code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://adventofcode.com/2018/day/
import numpy as np
with open("day_input.txt") as file:
puzzle_input = file.read().splitlines()
small_puzzle_input = 0
# ## Part 1
# ## Part 2
| Advent of Code 2018 - Day Template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Mattyughhh/CPEN-ECE-2-2/blob/main/Introduction_to_Python_Programming.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="9sfkL2STjVUj" outputId="a1121d50-e95d-48a0-e5fc-76be5d951664"
print ("Hello, World")
# + colab={"base_uri": "https://localhost:8080/"} id="kba_vKaFuUmo" outputId="893afa6f-b1ea-4bec-d50b-81249c56c5a7"
b = float (4)
print (round,((b),3))
# + colab={"base_uri": "https://localhost:8080/"} id="d9qrdw2XujDW" outputId="225ca97d-f103-42ab-e309-36f8bd7a48ae"
a = 4
A = "sally"
#A will not overwrite a
print (a)
print (A)
# + colab={"base_uri": "https://localhost:8080/"} id="eQwmv1sZxMs1" outputId="1ab569af-187b-46e9-8db0-dcb9e6c05bd0"
x, y ,z= "one", "two", "three"
print (z)
print (y)
print (x)
# + colab={"base_uri": "https://localhost:8080/"} id="KUrafFrLxaD6" outputId="700cabc5-a71b-4fb1-dc50-a7e2716903a0"
x=y=z="Four"
print (x)
print (y)
print (z)
# + colab={"base_uri": "https://localhost:8080/"} id="Afzpug9FtK2u" outputId="60c19f80-85fd-4692-8da6-84ad13efa03b"
x = "enjoying"
print ("Python Programming is "+ x)
# + colab={"base_uri": "https://localhost:8080/"} id="3AapJnIHxkzQ" outputId="950c7a32-6b77-42d0-e256-4aec33aa59a0"
x=5
y=3
sum=x+y
print (sum)
# + colab={"base_uri": "https://localhost:8080/"} id="PaZc9EPvvEpD" outputId="293806bb-1125-47ff-c11c-d85f8c11057f"
x = 5
y = 6
print (x+y)
# + colab={"base_uri": "https://localhost:8080/"} id="uOyGA0Y2vKcN" outputId="306d78ce-80dc-4ab9-bba3-ee81f52ae5e7"
a, b, c = 0, -1, 8
c%=3
print (c)
# + colab={"base_uri": "https://localhost:8080/"} id="QHk6Xr2sxxmX" outputId="c425a8ca-06ee-4002-c342-5bd1be94d4c0"
x=5
x<6 and x<10
# + colab={"base_uri": "https://localhost:8080/"} id="-uGpfI_JtyDO" outputId="b2010be2-3798-4e09-bd06-830f0b7cfa5f"
y = 6
z = 7
y is not z
| Introduction_to_Python_Programming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ##Notes: the approach taken in this baseline model is from this 2020 article
# ##by Lianne and Justin, thanks to them for sharing. They used
# ##ridge regression alpha = 0.001
#
# https://www.justintodata.com/improve-sports-betting-odds-guide-in-python/
#
#
# +
import pandas as pd
import numpy as np
from sklearn.metrics import accuracy_score, precision_score, recall_score, roc_auc_score, confusion_matrix
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score, f1_score
# +
##we will try the following models on the base-line data ... just win/loss and which teams
##note KNN or other clusters might be helpful group the teams in smart way ... but not now.
#models
##regression
from sklearn.linear_model import Ridge
from sklearn.ensemble import RandomForestRegressor
#classifiers (non-tree)
from sklearn.linear_model import RidgeClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
#tree-based classifiers
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import BaggingClassifier
from sklearn.ensemble import GradientBoostingClassifier
from xgboost import XGBClassifier
from xgboost import XGBRegressor
##regression models
lr = Ridge(alpha=0.001)
rfr = RandomForestRegressor(max_depth=3, random_state=0)
xgbr = XGBRegressor()
regr_models = [lr, rfr, xgbr]
##classifier models
lrc = RidgeClassifier()
gnb = GaussianNB()
lgr = LogisticRegression(random_state = 0)
svc = SVC()
#tree-based classifiers
rfc = RandomForestClassifier(max_depth=3, random_state=0)
bc = BaggingClassifier()
gbc = GradientBoostingClassifier()
xgbc = XGBClassifier( use_label_encoder=False, num_class = [0,1])
class_models= [lrc, gnb, lgr, svc, rfc, bc, gbc, xgbc]
# +
data = pd.read_csv("/Users/joejohns/data_bootcamp/GitHub/final_project_nhl_prediction/Data/Processed_Data/Approach_1 and_2_win_loss_and_cumul_1seas_Pisch_data/data_bet_stats_mp.csv")
data.drop(columns=[ 'Unnamed: 0'], inplace=True)
# +
data['won'] = data['won'].apply(int)
data_playoffs = data.loc[data['playoffGame'] == 1, :].copy() #set aside playoff games ... probably won't use them.
data= data.loc[data['playoffGame'] == 0, :].copy()
#sorted(data.columns)
# -
all_seasons = sorted(set(data['season']))
all_seasons
# +
def make_HA_data(X, season, list_var_names = None ):
X = X.loc[X['season'] == season, :].copy()
X_H = X.loc[X['HoA'] == 'home',:].copy()
X_A = X.loc[X['HoA'] == 'away',:].copy()
X_H['goal_difference'] = X_H['goalsFor'] - X_H['goalsAgainst'] ##note every thing is based in home data
X_H.reset_index(drop = True, inplace = True)
X_A.reset_index(drop = True, inplace = True)
df_visitor = pd.get_dummies(X_H['nhl_name'], dtype=np.int64)
df_home = pd.get_dummies(X_A['nhl_name'], dtype=np.int64)
df_model = df_home.sub(df_visitor)
df_model['date'] = X_H['date']
df_model['full_date'] = X_H['full_date']
df_model['game_id'] = X_H['game_id']
df_model['home_id'] = X_H['team_id']
df_model['away_id'] = X_A['team_id']
y = X_H.loc[:,['date', 'full_date','game_id', 'Open','goal_difference', 'won']].copy() ##these are from home team perspective; 'Open' is for betting
return (df_model, y)
# -
X_dic = {}
y_dic = {}
for sea in all_seasons:
X_dic[sea] = make_HA_data(data, sea)[0]
y_dic[sea] = make_HA_data(data, sea)[1]
# +
#this is for regressors predicting wins - losses, can use this to turn output into win prediction
def make_win(x):
if x <= 0:
return 0
if x >0:
return 1
v_make_win = np.vectorize(make_win)
#useage: v_make_win(y_pred)
# +
##naive method: train on first half of season, 600 games, test on second half of season
##with no further training
def naive_test_train_regr_models(model, cut_off = 600, regr = True):
all_seasons2 = [sea for sea in all_seasons if sea != 20122013]#2012 is shortened season
total_acc = 0
counter = 0
model_name = str(model)
print("results for ", model_name)
print(" ")
for sea in all_seasons2:
#set teh predictor variables, :-5 does the job, would be better
#and safer to name the columns explcitly ... but the columns are date
#and so on ... no leakage worries. OK for this base line
X = X_dic[sea].iloc[:, :-5].copy()
#select season, remove date, etc. select target y
if regr == True:
y = y_dic[sea].loc[:, 'goal_difference'].copy()
else:
y = y_dic[sea].loc[:, 'won'].copy()
#carry out naive train-test split
y_train = y[0: cut_off].copy()
y_test = y[cut_off :].copy()
X_train = X[0: cut_off].copy()
X_test = X[cut_off :].copy()
#train model, find predictions
model.fit(X_train, y_train)
y_pred = model.predict(X_test) #this is regression pred on Hg - Ag
y_pred_win = v_make_win(y_pred) #this is the pred of who wins HW =1, AW =0
y_test_win = v_make_win(y_test) #this gives the correct win, loss
#note: if y, y_pred and y_test are already 1, 0 then v_make_win will
#keep them the same (<= 0 --> 0, >0 ---> 1)
accuracy = accuracy_score(y_test_win, y_pred_win)
f1 = f1_score(y_test_win, y_pred_win) #, average = None)
counter+=1
total_acc+= accuracy
print("seaoson: ", sea)
print("acc: ", accuracy, " f1: ", f1)
avg_acc = total_acc/counter
print('avg acuracy: ', avg_acc)
print(" ")
#evaluate_regression(y_test, y_pred)
#evaluate_binary_classification(y_test_win, y_pred_win
# +
##try for ridge regression
naive_test_train_regr_models(model = lr, cut_off = 700, regr = True) ##ok looks like 20162017 is unusually good for some reason
# +
##avg is around 54% for ridge regression
# +
##now try all regressors
for model in regr_models:
naive_test_train_regr_models(model = model, cut_off = 700, regr = True)
# -
##now try all classifiers
for model in class_models:
naive_test_train_regr_models(model = model, cut_off = 700, regr = False)
# +
##conclusions: some of the average scores are around 55% and some of the top
##scores on a season are as high as 58, 59%
##next steps:
##1. tune the models
##2. look into partial_fit across the seaosn for appropriate models that have that
| Model 1_v6_clean_Aug_2021_version-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from sklearn.model_selection import train_test_split
from glob import glob
trainval = pd.read_csv('../data/caterpillar/train_set.csv')
test = pd.read_csv('../data/caterpillar/test_set.csv')
trainval_tube_assemblies = trainval['tube_assembly_id'].unique()
test_tube_assemblies = test['tube_assembly_id'].unique()
from sklearn.model_selection import train_test_split
train_tube_assemblies, val_tube_assemblies = train_test_split(
trainval_tube_assemblies, random_state=42
)
train = trainval[trainval.tube_assembly_id.isin(train_tube_assemblies)]
val = trainval[trainval.tube_assembly_id.isin(val_tube_assemblies)]
train.shape, val.shape, test.shape
# -
for path in glob('../data/caterpillar/*.csv'):
df = pd.read_csv(path)
shared_columns = set(df.columns) & set(train.columns)
if shared_columns:
print(path, df.shape)
print(df.columns.tolist(), '\n')
# +
def wrangle(data):
data = data.copy()
# Engineer date features
data['quote_date'] = pd.to_datetime(data['quote_date'], infer_datetime_format=True)
data['quote_date_year'] = data['quote_date'].dt.year
data['quote_date_month'] = data['quote_date'].dt.month
data = data.drop(columns='quote_date')
# Merge data
tube = pd.read_csv('../data/caterpillar/tube.csv')
bill_of_materials = pd.read_csv('../data/caterpillar/bill_of_materials.csv')
specs = pd.read_csv('../data/caterpillar/specs.csv')
data = data.merge(tube, how='left')
data = data.merge(bill_of_materials, how='left')
data = data.merge(specs, how='left')
# Drop tube_assembly_id because our goal is to predict unknown assemblies
data = data.drop(columns='tube_assembly_id')
return data
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
print(train.shape, val.shape, test.shape)
# +
from sklearn.metrics import mean_squared_error
def rmse(y_true, y_pred):
return np.sqrt(mean_squared_error(y_true, y_pred))
# +
import numpy as np
from sklearn.pipeline import make_pipeline
import category_encoders as ce
from xgboost import XGBRegressor
import matplotlib.pyplot as plt
target = 'cost'
X_train = train.drop(columns=target)
X_val = val.drop(columns=target)
y_train = train[target]
y_val = val[target]
y_train_log = np.log1p(y_train)
y_val_log = np.log1p(y_val)
pipeline = make_pipeline(
ce.OrdinalEncoder(),
XGBRegressor(max_depth=7,
n_estimators=1000,
n_jobs=-1,
gamma=.05,
colsample_bytree=.3,
reg_alpha=.1,
reg_lambda=.95,
objective='reg:squarederror')
)
pipeline.fit(X_train, np.array(y_train_log))
y_train_pred_log = pipeline.predict(X_train)
y_val_pred_log = pipeline.predict(X_val)
print(rmse(y_train_log, y_train_pred_log))
print(rmse(y_val_log, y_val_pred_log))
coefficients = pd.Series(pipeline[1].feature_importances_,
X_train.columns.tolist())
plt.style.use('dark_background')
plt.figure(figsize=(10,30))
coefficients.sort_values().plot.barh(color='grey');
plt.show()
# +
X_test = test.drop(columns='id')
y_pred_log = pipeline.predict(X_test)
y_pred = np.expm1(y_pred_log)
sample_submission = pd.read_csv('../data/caterpillar/sample_submission.csv')
submission = sample_submission.copy()
submission['cost'] = y_pred
submission.to_csv('nathan_van_wyck_submission.csv', index=False)
| module1-log-linear-regression/log_linear_regression_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This program finds the numerical solution using Euler's method of differential equation $\frac{dy}{dt} = f(t,y)$ with $y(t_0)=y_0$
# # <font color='red'> Euler's Method $y_{i+1} = y_i + f(t_i,y_i)(t_{i+1}-t_i), i = 0, 1,...,n-1$
# # <font color='green'> Example: $\frac{dy}{dt}=y, y(0)=1$. Find the solution on the time $0<t\le2$
# # Note its true solution is $y=exp(t)$
##f=lambda t,y,x:y**2+2*t+x
#f(5,1,2
from numpy import*
y=zeros(5)
y
# +
#import numpy as np
#import matplotlib.pyplot as plt
from matplotlib.pyplot import*
from numpy import*
f = lambda t,y: y #for matlab it looks like f=@(t,y)y
T = 2
n = 10
t = np.linspace(0,T,n)
y = np.zeros(n)
y[0] = 1
for i in range(0,n-1):
y[i+1] = y[i] + f(t[i],y[i])*(t[i+1]-t[i])
plot(t,y,'r')
grid()
import pandas as pd
data={}
data.update({"time":t, "displacement":y})
#print(data)
DF=pd.DataFrame(data)
print(DF)
DF.to_csv("euler.csv")
savefig("e.png")
# +
#import numpy as np
#import matplotlib.pyplot as plt
'''from matplotlib.pyplot import*
from numpy import*
f = lambda t,y:2*y #defining function, in matlab it is denoted by f=@(t,y)(y)
T = 2
n = 100
t = np.linspace(0,T,n)
y = np.zeros(n)
y[0] = 1
for i in range(0,n-1):
y[i+1] = y[i] + f(t[i],y[i])*(t[i+1]-t[i])
plot(t,y)
grid(True)
'''
import numpy as np
import matplotlib.pyplot as plt
#from matplotlib.pyplot import*
#from numpy import*
f = lambda t,y: y
T = 2
n = 50
t = np.linspace(0,T,n)
y = np.zeros(n)
y[0] = 1
true_soln=np.exp(t)
for i in range(0,n-1):
y[i+1] = y[i] + f(t[i],y[i])*(t[i+1]-t[i])
plt.plot(t,y,'o')
plt.plot(t,true_soln)
#plt.plot(t,np.exp(t))
plt.grid(True)
# -
# to see the data in table
import pandas as pd
data={} #making emptylist for data
data.update({"time":t,"yvaleu":y})
print(data) #here it print raw data
#to manage the data
DF=pd.DataFrame(data)
print(DF)
#import numpy as np
#import matplotlib.pyplot as plt
'''from matplotlib.pyplot import*
from numpy import*
f = lambda t,y: y
T = 2
n = 10
t = np.linspace(0,T,n)
y = np.zeros(n)
y[0] = 1
for i in range(0,n-1):
y[i+1] = y[i] + f(t[i],y[i])*(t[i+1]-t[i])
plot(t,y)
grid(True)
'''
# # <font color='green'>Find the numerical solution using Eular's method of differential equation
# # $\frac{dy}{dt}=f(t,y)$ with $y(t_0)=0$
# # Eulers formula:$ y_{i+1}=y_i+f(t_{i},y_i)(t_{i+1}- t_i)$
# # <font color='red'>Application:
# # Charging of capacitor $V = V_c +V_R$
# # 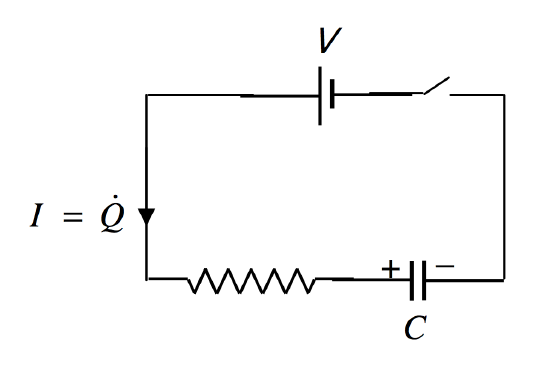
# # putting q=CV and $V_R=IR=\frac{dq}{dt}R$ we have $\frac{dq}{dt}=\frac{V}{R}-\frac{q}{RC}$,Given: q(0)=0(initial charge)
# #<font color='green'> true solution $ q=q_0(1-exp(-t/(RC)))$
# +
## now python coding
import numpy as np
import matplotlib.pyplot as plt
f=lambda t,q:(V/R)-(q/(R*C))
n=100
q = np.zeros(n)
q[0]=0
T= 5
R= 1
C= 1
V=5
qmax=C*V
t= np.linspace(0,T,n)
true_soln= qmax*(1-np.exp(-t/(R*C)))
for i in range(0,n-1):
q[i+1] = q[i]+f(t[i],q[i])*(t[i+1]-t[i])
# +
plt.plot(t,q)
plt.plot(t,true_soln,'or')
plt.grid(True)
plt.xlabel('time')
plt.ylabel('amount of charge')
plt.title('charging of capacitor')
# -
# To make the data table
import pandas as pd
data={}
data.update({"time":t,"charge":q})
#print (data)# this is raw data
DF=pd.DataFrame(data)
print(DF)
# +
## <font color='red'> Discharging of capacitor $V_C+V_R=0$
## put q=CV and $I= \frac{dq}{dt}$ , $\frac{dq}{dt}=-q/RC$ charge at t=0 is ,$q_0=CV$
## Euler's formula: $q_{i+1}=q_i +f(t[i],q[i])*(t[i+1]-t[i])$
##<font color='green'> true solution $q=q_0(exp(-t/(RC))$
# -
import numpy as np
import matplotlib.pyplot as plt
f=lambda t,q:-q/(R*C)
T=15
n=20
C=0.005
R=200
V=5
t=np.linspace(0,T,n)
q=np.zeros(n)
q[0]=C*V
true_soln=q[0]*(np.exp(-t/(R*C)))
for i in range(0,n-1):
q[i+1]=q[i]+f(t[i],q[i])*(t[i+1]-t[i])
plt.plot(t,q)
plt.plot(t,true_soln,'or')
plt.grid(True)
# +
import pandas as pd
data={}
data.update({"time":t,"charge":q})
print(data)
DF=pd.DataFrame(data)
print(DF)
# -
# # To make data table
# +
import pandas as pd
import numpy as np
n=11
x=np.linspace(0,20,n)
#y1=1/x
y2=x**2
y3=np.exp(x)
#y4=1/x**2
data={}
data.update({"value of x:":x,"reci.of x":y1,"square of x":y2,"exponential":y3,"reci of sqr":y4})
#print(data)
DF= pd.DataFrame(data)
print(DF)
# -
# # Solve first order diff eqn using Euler's method
# # Example: $ \frac{dy}{dt}+2y=2- \exp(-4t)$ ,$y_0=1$
# # making it into simpler form
# # $\frac{dy}{dt}=2-2y-exp(-4t)$ which is of form $\frac{dy}{dt}=f(t,y)$,$y_0=1$
# ## Euler's solution : $y_{i+1}=y_i+f(t_i,y_i)(t_{i+1}-t_{i})$
# +
# python code
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
n= 20
T= 5
t= np.linspace(0,T,n)
y= np.zeros(n)
y[0]=1
f= lambda t,y:2-2*y-np.exp(-4*t)
for i in range(0,n-1):
y[i+1]= y[i]+f(t[i],y[i])*(t[i+1]-t[i])
# -
plt.plot(t,y)
data={}
data.update({"time:":t, "function value y:":y})
DF=pd.DataFrame(data)
print(DF)
#
# # <font color='red'> Rise and decay of current in LR circuit
# # 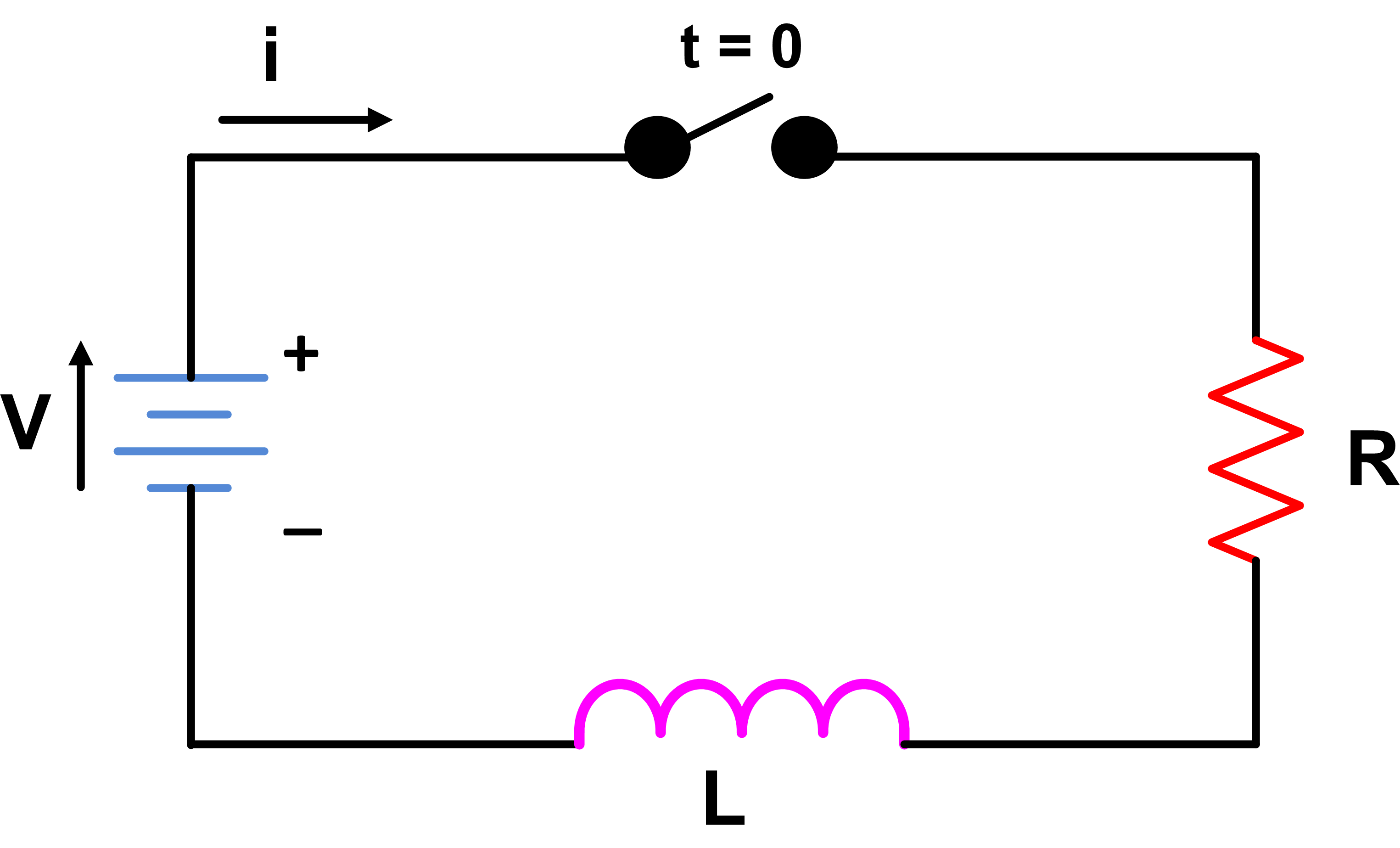
# # Growth of current:
# # When switch is on and applyinf kirchoff's voltage rule: $V=V_R +V_L$
# # where V is supply voltage,$V_R$ is potential across resistro and $V_L$ is induced emf across inductor
# # put $V_R=IR$, $V_L=L\frac{dI}{dt}$ in the above relation
# # we get
# # <font color='green'> $\frac{dI}{dt}= \frac{V}{L}-\frac{R}{L}I$ which is of the form of $\frac{dy}{dt}=f(t,y)$
# # here $I_0=0$, initially there is no current
# # True solution is $I=I_{max}(1-exp(-Rt/L))$,where $I_{max} =V/R$, max current in the ckt
# Python code for solving using Euler's method
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
f=lambda t,I: V/L-(R/L)*I
n=100
T=2
V=50
I=np.zeros(n)
I[0]=0
R=15
L=5
Imax=V/R
t=np.linspace(0,T,n)
truei=Imax*(1-np.exp(-R*t/L))
for i in range (0,n-1):
I[i+1]=I[i]+f(t[i],I[i])*(t[i+1]-t[i])
plt.plot(t,I,'red')
plt.plot(t,truei,'-og')
# -
plt.plot(t,truei)
# # Decay current:
# # when the battery is removed from the ckt(V=0) then charging condition becomes
# # $V=V_R+V_L$
# # $V_L=-V_R$
# # $ L\frac{dI}{dt}=-IR$
# # $\frac{dI}{dt}=-\frac{R}{L}I$
# # true solution= $I_{max}exp(-Rt/L)$, where $I_{max} $is max current $I_{max}= \frac{V}{R}$
# Code:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
f=lambda t,I: -(R/L)*I
n=20
T=2
V=50
I=np.zeros(n)
I[0]=Imax
R=15
L=5
Imax=V/R
t=np.linspace(0,T,n)
truei=Imax*(np.exp(-R*t/L))
for i in range (0,n-1):
I[i+1]=I[i]+f(t[i],I[i])*(t[i+1]-t[i])
plt.plot(t,I,'red')
plt.plot(t,truei,'-og')
plt.legend('decay')
# +
#
# +
# Code:while filling the tank of water
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
f=lambda r,h:2*r*np.pi*h
n=20
T=1
h=10
v=np.zeros(n)
v[0]=10
r=2
t=np.linspace(0,T,n)
for i in range (0,n-1):
v[i+1]=v[i]+f(t[i],v[i])*(t[i+1]-t[i])
plt.plot(t,v,'or')
# -
data={}
data.update({'time:':t,'volume:':v})
DF=pd.DataFrame(data)
print(DF)
df=pd.to_csv("water tank.csv")
DF.to_csv("water tank.csv")
df=pd.read_csv("water tank.csv")
# +
# Euler second order
# f
| .ipynb_checkpoints/EulersMethod-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="WTvhIdEILOpF"
# # ML 101
#
# ## Text Mining
#
# In this project, we will explore a simplified version of the DPWC model to draw a graph of similar personalities.
#
# The data will be consumed from Twitter, and for each search, we create a DWP profile with the most relevant words.
#
# After we apply a Matrix Factorization to estimate the "real" value of the word with zero weight.
#
# Finally, we use a thresholding method to create the entries on the graph.
# + id="W16qJeogxbbe"
# !pip install pyvis gensim python-twitter git+git://github.com/mariolpantunes/nmf@main#egg=nmf git+git://github.com/mariolpantunes/uts@main#egg=uts --upgrade
# + id="X66LmyIZ3UR4"
import pprint
import twitter
import numpy as np
from IPython.core.display import display, HTML
from pyvis.network import Network
import uts.thresholding as thres
import nltk
from nltk.corpus import stopwords
from nltk.stem import RSLPStemmer
from nltk.tokenize import RegexpTokenizer
from nmf.nmf import nmf_mu
import math
pp = pprint.PrettyPrinter(indent=2)
nltk.download('stopwords')
nltk.download('punkt')
nltk.download('wordnet')
nltk.download('omw-1.4')
nltk.download('rslp')
# + id="jFUqvRM_3e5l"
tokenizer = RegexpTokenizer(r'\w+')
stop_words = set(stopwords.words('portuguese'))
stop_words.update(set(stopwords.words('english')))
stop_words.add('https')
stemmer = RSLPStemmer()
db = {}
def cosine_similarity(a,b):
return np.dot(a, b)/(np.linalg.norm(a)*np.linalg.norm(b))
def generate_ngrams(tokens, n=2):
ngrams = zip(*[tokens[i:] for i in range(n)])
return [" ".join(ngram) for ngram in ngrams]
def tokenize(text):
tokens = tokenizer.tokenize(text)
tokens = [stemmer.stem(w.lower()) for w in tokens if not w in stop_words and w.isalpha() and len(w) > 3]
#ngrams = generate_ngrams(tokens)
#return tokens + ngrams
return tokens
def get_term_frequency(corpus, p=0.2):
tf = {}
# count the terms
for t in corpus:
if t not in tf:
tf[t] = 0
tf[t]+=1
# discard non-relevant items
neighborhood = [(k, v) for k, v in tf.items() if v > 1]
neighborhood.sort(key=lambda tup: tup[1], reverse=True)
limit = int(len(neighborhood)*p)
neighborhood = neighborhood[:limit]
# return
return neighborhood
# + id="OWmBDlBB7tqm"
api = twitter.Api(consumer_key='2lDgkNXdm03bxodf55vlY5IHo',
consumer_secret='<KEY>',
access_token_key='<KEY>',
access_token_secret='<KEY>')
terms = ["António Guterres", "<NAME>",
"<NAME>",
"<NAME>",
"<NAME>",
"<NAME>",
"<NAME>"]
for t in terms:
if t not in db:
results = api.GetSearch(term=t, count=300, lang='pt')
corpus = []
for r in results:
corpus.extend(tokenize(r.text.lower()))
tf = get_term_frequency(corpus)
db[t] = tf
pp.pprint(f'{db}')
# + id="4IFYRgY9dS92"
# create vector matrix
vocab = set()
for k in db:
tf = db[k]
for t,_ in tf:
vocab.add(t)
X = np.zeros((len(db), len(vocab)))
r = 0
for k in db:
tf = db[k]
c = 0
for t, v in tf:
vocab.add(t)
c += 1
X[r,c] = v
r +=1
rows, cols = X.shape
k = int(math.ceil(rows/2.0))
Xr, W, H, cost = nmf_mu(X, k=k, seed=42)
seeds = [3, 5, 7, 11, 13]
for s in seeds:
Xt, Wt, Ht, costt = nmf_mu(X, k=k, seed=s)
if costt < cost:
cost = costt
Xr = Xt
# compute the distance matrix (graph)
D = np.identity(len(db))
for i in range(0, len(Xr)):
for j in range(0, len(Xr)):
similarity = cosine_similarity(Xr[i], Xr[j])
D[i][j] = similarity
D[j][i] = similarity
# compute the ideal threshold
flat_distance = D.ravel()
#t = thres.isodata(flat_distance)
t = np.percentile(flat_distance, 75)
D[D<t] = 0
# + id="oQHrkE7ndS2u"
net = Network()
# Create the nodes
i=0
for k in db:
net.add_node(i, label=k)
i+=1
for i in range(0, len(D)-1):
for j in range(1, len(D)):
if (i!=j) and D[i][j] > 0:
net.add_edge(i, j, weight=D[i][j])
net.show('network.html')
display(HTML('network.html'))
| projects/06 text mining/06_tm_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"></ul></div>
# +
# %matplotlib inline
from time import sleep
import numpy as np
from livelossplot import PlotLosses
# +
liveplot = PlotLosses()
for i in range(10):
liveplot.update({
'accuracy': 1 - np.random.rand() / (i + 2.),
'val_accuracy': 1 - np.random.rand() / (i + 0.5),
'mse': 1. / (i + 2.),
'val_mse': 1. / (i + 0.5)
})
liveplot.draw()
sleep(1.)
# -
| examples/minimal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function
from bs4 import BeautifulSoup
import requests
# -
# # Beautiful soup on test data
#
# Here, we create some simple HTML that include some frequently used tags.
# Note, however, that we have also left one paragraph tag unclosed.
# +
source = """
<!DOCTYPE html>
<html>
<head>
<title>Scraping</title>
</head>
<body class="col-sm-12">
<h1>section1</h1>
<p>paragraph1</p>
<p>paragraph2</p>
<div class="col-sm-2">
<h2>section2</h2>
<p>paragraph3</p>
<p>unclosed
</div>
</body>
</html>
"""
soup = BeautifulSoup(source, "html.parser")
# -
# Once the soup object has been created successfully, we can execute a number of queries on the DOM.
# First we request all data from the `head` tag.
# Note that while it looks like a list of strings was returned, actually, a `bs4.element.Tag` type is returned.
# These examples expore how to extract tags, the text from tags, how to filter queries based on
# attributes, how to retreive attributes from a returned query, and how the BeautifulSoup engine
# is tolerant of unclosed tags.
print(soup.prettify())
print('Head:')
print('', soup.find_all("head"))
# [<head>\n<title>Scraping</title>\n</head>]
print('\nType of head:')
print('', map(type, soup.find_all("head")))
# [<class 'bs4.element.Tag'>]
print('\nTitle tag:')
print('', soup.find("title"))
# <title>Scraping</title>
print('\nTitle text:')
print('', soup.find("title").text)
# Scraping
divs = soup.find_all("div", attrs={"class": "col-sm-2"})
print('\nDiv with class=col-sm-2:')
print('', divs)
# [<div class="col-sm-2">....</div>]
print('\nClass of first div:')
print('', divs[0].attrs['class'])
# [u'col-sm-2']
print('\nAll paragraphs:')
print('', soup.find_all("p"))
# [<p>paragraph1</p>,
# <p>paragraph2</p>,
# <p>paragraph3</p>,
# <p>unclosed\n </p>]
# # Beautilful soup on real data
#
# In this example I will show how you can use BeautifulSoup to retreive information from live web pages.
# We make use of The Guardian newspaper, and retreive the HTML from an arbitrary article.
# We then create the BeautifulSoup object, and query the links that were discovered in the DOM.
# Since a large number are returned, we then apply attribute filters that let us reduce significantly
# the number of returned links.
# I selected the filters selected for this example in order to focus on the names in the paper.
# The parameterisation of the attributes was discovered by using the `inspect` functionality of Google Chrome
url = 'https://www.theguardian.com/technology/2017/jan/31/amazon-expedia-microsoft-support-washington-action-against-donald-trump-travel-ban'
req = requests.get(url)
source = req.text
soup = BeautifulSoup(source, 'html.parser')
print(source)
links = soup.find_all('a')
links
# +
links = soup.find_all('a', attrs={
'data-component': 'auto-linked-tag'
})
for link in links:
print(link['href'], link.text)
# -
# # Chaining queries
#
# Now, let us conisder a more general query that might be done on a website such as this.
# We will query the base technology page, and attempt to list all articles that pertain to this main page
url = 'https://www.theguardian.com/uk/technology'
req = requests.get(url)
source = req.text
soup = BeautifulSoup(source, 'html.parser')
# After inspecting the DOM (via the `inspect` tool in my browser), I see that the attributes that define
# a `technology` article are:
#
# class = "js-headline-text"
# +
articles = soup.find_all('a', attrs={
'class': 'js-headline-text'
})
for article in articles:
print(article['href'][:], article.text[:20])
# -
# With this set of articles, it is now possible to chain further querying, for example with code
# similar to the following
#
# ```python
# for article in articles:
# req = requests.get(article['href'])
# source = req.text
# soup = BeautifulSoup(source, 'html.parser')
#
# ... and so on...
# ```
#
# However, I won't go into much detail about this now. For scraping like this tools, such as `scrapy` are more
# appropriate than `BeautifulSoup` since they are designed for multithreadded web crawling.
# Once again, however, I urge caution and hope that before any crawling is initiated you determine whether
# crawling is within the terms of use of the website.
# If in doubt contact the website administrators.
#
# https://scrapy.org/
| 01_data_ingress/05_web_scraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Py3 research env
# language: python
# name: py3_research
# ---
# + [markdown] colab_type="text" id="13pL--6rycN3"
# ## Homework01: Three headed network in PyTorch
#
# This notebook accompanies the [week02 seminar](https://github.com/girafe-ai/ml-mipt/blob/advanced/week02_CNN_n_Vanishing_gradient/week02_CNN_for_texts.ipynb). Refer to that notebook for more comments.
#
# All the preprocessing is the same as in the classwork. *Including the data leakage in the train test split (it's still for bonus points).*
# + colab={} colab_type="code" id="P8zS7m-gycN5"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import nltk
import tqdm
from collections import Counter
# -
# If you have already downloaded the data on the Seminar, simply run through the next cells. Otherwise uncomment the next cell (and comment the another one ;)
# +
# uncomment and run this cell, if you don't have data locally yet.
# # !curl -L "https://www.dropbox.com/s/5msc5ix7ndyba10/Train_rev1.csv.tar.gz?dl=1" -o Train_rev1.csv.tar.gz
# # !tar -xvzf ./Train_rev1.csv.tar.gz
# data = pd.read_csv("./Train_rev1.csv", index_col=None)
# wget https://raw.githubusercontent.com/girafe-ai/ml-mipt/advanced_f20/homeworks_advanced/assignment1_02_Three_headed_network/network.py
# + colab={"base_uri": "https://localhost:8080/", "height": 143} colab_type="code" id="vwN72gd4ycOA" outputId="7b9e8549-3128-4041-c4be-33fb6f326c78"
# run this cell if you have downloaded the dataset on the seminar
data = pd.read_csv("../../week02_CNN_n_Vanishing_gradient/Train_rev1.csv", index_col=None)
# + colab={"base_uri": "https://localhost:8080/", "height": 265} colab_type="code" id="UuuKIKfrycOH" outputId="e5de0f94-a4f6-4b51-db80-9d11ddc1db31"
data['Log1pSalary'] = np.log1p(data['SalaryNormalized']).astype('float32')
text_columns = ["Title", "FullDescription"]
categorical_columns = ["Category", "Company", "LocationNormalized", "ContractType", "ContractTime"]
target_column = "Log1pSalary"
data[categorical_columns] = data[categorical_columns].fillna('NaN') # cast missing values to string "NaN"
data.sample(3)
data_for_autotest = data[-5000:]
data = data[:-5000]
# + colab={} colab_type="code" id="RUWkpd7PycOQ"
tokenizer = nltk.tokenize.WordPunctTokenizer()
# see task above
def normalize(text):
text = str(text).lower()
return ' '.join(tokenizer.tokenize(text))
data[text_columns] = data[text_columns].applymap(normalize)
print("Tokenized:")
print(data["FullDescription"][2::100000])
assert data["FullDescription"][2][:50] == 'mathematical modeller / simulation analyst / opera'
assert data["Title"][54321] == 'international digital account manager ( german )'
# Count how many times does each token occur in both "Title" and "FullDescription" in total
# build a dictionary { token -> it's count }
from collections import Counter
from tqdm import tqdm as tqdm
token_counts = Counter()# <YOUR CODE HERE>
for _, row in tqdm(data[text_columns].iterrows()):
for string in row:
token_counts.update(string.split())
# hint: you may or may not want to use collections.Counter
# -
token_counts.most_common(1)[0][1]
# + colab={"base_uri": "https://localhost:8080/", "height": 215} colab_type="code" id="GiOWbc15ycOb" outputId="1e807140-5513-4af0-d9a9-9f029059a553"
print("Total unique tokens :", len(token_counts))
print('\n'.join(map(str, token_counts.most_common(n=5))))
print('...')
print('\n'.join(map(str, token_counts.most_common()[-3:])))
assert token_counts.most_common(1)[0][1] in range(2500000, 2700000)
assert len(token_counts) in range(200000, 210000)
print('Correct!')
min_count = 10
# tokens from token_counts keys that had at least min_count occurrences throughout the dataset
tokens = [token for token, count in token_counts.items() if count >= min_count]# <YOUR CODE HERE>
# Add a special tokens for unknown and empty words
UNK, PAD = "UNK", "PAD"
tokens = [UNK, PAD] + sorted(tokens)
print("Vocabulary size:", len(tokens))
assert type(tokens) == list
assert len(tokens) in range(32000, 35000)
assert 'me' in tokens
assert UNK in tokens
print("Correct!")
token_to_id = {token: idx for idx, token in enumerate(tokens)}
assert isinstance(token_to_id, dict)
assert len(token_to_id) == len(tokens)
for tok in tokens:
assert tokens[token_to_id[tok]] == tok
print("Correct!")
# + colab={} colab_type="code" id="JEsLeBjVycOw"
UNK_IX, PAD_IX = map(token_to_id.get, [UNK, PAD])
def as_matrix(sequences, max_len=None):
""" Convert a list of tokens into a matrix with padding """
if isinstance(sequences[0], str):
sequences = list(map(str.split, sequences))
max_len = min(max(map(len, sequences)), max_len or float('inf'))
matrix = np.full((len(sequences), max_len), np.int32(PAD_IX))
for i,seq in enumerate(sequences):
row_ix = [token_to_id.get(word, UNK_IX) for word in seq[:max_len]]
matrix[i, :len(row_ix)] = row_ix
return matrix
# + colab={"base_uri": "https://localhost:8080/", "height": 179} colab_type="code" id="JiBlPkdKycOy" outputId="3866b444-1e2d-4d79-d429-fecc6d8e02a8"
print("Lines:")
print('\n'.join(data["Title"][::100000].values), end='\n\n')
print("Matrix:")
print(as_matrix(data["Title"][::100000]))
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="DpOlBp7ZycO6" outputId="30a911f2-7d35-4cb5-8991-60457b1e8bac"
from sklearn.feature_extraction import DictVectorizer
# we only consider top-1k most frequent companies to minimize memory usage
top_companies, top_counts = zip(*Counter(data['Company']).most_common(1000))
recognized_companies = set(top_companies)
data["Company"] = data["Company"].apply(lambda comp: comp if comp in recognized_companies else "Other")
categorical_vectorizer = DictVectorizer(dtype=np.float32, sparse=False)
categorical_vectorizer.fit(data[categorical_columns].apply(dict, axis=1))
# + [markdown] colab_type="text" id="yk4jmtAYycO8"
# ### The deep learning part
#
# Once we've learned to tokenize the data, let's design a machine learning experiment.
#
# As before, we won't focus too much on validation, opting for a simple train-test split.
#
# __To be completely rigorous,__ we've comitted a small crime here: we used the whole data for tokenization and vocabulary building. A more strict way would be to do that part on training set only. You may want to do that and measure the magnitude of changes.
#
#
# #### Here comes the simple one-headed network from the seminar.
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="TngLcWA0ycO_" outputId="6731b28c-07b1-41dc-9574-f76b01785bba"
from sklearn.model_selection import train_test_split
data_train, data_val = train_test_split(data, test_size=0.2, random_state=42)
data_train.index = range(len(data_train))
data_val.index = range(len(data_val))
print("Train size = ", len(data_train))
print("Validation size = ", len(data_val))
# + colab={} colab_type="code" id="2PXuKgOSycPB"
def make_batch(data, max_len=None, word_dropout=0):
"""
Creates a keras-friendly dict from the batch data.
:param word_dropout: replaces token index with UNK_IX with this probability
:returns: a dict with {'title' : int64[batch, title_max_len]
"""
batch = {}
batch["Title"] = as_matrix(data["Title"].values, max_len)
batch["FullDescription"] = as_matrix(data["FullDescription"].values, max_len)
batch['Categorical'] = categorical_vectorizer.transform(data[categorical_columns].apply(dict, axis=1))
if word_dropout != 0:
batch["FullDescription"] = apply_word_dropout(batch["FullDescription"], 1. - word_dropout)
if target_column in data.columns:
batch[target_column] = data[target_column].values
return batch
def apply_word_dropout(matrix, keep_prop, replace_with=UNK_IX, pad_ix=PAD_IX,):
dropout_mask = np.random.choice(2, np.shape(matrix), p=[keep_prop, 1 - keep_prop])
dropout_mask &= matrix != pad_ix
return np.choose(dropout_mask, [matrix, np.full_like(matrix, replace_with)])
# + colab={"base_uri": "https://localhost:8080/", "height": 251} colab_type="code" id="I6LpEQf0ycPD" outputId="e3520cae-fba1-46cc-a216-56287b6e4929"
a = make_batch(data_train[:3], max_len=10)
# -
# But to start with let's build the simple model using only the part of the data. Let's create the baseline solution using only the description part (so it should definetely fit into the Sequential model).
import torch
from torch import nn
import torch.nn.functional as F
# +
# You will need these to make it simple
class Flatten(nn.Module):
def forward(self, input):
return input.view(input.size(0), -1)
class Reorder(nn.Module):
def forward(self, input):
return input.permute((0, 2, 1))
# -
# To generate minibatches we will use simple pyton generator.
def iterate_minibatches(data, batch_size=256, shuffle=True, cycle=False, **kwargs):
""" iterates minibatches of data in random order """
while True:
indices = np.arange(len(data))
if shuffle:
indices = np.random.permutation(indices)
for start in range(0, len(indices), batch_size):
batch = make_batch(data.iloc[indices[start : start + batch_size]], **kwargs)
target = batch.pop(target_column)
yield batch, target
if not cycle: break
iterator = iterate_minibatches(data_train, 3)
batch, target = next(iterator)
# +
# Here is some startup code:
n_tokens=len(tokens)
n_cat_features=len(categorical_vectorizer.vocabulary_)
hid_size=64
simple_model = nn.Sequential()
simple_model.add_module('emb', nn.Embedding(num_embeddings=n_tokens, embedding_dim=hid_size))
simple_model.add_module('reorder', Reorder())
simple_model.add_module('conv1', nn.Conv1d(
in_channels=hid_size,
out_channels=hid_size,
kernel_size=2)
)
simple_model.add_module('relu1', nn.ReLU())
simple_model.add_module('adapt_avg_pool', nn.AdaptiveAvgPool1d(output_size=1))
simple_model.add_module('flatten1', Flatten())
simple_model.add_module('linear1', nn.Linear(in_features=hid_size, out_features=1))
# <YOUR CODE HERE>
# -
batch
# __Remember!__ We are working with regression problem and predicting only one number.
# Try this to check your model. `torch.long` tensors are required for nn.Embedding layers.
simple_model(torch.tensor(batch['FullDescription'], dtype=torch.long))
batch['FullDescription'].shape
# And now simple training pipeline (it's commented because we've already done that in class. No need to do it again).
# +
# from IPython.display import clear_output
# from random import sample
# epochs = 1
# model = simple_model
# opt = torch.optim.Adam(model.parameters())
# loss_func = nn.MSELoss()
# history = []
# for epoch_num in range(epochs):
# for idx, (batch, target) in enumerate(iterate_minibatches(data_train)):
# # Preprocessing the batch data and target
# batch = torch.tensor(batch['FullDescription'], dtype=torch.long)
# target = torch.tensor(target)
# predictions = model(batch)
# predictions = predictions.view(predictions.size(0))
# loss = loss_func(predictions, target)# <YOUR CODE HERE>
# # train with backprop
# loss.backward()
# opt.step()
# opt.zero_grad()
# # <YOUR CODE HERE>
# history.append(loss.data.numpy())
# if (idx+1)%10==0:
# clear_output(True)
# plt.plot(history,label='loss')
# plt.legend()
# plt.show()
# -
# ### Actual homework starts here
# __Your ultimate task is to code the three headed network described on the picture below.__
# To make it closer to the real world, please store the network code in file `network.py` in this directory.
# + [markdown] colab_type="text" id="0eI5h9UMycPF"
# #### Architecture
#
# Our main model consists of three branches:
# * Title encoder
# * Description encoder
# * Categorical features encoder
#
# We will then feed all 3 branches into one common network that predicts salary.
#
# <img src="https://github.com/yandexdataschool/nlp_course/raw/master/resources/w2_conv_arch.png" width=600px>
#
# This clearly doesn't fit into PyTorch __Sequential__ interface. To build such a network, one will have to use [__PyTorch nn.Module API__](https://pytorch.org/docs/stable/nn.html#torch.nn.Module).
# -
import network
# Re-run this cell if you updated the file with network source code
import imp
imp.reload(network)
model = network.ThreeInputsNet(
n_tokens=len(tokens),
n_cat_features=len(categorical_vectorizer.vocabulary_),
# this parameter defines the number of the inputs in the layer,
# which stands after the concatenation. In should be found out by you.
concat_number_of_features=<YOUR CODE HERE>
)
testing_batch, _ = next(iterate_minibatches(data_train, 3))
testing_batch = [
torch.tensor(testing_batch['Title'], dtype=torch.long),
torch.tensor(testing_batch['FullDescription'], dtype=torch.long),
torch.tensor(testing_batch['Categorical'])
]
assert model(testing_batch).shape == torch.Size([3, 1])
assert model(testing_batch).dtype == torch.float32
print('Seems fine!')
# Now train the network for a while (100 batches would be fine).
# +
# Training pipeline comes here (almost the same as for the simple_model)
# -
# Now, to evaluate the model it can be switched to `eval` state.
model.eval()
def generate_submission(model, data, batch_size=256, name="", three_inputs_mode=True, **kw):
squared_error = abs_error = num_samples = 0.0
output_list = []
for batch_x, batch_y in tqdm(iterate_minibatches(data, batch_size=batch_size, shuffle=False, **kw)):
if three_inputs_mode:
batch = [
torch.tensor(batch_x['Title'], dtype=torch.long),
torch.tensor(batch_x['FullDescription'], dtype=torch.long),
torch.tensor(batch_x['Categorical'])
]
else:
batch = torch.tensor(batch_x['FullDescription'], dtype=torch.long)
batch_pred = model(batch)[:, 0].detach().numpy()
output_list.append((list(batch_pred), list(batch_y)))
squared_error += np.sum(np.square(batch_pred - batch_y))
abs_error += np.sum(np.abs(batch_pred - batch_y))
num_samples += len(batch_y)
print("%s results:" % (name or ""))
print("Mean square error: %.5f" % (squared_error / num_samples))
print("Mean absolute error: %.5f" % (abs_error / num_samples))
batch_pred = [c for x in output_list for c in x[0]]
batch_y = [c for x in output_list for c in x[1]]
output_df = pd.DataFrame(list(zip(batch_pred, batch_y)), columns=['batch_pred', 'batch_y'])
output_df.to_csv('submission.csv', index=False)
generate_submission(model, data_for_autotest, name='Submission')
print('Submission file generated')
# __To hand in this homework, please upload `network.py` file with code and `submission.csv` to the google form.__
| ML-MIPT-advanced/homeworks_advanced/assignment1_02_Three_headed_network/assignment1_02_three_headed_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dev3.7]
# language: python
# name: conda-env-dev3.7-py
# ---
# # Counter
# In this activity, you will create a function that preprocesses and outputs a list of the most common words in a corpus.
from nltk.corpus import reuters, stopwords
from nltk.util import ngrams
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
import re
import pandas as pd
from collections import Counter
lemmatizer = WordNetLemmatizer()
# Corpus - list of articles about grains
ids = reuters.fileids(categories='grain')
corpus = [reuters.raw(i) for i in ids]
# Define preprocess function
def process_text(doc):
sw = set(stopwords.words('english'))
regex = re.compile("[^a-zA-Z ]")
re_clean = regex.sub('', doc)
words = word_tokenize(re_clean)
lem = [lemmatizer.lemmatize(word) for word in words]
output = [word.lower() for word in lem if word.lower() not in sw]
return output
# Define the counter function
def word_counter(corpus):
# Combine all articles in corpus into one large string
big_string = ' '.join(corpus)
processed = process_text(big_string)
top_10 = dict(Counter(processed).most_common(10))
return pd.DataFrame(list(top_10.items()), columns=['word', 'count'])
# Run the word_counter function
word_counter(corpus)
def bigram_counter(corpus):
# Combine all articles in corpus into one large string
big_string = ' '.join(corpus)
processed = process_text(big_string)
bigrams = ngrams(processed, n=2)
top_10 = dict(Counter(bigrams).most_common(10))
return pd.DataFrame(list(top_10.items()), columns=['bigram', 'count'])
# Run the bigram_counter function
bigram_counter(corpus)
def trigram_counter(corpus):
# Combine all articles in corpus into one large string
big_string = ' '.join(corpus)
processed = process_text(big_string)
bigrams = ngrams(processed, n=3)
top_10 = dict(Counter(bigrams).most_common(10))
return pd.DataFrame(list(top_10.items()), columns=['trigram', 'count'])
trigram_counter(corpus)
# +
def quadgram_counter(corpus):
# Combine all articles in corpus into one large string
big_string = ' '.join(corpus)
processed = process_text(big_string)
bigrams = ngrams(processed, n=4)
top_10 = dict(Counter(bigrams).most_common(10))
return pd.DataFrame(list(top_10.items()), columns=['quadgram', 'count'])
quadgram_counter(corpus)
# -
| PythonJupyterNotebooks/Week12-Day1-Activity8-counter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
pm_10 = pd.read_csv('yurukov/sofia_airpollution.csv')
# # Seasons
weather_raw = pd.read_csv("./data/sofia-weather-data.csv")
weather = weather_raw.dropna(axis=1, how='all')
weather.drop(axis=1,labels=['weather_icon'], inplace=True)
# +
weather["temp_cent"] = weather["temp"] - 272.15
weather["temp_max_cent"] = weather["temp_max"] - 272.15
weather["temp_min_cent"] = weather["temp_min"] - 272.15
weather['date'] = weather['dt_iso'].apply(lambda x : str.split(x, ' +')[0])
# f = {'temp_cent':['min','mean', 'max'], 'pressure':['mean'], 'humidity': ['mean'], 'wind_speed': ['min', 'mean', 'max']}
processed_weather = weather.groupby('date', as_index=False).mean()
# -
dates = pd.DataFrame(weather['date'].unique(), columns=["date"])
# weather = weather[weather['aaa'] != 1]
pm_10 = pm_10[pm_10['inv_PM10'] != 1]
pm_10 = pm_10[pm_10['inv_NO2'] != 1]
pm_10['timestamp'] = pm_10['timestamp'].astype("str")
processed_weather['date'] = processed_weather['date'].astype("str")
merged_with_date = pd.merge(processed_weather, pm_10, left_on='date', right_on="timestamp")
merged_with_date = merged_with_date.drop(axis=1, labels=['temp', 'dt', 'city_id', 'wind_deg', 'rain_1h', 'rain_3h',
'rain_today', 'weather_id', 'inv_PM10', 'inv_NO2', 'NO', 'inv_NO', 'C6H6', 'inv_C6H6', 'CO',
'inv_CO', 'O3', 'inv_O3', 'SO2', 'inv_SO2', 'HU', 'inv_HU', 'AP',
'inv_AP', 'WND', 'inv_WND', 'SUN', 'inv_SUN', 'TMP', 'inv_TMP', 'timestamp'])
#print(merged_with_date.columns)
#print(merged_with_date.head())
#print(merged_with_date.describe())
#print(len(merged_with_date))
merged_with_date
# # Shift weather one day forward - for predictions
# +
merged_with_date["temp_shifted"] = merged_with_date.temp_cent.shift(1)
merged_with_date["temp_min_shifted"] = merged_with_date.temp_min.shift(1)
merged_with_date["temp_max_shifted"] = merged_with_date.temp_max.shift(1)
#TODO
| yurukov_weather_merging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (QISKitenv)
# language: python
# name: qiskitenv
# ---
# <img src="../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# ## _*Shor's Algorithm for Integer Factorization*_
#
# The latest version of this tutorial notebook is available on https://github.com/qiskit/qiskit-tutorial.
#
# In this tutorial, we first introduce the problem of [integer factorization](#factorization) and describe how [Shor's algorithm](#shorsalgorithm) solves it in detail. We then [implement](#implementation) a version of it in Qiskit.
#
# ### Contributors
# <NAME>
# ***
# ## Integer Factorization <a id='factorization'></a>
#
# Integer factorization is the decomposition of an composite integer into a product of smaller integers, for example, the integer $100$ can be factored into $10 \times 10$. If these factors are restricted to prime numbers, the process is called prime factorization, for example, the prime factorization of $100$ is $2 \times 2 \times 5 \times 5$.
#
# When the integers are very large, no efficient classical integer factorization algorithm is known. The hardest factorization problems are semiprime numbers, the product of two prime numbers. In [2009](https://link.springer.com/chapter/10.1007/978-3-642-14623-7_18), a team of researchers factored a 232 decimal digit semiprime number (768 bits), spending the computational equivalent of more than two thousand years on a single core 2.2 GHz AMD Opteron processor with 2 GB RAM:
# ```
# RSA-768 = 12301866845301177551304949583849627207728535695953347921973224521517264005
# 07263657518745202199786469389956474942774063845925192557326303453731548268
# 50791702612214291346167042921431160222124047927473779408066535141959745985
# 6902143413
#
# = 33478071698956898786044169848212690817704794983713768568912431388982883793
# 878002287614711652531743087737814467999489
# × 36746043666799590428244633799627952632279158164343087642676032283815739666
# 511279233373417143396810270092798736308917
# ```
# The presumed difficulty of this semiprime factorization problem underlines many encryption algorithms, such as [RSA](https://www.google.com/patents/US4405829), which is used in online credit card transactions, amongst other applications.
# ***
# ## Shor's Algorithm <a id='shorsalgorithm'></a>
#
# Shor's algorithm, named after mathematician <NAME>, is a polynomial time quantum algorithm for integer factorization formulated in [1994](http://epubs.siam.org/doi/10.1137/S0097539795293172). It is arguably the most dramatic example of how the paradigm of quantum computing changed our perception of which computational problems should be considered tractable, motivating the study of new quantum algorithms and efforts to design and construct quantum computers. It also has expedited research into new cryptosystems not based on integer factorization.
#
# Shor's algorithm has been experimentally realised by multiple teams for specific composite integers. The composite $15$ was first factored into $3 \times 5$ in [2001](https://www.nature.com/nature/journal/v414/n6866/full/414883a.html) using seven NMR qubits, and has since been implemented using four photon qubits in 2007 by [two](https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.99.250504) [teams](https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.99.250505), three solid state qubits in [2012](https://www.nature.com/nphys/journal/v8/n10/full/nphys2385.html) and five trapped ion qubits in [2016](http://science.sciencemag.org/content/351/6277/1068). The composite $21$ has also been factored into $3 \times 7$ in [2012](http://www.nature.com/nphoton/journal/v6/n11/full/nphoton.2012.259.html) using a photon qubit and qutrit (a three level system). Note that these experimental demonstrations rely on significant optimisations of Shor's algorithm based on apriori knowledge of the expected results. In general, [$2 + \frac{3}{2}\log_2N$](https://link-springer-com.virtual.anu.edu.au/chapter/10.1007/3-540-49208-9_15) qubits are needed to factor the composite integer $N$, meaning at least $1,154$ qubits would be needed to factor $RSA-768$ above.
#
from IPython.display import HTML
HTML('<iframe width="560" height="315" src="https://www.youtube.com/embed/hOlOY7NyMfs?start=75&end=126" frameborder="0" allowfullscreen></iframe>')
# As <NAME> describes in the video above from [PhysicsWorld](http://physicsworld.com/cws/article/multimedia/2015/sep/30/what-is-shors-factoring-algorithm), Shor’s algorithm is composed of three parts. The first part turns the factoring problem into a period finding problem using number theory, which can be computed on a classical computer. The second part finds the period using the quantum Fourier transform and is responsible for the quantum speedup of the algorithm. The third part uses the period found to calculate the factors.
#
# The following sections go through the algorithm in detail, for those who just want the steps, without the lengthy explanation, refer to the [blue](#stepsone) [boxes](#stepstwo) before jumping down to the [implemention](#implemention).
# ### From Factorization to Period Finding
#
# The number theory that underlines Shor's algorithm relates to periodic modulo sequences. Let's have a look at an example of such a sequence. Consider the sequence of the powers of two:
# $$1, 2, 4, 8, 16, 32, 64, 128, 256, 512, 1024, ...$$
# Now let's look at the same sequence 'modulo 15', that is, the remainder after fifteen divides each of these powers of two:
# $$1, 2, 4, 8, 1, 2, 4, 8, 1, 2, 4, ...$$
# This is a modulo sequence that repeats every four numbers, that is, a periodic modulo sequence with a period of four.
#
# Reduction of factorization of $N$ to the problem of finding the period of an integer $x$ less than $N$ and greater than $1$ depends on the following result from number theory:
#
# > The function $\mathcal{F}(a) = x^a \bmod N$ is a periodic function, where $x$ is an integer coprime to $N$ and $a \ge 0$.
#
# Note that two numbers are coprime, if the only positive integer that divides both of them is 1. This is equivalent to their greatest common divisor being 1. For example, 8 and 15 are coprime, as they don't share any common factors (other than 1). However, 9 and 15 are not coprime, since they are both divisible by 3 (and 1).
#
# > Since $\mathcal{F}(a)$ is a periodic function, it has some period $r$. Knowing that $x^0 \bmod N = 1$, this means that $x^r \bmod N = 1$ since the function is periodic, and thus $r$ is just the first nonzero power where $x^r = 1 (\bmod N)$.
#
# Given this information and through the following algebraic manipulation:
# $$ x^r \equiv 1 \bmod N $$
# $$ x^r = (x^{r/2})^2 \equiv 1 \bmod N $$
# $$ (x^{r/2})^2 - 1 \equiv 0 \bmod N $$
# and if $r$ is an even number:
# $$ (x^{r/2} + 1)(x^{r/2} - 1) \equiv 0 \bmod N $$
#
# From this, the product $(x^{r/2} + 1)(x^{r/2} - 1)$ is an integer multiple of $N$, the number to be factored. Thus, so long as $(x^{r/2} + 1)$ or $(x^{r/2} - 1)$ is not a multiple of $N$, then at least one of $(x^{r/2} + 1)$ or $(x^{r/2} - 1)$ must have a nontrivial factor in common with $N$.
#
# So computing $\text{gcd}(x^{r/2} - 1, N)$ and $\text{gcd}(x^{r/2} + 1, N)$ will obtain a factor of $N$, where $\text{gcd}$ is the greatest common denominator function, which can be calculated by the polynomial time [Euclidean algorithm](https://en.wikipedia.org/wiki/Euclidean_algorithm).
# #### Classical Steps to Shor's Algorithm
#
# Let's assume for a moment that a period finding machine exists that takes as input coprime integers $x, N$ and outputs the period of $x \bmod N$, implemented by as a brute force search below. Let's show how to use the machine to find all prime factors of $N$ using the number theory described above.
# Brute force period finding algorithm
def find_period_classical(x, N):
n = 1
t = x
while t != 1:
t *= x
t %= N
n += 1
return n
# For simplicity, assume that $N$ has only two distinct prime factors: $N = pq$.
#
# <div class="alert alert-block alert-info"> <a id='stepsone'></a>
# <ol>
# <li>Pick a random integer $x$ between $1$ and $N$ and compute the greatest common divisor $\text{gcd}(x,N)$ using Euclid's algorithm.</li>
# <li>If $x$ and $N$ have some common prime factors, $\text{gcd}(x,N)$ will equal $p$ or $q$. Otherwise $\text{gcd}(x,N) = 1$, meaning $x$ and $N$ are coprime. </li>
# <li>Let $r$ be the period of $x \bmod N$ computed by the period finding machine. Repeat the above steps with different random choices of $x$ until $r$ is even.</li>
# <li>Now $p$ and $q$ can be found by computing $\text{gcd}(x^{r/2} \pm 1, N)$ as long as $x^{r/2} \neq \pm 1$.</li>
# </ol>
# </div>
#
# As an example, consider $N = 15$. Let's look at all values of $1 < x < 15$ where $x$ is coprime with $15$:
#
# | $x$ | $x^a \bmod 15$ | Period $r$ |$\text{gcd}(x^{r/2}-1,15)$|$\text{gcd}(x^{r/2}+1,15)$ |
# |:-----:|:----------------------------:|:----------:|:------------------------:|:-------------------------:|
# | 2 | 1,2,4,8,1,2,4,8,1,2,4... | 4 | 3 | 5 |
# | 4 | 1,4,1,4,1,4,1,4,1,4,1... | 2 | 3 | 5 |
# | 7 | 1,7,4,13,1,7,4,13,1,7,4... | 4 | 3 | 5 |
# | 8 | 1,8,4,2,1,8,4,2,1,8,4... | 4 | 3 | 5 |
# | 11 | 1,11,1,11,1,11,1,11,1,11,1...| 2 | 5 | 3 |
# | 13 | 1,13,4,7,1,13,4,7,1,13,4,... | 4 | 3 | 5 |
# | 14 | 1,14,1,14,1,14,1,14,1,14,1,,,| 2 | 1 | 15 |
#
# As can be seen, any value of $x$ except $14$ will return the factors of $15$, that is, $3$ and $5$. $14$ is an example of the special case where $(x^{r/2} + 1)$ or $(x^{r/2} - 1)$ is a multiple of $N$ and thus another $x$ needs to be tried.
#
# In general, it can be shown that this special case occurs infrequently, so on average only two calls to the period finding machine are sufficient to factor $N$.
# For a more interesting example, first let's find larger number N, that is semiprime that is relatively small. Using the [Sieve of Eratosthenes](https://en.wikipedia.org/wiki/Sieve_of_Eratosthenes) [Python implementation](http://archive.oreilly.com/pub/a/python/excerpt/pythonckbk_chap1/index1.html?page=last), let's generate a list of all the prime numbers less than a thousand, randomly select two, and muliply them.
# +
import random, itertools
# Sieve of Eratosthenes algorithm
def sieve( ):
D = { }
yield 2
for q in itertools.islice(itertools.count(3), 0, None, 2):
p = D.pop(q, None)
if p is None:
D[q*q] = q
yield q
else:
x = p + q
while x in D or not (x&1):
x += p
D[x] = p
# Creates a list of prime numbers up to the given argument
def get_primes_sieve(n):
return list(itertools.takewhile(lambda p: p<n, sieve()))
def get_semiprime(n):
primes = get_primes_sieve(n)
l = len(primes)
p = primes[random.randrange(l)]
q = primes[random.randrange(l)]
return p*q
N = get_semiprime(1000)
print("semiprime N =",N)
# -
# Now implement the [above steps](#stepsone) of Shor's Algorithm:
# +
import math
def shors_algorithm_classical(N):
x = random.randint(0,N) # step one
if(math.gcd(x,N) != 1): # step two
return x,0,math.gcd(x,N),N/math.gcd(x,N)
r = find_period_classical(x,N) # step three
while(r % 2 != 0):
r = find_period_classical(x,N)
p = math.gcd(x**int(r/2)+1,N) # step four, ignoring the case where (x^(r/2) +/- 1) is a multiple of N
q = math.gcd(x**int(r/2)-1,N)
return x,r,p,q
x,r,p,q = shors_algorithm_classical(N)
print("semiprime N = ",N,", coprime x = ",x,", period r = ",r,", prime factors = ",p," and ",q,sep="")
# -
# ### Quantum Period Finding <a id='quantumperiodfinding'></a>
#
# Let's first describe the quantum period finding algorithm, and then go through a few of the steps in detail, before going through an example. This algorithm takes two coprime integers, $x$ and $N$, and outputs $r$, the period of $\mathcal{F}(a) = x^a\bmod N$.
#
# <div class="alert alert-block alert-info"><a id='stepstwo'></a>
# <ol>
# <li> Choose $T = 2^t$ such that $N^2 \leq T \le 2N^2$. Initialise two registers of qubits, first an argument register with $t$ qubits and second a function register with $n = log_2 N$ qubits. These registers start in the initial state:
# $$\vert\psi_0\rangle = \vert 0 \rangle \vert 0 \rangle$$ </li>
# <li> Apply a Hadamard gate on each of the qubits in the argument register to yield an equally weighted superposition of all integers from $0$ to $T$:
# $$\vert\psi_1\rangle = \frac{1}{\sqrt{T}}\sum_{a=0}^{T-1}\vert a \rangle \vert 0 \rangle$$ </li>
# <li> Implement the modular exponentiation function $x^a \bmod N$ on the function register, giving the state:
# $$\vert\psi_2\rangle = \frac{1}{\sqrt{T}}\sum_{a=0}^{T-1}\vert a \rangle \vert x^a \bmod N \rangle$$
# This $\vert\psi_2\rangle$ is highly entangled and exhibits quantum parallism, i.e. the function entangled in parallel all the 0 to $T$ input values with the corresponding values of $x^a \bmod N$, even though the function was only executed once. </li>
# <li> Perform a quantum Fourier transform on the argument register, resulting in the state:
# $$\vert\psi_3\rangle = \frac{1}{T}\sum_{a=0}^{T-1}\sum_{z=0}^{T-1}e^{(2\pi i)(az/T)}\vert z \rangle \vert x^a \bmod N \rangle$$
# where due to the interference, only the terms $\vert z \rangle$ with
# $$z = qT/r $$
# have significant amplitude where $q$ is a random integer ranging from $0$ to $r-1$ and $r$ is the period of $\mathcal{F}(a) = x^a\bmod N$. </li>
# <li> Measure the argument register to obtain classical result $z$. With reasonable probability, the continued fraction approximation of $T / z$ will be an integer multiple of the period $r$. Euclid's algorithm can then be used to find $r$.</li>
# </ol>
# </div>
#
# Note how quantum parallelism and constructive interference have been used to detect and measure periodicity of the modular exponentiation function. The fact that interference makes it easier to measure periodicity should not come as a big surprise. After all, physicists routinely use scattering of electromagnetic waves and interference measurements to determine periodicity of physical objects such as crystal lattices. Likewise, Shor's algorithm exploits interference to measure periodicity of arithmetic objects, a computational interferometer of sorts.
# #### Modular Exponentiation
#
# The modular exponentiation, step 3 above, that is the evaluation of $x^a \bmod N$ for $2^t$ values of $a$ in parallel, is the most demanding part of the algorithm. This can be performed using the following identity for the binary representation of any integer: $x = x_{t-1}2^{t-1} + ... x_12^1+x_02^0$, where $x_t$ are the binary digits of $x$. From this, it follows that:
#
# \begin{aligned}
# x^a \bmod N & = x^{2^{(t-1)}a_{t-1}} ... x^{2a_1}x^{a_0} \bmod N \\
# & = x^{2^{(t-1)}a_{t-1}} ... [x^{2a_1}[x^{2a_0} \bmod N] \bmod N] ... \bmod N \\
# \end{aligned}
#
# This means that 1 is first multiplied by $x^1 \bmod N$ if and only if $a_0 = 1$, then the result is multiplied by $x^2 \bmod N$ if and only if $a_1 = 1$ and so forth, until finally the result is multiplied by $x^{2^{(s-1)}}\bmod N$ if and only if $a_{t-1} = 1$.
#
# Therefore, the modular exponentiation consists of $t$ serial multiplications modulo $N$, each of them controlled by the qubit $a_t$. The values $x,x^2,...,x^{2^{(t-1)}} \bmod N$ can be found efficiently on a classical computer by repeated squaring.
# #### Quantum Fourier Transform
#
# The Fourier transform occurs in many different versions throughout classical computing, in areas ranging from signal processing to data compression to complexity theory. The quantum Fourier transform (QFT), step 4 above, is the quantum implementation of the discrete Fourier transform over the amplitudes of a wavefunction.
#
# The classical discrete Fourier transform acts on a vector $(x_0, ..., x_{N-1})$ and maps it to the vector $(y_0, ..., y_{N-1})$ according to the formula
# $$y_k = \frac{1}{\sqrt{N}}\sum_{j=0}^{N-1}x_j\omega_N^{jk}$$
# where $\omega_N^{jk} = e^{2\pi i \frac{jk}{N}}$.
#
# Similarly, the quantum Fourier transform acts on a quantum state $\sum_{i=0}^{N-1} x_i \vert i \rangle$ and maps it to the quantum state $\sum_{i=0}^{N-1} y_i \vert i \rangle$ according to the formula
# $$y_k = \frac{1}{\sqrt{N}}\sum_{j=0}^{N-1}x_j\omega_N^{jk}$$
# with $\omega_N^{jk}$ defined as above. Note that only the amplitudes of the state were affected by this transformation.
#
# This can also be expressed as the map:
# $$\vert x \rangle \mapsto \frac{1}{\sqrt{N}}\sum_{y=0}^{N-1}\omega_N^{xy} \vert y \rangle$$
#
# Or the unitary matrix:
# $$ U_{QFT} = \frac{1}{\sqrt{N}} \sum_{x=0}^{N-1} \sum_{y=0}^{N-1} \omega_N^{xy} \vert y \rangle \langle x \vert$$
# As an example, we've actually already seen the quantum Fourier transform for when $N = 2$, it is the Hadamard operator ($H$):
# $$H = \frac{1}{\sqrt{2}}\begin{bmatrix} 1 & 1 \\ 1 & -1 \end{bmatrix}$$
# Suppose we have the single qubit state $\alpha \vert 0 \rangle + \beta \vert 1 \rangle$, if we apply the $H$ operator to this state, we obtain the new state:
# $$\frac{1}{\sqrt{2}}(\alpha + \beta) \vert 0 \rangle + \frac{1}{\sqrt{2}}(\alpha - \beta) \vert 1 \rangle
# \equiv \tilde{\alpha}\vert 0 \rangle + \tilde{\beta}\vert 1 \rangle$$
# Notice how the Hadamard gate performs the discrete Fourier transform for $N = 2$ on the amplitudes of the state.
#
# So what does the quantum Fourier transform look like for larger N? Let's derive a circuit for $N=2^n$, $QFT_N$ acting on the state $\vert x \rangle = \vert x_1...x_n \rangle$ where $x_1$ is the most significant bit.
# \begin{aligned}
# QFT_N\vert x \rangle & = \frac{1}{\sqrt{N}} \sum_{y=0}^{N-1}\omega_N^{xy} \vert y \rangle \\
# & = \frac{1}{\sqrt{N}} \sum_{y=0}^{N-1} e^{2 \pi i xy / 2^n} \vert y \rangle \:\text{since}\: \omega_N^{xy} = e^{2\pi i \frac{xy}{N}} \:\text{and}\: N = 2^n\\
# & = \frac{1}{\sqrt{N}} \sum_{y=0}^{N-1} e^{2 \pi i \left(\sum_{k=1}^n y_k/2^k\right) x} \vert y_1 ... y_n \rangle \:\text{rewriting in fractional binary notation}\: y = y_1...y_k, y/2^n = \sum_{k=1}^n y_k/2^k \\
# & = \frac{1}{\sqrt{N}} \sum_{y=0}^{N-1} \prod_{k=0}^n e^{2 \pi i x y_k/2^k } \vert y_1 ... y_n \rangle \:\text{after expanding the exponential of a sum to a product of exponentials} \\
# & = \frac{1}{\sqrt{N}} \bigotimes_{k=1}^n \left(\vert0\rangle + e^{2 \pi i x /2^k } \vert1\rangle \right) \:\text{after rearranging the sum and products, and expanding} \\
# & = \frac{1}{\sqrt{N}} \left(\vert0\rangle + e^{2 \pi i[0.x_n]} \vert1\rangle\right) \otimes...\otimes \left(\vert0\rangle + e^{2 \pi i[0.x_1.x_2...x_{n-1}.x_n]} \vert1\rangle\right) \:\text{as}\: e^{2 \pi i x/2^k} = e^{2 \pi i[0.x_k...x_n]}
# \end{aligned}
#
# This is a very useful form of the QFT for $N=2^n$ as only the last qubit depends on the the
# values of all the other input qubits and each further bit depends less and less on the input qubits. Furthermore, note that $e^{2 \pi i.0.x_n}$ is either $+1$ or $-1$, which resembles the Hadamard transform.
#
# Before we create the circuit code for general $N=2^n$, let's look at $N=8,n=3$:
# $$QFT_8\vert x_1x_2x_3\rangle = \frac{1}{\sqrt{8}} \left(\vert0\rangle + e^{2 \pi i[0.x_3]} \vert1\rangle\right) \otimes \left(\vert0\rangle + e^{2 \pi i[0.x_2.x_3]} \vert1\rangle\right) \otimes \left(\vert0\rangle + e^{2 \pi i[0.x_1.x_2.x_3]} \vert1\rangle\right) $$
#
# The steps to creating the circuit for $\vert y_1y_2x_3\rangle = QFT_8\vert x_1x_2x_3\rangle$, remembering the [controlled phase rotation gate](../tools/quantum_gates_and_linear_algebra.ipynb
# ) $CU_1$, would be:
# 1. Apply a Hadamard to $\vert x_3 \rangle$, giving the state $\frac{1}{\sqrt{2}}\left(\vert0\rangle + e^{2 \pi i.0.x_3} \vert1\rangle\right) = \frac{1}{\sqrt{2}}\left(\vert0\rangle + (-1)^{x_3} \vert1\rangle\right)$
# 2. Apply a Hadamard to $\vert x_2 \rangle$, then depending on $k_3$ (before the Hadamard gate) a $CU_1(\frac{\pi}{2})$, giving the state $\frac{1}{\sqrt{2}}\left(\vert0\rangle + e^{2 \pi i[0.x_2.x_3]} \vert1\rangle\right)$.
# 3. Apply a Hadamard to $\vert x_1 \rangle$, then $CU_1(\frac{\pi}{2})$ depending on $k_2$, and $CU_1(\frac{\pi}{4})$ depending on $k_3$.
# 4. Measure the bits in reverse order, that is $y_3 = x_1, y_2 = x_2, y_1 = y_3$.
#
# In Qiskit, this is:
# ```
# q3 = QuantumRegister(3, 'q3')
# c3 = ClassicalRegister(3, 'c3')
#
# qft3 = QuantumCircuit(q3, c3)
# qft3.h(q[0])
# qft3.cu1(math.pi/2.0, q3[1], q3[0])
# qft3.h(q[1])
# qft3.cu1(math.pi/4.0, q3[2], q3[0])
# qft3.cu1(math.pi/2.0, q3[2], q3[1])
# qft3.h(q[2])
# ```
#
# For $N=2^n$, this can be generalised, as in the `qft` function in [tools.qi](https://github.com/Q/qiskit-terra/blob/master/qiskit/tools/qi/qi.py):
# ```
# def qft(circ, q, n):
# """n-qubit QFT on q in circ."""
# for j in range(n):
# for k in range(j):
# circ.cu1(math.pi/float(2**(j-k)), q[j], q[k])
# circ.h(q[j])
# ```
# #### Example
#
# Let's factorize $N = 21$ with coprime $x=2$, following the [above steps](#stepstwo) of the quantum period finding algorithm, which should return $r = 6$. This example follows one from [this](https://arxiv.org/abs/quant-ph/0303175) tutorial.
#
# 1. Choose $T = 2^t$ such that $N^2 \leq T \le 2N^2$. For $N = 21$, the smallest value of $t$ is 9, meaning $T = 2^t = 512$. Initialise two registers of qubits, first an argument register with $t = 9$ qubits, and second a function register with $n = log_2 N = 5$ qubits:
# $$\vert\psi_0\rangle = \vert 0 \rangle \vert 0 \rangle$$
#
# 2. Apply a Hadamard gate on each of the qubits in the argument register:
# $$\vert\psi_1\rangle = \frac{1}{\sqrt{T}}\sum_{a=0}^{T-1}\vert a \rangle \vert 0 \rangle = \frac{1}{\sqrt{512}}\sum_{a=0}^{511}\vert a \rangle \vert 0 \rangle$$
#
# 3. Implement the modular exponentiation function $x^a \bmod N$ on the function register:
# \begin{eqnarray}
# \vert\psi_2\rangle
# & = & \frac{1}{\sqrt{T}}\sum_{a=0}^{T-1}\vert a \rangle \vert x^a \bmod N \rangle
# = \frac{1}{\sqrt{512}}\sum_{a=0}^{511}\vert a \rangle \vert 2^a \bmod 21 \rangle \\
# & = & \frac{1}{\sqrt{512}} \bigg( \;\; \vert 0 \rangle \vert 1 \rangle + \vert 1 \rangle \vert 2 \rangle +
# \vert 2 \rangle \vert 4 \rangle + \vert 3 \rangle \vert 8 \rangle + \;\; \vert 4 \rangle \vert 16 \rangle + \;\,
# \vert 5 \rangle \vert 11 \rangle \, + \\
# & & \;\;\;\;\;\;\;\;\;\;\;\;\;\, \vert 6 \rangle \vert 1 \rangle + \vert 7 \rangle \vert 2 \rangle + \vert 8 \rangle \vert 4 \rangle + \vert 9 \rangle \vert 8 \rangle + \vert 10 \rangle \vert 16 \rangle + \vert 11 \rangle \vert 11 \rangle \, +\\
# & & \;\;\;\;\;\;\;\;\;\;\;\;\, \vert 12 \rangle \vert 1 \rangle + \ldots \bigg)\\
# \end{eqnarray}
# Notice that the above expression has the following pattern: the states of the second register of each “column” are the same. Therefore we can rearrange the terms in order to collect the second register:
# \begin{eqnarray}
# \vert\psi_2\rangle
# & = & \frac{1}{\sqrt{512}} \bigg[ \big(\,\vert 0 \rangle + \;\vert 6 \rangle + \vert 12 \rangle \ldots + \vert 504 \rangle + \vert 510 \rangle \big) \, \vert 1 \rangle \, + \\
# & & \;\;\;\;\;\;\;\;\;\;\; \big(\,\vert 1 \rangle + \;\vert 7 \rangle + \vert 13 \rangle \ldots + \vert 505 \rangle + \vert 511 \rangle \big) \, \vert 2 \rangle \, + \\
# & & \;\;\;\;\;\;\;\;\;\;\; \big(\,\vert 2 \rangle + \;\vert 8 \rangle + \vert 14 \rangle \ldots + \vert 506 \rangle + \big) \, \vert 4 \rangle \, + \\
# & & \;\;\;\;\;\;\;\;\;\;\; \big(\,\vert 3 \rangle + \;\vert 9 \rangle + \vert 15 \rangle \ldots + \vert 507 \rangle + \big) \, \vert 8 \rangle \, + \\
# & & \;\;\;\;\;\;\;\;\;\;\; \big(\,\vert 4 \rangle + \vert 10 \rangle + \vert 16 \rangle \ldots + \vert 508 \rangle + \big) \vert 16 \rangle \, + \\
# & & \;\;\;\;\;\;\;\;\;\;\; \big(\,\vert 5 \rangle + \vert 11 \rangle + \vert 17 \rangle \ldots + \vert 509 \rangle + \big) \vert 11 \rangle \, \bigg]\\
# \end{eqnarray}
#
# 4. To simplify following equations, we'll measure the function register before performing a quantum Fourier transform on the argument register. This will yield one of the following numbers with equal probability: $\{1,2,4,6,8,16,11\}$. Suppose that the result of the measurement was $2$, then:
# $$\vert\psi_3\rangle = \frac{1}{\sqrt{86}}(\vert 1 \rangle + \;\vert 7 \rangle + \vert 13 \rangle \ldots + \vert 505 \rangle + \vert 511 \rangle)\, \vert 2 \rangle $$
# It does not matter what is the result of the measurement; what matters is the periodic pattern. The period of the states of the first register is the solution to the problem and the quantum Fourier transform can reveal the value of the period.
#
# 5. Perform a quantum Fourier transform on the argument register:
# $$
# \vert\psi_4\rangle
# = QFT(\vert\psi_3\rangle)
# = QFT(\frac{1}{\sqrt{86}}\sum_{a=0}^{85}\vert 6a+1 \rangle)\vert 2 \rangle
# = \frac{1}{\sqrt{512}}\sum_{j=0}^{511}\bigg(\big[ \frac{1}{\sqrt{86}}\sum_{a=0}^{85} e^{-2 \pi i \frac{6ja}{512}} \big] e^{-2\pi i\frac{j}{512}}\vert j \rangle \bigg)\vert 2 \rangle
# $$
#
# 6. Measure the argument register. The probability of measuring a result $j$ is:
# $$ \rm{Probability}(j) = \frac{1}{512 \times 86} \bigg\vert \sum_{a=0}^{85}e^{-2 \pi i \frac{6ja}{512}} \bigg\vert^2$$
# This peaks at $j=0,85,171,256,341,427$. Suppose that the result of the measement yielded $j = 85$, then using continued fraction approximation of $\frac{512}{85}$, we obtain $r=6$, as expected.
# ## Implementation <a id='implementation'></a>
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister
from qiskit import execute, register, get_backend, compile
from qiskit.tools.visualization import plot_histogram, circuit_drawer
# As mentioned [earlier](#shorsalgorithm), many of the experimental demonstrations of Shor's algorithm rely on significant optimisations based on apriori knowledge of the expected results. We will follow the formulation in [this](http://science.sciencemag.org/content/351/6277/1068) paper, which demonstrates a reasonably scalable realisation of Shor's algorithm using $N = 15$. Below is the first figure from the paper, showing various quantum circuits, with the following caption: _Diagrams of Shor’s algorithm for factoring $N = 15$, using a generic textbook approach (**A**) compared with Kitaev’s approach (**B**) for a generic base $a$. (**C**) The actual implementation for factoring $15$ to base $11$, optimized for the corresponding single-input state. Here $q_i$ corresponds to the respective qubit in the computational register. (**D**) Kitaev’s approach to Shor’s algorithm for the bases ${2, 7, 8, 13}$. Here, the optimized map of the first multiplier is identical in all four cases, and the last multiplier is implemented with full modular multipliers, as depicted in (**E**). In all cases, the single QFT qubit is used three times, which, together with the four qubits in the computation register, totals seven effective qubits. (**E**) Circuit diagrams of the modular multipliers of the form $a \bmod N$ for bases $a = {2, 7, 8, 11, 13}$._
#
# <img src="../../images/shoralgorithm.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="center">
#
# Note that we cannot run this version of Shor's algorithm on an IBM Quantum Experience device at the moment as we currently lack the ability to do measurement feedforward and qubit resetting. Thus we'll just be building the ciruits to run on the simulators for now. Based on <NAME> & <NAME>'s implementation, found [here](https://github.com/amitabhyadav/Shor-Algorithm-on-IBM-Quantum-Experience)
# First we'll construct the $a^1 \bmod 15$ circuits for $a = 2,7,8,11,13$ as in **E**:
# qc = quantum circuit, qr = quantum register, cr = classical register, a = 2, 7, 8, 11 or 13
def circuit_amod15(qc,qr,cr,a):
if a == 2:
qc.cswap(qr[4],qr[3],qr[2])
qc.cswap(qr[4],qr[2],qr[1])
qc.cswap(qr[4],qr[1],qr[0])
elif a == 7:
qc.cswap(qr[4],qr[1],qr[0])
qc.cswap(qr[4],qr[2],qr[1])
qc.cswap(qr[4],qr[3],qr[2])
qc.cx(qr[4],qr[3])
qc.cx(qr[4],qr[2])
qc.cx(qr[4],qr[1])
qc.cx(qr[4],qr[0])
elif a == 8:
qc.cswap(qr[4],qr[1],qr[0])
qc.cswap(qr[4],qr[2],qr[1])
qc.cswap(qr[4],qr[3],qr[2])
elif a == 11: # this is included for completeness
qc.cswap(qr[4],qr[2],qr[0])
qc.cswap(qr[4],qr[3],qr[1])
qc.cx(qr[4],qr[3])
qc.cx(qr[4],qr[2])
qc.cx(qr[4],qr[1])
qc.cx(qr[4],qr[0])
elif a == 13:
qc.cswap(qr[4],qr[3],qr[2])
qc.cswap(qr[4],qr[2],qr[1])
qc.cswap(qr[4],qr[1],qr[0])
qc.cx(qr[4],qr[3])
qc.cx(qr[4],qr[2])
qc.cx(qr[4],qr[1])
qc.cx(qr[4],qr[0])
# Next we'll build the rest of the period finding circuit as in **D**:
# qc = quantum circuit, qr = quantum register, cr = classical register, a = 2, 7, 8, 11 or 13
def circuit_aperiod15(qc,qr,cr,a):
if a == 11:
circuit_11period15(qc,qr,cr)
return
# Initialize q[0] to |1>
qc.x(qr[0])
# Apply a**4 mod 15
qc.h(qr[4])
# controlled identity on the remaining 4 qubits, which is equivalent to doing nothing
qc.h(qr[4])
# measure
qc.measure(qr[4],cr[0])
# reinitialise q[4] to |0>
qc.reset(qr[4])
# Apply a**2 mod 15
qc.h(qr[4])
# controlled unitary
qc.cx(qr[4],qr[2])
qc.cx(qr[4],qr[0])
# feed forward
if cr[0] == 1:
qc.u1(math.pi/2.,qr[4])
qc.h(qr[4])
# measure
qc.measure(qr[4],cr[1])
# reinitialise q[4] to |0>
qc.reset(qr[4])
# Apply a mod 15
qc.h(qr[4])
# controlled unitary.
circuit_amod15(qc,qr,cr,a)
# feed forward
if cr[1] == 1:
qc.u1(math.pi/2.,qr[4])
if cr[0] == 1:
qc.u1(math.pi/4.,qr[4])
qc.h(qr[4])
# measure
qc.measure(qr[4],cr[2])
# Next we build the optimised circuit for $11 \bmod 15$ as in **C**.
def circuit_11period15(qc,qr,cr):
# Initialize q[0] to |1>
qc.x(qr[0])
# Apply a**4 mod 15
qc.h(qr[4])
# controlled identity on the remaining 4 qubits, which is equivalent to doing nothing
qc.h(qr[4])
# measure
qc.measure(qr[4],cr[0])
# reinitialise q[4] to |0>
qc.reset(qr[4])
# Apply a**2 mod 15
qc.h(qr[4])
# controlled identity on the remaining 4 qubits, which is equivalent to doing nothing
# feed forward
if cr[0] == 1:
qc.u1(math.pi/2.,qr[4])
qc.h(qr[4])
# measure
qc.measure(qr[4],cr[1])
# reinitialise q[4] to |0>
qc.reset(qr[4])
# Apply 11 mod 15
qc.h(qr[4])
# controlled unitary.
qc.cx(qr[4],qr[3])
qc.cx(qr[4],qr[1])
# feed forward
if cr[1] == 1:
qc.u1(math.pi/2.,qr[4])
if cr[0] == 1:
qc.u1(math.pi/4.,qr[4])
qc.h(qr[4])
# measure
qc.measure(qr[4],cr[2])
# Let's build and run a circuit for $a = 7$, and plot the results:
# +
q = QuantumRegister(5, 'q')
c = ClassicalRegister(5, 'c')
shor = QuantumCircuit(q, c)
circuit_aperiod15(shor,q,c,7)
sim_job = execute([shor], 'local_qasm_simulator')
sim_result = sim_job.result()
sim_data = sim_result.get_counts(shor)
plot_histogram(sim_data)
# -
# We see here that the period, $r = 4$, and thus calculate the factors $p = \text{gcd}(a^{r/2}+1,15) = 3$ and $q = \text{gcd}(a^{r/2}-1,15) = 5$. Why don't you try seeing what you get for $a = 2, 8, 11, 13$?
| reference/algorithms/shor_algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# (image-segmentation:voronoi-otsu-labeling=)
# # Voronoi-Otsu-labeling
# This workflow for image segmentation is a rather simple and yet powerful approach, e.g. for detecting and segmenting nuclei in fluorescence micropscopy images. A nuclei marker such as nuclei-GFP, DAPI or histone-RFP in combination with various microscopy techniques can be used to generate images of suitable kind.
from skimage.io import imread, imshow
import matplotlib.pyplot as plt
import pyclesperanto_prototype as cle
import napari_segment_blobs_and_things_with_membranes as nsbatwm
# To demonstrate the workflow, we're using image data from the [Broad Bio Image Challenge](https://bbbc.broadinstitute.org/BBBC022):
# We used image set BBBC022v1 [Gustafsdottir et al., PLOS ONE, 2013](http://dx.doi.org/10.1371/journal.pone.0080999), available from the Broad Bioimage Benchmark Collection [Ljosa et al., Nature Methods, 2012](http://dx.doi.org/10.1038/nmeth.2083).
# +
input_image = imread("../../data/BBBC022/IXMtest_A02_s9.tif")[:,:,0]
input_crop = input_image[0:200, 200:400]
fig, axs = plt.subplots(1, 2, figsize=(15, 15))
cle.imshow(input_image, plot=axs[0])
cle.imshow(input_crop, plot=axs[1])
# -
# ## Applying the algorithm
# Voronoi-Otsu-labeling is a command in clesperanto, which asks for two sigma parameters. The first sigma controls how close detected cells can be (`spot_sigma`) and second controls how precise segmented objects are outlined (`outline_sigma`).
# +
sigma_spot_detection = 5
sigma_outline = 1
segmented = cle.voronoi_otsu_labeling(input_image, spot_sigma=sigma_spot_detection, outline_sigma=sigma_outline)
segmented_crop = segmented[0:200, 200:400]
fig, axs = plt.subplots(1, 2, figsize=(15, 15))
cle.imshow(segmented, labels=True, plot=axs[0])
cle.imshow(segmented_crop, labels=True, plot=axs[1])
# -
# ## How does it work?
# The Voronoi-Otsu-Labeling workflow is a combination of Gaussian blur, spot detection, thresholding and binary watershed. The interested reader might want to see the [open source code](https://github.com/clEsperanto/pyclesperanto_prototype/blob/master/pyclesperanto_prototype/_tier9/_voronoi_otsu_labeling.py). The approach is similar to applying a seeded watershed to a binary image, e.g. in [MorphoLibJ](https://imagej.net/plugins/marker-controlled-watershed) or [scikit-image](https://scikit-image.org/docs/dev/api/skimage.segmentation.html#skimage.segmentation.watershed). However, the seeds are computed automatically and cannot be passed.
#
# For demonstration purposes we do that only on the 2D cropped image shown above. If this algorithm is applied to 3D data, it is recommended to make it isotropic first.
image_to_segment = input_crop
print(image_to_segment.shape)
# As a first step, we blur the image with a given sigma and detect maxima in the resulting image.
# +
blurred = cle.gaussian_blur(image_to_segment, sigma_x=sigma_spot_detection, sigma_y=sigma_spot_detection, sigma_z=sigma_spot_detection)
detected_spots = cle.detect_maxima_box(blurred, radius_x=0, radius_y=0, radius_z=0)
number_of_spots = cle.sum_of_all_pixels(detected_spots)
print("number of detected spots", number_of_spots)
fig, axs = plt.subplots(1, 2, figsize=(15, 15))
cle.imshow(blurred, plot=axs[0])
cle.imshow(detected_spots, plot=axs[1])
# -
# Furthermore, we start again from the cropped image and blur it again, with a different sigma. Afterwards, we threshold the image using [Otsu's thresholding method (Otsu et al 1979)](https://doi.org/10.1109%2FTSMC.1979.4310076).
# +
blurred = cle.gaussian_blur(image_to_segment, sigma_x=sigma_outline, sigma_y=sigma_outline, sigma_z=sigma_outline)
binary = cle.threshold_otsu(blurred)
fig, axs = plt.subplots(1, 2, figsize=(15, 15))
cle.imshow(blurred, plot=axs[0])
cle.imshow(binary, plot=axs[1])
# -
# Afterwards, we take the binary spots image and the binary segmentation image and apply a `binary_and` operation to exclude spots which were detected in the background area. Those likely corresponded to noise.
# +
selected_spots = cle.binary_and(binary, detected_spots)
number_of_spots = cle.sum_of_all_pixels(selected_spots)
print("number of selected spots", number_of_spots)
fig, axs = plt.subplots(1, 3, figsize=(15, 15))
cle.imshow(detected_spots, plot=axs[0])
cle.imshow(binary, plot=axs[1])
cle.imshow(selected_spots, plot=axs[2])
# -
# Next, we separate the image space between the selected spots using a [Voronoi diagram](https://en.wikipedia.org/wiki/Voronoi_diagram#References) which is limited to the positive pixels in the binary image.
# +
voronoi_diagram = cle.masked_voronoi_labeling(selected_spots, binary)
fig, axs = plt.subplots(1, 3, figsize=(15, 15))
cle.imshow(selected_spots, plot=axs[0])
cle.imshow(binary, plot=axs[1])
cle.imshow(voronoi_diagram, labels=True, plot=axs[2])
# -
# ## Other Voronoi-Otsu-Labeling implementations
#
# There is an alternative implementation of the algorithm in the scriptable napari plugin [napari-segment-blobs-and-things-with-membranes](https://github.com/haesleinhuepf/napari-segment-blobs-and-things-with-membranes).
# The code here is almost identical to the code above. The major difference is that we call `nsbatwm.voronoi_otsu_labeling()` instead of `cle.voronoi_otsu_labeling()`.
# +
sigma_spot_detection = 5
sigma_outline = 1
segmented2 = nsbatwm.voronoi_otsu_labeling(input_image, spot_sigma=sigma_spot_detection, outline_sigma=sigma_outline)
segmented_crop2 = segmented2[0:200, 200:400]
fig, axs = plt.subplots(1, 2, figsize=(15, 15))
cle.imshow(segmented2, labels=True, plot=axs[0])
cle.imshow(segmented_crop2, labels=True, plot=axs[1])
| docs/20_image_segmentation/11_voronoi_otsu_labeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Unattacked Queens
# ===
# The properties of chess pieces play a part in many challenges, including in a group of problems about unattacked queens. Imagine three white queens and five black queens on a 5 × 5 chessboard. Can you arrange them so that no queen of one color can attack a queen of the other color? There is only one solution, excluding reflections and rotations.
# http://www.scientificamerican.com/article/martin-gardner-fans-try-these-mathematical-games/
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.table import Table
from copy import deepcopy
# %matplotlib inline
# +
fig, ax = plt.subplots()
min_val, max_val, diff = 0., 10., 1.
#imshow portion
N_points = (max_val - min_val) / diff
imshow_data = np.random.rand(N_points, N_points)
ax.imshow(imshow_data, interpolation='nearest')
#text portion
ind_array = np.arange(min_val, max_val, diff)
x, y = np.meshgrid(ind_array, ind_array)
for x_val, y_val in zip(x.flatten(), y.flatten()):
c = 'x' if (x_val + y_val)%2 else 'o'
ax.text(x_val, y_val, c, va='center', ha='center')
#set tick marks for grid
ax.set_xticks(np.arange(min_val-diff/2, max_val-diff/2))
ax.set_yticks(np.arange(min_val-diff/2, max_val-diff/2))
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.set_xlim(min_val-diff/2, max_val-diff/2)
ax.set_ylim(min_val-diff/2, max_val-diff/2)
ax.grid()
plt.show()
# -
N = 5
#board = [[(i*N+j)%2 for j in range(N)] for i in range(N)]
board = [[0 for j in range(N)] for i in range(N)]
temp_board = deepcopy(board)
plt.matshow(board, interpolation='nearest', cmap=plt.get_cmap("gray"))
plt.grid()
plt.show()
def addQueen(board, pos, val=-2):
x, y = pos
board[x][y] = val
val = (abs(val)+1)*val/abs(val)
for i in range(1,N):
board[(x+i)%N][y] = val
board[x][(y+i)%N] = val
for j in [1, -1]:
for k in [1,-1]:
if (x+i*j) >= 0 and (x+i*j) < N and (y+i*k) >= 0 and (y+i*k) < N:
board[(x+i*j)][(y+i*k)] = val
plt.matshow(board, interpolation='nearest', cmap=plt.get_cmap("Set3"), vmin=-3,vmax=3)
plt.grid()
plt.show()
return board
board = deepcopy(temp_board)
addQueen(board, (0,0))
addQueen(board, (N-2,N-1), val = 2)
plt.imshow(temp_board, interpolation='nearest', cmap=plt.get_cmap("gray"))
| Unattacked Queens.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="SNjbzn4bXtjU" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# !kill -9 -1
# + id="FKVmEEFNW0SY" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 36} outputId="b742fa1d-e886-4959-b934-037be1fe7b49" executionInfo={"status": "ok", "timestamp": 1522007750832, "user_tz": -60, "elapsed": 828, "user": {"displayName": "niren52", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "118152712183165316846"}}
# !ls
# + id="RxUQHoqmXw5E" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
import matplotlib.pyplot as plt
import pandas as pd
import plotly.plotly as py
import numpy as np
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# + id="L4xtcEEoZ4B0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Code to read csv file into colaboratory:
# !pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# 1. Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
#2. Get the file
downloadedTest = drive.CreateFile({'id':'1rGfRRaEqKAT0EwyF8uZRTDbfVBDajiE_'}) # replace the id with id of file you want to access
downloadedTest.GetContentFile('test.csv')
downloadedTrain = drive.CreateFile({'id':'14rKrVfwK_Q0YeDnCknCF9N5Gz-m5502U'}) # replace the id with id of file you want to access
downloadedTrain.GetContentFile('train.csv')
downloadedValidation = drive.CreateFile({'id':'16kjxp9JOgUVExDHlcTGGS4eGjoYP6m0N'}) # replace the id with id of file you want to access
downloadedValidation.GetContentFile('validation.csv')
#3. Read file as panda dataframe
test = pd.read_csv('test.csv')
train = pd.read_csv('train.csv')
validation = pd.read_csv('validation.csv')
# + id="L3VIEiJwjprC" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 583} outputId="e19878e0-677e-4389-bfdc-dffed97c900e" executionInfo={"status": "ok", "timestamp": 1522007836880, "user_tz": -60, "elapsed": 482, "user": {"displayName": "niren52", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "118152712183165316846"}}
train.info()
# + id="n-AuIaDWZ7gA" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
impressions = pd.value_counts(train['advertiser'])
# + id="f0VN4T_Kaas2" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 206} outputId="0c96903d-80c7-4397-f7dd-09c3dcb8022d" executionInfo={"status": "ok", "timestamp": 1522007852846, "user_tz": -60, "elapsed": 460, "user": {"displayName": "niren52", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "118152712183165316846"}}
impressions
# + id="TLlHQvrneD98" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
trainGroup = train.groupby('advertiser')
trainGroup2 = trainGroup[['click','payprice']]
# + id="EikBox_EahdG" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 363} outputId="9604e800-5b23-4f4b-a3ce-48e39a1585a6" executionInfo={"status": "ok", "timestamp": 1522008061702, "user_tz": -60, "elapsed": 442, "user": {"displayName": "niren52", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "118152712183165316846"}}
validationGroup2.sum()
# + id="2hurIwwfbIL8" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 138} outputId="c23af12d-ef45-4f30-9013-08fc2112dd6f" executionInfo={"status": "error", "timestamp": 1522005606332, "user_tz": -60, "elapsed": 316, "user": {"displayName": "niren52", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "118152712183165316846"}}
#Exported to Excel
PROBLEM WITH PANDAS STRUCTURES WHEN MERGING
# + id="O4oYbbRmh0y8" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# + id="U7ZWqgdOllvq" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
| EXPLORATORY STATISTICS/EXPLORATORY (TRAINING).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Packaging
#
# Once we've made a working program, we'd like to be able to share it with others.
#
# A good cross-platform build tool is the most important thing: you can always
# have collaborators build from source.
#
# ### Distribution tools
# Distribution tools allow one to obtain a working copy of someone else's package.
#
# - Language-specific tools:
# - python: PyPI,
# - ruby: Ruby Gems,
# - perl: CPAN,
# - R: CRAN
#
# - Platform specific packagers e.g.:
# - `brew` for MacOS,
# - `apt`/`yum` for Linux or
# - [`choco`](https://chocolatey.org/) for Windows.
# ### Laying out a project
#
# When planning to package a project for distribution, defining a suitable
# project layout is essential.
#
#
#
# + language="bash"
# tree --charset ascii greetings -I "doc|build|Greetings.egg-info|dist|*.pyc"
# -
# We can start by making our directory structure. You can create many nested directories at once using the `-p` switch on `mkdir`.
# + language="bash"
# mkdir -p greetings/greetings/test/fixtures
# mkdir -p greetings/scripts
# -
# ### Using setuptools
#
# To make python code into a package, we have to write a `setup.py` file:
# ```python
# from setuptools import setup, find_packages
#
# setup(
# name="Greetings",
# version="0.1.0",
# packages=find_packages(exclude=['*test']),
# )
# ```
# We can now install this code with
# ```
# pip install .
# ```
#
# And the package will be then available to use everywhere on the system.
#
from greetings.greeter import greet
greet("Terry","Gilliam")
# ### Convert the script to a module
#
# Of course, there's more to do when taking code from a quick script and turning it into a proper module:
# We need to add docstrings to our functions, so people can know how to use them.
# %pycat greetings/greetings/greeter.py
import greetings
help(greetings.greeter.greet)
# The documentation string explains how to use the function; don't worry about this for now, we'll consider
# this on [the next notebook](./04documentation.ipynb).
# ### Write an executable script
#
#
#
#
#
#
# %pycat greetings/greetings/command.py
#
#
#
# ### Specify dependencies
# We use the setup.py file to specify the packages we depend on:
# ```python
# setup(
# name="Greetings",
# version="0.1.0",
# packages=find_packages(exclude=['*test']),
# install_requires=['argparse']
# )
# ```
# ### Specify entry point
# This allows us to create a command to execute part of our library. In this case when we execute `greet` on the terminal, we will be calling the `process` function under `greetings/command.py`.
#
# %pycat greetings/setup.py
#
# And the scripts are now available as command line commands:
#
#
#
# + language="bash"
# greet --help
# + language="bash"
# greet <NAME>
# greet --polite <NAME>
# greet <NAME> --title Cartoonist
# -
# ### Installing from GitHub
#
# We could now submit "greeter" to PyPI for approval, so everyone could `pip install` it.
#
# However, when using git, we don't even need to do that: we can install directly from any git URL:
#
# ```
# pip install git+git://github.com/jamespjh/greeter
# ```
# + language="bash"
# greet Lancelot the-Brave --title Sir
# -
#
#
# ### Write a readme file
# e.g.:
# +
# %%writefile greetings/README.md
Greetings!
==========
This is a very simple example package used as part of the UCL
[Research Software Engineering with Python](development.rc.ucl.ac.uk/training/engineering) course.
Usage:
Invoke the tool with `greet <FirstName> <Secondname>`
# -
# ### Write a license file
# e.g.:
# +
# %%writefile greetings/LICENSE.md
(C) University College London 2014
This "greetings" example package is granted into the public domain.
# -
# ### Write a citation file
# e.g.:
# +
# %%writefile greetings/CITATION.md
If you wish to refer to this course, please cite the URL
http://github-pages.ucl.ac.uk/rsd-engineeringcourse/
Portions of the material are taken from [Software Carpentry](http://software-carpentry.org/)
# -
# You may well want to formalise this using the [codemeta.json](https://codemeta.github.io/) standard or the [citation file format](http://citation-file-format.github.io/) - these don't have wide adoption yet, but we recommend it.
# ### Define packages and executables
# + language="bash"
# touch greetings/greetings/test/__init__.py
# touch greetings/greetings/__init__.py
# -
# ### Write some unit tests
#
# Separating the script from the logical module made this possible:
#
#
#
#
#
#
# +
# %%writefile greetings/greetings/test/test_greeter.py
import yaml
import os
from ..greeter import greet
def test_greeter():
with open(os.path.join(os.path.dirname(__file__),
'fixtures',
'samples.yaml')) as fixtures_file:
fixtures = yaml.load(fixtures_file)
for fixture in fixtures:
answer = fixture.pop('answer')
assert greet(**fixture) == answer
# -
#
#
#
# Add a fixtures file:
#
#
#
#
#
#
# %%writefile greetings/greetings/test/fixtures/samples.yaml
- personal: Eric
family: Idle
answer: "Hey, <NAME>."
- personal: Graham
family: Chapman
polite: True
answer: "How do you do, <NAME>."
- personal: Michael
family: Palin
title: CBE
answer: "Hey, CBE <NAME>."
# + language="bash"
# pytest
# -
# However, this hasn't told us that also the third test is wrong! A better aproach is to parametrize the test as follows:
# +
# %%writefile greetings/greetings/test/test_greeter.py
import yaml
import os
import pytest
from ..greeter import greet
def read_fixture():
with open(os.path.join(os.path.dirname(__file__),
'fixtures',
'samples.yaml')) as fixtures_file:
fixtures = yaml.load(fixtures_file)
return fixtures
@pytest.mark.parametrize("fixture", read_fixture())
def test_greeter(fixture):
answer = fixture.pop('answer')
assert greet(**fixture) == answer
# -
# Now when we run `pytest`, we get a failure per element in our fixture and we know all that fails.
# + language="bash"
# pytest
# -
# ### Developer Install
#
# If you modify your source files, you would now find it appeared as if the program doesn't change.
#
# That's because pip install **copies** the files.
#
# If you want to install a package, but keep working on it, you can do:
# ```
# pip install --editable .
# ```
# ### Distributing compiled code
#
# If you're working in C++ or Fortran, there is no language specific repository.
# You'll need to write platform installers for as many platforms as you want to
# support.
#
# Typically:
#
# * `dpkg` for `apt-get` on Ubuntu and Debian
# * `rpm` for `yum`/`dnf` on Redhat and Fedora
# * `homebrew` on OSX (Possibly `macports` as well)
# * An executable `msi` installer for Windows.
#
# #### Homebrew
#
# Homebrew: A ruby DSL, you host off your own webpage
#
# See an [installer for the cppcourse example](http://github.com/jamespjh/homebrew-reactor)
#
# If you're on OSX, do:
#
# ```
# brew tap jamespjh/homebrew-reactor
# brew install reactor
# ```
| ch04packaging/03Packaging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="fU0Qi60TN2N6"
# Baseado no exemplo: https://www.kaggle.com/thebrownviking20/intro-to-recurrent-neural-networks-lstm-gru
# + colab={} colab_type="code" id="w_-ug3lgN2OC" outputId="0f093dd1-82e3-407c-f802-ea9f920ec4fa"
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from keras.layers import SimpleRNN
from keras.models import Sequential
from numpy.random import seed
#from tensorflow import set_random_seed
# + colab={} colab_type="code" id="guShsCZpN2OJ"
seed(42)
#set_random_seed(42)
plt.rcParams['figure.figsize'] = 16,4
# + colab={} colab_type="code" id="iiEq6m2lN2OM"
SPLIT_DATE = '2015/01/01'
WINDOW_SIZE = 30
# -
# # Importar Bases
import os.path
def path_base(base_name,folder):
current_dir = os.path.abspath(os.path.join(os.getcwd()))
print(current_dir)
data_dir = current_dir.replace('notebook',folder)
print(data_dir)
data_base = data_dir + '\\' + base_name
print(data_base)
return data_base
# + colab={} colab_type="code" id="iKFiNtmJN2OP" outputId="b0ee7046-83ce-4d46-9813-f2b2ce14bae7"
df = pd.read_csv(path_base('db_IBM_stocks.csv','data'),index_col='Date')
df.head(5)
# + colab={} colab_type="code" id="r1Bg5VyyN2OU" outputId="c158d083-2500-4315-df69-4b3e4fdef798"
df['Average'] = (df['High'] + df['Low'])/2
df = df[['Average']]
df.head(5)
# + colab={} colab_type="code" id="xJnQywVYN2OY" outputId="1144c7b6-ad64-4077-caf7-d2250cb56a78"
df.plot(legend=True)
plt.title('IBM stock price')
plt.show()
# + colab={} colab_type="code" id="sjDnF5pSN2Oc"
df_train = df[df.index < SPLIT_DATE].copy()
df_test = df[df.index >= SPLIT_DATE].copy()
SPLIT_DATE
# + colab={} colab_type="code" id="nKgyiI4WN2Of"
#scaler = MinMaxScaler()
#df_train['Average'] = scaler.fit_transform(df_train['Average'].values.reshape(-1, 1))
#df_test['Average'] = scaler.transform(df_test['Average'].values.reshape(-1, 1))
# + colab={} colab_type="code" id="Q_PlY59aN2Oi"
def gen_rnn_inputs(df, window_size):
X, y = [], []
averages = df['Average'].values
for i in range(window_size, len(df)):
X.append(averages[i-window_size: i])
y.append(averages[i])
return np.array(X), np.array(y)
# + colab={} colab_type="code" id="Mc_oajHkN2Ol"
X_train, y_train = gen_rnn_inputs(df_train, WINDOW_SIZE)
X_test, y_test = gen_rnn_inputs(df_test, WINDOW_SIZE)
# -
df_test
X_test
y_test
# + colab={} colab_type="code" id="XJkXo_C3N2Oo" outputId="46e227cd-ab6f-4170-db1a-03f4d9e97355"
model = Sequential()
model.add(SimpleRNN(1, activation='relu', input_shape=(WINDOW_SIZE, 1)))
model.compile(optimizer='adam',loss='mean_squared_error')
model.summary()
# + colab={} colab_type="code" id="hPhbIB8EN2Os" outputId="283adab5-f7fc-460c-a245-dd960989265a"
model.fit(np.expand_dims(X_train, axis=-1), y_train, epochs=100, batch_size=8, verbose=2)
# + colab={} colab_type="code" id="LPurNWDoN2Ox"
y_pred = model.predict(np.expand_dims(X_test, axis=-1))
np.expand_dims(X_test, axis=-1)
# + colab={} colab_type="code" id="jFLo7yj-N2O0"
df_test_preds = df_test.copy()
df_test_preds['Average'] = np.zeros(WINDOW_SIZE).tolist() + y_test.reshape(-1,1).squeeze().tolist()
df_test_preds['Pred'] = np.zeros(WINDOW_SIZE).tolist() + y_pred.reshape(-1,1).squeeze().tolist()
# + colab={} colab_type="code" id="7VtmKg-fN2O3" outputId="36138205-174f-42a1-848d-b015058767e9"
df_test_preds.plot(legend=True)
# -
| notebook/Keras_TimeSeries_RNN_ibm_stock_com.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# in the conversion to jpg or png, we're losing the image
# +
from custom_libraries import import_data
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
# multithreading and showing progress
import concurrent.futures
from tqdm import tqdm
# image processing
import pydicom
import cv2
# saving to cloud storage
from google.cloud import storage
from custom_libraries import gcloud_storage
# +
# train paths
train_csv_path = '../data/stage_1_train.csv'
train_image_folder_original = '../data/stage_1_train_images/'
train_image_folder_target = '../data/stage_1_train_images_jpg/'
# test paths
test_csv_path = '../data/stage_1_sample_submission.csv'
test_image_folder_original = '../data/stage_1_test_images/'
test_image_folder_target = '../data/stage_1_test_images_jpg/'
# +
def get_image_list(csv_path, image_folder_original, image_folder_target):
# load list from CSV
image_list = pd.read_csv(csv_path)
# drop 'Label' column
image_list.drop(columns=['Label'],inplace=True)
# standardize the ID to format ID_{number}
image_list.ID = image_list.ID.map(lambda id: 'ID_' + id.split('_')[1])
# drop duplicates keeping only unique IDs
image_list.drop_duplicates(subset='ID',inplace=True)
# reset index after dropping duplicated
image_list.reset_index(drop=True,inplace=True)
# add origin and target paths for each image
image_list['origin'] = image_list.ID.map(lambda id: image_folder_original + id + '.dcm')
image_list['target'] = image_list.ID.map(lambda id: image_folder_target + id + '.png')
return image_list
image_list = get_image_list(train_csv_path,train_image_folder_original,train_image_folder_target)
# -
# Some functions to extract the correct window-image. These have been adapted from https://www.kaggle.com/allunia/rsna-ih-detection-eda and references therin
# +
def window_image(img, window_center,window_width, intercept, slope, rescale=True):
img = (img*slope +intercept)
img_min = window_center - window_width//2
img_max = window_center + window_width//2
img[img<img_min] = img_min
img[img>img_max] = img_max
if rescale:
# Extra rescaling to 0-1, not in the original notebook
img = (img - img_min) / (img_max - img_min)
return img
def get_first_of_dicom_field_as_int(x):
#get x[0] as in int is x is a 'pydicom.multival.MultiValue', otherwise get int(x)
if type(x) == pydicom.multival.MultiValue:
return int(x[0])
else:
return int(x)
def get_windowing(data):
dicom_fields = [data[('0028','1050')].value, #window center
data[('0028','1051')].value, #window width
data[('0028','1052')].value, #intercept
data[('0028','1053')].value] #slope
return [get_first_of_dicom_field_as_int(x) for x in dicom_fields]
# -
def get_image_resized(img_path, dim = (512,512)):
"""Output windowed and resized jpg of dcm image"""
ds=pydicom.dcmread(img_path)
raw=ds.pixel_array
window_center , window_width, intercept, slope = get_windowing(ds)
windowed = window_image(raw, 50, 100, intercept, slope)
resized = cv2.resize(windowed, dim)
return resized
def get_image(img_path, dim = (512,512)):
"""Output three versions of image - raw, windowed and resized"""
ds=pydicom.dcmread(img_path)
raw=ds.pixel_array
window_center , window_width, intercept, slope = get_windowing(ds)
windowed = window_image(raw, 50, 100, intercept, slope)
resized = cv2.resize(windowed, dim)
return raw,windowed,resized
def display_image_processing(
image_paths = False,
n_images = 5,
random_state=12345,
bone_color = True,
**kwargs):
"""Display the versions of each image
Version:
1. raw
2. windowed
3. windowed and reduced
Keyword arguments:
image_paths (list of strings) - image paths
n_images (int) - number of different images to display
random_state (int) - random state for image selection
Output:
display (inline image) - images displayed inline
image_paths (list) - list of image paths
bone_color (bool) - if true, pass cmap=plt.cm.bone to plt.imshow()
"""
if not image_paths:
image_paths = list(image_list.origin.sample(
n=n_images,
replace=False,
random_state=12345))
# print(image_paths)
nrow = len(image_paths)
ncol = 3
fig = plt.figure(figsize=(15,5*nrow))
for row in range(0,nrow):
img_list = list( get_image(image_paths[row]) )
for sub_img in range(0,3):
fig.add_subplot(nrow,ncol,row*3+sub_img+1)
if bone_color:
plt.imshow(img_list[sub_img],cmap=plt.cm.bone,**kwargs)
else:
plt.imshow(img_list[sub_img],**kwargs)
# fig.add_subplot()
return image_paths
paths = display_image_processing(n_images=2, random_state=12, bone_color=False)
# +
# fig = plt.figure(figsize=(15,10))
# plt.imshow(get_image_resized(image_list.origin[350]),cmap=plt.cm.bone)
# plt.show()
# -
# +
def save_as_jpg_train(image_id,
image_folder_original ='../data/stage_1_train_images/',
image_folder_target='../data/stage_1_train_images_jpg/'):
img = get_image_resized(image_folder_original+image_id+'.dcm')
cv2.imwrite(image_folder_target+image_id+'.jpg',img)
def save_as_jpg_test(image_id,
image_folder_original ='../data/stage_1_test_images/',
image_folder_target='../data/stage_1_test_images_jpg/'):
img = get_image_resized(image_folder_original+image_id+'.dcm')
cv2.imwrite(image_folder_target+image_id+'.jpg',img)
def run_thread(f, iterable):
with concurrent.futures.ProcessPoolExecutor() as executor:
executor.map(f, iterable, chunksize=50)
# +
# convert all stage 1 train images with multithreading
# image_list = get_image_list(train_csv_path,train_image_folder_original,train_image_folder_target)
# image_list_ids = image_list['ID']
# run_thread(save_as_jpg_train, image_list_ids)
# print('stage 1 train complete')
# convert all stage 1 test images with multithreading
# image_list = get_image_list(test_csv_path,test_image_folder_original,test_image_folder_target)
# image_list_ids = image_list['ID']
# run_thread(save_as_jpg_test, image_list_ids)
# print('stage 1 test complete')
# -
"ID_"+paths[0].split('_')[-1].split('.')[0]
save_as_jpg_train("ID_"+paths[0].split('_')[-1].split('.')[0],
image_folder_original ='../data/stage_1_train_images/',
image_folder_target='test/')
plt.imshow(plt.imread('test/'+"ID_"+paths[0].split('_')[-1].split('.')[0]+".jpg"))
upload_blob("fi-capstone-data",
'test/'+"ID_"+paths[0].split('_')[-1].split('.')[0]+".jpg",
'test/'+"ID_"+paths[0].split('_')[-1].split('.')[0]+".jpg")
# [START storage_list_buckets]
def list_buckets():
"""Lists all buckets."""
storage_client = storage.Client()
buckets = storage_client.list_buckets()
for bucket in buckets:
print(bucket.name)
# [END storage_list_buckets]
list_buckets()
def upload_blob(bucket_name, source_file_name, destination_blob_name):
"""Uploads a file to the bucket."""
storage_client = storage.Client()
bucket = storage_client.get_bucket(bucket_name)
blob = bucket.blob(destination_blob_name)
blob.upload_from_filename(source_file_name)
print('File {} uploaded to {}.'.format(
source_file_name,
destination_blob_name))
upload_blob("fi-capstone-data","LICENSE","test/LICENSE")
# ## Finding images of a different size
def get_metadata(img_path):
ds=pydicom.dcmread(img_path)
try:
metadata_fields = [ds[('0028','0010')].value, #Rows
ds[('0028','0010')].value, #Columns
ds[('0028','0030')].value, #Pixel Spacing
ds[('0028','0100')].value, #Bits Allocated
ds[('0028','0101')].value, #Bits Stored
ds[('0028','0102')].value] #High Bits
raw=ds.pixel_array
raw = ds.pixel_array
metadata_fields.append(raw.shape[0])
metadata_fields.append(raw.shape[1])
except:
metadata_fields = [99,99,99,99,99,99,99,99]
return metadata_fields
# ### Search for issues in metadata as well as the pixel_array (image), results saved to csv.gz
# Create a df to get a list of all images
temp_df = image_list
temp_df['metadata'] = temp_df.origin.map(get_metadata)
temp_df['rows'] = temp_df.metadata.map(lambda x: x[0])
temp_df['columns'] = temp_df.metadata.map(lambda x: x[1])
temp_df['pixels_row_col'] = temp_df['rows'] * temp_df['columns']
temp_df['pixel_spacing'] = temp_df.metadata.map(lambda x: x[2])
temp_df['bits_allocated'] = temp_df.metadata.map(lambda x: x[3])
temp_df['bits_stored'] = temp_df.metadata.map(lambda x: x[4])
temp_df['high_bits'] = temp_df.metadata.map(lambda x: x[5])
temp_df['rows_from_array'] = temp_df.metadata.map(lambda x: x[6])
temp_df['columns_from_array'] = temp_df.metadata.map(lambda x: x[7])
temp_df['pixels_row_col_array'] = temp_df['rows_from_array'] * temp_df['columns_from_array']
# temp_df.head()
temp_df.to_csv('data_staging/metadata_2.csv.gz',index=False)
# ### Loading the saved dataset of corrupted images"
test = pd.read_csv('data_staging/metadata_2.csv.gz')
# image_list = pd.read_csv('data_staging/metadata_2.csv.gz')
# temp_df = image_list
# ### Searching for 99 gives us the corrupted image
# +
temp_df[temp_df.rows_from_array == 99]
# -
# we can check it here
ds = pydicom.dcmread('../data/stage_1_train_images/ID_6431af929.dcm') #ID_6431af929.dcm is the one that doesn't work
# raw = ds.pixel_array
# ### list of corrupted images
# +
# list(temp_df[(temp_df.pixels_row_col != 262144) | (temp_df.pixels_row_col_array != 262144)].ID)
| resources/dcm_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# hide
# %load_ext autoreload
# %autoreload 2
# %load_ext nb_black
# %load_ext lab_black
# +
# default_exp evaluation
# -
# # Evaluation
# > Compute evaluation metrics.
# ## Overview
#
# This section provides evaluation schemes for both Numerai Classic and Signals.
# The `Evaluator` takes a `NumerFrame` as input and returns a Pandas DataFrame containing metrics for each given prediction column.
# +
# export
import time
import json
import numpy as np
import pandas as pd
from tqdm.auto import tqdm
import matplotlib.pyplot as plt
from typing import Tuple, Union
from numerapi import SignalsAPI
from rich import print as rich_print
from numerblox.numerframe import NumerFrame, create_numerframe
from numerblox.postprocessing import FeatureNeutralizer
from numerblox.key import Key
# + pycharm={"name": "#%%\n"}
# export
# hide
FNCV3_FEATURES = ["feature_honoured_observational_balaamite", "feature_polaroid_vadose_quinze", "feature_untidy_withdrawn_bargeman", "feature_genuine_kyphotic_trehala", "feature_unenthralled_sportful_schoolhouse", "feature_divulsive_explanatory_ideologue", "feature_ichthyotic_roofed_yeshiva", "feature_waggly_outlandish_carbonisation", "feature_floriated_amish_sprite", "feature_iconoclastic_parietal_agonist", "feature_demolished_unfrightened_superpower", "feature_styloid_subdermal_cytotoxin", "feature_ironfisted_nonvintage_chlorpromazine", "feature_torose_unspiritualised_kylie", "feature_tearing_unkingly_adulthood", "feature_stylolitic_brown_spume", "feature_ferial_incumbent_engraving", "feature_litigant_unsizable_rhebok", "feature_floatiest_quintuplicate_carpentering", "feature_tuberculate_patelliform_paging", "feature_cuddlesome_undernamed_incidental", "feature_loony_zirconic_hoofer", "feature_indign_tardier_borough", "feature_fair_papal_vinaigrette", "feature_attack_unlit_milling", "feature_froggier_unlearned_underworkman", "feature_peninsular_pulsatile_vapor", "feature_midmost_perspiratory_hubert", "feature_laminable_unspecified_gynoecium", "feature_bally_bathymetrical_isadora", "feature_skim_unmeant_bandsman", "feature_ungenuine_sporophytic_evangelist", "feature_supercelestial_telic_dyfed", "feature_inconsiderate_unbooted_ricer", "feature_inured_conservable_forcer", "feature_glibber_deficient_jakarta", "feature_morbific_irredentist_interregnum", "feature_conjoint_transverse_superstructure", "feature_tingling_large_primordiality", "feature_phyllopod_unconstrainable_blubberer", "feature_deformable_unitary_schistosity", "feature_unprovisioned_aquatic_deuterogamy", "feature_equipped_undoubted_athanasian", "feature_inflammable_numb_anticline", "feature_kinky_benzal_holotype", "feature_ruptured_designing_interpolator", "feature_hierologic_expectable_maiolica", "feature_boiling_won_rama", "feature_lovelorn_aided_limiter", "feature_soviet_zibeline_profiler", "feature_altimetrical_muddled_symbolism", "feature_bratty_disrespectable_bookstand", "feature_unshaken_ahorse_wehrmacht", "feature_mightier_chivalric_kana", "feature_gambrel_unblessed_gigantomachy", "feature_ethiopic_anhedonic_stob", "feature_overstrung_dysmenorrheal_ingolstadt", "feature_rose_buttoned_dandy", "feature_recipient_perched_dendrochronologist", "feature_spikier_ordinate_taira", "feature_mercian_luddite_aganippe", "feature_faint_consociate_rhytidectomy", "feature_unpressed_mahratta_dah", "feature_maxillary_orphic_despicability", "feature_clasping_fast_menstruation", "feature_obeliscal_bewildered_reviewer", "feature_babist_moribund_myna", "feature_underdressed_tanagrine_prying", "feature_corniest_undue_scall", "feature_reduplicative_appalling_metastable", "feature_wrathful_prolix_colotomy", "feature_limonitic_issuable_melancholy", "feature_approximal_telautographic_sharkskin", "feature_fribble_gusseted_stickjaw", "feature_spec_subversive_plotter", "feature_unsinkable_dumbstruck_octuplet", "feature_integrative_reviviscent_governed", "feature_tamil_grungy_empathy", "feature_canopic_exigible_schoolgirl", "feature_plumular_constantinian_repositing", "feature_serpentiform_trinary_imponderability", "feature_gyroidal_embowed_pilcher", "feature_unlivable_armenian_wedge", "feature_flawed_demonological_toady", "feature_pruinose_raploch_roubaix", "feature_seediest_ramshackle_reclamation", "feature_hagiological_refer_vitamin", "feature_alcibiadean_lumpier_origan", "feature_encased_unamiable_hasidism", "feature_evocable_woollen_guarder", "feature_hunchbacked_unturning_meditation", "feature_circumnavigable_naughty_retranslation", "feature_testicular_slashed_ventosity", "feature_potential_subsessile_disconnection", "feature_unswaddled_inenarrable_goody", "feature_stellular_paler_centralisation", "feature_angevin_fitful_sultan", "feature_subinfeudatory_brainy_carmel", "feature_simpatico_cadential_pup", "feature_esculent_erotic_epoxy", "feature_milliary_hyperpyretic_medea", "feature_coraciiform_sciurine_reef", "feature_weightiest_protozoic_brawler", "feature_cooled_perkiest_electrodeposition", "feature_differing_peptizing_womaniser", "feature_gleaming_monosyllabic_scrod", "feature_unyielding_dismal_divertissement", "feature_rankine_meaty_port", "feature_southernmost_unhuman_arbiter", "feature_singhalese_cerographical_ego", "feature_malignant_campodeid_pluton", "feature_dure_jaspery_mugging", "feature_educational_caustic_mythologisation", "feature_diverted_astral_dunghill", "feature_degenerate_diaphragmatic_literalizer", "feature_laced_scraggly_grimalkin", "feature_wheezier_unjaundiced_game", "feature_unimpressed_uninflected_theophylline", "feature_shiite_overfed_mense", "feature_irritant_reciprocal_pelage", "feature_bricky_runed_bottleful", "feature_phyletic_separate_genuflexion", "feature_peckish_impetrative_kanpur", "feature_unshrinking_semiarid_floccule", "feature_heartier_salverform_nephew", "feature_geostrophic_adaptative_karla", "feature_navigational_enured_condensability", "feature_confusable_pursy_plosion", "feature_clenched_wayward_coelostat", "feature_developed_arbitrary_traditionalist", "feature_unnameable_abysmal_net", "feature_completive_pedantical_sinecurist", "feature_witchy_orange_muley", "feature_misfeatured_sometime_tunneler", "feature_agaze_lancinate_zohar", "feature_subservient_wedged_limping", "feature_urticant_ultracentrifugal_wane", "feature_pulverized_unified_dupery", "feature_stoichiometric_unanswerable_leveller", "feature_cyanophyte_emasculated_turpin", "feature_unruly_salian_impetuosity", "feature_ataractic_swept_rubeola", "feature_pansophical_agitato_theatricality", "feature_recreational_homiletic_nubian", "feature_burning_phrygian_axinomancy", "feature_protractive_moral_forswearing", "feature_certificated_putrescent_godship", "feature_dietetic_unscholarly_methamphetamine", "feature_vegetable_manlier_macaco", "feature_anthropoid_pithy_newscast", "feature_verifying_imagism_sublease", "feature_deckled_exaggerative_algol", "feature_songful_intercostal_frightener", "feature_additive_untrustworthy_hierologist", "feature_translative_quantitative_eschewer", "feature_coseismic_surpassable_invariance", "feature_blubbery_octahedral_bushfire", "feature_continued_conjugated_natalia", "feature_dissident_templed_shippon", "feature_wally_unrotted_eccrinology", "feature_unforgivable_airtight_reinsurance", "feature_unrelenting_intravascular_mesenchyme", "feature_linear_scummiest_insobriety", "feature_ovine_bramblier_leaven", "feature_uninforming_predictable_pepino", "feature_pluviometrical_biannual_saiga", "feature_affettuoso_taxidermic_greg", "feature_lateral_confervoid_belgravia", "feature_coalier_hircine_brokerage", "feature_undiverted_analyzed_accidie", "feature_favourable_swankiest_tympanist", "feature_refractory_topped_dependance", "feature_bustled_fieriest_doukhobor", "feature_isobilateral_olden_nephron", "feature_circassian_leathern_impugner", "feature_signed_ringent_sunna", "feature_cornute_potentiometric_tinhorn", "feature_veristic_parklike_halcyon", "feature_geochemical_unsavoury_collection", "feature_guerrilla_arrested_flavine", "feature_undependable_stedfast_donegal", "feature_bijou_penetrant_syringa", "feature_lamarckian_tarnal_egestion", "feature_horticultural_footworn_superscription", "feature_unwithered_personate_dilatation", "feature_wrought_muckier_temporality", "feature_rival_undepraved_countermarch", "feature_irrevocable_unlawful_oral", "feature_flawy_caller_superior", "feature_elohistic_totalitarian_underline", "feature_unrecognisable_waxier_paging", "feature_paraffinoid_flashiest_brotherhood", "feature_depauperate_armipotent_decentralisation", "feature_palpebral_univalve_pennoncel", "feature_received_veiniest_tamarix", "feature_scissile_dejected_kainite", "feature_narcotized_collectivist_evzone", "feature_jamesian_scutiform_ionium", "feature_gambogian_feudalist_diocletian", "feature_moneyed_mesophytic_lester", "feature_purblind_autarkic_pyrenoid", "feature_paleolithic_myalgic_lech", "feature_fortyish_neptunian_catechumenate", "feature_tricksiest_pending_voile", "feature_forcipate_laced_greenlet", "feature_overjoyed_undriven_sauna", "feature_small_cumulative_graywacke", "feature_incertain_catchable_zibet", "feature_unsustaining_chewier_adnoun", "feature_ruthenic_peremptory_truth", "feature_blind_concordant_tribalist", "feature_strigose_rugose_interjector", "feature_binding_lanky_rushing", "feature_carolean_tearable_smoothie", "feature_nappiest_unportioned_readjustment", "feature_sarmatia_foldable_eutectic", "feature_plum_anemometrical_guessing", "feature_gubernacular_liguloid_frankie", "feature_castigatory_hundredfold_hearthrug", "feature_pennsylvanian_sibylic_chanoyu", "feature_unreaving_intensive_docudrama", "feature_relinquished_incognizable_batholith", "feature_indusiate_canned_cosh", "feature_maglemosian_kittle_coachbuilding", "feature_unreeling_homeothermic_macedonia", "feature_asteriated_invigorated_penitence", "feature_anucleate_knotted_nonage", "feature_shrinelike_unreplaceable_nitrogenization", "feature_lacerable_backmost_vaseline", "feature_unreceipted_latest_lesser", "feature_unimaginable_sec_kaka", "feature_goidelic_gobelin_ledge", "feature_incondite_undisappointing_telephotograph", "feature_concoctive_symmetric_abulia", "feature_anglophobic_unformed_maneuverer", "feature_gravimetric_ski_enigma", "feature_balmiest_spinal_roundelay", "feature_required_bibliological_tonga", "feature_amoroso_wimpish_maturing", "feature_exertive_unmodernised_scaup", "feature_rude_booziest_ilium", "feature_uncompelled_curvy_amerindian", "feature_septuple_bonapartean_sanbenito", "feature_tottery_unmetalled_codder", "feature_tachygraphical_sedimentological_mesoderm", "feature_adsorbed_blizzardy_burlesque", "feature_wistful_tussive_cycloserine", "feature_superjacent_grubby_axillary", "feature_biological_caprine_cannoneer", "feature_unreversed_fain_jute", "feature_unexalted_rebel_kofta", "feature_doggish_mouthwatering_abelard", "feature_forfeit_contributing_joinder", "feature_necked_moresque_lowell", "feature_footling_unpuckered_lophophore", "feature_thorniest_laughable_hindustani", "feature_hotter_cattish_aridity", "feature_developing_behind_joan", "feature_ectodermal_mandaean_saffian", "feature_crimpier_gude_housedog", "feature_probationary_readying_roundelay", "feature_inserted_inconvertible_functioning", "feature_manifold_melodramatic_girl", "feature_drizzling_refrigerative_imperfection", "feature_sardonic_primary_shadwell", "feature_monocyclic_galliambic_par", "feature_smutty_prohibited_sullivan", "feature_productile_auriform_fil", "feature_accommodable_crinite_cleft", "feature_clipped_kurdish_grainer", "feature_dustproof_unafraid_stampede", "feature_neutered_postpositive_writ", "feature_twelve_haphazard_pantography", "feature_riskier_ended_typo", "feature_smaller_colored_immurement", "feature_snatchy_xylic_institution", "feature_conchal_angriest_oophyte", "feature_multiseriate_oak_benzidine", "feature_gobioid_transhuman_interconnection", "feature_reservable_peristomal_emden", "feature_inestimable_unmoral_extraversion", "feature_nubby_sissified_value", "feature_incorporating_abominable_daily", "feature_herbaged_brownish_consubstantialist", "feature_solemn_wordier_needlework", "feature_evangelistic_cruel_dissimilitude", "feature_impetratory_shuttered_chewer", "feature_referenced_biliteral_chiropody", "feature_eleatic_fellow_auctioneer", "feature_malpighian_vaporized_biogen", "feature_expiscatory_wriest_colportage", "feature_yelled_hysteretic_eath", "feature_bitterish_buttocked_turtleneck", "feature_percipient_atelectatic_cinnamon", "feature_gobony_premonitory_twinkler", "feature_twittery_tai_attainment", "feature_crooked_wally_lobation", "feature_crookback_workable_infringement", "feature_brawling_unpeppered_comedian", "feature_glyphographic_reparable_empyrean", "feature_noctilucent_subcortical_proportionality", "feature_guardian_frore_rolling", "feature_denuded_typed_wattmeter", "feature_unreachable_neritic_saracen", "feature_enzymatic_poorest_advocaat", "feature_wariest_vulnerable_unmorality", "feature_guttering_half_spondee", "feature_distressed_bloated_disquietude", "feature_leaky_overloaded_rhodium", "feature_unsapped_anionic_catherine", "feature_kissable_forfeit_egotism", "feature_unsizable_ancestral_collocutor", "feature_healthier_unconnected_clave", "feature_cirsoid_buddhism_vespa", "feature_rid_conveyable_cinchonization", "feature_donsie_folkish_renitency", "feature_agee_sold_microhabitat", "feature_newfangled_huddled_gest", "feature_clandestine_inkiest_silkworm", "feature_unutterable_softening_roper", "feature_balaamitical_electropositive_exhaustibility", "feature_unvalued_untangled_keener", "feature_undisturbing_quadrifid_reinhardt", "feature_bucked_costume_malagasy", "feature_joint_unreturning_basalt", "feature_coordinate_shyer_evildoing", "feature_carunculate_discursive_hectare", "feature_cynic_unreckonable_feoffment", "feature_cnidarian_micrologic_sousaphone", "feature_unperceivable_unrumpled_appendant", "feature_dissolvable_chrismal_obtund", "feature_choosier_uncongenial_coachwood", "feature_grimmest_prostate_doctrinaire", "feature_granulative_uncritical_agostini", "feature_convalescence_deuteranopic_lemuroid", "feature_disintegrable_snakier_zion", "feature_thoughtful_accommodable_lack", "feature_basophil_urdy_matzo", "feature_repellant_unwanted_clarinetist", "feature_antimonarchist_ordainable_quarterage", "feature_hardback_saturnalian_cyclometer", "feature_mythic_florentine_psammite", "feature_serpentiform_incomplete_bessarabia", "feature_unappeasable_employed_photoelectron", "feature_seaboard_adducent_polynesian", "feature_genoese_uncreditable_subregion", "feature_dexter_unstifled_snoring", "feature_protonematal_springtime_varioloid", "feature_orchitic_reported_coloration", "feature_stelliform_curling_trawler", "feature_athenian_pragmatism_isomorphism", "feature_abating_unadaptable_weakfish", "feature_instructional_desensitized_symmetallism", "feature_disarrayed_rarefactive_trisulphide", "feature_partible_amphibrachic_classicism", "feature_ecstatic_foundational_crinoidea", "feature_unimproved_courtliest_uncongeniality", "feature_cosy_microtonal_cedar", "feature_heedful_argyle_russianization", "feature_unhonoured_detested_xenocryst", "feature_sicker_spelaean_endplay", "feature_coordinated_astir_vituperation", "feature_stratocratic_aerodynamic_herero", "feature_uneasy_unaccommodating_immortality", "feature_professional_platonic_marten", "feature_detrital_respected_parlance", "feature_contraceptive_cartelist_beast", "feature_tapestried_madding_acclimatiser", "feature_optic_mycelial_whimper", "feature_liftable_direful_polyploid", "feature_objective_micro_langton", "feature_entopic_interpreted_subsidiary", "feature_saclike_hyphal_postulator", "feature_recent_shorty_preferment", "feature_strip_honoured_trail", "feature_unsheltered_doughtiest_episiotomy", "feature_acclimatisable_unfeigned_maghreb", "feature_galactopoietic_luckiest_protecting", "feature_scarcest_vaporized_max", "feature_spicier_unstripped_initial", "feature_hooly_chekhovian_phytogeographer", "feature_smouldering_underground_wingspan", "feature_phantasmal_extenuative_britain", "feature_sciurine_stibial_lintwhite", "feature_eucharistic_widowed_misfeasance", "feature_libratory_seizable_orlando", "feature_brackish_obstructed_almighty", "feature_translucid_neuroanatomical_sego", "feature_unheeded_stylar_planarian", "feature_preceptive_rushed_swedenborgian", "feature_sumerian_descendible_kalpa", "feature_jazziest_spellbinding_philabeg", "feature_dormie_sodden_steed", "feature_directoire_propositional_clydebank", "feature_triangled_rubber_skein", "feature_vendean_thwartwise_resistant", "feature_preoral_tonsorial_souk", "feature_virescent_telugu_neighbour", "feature_prefigurative_downstream_transvaluation", "feature_undepreciated_partitive_ipomoea", "feature_coactive_bandoleered_trogon", "feature_southerly_assonant_amicability", "feature_cortical_halt_catcher", "feature_queenliest_childing_ritual", "feature_antarthritic_syzygial_wonderland", "feature_revitalizing_rutilant_swastika", "feature_holy_chic_cali", "feature_hermitical_stark_serfhood", "feature_deformable_productile_piglet", "feature_lentissimo_ducky_quadroon", "feature_happening_tristful_yodeling", "feature_guardant_giocoso_natterjack", "feature_bootleg_clement_joe", "feature_thousandth_hierarchal_plight", "feature_unhoped_hex_ventriloquism", "feature_unappreciated_humiliated_misapprehension", "feature_cragged_sacred_malabo", "feature_idled_unwieldy_improvement", "feature_censorial_leachier_rickshaw", "feature_carbuncled_athanasian_ampul"]
MEDIUM_FEATURES = ["feature_abstersive_emotional_misinterpreter", "feature_accessorial_aroused_crochet", "feature_acerb_venusian_piety", "feature_affricative_bromic_raftsman", "feature_agile_unrespited_gaucho", "feature_agronomic_cryptal_advisor", "feature_alkaline_pistachio_sunstone", "feature_altern_unnoticed_impregnation", "feature_ambisexual_boiled_blunderer", "feature_amoebaean_wolfish_heeler", "feature_amygdaloidal_intersectional_canonry", "feature_antipathetical_terrorful_ife", "feature_antipodal_unable_thievery", "feature_antisubmarine_foregoing_cryosurgery", "feature_apomictical_motorized_vaporisation", "feature_apophthegmatical_catechetical_millet", "feature_apostate_impercipient_knighthood", "feature_appraisive_anagrammatical_tentacle", "feature_arillate_nickelic_hemorrhage", "feature_armoured_finable_skywriter", "feature_assenting_darn_arthropod", "feature_assertive_worsened_scarper", "feature_atlantic_uveal_incommunicability", "feature_attuned_southward_heckle", "feature_autarkic_constabulary_dukedom", "feature_autodidactic_gnarlier_pericardium", "feature_axillary_reluctant_shorty", "feature_aztecan_encomiastic_pitcherful", "feature_barest_kempt_crowd", "feature_basaltic_arid_scallion", "feature_base_ingrain_calligrapher", "feature_beady_unkind_barret", "feature_belgravian_salopian_sheugh", "feature_biannual_maleficent_thack", "feature_bifacial_hexastyle_hemialgia", "feature_bleeding_arabesque_pneuma", "feature_bloodied_twinkling_andante", "feature_brawny_confocal_frail", "feature_brickier_heterostyled_scrutiny", "feature_built_reincarnate_sherbet", "feature_bushwhacking_unaligned_imperturbability", "feature_busty_unfitted_keratotomy", "feature_buxom_curtained_sienna", "feature_caecilian_unexperienced_ova", "feature_caespitose_unverifiable_intent", "feature_cairned_fumiest_ordaining", "feature_calceolate_pudgy_armure", "feature_calculating_unenchanted_microscopium", "feature_calefactive_anapaestic_jerome", "feature_calycled_living_birmingham", "feature_camphorated_spry_freemartin", "feature_caressive_cognate_cubature", "feature_casemated_ibsenian_grantee", "feature_castrated_presented_quizzer", "feature_casuistic_barbarian_monochromy", "feature_centric_shaggier_cranko", "feature_cerebrovascular_weeny_advocate", "feature_chafed_undenominational_backstitch", "feature_chaldean_vixenly_propylite", "feature_chaotic_granitoid_theist", "feature_chartered_conceptual_spitting", "feature_cheering_protonemal_herd", "feature_chelonian_pyknic_delphi", "feature_chopfallen_fasciate_orchidologist", "feature_christadelphian_euclidean_boon", "feature_chuffier_analectic_conchiolin", "feature_churrigueresque_talc_archaicism", "feature_clawed_unwept_adaptability", "feature_clerkish_flowing_chapati", "feature_coalier_typhoid_muntin", "feature_collective_stigmatic_handfasting", "feature_commensurable_industrial_jungfrau", "feature_communicatory_unrecommended_velure", "feature_conceding_ingrate_tablespoonful", "feature_confiscatory_triennial_pelting", "feature_congealed_lee_steek", "feature_congenial_transmigrant_isobel", "feature_congenital_conched_perithecium", "feature_conjugal_postvocalic_rowe", "feature_consecrate_legislative_cavitation", "feature_contaminative_intrusive_tagrag", "feature_continuate_unprocurable_haversine", "feature_contused_festal_geochemistry", "feature_coordinated_undecipherable_gag", "feature_covalent_methodological_brash", "feature_covalent_unreformed_frogbit", "feature_crablike_panniered_gloating", "feature_criticisable_authentical_deprecation", "feature_croupiest_shaded_thermotropism", "feature_ctenoid_moaning_fontainebleau", "feature_culinary_pro_offering", "feature_curling_aurorean_iseult", "feature_curtained_gushier_tranquilizer", "feature_cyrenaic_unschooled_silurian", "feature_decent_solo_stickup", "feature_degenerate_officinal_feasibility", "feature_demisable_expiring_millepede", "feature_demure_groutiest_housedog", "feature_dendritic_prothallium_sweeper", "feature_dentilingual_removed_osmometer", "feature_descendent_decanal_hon", "feature_desiderative_commiserative_epizoa", "feature_designer_notchy_epiploon", "feature_dichasial_hammier_spawner", "feature_dipped_sent_giuseppe", "feature_discrepant_ventral_shicker", "feature_dismaying_chaldean_tallith", "feature_dispiriting_araeostyle_jersey", "feature_diverticular_punjabi_matronship", "feature_doggish_whacking_headscarf", "feature_dovetailed_winy_hanaper", "feature_draconic_contractible_romper", "feature_emmetropic_heraclitean_conducting", "feature_encompassing_skeptical_salience", "feature_endangered_unthreaded_firebrick", "feature_enlightening_mirthful_laurencin", "feature_epicurean_fetal_seising", "feature_epidermic_scruffiest_prosperity", "feature_epitaxial_loathsome_essen", "feature_eruptive_seasoned_pharmacognosy", "feature_escutcheoned_timocratic_kotwal", "feature_euterpean_frazzled_williamsburg", "feature_exacerbating_presentationism_apagoge", "feature_expressed_abhominable_pruning", "feature_extractable_serrulate_swing", "feature_fake_trident_agitator", "feature_faltering_tergal_tip", "feature_farcical_spinal_samantha", "feature_faustian_unventilated_lackluster", "feature_favoring_prescript_unorthodoxy", "feature_festering_controvertible_hostler", "feature_fierier_goofier_follicle", "feature_fissirostral_multifoliate_chillon", "feature_flakiest_fleecy_novelese", "feature_flavourful_seismic_erica", "feature_fleshly_bedimmed_enfacement", "feature_foamy_undrilled_glaciology", "feature_fragrant_fifteen_brian", "feature_frequentative_participial_waft", "feature_fumarolic_known_sharkskin", "feature_fustiest_voiced_janet", "feature_galvanometric_sturdied_billingsgate", "feature_ganoid_osiered_mineralogy", "feature_generative_honorific_tughrik", "feature_glare_factional_assessment", "feature_glyptic_unrubbed_holloway", "feature_gone_honduran_worshipper", "feature_gossamer_placable_wycliffite", "feature_grazed_blameful_desiderative", "feature_greedier_favorable_enthymeme", "feature_groggy_undescried_geosphere", "feature_gullable_sanguine_incongruity", "feature_gutta_exploitive_simpson", "feature_haematoid_runaway_nightjar", "feature_hawkish_domiciliary_duramen", "feature_headhunting_unsatisfied_phenomena", "feature_hellenistic_scraggly_comfort", "feature_helpable_chanciest_fractionisation", "feature_hemispherical_unabsolved_aeolipile", "feature_hendecagonal_deathly_stiver", "feature_hexametric_ventricose_limnology", "feature_hibernating_soritic_croupe", "feature_highland_eocene_berean", "feature_hillier_unpitied_theobromine", "feature_himyarite_tetragonal_deceit", "feature_horizontal_snug_description", "feature_hotfoot_behaviorist_terylene", "feature_huskiest_compartmental_jacquerie", "feature_hydrologic_cymric_nyctophobia", "feature_hypermetropic_unsighted_forsyth", "feature_hypersonic_volcanological_footwear", "feature_hypogastric_effectual_sunlight", "feature_hypothetic_distressing_endemic", "feature_hysteric_mechanized_recklinghausen", "feature_iconomatic_boozier_age", "feature_illiterate_stomachal_terpene", "feature_impractical_endorsed_tide", "feature_incitant_trochoidal_oculist", "feature_incommensurable_diffused_curability", "feature_indefatigable_enterprising_calf", "feature_indentured_communicant_tulipomania", "feature_indirect_concrete_canaille", "feature_induplicate_hoarse_disbursement", "feature_inexpugnable_gleg_candelilla", "feature_inflexed_lamaism_crit", "feature_inhabited_pettier_veinlet", "feature_inhibited_snowiest_drawing", "feature_inseminated_filarial_mesoderm", "feature_insociable_exultant_tatum", "feature_instrumentalist_extrovert_cassini", "feature_integrated_extroversive_ambivalence", "feature_intended_involute_highbinder", "feature_intercalative_helvetian_infirmarian", "feature_interdental_mongolian_anarchism", "feature_intermontane_vertical_moo", "feature_interrogatory_isohyetal_atacamite", "feature_intersubjective_juristic_sagebrush", "feature_intertwined_leeriest_suffragette", "feature_introvert_symphysial_assegai", "feature_intrusive_effluent_hokkaido", "feature_invalid_chromatographic_cornishman", "feature_invalid_extortionary_titillation", "feature_iridic_unpropertied_spline", "feature_irresponsive_compositive_ramson", "feature_irritant_euphuistic_weka", "feature_isotopic_hymenial_starwort", "feature_jerkwater_eustatic_electrocardiograph", "feature_jiggish_tritheist_probity", "feature_juvenalian_paunchy_uniformitarianism", "feature_kerygmatic_splashed_ziegfeld", "feature_koranic_rude_corf", "feature_leaky_maroon_pyrometry", "feature_learned_claustral_quiddity", "feature_leggiest_slaggiest_inez", "feature_leisurable_dehortatory_pretoria", "feature_leukemic_paler_millikan", "feature_levigate_kindly_dyspareunia", "feature_liege_unexercised_ennoblement", "feature_limitable_astable_physiology", "feature_lipogrammatic_blowsier_seismometry", "feature_log_unregenerate_babel", "feature_lordly_lamellicorn_buxtehude", "feature_loricate_cryptocrystalline_ethnology", "feature_lost_quirky_botel", "feature_loyal_fishy_pith", "feature_malacological_differential_defeated", "feature_malagasy_abounding_circumciser", "feature_massed_nonracial_ecclesiologist", "feature_mattery_past_moro", "feature_maximal_unobserving_desalinisation", "feature_mazy_superrefined_punishment", "feature_merovingian_tenebrism_hartshorn", "feature_methylated_necrophilic_serendipity", "feature_midget_noncognizable_plenary", "feature_migrant_reliable_chirurgery", "feature_mined_game_curse", "feature_misanthropic_knurliest_freebooty", "feature_more_hindoo_diageotropism", "feature_mucky_loanable_gastrostomy", "feature_multilinear_sharpened_mouse", "feature_myographic_gawkier_timbale", "feature_naval_edified_decarbonize", "feature_nebule_barmier_bibliomania", "feature_nubblier_plosive_deepening", "feature_nucleophilic_uremic_endogen", "feature_obeisant_vicarial_passibility", "feature_offshore_defamatory_catalog", "feature_outdated_tapered_speciation", "feature_outsized_admonishing_errantry", "feature_oversea_permed_insulter", "feature_ovular_powered_neckar", "feature_padded_peripteral_pericranium", "feature_palatalized_unsucceeded_induration", "feature_palmy_superfluid_argyrodite", "feature_pansophic_merino_pintado", "feature_paraffinoid_irreplevisable_ombu", "feature_paramagnetic_complex_gish", "feature_passerine_ultraist_neon", "feature_patristical_analysable_langouste", "feature_peaty_vulgar_branchia", "feature_peculiar_sheenier_quintal", "feature_peltate_okay_info", "feature_perceivable_gasiform_psammite", "feature_perigean_bewitching_thruster", "feature_periscopic_thirteenth_cartage", "feature_permanent_cottony_ballpen", "feature_pert_performative_hormuz", "feature_petitionary_evanescent_diallage", "feature_phellogenetic_vibrational_jocelyn", "feature_piffling_inflamed_jupiter", "feature_planar_unessential_bride", "feature_planned_superimposed_bend", "feature_plexiform_won_elk", "feature_polaroid_squalliest_applause", "feature_precooled_inoperable_credence", "feature_puberulent_nondescript_laparoscope", "feature_publishable_apiarian_rollick", "feature_quadratic_untouched_liberty", "feature_questionable_diplex_caesarist", "feature_quinsied_increased_braincase", "feature_ratlike_matrilinear_collapsability", "feature_recidivism_petitory_methyltestosterone", "feature_reclaimed_fallibilist_turpentine", "feature_reclinate_cruciform_lilo", "feature_reconciling_dauby_database", "feature_reduplicate_conoid_albite", "feature_refreshed_untombed_skinhead", "feature_reminiscent_unpained_ukulele", "feature_renegade_undomestic_milord", "feature_reported_slimy_rhapsody", "feature_reserved_cleanable_soldan", "feature_restricted_aggregately_workmanship", "feature_resuscitative_communicable_brede", "feature_retinoscopy_flinty_wool", "feature_revealable_aeonian_elvira", "feature_revitalizing_dashing_photomultiplier", "feature_rheumy_epistemic_prancer", "feature_rimmed_conditional_archipelago", "feature_roasting_slaked_reposition", "feature_roiling_trimeric_kurosawa", "feature_rowable_unshod_noise", "feature_rubblier_chlorotic_stogy", "feature_ruffianly_uncommercial_anatole", "feature_rural_inquisitional_trotline", "feature_rusted_unassisting_menaquinone", "feature_ruthenian_uncluttered_vocalizing", "feature_salian_suggested_ephemeron", "feature_sallowish_cognisant_romaunt", "feature_scenic_cormophytic_bilirubin", "feature_scenographical_dissentient_trek", "feature_scorbutic_intellectualism_mongoloid", "feature_scrobiculate_unexcitable_alder", "feature_seamier_jansenism_inflator", "feature_seclusive_emendatory_plangency", "feature_seemlier_reorient_monandry", "feature_severe_tricky_pinochle", "feature_sixteen_inbreed_are", "feature_sludgy_implemental_sicily", "feature_smoggy_niftiest_lunch", "feature_smugger_hydroponic_farnesol", "feature_softish_unseparated_caudex", "feature_sorted_ignitable_sagitta", "feature_spagyric_echt_alum", "feature_spookiest_expedite_overnighter", "feature_springlike_crackjaw_bheesty", "feature_squishiest_unsectarian_support", "feature_stelar_balmiest_pellitory", "feature_stereotypic_ebracteate_louise", "feature_strychnic_structuralist_chital", "feature_stylistic_honduran_comprador", "feature_subapostolic_dungy_fermion", "feature_subdued_spiffier_kano", "feature_subglobular_unsalable_patzer", "feature_substandard_permissible_paresthesia", "feature_sudsy_polymeric_posteriority", "feature_supergene_legible_antarthritic", "feature_synoptic_botryose_earthwork", "feature_syrian_coital_counterproof", "feature_tarry_meet_chapel", "feature_telephonic_shakable_bollock", "feature_terrific_epigamic_affectivity", "feature_tittering_virgilian_decliner", "feature_together_suppositive_aster", "feature_tonal_graptolitic_corsac", "feature_tortured_arsenical_arable", "feature_torturesome_estimable_preferrer", "feature_tossing_denominative_threshing", "feature_trabeate_eutherian_valedictory", "feature_tranquilizing_abashed_glyceria", "feature_transmontane_clerkly_value", "feature_travelled_semipermeable_perruquier", "feature_tribal_germinable_yarraman", "feature_trim_axial_suffocation", "feature_unaimed_yonder_filmland", "feature_unamazed_tumular_photomicrograph", "feature_unapplicable_jerkiest_klemperer", "feature_unbeaten_orological_dentin", "feature_unbreakable_nosological_comedian", "feature_unburied_exponent_pace", "feature_uncertified_myrmecological_nagger", "feature_uncharged_unovercome_smolder", "feature_unco_terefah_thirster", "feature_uncomplimentary_malignant_scoff", "feature_uncompromising_fancy_kyle", "feature_uncurtailed_translucid_coccid", "feature_undescribed_methylic_friday", "feature_undetermined_idle_aftergrowth", "feature_undirected_perdu_ylem", "feature_undisguised_whatever_gaul", "feature_undivorced_unsatisfying_praetorium", "feature_undrossy_serpentiform_sack", "feature_unextinct_smectic_isa", "feature_uninclosed_handcrafted_springing", "feature_univalve_abdicant_distrail", "feature_unknown_reusable_cabbage", "feature_unlawful_superintendent_brunet", "feature_unlivable_morbific_traveling", "feature_unliving_bit_bengaline", "feature_unluckiest_mulley_benzyl", "feature_unmalleable_resistant_kingston", "feature_unmodernized_vasodilator_galenist", "feature_unmoved_alt_spoonerism", "feature_unnetted_bay_premillennialist", "feature_unnourishing_indiscreet_occiput", "feature_unperfect_implemental_cellarage", "feature_unrated_intact_balmoral", "feature_unrelieved_rawish_cement", "feature_unrequired_waxing_skeptic", "feature_unscheduled_malignant_shingling", "feature_unsparing_moralistic_commissary", "feature_unsparred_scarabaeid_anthologist", "feature_unspotted_practiced_gland", "feature_unstacked_trackable_blizzard", "feature_unsurveyed_boyish_aleph", "feature_unsurveyed_chopped_feldspathoid", "feature_untellable_penal_allegorization", "feature_untouchable_unsolvable_agouti", "feature_untrimmed_monaxial_accompanist", "feature_untumbled_histologic_inion", "feature_unvaried_social_bangkok", "feature_unweary_congolese_captain", "feature_uretic_seral_decoding", "feature_urochordal_swallowed_curn", "feature_vedic_mitral_swiz", "feature_venatic_intermetallic_darling", "feature_vestmental_hoofed_transpose", "feature_vizierial_courtlier_hampton", "feature_volitional_ascensive_selfhood", "feature_voltairean_consolidative_parallel", "feature_voltairean_dyslogistic_epagoge", "feature_vulcanological_sepulchral_spean", "feature_wale_planned_tolstoy", "feature_westering_immunosuppressive_crapaud", "feature_whistleable_unbedimmed_chokey", "feature_whitened_remanent_blast", "feature_whopping_eminent_attempter", "feature_wieldable_defiled_aperitive", "feature_wombed_reverberatory_colourer", "feature_zarathustrian_albigensian_itch", "feature_zymotic_varnished_mulga"]
# -
# ## 0. Base
# `BaseEvaluator` implements all the evaluation logic that is common for Numerai Classic and Signals. This includes:
# - Mean, Standard Deviation and Sharpe for era returns.
# - Max drawdown
# - Annual Percentage Yield (APY)
# - Mean, Standard deviation and Sharpe for [MMC (Meta Model Contribution)](https://docs.numer.ai/tournament/metamodel-contribution) returns.
# - Correlation with example predictions
# - Max [feature exposure](https://forum.numer.ai/t/model-diagnostics-feature-exposure/899)
# - [Feature Neutral Mean (FNC)](https://docs.numer.ai/tournament/feature-neutral-correlation), Standard deviation and Sharpe
# - [Exposure Dissimilarity](https://forum.numer.ai/t/true-contribution-details/5128/4)
# - Mean, Standard Deviation and Sharpe for TB200 (Buy top 200 stocks and sell bottom 200 stocks).
# - Mean, Standard Deviation and Sharpe for TB500 (Buy top 500 stocks and sell bottom 500 stocks).
#
# export
class BaseEvaluator:
"""
Evaluation functionality that is relevant for both
Numerai Classic and Numerai Signals.
:param era_col: Column name pointing to eras. \n
Most commonly "era" for Numerai Classic and "friday_date" for Numerai Signals. \n
:param fast_mode: Will skip compute intensive metrics if set to True,
namely max_exposure, feature neutral mean, TB200 and TB500.
"""
def __init__(self, era_col: str = "era", fast_mode=False):
self.era_col = era_col
self.fast_mode = fast_mode
def full_evaluation(
self,
dataf: NumerFrame,
example_col: str,
pred_cols: list = None,
target_col: str = "target",
) -> pd.DataFrame:
"""
Perform evaluation for each prediction column in the NumerFrame
against give target and example prediction column.
"""
val_stats = pd.DataFrame()
cat_cols = dataf.get_feature_data.select_dtypes(include=['category']).columns.to_list()
if cat_cols:
rich_print(f":warning: WARNING: Categorical features detected that cannot be used for neutralization. Removing columns: '{cat_cols}' for evaluation. :warning:")
dataf.loc[:, dataf.feature_cols] = dataf.get_feature_data.select_dtypes(exclude=['category'])
dataf = dataf.fillna(0.5)
pred_cols = dataf.prediction_cols if not pred_cols else pred_cols
for col in tqdm(pred_cols, desc="Evaluation: "):
col_stats = self.evaluation_one_col(
dataf=dataf,
pred_col=col,
target_col=target_col,
example_col=example_col,
)
val_stats = pd.concat([val_stats, col_stats], axis=0)
return val_stats
def evaluation_one_col(
self,
dataf: NumerFrame,
pred_col: str,
target_col: str,
example_col: str,
):
"""
Perform evaluation for one prediction column
against given target and example prediction column.
"""
col_stats = pd.DataFrame()
# Compute stats
val_corrs = self.per_era_corrs(
dataf=dataf, pred_col=pred_col, target_col=target_col
)
mean, std, sharpe = self.mean_std_sharpe(era_corrs=val_corrs)
max_drawdown = self.max_drawdown(era_corrs=val_corrs)
apy = self.apy(era_corrs=val_corrs)
example_corr = self.example_correlation(
dataf=dataf, pred_col=pred_col, example_col=example_col
)
mmc_mean, mmc_std, mmc_sharpe = self.mmc(
dataf=dataf,
pred_col=pred_col,
target_col=target_col,
example_col=example_col,
)
col_stats.loc[pred_col, "target"] = target_col
col_stats.loc[pred_col, "mean"] = mean
col_stats.loc[pred_col, "std"] = std
col_stats.loc[pred_col, "sharpe"] = sharpe
col_stats.loc[pred_col, "max_drawdown"] = max_drawdown
col_stats.loc[pred_col, "apy"] = apy
col_stats.loc[pred_col, "mmc_mean"] = mmc_mean
col_stats.loc[pred_col, "mmc_std"] = mmc_std
col_stats.loc[pred_col, "mmc_sharpe"] = mmc_sharpe
col_stats.loc[pred_col, "corr_with_example_preds"] = example_corr
# Compute intensive stats
if not self.fast_mode:
max_feature_exposure = self.max_feature_exposure(
dataf=dataf, pred_col=pred_col
)
fn_mean, fn_std, fn_sharpe = self.feature_neutral_mean_std_sharpe(
dataf=dataf, pred_col=pred_col, target_col=target_col
)
tb200_mean, tb200_std, tb200_sharpe = self.tbx_mean_std_sharpe(
dataf=dataf, pred_col=pred_col, target_col=target_col, tb=200
)
tb500_mean, tb500_std, tb500_sharpe = self.tbx_mean_std_sharpe(
dataf=dataf, pred_col=pred_col, target_col=target_col, tb=500
)
ex_diss = self.exposure_dissimilarity(
dataf=dataf, pred_col=pred_col, example_col=example_col
)
col_stats.loc[pred_col, "max_feature_exposure"] = max_feature_exposure
col_stats.loc[pred_col, "feature_neutral_mean"] = fn_mean
col_stats.loc[pred_col, "feature_neutral_std"] = fn_std
col_stats.loc[pred_col, "feature_neutral_sharpe"] = fn_sharpe
col_stats.loc[pred_col, "tb200_mean"] = tb200_mean
col_stats.loc[pred_col, "tb200_std"] = tb200_std
col_stats.loc[pred_col, "tb200_sharpe"] = tb200_sharpe
col_stats.loc[pred_col, "tb500_mean"] = tb500_mean
col_stats.loc[pred_col, "tb500_std"] = tb500_std
col_stats.loc[pred_col, "tb500_sharpe"] = tb500_sharpe
col_stats.loc[pred_col, "exposure_dissimilarity"] = ex_diss
return col_stats
def per_era_corrs(
self, dataf: pd.DataFrame, pred_col: str, target_col: str
) -> pd.Series:
"""Correlation between prediction and target for each era."""
return dataf.groupby(dataf[self.era_col]).apply(
lambda d: self._normalize_uniform(d[pred_col].fillna(0.5)).corr(
d[target_col]
)
)
def mean_std_sharpe(
self, era_corrs: pd.Series
) -> Tuple[np.float64, np.float64, np.float64]:
"""
Average, standard deviation and Sharpe ratio for
correlations per era.
"""
mean = pd.Series(era_corrs.mean()).item()
std = pd.Series(era_corrs.std(ddof=0)).item()
sharpe = mean / std
return mean, std, sharpe
@staticmethod
def max_drawdown(era_corrs: pd.Series) -> np.float64:
"""Maximum drawdown per era."""
# Arbitrarily large window
rolling_max = (
(era_corrs + 1).cumprod().rolling(window=9000, min_periods=1).max()
)
daily_value = (era_corrs + 1).cumprod()
max_drawdown = -((rolling_max - daily_value) / rolling_max).max()
return max_drawdown
@staticmethod
def apy(era_corrs: pd.Series, stake_compounding_lag: int = 4) -> np.float64:
"""
Annual percentage yield.
:param era_corrs: Correlation scores by era
:param stake_compounding_lag: Compounding lag for Numerai rounds (4 for Numerai Classic)
"""
payout_scores = era_corrs.clip(-0.25, 0.25)
payout_daily_value = (payout_scores + 1).cumprod()
apy = (
((payout_daily_value.dropna().iloc[-1]) ** (1 / len(payout_scores)))
** (
52 - stake_compounding_lag
) # 52 weeks of compounding minus n for stake compounding lag
- 1
) * 100
return apy
def example_correlation(
self, dataf: Union[pd.DataFrame, NumerFrame], pred_col: str, example_col: str
):
"""Correlations with example predictions."""
return self.per_era_corrs(
dataf=dataf,
pred_col=pred_col,
target_col=example_col,
).mean()
def max_feature_exposure(
self, dataf: Union[pd.DataFrame, NumerFrame], pred_col: str
) -> np.float64:
"""Maximum exposure over all features."""
max_per_era = dataf.groupby(self.era_col).apply(
lambda d: d[dataf.feature_cols].corrwith(d[pred_col]).abs().max()
)
max_feature_exposure = max_per_era.mean(skipna=True)
return max_feature_exposure
def feature_neutral_mean_std_sharpe(
self, dataf: Union[pd.DataFrame, NumerFrame], pred_col: str, target_col: str, feature_names: list = None
) -> Tuple[np.float64, np.float64, np.float64]:
"""
Feature neutralized mean performance.
More info: https://docs.numer.ai/tournament/feature-neutral-correlation
"""
fn = FeatureNeutralizer(pred_name=pred_col,
feature_names=feature_names,
proportion=1.0)
neutralized_dataf = fn(dataf=dataf)
neutral_corrs = self.per_era_corrs(
dataf=neutralized_dataf,
pred_col=f"{pred_col}_neutralized_1.0",
target_col=target_col,
)
mean, std, sharpe = self.mean_std_sharpe(era_corrs=neutral_corrs)
return mean, std, sharpe
def tbx_mean_std_sharpe(
self, dataf: pd.DataFrame, pred_col: str, target_col: str, tb: int = 200
) -> Tuple[np.float64, np.float64, np.float64]:
"""
Calculate Mean, Standard deviation and Sharpe ratio
when we focus on the x top and x bottom predictions.
:param tb: How many of top and bottom predictions to focus on.
TB200 and TB500 are the most common situations.
"""
tb_val_corrs = self._score_by_date(
dataf=dataf, columns=[pred_col], target=target_col, tb=tb
)
return self.mean_std_sharpe(era_corrs=tb_val_corrs)
def mmc(
self, dataf: pd.DataFrame, pred_col: str, target_col: str, example_col: str
) -> Tuple[np.float64, np.float64, np.float64]:
"""
MMC Mean, standard deviation and Sharpe ratio.
More info: https://docs.numer.ai/tournament/metamodel-contribution
"""
mmc_scores = []
corr_scores = []
for _, x in dataf.groupby(self.era_col):
series = self._neutralize_series(
self._normalize_uniform(x[pred_col]), (x[example_col])
)
mmc_scores.append(np.cov(series, x[target_col])[0, 1] / (0.29 ** 2))
corr_scores.append(self._normalize_uniform(x[pred_col]).corr(x[target_col]))
val_mmc_mean = np.mean(mmc_scores)
val_mmc_std = np.std(mmc_scores)
corr_plus_mmcs = [c + m for c, m in zip(corr_scores, mmc_scores)]
corr_plus_mmc_sharpe = np.mean(corr_plus_mmcs) / np.std(corr_plus_mmcs)
return val_mmc_mean, val_mmc_std, corr_plus_mmc_sharpe
def exposure_dissimilarity(self, dataf: NumerFrame, pred_col: str, example_col: str) -> np.float32:
"""
Model pattern of feature exposure to the example column.
See TC details forum post: https://forum.numer.ai/t/true-contribution-details/5128/4
"""
U = dataf.get_feature_data.corrwith(dataf[pred_col]).values
E = dataf.get_feature_data.corrwith(dataf[example_col]).values
exp_dis = 1 - np.dot(U, E) / np.dot(E, E)
return exp_dis
@staticmethod
def _neutralize_series(series, by, proportion=1.0):
scores = series.values.reshape(-1, 1)
exposures = by.values.reshape(-1, 1)
# this line makes series neutral to a constant column so that it's centered and for sure gets corr 0 with exposures
exposures = np.hstack(
(exposures, np.array([np.mean(series)] * len(exposures)).reshape(-1, 1))
)
correction = proportion * (
exposures.dot(np.linalg.lstsq(exposures, scores, rcond=None)[0])
)
corrected_scores = scores - correction
neutralized = pd.Series(corrected_scores.ravel(), index=series.index)
return neutralized
def _score_by_date(
self, dataf: pd.DataFrame, columns: list, target: str, tb: int = None
):
"""
Get era correlation based on given TB (x top and bottom predictions).
:param tb: How many of top and bottom predictions to focus on.
TB200 is the most common situation.
"""
unique_eras = dataf[self.era_col].unique()
computed = []
for u in unique_eras:
df_era = dataf[dataf[self.era_col] == u]
era_pred = np.float64(df_era[columns].values.T)
era_target = np.float64(df_era[target].values.T)
if tb is None:
ccs = np.corrcoef(era_target, era_pred)[0, 1:]
else:
tbidx = np.argsort(era_pred, axis=1)
tbidx = np.concatenate([tbidx[:, :tb], tbidx[:, -tb:]], axis=1)
ccs = [
np.corrcoef(era_target[idx], pred[idx])[0, 1]
for idx, pred in zip(tbidx, era_pred)
]
ccs = np.array(ccs)
computed.append(ccs)
return pd.DataFrame(
np.array(computed), columns=columns, index=dataf[self.era_col].unique()
)
@staticmethod
def _normalize_uniform(df: pd.DataFrame) -> pd.Series:
"""Normalize predictions uniformly using ranks."""
x = (df.rank(method="first") - 0.5) / len(df)
return pd.Series(x, index=df.index)
def plot_correlations(
self,
dataf: NumerFrame,
pred_cols: list = None,
target_col: str = "target",
roll_mean: int = 20,
):
"""
Plot per era correlations over time.
:param roll_mean: How many eras should be averaged to compute a rolling score.
"""
validation_by_eras = pd.DataFrame()
pred_cols = dataf.prediction_cols if not pred_cols else pred_cols
for pred_col in pred_cols:
per_era_corrs = self.per_era_corrs(
dataf, pred_col=pred_col, target_col=target_col
)
validation_by_eras.loc[:, pred_col] = per_era_corrs
validation_by_eras.rolling(roll_mean).mean().plot(
kind="line",
marker="o",
ms=4,
title=f"Rolling Per Era Correlation Mean (rolling window size: {roll_mean})",
figsize=(15, 5),
)
plt.legend(
loc="upper center",
bbox_to_anchor=(0.5, -0.05),
fancybox=True,
shadow=True,
ncol=1,
)
plt.axhline(y=0.0, color="r", linestyle="--")
plt.show()
validation_by_eras.cumsum().plot(
title="Cumulative Sum of Era Correlations", figsize=(15, 5)
)
plt.legend(
loc="upper center",
bbox_to_anchor=(0.5, -0.05),
fancybox=True,
shadow=True,
ncol=1,
)
plt.axhline(y=0.0, color="r", linestyle="--")
plt.show()
return
# ## 1. Numerai Classic
# `NumeraiClassicEvaluator` extends the base evaluation scheme with metrics specific to Numerai Classic.
#
# Additional metrics specific to Numerai are:
#
# - [Feature Neutral Mean v3 (FNCv3)](https://forum.numer.ai/t/true-contribution-details/5128/4), Standard deviation v3 and Sharpe v3.
# export
class NumeraiClassicEvaluator(BaseEvaluator):
"""Evaluator for all metrics that are relevant in Numerai Classic."""
def __init__(self, era_col: str = "era", fast_mode=False):
super().__init__(era_col=era_col, fast_mode=fast_mode)
self.fncv3_features = FNCV3_FEATURES
self.medium_features = MEDIUM_FEATURES
def full_evaluation(
self,
dataf: NumerFrame,
example_col: str,
pred_cols: list = None,
target_col: str = "target",
) -> pd.DataFrame:
val_stats = pd.DataFrame()
dataf = dataf.fillna(0.5)
pred_cols = dataf.prediction_cols if not pred_cols else pred_cols
# Check if sufficient columns are present in dataf to compute FNCv3
if set(self.fncv3_features).issubset(set(dataf.columns)):
valid_features = self.fncv3_features
elif set(self.medium_features).issubset(set(dataf.columns)):
print("WARNING: 'v4/fncv3_features' are not present in the DataFrame. Falling back on 'v3/medium' features.")
valid_features = self.medium_features
else:
print("WARNING: neither 'v4/fncv3_features' nor 'v3/medium' features are defined in DataFrame. Skipping calculation of v3 metrics.")
valid_features = []
for col in tqdm(pred_cols, desc="Evaluation: "):
# Metrics that can be calculated for both Numerai Classic and Signals
col_stats = self.evaluation_one_col(
dataf=dataf,
pred_col=col,
target_col=target_col,
example_col=example_col,
)
# Numerai Classic specific metrics
if not self.fast_mode and valid_features:
fnc_v3, fn_std_v3, fn_sharpe_v3 = self.feature_neutral_mean_std_sharpe(
dataf=dataf, pred_col=col, target_col=target_col, feature_names=valid_features
)
col_stats.loc[col, "feature_neutral_mean_v3"] = fnc_v3
col_stats.loc[col, "feature_neutral_std_v3"] = fn_std_v3
col_stats.loc[col, "feature_neutral_sharpe_v3"] = fn_sharpe_v3
val_stats = pd.concat([val_stats, col_stats], axis=0)
return val_stats
def __load_json(self, json_path: str) -> dict:
with open(json_path, 'r') as f:
data = json.load(f)
return data
# ## 2. Numerai Signals
# `NumeraiSignalsEvaluator` extends the base evaluation scheme with metrics specific to Numerai Signals.
#
# export
class NumeraiSignalsEvaluator(BaseEvaluator):
"""Evaluator for all metrics that are relevant in Numerai Signals."""
def __init__(self, era_col: str = "friday_date", fast_mode=False):
super().__init__(era_col=era_col, fast_mode=fast_mode)
def get_neutralized_corr(self, val_dataf: pd.DataFrame, model_name: str, key: Key, timeout_min: int = 2) -> pd.Series:
"""
Retrieved neutralized validation correlation by era. \n
Calculated on Numerai servers. \n
:param val_dataf: A DataFrame containing prediction, friday_date, era and data_type columns. \n
data_type column should contain 'validation' instances. \n
:param model_name: Any model name for which you have authentication credentials. \n
:param key: Key object to authenticate upload of diagnostics. \n
:param timeout_min: How many minutes to wait on diagnostics processing on Numerai servers before timing out. \n
2 minutes by default. \n
:return: Pandas Series with era as index and neutralized validation correlations (validationCorr).
"""
api = SignalsAPI(public_id=key.pub_id, secret_key=key.secret_key)
model_id = api.get_models()[model_name]
api.upload_diagnostics(df=val_dataf, model_id=model_id)
data = self.__await_diagnostics(api=api, model_id=model_id, timeout_min=timeout_min)
dataf = pd.DataFrame(data['perEraDiagnostics']).set_index("era")['validationCorr']
dataf.index = pd.to_datetime(dataf.index)
return dataf
@staticmethod
def __await_diagnostics(api: SignalsAPI, model_id: str, timeout_min: int, interval_sec: int = 15):
"""
Wait for diagnostics to be uploaded.
Try every 'interval_sec' seconds until 'timeout_min' minutes have passed.
"""
timeout = time.time() + 60 * timeout_min
data = {"status": "not_done"}
while time.time() < timeout:
data = api.diagnostics(model_id=model_id)[0]
if data['status'] == 'done':
break
else:
print(f"Diagnostics not processed yet. Sleeping for another {interval_sec} seconds.")
time.sleep(interval_sec)
if not data['status'] == 'done':
raise Exception(f"Diagnostics couldn't be retrieved within {timeout_min} minutes after uploading. Check if Numerai API is offline.")
return data
# ### Example usage
# We will test `NumeraiClassicEvaluator` on version 2 evaluation data with example predictions. The baseline reference (`example_col`) will be random predictions.
# +
from numerblox.download import NumeraiClassicDownloader
directory = "eval_test_1234321/"
downloader = NumeraiClassicDownloader(directory_path=directory)
downloader.download_single_dataset(filename="example_validation_predictions.parquet",
dest_path=directory + "example_validation_predictions.parquet")
downloader.download_single_dataset(filename="numerai_validation_data.parquet",
dest_path=directory + "numerai_validation_data.parquet")
# +
# other
np.random.seed(1234)
test_dataf = create_numerframe(directory + "numerai_validation_data.parquet")
example_preds = pd.read_parquet(directory + "example_validation_predictions.parquet")
test_dataf = test_dataf.merge(example_preds, on="id", how="left")
test_dataf.loc[:, "prediction_random"] = np.random.uniform(size=len(test_dataf))
test_dataf.head(2)
# -
# #### Full evaluation
# other
evaluator = NumeraiClassicEvaluator()
val_stats = evaluator.full_evaluation(
dataf=test_dataf,
target_col="target",
pred_cols=["prediction", "prediction_random"],
example_col="prediction_random",
)
val_stats
# The `Evaluator` returns a Pandas DataFrame containing metrics for each prediction column defined.
# Note that any column can be used as example prediction. For practical use cases we recommend using proper example predictions (provided by Numerai) instead of random predictions.
# #### Fast evaluation
#
# `fast_mode` skips max. feature exposure, feature neutral mean, FNCv3, Exposure Dissimilarity, TB200 and TB500 calculations, which can take a while to compute on full Numerai datasets.
# other
evaluator = NumeraiClassicEvaluator(fast_mode=True)
val_stats_fast = evaluator.full_evaluation(
dataf=test_dataf,
target_col="target",
pred_cols=["prediction", "prediction_random"],
example_col="prediction_random",
)
val_stats_fast
# #### Plot correlations
# The `plot_correlations` method will use matplotlib to plot per era correlation scores over time. The plots default to a rolling window of 20 eras in order to best align with repuation scores as measured on the Numerai leaderboards.
# other
evaluator.plot_correlations(
test_dataf, pred_cols=["prediction", "prediction_random"], roll_mean=20
)
# Clean up environment
downloader.remove_base_directory()
# --------------------------------------------------
# +
# hide
# Run this cell to sync all changes with library
from nbdev.export import notebook2script
notebook2script()
# -
| nbs/07_evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using GECKO for metabolic engineering applications
#
# <NAME>, 2020-09-10
# +
# requirements
import numpy as np
import pandas as pd
import os
import sys
import wget
from cobra import Reaction
from cobra.flux_analysis import pfba
from cobra.io import load_json_model
from cameo.strain_design.deterministic.flux_variability_based import FSEOF
# -
# ## 1. Loading models
#
# Load the metabolic and enzyme constrained models:
# +
# Metabolic model:
wget.download("https://github.com/BenjaSanchez/notebooks/blob/master/caffeine-fix-yarrowia/iYali-model.json?raw=true", "model.json", bar = False)
model = load_json_model("model.json")
os.remove("model.json")
# Enzyme-constrained model:
wget.download("https://github.com/BenjaSanchez/notebooks/blob/master/caffeine-fix-yarrowia/iYali-ecModel.json?raw=true", "ec_model.json", bar = False)
ec_model = load_json_model("ec_model.json")
os.remove("ec_model.json")
# -
# The model has two differences with a standard COBRA model. First, the reactions contain another metabolite: the enyzme itself.
model.reactions.NDP7
ec_model.reactions.NDP7No1
ec_model.metabolites.prot_Q6CBD9
# **NB:** All protein ids follow the form `prot_UNIPROT`.
#
# The second difference is the existence of _protein exchange reactions_. These protein exchanges follow the naming `prot_UNIPROT_exchange`.
ec_model.reactions.prot_Q6CBD9_exchange
# By putting an upper bound on these exchanges, we can integrate proteomics data into the model and treat it as an usual COBRA model without further changes.
#
# **NB:** Without any additional constraints, both models predict more or less the same using simple FBA simulations:
model.optimize()
model.summary()
ec_model.optimize()
ec_model.summary()
# ## 2. Metabolic engineering applications of models
#
# We will optimize for the production of:
model.metabolites.pmtcoa_c
# For this we add to both models a reaction that "burns" the palmitoyl from the palmitoyl-CoA:
# +
def add_pmt_rxn(model):
reaction = Reaction('test')
reaction.name = 'test rxn'
reaction.lower_bound = 0
reaction.upper_bound = 1000
reaction.add_metabolites({
model.metabolites.pmtcoa_c: -1.0,
model.metabolites.coa_c: +1})
model.add_reactions([reaction])
print(reaction)
add_pmt_rxn(model)
add_pmt_rxn(ec_model)
# -
# Now we can perform any typical analysis, e.g. FSEOF:
# +
biomass_rxn_id = 'xBIOMASS'
test_rxn_id = 'test'
def sorted_fseof(model, biomass_rxn_id, test_rxn_id):
# Revert the model to its original state:
model.reactions.get_by_id(biomass_rxn_id).lower_bound = 0
model.reactions.get_by_id(test_rxn_id).lower_bound = 0
model.objective = biomass_rxn_id
# Run analysis
fseof = FSEOF(model)
fseof_result = fseof.run(target=model.reactions.get_by_id(test_rxn_id))
fseof_df = fseof_result.data_frame
# For each row, create a linear model with the test exchange as prediction, and store the slope of said model:
fseof_df["slope"] = np.nan
fseof_df["r2"] = np.nan
for index, row in fseof_df.iterrows():
if sum(row) == 0:
fseof_df.loc[index,"slope"] = 0
else:
x = row.iloc[:-2]
y = fseof_df.loc[test_rxn_id].iloc[:-2]
A = np.vstack([x, np.ones(len(x))]).T
m, c = np.linalg.lstsq(A, y, rcond=None)[0]
resid = np.linalg.lstsq(A, y, rcond=None)[1]
r2 = 1 - resid / (y.size * y.var())
fseof_df.loc[index,"slope"] = m
try:
fseof_df.loc[index,"r2"] = r2
except:
fseof_df.loc[index,"r2"] = 0
# Sort the dataframe by slope and print only rows with R2 > 0.5:
fseof_df = fseof_df.sort_values(by=["slope"], ascending=False)
print(fseof_df.loc[fseof_df.index != "test"].loc[fseof_df["r2"] > 0.5].iloc[:20, :])
return fseof_df
# Run the FSEOF analysis for both models:
fseof_df = sorted_fseof(model, biomass_rxn_id, test_rxn_id)
ec_fseof_df = sorted_fseof(ec_model, biomass_rxn_id, test_rxn_id)
# -
# ## 3. Integrating proteomics in an ecModel
#
# We will now load some data.
#
# **NB:** Data should come in mmol/gDW units.
proteomics = pd.read_csv("fake-data.csv", index_col=0, squeeze=True)
proteomics.items
# We now add the data to the ecModel:
# +
# Add simulations to python's search path & import simulation functions:
sys.path.append('../../simulations/src')
from simulations.modeling.driven import limit_proteins, flexibilize_proteomics
# Limit model with abundances:
limit_proteins(ec_model, proteomics)
# -
# Visualize results:
ec_model.optimize()
# Note that the model grows very slow! Probably due to some values in the data (it is fake after all...)
#
# ## 4. Flexibilization
#
# Experimental proteomics measurements can be too restrictive, due to instrument error and/or non-accurate kinetic data in the model. Thus, a flexibilization of the proteomics data is usually required to work with enzyme constrained models. Let's do that by requiring at least a growth rate of 0.1 1/h:
# +
# Convert proteomics to list opf dictionaries (required for flexibilizing):
ec_model.reactions.EX_glc__D_e_REV.upper_bound = +10
proteomics_list = []
for key, value in proteomics.items():
protein = {"identifier":key, "measurement":value, "uncertainty":0}
proteomics_list.append(protein)
#Flexibilize proteomics:
D = {"measurement":0.1, "uncertainty":0.01}
new_growth_rate, proteomics_filtered, warnings = flexibilize_proteomics(ec_model, biomass_rxn_id, D, proteomics_list, [])
print(f"new growth rate: {new_growth_rate} h-1")
print(f"filtered proteins: {len(proteomics) - len(proteomics_filtered)} proteins")
# -
ec_model.optimize()
# Now that we have a functional model, we can repeat the previous FSEOF analysis
# Run the FSEOF analysis, filtering out any reaction that is not a protein exchange pseudo-rxn:
ec_fseof_df = sorted_fseof(ec_model, biomass_rxn_id, test_rxn_id)
# ## 5. Enzyme Usage
#
# Finally, we can look at enzyme usage values between biomass production and our test production. For this, we should first perform simulations with the ecModel now that it has proteomics data integrated:
# +
def simulate_ec_model(model, rxn_id):
# First optimization: maximize input
model.objective = rxn_id
solution = model.optimize()
# Second optimization: minimize glucose
model.reactions.get_by_id(rxn_id).lower_bound = solution.fluxes[rxn_id]
glc_rxn = model.reactions.get_by_id("EX_glc__D_e_REV")
model.objective = {glc_rxn: -1}
solution = pfba(model)
return solution
def print_fluxes(model, solution, bio_rxn_id):
print("growth: " + str(solution.fluxes[bio_rxn_id]))
for rxn in model.reactions:
if (len(rxn.metabolites) == 1 or rxn.id == "test") and solution.fluxes[rxn.id] != 0 and "_exchange" not in rxn.id:
print(rxn.id + ": " + str(solution.fluxes[rxn.id]))
# -
# Condition 1: 100% of carbon going towards biomass.
ec_sol_biomass = simulate_ec_model(ec_model, biomass_rxn_id)
print_fluxes(ec_model, ec_sol_biomass, biomass_rxn_id)
# Condition 2: 20% going towards the desired metabolite, and the rest to biomass.
ec_model.reactions.get_by_id(biomass_rxn_id).lower_bound = 0.8 * ec_sol_biomass.fluxes[biomass_rxn_id]
ec_sol_test = simulate_ec_model(ec_model, test_rxn_id)
print_fluxes(ec_model, ec_sol_test, biomass_rxn_id)
# Now let's build a dataframe with all enzyme usages under both conditions. We are looking for this in the `prot_XXXXXX_exchange` rxns, and they are all in units of `mmol/gDW`.
# +
usage_df = pd.DataFrame()
for reaction in ec_model.reactions:
if reaction.id.startswith("prot_") and reaction.id.endswith("_exchange"):
new_line = pd.DataFrame(index = [reaction.id[5:11]],
data = {"gene":reaction.gene_reaction_rule,
"biomass_usage":[ec_sol_biomass.fluxes[reaction.id]],
"test_usage":[ec_sol_test.fluxes[reaction.id]]})
usage_df = usage_df.append(new_line)
print(usage_df)
# -
# Let's make sure all values are positive:
usage_df = usage_df.sort_values(by=['biomass_usage'])
print(usage_df.head(n=5))
usage_df = usage_df.sort_values(by=['test_usage'])
print(usage_df.head(n=5))
# There are a lot of rows with zero usage under both conditions, so let's filter them out:
usage_df = usage_df.query("biomass_usage > 0 or test_usage > 0")
print(usage_df)
# Now let's compute usage changes. We will look at both absolute changes (the difference between both conditions) and relative changes (the fold change or ratio between them).
usage_df["abs_changes"] = usage_df["test_usage"] - usage_df["biomass_usage"]
usage_df["rel_changes"] = usage_df["test_usage"] / usage_df["biomass_usage"]
print(usage_df)
# We can now sort and take a look at the top 20 of enzymes that:
#
# * Increased their absolute usage the most:
usage_df = usage_df.sort_values(by=['abs_changes'])
print(usage_df.tail(n=20).iloc[::-1])
# * Decreased their absolute usage the most:
print(usage_df.head(n=20))
# * Increased their relative usage the most:
usage_df = usage_df.sort_values(by=['rel_changes'])
print(usage_df.tail(n=20).iloc[::-1])
# * Decreased their relative usage the most:
print(usage_df.head(n=20))
# ## 6. Adding heterologous reactions with enzyme properties
#
# For this we need to know the kcat of the reaction being added:
# +
from cobra import Metabolite
new_prot = Metabolite("prot_P1234")
ec_model.add_metabolites([new_prot])
kcat = 1e6
reaction = Reaction('test2')
reaction.name = 'test rxn'
reaction.lower_bound = 0
reaction.upper_bound = 1000
reaction.add_metabolites({
ec_model.metabolites.pmtcoa_c: -1.0,
ec_model.metabolites.prot_P1234: -1/(kcat),
ec_model.metabolites.coa_c: +1})
ec_model.add_reactions([reaction])
print(reaction)
reaction = Reaction('prot_P1234_exchange')
reaction.name = 'prot exchange'
reaction.lower_bound = 0
reaction.upper_bound = 1000
reaction.add_metabolites({ec_model.metabolites.prot_P1234: +1})
ec_model.add_reactions([reaction])
print(reaction)
# -
| ec-model-yarrowia/gecko-yarrowia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="x9xv-59YNpl5" executionInfo={"status": "ok", "timestamp": 1626891418391, "user_tz": -180, "elapsed": 1217, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}}
import numpy as np
import pandas as pd
from sklearn.datasets import load_iris
from sklearn.preprocessing import StandardScaler
# + colab={"base_uri": "https://localhost:8080/", "height": 197} id="KAPgR2g1O1lp" executionInfo={"status": "ok", "timestamp": 1626891418400, "user_tz": -180, "elapsed": 37, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="b297309b-7699-4d62-c161-44e33070de28"
iris = load_iris()
df = pd.DataFrame(data= np.c_[iris['data'], iris['target']],
columns= iris['feature_names'] + ['target'])
df.head()
# + id="-2qXh1tYP9sb" executionInfo={"status": "ok", "timestamp": 1626891418402, "user_tz": -180, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}}
X = df.loc[:, ~df.columns.isin(['target'])]
y = df[['target']]
# + colab={"base_uri": "https://localhost:8080/"} id="3d6kLNY5O1pV" executionInfo={"status": "ok", "timestamp": 1626891418403, "user_tz": -180, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="f7e25c06-bd0c-436d-833b-21d057573e61"
X_scaled = StandardScaler().fit_transform(X)
X_scaled[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="GAMxjLY4O1tJ" executionInfo={"status": "ok", "timestamp": 1626891418404, "user_tz": -180, "elapsed": 27, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="e0698cc3-86b7-4651-ed71-746e4dad2dd0"
features = X_scaled.T
cov_matrix = np.cov(features)
cov_matrix[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="ZtPDaxMLO1vz" executionInfo={"status": "ok", "timestamp": 1626891418406, "user_tz": -180, "elapsed": 26, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="d37ae975-1b67-496e-87f6-cfacf1d0693f"
values, vectors = np.linalg.eig(cov_matrix)
values[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="coL_9kPvO1yg" executionInfo={"status": "ok", "timestamp": 1626891418407, "user_tz": -180, "elapsed": 24, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="0970c93e-04f8-42a7-a48b-2b2056eadc95"
vectors[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="th8KtcF6O11k" executionInfo={"status": "ok", "timestamp": 1626891418409, "user_tz": -180, "elapsed": 24, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="011b4245-fbc1-4ba5-cfe6-cb924d60db29"
explained_variances = []
for i in range(len(values)):
explained_variances.append(values[i] / np.sum(values))
print(np.sum(explained_variances), '\n', explained_variances)
# + colab={"base_uri": "https://localhost:8080/", "height": 197} id="dRGf75fdRsH5" executionInfo={"status": "ok", "timestamp": 1626891418410, "user_tz": -180, "elapsed": 23, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="178beead-8469-422f-a42d-129443c24fd5"
projected_1 = X_scaled.dot(vectors.T[0])
projected_2 = X_scaled.dot(vectors.T[1])
res = pd.DataFrame(projected_1, columns=['PC1'])
res['PC2'] = projected_2
res['Y'] = y
res.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 678} id="pmfSec-PRsK3" executionInfo={"status": "ok", "timestamp": 1626891419167, "user_tz": -180, "elapsed": 778, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="be455245-96f4-49d3-8d3b-d727f99ef13d"
import matplotlib.pyplot as plt
import seaborn as sns
plt.figure(figsize=(20, 10))
sns.scatterplot(res['PC1'], [0] * len(res), hue=res['Y'], s=200)
# + colab={"base_uri": "https://localhost:8080/", "height": 678} id="lhga42FhRsNS" executionInfo={"status": "ok", "timestamp": 1626891419630, "user_tz": -180, "elapsed": 482, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}} outputId="a91396e5-1a7e-4dc2-b973-dbb16961f0f4"
plt.figure(figsize=(20, 10))
sns.scatterplot(res['PC1'], len(res), hue=res['Y'], s=50)
# + id="OOpiizeCRsP6" executionInfo={"status": "ok", "timestamp": 1626891419632, "user_tz": -180, "elapsed": 17, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08851466340381164023"}}
| Chapter 6/Python/Dimension reduction/Towards DS PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Core routines for modeling is found in the file `modeling.py`. This notebook is for sample testing and analytics only.
# +
# %pylab inline
from matplotlib import pyplot as plt
from workflow.data import *
from workflow.features import *
import pandas as pd
import numpy as np
import seaborn as sns
from sqlalchemy import create_engine
from sqlalchemy_utils import database_exists, create_database
import psycopg2
plt.tight_layout
plt.rcParams.update({'font.size': 22})
rc('xtick', labelsize=15)
rc('ytick', labelsize=15)
figure(figsize(10,7))
cmap= sns.color_palette('Set1')
sns.set_palette(cmap)
# +
# connect to SQL database
username = 'psam071'
host = 'localhost'
dbname = 'citibike'
db = create_engine('postgres://%s%s/%s' % (username,host,dbname))
con = None
con = psycopg2.connect(database = dbname, user = username, host = host)
# -
# # Querying Data and transforming it
# +
# query stations from 2015 that existed at the beginning of the data
# collecting period
query_stations2015 = """
SELECT DISTINCT a.id, name, lat, long, neighborhood, borough
FROM features a
LEFT JOIN stations b ON a.id = b.id
LEFT JOIN neighborhoods c on a.id = c.id
WHERE a.date = '2015-03-01'
--AND tot_docks > 0
AND borough = 'Manhattan'
ORDER BY a.id;
"""
stations2015 = pd.read_sql_query(query_stations2015, con)
stations2015 = stations2015.dropna()
# +
# most unbalanced stations
query_unbal_stations = """
SELECT a.id, abs(a.bikes_in - a.bikes_out) as flux,
abs(a.rbikes_in - a.rbikes_out) as rflux,
a.bikes_in, a.bikes_out,
c.name, neighborhood, borough, long, lat
FROM (SELECT id, min(date) as date, sum(bikes_out) as bikes_out,
sum(bikes_in) as bikes_in,
sum(rbikes_in) as rbikes_in, sum(rbikes_out) as rbikes_out
FROM features
GROUP BY id) a
JOIN neighborhoods b ON a.id = b.id
JOIN stations c on b.id = c.id
WHERE date = '2015-03-01'
ORDER BY rflux DESC
LIMIT 100;
"""
# make query and filter stations that existed at the beginning of the data collection phase
df_unbal_stations = pd.read_sql_query(query_unbal_stations, con)
# df_unbal_stations = df_unbal_stations[df_unbal_stations.id.isin(stations2015.id)]
# -
# save list of top 100 unbalanced stations to pickle file for webapp
df_unbal_stations.to_pickle('websitetools/stations.pickle')
# +
def dfcol_into_sqllist(df, col):
# converts a column in a pandas dataframe into a string for sql queries
listy = list(df[col].unique())
listy = listy[0:10]
return "(" + str(listy)[1:-1] + ")"
string_of_unbal_stations = str(list(df_unbal_stations.id.unique()))[1:-1]
list_of_unbal_stations = list(df_unbal_stations.id)
df_unbal_stations.id.unique().shape
# +
# look at the patterns for the rebalanced stations
# make sure tot_docks > 0 (especially when calculating bikes available)
ids_to_see = dfcol_into_sqllist(df_unbal_stations, 'id')
# ids_to_see = '(' + str(['72'])[1:-1] + ')'
query = """
SELECT a.id, a.date, a.hour, bikes_out, bikes_in, dayofweek, month, is_weekday,
is_holiday, rbikes_out, rbikes_in, tot_docks, avail_bikes, avail_docks,
precip, temp, long, lat, neighborhood, borough
FROM features a
LEFT JOIN weather b ON a.date = b.date AND a.hour = b.hour
LEFT JOIN stations c ON a.id = c.id
LEFT JOIN neighborhoods d ON a.id = d.id
WHERE a.id in {}
AND tot_docks > 0
AND borough = 'Manhattan'
--WHERE tot_docks > 0
ORDER BY a.id, a.date, a.hour;
""".format(ids_to_see)
df = pd.read_sql_query(query, con)
df.date = pd.to_datetime(df.date)
# -
# make new features (percentages)`
df = new_features(df)
# +
# split data into 2015 (train) and 2016 (test) data
#
data_cols = ['id', 'long', 'lat', 'hour', 'month', 'dayofweek',
'is_weekday',
'is_holiday', 'precip', 'temp', 'pct_avail_bikes',
'pct_avail_docks']#, 'pct_avail_docks']
# df = make_categorical(df, ['id', 'hour', 'month', 'is_weekday', 'is_holiday'])
hist_cols = ['mean_flux', 'yest_flux', 'last_week_flux']
df2015 = df[(df.date.dt.year == 2015)]
df2016 = df[(df.date.dt.year == 2016)]
# -
# # Prepare pipeline
# ### TPOT Regressor
# +
# tpot regressor
# from tpot import TPOTRegressor
# from sklearn.model_selection import train_test_split
# # data = df[data_cols + hist_cols].sort_index()
# # target = df.pct_flux
# # X_train, X_test, y_train, y_test = train_test_split(data, target,
# # train_size = 0.75, test_size = 0.25)
# X_train = df2015[data_cols]
# y_train = df2015.pct_flux
# X_test = df2016[data_cols]
# y_test = df2016.pct_flux
# reg = TPOTRegressor(generations=2, population_size = 5, verbosity=2)
# reg.fit(X_train, y_train)
# pred = tpot.predict(X_test)
# -
# ### RandomForestRegressor
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import GridSearchCV
params = [{'min_samples_leaf': [12, 14, 16],
'min_samples_split': [6, 8, 10],
'max_features': [0.85,0.95,1.]}]
X_train = df2015[data_cols]
y_train = df2015.pct_flux#.apply(flux_conditions, 0.15)
X_test = df2016[data_cols]
y_test = df2016.pct_flux#.apply(flux_conditions, 0.15)
# reg = GridSearchCV(RandomForestRegressor(), params, cv=2, scoring = 'neg_mean_squared_error')
reg = RandomForestRegressor(min_samples_leaf=16, min_samples_split=6, max_features = 0.95,
n_jobs=-1)
reg.fit(X_train, y_train)
pred = reg.predict(X_test)
# +
reg.feature_importances_.round(2)
importances = list(reg.feature_importances_.round(2))
features_dict = {}
for importance, col in zip(importances, data_cols):
features_dict[col] = importance
feature_imp = pd.Series(features_dict)
values_to_plot = feature_imp.sort_values()
values_to_plot.rename(index = {'id':'Station ID',
'hour': 'Hour',
'pct_avail_bikes': 'Available Bikes',
'dayofweek': 'Day of the Week',
'is_weekday': 'Is a Weekday',
'temp': 'Temperature',
'precip': 'Precipitation',
'month': 'Month',
'lat': 'Station Latitude',
'long': 'Station Longitude',
'pct_avail_docks': 'Available Docks',
'is_holiday': 'Is a Holiday'}, inplace = True)
values_to_plot.plot(kind = 'barh', figsize=(7,7))
xlabel('Feature Importance', fontsize = 15)
# -
# # Test Model
# +
def merge_pred_test(pred, y_test):
tests = pd.DataFrame()
tests['pct_flux_test'] = y_test.reset_index().pct_flux
tests['pct_flux_pred'] = pred
return tests
def plot_pred_test(test,X_test):
tests.iloc[50050:51000].plot(figsize = (13,5), alpha = 0.5)
X_test.reset_index().iloc[50050:51000].pct_avail_bikes.plot(alpha = 0.3)
tests = merge_pred_test(pred, y_test)
# tests.pct_flux_test = tests.pct_flux_test.apply(flux_conditions, 0.2)
# tests.pct_flux_pred = tests.pct_flux_pred.apply(flux_conditions, 0.2)
plot_pred_test(tests, X_test)
# +
def merge_test_pred(X, y, pred):
pred_series = pd.Series(pred)
X = X.reset_index()
X['pct_flux_test'] = y.reset_index().pct_flux
X['pct_flux_pred'] = pred_series
return X
df_compare = merge_test_pred(X_test, y_test, pred)
# +
cols = ['pct_flux_pred', 'pct_flux_test', 'hour']
weekday = 1
dock_id = 477
grps = df_compare.groupby(['id','is_weekday', 'hour']).mean().reset_index()
grps_std = df_compare.groupby(['id','is_weekday', 'hour']).std().reset_index()
cond = (grps.is_weekday == weekday) & (grps.id == dock_id)
# grps[cond][cols].set_index('hour').plot()
hr_profile = grps[cond][cols].set_index('hour')
hr_profile_errors = grps_std[cond][cols].set_index('hour')
x = hr_profile.pct_flux_pred
error = hr_profile_errors.pct_flux_pred
ax=hr_profile.pct_flux_test.plot(label = '2016 Data', linewidth = 3, color = 'red')
x.plot(linewidth = 3, label = '2016 Prediction', color = 'steelblue')
fill_between(list(x.index), list(x - error), list(x + error), alpha = 0.2, color = 'steelblue')
labels = ['12 AM', '5 AM', '10 AM', '3 PM', '8 PM']
ax.set_xticklabels(labels)
xlabel('Time of Day', size = 20)
legend(loc = 2, prop = {'size':15})
ylim([-0.45,0.45])
suptitle('2016 Bike Flow for Station: {} (Weekday)'.format(station_name))
# +
from sklearn.metrics import mean_squared_error, r2_score
def scoring_metrics(predicted, labeled):
mse = mean_squared_error(predicted, labeled)
r2 = r2_score(predicted, labeled)
print 'MSE: {}'.format(mse)
print 'R2: {}'.format(r2)
scoring_metrics(hr_profile.pct_flux_pred, hr_profile.pct_flux_test)
# return mse, r2
# mean_squared_error(y_test, pred)
# r2_score(y_test, pred)
# explained_variance_score(y_test, pred)
# RegressorMixin.score(X_test, y_test)
# -
# # Miscellaneous Plotting
# +
# prediction/observation plot
line = linspace(-1,1, num = 50)
fig = plt.figure(figsize = (10,10))
pl = fig.add_subplot(111)
pl.scatter(pred, y_test, alpha = 0.1)
pl.plot(line, line, c = 'k', linestyle = '--')
xlabel('Predicted',fontsize = 15)
ylabel('Observed',fontsize = 15)
# +
# queries the entire features table to calculate pct_flux
ids_to_see = dfcol_into_sqllist(df_unbal_stations, 'id')
query = """
SELECT a.id, a.date, a.hour, bikes_out, bikes_in, dayofweek, month, is_weekday,
is_holiday, rebal_net_flux, tot_docks, avail_bikes, avail_docks,
precip, snow, temp, c.long, c.lat
FROM features a
LEFT JOIN weather b ON a.date = b.date AND a.hour = b.hour
LEFT JOIN stations c ON a.id = c.id
WHERE tot_docks > 0 AND a.id in {}
ORDER BY a.id, a.date, a.hour;
""".format(ids_to_see)
df = pd.read_sql_query(query, con)
df.date = pd.to_datetime(df.date)
df = new_features(df)
# +
# histogram of pct_flux compared to normal distribution
from scipy.stats import norm
x_axis = np.arange(-1,1,0.001)
df.pct_flux.plot(kind = 'hist', logy = True, bins=400, normed = True, alpha = 0.5)
plot(x_axis, norm.pdf(x_axis, df.pct_flux.mean(), df.pct_flux.std()))
xlim([-0.5,0.5])
ylim([0.001, None])
xlabel('Bike Flow', fontsize = 15)
ylabel('Frequency', fontsize = 15)
# +
# df['hours12'] = pd.to_datetime(df.hour, format='%H').dt.strftime('%I %p')
cols = ['pct_flux', 'pct_avail_bikes', 'hour']
weekday = 1
dock_id = 477
grps = df.groupby(['id','is_weekday', 'hour']).mean().reset_index()
cond = (grps.is_weekday == weekday) & (grps.id == dock_id)
plotter = grps[cond][cols].sort_values('hour').set_index('hour')
# plot1 = plotter['pct_avail_bikes'].plot(c = 'steelblue', label = 'Available Bikes')
# change ticklabels
# labels = [item.get_text() for item in ax.get_xticklabels()]
labels = ['12 AM', '5 AM', '10 AM', '3 PM', '8 PM']
plot1.set_xticklabels(labels)
plot2 = plotter['pct_flux'].plot(c = 'r', label = 'Flow')
legend(loc = 4, prop = {'size':15})
xlabel('Time of Day', fontsize = 15)
ylim([-.45, 0.7])
station_name = stations2015[stations2015.id == dock_id].name.iloc[0]
suptitle('Bike Activity for Station: {} (Weekday)'.format(station_name))
# -
stations2015[stations2015.id == dock_id].name.iloc[0]
# +
def flux_by_hour(df, cols, dock_id, day = 0, month = 1):
grp_cols = ['id','month','dayofweek', 'hour']
grps = df.groupby(grp_cols).mean().reset_index()
if month:
cond = (grps.dayofweek == day) & (grps.month == month) & (grps.id == dock_id)
else:
cond = (grps.dayofweek == day) & (grps.id == dock_id)
return grps[cond].set_index('hour')[cols]
def plot_by_hour(df, cols, dock_id, day = 0, month = 1):
df_hour = flux_by_hour(df, cols, dock_id, day = day, month = month)
df_hour.plot()
#plot formatting
labels = ['12 AM', '5 AM', '10 AM', '3 PM', '8 PM']
plot1.set_xticklabels(labels)
plt.xlabel('Time of Day', size = 15)
legend(loc = 4, prop = {'size':15})
# -
unbal_stations_list = df_unbal_stations[df_unbal_stations.id.isin(stations2015.id)].id
for id in list(unbal_stations_list)[0:5]:
plot_by_hour(df,['pct_flux', 'pct_avail_bikes'], id, day=3)
suptitle('Bike Activity for station {}'.format(id))
flux_profile = plot_by_hour(df,['pct_flux'], 477, day = 1, month=3)
# flux_profile
# +
aggregators = {'bikes_in': 'sum', 'bikes_out': 'sum', 'long': 'max', 'lat': 'max'}
df_morn = df_unbal_stations_byhr[df_unbal_stations_byhr.hour.isin([7,8,9])].groupby('id').agg(aggregators)
# +
# map plot of difference between 8 am and 6 pm bike activity
aggregators = {'bikes_in': 'sum', 'bikes_out': 'sum'}
morn_cond = df_unbal_stations_byhr.hour.isin([8])
even_cond = df_unbal_stations_byhr.hour.isin([17])
grp_cols = ['id', 'lat', 'long', 'name']
df_morn = df_unbal_stations_byhr[morn_cond].groupby(grp_cols).agg(aggregators).reset_index()
df_even = df_unbal_stations_byhr[even_cond].groupby(grp_cols).agg(aggregators).reset_index()
fig = plt.figure(figsize = (15,15))
pl1 = fig.add_subplot(111)
pl1.scatter(df_morn.long, df_morn.lat,
s = df_morn.bikes_out/50, color = 'r', alpha=0.9,
label = 'bikes out at 8 am')
pl1.scatter(df_even.long, df_even.lat,
s = df_even.bikes_out/50, color = 'g', alpha=0.6,
label = 'bikes out at 6 pm')
plt.axes().set_aspect('equal')
plt.legend(loc='best')
# +
import folium
# norm = Normalize(start_station['trip count'].min(), start_station['trip count'].max())
# Get dark tileset from CartoBD (https://cartodb.com/basemaps)
tileset = r'http://{s}.basemaps.cartocdn.com/light_all/{z}/{x}/{y}.png'
station_map = folium.Map(location = [40.73, -73.985], width = 400, height = 700,
tiles = tileset,
attr = '© <a href="http://www.openstreetmap.org/copyright">OpenStreetMap</a> contributors, © <a href="http://cartodb.com/attributions">CartoDB</a>',
zoom_start = 13)
for index, row in df_morn.iterrows():
morn_color = 'red' #rgb2hex(cm.YlOrRd(norm(row['trip count'])))
folium.CircleMarker(
location = [row['lat'], row['long']],
popup = row['name'],
radius = sqrt(row['bikes_out'])/15,
color = None, fill_color = morn_color).add_to(station_map)
for index, row in df_even.iterrows():
green_color = 'green' #rgb2hex(cm.YlOrRd(norm(row['trip count'])))
folium.CircleMarker(
location = [row['lat'], row['long']],
popup = row['name'],
radius = sqrt(row['bikes_out'])/15,
color = None, fill_color = green_color).add_to(station_map)
station_map
# station_map.save('station_map.html')
# -
# +
# map plot of flow activity vs. rebalancing activity
fig = plt.figure(figsize = (10,15))
pl1 = fig.add_subplot(111)
pl1.scatter(df_unbal_stations.long, df_unbal_stations.lat,
s = df_unbal_stations.flux/25, color = 'k', alpha=0.9,
label = 'total bike flow')
pl1.scatter(df_unbal_stations.long, df_unbal_stations.lat,
s = df_unbal_stations.rflux/25, color = 'y', alpha=0.6,
label = 'total rebalancing flow')
plt.axes().set_aspect('equal')
plt.legend(loc='best')
# -
# ## Sample Queries
# +
# find the most active stations by bikes_out
query_stations_out = """
SELECT a.id, bikes_out, c.name, neighborhood, borough
FROM (SELECT id, sum(bikes_out) as bikes_out
FROM features
GROUP BY id) a
JOIN neighborhoods b ON a.id = b.id
JOIN stations c on b.id = c.id
ORDER BY bikes_out DESC;
"""
df_stations_out = pd.read_sql_query(query_stations_out, con)
df_stations_out.head(20)
# +
# find the most active stations by bikes_in
query_stations_in = """
SELECT a.id, bikes_in, c.name, neighborhood, borough
FROM (SELECT id, sum(bikes_in) as bikes_in
FROM features
GROUP BY id) a
JOIN neighborhoods b ON a.id = b.id
JOIN stations c ON b.id = c.id
ORDER BY bikes_in DESC;
"""
df_stations_in = pd.read_sql_query(query_stations_in, con)
df_stations_in.head(20)
# +
# find the most active neighborhoods by bikes_out
query_hoods_out = """
SELECT sum(a.bikes_out) as bikes_out,
b.neighborhood, b.borough
FROM features a
JOIN neighborhoods b on a.id = b.id
GROUP BY borough, neighborhood
ORDER BY bikes_out DESC;
"""
df_hoods_out = pd.read_sql_query(query_hoods_out, con)
df_hoods_out.head(20)
# +
# find the most active neighborhoods by bikes_in
query_hoods_in = """
SELECT sum(a.bikes_in) as bikes_in,
b.neighborhood, b.borough
FROM features a
JOIN neighborhoods b on a.id = b.id
GROUP BY borough, neighborhood
ORDER BY bikes_in DESC;
"""
df_hoods_in = pd.read_sql_query(query_hoods_in, con)
df_hoods_in.head(20)
# +
# find the most unbalanced neighborhoods
query_hoods_in = """
SELECT sum(a.bikes_in - a.bikes_out) as flux,
b.neighborhood, b.borough
FROM features a
JOIN neighborhoods b on a.id = b.id
GROUP BY borough, neighborhood
ORDER BY flux DESC;
"""
df_hoods_in = pd.read_sql_query(query_hoods_in, con)
df_hoods_in.head(20)
# -
| modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Qiskit Syntax
# ### Installation
#
# Qiskit is a package in Python for doing everything you'll ever need with quantum computing.
#
# If you don't have it already, you need to install it. Once it is installed, you need to import it.
#
# There are generally two steps to installing Qiskit. The first one is to install Anaconda, a python package that comes with almost all dependencies that you will ever need. Once you've done this, Qiskit can then be installed by running the command
# ```
# pip install qiskit
# ```
# in your terminal. For detailed installation instructions, refer to [the documentation page here](https://qiskit.org/documentation/install.html).
#
# **Note: The rest of this section is intended for people who already know the fundamental concepts of quantum computing.** It can be used by readers who wish to skip straight to the later chapters in which those concepts are put to use. All other readers should read the [Introduction to Python and Jupyter notebooks](../ch-prerequisites/python-and-jupyter-notebooks.html), and then move on directly to the start of [Chapter 1](../ch-states/introduction.html).
# ### Quantum circuits
# The object at the heart of Qiskit is the quantum circuit. Here's how we create one, which we will call `qc`
from qiskit import QuantumCircuit
qc = QuantumCircuit()
# This circuit is currently completely empty, with no qubits and no outputs.
# ### Quantum registers
# To make the circuit less trivial, we need to define a register of qubits. This is done using a `QuantumRegister` object. For example, let's define a register consisting of two qubits and call it `qr`.
from qiskit import QuantumRegister
qr = QuantumRegister(2,'a')
# Giving it a label like `'a'` is optional.
#
# Now we can add it to the circuit using the `add_register` method, and see that it has been added by checking the `qregs` variable of the circuit object. This guide uses [Jupyter Notebooks](https://jupyter.org/). In Jupyter Notebooks, the output of the last line of a cell is displayed below the cell:
# +
qc.add_register( qr )
qc.qregs
# -
# Now our circuit has some qubits, we can use another attribute of the circuit to see what it looks like: <code>draw()</code>.
qc.draw()
# Our qubits are ready to begin their journey, but are currently just sitting there in state $\left|0\right\rangle$.
# #### Applying Gates
# To make something happen, we need to add gates. For example, let's try out <code>h()</code>.
# + tags=["raises-exception"]
qc.h()
# -
# Here we got an error, because we didn't tell the operation which qubit it should act on. The two qubits in our register `qr` can be individually addressed as `qr[0]` and `qr[1]`.
qc.h(qr[0])
# Ignore the output in the above. When the last line of a cell has no `=`, Jupyter notebooks like to print out what is there. In this case, it's telling us that there is a Hadamard as defined by Qiskit. To suppress this output, we could use a <code>;</code>.
#
# We can also add a controlled-NOT using `cx`. This requires two arguments: control qubit, and then target qubit.
qc.cx(qr[0], qr[1])
# Now our circuit has more to show
qc.draw()
# ### Statevector simulator
# We are now at the stage that we can actually look at an output from the circuit. Specifically, we will use the 'statevector simulator' to see what is happening to the state vector of the two qubits.
#
# To get this simulator ready to go, we use the following line.
from qiskit import Aer
sv_sim = Aer.get_backend('statevector_simulator')
# In Qiskit, we use *backend* to refer to the things on which quantum programs actually run (simulators or real quantum devices). To set up a job for a backend, we need to set up the corresponding backend object.
#
# The simulator we want is defined in the part of qiskit known as `Aer`. By giving the name of the simulator we want to the <code>get_backend()</code> method of Aer, we get the backend object we need. In this case, the name is `'statevector_simulator'`.
#
# A list of all possible simulators in Aer can be found using
for backend in Aer.backends():
print(backend)
# All of these simulators are 'local', meaning that they run on the machine on which Qiskit is installed. Using them on your own machine can be done without signing up to the IBMQ user agreement.
#
# To run this simulation, we need to assemble the circuit into a `Qobj` which contains the circuit, as well as other information about how to run the experiment (for example how many times we should run the circuit), but we will ignore these other options here.
#
# We then use the <code>.run()</code> method of the backend we want to use (in this case a simulator) to run the experiment. This is where the quantum computations happen!
from qiskit import assemble
qobj = assemble(qc)
job = sv_sim.run(qobj)
# This creates an object that handles the job, which here has been called `job`. All we need from this is to extract the result. Specifically, we want the state vector.
ket = job.result().get_statevector()
for amplitude in ket:
print(amplitude)
# This is the vector for a Bell state $\left( \left|00\right\rangle + \left|11\right\rangle \right)/\sqrt{2}$, which is what we'd expect given the circuit.
# While we have a nicely defined state vector, we can show another feature of Qiskit: it is possible to initialize a circuit with an arbitrary pure state.
# +
new_qc = QuantumCircuit(qr)
new_qc.initialize(ket, qr)
# -
# ### Classical registers and the qasm simulator
# In the above simulation, we got out a statevector. That's not what we'd get from a real quantum computer. For that we need measurement. And to handle measurement we need to define where the results will go. This is done with a `ClassicalRegister`. Let's define a two bit classical register, in order to measure both of our two qubits.
# +
from qiskit import ClassicalRegister
cr = ClassicalRegister(2,'creg')
qc.add_register(cr)
# -
# Now we can use the `measure` method of the quantum circuit. This requires two arguments: the qubit being measured, and the bit where the result is written.
#
# Let's measure both qubits, and write their results in different bits.
# +
qc.measure(qr[0],cr[0])
qc.measure(qr[1],cr[1])
qc.draw()
# -
# Now we can run this on a local simulator whose effect is to emulate a real quantum device. For this we need to add another input to the `assemble` function, `shots`, which determines how many times we run the circuit to take statistics. If you don't provide any `shots` value, you get the default of 1024.
qasm_sim = Aer.get_backend('qasm_simulator')
qobj = assemble(qc, shots=8192)
job = qasm_sim.run(qobj)
# The result is essentially a histogram in the form of a Python dictionary. We can use `print` to display this for us.
hist = job.result().get_counts()
print(hist)
# We can even get Qiskit to plot it as a histogram.
# +
from qiskit.visualization import plot_histogram
plot_histogram(hist)
# -
# For compatible backends we can also ask for and get the ordered list of results.
qobj = assemble(qc, shots=10)
job = qasm_sim.run(qobj, memory=True)
samples = job.result().get_memory()
print(samples)
# Note that the bits are labelled from right to left. So `cr[0]` is the one to the furthest right, and so on. As an example of this, here's an 8 qubit circuit with a Pauli $X$ on only the qubit numbered `7`, which has its output stored to the bit numbered `7`.
# +
qubit = QuantumRegister(8)
bit = ClassicalRegister(8)
qc_2 = QuantumCircuit(qubit,bit)
qc_2.x(qubit[7])
qc_2.measure(qubit,bit) # this is a way to do all the qc.measure(qr8[j],cr8[j]) at once
qobj = assemble(qc_2, shots=8192)
qasm_sim.run(qobj).result().get_counts()
# -
# The `1` appears at the left.
#
# This numbering reflects the role of the bits when they represent an integer.
#
#
#
# $$ b_{n-1} ~ b_{n-2} ~ \ldots ~ b_1 ~ b_0 = \sum_j ~ b_j ~ 2^j $$
#
#
#
# So the string we get in our result is the binary for $2^7$ because it has a `1` for the bit numbered `7`.
# ### Simplified notation
# Multiple quantum and classical registers can be added to a circuit. However, if we need no more than one of each, we can use a simplified notation.
#
# For example, consider the following.
qc = QuantumCircuit(3)
# The single argument to `QuantumCircuit` is interpreted as the number of qubits we want. So this circuit is one that has a single quantum register consisting of three qubits, and no classical register.
#
# When adding gates, we can then refer to the three qubits simply by their index: 0, 1 or 2. For example, here's a Hadamard on qubit 1.
# +
qc.h(1)
qc.draw()
# -
# To define a circuit with both quantum and classical registers, we can supply two arguments to `QuantumCircuit`. The first will be interpreted as the number of qubits, and the second will be the number of bits. For example, here's a two qubit circuit for which we'll take a single bit of output.
qc = QuantumCircuit(2,1)
# To see this in action, here is a simple circuit. Note that, when making a measurement, we also refer to the bits in the classical register by index.
# +
qc.h(0)
qc.cx(0,1)
qc.measure(1,0)
qc.draw()
# -
# ### Creating custom gates
# As we've seen, it is possible to combine different circuits to make bigger ones. We can also use a more sophisticated version of this to make custom gates. For example, here is a circuit that implements a `cx` between qubits 0 and 2, using qubit 1 to mediate the process.
# +
sub_circuit = QuantumCircuit(3, name='toggle_cx')
sub_circuit.cx(0,1)
sub_circuit.cx(1,2)
sub_circuit.cx(0,1)
sub_circuit.cx(1,2)
sub_circuit.draw()
# -
# We can now turn this into a gate
toggle_cx = sub_circuit.to_instruction()
# and then insert it into other circuits using any set of qubits we choose
# +
qr = QuantumRegister(4)
new_qc = QuantumCircuit(qr)
new_qc.append(toggle_cx, [qr[1],qr[2],qr[3]])
new_qc.draw()
# -
# ### Accessing on real quantum hardware
# Backend objects can also be set up using the `IBMQ` package. The use of these requires us to [sign with an IBMQ account](https://qiskit.org/documentation/install.html#access-ibm-q-systems). Assuming the credentials are already loaded onto your computer, you sign in with
# + tags=["uses-hardware"]
from qiskit import IBMQ
IBMQ.load_account()
# -
# Now let's see what additional backends we have available.
# + tags=["uses-hardware"]
provider = IBMQ.get_provider(hub='ibm-q')
for backend in provider.backends():
print(backend)
# -
# Here there is one simulator, but the rest are prototype quantum devices.
#
# We can see what they are up to with the <code>status()</code> method.
# + tags=["uses-hardware"]
for backend in provider.backends():
print(backend.status().to_dict())
# -
# Let's get the backend object for the largest public device.
# + tags=["uses-hardware"]
real_device = provider.get_backend('ibmq_16_melbourne')
# -
# We can use this to run a job on the device in exactly the same way as for the emulator.
#
# We can also extract some of its properties.
# + tags=["uses-hardware"]
properties = real_device.properties()
coupling_map = real_device.configuration().coupling_map
# -
# From this we can construct a noise model to mimic the noise on the device (we will discuss noise models further later in the textbook).
# + tags=["uses-hardware"]
from qiskit.test.mock import FakeAthens
athens = FakeAthens()
# -
# And then run the job on the emulator, with it reproducing all these features of the real device. Here's an example with a circuit that should output `'10'` in the noiseless case.
# + tags=["uses-hardware"]
qc = QuantumCircuit(5,5)
qc.x(0)
for q in range(4):
qc.cx(0,q+1)
qc.measure_all()
qc.draw()
# -
from qiskit.visualization import plot_gate_map
plot_gate_map(athens)
from qiskit import transpile
t_qc = transpile(qc, athens)
t_qc.draw()
# Now the very basics have been covered, let's learn more about what qubits and quantum circuits are all about.
qobj = assemble(t_qc)
counts = athens.run(qobj).result().get_counts()
plot_histogram(counts)
import qiskit
qiskit.__qiskit_version__
| notebooks/ch-appendix/qiskit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # DAE solver
#
# In this notebook, we show some examples of solving a DAE model. For the purposes of this example, we use the CasADi solver, but the syntax remains the same for other solvers
# +
# %pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import tests
import numpy as np
import os
import matplotlib.pyplot as plt
from pprint import pprint
os.chdir(pybamm.__path__[0]+'/..')
# Create solver
dae_solver = pybamm.CasadiSolver(mode="safe") # use safe mode so that solver stops at events
# -
# ## Integrating DAEs
# In PyBaMM, a model is solved by calling a solver with solve. This sets up the model to be solved, and then calls the method `_integrate`, which is specific to each solver. We begin by setting up and discretising a model
# +
# Create model
model = pybamm.BaseModel()
u = pybamm.Variable("u")
v = pybamm.Variable("v")
model.rhs = {u: -v} # du/dt = -v
model.algebraic = {v: 2 * u - v} # 2*v = u
model.initial_conditions = {u: 1, v: 2}
model.variables = {"u": u, "v": v}
# Discretise using default discretisation
disc = pybamm.Discretisation()
disc.process_model(model);
# -
# Now the model can be solved by calling `solver.solve` with a specific time vector at which to evaluate the solution
# +
# Solve #################################
t_eval = np.linspace(0, 2, 30)
solution = dae_solver.solve(model, t_eval)
#########################################
# Extract u and v
t_sol = solution.t
u = solution["u"]
v = solution["v"]
# Plot
t_fine = np.linspace(0,t_eval[-1],1000)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(13,4))
ax1.plot(t_fine, np.exp(-2 * t_fine), t_sol, u(t_sol), "o")
ax1.set_xlabel("t")
ax1.legend(["exp(-2*t)", "u"], loc="best")
ax2.plot(t_fine, 2 * np.exp(-2 * t_fine), t_sol, v(t_sol), "o")
ax2.set_xlabel("t")
ax2.legend(["2*exp(-2*t)", "v"], loc="best")
plt.tight_layout()
plt.show()
# -
# Note that, where possible, the solver makes use of the mass matrix and jacobian for the model. However, the discretisation or solver will have created the mass matrix and jacobian algorithmically, using the expression tree, so we do not need to calculate and input these manually.
# The solution terminates at the final simulation time:
solution.termination
# ### Events
#
# It is possible to specify events at which a solution should terminate. This is done by adding events to the `model.events` dictionary. In the following example, we solve the same model as before but add a termination event when `v=-0.2`.
# +
# Create model
model = pybamm.BaseModel()
u = pybamm.Variable("u")
v = pybamm.Variable("v")
model.rhs = {u: -v} # du/dt = -v
model.algebraic = {v: 2 * u - v} # 2*v = u
model.initial_conditions = {u: 1, v: 2}
model.events.append(pybamm.Event('v=0.2', v - 0.2)) # adding event here
model.variables = {"u": u, "v": v}
# Discretise using default discretisation
disc = pybamm.Discretisation()
disc.process_model(model)
# Solve #################################
t_eval = np.linspace(0, 2, 30)
solution = dae_solver.solve(model, t_eval)
#########################################
# Extract u and v
t_sol = solution.t
u = solution["u"]
v = solution["v"]
# Plot
t_fine = np.linspace(0,t_eval[-1],1000)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(13,4))
ax1.plot(t_fine, np.exp(-2 * t_fine), t_sol, u(t_sol), "o")
ax1.set_xlabel("t")
ax1.legend(["exp(-2*t)", "u"], loc="best")
ax2.plot(t_fine, 2 * np.exp(-2 * t_fine), t_sol, v(t_sol), "o", t_fine, 0.2 * np.ones_like(t_fine), "k")
ax2.set_xlabel("t")
ax2.legend(["2*exp(-2*t)", "v", "v = 0.2"], loc="best")
plt.tight_layout()
plt.show()
# -
# Now the solution terminates because the event has been reached
solution.termination
# ## Finding consistent initial conditions
#
# The solver will fail if initial conditions that are inconsistent with the algebraic equations are provided. However, before solving the DAE solvers automatically use `_set_initial_conditions` to obtain consistent initial conditions, starting from a guess of bad initial conditions, using a simple root-finding algorithm.
# +
# Create model
model = pybamm.BaseModel()
u = pybamm.Variable("u")
v = pybamm.Variable("v")
model.rhs = {u: -v} # du/dt = -v
model.algebraic = {v: 2 * u - v} # 2*v = u
model.initial_conditions = {u: 1, v: 1} # bad initial conditions, solver fixes
model.events.append(pybamm.Event('v=0.2', v - 0.2))
model.variables = {"u": u, "v": v}
# Discretise using default discretisation
disc = pybamm.Discretisation()
disc.process_model(model)
print(f"y0_guess={model.concatenated_initial_conditions.evaluate().flatten()}")
dae_solver.set_up(model)
dae_solver._set_initial_conditions(model, {}, True)
print(f"y0_fixed={model.y0}")
# -
# ## References
#
# The relevant papers for this notebook are:
pybamm.print_citations()
| examples/notebooks/solvers/dae-solver.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 420115
# # ปฏิบัติการฟิสิกส์อย่างสังเขป
# + [markdown] slideshow={"slide_type": "slide"}
#
# # การทดลองที่3
# ## การวัดความเร่งเนื่องจากแรงโน้มถ่วง
# + [markdown] slideshow={"slide_type": "slide"}
# ## วัตถุประสงค์
#
# 1. เพื่อศึกษาการแกว่งของลูกตุ้มนาฬิกาอย่างง่ายและลุกตุ้มนาฬิกาฟิสิกัล
#
# 2. เพื่อวัดค่าความเร่งโน้มถ่วงของโลก จากคาบการแกว่งของลูกตุ้มนาฬิกาอย่างง่าย
#
# 3. เพื่อวัดค่าความเร่งโน้มถ่วงของโลก จากคาบการแกว่งของลูกตุ้มนาฬิกาฟิสิกัล
# + [markdown] slideshow={"slide_type": "slide"}
# # การเคลื่อนที่แบบฮาร์มอนิก
#
# - การเคลื่อนที่แบบฮาร์มอนิกเป็นการเคลื่อนที่กลับไปกลับมาผ่านตำแหน่งสมดุล โดย
# การกระจัดของวัตถุเป็นฟังก์ชันไซน์หรือโคไซน์ ซึ่งเป็นฟังก์ชันฮาร์โมนิก จึงเรียกว่าการเคลื่อนที่
# แบบฮาร์โมนิก การเคลื่อนที่ที่ง่ายที่สุดของการเคลื่อนที่แบบฮาร์มอนิก เรียกว่า การเคลื่อนที่
# แบบฮาร์มอนิก อย่างง่าย
#
#
# <img src="figures/bull-dog-in-a-swing1.jpg" width="35%" height="35%" align="left">
# <img src="figures/Panthéon_Pendule_de_Foucault2.JPG" width="35%" height="35%" align="left">
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### ตอนที่1: ลูกตุ้มนาฬิกาอย่างง่าย(A Simple Pendulum)
#
# - วัดค่าความเร่งโน้มถ่วงของโลก,g , จากคาบการแกว่งของลูกตุ้มนาฬิกาอย่างง่าย
# <img src="figures/pendulo2.gif" width="40%" height="40%">
#
# + [markdown] colab_type="text" id="haQCFeQUKWmA" slideshow={"slide_type": "slide"}
# ## ทฤษฎี
# - ลูกตุ้มนาฬิกาอย่างง่าย เป็นโมเดลทางคณิตศาสตตร์ ประกอบด้วยลุุูกตุ้มมวล m ขนาดเล็กมากๆ (จุด)ห้อยแขวนไว้ด้วยเชือกที่ไม่มีมวล
# ถ้าดึงลูกตุ้มให้เคลื่อนที่ออกไปจากตำแหน่งสมดุล จนเส้นเชือกทำมุม θ น้อยๆ กับ
# แนวดิ่งแล้วปล่อย ลูกตุ้มจะเคลื่อนที่กลับไปกลับมาผ่านตำแหน่งสมดุล โดยมีคาบการแกว่งคงที่
#
# <img src="figures/Spendulum.jpeg.jpg" width="30%" height="30%">
#
# คาบเวลาของการแกว่ง,T ($s$), จะมีค่าตามความสัมพันธ์:
#
# >$T~=~2~\pi~\sqrt{\frac{L}{g}} $
#
# เมื่อ **g** เป็นค่าความเร่งจากแรงโน้มถ่วงของโลก ($m/s^{2}$)
# และ **L** เป็นความยาวของเส้นเชือก ($m$)
#
#
# >$T^2~=~4~\pi^2\frac{L}{g} $
#
# หากเขียนความสัมพันธ์โดยใช้กราฟเส้นตรงที่มี ความชัน=$slope$, $y=t^2$ และ $x=L$
#
# >$y~=~slope~x $
#
#
# เมื่อเปรียบเทียบสมการข้างบน ค่าความเร่งจากแรงโน้มถ่วง, g, จะหาได้จาก
#
#
# >$g~=~4\pi^2\left ({\frac{L}{T^2}} \right ) ~=~ \frac{ 4 \pi^2}{slope} $
#
# >$slope~=~\frac{T^2}{L} $
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### อุปกรณ์การทดลอง
# <img src="figures/IMG_6655.jpg" width="40%" height="30%" >
#
# 1. เสาตั้ง
# 2. ลูกตุ้ม
# 3. เชือก
# 4. ไม้เมตร
# 5. นาฬิกาจับเวลา
# 6. ฐานตั้งและแกนหมุน
# 7. ไม้เมตรที่เจาะรู
# 8. เครื่องชั่งมวล
# + [markdown] slideshow={"slide_type": "slide"}
# ### วิธีการทดลอง
# <img src="figures/procedure01.jpg" width="40%" height="30%" >
#
# 1. จัดอุปกรณ์ การทดลองดังรูป
# 2. วัดความยาวเชือกที่แขวนลูกตุ้ม ( $L$ ) (ความยาวเชือกบวกกับรัศมีของลูกตุ้ม)โดยเริ่มที่ 60 cm
# 3. จัดให้ลูกตุ้มแกว่งโดยมีมุมแกว่งไม่เกิน 10 องศา
# 4. จับเวลาที่ลูกตุ้มแกว่งครบ 10 รอบ คำนวณค่าคาบการแกว่งเฉลี่ยของลูกตุ้ม ( $T$ )
# 5. ทำการทดลองซ้ำข้อ 2-4 โดยเปลี่ยนค่าความยาวของลูกตุ้ม ที่ความยาว 70, 80, 90 และ 100 cm
# 6. คำนวณหาค่าคาบกำลังสอง ( $T^2$) เชียนกราฟเส้นตรงระหว่าง $T^2$ (แกนตั้ง) และ $L^2$ (แกนนอน) ในกระดาษกราฟหรือในโปรแกรม
# 7. หาค่าความชันของกราฟและนำไปนวณหาค่า g และ หาเปอร์เซนต์ความความคลาดเคลื่อน
# 8. สรุปผลการทดลอง
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### ผลการทดลอง: ภาพแสดงนาฬิกาจับเวลา เมื่อลูกคุ้มเคลื่อนที่ไป 10 รอบของการแกว่ง
#
# - ท่ี่ความยาวเชือก $L=60.00$ cm
# <img src="figures/resultp1/tsp60cm.png" width="40%" height="40%">
#
# - ท่ี่ความยาวเชือก $L=70.00$ cm
# <img src="figures/resultp1/tsp70cm.png" width="40%" height="40%">
#
# - ท่ี่ความยาวเชือก $L=80.00$ cm
# <img src="figures/resultp1/tsp80cm.png" width="40%" height="40%">
#
# - ท่ี่ความยาวเชือก $L=90.00$ cm
# <img src="figures/resultp1/tsp90cm.png" width="40%" height="40%">
#
# - ท่ี่ความยาวเชือก $L=100.00$ cm
# <img src="figures/resultp1/tsp100cm.png" width="40%" height="40%">
# + [markdown] slideshow={"slide_type": "slide"}
# ### ตารางบันทึกผล
#
# <img src="figures/datatablepart1.png" width="ุุุ30%" height="30%">
#
# + slideshow={"slide_type": "skip"}
import numpy as np
from IPython.display import display, HTML
import pandas as pd
an_array=np.full((5), [' '],dtype=str)#initialize empty chararray
df = pd.DataFrame({"ความยาว L (m)":0.10*np.arange(5)+0.60,
#"เวลาในการแกว่ง10รอบ t(s)":[14.48,16.78,17.97,18.60,19.65],
"เวลาในการแกว่ง10รอบ t(s)":[15.6,16.43,17.68,18.94,19.93],
"คาบการแกว่ง T":an_array,
r" $T^2$":an_array})
display(HTML(df.to_html(index=False)))
# + slideshow={"slide_type": "skip"}
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
#import seaborn as sns
T=0.1*df["เวลาในการแกว่ง10รอบ t(s)"].values
y=T**2
x= df["ความยาว L (m)"].values
x_new = np.linspace(0, 1.1, 100)
model = LinearRegression()
model.fit(x.reshape(-1, 1), y.reshape(-1, 1))
print('Intercept:', model.intercept_)
print('slope:',model.coef_)
print('g:',(4*np.pi**2)/model.coef_)
def plotLab3():
fig, ax = plt.subplots(1,1,figsize=(8,6))
ax.scatter(x, y, c="r", s=50,alpha=0.3,linewidths=2, edgecolors='r')
ax.plot(x_new, model.predict(x_new[:, np.newaxis]))
ax.set_title(r'Plot of pendulum length vs time period squared' ,fontsize=20)
ax.set_xlabel('Length (m)')
ax.set_ylabel(r' $T^2$ $s^2$' ,fontsize=20)
ax.text(0.6, 1.0, r'$slope= ...$ ', fontsize=15)
ax.text(0.71, 0.2, r'$g= ...$ ', fontsize=15)
#data=pd.DataFrame()
#sns.regplot(x="total_bill", y="tip", data=tips);
# + slideshow={"slide_type": "slide"}
# ตัวอย่างการเขียนกราฟ การทดลอง
plotLab3()
# + colab_type="text" id="mx6yjSBGAm6x" slideshow={"slide_type": "skip"}
from IPython.display import YouTubeVideo
id=["c7b5qhX7pSQ","QIILmDwgaVM","KQcMaotEznc","JPaXWr39zXk","PILcMx23O24"]
# + slideshow={"slide_type": "slide"}
# สาธิตปฏิบัติการที่ 3 รายวิชา 420115 ตอนที่ 1.1
YouTubeVideo(id[0],560,315,rel=0)
# + slideshow={"slide_type": "slide"}
#สาธิตปฏิบัติการที่ ตอนที่ 1.2(การแกว่งที่ระยะ 60 cm)
YouTubeVideo(id[1],560,315,rel=0)
# + [markdown] slideshow={"slide_type": "slide"}
# #### ตอนที่2: ลูกตุ้มนาฬิกาฟิสิกัล(Physical Pendulum)
#
# - วัดค่าความเร่งจากแรงโน้มถ่วงของโลก,g ,จากคาบการแกว่งของลูกตุ้มนาฬิกาฟิสิกัล
# <img src="figures/swing.gif" width="50%" height="50%">
# + [markdown] slideshow={"slide_type": "slide"}
# ## ทฤษฏี
# - ใช้อธิบายการแกว่งของวัตถุแข็งเกร็ง ที่ไม่สามารถโมเดลด้วยจุดมวล m หากแขวนวัตถุที่จุดหมุน O ขณะอยู่นิ่ง จุดศูนย์กลางมวลของวัตถุ (จุด C.M.) จะอยู่ในแนวดิ่งใต้จุด O เป็นระยะ h และโมเมนต์ความเฉื่อยของวัตถุรอบแกนที่ผ่านจุด O เป็น I
#
# <img src="figures/Ppendulum.jpg" width="30%" height="30%">
#
# หากเราแกว่งลูกตุ้มนาฬิกาฟิสิกัลเป็นมุมเล็กๆ คาบเวลาของการแกว่ง,T ($s$), จะมีค่าตามความสัมพันธ์:
#
# >$T~=~2~\pi~\sqrt{\frac{I}{mgh}} $
#
# - $g$ เป็นค่าความเร่งจากแรงโน้มถ่วงของโลก ($m/s^{2}$)
# - $I$ เป็นโมเมนต์ความเฉื่อยของวัตถุ ($kg m^2$)
# - $h$ เป็น ระยะหว่างจุดหมุน O และ จุดศูนย์กลางมวลของวัตถุ($m$)
#
# >$T^2~=~4~\pi^2\frac{I}{mgh} $
#
# ค่าความเร่งจากแรงโน้มถ่วง, g, จะหาได้จาก
# >$g~=~4\pi^2\left ({\frac{I}{mT^2h}} \right )$
#
# โมเมนต์ความเฉื่อยของไม้เมตรยาว L รอบแกนที่ผ่านจุด O คำนวณได้จาก
#
# >$I~=~ I_{CM} + mh^2$
#
# >$I~=~ \frac{1}{12}mL^2 + mh^2$
#
# เมื่อ $I_{CM}~=~ \frac{1}{12}mL^2$คือโมเมนต์ความเฉื่อยของไม้เมตรยาว L รอบแกนที่ผ่านจุดจุดศูนย์กลางมวล
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### วิธีการทดลอง
# <img src="figures/figp2config.jpg" width="40%" height="20%" >
#
# 1. ชั่งมวลของไม้เมตร
# 2. ระยะห่างจากจุดหมุน(รูบนไม้เมตร)ถึงจุดกึ่งกลางของไม้เมตร(จุดที่ความยาว 50 cmเป็นระยะ h
# 3. จัดอุปกรณ์ การทดลองดังรูป
# 4. จัดให้ไม้เมตรแกว่งโดยมีมุมแกว่งไม่เกิน 10 องศา
# 5. จับเวลาที่ไม้เมตรแกว่งครบ 10 รอบ คำนวณค่าคาบการแกว่งเฉลี่ยของลูกตุ้ม (T)
# 6. คํานวณหาค่า g และ หาเปอร์เซนต์ความแตกต่างจากค่า g ที่ได้จากการทดลองในตอนที่ 1
# 8. สรุปผลการทดลอง
#
# + [markdown] slideshow={"slide_type": "slide"}
# #### ผลการทดลอง ตอนที่ 2
#
# - ภาพเครื่องชั่งมวลของไม้เมตร
# <img src="figures/resultp2/weight_metrestick.jpg" width="40%" height="40%">
# - ภาพจุดที่เจาะรูสำหรับแขวนของไม้เมตร (จุดหมุนม ใช้สำหรับคำนาณค่า h)
# <img src="figures/resultp2/meterstickpivot.jpg" width="40%" height="40%">
# - ภาพแสดงนาฬิกาจับเวลา เมื่อไม้เมตรเคลื่อนที่ไป 10 รอบของการแกว่ง
# <img src="figures/resultp2/tphysicalpendulum1568.jpg" width="40%" height="40%">
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### ตารางบันทึกผล ตอนที่ 2
#
# <img src="figures/datatablepart2.png" width="ุุุ55%" height="55%">
# + [markdown] slideshow={"slide_type": "slide"}
# [hi]<iframe width="560" height="315" src="https://www.youtube.com/embed/c7b5qhX7pSQ" frameborder="0" allow="accelerometer; autoplay; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
# + slideshow={"slide_type": "skip"}
# สาธิตปฏิบัติการที่ ตอนที่ 2.1
YouTubeVideo(id[2],560,315,rel=0)
# + slideshow={"slide_type": "skip"}
# สาธิตปฏิบัติการที่ 3 ตอนที่ 2.2
YouTubeVideo(id[3],560,315,rel=0)
# + slideshow={"slide_type": "skip"}
# สาธิตปฏิบัติการที่ 3 ตอนที่ 2.3 (การวัดมวลไม้เมตร)
YouTubeVideo(id[4],560,315,rel=0)
# +
#jupyter nbconvert DJExp3revised.ipynb --to slide --to slides --post serve --SlidesExporter.reveal_scroll=True --SlidesExporter.reveal_theme=sky
| Lab3/DJExp3revised.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercícios de Classificação
# **Lista 3**
#
# Para este exercício, será utilizado o dataset de sobreviventes do titanic. Os dados já
# encontram-se separados em arquivos de treino (train.csv) e teste (test.csv). Neste dataset,
# encontram-se informações como sexo, idade, classe socio-ecônomica, entre outras. Abaixo,
# você encontra o dicionário dos dados, contendo a descrição do que cada atributo e seus
# respectivos valores representam.
# 1. Execute uma análise exploratória dos dados. Elabore gráficos, calcule estatísticas e
# obtenha inferências iniciais sobre os dados. Discorra sobre as inferências realizadas.
#
# 2. Realize o pré-processamento dos dados. Faça as limpezas e formatações que julgar
# necessárias para obter um conjunto de dados consistente. (Dica: você pode juntar os
# dois arquivos de dados em um único dataframe para facilitar a manipulação!).
#
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import sklearn
import seaborn as sns
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn import linear_model as lm
from sklearn import metrics
from sklearn.metrics import r2_score, mean_squared_error, accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split, KFold, cross_val_score, cross_val_predict
from sklearn.naive_bayes import BernoulliNB, MultinomialNB
import matplotlib.pyplot as plt
# %matplotlib inline
#from pandas_profiling import ProfileReport
# **Carrengando o dataset de treino**
df_test = pd.read_csv('https://raw.githubusercontent.com/hinessacaminha/mentoring-ml/main/exercicios/classificacao/test.csv')
df_test.head()
df_train = pd.read_csv('https://raw.githubusercontent.com/hinessacaminha/mentoring-ml/main/exercicios/classificacao/train.csv')
df_train.head()
# **Concatenando os dois DFs (df_test e df_train)**
# Nesta etapa foi necessário juntar os DFs para melhorar a exploração e limpeza dos dados. Para isso, utilizamos a função do pandas `concat` e o parâmetro `ignore_index` com valor `True` para garantir que não haveria a repetição dos índices . Observa-se que nessa junção a coluna `Survived` ficou à direta e com vários valores NaN, isto ocorreu porque esta coluna não constava no `df_test` e os valores de suas linhas foram separados das colunas, significa dizer que parte dos seus valores estão nas linhas e parte nas colunas. Provavelmente esta não é a melhor solução para junção destes Dfs. Em todo caso iremos tratar os NaN das colunas e continuar com essa abordagem.
df_unido = pd.concat([df_test, df_train], ignore_index=True)
df_unido.head()
# **Estudar como usa o Profiling**
#
# *Visão Geral do DF com `pandas_profiling`*
#
# *Função do Python que exibe todas as estatísticas*
# +
#profile = ProfileReport(df_unido)
#profile.to_notebook_iframe()
# -
# exibindo as colunas do DF após unir o df_test e df_train
df_unido.columns
# +
# Definindo a coluna PassengerId como índice
#df = df_unido.set_index('PassengerId', inplace=False) # coloquei a coluna PassengerId como índice
#df
# -
df_unido['Survived']
# porque estes dados ficaram como float e não inteiro?
# observando o arquivo de treino todas as células estão preenchidas o que não justifica a presença desses NaN que são interpretados
# desta forma quando as células não possuem valores.
# verificando se há NaN e somando todos por coluna
df_unido.isnull().sum()
# *Calculando o percentual de NaN nas colunas em relação ao total de linhas. Isto ajuda a verificar o impacto da ausência
# (caso opte por dropar) na análise dessas informações para a regra de negócio.*
# Soma dos valores NaN em relação as linhas (por isso o shape está setado em 0), multiplicado por 100
df_unido.isnull().sum()/df_unido.shape[0]*100
# *A coluna `Survived` é importante para a análise da questão 3, logo precisamos ter cuidado ao decidir o que será feito com os NaN. Provavelmente não deverá ser eliminada e deveremos optar pela a técnica de "imputação múltipla" que consite em substituir o NaN por uma valor mais recorrente na coluna do DF*
# **Eliminando colunas**
#
# *No caso desta coluna o mais recomendado seria eleminá-la uma vez que tem mais de 70% de NaN. Substituir os valores não seria uma boa opção, pois iria apenas "mascarar" as informações desta feature e poderia prejudicar as predições*
df_unido.drop(columns = ['Ticket','Fare','Cabin','Name'], inplace = True)
df_unido.head()
df_unido.info()
# **Tratando os NaN**
#
# *Utilizamos a técnica de imputação única através da função `SimpleImputer` da biblioteca do Sklearn com o parâmetro `strategy` definido por `mean` ,isto é, média. Para isso precisamos importar no módulo `impute` o `SimpleImputer`.*
# +
from sklearn.impute import SimpleImputer
# substituição dos valores NaN pela estratégia da média para a coluna Survived
most_imputer = SimpleImputer(missing_values=np.nan, strategy='most_frequent')
mean_imputer = SimpleImputer(missing_values=np.nan, strategy='mean')
# -
# *Para aplicar a estratégia `SimpleImputer` para a substituição na coluna é necessário fazer a combinação com `fit_transform`*
# substituição dos valores NaN pela estratégia da média combinado com fit_transform
df_unido.Survived = most_imputer.fit_transform(df_unido[['Survived']])
df_unido.Age = mean_imputer.fit_transform(df_unido[['Age']])
# substituição dos valores NaN pela estratégia da média para a coluna Embarked, pois é a estratégia recomendada para feature categórica
most_imputer = SimpleImputer(missing_values=np.nan, strategy='most_frequent')
df_unido.Embarked = most_imputer.fit_transform(df_unido[['Embarked']])
# o sample também funciona como o .head(), porém as linhas são aleatórias ao contrário do primeiro que me retorna as 5 primeiras linhas do DF.
df_unido.sample(5)
# **Análise do uso do SimpleImputer**
#
# *A estratégia da média no caso da coluna Survived não é a mais indicada, uma vez que irá gerar uma terceira coluna ao plotar o histograma na análise de sobreviventes por sexo. Além de preencher com valores muito que irão gerar erros e não informarão corretamente, pois não existe por exemplo o valor de 0.38 de vida ou morte, isto é ou é 0 (morto) ou é 1(vivo). Com isso, optei por mudar para a estratégia de mais frequente para esta feature. Se observarmos a plotagem do novo histograma veremos que a estratégia de `most_frequent` mostrou melhor análise e não gerou uma terceira coluna com os dados separados da feature `Suvived`*
# *Removendo a linha que contém o NaN na coluna Fare. Como é apenas um único dado e não ultrapassa de 10% da base de dados
# então este procedimento não se torna prejudicial para as análises*
# Armazenando em uma variável a remoção da linha que contém o NaN. O inplace setado como "False" para não aplicar a remoção no df_unido
df_clean = df_unido.dropna(axis=0, inplace = False)
df_unido.head()
# Calculando o percentual de NaN no df_clean, somente para visualizar se a remoção da linha que tinha NaN deu certo.
df_clean.isnull().sum()/df_clean.shape[0]*100
df_unido.dtypes
# ## Análise Exploratória dos Dados (AED)
df_unido.dtypes
df_unido.duplicated().value_counts()
df_unido.info()
df_unido.describe()
df_unido.corr('pearson')
df_unido['Age'].plot.hist(edgecolor='black', title= 'Sobreviventes por idade')
df_unido['Pclass'].plot.hist(edgecolor='black', title= 'Sobreviventes por classe (1ª, 2ª, 3ª)', color= 'coral')
df_unido['SibSp'].plot.hist(edgecolor='black', title= 'Sobreviventes por Quantidade de irmãos/cônjuges a bordo do Titanic', color= 'pink')
#gráfico de dispersão
plt.scatter(df_unido['Pclass'], df_unido['Survived'], color= 'green')
plt.scatter(df_unido['Age'], df_unido['Survived'], color = 'Plum')
plt.scatter(df_unido['SibSp'], df_unido['Survived'], color = 'yellow')
plt.scatter(df_unido['Parch'], df_unido['Survived'], color = 'CornflowerBlue')
# agrupando por sexo
colunas_selecionadas = ['Sex', 'Survived']
df_titanic = df_unido.filter(items=colunas_selecionadas)
df_titanic.head()
df_titanic.groupby(by=["Sex"]).count()
# Mulherese x homens que sobreviveram
df_titanic.groupby(by=["Sex"]).count().hist()
df_titanic.groupby(by=["Sex"]).describe()
# +
# transformando os dados das colunas Sex, Embarked em int ou float
# -
# **Atribuindo valores numéricos à coluna "Sex"**
# 1 = male
#
# 2 = female
df_unido.loc[df_unido.Sex=='male','Sex'] = 1
df_unido.loc[df_unido.Sex=='female','Sex'] = 2
df_unido['Sex'].plot.hist(edgecolor='black', title= 'Sobreviventes por Sexo', color= 'blue')
# **Atribuindo valores numéricos à coluna "Embarked"**
# 3 = C - Cherbourg
#
# 4 = Q - Queenstown
#
# 5 = S - Southampton
df_unido.loc[df_unido.Embarked=='S','Embarked'] = 3
df_unido.loc[df_unido.Embarked=='C','Embarked'] = 4
df_unido.loc[df_unido.Embarked=='Q','Embarked'] = 5
df_unido.head()
# **Verificando os tipos de dados das colunas**
df_unido.dtypes
df_unido['Embarked'].plot.hist(edgecolor='black', title= 'Sobreviventes por Porto de Embarque', color= 'purple')
# *Segundo o histograma acima a maior quantidade de sobreviventes do Titanic partiu do porto de `Cherbourg (3.00)`*
# **Tranformando os dados das colunas Sex, Embarked e Survived (`float`) em dados do tipo `int`**
df_unido['Sex']= df_unido['Sex'].astype(int)
df_unido['Embarked'] = df_unido['Embarked'].astype(int)
df_unido['Survived'] = df_unido['Survived'].astype(int)
df_unido
df_unido.dtypes()
#
# **Dividindo o dataset em treino e teste**
# Separando as colunas exceto a Survived que será o y
X = df_unido[df_unido.columns[:-1]]
X
# Separando os dados que irei estimar
y = df_unido[df_unido.columns[-1:]]
y
[X_train, X_test, y_train, y_test]= train_test_split(X, y,test_size = 0.3, random_state=2)
X_test
# Padronizando os dados em uma mesma escala
scaler = MinMaxScaler()
scaler.fit(X_train, y_train)
scaled = scaler.transform(X_test)
X_test
# 3. Crie um classificador para predizer se um passageiro sobreviveu ou não a partir dos
# atributos presentes no dataset. Utilize os algoritmos KNN, Regressão Logística e
# Naive Bayes para criar os modelos. Crie um modelo para cada algoritmo.
# ## **Regressão Logística**
log_reg = LogisticRegression()
model_reg = log_reg.fit(X_train,y_train)
pred_log_reg = model_reg.predict(X_test)
print("Regressão Logística:", model_reg.score(X_test, y_test))
# ## **KNN**
knn = KNeighborsClassifier(n_neighbors=5)
model_knn = knn.fit(X_train,y_train)
pred_knn = model_knn.predict(X_test)
print("KNN:", model_knn.score(X_test, y_test))
# **Acurácia**
accuracy_score(y_test, pred_log_reg)
# **Kappa**
print("Kappa:", metrics.cohen_kappa_score(y_test, pred_log_reg))
# **F1**
print("Todas:", metrics.precision_recall_fscore_support(y_test, pred_log_reg))
# ## **Naive Bayes**
naive = MultinomialNB()
model_naive = naive.fit(X_train,y_train)
pred_naive = model_naive.predict(X_test)
print("Naive Bayes:", model_naive.score(X_test, y_test))
# ## **Matriz de Confusão**
matrix = metrics.confusion_matrix(y_test, pred_log_reg)
matrix_bayes = metrics.confusion_matrix(y_test, pred_naive)
matrix_knn = metrics.confusion_matrix(y_test, pred_knn)
plt.figure(figsize=(9,9))
sns.heatmap(matrix, annot=True, fmt=".3f", linewidths=.5, square = True, cmap = 'Blues_r');
plt.ylabel('Actual label');
plt.xlabel('Predicted label');
all_sample_title = 'Accuracy Score: {0}'.format(model_reg.score(X_test, y_test))
plt.title(all_sample_title, size = 15);
plt.figure(figsize=(9,9))
sns.heatmap(matrix_bayes, annot=True, fmt=".3f", linewidths=.5, square = True, cmap = 'Blues_r');
plt.ylabel('Actual label');
plt.xlabel('Predicted label');
all_sample_title = 'Accuracy Score: {0}'.format(model_naive.score(X_test, y_test))
plt.title(all_sample_title, size = 15);
plt.figure(figsize=(9,9))
sns.heatmap(matrix_knn, annot=True, fmt=".3f", linewidths=.5, square = True, cmap = 'Blues_r');
plt.ylabel('Actual label');
plt.xlabel('Predicted label');
all_sample_title = 'Accuracy Score: {0}'.format(model_knn.score(X_test, y_test))
plt.title(all_sample_title, size = 15);
# **Referências Bibliográficas**
#
# https://minerandodados.com.br/analise-de-dados-com-python-usando-pandas/
#
# https://medium.com/data-hackers/tratamento-e-transforma%C3%A7%C3%A3o-de-dados-nan-uma-vis%C3%A3o-geral-e-pr%C3%A1tica-54efa9fc7a98
#
# https://www.youtube.com/watch?v=ojA65o8N0iM&t=684s
#
# https://datatofish.com/check-nan-pandas-dataframe/
#
# http://www.each.usp.br/lauretto/SIN5008_2011/aula01/aula1#:~:text=A%20%EF%AC%81nalidade%20da%20An%C3%A1lise%20Explorat%C3%B3ria%20de%20Dados%20%28AED%29,digita%C3%A7%C3%A3o%20de%20dados%20em%20um%20banco%20de%20dados
#
# https://pypi.org/project/pandas-profiling/
#
# https://www.youtube.com/watch?v=TRatkPjzHNE
#
# https://minerandodados.com.br/validacao-cruzada-aprenda-de-forma-simples-como-usar-essa-tecnica/#:~:text=Uma%20das%20maneiras%20de%20fazer%20a%20divis%C3%A3o%20desses,dos%20dados%20para%20treino%20e%2030%25%20para%20teste.
#
# https://minerandodados.com.br/7-tipos-graficos-cientista-de-dados/
#
# https://inferir.com.br/artigos/algoritimo-knn-para-classificacao/
#
# https://www.datacamp.com/community/tutorials/k-nearest-neighbor-classification-scikit-learn
| notebooks/lista_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/Tessellate-Imaging/monk_v1/blob/master/study_roadmaps/3_image_processing_deep_learning_roadmap/3_deep_learning_advanced/1_Blocks%20in%20Deep%20Learning%20Networks/11)%20Mobilenet%20V2%20Linear%20Bottleneck%20Block.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Goals
#
# ### 1. Learn to implement Mobilenet V2 Linear Bottleneck Block using monk
# - Monk's Keras
# - Monk's Pytorch
# - Monk's Mxnet
#
# ### 2. Use network Monk's debugger to create complex blocks
#
#
# ### 3. Understand how syntactically different it is to implement the same using
# - Traditional Keras
# - Traditional Pytorch
# - Traditional Mxnet
# # Mobilenet V2 Linear Bottleneck Block
#
# - Note: The block structure can have variations too, this is just an example
from IPython.display import Image
Image(filename='imgs/mobilenet_v2_linear_bottleneck.png')
# # Table of contents
#
# [1. Install Monk](#1)
#
#
# [2. Block basic Information](#2)
#
# - [2.1) Visual structure](#2-1)
#
# - [2.2) Layers in Branches](#2-2)
#
#
# [3) Creating Block using monk visual debugger](#3)
#
# - [3.1) Create the first branch](#3-1)
#
# - [3.2) Create the second branch](#3-2)
#
# - [3.3) Merge the branches](#3-3)
#
# - [3.4) Debug the merged network](#3-4)
#
# - [3.5) Compile the network](#3-5)
#
# - [3.6) Visualize the network](#3-6)
#
# - [3.7) Run data through the network](#3-7)
#
#
# [4) Creating Block Using MONK one line API call](#4)
#
# - [Mxnet Backend](#4-1)
#
# - [Pytorch Backend](#4-2)
#
# - [Keras Backend](#4-3)
#
#
#
# [5) Appendix](#5)
#
# - [Study Material](#5-1)
#
# - [Creating block using traditional Mxnet](#5-2)
#
# - [Creating block using traditional Pytorch](#5-3)
#
# - [Creating block using traditional Keras](#5-4)
#
# <a id='0'></a>
# # Install Monk
# ## Using pip (Recommended)
#
# - colab (gpu)
# - All bakcends: `pip install -U monk-colab`
#
#
# - kaggle (gpu)
# - All backends: `pip install -U monk-kaggle`
#
#
# - cuda 10.2
# - All backends: `pip install -U monk-cuda102`
# - Gluon bakcned: `pip install -U monk-gluon-cuda102`
# - Pytorch backend: `pip install -U monk-pytorch-cuda102`
# - Keras backend: `pip install -U monk-keras-cuda102`
#
#
# - cuda 10.1
# - All backend: `pip install -U monk-cuda101`
# - Gluon bakcned: `pip install -U monk-gluon-cuda101`
# - Pytorch backend: `pip install -U monk-pytorch-cuda101`
# - Keras backend: `pip install -U monk-keras-cuda101`
#
#
# - cuda 10.0
# - All backend: `pip install -U monk-cuda100`
# - Gluon bakcned: `pip install -U monk-gluon-cuda100`
# - Pytorch backend: `pip install -U monk-pytorch-cuda100`
# - Keras backend: `pip install -U monk-keras-cuda100`
#
#
# - cuda 9.2
# - All backend: `pip install -U monk-cuda92`
# - Gluon bakcned: `pip install -U monk-gluon-cuda92`
# - Pytorch backend: `pip install -U monk-pytorch-cuda92`
# - Keras backend: `pip install -U monk-keras-cuda92`
#
#
# - cuda 9.0
# - All backend: `pip install -U monk-cuda90`
# - Gluon bakcned: `pip install -U monk-gluon-cuda90`
# - Pytorch backend: `pip install -U monk-pytorch-cuda90`
# - Keras backend: `pip install -U monk-keras-cuda90`
#
#
# - cpu
# - All backend: `pip install -U monk-cpu`
# - Gluon bakcned: `pip install -U monk-gluon-cpu`
# - Pytorch backend: `pip install -U monk-pytorch-cpu`
# - Keras backend: `pip install -U monk-keras-cpu`
# ## Install Monk Manually (Not recommended)
#
# ### Step 1: Clone the library
# - git clone https://github.com/Tessellate-Imaging/monk_v1.git
#
#
#
#
# ### Step 2: Install requirements
# - Linux
# - Cuda 9.0
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu90.txt`
# - Cuda 9.2
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu92.txt`
# - Cuda 10.0
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu100.txt`
# - Cuda 10.1
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu101.txt`
# - Cuda 10.2
# - `cd monk_v1/installation/Linux && pip install -r requirements_cu102.txt`
# - CPU (Non gpu system)
# - `cd monk_v1/installation/Linux && pip install -r requirements_cpu.txt`
#
#
# - Windows
# - Cuda 9.0 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu90.txt`
# - Cuda 9.2 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu92.txt`
# - Cuda 10.0 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu100.txt`
# - Cuda 10.1 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu101.txt`
# - Cuda 10.2 (Experimental support)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cu102.txt`
# - CPU (Non gpu system)
# - `cd monk_v1/installation/Windows && pip install -r requirements_cpu.txt`
#
#
# - Mac
# - CPU (Non gpu system)
# - `cd monk_v1/installation/Mac && pip install -r requirements_cpu.txt`
#
#
# - Misc
# - Colab (GPU)
# - `cd monk_v1/installation/Misc && pip install -r requirements_colab.txt`
# - Kaggle (GPU)
# - `cd monk_v1/installation/Misc && pip install -r requirements_kaggle.txt`
#
#
#
# ### Step 3: Add to system path (Required for every terminal or kernel run)
# - `import sys`
# - `sys.path.append("monk_v1/");`
# # Imports
# Common
import numpy as np
import math
import netron
from collections import OrderedDict
from functools import partial
# +
#Using mxnet-gluon backend
# When installed using pip
from monk.gluon_prototype import prototype
# When installed manually (Uncomment the following)
#import os
#import sys
#sys.path.append("monk_v1/");
#sys.path.append("monk_v1/monk/");
#from monk.gluon_prototype import prototype
# -
# <a id='2'></a>
# # Block Information
# <a id='2_1'></a>
# ## Visual structure
from IPython.display import Image
Image(filename='imgs/mobilenet_v2_linear_bottleneck.png')
# <a id='2_2'></a>
# ## Layers in Branches
#
# - Number of branches: 2
#
#
# - Branch 1
# - identity
#
#
# - Branch 2
# - conv_1x1 -> batchnorm -> relu -> conv_3x3 -> batchnorm -> relu -> conv_1x1 -> batchnorm
#
#
# - Branches merged using
# - Elementwise addition
#
#
# (See Appendix to read blogs on mobilenet-v2)
# <a id='3'></a>
# # Creating Block using monk debugger
# +
# Imports and setup a project
# To use pytorch backend - replace gluon_prototype with pytorch_prototype
# To use keras backend - replace gluon_prototype with keras_prototype
from monk.gluon_prototype import prototype
# Create a sample project
gtf = prototype(verbose=1);
gtf.Prototype("sample-project-1", "sample-experiment-1");
# -
# <a id='3-1'></a>
# ## Create the first branch
def first_branch():
network = [];
network.append(gtf.identity());
return network;
# Debug the branch
branch_1 = first_branch()
network = [];
network.append(branch_1);
gtf.debug_custom_model_design(network);
# <a id='3-2'></a>
# ## Create the second branch
def second_branch(output_channels=128, bottleneck_width=4, stride=1):
network = [];
network.append(gtf.convolution(output_channels=output_channels*bottleneck_width,
kernel_size=1, stride=1));
network.append(gtf.batch_normalization());
network.append(gtf.relu());
network.append(gtf.convolution(output_channels=output_channels*bottleneck_width,
kernel_size=3, stride=stride));
network.append(gtf.batch_normalization());
network.append(gtf.relu());
network.append(gtf.convolution(output_channels=output_channels, kernel_size=1, stride=1));
network.append(gtf.batch_normalization());
return network;
# Debug the branch
branch_2 = second_branch(output_channels=128, stride=1)
network = [];
network.append(branch_2);
gtf.debug_custom_model_design(network);
# <a id='3-3'></a>
# ## Merge the branches
# +
def final_block(output_channels=128, bottleneck_width=4, stride=1):
network = [];
#Create subnetwork and add branches
subnetwork = [];
branch_1 = first_branch()
branch_2 = second_branch(output_channels=output_channels,
bottleneck_width=bottleneck_width,
stride=stride)
subnetwork.append(branch_1);
subnetwork.append(branch_2);
# Add merging element
subnetwork.append(gtf.add());
# Add the subnetwork
network.append(subnetwork)
return network;
# -
# <a id='3-4'></a>
# ## Debug the merged network
final = final_block(output_channels=128, stride=1)
network = [];
network.append(final);
gtf.debug_custom_model_design(network);
# <a id='3-5'></a>
# ## Compile the network
gtf.Compile_Network(network, data_shape=(128, 224, 224), use_gpu=False);
# <a id='3-6'></a>
# ## Run data through the network
import mxnet as mx
x = np.zeros((1, 128, 224, 224));
x = mx.nd.array(x);
y = gtf.system_dict["local"]["model"].forward(x);
print(x.shape, y.shape)
# <a id='3-7'></a>
# ## Visualize network using netron
gtf.Visualize_With_Netron(data_shape=(128, 224, 224))
# <a id='4'></a>
# # Creating Using MONK LOW code API
# <a id='4-1'></a>
# ## Mxnet backend
# +
from monk.gluon_prototype import prototype
gtf = prototype(verbose=1);
gtf.Prototype("sample-project-1", "sample-experiment-1");
network = [];
# Single line addition of blocks
network.append(gtf.mobilenet_v2_linear_bottleneck_block(output_channels=128, bottleneck_width=4));
gtf.Compile_Network(network, data_shape=(128, 224, 224), use_gpu=False);
# -
# <a id='4-2'></a>
# ## Pytorch backend
#
# - Only the import changes
# +
#Change gluon_prototype to pytorch_prototype
from monk.pytorch_prototype import prototype
gtf = prototype(verbose=1);
gtf.Prototype("sample-project-1", "sample-experiment-1");
network = [];
# Single line addition of blocks
network.append(gtf.mobilenet_v2_linear_bottleneck_block(output_channels=128, bottleneck_width=4));
gtf.Compile_Network(network, data_shape=(128, 224, 224), use_gpu=False);
# -
# <a id='4-3'></a>
# ## Keras backend
#
# - Only the import changes
# +
#Change gluon_prototype to keras_prototype
from monk.keras_prototype import prototype
gtf = prototype(verbose=1);
gtf.Prototype("sample-project-1", "sample-experiment-1");
network = [];
# Single line addition of blocks
network.append(gtf.mobilenet_v2_linear_bottleneck_block(output_channels=128, bottleneck_width=4));
gtf.Compile_Network(network, data_shape=(128, 224, 224), use_gpu=False);
# -
# <a id='5'></a>
# # Appendix
# <a id='5-1'></a>
# ## Study links
# - https://ai.googleblog.com/2018/04/mobilenetv2-next-generation-of-on.html
# - https://machinethink.net/blog/mobilenet-v2/
# - https://towardsdatascience.com/review-mobilenetv2-light-weight-model-image-classification-8febb490e61c
# - https://towardsdatascience.com/mobilenetv2-inverted-residuals-and-linear-bottlenecks-8a4362f4ffd5
# - https://medium.com/@luis_gonzales/a-look-at-mobilenetv2-inverted-residuals-and-linear-bottlenecks-d49f85c12423
# <a id='5-2'></a>
# ## Creating block using traditional Mxnet
#
# - Code credits - https://mxnet.incubator.apache.org/
# Traditional-Mxnet-gluon
import mxnet as mx
from mxnet.gluon import nn
from mxnet.gluon.nn import HybridBlock, BatchNorm
from mxnet.gluon.contrib.nn import HybridConcurrent, Identity
from mxnet import gluon, init, nd
# +
def _add_conv(out, channels=1, kernel=1, stride=1, pad=0,
num_group=1, active=True, relu6=False, norm_layer=BatchNorm, norm_kwargs=None):
out.add(nn.Conv2D(channels, kernel, stride, pad, groups=num_group, use_bias=False))
out.add(norm_layer(scale=True, **({} if norm_kwargs is None else norm_kwargs)))
if active:
out.add(nn.Activation('relu'))
class LinearBottleneck(nn.HybridBlock):
def __init__(self, in_channels, channels, t, stride,
norm_layer=BatchNorm, norm_kwargs=None, **kwargs):
super(LinearBottleneck, self).__init__(**kwargs)
self.use_shortcut = stride == 1 and in_channels == channels
with self.name_scope():
self.out = nn.HybridSequential()
if t != 1:
_add_conv(self.out,
in_channels * t,
relu6=True,
norm_layer=norm_layer, norm_kwargs=norm_kwargs)
_add_conv(self.out,
in_channels * t,
kernel=3,
stride=stride,
pad=1,
num_group=in_channels * t,
relu6=True,
norm_layer=norm_layer, norm_kwargs=norm_kwargs)
_add_conv(self.out,
channels,
active=False,
relu6=True,
norm_layer=norm_layer, norm_kwargs=norm_kwargs)
def hybrid_forward(self, F, x):
out = self.out(x)
if self.use_shortcut:
out = F.elemwise_add(out, x)
return out
# +
# Invoke the block
block = LinearBottleneck(64, 64, 4, 1)
# Initialize network and load block on machine
ctx = [mx.cpu()];
block.initialize(init.Xavier(), ctx = ctx);
block.collect_params().reset_ctx(ctx)
block.hybridize()
# Run data through network
x = np.zeros((1, 64, 224, 224));
x = mx.nd.array(x);
y = block.forward(x);
print(x.shape, y.shape)
# Export Model to Load on Netron
block.export("final", epoch=0);
netron.start("final-symbol.json", port=8082)
# -
# <a id='5-3'></a>
# ## Creating block using traditional Pytorch
#
# - Code credits - https://github.com/pytorch/vision/blob/master/torchvision/models/mobilenet.py
# Traiditional-Pytorch
import torch
from torch import nn
from torch.jit.annotations import List
import torch.nn.functional as F
# +
class ConvBNReLU(nn.Sequential):
def __init__(self, in_planes, out_planes, kernel_size=3, stride=1, groups=1):
padding = (kernel_size - 1) // 2
super(ConvBNReLU, self).__init__(
nn.Conv2d(in_planes, out_planes, kernel_size, stride, padding, groups=groups, bias=False),
nn.BatchNorm2d(out_planes),
nn.ReLU6(inplace=True)
)
class InvertedResidual(nn.Module):
def __init__(self, inp, oup, stride, expand_ratio):
super(InvertedResidual, self).__init__()
self.stride = stride
assert stride in [1, 2]
hidden_dim = int(round(inp * expand_ratio))
self.use_res_connect = self.stride == 1 and inp == oup
layers = []
if expand_ratio != 1:
# pw
layers.append(ConvBNReLU(inp, hidden_dim, kernel_size=1))
layers.extend([
# dw
ConvBNReLU(hidden_dim, hidden_dim, stride=stride, groups=hidden_dim),
# pw-linear
nn.Conv2d(hidden_dim, oup, 1, 1, 0, bias=False),
nn.BatchNorm2d(oup),
])
self.conv = nn.Sequential(*layers)
def forward(self, x):
if self.use_res_connect:
return x + self.conv(x)
else:
return self.conv(x)
# +
# Invoke the block
block = InvertedResidual(64, 64, 1, 4);
# Initialize network and load block on machine
layers = []
layers.append(block);
net = nn.Sequential(*layers);
# Run data through network
x = torch.randn(1, 64, 224, 224)
y = net(x)
print(x.shape, y.shape);
# Export Model to Load on Netron
torch.onnx.export(net, # model being run
x, # model input (or a tuple for multiple inputs)
"model.onnx", # where to save the model (can be a file or file-like object)
export_params=True, # store the trained parameter weights inside the model file
opset_version=10, # the ONNX version to export the model to
do_constant_folding=True, # whether to execute constant folding for optimization
input_names = ['input'], # the model's input names
output_names = ['output'], # the model's output names
dynamic_axes={'input' : {0 : 'batch_size'}, # variable lenght axes
'output' : {0 : 'batch_size'}})
netron.start('model.onnx', port=9998);
# -
# <a id='5-4'></a>
# ## Creating block using traditional Keras
#
# - Code credits: https://github.com/xiaochus/MobileNetV2/blob/master/mobilenet_v2.py
# Traditional-Keras
import keras
import keras.layers as kla
import keras.models as kmo
import tensorflow as tf
from keras.models import Model
backend = 'channels_last'
from keras import layers
from keras.layers import *
from keras import backend as K
# +
def relu6(x):
return K.relu(x, max_value=6.0)
def _conv_block(inputs, filters, kernel, strides):
channel_axis = 1 if K.image_data_format() == 'channels_first' else -1
x = Conv2D(filters, kernel, padding='same', strides=strides)(inputs)
x = BatchNormalization(axis=channel_axis)(x)
return Activation(relu6)(x)
def _bottleneck(inputs, filters, kernel, t, alpha, s, r=False):
channel_axis = 1 if K.image_data_format() == 'channels_first' else -1
# Depth
tchannel = K.int_shape(inputs)[channel_axis] * t
# Width
cchannel = int(filters * alpha)
x = _conv_block(inputs, tchannel, (1, 1), (1, 1))
x = DepthwiseConv2D(kernel, strides=(s, s), depth_multiplier=1, padding='same')(x)
x = BatchNormalization(axis=channel_axis)(x)
x = Activation(relu6)(x)
x = Conv2D(cchannel, (1, 1), strides=(1, 1), padding='same')(x)
x = BatchNormalization(axis=channel_axis)(x)
if r:
x = Add()([x, inputs])
return x
def create_model(input_shape, kernel_size, filters, bottleneck_width, stride=1):
img_input = layers.Input(shape=input_shape);
x = _bottleneck(img_input, filters, kernel_size, bottleneck_width, 1, stride, r=True)
return Model(img_input, x);
# +
# Invoke the block
kernel_size=3;
filters=64;
input_shape=(224, 224, 64);
model = create_model(input_shape, kernel_size, filters, 4);
# Run data through network
x = tf.placeholder(tf.float32, shape=(1, 224, 224, 64))
y = model(x)
print(x.shape, y.shape)
# Export Model to Load on Netron
model.save("final.h5");
netron.start("final.h5", port=8082)
# -
# # Goals Completed
#
# ### 1. Learn to implement Mobilenet V2 Linear Bottleneck Block using monk
# - Monk's Keras
# - Monk's Pytorch
# - Monk's Mxnet
#
# ### 2. Use network Monk's debugger to create complex blocks
#
#
# ### 3. Understand how syntactically different it is to implement the same using
# - Traditional Keras
# - Traditional Pytorch
# - Traditional Mxnet
| study_roadmaps/3_image_processing_deep_learning_roadmap/3_deep_learning_advanced/1_Blocks in Deep Learning Networks/11) Mobilenet V2 Linear Bottleneck Block.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import gym
import tensorflow as tf
env = gym.make('CartPole-v0')
print("Action space =", env.action_space)
print("Observation space =", env.observation_space)
print("Action space dimension =", env.action_space.n)
print("Observation space dimension =", env.observation_space.shape[0])
# +
model = tf.keras.Sequential([
tf.keras.layers.Dense(64, activation='relu',
input_shape=[env.observation_space.shape[0]]),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(env.action_space.n, activation="softmax")
])
model.summary()
| Exercise02/Exercise02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math, time, random, datetime
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('seaborn-whitegrid')
# %matplotlib inline
# +
# Loading data
import sys
sys.path.append('../data')
from dataset_pointers import graph_nodes, graph_edges
nodes = pd.read_csv(graph_nodes, low_memory=False)
edges = pd.read_csv(graph_edges, low_memory=False, index_col='edge_id')
no_nan = pd.read_csv('../data/raw/no_nan_data.csv', low_memory=False, index_col='node_id')
# -
clean_sample = no_nan.sample(n=10000)
clean = clean_sample.drop(['Account ID String', 'Address', 'Name'], axis=1)
clean.head()
clean[clean.isnull().any(1)]
clean = clean.drop('testingFlag', axis=1)
clean.head()
clean.ExtendedCaseGraphID.value_counts()[:4]
clean.CoreCaseGraphID.value_counts()[:4]
# +
# CLASSES ARE EXTREMELY UNBALANCED -> EXPECT SHITTY RESULT
# 300'000 nodes with case=0 ...
# -
| notebooks/initial_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ctarrington/try-colab/blob/master/normal.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="YMrBxSprx_wB" colab_type="code" colab={}
from math import factorial
import numpy as np
from scipy.stats import norm
from matplotlib.pyplot import plot, fill_between
# + id="YaiwPS-JzqLN" colab_type="code" colab={}
points_to_18 = np.linspace(10, 18)
points = np.linspace(10, 30)
# + [markdown] id="HuihgBiOyuiW" colab_type="text"
# ## probability that t < 18 when $ \mu_t = 20.6 $ and $ \sigma = 1.62 $
# + id="S39svYeaydB7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 299} outputId="38265247-7c37-473b-f3d0-9ccfe45590ff"
rv = norm(20.6, 1.62)
print('probability that t < 18 is', rv.cdf(18))
plot(points, rv.pdf(points))
fill_between(points_to_18, rv.pdf(points_to_18))
# + id="OgIuP-YRzJUT" colab_type="code" colab={}
| normal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../') # or just install the module
sys.path.append('../../fuzzy-tools') # or just install the module
sys.path.append('../../astro-lightcurves-handler') # or just install the module
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import numpy as np
import fuzzytools.files as ftfiles
from fuzzytools.datascience.cms import ConfusionMatrix
from fuzzytools.matplotlib.cm_plots import plot_custom_confusion_matrix
import matplotlib.pyplot as plt
from fuzzytools.datascience.xerror import XError
from IPython.display import display
from fuzzytools.strings import bf_alphabet_count
def get_kf_text(set_name, kf):
if kf=='.':
return f'($\\mu\\pm\\sigma$ from 5-fold cross-validation in {set_name}-set)'
else:
return f'($\\mu\\pm\\sigma$ from {kf} k-fold in {set_name}-set)'
def get_mode_text(mode, features):
if mode=='sne':
return f'Using {len(features)} SNe selected astrophysical features [{mode}]'
else:
return f'Using {len(features)} astrophysical features [{mode}]'
mode = 'all' # all spm sne
methods = [
'linear-fstw',
#'spm-mle-fstw',
#'spm-mle-estw',
#'spm-mcmc-fstw',
#'spm-mcmc-estw',
#'bspline-fstw',
]
RANDOM_STATE = 0
set_name = 'test'
kf = '.'
thday = 100
for method in methods:
print('='*30+method)
for train_config in ['r', 's', 'r+s']:
load_roodir = f'../save/exp=rf_eval~train_config={train_config}~mode={mode}/survey=alerceZTFv7.1~bands=gr~mode=onlySNe~method={method}'
print(load_roodir)
files, files_ids, kfs, all_kf_files = ftfiles.gather_files_by_kfold(load_roodir, kf, set_name,
fext='d',
disbalanced_kf_mode='oversampling', # error oversampling
random_state=RANDOM_STATE,
returns_all_kf_files=True,
)
print(f'{files_ids}({len(files_ids)}#)')
if len(files)==0:
continue
class_names = files[0]()['class_names']
features = files[0]()['features']
rank = files[0]()['rank']
for f in features:
#print(f)
pass
thdays = files[0]()['thdays']
thday = thdays[np.argmin((np.array(thdays)-thday)**2)]
xe_dict = {}
for metric_name in ['b-precision', 'b-recall', 'b-f1score']:
xe_metric = XError([f()['thdays_class_metrics_df'].loc[f()['thdays_class_metrics_df']['_thday']==thday][metric_name].item() for f in files])
xe_dict[metric_name] = xe_metric
bprecision_xe = xe_dict['b-precision']
brecall_xe = xe_dict['b-recall']
bf1score_xe = xe_dict['b-f1score']
new_order_class_names = ['SNIa', 'SNIbc', 'SNIIbn', 'SLSN']
cm = ConfusionMatrix([f()['thdays_cm'][thday] for f in files], class_names)
cm.reorder_classes(new_order_class_names)
true_label_d = {c:f'({k}#)' for c,k in zip(class_names, np.sum(files[0]()['thdays_cm'][thday], axis=1))}
### plot cm
_title = f'no-method [r]' if train_config=='r' else f'{method} [{train_config}]'
title = ''
title += f'{bf_alphabet_count(0)} train-set={_title}; features-mode={mode} ({len(features)}#)'+'\n'
#title += f'train-set={_title} - eval-set={kf}@{eval_set_name}'+'\n'
#title += f'b-p/r={bprecision_xe} / {brecall_xe}'+'\n'
#title += f'b-f1score={bf1score_xe}'+'\n'
title += f'b-Recall={brecall_xe}; b-$F_1$score={bf1score_xe}'+'\n'
title += f'th-day={thday:.3f} [days]'+'\n'
fig, ax = plot_custom_confusion_matrix(cm,
title=title[:-1],
figsize=(6,5),
true_label_d=true_label_d,
)
#accu = XError(np.diagonal(cm_norm*100, axis1=1, axis2=2).flatten().tolist())
#title += f'b-accu={accu}'+'\n'
#ax.set_title(title[:-1])
caption = ''
caption += f'Confusion matrix.'
caption += f' {get_mode_text(mode, features)} {get_kf_text(set_name, kf)}'
print(caption)
plt.show()
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
from fuzzytools.files import load_pickle, save_pickle
from fuzzytools.datascience.xerror import XError
from fuzzytools.dataframes import DFBuilder
from fuzzytools.latex.latex_tables import LatexTable
from fuzzytools.files import save_pickle
dmetrics = {
f'b-precision':{'k':1, 'mn':None},
f'b-recall':{'k':1, 'mn':None},
f'b-f1score':{'k':1, 'mn':None},
f'b-rocauc':{'k':1, 'mn':'b-AUCROC'},
#f'b-prauc':{'k':1, 'mn':'b-AUCPR'},
}
info_df = DFBuilder()
for train_config in ['r', 's', 'r+s']:
aux_r = []
for method in methods:
d = {}
for metric in dmetrics.keys():
mn = metric if dmetrics[metric]['mn'] is None else dmetrics[metric]['mn']
rootdir = f'../save/exp=rf_eval~train_config={train_config}~mode={mode}/survey=alerceZTFv7.1~bands=gr~mode=onlySNe~method={method}'
files, files_ids = gather_files_by_kfold(rootdir, kf, eval_set_name)
class_names = files[0]()['lcset_info']['class_names']
features = files[0]()['features']
metric_xe = XError([f()['metrics_dict'][metric]*dmetrics[metric]['k'] for f in files])
d[mn] = metric_xe
#print(len(metric_xe))
if not train_config=='r': # mean across methods in real case
info_df.append(f'synthetic-method={method} [{train_config}]', d)
else:
aux_r.append(d)
if train_config=='r': # mean across methods in real case
for r in aux_r:
print(r)
new_d = {k:sum([r[k] for r in aux_r]) for k in d.keys()}
info_df.append(f'synthetic-method=no-method [{train_config}]', new_d)
save_pickle(f'../temp/baseline~mode={mode}.df', info_df)
display(info_df())
ttest_metric = 'b-AUCROC'
info_df['synthetic-method=spm-mcmc-estw [r+s]'][ttest_metric].gt_ttest(info_df['synthetic-method=no-method [r]'][ttest_metric], verbose=1)
caption = ''
caption += f'Synthetic generation methods scores using only real samples [r], only synthetic samples [s], and the same proportion of real and synthetic samples [r+s].'
caption += f' {get_mode_text(mode, features)} {get_kf_text(eval_set_name, kf)}'
latex_kwargs = {
'caption':caption,
'label':'?',
'bold_axis':'columns',
}
latex_table = LatexTable(info_df(), **latex_kwargs)
print(latex_table)
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
import fuzzytools.files as ftfiles
from fuzzytools.datascience.xerror import XError
from fuzzytools.dataframes import DFBuilder
from fuzzytools.latex.latex_tables import LatexTable
info_df = DFBuilder()
train_config = 'r' # r s r+s
rank_n = 10
for k in range(0, rank_n):
d = {}
for method in methods:
rootdir = f'../save/exp=rf_eval~train_config={train_config}~mode={mode}/survey=alerceZTFv7.1~bands=gr~mode=onlySNe~method={method}'
files, files_ids = ftfiles.gather_files_by_kfold(load_roodir, kf, set_name,
fext='d',
disbalanced_kf_mode='oversampling', # error oversampling
random_state=RANDOM_STATE,
)
class_names = files[0]()['lcset_info']['class_names']
features = files[0]()['features']
rank = files[0]()['rank'] # just show one
feature_name, feature_p,_ = rank[k]
feature_name = feature_name.replace('_', '-')
d[f'{method} [{train_config}]'] = f'{feature_name} ({feature_p*100:.2f}%)'
info_df.append(f'k={k+1}', d)
caption = ''
caption += f'Random Forest astrophysical features ranking (top {rank_n}) using the same proportion of real and synthetic samples [r+s].'
caption += f' {get_mode_text(mode, features)}.'
latex_kwargs = {
'caption':caption,
'label':'?',
#'custom_tabular_align':'l|'+'c'*sum([m.split('-')[-1]=='fstw' for m in methods])+'|'+'c'*sum([m.split('-')[-1]=='estw' for m in methods]),
}
latex_table = LatexTable(info_df(), **latex_kwargs)
print(latex_table)
info_df()
# -
txt = ''
for k,idx in enumerate(rank.idxs):
f = rank.names[idx]
r = rank.values[idx]
txt += f'{k+1}. {f} ({r*100:.3f}\\%), '.replace('_', '\\textunderscore ')
if k>30-2:
break
print(txt)
| experiments/.ipynb_checkpoints/metrics_results-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/techonair/Machine-Learing-A-Z/blob/main/Model%20Selection/k-Fold%20Cross%20Verification/k_Fold_Cross_Verification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="fZ__xIu5pO1K"
# # k-Fold Cross Verification
#
# Predicting if a customer will buy a car or not based on their age and salary, data contains age salary and buy or not buy decision.
# + [markdown] id="EuSN6KblqONG"
# ## Importing libraries
# + id="b46pTxNZpE3E"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="jIcNoA6PqTOA"
# ## Importing the dataset
#
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 93} id="KrN-iTcbqdUg" outputId="92281a87-77f8-4bf3-89bc-541c9483cc9f"
from google.colab import files
files.upload()
# + id="01aLrYXTqj-l"
dataset = pd.read_csv('Social_Network_Ads.csv')
X = dataset.iloc[: , :-1].values
Y = dataset.iloc[:, -1].values
# + [markdown] id="ZFLM4ByMq67s"
# ## Spliting the data into training and test sets
# + id="5RdNm56srFRL"
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.20, random_state = 0)
# + [markdown] id="ywbp_EGBrbXZ"
# ## Feature Scaling
# + id="jFcF9HIsriU9"
from sklearn.preprocessing import StandardScaler
feat_scale = StandardScaler()
X_train = feat_scale.fit_transform(X_train)
X_test = feat_scale.fit_transform(X_test)
# + [markdown] id="2cLSdFhCslmw"
# ## Training the Kernal SVM Model on training set
# + colab={"base_uri": "https://localhost:8080/"} id="l9dEvrjDvOYC" outputId="1dc23529-c1e3-4e2a-be6d-e1c29edd34fa"
from sklearn.svm import SVC
classifier = SVC(kernel = 'rbf', random_state = 0)
classifier.fit(X_train, Y_train)
# + [markdown] id="BH-fOC1dheWS"
# ## Making Confusion Matrix
# + colab={"base_uri": "https://localhost:8080/"} id="Hp2l1iqOheo4" outputId="7cb4ba48-401c-4928-ea7d-659e0e5a8172"
from sklearn.metrics import confusion_matrix, accuracy_score
y_pred = classifier.predict(X_test)
confusionMatrix = confusion_matrix(Y_test, y_pred)
print(confusionMatrix)
accuracy_score(Y_test, y_pred)
# + [markdown] id="MZTPosepfR-r"
# ## Applying k-Fold Cross Verification
#
# It creates 10 training sets and looks at their accuracy and then finds the mean value.
# + colab={"base_uri": "https://localhost:8080/"} id="-Mg2YY4JfSPk" outputId="b8bebc0e-de43-4b14-906a-0823db7e5d4a"
from sklearn.model_selection import cross_val_score
accuraries = cross_val_score(estimator = classifier, X = X_train, y = Y_train, cv = 10)
print("Accuracy: {:2f} % ".format(accuraries.mean()*100))
print("Standard Deviation: {:2f} % ".format(accuraries.std()*100))
# + [markdown] id="rxUze_oDhe3p"
# ## Visulization of Training Set Result
# + colab={"base_uri": "https://localhost:8080/", "height": 349} id="7ya0WnSERe_w" outputId="2e1169f1-59b0-494b-b5f5-fcf37147000d"
from matplotlib.colors import ListedColormap
X_set, y_set = feat_scale.inverse_transform(X_train), Y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 10, stop = X_set[:, 0].max() + 10, step = 0.25),
np.arange(start = X_set[:, 1].min() - 1000, stop = X_set[:, 1].max() + 1000, step = 0.25))
plt.contourf(X1, X2, classifier.predict(feat_scale.transform(np.array([X1.ravel(), X2.ravel()]).T)).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Support Vector Machine (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
# + [markdown] id="bl59PoruhfuJ"
# ## Visulization of Test Set Result
# + id="wZEsJuuWVb3V" colab={"base_uri": "https://localhost:8080/", "height": 349} outputId="93f71dfa-577c-4775-b3ea-dddde5decb46"
from matplotlib.colors import ListedColormap
X_set, y_set = feat_scale.inverse_transform(X_test), Y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 10, stop = X_set[:, 0].max() + 10, step = 0.25),
np.arange(start = X_set[:, 1].min() - 1000, stop = X_set[:, 1].max() + 1000, step = 0.25))
plt.contourf(X1, X2, classifier.predict(feat_scale.transform(np.array([X1.ravel(), X2.ravel()]).T)).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Support Vector Machine (Test set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
| Model Selection/k-Fold Cross Verification/k_Fold_Cross_Verification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Final Exercise - Putting it All Together
#
# In this last exercise, you'll write a full ETL pipeline for the GDP data. That means you'll extract the World Bank data, transform the data, and load the data all in one go. In other words, you'll want one Python script that can do the entire process.
#
# Why would you want to do this? Imagine working for a company that creates new data every day. As new data comes in, you'll want to write software that periodically and automatically extracts, transforms, and loads the data.
#
# To give you a sense for what this is like, you'll extract the GDP data one line at a time. You'll then transform that line of data and load the results into a SQLite database. The code in this exercise is somewhat tricky.
#
# Here is an explanation of how this Jupyter notebook is organized:
# 1. The first cell connects to a SQLite database called worldbank.db and creates a table to hold the gdp data. You do not need to do anything in this code cell other than executing the cell.
# 2. The second cell has a function called extract_line(). You don't need to do anything in this code cell either besides executing the cell. This function is a [Python generator](https://wiki.python.org/moin/Generators). You don't need to understand how this works in order to complete the exercise. Essentially, a generator is like a regular function except instead of a return statement, a generator has a yield statement. Generators allow you to use functions in a for loop. In essence, this function will allow you to read in a data file one line at a time, run a transformation on that row of data, and then move on to the next row in the file.
# 3. The third cell contains a function called transform_indicator_data(). This function receives a line from the csv file and transforms the data in preparation for a load step.
# 4. The fourth cell contains a function called load_indicator_data(), which loads the trasnformed data into the gdp table in the worldbank.db database.
# 5. The fifth cell runs the ETL pipeilne
# 6. The sixth cell runs a query against the database to make sure everything worked correctly.
#
# You'll need to modify the third and fourth cells.
# +
# run this cell to create a database and a table, called gdp, to hold the gdp data
# You do not need to change anything in this code cell
import sqlite3
# connect to the database
# the database file will be worldbank.db
# note that sqlite3 will create this database file if it does not exist already
conn = sqlite3.connect('worldbank.db')
# get a cursor
cur = conn.cursor()
# drop the test table in case it already exists
cur.execute("DROP TABLE IF EXISTS gdp")
# create the test table including project_id as a primary key
cur.execute("CREATE TABLE gdp (countryname TEXT, countrycode TEXT, year INTEGER, \
gdp REAL, PRIMARY KEY (countrycode, year));")
conn.commit()
conn.close()
# -
# Generator for reading in one line at a time
# generators are useful for data sets that are too large to fit in RAM
# You do not need to change anything in this code cell
def extract_lines(file):
while True:
line = file.readline()
if not line:
break
yield line
# +
# TODO: fill out the code wherever you find a TODO in this cell
# This function has two inputs:
# data, which is a row of data from the gdp csv file
# colnames, which is a list of column names from the csv file
# The output should be a list of [countryname, countrycode, year, gdp] values
# In other words, the output would look like:
# [[Aruba, ABW, 1994, 1.330168e+09], [Aruba, ABW, 1995, 1.320670e+09], ...]
#
import pandas as pd
import numpy as np
import sqlite3
# transform the indicator data
def transform_indicator_data(data, colnames):
# get rid of quote marks
for i, datum in enumerate(data):
data[i] = datum.replace('"','')
# TODO: the data variable contains a list of data in the form [countryname, countrycode, 1960, 1961, 1962,...]
# since this is the format of the data in the csv file. Extract the countryname from the list
# and put the result in the country variable
country = data[0]
# these are "countryname" values that are not actually countries
non_countries = ['World',
'High income',
'OECD members',
'Post-demographic dividend',
'IDA & IBRD total',
'Low & middle income',
'Middle income',
'IBRD only',
'East Asia & Pacific',
'Europe & Central Asia',
'North America',
'Upper middle income',
'Late-demographic dividend',
'European Union',
'East Asia & Pacific (excluding high income)',
'East Asia & Pacific (IDA & IBRD countries)',
'Euro area',
'Early-demographic dividend',
'Lower middle income',
'Latin America & Caribbean',
'Latin America & the Caribbean (IDA & IBRD countries)',
'Latin America & Caribbean (excluding high income)',
'Europe & Central Asia (IDA & IBRD countries)',
'Middle East & North Africa',
'Europe & Central Asia (excluding high income)',
'South Asia (IDA & IBRD)',
'South Asia',
'Arab World',
'IDA total',
'Sub-Saharan Africa',
'Sub-Saharan Africa (IDA & IBRD countries)',
'Sub-Saharan Africa (excluding high income)',
'Middle East & North Africa (excluding high income)',
'Middle East & North Africa (IDA & IBRD countries)',
'Central Europe and the Baltics',
'Pre-demographic dividend',
'IDA only',
'Least developed countries: UN classification',
'IDA blend',
'Fragile and conflict affected situations',
'Heavily indebted poor countries (HIPC)',
'Low income',
'Small states',
'Other small states',
'Not classified',
'Caribbean small states',
'Pacific island small states']
# filter out country name values that are in the above list
if country not in non_countries:
# In this section, you'll convert the single row of data into a data frame
# The advantage of converting a single row of data into a data frame is that you can
# re-use code from earlier in the lesson to clean the data
# TODO: convert the data variable into a numpy array
# Use the ndmin=2 option
data_array = np.array(data, ndmin=2)
# TODO: reshape the data_array so that it is one row and 63 columns
data_array.
# TODO: convert the data_array variable into a pandas dataframe
# Note that you can specify the column names as well using the colnames variable
# Also, replace all empty strings in the dataframe with nan (HINT: Use the replace module and np.nan)
df = None
# TODO: Drop the 'Indicator Name' and 'Indicator Code' columns
# TODO: Reshape the data sets so that they are in long format
# The id_vars should be Country Name and Country Code
# You can name the variable column year and the value column gdp
# HINT: Use the pandas melt() method
# HINT: This was already done in a previous exercise
df_melt = None
# TODO: Iterate through the rows in df_melt
# For each row, extract the country, countrycode, year, and gdp values into a list like this:
# [country, countrycode, year, gdp]
# If the gdp value is not null, append the row (in the form of a list) to the results variable
# Finally, return the results list after iterating through the df_melt data
# HINT: the iterrows() method would be useful
# HINT: to check if gdp is equal to nan, you might want to convert gdp to a string and compare to the
# string 'nan
results = []
return results
# +
# TODO: fill out the code wherever you find a TODO in this cell
# This function loads data into the gdp table of the worldbank.db database
# The input is a list of data outputted from the transformation step that looks like this:
# [[Aruba, ABW, 1994, 1.330168e+09], [Aruba, ABW, 1995, 1.320670e+09], ...]
# The function does not return anything. Instead, the function iterates through the input and inserts each
# value into the gdp data set.
def load_indicator_data(results):
# TODO: connect to the worldbank.db database using the sqlite3 library
conn = None
# TODO: create a cursor object
cur = None
if results:
# iterate through the results variable and insert each result into the gdp table
for result in results:
# TODO: extract the countryname, countrycode, year, and gdp from each iteration
countryname, countrycode, year, gdp = None
# TODO: prepare a query to insert a countryname, countrycode, year, gdp value
sql_string = None
# connect to database and execute query
try:
cur.execute(sql_string)
# print out any errors (like if the primary key constraint is violated)
except Exception as e:
print('error occurred:', e, result)
# commit changes and close the connection
conn.commit()
conn.close()
return None
# +
# Execute this code cell to run the ETL pipeline
# You do not need to change anything in this cell
# open the data file
with open('../data/gdp_data.csv') as f:
# execute the generator to read in the file line by line
for line in extract_lines(f):
# split the comma separated values
data = line.split(',')
# check the length of the line because the first four lines of the csv file are not data
if len(data) == 63:
# check if the line represents column names
if data[0] == '"Country Name"':
colnames = []
# get rid of quote marks in the results to make the data easier to work with
for i, datum in enumerate(data):
colnames.append(datum.replace('"',''))
else:
# transform and load the line of indicator data
results = transform_indicator_data(data, colnames)
load_indicator_data(results)
# +
# Execute this code cell to output the values in the gdp table
# You do not need to change anything in this cell
# connect to the database
# the database file will be worldbank.db
# note that sqlite3 will create this database file if it does not exist already
conn = sqlite3.connect('worldbank.db')
# get a cursor
cur = conn.cursor()
# create the test table including project_id as a primary key
df = pd.read_sql("SELECT * FROM gdp", con=conn)
conn.commit()
conn.close()
df
# -
# # Conclusion
#
# ETL Pipelines involve extracting data from one source, which in this case was a csv file, then transforming the data into a more usable form, and finally loading the data somewhere else.
#
# The purpose of ETL pipelines is to make data more usable and accessible.
| lessons/ETLPipelines/18_final_exercise/.ipynb_checkpoints/18_final_exercise-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import gzip
import pandas as pd
import numpy as np
np.random.seed(1000)
from matplotlib import pyplot as plt
import seaborn as sns
plt.tight_layout()
sns.set(style="whitegrid")
sns.set_palette((sns.color_palette('colorblind', 8)))
dims = (11.7, 8.27)
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from matplotlib.colors import ListedColormap
from sklearn.linear_model import LogisticRegression
from sklearn.utils import check_array
from sklearn.preprocessing import StandardScaler
import time
import datetime
import math
import random
# +
def load_mnist(path, kind='train'):
## Load MNIST function. Retrieved from https://github.com/zalandoresearch/fashion-mnist
labels_path = os.path.join(path, '%s-labels-idx1-ubyte.gz' % kind)
images_path = os.path.join(path, '%s-images-idx3-ubyte.gz' % kind)
with gzip.open(labels_path, 'rb') as lbpath:
labels = np.frombuffer(lbpath.read(), dtype=np.uint8, offset=8)
with gzip.open(images_path, 'rb') as imgpath:
images = np.frombuffer(imgpath.read(), dtype=np.uint8, offset=16).reshape(len(labels), 784)
return images, labels
def preprocess(train, test):
sc = StandardScaler()
sc.fit(train)
xform_train = sc.transform(train)
xform_test = sc.transform(test)
return xform_train, xform_test
def refresh_data():
X_train, Y_train = load_mnist('C:/git/IST718/Lab3/data/fashion', kind='train')
X_test, Y_test = load_mnist('C:/git/IST718/Lab3/data/fashion', kind='t10k')
## Transforming the data
X_train, X_test = preprocess(X_train.astype('float64'), X_test.astype('float64'))
## Reducing the data to a decimal value
X_train /= 255
X_test /= 255
return X_train, X_test, Y_train, Y_test
def int_to_desc(i):
## Numeric dict for each value in the dataset
conv = {0: 'T-shirt/top', 1: 'Trouser', 2: 'Pullover', 3: 'Dress', 4: 'Coat', 5: 'Sandal',
6: 'Shirt', 7: 'Sneaker', 8: 'Bag', 9: 'Ankle boot'}
## Try to get the value for key i, else assign unknown
try:
ret = conv[i]
except:
ret = 'Unknown'
return ret
def check_random(n, x, y, p):
## Takes in integer N, X data, Y data, and predicted Y data and returns a plot with the information displayed
rows = math.ceil(n/5)
fig, ax = plt.subplots(nrows=rows, ncols=5, sharex=True, sharey=True,)
ax = ax.flatten()
for i in range(n):
j = random.randint(0,len(p)-1)
img = x[j].reshape(28, 28)
if p[j] != y[j]:
cmap = 'Reds'
else:
cmap = 'Greens'
ax[i].imshow(img, cmap=cmap, interpolation='nearest')
predicted = int_to_desc(p[j])
actual = int_to_desc(y[j])
ax[i].set_title('P: {}\n A: {}'.format(predicted,actual))
ax[0].set_xticks([])
ax[0].set_yticks([])
plt.tight_layout()
plt.show()
# +
solvers = ['newton-cg', 'sag', 'saga', 'lbfgs']
result_columns = ['solver', 'time', 'acc']
result_df = pd.DataFrame(columns=result_columns)
# -
for solver in solvers:
print('Refreshing data...')
X_train, X_test, Y_train, Y_test = refresh_data()
print('Running {0} solver'.format(solver))
## Starting timer
starttime = time.time()
## Compiling and fitting model
model = LogisticRegression(solver=solver)
fit = model.fit(X_train, Y_train)
## Ending timer
endtime = time.time()
## Total time
totaltime = endtime - starttime
## Evaluating model on test data
score = model.score(X_test, Y_test)
rownum = len(result_df)
result_df.at[rownum, 'solver'] = solver
result_df.at[rownum, 'acc'] = score
result_df.at[rownum, 'time'] = totaltime
print(result_df)
# +
print('Refreshing data...')
X_train, X_test, Y_train, Y_test = refresh_data()
model = LogisticRegression(solver='lbfgs')
fit = model.fit(X_train, Y_train)
pred = model.predict(X_test)
# -
check_random(n=15, x=X_test, y=Y_test, p=pred)
| Lab3/logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Functions
# ## Excercise 1
#
# ### Write a function that takes two numbers as parameters and returns the sum of two
# +
## Excercise 1 - code cell
def func_add(num1, num2):
## your code here
# + editable=false
## Run this cell to validate your code. ##
## This is a test cell for the Excercise 1 Cell ##
## Please do not modify the contents of this cell ##
## These are the test cases your code should pass ##
assert func_add(4,5) == 9
assert func_add(4.5,4.5) == 9
print("All Passed!")
###################################################
# -
# ## Excercise 2
#
# ### Write a function that takes lengths of two sides of a right angle triangle as parameters and returns the length of the hypotnuse
#
# #### Hint: hypotnuse**2 = side_1**2 + side_2**2
# +
## Excercise 2 - code cell
def func_hypot(side1, side2):
## your code here
# + editable=false
## Run this cell to validate your code. ##
## This is a test cell for the Excercise 2 Cell ##
## Please do not modify the contents of this cell ##
## These are the test cases your code should pass ##
assert func_hypot(3,4) == 5
assert func_hypot(16,30) == 34
print("All Passed!")
###################################################
# -
# ## Excercise 3
#
# ### Write a function that takes two numbers as parameters and returns a string as per the following flow chart -
#
# 
# +
## Excercise 3 - code cell
def func_compare(x, y):
## your code here
# + editable=false
## Run this cell to validate your code. ##
## This is a test cell for the Excercise 3 Cell ##
## Please do not modify the contents of this cell ##
## These are the test cases your code should pass ##
assert func_compare(3,4) == 'less'
assert func_compare(16,16) == 'equal'
assert func_compare(4,3) == 'greater'
print("All Passed!")
###################################################
# -
# ## Excercise 4
#
# ### Write a function that takes one, two, or three numbers as parameters and returns the sum of all
#
# ### Example - func_add2(1) = 1, func_add2(1,2) = 3, func_add2(3,7,10) = 20
# +
## Excercise 4 - code cell
def func_add2
## your code here
# + editable=false
## Run this cell to validate your code. ##
## This is a test cell for the Excercise 4 Cell ##
## Please do not modify the contents of this cell ##
## These are the test cases your code should pass ##
assert func_add2(1) == 1
assert func_add2(1,2) == 3
assert func_add2(3,7,10) == 20
print("All Passed!")
###################################################
# -
| Basic Python/Worksheets/03_Functions_Worksheet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import display, HTML
import warnings
warnings.filterwarnings('ignore')
import pickle
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.svm import SVC
from sklearn.gaussian_process.kernels import RBF
from sklearn.naive_bayes import GaussianNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import cross_val_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import confusion_matrix # evaluation metric
from sklearn.metrics import accuracy_score # evaluation metric
from sklearn.metrics import f1_score # evaluation metric
from sklearn.metrics import precision_recall_fscore_support
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV #Performing grid search
from sklearn.ensemble import RandomForestRegressor
df=pd.read_csv("green_tripdata_2016-12.csv", header=0)
df.head()
df.info()
# As shown above pd.read_csv is not able to read the rows and columns correctly. The data is not corrcectly split and there are 2 extra null columns at the end that are added.
# So I have to read the csv twice, once for the rows and then for the columns and drop the columns with all null values to align the rows to the columns.
# +
rows = pd.read_csv('green_tripdata_2016-12.csv', skiprows=[0], header = None)
# skip header line
rows = rows.dropna(axis=1, how='all')
# drop columns that only have NaNs
columns = pd.read_csv('green_tripdata_2016-12.csv', nrows=0).columns
# -
rows
columns = columns.drop('ehail_fee')
rows.columns = columns
#df1 is the dataframe with the rows and columns aligned
df1 = rows
df1
df1.info()
df1.dtypes
df1.shape
# Field Name Description
# 1. VendorID: A code indicating the LPEP provider that provided the record.
# 1= Creative Mobile Technologies, LLC;
# 2= VeriFone Inc.
# 2. lpep_pickup_datetime: The date and time when the meter was engaged.
# 3. lpep_dropoff_datetime The date and time when the meter was disengaged.
# 4. Passenger_count: The number of passengers in the vehicle.
# This is a driver-entered value.
# 5. Trip_distance: The elapsed trip distance in miles reported by the taximeter.
# 6. PULocationID: TLC Taxi Zone in which the taximeter was engaged
# 7. DOLocationID: TLC Taxi Zone in which the taximeter was disengaged
# 8. RateCodeID: The final rate code in effect at the end of the trip.
# 1= Standard rate
# 2=JFK
# 3=Newark
# 4=Nassau or Westchester
# 5=Negotiated fare
# 6=Group ride
# 9. Store_and_fwd_flag: This flag indicates whether the trip record was held in vehicle memory before sending to the vendor, aka “store and forward,” because the vehicle did not have a connection to the server.
# Y= store and forward trip
# N= not a store and forward trip
# 10. Payment_type A numeric code signifying how the passenger paid for the trip.
# 1= Credit card
# 2= Cash
# 3= No charge
# 4= Dispute
# 5= Unknown
# 6= Voided trip
# 11. Fare_amount: The time-and-distance fare calculated by the meter.
# 12. Extra: Miscellaneous extras and surcharges. Currently, this only includes
# the $0.50 and $1 rush hour and overnight charges.
# 13. MTA_tax: $0.50 MTA tax that is automatically triggered based on the metered rate in use.
#
# 14. Improvement_surcharge: $0.30 improvement surcharge assessed on hailed trips at the flag drop. The improvement surcharge began being levied in 2015.
# 15. Tip_amount Tip amount – This field is automatically populated for credit card tips. Cash tips are not included.
# 16. Tolls_amount: Total amount of all tolls paid in trip.
# 17. Total_amount: The total amount charged to passengers. Does not include cash tips.
# 18. Trip_type: A code indicating whether the trip was a street-hail or a dispatch that is automatically assigned based on the metered rate in use but can be altered by the driver.
# 1= Street-hail
# 2= Dispatch
df1.describe()
# ## Data Cleaning
# - droped ehail_fee- all transactions are NaNs
# - Remove negative observations in variables- fare_amount, extra, mta_tax, tip_amount, improvement_surcharge, total_amount.
# - Removed observations with 0 trip distance values
# - There were 166 trips with passenger count as. Removed observations with passenger count as 0
# - Removed outlier values in trip distance as seen from the box plot which are 3 standard deviations away from mean
# - converted the pickup time and drop off time into datetime
#### Selecting records where total amount is greater than zero.
dfc = df1[df1['total_amount']>0]
dfc.shape
#Removed negative observations in variables
dfc[dfc['fare_amount']<0]
dfc[dfc['extra']<0]
dfc[dfc['mta_tax']<0]
dfc[dfc['tip_amount']<0]
# Plotting a histogram of trip distance.
plt.hist(dfc["trip_distance"])
plt.title('Histogram of the trip distance')
plt.savefig('histogram of trip distance with outliers.png')
plt.show()
plt.figure(figsize=[30,2])
sns.boxplot(dfc['trip_distance'])
dfc['trip_distance'].describe()
(dfc['trip_distance'][dfc['trip_distance']==0]).count()
dfc['trip_distance'][(dfc['trip_distance']>=0) & (dfc['trip_distance'] <=11)]
# There are 13958 rows out of 1224158 which has 0 trip distance. We need to remove the records which has 0 trip distance and remove outliers as seen in the box plot. We can calculate z score to remove outliers or remove values which are 3 std deviations away from the mean
# +
#keeping records where trip distance is greater than 0.
dfc = dfc[dfc['trip_distance']>0]
#Removing outliers from trip distance
dfc = dfc[~(np.abs(dfc.trip_distance-dfc.trip_distance.mean()) > (3*dfc.trip_distance.std()))]
#Keeping records with passenger count greater than 0.
dfc = dfc[dfc['passenger_count']>0]
# -
dfc.shape
# ## histogram of trip distance
# - plotted the histogram with 50 bins
# - Outliers have been removed before plotting.
# - Outliers are defined as any point located further than 3 standard deviations from the mean
#Plotting histogram with 50 bins.
plt.hist(dfc["trip_distance"],bins = 50)
plt.title('Histogram of trip distance without outliers')
plt.xlabel('Trip Distance (in miles)')
plt.ylabel('Count')
plt.savefig('histogram of trip distance without outliers.png')
plt.show()
# - The trip distance is skewed to the right that means the mean is greater than the median.
# - That means most of the data is on the left side of the histogram.
# - This tells us most of the trips are short distance trips between 0-3 miles of distance.
dfc.head()
# ## Find interesting trip statistics grouped by hour
#converting the pickup time and drop off time into datetime
dfc['lpep_pickup_datetime']=pd.to_datetime(dfc['lpep_pickup_datetime'])
dfc['lpep_dropoff_datetime']=pd.to_datetime(dfc['lpep_dropoff_datetime'])
#we have pickup datetime and dropoff datetime converted into datetime
dfc.dtypes
# ### Feature Engineering
# - Added pickupday, pickupday_no, pickup_hour into dataframe df1.
# - these features are extracted from lpep_pickup_datetime and lpep_dropoff_datetime
# - added time of day feature- morning, afternoon, evening, late night
# - added is_weekday- 1 for weekday, 0 for weekend
# +
dfc['pickupday']=dfc['lpep_pickup_datetime'].dt.day_name()
dfc['dropoffday']=dfc['lpep_dropoff_datetime'].dt.day_name()
dfc['pickup_day_no']=dfc['lpep_pickup_datetime'].dt.weekday
dfc['dropoff_day_no']=dfc['lpep_dropoff_datetime'].dt.weekday
dfc['pickup_hour']=dfc['lpep_pickup_datetime'].dt.hour
dfc['dropoff_hour']=dfc['lpep_dropoff_datetime'].dt.hour
# -
dfc.head()
def time_of_day(x):
if x in range(6,12):
return 'Morning'
elif x in range(12,16):
return 'Afternoon'
elif x in range(16,22):
return 'Evening'
else:
return 'Late night'
dfc['pickup_timeofday']=dfc['pickup_hour'].apply(time_of_day)
dfc['dropoff_timeofday']=dfc['dropoff_hour'].apply(time_of_day)
#returns 1 for weekday and 0 for weekend
def is_weekday(x):
if x in range(0,5):
return 1
else:
return 0
# Integer Value Day of the week
#
# 0 Monday
# 1 Tuesday
# 2 Wednesday
# 3 Thursday
# 4 Friday
# 5 Saturday
# 6 Sunday
dfc['is_weekday'] = dfc['pickup_day_no'].apply(is_weekday)
# count of number of vendor ID grouped by pickup hour and sorted in descending order.
groupby_pickup_hour = ((dfc.groupby(dfc['pickup_hour']))['VendorID'].count()).reset_index(name='count') \
.sort_values(['count'], ascending=False)
groupby_pickup_hour.values
groupby_pickuphour = pd.DataFrame(((dfc.groupby(dfc['pickup_hour']))['VendorID'].count()).reset_index(name='count'))
# +
fig, ax = plt.subplots(figsize=(6, 4.5))
groupby_pickuphour['count'].plot()
# set axis labels
ax.set_xlabel('Pickup hour')
ax.set_ylabel('no of trips')
# update legend
ax.legend(ax.get_lines(), df.columns, loc='best', ncol=2)
plt.savefig('Graph plotting pickup hour against no of trips.jpg', dpi=fig.dpi)
plt.tight_layout()
# -
# From the above graph we can see that no of trips are
# - maximum around 7 p.m. which could be because people usually go home from work at that hour and
# - minimum at 5 a.m when everyone sleeps :)
sns.lineplot(x='pickup_hour',y='trip_distance',data=dfc)
plt.savefig('Graph_plotting_pickup_hour_against_trip_distance.jpg')
# The above graph plots trip distance against pickup hour and from the graph we can see:
# - The trip distance is maximum around 5 a.m. that maybe because of the long distance travel early morning rides. People who live far from there office have to start early to reach work.
# - The trip distance is minimum around 6 p.m- 7 p.m.
# ## The taxi drivers want to know what kind of trip yields better tips. Can you build a model for them and explain the model?
#
# - we will predict whether or not there will be a tip and how much the tip will be
# - Will have to build a classification and a regressor model for this
# - For example, we could identify that for this particular trip, if the duration is significantly shorter, then we might want to flag to this to the driver. The driver might then speed up a little so as to earn that extra tip. The downsides of this of course, would be that there might be dangerous driving, and the passenger might end up not tipping because of dangerous driving.
dfc.corr()
# From the above correlation we can see that tip_amount is highly correlated with trip_distance, fare amount, total amount and payment type.
dfc.describe()
# ### Data Cleaning for tip classification and prediction
# - Remove negative observations in variables- fare_amount, extra, mta_tax, tip_amount, improvement_surcharge, total_amount ;
# - Only select credit card transactions because only credit card tips can be captured in the system;
# - Remove variable payment_type, because there are only credit card payments in the remaining data;
# - Convert variables into proper (categorical / factor format / date time) formats.
# ### Payment_type
dfc.payment_type.value_counts()
sns.countplot(x='payment_type',data=dfc)
# Payment type
# 1= Credit card
# 2= Cash
#
# For tips- we have to consider only credit card payment
#created a new df dfcc which has data for only credit card payments
dfcc = dfc[dfc['payment_type']==1]
dfcc.drop('payment_type', axis=1, inplace=True)
dfcc.shape
dfcc.corr()
# ### Feature Engineering
# - Created variable- tip percentage which is percentage of tip amount from total amount
# - Created our target variable- tip_given- 1/0 for classification
# - created a categorical variable for trip distance
# - 0-2 miles: short distance
# - 2-7 miles: medium distance
# - 7 and above: large distance
# ##### tip_percentage
dfcc['tip_perc'] = round((dfcc['tip_amount']/dfcc['total_amount'])*100, 2)
dfcc.head()
dfcc.shape
dfcc['tip_perc'][dfcc['tip_perc']>0].count()/ dfcc['tip_perc'].count()
# 85% of the total rides have paid tips.
#create tip_given variable for classification
dfcc['tip_given'] = (dfcc.tip_perc>0)*1
dfcc['tip_given'].value_counts()
dfcc['tip_perc'].describe()
#lets look at the passenger count
dfcc['passenger_count'].describe()
# Number of passenger ranges from a minimum of 1 to a maximum of 9.
#this could be a class imbalanced dataset
sns.countplot(x='tip_given',data=dfcc)
plt.savefig('tip_given.jpg')
#Plotting histogram with 50 bins.
plt.hist(dfcc["tip_perc"],bins = 50)
plt.title('Histogram of tip percentage')
plt.xlabel('Tip Percentage (%)')
plt.ylabel('Count')
plt.savefig('Histogram of tip percentage.jpg')
plt.show()
# The distribution of the tip percentage is shown above. The mean tip percentage was 14.08% with a standard deviation of 7.5%. The tip percentage does not follow a normal distribution and is centered around a few typical values, i.e., 0%, 16-18%, 20%, 23%.
sns.catplot(x="pickupday",y="tip_perc",kind="bar",data=dfcc,height=5,aspect=1)
plt.title('The Average Trip percentage per PickUp Day of the week')
sns.catplot(x="dropoffday",y="tip_perc",kind="bar",data=dfcc,height=5,aspect=1)
plt.title('The Average Trip percentage per Dropoff Day of the week')
# The above graph shows us that the tip percentage was slightly higher on Saturday and Sunday but there is not much difference otherwise.
sort_by_trip_distance = dfcc.sort_values('trip_distance', ascending=False)
sort_by_trip_distance.describe()
def trip_dist_cat(x):
if x<2.0:
return "short_distance"
elif 2.0< x <7.0:
return "medium_distance"
else:
return "long distance"
dfcc['trip_distance_cat'] = dfcc['trip_distance'].apply(trip_dist_cat)
dfcc.head()
sns.countplot(x='trip_distance_cat',data=dfcc)
plt.savefig('tip_distance_category.jpg')
sns.catplot(x="trip_distance_cat",y="tip_perc",kind="strip",data=dfcc,height=5,aspect=1)
plt.title('The Average Tip percentage per trip distance category')
plt.savefig('The Average Tip percentage per trip distance category2.jpg')
sns.catplot(x="trip_distance_cat",y="tip_perc",kind="box",data=dfcc,height=5,aspect=1)
plt.title('The Average Tip percentage per trip distance category')
plt.savefig('The Average Tip percentage per trip distance category.jpg')
# From the above 2 plot we can see that
# - tip percentage are higher for short distances as compared to medium and long distances.
# - Also riders taking shorter trips between 0 and 2 miles are more likely to tip.
# - the average trip is around 14% for all distances.
sns.catplot(x="RatecodeID",y="tip_perc",kind="strip",data=dfcc,height=5,aspect=1)
plt.title('The Average Tip percentage per rate code ID')
plt.savefig('The Average Tip percentage per rate code ID.jpg')
# The rate code id 1 and 5 yeilds more tips as compared to other.
sns.catplot(x="mta_tax",y="tip_perc",kind="violin",data=dfcc,height=5,aspect=1)
plt.title('The Average Tip percentage per mta tax')
plt.savefig('The Average Tip percentage per mta tax.jpg')
# The tip percentage is higher for mta tax of 0.5.
sns.catplot(x="pickup_timeofday",y="tip_perc",hue="is_weekday",kind="strip",data=dfcc,height=5,aspect=1)
plt.title('The Average Tip percentage per pickup time of day')
plt.savefig('The Average Tip percentage per pickup time of day.jpg')
# The tips are usually given more
# - if its a weekend.
# - During Latenight and Evening times
sns.catplot(x="passenger_count",y="tip_perc",kind="strip",data=dfcc,height=5,aspect=1)
plt.title('The Average Tip percentage per passenger count')
plt.savefig('The Average Tip percentage per passenger count.jpg')
# Passenger count
# - 1 gives the most and the maximum tips
# - 2-6 gives lesser tips as compared to 1.
# - 7 and above give minimal tips.
# Classification
#
# - One-hot encoded categorical variables
# - Split data into features and labels
# - Converted to arrays
# - Split data into training and testing sets
# #### Encoding
# Next step is to convert categorical data: user, application and device_type into numeric values to be able to use in our model. We will be using one hot encoding to achieve the same.
dfcc.columns
s = dfcc['PULocationID'].value_counts()
s.describe()
dft = dfcc[['VendorID','store_and_fwd_flag', 'RatecodeID', 'PULocationID', 'DOLocationID',
'passenger_count', 'trip_distance', 'fare_amount', 'extra', 'mta_tax',
'tolls_amount', 'improvement_surcharge',
'trip_type', 'pickup_day_no', 'dropoff_day_no', 'pickup_timeofday',
'dropoff_timeofday', 'is_weekday','tip_given']]
dfx = pd.get_dummies(dft, columns=['VendorID','store_and_fwd_flag', 'RatecodeID', 'PULocationID', 'DOLocationID',
'passenger_count', 'improvement_surcharge',
'trip_type', 'pickup_day_no', 'dropoff_day_no', 'pickup_timeofday',
'dropoff_timeofday','is_weekday'])
list(dfx.columns)
target = ['tip_given']
test = dfx
test1 = test.drop(target, axis = 1)
test.shape, test1.shape
X_train, X_test, y_train, y_test = train_test_split(
np.array(test1),
np.array(test[target]),
test_size=0.4,
random_state=0)
X_train.shape, y_train.shape, X_test.shape, y_test.shape
# %%time
clf1 = RandomForestClassifier(n_estimators=500,
max_depth=3,
max_features=0.7,
random_state=0)
score1 = clf1.fit(X_train, y_train).predict(X_test)
# save the model to disk
pickle.dump(clf1, open('RandomForestClassifier_tip.p', 'wb'))
conf_matrix1 = confusion_matrix(y_test, score1)
precision_recall_fscore = precision_recall_fscore_support(y_test, score1, average='micro')
print(accuracy_score(y_test, score1))
print(precision_recall_fscore)
print(conf_matrix1)
# Random forest classification is giving an accuracy of 85.3% with a f1 score of 85.3%.
# What are the most important features?
import operator
dict_feat_imp = dict(zip(list(test1.columns.values),clf1.feature_importances_))
sorted_features = sorted(dict_feat_imp.items(), key=operator.itemgetter(1), reverse=True)
sorted_features
# From the above we can see that tips are paid if following conditions are true:
# - mta_tax is paid.
# - Pickup and drop off location number 42 which when checked online is Manhattan- Central Harlem North.
# - Rate code id 1 and 5- which are standard rate and negotiated fare resp.
# - Fare amount is higher
# - Pickup location id 255- Brooklyn Williamsburg (North Side)
# - trip distance is lower.
# ### Classification using gradient boosting algorithm
# - Sample size for training and optimization was chosen as 100000. This is a small sample size compared to the available data but the optimization was stable and good enough with 5 folds cross-validation
# - For this classification we have identified the predictors from our visualization analysis and feature importance obtained from our random forest algorithm
# - The number of trees were optimized using gridsearchCV
# - optimized number of trees: 70
#DATA
test[['mta_tax','DOLocationID_42','PULocationID_42', 'RatecodeID_5', 'RatecodeID_1', 'fare_amount',
'PULocationID_255', 'trip_distance', 'DOLocationID_74', 'PULocationID_247', 'DOLocationID_41',
'DOLocationID_138', 'PULocationID_130', 'PULocationID_159', 'PULocationID_66','PULocationID_166',
'DOLocationID_159','PULocationID_69', 'PULocationID_115', 'is_weekday_0','tolls_amount',
'passenger_count_1', 'passenger_count_2','pickup_timeofday_Evening', 'dropoff_timeofday_Evening',
'pickup_timeofday_Late night', 'tip_given']]
# +
## OPTIMIZATION & TRAINING OF THE CLASSIFIER
from sklearn.ensemble import GradientBoostingClassifier
print ("Optimizing the classifier...")
train = test.copy() # make a copy of the training set
# since the dataset is too big for my system, select a small sample size to carry on training and 5 folds cross validation
train = train.loc[np.random.choice(train.index,size=100000,replace=False)]
target = 'tip_given' # set target variable - it will be used later in optimization
tic = datetime.now() # initiate the timing
# for predictors start with candidates identified during the EDA and feature importance of previous model
predictors = ['mta_tax','DOLocationID_42','PULocationID_42', 'RatecodeID_5', 'RatecodeID_1', 'fare_amount',
'PULocationID_255', 'trip_distance', 'DOLocationID_74', 'PULocationID_247', 'DOLocationID_41',
'DOLocationID_138', 'PULocationID_130', 'PULocationID_159', 'PULocationID_66','PULocationID_166',
'DOLocationID_159','PULocationID_69', 'PULocationID_115', 'is_weekday_0','tolls_amount',
'passenger_count_1', 'passenger_count_2','pickup_timeofday_Evening', 'dropoff_timeofday_Evening',
'pickup_timeofday_Late night']
# optimize n_estimator through grid search
param_test = {'n_estimators':range(30,151,20)} # define range over which number of trees is to be optimized
# initiate classification model
model_cls = GradientBoostingClassifier(
learning_rate=0.1, # use default
min_samples_split=2,# use default
max_depth=5,
max_features='auto',
subsample=0.8, # try <1 to decrease variance and increase bias
random_state = 10)
# get results of the search grid
gs_cls = optimize_num_trees(model_cls,param_test,'roc_auc',train,predictors,target)
print (gs_cls.best_estimator_, gs_cls.best_params_, gs_cls.best_score_, gs_rfr.n_splits_)
# cross validate the best model with optimized number of estimators
modelfit(gs_cls.best_estimator_,train,predictors,target,'roc_auc')
# save the best estimator on disk as pickle for a later use
with open('gradient_classifier.p','wb') as fid:
pickle.dump(gs_cls.best_estimator_,fid)
fid.close()
print ("Processing time:", datetime.now()-tic)
# -
# ### Regression
# - To find out the tip percentage paid by the trips.
# - We will run the regression model for data which has tip>0
# - The factors which help increase the percentage of tips
# +
#PREPARING THE DATA
dfr = dfcc[['VendorID','store_and_fwd_flag', 'RatecodeID', 'PULocationID', 'DOLocationID',
'passenger_count', 'trip_distance', 'fare_amount', 'extra', 'mta_tax',
'tolls_amount', 'improvement_surcharge',
'trip_type', 'pickup_day_no', 'dropoff_day_no', 'pickup_timeofday',
'dropoff_timeofday', 'is_weekday','tip_perc','tip_given']]
dfrx = pd.get_dummies(dfr, columns=['RatecodeID', 'PULocationID', 'DOLocationID',
'passenger_count', 'improvement_surcharge',
'trip_type', 'pickup_day_no', 'dropoff_day_no', 'pickup_timeofday',
'dropoff_timeofday','is_weekday'])
targetr = ['tip_perc']
testr = dfrx[dfrx['tip_given']>0]
testr.drop('tip_given', axis=1, inplace= True)
testr1 = testr.drop(targetr, axis = 1)
print(testr.shape, testr1.shape)
# -
testr[['mta_tax','DOLocationID_42','PULocationID_42', 'RatecodeID_5', 'RatecodeID_1', 'fare_amount',
'PULocationID_255', 'trip_distance', 'DOLocationID_74', 'PULocationID_247', 'DOLocationID_41',
'DOLocationID_138', 'PULocationID_130', 'PULocationID_159', 'PULocationID_66','PULocationID_166',
'DOLocationID_159','PULocationID_69', 'PULocationID_115', 'is_weekday_0','tolls_amount',
'passenger_count_1', 'passenger_count_2','pickup_timeofday_Evening', 'dropoff_timeofday_Evening',
'pickup_timeofday_Late night']]
# +
# %%time
from datetime import datetime
print("start")
train = testr.copy()
train = train.loc[np.random.choice(train.index,size=100000,replace=False)]
indices = testr.index[~testr.index.isin(train.index)]
test = testr.loc[np.random.choice(indices,size=100000,replace=False)]
train['ID'] = train.index
IDCol = 'ID'
target = 'tip_perc'
predictors = ['mta_tax','DOLocationID_42','PULocationID_42', 'RatecodeID_5', 'RatecodeID_1', 'fare_amount',
'PULocationID_255', 'trip_distance', 'DOLocationID_74', 'PULocationID_247', 'DOLocationID_41',
'DOLocationID_138', 'PULocationID_130', 'PULocationID_159', 'PULocationID_66','PULocationID_166',
'DOLocationID_159','PULocationID_69', 'PULocationID_115', 'is_weekday_0','tolls_amount',
'passenger_count_1', 'passenger_count_2','pickup_timeofday_Evening', 'dropoff_timeofday_Evening',
'pickup_timeofday_Late night']
def optimize_num_trees(alg,param_test,scoring_method,train,predictors,target):
"""
This functions is used to tune paremeters of a predictive algorithm
alg: sklearn model,
param_test: dict, parameters to be tuned
scoring_method: str, method to be used by the cross-validation to valuate the model
train: pandas.DataFrame, training data
predictors: list, labels to be used in the model training process. They should be in the column names of dtrain
target: str, target variable
"""
gsearch = GridSearchCV(estimator=alg, param_grid = param_test, scoring=scoring_method,n_jobs=2,iid=False,cv=5)
gsearch.fit(train[predictors],train[target])
return gsearch
from sklearn.ensemble import RandomForestRegressor
# optimize n_estimator through grid search
param_test = {'n_estimators':range(50,200,25)} # define range over which number of trees is to be optimized
# initiate regression model
rfr = RandomForestRegressor()
print("regressor built")
# -
# get results of the search grid
gs_rfr = optimize_num_trees(rfr,param_test,'neg_mean_squared_error' ,train,predictors,target)
print (gs_rfr.best_estimator_, gs_rfr.best_params_, gs_rfr.best_score_, gs_rfr.n_splits_)
# Regression modeling:
# - Sample size for training and optimization was chosen as 100000. This is a small sample size compared to the available data but the optimization was stable and good enough with 5 folds cross-validation
# - The number of trees were optimized
# - Mean square
# Results:
#
# - optimized number of trees: 175
# - no of splits for cross validation: 5
# +
# define a function that help to train models and perform cv
def modelfit(alg,dtrain,predictors,target,scoring_method,performCV=True,printFeatureImportance=True,cv_folds=5):
"""
This functions train the model given as 'alg' by performing cross-validation. It works on both regression and classification
alg: sklearn model
dtrain: pandas.DataFrame, training set
predictors: list, labels to be used in the model training process. They should be in the column names of dtrain
target: str, target variable
scoring_method: str, method to be used by the cross-validation to valuate the model
performCV: bool, perform Cv or not
printFeatureImportance: bool, plot histogram of features importance or not
cv_folds: int, degree of cross-validation
"""
# train the algorithm on data
alg.fit(dtrain[predictors],dtrain[target])
#predict on train set:
dtrain_predictions = alg.predict(dtrain[predictors])
if scoring_method == 'roc_auc':
dtrain_predprob = alg.predict_proba(dtrain[predictors])[:,1]
#perform cross-validation
if performCV:
cv_score = cross_val_score(alg,dtrain[predictors],dtrain[target],cv=cv_folds,scoring=scoring_method)
#print model report
print ("\nModel report:")
if scoring_method == 'roc_auc':
print ("Accuracy:",metrics.accuracy_score(dtrain[target].values,dtrain_predictions))
print ("AUC Score (Train):",metrics.roc_auc_score(dtrain[target], dtrain_predprob))
if (scoring_method == 'neg_mean_squared_error'):
print ("Accuracy:",metrics.mean_squared_error(dtrain[target].values,dtrain_predictions))
if performCV:
print ("CV Score - Mean : %.7g | Std : %.7g | Min : %.7g | Max : %.7g" % (np.mean(cv_score),np.std(cv_score),np.min(cv_score),np.max(cv_score)))
#print feature importance
if printFeatureImportance:
if dir(alg)[0] == '_Booster': #runs only if alg is xgboost
feat_imp = pd.Series(alg.booster().get_fscore()).sort_values(ascending=False)
else:
feat_imp = pd.Series(alg.feature_importances_,predictors).sort_values(ascending=False)
feat_imp.plot(kind='bar',title='Feature Importances')
plt.ylabel('Feature Importance Score')
plt.show()
# cross validate the best model with optimized number of estimators
modelfit(gs_rfr.best_estimator_,train,predictors,target,'neg_mean_squared_error' )
# save the best estimator on disk as pickle for a later use
with open('regression_tip.p','wb') as fid:
pickle.dump(gs_rfr.best_estimator_,fid)
fid.close()
# -
# ### Conclusion
# * The most important features for getting better trips is as follows:
# - trip distance- lower the trip distance, higher the tip percentage
# - fare amount- higher the fare amount more the tip.
# - is weekday- get better tips on weekend
# - The tips are better for passenger count of 1-2.
# - Pickup and drop off time of evening and late night
# - tolls amount and mta tax also have some role to play in determining the tips.
# - Some pickup and dropoff locations in Manhattan and brooklyn are said to yeild better tips.
# ## Visualize the data to help understand trip patterns
dfcc.head()
# ### VendorID
# - vendor_id — a code for the provider associated with the trip record
dfcc.columns
dfcc['VendorID'].value_counts()
sns.countplot('VendorID',data=dfcc)
plt.savefig('VendorID.jpg')
# There is a 4 times difference between the trips taken by vendor ID 1 and 2 for December 2016.
# #### Trip Distance per VendorID
dfcc['trip_distance'].describe()
#sns.barplot(y='trip_distance',x='VendorID',data=dfcc, estimator=np.mean)
sns.catplot(y='trip_distance',x='VendorID',data=dfcc, kind='strip')
plt.savefig('visualizations/distribution_of_vendors_with_trip_distance.jpg')
# The distribution of both vendors in terms of trip distance is same. Which means there is no particular vendor who is taking longer or shorter trips.
# #### Passenger count per VendorID
sns.barplot(y='passenger_count',x='VendorID',data=dfcc)
plt.savefig('visualizations/passenger_count_per_vendors.jpg')
# This shows that vendor 2 generally carries more passengers as compared to vendor 1.
# ### store_and_fwd_flag
dfc['store_and_fwd_flag'].value_counts()
# There were less than 0.01% trips that were stored before forwarding
1881/1178533
sns.catplot(y='trip_distance',x='store_and_fwd_flag',data=dfcc, kind='strip')
plt.savefig('visualizations/storeandfwd_tripdist.jpg')
# - Most of the time the trip is not stored.
# - When the trips are stored, it is done mostly for shorter distances.
# ### RatecodeID
# 1= Standard rate
# 2=JFK
# 3=Newark
# 4=Nassau or Westchester
# 5=Negotiated fare
# 6=Group ride
dfc['RatecodeID'].value_counts()
sns.countplot('RatecodeID',data=dfcc)
plt.savefig('visualizations/ratecodeID.jpg')
# - Most of the trips are standard code trips and
# - 0.012% of those have negotiated fare.
# ### Passenger count
sns.distplot(dfcc['passenger_count'],kde=False)
plt.title('Distribution of Passenger Count')
plt.savefig('visualizations/Distribution_of_passenger_count.jpg')
figure,ax=plt.subplots(nrows=2,ncols=1,figsize=(10,10))
sns.countplot(x='pickupday',data=dfcc,ax=ax[0])
ax[0].set_title('Number of Pickups per day')
sns.countplot(x='dropoffday',data=dfcc,ax=ax[1])
ax[1].set_title('Number of Dropoffs per day')
plt.savefig('visualizations/no_of_pickups_dropoffs_per_day.jpg')
# The above graphs shows the distribution of number of pickups and dropoffs on each day of the week. We can see from the graph that maximum pickups and dropoffs are on Saturay followed by Friday. This could also be because December being holday season, People must be travelling to relatives and nearby places to spend time and enjoy.
figure,ax=plt.subplots(nrows=1,ncols=2,figsize=(15,5))
sns.countplot(x='pickup_timeofday',data=dfcc,ax=ax[0])
ax[0].set_title('The distribution of number of pickups on each part of the day')
sns.countplot(x='dropoff_timeofday',data=dfcc,ax=ax[1])
ax[1].set_title('The distribution of number of dropoffs on each part of the day')
plt.savefig('visualizations/no_of_pickups_dropoffs_each_time_of_day.jpg')
# The above graphs shows the distribution of number of pickups and dropoffs on each time of the day. We can see from the graph that maximum pickups and dropoffs are in the evening and least are in the afternoon.
figure,ax=plt.subplots(nrows=1,ncols=2,figsize=(10,5))
dfcc.pickup_hour.hist(bins=24,ax=ax[0])
ax[0].set_title('Distribution of pickup hours')
dfcc.dropoff_hour.hist(bins=24,ax=ax[1])
ax[1].set_title('Distribution of dropoff hours')
plt.savefig('visualizations/distribution_pickups_dropoffs_hours.jpg')
# - The above graph tells the same story as the distribution by time of day.
# - These graphs tells us the maximum pickups and dropoffs are between 6-8 p.m. in the evening which can be due to office going people and
# - It also suggests that people prefer taking public transport in the morning hours and take a cab during the evening hours when going back home.
# - minimum are between 4-6 a.m. in the morning.
# ### total amount
ax = sns.stripplot(x="pickupday", y="total_amount", data=dfcc, jitter=0.05, linewidth=1)
plt.savefig('visualizations/total_amount_billed_on_each day.jpg')
# The above plot shows the total amount billed on each day. We can see the amount is slightly higher on Friday as compared to other days.
sns.catplot(x="pickupday",y="trip_distance",kind="bar",data=dfcc,height=5,aspect=1)
plt.title('The Average Trip Distance per PickUp Day of the week')
sns.catplot(x="dropoffday",y="trip_distance",kind="bar",data=dfcc,height=5,aspect=1)
plt.title('The Average Trip Distance per Dropoff Day of the week')
plt.savefig('visualizations/avg_trip_distance_per_pickup_dropoff_day_of_week.jpg')
# The above graph shows the distribution of trip distance with days of the week. We can see from the above graph that maximum distances are travelled on Sunday.
# #### Conclusion and Future Work
# - Above is a basic visualization analysis of the trip data. In interest of time I will stop at this point.
# - As a future work, I will like to analyse the data further and understand how trip distance, fare etc are related to each other.
| NYC_green_taxi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
improt os
datasets = [
'AlOH_group_composition',
'AlOH_group_content',
'Ferrous_iron_index',
'Opaque_index',
'Ferric_oxide_content',
'FeOH_group_content',
'Ferric_oxide_composition',
'Kaolin_group_index',
'MgOH_group_content',
'Ferrous_iron_content_in_MgOH'
'Quartz_Index',
'Gypsum_Index',
'Silica_Index'
]
with rasterio.open('datasets/AlOH_Group_Content.tif') as src:
data = src.read(1)
plt.imshow(data)
bins, counts = numpy.unique(data, return_counts=True)
bins
plt.plot(bins[1:], counts[1:])
data.shape
data[:10, :10]
| get_aster_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="_wYSEntdD5QH"
# #Cleaning the dataset
# + id="DbiHec-aEFqf" colab={"base_uri": "https://localhost:8080/"} outputId="7aa068fa-4c22-4689-d9bd-22f652687b80"
# !pip install -q wordcloud
import wordcloud
import nltk
nltk.download('stopwords')
nltk.download('wordnet')
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
# Dataset: https://www.kaggle.com/amananandrai/clickbait-dataset=clickbait_data.csv
#https://towardsdatascience.com/is-this-headline-clickbait-86d27dc9b389
#https://medium.com/@sid321axn/fake-news-detection-using-nlp-and-machine-learning-in-python-wisdom-ml-6f548b0691a
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
pd.set_option('display.max_columns',100)
pd.set_option('display.max_rows',100)
import numpy as np
import warnings
warnings.filterwarnings('ignore')
import re
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.probability import FreqDist
from nltk.stem import PorterStemmer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.metrics import confusion_matrix
from sklearn.naive_bayes import MultinomialNB
from sklearn.model_selection import train_test_split
from sklearn.linear_model import PassiveAggressiveClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
from sklearn import metrics
from wordcloud import WordCloud, STOPWORDS
import string
from string import punctuation
from matplotlib import style
# + id="0hZM4g-VEkHk" colab={"base_uri": "https://localhost:8080/"} outputId="a94b9da9-dc97-4cdd-c434-355f8a3d3855"
# Loading the data
df = pd.read_csv('/content/sample_data/clickbait_data.csv')
df.info()
# + id="i29iBLCHVzuT"
from nltk.corpus import stopwords
stop = set(stopwords.words('english'))
punctuation = list(string.punctuation)
stop.update(punctuation)
def remove_stopwords(text):
final_text = []
for i in text.split():
if i.strip().lower() not in stop:
final_text.append(i.strip())
return " ".join(final_text)
df['headline']=df['headline'].apply(remove_stopwords)
# + colab={"base_uri": "https://localhost:8080/", "height": 199} id="4eUxTmaaV1IX" outputId="9d5c8b22-82a9-4c40-f795-2707f4ff5d58"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 199} id="vA7_tZzgV6lL" outputId="29b858c6-3d7a-4960-b861-eedeaffbaf82"
# converting to lowercase
df['headline']=df['headline'].str.lower()
df.head()
# + id="PhP7LTrRV_NK"
spec_chars = ["-","!",'"',"#","%","&","'","(",")",
"*","+",",","-",".","/",":",";","<",
"=",">","?","@","[","\\","]","^","_",
"`","{","|","}","~","–"]
for char in spec_chars:
df['headline'] = df['headline'].str.replace(char, '')
# + colab={"base_uri": "https://localhost:8080/", "height": 199} id="Pu5tJDJpWBZF" outputId="b673b04f-8eae-4bcd-8b4b-06af9ca0429e"
df.head()
# + id="AeY388GgWeM3"
from sklearn.model_selection import train_test_split
X = df['headline']
y = df.clickbait
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size=0.20, random_state=0)
# + id="FbH2UwdhWhOi"
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf_vectorizer = TfidfVectorizer(stop_words='english', max_df=0.8, ngram_range=(1,2))
tfidf_train_2 = tfidf_vectorizer.fit_transform(X_train)
tfidf_test_2 = tfidf_vectorizer.transform(X_test)
# + id="QV6ZzuM-Wibd"
count_vectorizer = CountVectorizer(stop_words='english')
count_train = count_vectorizer.fit_transform(X_train)
count_test = count_vectorizer.transform(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 346} id="adXs6m978IX6" outputId="2fd5618f-091a-4e23-f30d-237adb44a7ba"
pass_tf = PassiveAggressiveClassifier()
pass_tf.fit(tfidf_train_2, y_train)
pred = pass_tf.predict(tfidf_test_2)
score = metrics.accuracy_score(y_test, pred)
print("accuracy: %0.3f" % score)
cm = metrics.confusion_matrix(y_test, pred, labels=[1,0])
plot_confusion_matrix(cm, classes=[1,0])
# + id="OrPcl1BZ7oCj"
from sklearn.metrics import classification_report
# + colab={"base_uri": "https://localhost:8080/"} id="Hfo-IFMuWy76" outputId="c312ebe3-6cbc-4880-feed-cc7eddfdfcc1"
pass_tf = PassiveAggressiveClassifier()
pass_tf.fit(tfidf_train_2, y_train)
pred = pass_tf.predict(tfidf_test_2)
score = metrics.accuracy_score(y_test, pred)
print("accuracy: %0.3f" % score)
cm = metrics.confusion_matrix(y_test, pred, labels=[1,0])
print(cm)
matrix = classification_report(y_test,pred,labels=[1,0])
print('Classification report: \n',matrix)
| colab_notebook/Passive_Agressive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xarray as xr
import numpy as np
import matplotlib.pyplot as plt
import cmocean
# %matplotlib inline
dxr = xr.open_dataset('https://salishsea.eos.ubc.ca/erddap/griddap/ubcSSgSurfaceTracerFields1hV18-06')
dxr
ssh = dxr.ssh[24*30:24*60,520,230]
ssh
# +
ssh.plot()
| notebooks/SalishSeaCast/plot_ssh_ts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # User Defined Types
#
# User Defined Types (UDT) allow you to define a custom Matrix element type. For example in the shortest path problem, it can be useful to calculate not just the path length, but the number of hops along the shortest path, and a tree of shortest paths from every node back to the source. This example is a high level translate of the `LAGraph_BF_full` function implementing the [Bellman-Ford algorithm](https://en.wikipedia.org/wiki/Bellman%E2%80%93Ford_algorithm)
# +
from pygraphblas import *
class BF(Type):
members = ['double w', 'uint64_t h', 'uint64_t pi']
one = (lib.INFINITY, lib.UINT64_MAX, lib.UINT64_MAX)
@binop(boolean=True)
def EQ(z, x, y):
if not x.w == y.w and x.h == y.h and x.pi == y.pi:
z = True
else:
z = False
@binop()
def PLUS(z, x, y):
if (x.w < y.w
or x.w == y.w and x.h < y.h
or x.w == y.w and x.h == y.h and x.pi < y.pi):
if (z.w != x.w and z.h != x.h and z.pi != x.pi):
z.w = x.w
z.h = x.h
z.pi = x.pi
else:
z.w = y.w
z.h = y.h
z.pi = y.pi
@binop()
def TIMES(z, x, y):
z.w = x.w + y.w
z.h = x.h + y.h
if x.pi != lib.UINT64_MAX and y.pi != 0:
z.pi = y.pi
else:
z.pi = x.pi
# +
BF_monoid = BF.new_monoid(BF.PLUS, BF.one)
BF_semiring = BF.new_semiring(BF_monoid, BF.TIMES)
def shortest_path(matrix, start):
n = matrix.nrows
v = Vector.sparse(matrix.type, n)
for i, j, k in matrix:
if i == j:
matrix[i,j] = (0, 0, 0)
else:
matrix[i,j] = (k[0], 1, i)
v[start] = (0, 0, 0)
with BF_semiring, Accum(BF.PLUS):
for _ in range(matrix.nrows):
w = v.dup()
v @= matrix
if w.iseq(v):
break
return v
A = Matrix.sparse(BF, 6, 6)
A[0,1] = (9.0, 0, 0)
A[0,3] = (3.0, 0, 0)
A[1,2] = (8.0, 0, 0)
A[3,4] = (6.0, 0, 0)
A[3,5] = (1.0, 0, 0)
A[4,2] = (4.0, 0, 0)
A[1,5] = (7.0, 0, 0)
A[5,4] = (2.0, 0, 0)
# -
from pygraphblas.demo.gviz import draw
draw(A)
# ## Computing the shortest path
#
# Below is the result of labeling each node with the result of the shortest path computation. Each result is a tuple, the first value is the accumulated weight from the starting node. The second value is the number of hops from the starting node. The third value is the node "back" to the starting node, forming a shortest path tree with the starting node as the root.
draw(A, label_vector=shortest_path(A, 0))
| pygraphblas/demo/User-Defined-Types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.1
# language: julia
# name: julia-1.3
# ---
# # Compensated summation
# ## Accuracy and Stability of Numerical Algorithms
# N.Higham (pages 86-87)
#
# A good illustration of the benefits of compensated summation is provided by Euler's method for the ordinary differential equation initial value problem $\dot{y}=f(x,y), \ y(a)$ given , which generates an approximate solution according to $y_{k+1}=y_k+hf_k, \ y_0=y(a)$, We solved the equation $\dot{y}=-y$ with $y(0)=1$ over $[0,1]$ using $n$ steps of Euler's method ($nh=1$), with $n$ ranging from $10$ to $10^8$.
#
# Figure 4.2 shows the errors $e_n = |y(1) - \hat{y}_n|$ , where $\hat{y}_n$ is the computed approximation to $y(l)$. The computations were done in Fortran 90 in single precision arithmetic on a Sun SPARCstation ($ u \approx 6 x 10^{-8}$). Since Euler's method has global error of order $h$, the error curve on the plot should be approximately a straight line. For the standard implementation of Euler's method the errors $e_n$ start to increase steadily beyond $n = 20000$ because of the influence of rounding errors.
# <img src="Figure42.png" alt="" width="600"/>
# ## Experiment
using Pkg
using OrdinaryDiffEq,DiffEqDevTools
using IRKGaussLegendre,LinearAlgebra,Plots
using DoubleFloats
f=(du,u,p,t)->begin
du[1]=-u[1]
end
T= Float32(1.0)
tspan=(Float32(0.0),T)
u0=[Float32(1.0)];
prob=ODEProblem(f,u0,tspan);
n=8
out1=[]
out2=[]
out3=[]
out4=[]
for k in 1:n
h=Float32(1/10^k)
if (k<8) # k=8 fail
sol1=solve(prob,Euler(),dt=h,save_everystep=false)
u1=sol1.u[end]
push!(out1,u1[1])
end
sol2=solve(prob,IRKGL16(),adaptive=false, dt=Float32(h^(1/16)), save_everystep=false)
sol3=solve(prob,Vern9(),adaptive=false, dt=Float32(h^(1/9)), save_everystep=false)
sol4=solve(prob,RK4(),adaptive=false, dt=Float32((h)^(1/4)), save_everystep=false)
u2=sol2.u[end]
push!(out2,u2[1])
u3=sol3.u[end]
push!(out3,u3[1])
u4=sol4.u[end]
push!(out4,u4[1])
end
err1=norm.(out1.-exp(-1))
err2=norm.(out2.-exp(-1))
err3=norm.(out3.-exp(-1))
err4=norm.(out4.-exp(-1));
# +
hh=[10, 10^(-2),10^(-3),10^(-4), 10^(-5), 10^(-6) ]
ylimit1=-10
ylimit2=0
pl1=plot(log10.(abs.(err1)),ylims=(ylimit1,ylimit2), seriestype=:scatter, title="Euler", label="")
pl2=plot(log10.(abs.(err2)),ylims=(ylimit1,ylimit2), seriestype=:scatter, title="IRKGL16", label="")
pl3=plot(log10.(abs.(err3)),ylims=(ylimit1,ylimit2), seriestype=:scatter, title="Vern9", label="")
pl4=plot(log10.(abs.(err4)),ylims=(ylimit1,ylimit2), seriestype=:scatter, title="RK4", label="")
plot(pl1,pl4,pl3,pl2, layout=(2,2))
# -
| ODEProblems/.ipynb_checkpoints/Compesated Summation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [LEGALST-190] Lab 4/17: Feature Selection
#
# This lab will cover feature selection in order to train a machine learning model using `scikit-learn`. With complex datasets with 50+ features, we would run into the problem of overfitting your model and a long run time if all features were to be used. Feature selection is used in machine learning to avoid those type of issues.
#
# Estimated time: 35 minutes
#
# ### Table of Contents
# [The Data](#section data)<br>
# 1 - [Second Model](# section 1)<br>
# 2 - [Intro to Feature Removal Intuition](#section 2)<br>
# 3 - [Checking Results](#section 3)<br>
#
# +
# load all libraries
import numpy as np
from datascience import *
import datetime as dt
import pandas as pd
import seaborn as sns
#matplotlin
# %matplotlib inline
import matplotlib.pyplot as plt
#scikit-learn
from sklearn.feature_selection import RFE
from sklearn.feature_selection import VarianceThreshold
from sklearn.model_selection import train_test_split
from sklearn.linear_model import Ridge, Lasso, LinearRegression
from sklearn.model_selection import KFold
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.feature_selection import SelectFromModel
from sklearn import preprocessing
from sklearn import metrics
# -
# ## The Data: Bike Sharing<a id='section data'></a>
#
# By now, I'm sure you have been exposed to bike sharing dataset several times in this lab. This lab's data describes one such bike sharing system in Washington D.C., from UC Irvine's Machine Learning Repository.
#
# Information about the dataset: http://archive.ics.uci.edu/ml/datasets/Bike+Sharing+Dataset
# +
bike = pd.read_csv('data/day.csv', index_col=0)
# reformat the date column to integers that represent the day of the year, 001-366
bike['dteday'] = pd.to_datetime(bike['dteday'].unique()).strftime('%j')
# drop casual and registered riders because we want to predict the number of total riders
bike = bike.drop(['casual', 'registered'], axis = 1)
bike.head()
# -
# If you need to become familiar with this data set again, feel free to refer back to lab 2-22.
# To see how feature selection can change the accuracy for the better or worse, let's try to make a classifer that uses all features.
# +
# the features used to predict riders
X = bike.drop(['cnt'], axis = 1)
# the number of riders (target)
y = bike['cnt']
# set the random seed
np.random.seed(10)
# split the data with a 0.80 and 0.20 proportion respectively for train size and test size
# train_test_split returns 4 values: X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = train_test_split(X, y,
train_size=0.80, test_size=0.20)
# create a linear regression model
first_model_reg = LinearRegression()
#fit your model
first_model = first_model_reg.fit(X_train, y_train)
#predict X_train using your model
first_pred = first_model.predict(X_train)
#predict X_test using your model
test_pred = first_model.predict(X_test)
# -
# In order to check the error between the predicted values and the actual values, I have defined the root mean square error for you. Recall that the equation is the mean squared error of a predicted set of values.
def rmse(pred, actual):
return np.sqrt(np.mean((pred - actual) ** 2))
# +
# check the rmse of your models
first_train_error = rmse(first_pred, y_train)
first_test_error = rmse(test_pred, y_test)
print("Training RMSE:", first_train_error)
print("Test RMSE:", first_test_error)
# -
# ## Section 1: Second Model
#
# Our training and test errors seem to be pretty high. Let's see how we can improve our model by using feature selection. This process is often accompanied by lots of Exploratory Data Analysis (EDA). First we will look at which features correlate to our target feature (`cnt`).
#
# **Question 1.1:** Plot a few EDA yourself to become familar with the correlation values between certain features with the number of riders.
#
# **hint:** I recommend looking into heat maps
# +
#using the seaborn heatmaps library
#so it looks like this cannot take the dataframe 'bike' as its argument
#the following code comes from the Seaborn examples page
#https://seaborn.pydata.org/examples/many_pairwise_correlations.html
# Compute the correlation matrix
corr = bike.corr()
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.7, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# -
# **Question 1.2:** Looking at your EDA, how will that help you select which features to use?
# *Answer:* sample answer
# - Looking at the correlations between all features and cnt, I would pick the ones with correlation values higher than 0.50
#
# <b>*My answer*
# I would choose the really highly correlated features and select only one of them (for example, you probably don't need month and season, and you don't need both weekday and workingday). Looking at the dark colors on the heatmap in the cnt row is a good way to start, since those bivariate relationships are the strongest. You could then take the predictors that are most correlated and select one of them (like choose temp or atemp, not both).</b>
# **Question 1.3:** List out features that would be important to select for your model. Make sure to not include registered or casual riders in your features list.
features = ['yr', 'season', 'weathersit', 'atemp', 'windspeed'] #why are these strings?
features
# **Question 1.4:** Now create a `linear regression` model with the features that you have selected to predict the number of riders(`cnt`).
#
# First, separate your data into two parts, a dataframe containing the features used to make our prediction (X) and an array of the true values (y). To start, let's predict the total number of riders (y) using every feature that isn't a rider count (X). Then split the train_size and test_size containing 80% and 20% of the data respectively. Scikit-learn's `test_train_split function` will help here.
#
# You can refer back to lab 2-22 if needed.
#
# *Note that Lasso and Ridge models would use the same steps below.*
# +
# the features used to predict riders
X = bike[features]
# the number of riders
y = bike['cnt']
# set the random seed
np.random.seed(10)
# split the data
# train_test_split returns 4 values: X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = train_test_split(X, y,
train_size=0.80, test_size=0.20)
# -
# create a linear regression model
lin_reg = LinearRegression()
# fit the model
lin_model = lin_reg.fit(X_train, y_train)
# +
# predict the number of riders
lin_pred = lin_reg.predict(X_train)
# plot the residuals on a scatter plot
plt.scatter(y_train, lin_pred)
plt.title('Linear Model (OLS)')
plt.xlabel('actual value')
plt.ylabel('predicted value')
plt.show()
# -
# **Question 1.5:** What is the rmse for both the prediction of X_train and X_test?
# +
#predict your X_test here
lin_test_pred = lin_reg.predict(X_test)
second_train_error = rmse(lin_pred, y_train)
second_test_error = rmse(lin_test_pred, y_test)
print("Training RMSE:", second_train_error)
print("Test RMSE:", second_test_error)
# -
# hmm... maybe our selected features did not improve the error as much. Let's see how we can improve our model.
# ## Section 2: Introduction to Feature Removal Intuition<a id = 'section 2'></a>
#
# As a good rule of thumb, we typically wish to pick features that have roughly more than a 0.50 correlation with the target column. Also, even though not relevant to the bike sharing dataset, it is often best to remove columns that contain a high ratio of null values. However, sometimes null values represent 0 instead of data actually missing! So always be on the look out when you have to clean data.
#
# Of course, with any tedious and error prone process there is always a short cut that reduces time and human error. In part 1, you used your own intuition to pick out features that correlate the highest with the target feature. However, we can use `scikit-learn` to help pick the important features for us.
#
# Feature selection methods can give you useful information on the relative importance or relevance of features for a given problem. You can use this information to create filtered versions of your dataset and increase the accuracy of your model.
# ### Remove Features with Low Variance
#
# In removing features with low variance, all features whose variance does not meet some threshold are removed. In order to remove features that have low variance, you must use normalization on the columns before using VarianceThreshold. This is necessary to bring all the features to same scale. Ensuring standardised feature values implicitly weights all features equally. Otherwise, the variance estimates can be misleading between higher value features and lower value features. By default, normalization is not included in the function.
# +
# here we will reload the X and y values for you
X = bike.drop(['cnt'], axis = 1)
# the number of riders
y = bike['cnt']
# set the random seed
np.random.seed(10)
# normalize your data
X = preprocessing.normalize(X, norm = 'max', axis = 0)
# split the data
# train_test_split returns 4 values: X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = train_test_split(X, y,
train_size=0.80, test_size=0.20)
# -
# **Question 2.1:** What is the current shape of X_train?
# code answer here
X_train.shape
# Use `VarianceThreshold` to filter out features that match a 0.1 threshold. If your code produces an error ('No feature in X meets the variance threshold 0.10000') and you believe it is correct, try rerunning the `train_test_split` code block above and rerun the code below.
#
# Then you can use `transform` on the X_train. This will select features that match the threshold.
# +
#use VarianceThreshold
sel = VarianceThreshold(threshold=0.1)
sel.fit_transform(X_train)
# Subset features with transform
X_new = sel.transform(X_train)
#notice how many features are then selected compared to X_train's original features
print(X_new.shape)
#make sure to also transform X_test so it will match dimensions of X_train
X_new_test = sel.transform(X_test)
# -
# **Question 2.1:** How does the number of features from X_train compare to X_new?
# *Answer:* <b>There are far fewer features--we went from 12 to 3. This makes perfect sense since so many were closely correlated (like month and season).
# +
# New Linear Regression model for your X_new. Recall that X_new is the X_train with selected features.
new_lin_reg = LinearRegression()
# fit the model
new_lin_model = new_lin_reg.fit(X_new, y_train)
#predict X_new
new_lin_pred = new_lin_model.predict(X_new)
#predict X_new_test
new_test_pred = new_lin_model.predict(X_new_test)
# +
third_train_error = rmse(new_lin_pred, y_train)
third_test_error = rmse(new_test_pred, y_test)
print("Training RMSE:", third_train_error)
print("Test RMSE:", third_test_error)
# -
# **Question 2.2:** How does your root mean square error change compared to your model in section 1?
# *Answer:* <b>The root mean square error is much larger, probably because we were overfitting to the data in section 1.</b>
# ### Recursive Feature Elimiation with scikit-learn
#
# According to [Feature Selection in Python with Scikit-Learn](https://machinelearningmastery.com/feature-selection-in-python-with-scikit-learn/), recursive feature elimination works by “recursively removing attributes and building a model on those attributes that remain. It uses the model accuracy to identify which attributes (and combination of attributes) contribute the most to predicting the target attribute.”
#
# +
# here we will reload the X and y values
X = bike.drop(['cnt'], axis = 1)
# the number of riders
y = bike['cnt']
# set the random seed
np.random.seed(10)
# split the data
# train_test_split returns 4 values: X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = train_test_split(X, y,
train_size=0.80, test_size=0.20)
# +
# create a base classifier used to evaluate a subset of attributes
model = LinearRegression()
# create the RFE model and select 10 attributes
rfe = RFE(model, 10)
rfe.fit(X_train, y_train)
# -
# To check which features have been selected, we can use rfe.support_ to show mask of selected features.
print(rfe.support_)
# The feature ranking, such that ranking_[i] corresponds to the ranking position of the i-th feature. Selected (i.e., estimated best) features are assigned rank 1.
print(rfe.ranking_)
# +
# using rfe, predict your training set
new_pred = rfe.predict(X_train)
# now predict your test set
new_test_pred = rfe.predict(X_test)
# +
# time for errors
fourth_train_error = rmse(new_pred, y_train)
fourth_test_error = rmse(new_test_pred, y_test)
print("Training RMSE:", fourth_train_error)
print("Test RMSE:", fourth_test_error)
# -
# **Question 2.3:** How does recursive feature elimination change your error?
# *Answer:* <b>It reduced the root mean square error in both the training and test set below what it was for the regression in Part 1. The change wasn't huge, but I guess for OLS regression we would not expect it to be.
# ### Feature Importance
#
# Feature importance is selecting features that are most important from a previous classifier. For example, selecting the most important features from a number of randomized decision trees. "A decision tree can be used to visually and explicitly represent decisions and decision making. As the name goes, it uses a tree-like model of decisions." If you would like to read more, feel free to [click here](https://towardsdatascience.com/decision-trees-in-machine-learning-641b9c4e8052). The main idea behind using the randomized trees is to use many of them to perform prediction. This helps the model to be more robust.
#
# Methods that use ensembles of decision trees (like `Random Forest` or `Extra Trees`) can also compute the relative importance of each attribute. These importance values can be used to inform a feature selection process. In this lab, we will be using `Extra Trees`, Random forest will be introduced in the next lab.
#
# Below shows the construction of an Extra Trees ensemble of the bike share dataset and the display of the relative feature importance.
#
# Once you use `ExtraTreesClassifer` to create a new model, fit the model. Afterwards, you can use `SelectFromModel` to select features using the classifier. Make sure to `transform` your X_train to obtain the selected important features.
# +
# Fits a number of randomized decision trees. Use 15 estimators (this value was arbitrarily chosen)
# this allows us to select features
model = ExtraTreesClassifier(n_estimators=15)
#fit your model
model.fit(X_train, y_train)
# Select the important features of previous model
sel = SelectFromModel(model, prefit=True)
# Subset features
select_X_train = sel.transform(X_train)
# We want to create a train model *hint this model is exactly the same as model ^^
sel_model = ExtraTreesClassifier(n_estimators=15)
#fit your sel_model with the new X_train
sel_model.fit(select_X_train, y_train)
# +
#predict X_train using the new model
y_train_pred = sel_model.predict(select_X_train)
# we must also select features from X_test to have number of features match up with the model
select_X_test = sel.transform(X_test)
#predict y using select_X_test
y_test_pred = sel_model.predict(select_X_test)
fifth_train_error = rmse(y_train_pred, y_train)
fifth_test_error = rmse(y_test_pred, y_test)
print("Training RMSE:", fifth_train_error)
print("Test RMSE:", fifth_test_error)
# -
# **Question 2.4:** Which method gives the best results? Check errors rates between all methods mentioned in this section.
# *Answer:* <b>Whoa! There is zero root mean square error for the training set but much greater RMSE on the test set. Extreme case of overfitting?</b>
# ## Section 3: Checking Results<a id = 'section 3'></a>
#
# Note that since Linear Regression is not the only model option, you can use the above methods to fit a new model using either `Lasso` or `Ridge`.
# **Question 3.1:** Within the scope of this class, what are other methods that can be used to improve estimation?
# *Answer:* possible answers
# - testing multiple models
# - averging models
# - stacking models
# - change number of attributes in Recursive Feature Elimiation method
# - increase number of estimates in Feature Importance method
# **Question 3.1:** Now that we have gone through different methods of feature selection, let's see how the error changes with each method. I have created the dataframe for you, now graph it!
# +
labels = ['all_features', 'own_selection', 'variance_theshold', 'rfe', 'important']
methods = pd.DataFrame(columns = labels)
methods['all_features'] = [first_train_error, first_test_error]
methods['own_selection'] = [second_train_error, second_test_error]
methods['variance_theshold'] = [third_train_error, third_test_error]
methods['rfe'] = [fourth_train_error, fourth_test_error]
methods['important'] = [fifth_train_error, fifth_test_error]
methods = methods.rename(index={0: 'train'})
methods = methods.rename(index={1: 'test'})
methods
# +
#sample plot
methods.plot.bar()
# -
# ## Bibliography
#
# - <NAME>, An Introduction to Feature Selection. https://machinelearningmastery.com/an-introduction-to-feature-selection/
# - <NAME>, Feature Selection in Python with Scikit-Learn. https://machinelearningmastery.com/feature-selection-in-python-with-scikit-learn/
# - <NAME>, Why, How and When to apply Feature Selection. https://towardsdatascience.com/why-how-and-when-to-apply-feature-selection-e9c69adfabf2
# - Use of `Bike Share` data set adapted from UC Irvine's Machine Learning Repository. http://archive.ics.uci.edu/ml/datasets/Bike+Sharing+Dataset
# - Some code adapted from Celia Siu: https://csiu.github.io/blog/update/2017/03/06/day10.html
#
# ----
# Notebook developed by: <NAME>
#
# Data Science Modules: http://data.berkeley.edu/education/modules
| labs/4-17/4-17_Feature_Selection_Jon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Demo 5: Algorithms01
# + active=""
# This demo will demonstrate the options for plotting projections and images on TIGRE. The functions have been in previous demos, but in here an exaustive explanation and usage of them is given.
# NOTE: if you havent already downloaded the tigre_demo_file and navigated to the correct directory, do so before continuing with this demo.
# --------------------------------------------------------------------------
# This file is part of the TIGRE Toolbox
#
# Copyright (c) 2015, University of Bath and
# CERN-European Organization for Nuclear Research
# All rights reserved.
#
# License: Open Source under BSD.
# See the full license at
# https://github.com/CERN/TIGRE/license.txt
#
# Contact: <EMAIL>
# Codes: https://github.com/CERN/TIGRE/
# --------------------------------------------------------------------------
# Coded by: MATLAB (original code): <NAME>
# PYTHON : Reuben Lindroos
# -
# ## Define Geometry
import tigre
geo=tigre.geometry_default(high_quality=False)
# ## Load data and generate projections
import numpy as np
from tigre.Ax import Ax
from Test_data import data_loader
# define angles
angles=np.linspace(0,2*np.pi,dtype=np.float32)
# load head phantom data
head=data_loader.load_head_phantom(number_of_voxels=geo.nVoxel)
# generate projections
projections=Ax(head,geo,angles,'interpolated')
# ## Usage of FDK
# + active=""
# the FDK algorithm has been taken and modified from
# 3D Cone beam CT (CBCT) projection backprojection FDK, iterative reconstruction Matlab examples
# https://www.mathworks.com/matlabcentral/fileexchange/35548-3d-cone-beam-ct--cbct--projection-backprojection-fdk--iterative-reconstruction-matlab-examples
#
# The algorithm takes, as eny of them, 3 mandatory inputs:
# PROJECTIONS: Projection data
# GEOMETRY : Geometry describing the system
# ANGLES : Propjection angles
# And has a single optional argument:
# FILTER: filter type applied to the projections. Possible options are
# 'ram-lal' (default)
# 'shepp-logan'
# 'cosine'
# 'hamming'
# 'hann'
# The choice of filter will modify the noise and sopme discreatization
# errors, depending on which is chosen.
# -
import tigre.algorithms as algs
imgfdk1=algs.FDK(projections,geo,angles,filter='ram_lak')
imgfdk2=algs.FDK(projections,geo,angles,filter='hann')
# The look quite similar:
from tigre.utilities.plotImg import plotImg
plotImg(np.hstack((imgfdk1,imgfdk2)),slice=32,dim='x')
# + active=""
# On the other hand it can be seen that one has bigger errors in the whole
# image while the other just in the boundaries
# -
dif1=abs(head-imgfdk1)
dif2=abs(head-imgfdk2)
plotImg(np.hstack((dif1,dif2)),slice=32,dim='x')
| Python/tigre/demos/d05_Algorihtms01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem Statement
# Customer Churn is a burning problem for Telecom companies. Almost every telecom company pays a premium to get a customer on-board. Customer churn is a directly impacts company’s revenue.
#
# In this case-study, we simulate one such case of customer churn where we work on a data of post-paid customers with a contract. The data has information about customer usage behaviour, contract details, and payment details. The data also indicates which were the customers who cancelled their service.
#
# Based on this past data, Perform an EDA and build a model which can predict whether a customer will cancel their service in the future or not.
# # Data Dictionary
# * <b>Churn</b> - 1 if customer cancelled service, 0 if not
# * <b>AccountWeeks</b> - number of weeks customer has had active account
# * <b>ContractRenewal</b> - 1 if customer recently renewed contract, 0 if not
# * <b>DataPlan</b> - 1 if customer has data plan, 0 if not
# * <b>DataUsage</b> - gigabytes of monthly data usage
# * <b>CustServCalls</b> - number of calls into customer service
# * <b>DayMins</b> - average daytime minutes per month
# * <b>DayCalls</b> - average number of daytime calls
# * <b>MonthlyCharge</b> - average monthly bill
# * <b>OverageFee</b> - largest overage fee in last 12 months
# * <b>RoamMins</b> - average number of roaming minutes
#
#Import all necessary modules
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn import metrics,model_selection
from sklearn.preprocessing import scale
cell_df = pd.read_excel("Cellphone.xlsx")
# ## EDA
cell_df.head()
cell_df.info()
# There are missing values in some coumns.
# All variables are of numeric type and does not contain any data inconsistencies (causing numeric variables to be object due to some special characters present in the data).
# Churn is the target variable.
# Churn, ContractRenewal and DataPlan are binary variables.
cell_df[['AccountWeeks','DataUsage','CustServCalls','DayMins','DayCalls','MonthlyCharge','OverageFee','RoamMins']].describe()
# ### Check for Missing values
cell_df.isnull().sum()
# ### Imputing missing values
# Since, ContractRenewal and DataPlan are binary, we cannot substitute with mean values for these 2 variables. We will impute these two variables with their respective modal values.
# +
# Compute Mode for the 'Contract Renewal' and 'DataPlan' columns and impute the missing values
cols = ['ContractRenewal','DataPlan']
for column in cols:
cell_df.isnull().sum()
# -
# Now let us impute the rest of the continuous variables with the median. For that we are going to use the SimpleImputer sub module from sklearn.
from sklearn.impute import SimpleImputer
SI = SimpleImputer(strategy='median')
# +
#Now we need to fit and transform our respective data set to fill the missing values with the corresponding 'median' values
# -
# ### Checking for Duplicates
# Are there any duplicates ?
dups = cell_df.duplicated()
print('Number of duplicate rows = %d' % (dups.sum()))
# ### Proportion in the Target classes
cell_df.Churn.value_counts(normalize=True)
# ### Distribution of the variables Check using Histogram
cell_df[['AccountWeeks','DataUsage','CustServCalls','DayMins','DayCalls','MonthlyCharge','OverageFee','RoamMins']].hist();
# ### Outlier Check using boxplots
# +
cols=['AccountWeeks','DataUsage','CustServCalls','DayMins','DayCalls','MonthlyCharge','OverageFee','RoamMins'];
for i in cols:
sns.boxplot(cell_df[i])
plt.show()
# -
# ### Bi-Variate Analysis with Target variable
# <b>Account Weeks and Churn</b>
sns.boxplot(cell_df['Churn'],cell_df['AccountWeeks'])
# <b>Data Usage against Churn</b>
sns.boxplot(cell_df['Churn'],cell_df['DataUsage'])
# <b>DayMins against Churn</b>
sns.boxplot(cell_df['Churn'],cell_df['DayMins'])
# <b>DayCalls against Churn</b>
sns.boxplot(cell_df['Churn'],cell_df['DayCalls'])
# <b>MonthlyCharge against Churn</b>
sns.boxplot(cell_df['Churn'],cell_df['MonthlyCharge'])
# <b>OverageFee against Churn</b>
sns.boxplot(cell_df['Churn'],cell_df['OverageFee'])
# <b>RoamMins against Churn</b>
sns.boxplot(cell_df['Churn'],cell_df['RoamMins'])
# <b>CustServCalls against Churn</b>
sns.boxplot(cell_df['Churn'],cell_df['CustServCalls'])
# <b>Contract Renewal against Churn</b>
sns.countplot(cell_df['ContractRenewal'],hue=cell_df['Churn'])
# <b>Data Plan against Churn</b>
sns.countplot(cell_df['DataPlan'],hue=cell_df['Churn'])
# ### Train (70%) - Test(30%) Split
# ### LDA Model
# +
#Build LDA Model and fit the data
# -
# ### Prediction and Evaluation on both Training and Test Set using Confusion Matrix, Classification Report and AUC-ROC.
# +
# Predict it
# +
# Evaluation
| M5 Pridictive Modeling/M5 W3 Linear Dicriminant Analytics LDA/Predictive Modelling - Linear Discriminant Analysis - Student Version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Import Libraries
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
from sklearn.svm import SVR
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error
from sklearn.metrics import median_absolute_error
# +
#load boston data
BostonData = load_boston()
#X Data
X = BostonData.data
#y Data
y= BostonData.target
# +
#Splitting data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=40, shuffle=True)
# +
#Applying SVR Model
SVRModel = SVR(kernel='linear', C=1.0, shrinking=True, cache_size=300)
SVRModel.fit(X_train, y_train)
# +
#Calculating Details
print('SVRModel Train Score is : ' , SVRModel.score(X_train, y_train))
print('SVRModel Test Score is : ' , SVRModel.score(X_test, y_test))
# -
#Calculating Prediction
y_pred = SVRModel.predict(X_test)
print('Predicted Value for SVRModel is : ' , y_pred[:10])
#Calculating Mean Absolute Error
MAEValue = mean_absolute_error(y_test, y_pred, multioutput='uniform_average') # it can be raw_values
print('Mean Absolute Error Value is : ', MAEValue)
#Calculating Mean Squared Error
MSEValue = mean_squared_error(y_test, y_pred, multioutput='uniform_average') # it can be raw_values
print('Mean Squared Error Value is : ', MSEValue)
#Calculating Median Squared Error
MdSEValue = median_absolute_error(y_test, y_pred)
print('Median Squared Error Value is : ', MdSEValue )
# ## BY : KhaledNada
| Sklearn/SVR & SVC/2.4.2 SVR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.1 - Introduction to iterators
# #### > Iterating with a for loop
employees = ['Nick','Lore','Hugo']
for employee in employees:
print(employee, end=" ")
# #### > Iterating with a for loop
for letter in 'DataCamp':
print(letter)
for i in range(4):
print(i)
# #### > Iterating over iterables: next()
word = 'Da'
it = iter(word)
next(it)
print(it)
next(it)
next(it)
# #### > Iterating at once with *
word ='Data'
it = iter(word)
print(*it)
print(*it)
# #### > Iterating over dictionaries
pythonistas = {'hugo': 'bowne-anderson','francis': 'castro'}
for key, value in pythonistas.items():
print(key, value)
# #### > Iterating over file connections
file = open('file.txt')
it = iter(file)
print(next(it))
#This is the first line
print(next(it))
#This is the second line
# # 1.2 - Playing with iterators
# #### > Using enumerate()
avengers = ['hawkeye', 'iron man', 'thor', 'quicksilver']
e = enumerate(avengers)
print(type(e))
e_list = list(e)
print(e_list)
# #### > enumerate() and unpack
avengers = ['hawkeye', 'iron man', 'thor', 'quicksilver']
for index, value in enumerate(avengers):
print(index, value)
for index, value in enumerate(avengers, start=10):
print(index, value)
# #### > Using zip()
avengers = ['hawkeye', 'iron man', 'thor', 'quicksilver']
names = ['barton', 'stark', 'odinson', 'maximoff']
z = zip(avengers, names)
print(type(z))
z_list = list(z)
print(z_list)
# #### > zip() and unpack
avengers = ['hawkeye', 'iron man', 'thor', 'quicksilver']
names = ['barton', 'stark', 'odinson', 'maximoff']
for z1, z2 in zip(avengers, names):
print(z1, z2)
# #### > Print zip with *
avengers = ['hawkeye', 'iron man', 'thor', 'quicksilver']
names = ['barton', 'stark', 'odinson', 'maximoff']
z = zip(avengers, names)
print(*z)
# # 1.3 - Using iterators to load large files into memory
import pandas as pd
result = []
for chunk in pd.read_csv('data.csv', chunksize=1000):
result.append(sum(chunk['x']))
total = sum(result)
print(total)
import pandas as pd
total = 0
for chunk in pd.read_csv('data.csv', chunksize=1000):
total += sum(chunk['x'])
print(total)
# # 2.1 - List comprehensions
# #### > Populate a list with a for loop
nums = [12, 8, 21, 3, 16]
new_nums = []
for num in nums:
new_nums.append(num + 1)
print(new_nums)
# #### > A list comprehension
nums = [12, 8, 21, 3, 16]
new_nums = [num + 1 for num in nums]
print(new_nums)
# #### > For loop and list comprehension syntax
new_nums = [num + 1 for num in nums]
for num in nums:
new_nums.append(num + 1)
print(new_nums)
# #### > List comprehension with range()
result = [num for num in range(11)]
print(result)
# #### > Nested loops (1)
pairs_1 = []
for num1 in range(0, 2):
for num2 in range(6, 8):
pairs_1.append(num1, num2)
print(pairs_1)
# #### > Nested loops (2)
pairs_2 = [(num1, num2) for num1 in range(0, 2) for num2 in range(6,8)]
print(pairs_2)
# # 2.2 - Advanced comprehensions
# #### > Conditionals in comprehensions
[num ** 2 if num % 2 == 0 else 0 for num in range(10)]
# #### > Dict comprehensions
pos_neg = {num: -num for num in range(9)}
print(pos_neg)
print(type(pos_neg))
# # 2.3 - Introduction to generator expressions
# #### > Generator expressions
[2 * num for num in range(10)]
(2 * num for num in range(10))
# #### > Printing values from generators (1)
result = (num for num in range(6))
for num in result:
print(num)
result = (num for num in range(6))
print(list(result))
# #### > Printing values from generators (2)
result = (num for num in range(6))
print(next(result))
print(next(result))
print(next(result))
print(next(result))
print(next(result))
print(next(result))
# #### > Conditionals in generator expressions
even_nums = (num for num in range(10) if num % 2 == 0)
print(list(even_nums))
# #### > Build a generator function
def num_sequence(n):
"""Generate values from 0 to n."""
i = 0
while i < n:
yield i
i += 1
# #### > Use a generator function
result = num_sequence(5)
print(type(result))
for item in result:
print(item)
# # 3.1 - Welcome to the case study!
# #### > Using zip()
avengers = ['hawkeye', '<NAME>', 'thor', 'quicksilver']
names = ['barton', 'stark', 'odinson', 'maximoff']
z = zip(avengers, names)
print(type(z))
print(list(z))
# #### > Defining a function
def raise_both(value1, value2):
"""Raise value1 to the power of value2 and vice versa."""
new_value1 = value1 ** value2
new_value2 = value2 ** value1
new_tuple = (new_value1, new_value2)
return new_tuple
# # 3.2 - Using Python generators for streaming data
# #### > Build a generator function
def num_sequence(n):
"""Generate values from 0 to n."""
i = 0
while i < n:
yield i
i += 1
# # 3.3 - Using pandas 'read_csv' iterator for streaming data
# +
# read_csv() function and chunk_size argument
# -
| 08. Python Data Science Toolbox (Part 2)/Python Data Science Toolbox (Part 2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from selenium.common.exceptions import TimeoutException
import time
results_base_url = "https://elections.miottawa.org/ElectionResults/"
summary_base_url = "https://elections.miottawa.org//ElectionResults/Election/Summary/"
export_base_url = "https://elections.miottawa.org/ElectionResults/Election/Export/"
browser = webdriver.Firefox()
browser.get(results_base_url)
election_select = browser.find_element_by_id("ddElections")
election_options = election_select.find_elements_by_tag_name("option")
election_shorts = list()
for election_option in election_options:
if election_option.text=="Select":
continue
election_shorts.append(election_option.get_property("value"))
election_shorts
import requests
import os
for election_short in election_shorts:
r = requests.get(export_base_url+election_short)
with open(election_short+".xlsx", "wb") as fid:
fid.write(r.content)
| OttawaCountySelenium-Copy5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import panel as pn
from awesome_panel_extensions.widgets.pivot_table import PivotTable # Please note you need to import extensions before you run pn.extension("perspective")
# Expected to work for Panel >= 0.10
# C.f. https://github.com/holoviz/panel/issues/1529#event-3682125861
# pn.extension("perspective")
# Works for Panel < 0.10
PivotTable.config()
pn.extension()
from awesome_panel_extensions.awesome_panel import notebook
notebook.Header(folder="examples/reference/widgets", notebook="PivotTable.ipynb")
# -
# # PivotTable - Reference Guide
#
# [PivotTable.js](https://pivottable.js.org/examples/) is provides an awesome interactive pivot tables and charts.
#
# <img src="https://raw.githubusercontent.com/MarcSkovMadsen/awesome-panel-extensions/master/assets/videos/pivottable-official.gif" style="max-height:400px">
#
# The `PivotTable` widget enables the use of `pivottable.js` in Panel apps.
#
# #### Parameters:
#
# * **``value``** (DataFrame): The data loaded to the viewer.
#
# The `PivotTable` has the same layout and styling parameters as most other widgets. For example `width` and `sizing_mode`.
#
# ___
#
# Let's start by importing the **dependencies**
import param
import pandas as pd
import panel as pn
import time
# Let's load **S&P500 financials data**
# Source: https://datahub.io/core/s-and-p-500-companies-financials
DATA = "https://raw.githubusercontent.com/MarcSkovMadsen/awesome-panel/master/application/pages/awesome_panel_express_tests/PerspectiveViewerData.csv"
dataframe = pd.read_csv(DATA)
# Let's create a **PivotTable app** to display the data
DARK_BACKGROUND = "rgb(42, 44, 47)" # pylint: disable=invalid-name
top_app_bar = pn.Row(
pn.layout.Spacer(width=10),
pn.pane.HTML("<h1 style='color:white'> PivotTable.js</h1>", align="center"),
pn.layout.HSpacer(),
pn.pane.PNG("https://panel.holoviz.org/_static/logo_horizontal.png", height=50, align="center"),
pn.layout.Spacer(width=25),
background=DARK_BACKGROUND,
)
# pn.config.sizing_mode = "stretch_width"
# Source: https://datahub.io/core/s-and-p-500-companies-financials
data = "https://raw.githubusercontent.com/MarcSkovMadsen/awesome-panel/master/application/pages/awesome_panel_express_tests/PerspectiveViewerData.csv"
dataframe = pd.read_csv(data)
columns = [
"Name",
"Symbol",
"Sector",
"Price",
"52 Week Low",
"52 Week High",
"Price/Earnings",
"Price/Sales",
"Price/Book",
"Dividend Yield",
"Earnings/Share",
"Market Cap",
"EBITDA",
"SEC Filings",
]
dataframe=dataframe[columns]
pivot_table = PivotTable(
height=500,
value=dataframe.copy(deep=True),
sizing_mode="stretch_width",
)
app = pn.Column(
top_app_bar,
pn.Row(
pivot_table,
sizing_mode="stretch_width",
),
height=800,
sizing_mode="stretch_width",
)
app
# ## Known Issues
#
# - None so far
#
# ## Roadmap
#
# - Add more functionality like ability to set rows, column etc. And include plot options.
# - Get this included in the Panel package.
#
# ## Resources
#
# - [Pivottable.js](https://pivottable.js.org)
| examples/reference/widgets/PivotTable.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
print('This is a string {}'.format('INSERTED'))
print('The {} {} {}'.format('fox','brown','quick'))
print('The {2} {1} {0}'.format('fox','brown','quick'))
print('The {0} {0} {0}'.format('fox','brown','quick'))
print('The {q} {b} {f}'.format(f='fox',b='brown',q='quick'))
print('The {f} {f} {f}'.format(f='fox',b='brown',q='quick'))
# #### Float formatting "{value:width.precision f}"
result = 100/777
result
print("The result was {r:1.5f}".format(r=result))
result = 104.12345
print("The result was {r:1.2f}".format(r=result))
name = "Harshit"
print(f'Hello, his name is {name}')
name = "Sam"
age = 3
print(f'{name} is {age} years old')
| Section 3: Python Object and Data Structure Basics/Print Formatting and Strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Simulating From the Null Hypothesis
#
# Load in the data below, and use the exercises to assist with answering the quiz questions below.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
np.random.seed(42)
full_data = pd.read_csv('coffee_dataset.csv')
sample_data = full_data.sample(200)
# -
# `1.` If you were interested in studying whether the average height for coffee drinkers is the same as for non-coffee drinkers, what would the null and alternative hypotheses be? Write them in the cell below, and use your answer to answer the first quiz question below.
# **Since there is no directional component associated with this statement, a not equal to seems most reasonable.**
#
# $$H_0: \mu_{coff} - \mu_{no} = 0$$
#
#
# $$H_1: \mu_{coff} - \mu_{no} \neq 0$$
#
#
# **$\mu_{coff}$ and $\mu_{no}$ are the population mean values for coffee drinkers and non-coffee drinkers, respectivley.**
# `2.` If you were interested in studying whether the average height for coffee drinkers is less than non-coffee drinkers, what would the null and alternative be? Place them in the cell below, and use your answer to answer the second quiz question below.
# **In this case, there is a question associated with a direction - that is the average height for coffee drinkers is less than non-coffee drinkers. Below is one of the ways you could write the null and alternative. Since the mean for coffee drinkers is listed first here, the alternative would suggest that this is negative.**
#
# $$H_0: \mu_{coff} - \mu_{no} \geq 0$$
#
#
# $$H_1: \mu_{coff} - \mu_{no} < 0$$
#
#
# **$\mu_{coff}$ and $\mu_{no}$ are the population mean values for coffee drinkers and non-coffee drinkers, respectivley.**
# `3.` For 10,000 iterations: bootstrap the sample data, calculate the mean height for coffee drinkers and non-coffee drinkers, and calculate the difference in means for each sample. You will want to have three arrays at the end of the iterations - one for each mean and one for the difference in means. Use the results of your sampling distribution, to answer the third quiz question below.
# +
nocoff_means, coff_means, diffs = [], [], []
for _ in range(10000):
bootsamp = sample_data.sample(200, replace = True)
coff_mean = bootsamp[bootsamp['drinks_coffee'] == True]['height'].mean()
nocoff_mean = bootsamp[bootsamp['drinks_coffee'] == False]['height'].mean()
# append the info
coff_means.append(coff_mean)
nocoff_means.append(nocoff_mean)
diffs.append(coff_mean - nocoff_mean)
# -
np.std(nocoff_means) # the standard deviation of the sampling distribution for nocoff
np.std(coff_means) # the standard deviation of the sampling distribution for coff
np.std(diffs) # the standard deviation for the sampling distribution for difference in means
plt.hist(nocoff_means, alpha = 0.5);
plt.hist(coff_means, alpha = 0.5); # They look pretty normal to me!
plt.hist(diffs, alpha = 0.5); # again normal - this is by the central limit theorem
# `4.` Now, use your sampling distribution for the difference in means and [the docs](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.random.normal.html) to simulate what you would expect if your sampling distribution were centered on zero. Also, calculate the observed sample mean difference in `sample_data`. Use your solutions to answer the last questions in the quiz below.
# ** We would expect the sampling distribution to be normal by the Central Limit Theorem, and we know the standard deviation of the sampling distribution of the difference in means from the previous question, so we can use this to simulate draws from the sampling distribution under the null hypothesis. If there is truly no difference, then the difference between the means should be zero.**
null_vals = np.random.normal(0, np.std(diffs), 10000) # Here are 10000 draws from the sampling distribution under the null
plt.hist(null_vals); #Here is the sampling distribution of the difference under the null
| Practical_Statistics/Hypothesis_Testing/Simulating From the Null.ipynb |
# +
# Plots the posterior and plugin predictives for the Beta-Binomial distribution.
import numpy as np
import matplotlib.pyplot as plt
try:
import probml_utils as pml
except ModuleNotFoundError:
# %pip install -qq git+https://github.com/probml/probml-utils.git
import probml_utils as pml
from scipy.special import comb, beta
from scipy.stats import binom
N = 10 # Future sample size M
# Hyperparameters
a = 1
b = 1
N1 = 4
N0 = 1
ind = np.arange(N + 1)
post_a = a + N1
post_b = b + N0
# Compound beta-binomial distribution
distribution = []
for k in range(N + 1):
distribution.append(comb(N, k) * beta(k + post_a, N - k + post_b) / beta(post_a, post_b))
fig, ax = plt.subplots()
rects = ax.bar(ind, distribution, align="center")
ax.set_title("posterior predictive")
ax.set_xticks(list(range(N + 1)))
ax.set_xticklabels(list(range(N + 1)))
pml.savefig("BBpostpred.pdf")
plt.show()
# Plugin binomial distribution
mu = (post_a - 1) / float(post_a + post_b - 2) # MAP estimate
distribution = []
rv = binom(N, mu)
for k in range(N + 1):
distribution.append(rv.pmf(k))
fig, ax = plt.subplots()
rects = ax.bar(ind, distribution, align="center")
ax.set_title("plugin predictive")
ax.set_xticks(list(range(N + 1)))
ax.set_xticklabels(list(range(N + 1)))
pml.savefig("BBpluginpred.pdf")
plt.show()
| notebooks/book1/04/beta_binom_post_pred_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="w-A676oSu4HF"
# # HMM inference and MAP estimation in Gilbert-Elliot channel model
# ## TensorFlow implementation of potential based HMM and the parallel version of it.
#
# This notebook implements the algorithms from Ref.[2]
#
# [2] <NAME>, <NAME> and <NAME> (2021). __Temporal Parallelization of Inference in Hidden Markov Models__. *__IEEE Transactions on Signal Processing__*, Volume: 69, Pages 4875-4887. DOI: [10.1109/TSP.2021.3103338](https://doi.org/10.1109/TSP.2021.3103338)
#
#
# + [markdown] id="g8kqzPaGatUu"
# ### Imports and utilities
# + id="vK5Nq0QJu_6j"
# Python specific imports that will make our job easier and our code prettier
from collections import namedtuple
from functools import partial
import math
import time
from tqdm.auto import trange, tqdm
# TF specific imports that we will use to code the logic
from tensorflow import function
import tensorflow as tf
import tensorflow_probability as tfp
# Auxiliary libraries that we will use to report results and create the data
import matplotlib.pyplot as plt
import numpy as np
import scipy as sc
# Utilities
mm = tf.linalg.matmul
mv = tf.linalg.matvec
# device = '/TPU:0'
device = '/GPU:0'
# check if device is available
# tf.test.is_gpu_available()
# + [markdown] id="WHKrIHFHwy0Z"
# ### Model
# + [markdown] id="7ZuiV8_QatUw"
# We consider the Gilbert-Elliot channel model. This is a classical model used in the transmission of signals in digital communication channels. We consider two inputs: one for input signal and another for channel regime signal. They are binary signals and represented by $b_k$ and $s_k$, respectively.
#
# The input signal $b_k$ is flipped by an idependendent error. We can model this as
# $$
# y_k = b_k \oplus v_k
# $$
#
# The regime channel signal $s_k$ is represented as two states hidden Markov model, which can have either a good or a bad channel condition. The good state has a low probability error, while the bad state can have a high probability error. If $v_k = 1$ i.e. an error occurs, where $y_k \neq b_k$, then
#
# - $q_0$ represents the probability of the error in the good state.
# - $q_1$ represents the probability of the error in the bad state.
#
# The transition probability can be represented as
#
# - $p_0$: transition from high error state (bad state) to low error state (good state).
# - $p_1$: transition from low error state (good state) to high error state (bad state).
# - $p_2$: switching probability of $b_k$.
#
# The joint model $x_k = (s_k, b_k)$ is a four-state Markov chain. That is,
# $$
# x_k = \{(0, 0), (0, 1), (1, 0), (1, 1)\}
# $$
# and encoded as $\{0, 1, 2, 3\}$. Our transition and observation matrices are as follows:
#
#
# $$
# \Pi = p(x_k \mid x_{k-1}) = \begin{pmatrix}
# (1 - p_0)(1 - p_2) & p_0 (1 - p_2) & (1 - p_0)p_2 & p_0 p_2 \\
# p_1 (1 - p_2) & (1-p_1) (1-p_2) & p_1 p_2 & (1- p_1) p_2 \\
# (1 - p_0) & p_0 p_2 & (1 - p_0) (1 - p_2) & p_0 (1 - p_2) \\
# p_1 p_2 & (1 - p_1)p_2 & p_1 (1 - p_2) & (1 - p_1) (1 - p_2)
# \end{pmatrix}
# $$
#
# and
#
# $$
# O = p(y_k\mid x_k) = \begin{pmatrix}
# (1 - q_0) & q_0 \\
# (1 - q_1) & q_1 \\
# q_0 & (1 - q_0) \\
# q_1 & (1 - q_1)
# \end{pmatrix}
# $$
#
#
#
#
# + id="GiqvK_b_wHVK"
class HMMModel:
def __init__(self, p0=0.03, p1=0.1, p2=0.05, q0=0.01, q1=0.1):
"""Gilbert-Elliot channel model.
:math:``
p0: `P(S_{k+1} = 1 | S_{k} = 0)`
p1: `P(S_{k+1} = 0 | S_{k} = 1)`
p2: `P(B_{k+1} = 1 | B_{k} = 0) = P(B_{k+1} = 0 | B_{k} = 1)`
q0: `P(Y_k != b | B_k = b, S_k = 0)`
q1: `P(Y_k != b | B_k = b, S_k = 1)`
"""
# transition matrix
self.np_Pi = np.array([
[(1-p0)*(1-p2), p0*(1-p2), (1-p0)*p2, p0*p2],
[p1*(1-p2), (1-p1)*(1-p2), p1*p2, (1-p1)*p2],
[(1-p0)*p2, p0*p2, (1-p0)*(1-p2), p0*(1-p2)],
[p1*p2, (1-p1)*p2, p1*(1-p2), (1-p1)*(1-p2)]
])
# observation matrix
self.np_Po = np.array([
[(1-q0), q0],
[(1-q1), q1],
[q0, (1-q0)],
[q1, (1-q1)]
])
# prior
self.np_prior = np.array([0.25, 0.25, 0.25, 0.25])
# convert to tensor
self.Pi = tf.convert_to_tensor(self.np_Pi)
self.Po = tf.convert_to_tensor(self.np_Po)
self.prior = tf.convert_to_tensor(self.np_prior)
# dimensions
self.Pi_dim = self.Pi.shape[-1]
self.Po_dim = self.Po.shape[-1]
self.prior_dim = self.prior.shape[-1]
# transition, observation, and prior in log-space
self.log_Pi = tf.math.log(self.Pi)
self.log_Po = tf.math.log(self.Po)
self.log_prior = tf.math.log(self.prior)
# + id="WbLLiUbgbGVW"
def catrnd(rng, p):
if p.ndim > 1:
p = p.squeeze()
return rng.choice(p.shape[0], size=1, p=p)
def simulate_data(model: HMMModel, steps:int, seed:int=1):
rng = np.random.default_rng(seed)
X = np.zeros((steps, 1))
Y = np.zeros(steps, dtype=np.int32)
x = catrnd(rng, model.np_prior)
for k in range(steps):
if k > 0:
x = catrnd(rng, model.np_Pi[x])
y = catrnd(rng, model.np_Po[x])
X[k] = x
Y[k] = y
return X, Y
# + [markdown] id="Mkl1DYsDbJz2"
# #### Initialize the parameters
# + id="fjBXhfmWbOBO"
log10T = 5
nsteps = 100
T = np.arange(10 ** log10T)+1
# + [markdown] id="7sJxl956bkWC"
# #### Initialize the model
# + id="BtCSf2wWbjuw"
model = HMMModel()
# + [markdown] id="LG2nnFzobs54"
# #### Simulate the data
# + id="_LJMh9yLbtaV"
X,Y = simulate_data(model,10 ** log10T)
# + [markdown] id="uWLHqO9Vb4Qd"
# #### Plot the simulated data
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="xK3HGrJib3t_" outputId="5e40afc1-b3bb-4e49-95dd-1daaca9dc48d"
plt.plot(T[:nsteps], X[:nsteps], '*-', T[:nsteps], Y[:nsteps], 'o--')
plt.xlabel('Number of steps')
plt.ylabel('States')
plt.legend(['True states', 'Observed states'])
# + [markdown] id="1Ffp-XeIFXM8"
# ### Reference implementations for comparison and debugging
# Here, we have implemented forward-backward potential based filter-smoother summarized in Algorithm 1 of the article.
# + id="b_WJXy1-Fawx"
def ref_fwdbwd_filter_smoother(model, Y):
"""Reference Forward-Backward Filter and smoother
Parameters
----------
model:
HMM model
Y: array_like
Observations
Returns
-------
fest:
filtering estimates
sest:
smoother estimates
all_psi_f:
all forward potentials
all_psi_b:
all backward potentials
"""
all_psi_f = np.zeros((Y.shape[0], model.Pi_dim))
fest = np.zeros((Y.shape[0], model.Pi_dim))
for k in range(len(Y)):
if k > 0:
psi = model.np_Po[:, Y[k]] * model.np_Pi
psi_f = psi_f @ psi
else:
psi_f = model.np_Po[:, Y[k]] * model.np_prior
all_psi_f[k] = psi_f
fest[k] = psi_f / np.sum(psi_f)
psi_b = np.ones(psi_f.shape)
all_psi_b = all_psi_f.copy()
all_psi_b[-1] = psi_b
for k in reversed(range(len(Y)-1)):
psi = model.np_Po[:,Y[k+1]] * model.np_Pi
psi_b = psi @ psi_b
all_psi_b[k] = psi_b
sest = []
for f,b in zip(all_psi_f, all_psi_b):
p = f * b
p = p / np.sum(p)
sest.append(p)
return fest, sest, all_psi_f, all_psi_b
# + [markdown] id="T6-iEMuqatUy"
# ### Plots for reference Bayesian and forward-backward filters and smoothers
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="ZNh8-XjSs7O6" outputId="4eb9b873-eaa6-439e-fcae-85785595e0f3"
ref_fwdbwd_fs, ref_fwdbwd_ss, ref_fwdbwd_psi_f, ref_fwdbwd_psi_b = ref_fwdbwd_filter_smoother(model, Y)
print()
plt.clf()
plt.subplots_adjust(hspace=.5)
for i in range(4):
plt.subplot(2, 2, i+1)
plt.plot(T[:nsteps], [p[i] for p in ref_fwdbwd_fs[:nsteps]], T[:nsteps], [p[i] for p in ref_fwdbwd_ss[:nsteps]])
plt.title(('State x_k = %d ' % i ))
plt.legend(['Filtering', 'Smoothing'])
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="wchn8C20ra1H" outputId="af451580-41b1-4ef0-a58b-302532660b9a"
print('Is Nan?', tf.math.is_nan(tf.reduce_sum(ref_fwdbwd_ss)))
# + [markdown] id="03HAT2p6gc6F"
# The results of the smoothing distribution contain 'nan' values. Therefore, we were unable to view the smoother result.
# + [markdown] id="nQjmem2ZwHLL"
# ### Utilities for log-space computations
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="5lPjL7LvwMiq" outputId="969419d5-5935-4254-ce15-6a93aa0d78c7"
@tf.function
def log_mv(log_A, log_b, transpose_a=False):
Amax = tf.reduce_max(log_A, axis=(-1, -2), keepdims=True)
bmax = tf.reduce_max(log_b, axis=(-1), keepdims=True)
return tf.squeeze(Amax, axis=-1) + bmax + tf.math.log(mv(tf.math.exp(log_A - Amax), tf.math.exp(log_b - bmax), transpose_a=transpose_a))
@tf.function
def semilog_mv(A, log_b, transpose_a=False):
bmax = tf.reduce_max(log_b, axis=(-1), keepdims=True)
return bmax + tf.math.log(mv(A, tf.math.exp(log_b - bmax), transpose_a=transpose_a))
@tf.function
def log_mm(log_A,log_B,transpose_a=False,transpose_b=False):
Amax = tf.reduce_max(log_A, axis=(-1, -2), keepdims=True)
Bmax = tf.reduce_max(log_B, axis=(-1, -2), keepdims=True)
return Amax + Bmax + tf.math.log(mm(tf.math.exp(log_A - Amax), tf.math.exp(log_B - Bmax), transpose_a=transpose_a, transpose_b=transpose_b))
@tf.function
def log_normalize(log_p):
pmax = tf.reduce_max(log_p, axis=(-1), keepdims=True)
temp = tf.math.exp(log_p - pmax)
return tf.math.log(temp / tf.reduce_sum(temp, axis=-1, keepdims=True))
print('Test utility functions:')
with tf.device('/CPU:0'):
tf.random.set_seed(5)
A = tf.random.uniform(shape=[4, 4])
B = tf.random.uniform(shape=[4, 4])
log_A = tf.math.log(A)
log_B = tf.math.log(B)
print(mm(A, B))
print(tf.math.exp(log_mm(log_A, log_B)))
b = tf.random.uniform(shape=[4])
log_b = tf.math.log(b)
print(mv(A, b))
print(tf.math.exp(log_mv(log_A, log_b)))
print(tf.math.exp(semilog_mv(A, log_b)))
print(b / tf.reduce_sum(b, keepdims=True))
print(tf.math.exp(log_normalize(log_b)))
# + [markdown] id="WkV6b5vXhJFB"
# ### Sequential potential based filter and smoother
#
#
# This is done by forward and backward pass.
#
# Here, we have implemented forward-backward potential based filter-smoother summarized in Algorithm 1 of the article. This is implemented in tensorflow by leveraging the built-in sequential scan operator.
#
#
# + [markdown] id="-JCtZ9JveVwp"
# #### Forward pass routine
# + id="9p0Be6VkhOq-"
@tf.function
def potential_forward_pass(model, obs):
def body(carry, y):
k, p = carry # k is the iteration
p = tf.cond(k > 0, lambda: mv(model.Pi, p, transpose_a=True), lambda: p)
p = model.Po[:, y] * p
return (k+1, p)
ks, psi_f = tf.scan(body, obs, (0, model.prior))
return psi_f
# + [markdown] id="YAp0P7-_ecZu"
# #### Backward pass routine
# + id="lrRiXRmIebr5"
@tf.function
def potential_backward_pass(model, obs):
psi_full = tf.expand_dims(model.Po, 0) * tf.expand_dims(model.Pi, -1)
def body(carry, y):
p = carry
psi = psi_full[..., y]
p = mv(psi, p)
return p
psi_b_end = tf.ones_like(model.prior)
psi_b_rest = tf.scan(body, obs[1:], psi_b_end, reverse=True)
psi_b = tf.concat([psi_b_rest, tf.expand_dims(psi_b_end, 0)], 0)
return psi_b
# + [markdown] id="VZ1IzQiQeil2"
# #### Filter-smoother routine
# + id="d5JZTB9KeoqW"
@tf.function
def potential_smoother(model, obs):
psi_f = potential_forward_pass(model, obs)
psi_b = potential_backward_pass(model, obs)
p = psi_f * psi_b
p = p / tf.reduce_sum(p, axis=-1, keepdims=True)
return p
# + [markdown] id="412gFPLieUfA"
# #### Test and compare the reference and potential based forward-backward filter-smoothers.
#
# + colab={"base_uri": "https://localhost:8080/"} id="h1UPK8GqkEol" outputId="7b4c92b2-e4ec-49c3-b14b-43c393867b01"
with tf.device('/CPU:0'):
psi_f = potential_forward_pass(model, Y)
print('Difference between forward pass of potential based and reference: ', np.abs(psi_f - ref_fwdbwd_psi_f).max())
psi_b = potential_backward_pass(model, Y)
print('Difference between backward pass of potential based and reference: ', np.abs(psi_b - ref_fwdbwd_psi_b).max())
potential_ss = potential_smoother(model, Y)
print('Difference between potential based and reference smoothers: ', np.abs(potential_ss - ref_fwdbwd_ss).max())
# + [markdown] id="XAg3lQP2GiyP"
# ### Log-space sequential potential based filter and smoother
# + [markdown] id="1xF9uTvDfjdO"
# #### Forward pass routine
# + id="xBcU0KaUGk-K"
@tf.function
def log_potential_forward_pass(model, obs):
def body(carry, y):
k, log_p = carry # k is the iteration
log_p = tf.cond(k > 0, lambda: semilog_mv(model.Pi, log_p, transpose_a=True), lambda: log_p)
log_p = model.log_Po[:, y] + log_p
return (k+1, log_p)
ks, log_psi_f = tf.scan(body, obs, (0, model.log_prior))
return log_psi_f
# + [markdown] id="QteRbRLqfmUO"
# #### Backward pass routine
# + id="sQz7uCM_fluJ"
@tf.function
def log_potential_backward_pass(model, obs):
psi_full = tf.expand_dims(model.Po, 0) * tf.expand_dims(model.Pi, -1)
def body(carry, y):
log_p = carry
log_p = semilog_mv(psi_full[..., y], log_p)
return log_p
log_psi_b_end = tf.zeros_like(model.log_prior)
log_psi_b_rest = tf.scan(body, obs[1:], log_psi_b_end, reverse=True)
log_psi_b = tf.concat([log_psi_b_rest, tf.expand_dims(log_psi_b_end, 0)], 0)
return log_psi_b
# + [markdown] id="f4aquYVBfoAt"
# #### Smoother routine
# + id="EHf5Y11HfoR9"
@tf.function
def log_potential_smoother(model, obs, max_parallel=0):
log_psi_f = log_potential_forward_pass(model, obs)
log_psi_b = log_potential_backward_pass(model, obs)
log_p = log_psi_f + log_psi_b
log_p = log_normalize(log_p)
return log_p
# + [markdown] id="XuJ0HREafsD9"
# #### Test sequential potential based smoother routine in log space
# + colab={"base_uri": "https://localhost:8080/"} id="zkR_e65vHUVt" outputId="6bf3a9a7-fb45-4e46-b16a-a15f724adb02"
with tf.device('/CPU:0'):
psi_f = potential_forward_pass(model, Y)
log_psi_f = log_potential_forward_pass(model, Y)
print('Difference between forward and log-space-forward potentials: ', np.abs(psi_f - tf.math.exp(log_psi_f)).max())
psi_b = potential_backward_pass(model, Y)
log_psi_b = log_potential_backward_pass(model, Y)
print('Difference between backward and log-space-backward potentials: ', np.abs(psi_b - tf.math.exp(log_psi_b)).max())
potential_ss = potential_smoother(model, Y)
log_potential_ss = log_potential_smoother(model, Y)
print('Difference between potential based smoothers and log-space-smoothers: ', np.abs(potential_ss - tf.math.exp(log_potential_ss)).max())
print('Is the result of log-space smoothers contains NaNs? ', tf.math.is_nan(tf.reduce_mean(log_potential_ss)))
# + [markdown] id="S-IIsQRlwqC9"
# ### Parallel potential based filter and smoother
#
# We have Sequential Bayesian filter and smoother as a baseline. Now, we implement a parallel version of this. The parallel version is similar to Algorithm 3 described in the paper. However, there are differences with classical parallel hmm (See phmm_classical.ipynb) implementation. Here, we consider potential-based implementation. Moreover, the forward algorithm is equivalent to HMM filtering algorithm. However, the backward algorithm corresponds to backward pass of two-filtering algorithm.
# + [markdown] id="aU8bhvqRh_c3"
# #### Forward part
# + [markdown] id="FuV4CluHiEaG"
# #### Initialization routine
# + id="EYqDl_T-wu2t"
@tf.function
def parallel_forward_potential_init_first(model, y):
prior = tf.tile(tf.expand_dims(model.prior, axis=1), [1, model.prior.shape[0]])
psi = tf.multiply(model.Po[:, y], prior)
return psi
@tf.function
def parallel_forward_potential_init_rest(model, obs):
temp = tf.gather(tf.transpose(model.Po), obs, axis=0)
psi = tf.expand_dims(temp, 1) * tf.expand_dims(model.Pi, 0)
return psi
@tf.function
def parallel_forward_potential_init(model, obs):
"""A parallel forward potential init routine to initialize first and generic elements.
"""
first_elem = parallel_forward_potential_init_first(model, obs[0])
generic_elems = parallel_forward_potential_init_rest(model, obs[1:])
return tf.concat([tf.expand_dims(first_elem, 0), generic_elems], axis=0)
# + [markdown] id="_XvWmKuoisHY"
# #### Potential operator
# + id="MmP1NY_risTf"
@tf.function
def parallel_potential_op(elemij, elemjk):
"""A parallel potential operator."""
return elemij@elemjk
# + [markdown] id="DRUxwLXQisiX"
# #### Forward potential routine
# + id="M01_sH8Fiswv"
@tf.function
def parallel_forward_potential(model, obs, max_parallel=10000):
"""A parallel forward potential routine to compute forward potentials.
This function implements Theorem 1 from the paper.
"""
initial_elements = parallel_forward_potential_init(model, obs)
final_elements = tfp.math.scan_associative(parallel_potential_op,
initial_elements,
max_num_levels=math.ceil(math.log2(max_parallel)))
return final_elements[:,0]
# + [markdown] id="3pzn9BdNitAd"
# #### Backward part
# + [markdown] id="sOg3hOe4jNqv"
# #### Backward initialization routine
# + id="TwiMTgzcitJg"
@tf.function
def parallel_backward_potential_init_last(model):
psi = tf.ones_like(model.Pi)
return psi
@tf.function
def parallel_backward_potential_init_rest(model, obs):
temp = tf.gather(tf.transpose(model.Po), obs, axis=0)
psi = tf.expand_dims(temp, 1) * tf.expand_dims(model.Pi, 0)
return psi
@tf.function
def parallel_backward_potential_init(model, obs):
"""A parallel backward potential init routine to initialize last and generic elements."""
last_elem = parallel_backward_potential_init_last(model)
generic_elems = parallel_backward_potential_init_rest(model, obs[1:])
return tf.concat([generic_elems, tf.expand_dims(last_elem, 0)], axis=0)
# + [markdown] id="BCenogN6joxy"
# #### Backward potential operator
# + id="a86HF0Fljo9X"
@tf.function
def parallel_backward_potential_op(elemjk, elemij):
"""A parallel backward potential operator."""
return elemij@elemjk
# + [markdown] id="iR0QG_QQjpKO"
# #### Backward potential routine
# + id="v-6ix57PjpUe"
@tf.function
def parallel_backward_potential(model, obs, max_parallel=100000):
"""A parallel backward potential routine to compute backward elements.
This function implements Theorem 2 from the paper.
"""
initial_elements = parallel_backward_potential_init(model, obs)
reversed_elements = tf.reverse(initial_elements, axis=[0])
final_elements = tfp.math.scan_associative(parallel_backward_potential_op,
reversed_elements,
max_num_levels=math.ceil(math.log2(max_parallel)))
final_elements = tf.reverse(final_elements, axis=[0])
return final_elements[:, :, 0]
# + [markdown] id="Y-SoS_5ZjpeW"
# #### Parallel potential routine
# + id="piTr3lHUjpm4"
@tf.function
def parallel_potential_smoother(model, obs, max_parallel=10000):
"""A parallel potential smoother routine to compute forward and backward elements.
See Eq. (22) from the paper.
"""
psi_f = parallel_forward_potential(model, obs, max_parallel)
psi_b = parallel_backward_potential(model, obs, max_parallel)
sest = psi_f * psi_b
sest = sest / tf.reduce_sum(sest, axis=1, keepdims=True)
return sest
# + colab={"base_uri": "https://localhost:8080/"} id="d5DszmzmxBIX" outputId="7b4ff3b9-0802-461e-aba2-23eea1cffc45"
with tf.device('/CPU:0'):
psi_f = parallel_forward_potential(model, Y, max_parallel=100000)
print('Difference between reference and parallel forward potentials: ', np.abs(psi_f - ref_fwdbwd_psi_f).max())
psi_b = parallel_backward_potential(model, Y, max_parallel=100000)
print('Difference between reference and parallel backward potentials: ', np.abs(psi_b - ref_fwdbwd_psi_b).max())
potential_ss = parallel_potential_smoother(model, Y, max_parallel=100000)
print('Difference between reference and parallel smoothers: ', np.abs(potential_ss - ref_fwdbwd_ss).max())
# + [markdown] id="YoIo52PGJmqy"
# ### Log-space parallel potential based filter and smoother
# + [markdown] id="subNyV0YkyVC"
# #### Forward part
# + [markdown] id="_BjIapsRkzVz"
# #### Initialization routine
# + id="k3FUeU9DJ9zW"
@tf.function
def log_parallel_forward_potential_init_first(model, y):
return tf.math.log(parallel_forward_potential_init_first(model, y))
@tf.function
def log_parallel_forward_potential_init_rest(model, obs):
return tf.math.log(parallel_forward_potential_init_rest(model, obs))
@tf.function
def log_parallel_forward_potential_init(model, obs):
"""A parallel forward potential routine in log space to initialize first and generic elements."""
first_elem = log_parallel_forward_potential_init_first(model, obs[0])
generic_elems = log_parallel_forward_potential_init_rest(model, obs[1:])
return tf.concat([tf.expand_dims(first_elem, 0), generic_elems], axis=0)
# + [markdown] id="YSyPR9E-lt6V"
# #### Potential operator
# + id="7fFLG0bOluBc"
@tf.function
def log_parallel_potential_op(elemij, elemjk):
"""A parallel potential operator in log space."""
return log_mm(elemij,elemjk)
# + [markdown] id="dtKSIBDvluIf"
# #### Parallel forward potential
# + id="lbXbe0YGluPb"
@tf.function
def log_parallel_forward_potential(model, obs, max_parallel=100000):
"""A parallel forward potential routine to compute forward elements in log space."""
initial_elements = log_parallel_forward_potential_init(model, obs)
final_elements = tfp.math.scan_associative(log_parallel_potential_op,
initial_elements,
max_num_levels=math.ceil(math.log2(max_parallel)))
return final_elements[:,0]
# + [markdown] id="wQm4EYJAluV0"
# #### Backward part
# + [markdown] id="fciK-Y2_m4JC"
# #### Initialization routines
# + id="4eY2A59JlugH"
@tf.function
def log_parallel_backward_potential_init_last(model):
log_psi = tf.zeros_like(model.Pi)
return log_psi
@tf.function
def log_parallel_backward_potential_init_rest(model, obs):
temp = tf.gather(tf.transpose(model.log_Po), obs, axis=0)
log_psi = tf.expand_dims(temp, 1) + tf.expand_dims(model.log_Pi, 0)
return log_psi
@tf.function
def log_parallel_backward_potential_init(model, obs):
"""A parallel backward potential init routine to initialize backward elements in log space."""
last_elem = log_parallel_backward_potential_init_last(model)
generic_elems = log_parallel_backward_potential_init_rest(model, obs[1:])
return tf.concat([generic_elems, tf.expand_dims(last_elem, 0)], axis=0)
# + [markdown] id="W2fl8FZGlum4"
# #### Backward potential operator
# + id="5twHO94nluuv"
@tf.function
def log_parallel_backward_potential_op(elemjk, elemij):
"""A parallel backward potential op routine in log space."""
return log_mm(elemij,elemjk)
# + [markdown] id="AodmoD8nl6AZ"
# #### Parallel backward potentials
# + id="GON_JNp1l6Hy"
@tf.function
def log_parallel_backward_potential(model, obs, max_parallel=10000):
"""A parallel backward potential routine to compute backward elements in log space."""
initial_elements = log_parallel_backward_potential_init(model, obs)
reversed_elements = tf.reverse(initial_elements, axis=[0])
final_elements = tfp.math.scan_associative(log_parallel_backward_potential_op,
reversed_elements,
max_num_levels=math.ceil(math.log2(max_parallel)))
final_elements = tf.reverse(final_elements, axis=[0])
return final_elements[:, :, 0]
# + [markdown] id="MiJkj10Pl6QH"
# #### Parallel potential smoother in log space
# + id="Fup9OrCTl6YX"
@tf.function
def log_parallel_potential_smoother(model, obs, max_parallel=10000):
"""A parallel potential smoother routine to compute forward and backward elements in log space."""
log_psi_f = log_parallel_forward_potential(model, obs, max_parallel)
log_psi_b = log_parallel_backward_potential(model, obs, max_parallel)
log_sest = log_psi_f + log_psi_b
log_sest = log_normalize(log_sest)
return log_sest
# + [markdown] id="oHV6uOERl9JP"
# #### Comparison with different implementation
# + colab={"base_uri": "https://localhost:8080/"} id="xHSNpjLCK_Zv" outputId="5edac0c9-60ff-4a98-a309-9ce909120b2b"
with tf.device('/CPU:0'):
psi_f = parallel_forward_potential(model, Y, max_parallel=100000)
log_psi_f = log_parallel_forward_potential(model, Y, max_parallel=100000)
print('Difference between parallel and log-parallel forward potentials: ', np.abs(psi_f - tf.math.exp(log_psi_f)).max())
psi_b = parallel_backward_potential(model, Y, max_parallel=100000)
log_psi_b = log_parallel_backward_potential(model, Y, max_parallel=100000)
print('Difference between parallel and log-parallel backward potentials: ', np.abs(psi_b - tf.math.exp(log_psi_b)).max())
potential_ss = parallel_potential_smoother(model, Y, max_parallel=100000)
log_potential_ss = log_parallel_potential_smoother(model, Y, max_parallel=100000)
print('Difference between parallel and log-parallel potential smoothers: ', np.abs(potential_ss - tf.math.exp(log_potential_ss)).max())
print('Is the result of log-space potential based smoothers contains NaNs? ', tf.math.is_nan(tf.reduce_mean(log_potential_ss)))
# + id="67ZJOauhvaJy"
def get_average_runtimes(func, n_iter, model, observations):
runtimes = np.empty(input_sizes.shape)
for i, input_size in tqdm(enumerate(input_sizes), total=runtimes.shape[0]):
observation_slice = observations[:input_size]
max_parallel = int(input_size)
_ = func(model, observation_slice, max_parallel=max_parallel) # compilation run
tic = time.time()
for _ in trange(n_iter, leave=False):
res = func(model, observation_slice, max_parallel=max_parallel)
runtimes[i] = (time.time() - tic) / n_iter
return runtimes
# + id="nMJyFdJ9vhnO"
input_sizes = np.logspace(2, log10T, num=20, base=10).astype(int)
n_iter = 100
n_iter_seq = 10
with tf.device('/CPU:0'):
cpu_Y = tf.constant(Y)
cpu_sequential_runtimes = get_average_runtimes(log_potential_smoother, n_iter_seq, model, cpu_Y)
cpu_parallel_runtimes = get_average_runtimes(log_parallel_potential_smoother, n_iter, model, cpu_Y)
with tf.device(device):
xpu_Y = tf.constant(Y)
xpu_sequential_runtimes = get_average_runtimes(log_potential_smoother, n_iter_seq, model, xpu_Y)
xpu_parallel_runtimes = get_average_runtimes(log_parallel_potential_smoother, n_iter, model, xpu_Y)
# + colab={"base_uri": "https://localhost:8080/", "height": 568} id="13DKAy5Dzxsa" outputId="dc48bc66-874e-4a3d-f3d2-31742472e0f6"
fig = plt.figure(1, dpi=150)
plt.loglog(input_sizes, cpu_sequential_runtimes, label="PS Sequential-CPU", linestyle="-.", linewidth=3)
plt.loglog(input_sizes, cpu_parallel_runtimes, label="PS Parallel-CPU", linewidth=3)
plt.legend()
plt.xlabel("Number of data points")
plt.ylabel("Average run time (seconds)")
plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 568} id="xHA77tPvw4S0" outputId="0acd5037-9dfb-4588-d655-b883f9ccc515"
fig = plt.figure(1, dpi=150)
plt.loglog(input_sizes, xpu_sequential_runtimes, label="PS Sequential-GPU", linestyle="-.", linewidth=3)
plt.loglog(input_sizes, xpu_parallel_runtimes, label="PS Parallel-GPU", linewidth=3)
plt.legend()
plt.xlabel("Number of data points")
plt.ylabel("Average run time (seconds)")
plt.grid()
plt.show()
| python/temporal-parallelization-inference-in-HMMs/phmm_potentials.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp data.metadatasets
# -
# # Metadatasets: a dataset of datasets
#
# > This functionality will allow you to create a dataset from data stores in multiple, smaller datasets.
# * I'd like to thank <NAME> and <NAME> for the work they've done to make this possible.
# * This functionality allows you to use multiple numpy arrays instead of a single one, which may be very useful in many practical settings. I've tested it with 10k+ datasets and it works well.
#export
from tsai.imports import *
from tsai.utils import *
from tsai.data.validation import *
from tsai.data.core import *
# +
#export
class TSMetaDataset():
" A dataset capable of indexing mutiple datasets at the same time!"
def __init__(self, dataset_list, **kwargs):
if not is_listy(dataset_list): dataset_list = [dataset_list]
self.datasets = dataset_list
self.split = kwargs['split'] if 'split' in kwargs else None
self.mapping = self._mapping()
if hasattr(dataset_list[0], 'loss_func'):
self.loss_func = dataset_list[0].loss_func
else:
self.loss_func = None
def __len__(self):
if self.split is not None:
return len(self.split)
else:
return sum([len(ds) for ds in self.datasets])
def __getitem__(self, idx):
if self.split is not None: idx = self.split[idx]
idx = listify(idx)
idxs = self.mapping[idx]
idxs = idxs[idxs[:, 0].argsort()]
self.mapping_idxs = idxs
ds = np.unique(idxs[:, 0])
b = [self.datasets[d][idxs[idxs[:, 0] == d, 1]] for d in ds]
output = tuple(map(torch.cat, zip(*b)))
return output
def _mapping(self):
lengths = [len(ds) for ds in self.datasets]
idx_pairs = np.zeros((np.sum(lengths), 2)).astype(np.int32)
start = 0
for i,length in enumerate(lengths):
if i > 0:
idx_pairs[start:start+length, 0] = i
idx_pairs[start:start+length, 1] = np.arange(length)
start += length
return idx_pairs
@property
def vars(self):
s = self.datasets[0][0][0] if not isinstance(self.datasets[0][0][0], tuple) else self.datasets[0][0][0][0]
return s.shape[-2]
@property
def len(self):
s = self.datasets[0][0][0] if not isinstance(self.datasets[0][0][0], tuple) else self.datasets[0][0][0][0]
return s.shape[-1]
class TSMetaDatasets(FilteredBase):
def __init__(self, metadataset, splits):
store_attr()
self.mapping = metadataset.mapping
def subset(self, i):
return type(self.metadataset)(self.metadataset.datasets, split=self.splits[i])
@property
def train(self):
return self.subset(0)
@property
def valid(self):
return self.subset(1)
# -
# Let's create 3 datasets. In this case they will have different sizes.
dsets = []
for i in range(3):
size = np.random.randint(50, 150)
X = torch.rand(size, 5, 50)
y = torch.randint(0, 10, (size,))
tfms = [None, TSClassification()]
dset = TSDatasets(X, y, tfms=tfms)
dsets.append(dset)
dsets
metadataset = TSMetaDataset(dsets)
metadataset, metadataset.vars, metadataset.len
# We'll apply splits now to create train and valid metadatasets:
splits = TimeSplitter()(metadataset)
splits
metadatasets = TSMetaDatasets(metadataset, splits=splits)
metadatasets.train, metadatasets.valid
dls = TSDataLoaders.from_dsets(metadatasets.train, metadatasets.valid)
xb, yb = first(dls.train)
xb, yb
# There also en easy way to map any particular sample in a batch to the original dataset and id:
dls = TSDataLoaders.from_dsets(metadatasets.train, metadatasets.valid)
xb, yb = first(dls.train)
mappings = dls.train.dataset.mapping_idxs
for i, (xbi, ybi) in enumerate(zip(xb, yb)):
ds, idx = mappings[i]
test_close(dsets[ds][idx][0].data, xbi)
test_close(dsets[ds][idx][1].data, ybi)
# For example the 3rd sample in this batch would be:
dls.train.dataset.mapping_idxs[2]
#hide
out = create_scripts(); beep(out)
| nbs/002c_data.metadatasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What makes a movie a commercial success?
#
# ###### A data science approach to dissect the economics of movie-making business
#
#
# ## Section 1: Business Understanding
#
# In this notebook, I will use the IMDb publlic dataset and visualization tools in order to answer improtant questions about what factors lead to a commercial success of a movie. The key questions tht I will try to answer are as follows:
# 1. Do certain movie genres make more money than others?
# 2. Does the amount of funding for a movie have an impact on commercial success?
# 3. What impact does duration have on the success of a movie?
# 4. Do viewer ratings have an impact on how well a movie will do in the Box Office?
# 5. Does the number of votes impact the success of a movie?
#
# Answering these questions could dramatically improve the decision processes underlying moie production.
#
#
#
# ## Section 2: Data Understanding
#
# The IMDB movie dataset covers over 80000 movies and accounts for all the basic attributes one would expect such as duration, genre, country, language, and gross income. The dataset can be accessed here: https://www.imdb.com/interfaces/
#
# The following codeblock enables me to import the data and convert into a Pandas Dataframe
#import necessary libraries
import pandas as pd
import os
import sys
import seaborn as sns
import matplotlib.pyplot as plt
from forex_python.converter import CurrencyRates
cwd = os.getcwd()
import re
import math
import numpy as np
pd.reset_option('^display.', silent=True)
np.set_printoptions(suppress=True)
imdb_df = pd.read_csv(cwd+'/datasets/imdb.csv') #import imdb csv
# ## Section 3: Data Preparation
# In this section, we will try to understand the dataset on a colummn by column basis and figure out which columns are valauble and how we could still make the most out of seemingly not valuable ones and also address issues related to missing data.
#
imdb_df.columns #Printing columns to understand the dataset
# +
imdb_df.isna().sum()/imdb_df.shape[0] * 100 #Let's get a macroview of which columns are useful and which ones aren't
# -
#we will drop the Nan rows from output and also from metscore since 84% of its values are missing
imdb_df = imdb_df.drop(columns=['metascore'])
# +
#Then, we will remove the rows from imdb_df that do not have worldwide & usa revenue numbers as this is the output we are looking to compare with
imdb_df = imdb_df.dropna(subset=['worlwide_gross_income'])
# -
#Genre column can serve as a great categorical variable
imdb_df['genre'] = imdb_df['genre'].str.replace(' ', '') # But first, we need to split the genres separated by commas
genre_encoded = imdb_df['genre'].str.get_dummies(sep=',') #We encode the genres by using get_dummies
genre_encoded
#Now, we will make use of the encoded genre column
imdb_df = pd.concat([imdb_df,genre_encoded], axis=1) #join the encoded data with original dataframe
imdb_df.drop(columns=['genre']) #drop the original genre column
# +
#Next, we will attempt at converting the income related columns to one unified currency - USD
c = CurrencyRates() #instantiating the forex conversion module
def get_symbol(price):
""" function for reading in the price and returning the currency
inputs:
- price: amount in local currency
outputs:
- currency: currency of the local price
"""
import re
pattern = r'(\D*)\d*\.?\d*(\D*)'
g = re.match(pattern,price).groups()
return g[0]
def return_USD(budget):
""" function for reading in the currency and converting to USD
inputs:
- price: amount in local currency
outputs:
- price_in_USD: amount in USD
"""
if budget!='nan':
if '$' not in budget:
try:
return c.get_rate(get_symbol(budget).strip(), 'USD') * int(re.findall('\d+', budget)[0])
except:
return float('NaN')
else:
return int(re.findall('\d+', budget)[0])
else:
return float('NaN')
# -
#lambda function for applying the USD conversion to the budget column
imdb_df['budget'] = imdb_df['budget'].apply(lambda x: return_USD(str(x)))
imdb_df
#similarly, we'll convert the worldwide_gross_income and usa_gross_icome to USD
imdb_df['worlwide_gross_income'] = imdb_df['worlwide_gross_income'].apply(lambda x: return_USD(str(x)))
imdb_df['usa_gross_income'] = imdb_df['usa_gross_income'].apply(lambda x: return_USD(str(x)))
imdb_df.to_csv(cwd+'/datasets/imdb_clean.csv') #we will save the cleaned up dataframe to a csv in order to create a milestone
# We will now address the next few steps sequentially for each question
#
# ### Section 3.1 Prepare data for question 1
# First question - Do certain genres make more $ than others?
#
#We will extract the grenres columns and save them into a new dataframe
imdb_genres_df = imdb_df[['worlwide_gross_income','Animation', 'Biography', 'Comedy', 'Crime', 'Documentary', 'Drama',
'Family', 'Fantasy', 'Film-Noir', 'History', 'Horror', 'Music',
'Musical', 'Mystery', 'Romance', 'Sci-Fi', 'Sport', 'Thriller', 'War',
'Western' ]]
# # Section 4: Data Modeling
#
# We are not applying any machine learning techniques so we will skip this section
# # Section 5: Evaluating results
# ### Section 5.1 Evlauting results for Question 1
# Question: First question - Do certain genres make more $ than others?
#
# We will make use of seaborn to generate a heatmap of correlations between various genres and the worldwide gross income
#
fig, ax = plt.subplots(figsize=(15,15)) #instantiate the plot
income_corr = imdb_genres_df.corr() #calculate correlation
mask = np.zeros_like(income_corr[['worlwide_gross_income']], dtype=np.bool) #masking all the 1 values since they don't add any value
mask[np.triu_indices_from(mask)] = True #masking list
sns.heatmap(income_corr[['worlwide_gross_income']].sort_values(by=['worlwide_gross_income'],ascending=False), annot=True, fmt=".2f",linewidths=.5, ax=ax, vmin=-1, square=True, mask = mask, cmap='coolwarm');
# ###### Sci-fi, animation, and fantasy are clear winners. Drama and romance have a negative correlation implying that these genres lead to unimpressive returns.
# ### Section 5.2 Evlauting results for Question 2
#
# Question 2 - Does the amount of funding for a movie have an impact on commercial success?
# We will generate a scatter plot of the budget vs worldwide_gross_income to assess the distribution
#
imdb_df.plot.scatter(x='budget',
y='worlwide_gross_income',
c='DarkBlue', figsize=(20,10),style='plain') #simple scatter plot generation function that defines the axes and size of plot
# Based on the above plot, we can somewhat infer that budget and gross income are correlated. Let's see if we can draw a regression line to fit the plot
sns.lmplot(x='budget',y='worlwide_gross_income',data=imdb_df,fit_reg=True,line_kws={'color': 'red'},height=8, aspect=2)
#Clearly, there is correlation. Let's calculate the Pearson correltion between the two columns.
imdb_df['budget'].corr(imdb_df['worlwide_gross_income'])
# ##### Conclusion: Budget and worldwide gross income are highly correlated
# ### Section 5.3 Evlauting results for Question 3
#
# Question 2 - What impact does duration have on the success of a movie?
#
#Let's get an idea what the duration distribution looks like
imdb_df['duration'].describe()
# +
#We will generate a scatter plot of the duration vs worldwide_gross_income to assess the distribution
imdb_df.plot.scatter(x='duration',
y='worlwide_gross_income',
c='DarkBlue', figsize=(20,10))
# -
#The average length of a movie in our database is 105 minutes. Anything less or more tends to taper off the commercial value.
#We will seperate the duration into buckets
imdb_df['duration_binned'] = pd.cut(imdb_df['duration'], [0,30,60,90,120,150,180,210,240,270,300])
#We will then generate a bar chart distribution
imdb_df.groupby('duration_binned')['worlwide_gross_income'].mean().plot.bar()
#It seems the ideal movie falls within the bucket of 180 minutes to 210 minutes.
imdb_df.groupby('duration_binned')['worlwide_gross_income'].count().plot.bar()
#Turns out that the average commercial value in the 180-210 bucket which is high seems to be driven by a small number of highly successful movies.
imdb_df.groupby('duration')['worlwide_gross_income'].mean().idxmax() #For context, let's understand which movie is the source of greatest success
# ###### Conclusion: For production studios, perhaps this means pushing to fall within this bucket if they’ve got other variables right. But if they want to play it safe, falling within the 120 to 150 minutes will be a safer bet.
# ### Section 5.4 Evlauting results for Question 4
#
# Question 4 - Do viewer ratings have an impact on how well a movie will do in the Box Office?
#
# +
#We will generate a scatter plot of the avg_vote vs worldwide_gross_income to assess the distribution
imdb_df.plot.scatter(x='avg_vote',
y='worlwide_gross_income',
c='DarkBlue',figsize=(20,10))
# -
#We will draw a regression line to see if there's some correlation
sns.lmplot(x='avg_vote',y='worlwide_gross_income',data=imdb_df,fit_reg=True,line_kws={'color': 'red'},height=8, aspect=2)
#Shocking! Let's calculate the Pearson correltion between the two columns.
imdb_df['avg_vote'].corr(imdb_df['worlwide_gross_income'])
# ###### Conclusion: The average user rating has very little to no impact on worldwide gross income. In fact, the correlation between the two is just 13%! This is yet another incentive for movie studios to continue doing what they do best, and care little about ratings they receive from the audience.
# ### Section 5.5 Evaluating results for Question 5
#
# Question 5 - Does the number of votes impact the success of a movie?
#
# Similar to my approach in Question 4, I will perform an analysis and derive the correlation between the numbers of votes and the corresponding worldwide gross income.
#
sns.lmplot(x='votes',y='worlwide_gross_income',data=imdb_df,fit_reg=True,line_kws={'color': 'red'},height=8, aspect=2)
#Clearly, there is correlation. Let's calculate the Pearson correltion between the two columns.
imdb_df['votes'].corr(imdb_df['worlwide_gross_income'])
# ###### Conclusion: It seems the number of votes is a great proxy for the popularity of a movie, which corresponds to commercial success. Perhaps, the age-old proverb that “all publicity is good publicity” holds especially true for movies.
# # Conclusion summary
# 1. We discovered that certain movie genres generate greater success than others.
# 2. The budget and number of user ratings have a strong positive impact on the likelihood of success of a movie.
# 3. Movie ratings have very little impact on the success of a movie.
| analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import os
# %matplotlib inline
# -
cwd = os.getcwd()
data_path = os.path.join(os.path.dirname(cwd), 'output', 'summary_block_group_id.csv')
df = pd.read_csv(data_path)
# # Zero Workers
field = 'workers_0'
df.plot(kind='scatter', x=field+'_control', y=field+'_result')
# # 1 Worker
field = 'workers_1'
df.plot(kind='scatter', x=field+'_control', y=field+'_result')
# # 2 Workers
field = 'workers_2'
df.plot(kind='scatter', x=field+'_control', y=field+'_result')
# # 3 Or More Workers
# 3 Workers
field = 'workers_3_plus'
df.plot(kind='scatter', x=field+'_control', y=field+'_result')
| psrc/notebooks/hh_workers-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.9 64-bit
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import glob
import functools
import matplotlib.pyplot as plt
import plotly.express as px
import warnings
import seaborn as sns
# %matplotlib inline
# %precision 4
warnings.filterwarnings('ignore')
plt.style.use('seaborn')
np.set_printoptions(suppress=True)
pd.set_option("display.precision", 15)
def sentiment_per_day(dataframe, fromDate, toDate):
dataframe = dataframe[(dataframe['created_at']>=fromDate) & (dataframe['created_at']<=toDate)]
daily_sentiment = dataframe.groupby([dataframe['date'].dt.date]).agg({'positive':'sum', 'negative':'sum','neutral':'sum'})
# daily_sentiment = dataframe.groupby([dataframe['date'].dt.date]).agg({'positive':'sum', 'negative':'sum'})
tweets_per_day = (dataframe.groupby([dataframe['date'].dt.date])['tweet'].count()).to_frame('tweets_per_day')
average_sentiments_per_day = tweets_per_day.merge(daily_sentiment, how='inner', on='date')
average_sentiments_per_day['positive'] = np.round(average_sentiments_per_day['positive']/average_sentiments_per_day['tweets_per_day'],6)
average_sentiments_per_day['negative'] = np.round(average_sentiments_per_day['negative']/average_sentiments_per_day['tweets_per_day'],6)
average_sentiments_per_day['neutral'] = np.round(average_sentiments_per_day['neutral']/average_sentiments_per_day['tweets_per_day'],6)
return average_sentiments_per_day
pre_covid_from = '2017-01-01T00:00:00'
pre_covid_to = '2020-02-26 23:59:59'
during_covid_from = '2020-02-27 00:00:00'
during_covid_to = '2021-09-30T23:59:59'
# +
user_folder_path = '../../data/twitter/'
pharma_df = pd.concat([pd.read_csv(f, sep=',') for f in glob.glob(user_folder_path + "/pharma companies/*.csv")],ignore_index=True)
# pharma_df = pd.read_csv(user_folder_path+'pharma companies/biogen.csv')
pharma_df['date'] = pd.to_datetime(pharma_df['created_at'])
# -
pre_pharma_companies_average_sentiments_per_day = sentiment_per_day(pharma_df, pre_covid_from, pre_covid_to)
fig = px.line(pre_pharma_companies_average_sentiments_per_day, x=pre_pharma_companies_average_sentiments_per_day.index, y=['positive','negative','neutral'],title='Average Change in Sentiments-Pharma Companies',
labels={'variable':'sentiment', 'date':'Date','value':'Average sentiment per day'})
# fig = px.line(pre_pharma_companies_average_sentiments_per_day, x=pre_pharma_companies_average_sentiments_per_day.index, y=['positive','negative'],title='Average Change in Sentiments-Pharma Companies',
# labels={'variable':'sentiment', 'date':'Date','value':'Average sentiment per day'})
fig.show()
pre_pharma_companies_average_sentiments_per_day
| code/sentiment-analysis/daily-sentiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Refer to https://tensorflow.google.cn/beta/tutorials/generative/dcgan
from __future__ import absolute_import, division, print_function
import tensorflow as tf
import glob
import imageio
import matplotlib.pyplot as plt
import numpy as np
import os
import PIL
import tensorflow.keras.layers as layers
import time
from IPython import display
# ### Load dataset
(train_images, train_labels), (_, _) = tf.keras.datasets.mnist.load_data()
train_images = train_images.reshape(train_images.shape[0], 28, 28, 1).astype('float32')
train_images = (train_images - 127.5) / 127.5 # Normalize the images to [-1, 1]
BUFFER_SIZE = 60000
BATCH_SIZE = 256
# Batch and shuffle the data
train_dataset = tf.data.Dataset.from_tensor_slices(train_images).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
# ### Define generator
# The generator uses `tf.keras.layers.Conv2DTranspose` (upsampling) layers to produce an image from a seed (random noise). Start with a Dense layer that takes this seed as input, then upsample several times until you reach the desired image size of 28x28x1. Notice the `tf.keras.layers.LeakyReLU` activation for each layer, except the output layer which uses tanh.
# +
def make_generator_model():
model = tf.keras.Sequential()
model.add(layers.Dense(7*7*256, use_bias=False, input_shape=(100,)))
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Reshape((7, 7, 256)))
assert model.output_shape == (None, 7, 7, 256) # Note: None is the batch size
model.add(layers.Conv2DTranspose(128, (5, 5), strides=(1, 1), padding='same', use_bias=False))
assert model.output_shape == (None, 7, 7, 128)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(64, (5, 5), strides=(2, 2), padding='same', use_bias=False))
assert model.output_shape == (None, 14, 14, 64)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(1, (5, 5), strides=(2, 2), padding='same', use_bias=False, activation='tanh'))
assert model.output_shape == (None, 28, 28, 1)
return model
# -
# ### Define discriminator
# The discriminator is a CNN-based image classifier.
def make_discriminator_model():
model = tf.keras.Sequential()
model.add(layers.Conv2D(64, (5, 5), strides=(2, 2), padding='same',
input_shape=[28, 28, 1]))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(128, (5, 5), strides=(2, 2), padding='same'))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Flatten())
model.add(layers.Dense(1))
return model
# ### Init model
# The model will be trained to output positive values for real images, and negative values for fake images.
# +
generator = make_generator_model()
noise = tf.random.normal([1, 100])
generated_image = generator(noise, training=False)
plt.imshow(generated_image[0, :, :, 0], cmap='gray')
discriminator = make_discriminator_model()
decision = discriminator(generated_image)
print (decision)
# -
# ### Define loss & optimizers
# Define loss functions and optimizers for both models.
# This method returns a helper function to compute cross entropy loss
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
# #### Discriminator loss
# This method quantifies how well the discriminator is able to distinguish real images from fakes. It compares the discriminator's predictions on real images to an array of 1s, and the discriminator's predictions on fake (generated) images to an array of 0s.
def discriminator_loss(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output), fake_output)
total_loss = real_loss + fake_loss
return total_loss
# #### Generator loss
# The generator's loss quantifies how well it was able to trick the discriminator. Intuitively, if the generator is performing well, the discriminator will classify the fake images as real (or 1). Here, we will compare the discriminators decisions on the generated images to an array of 1s.
def generator_loss(fake_output):
return cross_entropy(tf.ones_like(fake_output), fake_output)
# #### optimizers
# The discriminator and the generator optimizers are different since we will train two networks separately.
generator_optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_optimizer = tf.keras.optimizers.Adam(1e-4)
# ### Save checkpoints
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
# ### Define the training loop
# +
EPOCHS = 10
noise_dim = 100
num_examples_to_generate = 16
# We will reuse this seed overtime (so it's easier)
# to visualize progress in the animated GIF)
seed = tf.random.normal([num_examples_to_generate, noise_dim])
# -
# The training loop begins with generator receiving a random seed as input. That seed is used to produce an image. The discriminator is then used to classify real images (drawn from the training set) and fakes images (produced by the generator). The loss is calculated for each of these models, and the gradients are used to update the generator and discriminator.
# Notice the use of `tf.function`
# This annotation causes the function to be "compiled".
@tf.function
def train_step(images):
noise = tf.random.normal([BATCH_SIZE, noise_dim])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(noise, training=True)
real_output = discriminator(images, training=True)
fake_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
# +
def train(dataset, epochs):
for epoch in range(epochs):
start = time.time()
for image_batch in dataset:
train_step(image_batch)
# Produce images for the GIF as we go
display.clear_output(wait=True)
generate_and_save_images(generator,
epoch + 1,
seed)
# Save the model every 15 epochs
if (epoch + 1) % 15 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time for epoch {} is {} sec'.format(epoch + 1, time.time()-start))
# Generate after the final epoch
display.clear_output(wait=True)
generate_and_save_images(generator,
epochs,
seed)
def generate_and_save_images(model, epoch, test_input):
# Notice `training` is set to False.
# This is so all layers run in inference mode (batchnorm).
predictions = model(test_input, training=False)
fig = plt.figure(figsize=(4,4))
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, 0] * 127.5 + 127.5, cmap='gray')
plt.axis('off')
plt.savefig('image_at_epoch_{:04d}.png'.format(epoch))
plt.show()
# -
# ### Train the model
# Call the train() method defined above to train the generator and discriminator simultaneously. Note, training GANs can be tricky. It's important that the generator and discriminator do not overpower each other (e.g., that they train at a similar rate).
#
# At the beginning of the training, the generated images look like random noise. As training progresses, the generated digits will look increasingly real. After about 50 epochs, they resemble MNIST digits.
# %%time
train(train_dataset, EPOCHS)
# Restore the latest checkpoint.
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
# ### Create a GIF
# Use imageio to create an animated gif using the images saved during training.
# +
anim_file = 'dcgan.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('image*.png')
filenames = sorted(filenames)
last = -1
for i,filename in enumerate(filenames):
frame = 2*(i**0.5)
if round(frame) > round(last):
last = frame
else:
continue
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
# -
# A hack to display the GIF inside this notebook
os.rename('dcgan.gif', 'dcgan.gif.png')
display.Image(filename="dcgan.gif.png")
| TensorFlow2_DCGAN_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from copy import deepcopy
import numpy as np
import pandas as pd
# %matplotlib notebook
import matplotlib.pyplot as plt
# a,bは0.5~2m、dは0.6mくらい、rは15mmくらい
# const [mm]
r = 15.725
d = 500.0
a = 2038.37
b = 2155.12
def terminal_trajectory(th, r, l, d):
"""
th: angle [rad]
r: pulley raidus [m]
l: wire length [m]
d: x-axis offset [m]
"""
x = (l - r * th) * np.cos(th) + r * np.sin(th) - d
y = (l - r * th) * np.sin(th) - r * np.cos(th)
return np.array([x, y])
df = pd.DataFrame()
# theta [rad]
dt = np.pi / (180)
df['theta'] = np.arange(0, 2*np.pi + dt, dt)
# x1, y1
f1 = terminal_trajectory(df.theta, r, a, 0)
df['x1'] = f1[0]
df['y1'] = f1[1]
# x2, y2
f2 = terminal_trajectory(df.theta, r, b, d)
df['x2'] = f2[0]
df['y2'] = f2[1]
# check
fig = plt.figure()
ax1 = fig.add_subplot(111)
ax1.set_aspect('equal')
ax1.scatter(df.x1, df.y1, s=1, c='red', label='Encoder 1')
ax1.scatter(df.x2, df.y2, s=1, c='blue', label='Encoder 2')
ax1.set_xlabel('x [mm]')
ax1.set_ylabel('y [mm]')
comment = "a = %s [mm]\nb = %s [mm]\nd = %s [mm]\nr = %s [mm]" % (a, b, d, r)
ax1.text(0, 0, comment, size=9, c="black")
ax1.legend(bbox_to_anchor=(1, 1), loc='lower right', borderaxespad=1, fontsize=9)
plt.show()
df
def fx_fy(theta_phi):
(th, ph) = theta_phi
(x1, y1) = terminal_trajectory(th, r, a, 0)
(x2, y2) = terminal_trajectory(ph, r, b, d)
fx = x1 - x2
fy = y1 - y2
return np.array([fx, fy])
def jacobian(theta_phi):
(th, ph) = theta_phi
a_ = a - r * th
b_ = b - r * ph
J11 = -a_ * np.sin(th)
J12 = b_ * np.sin(ph)
J21 = a_ * np.cos(th)
J22 = -b_ * np.cos(ph)
return np.array([[J11, J12], [J21, J22]])
# newton method
theta_phi = np.array([0.25*np.pi, 0.3*np.pi])
delta = np.array([1e3, 1e3])
i = 0
solutions = []
while (abs(delta[0]) > 1e-4 or abs(delta[1]) > 1e-4):
try:
J = jacobian(theta_phi)
delta = np.dot(np.linalg.inv(J), fx_fy(theta_phi))
theta_phi = theta_phi - delta
except:
theta_phi = np.array([np.nan, np.nan])
p1 = terminal_trajectory(theta_phi[0], r, a, 0)
solutions.append(p1)
print(i, delta, theta_phi, p1)
i = i + 1
if (i > 30):
print('not convergence')
break
# add solutions to plot
px=np.array(points).transpose()[0]
py=np.array(points).transpose()[1]
ax1.scatter(px, py, s=50, marker='+', c='limegreen', label='solution')
for i in range(len(points)):
ax1.text(px[i], py[i]-10, str(i), size=9, c='limegreen')
fig.savefig("a=%s[mm]b=%s[mm]d=%s[mm]r=%s[mm].png" % (a, b, d, r))
| solve-two-nonlinear-equations-with-newton-method.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tfs
# language: python
# name: tfs
# ---
# # Autoencoder: Denoising Images
# +
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ##### preprocess the data
# +
mnist = tf.keras.datasets.fashion_mnist
(x_train, _), (x_test, _) = mnist.load_data()
# reshape for input convultional layer
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1)
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
# -
x_test.shape
# ###### introduce random noise to the data
# +
noise = 50
mean, stddev = 0., 1.
x_train_noisy = x_train + noise * np.random.normal(loc=mean, scale=stddev, size=x_train.shape)
x_test_noisy = x_test + noise * np.random.normal(loc=mean, scale=stddev, size=x_test.shape)
# Clip to range [0 -1]
# x_train_noisy = np.clip(x_train_noisy, 0., 1.)
# x_test_noisy = np.clip(x_test_noisy, 0., 1.)
# +
# display the noisy images
fig=plt.figure(figsize=(11, 11))
columns = 6
rows = 2
w, h = 28, 28
for i in range(1, columns*rows +1):
img = np.reshape(x_train_noisy[i], [w, h])
ax = fig.add_subplot(rows, columns, i)
plt.imshow(img, cmap='Greys')
plt.tight_layout()
plt.show()
# -
# #### Create the model
# ##### - encoding layers
# +
input_img = tf.keras.layers.Input(shape=(28, 28, 1))
conv_1 = tf.keras.layers.Conv2D(filters=32, kernel_size=3,
activation='relu',
padding='same')(input_img) # [None, 28, 28, 32]
max_pool1 = tf.keras.layers.MaxPool2D(pool_size=(2, 2), padding='same')(conv_1) # [None, 14, 14, 32]
conv_2 = tf.keras.layers.Conv2D(filters=32, kernel_size=2,
activation='relu',
padding='same')(max_pool1) # [None, 14, 14, 32]
encoded = tf.keras.layers.MaxPool2D((2, 2), padding='same')(conv_2) # [None, 7, 7, 32]
# -
# ##### - decoding layers
# - Decoding reverses the encoding process.
#
#
# We use an UpSampling2D layer to `undo` the effect of MaxPooling
# +
conv_3 = tf.keras.layers.Conv2D(filters=32, kernel_size=3, activation='relu',
padding='same')(encoded) # [None, 7, 7, 32]
up_sampl1 = tf.keras.layers.UpSampling2D(size=[2, 2])(conv_3) # [None, 14, 14, 32]
conv_4 = tf.keras.layers.Conv2D(filters=32, kernel_size=3, activation='relu',
padding='same')(up_sampl1) # [None, 14, 14, 32]
up_sampl2 = tf.keras.layers.UpSampling2D(size=[2, 2])(conv_4) # [None, 28, 28, 32]
decoded = tf.keras.layers.Conv2D(filters=1, kernel_size=3, activation='sigmoid', padding='same')(up_sampl2)
# -
decoded.shape
# ##### train
# +
log_dir = './tensorboard'
epochs, batch_size = 100, 128
autoencoder = tf.keras.models.Model(inputs=input_img, outputs=decoded)
autoencoder.compile(optimizer='adam', loss='binary_crossentropy')
autoencoder.fit(x_train_noisy, x_train, epochs=epochs, batch_size=batch_size,
validation_data=(x_test_noisy, x_test),
callbacks=[tf.keras.callbacks.TensorBoard(log_dir)]
)
# -
decoded_imgs = autoencoder.predict(x_test_noisy)
# ##### display the denoised output
# +
fig=plt.figure(figsize=(9, 9))
columns = 6
rows = 2
w, h = 28, 28
for i in range(1, columns*rows +1):
img = np.reshape(decoded_imgs[i], [w, h])
ax = fig.add_subplot(rows, columns, i)
plt.imshow(img, cmap='Greys')
plt.tight_layout()
plt.show()
| notebooks/2.0 supervised_and_unsupervised_learning/5.2 autoencoder_denoising.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
import pickle
df_train = pd.read_csv("train.csv")
df_train = df_train.sample(frac=1).reset_index(drop=True)
df_train_cat0 = df_train.loc[ df_train['target'] == 0 ]
df_train_cat1 = df_train.loc[ df_train['target'] == 1 ]
len(df_train_cat1)
len(df_train_cat0)
len(df_train_cat1)
df_train = df_train_cat0[:40000].append(df_train_cat1 )
len(df_train)
df_train = df_train.sample(frac=1).reset_index(drop=True)
df_train_cat0[:3]
Y_labels = df_train['target']
df_train = df_train.drop("id", 1)
df_train = df_train.drop("target", 1)
all_cols = df_train.columns
len(all_cols)
# +
categoical_vars = [ ]
continous_vars = [ ]
categoical_binary_vars = [ ]
for col in all_cols:
#if "bin" in col :
# categoical_binary_vars.append(col)
if "cat" in col :
categoical_vars.append(col)
else:
continous_vars.append(col )
# -
print ("categorical binary vars: ", len(categoical_binary_vars))
print ("categorical non binary vars: ", len(categoical_vars))
print ("continues vars: ", len(continous_vars))
for cat_var in categoical_vars:
print (cat_var, df_train[cat_var].nunique())
len(df_train)
df_tr = df_train[:40000]
df_test = df_train[40000:]
Y_train = Y_labels[:40000]
Y_test = Y_labels[40000: ]
df_tr[:3]
# ## NOW LETS WORK WITH MODEL BUILDING
# ---
other_cols = [i for i in all_cols if i not in categoical_vars ]
# +
#understanding th eembedding size thing:
#say there is a string ["I am a man"]
#coresponding input will be ["12 56 89 74"]
#Now for each word will converted to some vector of lenght embedding size (say 2 here )
#coresponding output will be [[0.2 0.3], [0.12, 0.25], [78, 45]]
#And for multiple ips, multiple ops
# the model will take as input an integer matrix of size (batch, input_length).
# the largest integer (i.e. word index) in the input should be no larger than 999 (vocabulary size).
# now model.output_shape == (None, 10, 64), where None is the batch dimension.
# now model.output_shape == (None, 10, 64), where None is the batch dimension, 10 is embedding size and 64 is input length
# -
def preproc(X_train ) :
input_list_train = []
#the cols to be embedded: rescaling to range [0, # values)
for c in categoical_vars :
raw_vals = np.unique(X_train[c])
val_map = {}
for i in range(len(raw_vals)):
val_map[raw_vals[i]] = i
input_list_train.append(X_train[c].map(val_map).values)
#the rest of the columns
input_list_train.append(X_train[other_cols].values)
return input_list_train
df_tr[other_cols].ndim
df_tr = preproc( df_tr )
df_test = preproc( df_test )
len(df_tr)
len(df_tr[14] )
len(categoical_vars)
len(df_tr[13])
df_tr[14].ndim
len (df_tr[14][0] )
# +
from keras.layers import *
from keras.models import *
models = []
for categoical_var in categoical_vars :
model = Sequential()
no_of_unique_cat = df_train[categoical_var].nunique()
embedding_size = min(np.ceil((no_of_unique_cat)/2), 50 )
embedding_size = int(embedding_size)
model.add( Embedding( no_of_unique_cat+1, embedding_size, input_length = 1 ) )
model.add(Reshape(target_shape=(embedding_size,)))
models.append( model )
model_rest = Sequential()
model_rest.add(Dense(16, input_dim= 43 ))
models.append(model_rest)
full_model = Sequential()
full_model.add(Merge(models, mode='concat'))
full_model.add(Dense(1000))
full_model.add(Activation('relu'))
full_model.add(Dense(400))
full_model.add(Activation('relu'))
full_model.add(Dense(200))
full_model.add(Activation('sigmoid'))
full_model.add(Dense(2))
full_model.add(Activation('sigmoid'))
full_model.compile(loss='binary_crossentropy', optimizer='adam',metrics=['accuracy'])
# -
len(models)
full_model.summary()
from keras.utils.np_utils import to_categorical
Y_test_cat = to_categorical(Y_test.tolist() )
Y_train_cat = to_categorical(Y_train.tolist() )
history = full_model.fit( df_tr, Y_train_cat , validation_data= (df_test, Y_test_cat ), epochs= 10 )
history.params
# +
loss = history.history['loss']
acc = history.history['acc']
import matplotlib.pyplot as plt
# %matplotlib inline
plt.xkcd()
plt.xlabel("Epochs")
plt.ylabel("Loss")
plt.plot( loss )
# -
plt.xlabel("epoch")
plt.ylabel("train_acc")
plt.plot( acc )
| entity_embedding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Annotation Tutorial
#
# **NB**: please refer to the scVI-dev notebook for introduction of the scVI package.
#
# In this notebook, we investigate how semi-supervised learning combined with the probabilistic modelling of latent variables in scVI can help address the annotation problem.
#
# The annotation problem consists in labelling cells, ie. **inferring their cell types**, knowing only a part of the labels.
# cd ../..
# %matplotlib inline
# #### Loading Config
save_path = 'data/'
n_epochs_all = None
import os
import numpy as np
import torch
import matplotlib.pyplot as plt
from scvi.dataset import CortexDataset, LoomDataset
from scvi.models import SCANVI, VAE
from scvi.inference import UnsupervisedTrainer, JointSemiSupervisedTrainer, SemiSupervisedTrainer
# ## Annotating one dataset from another
#
#
# #### Synthetic data
#
# We perform the annotation task on synthetically generated data:
if not os.path.exists(os.path.join(save_path, 'simulation/')):
os.makedirs(os.path.join(save_path, 'simulation/'))
# +
simulation_1 = LoomDataset(filename='simulation_1.loom',
save_path=os.path.join(save_path, 'simulation/'),
url='https://github.com/YosefLab/scVI-data/raw/master/simulation/simulation_1.loom')
scanvi = SCANVI(simulation_1.nb_genes, simulation_1.n_batches, simulation_1.n_labels)
trainer_scanvi = SemiSupervisedTrainer(scanvi, simulation_1, frequency=5)
# -
# #### Introductory example
#
# In the semi-supervised setting, we first need to define the labelled and unlabelled set. These are each represented in as the `Posterior` attributes of `trainer` object, called `labelled_set` and `unlabelled_set`.
#
# To override these attributes, the method `.create_posterior()` of the trainer might take the indices of interest.
#
# In the scenario of annotating one dataset from another, we can set those indices according to the biological batch index, which determines the origin of the dataset. In the example below, we try to predict cell-types of batch 1 given cell-types of batch 0.
# +
n_epochs = 200 if n_epochs_all is None else n_epochs_all
trainer_scanvi.labelled_set = trainer_scanvi.create_posterior(indices=(simulation_1.batch_indices == 0).ravel())
trainer_scanvi.labelled_set.to_monitor = ['reconstruction_error', 'accuracy']
trainer_scanvi.unlabelled_set = trainer_scanvi.create_posterior(indices=(simulation_1.batch_indices == 1).ravel())
trainer_scanvi.unlabelled_set.to_monitor = ['reconstruction_error', 'accuracy']
trainer_scanvi.train(n_epochs)
# -
# We plot the evolution of the accuracy throughout the training.
accuracy_labelled_set = trainer_scanvi.history["accuracy_labelled_set"]
accuracy_unlabelled_set = trainer_scanvi.history["accuracy_unlabelled_set"]
x = np.linspace(0,n_epochs,(len(accuracy_labelled_set)))
plt.plot(x, accuracy_labelled_set, label="accuracy labelled")
plt.plot(x, accuracy_unlabelled_set, label="accuracy unlabelled")
# We can evaluate the final accuracy of SCANVI's classifier:
print("Labelled set accuracy : ", trainer_scanvi.labelled_set.accuracy())
print("Unlabelled set accuracy : ", trainer_scanvi.unlabelled_set.accuracy())
# As well as the Nearest Neighbors predictor accuracy in SCANVI's latent space:
print("NN accuracy in latent space : ", trainer_scanvi.labelled_set.nn_latentspace(trainer_scanvi.unlabelled_set))
# To get the predictions, `.compute_predictions()` returns the original (ground truth) and predicted labels. In the case where the second dataset actually came with no cell-type labelling at all, the first argument is just an array of "dummy" labels.
_, y_pred = trainer_scanvi.unlabelled_set.compute_predictions()
# The tsne for both datasets can be visualized with the `.full_dataset` Posterior attribute of the `trainer`.
n_samples_tsne = 1000
trainer_scanvi.full_dataset.show_t_sne(n_samples=n_samples_tsne, color_by='batches and labels')
trainer_scanvi.full_dataset.entropy_batch_mixing()
n_samples = 100
M_permutation = 10000
bayes_factors_list = trainer_scanvi.full_dataset.differential_expression_score(
simulation_1.labels.ravel()==0 , simulation_1.labels.ravel() == 4,
n_samples=n_samples, M_permutation=M_permutation
)
# ### Going further
#
# For benchmarking, let's wrap it all into a function. We might use VAE model before for pretraining.
# +
n_epochs_vae = 150 if n_epochs_all is None else n_epochs_all
n_epochs_scanvi = 50 if n_epochs_all is None else n_epochs_all
n_samples_tsne = 1000
def scanvi_both_ways(dataset):
print("Unsupervised training: warm-up phase using only the VAE.")
vae = VAE(dataset.nb_genes, dataset.n_batches, dataset.n_labels)
trainer = UnsupervisedTrainer(vae, dataset, train_size=1.0)
trainer.train(n_epochs_vae)
print("Entropy batch mixing : ", trainer.train_set.entropy_batch_mixing())
trainer.train_set.show_t_sne(n_samples=n_samples_tsne, color_by='batches and labels')
for i in [0,1]:
print("\nUsing batch %d as labelled set"%i)
scanvi = SCANVI(dataset.nb_genes, dataset.n_batches, dataset.n_labels,
classifier_parameters = {'dropout_rate':0.2, 'n_hidden':256, 'n_layers':2})
scanvi.load_state_dict(vae.state_dict(), strict=False)
trainer_scanvi = SemiSupervisedTrainer(scanvi, dataset)
trainer_scanvi.labelled_set = trainer_scanvi.create_posterior(indices=(dataset.batch_indices==i).ravel())
trainer_scanvi.unlabelled_set = trainer_scanvi.create_posterior(indices=(dataset.batch_indices==1-i).ravel())
print("VAE + KNN classifier: ", trainer_scanvi.labelled_set.nn_latentspace(trainer_scanvi.unlabelled_set))
trainer_scanvi.train(n_epochs_scanvi, lr=1e-4)
accuracy_score = trainer_scanvi.unlabelled_set.accuracy()
print("Unlabelled set accuracy : ", trainer_scanvi.unlabelled_set.accuracy())
# -
scanvi_both_ways(simulation_1)
# #### UMI / Non UMI simulation
#
# In this simulation, batch 0 is UMI counts whereas batch 1 is non UMI.
# +
simulation_2 = LoomDataset(filename='simulation_2.loom',
save_path=os.path.join(save_path, 'simulation/'),
url='https://github.com/YosefLab/scVI-data/raw/master/simulation/simulation_2.loom')
scanvi_both_ways(simulation_2)
# -
# #### EVF / not EVF
# +
simulation_3 = LoomDataset(filename='simulation_3.loom',
save_path=os.path.join(save_path, 'simulation/'),
url='https://github.com/YosefLab/scVI-data/raw/master/simulation/simulation_3.loom')
scanvi_both_ways(simulation_3)
# -
# We instantiate the `SCANVI` model and train it over 250 epochs. Only labels from the `data_loader_labelled` will be used, but to cross validate the results, the labels of `data_loader_unlabelled` will is used at test time. The accuracy of the `unlabelled` dataset reaches 93% here at the end of training.
#
# ## Annotating a dataset with few labels
#
# SCANVI also performs well in the context of annotating the rest of a dataset.
# +
gene_dataset = CortexDataset(save_path=save_path)
use_batches=False
use_cuda=True
n_epochs = 100 if n_epochs_all is None else n_epochs_all
n_cl = 10
scanvi = SCANVI(gene_dataset.nb_genes, gene_dataset.n_batches, gene_dataset.n_labels)
trainer = JointSemiSupervisedTrainer(scanvi, gene_dataset,
n_labelled_samples_per_class=n_cl,
classification_ratio=100)
trainer.train(n_epochs=n_epochs)
trainer.unlabelled_set.accuracy()
# -
# **Benchmarking against other algorithms**
#
# We can compare ourselves against the random forest and SVM algorithms, where we do grid search with 3-fold cross validation to find the best hyperparameters of these algorithms. This is automatically performed through the functions **`compute_accuracy_svc`** and **`compute_accuracy_rf`**.
#
# These functions should be given as input the numpy array corresponding to the equivalent dataloaders, which is the purpose of the **`get_raw_data`** method from `scvi.dataset.utils`.
#
# The format of the result is an Accuracy named tuple object giving higher granularity information about the accuracy ie, with attributes:
#
# - **unweighted**: the standard definition of accuracy
#
# - **weighted**: we might give the same weight to all classes in the final accuracy results. Informative only if the dataset is unbalanced.
#
# - **worst**: the worst accuracy score for the classes
#
# - **accuracy_classes** : give the detail of the accuracy per classes
#
#
# Compute the accuracy score for rf and svc
# +
from scvi.inference.annotation import compute_accuracy_rf, compute_accuracy_svc
data_train, labels_train = trainer.labelled_set.raw_data()
data_test, labels_test = trainer.unlabelled_set.raw_data()
svc_scores = compute_accuracy_svc(data_train, labels_train, data_test, labels_test)
rf_scores = compute_accuracy_rf(data_train, labels_train, data_test, labels_test)
print("\nSVC score test :\n", svc_scores[0][1])
print("\nRF score train :\n", rf_scores[0][1])
# -
def allow_notebook_for_test():
print("Testing the annotation notebook")
| tests/notebooks/annotation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NLP Information Extraction
# ---
# ## SparkContext and SparkSession
# +
from pyspark import SparkContext
sc = SparkContext(master = 'local')
from pyspark.sql import SparkSession
spark = SparkSession.builder \
.appName("Python Spark SQL basic example") \
.config("spark.some.config.option", "some-value") \
.getOrCreate()
# -
# ## Simple NLP pipeline architecture
#
# 
# **Reference:** Bird, Steven, <NAME>, and <NAME>. Natural language processing with Python: analyzing text with the natural language toolkit. " O'Reilly Media, Inc.", 2009.
#
# ## Example data
#
# The raw text is from the gutenberg corpus from the nltk package. The fileid is *milton-paradise.txt*.
#
# ### Get the data
#
# #### Raw text
# +
import nltk
from nltk.corpus import gutenberg
milton_paradise = gutenberg.raw('milton-paradise.txt')
# -
# ## Create a spark data frame to store raw text
#
# * Use the `nltk.sent_tokenize()` function to split text into sentences.
import pandas as pd
pdf = pd.DataFrame({
'sentences': nltk.sent_tokenize(milton_paradise)
})
df = spark.createDataFrame(pdf)
df.show(n=5)
# ## Tokenization and POS tagging
# +
from pyspark.sql.functions import udf
from pyspark.sql.types import *
## define udf function
def sent_to_tag_words(sent):
wordlist = nltk.word_tokenize(sent)
tagged_words = nltk.pos_tag(wordlist)
return(tagged_words)
## define schema for returned result from the udf function
## the returned result is a list of tuples.
schema = ArrayType(StructType([
StructField('f1', StringType()),
StructField('f2', StringType())
]))
## the udf function
sent_to_tag_words_udf = udf(sent_to_tag_words, schema)
# -
# #### Transform data
df_tagged_words = df.select(sent_to_tag_words_udf(df.sentences).alias('tagged_words'))
df_tagged_words.show(5)
# ## Chunking
#
# Chunking is the process of segmenting and labeling multitokens. The following example shows how to do a noun phrase chunking on the tagged words data frame from the previous step.
#
# First we define a *udf* function which chunks noun phrases from a list of pos-tagged words.
# +
import nltk
from pyspark.sql.functions import udf
from pyspark.sql.types import *
# define a udf function to chunk noun phrases from pos-tagged words
grammar = "NP: {<DT>?<JJ>*<NN>}"
chunk_parser = nltk.RegexpParser(grammar)
chunk_parser_udf = udf(lambda x: str(chunk_parser.parse(x)), StringType())
# -
# #### Transform data
df_NP_chunks = df_tagged_words.select(chunk_parser_udf(df_tagged_words.tagged_words).alias('NP_chunk'))
df_NP_chunks.show(2, truncate=False)
| notebooks/07-natural-language-processing/nlp-information-extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise Session 2: Convolution, Filtering and Gradients
# The goal of this exercise is to:
# * Understand and apply a convolutional filter to an image
# * Compare the computational complexity of separable and non-separable filters
# * Compute image gradients and apply them to real-world images
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import cv2
import time
import scipy.signal as conv
plt.rcParams['figure.figsize'] = (10, 10)
plt.rcParams['image.cmap'] = 'gray'
# -
# ## Exercise 1: Applying Convolutional Filters
# Assume we are given a gray-scale image $I[x, y]$, of size $W \times H$, such that $0 \leq x \leq W-1$,
# and $0 \leq y \leq H-1$. We want to apply a filter $F[i, j]$ to image $I$. The filter $F$ is of size $(2N + 1) \times (2M + 1)$, such that $−N \leq i \leq N$, and $−M \leq j \leq M$.
#
# The result can be computed as
# \begin{align}
# R[x, y] = (I ∗ F)[x, y] = \sum_{i=-N}^{N} \sum_{i=-M}^{M} I[x − i, y − j]~F[i, j]
# \end{align}
#
# * Implement a function ```R = applyImageFilter(I,F)``` that takes a
# gray-scale image $I$ and a filter $F$ as inputs, and returns the result of the convolution
# of the two.
# Note: There are many python libraries providing convolution function to convolve an image with a filter,
# but in this exercise you are requested to implement your own. This will help your understanding
# of how convolutional filters work.
# - To avoid numerical issues, make sure $I$ and $F$ are of type float. To understand
# why, think what would happen if you add two unsigned 8-bit numbers when
# computing the convolution, for example 240 and 80.
# - What happens when computing $R[x, y]$ near the border of the image? What
# would you propose to deal with this issue?
def applyImageFilter(I, F, padding='same'):
# First input parameter: I
# Input image. It should be a 2D matrix. According to the notation in the description, it has W rows and
# H columns.
# Second input parameter: F
# Filter used for the convolution. It should be a 2D matrix. According to the notation in the decription,
# it has (2N+1) rows and (2M+1) columns.
# If image is not of type float, convert it to float
if not np.issubdtype(I.dtype,float):
I = np.float64(I)
# If the filter is not of type float, convert it to float
if not np.issubdtype(F.dtype,float):
F = np.float64(F)
# Shape of Filter
N_, M_ = F.shape
# Check whether the dimensions of input are accurate, they should be odd
if not N_%2:
raise ValueError('Number of rows in the filter must be odd')
if not M_%2:
raise ValueError('Number of columns in the filter must be odd')
# Compute the values for N,M which is used in the above description.
N = np.int64((N_-1)/2)
M = np.int64((M_-1)/2)
# Shape of the input image
W, H = I.shape
# Initialize output matrix R
# "same" padding mode: allows conv at boundary pixels, final output image size = input image size
# default padding value = 0, initialize output matrix to the original input shape
if padding == 'same':
I = np.pad(I, ((N,N),(M,M)), 'constant', constant_values=0)
R = np.zeros((W,H), dtype=np.float64)
# "valid" padding mode: no conv for boundary pixels, output image size reduced to (W-2M)*(H-2N)
elif padding == 'valid':
R = np.zeros((W-2*M, H-2*N), dtype=np.float64)
# if not any mode, raise ValueError
else:
raise ValueError("Please choose padding='same' or 'valid'.")
# Output image size
W_R, H_R = R.shape
# Perform 2D convolution
for x in range(W_R):
for y in range(H_R):
# calculate R[x,y]
# iterate over N,M to get conv value
for i in range(-N, N+1):
for j in range(-M, M+1):
R[x,y] += I[x-i+N, y-j+M] * F[i+N,j+M]
return R
# +
# Below code is to verify your implementation of the convolution
# we compare your output with the scipy.signal implementation of the convolution
# and the error should be zero
# Read the sample image
img = cv2.imread('images/res1.png',0)
# Generate a random filter F of size 3 x 3
F = np.random.randn(5,3)
# Perform convolution with the function you implemented
output_1 = applyImageFilter(img, F, padding='same')
# Perform convolution with scipy.signal implementation of convolution
output_2 = conv.convolve2d(img, F , mode='same')
# Error
print(f'Input shape: {img.shape}')
print(f'Output shape: {output_1.shape}')
print(f'Error: {np.sum(np.abs(output_1 - output_2))}')
# if the implmentation is accurate, the error will be 0
# -
# ## Exercise 2: Image Smoothing
# Convolutional filters have many uses. A very common one is to smooth or soften an
# image. A typical smoothing filter is the Gaussian Filter, which follows the equation of a
# 2D Gaussian probability distribution. It is an important preprocessing step typically to reduce the amount of noise. When constructing a Gaussian filter we can specify the size and standard deviation (σ) of the underlying Gaussian function.
# * To create a Gaussian filter use the function given below.
def gaussian_filter(fSize, fSigma):
x, y = np.mgrid[-fSize//2 + 1:fSize//2 + 1, -fSize//2 + 1:fSize//2 + 1]
g = np.exp(-((x**2 + y**2)/(2.0*fSigma**2)))
return g/g.sum()
# * What are fSize and fSigma?
# * How do fSize and fSigma affect the filter’s shape and size?
# * Visualize different filters by giving different values to these parameters.
# * If you are given fSigma, how would you choose fSize?
# +
# Here we visualize some combinations of fSize and fSigma
size_list = [3, 5, 7]
sigma_list = [1, 3, 6]
fig, axes = plt.subplots(len(size_list), len(sigma_list))
for pos in [(i,j) for i in range(len(size_list)) for j in range(len(sigma_list))]:
axes[pos].imshow(gaussian_filter(size_list[pos[0]], sigma_list[pos[1]]), cmap='viridis')
# -
# **Answer to Questions**:
# 1. fSize is the width/length of the gaussian filter, fSigma is the standard deviation for the gaussian filter/kernel.
# 2. fSize controls the size(width, height) of the filter, fSigma controls how the elements in gaussian kernel variates.
# 3. A rule of thumb of choosing the filter size: $k = ceil(3\sigma)$ for k*k kernel, k is odd
# * Apply Gaussian filtering to 'res1.png'
# * To apply the filter to an image use ```cv2.GaussianBlur(img, fSize, fSigma)```
# * Experiment with different values of fSigma. How does the amount of smoothing vary with this parameter?
# +
img = cv2.imread('images/res1.png',0)
plt.imshow(img)
# apply filters, try different values of Sigma
size_list = [3, 5, 7]
sigma_list = [1, 3, 6]
fig, axes = plt.subplots(len(size_list), len(sigma_list))
for pos in [(i,j) for i in range(len(size_list)) for j in range(len(sigma_list))]:
axes[pos].imshow(cv2.GaussianBlur(img, [size_list[pos[0]],size_list[pos[0]]], sigma_list[pos[1]]))
# -
# As you saw in the first exercise of this session, applying the filter near the border of
# the image is not a well-defined problem. It is possible to deal with this issue through
# different approaches. Luckily, cv2.GaussianBlur implements different ways of dealing with border effects.
# Here you will try them out and understand what each of them does.
# * Apply a Gaussian filter with fSigma = 10 and fSize = 25 to 'res1.png'
# * Use borderType flag of cv2.GaussianBlur function to define what kind of border should be added around the image (cv2.BORDER_CONSTANT, cv2.BORDER_REFLECT, cv2.BORDER_REPLICATE).
#
# +
img = cv2.imread('images/res1.png',0)
# apply different border types for blur filters
fig, axes = plt.subplots(1, 3)
axes[0].imshow(cv2.GaussianBlur(img, [25,25], 10, borderType=cv2.BORDER_CONSTANT))
axes[1].imshow(cv2.GaussianBlur(img, [25,25], 10, borderType=cv2.BORDER_REFLECT))
axes[2].imshow(cv2.GaussianBlur(img, [25,25], 10, borderType=cv2.BORDER_REPLICATE))
# -
# ## Exercise 3: Edge detector
# Edge detection is one of the most important operations in Computer Vision. In this exercise we will investigate how it
# can be performed and what information can be obtained from it.
#
# One of the ways to detect edges is to compute image intensity gradients. They can be approximated by convolving image with Sobel filters. They consist of 2 $3 \times 3$ filters:
#
# \begin{equation*}
# S_x = \frac{1}{8}
# \begin{bmatrix}
# -1 & 0 & +1\\
# -2 & 0 & +2\\
# -1 & 0 & +1\\
# \end{bmatrix}
# \hspace{2cm}
# S_y = \frac{1}{8}
# \begin{bmatrix}
# -1 & -2 & -1\\
# 0 & 0 & 0\\
# +1 & +2 & +1\\
# \end{bmatrix}
# \end{equation*}
#
# where $S_x$ computes the partial derivative of the image in the horizontal direction, while $S_y$ does it in the vertical direction.
# * Compute the image derivatives in x- and y-directions using Sobel filters. You can use for that function ```python cv2.Sobel()```
# +
img_coins = cv2.imread('images/coins.png',0)
### Compute gradient in x-direction
grad_x = cv2.Sobel(img_coins, ddepth=cv2.CV_64F, dx=1, dy=0, ksize=3)
### Compute gradient in y-direction
grad_y = cv2.Sobel(img_coins, ddepth=cv2.CV_64F, dx=0, dy=1, ksize=3)
# -
# * Visualize the image gradients. How can you explain the differences between the two results?
fig = plt.figure()
plt.subplot(1,2,1)
grady = plt.imshow(grad_x,cmap="jet")
plt.title("Gradient x")
plt.colorbar(fraction=0.046, pad=0.04)
plt.subplot(1,2,2)
gradx = plt.imshow(grad_y,cmap="jet")
plt.title("Gradient y")
plt.colorbar(fraction=0.046, pad=0.04)
plt.tight_layout()
# Mathematically, the goal of the filters $S_x$ and $S_y$ is to approximate the derivatives of the image with respect to
# the horizontal and vertical directions respectively, such that
# \begin{equation*}
# \nabla I_x(x,y) = (I*S_x)[x,y]
# \quad\mathrm{and}\quad
# \nabla I_y(x,y) = (I*S_y)[x,y]
# \end{equation*}
#
# Therefore, the gradient of an image at each point is a 2D vector
# \begin{equation*}
# \nabla I =
# \begin{bmatrix}
# \nabla I_x\\
# \nabla I_y
# \end{bmatrix}
# \end{equation*}
#
# This vector can be computed for every pixel. Its magnitude and phase can be computed as
# \begin{equation*}
# || \nabla I || = \sqrt{(\nabla I_x)^2 + (\nabla I_y)^2} \\
# \angle \nabla I = atan2(\nabla I_x, \nabla I_y)
# \end{equation*}
#
# $\it{Note:}$ we use atan2() instead of atan() to be able to determine the right quadrant of the phase.
# * Using the previously computed image gradients and the above formula compute the gradient magnitude and gradient phase.
#
#
# +
### Compute gradient magnitude
grad_mag = np.sqrt(np.power(grad_x, 2) + np.power(grad_y, 2))
### Compute gradient phase
grad_phase = np.arctan2(grad_x, grad_y)
# -
# * Visualize the gradient magnitude and phase images along with the original image. Where does the gradient have high
# magnitude? How does the phase change along the coin border?
fig = plt.figure()
plt.subplot(1,3,1)
plt.imshow(img_coins)
plt.subplot(1,3,2)
grady = plt.imshow(grad_mag,cmap="jet")
plt.title("Gradient magnitude")
plt.colorbar(fraction=0.046, pad=0.04)
plt.subplot(1,3,3)
gradx = plt.imshow(grad_phase)
plt.title("Gradient phase")
plt.colorbar(fraction=0.046, pad=0.04)
plt.tight_layout()
# The gradient has got the highest magnitude at the borders of the coins. For that reason it can be considered a good
# proxy for detecting edges. The phase of the gradient changes smoothly along the circular coin border along with the
# normal of the boundary.
# ## Exercise 4: Separable Filters
#
# As seen in class, certain types of 2D filters can be thought of as the composition of two
# 1-dimensional filters. These are called Separable Filters, and can be computed more
# efficiently than those who are non-separable.
#
# For example, the Sobel filter $S_x$ can be decomposed as
#
# \begin{equation*}
# \begin{bmatrix}
# -1 & 0 & +1\\
# -2 & 0 & +2\\
# -1 & 0 & +1\\
# \end{bmatrix} =
# \begin{bmatrix}
# +1 \\
# +2 \\
# +1 \\
# \end{bmatrix}
# \begin{bmatrix}
# -1 & 0 & +1
# \end{bmatrix}
# \end{equation*}
#
# which means that the convolution of I with the 2D filter $S_x$ can be simplified to two
# convolutions with 1D filters,
#
# \begin{equation*}
# I * S_x =
# \frac{1}{8} I *
# \begin{bmatrix}
# -1 & 0 & +1\\
# -2 & 0 & +2\\
# -1 & 0 & +1\\
# \end{bmatrix}
# \end{equation*}
#
# \begin{equation*}
# =
# \frac{1}{8}\Bigg(I *
# \begin{bmatrix}
# +1\\
# +2\\
# +1\\
# \end{bmatrix} \Bigg) *
# \begin{bmatrix}
# -1 & 0 & +1
# \end{bmatrix}
# \end{equation*}
#
# For the comparison of the computational complexity of separable and non-separable
# filters, check out the lecture notes.
#
# * We will compare the performance of separable and non-separable filters.
# - Create a separable filter (for example a Gaussian filter, you can use ```gaussian_filter``` function given below)
# - Use ```decomposeSeparableFilter``` to decompose the seperable filter to two, 1D filters.
# - Implement ```applyImageSepFilter``` function. You can use ```applyImageFilter``` function implemented in Applying Convolutional Filters (Hint: two lines of code)
# - Perform convolution on ```img``` matrix and measure the time for ```decomposeSeparableFilter``` and ```applyImageSepFilter```.
# - Repeat the test with several increasing filter sizes.
# - measure the elapsed time for each filtering. To get the start and end time, use time.time()
# - Plot the results in a graph (elapsed time) vs (filter size).
# - What is your conclusion ? Is filtering with separable filters always faster than with non-separable ones?
# +
def decomposeSeparableFilter(F):
h = [1]
s = len(F)
for i in range(1,s):
h.append(np.sum(F[:,i])/(np.sum(F[:,0])))
h = np.asmatrix(np.array(h))
v = np.asmatrix(F[:,0]).transpose()
return v, h
# implement the convolution with two 1D filters and return the output
def applyImageSepFilter(I, F_v, F_h):
I = applyImageFilter(I, F_h, padding='same')
output = applyImageFilter(I, F_v, padding='same')
return output
# +
# Sample image
img = cv2.imread('images/res1.png',0)
img = img[150:200,150:200]
# We crop the image, becasue larger images take considerably longer time to
# perform convolution with large filters
# Filter sizes to try out.
# Since we consider filter with odd heigh and width,
# we start from 3 and increment it by 2 till 29.
filter_sizes = range(3,30,2)
# +
# ******************************************
# Perform the comparison here
# Hint: Iterate through the filter_sizes and in each iteration perform convolution with the 2D and 1D filters
# using applyImageFilter and applyImageSepFilter functions respetively. Measure the time for each execution.
# Store the execution times and plot them at the end
# Note: Performing convolutions with larger filters could take longer time (sometimes more than a minute)
# ******************************************
time_records = []
for size in filter_sizes:
F = gaussian_filter(fSize=size, fSigma=1)
F_v, F_h = decomposeSeparableFilter(F)
# measure time used for 2D filter conv
start_2D = time.time()
output_2D = applyImageFilter(img, F, padding='same')
end_2D = time.time()
time_2D = end_2D - start_2D
start_1D = time.time()
output_1D = applyImageSepFilter(img, F_v, F_h)
end_1D = time.time()
time_1D = end_1D - start_1D
# check if the error of two conv result is within an accepatable range
print("Different of 2D and 1D separate conv result: {}".format(np.sum(np.abs(output_2D-output_1D))))
time_records.append([time_2D, time_1D])
# +
time_records = np.array(time_records)
plt.figure(figsize=(8, 4))
plt.plot(filter_sizes, time_records[:, 0], label="2D Filter Conv")
plt.plot(filter_sizes, time_records[:, 1], label="1D Filter Conv")
plt.xlabel("Filter Size")
plt.ylabel("Time Elapsed 10^(-11)s")
plt.legend()
# -
| ExerciseSession2/Convolution, Filtering and Gradients.ipynb |
#!/usr/bin/env python
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: ipynb,py
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tfrl-cookbook
# language: python
# name: tfrl-cookbook
# ---
# Visual stock/share trading environment with indicators & real data
# Chapter 5, TensorFlow 2 Reinforcement Learning Cookbook | <NAME>
import os
import random
from typing import Dict
import cv2
import gym
import numpy as np
import pandas as pd
from gym import spaces
from trading_utils import TradeVisualizer
env_config = {
"ticker": "TSLA",
"opening_account_balance": 100000,
# Number of steps (days) of data provided to the agent in one observation
"observation_horizon_sequence_length": 30,
"order_size": 1, # Number of shares to buy per buy/sell order
}
class StockTradingVisualEnv(gym.Env):
def __init__(self, env_config: Dict = env_config):
"""Stock trading environment for RL agents
The observations are stock price info (OHLCV) over a horizon as specified
in env_config. Action space is discrete to perform buy/sell/hold trades.
Args:
ticker (str, optional): Ticker symbol for the stock. Defaults to "MSFT".
env_config (Dict): Env configuration values
"""
super(StockTradingVisualEnv, self).__init__()
self.ticker = env_config.get("ticker", "MSFT")
data_dir = "data"
self.ticker_file_stream = os.path.join(f"{data_dir}", f"{self.ticker}.csv")
assert os.path.isfile(
self.ticker_file_stream
), f"Historical stock data file stream not found at: data/{self.ticker}.csv"
# Stock market data stream. An offline file stream is used. Alternatively, a web
# API can be used to pull live data.
# Data-Frame: Date Open High Low Close Adj-Close Volume
self.ohlcv_df = pd.read_csv(self.ticker_file_stream)
self.opening_account_balance = env_config["opening_account_balance"]
# Action: 0-> Hold; 1-> Buy; 2 ->Sell;
self.action_space = spaces.Discrete(3)
self.observation_features = [
"Open",
"High",
"Low",
"Close",
"Adj Close",
"Volume",
]
self.obs_width, self.obs_height = 128, 128
self.horizon = env_config.get("observation_horizon_sequence_length")
self.observation_space = spaces.Box(
low=0,
high=255,
shape=(128, 128, 3),
dtype=np.uint8,
)
self.order_size = env_config.get("order_size")
self.viz = None # Visualizer
def step(self, action):
# Execute one step within the trading environment
self.execute_trade_action(action)
self.current_step += 1
reward = self.account_value - self.opening_account_balance # Profit (loss)
done = self.account_value <= 0 or self.current_step >= len(
self.ohlcv_df.loc[:, "Open"].values
)
obs = self.get_observation()
return obs, reward, done, {}
def reset(self):
# Reset the state of the environment to an initial state
self.cash_balance = self.opening_account_balance
self.account_value = self.opening_account_balance
self.num_shares_held = 0
self.cost_basis = 0
self.current_step = 0
self.trades = []
if self.viz is None:
self.viz = TradeVisualizer(
self.ticker,
self.ticker_file_stream,
"TFRL-Cookbook Ch4-StockTradingVisualEnv",
)
return self.get_observation()
def render(self, **kwargs):
# Render the environment to the screen
if self.current_step > self.horizon:
self.viz.render(
self.current_step,
self.account_value,
self.trades,
window_size=self.horizon,
)
def close(self):
if self.viz is not None:
self.viz.close()
self.viz = None
def get_observation(self):
"""Return a view of the Ticker price chart as image observation
Returns:
img_observation (np.ndarray): Image of ticker candle stick plot
with volume bars as observation
"""
img_observation = self.viz.render_image_observation(
self.current_step, self.horizon
)
img_observation = cv2.resize(
img_observation, dsize=(128, 128), interpolation=cv2.INTER_CUBIC
)
return img_observation
def execute_trade_action(self, action):
if action == 0: # Hold position
return
order_type = "buy" if action == 1 else "sell"
# Stochastically determine the current stock price based on Market Open & Close
current_price = random.uniform(
self.ohlcv_df.loc[self.current_step, "Open"],
self.ohlcv_df.loc[self.current_step, "Close"],
)
if order_type == "buy":
allowable_shares = int(self.cash_balance / current_price)
if allowable_shares < self.order_size:
# Not enough cash to execute a buy order
return
# Simulate a BUY order and execute it at current_price
num_shares_bought = self.order_size
current_cost = self.cost_basis * self.num_shares_held
additional_cost = num_shares_bought * current_price
self.cash_balance -= additional_cost
self.cost_basis = (current_cost + additional_cost) / (
self.num_shares_held + num_shares_bought
)
self.num_shares_held += num_shares_bought
self.trades.append(
{
"type": "buy",
"step": self.current_step,
"shares": num_shares_bought,
"proceeds": additional_cost,
}
)
elif order_type == "sell":
# Simulate a SELL order and execute it at current_price
if self.num_shares_held < self.order_size:
# Not enough shares to execute a sell order
return
num_shares_sold = self.order_size
self.cash_balance += num_shares_sold * current_price
self.num_shares_held -= num_shares_sold
sale_proceeds = num_shares_sold * current_price
self.trades.append(
{
"type": "sell",
"step": self.current_step,
"shares": num_shares_sold,
"proceeds": sale_proceeds,
}
)
if self.num_shares_held == 0:
self.cost_basis = 0
# Update account value
self.account_value = self.cash_balance + self.num_shares_held * current_price
if __name__ == "__main__":
env = StockTradingVisualEnv()
obs = env.reset()
num_episodes = 2 # Increase num_episodes
for _ in range(num_episodes):
action = env.action_space.sample()
next_obs, reward, done, _ = env.step(action)
env.render()
| Chapter05/stock_trading_visual_env.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
# -
from get_aq_data import get_flo_data, ID_to_name, TICKS_TWO_HOURLY
data, hourly_mean, daily_mean = get_flo_data()
subset = data
subset = subset.mean(axis=1)
just_sundays = subset[subset.index.dayofweek == 6].to_frame('Mean')
just_sundays['Marathon'] = 'Non-Marathon'
just_sundays.loc['2019-05-05', 'Marathon'] = 'Marathon'
just_sundays.loc['2019-07-21', 'Marathon'] = 'Just Ride'
marathon = just_sundays[just_sundays['Marathon'] == 'Marathon']
marathon = marathon.groupby(marathon.index.time).mean()
just_ride = just_sundays[just_sundays['Marathon'] == 'Just Ride']
just_ride = just_ride.groupby(just_ride.index.time).mean()
non_marathon = just_sundays[just_sundays['Marathon'] == 'Non-Marathon']
non_marathon_mean = non_marathon.groupby(non_marathon.index.time).mean()
non_marathon_std = non_marathon.groupby(non_marathon.index.time).std()
marathon = marathon['Mean']
just_ride = just_ride['Mean']
non_marathon_mean = non_marathon_mean['Mean']
non_marathon_std = non_marathon_std['Mean']
ax = marathon.plot(label='Marathon (5th May)', figsize=(10,6))
non_marathon_mean.plot(ax=ax, label='Other Sundays ($\pm$ 1SD)')
just_ride.plot(ax=ax, label="Let's Ride (21st July)")
plt.legend()
filled_top = non_marathon_mean + non_marathon_std
filled_bottom = non_marathon_mean - non_marathon_std
filled_bottom[filled_bottom < 0] = 0
plt.fill_between(non_marathon_mean.index, filled_bottom, filled_top, alpha=0.2, color='C1')
plt.suptitle('')
plt.title('$\mathrm{PM}_{2.5}$ on special event days vs other Sundays')
plt.title('7th Mar-23rd Aug 2019', loc='right', fontstyle='italic')
plt.ylabel(r'$\mathrm{PM}_{2.5}$ ($\mu g / m^3$)')
plt.xlabel('Time')
plt.xticks(TICKS_TWO_HOURLY)
plt.grid()
plt.tight_layout()
plt.savefig('graphs/SpecialEvent_TimeSeries_WholePeriod.png', dpi=300)
| Plot Special Event days - Whole Period.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rahul1990gupta/indic-nlp-datasets/blob/master/examples/Gettinng_started_with_processing_hindi_text.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="aqjCQxMnTUlo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="384913fe-0eed-455b-b0c9-1f650ac9bc8d"
# Set up the required libraries
# ! pip install indic-nlp-datasets==0.1.2
# + id="mT6vYEEqTjue" colab_type="code" colab={}
from idatasets import load_devdas
devdas = load_devdas()
# devdas.data is a generator of paragraphs
paragraphs = list(devdas.data)
text = " ".join(paragraphs)
words = text.split(" ")
# + id="haD0uuJxT7BH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="aa7df264-31c6-43b2-cdb2-7c652233db33"
# Let's print the most common words
from collections import Counter
cnt = Counter(words)
cnt.most_common(10)
# + id="lElg1KALVEM3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="3fce7a99-43ad-4978-b6f6-dcd3d874fe80"
# Let's remove the stop words before printing most common words
from spacy.lang.hi import Hindi
nlp = Hindi()
doc = nlp(text)
not_stop_words = []
for token in doc:
if token.is_stop:
continue
if token.is_punct or token.text =="|":
continue
not_stop_words.append(token.text)
not_stop_cnt = Counter(not_stop_words)
not_stop_cnt.most_common(10)
# + id="a1akbf-abkPE" colab_type="code" colab={}
# Let's render this in wordcloud
# first import the rquired libraries
from wordcloud import WordCloud
from spacy.lang.hi import STOP_WORDS as STOP_WORS_HI
import matplotlib.pyplot as plt
# + id="iypDqB00dsYx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="b9493ca9-da25-44e5-ec5d-ce4420280e08"
wordcloud = WordCloud(
width=400,
height=300,
max_font_size=50,
max_words=1000,
background_color="white",
stopwords=STOP_WORDS_HI,
).generate(text)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# + id="4KX7Q_9PdxqS" colab_type="code" colab={}
# That doesn't look right. We need to provide a custom font file to render it correctly.
# the issue is highlighted here: https://github.com/amueller/word_cloud/issues/70
import requests
url = "https://hindityping.info/download/assets/Hindi-Fonts-Unicode/gargi.ttf"
r = requests.get(url, allow_redirects=True)
font_path="gargi.ttf"
with open(font_path, "wb") as fw:
fw.write(r.content)
# + id="gRanB6w0gbBp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="b3a17d61-f204-42d6-96b7-b8fcb63d7e17"
wordcloud = WordCloud(
width=400,
height=300,
max_font_size=50,
max_words=1000,
background_color="white",
stopwords=STOP_WORDS_HI,
font_path=font_path
).generate(text)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# + id="ghnpMGzngcOn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="210066ec-b4a4-4928-84cf-e667a5e82159"
wordcloud = WordCloud(
width=400,
height=300,
max_font_size=50,
max_words=1000,
background_color="white",
stopwords=STOP_WORDS_HI,
regexp=r"[\u0900-\u097F]+",
font_path=font_path
).generate(text)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# + id="Ld2kLJ8rggOH" colab_type="code" colab={}
| examples/Getting_started_with_processing_hindi_text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Photodiode detector
#
# +
# sphinx_gallery_thumbnail_path = '../images/Basics_Photodiode.png'
def run(Plot, Save):
from PyMieSim.Detector import Photodiode
Detector = Photodiode(NA = 0.8,
Sampling = 1001,
GammaOffset = 0,
PhiOffset = 0)
if Plot:
Detector.Plot()
if Save:
from pathlib import Path
dir = f'docs/images/{Path(__file__).stem}'
Detector.SaveFig(dir)
if __name__ == '__main__':
run(Plot=True, Save=False)
| docs/source/Basics/Photdiode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''scratch'': conda)'
# name: python3
# ---
# # Cython for fast python
# > Trying out cython
#
# - comments: true
# - badges: true
# - categories: [programming]
# - publishes: true
# A philosophy I like to follow in Python is "Python is slow, let's code everything and than see if we have any bottleneck we should replace with something else". If you find such bottlenecks, you can replace them with a faster library or another language.
#
# Replacing Python by Cython is one of the ways to speedup such bottlenecks.
# Let's try to code a correlation computation function in cython and compare it to Python or Numpy.
# First, the easy numpy version
a = [1,2,3,4,5,6] * 1000
b = [2,3,4,5,7,6] * 1000
# +
import numpy as np
# %timeit np.corrcoef(a,b)
# -
# Now, a simple pupre Python version.
# +
def correlation(a_samples, b_samples):
a_mean = sum(a_samples) / len(a_samples)
b_mean = sum(b_samples) / len(b_samples)
diff_a_samples = [a - a_mean for a in a_samples]
diff_b_samples = [b - b_mean for b in b_samples]
covariance = sum([diff_a * diff_b for diff_a, diff_b in zip(diff_a_samples, diff_b_samples)])
variance_a = sum(diff_a ** 2 for diff_a in diff_a_samples)
variance_b = sum(diff_b ** 2 for diff_b in diff_b_samples)
correlation = covariance / (variance_a * variance_b) ** (1/2)
return correlation
# %timeit correlation(a,b)
# -
# Let's now try to build a version in cython.
#
# First, I have to transform the Python lists in C arrays.
# Then I compute the values one by one, making sure that I don't leave any Python operations.
# %load_ext cython
# + language="cython"
#
# import cython
#
# from libc.stdlib cimport malloc, free
# from libc.math cimport sqrt
#
# def cython_correlation(a_samples, b_samples):
# cdef int a_len = len(a_samples)
# cdef int b_len = len(b_samples)
#
# # First we convert the Python lists into C arrays
# a_samples_array = <int *>malloc(a_len*cython.sizeof(int))
# if a_samples_array is NULL:
# raise MemoryError
# b_samples_array = <int *>malloc(b_len*cython.sizeof(int))
# if b_samples_array is NULL:
# raise MemoryError
#
# cdef int i = 0
# for i in range(a_len):
# a_samples_array[i] = a_samples[i]
# b_samples_array[i] = b_samples[i]
#
# # Now we can compute the correlation
#
# # First compute the sum of the arrays
# cdef int a_sum = 0
# for i in range(a_len):
# a_sum += a_samples_array[i]
#
# cdef int b_sum = 0
# for i in range(a_len):
# b_sum += b_samples_array[i]
#
# # Then we can compute the means
# cdef double a_mean
# cdef double b_mean
# a_mean = a_sum / a_len
# b_mean = b_sum / b_len
#
# # We then put the difference to the means in new arrays
# diff_a_samples = <double *>malloc(a_len*cython.sizeof(double))
# if diff_a_samples is NULL:
# raise MemoryError
# diff_b_samples = <double *>malloc(b_len*cython.sizeof(double))
# if diff_b_samples is NULL:
# raise MemoryError
#
# for i in range(a_len):
# diff_a_samples[i] = a_samples_array[i] - a_mean
# diff_b_samples[i] = b_samples_array[i] - b_mean
#
# # This then allows us to easily compute the
# # covariance and variances.
# cdef double covariance = 0
# for i in range(a_len):
# covariance += diff_a_samples[i] * diff_b_samples[i]
#
# cdef double variance_a = 0
# cdef double variance_b = 0
# for i in range(a_len):
# variance_a += diff_a_samples[i] ** 2
# variance_b += diff_b_samples[i] ** 2
#
#
# cdef double correlation = 0
# cdef double variance_product = (variance_a * variance_b)
# correlation = covariance / sqrt(variance_product)
#
# free(a_samples_array)
# free(b_samples_array)
#
# return correlation
#
# +
# %timeit cython_correlation(a,b)
# 10000 loops, best of 5: 154 µs per loop
# -
# Nice! We got a 6X improvement compared to Numpy and 15X improvement compared to pure Python. Pretty cool.
| _notebooks/2021-10-15-cython.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="dD6Df_3PiH3M" colab_type="text"
# ### Mounting the G-Drive
# + id="Z59SiRYVhT-v" colab_type="code" outputId="b76c6640-3e10-480d-f397-2608aa3c6443" colab={"base_uri": "https://localhost:8080/", "height": 0}
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="OSa93CQPitTF" colab_type="text"
# ### Frame Extraction
# + id="MszhTpfHiL1a" colab_type="code" outputId="0521879e-1b11-4592-affa-54ab887b3d12" colab={"base_uri": "https://localhost:8080/", "height": 34}
# imports here
import sys
import argparse
import cv2
print(cv2.__version__)
import numpy as np
import os
# + id="-WwwwgHmjlqV" colab_type="code" colab={}
def extract_image_one_fps(video_source_path):
vidcap = cv2.VideoCapture(video_source_path)
count = 0
success = True
while success:
vidcap.set(cv2.CAP_PROP_POS_MSEC,(count*100))
success,image = vidcap.read()
## Stop when last frame is identified
image_last = cv2.imread("/content/gdrive/My Drive/Key Frame/output1/frame{}.png".format(count-1))
if np.array_equal(image,image_last):
break
cv2.imwrite("/content/gdrive/My Drive/Key Frame/output1/frame%d.png" % count, image) # save frame as PNG file
print('{}.sec reading a new frame: {} '.format(count,success))
count += 1
# + [markdown] id="mnTNZZSOtP7x" colab_type="text"
# #### Generating Frames
# + id="KdThx5vhldLJ" colab_type="code" outputId="be48743c-75e8-492f-ff06-dd8331bc2341" colab={"base_uri": "https://localhost:8080/", "height": 0}
video_source_path = '/content/gdrive/My Drive/Key Frame/input1.avi'
extract_image_one_fps(video_source_path)
# + id="hxqGWQ1mpGEX" colab_type="code" colab={}
# + [markdown] id="HMWMrDaktToV" colab_type="text"
# #### Displaying Some Frames
# + id="641mP5AVpF3R" colab_type="code" outputId="8ca84e54-ca16-484f-f0de-a2bd051eea12" colab={"base_uri": "https://localhost:8080/", "height": 4066}
# Displaying Images
# %pylab inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
for count in range(0, 159, 10):
img=mpimg.imread('/content/gdrive/My Drive/Key Frame/output1/frame%d.png' % count)
imgplot = plt.imshow(img)
plt.show()
# + id="QCMmtwgOpFz3" colab_type="code" colab={}
# + [markdown] id="MrxKI9Ctuf87" colab_type="text"
# #### Extraction Algorithm
# + id="Apg4rvM2vUJv" colab_type="code" outputId="2f144403-e954-485c-eea8-41a8a12d2d90" colab={"base_uri": "https://localhost:8080/", "height": 52}
from PIL import Image
img = Image.open('/content/gdrive/My Drive/Key Frame/output1/frame0.png').convert('LA')
#plt.imshow(img)
gray_img = np.asarray(img)
hist = cv2.calcHist([gray_img],[0],None,[256],[0,256])
#plt.hist(gray_img.ravel(),256,[0,256])
#plt.title('Histogram for gray scale picture')
#plt.show()
print(hist.shape)
print(gray_img.shape)
# + id="ORSCo0rZ1lgx" colab_type="code" colab={}
r, c, ch = gray_img.shape
# + id="gPc6LYUSwcA6" colab_type="code" outputId="8e7238bd-17d0-4d65-b9a3-40caa5a161e2" colab={"base_uri": "https://localhost:8080/", "height": 282}
pdf = hist / (r*c*ch)
plt.plot(np.array(range(0, 256)), pdf)
# + id="cdEnEQe725Dw" colab_type="code" outputId="9822fd9b-6222-4b22-bfba-a6bdb6ccd63b" colab={"base_uri": "https://localhost:8080/", "height": 34}
pdf.shape
# + id="-kBRsZzmmFkO" colab_type="code" colab={}
#pdf
# + id="sEtT5iLHwb3T" colab_type="code" colab={}
def Entropy(pdf):
pdf = pdf + 0.001
pdf = pdf / np.mean(pdf)
return -np.matmul(pdf.T, np.log(pdf))[0][0]
# + id="LwfiQ_mi2XHl" colab_type="code" outputId="4830f067-7d67-47b1-b171-c60230af4c13" colab={"base_uri": "https://localhost:8080/", "height": 34}
Entropy(pdf)
# + id="iRYMAa5wwbqC" colab_type="code" colab={}
# + id="zwolYKLSufoj" colab_type="code" outputId="8cae7e7f-907e-4b1d-cd5d-3594d06c6a1c" colab={"base_uri": "https://localhost:8080/", "height": 34}
E = []
for i in range(0, 159):
img = Image.open('/content/gdrive/My Drive/Key Frame/output1/frame%d.png' %i).convert('LA')
gray_img = np.asarray(img)
hist = cv2.calcHist([gray_img],[0],None,[256],[0,256])
r, c, ch = gray_img.shape
pdf = hist / (r*c*ch)
e = Entropy(pdf)
E.append(e)
len(E)
# + id="76Va4-T2pG-D" colab_type="code" outputId="55499a41-6067-4a28-e946-44fe205a64b0" colab={"base_uri": "https://localhost:8080/", "height": 282}
plt.plot(range(len(E)), E)
# + id="QOk8Bz7aldPt" colab_type="code" colab={}
frames = [0]*len(E)
# + id="v4_wpGujldTV" colab_type="code" colab={}
for i in range(1, len(E)-1):
if E[i] > E[i+1] and E[i] > E[i-1]:
frames[i] = 1
elif E[i+1] > E[i] and E[i-1] > E[i]:
frames[i] = 1
# + id="xr0RTmYjldWW" colab_type="code" outputId="f7bb1c0e-15b0-496c-bb4f-c36c42d8b034" colab={"base_uri": "https://localhost:8080/", "height": 34}
frames.count(1)
# + id="zkSsWP6v6oP-" colab_type="code" outputId="994d4bed-6416-4e50-e5db-8c238811bef9" colab={"base_uri": "https://localhost:8080/", "height": 34}
(np.array(frames)*np.array(E)).shape
# + id="nORY-gnLldZF" colab_type="code" outputId="d66c1dd4-8af0-4073-a315-2085bc6395e4" colab={"base_uri": "https://localhost:8080/", "height": 265}
arr = np.array(frames)*np.array(E)
x = np.where(arr != 0)[0]
y = []
for i in x:
y.append(arr[i])
y = np.array(y)
plt.scatter(x, y)
plt.show()
# + id="mzSVF0Roldbs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="32c4e5a7-6d45-4c3c-b9d0-58dd496d9874"
y.shape
# + [markdown] id="_Gr73K2750-i" colab_type="text"
# #### Density Clustering
# + id="6NwlIhAM5zMr" colab_type="code" colab={}
N = 10
# + id="LWvveKUC5zSh" colab_type="code" colab={}
from sklearn.cluster import KMeans
# + id="PX8M56Opt4ER" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="253bbd08-d75a-4638-8138-6a33fc1c3349"
x.reshape((x.shape[0], 1))
y.reshape((y.shape[0]), 1)
X = np.stack((x, y), axis=-1)
print(X.shape)
#print(X)
# + id="fyu1fjVF5zXX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="4ff099d5-7336-48c3-b732-fd152cea10a8"
kmeans = KMeans(n_clusters = N)
kmeans.fit(X)
# + id="KaMnGljR5zaV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8a79c592-8733-4f28-daa1-5faae7810be1"
pred = kmeans.predict(X)
pred = pred.reshape((pred.shape[0], 1))
pred.shape
# + id="y2X-WbcE5zdZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1597} outputId="c8aac652-fa26-46a3-b41c-c3792d13fb00"
t = (pred == 0) * X
m = 0
c = 0
for i in range(t.shape[0]):
if t[i][1] != 0:
m += t[i][1]
c += 1
m = m/c
print(t)
print(c)
print(m)
# + id="Q4O5aErf5I7t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e4aa1daf-86b6-4eb6-e7f7-e8670467ac38"
mm = t-np.array([0, m])
ind, m = mm[0][0], mm[0][1]**2
for j in range(t.shape[0]):
if m > mm[j][1]**2:
m = mm[j][1]**2
ind = mm[j][0]
ind
# + id="sjAF8bt35Iv3" colab_type="code" colab={}
# + id="J7Lb2TS40H3R" colab_type="code" colab={}
# + id="6VK797H25zgK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f79e8384-6fda-47dd-bd68-5dfbafb5c912"
correct_frames = []
for i in range(N):
t = (pred == i) * X
m = 0
c = 0
for j in range(t.shape[0]):
if t[j][1] != 0:
m += t[j][1]
c += 1
m = m/c
mm = t-np.array([0, m])
ind, m = mm[0][0], mm[0][1]**2
for j in range(mm.shape[0]):
if m > mm[j][1]**2:
m = mm[j][1]**2
ind = mm[j][0]
correct_frames.append(int(ind))
correct_frames
# + id="KRV2iCwL5zVh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2537} outputId="3a849fec-a394-48d0-a4ec-1524eadf51d3"
for i in sorted(correct_frames):
img=mpimg.imread('/content/gdrive/My Drive/Key Frame/output1/frame%d.png' % i)
imgplot = plt.imshow(img)
plt.show()
# + id="t-mWmzUy7888" colab_type="code" colab={}
# + [markdown] id="DGth-jwh790a" colab_type="text"
# #### References
# + id="uh8dCQKo5zQX" colab_type="code" colab={}
| Key_Frame_Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/diem-ai/datascience-projects/blob/master/twitter_bot/ebay_bot_w2v.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="fBvZqu--xI3x" colab_type="text"
# # Google Setup
# + id="3RWA7-vkHsqj" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
import sys
# To add a directory with your code into a list of directories
# which will be searched for packages
sys.path.append('/content/drive/My Drive/Colab Notebooks')
# !pip install PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + [markdown] id="bpK0BoWExEYs" colab_type="text"
# # Install packages
# + id="OgFHKRzXILOQ" colab_type="code" outputId="d36a0cb1-7d8a-4023-c620-c4824dd82f53" colab={"base_uri": "https://localhost:8080/", "height": 608}
# ! pip install --upgrade gensim
# ! pip install --upgrade emoji
# + [markdown] id="nNDWknuEw_r0" colab_type="text"
# # Libraries
# + id="bPmwKsUBIQCk" colab_type="code" outputId="cc1047ab-08b6-4c99-fcc4-6e6eaa0735cf" colab={"base_uri": "https://localhost:8080/", "height": 118}
import pandas as pd
import math
import re
import nltk
import string
#Download only once
nltk.download('punkt') # pre-trained tokenizer for English
nltk.download('wordnet') # lexical database for the English language
nltk.download('stopwords')
from gensim.test.utils import common_texts
from gensim.corpora import Dictionary
from gensim.models import Word2Vec , WordEmbeddingSimilarityIndex
from gensim.similarities import SoftCosineSimilarity, SparseTermSimilarityMatrix
from gensim.models import TfidfModel
from multiprocessing import cpu_count
# + [markdown] id="-5ARIR3jI3_P" colab_type="text"
# # Data Preparation
# Load twitter dataset into pandas dataframe
# + id="F5bCxs0MIWB3" colab_type="code" outputId="1a9260b5-376b-4a4b-8837-30fa92cb1758" colab={"base_uri": "https://localhost:8080/", "height": 195}
filepath = '/content/drive/My Drive/data/'
df = pd.read_csv(filepath + "twcs.csv.zip")
df.head()
# + [markdown] id="94OuBzPFJFbE" colab_type="text"
# Retrive list of companies providing customer support on twitter
# + id="GIfk2ybESDLG" colab_type="code" colab={}
pattern = r"\A[a-z]+\Z"
authors = []
authors = [ val for val in df["author_id"].values if (re.search(pattern, val.lower())) ]
# remove duplicated values in the list
authors = list(set(authors))
#print(re.search(pattern, "12345"))
# + [markdown] id="ByLIIV0SJN71" colab_type="text"
# Select eBay's data
# + id="fpUF8V2BRt8X" colab_type="code" outputId="cad6e0dc-a6bc-483a-f9dd-96a52939c7b4" colab={"base_uri": "https://localhost:8080/", "height": 195}
select_author = "AskeBay"
data = df [df["author_id"] == select_author]
data.head()
# + [markdown] id="B9cRD6Q1JUSd" colab_type="text"
# Select neccessary features for data processing
# + id="g8yXF3QpU01H" colab_type="code" outputId="0eb57d4e-4972-4bfd-cc9a-357472232a86" colab={"base_uri": "https://localhost:8080/", "height": 195}
# select sprintcare's responses to their customers questions
select_features = ["tweet_id", "text", "response_tweet_id", "in_response_to_tweet_id"]
data = data[select_features]
data.head()
# + [markdown] id="j-bjh8leJZUz" colab_type="text"
# Create a data frame with all responses from eBay
# + id="hNNJa0fqVhpX" colab_type="code" outputId="bcd70082-d112-4801-92e2-c40a677bd8fe" colab={"base_uri": "https://localhost:8080/", "height": 195}
# select eBay's questions
questions = df[ df["text"].str.contains("AskeBay") ]
questions.head()
# + id="8lhDkzvMXPUL" colab_type="code" outputId="d98a216b-1ad9-49a1-c5c1-466992fd2483" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(questions)
# + id="B_vGhpdFWinC" colab_type="code" outputId="208ba538-dcb2-47a7-80db-1cf0ca0ab58f" colab={"base_uri": "https://localhost:8080/", "height": 34}
questions.iloc[0]["text"]
# + id="SiWZXwb0WDdF" colab_type="code" outputId="2ebac995-c047-418e-f9ad-ed12f8db990e" colab={"base_uri": "https://localhost:8080/", "height": 70}
idx = 0
print("id: {}, question: {}".format(questions.iloc[idx]["author_id"]
, questions.iloc[idx]["text"]))
print("answer: {}".format(data.iloc[idx]["text"]))
# + id="8Ov6QyTFXOjQ" colab_type="code" outputId="1d63b6f7-c260-4b5e-b69c-1a40bb993ec2" colab={"base_uri": "https://localhost:8080/", "height": 70}
idx = 100
print("id: {}, question: {}".format(questions.iloc[idx]["author_id"]
, questions.iloc[idx]["text"]))
print("answer: {}".format(data.iloc[idx]["text"]))
# + id="VpmlwNzkXnZB" colab_type="code" outputId="7e6cf162-0929-4675-b19c-895b2128d86f" colab={"base_uri": "https://localhost:8080/", "height": 50}
idx = 500
print("id: {}, question: {}".format(questions.iloc[idx]["author_id"]
, questions.iloc[idx]["text"]))
print("answer: {}".format(data.iloc[idx]["text"]))
# + id="Sm0UZ5gzXy1v" colab_type="code" colab={}
def get_tweet_from_id(data, tweet_id):
"""
@data: [pandas dataframe] : pandas dataframe
@tweet_id: [integer] : an tweet ID
@return : [list] : a tweet corresponding with tweet_id input
"""
return data[data["tweet_id"] == tweet_id]["text"].values
# + [markdown] id="fcbox-mJJmlP" colab_type="text"
# Create a pair of list of questions and responses
# + id="naVFbDdwYAg-" colab_type="code" colab={}
ebay_questions = []
ebay_responses = []
for i in range(len(questions)):
tweet = questions.iloc[i]
text = tweet["text"]
owner = tweet["author_id"]
ebay_questions.append(owner + "-" + text)
rep_ids = tweet["response_tweet_id"]
if (rep_ids and (not pd.isna(rep_ids))):
if "," in rep_ids:
rep_ids = rep_ids.split(",")[0]
# convert an array into string
response = "".join( get_tweet_from_id(df, int(rep_ids)) )
ebay_responses.append(response)
else:
ebay_responses.append("")
# + id="6Qt4sNNGfyCE" colab_type="code" outputId="056b1d9b-85b9-4ef8-be49-c639987485ae" colab={"base_uri": "https://localhost:8080/", "height": 54}
ebay_questions[2]
# + id="OGDDo6jZf0k9" colab_type="code" outputId="1f5743e8-bd88-4157-e63e-0ec8a593b418" colab={"base_uri": "https://localhost:8080/", "height": 34}
ebay_responses[2]
# + [markdown] id="1y5LIW_gJt24" colab_type="text"
# # Data Processing
# + id="k5jkSp2o3Cvp" colab_type="code" outputId="92badc3f-85c8-41b9-939d-911c1a4560a6" colab={"base_uri": "https://localhost:8080/", "height": 54}
import spacy
nlp = spacy.load("en", disable=['parser', 'tagger', 'ner'])
stops = spacy.lang.en.stop_words.STOP_WORDS
from nltk.corpus import stopwords
SW = set(stopwords.words("english"))
SW.add("askebay")
def normalize(text):
"""
Lemmatize tokens and remove them if they are stop words
"""
text = nlp(text.lower())
lemmatized = list()
for word in text:
lemma = word.lemma_.strip()
if (lemma and lemma not in SW ):
lemmatized.append(lemma)
return lemmatized
# test normalize()
text = ["applesupport tried resetting my settings restarting my phone all that 🙃"
, "i need answers because it is annoying 🙃"
, "That's great it has iOS 111 as we can rule out being outdated Any steps tried since this started Do you recall when it started"]
print([normalize(sent) for sent in text])
# + id="DE2OULhH8LyO" colab_type="code" outputId="31f59dff-8ebc-4c69-ce85-cdb26dfc1183" colab={"base_uri": "https://localhost:8080/", "height": 70}
import emoji
def clean_emoji(text):
if (emoji.emoji_count(text) > 0):
return re.sub(emoji.get_emoji_regexp(), r"", text)
else:
return text
# test clean_emoji()
text = ["applesupport tried resetting my settings restarting my phone all that 🙃"
, "i️ need answers because it is annoying 🙃"
, "That's great it has iOS 111 as we can rule out being outdated Any steps tried since this started Do you recall when it started"]
print([clean_emoji(sent) for sent in text])
print([normalize(clean_emoji(sent)) for sent in text])
# + id="CyD_RvJF8q7r" colab_type="code" outputId="32e2ad25-b93a-4a46-a9d4-fbb23115421c" colab={"base_uri": "https://localhost:8080/", "height": 70}
# from string import maketrans
# Remove the punctuation of text
def clean_punctuation(text):
"""
@text: is a text with/without the punctuation
@return a text without punctuation
"""
re_punc = re.compile("[%s]" % re.escape(string.punctuation))
return re_punc.sub("", text)
# test clean_punctuation()
text = ["applesupport tried resetting my settings #@ restarting my phone && all that 🙃"
, "i️ need answers because it is annoying 🙃 ..."
, "That's great it has iOS 111 as we can rule out being outdated Any steps tried since this started Do you recall when it started"]
print([ clean_punctuation( sent ) for sent in text])
print([ normalize(clean_emoji(clean_punctuation(sent))) for sent in text])
# + id="XK-0vqwQ-vD3" colab_type="code" outputId="72223ddd-f11b-4209-de23-9196b063ef94" colab={"base_uri": "https://localhost:8080/", "height": 50}
def decontracted(text):
# specific
text = re.sub(r"won’t", "will not", text)
text = re.sub(r"won\'t", "will not", text)
text = re.sub(r"can’t", "can not", text)
text = re.sub(r"can\'t", "can not", text)
text = re.sub(r"dont’t", "do not", text)
text = re.sub(r"don\'t", "do not", text)
text = re.sub(r"doesn’t", "does not", text)
text = re.sub(r"doesn\'t", "does not", text)
text = re.sub(r"y’all", "you all", text)
text = re.sub(r"y\'all", "you all", text)
# general
text = re.sub(r"’t", " not", text)
text = re.sub(r"n\'t", " not", text)
text = re.sub(r"’re", " are", text)
text = re.sub(r"\'re", " are", text)
text = re.sub(r"’s", " is", text)
text = re.sub(r"\'s", " is", text)
text = re.sub(r"’d", " would", text)
text = re.sub(r"\'d", " would", text)
text = re.sub(r"’ll", " will", text)
text = re.sub(r"\'ll", " will", text)
text = re.sub(r"’t", " not", text)
text = re.sub(r"\'t", " not", text)
text = re.sub(r"’ve", " have", text)
text = re.sub(r"\'ve", " have", text)
text = re.sub(r"’m", " am", text)
text = re.sub(r"\'m", " am", text)
return text
text = "@applesupport y’all gotta fix this i.t problem on everyone is phone i just want to be able to say it again not i.t',"
print(decontracted(text))
print(normalize(clean_emoji(clean_punctuation(decontracted(text )))))
# + id="JtQkYXs5BJbl" colab_type="code" outputId="0260cb6e-1c64-4af3-a9fb-23fb07da4087" colab={"base_uri": "https://localhost:8080/", "height": 34}
def clean_num(text):
"""
@Remove numberics in the text
@return the text without numberics
"""
re_num = r"\d+"
return re.sub(re_num, "", text)
# test clean_num()
text = "115855 That's great it has iOS 11.1 as we can rule out being outdated. Any steps tried since this started?"
text = clean_punctuation(text)
print( clean_num(text) )
# + id="xR4t3DkC-6Ph" colab_type="code" outputId="aeb49cec-b52a-4f5b-e91e-44de658c53bd" colab={"base_uri": "https://localhost:8080/", "height": 34}
def preprocessing(text):
# text = clean_at(text)
text = clean_emoji(text)
text = decontracted(text)
text = clean_punctuation(text)
text = clean_num(text)
text = normalize(text)
return text
# test preprocessing()
text = ["@applesupport tried resetting my settings .. restarting my phone .. all that"
, "i need answers because it is annoying 🙃"
, "@115855 That's great it has iOS 11.1 as we can rule out being outdated. Any steps tried since this started?"]
print( [preprocessing(sent) for sent in text] )
# + id="HC5lmtMLsM-p" colab_type="code" outputId="f3b04f6f-495d-4584-9cda-06392157e12f" colab={"base_uri": "https://localhost:8080/", "height": 34}
def clean_at(text):
re_at = r"^@\S+"
return re.sub(re_at, "", text)
# test clean_at()
text_at = "@115855 That's great it has iOS 11.1 as we can rule out being outdated. Any steps tried since this started?"
clean_at(text_at)
# + id="N3ygB_lQKA50" colab_type="code" colab={}
# processing questions
processed_ebay_questions = [preprocessing(quest) for quest in ebay_questions]
ebay_responses = [clean_at(resp) for resp in ebay_responses]
# + id="r9lJ75lyKsTz" colab_type="code" outputId="882d0800-232a-4dea-ad17-91d8522e01a5" colab={"base_uri": "https://localhost:8080/", "height": 218}
processed_ebay_questions[:2]
# + id="imUScpDdO5NM" colab_type="code" outputId="7105a3b2-c5a1-4e1b-dfe1-9d2df66da041" colab={"base_uri": "https://localhost:8080/", "height": 50}
ebay_questions[:2]
# + id="4rfXR0E3PAe_" colab_type="code" outputId="345d2dd6-e14e-4215-ce4f-3175f3304f5a" colab={"base_uri": "https://localhost:8080/", "height": 70}
ebay_responses[:2]
# + id="oK4NC2JBcWzX" colab_type="code" outputId="1cfd8134-c8d8-4d66-9c1f-fae98aa96f93" colab={"base_uri": "https://localhost:8080/", "height": 218}
processed_ebay_questions[3:5]
# + id="-ZspKBcQcnuA" colab_type="code" outputId="1d076ebf-e871-486b-9b46-a8d04da58e4e" colab={"base_uri": "https://localhost:8080/", "height": 70}
ebay_responses[3:5]
# + [markdown] id="T6IMotA3rT81" colab_type="text"
# # Build model with small dataset
# + id="3p9EPBBNhc0d" colab_type="code" colab={}
## make a test on 10 questions and their 10 responses
test_questions = processed_ebay_questions[:10]
test_responses = ebay_responses[0:10]
test_dictionary = Dictionary(test_questions)
test_tfidf = TfidfModel(dictionary = test_dictionary)
test_word2vec = Word2Vec(test_questions
, workers=cpu_count()
, min_count=5
, size=300
, seed=12345)
test_sim_index = WordEmbeddingSimilarityIndex(test_word2vec.wv)
test_sim_matrix = SparseTermSimilarityMatrix(test_sim_index
, test_dictionary
, test_tfidf
, nonzero_limit=100)
test_bow_corpus = [test_dictionary.doc2bow(document) for document in test_questions]
test_docsim_index = SoftCosineSimilarity(test_bow_corpus, test_sim_matrix, num_best=5)
# + [markdown] id="AdxjSwHyrYCc" colab_type="text"
# # Make a test with small model
# + id="6ik9DSdnjGA8" colab_type="code" outputId="153b9f00-9387-40f7-a833-bbbafd21f8b5" colab={"base_uri": "https://localhost:8080/", "height": 188}
query = "I need to contact customer service to report an purchase issue"
pro_query = preprocessing(query)
test_vector = test_dictionary.doc2bow(pro_query)
test_vector_tfidf = test_tfidf[test_vector]
sims = test_docsim_index[test_vector_tfidf]
for i, score in sims:
print(ebay_responses[i], score)
# + [markdown] id="cs1jbHdm3rxG" colab_type="text"
# @115938 As the seller is responsible for the courier, they can claim for lost items from the courier. You still have to open a claim through eBay to be protected. Please follow the previous steps. We cant outbound DM. You can DM us if you have any other questions. ^SG 0.9561439156532288
#
# @AskeBay Need to contact customer service directly. Please DM contact email for customer service 0.842599630355835
# 0.842599630355835
#
# @115938 Hi there. Sorry to hear the item has not arrived. Is there tracking for the item? If not, I'd advise opening an item not received claim. You can learn more about that here> https://t.co/0toc8cKSB0. Any other questions, let us know. ^SG 0.6582105755805969
#
# 0.52857506275177
# /usr/local/lib/python3.6/dist-packages/gensim/similarities/termsim.py:358: RuntimeWarning: divide by zero encountered in true_divide
#
#
# + id="gUScb1EOlTyR" colab_type="code" outputId="a5e64618-49a3-467a-cc70-b7a54844753f" colab={"base_uri": "https://localhost:8080/", "height": 188}
query = "I need an assistance. The item is marked delivery but I never receive it"
pro_query = preprocessing(query)
test_vector = test_dictionary.doc2bow(pro_query)
test_vector_tfidf = test_tfidf[test_vector]
sims = test_docsim_index[test_vector_tfidf]
for i, score in sims:
print(ebay_responses[i], score)
# + [markdown] id="iBgTVyqV1qdE" colab_type="text"
# '''
# @115938 Hi there. Sorry to hear the item has not arrived. Is there tracking for the item? If not, I'd advise opening an item not received claim. You can learn more about that here> https://t.co/0toc8cKSB0. Any other questions, let us know. ^SG 0.9377691149711609
#
# @AskeBay Need to contact customer service directly. Please DM contact email for customer service 0.6280146837234497
#
# 0.448180615901947
# 0.3625844419002533
# @115938 As the seller is responsible for the courier, they can claim for lost items from the courier. You still have to open a claim through eBay to be protected. Please follow the previous steps. We cant outbound DM. You can DM us if you have any other questions. ^SG 0.27123868465423584
#
#
# + id="B0u9moTGk9_W" colab_type="code" outputId="f7e73e7b-1cb7-4bac-bec8-cdbbc24a0b1d" colab={"base_uri": "https://localhost:8080/", "height": 87}
print(ebay_questions[8])
print(test_questions[8])
print(test_responses[8])
# + [markdown] id="Vy3eXZAUlsBF" colab_type="text"
# # Build the mode with entire corpus
# + id="5B2hQf1ZlqhE" colab_type="code" colab={}
dictionary = Dictionary(processed_ebay_questions)
tfidf = TfidfModel(dictionary = dictionary)
word2vec_model = Word2Vec(processed_ebay_questions
, workers=cpu_count()
, min_count=5
, size=300
, seed=12345)
sim_index = WordEmbeddingSimilarityIndex(word2vec_model.wv)
sim_matrix = SparseTermSimilarityMatrix(sim_index
, dictionary
, tfidf
, nonzero_limit=100)
bow_corpus = [dictionary.doc2bow(document) for document in processed_ebay_questions]
tfidf_corpus = [tfidf[bow] for bow in bow_corpus]
docsim_index = SoftCosineSimilarity(tfidf_corpus, sim_matrix, num_best=10)
# + [markdown] id="LFtTS-RjrLYI" colab_type="text"
# # Make an example
# + [markdown] id="TZhGO7sEwbmX" colab_type="text"
# ### Example 1
# + id="3Go_Z_KMmMxy" colab_type="code" outputId="b7d03b2b-4081-4808-f8f8-25def3c07744" colab={"base_uri": "https://localhost:8080/", "height": 252}
query1 = "I need an assistance. The item is marked delivery but I never receive it"
_query1 = preprocessing(query1)
vector1 = dictionary.doc2bow(_query1)
vector1_tfidf = tfidf[vector1]
sims1 = docsim_index[vector1_tfidf]
for i, score in sims1:
print(ebay_responses[i], score)
# + [markdown] id="lhs08coC27U-" colab_type="text"
# - If you do run into this and the fee isn't credited, just DM us the details and we'll be happy to take a look! ^B 1.0
# - We'll take a look! DM us your item #, email address, first/last name and postal code on file with more information. ^B 1.0
# - I'm sorry to hear you're having these troubles. You should be able to print labels from the app itself. Are you not getting the option when you view the purchase? ^BL 1.0
# - If an item isn't as described a return may still be needed and we would help with that, but you can see https://t.co/QDgDg5dz42 if you think it's intentional so we can ensure it's not misused. Keep us posted! ^D 1.0
# - You’re welcome to relist now, just make sure to end the listing if the buyer ends up paying. Once you open the unpaid item case you can close it on the 4th day! ^JN 1.0
# 1.0
# - Hi there! Keep in mind, once a case is closed you can't reopen it so you wouldn't be able to ask for a refund for the replacement.^C 1.0
# 1.0
# - Apologies if the way I worded things upset you. I can definitely pass your concerns on to my manager but the best course of action is to submit your suggestion to the link I provided so the relevant team gets it and can review it for possible implementation. ^V 1.0
# - Sorry about that! When did you turn them off? It may take a bit. You may also want to toggle all notifications off, then on. ^S 1.0
# /usr/local/lib/python3.6/dist-packages/gensim/similarities/termsim.py:358: RuntimeWarning: divide by zero encountered in true_divide
# Y = np.multiply(Y, 1 / np.sqrt(Y_norm))
# /usr/local/lib/python3.6/dist-packages/gensim/similarities/termsim.py:358: RuntimeWarning: invalid value encountered in multiply
# Y = np.multiply(Y, 1 / np.sqrt(Y_norm))
# + [markdown] id="cxYk8taGwPO1" colab_type="text"
# ### Example 2
# + id="wAOIf4ujqPDd" colab_type="code" outputId="0742a5cc-bc98-4e5c-fab0-197a05f6cf3f" colab={"base_uri": "https://localhost:8080/", "height": 272}
query3 = "Please repond to DM"
_query3 = preprocessing(query3)
vector3 = dictionary.doc2bow(_query3)
vectors_tfidf = tfidf[vector3]
sims3 = docsim_index[vectors_tfidf]
for i, score in sims3:
print(processed_ebay_questions[i], ebay_responses[i], score)
# + [markdown] id="w5afCkQF3aBG" colab_type="text"
# ['contact', 'several', 'time', 'amp', 'hit', 'brick', 'wall', 'try', 'arrange', 'collection', 'via', 'msg', 'bfr', 'purchase', 'ban', 'amp', 'lose', 'day', 'taking', 'httpstcoskxdicitw'] Hey there! We'd be happy to investigate further if you send us over a DM with details. Sorry if we've missed anything! ^CR 1.0
# ['glad', 'help', 'obstructive', 'rude', 'want', 'escalation', 'contact', 'detail', 'staff', 'refuse', 'give'] 1.0
#
# ['product', 'undelivered', 'purchase', 'remove', 'system', 'unable', 'contact', 'support', 'seller', 'unresponsive', 'need', 'help'] We're here to help! Send us a DM with first/last name, email address, postal/zip code and info about the item and we'll review! ^CR 1.0
#
# ['dm', 'would', 'return', 'query', 'hope', 'help'] Hey we got it! We'll pick up the conversation over there! ^CR 1.0
#
# ['poor', 'service', 'restrict', 'high', 'volume', 'seller', 'discuss', 'transaction', 'via', 'msg', 'downhill', 'well', 'service'] We'd be happy to talk with you more about what caused the restriction. Feel free to DM us with more details and we'll review. ^CR 1.0
#
# ['need', 'change', 'numb', 'since', 'long', 'use', 'find', 'way', 'thati', 'trouble', 'list'] Hey Dewi, thanks for getting in touch. You can update that here: https://t.co/LL6QKfBFMr. Let us know if we can help. ^CM 1.0
#
# ['use', 'mobile', 'app', 'create', 'list', 'check', 'mobile', 'friendly', 'laptop', 'need', 'fix'] Hey Dennis. What do you mean that if you check the mobile-friendly on the laptop? Can you please send us a screenshot? Thanks ^A 1.0
#
# ['hey', 'ebay', 'kind', 'need', 'help', 'studiescan', 'give', 'docsinfos', 'datum', 'strategynext', 'move', 'ebay', 'thx', 'lot'] We love that you'll be finding out more about eBay! Check out https://t.co/pRy6K18nod for some great info. Best of luck w/ your studies! ^LB 1.0
#
# ['bid', 'restriction', 'help', 'find', 'restrict', 'remove', 'thank', 'lot', 'httpstcotyzndsmhk'] Sure! DM us your full name, registered email address, & registered postal code. We're happy to help! ^AE 1.0
#
# ['yes', 'also', 'report', 'via', 'phone', 'tell', 'would', 'contact', 'response', 'seller'] Since the seller has been notified, it’s up to them to reach out to you if they’d like their item back. ^BT 1.0
#
# + id="mCJ9P-gOumM9" colab_type="code" outputId="c830a4ac-dac3-4e2d-8082-c594bb67e9fa" colab={"base_uri": "https://localhost:8080/", "height": 50}
print(ebay_questions[700])
print(ebay_responses[700])
# + [markdown] id="sh7iTKXRwUUB" colab_type="text"
# ### Example 3
# + id="FB5NLNf2u4xS" colab_type="code" outputId="38b75f6d-914c-453c-f5a3-62b6ea1b9b93" colab={"base_uri": "https://localhost:8080/", "height": 272}
query4 = "I'm not sure if the bottle of perfume is considered as dangerous material?"
_query4 = preprocessing(query4)
vector4 = dictionary.doc2bow(_query4)
vector4_tfidf = tfidf[vector4]
sims4 = docsim_index[vector4_tfidf]
for i, score in sims4:
print(processed_ebay_questions[i], ebay_responses[i], score)
# + [markdown] id="DF_ho1oF3Kiy" colab_type="text"
# ['wait', 'perfume', 'bottle', 'consider', 'dangerous', 'material'] Looks like you can, if they don't contain alcohol: https://t.co/NBaDNpnGh6. Glad to hear you got it shipped out! ^TL 0.9667293429374695
#
# ['light', 'travel', 'straight', 'line', 'light', 'enter', 'degree', 'center', 'degree', 'field', 'lens', 'protude', 'lot', 'camera', 'advertise', 'doe', 'half', 'sphere', 'lens', 'front', 'false', 'advertise'] Thanks for the details. As it seems to be a concern with the manufacturer specs and not the listings themselves, it does not sound like something we can action here. You will want to discuss that with the manufacturer to determine why they claim 170 instead. ^D 0.812565803527832
#
# ['generate', 'fedex', 'label', 'overestimate', 'weight', 'come', 'much', 'light', 'adjust', 'price', 'thank', 'advance'] Yes! FedEx doesn't charge you until the item is delivered, so they'll charge you the correct amount based on the weight. ^BL 0.8121383190155029
#
# ['constitute', 'special', 'punctuation', 'mean', 'remove', 'normal', 'punctuation', 'like', 'full', 'stop', 'say', 'slash', 'constitute', 'swear', 'please', 'clarify'] It's difficult to identify for sure, but it looks likes the term "bit cheap" may not recognize the space there. I't try another word and see if that works! ^D 0.8120605945587158
#
# ['anytime', 'day', 'actually', 'think', 'something', 'go', 'sell', 'jack', 'use', 'sell', 'alot', 'search', 'engine', 'attempt', 'compete', 'amazon', 'seem', 'would', 'say', 'go', 'back', 'previous', 'format', 'use', 'original', 'search', 'engine', 'cassinni'] 0.8106344938278198
#
# ['amigooo', 'necesito', 'datos', 'de', 'cuenta', 'bancaria', 'para', 'el', 'reintegro', 'es', 'estado', 'actual', 'del', 'producto', 'como', 'llegó', 'destino', 'que', 'patetico', 'ustedes', 'de', 'ebay', 'hay', 'justicia', 'para', 'los', 'vendedores', 'si', 'la', 'hay', 'retiro', 'de', 'sus', 'politicas', 'absurdas'] Hi Nino. Unfortunately our Spanish support isn't available for the rest of the day. Although we recommend that you go here https://t.co/o1PE5TGxBo after logging in to find the details. ^KE 0.7033706903457642
# ['imply', 'ebay', 'feel', 'control', 'fb', 'favor', 'buyer', 'increase', 'revenue', 'let', 'face', 'amazon', 'trump', 'ebayebay', 'decline', 'past', 'year', 'ebay', 'cold', 'amp', 'unappreciative', 'sellersamazon', 'cust', 'serv', 'leave', 'ebay', 'dust', 'light', 'year'] 0.6342083811759949
# ['actually', 'week', 'miss', 'since', 'upset', 'barely', 'miss', 'coupon', 'three', 'hour', 'today', 'item', 'go', 'back', 'regular', 'price', 'also'] Sorry to hear that. I understand that would be tough, but hopefully you'll see another coupon/deal soon. ^BL 0.6339040994644165
#
# ['hello', 'touch', 'base', 'tomorrow', 'item', 'start', 'b', 'relisted', 'putt', 'thougt', 'issue', 'current', 'plan', 'action', 'order', 'upset', 'buyerscustomers', 'likely', 'affect', 'feedback', 'score', 'plan'] Following. Please understand Im doing this to put my customers at ease and protect the intergerty of my feedback. 1) for each item that the ebay computer adds a "Best Offer" to I will remove it.
# 2) I will also place a disclaimer in the listing letting customers know it was 0.6336686611175537
#
# ['ebay', 'list', 'sale', 'day', 'clearly', 'choose', 'general', 'problem', 'atm', 'ebayuk'] Are you listing on the site or app? It's possible that Good 'Til Cancelled is added to your fixed price items automatically. ^S 0.6332412362098694
#
#
| twitter_bot/ebay_bot_w2v.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importando Bibliotecas usada no problema
import pandas as pd
import numpy as np
import math
import scipy as sp
import matplotlib.pyplot as plt
# ## Ajustando configurações de plot
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
params = {'figure.figsize': [10, 5],
'axes.labelsize': 10,
'axes.titlesize':14,
'font.size': 10,
'legend.fontsize': 10,
'xtick.labelsize': 8,
'ytick.labelsize': 8
}
plt.rcParams.update(params)
# ## [2a]
# +
x = np.arange(0,10,0.20)
mean1 = 4
mean2 = 6
deav1 = math.sqrt(1)
deav2 = math.sqrt(1)
data1 = []
data2 = []
data3 = []
data4 = []
for i in x:
f_x_1 = (1/(deav1*math.sqrt(2*math.pi)))*math.exp(((-1/2)*((i-mean1)/deav1)**2))
f_x_2 = (1/(deav2*math.sqrt(2*math.pi)))*math.exp(((-1/2)*((i-mean2)/deav2)**2))
f_x_3 = f_x_1/2
f_x_4 = f_x_2/2
data1.append(f_x_1)
data2.append(f_x_2)
data3.append(f_x_3)
data4.append(f_x_4)
# -
plt.plot(x, data1, label="P(x/\u03C9\N{SUBSCRIPT ONE})")
plt.plot(x, data2, label="P(x/\u03C9\N{SUBSCRIPT TWO})")
section = np.arange(0,10,0.20)
plt.fill_between(section, data1, alpha = 0.6)
plt.fill_between(section, data2, alpha = 0.6)
plt.title("Probabilidade Condicional P(x/\u03C9\N{SUBSCRIPT ONE})")
plt.xlabel("Valor de x")
plt.ylabel("Probabilidade")
plt.legend()
plt.grid(alpha=0.2)
plt.show()
plt.plot(x, data3, label="P(\u03C9\N{SUBSCRIPT ONE})P(x/\u03C9\N{SUBSCRIPT ONE})")
plt.plot(x, data4, label="P(\u03C9\N{SUBSCRIPT TWO})P(x/\u03C9\N{SUBSCRIPT TWO})")
plt.fill_between(x, data3, alpha = 0.6)
plt.fill_between(x, data4, alpha = 0.6)
plt.title("Probabilidade Condicional P(x/\u03C9\N{SUBSCRIPT ONE}) x Probabilidade a priori P(\u03C9\N{SUBSCRIPT ONE}) ")
plt.xlabel("Valor de x")
plt.ylabel("Probabilidade")
plt.legend()
plt.grid(alpha=0.2)
plt.show()
# ## [2c]
# ### Criando um classificador Baysiano
## Classificador
def bayesanClassifier(mean1, deav1, pClass1, mean2, deav2, pClass2, data):
results = []
for x in data[0]:
class1 = 0
class2 = 0
f_x_1 = (1/(deav1*math.sqrt(2*math.pi)))*math.exp(((-1/2)*((x-mean1)/deav1)**2))
f_x_2 = (1/(deav2*math.sqrt(2*math.pi)))*math.exp(((-1/2)*((x-mean2)/deav2)**2))
class1 = pClass1*f_x_1
class2 = pClass2*f_x_2
if class1 >= class2:
results.append(0)
else:
results.append(1)
return results
# ### Gerando n = 1000 amostras
# +
##Criando os dados com 1000 amostras igualmente distribuidas entre classe1 e 2
class1Sample = np.random.normal(4, 1, 500)
class2Sample = np.random.normal(6, 1, 500)
class1 = np.full(500, 0)
dataClass1 = [class1Sample,class1]
class2 = np.full(500, 1)
dataClass2 = [class2Sample, class2]
dataFinal = np.concatenate((dataClass1, dataClass2), axis = 1)
# -
# ## Plotando as amostras
count, bins, ignored = plt.hist(class1Sample, 20, density=True, alpha = 0.6)
count2, bins2, ignored2 = plt.hist(class2Sample, 20, density=True, alpha = 0.6)
plt.plot(bins, 1/(1 * np.sqrt(2 * np.pi)) * np.exp( - (bins - 4)**2 / (2 * 1**2) ), linewidth=2, color='blue')
plt.plot(bins2, 1/(1 * np.sqrt(2 * np.pi)) * np.exp( - (bins2 - 6)**2 / (2 * 1**2) ), linewidth=2, color='orange')
plt.title("Probabilidade Condicional P(x/\u03C9\N{SUBSCRIPT ONE}) x Probabilidade a priori P(\u03C9\N{SUBSCRIPT ONE}) ")
plt.xlabel("Valor de x")
plt.ylabel("Probabilidade")
plt.grid(alpha=0.2)
plt.show()
# ### Classificação das Amostras
results = bayesanClassifier(mean1, deav1, 0.5, mean2, deav2, 0.5, dataFinal)
# ## Calculando o erro do classificador
# +
##Calculando os erros
erros = 0
for i in range(0, 1000):
if results[i] != dataFinal[1][i]:
erros += 1
print('Erro de',((erros)/1000)*100, '%')
# -
# ## Plotando os dados de acordo com sua classificação
#Separar os dados para cada classe
class1Plot = []
class2Plot = []
for i in range(0, 1000):
if results[i] == 0:
class1Plot.append(dataFinal[0][i])
else:
class2Plot.append(dataFinal[0][i])
count, bins, ignored = plt.hist(class1Plot, 20, density=True, alpha = 0.6)
count2, bins2, ignored2 = plt.hist(class2Plot, 20, density=True, alpha = 0.6)
plt.title("Distribuição das amostras identificada pela sua classificação no Classificador Bayesiano")
plt.xlabel("Valor de x")
plt.ylabel("Probabilidade")
plt.grid(alpha=0.2)
plt.legend("12")
plt.show()
| ML_LISTA/Q2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bigbossthecat/ml-learn/blob/master/intro_to_pandas.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Ml2B40GVNF1H" colab_type="text"
# **Learning Panda**
# + id="FDAONRmdNRVb" colab_type="code" outputId="8a110cbf-d279-43a5-b712-975009e6b7ad" colab={"base_uri": "https://localhost:8080/", "height": 34}
from __future__ import print_function
import pandas as pd
pd.__version__
# + [markdown] id="3EutRgYjQr77" colab_type="text"
# **Creating a Series Object, i.e., a column and create a Table (Data frame) from multiple columns **
# + id="zMqqZ9UGQ38d" colab_type="code" outputId="c65be853-e95a-4f20-fc6d-17a38b14d348" colab={"base_uri": "https://localhost:8080/", "height": 142}
city_names = pd.Series (['San Francisco', 'San Jose', 'Sacramento'])
population = pd.Series ([852469,1015785, 485199])
pd.DataFrame ({'City Name': city_names, 'Population': population})
# + [markdown] id="2bJrChyaTJt_" colab_type="text"
# **Get California Housing Data**
# + id="VM8e0slmTQ_a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2176} outputId="dfeeb6f4-aa72-4ad4-f469-b260d7cf2a29"
california_housing_dataframe = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv", sep=",")
california_housing_dataframe.describe
# + id="PFxCAaAcUUKs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2176} outputId="f169060e-fa7b-4995-ada8-65522ae94dd6"
california_housing_dataframe.head
# + id="btwwHaOmUfyt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 396} outputId="b2e30410-174f-445d-fec5-f5d48df23c6c"
california_housing_dataframe.hist('housing_median_age')
| intro_to_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Xgboost GPU(env)
# language: python
# name: xgbgpuenv
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import preprocessing
import xgboost as xgb
from sklearn.metrics import mean_absolute_error
from datetime import date
import warnings
warnings.filterwarnings(action='ignore')
# set the seed of random number generator, which is useful for creating simulations
# or random objects that can be reproduced.
import random
SEED=3
random.seed(SEED)
np.random.seed(SEED)
# + active=""
# # Load Train Data
# train = pd.read_csv('../data/processed/train_aggr.csv', sep=';')
# -
# Load Train Data
train = pd.read_pickle('../data/processed/train_nochanel_uniqueidpos_x_envios_feateng.pkl')
train.shape
train['fecha_venta_norm'] = pd.to_datetime(train['fecha_venta_norm'])
train['fecha_venta_norm'] = train['fecha_venta_norm'].dt.date
# + active=""
# # Filtramos los meses que consideramos buenos para el entrenamiento (11 y 12)
# train = train[train.fecha_venta_norm.isin([ #date(2012, 11, 1),
# date(2012, 12, 1),
# #date(2013, 11, 1),
# date(2013, 12, 1),
# #date(2014, 11, 1)
# ])]
# + active=""
# predictors = ['id_pos','canal', 'competidores',
# 'ingreso_mediana', 'densidad_poblacional',
# 'pct_0a5', 'pct_5a9', 'pct_10a14', 'pct_15a19', 'pct_20a24',
# 'pct_25a29', 'pct_30a34', 'pct_35a39', 'pct_40a44', 'pct_45a49',
# 'pct_50a54', 'pct_55a59', 'pct_60a64', 'pct_65a69', 'pct_70a74',
# 'pct_75a79', 'pct_80a84', 'pct_85ainf', 'pct_bachelors',
# 'pct_doctorados', 'pct_secundario', 'pct_master', 'pct_bicicleta',
# 'pct_omnibus', 'pct_subtes', 'pct_taxi', 'pct_caminata',
# 'mediana_valor_hogar', 'unidades']
# -
predictors = [
'id_pos',
#'canal',
'competidores',
'ingreso_mediana',
'ingreso_promedio',
'densidad_poblacional',
'pct_0a5',
'pct_5a9',
'pct_10a14',
'pct_15a19',
'pct_20a24',
'pct_25a29',
'pct_30a34',
'pct_35a39',
'pct_40a44',
'pct_45a49',
'pct_50a54',
'pct_55a59',
'pct_60a64',
'pct_65a69',
'pct_70a74',
'pct_75a79',
'pct_80a84',
'pct_85ainf',
'pct_bachelors',
'pct_doctorados',
'pct_secundario',
'pct_master',
'pct_bicicleta',
'pct_omnibus',
'pct_subtes',
'pct_taxi',
'pct_caminata',
'mediana_valor_hogar',
#'unidades_despachadas_sum',
#'unidades_despachadas_max',
#'unidades_despachadas_min',
#'unidades_despachadas_avg',
#'cantidad_envios_max',
#'cantidad_envios_min',
#'cantidad_envios_avg',
#'num_cantidad_envios',
#'unidades_despachadas_sum_acum',
#'unidades_despachadas_sum_acum_3p',
#'unidades_despachadas_sum_acum_6p',
#'unidades_despachadas_max_acum',
#'unidades_despachadas_min_acum',
#'num_cantidad_envios_acum',
#'num_cantidad_envios_acum_3per',
#'num_cantidad_envios_acum_6per',
#'diff_dtventa_dtenvio',
'unidades_before',
'num_ventas_before',
'rel_unidades_num_ventas',
'unidades_acum',
'num_ventas_acum',
'countacum', 'unidades_mean',
'num_ventas_mean',
'unidades_2time_before',
'unidades_diff',
'month',
'diff_dtventa_dtventa_before',
'unidades_pend',
]
train = train[predictors]
# #### encode catvars
le = preprocessing.LabelEncoder()
classes = train['canal'].unique()
classes = [i for i in classes]
classes.append('NAN')
le.fit(classes)
np.save('../models/canal_le.npy', le.classes_)
train['canal'] = le.transform(train['canal'].values)
X, y = train.iloc[:,:-1],train.iloc[:,-1]
# #### Building final model
model = xgb.XGBRegressor(seed = SEED)
model.set_params(objective = 'reg:squarederror')
model.set_params(gpu_id = 0)
model.set_params(max_bin= 16)
model.set_params(tree_method='gpu_hist')
model.set_params(learning_rate = 0.01)
model.set_params(n_estimators = 273)
model.set_params(max_depth = 4)
model.set_params(min_child_weight = 5)
model.set_params(gamma = 0.0)
model.set_params(colsample_bytree = 0.9)
model.set_params(subsample = 0.8)
model.set_params(reg_alpha = 1)
model.fit(X, y)
y_pred = model.predict(X)
print("MAE unidades: ",mean_absolute_error(y, y_pred))
print("median unidades: ", np.median(y))
print("median unidades pred: ", np.median(y_pred))
import pickle
#save model
pickle.dump(model, open("../models/xgboost_013.pkl","wb"))
| notebooks/xgboost/build_xgboost_model_boorar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: CMIP6 2019.10
# language: python
# name: cmip6-201910
# ---
import numpy as np
import xarray as xr
import pandas as pd
import scipy
from scipy import signal
import scipy.stats as stats
import statsmodels.api as sm
import matplotlib.pyplot as plt
import numpy.random as random
#import cartopy
# Create toy data set in space-time [x,y,t]. We start with random noise in [x,y], and then multiply it by a weighting function to make it a bit more similar to geophysical data. Then, we create a linear trend along the time dimension, add some noise to simulate interannual variability, and apply the trend to each x,y gridpoint.
time_range = np.arange(0,10000)
x_dim = 50
y_dim = 100
time_dim = 10000
#Create initial x-y array from random noise
noise_space_uwt = random.random([x_dim,y_dim,time_dim])
#apply wts in the x-direction to better emulate geophysical space.
#We'll create a weighting function and then add a bit of noise
y_wts = np.concatenate((np.arange(0,y_dim/2),np.arange(y_dim/2,0,-1)))/(y_dim*0.1)
y_wts = y_wts + random.random([y_dim,])
noise_space = y_wts[None,:,None]*noise_space_uwt
#Plot average over y and time, we should see a triangle (values in the middle weigh more)
plt.plot(np.nanmean(np.nanmean(noise_space,axis=2),axis=0),np.arange(0,y_dim))
#Now, we add a first-order trend in time. First we create the equation (and add some random noise).
trend_slope = 0.01 #slope of trend
trend_int = 0.001 #intercept of trend
#We create a line with the slope and intercept we defined, and then we add some random noise
trend_eqn = trend_slope*np.arange(0,time_dim) + 5*random.random([time_dim,])
#Now, apply the trend to each spatial gridpoint
trend_3d = trend_eqn[None,None,:]*noise_space
data_detrend = signal.detrend(trend_3d,axis=2)
print(data_detrend.shape)
xg,yg = np.meshgrid(np.arange(0,x_dim),np.arange(0,y_dim))
# Now, plot the spatial average of our trended data. We should see the data increasing with time, but it should have some wiggles from the noise we added
plt.plot(np.nanmean(np.nanmean(trend_3d,axis=1),axis=0))
plt.title('Spatial average of our data with a linear trend')
# Remove the time mean from each grid cell. With geophysical data, we'd want to remove a seasonal average. For example, January temperatures in Texas are naturally more variable than July temperatures (January can be warmish or chilly, July is always hot). So, we want to account for this in our preprocessing (note that there are a lot of ways to do this; exactly what you'd do would depend on your scientific problem and the time resolution of your data). Here, we will just take a mean in time and remove that to create an anomaly time series.
time_mean = np.nanmean(trend_3d,axis=2)
time_anom = trend_3d - time_mean[:,:,None]
plt.plot(np.nanmean(np.nanmean(time_anom,axis=1),axis=0))
plt.title('Spatial average of our data, time-mean removed')
# Now, following Hawkins and Sutton (2012), we'll create a global mean. We're going to average our data in X and Y to simulate a global mean time series. With geophysical data, we'd typically want to include an area weighting, usually cos(latitude), since a grid cell at the equator is more spatially extensive than one at the pole. Here, we will just assume our grid cells have equal area.
time_mean = np.nanmean(np.nanmean(time_anom,axis=1),axis=0)
# Create smoothed version of our global (spatially averaged) time series. We want to capture the trend over our full time domain while smoothing out some of the wiggles that are happening on shorter timescales. Here, we'll approximate our smoothed time series with a first-order polynomial using numpy's polyfit function.
# We note that we're cheating a bit--we created this synthetic data with a linear trend, so we know that it's best approximated with a linear trend. We note that Hawkins and Sutton do this with a 4-th order polynomial, because this best captures the overall time series of global mean surface temperature. As you can imagine, you might want to use a different smoothing approach that better approximates your specific data.
#Fit a first-order polynomial to our spatially-averaged data
smoothed_coefs = np.polyfit(time_range,time_mean,1)
smoothed_vals = np.poly1d(smoothed_coefs)
#Apply polynomial over our time domain
data_smoothed = smoothed_vals(time_range)
#plot our spatially averaged data (blue), and our smoothed data (red)
plt.plot(time_range,time_mean)
plt.plot(time_range,data_smoothed,'r')
plt.legend(['raw data','smoothed data'])
plt.title('Comparison of raw and smoothed spatially-averaged data')
# Now, we use linear regression to explore the question, "What portion of the variability at each gridpoint can be explained by the trend?". We're assuming that our smoothed global mean time series is a good approximation of our trend, and we'll regress this time series (our predictor) on our data at each gridpoint. For this example, we'll use the OLS package in statsmodel, but you could use any package that can perform a straightforward linear regression. We want our regression model to output 2 things at each gridpoint--a regression coefficient, alpha (a slope), and the residuals, beta (an intercept).
# Note that if we apply our OLS model to our anomaly data, our alphas (slopes), should be approximately zero; this will change if we apply our OLS model to the raw data. Our betas (intercepts), should be the same either way. First we'll just do an example at a single gridpoint
time_anom.shape
#To get the intercept coefficients, we need to use add_constant
x_regress_example = sm.OLS(time_anom[33,2,:],sm.tools.add_constant(data_smoothed)).fit()
x_regress_example.params
#version with raw data
#x_regress_example = sm.OLS(trend_3d[33,2,:],sm.tools.add_constant(data_smoothed)).fit()
#x_regress_example.params
print("alpha: ",x_regress_example.params[0]," beta: ",x_regress_example.params[1])
print(x_regress_example.summary())
# Apply regression to each gridpoint. I am unsophisticated and tired so I am going to do two loops, the matlab way. Obviously this is dumb and can probably be avoided with some kind of unraveling function but we will fix this tomorrow.
alphas_all = np.zeros([x_dim,y_dim])
betas_all = np.zeros([x_dim,y_dim])
for i in np.arange(0,x_dim):
for j in np.arange(0,y_dim):
ij_regress = sm.OLS(time_anom[i,j,:],sm.tools.add_constant(data_smoothed)).fit()
alphas_all[i,j] = ij_regress.params[0]
betas_all[i,j] = ij_regress.params[1]
# Plot our alphas and our betas as a function of space
fig1, (ax1, ax2) = plt.subplots(1,2, figsize=(10,5))
xgrid,ygrid = np.meshgrid(np.arange(0,x_dim),np.arange(0,y_dim))
pc1 = ax1.pcolormesh(xgrid,ygrid,np.transpose(alphas_all),cmap='RdBu_r',vmin=-5e-15,vmax=5e-15)
fig1.colorbar(pc1,ax=ax1)
ax1.set_xlabel('X')
ax1.set_ylabel('Y')
pc2 = ax2.pcolormesh(xgrid,ygrid,np.transpose(betas_all),cmap='Reds',vmin=0,vmax=2)
fig1.colorbar(pc2,ax=ax2)
ax2.set_xlabel('X')
ax2.set_ylabel('Y')
ax1.set_title('alphas')
ax2.set_title('betas')
#print(xgrid.shape)
#print(alphas_all.shape)
# Now, let's estimate noise. We'll keep it basic and call our "noise" the standard deviation of our anomaly time series at each gridpoint. This should roughly represent how big the wiggles around the red line (smoothed data) are
# +
noise = np.nanstd(noise_space,axis=2) #should have dimensions of x,y
noiseplot = plt.pcolormesh(xgrid,ygrid,np.transpose(noise))
plt.colorbar(noiseplot)
plt.title('noise (standard deviation in time)')
print('spatially averaged standard deviation is ',np.nanmean(np.nanmean(noise)))
# -
# Now, we estimate our signal to noise ratio. The signal is estimated from our regression coefficients, and the noise is estimated above (standard dev). We'll do a really naive prediction here and just do an in-sample prediction. In a real analysis, we'd probably want to divide things into training and testing data.
#
data_test_to_predict = sm.add_constant(time_anom[4,63,2999:-1])
print("dependent variable has shape of ",time_anom[33,2,:].shape)
print("independent variable has shape of ",sm.add_constant(data_smoothed).shape)
print("regression model has shape of ",x_regress_example.params.shape)
print("data to predict has shape of ",data_test_to_predict.shape)
data_prediction = x_regress_example.predict(np.squeeze(data_test_to_predict))
print(data_prediction.shape)
# +
plt.plot(time_anom[4,33,2999:-1])
plt.plot(data_prediction,'r')
plt.legend(['original data','predicted from our regression'])
plt.title('comparison of actual data and prediction from regression')
# -
# Finally, we compare the signal (red line in the above plot) to the noise to see when/if S > N
sig_noise = data_prediction/noise[4,33]
plt.plot(sig_noise)
| notebooks/ToE_synthetic_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Evaluating Classifiers
# The first model predicts a high risk of cancer for 800 out of 1000 patients’ images in the test set.
# Of these 800 images 50 actually show signs of skin cancer.
# Hence, all problematic images are correctly identified.
#
# +
TP = 50 # True Positive
TN = 200 # True Negative
FP = 750 # False Positive
FN = 0
correct = TP + TN
total = 1000
correct / total
# -
precision = TP / (TP + FP)
precision # bad, makes a lot of false alarms
recall = TP / (TP + FN)
recall # really good, it catches all the cancers
# The second classifier categorizes 100 out of 1000 images into the high risk group.
# 40 of the 100 images show real signs of cancer. 10 images are not identified and falsely classified as low-risk.
#
#
# +
TP = 40
TN = 890
FN = 10 # we predict healthy but it is cancer
FP = 60 # we predict cancer but it is healthy
(TP + TN) / 1000
# -
precision = TP / (TP + FP)
precision
recall = TP / (TP + FN)
recall # good, but not that great for cancer detection
# +
# "everybody is healthy"
TP = 0
TN = 950
FN = 50 # we predict healthy but it is cancer
FP = 0 # we predict cancer but it is healthy
(TP + TN) / 1000
# -
recall = TP / (TP + FN)
recall
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.compose import ColumnTransformer
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler
from sklearn.pipeline import make_pipeline
# ### 1. Load the data
df = pd.read_csv('data/penguins_simple.csv', sep=';')
df.shape
# ### 2. Train-Test Split
X = df.iloc[:, 1:]
y = df['Species']
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, random_state=42)
# ### 3. Define a ColumnTransformer
trans = ColumnTransformer([
#('kristians_onehot', OneHotEncoder(sparse=False, handle_unknown='ignore'), ['Sex']),
#('kristians_scale', MinMaxScaler(), ['Body Mass (g)', 'Culmen Depth (mm)']),
('do_nothing', 'passthrough', ['Culmen Length (mm)']),
])
# ### 4. fit + transform training data
# +
trans.fit(Xtrain)
Xtrain_transformed = trans.transform(Xtrain) # result is a single numpy array
Xtrain_transformed.shape
# -
# ### 5. fit a LogReg model
# +
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(max_iter=1000)
model.fit(Xtrain_transformed, ytrain)
# -
# ## Evaluation Metrics
# +
from sklearn.metrics import accuracy_score, classification_report
ypred = model.predict(Xtrain_transformed)
acc = accuracy_score(ytrain, ypred)
round(acc, 3)
# -
print(classification_report(ytrain, ypred))
# +
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(model, Xtrain_transformed, ytrain)
# -
# ### 6. transform test data
Xtest_transform = trans.transform(Xtest)
Xtest_transform.shape
#test acc
ypred = model.predict(Xtest_transform)
acc = accuracy_score(ytest, ypred)
round(acc, 3)
# +
# training acc: 73%, test acc: 78.6%
# -
# ### Interpretation
#
# * training and test accuracy are good: everything OK
# * training and test accuracy are bad: not OK, model does not work (underfitting)
# * training accuracy good, test worse: not OK, model does not generalize (overfitting)
# * test accuracy better thatn training: strange, sometimes in small datasets (sampling bias)
# ### Advanced: ROC Curve
# very detailed but difficult to interpret
# +
from sklearn.metrics import roc_curve
# works for Titanic but not for Penguins
probs = model.predict_proba(Xtrain_transformed)
roc_curve(ytrain, probs)
# -
| week_02/EvaluatingClassifiers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Blue Crow Analytics Exercise
# Write a method that assigns players on each team to a tactical unit (ie defensive line, midfield line, etc).
# 1. Describe your general approach and any difficulties you encountered
# 2. Where does your approach work well and where does it break down?
# 3. How do you think it could be improved given more resources?
# 4. What are some applications of this model?
import json
import pandas as pd
import numpy as np
from mplsoccer import Pitch
# ## Read the match_data file
# 1. Get the pitch dimensions
# 2. Identify the IDs of the referees and the ball, to be filtered out of the strcutured_data
# 3. Keep the players information in a data frame
# +
#available match IDs:
#match_id = 4039
#match_id = 3749
match_id = 3518
#match_id = 3442
#match_id = 2841
#match_id = 2440
#match_id = 2417
#match_id = 2269
#match_id = 2068
match_fname = '../data/matches/{}/match_data.json'.format(match_id)
with open(match_fname, 'r') as f:
match_data = json.loads(f.read())
pitch_length = match_data['pitch_length']
pitch_width = match_data['pitch_width']
referee_object_ids = [ ref['trackable_object'] for ref in match_data['referees'] ]
objects_to_filter = [ ref['trackable_object'] for ref in match_data['referees'] ]
ball_object_id = match_data['ball']['trackable_object']
objects_to_filter.append(ball_object_id)
print('Pitch length, width: {},{}'.format(pitch_length, pitch_width))
print('Ball object id:{}'.format(ball_object_id))
print('Referee object ids: {}'.format(referee_object_ids))
print('Object ids to filter: {}'.format(objects_to_filter))
# -
match_players = pd.json_normalize(match_data['players'])
# ## Read the structured_data file
# 1. Keep a subset of the json fields
# 2. Drop the frames with no info
# 3. Filter out the referees and ball info
# 4. **Flip the sign of the x coordinates from the second half of the game:** needed for accurate counting of each player's position in each tactical zone (done later)
# 5. Join the players info from match_data with their postional info from strutured_data
struct_fname = '../data/matches/{}/structured_data.json'.format(match_id)
with open(struct_fname, 'r') as f:
struct_data = json.loads(f.read())
struct_df = pd.json_normalize(struct_data,'data', ['period', 'frame', 'time', ['possession', 'trackable_object'], ['possession', 'group']])
struct_df.dropna(subset=['period', 'trackable_object'], inplace=True)
struct_df['trackable_object'] = struct_df['trackable_object'].astype(int)
for id_to_drop in objects_to_filter:
struct_df = struct_df[ struct_df['trackable_object'] != id_to_drop]
# Fold second half coordinates onto first half
mult = [1 if period == 1 else -1 for period in struct_df['period']]
struct_df['x'] = struct_df['x'] * mult
struct_df.head()
# ### Join the players info from match_data with their postional info from strutured_data
# This step is not strictly needed, but for this exploratory exercise it's helpful to have this data all in one dataframe for visual inspection and quicker insight. More efficient choices exist if one were to automate this notebook into a processing pipeline.
struct_df = struct_df.merge(match_players, on='trackable_object', how='left')
struct_df.head()
# ## Find the direction of play
# * **Why?** The algorithm divides the pitch in thirds, but to correctly distinguish the attacking third from the defensive third, we need to know the direction of play of each team. Since this information is not in the match data file we need to infer it.
# * **How?** The goalkeeper position determines the direction of play, because the goalkeeper almost never passes the midfield line. Note that we have previously folded the coordinates of the player positions from the second half onto the first half, so the direction of play does not change at half time for the purpose of this analysis.
# +
# Find the direction of play for the two teams
# Needed to distinguish attacking line from defensive line
keepers_df = struct_df[struct_df['player_role.name'] == 'Goalkeeper']
team1_id, team2_id = keepers_df['team_id'].unique() # there are only 2 teams per game, so OK to do this
team1_keeper = keepers_df[ keepers_df['team_id'] == team1_id ]
keeper1_position = team1_keeper.x.median()
if keeper1_position < 0:
team_direction_of_play = {team1_id: 'left_to_right', team2_id: 'right_to_left'}
else:
team_direction_of_play = {team2_id: 'left_to_right', team1_id: 'right_to_left'}
#Store the direction of play in the dataframe: this is not memory efficient, but it's OK for this interactive exploration
struct_df['team_direction'] = [ team_direction_of_play[team_id] for team_id in struct_df['team_id']]
struct_df.head()
# -
# ## Assign each player to one of three tactical units
# 1. Using 3 tactical units for this exercise: **'defensive_line', 'midfield_line', 'attacking_line'**
# 2. Each unit correponds to 1/3 of the pitchzones for tis exercise
# 3. Count how many times a player visits each of the tactical unit zones
# 4. Assign the player to the tactical unit with the most counts
tactical_unit_zones = ['defensive_line', 'midfield_line', 'attacking_line']
other_direction_zones = tactical_unit_zones.copy()
other_direction_zones.reverse()
label_dict = {'left_to_right': tactical_unit_zones, 'right_to_left': other_direction_zones}
label_dict
# +
pitch_thirds = list(np.array([pitch_length/3, pitch_length *2/3]) - pitch_length/2)
bins = [-pitch_length/2] + pitch_thirds + [pitch_length/2]
counts_dict = {'trackable_object': [], 'defensive_line': [], 'midfield_line': [], 'attacking_line': []}
byobject = struct_df.groupby('trackable_object')
for trackable_object, data in byobject:
this_player_direction = data['team_direction'].unique()[0]
labels = label_dict[this_player_direction]
counts = pd.cut(data.x, bins=bins, labels=labels).value_counts().to_dict()
for label in labels:
counts_dict[label].append(counts[label])
counts_dict['trackable_object'].append(trackable_object)
actions_per_unit = pd.DataFrame(counts_dict)
# This line is where we pick the tactical zone corresponding to the maximum-count column
actions_per_unit['tactical_unit'] = actions_per_unit[tactical_unit_zones].idxmax(axis=1)
# -
# Joining the result with the players info from match_data for visual inspection
columns_to_pick = ['trackable_object', 'player_role.name', 'last_name', 'first_name']
result = actions_per_unit.merge(match_players[columns_to_pick], on='trackable_object', how='left')
result
# A little sanity check: does the assigned tactical zone make sense compared to the player roles from the match_data file?
tmp = pd.DataFrame(result.groupby(['tactical_unit', 'player_role.name'])['tactical_unit'].count())
tmp.columns = ['counts']
tmp
# ## Make some plots
# Create some plots for the presentation
# Make sure the players selected below are part of the match analyzed above
# +
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_theme(context='talk')
import matplotlib.ticker as mtick
def make_player_kde_plot(player_last_name, data_df):
"""Create a kde plot of player positions."""
player_df = data_df[data_df['last_name'] == player_last_name]
pitch = Pitch(pitch_type='skillcorner', pitch_length=pitch_length, pitch_width=pitch_width, line_color='black', line_zorder=2)
fig,ax = pitch.draw()
kde = pitch.kdeplot(player_df.x, player_df.y, ax=ax,
# shade using 100 levels so it looks smooth
shade=True, levels=100,
# shade the lowest area so it looks smooth
# so even if there are no events it gets some color
shade_lowest=True,
cut=4) # extended the cut so it reaches the bottom edge
def make_player_zones_barplot(player_last_name, data_df, names=tactical_unit_zones):
"""Make a bar plot of counts of a player's visits to the tactical unit zones."""
player_df = data_df[data_df['last_name'] == player_last_name] #this is actually just one row from the result data frame
tmp = player_df[names].T
tmp = tmp/tmp.sum() * 100
title_str = '{}: Distribution of tactical units'.format(player_last_name)
ax = tmp.plot(kind='bar',rot=45, legend=False, title=title_str)
ax.yaxis.set_major_formatter(mtick.PercentFormatter())
# -
#Get an empty pitch for fun
pitch = Pitch(pitch_color='#aabb97', line_color='white',
stripe_color='#c2d59d', stripe=True) # optional stripes
fig, ax = pitch.draw()
make_player_kde_plot('Lukaku', struct_df)
#make_player_kde_plot('de Bruyne', struct_df)
make_player_zones_barplot('Lukaku', result, names=other_direction_zones)
#make_player_zones_barplot('de Bruyne', result)
make_player_kde_plot('Barella', struct_df)
make_player_zones_barplot('Barella', result, names=other_direction_zones)
make_player_kde_plot('Skriniar', struct_df)
make_player_zones_barplot('Skriniar', result, names=other_direction_zones)
| bluecrow/tactical_unit_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring Canadian naturalization records, 1915 to 1946
#
# This notebook explores data harvested from the Library and Archives Canada database of [Naturalization Records, 1915-1946](http://www.bac-lac.gc.ca/eng/discover/immigration/citizenship-naturalization-records/naturalized-records-1915-1951/Pages/introduction.aspx).
#
# To create this dataset, we first harvested records where the `country` was listed as 'China'. However, we realised that the wives and children of a naturalised man weren't assigned a country value and so will be missing from the harvested data. We attempted to overcome this by adding in records of what appeared to be family members, but this data might be inaccurate or incomplete.
#
# The harvested data was saved as a [CSV file](lac-naturalisations-china-with-families.csv).
#
# For full details of the harvesting process, see the [LAC section](https://glam-workbench.net/lac/) of the GLAM Workbench.
import pandas as pd
import altair as alt
df = pd.read_csv('lac-naturalisations-china-with-families.csv')
df.head()
# How many records are there?
df.shape[0]
# How many records are relations (ie wives and children)?
df['relation'].value_counts()
# Some years are recorded as a range – let's put the first year mentioned into a separate field for aggregation.
df['year_int'] = df['year'].str.slice(0,4)
# Let's look at the number of records per year.
df['year_int'].value_counts()
# Let's include the `relation` field as well, so we can highlight women and children.
df['relation'].fillna('Not recorded', inplace=True)
year_counts = df.value_counts(['year_int', 'relation']).to_frame().reset_index()
year_counts.columns = ['year', 'relation', 'count']
alt.Chart(year_counts).mark_bar(size=15).encode(
x=alt.X('year:Q', axis=alt.Axis(format='c')),
y=alt.Y('count:Q', stack=True),
color='relation:N',
tooltip=['year', 'relation', 'count']
).properties(width=700)
| lac_canada_naturalisations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Dog():
def __init__(self,mybreed):
self.breed=mybreed
def bark(self):
print(self.breed)
my_dog=Dog(mybreed="Lab")
type(my_dog)
my_dog.breed
my_dog.bark()
class a():
def text(self):
print("hello")
b=a()
b
type(b)
class circle():
#class object attribute
pi=3.14
def __init__(self,radius):
self.radius=radius
def circumference(self):
return 2*self.pi*self.radius
a=circle(2)
type(a)
a.circumference()
class Animal():
def __init__(self):
print("animal created")
def identity(self):
print("i am an animal")
class Dog(Animal):
def __init__(self):
print("I am a dog")
a=Dog()
type(a)
a.identity()
class Bank():
def __init__(self,owner,balance):
self.owner=owner
self.balance=balance
def deposit(self):
amt=int(input("enter amt to be deposited"))
self.balance=amt+self.balance
print(f"The balance now is{self.balance}")
def withdraw(self):
cash=int(input("Enter amt to be withdrawn"))
if(cash>self.balance):
print("Low balance!")
else:
self.balance=self.balance-cash
print(f"the balance now is {self.balance}")
b=Bank("h",1500)
b.withdraw()
# b.deposit()
# b.owner
while True:
try:
a=int(input("enter number"))
print(a**2)
except:
print("That is not an integer")
else:
print("Thank you")
break
finally:
print("All done")
#
def decorator(original):
def new():
print("Before code")
original()
print("After code")
return new
def original():
print("hello")
original=decorator(original)
original()
for a in range(10):
print(a)
def f(n):
for a in range(n):
yield a**5
for a in f(3):
print(a)
f(3)
a=f(3)
next(a)
next(a)
next(a)
a=iter("hello")
next(a)
next(a)
next(a)
next(a)
next(a)
for a in range(3,7,2):
print(a)
a=[1,2]
a.append(2)
a
| Basics_Python/python5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dilaragokay/3D-point-capsule-networks/blob/colab/Perturbation_3DCapsNet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="3_zvdaH4-drr"
# # 1. Install dependencies
# + id="C4kuG295KFjs"
# !sudo apt-get install libhdf5-dev
# !sudo pip install h5py
# !pip install open3d
# !pip install gdown
# + id="s0zoVlKAKcXx"
# %cd models/nndistance
# !python build.py install
# + [markdown] id="imynwMYfCMc5"
# # 2. Install ShapeNetPart Dataset
# + id="wsXvYT4eKu9n"
# %cd ../../dataset
# !bash download_shapenet_part16_catagories.sh
# + [markdown] id="yg_fYo1lpoY7"
# # 3. Download checkpoints
# + id="GF5nr99DNPg-"
# !mkdir ../checkpoints
# %cd ../checkpoints
# Checkpoint link is obtained from https://github.com/yongheng1991/3D-point-capsule-networks/blob/master/README.md
# !gdown https://drive.google.com/uc?id=1so0OiVPS93n7Ed36yiMHQb-u0f1zJSzJ
# + [markdown] id="zISG0CBzqOnj"
# # 4. Perturb example point cloud
# <img src="https://i.ibb.co/jwGwVSV/Screenshot-2020-11-05-at-00-58-39.png">
#
# *Image source: [1]*
#
# * I picked one object (plane) for the sake of simplicity.
#
# * After dynamic routing, 64 latent capsules are obtained. Since some of the capsules seem to have irregular points, plus, visualizing all of the capsules is not feasible, I've picked 3 capsules which seem to represent the object well. These capsules represent **wings, rudder, and bottom**.
# * Each capsule is a 64x1 vector. I've picked 3 features from these capsules. I've picked the ones on position 10, 20, and 30 (with no particular reason). [2] perturbs features in the capsules within [-0.5, 0.5]. I chose {-0.25, 0.25} in order not to have too many combinations to visualize.
# * At each attempt, I've changed one of the variables above and kept the rest constant. In a nutshell,
#
# ```
# for capsule in capsule_set:
# for feature in feature_set:
# for amount in perturbations:
# perturb the feature in the capsule by amount then save
# ```
#
# Changes are in `viz_perturbation.py`.
#
#
# + id="57_Hi34QNlv6"
# %cd ../../mini_example/AE/
# + id="4n5kBYWuN4Iz" colab={"base_uri": "https://localhost:8080/"} outputId="c3a2336a-fbe0-4032-907a-f6b283016f5d"
# !python viz_perturbation.py --model ../../checkpoints/shapenet_part_dataset_ae_200.pth
# + [markdown] id="7dieQmcHwXdm"
# ## 5. Show results in 3D plot
# + id="HcDtTTKitm_j"
import open3d as o3d
import plotly.graph_objects as go
from plotly.subplots import make_subplots
import numpy as np
import glob
# + id="n2efaqLXtx9h"
pcd_files = glob.glob('*.pcd')
# + id="FuBciOvzuHmM"
dictionary = {}
for f in pcd_files:
key = '_'.join(f.split('_')[:2])
if key in dictionary:
dictionary[key].append(f)
else:
dictionary[key] = [f]
for key in dictionary:
dictionary[key] = sorted(dictionary[key])
# + [markdown] id="f3EZP_Pxw6X5"
# ### Visualize the first point cloud
# + id="85nHXknjPfqo"
def plot_perturbations(minus_pert, zero_pert, plus_pert, specs):
# Initialize figure with 9 3D subplots
fig = make_subplots(
rows=3,
cols=1,
specs=[[{"type": "scene"}], [{"type": "scene"}], [{"type": "scene"}]],
shared_xaxes=True,
subplot_titles=("-0.25 perturbation", "No perturbation", "+0.25 perturbation"))
# Generate data
pcd = o3d.io.read_point_cloud(minus_pert)
x1 = np.asarray(pcd.points)[:, 0]
y1 = np.asarray(pcd.points)[:, 1]
z1 = np.asarray(pcd.points)[:, 2]
c1 = np.asarray(pcd.colors)
pcd = o3d.io.read_point_cloud(zero_pert)
x2 = np.asarray(pcd.points)[:, 0]
y2 = np.asarray(pcd.points)[:, 1]
z2 = np.asarray(pcd.points)[:, 2]
c2 = np.asarray(pcd.colors)
pcd = o3d.io.read_point_cloud(plus_pert)
x3 = np.asarray(pcd.points)[:, 0]
y3 = np.asarray(pcd.points)[:, 1]
z3 = np.asarray(pcd.points)[:, 2]
c3 = np.asarray(pcd.colors)
# adding point clouds to subplots.
fig.add_trace(go.Scatter3d(x=x1, y=y1, z=z1,mode='markers',
marker=dict(
size=2,
color=c1, # set color to an array/list of desired values
opacity=0.4
)),
row=1, col=1)
fig.add_trace(go.Scatter3d(x=x2, y=y2, z=z2, mode='markers',
marker=dict(
size=2,
color=c2, # set color to an array/list of desired values
opacity=0.4
)),
row=2, col=1)
fig.add_trace(go.Scatter3d(x=x3, y=y3, z=z3, mode='markers',
marker=dict(
size=2,
color=c3, # set color to an array/list of desired values
opacity=0.4
)),
row=3, col=1)
fig.update_layout(
title_text=specs,
height=1000,
width=1000
)
fig.show()
# + id="-3rV7cV09me1"
key = list(dictionary.keys())[0] # key for the first point cloud
point_clouds = dictionary[key]
num_to_color = {
'29': 'red',
'40': 'green',
'16': 'blue'
}
num_to_name = {
'29': 'rudder',
'40': 'wings',
'16': 'bottom'
}
i = 0
# + [markdown] id="o0_vWP7K9zpL"
# Run the following cell as long as there are more combinations. All point clouds are not visualized at once as it can crash the browser.
# + id="MdTApmTz3MPv" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3710e349-d275-442e-bd18-faae55b1c788"
if i < len(point_clouds):
minus = point_clouds[i]
plus = point_clouds[i+1]
zero = point_clouds[i+2]
name_splitted = plus.split('_')
caps_no = name_splitted[2][4:]
feat_no = name_splitted[3][4:]
amount = name_splitted[5][:-4]
specs = "Feature #{} of the capsule for {}. Capsule is shown in {}".format(
feat_no,
num_to_name[caps_no],
num_to_color[caps_no])
plot_perturbations(minus, zero, plus, specs)
i+=3
# + [markdown] id="gFoyrNSsJ06P"
# # 6. Visualize precaptured images from 3 different viewpoints
#
# In images below, each 3x3 figure shows how changing certain feature in a certain capsule affects view from side, front, and top of the plane. In short, each subfigure has been organized the following way.
#
# | | side view | front view | top view |
# |-------|-----------|------------|----------|
# | -0.25 | img | img | img |
# | 0 | img | img | img |
# | 0.25 | img | img | img |
#
# Note that these figures have been captured by hand so not all these images have been captured from the same angle i.e. some images might be slightly rotated.
#
# ## 6.1. Change in capsule that corresponds to the bottom of the plane (blue points)
# ### 6.1.1. Perturbation on feature #10
#
# <img src="https://drive.google.com/uc?export=view&id=1ZJpBLMVfL3R1pLoge1ZiDrXN2OJiF_OX">
#
# ### 6.1.2. Perturbation on feature #20
#
# <img src="https://drive.google.com/uc?export=view&id=1tVz8mhOhHFsJbvZR3XnUzLSHxvHvzXws">
#
# ### 6.1.3. Perturbation on feature #30
#
# <img src="https://drive.google.com/uc?export=view&id=1fhTCQXEP25SrFT1kMR0ZrKoXFenRkYjC">
#
# ## 6.2. Change in capsule that corresponds to the rudder of the plane (red points)
# ### 6.2.1. Perturbation on feature #10
#
# <img src="https://drive.google.com/uc?export=view&id=1uELPomS0IZw9IqrRoF5uCui6_uQdd8nr">
#
# ### 6.2.2. Perturbation on feature #20
#
# <img src="https://drive.google.com/uc?export=view&id=1yaLi7Ed7mI9ttixmxcAMcdlWub421vrd">
#
# ### 6.2.3. Perturbation on feature #30
#
# <img src="https://drive.google.com/uc?export=view&id=1jsvh0CbT263KbtarXo8zFAdxwdpwvIpr">
#
# ## 6.3. Change in capsule that corresponds to the wings of the plane (green points)
# ### 6.3.1. Perturbation on feature #10
#
# <img src="https://drive.google.com/uc?export=view&id=15VStNiDnyovnnWYCX9xQfs9hkdPHUul2">
#
# ### 6.3.2. Perturbation on feature #20
#
# <img src="https://drive.google.com/uc?export=view&id=1Z_kIv7yCX5inAnlQeQ5gxD06CATFjgkd">
#
# ### 6.3.3. Perturbation on feature #30
#
# <img src="https://drive.google.com/uc?export=view&id=1C87QsMgY0I8lsFIvaaU2PyJSm-vTU9kT">
# + [markdown] id="PU1HsYtW3A0G"
# # 7. Observations (w/ [@evinpinar](https://github.com/evinpinar))
#
# * Changes in feature #30 of rudder,
# * Changes in feature #20 of wing,
# * Changes in feature #30 of wing
# causes rotation and translation.
# + [markdown] id="wuBDV-ahs2Tk"
# # Reference
#
# [1] Zhao, Yongheng, et al. "3D point capsule networks." *Proceedings of the IEEE conference on computer vision and pattern recognition*. 2019.
# [2] Sabour, Sara, <NAME>, and <NAME>. "Dynamic routing between capsules." *Advances in neural information processing systems*. 2017.
| Perturbation_3DCapsNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import os
import sys
import copy
import warnings
import cProfile
from time import time
from astropy.stats import sigma_clip
from astropy.table import Table, Column, vstack
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as patches
from asap import io
from asap import smf
from asap import config
from asap import fitting
from asap import ensemble
from asap import plotting
from asap.parameters import AsapParams
from asap.likelihood import ln_likelihood, ln_probability
from asap.predictions import (predict_mstar_basic, predict_smf, predict_mhalo,
predict_dsigma_profiles, make_model_predictions)
plt.rc('text', usetex=True)
# -
# ## Corner and traceplot of the model
#
# * This is the default `A.S.A.P` model
# * Three stages burn-in with 256 walkers using the "Snooker" moves, each stage has 250 steps.
# * Using the walker position with the best likelihood as initial positions for the next stage.
# * Final sampling process has 400 steps.
# * **SMF**: use the covariance matrix of SMFs
# * **DeltaSigma** profiles: fit the radius between 0.15 to 15 Mpc.
# +
test_dir = '../model/'
model_str = 'final_default'
# The configuration file
config_file = os.path.join(test_dir, 'asap_%s.yaml' % model_str)
# The results from the 3-stage burn-in results
burnin_file_1 = os.path.join(test_dir, 'asap_%s_burnin_1.npz' % model_str)
burnin_file_2 = os.path.join(test_dir, 'asap_%s_burnin_2.npz' % model_str)
burnin_file_3 = os.path.join(test_dir, 'asap_%s_burnin_3.npz' % model_str)
# The results of the final sampling process
result_file = os.path.join(test_dir, 'asap_%s_sample.npz' % model_str)
# Initialize the model, load the data
cfg, params, obs_data, um_data = fitting.initial_model(config_file, verbose=True)
# Load the burn-in results
(mod_burnin_samples_1,
mod_burnin_chains_1,
mod_burnin_lnprob_1,
mod_burnin_best_1, _, _) = io.load_npz_results(burnin_file_1)
(mod_burnin_samples_2,
mod_burnin_chains_2,
mod_burnin_lnprob_2,
mod_burnin_best_2, _, _) = io.load_npz_results(burnin_file_2)
#(mod_burnin_samples_3,
# mod_burnin_chains_3,
# mod_burnin_lnprob_3,
# mod_burnin_best_3, _, _) = io.load_npz_results(burnin_file_3)
mod_burnin_chains = np.concatenate([
mod_burnin_chains_1, mod_burnin_chains_2], axis=1)
# Load in the final sampling results
(mod_result_samples,
mod_result_chains,
mod_result_lnprob,
mod_result_best, _, _) = io.load_npz_results(result_file)
_, n_step, n_dim = mod_result_chains.shape
print(np.nanmax(mod_result_lnprob), mod_result_best)
mod_result_best = np.nanmean(mod_result_chains[:, -int(n_step * 0.1):, :].reshape([-1, n_dim]), axis=0)
print(mod_result_best)
# -
# ### Corner plot
# +
params_label = [r'$a$', r'$b$', r'$c$', r'$d$',
r'$f_{\rm ins}$', r'$A_{\rm exs}$', r'$B_{\rm exs}$']
params_range = [(0.585, 0.599), (11.831, 11.854),
(-0.024, 0.007), (-0.005, 0.0039),
(0.629, 0.679),
(-0.22, -0.172), (0.301, 0.41)]
title_fmt = '.3f'
mod_samples_use = mod_result_chains[:, 1000:, :].reshape([-1, n_dim])
mod_corner = plotting.plot_mcmc_corner(
mod_samples_use, params_label, truths=mod_result_best, truth_color='skyblue',
**{'title_fmt': title_fmt, 'ranges': params_range, 'plot_datapoints': False})
# -
# ### Trace plot
mod_trace = plotting.plot_mcmc_trace(
mod_result_chains, params_label,
mcmc_best=mod_result_best, mcmc_burnin=mod_burnin_chains,
burnin_alpha=0.12, trace_alpha=0.15)
# ### Quick check of the SMFs
parameters = mod_result_best
parameters = np.nanmean(mod_result_chains[:, -int(n_step * 0.1):, :].reshape([-1, n_dim]), axis=0)
# +
# Predict the stellar mass in inner and outer apertures
logms_inn, logms_tot, sig_logms, mask_use = predict_mstar_basic(
um_data['um_mock'], parameters, min_logms=10.6,
logmh_col=cfg['um']['logmh_col'], min_scatter=cfg['um']['min_scatter'],
pivot=cfg['um']['pivot_logmh'])
# Predict the SMFs and DeltaSigma profiles
um_smf_tot, um_smf_inn, um_dsigma = make_model_predictions(
mod_result_best, cfg, obs_data, um_data)
# Check the likelihood for SMF and DeltaSigma profiles
lnlike_smf, lnlike_dsigma = ln_likelihood(
mod_result_best, cfg, obs_data, um_data, sep_return=True)
print("# ln(Likelihood) for SMFs : %8.4f" % lnlike_smf)
print("# ln(Likelihood) for DSigma : %8.4f" % lnlike_dsigma)
# +
um_table = Table(um_data['um_mock'][mask_use])
um_table.add_column(Column(data=logms_inn, name='logms_inn_asap'))
um_table.add_column(Column(data=logms_tot, name='logms_tot_asap'))
um_table.write('../data/asap_smdpl_mock_default.fits', overwrite=True)
# +
um_smf_tot_all = smf.get_smf_bootstrap(logms_tot, cfg['um']['volume'],
10, 10.9, 12.4, n_boots=1)
mod_smf_plot = plotting.plot_mtot_minn_smf(
obs_data['smf_tot'], obs_data['smf_inn'], obs_data['mtot'], obs_data['minn'],
um_smf_tot, um_smf_inn, logms_tot, logms_inn, obs_smf_full=obs_data['smf_full'],
um_smf_tot_all=um_smf_tot_all, not_table=True, x_label='Max', y_label='10\ kpc')
# -
# ### Save the figures
# +
mod_corner.savefig('fig/fig3_corner_default.pdf', dpi=130)
mod_trace.savefig('fig/fig3_trace_default.png', dpi=120)
mod_smf_plot.savefig('fig/fig3_smf_default.png', dpi=120)
# -
| note/fig3_default.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
with open("../rpn_ret.pkl", "rb") as f:
rpn = pickle.load(f)
rpn.keys()
ex = rpn["rois_fpn2"][0]
rpn["rois"][511] in rpn["rois_fpn2"]
idx = rpn['rois_idx_restore_int32']
N2 = len(rpn["rois_fpn2"])
count = 0
for x in rpn["rois"][:N2]:
if x in rpn["rois_fpn2"]:
count+=1
count = 0
N3 = N2 + len(rpn["rois_fpn3"])
for x in rpn["rois"][N2:N3]:
if x in rpn["rois_fpn3"]:
count+=1
len(rpn["rois_fpn3"]), count
# +
count = 0
N4 = N3 + len(rpn["rois_fpn4"])
for x in rpn["rois"][N3:N4]:
if x in rpn["rois_fpn4"]:
count+=1
# -
len(rpn["rois_fpn4"]), count
| notebooks/Rpn_ret debuggin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Specifying Intent in Lux
# Lux provides a flexible language for communicating your analysis intent to the system, so that Lux can provide better and more relevant recommendations to you. In this tutorial, we will see different ways of specifying the intent, including the attributes and values that you are interested or not interested in, enumeration specifiers, as well as any constraints on the visualization encoding.
#
# The primary way to set the current intent associated with a dataframe is by setting the `intent` property of the dataframe, and providing a list of specification as input. We will first describe how intent can be specified through convenient shorthand descriptions as string inputs, then we will describe advance usage via the `lux.Clause` object.
#
# ## Basic descriptions
# Continuing with our college dataset example from earlier,
import pandas as pd
import lux
# Collecting basic usage statistics for Lux (For more information, see: https://tinyurl.com/logging-consent)
lux.logger = True # Remove this line if you do not want your interactions recorded
df = pd.read_csv("../data/college.csv")
lux.config.default_display = "lux"
# ### Specifying attributes of interest
# You can indicate that you are interested in an attribute, let's say `AverageCost`.
df.intent = ['AverageCost']
df
# You might be interested in multiple attributes, for instance you might want to look at both `AverageCost` and `FundingModel`. When multiple clauses are specified, Lux applies all the clauses in the intent and searches for visualizations that are relevant to `AverageCost` **and** `FundingModel`.
#
df.intent = ['AverageCost','FundingModel']
df
# Let's say that in addition to `AverageCost`, you are interested in the looking at a list of attributes that are related to different financial measures, such as `Expenditure` or `MedianDebt`, and how they breakdown with respect to `FundingModel`.
#
# You can specify a list of desired attributes separated by the `|` symbol, which indicates an `OR` relationship between the list of attributes. If multiple clauses are specified, Lux automatically create combinations of the specified attributes.
possible_attributes = "AverageCost|Expenditure|MedianDebt|MedianEarnings"
df.intent = [possible_attributes,"FundingModel"]
df
# Alternatively, you could also provide the specification as a list:
possible_attributes = ['AverageCost','Expenditure','MedianDebt','MedianEarnings']
df.intent = [possible_attributes,"FundingModel"]
df
# ### Specifying values of interest
# In Lux, you can also specify particular values corresponding to subsets of the data that you might be interested in. For example, you may be interested in only colleges located in New England.
#
df.intent = ["Region=New England"]
df
# You can also specify multiple values of interest using the same `|` notation that we saw earlier. For example, you can compare the median debt of students from colleges in New England, Southeast, and Far West.
df.intent = ["MedianDebt","Region=New England|Southeast|Far West"]
df
# Note that since there are three different visualizations that is generated based on the intent, we only display these possible visualization, rather than the recommendations.
df.clear_intent()
# #### Note: Applying Filters v.s. Expressing Filter Intent
# You might be wondering what is the difference between specifying values of interest through the intent in Lux versus applying a filter directly on the dataframe through Pandas. By specifying the intent directly via Pandas, Lux is not aware of the specified inputs to Pandas, so these values of interest will not be reflected in the recommendations.
df[df["Region"]=="New England"]
# Specifying the values through `set_intent` tells Lux that you are interested in colleges in New England. In the resulting Filter action, we see that Lux suggests visualizations in other `Region`s as recommendations.
df.intent = ["Region=New England"]
df
# So while both approaches applies the filter on the specified visualization, the subtle difference between *applying* a filter and *indicating* a filter intent leads to different sets of resulting recommendations. In general, we encourage using Pandas for filtering if you are certain about applying the filter (e.g., a cleaning operation deleting a specific data subset), and specify the intent through Lux if you might want to experiment and change aspects related to the filter in your analysis.
# ### Advanced intent specification through `lux.Clause`
# The basic string-based description provides a convenient way of specifying the intent. However, not all specification can be expressed through the string-based descriptions, more complex specification can be expressed through the `lux.Clause` object. The two modes of specification is essentially equivalent, with the Parser parsing the `description` field in the `lux.Clause` object.
# #### Specifying attributes or values of interest
# To see an example of how lux.Clause is used, we rewrite our earlier example of expressing interest in `AverageCost` as:
df.intent = [lux.Clause(attribute='AverageCost')]
# Similarly, we can use `lux.Clause` to specify values of interest:
df.intent = ['MedianDebt',
lux.Clause(attribute='Region',filter_op='=', value=['New England','Southeast','Far West'])]
# Both the `attribute` and `value` fields can take in either a single string or a list of attributes to specify items of interest. This example also demonstrates how we can intermix the `lux.Clause` specification alongside the basic string-based specification for convenience.
# #### Adding constraints to override auto-inferred details
# So far, we have seen examples of how Lux takes in a loosely specified intent and automatically fills in many of the details that is required to generate the intended visualizations. There are situations where the user may want to override these auto-inferred values. For example, you might be interested in fixing an attribute to show up on a particular axis, ensuring that an aggregated attribute is summed up instead of averaged by default, or picking a specific bin size for a histogram. Additional properties specified on lux.Clause acts as constraints to the specified intent.
# <ins>Fixing attributes to specific axis channels<ins>
# As we saw earlier, when we set `AverageCost` as the intent, Lux generates a histogram with `AverageCost` on the x-axis.
# While this is unconventional, let's say that instead we want to set `AverageCost` to the y axis. We would specify this as additional properties to constrain the intent clause.
df.intent = [lux.Clause(attribute='AverageCost', channel='y')]
df
# <ins>Changing aggregation function applied<ins>
# We can also set constraints on the type of aggregation that is used. For example, by default, we use `mean` as the default aggregation function for quantitative attributes.
df.intent = ["HighestDegree","AverageCost"]
df
# We can override the aggregation function to be `sum` instead.
#
df.intent = ["HighestDegree",lux.Clause("AverageCost",aggregation="sum")]
df
# The possible aggregation values are the same as the ones supported in Pandas's [agg](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.agg.html) function, which can either be a string shorthand (e.g., "sum", "count", "min", "max", "median") or as a numpy aggregation function.
#
# For example, we can change the aggregation function to be the point-to-point value ([np.ptp](https://numpy.org/doc/stable/reference/generated/numpy.ptp.html)) by inputting the numpy function.
import numpy as np
df.intent = ["HighestDegree",lux.Clause("AverageCost",aggregation=np.ptp)]
df
# ### Specifying wildcards
# Let's say that you are interested in *any* attribute with respect to `AverageCost`. Lux support *wildcards* (based on [CompassQL](https://idl.cs.washington.edu/papers/compassql/) ), which specifies the enumeration of any possible attribute or values that satisfies the provided constraints.
df.intent = ['AverageCost',lux.Clause('?')]
df
# The space of enumeration can be narrowed based on constraints. For example, you might only be interested in looking at scatterplots of `AverageCost` with respect to quantitative attributes. This narrows the 15 visualizations that we had earlier to only 9 visualizations now, involving only quantitative attributes.
df.intent = ['AverageCost',lux.Clause('?',data_type='quantitative')]
df
# The enumeration specifier can also be placed on the value field. For example, you might be interested in looking at how the distribution of `AverageCost` varies for all possible values of `Geography`.
#
df.intent = ['AverageCost','Geography=?']
# OR
df.intent = ['AverageCost',lux.Clause(attribute='Geography',filter_op='=',value='?')]
df
| tutorial/1-specifying-intent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BrunaKuntz/Python-Curso-em-Video/blob/main/Mundo03/Desafio100.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xolHFjDi_Hj4"
#
# # **Desafio 100**
# **Python 3 - 3º Mundo**
#
# Descrição: Faça um programa que tenha uma lista chamada números e duas funções chamadas sorteia() e somaPar(). A primeira função vai sortear 5 números e vai colocá-los dentro da lista e a segunda função vai mostrar a soma entre todos os valores pares sorteados pela função anterior.
#
# Link: https://www.youtube.com/watch?v=MEs-41JcuhM&t=25s
# + id="8Lylf4Su-6aB"
from random import randint
from time import sleep
def sorteia(numeros):
print('Sorteando 5 valores da lista: ', end='')
for i in range(5):
n = randint(1, 10)
print(f'{n} ', end='')
sleep(0.3)
numeros.append(n)
print('PRONTO!')
def somapar(numeros):
s = 0
for i in numeros:
if i % 2 == 0:
s += i
print(f'Somando os valores pares de {numeros}, temos {s}')
numeros = []
sorteia(numeros)
somapar(numeros)
| Mundo03/Desafio100.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="9-ivKRhSp-9y"
# Data is from Yelp - https://www.yelp.com/dataset/documentation/main
# + [markdown] id="1qegw3OJuLJb"
# # Ideas
# - Create a corpus of reviews from top reviewers
# - try to predict ratings based on review nlp
# - use ANN or LSTM to asses review classifier prediction.
# - calculate error of ML rating vs actual rating.
# - see if there is an argument for an adjusted rating as there are too many 5 star reviews..
#
# - Create clusters of reviewers on particular metrics to find similarities or elite reviewers in the making.
# - Do this based on review NLP metrics, are they using language of top reviewers
# - maybe include a ration of reviews / fans in as a feature to normalize.
# -
# - After training, consider adding a chatbot trained on reviews.
#
# - stars can be a target variable
# - techniques to consider - bag of words , tf- idf, word2vec
# - for lstm - over 100 words is a good threshold
# - consider the frequency dist of words in the dataset - sort by lemm
# - train models by city / category
# - look at confusion matrix on models that are by city vs all cities model.
# - clean docs in whole prior to slicing for tf-idf / vectorizaion (word2vec)
# + [markdown] id="j-K-tsEsiTcT"
# ## Imports,
# + id="I_ldGnWPI10A" executionInfo={"status": "ok", "timestamp": 1616107797905, "user_tz": 420, "elapsed": 4206, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}}
import json
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tqdm
from nltk.corpus import stopwords
import spacy
from collections import Counter
import nltk
import spacy
import re
from keras.preprocessing.text import Tokenizer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
# + colab={"base_uri": "https://localhost:8080/"} id="I9xzl6rNhsyp" executionInfo={"status": "ok", "timestamp": 1616107798259, "user_tz": 420, "elapsed": 4551, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="110a8620-89ee-4289-c373-0dde99888849"
from nltk import word_tokenize
from nltk.corpus import stopwords
from nltk.stem.snowball import SnowballStemmer
from nltk.stem.wordnet import WordNetLemmatizer
from nltk.corpus.reader.plaintext import PlaintextCorpusReader
from sklearn.cluster import KMeans
from sklearn.cluster import AgglomerativeClustering
from sklearn.feature_extraction.text import TfidfVectorizer
import matplotlib.pyplot as plt
from yellowbrick.cluster import KElbowVisualizer
import nltk
nltk.download('wordnet')
nltk.download('punkt')
nltk.download('stopwords')
nltk.download('gutenberg')
# + colab={"base_uri": "https://localhost:8080/"} id="Z3FeUQSwwZpC" executionInfo={"status": "ok", "timestamp": 1616107801806, "user_tz": 420, "elapsed": 8088, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="fc7f5eeb-759d-4e76-9ad5-b0ec221c9975"
# !python -m spacy download en --quiet
# + id="z2KMCCx776wi" executionInfo={"status": "ok", "timestamp": 1616107801807, "user_tz": 420, "elapsed": 8083, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}}
def clean_cats(x):
tokens = word_tokenize(x)
cleaned_tokens = [token.lower() for token in tokens if token.lower() not in stopwords.words('english') if token.isalpha()] # is alpha is only A-Z
sorted_tokens = sorted(cleaned_tokens)
return ' '.join(sorted_tokens)
# + [markdown] id="OM4xUIu8fUD5"
# ## Download Data and perform basic EDA, and Merge DataFrames
# - Biz Data
# - User Data
# - Review Data
# + colab={"base_uri": "https://localhost:8080/"} id="lyHaBDYR22zw" executionInfo={"status": "ok", "timestamp": 1616107804248, "user_tz": 420, "elapsed": 10518, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="d3631fee-457e-4983-df86-d4928140afb3"
data = {'business_id': [], 'name':[], 'city': [], 'stars':[], 'review_count': [], 'categories': []}
with open('/content/drive/MyDrive/Colab Notebooks/Yelp_dataset/yelp_academic_dataset_business.json') as f:
for line in tqdm(f):
review = json.loads(line)
data['business_id'].append(review['business_id'])
data['name'].append(review['name'])
data['city'].append(review['city'])
data['stars'].append(review['stars'])
data['review_count'].append(review['review_count'])
data['categories'].append(review['categories'])
biz_df = pd.DataFrame(data)
biz_df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 80} id="KG2C-JvqwHrv" executionInfo={"status": "ok", "timestamp": 1616107804249, "user_tz": 420, "elapsed": 10512, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="01a640c2-2d38-42d5-aece-cfea3358ff9f"
biz_df[biz_df.business_id == 'ISBs1ARjIFCXGWqPmGSMog']
# + colab={"base_uri": "https://localhost:8080/"} id="MRkMmE66j9mF" executionInfo={"status": "ok", "timestamp": 1616107804250, "user_tz": 420, "elapsed": 10504, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="e70ddd9e-0f20-4f7e-ad7e-353fd5cb5515"
biz_df.isna().sum()
# + id="D4682b76lPAM" executionInfo={"status": "ok", "timestamp": 1616107804250, "user_tz": 420, "elapsed": 10496, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}}
biz_df = biz_df.dropna()
# + colab={"base_uri": "https://localhost:8080/"} id="iysn_Ms6I1wB" executionInfo={"status": "ok", "timestamp": 1616107915473, "user_tz": 420, "elapsed": 121714, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="07952d7b-c780-4432-b36e-66fc520f200f"
data = {'review_id':[], 'user_id': [],'business_id': [], 'stars':[], 'text': [], 'useful': [], 'funny': [], 'cool': []}
with open('/content/drive/MyDrive/Colab Notebooks/Yelp_dataset/yelp_academic_dataset_review.json') as f:
for line in tqdm(f):
review = json.loads(line)
data['review_id'].append(review['review_id'])
data['user_id'].append(review['user_id'])
data['business_id'].append(review['business_id'])
data['stars'].append(review['stars'])
data['text'].append(review['text'])
data['useful'].append(review['useful'])
data['funny'].append(review['funny'])
data['cool'].append(review['cool'])
rev_df = pd.DataFrame(data)
rev_df.info()
# + colab={"base_uri": "https://localhost:8080/"} id="GwIk09xpsgeV" executionInfo={"status": "ok", "timestamp": 1616107970865, "user_tz": 420, "elapsed": 177099, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="4128bb43-96ce-4ebb-af01-d2d783fe738b"
data = {'user_id': [],'review_count': [], 'fans':[], 'average_stars': []}
#data = {'business_id': []}
with open('/content/drive/MyDrive/Colab Notebooks/Yelp_dataset/yelp_academic_dataset_user.json') as f:
for line in tqdm(f):
review = json.loads(line)
data['user_id'].append(review['user_id'])
data['review_count'].append(review['review_count'])
data['fans'].append(review['fans'])
data['average_stars'].append(review['average_stars'])
user_df = pd.DataFrame(data)
user_df.info()
# + id="dfkJg_LuHSUH" executionInfo={"status": "ok", "timestamp": 1616107989200, "user_tz": 420, "elapsed": 195427, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}}
df = rev_df.merge(user_df, on = 'user_id', how = 'left').merge(biz_df, on = 'business_id', how = 'left')
# + colab={"base_uri": "https://localhost:8080/"} id="h1T7t4EeYmJJ" executionInfo={"status": "ok", "timestamp": 1616108003084, "user_tz": 420, "elapsed": 209305, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="cc5c1a32-6031-41ec-98c1-f23de0624f5e"
df = df.dropna()
df.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="OuK01SlTE_Lb" executionInfo={"status": "ok", "timestamp": 1616108003825, "user_tz": 420, "elapsed": 210040, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="7a89787e-aff2-4358-89dc-d48133e0a3c1"
df = df.sample(frac = .01, replace = 'disallow')
df.shape
# + id="_8OrgO_mtr5I" executionInfo={"status": "ok", "timestamp": 1616108111854, "user_tz": 420, "elapsed": 318061, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}}
# Clean df - rename columns, drop unneeded columns
df['cur_review_star'] = df['stars_x']
df['user_rev_sum'] = df['review_count_x']
df['biz_name'] = df['name']
df['user_avg_star'] = df['average_stars']
df['biz_star'] = df['stars_y']
df['biz_rev_sum'] = df['review_count_y']
df['rev_fb'] = (df['useful'] + df['funny'] + df['cool']) # summing positive feedback actions from other users.
df['clean_cats'] = df.categories.apply(clean_cats)
df['star_diff'] = (df['cur_review_star'] - df['biz_star'])/ df['user_avg_star'] # did user give a boost or decline to biz star avg.
df['fb_ratio'] = (df['rev_fb']+.01) / np.abs(df['star_diff']+.01) # Get a normalized metric for how valuable the review is to the readers.
df = df.drop(columns= [
'fans',
'stars_x',
'review_count_x',
'name',
'average_stars',
'stars_y',
'review_count_y',
'categories',
'useful',
'funny',
'cool',
])
# + colab={"base_uri": "https://localhost:8080/", "height": 534} id="6VgR_O01liC7" executionInfo={"status": "ok", "timestamp": 1616108111855, "user_tz": 420, "elapsed": 318055, "user": {"displayName": "Jason B", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="182db2b9-e735-47a7-bd4b-8cad8cef3dbc"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="j_4y9HPqOBHc" executionInfo={"status": "ok", "timestamp": 1616108111855, "user_tz": 420, "elapsed": 318049, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="8c301d05-93b2-4cdb-c577-ecf704a464de"
df.shape
# + id="i-6MrMkolq4q" executionInfo={"status": "ok", "timestamp": 1616108114197, "user_tz": 420, "elapsed": 320384, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}}
df.to_csv('merged_data.csv', index = False)
# !cp merged_data.csv '/content/drive/MyDrive/Colab Notebooks/Yelp NLP Project'
# + colab={"base_uri": "https://localhost:8080/", "height": 294} id="Vk5SJ-B95MUW" executionInfo={"status": "ok", "timestamp": 1616108114418, "user_tz": 420, "elapsed": 320597, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="6086d2cc-9380-4cd7-9e40-8d715284e70f"
df.describe()
# + id="1n_Ensn71OVm" executionInfo={"status": "ok", "timestamp": 1616108114419, "user_tz": 420, "elapsed": 320591, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}}
#to - do
# visualize the data
# + colab={"base_uri": "https://localhost:8080/", "height": 362} id="txcVEsmo1OR2" executionInfo={"status": "ok", "timestamp": 1616108114754, "user_tz": 420, "elapsed": 320919, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="5a9f170f-87dd-47ee-d563-b2b9f64bc1ff"
df.biz_star.hist()
plt.title('Star Avg for Businesses on Yelp')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 362} id="ynIi09tQ1OON" executionInfo={"status": "ok", "timestamp": 1616111506860, "user_tz": 420, "elapsed": 493, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="95c75f1a-3f9a-46f8-aa7f-a3c294b5dbcb"
df.cur_review_star.hist()
plt.title('Typical Stars Given Per Review on Yelp')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 364} id="eK_eDGlI1OHR" executionInfo={"status": "ok", "timestamp": 1616108114965, "user_tz": 420, "elapsed": 321110, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="5a4ed4dd-6942-4e9d-9506-a2e0fe46d5d7"
df.user_avg_star.hist()
# + colab={"base_uri": "https://localhost:8080/"} id="sQ00OhIX1OD5" executionInfo={"status": "ok", "timestamp": 1616108114965, "user_tz": 420, "elapsed": 321104, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="dd5cca00-8eca-4997-b403-bf28cddc6920"
print(df.star_diff.min())
df.star_diff.max()
# + [markdown] id="_wpoM6X5z3aq"
# ## Dimension Reduce by category filters
# - Food
# - Beauty
# + colab={"base_uri": "https://localhost:8080/", "height": 362} id="gzzho_6TkdPm" executionInfo={"status": "ok", "timestamp": 1616111738971, "user_tz": 420, "elapsed": 686, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="f6ffc8be-e031-47ac-9b4b-180d3fb9c4db"
beauty_df = df[df.clean_cats.str.contains(pat = 'beauty')]
beauty_df.cur_review_star.hist()
plt.title("Stars Given Per Review - Beauty")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="VNVZrl-sudz3" executionInfo={"status": "ok", "timestamp": 1616111666011, "user_tz": 420, "elapsed": 915, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="5aff5aba-5e87-423a-af96-0a0747faa1fb"
beauty_df.head(2)
# + colab={"base_uri": "https://localhost:8080/"} id="y-U_ycrOXKmY" executionInfo={"status": "ok", "timestamp": 1616111670399, "user_tz": 420, "elapsed": 480, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="c40c7c4a-7ca4-4100-b486-77d0b20d5ffb"
rest_df = df[df.clean_cats.str.contains(pat = 'restaurant|food')]
len(rest_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 362} id="py3P1vDTlFhO" executionInfo={"status": "ok", "timestamp": 1616111672365, "user_tz": 420, "elapsed": 430, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="5bef987c-0082-405f-8b62-4080861cb2a1"
rest_df.cur_review_star.hist()
plt.title("Stars Given Per Review - Food")
plt.show()
# + id="z3gG_NbBqMBp" colab={"base_uri": "https://localhost:8080/", "height": 362} executionInfo={"status": "ok", "timestamp": 1616111796896, "user_tz": 420, "elapsed": 574, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="ac2b9005-6f8b-4873-8f90-e52395b56bf9"
top_r_revs = rest_df[rest_df['fb_ratio'] >= 39]
top_r_revs.cur_review_star.hist()
plt.title("More Useful Reviews by Feedback - Restaurant Stars Given")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="vifZ-NqMpy5n" executionInfo={"status": "ok", "timestamp": 1616111680351, "user_tz": 420, "elapsed": 488, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="d2ed025e-a3d4-424a-bc83-439f6abbc076"
hotel_df = df[df.clean_cats.str.contains(pat = 'hotel')]
len(hotel_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 362} id="obWR2fh5gomf" executionInfo={"status": "ok", "timestamp": 1616111694801, "user_tz": 420, "elapsed": 523, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="fae82333-a9a9-4bd6-a95c-188006c93d78"
hotel_df.cur_review_star.hist()
plt.title("Stars Given Per Review - Hotel")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="xHb6iBpx7D9v" executionInfo={"status": "ok", "timestamp": 1616111831895, "user_tz": 420, "elapsed": 326, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="e9dbd651-28ea-485b-ef6b-482ac33de1ef"
five_cities = df.city.value_counts().head(5).reset_index()
five_cities['index'].values # top 5 cities for review amounts
# + colab={"base_uri": "https://localhost:8080/"} id="pyxPnV-F8MjB" executionInfo={"status": "ok", "timestamp": 1616111834125, "user_tz": 420, "elapsed": 309, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjbKOYPQyDmJL_lXR9vLi9QTBw4qZ0BrK7aT_IFHw=s64", "userId": "04496914521905721027"}} outputId="78456390-ce35-4ef2-82b8-13a56564332b"
top_c = df.loc[df['city'].isin(five_cities['index'].values)]
top_c.clean_cats.value_counts().head(10) # top 10 categories by freq in top 5 cities
# + [markdown] id="J0f3xV0Ks3xN"
# # Notes -
# - reduce dimensionality
# - can do umap style
# - can remove words not in dictionary
#
#
# Notes with Reza -
# from nltk.corpus import words as nltk_words
#
# dictionary = dict.fromkeys(nltk_words.words(), None)
#
# def spellchk(word): try: if dictionary[word.lower()] == None: return True except KeyError: return False
#
# def no_number_preprocessor(tokens): r = re.sub('(\d)+', 'NUM', tokens.lower()) # This alternative just removes numbers: # r = re.sub('(\d)+', '', tokens.lower()) return r
#
# def remove_non_ascii(text): return ''.join(i for i in text if ord(i)<128)
#
#
# - Try topic modeling clusters - then analyze in a df.
| Final_CSTONE_Research.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import tubesml as tml
from sklearn.pipeline import Pipeline
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import KFold
from sklearn.linear_model import Lasso, Ridge, SGDRegressor
from sklearn.ensemble import RandomForestRegressor, ExtraTreesRegressor
from sklearn.svm import SVR
from sklearn.neighbors import KNeighborsRegressor
from sklearn.neural_network import MLPRegressor
import xgboost as xgb
import lightgbm as lgb
from source import utility as ut
from source import transform as tr
from source.train import train_model
import warnings
pd.set_option('max_columns', 100)
# -
df_train = pd.read_csv('data/train.csv')
df_test = pd.read_csv('data/test.csv')
subs = pd.read_csv('data/sample_submission.csv')
# +
train_set, test_set = ut.make_test(df_train, 0.25, random_state=516, strat_feat='cat9')
train_set.head()
# -
del train_set['id']
del test_set['id']
# +
numeric_pipe = Pipeline([('fs', tml.DtypeSel('numeric'))])
cat_pipe = Pipeline([('fs', tml.DtypeSel('category')),
('tarenc', tr.TargetEncoder(to_encode=[f'cat{c}' for c in range(3,10)])),
('dummies', tml.Dummify(match_cols=True, drop_first=True))])
processing_pipe = tml.FeatureUnionDf(transformer_list=[('cat_pipe', cat_pipe),
('num_pipe', numeric_pipe)])
full_pipe = Pipeline([('processing', processing_pipe),
('scaler', tml.DfScaler())])
# +
tmp = train_set.copy()
full_pipe.fit_transform(tmp, train_set['target']).head()
# +
del tmp
y = train_set['target'].copy()
del train_set['target']
y_test = test_set['target'].copy()
del test_set['target']
# +
models = [('lasso', Lasso(alpha=0.01)), ('ridge', Ridge()), ('sgd', SGDRegressor()),
('xgb', xgb.XGBRegressor(n_estimators=200, objective='reg:squarederror', n_jobs=5)),
('lgb', lgb.LGBMRegressor(n_estimators=200, n_jobs=5))]
mod_name = []
rmse_train = []
rmse_test = []
folds = KFold(5, shuffle=True, random_state=541)
warnings.filterwarnings("ignore",
message="The dummies in this set do not match the ones in the train set, we corrected the issue.")
for model in models:
train = train_set.copy()
test = test_set.copy()
print(model[0])
mod_name.append(model[0])
pipe = [('processing', processing_pipe),
('scaler', tml.DfScaler())] + [model]
model_pipe = Pipeline(pipe)
inf_preds = tml.cv_score(data=train, target=y, cv=folds, estimator=model_pipe)
model_pipe.fit(train, y) # refit on full train set
preds = model_pipe.predict(test)
rmse_train.append(mean_squared_error(y, inf_preds))
rmse_test.append(mean_squared_error(y_test, preds))
print(f'\tTrain set RMSE: {round(np.sqrt(mean_squared_error(y, inf_preds)), 4)}')
print(f'\tTest set RMSE: {round(np.sqrt(mean_squared_error(y_test, preds)), 4)}')
print('_'*40)
print('\n')
results = pd.DataFrame({'model_name': mod_name,
'rmse_train': rmse_train, 'rmse_test': rmse_test})
results
# +
mod = xgb.XGBRegressor(n_estimators=2000, objective='reg:squarederror', n_jobs=5)
folds = KFold(10, shuffle=True, random_state=541)
oof, pred, imp = train_model(train_set, test_set, y, full_pipe, mod, folds, early_stopping=100, verbose=False)
print(np.sqrt(mean_squared_error(y_pred=oof, y_true=y)))
print(np.sqrt(mean_squared_error(y_pred=pred, y_true=y_test)))
imp
# +
mod = lgb.LGBMRegressor(n_estimators=2000, n_jobs=5)
folds = KFold(10, shuffle=True, random_state=541)
oof, pred, imp = train_model(train_set, test_set, y, full_pipe, mod, folds, early_stopping=100, verbose=False)
print(np.sqrt(mean_squared_error(y_pred=oof, y_true=y)))
print(np.sqrt(mean_squared_error(y_pred=pred, y_true=y_test)))
imp.head(15)
# +
numeric_pipe = Pipeline([('fs', tml.DtypeSel('numeric'))])
cat_pipe = Pipeline([('fs', tml.DtypeSel('category')),
('tarenc', tr.TargetEncoder(to_encode=[f'cat{c}' for c in range(3,10)])),
('dummies', tml.Dummify(match_cols=True, drop_first=True))])
processing_pipe = tml.FeatureUnionDf(transformer_list=[('cat_pipe', cat_pipe),
('num_pipe', numeric_pipe)])
full_pipe = Pipeline([('processing', processing_pipe),
('scaler', tml.DfScaler()),
('pca', tr.PCADf(n_components=0.9))])
# +
mod = xgb.XGBRegressor(n_estimators=2000, objective='reg:squarederror', n_jobs=5)
folds = KFold(10, shuffle=True, random_state=541)
oof, pred, imp = train_model(train_set, test_set, y, full_pipe, mod, folds, early_stopping=100, verbose=False)
print(np.sqrt(mean_squared_error(y_pred=oof, y_true=y)))
print(np.sqrt(mean_squared_error(y_pred=pred, y_true=y_test)))
imp
# +
mod = lgb.LGBMRegressor(n_estimators=2000, n_jobs=5)
folds = KFold(10, shuffle=True, random_state=541)
oof, pred, imp = train_model(train_set, test_set, y, full_pipe, mod, folds, early_stopping=100, verbose=False)
print(np.sqrt(mean_squared_error(y_pred=oof, y_true=y)))
print(np.sqrt(mean_squared_error(y_pred=pred, y_true=y_test)))
imp.head(15)
# +
numeric_pipe = Pipeline([('fs', tml.DtypeSel('numeric'))])
cat_pipe = Pipeline([('fs', tml.DtypeSel('category')),
('tarenc', tr.TargetEncoder(to_encode=[f'cat{c}' for c in range(3,10)], agg_func='skew')),
('dummies', tml.Dummify(match_cols=True, drop_first=True))])
processing_pipe = tml.FeatureUnionDf(transformer_list=[('cat_pipe', cat_pipe),
('num_pipe', numeric_pipe)])
full_pipe = Pipeline([('processing', processing_pipe),
('scaler', tml.DfScaler())])
# +
mod = xgb.XGBRegressor(n_estimators=2000, objective='reg:squarederror', n_jobs=5)
folds = KFold(10, shuffle=True, random_state=541)
oof, pred, imp = train_model(train_set, test_set, y, full_pipe, mod, folds, early_stopping=100, verbose=False)
print(np.sqrt(mean_squared_error(y_pred=oof, y_true=y)))
print(np.sqrt(mean_squared_error(y_pred=pred, y_true=y_test)))
imp
# +
mod = lgb.LGBMRegressor(n_estimators=2000, n_jobs=5)
folds = KFold(10, shuffle=True, random_state=541)
oof, pred, imp = train_model(train_set, test_set, y, full_pipe, mod, folds, early_stopping=100, verbose=False)
print(np.sqrt(mean_squared_error(y_pred=oof, y_true=y)))
print(np.sqrt(mean_squared_error(y_pred=pred, y_true=y_test)))
imp.head(15)
# -
| tabular_playground/february/03_simple_model_sel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 - rstudio
# language: python
# name: rstudio-user-3.9.5
# ---
# # Lab 7: git and GitHub
#
#
# ## Acknowledgements
#
# Much of the material for this lesson was borrowed from or inspired by <NAME>' NCEAS Reproducible Research Techniques for Synthesis workshop](https://learning.nceas.ucsb.edu/2020-02-RRCourse/)
#
#
# ## Learning Objectives
#
# In this lab, you will learn:
#
# - What computational reproducibility is and why it is useful
# - How version control can increase computational reproducibility
# - to set up git on your computer
# - to use git and github to track changes to your work over time
#
#
# ## Reproducible Research
#
# Reproducibility is the hallmark of science, which is based on empirical observations
# coupled with explanatory models. While reproducibility encompasses
# the full science lifecycle, and includes issues such as methodological consistency and
# treatment of bias, in this course we will focus on **computational reproducibility**:
# the ability to document data, analyses, and models sufficiently for other researchers
# to be able to understand and ideally re-execute the computations that led to
# scientific results and conclusions.
#
# ### What is needed for computational reproducibility?
#
# The first step towards addressing these issues is to be able to evaluate the data,
# analyses, and models on which conclusions are drawn. Under current practice,
# this can be difficult because data are typically unavailable, the method sections
# of papers do not detail the computational approaches used, and analyses and models
# are often conducted in graphical programs, or, when scripted analyses are employed,
# the code is not available.
#
# And yet, this is easily remedied. Researchers can achieve computational
# reproducibility through open science approaches, including straightforward steps
# for archiving data and code openly along with the scientific workflows describing
# the provenance of scientific results (e.g., @hampton_tao_2015, @munafo_manifesto_2017).
#
# ### Conceptualizing workflows
#
# Scientific workflows encapsulate all of the steps from data acquisition, cleaning,
# transformation, integration, analysis, and visualization.
#
# 
#
# Workflows can range in detail from simple flowcharts
# to fully executable scripts. R scripts and python scripts are a textual form
# of a workflow, and when researchers publish specific versions of the scripts and
# data used in an analysis, it becomes far easier to repeat their computations and
# understand the provenance of their conclusions.
#
# ### The problem with filenames
#
# Every file in the scientific process changes. Manuscripts are edited.
# Figures get revised. Code gets fixed when problems are discovered. Data files
# get combined together, then errors are fixed, and then they are split and
# combined again. In the course of a single analysis, one can expect thousands of
# changes to files. And yet, all we use to track this are simplistic *filenames*.
# You might think there is a better way, and you'd be right: __version control__.
#
# Version control systems help you track all of the changes to your files, without
# the spaghetti mess that ensues from simple file renaming. In version control systems
# like `git`, the system tracks not just the name of the file, but also its contents,
# so that when contents change, it can tell you which pieces went where. It tracks
# which version of a file a new version came from. So its easy to draw a graph
# showing all of the versions of a file, like this one:
#
# 
#
# Version control systems assign an identifier to every version of every file, and
# track their relationships. They also allow branches in those versions, and merging
# those branches back into the main line of work. They also support having
# *multiple copies* on multiple computers for backup, and for collaboration.
# And finally, they let you tag particular versions, such that it is easy to return
# to a set of files exactly as they were when you tagged them. For example, the
# exact versions of data, code, and narrative that were used when a manuscript was originally
# submitted might be `eco-ms-1` in the graph above, and then when it was revised and resubmitted,
# it was done with tag `eco-ms-2`. A different paper was started and submitted with tag `dens-ms-1`, showing that you can be working on multiple manuscripts with closely related but not identical sets of code and data being used for each, and keep track of it all.
#
#
# ## Version control and Collaboration using Git and GitHub
#
# First, just what are `git` and GitHub?
#
# - __git__: version control software used to track files in a folder (a repository)
# - git creates the versioned history of a repository
# - __GitHub__: web site that allows users to store their git repositories and share them with others
#
#
# ### Getting started on GitHub
#
# Go to https://github.com/ and sign up for an account. This is a good opportunity to create a professional presence in the bioinformatics and data science world.
#
# ### Creating a personal access token
#
# Starting this fall GitHub is requiring personal access tokens. To generate one follow https://docs.github.com/en/authentication/keeping-your-account-and-data-secure/creating-a-personal-access-token This will serve as your password later.
#
# ### Cloning a repository
#
# You can clone any Github repository. This is the whole purpose, to openly share code. To do this you must install git on your computer. Fortunately for us git is already
#
#
# Let's start by cloning the course repository - https://github.com/jeffreyblanchard/EvoGeno2021PyTo do so, locate the green "CODE" button and copy the URL. Now go to RStudio Cloud or Unity and open a terminal. Type
#
git clone https://github.com/jeffreyblanchard/EvoGeno2021Py.git
# If you type ls you will now see a new directory "EvoGeno2021Py" with all of the class materials. To update the directory use the git pull command when you are inside the directory
# cd EvoGeno2021Py
git pull
# *** Note: If you change the contents of the directory in any way it will be out of sync with the course git hub repo and you will get an error when you try to pull. More on this in a bit
# ### Create your own repository on GitHub
#
# Next create a repository on GitHub, then we'll edit some files.
#
# - Log into [GitHub](https://github.com)
# - Click on the Repositories tab
# - Click the New Repository button
# - Name it `genomics-course` or something similar
# - Create a README.md
# - Set the LICENSE to Apache 2.0
#
# You've now created your first repository! It has a couple of files that GitHub created
# for you, like the README.md file, and the LICENSE file
#
# For simple changes to text files, you can make edits right in the GitHub web interface. For example,
# navigate to the `README.md` file in the file listing, and edit it by clicking on the *pencil* icon.
# This is a regular Markdown file, so you can just add text, and when done, add a commit message, and
# hit the `Commit changes` button.
#
# Congratulations, you've now authored your first versioned commit. If you navigate back to the GitHub page for the repository, you'll see your commit listed there, as well as the
# rendered README.md file.
#
# Now locate the green `CODE` button and copy the URL. Now go to RStudio Cloud or Unity and open a terminal. Type git clone and the name of your URL as you did above.
# ### Pushing and Pulling changes.
#
# Make a simple change to your README file directly in your GitHub repo. In order for your RStudio Cloud or Unity to update that change move into your GitHub directory in RStudio Cloud or Unity and
git pull
# ***Once you have created a Git repository on RStudio Cloud, Unity or your laptop I recommend NOT making changes on the Github site as it is easy to get the repositories out of sync
# Now create a new Jupyter notebook file in your repo/directory on RStudio Cloud or Unity and put some simple text and code in the notebook. Save it as test.ipynb
#
# To make the notebook file or any changes to it visible on GitHub we have to "Push" the changes. This involves a series of steps which include indicate which specific which changes to the local working files should be staged for versioning (using the `git add` command), and when to record those changes as a version in the local repository (using the command `git commit`).
#
# The remaining concepts are involved in synchronizing the changes in your local repository with changes in a remote repository. The `git push` command is used to send local changes up to the remote repository on GitHub,
#
# 
#
# In your terminal type
git add test.ipynb
# then
git commit
# When using the git commit command in RStudio or Unity it will open the Unix text editors `vim` and `nano` respectively. You will need to learn a few import commands
#
#
# - `vim` starts in the command mode. To insert text type `i` and add your commit message (e.g. Adding test file). To exit the insert mode hit `esc`. To save and exit `vim` `shift ZZ` with shift held down for both Zs. For more info on `vim` https://www.vim.org/
#
# - In `nano` you can start typing your commit message like 'Adding test file'. Then use `ctr x` to exit and save the changes to the file. For more info on `nano` https://www.nano-editor.org/docs.php
#
# Now you can `Push` the changes to your GitHub repo
git push
# The first time doing this you will need to add the email you used for Github and your username following the suggested command line syntax
# ### Deleting files in git
#
# If you delete a file from your RStudio Cloud or Unity git repository, you still need to tell Github about it. The process is still the same as shown with the example.txt file
git add example.txt
git commit
git push
# ### On good commit messages
#
# Clearly, good documentation of what you've done is critical to making the version history of your repository meaningful and helpful. Its tempting to skip the commit message altogether, or to add some stock blurd like 'Updates'. Its better to use messages that will be helpful to your future self in deducing not just what you did, but why you did it. Also, commit messaged are best understood if they follow the active verb convention. For example, you can see that my commit messages all started with a past tense verb, and then explained what was changed.
#
# While some of the changes we illustrated here were simple and so easily explained in a short phrase, for more complext changes, its best to provide a more complete message. The convention, however, is to always have a short, terse first sentence, followed by a more verbose explanation of the details and rationale for the change. This keeps the high level details readable in the version log. I can't count the number of times I've looked at the commit log from 2, 3, or 10 years prior and been so grateful for diligence of my past self and collaborators.
# ### Github web pages
#
# You can enable Github pages to create a web presence for your project.
#
# - Go to your repository you just created
# - Click on Settings
# - Scroll down to GitHub pages
# - Select Master branch
# - Click on Save
# (Do not choose a theme today. You have the option of choosing a theme later).
# - It will create a GitHub page (e.g. https://jeffreyblanchard.github.io/jeffblanchard/)
# - Copy the link to your GitHub page
# - Go to the main page for your web (e.g. jeffblanchard) repository.
# - In the `about section` add the url to your GitHub repo page
#
# Under the settings tab enable Github pages. It takes about 10 min for the web site to appear. The default web pages in the README.md file, but if you create and upload an index.html page this will be your new default. This provides a way to see the html files in your browser as you intended them to appear (not just the html code).
#
# * Note: It is critical that you use a small `i` in `index.html` and not a captial `I`
#
#
# ### Github project management
#
# You can keep tract of ideas, todos and fixes by creating a wiki or using the Project
#
# 
#
#
#
# ### Collaboration and conflict free workflows (we walk talk more about this later in the class)
#
# Up to now, we have been focused on using Git and GitHub for yourself, which is a great use. But equally powerful is to share a GitHib repository with other researchers so that you can work on code, analyses, and models together. When working together, you will need to pay careful attention to the state of the remote repository to avoid and handle merge conflicts. A *merge conflict* occurs when two collaborators make two separate commits that change the same lines of the same file. When this happens, git can't merge the changes together automatically, and will give you back an error asking you to resolve the conflict. Don't be afraid of merge conflicts, they are pretty easy to handle. and there are some
# [great](https://help.github.com/articles/resolving-a-merge-conflict-using-the-command-line/) [guides](https://stackoverflow.com/questions/161813/how-to-resolve-merge-conflicts-in-git).
#
# That said, its truly painless if you can avoid merge conflicts in the first place. You can minimize conflicts by:
#
# - Ensure that you pull down changes just before you commit
# + Ensures that you have the most recent changes
# + But you may have to fix your code if conflict would have occurred
# - Coordinate with your collaborators on who is touching which files
# + You still need to comunicate to collaborate
#
# ### More with git
#
# There's a lot we haven't covered in this brief tutorial. There are some good longer tutorials that cover additional topics:
#
# - Git Guides - https://github.com/git-guides/git-push
# - Git cheatsheat - https://github.com/git-guides/git-push
# - Git Learning Lab - https://lab.github.com/
# - [Happy Git and Github for the useR](https://happygitwithr.com/)
# - [Try Git](https://try.github.io) a great interactive tutorial
# - Software Carpentry [Version Control with Git](http://swcarpentry.github.io/git-novice/)
#
#
# ### Example Github repositories and pages
#
# - <NAME> - https://github.com/nickreich
# - <NAME> - http://seedscape.github.io/BeckmanLab/Beckman.html
# - Women In Soil Ecology - https://womeninsoilecology.github.io/
# - <NAME> - https://github.com/faylward
# ## Exercises
#
# You only need to turn in the link to your GitHub repository. In it should be your new course repo and a test file.
| EvoGeno_Lab7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src='./img/intel-logo.jpg' width=30%, Fig1>
#
# # 파이썬 기초강의
# <font size=5><b>01. 연 습 문 제<b></font>
#
# <div align='right'>이 인 구 (<NAME>)</div>
#
# ## Example : 1
#
# ### 직육면체의 부피를 구해보자
# <img src='./img/volume.png' width=50%, Fig2>
# 변수 설정
length = 5
height = 5
width = 20
# +
volume = length*width*height
print('직육면체의 부피 : %d'%volume)
# +
length = 10 # 다시 할당하면 됨
volume = length*width*height
print('직육면체의 부피 : %d'%volume)
# -
# ## Example : 2
# ### for 를 사용해서 암컷 개를 찾으세요. and를 사용해서 일치하는것을 찾으세요
# suspects = [['낙타', '포유류','암컷'], ['상어','어류','숫컷'], ['푸들','개','암컷']]
#
#
#
#
#
#
#
#
# +
suspects = [['낙타', '포유류','암컷'], ['상어','어류','숫컷'], ['푸들','개','암컷']]
for suspect in suspects:
if suspect[1] == '개' and suspect[2] =='암컷':
print('범인은', suspect[0], '입니다')
# -
# ## Example : 3
# ### 연이율구하기
# ```
# 2017년 7월 2일 연이율 3% 계좌를 생성하여 3000000원을 입금한 경우
# 2018년 7월 2일 계좌 총액을 계산하여 출력하는 프로그램을 작성하십시오.
# 프로그램에서 입금액을 위한 변수, 연이율을 위한 변수를 만들어 사용하십시오.
#
# 위의 프로그램을 입금액과 연이율을 입력받아 총액을 출력하도록 변경하십시오.
# 언어 : python3
# 입력 설명 :
#
# 다음은 입금액과 연이율의 입력예입니다.
# ===============================
# 입금액(원), 연이율(%):: 4000, 3
# 출력 설명 :
#
# 다음과 같이 1년 후 총액을 출력합니다.
# ===============================
# 4120.0
#
# 샘플 입력 : 4000, 3
# 샘플 출력 : 4120.0
# ```
# +
money, ratio = eval(input('입금액(원), 연이율(%)::'))
print(money*(1+(1/100)*ratio))
# -
# ## Example : 4
# ### 삼각형 넓이를 구하시오
# ```
#
# 삼각형의 세변의 길이가 3,4,5인 경우 삼각형 넓이는 다음과 같이 계산합니다.
# x = (3 + 4 + 5)/2
# 넒이는 x(x-3)(x-4)(x-5) 의 양의 제곱근
# 언어 : python3
# 입력 설명 :
#
# 다음과 같이 삼각형의 세변의 길이를 입력합니다.
# ======================
# 삼각형 세변의 길이(comma로 구분): 3,4,5
# 출력 설명 :
#
# 다음과 같이 삼각형의 넓이를 출력합니다.
# ======================
# 6.0
#
# 샘플 입력 : 3,4,5
# 샘플 출력 : 6.0
# ```
# +
a,b,c = eval(input())
x = (a+b+c)/2
area = (x*(x-a)*(x-b)*(x-c))**(0.5)
print(area)
# -
# <img src='./img/C1.jpg'>
#
# ## Example : 5
# ### for 를 사용해서 암컷 개를 찾으세요. and를 사용해서 일치하는것을 찾으세요
# +
suspects = [['낙타', '포유류','암컷'], ['상어','어류','숫컷'], ['푸들','개','암컷']]
for suspect in suspects:
if suspect[1] == '개' and suspect[2] =='암컷':
print('범인은', suspect[0], '입니다')
# -
# ## Example : 6
# ### random을 이용해서 중복되지 않는 카드 두 장을 뽑도록 빈칸을 채우세요.
# +
import random
cities = ['서울','부산','울산','인천' ]
print(random.sample(cities, 2))
# -
# ## Example : 7
# ### 다음중 하나를 무작위로 뽑아주세요!
# annimals = 얼룩말, 황소, 개구리, 참새.
#
# +
#리스트[]
import random
annimals = ['얼룩말','황소', '개구리', '참새']
print(random.choice(annimals))
# -
# ## Example : 8
# ### def 를 이용해서 서로에게 인사하는 문구를 만들어 보세요!
# 가브리엘 님 안녕하세요? \
# 엘리스 님 안녕하세요?
# +
def welcome(name):
print(name,'님 안녕하세요?')
welcome('가브리엘')
welcome('엘리스')
# -
# ## Example : 9
# ### 점수에 따라 학점을 출력 해주세요.
# 철수의 점수는 75점 입니다. 몇 학점 인지 표시해 주세요.
# A학점은 80< score <=100
# B학점은 60< score <=80
# C학점은 40< score <=60
score =75
if 80< score <=100:
print('학점은 A 입니다')
if 60< score <=80:
print('학점은 B 입니다')
if 40< score <=60:
print('학점은 C 입니다')
# ## Example : 10
# ### 변수를 사용해서 매출액을 계산해 주세요.
# 주문서1 - 커피2잔, 홍차4잔, 레몬티5잔
# 주문서2 - 커피1잔, 홍차1잔, 레몬티5잔
# 주문서3 - 커피2잔, 홍차3잔, 레몬티1잔
coffee =4000
tea = 3000
lemon =200
order1 = (coffee*2 + tea*4 + lemon*5)
order2 = (coffee*1 + tea*1 + lemon*5)
order3 = (coffee*2 + tea*3 + lemon*1)
print(order1+order2+order3)
# ## Example : 11
# ### 5바퀴를 도는 레이싱 경주를 하고 있습니다. while 코드를 이용해서 트랙의 수를 카운트하고 5바퀴를 돌면 종료 멧세지를 주세요.
# 반복할 때마다 몇 번째 바퀴인지 출력하세요. \
# 5바퀴를 돌면 종료 멧세지와 함께 종료해 주세요.
#
count = 0
while count <5:
count =count +1
print(count, "번째 바퀴입니다.")
print('경주가 종료되었습니다!')
# ## Example : 12
# ### 정답을 맟춰보세요.
# 미국이 수도는 어디인기요? \
# 보기에서 찾아서 답하게 하세요.
# 런던,오타와, 파리, 뉴욕
# 틀린 답을 말하면 어느 나라의 수도인지 말해주세요.
#
while True:
answer = input('런던,오타와,파리,뉴욕 중 미국이 수도는 어디일까요?')
if answer == '뉴욕':
print('정답입니다. 뉴욕은 미국의 수도 입니다')
break
elif answer == '오타와':
print('오타와는 캐나다의 수도 입니다')
elif answer == '파리':
print('파리는 프랑스의 수도 입니다')
elif answer == '런던':
print('런던은 영국의 수도 입니다')
else:
print('보기에서 골라주세요')
# ## Example : 13
# ### 물건을 교환 해주세요
# 철수는 마트에서 형광등을 샀습니다. 그런데 LED 전구가 전기 효율이 좋아 형광등을 LED 전구로 교환 하고자 합니다.
# 형광등 3개를 LED 3개로 바꾸어 주세요.
# 형광등, 형광등, 형광등 ==> LED 전구, LED 전구, LED전구
전구 = ['형광등', '형광등', '형광등']
for i in range(3):
전구[i] = 'LED 전구'
print(전구)
# ## Example : 14
# ### 반복하기
# 동물원 원숭이 10 마리에게 인사하기.
# for을 사용해서 10마리에게 한번에 인사하기 코드를 적어주세요.
#
for num in range(10):
print ('안녕 원숭이', num)
my_str ='My name is %s' % 'Lion'
print(my_str)
'%d %d' % (1,2)
'%f %f' % (1,2)
# ### print Options
print('집단지성', end='/')
print('집단지성', end='통합하자')
| python/examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Best Practices in Coding
# ## Introduction
#
# * Many scientists write code regularly but few have been formally trained to do so
# * Best practices evolved from programmer’s folk wisdom
# * They increase productivity and decrease stress
# * Development methodologies, such as Agile Programming and Test Driven Development, are established in the software engineering industry
# * We can learn a lot from them to improve our coding skills
# # Coding Style
#
# * Readability counts
# * Explicit is better than implicit
# * Beautiful is better than ugly
# * Give your variables intention revealing names
# * For example: `numbers` instead of `nu`
# * For example: `numbers instead of list_of_float_numbers`
# Example:
def my_product(numbers):
""" Compute the product of a sequence of numbers. """
total = 1
for item in numbers:
total *= item
return total
# ## Formatting Code
#
# * Format code to coding conventions for example:
# * PEP-8
# * OR use a consistent style (especially when collaborating)
# * Conventions Specify:
# * variable naming convention
# * indentation
# * import
# * maximum line length
# * blank lines, whitespace, comments
# * Use automated tools to check adherence (aka static checking):
# * `pep8`
# * `pyflakes`
# * flake8 (combination of pep8 and pyflakes)
# * pylint
# * pycheker
#
# ## About PEP8
#
# The name of the pep8 stems from the
# [Python Enhancement Proposal #8](http://www.python.org/dev/peps/pep-0008/).
#
# This proposal is about coding conventions for the Python code comprising the
# standard library in the main Python distribution. But it is considered a
# **good practice** following it in your own code.
# ## Exercise:
#
# Provided the code below:
# +
# %%file downsample.py
import os, sys
import numpy as np
from time import time
# This function downsamples a certain image by getting the mean in a certain cell shape
def downsample(x, cell):
c0, c1 = cell
yshape = (x.shape[0] // c0, x.shape[1] // c1)
y = np.empty(yshape, x.dtype)
for i in range(yshape[0]):
for j in range(yshape[1]):
y[i, j] = x[i*c0:(i+1)*c0,j*c1:(j+1)*c1].mean()
return y
# Create a sample image
if len(sys.argv) > 1:
img_shape = int(sys.argv[1]), int(sys.argv[2])
else:
img_shape = 2**24
img = np.arange(img_shape[0]*img_shape[1], dtype=np.float32).reshape(img_shape)
t0 = time()
dsimg = downsample_(img, (16,16))
print("The time for downsampling: %.3f" % (time() - t0))
print("Initial shape: %s. Final shape: %s" % (img.shape, dsimg.shape))
# -
# Make it `pep8` and `pyflakes` clean:
# ! pep8 downsample.py
# ! pyflakes downsample.py
#
# ## Documenting Code
#
# * Minimum requirement: at least a single line docstring
# * Not only for others, but also for yourself!
# * Serves as on-line help in the interpreter
# * Document arguments and return objects, including types
# * Use the numpy docstring conventions
# * Use tools to automatically generate website from docstrings
# * pydoc
# * epydoc
# * sphinx (recommended one nowadays)
# * For complex algorithms, document every line, and include equations in docstring
# * When your project gets bigger: provide a how-to, FAQ or quick-start on your website
# ### Example of docstring
def array(object, dtype=None, copy=True, order=None, subok=False, ndmin=0):
"""
Create an array.
Parameters
----------
object : array_like
An array, any object exposing the array interface, an
object whose __array__ method returns an array, or any
(nested) sequence.
dtype : data-type, optional
The desired data-type for the array. If not given, then
the type will be determined as the minimum type required
to hold the objects in the sequence. This argument can only
be used to 'upcast' the array. For downcasting, use the
.astype(t) method.
[clip]
"""
# Implementation here...
# Example of rendered output in a console:
# +
import numpy
# numpy.array?
# -
# Example of rendered output using Sphinx:
#
# http://docs.scipy.org/doc/numpy/reference/generated/numpy.array.html#numpy.array
# ### Exercise
#
# Document the `downsample` function above following the same structure than the `array`
# docstrings above.
# # Keep it Simple (Stupid) (KIS(S) Principle)
#
# * Resist the urge to over-engineer
# * Write only what you need now
# * Simple is better than complex
# * Complex is better than complicated
# * Special cases aren’t special enough to break the rules
# * Although practicality beats purity
# # The Zen of Python
import this
# ## More info:
# * [Idiomatic Python](http://python.net/~goodger/projects/pycon/2007/idiomatic/handout.html) by <NAME>
# * PEP8: http://www.python.org/dev/peps/pep-0008/
# * About pyflakes: http://www.blog.pythonlibrary.org/2012/06/13/pyflakes-the-passive-checker-of-python-programs/
# * Nice blog on static code analyzers: http://doughellmann.com/2008/03/static-code-analizers-for-python.html
| best_practices/best_practices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Library Import
import tensorflow as tf
physical_devices = tf.config.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], True)
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, LSTM, Reshape, Flatten#,GRU
import numpy as np
import pandas as pd
import gc
import math
import os.path
import time
import matplotlib.pyplot as plt
from datetime import timedelta, datetime
from dateutil import parser
from tqdm import tqdm
import copy
import warnings
warnings.filterwarnings("ignore")
# # 2. 데이터 가공
# ## 2.1 read_csv
data_path = './data'
train_x_df = pd.read_csv(data_path + "/train_x_df.csv")
train_y_df = pd.read_csv(data_path + "/train_y_df.csv")
test_x_df = pd.read_csv(data_path + "/test_x_df.csv")
# ## 2.2 numpy array로 변환
def df2d_to_array3d(df_2d):
# 입력 받은 2차원 데이터 프레임을 3차원 numpy array로 변경하는 함수
feature_size = df_2d.iloc[:,2:].shape[1]
time_size = len(df_2d.time.value_counts())
sample_size = len(df_2d.sample_id.value_counts())
array_3d = df_2d.iloc[:,2:].values.reshape([sample_size, time_size, feature_size])
return array_3d
train_x_array = df2d_to_array3d(train_x_df) #(7362, 1380, 10)
train_y_array = df2d_to_array3d(train_y_df) #(7362, 120, 10)
test_x_array = df2d_to_array3d(test_x_df) #(529, 1380, 10)
# # 3. 모델 훈련하기
# ## 3.1 훈련 층 쌓기(모델 구성)
model=Sequential([
LSTM(units = 32, input_shape=[1380, 1]),
Dense(120 * 1),
Reshape([120, 1])
])
# +
# model = Sequential()
# model.add(LSTM(7, input_shape = (1380,1), activation = 'relu')) #
# model.add(Dense(4))
# # model.add(Flatten())
# model.add(Dense(1))
# -
model.compile(optimizer = 'adam', loss = 'mse', metrics = ['mse'])
model.summary
# # 결과 불만족 스러우면 optimizer 바꿔보고 sample_id도 증가시키고 LSTM층도 더 쌓아서 학습 파라미터 수 늘리기
# ## 3.2 validation set 훈련하기
# ### 3.2.1 validation 데이터 생성하기
# +
# train 데이터 상단의 300개 샘플로 validation set를 구성하여 학습 및 추론
valid_x_df = train_x_df[train_x_df.sample_id < 100]
valid_y_df = train_y_df[train_y_df.sample_id < 100]
valid_x_array = df2d_to_array3d(valid_x_df)
valid_y_array = df2d_to_array3d(valid_y_df)
valid_pred_array = np.zeros([100, 120, 1])
# -
# ### 3.2.2 학습 및 예측하기(compile구성 및 fit 포함)
# + tags=[]
for idx in tqdm(range(valid_x_array.shape[0])):
#학습
x_series = np.reshape(valid_x_array[idx,:,1], (1, 1380,1))
y_series = np.reshape(valid_y_array[idx,:,1], (1, 120, 1))
model.fit(x_series, y_series, epochs = 10, batch_size = 32)
# -
model.summary()
model.save('my_model.h5')
model = tf.keras.models.load_model('./my_model.h5')
# ## 3.3. 훈련한 모델 테스트
new_valid_x_df = train_x_df[train_x_df.sample_id > 100]
# # 4. valid_pred_array 로부터 buy_quantity, sell_time 구하기
# + tags=[]
#valid data predict
train_pred_array = np.zeros([100, 120, 1])
for idx in tqdm(range(valid_x_array.shape[0])): #1380
train_x = np.reshape(valid_x_array[idx,:,1], (1, 1380,1))
# test_x = tf.reshape(test_x_df.iloc[:,3].values, [-1, 1380, 1]) #open 값의 3차원형
preds = model.predict(train_x)
train_pred_array[idx,:] = preds
# model = ARIMA(x_series, order=(5,1,1))
# fit = model.fit()
# -
train_pred_array.shape
# +
#valid_pred_array 3차원에서 2차원으로 바꾸기
new_pred_array = np.zeros([100, 120])
for idx in tqdm(range(train_pred_array.shape[0])):
val_open = train_pred_array[idx, :, 0]
new_pred_array[idx, :] = val_open
new_pred_array
# -
new_pred_array.shape
def array_to_submission(pred_array):
submission = pd.DataFrame(np.zeros([pred_array.shape[0],2], np.int64),
columns = ['buy_quantity', 'sell_time'])
submission = submission.reset_index()
submission.loc[:, 'buy_quantity'] = 0.1
buy_price = []
for idx, sell_time in enumerate(np.argmax(pred_array, axis = 1)):
buy_price.append(pred_array[idx, sell_time])
buy_price = np.array(buy_price)
submission.loc[:, 'buy_quantity'] = (buy_price > 1.15) * 1
submission['sell_time'] = np.argmax(pred_array, axis = 1)
submission.columns = ['sample_id','buy_quantity', 'sell_time']
return submission
# + tags=[]
valid_submission = array_to_submission(new_pred_array)
valid_submission
# -
# 전체 300가지 sample에 대해
# 9가지 case에서 115% 이상 상승한다고 추론함.
valid_submission.buy_quantity.value_counts()
# # 5. 실제 test x 데이터로 test y 예측
# + tags=[]
#predict
test_pred_array = np.zeros([529, 120, 1])
for idx in tqdm(range(test_x_array.shape[0])): #529
test_x = np.reshape(test_x_array[idx,:,1], (1, 1380,1)) #각 open 값의 3차원형
# test_x = tf.reshape(test_x_df.iloc[:,3].values, [-1, 1380, 1]) #open 값의 3차원형
preds = model.predict(test_x)
test_pred_array[idx,:] = preds
# model = ARIMA(x_series, order=(5,1,1))
# fit = model.fit()
# -
preds.shape
# + tags=[]
test_pred_array.shape
# +
#test_pred_array 3차원에서 2차원으로 바꾸기
final_pred_array = np.zeros([529, 120])
for idx in tqdm(range(test_pred_array.shape[0])):
val_open = test_pred_array[idx, :, 0]
final_pred_array[idx, :] = val_open
final_pred_array
# -
final_pred_array
submission= array_to_submission(final_pred_array)
submission
# ## 3) 샘플 확인하기
def plot_series(x_series, y_series):
#입력 series와 출력 series를 연속적으로 연결하여 시각적으로 보여주는 코드 입니다.
plt.plot(x_series, label = 'input_series')
plt.plot(np.arange(len(x_series), len(x_series)+len(y_series)),
y_series, label = 'output_series')
plt.axhline(1, c = 'red')
plt.legend()
len(test_pred_array[500])
# +
idx = 300
# train data 중 sample_id 1121에 해당하는 x_series로 arima 모델을 학습한 후
# y_sereis를 추론
x_series = train_x_array[idx,:,1]
y_series = train_y_array[idx,:,1]
# pred_dim1 = np.zeros(120)
# pred_dim1 = preds[0,,0]
# pred_dim1.shape
plot_series(x_series, y_series)
plt.plot(np.arange(1380, 1380+120), test_pred_array[idx], label = 'prediction')
plt.legend()
plt.show()
# -
# 제대로 학습되지 않고 있다는 것을 알 수 있다.
# + active=""
# 정확도 측정(수정필요)
# # Test RMSE 측정하기
# from sklearn.metrics import mean_squared_error, r2_score
# from math import sqrt
#
# rmse = sqrt(mean_squared_error(pred_y, test_y))
# print(rmse)
# -
submission.to_csv('submission1.csv', index = False)
| Trials/Deep_LSTM-DESKTOP-EF7794D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# +
import pandas as pd
old_data = pd.DataFrame.from_csv('./Documents/Cornell/Courses/MPS Project/polarity_specific.csv')
list_pos = []
list_neg = []
list_neu = []
list_com = []
for i in range(1,old_data.shape[1]):
if i % 4 == 1:
list_pos.append(old_data.ix[:,i])
elif i % 4 == 2:
list_neg.append(old_data.ix[:,i])
elif i % 4 == 3:
list_neu.append(old_data.ix[:,i])
elif i % 4 == 0:
list_com.append(old_data.ix[:,i])
positive = pd.DataFrame(list_pos).T
# del positive['Date']
# del positive[0]
negative = pd.DataFrame(list_neg).T
neutral = pd.DataFrame(list_neu).T
compound = pd.DataFrame(list_com).T
output = pd.DataFrame([positive.mean(axis=1),negative.mean(axis=1),neutral.mean(axis=1),compound.mean(axis=1)],
['pos','neg','neutral','compound']).T
# positive.to_csv("./Documents/Cornell/Courses/MPS Project/Matlab/positive.csv")
# negative.to_csv("./Documents/Cornell/Courses/MPS Project/Matlab/negative.csv")
# neutral.to_csv("./Documents/Cornell/Courses/MPS Project/Matlab/neutral.csv")
# compound.to_csv("./Documents/Cornell/Courses/MPS Project/Matlab/compound.csv")
output
# -
# ori_price = pd.DataFrame.from_csv('./Documents/Cornell/Courses/MPS Project/DJIA_table.csv',index_col=0)
# ori_price
positive.describe()
negative.describe()
neutral.describe()
compound.describe()
| _data_analysis/ipynb/ProcessPolarityData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function
import h5py
import numpy as np
import matplotlib.pyplot as plt
from sklearn.utils import class_weight
from sklearn.metrics import classification_report
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Input, Concatenate, Reshape
from keras.layers import Conv2D, MaxPooling2D, AveragePooling2D
from keras.utils import plot_model
from keras.models import Model
from keras.optimizers import adadelta as ada
from PIL import Image
import matplotlib.pyplot as plt
import pandas as pd
import copy
import pydot
from keras.utils import multi_gpu_model
from keras import backend as K
import gc
import time
# %matplotlib inline
# +
hdf5_path = 'Data/dataset.hdf5'
subtract_mean = True
hdf5_file = h5py.File(hdf5_path, "r")
if subtract_mean:
mm = hdf5_file["train_mean"][0, ...]
mm = mm[np.newaxis, ...]
data_num = hdf5_file["train_flow"].shape[0]
num_classes = 2
epochs = 30
flow_rows, flow_cols = 298, 17
x_train = hdf5_file["train_flow"][:,...]
if subtract_mean:
x_train -= mm
y_train = hdf5_file["train_labels"][:, ...]
hdf5_file.close()
hdf5_path = 'Data/dataset-IoT.hdf5'
hdf5_file = h5py.File(hdf5_path, "r")
x_test = hdf5_file["IoT_flow"][:,...]
if subtract_mean:
x_test -= mm
y_test = hdf5_file["labels"][:, ...]
hdf5_file.close()
class_weights = class_weight.compute_class_weight('balanced',
np.unique(y_train),
y_train)
d_class_weights = dict(enumerate(class_weights))
packets = 200
x_train = x_train[:,:packets,:,:]
x_test = x_test[:,:packets,:,:]
input_shape = (x_train.shape[1], x_train.shape[2], x_train.shape[3])
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# +
AccScores = []
Times = []
LossScores = []
#AucScores = []
#F1Scores = []
#PrecisionScores = []
t2 = time.time()
for i in range (1,50):
batch_size = 1024
epochs = 30
filters= 32
kernel_size= (3,3)
activations= "sigmoid"
pool_size= (2,2)
dropout = 0.29104403739531765
lr = 0.001433773291970261
rmsprop = keras.optimizers.RMSprop(lr=lr)
optim = rmsprop
layers = 2
model = Sequential()
model.add(Conv2D(filters, kernel_size=kernel_size,activation=activations, input_shape=input_shape,padding = "same"))
for i in range(layers-1):
model.add(Conv2D(filters,kernel_size=kernel_size, activation=activations, padding = "valid"))
model.add(MaxPooling2D(pool_size=(pool_size)))
model.add(Flatten())
model.add(Dropout(dropout))
model.add(Dense(64, activation='relu'))
model.add(Dense(32, activation='relu'))
model.add(Dropout(dropout))
model.add(Dense(num_classes, activation='softmax'))
#model.summary()
try:
model = multi_gpu_model(model, gpus = 4)
except:
pass
model.compile(loss='binary_crossentropy', optimizer=optim, metrics=['accuracy'])
t0 = time.time()
model.fit(x_train,y_train, batch_size=batch_size, epochs=epochs, verbose=0, class_weight=class_weights, shuffle=True)
scores = model.evaluate(x_test, y_test, verbose=0)
t1 = time.time()
total_run = t1-t0
Times.append(total_run)
print("%s: %.5f%%" % (model.metrics_names[1], scores[1]*100))
print("%s: %.5f%%" % (model.metrics_names[0], scores[0]))
print('Fit and Evaluate Time:',total_run)
AccScores.append(scores[1] * 100)
LossScores.append(scores[0])
del model
gc.collect()
K.clear_session()
gc.collect()
print("%.5f%% (+/- %.5f%%)" % (np.mean(AccScores), np.std(AccScores)))
print("%.5f%% (+/- %.5f%%)" % (np.mean(LossScores), np.std(LossScores)))
print("%.5f%% (+/- %.5f%%)" % (np.mean(Times), np.std(Times)))
t3 = time.time()
total_time = t3-t2
print('Time to do everything:',total_time)
# -
AccScores
LossScores
Times
| CNN 2D Packet Select.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Neuroinformatics: Using Python for literature searches
# ### MAY 24, 2022
# ### Guest lecture by <NAME>
#
# In this notebook, you will learn to:
# * Identify the conceptual and technical tools used to conduct informatics research (e.g. APIs, ontologies, bioentrez, BLAST)
# * Identify the structure and use of json format
# * Define MESH terms & describe their role in informatics research
# * Explain the role and importance of informatics research
# * Conduct a pubmed search using bioentrez
# -
# ## Setup
# We'll need many new packages that aren't in our DataHub for today's lab. To get everything setup, run our setup script below.
# +
# Install packages that do not exist in DataHub
# ! pip install xmljson
# ! pip install xmltodict
# ! pip install Biopython
# Import many packages!
# %run informatics_module/informatics_setup.py
# -
# Since we just did several things behind the scenes, let's take a look at the packages we imported.
# %whos
# ## Accessing NCBI databases with Biopython
# Biopython is a set of freely available tools for biological computation written in Python. It contains a collection of python modules to search to deal with DNA, RNA & protein sequence operations such as reverse complementing of a DNA string, finding motifs in protein sequences, etc.
#
# Bio.Entrez is the module within the BioPython package that provides code to access NCBI over the World Wide Web to retrieve various sorts of information. This module provides a number of functions which will return the data as a handle object. This is the standard interface used in Python for reading data from a file and provides methods or offers iteration over the contents line by line.
#
# ### Bio.Entrez is not the only sub-module in Biopython. [Other packages include](https://biopython.org/docs/1.75/api/index.html):
# - Bio.GEO - Access to data from the Gene Expression Omibus database.
# - Bio.KEGG - Access to data from the KEGG database.
# - Bio.motifs - Access to tools for sequence motif analysis.
#
# ### Best practice/required information:
# - Always provide your email with Entrez.email = "youremail.com"
# ### Functions used in Bio.entrez
#
# Bio.Entrez has a ton of different functions. We'll use a few (highlighted) in our notebook today. **Read more about these functions on [the website](https://www.ncbi.nlm.nih.gov/books/NBK25499/).**
#
# - **eInfo** - Provides the number of records indexed in each field of a given database, the date of the last update of the database, and the available links from the database to other Entrez databases.
# - **eSearch** - Responds to a text query with the list of matching **Entrez Unique Identifier (UIDs)** in a given database (for later use in ESummary, EFetch or ELink), along with the term translations of the query.
# - ePost - Accepts a list of UIDs from a given database, stores the set on the History Server, and responds with a query key and web environment for the uploaded dataset.
# - efetch - Retrieves records in the requested format from a list of one or more primary IDs or from the user’s environment
# - elink - Checks for the existence of an external or Related Articles link from a list of one or more primary IDs. Retrieves primary IDs and relevancy scores for links to Entrez databases or Related Articles; creates a hyperlink to the primary LinkOut provider for a specific ID and database, or lists LinkOut URLs and Attributes for multiple IDs.
# - eSummary - Retrieves document summaries from a list of primary IDs or from the user’s environment.
# - egQuery - Provides Entrez database counts in XML for a single search using Global Query.
# - eSpell - Retrieves spelling suggestions.
# - eCitmatch - Retrieves PubMed IDs (PMIDs) that correspond to a set of input citation strings.
# - **read** - Parses the XML results returned by any of the above functions.
#
#
# <div class="alert alert-success">
#
# **Task**: Write your email below.
#
# </div>
### provide your email as a string
Entrez.email = ...
# As previously mentioned in the slides, the BioEntrez package provides access to multiple biomedical databases. Below, we'll use `einfo` and `read` to show the list of possible databases.
# +
### access API to search for a list of databases in Bio.Entrez
# declare a variable (e.g.handle) where the results will be stored
# pass within the Entrez.einfo() function.
handle = Entrez.einfo()
record = Entrez.read(handle)
print (record)
# -
# <div class="alert alert-success">
#
# **Task**: Look for info about PubMed by specifying the `db="pubmed"` argument in our call to `Entrez.einfo` (otherwise, same code as above!).
#
# </div>
# Look for info about a database - PubMed
# As you can see, this is a very big, complicated dictionary! It actually has multiple layers, kind of like an onion. If we just look for the keys, we'll only find one thing:
record.keys()
# So the first and only dictionary key is `DbInfo`. We need to then dig *into* the DbInfo dictionary for more information.
# <div class="alert alert-success">
#
# **Task**:
# 1. Print the list of keys for the `DbInfo` dictionary.
# 2. Print the `FieldList` within the DbInfo. **What kind of object is this? What is contained in it?**
# 3. Using `FieldList`, figure out how many authors (`AUTH`), (`JOUR`), and MESH terms (`MESH`) are in this database.
#
# </div>
# Work with the record dictionary
# ### What are MESH terms?
# - abbreviation for Medical Subject Headings
# - Controlled vocabulary thesaurus produced by the National Library of Medicine (NLM)
# - It consists of sets of terms naming descriptors in a hierarchical structure that permits searching at various levels of specificity.
#
# *** Search for MESH terms - https://www.nlm.nih.gov/mesh/meshhome.html
# + [markdown] slideshow={"slide_type": "slide"}
# .
#
# <NAME>., <NAME>., & <NAME>. (2017). Implementation of a classification server to support metadata organization for long term preservation systems. VOEB-Mitteilungen, 70(2), 225–243. https://doi.org/10.31263/voebm.v70i2.1897
# -
# <div class="alert alert-success">
#
# **Task**:
#
# 1. Create a pubmed search using Bio.Entrez. You'll use the `esearch` function first. This will return the top 20 results by default. You'll use the same syntax as before, first defining a handle, then using the read function. [This website may help!](https://biopython-tutorial.readthedocs.io/en/latest/notebooks/09%20-%20Accessing%20NCBIs%20Entrez%20databases.html)
#
# 2. How do the search results differ from the PubMed user interface?
#
# 3. Assign the `IdList` field to a variable `paper_ids`.
#
# 4. Use `paper_ids` and `efetch` to fetch the abstracts for these papers, using this syntax:
#
# ```
# abstracts = Entrez.efetch(db='pubmed', id=paper_ids, rettype='abstract')
# abstracts.read()
# ```
#
#
# </div>
# Create your pubmed search here
# Now, let's do a more complicated search using the same basic concepts.
#
# We'll define a function below to iterate over multiple terms. Then, we will locate PubMed articles that may identify gene product in primary cell types in the hippocampus and then pull the text from articles that are accessible in PubMed Central.
# +
# Define the function
def comb_list(brain_region, cell_type, method):
'''iterate over terms given - in this case brain region, cell type and method'''
all_list = list(product(brain_region, method, cell_type))
new_list = [list(i) for i in all_list]
terms = [" AND ".join(i) for i in new_list]
return terms
# +
# Use the function
brain_region = ["hippocampus"]
cell_type = ["CA1 pyramidal cell", "CA2 pyramidal cell", "CA3 pyramidal cell", "CA4 Pyramidal Cell", "Dentate Gyrus Granule Cell", "Dentate Gyrus Basket Cell", "CA1 Basket Cell"]
method = ["rna seq* ","microarray","in situ hybridization", "polymerase chain reaction"]
terms = comb_list(brain_region, cell_type, method)
terms
# -
# Below, we'll use a script (`informatics_functions.py`) to define two short functions and a couple lengthy functions in the file `informatics_functions.py`.
#
# <div class="alert alert-success">
#
# Take a look at the `findformat_abstract` function within the script closely -- what is it doing?
#
# </div>
# Run functions script and show functions that we have available
# %run informatics_module/informatics_functions.py
# %whos
# +
#get all abstracts
new_abstracts = {}
gene_abstracts = findformat_abstract(terms)
#only get the abstracts with pmc ids
pmc_abstracts = {k: v for k, v in gene_abstracts.items() if len(v['PMC']) > 0}
# This is a way to make a copy of dictionary, as a backup in case
# Deepcopy () copies all the elements of an object as well as the memory location that contains data rather than containing the data itself.
gene_abstract_cp = deepcopy(gene_abstracts)
pmc_abstract_cp = deepcopy(pmc_abstracts)
# -
# Print gene abstracts
gene_abstracts
print ('original count', len(gene_abstracts))
print('PMC:', len(pmc_abstracts))
print('difference =', len(gene_abstracts)-len(pmc_abstracts))
# <div class="alert alert-success">
#
# The length of `pmc_abstracts` is less than that of `gene_abstract`. Why is that?
#
# </div>
# ## Read results as a dataframe
# It's difficult to visually parse dictionaries. Thankfully we have another tool at our disposal: pandas.
#
# <div class="alert alert-success">
#
# **Task**: Turn `gene_abstract` into a pandas dataframe called `gene_abstract_df`.
#
# </div>
# Turn gene_abstract into a df
# Hmm, it would make a lot more sense if each paper had its own row -- that's how we typically conceptualize dataframes, with each row as a different observation, patient, cell, etc. We can **transpose** the dataframe using the [`transpose`](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.transpose.html) (or `T` for short) method.
#transposed data frame
papers_df = gene_abstract_df.T
papers_df
# <div class="alert alert-success">
#
# **Task**: Using the `iloc` method, view *just* the first abstract.
#
# </div>
# Look at the first abstract here
#format the Methods and Results section
g_updated_records = getTexts(gene_abstract)
#make another copy of the file since a lot of information is in here.
pmc_papers = deepcopy(g_updated_records)
pmc_papers
# ## Save results as a json & excel file
#
# Below, we'll save our findings as both a json and an Excel file.
# **Json?**
# JavaScript Object Notation (JSON) is a standardized format commonly used to transfer data between systems and used by a lot of databases and APIs.
# Like Python dictionaries, it represents objects as name/value pairs.
#
# +
#save file as a json file
with open('g_updated_records.json', 'w') as outfile:
json.dump(g_updated_records, outfile)
#read in json file
with open('g_updated_records.json', 'r') as newfile:
g_updated_records = json.load(newfile)
#save file to an excel file - save file as pandas dataframe, save to excel
df_updated_records = dp = pd.DataFrame(gene_abstract).T
g_update_records.to_excel('g_updated_records.json.xlsx')
| 14-Informatics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: html
# language: python
# name: html
# ---
# # Multi-fidelity demo
#
# In this notebook, we show how we can already obtain a good idea of where the best hyperparameters lie with low fidelity models.
#
# Low fidelity models are those that would deliver a sub-optimal performance but at a much lower computational cost.
# +
import datetime
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import (
RandomizedSearchCV,
train_test_split,
)
# the models to optimize
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
# +
# load dataset
breast_cancer_X, breast_cancer_y = load_breast_cancer(return_X_y=True)
X_train = pd.DataFrame(breast_cancer_X)
y_train = pd.Series(breast_cancer_y).map({0:1, 1:0})
X_train.head()
# +
# the target:
# percentage of benign (0) and malign tumors (1)
y_train.value_counts() / len(y_train)
# -
# ## Multi-fidelity
#
# ### data size
# +
# set up the model
svm_model = SVC(kernel='rbf', random_state=0)
# determine the hyperparameter space
# we will search over 2 hyperparameters for this demo
param_grid = dict(
C = stats.loguniform(0.001, 100),
gamma = stats.loguniform(0.001, 100),
)
# +
# set up the search
search = RandomizedSearchCV(svm_model,
param_grid,
scoring='accuracy',
cv=2,
n_iter = 50,
random_state=10,
n_jobs=4,
refit=False,
)
# +
# determine the best hyperparameters and performance
# searching over datasets of different sizes
# that is, different fidelities
print('Darker orange means better accuracy')
for size in [0.1, 0.2, 0.4, 0.8]:
# subsample the data
n_samples = int(size * len(X_train))
X_train_ = X_train.sample(n_samples, random_state=0)
y_train_ = y_train[X_train_.index]
# search
ts = datetime.datetime.now()
search.fit(X_train_, y_train_)
td = (datetime.datetime.now()-ts).microseconds
results = pd.DataFrame(search.cv_results_)
# plot results
plt.scatter(results['param_gamma'],
results['param_C'],
c=results['mean_test_score'],
cmap='Wistia',
)
plt.title(f"Data fraction: {size}, time: {td}")
plt.xlabel('gamma')
plt.ylabel('C')
plt.show()
# -
# For this particular dataset, utilizing 20% of the data already tells us where the best hyperparameter combination lies.
# ## number of estimators
# +
# set up the model
rf = RandomForestClassifier(random_state=0)
# determine the hyperparameter space
# we will search over 2 hyperparameters for this demo
param_grid = dict(
min_samples_split=stats.uniform(0, 1),
max_depth=stats.randint(1, 5),
)
# +
# set up the search
search = RandomizedSearchCV(rf,
param_grid,
scoring='accuracy',
cv=2,
n_iter = 50,
random_state=10,
n_jobs=4,
refit=False,
)
# +
# determine the best hyperparameters and performance
# searching over models with different number of trees
# that is, different fidelities
print('Darker orange means better accuracy')
for n_estimators in [5, 10, 20, 50]:
# fix the n_estimators parameter
rf.set_params(**{'n_estimators': n_estimators})
# search
ts = datetime.datetime.now()
search.fit(X_train, y_train)
td = (datetime.datetime.now()-ts).seconds
results = pd.DataFrame(search.cv_results_)
# plot results
plt.scatter(results['param_min_samples_split'],
results['param_max_depth'],
c=results['mean_test_score'],
cmap='Wistia',
)
plt.title(f"n_estimators: {n_estimators}, time: {td}")
plt.xlabel('min_samples_split')
plt.ylabel('max_depth')
plt.show()
# -
# We see that a random forest with 5 estimators, already tells us where the other hyperparameters best values are.
#
# For this particular example, if we use the number of estimators as the fidelity, then, we can't optimize this hyperparameter per se. But for the purpose of this demo, it helps us understand the different ways in which we can train lower fidelity algorithms.
| Section-08-Multi-fidelity-Optimization/01-Multi--fidelity-demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pickle
import pandas as pd
# -
data_dir = 'data_0419_0'
with open(os.path.join(data_dir, 'pickled_model_gbm_smote'), 'rb') as f:
model = pickle.load(f)
df = pd.DataFrame({'cust_mean_buy_price':{'0':11.61,'1':11.61,'2':11.61},'cust_total_coupons':{'0':701,'1':701,'2':701},'cust_mean_discount':{'0':9.71,'1':9.71,'2':9.71},'cust_unique_products':{'0':1309,'1':1309,'2':1309},'cust_unique_products_coupon':{'0':485,'1':485,'2':485},'cust_total_products':{'0':2328,'1':2328,'2':2328},'coupon_discount':{'0':57,'1':18,'2':15},'coupon_how_many':{'0':1,'1':4,'2':2},'coupon_mean_prod_price':{'0':18.23,'1':12.39,'2':9.22},'coupon_prods_avail':{'0':641,'1':4,'2':1},'cust_age_young':{'0':0,'1':0,'2':0},'cust_age_mid':{'0':0,'1':0,'2':0},'cust_age_old':{'0':1,'1':1,'2':1},'cust_gender_F':{'0':1,'1':1,'2':1},'cust_gender_M':{'0':0,'1':0,'2':0},'coupon_type_buy_all':{'0':0,'1':1,'2':0},'coupon_type_buy_more':{'0':0,'1':0,'2':1},'coupon_type_department':{'0':1,'1':0,'2':0},'coupon_type_just_discount':{'0':0,'1':0,'2':0},'coupon_dpt_Boys':{'0':0,'1':0,'2':0},'coupon_dpt_Girls':{'0':0,'1':0,'2':0},'coupon_dpt_Men':{'0':0,'1':0,'2':0},'coupon_dpt_Sports':{'0':0,'1':0,'2':0},'coupon_dpt_Women':{'0':0,'1':0,'2':0}})
df
preds = model.predict_proba(df)
preds
| training-with-artificial-data/delete_me.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
# -
# # Question
#
# We observed that the credit amount of people with bad credibility (Risk = 1) is approximately 1,000 units above that of people with good credibility (Risk = 0). Is this statistically meaningful?
# ### 1. Import Dataset
german_dataset = pd.read_csv('../statistics/german_dataset.csv')
german_dataset.head()
# ### 2. Find Each Group's Mean
# +
mean_good = german_dataset[german_dataset['Risk'] == 0]['Credit amount'].mean()
mean_bad = german_dataset[german_dataset['Risk'] == 1]['Credit amount'].mean()
print('Credit amount of Good:', mean_good)
print('Credit amount of bad:', mean_bad)
# -
#find length of each groups
print(len(german_dataset[german_dataset['Risk'] == 0]))
print(len(german_dataset[german_dataset['Risk'] == 1]))
# ### 3. Permutation Test
# +
len_good = len(german_dataset[german_dataset['Risk'] == 0])
len_bad = len(german_dataset[german_dataset['Risk'] == 1])
len_good, len_bad
# +
random_generator = np.random.RandomState(42)
pm = []
total_indice = np.arange(len(german_dataset))
for i in range(1000):
copy_indice = total_indice.copy()
random_generator.shuffle(copy_indice)
perm_a = german_dataset.iloc[copy_indice[:len_good]]
perm_b = german_dataset.iloc[copy_indice[len_good:]]
pm.append(perm_b['Credit amount'].mean() - perm_a['Credit amount'].mean())
# -
sns.distplot(pm, kde=False)
# ### 4. Conclusion
#
# The observed difference between the two groups (approximately $1,000) seems to have a statistical significance. Looking at the graph above, it is hard to say that the observed difference occured to a mere chance
| algorithm_exercise/statistics/permutation_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.5
# language: python
# name: python3
# ---
# # DAT257x: Reinforcement Learning Explained
#
# ## Lab 2: Bandits
#
# ### Exercise 2.1A: Greedy policy
# +
import numpy as np
import sys
if "../" not in sys.path:
sys.path.append("../")
from lib.envs.bandit import BanditEnv
from lib.simulation import Experiment
# -
# Let's define an interface of a policy. For a start, the policy should know how many actions it can take and able to take a particular action given that policy
#Policy interface
class Policy:
#num_actions: (int) Number of arms [indexed by 0 ... num_actions-1]
def __init__(self, num_actions):
self.num_actions = num_actions
def act(self):
pass
def feedback(self, action, reward):
pass
# Now let's implement a greedy policy based on the policy interface. The greedy policy will take the most rewarding action (i.e greedy). This is implemented in the act() function. In addition, we will maintain the name of the policy (name), the rewards it has accumulated for each action (total_rewards), and the number of times an action has been performed (total_counts).
#Greedy policy
class Greedy(Policy):
def __init__(self, num_actions):
Policy.__init__(self, num_actions)
self.name = "Greedy"
self.total_rewards = np.zeros(num_actions, dtype = np.longdouble)
self.total_counts = np.zeros(num_actions, dtype = np.longdouble)
def act(self):
current_averages = np.divide(self.total_rewards, self.total_counts, where = self.total_counts > 0)
current_averages[self.total_counts <= 0] = 0.5 #Correctly handles Bernoulli rewards; over-estimates otherwise
current_action = np.argmax(current_averages)
return current_action
def feedback(self, action, reward):
self.total_rewards[action] += reward
self.total_counts[action] += 1
# We are now ready to perform our first simulation. Let's set some parameters.
evaluation_seed = 8026
num_actions = 5
trials = 10000
distribution = "bernoulli"
# Now, put the pieces together and run the experiment.
env = BanditEnv(num_actions, distribution, evaluation_seed)
agent = Greedy(num_actions)
experiment = Experiment(env, agent)
experiment.run_bandit(trials)
# Observe the above results and answer the lab questions!
| Module 2/Ex2.1A Greedy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
path = "./data/moses/test.txt"
pd.read_csv(path,squeeze=True).astype(str).tolist()
# +
def a():
print("aaaa")
def b():
print("bbbb")
def c(x,y):
return x+y
def d(q,w,e,m,n):
q()
z=w(m,n)
e()
return z
# -
p=d(a,c,b,1,2)
print(p)
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ig1HGm_sXvHx"
# # Twitter Sentiment Analysis - Feed-forward Neural Networks
# ## <div> <NAME> </div>
# + id="lQ0xV4tFJZTI" colab={"base_uri": "https://localhost:8080/"} outputId="0e4173ed-ba10-494f-f938-29de90589898"
# !pip install d2l==0.15.0
# + id="L3VQYcefW8Hf"
import pandas as pd
import numpy as np
import re
import warnings
import torch
import matplotlib.pyplot as plt
import pickle
warnings.filterwarnings('ignore')
# + [markdown] id="sStnn3cnXAOM"
# ## Load Data
# + id="FyzcY9_IXCal" colab={"base_uri": "https://localhost:8080/"} outputId="dd831744-f08a-4cac-f67c-87e89f813152"
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="_LM1FQO7XNK6"
# #### Create a dataframe from SentimentTweets.csv file data
# + id="Z54N_yxEXG2r" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="5accc59f-e159-4187-c8ae-38c8c59aafa8"
df=pd.read_csv('gdrive/My Drive/Colab Notebooks/SentimentTweets.csv', usecols=['target','id','date','flag','user','text'])
df.head()
# + id="9Cu7EnZPXad-" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="f69299ef-1ab4-422f-a1c3-a8da7b775876"
df.drop(columns=['id', 'date', 'flag', 'user'], axis=1, inplace=True) #drop useless columns
df.head()
# + id="r64ZFbZwXdIc"
# split 'target' and 'text' columns
X = df[['text']]
y = df[['target']]
# + id="Cv0fmkO5Xela"
from sklearn.model_selection import train_test_split
# get train and test dataframes
train_X, test_X, train_Y, test_Y = train_test_split(X, y, test_size=0.2, stratify=df['target'], random_state = 42)
# + [markdown] id="4qt2539dXho5"
# #### Display train and test sets after split
# + id="EM1VLqxVXgI1" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="7bf0bf56-be11-4705-ee04-5de35d578db5"
train_X.head()
# + id="OBaNuTQZXksQ" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="0bcf72ee-ccae-44fe-90c4-c79efaab432c"
test_X.head()
# + [markdown] id="4ClyD5k4Xl61"
# ## Labels Pre-processing
# + id="T6o-BhRXXnlJ"
# replace 4 with 1 to create binary labels
train_Y = train_Y.replace(4,1)
test_Y = test_Y.replace(4,1)
# create numpy arrays for sets' labels
train_y = np.asarray(train_Y['target'].tolist())
test_y = np.asarray(test_Y['target'].tolist())
# covert numpy arrays to torch tensors
train_y, test_y = map(torch.tensor, (train_y, test_y))
# covert tensors data to float
train_y, test_y = train_y.float(), test_y.float()
# + [markdown] id="ZzDrAI8xXpzc"
# ## Data Pre-processing
# + [markdown] id="JhfmSB4kcQDV"
# #### Load cleansed data from csv
#
# + id="YoFIMphscMsR"
# restore processed data
train_X = pd.read_csv('gdrive/My Drive/Colab Notebooks/CleanedTrain.csv', usecols=['text','processedText'])
test_X = pd.read_csv('gdrive/My Drive/Colab Notebooks/CleanedTest.csv', usecols=['text','processedText'])
# + [markdown] id="a8UMJDrgchTW"
# ### Cleanse data
# + id="RtrcZJSWXsb7"
# function that removes all @mentions, links and non alphabetic strings
def clean_content(text):
text = re.sub(r'@[A-Za-z0-9_]+', '', text) # remove text with @ prefix
text = re.sub(r'http\S+', '', text) # remove text with http prefix (links)
text = re.sub(r'www\S+', '', text) # remove text with www prefix (links)
text = re.sub(r'\\\w+', '', str(text)) # remove text after backslash
text = re.sub(r'\b\w{1,2}\b', '', text) # remove text containing 2 or less characters
text = ''.join(ch for ch in text if ch.isalpha() or ch == ' ')
text = text.lower() # convert text into lowercase
return text
# + id="TRZXeA2nXt8L"
# create a column for each set containing the processed text data
for index, row in train_X.iterrows():
train_X.loc[index,'processedText'] = clean_content(train_X.loc[index,'text'])
for index, row in test_X.iterrows():
test_X.loc[index,'processedText'] = clean_content(test_X.loc[index,'text'])
# + [markdown] id="Pt5LUmFqXwzV"
# #### Display train and test sets after text cleansing
# + id="YPZLdglyXvZh" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="8ba51626-3f1f-4625-ca79-c892cb7c9325"
train_X.head()
# + id="8_dyGR4IXzTG" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="45c16a58-b419-4d90-d94f-667e9ecfb162"
test_X.head()
# + id="OS56PPZjg6kT"
# # store processed data to a csv file
# train_X.to_csv('gdrive/My Drive/Colab Notebooks/CleanedTrain.csv', index = True, header=True)
# test_X.to_csv('gdrive/My Drive/Colab Notebooks/CleanedTest.csv', index = True, header=True)
# + [markdown] id="gdbg3VI4X2M_"
# ### Tf-idf Vectorization
# + id="kH2OoLFGX0n2"
from sklearn.feature_extraction.text import TfidfVectorizer
tfidfVectorizer = TfidfVectorizer(max_df=0.99, min_df=1, stop_words='english', max_features=150)
# apply tf-idf transformation to both train and test sets data
tr_tfidf = tfidfVectorizer.fit_transform(train_X['processedText'])
te_tfidf = tfidfVectorizer.transform(test_X['processedText'])
# insert transformed data to numpy arrays for both training and test sets
train_tfidf = tr_tfidf.toarray()
test_tfidf = te_tfidf.toarray()
# + [markdown] id="7JW0sbRl3Wza"
# #### Convert input vectors to torch tensors
# + id="-s2GcdA13RV6"
# covert numpy arrays to torch tensors
train_tfidf, test_tfidf = map(torch.tensor, (train_tfidf, test_tfidf))
# covert tensors data to float
train_tfidf, test_tfidf = train_tfidf.float(), test_tfidf.float()
# + [markdown] id="Lu0tIeExpLPd"
# ### GloVe Pre-trained Embeddings
# + id="L91p4LurpJ2Z"
from torchtext.data import Field
from torchtext.vocab import GloVe
# + [markdown] id="oo6oCRsY2Fxx"
# #### Load glove embeddings from disk
# + id="0AoRegLWEdka"
with open('gdrive/My Drive/Colab Notebooks/train_glove.pkl', 'rb') as fp:
train_glove = pickle.load(fp)
with open('gdrive/My Drive/Colab Notebooks/test_glove.pkl', 'rb') as fp:
test_glove = pickle.load(fp)
# + [markdown] id="RITdTeFH3HIr"
# #### Convert input vectors to torch tensors
# + id="SpCzi6Bo3HIx"
#covert numpy arrays to torch tensors
train_glove, test_glove = map(torch.tensor, (train_glove, test_glove))
# covert tensors data to float
train_glove, test_glove = train_glove.float(), test_glove.float()
# + [markdown] id="Qd2UmWYYd5PR"
# ### Produce embedding input vectors
# + id="fJj7sins3DWE" colab={"base_uri": "https://localhost:8080/"} outputId="754e12ca-0f1d-4c1f-9dcb-c3de13ca9a22"
emb_size=100
embedding = GloVe(name='6B', dim=emb_size) # use "glove.6B.100d" as embedding
# + id="MQJ02_a_3Mzv"
# function to tokenize preprocessed text and add the necessary padding tokens
def tokenize(df, padding):
text_field = Field(
sequential=True,
tokenize='spacy',
fix_length=padding,
lower=True
)
label_field = Field(sequential=False, use_vocab=False)
preprocessed_text = df['processedText'].apply(
lambda x: text_field.preprocess(x)
)
return preprocessed_text
# + id="yL932Qp2rW1K"
# function that creates a mean vector for a series of tokens
def tokens_to_vector(tokens, embedding, emb_size):
vectors = []
for token in tokens:
try:
vectors.append(embedding[token].cpu().detach().numpy()) # add word's vector to list if word belongs to embedding dictionary
except KeyError:
vectors.append(np.zeros(emb_size).astype('float32') ) # else, add a zero vector to the list to represent unknown words
total = np.zeros(emb_size).astype('float32')
# compute the mean vector
for v in vectors:
total += v
mean = total / len(vectors)
return mean.tolist() # return vector in list format
# + id="5rNqilsfHFCI"
padding = train_X.processedText.map(lambda x: len(x)).max() # find the largest 'processedText' in dataframe to determine the padding's length
# + id="BKVv3EeVno2f" colab={"base_uri": "https://localhost:8080/"} outputId="946b34e7-8aae-4ab9-e<PASSWORD>"
# apply tokenization to both train and test sets
train_toks = tokenize(train_X, padding)
test_toks = tokenize(test_X, padding)
# + id="_Eqxp7EkBBOu"
tr_glove = []
# create a list of vectors for every series of tokens in train set
for i, tokens in enumerate(train_toks):
tr_glove.append(tokens_to_vector(tokens, embedding, 100))
te_glove = []
# create a list of vectors for every series of tokens in test set
for i, tokens in enumerate(test_toks):
te_glove.append(tokens_to_vector(tokens, embedding, 100))
# + id="S-e5AoOBFuSX"
# convert list of vectors to numpy array
train_glove = np.array(tr_glove)
test_glove = np.array(te_glove)
# + id="4Gauk8vx3QtE"
# # store glove embeddings to disk
# with open('gdrive/My Drive/Colab Notebooks/train_glove.pkl', 'wb') as fp:
# pickle.dump(train_glove, fp)
# with open('gdrive/My Drive/Colab Notebooks/test_glove.pkl', 'wb') as fp:
# pickle.dump(test_glove, fp)
# + [markdown] id="9CLiNV0XfAde"
# #### Convert input vectors to torch tensors
# + id="zR59A_DfrZXE"
#covert numpy arrays to torch tensors
train_glove, test_glove = map(torch.tensor, (train_glove, test_glove))
# covert tensors data to float
train_glove, test_glove = train_glove.float(), test_glove.float()
# + [markdown] id="YhSyXl904xb_"
# ## Feed-Forward Neural Networks
# + id="x0KLu_hZeYKx"
import torch.nn as nn
import torch.nn.functional as F
from torch import optim
# + [markdown] id="P3c0KOKtMziW"
# ### Evaluation Functions
# + id="_CE6lI8iYIVE"
# function that creates a dataframe to display a user-given field of the Neural Networks
def display_func(net1, net2, net3, net4, column):
nn_dic = {net1:['-'], net2:['-'], net3:['-'], net4:['-']}
nn_df = pd.DataFrame.from_dict(nn_dic, orient='index', columns=[column])
return nn_df
# function that creates a dataframe to display the Precision, Recall and F1-score of the Neural Networks
def display_metrics(tfidf, glove):
nn_dic = {tfidf:['-','-','-'], glove:['-','-','-']}
nn_df = pd.DataFrame.from_dict(nn_dic, orient='index', columns=['Precision','Recall','F1-Score'])
return nn_df
# function that implements Root Mean Square Error
def RMSELoss(yhat,y):
return torch.sqrt(torch.mean((yhat - y) ** 2))
# + id="HvnVsVT6sjSI"
from sklearn.metrics import roc_curve, auc
from sklearn.metrics import roc_auc_score
# function to plot loss per epoch
def loss_per_epoch(loss, title):
plt.figure(figsize=(8, 5))
plt.style.use('seaborn-whitegrid')
plt.plot(range(len(loss)), loss)
plt.title('Loss - Epoch Diagram (' + title + ')', size=20)
plt.xlim=[1, len(loss)]
plt.xlabel('Epochs', size=20)
plt.yticks(loss, [i for i in loss])
plt.yscale('logit')
plt.ylabel('Loss', size=20)
plt.show()
# function that plots the ROC curve on the test set
def roc_plot(net, test_X, test_y, batch_size, title):
fpr = dict()
tpr = dict()
roc_auc = dict()
test_iter = d2l.load_array((test_X, test_y), batch_size) # split train set in batches
fin_y_pred = []
prec, rec, f1 = 0, 0, 0
for X, y in test_iter:
dim = X.shape[1]
X = X.unsqueeze(1)
X = X.expand(X.shape[0], dim, dim)
# add parameters to device
X = X.to(device)
y = y.to(device)
y_pred = net.predict(X) # execute predict on test set
y_pred = torch.mean(y_pred, dim=1) # calculate mean
y_pred = y_pred.cpu().data.numpy() # convert to numpy
y = y.cpu().data.numpy() # convert to numpy
for i in range(2):
fpr[i], tpr[i], _ = roc_curve(y, y_pred)
roc_auc[i] = auc(fpr[i], tpr[i])
plt.figure(figsize=(8, 5))
plt.style.use('seaborn-whitegrid')
plt.plot(fpr[1], tpr[1])
plt.title('Receiver operating characteristic (' + title + ')', size=20)
plt.xlabel('False Positive Rate', size=20)
plt.ylabel('True Positive Rate', size=20)
plt.show()
roc_score = "%.3f%%" % (roc_auc[i] * 100)
print("\nROC Score = ", roc_score)
# + [markdown] id="fo9Vk5aQZM48"
# ### Neural Network (Basic Edition)
# + id="CWNHg4E84xlF"
class BasicNet(nn.Module):
'''
Basic Edition of our Feed-Forward Neural Net
Structure:
Linear input, hidden and output layers
Use of ReLu activation between every pair of layers for better performance and faster learning of train data
Output layer:
Softmax activation function
'''
def __init__(self, input_dim, hidden_dim, output_dim):
super(BasicNet, self).__init__()
self.l1 = nn.Linear(input_dim, hidden_dim)
self.relu1 = nn.ReLU()
self.l2 = nn.Linear(hidden_dim, hidden_dim)
self.relu2 = nn.ReLU()
self.l3 = nn.Linear(hidden_dim, output_dim)
def forward(self, X):
out = self.l1(X)
out = self.relu1(out)
out = self.l2(out)
out = self.relu2(out)
out = self.l3(out)
return F.softmax(out, dim=1)
def predict(self, X):
Y_pred = self.forward(X)
return Y_pred
# + id="ydZguWPPZwwo"
# function that trains a Neural Net and returns the loss history
def fit(net, train_X, train_y, epochs=10, learning_rate=0.1, batch_size=1):
train_ls = []
train_iter = d2l.load_array((train_X, train_y), batch_size) # split train set in batches
optimizer = torch.optim.Adagrad(net.parameters(), lr=learning_rate) # AdaGrad Optimizer
for epoch in range(epochs):
epoch_ls = 0
for X, y in train_iter:
criterion = RMSELoss # use RMSE to calucalate the loss
l = criterion(net(X.float()), y.long()) # calculate batch's loss
epoch_ls += l.item() # sum all batches' losses of current epoch
optimizer.zero_grad()
l.backward()
optimizer.step()
mean_ls = epoch_ls / len(train_iter) # calculate the mean loss of the current epoch
print(f'epoch {epoch + 1}, loss {mean_ls:f}')
train_ls.append(mean_ls)
return train_ls
# function that executes the Neural Net training and prints the loss per epoch
def train_net(device, net, train_X, train_y, sample, epochs=10, lr=0.1, batch=1):
# add parameters to device
train_X = train_X.to(device)
train_y = train_y.to(device)
net.to(device)
# execute fit on train set
train_ls = fit(net, train_X[:sample], train_y[:sample], epochs=epochs, learning_rate=lr, batch_size=batch)
print(f'\nFinal loss {train_ls[-1]:f}')
return train_ls
# + id="dKTELKj0eU5i"
from sklearn import metrics
# function that executes prediction on test set
def predict_on_test(device, net, test_X, test_y):
# add parameters to device
test_X = test_X.to(device)
test_y = test_y.to(device)
y_pred = net.predict(test_X) # execute predict on test set
test_y, y_pred = test_y.cpu().data.numpy(), y_pred.cpu().data.numpy() # convert to numpy
prec = "%.3f%%" % (metrics.precision_score(test_y, y_pred) * 100)
rec = "%.3f%%" % (metrics.recall_score(test_y, y_pred) * 100)
f1 = "%.3f%%" % (metrics.f1_score(test_y, y_pred) * 100)
return y_pred, prec, rec, f1
# + [markdown] id="ASbjSirM24cj"
# ### Train & Predict
# + id="pVzvILvo3GCw"
# initialize dataframe to display metrics
bnn_df = display_metrics('BasicNet-tfidf', 'BasicNet-glove')
# + id="LODecKOGVtZs" colab={"base_uri": "https://localhost:8080/"} outputId="aef42eb4-d5f5-4514-a26b-5c6bb95ef44e"
# enable gpu for faster execution
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("Device available for running: ")
print(device)
# + [markdown] id="Be8NXB0gfxoD"
# #### Tf-idf
# + id="ys1OzarFuTst"
# initialize parameters
sample1=train_tfidf.shape[0] # the whole training set
epochs1=5
lr1=0.001
batch1=1024
# initialize Neural Net
net1 = BasicNet(train_tfidf.shape[1], train_tfidf.shape[1], 1)
# + colab={"base_uri": "https://localhost:8080/"} id="zRsY2ZckUrWf" outputId="602cd4e0-270a-482d-8899-5b4196b37233"
# train Neural Net
btrain_tfidf_ls = train_net(device, net1, train_tfidf, train_y, sample1, epochs1, lr1, batch1)
# predict on test set
(bpred_tfidf,
bnn_df.loc['BasicNet-tfidf','Precision'],
bnn_df.loc['BasicNet-tfidf','Recall'],
bnn_df.loc['BasicNet-tfidf','F1-Score']) = predict_on_test(device, net1, test_tfidf, test_y)
# + [markdown] id="HvfmXHhAhvYI"
# **Our first Neural network, BasicNet, is trained with RMSE loss function and uses softmax activation function (adapted for binary classification) in the output layer. Moreover, we choose the number of neurons in the hidden layer to be equal to the number of neurons in the input layer. <br> From our first experiment with BasicNet, using tf-idf input vectors, it's obvious that our model doesn't converge because it's overfitting. This can be easily concluded both from the ups and downs between the pairs of epochs and from the below Loss-Epoch Diagram. <br> This is a problem we will solve with our next Neural Network, UpgradedNet.**
#
# + id="nSf_9I4Dx-5a" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="1c1d6fc1-3225-402d-f29f-e7994610bbae"
loss_per_epoch(btrain_tfidf_ls, 'BasicNet-tfidf')
# + [markdown] id="EUsO_NXygAL2"
# #### GloVe
# + id="iJvtif7wPnd5"
# initialize parameters
sample2=train_glove.shape[0] # the whole training set
epochs2=5
lr2=0.001
batch2=1024
# initialize Neural Net
net2 = BasicNet(train_glove.shape[1], train_glove.shape[1], 1)
# + colab={"base_uri": "https://localhost:8080/"} id="G_hLfGBlsNR1" outputId="7183d308-eda6-4a00-bb29-c0c54f35b197"
# train Neural Net
btrain_glove_ls = train_net(device, net2, train_glove, train_y, sample2, epochs2, lr2, batch2)
# predict on test set
(bpred_glove,
bnn_df.loc['BasicNet-glove','Precision'],
bnn_df.loc['BasicNet-glove','Recall'],
bnn_df.loc['BasicNet-glove','F1-Score']) = predict_on_test(device, net2, test_glove, test_y)
# + [markdown] id="ewbtN2q4mqzm"
# **Our second experiment with BasicNet, using glove embeddings input vectors, produces more less the same results with our first experiment. Our model is overfitting, again. This means that pretrained embeddings can not change the efficiency of a whole model on their own. <br> We are obliged to make changes to BasicNet's layers and activations functions, in order to produce better results.**
# + colab={"base_uri": "https://localhost:8080/", "height": 360} id="rPZd9LuWsNR3" outputId="dae9791a-a4f2-4835-b169-d363c6417c6b"
loss_per_epoch(btrain_glove_ls, 'BasicNet-glove')
# + [markdown] id="qFqzo5-DismJ"
# ### BasicNet Prediction Evaluation
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="MwdbdV51rE3K" outputId="b14dc80b-23a6-4e04-9400-9cc6e70caafd"
bnn_df
# + [markdown] id="_1oDP94voQTe"
# **Our first two experiments with BasicNet didn't provide us the results we would like, in terms of prediction on the test set. Examining the metrics at first glance, we may consider that the F1 Score seems satisfying. That's not entirely true because there is a big imbalance between the Precision and Recall metrics. This imbalance (Low Precision - High Recall) practically means that the Neural Network classifies all the positive sentiments correctly but it also classifies almost half of the negative sentiments as positive. As a result F1 Score gets a value between the values of Precision and Recall. However, this F1 Score value isn't representative of the effectiveness of the classifier, due to the fact that Precision is really bad. Another observation that is not so much expected is the fact that we get the same prediction results for both our models trained with different input vectors.**
#
# + [markdown] id="ir5xbfh2-RaN"
# ### Neural Network (Upgraded Edition)
# + id="rKVUnt9E-RaN"
class UpgradedNet(nn.Module):
'''
Upgraded Edition of our Feed-Forward Neural Net
Structure:
Linear input, hidden and output layers
Use of LeakyReLu activation between every pair of layers to deal with the dying ReLU problem
Output layer:
Sigmoid activation function
'''
def __init__(self, input_dim, hidden_dim, output_dim):
super(UpgradedNet, self).__init__()
self.l1 = nn.Linear(input_dim, hidden_dim)
self.relu1 = nn.LeakyReLU() # switch ReLu activation function with LeakyReLu activation function
self.hd1 = (hidden_dim / 3) * 2 + 1 # calculate the number of output neurons of the first hidden layer
self.l2 = nn.Linear(hidden_dim, int(self.hd1))
self.relu2 = nn.LeakyReLU() # switch ReLu activation function with LeakyReLu activation function
self.hd2 = (int(self.hd1) / 3) * 2 + 1 # calculate the number of output neurons of the second hidden layer
self.l3 = nn.Linear(int(self.hd1), int(self.hd2))
self.relu3 = nn.LeakyReLU() # switch ReLu activation function with LeakyReLu activation function
self.l4 = nn.Linear(int(self.hd2), output_dim)
def forward(self, X):
out = self.l1(X)
out = self.relu1(out)
out = self.l2(out)
out = self.relu2(out)
out = self.l3(out)
out = self.relu3(out)
out = self.l4(out)
return F.sigmoid(out) # replace softmax of BasicNet with sigmoid
def predict(self, X):
Y_pred = self.forward(X)
return Y_pred
# + id="HM6HJA6ZspsR"
# function that trains a Neural Net and returns the loss history
# with manual implementation of Early Stopping and BCELoss loss function
def fit_with_early_stopping(net, train_X, train_y, epochs=10, learning_rate=0.1, batch_size=1, patience=3):
not_improved = 0
early_stop = False
min_ls = np.Inf # initialize minimum loss
train_ls = []
train_iter = d2l.load_array((train_X, train_y), batch_size) # split train set in batches
optimizer = torch.optim.Adagrad(net.parameters(), lr=learning_rate) # AdaGrad Optimizer
for epoch in range(epochs):
epoch_ls = 0
for X, y in train_iter:
criterion = nn.BCELoss() # use BCELoss to calucalate the loss
y = y.unsqueeze(1)
l = criterion(net(X.float()), y) # calculate batch's loss
epoch_ls += l.item() # sum all batches' losses of current epoch
optimizer.zero_grad()
l.backward()
optimizer.step()
mean_ls = epoch_ls / len(train_iter) # calculate the mean loss of the current epoch
mean_ls = round(mean_ls, 6) # round up the loss to the 6th decimal digit
if mean_ls < min_ls: # store minimum loss
not_improved = 0
min_ls = mean_ls
else:
not_improved += 1 # raise not_improved counter
if (epoch + 1) >= patience and not_improved == patience: # apply early stopping
print("Early stopping! (patience = " + str(patience) + ")")
early_stop = True
break
print(f'epoch {epoch + 1}, loss {mean_ls:f}')
train_ls.append(mean_ls)
return train_ls
# function that executes the Neural Net training and prints the loss per epoch, applying Early Stopping
def train_net_with_early_stopping(device, net, train_X, train_y, sample, epochs=10, lr=0.1, batch=1, patience=3):
# add parameters to device
train_X = train_X.to(device)
train_y = train_y.to(device)
net.to(device)
# execute fit with Early Stopping on train set
train_ls = fit_with_early_stopping(net, train_X[:sample], train_y[:sample], epochs=epochs, learning_rate=lr, batch_size=batch, patience=patience)
print(f'\nFinal loss {train_ls[-1]:f}')
return train_ls
# + id="OZu7Nyttmh6x"
# function that executes prediction on test set using sigmoid activation function on output layer
def predict_on_test_with_sigmoid(device, net, test_X, test_y):
# add parameters to device
test_X = test_X.to(device)
test_y = test_y.to(device)
y_pred = net.predict(test_X) # execute predict on test set
y_pred = y_pred.cpu().data.numpy() # convert to numpy
y_pred = np.where(y_pred >= 0.5, 1, 0) # if prediction is >= 0.5 set it to 1
test_y = test_y.cpu().data.numpy() # convert to numpy
prec = "%.3f%%" % (metrics.precision_score(test_y, y_pred) * 100)
rec = "%.3f%%" % (metrics.recall_score(test_y, y_pred) * 100)
f1 = "%.3f%%" % (metrics.f1_score(test_y, y_pred) * 100)
return y_pred, prec, rec, f1
# + [markdown] id="Q0Dzsc_13Ch8"
# ### Train & Predict
# + id="aqGlRs5_53PW"
# initialize dataframe to display metrics
unn_df = display_metrics('UpgradedNet-tfidf', 'UpgradedNet-glove')
# + colab={"base_uri": "https://localhost:8080/"} id="mPZfeCWpwjdn" outputId="a76c5cf7-fd7a-43e5-ef77-3ceabb36b123"
# enable gpu for faster execution
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("Device available for running: ")
print(device)
# + [markdown] id="0hLrkISE-RaN"
# #### Tf-idf
# + id="ZNqpNySA-RaN"
# initialize parameters
sample3=train_tfidf.shape[0] # the whole training set
epochs3=20
lr3=0.005
batch3=1024
patience=3 # number of allowed epochs with worse loss
# initialize Neural Net
net3 = UpgradedNet(train_tfidf.shape[1], train_tfidf.shape[1], 1)
# + colab={"base_uri": "https://localhost:8080/"} id="9nAOzl9S-RaN" outputId="ccd05f20-cf23-46a7-89c1-58d35a7cb372"
# train Neural Net
utrain_tfidf_ls = train_net_with_early_stopping(device, net3, train_tfidf, train_y, sample3, epochs3, lr3, batch3, patience)
# predict on test set
(upred_tfidf,
unn_df.loc['UpgradedNet-tfidf','Precision'],
unn_df.loc['UpgradedNet-tfidf','Recall'],
unn_df.loc['UpgradedNet-tfidf','F1-Score']) = predict_on_test_with_sigmoid(device, net3, test_tfidf, test_y)
# + [markdown] id="2oqXd6vtzs4O"
# **We upgrade our first Neural network, BasicNet, into UpgradedNet. This time, we train UpgradedNet with BCELoss loss function and we use sigmoid activation function (appropriate for binary classification by default) in the output layer. We, also replace the ReLu activation functions with LeakyReLu ones. The reason we apply this change is related to the prediction results on the test set and we will analyze it below. Moreover, we choose the number of neurons in each hidden layer to be 2/3 the size of the neurons of the previous layer, plus the size of the output layer. The best learning rate that occured after experementation is 0.005, that's why we use it in both our UpgradedNet models. Finally, trying to avoid overfitting we train UpgradedNet with the Early Stopping technique, with patience=3 and after experementation it is indeed applied for a big number of epochs ( > 100). <br> In the above experiment we choose to train the model for 20 epochs to reduce time execution. Besides, the model already converges just fine and the loss is now reduced by more than 0.1 compared to the relative experiment we conducted with the BasicNet model. After executing the above experiment we can, in fact, confirm the bad effect of RMSE loss function in binary classification models. We know that MSE is not guaranteed to minimize the loss function, because MSE function expects real-valued inputs in range (-∞, ∞), while binary classification models output probabilities in range(0,1) through the sigmoid function. In a few words, RMSE was the main factor that caused overfitting in BasicNet models. <br> The above results are visualized in the Loss-Epoch Diagram.**
#
# + colab={"base_uri": "https://localhost:8080/", "height": 360} id="dJoFK-NQ-RaO" outputId="1b460ba2-59f3-4d28-cfa9-bdeaffa9d8cc"
loss_per_epoch(utrain_tfidf_ls, 'UpgradedNet-tfidf')
# + [markdown] id="56JssDknTOhi"
# #### GloVe
#
# + id="Ql7_Ph-gTOhi"
# initialize parameters
sample4=train_glove.shape[0] # the whole training set
epochs4=20
lr4=0.005
batch4=1024
patience=3 # number of allowed epochs with worse loss
# initialize Neural Net
net4 = UpgradedNet(train_glove.shape[1], train_glove.shape[1], 1)
# + colab={"base_uri": "https://localhost:8080/"} id="ShzF80JgTOhi" outputId="c358ce7e-ebe2-4bff-ac29-d46f92658091"
# train Neural Net
utrain_glove_ls = train_net_with_early_stopping(device, net4, train_glove, train_y, sample4, epochs4, lr4, batch4, patience)
# predict on test set
(upred_glove,
unn_df.loc['UpgradedNet-glove','Precision'],
unn_df.loc['UpgradedNet-glove','Recall'],
unn_df.loc['UpgradedNet-glove','F1-Score']) = predict_on_test_with_sigmoid(device, net4, test_glove, test_y)
# + [markdown] id="fvBkLhcJ7syZ"
# **Our second experiment with UpgradedNet, using glove embeddings input vectors, produces similar results with our previous experiment with UpgradedNet, concerning the loss. We train our model using the Early Stopping technique, with patience=3 and Early Stopping is again applied for a big number of epochs. In this experiment we also manage to avoid overfitting and we observe an even better loss reduction, since the loss is almost 0.2 lower compared to the relative loss of BasicNet model. The above results are visualized in the Loss-Epoch Diagram.**
#
# + colab={"base_uri": "https://localhost:8080/", "height": 360} id="pEhg5lfOTOhj" outputId="d330c827-e137-47f1-d679-15841c8ea70e"
loss_per_epoch(utrain_glove_ls, 'UpgradedNet-glove')
# + [markdown] id="vY8CqNrhi1iQ"
# ### UpgradedNet Prediction Evaluation
# + id="qVaL1mG3gDYa" colab={"base_uri": "https://localhost:8080/", "height": 111} outputId="12a32540-9746-4d89-d1c0-88a7331cc2d5"
unn_df
# + [markdown] id="bo0E9KpjzyTr"
# **Our two experiments with UpgradedNet helped us optimize the results concerning the loss convergence. Specifically, we managed to stop overfitting and we reduced the loss by almost 15% in the first UpgradedNet model and by 30% in the second. Let's see what happened with the prediction on the test set. <br> At first glance, there is a significant improvement in the prediction results compared to the corresponding ones of our BasicNet models. After experementation, we concluded that the main reason that led to this improvement of the prediction results, is the use of the sigmoid function in the output layer combined with the changes we made in the hidden layers architecture. Let's break it down. <br> First of all, the switching of the output activation function from softmax to sigmoid had a big impact in loss reduction. More precisely, by using the sigmoid function the loss automatically dropped by 0.5 and the models began to converge normally. Another important factor, was the hidden layers architecture. The way we chose to gradually reduce the neurons layer by layer, led not only in faster loss convergence but also raised the prediction metrics by almost 4%. Continuing the analysis on model's architecture, the use of LeakyReLu activation function instead of ReLu had a small but significant effect in the metrics' values. We came up using LeakyReLu after experimenting with all respective activation functions like ReLu, ELU and SELU. LeakyReLu produced the best metrics among all of them by 2%. LeakyReLu turned out to have a good loss convergence even though it didn't converge faster than ReLu. ELU was a step behind concerning efficiency in both loss convergence and prediction metrics. Finally, SELU had a terrible convergence compared to the other 3 activation functions. <br> On the other hand, adding a dropout layer didn't have a good effect on the model. We tried putting a dropout layer in different positions of our Neural Network but in no case it had a positive impact. Specifically, the model was converging much slower and the metrics were similar with the ones of BasicNet. <br> Returning to the positive factors of prediction's improvment we should notice the importance of the size of input vectors. For example, at first we used tf-idf input vectors of size 100. The prediction results we got were worse than the BasicNet ones! Once, we changed the input size to 150 we noticed a huge raise of almost 10% on all metrics, that eventually led to the prediction results we show on the above dataframes. Last but not least, we should mention the importance of picking the correct optimizer. We already knew that theoretically AdaGrad is usually a good optimizer for binary classification problems. In order to be sure, we also, experimented with Stochastic Gradient Descent optimizer and the prediction results were very close to AdaGrad's, but the converge was much slower. Finally, we tried Adam optimizer as well and we observed a fast convergence. Yet, the prediction on the test set was much worse, since all metrics were almost 10% lower than metrics we got using AdaGrad.**
#
# + [markdown] id="6R7VR_H9Mziy"
# ## Final Evaluation
# + [markdown] id="TwJ5hBlKDB7n"
# #### Sort models from best to worst based on lowest Final Loss
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="tTYg0HJPB9Ba" outputId="49010bbc-37e1-4054-9ff3-74a5df90a0b3"
# initialize a dataframe to display loss
ls_df = display_func('BasicNet-tfidf', 'BasicNet-glove', 'UpgradedNet-tfidf', 'UpgradedNet-glove', 'Final-Loss')
# store each model's final loss
ls_df.loc['BasicNet-tfidf','Final-Loss'] = btrain_tfidf_ls[-1]
ls_df.loc['BasicNet-glove','Final-Loss'] = btrain_glove_ls[-1]
ls_df.loc['UpgradedNet-tfidf','Final-Loss'] = utrain_tfidf_ls[-1]
ls_df.loc['UpgradedNet-glove','Final-Loss'] = utrain_glove_ls[-1]
# sort Neural Nets based on Final-Loss column
sort_ls_df = ls_df.sort_values(by=['Final-Loss'], ascending=True)
sort_ls_df
# + [markdown] id="TyPExtEwAdQJ"
# #### Sort models from best to worst based on highest F1 Score
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="BWsBiUP3ASth" outputId="3fd099f8-76c8-4e6c-8b33-62ae24086e67"
#concatenate dataframes' rows to create a new dataframe
nn_df = pd.concat([bnn_df, unn_df])
#sort Neural Nets based on F1-Score
sort_nn_df = nn_df.sort_values(by=['F1-Score'], ascending=False)
sort_nn_df
# + [markdown] id="Tn4w-NcnEzDa"
# #### Dominant Model
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="it8WkDaVE9Z2" outputId="aeb98fa0-070b-4778-cd1e-d5431fdca576"
sort_ls_df.head(2)
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="p_Xtr6xmFIR2" outputId="ba569053-aaff-4e26-fd8b-be54bc2fc16a"
sort_nn_df.head(2)
# + [markdown] id="-bop2sMGGpZ_"
# **There is no doubt that our UpgradedNet Neural Network is far better than the BasicNet one. UpgradedNet is dominating BasicNet in every way. Not only it dealt with the overfitting problem and reduced the loss by almost 30% but it also led to a very trustworthy prediction on the test set. As we can see, there is a great balance between Pricison and Recall metrics, specially in the model, where glove embeddings are used. This balance leads to an equally great F1 Score, which is a determinant representative of a model's efficiency. <br> We chose to implement two totally contradictory models, in order to gradually present the changes we should make to reach a very satisfying result, as well as to emphasize the bad choices someone must avoid, so as not to be led to a frustrating result.**
# + [markdown] id="1of_QkvmF7Rr"
# ### ROC Curve
# + [markdown] id="Jn55nlXEoipb"
# **The Auc Score of UpgradedNet Neural Network, using pre-trained glove embeddings input vectors is ~80% and it's a decent score. As a matter of fact, the Auc Score is far better from the relative scores of the two BasicNet models and much better than the score of UpgradedNet, using tf-idf transformations as input vectors. The high Auc Score means that the model has a good ability at measuring how often a sentiment rating is correct. We should be happy with our result, specially if we consider that Feed-Forward Neural Networks are, usually, good when classification is determined by a long range semantic dependency and they are not so effective at extracting local and position-invariant features. For tasks like Sentiment Analysis, where feature detection in text is more important (searching for angry terms, irony, sadness etc.), a Convolution Neural Network would surely produce the best results possible. <br> The Auc Score is visualized in the below ROC diagram, where the outcome is almost a perfect curve.**
# + id="0-ZqQmixF7gn" colab={"base_uri": "https://localhost:8080/", "height": 394} outputId="cbcf5905-ee55-4dc6-b6ff-7c6347681a75"
# display the best ROC diagram based on Auc Score
roc_plot(net4, test_glove, test_y, batch4, 'UpgradedNet-glove')
| feed_forward_neural_net/sentiment_analysis_ffnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # NLTK : présentation
# + [markdown] slideshow={"slide_type": "-"}
# **NLTK :** *Natural Language ToolKit*
#
# Librairie Python pour le traitement et l’analyse automatique du texte
#
# **Documentation :** [http://www.nltk.org/](http://www.nltk.org/)
# + [markdown] slideshow={"slide_type": "subslide"}
# Une fois la librairie installée, il faut l’importer :
# + slideshow={"slide_type": "-"}
import nltk
# + [markdown] slideshow={"slide_type": "fragment"}
# Puis installer les outils nécessaires (corpus, grammaires…) :
# + slideshow={"slide_type": "-"}
# nltk.download()
# + [markdown] slideshow={"slide_type": "subslide"}
# Exemple d’accès au corpus de textes [Gutenberg](http://www.gutenberg.org/) :
# + slideshow={"slide_type": "-"}
from nltk.corpus import gutenberg
# + [markdown] slideshow={"slide_type": "fragment"}
# Quels sont les textes disponibles ?
# + slideshow={"slide_type": "-"}
print(gutenberg.fileids())
# + [markdown] slideshow={"slide_type": "fragment"}
# Combien de mots compte chaque texte ?
# + slideshow={"slide_type": "-"}
words = [len(gutenberg.words(text)) for text in gutenberg.fileids()]
print(words)
# + [markdown] slideshow={"slide_type": "subslide"}
# Accéder au texte brut :
# + slideshow={"slide_type": "-"}
raw = gutenberg.raw('austen-emma.txt')
# + [markdown] slideshow={"slide_type": "fragment"}
# À la liste des mots :
# + slideshow={"slide_type": "-"}
words = gutenberg.words('austen-emma.txt')
# + [markdown] slideshow={"slide_type": "fragment"}
# À la liste des phrases, découpées en liste de mots :
# + slideshow={"slide_type": "-"}
sents_words = gutenberg.sents('austen-emma.txt')
# -
# À la liste des paragraphes, découpés en listes de phrases elles-mêmes découpées en listes de mots :
paras_words = gutenberg.paras('austen-emma.txt')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Importer un corpus textuel
#
# Les textes précédemment chargés sont des instances d’une classe `PlaintextCorpusReader` qui fournit les méthodes `raw()`, `words()` et `sents()` sur chacun des textes du corpus.
# + [markdown] slideshow={"slide_type": "subslide"}
# Pour bénéficier de ces méthodes, il suffit de créer une instance de cette classe :
# + slideshow={"slide_type": "-"}
from nltk.corpus import PlaintextCorpusReader
corpus = PlaintextCorpusReader('./data', '.*', encoding='utf8')
# + [markdown] slideshow={"slide_type": "subslide"}
# Accéder aux identifiants, au texte brut… :
# + slideshow={"slide_type": "-"}
print(corpus.fileids())
# + slideshow={"slide_type": "fragment"}
print(len(corpus.words('salammbo.txt')))
# + slideshow={"slide_type": "fragment"}
print(corpus.raw('dormeur-du-val.txt'))
# + [markdown] slideshow={"slide_type": "fragment"}
# Et pour exclure par exemple le sous-dossier *allocine* :
# + slideshow={"slide_type": "-"}
# A filename with a "/" in it will be ignored
corpus = PlaintextCorpusReader('./data', '[^\/]*', encoding='utf8')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Importer un corpus catégorisé
#
# Si un corpus est organisé en sous-dossiers, peu importe le système de classement (par thème, par année…), la classe `CategorizedPlaintextCorpusReader` permet d’effectuer une importation de tout le corpus :
# + slideshow={"slide_type": "-"}
from nltk.corpus import CategorizedPlaintextCorpusReader
corpus = CategorizedPlaintextCorpusReader('./data/allocine', '.*', encoding='utf8', cat_pattern = r'(.*)[/]')
# + [markdown] slideshow={"slide_type": "subslide"}
# En plus des méthodes héritées de `PlaintextCorpusReader`, une méthode `categories()` permet de refléter l’organisation du corpus :
# + slideshow={"slide_type": "-"}
print([cat for cat in corpus.categories()])
| 4.text-processing/0.nltk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Lab 7: Qubit Spectroscopy
# In this lab, you will take what you learned about the interactions between qubits and resonators to perform transmon spectroscopy with the pulse simulator.
# ### Installing Necessary Packages
# Before we begin, you will need to install some prerequisites into your environment. Run the cell below to complete these installations. At the end, the cell outputs will be cleared.
# +
# !pip install -U -r grading_tools/requirements.txt
from IPython.display import clear_output
clear_output()
# -
# ## Simulating the Transmon as a Duffing Oscillator
# As you learned in Lecture 6, the transmon can be understood as a Duffing oscillator specified by a frequency $\nu$, anharmonicity $\alpha$, and drive strength $r$, which results in the Hamiltonian
# $$
# \hat{H}_{\rm Duff}/\hbar = 2\pi\nu a^\dagger a + \pi \alpha a^\dagger a(a^\dagger a - 1) + 2 \pi r (a + a^\dagger) \times D(t),
# $$
#
# where $D(t)$ is the signal on the drive channel for the qubit, and $a^\dagger$ and $a$ are, respectively, the creation and annihilation operators for the qubit. Note that the drive strength $r$ sets the scaling of the control term, with $D(t)$ assumed to be a complex and unitless number satisfying $|D(t)| \leq 1$.
# ## Qiskit Pulse Overview
#
# As a brief overview, Qiskit Pulse schedules (experiments) consist of Instructions (i.e., Play) acting on Channels (i.e., the drive channel). Here is a summary table of available Instructions and Channels:
#
# 
#
# For more detail, this table summarizes the interaction of the channels with the actual quantum hardware:
#
# 
#
# However, we find it is more instructive to begin with guided programming in Pulse. Below you will learn how to create pulses, schedules, and run experiments on a simulator. These lessons can be immediately applied to actual pulse-enabled quantum hardware, in particular [`ibmq_armonk`](https://www.ibm.com/blogs/research/2019/12/qiskit-openpulse/).
# ## Let's get started!
# In most of the cells below, nothing needs to be modified. **However, you will need to execute the cells by pressing `shift+Enter` in each code block**. In order to keep things tidy and focus on the important aspects of Qiskit Pulse, the following cells make use of methods from the `helper` module. For the gory details, please refer back to the [Lab 7 notebook](lab7-jc-spect-readout.ipynb). Just as in Lab 6, before coming to the discussion of **Sideband Modulation**, the following code blocks
#
# - create backend pulse simulator and instantiate the transmon as a Duffing oscillator of frequency $\sim 5$ GHz
# - import libraries for numerics and visualization, and define helpful constants
# - create the channels for the pulse schedule and define measurment schedule (we will only work with the drive channel)
# +
# our backend is the Pulse Simulator
from resources import helper
from qiskit.providers.aer import PulseSimulator
backend_sim = PulseSimulator()
# sample duration for pulse instructions
dt = 1e-9
# create the model
duffing_model = helper.get_transmon(dt)
# get qubit frequency from Duffing model
qubit_lo_freq = duffing_model.hamiltonian.get_qubit_lo_from_drift()
# +
import numpy as np
# visualization tools
import matplotlib.pyplot as plt
plt.style.use('dark_background')
# unit conversion factors -> all backend properties returned in SI (Hz, sec, etc)
GHz = 1.0e9 # Gigahertz
MHz = 1.0e6 # Megahertz
kHz = 1.0e3 # kilohertz
us = 1.0e-6 # microseconds
ns = 1.0e-9 # nanoseconds
# -
# ### Instantiate channels and create measurement schedule
#
# We will use the same measurement schedule throughout, whereas the drive schedules will vary. This must be built for the simulator, for a real backend we can ask for its default measurement pulse.
# +
from qiskit import pulse
from qiskit.pulse import Play, Acquire
from qiskit.pulse.pulse_lib import GaussianSquare
# qubit to be used throughout the notebook
qubit = 0
### Collect the necessary channels
drive_chan = pulse.DriveChannel(qubit)
meas_chan = pulse.MeasureChannel(qubit)
acq_chan = pulse.AcquireChannel(qubit)
# Construct a measurement schedule and add it to an InstructionScheduleMap
meas_samples = 1200
meas_pulse = GaussianSquare(duration=meas_samples, amp=0.025, sigma=4, width=1150)
measure_sched = Play(meas_pulse, meas_chan) | Acquire(meas_samples, acq_chan, pulse.MemorySlot(qubit))
inst_map = pulse.InstructionScheduleMap()
inst_map.add('measure', [qubit], measure_sched)
# save the measurement/acquire pulse for later
measure = inst_map.get('measure', qubits=[qubit])
# -
# ## Sideband Modulation
#
# Unlike the case of running on an actual device, with the simulator we can only set the (local) oscillator frequency of the drive, $f_{\rm LO}$, to a single value. In order to sweep frequencies to perform spectroscopy, we use a trick called *sideband modulation*, where we modulate our spectroscopy pulse by a sideband frequency $f_{\rm SB}$ so that the pulse applied to the qubit is of (radio) frequency
#
# $$ f_{\rm RF} = f_{\rm LO} + f_{\rm SB}. $$
#
# This is achieved by multiplying each sample amplitude by a complex exponential
#
# $$ d_j^{\rm SB} = \sum_j e^{2\pi f_{\rm SB} t_j} d_j $$
#
# but we will tuck the details away in the `helper` module. The important thing is that we must apply the sideband for each pulse in order to change its frequency.
# Now, instead of `assemble`'ing a single schedule with an array of schedule LO's as, we will create a schedule of the same pulse *sidebanded* by an array of sideband frequecies at a fixed LO frequency. Since we are now considering a transmon, we have multiple energy levels we can perform spectroscopy on. We will being with spectroscopy of the $|0\rangle \to |1\rangle$ transition, which is the one used as the qubit, often called the *computational basis*.
# +
from qiskit.pulse import pulse_lib
# the same spect pulse used in every schedule
drive_amp = 0.9
drive_sigma = 16
drive_duration = 128
spec_pulse = pulse_lib.gaussian(duration=drive_duration, amp=drive_amp,
sigma=drive_sigma, name=f"Spec drive amplitude = {drive_amp}")
# Construct an np array of the frequencies for our experiment
spec_freqs_GHz = np.arange(5.0, 5.2, 0.005)
# Create the base schedule
# Start with drive pulse acting on the drive channel
spec_schedules = []
for freq in spec_freqs_GHz:
sb_spec_pulse = helper.apply_sideband(spec_pulse, qubit_lo_freq[0]-freq*GHz, dt)
spec_schedule = pulse.Schedule(name='SB Frequency = {}'.format(freq))
spec_schedule += Play(sb_spec_pulse, drive_chan)
# The left shift `<<` is special syntax meaning to shift the start time of the schedule by some duration
spec_schedule += measure << spec_schedule.duration
spec_schedules.append(spec_schedule)
# -
spec_schedules[0].draw()
# +
from qiskit import assemble
# assemble the schedules into a Qobj
spec01_qobj = assemble(**helper.get_params('spec01', globals()))
# -
# run the simulation
spec01_result = backend_sim.run(spec01_qobj, duffing_model).result()
# retrieve the data from the experiment
spec01_values = helper.get_values_from_result(spec01_result, qubit)
# We will fit the spectroscopy signal to a Lorentzian function of the form
#
# $$ \frac{AB}{\pi[(f-f_{01})^2 + B^2]} + C $$
#
# to find the qubit frequency $f_{01}$.
# +
fit_params, y_fit = helper.fit_lorentzian(spec_freqs_GHz, spec01_values, [5, 5, 1, 0])
f01 = fit_params[1]
plt.scatter(spec_freqs_GHz, np.real(spec01_values), color='white') # plot real part of sweep values
plt.plot(spec_freqs_GHz, y_fit, color='red')
plt.xlim([min(spec_freqs_GHz), max(spec_freqs_GHz)])
plt.xlabel("Frequency [GHz]")
plt.ylabel("Measured Signal [a.u.]")
plt.show()
print("01 Spectroscopy yields %f GHz"%f01)
# -
# # Exercise 1: Spectroscopy of 1->2 transition
#
# In order to observe the transition between the $|1\rangle$ and $|2\rangle$ states of the transmon, we must apply an $X_\pi$ pulse to transition the qubit from $|0\rangle$ to $|1\rangle$ first. Because we are using the simulator, we must first define our $X_\pi$ pulse from the Rabi experiment in Lab 6.
# +
x180_amp = 0.629070 #from lab 6 Rabi experiment
x_pulse = pulse_lib.gaussian(duration=drive_duration,
amp=x180_amp,
sigma=drive_sigma,
name='x_pulse')
# -
# The anharmonicity of our transmon qubits is typically around $-300$ MHz, so we will sweep around that value.
# +
anharmonicity_guess_GHz = -0.3
def build_spec12_pulse_schedule(freq):
sb12_spec_pulse = helper.apply_sideband(spec_pulse, (freq + anharmonicity_guess_GHz)*GHz, dt)
### create a 12 spectroscopy pulse schedule spec12_schedule (already done)
### play an x pulse on the drive channel
### play sidebanded spec pulse on the drive channel
### add measurement pulse to schedule
spec12_schedule = pulse.Schedule()
### WRITE YOUR CODE BETWEEN THESE LINES - START
spec12_schedule += Play(x_pulse, drive_chan)
spec12_schedule += Play(sb12_spec_pulse, drive_chan)
spec12_schedule += measure << spec12_schedule.duration
### WRITE YOUR CODE BETWEEN THESE LINES - END
return spec12_schedule
# +
sb_freqs_GHz = np.arange(-.1, .1, 0.005) # sweep +/- 100 MHz around guess
# now vary the sideband frequency for each spec pulse
spec_schedules = []
for freq in sb_freqs_GHz:
spec_schedules.append(build_spec12_pulse_schedule(freq))
# -
spec_schedules[0].draw()
# assemble the schedules into a Qobj
spec12_qobj = assemble(**helper.get_params('spec12', globals()))
answer1 = spec12_qobj
# run the simulation
spec12_result = backend_sim.run(spec12_qobj, duffing_model).result()
# retrieve the data from the experiment
spec12_values = helper.get_values_from_result(spec12_result, qubit)
# We will again fit the spectroscopy signal to a Lorentzian function of the form
#
# $$ \frac{AB}{\pi[(f-f_{12})^2 + B^2]} + C $$
#
# to find the frequency of the $|1\rangle \to |2\rangle$ transition $f_{12}$.
# +
anharm_offset = qubit_lo_freq[0]/GHz + anharmonicity_guess_GHz
fit_params, y_fit = helper.fit_lorentzian(anharm_offset + sb_freqs_GHz, spec12_values, [5, 4.5, .1, 3])
f12 = fit_params[1]
plt.scatter(anharm_offset + sb_freqs_GHz, np.real(spec12_values), color='white') # plot real part of sweep values
plt.plot(anharm_offset + sb_freqs_GHz, y_fit, color='red')
plt.xlim([anharm_offset + min(sb_freqs_GHz), anharm_offset + max(sb_freqs_GHz)])
plt.xlabel("Frequency [GHz]")
plt.ylabel("Measured Signal [a.u.]")
plt.show()
print("12 Spectroscopy yields %f GHz"%f12)
print("Measured transmon anharmonicity is %f MHz"%((f12-f01)*GHz/MHz))
# -
# **Help us improve our educational tools by submitting your code**<br>
# If you would like to help us learn how to improve our educational materials and offerings, you can opt in to send us a copy of your Jupyter notebook. By executing the cell below, you consent to sending us the code in your Jupyter notebook. All of the personal information will be anonymized.
from IPython.display import display, Javascript;display(Javascript('IPython.notebook.save_checkpoint();'));
from grading_tools import send_code;send_code('ex1.ipynb')
# # Additional Resources
#
# - The Qiskit textbook sections that cover this material are
# - [Circuit Quantum Electrodynamics](https://qiskit.org/textbook/ch-quantum-hardware/cQED-JC-SW.html)
# - [Accessing Higher Energy States](https://qiskit.org/textbook/ch-quantum-hardware/accessing_higher_energy_states.html)
#
# - Watch the videos
# - [Quantum Coding with Lauren Capelluto](https://www.youtube.com/watch?v=ZvipHRY-URs)
# - ["Qiskit Pulse: Programming Quantum Computers Through the Cloud with Pulses"](https://www.youtube.com/watch?v=V_as5PufUiU) webinar at CQT by yours truly
| lab7/ex1.ipynb |