + hide_input=true
show_doc(Learner.to_parallel, full_name='to_parallel')
# -
# ### Discriminative layer training
# When fitting a model you can pass a list of learning rates (and/or weight decay amounts), which will apply a different rate to each *layer group* (i.e. the parameters of each module in `self.layer_groups`). See the [Universal Language Model Fine-tuning for Text Classification](https://arxiv.org/abs/1801.06146) paper for details and experimental results in NLP (we also frequently use them successfully in computer vision, but have not published a paper on this topic yet). When working with a [`Learner`](/basic_train.html#Learner) on which you've called `split`, you can set hyperparameters in four ways:
#
# 1. `param = [val1, val2 ..., valn]` (n = number of layer groups)
# 2. `param = val`
# 3. `param = slice(start,end)`
# 4. `param = slice(end)`
#
# If we chose to set it in way 1, we must specify a number of values exactly equal to the number of layer groups. If we chose to set it in way 2, the chosen value will be repeated for all layer groups. See [`Learner.lr_range`](/basic_train.html#Learner.lr_range) for an explanation of the `slice` syntax).
#
# Here's an example of how to use discriminative learning rates (note that you don't actually need to manually call [`Learner.split`](/basic_train.html#Learner.split) in this case, since fastai uses this exact function as the default split for `resnet18`; this is just to show how to customize it):
# creates 3 layer groups
learn.split(lambda m: (m[0][6], m[1]))
# only randomly initialized head now trainable
learn.freeze()
learn.fit_one_cycle(1)
# all layers now trainable
learn.unfreeze()
# optionally, separate LR and WD for each group
learn.fit_one_cycle(1, max_lr=(1e-4, 1e-3, 1e-2), wd=(1e-4,1e-4,1e-1))
# + hide_input=true
show_doc(Learner.lr_range)
# -
# Rather than manually setting an LR for every group, it's often easier to use [`Learner.lr_range`](/basic_train.html#Learner.lr_range). This is a convenience method that returns one learning rate for each layer group. If you pass `slice(start,end)` then the first group's learning rate is `start`, the last is `end`, and the remaining are evenly geometrically spaced.
#
# If you pass just `slice(end)` then the last group's learning rate is `end`, and all the other groups are `end/10`. For instance (for our learner that has 3 layer groups):
learn.lr_range(slice(1e-5,1e-3)), learn.lr_range(slice(1e-3))
# + hide_input=true
show_doc(Learner.unfreeze)
# -
# Sets every layer group to *trainable* (i.e. `requires_grad=True`).
# + hide_input=true
show_doc(Learner.freeze)
# -
# Sets every layer group except the last to *untrainable* (i.e. `requires_grad=False`).
#
# What does '**the last layer group**' mean?
#
# In the case of transfer learning, such as `learn = cnn_learner(data, models.resnet18, metrics=error_rate)`, `learn.model`will print out two large groups of layers: (0) Sequential and (1) Sequental in the following structure. We can consider the last conv layer as the break line between the two groups.
# ```
# Sequential(
# (0): Sequential(
# (0): Conv2d(3, 64, kernel_size=(7, 7), stride=(2, 2), padding=(3, 3), bias=False)
# (1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
# (2): ReLU(inplace)
# ...
#
# (conv2): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
# (bn2): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
# )
# )
# )
# (1): Sequential(
# (0): AdaptiveConcatPool2d(
# (ap): AdaptiveAvgPool2d(output_size=1)
# (mp): AdaptiveMaxPool2d(output_size=1)
# )
# (1): Flatten()
# (2): BatchNorm1d(1024, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
# (3): Dropout(p=0.25)
# (4): Linear(in_features=1024, out_features=512, bias=True)
# (5): ReLU(inplace)
# (6): BatchNorm1d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
# (7): Dropout(p=0.5)
# (8): Linear(in_features=512, out_features=12, bias=True)
# )
# )
# ```
#
# `learn.freeze` freezes the first group and keeps the second or last group free to train, including multiple layers inside (this is why calling it 'group'), as you can see in `learn.summary()` output. How to read the table below, please see [model summary docs](/callbacks.hooks.html#model_summary).
#
# ```
# ======================================================================
# Layer (type) Output Shape Param # Trainable
# ======================================================================
# ...
# ...
# ...
# ______________________________________________________________________
# Conv2d [1, 512, 4, 4] 2,359,296 False
# ______________________________________________________________________
# BatchNorm2d [1, 512, 4, 4] 1,024 True
# ______________________________________________________________________
# AdaptiveAvgPool2d [1, 512, 1, 1] 0 False
# ______________________________________________________________________
# AdaptiveMaxPool2d [1, 512, 1, 1] 0 False
# ______________________________________________________________________
# Flatten [1, 1024] 0 False
# ______________________________________________________________________
# BatchNorm1d [1, 1024] 2,048 True
# ______________________________________________________________________
# Dropout [1, 1024] 0 False
# ______________________________________________________________________
# Linear [1, 512] 524,800 True
# ______________________________________________________________________
# ReLU [1, 512] 0 False
# ______________________________________________________________________
# BatchNorm1d [1, 512] 1,024 True
# ______________________________________________________________________
# Dropout [1, 512] 0 False
# ______________________________________________________________________
# Linear [1, 12] 6,156 True
# ______________________________________________________________________
#
# Total params: 11,710,540
# Total trainable params: 543,628
# Total non-trainable params: 11,166,912
# ```
#
# + hide_input=true
show_doc(Learner.freeze_to)
# -
# From above we know what is layer group, but **what exactly does `freeze_to` do behind the scenes**?
#
# The `freeze_to` source code can be understood as the following pseudo-code:
# ```python
# def freeze_to(self, n:int)->None:
# for g in self.layer_groups[:n]: freeze
# for g in self.layer_groups[n:]: unfreeze
# ```
# In other words, for example, `freeze_to(1)` is to freeze layer group 0 and unfreeze the rest layer groups, and `freeze_to(3)` is to freeze layer groups 0, 1, and 2 but unfreeze the rest layer groups (if there are more layer groups left).
#
# Both `freeze` and `unfreeze` [sources](https://github.com/fastai/fastai/blob/master/fastai/basic_train.py#L216) are defined using `freeze_to`:
# - When we say `freeze`, we mean that in the specified layer groups the [`requires_grad`](/torch_core.html#requires_grad) of all layers with weights (except BatchNorm layers) are set `False`, so the layer weights won't be updated during training.
# - when we say `unfreeze`, we mean that in the specified layer groups the [`requires_grad`](/torch_core.html#requires_grad) of all layers with weights (except BatchNorm layers) are set `True`, so the layer weights will be updated during training.
# + hide_input=true
show_doc(Learner.split)
# -
# A convenience method that sets `layer_groups` based on the result of [`split_model`](/torch_core.html#split_model). If `split_on` is a function, it calls that function and passes the result to [`split_model`](/torch_core.html#split_model) (see above for example).
# ### Saving and loading models
# Simply call [`Learner.save`](/basic_train.html#Learner.save) and [`Learner.load`](/basic_train.html#Learner.load) to save and load models. Only the parameters are saved, not the actual architecture (so you'll need to create your model in the same way before loading weights back in). Models are saved to the `path`/`model_dir` directory.
# + hide_input=true
show_doc(Learner.save)
# -
# If agument `name` is a pathlib object that's an absolute path, it'll override the default base directory (`learn.path`), otherwise the model will be saved in a file relative to `learn.path`.
learn.save("trained_model")
learn.save("trained_model", return_path=True)
# + hide_input=true
show_doc(Learner.load)
# -
# This method only works after `save` (don't confuse with `export`/[`load_learner`](/basic_train.html#load_learner) pair).
#
# If the `purge` argument is `True` (default) `load` internally calls `purge` with `clear_opt=False` to presever `learn.opt`.
learn = learn.load("trained_model")
# ### Deploying your model
# When you are ready to put your model in production, export the minimal state of your [`Learner`](/basic_train.html#Learner) with:
# + hide_input=true
show_doc(Learner.export)
# -
# If agument `fname` is a pathlib object that's an absolute path, it'll override the default base directory (`learn.path`), otherwise the model will be saved in a file relative to `learn.path`.
# Passing `destroy=True` will destroy the [`Learner`](/basic_train.html#Learner), freeing most of its memory consumption. For specifics see [`Learner.destroy`](/basic_train.html#Learner.destroy).
#
# This method only works with the [`Learner`](/basic_train.html#Learner) whose [`data`](/vision.data.html#vision.data) was created through the [data block API](/data_block.html).
#
# Otherwise, you will have to create a [`Learner`](/basic_train.html#Learner) yourself at inference and load the model with [`Learner.load`](/basic_train.html#Learner.load).
learn.export()
learn.export('trained_model.pkl')
path = learn.path
path
# + hide_input=true
show_doc(load_learner)
# -
# This function only works after `export` (don't confuse with `save`/`load` pair).
#
# The `db_kwargs` will be passed to the call to `databunch` so you can specify a `bs` for the test set, or `num_workers`.
learn = load_learner(path)
learn = load_learner(path, 'trained_model.pkl')
# WARNING: If you used any customized classes when creating your learner, you must first define these classes first before executing [`load_learner`](/basic_train.html#load_learner).
#
# You can find more information and multiple examples in [this tutorial](/tutorial.inference.html).
# ### Freeing memory
#
# If you want to be able to do more without needing to restart your notebook, the following methods are designed to free memory when it's no longer needed.
#
# Refer to [this tutorial](/tutorial.resources.html) to learn how and when to use these methods.
# + hide_input=true
show_doc(Learner.purge)
# -
# If `learn.path` is read-only, you can set `model_dir` attribute in Learner to a full `libpath` path that is writable (by setting `learn.model_dir` or passing `model_dir` argument in the [`Learner`](/basic_train.html#Learner) constructor).
# + hide_input=true
show_doc(Learner.destroy)
# -
# If you need to free the memory consumed by the [`Learner`](/basic_train.html#Learner) object, call this method.
#
# It can also be automatically invoked through [`Learner.export`](/basic_train.html#Learner.export) via its `destroy=True` argument.
# ### Other methods
# + hide_input=true
show_doc(Learner.init)
# -
# Initializes all weights (except batchnorm) using function `init`, which will often be from PyTorch's [`nn.init`](https://pytorch.org/docs/stable/nn.html#torch-nn-init) module.
# + hide_input=true
show_doc(Learner.mixup)
# -
# Uses [`MixUpCallback`](/callbacks.mixup.html#MixUpCallback).
# + hide_input=true
show_doc(Learner.backward)
# + hide_input=true
show_doc(Learner.create_opt)
# -
# You generally won't need to call this yourself - it's used to create the [`optim`](https://pytorch.org/docs/stable/optim.html#module-torch.optim) optimizer before fitting the model.
# + hide_input=true
show_doc(Learner.dl)
# -
learn.dl()
learn.dl(DatasetType.Train)
# + hide_input=true
show_doc(Recorder, title_level=2)
# -
# A [`Learner`](/basic_train.html#Learner) creates a [`Recorder`](/basic_train.html#Recorder) object automatically - you do not need to explicitly pass it to `callback_fns` - because other callbacks rely on it being available. It stores the smoothed loss, hyperparameter values, and metrics for each batch, and provides plotting methods for each. Note that [`Learner`](/basic_train.html#Learner) automatically sets an attribute with the snake-cased name of each callback, so you can access this through `Learner.recorder`, as shown below.
# + [markdown] hide_input=true
# ### Plotting methods
# + hide_input=true
show_doc(Recorder.plot)
# -
# This is mainly used with the learning rate finder, since it shows a scatterplot of loss vs learning rate.
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
learn = cnn_learner(data, models.resnet18, metrics=accuracy)
learn.lr_find()
learn.recorder.plot()
# + hide_input=true
show_doc(Recorder.plot_losses)
# -
# Note that validation losses are only calculated once per epoch, whereas training losses are calculated after every batch.
learn.fit_one_cycle(5)
learn.recorder.plot_losses()
# + hide_input=true
show_doc(Recorder.plot_lr)
# -
learn.recorder.plot_lr()
learn.recorder.plot_lr(show_moms=True)
# + hide_input=true
show_doc(Recorder.plot_metrics)
# -
# Note that metrics are only collected at the end of each epoch, so you'll need to train at least two epochs to have anything to show here.
learn.recorder.plot_metrics()
# ### Callback methods
# You don't call these yourself - they're called by fastai's [`Callback`](/callback.html#Callback) system automatically to enable the class's functionality. Refer to [`Callback`](/callback.html#Callback) for more details.
# + hide_input=true
show_doc(Recorder.on_backward_begin)
# + hide_input=true
show_doc(Recorder.on_batch_begin)
# + hide_input=true
show_doc(Recorder.on_epoch_end)
# + hide_input=true
show_doc(Recorder.on_train_begin)
# -
# ### Inner functions
# The following functions are used along the way by the [`Recorder`](/basic_train.html#Recorder) or can be called by other callbacks.
# + hide_input=true
show_doc(Recorder.add_metric_names)
# + hide_input=true
show_doc(Recorder.format_stats)
# -
# ## Module functions
# Generally you'll want to use a [`Learner`](/basic_train.html#Learner) to train your model, since they provide a lot of functionality and make things easier. However, for ultimate flexibility, you can call the same underlying functions that [`Learner`](/basic_train.html#Learner) calls behind the scenes:
# + hide_input=true
show_doc(fit)
# -
# Note that you have to create the [`Optimizer`](https://pytorch.org/docs/stable/optim.html#torch.optim.Optimizer) yourself if you call this function, whereas [`Learn.fit`](/basic_train.html#fit) creates it for you automatically.
# + hide_input=true
show_doc(train_epoch)
# -
# You won't generally need to call this yourself - it's what [`fit`](/basic_train.html#fit) calls for each epoch.
# + hide_input=true
show_doc(validate)
# -
# This is what [`fit`](/basic_train.html#fit) calls after each epoch. You can call it if you want to run inference on a [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) manually.
# + hide_input=true
show_doc(get_preds)
# + hide_input=true
show_doc(loss_batch)
# -
# You won't generally need to call this yourself - it's what [`fit`](/basic_train.html#fit) and [`validate`](/basic_train.html#validate) call for each batch. It only does a backward pass if you set `opt`.
# ## Other classes
# + hide_input=true
show_doc(LearnerCallback, title_level=3)
# + hide_input=true
show_doc(RecordOnCPU, title_level=3)
# -
# ## Undocumented Methods - Methods moved below this line will intentionally be hidden
# + hide_input=true
show_doc(Learner.tta_only)
# + hide_input=true
show_doc(Learner.TTA)
# + hide_input=true
show_doc(RecordOnCPU.on_batch_begin)
# -
# ## New Methods - Please document or move to the undocumented section
|
docs_src/basic_train.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Glue Tests
#
from myst_nb import glue
glue("key_text1", "text1")
glue("key_float", 3.14159)
glue("key_undisplayed", "undisplayed", display=False)
import pandas as pd
df = pd.DataFrame({"header": [1, 2, 3]})
glue("key_df", df)
import matplotlib.pyplot as plt
plt.plot([1, 2, 3])
glue("key_plt", plt.gcf(), display=False)
# ## Referencing the figs
#
# {glue:any}`key_text1`, {glue:}`key_plt`
#
# ```{glue:any} key_df
# ```
#
# and {glue:text}`key_text1` inline...
#
# and formatted {glue:text}`key_float:.2f`
#
# ```{glue:} key_plt
# ```
#
# and {glue:text}`key_undisplayed` inline...
#
#
# ```{glue:figure} key_plt
# :name: abc
#
# A caption....
# ```## A test title {glue:any}`key_text1`
#
#
# ## Math
import sympy as sym
f = sym.Function('f')
y = sym.Function('y')
n = sym.symbols(r'\alpha')
f = y(n)-2*y(n-1/sym.pi)-5*y(n-2)
glue("sym_eq", sym.rsolve(f,y(n),[1,4]))
# ```{glue:math} sym_eq
# :label: eq-sym
# ```
|
tests/notebooks/with_glue.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# 
# <br>
# <center> <NAME> | NYU Stern School of Business | Spring 2016 </center>
# ______________________
#
# ## To get started:
# ** Click 'Cell ➤ Run All' above. *When the program runs, it will skip down the page. After it stops skipping, scroll back up to the top to continue the lesson. Just a weird quirk of Jupyter.***
# 
# -
from IPython.display import display, HTML, clear_output
HTML('''<script> code_show=true; function code_toggle() {if (code_show){$('div.input').hide();}
else {$('div.input').show();}code_show = !code_show} $( document ).ready(code_toggle);
</script> <form action="javascript:code_toggle()"><input type="submit" value="Hide Raw Code"></form>''')
from ipywidgets import interact, interactive, fixed, widgets
import pandas as pd
import sqlite3
import re
# +
# just testing out the youtube player capabilites of Jupyter
#from IPython.display import YouTubeVideo
#YouTubeVideo("a1Y73sPHKxw", width=700, height=500)
# +
# if this .sqlite db doesn't already exists, this will create it
# if the .sqlite db *does* already exist, this establishes the desired connection
con = sqlite3.connect("sql_sample_db_new.sqlite")
book_table = pd.read_csv('https://raw.githubusercontent.com/DaveBackus/Data_Bootcamp/master/Code/SQL/book_table.csv')
auth_table = pd.read_csv('https://raw.githubusercontent.com/DaveBackus/Data_Bootcamp/master/Code/SQL/author_table.csv')
sales_table = pd.read_csv('https://raw.githubusercontent.com/DaveBackus/Data_Bootcamp/master/Code/SQL/sales_table.csv')
tech_cos = pd.read_csv('https://raw.githubusercontent.com/DaveBackus/Data_Bootcamp/master/Code/SQL/tech_cos.csv')
public_cos = pd.read_csv('https://raw.githubusercontent.com/DaveBackus/Data_Bootcamp/master/Code/SQL/public_cos.csv')
movie_table = pd.read_csv('https://raw.githubusercontent.com/DaveBackus/Data_Bootcamp/master/Code/SQL/movie_table.csv')
tables = [book_table,
auth_table,
sales_table,
tech_cos,
public_cos,
movie_table]
table_names = ['book_table',
'auth_table',
'sales_table',
'tech_cos',
'public_cos',
'movie_table']
# drop each table name if it already exists to avoid error if you rerun this bit of code
# then add it back (or add it for the first time, if the table didn't already exist)
for i in range(len(tables)):
table_name = table_names[i]
table = tables[i]
con.execute("DROP TABLE IF EXISTS {}".format(table_name))
pd.io.sql.to_sql(table, "{}".format(table_name), con, index=False)
# +
# Function to make it easy to run queries on this mini-database
def run(query):
try:
results = pd.read_sql("{}".format(query), con).fillna(' ')
return results
except:
pass
def run_q(query, button):
def on_button_clicked(b):
clear_output()
new_value = query.value.replace('\n', ' ')
if new_value != '':
df = run(new_value)
try:
output = HTML(df.to_html(index=False))
display(output)
#except AttributeError:
except:
print('''SQL error! Check your query:
1. Text values are in quotation marks and capitalized correctly
2. Items in the SELECT clause are comma-separated
3. No dangling comma in the SELECT clause right before the FROM clause
4. If you are joining tables that have columns with the same name, use table_name.column_name format
5. Try "PRAGMA TABLE_INFO(table_name) to double-check the column names in the table
6. Correct order of clauses:
SELECT
FROM
JOIN...ON
WHERE
GROUP BY
ORDER BY
LIMIT
''')
button.on_click(on_button_clicked)
on_button_clicked(None)
def cheat(answer):
def f(Reveal):
if Reveal == False:
clear_output()
else:
print(answer)
interact(f, Reveal=False)
clear_output()
# + [markdown] slideshow={"slide_type": "slide"}
# ____________
# ____________
# ____________
# <a id='table_of_contents'></a>
# # Table of Contents
#
# [Course Details](#course_details)
#
# [Introduction to SQL](#introduction)
#
# [Structure and Formatting Basics](#formatting)
#
# [Determine a table's strucure.............................`PRAGMA TABLE_INFO()`](#pragma_table)
#
# [Query building blocks.......................................`SELECT` & `FROM`](#select_from)
#
# [Filter your data..................................................`WHERE`](#where)
#
# [Wildcards and vague search.............................`LIKE` and `%`](#where_like)
#
# [Sort your data...................................................`ORDER BY`](#order_by)
#
# [Limit the number of rows you see.....................`LIMIT`](#limit)
#
# [Combining tables..............................................`JOIN`](#join_tables) <br>
# • [Combining 3+ tables........................................Multiple `JOIN`'s](#multi_join) <br>
# • [Different `JOIN` Types.......................................Overview](#join_types)<br>
# • [Simple Join......................................................`INNER JOIN` aka `JOIN`](#inner_joins)<br>
# • [One-Sided Join................................................`LEFT JOIN`](#left_joins)<br>
# • [Full Join............................................................`OUTER JOIN`](#outer_joins)<br>
# • [Practice combining tables...............................`JOIN` Drills](#join_drills)
#
# [Column & Table Aliases.....................................`AS`](#as)
#
# [Add, subtract, multiply, & divide data................Operators](#operators)
#
# [Apply functions to columns...............................Functions](#functions)
#
# [Group data by categories..................................`GROUP BY`](#group_by)
#
# [Filter out certain groups.....................................`HAVING`](#having)
#
# [Conditional values..............................................`IF` & `CASE WHEN`](#case_when)
#
# [SQL-ception: Queries within queries..................Nesting](#nesting)
#
# [Run multiple queries at once..............................`UNION` & `UNION ALL`](#union)
#
# [Add a summaryt/total row..................................`ROLLUP`](#rollup)
#
# [Summary](#wrapping_up)
#
# [Full table of RDBMS dialect differences](#dialect_differences)
#
# ### Additional Resources:
# - [Syllabus](https://www.dropbox.com/s/b4orgekbbom40x6/SQL_Bootcamp_Syllabus.pdf?dl=0)
# - [Cheatsheet](https://www.dropbox.com/s/oo0uhi7xm2sfy5e/SQLBOOTCAMPCHEATSHEET.pdf?dl=0)
# - [Google Group](https://groups.google.com/forum/#!forum/nyu_data_bootcamp)
# + [markdown] slideshow={"slide_type": "slide"}
# _______
# _______
# _______
# <center> <a id='introduction'></a> [Table of Contents](#table_of_contents) | [Next](#course_details)
# </center>
#
# 
#
#
# ### SQL, "sequel", "ESS CUE ELL"
# SQL stands for "Structured Query Language", but no one calls it that. You can pronouce it as either "S-Q-L" or "sequel". Some people feel strongly in favor of a particular pronunciation. I don't. I'll say "sequel" in class, but I'll never correct you for saying S-Q-L.
#
# SQL is the database language of choice for most businesses; you use it to communicate with databases. "Communication" can take the form of creating, reading, updating, and deleting data. This course only covers reading data. That's all most MBAs do with SQL.
#
# ____________
# ### Relational Databases
#
# Companies use relational databases because they can store and easily recall A LOT of data. Excel can't handle more than a million rows. If you're Amazon and you need to record every click, Excel is useless. Relational databases are much more efficient.
#
# What do we mean by "efficient"? Every recorded "bit" takes up server space, which costs money. It also slows everything down. So an efficient database should allow you to record and recall a lot of information using the minimal number of bits.
#
# Imagine you want to store the names of four books and some information about their authors. Think of the character count as a proxy for how many bits of storage your table takes up:
# 
#
# Now imagine you want to add some more books by each of those authors. Some of the information gets redundant. Imagine if you had to do this for millions of different books:
# 
#
# This is where relational databases can help. With a relational database, you'd create two separate tables that *relate* to each other. You still storing the same information, but you're doing it by using fewer characters. You've eliminated the need to repeat yourself, so you've made a much more efficient database.
# 
# <br>
# _______
# ### SQL Dialects
#
# There are different softwares that can manage relational databases. SQL varies a little from software to software, just like English varies a little between England and the U.S. We'll address these instances whenever possible.
#
# Each software is called a Relational Database Management System, or RDBMS. These are some of the most popular that you might encounter at work:
#
# <font color='#1f5fd6'> Microsoft SQL Server | <font color='#1f5fd6'> MySQL | <font color='#1f5fd6'> Oracle | <font color='#1f5fd6'> SQLite </font>|
# :------------------: | :---: | :----: | :----: |
# Proprietary, more common at older companies | Open source, frequently used by startups and tech companies | Proprietary, more common at older companies | Frequently used for mobile apps (and this class!)
# <br>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ________
# _______
# _______
# <a id='course_details'></a>
# <center> [Previous](#introduction) | [Table of Contents](#table_of_contents) | [Next](#formatting)
# </center>
#
# 
#
# The goal is to start simple and practice often. By the end of this class, you should feel extremely comfortable writing moderately complicated SQL code, which will save you countless hours trying to figure out SQL on the job or waiting for someone else with SQL knowledge to pull data for you. Using this interactive program, we'll explore a small sample database by learning new SQL concepts one at a time. Concepts will build on each other.
#
# ### Quick Exercises
# Sometimes you'll be asked to edit or delete parts of a provided query. Rerun the query with each step, taking care to understand what changed with the output each time. Note that none of the changes that you make to these queries will be saved when you close this program.
#
#
# **Try it by changing something in the cell below and hitting "Run"!**
# -
test_ex = widgets.Textarea(value=
'''You can change text in these boxes to edit and re-run queries!''',
width = '50em', height = '7em')
display(test_ex)
text_ex_button = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(text_ex_button)
def on_button_clicked2(b):
clear_output()
print('''Here's the output from the cell above:
''', test_ex.value)
text_ex_button.on_click(on_button_clicked2)
# ### Challenges
# After we've learned a new concept and you've practiced with some quick exercises, you'll be challenged to write your own query. Read each challenge carefully, and keep re-running it until you get the results you are looking for.
#
# **Need to cheat a little?** Check the "Reveal" box to see the answer to a challenge.
# <br>
# <img align="left" src="http://i.imgur.com/FhCJTqa.png">
# + slideshow={"slide_type": "subslide"}
chall ='''When you click a checkbox, you reveal the answer to a challenge!
Uncheck it to hide the answer again.'''
cheat(chall)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Using this program
# The content you're currently reading is written in Python, Markdown and HTML and runs in a Jupyter Notebook. No need to know what any of that means, I only told you in case you were terribly curious.
#
# **You will not be using this interface at work** - the point of this class is to teach you SQL the language, which can be typed into a variety of different software programs. You'll be able to learn the quirks of a different software program pretty easily as long as you know SQL.
#
# Still, there are some things you should know about this program to help you with the class:
# * If you accidentally double-click on a block of text, and suddenly it looks like code, hit **`SHIFT-RETURN`** or **'Cell ➤ Run'**.
# * If you try to run a query and the output doesn't refresh, select **'Cell ➤ Run All'** to reboot the program.
# * If you accidentally delete a cell, click **'Edit ➤ Undo Delete Cell'**
# * Nothing that you write in the challenges and exercises will save after you close this program.
# * If you want to save something that you've written, follow the steps below:
# <br>
# <img align="left" src="http://i.imgur.com/qkh6TiN.png">
# + active=""
# A grey "cell" like this will appear when you follow Step 1 (click the + sign in the toolbar) and Step 2 (change the cell to Raw NBConvert). Write your notes in the cell; the program will automatically save it.
# + [markdown] slideshow={"slide_type": "slide"}
# _______
# _______
# ________
# _______
# ________
# ________
# ________
# 
#
# _______
# ________
# __________
# ________
# ________
# ________
# ________
#
# <a id='formatting'></a>
# <center>
# [Previous](#course_details) | [Table of Contents](#table_of_contents) | [Next](#pragma_table)
# </center>
# ### Structure and Formatting Query Basics:
# Below is an example of a **query**, SQL code that requests data from a database. Try to make a habit of writing queries by following these formatting conventions. Queries can get very long and complicated, and formatting makes them easier to read.
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# ______
# ______
# ______
#
# <a id='pragma_table'></a>
#
# <center>
# [Previous](#formatting) | [Table of Contents](#table_of_contents) | [Next](#select_from)
# </center>
#
# 
#
#
# <font color='#1f5fd6'>Microsoft SQL Server | <font color='#1f5fd6'>MySQL | <font color='#1f5fd6'>Oracle | <font color='#1f5fd6'>SQLite </font>
# :------------------: | :---: | :----: | :----:
# `SP_Help some_table` | `DESCRIBE some_table` | `DESCRIBE some_table` | `PRAGMA TABLE_INFO(some_table)`
#
# ### SQLite version that we'll be using for this class:
# > **`PRAGMA TABLE_INFO(some_table)`** ➞ result-set lists the column names and data types within the table
#
# We're using SQLite, so we're going to be using the **`PRAGMA TABLE_INFO()`** option. Put the name of a table in the parentheses, and the output tells you the names and data types in each column in the table.
#
# So far, we've learned about 2 tables in our relational database, which we'll call `book_table` and `auth_table`. We're also going to use a `sales_table`, which we'll take a look at later on. Combined, these three tables will make up the "database" of a very tiny, very limited, very imaginary bookstore.
#
# We'll start by reviewing the `book_table` and `auth_table`:
# 
#
# Now, we'll use `PRAGMA TABLE_INFO()` to read the table structure of the `book_table`. In plain English, the query below says "*show me the names of the columns in the `book_table`, and what type of data (text, numbers?) is in each column*."
# + slideshow={"slide_type": "slide"}
pragma = widgets.Textarea(
value='''PRAGMA TABLE_INFO(book_table)''',
width = '50em', height = '3em')
display(pragma)
# + slideshow={"slide_type": "-"}
prag_button = widgets.Button(description='Run',width='10em', height='2.5em', color='white', background_color='black', border_color='black')
display(prag_button)
run_q(pragma, prag_button)
# -
# - **name** tells us the names of each column in the table. So now we know that the **`book_table`** has columns headed **`Book`**, **`COGs`**, and **`Author`**
# - **type** tells us what type of data is in each column. So now we know that the **`Book`** column has TEXT data, and that **`COGs`** contains REAL numbers - numbers that can have a fractional value.
# - **All other columns** you can ignore. Seriously.
#
#
# ### Quick Exercise:
# Change the query above to look at the **`auth_table`** instead. Why is the author's **`birth_year`** data type not REAL like we saw with **`COGs`**?
# _______
#
# ### Challenge:
# Rewrite the query above to take a look at the **`sales_table`** structure. Judging from what your query returns, can you guess what you'll probably see once you actually look all the data in the **`sales_table`**?
pragma_sales = widgets.Textarea(
value='',
width = '50em',
height = '4em'
)
display(pragma_sales)
prag_sales_button = widgets.Button(description='Run',width='10em', height='2.5em', color='white', background_color='black', border_color='black')
display(prag_sales_button)
run_q(pragma_sales, prag_sales_button)
prag_sales_answer = 'PRAGMA TABLE_INFO(sales_table)'
cheat(prag_sales_answer)
# _________
# _________
# ________
# <a id='select_from'></a>
# <center>
# [Previous](#pragma_table) | [Table of Contents](#table_of_contents) | [Next](#where)
# </center>
#
# 
# <!--<center>
# [Jump to: Selecting specific columns](#select_col) | [Jump to: Selecting distinct values](#select_distinct)
# </center>-->
#
# > **`SELECT` <br>
# `*` **➞ an asterisk means "all columns"** <br>
# `FROM` <br>
# `table_name`**
#
#
# To see the actual data in a table, we'll use **[`SELECT`](http://www.w3schools.com/sql/sql_select.asp)** and **`FROM`** clauses. In the `SELECT` clause, you tell SQL which columns you want to see. In the `FROM` clause, you tell SQL the table where those columns are located. An **asterisk returns all columns from a particular table.**
#
#
# In plain English, the query below says: "*Show me all columns and their data from the `book_table`*"
select = widgets.Textarea(value=
'''SELECT
*
FROM
book_table''',
width = '50em', height = '7em')
display(select)
select_button = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(select_button)
run_q(select, select_button)
# ### Quick Exercise:
# Change the query above to show us all columns and their data from the **`auth_table`** instead of the **`book_table`**
# _____________
#
# ### Challenge:
# Write a query to view all columns and their data from the **`sales_table`**
select_c = widgets.Textarea(value='', width = '50em')
display(select_c)
select_c_button = widgets.Button(
description='Run',
width='10em',
height='2.5em',
color='white',
background_color='black',
border_color='black')
display(select_c_button)
run_q(select_c, select_c_button)
select_answer ='''SELECT
*
FROM
sales_table'''
cheat(select_answer)
# <img align="left" src="http://i.imgur.com/p6d18FV.png"> <br><br>
# **Use asterisks sparingly**. Usually, you'll select specific columns from a table rather than all columns. Using an asterisk to select all columns is okay when the table is small or when you tightly constrain your selection of rows. Otherwise, select specific columns and use WHERE and LIMIT (taught below) to go easy on your servers.
# <a id='select_col'></a>
# _____________
#
# ## `SELECT` specific columns:
# > **`SELECT` <br>
# `column_a,`** ➞ separate multiple columns with commas <br>
# ** `column_b`** ➞ optional, but conventional, to also use a return <br>
# **`FROM` <br>
# `table_name`** <br>
#
# Instead of using an asterisk for "all columns", you can specify a particular column or columns. In plain English: "*Show me the data in the `book` and `author` columns from the `book_table`"*
select_col = widgets.Textarea(value=
'''SELECT
book,
author
FROM
book_table''',
width = '50em', height = '8em')
display(select_col)
select_col_button = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(select_col_button)
run_q(select_col, select_col_button)
# ________
# ### Challenge:
# Write a query to show the **`first_name`** and ** `last_name` ** columns from the **`auth_table`**
select_cols_chall = widgets.Textarea(value=
'',
width = '50em', height = '8em')
display(select_cols_chall)
select_cols_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(select_cols_chall_b)
run_q(select_cols_chall, select_cols_chall_b)
select_cols_chall_cheat ='''SELECT
first_name,
last_name
FROM
auth_table'''
cheat(select_cols_chall_cheat)
# ________
# ### Challenge:
# Write a query to select only the **`book`** column from the **`sales_table`**
select_cols_chall2 = widgets.Textarea(value='', width = '50em', height = '8em')
display(select_cols_chall2)
select_cols_chall_b2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(select_cols_chall_b2)
run_q(select_cols_chall2, select_cols_chall_b2)
select_cols_chall_cheat2 ='''SELECT
book
FROM
sales_table'''
cheat(select_cols_chall_cheat2)
# <a id='select_distinct'></a>
# _________
#
# ## `SELECT DISTINCT`:
# > **`SELECT` <br>
# `DISTINCT column_a`** ➞ *returns only unique values* <br>
# **`FROM` <br>
# `table_name`** <br>
#
# Use **`DISTINCT`** to return unique values from a column, so if there are any repeats in a column, your **output will include each value just once**. The query below displays each book in the `sales_table` just once, even though we know each shows up multiple times in the table.
distinct_q = widgets.Textarea(value=
'''SELECT
DISTINCT book
FROM
sales_table''',
width = '50em', height = '7em')
display(distinct_q)
distinct_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(distinct_b)
run_q(distinct_q, distinct_b)
# ______
# ### Challenge:
# Write a query to return each author from the **`book_table`** without any names repeating.
distinct_q_chall = widgets.Textarea(value='', width = '50em', height = '7em')
display(distinct_q_chall)
distinct_q_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(distinct_q_chall_b)
run_q(distinct_q_chall, distinct_q_chall_b)
REP ='''SELECT
DISTINCT author
FROM
book_table'''
cheat(REP)
# _____
# ____
# ____
# <a id='where'></a>
# <center>
# [Previous](#select_from) | [Table of Contents](#table_of_contents) | [Next](#where_like)
# </center>
# 
#
# <!--
# [Jump to: WHERE & Text Values](#where_text) [Jump to: Where & Numbers](#table_of_contents) [Jump to: WHERE & Multiple Requirements](#where_and)
# -->
#
# >**`SELECT` <br>
# `column_a` <br>
# `FROM` <br>
# `table_name` <br>
# `WHERE` <br>
# `column_a = x`** ➞ result-set will only include rows where value of column_a is x
#
# [**`WHERE`**](http://www.w3schools.com/sql/sql_where.asp) lets you filter results so you only see rows that specifically match your criteria. Below there are few more options for the **`WHERE`** clause:
#
# Options for WHERE | Description
# :------- | :-------------
# `col = 'some_text'` | Put text in quotations. Capitalization is important!
# `col != x` | Return rows where col's values DO NOT equal x
# `col < x` | Return rows where col's value is less than x
# `col <= x` | Return rows where col's value is less OR EQUAL TO than x
# `col IN (x, y)` | Values can equal EITHER x OR y
# `col NOT IN (x, y)` | Value are NEITHER x NOR y
# `col BETWEEN x AND y` | Values are between x and y
# `col = x AND another_col = y` | Returns rows when col's values are x AND another_col's values are y
# `col = x OR another_col = y` | Returns rows when col's values are x OR another_col's values are y
#
# __________
#
# ## `WHERE` & text values
#
# Below, we use **`WHERE`** to tell SQL to only show us rows in the **`book_table`** when Hemingway is the author. In plain English, we're saying "*Show me information about books that are written by Hemingway in the `book_table`*"
where_q = widgets.Textarea(value=
'''SELECT
*
FROM
book_table
WHERE
author = 'Hemingway' ''',
width = '50em', height = '10em')
display(where_q)
where_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(where_b)
run_q(where_q, where_b)
# ### Quick Exercises:
# 1. Above, change the name from **`'Hemingway'`** to **`'Shakespeare'`**, rerun
# 2. Delete the quotation marks around the word `Shakespeare`, rerun. Why the error?
# 2. Put **double** quotation marks, rerun
# 3. Change **`"Shakespeare"`** to **`"shakespeare"`**, rerun
# 4. Change **`"shakespeare"`** to **`"Twain"`**, rerun
# 5. Change **`"Twain"`** to **`'Hemingway'`**, rerun to get back to where we started
# 6. Change = in the **`WHERE`** clause to !=, rerun
# _______
# ### Challenge:
#
# Write a query to return all columns of the **`auth_table`**, but only rows where the author's country is England.
where_q_chall = widgets.Textarea(value='',
width = '50em', height = '10em')
display(where_q_chall)
where_b_chall = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(where_b_chall)
run_q(where_q_chall, where_b_chall)
where_q_chall_cheat ='''SELECT
*
FROM
auth_table
WHERE
country = 'England' '''
cheat(where_q_chall_cheat)
# We use **`IN (value_1, value_2)`** to return rows that can match more than one value. In plain English, the query below says, "*Show me all columns from the book table when the author is EITHER Hemingway OR Austen*"
in_q = widgets.Textarea(value=
'''SELECT
*
FROM
book_table
WHERE
author IN ('Hemingway', 'Austen')''',
width = '50em', height = '10em')
display(in_q)
in_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(in_b)
run_q(in_q, in_b)
# ### Quick Exercise:
# 1. Add **`'Faulkner'`** to the list, rerun.
# 2. Replace **`IN`** with **`NOT IN`**, rerun.
# 3. Delete the whole last line and replace it so that the query returns all books except for <u>Emma</u> and <u>Macbeth</u>.
#
# <a id='where_numbers'></a>
# ________
# ## `WHERE` & number values
# The **`WHERE`** clause is useful with numbers as well. We can start throwing in comparisons like less than (<) and greater than (>):
greater_q = widgets.Textarea(value=
'''SELECT
*
FROM
sales_table
WHERE
revenue > 18''',
width = '50em', height = '10em')
display(greater_q)
greater_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(greater_b)
run_q(greater_q, greater_b)
# ### Quick Exercises:
# 1. Replace > with <, rerun
# 2. Add an = directly after the <, rerun
# 2. Change the line to **`revenue BETWEEN 10 AND 12`**, rerun
#
# _____
#
# ### Challenge:
# Write a query that returns all columns from the `auth_table` for authors with a `birth_year` before 1800:
born_chall = widgets.Textarea(value='',width = '50em', height = '10em')
display(born_chall)
born_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(born_chall_b)
run_q(born_chall, born_chall_b)
born_chall_c ='''SELECT
*
FROM
auth_table
WHERE
birth_year < 1800
'''
cheat(born_chall_c)
# <a id='where_and'></a>
# _____________
#
# # `WHERE` with `AND`/`OR`
#
# So far, we've only filtered by a specific column (like the revenue column , country, or author columns). Sometimes you'll want to filter by multiple columns. This is where **`AND`** and **`OR`** come in handy.
and_q = widgets.Textarea(value=
'''SELECT
*
FROM
book_table
WHERE
author = 'Hemingway'
AND cogs > 11''',
width = '50em', height = '12em')
display(and_q)
and_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(and_b)
run_q(and_q, and_b)
# ### Quick Exercises:
# 1. Delete **`AND cogs > 11`** and rerun the query. Then replace it and run it again.
# 2. Change the word **`AND`** to **`OR`**, rerun. What's going on?
# _________
# ### Challenge:
#
# Write a query to pull the **`last_name`**, **`country`**, and **`birth_year`** of authors who were from England AND born after 1650
and_chall = widgets.Textarea(value=
'',
width = '50em', height = '14em')
display(and_chall)
and_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(and_chall_b)
run_q(and_chall, and_chall_b)
and_chall_c ='''SELECT
last_Name,
country,
birth_Year
FROM
auth_table
WHERE
country = 'England'
AND birth_Year > 1650 '''
cheat(and_chall_c)
# ________
# ### Challenge:
# Write a query to see all columns from the **`sales_table`** where the book name is <u>Macbeth</u> OR **`revenue`** was greater than $17.
or_chall = widgets.Textarea(value=
'',
width = '50em', height = '12em')
display(or_chall)
or_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(or_chall_b)
run_q(or_chall, or_chall_b)
or_chall_cheat ='''SELECT
*
FROM
sales_table
WHERE
book = 'Macbeth'
OR revenue > 17 '''
cheat(or_chall_cheat)
# __________
# <a id='where_like'></a>
# <center>
# [Previous](#where) | [Table of Contents](#table_of_contents) | [Next](#order_by)
# </center>
# 
#
# >**`SELECT` <br>
# `column_a` <br>
# `FROM` <br>
# `table_name` <br>
# `WHERE` <br>
# `column_a LIKE 's%Me_t%xT'`** ➞ correct capitalization isn't necessary with `LIKE`, and `%` stands in for any missing character
#
# [**`LIKE`**](http://www.w3schools.com/sql/sql_like.asp) lets you search for a value even if you capitalize it incorrectly. It also allows you to work with percentage signs that act as [wildcards](http://www.w3schools.com/sql/sql_wildcards.asp), which stand in for an unlimited number of missing characters (helpful if you don't know how to spell something). Take a look at the query below. **Recall that earlier when we wrote `author = 'shakespeare'`, we got no results.**
like_q = widgets.Textarea(value=
'''SELECT
*
FROM
book_table
WHERE
author LIKE 'hemingway' ''',
width = '50em', height = '10em')
display(like_q)
like_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(like_b)
run_q(like_q, like_b)
# ### Quick Exercises:
# 1. Replace **`'hemingway'`** with **`'hemingWAY'`**, rerun
# 2. Replace **`LIKE`** with =, rerun
# 3. Replace = with **`LIKE`** again, but change '**`hemingWAY`**' to '**`Hemmingway`**', rerun
# 4. Replace '**`Hemmingway`**' with **`'Hem'`**, rerun
#
# ____________
#
# ## Using % as a "wildcard"
# With exercises #3 and #4, you saw that **`LIKE`** alone has a limitation - it only lets you mess with capitalization. You need **wildcards** to do more with **`LIKE`**. Let's say you can't remember if Hemingway is spelled with 1 "m" or 2. Use a percentage sign (%) to get the value you're looking for:
like_q2 = widgets.Textarea(value=
'''SELECT
*
FROM
book_table
WHERE
author LIKE 'He%ingway' ''',
width = '50em', height = '11em')
display(like_q2)
like_b2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(like_b2)
run_q(like_q2, like_b2)
# ### Quick Exercises:
# 1. Change **`'He%ingway'`** to **`'Hemm%ingway'`**. Why doesn't this work?
# 2. Change **`'Hemm%ingway'`** to **`'Hem%'`**, rerun
# 3. Change **`'Hem%'`** to **`'%us%'`**, rerun
# 4. Change **`LIKE`** to =, rerun (see how wildcards only work with **`LIKE`**?)
# __________
# ### Challenge:
#
# Write a query to pull the **`book`** and **`author` ** columns from the **`book_table`**. Pretend you can't remember the full name of the book you're looking for. You just know it starts with the word "Pride".
like_chall = widgets.Textarea(value=
'',
width = '50em', height = '12em')
display(like_chall)
like_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(like_chall_b)
run_q(like_chall, like_chall_b)
like_chall_c ='''SELECT
book,
author
FROM
book_table
WHERE
book LIKE 'Pride %' '''
cheat(like_chall_c)
# <img align="left" src="http://i.imgur.com/p6d18FV.png"> <br><br>
# **Use `LIKE` sparingly:** It's a great tool, but it really puts the strain on your database's servers. Use it only when a table is pretty small or when you've limited your result-set by using additional filters in the `WHERE` clause.
# __________________________
# __________
# __________________________
# <a id='order_by'></a>
# <center>
# [Previous](#where_like) | [Table of Contents](#table_of_contents) | [Next](#limit)
# </center>
# 
#
# > **`SELECT` <br>
# `column_a` <br>
# `FROM` <br>
# `table_name`** <br>
# `[WHERE clause, optional]` <br>
# **`ORDER BY`** ➞ sorts the result-set by column_a <br>
# **`column_a DESC`** ➞ `DESC` is optional, it sorts results in descending order
#
# Without an `ORDER BY` clause, the default result-set will be sorted by however it appears in the database (which is crap-shoot depending on the type of table). **Use [`ORDER BY`](http://www.w3schools.com/sql/sql_orderby.asp) to sort your result-set by a particular column**, and add **`DESC`** to sort in descending order (Z→A, 100→1).
order_q = widgets.Textarea(value=
'''SELECT
*
FROM
book_table
ORDER BY
book''',
width = '50em', height = '11em')
display(order_q)
order_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(order_b)
run_q(order_q, order_b)
# ### Quick Exercises:
# 1. Change the query so it sorts by **`author`** instead
# 2. Add **`DESC`** and rerun
# 3. Delete **`author DESC`**, replace it with **`author, book`**, rerun
# 3. Add **`DESC`** so it reads **`author, book DESC`**, rerun
# 4. Change the line to **`author DESC, book`**
# ________
#
# ### Challenge:
#
# Write a query to see the **`book`** and **`revenue`** columns from the **`sales_table`**, and sort the results by **`revenue`** in descending order.
order_chall = widgets.Textarea(value=
'',
width = '50em', height = '12em')
display(order_chall)
order_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(order_chall_b)
run_q(order_chall, order_chall_b)
order_chall_c ='''SELECT
book,
revenue
FROM
sales_table
ORDER BY
revenue DESC'''
cheat(order_chall_c)
# ______
# ### Challenge:
# Write a query to view all columns from the **`book_table`**, but only where the author's name is something like "pear" or **`COGs`** are over $12. Sort your results by **`COGs`** with the cheapest book first.
order_chall2 = widgets.Textarea(value=
'',
width = '50em', height = '16em')
display(order_chall2)
order_chall_b2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(order_chall_b2)
run_q(order_chall2, order_chall_b2)
order_chall_c2 ='''SELECT
*
FROM
book_table
WHERE
author LIKE '%pear%'
OR cogs > 12
ORDER BY
cogs'''
cheat(order_chall_c2)
# _____
# ____
# ____
# <a id='limit'></a>
# <center>
# [Previous](#order_by) | [Table of Contents](#table_of_contents) | [Next](#join_tables)
# </center>
#
# 
# <font color='#1f5fd6'>Microsoft SQL Server | <font color='#1f5fd6'>MySQL | <font color='#1f5fd6'>Oracle | <font color='#1f5fd6'>SQLite </font>
# :------------------: | :---: | :----: | :----:
# `SELECT TOP N column_name` | `LIMIT N` | `WHERE ROWNUM <= N` | `LIMIT N`
#
# >**`SELECT`** <br>
# **`column_a`** <br>
# **`FROM`** <br>
# **`table_name`** <br>
# `[WHERE clause]` <br>
# `[ORDER BY clause]` <br>
# **`LIMIT N`** ➞ *limits the result-set to N rows*
#
# [**`LIMIT`**](http://www.w3schools.com/sql/sql_top.asp) lets you set a maximum limit to the number of rows that your query returns. You've seen the query below before, but now we've added a `LIMIT` clause:
limit_q = widgets.Textarea(value=
'''SELECT
*
FROM
sales_table
LIMIT 5''',
width = '50em', height = '14em')
display(limit_q)
limit_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(limit_b)
run_q(limit_q, limit_b)
# ### Quick Exercises:
# 1. Change 5 to 10, rerun
# 2. Add an **`ORDER BY`** clause so that you see the top 10 transactions in terms of **`revenue`**, rerun
# 3. Add a **`WHERE`** clause so you only see transactions relating to <u>Emma</u>
#
# ### Challenge:
# Write a query to view the **`title`** and **`cogs`** of the two books with the lowest cogs in the **`book_table`**
limit_chall = widgets.Textarea(value=
'',
width = '50em', height = '12em')
display(limit_chall)
limit_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(limit_chall_b)
run_q(limit_chall, limit_chall_b)
limit_chall_c ='''SELECT
book,
cogs
FROM
book_table
ORDER BY
cogs
LIMIT 2'''
cheat(limit_chall_c)
# ### Challenge:
# Write a query to view the **`book`** and **`revenue`** columns from the **`sales_table`** and sort by **`book`** title first, then by **`revenue`** (ascending). Limit your results to 15 rows.
limit_chall2 = widgets.Textarea(value=
'',
width = '50em', height = '12em')
display(limit_chall2)
limit_chall2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(limit_chall2_b)
run_q(limit_chall2, limit_chall2_b)
limit_chall2_c ='''SELECT
book,
revenue
FROM
sales_table
ORDER BY
book, revenue
LIMIT 15'''
cheat(limit_chall2_c)
# _________
# _________
# __________
# <a id='join_tables'></a>
# <center>
# [Previous](#limit) | [Table of Contents](#table_of_contents) | [Next](#multi_join)
# </center>
# 
#
# > **`SELECT` <br>
# `table_x.column_a,`** ➞ read this as "column_a from table_x"<br>
# ** `table_y.column_b,`** ➞ "column_b from table_y"<br>
# **`FROM`<br>
# `table_x`<br>
# `JOIN table_y`** <br>
# **`ON table_x.key_column_x = table_y.key_column_y`** <br> ➞ table_x's key_column_x has corresponding values with table_y's key_column_y<br>
# `[WHERE clause]` <br>
# `[ORDER BY clause]` <br>
# `[LIMIT clause]` <br>
#
# The ability to [join](http://www.w3schools.com/sql/sql_join.asp) tables is the most fundamental and useful part about relational databases. Different tables have columns with corresponding values, and you can use these columns as "keys" to join the two tables.
#
# The format `table_x.key_column` can be read as "`key_column` from `table_x`"; it tells SQL the tables where a column is located. We didn't need this before because we were only using one table at a time, so SQL knew exactly which table we were talking about. When we deal with more than one table, we need to be more specific. So for example, **`book_table.book` means "the `book` column from the `book_table`"**, and `auth_table.last_name` means "the `last_name` column from the `auth_table`."
#
# Think back to our original discussion of splitting up our author and book data onto two separate tables:
# 
#
# You could think of the columns in these tables in terms of a Venn Diagram. Again, the format `table_x.key_column` is read as "`key_column` from `table_x`", so `book_table.author` means "the `author` column from the `book_table`":
# 
#
# **The `author` column from the `book_table` corresponds with the `last_name` column in the `auth_table`** - they both list the last names of the writers. Whenever you have two tables corresponding columns, you can "join" them by telling SQL to use these corresponding columns as keys. **`book_table.author` and `auth_table.last_name` are the key columns for our join**.
join_q = widgets.Textarea(value=
'''SELECT
*
FROM
book_table
JOIN auth_table
ON book_table.author = auth_table.last_name''',
width = '50em', height = '11em')
display(join_q)
join_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_b)
run_q(join_q, join_b)
# What just happened? SQL went through these steps:
#
# - First, SQL pulled up the two tables that we named in the `FROM` clause: **`FROM book_table JOIN auth_table`**.
# - Then it identified the "key" columns that we named with `ON`: **`ON book_table.author = auth_table.last_name`**.
# - Next, it matched up the values on the key columns:
# 
#
# - Whenever it found a match, it made a kind of copy of the row from the **`auth_table`** and pasted it to the **`book_table`**
# - Finally, it literally "joined" the two tables by returning their columns in a single table
# 
#
# ### Quick Exercises:
# The **`JOIN`** query we've been discussing has been reproduced in the box below for these exercises.
# 1. Change the query so you only see the **`book`**, **`first_name`** and the author's last name (you can do this with either **`author`** or **`last_name`**), and the **`birth_year`**, then rerun.
# 2. Add a **`WHERE`** clause so that you only see books by Hemingway and Austen, rerun.
# 3. Add an **`ORDER BY`** clause so that the author born first appears first, and so that their books appear in alphabetical order. Rerun.
#
join_q2 = widgets.Textarea(value=
'''SELECT
*
FROM
book_table
JOIN auth_table
ON book_table.author = auth_table.last_name''',
width = '50em', height = '18em')
display(join_q2)
join_b2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_b2)
run_q(join_q2, join_b2)
# ### Challenge:
# Our "database" has another Venn Diagram relationship: the **`book_table`** is related to the **`sales_table`**. Write a query to join these tables and view all their columns but limit your results to 20 rows. Use the Venn Diagram below as a guide:
# 
join_chall = widgets.Textarea(value=
'',
width = '50em', height = '15em')
display(join_chall)
join_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_chall_b)
run_q(join_chall, join_chall_b)
join_chall_c ='''SELECT
*
FROM
book_table
JOIN sales_table
ON book_table.book = sales_table.book
LIMIT 20'''
cheat(join_chall_c)
# ### Challenge Continued:
# Start by copying the query that you wrote in the previous challenge and pasting it in the box below.
# 1. Change the line **`book_table.book = sales_table.book`** to **`book = book`** and rerun. What's going wrong? Fix it and rerun.
# 2. Change the query so that you only see the book title listed once. If you get stuck, remember that **`table_x.column_a`** means "column_a from table_x".
join_chall2 = widgets.Textarea(value=
'',
width = '50em', height = '17em')
display(join_chall2)
join_chall2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_chall2_b)
run_q(join_chall2, join_chall2_b)
join_chall_c2 ='''SELECT
book_table.book, [NOTE: or you could use "sales_table.book" instead]
cogs,
author,
id,
revenue
FROM
book_table
JOIN sales_table
ON book_table.book = sales_table.book
LIMIT 20'''
cheat(join_chall_c2)
# _________
# __________
# ____________
# <a id='multi_join'></a>
# <center>
# [Previous](#join_tables) | [Table of Contents](#table_of_contents) | [Next](#join_types)
# </center>
# 
#
# > **`SELECT` <br>
# `table_x.column_a,` <br>
# `table_y.column_b` <br>
# `table_z.column_c`<br>
# `FROM`<br>
# `table_x`<br>
# `JOIN table_y ON table_x.key_column = table_y.key_column`<br>
# `JOIN table_z ON table_x.other_key_column = table_z.other_key_column`<br>**
# `[WHERE clause]` <br>
# `[ORDER BY clause]` <br>
# `[LIMIT clause]` <br>
#
# As long as they are directly related or related by the transitive property, you can join multiple tables. Consider the `sales_table` and the `auth_table` in a Venn Diagram - there's no relation at all:
#
# 
#
# However, when the `book_table` enters the picture, suddenly the `sales_table` and `auth_table` have a connection:
#
# 
#
# Now we have an opportunity to join all three!
#
# ### Challenge:
# Write a query to show the first and last name of the author, the book title, the COGs, and the revenue from each transaction.
#
# ** Extra credit once you've completed the challenge**: Only return rows where the book was written by an English author. Sort your results so that the transaction with the most revenue appears first.
multi_join_chall = widgets.Textarea(value=
'',
width = '50em', height = '25em')
display(multi_join_chall)
multi_join_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(multi_join_chall_b)
run_q(multi_join_chall, multi_join_chall_b)
multi_join_chall_c ='''SELECT
first_name,
last_name,
book_table.book,
cogs,
revenue
FROM
book_table
JOIN auth_table
ON book_table.author = auth_table.last_name
JOIN
sales_table
ON book_table.book = sales_table.book'''
cheat(multi_join_chall_c)
# Answer to the extra credit:
multi_join_chall_c2 ='''SELECT
first_name,
last_name,
book_table.book,
cogs,
revenue
FROM
book_table
JOIN auth_table
ON book_table.author = auth_table.last_name
JOIN
sales_table
ON book_table.book = sales_table.book
WHERE
country = 'England'
ORDER BY
revenue DESC'''
cheat(multi_join_chall_c2)
# <img align="left" src="http://i.imgur.com/p6d18FV.png"> <br>
# **Use multiple joins sparingly:** Multiple joins can put a lot of strain on servers because SQL has to do a lot of work matching up all that data. The more tangential the relationship, the worse it gets. Avoid more than 2 degrees of separatation, and avoid joining 2 or more large tables. It's ok if one of your tables is big, but the others should be small.
# <a id='join_types'></a>
# <center>
# [Previous](#multi_join) | [Table of Contents](#table_of_contents) | [Next](#inner_joins)
# </center>
# 
#
# There are more ways to join two tables than the method we just covered. However, not all RDBMS support these different join methods. We'll learn about each of these methods, even if we can only practice 2 of them in SQLite.
#
# <font color='#1f5fd6'> Join Type |<font color='#1f5fd6'>Microsoft SQL Server | <font color='#1f5fd6'>MySQL | <font color='#1f5fd6'>Oracle | <font color='#1f5fd6'>SQLite </font>
# :----: | :------------------: | :---: | :----: | :----:
# `JOIN` or `INNER JOIN` | ✓ | ✓ | ✓ | ✓
# `LEFT JOIN` or `LEFT OUTER JOIN` | ✓ | ✓ | ✓ | ✓
# `RIGHT JOIN` or `RIGHT OUTER JOIN` | ✓ | ✓ | ✓ | not supported
# `OUTER JOIN` or `FULL OUTER JOIN` | ✓ | not supported | ✓ | not supported
#
# We're going to leave behind the book database for the next lesson, since a different data set will help illustrate the point a little better.
#
# Until now, the tables that we've joined have had columns that correspond perfectly - that is to say, every value that appears in one table also appears in the other. There aren't any authors that appear in the auth_table that don't also appear at least once in the book_table, and vice versa.
#
#
# Sometimes, you'll have two tables with corresponding columns, but they don't match perfectly.
# Consider the two tables below. The first lists tech companies and their CEOs, the second lists publicly traded companies and their share price.
#
# 
#
# Amazon, Alphabet, and Microsoft all appear on both tables. But Uber, SpaceX and AirBnB - which haven't IPO'd - aren't on the `public_cos` table. Conversely, Walmart, GE and P&G only appear on the `public_cos` table.
#
# Even though it's not a perfect match, there is some overlap. So, we can still use the `company` columns from each table as keys to join the tables:
# 
# ___________
# <a id='inner_joins'></a>
# <center>
# [Previous](#join_types) | [Table of Contents](#table_of_contents) | [Next](#left_joins)
# </center>
# 
# > **`SELECT` <br>
# `table_x.column_a,`** ➞ read this as "column_a from table_x"<br>
# ** `table_y.column_b,`** ➞ "column_b from table_y"<br>
# **`FROM`<br>
# `table_x`<br>
# `JOIN table_y`** ➞ SQL interprets `JOIN` and `INNER JOIN` as the same thing <br>
# **`ON table_x.key_column = table_y.key_column`**
#
# So what would happen if we tried to join `public_cos` and `tech_cos` using the method we just learned with the book database? We'll give it a shot:
inner_join_q = widgets.Textarea(value=
'''SELECT
*
FROM
tech_cos
JOIN public_cos
ON tech_cos.company = public_cos.company''',
width = '50em', height = '11em')
display(inner_join_q)
inner_join_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(inner_join_b)
run_q(inner_join_q, inner_join_b)
# This time, SQL can't find a match for every value in the two different **`company`** columns:
# 
#
# So it performs an **"INNER JOIN"**. You can write either **`JOIN`** or **`INNER JOIN`** - SQL will interpret them as the same thing. This eliminates any rows that don't have matching values, then combined the tables:
# 
# ____________
# <a id='left_joins'></a>
# <center>
# [Previous](#inner_joins) | [Table of Contents](#table_of_contents) | [Next](#outer_joins)
# </center>
# 
#
# > **`SELECT` <br>
# `table_x.column_a,`<br>
# `table_y.column_b,`<br>
# `FROM`<br>
# `table_x`<br>
# `LEFT JOIN table_y`** ➞ see all results from the first ("left") table & results *when available* from second table <br>
# **`ON table_x.key_column = table_y.key_column`**
#
#
# If you want to make sure you see all the rows from a particular table - even if there's no match in the other table - you can do a **`LEFT JOIN`** instead. It lets you prioritize the results from one table over another. Let's say your priority is to see all tech companies in your result-set, but you also want to see the `share_price` when that data is available:
left_q = widgets.Textarea(value=
'''SELECT
*
FROM
tech_cos
LEFT JOIN public_cos
ON tech_cos.company = public_cos.company''',
width = '50em', height = '11em')
display(left_q)
left_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(left_b)
run_q(left_q, left_b)
# With a **`LEFT JOIN`**, SQL still starts by looking for matching values.
# 
#
# When it fails to find a match, it will still keep the values on the "left" table, but get rid of the unmatched values on the "right" table.
# 
#
#
# ### Quick Exercise:
# The query from above has been reproduced below for these exercises.
# 1. Figure out how to change the query above so that you only see the company column appear once.
# 2. Rewrite the query so that **`public_cos`** becomes the priority table instead.
#
#
left_q2 = widgets.Textarea(value=
'''SELECT
*
FROM
tech_cos
LEFT JOIN public_cos
ON tech_cos.company = public_cos.company''',
width = '50em', height = '11em')
display(left_q2)
left_b2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(left_b2)
run_q(left_q2, left_b2)
# ________
#
# 
# > **`SELECT` <br>
# `table_x.column_a,`<br>
# `table_y.column_b,`<br>
# `FROM`<br>
# `table_x`<br>
# `RIGHT JOIN table_y`** ➞ see all results from the second ("right") table, results where available from first table <br>
# **`ON table_x.key_column = table_y.key_column`**
#
# It's exactly the same as `LEFT JOIN`, except it prioritizes the second table (the "right" table) over the first ("left") table. We can't practice it because SQLite doesn't support it, and it's super redundant anyway because we can just use `LEFT JOIN`. Boom. We're done with `RIGHT JOIN`.
#
# _____
# <a id='outer_joins'></a>
# <center>
# [Previous](#left_joins) | [Table of Contents](#table_of_contents) | [Next](#join_drills)
# </center>
# 
#
# > **`SELECT` <br>
# `table_x.column_a,`<br>
# `table_y.column_b,`<br>
# `FROM`<br>
# `table_x`<br>
# `OUTER JOIN table_y`** ➞ see all results from BOTH the first and second table <br>
# **`ON table_x.key_column = table_y.key_column`**
#
# What if you want to **see all values from both tables**? You can do this with an **`OUTER JOIN`**. Unfortunately, MySQL and SQLite (what we're using right now!) doesn't support it, so we can't practice it.
#
# If you are using Oracle or Microsoft SQL, then you'd use the example code above. For MySQL and SQLite, there is a workaround. You don't need to understand what's going on in the code for now, just look at the output to make sure you understand what the **`OUTER JOIN`** output *should* look like.
full_join_q = widgets.Textarea(value=
'''SELECT
ceo,
tech_cos.company,
public_cos.company,
share_price
FROM
tech_cos
LEFT JOIN public_cos ON tech_cos.company = public_cos.company
UNION ALL
SELECT
' ',
' ',
public_cos.company,
share_price
FROM
public_cos
WHERE
public_cos.company NOT IN (SELECT company FROM tech_cos)''',
width = '50em', height = '26em')
display(full_join_q)
full_join_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(full_join_b)
run_q(full_join_q, full_join_b)
# The **`OUTER JOIN`** starts out the same as the **`INNER JOIN`** and **`LEFT JOIN`**, trying to find matches wherever it can:
# 
#
# But when it can't find a match, instead of eliminating any the rows, it makes room for them:
# 
# __________
# ___________
# ___________
# <a id='join_drills'></a><center>
# [Previous](#outer_joins) | [Table of Contents](#table_of_contents) | [Next](#as)
# </center>
# 
insert_b = widgets.Button(description='Click here JUST ONCE before starting', width='20em', height='3em', color='white',background_color='#1f5fd6', border_color='#1f5fd6')
display(insert_b)
def insert_button(b):
insert_q1 = '''INSERT INTO auth_table VALUES ('Tolstoy', 'Leo', 'Russia', 1828)'''
insert_q2 = '''INSERT INTO auth_table VALUES ('Twain', 'Mark', 'USA', 1835)'''
insert_q3 = '''INSERT INTO book_table VALUES ('Jude the Obscure', '11.25', 'Hardy')'''
insert_q4 = '''INSERT INTO book_table VALUES ('The Age of Innocence', '14.20', 'Wharton')'''
query_list = [insert_q1, insert_q2, insert_q3, insert_q4]
for query in query_list:
run(query)
print('New rows have been added to auth_table and book_table!')
insert_b.on_click(insert_button)
# **We've now added some rows to `auth_table` and `book_table`** so we can practice our new join skills!
#
# ### Challenge
# Start simple by taking a look at the new rows we've added. Write a query to see all columns and rows from **`book_table`**, then change the query so you can take a look at **`auth_table`** instead:
join_drill1 = widgets.Textarea(value='',width = '50em', height = '7em')
display(join_drill1)
join_drill1_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_drill1_b)
run_q(join_drill1, join_drill1_b)
join_drill1_c ='''SELECT
*
FROM
auth_table [change to "book_table" for second part of challenge]'''
cheat(join_drill1_c)
# Now auth_table has 2 authors listed (Tolstoy and Twain) that don't appear on the `book_table`, and the `book_table` has two books (<u>Jude the Obscure</u> and <u>The Age of Innocence</u>) whose authors don't appear in the `auth_table`.
#
# ### Challenge:
# Write a query to view the book titles, first names, and last names of authors that appear on *both* the **`auth_table`** and the **`book_table`**.
join_drill2 = widgets.Textarea(value='', width = '50em', height = '12em')
display(join_drill2)
join_drill2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_drill2_b)
run_q(join_drill2, join_drill2_b)
join_drill2_c ='''SELECT
book,
author,
first_name
FROM
book_table
JOIN auth_table ON book_table.author = auth_table.last_name'''
cheat(join_drill2_c)
# _______
# ### Challenge:
# Write a query to see the titles of all the books from the **`book_table`**, and the author's **`country`** when that information is available.
#
join_drill3 = widgets.Textarea(value='', width = '50em', height = '13em')
display(join_drill3)
join_drill3_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_drill3_b)
run_q(join_drill3, join_drill3_b)
join_drill3_b ='''SELECT
book,
country
FROM
book_table
LEFT JOIN auth_table
ON book_table.author = auth_table.last_name'''
cheat(join_drill3_b)
# ### Quick Exercise:
# 1. Edit the query above so that you only see books by authors from England
# 2. Edit it so that you only see books by authors NOT from England.
# ________
# ### Challenge
# Write a query to see the first names of all authors in the **`auth_table`**, and the books they've written when that information is available.
join_drill4 = widgets.Textarea(value='',width = '50em', height = '16em')
display(join_drill4)
join_drill4_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_drill4_b)
run_q(join_drill4, join_drill4_b)
join_drill4_c ='''SELECT
first_name,
book
FROM
auth_table
LEFT JOIN book_table ON auth_table.last_name = book_table.author'''
cheat(join_drill4_c)
# ### Quick Exercise:
# 1. Change the query so that you only see results when the writer's first name is William, rerun
# 2. Change the query to sort the books in alphabethical order, rerun
# 3. Limit the number of rows to 3, rerun
join_drill4_ex_c ='''SELECT
first_name,
book
FROM
auth_table
LEFT JOIN book_table ON auth_table.last_name = book_table.author
WHERE
first_name = 'William'
ORDER BY
book
LIMIT 3'''
cheat(join_drill4_ex_c)
# ______
# ### Challenge:
# Write a query to return all books from the **`book_table`**, and their **`revenue`** data whenever that information is available. Try to figure out how you might sort your results so that you see books with no sales first.
join_drill4 = widgets.Textarea(value='',width = '50em', height = '12em')
display(join_drill4)
join_drill4_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(join_drill4_b)
run_q(join_drill4, join_drill4_b)
join_drill4_c ='''SELECT
book_table.book,
revenue
FROM
book_table
LEFT JOIN sales_table ON book_table.book = sales_table.book
ORDER BY
revenue'''
cheat(join_drill4_c)
# __________
# ___________
# ___________
# <a id='as'></a>
# <center>
# [Previous](#join_drills) | [Table of Contents](#table_of_contents) | [Next](#operators)
# </center>
# 
#
# # Assigning aliases to columns
#
# > **`SELECT` <br>
# `column_a AS alias_a`** ➞ creates an alias for column_a <br>
# **`FROM`**<br>
# **`table_name`**<br>
# `WHERE`<br>
# `alias_a = x` ➞ optional; use the alias in the `WHERE` clause<br>
# `ORDER BY`<br>
# `alias_a` ➞ optional; use the alias in the `ORDER BY` clause<br>
# `[LIMIT clause]` <br>
#
# **[Aliases](http://www.w3schools.com/sql/sql_alias.asp) allow you to rename columns and tables** in your query. They will come in handy as we learn to do more with the data.
#
# In plain English, the query below can be read as "*Show me the `book` column from the `book_table`, but rename the column to `book_title`*.
as_q = widgets.Textarea(value=
'''SELECT
book AS book_title
FROM
book_table''',
width = '50em', height = '14em')
display(as_q)
as_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(as_b)
run_q(as_q, as_b)
# ### Quick Exercises:
# 1. Change the query to rename the column to **`titles`**.
# 2. Delete the word **`AS`** and rerun. (You'll see that `AS` is totally optional when assigning aliases. It just make the query easier to read.)
# 3. Change the query so that you also pull the author, but rename the column **`author_name`**.
# 4. Change the query so that you only see books by **`Austen`**. Use the column's alias in your **`WHERE`** clause.
# 5. Order results by book in reverse alphabetical order. Use the column's alias in your **`ORDER BY`** clause.
as_ex_c ='''SELECT
book titles,
author AS author_name
FROM
book_table
WHERE
author_name = 'Austen'
ORDER BY
titles DESC'''
cheat(as_ex_c)
# ________
# ### Challenge:
# - Write a query to pull:
# - **`last_name`**, but renamed **`author`**
# - **`country`**, but renamed **`nationality`**
# - **`birth_year`**, but renamed **`year_born`**
# - Use each column's alias in the **`WHERE`** clause; use **`WHERE`** clause to only return results where the author is not from England, AND was born between 1800 and 1850.
as_chall = widgets.Textarea(value='', width = '50em', height = '15em')
display(as_chall)
as_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(as_chall_b)
run_q(as_chall, as_chall_b)
as_chall_c ='''SELECT
last_name AS author,
country as nationality,
birth_year AS year_born
FROM
auth_table
WHERE
nationality != 'England'
AND year_born BETWEEN 1800 and 1850'''
cheat(as_chall_c)
# _________
#
# # Assigning aliases to tables
# > **`SELECT` <br>
# `X.column_a,` <br>
# `Y.column_b` <br>
# `FROM` <br>
# `table_x X` ** ➞ assigns table_x the alias X <br>
# ** `JOIN table Y`** ➞ assigns table_y the alias Y <br>
# **`ON X.key_column = Y.key_column`** ➞ table aliases can be used as substitutes in the `table_x.column_a` format <br>
#
# When dealing with one or more tables in a query, we commonly assign capitalized one-letter aliases to tables. Writing `X.key_column` is much shorter than `table_x.key_column`, and coders like shortcuts. They also typically won't use `AS` when assigning aliases to tables (although it makes no difference either way).
#
# When you're dealing with only one table, it's unnecessary to use table aliases because SQL knows exactly what columns you are referring to. However, when you are dealing with 2 or more tables, particularly tables that have columns with the same names (like `book`, which is a column in both **`sales_table`** and **`book_table`**), then aliases are extremely handy.
as_table_q = widgets.Textarea(value=
'''SELECT
S.book,
S.revenue,
B.cogs
FROM
book_table B
JOIN sales_table S
ON S.book = B.book''',
width = '50em', height = '12em')
display(as_table_q)
as_table_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(as_table_b)
run_q(as_table_q, as_table_b)
# ### Quick Exercises:
# 1. In the **`SELECT`** clause, change **`S.book`** to just **`book`**, rerun. What's going wrong?
# 2. Now change **`book`** to **`B.book`**, rerun.
# 3. Change **`S.revenue`** to **`B.revenue`**, rerun. What's going wrong?
# 4. Change **`B.revenue`** to just **`revenue`** and rerun. Note that when you're joining tables, it's standard to use table aliases even on columns that don't need them. This makes it easier for someone to read your query even if they are unfamaliar with the tables that you're working with. However, as you see, it's not technically necessary.
# 5. Give each of these columns an alias (any alias) and rerun.
#
# _____
# ### Challenge:
# Write a query to view books and the author's country by joining **`auth_table`** and **`book_table`**. Give **`auth_table`** the alias **`A`** and **`book_table`** the alias **`B`**. Use the aliases in the **`ON`** part of the **`JOIN`** clause.
as_table_chall = widgets.Textarea(value='', width = '50em', height = '10em')
display(as_table_chall)
as_table_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(as_table_chall_b)
run_q(as_table_chall, as_table_chall_b)
as_table_chall_c ='''SELECT
B.book,
A.country
FROM
book_table B
JOIN auth_table A ON B.author = A.last_name'''
cheat(as_table_chall_c)
# _____
# ### Challenge:
# - Write a query to view these columns:
# - book titles with the alias **`titles`**
# - revenue with the alias **`earnings`**
# - author's last name with the alias **`author_name`**
# - year in which the author was born with the alias **`year_born`**
# - Use one-letter aliases for table names in your **`SELECT`** and **`JOIN`** clauses
# - For your **`WHERE`** and **`ORDER BY`** clauses:
# - Use column aliases
# - Only view results where author was born between 1700 and 1900 AND where revenue is more than $12.
# - Sort your results so that earnings appear in ascending order
# - Limit your results to 20 rows
as_chall2 = widgets.Textarea(value='', width = '50em', height = '20em')
display(as_chall2)
as_chall2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(as_chall2_b)
run_q(as_chall2, as_chall2_b)
as_chall2_c ='''SELECT
B.book as titles,
S.revenue as earnings,
A.birth_year as year_born
FROM
book_table B
JOIN auth_table A ON B.author = A.last_name
JOIN sales_table S on B.book = S.book
WHERE
year_born BETWEEN 1700 and 1900
AND earnings > 12
ORDER BY
earnings
LIMIT 20'''
cheat(as_chall2_b)
# __________
# ___________
# ___________
# <a id='operators'></a>
# <center>
# [Previous](#as) | [Table of Contents](#table_of_contents) | [Next](#functions)
# </center>
# 
#
# > **`SELECT` <br>
# `column_a + column_b,`** ➞ adds the values in `column_a` and `columns_b`<br>
# **`column_a - column_b,`** ➞ subtracts<br>
# **`column_a * column_b,`** ➞ multiplies<br>
# **`column_a / column_b,`** ➞ divides<br>
# **`(column_a + column_b) * column_c,`** ➞ use parentheses to make more complex calculations<br>
# **`FROM` <br>
# `table_name`** <br>
# `[WHERE clause]` <br>
# `[ORDER BY clause]` <br>
# `[LIMIT clause]` <br>
#
# This is pretty straightforward. Let's start by calculating gross profit per transaction: **`revenue`** minus **`cogs`**. Recall again that we can use `S.book` or `B.book` - we'll get the same results.
op_q = widgets.Textarea(value=
'''SELECT
B.book,
S.revenue,
B.cogs,
S.revenue - B.cogs
FROM
book_table B
JOIN sales_table S ON B.book = S.Book''',
width = '50em', height = '18em')
display(op_q)
op_q_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(op_q_b)
run_q(op_q, op_q_b)
# ### Quick Exercise:
# 1. Give the calculated column the alias **`gross_grofit`**, rerun. See how nice aliases are?
# 2. Add a **`WHERE`** clause to only see transactions where **`gross_grofit`** is over $5, rerun.
# 3. Add an **`ORDER BY`** clause to sort by **`gross_profit`** with the most profitable transaction is listed first, rerun.
#
# ____
# ### Challenge:
# - Pull book name and author's last name
# - Calculate the gross margin per transaction, give the calculated column the alias **`gross_margin`**
# - Use one-letter aliases for all the table names
# - Only return rows where the author's name is NOT Faulkner or Austen
# - Sort your results with the highest margin transaction listed first
# - Limit your results to 10 rows
op_chall = widgets.Textarea(value='',width = '50em', height = '18em')
display(op_chall)
op_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(op_chall_b)
run_q(op_chall, op_chall_b)
op_chall_c ='''SELECT
B.book,
B.author,
(S.revenue - B.cogs) / S.revenue AS gross_margin
FROM
book_table B
JOIN sales_table S ON B.book = S.book
WHERE
B.author NOT IN ('Falkner', 'Austen')
ORDER BY
gross_margin DESC
LIMIT 10'''
cheat(op_chall_c)
# # Concatonating
# <font color='#1f5fd6'>Microsoft SQL Server | <font color='#1f5fd6'>MySQL | <font color='#1f5fd6'>Oracle | <font color='#1f5fd6'>SQLite </font>
# :------------------: | :---: | :----: | :----:
# `CONCAT(column_a, column_b)` or `+` | `CONCAT(column_a, column_b)` | `CONCAT(column_a, column_b)` or || | ||
#
# > **`SELECT` <br>
# `column_a || column_b,`** ➞ *combines the characters of column_a & column_b *<br>
# ** `column_a || ' ' || column_b`** ➞ * combines the characters of column_a & column_b with a space inbetween *<br>
# **`FROM` <br>
# `table_name`**
#
# This one is extremely straightforward. This allows you to non-mathematically combine values. So "**`some || word`**" becomes "**`someword`**", and "**`some || ' ' || word`**" becomes "**`some word`**".
conc_q = widgets.Textarea(value=
'''SELECT
first_name || last_name
FROM
auth_table''',
width = '50em', height = '7em')
display(conc_q)
conc_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(conc_b)
run_q(conc_q, conc_b)
# ### Quick Exercises:
# 1. Fix the query so that there is a space between the names, rerun
# 2. Give the concatenated column an alias, rerun
# 3. Rewrite the query so that it follows the format "last_name, first_name" instead
# __________
# ___________
# ___________
# <a id='functions'></a>
# <center>
# [Previous](#operators) | [Table of Contents](#table_of_contents) | [Next](#group_by)
# </center>
# 
#
# > **`SELECT`** <br>
# **`SOME_FUNCTION(column_a),`** ➞ performs the function on the column <br>
# **`FROM`** <br>
# **`table_name`**<br>
# `[WHERE clause]` <br>
# `[ORDER BY clause]` <br>
# `[LIMIT clause]` <br>
#
# [Functions](http://www.w3schools.com/sql/sql_functions.asp) work similar to funcitons in Excel - you can apply them to entire columns. There are tons more functions than the ones listed below, just Google what you want to do to find more.
#
# #### Short List of Functions:
# FUNCTION | DESCRIPTION
# :------- | :-------------
# `AVG(col)` | Averages values
# `COUNT(col)` | Counts the number of rows with non-null values in the column
# `COUNT(*)` | Counts the number of rows in the table
# `COUNT(DISTINCT(col))` | Counts the number of unique values in the column
# `GROUP_CONCAT(col, 'separator')` | Returns a comma-separated list of values, specify a separator in quotes
# `MAX(col)` | Returns the maximum value
# `MIN(col)` | Returns the minimum value
# `ROUND(AVG(col), x)` | Rounds value to x decimals
# `SUM(col)` | Sums values
# `UPPER(col)` | If column is text, it will return all-caps version of the text
#
#
# First, we'll start with **`SUM()`** to find the total revenue for all our transactions.
sum_q = widgets.Textarea(value=
'''SELECT
sum(revenue)
FROM
sales_table''',
width = '50em', height = '17em')
display(sum_q)
sum_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(sum_b)
run_q(sum_q, sum_b)
# ### Quick Exercises:
# 1. Give the calculated column an alias, rerun
# 2. Add a line to the **`SELECT`** clause to find the average revenue per transaction and give the column an alias, rerun
# 3. Edit the average column so that your results are rounded to the nearest cent, rerun
# 4. Add a line to the **`SELECT`** clause to count the total number of transactions and give the column an alias, rerun
# 5. Add a 2 lines to the **`SELECT`** clause to see the minimum and maximum revenue earned on a single transaction, rerun
# 6. Add a line to the **`SELECT`** clause to see a count of the number of *distinct* books that appear in **`sales_table`**
# 7. Add a **`WHERE`** clause to only see results for the books "For Whom the Bell Tolls" and "Emma"
sum_q_c ='''SELECT
SUM(revenue) AS total_rev,
ROUND(AVG(revenue), 2) AS avg_rev,
COUNT(revenue) AS total_transactions,
MAX(revenue) AS max_rev,
MIN(revenue) AS min_rev,
COUNT(DISTINCT(book)) AS distinct_books
FROM
sales_table
WHERE
book IN ('For Whom the Bell Tolls', 'Emma') '''
cheat(sum_q_c)
# ______
# ### Challenge:
# Write a query to find the average cost of goods for books whose authors are from the US (**`USA`**). Round the number to the nearest cent. Use an alias for your column.
function_chall = widgets.Textarea(value='', width = '50em', height = '11em')
display(function_chall)
function_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(function_chall_b)
run_q(function_chall, function_chall_b)
function_chall_c ='''SELECT
ROUND(AVG(B.cogs), 2) AS avg_cogs
FROM
book_table B
JOIN auth_table A ON B.author = A.last_name
WHERE
A.country = 'USA'
'''
cheat(function_chall_c)
# ________
# ### Challenge:
# Try out the **`GROUP_CONCAT`** function. Write a query to select **`GROUP_CONCAT(last_name)`** from the **`auth_table`**, only return results where the author is NEITHER Austen NOR Shakespeare. After you get your query to work, change it to **`GROUP_CONCAT(last_name, ' / ')`** and rerun.
function_chall2 = widgets.Textarea(value='',width = '50em', height = '10em')
display(function_chall2)
function_chall2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(function_chall2_b)
run_q(function_chall2, function_chall2_b)
function_chall2_c ='''SELECT
GROUP_CONCAT(last_name)
FROM
auth_table
WHERE
last_name NOT IN ('Austen', 'Shakespeare')
'''
cheat(function_chall2_c)
# # `COUNT(*)` vs `COUNT(column_name)`
insert_null_b = widgets.Button(description='Click here JUST ONCE before starting', width='20em', height='3em', color='white',background_color='#1f5fd6', border_color='#1f5fd6')
display(insert_null_b)
def insert_button(b):
null_query1 = '''INSERT INTO auth_table VALUES ('Homer', NULL, 'Greece', NULL)'''
run(null_query1)
print('A new row has been added to the auth_table!')
insert_null_b.on_click(insert_button)
# It's much more common to use **`COUNT(*)`** than **`COUNT(column_name)`** when you are trying to get a count of the number of rows in your result-set. This is because **`COUNT(*)`** will capture all rows, while **`COUNT(column_name)`** will skip over NULL values in that particular row.
#
# ### Challenge:
# We've just added new a new row to the **`auth_table`** that has some NULL (blank) values. Start by writing a query to view everything (`SELECT *`) in the **`auth_table`** and make a note of the new row.
star_chall = widgets.Textarea(value='', width = '50em', height = '12em')
display(star_chall)
star_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(star_chall_b)
run_q(star_chall, star_chall_b)
# ### Challenge Continued...
# 1. Delete **`*`** and replace it with **`COUNT(first_name)`**, rerun
# 2. Add a line (but don't erase anything) to the `SELECT` clause: **`COUNT(*)`**
star_chall_c ='''SELECT
COUNT(first_name),
COUNT(*)
FROM
auth_table
'''
cheat(star_chall_c)
# ### What's going on:
# When you use `COUNT(first_name)`, SQL skipped over the Homer row because there was no value in the first_name column. `COUNT(*)`, on the other hand, looks across all columns, so as long as a row has a value in at least one column, it'll get included in the count. You might argue that you could just use `country` or `last_name`, but they fact is that **`*`** is just way easier and less time-consuming to type out. **Overwhelmingly, people opt for `COUNT(*)` instead of `COUNT(column_name)`** unless they are interested in overlooking NULL values.
# _______
# # Functions + Operators
#
# You can use functions together with operators to do more complex calculations. Below, we've calculated our total gross profit using both the `SUM()` function and subtraction:
fun_op_q = widgets.Textarea(value=
'''SELECT
SUM(S.revenue) - SUM(B.cogs) AS gross_profit
FROM
book_table B
JOIN sales_table S ON S.book = B.book''',
width = '50em', height = '8em')
display(fun_op_q)
fun_op_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(fun_op_b)
run_q(fun_op_q, fun_op_b)
# ### Quick Exercise:
# Rewrite the `SELECT` clause so that you get the same results but only have to use `SUM` once.
# ___________
# ### Challenge:
# Write a query to view gross *margin* for all transactions using functions in conjunction with operators. Extra credit: round your results to the nearest cent.
fun_op_chall = widgets.Textarea(value='', width = '50em', height = '8em')
display(fun_op_chall)
fun_op_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(fun_op_chall_b)
run_q(fun_op_chall, fun_op_chall_b)
fun_op_chall_c ='''SELECT
SUM(S.revenue - B.cogs) / SUM(S.revenue) AS gross_margin
FROM
book_table B
JOIN sales_table S ON S.book = B.book
- OR - you can use AVG() instead of SUM() for all functions'''
cheat(fun_op_chall_c)
# __________
# ___________
# ___________
# <a id='group_by'></a>
# <center>
# [Previous](#functions) | [Table of Contents](#table_of_contents) | [Next](#having)
# </center>
# 
#
# > **`SELECT`** <br>
# **`column_a,`**
# **`SUM(column_b)`** ➞ sums up the values in column_b <br>
# **`FROM`** <br>
# **`table_name`** <br>
# `[WHERE clause]` <br>
# **`GROUP BY`** ➞ creates one group for each unique value in column_a <br>
# **`column_a`** <br>
# `[ORDER BY clause]` <br>
# `[LIMIT clause]`
#
#
# [**`GROUP BY`**](http://www.w3schools.com/sql/sql_groupby.asp) creates a group for each unique value in the column you specify. You'll always use it in conjunction with functions - it creates segments for your results. In plain English, the query below says: "*Show me the average `revenue` per `book` from the `sales_table`*"
group_q = widgets.Textarea(value=
'''SELECT
book,
AVG(revenue)
FROM
sales_table
GROUP BY
book''',
width = '50em', height = '20em')
display(group_q)
group_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(group_b)
run_q(group_q, group_b)
# ### Quick Exercises:
# 1. Change **`AVG()`** to **`SUM()`**, rerun
# 2. Give the **`book`** column the alias **`book_title`**, then use the alias in the **`GROUP BY`** clause, rerun
# 3. Sort the results so that the most profitable book is listed first, rerun
# 4. Add this to the **`SELECT`** clause: **`COUNT(*)`**, rerun
# 5. Add a **`WHERE`** clause to only return results that are **not** written by Faulkner (hint: you'll have to join a table for this)
#
# _________
# ### Challenge:
# Write a query to count the number of books each author has listed in the **`book_table`**.
group_chall = widgets.Textarea(value='', width = '50em', height = '12em')
display(group_chall)
group_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(group_chall_b)
run_q(group_chall, group_chall_b)
group_chall_c ='''SELECT
author,
count(*)
FROM
book_table
GROUP BY
author'''
cheat(group_chall_c)
# ______
# ### Challenge:
# Write a query that joins the **`book_table`** and the **`sales_table`** to see total revenue per author.
group_chall2 = widgets.Textarea(value='', width = '50em', height = '12em')
display(group_chall2)
group_chall2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(group_chall2_b)
run_q(group_chall2, group_chall2_b)
group_chall2_c ='''SELECT
B.author,
SUM(S.revenue)
FROM
book_table B
JOIN sales_table S ON B.book = S.book
GROUP BY
B.author'''
cheat(group_chall2_c)
# ____
# ### Challenge:
# Write a query to see the maximum and minimum prices that each book sold for, but don't include Macbeth or Hamlet in your result-set:
group_chall3 = widgets.Textarea(value='',width = '50em', height = '15em')
display(group_chall3)
group_chall3_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(group_chall3_b)
run_q(group_chall3, group_chall3_b)
group_chall3_c ='''SELECT
book,
MAX(revenue),
MIN(revenue)
FROM
sales_table
WHERE
book NOT IN ('Macbeth','Hamlet')
GROUP BY
book'''
cheat(group_chall3_c)
# ______
# ______
# # `GROUP BY` + Functions + Operators
#
# You can use `GROUP BY` with functions and operators to do more complex analysis. Below, we use `SUM()` and the subtraction operator to see gross profit for each book.
group_fun_op_q = widgets.Textarea(value=
'''SELECT
B.book,
SUM(S.revenue) - SUM(B.cogs) AS gross_profit
FROM
sales_table S
JOIN book_table B ON S.book = B.book
GROUP BY
B.book''',
width = '50em', height = '12em')
display(group_fun_op_q)
group_fun_op_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(group_fun_op_b)
run_q(group_fun_op_q, group_fun_op_b)
# ________
# ### Challenge:
# Write a query to find the gross margin per author using **`GROUP BY`, functions and operators.** Give the gross margin column an alias.
group_func_op_chall = widgets.Textarea(value='', width = '50em', height = '13em')
display(group_func_op_chall)
group_func_op_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(group_func_op_chall_b)
run_q(group_func_op_chall, group_func_op_chall_b)
group_func_op_chall_c ='''SELECT
B.author,
(SUM(S.revenue) - SUM(B.cogs)) / SUM(S.revenue) AS gross_margin
FROM
book_table B
JOIN sales_table S ON B.book = S.book
GROUP BY
B.author'''
cheat(group_func_op_chall_c)
# _______
# ### Challenge:
# - Copy and paste the query you just wrote for the previous challenge.
# - Add the following columns in the **`SELECT`** clause (in addition to author and gross margin columns), and give every column an alias:
# - total revenue
# - total cogs
# - a count of the number of individual transactions
# - BONUS: a count of the distinct book titles sold
# - BONUS: a comma-separated list of the book titles with no repeats
# - Only include results where the author isn't Faulker and the book isn't Hamlet
# - Sort your results so that the author with the highest average gross margin is listed first
group_chall3 = widgets.Textarea(value='', width = '50em', height = '26em')
display(group_chall3)
group_chall3_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(group_chall3_b)
run_q(group_chall3, group_chall3_b)
group_chall3_c ='''SELECT
B.author AS author_name,
SUM(S.revenue) AS total_revenue,
SUM(B.cogs) AS total_cogs,
(SUM(S.revenue) - SUM(B.cogs))/SUM(S.revenue) AS gross_margin,
COUNT(*) AS transaction_count,
COUNT(DISTINCT(S.book)) AS distinct_book_titles,
GROUP_CONCAT(DISTINCT(S.book)) AS book_list
FROM
book_table B
JOIN sales_table S ON B.book = S.book
WHERE
author_name != 'Faulkner'
AND S.book != 'Hamlet'
GROUP BY
author_name
ORDER BY
gross_margin DESC'''
cheat(group_chall3_c)
# __________
# ___________
# ___________
# <a id='having'></a>
# <center>
# [Previous](#group_by) | [Table of Contents](#table_of_contents) | [Next](#case_when)
# </center>
# 
#
# > **`SELECT`** <br>
# **`column_a,`** <br>
# **`FUNCTION(column_b)`** <br>
# **`FROM`** <br>
# **`table_name`** <br>
# `[WHERE clause]` <br>
# **`GROUP BY`** <br>
# **`column_a HAVING FUNCTION(column_b) > x`** ➞ returns groups whose value is greater than x <br>
# `[ORDER BY clause]` <br>
# `[LIMIT clause]` <br>
#
# Use [**`HAVING`**](http://www.w3schools.com/sql/sql_having.asp) with `GROUP BY` in order to filter out groups that don't meet your criteria. Below, the plain English translation of this query says, "*show my the total revenue for each book, but only show me books that have total revenue over $100*"
having_q = widgets.Textarea(value=
'''SELECT
book,
SUM(revenue)
FROM
sales_table
GROUP BY
book HAVING SUM(revenue) > 100''',
width = '50em', height = '11em')
display(having_q)
having_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(having_b)
run_q(having_q, having_b)
# ### Quick Exercises:
# 1. Change the > to <, rerun
# 2. Give the **`SUM(revenue)`** column an alias, and change the **`GROUP BY`** clause so that you're using the alias instead, rerun
# 3. Think about why this is different from **`WHERE`**. Take a moment to discuss this with your partner in class.
# ____
# ### Challenge:
# Write a query to see average COGs per author, but use `HAVING` to return authors whose average COGs is greater than $10. Assign the average COGs column an alias and use it in the `GROUP BY` clause.
having_chall = widgets.Textarea(value='',width = '50em', height = '10em')
display(having_chall)
having_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(having_chall_b)
run_q(having_chall, having_chall_b)
having_chall_c ='''SELECT
author,
AVG(cogs) AS avg_cogs
FROM
book_table
GROUP BY
author HAVING avg_cogs > 10'''
cheat(having_chall_c)
# ### `HAVING` vs. `WHERE`
# `HAVING` and `WHERE` both let you change the results you see in your result-set, but they operate quite differently. Take a look at the query below. It looks at the average cogs per author, but uses a `WHERE` clause to filter out 'Faulkner':
hw_q1 = widgets.Textarea(value=
'''SELECT
author,
AVG(cogs)
FROM
book_table
WHERE
author != 'Faulkner'
GROUP BY
author''',
width = '50em', height = '14em')
display(hw_q1)
hw_b1 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(hw_b1)
run_q(hw_q1, hw_b1)
# ### Quick Exercise:
# Now let's say you want to filter out Faulkner AND you only want to see authors whose average COGs are over $11. Your first thought might be to use the `WHERE` clause. Add `AND AVG(cogs) > 11`, rerun. Why do you think you're hitting an error?
#
# ______
# You hit an error because the `AVG(cogs)` column was created by a function, and SQL doesn't let you put function-generated columns in the `WHERE` clause. You have to use `HAVING` instead. The query below will accomplish what we're trying to do, and returns a result-set that doesn't include Faulkner AND only shows authors whose average COGs are over $11.
hw_q2 = widgets.Textarea(value=
'''SELECT
author,
AVG(cogs)
FROM
book_table
WHERE
author != 'Faulkner'
GROUP BY
author HAVING AVG(cogs) > 11''',
width = '50em', height = '14em')
display(hw_q2)
hw_b2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(hw_b2)
run_q(hw_q2, hw_b2)
# This seems relatively straightforward - but it's easy to forget and wind up with inaccurate results. Consider the following query. Why would this be wrong?
hw_q3 = widgets.Textarea(value=
'''SELECT
author,
AVG(cogs)
FROM
book_table
WHERE
author != 'Faulkner'
AND cogs > 11
GROUP BY
author''',
width = '50em', height = '15em')
display(hw_q3)
hw_b3 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(hw_b3)
run_q(hw_q3, hw_b3)
# These results are inaccurate because instead of telling SQL to only return authors with average COGs over \$11, we've told SQL "only consider rows where COGs are over \$11". SQL dropped the rows with COGs under \$11 *before* it started grouping and averaging.
#
# ### SQL's Order of Execution
# When we read, we start at the top of a page and work our way to the bottom. That's not how SQL works. It actually starts with the `FROM` clause and jumps around. It's helpful to understand the order it follows to determine when to use `HAVING` and when to use `WHERE`.
#
# <!-- This will also help clear up some other issues. Ever wonder why you can mention a column in the `WHERE` clause that you don't mention in the `SELECT` clause? Or why SQL knows the table alias that you're referring to in the `SELECT` clause even though you don't assign aliases to tables until later in the query? This is why.-->
#
# We *write* the clauses in this order:
# > `SELECT` <br>
# `FROM` <br>
# `JOIN...ON` <br>
# `WHERE` <br>
# `GROUP BY...HAVING` <br>
# `ORDER BY` <br>
# `LIMIT` <br>
#
# However, SQL *reads and executes* the clauses in this order:
# > `FROM` <br>
# `JOIN...ON` <br>
# `WHERE` <br>
# `SELECT` <br>
# `GROUP BY...HAVING` <br>
# `ORDER BY` <br>
# `LIMIT` <br>
#
# Here's a query we've seen before, but now we've added a few more clauses so that we can see all of them in action:
sql_order_q = widgets.Textarea(value=
'''SELECT
author,
AVG(cogs)
FROM
book_table
WHERE
author != 'Faulkner'
GROUP BY
author HAVING AVG(cogs) > 11
ORDER BY
AVG(cogs)
LIMIT 3''',
width = '50em', height = '17.5em')
display(sql_order_q)
sql_order_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(sql_order_b)
run_q(sql_order_q, sql_order_b)
# The GIF below shows the order that SQL follows the steps:
# 
# Let's revisit the query that gave us the skewed average:
hw_q4 = widgets.Textarea(value=
'''SELECT
author,
AVG(cogs)
FROM
book_table
WHERE
author != 'Faulkner'
AND cogs > 11
GROUP BY
author''',
width = '50em', height = '15em')
display(hw_q4)
hw_b4 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(hw_b4)
run_q(hw_q4, hw_b4)
# Now that we know the order in which SQL executes commands, we can see what went wrong. The rows with COGs under $11 were eliminated before SQL averaged COGs for each group:
# 
# ### Challenge:
# Write a query to join `book_table` and `sales_table`. Select author and total revenue, but only return authors whose total revenue was over $200
have_chall2 = widgets.Textarea(value='',width = '50em', height = '12em')
display(have_chall2)
have_chall2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(have_chall2_b)
run_q(have_chall2, have_chall2_b)
have_chall2_c ='''SELECT
B.author,
SUM(S.revenue) AS total_rev
FROM
book_table B
JOIN sales_table S ON B.book = S.book
GROUP BY
author HAVING total_rev > 200'''
cheat(have_chall2_c)
# ______
# ### Challenge:
# Write a query to join the `auth_table` with the `sales_table` (remember that this requires multiple joins). Count the number of sales per country (author's country of origin in the `auth_table`), but don't include sales from Hemingway.
have_chall3 = widgets.Textarea(value='', width = '50em', height = '16em')
display(have_chall3)
have_chall3_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(have_chall3_b)
run_q(have_chall3, have_chall3_b)
have_chall3_c ='''SELECT
A.country,
COUNT(*) AS count_of_sales
FROM
sales_table S
JOIN book_table B ON S.book = B.book
JOIN auth_table A ON A.last_name = B.author
WHERE
A.last_name != 'Hemingway'
GROUP BY
A.country
NOTE: you can also use "B.author != 'Hemingway'" in the WHERE clause to get the same results'''
cheat(have_chall3_c)
# _________
# _________
# _________
# <a id='case_when'></a>
# <center>
# [Previous](#having) | [Table of Contents](#table_of_contents) | [Next](#nesting)
# </center>
# 
#
# Conditional Type | <font color='#1f5fd6'>Microsoft SQL Server | <font color='#1f5fd6'>MySQL | <font color='#1f5fd6'>Oracle | <font color='#1f5fd6'>SQLite </font>
# :--------------- |:------------------ |:--- |:---- |:----
# IF | `IF logical_test PRINT value_if_true ` | `IF(logical_test, value_if_true, value_if_false)` (same as Excel) | `IF logical_test THEN value_if_true ELSIF...END IF` | NOT SUPPORTED
# CASE WHEN | ✓ | ✓ | ✓ | ✓
#
# > **`SELECT`** <br>
# **`CASE WHEN some_column = x THEN value_if_true`** <br>
# **`WHEN some_column = y THEN other_value_if_true`** <br>
# **`ELSE value_if_false`** <br>
# **`END`** <br>
# **`FROM`** <br>
# **`some_table`** <br>
#
# Because SQLite doesn't support `IF` statements, we're going to focus on `CASE WHEN`. `CASE WHEN` lets you accomplish the same thing by setting logical tests and conditional values, but it has the added bonus of freeing you from ever needing to nest multiple `IF` statements.
#
# Let's start very simple. The following query uses a logical test to create a column where the value is "true" if the author is Austen, and "false" if the author is not Austen:
case1_q = widgets.Textarea(value=
'''SELECT
last_name,
CASE WHEN last_name = 'Austen' THEN 'True'
ELSE 'False'
END
FROM
auth_table''',
width = '50em', height = '13em')
display(case1_q)
case1_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case1_b)
run_q(case1_q, case1_b)
# ### Quick Exercises:
# 1. Give the `CASE WHEN` column an alias (immediately after `END`), rerun
# 2. Change the query so that instead of "True", the query returns Austen's first name (use the first_name column), rerun
# 3. Add something to the `CASE WHEN` column so that the query returns Faulkner's first name as well (look at the example code above for help), rerun.
#
# ___
#
# ### Using `CASE WHEN` to create categories
# `CASE WHEN` allows you to set multiple logical tests, which can help you create buckets or categories. In Excel, you'd have to nest multiple logical tests in an `IF` statement (ie. IF(logical_test, value_if_true, IF(other_logical_test, value_if_true, value_if_false))...very messy). With `CASE WHEN`, you can add an infinite number very easily.
#
# Let's say that rather than caring about the exact revenue for each transaction, you only cared whether it was under \$10, between \$10 and \$15, or over \$15. That's easy to do with `CASE WHEN`. We'll include the `revenue` column as well so you can more easily see what's going on:
case2_q = widgets.Textarea(value=
'''SELECT
book,
revenue,
CASE WHEN revenue < 10 THEN "<$10"
WHEN revenue BETWEEN 10 AND 15 THEN "$10-15"
WHEN revenue > 15 THEN ">$15"
END AS revenue_category
FROM
sales_table
''',
width = '50em', height = '15em')
display(case2_q)
case2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case2_b)
run_q(case2_q, case2_b)
# This might not immediately seem useful, but when you start grouping by your newly created categories, you'll be able to do all kinds of new analysis. Consider the query below. We use `CASE WHEN` to create a column that we then use in the `GROUP BY` clause. We've essentially created new groups where there were none before, and now we can assess the number of sales and total revenue from each revenue group.
case3_q = widgets.Textarea(value=
'''SELECT
CASE WHEN revenue < 10 THEN "<$10"
WHEN revenue BETWEEN 10 AND 15 THEN "$10-15"
WHEN revenue > 15 THEN ">$15"
END AS revenue_category,
COUNT(*) AS total_sales,
SUM(revenue) AS total_revenue
FROM
sales_table
GROUP BY
revenue_category
''',
width = '50em', height = '18em')
display(case3_q)
case3_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case3_b)
run_q(case3_q, case3_b)
# ____
# ### Challenge:
# Suppose you want to see total revenues broken out by male vs. female authors. Use `CASE WHEN` to create these groups - with Austen in the "female" group and Faulkner, Hemingway, and Shakespeare in the "male" group.
case_chall = widgets.Textarea(value='', width = '50em', height = '16em')
display(case_chall)
case_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case_chall_b)
run_q(case_chall, case_chall_b)
case_chall_c ='''SELECT
CASE WHEN
B.author = 'Austen' THEN 'Female'
ELSE 'Male' --- or you can say, "WHEN B.author IN ('Faulkner', 'Shakespeare', 'Hemingway') THEN 'Male' "
END AS gender,
SUM(S.revenue) AS total_revenue
FROM
book_table B
JOIN sales_table S ON B.book = S.book
GROUP BY
gender'''
cheat(case_chall_c)
# ____
# ## Using `CASE WHEN` to create a pivot table
# Say you want to see revenue broken out by gender *and* by date. Right now, the only way we know how to do this is to add "date" to the `GROUP BY` clause. The query below is the same as the one from your last challenge, only we've added `date` to both the `SELECT` clause and the `GROUP BY` clause.
case_pivot_ex = widgets.Textarea(value='''SELECT
date,
CASE WHEN
B.author = 'Austen' THEN 'Female'
ELSE 'Male'
END AS gender,
SUM(S.revenue) AS total_revenue
FROM
book_table B
JOIN sales_table S ON B.book = S.book
GROUP BY
date, gender'''
, width = '50em', height = '18em')
display(case_pivot_ex)
case_pivot_ex_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case_pivot_ex_b)
run_q(case_pivot_ex, case_pivot_ex_b)
# The result-set above is almost useless - it's impossible to do meaningful analysis when you group by multiple columns. Instead, we'll use `CASE WHEN` nested inside a function to essentially create a pivot table:
case_pivot = widgets.Textarea(value=
'''SELECT
date,
SUM(CASE WHEN B.author = 'Austen' THEN revenue END) AS Female_Rev,
SUM(CASE WHEN B.author != 'Austen' THEN revenue END) AS Male_Rev
FROM
book_table B
JOIN sales_table S ON B.book = S.book
GROUP BY
date''',
width = '50em', height = '14em')
display(case_pivot)
case_pivot_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case_pivot_b)
run_q(case_pivot, case_pivot_b)
# Note that while this `CASE WHEN` method will work in other RDBMSs, it's more common to use `IF` when you are only using a single logical test. In MySQL, for instance, the line for Female_Rev would look like this instead, which would translate to *"sum up the revenue for any row where the author is Austen, and the number 0 whenever the author is not Austen":*
# > **`SUM(IF(B.author = 'Austen', revenue, 0))`**
#
# ____
# ### Quick Exercises:
# The query from above has been reproduced below for these exercises (so you don't have to keep scrolling up and down).
# 1. Change the query so that you have separate columns for each individual author's revenue, rerun
# 2. Change `SUM` to `AVG`, rerun
# 3. Change `AVG` to `COUNT` - note that with conditional statements, you don't use an asterisk with `COUNT`. You need to stick with a specific column name
case_pivot1 = widgets.Textarea(value=
'''SELECT
date,
SUM(CASE WHEN B.author = 'Austen' THEN revenue END) AS Female_Rev,
SUM(CASE WHEN B.author != 'Austen' THEN revenue END) AS Male_Rev
FROM
book_table B
JOIN sales_table S ON B.book = S.book
GROUP BY
date''',
width = '50em', height = '18em')
display(case_pivot1)
case_pivot_b1 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case_pivot_b1)
run_q(case_pivot1, case_pivot_b1)
# ______
# ### Challenge:
# - In the **`SELECT`** clause:
# - Use **`CASE WHEN`** to create a column that creates buckets for author's **`birth_year`**: "Before 1700", "1700-1800", "After 1800"
# - Use **`CASE WHEN`** to create a column that returns the count of books by authors from USA
# - Use **`CASE WHEN`** to create a column that returns the count of books by authors from England
# - **`GROUP BY`** the birth_year bucket column that you created
case_pivot_chall = widgets.Textarea(value='',width = '50em', height = '18em')
display(case_pivot_chall)
case_pivot_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case_pivot_chall_b)
run_q(case_pivot_chall, case_pivot_chall_b)
case_pivot_chall_c ='''SELECT
CASE WHEN A.birth_year < 1700 THEN "Before 1700"
WHEN A.birth_year BETWEEN 1700 AND 1800 THEN "1700-1800"
WHEN A.birth_year > 1800 THEN "After 1800"
END AS era,
COUNT(CASE WHEN A.country = 'USA' THEN book END) AS count_from_USA,
COUNT(CASE WHEN A.country = 'England' THEN book END) AS count_from_England
FROM
book_table B
JOIN auth_table A ON B.author = A.last_name
GROUP BY
era
'''
cheat(case_pivot_chall_c)
# ____
# ### Challenge:
# - Write a query that returns a *daily* count of the sales of:
# - <u>For Whom the Bell Tolls</u> (in its own column)
# - <u>Emma</u> (in its own column)
# - <u>Macbeth</u> and <u>Hamlet</u> (in a combined column)
case_pivot_chall2 = widgets.Textarea(value='', width = '50em', height = '14em')
display(case_pivot_chall2)
case_pivot_chall2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(case_pivot_chall2_b)
run_q(case_pivot_chall2, case_pivot_chall2_b)
case_pivot_chall2_c ='''SELECT
date,
COUNT(CASE WHEN book = 'For Whom the Bell Tolls' THEN revenue END) Bell_Tolls_Count,
COUNT(CASE WHEN book = 'Emma' THEN revenue END) Emma_Count,
COUNT(CASE WHEN book IN ('Macbeth', 'Hamlet') THEN revenue END) Macbeth_Hamlet_Count
FROM
sales_table
GROUP BY
date'''
cheat(case_pivot_chall2_c)
# __________
# ___________
# ___________
# <a id='nesting'></a>
# <center>
# [Previous](#case_when) | [Table of Contents](#table_of_contents) | [Next](#union)
# </center>
# 
# >**`SELECT`** <br>
# **`column_a`** <br>
# **`FROM`** <br>
# **`table_x`** <br>
# **`WHERE`** <br>
# **`column_a IN (SELECT column_b FROM table_y)`**
#
# **Read first**: For this section, we'll need the extra rows in the `auth_table` and `book_table` that we added during the `JOIN` exercises. If you've closed the program or re-run it since you last added those rows, then click the button below to re-add them.
insert_b = widgets.Button(description="Read the paragraph above before clicking", width='25em', height='3em', color='white',background_color='#1f5fd6', border_color='#1f5fd6')
display(insert_b)
def insert_button(b):
insert_q1 = '''INSERT INTO auth_table VALUES ('Tolstoy', 'Leo', 'Russia', 1828)'''
insert_q2 = '''INSERT INTO auth_table VALUES ('Twain', 'Mark', 'USA', 1835)'''
insert_q3 = '''INSERT INTO book_table VALUES ('<NAME>', '11.25', 'Hardy')'''
insert_q4 = '''INSERT INTO book_table VALUES ('The Age of Innocence', '14.20', 'Wharton')'''
query_list = [insert_q1, insert_q2, insert_q3, insert_q4]
for query in query_list:
run(query)
print('New rows have been added to auth_table and book_table!')
insert_b.on_click(insert_button)
# To be totally honest, you likely won't be writing nested queries yourself until you've become much more comfortable with SQL. However, it's good to learn about them because you'll likely encounter them when coworkers share queries with you.
#
# Start by looking at the two queries and their outputs below:
nested_q1 = widgets.Textarea(value=
'''SELECT
COUNT(DISTINCT(book)) AS Count_of_Distinct_Books
FROM
sales_table''',
width = '50em', height = '7em')
display(nested_q1)
nested_b1 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(nested_b1)
run_q(nested_q1, nested_b1)
# ** NOTE THAT YOU NEED TO HIT "RUN" AGAIN FOR THE QUERY BELOW**
# (It should return the number 13. If it doesn't, click the blue button above to update the `book_table`, then re-run the query below)
nested_q2 = widgets.Textarea(value=
'''SELECT
COUNT(DISTINCT(book)) AS Count_of_Distinct_Books
FROM
book_table''',
width = '50em', height = '7em')
display(nested_q2)
nested_b2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(nested_b2)
run_q(nested_q2, nested_b2)
# From the count of distinct books in each table, we see that there are two books in our `book_table` (our inventory) that haven't made a single sale. Imagine if both tables had thousands of rows - it'd be a nightmare to try to figure out which were the books with no sales. However, a nested query can help us out.
#
# The query below uses a nested query in the `WHERE` clause. In plain English, it says "*Show me the books from the book_table, but not the ones that also show up in the sales_table*":
nested_q3 = widgets.Textarea(value=
'''SELECT
book
FROM
book_table
WHERE
book NOT IN (SELECT book FROM sales_table)''',
width = '50em', height = '10em')
display(nested_q3)
nested_q3_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(nested_q3_b)
run_q(nested_q3, nested_q3_b)
# ____
# ### Challenge:
# Write a query to see authors who appear in the `auth_table` but don't show up in the `book_table`.
nest_chall = widgets.Textarea(value='', width = '50em', height = '12em')
display(nest_chall)
nest_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(nest_chall_b)
run_q(nest_chall, nest_chall_b)
nest_chall_c ='''SELECT
last_name
FROM
auth_table
WHERE
last_name NOT IN (SELECT author FROM book_table)
'''
cheat(nest_chall_c)
# You can also use nested queries to avoid the need for multiple `JOIN` clauses. Suppose you wanted to see the total revenue for books by authors from England. Previously, we would have joined the `sales_table` to the `book_table`, and then the `book_table` to the `auth_table` in order to be able to work with both the `revenue` column and the `country` column:
nest_q3 = widgets.Textarea(value=
'''SELECT
SUM(revenue)
FROM
sales_table
WHERE
book IN (SELECT
book
FROM
book_table B
JOIN auth_table A ON B.author = A.last_name
WHERE
A.country = 'England')''',
width = '50em', height = '18em')
display(nest_q3)
nest_b3 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(nest_b3)
run_q(nest_q3, nest_b3)
# Let's break down what's going on here. First, copy and paste the *nested* query (the part in parentheses) in the cell below, then run it:
nest_q3_explained = widgets.Textarea(value='',width = '50em', height = '12em')
display(nest_q3_explained)
nest_b3_explained = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(nest_b3_explained)
run_q(nest_q3_explained, nest_b3_explained)
# Next, take the output that you just produced and: <br>
# • comma-separate each book <br>
# • wrap each book in quotation marks <br>
# • paste your list between the parentheses in the `WHERE` clause below: <br>
# • rerun the query <br>
nest_q3_explained2 = widgets.Textarea(value=
'''SELECT
SUM(revenue)
FROM
sales_table
WHERE
book IN ( )''',
width = '50em', height = '12em')
display(nest_q3_explained2)
nest_b3_explained2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(nest_b3_explained2)
run_q(nest_q3_explained2, nest_b3_explained2)
# This is essentially the same process that SQL walks through when you run a nested query. It pulls the list of books from the nested query, then uses that list in the `WHERE` clause of the dominant query.
#
# <img align="left" src="http://i.imgur.com/sRHO5xr.png"> <br>
# The query above produces the same results as if you'd done a multiple join, but it's much more efficient. That's because SQL can just get the book titles it needs from the nested query and plug them into the dominent query, rather than needing to do all the work of duplicating rows to create joined tables. Use nested queries for more efficient joins whenever possible.
# __________
# ___________
# ___________
# <a id='union'></a>
# <center>
# [Previous](#case_when) | [Table of Contents](#table_of_contents) | [Next](#rollup)
# </center>
# 
# >**`SELECT`** <br>
# **`some_column`** <br>
# **`FROM`** <br>
# **`table_x`** <br>
#
# >**`UNION`** ➞ or use **`UNION ALL`**, see explanation below <br>
#
# >**`SELECT`** <br>
# **`some_other_column`** <br>
# **`FROM`** <br>
# **`table_y`** <br>
#
# **`UNION`** and **`UNION ALL`** allow you to attach two completely separate queries. **`UNION`** will result in the output from the first query and the second query to be sorted by default (or you can add an `ORDER BY` clause). **`UNION ALL`** will ensure that the results from the second query will all appear after the results from the first query.
#
# We'll start with a very, very simple illustration and work our way into more complex versions of `UNION` queries. First, consider the query below. We're pulling all books from the book table with the first query, and all the authors' first names from the auth_table with the second query. By using `UNION`, we're telling SQL to return the results of both these queries in the same column.
union_q = widgets.Textarea(value=
'''SELECT
book AS selection
FROM
book_table
UNION
SELECT
first_name AS selection
FROM
auth_table''',
width = '50em', height = '16em')
display(union_q)
union_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(union_b)
run_q(union_q, union_b)
# ### Quick Exercise:
# 1. Change the query above to use **`UNION ALL`** instead of **`UNION`** and re-run. Make sure you understand how the output changes.
# 2. Delete **`AS selection`** in the second query and rerun.
# 3. Rename **`selection`** to something else in the first query and rerun.
# 4. Add **`cogs`** in the **`SELECT`** clause in the first query and **`country`** to the **`SELECT`** clause in the second query, rerun.
# 5. Delete **`cogs`** in the first query and rerun. Can you think why you're hitting an error?
# ### Useful applications for `UNION`
# The above example is just a simple illustration of how `UNION` functions, but it's not very useful as a practical application. Now lets try `UNION` in a more useful way.
#
# Let's say that our imaginary book store decides to start stocking a few movies, so we've created a new table to manage this new inventory:
# 
#
# Now let's say we wanted to view *all* of our store inventory, COGs, and the author or director of each item. We don't want to join the `movie_table` and the `book_table` - there's nothing to really join them on. However, it'd be useful to stack them.
#
# ### Challenge:
# Use **`UNION`** to write a query to view the contents of both **`movie_table`** and **`book_table`** in a single table. The column-headers should be: **`Item`**, **`COGs`**, and **`Creator`**. Order by item title (hint: with **`UNION`**, the **`ORDER BY`** clause can only go after the second query).
union_chall = widgets.Textarea(value='',width = '50em', height = '25em')
display(union_chall)
union_chall_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(union_chall_b)
run_q(union_chall, union_chall_b)
union_chall_c ='''SELECT
book as Item,
cogs as COGs,
author as Creator
FROM
book_table
UNION
SELECT
film,
cogs,
director
FROM
movie_table
ORDER BY
Item'''
cheat(union_chall_c)
# ### Using `UNION` to add totals and subtotals:
# Take a look at the query below. You'll see it pulls the COGs and book title for each book. It also uses `UNION ALL` to add a final line - a summary row averaging all cogs:
union_q2 = widgets.Textarea(value=
'''SELECT
book,
cogs
FROM
book_table
UNION ALL
SELECT
'Average COGs',
ROUND(AVG(cogs), 2)
FROM
book_table''',
width = '50em', height = '19em')
display(union_q2)
union_b2 = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(union_b2)
run_q(union_q2, union_b2)
# ### Quick Exercise:
# 1. Remove the **`ALL`** from **`UNION ALL`** and rerun. See how **`ALL`** can be useful?
# 2. Delete **` 'Average COGs',`** from the second query and rerun. Make sure you understand why there's an error. Fix it and reruun.
#
# ### Challenge:
# Write a query that totals revenue per book from the **`sales_table`**. Use **`UNION ALL`** to add a summary line that totals revenue for all books.
union_chall1 = widgets.Textarea(value='',
width = '50em', height = '22em')
display(union_chall1)
union_chall1_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(union_chall1_b)
run_q(union_chall1, union_chall1_b)
union_chall1_c ='''SELECT
book,
SUM(revenue) AS total_revenue
FROM
sales_table
GROUP BY
book
UNION ALL
SELECT
'NULL',
SUM(revenue)
FROM
sales_table
'''
cheat(union_chall1_c)
# ### Extra Challenging Challenge:
# Write a query that totals revenue per book. Add subtotal lines for each author's revenue above their books. The output should look like this (use Google to figure out how to capitalize the authors' names for the subtotal rows).
HTML(run('''SELECT
B.author AS author_last_name,
B.book AS book_title,
SUM(S.revenue) AS sum_revenue
FROM
sales_table S
JOIN book_table B on S.book = B.book
GROUP BY
B.book
UNION
SELECT
UPPER(B.author),
'TOTAL REVENUE',
SUM(S.revenue)
FROM
sales_table S
JOIN book_table B on S.book = B.book
GROUP BY
B.author
''').to_html(index=False))
union_chall2 = widgets.Textarea(value='', width = '50em', height = '30em')
display(union_chall2)
union_chall2_b = widgets.Button(description='Run', width='10em', height='2.5em', color='white',background_color='black', border_color='black')
display(union_chall2_b)
run_q(union_chall2, union_chall2_b)
union_chall2_c ='''SELECT
B.author AS author_last_name,
B.book AS book_title,
SUM(S.revenue) AS sum_revenue
FROM
sales_table S
JOIN book_table B on S.book = B.book
GROUP BY
B.book
UNION
SELECT
UPPER(B.author),
'TOTAL REVENUE',
SUM(S.revenue)
FROM
sales_table S
JOIN book_table B on S.book = B.book
GROUP BY
B.author'''
cheat(union_chall2_c)
# __________
# ___________
# ___________
# <a id='rollup'></a>
# <center>
# [Previous](#union) | [Table of Contents](#table_of_contents) | [Next](#wrapping_up)
# </center>
# 
#
# <font color='#1f5fd6'>Microsoft SQL Server | <font color='#1f5fd6'>MySQL | <font color='#1f5fd6'>Oracle | <font color='#1f5fd6'>SQLite </font>
# :------------------: | :---: | :----: | :----:
# `GROUP BY column_a WITH ROLLUP` | `GROUP BY column_a WITH ROLLUP` | `GROUP BY ROLLUP (column_a)` | not supported
#
# Unfortunately, SQLite doesn't have a simple way to do ROLLUP like the other RDBMSs, so we can't practice it here. However, the concept is very straightforward: it's exactly like using `UNION` to add a summary row, except way simpler. Below is what the query *would* look like if we were using Microsoft or MySQL. Take a look at the query and the output to understand what's going on, even if you can't practice it:
#
# > **`SELECT` <br>
# `book,` <br>
# `SUM(revenue) AS total_revenue,` <br>
# `COUNT(*) AS count_of_sales` <br>
# `FROM` <br>
# `sales_table` <br>
# `GROUP BY` <br>
# `book WITH ROLLUP`** <br>
HTML(run('''SELECT
book,
SUM(revenue) AS total_revenue,
COUNT(*) AS count_of_sales
FROM
sales_table
GROUP BY
book
UNION ALL
SELECT
'NULL',
SUM(revenue),
COUNT(*)
FROM
sales_table''').to_html(index=False))
# Note that rollup produces the word "NULL" for any row that it cannot sum up. Essentially its function is to find all columns with numbers and add these up. Numberless columns can't be added, so rollup just skips them.
# __________
# ___________
# ___________
# <a id='wrapping_up'></a>
# <center>
# [Previous](#rollup) | [Table of Contents](#table_of_contents)
# </center>
# 
#
#
# ### One or two days before your job or internship:
# Review and PRACTICE! Review the lessons and terms, re-do the quick exercises and challenges. Seriously. This stuff is easily forgotten if you don't use it, so be sure to refresh what you've learned before you start working.
#
# Here's some [additional reading](http://www.w3schools.com/sql/sql_intro.asp) on SQL if you're interested.
#
# ### When you first start work:
# 1. Find out what relational database management system your company uses, and get acquainted with how that system differs from what we've learned in class (use the table below as a quide).
# 2. Ask coworkers if they have any pre-written queries that will be useful to your work. Read through them and make sure you understand them.
# 4. Read the structure of any table that seems important so you understand what data can be found in each.
# 5. Figure out how tables join to one another, and which columns come from which tables
# 6. ALWAYS ALWAYS avoid "slow server" traps when you are exploring your database. That means:
# - NEVER Run a simple **`SELECT * FROM table_name`** query unless you are absolutely certain that the table is very, very small
# - Avoid joining 3 or more tables whenever possible. If you find yourself needing them, try to see if you can use a nested query to cut one of the **`JOIN`** clauses out.
# - If your table is recording dates, use these to limit how much data you pull. Depending on the table, more than 1-2 months at a time will usually slow down a system.
# - Use **`LIKE`** and % sparingly or use **`WHERE`** to limit your search as much as possible unless you are dealing with a small-ish table
#
# ### What do I mean by small vs. large tables?
# Several times in class we've discussed certain practices that should only be applied to small tables to avoid strain on your server. You can use `SELECT COUNT(*) FROM table_name` to see how many rows it has. You can also think of it like this: the more often the table is updated, the larger it probably is. A table that adds a row every time a user views a webpage is updated constantly and is probably huge. A table that simply lists all the ZIP codes in the US probably doesn't get updated often, it'll be pretty small.
# <a id='dialect_differences'></a>
# # Dialect Differences:
# <font color='#1f5fd6'> Description | <font color='#1f5fd6'> Microsoft SQL Server | <font color='#1f5fd6'> MySQL | <font color='#1f5fd6'> Oracle | <font color='#1f5fd6'> SQLite </font>|
# :--------: | :------------------: | :---: | :----: | :----: |
# **Reading a table's structure** | `SP_Help tablename` | `DESCRIBE tablename` | `DESCRIBE tablename` | `PRAGMA TABLE_INFO(tablename)`
# **Limiting rows** | `SELECT TOP N column_name` | `LIMIT N` | `WHERE ROWNUM <= N` | `LIMIT N`
# **`JOIN` or `INNER JOIN`** | ✓ | ✓ | ✓ | ✓
# **`LEFT JOIN` or `LEFT OUTER JOIN`** | ✓ | ✓ | ✓ | ✓
# **`RIGHT JOIN` or `RIGHT OUTER JOIN`** | ✓ | ✓ | ✓ | not supported
# **`OUTER JOIN` or `FULL OUTER JOIN`** | ✓ | not supported | ✓ | not supported
# **`IF`** | `IF logical_test PRINT value_if_true ` | `IF(logical_test, value_if_true, value_if_false)` (same as Excel) | `IF logical_test THEN value_if_true ELSIF...END IF` | NOT SUPPORTED
# **`CASE WHEN`** | ✓ | ✓ | ✓ | ✓
# **`ROLLUP`** | `GROUP BY column_a WITH ROLLUP` | `GROUP BY column_a WITH ROLLUP` | `GROUP BY ROLLUP (column_a)` | not supported
|
notebooks/SQL_Bootcamp_Stern_2016.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="8F4l1VYwjCyU" colab_type="code" colab={}
import numpy as np
import pandas as pd
import os
import random
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from sklearn.dummy import DummyRegressor
from sklearn.metrics import r2_score
# + id="8il-gBGgjqE8" colab_type="code" outputId="432ce253-a928-41e0-d36e-beda2c3c8d29" executionInfo={"status": "ok", "timestamp": 1582455021679, "user_tz": 300, "elapsed": 32454, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 139}
from google.colab import drive
drive.mount('../content/drive')
work_dir = '../content/drive/Shared drives/brown datathon'
print(os.listdir(work_dir))
# + id="h7eHMI-d8XA-" colab_type="code" outputId="d779e9ca-88b0-407e-a89f-bbe1578cdb82" executionInfo={"status": "error", "timestamp": 1582455235173, "user_tz": 300, "elapsed": 915, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 232}
# Predict 1a
for file in os.listdir(work_dir + "/1/Testing")[:-1]:
df_temp = pd.read_csv(work_dir + f"/1/Testing/{file}").drop(["Unnamed: 4"], axis=1)
df_temp["power"] = float(file[-8:-5])
df_temp["speed"] = float(file[0] + "." + file[2:4])
y_pred_temp = model_1a.predict(df_temp)
df_temp["Temp"] = y_pred_temp
df_temp.drop(["power", "speed"], axis=1, inplace=True)
df_temp#.to_csv(work_dir + f"\\1\\Testing\\{file}", index=False)
# + id="EtU1SFYRlDhH" colab_type="code" outputId="0c64b82f-9346-4845-dd3d-7858dfbfd49f" executionInfo={"status": "ok", "timestamp": 1582410360627, "user_tz": 300, "elapsed": 3311, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
laser_speeds = np.linspace(0.6, 1.5, 19)
laser_powers = np.linspace(100, 400, 13)
train_df = pd.DataFrame()
test_df = pd.DataFrame()
for speed in [round(s, 2) for s in laser_speeds]:
flag=True
if np.random.rand()>0.8:
flag=False
for power in [int(p) for p in laser_powers]:
flag=True
if np.random.rand()>0.8:
flag=False
file_name = (str(speed).replace('.', '_')
+ 'ms_'
+ str(power)
+ 'W.csv')
try:
file_df = pd.read_csv(os.path.join(work_dir, '1/Training', file_name), index_col = False)
file_df = file_df.assign(speed = lambda x: speed, power = lambda x: power)
if flag:
train_df = pd.concat([train_df, file_df])
else:
test_df = pd.concat([test_df, file_df])
print(file_name)
except:
pass
#file_df = pd.read_csv(os.path.join(work_dir, '1/Validation', file_name), index_col = False)
print(train_df.shape)
print(test_df.shape)
# + id="z15ni8zg_Hqn" colab_type="code" colab={}
train_df.to_csv(work_dir+'/1/Training/temp_train.csv')
test_df.to_csv(work_dir+'/1/Training/temp_test.csv')
# + id="EmMA8CFOl_ES" colab_type="code" colab={}
#full_df_temp.to_csv(work_dir+'/1/Validation/full_temperature_cv.csv')
# + id="i0LwpkvfrsHS" colab_type="code" colab={}
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
# + id="wZx6dyP0tOkT" colab_type="code" colab={}
train_X, test_X, train_y, test_y = train_test_split(full_df_temp.drop('Temp',axis=1), full_df_temp['Temp'], test_size=0.33)
# + id="nG5NxEzpttRy" colab_type="code" outputId="30e66d58-d02b-41b6-fbbb-44809061da58" executionInfo={"status": "ok", "timestamp": 1582405618091, "user_tz": 300, "elapsed": 363, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
reg = LinearRegression()
reg.fit(train_X, train_y)
reg.score(test_X, test_y)
# + id="bFCmhX2auAnA" colab_type="code" colab={}
train_X1 = train_X.assign(distance = lambda x: np.sqrt(x.X_Coord**2+x.Y_Coord**2+x.Z_Coord**2))
test_X1 = test_X.assign(distance = lambda x: np.sqrt(x.X_Coord**2+x.Y_Coord**2+x.Z_Coord**2))
# + id="1WyM-9_MuzRe" colab_type="code" outputId="90c6e946-898f-4179-e587-1936b075e1f6" executionInfo={"status": "ok", "timestamp": 1582405919725, "user_tz": 300, "elapsed": 463, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
reg = LinearRegression()
reg.fit(train_X1, train_y)
reg.score(test_X1, test_y)
# + id="kUzpUfggviG9" colab_type="code" colab={}
train_X1 = train_X1.assign(distance_05 = lambda x: x.distance**0.5, distance_2 = lambda x: x.distance**2)
test_X1 = test_X1.assign(distance_05 = lambda x: x.distance**0.5, distance_2 = lambda x: x.distance**2)
# + id="MsgzenbowyOJ" colab_type="code" outputId="6b752aff-a74a-4e87-b2d1-b7d24e5f903c" executionInfo={"status": "ok", "timestamp": 1582406253475, "user_tz": 300, "elapsed": 673, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
reg = LinearRegression()
reg.fit(train_X1, train_y)
reg.score(test_X1, test_y)
# + id="UP7fbYqIwzjC" colab_type="code" outputId="ced3a272-2c15-4488-dc03-05512326405a" executionInfo={"status": "ok", "timestamp": 1582406449596, "user_tz": 300, "elapsed": 2343, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 282}
y_pred = reg.predict(test_X1)
import matplotlib.pyplot as plt
plt.scatter(test_y, y_pred)
# + id="baV1lQPfxc1W" colab_type="code" outputId="4e2b2cb2-4960-4c17-cecf-f282b0c529dc" executionInfo={"status": "error", "timestamp": 1582407463943, "user_tz": 300, "elapsed": 35787, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 375}
from sklearn.ensemble import RandomForestRegressor
reg = RandomForestRegressor(n_jobs=-1)
reg.fit(train_X1, train_y)
reg.score(test_X1, test_y)
#y_pred = reg.predict(test_X1)
#plt.scatter(test_y, y_pred)
# + id="4RmqC7n3yMVe" colab_type="code" outputId="01c76f1f-383c-4106-d1a3-3b0cbb75ef3f" executionInfo={"status": "ok", "timestamp": 1582407230269, "user_tz": 300, "elapsed": 6324, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12439108135450337217"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
print(reg.score(test_X1, test_y))
# + id="xBElqLg00goo" colab_type="code" colab={}
|
src/MergingData_TempPrediction .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><img src="../images/DLI Header.png" alt="Header" style="width: 400px;"/></center>
# # Getting Started with AI on Jetson Nano
# ### Interactive Classification Tool
# This notebook is an interactive data collection, training, and testing tool, provided as part of the NVIDIA Deep Learning Institute (DLI) course, "Getting Started with AI on Jetson Nano". It is designed to be run on the Jetson Nano in conjunction with the detailed instructions provided in the online DLI course pages.
#
# To start the tool, set the **Camera** and **Task** code cell definitions, then execute all cells. The interactive tool widgets at the bottom of the notebook will display. The tool can then be used to gather data, add data, train data, and test data in an iterative and interactive fashion!
#
# The explanations in this notebook are intentionally minimal to provide a streamlined experience. Please see the DLI course pages for detailed information on tool operation and project creation.
# ### Camera
# First, create your camera and set it to `running`. Uncomment the appropriate camera selection lines, depending on which type of camera you're using (USB or CSI). This cell may take several seconds to execute.
# <div style="border:2px solid black; background-color:#e3ffb3; font-size:12px; padding:8px; margin-top: auto;"><i>
# <h4><i>Tip</i></h4>
# <p>There can only be one instance of CSICamera or USBCamera at a time. Before starting a new project and creating a new camera instance, you must first release the existing one. To do so, shut down the notebook's kernel from the JupyterLab pull-down menu: <strong>Kernel->Shutdown Kernel</strong>, then restart it with <strong>Kernel->Restart Kernel</strong>.</p>
# <ul><code>sudo systemctl restart nvargus-daemon</code> with password:<code><PASSWORD></code> is included to then force a reset of the camera daemon.</ul>
# +
# Full reset of the camera
# !echo 'dlinano' | sudo -S systemctl restart nvargus-daemon && printf '\n'
# Check device number
# !ls -ltrh /dev/video*
# USB Camera (Logitech C270 webcam)
# from jetcam.usb_camera import USBCamera
# camera = USBCamera(width=224, height=224, capture_device=0) # confirm the capture_device number
# CSI Camera (Raspberry Pi Camera Module V2)
from jetcam.csi_camera import CSICamera
camera = CSICamera(width=224, height=224)
camera.running = True
print("camera created")
# -
# ### Task
# Next, define your project `TASK` and what `CATEGORIES` of data you will collect. You may optionally define space for multiple `DATASETS` with names of your choosing.
# Uncomment/edit the associated lines for the classification task you're building and execute the cell.
# This cell should only take a few seconds to execute.
# +
import torchvision.transforms as transforms
from dataset import ImageClassificationDataset
TASK = 'thumbs'
# TASK = 'emotions'
# TASK = 'fingers'
# TASK = 'diy'
CATEGORIES = ['thumbs_up', 'thumbs_down']
# CATEGORIES = ['none', 'happy', 'sad', 'angry']
# CATEGORIES = ['1', '2', '3', '4', '5']
# CATEGORIES = [ 'diy_1', 'diy_2', 'diy_3']
DATASETS = ['A', 'B']
# DATASETS = ['A', 'B', 'C']
TRANSFORMS = transforms.Compose([
transforms.ColorJitter(0.2, 0.2, 0.2, 0.2),
transforms.Resize((224, 224)),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
datasets = {}
for name in DATASETS:
datasets[name] = ImageClassificationDataset(TASK + '_' + name, CATEGORIES, TRANSFORMS)
print("{} task with {} categories defined".format(TASK, CATEGORIES))
# -
# ### Data Collection
# Execute the cell below to create the data collection tool widget. This cell should only take a few seconds to execute.
# +
import ipywidgets
import traitlets
from IPython.display import display
from jetcam.utils import bgr8_to_jpeg
# initialize active dataset
dataset = datasets[DATASETS[0]]
# unobserve all callbacks from camera in case we are running this cell for second time
camera.unobserve_all()
# create image preview
camera_widget = ipywidgets.Image()
traitlets.dlink((camera, 'value'), (camera_widget, 'value'), transform=bgr8_to_jpeg)
# create widgets
dataset_widget = ipywidgets.Dropdown(options=DATASETS, description='dataset')
category_widget = ipywidgets.Dropdown(options=dataset.categories, description='category')
count_widget = ipywidgets.IntText(description='count')
save_widget = ipywidgets.Button(description='add')
# manually update counts at initialization
count_widget.value = dataset.get_count(category_widget.value)
# sets the active dataset
def set_dataset(change):
global dataset
dataset = datasets[change['new']]
count_widget.value = dataset.get_count(category_widget.value)
dataset_widget.observe(set_dataset, names='value')
# update counts when we select a new category
def update_counts(change):
count_widget.value = dataset.get_count(change['new'])
category_widget.observe(update_counts, names='value')
# save image for category and update counts
def save(c):
dataset.save_entry(camera.value, category_widget.value)
count_widget.value = dataset.get_count(category_widget.value)
save_widget.on_click(save)
data_collection_widget = ipywidgets.VBox([
ipywidgets.HBox([camera_widget]), dataset_widget, category_widget, count_widget, save_widget
])
# display(data_collection_widget)
print("data_collection_widget created")
# -
# ### Model
# Execute the following cell to define the neural network and adjust the fully connected layer (`fc`) to match the outputs required for the project. This cell may take several seconds to execute.
# +
import torch
import torchvision
device = torch.device('cuda')
# ALEXNET
# model = torchvision.models.alexnet(pretrained=True)
# model.classifier[-1] = torch.nn.Linear(4096, len(dataset.categories))
# SQUEEZENET
# model = torchvision.models.squeezenet1_1(pretrained=True)
# model.classifier[1] = torch.nn.Conv2d(512, len(dataset.categories), kernel_size=1)
# model.num_classes = len(dataset.categories)
# RESNET 18
model = torchvision.models.resnet18(pretrained=True)
model.fc = torch.nn.Linear(512, len(dataset.categories))
# RESNET 34
# model = torchvision.models.resnet34(pretrained=True)
# model.fc = torch.nn.Linear(512, len(dataset.categories))
model = model.to(device)
model_save_button = ipywidgets.Button(description='save model')
model_load_button = ipywidgets.Button(description='load model')
model_path_widget = ipywidgets.Text(description='model path', value='my_model.pth')
def load_model(c):
model.load_state_dict(torch.load(model_path_widget.value))
model_load_button.on_click(load_model)
def save_model(c):
torch.save(model.state_dict(), model_path_widget.value)
model_save_button.on_click(save_model)
model_widget = ipywidgets.VBox([
model_path_widget,
ipywidgets.HBox([model_load_button, model_save_button])
])
# display(model_widget)
print("model configured and model_widget created")
# -
# ### Live Execution
# Execute the cell below to set up the live execution widget. This cell should only take a few seconds to execute.
# +
import threading
import time
from utils import preprocess
import torch.nn.functional as F
state_widget = ipywidgets.ToggleButtons(options=['stop', 'live'], description='state', value='stop')
prediction_widget = ipywidgets.Text(description='prediction')
score_widgets = []
for category in dataset.categories:
score_widget = ipywidgets.FloatSlider(min=0.0, max=1.0, description=category, orientation='vertical')
score_widgets.append(score_widget)
def live(state_widget, model, camera, prediction_widget, score_widget):
global dataset
while state_widget.value == 'live':
image = camera.value
preprocessed = preprocess(image)
output = model(preprocessed)
output = F.softmax(output, dim=1).detach().cpu().numpy().flatten()
category_index = output.argmax()
prediction_widget.value = dataset.categories[category_index]
for i, score in enumerate(list(output)):
score_widgets[i].value = score
def start_live(change):
if change['new'] == 'live':
execute_thread = threading.Thread(target=live, args=(state_widget, model, camera, prediction_widget, score_widget))
execute_thread.start()
state_widget.observe(start_live, names='value')
live_execution_widget = ipywidgets.VBox([
ipywidgets.HBox(score_widgets),
prediction_widget,
state_widget
])
# display(live_execution_widget)
print("live_execution_widget created")
# -
# ### Training and Evaluation
# Execute the following cell to define the trainer, and the widget to control it. This cell may take several seconds to execute.
# +
BATCH_SIZE = 8
optimizer = torch.optim.Adam(model.parameters())
# optimizer = torch.optim.SGD(model.parameters(), lr=1e-3, momentum=0.9)
epochs_widget = ipywidgets.IntText(description='epochs', value=1)
eval_button = ipywidgets.Button(description='evaluate')
train_button = ipywidgets.Button(description='train')
loss_widget = ipywidgets.FloatText(description='loss')
accuracy_widget = ipywidgets.FloatText(description='accuracy')
progress_widget = ipywidgets.FloatProgress(min=0.0, max=1.0, description='progress')
def train_eval(is_training):
global BATCH_SIZE, LEARNING_RATE, MOMENTUM, model, dataset, optimizer, eval_button, train_button, accuracy_widget, loss_widget, progress_widget, state_widget
try:
train_loader = torch.utils.data.DataLoader(
dataset,
batch_size=BATCH_SIZE,
shuffle=True
)
state_widget.value = 'stop'
train_button.disabled = True
eval_button.disabled = True
time.sleep(1)
if is_training:
model = model.train()
else:
model = model.eval()
while epochs_widget.value > 0:
i = 0
sum_loss = 0.0
error_count = 0.0
for images, labels in iter(train_loader):
# send data to device
images = images.to(device)
labels = labels.to(device)
if is_training:
# zero gradients of parameters
optimizer.zero_grad()
# execute model to get outputs
outputs = model(images)
# compute loss
loss = F.cross_entropy(outputs, labels)
if is_training:
# run backpropogation to accumulate gradients
loss.backward()
# step optimizer to adjust parameters
optimizer.step()
# increment progress
error_count += len(torch.nonzero(outputs.argmax(1) - labels).flatten())
count = len(labels.flatten())
i += count
sum_loss += float(loss)
progress_widget.value = i / len(dataset)
loss_widget.value = sum_loss / i
accuracy_widget.value = 1.0 - error_count / i
if is_training:
epochs_widget.value = epochs_widget.value - 1
else:
break
except e:
pass
model = model.eval()
train_button.disabled = False
eval_button.disabled = False
state_widget.value = 'live'
train_button.on_click(lambda c: train_eval(is_training=True))
eval_button.on_click(lambda c: train_eval(is_training=False))
train_eval_widget = ipywidgets.VBox([
epochs_widget,
progress_widget,
loss_widget,
accuracy_widget,
ipywidgets.HBox([train_button, eval_button])
])
# display(train_eval_widget)
print("trainer configured and train_eval_widget created")
# -
# ### Display the Interactive Tool!
# The interactive tool includes widgets for data collection, training, and testing.
# <center><img src="../images/classification_tool_key2.png" alt="tool key" width=500/></center>
# <br>
# <center><img src="../images/classification_tool_key1.png" alt="tool key"/></center>
# Execute the cell below to create and display the full interactive widget. Follow the instructions in the online DLI course pages to build your project.
# +
# Combine all the widgets into one display
all_widget = ipywidgets.VBox([
ipywidgets.HBox([data_collection_widget, live_execution_widget]),
train_eval_widget,
model_widget
])
display(all_widget)
# -
# <center><img src="../images/DLI Header.png" alt="Header" style="width: 400px;"/></center>
|
nvdli-nano/classification/classification_interactive.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # 11 ODE Applications (Projectile motion) – Part 1
# Let's apply our ODE solvers to some problems involving balls and projectiles.
#
# The `integrators.py` file from [Lesson 10](http://asu-compmethodsphysics-phy494.github.io/ASU-PHY494//2018/02/20/10_ODEs/) is used here (and named [`ode.py`](https://github.com/ASU-CompMethodsPhysics-PHY494/PHY494-resources/tree/master/11_ODE_applications/ode.py)).
#
# *Note: Incomplete notebook for students to work on*
#
# ## Contents
#
# 1. Projectile with linear air-resistance (theory and full code)
# 2. Baseball physics (theory and skeleton code)
import numpy as np
import ode
# %matplotlib inline
import matplotlib.pyplot as plt
plt.matplotlib.style.use('ggplot')
# ## Projectile with linear air-resistance
# Linear drag force
#
# $$
# \mathbf{F}_1 = -b_1 \mathbf{v}
# $$
#
# Equations of motion with force due to gravity $\mathbf{g} = -g \hat{\mathbf{e}}_y$
#
# \begin{align}
# \frac{d\mathbf{r}}{dt} &= \mathbf{v}\\
# \frac{d\mathbf{v}}{dt} &= - g \hat{\mathbf{e}}_y -\frac{b_1}{m} \mathbf{v}
# \end{align}
# Bring into standard ODE form for
#
# $$
# \frac{d\mathbf{y}}{dt} = \mathbf{f}(t, \mathbf{y})
# $$
# as
#
# $$
# \mathbf{y} = \begin{pmatrix}
# x\\
# y\\
# v_x\\
# v_y
# \end{pmatrix}, \quad
# \mathbf{f} = \begin{pmatrix}
# v_x\\
# v_y\\
# -\frac{b_1}{m} v_x\\
# -g -\frac{b_1}{m} v_y
# \end{pmatrix}
# $$
# (Based on Wang 2016, Ch 3.3.1)
# +
def simulate(v0, h=0.01, b1=0.2, g=9.81, m=0.5):
def f(t, y):
# y = [x, y, vx, vy]
return np.array([y[2], y[3], -b1/m * y[2], -g - b1/m * y[3]])
vx, vy = v0
t = 0
positions = []
y = np.array([0, 0, vx, vy], dtype=np.float64)
while y[1] >= 0:
positions.append([t, y[0], y[1]]) # record t, x and y
y[:] = ode.rk4(y, f, t, h)
t += h
return np.array(positions)
def initial_v(v, theta):
x = np.deg2rad(theta)
return v * np.array([np.cos(x), np.sin(x)])
# -
r = simulate(initial_v(200, 30), h=0.01, b1=1)
plt.plot(r[:, 1], r[:, 2])
plt.xlabel(r"distance $x$ (m)")
plt.ylabel(r"height $y$ (m)");
for angle in (5, 7.5, 10, 20, 30, 45):
r = simulate(initial_v(200, angle), h=0.01, b1=1)
plt.plot(r[:, 1], r[:, 2], label=r"$\theta = {}^\circ$".format(angle))
plt.legend(loc="best")
plt.xlabel(r"distance $x$ (m)")
plt.ylabel(r"height $y$ (m)");
# ## Simple Baseball physics
#
# - quadratic air resistance (with velocity-dependent drag coefficient)
# - Magnus force due to spin
#
# ### Quadratic air resistance
# Occurs at high Reynolds numbers, i.e., turbulent flow. Only approximate:
#
# $$
# \mathbf{F}_2 = -b_2 v \mathbf{v}
# $$
# ### Magnus effect
# **Magnus effect**: airflow is changed around a spinning object. The Magnus force is
#
# $$
# \mathbf{F}_M = \alpha \boldsymbol{\omega} \times \mathbf{v}
# $$
#
# where $\boldsymbol{\omega}$ is the ball's angular velocity in rad/s (e.g., 200/s for a baseball).
#
# For a sphere the proportionality constant $\alpha$ can be written
#
# $$
# \mathbf{F}_M = \frac{1}{2} C_L \rho A \frac{v}{\omega} \boldsymbol{\omega} \times \mathbf{v}
# $$
#
# where $C_L$ is the lift coefficient, $\rho$ the air density, $A$ the ball's cross section. (Advantage of defining $C_L$ this way: when spin and velocity are perpendicular, the Magnus force is simply $F_M = \frac{1}{2} C_L \rho A v^2$.)
# $C_L$ is mainly a function of the *spin parameter*
#
# $$
# S = \frac{r\omega}{v}
# $$
#
# with the radius $r$ of the ball. In general we write
#
# $$
# \mathbf{F}_M = \frac{1}{2} C_L \frac{\rho A r}{S} \boldsymbol{\omega} \times \mathbf{v}
# $$
# For a baseball, experimental data show approximately a power law dependence of $C_L$ on $S$
#
# $$
# C_L = 0.62 \times S^{0.7}
# $$
# All together:
#
# \begin{align}
# \mathbf{F}_M &= \alpha\ \boldsymbol{\omega} \times \mathbf{v}\\
# v &= \sqrt{\mathbf{v}\cdot\mathbf{v}}\\
# S &= \frac{r\omega}{v}\\
# C_L &= 0.62 \times S^{0.7}\\
# \alpha &= \frac{1}{2} C_L \frac{\rho A r}{S}
# \end{align}
#
# ### Equations of motion
#
# \begin{align}
# \frac{d\mathbf{r}}{dt} &= \mathbf{v}\\
# \frac{d\mathbf{v}}{dt} &= -g \hat{\mathbf{e}}_y \mathbf{v} -\frac{b_2}{m} v \mathbf{v} + \alpha\ \boldsymbol{\omega} \times \mathbf{v}\\
# \end{align}
#
# (quadratic drag $-\frac{b_2}{m} v \mathbf{v}$ included.)
#
# ### Baseball simulation
#
# Implement the full baseball equations of motions:
# - gravity $a_\text{gravity}$
# - quadratic drag $a_\text{drag}$
# - Magnus effect $a_\text{Magnus}$
#
# For the cross product you can look at [numpy.cross()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.cross.html).
# We will live-code the baseball simulation in class (i.e., build it from scratch), but if you want to work on this problem on your own and need some starter code, see below.
# #### Baseball skeleton code
# (Incomplete, full solution will be posted as `baseball_solution.ipynb`.)
# +
def C_L(S):
return 0.62 * S**0.7
def simulate_baseball(v0, omega=200.*np.array([0,1,1]), r0=np.array([0, 2.]),
h=0.01, b2=0.0013, g=9.81, rho = 1.225,
r=0.07468/2, m=0.14883, R_homeplate=18.4):
# make sure that omega is a numpy array
omega = np.asarray(omega)
# all SI units (kg, m)
# air density rho in kg/m^3
domega = np.linalg.norm(omega)
A = np.pi*r**2
rhoArm = rho * A * r / m
# internally, use 3d coordinates [x,y,z];
# y = [x, y, z, vx, vy, vz]
a_gravity = np.array([0, -g, 0])
def f(t, y):
# y = [x, y, z, vx, vy, vz]
v = y[3:]
dv = np.linalg.norm(v)
# COMPLETE
# 1. acceleration due to drag
# 2. acceleration due to Magnus effect
# 3. acceleration due to gravity (a_gravity)
# need to return array f of length 6!
raise NotImplementedError
x0, y0 = r0
vx, vy = v0
t = 0
positions = []
# initialize 3D!
y = np.array([x0, y0, 0, vx, vy, 0], dtype=np.float64)
# IMPLEMENT integration loop
# - use ode.rk4()
# - stop when x >= R_homeplate or y < 0.2 (i.e. cannot be caught)
return np.array(positions)
# -
# #### Simulate throws
# Simulate baseball throw for initial velocity $\mathbf{v} = (30\,\text{m/s}, 0)$.
#
# Plot x vs y and x vs z (to see curving).
#
# Try out different spins; a good value is $\boldsymbol{\omega} = 200\,\text{rad/s} \times (0, 1, 1)$.
# Simulate the baseball throw with
# - almost no spin: $\omega = 0.001 \times (0, 0, 1)$ (our code does not handle $\omega = 0$ gracefully...)
# - full upward spin: $\omega = 200 \times (0, 0, 1)$
# - sideways spin: $\omega = 200 \times (0, 1, 1)$
r = simulate_baseball([30, 0], omega=0.001*np.array([0,0,1]))
rz = simulate_baseball([30, 0], omega=200.*np.array([0,0,1]))
rzy = simulate_baseball([30, 0], omega=200.*np.array([0,1,1]))
# #### Plotting
#
# Plot the three scenarios in 2D planes: x-y (side view) and x-z (top view).
# #### 3D plot
# Use simple `matplotlib` 3D plot. (BONUS: Make it work with vpython)
# If we use the [`notebook` backend for matplotlib](http://ipython.readthedocs.io/en/stable/interactive/plotting.html) then we will be able to interactively rotate our [matplotlib 3D graphics](http://matplotlib.org/mpl_toolkits/mplot3d/tutorial.html). (Note: If this does not seem to work, disable adblockers and allow javascript on the page.)
# %matplotlib notebook
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot(r[:,1], r[:,3], r[:,2], 'o', label="no spin")
# ...
# hand of the catcher, 0.2m above homeplate
ax.plot([18.4, 18.4], [0, 0], [0, 0.2], color="black", lw=6)
ax.set_xlabel("$x$ (m)")
ax.set_ylabel("$z$ (m)")
ax.set_zlabel("$y$ (m)")
ax.legend(loc="upper left", numpoints=1)
ax.figure.tight_layout()
# -
# ## Reynolds number
#
# $$
# \text{Re} = \frac{\rho v L}{\mu}\\
# \text{Re} > 2300\quad\text{flow turbulent}
# $$
#
# * density $\rho$: air 1.275 kg/m^3 (kilograms per cubic meter)
# * fluid viscosity $\mu$: air 1.845×10^-5 Pa s (pascal seconds) (at 25 °C)
#
# (from Wolfram Alpha)
rho_air = 1.275 # kg/m^3
mu_air = 1.845e-5 # Pa s
L = 0.05 # m
v = 200 # m/s
def ReynoldsNumber(v, L, rho=rho_air, mu=mu_air):
return rho*v*L/mu
ReynoldsNumber(v, L)
# This means that we really should be using quadratic air resistance for the projectile.
|
11_ODE_applications/11-ODE-applications.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Importações
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import RepeatedKFold, StratifiedKFold, cross_val_score
from yellowbrick.model_selection import LearningCurve, ValidationCurve
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.linear_model import SGDClassifier
from sklearn.ensemble import RandomForestClassifier
from imblearn.over_sampling import SMOTE
from collections import Counter
import warnings
warnings.filterwarnings("ignore")
print("Seleção de modelo iniciada")
# +
def load_data(file):
"""
Função para carregamento de arquivo em qualquer diretório.
Parametro:
file: Nome do arquivo , string, a ser carregado.
"""
path = input('Por favor adicione o diretório de trabalho?')
for dirname, _, filename in os.walk(path, topdown = True):
for filename in filename:
if filename == file:
treino = pd.read_csv(os.path.join(dirname,filename), header = None)
else:
pass
print("Carregamento finalizado!!!")
return treino
X_treino = load_data('X_treino.csv')
y_treino = load_data('y_treino.csv')
X_teste = load_data('X_teste.csv')
y_teste = load_data('y_teste.csv')
# +
# Preparando a lista de modelos
modelos = []
modelos.append(('LR', LogisticRegression()))
modelos.append(('LDA', LinearDiscriminantAnalysis()))
modelos.append(('CART', DecisionTreeClassifier()))
modelos.append(('RF', RandomForestClassifier()))
modelos.append(('SGD', SGDClassifier()))
def model_selection(models, x_treino, y_treino,x_teste,y_teste, num_folds = 10):
resultados = []
nomes = []
resultados_teste = []
nomes_teste = []
print('Dados de treino...')
for nome, modelo in models:
kfold = KFold(n_splits = num_folds, random_state = 42)
cv_results = cross_val_score(modelo, x_treino, y_treino, cv = kfold, scoring = 'roc_auc')
resultados.append(cv_results)
nomes.append(nome)
print(f'{nome}- ROCAUC: {cv_results.mean()} Std: {cv_results.std()}')
print('\n')
print('Dados de teste...')
for nome, modelo in models:
kfold = KFold(n_splits = num_folds, random_state = 24)
cv_results_teste = cross_val_score(modelo, x_teste, y_teste, cv = kfold, scoring = 'roc_auc')
resultados_teste.append(cv_results_teste)
nomes_teste.append(nome)
print(f'{nome}- ROCAUC: {cv_results_teste.mean()} Std: {cv_results_teste.std()}')
return modelos
# -
"""
Temos somente o RF acima de 0.90!
Assim como com LR, LDA e SGD como pequenas quedas de performance.
Já que o SGD esta com o menor desvio padrão seguirei com RF, SGD para a etapa de modelagem
"""
model_selection(modelos, X_treino, y_treino, X_teste, y_teste)
# +
models = [modelos[3][1], modelos[4][1]]
def learning_curves(models, X, y):
cv_strategy = StratifiedKFold(n_splits=3)
for model in models:
sizes = np.linspace(0.3,1.0,10)
viz = LearningCurve(model, cv=cv_strategy, scoring='roc_auc', train_sizes=sizes, n_jobs=4)
viz.fit(X, y)
viz.show()
learning_curves(models, X_treino, y_treino)
# -
|
notebooks/03_ModelSelection/20200101_ModelSelection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Functions
# +
def my_first_function():
print('Hello world!')
print('type: {}'.format(my_first_function))
my_first_function() # Calling a function
# -
# ### Arguments
# +
def greeting(arg1, arg2):
print('Hello {} and {}!'.format(arg1, arg2))
greeting('Ragnar', 'Uhtred')
# +
# Function with return value
def strip_and_lowercase(original):
modified = original.strip().lower()
return modified
uggly_string = ' MixED CaSe '
pretty = strip_and_lowercase(uggly_string)
print('pretty: {}'.format(pretty))
# -
# ### Keyword arguments
# +
def calculator(first, second, third):
return first + second - third
print(calculator(3, 2, 1))
print(calculator(first=3, second=2, third=1))
# With keyword arguments you can mix the order
print(calculator(third=1, first=3, second=2))
# You can mix arguments and keyword arguments but you have to start with arguments
print(calculator(3, third=1, second=2))
# -
# ### Default arguments
# +
def create_person_info(name, age, job=None, salary=300):
info = {'name': name, 'age': age, 'salary': salary}
# Tambahkan 'job' jika job tidak ada dalam argument
if job:
info.update(dict(job=job))
return info
person1 = create_person_info('Ragnar', 82) #salary nya?
person2 = create_person_info('Uhtred', 22, 'warrior', 10000)
print(person1)
print(person2)
# -
# **Don't use mutable objects as default arguments!**
# +
def simpan_jika_kelipatan_lima(number, magical_list=[]):
if number % 5 == 0:
magical_list.append(number)
return magical_list
print(simpan_jika_kelipatan_lima(100))
print(simpan_jika_kelipatan_lima(105))
print(simpan_jika_kelipatan_lima(123))
print(simpan_jika_kelipatan_lima(123, []))
print(simpan_jika_kelipatan_lima(123))
# -
# Here's how you can achieve desired behavior:
# +
def simpan_jika_kelipatan_lima(number, magical_list=None):
if not magical_list:
magical_list = []
if number % 5 == 0:
magical_list.append(number)
return magical_list
print(simpan_jika_kelipatan_lima(100))
print(simpan_jika_kelipatan_lima(105))
print(simpan_jika_kelipatan_lima(123))
print(simpan_jika_kelipatan_lima(123, []))
print(simpan_jika_kelipatan_lima(123))
# -
# ### Docstrings
# Strings for documenting your functions, methods, modules and variables.
# +
def fungsi_penjumlahan(val1, val2):
"""Fungsi ini berguna untuk menjumlahkan angka yg kurang penting :)"""
print('jumlah: {}'.format(val1 + val2))
print(help(fungsi_penjumlahan))
# +
def penjumlahan(val1, val2):
"""Fungsi ini berguna untuk menjumlahkan angka yg kurang penting :)
Args:
val1: Parameter atau angka pertama.
val2: Parameter atau angka kedua.
Returns:
Total val1 + val2.
"""
return val1 + val2
print(help(penjumlahan))
# -
# ### [`pass`](https://docs.python.org/3/reference/simple_stmts.html#the-pass-statement) statement
# `pass` adalah sebuah statemen yang tidak melakukan apa apa ketika dijalankan.
# +
def pass_function(some_argument):
pass
def another_pass_function():
pass
|
dasar/function/function.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://cocl.us/pytorch_link_top">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/Pytochtop.png" width="750" alt="IBM Product " />
# </a>
#
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/cc-logo-square.png" width="200" alt="cognitiveclass.ai logo" />
#
# <h1>Logistic Regression and Bad Initialization Value</h1>
#
# <h2>Objective</h2><ul><li> How bad initialization value can affects the accuracy of model. .</li></ul>
#
# <h2>Table of Contents</h2>
# <p>In this lab, you will see what happens when you use the root mean square error cost or total loss function and select a bad initialization value for the parameter values.</p>
#
# <ul>
# <li><a href="#Makeup_Data">Make Some Data</a></li>
# <li><a href="#Model_Cost">Create the Model and Cost Function the PyTorch way</a></li>
# <li><a href="#BGD">Train the Model:Batch Gradient Descent</a></li>
# </ul>
#
# <br>
# <p>Estimated Time Needed: <strong>30 min</strong></p>
#
# <hr>
#
# <h2>Preparation</h2>
#
# We'll need the following libraries:
#
# +
# Import the libraries we need for this lab
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
import torch
from torch.utils.data import Dataset, DataLoader
import torch.nn as nn
# -
# Helper functions
#
# The class <code>plot_error_surfaces</code> is just to help you visualize the data space and the Parameter space during training and has nothing to do with Pytorch.
#
# +
# Create class for plotting and the function for plotting
class plot_error_surfaces(object):
# Construstor
def __init__(self, w_range, b_range, X, Y, n_samples = 30, go = True):
W = np.linspace(-w_range, w_range, n_samples)
B = np.linspace(-b_range, b_range, n_samples)
w, b = np.meshgrid(W, B)
Z = np.zeros((30, 30))
count1 = 0
self.y = Y.numpy()
self.x = X.numpy()
for w1, b1 in zip(w, b):
count2 = 0
for w2, b2 in zip(w1, b1):
Z[count1, count2] = np.mean((self.y - (1 / (1 + np.exp(-1*w2 * self.x - b2)))) ** 2)
count2 += 1
count1 += 1
self.Z = Z
self.w = w
self.b = b
self.W = []
self.B = []
self.LOSS = []
self.n = 0
if go == True:
plt.figure()
plt.figure(figsize=(7.5, 5))
plt.axes(projection='3d').plot_surface(self.w, self.b, self.Z, rstride=1, cstride=1, cmap='viridis', edgecolor='none')
plt.title('Loss Surface')
plt.xlabel('w')
plt.ylabel('b')
plt.show()
plt.figure()
plt.title('Loss Surface Contour')
plt.xlabel('w')
plt.ylabel('b')
plt.contour(self.w, self.b, self.Z)
plt.show()
# Setter
def set_para_loss(self, model, loss):
self.n = self.n + 1
self.W.append(list(model.parameters())[0].item())
self.B.append(list(model.parameters())[1].item())
self.LOSS.append(loss)
# Plot diagram
def final_plot(self):
ax = plt.axes(projection='3d')
ax.plot_wireframe(self.w, self.b, self.Z)
ax.scatter(self.W, self.B, self.LOSS, c='r', marker='x', s=200, alpha=1)
plt.figure()
plt.contour(self.w, self.b, self.Z)
plt.scatter(self.W, self.B, c='r', marker='x')
plt.xlabel('w')
plt.ylabel('b')
plt.show()
# Plot diagram
def plot_ps(self):
plt.subplot(121)
plt.ylim
plt.plot(self.x, self.y, 'ro', label="training points")
plt.plot(self.x, self.W[-1] * self.x + self.B[-1], label="estimated line")
plt.plot(self.x, 1 / (1 + np.exp(-1 * (self.W[-1] * self.x + self.B[-1]))), label='sigmoid')
plt.xlabel('x')
plt.ylabel('y')
plt.ylim((-0.1, 2))
plt.title('Data Space Iteration: ' + str(self.n))
plt.show()
plt.subplot(122)
plt.contour(self.w, self.b, self.Z)
plt.scatter(self.W, self.B, c='r', marker='x')
plt.title('Loss Surface Contour Iteration' + str(self.n))
plt.xlabel('w')
plt.ylabel('b')
# Plot the diagram
def PlotStuff(X, Y, model, epoch, leg=True):
plt.plot(X.numpy(), model(X).detach().numpy(), label=('epoch ' + str(epoch)))
plt.plot(X.numpy(), Y.numpy(), 'r')
if leg == True:
plt.legend()
else:
pass
# -
# Set the random seed:
#
# +
# Set random seed
torch.manual_seed(0)
# -
# <!--Empty Space for separating topics-->
#
# <h2 id="Makeup_Data">Get Some Data </h2>
#
# Create the <code>Data</code> class
#
# +
# Create the data class
class Data(Dataset):
# Constructor
def __init__(self):
self.x = torch.arange(-1, 1, 0.1).view(-1, 1)
self.y = torch.zeros(self.x.shape[0], 1)
self.y[self.x[:, 0] > 0.2] = 1
self.len = self.x.shape[0]
# Getter
def __getitem__(self, index):
return self.x[index], self.y[index]
# Get Length
def __len__(self):
return self.len
# -
# Make <code>Data</code> object
#
# +
# Create Data object
data_set = Data()
# -
# <!--Empty Space for separating topics-->
#
# <h2 id="Model_Cost">Create the Model and Total Loss Function (Cost)</h2>
#
# Create a custom module for logistic regression:
#
# +
# Create logistic_regression class
class logistic_regression(nn.Module):
# Constructor
def __init__(self, n_inputs):
super(logistic_regression, self).__init__()
self.linear = nn.Linear(n_inputs, 1)
# Prediction
def forward(self, x):
yhat = torch.sigmoid(self.linear(x))
return yhat
# -
# Create a logistic regression object or model:
#
# +
# Create the logistic_regression result
model = logistic_regression(1)
# -
# Replace the random initialized variable values with some predetermined values that will not converge:
#
# +
# Set the weight and bias
model.state_dict() ['linear.weight'].data[0] = torch.tensor([[-5]])
model.state_dict() ['linear.bias'].data[0] = torch.tensor([[-10]])
print("The parameters: ", model.state_dict())
# -
# Create a <code> plot_error_surfaces</code> object to visualize the data space and the parameter space during training:
#
# +
# Create the plot_error_surfaces object
get_surface = plot_error_surfaces(15, 13, data_set[:][0], data_set[:][1], 30)
# -
# Define the dataloader, the cost or criterion function, the optimizer:
#
# +
# Create dataloader object, crierion function and optimizer.
trainloader = DataLoader(dataset=data_set, batch_size=3)
criterion_rms = nn.MSELoss()
learning_rate = 2
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
# -
# <a id="ref2"></a>
#
# <h2 align=center>Train the Model via Batch Gradient Descent </h2>
#
# Train the model
#
# +
# Train the model
def train_model(epochs):
for epoch in range(epochs):
for x, y in trainloader:
yhat = model(x)
loss = criterion_rms(yhat, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
get_surface.set_para_loss(model, loss.tolist())
if epoch % 20 == 0:
get_surface.plot_ps()
train_model(100)
# -
# Get the actual class of each sample and calculate the accuracy on the test data:
#
# +
# Make the Prediction
yhat = model(data_set.x)
label = yhat > 0.5
print("The accuracy: ", torch.mean((label == data_set.y.type(torch.ByteTensor)).type(torch.float)))
# -
# Accuracy is 60% compared to 100% in the last lab using a good Initialization value.
#
# <!--Empty Space for separating topics-->
#
# <a href="http://cocl.us/pytorch_link_bottom">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/notebook_bottom%20.png" width="750" alt="PyTorch Bottom" />
# </a>
#
# <h2>About the Authors:</h2>
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/"><NAME></a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
#
# Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/"><NAME></a>, <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a"><NAME></a>
#
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ---------- | ----------------------------------------------------------- |
# | 2020-09-23 | 2.0 | Shubham | Migrated Lab to Markdown and added to course repo in GitLab |
#
# <hr>
#
# Copyright © 2018 <a href="cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu">cognitiveclass.ai</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.
#
|
5.2.2bad_inshilization_logistic_regression_with_mean_square_error_v2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Working out some issues with the Ticktock class and keeping it in this Notebook because that's what I want to DO.
import spacepy.time as spt
import datetime as dt
dts = spt.doy2date([2002]*4, range(186,190), dtobj=True)
dts
dts = spt.Ticktock(dts,'UTC')
dts.DOY
# Ticktock object creation
isodates = []
|
cuspStudy/.ipynb_checkpoints/Datetime Notebook-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pyenv_tesseract
# language: python
# name: pyenv_tesseract
# ---
from __future__ import absolute_import, division, print_function
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
print(train_images.shape)
print(train_labels.shape)
print(train_labels[0])
# ## Show some sample images
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
plt.figure(figsize=(10,10))
for i in range(20):
plt.subplot(5,5,i+1)
plt.xticks([]) # do not show axis
plt.yticks([]) # do not show axis
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
train_images = train_images / 255.0
test_images = test_images / 255.0
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28,28)),
keras.layers.Dense(512, activation=tf.nn.relu),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
fit = model.fit(train_images, train_labels, epochs=5, batch_size=32)
model.summary()
# +
def plot_history_loss(fit, axL):
# Plot the loss in the history
axL.plot(fit.history['loss'],label="loss for training")
# axL.plot(fit.history['val_loss'],label="loss for validation")
axL.set_title('model loss')
axL.set_xlabel('epoch')
axL.set_ylabel('loss')
axL.legend(loc='upper right')
# acc
def plot_history_acc(fit, axR):
# Plot the loss in the history
axR.plot(fit.history['acc'],label="loss for training")
# axR.plot(fit.history['val_acc'],label="loss for validation")
axR.set_title('model accuracy')
axR.set_xlabel('epoch')
axR.set_ylabel('accuracy')
axR.legend(loc='upper right')
# -
fig, (axL, axR) = plt.subplots(ncols=2, figsize=(10,4))
plot_history_loss(fit, axL)
plot_history_acc(fit, axR)
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('Test accuracy:', test_acc)
predictions = model.predict(test_images)
print(np.argmax(predictions[0]))
print(test_labels[0])
# +
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
print(predictions_array)
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# -
i = 12
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
plt.show()
|
mnist_fasshion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: rocketPy
# language: python
# name: rocketpy
# ---
import numpy as np
# +
class Quaternion(np.ndarray):
"""Only works with unit quaternions (as needed to describe rotations)"""
def __new__(cls, input_array=[0.,1.,0.,0.]):
"""By default works with 0 rotation about x axis"""
input_array = np.array(input_array)/np.linalg.norm(input_array)
obj = np.asarray(input_array).view(cls)
return obj
def __array__finalize(self, obj):
pass
@classmethod
def from_angle(cls, theta, axis):
"""A rotation of angle `theta` about the axis `ax, ay, az`. Allows the axis to be not normalized"""
axis = np.array(axis)/np.linalg.norm(axis)
s = np.cos(theta / 2)
v = np.sin(theta / 2) * axis
return cls([s, *v])
def normalize(self):
"""Convert quaternion to unit vector"""
self/np.linalg.norm(self)
def rot_matrix(self):
"""Generate a rotation matrix"""
R = np.array([[1 - 2 * self[2]**2 - 2 * self[3]**2,
2 * self[1] * self[2] - 2 * self[0] * self[3],
2 * self[1] * self[3] + 2 * self[0] * self[2]],
[2 * self[1] * self[2] + 2 * self[0] * self[3],
1 - 2 * self[1]**2 - 2 * self[3]**2,
2 * self[2] * self[3] - 2 * self[0] * self[1]],
[2 * self[1] * self[3] - 2 * self[0] * self[2],
2 * self[2] * self[3] + 2 * self[0] * self[1],
1 - 2 * self[1]**2 - 2 * self[2]**2]])
return R
def rate_of_change(self, omega):
"""Return the rate of change of the quaternion based on an angular velocity"""
# follows the formulation in Box
# note that there are other methods, which may be more accurate
omega = np.array(omega)
s = self[0]
v = np.array(self[1:])
sdot = - 0.5 * (omega @ v) # mistake in Box eqn 7 (based on https://www.sciencedirect.com/topics/computer-science/quaternion-multiplication and http://web.cs.iastate.edu/~cs577/handouts/quaternion.pdf)
vdot = 0.5 * (s * omega + np.cross(omega, v))
# returns a quaternion object as a rate of change is really just a quaternion
return Quaternion([sdot, *vdot])
def __mul__(self, other):
raise NotImplementedError
# -
q = Quaternion()
q
q.rot_matrix()
q.rate_of_change(omega=[0.,1,1])
[0,1,1] @ np.array(q[1:])
q[0]
|
examples/quaternion_rewrite.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/94JuHo/study_for_deeplearning/blob/master/MS_AI/03.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="qcd9X5gVXNVq" colab_type="code" outputId="f22709e2-593f-4c09-866b-3a953f25bb48" colab={"base_uri": "https://localhost:8080/", "height": 68}
print("Enter your name:")
somebody = input()
print('Hi', somebody, "How are you today?")
# + id="R_3TGaeUX2ig" colab_type="code" outputId="ff48c8e9-97a6-484a-8bc4-76f890502307" colab={"base_uri": "https://localhost:8080/", "height": 51}
temperature = float(input("온도를 입력하세요: "))
print(temperature)
# + id="VS6ldDXOlJDg" colab_type="code" outputId="1d41cf52-eb0a-4af5-8115-1df21c0f32bc" colab={"base_uri": "https://localhost:8080/", "height": 68}
colors = ['red', 'blue', 'green']
print(colors[0])
print(colors[2])
print(len(colors))
# + id="OuiVHJq2ld3x" colab_type="code" outputId="454e99a2-ea1b-408d-9e09-2ea5d7f74557" colab={"base_uri": "https://localhost:8080/", "height": 34}
cities = ['서울', '부산', '인천', '대구', '대전', '광주', '울산', '수원']
cities[0:6]
# + id="ZTVnwKhnlosb" colab_type="code" outputId="e32e259b-bbe1-4c85-8419-01f8371ded68" colab={"base_uri": "https://localhost:8080/", "height": 34}
cities[0:5]
# + id="MJ-1VN4tlrb_" colab_type="code" outputId="464e6c24-d49d-4703-a219-1ad8d56a8013" colab={"base_uri": "https://localhost:8080/", "height": 34}
cities[5:]
# + id="FubtkQN_ls_h" colab_type="code" outputId="a1b88254-01ac-4dd4-b66a-fbc8af64d18c" colab={"base_uri": "https://localhost:8080/", "height": 34}
cities = ['서울', '부산', '인천', '대구', '대전', '광주', '울산', '수원']
print(cities[-8:])
# + id="f2uZS8zymCxX" colab_type="code" outputId="fcbad380-3def-451a-afd0-c3b671d27276" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(cities[:])
# + id="1AtGR-QCmUHi" colab_type="code" outputId="c28aa112-0c26-435f-d826-eef995550054" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(cities[-50:50])
# + id="mfLnCLHpmZl8" colab_type="code" outputId="4d045eec-96e2-4973-90a2-26baf0069e72" colab={"base_uri": "https://localhost:8080/", "height": 34}
cities[::2]
# + id="4ZFFl3SGmdha" colab_type="code" outputId="d37104b3-8c34-4897-b4bc-172d0e422078" colab={"base_uri": "https://localhost:8080/", "height": 34}
cities[::-1]
# + id="5VB53NntmiJr" colab_type="code" outputId="0516a768-709d-4aac-afa1-893e0d0239b7" colab={"base_uri": "https://localhost:8080/", "height": 34}
color1 = ['red', 'blue', 'green']
color2 = ['orange', 'black', 'white']
print(color1 + color2)
# + id="iM6Up3wFm9e3" colab_type="code" outputId="e35586f4-fa97-444d-f72c-042ca3e67539" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(color1)
# + id="_iqYBOSLnbwX" colab_type="code" outputId="53faf705-6128-4177-b6e3-66fedaaeb442" colab={"base_uri": "https://localhost:8080/", "height": 34}
total_color = color1 + color2
total_color
# + id="ql4svww5nePC" colab_type="code" outputId="78818bf3-963a-452d-d21b-09230bb06d99" colab={"base_uri": "https://localhost:8080/", "height": 34}
color1 * 2
# + id="ObLhofIJoS1Y" colab_type="code" outputId="5d2d47d4-c3da-49cf-df47-a652410c7930" colab={"base_uri": "https://localhost:8080/", "height": 34}
'blue' in color2
# + id="31HyDmlIoXgc" colab_type="code" outputId="074e7374-6324-4743-c9ef-a303b134a61c" colab={"base_uri": "https://localhost:8080/", "height": 34}
color = ['red', 'blue', 'green']
color.append('white')
color
# + id="vRScPHlSoe0c" colab_type="code" outputId="d48f5bbc-8841-4589-dfab-030f1c571c50" colab={"base_uri": "https://localhost:8080/", "height": 34}
color = ['red', 'blue', 'green']
color.extend(['black', 'purple'])
color
# + id="O0ZAuctypIr8" colab_type="code" outputId="49c9550b-5310-4944-f280-997847c2ca6d" colab={"base_uri": "https://localhost:8080/", "height": 34}
color = ['red', 'green', 'blue']
color.insert(0, 'orange')
color
# + id="MGIH2pxjpPS9" colab_type="code" outputId="6712fc74-f82a-4222-df9c-383cc4a3c564" colab={"base_uri": "https://localhost:8080/", "height": 34}
color.remove('red')
color
# + id="hfXH_QuipVPD" colab_type="code" outputId="9287d923-c402-4929-d915-5b2a1bebc044" colab={"base_uri": "https://localhost:8080/", "height": 34}
color = ['red', 'blue', 'green']
color[0] = 'orange'
color
# + id="eh0Z1I_lpb2q" colab_type="code" outputId="52b54095-13ef-4fef-dcd0-a6bb6040821b" colab={"base_uri": "https://localhost:8080/", "height": 34}
del color[0]
color
# + id="vRvXz4lNpd40" colab_type="code" outputId="bc4b22db-c604-4320-c181-ce47d5dbe764" colab={"base_uri": "https://localhost:8080/", "height": 34}
t = [1, 2, 3]
a, b, c = t
print(t, a, b, c)
# + id="UVbUWk2DpqSm" colab_type="code" outputId="3ffe8632-3fe3-47b5-c22a-e9dc5ea5b8fa" colab={"base_uri": "https://localhost:8080/", "height": 181}
t = [ 1, 2, 3]
a, b, c, d, e = t
# + id="37znNKioptZ0" colab_type="code" outputId="5c458f0a-05b2-494f-900e-e3e8162dcd8f" colab={"base_uri": "https://localhost:8080/", "height": 34}
kor_score = [49, 79, 20, 100, 80]
math_score = [43, 59, 85, 30, 90]
eng_score = [49, 79, 48, 80, 100]
midterm_score = [kor_score, math_score, eng_score]
midterm_score
# + id="weMtwCSIqDWo" colab_type="code" outputId="720f43ae-ce7b-48dc-8211-f24a0626a013" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(midterm_score[0][2])
# + id="nRkWAff2uyZK" colab_type="code" outputId="7dd58ce1-0b74-4dd0-b501-602d7b26d00e" colab={"base_uri": "https://localhost:8080/", "height": 34}
math_score[0] = 1000
midterm_score
# + id="TXxvte_ju4PR" colab_type="code" outputId="ac8314c6-e4fe-4a29-fbaa-94effcbb2880" colab={"base_uri": "https://localhost:8080/", "height": 34}
a = 300
b = 300
a is b
# + id="ggP8B6Ivu8Vw" colab_type="code" outputId="42084edd-bd56-45d7-d591-68274f58b85d" colab={"base_uri": "https://localhost:8080/", "height": 34}
a == b
# + id="xNXpTN2ju9pQ" colab_type="code" outputId="b01d13d6-e570-460a-a936-5f3addf71ac2" colab={"base_uri": "https://localhost:8080/", "height": 34}
a = 1
b = 1
a is b
# + id="AyI5cQzwvOL_" colab_type="code" outputId="996ae7aa-68ea-4baa-c6e2-8cf0b76967b3" colab={"base_uri": "https://localhost:8080/", "height": 34}
a == b
# + id="-AZTfClNvO9u" colab_type="code" outputId="de0bcfea-a7b6-43be-ac52-18ab425429b0" colab={"base_uri": "https://localhost:8080/", "height": 34}
a = ["color", 1, 0.2]
color = ['yellow', 'blue', 'green', 'black', 'purple']
a[0] = color
print(a)
# + id="xC1hb5K2v33v" colab_type="code" outputId="4c1126b7-53b3-4a9d-9601-da4119277dd7" colab={"base_uri": "https://localhost:8080/", "height": 34}
a = [5, 4, 3, 2, 1]
b = [1, 2, 3, 4, 5]
b = a
print(b)
# + id="8P7XqnQ3v_zF" colab_type="code" outputId="98684069-e440-4864-d9f8-bbcdd37b50e9" colab={"base_uri": "https://localhost:8080/", "height": 34}
a.sort()
print(b)
# + id="Zl_MtaNNwDuV" colab_type="code" outputId="1c7fe0d8-1733-4008-b3d0-782a661a6981" colab={"base_uri": "https://localhost:8080/", "height": 34}
b = [6, 7, 8, 9, 10]
print(a, b)
# + id="VbG-jzvlwIm3" colab_type="code" colab={}
|
MS_AI/03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="_jQ1tEQCxwRx"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="V_sgB_5dx1f1"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="rF2x3qooyBTI"
# # Deep Convolutional Generative Adversarial Network
# + [markdown] colab_type="text" id="0TD5ZrvEMbhZ"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/beta/tutorials/generative/dcgan">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/generative/dcgan.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/generative/dcgan.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/r2/tutorials/generative/dcgan.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="ITZuApL56Mny"
# This tutorial demonstrates how to generate images of handwritten digits using a [Deep Convolutional Generative Adversarial Network](https://arxiv.org/pdf/1511.06434.pdf) (DCGAN). The code is written using the [Keras Sequential API](https://www.tensorflow.org/guide/keras) with a `tf.GradientTape` training loop.
# + [markdown] colab_type="text" id="2MbKJY38Puy9"
# ## What are GANs?
# [Generative Adversarial Networks](https://arxiv.org/abs/1406.2661) (GANs) are one of the most interesting ideas in computer science today. Two models are trained simultaneously by an adversarial process. A *generator* ("the artist") learns to create images that look real, while a *discriminator* ("the art critic") learns to tell real images apart from fakes.
#
# 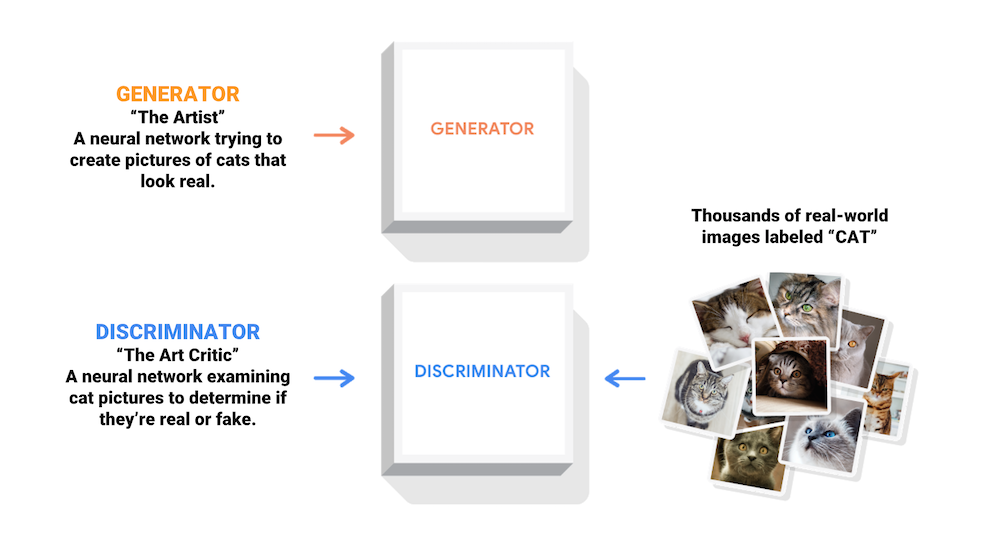
#
# During training, the *generator* progressively becomes better at creating images that look real, while the *discriminator* becomes better at telling them apart. The process reaches equilibrium when the *discriminator* can no longer distinguish real images from fakes.
#
# 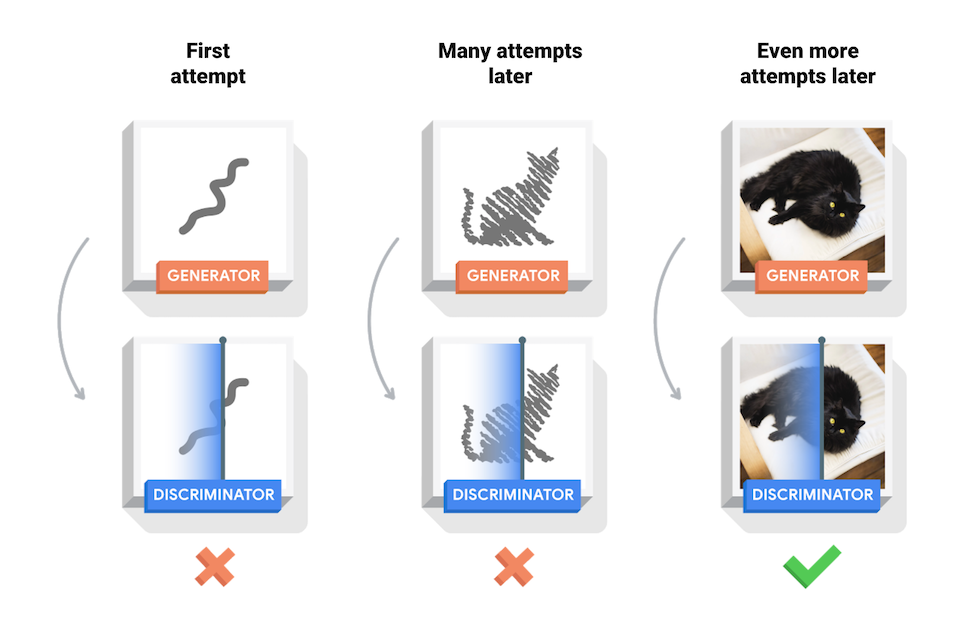
#
# This notebook demonstrates this process on the MNIST dataset. The following animation shows a series of images produced by the *generator* as it was trained for 50 epochs. The images begin as random noise, and increasingly resemble hand written digits over time.
#
# 
#
# To learn more about GANs, we recommend MIT's [Intro to Deep Learning](http://introtodeeplearning.com/) course.
# + [markdown] colab_type="text" id="e1_Y75QXJS6h"
# ### Import TensorFlow and other libraries
# + colab={} colab_type="code" id="J5oue0oqCkZZ"
from __future__ import absolute_import, division, print_function, unicode_literals
# + colab={} colab_type="code" id="g5RstiiB8V-z"
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
# + colab={} colab_type="code" id="WZKbyU2-AiY-"
import tensorflow as tf
# + colab={} colab_type="code" id="wx-zNbLqB4K8"
tf.__version__
# + colab={} colab_type="code" id="YzTlj4YdCip_"
# To generate GIFs
# !pip install imageio
# + colab={} colab_type="code" id="YfIk2es3hJEd"
import glob
import imageio
import matplotlib.pyplot as plt
import numpy as np
import os
import PIL
from tensorflow.keras import layers
import time
from IPython import display
# + [markdown] colab_type="text" id="iYn4MdZnKCey"
# ### Load and prepare the dataset
#
# You will use the MNIST dataset to train the generator and the discriminator. The generator will generate handwritten digits resembling the MNIST data.
# + colab={} colab_type="code" id="a4fYMGxGhrna"
(train_images, train_labels), (_, _) = tf.keras.datasets.mnist.load_data()
# + colab={} colab_type="code" id="NFC2ghIdiZYE"
train_images = train_images.reshape(train_images.shape[0], 28, 28, 1).astype('float32')
train_images = (train_images - 127.5) / 127.5 # Normalize the images to [-1, 1]
# + colab={} colab_type="code" id="S4PIDhoDLbsZ"
BUFFER_SIZE = 60000
BATCH_SIZE = 256
# + colab={} colab_type="code" id="-yKCCQOoJ7cn"
# Batch and shuffle the data
train_dataset = tf.data.Dataset.from_tensor_slices(train_images).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
# + [markdown] colab_type="text" id="THY-sZMiQ4UV"
# ## Create the models
#
# Both the generator and discriminator are defined using the [Keras Sequential API](https://www.tensorflow.org/guide/keras#sequential_model).
# + [markdown] colab_type="text" id="-tEyxE-GMC48"
# ### The Generator
#
# The generator uses `tf.keras.layers.Conv2DTranspose` (upsampling) layers to produce an image from a seed (random noise). Start with a `Dense` layer that takes this seed as input, then upsample several times until you reach the desired image size of 28x28x1. Notice the `tf.keras.layers.LeakyReLU` activation for each layer, except the output layer which uses tanh.
# + colab={} colab_type="code" id="6bpTcDqoLWjY"
def make_generator_model():
model = tf.keras.Sequential()
model.add(layers.Dense(7*7*256, use_bias=False, input_shape=(100,)))
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Reshape((7, 7, 256)))
assert model.output_shape == (None, 7, 7, 256) # Note: None is the batch size
model.add(layers.Conv2DTranspose(128, (5, 5), strides=(1, 1), padding='same', use_bias=False))
assert model.output_shape == (None, 7, 7, 128)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(64, (5, 5), strides=(2, 2), padding='same', use_bias=False))
assert model.output_shape == (None, 14, 14, 64)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(1, (5, 5), strides=(2, 2), padding='same', use_bias=False, activation='tanh'))
assert model.output_shape == (None, 28, 28, 1)
return model
# + [markdown] colab_type="text" id="GyWgG09LCSJl"
# Use the (as yet untrained) generator to create an image.
# + colab={} colab_type="code" id="gl7jcC7TdPTG"
generator = make_generator_model()
noise = tf.random.normal([1, 100])
generated_image = generator(noise, training=False)
plt.imshow(generated_image[0, :, :, 0], cmap='gray')
# + [markdown] colab_type="text" id="D0IKnaCtg6WE"
# ### The Discriminator
#
# The discriminator is a CNN-based image classifier.
# + colab={} colab_type="code" id="dw2tPLmk2pEP"
def make_discriminator_model():
model = tf.keras.Sequential()
model.add(layers.Conv2D(64, (5, 5), strides=(2, 2), padding='same',
input_shape=[28, 28, 1]))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(128, (5, 5), strides=(2, 2), padding='same'))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Flatten())
model.add(layers.Dense(1))
return model
# + [markdown] colab_type="text" id="QhPneagzCaQv"
# Use the (as yet untrained) discriminator to classify the generated images as real or fake. The model will be trained to output positive values for real images, and negative values for fake images.
# + colab={} colab_type="code" id="gDkA05NE6QMs"
discriminator = make_discriminator_model()
decision = discriminator(generated_image)
print (decision)
# + [markdown] colab_type="text" id="0FMYgY_mPfTi"
# ## Define the loss and optimizers
#
# Define loss functions and optimizers for both models.
#
# + colab={} colab_type="code" id="psQfmXxYKU3X"
# This method returns a helper function to compute cross entropy loss
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
# + [markdown] colab_type="text" id="PKY_iPSPNWoj"
# ### Discriminator loss
#
# This method quantifies how well the discriminator is able to distinguish real images from fakes. It compares the discriminator's predictions on real images to an array of 1s, and the discriminator's predictions on fake (generated) images to an array of 0s.
# + colab={} colab_type="code" id="wkMNfBWlT-PV"
def discriminator_loss(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output), fake_output)
total_loss = real_loss + fake_loss
return total_loss
# + [markdown] colab_type="text" id="Jd-3GCUEiKtv"
# ### Generator loss
# The generator's loss quantifies how well it was able to trick the discriminator. Intuitively, if the generator is performing well, the discriminator will classify the fake images as real (or 1). Here, we will compare the discriminators decisions on the generated images to an array of 1s.
# + colab={} colab_type="code" id="90BIcCKcDMxz"
def generator_loss(fake_output):
return cross_entropy(tf.ones_like(fake_output), fake_output)
# + [markdown] colab_type="text" id="MgIc7i0th_Iu"
# The discriminator and the generator optimizers are different since we will train two networks separately.
# + colab={} colab_type="code" id="iWCn_PVdEJZ7"
generator_optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_optimizer = tf.keras.optimizers.Adam(1e-4)
# + [markdown] colab_type="text" id="mWtinsGDPJlV"
# ### Save checkpoints
# This notebook also demonstrates how to save and restore models, which can be helpful in case a long running training task is interrupted.
# + colab={} colab_type="code" id="CA1w-7s2POEy"
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
# + [markdown] colab_type="text" id="Rw1fkAczTQYh"
# ## Define the training loop
#
#
# + colab={} colab_type="code" id="NS2GWywBbAWo"
EPOCHS = 50
noise_dim = 100
num_examples_to_generate = 16
# We will reuse this seed overtime (so it's easier)
# to visualize progress in the animated GIF)
seed = tf.random.normal([num_examples_to_generate, noise_dim])
# + [markdown] colab_type="text" id="jylSonrqSWfi"
# The training loop begins with generator receiving a random seed as input. That seed is used to produce an image. The discriminator is then used to classify real images (drawn from the training set) and fakes images (produced by the generator). The loss is calculated for each of these models, and the gradients are used to update the generator and discriminator.
# + colab={} colab_type="code" id="3t5ibNo05jCB"
# Notice the use of `tf.function`
# This annotation causes the function to be "compiled".
@tf.function
def train_step(images):
noise = tf.random.normal([BATCH_SIZE, noise_dim])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(noise, training=True)
real_output = discriminator(images, training=True)
fake_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
# + colab={} colab_type="code" id="2M7LmLtGEMQJ"
def train(dataset, epochs):
for epoch in range(epochs):
start = time.time()
for image_batch in dataset:
train_step(image_batch)
# Produce images for the GIF as we go
display.clear_output(wait=True)
generate_and_save_images(generator,
epoch + 1,
seed)
# Save the model every 15 epochs
if (epoch + 1) % 15 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time for epoch {} is {} sec'.format(epoch + 1, time.time()-start))
# Generate after the final epoch
display.clear_output(wait=True)
generate_and_save_images(generator,
epochs,
seed)
# + [markdown] colab_type="text" id="2aFF7Hk3XdeW"
# **Generate and save images**
#
#
# + colab={} colab_type="code" id="RmdVsmvhPxyy"
def generate_and_save_images(model, epoch, test_input):
# Notice `training` is set to False.
# This is so all layers run in inference mode (batchnorm).
predictions = model(test_input, training=False)
fig = plt.figure(figsize=(4,4))
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, 0] * 127.5 + 127.5, cmap='gray')
plt.axis('off')
plt.savefig('image_at_epoch_{:04d}.png'.format(epoch))
plt.show()
# + [markdown] colab_type="text" id="dZrd4CdjR-Fp"
# ## Train the model
# Call the `train()` method defined above to train the generator and discriminator simultaneously. Note, training GANs can be tricky. It's important that the generator and discriminator do not overpower each other (e.g., that they train at a similar rate).
#
# At the beginning of the training, the generated images look like random noise. As training progresses, the generated digits will look increasingly real. After about 50 epochs, they resemble MNIST digits. This may take about one minute / epoch with the default settings on Colab.
# + colab={} colab_type="code" id="Ly3UN0SLLY2l"
# %%time
train(train_dataset, EPOCHS)
# + [markdown] colab_type="text" id="rfM4YcPVPkNO"
# Restore the latest checkpoint.
# + colab={} colab_type="code" id="XhXsd0srPo8c"
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
# + [markdown] colab_type="text" id="P4M_vIbUi7c0"
# ## Create a GIF
#
# + colab={} colab_type="code" id="WfO5wCdclHGL"
# Display a single image using the epoch number
def display_image(epoch_no):
return PIL.Image.open('image_at_epoch_{:04d}.png'.format(epoch_no))
# + colab={} colab_type="code" id="5x3q9_Oe5q0A"
display_image(EPOCHS)
# + [markdown] colab_type="text" id="NywiH3nL8guF"
# Use `imageio` to create an animated gif using the images saved during training.
# + colab={} colab_type="code" id="IGKQgENQ8lEI"
anim_file = 'dcgan.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('image*.png')
filenames = sorted(filenames)
last = -1
for i,filename in enumerate(filenames):
frame = 2*(i**0.5)
if round(frame) > round(last):
last = frame
else:
continue
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
import IPython
if IPython.version_info > (6,2,0,''):
display.Image(filename=anim_file)
# + [markdown] colab_type="text" id="cGhC3-fMWSwl"
# If you're working in Colab you can download the animation with the code below:
# + colab={} colab_type="code" id="uV0yiKpzNP1b"
try:
from google.colab import files
except ImportError:
pass
else:
files.download(anim_file)
# + [markdown] colab_type="text" id="k6qC-SbjK0yW"
# ## Next steps
#
# + [markdown] colab_type="text" id="xjjkT9KAK6H7"
# This tutorial has shown the complete code necessary to write and train a GAN. As a next step, you might like to experiment with a different dataset, for example the Large-scale Celeb Faces Attributes (CelebA) dataset [available on Kaggle](https://www.kaggle.com/jessicali9530/celeba-dataset/home). To learn more about GANs we recommend the [NIPS 2016 Tutorial: Generative Adversarial Networks](https://arxiv.org/abs/1701.00160).
#
|
site/en/r2/tutorials/generative/dcgan.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle
import numpy as np
import matplotlib.pyplot as plt
# from keras.models import Model
# from keras.layers import Input, Dense
# import tensorflow.python.util.deprecation as deprecation
# deprecation._PRINT_DEPRECATION_WARNINGS = False
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input, Dense
# -
with open('mnist.pkl', 'rb') as f:
images = pickle.load(f)['images']
images = images.reshape((-1, 28 ** 2))
images = images / 255.
input_stage = Input(shape=(784,))
encoding_stage = Dense(100, activation='relu')(input_stage)
decoding_stage = Dense(784, activation='sigmoid')(encoding_stage)
autoencoder = Model(input_stage, decoding_stage)
autoencoder.compile(loss='binary_crossentropy',
optimizer='adadelta')
autoencoder.fit(images, images, epochs=100)
encoder_output = Model(input_stage, encoding_stage).predict(images[:5])
encoder_output = encoder_output.reshape((-1, 10, 10)) * 255
decoder_output = autoencoder.predict(images[:5])
decoder_output = decoder_output.reshape((-1, 28, 28)) * 255
images = images.reshape((-1, 28, 28))
plt.figure(figsize=(10, 7))
for i in range(5):
plt.subplot(3, 5, i + 1)
plt.imshow(images[i], cmap='gray')
plt.axis('off')
plt.subplot(3, 5, i + 6)
plt.imshow(encoder_output[i], cmap='gray')
plt.axis('off')
plt.subplot(3, 5, i + 11)
plt.imshow(decoder_output[i], cmap='gray')
plt.axis('off')
# +
# Unit Test
# +
import unittest
class TestEncodeDecodeMNIST(unittest.TestCase):
def test_images(self):
self.assertEqual(len(images),10000)
def test_input_stage_shape(self):
self.assertListEqual(list(input_stage.shape),[None, 784])
def test_encoding_stage_shape(self):
self.assertListEqual(list(encoding_stage.shape),[None, 100])
def test_decoding_stage_spape(self):
self.assertListEqual(list(decoding_stage.shape),[None, 784])
def test_encoder_output_len(self):
self.assertEqual(len(encoder_output),5)
def test_decoder_output_len(self):
self.assertEqual(len(decoder_output),5)
# -
suite = unittest.TestLoader().loadTestsFromTestCase(TestEncodeDecodeMNIST)
unittest.TextTestRunner(verbosity=2).run(suite)
|
Chapter05/tests/Activity5.02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Importing libraries for the model**
# import the basic libraries to work with dataframes and arrays
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# ema related imports
from ema_workbench import (Model, MultiprocessingEvaluator, Policy, Scenario, perform_experiments, ema_logging, ScalarOutcome)
from ema_workbench.em_framework.evaluators import SOBOL
from ema_workbench.em_framework.samplers import sample_uncertainties, sample_levers
from ema_workbench.util import ema_logging
from ema_workbench.em_framework.salib_samplers import get_SALib_problem
from SALib.analyze import sobol
import functools
from ema_workbench.em_framework.evaluators import BaseEvaluator
from ema_workbench.em_framework.optimization import (HyperVolume,
EpsilonProgress)
# **Preparing optimization with MORDM results, constraints and the robustness function**
#results from the MORDM optimization in the final assignment file
results_optimize_MORDM = pd.read_csv('results_optimize_MORDM_1.csv')
#the outlook of the dataframe constructed with the MORDM outcomes
results_optimize_MORDM.head()
#retreiving the lowest/worst 3 quartiles for all objectives
#this is used to set the boundaries
#it functions as selecting the cases of interest with a size of 75%
#dumps the best performing quartile
death_optimize_constrain = np.percentile(results_optimize_MORDM['Total Number of Deaths'],75)
annualdamage_optimize_constrain = np.percentile(results_optimize_MORDM['Expected Annual Damage'],75)
totalinvestment_optimize_constrain = np.percentile(results_optimize_MORDM['Total Investment Costs'],75)
allcosts_optimize_constrain = np.percentile(results_optimize_MORDM['All Costs'],75)
dike_optimize_constrain = np.percentile(results_optimize_MORDM['Dike Investment Costs'],75)
rfr_optimize_constrain = np.percentile(results_optimize_MORDM['RfR Total Costs'],75)
evacuation_optimize_constrain = np.percentile(results_optimize_MORDM['Expected Evacuation Costs'],75)
# +
#robustness function, changing the objection directions to what Rijkswaterstaat prefers.
def robustness(direction, threshold, data):
#making clear the direction is correctly implemented
#mostly needed in case any of the objectives would be maximized as all are minimized now.
if direction == SMALLER:
return np.sum(data<=threshold)/data.shape[0]
else:
return np.sum(data>=threshold)/data.shape[0]
SMALLER = 'SMALLER'
LARGER = 'LARGER'
moro_investment = functools.partial(robustness, SMALLER, totalinvestment_optimize_constrain)
moro_annual_damage = functools.partial(robustness, SMALLER, annualdamage_optimize_constrain)
moro_death = functools.partial(robustness, SMALLER, death_optimize_constrain)
moro_dike = functools.partial(robustness, SMALLER, dike_optimize_constrain)
moro_rfr = functools.partial(robustness, SMALLER, rfr_optimize_constrain)
moro_evacuation = functools.partial(robustness, SMALLER, evacuation_optimize_constrain)
# +
MAXIMIZE = ScalarOutcome.MAXIMIZE
MINIMIZE = ScalarOutcome.MINIMIZE
#creating the objectives to match the type of outcome wanted
#the objectives are defined as scalaroutcomes
#the intended direction for optimization is added
robustnes_functions = [ScalarOutcome('total investment', kind=MINIMIZE,
variable_name='Total Investment Costs', function=moro_investment),
ScalarOutcome('annual damage', kind=MINIMIZE,
variable_name='Expected Annual Damage', function=moro_annual_damage),
ScalarOutcome('expected death', kind=MINIMIZE,
variable_name='Total Number of Deaths', function=moro_death),
ScalarOutcome('dike investment', kind=MINIMIZE,
variable_name='Dike Investment Costs', function=moro_dike),
ScalarOutcome('RfR Investment', kind=MINIMIZE,
variable_name='RfR Total Costs', function=moro_rfr),
ScalarOutcome('Evacuation Costs', kind=MINIMIZE,
variable_name='Expected Evacuation Costs', function=moro_evacuation)]
# -
# **Using MORO for optimization**
#importing the problem_formulation function from the python file
from problem_formulation_2 import get_model_for_problem_formulation
#the newly defined problem formulation from the problem formulation file
#this problem formulation specifies the outcomes of interest
dike_model = get_model_for_problem_formulation(4)
#creating the number of scenarios and reference scenario for optimization with moro
n_scenarios_moro = 50
ref_scenarios_moro = sample_uncertainties(dike_model, n_scenarios_moro)
#logging function to track the progress of the moro optimization
ema_logging.log_to_stderr(ema_logging.INFO)
# +
#setting boundaries
convergence_value = [HyperVolume(minimum=[0,0,0,0,0,0], maximum=[1.1,1.1,1.1,1.1,1.1,1.1]),
EpsilonProgress()]
#The number of iterations/function evaluations the optimization uses
nfe = 3000
#optimization processs with all results and experiments being stored in two variables
#namely archive_moro and convergence_moro
#setting the epsilons to 0.05. This can be seen as the accuracy required by the optimization
with MultiprocessingEvaluator(dike_model) as evaluator:
archive_moro, convergence_moro = evaluator.robust_optimize(robustnes_functions, ref_scenarios_moro,
nfe=nfe, convergence=convergence_value,
epsilons=[0.05,0.05,0.05,0.05,0.05,0.05]*len(robustnes_functions))
# -
#the most promising policies given the optimization above
archive_moro
#creating the csv of those promising policies in the same folder
archive_moro.to_csv('policies.csv')
# +
#creating subplots reporting change in the epsilon
fig, (ax1, ax2) = plt.subplots(ncols=2, sharex=True, figsize=(8,4))
ax1.plot(convergence_moro.nfe, convergence_moro.epsilon_progress)
ax1.set_ylabel('$\epsilon$-progress')
ax2.plot(convergence_moro.nfe, convergence_moro.hypervolume)
ax2.set_ylabel('hypervolume')
ax1.set_xlabel('number of function evaluations')
ax2.set_xlabel('number of function evaluations')
plt.show()
# -
#creating a set of list of all policies and their respective outcomes for the policies
policies_moro = [Policy("policy_1", **archive_moro.to_dict('records')[0]),
Policy("policy_2", **archive_moro.to_dict('records')[1]),
Policy("policy_3", **archive_moro.to_dict('records')[2]),
Policy("policy_4", **archive_moro.to_dict('records')[3]),
Policy("policy_5", **archive_moro.to_dict('records')[4])]
#
n_scenarios_moro_open = 400
with MultiprocessingEvaluator(dike_model) as evaluator:
results_moro_open = evaluator.perform_experiments(n_scenarios_moro_open, policies_moro)
# **Visualizing the different policies**
#importing seaborn
import seaborn as sns
#unpacking the results into two storage variables
#the first a ndarray the latter as a dictionarry
experiments, outcomes = results_moro_open
# +
#importing the plotting tools to create a 3d stacking plot
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
colors = sns.color_palette('Dark2', 5)
markers = ['o', '^']
oois = ['Total Number of Deaths', 'Total Investment Costs','Expected Annual Damage']
#setting the axis of the 3d plot with the aformentioned performance indicators/objectives
ax.set_xlabel(oois[0])
ax.set_ylabel(oois[1])
ax.set_zlabel(oois[2])
#looping through the assessed combinations of the policy levers in the experiments
for p, policy in enumerate(set(experiments['policy'])):
#selecting based on policy to form grouping in the plot
logical = experiments['policy']==policy
new_outcomes = {key:value[logical] for key, value in outcomes.items()}
new_experiments = experiments[logical]
#logical_index = (new_outcomes['infected fraction R1'][:, 99]>0.1) &\
# (new_experiments['infection ratio region 1']>0.1)
#x_ = new_outcomes[oois[0]][logical_index]
x = new_outcomes[oois[0]]
# y_ = new_outcomes[oois[1]][logical_index,-1]
y = new_outcomes[oois[1]]
# z_ = new_outcomes[oois[2]][logical_index]
z = new_outcomes[oois[2]]
ax.scatter(x, y, z, c=colors[p], marker=markers[1], s=60, label=policy)
#ax.scatter(x_, y_, z_, c=colors[p], marker=markers[1], s=60, label=policy+'*')
ax.legend(loc=2, scatterpoints = 1)
#ax.set_xlim([0, 0.5])
#ax.set_ylim([0, 5e+07])
#ax.set_zlim([0, 50])
fig.set_figheight(8)
fig.set_figwidth(12)
plt.show()
# +
# Show in dimensional stacking
df = pd.DataFrame.from_dict(outcomes)
df = df.assign(policy=experiments['policy'])
# use seaborn to plot the dataframe
grid = sns.pairplot(df, hue='policy', vars=outcomes.keys())
ax = plt.gca()
plt.show()
# -
#changing structured arrays require this library
import numpy.lib.recfunctions as rf
#cleaning experiments to only contain useful information to prim
lever_names_open = [l.name for l in dike_model.levers]
experiments2 = rf.drop_fields(experiments, drop_names=lever_names_open+['policy'],
asrecarray=True)
# **Searching for the policy attributes of interest with Prim**
from ema_workbench.analysis import prim
# +
#using the cleaned experiments from the previous section
#comparing it on number of deaths
x = experiments2
y = outcomes['Total Number of Deaths'] >= np.percentile(outcomes['Total Number of Deaths'], 75)
#using prim to see the amount of cases of interest in a certain box
#these cases of interest have the largest extent of bad performing scenarios on number of deaths
prim_alg = prim.Prim(x,y, threshold=0.66)
box1 = prim_alg.find_box()
# -
#the density/coverage trade-off found using prim
box1.show_tradeoff()
plt.show()
#inspecting the properties of the most promising box
box1.inspect()
#inspection of which attributes of the policies have most impact
box1.inspect(26, style='graph')
plt.show()
policies_moro
policy1 = {'A.1_DikeIncrease':0, 'A.2_DikeIncrease':9, 'A.3_DikeIncrease':0, 'A.4_DikeIncrease':1, 'A.5_DikeIncrease':1, '0_RfR':0, '1_RfR':1, '2_RfR':0, '3_RfR':1, '4_RfR':1, 'EWS_DaysToThreat':3}
policy2 = {'A.1_DikeIncrease':1, 'A.2_DikeIncrease':0, 'A.3_DikeIncrease':0, 'A.4_DikeIncrease':0, 'A.5_DikeIncrease':5, '0_RfR':0, '1_RfR':1, '2_RfR':0, '3_RfR':0, '4_RfR':1, 'EWS_DaysToThreat':0}
policy3 = {'A.1_DikeIncrease':10, 'A.2_DikeIncrease':10, 'A.3_DikeIncrease':0, 'A.4_DikeIncrease':4, 'A.5_DikeIncrease':1, '0_RfR':0, '1_RfR':1, '2_RfR':0, '3_RfR':1, '4_RfR':1, 'EWS_DaysToThreat':0}
policy4 = {'A.1_DikeIncrease':9, 'A.2_DikeIncrease':10, 'A.3_DikeIncrease':0, 'A.4_DikeIncrease':3, 'A.5_DikeIncrease':8, '0_RfR':1, '1_RfR':1, '2_RfR':0, '3_RfR':0, '4_RfR':1, 'EWS_DaysToThreat':0}
policy5 = {'A.1_DikeIncrease':0, 'A.2_DikeIncrease':0, 'A.3_DikeIncrease':0, 'A.4_DikeIncrease':5, 'A.5_DikeIncrease':0, '0_RfR':0, '1_RfR':1, '2_RfR':0, '3_RfR':1, '4_RfR':1, 'EWS_DaysToThreat':0}
outcomes
# +
# outcomes?
# -
pol1 = outcomes("Policy_1", outcomes.to_dict('records')[0])
|
Model/final assignment part2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### EXP: Beta2 QC rating
# - **Aim:** Second attempt of assesing quality control (QC) of brain registration on the Zooniverse platform. Raters are some of zooniverse users who agreed to test new projects and give feedback ( ref: https://www.zooniverse.org/projects/simexp/brain-match ).
#
# - **Exp:**
# - We choose 100 anatomical brain images (?? OK, ?? Maybe and ?? Fail) preprocced with NIAK pipelines from ADHD200 and COBRE datsets.
# - We asked raters on the Zooniverse platform to QC images based on the tutorial and the rated sample images.
import os
import pandas as pd
import numpy as np
import json
import itertools
import seaborn as sns
from sklearn import metrics
from matplotlib import gridspec as gs
import matplotlib.pyplot as plt
from functools import reduce
# %matplotlib inline
# %load_ext rpy2.ipython
sns.set(style="white")
def CustomParser(data):
j1 = json.loads(data)
return j1
# Read raw table
classifications = pd.read_csv('../data/rating/brain-match-classifications-12-10-2018.csv',
converters={'metadata':CustomParser,
'annotations':CustomParser,
'subject_data':CustomParser},
header=0)
# List all workflows
classifications.workflow_name.unique()
# Filter out only specific workflow
ratings = classifications.loc[classifications['workflow_name'].isin(['Start Project'])]
ratings.count()
# extract tagging count
ratings.loc[:,"n_tagging"] = [ len(q[0]['value']) for q in ratings.annotations]
# extract rating count
ratings.loc[:,"rating"] = [ q[1]['value'] for q in ratings.annotations]
# extract subjects id
ratings.loc[:,"ID"] = [ row.subject_data[str(ratings.subject_ids.loc[ind])]['subject_ID'] for ind,row in ratings.iterrows()]
# extract files name
ratings.loc[:,"imgnm"] = [ row.subject_data[str(ratings.subject_ids.loc[ind])]['image1'] for ind,row in ratings.iterrows()]
# How many rating per user
user_count = ratings.user_name.value_counts()
user_count
#select only users that have rated a certain ammount of images
list_user = user_count.index
list_user = list_user[user_count.values>=10]
user_count[list_user]
# remove users with less rating then the selected threshold
ratings = ratings[ratings.user_name.isin(list_user)]
# Drop my test ratings (yassinebha)
mask = [x.user_name != 'Yassinebha' for ind,x in ratings.iterrows()]
ratings = ratings[mask]
ratings.count()
# drop duplicated rating
inc = 0
sum_dup = 0
for ind,user in enumerate(ratings.user_name.unique()):
user_select_df = ratings[ratings.user_name.isin([user])]
mask=~user_select_df.ID.duplicated()
dup = len([m for m in mask if m == False])
sum_dup = sum_dup+ dup
if dup > 0 :
print('{} have {} duplicated ratings'.format(user,dup))
if ind == 0 and inc == 0:
classi_unique= user_select_df[mask]
inc+=1
else:
classi_unique = classi_unique.append(user_select_df[~user_select_df.ID.duplicated()])
inc+=1
print('Total number of duplicated ratings = {}'.format(sum_dup))
# Get the final rating numbers per subject
user_count = classi_unique.user_name.value_counts()
user_count
# plot rating per image distribution
image_count = classi_unique.subject_ids.value_counts()
image_count.plot.hist(grid=True,rwidth=0.9, bins=13,color='#607c8e')
plt.title('Frequency of rating per images')
plt.xlabel('Nuber of Images')
plt.ylabel('Frequency')
plt.grid(axis='y', alpha=0.75)
# +
#Create Users rating dataframe
list_user = user_count.index
concat_rating = [classi_unique[classi_unique.user_name == user][['ID','rating']].rename(columns={'rating': user})
for user in list_user]
df_ratings = reduce(lambda left,right: pd.merge(left,right,how='outer',on='ID'), concat_rating)
df_ratings.head()
# -
# remove duplicates
df_ratings = df_ratings[~df_ratings.ID.duplicated()]
# ### Explore the concensus of rating between images
# Get ratings from images rated more tan N different rates
n = 4 # Minimun number of ratings per image
stuff = np.array([[row.ID,
np.sum(row[1:].values=='Fail'),
np.sum(row[1:].values=='Maybe'),
np.sum(row[1:].values=='OK')]
for ind, row in df_ratings.iterrows() if np.sum([np.sum(row[1:-1].values=='Fail'),
np.sum(row[1:-1].values=='Maybe'),
np.sum(row[1:-1].values=='OK')]) >= n])
df_score = pd.DataFrame(data=stuff, columns=['ID','Fail', 'Maybe', 'OK'])
df_score.head()
# Normalise table's row
df_score_tmp = df_score[['Fail','Maybe','OK']].astype('int')
nb_rating = df_score[['Fail','Maybe','OK']].astype('int').sum(axis="columns")
df_norm = pd.DataFrame( index=df_score.index,columns=['ID','Fail', 'Maybe', 'OK'])
for status in ['Fail','Maybe','OK']:
for image in df_score.index:
df_norm[status][image] = np.int(df_score[status][image])/nb_rating[image]
df_norm['ID'][image] = df_score['ID'][image]
# get max value
max_value = [row.iloc[1:].get_values().max() for ind,row in df_norm.iterrows()]
df_norm.loc[:,'max_value_NoExp'] = max_value
# get concensus rating
s = ['Fail', 'Maybe', 'OK']
#max_rate = [row.iloc[1:].idxmax(axis=1) for ind,row in df_norm.iterrows()]
max_rate = [s[row[1:].values.argmax()] for rid, row in df_norm.iterrows()]
df_norm.loc[:,'concensus_NoExp'] = max_rate
df_norm.head()
# +
#Setting the figure with matplotlib
plt.figure(figsize=(7,5))
#plt.xticks(rotation=90)
plt.rcParams["axes.labelsize"] = 12
#Creating the desired plot
sns.violinplot(x='concensus_NoExp',y='max_value_NoExp',data=df_norm,
inner=None #removes the inner bars inside the violins
)
sns.swarmplot(x='concensus_NoExp',y='max_value_NoExp',data=df_norm,
color='k',#for making the points black
alpha=0.6) #value of alpha will increase the transparency
#Title for the plot
plt.grid(axis='y', alpha=0.75)
plt.title('Distribution of rating concensus')
plt.xlabel('')
plt.ylabel('Concensus rating')
# -
count_ = df_norm.concensus_NoExp[[0 <= row.max_value_NoExp < 0.5 for ind, row in df_norm.iterrows() ]].value_counts()
axes = count_.plot.bar(title = 'Frequency of rating for low concensus')
count_ = df_norm.concensus_NoExp[[0.5 <= row.max_value_NoExp < 0.65 for ind, row in df_norm.iterrows() ]].value_counts()
axes = count_.plot.bar(title = 'Frequency of rating for medium concensus')
count_ = df_norm.concensus_NoExp[[0.6 < row.max_value_NoExp <= 1 for ind, row in df_norm.iterrows() ]].value_counts()
axes = count_.plot.bar(title = 'Frequency of rating for high concensus')
# ### Merge Pilot3 and Beta2 rating and get Kappa score
pilot3 = pd.read_csv('../data/rating/Pilot3_internal_rating.csv')
pilot3.head()
# Merge
merge_ratings = pd.merge(pilot3,df_norm[['ID','concensus_NoExp']],on='ID',how='inner').apply(lambda x: x.str.strip() if x.dtype == "object" else x)
merge_ratings.rename(columns={'concensus_NoExp':'Cons_B2'},inplace=True)
merge_ratings
# Replace OK with 1 , Maybe with 2 and Fail with 3
merge_ratings.replace({'OK':1,'Maybe':2, 'Fail':3}, inplace=True)
merge_ratings.rename(columns={'bpinsard':'Bpin',
'saradupont':'Sdup',
'angelatam':'Atam',
'hereinlies':'Mpel',
'benjamindeleener':'Bdel'},inplace=True)
merge_ratings = merge_ratings[['ID','Yben','Pbel','Atam','Bdel','Sdup','Mpel','Bpin','Cons_B2']]
merge_ratings.head()
# + language="R"
# suppressPackageStartupMessages(library(dplyr))
# #install.packages("irr")
# library(irr)
# -
# Percenteage of agrrement between raters with R package IRR
agree_ = merge_ratings.drop(['ID'],axis=1)
# %Rpush agree_
# agree_n = %R agree(agree_)
print(agree_n)
# +
# FDR correction
from statsmodels.sandbox.stats import multicomp as smi
def fdr_transf(mat,log10 = False):
'''compute fdr of a given matrix'''
row = mat.shape[0]
col = mat.shape[1]
flatt = mat.flatten()
fdr_2d = smi.multipletests(flatt, alpha=0.05, method='fdr_bh')[1]
if log10 == True:
fdr_2d = [-np.log10(ii) if ii != 0 else 50 for ii in fdr_2d ]
fdr_3d = np.reshape(fdr_2d,(row,col))
return fdr_3d
# -
# Kappa calculation
def kappa_score(k_df,log10 = False):
'''compute Kappa between diferent raters organized in dataframe'''
k_store = np.zeros((len(k_df.columns), len(k_df.columns)))
p_store = np.zeros((len(k_df.columns), len(k_df.columns)))
# %Rpush k_df
for user1_id, user1 in enumerate(k_df.columns):
for user2_id, user2 in enumerate(k_df.columns):
weight = np.unique(kappa_df[[user1,user2]])
# %Rpush user1_id user1 user2_id user2 weight
# kappaR = %R kappa2(k_df[,c(user1,user2)],weight)
# store the kappa
k_store[user1_id, user2_id] = [kappaR[x][0] for x in range(np.shape(kappaR)[0])][4]
p_store[user1_id, user2_id] = [kappaR[x][0] for x in range(np.shape(kappaR)[0])][-1]
# FDR Correction
p_store = fdr_transf(p_store,log10)
return k_store, p_store
# +
# Get Kappa score out of all different combination of ratings
kappa_df = merge_ratings.drop(['ID'],axis=1)
kappa_store, Pval_store = kappa_score(kappa_df)
mean_kap = np.mean(kappa_store[np.triu_indices(len(kappa_store),k=1)])
std_kap = np.std(kappa_store[np.triu_indices(len(kappa_store),k=1)])
print('Mean Kappa : {0:.2f} , std : {1:.2f}\n'.format(mean_kap, std_kap))
#calculte the over all kappa values of all ratings
# %Rpush kappa_df
# fleiss_kappa = %R kappam.fleiss(kappa_df,c(0,1,2))
print(fleiss_kappa)
# +
# Plot kappa matrix
kappa_out = pd.DataFrame(kappa_store,
index=kappa_df.columns.get_values(),
columns=kappa_df.columns.get_values())
# Set up the matplotlib figure
f, axes = plt.subplots(figsize = (10,7))
f.subplots_adjust(hspace= .8)
f.suptitle('Pilot3 Vs Beta2 QC',x=0.49,y=1.05, fontsize=14, fontweight='bold')
# Draw kappa heat map
sns.heatmap(kappa_out,vmin=0,vmax=1,cmap="YlGnBu",
square=True,
annot=True,
linewidths=.5,
cbar_kws={"shrink": .9,"label": "Cohen's Kappa"},
ax=axes)
axes.set_yticks([x+0.5 for x in range(len(kappa_df.columns))])
axes.set_yticklabels(kappa_df.columns,rotation=0)
axes.set_title("Cohen's Kappa matrix for {} images from \n 7 raters with different level of QC expertise \n and the Beta2 concensus ratings ".format(len(merge_ratings)),
pad=20,fontsize=12)
#axes.annotate('Low', xy=(-0.17, 0.97),xytext=(-0.2, -0), xycoords='axes fraction',
# arrowprops=dict(arrowstyle="fancy,tail_width=1.2,head_width=01",
# fc="0.7", ec="none",
# linewidth =2))
# Caption
pval = np.unique(Pval_store)[-1]
txt = '''
Fig1: Kappa matrix for 7 raters and one concensus from zooniverse rating. Each
of the 7 subject is ranked according to his level of expertise in QC of
brain images. Kappa's P-values range from {:.2g} to {:.2g} '''.format(Pval_store.min(), Pval_store.max())
f.text(0.1,-0.1,txt,fontsize=12)
#f.text(0.11,0.88,'High',fontsize=12)
#f.text(0.10,0.62,'Level of QC expertise',fontsize=12,rotation=90)
# Save figure
f.savefig('../reports/figures/Pilot3-vs-Beta2_qc.svg')
# -
from IPython.display import Image
Image(url= "https://i.stack.imgur.com/kYNd6.png" ,width=600, height=600)
# ### Comapre beta1 and 2 regards to expert raters
beta1 = pd.read_csv('../data/rating/Beta1_zooniverse_rating.csv')
beta1.head()
pilot2 = pd.read_csv('../data/rating/Pilot2_internal_rating-PB_YB.csv')
pilot2.head()
full_launch = pd.read_csv('../data/rating/all_experts_ratings.csv')
full_launch.head()
# Merge
dfs = [df_norm[['ID','concensus_NoExp']],beta1,pilot2,full_launch[['ID','Econ','Zcon']]]
merge_ratings = reduce(lambda left,right: pd.merge(left,right,how='inner',on='ID'), dfs).apply(lambda x: x.str.strip() if x.dtype == "object" else x)
merge_ratings.rename(columns={'concensus_NoExp':'Zcon_B2',''},inplace=True)
merge_ratings.head()
# Replace OK with 1 , Maybe with 2 and Fail with 3
merge_ratings.replace({'OK':1,'Maybe':2, 'Fail':3}, inplace=True)
merge_ratings.head()
# + language="R"
# suppressPackageStartupMessages(library(dplyr))
# #install.packages("irr")
# library(irr)
# -
# Percenteage of agrrement between raters with R package IRR
agree_ = merge_ratings.drop(['ID'],axis=1)
# %Rpush agree_
# agree_n = %R agree(agree_)
print(agree_n)
# +
# FDR correction
from statsmodels.sandbox.stats import multicomp as smi
def fdr_transf(mat,log10 = False):
'''compute fdr of a given matrix'''
row = mat.shape[0]
col = mat.shape[1]
flatt = mat.flatten()
fdr_2d = smi.multipletests(flatt, alpha=0.05, method='fdr_bh')[1]
if log10 == True:
fdr_2d = [-np.log10(ii) if ii != 0 else 50 for ii in fdr_2d ]
fdr_3d = np.reshape(fdr_2d,(row,col))
return fdr_3d
# -
# Kappa calculation
def kappa_score(k_df,log10 = False):
'''compute Kappa between diferent raters organized in dataframe'''
k_store = np.zeros((len(k_df.columns), len(k_df.columns)))
p_store = np.zeros((len(k_df.columns), len(k_df.columns)))
# %Rpush k_df
for user1_id, user1 in enumerate(k_df.columns):
for user2_id, user2 in enumerate(k_df.columns):
weight = np.unique(kappa_df[[user1,user2]])
# %Rpush user1_id user1 user2_id user2 weight
# kappaR = %R kappa2(k_df[,c(user1,user2)],weight)
# store the kappa
k_store[user1_id, user2_id] = [kappaR[x][0] for x in range(np.shape(kappaR)[0])][4]
p_store[user1_id, user2_id] = [kappaR[x][0] for x in range(np.shape(kappaR)[0])][-1]
# FDR Correction
p_store = fdr_transf(p_store,log10)
return k_store, p_store
# +
# Get Kappa score out of all different combination of ratings
kappa_df = merge_ratings.drop(['ID'],axis=1)
kappa_store, Pval_store = kappa_score(kappa_df)
mean_kap = np.mean(kappa_store[np.triu_indices(len(kappa_store),k=1)])
std_kap = np.std(kappa_store[np.triu_indices(len(kappa_store),k=1)])
print('Mean Kappa : {0:.2f} , std : {1:.2f}\n'.format(mean_kap, std_kap))
#calculte the over all kappa values of all ratings
# %Rpush kappa_df
# fleiss_kappa = %R kappam.fleiss(kappa_df,c(0,1,2))
print(fleiss_kappa)
# +
# Plot kappa matrix
kappa_out = pd.DataFrame(kappa_store,
index=kappa_df.columns.get_values(),
columns=kappa_df.columns.get_values())
# Set up the matplotlib figure
f, axes = plt.subplots(figsize = (10,7))
f.subplots_adjust(hspace= .8)
f.suptitle('Pilot3 Vs Beta2 QC',x=0.49,y=1.05, fontsize=14, fontweight='bold')
# Draw kappa heat map
sns.heatmap(kappa_out,vmin=0,vmax=1,cmap="YlGnBu",
square=True,
annot=True,
linewidths=.5,
cbar_kws={"shrink": .9,"label": "Cohen's Kappa"},
ax=axes)
axes.set_yticks([x+0.5 for x in range(len(kappa_df.columns))])
axes.set_yticklabels(kappa_df.columns,rotation=0)
axes.set_title("Cohen's Kappa matrix for {} images from \n 7 raters with different level of QC expertise \n and the Beta2 concensus ratings ".format(len(merge_ratings)),
pad=20,fontsize=12)
#axes.annotate('Low', xy=(-0.17, 0.97),xytext=(-0.2, -0), xycoords='axes fraction',
# arrowprops=dict(arrowstyle="fancy,tail_width=1.2,head_width=01",
# fc="0.7", ec="none",
# linewidth =2))
# Caption
pval = np.unique(Pval_store)[-1]
txt = '''
Fig1: Kappa matrix for 7 raters and one concensus from zooniverse rating. Each
of the 7 subject is ranked according to his level of expertise in QC of
brain images. Kappa's P-values range from {:.2g} to {:.2g} '''.format(Pval_store.min(), Pval_store.max())
f.text(0.1,-0.1,txt,fontsize=12)
#f.text(0.11,0.88,'High',fontsize=12)
#f.text(0.10,0.62,'Level of QC expertise',fontsize=12,rotation=90)
# Save figure
f.savefig('../reports/figures/Pilot3-vs-Beta1-and-2_qc.svg')
# -
# ### Report tagging from Beta1 raters
# +
# output markings from classifications
clist=[]
for index, c in classi_unique.iterrows():
if c['n_tagging'] > 0:
for q in c.annotations[0]['value']:
clist.append({'ID':c.ID, 'workflow_name':c.workflow_name,'user_name':c.user_name, 'rating':c.rating,'imgnm':c.imgnm,
'x':q['x'], 'y':np.round(q['y']).astype(int), 'r':'1.5','n_tagging':c.n_tagging ,'frame':q['frame']})
else:
clist.append({'ID':c.ID, 'workflow_name':c.workflow_name, 'user_name':c.user_name,'rating':c.rating,'imgnm':c.imgnm,
'x':float('nan'), 'y':float('nan'), 'r':float('nan'),'n_tagging':c.n_tagging ,'frame':'1'})
col_order=['ID','workflow_name','user_name','rating','x','y','r','n_tagging','imgnm','frame']
out_tag = pd.DataFrame(clist)[col_order]
out_tag.user_name.replace({'simexp':'PB','Yassinebha':'YB'},inplace=True)
out_tag.head()
# -
# Extract unique IDs for each image
ids_imgnm = np.reshape([out_tag.ID.unique(),out_tag.imgnm.unique()],(2,np.shape(out_tag.ID.unique())[0]))
df_ids_imgnm = pd.DataFrame(np.sort(ids_imgnm.T, axis=0),columns=['ID', 'imgnm'])
df_ids_imgnm.head()
# +
# Create custom color map
from matplotlib.colors import LinearSegmentedColormap , ListedColormap
from PIL import Image
def _cmap_from_image_path(img_path):
img = Image.open(img_path)
img = img.resize((256, img.height))
colours = (img.getpixel((x, 0)) for x in range(256))
colours = [(r/255, g/255, b/255, a/255) for (r, g, b, a) in colours]
return colours,LinearSegmentedColormap.from_list('from_image', colours)
coll,a=_cmap_from_image_path('../data/Misc/custom_ColBar.png')
#invert color map
coll_r = ListedColormap(coll[::-1])
# -
# set color different for each rater
list_tagger = out_tag.user_name.unique()
colors_tagger = sns.color_palette("Set2", len(list_tagger))
# ### Plot heat map for all tagging
# +
from heatmappy import Heatmapper
from PIL import Image
patches=list()
for ind, row in df_ids_imgnm.iterrows():
out_tmp = out_tag[out_tag['ID'] == row.ID]
patches.append([(row.x,row.y) for ind,row in out_tmp.iterrows()])
patches = [x for x in sum(patches,[]) if str(x[0]) != 'nan']
# plot heat map on the template
f, axes = plt.subplots(1, 1,figsize = (10,14))
f.subplots_adjust(hspace= .8)
f.suptitle('Beta2 Zooniverse QC',x=0.49,y=.83, fontsize=14, fontweight='bold')
img = Image.open('../data/Misc/template_stereotaxic_v3.png')
axes.set_title('Tagging from all Beta2 raters')
heatmapper = Heatmapper(opacity=0.5,
point_diameter=15,
point_strength = 0.5,
colours=a)
heatmap= heatmapper.heatmap_on_img(patches, img)
im = axes.imshow(heatmap,cmap=coll_r)
axes.set_yticklabels([])
axes.set_xticklabels([])
cbar = plt.colorbar(im, orientation='vertical', ticks=[0, 125, 255],fraction=0.046, pad=0.04,ax=axes)
cbar.ax.set_yticklabels(['0', '5', '> 10'])
img.close()
heatmap.close()
f.savefig('../reports/figures/Beta2_qc_heatmap_tags.svg')
# -
|
notebooks/Beta2_zooniverse_QC_results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## Compare candidate-entities_v1 vs candidate-entities_v3
# candidate-entities_v3 includes some spelling mistakes added by <NAME> to the synonyms column
# import libraries
import pandas as pd
# import files
df_v1 = pd.read_csv('../../electionBot_sBox/candidates-Entities.csv')
df_v3 = pd.read_csv('../../electionBot_sBox/candidates-Entities_v3.csv')
df_v1.head()
df_v3.head()
# reimport v3 with delimiter
df_v3 = pd.read_csv('../../electionBot_sBox/candidates-Entities_v3.csv', delimiter=';')
df_v3.head()
df_v1.info()
df_v3.info()
# rexport v3 with correct format
df_v3.to_csv('../../electionBot_sBox/candidates-Entities_v3_edt.csv',index=False)
|
DATA_format_JSON/python_stuff/Comparing_candidate_entitiesv1_v3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %run ../common-imports.ipynb
# # DBSCAN common utilities
# +
from matplotlib.axes import Axes
def dbscan_cluster_plot(eps: float, nPts:int, data:pd.DataFrame, ax:Axes) -> DBSCAN:
data['x1'] = data.iloc[:,0]
data['x2'] = data.iloc[:,1]
X = data[['x1', 'x2']]
clusterer = DBSCAN(eps=eps, min_samples=nPts)
ŷ = clusterer.fit_predict(X)
data['yhat'] = ŷ
unique_labels= data.yhat.unique()
n_clusters = len(np.unique(clusterer.labels_)) - (1 if -1 in clusterer.labels_ else 0)
# First, separate the clustered-points from outliers
clusters = data[data.yhat != -1]
outliers = data[data.yhat == -1]
colors = [plt.cm.Spectral(each) for each in np.linspace(0,1, len(unique_labels))]
# plot clusters
ax.scatter(clusters.x1,
clusters.x2,
c=clusters.yhat,
s=150,
cmap='Spectral',
alpha=0.5, edgecolor='black');
# plot outliers as dark black points
ax.scatter(outliers.x1,
outliers.x2,
c='black',
s=50,
cmap='Paired',
alpha=0.3);
ax.set_title(f'eps:{eps}, nPts: {nPts}, clusters: {n_clusters}')
return clusterer
# -
from typing import List
def dbscan_cluster(epsilons: List[float], neighbors:List[int], data:pd.DataFrame) -> pd.DataFrame:
is_labeled = 'label' in data.columns
columns = ['epsilon',
'nPts',
'clusters',
'silhouette score']
if is_labeled:
columns.extend(['homogeniety',
'completeness',
'v-measure',
'adjusted rand index',
'adjusted mutual information'])
# Create an empty dataframe to store the clustering quality metrics
quality = pd.DataFrame(columns=columns)
# Subplots
row_count = len(neighbors)
col_count = len(epsilons)
fig, axes = plt.subplots(row_count, col_count, figsize=(8*row_count,5*col_count))
# Cluster for each combination of the hyper-parameters
for row, nPts in enumerate(neighbors):
for col, ϵ in enumerate(epsilons):
clusterer = dbscan_cluster_plot(ϵ, nPts, data, axes[row][col])
n_clusters = len(np.unique(clusterer.labels_)) - (1 if -1 in clusterer.labels_ else 0)
silhouette = 0 if len(np.unique(clusterer.labels_)) == 1 else metrics.silhouette_score(data, clusterer.labels_)
values = [ϵ, nPts, n_clusters, silhouette]
if is_labeled:
y = data.label
ŷ = clusterer.labels_
values.extend ([metrics.homogeneity_score(y, ŷ),
metrics.completeness_score(y, ŷ),
metrics.v_measure_score(y, ŷ),
metrics.adjusted_rand_score(y, ŷ),
metrics.adjusted_mutual_info_score(y, ŷ),])
quality.loc[len(quality.index)] = values
plt.suptitle(r'\textbf{\Huge Sensitivity of DBSCAN to choices of the hyperparameters}');
plt.tight_layout()
return quality
|
notebook/cluster/dbscan_common.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="vhzqCR6gjMv9" colab_type="text"
# **Coding Challenge** #** 2** - Collaborative Filtering
# + [markdown] id="hJZzrf7qg6FU" colab_type="text"
# **Coding Challenge:** **Context**
#
# With collaborative filtering, an application can find users with similar tastes and can look at ietms they like and combine them to create a ranked list of suggestions which is known as user based recommendation. Or can also find items which are similar to each other and then suggest the items to users based on their past purchases which is known as item based recommendation. The first step in this technique is to find users with similar tastes or items which share similarity.
#
# There are various similarity models like** Cosine Similarity, Euclidean Distance Similarity and Pearson Correlation Similarity** which can be used to find similarity between users or items.
# + [markdown] id="E3zcH1mQppxI" colab_type="text"
# In this coding challenge, you will go through the process of identifying users that are similar (i.e. User Similarity) and items that are similar (i.e. "Item Similarity")
#
# **User Similarity:**
#
# **1a)** Compute "User Similarity" based on cosine similarity coefficient (fyi, the other commonly used similarity coefficients are Pearson Correlation Coefficient and Euclidean)
#
# **1b)** Based on the cosine similarity coefficient, identify 2 users who are similar and then discover common movie names that have been rated by the 2 users; examine how the similar users have rated the movies
#
# **Item Similarity:**
#
# **2a) ** Compute "Item Similarity" based on the Pearson Correlation Similarity Coefficient
#
# **2b)** Pick 2 movies and find movies that are similar to the movies you have picked
#
# **Challenges:**
#
# **3)** According to you, do you foresee any issue(s) associated with Collaborative Filtering?
#
# **Dataset: ** For the purposes of this challenge, we will leverage the data set accessible via https://grouplens.org/datasets/movielens/
#
# The data set is posted under the section: ***recommended for education and development*** and we will stick to the small version of the data set with 100,000 ratings
# + id="Ghz8ZdKKgWSd" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
import zipfile
import pandas as pd
import numpy as np
from scipy.stats import pearsonr
from scipy.spatial.distance import pdist, squareform
# + id="FvWUQasskD0a" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 204} outputId="5c496843-87ac-485e-ebc1-7dcd71711f0a" executionInfo={"status": "ok", "timestamp": 1527291536243, "user_tz": 420, "elapsed": 1834, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
# ! wget 'http://files.grouplens.org/datasets/movielens/ml-latest-small.zip'
# + id="4N4flwoOjvMi" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
folder = zipfile.ZipFile('ml-latest-small.zip')
# + id="kJImatYTkZCZ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 119} outputId="0f516f53-14dd-4c97-b9d3-ec54b981aaaa" executionInfo={"status": "ok", "timestamp": 1527291541284, "user_tz": 420, "elapsed": 542, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
folder.infolist()
# + id="KwSzpR2JgyYI" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
ratings = pd.read_csv(folder.open('ml-latest-small/ratings.csv'))
movies = pd.read_csv(folder.open('ml-latest-small/movies.csv'))
# + id="mD4mNlK0k0tK" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 391} outputId="2ef6890a-c6e2-4711-a9a9-6de36b95843c" executionInfo={"status": "ok", "timestamp": 1527291546967, "user_tz": 420, "elapsed": 570, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
display(ratings.head())
display(movies.head())
# + [markdown] id="Nu2wj2E7lPkC" colab_type="text"
# ## User Similarity
# + id="U4U7EQMelK_S" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 312} outputId="e2fccaea-c2f8-4a7f-bf33-d1b205628bf1" executionInfo={"status": "ok", "timestamp": 1527291549746, "user_tz": 420, "elapsed": 721, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
ratings_pivot = pd.pivot_table(ratings.drop('timestamp', axis=1),
index='userId', columns='movieId',
aggfunc=np.max).fillna(0)
print(ratings_pivot.shape)
ratings_pivot.head()
# + id="8jgOZ-fDlfa8" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 238} outputId="1ac75483-c80c-40fe-c827-bc42e6945642" executionInfo={"status": "ok", "timestamp": 1527291982501, "user_tz": 420, "elapsed": 3881, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
distances = pdist(ratings_pivot.as_matrix(), 'cosine')
squareform(distances)
# + [markdown] id="V1fnnS3_sNyp" colab_type="text"
# Since pdist calculates $1 - \frac{u\cdot v}{|u||v|}$ instead of cosine similarity, I will have to subtract the result from 1.
# + id="ScHoH2sDsgfV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 255} outputId="7391bf69-4002-483c-efc0-b894ffbe4fa9" executionInfo={"status": "ok", "timestamp": 1527292767427, "user_tz": 420, "elapsed": 566, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
similarities = squareform(1-distances)
print(similarities.shape)
similarities
# + id="5XKNoj3wssNp" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} outputId="3be6241d-0c03-4ed2-b4de-bf676daa6ab2" executionInfo={"status": "ok", "timestamp": 1527292769633, "user_tz": 420, "elapsed": 608, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
ix = np.unravel_index(np.argmax(similarities), similarities.shape)
print(ix)
print(similarities[ix])
# + [markdown] id="wHcwUoA1oRE7" colab_type="text"
# Users 151 and 369 appear to be similar, with a cosine similarity of 0.84
# + id="V95DddNpoitx" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1616} outputId="8e852cbd-784f-40c4-e577-f8d1d3170355" executionInfo={"status": "ok", "timestamp": 1527292771126, "user_tz": 420, "elapsed": 428, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
print('Common movies rated')
display(ratings_pivot.iloc[[150, 368], :].T[(ratings_pivot.iloc[150]>0)
& (ratings_pivot.iloc[368]>0)])
# + [markdown] id="Izw4Ziyvtz7E" colab_type="text"
# ## Item Similarity
# + id="h5aVR4lBt3tw" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 238} outputId="faa18ce5-6bf5-4f32-cac1-57e3da07c6a3" executionInfo={"status": "ok", "timestamp": 1527292825344, "user_tz": 420, "elapsed": 37381, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
correlations = squareform(1-pdist(ratings_pivot.as_matrix().T, 'correlation'))
correlations
# + id="Ob6w02Jiwq9m" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="d3e6c57d-ec27-4d71-ae3d-90279a8eb69b" executionInfo={"status": "ok", "timestamp": 1527293151039, "user_tz": 420, "elapsed": 466, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
np.argsort(correlations[0])[::-1]
# + id="rrBBkps-w2-3" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} outputId="e53ab141-1c02-4d3d-9ffc-753697f7dc2c" executionInfo={"status": "ok", "timestamp": 1527293162018, "user_tz": 420, "elapsed": 1164, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
correlations[0][np.argsort(correlations[0])[::-1]]
# + id="cnXPtLu-wT4j" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 204} outputId="a115e1c0-eda3-4bfd-d431-04396d48a5cd" executionInfo={"status": "ok", "timestamp": 1527292977805, "user_tz": 420, "elapsed": 910, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
movies.head()
# + [markdown] id="4Qcb65_NwXVm" colab_type="text"
# I will see which movies correlate the most with "Toy Story" and "Jumanji."
# + id="0JQdbYDoyI3S" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="94c74b64-5aa2-4a9a-ae9f-22a7534d666e" executionInfo={"status": "ok", "timestamp": 1527293548883, "user_tz": 420, "elapsed": 436, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
np.argsort(correlations[1])[::-1][:5] + 1
# + id="cCxA_LRdwV6M" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def most_correlated_movies(movieId, corr_matrix, n=5):
ix = movieId - 1
return np.argsort(correlations[ix])[::-1][:n] + 1
# + id="JpWUq_0zxUMR" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 142} outputId="929a601b-0fdc-4d20-9939-3f5d44b40beb" executionInfo={"status": "ok", "timestamp": 1527293433170, "user_tz": 420, "elapsed": 435, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
toy_story_similar = most_correlated_movies(1, correlations)
movies[movies['movieId'].isin(toy_story_similar)]
# + id="5z99NK_-x68c" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 142} outputId="e26655b7-9df3-472d-80e2-f4d76e399425" executionInfo={"status": "ok", "timestamp": 1527293570385, "user_tz": 420, "elapsed": 432, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
jumanji_similar = most_correlated_movies(2, correlations)
movies[movies['movieId'].isin(jumanji_similar)]
# + [markdown] id="WtZ4I523yw6H" colab_type="text"
# It seems that there are less movies in DataFrame matching IDs to titles, so not every movie ID found by the `most_correlated_movies` function correponds to a named entry.
# + id="ddaDvEYKysBx" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="a59a4145-7658-4def-8e0e-16a100c9c8f2" executionInfo={"status": "ok", "timestamp": 1527293594043, "user_tz": 420, "elapsed": 591, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-BMlr5I5Dhow/AAAAAAAAAAI/AAAAAAAAABc/XW4PF5A8K2Q/s50-c-k-no/photo.jpg", "userId": "116545933704048584401"}}
movies.shape
|
Week 08 Unsupervised Learning/Code Challenges/Day 4 Collaborative Filtering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: covid19_env
# language: python
# name: covid19_env
# ---
# # USAID Sites and Geospatial Intelligence
# This notebook covers the creating of geospatial data
# +
import os
import pandas as pd
import numpy as np
# +
#os.mkdir("geospatial_data")
# -
# %matplotlib inline
# Load the site data provided by USAID
site_data = pd.read_csv("final_data/service_delivery_site_data.csv")
site_data.head()
import geopandas as gpd
# +
# Use the data provided by UN Geospatial Repsitory
gdf = gpd.read_file('civ_admbnda_adm3_cntig_ocha_itos_20180706/civ_admbnda_adm3_cntig_ocha_itos_20180706.shp')
gdf.head()
# -
# ### Lets start trying to find a shared column to match on
districts = site_data['site_district'].unique()
print(len(districts))
districts.sort()
districts
site_data.head()
# 'ABOBO-EST' is a neighborhood in Abidjan
#
# Match city to district then aggregate the district level.
#
# Need to do some manual analysis to figure this whole matching up.
# #### Data Processing
# - Observe there is a missing I so you think perhaps the two will match on ADM2_PCODE but this is a mistake. **There is no relationship between the two**
#
# ```
# #site_data['site_code'].head()
#
# #gdf['ADM2_PCODE'].head()
#
# # Insert String
# #ins_char = lambda x: x[0:1]+"I"+x[1:]
# #site_data['ADM2_PCODE'] = site_data['site_code'].apply(ins_char)
# ```
# ## Data Processing
# ### The codes are not matching up between the two dataframes
# - On inspection we can see there is an I missing, lets try tp add that and see if that fixes thing
# +
print("Num sites: ", len(site_data))
print("Num boundary shapes at ADM3: ",len(gdf))
# -
# ## Geospatial Data
#
#
# ### Regional Data
#
# - Wikipedia data
# - Data from UN Geospatial Data Repository
# #### Notes
# https://en.wikipedia.org/wiki/Subdivisions_of_Ivory_Coast
# https://www.youtube.com/watch?v=6pYorKr3XFQ&ab_channel=AlJazeeraEnglish
# https://www.youtube.com/watch?v=O1_wpzPX7C8&ab_channel=FRANCE24English
# https://fr.wikipedia.org/wiki/R%C3%A9gions_de_C%C3%B4te_d%27Ivoire
# +
from io import StringIO
# Taken from this Wikipedia Page
# https://fr.wikipedia.org/wiki/R%C3%A9gions_de_C%C3%B4te_d%27Ivoire
wikipedia_table = """
District Chef-lieu de district Région Chef-lieu de région
Zanzan Bondoukou Bounkani Bouna
Zanzan Bondoukou Gontougo Bondoukou
Yamoussoukro (district autonome) — — —
Woroba Séguéla Béré Mankono
Woroba Séguéla Bafing Touba
Woroba Séguéla Worodougou Séguéla
Vallée du Bandama Bouaké Hambol Katiola
Vallée du Bandama Bouaké Gbêkê Bouaké
Savanes Korhogo Poro Korhogo
Savanes Korhogo Tchologo Ferkessédougou
Savanes Korhogo Bagoué Boundiali
Sassandra-Marahoué Daloa Haut-Sassandra Daloa
Sassandra-Marahoué Daloa Marahoué Bouaflé
Montagnes Man Tonkpi Man
Montagnes Man Cavally Guiglo
Montagnes Man Guémon Duékoué
Lagunes Dabou Agnéby-Tiassa Agboville
Lagunes Dabou Mé Adzopé
Lagunes Dabou Grands Ponts Dabou
Lacs Dimbokro N’Zi Dimbokro
Lacs Dimbokro Iffou Daoukro
Lacs Dimbokro Bélier Toumodi
Lacs Dimbokro Moronou Bongouanou
Gôh-Djiboua Gagnoa Gôh Gagnoa
Gôh-Djiboua Gagnoa Lôh-Djiboua Divo
Denguélé Odienné Folon Minignan
Denguélé Odienné Kabadougou Odienné
Comoé Abengourou Indénié-Djuablin Abengourou
Comoé Abengourou Sud-Comoé Aboisso
Bas-Sassandra San-Pédro Nawa Soubré
Bas-Sassandra San-Pédro San-Pédro San-Pédro
Bas-Sassandra San-Pédro Gbôklé Sassandra
Abidjan (district autonome) — — —"""
wiki_region_mappings = pd.read_csv(StringIO(wikipedia_table),sep="\t")
# -
gdf.groupby(['ADM1_FR','ADM2_FR']).size().sort_values(ascending =False)
# #### Site Data
site_data.head()
site_data.groupby(['site_region','site_district'])['site_code'].size().sort_values(ascending=False).head(10)
# #### Analysis of Site Data Boundary Structure
# - Data is organized by Region and then by Site, this is contrary to the way that Cote d'Ivoire organizes itself which is by:
# 1. District
# 2. Region
# 3. Department
# 4. Village
# 5. Commune
#
# - The USAID dataset seems to be presented as
# 1. Region: regions or multiple regions combined under one administrative boundary
# 2. District: regions and departments
# ### Fuzz-Matching of regional names in the dataset
#
# - Calculate the character lexical similarity in the strings to determine the best matches.
# +
from fuzzywuzzy import fuzz
from fuzzywuzzy import process
def get_fuzzy_match_results(ref_array,custom_array):
# Create a dictionary to hold all the string matching calculations
custom_mapping = {}
# Lambda functions to extract the match name from the tuple
get_match_name = lambda x: match_dict[x][0]
# Lambda functions to extract the match score from the tuple
get_match_score = lambda x: match_dict[x][1]
# Create_Reference_table
reference_table = pd.DataFrame({'ref':ref_array})
# Iterate over every commune name in the reference table
for custom_label in custom_array:
# Skip values if not string
if type(custom_label) == str:
ffuzzy_match = lambda x : fuzz.partial_ratio(custom_label,x)
reference_table[custom_label] = reference_table['ref'].apply(ffuzzy_match)
#reference_table[custom_label] = fuzzy_ratios
ref_max_value = reference_table[custom_label].max()
# Identify the record that has the highest score with the provided custom_label
matching_recs = reference_table.loc[reference_table[custom_label]==ref_max_value]
# If there are two communes that have an equal score, select the first value
if len(matching_recs)>1:
#print("Multiple matches: ",custom_label)
match_site_name = matching_recs['ref'].values[0]
else:
match_site_name = matching_recs['ref'].values[0]
# Update the match_dict with a tuple to store the final string and its corresponding score
custom_mapping[custom_label] = {'est_site_name':match_site_name,'fratio':ref_max_value}
else:
custom_mapping[custom_label] = None
mapping_df = pd.DataFrame(custom_mapping).transpose()
ref_df = pd.DataFrame(reference_table).set_index('ref')
return mapping_df,ref_df
# -
# ### Create a series of vectors to perform fuzzy wuzzy matching to create mappings
import seaborn as sns; sns.set()
# +
## USAID
# Isolate USAID Region values
usaid_civ_site_region = site_data['site_region'].str.upper().unique()
#print(len(usaid_civ_site_region))
#print(usaid_civ_site_region)
# Isolate USAID District values
usaid_civ_site_district = site_data['site_district'].str.upper().unique()
#print(len(usaid_civ_site_district))
#print(usaid_civ_site_district)
## Wikipedia
# Isolate Wikipedia Region labels
wiki_admn_region = wiki_region_mappings['Région'].str.upper().unique()
#print(len(wiki_admn_region))
#print(wiki_admn_region)
# Isolate Wikipedia Department labels
wiki_admn_dept = wiki_region_mappings['Chef-lieu de région'].str.upper().unique()
print(len(wiki_admn_dept))
#print(wiki_admn_dept)
## UN Geospatial
# Isolate UN ADM1_FR labels
geospatial_admn_1 = gdf['ADM1_FR'].str.upper().unique()
#print(len(geospatial_admn_1))
#print(geospatial_admn_1)
# Isolate UN ADM3_FR labels
geospatial_admn_2 = gdf['ADM2_FR'].str.upper().unique()
print(len(geospatial_admn_2))
#print(geospatial_admn_2)
# Isolate UN ADM4_FR labels
geospatial_admn_3 = gdf['ADM3_FR'].str.upper().unique()
print(len(geospatial_admn_3))
#print(geospatial_admn_3)
# -
import matplotlib.pyplot as plt
def make_fuzzy_matching_evaluation(ref,labels,ref_name='x',label_name='y'):
mapping ,ref_table = get_fuzzy_match_results(ref,labels)
mapping['fratio'] = pd.to_numeric(mapping['fratio'])
plt.subplots(figsize=(8,6))
sns.heatmap(ref_table_1)
plt.title(f"{ref_name} - {len(ref)} vs {label_name} - {len(labels)}",fontdict={'fontsize':20})
plt.show()
print(mapping.fratio.describe())
return mapping ,ref_table
wu_reg_reg_map, wu_reg_reg_tbl = make_fuzzy_matching_evaluation(wiki_admn_region,usaid_civ_site_region,'Wikipedia-Regions','USAID-Regions')
# #### Wikipedia: Spatial-Regions vs USAID-Regions
# - First we capitalize the results and there started to be several more higher fuzzy ratio scores calculated. Initially there were very weak results 'Me' was the highest match for all of the initial values.
#
#
# *Lets try Wikipedia Regions vs USAID - districts....*
#
# +
wu_reg_dis_map, wu_reg_dis_tbl = make_fuzzy_matching_evaluation(
wiki_admn_region,
usaid_civ_site_district,'Wikipedia-Regions','USAID-Districts')
# -
# #### Results: Wikipedia-Regions vs USAID-Regions
# - Great results startings to see many matches over 80
#
#
# *Lets try Wikipedia-departments_ with USAID-regions*
# +
wu_dep_dis_map, wu_dep_dis_tbl = make_fuzzy_matching_evaluation(
wiki_admn_dept,
usaid_civ_site_district,'Wikipedia-Departments','USAID-Districts')
# -
# ## Matching with the geospatial data and making custom maps
# #### ADM1_FR
# +
# Compare ADM1_FR and USAID's Region Codes
ug_reg_dis_map, ug_reg_dis_tbl = make_fuzzy_matching_evaluation(
usaid_civ_site_region,
geospatial_admn_1,'USAID-Regions','UN Geospatial ADM1_FR')
# +
# Using the matches from fuzzy model as a base with ADM1_FR and USAID Region
# Region vs Region
# This will drop one region from the reference index
# Drop Yamaoussoukora
civ_strong_matches = ug_reg_dis_map[ug_reg_dis_map.fratio>50]
print(len(civ_strong_matches))
print(civ_strong_matches)
civ_region_adm1_mapping = {index:data['est_site_name'] for index, data in civ_strong_matches.iterrows()}
# -
# ### Lets compare ADM2_Fr and USAID District
#
# In the past example we used 50 as our cut-off criteria now as we have stronger matches we will increase it to 75
#
# #### ADM2_FR
# +
# Compare ADM1_FR and USAID's Region Codes
gu_dep_dis_map, gu_dep_dis_tbl = make_fuzzy_matching_evaluation(
geospatial_admn_2,
usaid_civ_site_district,'UN Geospatial ADM1_FR','USAID-Districts')
strong_matches = gu_dep_dis_map[gu_dep_dis_map.fratio>50]
print("Strong matches")
print(len(strong_matches))
print(strong_matches.sort_values(by='fratio',ascending=False).head(10))
weak_matches = gu_dep_dis_map[gu_dep_dis_map.fratio<70]
print("Weak matches")
print(len(weak_matches))
print(weak_matches)
# Drop values with a bad mapping
# Using the weak matches we can identify the worst performing matches
# We want to keep the last records though, it seems the accent is being interpretted poorly
drop_indices = weak_matches.index[:-1]
civ_dist_adm2_mapping = {index:data['est_site_name'] for index, data in gu_dep_dis_map.drop(drop_indices).iterrows()}
# -
# #### ADM3_FR
# #### Lets compare ADM3_Fr and USAID District
#
# I have already pruned the matches by evaluating weak matches.
# +
geospatial_admn_3 = gdf['ADM3_FR'].str.upper().unique()
print(len(geospatial_admn_3))
#print(civ_admn_dept)
print(len(usaid_civ_site_district))
site_dist_geo_admn3_res,site_dist_geo_admn3_tbl = get_fuzzy_match(geospatial_admn_3,usaid_civ_site_district)
sns.heatmap(site_dist_geo_admn3_tbl.set_index('ref'))
# Compare ADM1_FR and USAID's Region Codes
gu_dep3_dis_map, gu_dep3_dis_tbl = make_fuzzy_matching_evaluation(
geospatial_admn_3,
usaid_civ_site_district,'UN Geospatial ADM1_FR','USAID-Districts')
strong_matches = gu_dep3_dis_map[gu_dep3_dis_map.fratio>50]
print("Strong matches")
print(len(strong_matches))
print(strong_matches.sort_values(by='fratio',ascending=False).head(10))
weak_matches = gu_dep3_dis_map[gu_dep3_dis_map.fratio<70]
print("Weak matches")
print(len(weak_matches))
print(weak_matches)
# Drop values with a bad mapping
# Using the weak matches we can identify the worst performing matches
drop_indices = weak_matches.index[:-1]
civ_dist_adm3_mapping = {index:data['est_site_name'] for index, data in gu_dep3_dis_map.drop(drop_indices).iterrows()}
# -
# #### Lets Apply the mappings
#
# - Starting from region, create a mapping to map each value in the USAID data set with our pruned mappings
site_data['adm3_fr'] = site_data['site_district'].map(civ_dist_adm3_mapping)
site_data['adm2_fr'] = site_data['site_district'].map(civ_dist_adm2_mapping)
# ##### Apply the mappings to the geospatial data
# - Because of how the USAID regions incorporated multiple existing geospatial boundaries, I applied the USAID regional mapping onto the existing ADM1_FR names in order to gain access to map projections with those groupings.
# +
gdf['usaid_admin_region'] = gdf['ADM1_FR'].str.upper().map(civ_region_adm1_mapping)
gdf.to_file("geospatial_data/Custom_CIV.shp")
gdf['usaid_admin_region'].head()
|
Connecting Site and Geospatial data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # パーセプトロン
# ## 簡単な実装
# 入力が2つの場合の実装としてANDゲートのイメージ
# |x1|x2|y|
# |:--|:--|:--|
# |0|0|0|
# |1|0|1|
# |0|1|1|
# |1|1|1|
def AND(x1, x2):
w1, w2, theta = 0.5, 0.5, 0.7
tmp = x1 * w1 + x2 * w2
if tmp <= theta:
return 0
elif tmp > theta:
return 1
AND(0, 0)
AND(1, 1)
# # 重みとバイアス
import numpy as np
x = np.array([0, 1])
w = np.array([0.5, 0.5])
b = -0.7
w * x
np.sum(w * x)
np.sum(w * x) + b
def AND(x1, x2):
x = np.array([x1, x2])
w = np.array([0.5, 0.5])
b = -0.7
tmp = np.sum(w*x) + b
if tmp <= 0:
return 0
elif tmp > 0:
return 1
AND(1, 0)
AND(1, 1)
# ## NANDとORゲート
# NANDは重みとバイアスの符号を変えただけ
# ORは重みとバイアスの値を変えるだけ
# |x1|x2|y|
# |:--|:--|:--|
# |0|0|1|
# |1|0|0|
# |0|1|0|
# |1|1|0|
def NAND(x1, x2):
x = np.array([x1, x2])
w = np.array([-0.5, -0.5]) #重み
b = 0.7 #バイアス
tmp = np.sum(w*x) + b
if tmp <= 0:
return 0
elif tmp > 0:
return 1
# |x1|x2|y|
# |:--|:--|:--|
# |0|0|0|
# |1|0|0|
# |0|1|0|
# |1|1|1|
def OR(x1, x2):
x = np.array([x1, x2])
w = np.array([0.5, 0.5])
b = -0.2 #ここだけ
tmp = np.sum(w*x) + b
if tmp <= 0:
return 0
elif tmp > 0:
return 1
# ### XORゲート
# パーセプトロン(一本の直線で分離できない(線形ではむり))
# |x1|x2|y|
# |:--|:--|:--|
# |0|0|0|
# |1|0|1|
# |0|1|1|
# |1|1|0|
# +
import numpy as np
import matplotlib.pyplot as plt
x = np.arange(-2, 2, 0.1)
y1 = -x + 0.6
y2 = -x + 1.4
fig = plt.figure()
ax = fig.add_subplot(111)
ax.set_xlim(0, 1)
ax.set_ylim(0, 1)
# x軸に補助目盛線を設定
ax.grid(which = "major", axis = "x", color = "blue", alpha = 0.5,
linestyle = "-", linewidth = 1)
# y軸に目盛線を設定
ax.grid(which = "major", axis = "y", color = "blue", alpha = 0.5,
linestyle = "-", linewidth = 1)
plt.plot(x,y1, label="y1")
plt.plot(x,y2, linestyle = "--", label="y2")
plt.xlabel("x")
plt.ylabel("y")
plt.title("XOR")
plt.figlegend()
plt.show()
# -
# |x1|x2|S1|S2|y|
# |:--|:--|:--|:--|:--|
# |0|0|1|0|0|
# |1|0|1|1|1|
# |0|1|1|1|1|
# |1|1|0|1|0|
def XOR(x1, x2):
s1 = NAND(x1, x2)
s2 = OR(x1, x2)
y = AND(s1, s2)
return y
XOR(0, 0)
XOR(1, 0)
XOR(0, 1)
XOR(1, 1)
|
day003/perceptron.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/prikmm/MLprojects/blob/main/notebooks/ShakespeareanText_Generator.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="U-Q4awBfreku"
import tensorflow as tf
from tensorflow import keras
import numpy as np
# + id="xmJm31stsC4u"
shakespeare_url = "https://homl.info/shakespeare"
filepath = keras.utils.get_file("shakespeare.txt", shakespeare_url)
with open(filepath) as f:
shakespeare_text = f.read()
# + [markdown] id="wseyjkDluPmu"
# ## Encoding using Tokenizer:
# + colab={"base_uri": "https://localhost:8080/"} id="wuvMnbnJ1k_y" outputId="0b63f44a-568a-4a43-eaa3-fdacccf05062"
print(shakespeare_text[:148])
# + id="9d7mHdpZsaGh"
tokenizer = keras.preprocessing.text.Tokenizer(char_level=True)
tokenizer.fit_on_texts(shakespeare_text)
# + colab={"base_uri": "https://localhost:8080/"} id="bwxMumG9s8UN" outputId="fadc5153-1b22-4d31-d186-34464a5cbf69"
tokenizer.texts_to_sequences(["HIIII", "hiiii", "Hey there"])
# + colab={"base_uri": "https://localhost:8080/"} id="z19QvVZwtMjl" outputId="7f676929-d03b-44d8-a900-91aa8eed52b3"
tokenizer.sequences_to_texts([[20, 6, 9, 3, 4]])
# + colab={"base_uri": "https://localhost:8080/"} id="7nA1jHXytXQh" outputId="6d9fca28-ad41-4a14-d31c-4eef1cb02ffe"
max_id = len(tokenizer.word_index) # no.of distinct characters
dataset_size = tokenizer.document_count
print(max_id, dataset_size)
# + colab={"base_uri": "https://localhost:8080/"} id="5kJy832Qts6K" outputId="39b4efab-3d96-42dc-e352-b40dc2f39edc"
[encoded] = np.array(tokenizer.texts_to_sequences([shakespeare_text])) - 1
print(encoded)
# + [markdown] id="Fwg_GxSxt8IT"
# ## Splitting a Sequential Dataset:
# + colab={"base_uri": "https://localhost:8080/"} id="V9RX6hiAuXjN" outputId="56e0e835-9061-434f-c3ee-d7f642b3c561"
train_size = dataset_size * 90 //100
dataset = tf.data.Dataset.from_tensor_slices(encoded[:train_size])
for item in dataset.take(10):
print(item)
# + colab={"base_uri": "https://localhost:8080/"} id="EqlTkwTvv8vi" outputId="60f26507-5dd7-458d-ea5a-56a437a64a14"
n_steps = 100
window_length = n_steps + 1
dataset = dataset.window(window_length, shift=1, drop_remainder=True)
for item in dataset.take(1):
print(item)
# + colab={"base_uri": "https://localhost:8080/"} id="NMvrM9XsyoAz" outputId="925e299d-971d-4ae7-9618-5b0f39bd5435"
dataset = dataset.flat_map(lambda window: window.batch(window_length))
for item in dataset.take(1):
print(item)
# + id="yh0S_9pVzX5-"
batch_size = 32
dataset = dataset.shuffle(10000).batch(batch_size)
# + colab={"base_uri": "https://localhost:8080/"} id="54JFoIzY3zl2" outputId="fe4b6791-80ea-451f-a91a-92a09de88b83"
for item in dataset.take(1):
print(item)
# + id="sTMkqR3T3x6w"
dataset = dataset.map(lambda windows: (windows[:, :-1], windows[:, 1:]))
# + colab={"base_uri": "https://localhost:8080/"} id="phH9GfPnz6I-" outputId="f7cf830f-c8b1-408d-83b8-e16fe013c6e1"
for item in dataset.take(1):
print(item)
# + id="W5Ex6sjg3896"
dataset = dataset.map(lambda X_batch, Y_batch: (tf.one_hot(X_batch, depth=max_id), Y_batch))
# + colab={"base_uri": "https://localhost:8080/"} id="YGKITH_44em_" outputId="04a26689-381e-4ce1-f923-ce87868d775a"
for X, y in dataset.take(1):
print(X.shape, y.shape)
# + id="8hmbb9O04i8E"
dataset = dataset.prefetch(1)
# + [markdown] id="J88YgPzQ4noT"
# ## Building Model:
# + colab={"base_uri": "https://localhost:8080/"} id="fZMKSxKc5LYw" outputId="cb59bad1-2123-4a39-e93b-ba67949c58ea"
shakespearean_model = keras.models.Sequential([
keras.layers.GRU(128, return_sequences=True, input_shape=[None, max_id],
dropout=0.2),#, recurrent_dropout=0.2),
keras.layers.GRU(128, return_sequences=True,
dropout=0.2),#, recurrent_dropout=0.2),
keras.layers.TimeDistributed(keras.layers.Dense(max_id, activation="softmax")),
])
shakespearean_model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.RMSprop(4e-4),
metrics=["accuracy"])
history = shakespearean_model.fit(dataset, epochs=10)
# + [markdown] id="LAqxRFFr6JuN"
# ## Predicting a Character:
# + id="JjYSQe0GdFZS"
def preprocess(texts):
X = np.array(tokenizer.texts_to_sequences(texts)) - 1
return tf.one_hot(X, max_id)
# + id="wWYaOBVKdPyh"
X_new = preprocess(["How are yo"])
Y_pred = shakespearean_model.predict_classes(X_new)
tokenizer.sequences_to_texts(Y_pred + 1)[0][-1]
# + [markdown] id="Jbu9qLr7dkO7"
# ## Predicting multilpe characters:
# + id="Q4-eTpIleKA2"
def next_char(text, temperature=1):
X_new = preprocess([text])
y_proba = shakespearean_model.predict(X_new)[0, -1:, :]
rescaled_logits = tf.math.log(y_proba) / temperature
char_id = tf.random.categorical(rescaled_logits, num_samples=1) + 1
return tokenizer.sequences_to_texts(char_id.numpy())[0]
# + id="defN9jVXergh"
def complete_text(text, n_chars=50, temperature=1):
for _ in range(n_chars):
text += next_char(text, temperature)
return text
# + id="XOyfO84ClHvj"
print(complete_text("t", temperature=0.2))
# + id="8NJOEqB1lWCd"
print(complete_text("a", temperature=0.5))
# + id="9FBpYJITldtB"
print(complete_text("s", temperature=1))
# + id="0z9ixZrZlhpR"
print(complete_text("r", temperature=2))
# + [markdown] id="e2OY4UNOlmrt"
# ## Stateful RNN:
# + [markdown] id="rWdgmVYPRXLa"
# Fabien Chollet gives this definition of STATEFULNESS:
# <br>Boolean (default False). If True, the last state for each sample at index i in a batch will be used as initial state for the sample of index i in the following batch.
# <br>
#
# By default, Keras shuffles (permutes) the samples in X and the dependencies between Xi and Xi+1 are lost. Let’s assume there’s no shuffling in our explanation.
#
# If the model is stateless, the cell states are reset at each sequence. With the stateful model, all the states are propagated to the next batch. It means that the state of the sample located at index i, Xi will be used in the computation of the sample Xi+bs in the next batch, where bs is the batch size (no shuffling).
#
# + id="TrqQl60LEBEP"
batch_size = 32
encoded_parts = np.array_split(encoded[:train_size], batch_size)
datasets = []
for encoded_part in encoded_parts:
dataset = tf.data.Dataset.from_tensor_slices(encoded_part)
dataset = dataset.window(window_length, shift=n_steps, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_length))
datasets.append(dataset)
dataset = tf.data.Dataset.zip(tuple(datasets)).map(lambda *windows: tf.stack(windows))
dataset = dataset.repeat().map(lambda windows: (windows[:, :-1], windows[:, 1:]))
dataset = dataset.map(
lambda X_batch, Y_batch: (tf.one_hot(X_batch, depth=max_id), Y_batch))
dataset = dataset.prefetch(1)
# + id="4YgCGoUnSTdg"
stateful_model = keras.models.Sequential([
keras.layers.GRU(128, return_sequences=True, stateful=True,
dropout=0.2, recurrent_dropout=0.2,
batch_input_shape=[batch_size, None, max_id]),
keras.layers.GRU(128, return_sequences=True, stateful=True,
dropout=0.2, recurrent_dropout=0.2),
keras.layers.TimeDistributed(keras.layers.Dense(max_id,
activation="softmax"))
])
# + id="nQoKh-qBTYOV"
class ResetStatesCallback(keras.callbacks.Callback):
def on_epoch_begin(self, epoch, logs):
self.model.reset_states()
# + colab={"base_uri": "https://localhost:8080/"} id="Pb4NlA47Tpf2" outputId="55cfef05-c6a2-49fc-ca4d-7121fe8719d9"
stateful_model.compile(loss="sparse_categorical_crossentropy",
optimizer="adam")
steps_per_epoch = train_size // batch_size // n_steps
stateful_model.fit(dataset, steps_per_epoch=steps_per_epoch,
epochs=50, callbacks=[ResetStatesCallback()])
# + id="w4Up-rj8qQQZ"
|
notebooks/ShakespeareanText_Generator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:p3]
# language: python
# name: conda-env-p3-py
# ---
# Upload data to sqlite tables.
# +
import pandas as pd
import numpy as np
from IPython import display as dis
import scipy.io.wavfile as wav
import tensorflow as tf
from tensorflow.contrib.legacy_seq2seq.python.ops.seq2seq import basic_rnn_seq2seq
from tensorflow.contrib.rnn import RNNCell, LSTMCell, MultiRNNCell
from scipy import signal
from librosa import core
# %matplotlib inline
# -
dis.Audio("dataset/wav/Ses01F_impro01/Ses01F_impro01_F000.wav")
(sig,rate) = core.load("dataset/wav/Ses01F_impro01/Ses01F_impro01_F000.wav", sr = 4000)
print(sig, rate)
print(len(sig))
dis.Audio(data = sig, rate = rate)
class network(object):
time_step = 7783
hidden_layers = 1
latent_dim=61
batch_size = 1
def __init__(self):
pass
def build_layers(self):
tf.reset_default_graph()
#learning_rate = tf.Variable(initial_value=0.001)
time_step = self.time_step
hidden_layers = self.hidden_layers
latent_dim = self.latent_dim
batch_size = self.batch_size
with tf.variable_scope("Input"):
self.x_input = tf.placeholder("float", shape=[batch_size, time_step, 1])
self.y_input_ = tf.placeholder("float", shape=[batch_size, time_step, 1])
self.keep_prob = tf.placeholder("float")
self.lr = tf.placeholder("float")
self.x_list = tf.unstack(self.x_input, axis= 1)
self.y_list_ = tf.unstack(self.y_input_, axis = 1)
with tf.variable_scope("lstm"):
multi_cell = MultiRNNCell([LSTMCell(latent_dim) for i in range(hidden_layers)] )
self.y, states = basic_rnn_seq2seq(self.x_list, self.y_list_, multi_cell)
#self.y = tf.slice(self.y, [0, 0], [-1,2])
#self.out = tf.squeeze(self.y)
#self.y = tf.layers.dense(self.y[0], classes, activation = None)
#self.y = tf.slice(self.y[0], [0, 0], [-1,2])
self.y = tf.slice(self.y, [0, 0, 0], [-1,-1,1])
with tf.variable_scope("Loss"):
self.pred = tf.stack(self.y)
self.regularized_loss = tf.losses.mean_squared_error(self.y, self.y_list_)
with tf.variable_scope("Optimizer"):
learning_rate=self.lr
optimizer = tf.train.AdamOptimizer(learning_rate)
gradients, variables = zip(*optimizer.compute_gradients(self.regularized_loss))
gradients = [
None if gradient is None else tf.clip_by_value(gradient, -1, 1)
for gradient in gradients]
self.train_op = optimizer.apply_gradients(zip(gradients, variables))
#self.train_op = optimizer.minimize(self.regularized_loss)
# add op for merging summary
#self.summary_op = tf.summary.merge_all()
# add Saver ops
self.saver = tf.train.Saver()
# +
import collections
y_pred = None
class Train:
def train(epochs, net, lrs):
global y_pred
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
start_time = time.perf_counter()
for c, lr in enumerate(lrs):
for epoch in range(1, (epochs+1)):
print("Step {} ".format(train_loss))
_, train_loss = sess.run([net.train_op, net.regularized_loss], #net.summary_op
feed_dict={net.x_input: x_train[np.newaxis,...],
net.y_input_: y_train[np.newaxis,...],
net.lr:lr})
print("Training Loss: {:.6f}".format(train_loss))
#valid_accuracy,valid_loss = sess.run([net.tf_accuracy, net.regularized_loss], #net.summary_op
# feed_dict={net.x_input: x_valid[np.newaxis,...],
# net.y_input_: y_valid[np.newaxis,...],
# net.lr:lr})
accuracy, y_pred = sess.run([net.regularized_loss,
net.pred],
feed_dict={net.x_input: x_train[np.newaxis,...],
net.y_input_: y_train[np.newaxis,...],
net.lr:lr})
# +
import itertools
class Hyperparameters:
def start_training():
epochs = 1
lrs = [1e-5]
n = network()
n.build_layers()
Train.train(epochs, n, lrs)
# -
sig = np.reshape(sig, (1,-1, 1))
print(sig.shape)
x_train = y_train = sig
Hyperparameters.start_training()
y_pred
|
feature_extraction_seq2seq.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pilatus on a goniometer at ID28
#
# <NAME> who was post-doc at ESRF-ID28 enquired about a potential bug in pyFAI in October 2016: he calibrated 3 images taken with a Pilatus-1M detector at various detector angles: 0, 17 and 45 degrees.
# While everything looked correct, in first approximation, one peak did not overlap properly with itself depending on the detector angle. This peak correspond to the peak in the angle of the detector, at 23.6° ...
#
# This notebook will guide you through the calibration of the goniometer setup.
#
# Let's first retrieve the images and initialize the environment:
# %matplotlib inline
import os, sys, time
start_time = time.perf_counter()
print(sys.version)
import numpy
import fabio, pyFAI
print(f"Using pyFAI version: {pyFAI.version}")
from os.path import basename
from pyFAI.gui import jupyter
from pyFAI.calibrant import get_calibrant
from silx.resources import ExternalResources
from scipy.interpolate import interp1d
from scipy.optimize import bisect
from matplotlib.pyplot import subplots
from matplotlib.lines import Line2D
downloader = ExternalResources("thick", "http://www.silx.org/pub/pyFAI/testimages")
all_files = downloader.getdir("gonio_ID28.tar.bz2")
for afile in all_files:
print(basename(afile))
# There are 3 images stored as CBF files and the associated control points as npt files.
# +
images = [i for i in all_files if i.endswith("cbf")]
images.sort()
mask = None
fig, ax = subplots(1,3, figsize=(9,3))
for i, cbf in enumerate(images):
fimg = fabio.open(cbf)
jupyter.display(fimg.data, label=basename(cbf), ax=ax[i])
if mask is None:
mask = fimg.data<0
else:
mask |= fimg.data<0
numpy.save("mask.npy", mask)
# -
# To be able to calibrate the detector position, the calibrant used is LaB6 and the wavelength was 0.69681e-10m
# +
wavelength=0.6968e-10
calibrant = get_calibrant("LaB6")
calibrant.wavelength = wavelength
print(calibrant)
detector = pyFAI.detector_factory("Pilatus1M")
# +
# Define the function that extracts the angle from the filename:
def get_angle(basename):
"""Takes the basename (like det130_g45_0001.cbf ) and returns the angle of the detector"""
return float(os.path.basename((basename.split("_")[-2][1:])))
for afile in images:
print('filename', afile, "angle:",get_angle(afile))
# +
#Define the transformation of the geometry as function of the goniometrer position.
# by default scale1 = pi/180 (convert deg to rad) and scale2 = 0.
from pyFAI.goniometer import GeometryTransformation, GoniometerRefinement, Goniometer
goniotrans2d = GeometryTransformation(param_names = ["dist", "poni1", "poni2",
"rot1", "rot2",
"scale1", "scale2"],
dist_expr="dist",
poni1_expr="poni1",
poni2_expr="poni2",
rot1_expr="scale1 * pos + rot1",
rot2_expr="scale2 * pos + rot2",
rot3_expr="0.0")
# +
epsilon = numpy.finfo(numpy.float32).eps
#Definition of the parameters start values and the bounds
param = {"dist":0.30,
"poni1":0.08,
"poni2":0.08,
"rot1":0,
"rot2":0,
"scale1": numpy.pi/180., # rot2 is in radians, while the motor position is in degrees
"scale2": 0
}
#Defines the bounds for some variables. We start with very strict bounds
bounds = {"dist": (0.25, 0.31),
"poni1": (0.07, 0.1),
"poni2": (0.07, 0.1),
"rot1": (-0.01, 0.01),
"rot2": (-0.01, 0.01),
"scale1": (numpy.pi/180.-epsilon, numpy.pi/180.+epsilon), #strict bounds on the scale: we expect the gonio to be precise
"scale2": (-epsilon, +epsilon) #strictly bound to 0
}
# -
gonioref2d = GoniometerRefinement(param, #initial guess
bounds=bounds,
pos_function=get_angle,
trans_function=goniotrans2d,
detector=detector,
wavelength=wavelength)
print("Empty goniometer refinement object:")
print(gonioref2d)
# +
# Populate with the images and the control points
for fn in images:
base = os.path.splitext(fn)[0]
bname = os.path.basename(base)
fimg = fabio.open(fn)
sg =gonioref2d.new_geometry(bname, image=fimg.data, metadata=bname,
control_points=base+".npt",
calibrant=calibrant)
print(sg.label, "Angle:", sg.get_position())
print("Filled refinement object:")
print(gonioref2d)
# +
# Initial refinement of the goniometer model with 5 dof
gonioref2d.refine2()
# -
# Remove constrains on the refinement:
gonioref2d.bounds=None
gonioref2d.refine2()
# +
# Check the calibration on all 3 images
fig, ax = subplots(1, 3, figsize=(18, 6) )
for idx,lbl in enumerate(gonioref2d.single_geometries):
sg = gonioref2d.single_geometries[lbl]
if sg.control_points.get_labels():
sg.geometry_refinement.set_param(gonioref2d.get_ai(sg.get_position()).param)
a=jupyter.display(sg=sg, ax=ax[idx])
# +
#Create a MultiGeometry integrator from the refined geometry:
angles = []
images = []
for sg in gonioref2d.single_geometries.values():
angles.append(sg.get_position())
images.append(sg.image)
multigeo = gonioref2d.get_mg(angles)
multigeo.radial_range=(0, 63)
print(multigeo)
# Integrate the whole set of images in a single run:
res_mg = multigeo.integrate1d(images, 10000)
fig, ax = subplots(1, 2, figsize=(12,4))
ax0 = jupyter.plot1d(res_mg, label="multigeo", ax=ax[0])
ax1 = jupyter.plot1d(res_mg, label="multigeo", ax=ax[1])
# Let's focus on the inner most ring on the image taken at 45°:
for lbl, sg in gonioref2d.single_geometries.items():
ai = gonioref2d.get_ai(sg.get_position())
img = sg.image * ai.dist * ai.dist / ai.pixel1 / ai.pixel2
res = ai.integrate1d(img, 5000, unit="2th_deg", method="splitpixel")
ax0.plot(*res, "--", label=lbl)
ax1.plot(*res, "--", label=lbl)
ax1.set_xlim(29,29.3)
ax0.set_ylim(0, 1.5e12)
ax1.set_ylim(0, 7e11)
p8tth = numpy.rad2deg(calibrant.get_2th()[7])
ax1.set_title("Zoom on peak #8 at %.4f°"%p8tth)
l = Line2D([p8tth, p8tth], [0, 2e12])
ax1.add_line(l)
ax0.legend()
ax1.legend().remove()
pass
# -
# On all three imges, the rings on the outer side of the detector are shifted in compatison with the average signal comming from the other two images.
# This phenomenon could be related to volumetric absorption of the photon in the thickness of the detector.
#
# To be able to investigate this phenomenon further, the goniometer geometry is saved in a JSON file:
# +
gonioref2d.save("id28.json")
with open("id28.json") as f:
print(f.read())
# -
# ## Peak profile
#
# Let's plot the full-width at half maximum for every peak in the different intergated profiles:
# +
#Peak profile
def calc_fwhm(integrate_result, calibrant):
"calculate the tth position and FWHM for each peak"
delta = integrate_result.intensity[1:] - integrate_result.intensity[:-1]
maxima = numpy.where(numpy.logical_and(delta[:-1]>0, delta[1:]<0))[0]
minima = numpy.where(numpy.logical_and(delta[:-1]<0, delta[1:]>0))[0]
maxima += 1
minima += 1
tth = []
FWHM = []
for tth_rad in calibrant.get_2th():
tth_deg = tth_rad*integrate_result.unit.scale
if (tth_deg<=integrate_result.radial[0]) or (tth_deg>=integrate_result.radial[-1]):
continue
idx_theo = abs(integrate_result.radial-tth_deg).argmin()
id0_max = abs(maxima-idx_theo).argmin()
id0_min = abs(minima-idx_theo).argmin()
I_max = integrate_result.intensity[maxima[id0_max]]
I_min = integrate_result.intensity[minima[id0_min]]
tth_maxi = integrate_result.radial[maxima[id0_max]]
I_thres = (I_max + I_min)/2.0
if minima[id0_min]>maxima[id0_max]:
if id0_min == 0:
min_lo = integrate_result.radial[0]
else:
min_lo = integrate_result.radial[minima[id0_min-1]]
min_hi = integrate_result.radial[minima[id0_min]]
else:
if id0_min == len(minima) -1:
min_hi = integrate_result.radial[-1]
else:
min_hi = integrate_result.radial[minima[id0_min+1]]
min_lo = integrate_result.radial[minima[id0_min]]
f = interp1d(integrate_result.radial, integrate_result.intensity-I_thres)
tth_lo = bisect(f, min_lo, tth_maxi)
tth_hi = bisect(f, tth_maxi, min_hi)
FWHM.append(tth_hi-tth_lo)
tth.append(tth_deg)
return tth, FWHM
fig, ax = subplots()
ax.plot(*calc_fwhm(res_mg, calibrant), "o", label="multi")
for lbl, sg in gonioref2d.single_geometries.items():
ai = gonioref2d.get_ai(sg.get_position())
img = sg.image * ai.dist * ai.dist / ai.pixel1 / ai.pixel2
res = ai.integrate1d(img, 5000, unit="2th_deg", method="splitpixel")
t,w = calc_fwhm(res, calibrant=calibrant)
ax.plot(t, w,"-o", label=lbl)
ax.set_title("Peak shape as function of the angle")
ax.set_xlabel(res_mg.unit.label)
ax.legend()
pass
# -
# ## Conclusion:
# Can the FWHM and peak position be corrected using raytracing and deconvolution ?
print(f"Total execution time: {time.perf_counter()-start_time:.3f} s")
|
doc/source/usage/tutorial/ThickDetector/goniometer_id28.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def apply_word_dropout(matrix, keep_prop, replace_with=UNK_IX, pad_ix=PAD_IX,):
dropout_mask = np.random.choice(2, np.shape(matrix), p=[keep_prop, 1 - keep_prop])
dropout_mask &= matrix != pad_ix
return np.choose(dropout_mask, [matrix, np.full_like(matrix, replace_with)])
np.choose()
# +
import numpy as np
matrix = np.random.normal(size=(3, 4))
keep_prop = .1
matrix
# -
dropout_mask = np.random.choice(2, np.shape(matrix), p=[keep_prop, 1 - keep_prop])
dropout_mask
pad_ix = 2
dropout_mask &= matrix != pad_ix
dropout_mask
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import os
import gc
import time
import math
from sklearn.metrics import roc_auc_score
from gensim.models import Word2Vec
SEED = 41
np.random.seed(SEED)
# +
RAW_DATA_PATH = '../../dl_nlp/data/jigsaw_toxic/raw/'
PROCESSED_DATA_PATH = '../../dl_nlp/data/jigsaw_toxic/processed/'
MAX_LEN = 100
# -
# ### Load Data
# +
# %%time
train = pd.read_csv(os.path.join(RAW_DATA_PATH, 'train.csv'))
test = pd.read_csv(os.path.join(RAW_DATA_PATH, 'test.csv'))
test_labels = pd.read_csv(os.path.join(RAW_DATA_PATH, 'test_labels.csv'))
# -
# #### Define target columns
TARGET_COLS = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']
# #### Load Sample
# +
# train = pd.read_csv(os.path.join(PROCESSED_DATA_PATH, 'train_sample.csv'))
# -
# #### Process dataset
train['decent'] = 1 - train.loc[:, TARGET_COLS].max(axis=1)
TARGET_COLS += ['decent']
# #### Preprocessing
import nltk
tokenizer = nltk.tokenize.WordPunctTokenizer()
# %%time
train_tokenized_comments = list(map(tokenizer.tokenize, train.comment_text))
# %%time
train['tokenized_comments'] = list(map(' '.join, map(tokenizer.tokenize, train.comment_text)))
test['tokenized_comments'] = list(map(' '.join, map(tokenizer.tokenize, test.comment_text)))
# #### Define Word2Vec model
# +
# %%time
model = Word2Vec(train_tokenized_comments,
size=32,
min_count=10,
window=5).wv
# -
words = sorted(model.vocab.keys(),
key=lambda word: model.vocab[word].count,
reverse=True)
# #### Load pretrained embeddings
word_vectors = model.vectors[[model.vocab[word].index for word in words]]
emb_mean,emb_std = word_vectors.mean(), word_vectors.std()
emb_mean,emb_std
# +
UNK, PAD = 'UNK', 'PAD'
UNK_IX, PAD_IX = len(words), len(words) + 1
nb_words = len(words) + 2
embed_size = 32
embedding_matrix = np.random.normal(emb_mean, emb_std, (nb_words, embed_size))
for word in words + [UNK, PAD]:
if word in model.vocab:
word_idx = model.vocab[word].index
embedding_vector = model.vectors[model.vocab[word].index]
embedding_matrix[word_idx] = embedding_vector
# +
token_to_id = {word: model.vocab[word].index for word in words}
token_to_id[UNK] = UNK_IX
token_to_id[PAD] = PAD_IX
# +
UNK_IX, PAD_IX = map(token_to_id.get, [UNK, PAD])
def as_matrix(sequences, max_len=None):
""" Convert a list of tokens into a matrix with padding """
if isinstance(sequences[0], str):
sequences = list(map(str.split, sequences))
max_len = min(max(map(len, sequences)), max_len or float('inf'))
matrix = np.full((len(sequences), max_len), np.int32(PAD_IX))
for i,seq in enumerate(sequences):
row_ix = [token_to_id.get(word, UNK_IX) for word in seq[:max_len]]
matrix[i, :len(row_ix)] = row_ix
return matrix
# +
from sklearn.model_selection import train_test_split
data_train, data_val = train_test_split(train, test_size=0.2, random_state=42)
data_train.index = range(len(data_train))
data_val.index = range(len(data_val))
print("Train size = ", len(data_train))
print("Validation size = ", len(data_val))
# -
def iterate_batches(matrix, labels, batch_size, predict_mode='train'):
indices = np.arange(len(matrix))
if predict_mode == 'train':
np.random.shuffle(indices)
for start in range(0, len(matrix), batch_size):
end = min(start + batch_size, len(matrix))
batch_indices = indices[start: end]
X = matrix[batch_indices]
if predict_mode != 'train': yield X
else: yield X, labels[batch_indices]
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
class TConvolution(nn.Module):
def __init__(self, pre_trained_embeddings, vocab_size, hidden_dim, num_classes, PAD_IX):
super(TConvolution, self).__init__()
self.hidden_dim = hidden_dim
self.vocab_size = vocab_size
self.num_classes = num_classes
self.Cin = 1
self.Cout = 1
self.embedding = nn.Embedding(self.vocab_size, self.hidden_dim, padding_idx=PAD_IX)
self.embedding.weight = nn.Parameter(pre_trained_embeddings)
self.conv_layer1 = nn.Conv2d(self.Cout, self.Cin, kernel_size=(3, self.hidden_dim))
self.conv_layer2 = nn.Conv2d(self.Cout, self.Cin, kernel_size=(4, self.hidden_dim))
self.relu = nn.ReLU()
self.fc = nn.Linear(2, self.num_classes)
def forward(self, x):
emb = self.embedding(x)
# create a matrix of shape ( N, Cin, max_len, embedding_dim)
emb = emb.unsqueeze(1)
# pass it through convolutional layer to calculate unigrams
out1 = self.conv_layer1(emb)
out1 = self.relu(out1)
out1 = out1.squeeze(3)
out2 = self.conv_layer2(emb)
out2 = self.relu(out2)
out2 = out2.squeeze(3)
# global max pool
out1, _ = torch.max(out1, dim=-1)
out2, _ = torch.max(out2, dim=-1)
out = torch.cat((out1, out2), dim=1)
# fully connected layer
out = self.fc(out)
return out
# +
def do_epoch(model, criterion, data, batch_size, optimizer=None):
epoch_loss, total_size = 0, 0
per_label_preds = [[], [], [], [], [], [], []]
per_label_true = [[], [], [], [], [], [], []]
is_train = not optimizer is None
model.train(is_train)
data, labels = data
batchs_count = math.ceil(data.shape[0] / batch_size)
with torch.autograd.set_grad_enabled(is_train):
for i, (X_batch, y_batch) in enumerate(iterate_batches(data, labels, batch_size)):
X_batch, y_batch = torch.cuda.LongTensor(X_batch), torch.cuda.FloatTensor(y_batch)
logits = model(X_batch)
loss = criterion(logits, y_batch)
if is_train:
loss.backward()
optimizer.step()
optimizer.zero_grad()
# convert true target
batch_target = y_batch.cpu().detach().numpy()
logits_cpu = logits.cpu().detach().numpy()
# per_label_preds
for j in range(7):
label_preds = logits_cpu[:, j]
per_label_preds[j].extend(label_preds)
per_label_true[j].extend(batch_target[:, j])
# calculate log loss
epoch_loss += loss.item()
print('\r[{} / {}]: Loss = {:.4f}'.format(
i, batchs_count, loss.item(), end=''))
label_auc = []
for i in range(7):
label_auc.append(roc_auc_score(per_label_true[i], per_label_preds[i]))
return epoch_loss / batchs_count, np.mean(label_auc)
def fit(model, criterion, optimizer, train_data, epochs_count=1,
batch_size=32, val_data=None, val_batch_size=None):
if not val_data is None and val_batch_size is None:
val_batch_size = batch_size
for epoch in range(epochs_count):
start_time = time.time()
train_loss, train_auc = do_epoch(
model, criterion, train_data, batch_size, optimizer
)
output_info = '\rEpoch {} / {}, Epoch Time = {:.2f}s: Train Loss = {:.4f}, Train AUC = {:.4f}'
if not val_data is None:
val_loss, val_auc = do_epoch(model, criterion, val_data, val_batch_size, None)
epoch_time = time.time() - start_time
output_info += ', Val Loss = {:.4f}, Val AUC = {:.4f}'
print(output_info.format(epoch+1, epochs_count, epoch_time,
train_loss,
train_auc,
val_loss,
val_auc
))
else:
epoch_time = time.time() - start_time
print(output_info.format(epoch+1, epochs_count, epoch_time, train_loss, train_auc))
# +
model = TConvolution(pre_trained_embeddings=torch.FloatTensor(embedding_matrix),
vocab_size=len(token_to_id),
hidden_dim=embed_size,
num_classes=7,
PAD_IX=PAD_IX).cuda()
criterion = nn.BCEWithLogitsLoss().cuda()
optimizer = optim.Adam([param for param in model.parameters() if param.requires_grad], lr=0.01)
X_train = as_matrix(data_train['tokenized_comments'])
train_labels = data_train.loc[:, TARGET_COLS].values
X_test = as_matrix(data_val['tokenized_comments'])
test_labels = data_val.loc[:, TARGET_COLS].values
fit(model, criterion, optimizer, train_data=(X_train, train_labels), epochs_count=5,
batch_size=512, val_data=(X_test, test_labels), val_batch_size=1024)
# -
# #### Token to count
from collections import Counter
token_counts = Counter()
# +
# %%time
for comment in train.tokenized_comments:
token_counts.update(comment.split())
# -
token_counts.most_common(10)
print('Total unique tokens: {}'.format(len(token_counts)))
print('\n'.join(map(str, token_counts.most_common(5))))
print()
print('\n'.join(map(str, token_counts.most_common()[-3:])))
plt.hist(token_counts.values(), range=(0, 10 ** 4), log=True, bins=50)
plt.xlabel('Token Counts');
# #### Remove rare words from the dictionary.
min_count = 10
tokens = {token: count for token, count in token_counts.items() if count >= min_count}
# +
UNK, PAD = 'UNK', 'PAD'
tokens = [UNK, PAD] + sorted(tokens)
print('Vocabulary size:', len(tokens))
assert type(tokens) == list
assert 'me' in tokens
assert UNK in tokens
print("Correct!")
# -
# #### Mapping from token to id
# %%time
token_to_id = dict(map(reversed, zip(range(len(tokens)), tokens)))
# And finally, let's use the vocabulary you've built to map text lines into neural network-digestible matrices.
# +
UNK_IX, PAD_IX = map(token_to_id.get, [UNK, PAD])
def as_matrix(sequences, max_len=None):
""" Convert a list of tokens into a matrix with padding """
if isinstance(sequences[0], str):
sequences = list(map(str.split, sequences))
max_len = min(max(map(len, sequences)), max_len or float('inf'))
matrix = np.full((len(sequences), max_len), np.int32(PAD_IX))
for i,seq in enumerate(sequences):
row_ix = [token_to_id.get(word, UNK_IX) for word in seq[:max_len]]
matrix[i, :len(row_ix)] = row_ix
return matrix
# -
print('Lines:')
print('\n'.join(train['tokenized_comments'][:2].values), end='\n\n')
print('Matrix:')
print(as_matrix(train['tokenized_comments'])[:2])
# ### Split into training and test set
# +
from sklearn.model_selection import train_test_split
data_train, data_val = train_test_split(train, test_size=0.2, random_state=42)
data_train.index = range(len(data_train))
data_val.index = range(len(data_val))
print("Train size = ", len(data_train))
print("Validation size = ", len(data_val))
# -
# ### Create Batches
#
# ```
# [[1, 2, 3, 4],
# [4, 1, 4, 1],
# ....
# ] -> [[0, 1, 1], [1, 1, 0]]
# ```
def iterate_batches(matrix, labels, batch_size, predict_mode='train'):
indices = np.arange(len(matrix))
if predict_mode == 'train':
np.random.shuffle(indices)
for start in range(0, len(matrix), batch_size):
end = min(start + batch_size, len(matrix))
batch_indices = indices[start: end]
X = matrix[batch_indices]
if predict_mode != 'train': yield X
else: yield X, labels[batch_indices]
# +
matrix = as_matrix(data_train['tokenized_comments'], max_len=MAX_LEN)
labels = data_train.loc[:, TARGET_COLS].values
X, y = next(iterate_batches(matrix, labels, batch_size=2))
# -
# ### Model
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
class TConvolution(nn.Module):
def __init__(self, vocab_size, hidden_dim, num_classes, PAD_IX):
super(TConvolution, self).__init__()
self.hidden_dim = hidden_dim
self.vocab_size = vocab_size
self.num_classes = num_classes
self.Cin = 1
self.Cout = 1
self.embedding = nn.Embedding(self.vocab_size, self.hidden_dim, padding_idx=PAD_IX)
self.conv_layer1 = nn.Conv2d(self.Cout, self.Cin, kernel_size=(1, self.hidden_dim))
self.conv_layer2 = nn.Conv2d(self.Cout, self.Cin, kernel_size=(2, self.hidden_dim))
self.relu = nn.ReLU()
self.fc = nn.Linear(2, self.num_classes)
def forward(self, x):
emb = self.embedding(x)
# create a matrix of shape ( N, Cin, max_len, embedding_dim)
emb = emb.unsqueeze(1)
# pass it through convolutional layer to calculate unigrams
out1 = self.conv_layer1(emb)
out1 = self.relu(out1)
out1 = out1.squeeze(3)
out2 = self.conv_layer2(emb)
out2 = self.relu(out2)
out2 = out2.squeeze(3)
# global max pool
out1, _ = torch.max(out1, dim=-1)
out2, _ = torch.max(out2, dim=-1)
out = torch.cat((out1, out2), dim=1)
# fully connected layer
out = self.fc(out)
return out
model = TConvolution(len(token_to_id), hidden_dim=32, num_classes=7, PAD_IX=PAD_IX).cuda()
criterion = nn.BCEWithLogitsLoss().cuda()
# ### Run on a single batch
# +
X = torch.cuda.LongTensor(X)
y = torch.cuda.FloatTensor(y)
logits = model(X)
# -
loss = criterion(logits, y)
loss.item()
# ### Training Loop
# ```
# Evaluation:
#
# Mean columnwise auc score: In other words, the score is the average of the individual AUCs of each predicted column.
#
# example:
#
# TRUE:
# a b c
# [[1, 0, 1],
# [0, 1, 0],
# [0, 0, 1],
# [0, 1, 1]
# ]
#
# PREDS:
# a b c
# [[0.3, 0.6, 0.1],
# [0.1, 0.1, 0.8],
# [0.2, 0.2, 0.6]
# ]
#
# AUC score for column (a) : auc_a = roc_auc_score(true_a, preds_a)
# AUC score for column (b) : auc_b = roc_auc_score(true_b, preds_b)
# AUC score for column (c) : auc_c = roc_auc_score(true_c, preds_c)
#
# Mean score = (auc_a + auc_b + auc_c) / 3
#
#
# Required:
#
# true matrix = one hot encoded data frame of target labels
#
# ```
# +
def do_epoch(model, criterion, data, batch_size, optimizer=None):
epoch_loss, total_size = 0, 0
per_label_preds = [[], [], [], [], [], [], []]
per_label_true = [[], [], [], [], [], [], []]
is_train = not optimizer is None
model.train(is_train)
data, labels = data
batchs_count = math.ceil(data.shape[0] / batch_size)
with torch.autograd.set_grad_enabled(is_train):
for i, (X_batch, y_batch) in enumerate(iterate_batches(data, labels, batch_size)):
X_batch, y_batch = torch.cuda.LongTensor(X_batch), torch.cuda.FloatTensor(y_batch)
logits = model(X_batch)
loss = criterion(logits, y_batch)
if is_train:
loss.backward()
optimizer.step()
optimizer.zero_grad()
# convert true target
batch_target = y_batch.cpu().detach().numpy()
logits_cpu = logits.cpu().detach().numpy()
# per_label_preds
for j in range(7):
label_preds = logits_cpu[:, j]
per_label_preds[j].extend(label_preds)
per_label_true[j].extend(batch_target[:, j])
# calculate log loss
epoch_loss += loss.item()
print('\r[{} / {}]: Loss = {:.4f}'.format(
i, batchs_count, loss.item(), end=''))
label_auc = []
for i in range(7):
label_auc.append(roc_auc_score(per_label_true[i], per_label_preds[i]))
return epoch_loss / batchs_count, np.mean(label_auc)
def fit(model, criterion, optimizer, train_data, epochs_count=1,
batch_size=32, val_data=None, val_batch_size=None):
if not val_data is None and val_batch_size is None:
val_batch_size = batch_size
for epoch in range(epochs_count):
start_time = time.time()
train_loss, train_auc = do_epoch(
model, criterion, train_data, batch_size, optimizer
)
output_info = '\rEpoch {} / {}, Epoch Time = {:.2f}s: Train Loss = {:.4f}, Train AUC = {:.4f}'
if not val_data is None:
val_loss, val_auc = do_epoch(model, criterion, val_data, val_batch_size, None)
epoch_time = time.time() - start_time
output_info += ', Val Loss = {:.4f}, Val AUC = {:.4f}'
print(output_info.format(epoch+1, epochs_count, epoch_time,
train_loss,
train_auc,
val_loss,
val_auc
))
else:
epoch_time = time.time() - start_time
print(output_info.format(epoch+1, epochs_count, epoch_time, train_loss, train_auc))
# +
model = TConvolution(len(token_to_id), hidden_dim=64, num_classes=7).cuda()
criterion = nn.BCEWithLogitsLoss().cuda()
optimizer = optim.Adam([param for param in model.parameters() if param.requires_grad], lr=0.01)
X_train = as_matrix(data_train['tokenized_comments'])
train_labels = data_train.loc[:, TARGET_COLS].values
X_test = as_matrix(data_val['tokenized_comments'])
test_labels = data_val.loc[:, TARGET_COLS].values
fit(model, criterion, optimizer, train_data=(X_train, train_labels), epochs_count=5,
batch_size=512, val_data=(X_test, test_labels), val_batch_size=1024)
# -
# ### Full Training
# ```
# Full Training
#
# a) Train model for 3 epochs using our Convolutional Model.
# b) Store all the hyper-parameters used for the experiment.
# c) Store model to disk using Pytorch best practices.
# d) Load model from disk in the prediction phase ( full test set )
# e) Write a method that takes in model and the test dataset and returns back all the predictions in the same
# order.
# f) Generate logits for every label ( we would generate logits for all the 7 labels but we need sigmoid
# for only 6 labels ).
# g) Apply sigmoid across all rows for all the 6 not 7 labels.
# h) Using Kaggle API, submit predictions to kaggle and note down the public and private evaluation score.
# ```
def predict(model, data, batch_size):
is_train = False
model.train(is_train)
batchs_count = math.ceil(data.shape[0] / batch_size)
preds = []
with torch.autograd.set_grad_enabled(is_train):
for i, X_batch in enumerate(iterate_batches(data, labels=[], batch_size=batch_size, predict_mode='test')):
X_batch = torch.cuda.LongTensor(X_batch)
logits = model(X_batch)
p = torch.sigmoid(logits).cpu().detach().numpy()
preds.append(p)
return np.vstack(preds)
# +
model = TConvolution(len(token_to_id), hidden_dim=64, num_classes=7, PAD_IX=PAD_IX).cuda()
criterion = nn.BCEWithLogitsLoss().cuda()
optimizer = optim.Adam([param for param in model.parameters() if param.requires_grad], lr=0.01)
X_train = as_matrix(train['tokenized_comments'])
train_labels = train.loc[:, TARGET_COLS].values
# train model
fit(model,
criterion,
optimizer,
train_data=(X_train, train_labels),
epochs_count=3,
batch_size=512)
# -
# generate Xtest matrix
X_test = as_matrix(test['tokenized_comments'])
preds = predict(model, X_test, batch_size=512)
# create generator for test set
preds.shape
train.head()
test_labels.iloc[:, 1:] = preds[:, :-1]
test_labels.to_csv('./conv1d_sub.csv', index=False)
# !/home/jupyter/.local/bin/kaggle competitions submit jigsaw-toxic-comment-classification-challenge -f ./conv1.csv -m 'Baseline sub'
|
notebooks/Experiments_Text_Classification_WV.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="fSIfBsgi8dNK" colab_type="code" colab={}
#@title Copyright 2020 Google LLC. { display-mode: "form" }
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="aV1xZ1CPi3Nw" colab_type="text"
# <table class="ee-notebook-buttons" align="left"><td>
# <a target="_blank" href="http://colab.research.google.com/github/google/earthengine-api/blob/master/python/examples/ipynb/Manifest_image_upload_demo.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a>
# </td><td>
# <a target="_blank" href="https://github.com/google/earthengine-api/blob/master/python/examples/ipynb/Manifest_image_upload_demo.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td></table>
# + [markdown] id="RPBL-XjRFNop" colab_type="text"
# # Uploading an image from tiles using a manifest
#
# This notebook demonstrates uploading a set of image tiles into a single asset using a manifest file. See [this doc](https://developers.google.com/earth-engine/image_manifest) for more details about manifest upload using the Earth Engine command line tool.
#
# 10-meter land cover images derived from Sentinel-2 ([reference](https://doi.org/10.1016/j.scib.2019.03.002)) from the [Finer Resolution Global Land Cover Mapping (FROM-GLC) website](http://data.ess.tsinghua.edu.cn/) are downloaded directly to a Cloud Storage bucket and uploaded to a single Earth Engine asset from there. A manifest file, described below, is used to configure the upload.
# + [markdown] id="K57gwmayH24H" colab_type="text"
# First, authenticate with Google Cloud, so you can access Cloud Storage buckets.
# + id="a0WqP4vKIM5v" colab_type="code" colab={}
from google.colab import auth
auth.authenticate_user()
# + [markdown] id="1tPSX8ABIB36" colab_type="text"
# ## Download to Cloud Storage
#
# Paths from [the provider website](http://data.ess.tsinghua.edu.cn/fromglc10_2017v01.html) are manually copied to a list object as demonstrated below. Download directly to a Cloud Storage bucket to which you can write.
# + id="TQGLIdH6IQmn" colab_type="code" colab={}
# URLs of a few tiles.
urls = [
'http://data.ess.tsinghua.edu.cn/data/fromglc10_2017v01/fromglc10v01_36_-120.tif',
'http://data.ess.tsinghua.edu.cn/data/fromglc10_2017v01/fromglc10v01_36_-122.tif',
'http://data.ess.tsinghua.edu.cn/data/fromglc10_2017v01/fromglc10v01_36_-124.tif',
'http://data.ess.tsinghua.edu.cn/data/fromglc10_2017v01/fromglc10v01_38_-120.tif',
'http://data.ess.tsinghua.edu.cn/data/fromglc10_2017v01/fromglc10v01_38_-122.tif',
'http://data.ess.tsinghua.edu.cn/data/fromglc10_2017v01/fromglc10v01_38_-124.tif'
]
# You need to have write access to this bucket.
bucket = 'your-bucket-folder'
# Pipe curl output to gsutil.
for f in urls:
filepath = bucket + '/' + f.split('/')[-1]
# !curl {f} | gsutil cp - {filepath}
# + [markdown] id="nIOsWbLf66F-" colab_type="text"
# ## Build the manifest file
#
# Build the manifest file from a dictionary. Turn the dictionary into JSON. Note the use of the `gsutil` tool to get a listing of files in a Cloud Storage bucket ([learn more about `gsutil`](https://cloud.google.com/storage/docs/gsutil)). Also note that the structure of the manifest is described in detail [here](https://developers.google.com/earth-engine/image_manifest#manifest-structure-reference). Because the data are categorical, a `MODE` pyramiding policy is specified. Learn more about how Earth Engine builds image pyramids [here](https://developers.google.com/earth-engine/scale).
# + id="DPddpXYrJlap" colab_type="code" colab={}
# List the contents of the cloud folder.
# cloud_files = !gsutil ls {bucket + '/*.tif'}
# Get the list of source URIs from the gsutil output.
sources_uris = [{'uris': [f]} for f in cloud_files]
asset_name = 'path/to/your/asset'
# The enclosing object for the asset.
asset = {
'name': asset_name,
'tilesets': [
{
'sources': sources_uris
}
],
'bands': [
{
'id': 'cover_code',
'pyramiding_policy': 'MODE',
'missing_data': {
'values': [0]
}
}
]
}
import json
print(json.dumps(asset, indent=2))
# + [markdown] id="D2j6_TbCUiwZ" colab_type="text"
# Inspect the printed JSON for errors. If the JSON is acceptable, write it to a file and ensure that the file matches the printed JSON.
# + id="frZyXUDnFHVv" colab_type="code" colab={}
file_name = 'gaia_manifest.json'
with open(file_name, 'w') as f:
json.dump(asset, f, indent=2)
# + [markdown] id="k9WBqTW6XAwn" colab_type="text"
# Inspect the written file for errors.
# + id="wjunR9SLWn2A" colab_type="code" colab={}
# !cat {file_name}
# + [markdown] id="4MWm6WWbXG9G" colab_type="text"
# ## Upload to Earth Engine
#
# If you are able to `cat` the written file, run the upload to Earth Engine. First, import the Earth Engine library, authenticate and initialize.
# + id="hLFVQeDPXPE0" colab_type="code" colab={}
import ee
ee.Authenticate()
ee.Initialize()
# + id="A3ztutjFYqmt" colab_type="code" colab={}
# Do the upload.
# !earthengine upload image --manifest {file_name}
# + [markdown] id="vELn42MrZxwY" colab_type="text"
# ## Visualize the uploaded image with folium
#
# This is what [FROM-GLC](http://data.ess.tsinghua.edu.cn/) says about the classification system:
#
# | Class | Code |
# | ------------- | ------------- |
# | Cropland | 10 |
# | Forest | 20 |
# | Grassland | 30 |
# | Shrubland | 40 |
# | Wetland | 50 |
# | Water | 60 |
# | Tundra | 70 |
# | Impervious | 80 |
# | Bareland | 90 |
# | Snow/Ice | 100 |
#
# Use a modified FROM-GLC palette to visualize the results.
# + id="mKQOEbkvPAS0" colab_type="code" colab={}
palette = [
'a3ff73', # farmland
'267300', # forest
'ffff00', # grassland
'70a800', # shrub
'00ffff', # wetland
'005cff', # water
'004600', # tundra
'c500ff', # impervious
'ffaa00', # bare
'd1d1d1', # snow, ice
]
vis = {'min': 10, 'max': 100, 'palette': palette}
ingested_image = ee.Image('projects/ee-nclinton/assets/fromglc10_demo')
map_id = ingested_image.getMapId(vis)
import folium
map = folium.Map(location=[37.6413, -122.2582])
folium.TileLayer(
tiles=map_id['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
overlay=True,
name='fromglc10_demo',
).add_to(map)
map.add_child(folium.LayerControl())
map
|
python/examples/ipynb/Uploading_image_tiles_as_a_single_asset_using_a_manifest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
import collections
import numpy as py
import pandas as pd
import matplotlib.pyplot as pp
# %matplotlib inline
# -
nephews = ["Huey", "Duey", "Lewie"]
nephews
len(nephews)
nephews[0]
nephews[1]
nephews[2]
nephews[3]
print(nephews[-1])
print(nephews[-2])
for i in range(3):
nephews[i] = nephews[i] + "Duck"
nephews
for i in range(3):
nephews[i] = nephews[i] + "Zumran"
#gives hueyduckzumran dueyduckzumran and lewieduckzumran
nephews
for i in range(3):
nephews[i] = nephews[i] - "umran"
mix_it_up = [1,[2,3], 'Alpha']
mix_it_up
nephews.append('April Duck')
nephews
nephews.append(mix_it_up)
#can append lists within lists
nephews
ducks = nephews + ["zumranduck"]
#also appends it to teh list same way
ducks
ducks.extend(["Johnduck", "Peterduck"])
ducks
ducks.extend(["Wow", "Now"])
ducks
ducks.insert(0, "zahin")
ducks
ducks.insert(2, "zumran")
ducks
ducks.remove("zahin")
ducks
del ducks[0]
ducks
del ducks[1]
ducks
del ducks[2]
del ducks[3]
ducks
ducks.sort()
ducks
|
working with lists.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''base'': conda)'
# name: python3
# ---
# # Leaflet cluster map of talk locations
#
# Run this from the _talks/ directory, which contains .md files of all your talks. This scrapes the location YAML field from each .md file, geolocates it with geopy/Nominatim, and uses the getorg library to output data, HTML, and Javascript for a standalone cluster map.
# !pip install getorg --upgrade
import glob
import getorg
from geopy import Nominatim
g = glob.glob("*.md")
geocoder = Nominatim(user_agent='my_application')
location_dict = {}
location = ""
permalink = ""
title = ""
# +
location_count = {}
for file in g:
with open(file, 'r') as f:
lines = f.read()
if lines.find('location: "') > 1:
loc_start = lines.find('location: "') + 11
lines_trim = lines[loc_start:]
loc_end = lines_trim.find('"')
location = lines_trim[:loc_end]
try:
location_count[location] += 1
except KeyError:
location_count[location] = 1
if location_count[location] > 1:
location_dict[f'{location} {location_count[location]}'] = geocoder.geocode(location)
print(location, "\n", location_dict[f'{location} {location_count[location]}'])
else:
location_dict[location] = geocoder.geocode(location)
print(location, "\n", location_dict[location])
# -
location_dict
m = getorg.orgmap.create_map_obj()
getorg.orgmap.output_html_cluster_map(location_dict, folder_name="../talkmap", hashed_usernames=False)
|
_all_talks_for_maps/talkmap.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# # Lotka-Volterra Work-Precision Diagrams
#
# Adapted from
# [SciMLBenchmarks.jl Lotka-Volterra benchmark](https://benchmarks.sciml.ai/html/NonStiffODE/LotkaVolterra_wpd.html).
# Imports
using LinearAlgebra, Statistics
using OrdinaryDiffEq, ParameterizedFunctions, ODEInterfaceDiffEq, LSODA, Sundials, DiffEqDevTools
using Plots
using ProbNumDiffEq
# Plotting theme
theme(:dao;
linewidth=8,
linealpha=0.7,
markersize=5,
markerstrokewidth=0.5,
legend=:outerright,
)
# ## Problem Definition
# +
# Problem definition and reference solution
f = @ode_def LotkaVolterra begin
dx = a*x - b*x*y
dy = -c*y + d*x*y
end a b c d
p = [1.5,1.0,3.0,1.0]
prob = ODEProblem(f,[1.0;1.0],(0.0,10.0),p)
abstols = 1.0 ./ 10.0 .^ (6:13)
reltols = 1.0 ./ 10.0 .^ (3:10);
sol = solve(prob,Vern7(),abstol=1/10^14,reltol=1/10^14)
test_sol = TestSolution(sol)
plot(sol, title="Lotka-Volterra Solution", linewidth=2)
# -
# ## Low Order
setups = [Dict(:alg=>DP5())
#Dict(:alg=>ode45()) # fail
Dict(:alg=>dopri5())
Dict(:alg=>Tsit5())
Dict(:alg=>Vern6())
Dict(:alg=>EK0(order=4, smooth=false))
Dict(:alg=>EK1(order=5, smooth=false))
]
wp = WorkPrecisionSet(prob,abstols,reltols,setups;appxsol=test_sol,save_everystep=false,maxiters=100000,numruns=10)
plot(wp)
# ### Interpolation Error
setups = [Dict(:alg=>DP5())
#Dict(:alg=>ode45())
Dict(:alg=>dopri5())
Dict(:alg=>Tsit5())
#Dict(:alg=>Vern6()) # does not work currently for some reason
Dict(:alg=>EK0(order=4))
Dict(:alg=>EK1(order=5))
]
wp = WorkPrecisionSet(prob,abstols,reltols,setups;
appxsol=test_sol,maxiters=1000000,error_estimate=:L2,dense_errors=true,numruns=10)
plot(wp)
# ## Higher Order
setups = [Dict(:alg=>DP8())
#Dict(:alg=>ode78()) # fails
Dict(:alg=>Vern7())
Dict(:alg=>Vern8())
Dict(:alg=>dop853())
Dict(:alg=>Vern6())
Dict(:alg=>EK1(order=6, smooth=false))
Dict(:alg=>EK1(order=7, smooth=false))
Dict(:alg=>EK1(order=8, smooth=false))
]
wp = WorkPrecisionSet(prob,abstols,reltols,setups;appxsol=test_sol,save_everystep=false,maxiters=100000,numruns=10)
plot(wp)
setups = [Dict(:alg=>odex())
Dict(:alg=>ddeabm())
Dict(:alg=>Vern7())
Dict(:alg=>Vern8())
Dict(:alg=>CVODE_Adams())
Dict(:alg=>lsoda())
Dict(:alg=>Vern6())
Dict(:alg=>ARKODE(Sundials.Explicit(),order=6))
Dict(:alg=>EK1(order=6, smooth=false))
Dict(:alg=>EK1(order=7, smooth=false))
Dict(:alg=>EK1(order=8, smooth=false))
]
wp = WorkPrecisionSet(prob,abstols,reltols,setups;
appxsol=test_sol,save_everystep=false,maxiters=100000,numruns=10)
plot(wp)
# ### Interpolation Error
setups = [Dict(:alg=>DP8())
#Dict(:alg=>ode78())
# For some reasons the Vern methods do not work right now
#Dict(:alg=>Vern7())
#Dict(:alg=>Vern8())
#Dict(:alg=>Vern6())
Dict(:alg=>EK1(order=6))
Dict(:alg=>EK1(order=7))
Dict(:alg=>EK1(order=8))
]
wp = WorkPrecisionSet(prob,abstols,reltols,setups;
appxsol=test_sol,dense=true,maxiters=100000,error_estimate=:L2,numruns=10)
plot(wp)
|
benchmarks/nonstiff-lotkavolterra.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## LiDAR Basics with VLP-16
#
# LIDAR—Light Detection and Ranging—is used to find the precise distance of objects in relation to us. Velodyne LiDAR sensors use time-of-flight methodology for doing this.
#
# When a laser pulse is emited, its time-of-shooting and direction are registered. The laser pulse travels through air until it hits an obstacle which reflects some of the energy. The time-of-acquisition and power received are registered by sensor after recieving the portion of energy. The spherical coordinates of the obstacle is calculated using time-of-acquisition which is returned along with power received(as relectance) after each scan.
#
# As LiDAR sensor returns reading in spherical coordinates, let's brush up with the spherical coordinate system.
# ___
# To know more on LiDAR's history and how it works this [blog](https://news.voyage.auto/an-introduction-to-lidar-the-key-self-driving-car-sensor-a7e405590cff) by <NAME> is helpful.
# ### Spherical Coordinate System
# In a spherical coordinate system, a point is defined by a distance and two angles. To represent the two angles we use azimuth($\theta$) and polar angle($\phi$) convention. Thus a point is defined by $(\text{r}, \theta, \phi)$.
#
# : radial distance r, polar angle θ (theta), and azimuthal angle φ (phi). The symbol ρ (rho) is often used instead of r. This diagram is by Andeggs from Wikimedia Commons.")
#
# As you can see from above diagram, the azimuth angle is in X-Y plane measured from X-axis and polar angle is in Z-Y plane measured from Z axis.
#
# From above diagram, we can get the following equations for converting a cartesian coordinate to spherical coordinates.
#
# <math>\begin{align} r&=\sqrt{x^2 + y^2 + z^2} \\ \theta &= \arccos\frac{z}{\sqrt{x^2 + y^2 + z^2}} = \arccos\frac{z}{r} \\ \varphi &= \arctan \frac{y}{x} \end{align}</math>
#
# We can derive cartesian coordinates from spherical coordinates using below equations.
#
# <math>\begin{align} x&=r \, \sin\theta \, \cos\varphi \\ y&=r \, \sin\theta \, \sin\varphi \\ z&=r \, \cos\theta\end{align}</math>
#
# ___
# Read [more](https://en.wikipedia.org/wiki/Spherical_coordinate_system) about Spherical coordinate system at Wikipedia.
# ## VLP-16 coordinate system
# Velodyne VLP-16 returns reading in spherical coordinates. But there is a slight difference from the above discussed convention.
#
# In sensor coordinate system, a point is defined by (radius $\text{r}$, elevation $\omega$, azimuth $\alpha$). Elevation angle, $\omega$ is in Z-Y plane measured from Y-axis. Azimuth angle, $\alpha$ is in X-Y plane measured from Y-axis. Below is the diagram.
#
# 
#
# Cartesian coordinates can be derived by following equations.
#
# <math>\begin{align} x&=r \, \cos\omega \, \sin\alpha \\ y&=r \, \cos\omega \, \cos\alpha \\ z&=r \, \sin\omega\end{align}</math>
#
# A computation is necessary to convert the spherical data from the sensor to Cartesian coordinates using above equations. This can be done using a ros package.
#
# 
# Above diagram shows the coordinate system of sensor mounted on a car.
# ___
# This [manual](https://velodynelidar.com/docs/manuals/63-9243%20REV%20D%20MANUAL,USERS,VLP-16.pdf) is a good start to know more about Velodyne VLP-16.
# ## Data format
# An unstructured point cloud is returned after each scan by sensor. Even though LiDAR returns reading in spherical coordinates, widely used coordinate system is Cartesian.
#
# A point in point cloud is defined by it's coordinates and reflectance. Reflectance tells us the reflectivity of the surface. A zero value in reflectance denotes that the laser pulse didn't result in a measurement.
#
# There are many formats to store and process point clouds like PCD, PCL but we can treat point cloud as a `Numpy` array. Each element of the array will have `(x,y,z,r)`. For processing point cloud we can use `Numpy`.
#
|
tutorial/LiDAR Basics with VLP-16.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocess data with TensorFlow Transform
# credits: [TFX Tutorials Transform](https://www.tensorflow.org/tfx/transform/get_started)
# !pip install tensorflow-transform
# ### Preprocessing function
import tensorflow as tf
import tensorflow_transform as tft
import tensorflow_transform.beam as tft_beam
def preprocessing_fn(inputs):
"""Preprocessing function transforms each of three inputs in different ways
Note:
Input `inputs` must be a dictionary of `Tensor` or `SparseTensor`
Args:
inputs: Dictionary of `Tensor` or `SparseTensor` of the raw data
Returns:
A dictionary of `Tensor` or `SparseTensor` containing the transformed values
"""
x = inputs['x']
y = inputs['y']
s = inputs['s']
x_centered = x - tft.mean(x)
y_normalized = tft.scale_to_0_1(y)
s_integerized = tft.compute_and_apply_vocabulary(s)
x_centered_times_y_normalized = x_centered * y_normalized
return {
'x_centered': x_centered,
'y_normalized': y_normalized,
'x_centered_times_y_normalized': x_centered_times_y_normalized,
's_integerized': s_integerized
}
raw_data = [
{'x': 1, 'y': 1, 's': 'hello'},
{'x': 2, 'y': 2, 's': 'world'},
{'x': 3, 'y': 3, 's': 'hello'}
]
# +
from tensorflow_transform.tf_metadata import dataset_metadata
from tensorflow_transform.tf_metadata import schema_utils
raw_data_metadata = dataset_metadata.DatasetMetadata(
schema_utils.schema_from_feature_spec({
'x': tf.io.FixedLenFeature([], tf.int64),
'y': tf.io.FixedLenFeature([], tf.int64),
's': tf.io.FixedLenFeature([], tf.string),
})
)
# +
import tensorflow_transform.beam as tft_beam
transformed_dataset, transform_fn = (
(raw_data, raw_data_metadata) | tft_beam.AnalyzeAndTransformDataset(
preprocessing_fn))
# transformed_data, transformed_metadata = transformed_dataset
# -
transformed_data
|
Learning_TFX/TF_Tutorial/TF Transform .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''KickstarterSuccess-Diw2u4NI'': pipenv)'
# name: python385jvsc74a57bd0849be1930e84a28de76efd7f21d926cca918e7072377246a7749ae7f6697b53e
# ---
# +
# Import everything
import sys
import pandas as pd
import numpy as np
import sklearn
from sklearn.model_selection import train_test_split
from category_encoders import OneHotEncoder, OrdinalEncoder
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.metrics import accuracy_score
import xgboost
from xgboost import XGBClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
import pickle
import joblib
# -
df = pd.read_csv('KickstarterCleanedv4.csv')
df.head()
df.drop(columns=['Unnamed: 0','goal','spotlight'],inplace=True)
df.columns
df = df.drop_duplicates()
df.shape
# +
#df.to_csv('Kickstarter_FinalCleaned.csv')
# +
# Extracting the target and feature matrix
target = 'state'
y = df[target]
X = df.drop(columns=target)
print(X.shape)
print(y.shape)
# -
# Splitting into train test split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = .4)
print(X_train.shape,X_test.shape)
print(y_train.shape,y_test.shape)
# +
#Baseline
print('baseline accuracy', y.value_counts(normalize=True).max())
# +
# Random Forest
model_rf = make_pipeline(OrdinalEncoder(),
SimpleImputer(strategy="mean"),
RandomForestClassifier( n_jobs=-1, random_state=42))
# -
# Decision Tree
model_dt = make_pipeline(OrdinalEncoder(),
SimpleImputer(strategy="mean"),
DecisionTreeClassifier(random_state=42))
# +
# XGBoost
model_xgb = make_pipeline(OrdinalEncoder(),
SimpleImputer(strategy="mean"),
XGBClassifier(random_state=42))
# +
# Gradient Boost
model_gb = make_pipeline(OrdinalEncoder(),
SimpleImputer(strategy="mean"),
GradientBoostingClassifier(random_state=42))
# -
model_rf.fit(X_train,y_train)
model_dt.fit(X_train,y_train)
model_xgb.fit(X_train,y_train)
model_gb.fit(X_train,y_train)
#Check Metrics on training
print('model_dt accuracy score', accuracy_score(y_train, model_dt.predict(X_train)))
print('model_rf accuracy score', accuracy_score(y_train, model_rf.predict(X_train)))
print('model_xgb accuracy score', accuracy_score(y_train, model_xgb.predict(X_train)))
print('model_gb accuracy score', accuracy_score(y_train, model_gb.predict(X_train)))
# +
# Metrics with test data
# print('model_dt accuracy score', accuracy_score(y_test, model_dt.predict(X_test)))
# print('model_rf accuracy score', accuracy_score(y_test, model_rf.predict(X_test)))
# print('model_xgb accuracy score', accuracy_score(y_test, model_xgb.predict(X_test)))
# print('model_gb accuracy score', accuracy_score(y_test, model_gb.predict(X_test)))
# -
# saving models using pickle
saved_model_rf = pickle.dumps(model_rf)
saved_model_xgb = pickle.dumps(model_xgb)
# +
joblib_file = "joblib_RF_Model.pkl"
joblib.dump(model_rf, 'assets/model_rf')
# -
joblib_file = "joblib_XGB_Model.pkl"
joblib.dump(model_xgb, 'assets/model_xgb')
# +
#Testing if model saved and working correctly
# # Load from file
# load_xgb_model = joblib.load('assets/model_xgb')
# load_xgb_model
# +
# # Use the Reloaded Joblib Model to
# # Calculate the accuracy score and predict target values
# # Calculate the Score
# score = load_xgb_model.score(X_test, y_test)
# # Print the Score
# print("Test score: {0:.2f} %".format(100 * score))
# # Predict the Labels using the reloaded Model
# Ypredict = load_xgb_model.predict(X_test)
# Ypredict
# -
|
model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: drlnd
# language: python
# name: drlnd
# ---
# # Tennis
#
# ---
#
# In this notebook, we will train MADDPG(Multi Agent Deep Deterministic Policy Gradient) agents to control rackets in Unity's Tennis environment.
#
# ### 1. Start the Environment
#
# We begin by importing some necessary packages and starting Unity environment. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed necessary packages.
# +
import sys
sys.path.append('../')
from unityagents import UnityEnvironment
from collections import deque
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import utils.config
import pprint
import torch
env = UnityEnvironment(file_name="Tennis_Linux/Tennis.x86", no_graphics=False)
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
# -
# ### 2. Create instance of agent
#
# Agent's hyperparameters are saved and loaded from config.py file in utils folder. Current values were selected after coarse hyperparameter tuing. But you can try different hyperparameter values if you want.
#
# If you just want to see the trained agent jump to cell 5.
#
# +
from agents.maddpg_agent import MADDPG
# Load parameters from file
params = utils.config.HYPERPARAMS['Tennis']
hparams = params['agent']
# Create agent instance
agent = MADDPG(hparams)
print("Created agent with following hyperparameter values:")
pprint.pprint(hparams)
# -
# ### 3. Train an agent!
# +
# Reset and set environment to training mode
env_info = env.reset(train_mode=True)[brain_name]
# Maximum number of training episodes
n_episodes = params['n_episodes']
# List containing scores from each episode
scores = []
# Store last 100 scores
scores_window = deque(maxlen=params['scores_window_size'])
# Flag to indicate environment is solved
solved = False
# Train loop
for i_episode in range(1, n_episodes+1):
# Reset environment
env_info = env.reset(train_mode=True)[brain_name]
# Observe current state
states = env_info.vector_observations
# Reset score
agent_scores = np.zeros(num_agents)
# Reset agent
agent.reset()
# Loop each episode
while True:
# Select action
actions = agent.act(states)
# Take action
env_info = env.step(actions)[brain_name]
# Get next state, reward and done
next_states = env_info.vector_observations
rewards = env_info.rewards
dones = env_info.local_done
# Store experience and learn
agent.step(states, actions, rewards, next_states, dones)
# State transition
states = next_states
# Update total score
agent_scores += rewards
# Exit loop if episode finished
if np.any(dones):
break
# Save most recent score
scores_window.append(np.max(agent_scores))
scores.append([np.max(agent_scores), np.mean(scores_window)])
# Print learning progress
print('\rEpisode {}\tMax Score: {:.6f}\tAverage Score: {:.6f}'.format(i_episode, np.max(agent_scores), np.mean(scores_window)), end="")
if i_episode % params['scores_window_size'] == 0:
print('\rEpisode {}\tMax Score: {:.6f}\tAverage Score: {:.6f}'.format(i_episode, np.max(agent_scores), np.mean(scores_window)))
if np.mean(scores_window)>=params['solve_score']:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_window)))
filename = "{:s}_lra{:.0E}_lrc{:.0E}_tau{:.0E}_nstart{:.1f}_ndecay{}_solved{:d}"
filename = filename.format(hparams['agent_name'], hparams['lr_actor'], hparams['lr_critic'],
hparams['tau'], hparams['noise_start'], hparams['noise_decay'], i_episode-100)
solved = True
agent.save(filename)
break
# -
# ### 4. Save and plot the score
# +
# Save score if environment is solved
df = pd.DataFrame(scores,columns=['scores','average_scores'])
if solved:
df.to_csv('scores/{:s}.csv'.format(filename))
# Plot scores
plt.figure(figsize=(10,5))
plt.axhline(params['solve_score'], color='red', lw=1, alpha=0.3)
plt.plot( df.index, 'scores', data=df, color='lime', lw=1, label="score", alpha=0.4)
plt.plot( df.index, 'average_scores', data=df, color='green', lw=2, label="average score")
# Set labels and legends
plt.xlabel('Episode')
plt.xlim(0, len(df.index))
plt.ylabel('Score')
if solved:
plt.title(filename)
plt.grid(True, alpha=0.3, linestyle='--')
plt.legend()
plt.show()
# Save figure if environment is solved
if solved:
plt.savefig('docs/plots/{:s}.png'.format(filename), bbox_inches='tight')
# -
# ### 5. Watch smart agent
#
# If you skipped training and just want to see trained agent, specify filename for pre-trained network model.
#
# +
# Speed (False: Real time, True: Fast)
train_mode = False
# reset the environment
env_info = env.reset(train_mode=train_mode)[brain_name]
# Load learned model weight.
#filename = 'MADDPG_lra1E-03_lrc1E-03_tau1E-01_nstart7.0_ndecay0.999_solved540' #Uncomment if skipped training!
agent.load(filename)
# Number of episodes to run
n_episodes = 1
# Run loop
for i_episode in range(1, n_episodes+1):
# Reset environment
env_info = env.reset(train_mode=train_mode)[brain_name]
# Observe current state
states = env_info.vector_observations
# Reset score and done flag
score = np.zeros(num_agents)
# Episode loop
while True:
# Select action with greedy policy
actions = agent.act(states, add_noise=False)
# Take action
env_info = env.step(actions)[brain_name]
# Observe the next state
next_states = env_info.vector_observations
# Get the reward
rewards = env_info.rewards
# Check if episode is finished
dones = env_info.local_done
# State transition
states = next_states
# Update total score
score += rewards
# Exit loop if episode finished
if np.any(dones):
break
# Print episode summary
print('Episode {} Score:{:.4f}'.format(i_episode, np.mean(score)))
# -
# When finished, close the environment.
#
env.close()
|
Tennis/Tennis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import missingno as msno
df = pd.read_csv('../../fixtures/cleaned_data/2019RedZoneStats_cleaned.csv')
# -
print(df)
# see how many rows and columns are in this dataset
shape_info = df.shape
print('This dataset contains {} rows and {} columns'
.format(shape_info[0],
shape_info[1]))
# look at first 5 rows of data
df.head()
# look at last 5 rows of data
df.tail()
# see list of all columns
list(df)
# +
#removing columns don't need
df = df.drop(['FantasyPointsPerGamePPR', 'Rank'], axis=1)
# +
new_shape = df.shape[1]
print('{} columns have been removed from the dataset'
.format(
abs(new_shape-shape_info[1])))
# +
shape_info = df.shape
print('This dataset evaluates {} running backs based on data from {} columns'
.format(shape_info[0],
shape_info[1]))
# -
display(df)
# +
# rename columns to label as red zone specific stats
# create a dictionary
# key = old name
# value = new name
dict = {'Opponent':'OpponentRZ',
'PassingYards':'PassingYardsRZ',
'PassingTouchdowns':'PassingTouchdownsRZ',
'PassingInterceptions':'PassingInterceptionsRZ',
'RushingYards':'RushingYardsRZ',
'RushingTouchdowns':'RushingTouchdownsRZ',
'Receptions':'ReceptionsRZ',
'ReceivingYards':'ReceivingYardsRZ',
'ReceivingTouchdowns':'ReceivingTouchdownsRZ',
'Sacks':'SacksRZ',
'Interceptions':'InterceptionsRZ',
'FumblesForced':'FumblesForcedRZ',
'FumblesRecovered':'FumlbesRecoveredRZ',
'FantasyPointsPPR':'FantasyPointsRZ'}
# call rename () method
df.rename(columns=dict,
inplace=True)
# print Data frame after rename columns
display(df)
# -
list(df)
#check for NAs
msno.matrix(df)
# There are no NA values in the data but there are a lot of 0s
df['Season'] = 2019
list(df)
df.head()
# starting to explore data and see which team has the most fantasy points in the red zone for 2019 season
df.groupby('Team', as_index=False).agg({"FantasyPointsRZ": "sum"}).sort_values(by=['FantasyPointsRZ'],ascending=False)
# what position has the most fantasy points in the red zone for the 2019 season?
df.groupby('Position', as_index=False).agg({"FantasyPointsRZ": "sum"}).sort_values(by=['FantasyPointsRZ'],ascending=False)
# which defense allowed the most points in the red zone for 2019 season?
df.groupby('OpponentRZ', as_index=False).agg({"FantasyPointsRZ": "sum"}).sort_values(by=['FantasyPointsRZ'],ascending=False)
# which week in the 2019 season had the most fantasy points in the red zone?
df.groupby('Week', as_index=False).agg({"FantasyPointsRZ": "sum"}).sort_values(by=['FantasyPointsRZ'],ascending=False)
# which players had the most fantasy points in the red zone for the 2019 season
df.groupby('Name', as_index=False).agg({"FantasyPointsRZ": "sum"}).sort_values(by=['FantasyPointsRZ'],ascending=False)
pd.set_option("display.max_rows", None, "display.max_columns", None)
df.groupby(['Team', 'Position'])['PassingTouchdownsRZ'].sum()
# pivot data and show 2019 red zone fantasy points by team and position
pd.set_option("display.max_rows", None, "display.max_columns", None)
points_by_team = pd.pivot_table(df, index = ["Team","Position"], values = "FantasyPointsRZ", aggfunc = "sum")
print(points_by_team)
# export data frame to a csv
df.to_csv('2019RedZoneStats_cleaned.csv')
|
notebooks/cleaning/2019RZcleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building your Deep Neural Network: Step by Step
#
# Welcome to your week 4 assignment (part 1 of 2)! You have previously trained a 2-layer Neural Network (with a single hidden layer). This week, you will build a deep neural network, with as many layers as you want!
#
# - In this notebook, you will implement all the functions required to build a deep neural network.
# - In the next assignment, you will use these functions to build a deep neural network for image classification.
#
# **After this assignment you will be able to:**
# - Use non-linear units like ReLU to improve your model
# - Build a deeper neural network (with more than 1 hidden layer)
# - Implement an easy-to-use neural network class
#
# **Notation**:
# - Superscript $[l]$ denotes a quantity associated with the $l^{th}$ layer.
# - Example: $a^{[L]}$ is the $L^{th}$ layer activation. $W^{[L]}$ and $b^{[L]}$ are the $L^{th}$ layer parameters.
# - Superscript $(i)$ denotes a quantity associated with the $i^{th}$ example.
# - Example: $x^{(i)}$ is the $i^{th}$ training example.
# - Lowerscript $i$ denotes the $i^{th}$ entry of a vector.
# - Example: $a^{[l]}_i$ denotes the $i^{th}$ entry of the $l^{th}$ layer's activations).
#
# Let's get started!
# ## 1 - Packages
#
# Let's first import all the packages that you will need during this assignment.
# - [numpy](www.numpy.org) is the main package for scientific computing with Python.
# - [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
# - dnn_utils provides some necessary functions for this notebook.
# - testCases provides some test cases to assess the correctness of your functions
# - np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work. Please don't change the seed.
# +
import numpy as np
import h5py
import matplotlib.pyplot as plt
from testCases_v4 import *
from dnn_utils_v2 import sigmoid, sigmoid_backward, relu, relu_backward
# %matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# %load_ext autoreload
# %autoreload 2
np.random.seed(1)
# -
# ## 2 - Outline of the Assignment
#
# To build your neural network, you will be implementing several "helper functions". These helper functions will be used in the next assignment to build a two-layer neural network and an L-layer neural network. Each small helper function you will implement will have detailed instructions that will walk you through the necessary steps. Here is an outline of this assignment, you will:
#
# - Initialize the parameters for a two-layer network and for an $L$-layer neural network.
# - Implement the forward propagation module (shown in purple in the figure below).
# - Complete the LINEAR part of a layer's forward propagation step (resulting in $Z^{[l]}$).
# - We give you the ACTIVATION function (relu/sigmoid).
# - Combine the previous two steps into a new [LINEAR->ACTIVATION] forward function.
# - Stack the [LINEAR->RELU] forward function L-1 time (for layers 1 through L-1) and add a [LINEAR->SIGMOID] at the end (for the final layer $L$). This gives you a new L_model_forward function.
# - Compute the loss.
# - Implement the backward propagation module (denoted in red in the figure below).
# - Complete the LINEAR part of a layer's backward propagation step.
# - We give you the gradient of the ACTIVATE function (relu_backward/sigmoid_backward)
# - Combine the previous two steps into a new [LINEAR->ACTIVATION] backward function.
# - Stack [LINEAR->RELU] backward L-1 times and add [LINEAR->SIGMOID] backward in a new L_model_backward function
# - Finally update the parameters.
#
# <img src="images/final outline.png" style="width:800px;height:500px;">
# <caption><center> **Figure 1**</center></caption><br>
#
#
# **Note** that for every forward function, there is a corresponding backward function. That is why at every step of your forward module you will be storing some values in a cache. The cached values are useful for computing gradients. In the backpropagation module you will then use the cache to calculate the gradients. This assignment will show you exactly how to carry out each of these steps.
# ## 3 - Initialization
#
# You will write two helper functions that will initialize the parameters for your model. The first function will be used to initialize parameters for a two layer model. The second one will generalize this initialization process to $L$ layers.
#
# ### 3.1 - 2-layer Neural Network
#
# **Exercise**: Create and initialize the parameters of the 2-layer neural network.
#
# **Instructions**:
# - The model's structure is: *LINEAR -> RELU -> LINEAR -> SIGMOID*.
# - Use random initialization for the weight matrices. Use `np.random.randn(shape)*0.01` with the correct shape.
# - Use zero initialization for the biases. Use `np.zeros(shape)`.
# +
# GRADED FUNCTION: initialize_parameters
def initialize_parameters(n_x, n_h, n_y):
"""
Argument:
n_x -- size of the input layer
n_h -- size of the hidden layer
n_y -- size of the output layer
Returns:
parameters -- python dictionary containing your parameters:
W1 -- weight matrix of shape (n_h, n_x)
b1 -- bias vector of shape (n_h, 1)
W2 -- weight matrix of shape (n_y, n_h)
b2 -- bias vector of shape (n_y, 1)
"""
np.random.seed(1)
### START CODE HERE ### (≈ 4 lines of code)
W1 = np.random.randn(n_h,n_x)*0.01
b1 = np.zeros(shape=(n_h, 1))
W2 = np.random.randn(n_y, n_h) * 0.01
b2 = np.zeros(shape=(n_y, 1))
### END CODE HERE ###
assert(W1.shape == (n_h, n_x))
assert(b1.shape == (n_h, 1))
assert(W2.shape == (n_y, n_h))
assert(b2.shape == (n_y, 1))
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return parameters
# -
parameters = initialize_parameters(3,2,1)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
# **Expected output**:
#
# <table style="width:80%">
# <tr>
# <td> **W1** </td>
# <td> [[ 0.01624345 -0.00611756 -0.00528172]
# [-0.01072969 0.00865408 -0.02301539]] </td>
# </tr>
#
# <tr>
# <td> **b1**</td>
# <td>[[ 0.]
# [ 0.]]</td>
# </tr>
#
# <tr>
# <td>**W2**</td>
# <td> [[ 0.01744812 -0.00761207]]</td>
# </tr>
#
# <tr>
# <td> **b2** </td>
# <td> [[ 0.]] </td>
# </tr>
#
# </table>
# ### 3.2 - L-layer Neural Network
#
# The initialization for a deeper L-layer neural network is more complicated because there are many more weight matrices and bias vectors. When completing the `initialize_parameters_deep`, you should make sure that your dimensions match between each layer. Recall that $n^{[l]}$ is the number of units in layer $l$. Thus for example if the size of our input $X$ is $(12288, 209)$ (with $m=209$ examples) then:
#
# <table style="width:100%">
#
#
# <tr>
# <td> </td>
# <td> **Shape of W** </td>
# <td> **Shape of b** </td>
# <td> **Activation** </td>
# <td> **Shape of Activation** </td>
# <tr>
#
# <tr>
# <td> **Layer 1** </td>
# <td> $(n^{[1]},12288)$ </td>
# <td> $(n^{[1]},1)$ </td>
# <td> $Z^{[1]} = W^{[1]} X + b^{[1]} $ </td>
#
# <td> $(n^{[1]},209)$ </td>
# <tr>
#
# <tr>
# <td> **Layer 2** </td>
# <td> $(n^{[2]}, n^{[1]})$ </td>
# <td> $(n^{[2]},1)$ </td>
# <td>$Z^{[2]} = W^{[2]} A^{[1]} + b^{[2]}$ </td>
# <td> $(n^{[2]}, 209)$ </td>
# <tr>
#
# <tr>
# <td> $\vdots$ </td>
# <td> $\vdots$ </td>
# <td> $\vdots$ </td>
# <td> $\vdots$</td>
# <td> $\vdots$ </td>
# <tr>
#
# <tr>
# <td> **Layer L-1** </td>
# <td> $(n^{[L-1]}, n^{[L-2]})$ </td>
# <td> $(n^{[L-1]}, 1)$ </td>
# <td>$Z^{[L-1]} = W^{[L-1]} A^{[L-2]} + b^{[L-1]}$ </td>
# <td> $(n^{[L-1]}, 209)$ </td>
# <tr>
#
#
# <tr>
# <td> **Layer L** </td>
# <td> $(n^{[L]}, n^{[L-1]})$ </td>
# <td> $(n^{[L]}, 1)$ </td>
# <td> $Z^{[L]} = W^{[L]} A^{[L-1]} + b^{[L]}$</td>
# <td> $(n^{[L]}, 209)$ </td>
# <tr>
#
# </table>
#
# Remember that when we compute $W X + b$ in python, it carries out broadcasting. For example, if:
#
# $$ W = \begin{bmatrix}
# j & k & l\\
# m & n & o \\
# p & q & r
# \end{bmatrix}\;\;\; X = \begin{bmatrix}
# a & b & c\\
# d & e & f \\
# g & h & i
# \end{bmatrix} \;\;\; b =\begin{bmatrix}
# s \\
# t \\
# u
# \end{bmatrix}\tag{2}$$
#
# Then $WX + b$ will be:
#
# $$ WX + b = \begin{bmatrix}
# (ja + kd + lg) + s & (jb + ke + lh) + s & (jc + kf + li)+ s\\
# (ma + nd + og) + t & (mb + ne + oh) + t & (mc + nf + oi) + t\\
# (pa + qd + rg) + u & (pb + qe + rh) + u & (pc + qf + ri)+ u
# \end{bmatrix}\tag{3} $$
# **Exercise**: Implement initialization for an L-layer Neural Network.
#
# **Instructions**:
# - The model's structure is *[LINEAR -> RELU] $ \times$ (L-1) -> LINEAR -> SIGMOID*. I.e., it has $L-1$ layers using a ReLU activation function followed by an output layer with a sigmoid activation function.
# - Use random initialization for the weight matrices. Use `np.random.randn(shape) * 0.01`.
# - Use zeros initialization for the biases. Use `np.zeros(shape)`.
# - We will store $n^{[l]}$, the number of units in different layers, in a variable `layer_dims`. For example, the `layer_dims` for the "Planar Data classification model" from last week would have been [2,4,1]: There were two inputs, one hidden layer with 4 hidden units, and an output layer with 1 output unit. This means `W1`'s shape was (4,2), `b1` was (4,1), `W2` was (1,4) and `b2` was (1,1). Now you will generalize this to $L$ layers!
# - Here is the implementation for $L=1$ (one layer neural network). It should inspire you to implement the general case (L-layer neural network).
# ```python
# if L == 1:
# parameters["W" + str(L)] = np.random.randn(layer_dims[1], layer_dims[0]) * 0.01
# parameters["b" + str(L)] = np.zeros((layer_dims[1], 1))
# ```
# +
# GRADED FUNCTION: initialize_parameters_deep
def initialize_parameters_deep(layer_dims):
"""
Arguments:
layer_dims -- python array (list) containing the dimensions of each layer in our network
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
Wl -- weight matrix of shape (layer_dims[l], layer_dims[l-1])
bl -- bias vector of shape (layer_dims[l], 1)
"""
np.random.seed(3)
parameters = {}
L = len(layer_dims) # number of layers in the network
for l in range(1, L):
### START CODE HERE ### (≈ 2 lines of code)
parameters['W' + str(l)] = np.random.randn(layer_dims[l], layer_dims[l - 1]) * 0.01
parameters['b' + str(l)] = np.zeros((layer_dims[l], 1))
### END CODE HERE ###
assert(parameters['W' + str(l)].shape == (layer_dims[l], layer_dims[l-1]))
assert(parameters['b' + str(l)].shape == (layer_dims[l], 1))
return parameters
# -
parameters = initialize_parameters_deep([5,4,3])
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
# **Expected output**:
#
# <table style="width:80%">
# <tr>
# <td> **W1** </td>
# <td>[[ 0.01788628 0.0043651 0.00096497 -0.01863493 -0.00277388]
# [-0.00354759 -0.00082741 -0.00627001 -0.00043818 -0.00477218]
# [-0.01313865 0.00884622 0.00881318 0.01709573 0.00050034]
# [-0.00404677 -0.0054536 -0.01546477 0.00982367 -0.01101068]]</td>
# </tr>
#
# <tr>
# <td>**b1** </td>
# <td>[[ 0.]
# [ 0.]
# [ 0.]
# [ 0.]]</td>
# </tr>
#
# <tr>
# <td>**W2** </td>
# <td>[[-0.01185047 -0.0020565 0.01486148 0.00236716]
# [-0.01023785 -0.00712993 0.00625245 -0.00160513]
# [-0.00768836 -0.00230031 0.00745056 0.01976111]]</td>
# </tr>
#
# <tr>
# <td>**b2** </td>
# <td>[[ 0.]
# [ 0.]
# [ 0.]]</td>
# </tr>
#
# </table>
# ## 4 - Forward propagation module
#
# ### 4.1 - Linear Forward
# Now that you have initialized your parameters, you will do the forward propagation module. You will start by implementing some basic functions that you will use later when implementing the model. You will complete three functions in this order:
#
# - LINEAR
# - LINEAR -> ACTIVATION where ACTIVATION will be either ReLU or Sigmoid.
# - [LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID (whole model)
#
# The linear forward module (vectorized over all the examples) computes the following equations:
#
# $$Z^{[l]} = W^{[l]}A^{[l-1]} +b^{[l]}\tag{4}$$
#
# where $A^{[0]} = X$.
#
# **Exercise**: Build the linear part of forward propagation.
#
# **Reminder**:
# The mathematical representation of this unit is $Z^{[l]} = W^{[l]}A^{[l-1]} +b^{[l]}$. You may also find `np.dot()` useful. If your dimensions don't match, printing `W.shape` may help.
# +
# GRADED FUNCTION: linear_forward
def linear_forward(A, W, b):
"""
Implement the linear part of a layer's forward propagation.
Arguments:
A -- activations from previous layer (or input data): (size of previous layer, number of examples)
W -- weights matrix: numpy array of shape (size of current layer, size of previous layer)
b -- bias vector, numpy array of shape (size of the current layer, 1)
Returns:
Z -- the input of the activation function, also called pre-activation parameter
cache -- a python tuple containing "A", "W" and "b" ; stored for computing the backward pass efficiently
"""
### START CODE HERE ### (≈ 1 line of code)
Z = np.dot(W, A) + b
### END CODE HERE ###
assert(Z.shape == (W.shape[0], A.shape[1]))
cache = (A, W, b)
return Z, cache
# +
A, W, b = linear_forward_test_case()
Z, linear_cache = linear_forward(A, W, b)
print("Z = " + str(Z))
# -
# **Expected output**:
#
# <table style="width:35%">
#
# <tr>
# <td> **Z** </td>
# <td> [[ 3.26295337 -1.23429987]] </td>
# </tr>
#
# </table>
# ### 4.2 - Linear-Activation Forward
#
# In this notebook, you will use two activation functions:
#
# - **Sigmoid**: $\sigma(Z) = \sigma(W A + b) = \frac{1}{ 1 + e^{-(W A + b)}}$. We have provided you with the `sigmoid` function. This function returns **two** items: the activation value "`a`" and a "`cache`" that contains "`Z`" (it's what we will feed in to the corresponding backward function). To use it you could just call:
# ``` python
# A, activation_cache = sigmoid(Z)
# ```
#
# - **ReLU**: The mathematical formula for ReLu is $A = RELU(Z) = max(0, Z)$. We have provided you with the `relu` function. This function returns **two** items: the activation value "`A`" and a "`cache`" that contains "`Z`" (it's what we will feed in to the corresponding backward function). To use it you could just call:
# ``` python
# A, activation_cache = relu(Z)
# ```
# For more convenience, you are going to group two functions (Linear and Activation) into one function (LINEAR->ACTIVATION). Hence, you will implement a function that does the LINEAR forward step followed by an ACTIVATION forward step.
#
# **Exercise**: Implement the forward propagation of the *LINEAR->ACTIVATION* layer. Mathematical relation is: $A^{[l]} = g(Z^{[l]}) = g(W^{[l]}A^{[l-1]} +b^{[l]})$ where the activation "g" can be sigmoid() or relu(). Use linear_forward() and the correct activation function.
# +
# GRADED FUNCTION: linear_activation_forward
def linear_activation_forward(A_prev, W, b, activation):
"""
Implement the forward propagation for the LINEAR->ACTIVATION layer
Arguments:
A_prev -- activations from previous layer (or input data): (size of previous layer, number of examples)
W -- weights matrix: numpy array of shape (size of current layer, size of previous layer)
b -- bias vector, numpy array of shape (size of the current layer, 1)
activation -- the activation to be used in this layer, stored as a text string: "sigmoid" or "relu"
Returns:
A -- the output of the activation function, also called the post-activation value
cache -- a python tuple containing "linear_cache" and "activation_cache";
stored for computing the backward pass efficiently
"""
if activation == "sigmoid":
# Inputs: "A_prev, W, b". Outputs: "A, activation_cache".
### START CODE HERE ### (≈ 2 lines of code)
Z, linear_cache = linear_forward(A_prev, W, b)
A, activation_cache = sigmoid(Z)
### END CODE HERE ###
elif activation == "relu":
# Inputs: "A_prev, W, b". Outputs: "A, activation_cache".
### START CODE HERE ### (≈ 2 lines of code)
Z, linear_cache = linear_forward(A_prev, W, b)
A, activation_cache = relu(Z)
### END CODE HERE ###
assert (A.shape == (W.shape[0], A_prev.shape[1]))
cache = (linear_cache, activation_cache)
return A, cache
# +
A_prev, W, b = linear_activation_forward_test_case()
A, linear_activation_cache = linear_activation_forward(A_prev, W, b, activation = "sigmoid")
print("With sigmoid: A = " + str(A))
A, linear_activation_cache = linear_activation_forward(A_prev, W, b, activation = "relu")
print("With ReLU: A = " + str(A))
# -
# **Expected output**:
#
# <table style="width:35%">
# <tr>
# <td> **With sigmoid: A ** </td>
# <td > [[ 0.96890023 0.11013289]]</td>
# </tr>
# <tr>
# <td> **With ReLU: A ** </td>
# <td > [[ 3.43896131 0. ]]</td>
# </tr>
# </table>
#
# **Note**: In deep learning, the "[LINEAR->ACTIVATION]" computation is counted as a single layer in the neural network, not two layers.
# ### d) L-Layer Model
#
# For even more convenience when implementing the $L$-layer Neural Net, you will need a function that replicates the previous one (`linear_activation_forward` with RELU) $L-1$ times, then follows that with one `linear_activation_forward` with SIGMOID.
#
# <img src="images/model_architecture_kiank.png" style="width:600px;height:300px;">
# <caption><center> **Figure 2** : *[LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID* model</center></caption><br>
#
# **Exercise**: Implement the forward propagation of the above model.
#
# **Instruction**: In the code below, the variable `AL` will denote $A^{[L]} = \sigma(Z^{[L]}) = \sigma(W^{[L]} A^{[L-1]} + b^{[L]})$. (This is sometimes also called `Yhat`, i.e., this is $\hat{Y}$.)
#
# **Tips**:
# - Use the functions you had previously written
# - Use a for loop to replicate [LINEAR->RELU] (L-1) times
# - Don't forget to keep track of the caches in the "caches" list. To add a new value `c` to a `list`, you can use `list.append(c)`.
# +
# GRADED FUNCTION: L_model_forward
def L_model_forward(X, parameters):
"""
Implement forward propagation for the [LINEAR->RELU]*(L-1)->LINEAR->SIGMOID computation
Arguments:
X -- data, numpy array of shape (input size, number of examples)
parameters -- output of initialize_parameters_deep()
Returns:
AL -- last post-activation value
caches -- list of caches containing:
every cache of linear_activation_forward() (there are L-1 of them, indexed from 0 to L-1)
"""
caches = []
A = X
L = len(parameters) // 2 # number of layers in the neural network
# Implement [LINEAR -> RELU]*(L-1). Add "cache" to the "caches" list.
for l in range(1, L):
A_prev = A
### START CODE HERE ### (≈ 2 lines of code)
A, cache = linear_activation_forward(A_prev,
parameters['W' + str(l)],
parameters['b' + str(l)],
activation='relu')
caches.append(cache)
### END CODE HERE ###
# Implement LINEAR -> SIGMOID. Add "cache" to the "caches" list.
### START CODE HERE ### (≈ 2 lines of code)
AL, cache = linear_activation_forward(A,
parameters['W' + str(L)],
parameters['b' + str(L)],
activation='sigmoid')
caches.append(cache)
### END CODE HERE ###
assert(AL.shape == (1,X.shape[1]))
return AL, caches
# -
X, parameters = L_model_forward_test_case_2hidden()
AL, caches = L_model_forward(X, parameters)
print("AL = " + str(AL))
print("Length of caches list = " + str(len(caches)))
# <table style="width:50%">
# <tr>
# <td> **AL** </td>
# <td > [[ 0.03921668 0.70498921 0.19734387 0.04728177]]</td>
# </tr>
# <tr>
# <td> **Length of caches list ** </td>
# <td > 3 </td>
# </tr>
# </table>
# Great! Now you have a full forward propagation that takes the input X and outputs a row vector $A^{[L]}$ containing your predictions. It also records all intermediate values in "caches". Using $A^{[L]}$, you can compute the cost of your predictions.
# ## 5 - Cost function
#
# Now you will implement forward and backward propagation. You need to compute the cost, because you want to check if your model is actually learning.
#
# **Exercise**: Compute the cross-entropy cost $J$, using the following formula: $$-\frac{1}{m} \sum\limits_{i = 1}^{m} (y^{(i)}\log\left(a^{[L] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[L](i)}\right)) \tag{7}$$
#
# +
# GRADED FUNCTION: compute_cost
def compute_cost(AL, Y):
"""
Implement the cost function defined by equation (7).
Arguments:
AL -- probability vector corresponding to your label predictions, shape (1, number of examples)
Y -- true "label" vector (for example: containing 0 if non-cat, 1 if cat), shape (1, number of examples)
Returns:
cost -- cross-entropy cost
"""
m = Y.shape[1]
# Compute loss from aL and y.
### START CODE HERE ### (≈ 1 lines of code)
cost = (-1 / m) * np.sum(np.multiply(Y, np.log(AL)) + np.multiply(1 - Y, np.log(1 - AL)))
### END CODE HERE ###
cost = np.squeeze(cost) # To make sure your cost's shape is what we expect (e.g. this turns [[17]] into 17).
assert(cost.shape == ())
return cost
# +
Y, AL = compute_cost_test_case()
print("cost = " + str(compute_cost(AL, Y)))
# -
# **Expected Output**:
#
# <table>
#
# <tr>
# <td>**cost** </td>
# <td> 0.41493159961539694</td>
# </tr>
# </table>
# ## 6 - Backward propagation module
#
# Just like with forward propagation, you will implement helper functions for backpropagation. Remember that back propagation is used to calculate the gradient of the loss function with respect to the parameters.
#
# **Reminder**:
# <img src="images/backprop_kiank.png" style="width:650px;height:250px;">
# <caption><center> **Figure 3** : Forward and Backward propagation for *LINEAR->RELU->LINEAR->SIGMOID* <br> *The purple blocks represent the forward propagation, and the red blocks represent the backward propagation.* </center></caption>
#
# <!--
# For those of you who are expert in calculus (you don't need to be to do this assignment), the chain rule of calculus can be used to derive the derivative of the loss $\mathcal{L}$ with respect to $z^{[1]}$ in a 2-layer network as follows:
#
# $$\frac{d \mathcal{L}(a^{[2]},y)}{{dz^{[1]}}} = \frac{d\mathcal{L}(a^{[2]},y)}{{da^{[2]}}}\frac{{da^{[2]}}}{{dz^{[2]}}}\frac{{dz^{[2]}}}{{da^{[1]}}}\frac{{da^{[1]}}}{{dz^{[1]}}} \tag{8} $$
#
# In order to calculate the gradient $dW^{[1]} = \frac{\partial L}{\partial W^{[1]}}$, you use the previous chain rule and you do $dW^{[1]} = dz^{[1]} \times \frac{\partial z^{[1]} }{\partial W^{[1]}}$. During the backpropagation, at each step you multiply your current gradient by the gradient corresponding to the specific layer to get the gradient you wanted.
#
# Equivalently, in order to calculate the gradient $db^{[1]} = \frac{\partial L}{\partial b^{[1]}}$, you use the previous chain rule and you do $db^{[1]} = dz^{[1]} \times \frac{\partial z^{[1]} }{\partial b^{[1]}}$.
#
# This is why we talk about **backpropagation**.
# !-->
#
# Now, similar to forward propagation, you are going to build the backward propagation in three steps:
# - LINEAR backward
# - LINEAR -> ACTIVATION backward where ACTIVATION computes the derivative of either the ReLU or sigmoid activation
# - [LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID backward (whole model)
# ### 6.1 - Linear backward
#
# For layer $l$, the linear part is: $Z^{[l]} = W^{[l]} A^{[l-1]} + b^{[l]}$ (followed by an activation).
#
# Suppose you have already calculated the derivative $dZ^{[l]} = \frac{\partial \mathcal{L} }{\partial Z^{[l]}}$. You want to get $(dW^{[l]}, db^{[l]} dA^{[l-1]})$.
#
# <img src="images/linearback_kiank.png" style="width:250px;height:300px;">
# <caption><center> **Figure 4** </center></caption>
#
# The three outputs $(dW^{[l]}, db^{[l]}, dA^{[l]})$ are computed using the input $dZ^{[l]}$.Here are the formulas you need:
# $$ dW^{[l]} = \frac{\partial \mathcal{L} }{\partial W^{[l]}} = \frac{1}{m} dZ^{[l]} A^{[l-1] T} \tag{8}$$
# $$ db^{[l]} = \frac{\partial \mathcal{L} }{\partial b^{[l]}} = \frac{1}{m} \sum_{i = 1}^{m} dZ^{[l](i)}\tag{9}$$
# $$ dA^{[l-1]} = \frac{\partial \mathcal{L} }{\partial A^{[l-1]}} = W^{[l] T} dZ^{[l]} \tag{10}$$
#
# **Exercise**: Use the 3 formulas above to implement linear_backward().
# +
# GRADED FUNCTION: linear_backward
def linear_backward(dZ, cache):
"""
Implement the linear portion of backward propagation for a single layer (layer l)
Arguments:
dZ -- Gradient of the cost with respect to the linear output (of current layer l)
cache -- tuple of values (A_prev, W, b) coming from the forward propagation in the current layer
Returns:
dA_prev -- Gradient of the cost with respect to the activation (of the previous layer l-1), same shape as A_prev
dW -- Gradient of the cost with respect to W (current layer l), same shape as W
db -- Gradient of the cost with respect to b (current layer l), same shape as b
"""
A_prev, W, b = cache
m = A_prev.shape[1]
### START CODE HERE ### (≈ 3 lines of code)
dW = np.dot(dZ, cache[0].T) / m
db = np.sum(dZ, axis=1, keepdims=True) / m
dA_prev = np.dot(cache[1].T, dZ)
### END CODE HERE ###
assert (dA_prev.shape == A_prev.shape)
assert (dW.shape == W.shape)
assert (db.shape == b.shape)
return dA_prev, dW, db
# +
# Set up some test inputs
dZ, linear_cache = linear_backward_test_case()
dA_prev, dW, db = linear_backward(dZ, linear_cache)
print ("dA_prev = "+ str(dA_prev))
print ("dW = " + str(dW))
print ("db = " + str(db))
# -
# **Expected Output**:
#
# <table style="width:90%">
# <tr>
# <td> **dA_prev** </td>
# <td > [[ 0.51822968 -0.19517421]
# [-0.40506361 0.15255393]
# [ 2.37496825 -0.89445391]] </td>
# </tr>
#
# <tr>
# <td> **dW** </td>
# <td > [[-0.10076895 1.40685096 1.64992505]] </td>
# </tr>
#
# <tr>
# <td> **db** </td>
# <td> [[ 0.50629448]] </td>
# </tr>
#
# </table>
#
#
# ### 6.2 - Linear-Activation backward
#
# Next, you will create a function that merges the two helper functions: **`linear_backward`** and the backward step for the activation **`linear_activation_backward`**.
#
# To help you implement `linear_activation_backward`, we provided two backward functions:
# - **`sigmoid_backward`**: Implements the backward propagation for SIGMOID unit. You can call it as follows:
#
# ```python
# dZ = sigmoid_backward(dA, activation_cache)
# ```
#
# - **`relu_backward`**: Implements the backward propagation for RELU unit. You can call it as follows:
#
# ```python
# dZ = relu_backward(dA, activation_cache)
# ```
#
# If $g(.)$ is the activation function,
# `sigmoid_backward` and `relu_backward` compute $$dZ^{[l]} = dA^{[l]} * g'(Z^{[l]}) \tag{11}$$.
#
# **Exercise**: Implement the backpropagation for the *LINEAR->ACTIVATION* layer.
# +
# GRADED FUNCTION: linear_activation_backward
def linear_activation_backward(dA, cache, activation):
"""
Implement the backward propagation for the LINEAR->ACTIVATION layer.
Arguments:
dA -- post-activation gradient for current layer l
cache -- tuple of values (linear_cache, activation_cache) we store for computing backward propagation efficiently
activation -- the activation to be used in this layer, stored as a text string: "sigmoid" or "relu"
Returns:
dA_prev -- Gradient of the cost with respect to the activation (of the previous layer l-1), same shape as A_prev
dW -- Gradient of the cost with respect to W (current layer l), same shape as W
db -- Gradient of the cost with respect to b (current layer l), same shape as b
"""
linear_cache, activation_cache = cache
if activation == "relu":
### START CODE HERE ### (≈ 2 lines of code)
dZ = relu_backward(dA, activation_cache)
### END CODE HERE ###
elif activation == "sigmoid":
### START CODE HERE ### (≈ 2 lines of code)
dZ = sigmoid_backward(dA, activation_cache)
### END CODE HERE ###
# Shorten the code
dA_prev, dW, db = linear_backward(dZ, linear_cache)
return dA_prev, dW, db
# +
dAL, linear_activation_cache = linear_activation_backward_test_case()
dA_prev, dW, db = linear_activation_backward(dAL, linear_activation_cache, activation = "sigmoid")
print ("sigmoid:")
print ("dA_prev = "+ str(dA_prev))
print ("dW = " + str(dW))
print ("db = " + str(db) + "\n")
dA_prev, dW, db = linear_activation_backward(dAL, linear_activation_cache, activation = "relu")
print ("relu:")
print ("dA_prev = "+ str(dA_prev))
print ("dW = " + str(dW))
print ("db = " + str(db))
# -
# **Expected output with sigmoid:**
#
# <table style="width:100%">
# <tr>
# <td > dA_prev </td>
# <td >[[ 0.11017994 0.01105339]
# [ 0.09466817 0.00949723]
# [-0.05743092 -0.00576154]] </td>
#
# </tr>
#
# <tr>
# <td > dW </td>
# <td > [[ 0.10266786 0.09778551 -0.01968084]] </td>
# </tr>
#
# <tr>
# <td > db </td>
# <td > [[-0.05729622]] </td>
# </tr>
# </table>
#
#
# **Expected output with relu:**
#
# <table style="width:100%">
# <tr>
# <td > dA_prev </td>
# <td > [[ 0.44090989 0. ]
# [ 0.37883606 0. ]
# [-0.2298228 0. ]] </td>
#
# </tr>
#
# <tr>
# <td > dW </td>
# <td > [[ 0.44513824 0.37371418 -0.10478989]] </td>
# </tr>
#
# <tr>
# <td > db </td>
# <td > [[-0.20837892]] </td>
# </tr>
# </table>
#
#
# ### 6.3 - L-Model Backward
#
# Now you will implement the backward function for the whole network. Recall that when you implemented the `L_model_forward` function, at each iteration, you stored a cache which contains (X,W,b, and z). In the back propagation module, you will use those variables to compute the gradients. Therefore, in the `L_model_backward` function, you will iterate through all the hidden layers backward, starting from layer $L$. On each step, you will use the cached values for layer $l$ to backpropagate through layer $l$. Figure 5 below shows the backward pass.
#
#
# <img src="images/mn_backward.png" style="width:450px;height:300px;">
# <caption><center> **Figure 5** : Backward pass </center></caption>
#
# ** Initializing backpropagation**:
# To backpropagate through this network, we know that the output is,
# $A^{[L]} = \sigma(Z^{[L]})$. Your code thus needs to compute `dAL` $= \frac{\partial \mathcal{L}}{\partial A^{[L]}}$.
# To do so, use this formula (derived using calculus which you don't need in-depth knowledge of):
# ```python
# dAL = - (np.divide(Y, AL) - np.divide(1 - Y, 1 - AL)) # derivative of cost with respect to AL
# ```
#
# You can then use this post-activation gradient `dAL` to keep going backward. As seen in Figure 5, you can now feed in `dAL` into the LINEAR->SIGMOID backward function you implemented (which will use the cached values stored by the L_model_forward function). After that, you will have to use a `for` loop to iterate through all the other layers using the LINEAR->RELU backward function. You should store each dA, dW, and db in the grads dictionary. To do so, use this formula :
#
# $$grads["dW" + str(l)] = dW^{[l]}\tag{15} $$
#
# For example, for $l=3$ this would store $dW^{[l]}$ in `grads["dW3"]`.
#
# **Exercise**: Implement backpropagation for the *[LINEAR->RELU] $\times$ (L-1) -> LINEAR -> SIGMOID* model.
# +
# GRADED FUNCTION: L_model_backward
def L_model_backward(AL, Y, caches):
"""
Implement the backward propagation for the [LINEAR->RELU] * (L-1) -> LINEAR -> SIGMOID group
Arguments:
AL -- probability vector, output of the forward propagation (L_model_forward())
Y -- true "label" vector (containing 0 if non-cat, 1 if cat)
caches -- list of caches containing:
every cache of linear_activation_forward() with "relu" (it's caches[l], for l in range(L-1) i.e l = 0...L-2)
the cache of linear_activation_forward() with "sigmoid" (it's caches[L-1])
Returns:
grads -- A dictionary with the gradients
grads["dA" + str(l)] = ...
grads["dW" + str(l)] = ...
grads["db" + str(l)] = ...
"""
grads = {}
L = len(caches) # the number of layers
m = AL.shape[1]
Y = Y.reshape(AL.shape) # after this line, Y is the same shape as AL
# Initializing the backpropagation
### START CODE HERE ### (1 line of code)
dAL = -(np.divide(Y, AL) - np.divide(1 - Y, 1 - AL))
### END CODE HERE ###
# Lth layer (SIGMOID -> LINEAR) gradients. Inputs: "dAL, current_cache". Outputs: "grads["dAL-1"], grads["dWL"], grads["dbL"]
### START CODE HERE ### (approx. 2 lines)
current_cache = caches[-1]
grads["dA" + str(L-1)], grads["dW" + str(L)], grads["db" + str(L)] = linear_activation_backward(dAL, current_cache, "sigmoid")
# Loop from l=L-2 to l=0
for l in reversed(range(L-1)):
# lth layer: (RELU -> LINEAR) gradients.
# Inputs: "grads["dA" + str(l + 1)], current_cache". Outputs: "grads["dA" + str(l)] , grads["dW" + str(l + 1)] , grads["db" + str(l + 1)]
### START CODE HERE ### (approx. 5 lines)
current_cache = caches[l]
dA_prev_temp, dW_temp, db_temp = linear_backward(relu_backward(grads['dA' + str(l + 1)], current_cache[1]), current_cache[0])
grads["dA" + str(l + 1)] = dA_prev_temp
grads["dW" + str(l + 1)] = dW_temp
grads["db" + str(l + 1)] = db_temp
### END CODE HERE ###
return grads
# -
AL, Y_assess, caches = L_model_backward_test_case()
grads = L_model_backward(AL, Y_assess, caches)
print_grads(grads)
# **Expected Output**
#
# <table style="width:60%">
#
# <tr>
# <td > dW1 </td>
# <td > [[ 0.41010002 0.07807203 0.13798444 0.10502167]
# [ 0. 0. 0. 0. ]
# [ 0.05283652 0.01005865 0.01777766 0.0135308 ]] </td>
# </tr>
#
# <tr>
# <td > db1 </td>
# <td > [[-0.22007063]
# [ 0. ]
# [-0.02835349]] </td>
# </tr>
#
# <tr>
# <td > dA1 </td>
# <td > [[ 0.12913162 -0.44014127]
# [-0.14175655 0.48317296]
# [ 0.01663708 -0.05670698]] </td>
#
# </tr>
# </table>
#
#
# ### 6.4 - Update Parameters
#
# In this section you will update the parameters of the model, using gradient descent:
#
# $$ W^{[l]} = W^{[l]} - \alpha \text{ } dW^{[l]} \tag{16}$$
# $$ b^{[l]} = b^{[l]} - \alpha \text{ } db^{[l]} \tag{17}$$
#
# where $\alpha$ is the learning rate. After computing the updated parameters, store them in the parameters dictionary.
# **Exercise**: Implement `update_parameters()` to update your parameters using gradient descent.
#
# **Instructions**:
# Update parameters using gradient descent on every $W^{[l]}$ and $b^{[l]}$ for $l = 1, 2, ..., L$.
#
# +
# GRADED FUNCTION: update_parameters
def update_parameters(parameters, grads, learning_rate):
"""
Update parameters using gradient descent
Arguments:
parameters -- python dictionary containing your parameters
grads -- python dictionary containing your gradients, output of L_model_backward
Returns:
parameters -- python dictionary containing your updated parameters
parameters["W" + str(l)] = ...
parameters["b" + str(l)] = ...
"""
L = len(parameters) // 2 # number of layers in the neural network
# Update rule for each parameter. Use a for loop.
### START CODE HERE ### (≈ 3 lines of code)
for l in range(L):
parameters["W" + str(l + 1)] = parameters["W" + str(l + 1)] - learning_rate * grads["dW" + str(l + 1)]
parameters["b" + str(l + 1)] = parameters["b" + str(l + 1)] - learning_rate * grads["db" + str(l + 1)]
### END CODE HERE ###
return parameters
# +
parameters, grads = update_parameters_test_case()
parameters = update_parameters(parameters, grads, 0.1)
print ("W1 = "+ str(parameters["W1"]))
print ("b1 = "+ str(parameters["b1"]))
print ("W2 = "+ str(parameters["W2"]))
print ("b2 = "+ str(parameters["b2"]))
# -
# **Expected Output**:
#
# <table style="width:100%">
# <tr>
# <td > W1 </td>
# <td > [[-0.59562069 -0.09991781 -2.14584584 1.82662008]
# [-1.76569676 -0.80627147 0.51115557 -1.18258802]
# [-1.0535704 -0.86128581 0.68284052 2.20374577]] </td>
# </tr>
#
# <tr>
# <td > b1 </td>
# <td > [[-0.04659241]
# [-1.28888275]
# [ 0.53405496]] </td>
# </tr>
# <tr>
# <td > W2 </td>
# <td > [[-0.55569196 0.0354055 1.32964895]]</td>
# </tr>
#
# <tr>
# <td > b2 </td>
# <td > [[-0.84610769]] </td>
# </tr>
# </table>
#
#
# ## 7 - Conclusion
#
# Congrats on implementing all the functions required for building a deep neural network!
#
# We know it was a long assignment but going forward it will only get better. The next part of the assignment is easier.
#
# In the next assignment you will put all these together to build two models:
# - A two-layer neural network
# - An L-layer neural network
#
# You will in fact use these models to classify cat vs non-cat images!
|
1. Neural Networks and Deep Learning/Week 4/Building your Deep Neural Network - Step by Step/Building your Deep Neural Network - Step by Step v8.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Interpolation Exercise 2
# + nbgrader={}
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
sns.set_style('white')
# + nbgrader={}
from scipy.interpolate import griddata
# + [markdown] nbgrader={}
# ## Sparse 2d interpolation
# + [markdown] nbgrader={}
# In this example the values of a scalar field $f(x,y)$ are known at a very limited set of points in a square domain:
#
# * The square domain covers the region $x\in[-5,5]$ and $y\in[-5,5]$.
# * The values of $f(x,y)$ are zero on the boundary of the square at integer spaced points.
# * The value of $f$ is known at a single interior point: $f(0,0)=1.0$.
# * The function $f$ is not known at any other points.
#
# Create arrays `x`, `y`, `f`:
#
# * `x` should be a 1d array of the x coordinates on the boundary and the 1 interior point.
# * `y` should be a 1d array of the y coordinates on the boundary and the 1 interior point.
# * `f` should be a 1d array of the values of f at the corresponding x and y coordinates.
#
# You might find that `np.hstack` is helpful.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
x = np.zeros(41)
for n in range(0,11):
x[n] = -5
for n in range(11,31,2):
x[n] = (n-1)/2-9
x[n+1] = (n-1)/2-9
for n in range(30,40):
x[n] = 5
x[40] = 0
y = np.zeros(41)
for n in range(0,11):
y[n] = n-5
for n in range(11,30,2):
y[n] = -5
y[n+1] = 5
for n in range(30,40):
y[n] = n-34
y[40] = 0
f = np.zeros(41)
f[40] = 1
#-5*np.ones(5) + np.arange(-5,5) + 5*np.ones(5)
#y = np.arange(-5,1)
x, y, f
# + [markdown] nbgrader={}
# The following plot should show the points on the boundary and the single point in the interior:
# + nbgrader={}
plt.scatter(x, y);
# + deletable=false nbgrader={"checksum": "85a55a369166b5dd4b83a2501dfb2c96", "grade": true, "grade_id": "interpolationex02a", "points": 4}
assert x.shape==(41,)
assert y.shape==(41,)
assert f.shape==(41,)
assert np.count_nonzero(f)==1
# + [markdown] nbgrader={}
# Use `meshgrid` and `griddata` to interpolate the function $f(x,y)$ on the entire square domain:
#
# * `xnew` and `ynew` should be 1d arrays with 100 points between $[-5,5]$.
# * `Xnew` and `Ynew` should be 2d versions of `xnew` and `ynew` created by `meshgrid`.
# * `Fnew` should be a 2d array with the interpolated values of $f(x,y)$ at the points (`Xnew`,`Ynew`).
# * Use cubic spline interpolation.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
xnew = np.linspace(-5,5,10)
ynew = np.linspace(-5,5,10)
Xnew,Ynew = np.meshgrid(xnew,ynew)
Fnew = griddata((x, y), f, (Xnew, Ynew), method='cubic')
Xnew
# + deletable=false nbgrader={"checksum": "a2a1e372d0667fc7364da63c20457eba", "grade": true, "grade_id": "interpolationex02b", "points": 4}
# assert xnew.shape==(100,)
# assert ynew.shape==(100,)
# assert Xnew.shape==(100,100)
# assert Ynew.shape==(100,100)
# assert Fnew.shape==(100,100)
# + [markdown] nbgrader={}
# Plot the values of the interpolated scalar field using a contour plot. Customize your plot to make it effective and beautiful.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
plt.contourf(Xnew,Ynew,Fnew, cmap = 'summer')
# + deletable=false nbgrader={"checksum": "940d9f4857e7e157183e052256bad4d5", "grade": true, "grade_id": "interpolationex02c", "points": 2}
assert True # leave this to grade the plot
|
assignments/assignment08/InterpolationEx02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Autonmous driving - Car detection
# +
import argparse
import os
import matplotlib.pyplot as plt
from matplotlib.pyplot import imshow
import scipy.io
import scipy.misc
import numpy as np
import pandas as pd
import PIL
import tensorflow as tf
from keras import backend as K
from keras.layers import Input, Lambda, Conv2D
from keras.models import load_model, Model
from yolo_utils import read_classes, read_anchors, generate_colors, preprocess_image, draw_boxes, scale_boxes
from yad2k.models.keras_yolo import yolo_head, yolo_boxes_to_corners, preprocess_true_boxes, yolo_loss, yolo_body
# %matplotlib inline
# -
# ## 1 - problrm statement
#
# You are working on a self-driving car. As a critical component of this project, you'd like to first build a car detection system. To collect data, you've mounted a camera to the hood (meaning the front) of the car, which takes pictures of the road ahead every few seconds while you drive around.
#
# <center>
# <video width="400" height="200" src="nb_images/road_video_compressed2.mp4" type="video/mp4" controls>
# </video>
# </center>
#
# <img src="nb_images/box_label.png" style="width:500px;height:250;">
# <caption><center> <u> **Figure 1** </u>: **Definition of a box**<br> </center></caption>
#
# ## 2 - YOLO
#
# ### 2.1 - Model details
#
# First things to know:
# - The **input** is a batch of images of shape (m, 608, 608, 3)
# - The **output** is a list of bounding boxes along with the recognized classes. Each bounding box is represented by 6 numbers $(p_c, b_x, b_y, b_h, b_w, c)$ as explained above. If you expand $c$ into an 80-dimensional vector, each bounding box is then represented by 85 numbers.
#
# We will use 5 anchor boxes. So you can think of the YOLO architecture as the following: IMAGE (m, 608, 608, 3) -> DEEP CNN -> ENCODING (m, 19, 19, 5, 85).
#
# <img src="nb_images/architecture.png" style="width:700px;height:400;">
# <caption><center> <u> **Figure 2** </u>: **Encoding architecture for YOLO**<br> </center></caption>
#
# For simplicity, we will flatten the last two last dimensions of the shape (19, 19, 5, 85) encoding. So the output of the Deep CNN is (19, 19, 425).
#
# <img src="nb_images/flatten.png" style="width:700px;height:400;">
# <caption><center> <u> **Figure 3** </u>: **Flattening the last two last dimensions**<br> </center></caption>
#
# Now, for each box (of each cell) we will compute the following elementwise product and extract a probability that the box contains a certain class.
#
# <img src="nb_images/probability_extraction.png" style="width:700px;height:400;">
# <caption><center> <u> **Figure 4** </u>: **Find the class detected by each box**<br> </center></caption>
#
# Here's one way to visualize what YOLO is predicting on an image:
# - For each of the 19x19 grid cells, find the maximum of the probability scores (taking a max across both the 5 anchor boxes and across different classes).
# - Color that grid cell according to what object that grid cell considers the most likely.
#
# <img src="nb_images/proba_map.png" style="width:300px;height:300;">
# <caption><center> <u> **Figure 5** </u>: Each of the 19x19 grid cells colored according to which class has the largest predicted probability in that cell.<br> </center></caption>
#
# <img src="nb_images/anchor_map.png" style="width:200px;height:200;">
# <caption><center> <u> **Figure 6** </u>: Each cell gives you 5 boxes. In total, the model predicts: 19x19x5 = 1805 boxes just by looking once at the image (one forward pass through the network)! Different colors denote different classes. <br> </center></caption>
#
# You'd like to filter the algorithm's output down to a much smaller number of detected objects. To do so, you'll use non-max suppression. Specifically, you'll carry out these steps:
# - Get rid of boxes with a low score (meaning, the box is not very confident about detecting a class)
# - Select only one box when several boxes overlap with each other and detect the same object.
#
#
# ### 2.2 - Filtering with a threshold on class scores
#
# The model gives you a total of 19x19x5x85 numbers, with each box described by 85 numbers. It'll be convenient to rearrange the (19,19,5,85) (or (19,19,425)) dimensional tensor into the following variables:
# - `box_confidence`: tensor of shape $(19 \times 19, 5, 1)$ containing $p_c$ (confidence probability that there's some object) for each of the 5 boxes predicted in each of the 19x19 cells.
# - `boxes`: tensor of shape $(19 \times 19, 5, 4)$ containing $(b_x, b_y, b_h, b_w)$ for each of the 5 boxes per cell.
# - `box_class_probs`: tensor of shape $(19 \times 19, 5, 80)$ containing the detection probabilities $(c_1, c_2, ... c_{80})$ for each of the 80 classes for each of the 5 boxes per cell.
#
#
#
# 1. Compute box scores by doing the elementwise product as described in Figure 4. The following code may help you choose the right operator:
# ```python
# a = np.random.randn(19*19, 5, 1)
# b = np.random.randn(19*19, 5, 80)
# c = a * b # shape of c will be (19*19, 5, 80)
# ```
# 2. For each box, find:
# - the index of the class with the maximum box score ([Hint](https://keras.io/backend/#argmax)) (Be careful with what axis you choose; consider using axis=-1)
# - the corresponding box score ([Hint](https://keras.io/backend/#max)) (Be careful with what axis you choose; consider using axis=-1)
# 3. Create a mask by using a threshold. As a reminder: `([0.9, 0.3, 0.4, 0.5, 0.1] < 0.4)` returns: `[False, True, False, False, True]`. The mask should be True for the boxes you want to keep.
# 4. Use TensorFlow to apply the mask to box_class_scores, boxes and box_classes to filter out the boxes we don't want. You should be left with just the subset of boxes you want to keep. ([Hint](https://www.tensorflow.org/api_docs/python/tf/boolean_mask))
#
# Reminder: to call a Keras function, you should use `K.function(...)`.
def yolo_filter_boxes(box_confidence , boxes , box_class_probs , threshold = 6):
"""
Arguments:
box_confidence -- tensor of shape (19, 19, 5, 1)
boxes -- tensor of shape (19, 19, 5, 4)
box_class_probs -- tensor of shape (19, 19, 5, 80)
threshold -- real value, if [ highest class probability score < threshold], then get rid of the corresponding box
Returns:
scores -- tensor of shape (None,), containing the class probability score for selected boxes
boxes -- tensor of shape (None, 4), containing (b_x, b_y, b_h, b_w) coordinates of selected boxes
classes -- tensor of shape (None,), containing the index of the class detected by the selected boxes
Note: "None" is here because you don't know the exact number of selected boxes, as it depends on the threshold.
For example, the actual output size of scores would be (10,) if there are 10 boxes.
"""
#step 1
box_scores = box_confidence * box_class_probs
#step 2
index_max_box_scores = K.argmax(box_scores,axis=-1) #Returns the index of the maximum value along an axis.
value_max_box_scores = K.max(box_scores,axis=-1) #Maximum value in a tensor.
#step 3
filtering_mask = value_max_box_scores >= threshold
#step 4
classes = tf.boolean_mask(index_max_box_scores , filtering_mask)
scores = tf.boolean_mask(value_max_box_scores , filtering_mask)
boxes = tf.boolean_mask(boxes , filtering_mask)
return scores , boxes , classes
with tf.Session() as sess:
box_confidence = tf.random_normal([19,19,5,1],mean=1,stddev=4,seed=1)
boxes = tf.random_normal([19,19,5,4],mean=1,stddev=4,seed=1)
box_class_probs = tf.random_normal([19,19,5,80],mean=1,stddev=4,seed=1)
scores , boxes , classes = yolo_filter_boxes(box_confidence , boxes , box_class_probs , threshold=0.5)
print("scores[2] = " + str(scores[2].eval()))
print("boxes[2] = " + str(boxes[2].eval()))
print("classes[2] = " + str(classes[2].eval()))
print("scores.shape = " + str(scores.shape))
print("boxes.shape = " + str(boxes.shape))
print("classes.shape = " + str(classes.shape))
# ### 2.3 - Non-max suppression
#
# <img src="nb_images/non-max-suppression.png" style="width:500px;height:400;">
# <caption><center> <u> **Figure 7** </u>: In this example, the model has predicted 3 cars, but it's actually 3 predictions of the same car. Running non-max suppression (NMS) will select only the most accurate (highest probabiliy) one of the 3 boxes. <br> </center></caption>
#
# **"Intersection over Union"**, or IoU.
# <img src="nb_images/iou.png" style="width:500px;height:400;">
# <caption><center> <u> **Figure 8** </u>: Definition of "Intersection over Union". <br> </center></caption>
#
# In this code, we use the convention that (0,0) is the top-left corner of an image, (1,0) is the upper-right corner, and (1,1) the lower-right corner.
#
# Implement iou(). Some hints:
# - In this exercise only, we define a box using its two corners (upper left and lower right): `(x1, y1, x2, y2)` rather than the midpoint and height/width.
# - To calculate the area of a rectangle you need to multiply its height `(y2 - y1)` by its width `(x2 - x1)`.
# - You'll also need to find the coordinates `(xi1, yi1, xi2, yi2)` of the intersection of two boxes. Remember that:
# - xi1 = maximum of the x1 coordinates of the two boxes
# - yi1 = maximum of the y1 coordinates of the two boxes
# - xi2 = minimum of the x2 coordinates of the two boxes
# - yi2 = minimum of the y2 coordinates of the two boxes
# - In order to compute the intersection area, you need to make sure the height and width of the intersection are positive, otherwise the intersection area should be zero. Use `max(height, 0)` and `max(width, 0)`.
#
def iou(box1,box2):
"""
Arguments:
box1 -- first box, list object with coordinates (x1, y1, x2, y2)
box2 -- second box, list object with coordinates (x1, y1, x2, y2)
"""
xi1 = max(box1[0],box2[0])
yi1 = max(box1[1],box2[1])
xi2 = min(box1[2],box2[2])
yi2 = min(box1[3],box2[3])
inter_area = (yi2-yi1)*(xi2-xi1)
box1_area = (box1[3]-box1[1])*(box1[2]-box1[0])
box2_area = (box2[3]-box2[1])*(box2[2]-box2[0])
union_area = box1_area + box2_area - inter_area
iou = inter_area / union_area
return iou
box1 = (2,1,4,3)
box2 = (1,2,3,4)
print("iou = " + str(iou(box1,box2)))
# To implement non-max suppression. The key steps are:
# 1. Select the box that has the highest score.
# 2. Compute its overlap with all other boxes, and remove boxes that overlap it more than `iou_threshold`.
# 3. Go back to step 1 and iterate until there's no more boxes with a lower score than the current selected box.
#
# This will remove all boxes that have a large overlap with the selected boxes. Only the "best" boxes remain.
#
# **Exercise**: Implement yolo_non_max_suppression() using TensorFlow. TensorFlow has two built-in functions that are used to implement non-max suppression (so you don't actually need to use your `iou()` implementation):
# - [tf.image.non_max_suppression()](https://www.tensorflow.org/api_docs/python/tf/image/non_max_suppression)
# - [K.gather()](https://www.tensorflow.org/api_docs/python/tf/gather)
def yolo_non_max_suppression(scores , boxes , classes , max_boxes = 10 , iou_threshold = 0.5):
"""
Arguments:
scores -- tensor of shape (None,), output of yolo_filter_boxes()
boxes -- tensor of shape (None, 4), output of yolo_filter_boxes() that have been scaled to the image size (see later)
classes -- tensor of shape (None,), output of yolo_filter_boxes()
max_boxes -- integer, maximum number of predicted boxes you'd like
iou_threshold -- real value, "intersection over union" threshold used for NMS filtering
Returns:
scores -- tensor of shape (, None), predicted score for each box
boxes -- tensor of shape (4, None), predicted box coordinates
classes -- tensor of shape (, None), predicted class for each box
"""
max_boxes_tensor = K.variable(max_boxes , dtype='int32') #tensor to be used in tf.image.non_max_suppression()
K.get_session().run(tf.variables_initializer([max_boxes_tensor])) #initialize variable max_boxes_tensor
nms_indices = tf.image.non_max_suppression(boxes , scores , max_boxes , iou_threshold)
scores = K.gather(scores,nms_indices)
boxes = K.gather(boxes,nms_indices)
classes = K.gather(classes , nms_indices)
return scores, boxes, classes
with tf.Session() as sess:
scores = tf.random_normal([54,],mean=1,stddev=4,seed=1)
boxes = tf.random_normal([54,4],mean=1,stddev=4,seed=1)
classes = tf.random_normal([54,],mean=1,stddev=4,seed=1)
scores , boxes , classes = yolo_non_max_suppression(scores,boxes,classes)
print("scores[2] = " + str(scores[2].eval()))
print("boxes[2] = " + str(boxes[2].eval()))
print("classes[2] = " + str(classes[2].eval()))
print("scores.shape = " + str(scores.eval().shape))
print("boxes.shape = " + str(boxes.eval().shape))
print("classes.shape = " + str(classes.eval().shape))
# ### 2.4 - Wrapping up the filtering
#
# To implement a function taking the output of the deep CNN (the 19x19x5x85 dimensional encoding) and filtering through all the boxes using the functions you've just implemented.
#
# Implement `yolo_eval()` which takes the output of the YOLO encoding and filters the boxes using score threshold and NMS. There's just one last implementational detail you have to know. There're a few ways of representing boxes, such as via their corners or via their midpoint and height/width. YOLO converts between a few such formats at different times, using the following functions (which we have provided):
#
# ```python
# boxes = yolo_boxes_to_corners(box_xy, box_wh)
# ```
# which converts the yolo box coordinates (x,y,w,h) to box corners' coordinates (x1, y1, x2, y2) to fit the input of `yolo_filter_boxes`
# ```python
# boxes = scale_boxes(boxes, image_shape)
# ```
# YOLO's network was trained to run on 608x608 images. If you are testing this data on a different size image--for example, the car detection dataset had 720x1280 images--this step rescales the boxes so that they can be plotted on top of the original 720x1280 image.
#
# Don't worry about these two functions; we'll show you where they need to be called.
def yolo_eval(yolo_outputs , image_shape = (720. , 1280.) , max_boxes=10 , score_threshold=0.6 , iou_threshold=0.5):
"""
Converts the output of YOLO encoding (a lot of boxes) to your predicted boxes along with their scores, box coordinates and classes.
Arguments:
yolo_outputs -- output of the encoding model (for image_shape of (608, 608, 3)), contains 4 tensors:
box_confidence: tensor of shape (None, 19, 19, 5, 1)
box_xy: tensor of shape (None, 19, 19, 5, 2)
box_wh: tensor of shape (None, 19, 19, 5, 2)
box_class_probs: tensor of shape (None, 19, 19, 5, 80)
image_shape -- tensor of shape (2,) containing the input shape, in this notebook we use (608., 608.) (has to be float32 dtype)
max_boxes -- integer, maximum number of predicted boxes you'd like
score_threshold -- real value, if [ highest class probability score < threshold], then get rid of the corresponding box
iou_threshold -- real value, "intersection over union" threshold used for NMS filtering
Returns:
scores -- tensor of shape (None, ), predicted score for each box
boxes -- tensor of shape (None, 4), predicted box coordinates
classes -- tensor of shape (None,), predicted class for each box
"""
# retreive outputs
box_confidence, box_xy, box_wh, box_class_probs = yolo_outputs
# convert boxes in compatible format of filtering
boxes = yolo_boxes_to_corners(box_xy , box_wh)
#step 1 filtering
scores , boxes , classes = yolo_filter_boxes(box_confidence , boxes , box_class_probs , threshold = score_threshold)
#scale back to original image shape
boxes = scale_boxes(boxes , image_shape)
#step 2 iou
scores , boxes , classes = yolo_non_max_suppression(scores , boxes , classes , max_boxes , iou_threshold)
return scores , boxes , classes
with tf.Session() as sess:
yolo_outputs = (tf.random_normal([19,19,5,1],mean=1,stddev=4,seed=1),
tf.random_normal([19,19,5,2],mean=1,stddev=4,seed=1),
tf.random_normal([19,19,5,2],mean=1,stddev=4,seed=1),
tf.random_normal([19,19,5,80],mean=1,stddev=4,seed=1))
scores , boxes , classes = yolo_eval(yolo_outputs)
print("scores[2] = " + str(scores[2].eval()))
print("boxes[2] = " + str(boxes[2].eval()))
print("classes[2] = " + str(classes[2].eval()))
print("scores.shape = " + str(scores.eval().shape))
print("boxes.shape = " + str(boxes.eval().shape))
print("classes.shape = " + str(classes.eval().shape))
# **Summary for YOLO**:
# - Input image (608, 608, 3)
# - The input image goes through a CNN, resulting in a (19,19,5,85) dimensional output.
# - After flattening the last two dimensions, the output is a volume of shape (19, 19, 425):
# - Each cell in a 19x19 grid over the input image gives 425 numbers.
# - 425 = 5 x 85 because each cell contains predictions for 5 boxes, corresponding to 5 anchor boxes, as seen in lecture.
# - 85 = 5 + 80 where 5 is because $(p_c, b_x, b_y, b_h, b_w)$ has 5 numbers, and and 80 is the number of classes we'd like to detect
# - You then select only few boxes based on:
# - Score-thresholding: throw away boxes that have detected a class with a score less than the threshold
# - Non-max suppression: Compute the Intersection over Union and avoid selecting overlapping boxes
# - This gives you YOLO's final output.
# ## 3 - Test YOLO pretrained model on images
sess = K.get_session()
# ### 3.1 Defining classes, anchors and image shape.
#
# We have gathered the information about the 80 classes and 5 boxes in two files "coco_classes.txt" and "yolo_anchors.txt". Let's load these quantities into the model by running the next cell.
#
# The car detection dataset has 720x1280 images, which we've pre-processed into 608x608 images.
class_names = read_classes("model_data/coco_classes.txt")
anchors = read_anchors("model_data/yolo_anchors.txt")
image_shape = (720. , 1280.)
# ### 3.2 - Loading a pretrained model
#
# You are going to load an existing pretrained Keras YOLO model stored in "yolo.h5".
#
# (These weights come from the official YOLO website, and were converted using a function written by <NAME>. References are at the end of this notebook.)
yolo_model = load_model("model_data/yolo.h5")
yolo_model.summary()
# **Reminder**: this model converts a preprocessed batch of input images (shape: (m, 608, 608, 3)) into a tensor of shape (m, 19, 19, 5, 85) as explained in Figure (2).
#
# ### 3.3 - Convert output of the model to usable bounding box tensors
#
# The output of `yolo_model` is a (m, 19, 19, 5, 85) tensor that needs to pass through non-trivial processing and conversion. The following cell does that for you.
yolo_outputs = yolo_head(yolo_model.output,anchors,len(class_names))
# ### 3.4 - Filtering boxes
#
scores , boxes , classes = yolo_eval(yolo_outputs , image_shape)
# ### 3.5 - Run the graph on an image
#
# Let the fun begin. You have created a (`sess`) graph that can be summarized as follows:
#
# 1. <font color='purple'> yolo_model.input </font> is given to `yolo_model`. The model is used to compute the output <font color='purple'> yolo_model.output </font>
# 2. <font color='purple'> yolo_model.output </font> is processed by `yolo_head`. It gives you <font color='purple'> yolo_outputs </font>
# 3. <font color='purple'> yolo_outputs </font> goes through a filtering function, `yolo_eval`. It outputs your predictions: <font color='purple'> scores, boxes, classes </font>
def predict(sess,image_file):
image,image_data = preprocess_image("images/"+image_file,model_image_size=(608,608))
out_scores , out_boxes , out_classes = sess.run([scores , boxes , classes], feed_dict={yolo_model.input:image_data,K.learning_phase():0})
print('Found {} boxes for {}'.format(len(out_boxes),image_file))
colors = generate_colors(class_names)
draw_boxes(image,out_scores,out_boxes,out_classes,class_names,colors)
image.save(os.path.join("out",image_file),quality=90)
output_image = scipy.misc.imread(os.path.join("out", image_file))
imshow(output_image)
return out_scores, out_boxes, out_classes
out_scores, out_boxes, out_classes = predict(sess, "0015.jpg")
#
# If you were to run your session in a for loop over all your images. Here's what you would get:
#
# <center>
# <video width="400" height="200" src="nb_images/pred_video_compressed2.mp4" type="video/mp4" controls>
# </video>
# </center>
#
# <caption><center> Predictions of the YOLO model on pictures taken from a camera while driving around the Silicon Valley <br> Thanks [drive.ai](https://www.drive.ai/) for providing this dataset! </center></caption>
|
MyNotebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="PaDZ4sMpmgsE" colab_type="text"
# # Load library
# + id="4dggTO9llzzj" colab_type="code" colab={}
# #!pip install shap
# !pip install pyitlib
# + id="n17VUTTpmEb0" colab_type="code" outputId="27375e38-6dab-4d35-a292-a3dbc50c12e4" executionInfo={"status": "ok", "timestamp": 1584842643368, "user_tz": 300, "elapsed": 39218, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 139}
from google.colab import drive
drive.mount('/content/drive')
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
your_module = drive.CreateFile({'id':'14_Wk3mGMp53nEiSjMUkeCno_LvRtyQBH'})
your_module.GetContentFile('Bayesian_class.py')
from Bayesian_class import *
import os
os.path.abspath(os.getcwd())
os.chdir('/content/drive/My Drive/Protein project')
os.path.abspath(os.getcwd())
# + [markdown] id="hN1BDWJcmtpc" colab_type="text"
# ### P450
# + id="80g_xJh4mL-L" colab_type="code" colab={}
def readData(filename):
fr = open(filename)
returnData = []
headerLine=fr.readline()###move cursor
for line in fr.readlines():
lineStrip = line.strip().replace('"','')
lineList = lineStrip.split('\t')
returnData.append(lineList)###['3','2',...]
return returnData
"""first case P450 = [['1','1',....],[],[].....,[]] second case P450 = array([['1','1',....],[],[].....,[]]), third case P450 = """
P450 = readData('P450.txt') ### [[],[],[],....[]]
P450 = np.array(P450) ### either [['1','1',....],[],[].....,[]] or array([['1','1',....],[],[].....,[]]) works, but note that keys are '1', '0'
#P450 = P450.astype(int) ### for shap array [[1,1,....],[],[].....,[]], keys are 1, 0
M=np.matrix([[245, 9, 0, 3, 0, 2, 65, 8],
[9, 218, 17, 17, 49, 10, 50, 17],
[0, 17, 175, 16, 25, 13, 0, 46],
[3, 17, 16, 194, 19, 0, 0, 3],
[0, 49, 25, 19, 199, 10, 0, 3],
[2, 10, 13, 0, 10, 249, 50, 74],
[65, 50, 0, 0, 0, 50, 262, 11],
[8, 17, 46, 3, 3, 74, 11, 175]])
X = P450[:,0:8]
y = P450[:,-1]
# + [markdown] id="orpgFiYJnGnk" colab_type="text"
# ### Lactamase
# + id="zv0TMePRnCme" colab_type="code" colab={}
def readData2(filename):
fr = open(filename)
returnData = []
headerLine=fr.readline()###move cursor
for line in fr.readlines():
linestr = line.strip().replace(', ','')
lineList = list(linestr)
returnData.append(lineList)###['3','2',...]
return returnData
lactamase = readData2('lactamase.txt')
lactamase = np.array(lactamase)
#lactamase = lactamase.astype(int)
M2 = np.matrix([[101, 5, 0, 2, 0, 14, 4, 37],
[5 ,15, 14 ,1 ,7 ,7, 0 ,19],
[0, 14, 266, 15, 14, 2, 26, 4],
[2, 1, 15, 28, 2 ,15, 4, 0],
[0, 7, 14, 2, 32, 9 ,0, 8],
[14, 7, 2 ,15, 9, 29, 7, 9],
[4, 0, 26, 4 ,0 ,7 ,72, 21],
[37, 19, 4, 0, 8, 9, 21, 211]])
X2 = lactamase[:,0:8]
y2 = lactamase[:,-1]
# + [markdown] id="d5VERFZJnK-0" colab_type="text"
# ### lymph
# + id="TGb6uPKRuJEz" colab_type="code" colab={}
def readarff(filename):
arrfFile = open(filename)
lines = [line.rstrip('\n') for line in arrfFile]
data = [[]]
index = 0
for line in lines :
if(line.startswith('@attribute')) :
index+=1
elif(not line.startswith('@data') and not line.startswith('@relation') and not line.startswith('%')) :
data.append(line.split(','))
del data[0]
return data
lymph_train = readarff("others/lymph_train.arff.txt"); lymph_train = np.array(lymph_train)
lymph_test = readarff("others/lymph_test.arff.txt") ;lymph_test = np.array(lymph_test)
lymph = np.concatenate((lymph_train,lymph_test))
X10 = lymph[:,0:18]
y10 = lymph[:,-1]
# + [markdown] id="bEx6fEzQu0DH" colab_type="text"
# ### Vote
# + id="IXDQRBCXu2BX" colab_type="code" colab={}
vote_train = readarff("others/vote_train.arff.txt") ;vote_train = np.array(vote_train)
vote_test = readarff("others/vote_test.arff.txt") ; vote_test = np.array(vote_test)
vote = np.concatenate((vote_train,vote_test))
X11 = vote[:,0:16]
y11 = vote[:,-1]
# + [markdown] id="utT2vQgXwQiW" colab_type="text"
# ### german
# + id="Nk_3gl3VwXag" colab_type="code" colab={}
def readData3(filename):
fr = open(filename)
returnData = []
for line in fr.readlines():
lineList = line.strip().split(' ')
returnData.append(lineList)###['3','2',...]
return returnData
german = readData3("data folder/german.data")
german = np.array(german)
X15 = german[:,0:20]
y15 = german[:,-1]
# + [markdown] id="yQc_xPq9xKdz" colab_type="text"
# ### balance scale
# + id="xpixkuBOxP_m" colab_type="code" colab={}
def readData3(filename):
fr = open(filename)
returnData = []
for line in fr.readlines():
lineList = line.strip().split(',')
returnData.append(lineList)###['3','2',...]
return returnData
balance = readData3("data folder/balance-scale.data")
balance = np.array(balance)
X14 = balance[:,1:5]
y14 = balance[:,0]
# + [markdown] id="fJ4x_fVH_SxU" colab_type="text"
# ### Hayes
# + id="kSfrDLBw_YrO" colab_type="code" colab={}
def readData3(filename):
fr = open(filename)
returnData = []
for line in fr.readlines():
lineList = line.strip().split(',')
returnData.append(lineList)###['3','2',...]
return returnData
hayes = readData3("data folder/hayes-roth.data")
hayes = np.array(hayes)
X13 = hayes[:,1:5] ## remove row index
y13 = hayes[:,-1]
# + [markdown] id="IfAhrJfE_4sh" colab_type="text"
# ### mushroom
# + id="EfT829N9_73b" colab_type="code" colab={}
def readData3(filename):
fr = open(filename)
returnData = []
for line in fr.readlines():
lineList = line.strip().split(',')
returnData.append(lineList)###['3','2',...]
return returnData
mushroom = readData3("data folder/agaricus-lepiota.data")
mushroom = np.array(mushroom)
X12 = mushroom[:,1:23]
y12 = mushroom[:,0] ## first column is y
# + [markdown] id="Sq6-FG8gSxz9" colab_type="text"
# ### connect 4
# + id="JvBMy7EYS0fJ" colab_type="code" colab={}
def readData4(filename):
fr = open(filename)
returnData = []
for line in fr.readlines():
lineList = line.strip().split(',')
returnData.append(lineList)###['3','2',...]
return returnData
connect_4 = readData4("data folder/connect-4.data")
connect_4 = np.array(connect_4)
X4 = connect_4[:,0:42]
y4 = connect_4[:,-1]
# + [markdown] id="CK3ywA1aSi7q" colab_type="text"
# ### cars
# + id="WwYpFbbCSp1S" colab_type="code" colab={}
car = readData4("data folder/car.data")
car = np.array(car)
X6 = car[:,0:6]
y6 = car[:,-1]
# + [markdown] id="NMZmO1kWUUcv" colab_type="text"
# ### krkopt
# + id="d2yWtc-xUWdN" colab_type="code" colab={}
krkopt = readData4("data folder/krkopt.data")
krkopt = np.array(krkopt)
X5 = krkopt[:,0:6]
y5 = krkopt[:,-1]
# + [markdown] id="s0OU35oigski" colab_type="text"
# ### Cmc
# + id="ZNnnkjU2g4fP" colab_type="code" colab={}
cmc = readData4("data folder/cmc.data")
cmc = np.array(cmc)
X7 = cmc[:,0:9]
y7 = cmc[:,-1]
# + [markdown] id="pQH59QvIhjYQ" colab_type="text"
# ### nurse
# + id="I87Lj9OThk61" colab_type="code" colab={}
def readData3(filename):
fr = open(filename)
returnData = []
for line in fr.readlines():
lineList = line.strip().split(',')
returnData.append(lineList)###['3','2',...]
return returnData
nurse = readData3("data folder/nursery.data")
nurse.pop()
nurse = np.array(nurse)
X3 = nurse[:,0:8]
y3 = nurse[:,-1]
# + [markdown] id="OOj9GzQ4nTee" colab_type="text"
# # Cross validation
# + [markdown] id="NO2-zlt3hrDo" colab_type="text"
# ### Nurse
# + id="llew9hauhsYC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="ed2cf430-c62f-4c57-94d8-0f202c1406cc" executionInfo={"status": "ok", "timestamp": 1584851779324, "user_tz": 300, "elapsed": 6073, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X3,y3,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="4eXMHOv8hsa-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="b21f1006-8373-4cd3-9ece-aa6caeaf199b" executionInfo={"status": "ok", "timestamp": 1584851815598, "user_tz": 300, "elapsed": 34279, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X3,y3,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="BGNQJOJZhsed" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="72029a48-bbc4-49b5-f3e7-dee33923c1e8" executionInfo={"status": "ok", "timestamp": 1584852115511, "user_tz": 300, "elapsed": 268766, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X3,y3,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="ep5MJBkEg6HT" colab_type="text"
# ### Cmc
# + id="37IrsfThg7un" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="de319783-b576-4510-c4a4-b9011e4af517" executionInfo={"status": "ok", "timestamp": 1584851582617, "user_tz": 300, "elapsed": 818, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X7,y7,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="J2lu4l1Rg7xc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="c56449f7-258c-4616-c1b2-71cedb11f15a" executionInfo={"status": "ok", "timestamp": 1584851602380, "user_tz": 300, "elapsed": 9960, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X7,y7,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="kWJmssQTg7zt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="9684208a-3830-4adc-fd63-cf2f4cb26847" executionInfo={"status": "ok", "timestamp": 1584851687410, "user_tz": 300, "elapsed": 82363, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X7,y7,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="vCgaMNlxUYP0" colab_type="text"
# ### krkopt
# + id="5p5Jk5PfUZ0h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="7d0c7cad-0193-4cd0-d524-4c9cea7ff695" executionInfo={"status": "ok", "timestamp": 1584848324697, "user_tz": 300, "elapsed": 32741, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X5,y5,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="kb70xyMtUZ3U" colab_type="code" colab={}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X5,y5,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="iMq37ypSUZ6f" colab_type="code" colab={}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X5,y5,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="zytcDkXYS3hI" colab_type="text"
# ### cars
# + id="aGp1XAosS4wb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="a9215200-1d3b-408f-cdb8-aeb27aecfe9a" executionInfo={"status": "ok", "timestamp": 1584847897113, "user_tz": 300, "elapsed": 1114, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X6,y6,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="PCkcBx96S4zu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="e4676bb1-7a5b-47e2-b234-252ad258b321" executionInfo={"status": "ok", "timestamp": 1584847915401, "user_tz": 300, "elapsed": 4193, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X6,y6,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="TfwWcifCS46Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="99ae0212-5751-4c4b-9d02-ef5bf9c56517" executionInfo={"status": "ok", "timestamp": 1584847940213, "user_tz": 300, "elapsed": 22574, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X6,y6,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="Qh7CX4K6__M8" colab_type="text"
# ### Mushroom
# + id="xxn9EGoNABhF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="1777d3c1-ffc3-473b-e87f-189e3c2d8c2c" executionInfo={"status": "ok", "timestamp": 1584842954640, "user_tz": 300, "elapsed": 3985, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X12,y12,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="m-oxODDyABkC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="dce0b82a-3bd0-4fc9-a249-352978e6e011" executionInfo={"status": "ok", "timestamp": 1584843072810, "user_tz": 300, "elapsed": 110994, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X12,y12,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="rOciQqDYABv_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="4fc1d6f5-85ed-41f1-89b8-62147173c656" executionInfo={"status": "ok", "timestamp": 1584845520746, "user_tz": 300, "elapsed": 2422792, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X12,y12,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="SUDYn1tX_pHh" colab_type="text"
# ### Hayes
# + id="hP93DAAh_qip" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="52d152e4-76c3-41ff-d5c0-8171281e2f30" executionInfo={"status": "ok", "timestamp": 1584842867087, "user_tz": 300, "elapsed": 408, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X13,y13,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="Y3PwZRSa_qmF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="88ca4b5a-0c6a-4635-b43d-ddc25262dc3b" executionInfo={"status": "ok", "timestamp": 1584842877121, "user_tz": 300, "elapsed": 1135, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X13,y13,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="5DgmyZMG_qpq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="3c086cf8-084c-4cf1-8d8b-bd2e02255752" executionInfo={"status": "ok", "timestamp": 1584842888762, "user_tz": 300, "elapsed": 3160, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X13,y13,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="F2Q-ujWAxUiQ" colab_type="text"
# ### balance scale
# + id="Z26k7KwaxWmA" colab_type="code" outputId="09b80931-2b93-4d5a-a7ae-07e1e761ce6d" executionInfo={"status": "ok", "timestamp": 1584821818949, "user_tz": 300, "elapsed": 411, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X14,y14,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="q-YGxU48xWo3" colab_type="code" outputId="f32e10c6-6a34-49d7-b607-a457d69e7539" executionInfo={"status": "ok", "timestamp": 1584821829273, "user_tz": 300, "elapsed": 1483, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X14,y14,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="EMZWquZyxWrH" colab_type="code" outputId="443c8e3a-3721-44a6-f6a0-1ee7d02547c0" executionInfo={"status": "ok", "timestamp": 1584821838372, "user_tz": 300, "elapsed": 4477, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X14,y14,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="bMGtnmFbwZAm" colab_type="text"
# ### german
# + id="JStmZSDRwaqW" colab_type="code" outputId="af6f7a29-799f-4edc-8a6f-e8d6b4050486" executionInfo={"status": "ok", "timestamp": 1584821869397, "user_tz": 300, "elapsed": 743, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X15,y15,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="dJJtAVLZwas5" colab_type="code" outputId="d5c36801-305e-48f8-b221-30f7b37b1bc3" executionInfo={"status": "ok", "timestamp": 1584822025190, "user_tz": 300, "elapsed": 149772, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X15,y15,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="Ycj_EFiGwavm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="61b39e64-aab1-4e96-c7dc-6e1b6b22de79" executionInfo={"status": "ok", "timestamp": 1584825101535, "user_tz": 300, "elapsed": 3031918, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X15,y15,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="7Gqsymp6vNtP" colab_type="text"
# ### vote
# + id="4ps2zIzBvPi5" colab_type="code" outputId="61e00537-6880-4556-b134-01bf237e8c6c" executionInfo={"status": "ok", "timestamp": 1584827755737, "user_tz": 300, "elapsed": 541, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X11,y11,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="z5R-419XvlzC" colab_type="code" outputId="582fb0c7-0139-402c-d16b-d65d03ed386e" executionInfo={"status": "ok", "timestamp": 1584827771614, "user_tz": 300, "elapsed": 12655, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X11,y11,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="hRnnSsUfvoaQ" colab_type="code" outputId="91090d64-df5f-4672-94dd-90f86e6c9d16" executionInfo={"status": "ok", "timestamp": 1584827968846, "user_tz": 300, "elapsed": 194857, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X11,y11,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="tmq3FMq2uEmh" colab_type="text"
# ### lymph
# + id="wZUWrDvGuGHK" colab_type="code" outputId="cef38789-068a-415d-f5cf-93a4e9efed99" executionInfo={"status": "ok", "timestamp": 1584829617464, "user_tz": 300, "elapsed": 398, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X10,y10,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="WlgbgcEbua5c" colab_type="code" outputId="24d73ca6-aadd-44ce-9934-46133596dc5b" executionInfo={"status": "ok", "timestamp": 1584829633335, "user_tz": 300, "elapsed": 13499, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X10,y10,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="8gPOcvfSudDr" colab_type="code" outputId="3d80ec29-7022-472b-8623-a5c6dcb03e18" executionInfo={"status": "ok", "timestamp": 1584830346435, "user_tz": 300, "elapsed": 236869, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X10,y10,None,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="JrP7SZ92n8z0" colab_type="text"
# ### P450
# + id="JxpMIz_dnVZ5" colab_type="code" outputId="caa83ca1-1ee8-43d7-f15d-926920bf73c3" executionInfo={"status": "ok", "timestamp": 1584842678775, "user_tz": 300, "elapsed": 526, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X,y,M,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="Rz0U20wcn3MM" colab_type="code" outputId="3209e896-da32-4698-c22a-86db0b1ef20d" executionInfo={"status": "ok", "timestamp": 1584770081703, "user_tz": 300, "elapsed": 4099, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X,y,M,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="mWfoUpXln7lk" colab_type="code" outputId="55402492-8c78-43fb-e2cc-3a6c62f64a52" executionInfo={"status": "ok", "timestamp": 1584770083861, "user_tz": 300, "elapsed": 770, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(STAN,X,y,M,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="QhhMpj-NoBS2" colab_type="code" outputId="46d62ac3-748a-4fda-c6ae-9de2f4a865fa" executionInfo={"status": "ok", "timestamp": 1584770118475, "user_tz": 300, "elapsed": 30732, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X,y,M,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="f4aHzwqOoJ8n" colab_type="code" outputId="bc272d2d-fcbf-466e-c6bf-3ed2e86d229b" executionInfo={"status": "ok", "timestamp": 1584770121651, "user_tz": 300, "elapsed": 27717, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(STAN_bagging,X,y,M,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="HCl98escoMER" colab_type="code" outputId="c5bba799-6633-4984-d6f0-e274094f5f98" executionInfo={"status": "ok", "timestamp": 1584770161074, "user_tz": 300, "elapsed": 33630, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(STAN_TAN_bagging,X,y,M,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + [markdown] id="0bwEzEzOoyVw" colab_type="text"
# ### lactamase
# + id="_F5kyeWno1U_" colab_type="code" outputId="04eecae8-7193-49ae-8bb0-cee0106bd05f" executionInfo={"status": "ok", "timestamp": 1584842708819, "user_tz": 300, "elapsed": 480, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(NB,X2,y2,M2,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="R_dRyVtmo1dn" colab_type="code" outputId="e0dfa4c5-3561-4bd8-eb8f-011754118572" executionInfo={"status": "ok", "timestamp": 1584770358319, "user_tz": 300, "elapsed": 3688, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN,X2,y2,M2,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="XCJW2pfbo7uy" colab_type="code" outputId="d7cfd492-ef23-485d-8fdb-06add4dcb933" executionInfo={"status": "ok", "timestamp": 1584770455385, "user_tz": 300, "elapsed": 457, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(STAN,X2,y2,M2,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="cGcGNTmKo_VD" colab_type="code" outputId="e44872a8-0a8e-4968-d0cc-f500f05e617b" executionInfo={"status": "ok", "timestamp": 1584770416569, "user_tz": 300, "elapsed": 25450, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(TAN_bagging,X2,y2,M2,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="zMikb_q8pNZV" colab_type="code" outputId="fd4aab2b-962a-4d37-965f-444a2b90284e" executionInfo={"status": "ok", "timestamp": 1584770475252, "user_tz": 300, "elapsed": 2268, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(STAN_bagging,X2,y2,M2,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
# + id="noJ-0FhpppUz" colab_type="code" outputId="ddcdb10e-7f51-49df-94bd-1f8c03a223b3" executionInfo={"status": "ok", "timestamp": 1584770514870, "user_tz": 300, "elapsed": 28272, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02205225540197780949"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
Accuracy, CLL, training_time,Precision,Recall,F1 = get_cv(STAN_TAN_bagging,X2,y2,M2,10,verbose=False)
print("%s +/- %s" % (np.mean(Accuracy), np.std(Accuracy) ) )
print("%s +/- %s" % (np.mean(CLL), np.std(CLL) ) )
print("%s +/- %s" % (np.mean(Precision), np.std(Precision) ) )
print("%s +/- %s" % (np.mean(Recall), np.std(Recall) ) )
print("%s +/- %s" % (np.mean(F1), np.std(F1) ) )
print("%s +/- %s" % (np.mean(np.array(training_time),axis=0), np.std(np.array(training_time),axis=0) ) )
|
examples/Simulation_supervised.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import datetime
cities = pd.read_csv("./Resources/cities.csv")
cities.head()
cities['Date'] = pd.to_datetime(cities['Date'],unit='s')
cities['Date'] = cities['Date'].dt.strftime('%Y-%m-%d')
cities
cities.set_index('City_ID', inplace=True)
cities
cities.dtypes
cities_html = cities.to_html()
print(cities_html)
|
Assets/cities.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="SFDpSxHixpvk"
# <h1><font size=12>
# Weather Derivatites </h1>
# <h1> Rainfall Simulator -- LSTM <br></h1>
#
# Developed by [<NAME>](mailto:<EMAIL>) <br>
# 16 September 2018
#
# + colab={} colab_type="code" id="sm2luX0Vxpvm"
# Import needed libraries.
import numpy as np
import pandas as pd
import random as rand
import matplotlib.pyplot as plt
from scipy.stats import bernoulli
from scipy.stats import gamma
import pickle
import time
import datetime
from keras.models import load_model
# + colab={"base_uri": "https://localhost:8080/", "height": 1394} colab_type="code" id="E16mD9Xyxuyb" outputId="b2038382-a0bd-412b-b783-75010863119e"
# Download files.
# ! wget https://github.com/jesugome/WeatherDerivatives/raw/master/datasets/ensoForecastProb/ensoForecastProbabilities.pickle
# ! wget https://raw.githubusercontent.com/jesugome/WeatherDerivatives/master/results/visibleMarkov/transitionsParametersDry.csv
# ! wget https://raw.githubusercontent.com/jesugome/WeatherDerivatives/master/results/visibleMarkov/transitionsParametersWet.csv
# ! wget https://raw.githubusercontent.com/jesugome/WeatherDerivatives/master/results/visibleMarkov/amountGamma.csv
# ! wget https://github.com/jesugome/WeatherDerivatives/raw/master/results/visibleMarkov/rainfall_lstmDry_LSTM.h5
# ! wget https://github.com/jesugome/WeatherDerivatives/raw/master/results/visibleMarkov/rainfall_lstmWet_LSTM.h5
# + [markdown] colab_type="text" id="F64ZbFy6xpvv"
# # Generate artificial Data
# + colab={"base_uri": "https://localhost:8080/", "height": 359} colab_type="code" id="aaitHj5Oxpvw" outputId="c038ffb6-e23b-491f-e79b-871f4b5632e1"
### ENSO probabilistic forecast.
# Open saved data.
#ensoForecast = pickle.load(open('../datasets/ensoForecastProb/ensoForecastProbabilities.pickle','rb'))
ensoForecast = pickle.load(open('ensoForecastProbabilities.pickle','rb'))
# Print an example .. ( Format needed)
ensoForecast['2005-01']
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="_3uON7aXxpv5" outputId="9cba99bf-8623-411f-cc6f-60ddffe4222f"
### Create total dataframe.
def createTotalDataFrame(daysNumber, startDate , initialState , initialPrep , ensoForecast ):
# Set variables names.
totalDataframeColumns = ['state','Prep','Month','probNina','probNino', 'nextState']
# Create dataframe.
allDataDataframe = pd.DataFrame(columns=totalDataframeColumns)
# Number of simulation days(i.e 30, 60)
daysNumber = daysNumber
# Simulation start date ('1995-04-22')
startDate = startDate
# State of rainfall last day before start date --> Remember 0 means dry and 1 means wet.
initialState = initialState
initialPrep = initialPrep # Only fill when initialState == 1
dates = pd.date_range(startDate, periods = daysNumber + 2 , freq='D')
for date in dates:
# Fill precipitation amount.
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'Prep'] = np.nan
# Fill month of date
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'Month'] = date.month
# Fill El Nino ENSO forecast probability.
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'probNino'] = float(ensoForecast[date.strftime('%Y-%m')].loc[0,'El Niño'].strip('%').strip('~'))/100
# Fill La Nina ENSO forecast probability.
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'probNina'] = float(ensoForecast[date.strftime('%Y-%m')].loc[0,'La Niña'].strip('%').strip('~'))/100
# Fill State.
allDataDataframe.loc[date.strftime('%Y-%m-%d'),'state'] = np.nan
simulationDataFrame = allDataDataframe[:-1]
# Fill initial conditions.
simulationDataFrame['state'][0] = initialState
if initialState == 1:
simulationDataFrame['Prep'][0] = initialPrep
else:
simulationDataFrame['Prep'][0] = 0.0
return simulationDataFrame
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-08-18', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
simulationDataFrame.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 306} colab_type="code" id="vgTGzSctxpwA" outputId="07cb1885-c0d1-4506-a8c3-7447d892417b"
### Load transitions and amount parameters.
# Transitions probabilites.
transitionsParametersDry = pd.read_csv('transitionsParametersDry.csv', sep = ' ', header=None, names = ['variable', 'value'])
#transitionsParametersDry = pd.read_csv('../results/visibleMarkov/transitionsParametersDry.csv', sep = ' ', header=None, names = ['variable', 'value'])
transitionsParametersDry.index += 1
transitionsParametersDry
transitionsParametersWet = pd.read_csv('transitionsParametersWet.csv', sep = ' ', header=None, names = ['variable', 'value'])
#transitionsParametersWet = pd.read_csv('../results/visibleMarkov/transitionsParametersWet.csv', sep = ' ', header=None, names = ['variable', 'value'])
transitionsParametersWet.index += 1
transitionsParametersWet
amountParametersGamma = pd.read_csv('amountGamma.csv', sep = ' ', header=None, names = ['variable', 'loge(mu)', 'loge(shape)'])
#amountParametersGamma = pd.read_csv('../results/visibleMarkov/amountGamma.csv', sep = ' ', header=None, names = ['variable', 'loge(mu)', 'loge(shape)'])
amountParametersGamma.index += 1
print(amountParametersGamma)
print('\n * Intercept means firts month (January) ')
# Load neural network.
lstmModelDry = load_model('rainfall_lstmDry_LSTM.h5')
#lstmModel = load_model('../results/visibleMarkov/rainfall_lstmDry.h5')
# Load neural network.
lstmModelWet = load_model('rainfall_lstmWet_LSTM.h5')
#lstmModel = load_model('../results/visibleMarkov/rainfall_lstmWet.h5')
# + [markdown] colab_type="text" id="otpZv7iZxpwF"
# ## Simulation Function Core
# + colab={} colab_type="code" id="TeNMDSL7xpwI"
### Build the simulation core.
# Updates the state of the day based on yesterday state.
def updateState(yesterdayIndex, simulationDataFrame, transitionsParametersDry, transitionsParametersWet):
# Additional data of day.
yesterdayState = simulationDataFrame['state'][yesterdayIndex]
yesterdayPrep = simulationDataFrame['Prep'][yesterdayIndex]
yesterdayProbNino = simulationDataFrame['probNino'][yesterdayIndex]
yesterdayProbNina = simulationDataFrame['probNina'][yesterdayIndex]
yesterdayMonth = simulationDataFrame['Month'][yesterdayIndex]
# Calculate transition probability.
if yesterdayState == 0:
xPredict = np.array([(yesterdayMonth-1)/11,yesterdayProbNino,yesterdayProbNina])
xPredict = np.reshape(xPredict, ( 1, 1 , xPredict.shape[0]))
# Includes month factor + probNino value + probNino value.
successProbability = lstmModelDry.predict(xPredict)[0][0]
elif yesterdayState == 1:
xPredict = np.array([yesterdayPrep ,(yesterdayMonth-1)/11,yesterdayProbNino,yesterdayProbNina])
xPredict = np.reshape(xPredict, ( 1, 1 , xPredict.shape[0]))
# Includes month factor + probNino value + probNino value.
successProbability = lstmModelWet.predict(xPredict)[0][0]
else:
print('State of date: ', simulationDataFrame.index[yesterdayIndex],' not found.')
#print(successProbability)
todayState = bernoulli.rvs(successProbability)
return todayState
# + colab={} colab_type="code" id="wXTy_M15xpwN"
# Simulates one run of simulation.
def oneRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma):
# Define the total rainfall amount over the simulation.
rainfall = 0
# Loop over days in simulation to calculate rainfall ammount.
for day in range(1,len(simulationDataFrame)):
# Get today date.
dateOfDay = datetime.datetime.strptime(simulationDataFrame.index[day],'%Y-%m-%d')
# Update today state based on the yesterday state.
todayState = updateState(day-1, simulationDataFrame, transitionsParametersDry, transitionsParametersWet)
# Write new day information.
simulationDataFrame['state'][day] = todayState
simulationDataFrame['nextState'][day-1] = todayState
# Computes total accumulated rainfall.
if todayState == 1:
# Additional data of day.
todayProbNino = simulationDataFrame['probNino'][day]
todayProbNina = simulationDataFrame['probNina'][day]
todayMonth = simulationDataFrame['Month'][day]
# Calculates gamma log(mu).
gammaLogMU = amountParametersGamma['loge(mu)'][todayMonth]+ todayProbNino*amountParametersGamma['loge(mu)'][13]+todayProbNino*amountParametersGamma['loge(mu)'][13]
# Calculates gamma scale
gammaLogShape = amountParametersGamma['loge(shape)'][1]
# Update mu
gammaMu = np.exp(gammaLogMU)
# Update shape
gammaShape = np.exp(gammaLogShape)
# Calculate gamma scale.
gammaScale = gammaMu / gammaShape
# Generate random rainfall.
todayRainfall = gamma.rvs(a = gammaShape, scale = gammaScale)
# Write new day information.
simulationDataFrame['Prep'][day] = todayRainfall
# Updates rainfall amount.
rainfall += todayRainfall
else:
# Write new day information.
simulationDataFrame['Prep'][day] = 0
yesterdayState = todayState
return rainfall
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="CZxTMJwGxpwR" outputId="f1f9b25b-075b-4c4e-85f7-2226585b3da0"
updateState(0, simulationDataFrame, transitionsParametersDry, transitionsParametersWet)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="B_Lko5RExpwY" outputId="f84b4cb1-c131-4ab9-c123-486d4efe91c4"
# Run only one iteration(Print structure of results)
# Simulations iterations.
iterations = 10000
oneRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma)
# + [markdown] colab_type="text" id="q9b5zyXixpwe"
# ## Complete Simulation
# + colab={} colab_type="code" id="82jNLg_Kxpwf"
# Run total iterations.
def totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations):
# Initialize time
startTime = time.time()
# Array to store all precipitations.
rainfallPerIteration = [None]*iterations
# Loop over each iteration(simulation)
for i in range(iterations):
simulationDataFrameC = simulationDataFrame.copy()
iterationRainfall = oneRun(simulationDataFrameC, transitionsParametersDry, transitionsParametersWet, amountParametersGamma)
rainfallPerIteration[i] = iterationRainfall
# Calculate time
currentTime = time.time() - startTime
# Logging time.
print('The elapsed time over simulation is: ', currentTime, ' seconds.')
return rainfallPerIteration
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" id="n77y1lh7xpwl" outputId="6756a8ad-f22d-467b-e129-f431ea38ebe5"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-08-18', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
simulationDataFrame.head()
# + [markdown] colab_type="text" id="MkBHHPb6xpws"
# ## Final Results
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="6qIrbBFsxpwu" outputId="93246f09-1781-4a4f-d09a-172c6e464230"
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
# + colab={"base_uri": "https://localhost:8080/", "height": 620} colab_type="code" id="90cm0lbxxpwy" outputId="eb762372-995e-467b-d273-5075d05dbe93"
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + [markdown] colab_type="text" id="O6HjUS9rKDka"
# ### Enero
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="SRCwtMqVJ2xC" outputId="b2f41a5d-0af4-49f1-cf33-9581e1cf03b7"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-01-01', initialState = 0 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='lightgreen',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="RCZvC9IXKGTn" outputId="f0fff46b-bc80-4932-bb5e-f8627b760096"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-01-01', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='skyblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="KpQnAFIAKZ9_" outputId="4d426d3a-c011-4e98-ee14-f5096d30b118"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-01-01', initialState = 1 , initialPrep = 2.0 , ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + [markdown] colab_type="text" id="XnLdCFkRMdK7"
# ### Abril
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="DDnBPj-jMdK8" outputId="acbf117b-eadf-4fa5-d581-6725f6b930b5"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-04-01', initialState = 0 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='lightgreen',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="Pr7TTg8wMdLB" outputId="0bbf70be-2257-4325-b07c-e7b59886c950"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-04-01', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='skyblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="zApFwpupMdLG" outputId="0b48d535-4440-4724-e865-135760b5cadc"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-04-01', initialState = 1 , initialPrep = 2.0 , ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + [markdown] colab_type="text" id="M_wUgFfPMxTX"
# ### Octubre
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="_yMlAHPxMxTY" outputId="9fa2aa27-4e9c-4ddf-f7f3-04dee2d9b106"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-10-01', initialState = 0 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='lightgreen',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="SOictOlNMxTc" outputId="9ab61a60-8786-4362-c9d7-39acaa9a54ee"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-10-01', initialState = 1 , initialPrep = 0.4, ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='skyblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 722} colab_type="code" id="q3uiw2cLMxTm" outputId="8220bb00-b21f-4b5b-99d3-f135f835dfc4"
#### Define parameters simulation.
# Simulations iterations.
iterations = 1000
# Create dataframe to simulate.
simulationDataFrame = createTotalDataFrame(daysNumber= 30, startDate = '2017-10-01', initialState = 1 , initialPrep = 2.0 , ensoForecast = ensoForecast)
# Final Analysis.
finalSimulation = totalRun(simulationDataFrame, transitionsParametersDry, transitionsParametersWet, amountParametersGamma,iterations)
fig = plt.figure(figsize=(20, 10))
plt.hist(finalSimulation,facecolor='steelblue',bins=100, density=True,
histtype='stepfilled', edgecolor = 'black' , hatch = '+')
plt.title('Rainfall Simulation')
plt.xlabel('Rainfall Amount [mm]')
plt.ylabel('Probability ')
plt.grid()
plt.show()
# + colab={} colab_type="code" id="t7EaOsvEM3Za"
|
code/rainfallSimulator_NNVersion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import Needed Libraries
# +
import numpy as np
import pandas as pd
from sklearn.metrics import classification_report,confusion_matrix
# %matplotlib inline
# -
# ### Load Pokémon Names Data
#
# In the later generations we get "mega" evolutions, for this reason we want to only keep the default Pokémon
# +
url = 'https://raw.githubusercontent.com/veekun/pokedex/master/pokedex/data/csv/pokemon.csv'
all_pokemon = pd.read_csv(url, index_col = 0)
all_pokemon = all_pokemon[all_pokemon.is_default == 1]
all_pokemon.head(5)
# -
# ### Assign Generations to Each Pokémon
# +
def assign_generation(row):
if 0 < row['species_id'] <= 151:
return 'Generation I'
elif 151 < row['species_id'] <= 251:
return 'Generation II'
elif 251 < row['species_id'] <= 386:
return 'Generation III'
elif 386 < row['species_id'] <= 493:
return 'Generation IV'
elif 493 < row['species_id'] <= 649:
return 'Generation V'
elif 649 < row['species_id'] <= 721:
return 'Generation VI'
elif 721 < row['species_id'] <= 807:
return 'Generation VII'
else:
return 'other'
all_pokemon['generation'] = all_pokemon.apply(assign_generation, axis=1)
all_pokemon.head(5)
# -
# ## Build a Markov Chain
#
# This is based off of https://www.kaggle.com/naldershof/tweet-like-the-president-simple-markov
# We build a function that takes a series of strings and builds a dictionary of each letter and all letters that follow it - including the end of the word. While looping through the data, we also collect a list of starting letters and get the longest and shortest name.
def build_mc(corpus):
markov_dict = {'<EOT>':[]}
starting_letters = []
max_length = 0
min_length = 1000
for word in corpus:
tok = list(word) #make character list [l,i,k,e, ,t,h,i,s]
letter_count = len(tok) #length of word
#storing the max & min values of names
if(letter_count > max_length):
max_length = letter_count
if(letter_count < min_length):
min_length = letter_count
for index, letter in enumerate(tok):
#add letter if we haven't yet
if letter not in markov_dict.keys():
markov_dict[letter] = []
#add first letters to start list
if index == 0:
starting_letters.append(letter)
#add end of text to last letters of names
if index == letter_count - 1:
markov_dict[letter].append("<EOT>")
#add next letter to non-last letters
else:
markov_dict[letter].append(tok[index+1])
return markov_dict, starting_letters, max_length, min_length
# ## Build Markov Chains for each Generation of Pokémon
#
# For each generation we build a seperate model so that we can understand the differences
#hard code for each generation
markov_dict_1, starting_letters_1, max_length_1, min_length_1 = build_mc(all_pokemon[all_pokemon.generation == 'Generation I']['identifier'])
markov_dict_2, starting_letters_2, max_length_2, min_length_2 = build_mc(all_pokemon[all_pokemon.generation == 'Generation II']['identifier'])
markov_dict_3, starting_letters_3, max_length_3, min_length_3 = build_mc(all_pokemon[all_pokemon.generation == 'Generation III']['identifier'])
markov_dict_4, starting_letters_4, max_length_4, min_length_4 = build_mc(all_pokemon[all_pokemon.generation == 'Generation IV']['identifier'])
markov_dict_5, starting_letters_5, max_length_5, min_length_5 = build_mc(all_pokemon[all_pokemon.generation == 'Generation V']['identifier'])
markov_dict_6, starting_letters_6, max_length_6, min_length_6 = build_mc(all_pokemon[all_pokemon.generation == 'Generation VI']['identifier'])
markov_dict_7, starting_letters_7, max_length_7, min_length_7 = build_mc(all_pokemon[all_pokemon.generation == 'Generation VII']['identifier'])
# See what follows an x in each generation
print(markov_dict_1['x'])
print(markov_dict_2['x'])
print(markov_dict_3['x'])
print(markov_dict_4['x'])
print(markov_dict_5['x'])
print(markov_dict_6['x'])
print(markov_dict_7['x'])
# ## Generating New Pokémon
#
# We can do random walks on each Markov Chain to invent some new Pokémon - notice the differences in generations, for example we have a lot more dashes in our last generation.
#
# My personal favorite is telelucry :)
def new_pokemon_name(starting_letter, mc, max_length, min_length):
new_name = starting_letter
current_letter = starting_letter
while len(new_name) < max_length:
next_letter = np.random.choice(mc[current_letter])
#names have to be a least a certain length
while( (len(new_name) < min_length) & (next_letter == '<EOT>') ):
next_letter = np.random.choice(mc[current_letter])
if next_letter == '<EOT>':
return new_name
new_name = new_name + next_letter
current_letter = next_letter
return new_name
print('Generation I')
for x in range(0,5):
print(new_pokemon_name(np.random.choice(starting_letters_1), markov_dict_1, max_length_1,min_length_1))
print('\nGeneration II')
for x in range(0,5):
print(new_pokemon_name(np.random.choice(starting_letters_2), markov_dict_2, max_length_2,min_length_2))
print('\nGeneration III')
for x in range(0,5):
print(new_pokemon_name(np.random.choice(starting_letters_3), markov_dict_3, max_length_3,min_length_3))
print('\nGeneration IV')
for x in range(0,5):
print(new_pokemon_name(np.random.choice(starting_letters_4), markov_dict_4, max_length_4,min_length_4))
print('\nGeneration V')
for x in range(0,5):
print(new_pokemon_name(np.random.choice(starting_letters_5), markov_dict_5, max_length_5,min_length_5))
print('\nGeneration VI')
for x in range(0,5):
print(new_pokemon_name(np.random.choice(starting_letters_6), markov_dict_6, max_length_6,min_length_6))
print('\nGeneration VII')
for x in range(0,5):
print(new_pokemon_name(np.random.choice(starting_letters_7), markov_dict_7, max_length_7,min_length_7))
# ## Predict the Generation of Pokémon
#
# Now that we've invented some new Pokés we're going to predict the generation of a Pokémon based just on it's name. Because each model is built on a tiny dataset (80 to 160 names) we are __absolutely going to cheat__ and use models built on the full data set. In the real work we would train test split when checking to see if our models are working or not.
#
# We calculate the probability of one letter following another by going to the key, counting the number of times the next value happens, and dividing this by the total letters. This gives us a precent of the time that one letter follows the next. We then multiply the probabilities together and also multiply this by the probability of the starting letter.
#
# After getting the likelihood of a word in every model, we choose the most likely as our prediction.
#
# If the word is impossible in every model (for example: 666) it will return "No Prediction".
def generation_probability(word,starting_letters,markov_dict):
tok_word = list(word)
letter_count = len(tok_word) #length of word
probability = 1
for index, letter in enumerate(tok_word):
if(index == 0):
probability = probability * starting_letters.count('m') / starting_letters.__len__()
if index == letter_count - 1:
return probability
else:
probability = probability * markov_dict[letter].count(tok_word[index+1]) / markov_dict[letter].__len__()
def predicted_generation(row):
probabilities = pd.concat([
pd.DataFrame([[row['identifier'],'Generation I',generation_probability(row['identifier'],starting_letters_1,markov_dict_1)]]
,columns = ['identifier','generation','probability'])
,pd.DataFrame([[row['identifier'],'Generation II',generation_probability(row['identifier'],starting_letters_2,markov_dict_2)]]
,columns = ['identifier','generation','probability'])
,pd.DataFrame([[row['identifier'],'Generation III',generation_probability(row['identifier'],starting_letters_3,markov_dict_3)]]
,columns = ['identifier','generation','probability'])
,pd.DataFrame([[row['identifier'],'Generation IV',generation_probability(row['identifier'],starting_letters_4,markov_dict_4)]]
,columns = ['identifier','generation','probability'])
,pd.DataFrame([[row['identifier'],'Generation V',generation_probability(row['identifier'],starting_letters_5,markov_dict_5)]]
,columns = ['identifier','generation','probability'])
,pd.DataFrame([[row['identifier'],'Generation VI',generation_probability(row['identifier'],starting_letters_6,markov_dict_6)]]
,columns = ['identifier','generation','probability'])
,pd.DataFrame([[row['identifier'],'Generation VII',generation_probability(row['identifier'],starting_letters_7,markov_dict_7)]]
,columns = ['identifier','generation','probability'])
])
highest_prob = probabilities['probability'].max()
if(highest_prob == 0):
return 'No Prediction'
# return np.random.choice(['Generation I','Generation II','Generation III'
# ,'Generation IV', 'Generation V', 'Generation VI','Generation VII'])
return probabilities[probabilities.probability == highest_prob]['generation']
all_pokemon['prediction'] = all_pokemon.apply(predicted_generation, axis=1)
# #### We've Got Impressive Results!
#
# If we were to just guess the generation randomly, we would expect accuracies of ~1/7 or 14%. We know that we are giving the models a big advantage by training and testing on the same data. Even so, our prediction results are much much better than 14%. It's tempting to then claim that the names of Pokémon really did change from season to season, we proved it! And yes, there were some changes like loner names and more dashes. However, our training data sets are so tiny that we definitely just have over fitted models :)
print(classification_report(all_pokemon.generation,all_pokemon.prediction))
print(confusion_matrix(all_pokemon.generation,all_pokemon.prediction))
# ## What Generation of Pokémon Am I ???
#
# Finally, let's take some non-pokemon words and see what generation they are most likely to be from
mt = pd.DataFrame(['michelle','tanco','hunter','teradata','xx'], columns =['identifier'])
mt['generation'] = mt.apply(predicted_generation, axis=1)
mt
|
_jupyter/2018-10-09-mc-pokemon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Sentiment Analysis
#
# ## Using XGBoost in SageMaker
#
# _Deep Learning Nanodegree Program | Deployment_
#
# ---
#
# In this example of using Amazon's SageMaker service we will construct a random tree model to predict the sentiment of a movie review. You may have seen a version of this example in a pervious lesson although it would have been done using the sklearn package. Instead, we will be using the XGBoost package as it is provided to us by Amazon.
#
# ## Instructions
#
# Some template code has already been provided for you, and you will need to implement additional functionality to successfully complete this notebook. You will not need to modify the included code beyond what is requested. Sections that begin with '**TODO**' in the header indicate that you need to complete or implement some portion within them. Instructions will be provided for each section and the specifics of the implementation are marked in the code block with a `# TODO: ...` comment. Please be sure to read the instructions carefully!
#
# In addition to implementing code, there may be questions for you to answer which relate to the task and your implementation. Each section where you will answer a question is preceded by a '**Question:**' header. Carefully read each question and provide your answer below the '**Answer:**' header by editing the Markdown cell.
#
# > **Note**: Code and Markdown cells can be executed using the **Shift+Enter** keyboard shortcut. In addition, a cell can be edited by typically clicking it (double-click for Markdown cells) or by pressing **Enter** while it is highlighted.
# ## Step 1: Downloading the data
#
# The dataset we are going to use is very popular among researchers in Natural Language Processing, usually referred to as the [IMDb dataset](http://ai.stanford.edu/~amaas/data/sentiment/). It consists of movie reviews from the website [imdb.com](http://www.imdb.com/), each labeled as either '**pos**itive', if the reviewer enjoyed the film, or '**neg**ative' otherwise.
#
# > Maas, <NAME>., et al. [Learning Word Vectors for Sentiment Analysis](http://ai.stanford.edu/~amaas/data/sentiment/). In _Proceedings of the 49th Annual Meeting of the Association for Computational Linguistics: Human Language Technologies_. Association for Computational Linguistics, 2011.
#
# We begin by using some Jupyter Notebook magic to download and extract the dataset.
# %mkdir ../data
# !wget -O ../data/aclImdb_v1.tar.gz http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz
# !tar -zxf ../data/aclImdb_v1.tar.gz -C ../data
# ## Step 2: Preparing the data
#
# The data we have downloaded is split into various files, each of which contains a single review. It will be much easier going forward if we combine these individual files into two large files, one for training and one for testing.
# +
import os
import glob
def read_imdb_data(data_dir='../data/aclImdb'):
data = {}
labels = {}
for data_type in ['train', 'test']:
data[data_type] = {}
labels[data_type] = {}
for sentiment in ['pos', 'neg']:
data[data_type][sentiment] = []
labels[data_type][sentiment] = []
path = os.path.join(data_dir, data_type, sentiment, '*.txt')
files = glob.glob(path)
for f in files:
with open(f) as review:
data[data_type][sentiment].append(review.read())
# Here we represent a positive review by '1' and a negative review by '0'
labels[data_type][sentiment].append(1 if sentiment == 'pos' else 0)
assert len(data[data_type][sentiment]) == len(labels[data_type][sentiment]), \
"{}/{} data size does not match labels size".format(data_type, sentiment)
return data, labels
# -
data, labels = read_imdb_data()
print("IMDB reviews: train = {} pos / {} neg, test = {} pos / {} neg".format(
len(data['train']['pos']), len(data['train']['neg']),
len(data['test']['pos']), len(data['test']['neg'])))
# +
from sklearn.utils import shuffle
def prepare_imdb_data(data, labels):
"""Prepare training and test sets from IMDb movie reviews."""
#Combine positive and negative reviews and labels
data_train = data['train']['pos'] + data['train']['neg']
data_test = data['test']['pos'] + data['test']['neg']
labels_train = labels['train']['pos'] + labels['train']['neg']
labels_test = labels['test']['pos'] + labels['test']['neg']
#Shuffle reviews and corresponding labels within training and test sets
data_train, labels_train = shuffle(data_train, labels_train)
data_test, labels_test = shuffle(data_test, labels_test)
# Return a unified training data, test data, training labels, test labets
return data_train, data_test, labels_train, labels_test
# -
train_X, test_X, train_y, test_y = prepare_imdb_data(data, labels)
print("IMDb reviews (combined): train = {}, test = {}".format(len(train_X), len(test_X)))
train_X[100]
# ## Step 3: Processing the data
#
# Now that we have our training and testing datasets merged and ready to use, we need to start processing the raw data into something that will be useable by our machine learning algorithm. To begin with, we remove any html formatting that may appear in the reviews and perform some standard natural language processing in order to homogenize the data.
import nltk
nltk.download("stopwords")
from nltk.corpus import stopwords
from nltk.stem.porter import *
stemmer = PorterStemmer()
# +
import re
from bs4 import BeautifulSoup
def review_to_words(review):
text = BeautifulSoup(review, "html.parser").get_text() # Remove HTML tags
text = re.sub(r"[^a-zA-Z0-9]", " ", text.lower()) # Convert to lower case
words = text.split() # Split string into words
words = [w for w in words if w not in stopwords.words("english")] # Remove stopwords
words = [PorterStemmer().stem(w) for w in words] # stem
return words
# +
import pickle
cache_dir = os.path.join("../cache", "sentiment_analysis") # where to store cache files
os.makedirs(cache_dir, exist_ok=True) # ensure cache directory exists
def preprocess_data(data_train, data_test, labels_train, labels_test,
cache_dir=cache_dir, cache_file="preprocessed_data.pkl"):
"""Convert each review to words; read from cache if available."""
# If cache_file is not None, try to read from it first
cache_data = None
if cache_file is not None:
try:
with open(os.path.join(cache_dir, cache_file), "rb") as f:
cache_data = pickle.load(f)
print("Read preprocessed data from cache file:", cache_file)
except:
pass # unable to read from cache, but that's okay
# If cache is missing, then do the heavy lifting
if cache_data is None:
# Preprocess training and test data to obtain words for each review
#words_train = list(map(review_to_words, data_train))
#words_test = list(map(review_to_words, data_test))
words_train = [review_to_words(review) for review in data_train]
words_test = [review_to_words(review) for review in data_test]
# Write to cache file for future runs
if cache_file is not None:
cache_data = dict(words_train=words_train, words_test=words_test,
labels_train=labels_train, labels_test=labels_test)
with open(os.path.join(cache_dir, cache_file), "wb") as f:
pickle.dump(cache_data, f)
print("Wrote preprocessed data to cache file:", cache_file)
else:
# Unpack data loaded from cache file
words_train, words_test, labels_train, labels_test = (cache_data['words_train'],
cache_data['words_test'], cache_data['labels_train'], cache_data['labels_test'])
return words_train, words_test, labels_train, labels_test
# -
# Preprocess data
train_X, test_X, train_y, test_y = preprocess_data(train_X, test_X, train_y, test_y)
# ### Extract Bag-of-Words features
#
# For the model we will be implementing, rather than using the reviews directly, we are going to transform each review into a Bag-of-Words feature representation. Keep in mind that 'in the wild' we will only have access to the training set so our transformer can only use the training set to construct a representation.
# +
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.externals import joblib
# joblib is an enhanced version of pickle that is more efficient for storing NumPy arrays
def extract_BoW_features(words_train, words_test, vocabulary_size=5000,
cache_dir=cache_dir, cache_file="bow_features.pkl"):
"""Extract Bag-of-Words for a given set of documents, already preprocessed into words."""
# If cache_file is not None, try to read from it first
cache_data = None
if cache_file is not None:
try:
with open(os.path.join(cache_dir, cache_file), "rb") as f:
cache_data = joblib.load(f)
print("Read features from cache file:", cache_file)
except:
pass # unable to read from cache, but that's okay
# If cache is missing, then do the heavy lifting
if cache_data is None:
# Fit a vectorizer to training documents and use it to transform them
# NOTE: Training documents have already been preprocessed and tokenized into words;
# pass in dummy functions to skip those steps, e.g. preprocessor=lambda x: x
vectorizer = CountVectorizer(max_features=vocabulary_size,
preprocessor=lambda x: x, tokenizer=lambda x: x) # already preprocessed
features_train = vectorizer.fit_transform(words_train).toarray()
# Apply the same vectorizer to transform the test documents (ignore unknown words)
features_test = vectorizer.transform(words_test).toarray()
# NOTE: Remember to convert the features using .toarray() for a compact representation
# Write to cache file for future runs (store vocabulary as well)
if cache_file is not None:
vocabulary = vectorizer.vocabulary_
cache_data = dict(features_train=features_train, features_test=features_test,
vocabulary=vocabulary)
with open(os.path.join(cache_dir, cache_file), "wb") as f:
joblib.dump(cache_data, f)
print("Wrote features to cache file:", cache_file)
else:
# Unpack data loaded from cache file
features_train, features_test, vocabulary = (cache_data['features_train'],
cache_data['features_test'], cache_data['vocabulary'])
# Return both the extracted features as well as the vocabulary
return features_train, features_test, vocabulary
# -
# Extract Bag of Words features for both training and test datasets
train_X, test_X, vocabulary = extract_BoW_features(train_X, test_X)
# ## Step 4: Classification using XGBoost
#
# Now that we have created the feature representation of our training (and testing) data, it is time to start setting up and using the XGBoost classifier provided by SageMaker.
#
# ### Writing the dataset
#
# The XGBoost classifier that we will be using requires the dataset to be written to a file and stored using Amazon S3. To do this, we will start by splitting the training dataset into two parts, the data we will train the model with and a validation set. Then, we will write those datasets to a file and upload the files to S3. In addition, we will write the test set input to a file and upload the file to S3. This is so that we can use SageMakers Batch Transform functionality to test our model once we've fit it.
# +
import pandas as pd
val_X = pd.DataFrame(train_X[:10000])
train_X = pd.DataFrame(train_X[10000:])
val_y = pd.DataFrame(train_y[:10000])
train_y = pd.DataFrame(train_y[10000:])
test_y = pd.DataFrame(test_y)
test_X = pd.DataFrame(test_X)
# -
# The documentation for the XGBoost algorithm in SageMaker requires that the saved datasets should contain no headers or index and that for the training and validation data, the label should occur first for each sample.
#
# For more information about this and other algorithms, the SageMaker developer documentation can be found on __[Amazon's website.](https://docs.aws.amazon.com/sagemaker/latest/dg/)__
# First we make sure that the local directory in which we'd like to store the training and validation csv files exists.
data_dir = '../data/xgboost'
if not os.path.exists(data_dir):
os.makedirs(data_dir)
# +
# First, save the test data to test.csv in the data_dir directory. Note that we do not save the associated ground truth
# labels, instead we will use them later to compare with our model output.
# Solution:
# The test data shouldn't contain the ground truth labels as they are what the model is
# trying to predict. We will end up using them afterward to compare the predictions to.
# pd.concat([test_y, test_X], axis=1).to_csv(os.path.join(data_dir, 'test.csv'), header=False, index=False)
pd.DataFrame(test_X).to_csv(os.path.join(data_dir, 'test.csv'), header=False, index=False)
pd.concat([val_y, val_X], axis=1).to_csv(os.path.join(data_dir, 'validation.csv'), header=False, index=False)
pd.concat([train_y, train_X], axis=1).to_csv(os.path.join(data_dir, 'train.csv'), header=False, index=False)
# +
# To save a bit of memory we can set text_X, train_X, val_X, train_y and val_y to None.
train_X = val_X = train_y = val_y = None
# -
# ### Uploading Training / Validation files to S3
#
# Amazon's S3 service allows us to store files that can be access by both the built-in training models such as the XGBoost model we will be using as well as custom models such as the one we will see a little later.
#
# For this, and most other tasks we will be doing using SageMaker, there are two methods we could use. The first is to use the low level functionality of SageMaker which requires knowing each of the objects involved in the SageMaker environment. The second is to use the high level functionality in which certain choices have been made on the user's behalf. The low level approach benefits from allowing the user a great deal of flexibility while the high level approach makes development much quicker. For our purposes we will opt to use the high level approach although using the low-level approach is certainly an option.
#
# Recall the method `upload_data()` which is a member of object representing our current SageMaker session. What this method does is upload the data to the default bucket (which is created if it does not exist) into the path described by the key_prefix variable. To see this for yourself, once you have uploaded the data files, go to the S3 console and look to see where the files have been uploaded.
#
# For additional resources, see the __[SageMaker API documentation](http://sagemaker.readthedocs.io/en/latest/)__ and in addition the __[SageMaker Developer Guide.](https://docs.aws.amazon.com/sagemaker/latest/dg/)__
# +
import sagemaker
session = sagemaker.Session() # Store the current SageMaker session
# S3 prefix (which folder will we use)
prefix = 'sentiment-xgboost'
test_location = session.upload_data(os.path.join(data_dir, 'test.csv'), key_prefix=prefix)
val_location = session.upload_data(os.path.join(data_dir, 'validation.csv'), key_prefix=prefix)
train_location = session.upload_data(os.path.join(data_dir, 'train.csv'), key_prefix=prefix)
# -
# ### (TODO) Creating a tuned XGBoost model
#
# Now that the data has been uploaded it is time to create the XGBoost model. As in the Boston Housing notebook, the first step is to create an estimator object which will be used as the *base* of your hyperparameter tuning job.
# +
from sagemaker import get_execution_role
# Our current execution role is require when creating the model as the training
# and inference code will need to access the model artifacts.
role = get_execution_role()
# +
# We need to retrieve the location of the container which is provided by Amazon for using XGBoost.
# As a matter of convenience, the training and inference code both use the same container.
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(session.boto_region_name, 'xgboost')
# +
# TODO: Create a SageMaker estimator using the container location determined in the previous cell.
# It is recommended that you use a single training instance of type ml.m4.xlarge. It is also
# recommended that you use 's3://{}/{}/output'.format(session.default_bucket(), prefix) as the
# output path.
xgb = None
# Solution:
xgb = sagemaker.estimator.Estimator(container, # The location of the container we wish to use
role, # What is our current IAM Role
train_instance_count=1, # How many compute instances
train_instance_type='ml.m4.xlarge', # What kind of compute instances
output_path='s3://{}/{}/output'.format(session.default_bucket(), prefix),
sagemaker_session=session)
# TODO: Set the XGBoost hyperparameters in the xgb object. Don't forget that in this case we have a binary
# label so we should be using the 'binary:logistic' objective.
# Solution:
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
silent=0,
objective='binary:logistic',
early_stopping_rounds=10,
num_round=500)
# -
# ### (TODO) Create the hyperparameter tuner
#
# Now that the base estimator has been set up we need to construct a hyperparameter tuner object which we will use to request SageMaker construct a hyperparameter tuning job.
#
# **Note:** Training a single sentiment analysis XGBoost model takes longer than training a Boston Housing XGBoost model so if you don't want the hyperparameter tuning job to take too long, make sure to not set the total number of models (jobs) too high.
# +
# First, make sure to import the relevant objects used to construct the tuner
from sagemaker.tuner import IntegerParameter, ContinuousParameter, HyperparameterTuner
# TODO: Create the hyperparameter tuner object
xgb_hyperparameter_tuner = None
# Solution:
xgb_hyperparameter_tuner = HyperparameterTuner(estimator = xgb, # The estimator object to use as the basis for the training jobs.
objective_metric_name = 'validation:rmse', # The metric used to compare trained models.
objective_type = 'Minimize', # Whether we wish to minimize or maximize the metric.
max_jobs = 6, # The total number of models to train
max_parallel_jobs = 3, # The number of models to train in parallel
hyperparameter_ranges = {
'max_depth': IntegerParameter(3, 12),
'eta' : ContinuousParameter(0.05, 0.5),
'min_child_weight': IntegerParameter(2, 8),
'subsample': ContinuousParameter(0.5, 0.9),
'gamma': ContinuousParameter(0, 10),
})
# -
# ### Fit the hyperparameter tuner
#
# Now that the hyperparameter tuner object has been constructed, it is time to fit the various models and find the best performing model.
s3_input_train = sagemaker.s3_input(s3_data=train_location, content_type='csv')
s3_input_validation = sagemaker.s3_input(s3_data=val_location, content_type='csv')
xgb_hyperparameter_tuner.fit({'train': s3_input_train, 'validation': s3_input_validation})
# Remember that the tuning job is constructed and run in the background so if we want to see the progress of our training job we need to call the `wait()` method.
xgb_hyperparameter_tuner.wait()
# ### (TODO) Testing the model
#
# Now that we've run our hyperparameter tuning job, it's time to see how well the best performing model actually performs. To do this we will use SageMaker's Batch Transform functionality. Batch Transform is a convenient way to perform inference on a large dataset in a way that is not realtime. That is, we don't necessarily need to use our model's results immediately and instead we can peform inference on a large number of samples. An example of this in industry might be peforming an end of month report. This method of inference can also be useful to us as it means to can perform inference on our entire test set.
#
# Remember that in order to create a transformer object to perform the batch transform job, we need a trained estimator object. We can do that using the `attach()` method, creating an estimator object which is attached to the best trained job.
# +
# TODO: Create a new estimator object attached to the best training job found during hyperparameter tuning
xgb_attached = None
# Solution:
xgb_attached = sagemaker.estimator.Estimator.attach(xgb_hyperparameter_tuner.best_training_job())
# -
# Now that we have an estimator object attached to the correct training job, we can proceed as we normally would and create a transformer object.
# +
# TODO: Create a transformer object from the attached estimator. Using an instance count of 1 and an instance type of ml.m4.xlarge
# should be more than enough.
xgb_transformer = None
# Solution:
xgb_transformer = xgb_attached.transformer(instance_count = 1, instance_type = 'ml.m4.xlarge')
# -
# Next we actually perform the transform job. When doing so we need to make sure to specify the type of data we are sending so that it is serialized correctly in the background. In our case we are providing our model with csv data so we specify `text/csv`. Also, if the test data that we have provided is too large to process all at once then we need to specify how the data file should be split up. Since each line is a single entry in our data set we tell SageMaker that it can split the input on each line.
# TODO: Start the transform job. Make sure to specify the content type and the split type of the test data.
xgb_transformer.transform(test_location, content_type='text/csv', split_type='Line')
# Currently the transform job is running but it is doing so in the background. Since we wish to wait until the transform job is done and we would like a bit of feedback we can run the `wait()` method.
xgb_transformer.wait()
# Now the transform job has executed and the result, the estimated sentiment of each review, has been saved on S3. Since we would rather work on this file locally we can perform a bit of notebook magic to copy the file to the `data_dir`.
# !aws s3 cp --recursive $xgb_transformer.output_path $data_dir
# The last step is now to read in the output from our model, convert the output to something a little more usable, in this case we want the sentiment to be either `1` (positive) or `0` (negative), and then compare to the ground truth labels.
predictions = pd.read_csv(os.path.join(data_dir, 'test.csv.out'), header=None)
predictions = [round(num) for num in predictions.squeeze().values]
from sklearn.metrics import accuracy_score
accuracy_score(test_y, predictions)
# ## Optional: Clean up
#
# The default notebook instance on SageMaker doesn't have a lot of excess disk space available. As you continue to complete and execute notebooks you will eventually fill up this disk space, leading to errors which can be difficult to diagnose. Once you are completely finished using a notebook it is a good idea to remove the files that you created along the way. Of course, you can do this from the terminal or from the notebook hub if you would like. The cell below contains some commands to clean up the created files from within the notebook.
# +
# First we will remove all of the files contained in the data_dir directory
# !rm $data_dir/*
# And then we delete the directory itself
# !rmdir $data_dir
# Similarly we will remove the files in the cache_dir directory and the directory itself
# !rm $cache_dir/*
# !rmdir $cache_dir
# -
|
Mini-Projects/IMDB Sentiment Analysis - XGBoost (Hyperparameter Tuning) - Solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# 
# 
# # **DistROOT: CMS Example Notebook**
# <hr style="border-top-width: 4px; border-top-color: #34609b;">
# Get user credentials.
# +
import getpass
import os, sys
krb5ccname = '/tmp/krb5cc_' + os.environ['USER']
print("Please enter your password")
ret = os.system("echo \"%s\" | kinit -c %s" % (getpass.getpass(), krb5ccname))
if ret == 0: print("Credentials created successfully")
else: sys.stderr.write('Error creating credentials, return code: %s\n' % ret)
# -
# Import Spark modules.
from pyspark import SparkConf, SparkContext
# Create Spark configuration and context.
# +
conf = SparkConf()
# Generic for SWAN-Spark prototype
conf.set('spark.driver.host', os.environ['SERVER_HOSTNAME'])
conf.set('spark.driver.port', os.environ['SPARK_PORT_1'])
conf.set('spark.fileserver.port', os.environ['SPARK_PORT_2'])
conf.set('spark.blockManager.port', os.environ['SPARK_PORT_3'])
conf.set('spark.ui.port', os.environ['SPARK_PORT_4'])
conf.set('spark.master', 'yarn')
# DistROOT specific
conf.setAppName("ROOT")
conf.set('spark.executor.extraLibraryPath', os.environ['LD_LIBRARY_PATH'])
conf.set('spark.submit.pyFiles', os.environ['HOME'] + '/.local/lib/python2.7/site-packages/DistROOT.py')
conf.set('spark.executorEnv.KRB5CCNAME', krb5ccname)
conf.set('spark.yarn.dist.files', krb5ccname + '#krbcache')
# Resource allocation
conf.set('spark.executor.instances', 4)
conf.set('spark.driver.memory', '2g')
sc = SparkContext(conf = conf)
# -
# Import DistROOT.
import ROOT
from DistROOT import DistTree
# Define the mapper and reducer functions.
# +
def fillCMS(reader):
import ROOT
ROOT.TH1.AddDirectory(False)
ROOT.gInterpreter.Declare('#include "file.h"')
myAnalyzer = ROOT.wmassAnalyzer(reader)
return myAnalyzer.GetHistosList()
def mergeCMS(l1, l2):
for i in xrange(l1.GetSize()):
l1.At(i).Add(l2.At(i))
return l1
# -
# Build the DistTree and trigger the parallel processing.
# +
files = [ "data.root",
"data2.root" ]
dTree = DistTree(filelist = files,
treename = "random_test_tree",
npartitions = 8)
histList = dTree.ProcessAndMerge(fillCMS, mergeCMS)
# -
# Store resulting histograms in a file.
f = ROOT.TFile("output.root", "RECREATE")
for h in histList:
h.Write()
f.Close()
# Draw one of the histograms we filled using Spark and ROOT.
c = ROOT.TCanvas()
histList[0].Draw()
c.Draw()
|
notebooks/DistROOT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
"""
@description: Shopping List
@author:Yee
"""
from __future__ import print_function
from __future__ import unicode_literals
from matplotlib import pyplot as plt
from enum import Enum
from prettytable import PrettyTable
from IPython.display import display
import matplotlib.ticker as mticker
import pandas as pd
import seaborn as sns
import numpy as np
import datetime
import xlrd
import time
# +
""" @:parameter type: different share type as an enum"""
class ShareType(Enum):
AllShare = 1
Chris = 2
JoJo = 3
Yee = 4
ChrisAndJo = 5
ChrisAndYee = 6
JoAndYee = 7
# +
""" show shopping list in a pretty table"""
def showPersonList(index):
t = PrettyTable()
total = 0
t.field_names = ["ID", "Name", "Price", "Yee", "Chris", "Jossie", "Tax"]
print(time.strftime("%Y-%m-%d %H:%M:%S"))
for i in range(size):
if data[i][index] == 1:
t.add_row([i, data[i][0], data[i][1], data[i][2], data[i][3], data[i][4], data[i][5]])
if data[i][5] == 1:
total += data[i][1] * 1.15
else:
total += data[i][1]
totalPrice.append(total)
print(t)
print("Total:", total)
# +
""" show shopping list in a pandas"""
def showPersonList2(index):
# Total amount
total = 0
# Selected rows from whole dataFrame, for which the according person should pay
dataList = list()
columnNames = ["ID", "Name", "Price", "Yee", "Chris", "Jossie", "Tax", "Payer"]
for i in range(size):
if data[i][index] != 1:
continue
dataList.append([i, data[i][0], data[i][1], data[i][2], data[i][3], data[i][4], data[i][5], data[i][6]])
if data[i][5] == 1:
total += data[i][1] * 1.15
else:
total += data[i][1]
dFrame = pd.DataFrame(dataList, columns=columnNames)
display(dFrame)
payer = dFrame.Payer.unique()
print("Shopping List of", name[index-2])
for p in payer:
subtotal = 0
for i in range(dFrame.shape[0]):
if dataList[i][7] == p:
if dataList[i][6] == 1:
subtotal += dataList[i][2] * 1.15
else:
subtotal += dataList[i][2]
print(p, subtotal)
print("Total:", total)
# +
""" show item price distribution"""
def showDistribution(data):
for i in range(size):
priceList.append(data[i][1])
sns.displot(priceList, bins=100)
plt.style.use("dark_background")
plt.title("Distribution of price", fontsize=15, color = 'white')
plt.show()
# +
""" show subtotal pie graph"""
def showPiePlot(data):
sns.set_style("whitegrid")
plt.plot(np.arange(10))
plt.show()
# +
""" show item price distribution bar graph"""
def showBarGraph(data):
fig, ax = plt.subplots(1)
fig.set_size_inches(20,10)
position = np.arange(0, size*2, 2)
ax.bar(position, df['Price'].tolist(), width=1, color='darkslateblue' )
ax.set_xticklabels(df['Name'].tolist())
ax.set_xticks(position)
ax.set_ylabel('Price', fontsize=12)
ax.set_xlabel('Item', fontsize=12)
plt.title('Price distribution ', fontsize=26)
plt.xticks(rotation=90, fontsize=10)
plt.rcParams['font.sans-serif']=['SimHei']
# -
priceList = list()
name = ['Yee', 'Chris', 'Jo']
totalPrice = list()
df = pd.read_excel ('D:\Desktop\Shopping list.xlsx', 'Sheet5')
data = df.values
size = int(data.shape[0])
display(df)
df.info()
# +
for i in range(size):
divide = data[i][2] + data[i][3] + data[i][4]
data[i][1] = data[i][1] / divide
for i in range(2,5):
showPersonList2(i)
# showDistribution(data)
# showPiePlot(data)
showBarGraph(data)
# -
|
Python/Shopping-list.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torchvision
import torchvision.transforms as transforms
# +
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=4,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=4,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# +
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = Net()
# +
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
# +
for epoch in range(1): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
print('Finished Training')
# -
|
notebooks/torchvision-example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Batman, ecuaciones y python
# *Esta notebook fue creada originalmente como un blog post por [<NAME>](http://relopezbriega.com.ar/) en [Mi blog sobre Python](http://relopezbriega.github.io). El contenido esta bajo la licencia BSD.*
# <img alt="Ecuación Batman" title="Ecuación Batman" src="http://relopezbriega.github.io/images/ecnbatmancolor.png">
# [Batman](https://es.wikipedia.org/wiki/Batman) siempre fue mi superhéroe favorito porque es uno de los pocos héroes que no posee ningún superpoder, sino que debe recurrir a su intelecto y a la ciencia para construir las bati-herramientas que utiliza para combatir al crimen. Además posee ese toque de oscuridad producto de la dualidad entre realizar el bien, protegiendo a la gente de ciudad gótica, y su sed de venganza contra el crimen y la corrupción que acabó con la vida de su familia.
#
# Es un personaje con muchos recursos, en cada nueva aparación podemos verlo utilizar nuevas y muy modernas bati-herramientas; su intelecto es tan agudo que incluso escondió una ecuación matemática en su bati-señal!!
#
# La [ecuación de batman](http://www.wolframalpha.com/input/?i=batman+equation) fue creada por el profesor de matemáticas [Matthew Register](http://www.quora.com/J-Matthew-Register) y se pupolarizó a través de un post de uno de sus alumnos en la red social [reddit](https://www.reddit.com/r/pics/comments/j2qjc/do_you_like_batman_do_you_like_math_my_math/); su expesión matemática es la siguiente:
# $$
# ((\frac{x}{7})^2 \cdot \sqrt{\frac{||x|-3|}{(|x|-3)}}+ (\frac{y}{3})^2 \cdot \sqrt{\frac{|y+3 \cdot \frac{\sqrt{33}}{7}|}{y+3 \cdot \frac{\sqrt{33}}{7}}}-1) \cdot (|\frac{x}{2}|-((3 \cdot \frac{\sqrt{33}-7)}{112}) \cdot x^2-3+\sqrt{1-(||x|-2|-1)^2}-y) \cdot (9 \cdot \sqrt{\frac{|(|x|-1) \cdot (|x|-0.75)|}{((1-|x|)*(|x|-0.75))}}-8 \cdot |x|-y) \cdot (3 \cdot |x|+0.75 \cdot \sqrt{\frac{|(|x|-0.75) \cdot (|x|-0.5)|}{((0.75-|x|) \cdot (|x|-0.5))}}-y) \cdot (2.25 \cdot \sqrt{\frac{|(x-0.5) \cdot (x+0.5)|}{((0.5-x) \cdot (0.5+x))}}-y) \cdot (\frac{6 \cdot \sqrt{10}}{7}+(1.5-0.5 \cdot |x|) \cdot \sqrt{\frac{||x|-1|}{|x|-1}}-(\frac{6 \cdot \sqrt{10}}{14}) \cdot \sqrt{4-(|x|-1)^2}-y) =0
# $$
# Si bien a simple vista la ecuación parace sumamente compleja e imposible de graficar, la misma se puede descomponer en seis curvas distintas, mucho más simples.
#
# La primera de estas curvas, es la función del elipse $(\frac{x}{7})^2 + (\frac{y}{3})^2 = 1$, restringida a la región $\sqrt{\frac{||x|-3|}{(|x|-3)}}$ y $\sqrt{\frac{|y+3 \cdot \frac{\sqrt{33}}{7}|}{y+3 \cdot \frac{\sqrt{33}}{7}}}$ para cortar la parte central.
#
# Los cinco términos siguientes pueden ser entendidos como simples funciones de x, tres de los cuales son lineales.
# Por ejemplo, la siguente función es la que grafica las curvas de la parte inferior de la bati-señal.
#
# $y = |\frac{x}{2}|-(\frac{3 \cdot \sqrt{33} -7}{112})\cdot x^2 - 3 + \sqrt{1-(||x|-2| -1)^2}$
#
# Las restantes ecuaciones de las curvas que completan el gráfico, son las siguientes:
#
# $y = \frac{6\cdot\sqrt{10}}{7} + (-0.5|x| + 1.5) - \frac{3\cdot\sqrt{10}}{7}\cdot\sqrt{4 - (|x|-1)^2}, |x| > 1$
#
# $y = 9 -8|x|, 0.75 < |x| < 1$
#
# $y = 3|x| + 0.75, 0.5 < |x| < 0.75$
#
# $y = 2.25, |x| < 0.5$
#
# La [ecuación de batman](http://www.wolframalpha.com/input/?i=batman+equation) puede ser facilmente graficada utilizando [Matplotlib](http://matplotlib.org/) del siguiente modo:
# graficos embebidos
# %matplotlib inline
# Importando lo necesario para los cálculos
import matplotlib.pyplot as plt
from numpy import sqrt
from numpy import meshgrid
from numpy import arange
# +
# Graficando la ecuación de Batman.
xs = arange(-7.25, 7.25, 0.01)
ys = arange(-5, 5, 0.01)
x, y = meshgrid(xs, ys)
eq1 = ((x/7)**2*sqrt(abs(abs(x)-3)/(abs(x)-3))+(y/3)**2*sqrt(abs(y+3/7*sqrt(33))/(y+3/7*sqrt(33)))-1)
eq2 = (abs(x/2)-((3*sqrt(33)-7)/112)*x**2-3+sqrt(1-(abs(abs(x)-2)-1)**2)-y)
eq3 = (9*sqrt(abs((abs(x)-1)*(abs(x)-.75))/((1-abs(x))*(abs(x)-.75)))-8*abs(x)-y)
eq4 = (3*abs(x)+.75*sqrt(abs((abs(x)-.75)*(abs(x)-.5))/((.75-abs(x))*(abs(x)-.5)))-y)
eq5 = (2.25*sqrt(abs((x-.5)*(x+.5))/((.5-x)*(.5+x)))-y)
eq6 = (6*sqrt(10)/7+(1.5-.5*abs(x))*sqrt(abs(abs(x)-1)/(abs(x)-1))-(6*sqrt(10)/14)*sqrt(4-(abs(x)-1)**2)-y)
for f, c in [(eq1, "red"), (eq2, "purple"), (eq3, "green"),
(eq4, "blue"), (eq5, "orange"), (eq6, "black")]:
plt.contour(x, y, f, [0], colors=c)
plt.show()
# -
# Ahora ya saben...si están en algún apuro y necesitan la ayuda del bati-héroe, solo necesitan graficar una ecuación para llamarlo con la bati-señal!
#
# Saludos!
#
# *Este post fue escrito utilizando IPython notebook. Pueden descargar este [notebook](https://github.com/relopezbriega/relopezbriega.github.io/blob/master/downloads/Batman.ipynb) o ver su version estática en [nbviewer](http://nbviewer.ipython.org/github/relopezbriega/relopezbriega.github.io/blob/master/downloads/Batman.ipynb).*
|
content/notebooks/Batman.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp pybrms
# +
#export
#hide
import typing
import pandas as pd
import numpy as np
import pystan
import re
import rpy2.robjects.packages as rpackages
from rpy2.robjects import default_converter, pandas2ri, numpy2ri, ListVector, DataFrame, StrVector
from rpy2.robjects.conversion import localconverter
try:
brms = rpackages.importr("brms")
except:
utils = rpackages.importr("utils")
utils.chooseCRANmirror(ind=1)
utils.install_packages(StrVector(('brms',)))
brms = rpackages.importr("brms")
# -
# # Documentation
#export
def get_brms_data(dataset_name:str):
"A helper function for importing different datasets included in brms."
with localconverter(default_converter + pandas2ri.converter + numpy2ri.converter) as cv:
return pd.DataFrame(rpackages.data(brms).fetch(dataset_name)[dataset_name])
assert isinstance(get_brms_data("epilepsy"),pd.DataFrame)
assert isinstance(get_brms_data("kidney"),pd.DataFrame)
assert isinstance(get_brms_data("inhaler"),pd.DataFrame)
#export
def _convert_python_to_R(data: typing.Union[dict, pd.DataFrame]):
"""
Converts a python object to an R object brms can handle:
* python dict -> R list
* python dataframe -> R dataframe
"""
with localconverter(default_converter + pandas2ri.converter + numpy2ri.converter) as cv:
if isinstance(data, pd.DataFrame):
return DataFrame(data)
elif isinstance(data, dict):
return ListVector(data)
else:
raise ValueError("Data should be either a pandas dataframe or a dictionary")
assert isinstance(_convert_python_to_R(dict(a=1, b=2)),ListVector)
assert isinstance(_convert_python_to_R(get_brms_data("inhaler")),DataFrame)
#export
def get_stan_code(
formula: str,
data: typing.Union[dict, pd.DataFrame],
priors: list,
family: str,
sample_prior: str="no"
):
if len(priors)>0:
return brms.make_stancode(
formula=formula, data=data, prior=priors, family=family, sample_prior=sample_prior
)[0]
else:
return brms.make_stancode(
formula=formula, data=data, family=family, sample_prior=sample_prior
)[0]
#export
def _convert_R_to_python(
formula: str, data: typing.Union[dict, pd.DataFrame], family: str
):
# calls brms to preprocess the data; returns an R ListVector
model_data = brms.make_standata(formula, data, family=family)
# a context manager for conversion between R objects and python/pandas/numpy
# we're not activating it globally because it conflicts with creation of priors
with localconverter(default_converter + pandas2ri.converter + numpy2ri.converter) as cv:
model_data = dict(model_data.items())
return model_data
#export
def _coerce_types(stan_code, stan_data):
pat_data = re.compile(r'(?<=data {)[^}]*')
pat_identifiers = re.compile(r'([\w]+)')
# extract the data block and separate lines
data_lines = pat_data.findall(stan_code)[0].split('\n')
# remove commets, <>-style bounds and []-style data size declarations
data_lines_no_comments = [l.split('//')[0] for l in data_lines]
data_lines_no_bounds = [re.sub('<[^>]+>', '',l) for l in data_lines_no_comments]
data_lines_no_sizes = [re.sub('\[[^>]+\]', '',l) for l in data_lines_no_bounds]
# extract identifiers - first one should be the type, last one should be the name
identifiers = [pat_identifiers.findall(l) for l in data_lines_no_sizes]
var_types = [l[0] for l in identifiers if len(l)>0]
var_names = [l[-1] for l in identifiers if len(l)>0]
var_dict = dict(zip(var_names, var_types))
# coerce integers to int and 1-size arrays to scalars
for k,v in stan_data.items():
if k in var_names and var_dict[k]=="int":
stan_data[k] = v.astype(int)
if v.size==1:
stan_data[k] = stan_data[k][0]
return stan_data
#export
def fit(
formula: str,
data: typing.Union[dict, pd.DataFrame],
priors: list = [],
family: str = "gaussian",
sample_prior: str = "no",
sample:bool = "yes",
**pystan_args,
):
formula = brms.bf(formula)
data = _convert_python_to_R(data)
if len(priors)>0:
brms_prior = brms.prior_string(*priors[0])
for p in priors[1:]:
brms_prior = brms_prior + brms.prior_string(*p)
assert brms.is_brmsprior(brms_prior)
else:
brms_prior = []
model_code = get_stan_code(
formula=formula,
data=data,
family=family,
priors=brms_prior,
sample_prior=sample_prior,
)
model_data = _convert_R_to_python(formula, data, family)
model_data = _coerce_types(model_code, model_data)
sm = pystan.StanModel(model_code=model_code)
if sample==False:
return sm
else:
fit = sm.sampling(data=model_data, **pystan_args)
return fit
from nbdev.showdoc import *
from nbdev.export import *
notebook2script()
|
core.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3_UZH
# language: python
# name: python3_uzh
# ---
# +
import os
import os.path
import random
from operator import add
from datetime import datetime, date, timedelta
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
import shutil
import time
from scipy.integrate import simps
from numpy import trapz
from decimal import Decimal, ROUND_DOWN, ROUND_UP
# -
pd.set_option('display.max_columns',69)
pd.set_option('display.max_rows',119)
# +
# load the data
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case1_sattel-hochstuckli\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case2_Atzmaening\setup1'
root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case3_hoch-ybrig\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case4_villars-diablerets_elevations_b1339\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case4_villars-diablerets_elevations_b1822\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case4_villars-diablerets_elevations_b2000\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case4_villars-diablerets_elevations_b2500\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case5_champex\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case6_davos_elevations_b1564\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case6_davos_elevations_b2141\setup1'
#root = r'C:\Saeid\Prj100\SA_2\snowModelUZH\case6_davos_elevations_b2584\setup1'
rootOut = os.path.join(root, 'Results_3')
df_final_tipping_point_1980 = pd.read_csv(os.path.join(rootOut, 'df_final_tipping_point_1980.csv'))
df_final_tipping_point_2020 = pd.read_csv(os.path.join(rootOut, 'df_final_tipping_point_2020.csv'))
df_final_tipping_point_2050 = pd.read_csv(os.path.join(rootOut, 'df_final_tipping_point_2050.csv'))
df_final_tipping_point_2070 = pd.read_csv(os.path.join(rootOut, 'df_final_tipping_point_2070.csv'))
df_policies = pd.read_csv(os.path.join(rootOut, 'df_policies.csv'))
# -
df_final_tipping_point_1980
# +
#####Scenario_26####
tippingPoint26_All_1980 = df_final_tipping_point_1980['tippingPoint26_1980'].value_counts()
tippingPoint26_Accepted_1980 = df_final_tipping_point_1980['tippingPoint26_1_1980'].value_counts()
scenario26_1980 = df_final_tipping_point_1980['scenario26_1980'].value_counts()
policy26_1980 = df_final_tipping_point_1980['policy26_1980'].value_counts()
tippingPoint26_All_2020 = df_final_tipping_point_2020['tippingPoint26_2020'].value_counts()
tippingPoint26_Accepted_2020 = df_final_tipping_point_2020['tippingPoint26_1_2020'].value_counts()
scenario26_2020 = df_final_tipping_point_2020['scenario26_2020'].value_counts()
policy26_2020 = df_final_tipping_point_2020['policy26_2020'].value_counts()
tippingPoint26_All_2050 = df_final_tipping_point_2050['tippingPoint26_2050'].value_counts()
tippingPoint26_Accepted_2050 = df_final_tipping_point_2050['tippingPoint26_1_2050'].value_counts()
scenario26_2050 = df_final_tipping_point_2050['scenario26_2050'].value_counts()
policy26_2050 = df_final_tipping_point_2050['policy26_2050'].value_counts()
tippingPoint26_All_2070 = df_final_tipping_point_2070['tippingPoint26_2070'].value_counts()
tippingPoint26_Accepted_2070 = df_final_tipping_point_2070['tippingPoint26_1_2070'].value_counts()
scenario26_2070 = df_final_tipping_point_2070['scenario26_2070'].value_counts()
policy26_2070 = df_final_tipping_point_2070['policy26_2070'].value_counts()
#####Scenario_45####
tippingPoint45_All_1980 = df_final_tipping_point_1980['tippingPoint45_1980'].value_counts()
tippingPoint45_Accepted_1980 = df_final_tipping_point_1980['tippingPoint45_1_1980'].value_counts()
scenario45_1980 = df_final_tipping_point_1980['scenario45_1980'].value_counts()
policy45_1980 = df_final_tipping_point_1980['policy45_1980'].value_counts()
tippingPoint45_All_2020 = df_final_tipping_point_2020['tippingPoint45_2020'].value_counts()
tippingPoint45_Accepted_2020 = df_final_tipping_point_2020['tippingPoint45_1_2020'].value_counts()
scenario45_2020 = df_final_tipping_point_2020['scenario45_2020'].value_counts()
policy45_2020 = df_final_tipping_point_2020['policy45_2020'].value_counts()
tippingPoint45_All_2050 = df_final_tipping_point_2050['tippingPoint45_2050'].value_counts()
tippingPoint45_Accepted_2050 = df_final_tipping_point_2050['tippingPoint45_1_2050'].value_counts()
scenario45_2050 = df_final_tipping_point_2050['scenario45_2050'].value_counts()
policy45_2050 = df_final_tipping_point_2050['policy45_2050'].value_counts()
tippingPoint45_All_2070 = df_final_tipping_point_2070['tippingPoint45_2070'].value_counts()
tippingPoint45_Accepted_2070 = df_final_tipping_point_2070['tippingPoint45_1_2070'].value_counts()
scenario45_2070 = df_final_tipping_point_2070['scenario45_2070'].value_counts()
policy45_2070 = df_final_tipping_point_2070['policy45_2070'].value_counts()
#####Scenario_85####
tippingPoint85_All_1980 = df_final_tipping_point_1980['tippingPoint85_1980'].value_counts()
tippingPoint85_Accepted_1980 = df_final_tipping_point_1980['tippingPoint85_1_1980'].value_counts()
scenario85_1980 = df_final_tipping_point_1980['scenario85_1980'].value_counts()
policy85_1980 = df_final_tipping_point_1980['policy85_1980'].value_counts()
tippingPoint85_All_2020 = df_final_tipping_point_2020['tippingPoint85_2020'].value_counts()
tippingPoint85_Accepted_2020 = df_final_tipping_point_2020['tippingPoint85_1_2020'].value_counts()
scenario85_2020 = df_final_tipping_point_2020['scenario85_2020'].value_counts()
policy85_2020 = df_final_tipping_point_2020['policy85_2020'].value_counts()
tippingPoint85_All_2050 = df_final_tipping_point_2050['tippingPoint85_2050'].value_counts()
tippingPoint85_Accepted_2050 = df_final_tipping_point_2050['tippingPoint85_1_2050'].value_counts()
scenario85_2050 = df_final_tipping_point_2050['scenario85_2050'].value_counts()
policy85_2050 = df_final_tipping_point_2050['policy85_2050'].value_counts()
tippingPoint85_All_2070 = df_final_tipping_point_2070['tippingPoint85_2070'].value_counts()
tippingPoint85_Accepted_2070 = df_final_tipping_point_2070['tippingPoint85_1_2070'].value_counts()
scenario85_2070 = df_final_tipping_point_2070['scenario85_2070'].value_counts()
policy85_2070 = df_final_tipping_point_2070['policy85_2070'].value_counts()
# +
a1 = pd.DataFrame(tippingPoint26_All_1980.reset_index().values, columns=["freq26_tip_all_1980", "tippingPoint26_all_1980"])
a2 = pd.DataFrame(tippingPoint26_Accepted_1980.reset_index().values, columns=["freq26_tipacc_1980", "tippingPoint26_acc_1980"])
a3 = pd.DataFrame(scenario26_1980.reset_index().values, columns=["Policy26_1980", "freq26_1980_policy"])
a4 = pd.DataFrame(policy26_1980.reset_index().values, columns=["scenario26_1980", "freq26_1980_scenario"])
a5 = pd.DataFrame(tippingPoint26_All_2020.reset_index().values, columns=["freq26_tip_all_2020", "tippingPoint26_all_2020"])
a6 = pd.DataFrame(tippingPoint26_Accepted_2020.reset_index().values, columns=["freq26_tipacc_2020", "tippingPoint26_acc_2020"])
a7 = pd.DataFrame(scenario26_2020.reset_index().values, columns=["Policy26_2020", "freq26_2020_policy"])
a8 = pd.DataFrame(policy26_2020.reset_index().values, columns=["scenario26_2020", "freq26_2020_scenario"])
a9 = pd.DataFrame(tippingPoint26_All_2050.reset_index().values, columns=["freq26_tip_all_2050", "tippingPoint26_all_2050"])
a10 = pd.DataFrame(tippingPoint26_Accepted_2050.reset_index().values, columns=["freq26_tipacc_2050", "tippingPoint26_acc_2050"])
a11 = pd.DataFrame(scenario26_2050.reset_index().values, columns=["Policy26_2050", "freq26_2050_policy"])
a12 = pd.DataFrame(policy26_2050.reset_index().values, columns=["scenario26_2050", "freq26_2050_scenario"])
a13 = pd.DataFrame(tippingPoint26_All_2070.reset_index().values, columns=["freq26_tip_all_2070", "tippingPoint26_all_2070"])
a14 = pd.DataFrame(tippingPoint26_Accepted_2070.reset_index().values, columns=["freq26_tipacc_2070", "tippingPoint26_acc_2070"])
a15 = pd.DataFrame(scenario26_2070.reset_index().values, columns=["Policy26_2070", "freq26_2070_policy"])
a16 = pd.DataFrame(policy26_2070.reset_index().values, columns=["scenario26_2070", "freq26_2070_scenario"])
b1= pd.DataFrame(tippingPoint45_All_1980.reset_index().values, columns=["freq45_tip_all_1980", "tippingPoint45_all_1980"])
b2= pd.DataFrame(tippingPoint45_Accepted_1980.reset_index().values, columns=["freq45_tipacc_1980", "tippingPoint45_acc_1980"])
b3= pd.DataFrame(scenario45_1980.reset_index().values, columns=["Policy45_1980", "freq45_1980_policy"])
b4= pd.DataFrame(policy45_1980.reset_index().values, columns=["scenario45_1980", "freq45_1980_scenario"])
b5 = pd.DataFrame(tippingPoint45_All_2020.reset_index().values, columns=["freq45_tip_all_2020", "tippingPoint45_all_2020"])
b6 = pd.DataFrame(tippingPoint45_Accepted_2020.reset_index().values, columns=["freq45_tipacc_2020", "tippingPoint45_acc_2020"])
b7 = pd.DataFrame(scenario45_2020.reset_index().values, columns=["Policy45_2020", "freq45_2020_policy"])
b8 = pd.DataFrame(policy45_2020.reset_index().values, columns=["scenario45_2020", "freq45_2020_scenario"])
b9 = pd.DataFrame(tippingPoint45_All_2050.reset_index().values, columns=["freq45_tip_all_2050", "tippingPoint45_all_2050"])
b10 = pd.DataFrame(tippingPoint45_Accepted_2050.reset_index().values, columns=["freq45_tipacc_2050", "tippingPoint45_acc_2050"])
b11 = pd.DataFrame(scenario45_2050.reset_index().values, columns=["Policy45_2050", "freq45_2050_policy"])
b12 = pd.DataFrame(policy45_2050.reset_index().values, columns=["scenario45_2050", "freq45_2050_scenario"])
b13 = pd.DataFrame(tippingPoint45_All_2070.reset_index().values, columns=["freq45_tip_all_2070", "tippingPoint45_all_2070"])
b14 = pd.DataFrame(tippingPoint45_Accepted_2070.reset_index().values, columns=["freq45_tipacc_2070", "tippingPoint45_acc_2070"])
b15 = pd.DataFrame(scenario45_2070.reset_index().values, columns=["Policy45_2070", "freq45_2070_policy"])
b16 = pd.DataFrame(policy45_2070.reset_index().values, columns=["scenario45_2070", "freq45_2070_scenario"])
c1= pd.DataFrame(tippingPoint45_All_1980.reset_index().values, columns=["freq85_tip_all_1980", "tippingPoint85_all_1980"])
c2= pd.DataFrame(tippingPoint45_Accepted_1980.reset_index().values, columns=["freq85_tipacc_1980", "tippingPoint85_acc_1980"])
c3= pd.DataFrame(scenario45_1980.reset_index().values, columns=["Policy85_1980", "freq85_1980_policy"])
c4= pd.DataFrame(policy45_1980.reset_index().values, columns=["scenario85_1980", "freq85_1980_scenario"])
c5 = pd.DataFrame(tippingPoint45_All_2020.reset_index().values, columns=["freq85_tip_all_2020", "tippingPoint85_all_2020"])
c6 = pd.DataFrame(tippingPoint45_Accepted_2020.reset_index().values, columns=["freq85_tipacc_2020", "tippingPoint85_acc_2020"])
c7 = pd.DataFrame(scenario45_2020.reset_index().values, columns=["Policy85_2020", "freq85_2020_policy"])
c8 = pd.DataFrame(policy45_2020.reset_index().values, columns=["scenario85_2020", "freq85_2020_scenario"])
c9 = pd.DataFrame(tippingPoint45_All_2050.reset_index().values, columns=["freq85_tip_all_2050", "tippingPoint85_all_2050"])
c10 = pd.DataFrame(tippingPoint45_Accepted_2050.reset_index().values, columns=["freq85_tipacc_2050", "tippingPoint85_acc_2050"])
c11 = pd.DataFrame(scenario45_2050.reset_index().values, columns=["Policy85_2050", "freq85_2050_policy"])
c12 = pd.DataFrame(policy45_2050.reset_index().values, columns=["scenario85_2050", "freq85_2050_scenario"])
c13 = pd.DataFrame(tippingPoint85_All_2070.reset_index().values, columns=["freq85_tip_all_2070", "tippingPoint85_all_2070"])
c14 = pd.DataFrame(tippingPoint85_Accepted_2070.reset_index().values, columns=["freq85_tipacc_2070", "tippingPoint85_acc_2070"])
c15 = pd.DataFrame(scenario85_2070.reset_index().values, columns=["Policy85_2070", "freq85_2070_policy"])
c16 = pd.DataFrame(policy85_2070.reset_index().values, columns=["scenario85_2070", "freq85_2070_scenario"])
# -
df_final_tipping_point_1980_2070 = pd.concat((a1, a2, a3, a4, a5, a6, a7, a8, a9, a10, a11, a12, a13, a14, a15, a16,
b1, b2, b3, b4, b5, b6, b7, b8, b9, b10, b11, b12, b13, b14, b15, b16,
c1, c2, c3, c4, c5, c6, c7, c8, c9, c10, c11, c12, c13, c14, c15, c16), axis = 1)
df_final_tipping_point_1980_2070.to_csv(os.path.join(rootOut, 'df_final_tipping_point_1980_2070.csv'), index = False)
df_final_tipping_point_1980_2070.shape
df_final_tipping_point_1980_2070.head(67)
# +
adpation_Option1 = np.array(df_policies['x1SnowThershold'])
adpation_Option2 = np.array(df_policies['xGoodDays'])
all_Policies = []
for i in range(len(adpation_Option1)):
all_Policies.append('P' + '_' + str(int(adpation_Option1[i])) + '_' + str(int(adpation_Option2[i])))
# +
#all_Policies
# +
#df_final_tipping_point_1980_2070['Policy26_2020'] = df_final_tipping_point_1980_2070['Policy26_2020'].apply(lambda x: 'Plc_' + str(x))
# +
#df_final_tipping_point_1980_2070['Policy26_2050'] = df_final_tipping_point_1980_2070['Policy26_2050'].apply(lambda x: 'Plc_' + str(x))
# +
#df_final_tipping_point_1980_2070['Policy26_2070'] = df_final_tipping_point_1980_2070['Policy26_2070'].apply(lambda x: 'Plc_' + str(x))
# -
x_26 = df_final_tipping_point_1980_2070.loc[0:44, 'Policy26_2020'].to_list()
y_26 = df_final_tipping_point_1980_2070.loc[0:44, 'freq26_2020_policy'].to_list()
x1_26 = df_final_tipping_point_1980_2070.loc[0:44, 'Policy26_2050'].to_list()
y1_26 = df_final_tipping_point_1980_2070.loc[0:44, 'freq26_2050_policy'].to_list()
x2_26 = df_final_tipping_point_1980_2070.loc[0:44, 'Policy26_2070'].to_list()
y2_26 = df_final_tipping_point_1980_2070.loc[0:44, 'freq26_2070_policy'].to_list()
x_45 = df_final_tipping_point_1980_2070.loc[0:44, 'Policy45_2020'].to_list()
y_45 = df_final_tipping_point_1980_2070.loc[0:44, 'freq45_2020_policy'].to_list()
x1_45 = df_final_tipping_point_1980_2070.loc[0:41, 'Policy45_2050'].to_list()
y1_45 = df_final_tipping_point_1980_2070.loc[0:41, 'freq45_2050_policy'].to_list()
x2_45 = df_final_tipping_point_1980_2070.loc[0:41, 'Policy45_2070'].to_list()
y2_45 = df_final_tipping_point_1980_2070.loc[0:41, 'freq45_2070_policy'].to_list()
x_85 = df_final_tipping_point_1980_2070.loc[0:44, 'Policy85_2020'].to_list()
y_85 = df_final_tipping_point_1980_2070.loc[0:44, 'freq85_2020_policy'].to_list()
x1_85 = df_final_tipping_point_1980_2070.loc[0:41, 'Policy85_2050'].to_list()
y1_85 = df_final_tipping_point_1980_2070.loc[0:41, 'freq85_2050_policy'].to_list()
x2_85 = df_final_tipping_point_1980_2070.loc[0:3, 'Policy85_2070'].to_list()
y2_85 = df_final_tipping_point_1980_2070.loc[0:3, 'freq85_2070_policy'].to_list()
x2_85
y2_85
# +
#y.reverse()
# -
x_26_arr = np.array(x_26) - 0.25
x1_26_arr = np.array(x1_26)
x2_26_arr = np.array(x2_26) + 0.25
x_26_arr
title_Figs = 'case3_Hoch-Ybrig (1050-1820m)'
# +
fig35, ax1 = plt.subplots(figsize=(20,7.5))
width = 0.25
ax1.bar(x_26_arr, y_26, width = width, color = 'Blue', label = "2020")
ax1.bar(x1_26_arr, y1_26, width = width, color = 'Green', label = "2050")
ax1.bar(x2_26_arr, y2_26, width = width, color = 'Red', label = "2070")
#X-Axis
xticks = np.arange(0, 45, 1)
ax1.set_xticks(xticks)
#ax1.set_xticks(xticks, all_Policies)
xlabels = all_Policies
#ax1.set_xticklabels(xlabels)
#plt.setp(ax1.get_xticklabels(), rotation=60, size = 10, ha="right", rotation_mode="anchor")
#plt.setp(ax1.get_xticklabels(), rotation=60, size = 10)
plt.xticks(xticks, xlabels, fontsize=16)
plt.setp(ax1.get_xticklabels(), rotation=90, size = 10)
#Y-Axis
yticks = np.arange(0, 70, 3)
#ax1.set_yticks(yticks)
#plt.setp(ax1.get_yticklabels(), rotation=0, size = 15, ha="right", rotation_mode="anchor")
ax1.set_yticks(yticks)
ax1.set_title(title_Figs + ', RCP2.6', size = 30)
ax1.set_xlabel('45 Adaptation Options', size = 20)
ax1.set_ylabel('Frequency', size = 20)
ax1.axhline(y=67, color='green', alpha=0.8)
ax1.axhline(y=50, color='green', alpha=0.8)
ax1.axhline(y=33, color='orange', alpha=0.8)
ax1.axhline(y=16, color='red', alpha=0.8)
#ax1.set_ylim(bottom=0, top =70)
#ax1.y_axis = np.arange(0, 70)
fig35.savefig(os.path.join(rootOut, 'tipping_point_All_new_3_RCP26.tiff'), format='tiff', dpi=150)
# -
x_45_arr = np.array(x_45) - 0.25
x1_45_arr = np.array(x1_45)
x2_45_arr = np.array(x2_45) + 0.25
y2_45
# +
fig36, ax1 = plt.subplots(figsize=(20,7.5))
width = 0.25
ax1.bar(x_45_arr, y_45, width = width, color = 'Blue', label = "2020")
ax1.bar(x1_45_arr, y1_45, width = width, color = 'Green', label = "2050")
ax1.bar(x2_45_arr, y2_45, width = width, color = 'Red', label = "2070")
#X-Axis
xticks = np.arange(0, 45, 1)
ax1.set_xticks(xticks)
#ax1.set_xticks(xticks, all_Policies)
xlabels = all_Policies
#ax1.set_xticklabels(xlabels)
#plt.setp(ax1.get_xticklabels(), rotation=60, size = 10, ha="right", rotation_mode="anchor")
#plt.setp(ax1.get_xticklabels(), rotation=60, size = 10)
plt.xticks(xticks, xlabels, fontsize=16)
plt.setp(ax1.get_xticklabels(), rotation=90, size = 10)
#Y-Axis
yticks = np.arange(0, 70, 3)
#ax1.set_yticks(yticks)
#plt.setp(ax1.get_yticklabels(), rotation=0, size = 15, ha="right", rotation_mode="anchor")
ax1.set_yticks(yticks)
ax1.set_title(title_Figs + ', RCP4.5', size = 30)
ax1.set_xlabel('45 Adaptation Options', size = 20)
ax1.set_ylabel('Frequency', size = 20)
ax1.axhline(y=67, color='green', alpha=0.8)
ax1.axhline(y=50, color='green', alpha=0.8)
ax1.axhline(y=33, color='orange', alpha=0.8)
ax1.axhline(y=16, color='red', alpha=0.8)
#ax1.set_ylim(bottom=0, top =70)
#ax1.y_axis = np.arange(0, 70)
fig36.savefig(os.path.join(rootOut, 'tipping_point_All_new_3_RCP45.tiff'), format='tiff', dpi=150)
# -
x_85_arr = np.array(x_85) - 0.25
x1_85_arr = np.array(x1_85)
x2_85_arr = np.array(x2_85) + 0.25
y2_85
x2_85_arr
# +
fig37, ax1 = plt.subplots(figsize=(20,7.5))
width = 0.25
ax1.bar(x_85_arr, y_85, width = width, color = 'Blue', label = "2020")
ax1.bar(x1_85_arr, y1_85, width = width, color = 'Green', label = "2050")
ax1.bar(x2_85_arr, y2_85, width = width, color = 'Red', label = "2070")
#X-Axis
xticks = np.arange(0, 45, 1)
ax1.set_xticks(xticks)
#ax1.set_xticks(xticks, all_Policies)
xlabels = all_Policies
#ax1.set_xticklabels(xlabels)
#plt.setp(ax1.get_xticklabels(), rotation=60, size = 10, ha="right", rotation_mode="anchor")
#plt.setp(ax1.get_xticklabels(), rotation=60, size = 10)
plt.xticks(xticks, xlabels, fontsize=16)
plt.setp(ax1.get_xticklabels(), rotation=90, size = 10)
#Y-Axis
yticks = np.arange(0, 70, 3)
#ax1.set_yticks(yticks)
#plt.setp(ax1.get_yticklabels(), rotation=0, size = 15, ha="right", rotation_mode="anchor")
ax1.set_yticks(yticks)
ax1.set_title(title_Figs + ', RCP8.5', size = 30)
ax1.set_xlabel('45 Adaptation Options', size = 20)
ax1.set_ylabel('Frequency', size = 20)
ax1.axhline(y=67, color='green', alpha=0.8)
ax1.axhline(y=50, color='green', alpha=0.8)
ax1.axhline(y=33, color='orange', alpha=0.8)
ax1.axhline(y=16, color='red', alpha=0.8)
#ax1.set_ylim(bottom=0, top =70)
#ax1.y_axis = np.arange(0, 70)
fig37.savefig(os.path.join(rootOut, 'tipping_point_All_new_3_RCP85.tiff'), format='tiff', dpi=150)
# -
|
tipping_points_case3_hoch-ybrig.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Design of a Cold Weather Fuel
# The venerable alcohol stove has been invaluable camping accessory for generations. They are simple, reliable, and in a pinch, can be made from aluminum soda cans.
#
# 
#
# Alcohol stoves are typically fueled with denatured alcohol. Denatured alcohol, sometimes called methylated spirits, is a generally a mixture of ethanol and other alcohols and compounds designed to make it unfit for human consumption. An MSDS description of one [manufacturer's product](https://www.korellis.com/wordpress/wp-content/uploads/2016/05/Alcohol-Denatured.pdf) describes a roughly fifity/fifty mixture of ethanol and methanol.
#
# The problem with alcohol stoves is they can be difficult to light in below freezing weather. The purpose of this notebook is to design of an alternative cold weather fuel that could be mixed from other materials commonly available from hardware or home improvement stores.
# ## Data
#
# The following data was collected for potential fuels commonly available at hardware and home improvement stores. The data consists of price (\$/gal.) and parameters to predict vapor pressure using the Antoine equation,
#
# \begin{align}
# \log_{10}P^{vap}_{s}(T) & = A_s - \frac{B_s}{T + C_s}
# \end{align}
#
# where the subscript $s$ refers to species, temperature $T$ is in units of degrees Celcius, and pressure $P$ is in units of mmHg. The additional information for molecular weight and specific gravity will be needed to present the final results in volume fraction.
data = {
'ethanol' : {'MW': 46.07, 'SG': 0.791, 'A': 8.04494, 'B': 1554.3, 'C': 222.65},
'methanol' : {'MW': 32.04, 'SG': 0.791, 'A': 7.89750, 'B': 1474.08, 'C': 229.13},
'isopropyl alcohol': {'MW': 60.10, 'SG': 0.785, 'A': 8.11778, 'B': 1580.92, 'C': 219.61},
'acetone' : {'MW': 58.08, 'SG': 0.787, 'A': 7.02447, 'B': 1161.0, 'C': 224.0},
'xylene' : {'MW': 106.16, 'SG': 0.870, 'A': 6.99052, 'B': 1453.43, 'C': 215.31},
'toluene' : {'MW': 92.14, 'SG': 0.865, 'A': 6.95464, 'B': 1344.8, 'C': 219.48},
}
# ## Denatured Alcohol
#
# The first step is to determine the vapor pressure of denatured alcohol over a typical range of operating temperatures. For this we assume denatured alcohol is a 40/60 (mole fraction) mixture of ethanol and methanol.
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
def Pvap(T, s):
return 10**(data[s]['A'] - data[s]['B']/(T + data[s]['C']))
def Pvap_denatured(T):
return 0.4*Pvap(T, 'ethanol') + 0.6*Pvap(T, 'methanol')
T = np.linspace(0, 40, 200)
plt.plot(T, Pvap_denatured(T))
plt.title('Vapor Pressure of denatured alcohol')
plt.xlabel('temperature / °C')
plt.ylabel('pressure / mmHg')
plt.grid()
print("Vapor Pressure at 0C =", round(Pvap_denatured(0),1), "mmHg")
# -
# ## Cold Weather Product Requirements
#
# We seek a cold weather fuel with increased vapor pressure at 0°C and lower, and also provides safe and normal operation of the alcohol stove at higher operating temperatures.
#
# For this purpose, we seek a mixture of commonly available liquids with a vapor pressure of at least 22 mmHg at the lowest possible temperature, and no greater than the vapor pressure of denatured alcohol at temperatures 30°C and above.
for s in data.keys():
plt.plot(T, Pvap(T,s))
plt.plot(T, Pvap_denatured(T), 'k', lw=3)
plt.legend(list(data.keys()) + ['denatured alcohol'])
plt.title('Vapor Pressure of selected compounds')
plt.xlabel('temperature / °C')
plt.ylabel('pressure / mmHg')
plt.grid()
# ## Optimization Model
#
# The first optimization model is to create a mixture that maximizes the vapor pressure at -10°C while having a vapor pressure less than or equal to denatured alcohol at 30°C and above.
#
# The decision variables in the optimization model correspond to $x_s$, the mole fraction of each species $s \in S$ from the set of available species $S$. By definition, the mole fractions must satisfy
#
# \begin{align}
# x_s & \geq 0 & \forall s\in S \\
# \sum_{s\in S} x_s & = 1
# \end{align}
#
# The objective is to maximize the vapor pressure at low temperatures, say -10°C, while maintaing a vapor pressure less than or equal to denatured alcohol at 30°C. Using Raoult's law for ideal mixtures,
#
# \begin{align}
# \max_{x_s} \sum_{s\in S} x_s P^{vap}_s(-10°C) \\
# \end{align}
# subject to
# \begin{align}
# \sum_{s\in S} x_s P^{vap}_s(30°C) & \leq P^{vap}_{denatured\ alcohol}(30°C) \\
# \end{align}
#
# This optimization model is implemented in Pyomo in the following cell.
# +
import pyomo.environ as pyomo
m = pyomo.ConcreteModel()
S = data.keys()
m.x = pyomo.Var(S, domain=pyomo.NonNegativeReals)
def Pmix(T):
return sum(m.x[s]*Pvap(T,s) for s in S)
m.obj = pyomo.Objective(expr = Pmix(-10), sense=pyomo.maximize)
m.cons = pyomo.ConstraintList()
m.cons.add(sum(m.x[s] for s in S)==1)
m.cons.add(Pmix(30) <= Pvap_denatured(30))
m.cons.add(Pmix(40) <= Pvap_denatured(40))
solver = pyomo.SolverFactory('glpk')
solver.solve(m)
print("Vapor Pressure at -10°C =", m.obj(), "mmHg")
T = np.linspace(-10,40,200)
plt.plot(T, Pvap_denatured(T), 'k', lw=3)
plt.plot(T, [Pmix(T)() for T in T], 'r', lw=3)
plt.legend(['denatured alcohol'] + ['cold weather blend'])
plt.title('Vapor Pressure of selected compounds')
plt.xlabel('temperature / °C')
plt.ylabel('pressure / mmHg')
plt.grid()
# -
# ## Display Composition
# +
import pandas as pd
s = data.keys()
results = pd.DataFrame.from_dict(data).T
for s in S:
results.loc[s,'mole fraction'] = m.x[s]()
MW = sum(m.x[s]()*data[s]['MW'] for s in S)
for s in S:
results.loc[s,'mass fraction'] = m.x[s]()*data[s]['MW']/MW
vol = sum(m.x[s]()*data[s]['MW']/data[s]['SG'] for s in S)
for s in S:
results.loc[s,'vol fraction'] = m.x[s]()*data[s]['MW']/data[s]['SG']/vol
results
# -
|
Mathematics/Mathematical Modeling/06.07-Design-of-a-Cold-Weather-Fuel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
from collections import defaultdict
import numpy as np
total_output = """timestamps LikelihoodGradientJob::update_workers_state: 112610684029167 112610701565912
timestamps LikelihoodGradientJob::receive_results_on_master: 112644816902697 112644859488707
timestamps LikelihoodGradientJob::update_workers_state: 112645713883737 112645753127370
timestamps LikelihoodGradientJob::receive_results_on_master: 112665210540385 112665250951149
timestamps LikelihoodGradientJob::update_workers_state: 112665812374455 112665831142249
timestamps LikelihoodGradientJob::receive_results_on_master: 112685004710061 112685042886471
timestamps LikelihoodGradientJob::update_workers_state: 112685532735260 112685545556799
timestamps LikelihoodGradientJob::receive_results_on_master: 112702476907290 112702524958786
MnSeedGenerator: Negative G2 found - new state: - FCN = 440.6948853451 Edm = 761.327 NCalls = 24
VariableMetric: start iterating until Edm is < 1000
VariableMetric: Initial state - FCN = 440.6948853451 Edm = 761.327 NCalls = 24
VariableMetric: Iteration # 0 - FCN = 440.6948853451 Edm = 761.327 NCalls = 24
timestamps LikelihoodGradientJob::update_workers_state: 112702757048781 112702775713451
timestamps LikelihoodGradientJob::receive_results_on_master: 112719266431520 112719300644175
VariableMetric: Iteration # 1 - FCN = 334.9853991888 Edm = 664.469 NCalls = 26
timestamps LikelihoodGradientJob::update_workers_state: 112719423603118 112719438922361
timestamps LikelihoodGradientJob::receive_results_on_master: 112737191127063 112737225931912
"""
timestamps_lines = (x.split(': ') for x in (x for x in total_output.splitlines() if 'timestamps' in x))
timestamps = defaultdict(list)
times = defaultdict(list)
for x, y in timestamps_lines:
key = " ".join(x.split()[1:])
stamps = [int(stamp) for stamp in y.split()]
timestamps[key].append(stamps)
times[key].append(np.diff(stamps))
timestamps
for key, task_times in times.items():
print(key)
print('average:', np.mean(task_times) / 1e9, 'seconds')
print('total: ', np.sum(task_times) / 1e9, 'seconds')
|
20210410_analyze_performance_timestamps/kladblok timestamps verwerken.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis of Models using only MIMIC Notes
# ## Imports & Inits
# +
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append('../')
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("darkgrid")
# %matplotlib inline
import pickle
import numpy as np
import pandas as pd
from sklearn.metrics import mean_squared_error
from pathlib import Path
from utils.plots import *
# -
from args import args
vars(args)
# +
subsets = ['s', 'u', 'u+s']
rmsle = {}
for subset in subsets:
rmsle[subset] = []
with open(args.workdir/f'{subset}_preds.pkl', 'rb') as f:
targs = pickle.load(f)
preds = pickle.load(f)
for targ, pred in zip(targs, preds):
rmsle[subset].append(np.sqrt(mean_squared_error(pred, targ)))
# -
save = True
# +
df = pd.DataFrame.from_dict(rmsle)
df.columns = ['Structured (S)', 'Unstructured (U)', 'Multimodal (U+S)']
means = [np.round(value.mean(), 3) for colname, value in df.iteritems()]
df = pd.melt(df)
df.columns = ['', 'RMSLE']
# +
fig, ax = plt.subplots(1, 1, figsize=(11, 8))
g = sns.pointplot(x='', y='RMSLE', data=df, ax=ax, estimator=np.mean)
ax.set_ylabel('Average RMSLE')
[ax.text(p[0], p[1]+0.002, p[1], color='r') for p in zip(ax.get_xticks(), means)]
if save:
fig.savefig(args.figdir/'rmsle_pointplot.pdf', dpi=300, bbox_inches='tight', pad_inches=0)
# -
# ## Box Plot
plot_dfs = []
# +
prefix,cohort = 'full_common_vital','Notes & Vitals (U+S)'
bams = pickle.load(open(workdir/f'{prefix}_bams.pkl', 'rb'))
final_metrics = pd.read_csv(workdir/f'{prefix}_metrics.csv', index_col=0)
best_models = pd.read_csv(workdir/f'{prefix}_best_models.csv', index_col=0)
ttests = pd.read_csv(workdir/f'{prefix}_ttests.csv', index_col=0)
for k in bams.keys():
bams[k.upper()] = bams.pop(k)
bams['AVG-ALL'] = bams.pop('AVG-LR-RF-GBM')
bams['MAX-ALL'] = bams.pop('MAX-LR-RF-GBM')
itr = iter(bams.keys())
bams.keys()
metrics = {}
for md in itr:
df = pd.DataFrame()
for k, m in bams[md].yield_metrics():
df[k] = m
df['Model'] = md
cols = list(df.columns)
cols = [cols[-1]] + cols[:-1]
df = df[cols]
metrics[md] = df
plot_df = pd.concat(metrics.values())
plot_df['Cohort'] = cohort
plot_dfs.append(plot_df)
# -
plot_df = pd.concat(plot_dfs)
plot_df[['Sensitivity', 'Specificity', 'PPV', 'AUC']] = plot_df[['Sensitivity', 'Specificity', 'PPV', 'AUC']] * 100
plot_df.shape
# +
met = 'Sensitivity'
fig, ax = plt.subplots(1,1,figsize=(20,10))
sns.boxplot(x='Model', y=met, hue='Cohort', data=plot_df, ax=ax)
# for i in range(10): plt.axvline(x=i+0.5, ls='-.', color='black')
ax.set_xlabel('')
# -
save = True
if save:
fig.savefig(figdir/f'nxv_{met.lower()}_box_plot.pdf', dpi=300)
# ## Mean AUC
def get_mean_tprs(bams, base_fpr):
mean_tprs = {}
for model, bam in bams.items():
tprs = []
for i, (targs, probs) in enumerate(zip(bam.targs, bam.pos_probs)):
fpr, tpr, _ = roc_curve(targs, probs)
tpr = interp(base_fpr, fpr, tpr)
tpr[0] = 0.0
tprs.append(tpr)
tprs = np.array(tprs)
mean_tprs[model] = tprs.mean(axis=0)
return mean_tprs
save = True
# +
prefix = 'full_common_all'
bams = pickle.load(open(workdir/f'{prefix}_bams.pkl', 'rb'))
final_metrics = pd.read_csv(workdir/f'{prefix}_metrics.csv', index_col=0)
best_models = pd.read_csv(workdir/f'{prefix}_best_models.csv', index_col=0)
ttests = pd.read_csv(workdir/f'{prefix}_ttests.csv', index_col=0)
for k in bams.keys():
bams[k.upper()] = bams.pop(k)
bams['AVG-ALL'] = bams.pop('AVG-LR-RF-GBM')
bams['MAX-ALL'] = bams.pop('MAX-LR-RF-GBM')
# +
des = 'all_'
if not des:
plot_bams = {k: bams[k] for k in bams.keys() if '-' not in k}
des = ''
names = plot_bams.keys()
aucs = [model.auroc_avg() for _, model in plot_bams.items()]
legends = [f'{model} ({auc})' for model, auc in zip(names, aucs)]
elif des == 'avg_':
plot_bams = {k: bams[k] for k in bams.keys() if 'avg' in k}
names = [name[4:] for name in plot_bams.keys()]
aucs = [model.auroc_avg() for _, model in plot_bams.items()]
legends = [f'{model} ({auc})' for model, auc in zip(names, aucs)]
elif des == 'max_':
plot_bams = {k: bams[k] for k in bams.keys() if 'max' in k}
names = [name[4:] for name in plot_bams.keys()]
aucs = [model.auroc_avg() for _, model in plot_bams.items()]
legends = [f'{model} ({auc})' for model, auc in zip(names, aucs)]
elif des == 'all_':
plot_bams = bams
names = plot_bams.keys()
aucs = [model.auroc_avg() for _, model in plot_bams.items()]
legends = [f'{model} ({auc})' for model, auc in zip(names, aucs)]
legends
# +
base_fpr = np.linspace(0, 1, 100)
mean_tprs = get_mean_tprs(plot_bams, base_fpr)
fig, ax = plt.subplots(1, 1, figsize=(11, 8))
for i, (model, mean_tpr) in enumerate(mean_tprs.items()):
ax.plot(base_fpr, mean_tpr)
ax.plot([0, 1], [0, 1], linestyle=':')
ax.grid(b=True, which='major', color='#d3d3d3', linewidth=1.0)
ax.grid(b=True, which='minor', color='#d3d3d3', linewidth=0.5)
ax.set_ylabel('Sensitivity')
ax.set_xlabel('1 - Specificity')
ax.legend(legends)
if save:
fig.savefig(figdir/f'{prefix}_{des}mean_auc.pdf', dpi=300)
|
mercari/results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
# # I. Overview of Machine Learning
# ## Scenario
# In this module, we'll build a classifier to predict whether or not a patient has diabetes.
#
# Let's say that you're assigned the task of determining whether a set of 100 patients has diabetes. You don't get to meet with the patient or run any tests. All that you're given is some information that was collected for each of them, such as how many pregnancies they've had and what their current glucose levels are.
#
# **Option 1**
#
# If you're a clinician and an expert in diabetes, this information might be enough for you to make your decisions. You might follow existing guidelines, such as these provided by the [National Diabetes Education Initiative](http://www.ndei.org/ADA-diabetes-management-guidelines-diagnosis-A1C-testing.aspx.html). For each patient, you consult the guidelines and make a decision of whether they **have diabetes** or **do not have diabetes**.
#
# **Option 2**
#
# However, maybe the information that you've been provided isn't detailed or relevant enough. Or maybe you don't have the medical knowledge to make such a complex decision. In that case, an alternative approach could be to compare the patients you've been asked to classify to other similar patients who have already been diagnosed. You ask your data warehouse managers to retrieve the 8 columns of information for 900 other patients in the same population, plus whether or not they had diabetes. Now, you can see what kind of patterns occur in patients who have diabetes and those who do not, and you can use those patterns to make decisions about whether these new patients have diabetes.
#
# Machine learning uses this second option to make decisions. A **classifier** is an algorithm that takes data as input and learns how to make decisions based on that data. In our case, our classifier will decide whether a patient has diabetes.
#
# ## Definitions
# - **Task** - what we want our classifier to do. We want our classifier to predict if a patient is **positive** or **negative** for diabetes
# - **Model/Classifier** - this is the algorithm that we will use to make predictions
# - **Training Data** - the data that we provide our algorithm to learn patterns. In our scenario, the training data is the information for the 900 patients who have already been diagnosed with diabetes or been determined to not have diabetes
# - **Features** - the information that is collected for each patient in the dataset, such as number of pregnancies and glucose levels
# - **Label/Outcome** - a classification for each patient in the training data. For example, if a patient has a diabetes, then we might put a "1" in the outcome column, and if they don't we might put a "0"
# - **Training** - how our model learns to make predictions
# - **Evaluation** - once
# # Our Dataset
# We will use the [Pima Indians Diabetes Dataset](https://www.kaggle.com/uciml/pima-indians-diabetes-database/home), which can be downloaded from Kaggle. This dataset was originally created by the National Institute of Diabetes and Digestive and Kidney Diseases and contains data for a number of patients. Each patient is a female at least 21 years old of [Pima Indian heritage](https://en.wikipedia.org/wiki/Pima_people).
#
# ## What's in the Data?
# Let's take a look at our dataset. We'll read in the data from a comma-separated file and look at it as a table where each row represents a different patient:
df = pd.read_csv('diabetes.csv')
df.head()
# ### Features
# The **"features"** in a dataset are the information collected for each data point. In this scenario, the features are the 8 types of information collected for each patient.
#
# Take a few minutes with your group and look through some of the features. Try to get a sense for what each attribute is measuring. Optionally, do some programmatic analysis to look at the mean, standard deviations, etc.
#
# - **Pregnancies**: Number of times pregnant
# - **Glucose**: Plasma glucose concentration a 2 hours in an oral glucose tolerance test
# - **BloodPressure**: Diastolic blood pressure (mm Hg)
# - **SkinThickness**: Triceps skin fold thickness (mm)
# - **Insulin**: 2-Hour serum insulin (mu U/ml)
# - **BMI**: Body mass index (weight in kg/(height in m)^2)
# - **DiabetesPedigreeFunction**: Diabetes pedigree function which considers family history of diabetes
# - **Age**: Age (years)
# Optional: Do additional analysis
df.Pregnancies.describe()
# ### Label
# The **label** signifies what **class** each row belongs to. A **"1"** means that the patient has diabetes (positive class), while a **"0"** means that the patient does not have diabetes (negative class). This is contained in the *Outcome* column.
df.Outcome.value_counts()
# # Up Next
# Next, we'll look more closely at our dataset and analyze our features and class distribution.
#
# [II. Data Analysis](./II_DataAnalysis.ipynb)
|
ai_etc/I_Overview.ipynb
|